code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--NOTEBOOK_HEADER-->
# *This notebook contains material from [PyRosetta](https://RosettaCommons.github.io/PyRosetta.notebooks);
# content is available [on Github](https://github.com/RosettaCommons/PyRosetta.notebooks.git).*
# <!--NAVIGATION-->
# < [Packing & Design](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/06.00-Introduction-to-Packing-and-Design.ipynb) | [Contents](toc.ipynb) | [Index](index.ipynb) | [Packing and Relax](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/06.02-Packing-design-and-regional-relax.ipynb) ><p><a href="https://colab.research.google.com/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/06.01-Side-Chain-Conformations-and-Dunbrack-Energies.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a>
# # Side Chain Conformations and Dunbrack Energies
# Keywords: phi(), psi(), energies(), residue_total_energies()
#
# # References
# 1. Original Rotamer paper
# 2. New rotamer paper
#
# # Recommended Resources
# 1. Rotamer Library Homepage: http://dunbrack.fccc.edu/bbdep2010/
# 2. Rotamer Library Interactive Overview: http://dunbrack.fccc.edu/bbdep2010/ImagesMovies.php
# +
# Notebook setup
import sys
if 'google.colab' in sys.modules:
# !pip install pyrosettacolabsetup
import pyrosettacolabsetup
pyrosettacolabsetup.mount_pyrosetta_install()
print ("Notebook is set for PyRosetta use in Colab. Have fun!")
from pyrosetta import *
from pyrosetta.teaching import *
init()
# -
# Begin by moving to the directory with your pdb files and loading cetuximab from 1YY8.clean.pdb used in previous workshops.
#
# ```
# # cd google_drive/My\ Drive/student-notebooks/
# pose = pose_from_pdb("1YY8.clean.pdb")
# start_pose = Pose()
# start_pose.assign(pose)
# ```
pose = pose_from_pdb("inputs/1YY8.clean.pdb")
start_pose = Pose()
start_pose.assign(pose)
# __Question:__ What are the φ, ψ, and χ angles of residue K49?
print(pose.residue(49).name())
print("Phi: %.5f\nPsi: %.5f\n" %(pose.phi(49), pose.psi(49)))
print("Chi 1: %.5f\nChi 2: %.5f\nChi 3: %.5f\nChi 4: %.5f" %(pose.chi(1, 49), pose.chi(2, 49), pose.chi(3, 49), pose.chi(4, 49)))
# Score your pose with the standard full-atom score function. What is the energy of K49? Note the Dunbrack energy component (`fa_dun`), which represents the side-chain conformational probability given phi/psi (Ie backbone-dependant).
#
# Any energy can be converted to a probability. Use this energy (E) to calculate the approximate probability of the rotamer (p): p=e^(-E)
# ```
# scorefxn = get_fa_scorefxn()
# scorefxn(pose)
# energies = pose.energies()
# print(energies.residue_total_energies(49))
# print(energies.residue_total_energies(49)[pyrosetta.rosetta.core.scoring.fa_dun])
# ```
# + nbgrader={"grade": true, "grade_id": "cell-f210fdda6b087d77", "locked": false, "points": 0, "schema_version": 3, "solution": true}
### BEGIN SOLUTION
scorefxn = get_fa_scorefxn()
scorefxn(pose)
energies = pose.energies()
print(energies.residue_total_energies(49))
print(energies.residue_total_energies(49)[pyrosetta.rosetta.core.scoring.fa_dun])
### END SOLUTION
# -
# Use `pose.set_chi(<i>, <res_num>, <chi>)` to set the side chain of residue 49 to the all-anti conformation. (Here, `i` is the χ index, and `chi` is the new torsion angle in degrees.) Re-score the pose and note the Dunbrack energy.
# + nbgrader={"grade": true, "grade_id": "cell-74a711eb920136be", "locked": false, "points": 0, "schema_version": 3, "solution": true}
for i in range(1, 5):
pose.set_chi(i, 49, 180)
### BEGIN SOLUTION
scorefxn(pose)
print(energies.residue_total_energies(49))
print(energies.residue_total_energies(49)[pyrosetta.rosetta.core.scoring.fa_dun])
### END SOLUTION
# -
# <!--NAVIGATION-->
# < [Packing & Design](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/06.00-Introduction-to-Packing-and-Design.ipynb) | [Contents](toc.ipynb) | [Index](index.ipynb) | [Packing and Relax](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/06.02-Packing-design-and-regional-relax.ipynb) ><p><a href="https://colab.research.google.com/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/06.01-Side-Chain-Conformations-and-Dunbrack-Energies.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a>
|
notebooks/06.01-Side-Chain-Conformations-and-Dunbrack-Energies.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.3 ('base')
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import statsmodels.api as sm
# %matplotlib inline
df = pd.read_excel(r'D:\Uni\2курс(2021-2022)\Прикладная эконометрика\Econometrics1\DTP.xls')
df.to_csv (r'D:\Uni\2курс(2021-2022)\Прикладная эконометрика\Econometrics1\DTP.csv', index = None, header=True)
df = pd.read_csv(r'D:\Uni\2курс(2021-2022)\Прикладная эконометрика\Econometrics1\DTP.csv')
df.head()
df.describe()
df.columns
# +
y = df['lnDTP']
x = df[['lnCARS','DEV','lnLENTH','lnALC']]
x = sm.add_constant(x)
model1 = sm.OLS(y, x)
result = model1.fit()
result.summary()
# +
y = df['lnDTP']
x = df[['DEV','lnALC']]
x = sm.add_constant(x)
model = sm.OLS(y, x)
result = model.fit()
result.summary()
# +
y = df['lnDTP']
x = df[['lnCARS','lnLENTH']]
x = sm.add_constant(x)
model2 = sm.OLS(y, x)
result = model2.fit()
result.summary()
# -
# Так как теста Рамсея здесь нет, я его провела через Stata.
# Ramsey RESET test using powers of the fitted values of lnDTP
# Ho: model has no omitted variables
# F(3, 136) = 2.15
# Prob > F = 0.0964
# Исходя из таблицы Фишера, гипотеза об отсутствии упущенных переменных подтвердилась.
# Answers: 2 -A, 3 - A, 4 - A, 5 - A
#
|
DTP.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pickle
import numpy as np
import matplotlib.pyplot as plt
# -
"""
'this file check and convert the pose data'
'-----------------'
0'Right Ankle', 3
1'Right Knee', 2
2'Right Hip', 1
3'Left Hip', 4
4'Left Knee', 5
5'Left Ankle', 6
6'Right Wrist', 15
7'Right Elbow', 14
8'Right Shoulder', 13
9'Left Shoulder', 10
10'Left Elbow', 11
11'Left Wrist', 12
12'Neck', 8
13'Top of Head', 9
14'Pelvis)', 0
15'Thorax', 7
16'Spine', mpi3d
17'Jaw', mpi3d
18'Head', mpi3d
mpi3dval: reorder = [14,2,1,0,3,4,5,16,12,18,9,10,11,8,7,6]
"""
# +
import matplotlib.pyplot as plt
def show2Dpose(channels, ax, lcolor="#3498db", rcolor="#e74c3c", add_labels=True):
"""
Visualize a 2d skeleton
Args
channels: 64x1 vector. The pose to plot.
ax: matplotlib axis to draw on
lcolor: color for left part of the body
rcolor: color for right part of the body
add_labels: whether to add coordinate labels
Returns
Nothing. Draws on ax.
"""
vals = np.reshape( channels, (-1, 2) )
#plt.plot(vals[:,0], vals[:,1], 'ro')
I = np.array([0,1,2,0,4,5,0,7,8,8,10,11,8,13,14]) # start points
J = np.array([1,2,3,4,5,6,7,8,9,10,11,12,13,14,15]) # end points
LR = np.array([1,1,1,0,0,0,0, 0, 0, 0, 0, 0, 1, 1, 1], dtype=bool)
# Make connection matrix
for i in np.arange( len(I) ):
x, y = [np.array( [vals[I[i], j], vals[J[i], j]] ) for j in range(2)]
ax.plot(x, -y, lw=2, c=lcolor if LR[i] else rcolor)
RADIUS = 1 # space around the subject
xroot, yroot = vals[0,0], vals[0,1]
ax.set_xlim([-1, 1])
ax.set_ylim([-1, 1])
if add_labels:
ax.set_xlabel("x")
ax.set_ylabel("-y")
ax.set_aspect('equal')
# +
import matplotlib.pyplot as plt
import numpy as np
import os
from mpl_toolkits.mplot3d import Axes3D
def show3Dpose(channels, ax, lcolor="#3498db", rcolor="#e74c3c", add_labels=True,
gt=False,pred=False): # blue, orange
"""
Visualize a 3d skeleton
Args
channels: 96x1 vector. The pose to plot.
ax: matplotlib 3d axis to draw on
lcolor: color for left part of the body
rcolor: color for right part of the body
add_labels: whether to add coordinate labels
Returns
Nothing. Draws on ax.
"""
# assert channels.size == len(data_utils.H36M_NAMES)*3, "channels should have 96 entries, it has %d instead" % channels.size
vals = np.reshape( channels, (16, -1) )
I = np.array([0,1,2,0,4,5,0,7,8,8,10,11,8,13,14]) # start points
J = np.array([1,2,3,4,5,6,7,8,9,10,11,12,13,14,15]) # end points
LR = np.array([1,1,1,0,0,0,0, 0, 0, 0, 0, 0, 0, 1, 1, 1], dtype=bool)
# Make connection matrix
for i in np.arange( len(I) ):
x, y, z = [np.array( [vals[I[i], j], vals[J[i], j]] ) for j in range(3)]
if gt:
ax.plot(x,z, -y, lw=2, c='k')
elif pred:
ax.plot(x,z, -y, lw=2, c='r')
else:
ax.plot(x, z, -y, lw=2, c=lcolor if LR[i] else rcolor)
RADIUS = 1 # space around the subject
xroot, yroot, zroot = vals[0,0], vals[0,1], vals[0,2]
ax.set_xlim3d([-RADIUS+xroot, RADIUS+xroot])
ax.set_ylim3d([-RADIUS+zroot, RADIUS+zroot])
ax.set_zlim3d([-RADIUS-yroot, RADIUS-yroot])
if add_labels:
ax.set_xlabel("x")
ax.set_ylabel("z")
ax.set_zlabel("-y")
# Get rid of the panes (actually, make them white)
white = (1.0, 1.0, 1.0, 0.0)
ax.w_xaxis.set_pane_color(white)
ax.w_yaxis.set_pane_color(white)
# Keep z pane
# Get rid of the lines in 3d
ax.w_xaxis.line.set_color(white)
ax.w_yaxis.line.set_color(white)
ax.w_zaxis.line.set_color(white)
# -
'''deal with mpi3dval'''
# +
# load the download data
mpi3dval_path = './dataset_extras/mpi_inf_3dhp_valid.npz'
mpi_inf_3dhp_valid = np.load(mpi3dval_path)
print(mpi_inf_3dhp_valid.files)
# +
# convert the data to a list to processing.
mpi3d_val_list = []
for i in range(2929):
tmp_dict = {}
tmp_dict['filename'] = mpi_inf_3dhp_valid['imgname'][i]
tmp_dict['kpts2d'] = mpi_inf_3dhp_valid['part'][i]
tmp_dict['kpts3d'] = mpi_inf_3dhp_valid['S'][i]
# prepare the image width for 2D keypoint normalization.
if '/TS5/' in tmp_dict['filename']:
tmp_dict['width'] = 1920
tmp_dict['height'] = 1080
elif '/TS6/' in tmp_dict['filename']:
tmp_dict['width'] = 1920
tmp_dict['height'] = 1080
else:
tmp_dict['width'] = 2048
tmp_dict['height'] = 2048
mpi3d_val_list.append(tmp_dict)
# -
# +
def normalize_screen_coordinates(X,mask, w, h):
assert X.shape[-1] == 2
# Normalize so that [0, w] is mapped to [-1, 1], while preserving the aspect ratio
return (X / w * 2 - [1, h / w] ) * mask
def get_2d_pose_reorderednormed(source):
reorder = [14,2,1,0,3,4,5,16,12,18,9,10,11,8,7,6]
tmp_array = source['kpts2d'][reorder][:, :2]
mask = source['kpts2d'][reorder][:, 2:]
tmp_array1 = normalize_screen_coordinates(tmp_array, mask, source['width'], source['height'])
return tmp_array1, mask
def get_3d_pose_reordered(source):
reorder = [14,2,1,0,3,4,5,16,12,18,9,10,11,8,7,6]
tmp_array = source['kpts3d'][reorder][:, :3]
mask = source['kpts3d'][reorder][:, 3:]
return tmp_array, mask
# +
# convert the pose to 16 joints and put into array
mpi3d_mask_check = []
mpi3d_data_2dpose = []
mpi3d_data_3dpose = []
for source in mpi3d_val_list:
tmp2d, tmp_2dmask = get_2d_pose_reorderednormed(source)
tmp3d, tmp_3dmask = get_3d_pose_reordered(source)
assert np.sum(np.abs(tmp_2dmask - tmp_3dmask))== 0
# mask_check.append(tmp_2dmask)
if not np.sum(tmp_2dmask) == (len(tmp_2dmask)):
mpi3d_mask_check.append(tmp_2dmask)
mpi3d_data_2dpose.append(tmp2d)
mpi3d_data_3dpose.append(tmp3d)
mpi3d_data_2dpose = np.array(mpi3d_data_2dpose)
mpi3d_data_3dpose = np.array(mpi3d_data_3dpose)
# -
len(mpi3d_mask_check)
# +
# save the npz for test purpose
print(mpi3d_data_3dpose.shape)
print(mpi3d_data_2dpose.shape)
np.savez('./test_set/test_3dhp.npz',pose3d=mpi3d_data_3dpose,pose2d=mpi3d_data_2dpose)
# -
# load and check
mpi3d_npz = np.load('./test_set/test_3dhp.npz')
np.concatenate([mpi3d_npz['pose2d']]).shape
# show some pose from self defined idx
idx = 1200
mpi3d_val_list[idx]['filename']
vals2d = mpi3d_data_2dpose[idx]
ax2d = plt.axes()
show2Dpose(vals2d, ax2d)
plt.show()
vals3d = mpi3d_data_3dpose[idx]
fig3d = plt.figure()
# ax = fig.add_subplot(1, 1, 1, projection='3d')
ax3d = Axes3D(fig3d)
show3Dpose(vals3d, ax3d)
plt.show()
|
data_extra/prepare_data_3dhp.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LASP REU Tutorial Day 1 Instructions
#
# ## Write a simple non-interactive script
#
# In a terminal, type the following to open a text editor:
# ```bash
# ~$ gedit &
# ```
#
# Type in a program that
# * prints out the numbers from 10 to 30
# * with a step of 2,
#
# using a `for` loop and the `range` function.
#
# Save the program as "simple_loop.py" into your "Home" directory.
#
# [The terminal might show some `gedit` related errors, ignore them by pressing return and getting a new input prompt.]
#
# Go back to the terminal, and execute your program using:
#
# ```bash
# ~$ python simple_loop.py
# ```
# ## Work with Jupyter notebook
#
# Close gedit.
#
# Launch a jupyter notebook with
# ```bash
# ~$ jupyter notebook &
# ```
# (The `&` symbol means "Launch an independent process and give me back the terminal prompt")
#
# A webbrowser should open. (If asked which one, pick Chromium, which has a good support for Jupyter notebook)
#
# In the upper right, click on the button named "New" and in the dropdown menu, pick the item in the "notebooks" subsection that is most similar to Python3.
#
# ### Write a function to calculate the factorial of a number
#
# There are several ways to implement this.
#
# As a reminder, the factorial of 3 is `1*2*3`, the factorial of 5 is `1*2*3*4*5`.
# You should have learned enough to be able to come up with a combo of some kind of loop, either `for` or `while`.
#
# As a reminder about functions, your skeleton function should look like this:
#
# ```python
# def calc_factorial(n):
# <do the right stuff>
# print("The factorial of",n,"is",result) # if you store the result in `result`
# ```
# ### Dictionaries
#
# The last very important data container in Python that we did not discuss yet are called `dictionaries`.
#
# They work by storing a value with a certain key, so that that key can be used later to get that value back.
#
# Similar to how a list is initialised and recognized via the square brackets `[]`, dictionaries are recognized and initialized by curly brackets `{}`:
mydict = {}
mydict['a'] = 5
mydict['b'] = 10
mydict[4] = 'mystring'
mydict
# Note, that anything that is an unambigously identifiable Python object can be used as a key.
#
# Which is why the only simple certain choices are integers and strings.
# One can loop over the dictionary values and get both key and value at once:
for k, v in mydict.items():
print(k, v)
# Note, that dictionaries don't store things in the order they have received it, for maximum performance (optimzing memory). (there are sorted versions if required).
# ### Task: create function that combines 2 lists into dictionary
#
# Program a function that takes a list of first_names and last_names and stores them into a dictionary.
#
# As you might have guessed, keys have to be unique, but values don't have to.
# So don't try this with a list of first_names that is non-unique (unless you want to learn about error handling. ;)
#
# So, take the list ['john', 'james', 'jacob'] as first_names for example, and the list ['smith', 'baker', 'anderson'] as last_names and give them to the function.
#
# Your function skeleton would look like this:
#
# ```python
# def my_dict_creator(list1, list2):
# <do the right stuff and create mydict>
# return mydict
# ```
|
lasp_reu_python_tutorial_day1_exercises.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload
# %matplotlib inline
import pandas as pd
import numpy as np
from IPython.core.debugger import set_trace
from tqdm import tqdm_notebook
import texcrapy
#from konlpy.corpus import word
from ckonlpy.tag import Twitter, Postprocessor
import json
from soynlp.word import WordExtractor
from soynlp.tokenizer import MaxScoreTokenizer, LTokenizer
from soynlp.noun import LRNounExtractor_v2
from soynlp.pos.tagset import tagset
from soynlp.postagger import Dictionary as Dict
from soynlp.postagger import LRTemplateMatcher
from soynlp.postagger import LREvaluator
from soynlp.postagger import SimpleTagger
from soynlp.postagger import UnknowLRPostprocessor
import nltk
from nltk import Text
from nltk.corpus import stopwords as STOPWORDS
from nltk.corpus import words as WORDS
from nltk.tag import untag
import math
import os
import re
from threading import Thread
#from multiprocessing import Process
import dask
from dask import compute, delayed
import dask.multiprocessing
import dask.bag as db
import jpype
from gensim.models import Word2Vec, FastText
from gensim.corpora import Dictionary
from gensim.models.tfidfmodel import TfidfModel
from gensim.matutils import sparse2full
import seaborn as sns
from matplotlib import rcParams
from matplotlib import pyplot as plt
from sklearn import preprocessing
from sklearn.decomposition import PCA
from sklearn.cluster import KMeans, DBSCAN, AffinityPropagation
from wordcloud import WordCloud
sns.set(style='ticks')
rcParams['font.family'] = u'Malgun Gothic'
# -
nltk.download('stopwords')
nltk.download('words')
# # Scraping
# +
df = pd.read_excel('keywords and logos.xlsx', sheet_name='20190215')[['shortname','kw_supporter','kw_supported','keywords']]; df
_or = lambda kw: ' OR '.join(['#' + k.strip() for k in kw.split(',')])
qry_base = {row.shortname:_or(row.keywords) for row in df.itertuples()}
supporters = df.shortname[df.kw_supporter==True]
qry_sup = ' OR '.join([qry_base[sup] for sup in supporters]); qry_sup
qry = {row.shortname: '(' + qry_base[row.shortname] + ') AND (' + qry_sup + ')' if row.kw_supported==True else qry_base[row.shortname] for row in df.itertuples()}
# -
# %%time
what = ['id', 'timestamp', 'text']
texcrapy.scrap(qry, what=what, lang='ko', end='2019-01-31', download_to='scrapped/twitter')
# # Making Corpus
# +
def preproc(text, remove_url=True, remove_mention=True, remove_hashtag=False):
LINEBREAK = r'\n' # str.replace에서는 r'\n'으로 검색이 안된다
RT = '((?: rt)|(?:^rt))[^ @]?'
EMOJI = r'[\U00010000-\U0010ffff]'
DOTS = '…'
LONG_BLANK = r'[ ]+'
SPECIALS = r'([^ a-zA-Z0-9_\u3131-\u3163\uac00-\ud7a3]+)|([ㄱ-ㅣ]+)'
# \u3131-\u3163\uac00-\ud7a3 는 한글을 의미함
# URL = r'(?P<url>(https?://)?(www[.])?[^ \u3131-\u3163\uac00-\ud7a3]+[.][a-z]{2,6}\b([^ \u3131-\u3163\uac00-\ud7a3]*))'
URL1 = r'(?:https?:\/\/)?(?:www[.])?[^ :\u3131-\u3163\uac00-\ud7a3]+[.][a-z]{2,6}\b(?:[^ \u3131-\u3163\uac00-\ud7a3]*)'
URL2 = r'pic.twitter.com/[a-zA-Z0-9_]+'
URL = '|'.join((URL1, URL2))
HASHTAG = r'#(?P<inner_hashtag>[^ #@]+)'
MENTION = r'@(?P<inner_mention>[^ #@]+)'
text = text.lower()
if remove_url:
text = re.sub(URL, ' ', text)
if remove_mention:
text = re.sub(MENTION, ' ', text)
else:
text = re.sub(MENTION, ' \g<inner_mention>', text)
if remove_hashtag:
text = re.sub(HASHTAG, ' ', text)
else:
text = re.sub(HASHTAG, ' \g<inner_hashtag>', text)
text = re.sub('|'.join((LINEBREAK, RT, EMOJI, DOTS, SPECIALS)), ' ', text)
return re.sub(LONG_BLANK, ' ', text).strip()
class JsonCorpus:
def __init__(self, *fnames, textkey='text'):
self.fnames = fnames
self.textkey = textkey
self.corpus = self._corpus()
def _corpus2(self):
corpus = {}
nfiles = len(self.fnames)
_preproc = lambda doc: preproc(doc[self.textkey])
f_preproc = np.vectorize(_preproc)
for i, fname in enumerate(self.fnames):
with open(fname, encoding='UTF-8-sig') as f:
for item, docs in json.load(f).items():
corpus[item] = f_preproc(docs) if len(docs)>0 else []
pct = '%.2f' % (100 * (i+1) / nfiles)
print('\r {pct}% completed'.format(pct=pct), end='')
print('\n')
return corpus
def _corpus(self):
corpus = {}
nfiles = len(self.fnames)
for i, fname in enumerate(self.fnames):
with open(fname, encoding='UTF-8-sig') as f:
for item, docs in json.load(f).items():
corpus[item] = [preproc(doc[self.textkey]) for doc in docs]
pct = '%.2f' % (100 * (i+1) / nfiles)
print('\r {pct}% completed'.format(pct=pct), end='')
print('\n')
return corpus
def __iter__(self):
for sents in self.corpus.values():
yield from sents
def __len__(self):
return sum([len(sents) for sents in self.corpus.values()])
def tokenize(self, tagger):
return Tokens(tagger, **self.corpus)
class Tokens:
def __init__(self, tagger, **corpora):
self.tagger = tagger
self.tokensdict = self._get_tokens2(**corpora)
def _get_tokens(self, **corpora):
tokens = [] #{}
ths = []
n_items = len(corpora)
for item, corpus in list(corpora.items())[:50]:
th = Thread(target=do_concurrent_tagging, args=(item, corpus, n_items, tokens))
ths.append(th)
for th in ths: th.start()
for th in ths: th.join()
print('\n')
return tokens
def _get_tokens2(self, **corpora):
tokens = {}
for item, corpus in tqdm_notebook(list(corpora.items())[:]):
tokens[item] = [[w[0] for w in self.tagger.tag(corp) if w[1] is not None] for corp in set(corpus)]
return tokens
def freq(self):
return {
item: Text(sum(toks, [])).vocab()
for item, toks in tqdm_notebook(self.tokensdict.items())
if item not in ['ootd','fashion','category']
}
def __iter__(self):
for toks in self.tokensdict.values():
yield from toks
def __len__(self):
return sum([len(toks) for toks in self.tokensdict.values()])
# -
# %%time
fnames = ['scrapped/twitter/' + fname for fname in os.listdir('scrapped/twitter')]
jcorpus = JsonCorpus(*fnames)
# # Making Dictionary
# ### 1. From Scraping keywords
df_keywords = pd.read_excel('keywords and logos.xlsx', sheet_name='20190304')['keywords']
keywords = {w.strip() for w in ', '.join(df_keywords).split(',')}
# ### 2. Soynlp nouns
noun_extractor = LRNounExtractor_v2(verbose=True)
_soynouns = noun_extractor.train_extract(jcorpus, min_noun_score=0.3, min_noun_frequency=5)
soynouns = _soynouns.keys()
soyngrams = {v for k,v in noun_extractor._compounds_components.items() if k in soynouns}
word_extractor = WordExtractor()
word_extractor.train(jcorpus)
_soywords = word_extractor.extract()
# +
def word_score(score):
return score.cohesion_forward * math.exp(score.right_branching_entropy)
soywords = {word for word, score in _soywords.items() if word_score(score)>0.1}
# -
# ### 3. Korean words
# +
with open('dic_system.txt', encoding='UTF-8-sig') as f:
lines = f.readlines()
kowords = {tok.split('\t')[0] for tok in lines}
# -
# ### 4. English words
enwords = set(WORDS.words())
# ### 5. Custom words
# +
cwords = '''
'''
cwords = set(re.findall(r'[^ ,]+', re.sub(r'\n', '', cwords)))
# +
pos_dict = {
'Adverb': {},
'Noun': keywords | soynouns | soywords | kowords | enwords | cwords,
'Josa': {},
'Verb': {},
'Adjective': {},
'Exclamation': {},
}
dictionary = Dict(pos_dict)
generator = LRTemplateMatcher(dictionary)
evaluator = LREvaluator()
postprocessor = UnknowLRPostprocessor()
tagger = SimpleTagger(generator, evaluator, postprocessor)
# -
tagger.tag(jcorpus.corpus['crocs'][250])
# %%time
tokens = jcorpus.tokenize(tagger)
set(jcorpus.corpus['crocs']);
tokens.tokensdict['crocs'];
# # Tagger setup: 현재 이부분은 안쓴다
# ### 1. Passtags
stoptags = {'Foreign','Punctuation','KoreanParticle','Josa','Eomi','PreEomi','Exclamation','Determiner'}
passtags = set(twitter.tagset.keys()) - stoptags
# ### 2. Stopwords
# +
stopwords_en = STOPWORDS.words('english')
with open('stopwords-ko.json', encoding='UTF-8-sig') as f:
stopwords_ko = json.load(f)
stopwords_custom = '''
엔, 줄, 내, 에는, 잖아요, 라는, 데, 는, 것은, 는, 인가요, 란, 말인가, 시, 은, 하네, 이나, 지, 있음, 수, 있는,
있으신가, 요, 그렇다면, 하고, 내는, 후, 생, 가거, 한, 하시겠어요, 만, 되네, 하려면, 하다, 랑, 하게, 근데, pic, 님의,
호, 하세요, 흑, 적인, 다욧, 니, 보니, 건, 다, 위해, 히, 께, 난, 해, 면서, 용, 보자, 가서, 하는, 중, 도,
잉, 인, 울, 하러가기, 인, 준, 든, 이셨으면, 와의,
드는, 되네요, 크흐, 이걸, 서, 에도, 됩니다, 하다고, 날,
하네요, 했습니다, 해지네요, 이였냐, 더,
되니, 하지않을, 아아, htm, 고, 냐고, 이야, 아니야, 너네, 된, 이냐, 해봐라, 해서,
vi,
'''
stopwords_custom = re.findall(r'[^ ,]+', re.sub(r'\n','',stopwords_custom))
stopwords = set(stopwords_en + stopwords_ko)# + stopwords_custom)
# -
# ### 3. N-grams
# +
cngrams = {
('캘빈','클라인'),('켈빈','클라인'),
}
# ('the','boyz'),
# ('어글리','슈즈'),
# ('리미티드','에디션'),
# ('키코','코스타디노브'), ('kiko','kostadinov'),
# ('젤','카야노'),
# ('젤','라이트'),('gel','lyte'),
# ('스튜어트','와이츠먼'),
# ('주니','블랙'),
# ('jun','hyo','seong'),
# ('gel','ptg'),
# ('오니츠카','타이거'),
# ('노박','조코비치'),
# ('후디','티셔츠'),
# ('최종병기','활'),
# ('아이제아','토마스'),
# ('gel','ace')
# ]
ngrams = list(cngrams | soyngrams)
# -
tagger = Postprocessor(twitter)#, stopwords=stopwords, passtags=passtags, ngrams=ngrams)
# %%time
tokens = jcorpus.tokenize(tagger)
def do_concurrent(tagger, item, corpus):
return item, [untag(tagger.tag(corp)) for corp in corpus]
# %%time
if __name__ == '__main__':
values = [delayed(do_concurrent)(tagger, item, corpus) for item, corpus in list(jcorpus.corpus.items())[:5]]
result = compute(*values, scheduler='processes')
# %%time
test = [do_concurrent(tagger, item, corpus) for item, corpus in list(jcorpus.corpus.items())[:5]]
tokensfreq = tokens.freq()
with open('model/tokensfreq.json', 'w', encoding='UTF-8-sig') as f:
json.dump(tokensfreq, f, ensure_ascii=False)
with open('model/tokensfreq.json', encoding='UTF-8-sig') as f:
tokensfreq = json.load(f)
tokensfreq.keys();
# # Word2vec
w2v = Word2Vec(tokens, size=100, window=5, min_count=10, workers=4, sg=1)
w2v.save('model/word2vec.model')
w2v.init_sims(replace=True)
# %%time
w2v = Word2Vec.load('model/word2vec.model')
# +
# blist = pd.read_excel('keywords and logos.xlsx', sheet_name='20190304')[['shortname','keywords']][3:].to_dict('record')
# +
# res = []
# for brand1 in blist:
# for brand2 in blist:
# b1 = [w.strip() for w in brand1['keywords'].split(',')]
# b2 = [w.strip() for w in brand2['keywords'].split(',')]
# try:
# sim = w2v.wv.n_similarity(b1, b2)
# res.append((brand1['shortname']+'-'+brand2['shortname'], sim))
# except:
# _b1 = [w for w in b1 if w in w2v.wv.vocab]
# _b2 = [w for w in b2 if w in w2v.wv.vocab]
# if len(_b1)*len(_b2)!=0:
# sim = w2v.wv.n_similarity(_b1, _b2)
# res.append((brand1['shortname']+'-'+brand2['shortname'], sim))
# +
# sorted(res, key=lambda x:x[1])
# +
# # %%time
# dd = dict(w2v.wv.most_similar(positive=['사랑','가성비','스포티'], topn=1000))
# +
# # %%time
# out = {}
# for bname in tokensfreq.keys():
# try:
# out[bname] = w2v.wv.n_similarity([bname], ['편한','여성스러운'])
# except:
# pass
# out = sorted(out.items(), key=lambda x: x[1])
# -
w2v.wv.most_similar(positive=['머래요'], topn=10)
id_pools = {
'럭셔리': '럭셔리,고급,호화,과시,luxury,esteem',
'캐주얼': '캐주얼,캐쥬얼,casual',
'유니크': '유니크,독특,독창,unique,개성,only,참신,신선,특이,아이디어,철학',
'대중성': '대중,popular,널리,흔한',
'정통성': '정통,traditional,트레디셔널,클래식,classic,품격,약속,신뢰,예측',
'트렌디': '트렌디,트랜디,트렌드,트랜드,유행,trend,trendy,변화,새로운',#,'민감'],
'포멀': '포멀,노멀,normal,평범,일상,무난,기본',
'액티브': '화제,인기,hot,튀는,신상,최신,액티브,active,앞서가는,실험',
# '화제성': '화제,인기,hot,튀는',
#'가성비': ['가성비','저렴','효율','성능','실용'],
#'신뢰성': ['신뢰','믿음','trust','견고','품질','안정','클래식','classic'],
#'활동성': ['활동','활발','운동','액티브','스포츠','active','sport','sports','sporty'],
#'과감함': ['과감','선도','선구','대담','강렬','선명','예술','art'],
# 철학, 환상, 신상, 새로운,
}
# +
df = pd.read_excel('keywords and logos.xlsx', sheet_name='20190304')[['shortname','keywords']]
_kws = lambda kws: [kw.strip() for kw in kws.split(',') if kw.strip() in w2v.wv.vocab]
# _sim = lambda kws: {k:1 for k,v in id_pools.items()}
_sim = lambda kws: {k:w2v.wv.n_similarity(_kws(kws), v.split(',')) for k,v in id_pools.items()}
idty = {}
for row in list(df.itertuples())[3:]:
try:
idty[row.shortname] = _sim(row.keywords)
except:
pass
# -
idty
dct = Dictionary(tokens)
# dct.filter_extremes(no_below=20, no_above=0.2)
dct.compactify()
corp = {item:[dct.doc2bow(tok) for tok in toks] for item, toks in tqdm_notebook(tokens.tokensdict.items())}
model_tfidf = TfidfModel(sum(corp.values(),[]), id2word=dct)
wordlist = list(dct.values())
identities = list(id_pools.keys())
brands = list(tokens.tokensdict.keys())
# +
# sim_mat = np.zeros((len(identities),len(wordlist)))
# for i, idty in enumerate(tqdm_notebook(identities)):
# for j, word in enumerate(wordlist):
# sims = []
# for w in id_pools[idty].split(','):
# try:
# sims.append(w2v.wv.similarity(w, word))
# except:
# pass
# sim_mat[i,j] = np.mean(sims)
# -
len(w2v.wv.vocab), len(wordlist)
# # 여기서 주의할점:
# 여기에서는 각 브랜드의 token을 가지고 similarity를 계산하지만,
# 검색(Search)할때에는 빠른 검색을 위해 각 브랜드명 자체를 가지고 계산함
# --> 이제는 여기에서도 각 브랜드명 자체를 가지고 계산하자!
sim_mat = np.zeros((len(identities),len(wordlist)))
for i, idty in enumerate(tqdm_notebook(identities)):
for j, word in enumerate(wordlist):
if word in w2v.wv.vocab:
sim_mat[i,j] = w2v.wv.n_similarity([word], id_pools[idty].split(','))
else:
sim_mat[i,j] = 0
sim_mat.shape
# +
def get_brand_vecs(bname):
brand_tfidf = model_tfidf[corp[bname]]
return np.vstack([sparse2full(c, len(dct)) for c in brand_tfidf]).mean(axis=0)
def plot_id(bname):
val = sim_mat.dot(get_brand_vecs(bname))
#val /= val.sum()
pd.Series(val, index=identities).plot.barh()
# -
id_dict = {}
for bname in tqdm_notebook(brands):
if bname not in ['ootd','fashion','category']:
try:
id_dict[bname] = sim_mat.dot(get_brand_vecs(bname))
except:
print(bname)
pd.DataFrame(id_dict, index=identities).to_pickle('model/id_dict.pkl')
# df = pd.DataFrame(id_dict, index=identities) #
df = pd.read_pickle('model/id_dict.pkl')
df[['adidas','chanel']].plot.barh(figsize=(5,7))
# +
# df_id = pd.DataFrame(columns=df.columns)
# df_id.loc['럭셔리-캐주얼'] = df.loc['럭셔리'] / (df.loc['럭셔리'] + df.loc['캐주얼'])
# df_id.loc['유니크-대중성'] = df.loc['유니크'] / (df.loc['유니크'] + df.loc['대중성'])
# df_id.loc['정통성-트렌디'] = df.loc['정통성'] / (df.loc['정통성'] + df.loc['트렌디'])
# df_id.loc['포멀-화제성'] = df.loc['포멀'] / (df.loc['포멀'] + df.loc['화제성'])
# +
# df_id.to_pickle('model/id_dict.pkl')
# +
# df_id[['nike','crocs','chanel']].plot.barh(figsize=(8,5))
# -
# # 여기서 normalize 축을 바꾸면 어떨까?
#df = pd.read_pickle('model/id_dict.pkl')
min_max_scaler = preprocessing.MinMaxScaler(feature_range=(0.1, 1))
X_train_minmax = min_max_scaler.fit_transform(df_id)
df2 = df_id.copy()
df2[:] = X_train_minmax
df2[['nike','crocs']].plot.barh(figsize=(5,7))
pca = PCA(n_components=0.9)
X_reduced = pca.fit_transform(df2.T)
# cluster = KMeans(n_clusters=10, random_state=0).fit(X_reduced)
# cluster = DBSCAN().fit(X_reduced)
cluster = AffinityPropagation().fit(X_reduced)
clustered = pd.Series(cluster.labels_, index=df2.columns); clustered
clrd = pd.DataFrame(clustered.sort_values()).reset_index()
clrd.columns = ['bname', 'cluster']
clrd.to_excel('model/clustered.xlsx')
clrd[clrd.bname=='chanel']
clustered.index[clustered==10]
df2.loc['신뢰성']
df_kw = pd.read_excel('keywords and logos.xlsx', sheet_name='20190215')[['shortname','keywords']].set_index('shortname')
df_kw;
def plot_wc(bname):
with open('model/tokensfreq.json', encoding='UTF-8-sig') as f:
j = json.load(f)[bname]
kws = [kw.strip() for kw in df_kw.keywords.loc[bname].split(',')]
[j.pop(kw, None) for kw in kws]
x,y = np.ogrid[:300, :300]
mask = (x-150)**2 + (y-150)**2 > 130**2
mask = 255 * mask.astype(int)
font_path = r'c:\Windows\Fonts\NanumBarunGothic.ttf'
wc = WordCloud(width=100, height=100, background_color='white', font_path=font_path, random_state=0, mask=mask)
try:
fig, ax = plt.subplots(figsize=(7,7))
ax.imshow(wc.generate_from_frequencies(j), interpolation='bilinear')
ax.axis('off')
fig.savefig('model/wordcloud/wc_' + bname + '.png', format='png')
except:
print(bname)
finally:
plt.close(fig)
for row in tqdm_notebook(df_kw.itertuples()):
bname = row.Index
fname = 'model/wordcloud/wc_' + bname + '.png'
if os.path.isfile(fname):
continue
if bname not in ['ootd','fashion','category']:
plot_wc(bname)
df2.T['신뢰성'].sort_values()
|
archive/texcrapy4 2019.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.9 64-bit
# language: python
# name: python36964bitb8e2b9000e0f47189ecd49f9d9b8a2b7
# ---
# + [markdown] tags=["pdf-title"]
# # Convolutional Networks
#
# So far we have worked with deep fully-connected networks, using them to explore different optimization strategies and network architectures. Fully-connected networks are a good testbed for experimentation because they are very computationally efficient, but in practice all state-of-the-art results use convolutional networks instead.
#
# First you will implement several layer types that are used in convolutional networks. You will then use these layers to train a convolutional network on the CIFAR-10 dataset.
# + tags=[]
# As usual, a bit of setup
import numpy as np
import matplotlib.pyplot as plt
from cs231n.classifiers.cnn import *
from cs231n.data_utils import get_CIFAR10_data
from cs231n.gradient_check import eval_numerical_gradient_array, eval_numerical_gradient
from cs231n.layers import *
from cs231n.fast_layers import *
from cs231n.solver import Solver
# %matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# for auto-reloading external modules
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
def rel_error(x, y):
""" returns relative error """
return np.max(np.abs(x - y) / (np.maximum(1e-8, np.abs(x) + np.abs(y))))
# + tags=[]
# Load the (preprocessed) CIFAR10 data.
data = get_CIFAR10_data()
for k, v in data.items():
print('%s: ' % k, v.shape)
# -
# # Convolution: Naive forward pass
# The core of a convolutional network is the convolution operation. In the file `cs231n/layers.py`, implement the forward pass for the convolution layer in the function `conv_forward_naive`.
#
# You don't have to worry too much about efficiency at this point; just write the code in whatever way you find most clear.
#
# You can test your implementation by running the following:
# + tags=[]
x_shape = (2, 3, 4, 4)
w_shape = (3, 3, 4, 4)
x = np.linspace(-0.1, 0.5, num=np.prod(x_shape)).reshape(x_shape)
w = np.linspace(-0.2, 0.3, num=np.prod(w_shape)).reshape(w_shape)
b = np.linspace(-0.1, 0.2, num=3)
conv_param = {'stride': 2, 'pad': 1}
out, _ = conv_forward_naive(x, w, b, conv_param)
correct_out = np.array([[[[-0.08759809, -0.10987781],
[-0.18387192, -0.2109216 ]],
[[ 0.21027089, 0.21661097],
[ 0.22847626, 0.23004637]],
[[ 0.50813986, 0.54309974],
[ 0.64082444, 0.67101435]]],
[[[-0.98053589, -1.03143541],
[-1.19128892, -1.24695841]],
[[ 0.69108355, 0.66880383],
[ 0.59480972, 0.56776003]],
[[ 2.36270298, 2.36904306],
[ 2.38090835, 2.38247847]]]])
# Compare your output to ours; difference should be around e-8
print('Testing conv_forward_naive')
print('difference: ', rel_error(out, correct_out))
# -
# # Aside: Image processing via convolutions
#
# As fun way to both check your implementation and gain a better understanding of the type of operation that convolutional layers can perform, we will set up an input containing two images and manually set up filters that perform common image processing operations (grayscale conversion and edge detection). The convolution forward pass will apply these operations to each of the input images. We can then visualize the results as a sanity check.
# ## Colab Users Only
#
# Please execute the below cell to copy two cat images to the Colab VM.
# +
# Colab users only!
# # %mkdir -p cs231n/notebook_images
# # %cd drive/My\ Drive/$FOLDERNAME/cs231n
# # %cp -r notebook_images/ /content/cs231n/
# # %cd /content/
# + tags=["pdf-ignore-input"]
from imageio import imread
from PIL import Image
kitten = imread('cs231n/notebook_images/kitten.jpg')
puppy = imread('cs231n/notebook_images/puppy.jpg')
# kitten is wide, and puppy is already square
d = kitten.shape[1] - kitten.shape[0]
kitten_cropped = kitten[:, d//2:-d//2, :]
img_size = 200 # Make this smaller if it runs too slow
resized_puppy = np.array(Image.fromarray(puppy).resize((img_size, img_size)))
resized_kitten = np.array(Image.fromarray(kitten_cropped).resize((img_size, img_size)))
x = np.zeros((2, 3, img_size, img_size))
x[0, :, :, :] = resized_puppy.transpose((2, 0, 1))
x[1, :, :, :] = resized_kitten.transpose((2, 0, 1))
# Set up a convolutional weights holding 2 filters, each 3x3
w = np.zeros((2, 3, 3, 3))
# The first filter converts the image to grayscale.
# Set up the red, green, and blue channels of the filter.
w[0, 0, :, :] = [[0, 0, 0], [0, 0.3, 0], [0, 0, 0]]
w[0, 1, :, :] = [[0, 0, 0], [0, 0.6, 0], [0, 0, 0]]
w[0, 2, :, :] = [[0, 0, 0], [0, 0.1, 0], [0, 0, 0]]
# Second filter detects horizontal edges in the blue channel.
w[1, 2, :, :] = [[1, 2, 1], [0, 0, 0], [-1, -2, -1]]
# Vector of biases. We don't need any bias for the grayscale
# filter, but for the edge detection filter we want to add 128
# to each output so that nothing is negative.
b = np.array([0, 128])
# Compute the result of convolving each input in x with each filter in w,
# offsetting by b, and storing the results in out.
out, _ = conv_forward_naive(x, w, b, {'stride': 1, 'pad': 1})
def imshow_no_ax(img, normalize=True):
""" Tiny helper to show images as uint8 and remove axis labels """
if normalize:
img_max, img_min = np.max(img), np.min(img)
img = 255.0 * (img - img_min) / (img_max - img_min)
plt.imshow(img.astype('uint8'))
plt.gca().axis('off')
# Show the original images and the results of the conv operation
plt.subplot(2, 3, 1)
imshow_no_ax(puppy, normalize=False)
plt.title('Original image')
plt.subplot(2, 3, 2)
imshow_no_ax(out[0, 0])
plt.title('Grayscale')
plt.subplot(2, 3, 3)
imshow_no_ax(out[0, 1])
plt.title('Edges')
plt.subplot(2, 3, 4)
imshow_no_ax(kitten_cropped, normalize=False)
plt.subplot(2, 3, 5)
imshow_no_ax(out[1, 0])
plt.subplot(2, 3, 6)
imshow_no_ax(out[1, 1])
plt.show()
# -
# # Convolution: Naive backward pass
# Implement the backward pass for the convolution operation in the function `conv_backward_naive` in the file `cs231n/layers.py`. Again, you don't need to worry too much about computational efficiency.
#
# When you are done, run the following to check your backward pass with a numeric gradient check.
# + tags=[]
np.random.seed(231)
x = np.random.randn(4, 3, 5, 5)
w = np.random.randn(2, 3, 3, 3)
b = np.random.randn(2,)
dout = np.random.randn(4, 2, 5, 5)
conv_param = {'stride': 1, 'pad': 1}
dx_num = eval_numerical_gradient_array(lambda x: conv_forward_naive(x, w, b, conv_param)[0], x, dout)
dw_num = eval_numerical_gradient_array(lambda w: conv_forward_naive(x, w, b, conv_param)[0], w, dout)
db_num = eval_numerical_gradient_array(lambda b: conv_forward_naive(x, w, b, conv_param)[0], b, dout)
out, cache = conv_forward_naive(x, w, b, conv_param)
dx, dw, db = conv_backward_naive(dout, cache)
# Your errors should be around e-8 or less.
print('Testing conv_backward_naive function')
print('dx error: ', rel_error(dx, dx_num))
print('dw error: ', rel_error(dw, dw_num))
print('db error: ', rel_error(db, db_num))
# -
# # Max-Pooling: Naive forward
# Implement the forward pass for the max-pooling operation in the function `max_pool_forward_naive` in the file `cs231n/layers.py`. Again, don't worry too much about computational efficiency.
#
# Check your implementation by running the following:
# + tags=[]
x_shape = (2, 3, 4, 4)
x = np.linspace(-0.3, 0.4, num=np.prod(x_shape)).reshape(x_shape)
pool_param = {'pool_width': 2, 'pool_height': 2, 'stride': 2}
out, _ = max_pool_forward_naive(x, pool_param)
correct_out = np.array([[[[-0.26315789, -0.24842105],
[-0.20421053, -0.18947368]],
[[-0.14526316, -0.13052632],
[-0.08631579, -0.07157895]],
[[-0.02736842, -0.01263158],
[ 0.03157895, 0.04631579]]],
[[[ 0.09052632, 0.10526316],
[ 0.14947368, 0.16421053]],
[[ 0.20842105, 0.22315789],
[ 0.26736842, 0.28210526]],
[[ 0.32631579, 0.34105263],
[ 0.38526316, 0.4 ]]]])
# Compare your output with ours. Difference should be on the order of e-8.
print('Testing max_pool_forward_naive function:')
print('difference: ', rel_error(out, correct_out))
# -
# # Max-Pooling: Naive backward
# Implement the backward pass for the max-pooling operation in the function `max_pool_backward_naive` in the file `cs231n/layers.py`. You don't need to worry about computational efficiency.
#
# Check your implementation with numeric gradient checking by running the following:
# + tags=[]
np.random.seed(231)
x = np.random.randn(3, 2, 8, 8)
dout = np.random.randn(3, 2, 4, 4)
pool_param = {'pool_height': 2, 'pool_width': 2, 'stride': 2}
dx_num = eval_numerical_gradient_array(lambda x: max_pool_forward_naive(x, pool_param)[0], x, dout)
out, cache = max_pool_forward_naive(x, pool_param)
dx = max_pool_backward_naive(dout, cache)
# Your error should be on the order of e-12
print('Testing max_pool_backward_naive function:')
print('dx error: ', rel_error(dx, dx_num))
# -
# # Fast layers
#
# Making convolution and pooling layers fast can be challenging. To spare you the pain, we've provided fast implementations of the forward and backward passes for convolution and pooling layers in the file `cs231n/fast_layers.py`.
# The fast convolution implementation depends on a Cython extension; to compile it either execute the local development cell (option A) if you are developing locally, or the Colab cell (option B) if you are running this assignment in Colab.
#
# ---
#
# **Very Important, Please Read**. For **both** option A and B, you have to **restart** the notebook after compiling the cython extension. In Colab, please save the notebook `File -> Save`, then click `Runtime -> Restart Runtime -> Yes`. This will restart the kernel which means local variables will be lost. Just re-execute the cells from top to bottom and skip the cell below as you only need to run it once for the compilation step.
#
# ---
# ## Option A: Local Development
#
# Go to the cs231n directory and execute the following in your terminal:
#
# ```bash
# python setup.py build_ext --inplace
# ```
# ## Option B: Colab
#
# Execute the cell below only only **ONCE**.
# +
# # %cd drive/My\ Drive/$FOLDERNAME/cs231n/
# # !python setup.py build_ext --inplace
# -
# The API for the fast versions of the convolution and pooling layers is exactly the same as the naive versions that you implemented above: the forward pass receives data, weights, and parameters and produces outputs and a cache object; the backward pass recieves upstream derivatives and the cache object and produces gradients with respect to the data and weights.
#
# **NOTE:** The fast implementation for pooling will only perform optimally if the pooling regions are non-overlapping and tile the input. If these conditions are not met then the fast pooling implementation will not be much faster than the naive implementation.
#
# You can compare the performance of the naive and fast versions of these layers by running the following:
'''
Note: if running the following cell returns the error, "global name 'col2im_6d_cython' is not defined", re-run the cython compilation script above with python3 instead, e.g.
python3 setup.py build_ext --inplace
You may need to install Cython with
pip3 install Cython
'''
# + tags=[]
# Rel errors should be around e-9 or less
from cs231n.fast_layers import conv_forward_fast, conv_backward_fast
from time import time
np.random.seed(231)
x = np.random.randn(100, 3, 31, 31)
w = np.random.randn(25, 3, 3, 3)
b = np.random.randn(25,)
dout = np.random.randn(100, 25, 16, 16)
conv_param = {'stride': 2, 'pad': 1}
t0 = time()
out_naive, cache_naive = conv_forward_naive(x, w, b, conv_param)
t1 = time()
out_fast, cache_fast = conv_forward_fast(x, w, b, conv_param)
t2 = time()
print('Testing conv_forward_fast:')
print('Naive: %fs' % (t1 - t0))
print('Fast: %fs' % (t2 - t1))
print('Speedup: %fx' % ((t1 - t0) / (t2 - t1)))
print('Difference: ', rel_error(out_naive, out_fast))
t0 = time()
dx_naive, dw_naive, db_naive = conv_backward_naive(dout, cache_naive)
t1 = time()
dx_fast, dw_fast, db_fast = conv_backward_fast(dout, cache_fast)
t2 = time()
print('\nTesting conv_backward_fast:')
print('Naive: %fs' % (t1 - t0))
print('Fast: %fs' % (t2 - t1))
print('Speedup: %fx' % ((t1 - t0) / (t2 - t1)))
print('dx difference: ', rel_error(dx_naive, dx_fast))
print('dw difference: ', rel_error(dw_naive, dw_fast))
print('db difference: ', rel_error(db_naive, db_fast))
# + tags=[]
# Relative errors should be close to 0.0
from cs231n.fast_layers import max_pool_forward_fast, max_pool_backward_fast
np.random.seed(231)
x = np.random.randn(100, 3, 32, 32)
dout = np.random.randn(100, 3, 16, 16)
pool_param = {'pool_height': 2, 'pool_width': 2, 'stride': 2}
t0 = time()
out_naive, cache_naive = max_pool_forward_naive(x, pool_param)
t1 = time()
out_fast, cache_fast = max_pool_forward_fast(x, pool_param)
t2 = time()
print('Testing pool_forward_fast:')
print('Naive: %fs' % (t1 - t0))
print('fast: %fs' % (t2 - t1))
print('speedup: %fx' % ((t1 - t0) / (t2 - t1)))
print('difference: ', rel_error(out_naive, out_fast))
t0 = time()
dx_naive = max_pool_backward_naive(dout, cache_naive)
t1 = time()
dx_fast = max_pool_backward_fast(dout, cache_fast)
t2 = time()
print('\nTesting pool_backward_fast:')
print('Naive: %fs' % (t1 - t0))
print('fast: %fs' % (t2 - t1))
print('speedup: %fx' % ((t1 - t0) / (t2 - t1)))
print('dx difference: ', rel_error(dx_naive, dx_fast))
# -
# # Convolutional "sandwich" layers
# Previously we introduced the concept of "sandwich" layers that combine multiple operations into commonly used patterns. In the file `cs231n/layer_utils.py` you will find sandwich layers that implement a few commonly used patterns for convolutional networks. Run the cells below to sanity check they're working.
# + tags=[]
from cs231n.layer_utils import conv_relu_pool_forward, conv_relu_pool_backward
np.random.seed(231)
x = np.random.randn(2, 3, 16, 16)
w = np.random.randn(3, 3, 3, 3)
b = np.random.randn(3,)
dout = np.random.randn(2, 3, 8, 8)
conv_param = {'stride': 1, 'pad': 1}
pool_param = {'pool_height': 2, 'pool_width': 2, 'stride': 2}
out, cache = conv_relu_pool_forward(x, w, b, conv_param, pool_param)
dx, dw, db = conv_relu_pool_backward(dout, cache)
dx_num = eval_numerical_gradient_array(lambda x: conv_relu_pool_forward(x, w, b, conv_param, pool_param)[0], x, dout)
dw_num = eval_numerical_gradient_array(lambda w: conv_relu_pool_forward(x, w, b, conv_param, pool_param)[0], w, dout)
db_num = eval_numerical_gradient_array(lambda b: conv_relu_pool_forward(x, w, b, conv_param, pool_param)[0], b, dout)
# Relative errors should be around e-8 or less
print('Testing conv_relu_pool')
print('dx error: ', rel_error(dx_num, dx))
print('dw error: ', rel_error(dw_num, dw))
print('db error: ', rel_error(db_num, db))
# + tags=[]
from cs231n.layer_utils import conv_relu_forward, conv_relu_backward
np.random.seed(231)
x = np.random.randn(2, 3, 8, 8)
w = np.random.randn(3, 3, 3, 3)
b = np.random.randn(3,)
dout = np.random.randn(2, 3, 8, 8)
conv_param = {'stride': 1, 'pad': 1}
out, cache = conv_relu_forward(x, w, b, conv_param)
dx, dw, db = conv_relu_backward(dout, cache)
dx_num = eval_numerical_gradient_array(lambda x: conv_relu_forward(x, w, b, conv_param)[0], x, dout)
dw_num = eval_numerical_gradient_array(lambda w: conv_relu_forward(x, w, b, conv_param)[0], w, dout)
db_num = eval_numerical_gradient_array(lambda b: conv_relu_forward(x, w, b, conv_param)[0], b, dout)
# Relative errors should be around e-8 or less
print('Testing conv_relu:')
print('dx error: ', rel_error(dx_num, dx))
print('dw error: ', rel_error(dw_num, dw))
print('db error: ', rel_error(db_num, db))
# -
# # Three-layer ConvNet
# Now that you have implemented all the necessary layers, we can put them together into a simple convolutional network.
#
# Open the file `cs231n/classifiers/cnn.py` and complete the implementation of the `ThreeLayerConvNet` class. Remember you can use the fast/sandwich layers (already imported for you) in your implementation. Run the following cells to help you debug:
# ## Sanity check loss
# After you build a new network, one of the first things you should do is sanity check the loss. When we use the softmax loss, we expect the loss for random weights (and no regularization) to be about `log(C)` for `C` classes. When we add regularization the loss should go up slightly.
# + tags=[]
model = ThreeLayerConvNet()
N = 50
X = np.random.randn(N, 3, 32, 32)
y = np.random.randint(10, size=N)
loss, grads = model.loss(X, y)
print('Initial loss (no regularization): ', loss)
model.reg = 0.5
loss, grads = model.loss(X, y)
print('Initial loss (with regularization): ', loss)
# -
# ## Gradient check
# After the loss looks reasonable, use numeric gradient checking to make sure that your backward pass is correct. When you use numeric gradient checking you should use a small amount of artifical data and a small number of neurons at each layer. Note: correct implementations may still have relative errors up to the order of e-2.
# + tags=[]
num_inputs = 2
input_dim = (3, 16, 16)
reg = 0.0
num_classes = 10
np.random.seed(231)
X = np.random.randn(num_inputs, *input_dim)
y = np.random.randint(num_classes, size=num_inputs)
model = ThreeLayerConvNet(num_filters=3, filter_size=3,
input_dim=input_dim, hidden_dim=7,
dtype=np.float64)
loss, grads = model.loss(X, y)
# Errors should be small, but correct implementations may have
# relative errors up to the order of e-2
for param_name in sorted(grads):
f = lambda _: model.loss(X, y)[0]
param_grad_num = eval_numerical_gradient(f, model.params[param_name], verbose=False, h=1e-6)
e = rel_error(param_grad_num, grads[param_name])
print('%s max relative error: %e' % (param_name, rel_error(param_grad_num, grads[param_name])))
# -
# ## Overfit small data
# A nice trick is to train your model with just a few training samples. You should be able to overfit small datasets, which will result in very high training accuracy and comparatively low validation accuracy.
# + tags=[]
np.random.seed(231)
num_train = 100
small_data = {
'X_train': data['X_train'][:num_train],
'y_train': data['y_train'][:num_train],
'X_val': data['X_val'],
'y_val': data['y_val'],
}
model = ThreeLayerConvNet(weight_scale=1e-2)
solver = Solver(model, small_data,
num_epochs=15, batch_size=50,
update_rule='adam',
optim_config={
'learning_rate': 1e-3,
},
verbose=True, print_every=1)
solver.train()
# + id="small_data_train_accuracy" tags=[]
# Print final training accuracy
print(
"Small data training accuracy:",
solver.check_accuracy(small_data['X_train'], small_data['y_train'])
)
# + id="small_data_validation_accuracy" tags=[]
# Print final validation accuracy
print(
"Small data validation accuracy:",
solver.check_accuracy(small_data['X_val'], small_data['y_val'])
)
# -
# Plotting the loss, training accuracy, and validation accuracy should show clear overfitting:
# +
plt.subplot(2, 1, 1)
plt.plot(solver.loss_history, 'o')
plt.xlabel('iteration')
plt.ylabel('loss')
plt.subplot(2, 1, 2)
plt.plot(solver.train_acc_history, '-o')
plt.plot(solver.val_acc_history, '-o')
plt.legend(['train', 'val'], loc='upper left')
plt.xlabel('epoch')
plt.ylabel('accuracy')
plt.show()
# -
# ## Train the net
# By training the three-layer convolutional network for one epoch, you should achieve greater than 40% accuracy on the training set:
# + tags=[]
model = ThreeLayerConvNet(weight_scale=0.001, hidden_dim=500, reg=0.001)
solver = Solver(model, data,
num_epochs=1, batch_size=50,
update_rule='adam',
optim_config={
'learning_rate': 1e-3,
},
verbose=True, print_every=20)
solver.train()
# + id="full_data_train_accuracy" tags=[]
# Print final training accuracy
print(
"Full data training accuracy:",
solver.check_accuracy(small_data['X_train'], small_data['y_train'])
)
# + id="full_data_validation_accuracy" tags=[]
# Print final validation accuracy
print(
"Full data validation accuracy:",
solver.check_accuracy(data['X_val'], data['y_val'])
)
# -
# ## Visualize Filters
# You can visualize the first-layer convolutional filters from the trained network by running the following:
# +
from cs231n.vis_utils import visualize_grid
grid = visualize_grid(model.params['W1'].transpose(0, 2, 3, 1))
plt.imshow(grid.astype('uint8'))
plt.axis('off')
plt.gcf().set_size_inches(5, 5)
plt.show()
# -
# # Spatial Batch Normalization
# We already saw that batch normalization is a very useful technique for training deep fully-connected networks. As proposed in the original paper (link in `BatchNormalization.ipynb`), batch normalization can also be used for convolutional networks, but we need to tweak it a bit; the modification will be called "spatial batch normalization."
#
# Normally batch-normalization accepts inputs of shape `(N, D)` and produces outputs of shape `(N, D)`, where we normalize across the minibatch dimension `N`. For data coming from convolutional layers, batch normalization needs to accept inputs of shape `(N, C, H, W)` and produce outputs of shape `(N, C, H, W)` where the `N` dimension gives the minibatch size and the `(H, W)` dimensions give the spatial size of the feature map.
#
# If the feature map was produced using convolutions, then we expect every feature channel's statistics e.g. mean, variance to be relatively consistent both between different images, and different locations within the same image -- after all, every feature channel is produced by the same convolutional filter! Therefore spatial batch normalization computes a mean and variance for each of the `C` feature channels by computing statistics over the minibatch dimension `N` as well the spatial dimensions `H` and `W`.
#
#
# [1] [<NAME> and <NAME>, "Batch Normalization: Accelerating Deep Network Training by Reducing
# Internal Covariate Shift", ICML 2015.](https://arxiv.org/abs/1502.03167)
# ## Spatial batch normalization: forward
#
# In the file `cs231n/layers.py`, implement the forward pass for spatial batch normalization in the function `spatial_batchnorm_forward`. Check your implementation by running the following:
# + tags=[]
np.random.seed(231)
# Check the training-time forward pass by checking means and variances
# of features both before and after spatial batch normalization
N, C, H, W = 2, 3, 4, 5
x = 4 * np.random.randn(N, C, H, W) + 10
print('Before spatial batch normalization:')
print(' Shape: ', x.shape)
print(' Means: ', x.mean(axis=(0, 2, 3)))
print(' Stds: ', x.std(axis=(0, 2, 3)))
# Means should be close to zero and stds close to one
gamma, beta = np.ones(C), np.zeros(C)
bn_param = {'mode': 'train'}
out, _ = spatial_batchnorm_forward(x, gamma, beta, bn_param)
print('After spatial batch normalization:')
print(' Shape: ', out.shape)
print(' Means: ', out.mean(axis=(0, 2, 3)))
print(' Stds: ', out.std(axis=(0, 2, 3)))
# Means should be close to beta and stds close to gamma
gamma, beta = np.asarray([3, 4, 5]), np.asarray([6, 7, 8])
out, _ = spatial_batchnorm_forward(x, gamma, beta, bn_param)
print('After spatial batch normalization (nontrivial gamma, beta):')
print(' Shape: ', out.shape)
print(' Means: ', out.mean(axis=(0, 2, 3)))
print(' Stds: ', out.std(axis=(0, 2, 3)))
# + tags=[]
np.random.seed(231)
# Check the test-time forward pass by running the training-time
# forward pass many times to warm up the running averages, and then
# checking the means and variances of activations after a test-time
# forward pass.
N, C, H, W = 10, 4, 11, 12
bn_param = {'mode': 'train'}
gamma = np.ones(C)
beta = np.zeros(C)
for t in range(50):
x = 2.3 * np.random.randn(N, C, H, W) + 13
spatial_batchnorm_forward(x, gamma, beta, bn_param)
bn_param['mode'] = 'test'
x = 2.3 * np.random.randn(N, C, H, W) + 13
a_norm, _ = spatial_batchnorm_forward(x, gamma, beta, bn_param)
# Means should be close to zero and stds close to one, but will be
# noisier than training-time forward passes.
print('After spatial batch normalization (test-time):')
print(' means: ', a_norm.mean(axis=(0, 2, 3)))
print(' stds: ', a_norm.std(axis=(0, 2, 3)))
# -
# ## Spatial batch normalization: backward
# In the file `cs231n/layers.py`, implement the backward pass for spatial batch normalization in the function `spatial_batchnorm_backward`. Run the following to check your implementation using a numeric gradient check:
# + tags=[]
np.random.seed(231)
N, C, H, W = 2, 3, 4, 5
x = 5 * np.random.randn(N, C, H, W) + 12
gamma = np.random.randn(C)
beta = np.random.randn(C)
dout = np.random.randn(N, C, H, W)
bn_param = {'mode': 'train'}
fx = lambda x: spatial_batchnorm_forward(x, gamma, beta, bn_param)[0]
fg = lambda a: spatial_batchnorm_forward(x, gamma, beta, bn_param)[0]
fb = lambda b: spatial_batchnorm_forward(x, gamma, beta, bn_param)[0]
dx_num = eval_numerical_gradient_array(fx, x, dout)
da_num = eval_numerical_gradient_array(fg, gamma, dout)
db_num = eval_numerical_gradient_array(fb, beta, dout)
#You should expect errors of magnitudes between 1e-12~1e-06
_, cache = spatial_batchnorm_forward(x, gamma, beta, bn_param)
dx, dgamma, dbeta = spatial_batchnorm_backward(dout, cache)
print('dx error: ', rel_error(dx_num, dx))
print('dgamma error: ', rel_error(da_num, dgamma))
print('dbeta error: ', rel_error(db_num, dbeta))
# -
# # Group Normalization
# In the previous notebook, we mentioned that Layer Normalization is an alternative normalization technique that mitigates the batch size limitations of Batch Normalization. However, as the authors of [2] observed, Layer Normalization does not perform as well as Batch Normalization when used with Convolutional Layers:
#
# >With fully connected layers, all the hidden units in a layer tend to make similar contributions to the final prediction, and re-centering and rescaling the summed inputs to a layer works well. However, the assumption of similar contributions is no longer true for convolutional neural networks. The large number of the hidden units whose
# receptive fields lie near the boundary of the image are rarely turned on and thus have very different
# statistics from the rest of the hidden units within the same layer.
#
# The authors of [3] propose an intermediary technique. In contrast to Layer Normalization, where you normalize over the entire feature per-datapoint, they suggest a consistent splitting of each per-datapoint feature into G groups, and a per-group per-datapoint normalization instead.
#
# <p align="center">
# <img src="https://raw.githubusercontent.com/cs231n/cs231n.github.io/master/assets/a2/normalization.png">
# </p>
# <center>Visual comparison of the normalization techniques discussed so far (image edited from [3])</center>
#
# Even though an assumption of equal contribution is still being made within each group, the authors hypothesize that this is not as problematic, as innate grouping arises within features for visual recognition. One example they use to illustrate this is that many high-performance handcrafted features in traditional Computer Vision have terms that are explicitly grouped together. Take for example Histogram of Oriented Gradients [4]-- after computing histograms per spatially local block, each per-block histogram is normalized before being concatenated together to form the final feature vector.
#
# You will now implement Group Normalization. Note that this normalization technique that you are to implement in the following cells was introduced and published to ECCV just in 2018 -- this truly is still an ongoing and excitingly active field of research!
#
# [2] [Ba, <NAME>, <NAME>, and <NAME>. "Layer Normalization." stat 1050 (2016): 21.](https://arxiv.org/pdf/1607.06450.pdf)
#
#
# [3] [<NAME>, and <NAME>. "Group Normalization." arXiv preprint arXiv:1803.08494 (2018).](https://arxiv.org/abs/1803.08494)
#
#
# [4] [<NAME> and <NAME>. Histograms of oriented gradients for
# human detection. In Computer Vision and Pattern Recognition
# (CVPR), 2005.](https://ieeexplore.ieee.org/abstract/document/1467360/)
# ## Group normalization: forward
#
# In the file `cs231n/layers.py`, implement the forward pass for group normalization in the function `spatial_groupnorm_forward`. Check your implementation by running the following:
# + tags=[]
np.random.seed(231)
# Check the training-time forward pass by checking means and variances
# of features both before and after spatial batch normalization
N, C, H, W = 2, 6, 4, 5
G = 2
x = 4 * np.random.randn(N, C, H, W) + 10
x_g = x.reshape((N*G,-1))
print('Before spatial group normalization:')
print(' Shape: ', x.shape)
print(' Means: ', x_g.mean(axis=1))
print(' Stds: ', x_g.std(axis=1))
# Means should be close to zero and stds close to one
gamma, beta = np.ones((1,C,1,1)), np.zeros((1,C,1,1))
bn_param = {'mode': 'train'}
out, _ = spatial_groupnorm_forward(x, gamma, beta, G, bn_param)
out_g = out.reshape((N*G,-1))
print('After spatial group normalization:')
print(' Shape: ', out.shape)
print(' Means: ', out_g.mean(axis=1))
print(' Stds: ', out_g.std(axis=1))
# -
# ## Spatial group normalization: backward
# In the file `cs231n/layers.py`, implement the backward pass for spatial batch normalization in the function `spatial_groupnorm_backward`. Run the following to check your implementation using a numeric gradient check:
# + tags=[]
np.random.seed(231)
N, C, H, W = 2, 6, 4, 5
G = 2
x = 5 * np.random.randn(N, C, H, W) + 12
gamma = np.random.randn(1,C,1,1)
beta = np.random.randn(1,C,1,1)
dout = np.random.randn(N, C, H, W)
gn_param = {}
fx = lambda x: spatial_groupnorm_forward(x, gamma, beta, G, gn_param)[0]
fg = lambda a: spatial_groupnorm_forward(x, gamma, beta, G, gn_param)[0]
fb = lambda b: spatial_groupnorm_forward(x, gamma, beta, G, gn_param)[0]
dx_num = eval_numerical_gradient_array(fx, x, dout)
da_num = eval_numerical_gradient_array(fg, gamma, dout)
db_num = eval_numerical_gradient_array(fb, beta, dout)
_, cache = spatial_groupnorm_forward(x, gamma, beta, G, gn_param)
dx, dgamma, dbeta = spatial_groupnorm_backward(dout, cache)
#You should expect errors of magnitudes between 1e-12~1e-07
print('dx error: ', rel_error(dx_num, dx))
print('dgamma error: ', rel_error(da_num, dgamma))
print('dbeta error: ', rel_error(db_num, dbeta))
# -
|
legacy/assignments/local/2020/assignment2/ConvolutionalNetworks.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Study of Correlation Between Building Demolition and Associated Features
#
# + Capstone Project for Data Science at Scale on Coursera
# + Repo is located [here](https://github.com/cyang019/blight_fight)
#
# **<NAME>** [<EMAIL>](<EMAIL>)
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from IPython.display import Image
# %matplotlib inline
# ## Objective
#
# Build a model to make predictions on blighted buildings based on real data from [data.detroitmi.gov](https://data.detroitmi.gov/Property-Parcels/Parcel-Map/fxkw-udwf/data) as given by coursera.
#
# Building demolition is very important for the city to turn around and revive its economy. However, it's no easy task. Accurate predictions can provide guidance on potential blighted buildings and help avoid complications at early stages.
# ## Building List
#
# The buildings were defined as described below:
#
# 1. Building sizes were estimated using parcel info downloaded [here](https://data.detroitmi.gov/Property-Parcels/Parcel-Map/fxkw-udwf/data) at data.detroitmi.gov. Details can be found in [this notebook](https://github.com/cyang019/blight_fight/blob/master/src/Building_size_estimation.ipynb).
# 2. A event table was constructed from the 4 files (detroit-311.csv, detroit-blight-violations.csv, detroit-crime.csv, and detroit-demolition-permits.tsv) using their coordinates, as shown [here](https://github.com/cyang019/blight_fight/blob/master/src/Cleaning_data.ipynb).
# 3. Buildings were defined using these coordinates with an estimated building size (median of all parcels). Each building was represented as a same sized rectangle.
# The resulted buildings:
Image("./data/buildings_distribution.png")
# ## Features
#
# Three kinds (311-calls, blight-violations, and crimes) of incident counts and coordinates (normalized) was used in the end. I also tried to generate more features by differentiating each kind of crimes or each kind of violations in this [notebook](https://github.com/cyang019/blight_fight/blob/master/src/Feature_Engineering.ipynb). However, these differentiated features lead to smaller AUC scores.
# ## Data
# + The buildings were down-sampled to contain same number of blighted buildings and non-blighted ones.
# + The ratio between train and test was set at a ratio of 80:20.
# + During training using xgboost, the train data was further separated into train and evaluation with a ratio of 80:20 for monitoring.
# ## Model
# + A **Gradient Boosted Tree** model using [Xgboost](xgboost.readthedocs.io) achieved AUC score of 0.85 on evaluation data set:
Image('./data/train_process.png')
# + This model resulted in an AUC score of **0.858** on test data. Feature importances are shown below:
Image('./data/feature_f_scores.png')
# Locations were most important features in this model. Although I tried using more features generated by differentiating different kind of crimes or violations, the AUC scores did not improve.
# + Feature importance can also be viewed using tree representation:
Image('./data/bst_tree.png')
# + To reduce variance of the model, since overfitting was observed during training. I also tried to reduce variance by including in more nonblighted buildings by sampling again multiple times with replacement (**bagging**).
# + A final AUC score of **0.8625** was achieved. The resulted ROC Curve on test data is shown below:
Image('./data/ROC_Curve_combined.png')
# ## Discussion
# Several things worth trying:
#
# + Using neural net to study more features generated from differentiated crimes or violations if given more time.
# + Taken into account possibilities that a building might blight in the future.
# # Thanks for your time reading the report!
|
Final_Report.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import warnings
import itertools
import pandas as pd
import numpy as np
import statsmodels.api as sm
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
index = pd.DatetimeIndex(start=data.data['date'][0].decode('utf-8'), periods=len(data.data), freq='W-SAT')
y = pd.DataFrame(data.data['co2'], index=index, columns=['co2'])
y.head()
y.shape
y.plot(figsize=(15, 6))
plt.show()
# +
# The 'MS' string groups the data in buckets by start of the month
y = y['co2'].resample('MS').mean()
# The term bfill means that we use the value before filling in missing values
y = y.fillna(y.bfill())
y.plot(figsize=(15, 6))
plt.show()
# +
# Define the p, d and q parameters to take any value between 0 and 2
p = d = q = range(0, 2)
# Generate all different combinations of p, q and q triplets
pdq = list(itertools.product(p, d, q))
# Generate all different combinations of seasonal p, q and q triplets
seasonal_pdq = [(x[0], x[1], x[2], 12) for x in list(itertools.product(p, d, q))]
print('Examples of parameter combinations for Seasonal ARIMA...')
print('SARIMAX: {} x {}'.format(pdq[1], seasonal_pdq[1]))
print('SARIMAX: {} x {}'.format(pdq[1], seasonal_pdq[2]))
print('SARIMAX: {} x {}'.format(pdq[2], seasonal_pdq[3]))
print('SARIMAX: {} x {}'.format(pdq[2], seasonal_pdq[4]))
# +
warnings.filterwarnings("ignore") # specify to ignore warning messages
for param in pdq:
for param_seasonal in seasonal_pdq:
try:
mod = sm.tsa.statespace.SARIMAX(y,
order=param,
seasonal_order=param_seasonal,
enforce_stationarity=False,
enforce_invertibility=False)
results = mod.fit()
print('ARIMA{}x{}12 - AIC:{}'.format(param, param_seasonal, results.aic))
except:
continue
# +
mod = sm.tsa.statespace.SARIMAX(y,
order=(1, 1, 1),
seasonal_order=(1, 1, 1, 12),
enforce_stationarity=False,
enforce_invertibility=False)
results = mod.fit()
print(results.summary().tables[1])
# -
results.plot_diagnostics(figsize=(15, 12))
plt.show()
|
models/ARIMA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Statistical treatment for PASTIS
#
# Essentially a copy of notebook #14 so that we can mess around with things here.
#
# 1. set target contrat in code cell 2 (e.g. `1e-10`)
# 2. set apodizer design in code cell 3 (e.g. `small`)
# 3. comment in correct data path in code cell 3 (e.g. `[...]/2020-01-27T23-57-00_luvoir-small`)
# +
# Imports
import os
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm
# %matplotlib inline
from astropy.io import fits
import astropy.units as u
import hcipy as hc
from hcipy.optics.segmented_mirror import SegmentedMirror
os.chdir('../../pastis/')
import util_pastis as util
from e2e_simulators.luvoir_imaging import LuvoirAPLC
# -
nmodes = 120
c_target = 1e-10
# ## Instantiate LUVOIR telescope for full functionality
# +
apodizer_design = 'large'
#savedpath = '/Users/ilaginja/Documents/data_from_repos/pastis_data/2020-01-27T23-57-00_luvoir-small'
#savedpath = '/Users/ilaginja/Documents/data_from_repos/pastis_data/2020-01-28T02-17-18_luvoir-medium'
savedpath = '/Users/ilaginja/Documents/data_from_repos/pastis_data/2020-01-28T04-45-55_luvoir-large'
# +
# Instantiate LUVOIR
sampling = 4
# This path is specific to the paths used in the LuvoirAPLC class
optics_input = '/Users/ilaginja/Documents/LabWork/ultra/LUVOIR_delivery_May2019/'
luvoir = LuvoirAPLC(optics_input, apodizer_design, sampling)
# -
# Make reference image
luvoir.flatten()
psf_unaber, ref = luvoir.calc_psf(ref=True)
norm = ref.max()
# +
# Make dark hole
dh_outer = hc.circular_aperture(2*luvoir.apod_dict[apodizer_design]['owa'] * luvoir.lam_over_d)(luvoir.focal_det)
dh_inner = hc.circular_aperture(2*luvoir.apod_dict[apodizer_design]['iwa'] * luvoir.lam_over_d)(luvoir.focal_det)
dh_mask = (dh_outer - dh_inner).astype('bool')
plt.figure(figsize=(18, 6))
plt.subplot(131)
hc.imshow_field(psf_unaber/norm, norm=LogNorm())
plt.subplot(132)
hc.imshow_field(dh_mask)
plt.subplot(133)
hc.imshow_field(psf_unaber/norm, norm=LogNorm(), mask=dh_mask)
# -
dh_intensity = psf_unaber/norm * dh_mask
baseline_contrast = util.dh_mean(dh_intensity, dh_mask)
#np.mean(dh_intensity[np.where(dh_intensity != 0)])
print('Baseline contrast:', baseline_contrast)
# +
# Load PASTIS modes - piston value per segment per mode
pastismodes = np.loadtxt(os.path.join(savedpath, 'results', 'pastis_modes.txt'))
print('pastismodes.shape: {}'.format(pastismodes.shape))
# pastismodes[segs, modes]
# Load PASTIS matrix
pastismatrix = fits.getdata(os.path.join(savedpath, 'matrix_numerical', 'PASTISmatrix_num_piston_Noll1.fits'))
# Load sigma vector
sigmas = np.loadtxt(os.path.join(savedpath, 'results', 'mode_requirements_1e-10_uniform.txt'))
#print(sigmas)
# -
# Calculate the inverse of the PASTIS mode matrix
# This is ModeToSegs in Mathematica
modestosegs = np.linalg.pinv(pastismodes)
# modestosegs[modes, segs]
# ## Static sigmas to avg contrast
# +
# Calculate mean contrast of all modes with PASTIS matrix AND the sigmas, to make sure this works
c_avg_sigma = []
for i in range(nmodes):
c_avg_sigma.append(util.pastis_contrast(sigmas[i] * pastismodes[:,i]*u.nm, pastismatrix))
print(c_avg_sigma)
# -
# Confirm that all of of them, with the baseline contarst, add up to the target contrast.
np.sum(c_avg_sigma) + baseline_contrast
# Draw random numbers
# +
# Create x-array
x_vals = np.zeros_like(sigmas)
for i, sig in enumerate(sigmas):
x_vals[i] = np.random.normal(loc=0, scale=sig)
print(x_vals.shape)
print(x_vals)
# +
# Calculate mean contrast of all modes with PASTIS matrix AND the sigmas, to make sure this works
c_avg_sigma = []
for i in range(nmodes):
c_avg_sigma.append(util.pastis_contrast(x_vals[i] * pastismodes[:,i]*u.nm, pastismatrix))
print(c_avg_sigma)
# -
np.sum(c_avg_sigma) + baseline_contrast
# Loop it up - cumulatively
# +
runnum = 1000
xs_list = []
for l in range(runnum):
x_vals = np.zeros_like(sigmas)
for i, sig in enumerate(sigmas):
x_vals[i] = np.random.normal(loc=0, scale=sig)
#x_vals[i] = np.random.uniform(-sig, sig)
xs_list.append(x_vals)
xs_list = np.array(xs_list)
xs_list.shape
# -
c_avg_list = []
for a in range(runnum):
c_avg_sigma = []
for i in range(nmodes):
c_avg_sigma.append(util.pastis_contrast(xs_list[a][i] * pastismodes[:,i]*u.nm, pastismatrix))
c_avg_list.append(np.sum(c_avg_sigma) + baseline_contrast)
print(np.mean(c_avg_list))
print(np.std(c_avg_list))
# Which results in the input target contrast.
#
# Same thing for a single mode at a time
# +
modechoice = 50
runnum = 1000
xs_list = []
for l in range(runnum):
x_vals = np.zeros_like(sigmas)
x_vals[modechoice] = np.random.normal(loc=0, scale=sigmas[modechoice])
xs_list.append(x_vals)
xs_list = np.array(xs_list)
xs_list.shape
# -
c_avg_list = []
for a in range(runnum):
c_avg_sigma = util.pastis_contrast(xs_list[a][modechoice] * pastismodes[:,modechoice]*u.nm, pastismatrix)
c_avg_list.append(c_avg_sigma + baseline_contrast)
print(np.mean(c_avg_list))
print(np.std(c_avg_list))
# Which is equivalent to:
c_target/120 + baseline_contrast
# We are able to define a static sigma value from the target contrast and singular values (See equation for sigmas). With this static sigmas, you get the correct contrast in a deterministic sense, see cumulative contrast plot.
#
# We have verified in this notebook that a Gaussian distribution with a zero mean and std = sigma prodices an average (statistical) DH mean (spatial) contrast. It's the statistical average of the mean contrast. We have checked this both for individual modes as well as the ensemble of modes all toghether.
#
# ## Segment requirements
#
# Now we want to transform this into segment-based requirements/tolerances/constraints $\mu$.
#
# We now want to verify that this works in segment space (as opposed to the mode-base the sigmas are defined in).
#
# We're doing $ y = M \cdot x$ here.
# +
runnum = 1000
ys_list = []
xs_list = []
for l in range(runnum):
x_vals = np.zeros_like(sigmas)
for i, sig in enumerate(sigmas):
x_vals[i] = np.random.normal(loc=0, scale=sig)
xs_list.append(x_vals)
# Calculate y vector
y_vals = np.dot(pastismodes, x_vals)
ys_list.append(y_vals)
ys_list = np.array(ys_list)
xs_list = np.array(xs_list)
ys_list.shape
# -
c_avg_list = []
for a in range(runnum):
c_avg_sigma = util.pastis_contrast(ys_list[a]*u.nm, pastismatrix)
c_avg_list.append(c_avg_sigma + baseline_contrast)
#c_avg_list.append(c_avg_sigma)
print(np.mean(c_avg_list))
print(np.std(c_avg_list))
print(np.mean(ys_list))
print(np.mean(xs_list))
# This means it is completely equivalent to work in the mode basis or in the segment basis.
#
# We have now a bunch of ys for which this works. Now we want to figure out what distributions these y maps (correct realizations of $\mu$s follow, and what we can quote as segment requirements.
# The y capture everything, including the cross-terms of the covariance matrix. Just averaging the ys will probably not be enough, but we can run them through the MC and see. If the off-diagonal terms of the Covariance matrix are not large though, this will be very similar.
#
# We need to assemble the Covariance matrix for y, $C_y$.
#
# ## Covariance matrix $C_y$
#
# Build $C_x$ by hand by dumping the square of the std (= variance) into the diagonal of a properly sized matrix.
cx = np.diag(np.square(sigmas))
#print(cx)
# $$y = M \cdot x$$
# $$C_x = E(x \cdot x^T)$$
# $$C_y = E(y \cdot y^T) = M \cdot E(x \cdot x^T) \cdot M^T$$
# $$C_y = M \cdot C_x \cdot M^T$$
cy = np.dot(pastismodes, np.dot(cx, np.transpose(pastismodes)))
plt.figure(figsize=(10,10))
plt.imshow(cy)
plt.xlabel('segment')
plt.ylabel('segment')
plt.colorbar()
cy[0,0]
testmap = np.sqrt(np.diag(cy))
print(testmap)
luvoir.flatten()
testmap *= u.nm
for seg, mu in enumerate(testmap):
luvoir.set_segment(seg+1, (mu).to(u.m).value/2, 0, 0)
psf, ref = luvoir.calc_psf(ref=True, display_intermediate=True)
testmap
# +
# Recalculate the xs
runnum = 1000
xs_list = []
for l in range(runnum):
x_vals = np.zeros_like(sigmas)
for i, sig in enumerate(sigmas):
x_vals[i] = np.random.normal(loc=0, scale=sig) #sig
#x_vals[i] = np.random.uniform(-sig, sig)
xs_list.append(x_vals)
xs_list = np.array(xs_list)
xs_list.shape
# +
# Empirical Cx
# !! Make sure to rerun the correct xs_list
pall = []
for i in range(runnum):
pall.append(np.outer(xs_list[i], np.transpose(xs_list[i])))
pall = np.array(pall)
# -
cx_emp = np.mean(pall, axis=0)
print(cx_emp.shape)
plt.figure(figsize=(10, 10))
plt.imshow(np.log10(cx_emp))
plt.colorbar()
np.sqrt(np.diag(cx_emp)) # these should be equivlaneet to the sigmas
plt.plot(sigmas)
plt.semilogy()
sigmas
np.allclose(cx, cx_emp)
plt.figure(figsize=(10, 10))
plt.imshow(np.log10(cx))
plt.colorbar()
np.sqrt(np.diag(cx))
np.sqrt(np.diag(cy))
# +
# Empirical Cy
pall_y = []
for i in range(ys_list.shape[0]):
pall_y.append(np.outer(ys_list[i], np.transpose(ys_list[i])))
print('runnum = {}'.format(ys_list.shape[0]))
pall_y = np.array(pall_y)
# -
cy_emp = np.mean(pall_y, axis=0)
print(cy_emp.shape)
plt.figure(figsize=(10, 10))
plt.imshow(cy_emp)
plt.colorbar()
# Get mean of the ys directly, ignoring the Covariance matrix.
y_direkt_mean = np.mean(ys_list, axis=0)
print(y_direkt_mean.shape)
print(y_direkt_mean)
np.mean(y_direkt_mean)
# Put those mean ys/mus on the simulator
luvoir.flatten()
y_direkt_mean *= u.nm
for seg, mu in enumerate(y_direkt_mean):
luvoir.set_segment(seg+1, (mu).to(u.m).value/2, 0, 0)
psf, ref = luvoir.calc_psf(ref=True, display_intermediate=True)
avg_con = util.pastis_contrast(y_direkt_mean, pastismatrix) + baseline_contrast
print(avg_con)
# ### Some tests
#
# Use an x vector with only one entry (=1) and see whether $y = M \cdot x$ yields a PASTIS mode.
# +
testmode = -2
x_test = np.zeros(nmodes)
x_test[testmode] = 1
print(x_test)
# -
y_test = np.dot(pastismodes, x_test)
y_test *= u.nm
# Put the resulting y on the simulator
luvoir.flatten()
for seg, coef in enumerate(y_test):
luvoir.set_segment(seg+1, (coef).to(u.m).value/2, 0, 0)
psf, ref = luvoir.calc_psf(ref=True, display_intermediate=True)
|
Jupyter Notebooks/LUVOIR/15_Debug contrast floor.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="Lym_glPd23n7"
# !nvidia-smi
# + id="61KeHP4iDHOc"
# %cd /content/
# !git clone -b reduced-data https://github.com/westphal-jan/peer-data
# %cd /content/peer-data
# # !git checkout huggingface
# !git submodule update --init --recursive
# + id="ZzxkH1uGFGKb"
# # !pip install pytorch-lightning wandb python-dotenv catalyst sentence-transformers numpy requests
# !pip install wandb transformers nltk pytorch-lightning
# + id="W1ludytPTqS4"
import os
import torch
import json
import glob
from pathlib import Path
from tqdm import tqdm
# from sklearn.model_selection import train_test_split
from transformers import DistilBertTokenizerFast, DistilBertForSequenceClassification, Trainer, TrainingArguments, TrainerCallback, TrainerState, TrainerControl, AdamW, AutoTokenizer, AutoModel, AutoModelForSequenceClassification, AutoModelWithLMHead, T5ForConditionalGeneration
import wandb
from datetime import datetime
import pickle
import numpy as np
# import nlpaug.augmenter.word as naw
from torch.utils.data import DataLoader, WeightedRandomSampler
from torch import nn, optim
import copy
from datetime import datetime
import matplotlib.pyplot as plt
from matplotlib.ticker import PercentFormatter
from collections import defaultdict, Counter
from nltk.corpus import stopwords
import nltk
nltk.download("stopwords")
# + id="5IZktYlUTlqq"
class PaperDataset(torch.utils.data.Dataset):
def __init__(self, encodings, labels):
self.encodings = encodings
self.labels = labels
def __getitem__(self, idx):
item = {key: torch.tensor(val[idx]).float() for key, val in self.encodings.items()}
item['labels'] = torch.tensor(self.labels[idx]).float()
return item
def __len__(self):
return len(self.labels)
def raw_read_dataset(data_dir: Path, num_texts=None):
file_paths = glob.glob(f"{data_dir}/*.json")
if num_texts != None:
file_paths = file_paths[:num_texts]
raws = []
for i, file_path in enumerate(tqdm(file_paths)):
with open(file_path) as f:
paper_json = json.load(f)
raws.append(paper_json)
return raws
def read_dataset(data_dirs, num_texts=None, restrict_file=None):
if not isinstance(data_dirs, list):
data_dirs = [data_dirs]
# with open(restrict_file, "r") as f:
# filter_file_names = f.read().splitlines()
# for data_dir in data_dirs:
# file_paths = glob.glob(f"{data_dir}/*.json")
# file_paths = [p for p in file_paths if p.split("/")[-1] in filter_file_names]
# print(data_dir, len(file_paths))
file_paths = []
for data_dir in data_dirs:
file_paths.extend(glob.glob(f"{data_dir}/*.json"))
if restrict_file:
with open(restrict_file, "r") as f:
filter_file_names = f.read().splitlines()
file_paths = [p for p in file_paths if p.split("/")[-1] in filter_file_names]
if num_texts != None:
file_paths = file_paths[:num_texts]
abstracts = []
sections = []
labels = []
for i, file_path in enumerate(tqdm(file_paths)):
with open(file_path) as f:
paper_json = json.load(f)
accepted = paper_json["review"]["accepted"]
abstract = paper_json["review"]["abstract"]
_sections = paper_json["pdf"]["metadata"]["sections"]
_sections = _sections if _sections else []
abstracts.append(abstract)
labels.append(int(accepted))
sections.append(_sections)
return abstracts, sections, labels
# + id="_ZzdLZ2ppaGw"
data_dir = "data/original"
augmented = ["data/back-translations-train-accepted", "data/back-translations-train-rejected"]
data_dirs = [data_dir]# + augmented
#8121, 122194
_, train_sections, train_labels = read_dataset(data_dirs, restrict_file="data/train.txt")
_, val_sections, val_labels = read_dataset(data_dir, restrict_file="data/val.txt")
_, test_sections, test_labels = read_dataset(data_dir, restrict_file="data/test.txt")
# + id="ge-mQB1zS2M0"
# def label_distribution(labels):
# num_rejected, num_accepted = labels.count(0), labels.count(1)
# print(num_rejected, num_rejected / len(labels), num_accepted, num_accepted / len(labels))
# label_distribution(train_labels)
# label_distribution(val_labels)
# label_distribution(test_labels)
# print(len(np.nonzero(list(map(lambda x: len(x), train_sections)))[0]))
# print(len(np.nonzero(list(map(lambda x: len(x), val_sections)))[0]))
# print(len(np.nonzero(list(map(lambda x: len(x), test_sections)))[0]))
# + id="3fQpyNN7X1J7"
# num_sections = list(map(lambda x: len(x), train_sections))
# num_sections = sorted(num_sections)
# len(num_sections)
# q = 0.95
# index = int(len(num_sections) * q)
# percentile = num_sections[index]
# print(q, percentile, index, len(num_sections) - index)
# print(num_sections[index:])
# + id="Phhp-sMYpdss"
def extract_sections(sections, labels):
all_sections = []
all_labels = []
assignments = []
section_counter = 0
for _sections, label in zip(sections, labels):
if len(sections) == 0:
continue
texts = list(map(lambda x: x["text"], _sections))
all_sections.extend(texts)
all_labels.extend([label] * len(_sections))
# Create mapping from original submission to flattened sections
new_section_counter = section_counter + len(_sections)
assignments.append((section_counter, new_section_counter))
section_counter = new_section_counter
return all_sections, all_labels, assignments
flattened_train_sections, flattened_train_labels, _ = extract_sections(train_sections, train_labels)
flattened_val_sections, flattened_val_labels, val_assignemnts = extract_sections(val_sections, val_labels)
flattened_test_sections, flattened_test_labels, test_assignemnts = extract_sections(test_sections, test_labels)
print(len(flattened_train_sections), len(flattened_val_sections), len(flattened_test_sections))
num_accepted, num_rejected = flattened_train_labels.count(1), flattened_train_labels.count(0)
print(num_accepted, num_rejected)
label_weight = num_rejected / np.array([num_rejected, num_accepted])
print(label_weight)
# + id="E-B3-ZhNp4Jf"
tokenizer = AutoTokenizer.from_pretrained('sentence-transformers/paraphrase-TinyBERT-L6-v2')
# tokenizer = DistilBertTokenizerFast.from_pretrained('distilbert-base-cased')
# tokenizer = AutoTokenizer.from_pretrained('allenai/scibert_scivocab_uncased')
train_encodings = tokenizer(flattened_train_sections, truncation=True, padding="max_length", max_length=512)
val_encodings = tokenizer(flattened_val_sections, truncation=True, padding="max_length", max_length=512)
test_encodings = tokenizer(flattened_test_sections, truncation=True, padding="max_length", max_length=512)
# print(np.array(train_encodings["input_ids"]).shape)
train_dataset = PaperDataset(train_encodings, flattened_train_labels)
val_dataset = PaperDataset(val_encodings, flattened_val_labels)
test_dataset = PaperDataset(test_encodings, flattened_test_labels)
# + id="NewpWgWk8PPP"
# special_characters = set(['-', '\'', '.', ',', '!','"','#','$','%','&','(',')','*','+','/',':',';','<','=','>','@','[','\\',']','^','`','{','|','}','~','\t'])
# english_stopwords = set(stopwords.words('english'))
# def tokenize(text: str, token_to_id=None, encode=False):
# for c in special_characters:
# text = text.replace(c, ' ')
# text = text.lower()
# tokens = [t for t in text.split(' ') if t]
# tokens = [t for t in tokens if t.isalpha()]
# tokens = [t for t in tokens if not t in english_stopwords]
# if token_to_id:
# tokens = [t for t in tokens if t in token_to_id]
# if encode:
# tokens = [token_to_id[t] for t in tokens]
# onehot = np.zeros(len(token_to_id))
# onehot[tokens] = 1
# return onehot
# return tokens
# train_tokens = list(map(tokenize, flattened_train_sections))
# flattened_tokens = [item for sublist in train_tokens for item in sublist]
# vocab = sorted(list(set(flattened_tokens)))
# token_to_id = {t: i for i, t in enumerate(vocab)}
# val_tokens = list(map(lambda x: tokenize(x, token_to_id), flattened_val_sections))
# test_tokens = list(map(lambda x: tokenize(x, token_to_id), flattened_test_sections))
# print("Vocab Size:", len(vocab))
# + id="q5JJ-iuG8WHM"
train_encodings = list(map(lambda x: tokenize(x, token_to_id, True), flattened_train_sections))
val_encodings = list(map(lambda x: tokenize(x, token_to_id, True), flattened_val_sections))
test_encodings = list(map(lambda x: tokenize(x, token_to_id, True), flattened_test_sections))
train_dataset = PaperDataset({"tokens": train_encodings}, flattened_train_labels)
val_dataset = PaperDataset({"tokens": val_encodings}, flattened_val_labels)
test_dataset = PaperDataset({"tokens": test_encodings}, flattened_test_labels)
# + id="muf4YwxMaeeT"
samples_weight = [label_weight[t] for t in flattened_train_labels]
samples_weight = torch.Tensor(samples_weight)
sampler = WeightedRandomSampler(samples_weight, len(samples_weight))
# + id="PgxRi1Rcf2_5"
def compute_metrics(logits, labels, prefix="eval"):
# predictions = np.argmax(logits, axis=1)
predictions = np.array(logits) >= 0
actual = np.array(labels)
tp = ((predictions == 1) & (actual == 1)).sum()
fp = ((predictions == 1) & (actual == 0)).sum()
fn = ((predictions == 0) & (actual == 1)).sum()
tn = ((predictions == 0) & (actual == 0)).sum()
precision = tp / (tp + fp) if tp + fp > 0 else 0.0
recall = tp / (tp + fn) if tp + fn > 0 else 0.0
f1 = 2 * precision * recall / (precision + recall) if precision + recall > 0 else 0.0
accuracy = (tp + tn) / (tp + tn + fp + fn)
mcc = (tp * tn - fp * fn) / (((tp + fp) * (tp + fn) * (tn + fp) * (tn + fn))**0.5)
mcc = mcc if np.isnan(mcc) else 0.0
p = tp + fn
n = tn + fp
metrics = {"metric/accuracy": accuracy, "metric/precision": precision, "metric/recall": recall, "metric/f1": f1, "metric/mcc": mcc,
"classification/tp": tp, "classification/fp": fp, "classification/fn": fn, "classification/tn": tn, "classification/n": n, "classification/p": p}
metrics = {f"{prefix}/{key}": metric for key, metric in metrics.items()}
# "augmentation/train": train_dataset.num_augmentations, "augmentation/val": val_dataset.num_augmentations
return metrics
def compute_original_metrics(logits, original_labels, assignments, prefix="eval"):
assert len(original_labels) == len(assignments)
new_logits = []
for start_idx, end_idx in assignments:
_assignment = np.array(logits[start_idx:end_idx])
# TODO: Sum could also be possible but should only change confidence and not the outcome
reduced_logits = _assignment.mean(axis=0)
new_logits.append(reduced_logits.tolist())
return compute_metrics(new_logits, original_labels, prefix)
logits = [[1.0, 2.5], [1.3, 0.7]]
labels = [0, 1]
result = compute_metrics(logits, labels)
print(result)
logits = [[1.0, 2.5], [1.3, 0.7], [1.0, 2.5], [1.0, 2.5]]
assignments = [(0, 2), (2, 4)]
result = compute_original_metrics(logits, labels, assignments)
print(result)
# + id="bn9V7QNw4Pdv"
# class MyModel(nn.Module):
# def __init__(self):
# super().__init__()
# self.base_model = T5ForConditionalGeneration.from_pretrained("t5-base")
# self.classifier = nn.Linear(768, 2, bias=False)
# def forward(self, x):
# print(x)
# emb = self.base_model(**x)
# print(emb.shape)
# y = self.classifier(emb)
# print(y.shape)
# return y
# device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
# model = MyModel()
# model.to(device)
# loader = DataLoader(train_dataset, batch_size=4, shuffle=False)
# for _batch in loader:
# inputs = {key: val.to(device) for key, val in _batch.items()}
# labels = inputs.pop("labels")
# y = model(inputs)
# break
# + id="NTTuzfhv8EWJ"
# from klib.misc import kdict
# class BowClassifier(nn.Module):
# def __init__(self, input_size, output_size=1):
# super().__init__()
# self.input_size = input_size
# self.output_size = output_size
# self.classifier = nn.Linear(self.input_size, self.output_size, bias=False)
# def forward(self, tokens):
# y = self.classifier(tokens)
# y = y.squeeze()
# return kdict(logits=y)
# + id="D7iddk0kKS7x"
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
# model = BowClassifier(len(vocab)).float()
model = AutoModelForSequenceClassification.from_pretrained('sentence-transformers/paraphrase-TinyBERT-L6-v2', num_labels=2)
# model = AutoModelForSequenceClassification.from_pretrained('allenai/scibert_scivocab_uncased', num_labels=2)
# model = DistilBertForSequenceClassification.from_pretrained('distilbert-base-cased', num_labels=2)
model.to(device)
train_batch_size, val_batch_size = 64, 64
# train_loader = DataLoader(train_dataset, batch_size=train_batch_size, shuffle=True)
train_loader = DataLoader(train_dataset, batch_size=train_batch_size, sampler=sampler)
val_loader = DataLoader(val_dataset, batch_size=val_batch_size, shuffle=False)
# optimizer = AdamW(model.parameters(), lr=5e-5)
optimizer = optim.AdamW(model.parameters(), lr=0.001)
# _label_weight = torch.from_numpy(label_weight).float().to(device)
# loss_func = torch.nn.CrossEntropyLoss(_label_weight, reduction="sum")
loss_func = torch.nn.CrossEntropyLoss()
pos_weight = torch.tensor(label_weight[1]).float().to(device)
weighted_loss_func = nn.BCEWithLogitsLoss(pos_weight=pos_weight)
loss_func = nn.BCEWithLogitsLoss()
num_epochs = 3
wandb_logging = False
run_name = datetime.now().strftime('%d-%m-%Y_%H_%M_%S')
print("Run name:", run_name)
if wandb_logging:
wandb.login()
wandb.init(entity="paper-judging", project="huggingface", name=run_name)
output_dir=f'results/{run_name}'
os.makedirs(output_dir)
# logging_steps, eval_steps = 200, 2000
logging_steps, eval_steps = 400, 2000
# logging_steps, eval_steps = 5, 10
_steps = 0
def run_model(_model, _batch):
inputs = {key: val.to(device) for key, val in _batch.items()}
labels = inputs.pop("labels")
outputs = _model(**inputs)
logits = outputs.logits
loss = loss_func(logits, labels)
return loss, logits
min_val_loss, best_model, best_metrics, best_loss_epoch = None, None, None, None
best_f1, best_epoch = None, None
for epoch in range(num_epochs):
print(f"Epoch: {epoch + 1}/{num_epochs}")
train_losses = []
for batch in tqdm(train_loader, position=0, leave=True):
model.train()
optimizer.zero_grad()
train_loss, _ = run_model(model, batch)
train_losses.append(train_loss)
train_loss.backward()
optimizer.step()
metrics = {}
if _steps % logging_steps == 0:
_train_loss = sum(train_losses) / len(train_losses)
metrics.update({"train/loss": _train_loss.item(), "steps": _steps})
train_losses = []
if _steps % eval_steps == 0:
print("EVAL")
model.eval()
val_losses = []
logits = []
with torch.no_grad():
for val_batch in val_loader:
_val_loss, _logits = run_model(model, val_batch)
val_losses.append(_val_loss.item())
logits.extend(_logits.tolist())
val_loss = sum(val_losses) / len(val_losses)
_metrics = compute_metrics(logits, flattened_val_labels, "eval/flattened")
_original_metrics = compute_original_metrics(logits, val_labels, val_assignemnts, "eval")
metrics["eval/loss"] = val_loss
metrics.update(_metrics)
metrics.update(_original_metrics)
if min_val_loss == None:
min_val_loss = val_loss
elif min_val_loss > val_loss:
min_val_loss = val_loss
best_loss_epoch = epoch
f1 = metrics["eval/metric/f1"]
if best_f1 == None:
best_f1 = f1
best_model = copy.deepcopy(model)
elif f1 > best_f1:
best_f1 = f1
best_model = copy.deepcopy(model)
best_metrics = metrics
best_epoch = epoch
if metrics:
print(metrics)
if wandb_logging:
wandb.log(metrics)
_steps += 1
snapshot_dir = f"results/{run_name}/network-snapshot-latest"
# best_model.save_pretrained(snapshot_dir)
print(f"Best val metrics during training epoch {best_epoch}:")
print(best_metrics)
test_loader = DataLoader(test_dataset, batch_size=val_batch_size, shuffle=False)
best_model.eval()
with torch.no_grad():
logits = []
for val_batch in val_loader:
_, _logits = run_model(best_model, val_batch)
logits.extend(_logits.tolist())
metrics = compute_metrics(logits, flattened_val_labels, "eval/flattened")
original_metrics = compute_original_metrics(logits, val_labels, val_assignemnts, "eval")
print("Val metrics:")
print(metrics)
print(original_metrics)
logits = []
for test_batch in test_loader:
_, _logits = run_model(best_model, test_batch)
logits.extend(_logits.tolist())
metrics = compute_metrics(logits, flattened_test_labels, "test/flattened")
original_metrics = compute_original_metrics(logits, test_labels, test_assignemnts, "test")
print("Test metrics:")
print(metrics)
print(original_metrics)
if wandb_logging:
wandb.save(f"{snapshot_dir}/*", os.path.dirname(snapshot_dir))
wandb.finish()
# + id="UW6Q0AmmAE3m"
# wandb.finish()
# + id="dZPWnjVxU0or"
# device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
# model = DistilBertForSequenceClassification.from_pretrained('distilbert-base-uncased')
# model.to(device)
# + id="pOjvyeLkTUBz"
# val_batch_size = 16
# val_loader = DataLoader(val_dataset, batch_size=val_batch_size, shuffle=False)
# _label_weight = torch.from_numpy(label_weight).float().to(device)
# loss_func = torch.nn.CrossEntropyLoss(_label_weight, reduction="sum")
# model.eval()
# val_losses = []
# logits = []
# with torch.no_grad():
# for val_batch in tqdm(val_loader):
# inputs = {key: val.to(device) for key, val in val_batch.items()}
# _labels = inputs.pop("labels")
# _outputs = model(**inputs)
# _logits = _outputs.logits
# _val_loss = loss_func(_logits, _labels)
# val_losses.append(_val_loss.item())
# logits.extend(_logits.tolist())
# val_loss = sum(val_losses) / len(val_losses)
# metrics = compute_metrics((logits, val_labels))
# + id="M2AZLJF9zFCo"
# snapshot_dir = f"results/{run_name}/network-snapshot-latest"
# model.save_pretrained(snapshot_dir)
# if wandb_logging:
# wandb.save(f"{snapshot_dir}/*", os.path.dirname(snapshot_dir))
# wandb.finish()
# + id="q6gdbuMQsaZT"
# # !pip install wandb
# import wandb
# wandb.login()
# wandb.finish()
# + id="s-2Ct3_vc921"
# from transformers import AutoTokenizer, AutoModelWithLMHead
# t5_tokenizer = AutoTokenizer.from_pretrained("t5-base")
# t5_model = AutoModelWithLMHead.from_pretrained("t5-base").to(device)
# + id="Md8S9addYunG"
# input = "That's great"
# input_enc = t5_tokenizer(input, truncation=True, padding=True, return_tensors="pt")
# output = t5_model(**input_enc.to(device))
# print(output)
# print(torch.softmax(output.logits, dim=1))
# + id="odxHty8Cd9Xj"
# model_path = f"network-snapshot-latest-279194.pt"
# model = BowClassifier(len(vocab))
# model.load_state_dict(torch.load(model_path, map_location=torch.device('cpu')))
# def find_word(word, tokens, labels):
# total = [0, 0]
# classes = [0, 0]
# for i, _tokens in enumerate(tokens):
# _tokens = set(_tokens)
# label = labels[i]
# if word in _tokens:
# classes[label] += 1
# total[label] += 1
# # print(f"Not accepted: {classes[0]}/{total[0]} ({classes[0]/total[0]}), Accepted: {classes[1]}/{total[1]} ({classes[1]/total[1]})")
# return classes[0]/total[0], classes[1]/total[1]
# def analyze(params, k, tokens, labels):
# val, ind = params.topk(k)
# for i in ind:
# word = vocab[i]
# rejected, accepted = find_word(word, tokens, labels)
# n = 4
# print(f"{word} & {np.round(params[i].item(), 3)} & {np.round(rejected*100, n)} & {np.round(accepted*100, n)}")
# tokens = train_tokens + val_tokens + test_tokens
# print(len(train_tokens), len(val_tokens), len(test_tokens))
# labels = flattened_train_labels + flattened_val_labels + flattened_test_labels
# print(len(flattened_train_labels), len(flattened_val_labels), len(flattened_test_labels))
# params = list(model.parameters())[0][0]
# k = 8
# print(len(tokens), len(labels))
# print("Positive:")
# analyze(params, k, tokens, labels)
# print()
# print("Negative:")
# analyze(-params, k, tokens, labels)
# print()
# print("Unnecessary:")
# val, ind = (-(params.abs())).topk(k)
# for i in ind:
# print(vocab[i], params[i].item())
# + id="iXJP6lSnhSJU"
# paper_abstract = """Generative adversarial networks (GANs) have shown
# outstanding performance on a wide range of problems in
# computer vision, graphics, and machine learning, but often require numerous training data and heavy computational resources. To tackle this issue, several methods introduce a transfer learning technique in GAN training. They,
# however, are either prone to overfitting or limited to learning small distribution shifts. In this paper, we show that
# simple fine-tuning of GANs with frozen lower layers of
# the discriminator performs surprisingly well. This simple
# baseline, FreezeD, significantly outperforms previous techniques used in both unconditional and conditional GANs.
# We demonstrate the consistent effect using StyleGAN and
# SNGAN-projection architectures on several datasets of Animal Face, Anime Face, Oxford Flower, CUB-200-2011, and
# Caltech-256 datasets. The code and results are available at
# https://github.com/sangwoomo/FreezeD."""
# + id="BDqbwDCReV9a"
# input = ["It's incredibly bad", paper_abstract, "hello", "darkness"]
# input_enc = tokenizer(input, truncation=True, padding=True, return_tensors="pt")
# output = model(**input_enc.to(device))
# print(output)
# # print(torch.softmax(output.logits, dim=1))
# + id="ti9x-w_erAaX"
# prediction = output.logits.argmax(dim=1)
# actual = prediction
# n = tp = fp = fn = tn = 0
# tp += (prediction == 1) & (actual == 1)
# fp += (prediction == 1) & (actual == 0)
# fn += (prediction == 0) & (actual == 1)
# tn += (prediction == 0) & (actual == 0)
# + [markdown] id="dy500R68NdLG"
# # LIME
#
# + id="jnEQ1UWBNy5k"
# # !pip install lime transformers
# + id="N-vIZ4vhNfTX"
# import numpy as np
# import lime
# import torch
# import torch.nn.functional as F
# from lime.lime_text import LimeTextExplainer
# from transformers import AutoTokenizer, AutoModelForSequenceClassification
# tokenizer = AutoTokenizer.from_pretrained("ProsusAI/finbert")
# model = AutoModelForSequenceClassification.from_pretrained("ProsusAI/finbert")
# class_names = ['positive','negative', 'neutral']
# def predictor(texts):
# print(len(texts))
# outputs = model(**tokenizer(texts, return_tensors="pt", padding=True))
# probas = F.softmax(outputs.logits).detach().numpy()
# return probas
# explainer = LimeTextExplainer(class_names=class_names)
# str_to_predict = "surprising increase in revenue in spite of decrease in market share"
# exp = explainer.explain_instance(str_to_predict, predictor, num_features=20, num_samples=100)
# exp.show_in_notebook(text=str_to_predict)
|
NLP_Paper_Judge_Sections.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="xGfqtHThfPiE" colab_type="code" outputId="428bcf52-6bcf-4fac-85b6-7d46253adcc7" colab={"base_uri": "https://localhost:8080/", "height": 403}
# !pip install transformers==2.8.0
# + id="jZLCO1Jvfr4O" colab_type="code" outputId="267991b4-c2f4-4d8d-f603-7d80a7a6673b" colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd ~/../content
# + id="2fDvof2Nfy-5" colab_type="code" colab={}
import os
# + id="Eq9YdbeZf0Hw" colab_type="code" outputId="089414c6-3f0a-40b4-d1b3-2c38c000d728" colab={"base_uri": "https://localhost:8080/", "height": 34}
WORK_DIR = os.path.join('drive', 'My Drive', 'Colab Notebooks', 'NLU')
os.path.exists(WORK_DIR)
# + id="wVh5Qs1x0ENz" colab_type="code" outputId="92acc8ea-ad8a-4a8e-a45e-ba0d4073b6d7" colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd $WORK_DIR
# + id="jPWXQsot0ESU" colab_type="code" colab={}
GLUE_DIR="data/glue"
# + id="Ftm25a7O0G0J" colab_type="code" outputId="a28c8bf4-f707-4247-cdb7-6e56c8ce96b5" colab={"base_uri": "https://localhost:8080/", "height": 34}
# !echo $GLUE_DIR
# + id="EbNIkD8P0HDf" colab_type="code" colab={}
TASK_NAME="RTE"
# + id="DtZqPnCb0HF7" colab_type="code" outputId="3d179999-d76b-47f9-c7a3-7ac32899922e" colab={"base_uri": "https://localhost:8080/", "height": 34}
# !echo $TASK_NAME
# + id="rzPjt-Qhf3To" colab_type="code" outputId="519b834d-7272-41ca-d885-02ff5062e7fa" colab={"base_uri": "https://localhost:8080/", "height": 185}
# !python download_glue_data.py --help
# + id="ZiGvhtT_gAdr" colab_type="code" outputId="95b311a3-102d-46f9-972f-18240d2fa6ac" colab={"base_uri": "https://localhost:8080/", "height": 50}
# !python download_glue_data.py --data_dir $GLUE_DIR --tasks $TASK_NAME
# + id="oQaZ3_-SgB6r" colab_type="code" outputId="c16ad4ef-88b5-403c-c497-ae4689dddec4" colab={"base_uri": "https://localhost:8080/", "height": 34}
MODEL_DIR = os.path.join('model', 'roberta_token_discrimination_lr_1e-4_p_0.3')
os.path.exists(MODEL_DIR)
# + id="swzTjNDJgDVp" colab_type="code" outputId="b95debbf-4285-44a4-f06b-5b9b8c465be8" colab={"base_uri": "https://localhost:8080/", "height": 302}
# !nvidia-smi
# + id="Cw5WkGv6NfP3" colab_type="code" outputId="af066db5-ab8c-4fed-a1b6-2edc14a67749" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !python run_glue.py --help
# + id="990TFGa3gEOM" colab_type="code" outputId="be373091-3140-4ca2-91d6-1149dbc5d153" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !python run_glue.py \
# --model_type roberta \
# --model_name_or_path $MODEL_DIR \
# --task_name $TASK_NAME \
# --do_train \
# --do_eval \
# --data_dir $GLUE_DIR/$TASK_NAME \
# --max_seq_length 128 \
# --per_gpu_eval_batch_size=64 \
# --per_gpu_train_batch_size=64 \
# --learning_rate 2e-5 \
# --num_train_epochs 3 \
# --output_dir run \
# --overwrite_output_dir
# + id="GVeUbBYKgGiW" colab_type="code" colab={}
|
notebooks_GLUE/notebooks_roberta_jumbled_token_discrimination_lr_e-4_prob_0.3/rte.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://colab.research.google.com/github/timeseriesAI/tsai/blob/master/tutorial_nbs/10_Time_Series_Classification_and_Regression_with_MiniRocket.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# created by <NAME> and <NAME> (<EMAIL>) based on:
#
# * <NAME>., <NAME>., & <NAME>. (2020). MINIROCKET: A Very Fast (Almost) Deterministic Transform for Time Series Classification. arXiv preprint arXiv:2012.08791.
#
# * Original paper: https://arxiv.org/abs/2012.08791
#
# * Original code: https://github.com/angus924/minirocket
# # MiniRocket 🚀
#
# > A Very Fast (Almost) Deterministic Transform for Time Series Classification.
# ROCKET is a type of time series classification and regression methods that is different to the
# ones you may be familiar with. Typical machine learning classifiers will
# optimize the weights of convolutions, fully-connected, and pooling layers,
# learning a configuration of weights that classifies the time series.
#
# In contrast, ROCKET applies a large number of fixed, non-trainable, independent convolutions
# to the timeseries. It then extracts a number of features from each convolution
# output (a form of pooling), generating typically 10000 features per sample. (These
# features are simply floating point numbers.)
#
# The features are stored so that they can be used multiple times.
# It then learns a simple linear head to predict each time series sample from its features.
# Typical PyTorch heads might be based on Linear layers. When the number of training samples is small,
# sklearn's RidgeClassifier is often used.
#
# The convolutions' fixed weights and the pooling method have been chosen experimentally to
# effectively predict a broad range of real-world time series.
#
# The original ROCKET method used a selection of fixed convolutions with weights
# chosen according to a random distribution. Building upon the lessons learned
# from ROCKET, MiniRocket refines the convolutions to a specific pre-defined set
# that proved to be at least as effective ROCKET's. It is also much faster
# to calculate than the original ROCKET. Actually, the paper authors "suggest that MiniRocket should now be considered and used as the default variant of Rocket."
#
# MiniROCKET was implemented in Python using numba acceleration and mathematical
# speedups specific to the algorithm. It runs quite fast, utilizing CPU cores in
# parallel. Here we present a 2 implementations of MiniRocket:
# * a cpu version with an sklearn-like API (that can be used with small datasets - <10k samples), and
# * a PyTorch implementation of MiniRocket, optimized for
# the GPU. It runs faster (3-25x depending on your GPU) than the CPU version and offers some flexibility for further experimentation.
#
# We'll demonstrate how you can use both of them througout this notebook.
# # Import libraries 📚
# +
# ## NOTE: UNCOMMENT AND RUN THIS CELL IF YOU NEED TO INSTALL/ UPGRADE TSAI
# stable = False # True: stable version in pip, False: latest version from github
# if stable:
# # !pip install tsai -U >> /dev/null
# else:
# # !pip install git+https://github.com/timeseriesAI/tsai.git -U >> /dev/null
# ## NOTE: REMEMBER TO RESTART (NOT RECONNECT/ RESET) THE KERNEL/ RUNTIME ONCE THE INSTALLATION IS FINISHED
# -
# Restart your runtime before running the cells below.
from tsai.all import *
computer_setup()
# # Using MiniRocket 🚀
# * First, create the features for each timeseries sample using the MiniRocketFeatures module (MRF).
# MRF takes a minibatch of time series samples and outputs their features. Choosing an appropriate minibatch size
# allows training sets of any size to be used without exhausting CPU or GPU memory.
#
# Typically, 10000 features will characterize each sample. These features are relatively
# expensive to create, but once created they are fixed and may be used as the
# input for further training. They might be saved for example in memory or on disk.
#
#
# * Next, the features are sent to a linear model. The original
# MiniRocket research used sklearn's RidgeClassifier. When the number of samples
# goes beyond the capacity of RidgeClassifier, a deep learning "Head" can be
# used instead to learn the classification/regression from minibatches of features.
#
# For the following demos, we use the tsai package to handle timeseries efficiently and clearly. tsai is fully integrated with fastai, allowing fastai's training loop and other convenience to be used. To learn more about tsai, please check out the docs and tutorials at https://github.com/timeseriesAI/tsai
#
# Let's get started.
# ## sklearn-type API (<10k samples) 🚶🏻♂️
# ### Classifier
# +
# Univariate classification with sklearn-type API
dsid = 'OliveOil'
X_train, y_train, X_valid, y_valid = get_UCR_data(dsid) # Download the UCR dataset
# Computes MiniRocket features using the original (non-PyTorch) MiniRocket code.
# It then sends them to a sklearn's RidgeClassifier (linear classifier).
model = MiniRocketClassifier()
timer.start(False)
model.fit(X_train, y_train)
t = timer.stop()
print(f'valid accuracy : {model.score(X_valid, y_valid):.3%} time: {t}')
# -
# Multivariate classification with sklearn-type API
dsid = 'LSST'
X_train, y_train, X_valid, y_valid = get_UCR_data(dsid)
model = MiniRocketClassifier()
timer.start(False)
model.fit(X_train, y_train)
t = timer.stop()
print(f'valid accuracy : {model.score(X_valid, y_valid):.3%} time: {t}')
# One way to try to improve performance is to use an ensemble (that uses majority vote). Bear in mind that the ensemble will take longer since multiple models will be fitted.
# Multivariate classification ensemble with sklearn-type API
dsid = 'LSST'
X_train, y_train, X_valid, y_valid = get_UCR_data(dsid)
model = MiniRocketVotingClassifier(n_estimators=5)
timer.start(False)
model.fit(X_train, y_train)
t = timer.stop()
print(f'valid accuracy : {model.score(X_valid, y_valid):.3%} time: {t}')
# In this case, we see an increase in accuracy although this may not be the case with other datasets.
# Once a model is trained, you can always save it for future inference:
dsid = 'LSST'
X_train, y_train, X_valid, y_valid = get_UCR_data(dsid)
model = MiniRocketClassifier()
model.fit(X_train, y_train)
model.save(f'MiniRocket_{dsid}')
del model
model = load_minirocket(f'MiniRocket_{dsid}')
print(f'valid accuracy : {model.score(X_valid, y_valid):.3%}')
# ### Regressor
# Univariate regression with sklearn-type API
from sklearn.metrics import mean_squared_error
dsid = 'Covid3Month'
X_train, y_train, X_valid, y_valid = get_Monash_regression_data(dsid)
rmse_scorer = make_scorer(mean_squared_error, greater_is_better=False)
model = MiniRocketRegressor(scoring=rmse_scorer)
timer.start(False)
model.fit(X_train, y_train)
t = timer.stop()
y_pred = model.predict(X_valid)
rmse = mean_squared_error(y_valid, y_pred, squared=False)
print(f'valid rmse : {rmse:.5f} time: {t}')
# Univariate regression ensemble with sklearn-type API
from sklearn.metrics import mean_squared_error
dsid = 'Covid3Month'
X_train, y_train, X_valid, y_valid = get_Monash_regression_data(dsid)
rmse_scorer = make_scorer(mean_squared_error, greater_is_better=False)
model = MiniRocketVotingRegressor(n_estimators=5, scoring=rmse_scorer)
timer.start(False)
model.fit(X_train, y_train)
t = timer.stop()
y_pred = model.predict(X_valid)
rmse = mean_squared_error(y_valid, y_pred, squared=False)
print(f'valid rmse : {rmse:.5f} time: {t}')
# Multivariate regression with sklearn-type API
from sklearn.metrics import mean_squared_error
dsid = 'AppliancesEnergy'
X_train, y_train, X_valid, y_valid = get_Monash_regression_data(dsid)
rmse_scorer = make_scorer(mean_squared_error, greater_is_better=False)
model = MiniRocketRegressor(scoring=rmse_scorer)
timer.start(False)
model.fit(X_train, y_train)
t = timer.stop()
y_pred = model.predict(X_valid)
rmse = mean_squared_error(y_valid, y_pred, squared=False)
print(f'valid rmse : {rmse:.5f} time: {t}')
# Multivariate regression ensemble with sklearn-type API
from sklearn.metrics import mean_squared_error
dsid = 'AppliancesEnergy'
X_train, y_train, X_valid, y_valid = get_Monash_regression_data(dsid)
rmse_scorer = make_scorer(mean_squared_error, greater_is_better=False)
model = MiniRocketVotingRegressor(n_estimators=5, scoring=rmse_scorer)
timer.start(False)
model.fit(X_train, y_train)
t = timer.stop()
y_pred = model.predict(X_valid)
rmse = mean_squared_error(y_valid, y_pred, squared=False)
print(f'valid rmse : {rmse:.5f} time: {t}')
# We'll also save this model for future inference:
# Multivariate regression ensemble with sklearn-type API
from sklearn.metrics import mean_squared_error
dsid = 'AppliancesEnergy'
X_train, y_train, X_valid, y_valid = get_Monash_regression_data(dsid)
rmse_scorer = make_scorer(mean_squared_error, greater_is_better=False)
model = MiniRocketVotingRegressor(n_estimators=5, scoring=rmse_scorer)
model.fit(X_train, y_train)
model.save(f'MRVRegressor_{dsid}')
del model
model = load_minirocket(f'MRVRegressor_{dsid}')
y_pred = model.predict(X_valid)
rmse = mean_squared_error(y_valid, y_pred, squared=False)
print(f'valid rmse : {rmse:.5f}')
# ## Pytorch implementation (any # samples) 🏃
# ### Offline feature calculation
# In the offline calculation, all features will be calculated in a first stage and then passed to the dataloader that will create batches. This features will ramain the same throughout training.
#
# ⚠️ In order to avoid leakage when using the offline feature calculation, it's important to fit MiniRocketFeatures using just the train samples.
# Create the MiniRocket features and store them in memory.
dsid = 'LSST'
X, y, splits = get_UCR_data(dsid, split_data=False)
mrf = MiniRocketFeatures(X.shape[1], X.shape[2]).to(default_device())
X_train = X[splits[0]]
mrf.fit(X_train)
X_feat = get_minirocket_features(X, mrf, chunksize=1024, to_np=True)
X_feat.shape, type(X_feat)
# 👀 Note that X_train may be a np.ndarray or a torch.Tensor. In this case we'll pass a np.ndarray.
#
# If a torch.Tensor is passed, the model will move it to the right device (cuda) if necessary, so that it matches the model.
# We'll save this model, as we'll need it to create features in the future.
PATH = Path("./models/MRF.pt")
PATH.parent.mkdir(parents=True, exist_ok=True)
torch.save(mrf.state_dict(), PATH)
# As you can see the shape of the minirocket features is [sample_size x n_features x 1]. The last dimension (1) is added because `tsai` expects input data to have 3 dimensions, although in this case there's no longer a temporal dimension.
#
# Once the features are calculated, we'll need to train a Pytorch model. We'll use a simple linear model:
# +
# Using tsai/fastai, create DataLoaders for the features in X_feat.
tfms = [None, TSClassification()]
batch_tfms = TSStandardize(by_sample=True)
dls = get_ts_dls(X_feat, y, splits=splits, tfms=tfms, batch_tfms=batch_tfms)
# model is a linear classifier Head
model = build_ts_model(MiniRocketHead, dls=dls)
model.head
# +
# Using tsai/fastai, create DataLoaders for the features in X_feat.
tfms = [None, TSClassification()]
batch_tfms = TSStandardize(by_sample=True)
dls = get_ts_dls(X_feat, y, splits=splits, tfms=tfms, batch_tfms=batch_tfms)
# model is a linear classifier Head
model = build_ts_model(MiniRocketHead, dls=dls)
# Drop into fastai and use it to find a good learning rate.
learn = Learner(dls, model, metrics=accuracy, cbs=ShowGraph())
learn.lr_find()
# -
# As above, use tsai to bring X_feat into fastai, and train.
tfms = [None, TSClassification()]
batch_tfms = TSStandardize(by_sample=True)
dls = get_ts_dls(X_feat, y, splits=splits, tfms=tfms, batch_tfms=batch_tfms)
model = build_ts_model(MiniRocketHead, dls=dls)
learn = Learner(dls, model, metrics=accuracy, cbs=ShowGraph())
timer.start()
learn.fit_one_cycle(10, 3e-4)
timer.stop()
# We'll now save the learner for inference:
PATH = Path('./models/MRL.pkl')
PATH.parent.mkdir(parents=True, exist_ok=True)
learn.export(PATH)
# #### Inference:
# For inference we'll need to follow the same process as before:
#
# 1. Create the features
# 2. Create predictions for those features
# Let's recreate mrf (MiniRocketFeatures) to be able to create new features:
mrf = MiniRocketFeatures(X.shape[1], X.shape[2]).to(default_device())
PATH = Path("./models/MRF.pt")
mrf.load_state_dict(torch.load(PATH))
# We'll create new features. In this case we'll use the valid set to confirm the predictions accuracy matches the one at the end of training, but you can use any data:
new_feat = get_minirocket_features(X[splits[1]], mrf, chunksize=1024, to_np=True)
new_feat.shape, type(new_feat)
# We'll now load the saved learner:
PATH = Path('./models/MRL.pkl')
learn = load_learner(PATH, cpu=False)
# and pass the newly created features
probas, _, preds = learn.get_X_preds(new_feat)
preds
skm.accuracy_score(y[splits[1]], preds)
# Ok, so the predictions match the ones at the end of training as this accuracy is the same on we got in the end.
# ### Online feature calculation
# MiniRocket can also be used online, re-calculating the features each minibatch. In this scenario, you do not calculate fixed features one time. The online mode is a bit slower than the offline scanario, but offers more flexibility. Here are some potential uses:
#
# * You can experiment with different scaling techniques (no standardization, standardize by sample, normalize, etc).
# * You can use data augmentation is applied to the original time series.
# * Another use of online calculation is to experiment with training the kernels and biases.
# To do this requires modifications to the MRF code.
tfms = [None, TSClassification()]
batch_tfms = TSStandardize(by_sample=True)
dls = get_ts_dls(X, y, splits=splits, tfms=tfms, batch_tfms=batch_tfms)
model = build_ts_model(MiniRocket, dls=dls)
learn = Learner(dls, model, metrics=accuracy, cbs=ShowGraph())
learn.lr_find()
# Notice 2 important differences with the offline scenario:
#
# * in this case we pass X to the dataloader instead of X_tfm. The featurew will be calculated within the model.
# * we use MiniRocket instead of MiniRocketHead. MiniRocket is a Pytorch version that calculates features on the fly before passing them to a linear head.
tfms = [None, TSClassification()]
batch_tfms = TSStandardize(by_sample=True)
dls = get_ts_dls(X, y, splits=splits, tfms=tfms, batch_tfms=batch_tfms)
model = build_ts_model(MiniRocket, dls=dls)
model
tfms = [None, TSClassification()]
batch_tfms = TSStandardize(by_sample=True)
dls = get_ts_dls(X, y, splits=splits, tfms=tfms, batch_tfms=batch_tfms)
model = build_ts_model(MiniRocket, dls=dls)
learn = Learner(dls, model, metrics=accuracy, cbs=ShowGraph())
timer.start()
learn.fit_one_cycle(10, 3e-4)
timer.stop()
# Since we calculate the minirocket features within the model, we now have the option to use data augmentation for example:
# MiniRocket with data augmentation
tfms = [None, TSClassification()]
batch_tfms = [TSStandardize(by_sample=True), TSMagScale(), TSWindowWarp()]
dls = get_ts_dls(X, y, splits=splits, tfms=tfms, batch_tfms=batch_tfms)
model = build_ts_model(MiniRocket, dls=dls)
learn = Learner(dls, model, metrics=accuracy, cbs=[ShowGraph()])
learn.fit_one_cycle(20, 3e-4)
# In this case, we can see that using MiniRocket (Pytorch implementation) with data augmentation achieves an accuracy of 69%+, compared to the sklearn-API implementation which is around 65%.
# Once you have trained the model, you can always save if for future use. We just need to export the learner:
PATH = Path('./models/MiniRocket_aug.pkl')
PATH.parent.mkdir(parents=True, exist_ok=True)
learn.export(PATH)
del learn
# #### Inference
# Let's first recreate the learner:
PATH = Path('./models/MiniRocket_aug.pkl')
learn = load_learner(PATH, cpu=False)
# We are now ready to generate predictions. We'll confirm it worls well with the valid dataset:
probas, _, preds = learn.get_X_preds(X[splits[1]])
preds
# We can see that the validation loss & metrics are the same we had when we saved it.
skm.accuracy_score(y[splits[1]], preds)
# # Conclusion ✅
# MiniRocket is a new type of algorithm that is significantly faster than any other method of comparable accuracy (including Rocket), and significantly more accurate than any other method of even roughly-similar computational expense.
#
# `tsai` supports the 2 variations of MiniRocket introduced in this notebook. A cpu version (that can be used with relatively small datasets, with <10k samples) and a gpu (Pytorch) version that can be used with datasets of any size. The Pytorch version can be used in an offline mode (pre-calculating all features before fitting the model) or in an online mode (calculating features on the fly).
#
# We believe MiniRocket is a great new tool, and encourange you to try it in your next Time Series Classification or Regression task.
|
tutorial_nbs/10_Time_Series_Classification_and_Regression_with_MiniRocket.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# language: python
# name: python388jvsc74a57bd0862565ed761fef69c13c358536270023be20fe86f4b6648eb3a25208bbc233f0
# ---
import pandas as pd
import os
# +
import pandas as pd
import os
import glob
# use glob to get all the csv files
# in the folder
path = '/Users/parthshah/Documents/Northeastern/Spring2022/BigDataAnalytics/Assignment_1/Assignment_1/assignment_1/data/raw/NOAA/'
print(path)
csv_files = glob.glob(os.path.join(path, "*.csv"))
print(len(csv_files))
# loop over the list of csv files
df = pd.read_csv('/Users/parthshah/Documents/Northeastern/Spring2022/BigDataAnalytics/Assignment_1/Assignment_1/assignment_1/data/raw/NOAA/StormEvents_details-ftp_v1.0_d1977_c20210803.csv')
list1 = []
for f in csv_files:
if 'details' in f:
df = pd.read_csv(f)
list1.append(df)
NOOA_Data_Merged = pd.concat(list1)
# -
NOOA_Data_Merged[NOOA_Data_Merged["EVENT_ID"] == 835047].columns.to_list()
result.to_csv('NOAA_All_Data.csv',index = False)
NOOA_Data_Merged[NOOA_Data_Merged['EVENT_ID'] == 728503]
catlog = pd.read_csv('https://raw.githubusercontent.com/MIT-AI-Accelerator/eie-sevir/master/CATALOG.csv')
catlog['time_utc'].max()
catlog['time_utc'].min()
NOOA_Data_Merged.columns.to_list()
catlog['time_utc'] = pd.to_datetime(catlog['time_utc'])
catlog_storm_events = catlog[catlog['event_id'].isna() == False]
catlog_storm_events.reset_index(inplace = True)
catlog_storm_events['event_id']
DEF = pd.merge(catlog_storm_events,NOOA_Data_Merged,left_on = "event_id", right_on = "EVENT_ID")
DEF = pd.merge(NOOA_Data_Merged,catlog_storm_events,left_on = "EVENT_ID", right_on = "event_id",how = "left")
NOOA_Data_Merged = NOOA_Data_Merged[NOOA_Data_Merged["END_YEARMONTH"]> 201706]
NOOA_Data_Merged = NOOA_Data_Merged[NOOA_Data_Merged["END_YEARMONTH"] < 201912]
NOOA_Data_Merged = NOOA_Data_Merged.reset_index()
NOOA_Data_Merged.to_csv("NOOA_2017_2019.csv",index = False)
catlog_list = catlog_storm_events.columns.to_list()
NOAA_List = NOOA_Data_Merged.columns.to_list()
catlog_data = DEF[catlog_list]
NOAA_Data = DEF[NOAA_List]
print(catlog_data.shape)
print(NOAA_Data.shape)
catlog_data.to_csv("catlog_data.csv",index = False)
NOAA_Data.to_csv("NOAA_Data.csv",index = False)
catlog[catlog['event_id'] == 835047]
ir107/2019/SEVIR_IR107_STORMEVENTS_2019_0101_0...
ir107/2019/SEVIR_IR107_STORMEVENTS_2019_0101_0630.h5
vil/2019/SEVIR_VIL_STORMEVENTS_2019_0101_0630.h5
ir069/2019/SEVIR_IR069_STORMEVENTS_2019_0101_0630.h5
vis/2019/SEVIR_VIS_STORMEVENTS_2019_0601_0630.h5
ght/2019/SEVIR_LGHT_ALLEVENTS_2019_0601_0701.h5
# +
aws s3 cp --no-sign-request s3://sevir/data/ir107/2019/SEVIR_IR107_STORMEVENTS_2019_0101_0630.h5 SEVIR_IR107_STORMEVENTS_2019_0101_0630.h5
aws s3 cp --no-sign-request s3://sevir/data/vil/2019/SEVIR_VIL_STORMEVENTS_2019_0101_0630.h5 vil/2019/SEVIR_VIL_STORMEVENTS_2019_0101_0630.h5
aws s3 cp --no-sign-request s3://sevir/data/vis/2019/SEVIR_VIS_STORMEVENTS_2019_0601_0630.h5 SEVIR_VIS_STORMEVENTS_2019_0601_0630.h5
aws s3 cp --no-sign-request s3://sevir/data/ir069/2019/SEVIR_IR069_STORMEVENTS_2019_0101_0630.h5 SEVIR_IR069_STORMEVENTS_2019_0101_0630.h5
|
notebooks/NOAA_Sample.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="54TDqPjGFCSw" colab_type="code" colab={}
# + id="jPTalRawFFMe" colab_type="code" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 193} outputId="27685046-3fbf-45e9-f49c-dd3d67a720a2" executionInfo={"status": "ok", "timestamp": 1561725493204, "user_tz": -330, "elapsed": 17303, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-soqutdudy8A/AAAAAAAAAAI/AAAAAAAAE-U/fYLvFF1OU5s/s64/photo.jpg", "userId": "10015902151693891742"}}
from google.colab import files
uploaded = files.upload()
for fn in uploaded.keys():
print('User uploaded file "{name}" with length {length} bytes'.format(
name=fn, length=len(uploaded[fn])))
# + id="czx2EouJFbRT" colab_type="code" colab={}
# !cd sample_data
# + id="nOCEB-YRFxif" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="30b6d38c-1394-4c28-854a-da78bfc64998" executionInfo={"status": "ok", "timestamp": 1561725446619, "user_tz": -330, "elapsed": 718, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-soqutdudy8A/AAAAAAAAAAI/AAAAAAAAE-U/fYLvFF1OU5s/s64/photo.jpg", "userId": "10015902151693891742"}}
pwd
# + id="bzLmZGlTFzNu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="eb8e8990-8563-4116-b8b2-943a145e8a04" executionInfo={"status": "ok", "timestamp": 1561725500105, "user_tz": -330, "elapsed": 1904, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-soqutdudy8A/AAAAAAAAAAI/AAAAAAAAE-U/fYLvFF1OU5s/s64/photo.jpg", "userId": "10015902151693891742"}}
# ls
# + id="aIXf6k71E-bZ" colab_type="code" colab={}
import pandas as pd
import numpy as np
import spacy
from tqdm import tqdm
import re
import time
import pickle
pd.set_option('display.max_colwidth', 200)
# + id="xtQyDmKOE-bb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="eabb406f-2a6a-4f2c-8ead-31245cbe9f32" executionInfo={"status": "ok", "timestamp": 1561725523814, "user_tz": -330, "elapsed": 646, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-soqutdudy8A/AAAAAAAAAAI/AAAAAAAAE-U/fYLvFF1OU5s/s64/photo.jpg", "userId": "10015902151693891742"}}
# read data
train = pd.read_csv("train_2kmZucJ.csv")
test = pd.read_csv("test_oJQbWVk.csv")
train.shape, test.shape
# + id="ipGO9PHXE-bd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="666c0240-57e2-42d1-9988-3f5b6ff146e0" executionInfo={"status": "ok", "timestamp": 1561725526693, "user_tz": -330, "elapsed": 731, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-soqutdudy8A/AAAAAAAAAAI/AAAAAAAAE-U/fYLvFF1OU5s/s64/photo.jpg", "userId": "10015902151693891742"}}
train['label'].value_counts(normalize = True)
# + id="_-rR5YIjE-bg" colab_type="code" colab={}
#Here, 1 represents a negative tweet while 0 represents a non-negative tweet.
# + id="qnnrXefVE-bh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="784668da-1cf3-48c9-c260-b6d1b8818c34" executionInfo={"status": "ok", "timestamp": 1561725528918, "user_tz": -330, "elapsed": 450, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-soqutdudy8A/AAAAAAAAAAI/AAAAAAAAE-U/fYLvFF1OU5s/s64/photo.jpg", "userId": "10015902151693891742"}}
train.head()
# + id="9Fm6nFQuE-bj" colab_type="code" colab={}
# remove URL's from train and test
train['clean_tweet'] = train['tweet'].apply(lambda x: re.sub(r'http\S+', '', x))
test['clean_tweet'] = test['tweet'].apply(lambda x: re.sub(r'http\S+', '', x))
# + id="cQbY0qX2E-bl" colab_type="code" colab={}
# remove punctuation marks
punctuation = '!"#$%&()*+-/:;<=>?@[\\]^_`{|}~'
train['clean_tweet'] = train['clean_tweet'].apply(lambda x: ''.join(ch for ch in x if ch not in set(punctuation)))
test['clean_tweet'] = test['clean_tweet'].apply(lambda x: ''.join(ch for ch in x if ch not in set(punctuation)))
# convert text to lowercase
train['clean_tweet'] = train['clean_tweet'].str.lower()
test['clean_tweet'] = test['clean_tweet'].str.lower()
# remove numbers
train['clean_tweet'] = train['clean_tweet'].str.replace("[0-9]", " ")
test['clean_tweet'] = test['clean_tweet'].str.replace("[0-9]", " ")
# remove whitespaces
train['clean_tweet'] = train['clean_tweet'].apply(lambda x:' '.join(x.split()))
test['clean_tweet'] = test['clean_tweet'].apply(lambda x: ' '.join(x.split()))
# + id="U7O33tGXE-bm" colab_type="code" colab={}
nlp = spacy.load("en_core_web_sm")
# + id="kYDj3MueE-bo" colab_type="code" colab={}
# import spaCy's language model
# nlp = spacy.load('en', disable=['parser', 'ner'])
# function to lemmatize text
def lemmatization(texts):
output = []
for i in texts:
s = [token.lemma_ for token in nlp(i)]
output.append(' '.join(s))
return output
# + id="uN3uzFAlE-bp" colab_type="code" colab={}
train['clean_tweet'] = lemmatization(train['clean_tweet'])
test['clean_tweet'] = lemmatization(test['clean_tweet'])
# + id="sFF_gag1E-bs" colab_type="code" colab={}
train.sample(10)
# + id="vY-3umnmE-bu" colab_type="code" colab={}
# !pip install "tensorflow_hub==0.4.0"
# !pip install "tf-nightly"
# + id="dkkdkGrcE-bw" colab_type="code" colab={}
import tensorflow_hub as hub
import tensorflow as tf
elmo = hub.Module("https://tfhub.dev/google/elmo/2", trainable=True)
# + id="8jXJoYkOE-by" colab_type="code" colab={}
elmo = hub.Module("https://tfhub.dev/google/elmo/2", trainable=True)
embeddings = elmo(["the cat is on the mat", "dogs are in the fog"],signature="default",as_dict=True)["elmo"]
embeddings
# + id="JnoGMYLcE-b0" colab_type="code" colab={}
embeddings.shape
# + [markdown] id="lIA7_4f9E-b3" colab_type="text"
# ## The output is a 3 dimensional tensor of shape (1, 8, 1024):
#
# The first dimension of this tensor represents the number of training samples. This is 1 in our case
# The second dimension represents the maximum length of the longest string in the input list of strings. Since we have only 1 string in our input list, the size of the 2nd dimension is equal to the length of the string – 8
# The third dimension is equal to the length of the ELMo vector
# Hence, every word in the input sentence has an ELMo vector of size 1024.
# + id="mZ-5tbLvE-b3" colab_type="code" colab={}
def elmo_vectors(x):
embeddings = elmo(x.tolist(), signature="default", as_dict=True)["elmo"]
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
sess.run(tf.tables_initializer())
# return average of ELMo features
return sess.run(tf.reduce_mean(embeddings,1))
# + id="mWC7T910E-b5" colab_type="code" colab={}
list_train = [train[i:i+100] for i in range(0,train.shape[0],100)]
list_test = [test[i:i+100] for i in range(0,test.shape[0],100)]
# + id="le8zICXyE-b7" colab_type="code" colab={}
list_train
# + id="C3C2jSgzE-cA" colab_type="code" colab={}
# Extract ELMo embeddings
elmo_train = [elmo_vectors(x['clean_tweet']) for x in list_train]
elmo_test = [elmo_vectors(x['clean_tweet']) for x in list_test]
# + id="Ut6S9KjHE-cD" colab_type="code" colab={}
elmo_train_new = np.concatenate(elmo_train, axis = 0)
elmo_test_new = np.concatenate(elmo_test, axis = 0)
# + id="G9k5bHfKE-cE" colab_type="code" colab={}
# save elmo_train_new
pickle_out = open("elmo_train_03032019.pickle","wb")
pickle.dump(elmo_train_new, pickle_out)
pickle_out.close()
# save elmo_test_new
pickle_out = open("elmo_test_03032019.pickle","wb")
pickle.dump(elmo_test_new, pickle_out)
pickle_out.close()
# + id="V8OwXe67E-cH" colab_type="code" colab={}
# load elmo_train_new
pickle_in = open("elmo_train_03032019.pickle", "rb")
elmo_train_new = pickle.load(pickle_in)
# load elmo_train_new
pickle_in = open("elmo_test_03032019.pickle", "rb")
elmo_test_new = pickle.load(pickle_in)
# + id="1Nv5NBCtE-cI" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
xtrain, xvalid, ytrain, yvalid = train_test_split(elmo_train_new,
train['label'],
random_state=42,
test_size=0.2)
# + id="wy48wF5IE-cK" colab_type="code" colab={}
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import f1_score
lreg = LogisticRegression()
lreg.fit(xtrain, ytrain)
# + id="VskfAoAgE-cL" colab_type="code" colab={}
preds_valid = lreg.predict(xvalid)
# + id="6D7-VgDME-cM" colab_type="code" colab={}
f1_score(yvalid, preds_valid)
# + id="u5PMPM6jE-cO" colab_type="code" colab={}
# make predictions on test set
preds_test = lreg.predict(elmo_test_new)
# + id="Glso9wnZE-cP" colab_type="code" colab={}
# prepare submission dataframe
sub = pd.DataFrame({'id':test['id'], 'label':preds_test})
# write predictions to a CSV file
sub.to_csv("sub_lreg.csv", index=False)
# + id="tGctZXDQE-cQ" colab_type="code" colab={}
|
spam_ham_classfication.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
x = 11
x & 1
2**4
8 >> 2
# x(x-1) = x with its lowest set bit erased
x=10
x&(x-1)
x=8
x&(x-1)
# Best ways to save time:
# 1. Processing multiple bits at a time.
# 2. Caching results in an array-based lookup table
# x&~(x-1) = the lowest set bit of x
x = 9
x&~(x-1)
x = 8
x&~(x-1)
# 4.2
# 1001 = 9
# 3210
# Swap index 2 and 0
def swap_bits(x,i,j):
ith = x >> i & 1
jth = x >> j & 1
if ith != jth:
x = ((1 << i) | (1 << j)) ^ x
return x
swap_bits(9, 0, 2)
str(bin(43)[2:])[::-1]
# +
# 4.3
# +
# Reverse the bits in a word
# 10000111 -> 11100001 == 135 -> 225
# My way:
import math
def reverse_bits(x):
if x == 0:
return 0
return (x & 1) << int(math.log(x, 2)) | reverse_bits(x >> 1)
reverse_bits(135)
# -
# Their efficient cache way:
import array
array.array('i', [1, 2])
array
|
Chapter 4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
library(ggplot2) # Data visualization
library(readr) # CSV file I/O, e.g. the read_csv function
library(gridExtra)
library(grid)
library(plyr)
# Load the dataset
iris=read.csv('../input/Iris.csv')
# First let's get a random sampling of the data
iris[sample(nrow(iris),10),]
# +
# Density & Frequency analysis - Histogram
# Sepal length
HisSl <- ggplot(data=iris, aes(x=SepalLengthCm))+
geom_histogram(binwidth=0.2, color="black", aes(fill=Species)) +
xlab("Sepal Length (cm)") +
ylab("Frequency") +
theme(legend.position="none")+
ggtitle("Histogram of Sepal Length")+
geom_vline(data=iris, aes(xintercept = mean(SepalLengthCm)),linetype="dashed",color="grey")
# Sepal width
HistSw <- ggplot(data=iris, aes(x=SepalWidthCm)) +
geom_histogram(binwidth=0.2, color="black", aes(fill=Species)) +
xlab("Sepal Width (cm)") +
ylab("Frequency") +
theme(legend.position="none")+
ggtitle("Histogram of Sepal Width")+
geom_vline(data=iris, aes(xintercept = mean(SepalWidthCm)),linetype="dashed",color="grey")
# Petal length
HistPl <- ggplot(data=iris, aes(x=PetalLengthCm))+
geom_histogram(binwidth=0.2, color="black", aes(fill=Species)) +
xlab("Petal Length (cm)") +
ylab("Frequency") +
theme(legend.position="none")+
ggtitle("Histogram of Petal Length")+
geom_vline(data=iris, aes(xintercept = mean(PetalLengthCm)),
linetype="dashed",color="grey")
# Petal width
HistPw <- ggplot(data=iris, aes(x=PetalWidthCm))+
geom_histogram(binwidth=0.2, color="black", aes(fill=Species)) +
xlab("Petal Width (cm)") +
ylab("Frequency") +
theme(legend.position="right" )+
ggtitle("Histogram of Petal Width")+
geom_vline(data=iris, aes(xintercept = mean(PetalWidthCm)),linetype="dashed",color="grey")
# Plot all visualizations
grid.arrange(HisSl + ggtitle(""),
HistSw + ggtitle(""),
HistPl + ggtitle(""),
HistPw + ggtitle(""),
nrow = 2,
top = textGrob("Iris Frequency Histogram",
gp=gpar(fontsize=15))
)
# +
# Density plot
DhistPl <- ggplot(iris, aes(x=PetalLengthCm, colour=Species, fill=Species)) +
geom_density(alpha=.3) +
geom_vline(aes(xintercept=mean(PetalLengthCm), colour=Species),linetype="dashed",color="grey", size=1)+
xlab("Petal Length (cm)") +
ylab("Density")+
theme(legend.position="none")
DhistPw <- ggplot(iris, aes(x=PetalWidthCm, colour=Species, fill=Species)) +
geom_density(alpha=.3) +
geom_vline(aes(xintercept=mean(PetalWidthCm), colour=Species),linetype="dashed",color="grey", size=1)+
xlab("Petal Width (cm)") +
ylab("Density")
DhistSw <- ggplot(iris, aes(x=SepalWidthCm, colour=Species, fill=Species)) +
geom_density(alpha=.3) +
geom_vline(aes(xintercept=mean(SepalWidthCm), colour=Species), linetype="dashed",color="grey", size=1)+
xlab("Sepal Width (cm)") +
ylab("Density")+
theme(legend.position="none")
DhistSl <- ggplot(iris, aes(x=SepalLengthCm, colour=Species, fill=Species)) +
geom_density(alpha=.3) +
geom_vline(aes(xintercept=mean(SepalLengthCm), colour=Species),linetype="dashed", color="grey", size=1)+
xlab("Sepal Length (cm)") +
ylab("Density")+
theme(legend.position="none")
# Plot all density visualizations
grid.arrange(DhistSl + ggtitle(""),
DhistSw + ggtitle(""),
DhistPl + ggtitle(""),
DhistPw + ggtitle(""),
nrow = 2,
top = textGrob("Iris Density Plot",
gp=gpar(fontsize=15))
)
# +
# Boxplot - Petal length vs Species
ggplot(iris, aes(Species, PetalLengthCm, fill=Species)) +
geom_boxplot()+
scale_y_continuous("Petal Length (cm)", breaks= seq(0,30, by=.5))+
labs(title = "Iris Petal Length Box Plot", x = "Species")
# +
# Boxplot
BpSl <- ggplot(iris, aes(Species, SepalLengthCm, fill=Species)) +
geom_boxplot()+
scale_y_continuous("Sepal Length (cm)", breaks= seq(0,30, by=.5))+
theme(legend.position="none")
BpSw <- ggplot(iris, aes(Species, SepalWidthCm, fill=Species)) +
geom_boxplot()+
scale_y_continuous("Sepal Width (cm)", breaks= seq(0,30, by=.5))+
theme(legend.position="none")
BpPl <- ggplot(iris, aes(Species, PetalLengthCm, fill=Species)) +
geom_boxplot()+
scale_y_continuous("Petal Length (cm)", breaks= seq(0,30, by=.5))+
theme(legend.position="none")
BpPw <- ggplot(iris, aes(Species, PetalWidthCm, fill=Species)) +
geom_boxplot()+
scale_y_continuous("Petal Width (cm)", breaks= seq(0,30, by=.5))+
labs(title = "Iris Box Plot", x = "Species")
# Plot all visualizations
grid.arrange(BpSl + ggtitle(""),
BpSw + ggtitle(""),
BpPl + ggtitle(""),
BpPw + ggtitle(""),
nrow = 2,
top = textGrob("Sepal and Petal Box Plot",
gp=gpar(fontsize=15))
)
# +
# Violin plot - Shows mean, median for interquartile range with width of the shape representing the number of points at a particular value by width of the shapes
VpSl <- ggplot(iris, aes(Species, SepalLengthCm, fill=Species)) +
geom_violin(aes(color = Species), trim = T)+
scale_y_continuous("Sepal Length", breaks= seq(0,30, by=.5))+
geom_boxplot(width=0.1)+
theme(legend.position="none")
VpSw <- ggplot(iris, aes(Species, SepalWidthCm, fill=Species)) +
geom_violin(aes(color = Species), trim = T)+
scale_y_continuous("Sepal Width", breaks= seq(0,30, by=.5))+
geom_boxplot(width=0.1)+
theme(legend.position="none")
VpPl <- ggplot(iris, aes(Species, PetalLengthCm, fill=Species)) +
geom_violin(aes(color = Species), trim = T)+
scale_y_continuous("Petal Length", breaks= seq(0,30, by=.5))+
geom_boxplot(width=0.1)+
theme(legend.position="none")
VpPw <- ggplot(iris, aes(Species, PetalWidthCm, fill=Species)) +
geom_violin(aes(color = Species), trim = T)+
scale_y_continuous("Petal Width", breaks= seq(0,30, by=.5))+
geom_boxplot(width=0.1)+
labs(title = "Iris Box Plot", x = "Species")
# Plot all visualizations
grid.arrange(VpSl + ggtitle(""),
VpSw + ggtitle(""),
VpPl + ggtitle(""),
VpPw + ggtitle(""),
nrow = 2,
top = textGrob("Sepal and Petal Violin Plot",
gp=gpar(fontsize=15))
)
# +
# Scatterplot
ggplot(data = iris, aes(x = PetalLengthCm, y = PetalWidthCm))+
xlab("Petal Length")+
ylab("Petal Width") +
geom_point(aes(color = Species,shape=Species))+
geom_smooth(method='lm')+
ggtitle("Petal Length vs Width")
library(car)
scatterplot(iris$PetalLengthCm,iris$PetalWidthCm)
# -
# Scatterplot - Sepal Length vs Width
ggplot(data=iris, aes(x = SepalLengthCm, y = SepalWidthCm)) +
geom_point(aes(color=Species, shape=Species)) +
xlab("Sepal Length") +
ylab("Sepal Width") +
ggtitle("Sepal Length vs Width")
# +
# Pairwise correlation
library(GGally)
ggpairs(data = iris[1:4],
title = "Iris Correlation Plot",
upper = list(continuous = wrap("cor", size = 5)),
lower = list(continuous = "smooth")
)
# +
# Heatmap
irisMatix <- as.matrix(iris[1:150, 1:4])
irisTransposedMatrix <- t(irisMatix)[,nrow(irisMatix):1]
image(1:4, 1:150, irisTransposedMatrix)
# -
|
iris/iris-dataset-classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Multi-Region IoT - Persist Data
#
# Persist data accross regions by using [Amazon Simple Queue Service](https://docs.aws.amazon.com/AWSSimpleQueueService/latest/SQSDeveloperGuide/welcome.html) (SQS) or DynamoDB global tables.
# ## Libraries
# +
import boto3
import datetime
import json
import logging
import time
from AWSIoTPythonSDK.MQTTLib import AWSIoTMQTTClient
from os.path import join
# -
# #### Note: If you get an error that the AWSIoTPythonSDK is not installed, install the SDK with the command below and import the libraries again!
# !pip install AWSIoTPythonSDK -t .
# Restore variable that have been defined in the notebook which has been used to create the ACM PCA setup.
# %store -r config
print("config: {}".format(json.dumps(config, indent=4, default=str)))
# ## Global Vars
#
# For several actions in AWS IoT topic rules a service role is required. This roles allows the IoT Core to access other services. A role has already been created by the CloudFormation stack for the master region.
#
# The arn for this role can be found in the outputs section of your CloudFormation stack next to the key **IoTAccessServicesRoleArn**.
#
# Set the variable **topic_rule_role_arn** to contain this arn.
# +
topic_rule_role_arn = 'YOUR_ROLE_ARN_HERE'
topic_rule_name_sqs = 'IoTMRCrossRegionSQSRule'
topic_rule_name_sns = 'IoTMRCrossRegionSNSRule'
queue_name = 'IoTCrossRegion'
topic_name = 'IoTCrossRegion'
# -
# ## IoT Endpoint
# To connect a device to the AWS IoT master region we need to get the iot endpoint.
c_iot = boto3.client('iot', region_name = config['aws_region_master'])
# Get the iot endpoint.
response = c_iot.describe_endpoint(endpointType='iot:Data-ATS')
iot_endpoint = response['endpointAddress']
print("iot_endpoint: {}".format(iot_endpoint))
# ## Transfer data with Amazon Simple Queue Service (SQS)
#
# By using SQS in a topic rule data can be send across regions.
#
# An SQS queue in the slave region will be created and a topic rule in the master region which transfers incoming messages to the SQS queue in the slave region.
# SQS client in slave region.
c_sqs = boto3.client('sqs', region_name = config['aws_region_slave'])
# ### Queue operations
# Create the queue and get the queue url. The queue url is required to create the IoT topic rule.
#
# #### Create the queue
response = c_sqs.create_queue(QueueName=queue_name)
print("response: {}".format(json.dumps(response, indent=4, default=str)))
# #### Get the queue url
response = c_sqs.get_queue_url(QueueName=queue_name)
print("response: {}\n".format(json.dumps(response, indent=4, default=str)))
queue_url = response['QueueUrl']
print("queue_url: {}".format(queue_url))
# ## Topic rule
#
# Create the rule the topic rule.
# Create the topic rule
# +
response = c_iot.create_topic_rule(
ruleName=topic_rule_name_sqs,
topicRulePayload={
'awsIotSqlVersion': '2016-03-23',
'sql': 'SELECT * FROM \'cmd/+/cross/region\'',
'actions': [{
'sqs': {
'roleArn': topic_rule_role_arn,
'queueUrl': queue_url,
'useBase64': False
}
}],
'ruleDisabled': False
}
)
print("response: {}".format(json.dumps(response, indent=4, default=str)))
# -
# ## Verify
# Get the topic rule to verify that it has been created successfully.
response = c_iot.get_topic_rule(
ruleName=topic_rule_name_sqs
)
print("response: {}".format(json.dumps(response, indent=4, default=str)))
# ## Transfer data with Amazon Simple Notification Service (SNS) and AWS Lambda
#
# Another option to transfer data across regions is the use of SNS in combination with a Lambda function. In this example an SNS topic in the master region will be created. A Lambda function in the slave region was already created by CloudFormation. The Lambda endpoint from the slave region will be subscribed to the SNS topic.
c_sns = boto3.client('sns', region_name = config['aws_region_master'])
# Create the topic.
response = c_sns.create_topic(Name=topic_name)
print("response: {}".format(json.dumps(response, indent=4, default=str)))
# Get the topic arn.
topic_arn = response['TopicArn']
print("topic_arn: {}".format(topic_arn))
# ### Subscribe to SNS topic
#
# The Lambda function in the slave region has been created by the slave CloudFormation stack.
#
# Set the variable **lambda_arn** to this arn.
#
# The arn for the Lambda can be found in the outputs section of your CloudFormation stack next to the key **CrossRegionLambdaFunctionArn**.
lambda_arn = 'YOUR_LAMBDA_ARN_HERE'
lambda_name = lambda_arn.split(':')[-1]
statement_id = str(int(time.time()))
print("lambda_name: {} statement_id: {}".format(lambda_name, statement_id))
# ### Add permission to the Lambda function
#
# To allow SNS to invoke the lambda function a permission must be added to the function.
c_lambda = boto3.client('lambda', region_name = config['aws_region_slave'])
# +
response = c_lambda.add_permission(
FunctionName=lambda_name,
StatementId=statement_id,
Action='lambda:invokeFunction',
Principal='sns.amazonaws.com',
SourceArn=topic_arn
)
print("response: {}".format(json.dumps(response, indent=4, default=str)))
# +
response = c_sns.subscribe(
TopicArn=topic_arn,
Protocol='lambda',
Endpoint=lambda_arn,
ReturnSubscriptionArn=True
)
print("response: {}\n".format(json.dumps(response, indent=4, default=str)))
subscription_arn = response['SubscriptionArn']
print("subscription_arn: {}".format(subscription_arn))
# -
# ## Create topic rule
#
# Create a topic rule to forward messages to the SNS topic.
# +
response = c_iot.create_topic_rule(
ruleName=topic_rule_name_sns,
topicRulePayload={
'awsIotSqlVersion': '2016-03-23',
'sql': 'SELECT * FROM \'cmd/+/cross/region\'',
'actions': [{
'sns': {
'targetArn': topic_arn,
'roleArn': topic_rule_role_arn,
'messageFormat': 'RAW'
}
}],
'ruleDisabled': False
}
)
print("response: {}".format(json.dumps(response, indent=4, default=str)))
# -
# ## Verify
# Get the topic rule to verify that it has been created successfully.
response = c_iot.get_topic_rule(ruleName=topic_rule_name_sns)
print("response: {}".format(json.dumps(response, indent=4, default=str)))
# ## Connect a Device
# Connect a device that you created earlier to the message broker from AWS IoT and send some messages. These messages will be forwarded by the topic rules that you created to SQS as well as SNS.
# +
thing_name = 'thing-mr04'
root_ca = 'AmazonRootCA1.pem'
device_key_file = '{}.device.key.pem'.format(thing_name)
device_cert_file = '{}.device.cert.pem'.format(thing_name)
# AWS IoT Python SDK needs logging
logger = logging.getLogger("AWSIoTPythonSDK.core")
#logger.setLevel(logging.DEBUG)
logger.setLevel(logging.INFO)
streamHandler = logging.StreamHandler()
formatter = logging.Formatter("[%(asctime)s - %(levelname)s - %(filename)s:%(lineno)s - %(funcName)s - %(message)s")
streamHandler.setFormatter(formatter)
logger.addHandler(streamHandler)
myAWSIoTMQTTClient = None
myAWSIoTMQTTClient = AWSIoTMQTTClient(thing_name)
myAWSIoTMQTTClient.configureEndpoint(iot_endpoint, 8883)
myAWSIoTMQTTClient.configureCredentials(root_ca,
join(config['PCA_directory'], device_key_file),
join(config['PCA_directory'], device_cert_file))
# AWSIoTMQTTClient connection configuration
myAWSIoTMQTTClient.configureAutoReconnectBackoffTime(1, 32, 20)
myAWSIoTMQTTClient.configureOfflinePublishQueueing(-1) # Infinite offline Publish queueing
myAWSIoTMQTTClient.configureDrainingFrequency(2) # Draining: 2 Hz
myAWSIoTMQTTClient.configureConnectDisconnectTimeout(10) # 10 sec
myAWSIoTMQTTClient.configureMQTTOperationTimeout(5) # 5 sec
# Connect and reconnect to AWS IoT
try:
myAWSIoTMQTTClient.connect()
except Exception as e:
logger.error('{}'.format(e))
time.sleep(5)
myAWSIoTMQTTClient.connect()
# -
# ## Publish messages
# Publish some messages that should be transferred to the SQS queue in the other region.
#
# **Hint:** Before publishing message subscribe in the test client in the master region in the AWS IoT Core console to the topic `cmd/+/cross/region`. By doing so you can verify that your messages are reaching the IoT Core.
# +
topic = 'cmd/{}/cross/region'.format(thing_name)
print("topic: {}".format(topic))
for i in range(5):
date_time = datetime.datetime.now().isoformat()
message = {"thing_name": "{}".format(thing_name), "date_time": date_time, "i": i}
print("publish: message: {}".format(message))
myAWSIoTMQTTClient.publish(topic, json.dumps(message), 0)
time.sleep(1)
# -
# ## Poll the SQS queue
#
# To verify that the messages has been sent to SQS in the slave region poll the queue for messages. You should get messages from the queue. Feel free to execute polling multiple times.
# +
# Long poll for message on provided SQS queue
response = c_sqs.receive_message(
QueueUrl=queue_url,
AttributeNames=[
'All'
],
MaxNumberOfMessages=10,
MessageAttributeNames=[
'All'
],
WaitTimeSeconds=20
)
print("queue_url: {}\n".format(queue_url))
for message in response['Messages']:
body = message['Body']
message_id = message['MessageId']
#print(message)
print("message_id: {}\nbody: {}\n".format(message_id, body))
# -
# ## SNS/Lambda
#
# To verify that message have been sent also to the slave region with SNS and Lambda watch at CloudWatch in the slave region for the logs of your Lambda function.
#
# Logs can be found in `CloudWatch -> Logs -> /aws/lambda/<LAMBDA_FUNCTION_NAME`.
# ## Disconnet
# Disconnect the device from AWS IoT Core.
myAWSIoTMQTTClient.disconnect()
# ## Clean Up
# Clean up your environment:
#
# * Remove permissions from the Lambda function
# * Unsubscribe the Lambda from the SNS topic
# * Delete SNS topic
# * Delete IoT topic rules
response = c_lambda.remove_permission(
FunctionName=lambda_name,
StatementId=statement_id
)
print("response: {}".format(json.dumps(response, indent=4, default=str)))
response = c_sns.unsubscribe(SubscriptionArn=subscription_arn)
print("response: {}".format(json.dumps(response, indent=4, default=str)))
response = c_sns.delete_topic(TopicArn=topic_arn)
print("response: {}".format(json.dumps(response, indent=4, default=str)))
response = c_iot.delete_topic_rule(ruleName=topic_rule_name_sqs)
print("response: {}".format(json.dumps(response, indent=4, default=str)))
response = c_iot.delete_topic_rule(ruleName=topic_rule_name_sns)
print("response: {}".format(json.dumps(response, indent=4, default=str)))
response = c_sns.delete_topic(TopicArn=topic_arn)
print("response: {}".format(json.dumps(response, indent=4, default=str)))
|
jupyter/10_Multi_Region_Data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Automatic Differentiation with torch.autograd
# When training neural networks, the most frequently used algorithm is back propagation. In this algorithm, parameters (model weights) are adjusted according to the gradient of the loss function with respect to the given parameter.
#
# To compute those gradients, PyTorch has a built-in differentiation engine called torch.autograd. It supports automatic computation of gradient for any computational graph.
#
# Consider the simplest one-layer neural network, with input x, parameters w and b, and some loss function. It can be defined in PyTorch in the following manner:
# +
import torch
x = torch.ones(5) # input tensor
y = torch.zeros(3) #expected output
w = torch.randn(5,3, requires_grad=True)
b = torch.randn(3, requires_grad=True)
z = torch.matmul(x,w) + b
loss = torch.nn.functional.binary_cross_entropy_with_logits(z,y)
# -
# ## Tensors, Functions and Computational graph
#
# This code defines the following computational graph:
# 
# In this network, w and b are parameters, which we need to optimize. Thus, we need to be able to compute the gradients of loss function with respect to those variables. In order to do that, we set the requires_grad property of those tensors.
#
# You can set the value of requires_grad when creating a tensor, or later by using x.requires_grad_(True) method.
#
# A function that we apply to tensors to construct computational graph is in fact an object of class Function. This object knows how to compute the function in the forward direction, and also how to compute its derivative during the backward propagation step. A reference to the backward propagation function is stored in grad_fn property of a tensor. You can find more information of Function in the documentation.
print('Gradient function for z = ', z.grad_fn)
print('Gradient function for loss = ', loss.grad_fn)
# # Computing Gradients
#
# To optimize weights of parameters in the neural network, we need to compute the derivatives of our loss function with respect to parameters, namely, we need
# dL/dW and dL/db
# under some fixed values of x and y. To compute those derivatives, we call loss.backward(), and then retrieve the values from w.grad and b.grad:
loss.backward()
print(w.grad)
print(b.grad)
# Note
# - We can only obtain the grad properties for the leaf nodes of the computational graph, which have requires_grad property set to True. For all other nodes in our graph, gradients will not be available.
# - We can only perform gradient calculations using backward once on a given graph, for performance reasons. If we need to do several backward calls on the same graph, we need to pass retain_graph=True to the backward call.
# ## Disabling Gradient Tracking
# By default, all tensors with requires_grad=True are tracking their computational history and support gradient computation. However, there are some cases when we do not need to do that, for example, when we have trained the model and just want to apply it to some input data, i.e. we only want to do forward computations through the network. We can stop tracking computations by surrounding our computation code with torch.no_grad() block:
z = torch.matmul(x, w)+b
print(z.requires_grad)
with torch.no_grad():
z = torch.matmul(x,w)+b
print(z.requires_grad)
# Another way to achieve the same result is to use the detach() method on the tensor:
z = torch.matmul(x,w)+b
z_det = z.detach()
print(z_det.requires_grad)
# There are reasons you might want to disable gradient tracking:
# - To mark some parameters in your neural network at frozen parameters. This is a very common scenario for finetuning a pretrained network
# - To speed up computations when you are only doing forward pass, because computations on tensors that do not track gradients would be more efficient.
# ## More on Computational Graphs
# Conceptually, autograd keeps a record of data (tensors) and all executed operations (along with the resulting new tensors) in a directed acyclic graph (DAG) consisting of Function objects. In this DAG, leaves are the input tensors, roots are the output tensors. By tracing this graph from roots to leaves, you can automatically compute the gradients using the chain rule.
#
# In a forward pass, autograd does two things simultaneously:
# - run the requested operation to compute a resulting tensor
# - maintain the operation’s gradient function in the DAG.
#
# The backward pass kicks off when .backward() is called on the DAG root. autograd then:
# - computes the gradients from each .grad_fn,
# - accumulates them in the respective tensor’s .grad attribute
# - using the chain rule, propagates all the way to the leaf tensors.
#
# DAGs are dynamic in PyTorch An important thing to note is that the graph is recreated from scratch; after each .backward() call, autograd starts populating a new graph. This is exactly what allows you to use control flow statements in your model; you can change the shape, size and operations at every iteration if needed.
# ## Optional Reading: Tensor Gradients and Jacobian Products
# In many cases, we have a scalar loss function, and we need to compute the gradient with respect to some parameters. However, there are cases when the output function is an arbitrary tensor. In this case, PyTorch allows you to compute so-called Jacobian product, and not the actual gradient.
#
# 
inp = torch.eye(5, requires_grad=True)
print(inp)
out = (inp+1).pow(2)
out.backward(torch.ones_like(inp), retain_graph=True)
print('Frist call \n', inp.grad)
out.backward(torch.ones_like(inp), retain_graph=True)
print('Second call\n', inp.grad)
inp.grad.zero_()
out.backward(torch.ones_like(inp), retain_graph=True)
print('Call after zeroing gradients \n', inp.grad)
# Notice that when we call backward for the second time with the same argument, the value of the gradient is different. This happens because when doing backward propagation, PyTorch accumulates the gradients, i.e. the value of computed gradients is added to the grad property of all leaf nodes of computational graph. If you want to compute the proper gradients, you need to zero out the grad property before. In real-life training an optimizer helps us to do this.
# Note
# - Previously we were calling backward() function without parameters. This is essentially equivalent to calling backward(torch.tensor(1.0)), which is a useful way to compute the gradients in case of a scalar-valued function, such as loss during neural network training.
#
# Further Reading
# - Autograd Mechanics https://pytorch.org/docs/stable/notes/autograd.html
|
part-ax-pytorch/06-autograd.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 作業
# ### 請根據不同的 HOUSETYPE_MODE 對 AMT_CREDIT 繪製 Histogram
# +
# Import 需要的套件
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns # 另一個繪圖-樣式套件
# %matplotlib inline
plt.style.use('ggplot')
import warnings
warnings.filterwarnings('ignore')
# 設定 data_path
dir_data = './data/'
# -
f_app = os.path.join(dir_data, 'application_train.csv')
print('Path of read in data: %s' % (f_app))
app_train = pd.read_csv(f_app)
app_train.head()
# Continous to Discrete
app_train['AMT_CREDIT_BIN'] = pd.cut(app_train['AMT_CREDIT'],bins = 10) #切 10 組
print(app_train['AMT_CREDIT_BIN'].value_counts())
# +
"""
Your Code Here
"""
unique_house_type = app_train['NAME_HOUSING_TYPE'].unique()
nrows = len(unique_house_type)
ncols = nrows // 2
plt.figure(figsize=(10,30))
for i in range(len(unique_house_type)):
plt.subplot(nrows, ncols, i+1)
"""
Your Code Here
"""
#app_train.loc[ , ].hist()
app_train.loc[app_train['NAME_HOUSING_TYPE'] == unique_house_type[i],'AMT_CREDIT'].hist(label = 'AMT_CREDIT')
plt.legend()
plt.title(str(unique_house_type[i]))
plt.show()
# -
|
Day_014_subplot.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pysam
import pandas as pd
from tqdm.auto import tqdm
import numpy as np
import itertools
bam_file = "/home/dbeb/btech/bb1160039/scratch/project/295_possorted_genome_bam.bam"
bai_file = "/home/dbeb/btech/bb1160039/scratch/project/295_possorted_genome_bam.bam.bai"
samf = pysam.Samfile(bam_file, "rb")
replicon_dict = dict([[replicon, {'seq_start_pos': 0,'seq_end_pos': length}] for replicon, length in zip(samf.references, samf.lengths)])
print(replicon_dict['1']['seq_start_pos'])
print(replicon_dict['1']['seq_end_pos'])
samfile = pysam.AlignmentFile(bam_file, "rb", index_filename = bai_file)
# +
#read all non-genes
# x=['1','2','3','4','5','6','7','8','9','10','11','12','13','14','15','16','17','18','19','X','Y']
x=['1','2','3','4','5','6','7','8','9','10','11','12','13','14','15','16','17','18','19','20','21','22','X','Y']
list_tags_all = []
for i in tqdm(range(0,len(x))):
count=0
for read in samfile.fetch(x[i],replicon_dict[x[i]]['seq_start_pos'],replicon_dict[x[i]]['seq_end_pos']):
try:
if (read.has_tag("GX")==False and read.get_tag("NH")==1):
list_tags_all.append([read.get_tag("CB"),read.get_tag("UB"),str("gene" + "-" + str(read.reference_name) + "-" +str(read.get_reference_positions()[0])+'-'+ str(len(read.get_reference_positions())))])
except KeyError:
continue
# -
# %%time
list_tags_all_rm_dup = list(list_tags_all for list_tags_all,_ in itertools.groupby(list_tags_all))
print(len(list_tags_all))
print(len(list_tags_all_rm_dup))
# +
start=0
val=int(list_tags_all_rm_dup[start][2].split("-")[2])
for i in tqdm(range(1,len(list_tags_all_rm_dup))):
if (int(list_tags_all_rm_dup[i][2].split("-")[2]) - val>50):
if i-start>1:
#update the inner elements
for inner in list_tags_all_rm_dup[start:i]:
inner[2] = list_tags_all_rm_dup[start][2]
#update start to point to this pos
start = i
#update val to the val at this pos
val = int(list_tags_all_rm_dup[i][2].split("-")[2])
# -
# %%time
list_tags_all_rm_dup_final = list(list_tags_all_rm_dup for list_tags_all_rm_dup,_ in itertools.groupby(list_tags_all_rm_dup))
n=len(list_tags_all_rm_dup_final)
n
# %%time
df_all_p1 = pd.DataFrame(list_tags_all_rm_dup_final[:int(n/10)], columns=["celltag","moltag","pseudoname"])
df_all_p2 = pd.DataFrame(list_tags_all_rm_dup_final[int(n/10):int(2*n/10)], columns=["celltag","moltag","pseudoname"])
df_all_p3 = pd.DataFrame(list_tags_all_rm_dup_final[int(2*n/10):int(3*n/10)], columns=["celltag","moltag","pseudoname"])
df_all_p4 = pd.DataFrame(list_tags_all_rm_dup_final[int(3*n/10):int(4*n/10)], columns=["celltag","moltag","pseudoname"])
df_all_p5 = pd.DataFrame(list_tags_all_rm_dup_final[int(4*n/10):int(5*n/10)], columns=["celltag","moltag","pseudoname"])
df_all_p6 = pd.DataFrame(list_tags_all_rm_dup_final[int(5*n/10):int(6*n/10)], columns=["celltag","moltag","pseudoname"])
df_all_p7 = pd.DataFrame(list_tags_all_rm_dup_final[int(6*n/10):int(7*n/10)], columns=["celltag","moltag","pseudoname"])
df_all_p8 = pd.DataFrame(list_tags_all_rm_dup_final[int(7*n/10):int(8*n/10)], columns=["celltag","moltag","pseudoname"])
df_all_p9 = pd.DataFrame(list_tags_all_rm_dup_final[int(8*n/10):int(9*n/10)], columns=["celltag","moltag","pseudoname"])
df_all_p10 = pd.DataFrame(list_tags_all_rm_dup_final[int(9*n/10):], columns=["celltag","moltag","pseudoname"])
# %%time
c1 = df_all_p1['celltag'].value_counts()
c2 = df_all_p2['celltag'].value_counts()
c3 = df_all_p3['celltag'].value_counts()
c4 = df_all_p4['celltag'].value_counts()
c5 = df_all_p5['celltag'].value_counts()
c6 = df_all_p6['celltag'].value_counts()
c7 = df_all_p7['celltag'].value_counts()
c8 = df_all_p8['celltag'].value_counts()
c9 = df_all_p9['celltag'].value_counts()
c10 = df_all_p10['celltag'].value_counts()
c11 = df_all_p1['pseudoname'].value_counts()
c22 = df_all_p2['pseudoname'].value_counts()
c33 = df_all_p3['pseudoname'].value_counts()
c44 = df_all_p4['pseudoname'].value_counts()
c55 = df_all_p5['pseudoname'].value_counts()
c66 = df_all_p6['pseudoname'].value_counts()
c77 = df_all_p7['pseudoname'].value_counts()
c88 = df_all_p8['pseudoname'].value_counts()
c99 = df_all_p9['pseudoname'].value_counts()
c1010 = df_all_p10['pseudoname'].value_counts()
# %time
df_all_p1_subset = df_all_p1[df_all_p1["celltag"].isin(c1[c1>10].index)]
df_all_p1_subset = df_all_p1_subset[df_all_p1_subset["pseudoname"].isin(c11[c11>10].index)]
df_all_p2_subset = df_all_p2[df_all_p2["celltag"].isin(c2[c2>10].index)]
df_all_p2_subset = df_all_p2_subset[df_all_p2_subset["pseudoname"].isin(c22[c22>10].index)]
df_all_p3_subset = df_all_p3[df_all_p3["celltag"].isin(c3[c3>10].index)]
df_all_p3_subset = df_all_p3_subset[df_all_p3_subset["pseudoname"].isin(c33[c33>10].index)]
df_all_p4_subset = df_all_p4[df_all_p4["celltag"].isin(c4[c4>10].index)]
df_all_p4_subset = df_all_p4_subset[df_all_p4_subset["pseudoname"].isin(c44[c44>10].index)]
df_all_p5_subset = df_all_p5[df_all_p5["celltag"].isin(c5[c5>10].index)]
df_all_p5_subset = df_all_p5_subset[df_all_p5_subset["pseudoname"].isin(c55[c55>10].index)]
df_all_p6_subset = df_all_p6[df_all_p6["celltag"].isin(c6[c6>10].index)]
df_all_p6_subset = df_all_p6_subset[df_all_p6_subset["pseudoname"].isin(c66[c66>10].index)]
df_all_p7_subset = df_all_p7[df_all_p7["celltag"].isin(c7[c7>10].index)]
df_all_p7_subset = df_all_p7_subset[df_all_p7_subset["pseudoname"].isin(c77[c77>10].index)]
df_all_p8_subset = df_all_p8[df_all_p8["celltag"].isin(c8[c8>10].index)]
df_all_p8_subset = df_all_p8_subset[df_all_p8_subset["pseudoname"].isin(c88[c88>10].index)]
df_all_p9_subset = df_all_p9[df_all_p9["celltag"].isin(c9[c9>10].index)]
df_all_p9_subset = df_all_p9_subset[df_all_p9_subset["pseudoname"].isin(c99[c99>10].index)]
df_all_p10_subset = df_all_p10[df_all_p10["celltag"].isin(c10[c10>10].index)]
df_all_p10_subset = df_all_p10_subset[df_all_p10_subset["pseudoname"].isin(c1010[c1010>10].index)]
# %%time
counts_p1=df_all_p1_subset.groupby(['pseudoname','celltag']).size().unstack('celltag', fill_value=0)
counts_p2=df_all_p2_subset.groupby(['pseudoname','celltag']).size().unstack('celltag', fill_value=0)
counts_p3=df_all_p3_subset.groupby(['pseudoname','celltag']).size().unstack('celltag', fill_value=0)
counts_p4=df_all_p4_subset.groupby(['pseudoname','celltag']).size().unstack('celltag', fill_value=0)
counts_p5=df_all_p5_subset.groupby(['pseudoname','celltag']).size().unstack('celltag', fill_value=0)
counts_p6=df_all_p6_subset.groupby(['pseudoname','celltag']).size().unstack('celltag', fill_value=0)
counts_p7=df_all_p7_subset.groupby(['pseudoname','celltag']).size().unstack('celltag', fill_value=0)
counts_p8=df_all_p8_subset.groupby(['pseudoname','celltag']).size().unstack('celltag', fill_value=0)
counts_p9=df_all_p9_subset.groupby(['pseudoname','celltag']).size().unstack('celltag', fill_value=0)
counts_p10=df_all_p10_subset.groupby(['pseudoname','celltag']).size().unstack('celltag', fill_value=0)
cell_bcode=set(counts_p1.columns).intersection(set(counts_p2.columns)).intersection(set(counts_p3.columns)).intersection(set(counts_p4.columns)).intersection(set(counts_p5.columns)).intersection(set(counts_p6.columns)).intersection(set(counts_p7.columns)).intersection(set(counts_p8.columns)).intersection(set(counts_p9.columns)).intersection(set(counts_p10.columns))
# %%time
counts_p1 = counts_p1[cell_bcode]
counts_p2 = counts_p2[cell_bcode]
counts_p3 = counts_p3[cell_bcode]
counts_p4 = counts_p4[cell_bcode]
counts_p5 = counts_p5[cell_bcode]
counts_p6 = counts_p6[cell_bcode]
counts_p7 = counts_p7[cell_bcode]
counts_p8 = counts_p8[cell_bcode]
counts_p9 = counts_p9[cell_bcode]
counts_p10 = counts_p10[cell_bcode]
# %%time
counts_full = pd.concat([counts_p1,counts_p2, counts_p3, counts_p4, counts_p5, counts_p6, counts_p7, counts_p8, counts_p9, counts_p10])
# +
# counts_full.to_csv("/home/dbeb/btech/bb1160039/scratch/project/counts_non_genes_pbmc_10k_rmmulti.csv")
# -
|
examples/processing codes/.ipynb_checkpoints/Processing for Non-genes Brain Human II-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!/usr/bin/env python
# encoding: utf-8
"""
PegInHole
Use the scene:UR5PegInHole2.ttt
@Author: Zane
@Contact: <EMAIL>
@File: UR5PegInHole.py
@Time: 2019-07-29 15:55
"""
import sim as vrep
import sys
import numpy as np
import math
import matplotlib.pyplot as mpl
import time
##### Definition of parameters
RAD2DEG = math.pi / 180
# Define step size and timeout of simulation
step = 0.005
TIMEOUT = 5000
# Define parameters of joint
jointNum = 6
jointName = 'UR5_joint'
##### Python connect to the V-REP client
print('Program started')
# Close potential connections
vrep.simxFinish(-1)
clientID = vrep.simxStart('127.0.0.1', 19997, True, True, 5000, 5)
print("Connection success")
# Start simulation
vrep.simxStartSimulation(clientID, vrep.simx_opmode_blocking)
print("Simulation start")
i = 0
ur5ready = 0
while i < TIMEOUT and ur5ready == 0:
i = i + 1
errorCode, ur5ready = vrep.simxGetIntegerSignal(clientID, 'UR5READY', vrep.simx_opmode_blocking)
time.sleep(step)
if i >= TIMEOUT:
print('An error occurred in your V-REP server')
vrep.simxFinish(clientID)
##### Obtain the handle
jointHandle = np.zeros((jointNum,), dtype=np.int) # 注意是整型
for i in range(jointNum):
errorCode, returnHandle = vrep.simxGetObjectHandle(clientID, jointName + str(i + 1), vrep.simx_opmode_blocking)
jointHandle[i] = returnHandle
time.sleep(2)
errorCode, holeHandle = vrep.simxGetObjectHandle(clientID, 'Hole', vrep.simx_opmode_blocking)
errorCode, ikTipHandle = vrep.simxGetObjectHandle(clientID, 'UR5_ikTip', vrep.simx_opmode_blocking)
errorCode, connectionHandle = vrep.simxGetObjectHandle(clientID, 'UR5_connection', vrep.simx_opmode_blocking)
print('Handles available!')
##### Obtain the position of the hole
errorCode, targetPosition = vrep.simxGetObjectPosition(clientID, holeHandle, -1, vrep.simx_opmode_streaming)
time.sleep(0.5)
errorCode, targetPosition = vrep.simxGetObjectPosition(clientID, holeHandle, -1, vrep.simx_opmode_buffer)
print('Position available!')
###### Joint space control
# Action1 of initConfig
initConfig = [0, 22.5 * RAD2DEG, 67.5 * RAD2DEG, 0, -90 * RAD2DEG, 0]
vrep.simxPauseCommunication(clientID, True)
for i in range(jointNum):
vrep.simxSetJointTargetPosition(clientID, jointHandle[i], initConfig[i], vrep.simx_opmode_oneshot)
vrep.simxPauseCommunication(clientID, False)
# Make sure the step of simulation has been done
vrep.simxGetPingTime(clientID)
time.sleep(1)
# Get Object Quaternion
errorCode, tipQuat = vrep.simxGetObjectQuaternion(clientID, ikTipHandle, -1, vrep.simx_opmode_blocking)
# Action2 of targetPosition1
# The position of action2
targetPosition[2] = targetPosition[2] + 0.15
# Sent the signal of movement
vrep.simxPauseCommunication(clientID, 1)
vrep.simxSetIntegerSignal(clientID, 'ICECUBE_0', 21, vrep.simx_opmode_oneshot)
for i in range(1, 4):
vrep.simxSetFloatSignal(clientID, 'ICECUBE_' + str(i), targetPosition[i - 1], vrep.simx_opmode_oneshot)
for i in range(4, 8):
vrep.simxSetFloatSignal(clientID, 'ICECUBE_' + str(i), tipQuat[i - 4], vrep.simx_opmode_oneshot)
vrep.simxPauseCommunication(clientID, 0)
# Wait
j = 0
signal = 99
while j <= TIMEOUT and signal != 0:
j = j + 1
errorCode, signal = vrep.simxGetIntegerSignal(clientID, 'ICECUBE_0', vrep.simx_opmode_blocking)
time.sleep(step)
errorCode = vrep.simxSetIntegerParameter(clientID, vrep.sim_intparam_current_page, 1, vrep.simx_opmode_blocking)
# Action3 of targetPosition2
# The position2 of action3
targetPosition[2] = targetPosition[2] - 0.05
time.sleep(2)
# Sent the signal of movement
vrep.simxPauseCommunication(clientID, 1)
vrep.simxSetIntegerSignal(clientID, 'ICECUBE_0', 21, vrep.simx_opmode_oneshot)
for i in range(1, 4):
vrep.simxSetFloatSignal(clientID, 'ICECUBE_' + str(i), targetPosition[i - 1], vrep.simx_opmode_oneshot)
for i in range(4, 8):
vrep.simxSetFloatSignal(clientID, 'ICECUBE_' + str(i), tipQuat[i - 4], vrep.simx_opmode_oneshot)
vrep.simxPauseCommunication(clientID, 0)
# Wait
j = 0
signal = 99
while j <= TIMEOUT and signal != 0:
j = j + 1
errorCode, signal = vrep.simxGetIntegerSignal(clientID, 'ICECUBE_0', vrep.simx_opmode_blocking)
time.sleep(step)
time.sleep(1)
errorCode = vrep.simxSetIntegerParameter(clientID, vrep.sim_intparam_current_page, 0, vrep.simx_opmode_blocking)
time.sleep(2)
##### Stop simulation
vrep.simxStopSimulation(clientID, vrep.simx_opmode_blocking)
errorCode = vrep.simxSetIntegerSignal(clientID, 'ICECUBE_0', 1, vrep.simx_opmode_blocking)
time.sleep(0.5)
##### Close the connection to V-REP
vrep.simxFinish(clientID)
print('Program end')
# -
|
Example_UR5_PegInHole/UR5PegInHole.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Manifold Learning
# * https://jakevdp.github.io/PythonDataScienceHandbook/05.10-manifold-learning.html
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import os
import sys
p = os.path.join(os.path.dirname('__file__'), '..')
sys.path.append(p)
from common import *
sns.set()
# Manifold Learning
# * Dimensionality reduction technique
# * Given high-dimensional data, it seeks a low-dimensional representation of the data that preserves certain relationships within the data.
#
# Use cases
# * Good for nonlinear relationships (PCA fails)
#
# Manifold
# * Imagine a sheet of paper: this is a two-dimensional object that lives in our familiar three-dimensional world, and can be bent or rolled in two dimensions.
# * We can think of this sheet as a two-dimensional manifold embedded in three-dimensional space.
# * Rotating, re-orienting, or stretching the piece of paper in three-dimensional space doesn't change the flat geometry of the paper: such operations are akin to linear embeddings.
# * If you bend, curl, or crumple the paper, it is still a two-dimensional manifold, but the embedding into the three-dimensional space is no longer linear.
#
# Algorithms
# * MDS (multi-dimensional scaling)
# * Quantity preserved is the distance between every pair of points
# * LLE (locally linear embedding)
# * IsoMap (isometric mapping)
# ### Data
def make_hello(N=1000, rseed=42):
# Make a plot with "HELLO" text; save as PNG
fig, ax = plt.subplots(figsize=(4, 1))
fig.subplots_adjust(left=0, right=1, bottom=0, top=1)
ax.axis('off')
ax.text(0.5, 0.4, 'HELLO', va='center', ha='center', weight='bold', size=85)
fig.savefig('hello.png')
plt.close(fig)
# Open this PNG and draw random points from it
from matplotlib.image import imread
data = imread('hello.png')[::-1, :, 0].T
rng = np.random.RandomState(rseed)
X = rng.rand(4 * N, 2)
i, j = (X * data.shape).astype(int).T
mask = (data[i, j] < 1)
X = X[mask]
X[:, 0] *= (data.shape[0] / data.shape[1])
X = X[:N]
return X[np.argsort(X[:, 0])]
X = make_hello(1000)
colorize = dict(c=X[:, 0], cmap=plt.cm.get_cmap('rainbow', 5))
plt.scatter(X[:, 0], X[:, 1], **colorize)
plt.axis('equal');
# ### MDS (Multidimensional Scaling)
# * Quantity preserved is the distance between every pair of points.
# * Doesn't work for nonlinear embeddings
#
# * For rigid body transforms (translate, rotate, scale), the relationship between points (distance) doesn't change
# +
def rotate(X, angle):
theta = np.deg2rad(angle)
R = [[np.cos(theta), -np.sin(theta)],
[np.sin(theta), np.cos(theta)]]
return np.dot(X, R)
X2 = rotate(X, 20) + 5
plt.scatter(X2[:, 0], X2[:, 1], **colorize)
plt.axis('equal');
# -
# So lets figure out the distance between points using <b>pairwise distance</b>
#
# Pairwise Distance Matrix
# * For N points, we construct an N by N array such that entry (i, j) contains the distance between point i and point j.
from sklearn.metrics import pairwise_distances
D = pairwise_distances(X)
D.shape
plt.imshow(D, zorder=2, cmap='Blues', interpolation='nearest')
plt.colorbar();
# Given a distance matrix between points, MDS recovers a D-dimensional coordinate representation of the data.
from sklearn.manifold import MDS
model = MDS(n_components=2, dissimilarity='precomputed', random_state=1)
out = model.fit_transform(D)
plt.scatter(out[:, 0], out[:, 1], **colorize)
plt.axis('equal');
# So MDS can recover the datapoints given only a pairwise distance matrix
#
# Why is this useful?
# * It works for any number of dimensions...
# +
# Let's project to 3D
def random_projection(X, dimension=3, rseed=42):
assert dimension >= X.shape[1]
rng = np.random.RandomState(rseed)
C = rng.randn(dimension, dimension)
e, V = np.linalg.eigh(np.dot(C, C.T))
return np.dot(X, V[:X.shape[1]])
X3 = random_projection(X, 3)
X3.shape
# -
from mpl_toolkits import mplot3d
ax = plt.axes(projection='3d')
ax.scatter3D(X3[:, 0], X3[:, 1], X3[:, 2],
**colorize)
ax.view_init(azim=70, elev=50)
model = MDS(n_components=2, random_state=1)
out3 = model.fit_transform(X3)
plt.scatter(out3[:, 0], out3[:, 1], **colorize)
plt.axis('equal');
# This is essentially the goal of a manifold learning estimator: given high-dimensional embedded data, it seeks a low-dimensional representation of the data that preserves certain relationships within the data. In the case of MDS, the quantity preserved is the distance between every pair of points.
# ### LLE (Locally Linear Embedding)
# * MDS fails for nonlinear datasets, like this one
# * LLE solves this by relaxing the constraint on preserving distance between far away points
# * It focuses instead on preserving distance between nearby points
#
# 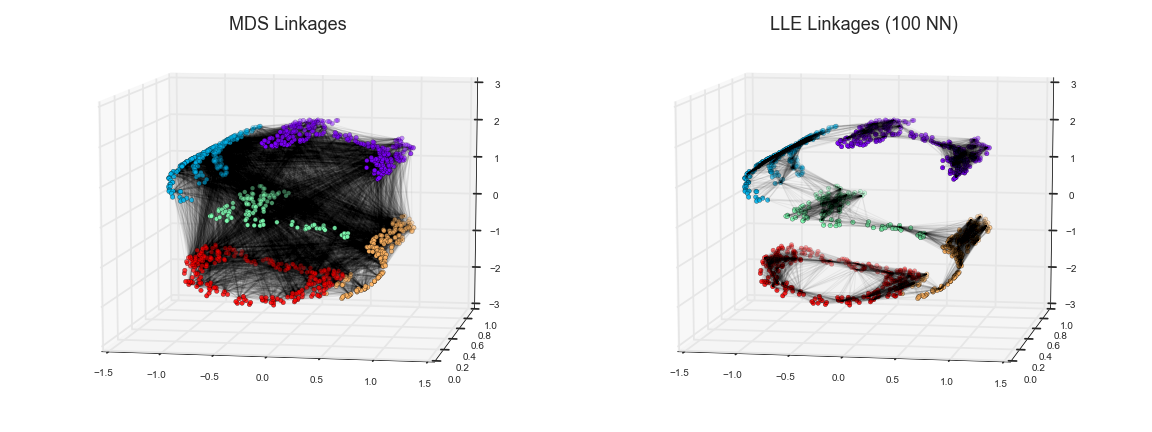
#
# * MDS: tries to preserve the distances between each pair of points in the dataset
# * LLE: rather than preserving all distances, it instead tries to preserve only the distances between neighboring points: in this case, the nearest 100 neighbors of each point.
# +
def make_hello_s_curve(X):
t = (X[:, 0] - 2) * 0.75 * np.pi
x = np.sin(t)
y = X[:, 1]
z = np.sign(t) * (np.cos(t) - 1)
return np.vstack((x, y, z)).T
XS = make_hello_s_curve(X)
# -
from mpl_toolkits import mplot3d
ax = plt.axes(projection='3d')
ax.scatter3D(XS[:, 0], XS[:, 1], XS[:, 2],
**colorize);
# Manifold Learning Fails to recover Hello (when it's warped in a non-linear way)
from sklearn.manifold import MDS
model = MDS(n_components=2, random_state=2)
outS = model.fit_transform(XS)
plt.scatter(outS[:, 0], outS[:, 1], **colorize)
plt.axis('equal');
# +
# LLE recovers Hello from the non-linearly distored representatino in higher dimensions
from sklearn.manifold import LocallyLinearEmbedding
model = LocallyLinearEmbedding(n_neighbors=100, n_components=2, method='modified',
eigen_solver='dense')
out = model.fit_transform(XS)
fig, ax = plt.subplots()
ax.scatter(out[:, 0], out[:, 1], **colorize)
ax.set_ylim(0.15, -0.15);
# -
# ### Summary
# * Manifold learning techniques are rarely used in practice
# * Use cases
# * Visualization of high-dimensional data
# * Pros
# * Preserve non-linear relationships in the data
# * Cons
# * Don't handle noise very well
# * Need to choose optimal # of neighbors
# * Computationally expensive (n^2 or n^3)
# ### Links
#
# * http://scikit-learn.org/stable/modules/manifold.html
# * https://jakevdp.github.io/PythonDataScienceHandbook/05.10-manifold-learning.html
|
theory/ManifoldLearning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Devan0369/AI-Project/blob/master/Copy_of_ALIFE_Stella_Chatbot.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="Gsv_5v7g583V" colab_type="code" colab={}
#Description: This is a 'self learning' chatbot program
# + id="FZXf8zUd6OBS" colab_type="code" colab={}
#Install the package NLTK
pip install nltk
# + id="bvDzZxZq6hyL" colab_type="code" colab={}
#Install the package newspaper3k
pip install newspaper3k
# + id="d-WbkdWc7D5r" colab_type="code" colab={}
#Import Libraries
from newspaper import Article
import random
import string
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics.pairwise import cosine_similarity
import nltk
import numpy as np
import warnings
# + id="IXX0jDqw8VUZ" colab_type="code" colab={}
#Ingore any warning messages
warnings.filterwarnings('ignore')
# + id="cX_CQCjZ8pNg" colab_type="code" colab={}
#Download the packages from NLTK
nltk.download('punkt', quiet=True)
nltk.download('wordnet', quiet=True)
# + id="6meA4J5A9W2P" colab_type="code" colab={}
#Get the article URL
article = Article('https://www.mayoclinic.org/diseases-conditions/coronavirus/symptoms-causes/syc-20479963')
article.download()
article.parse()
article.nlp()
corpus = article.text
#Print the corpus/text
print(corpus)
# + id="5uFWe2zp-ihR" colab_type="code" colab={}
#Tokenization
text = corpus
sent_tokens = nltk.sent_tokenize(text) #Convert the text into a list of sentences
#Print the list of sentences
print(sent_tokens)
# + id="jBJMKB2l_ZFP" colab_type="code" colab={}
#Create a dictornary (key:value) pair to remove punctuations
remove_punct_dict = dict( (ord(punct),None) for punct in string.punctuation)
#Print the punctuations
print(string.punctuation)
#Print the dictionary
print(remove_punct_dict)
# + id="yJX6k4OUB-CI" colab_type="code" colab={}
#Create a function to return a list of lemmatized lower case words after removing punctuations
def LemNormalize(text):
return nltk.word_tokenize(text.lower().translate(remove_punct_dict))
#Print the tokenization text
print(LemNormalize(text))
# + id="ApQtsHmjJdr7" colab_type="code" colab={}
#Keyword Matching
#Greeting Inputs
GREETING_INPUTS = ["hi","hello","hola","greetings","wassup","hey"]
#Greeting Responses back to user
GREETING_RESPONSES = ["howdy","hi","hello","hey","whatsup","hey there"]
#Function to return a random greeting response to a user greeting
def greeting(sentence):
#if the user input is a greeting, then return a randomly chosen greeting response
for word in sentence.split():
if word.lower() in GREETING_INPUTS:
return random.choice(GREETING_RESPONSES)
# + [markdown] id="NOGhPkYSNiPR" colab_type="text"
#
# + id="PrBTZG7GMfX0" colab_type="code" colab={}
#Generate the response
def response(user_response):
#The users response / query
#user_response = 'What is Coronavirus'
user_response = user_response.lower() #Make the response lower case
###Print the user query / response
#print(user_response)
#Set the chatbot response to an empty string
robo_response = ''
#Append the users response to the sentence list
sent_tokens.append(user_response)
###Print the sentence list after appending the users response
#print(sent_tokens)
#Create a TfidfVectorizer Object
TfidfVec = TfidfVectorizer(tokenizer = LemNormalize, stop_words='english')
#Convert the text to a matrix of TF-IDF features
tfidf = TfidfVec.fit_transform(sent_tokens)
###Print the TF-IDF Features
#print(tfidf)
#Get the measure of similarity (similarity scores)
vals = cosine_similarity(tfidf[-1], tfidf)
###Print the similarity scores
#print(vals)
#Get the index of the most similar text/sentense to the user response
idx = vals.argsort()[0][-2]
#Reduce the dimensionality of vals
flat = vals.flatten()
#sort the list in ascending order
flat.sort()
#Get the most similar score to the users response
score = flat[-2]
###Print the similarity score
#print(score)
#If the variable 'score' is 0 then there is no text to users response
if(score == 0):
robo_response = robo_response+"I apologise, I don't understand, kindly rephrase your questions."
else:
robo_response = robo_response+sent_tokens[idx]
#Print the chatbot response
#print(robo_response)
#Remove the users response from the senstence token list
sent_tokens.remove(user_response)
return robo_response
# + id="EBiAFniXfNjM" colab_type="code" colab={}
flag = True
print("Stella: I am an ALIFE Air Health Bot. I will help you understand all you need about COVID 19. You may exit anytime, just type Bye!")
while(flag == True):
user_response = input()
user_response = user_response.lower()
if(user_response != 'bye'):
if(user_response == 'thanks' or user_response =='thank you'):
flag=False
print("Stella: You are welcome !")
else:
if(greeting(user_response) != None):
print("Stella: "+greeting(user_response))
else:
print("Stella: "+response(user_response))
else:
flag = False
print("Stella: See you later !")
|
Copy_of_ALIFE_Stella_Chatbot.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="NQt_zSDId4ku"
# # Hypothesis Testing
# + colab={} colab_type="code" id="L-vLlLqCXRsg"
import numpy as np
import matplotlib.pyplot as plt
import random
plt.rcParams.update({'font.size': 22})
# %matplotlib inline
# + [markdown] colab_type="text" id="2mCeIN2OTdYa"
# ### Hypothesis Testing Overview
# -
# ##### Concepts Explanation with Example
# Generate the data
random.seed(2020)
math2019 = [random.normalvariate(75,5) for _ in range(900)]
math2020 = [random.normalvariate(73,5) for _ in range(900)]
# Plot the data
plt.figure(figsize=(10,6))
plt.rcParams.update({'font.size': 22})
plt.hist(math2020,bins=np.linspace(50,100,50),alpha=0.5,label="math2020")
plt.hist(math2019,bins=np.linspace(50,100,50),alpha=0.5,label="math2019")
plt.legend();
# Calcualte the statistics
from scipy import stats
stats.describe(math2020)
# ### P-value, example 1
random.seed(2020)
results = []
for _ in range(1000000):
results.append(sum([random.random() < 0.5 for i in range(6)]))
from collections import Counter
from math import factorial as factorial
counter = Counter(results)
for head in sorted(counter.keys()):
comput = counter[head]/1000000
theory = 0.5**6*factorial(6)/factorial(head)/factorial(6-head)
print("heads: {}; Computational: {}; Theoretical: {}".format(head,comput, theory))
# ### P-value, example 2
# +
from scipy.stats import f
plt.figure(figsize=(10,8))
styles = ["-",":","--","-."]
for i, [dfn, dfd] in enumerate([[20,30],[20,60],[50,30],[50,60]]):
x = np.linspace(f.ppf(0.001, dfn, dfd), f.ppf(0.999, dfn, dfd), 100)
plt.plot(x, f.pdf(x, dfn, dfd), linestyle= styles[i],
lw=4, alpha=0.6,
label='{} {}'.format(dfn,dfd))
plt.legend();
# -
plt.figure(figsize=(10,8))
[dfn, dfd] =[20,60]
x = np.linspace(f.ppf(0.001, dfn, dfd), f.ppf(0.999, dfn, dfd), 100)
plt.plot(x,
f.pdf(x, dfn, dfd),
linestyle= "--",
lw=4, alpha=0.6,
label='{} {}'.format(dfn,dfd))
right = x[x>1.5]
left = x[f.pdf(x, dfn, dfd) < f.pdf(right,dfn,dfd)[0]][0:8]
plt.fill_between(right,f.pdf(right,dfn,dfd),alpha=0.4,color="r")
plt.fill_between(left,f.pdf(left,dfn,dfd),alpha=0.4,color="r")
plt.legend();
# P-value
f.cdf(left[-1],dfn,dfd) + (1-f.cdf(right[0],dfn,dfd))
# ### t-distributions
# +
from scipy.stats import t, norm
plt.figure(figsize=(12,6))
DOFs = [2,4,8]
linestyles= [":","--","-."]
for i, df in enumerate(DOFs):
x = np.linspace(-4, 4, 100)
rv = t(df)
plt.plot(x, rv.pdf(x), 'k-', lw=2, label= "DOF = " + str(df),linestyle=linestyles[i]);
plt.plot(x,norm(0,1).pdf(x),'k-', lw=2, label="Standard Normal")
plt.legend(loc=[0.6,0.6]);
# -
# #### t-statistic and their corresponding locations
plt.figure(figsize=(10,6))
df = 5
x = np.linspace(-8, 8, 200)
rv = t(df)
plt.plot(x, rv.pdf(x), 'k-', lw=4,linestyle="--");
alphas = [0.1,0.05,0.025,0.01,0.005,0.001,0.0005]
threasholds = [1.476,2.015,2.571,3.365,4.032,5.894,6.869]
for thre, alpha in zip(threasholds,alphas):
plt.plot([thre,thre],[0,rv.pdf(thre)] ,label = "{}".format(str(alpha)),linewidth=4)
plt.legend();
plt.figure(figsize=(10,6))
df = 5
x = np.linspace(-8, 8, 200)
rv = t(df)
plt.plot(x, rv.pdf(x), 'k-', lw=4,linestyle="--");
alphas = [0.1,0.05,0.025,0.01,0.005,0.001,0.0005]
threasholds = [1.476,2.015,2.571,3.365,4.032,5.894,6.869]
for thre, alpha in zip(threasholds,alphas):
plt.plot([thre,thre],[0,rv.pdf(thre)] ,label = "{}".format(str(alpha)),linewidth=4)
plt.xlim(-2,8)
plt.ylim(0,0.15)
plt.legend();
# The t-statistic of math score example
(np.mean(math2020)-75)/(np.std(math2020)/30)
# ### Compare two-tail and one-tail significance level
# +
plt.figure(figsize=(10,6))
df = 5
x = np.linspace(-8, 8, 200)
rv = t(df)
plt.plot(x, rv.pdf(x), 'k-', lw=4,linestyle="--");
alpha=0.01
one_tail = 3.365
two_tail = 4.032
plt.plot([one_tail,one_tail],[0,rv.pdf(one_tail)] ,
label = "one_tail",linewidth=4,linestyle="--")
plt.plot([two_tail,two_tail],[0,rv.pdf(two_tail)] ,
label = "two tail",linewidth=4,color="r",linestyle=":")
plt.plot([-two_tail,-two_tail],[0,rv.pdf(two_tail)] ,
label = "two tail",linewidth=4,color="r",linestyle=":")
plt.fill_between(np.linspace(-8,-two_tail,200),
rv.pdf(np.linspace(-8,-two_tail,200)),color="g")
plt.fill_between(np.linspace(one_tail,two_tail,200),
rv.pdf(np.linspace(one_tail,two_tail,200)),color="g")
plt.ylim(0,0.02)
plt.legend();
# -
# ## SciPy Examples
# ### Example 1, t-test
from scipy import stats
stats.ttest_1samp(math2020,75.0)
# #### Two-sample t-test
np.random.seed(2020)
sample1 = np.random.normal(2,1,400)
sample2 = np.random.normal(2.1,1,400)
plt.figure(figsize=(10,6))
plt.hist(sample1,bins=np.linspace(-1,5,10),alpha=0.5,label="sample1")
plt.hist(sample2,bins=np.linspace(-1,5,10),alpha=0.5,label="sample2")
plt.legend();
stats.ttest_ind(sample1,sample2)
np.random.seed(2020)
p_values = []
for _ in range(100):
sample1 = np.random.normal(2,1,900)
sample2 = np.random.normal(2.1,1,900)
p_values.append(stats.ttest_ind(sample1,sample2)[1])
plt.figure(figsize=(10,6))
plt.boxplot(p_values);
# #### two-sample t-test, different variance
np.random.seed(2020)
sample1 = np.random.uniform(2,10,400)
sample2 = np.random.uniform(1,12,900)
plt.figure(figsize=(10,6))
plt.hist(sample1,bins=np.linspace(0,15,20),alpha=0.5,label="sample1")
plt.hist(sample2,bins=np.linspace(0,15,20),alpha=0.5,label="sample2")
plt.legend();
stats.ttest_ind(sample1,sample2,equal_var=False)
# ### Example 2, normality test
# +
from scipy.stats import t, norm
plt.figure(figsize=(12,6))
DOFs = [1,2,10]
linestyles= [":","--","-."]
for i, df in enumerate(DOFs):
x = np.linspace(-4, 4, 100)
rv = t(df)
plt.plot(x, rv.pdf(x), 'k-', lw=2, label= "DOF = " + str(df),linestyle=linestyles[i]);
plt.plot(x,norm(0,1).pdf(x),'k-', lw=2, label="Standard Normal")
plt.legend();
# -
from scipy.stats import chi2
plt.figure(figsize=(10,6))
DOFs = [4,8,16,32]
linestyles= [":","--","-.","-"]
for i, df in enumerate(DOFs):
x = np.linspace(chi2.ppf(0.01, df),chi2.ppf(0.99, df), 100)
rv = chi2(df)
plt.plot(x, rv.pdf(x), 'k-', lw=4,
label= "DOF = " + str(df),linestyle=linestyles[i]);
plt.legend();
# #### Generate samples
np.random.seed(2020)
sample1= np.random.chisquare(8,400)
sample2 = np.random.chisquare(32,400)
plt.figure(figsize=(10,6))
plt.hist(sample1,bins=np.linspace(0,60,20),alpha=0.5,label="sample1")
plt.hist(sample2,bins=np.linspace(0,60,20),alpha=0.5,label="sample2")
plt.legend();
# ##### Test the normality
from scipy.stats import shapiro, anderson
print("Results for Shapiro-Wilk Test: ")
print("Sample 1:", shapiro(sample1))
print("Sample 2:", shapiro(sample2))
print()
print("Results for Anderson-Darling Test:")
print("Sample 1:", anderson(sample1))
print("Sample 2:", anderson(sample2))
# Test a real normal distributed data
sample3 = np.random.normal(0,1,400)
print("Results for Shapiro-Wilk Test: ")
print("Sample 3:", shapiro(sample3))
print()
print("Results for Anderson-Darling Test:")
print("Sample 3:", anderson(sample3))
# ### Example 3, Goodness of Fit Test
from scipy.special import comb
P = [comb(39,3-i)*comb(13,i)/comb(52,3) for i in range(4)]
expected = [1023*p for p in P]
observed = [460,451,102,10]
x = np.array([0,1,2,3])
plt.figure(figsize=(10,6))
plt.bar(x-0.2,expected,width=0.4,label="Expected")
plt.bar(x+0.2,observed,width=0.4, label= "Observed")
plt.legend()
plt.xticks(ticks=[0,1,2,3])
plt.xlabel("Number of Hearts")
plt.ylabel("Count");
# ##### Do the test
from scipy.stats import chisquare
chisquare(observed,expected)
plt.figure(figsize=(10,6))
x = np.linspace(chi2.ppf(0.001, 3),chi2.ppf(0.999, 3), 100)
rv = chi2(3)
plt.plot(x, rv.pdf(x), 'k-', lw=4,
label= "DOF = " + str(3),linestyle="--");
# ##### Numerical evaluation
SF = np.array([120000,110300,127800,68900,79040,208000,159000,89000])
LA = np.array([65700,88340,240000,190000,45080,25900,69000,120300])
BO = np.array([87999,86340,98000,124000,113800,98000,108000,78080])
NY = np.array([300000,62010,45000,130000,238000,56000,89000,123000])
mu = np.mean(np.concatenate((SF,LA,BO,NY)))
ST = np.sum((np.concatenate((SF,LA,BO,NY)) - mu)**2)
SW = np.sum((SF-np.mean(SF))**2) +np.sum((LA-np.mean(LA))**2) + \
np.sum((BO-np.mean(BO))**2)+ np.sum((NY-np.mean(NY))**2)
SB = 8*(np.mean(SF)-mu)**2 + 8*(np.mean(LA)-mu)**2 + \
8*(np.mean(BO)-mu)**2 + 8*(np.mean(NY)-mu)**2
ST == SW+SB
print(ST,SW,SB)
F = SB/(4-1)/(SW/(4*8-4))
F
from scipy.stats import f
plt.figure(figsize=(10,6))
x = np.linspace(f.ppf(0.001, 3, 28),f.ppf(0.999, 3, 28), 100)
rv = f(dfn=3,dfd=28)
plt.plot(x, rv.pdf(x), 'k-', lw=4,linestyle="--");
from scipy.stats import f_oneway
f_oneway(LA,NY,SF,BO)
# ### Statistical Test for Time Series Model
# ##### White noise
np.random.seed(2020)
plt.figure(figsize=(10,6))
white_noise = [np.random.normal() for _ in range(100)]
plt.xlabel("Time step")
plt.ylabel("Value")
plt.plot(white_noise);
# ##### Random walk and modified random walk
plt.figure(figsize=(10,6))
np.random.seed(2020)
white_noise = [np.random.normal() for _ in range(500)]
random_walk_modified = [white_noise[0]]
for i in range(1,500):
random_walk_modified.append(random_walk_modified[-1]*0.8 \
+ white_noise[i])
random_walk = np.cumsum(white_noise)
plt.plot(white_noise, label = "white noise",linestyle=":")
plt.plot(random_walk, label = "standard random walk")
plt.plot(random_walk_modified, label = "modified random walk",linestyle="-.")
plt.xlabel("Time step")
plt.ylabel("Value")
plt.legend(loc=[0,0]);
# ##### Another 2nd order auto-regressive model
# roots in unit circle
plt.rcParams.update({'font.size': 10})
for root in np.roots([1.2,-0.6,1]):
plt.polar([0,np.angle(root)],[0,abs(root)],marker='o')
plt.rcParams.update({'font.size': 22})
# ##### oscillating behavior
plt.figure(figsize=(10,6))
np.random.seed(2020)
white_noise = [np.random.normal() for _ in range(200)]
series = [white_noise[0],white_noise[1]]
for i in range(2,200):
series.append(series[i-1]*0.6-series[i-2]*1.2 + white_noise[i])
plt.plot(series, label = "oscillating")
plt.xlabel("Time step")
plt.ylabel("Value")
plt.legend();
# ##### ADF test
from statsmodels.tsa.stattools import adfuller
adfuller(white_noise)
adfuller(random_walk)
adfuller(random_walk_modified)
# not valid for the `series` time series
adfuller(series)
# ### A/B Testing
random.seed(2020)
def build_sample():
device = "mobile" if np.random.random() < 0.6 else "desktop"
browser = "chrome" if np.random.random() < 0.9 else "IE"
wifi = "strong" if np.random.random() < 0.8 else "weak"
scheme = "warm" if np.random.random() < 0.5 else "cold"
return (device, browser, wifi, scheme)
from collections import Counter
results = [build_sample() for _ in range(100)]
counter = Counter(results)
for key in sorted(counter, key = lambda x: counter[x]):
print(key, counter[key])
# ##### more samples
results = [build_sample() for _ in range(10000)]
counter = Counter(results)
for key in sorted(counter, key = lambda x: counter[x]):
print(key, counter[key])
|
Chapter07/Chapter7.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--BOOK_INFORMATION-->
# <img align="left" style="padding-right:10px;" src="figures/PDSH-cover-small.png">
#
# *This notebook contains an excerpt from the [Python Data Science Handbook](http://shop.oreilly.com/product/0636920034919.do) by <NAME>; the content is available [on GitHub](https://github.com/jakevdp/PythonDataScienceHandbook).*
#
# *The text is released under the [CC-BY-NC-ND license](https://creativecommons.org/licenses/by-nc-nd/3.0/us/legalcode), and code is released under the [MIT license](https://opensource.org/licenses/MIT). If you find this content useful, please consider supporting the work by [buying the book](http://shop.oreilly.com/product/0636920034919.do)!*
# <!--NAVIGATION-->
# < [Operating on Data in Pandas](03.03-Operations-in-Pandas.ipynb) | [Contents](Index.ipynb) | [Hierarchical Indexing](03.05-Hierarchical-Indexing.ipynb) >
#
# <a href="https://colab.research.google.com/github/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/03.04-Missing-Values.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
#
# # Handling Missing Data
# The difference between data found in many tutorials and data in the real world is that real-world data is rarely clean and homogeneous.
# In particular, many interesting datasets will have some amount of data missing.
# To make matters even more complicated, different data sources may indicate missing data in different ways.
#
# In this section, we will discuss some general considerations for missing data, discuss how Pandas chooses to represent it, and demonstrate some built-in Pandas tools for handling missing data in Python.
# Here and throughout the book, we'll refer to missing data in general as *null*, *NaN*, or *NA* values.
# ## Trade-Offs in Missing Data Conventions
#
# There are a number of schemes that have been developed to indicate the presence of missing data in a table or DataFrame.
# Generally, they revolve around one of two strategies: using a *mask* that globally indicates missing values, or choosing a *sentinel value* that indicates a missing entry.
#
# In the masking approach, the mask might be an entirely separate Boolean array, or it may involve appropriation of one bit in the data representation to locally indicate the null status of a value.
#
# In the sentinel approach, the sentinel value could be some data-specific convention, such as indicating a missing integer value with -9999 or some rare bit pattern, or it could be a more global convention, such as indicating a missing floating-point value with NaN (Not a Number), a special value which is part of the IEEE floating-point specification.
#
# None of these approaches is without trade-offs: use of a separate mask array requires allocation of an additional Boolean array, which adds overhead in both storage and computation. A sentinel value reduces the range of valid values that can be represented, and may require extra (often non-optimized) logic in CPU and GPU arithmetic. Common special values like NaN are not available for all data types.
#
# As in most cases where no universally optimal choice exists, different languages and systems use different conventions.
# For example, the R language uses reserved bit patterns within each data type as sentinel values indicating missing data, while the SciDB system uses an extra byte attached to every cell which indicates a NA state.
# ## Missing Data in Pandas
#
# The way in which Pandas handles missing values is constrained by its reliance on the NumPy package, which does not have a built-in notion of NA values for non-floating-point data types.
#
# Pandas could have followed R's lead in specifying bit patterns for each individual data type to indicate nullness, but this approach turns out to be rather unwieldy.
# While R contains four basic data types, NumPy supports *far* more than this: for example, while R has a single integer type, NumPy supports *fourteen* basic integer types once you account for available precisions, signedness, and endianness of the encoding.
# Reserving a specific bit pattern in all available NumPy types would lead to an unwieldy amount of overhead in special-casing various operations for various types, likely even requiring a new fork of the NumPy package. Further, for the smaller data types (such as 8-bit integers), sacrificing a bit to use as a mask will significantly reduce the range of values it can represent.
#
# NumPy does have support for masked arrays – that is, arrays that have a separate Boolean mask array attached for marking data as "good" or "bad."
# Pandas could have derived from this, but the overhead in both storage, computation, and code maintenance makes that an unattractive choice.
#
# With these constraints in mind, Pandas chose to use sentinels for missing data, and further chose to use two already-existing Python null values: the special floating-point ``NaN`` value, and the Python ``None`` object.
# This choice has some side effects, as we will see, but in practice ends up being a good compromise in most cases of interest.
# ### ``None``: Pythonic missing data
#
# The first sentinel value used by Pandas is ``None``, a Python singleton object that is often used for missing data in Python code.
# Because it is a Python object, ``None`` cannot be used in any arbitrary NumPy/Pandas array, but only in arrays with data type ``'object'`` (i.e., arrays of Python objects):
# + jupyter={"outputs_hidden": true}
import numpy as np
import pandas as pd
# + jupyter={"outputs_hidden": false}
vals1 = np.array([1, None, 3, 4])
vals1
# -
# This ``dtype=object`` means that the best common type representation NumPy could infer for the contents of the array is that they are Python objects.
# While this kind of object array is useful for some purposes, any operations on the data will be done at the Python level, with much more overhead than the typically fast operations seen for arrays with native types:
# + jupyter={"outputs_hidden": false}
for dtype in ['object', 'int']:
print("dtype =", dtype)
# %timeit np.arange(1E6, dtype=dtype).sum()
print()
# -
# The use of Python objects in an array also means that if you perform aggregations like ``sum()`` or ``min()`` across an array with a ``None`` value, you will generally get an error:
# + jupyter={"outputs_hidden": false}
vals1.sum()
# -
# This reflects the fact that addition between an integer and ``None`` is undefined.
# ### ``NaN``: Missing numerical data
#
# The other missing data representation, ``NaN`` (acronym for *Not a Number*), is different; it is a special floating-point value recognized by all systems that use the standard IEEE floating-point representation:
# + jupyter={"outputs_hidden": false}
vals2 = np.array([1, np.nan, 3, 4])
vals2.dtype
# -
# Notice that NumPy chose a native floating-point type for this array: this means that unlike the object array from before, this array supports fast operations pushed into compiled code.
# You should be aware that ``NaN`` is a bit like a data virus–it infects any other object it touches.
# Regardless of the operation, the result of arithmetic with ``NaN`` will be another ``NaN``:
# + jupyter={"outputs_hidden": false}
1 + np.nan
# + jupyter={"outputs_hidden": false}
0 * np.nan
# -
# Note that this means that aggregates over the values are well defined (i.e., they don't result in an error) but not always useful:
# + jupyter={"outputs_hidden": false}
vals2.sum(), vals2.min(), vals2.max()
# -
# NumPy does provide some special aggregations that will ignore these missing values:
# + jupyter={"outputs_hidden": false}
np.nansum(vals2), np.nanmin(vals2), np.nanmax(vals2)
# -
# Keep in mind that ``NaN`` is specifically a floating-point value; there is no equivalent NaN value for integers, strings, or other types.
# ### NaN and None in Pandas
#
# ``NaN`` and ``None`` both have their place, and Pandas is built to handle the two of them nearly interchangeably, converting between them where appropriate:
# + jupyter={"outputs_hidden": false}
pd.Series([1, np.nan, 2, None])
# -
# For types that don't have an available sentinel value, Pandas automatically type-casts when NA values are present.
# For example, if we set a value in an integer array to ``np.nan``, it will automatically be upcast to a floating-point type to accommodate the NA:
# + jupyter={"outputs_hidden": false}
x = pd.Series(range(2), dtype=int)
x
# + jupyter={"outputs_hidden": false}
x[0] = None
x
# -
# Notice that in addition to casting the integer array to floating point, Pandas automatically converts the ``None`` to a ``NaN`` value.
# (Be aware that there is a proposal to add a native integer NA to Pandas in the future; as of this writing, it has not been included).
#
# While this type of magic may feel a bit hackish compared to the more unified approach to NA values in domain-specific languages like R, the Pandas sentinel/casting approach works quite well in practice and in my experience only rarely causes issues.
#
# The following table lists the upcasting conventions in Pandas when NA values are introduced:
#
# |Typeclass | Conversion When Storing NAs | NA Sentinel Value |
# |--------------|-----------------------------|------------------------|
# | ``floating`` | No change | ``np.nan`` |
# | ``object`` | No change | ``None`` or ``np.nan`` |
# | ``integer`` | Cast to ``float64`` | ``np.nan`` |
# | ``boolean`` | Cast to ``object`` | ``None`` or ``np.nan`` |
#
# Keep in mind that in Pandas, string data is always stored with an ``object`` dtype.
# ## Operating on Null Values
#
# As we have seen, Pandas treats ``None`` and ``NaN`` as essentially interchangeable for indicating missing or null values.
# To facilitate this convention, there are several useful methods for detecting, removing, and replacing null values in Pandas data structures.
# They are:
#
# - ``isnull()``: Generate a boolean mask indicating missing values
# - ``notnull()``: Opposite of ``isnull()``
# - ``dropna()``: Return a filtered version of the data
# - ``fillna()``: Return a copy of the data with missing values filled or imputed
#
# We will conclude this section with a brief exploration and demonstration of these routines.
# ### Detecting null values
# Pandas data structures have two useful methods for detecting null data: ``isnull()`` and ``notnull()``.
# Either one will return a Boolean mask over the data. For example:
# + jupyter={"outputs_hidden": true}
data = pd.Series([1, np.nan, 'hello', None])
# + jupyter={"outputs_hidden": false}
data.isnull()
# -
# As mentioned in [Data Indexing and Selection](03.02-Data-Indexing-and-Selection.ipynb), Boolean masks can be used directly as a ``Series`` or ``DataFrame`` index:
# + jupyter={"outputs_hidden": false}
data[data.notnull()]
# -
# The ``isnull()`` and ``notnull()`` methods produce similar Boolean results for ``DataFrame``s.
# ### Dropping null values
#
# In addition to the masking used before, there are the convenience methods, ``dropna()``
# (which removes NA values) and ``fillna()`` (which fills in NA values). For a ``Series``,
# the result is straightforward:
# + jupyter={"outputs_hidden": false}
data.dropna()
# -
# For a ``DataFrame``, there are more options.
# Consider the following ``DataFrame``:
# + jupyter={"outputs_hidden": false}
df = pd.DataFrame([[1, np.nan, 2],
[2, 3, 5],
[np.nan, 4, 6]])
df
# -
# We cannot drop single values from a ``DataFrame``; we can only drop full rows or full columns.
# Depending on the application, you might want one or the other, so ``dropna()`` gives a number of options for a ``DataFrame``.
#
# By default, ``dropna()`` will drop all rows in which *any* null value is present:
# + jupyter={"outputs_hidden": false}
df.dropna()
# -
# Alternatively, you can drop NA values along a different axis; ``axis=1`` drops all columns containing a null value:
# + jupyter={"outputs_hidden": false}
df.dropna(axis='columns')
# -
# But this drops some good data as well; you might rather be interested in dropping rows or columns with *all* NA values, or a majority of NA values.
# This can be specified through the ``how`` or ``thresh`` parameters, which allow fine control of the number of nulls to allow through.
#
# The default is ``how='any'``, such that any row or column (depending on the ``axis`` keyword) containing a null value will be dropped.
# You can also specify ``how='all'``, which will only drop rows/columns that are *all* null values:
# + jupyter={"outputs_hidden": false}
df[3] = np.nan
df
# + jupyter={"outputs_hidden": false}
df.dropna(axis='columns', how='all')
# -
# For finer-grained control, the ``thresh`` parameter lets you specify a minimum number of non-null values for the row/column to be kept:
# + jupyter={"outputs_hidden": false}
df.dropna(axis='rows', thresh=3)
# -
# Here the first and last row have been dropped, because they contain only two non-null values.
# ### Filling null values
#
# Sometimes rather than dropping NA values, you'd rather replace them with a valid value.
# This value might be a single number like zero, or it might be some sort of imputation or interpolation from the good values.
# You could do this in-place using the ``isnull()`` method as a mask, but because it is such a common operation Pandas provides the ``fillna()`` method, which returns a copy of the array with the null values replaced.
#
# Consider the following ``Series``:
# + jupyter={"outputs_hidden": false}
data = pd.Series([1, np.nan, 2, None, 3], index=list('abcde'))
data
# -
# We can fill NA entries with a single value, such as zero:
# + jupyter={"outputs_hidden": false}
data.fillna(0)
# -
# We can specify a forward-fill to propagate the previous value forward:
# + jupyter={"outputs_hidden": false}
# forward-fill
data.fillna(method='ffill')
# -
# Or we can specify a back-fill to propagate the next values backward:
# + jupyter={"outputs_hidden": false}
# back-fill
data.fillna(method='bfill')
# -
# For ``DataFrame``s, the options are similar, but we can also specify an ``axis`` along which the fills take place:
# + jupyter={"outputs_hidden": false}
df
# + jupyter={"outputs_hidden": false}
df.fillna(method='ffill', axis=1)
# -
# Notice that if a previous value is not available during a forward fill, the NA value remains.
# <!--NAVIGATION-->
# < [Operating on Data in Pandas](03.03-Operations-in-Pandas.ipynb) | [Contents](Index.ipynb) | [Hierarchical Indexing](03.05-Hierarchical-Indexing.ipynb) >
#
# <a href="https://colab.research.google.com/github/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/03.04-Missing-Values.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
#
|
PythonDataScienceHandbook-master/notebooks/03.04-Missing-Values.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # This example notebook shows how to create a source catalog through the JWST pipeline source_catalog step
#
# The specifics are done for a NIRCAM Grism direct image, since the catalog is needed to define the source locations in the grism observation so that the spectral bounding boxes for each order can be extracted for each object.
# %matplotlib inline
import os
import jwst
import numpy as np
from astropy.io import fits
print("Using jwst pipeline version: {}".format(jwst.__version__))
# ### I'm going to use a simulated image as an example, the cell below details its file structure
# **Make sure that you have set the JWST_NOTEBOOK_DATA environment variable in the terminal from which you started Jupyter Notebook.**
#
# The data will be read from that directory, and the pipeline should write to the current working directory, avoiding clobbers.
data_dir = os.environ['JWST_NOTEBOOK_DATA']
nircam_data= data_dir + 'data/nircam/'
grism_direct_image = nircam_data + 'nircam_grism_direct_image.fits'
fits.info(grism_direct_image)
# ### Let's take a visual look at the image we are using
from matplotlib import pyplot as plt
dirim = fits.getdata(grism_direct_image, ext=1)
xs,ys=dirim.shape
fig = plt.figure(figsize=(4,4), dpi=150)
ax = fig.add_subplot(1, 1, 1)
ax.set_adjustable('box-forced')
ax.set_title(grism_direct_image.split("/")[-1], fontsize=8)
ax.imshow(dirim, origin='lower', extent=[0,xs,0,ys], vmin=-10, vmax=10)
# ### Import the catalog creation step from the pipeline
from jwst.source_catalog import source_catalog_step
# ### Create the step object, it knows how to create the catalog when it's given a data model
sc=source_catalog_step.SourceCatalogStep()
# ### These are the default step parameters
print(sc.spec)
# ### This step only works on Drizzle products and single drizzle FITS images
# You can open the fits image into a datamodel or you can supply the step with the name of the FITS image directly
# Here we'll use a DataModel
from jwst.datamodels import DrizProductModel
dpm=DrizProductModel(grism_direct_image)
sc.process(dpm)
from astropy.table import QTable
source_catalog = 'nircam_grism_direct_image_cat.ecsv' # this name is listed in the output of the process above
# ### The INFO log suggests that's too many sources, let's see how we did
# The catalog cat be read is as an astropy quantities table
catalog = QTable.read(source_catalog, format='ascii.ecsv')
catalog
# ### Overplot the object detections on our image
# +
import matplotlib.patches as patches
dirim = fits.getdata(grism_direct_image, ext=1)
xs,ys=dirim.shape
fig = plt.figure(figsize=(4,4), dpi=150)
ax.ticklabel_format(useOffset=False)
ax = fig.add_subplot(1, 1, 1)
ax.ticklabel_format(useOffset=False)
ax.set_adjustable('box-forced')
ax.set_title(grism_direct_image.split("/")[-1], fontsize=8)
ax.imshow(dirim, origin='lower', extent=[0,xs,0,ys], vmin=-10, vmax=10)
# rectangle patches are xmin, ymin, width, height
plist1=[]
for obj in catalog:
plist1.append(patches.Circle((obj['xcentroid'].value, obj['ycentroid'].value),10))
for p in plist1:
ax.add_patch(p)
ax.imshow(dirim, origin='lower', extent=[0,xs,0,ys], vmin=-10, vmax=10)
# -
# ### Looks like we should edit the defaults so that we can restrict the detections to the visible objects
# as a reminder, these are the defaults
print(sc.spec)
# ### The defaults can be changes in the step object directly
sc.npixels=20
sc.snr_threshold=5
sc(dpm)
# +
# Read the new table that was created
catalog = QTable.read(source_catalog, format='ascii.ecsv')
dirim = fits.getdata(grism_direct_image, ext=1)
xs,ys=dirim.shape
fig = plt.figure(figsize=(4,4), dpi=150)
ax.ticklabel_format(useOffset=False)
ax = fig.add_subplot(1, 1, 1)
ax.ticklabel_format(useOffset=False)
ax.set_adjustable('box-forced')
ax.set_title(grism_direct_image.split("/")[-1], fontsize=8)
ax.imshow(dirim, origin='lower', extent=[0,xs,0,ys], vmin=-10, vmax=10)
# rectangle patches are xmin, ymin, width, height
plist1=[]
for obj in catalog:
plist1.append(patches.Circle((obj['xcentroid'].value, obj['ycentroid'].value),10))
for p in plist1:
ax.add_patch(p)
ax.imshow(dirim, origin='lower', extent=[0,xs,0,ys], vmin=-10, vmax=10)
# -
# ### Not bad! There were two nearby sources that were not deblended, but we'll leave that for now.
# checkout the final catalog
print(source_catalog)
catalog
|
general/Create-Source-Catalog.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Code for Module 3, Data Science Course
#
# Code is broken down by unit.
print(2+2)
# ## Unit 2
from pandas import Series
coins = Series([1,5,10,25],index=['penny','nickel','dime','quarter'])
coins
coins.min(), coins.max(), coins.sum()
mylist = ['penny','dime','loonie']
obj2 = Series(coins,index=mylist)
obj2
from pandas import DataFrame
coin_df = DataFrame([1,5,10,25],index=['penny', 'nickel', 'dime', 'quarter'])
coin_df
coin_df['weight']= [2.35, 3.95, 1.75, 4.4]
coin_df
print('The weight of a penny is', coin_df.loc['penny']['weight'],'grams.')
# ## Unit 3
# +
from pandas import DataFrame
data = {'name': ['penny', 'nickel', 'dime', 'quarter'],
'value': [1, 5, 10, 25],
'weight': [2.35, 3.95, 1.75, 4.4],
'design': ['Maple Leaves', 'Beaver', 'Schooner', 'Caribou'] }
dff=DataFrame( data )
# -
data = {'value': [1, 5, 10, 25],
'weight': [2.35, 3.95, 1.75, 4.4],
'design': ['Maple Leaves', 'Beaver', 'Schooner', 'Caribou'] }
coin_df=DataFrame( data ,index =['penny', 'nickel', 'dime', 'quarter'] )
coin_df
coin_df['design']=coin_df['design'].map(str.lower)
coin_df
dff = DataFrame(data,index=['a','b','c','d'])
dff.drop(index='a',inplace=True)
dff['weight'].map(lambda x: x+x)
DataFrame([[1,1,1],[2,4,8],[3,9,27],[4,16,64]])
DataFrame([[1,1,1],[2,4,8],[3,9,27],[4,16,64]],
columns =['base','square','cube'],
index=['one','two','three','four'])
# %%writefile Colours.csv
Colour,Emotion,Preference,Length
Yellow,Joy,2,3
Blue,Sadness,3,7
Red,Anger,6,5
Purple,Fear,5,4
Green,Disgust,4,7
Black,Grief,7,5
White,Ecstasy,1,7
# !cat Colours.csv
from pandas import read_csv
colours_df = read_csv('Colours.csv')
colours_df
from pandas import read_csv
colours_df = read_csv('https://tinyurl.com/CallyColours')
colours_df
read_csv('Colours.csv',nrows=5)
from pandas import read_table
read_table('Colours.csv',sep=',')
url = "https://tinyurl.com/y916axtz/pets.csv"
pets = read_csv(url)
pets
from pandas import DataFrame
import requests
import json
s1 = requests.get('https://www150.statcan.gc.ca/n1/dai-quo/ssi/homepage/ind-econ.json')
s1.json()
DataFrame(s1.json()['results']['geo'])
from pandas import read_html
url = 'https://en.wikipedia.org/wiki/Baseball_statistics'
df_list = read_html(url)
df_list[3]
table_list[3]
# ## Unit 4
#
pets.head()
pets.columns
pets['Age (years)']
pets.loc[3]
pets.loc[3,'Name']
pets.loc[3,'Name'] = '<NAME>'
pets['Age (years)'] = 12*pets['Age (years)']
pets.head()
pets.rename(columns = {'Age (years)':'Age (months)'},inplace = True)
pets.head()
print(
pets['Age (months)'].min(),
pets['Age (months)'].max(),
pets['Age (months)'].mean())
pets.plot(kind='bar',x='Name',y='Age (months)');
import plotly.express as px
fig = px.bar(pets, x='Name', y='Age (months)')
fig.show()
# ## Unit 5
from pandas import DataFrame
data = {'name': ['penny', 'nickel', 'dime', 'quarter'],
'value': [1, 5, 10, 25],
'weight': [2.35, 3.95, 1.75, 4.4],
'image': ['Leaf','Beaver','Blue Nose','Moose']}
coin_df = DataFrame(data)
coin_df[coin_df['value']>5]
coin_df.reindex([2,1,0,3])
coin_df.loc[5]= ['toonie',200,112.64,'Polar Bear']
coin_df
coin_df['weight'] = coin_df['weight']/28.35
coin_df
coin_df['image'].map(str.lower)
coin_df['weight'].map(lambda x: x*28.35)
coin_df['image'].replace('Beaver','Castor')
pets['Age (years)'].max()
pets['Weight (lbs)'].idxmax()
pets.loc[20]
# ## Unit 6
# ## Unit 7
from pandas import DataFrame
data = {'name': ['penny', 'nickel', 'dime', 'quarter'],
'value': [1, 5, 10, 25],
'weight': [2.35, 3.95, 1.75, 4.4],
'image': ['Leaf','Beaver','Blue Nose','Moose']}
coin_df = DataFrame(data)
coin_df
mint_df = DataFrame({'name': ['penny', 'nickel', 'dime', 'quarter'],
'minted (millions)': [199, 203, 292, 213]})
mint_df
mint_df = DataFrame({'name': ['quarter', 'dime', 'nickel', 'penny'],
'minted (millions)': [213, 292, 203, 199]})
mint_df
mint_df
from pandas import merge
merge(coin_df,mint_df)
merge(coin_df,mint_df, how='outer')
merge(coin_df,mint_df,on='name')
merge(coin_df,mint_df,on='weight',how='outer')
data_fr = {'name': ['sou', 'cinq sous', 'dix sous', 'vingt-cinq sous'],
'value': [1, 5, 10, 25],
'weight': [2.35, 3.95, 1.75, 4.4],
'image': ['Feuille','Castor','Voilier','Orignal']}
coin_fr = DataFrame(data_fr)
from pandas import concat
concat([coin_df,coin_fr])
result = concat([coin_df,coin_fr],keys=['English','French'])
result
result.sort_index(inplace=True)
result
result.loc['French']
result.loc[('English',0)]
pets.stack()
new_pets = pets.set_index(['Species','Name'])
new_pets
concat([pets[pets['Species']=='cat'],pets[pets['Species']=='dog']])
pets.sort_values('Species')
coin_df.T
from pandas import pivot
data = {'day':['Monday','Monday','Monday',
'Tuesday','Tuesday','Tuesday','Tuesday',
'Wednesday','Wednesday','Wednesday'],
'meal':['breakfast','lunch','dinner',
'breakfast','lunch','dinner','snack',
'breakfast','lunch','dinner'],
'calories':[400,500,1000,
400,600,1000,200,
300,700,1200]}
cal_count=DataFrame(data)
cal_count
cal_count.pivot('day','meal','calories')
pets.pivot('Name','Species')
# ## Unit 8
s1 = requests.get('https://www12.statcan.gc.ca/rest/census-recensement/CPR2016.json')
s1.text
s2 = s1.text[2:]
s3 = json.loads(s2)
s3
s3
df = pd.DataFrame(s3['DATA'],columns=s3['COLUMNS'])
df
df[0:20]['TEXT_NAME_NOM']
df.columns
df.head()
df.plot(kind='bar',x='num_children',y='num_pets',color='red')
df1 = DataFrame(df,
index =[9,10,11,13,14,15,16,17,18,19,20,21,22,24,25,26,27,29,30,31,32],
columns=['TEXT_NAME_NOM','T_DATA_DONNEE','F_DATA_DONNEE'])
df1.plot(kind='bar',x='TEXT_NAME_NOM',y='T_DATA_DONNEE');
df1.plot(kind='bar',x='TEXT_NAME_NOM',y='T_DATA_DONNEE');
import plotly.express as px
fig = px.bar(df1, x='TEXT_NAME_NOM', y='T_DATA_DONNEE')
fig.show()
df[100:200]['PROV_TERR_NAME_NOM']
df1 = DataFrame(df,
index =[9,10,11,13,14,15,16,17,18,19,20,21,22,24,25,26,27,29,30,31,32],
columns=['TEXT_NAME_NOM','T_DATA_DONNEE','M_DATA_DONNEE','F_DATA_DONNEE'])
fig = px.bar(df1, x='TEXT_NAME_NOM', y='M_DATA_DONNEE')
fig.show()
fig = px.bar(df1, x='TEXT_NAME_NOM', y='F_DATA_DONNEE')
fig.show()
# +
animals=['giraffes', 'orangutans', 'monkeys']
fig = go.Figure(data=[
go.Bar(name='SF Zoo', x=animals, y=[20, 14, 23]),
go.Bar(name='LA Zoo', x=animals, y=[12, 18, 29])
])
# Change the bar mode
fig.update_layout(barmode='group')
fig.show()
# +
url = "https://swift-yeg.cloud.cybera.ca:8080/v1/AUTH_233e84cd313945c992b4b585f7b9125d/callysto-open-data/pets.csv"
pets = pd.read_csv(url)
pets
# -
pets['Species']=='cat'
coins
frame.loc['image']
frame
frame.loc[1,'value']
frame[frame['value']>5]
frame['value']>5
frame
frame2=frame.reindex([2,1,0,3,5])
frame2.loc[5] = ['toonie',200,112.64,'Polar Bear']
frame2
frame['weight'] = frame['weight']/28.35
frame
import pandas as pd
url = "https://tinyurl.com/y9l6axtz/pets.csv"
pets = pd.read_csv(url)
pets
pets.head()
pets.columns
pets['Age (years)']
pets.loc[3]
pets.loc[3,'Name']
pets.loc[3,'Name'] = '<NAME>'
pets['Age (years)'] = 12*pets['Age (years)']
pets.head()
pets = pets.rename(columns = {'Age (years)':'Age (months)'})
pets.plot(kind='bar',x='Name',y='Age (years)');
import plotly.express as px
fig = px.bar(pets, x='Name', y='Age (years)')
fig.show()
print(
pets['Age (years)'].min(),
pets['Age (years)'].max(),
pets['Age (years)'].mean())
pets.head()
pets.plot(kind='scatter',x='Age (months)',y='Time to Adoption (weeks)');
s3 = json.loads(s1.text)
s3
s3['results']
len(s3['results']['indicators'])
df = pd.DataFrame(s3['results']['indicators'])
df
df['growth_rate'].head
df1
df2 = pd.DataFrame(df1.loc['results'])
df2
s1 = requests.get('https://www150.statcan.gc.ca/n1/dai-quo/ssi/homepage/ind-econ.json')
s1.json()
pd.DataFrame(s1.json()['results']['geo'])
data = {'name': ['penny', 'nickel', 'dime', 'quarter'],
'value': [1, 5, 10, 25],
'weight': [2.35, 3.95, 1.75, 4.4],
'image': ['Leaf','Beaver','Blue Nose','Moose']}
frame = pd.DataFrame(data)
frame
frame['weight'].map(lambda x: x/28.35)
frame
frame['image'].map(str.lower)
pets['Species'].replace('cat','feline')
pets['Legs'].replace(4,4.1)
from googletrans import Translator
# !pip install googletrans
translator = Translator()
translator.translate('bonjour').text
translator.translate('안녕하세요.').text
translator.translate('mains',src='fr',dest='en').text
pets['Name'].map(lambda x: translator.translate(x,dest='fr').text)
pets
translator.translate(pets['Species'][1],dest='fr').text
# !pip install pyspellchecker
from spellchecker import SpellChecker
from spellchecker import SpellChecker
spell = SpellChecker()
spell.correction("heey")
aa
pets['Name'].map(lambda x: spell.correction(x))
# !pip install gTTS
from gtts import gTTS
tts = gTTS( 'Hello, world' )
tts.save('hello.mp3')
# [](https://github.com/callysto/curriculum-notebooks/blob/master/LICENSE.md)
|
CallystoAndDataScience/DataScienceTests.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .ps1
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: .NET (PowerShell)
# language: PowerShell
# name: .net-powershell
# ---
# # T1016 - System Network Configuration Discovery
# Adversaries may look for details about the network configuration and settings of systems they access or through information discovery of remote systems. Several operating system administration utilities exist that can be used to gather this information. Examples include [Arp](https://attack.mitre.org/software/S0099), [ipconfig](https://attack.mitre.org/software/S0100)/[ifconfig](https://attack.mitre.org/software/S0101), [nbtstat](https://attack.mitre.org/software/S0102), and [route](https://attack.mitre.org/software/S0103).
#
# Adversaries may use the information from [System Network Configuration Discovery](https://attack.mitre.org/techniques/T1016) during automated discovery to shape follow-on behaviors, including whether or not the adversary fully infects the target and/or attempts specific actions.
# ## Atomic Tests
#Import the Module before running the tests.
# Checkout Jupyter Notebook at https://github.com/cyb3rbuff/TheAtomicPlaybook to run PS scripts.
Import-Module /Users/0x6c/AtomicRedTeam/atomics/invoke-atomicredteam/Invoke-AtomicRedTeam.psd1 - Force
# ### Atomic Test #1 - System Network Configuration Discovery on Windows
# Identify network configuration information
#
# Upon successful execution, cmd.exe will spawn multiple commands to list network configuration settings. Output will be via stdout.
#
# **Supported Platforms:** windows
# #### Attack Commands: Run with `command_prompt`
# ```command_prompt
# ipconfig /all
# netsh interface show interface
# arp -a
# nbtstat -n
# net config
# ```
Invoke-AtomicTest T1016 -TestNumbers 1
# ### Atomic Test #2 - List Windows Firewall Rules
# Enumerates Windows Firewall Rules using netsh.
#
# Upon successful execution, cmd.exe will spawn netsh.exe to list firewall rules. Output will be via stdout.
#
# **Supported Platforms:** windows
# #### Attack Commands: Run with `command_prompt`
# ```command_prompt
# netsh advfirewall firewall show rule name=all
# ```
Invoke-AtomicTest T1016 -TestNumbers 2
# ### Atomic Test #3 - System Network Configuration Discovery
# Identify network configuration information.
#
# Upon successful execution, sh will spawn multiple commands and output will be via stdout.
#
# **Supported Platforms:** macos, linux
# #### Attack Commands: Run with `sh`
# ```sh
# if [ -x "$(command -v arp)" ]; then arp -a; else echo "arp is missing from the machine. skipping..."; fi;
# if [ -x "$(command -v ifconfig)" ]; then ifconfig; else echo "ifconfig is missing from the machine. skipping..."; fi;
# if [ -x "$(command -v ip)" ]; then ip addr; else echo "ip is missing from the machine. skipping..."; fi;
# if [ -x "$(command -v netstat)" ]; then netstat -ant | awk '{print $NF}' | grep -v '[a-z]' | sort | uniq -c; else echo "netstat is missing from the machine. skipping..."; fi;
# ```
Invoke-AtomicTest T1016 -TestNumbers 3
# ### Atomic Test #4 - System Network Configuration Discovery (TrickBot Style)
# Identify network configuration information as seen by Trickbot and described here https://www.sneakymonkey.net/2019/10/29/trickbot-analysis-part-ii/
#
# Upon successful execution, cmd.exe will spawn `ipconfig /all`, `net config workstation`, `net view /all /domain`, `nltest /domain_trusts`. Output will be via stdout.
#
# **Supported Platforms:** windows
# #### Attack Commands: Run with `command_prompt`
# ```command_prompt
# ipconfig /all
# net config workstation
# net view /all /domain
# nltest /domain_trusts
# ```
Invoke-AtomicTest T1016 -TestNumbers 4
# ### Atomic Test #5 - List Open Egress Ports
# This is to test for what ports are open outbound. The technique used was taken from the following blog:
# https://www.blackhillsinfosec.com/poking-holes-in-the-firewall-egress-testing-with-allports-exposed/
#
# Upon successful execution, powershell will read top-128.txt (ports) and contact each port to confirm if open or not. Output will be to Desktop\open-ports.txt.
#
# **Supported Platforms:** windows
# #### Dependencies: Run with `powershell`!
# ##### Description: Test requires #{port_file} to exist
#
# ##### Check Prereq Commands:
# ```powershell
# if (Test-Path "PathToAtomicsFolder\T1016\src\top-128.txt") {exit 0} else {exit 1}
#
# ```
# ##### Get Prereq Commands:
# ```powershell
# New-Item -Type Directory (split-path PathToAtomicsFolder\T1016\src\top-128.txt) -ErrorAction ignore | Out-Null
# Invoke-WebRequest "https://github.com/redcanaryco/atomic-red-team/raw/master/atomics/T1016/src/top-128.txt" -OutFile "PathToAtomicsFolder\T1016\src\top-128.txt"
#
# ```
Invoke-AtomicTest T1016 -TestNumbers 5 -GetPreReqs
# #### Attack Commands: Run with `powershell`
# ```powershell
# $ports = Get-content PathToAtomicsFolder\T1016\src\top-128.txt
# $file = "$env:USERPROFILE\Desktop\open-ports.txt"
# $totalopen = 0
# $totalports = 0
# New-Item $file -Force
# foreach ($port in $ports) {
# $test = new-object system.Net.Sockets.TcpClient
# $wait = $test.beginConnect("allports.exposed", $port, $null, $null)
# $wait.asyncwaithandle.waitone(250, $false) | Out-Null
# $totalports++ | Out-Null
# if ($test.Connected) {
# $result = "$port open"
# Write-Host -ForegroundColor Green $result
# $result | Out-File -Encoding ASCII -append $file
# $totalopen++ | Out-Null
# }
# else {
# $result = "$port closed"
# Write-Host -ForegroundColor Red $result
# $totalclosed++ | Out-Null
# $result | Out-File -Encoding ASCII -append $file
# }
# }
# $results = "There were a total of $totalopen open ports out of $totalports ports tested."
# $results | Out-File -Encoding ASCII -append $file
# Write-Host $results
# ```
Invoke-AtomicTest T1016 -TestNumbers 5
# ## Detection
# System and network discovery techniques normally occur throughout an operation as an adversary learns the environment. Data and events should not be viewed in isolation, but as part of a chain of behavior that could lead to other activities, such as Lateral Movement, based on the information obtained.
#
# Monitor processes and command-line arguments for actions that could be taken to gather system and network information. Remote access tools with built-in features may interact directly with the Windows API to gather information. Information may also be acquired through Windows system management tools such as [Windows Management Instrumentation](https://attack.mitre.org/techniques/T1047) and [PowerShell](https://attack.mitre.org/techniques/T1059/001).
# ## Shield Active Defense
# ### Decoy Content
# Seed content that can be used to lead an adversary in a specific direction, entice a behavior, etc.
#
# Decoy Content is the data used to tell a story to an adversary. This content can be legitimate or synthetic data which is used to reinforce or validate your defensive strategy. Examples of decoy content are files on a storage object, entries in the system registry, system shortcuts, etc.
# #### Opportunity
# There is an opportunity to influence an adversary to move toward systems you want them to engage with.
# #### Use Case
# A defender can create breadcrumbs or honeytokens to lure the attackers toward the decoy systems or network services.
# #### Procedures
# Create directories and files with names and contents using key words that may be relevant to an adversary to see if they examine or exfiltrate the data.
# Seed a file system with content that is of no value to the company but reinforces the legitimacy of the system if viewed by an adversary.
|
playbook/tactics/discovery/T1016.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/bradonberner/PRML/blob/master/Copy_of_ch01_Introduction.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ceRmVgOq6XmY"
# # 1. Introduction
# + [markdown] id="Wj_RTULpriir"
# <table class="tfo-notebook-buttons" align="left">
#
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/pantelis/PRML/blob/master/notebooks/ch01_Introduction.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# </table>
# + id="TMxmVuag7eyA" colab={"base_uri": "https://localhost:8080/"} outputId="8b000f82-8338-4f4b-ce92-d7e40cc5180f"
from google.colab import drive
drive.mount('/content/drive')
# + id="F3rYRMAD6ynO" colab={"base_uri": "https://localhost:8080/"} outputId="737c5cd8-a369-4d1b-998b-cce1fe45f61c"
# You need to adjust the directory names below for your own account
# e.g. you may elect to create ms-notebooks dir or not
# Execute this cell once
# 1. Download the repo and set it as the current directory
# %cd /content/drive/My Drive/Colab-Notebooks
# !git clone https://github.com/bradonberner/PRML
# %cd /content/drive/My Drive/Colab-Notebooks/PRML
# 2. install the project/module
# !python setup.py install
# + id="iv9ADzLqiNsU" colab={"base_uri": "https://localhost:8080/"} outputId="3311b216-e056-404a-c745-2bee1a0fd0f1"
# 3. Add the project directory to the path
# %cd /content/drive/My Drive/Colab-Notebooks/PRML
import os, sys
sys.path.append(os.getcwd())
# + id="qwxjFZSR_vuX"
# Import seaborn
import seaborn as sns
# Apply the default theme
sns.set_theme()
# + id="pxBLdL3r6XmZ"
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from prml.preprocess import PolynomialFeature
from prml.linear import (
LinearRegression,
RidgeRegression,
BayesianRegression
)
np.random.seed(1234)
# + [markdown] id="TWjYKLNc6Xmd"
# ## 1.1. Example: Polynomial Curve Fitting
# + [markdown] id="6XPE-VC_-OYK"
# The cell below defines $p_{data}(y|x)$ and generates the $\hat p_{data}(y|x)$
# + id="Tj4RTV3X6Xmd" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="7448cce9-7bd5-4ee2-a65a-8c3c946580fa"
def create_toy_data(func, sample_size, std):
x = np.linspace(0, 1, sample_size) # p(x)
y = func(x) + np.random.normal(scale=std, size=x.shape)
return x, y
def func(x):
return np.sin(2 * np.pi * x)
x_train, y_train = create_toy_data(func, 10, 0.25)
x_test = np.linspace(0, 1, 100)
y_test = func(x_test)
plt.scatter(x_train, y_train, facecolor="none", edgecolor="b", s=50, label="training data")
plt.plot(x_test, y_test, c="g", label="$\sin(2\pi x)$")
plt.legend()
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.show()
# + id="diJlsNGo6Xmg" colab={"base_uri": "https://localhost:8080/", "height": 550} outputId="f8ba65be-073a-4fa7-c6c3-726c319b3e5b"
plt.subplots(figsize=(20, 10))
for i, degree in enumerate([0, 1, 3, 9]):
plt.subplot(2, 2, i + 1)
feature = PolynomialFeature(degree)
X_train = feature.transform(x_train)
X_test = feature.transform(x_test)
model = LinearRegression()
model.fit(X_train, y_train)
y = model.predict(X_test)
plt.scatter(x_train, y_train, facecolor="none", edgecolor="b", s=50, label="training data")
plt.plot(x_test, y_test, c="g", label="$\sin(2\pi x)$")
plt.plot(x_test, y, c="r", label="hypothesis")
plt.ylim(-1.5, 1.5)
plt.annotate("M={}".format(degree), xy=(-0.15, 1))
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.legend(bbox_to_anchor=(1.05, 0.64), loc=2, borderaxespad=0.)
plt.show()
# + id="nfpg434z6Xmj" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="aca98af4-8501-49b7-8167-7ba145eb82d5"
def rmse(a, b):
return np.sqrt(np.mean(np.square(a - b)))
training_errors = []
test_errors = []
for i in range(10):
feature = PolynomialFeature(i)
X_train = feature.transform(x_train)
X_test = feature.transform(x_test)
model = LinearRegression()
model.fit(X_train, y_train)
y = model.predict(X_test)
training_errors.append(rmse(model.predict(X_train), y_train))
test_errors.append(rmse(model.predict(X_test), y_test + np.random.normal(scale=0.25, size=len(y_test))))
plt.plot(training_errors, 'o-', mfc="none", mec="b", ms=10, c="b", label="Training")
plt.plot(test_errors, 'o-', mfc="none", mec="r", ms=10, c="r", label="Test")
plt.legend()
plt.xlabel("model capacity (degree)")
plt.ylabel("RMSE")
plt.show()
# + [markdown] id="bIa4pVbx8C8y"
# **Model Complexity:**
# Test error, that is the error between the model and the data points not used in making said model, is shown with the red line above. While the M=9 model is more complex then the M=3 model, it has a higher RMSE then M=3. This shows that simpler models can be more accurate than more complex ones.
# + [markdown] id="qZQUK1k_40Nv"
# **The Loss Function:**
# The loss function is called the root mean squared error because it takes the Error from all data points, Sqaures all of them, takes their Mean, than finally sqaure Roots this value. It uses the square root to negate the square performed earlier. It squared the data earlier to convert all negative errors to positive ones. This number helps explain how well the model fits the data.
# + id="Djzk4019_Pfj" colab={"base_uri": "https://localhost:8080/"} outputId="8e9a123d-6e0a-4c18-d4e9-1ab34dd1cfc8"
for i, degree in enumerate([0, 1, 3, 9]):
feature = PolynomialFeature(degree)
X_train = feature.transform(x_train)
X_test = feature.transform(x_test)
model = LinearRegression()
model.fit(X_train, y_train)
print("m",degree,"=",end=" ")
for num in model.w:
print(num,end=" ")
print()
# + [markdown] id="Ui1QFMFVXmBL"
# **Parameter Table:**
# As m and the number of w terms increase, their coefficents appear to exponentially increase in magnitude. For example, w1's coefficent in m3, 9.29, is much smaller in absolute value to w1's coefficent in m9, 32.23.
# + [markdown] id="PTOLlihm6Xml"
# #### Regularization
# + id="18aoGaUg6Xml" colab={"base_uri": "https://localhost:8080/", "height": 289} outputId="a061fa26-12bb-4daa-d2a0-fc5fc43ff5da"
feature = PolynomialFeature(9)
X_train = feature.transform(x_train)
X_test = feature.transform(x_test)
model = RidgeRegression(alpha=1e-3)
model.fit(X_train, y_train)
y = model.predict(X_test)
#y = model.predict(X_test)
plt.scatter(x_train, y_train, facecolor="none", edgecolor="b", s=50, label="training data")
plt.plot(x_test, y_test, c="g", label="$\sin(2\pi x)$")
plt.plot(x_test, y, c="r", label="hypothesis")
plt.ylim(-1.5, 1.5)
plt.legend()
plt.annotate("M=9", xy=(-0.15, 1))
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.show()
# + [markdown] id="xeg1JevvV89f"
# **Regularization:**
# Regularization models the data like regression, except that it penalizes wild osccilations that result from more complex models. It does this by factoring in the norm of w, along with the hyperparameter lamda.
# + id="FxScTn80NcUt" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="06b2cb9e-b5e1-4ae6-e93a-a02c56d09064"
def rmse(a, b):
return np.sqrt(np.mean(np.square(a - b)))
training_errors = []
test_errors = []
for i in range(10):
feature = PolynomialFeature(i)
X_train = feature.transform(x_train)
X_test = feature.transform(x_test)
model = RidgeRegression(alpha=1e-3)
model.fit(X_train, y_train)
y = model.predict(X_test)
training_errors.append(rmse(model.predict(X_train), y_train))
test_errors.append(rmse(model.predict(X_test), y_test + np.random.normal(scale=0.25, size=len(y_test))))
plt.plot(training_errors, 'o-', mfc="none", mec="b", ms=10, c="b", label="Training")
plt.plot(test_errors, 'o-', mfc="none", mec="r", ms=10, c="r", label="Test")
plt.legend()
plt.xlabel("model capacity (degree)")
plt.ylabel("RMSE")
plt.show()
# + [markdown] id="kfmJy1-96Xmo"
# ### 1.2.6 Bayesian curve fitting
# + id="GFCXxwiz6Xmo" colab={"base_uri": "https://localhost:8080/", "height": 289} outputId="78ca34ae-1564-4617-b5f1-85ea81138d46"
model = BayesianRegression(alpha=2e-3, beta=2)
model.fit(X_train, y_train)
y, y_err = model.predict(X_test, return_std=True)
plt.scatter(x_train, y_train, facecolor="none", edgecolor="b", s=50, label="training data")
plt.plot(x_test, y_test, c="g", label="$\sin(2\pi x)$")
plt.plot(x_test, y, c="r", label="mean")
plt.fill_between(x_test, y - y_err, y + y_err, color="pink", label="std.", alpha=0.5)
plt.xlim(-0.1, 1.1)
plt.ylim(-1.5, 1.5)
plt.annotate("M=9", xy=(0.8, 1))
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.legend(bbox_to_anchor=(1.05, 1.), loc=2, borderaxespad=0.)
plt.show()
# + id="i4VIskNS6Xmt"
|
Copy_of_ch01_Introduction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ENTRADA Y SALIDA DE DATOS
# ## 1. Entrada de Información por teclado (Input)
# En python la funcion input() nos ayudará con la tarea de capturar datos del usuario
# input por defecto nos devuelve datos de tipo texto
# SHIFT + TAB -> Documentacion de la palabra a ejecutar
decimal = input("Introduce un número decimal con punto: ")
type(decimal)
numero = float(decimal)
# tab autocompleta texto
numero
# ## 2. Entrada de datos por Argumentos
# Para poder enviar información a un script y manejarla, tenemos que utilizar la librería de sistema [sys](https://www.geeksforgeeks.org/how-to-use-sys-argv-in-python/). En ella encontraremos la lista argv que almacena los argumentos enviados al script.
import sys
print(sys.argv)
# ## 3. Salida de información (Output)
# La función print nos ayudará a realizar salidas de información
print('hola')
# # EJERCICIOS
# #### 1. Realiza un programa que lea 2 números por teclado y determine los siguientes aspectos (es suficiene con mostrar True o False):
#
# - Si los dos números son iguales
# - Si los dos números son diferentes
# - Si el primero es mayor que el segundo
# - Si el segundo es mayor o igual que el primero
a = float(input("Primer número: " ))
b = float(input("Segundo número: " ))
print ("Los dos números son iguales: ", a == b)
print ("Los dos números son diferentes: ", a != b)
print ("El primero es mayor que el segundo: ", a > b)
print ("El segundo es mayor o igual que el primero: ", a <= b)
# ### 2. Escribir un programa que pregunte el nombre del usuario en la consola y después de que el usuario lo introduzca muestre por pantalla la cadena ¡Hola "nombre"!, donde "nombre" es el nombre que el usuario haya introducido.
#
#
#
nombre = input("Introduzca nombre: ")
print("Hola" , " " , nombre , " ! ")
|
MODULO 1/4. Entrada y Salida de Datos.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from fastai2.vision.all import *
path = untar_data(URLs.PETS)
path.ls()
pets = DataBlock(blocks=(ImageBlock,CategoryBlock),get_items=get_image_files,
splitter=RandomSplitter(seed=42),
get_y=using_attr(RegexLabeller(r'(.+)_\d+.jpg$'),'name'),
item_tfms=Resize(460),
batch_tfms=aug_transforms(size=224,min_scale=0.75))
dls = pets.dataloaders(path/"images")
dls.show_batch(nrows=1,ncols=3)
learn = cnn_learner(dls,resnet34,metrics=error_rate)
learn.fine_tune(2)
#look at activations
x,y = dls.one_batch()
y
#get preds
preds,_ = learn.get_preds(dl=[(x,y)])
preds[1]
print(preds[1].shape) #37 preds
preds[1].sum() #sum to 1
def softmax(x):
return torch.exp(x)/torch.exp(x).sum(dim=1,keepdim=True)
acts = torch.randn((3,2))
acts = softmax(acts)
acts
def log_likeli(inputs,targets):
inputs = inputs.sigmoid()
return torch.where(targets==1,1-inputs,inputs).mean()
targ = torch.tensor([1,0,1])
idx = range(3)
idx
acts[idx,targ]
interp = ClassificationInterpretation.from_learner(learn)
interp.most_confused(min_val=5)
learn = cnn_learner(dls,resnet34,metrics=error_rate)
lr_min, lr_steep = learn.lr_find()
learn = cnn_learner(dls,resnet34,metrics=error_rate)
learn.fine_tune(2,base_lr =3e-3)
learn = cnn_learner(dls,resnet34,metrics=error_rate)
learn.fit_one_cycle(3,3e-3)
learn.unfreeze()
learn.lr_find()
# fit_one_cycle is used for transfer learning when fine_tune is not used, it does allow more manual mods, fine_tune does :
# 1) trains added layers for one epochs, with others frozen
# 2) unfreeze all layers, trains them for # of epochs requested
learn.fit_one_cycle(6,lr_max=5e-5)
learn = cnn_learner(dls,resnet34,metrics=error_rate)
learn.fit_one_cycle(3,3e-3)
learn.unfreeze()
learn.fit_one_cycle(12,lr_max=slice(1e-6,1e-4))
learn.recorder.plot_loss()
|
book/Chapter5/Chapter5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
from airsenal.framework.utils import *
from airsenal.framework.bpl_interface import get_fitted_team_model
from airsenal.framework.season import get_current_season, CURRENT_TEAMS
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import jax.numpy as jnp
# %matplotlib inline
# -
model_team = get_fitted_team_model(get_current_season(), NEXT_GAMEWEEK, session)
# +
# extract indices of current premier league teams
# val-1 because 1-indexed in model but 0-indexed in python
current_idx = {team: idx for idx, team in enumerate(model_team.teams)
if team in CURRENT_TEAMS}
top6 = ['MCI', 'LIV', 'TOT', 'CHE', 'MUN', 'ARS']
# +
ax = plt.figure(figsize=(15, 5)).gca()
for team, idx in current_idx.items():
sns.kdeplot(model_team.attack[:, idx], label=team)
plt.title('attack')
plt.legend()
ax = plt.figure(figsize=(15, 5)).gca()
for team, idx in current_idx.items():
sns.kdeplot(model_team.defence[:, idx], label=team)
plt.title('defence')
plt.legend()
ax = plt.figure(figsize=(15, 5)).gca()
for team, idx in current_idx.items():
sns.kdeplot(model_team.home_advantage[:, idx], label=team)
plt.title('home advantage')
plt.legend()
# +
a_mean = model_team.attack.mean(axis=0)
b_mean = model_team.defence.mean(axis=0)
a_conf95 = np.abs(np.quantile(model_team.attack,[0.025, 0.975], axis=0) - a_mean)
b_conf95 = np.abs(np.quantile(model_team.defence, [0.025, 0.975], axis=0) - b_mean)
a_conf80 = np.abs(np.quantile(model_team.attack,[0.1, 0.9], axis=0) - a_mean)
b_conf80 = np.abs(np.quantile(model_team.defence, [0.1, 0.9], axis=0) - b_mean)
fig, ax = plt.subplots(1, 1, figsize=(10,10))
ax.set_aspect('equal')
select_idx = jnp.array(list(current_idx.values()), dtype=int)
plt.errorbar(a_mean[select_idx],
b_mean[select_idx],
xerr=a_conf80[:, select_idx],
yerr=b_conf80[:, select_idx],
marker='o', markersize=10,
linestyle='', linewidth=0.5)
plt.xlabel('attack', fontsize=14)
plt.ylabel('defence', fontsize=14)
for team, idx in current_idx.items():
ax.annotate(team,
(a_mean[idx]-0.03, b_mean[idx]+0.02),
fontsize=12)
# +
plt.figure()
for i in range(model_team.attack_coefficients.shape[1]):
sns.kdeplot(model_team.attack_coefficients[:, i])
plt.title("Attack Coefficients (Covariates)")
plt.figure()
for i in range(model_team.defence_coefficients.shape[1]):
sns.kdeplot(model_team.defence_coefficients[:, i])
plt.title("Defence Coefficients (Covariates)")
# +
beta_a_mean = model_team.attack_coefficients.mean(axis=0)
beta_b_mean = model_team.defence_coefficients.mean(axis=0)
beta_a_conf95 = np.abs(np.quantile(model_team.attack_coefficients,[0.025, 0.975], axis=0) - beta_a_mean)
beta_b_conf95 = np.abs(np.quantile(model_team.defence_coefficients, [0.025, 0.975], axis=0) - beta_b_mean)
beta_a_conf80 = np.abs(np.quantile(model_team.attack_coefficients,[0.1, 0.9], axis=0) - beta_a_mean)
beta_b_conf80 = np.abs(np.quantile(model_team.defence_coefficients, [0.1, 0.9], axis=0) - beta_b_mean)
fig, ax = plt.subplots(1, 1, figsize=(7,7))
ax.set_aspect('equal')
plt.errorbar(beta_a_mean,
beta_b_mean,
xerr=beta_a_conf80,
yerr=beta_b_conf80,
marker='o', markersize=10,
linestyle='', linewidth=0.5)
plt.xlabel('beta_a', fontsize=14)
plt.ylabel('beta_b', fontsize=14)
plt.title('FIFA Ratings')
for idx, feat in enumerate(["att", "mid", "defn", "ovr"]):
ax.annotate(feat,
(beta_a_mean[idx]-0.03, beta_b_mean[idx]+0.02),
fontsize=12)
xlim = ax.get_xlim()
ylim = ax.get_ylim()
plt.plot([0, 0], ylim, color='k', linewidth=0.75)
plt.plot(xlim, [0, 0], color='k', linewidth=0.75)
plt.xlim(xlim)
plt.ylim(ylim)
# -
sns.kdeplot(model_team.rho)
plt.title("rho")
team_h = "MCI"
team_a = "MUN"
model_team.predict_concede_n_proba(2, team_h, team_a)
model_team.predict_score_n_proba(2, team_h, team_a)
model_team.predict_outcome_proba(team_h, team_a)
model_team.predict_score_proba(team_h, team_a, 2, 2)
# +
max_goals = 10
prob_score_h = [model_team.predict_score_n_proba(n, team_h, team_a)[0] for n in range(max_goals)]
print(team_h, "exp goals", sum([n*prob_score_h[n] for n in range(max_goals)])/sum(prob_score_h))
prob_score_a = [model_team.predict_score_n_proba(n, team_a, team_h, home=False)[0] for n in range(max_goals)]
print(team_a, "exp goals", sum([n*prob_score_a[n] for n in range(max_goals)])/sum(prob_score_a))
max_prob = 1.1*max(prob_score_h + prob_score_a)
plt.figure(figsize=(15,5))
plt.subplot(1,2,1)
plt.bar(range(max_goals), prob_score_h)
plt.ylim([0, max_prob])
plt.xlim([-1, max_goals])
plt.title(team_h)
plt.subplot(1,2,2)
plt.bar(range(max_goals), prob_score_a)
plt.ylim([0, max_prob])
plt.xlim([-1, max_goals])
plt.title(team_a);
# -
|
notebooks/team_model_bpl_next.ipynb
|
;; -*- coding: utf-8 -*-
;; ---
;; jupyter:
;; jupytext:
;; text_representation:
;; extension: .scm
;; format_name: light
;; format_version: '1.5'
;; jupytext_version: 1.14.4
;; kernelspec:
;; display_name: Calysto Scheme 3
;; language: scheme
;; name: calysto_scheme
;; ---
;; ### 2.3.2 例:記号微分
;;
;; ここでは導関数を返す手続きを考えます。
;; 引数として、式
;; $ax^2 + bx + c$
;; と変数名$x$を与えると、
;; $2ax + b$
;; を返すというものです。
(symbol? "test")
(define x "test")
(symbol? x)
(symbol? 'x)
(symbol? 'y)
;; +
; eは変数か?
(define (variable? x)
(symbol? x))
; v1とv2は同じ変数か?
(define (same-variable? v1 v2)
(and (variable? v1) (variable? v2) (eq? v1 v2)))
; eは和か?
(define (sum? x)
(and (pair? x) (eq? (car x) '+)))
; 和eの加数
(define (addend s)
(cadr s))
; 和eの被加数
(define (augend s)
(caddr s))
; a1とa2の和を構築する
(define (make-sum a1 a2)
(list '+ a1 a2))
;eは積か?
(define (product? x)
(and (pair? x) (eq? (car x) '*)))
; 積eの乗数
(define (multiplier p)
(cadr p))
;積eの被乗数
(define (multiplicand p)
(caddr p))
; m1とm2の積を構築する
(define (make-product m1 m2)
(list '* m1 m2))
;; -
(define (deriv exp var)
(cond ((number? exp) 0)
((variable? exp) (if (same-variable? exp var) 1 0))
((sum? exp) (make-sum (deriv (addend exp) var)
(deriv (augend exp) var)))
((product? exp) (make-sum (make-product (multiplier exp)
(deriv (multiplicand exp) var))
(make-product (deriv (multiplier exp) var) (multiplicand exp))))
(else (error "unknown expression type: DERIV" exp)))
)
(deriv '(+ x 3) 'x)
(deriv '(* x y) 'x)
(deriv '(* (* x y) (+ x 3)) 'x)
(define (make-sum a1 a2)
(cond ((=number? a1 0) a2)
((=number? a2 0) a1)
((and (number? a1) (number? a2)) (+ a1 a2))
(else (list '+ a1 a2)))
)
(define (=number? exp num)
(and (number? exp) (= exp num)))
(define (make-product m1 m2)
(cond ((or (=number? m1 0) (=number? m2 0)) 0)
((=number? m1 1) m2) ((=number? m2 1) m1)
((and (number? m1) (number? m2)) (* m1 m2))
(else (list '* m1 m2)))
)
(deriv '(+ x 3) 'x)
(deriv '(* x y) 'x)
(deriv '(* (* x y) (+ x 3)) 'x)
(number? 9)
(number? x)
(define (calc exp)
(cond ((number? exp) exp)
((variable? exp) exp)
((sum? exp) (make-sum (calc (addend exp))
(calc (augend exp))))
((product? exp) (make-product (calc (multiplier exp))
(calc (multiplicand exp))))
(else (error "unknown expression type: calc" exp)))
)
'(+ (+ (+ -2 3) 4) 8)
(calc '(+ (+ (+ -2 3) 4) 8))
(calc '(* (* 2 -3) 5))
(calc '(+ (* (+ 2 3) 4) 8))
(calc '(* (+ (* (+ 2 -3) 4) 8) a))
(define a 10)
(eval (calc '(* (+ (* (+ 2 -3) 4) 8) a)))
(* (+ (* (+ 2 -3) 4) 8) 4)
;; +
#define test aaa ## bbb
void test() -> aaabbb()
{
}
;; -
;; #### 練習問題
;;
;; - [練習問題2.56 べき乗の微分](../exercises/2.56.ipynb)
;; - [練習問題2.57 3以上の項数の和と積の微分](../exercises/2.57.ipynb)
;; - [練習問題2.58 中間記法の式](../exercises/2.58.ipynb)
|
texts/2.3.2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Topics
#
# - lists
# - indexing
# - slicing
# - mutable property of lists
# - list functions
# - list methods
# - tuples
# - immutable property of tuples
# - why use list over tuple, or tuple over list?
days = ['S', 'M', 'T', 'W', 'Th', 'F', 'Sa']
days
mix_bag = [20, 'weekend', 'monkey', 7.8]
mix_bag = [20, 'weekend', 'monkey', 7.8, [21, 'weekend', 'monkey', 7.8]]
mix_bag
days[0]
days[-1]
days[1:-1]
# create a list of month
# what is the slice to get feb to july
month = [1 , 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12]
index = [0 , 1 , 2, 3, 4, 5, 6, 7, 8, 9, 10, 11]
x = 1
y = 7
month_feb_to_juyl = month[x:y]
month_feb_to_juyl
len(month) - 5
month[0] = 'Rasbarry'
month
len(month)
days = ['S', 'M', 'T', 'W', 'Th', 'F', 'Sa']
sorted(days)
my_list = [1, 2, 3, 4 ,5]
sum(my_list)
max(my_list)
my_list.append(6)
my_list
my_list.pop()
7 in my_list
4 in my_list
# +
## Tuples
# -
# ### Exercise
# +
# swap the value of a and b
a = 10
b = 20
# fill in the blanks
temp = a
a = b
b = temp
print (a)
print (b)
# -
a , b = b, a
a, b = 5, 10
# a = 5
# b = 10
# a, b = 5, 10
KISS philosophy
# write a function that receive a number as its only argument (no default arg required),
# if it can be divided by 3: print '3 le huncha'
# if it can be divided by 5: print '5 le huncha'
# if it can be divided by 2: print 'even'
# otherwise print: 'nope'
def my_func(a):
final = ''
if (a%3 == 0):
final = final + "3 le huncha"
if (a%5 == 0):
final = final + "5 le huncha"
if (a%2==0):
final = final + "even"
if (final==''):
final = 'nope'
return final
my_func(10)
# ## Fizz Buzz Problem:
# Print integers 1 to 50, but print “Fizz” if an integer is divisible by 3, “Buzz” if an integer is divisible by 5, and “FizzBuzz” if an integer is divisible by both 3 and 5.
# +
# 1
# 2
# Fizz
# 4
# Buzz
# Fizz
# 7
# 8
# Fizz
# Buzz
# 11
# Fizz
# 13
# 14
# FizzBuzz
# -
for i in range(11):
print (i)
for i in range (1, 100):
final = ''
if i % 3 == 0:
final = final + 'Fizz'
if i % 5 == 0:
final = final + 'Buzz'
if final == '':
final = i
print (final)
|
python/4_lists_tuples.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Goals
#
#
#
# ### Train a blood cell type classifier using resnet variants
#
# ### Understand what all differences happen when switching between resnets variants
#
# ### Understand bigger and deeper network not always means better results
#
# #### For this experiment you will be using mxnet backend
# # What is resnet
#
# ## Readings on resnet
#
# 1) Points from https://towardsdatascience.com/an-overview-of-resnet-and-its-variants-5281e2f56035
# - The core idea of ResNet is introducing a so-called “identity shortcut connection” that skips one or more layers
# - The deeper model should not produce a training error higher than its shallower counterparts.
# - solves the problem of vanishing gradiens as network depth increased - https://medium.com/@anishsingh20/the-vanishing-gradient-problem-48ae7f501257
#
#
#
# 2) Points from https://medium.com/@14prakash/understanding-and-implementing-architectures-of-resnet-and-resnext-for-state-of-the-art-image-cf51669e1624
# - Won 1st place in the ILSVRC 2015 classification competition with top-5 error rate of 3.57% (An ensemble model)
# - Efficiently trained networks with 100 layers and 1000 layers also.
# - Replacing VGG-16 layers in Faster R-CNN with ResNet-101. They observed a relative improvements of 28%
#
#
# 3) Read more here
# - https://arxiv.org/abs/1512.03385
# - https://d2l.ai/chapter_convolutional-modern/resnet.html
# - https://cv-tricks.com/keras/understand-implement-resnets/
# - https://mc.ai/resnet-architecture-explained/
#
#
#
# # Table of Contents
#
#
# ## [0. Install](#0)
#
#
# ## [1. Train experiment with resnet-18 architecture and validate](#1)
#
#
# ## [2. Train experiment with resnet-32 architecture and validate](#2)
#
#
# ## [3. Train experiment with resnet-50 architecture and validate](#3)
#
#
# ## [4. Train experiment with resnet-101 architecture and validate](#4)
#
#
# ## [5. Train experiment with resnet-152 architecture and validate](#5)
#
#
# ## [6. Compare all the 4 experiments](#11)
# <a id='0'></a>
# # Install Monk
#
# - git clone https://github.com/Tessellate-Imaging/monk_v1.git
#
# - cd monk_v1/installation/Linux && pip install -r requirements_cu9.txt
# - (Select the requirements file as per OS and CUDA version)
# !git clone https://github.com/Tessellate-Imaging/monk_v1.git
# Select the requirements file as per OS and CUDA version
# !cd monk_v1/installation/Linux && pip install -r requirements_cu9.txt
# ## Dataset - Blood Cell Type Classification
# - https://www.kaggle.com/paultimothymooney/blood-cells
# ! wget --load-cookies /tmp/cookies.txt "https://docs.google.com/uc?export=download&confirm=$(wget --save-cookies /tmp/cookies.txt --keep-session-cookies --no-check-certificate 'https://docs.google.com/uc?export=download&id=1KhXKL58mnXL1G1uRDsCmCXMk7ubwXjps' -O- | sed -rn 's/.*confirm=([0-9A-Za-z_]+).*/\1\n/p')&id=1KhXKL58mnXL1G1uRDsCmCXMk7ubwXjps" -O blood-cells.zip && rm -rf /tmp/cookies.txt
# ! unzip -qq blood-cells.zip
# # Imports
# Monk
import os
import sys
sys.path.append("monk_v1/monk/");
#Using mxnet-gluon backend
from gluon_prototype import prototype
# <a id='1'></a>
# # Train experiment with resnet-18 architecture and validate
# +
# Load experiment
gtf = prototype(verbose=1);
gtf.Prototype("Compare-resnet-v1-depth", "resnet18-v1");
# Insert data and set params in default mode
gtf.Default(dataset_path="blood-cells/train",
model_name="resnet18_v1",
freeze_base_network=False,
num_epochs=5);
# +
#Start Training
gtf.Train();
#Read the training summary generated once you run the cell and training is completed
# -
# +
# Load for validation
gtf = prototype(verbose=1);
gtf.Prototype("Compare-resnet-v1-depth", "resnet18-v1", eval_infer=True);
# Set dataset
gtf.Dataset_Params(dataset_path="blood-cells/val");
gtf.Dataset();
# Validate
accuracy, class_based_accuracy = gtf.Evaluate();
# -
# <a id='2'></a>
# # Train experiment with resnet-34 architecture and validate
# +
# Load experiment
gtf = prototype(verbose=1);
gtf.Prototype("Compare-resnet-v1-depth", "resnet34-v1");
# Insert data and set params in default mode
gtf.Default(dataset_path="blood-cells/train",
model_name="resnet34_v1",
freeze_base_network=False,
num_epochs=5);
# -
# +
#Start Training
gtf.Train();
#Read the training summary generated once you run the cell and training is completed
# -
# +
# Load for validation
gtf = prototype(verbose=1);
gtf.Prototype("Compare-resnet-v1-depth", "resnet34-v1", eval_infer=True);
# Set dataset
gtf.Dataset_Params(dataset_path="blood-cells/val");
gtf.Dataset();
# Validate
accuracy, class_based_accuracy = gtf.Evaluate();
# -
# <a id='3'></a>
# # Train experiment with resnet-50 architecture and validate
# +
# Load experiment
gtf = prototype(verbose=1);
gtf.Prototype("Compare-resnet-v1-depth", "resnet50-v1");
# Insert data and set params in default mode
gtf.Default(dataset_path="blood-cells/train",
model_name="resnet50_v1",
freeze_base_network=False,
num_epochs=5);
# -
# +
#Start Training
gtf.Train();
#Read the training summary generated once you run the cell and training is completed
# -
# +
# Load for validation
gtf = prototype(verbose=1);
gtf.Prototype("Compare-resnet-v1-depth", "resnet50-v1", eval_infer=True);
# Set dataset
gtf.Dataset_Params(dataset_path="blood-cells/val");
gtf.Dataset();
# Validate
accuracy, class_based_accuracy = gtf.Evaluate();
# -
# <a id='4'></a>
# # Train experiment with resnet-101 architecture and validate
# +
# Load experiment
gtf = prototype(verbose=1);
gtf.Prototype("Compare-resnet-v1-depth", "resnet101-v1");
# Insert data and set params in default mode
gtf.Default(dataset_path="blood-cells/train",
model_name="resnet101_v1",
freeze_base_network=False,
num_epochs=5);
# -
# +
#Start Training
gtf.Train();
#Read the training summary generated once you run the cell and training is completed
# -
# +
# Load for validation
gtf = prototype(verbose=1);
gtf.Prototype("Compare-resnet-v1-depth", "resnet101-v1", eval_infer=True);
# Set dataset
gtf.Dataset_Params(dataset_path="blood-cells/val");
gtf.Dataset();
# Validate
accuracy, class_based_accuracy = gtf.Evaluate();
# -
# <a id='5'></a>
# # Train experiment with resnet-152 architecture and validate
# +
# Load experiment
gtf = prototype(verbose=1);
gtf.Prototype("Compare-resnet-v1-depth", "resnet152-v1");
# Insert data and set params in default mode
gtf.Default(dataset_path="blood-cells/train",
model_name="resnet152_v1",
freeze_base_network=False,
num_epochs=5);
# -
# +
#Start Training
gtf.Train();
#Read the training summary generated once you run the cell and training is completed
# -
# +
# Load for validation
gtf = prototype(verbose=1);
gtf.Prototype("Compare-resnet-v1-depth", "resnet152-v1", eval_infer=True);
# Set dataset
gtf.Dataset_Params(dataset_path="blood-cells/val");
gtf.Dataset();
# Validate
accuracy, class_based_accuracy = gtf.Evaluate();
# -
# <a id='11'></a>
# # Comparing all the experiments
# Invoke the comparison class
from compare_prototype import compare
# ### Creating and managing comparison experiments
# - Provide project name
# Create a project
gtf = compare(verbose=1);
gtf.Comparison("Compare-effect-of-network-depth");
# ### This creates files and directories as per the following structure
#
# workspace
# |
# |--------comparison
# |
# |
# |-----Compare-effect-of-network-depth
# |
# |------stats_best_val_acc.png
# |------stats_max_gpu_usage.png
# |------stats_training_time.png
# |------train_accuracy.png
# |------train_loss.png
# |------val_accuracy.png
# |------val_loss.png
#
# |
# |-----comparison.csv (Contains necessary details of all experiments)
# ### Add the experiments
# - First argument - Project name
# - Second argument - Experiment name
gtf.Add_Experiment("Compare-resnet-v1-depth", "resnet18-v1");
gtf.Add_Experiment("Compare-resnet-v1-depth", "resnet34-v1");
gtf.Add_Experiment("Compare-resnet-v1-depth", "resnet50-v1");
gtf.Add_Experiment("Compare-resnet-v1-depth", "resnet101-v1");
gtf.Add_Experiment("Compare-resnet-v1-depth", "resnet152-v1");
# ## Run Analysis
gtf.Generate_Statistics();
# ## Visualize and study comparison metrics
# ### Training Accuracy Curves
from IPython.display import Image
Image(filename="workspace/comparison/Compare-effect-of-network-depth/train_accuracy.png")
# ### Training Loss Curves
from IPython.display import Image
Image(filename="workspace/comparison/Compare-effect-of-network-depth/train_loss.png")
# ### Validation Accuracy Curves
from IPython.display import Image
Image(filename="workspace/comparison/Compare-effect-of-network-depth/val_accuracy.png")
# ### Validation loss curves
from IPython.display import Image
Image(filename="workspace/comparison/Compare-effect-of-network-depth/val_loss.png")
# ### Training Times and max gpu usages
from IPython.display import Image
Image(filename="workspace/comparison/Compare-effect-of-network-depth/stats_training_time.png")
from IPython.display import Image
Image(filename="workspace/comparison/Compare-effect-of-network-depth/stats_max_gpu_usage.png")
# # Comparisons
# #### You may get differet results
#
# Network | Val Acc | Training time (sec) | Gpu Usage (mb)
#
#
# resnet-18 | 88.1 | 220 | 1250
#
#
# resnet-34 | 87.4 | 310 | 1600
#
#
#
# resnet-50 | 88.4 | 580 | 2250
#
#
#
# resnet-101 | 86.2 | 840 | 2600
#
#
#
# resnet-152 | 85.8 | 1120 | 3700
#
#
#
|
study_roadmaps/4_image_classification_zoo/Blood Cell Type Classification_ Understanding the impact of depth in network.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Prelude
# %load_ext autoreload
# %autoreload 2
# +
# First, I have to laod different modules that I use for analyzing the data and for plotting:
import sys, os, collections
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt; plt.rcdefaults()
from matplotlib.pyplot import figure
from collections import Counter
# Second, I have to load the Text Fabric app
from tf.fabric import Fabric
from tf.app import use
# -
A = use('bhsa', hoist=globals())
# # Jer 1 queries
# ## The word happening
#
# ### Task #1:
#
# Try to investiage the formulation "The word happening" and do a statistical analysis of its appearance. How could you best understand the construction? Do a statistical survey by exporting your results and loading them back as a pandas dataframe (see the first TF notebook of this class to get instructions). Create some charts that help you get a better graphical representations of the phenomenons distribution.
WordHappens1='''
clause typ=WayX
phrase function=Pred
word lex=HJH[
phrase function=Subj
word lex=DBR/ st=c
<: word lex*
'''
WordHappens1 = A.search(WordHappens1)
A.table(WordHappens1, start=1, end=2, condensed=False, colorMap={3:'pink', 4:'lime'})
A.export(WordHappens1, toDir='D:/OneDrive/1200_AUS-research/Fabric-TEXT', toFile='WordHappens1.tsv')
WordHappens1=pd.read_csv('D:/OneDrive/1200_AUS-research/Fabric-TEXT/WordHappens1.tsv',delimiter='\t',encoding='utf-16')
pd.set_option('display.max_columns', 50)
WordHappens1.head()
WordHappens1["lex6"].value_counts()
# +
fig, ax = plt.subplots()
fig.set_size_inches(15, 10)
for S1, df in WordHappens1.groupby('lex6'):
ax.scatter(x="S1", y="lex6", data=df, label=S1)
ax.set_xlabel("predicates used by YHWH as subject")
ax.set_ylabel("book")
ax.legend();
# -
sns.lmplot(x="S1", y="R", data=WordHappens1, hue='lex6', height=20, aspect=2/3, fit_reg=False, scatter_kws={"s": 200})
ax = plt.gca()
ax.set_ylabel('Number of occurence of ImpChainType')
ax.set_xlabel('OT books')
WordHappens2='''
clause typ=WayX
phrase function=Pred
word lex=HJH[
phrase function=Subj
word lex=DBR/ st=c
<: word sp#nmpr
'''
WordHappens2 = A.search(WordHappens2)
A.table(WordHappens2, start=1, end=2, condensed=False, colorMap={3:'pink', 4:'lime'})
WordHappens3='''
clause typ=WayX
phrase function=Pred
word lex=HJH[ nu=sg
phrase function=Subj
:: word lex#DBR/ sp#nmpr lex#>LHJM/
'''
WordHappens3 = A.search(WordHappens3)
A.table(WordHappens3, start=1, end=20, condensed=False, colorMap={3:'pink', 4:'lime'})
WordHappens4='''
clause typ=WayX
phrase function=Pred
word lex=HJH[ nu=sg
phrase function=Subj
word lex=DBR/ nu=sg st=c
<: word sp=nmpr
'''
WordHappens4 = A.search(WordHappens4)
A.table(WordHappens4, start=1, end=20, condensed=False, colorMap={3:'pink', 4:'lime'})
# ## Prophet
# ### Task 2:
#
# Search in Bible Software (Logos/Accordance) how often NBJ> appears in the Bible. What insights could you generate from your findings?
NBJ='''
word lex=NBJ>/
'''
NBJ = A.search(NBJ)
A.table(NBJ, start=1, end=2, condensed=True)
A.export(NBJ, toDir='D:/OneDrive/1200_AUS-research/Fabric-TEXT', toFile='NBJ.tsv')
NBJ=pd.read_csv('D:/OneDrive/1200_AUS-research/Fabric-TEXT/NBJ.tsv',delimiter='\t',encoding='utf-16')
pd.set_option('display.max_columns', 50)
NBJ.head()
NBJ["S1"].value_counts()
# ## I am with you
# ### Task 3:
#
# Study the construction "I am with you" and create searches that help you to find these and other similar constructions of that nature. What links (if any) do you think are estabslihed by Jer's word usage?
withYou1='''
clause typ=NmCl
phrase function=PreC
word lex=>T==|<M
phrase function=Subj
word lex=>NKJ|>NJ
'''
withYou1 = A.search(withYou1)
A.table(withYou1, start=1, end=20, condensed=False, colorMap={3:'pink', 4:'lime'})
withYou2='''
chapter
clause typ=NmCl
phrase function=PreC
word lex=>T==|<M
phrase function=Subj
word lex=>NKJ|>NJ
clause
word lex=JR>[
'''
withYou2 = A.search(withYou2)
A.table(withYou2, start=1, end=20, condensed=False, colorMap={3:'pink', 4:'lime', 7:'blue', 8:'cyan'})
# ## Object of JYR[
# ### Task 4:
#
# Try to understand what the meaning of JYR is by searching for its typical objects (Objc, PreO).
JYR1='''
clause
phrase function=PreO
word lex=JYR[
'''
JYR1 = A.search(JYR1)
A.table(JYR1, start=1, end=20, condensed=True)
JYR2='''
clause
phrase function=Pred
word lex=JYR[
phrase function=Objc
word lex*
'''
JYR2 = A.search(JYR2)
A.table(JYR2, start=1, end=2, condensed=False, colorMap={3:'pink', 4:'lime'})
# ## Rhetoric
# ### Task 5 (Voluntary):
#
# Do you find an interesting rhetoric construction in Jer 1:11-12? If so, which one is it? Think about how you would build a query that would be able to find more of these cases. How do you think this could be done?
DiffLexSameCons = '''
chapter book=Jeremia chapter=1
w1:word
<20: w2:word
w1 < w2
w1 .g_cons=g_cons. w2
w1 .lex#lex. w2
'''
DiffLexSameCons = A.search(DiffLexSameCons)
A.table(DiffLexSameCons, start=1, end=2, condensed=True)
|
profNBs/0000_ETCBC-TF_BHS_0004_Jer01_o.glanz_prof-edition.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os as os
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib as mpl
import glob as glob
from scipy import stats
# ### Function to import kofamscan outputs for each MAG, remove empty rows, add col with MAG name
# +
#The D7 MAGS will have to be re-named to fit this scheme
x = "human_MAGs_kofams/H1.001.fasta.faa.kofam.filt"
def make_mag_ko_matrix(file):
a = file.split('.')
name = a[0] + '_' + a[1]
ko = pd.read_csv(file, sep='\t', names=[])
print(name)
make_mag_ko_matrix(x)
# +
# Try to run Roth's script to convert the kofamscan results into a table.
#os.chdir('cow_MAGs_kofams/')
# #%run -i 03c_KofamScan_Filter_Convert.py -i 'cow_MAGs_kofams/cow4.001.fasta.faa.kofam.filt' -o test
for file in "*MAGs_kofams/*.filt":
a = file.split('.')
name = a[0] + '_' + a[1]
# %run -i 01b_Parse_KEGG_Hierarchy.py -i file -o name.table
# -
# ### Import the KO count matrix generated from previous steps and pivot the rows and colums
# +
KO_counts = pd.read_csv("KO_counts/KO_counts_table.txt", sep='\t', index_col=0)
#KO_counts.shape => (3562, 81)
KO_counts_pivot = pd.pivot_table(KO_counts,columns="Tag")
KO_counts_pivot.tail()
# +
##drop columns whose sum is <2 (i.e. singleton KOS found in only 1 MAG)!
sum = KO_counts_pivot.sum(axis=0)
KO_lessThan2 = list()
KO_lessThan3 = list()
for col in range(len(sum)):
if sum[col] < 2:
KO_lessThan2.append(col)
#len(KO_lessThan2) => 757 KOs are singletons (only found in 1 MAG)
#len(KO_lessThan3) => 1098 KOs are doubletons
KO_drop_singletons = KO_counts_pivot.loc[: , (KO_counts_pivot.sum(axis=0) !=1)]
KO_drop_singletons.tail()
# -
# ### Run t-tests on KO values
# +
## Add rows (i.e. Series) to the dataframe
KO_ttest = KO_counts_pivot.append(pd.Series(name='ttest'))
KO_ttest = KO_ttest.append(pd.Series(name='p-value'))
print(KO_ttest.shape)
ttest = KO_ttest.iloc[81]
type(ttest)
pval = KO_ttest.iloc[82]
##Split the df up by host type:
h_c = KO_ttest.iloc[:31, :]
h_c = h_c.append(pd.Series(name='ttest'))
h_c = h_c.append(pd.Series(name='p-value'))
# ## rm the KO COLUMNS that are zero across all rows
h_c = h_c.loc[: , (h_c.sum(axis=0) !=1)]
h_c.shape
# +
## Fill new rows with results for scipy t-tests
human_MAGS = KO_ttest.index.values[:13]
cow = KO_ttest.index.values[13:31]
pig = KO_ttest.index.values[31:81]
KO_ttest.iloc[81] = np.array(stats.ttest_ind(KO_ttest.iloc[:13, :], KO_ttest.iloc[13:31, :], axis=0, equal_var=False))[0]
KO_ttest.iloc[82] = np.array(stats.ttest_ind(KO_ttest.iloc[:13, :], KO_ttest.iloc[13:31, :], axis=0, equal_var=False))[1]
KO_ttest.tail()
# -
## Run the t-tests
h_c.iloc[31] = np.array(stats.ttest_ind(h_c.iloc[:13, :], h_c.iloc[13:31, :], axis=0, equal_var=False))[0]
h_c.iloc[32] = np.array(stats.ttest_ind(h_c.iloc[:13, :], h_c.iloc[13:31, :], axis=0, equal_var=False))[1]
h_c.iloc[31]
# ### Funciton to calc fold difference between MAGs from different host type
def fold_difference(df, num_symp_rows, num_sample_rows):
set1 = df.iloc[0:num_symp_rows]
set2 = df.iloc[num_symp_rows:num_sample_rows]
mean_diff = np.mean(set1) / np.mean(set2)
#print(mean_diff)
return(mean_diff)
# #### Apply func to df and calcualte FOLD difference of each column
h_c = h_c.replace(0, 0.001)
#np.log2(h_c)
h_c.loc['fold change']= fold_difference(h_c, 13, 31)
h_c.tail()
# #### Select only the KOs that are significantly different between the groups (P<0.01)
# +
h_c_signif = h_c.loc[:, h_c.loc['p-value'] < 0.01]
h_c_signif.shape
# (34, 81) --> only 81 KOs are significant
## Save the list of Significantly Different KOs
h_c_sd_list = list(h_c_signif)
# +
## import the mast KO htext file
ko_master = "KO_Orthology_ko00001.txt"
## Get the total list of KOs:
kos_unfilt = list(KO_counts_pivot.columns)
with open(ko_master, 'r') as file, open('MAG_KOs_funcs_unfilt.tsv', 'w') as outfile:
for line in file:
X = line.rstrip().split('\t')
konumber = X[3].split(' ')[0]
if konumber in kos_unfilt:
outfile.write(line)
# -
with open(ko_master, 'r') as file, open('MAG_KOs_funcs_unfilt.tsv', 'w') as outfile:
for line in file:
X = line.rstrip().split('\t')
konumber = X[3].split(' ')[0]
if konumber in kos_unfilt:
outfile.write(line)
kos = pd.read_csv('MAG_KOs_funcs_unfilt.tsv', sep='\t', names=['Group', 'Subgroup1', 'Subgroup2', 'KO'])
print(kos.Group.unique())
print(kos.Subgroup1.unique())
kos.shape
# +
## Remove irrelevant KEGG categories ( not relevant to microbial gene functions)
rm_groups = ['Organismal Systems','Human Diseases']
kos2 = kos.loc[~kos['Group'].str.contains('|'.join(rm_groups))]
kos2.shape
# -
# #### Reformat the df tomake it more manageable/readable
kos2[['KO', 'Function']] = kos2['KO'].str.split(" ", n=1, expand=True)
#kos2 = kos.drop('PATH', axis=1)
kos2.head()
## Split out the "PATH" part of Subgroup2 label
kos2[['Subgroup2', 'PATH']] = kos2['Subgroup2'].str.split("[", n=1, expand=True)
kos2 = kos2.drop('PATH', axis=1)
kos2.head()
## Select only the KOs that are in the h_c_signif dataframe
h_c_sd_funct = kos2.loc[kos2['KO'].isin(h_c_sd_list), :]
h_c_sd_funct.shape
# ## Format for heatmap
# ### Exclude the last two rows (p-value and t-test)
df_hmap = h_c_signif.iloc[:31]
df_hmap
# +
#Sq. root transform if needed
#df_hmap_sqrt = np.sqrt(df_hmap.replace(0, 0.00000000001))
# -
# Quick and dirty to check
p = sns.clustermap(df_hmap, cmap='YlGnBu', figsize=(20,10), xticklabels=True, yticklabels=True,
row_cluster=False, method='ward', metric='euclidean')
p.savefig("Cow_Hum_SD_KOs_heatmap.png")
|
KO_MAG_t-test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## 第九章 幻灯片制作
#
# ### 导言
#
# 2003年,柏林工业大学的博士生<NAME>发布了一款用于制作演示文稿的工具Beamer。Beamer是Till Tantau利用业余时间开发的,他的初衷是方便自己使用LaTeX制作幻灯片,不过出乎意料的是,后来Beamer的受欢迎程度完全超出了他的想象。在Till Tantau开发Beamer期间,他收到了很多人的建议和反馈,这些都直接推动了开发工作。2010年,Till Tantau将Beamer交由他人维护和管理。
#
# Beamer作为LaTeX的一种特殊文档类型,它的出现无疑丰富了LaTeX制作演示文稿的功能。虽然Beamer并非LaTeX中第一款用于制作演示文稿的工具,但直到今日,Beamer却是最受大家欢迎的一款。Beamer的使用方式简单灵活,只需设定LaTeX文档类型为beamer,便可开始创作。同时,Beamer提供了大量的幻灯片模板,这些模板包含了丰富多彩的视觉效果,创作者可以直接使用。毫不夸张地说,Beamer的出现极大地提高了人们使用LaTeX制作幻灯片的热情。值得一提的是,在2005年,Till Tantau又发布了一款非常便捷的LaTeX绘图工具TikZ。TikZ不仅可以辅助Beamer幻灯片的制作,也可以应用于科技文档中的各类绘图任务。
#
# Beamer是随着LaTeX的发展而衍生出来的一种特殊文档类型,也常常被看作是一个功能强大的宏包,可以支撑科研工作者制作幻灯片的需求。使用Beamer制作幻灯片与Office办公软件(如PowerPoint)完全不同,从本质上来说,使用Beamer制作幻灯片其实和LaTeX中的其他文档类型没有太大区别:代码结构都是由前导代码和主体代码组成,完全沿用了LaTeX的文档环境与基本命令。因此,使用Beamer制作幻灯片也有一个缺点,那就是必须掌握LaTeX制作文档的基础。
#
# 在呈现方式上,Beamer制作的幻灯片最终会在LaTeX中被编译成PDF文档,在不同的系统上打开幻灯片时不存在不兼容等问题。在功能上,使用Beamer制作幻灯片时,我们可以对常规文本、公式、列表、图表甚至动画效果、视觉效果和主题样式等进行调整,最终达到想要的视觉效果。
#
# 事实上,LaTeX中用于制作演示文稿的工具并非只有Beamer,但相比其他工具,Beamer具有如下优点:
#
# - 拥有海量的模板和丰富的主题样式,且使用方便;
# - 能满足制作幻灯片的功能性需求,从创建标题、文本和段落到插入图表、参考文献等操作,且与常规文档中的使用规则几乎一致;
# - 使用方式简单,在主体代码中使用`\begin{frame} \end{frame}`环境就能创建一页幻灯片。
#
# 本章主要包括以下内容:Beamer的基本使用方式、在演示稿中添加动画效果、添加文本框等框元素、设置主题样式、插入程序源代码、添加参考文献、插入表格、插入与调整图片。
# 【继续】[**9.1 基本介绍**](https://nbviewer.jupyter.org/github/xinychen/latex-cookbook/blob/main/chapter-9/section1.ipynb)
# ### License
#
# <div class="alert alert-block alert-danger">
# <b>This work is released under the MIT license.</b>
# </div>
|
chapter-9/section0.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Realtime VAD
#
# Let say you want to cut your realtime recording audio by using VAD, malaya-speech able to do that.
# <div class="alert alert-info">
#
# This tutorial is available as an IPython notebook at [malaya-speech/example/realtime-vad](https://github.com/huseinzol05/malaya-speech/tree/master/example/realtime-vad).
#
# </div>
# <div class="alert alert-info">
#
# This module is language independent, so it save to use on different languages. Pretrained models trained on multilanguages.
#
# </div>
# <div class="alert alert-warning">
#
# This is an application of malaya-speech Pipeline, read more about malaya-speech Pipeline at [malaya-speech/example/pipeline](https://github.com/huseinzol05/malaya-speech/tree/master/example/pipeline).
#
# </div>
import malaya_speech
from malaya_speech import Pipeline
# ### Load VAD model
#
# Fastest and common model people use, is webrtc. Read more about VAD at https://malaya-speech.readthedocs.io/en/latest/load-vad.html
webrtc = malaya_speech.vad.webrtc()
# ### Recording interface
#
# So, to start recording audio including realtime VAD, we need to use `malaya_speech.streaming.record`. We use `pyaudio` library as the backend.
#
# ```python
# def record(
# vad,
# asr_model = None,
# classification_model = None,
# device = None,
# input_rate: int = 16000,
# sample_rate: int = 16000,
# blocks_per_second: int = 50,
# padding_ms: int = 300,
# ratio: float = 0.75,
# min_length: float = 0.1,
# filename: str = None,
# spinner: bool = False,
# ):
# """
# Record an audio using pyaudio library. This record interface required a VAD model.
#
# Parameters
# ----------
# vad: object
# vad model / pipeline.
# asr_model: object
# ASR model / pipeline, will transcribe each subsamples realtime.
# classification_model: object
# classification pipeline, will classify each subsamples realtime.
# device: None
# `device` parameter for pyaudio, check available devices from `sounddevice.query_devices()`.
# input_rate: int, optional (default = 16000)
# sample rate from input device, this will auto resampling.
# sample_rate: int, optional (default = 16000)
# output sample rate.
# blocks_per_second: int, optional (default = 50)
# size of frame returned from pyaudio, frame size = sample rate / (blocks_per_second / 2).
# 50 is good for WebRTC, 30 or less is good for Malaya Speech VAD.
# padding_ms: int, optional (default = 300)
# size of queue to store frames, size = padding_ms // (1000 * blocks_per_second // sample_rate)
# ratio: float, optional (default = 0.75)
# if 75% of the queue is positive, assumed it is a voice activity.
# min_length: float, optional (default=0.1)
# minimum length (s) to accept a subsample.
# filename: str, optional (default=None)
# if None, will auto generate name based on timestamp.
# spinner: bool, optional (default=False)
# if True, will use spinner object from halo library.
#
#
# Returns
# -------
# result : [filename, samples]
# """
# ```
# **Once you start to run the code below, it will straight away recording your voice**. Right now I am using built-in microphone.
#
# If you run in jupyter notebook, press button stop up there to stop recording, if in terminal, press `CTRL + c`.
file, samples = malaya_speech.streaming.record(webrtc)
file
# get the audio at [malaya-speech/speech/record](https://github.com/huseinzol05/malaya-speech/tree/master/speech/record).
# As you can see, it will automatically save to a file, and you can check the length of samples.
len(samples)
# means, we got 4 subsamples!
import IPython.display as ipd
ipd.Audio(file)
ipd.Audio(malaya_speech.astype.to_ndarray(samples[0][0]), rate = 16000)
ipd.Audio(malaya_speech.astype.to_ndarray(samples[1][0]), rate = 16000)
ipd.Audio(malaya_speech.astype.to_ndarray(samples[2][0]), rate = 16000)
ipd.Audio(malaya_speech.astype.to_ndarray(samples[3][0]), rate = 16000)
# ### Use pipeline
#
# We know, webrtc does not work really good in noisy environment, so to improve that, we can use VAD deep model from malaya-speech.
vad_model = malaya_speech.vad.deep_model(model = 'vggvox-v2', quantized = True)
# **pyaudio will returned int16 bytes, so we need to change to numpy array, normalize it to -1 and +1 floating point**.
p = Pipeline()
pipeline = (
p.map(malaya_speech.astype.to_ndarray)
.map(malaya_speech.astype.int_to_float)
.map(vad_model)
)
p.visualize()
# **Once you start to run the code below, it will straight away recording your voice**. Right now I am using built-in microphone.
#
# If you run in jupyter notebook, press button stop up there to stop recording, if in terminal, press `CTRL + c`.
file, samples = malaya_speech.streaming.record(p)
file
# get the audio at [malaya-speech/speech/record](https://github.com/huseinzol05/malaya-speech/tree/master/speech/record).
len(samples)
ipd.Audio(file)
ipd.Audio(malaya_speech.astype.to_ndarray(samples[0]), rate = 16000)
ipd.Audio(malaya_speech.astype.to_ndarray(samples[1]), rate = 16000)
ipd.Audio(malaya_speech.astype.to_ndarray(samples[2]), rate = 16000)
ipd.Audio(malaya_speech.astype.to_ndarray(samples[3]), rate = 16000)
|
example/realtime-vad/realtime-vad.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="85sNl1ailHU2"
# # Práctico 9: Difusión de epidemias
# + [markdown] id="nrZXWpp8m-12"
# # Introducción
#
# El objectivo de este práctico es explorar como la topología de la red de contactos (grafo de potenciales contagios entre individuos) puede impactar en el comportamiento de una epidemia.
#
# Para esto, vamos a implementar un simulador que nos ayude a corroborar hipótesis acerca de la difusión de procesos dinámicos en grafos.
# + id="c228a09d"
# %matplotlib inline
# + id="157696a5"
import networkx as nx
import pandas as pd
import numpy as np
import operator
import matplotlib.pyplot as plt
from ipywidgets import interact, fixed, IntSlider
from matplotlib.patches import Patch
plt.rcParams["figure.figsize"] = (12, 8)
# + id="566e910d"
STATE = "state"
STATE_HISTORY = "state_history"
SUSCEPTIBLE = 0
INFECTED = 1
RECOVERED = 2
STATE_COLORS = {
SUSCEPTIBLE: "blue",
INFECTED: "red",
RECOVERED: "green"
}
SEED = 42
# + id="c0f5e672"
def SIR_node_update(cur_state, neig_states, random_state, beta, gamma, **kwargs):
"""
This function implementsthe SIR model node update. It receives the
state of a node, the state of all nodes surrounding it and calculates
the probabilistic node update based on the SIR model formulation.
Paramteres
----------
cur_state: int
The state of the node for which the update will be calculated
neig_states: list of ints
The state of all the neighbours of the current state.
random_state: np.random.RandomState
This is the random state generator object. Use it to generate
any random number required within this function. This gurantees
a consisten output.
beta: float
The beta paramteter from the SIR model
gamma: float
The gamma paramters from the SIR model
"""
### START CODE HERE
### END CODE HERE
return new_state
# + id="c1e80e59"
def simulate_difussion(G, initial_states, random_state, update_func = None, **kwargs):
if update_func is None:
update_func = SIR_node_update
[G.nodes[k].update({STATE: v}) for k, v in initial_states.items()]
[G.nodes[k].update({STATE_HISTORY: []}) for k, v in initial_states.items()]
T = 1
while True:
new_states = dict()
for cur in G.nodes():
neigs = list(G.neighbors(cur))
state_cur = G.nodes[cur][STATE]
state_neigs = [G.nodes[n][STATE] for n in neigs]
new_state = update_func(state_cur, state_neigs, random_state, **kwargs)
new_states[cur] = new_state
for cur in G.nodes():
old_state = G.nodes[cur][STATE]
new_state = new_states[cur]
G.nodes[cur][STATE_HISTORY].append(old_state)
G.nodes[cur].update({STATE: new_state})
if sum(v == INFECTED for v in new_states.values()) == 0:
break
else:
T += 1
T += 1
for n in G.nodes(): G.nodes[n][STATE_HISTORY].append(G.nodes[n][STATE])
node_colors = dict(
(t, [STATE_COLORS[v[STATE_HISTORY][t]] for k, v in G.nodes(data=True)]) for t in range(T)
)
return node_colors, T
# + id="6b5f7f9c"
def plot_difussion(t, G, colors, layout):
legend_elements = [
Patch(facecolor=STATE_COLORS[SUSCEPTIBLE], edgecolor='k', label="Susceptible"),
Patch(facecolor=STATE_COLORS[INFECTED], edgecolor='k', label='Infected'),
Patch(facecolor=STATE_COLORS[RECOVERED], edgecolor='k', label='Recovered')
]
nx.draw_networkx_nodes(G, pos=layout, node_color=node_colors[t])
_ = nx.draw_networkx_labels(G, pos=layout)
nx.draw_networkx_edges(G, pos=layout)
plt.legend(handles=legend_elements, loc="upper center", bbox_to_anchor=(0.5, 1.1), ncol=3)
plt.title(f"Timeslot: {t}")
plt.show()
# + id="29B3u11uKj6-"
def stats_difussion(G, view=True):
t_range = len(G.nodes[0][STATE_HISTORY])
susceptible = [sum([v[STATE_HISTORY][t] == SUSCEPTIBLE for k, v in G.nodes(data=True)]) for t in range(t_range)]
infected = [sum([v[STATE_HISTORY][t] == INFECTED for k, v in G.nodes(data=True)]) for t in range(t_range)]
recovered = [sum([v[STATE_HISTORY][t] == RECOVERED for k, v in G.nodes(data=True)]) for t in range(t_range)]
if view:
plt.plot(range(t_range), susceptible, color=STATE_COLORS[SUSCEPTIBLE], linestyle = 'dashed', alpha=0.3)
plt.plot(range(t_range), infected, color=STATE_COLORS[INFECTED])
plt.plot(range(t_range), recovered, color=STATE_COLORS[RECOVERED], linestyle = 'dashed', alpha=0.3)
legend_elements = [
Patch(facecolor=STATE_COLORS[SUSCEPTIBLE], edgecolor='k', label="Susceptible"),
Patch(facecolor=STATE_COLORS[INFECTED], edgecolor='k', label='Infected'),
Patch(facecolor=STATE_COLORS[RECOVERED], edgecolor='k', label='Recovered')
]
plt.legend(handles=legend_elements, loc="upper center", bbox_to_anchor=(0.5, 1.1), ncol=3)
plt.title("Epidemic evolution")
plt.xlabel("Time")
plt.ylabel("# Population")
plt.show()
return susceptible,infected,recovered
# + id="FGCO6UZ4Tsfx"
def stats_trials(infected_trials, title='Epidemic evolution', view=True):
t_range = max([len(i) for i in infected_trials])
sum_trials = np.zeros((t_range))
len_trials = np.zeros((t_range))
for i in infected_trials:
i = np.array(i)
l = np.ones(i.shape)
l.resize(sum_trials.shape)
len_trials = len_trials + l
i.resize(sum_trials.shape)
sum_trials = sum_trials + i
infected_mean = sum_trials/len_trials
if view:
for i in range(len(infected_trials)):
plt.plot(range(len(infected_trials[i])), infected_trials[i], color=STATE_COLORS[INFECTED], alpha=0.1)
plt.plot(range(len(infected_mean)), infected_mean, color=STATE_COLORS[INFECTED])
plt.title(title)
plt.xlabel("Time")
plt.ylabel("# Population")
plt.show()
return infected_mean
# + [markdown] id="3W-VDXx4PpXw"
# # 1) Fundamentos en los procesos dinámicos de epidemias
#
#
# Seguir la Sección 8.5 del libro [SANDR], entendiendo los procesos epidémicos tradicionales del tipo SIR.
# + [markdown] id="3182a511"
# #2) Primer ejemplo, difusión en grafos lineales (caminos)
#
# Para empezar a explorar el primer modelo de difusión vamos a trabajar con uno de los grafos más simples de todos: el camino.
#
# El grafo camino es interesante desde el punto de vista de una epidemia, dado que una vez que un nodo se recupera, este impide la difusión del grafo entre los dos grupos de nodos a sus lados.
#
# Para empezar vamos a generar un grafo.
# + id="f08c8d06"
N=10
G = nx.path_graph(N)
initial_states = dict((n, SUSCEPTIBLE) for n in G.nodes())
initial_states[1] = INFECTED
layout = nx.spring_layout(G)
# + id="d185d4d7"
r = np.random.RandomState(SEED - 1)
node_colors, total_timeslots = simulate_difussion(
G,
initial_states,
r,
SIR_node_update,
beta=0.25,
gamma=0.1
)
# + id="30cbb2dd"
len(G.nodes(data=True)[8][STATE_HISTORY])
# + [markdown] id="JgqkkmweGH1q"
# Es posible ver la difusión de la epidemia, moviendose en el tiempo utilizando la barra de la siguiente figura.
# + id="98ece1d4"
interact(
plot_difussion,
t=IntSlider(value=0, min=0, max=total_timeslots - 1),
G=fixed(G),
colors=fixed(node_colors),
layout=fixed(layout)
)
# + [markdown] id="exBoJgqIOx3_"
# También podemos ver como evoluciona la cantidad de infectados.
# + id="ESyt4EAyLk2N"
_,_,_ = stats_difussion(G)
# + [markdown] id="c37eecfa"
# #3) Difusión en grafo Erdös-Renyi
#
# Como segundo ejemplo de difusión vamos a considerar el caso de un grafo aleatorio Erdös-Renyi
# + id="581cb8c4"
N, p = 28, 0.3
G = nx.erdos_renyi_graph(n=N, p=p, seed=SEED)
initial_states = dict((n, SUSCEPTIBLE) for n in G.nodes())
initial_states[1] = INFECTED
layout = nx.spring_layout(G)
# + id="ae757ca5"
r = np.random.RandomState(SEED)
node_colors, total_timeslots = simulate_difussion(
G,
initial_states,
r,
SIR_node_update,
beta=0.1,
gamma=0.1
)
# + id="3a40b407"
interact(
plot_difussion,
t=IntSlider(value=0, min=0, max=total_timeslots - 1),
G=fixed(G),
colors=fixed(node_colors),
layout=fixed(layout)
)
_,_,_ = stats_difussion(G)
# + [markdown] id="0f1758c4"
# ##3.1) ¿Qué puede decir de la diferencia de velocidad con la que el proceso se esparse a través de la red para el caso del grafo camino y para el grafo Erdös-Renyi?
# + id="428b777b"
### START CODE HERE
### END CODE HERE
# + [markdown] id="AoD7BD6Rxm0F"
# # 4) Difusión en grafos de distintos modelos
#
# Compararemos la difusión de la epidemia para los modelos:
#
# * Erdös-Renyi
# * <NAME>
# * <NAME>
#
# Para ver estadísticamente el resultado, realizaremos 20 pruebas por modelo, donde sortearemos la toppología y el vértice inicial.
#
# Para ser justos, intentaremos que todos los grafos tengan 250 vértices y 1250 aristias (aproximadamente).
#
# + id="y20ZZgEwxm0G"
#parámetros en común
N = 250 #vertices
#epidemia
beta=0.5/20
gamma=1.0/20
trials = 20 #pruebas
#mismo estado inicial para todos los grafos
initial_states = dict((n, SUSCEPTIBLE) for n in range(N))
initial_infected = np.random.randint(0,N-1, size=1)[0]
#initial_states[1] = INFECTED
initial_states[initial_infected] = INFECTED
# + [markdown] id="sMx8HNVQxm0H"
# Simular Erdös-Renyi
# + id="JY8Nk3Njxm0H"
p = 0.041
infected_trials = []
for i in range(trials):
G = nx.erdos_renyi_graph(n=N, p=p)
node_colors, total_timeslots = simulate_difussion(
G,
initial_states,
np.random.RandomState(SEED),
SIR_node_update,
beta=beta,
gamma=gamma
)
print(G.number_of_edges())
_, infected_i,_ = stats_difussion(G, view=False)
infected_trials.append(infected_i)
infected_mean_erdosrenyi = stats_trials(infected_trials, title="Epidemic evolution: Erdös-Renyi")
# + [markdown] id="BtlVOwMUxm0M"
# Simular <NAME>
# + id="L2MECbSjxm0M"
### START CODE HERE
### END CODE HERE
infected_mean_barabasi = stats_trials(infected_trials, title="Epidemic evolution: Barabasi Albert")
# + [markdown] id="KUQOWdrLxm0N"
# Simular Watts Strogatz
# + id="_Mk_umRCxm0O"
### START CODE HERE
### END CODE HERE
infected_mean_ws = stats_trials(infected_trials, title="Epidemic evolution: Watts Strogatz")
# + [markdown] id="XRgbR9mD3BTG"
# Resultado de la comparación
# + id="eUHsFO3c1-UQ"
plt.plot(infected_mean_erdosrenyi, color='red')
plt.plot(infected_mean_barabasi, color='green')
plt.plot(infected_mean_ws, color='blue')
plt.title("Evolución promedio de los distintos modelos")
plt.xlabel("Time")
plt.ylabel("# Population")
legend_elements = [
Patch(facecolor='red', edgecolor='k', label="<NAME>"),
Patch(facecolor='green', edgecolor='k', label='<NAME>'),
Patch(facecolor='blue', edgecolor='k', label='<NAME>')
]
plt.legend(handles=legend_elements, loc="upper center", bbox_to_anchor=(0.5, 1.1), ncol=3)
plt.show()
# + [markdown] id="f40bfb23"
# #5) Limitando el número máximo de infectados
#
# En esta sección intentaremos encontrar buenas estrategias para limitar y disminuir la máxima cantidad de nodos infectados a lo largo del proceso.
#
# Para esto vamos a permitirnos `inmunizar` a dos nodos (asignarles el estado inicial `RECUPERADO`) y evaluar el efecto de tal política en la epidemia.
#
# Para estos experimentos vamos a usar el [grafo de Tutte](https://en.wikipedia.org/wiki/Tutte_graph)
#
# Para simplificar la experimentación, vamos a asumir que la epidemia comienza en el nodo `3`.
# + id="d687ea87"
G = nx.tutte_graph()
layout = nx.spring_layout(G, iterations=300, seed=SEED)
initial_states = dict((n, SUSCEPTIBLE) for n in G.nodes())
initial_states[3] = INFECTED
# + id="92a96023"
nx.draw_networkx(G, pos=layout)
# + id="f0ccd176"
r = np.random.RandomState(SEED)
node_colors, total_timeslots = simulate_difussion(
G,
initial_states,
r,
SIR_node_update,
beta=0.3,
gamma=0.05
)
# + id="accc091c"
interact(
plot_difussion,
t=IntSlider(value=0, min=0, max=total_timeslots - 1),
G=fixed(G),
colors=fixed(node_colors),
layout=fixed(layout)
)
# + id="63ba891f"
def maximum_infected(G):
"""
Calculates the total maximum of infected nodes
in any given timeslot.
Parameters:
------------
G: nx.Graph
The graph of the network. Assume that the epidemic has
already been simulated and that each node in the graph
contains the attribute `STATE_HISTORY`
Returns
-------
max_infected: int
The maximum number of infected nodes in any timeslot.
"""
### START CODE HERE
### END CODE HERE
# + id="d5e42942"
assert maximum_infected(G) == 29 # Hay como máximo 29 infectados en la corrida por defecto
# + [markdown] id="de40a405"
# ##5.1) Eligiendo el nodo para "inmunizar"
#
# Vamos a comprar las siguientes estrategias de inmunización:
#
# * Un vécino del primer nodo infectado (19)
# * El nodo con mayor `betweeness centrality`
# * Un nodo poco relacionado (nodo 37)
#
# Todos los otros parámetros permaneceran iguales.
# + [markdown] id="84361dfc"
# ### Estratégia: Un vécino
# + id="cddaaf21"
initial_states_copy = initial_states.copy()
initial_states_copy[19] = RECOVERED
# + id="ce82f5f0"
r = np.random.RandomState(SEED)
node_colors, total_timeslots = simulate_difussion(
G,
initial_states_copy,
r,
SIR_node_update,
beta=0.3,
gamma=0.05
)
# + id="f92d009a"
maximum_infected(G)
# + [markdown] id="3d2bdfd7"
# Logramos reducir la maxima cantidad de infectados en 1.
# + [markdown] id="adff5171"
# ### Estrategia: Mayor `betweeness centrality`
# + id="1799accb"
### START CODE HERE
### END CODE HERE
# + id="83d7af4c"
initial_states_copy = initial_states.copy()
initial_states_copy[selected_node] = RECOVERED
# + id="d358293b"
r = np.random.RandomState(SEED)
node_colors, total_timeslots = simulate_difussion(
G,
initial_states_copy,
r,
SIR_node_update,
beta=0.3,
gamma=0.05
)
# + id="1906a99b"
maximum_infected(G)
# + [markdown] id="597caa16"
# Logramos reducir la maxima cantidad de infectados en 5.
# + [markdown] id="8801683e"
# ### Estrategia: Nodo poco relacionado (37)
# + id="fe00ee00"
initial_states_copy = initial_states.copy()
initial_states_copy[37] = RECOVERED
# + id="a260345f"
r = np.random.RandomState(SEED)
node_colors, total_timeslots = simulate_difussion(
G,
initial_states_copy,
r,
SIR_node_update,
beta=0.3,
gamma=0.05
)
# + id="269c610e"
maximum_infected(G)
# + [markdown] id="00ba51b7"
# Logramos reducir la maxima cantidad de infectados en 2.
|
homework_09_epidemics/public_homework_09_epidemics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Gradient Checking
#
# Welcome to the final assignment for this week! In this assignment you will learn to implement and use gradient checking.
#
# You are part of a team working to make mobile payments available globally, and are asked to build a deep learning model to detect fraud--whenever someone makes a payment, you want to see if the payment might be fraudulent, such as if the user's account has been taken over by a hacker.
#
# But backpropagation is quite challenging to implement, and sometimes has bugs. Because this is a mission-critical application, your company's CEO wants to be really certain that your implementation of backpropagation is correct. Your CEO says, "Give me a proof that your backpropagation is actually working!" To give this reassurance, you are going to use "gradient checking".
#
# Let's do it!
# Packages
import numpy as np
from testCases import *
from gc_utils import sigmoid, relu, dictionary_to_vector, vector_to_dictionary, gradients_to_vector
# ## 1) How does gradient checking work?
#
# Backpropagation computes the gradients $\frac{\partial J}{\partial \theta}$, where $\theta$ denotes the parameters of the model. $J$ is computed using forward propagation and your loss function.
#
# Because forward propagation is relatively easy to implement, you're confident you got that right, and so you're almost 100% sure that you're computing the cost $J$ correctly. Thus, you can use your code for computing $J$ to verify the code for computing $\frac{\partial J}{\partial \theta}$.
#
# Let's look back at the definition of a derivative (or gradient):
# $$ \frac{\partial J}{\partial \theta} = \lim_{\varepsilon \to 0} \frac{J(\theta + \varepsilon) - J(\theta - \varepsilon)}{2 \varepsilon} \tag{1}$$
#
# If you're not familiar with the "$\displaystyle \lim_{\varepsilon \to 0}$" notation, it's just a way of saying "when $\varepsilon$ is really really small."
#
# We know the following:
#
# - $\frac{\partial J}{\partial \theta}$ is what you want to make sure you're computing correctly.
# - You can compute $J(\theta + \varepsilon)$ and $J(\theta - \varepsilon)$ (in the case that $\theta$ is a real number), since you're confident your implementation for $J$ is correct.
#
# Lets use equation (1) and a small value for $\varepsilon$ to convince your CEO that your code for computing $\frac{\partial J}{\partial \theta}$ is correct!
# ## 2) 1-dimensional gradient checking
#
# Consider a 1D linear function $J(\theta) = \theta x$. The model contains only a single real-valued parameter $\theta$, and takes $x$ as input.
#
# You will implement code to compute $J(.)$ and its derivative $\frac{\partial J}{\partial \theta}$. You will then use gradient checking to make sure your derivative computation for $J$ is correct.
#
# <img src="images/1Dgrad_kiank.png" style="width:600px;height:250px;">
# <caption><center> <u> **Figure 1** </u>: **1D linear model**<br> </center></caption>
#
# The diagram above shows the key computation steps: First start with $x$, then evaluate the function $J(x)$ ("forward propagation"). Then compute the derivative $\frac{\partial J}{\partial \theta}$ ("backward propagation").
#
# **Exercise**: implement "forward propagation" and "backward propagation" for this simple function. I.e., compute both $J(.)$ ("forward propagation") and its derivative with respect to $\theta$ ("backward propagation"), in two separate functions.
# +
# GRADED FUNCTION: forward_propagation
def forward_propagation(x, theta):
"""
Implement the linear forward propagation (compute J) presented in Figure 1 (J(theta) = theta * x)
Arguments:
x -- a real-valued input
theta -- our parameter, a real number as well
Returns:
J -- the value of function J, computed using the formula J(theta) = theta * x
"""
### START CODE HERE ### (approx. 1 line)
J = np.dot(theta, x)
### END CODE HERE ###
return J
# -
x, theta = 2, 4
J = forward_propagation(x, theta)
print ("J = " + str(J))
# **Expected Output**:
#
# <table style=>
# <tr>
# <td> ** J ** </td>
# <td> 8</td>
# </tr>
# </table>
# **Exercise**: Now, implement the backward propagation step (derivative computation) of Figure 1. That is, compute the derivative of $J(\theta) = \theta x$ with respect to $\theta$. To save you from doing the calculus, you should get $dtheta = \frac { \partial J }{ \partial \theta} = x$.
# +
# GRADED FUNCTION: backward_propagation
def backward_propagation(x, theta):
"""
Computes the derivative of J with respect to theta (see Figure 1).
Arguments:
x -- a real-valued input
theta -- our parameter, a real number as well
Returns:
dtheta -- the gradient of the cost with respect to theta
"""
### START CODE HERE ### (approx. 1 line)
dtheta = x
### END CODE HERE ###
return dtheta
# -
x, theta = 2, 4
dtheta = backward_propagation(x, theta)
print ("dtheta = " + str(dtheta))
# **Expected Output**:
#
# <table>
# <tr>
# <td> ** dtheta ** </td>
# <td> 2 </td>
# </tr>
# </table>
# **Exercise**: To show that the `backward_propagation()` function is correctly computing the gradient $\frac{\partial J}{\partial \theta}$, let's implement gradient checking.
#
# **Instructions**:
# - First compute "gradapprox" using the formula above (1) and a small value of $\varepsilon$. Here are the Steps to follow:
# 1. $\theta^{+} = \theta + \varepsilon$
# 2. $\theta^{-} = \theta - \varepsilon$
# 3. $J^{+} = J(\theta^{+})$
# 4. $J^{-} = J(\theta^{-})$
# 5. $gradapprox = \frac{J^{+} - J^{-}}{2 \varepsilon}$
# - Then compute the gradient using backward propagation, and store the result in a variable "grad"
# - Finally, compute the relative difference between "gradapprox" and the "grad" using the following formula:
# $$ difference = \frac {\mid\mid grad - gradapprox \mid\mid_2}{\mid\mid grad \mid\mid_2 + \mid\mid gradapprox \mid\mid_2} \tag{2}$$
# You will need 3 Steps to compute this formula:
# - 1'. compute the numerator using np.linalg.norm(...)
# - 2'. compute the denominator. You will need to call np.linalg.norm(...) twice.
# - 3'. divide them.
# - If this difference is small (say less than $10^{-7}$), you can be quite confident that you have computed your gradient correctly. Otherwise, there may be a mistake in the gradient computation.
#
# +
# GRADED FUNCTION: gradient_check
def gradient_check(x, theta, epsilon = 1e-7):
"""
Implement the backward propagation presented in Figure 1.
Arguments:
x -- a real-valued input
theta -- our parameter, a real number as well
epsilon -- tiny shift to the input to compute approximated gradient with formula(1)
Returns:
difference -- difference (2) between the approximated gradient and the backward propagation gradient
"""
# Compute gradapprox using left side of formula (1). epsilon is small enough, you don't need to worry about the limit.
### START CODE HERE ### (approx. 5 lines)
thetaplus = theta + epsilon # Step 1
thetaminus = theta - epsilon # Step 2
J_plus = thetaplus*x # Step 3
J_minus = thetaminus*x # Step 4
gradapprox = (J_plus - J_minus)/(2*epsilon) # Step 5
### END CODE HERE ###
# Check if gradapprox is close enough to the output of backward_propagation()
### START CODE HERE ### (approx. 1 line)
grad = x
### END CODE HERE ###
### START CODE HERE ### (approx. 1 line)
numerator = np.linalg.norm((grad-gradapprox),keepdims=True) # Step 1'
denominator = np.linalg.norm((grad),keepdims=True) + np.linalg.norm((gradapprox),keepdims=True) # Step 2'
difference = numerator/denominator # Step 3'
### END CODE HERE ###
if difference < 1e-7:
print ("The gradient is correct!")
else:
print ("The gradient is wrong!")
return difference
# -
x, theta = 2, 4
difference = gradient_check(x, theta)
print("difference = " + str(difference))
# **Expected Output**:
# The gradient is correct!
# <table>
# <tr>
# <td> ** difference ** </td>
# <td> 2.9193358103083e-10 </td>
# </tr>
# </table>
# Congrats, the difference is smaller than the $10^{-7}$ threshold. So you can have high confidence that you've correctly computed the gradient in `backward_propagation()`.
#
# Now, in the more general case, your cost function $J$ has more than a single 1D input. When you are training a neural network, $\theta$ actually consists of multiple matrices $W^{[l]}$ and biases $b^{[l]}$! It is important to know how to do a gradient check with higher-dimensional inputs. Let's do it!
# ## 3) N-dimensional gradient checking
# The following figure describes the forward and backward propagation of your fraud detection model.
#
# <img src="images/NDgrad_kiank.png" style="width:600px;height:400px;">
# <caption><center> <u> **Figure 2** </u>: **deep neural network**<br>*LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SIGMOID*</center></caption>
#
# Let's look at your implementations for forward propagation and backward propagation.
def forward_propagation_n(X, Y, parameters):
"""
Implements the forward propagation (and computes the cost) presented in Figure 3.
Arguments:
X -- training set for m examples
Y -- labels for m examples
parameters -- python dictionary containing your parameters "W1", "b1", "W2", "b2", "W3", "b3":
W1 -- weight matrix of shape (5, 4)
b1 -- bias vector of shape (5, 1)
W2 -- weight matrix of shape (3, 5)
b2 -- bias vector of shape (3, 1)
W3 -- weight matrix of shape (1, 3)
b3 -- bias vector of shape (1, 1)
Returns:
cost -- the cost function (logistic cost for one example)
"""
# retrieve parameters
m = X.shape[1]
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
W3 = parameters["W3"]
b3 = parameters["b3"]
# LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SIGMOID
Z1 = np.dot(W1, X) + b1
A1 = relu(Z1)
Z2 = np.dot(W2, A1) + b2
A2 = relu(Z2)
Z3 = np.dot(W3, A2) + b3
A3 = sigmoid(Z3)
# Cost
logprobs = np.multiply(-np.log(A3), Y) + np.multiply(-np.log(1 - A3), 1 - Y)
cost = 1. / m * np.sum(logprobs)
cache = (Z1, A1, W1, b1, Z2, A2, W2, b2, Z3, A3, W3, b3)
return cost, cache
# Now, run backward propagation.
def backward_propagation_n(X, Y, cache):
"""
Implement the backward propagation presented in figure 2.
Arguments:
X -- input datapoint, of shape (input size, 1)
Y -- true "label"
cache -- cache output from forward_propagation_n()
Returns:
gradients -- A dictionary with the gradients of the cost with respect to each parameter, activation and pre-activation variables.
"""
m = X.shape[1]
(Z1, A1, W1, b1, Z2, A2, W2, b2, Z3, A3, W3, b3) = cache
dZ3 = A3 - Y
dW3 = 1. / m * np.dot(dZ3, A2.T)
db3 = 1. / m * np.sum(dZ3, axis=1, keepdims=True)
dA2 = np.dot(W3.T, dZ3)
dZ2 = np.multiply(dA2, np.int64(A2 > 0))
dW2 = 1. / m * np.dot(dZ2, A1.T) * 2 # Should not multiply by 2
db2 = 1. / m * np.sum(dZ2, axis=1, keepdims=True)
dA1 = np.dot(W2.T, dZ2)
dZ1 = np.multiply(dA1, np.int64(A1 > 0))
dW1 = 1. / m * np.dot(dZ1, X.T)
db1 = 4. / m * np.sum(dZ1, axis=1, keepdims=True) # Should not multiply by 4
gradients = {"dZ3": dZ3, "dW3": dW3, "db3": db3,
"dA2": dA2, "dZ2": dZ2, "dW2": dW2, "db2": db2,
"dA1": dA1, "dZ1": dZ1, "dW1": dW1, "db1": db1}
return gradients
# You obtained some results on the fraud detection test set but you are not 100% sure of your model. Nobody's perfect! Let's implement gradient checking to verify if your gradients are correct.
# **How does gradient checking work?**.
#
# As in 1) and 2), you want to compare "gradapprox" to the gradient computed by backpropagation. The formula is still:
#
# $$ \frac{\partial J}{\partial \theta} = \lim_{\varepsilon \to 0} \frac{J(\theta + \varepsilon) - J(\theta - \varepsilon)}{2 \varepsilon} \tag{1}$$
#
# However, $\theta$ is not a scalar anymore. It is a dictionary called "parameters". We implemented a function "`dictionary_to_vector()`" for you. It converts the "parameters" dictionary into a vector called "values", obtained by reshaping all parameters (W1, b1, W2, b2, W3, b3) into vectors and concatenating them.
#
# The inverse function is "`vector_to_dictionary`" which outputs back the "parameters" dictionary.
#
# <img src="images/dictionary_to_vector.png" style="width:600px;height:400px;">
# <caption><center> <u> **Figure 2** </u>: **dictionary_to_vector() and vector_to_dictionary()**<br> You will need these functions in gradient_check_n()</center></caption>
#
# We have also converted the "gradients" dictionary into a vector "grad" using gradients_to_vector(). You don't need to worry about that.
#
# **Exercise**: Implement gradient_check_n().
#
# **Instructions**: Here is pseudo-code that will help you implement the gradient check.
#
# For each i in num_parameters:
# - To compute `J_plus[i]`:
# 1. Set $\theta^{+}$ to `np.copy(parameters_values)`
# 2. Set $\theta^{+}_i$ to $\theta^{+}_i + \varepsilon$
# 3. Calculate $J^{+}_i$ using to `forward_propagation_n(x, y, vector_to_dictionary(`$\theta^{+}$ `))`.
# - To compute `J_minus[i]`: do the same thing with $\theta^{-}$
# - Compute $gradapprox[i] = \frac{J^{+}_i - J^{-}_i}{2 \varepsilon}$
#
# Thus, you get a vector gradapprox, where gradapprox[i] is an approximation of the gradient with respect to `parameter_values[i]`. You can now compare this gradapprox vector to the gradients vector from backpropagation. Just like for the 1D case (Steps 1', 2', 3'), compute:
# $$ difference = \frac {\| grad - gradapprox \|_2}{\| grad \|_2 + \| gradapprox \|_2 } \tag{3}$$
# +
# GRADED FUNCTION: gradient_check_n
def gradient_check_n(parameters, gradients, X, Y, epsilon=1e-7):
"""
Checks if backward_propagation_n computes correctly the gradient of the cost output by forward_propagation_n
Arguments:
parameters -- python dictionary containing your parameters "W1", "b1", "W2", "b2", "W3", "b3":
grad -- output of backward_propagation_n, contains gradients of the cost with respect to the parameters.
x -- input datapoint, of shape (input size, 1)
y -- true "label"
epsilon -- tiny shift to the input to compute approximated gradient with formula(1)
Returns:
difference -- difference (2) between the approximated gradient and the backward propagation gradient
"""
# Set-up variables
parameters_values, _ = dictionary_to_vector(parameters)
grad = gradients_to_vector(gradients)
num_parameters = parameters_values.shape[0]
J_plus = np.zeros((num_parameters, 1))
J_minus = np.zeros((num_parameters, 1))
gradapprox = np.zeros((num_parameters, 1))
# Compute gradapprox
for i in range(num_parameters):
# Compute J_plus[i]. Inputs: "parameters_values, epsilon". Output = "J_plus[i]".
# "_" is used because the function you have to outputs two parameters but we only care about the first one
### START CODE HERE ### (approx. 3 lines)
thetaplus = np.copy(parameters_values) # Step 1
thetaplus[i][0] = thetaplus[i][0] + epsilon # Step 2
J_plus[i], _ = forward_propagation_n(X, Y, vector_to_dictionary(thetaplus)) # Step 3
### END CODE HERE ###
# Compute J_minus[i]. Inputs: "parameters_values, epsilon". Output = "J_minus[i]".
### START CODE HERE ### (approx. 3 lines)
thetaminus = np.copy(parameters_values) # Step 1
thetaminus[i][0] = thetaminus[i][0] - epsilon # Step 2
J_minus[i], _ = forward_propagation_n(X, Y, vector_to_dictionary(thetaminus)) # Step 3
### END CODE HERE ###
# Compute gradapprox[i]
### START CODE HERE ### (approx. 1 line)
gradapprox[i] = (J_plus[i] - J_minus[i]) / (2 * epsilon)
### END CODE HERE ###
# Compare gradapprox to backward propagation gradients by computing difference.
### START CODE HERE ### (approx. 1 line)
numerator = np.linalg.norm(grad - gradapprox) # Step 1'
denominator = np.linalg.norm(grad) + np.linalg.norm(gradapprox) # Step 2'
difference = numerator / denominator # Step 3'
### END CODE HERE ###
if difference > 1e-7:
print("\033[93m" + "There is a mistake in the backward propagation! difference = " + str(difference) + "\033[0m")
else:
print("\033[92m" + "Your backward propagation works perfectly fine! difference = " + str(difference) + "\033[0m")
return difference
# +
X, Y, parameters = gradient_check_n_test_case()
cost, cache = forward_propagation_n(X, Y, parameters)
gradients = backward_propagation_n(X, Y, cache)
difference = gradient_check_n(parameters, gradients, X, Y)
# -
# **Expected output**:
#
# <table>
# <tr>
# <td> ** There is a mistake in the backward propagation!** </td>
# <td> difference = 0.285093156781 </td>
# </tr>
# </table>
# It seems that there were errors in the `backward_propagation_n` code we gave you! Good that you've implemented the gradient check. Go back to `backward_propagation` and try to find/correct the errors *(Hint: check dW2 and db1)*. Rerun the gradient check when you think you've fixed it. Remember you'll need to re-execute the cell defining `backward_propagation_n()` if you modify the code.
#
# Can you get gradient check to declare your derivative computation correct? Even though this part of the assignment isn't graded, we strongly urge you to try to find the bug and re-run gradient check until you're convinced backprop is now correctly implemented.
#
# **Note**
# - Gradient Checking is slow! Approximating the gradient with $\frac{\partial J}{\partial \theta} \approx \frac{J(\theta + \varepsilon) - J(\theta - \varepsilon)}{2 \varepsilon}$ is computationally costly. For this reason, we don't run gradient checking at every iteration during training. Just a few times to check if the gradient is correct.
# - Gradient Checking, at least as we've presented it, doesn't work with dropout. You would usually run the gradient check algorithm without dropout to make sure your backprop is correct, then add dropout.
#
# Congrats, you can be confident that your deep learning model for fraud detection is working correctly! You can even use this to convince your CEO. :)
#
# <font color='blue'>
# **What you should remember from this notebook**:
# - Gradient checking verifies closeness between the gradients from backpropagation and the numerical approximation of the gradient (computed using forward propagation).
# - Gradient checking is slow, so we don't run it in every iteration of training. You would usually run it only to make sure your code is correct, then turn it off and use backprop for the actual learning process.
|
Hyperparameter Tuning/Gradient+Checking+v1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import os
import sys
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
df = pd.read_csv("combined_data_20211217.csv")
df.columns
# ## Exploration: TNR vs confidence score
df1 = df[['positive_confidence', 'rating', 'neutral_confidence', 'negative_confidence', 'ads_domain_total_count']]
df1
# +
df1 = df1[df1.positive_confidence != "ERROR"]
df1 = df1[df1['positive_confidence'].notna()]
df1 = df1[df1.neutral_confidence != "ERROR"]
df1 = df1[df1['neutral_confidence'].notna()]
df1 = df1[df1.negative_confidence != "ERROR"]
df1 = df1[df1['negative_confidence'].notna()]
df1
# -
df1["positive_confidence"] = pd.to_numeric(df1["positive_confidence"])
df1["neutral_confidence"] = pd.to_numeric(df1["neutral_confidence"])
df1["negative_confidence"] = pd.to_numeric(df1["negative_confidence"])
t_pos = df1.loc[df1['rating'] == 'T', 'positive_confidence'].mean()
t_pos
n_pos = df1.loc[df1['rating'] == 'N', 'positive_confidence'].mean()
n_pos
r_pos = df1.loc[df1['rating'] == 'R', 'positive_confidence'].mean()
r_pos
# assign data of lists.
data = {'rating': ['T', 'N', 'R'], 'positive_confidence': [t_pos, n_pos, r_pos]}
# Create DataFrame.
df2 = pd.DataFrame(data)
# Print the output.
print(df2)
t_neu = df1.loc[df1['rating'] == 'T', 'neutral_confidence'].mean()
n_neu = df1.loc[df1['rating'] == 'N', 'neutral_confidence'].mean()
r_neu = df1.loc[df1['rating'] == 'R', 'neutral_confidence'].mean()
data = {'rating': ['T', 'N', 'R'], 'neutral_confidence': [t_neu, n_neu, r_neu]}
# Create DataFrame.
dfp = pd.DataFrame(data)
# Print the output.
print(dfp)
df2['neutral_confidence'] = dfp['neutral_confidence']
df2
# +
t_neg = df1.loc[df1['rating'] == 'T', 'negative_confidence'].mean()
n_neg = df1.loc[df1['rating'] == 'N', 'negative_confidence'].mean()
r_neg = df1.loc[df1['rating'] == 'R', 'negative_confidence'].mean()
data = {'rating': ['T', 'N', 'R'], 'negative_confidence': [t_neg, n_neg, r_neg]}
# Create DataFrame.
dfp = pd.DataFrame(data)
# Print the output.
print(dfp)
# -
df2['negative_confidence'] = dfp['negative_confidence']
df2
sns.barplot(x ='rating', y ='positive_confidence', data = df2, hue ='rating')
plt.legend([],[], frameon=False)
sns.barplot(x ='rating', y ='neutral_confidence', data = df2, hue ='rating')
plt.legend([],[], frameon=False)
sns.barplot(x ='rating', y ='negative_confidence', data = df2, hue ='rating')
plt.legend([],[], frameon=False)
# ## Sentiment Confidence Score
# +
fig, axes = plt.subplots(1, 3, figsize=(15, 5), sharey=True)
fig.suptitle('Average Confidence Scorces by Rating', size=20)
# Bulbasaur
sns.barplot(ax=axes[0], x=df2.rating, y=df2.positive_confidence)
axes[0].set_title("Average positive_confidence")
# Charmander
sns.barplot(ax=axes[1], x=df2.rating, y=df2.neutral_confidence)
axes[1].set_title("Average neutral_confidence")
# Squirtle
sns.barplot(ax=axes[2], x=df2.rating, y=df2.negative_confidence)
axes[2].set_title("Average negative_confidence")
# -
|
Code/Visualization Dashboard/Python_EDA_sentiment_confidence_score.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="rNdWfPXCjTjY"
# # Improving Data Quality
#
# **Learning Objectives**
#
#
# 1. Resolve missing values
# 2. Convert the Date feature column to a datetime format
# 3. Rename a feature column, remove a value from a feature column
# 4. Create one-hot encoding features
# 5. Understand temporal feature conversions
#
#
# ## Introduction
#
# Recall that machine learning models can only consume numeric data, and that numeric data should be "1"s or "0"s. Data is said to be "messy" or "untidy" if it is missing attribute values, contains noise or outliers, has duplicates, wrong data, upper/lower case column names, and is essentially not ready for ingestion by a machine learning algorithm.
#
# This notebook presents and solves some of the most common issues of "untidy" data. Note that different problems will require different methods, and they are beyond the scope of this notebook.
#
# Each learning objective will correspond to a _#TODO_ in this student lab notebook -- try to complete this notebook first and then review the [solution notebook](https://github.com/GoogleCloudPlatform/training-data-analyst/blob/master/courses/machine_learning/deepdive2/launching_into_ml/solutions/improve_data_quality.ipynb).
# -
# !sudo chown -R jupyter:jupyter /home/jupyter/training-data-analyst
# Ensure the right version of Tensorflow is installed.
# !pip freeze | grep tensorflow==2.1 || pip install tensorflow==2.1
# Please ignore any incompatibility warnings and errors and re-run the cell to view the installed tensorflow version.
#
# + [markdown] colab_type="text" id="VxyBFc_kKazA"
# Start by importing the necessary libraries for this lab.
# -
# ### Import Libraries
import os
import pandas as pd # First, we'll import Pandas, a data processing and CSV file I/O library
import numpy as np
from datetime import datetime
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# ### Load the Dataset
# The dataset is based on California's [Vehicle Fuel Type Count by Zip Code](https://data.ca.gov/dataset/vehicle-fuel-type-count-by-zip-codeSynthetic) report. The dataset has been modified to make the data "untidy" and is thus a synthetic representation that can be used for learning purposes.
#
# Let's download the raw .csv data by copying the data from a cloud storage bucket.
#
if not os.path.isdir("../data/transport"):
os.makedirs("../data/transport")
# !gsutil cp gs://cloud-training-demos/feat_eng/transport/untidy_vehicle_data.csv ../data/transport
# !ls -l ../data/transport
# ### Read Dataset into a Pandas DataFrame
# + [markdown] colab_type="text" id="lM6-n6xntv3t"
# Next, let's read in the dataset just copied from the cloud storage bucket and create a Pandas DataFrame. We also add a Pandas .head() function to show you the top 5 rows of data in the DataFrame. Head() and Tail() are "best-practice" functions used to investigate datasets.
# + colab={"base_uri": "https://localhost:8080/", "height": 222} colab_type="code" id="REZ57BXCLdfG" outputId="a6ef2eda-c7eb-4e2d-92e4-e7fcaa20b0af"
df_transport = pd.read_csv('../data/transport/untidy_vehicle_data.csv')
df_transport.head() # Output the first five rows.
# -
# ### DataFrame Column Data Types
#
# DataFrames may have heterogenous or "mixed" data types, that is, some columns are numbers, some are strings, and some are dates etc. Because CSV files do not contain information on what data types are contained in each column, Pandas infers the data types when loading the data, e.g. if a column contains only numbers, Pandas will set that column’s data type to numeric: integer or float.
#
# Run the next cell to see information on the DataFrame.
df_transport.info()
# From what the .info() function shows us, we have six string objects and one float object. Let's print out the first and last five rows of each column. We can definitely see more of the "string" object values now!
print(df_transport)
# ### Summary Statistics
#
# At this point, we have only one column which contains a numerical value (e.g. Vehicles). For features which contain numerical values, we are often interested in various statistical measures relating to those values. We can use .describe() to see some summary statistics for the numeric fields in our dataframe. Note, that because we only have one numeric feature, we see only one summary stastic - for now.
df_transport.describe()
# Let's investigate a bit more of our data by using the .groupby() function.
df_transport.groupby('Fuel').first() # Get the first entry for each month.
# #### Checking for Missing Values
#
# Missing values adversely impact data quality, as they can lead the machine learning model to make inaccurate inferences about the data. Missing values can be the result of numerous factors, e.g. "bits" lost during streaming transmission, data entry, or perhaps a user forgot to fill in a field. Note that Pandas recognizes both empty cells and “NaN” types as missing values.
# Let's show the null values for all features in the DataFrame.
df_transport.isnull().sum()
# To see a sampling of which values are missing, enter the feature column name. You'll notice that "False" and "True" correpond to the presence or abscence of a value by index number.
print (df_transport['Date'])
print (df_transport['Date'].isnull())
print (df_transport['Make'])
print (df_transport['Make'].isnull())
print (df_transport['Model Year'])
print (df_transport['Model Year'].isnull())
# ### What can we deduce about the data at this point?
#
# First, let's summarize our data by row, column, features, unique, and missing values,
print ("Rows : " ,df_transport.shape[0])
print ("Columns : " ,df_transport.shape[1])
print ("\nFeatures : \n" ,df_transport.columns.tolist())
print ("\nUnique values : \n",df_transport.nunique())
print ("\nMissing values : ", df_transport.isnull().sum().values.sum())
# Let's see the data again -- this time the last five rows in the dataset.
df_transport.tail()
# ### What Are Our Data Quality Issues?
#
# 1. **Data Quality Issue #1**:
# > **Missing Values**:
# Each feature column has multiple missing values. In fact, we have a total of 18 missing values.
# 2. **Data Quality Issue #2**:
# > **Date DataType**: Date is shown as an "object" datatype and should be a datetime. In addition, Date is in one column. Our business requirement is to see the Date parsed out to year, month, and day.
# 3. **Data Quality Issue #3**:
# > **Model Year**: We are only interested in years greater than 2006, not "<2006".
# 4. **Data Quality Issue #4**:
# > **Categorical Columns**: The feature column "Light_Duty" is categorical and has a "Yes/No" choice. We cannot feed values like this into a machine learning model. In addition, we need to "one-hot encode the remaining "string"/"object" columns.
# 5. **Data Quality Issue #5**:
# > **Temporal Features**: How do we handle year, month, and day?
#
# #### Data Quality Issue #1:
# ##### Resolving Missing Values
# Most algorithms do not accept missing values. Yet, when we see missing values in our dataset, there is always a tendency to just "drop all the rows" with missing values. Although Pandas will fill in the blank space with “NaN", we should "handle" them in some way.
#
# While all the methods to handle missing values is beyond the scope of this lab, there are a few methods you should consider. For numeric columns, use the "mean" values to fill in the missing numeric values. For categorical columns, use the "mode" (or most frequent values) to fill in missing categorical values.
#
# In this lab, we use the .apply and Lambda functions to fill every column with its own most frequent value. You'll learn more about Lambda functions later in the lab.
# Let's check again for missing values by showing how many rows contain NaN values for each feature column.
# **Lab Task #1a:** Check for missing values by showing how many rows contain NaN values for each feature column.
# TODO 1a
# TODO -- Your code here.
# **Lab Task #1b:** Apply the lambda function.
# TODO 1b
# TODO -- Your code here.
# **Lab Task #1c:** Check again for missing values.
# TODO 1c
# TODO -- Your code here.
# #### Data Quality Issue #2:
# ##### Convert the Date Feature Column to a Datetime Format
# The date column is indeed shown as a string object.
# **Lab Task #2a:** Convert the datetime datatype with the to_datetime() function in Pandas.
# TODO 2a
# TODO -- Your code here.
# **Lab Task #2b:** Show the converted Date.
# TODO 2b
# TODO -- Your code here.
# Let's parse Date into three columns, e.g. year, month, and day.
df_transport['year'] = df_transport['Date'].dt.year
df_transport['month'] = df_transport['Date'].dt.month
df_transport['day'] = df_transport['Date'].dt.day
#df['hour'] = df['date'].dt.hour - you could use this if your date format included hour.
#df['minute'] = df['date'].dt.minute - you could use this if your date format included minute.
df_transport.info()
# Next, let's confirm the Date parsing. This will also give us a another visualization of the data.
# +
# Here, we are creating a new dataframe called "grouped_data" and grouping by on the column "Make"
grouped_data = df_transport.groupby(['Make'])
# Get the first entry for each month.
df_transport.groupby('month').first()
# -
# Now that we have Dates as a integers, let's do some additional plotting.
plt.figure(figsize=(10,6))
sns.jointplot(x='month',y='Vehicles',data=df_transport)
#plt.title('Vehicles by Month')
# #### Data Quality Issue #3:
# ##### Rename a Feature Column and Remove a Value.
#
# Our feature columns have different "capitalizations" in their names, e.g. both upper and lower "case". In addition, there are "spaces" in some of the column names. In addition, we are only interested in years greater than 2006, not "<2006".
# **Lab Task #3a:** Remove all the spaces for feature columns by renaming them.
# TODO 3a
# TODO -- Your code here.
# **Note:** Next we create a copy of the dataframe to avoid the "SettingWithCopyWarning: A value is trying to be set on a copy of a slice from a DataFrame" warning. Run the cell to remove the value '<2006' from the modelyear feature column.
# **Lab Task #3b:** Create a copy of the dataframe to avoid copy warning issues.
# TODO 3b
# TODO -- Your code here.
# Next, confirm that the modelyear value '<2006' has been removed by doing a value count.
df['modelyear'].value_counts(0)
# #### Data Quality Issue #4:
# ##### Handling Categorical Columns
#
# The feature column "lightduty" is categorical and has a "Yes/No" choice. We cannot feed values like this into a machine learning model. We need to convert the binary answers from strings of yes/no to integers of 1/0. There are various methods to achieve this. We will use the "apply" method with a lambda expression. Pandas. apply() takes a function and applies it to all values of a Pandas series.
#
# ##### What is a Lambda Function?
#
# Typically, Python requires that you define a function using the def keyword. However, lambda functions are anonymous -- which means there is no need to name them. The most common use case for lambda functions is in code that requires a simple one-line function (e.g. lambdas only have a single expression).
#
# As you progress through the Course Specialization, you will see many examples where lambda functions are being used. Now is a good time to become familiar with them.
# First, lets count the number of "Yes" and"No's" in the 'lightduty' feature column.
df['lightduty'].value_counts(0)
# Let's convert the Yes to 1 and No to 0. Pandas. apply() . apply takes a function and applies it to all values of a Pandas series (e.g. lightduty).
df.loc[:,'lightduty'] = df['lightduty'].apply(lambda x: 0 if x=='No' else 1)
df['lightduty'].value_counts(0)
# +
# Confirm that "lightduty" has been converted.
df.head()
# -
# #### One-Hot Encoding Categorical Feature Columns
#
# Machine learning algorithms expect input vectors and not categorical features. Specifically, they cannot handle text or string values. Thus, it is often useful to transform categorical features into vectors.
#
# One transformation method is to create dummy variables for our categorical features. Dummy variables are a set of binary (0 or 1) variables that each represent a single class from a categorical feature. We simply encode the categorical variable as a one-hot vector, i.e. a vector where only one element is non-zero, or hot. With one-hot encoding, a categorical feature becomes an array whose size is the number of possible choices for that feature.
# Panda provides a function called "get_dummies" to convert a categorical variable into dummy/indicator variables.
# +
# Making dummy variables for categorical data with more inputs.
data_dummy = pd.get_dummies(df[['zipcode','modelyear', 'fuel', 'make']], drop_first=True)
data_dummy.head()
# -
# **Lab Task #4a:** Merge (concatenate) original data frame with 'dummy' dataframe.
# TODO 4a
# TODO -- Your code here.
# **Lab Task #4b:** Drop attributes for which we made dummy variables.
# TODO 4b
# TODO -- Your code here.
# +
# Confirm that 'zipcode','modelyear', 'fuel', and 'make' have been dropped.
df.head()
# -
# #### Data Quality Issue #5:
# ##### Temporal Feature Columns
#
# Our dataset now contains year, month, and day feature columns. Let's convert the month and day feature columns to meaningful representations as a way to get us thinking about changing temporal features -- as they are sometimes overlooked.
#
# Note that the Feature Engineering course in this Specialization will provide more depth on methods to handle year, month, day, and hour feature columns.
#
# First, let's print the unique values for "month" and "day" in our dataset.
print ('Unique values of month:',df.month.unique())
print ('Unique values of day:',df.day.unique())
print ('Unique values of year:',df.year.unique())
# Next, we map each temporal variable onto a circle such that the lowest value for that variable appears right next to the largest value. We compute the x- and y- component of that point using sin and cos trigonometric functions. Don't worry, this is the last time we will use this code, as you can develop an input pipeline to address these temporal feature columns in TensorFlow and Keras - and it is much easier! But, sometimes you need to appreciate what you're not going to encounter as you move through the course!
#
# Run the cell to view the output.
# **Lab Task #5:** Drop month, and day
# +
df['day_sin'] = np.sin(df.day*(2.*np.pi/31))
df['day_cos'] = np.cos(df.day*(2.*np.pi/31))
df['month_sin'] = np.sin((df.month-1)*(2.*np.pi/12))
df['month_cos'] = np.cos((df.month-1)*(2.*np.pi/12))
# TODO 5
# TODO -- Your code here.
# -
# scroll left to see the converted month and day coluumns.
df.tail(4)
# ### Conclusion
#
# This notebook introduced a few concepts to improve data quality. We resolved missing values, converted the Date feature column to a datetime format, renamed feature columns, removed a value from a feature column, created one-hot encoding features, and converted temporal features to meaningful representations. By the end of our lab, we gained an understanding as to why data should be "cleaned" and "pre-processed" before input into a machine learning model.
# Copyright 2020 Google Inc.
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
|
courses/machine_learning/deepdive2/launching_into_ml/labs/improve_data_quality.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W1D4_GeneralizedLinearModels/W1D4_Outro.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# -
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W1D4_GeneralizedLinearModels/W1D4_Outro.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] pycharm={"name": "#%% md\n"}
# # Outro
#
# -
# **Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
#
# <p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Video
# + cellView="form" pycharm={"name": "#%%\n"}
# @markdown
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1d5411Y7mN", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"NXVG9ORBYXQ", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# ## Daily survey
#
# Don't forget to complete your reflections and content check in the daily survey! Please be patient after logging in as there is
# a small delay before you will be redirected to the survey.
#
# <a href="https://portal.neuromatchacademy.org/api/redirect/to/245cbea5-780c-443e-a870-469a423bbe58"><img src="https://github.com/NeuromatchAcademy/course-content/blob/master/tutorials/static/button.png?raw=1" alt="button link to survey" style="width:410px"></a>
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Slides
# + cellView="form" pycharm={"name": "#%%\n"}
# @markdown
from IPython.display import IFrame
IFrame(src=f"https://mfr.ca-1.osf.io/render?url=https://osf.io/pzw7s/?direct%26mode=render%26action=download%26mode=render", width=854, height=480)
|
tutorials/W1D4_GeneralizedLinearModels/W1D4_Outro.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Running a Regression in Python
# *Suggested Answers follow (usually there are multiple ways to solve a problem in Python).*
# *A teacher at school decided her students should take an IQ test. She prepared 5 tests she believed were aligned with the requirements of the IQ examination.
# The father of one child in the class turned out to be an econometrician, so he asked her for the results of the 30 kids. The file contained the points they earned on each test and the final IQ score.*
# Load the IQ_data excel file.
# Prepare the data for a univariate regression of Test 1 based on the IQ result. Store the Test 1 scores in a variable, called X, and the IQ points in another variable, named Y.
# ### Univariate Regression
# Create a well-organized scatter plot. Use the “axis” method with the following start and end points: [0, 120, 0, 150]. Label the axes “Test 1” and “IQ”, respectively.
# Just by looking at the graph, do you believe Test 1 is a good predictor of the final IQ score?
|
23 - Python for Finance/4_Using Regressions for Financial Analysis/2_Running a Regression in Python (6:35)/Running a Regression in Python - Exercise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# paddle模式词性和专名类别标签集合如下表,其中词性标签 24 个(小写字母),专名类别标签 4 个(大写字母)。
#
# 标签 含义 标签 含义 标签 含义 标签 含义
#
# n 普通名词 f 方位名词 s 处所名词 t 时间
#
# nr 人名 ns 地名 nt 机构名 nw 作品名
#
# nz 其他专名 v 普通动词 vd 动副词 vn 名动词
#
# a 形容词 ad 副形词 an 名形词 d 副词
#
# m 数量词 q 量词 r 代词 p 介词
#
# c 连词 u 助词 xc 其他虚词 w 标点符号
#
# PER 人名 LOC 地名 ORG 机构名 TIME 时间
#
# +
## Demo
import jieba.posseg as pseg
words = pseg.cut("我爱北京天安门")
for (word,flag) in words:
print(word, flag)
# +
import json
import jieba.posseg as pseg
with open('../assets/hsk-level-1.json') as file:
hsk1_data = json.load(file)
hsk1_words = []
for item in hsk1_data:
hsk1_words.append(item['hanzi'])
with open('../assets/hsk-level-2.json') as file:
hsk2_data = json.load(file)
hsk2_words = []
for item in hsk2_data:
hsk2_words.append(item['hanzi'])
with open('../assets/hsk-level-3.json') as file:
hsk3_data = json.load(file)
hsk3_words = []
for item in hsk3_data:
hsk3_words.append(item['hanzi'])
# -
hsk1_data[0]
# +
for item in hsk1_data:
for word,flag in pseg.cut(item['hanzi']):
print(word,flag, sep=',')
item['attribute']=flag
# print('',item['id'] , item['pinyin'],item['translations'] , sep=',')
print("---------------,-")
for item in hsk2_data:
for word,flag in pseg.cut(item['hanzi']):
print(word, ",",flag, sep='')
item['attribute']=flag
print("---------------,-")
for item in hsk3_data:
for word,flag in pseg.cut(item['hanzi']):
print(word, ",",flag, sep='')
item['attribute']=flag
# -
hsk1_data[0]
# json.dumps(hsk1_data)
with open('tmp.json', 'w') as json_file:
json.dump(hsk1_data + hsk2_data + hsk3_data, json_file, ensure_ascii=False)
# +
# json.dumps(hsk2_data, ensure_ascii=False)
# -
|
others/get_attribute.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Author : <NAME>
# github link : https://github.com/amirshnll/Speaker-Accent-Recognition
# dataset link : http://archive.ics.uci.edu/ml/datasets/Speaker+Accent+Recognition
# email : <EMAIL>
# +
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import classification_report, confusion_matrix
df = pd.read_csv('dataset.csv')
x = df.iloc[:, 1:] # features data
y = df.iloc[:, :1] # class data
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.20) # split train data and validation (test) data with test data including 20 percent of whole data
# ========== (Begin) This block of code perform normalizing or scaling the features which is a good practice so that all of features can be uniformly evaluated
# scaler = StandardScaler()
# scaler.fit(x_train)
# x_train = scaler.transform(x_train)
# x_test = scaler.transform(x_test)
# ========== (End) Feature scaling
classifier = KNeighborsClassifier(n_neighbors=6)
classifier.fit(x_train, y_train)
y_pred = classifier.predict(x_test)
print('========== Test Features ============')
print(x_test)
print('============ Predicted Values ===========')
print(y_pred)
# ============ Evaluating the algorithm
print(confusion_matrix(y_test, y_pred))
print(classification_report(y_test, y_pred))
print('\n\n============================ Comparing Error rate with different k value ============================= \n\n')
error = []
# Calculating error for K values between 1 and 10
for i in range(1, 10):
knn = KNeighborsClassifier(n_neighbors=i)
knn.fit(x_train, y_train)
pred_i = knn.predict(x_test)
pred_i=pred_i.reshape(len(x_test),1)
error.append(np.mean(pred_i != y_test))
plt.figure(figsize=(12, 6))
plt.plot(range(1, 10), error, color='red', linestyle='dashed', marker='o',
markerfacecolor='blue', markersize=10)
plt.title('Error Rate K Value')
plt.xlabel('K Value')
plt.ylabel('Mean Error')
# -
|
knn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from __future__ import print_function
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from scipy import stats
# +
from sklearn import metrics
from sklearn.metrics import classification_report
from sklearn import preprocessing
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten, Reshape, GlobalAveragePooling1D
from keras.layers import Conv2D, MaxPooling2D, Conv1D, MaxPooling1D
from keras.utils import np_utils
# -
def feature_normalize(dataset):
mu = np.mean(dataset, axis=0)
sigma = np.std(dataset, axis=0)
return (dataset - mu)/sigma
def show_basic_dataframe_info(dataframe,
preview_rows=20):
"""
This function shows basic information for the given dataframe
Args:
dataframe: A Pandas DataFrame expected to contain data
preview_rows: An integer value of how many rows to preview
Returns:
Nothing
"""
# Shape and how many rows and columns
print("Number of columns in the dataframe: %i" % (dataframe.shape[1]))
print("Number of rows in the dataframe: %i\n" % (dataframe.shape[0]))
print("First 20 rows of the dataframe:\n")
# Show first 20 rows
print(dataframe.head(preview_rows))
print("\nDescription of dataframe:\n")
# Describe dataset like mean, min, max, etc.
# print(dataframe.describe())
def read_data(file_path):
"""
This function reads the accelerometer data from a file
Args:
file_path: URL pointing to the CSV file
Returns:
A pandas dataframe
"""
data = np.load(file_path)
column_names = ['x-axis',
'y-axis',
'labels']
df = pd.DataFrame(data, columns=column_names)
return df
def plot_axis(ax, x, y, title):
ax.plot(x, y)
ax.set_title(title)
ax.xaxis.set_visible(False)
ax.set_ylim([min(y) - np.std(y), max(y) + np.std(y)])
ax.set_xlim([min(x), max(x)])
ax.grid(True)
def create_segments_and_labels(df, time_steps, step, label_name):
"""
This function receives a dataframe and returns the reshaped segments
of x,y acceleration as well as the corresponding labels
Args:
df: Dataframe in the expected format
time_steps: Integer value of the length of a segment that is created
Returns:
reshaped_segments
labels:
"""
# x, y acceleration as features
N_FEATURES = 2
# Number of steps to advance in each iteration (for me, it should always
# be equal to the time_steps in order to have no overlap between segments)
# step = time_steps
segments = []
l = []
labels = []
for i in range(0, len(df) - time_steps, step):
xs = df['x-axis'].values[i: i + time_steps]
ys = df['y-axis'].values[i: i + time_steps]
# Retrieve the most often used label in this segment
label = stats.mode(df[label_name][i: i + time_steps])[0][0]
segments = np.dstack((xs, ys))
l.append(segments)
#print(segments.shape)
labels.append(label)
#break
# Bring the segments into a better shape
reshaped_segments = np.asarray(l, dtype= np.float32).reshape(-1, time_steps, N_FEATURES)
#print(reshaped_segments.shape)
labels = np.asarray(labels)
return reshaped_segments, labels
LABELS = ["bhujangasan",
"padamasan",
"shavasan",
"tadasan",
"trikonasan",
"vrikshasan"]
TIME_PERIODS = 25
STEP_DISTANCE = 25
df_train = read_data('data_files/combined_xylabel/data_train_xylabel.npy')
df_train.head()
df_test = read_data('data_files/combined_xylabel/data_test_xylabel.npy')
df_val = read_data('data_files/combined_xylabel/data_val_xylabel.npy')
LABEL = 'labels'
# +
df_train['x-axis'] = feature_normalize(df_train['x-axis'])
df_test['x-axis'] = feature_normalize(df_test['x-axis'])
df_val['x-axis'] = feature_normalize(df_val['x-axis'])
df_train['y-axis'] = feature_normalize(df_train['y-axis'])
df_test['y-axis'] = feature_normalize(df_test['y-axis'])
df_val['y-axis'] = feature_normalize(df_val['y-axis'])
# -
X_train, y_train = create_segments_and_labels(df_train,
TIME_PERIODS,
STEP_DISTANCE,
LABEL)
X_train.shape
X_test, y_test = create_segments_and_labels(df_test,
TIME_PERIODS,
STEP_DISTANCE,
LABEL)
X_test.shape
y_test.shape
X_val, y_val = create_segments_and_labels(df_val,
TIME_PERIODS,
STEP_DISTANCE,
LABEL)
num_time_periods, num_sensors = X_train.shape[1], X_train.shape[2]
num_classes = 6
y_train.shape
input_shape = (num_time_periods*num_sensors)
X_train = X_train.reshape(X_train.shape[0], input_shape)
X_train = X_train.reshape(X_train.shape[0], input_shape)
print('X_train shape:', X_train.shape)
X_train = X_train.astype("float32")
y_train = y_train.astype("float32")
y_train = np_utils.to_categorical(y_train, num_classes)
print('New y_train shape: ', y_train.shape)
print("\n--- Create neural network model ---\n")
# +
model_m = Sequential()
model_m.add(Reshape((TIME_PERIODS, num_sensors), input_shape=(input_shape,)))
print('here1')
model_m.add(Conv1D(100, 10, activation='relu', input_shape=(TIME_PERIODS, num_sensors)))
print('here2')
model_m.add(Conv1D(100, 10, activation='relu'))
print('here3')
model_m.add(MaxPooling1D(3))
print('here4')
model_m.add(Conv1D(160, 10, activation='relu', padding="same"))
print('here5')
model_m.add(Conv1D(160, 10, activation='relu', padding="same"))
print('here6')
model_m.add(GlobalAveragePooling1D())
model_m.add(Dropout(0.5))
model_m.add(Dense(num_classes, activation='softmax'))
print(model_m.summary())
# -
print("\n--- Fit the model ---\n")
callbacks_list = [
keras.callbacks.ModelCheckpoint(
filepath='best_model.{epoch:02d}-{val_acc:.2f}.h5',
monitor='val_loss', save_best_only=True),
#keras.callbacks.EarlyStopping(monitor='acc', patience=1)
]
model_m.compile(loss='categorical_crossentropy',
optimizer='adam', metrics=['accuracy'])
BATCH_SIZE = 400
EPOCHS = 50
# +
X_val = X_val.reshape(X_val.shape[0], input_shape)
X_val = X_val.astype("float32")
y_val = y_val.astype("float32")
y_val = np_utils.to_categorical(y_val, num_classes)
print('New y_val shape: ', y_val.shape)
# -
history = model_m.fit(X_train,
y_train,
batch_size=BATCH_SIZE,
epochs=EPOCHS,
callbacks=callbacks_list,
validation_data=(X_val, y_val),
verbose=1)
print("\n--- Learning curve of model training ---\n")
plt.figure(figsize=(6, 4))
plt.plot(history.history['acc'], "g--", label="Accuracy of training data")
plt.plot(history.history['val_acc'], "g", label="Accuracy of validation data")
#plt.plot(history.history['loss'], "r--", label="Loss of training data")
#plt.plot(history.history['val_loss'], "r", label="Loss of validation data")
plt.title('Model Accuracy and Loss')
plt.ylabel('Accuracy and Loss')
plt.xlabel('Training Epoch')
plt.ylim(0)
plt.legend()
plt.show()
model = model_m
#load best weights from current training
model.load_weights("best_model.04-0.93.h5")
X_test = X_test.reshape(X_test.shape[0], input_shape)
X_test = X_test.astype("float32")
y_test = y_test.astype("float32")
y_test = np_utils.to_categorical(y_test, num_classes)
y_test.shape
score = model.evaluate(X_test, y_test, verbose=1)
print("\nAccuracy on test data: %0.2f" % score[1])
print("\nLoss on test data: %0.2f" % score[0])
|
CNN-LSTM-model/models/25-joint-data/Model_Preparation_FrameWise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/danielsoy/ALOCC-CVPR2018/blob/master/train%20github.com%20ktr-hubrt%20WSAL.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="46PZumgYhCHU"
from __future__ import print_function
from __future__ import division
from torch.utils.data import DataLoader
from torch import optim
from torch.autograd import Variable
from torch.nn.functional import softmax
from torch.nn.functional import sigmoid
from torch.nn import MSELoss
from torch.nn import L1Loss
from torch.nn import SmoothL1Loss
import torch
import numpy as np
from tqdm import tqdm
from sklearn.metrics import roc_auc_score
from matplotlib import pyplot as plt
from models import HOE_model
from dataset import dataset_h5
from utils import *
import pdb
import os
import time
from torch import nn
# + id="rPT4kd70hCHX"
def train_wsal():
torch.cuda.empty_cache()
ver ='WSAL'
videos_pkl_train = "/test/UCF-Crime/UCF/Anomaly_Detection_splits/Anomaly_Train.txt"
hdf5_path = "/test/UCF-Crime/UCF/gcn_feas.hdf5"
mask_path = "/test/UCF-Crime/UCF/gcn_mask.hdf5"
# + id="bfb4bMFhhCHY"
modality = "rgb"
gpu_id = 0
batch_size = 30
iter_size = 30//batch_size
random_crop = False
train_loader = torch.utils.data.DataLoader(dataset_h5(videos_pkl_train, hdf5_path, mask_path),
batch_size=batch_size, shuffle=True, num_workers=0, pin_memory=True, drop_last=True)
model = HOE_model(nfeat=1024, nclass=1)
criterion = torch.nn.CrossEntropyLoss(reduction = 'none')
Rcriterion = torch.nn.MarginRankingLoss(margin=1.0, reduction = 'mean')
if gpu_id != -1:
model = model.cuda(gpu_id)
criterion = criterion.cuda(gpu_id)
Rcriterion = Rcriterion.cuda(gpu_id)
optimizer = optim.Adagrad(model.parameters(), lr=0.001)
opt_scheduler = optim.lr_scheduler.MultiStepLR(optimizer, milestones=[300,400], gamma=0.5)
start_epoch = 0
resume = './weights/WSAL_1.1/rgb_300.pth'
if os.path.isfile(resume):
print("=> loading checkpoint '{}'".format(resume))
checkpoint = torch.load(resume)
start_epoch = checkpoint['epoch']
opt_scheduler.load_state_dict(checkpoint['scheduler'])
model.load_state_dict(checkpoint['state_dict'])
optimizer.load_state_dict(checkpoint['optimizer'])
iter_count = 0
alpha = 0.5
vid2mean_pred = {}
losses = AverageMeter()
data_time = AverageMeter()
model.train()
for epoch in range(start_epoch, 500):
end = time.time()
pbar = tqdm(total=len(train_loader))
for step, data in enumerate(train_loader):
data_time.update(time.time() - end)
an_feats, no_feats, preds = data
an_feat, no_feat, pred = Variable(an_feats), Variable(no_feats), Variable(preds)
if gpu_id != -1:
an_feat = an_feat.cuda(gpu_id)
no_feat = no_feat.cuda(gpu_id)
pred = pred.cuda(gpu_id).float()
if iter_count % iter_size == 0:
optimizer.zero_grad()
ano_ss, ano_fea = model(an_feat)
nor_ss, nor_fea = model(no_feat)
ano_cos = torch.cosine_similarity(ano_fea[:,1:], ano_fea[:,:-1], dim=2)
dynamic_score_ano = 1-ano_cos
nor_cos = torch.cosine_similarity(nor_fea[:,1:], nor_fea[:,:-1], dim=2)
dynamic_score_nor = 1-nor_cos
ano_max = torch.max(dynamic_score_ano,1)[0]
nor_max = torch.max(dynamic_score_nor,1)[0]
loss_dy = Rcriterion(ano_max, nor_max, pred[:,0])
semantic_margin_ano = torch.max(ano_ss,1)[0]-torch.min(ano_ss,1)[0]
semantic_margin_nor = torch.max(nor_ss,1)[0]-torch.min(nor_ss,1)[0]
loss_se = Rcriterion(semantic_margin_ano, semantic_margin_nor, pred[:,0])
loss_3 = torch.mean(torch.sum(dynamic_score_ano,1))+torch.mean(torch.sum(dynamic_score_nor,1))+torch.mean(torch.sum(ano_ss,1))+torch.mean(torch.sum(nor_ss,1))
loss_5 = torch.mean(torch.sum((dynamic_score_ano[:,:-1]-dynamic_score_ano[:,1:])**2,1))+torch.mean(torch.sum((ano_ss[:,:-1]-ano_ss[:,1:])**2,1))
loss_train = loss_se + loss_dy+ loss_3*0.00008+ loss_5*0.00008
iter_count += 1
loss_train.backward()
losses.update(loss_train.item(), 1)
if (iter_count + 1) % iter_size == 0:
optimizer.step()
pbar.set_postfix({
'Data': '{data_time.val:.3f}({data_time.avg:.4f})\t'.format(data_time=data_time),
ver: '{0}'.format(epoch),
'lr': '{lr:.5f}\t'.format(lr=optimizer.param_groups[-1]['lr']),
'Loss': '{loss.val:.4f}({loss.avg:.4f})\t '.format(loss=losses)
})
pbar.update(1)
pbar.close()
model_path = 'weights/'+ver+'/'
if not os.path.isdir(model_path):
os.mkdir(model_path)
if epoch%50==0:
state = {
'epoch': epoch,
'state_dict': model.state_dict(),
'optimizer' : optimizer.state_dict(),
'scheduler': opt_scheduler.state_dict(),
}
torch.save(state, model_path+"rgb_%d.pth" % epoch)
# model = model.cuda(gpu_id)
# if epoch%25==0:
losses.reset()
opt_scheduler.step()
|
train github.com ktr-hubrt WSAL.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import numpy as np
import torch
import os
import pickle
import matplotlib.pyplot as plt
# # %matplotlib inline
# plt.rcParams['figure.figsize'] = (20, 20)
# plt.rcParams['image.interpolation'] = 'bilinear'
from argparse import ArgumentParser
from torch.optim import SGD, Adam
from torch.autograd import Variable
from torch.utils.data import DataLoader, Dataset
from torchvision.transforms import Normalize
from torchvision.transforms import ToTensor, ToPILImage
import torchvision
import torchvision.transforms as T
import torch.nn.functional as F
import torch.nn as nn
from torch.optim import lr_scheduler
from networks.SegUNet import SegUNet
import collections
import numbers
import random
import math
from PIL import Image, ImageOps, ImageEnhance
import logging
import time
import tool
# import bcolz
# -
CROSS_VALIDATION_FOLD = 0 # 0-4
SEED = CROSS_VALIDATION_FOLD * 100
NUM_CHANNELS = 3
NUM_CLASSES = 2
model_name = 'SegUNet'
log_path = 'log/'
save_weights_path = '../_weights/'
if not os.path.exists(save_weights_path):
os.makedirs(save_weights_path)
if not os.path.exists(log_path):
os.makedirs(log_path)
log_filename = log_path + model_name + '-fold'+str(CROSS_VALIDATION_FOLD)+'.log'
logging.basicConfig(filename=log_filename, level=logging.INFO,
format='%(asctime)s:%(levelname)s:%(message)s')
def log(message):
print(message)
logging.info(message)
log('='*50 + 'start run' + '='*50)
# ## dataset
# +
# NUM_CHANNELS = 3
# NUM_CLASSES = 2 # car is 1, background is 0
# color_transform = Colorize(n=NUM_CLASSES)
# image_transform = ToPILImage()
# -
class Crop_different_size_for_image_and_label(object):
def __init__(self, image_size=572, label_size=388):
self.image_size = image_size
self.label_size = label_size
self.bound = (self.image_size - self.label_size) // 2
def __call__(self, img_and_label):
img, label = img_and_label
w, h = img.size
xcenter = 1000
ycenter = 600
img = img.crop((xcenter - self.image_size // 2, ycenter - self.image_size // 2, xcenter + self.image_size // 2, ycenter + self.image_size // 2))
label = label.crop((xcenter - self.label_size // 2, ycenter - self.label_size // 2, xcenter + self.label_size // 2, ycenter + self.label_size // 2))
return img, label
# random_rotate = tool.Random_Rotate_Crop(maxAngle = 10)
# crop = tool.Random_Rotate_Crop(maxAngle = 0)
crop_512 = tool.RandomCrop(crop_size = 512)
# crop_572 = tool.RandomCrop_different_size_for_image_and_label(image_size=572, label_size=388)
# random_color = tool.RandomColor()
to_tensor_label = tool.ToTensor_Label()
normalize = tool.ImageNormalize([.485, .456, .406], [.229, .224, .225])
train_transforms = tool.Compose([crop_512, to_tensor_label, normalize])
val_transforms = tool.Compose([crop_512, to_tensor_label, normalize])
image_path = '../../data/images/train/'
mask_path = '../../data/images/train_masks/'
with open('./train_shuffle_names.pk', 'rb') as f:
filenames = pickle.load(f)
# +
fold_num = len(filenames) // 5
folds = []
for i in range(5):
if i == 4:
folds.append(filenames[i * fold_num :])
else:
folds.append(filenames[i * fold_num : (i + 1) * fold_num])
train_filenames = []
for i in range(5):
if i == CROSS_VALIDATION_FOLD:
val_filenames = folds[i]
else:
train_filenames += folds[i]
# +
# train_filenames = train_filenames[:4]
# val_filenames = val_filenames[:1]
# -
train_set = tool.Car_dataset(image_path, mask_path, train_filenames, train_transforms, ifFlip=True)
val_set = tool.Car_dataset(image_path, mask_path, val_filenames, val_transforms, ifFlip=True) # for validation set
train_loader = DataLoader(train_set, num_workers=4, batch_size=4, shuffle=True)
val_loader = DataLoader(val_set, num_workers=4, batch_size=1) # for validation set
# ## dataset examples
# +
# inp, tar = train_loader.__iter__().next()
# +
# i = 0
# inp = Variable(inp)
# tar = Variable(tar)
# # tar[:, 0]
# t = tar[i].cpu().data.numpy()
# inpu = inp[i].cpu().data.numpy()
# +
# img_tensor = inp[i]
# for ten, m, s in zip(img_tensor, [.229, .224, .225], [.485, .456, .406]):
# ten.mul_(m).add_(s)
# img = ToPILImage()(img_tensor)
# +
# img
# +
# b = (572-388)//2
# +
# img = img.crop((b, b, b + 388, b + 388))
# +
# img
# +
# label = Image.fromarray(t[0].astype('uint8') * 255)
# label
# -
# ## train
# +
def load_model(filename, model, optimizer):
checkpoint = torch.load(filename)
model.load_state_dict(checkpoint['model_state'])
optimizer.load_state_dict(checkpoint['optimizer_state'])
def save_model(filename, model, optimizer):
torch.save({'model_state': model.state_dict(),
'optimizer_state': optimizer.state_dict()},
filename)
# -
class CrossEntropyLoss2d(nn.Module):
def __init__(self, weight=None, size_average=True):
super(CrossEntropyLoss2d, self).__init__()
self.loss = nn.NLLLoss(weight, size_average)
def forward(self, outputs, targets):
return self.loss(F.log_softmax(outputs, dim=1), targets)
def train(epoch, steps_plot=0):
model.train()
weight = torch.ones(NUM_CLASSES)
criterion = CrossEntropyLoss2d(weight.cuda()) # loss function
epoch_loss = []
step_loss = []
for step, (images, labels) in enumerate(train_loader):
images = images.cuda()
labels = labels.cuda()
inputs = Variable(images)
targets = Variable(labels)
outputs = model(inputs)
optimizer.zero_grad()
loss = criterion(outputs, targets[:, 0])
loss.backward()
optimizer.step()
epoch_loss.append(loss.item())
step_loss.append(loss.item())
if step % 10 == 0:
average_step_loss = sum(step_loss) / len(step_loss)
message = 'Epoch[{}]({}/{}): \tloss: {:.4}'.format(epoch, step, len(train_loader), average_step_loss)
log(message)
step_loss = []
average = sum(epoch_loss) / len(epoch_loss)
message = 'Train: Epoch[{}] \taverage loss: {:.4}'.format(epoch, average)
log(message)
# +
def test(steps_plot = 0):
model.eval()
weight = torch.ones(NUM_CLASSES)
criterion = CrossEntropyLoss2d(weight.cuda())
# for epoch in range(start_epoch, end_epochs+1):
total_loss = []
for step, (images, labels) in enumerate(val_loader):
images = images.cuda()
labels = labels.cuda()
inputs = Variable(images)
targets = Variable(labels)
outputs = model(inputs)
loss = criterion(outputs, targets[:, 0])
total_loss.append(loss.item())
average = sum(total_loss) / len(total_loss)
message = 'Validation: \taverage loss: {:.4}'.format(average)
log(message)
return average
# -
# ## train
#
torch.cuda.manual_seed_all(SEED)
model = SegUNet(in_channels=NUM_CHANNELS, n_classes=NUM_CLASSES)
model = model.cuda()
optimizer = Adam(model.parameters(), lr = 1e-3)
# load_model('save_model_test.pth', model, optimizer)
# +
save_weights_path = '../_weights/'
scheduler = lr_scheduler.StepLR(optimizer, step_size=40, gamma=0.5)
val_losses = []
start_time = time.time()
epoch_num = 250
for epoch in range(epoch_num):
scheduler.step(epoch)
message = 'learning rate: ' + str(scheduler.get_lr()[0])
log(message)
train(epoch)
log('-'*100)
if epoch == 0:
t1 = time.time()
message = 'one epoch time: ' + str(t1 - start_time) + 's'
log(message)
log('-'*100)
val_loss = test()
log('-'*100)
val_losses.append(val_loss)
if val_loss == min(val_losses) and epoch >= 100:
save_file_name = save_weights_path+model_name+'-fold'+str(CROSS_VALIDATION_FOLD)+'-%.5f' % val_loss+'.pth'
save_model(save_file_name, model, optimizer)
end_time = time.time()
total_time = end_time - start_time
average_time = total_time / epoch_num
message = 'total_time: ' + str(total_time) + 's' + '\t\taverage_time: ' + str(average_time) + 's'
log(message)
# -
|
train/train_SegUNet.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MULTIPLY SAR Data Access and Pre-Processing
# The purpose of this Jupyter Notebook is to show how the MULTIPLY platform can be used to retrieve S1 SLC Data from the Data Access Component and how it can be processed into S1 GRD Data using the SAR Pre-Processing functionality.
#
# First, let's define working directories.
# +
from vm_support import get_working_dir
name = 'm1'
# create and/or clear working directory
working_dir = get_working_dir(name)
print('Working directory is {}'.format(working_dir))
s1_slc_directory = '{}/s1_slc'.format(working_dir)
s1_grd_directory = '{}/s1_grd'.format(working_dir)
# -
# Now to ensure that all data stores are set up (step over this part if you have already done it). Please also set your Earth Data Authentication if you execute this.
# +
#from vm_support import set_earth_data_authentication, set_up_data_stores
#set_up_data_stores()
#username = ''
#password = ''
#set_earth_data_authentication(username, password) # to download modis data, needs only be done once
# -
# We need to define start and end times and a region of interest.
start_time_as_string = '2017-06-01'
stop_time_as_string = '2017-06-10'
roi = 'POLYGON((9.99 53.51,10.01 53.51,10.01 53.49, 9.99 53.49, 9.99 53.51))'
# For the SAR Pre-Processing we require a config file. Let's create it.
from vm_support import create_sar_config_file
create_sar_config_file(temp_dir=working_dir, roi=roi, start_time=start_time_as_string, end_time=stop_time_as_string,
s1_slc_directory=s1_slc_directory, s1_grd_directory=s1_grd_directory)
config_file = f'{working_dir}/sar_config.yaml'
# Next set up the Data Access Component.
from multiply_data_access import DataAccessComponent
dac = DataAccessComponent()
# For the SAR Pre-Processing we need to have 15 products at least: 7 before and 7 after the product in question. However, the SAR Pre-Processing does not count all products as full products: If the products are located close to a border, they are counted as half products. As the determination whether a product is counted as a full or half a product is made by the SAR Pre-Processor, we need it to determine he products that are required. To do so, it is necessary to access the products, so we might already need to download.
#
# Let's start with determining the actual start date:
# +
import datetime
import logging
import os
from multiply_orchestration import create_sym_links
from sar_pre_processing import SARPreProcessor
one_day = datetime.timedelta(days=1)
before_sar_dir = f'{s1_slc_directory}/before'
if not os.path.exists(before_sar_dir):
os.makedirs(before_sar_dir)
start = datetime.datetime.strptime(start_time_as_string, '%Y-%m-%d')
before = start
num_before = 0
while num_before < 7:
before -= one_day
before_date = datetime.datetime.strftime(before, '%Y-%m-%d')
data_urls_before = dac.get_data_urls(roi, before_date, start_time_as_string, 'S1_SLC')
create_sym_links(data_urls_before, before_sar_dir)
processor = SARPreProcessor(config=config_file, input=before_sar_dir, output=before_sar_dir)
list = processor.create_processing_file_list()
num_before = len(list[0]) + (len(list[1]) / 2.)
logging.info(f'Set start date for collecting S1 SLC products to {before_date}.')
# -
# Now the actual end date. Take care not to set it in the future.
# +
after_sar_dir = f'{s1_slc_directory}/after'
if not os.path.exists(after_sar_dir):
os.makedirs(after_sar_dir)
end = datetime.datetime.strptime(stop_time_as_string, '%Y-%m-%d')
after = end
num_after = 0
while num_after < 7 and after < datetime.datetime.today():
after += one_day
after_date = datetime.datetime.strftime(after, '%Y-%m-%d')
data_urls_after = dac.get_data_urls(roi, stop_time_as_string, after_date, 'S1_SLC')
create_sym_links(data_urls_after, after_sar_dir)
processor = SARPreProcessor(config=config_file, input=after_sar_dir, output=after_sar_dir)
list = processor.create_processing_file_list()
num_after = len(list[0]) + (len(list[1]) / 2.)
logging.info(f'Set end date for collecting S1 SLC products to {after_date}.')
# -
# We created extra directories for collecting the products. Let's clean up here.
# +
import shutil
shutil.rmtree(before_sar_dir)
shutil.rmtree(after_sar_dir)
# -
# Now, we are finally set to collect the data:
sar_data_urls = dac.get_data_urls(roi, before_date, after_date, 'S1_SLC')
create_sym_links(sar_data_urls, s1_slc_directory)
# Now that the data has been collected, we can run the actual SAR Pre-Processing. The Processing consists of three steps. The first two steps create one output product for one input product, while the third step merges information from multiple products. We can run steps 1 and 2 safely now on all the input folders.
processor = SARPreProcessor(config=config_file, input=s1_slc_directory, output=s1_grd_directory)
processor.create_processing_file_list()
logging.info('Start Pre-processing step 1')
processor.pre_process_step1()
logging.info('Finished Pre-processing step 1')
logging.info('Start Pre-processing step 2')
processor.pre_process_step2()
logging.info('Finished Pre-processing step 2')
# Step 3 needs to be performed for each product separately. To do this, we need to make sure we hand in the correct products only. The output of the second step is located in an intermediate folder. First, we collect all these files and sort them temporally.
# +
import glob
output_step2_dir = f'{s1_grd_directory}/step2'
sorted_input_files = glob.glob(f'{output_step2_dir}/*.dim')
sorted_input_files.sort(key=lambda x: x[len(output_step2_dir) + 18:len(output_step2_dir) + 33])
sorted_input_files
# -
# Now we can run the thrird step of the SAR Pre-Processing for every product for which there are at least 7 products before and 7 products after it available. For this, it is necessary to first create the file list, then to remove all files from it that shall not be considered during this step.
# +
output_step3_dir = f'{s1_grd_directory}/step3'
for end in range(14, len(sorted_input_files)):
file_list = processor.create_processing_file_list()
start = end-14
sub_list = sorted_input_files[start:end]
for i, list in enumerate(file_list):
for file in list:
processed_name = file.replace('.zip', '_GC_RC_No_Su_Co.dim')
processed_name = processed_name.replace(s1_slc_directory, output_step2_dir)
if processed_name not in sub_list:
list.remove(file)
processor.set_file_list(file_list)
logging.info(f'Start Pre-processing step 3, run {start}')
processor.pre_process_step3()
logging.info(f'Finished Pre-processing step 3, run {start}')
files = os.listdir(output_step3_dir)
for file in files:
shutil.move(os.path.join(output_step3_dir, file), s1_grd_directory)
# -
|
notebooks/MULTIPLY_SAR_Data_Access_and_Pre-Processing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Week 11, Part 2
#
# ### Topic
# 1. Fitting Hubble's law using Linear Regression
# 1. Estimating the beginning of time using regression coefficients
#
#
# resize
require(repr)
options(repr.plot.width=8, repr.plot.height=4)
# ## 1. Fitting Hubble's law using Linear Regression
# Let's read in the Hubble data:
hubble = read.csv('hubble.csv',stringsAsFactors=TRUE)
# Let's do a bit of data formatting to translate from weird astronomy units (parsecs) to kilometers:
distance = hubble[,1]/1e6 # pc -> Mpc, PS: 1 pc = 30,856,776,000,000.00 km or ~30 trillion km, or 3.26 light years
# Please note: parsecs is a unit of distance NOT time (sorry Star Wars).
# Luckily for us, the velocity is in units that are understandable as km/s:
vel = hubble[,2] # km/s
# Recall from lecture notes: looking at the *redshift* of the lights of galaxies $\rightarrow$ redder means receding further.
# Questions:
# 1. How linear is the relationship? (R)
# 1. Fit a line
# 1. Is our fit justified? (residuals, qqnorm)
# 1. Are there any outliers we should worry about?
#
# First things first, lets plot!
plot(distance, vel, pch=16, col=30, xlab='Distance in Mpc', ylab = 'Recessional Velocity in km/s')
# Note: there are things that have velocities <0, which means they are moving towards us.
# How linear is this thing?
R = cor(distance,vel)
print(R)
# So, pretty linear. Let's fit a line:
myLine = lm(formula = vel ~ distance, data = data.frame(distance,vel))
summary(myLine)
# We can see that we have a very small p-value for our slope - so probably a line is a good fit!
# Let's analyze our residuals:
# +
par(mfrow=c(1,2))
myResiduals = resid(myLine)
plot(myResiduals)
abline(h=0, col='red') # if perfect fit
# we can also check if our residuals are normal (one of our fit conditions)
qqnorm(myResiduals)
qqline(myResiduals)
# -
# While our residuals look pretty normal, we see that there are some variations these variations are actually "cosmic scatter" and its because there are groups of galaxies that are gravitationally interacting and changing the relation from a straight line.
#
# Let's plot our data & fit, first by grabbing the coefficients of our fit:
b0 = myLine$coefficients[1] # intercept
b1 = myLine$coefficients[2] # slope
# ... and then using the same sorts of plots we used before for our BAC-Beers dataset:
# +
options(repr.plot.width=7, repr.plot.height=5)
par(mfrow=c(2,2))
x = seq(-0.25, 2.5) # little negative just so we can see the points
myNewLine = b0 + b1*x
plot(x, myNewLine, type='l', col="green", xlab="Distance in Mpc", ylab="Recessional Velocity in km/s")
points(distance, vel, pch=16, col=30) # over plot observation points
plot(myLine, which=1) # Residuals plot
plot(myLine, which=2) # qq-norm plot
# now, lets add another plot to our meagurie
plot(myLine, which=5) # residuals vs. leverage
# -
# ## 2. Estimating the beginning of time using regression coefficients
#
# Just for fun, let's use our linear fit's coefficients to calculate the beginning of time.
#
# This brings us to hubbles law: $V = H_0 \times D$ where $H_0$ is a constant in units of velocity/Mpc = km/s/Mpc (weird) where $H_0$ is hubble's constant = b1, i.e. $H_0$ is just the slope of our linear fit!
hubbles_const = b1
print(unname(hubbles_const)) # in km/s/Mpc
# Note: actual value is like ~70 km/s/Mpc. But even though it's pretty off, it was a huge discovery!!
#
# If we assume we are not in a special place in the universe, this means that the Universe is expanding away from itself you can think of for, example 2D universe represnted as a balloon and blowing up the balloon as the expanding universe you can draw some dots on the balloon and see that they expand away from eachother and the ones furthest from your mouth expand away from you fastest.
#
# <img src="https://www.universetoday.com/wp-content/uploads/2015/02/Balloon-universe-expansion.jpg" width='600px'>
#
# So, let's real quick, calculate the begining of time ok?
#
# We recall that velocity = distance/time, so then time = distance/velocity we have a realationship for velocity and distance in the hubble constant which is the inverse of this!
beginning_of_time = 1.0/( unname(hubbles_const) * (1.0/3.08e19) )
# the last bit of numbers is just a unit conversion from Mpc to km
beginning_of_time
# That is in seconds, so lets make it in years:
beginning_of_time_in_yrs = beginning_of_time/3.15e7 # into years
beginning_of_time_in_yrs
# This is still a large number, so let's finally transform it into Gyrs aka billions of years:
beginning_of_time_in_Gyrs = beginning_of_time/3.15e7/1e9 # into Gyears
beginning_of_time_in_Gyrs
# So, this says the Universe is ~2 Gyrs old. In realaity, here we have over estimated the hubble's constant so we are underestimating the age of the universe, which is actually about 13.5 Gyrs (or 13.5 billion years) old.
#
# But in general, good job everybody! With linear regression you just figured out (approximately) when the beginning of time was! Hurray!! Nobel prizes for everybody :D
|
week11/prep_hubblesExample_part2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: LVV-RI-NEW
# language: python
# name: lvv-ri-new
# ---
# **0.0-test.ipynb** for testing your `lvv-ri-new` environment
#
# # python, numpy, pandas, matplotlib, rpy2 and more
# If you successfully run through this notebook, then your Python environment is configured correctly.
# # How to use the Jupyter Notebook?
# [Jupyter Notebook](http://jupyter.org/) is a useful tool for experimenting with code. Everything of code and text is written in HTML, Markdown and Python.
#
# Use arrow keys to navigate between cells. Hit ENTER on a cell to switch to editing mode. ESC to get back.
print("This is a Jupyter cell containing Python code. Hit 'Run' in the menu to execute the cell. ")
# You can also run cells by typing **Shift+Enter** or **Ctrl+Enter**. Try running the cell above by using both of these.
# You will find more information via the Help menu above.
# `lvv-ri-new` is using Jupyter Notebooks with both Python and R for the coding. <br>Here is a good tutorial on Jupyter Notebook: [Jupyter Notebook Tutorial: The Definitive Guide](https://www.datacamp.com/community/tutorials/tutorial-jupyter-notebook).
# # Import libraries
# These are libaries that will be frequently used in the course:
# To diosplay plots directly in the notebook:
# %matplotlib inline
# A frequently used plotting libraryp:
import matplotlib
import matplotlib.pyplot as plt
# An extension of matplotlib making it easy to generate even nicer plots:
import seaborn as sns
# A numerical libray for efficient manipulation of matrices (and more):
import numpy as np
# To read, write and proecess tabular data:
import pandas as pd
# For machine learning:
import sklearn
# +
# For machine learning:
#import pandas_ml (not compatible any longer)
# +
# For rpy2:
# import tzlocal
# +
# For using R:
#import rpy2
#from rpy2.robjects import r, pandas2ri
# +
#from rpy2.robjects.packages import importr
#utils = importr('utils')
# If needed to install - uncomment the following. NOTE: MacOS and 'stringi' might cause a challenge
# see https://stackoverflow.com/questions/31038636/package-stringi-does-not-work-after-updating-to-r3-2-1
# utils.install_packages('stringi', repos='http://cran.rstudio.com')
#utils.install_packages('tidyr')
#utils.install_packages('lazyeval')
#utils.install_packages('lme4')
#utils.install_packages('ggplot2')
#utils.install_packages('GGally')
#utils.install_packages('foreign')
# -
# Supress some warnings:
import warnings
warnings.filterwarnings('ignore')
# +
# For using pandas2ri:
#pandas2ri.activate()
#from rpy2.robjects.lib.tidyr import DataFrame
# -
# If errors like<br>
# ```python
# RRuntimeError: Error in loadNamespace(name) : there is no package called 'tidyr'
# ```
# see cell below:
# +
# #%reload_ext rpy2.ipython
# +
# #%%R
#$R.version$system
# +
# #%%R
#R.version$version.str
# +
# #%R library(foreign); #library(readxl)
# -
# # Test libraries
# **REMARK:** The aim of the following is to test the installation, not doing relevant `lvv-ri`analysis.
# ## `Numpy`
import numpy as np
a = np.array([1, 2, 3])
print(type(a))
e = np.random.random((3,3))
e
# ## `matplotlib`: a simple plot
# %matplotlib inline
import matplotlib.pyplot as plt
# The following will give a figure displaying a sine function:
# +
# Data to be plotted (generated by Numpy)
t = np.arange(0.0, 2.0, 0.01)
s = 1 + np.sin(2 * np.pi * t)
# Make a figure of given size
f, ax = plt.subplots(figsize=(8,4))
# Plot t versus s
plt.plot(t, s)
# Add a title and labels:
plt.title('A simple plot')
plt.xlabel('time (s)')
plt.ylabel('voltage')
# Show plot:
plt.show()
# -
# ## `Seaborn`: a more advanced plot
import seaborn as sns
# Source: [Link](https://seaborn.pydata.org/examples/scatterplot_categorical.html)
# +
sns.set(style="whitegrid", palette="muted")
# Load the iris dataset
iris = sns.load_dataset("iris")
# "Melt" dataset to "long-form" or "tidy" representation
iris = pd.melt(iris, "species", var_name="measurement")
# Make a figure with given size
f, ax = plt.subplots(figsize=(8,8))
# Draw a categorical scatterplot to show each observation
sns.swarmplot(x="measurement", y="value", hue="species", data=iris, size=5, ax=ax)
plt.show()
# -
# ## `Pandas`
import pandas as pd
df = pd.read_csv('../data/lvv_ri_data.csv')
df.head()
df['AcquisitionYearsW1'].hist()
plt.title("Histogram of age at inclusion")
plt.xlabel("Age")
plt.show()
# ## sklearn.metrics: `confusion_matrix`
from sklearn.metrics import confusion_matrix
y_true = ['slow', 'slow', 'slow', 'fast', 'slow', 'slow', 'fast', 'medium',
'fast', 'medium', 'slow', 'medium', 'fast', 'medium', 'medium',
'medium', 'medium', 'slow', 'medium', 'medium', 'fast', 'fast',
'fast', 'fast', 'medium', 'slow', 'slow', 'fast', 'medium',
'medium', 'slow', 'slow', 'slow', 'slow', 'fast', 'fast', 'fast',
'medium', 'medium', 'medium', 'fast', 'fast', 'medium', 'medium',
'medium', 'medium', 'fast', 'slow', 'slow', 'slow', 'slow', 'slow',
'medium', 'fast', 'slow', 'slow', 'fast', 'slow', 'medium', 'fast',
'slow', 'medium', 'slow', 'slow', 'medium', 'fast', 'fast', 'fast',
'fast', 'fast', 'fast', 'fast', 'medium', 'slow']
y_pred = ['fast', 'medium', 'slow', 'medium', 'medium', 'slow', 'medium',
'fast', 'slow', 'medium', 'fast', 'fast', 'slow', 'medium',
'medium', 'medium', 'fast', 'fast', 'slow', 'fast', 'fast', 'fast',
'fast', 'slow', 'slow', 'slow', 'slow', 'slow', 'medium', 'fast',
'fast', 'fast', 'medium', 'fast', 'medium', 'medium', 'medium',
'slow', 'slow', 'medium', 'fast', 'fast', 'medium', 'medium',
'medium', 'fast', 'slow', 'medium', 'slow', 'slow', 'medium',
'fast', 'fast', 'fast', 'fast', 'slow', 'slow', 'fast', 'slow',
'fast', 'slow', 'medium', 'slow', 'slow', 'slow', 'fast', 'fast',
'fast', 'fast', 'fast', 'fast', 'fast', 'fast', 'fast']
y_confusion_matrix = confusion_matrix(y_true, y_pred)
print("y Confusion matrix:\n%s" % y_confusion_matrix)
# +
#y_confusion_matrix.stats()
#y_confusion_matrix.print_stats()
# -
# From previous pandas_ml
# ```code
# Confusion Matrix:
#
# Predicted fast medium slow __all__
# Actual
# fast 14 5 6 25
# medium 8 10 6 24
# slow 10 5 10 25
# __all__ 32 20 22 74
#
#
# Overall Statistics:
#
# Accuracy: 0.4594594594594595
# 95% CI: (0.3429195917062334, 0.5793443273251775)
# No Information Rate: ToDo
# P-Value [Acc > NIR]: 0.3608177987401836
# Kappa: 0.18815139879319806
# Mcnemar's Test P-Value: ToDo
#
#
# Class Statistics:
#
# Classes fast medium slow
# Population 74 74 74
# P: Condition positive 25 24 25
# N: Condition negative 49 50 49
# Test outcome positive 32 20 22
# Test outcome negative 42 54 52
# TP: True Positive 14 10 10
# TN: True Negative 31 40 37
# FP: False Positive 18 10 12
# FN: False Negative 11 14 15
# TPR: (Sensitivity, hit rate, recall) 0.56 0.416667 0.4
# TNR=SPC: (Specificity) 0.632653 0.8 0.755102
# PPV: Pos Pred Value (Precision) 0.4375 0.5 0.454545
# NPV: Neg Pred Value 0.738095 0.740741 0.711538
# FPR: False-out 0.367347 0.2 0.244898
# FDR: False Discovery Rate 0.5625 0.5 0.545455
# FNR: Miss Rate 0.44 0.583333 0.6
# ACC: Accuracy 0.608108 0.675676 0.635135
# F1 score 0.491228 0.454545 0.425532
# MCC: Matthews correlation coefficient 0.183927 0.228387 0.160499
# Informedness 0.192653 0.216667 0.155102
# Markedness 0.175595 0.240741 0.166084
# Prevalence 0.337838 0.324324 0.337838
# LR+: Positive likelihood ratio 1.52444 2.08333 1.63333
# LR-: Negative likelihood ratio 0.695484 0.729167 0.794595
# DOR: Diagnostic odds ratio 2.19192 2.85714 2.05556
# FOR: False omission rate 0.261905 0.259259 0.288462
# ```
# +
# Previously, using pandas_ml
#y_confusion_matrix.plot()
#plt.show()
# +
import seaborn as sn
class_names = ['fast', 'medium', 'slow']
df_cm = pd.DataFrame(y_confusion_matrix, index = [i for i in class_names],
columns = [i for i in class_names])
plt.figure(figsize = (10,7))
ax=sn.heatmap(df_cm, annot=True, fmt='d', square=True)
bottom, top = ax.get_ylim()
ax.set_ylim(bottom + 0.5, top - 0.5)
plt.title('Confusion matrix', fontsize= 16)
plt.xlabel('Predicted')
plt.ylabel('Actual')
plt.show()
# -
#df_y_cm = y_confusion_matrix.to_dataframe()
df_y_cm = pd.DataFrame.from_records(y_confusion_matrix)
df_y_cm
x = df_y_cm.values.copy()
x[0][0] = df_y_cm.values[2][2]
x[0][2] = df_y_cm.values[2][0]
x[1][0] = df_y_cm.values[1][2]
x[1][2] = df_y_cm.values[1][0]
x[2][0] = df_y_cm.values[0][2]
x[2][2] = df_y_cm.values[0][0]
x
# ## `utils.py`
# +
from utils import plot_confusion_matrix, plot_confusion_matrix_with_colorbar
cm = x
classes = ['slow', 'medium', 'fast']
plot_confusion_matrix_with_colorbar(cm, classes=classes,
title='Confusion matrix\n',
# cmap='gray',
figsize=(5,5))
plt.ylabel('Observed RI')
plt.xlabel('Predicted RI')
# fn_cm_pdf = '../figures/y_confusion_matrix.pdf'
#plt.savefig(fn_cm_pdf, transparent=True)
plt.show()
# -
# ## `scikit-learn`: machine learning
from sklearn import datasets
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
data = datasets.load_breast_cancer()
X = data['data']
y = data['target']
features = data['feature_names']
labels = data['target_names']
print(features)
print(labels)
X_train, X_test, y_train, y_test = train_test_split(X, y)
rf = RandomForestClassifier(n_estimators=100)
rf.fit(X_train, y_train)
predictions = rf.predict(X_test)
accuracy_score(y_test, predictions) * 100
# ## `rpy2`, `lmer` and `ggplot`: Analysis in R
# +
# #%%R
# library(foreign)
#Rdf <- read.csv(file="../data/lvv_ri_data.csv",head=TRUE, sep=",")
# +
# #%%R
#names(Rdf)
# +
# #%%R
#dim(Rdf)
# +
# #%%R
#Rdf$Stroop_3_R_W3
# +
# #%%R
#mean(Rdf$Left.Lateral.Ventricle_W1)
# +
# #%%R
#sd(Rdf$Left.Lateral.Ventricle_W1)
# +
# #%%R
#min(Rdf$AcquisitionYearsW1)
# +
# #%%R
#max(Rdf$AcquisitionYearsW1)
# +
# #%%R
#RdfS <- na.omit(Rdf, select=c(Subject, Stroop_3_R_W3, Left.Lateral.Ventricle_W1, Left.Lateral.Ventricle_W2, Left.Lateral.Ventricle_W3))#$RfitLVV <- lm(Stroop_3_R_W3 ~ Left.Lateral.Ventricle_W1 + Left.Lateral.Ventricle_W2 + Left.Lateral.Ventricle_W3, data=RdfS)
# +
# #%%R
#names(RfitLVV)
# +
# #%%R
#head(RfitLVV)
# +
# from rpy2.robjects import r, pandas2ri
# pandas2ri.activate()
# pd_RfitLVV = r['RfitLVV']
# temp = pandas2ri.ri2py(pd_RfitLVV)
# print(temp[0])
#print(temp[1])
#print(temp[2])
# print(temp[3])
#print(temp[4])
#print(temp[5])
#print(temp[6])
#print(temp[7])
#print(temp[8])
# +
# print(pd_RfitLVV.names)
# pandas2ri.ri2py(pd_RfitLVV).names
# +
# # %%R
# summary(RfitLVV)
# +
# # %R anova(RfitLVV)
# +
# # %R plot(RfitLVV$resid)
# +
# # %R hist(RfitLVV$resid)
# -
#
# A group of diagnostic plots (residual, qq, scale-location, leverage) to assess model performance when applied to a fitted linear regression model.
#
# https://data.library.virginia.edu/diagnostic-plots/
# +
# # %R par(mfrow=c(2,2)); plot(RfitLVV) # Plot four panels on the same figure
# -
df = pd.read_csv('../data/lvv_ri_data.csv')
df.head(7).T
# Reading a Pandas dataframe (df) into R (data)
# +
# Previously:
# #%R -i df
#
#data = df
# +
# import pandas as pd
# import rpy2.robjects as ro
# from rpy2.robjects.packages import importr
# from rpy2.robjects import pandas2ri
# from rpy2.robjects.conversion import localconverter
# with localconverter(ro.default_converter + pandas2ri.converter):
# data = ro.conversion.py2ro(df)
# print(data)
# +
# # %%R
# names(data)
# -
# ```
# # %%R
#
# library(ggplot2)
# library(GGally)
#
# data$Gender = data$Sex
#
# pm <- ggpairs(
# data, mapping = aes(color = Gender),
# title = "Pairs plot by gender",
# legend = 1,
# columns = c('AcquisitionYearsW1',
# 'Left.Lateral.Ventricle_W1',
# 'Left.Lateral.Ventricle_W2',
# 'Left.Lateral.Ventricle_W3',
# 'EstimatedTotalIntraCranialVol_W3',
# 'Stroop_3_R_W3'),
# lower = list(
# continuous = 'smooth'
# ))
#
#
# pm = pm + theme(axis.text.x = element_text(angle = 90, hjust = 1))
#
# pm = pm + theme(
# axis.text = element_text(size = 10),
# axis.title = element_text(size = 10),
# legend.background = element_rect(fill = "white"),
# panel.grid.major = element_line(colour = NA),
# # panel.grid.minor = element_blank(),
# panel.grid.minor = element_line(size = 0.25, linetype = 'solid', colour = "white"),
# panel.background = element_rect(fill = "grey95"),
# plot.title = element_text(size=24)
# )
#
# print(pm)
# ```
# Converting the R dataframes to Pandas DataFrames using `rpy2` <br>
# ```
# with localconverter(ro.default_converter + pandas2ri.converter):
# pd_data = ro.conversion.ri2ro(data)
#
# pd_data
# ```
# +
# pd_data.head()
# -
# LME example uning `lmer` from `lme4` and the ChickWeight data https://stat.ethz.ch/R-manual/R-devel/library/datasets/html/ChickWeight.html
# +
# # %%R
# library(lme4)
# data = ChickWeight
# +
# from rpy2.robjects import r, pandas2ri
# pandas2ri.activate()
# pd_data = r['data']
# +
# pd_data.names
# +
#import rpy2.ipython.html
#rpy2.ipython.html.init_printing()
#print(pd_data.head(12))
# -
# ##### Now using https://stackoverflow.com/questions/35757994/converting-lme4-ranef-output-to-data-frame-with-rpy2
# +
# # %reload_ext rpy2.ipython
# # %Rpush pd_data
# -
# ```
# # %%R
#
# library(lme4)
#
# m <- lmer(weight ~ Time * Diet + (1 + Time | Chick), data=pd_data, REML=F)
#
# rfs <- ranef(m)$cat
# ffs <- fixef(m)
#
# print(names(ffs))
# ```
# +
# #%Rpull ffs
# +
#print(ffs[0])
#print(ffs[1])
#print(ffs[2])
#print(ffs[3])
# +
# #%R summary(m)
# +
# #%%R
#plot(m)
# -
# ## Python and statsmodels
#
# https://medium.com/@emredjan/emulating-r-regression-plots-in-python-43741952c034
#
# https://emredjan.github.io/blog/2017/07/11/emulating-r-plots-in-python/
#
# Think Stats: Exploratory Data Analysis in Python (http://www.greenteapress.com/thinkstats2/html/index.html)
# %matplotlib inline
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import statsmodels.formula.api as smf
import statsmodels.stats.multicomp
from statsmodels.graphics.gofplots import ProbPlot
cwit = pd.read_csv('../data/cwit_data.csv')
cwit.describe()
# +
model_f12 = 'Stroop_3_R_W3 ~ Stroop_1_R_W3 + Stroop_2_R_W3'
model_f2 = 'Stroop_3_R_W3 ~ Stroop_2_R_W3'
model_f1 = 'Stroop_3_R_W3 ~ Stroop_1_R_W3'
model12 = smf.ols(formula=model_f12, data=cwit)
model12_fit = model12.fit()
model2 = smf.ols(formula=model_f2, data=cwit)
model2_fit = model2.fit()
model1 = smf.ols(formula=model_f1, data=cwit)
model1_fit = model1.fit()
# -
# **model1: 'Stroop_3_R_W3 ~ Stroop_1_R_W3'**
# Seeing if the overall model is significant
print(f"Overall model F({model1_fit.df_model: .0f},{model1_fit.df_resid: .0f}) = {model1_fit.fvalue: .3f}, p = {model1_fit.f_pvalue: .4f}")
from IPython.core.display import HTML
HTML(model1_fit.summary().tables[0].as_html())
HTML(model1_fit.summary().tables[1].as_html())
HTML(model1_fit.summary().tables[2].as_html())
# The *Durban-Watson* tests is to detect the presence of autocorrelation, *Jarque-Bera* tests the assumption of normality, *Omnibus* tests the assumption of homogeneity of variance, and the *Condition Number* assess multicollinearity. Condition Number values over 20 are indicative of multicollinearity.
# **model2: 'Stroop_3_R_W3 ~ Stroop_2_R_W3'**
# Seeing if the overall model is significant
print(f"Overall model F({model2_fit.df_model: .0f},{model2_fit.df_resid: .0f}) = {model2_fit.fvalue: .3f}, p = {model2_fit.f_pvalue: .4f}")
HTML(model2_fit.summary().tables[0].as_html())
HTML(model2_fit.summary().tables[1].as_html())
HTML(model2_fit.summary().tables[2].as_html())
# **model12: 'Stroop_3_R_W3 ~ Stroop_1_R_W3 + Stroop_2_R_W3'**
# Seeing if the overall model is significant
print(f"Overall model F({model12_fit.df_model: .0f},{model12_fit.df_resid: .0f}) = {model12_fit.fvalue: .3f}, p = {model12_fit.f_pvalue: .4f}")
HTML(model12_fit.summary().tables[0].as_html())
HTML(model12_fit.summary().tables[1].as_html())
HTML(model12_fit.summary().tables[2].as_html())
# **model2: Get an instance of Influence with influence and outlier measures**
infl12 = model12_fit.get_influence()
df_infl12 = infl12.summary_frame()
df_infl12.head()
# **Anova table for one or more fitted linear models**<br>
# Model statistics are given in the order of args. Models must have been fit using the formula api.
#
# Python for Data Science (https://pythonfordatascience.org/anova-2-way-n-way)
#
# There are 3 types of sum of squares that should be considered when conducting an ANOVA, by default Python and R uses Type 2, whereas SAS tends to use Type 3. The differences in the types of sum of squares is out of this page’s scope; but you should research the differences to decide which type you should use for your study.
import statsmodels.api as sm
sm.stats.anova_lm(model2_fit, typ=2)
anov12 = sm.stats.anova_lm(model12_fit, typ=2)
anov12
# i.e. each factor has an independent significant effect on the mean Stroop_3_R_W3
# **Calculating effect size**
# +
def anova_table(aov):
aov['mean_sq'] = aov[:]['sum_sq']/aov[:]['df']
aov['eta_sq'] = aov[:-1]['sum_sq']/sum(aov['sum_sq'])
aov['omega_sq'] = (aov[:-1]['sum_sq']-(aov[:-1]['df']*aov['mean_sq'][-1]))/(sum(aov['sum_sq'])+aov['mean_sq'][-1])
cols = ['sum_sq', 'mean_sq', 'df', 'F', 'PR(>F)', 'eta_sq', 'omega_sq']
aov = aov[cols]
return aov
anova_table(anov12)
# -
# $\omega^2$ is a better measure of effect size since it’s unbiased in it’s calculation. It takes into account the degrees of freedom, whereas $\eta^2$ does not. Side note, $\eta^2$ and $R^2$ are the same thing in the ANOVA framework. Each factor, Stroop_1 and Stroop_2, has a small effect on the mean Stroop_3_R_W3 reaction time
# **Comparing models**<br>
# When you use `anova(lm.1,lm.2,test="Chisq")`, it performs the Chi-square test to compare lm.1 and lm.2 (i.e. it tests whether reduction in the residual sum of squares are statistically significant or not). Note that this makes sense only if lm.1 and lm.2 are nested models.<br>
# https://stat.ethz.ch/R-manual/R-patched/library/stats/html/anova.lm.html
from statsmodels.stats.api import anova_lm
table1_12 = anova_lm(model1_fit, model12_fit, test="F")
print(table1_12)
table2_12 = anova_lm(model2_fit, model12_fit, test="F")
print(table2_12)
# +
model = model12
model_fit = model12_fit
# fitted values (need a constant term for intercept)
model_fitted_y = model_fit.fittedvalues
# model residuals
model_residuals = model_fit.resid
# normalized residuals
model_norm_residuals = model_fit.get_influence().resid_studentized_internal
# absolute squared normalized residuals
model_norm_residuals_abs_sqrt = np.sqrt(np.abs(model_norm_residuals))
# absolute residuals
model_abs_resid = np.abs(model_residuals)
# leverage, from statsmodels internals
model_leverage = model_fit.get_influence().hat_matrix_diag
# cook's distance, from statsmodels internals
model_cooks = model_fit.get_influence().cooks_distance[0]
# -
plt.hist(model_residuals, facecolor='w', edgecolor='k', lw=2, alpha=1.0)
plt.title('Residuals')
plt.show()
# **Residual plot**
# +
plot_lm_1 = plt.figure(1)
plot_lm_1.set_figheight(8)
plot_lm_1.set_figwidth(12)
plot_lm_1.axes[0] = sns.residplot(model_fitted_y, 'Stroop_3_R_W3', data=cwit,
lowess=True,
scatter_kws={'alpha': 0.5},
line_kws={'color': 'red', 'lw': 1, 'alpha': 0.8})
plot_lm_1.axes[0].set_title('Residuals vs Fitted')
plot_lm_1.axes[0].set_xlabel('Fitted values')
plot_lm_1.axes[0].set_ylabel('Residuals')
# annotations
abs_resid = model_abs_resid.sort_values(ascending=False)
abs_resid_top_3 = abs_resid[:3]
for i in abs_resid_top_3.index:
plot_lm_1.axes[0].annotate(i,
xy=(model_fitted_y[i],
model_residuals[i]));
# -
# **QQ-plot**
# +
QQ = ProbPlot(model_norm_residuals)
plot_lm_2 = QQ.qqplot(line='45', alpha=0.5, color='#4C72B0', lw=1)
plot_lm_2.set_figheight(8)
plot_lm_2.set_figwidth(12)
plot_lm_2.axes[0].set_title('Normal Q-Q')
plot_lm_2.axes[0].set_xlabel('Theoretical Quantiles')
plot_lm_2.axes[0].set_ylabel('Standardized Residuals');
# annotations
abs_norm_resid = np.flip(np.argsort(np.abs(model_norm_residuals)), 0)
abs_norm_resid_top_3 = abs_norm_resid[:3]
for r, i in enumerate(abs_norm_resid_top_3):
plot_lm_2.axes[0].annotate(i,
xy=(np.flip(QQ.theoretical_quantiles, 0)[r],
model_norm_residuals[i]));
# -
# **Scale-location plot**
# +
plot_lm_3 = plt.figure(3)
plot_lm_3.set_figheight(8)
plot_lm_3.set_figwidth(12)
plt.scatter(model_fitted_y, model_norm_residuals_abs_sqrt, alpha=0.5)
sns.regplot(model_fitted_y, model_norm_residuals_abs_sqrt,
scatter=False,
ci=False,
lowess=True,
line_kws={'color': 'red', 'lw': 1, 'alpha': 0.8})
plot_lm_3.axes[0].set_title('Scale-Location')
plot_lm_3.axes[0].set_xlabel('Fitted values')
plot_lm_3.axes[0].set_ylabel('$\sqrt{|Standardized Residuals|}$');
# annotations
abs_sq_norm_resid = np.flip(np.argsort(model_norm_residuals_abs_sqrt), 0)
abs_sq_norm_resid_top_3 = abs_sq_norm_resid[:3]
for i in abs_norm_resid_top_3:
plot_lm_3.axes[0].annotate(i,
xy=(model_fitted_y[i],
model_norm_residuals_abs_sqrt[i]));
# -
# **Leverage plot**
# +
plot_lm_4 = plt.figure(4)
plot_lm_4.set_figheight(8)
plot_lm_4.set_figwidth(12)
plt.scatter(model_leverage, model_norm_residuals, alpha=0.5)
sns.regplot(model_leverage, model_norm_residuals,
scatter=False,
ci=False,
lowess=True,
line_kws={'color': 'red', 'lw': 1, 'alpha': 0.8})
plot_lm_4.axes[0].set_xlim(0, 0.20)
plot_lm_4.axes[0].set_ylim(-3, 5)
plot_lm_4.axes[0].set_title('Residuals vs Leverage')
plot_lm_4.axes[0].set_xlabel('Leverage')
plot_lm_4.axes[0].set_ylabel('Standardized Residuals')
# annotations
leverage_top_3 = np.flip(np.argsort(model_cooks), 0)[:3]
for i in leverage_top_3:
plot_lm_4.axes[0].annotate(i,
xy=(model_leverage[i],
model_norm_residuals[i]))
# shenanigans for cook's distance contours
def graph(formula, x_range, label=None):
x = x_range
y = formula(x)
plt.plot(x, y, label=label, lw=1, ls='--', color='red')
p = len(model_fit.params) # number of model parameters
graph(lambda x: np.sqrt((0.5 * p * (1 - x)) / x),
np.linspace(0.001, 0.200, 50),
'Cook\'s distance') # 0.5 line
graph(lambda x: np.sqrt((1 * p * (1 - x)) / x),
np.linspace(0.001, 0.200, 50)) # 1 line
plt.legend(loc='upper right');
# -
cwit['resid12'] = np.array(model_residuals)
cwit.head()
cwit_resid12 = pd.DataFrame([cwit['Subject'], cwit['Stroop_3_R_W3'], cwit['resid12']]).T
cwit_resid12.head()
# +
#cwit_resid12.to_csv('../results/cwit_resid12.csv', index=False)
# -
|
notebooks/0.0-test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# I used `pygetpapers` to download all the suplimentary files analysed in the notebook.
#
# Query:
# ```
# pygetpapers --terms C:\Users\shweata\tps_pmcid.txt -k 400 -o "tps_300" --supp
# ```
# Instead of specifying the query, I created a custom corpus by specifying PMCIDs. To do so, I created a text file with PMCIDS (comma-separted), and used `--terms` flags. You can check the text file, [here](https://github.com/petermr/dictionary/blob/main/tps_data_availability/tps_pmcid.txt).
#
# The corpus has 283 CTrees. But all of them wouldn't have supplementary data.
#
# The purpose of this notebook is to analyse how many of these papers have supplemental data, and what formats they are in.
import os
from glob import glob
import pathlib
from collections import Counter
import pandas as pd
import zipfile
HOME= os.path.expanduser("~") # gets home directory
TPS_DIRECTORY = 'tps_300' # CProject directory
get_num_supp = (glob(os.path.join(HOME, TPS_DIRECTORY, "*", "supplementaryfiles")))
supp_glob = (glob(os.path.join(HOME, TPS_DIRECTORY, "*", "supplementaryfiles", "*"),recursive=True))
#C:\Users\shweata\tps_300\PMC3195254\supplementaryfiles
len(get_num_supp)
supp_extension = []
for supp_file_name in supp_glob:
supp_extension.append((pathlib.Path(supp_file_name).suffix).lower())
ext_counts = Counter(supp_extension)
df = pd.DataFrame.from_dict(ext_counts, orient='index')
df.plot(kind='bar')
ext_counts
# +
zip_file_names = []
for supp_file_name in supp_glob:
filename, file_extension = os.path.splitext(supp_file_name)
if file_extension.lower() == '.zip':
with zipfile.ZipFile(supp_file_name, 'r') as my_zip:
zip_file_names.extend((my_zip.namelist()))
len(zip_file_names)
# -
zip_supp_extension = []
for zip_file_name in zip_file_names:
zip_supp_extension.append((pathlib.Path(zip_file_name).suffix).lower())
zip_ext_counts = Counter(zip_supp_extension)
df = pd.DataFrame.from_dict(zip_ext_counts, orient='index')
df.plot(kind='bar')
zip_ext_counts
df = pd.DataFrame(get_num_supp)
df.columns=['path']
PMCID = []
for index, row in df.iterrows():
split_path = row["path"].split('\\')
PMCID.append(split_path[4])
df["PMCID"] = PMCID
df.to_csv('supp_exists.csv')
|
tps_data_availability/data_formats_tps.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using Remix's Partridge with SFMTA GTFS data
import partridge as ptg
sfmta_data = 'data/downloads/sfmta/'
# +
service_ids = ptg.read_busiest_date(sfmta_data)[1]
view = {'trips.txt': {'service_id': service_ids}}
feed = ptg.load_geo_feed(sfmta_data, view)
# -
feed.shapes.head()
|
GTFS/partridge.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Spyder)
# language: python3
# name: python3
# ---
# + id="optical-frame" outputId="ee4748b3-0d3f-482c-a9c8-8bc504857d09"
# TensorFlow is an open source machine learning library
import tensorflow as tf
# NumPy is a math library
import numpy as np
# Matplotlib is a graphing library
import matplotlib.pyplot as plt
# math is Python's math library
import math
# We'll use Keras to create a simple model architecture
# Note: Changed tf.keras below to tensorflow.keras (code from the book)
from tensorflow.keras import layers
# Define the number of nodes in each layer of the network
# Layer1 with 24 and Layer2 with 16 nodes is better than 16 and 24 respectively.
DENSE1_SIZE = 32
DENSE2_SIZE = 16
DENSE3_SIZE = 8
NUM_OF_EPOCHS = 100 #Changed 600 to 500 because no change in loss after 500
BATCH_SIZE = 16
# We'll generate this many sample datapoints
SAMPLES = 1200
# Set a "seed" value, so we get the same random numbers each time we run this
# notebook. Any number can be used here.
SEED = 1523
np.random.seed(SEED)
tf.random.set_seed(SEED)
# Generate a uniformly distributed set of random numbers in the range from
# 0 to 2π, which covers a complete Sine wave oscillation
# You can generate only a half of the Sine wave by changing high from 2pi to pi
x_values = np.random.uniform(low=0, high=2*math.pi, size=SAMPLES)
# Shuffle the values to guarantee they're not in order
np.random.shuffle(x_values)
y_values = np.sin(x_values)
# Calculate the corresponding sine values
np.random.uniform(low=0, high=2*math.pi, size=SAMPLES)
# Plot our data. The 'b.' argument tells the library to print blue dots.
plt.plot(x_values, y_values, 'b.')
plt.show()
# Add a small random number to each y value
# When the number of Layer 1 and 2 nodes are increased, the noise level
# from 0.001 to some higher value (0.01, etc) still impacts less the quality
# of predictions.
y_values += 0.1 * np.random.randn(*y_values.shape)
# Plot our data
plt.plot(x_values, y_values, 'b.')
plt.show()
# We'll use 60% of our data for training and 20% for testing. The remaining 20%
# will be used for validation. Calculate the indices of each section.
TRAIN_SPLIT = int(0.6 * SAMPLES)
TEST_SPLIT = int(0.2 * SAMPLES + TRAIN_SPLIT)
# Use np.split to chop our data into three parts.
# The second argument to np.split is an array of indices where the data will be
# split. We provide two indices, so the data will be divided into three chunks.
x_train, x_validate, x_test = np.split(x_values, [TRAIN_SPLIT, TEST_SPLIT])
y_train, y_validate, y_test = np.split(y_values, [TRAIN_SPLIT, TEST_SPLIT])
# Double check that our splits add up correctly
assert (x_train.size + x_validate.size + x_test.size) == SAMPLES
# Plot the data in each partition in different colors:
plt.plot(x_train, y_train, 'b.', label="Train")
plt.plot(x_validate, y_validate, 'y.', label="Validate")
plt.plot(x_test, y_test, 'r.', label="Test")
plt.legend()
plt.show()
# Activation function used here is relu (Rectified Linear Unit)
def relu(input):
return max(0.0, input)
model_3 = tf.keras.Sequential()
# First layer takes a scalar input and feeds it through 16 "neurons." The
# neurons decide whether to activate based on the 'relu' activation function.
model_3.add(layers.Dense(DENSE1_SIZE, activation='relu', input_shape=(1,)))
# The new second layer may help the network learn more Sine representations
model_3.add(layers.Dense(DENSE2_SIZE, activation='relu'))
model_3.add(layers.Dense(DENSE3_SIZE, activation='relu'))
# Final layer is a single neuron, since we want to output a single value
model_3.add(layers.Dense(1))
# Compile the model using a standard optimizer and loss function for regression
model_3.compile(optimizer='rmsprop', loss='mse', metrics=['mae'])
# Show a summary of the model
model_3.summary()
# + id="advised-facing" outputId="18ccf507-97fd-4995-f214-eccc9d2dfb8b"
# 0. Did not get the same accurate predection as shown in the book epochs=600
# 1. So, increased the epochs from 600 to 1000 to see if there is any improvement
# Saw a great improvment on the negative cycle, which was closer to actual
# 2. Increased the epochs still further from 1000 to 1200 to see if it improves further
# 3. Still some error near negative peak, so changed it from 1200 to 1500
# 4. So far, the batch size was kept at 16. Now epochs=1500 and batch_size=32 tried out
# The previous run +v peak also got affected a bit, but -ve peak became better
# 5. Change now epochs=1500 and batch_size=64, it became worse.
# 6. Went back to epochs=1500 and batch_size=16
# 7. Error does not improve beyond Epoch=1000, so reduced it from 1500 to 1000
history_3 = model_3.fit(x_train, y_train, epochs=NUM_OF_EPOCHS, batch_size=BATCH_SIZE,
validation_data=(x_validate, y_validate))
# + id="abandoned-fruit" outputId="b096154e-30de-4899-8dbb-68c37e161a65"
# Draw a graph of the loss, which is the distance between
# the predicted and actual values during training and validation.
loss = history_3.history['loss']
val_loss = history_3.history['val_loss']
epochs = range(1, len(loss) + 1)
plt.plot(epochs, loss, 'g.', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# + id="suitable-seafood" outputId="03483fdb-35fb-4d52-a12e-4a38afb90785"
# Exclude the first few epochs so the graph is easier to read
SKIP = 10
plt.clf()
plt.plot(epochs[SKIP:], loss[SKIP:], 'g.', label='Training loss')
plt.plot(epochs[SKIP:], val_loss[SKIP:], 'b.', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# + id="complete-clearing" outputId="3213c455-8109-4a0d-fbc4-3d1bb6399e85"
#Finally, we plot the mean absolute error for the same set of epochs:
plt.clf()
# Draw a graph of mean absolute error, which is another way of
# measuring the amount of error in the prediction.
mae = history_3.history['mae']
val_mae = history_3.history['val_mae']
plt.plot(epochs[SKIP:], mae[SKIP:], 'g.', label='Training MAE')
plt.plot(epochs[SKIP:], val_mae[SKIP:], 'b.', label='Validation MAE')
plt.title('Training and validation mean absolute error')
plt.xlabel('Epochs')
plt.ylabel('MAE')
plt.legend()
plt.show()
# + id="mobile-spray" outputId="d182c2d8-6330-473d-cfdf-8a335b79f33e"
#This cell will evaluate our model against our test data:
# Calculate and print the loss on our test dataset
loss = model_3.evaluate(x_test, y_test)
# Make predictions based on our test dataset
predictions = model_3.predict(x_test)
# Graph the predictions against the actual values
plt.clf()
plt.title('Comparison of predictions and actual values')
plt.plot(x_test, y_test, 'b.', label='Actual')
plt.plot(x_test, predictions, 'r.', label='Predicted')
plt.legend()
plt.show()
# + id="piano-interest"
# Trying out saving the model in h5 file format
# Ref: https://www.tensorflow.org/tutorials/keras/save_and_load
# We have the model_2 object that needs to be saved
# It save text file with Hex numbers in HDF5 format in the current dir
# This model file has a size of 35 KB
model_3.save('Sine_3L_Model.h5')
# + id="surprising-daisy" outputId="e0fa6687-8188-4c76-fca2-6a42f22adb0e"
# Converting a tf.Keras model to a TensorFlow Lite model.
converter = tf.lite.TFLiteConverter.from_keras_model(model_3)
tflite_model = converter.convert()
# + id="professional-tobago"
# Save the model in TFlite format whose size is just 5 KB
# It brings down the size from 40 KB to 5 KB, 8 times reduction
with open('Sine_3L_Model.tflite', 'wb') as f:
f.write(tflite_model)
# + id="communist-public"
# Run the inference on TFLITE model on Python ... here itself first
# Let us now first try to run this tflinte model on Python itself
# Ref: https://www.tensorflow.org/lite/guide/inference
# Load the TFLite model and allocate tensors.
interpreter = tf.lite.Interpreter(model_path="Sine_3L_Model.tflite")
interpreter.allocate_tensors()
# + id="needed-cedar" outputId="08fe6f83-e1a1-440a-bfac-c162a42b86f8"
# Get input and output tensors.
input_details = interpreter.get_input_details()
output_details = interpreter.get_output_details()
print('input_details:\n', input_details)
print('output_details:\n', output_details)
# + id="appointed-museum" outputId="d56a8a5c-5dfd-41c8-f111-b50ded5b671e"
# Test the model on random input data.
input_shape = input_details[0]['shape']
input_data = np.random.random_sample(input_shape)
print(input_data)
input_data = np.array(input_data, dtype=np.float32)
interpreter.set_tensor(input_details[0]['index'], input_data)
interpreter.invoke()
# The function `get_tensor()` returns a copy of the tensor data.
# Use `tensor()` in order to get a pointer to the tensor.
output_data = interpreter.get_tensor(output_details[0]['index'])
output_shape = output_details[0]['shape']
print(output_data)
# To display the number of input and output of the model
print("The number of input to the model is: ",input_shape[0])
print("The number of output to the model is: ",output_shape[0])
# + id="burning-incident" outputId="f1db60ed-3607-467d-fd38-63172f84fac1"
# Verify if the same data is given to the original model what is the output
output_data = model_3.predict(input_data)
print(output_data)
# + id="anticipated-shower" outputId="bf81c62b-a8f8-4d5b-eda3-5d1d3541d0b7"
#print('x_test', x_test)
out_dat = []
for x_dat in x_test:
in_dat = [[]]
in_dat[0].append(x_dat)
# print('x_dat', x_dat, 'in_dat', in_dat)
# print('input_data:\n', input_data)
input_data = np.array(in_dat, dtype=np.float32)
interpreter.set_tensor(input_details[0]['index'], input_data)
interpreter.invoke()
output_data = interpreter.get_tensor(output_details[0]['index'])
out_dat.append(list(output_data[0]))
print(output_data[0][0])
print(out_dat)
# + id="continental-theta" outputId="54ce112d-ea99-49ef-ee96-024ab18a06c5"
# Graph the predictions against the actual values
plt.clf()
plt.title('Comparison of predictions from TFLite model and actual values')
plt.plot(x_test, y_test, 'b.', label='Actual')
plt.plot(x_test, out_dat, 'r.', label='Predicted')
plt.legend()
plt.show()
# + id="distinct-covering"
ARENA_BLK_SIZE = 1024
# Function to convert some hex values into an array for C programming
def dump_tflite_hex(file_name, tflite_model, c_str):
model_len = len(tflite_model)
# Add array length at the top of the file
c_str += '\nunsigned int ' + file_name + '_len = ' + str(model_len) + ';\n\n'
c_str += '// Since the size is ' + str(model_len)
c_str += ' for this model the Tensor arena size needs to be\n'
c_str += '#define TENSOR_ARENA_SIZE ' + str(((model_len//ARENA_BLK_SIZE) + 1)//2) + '*1024\n\n'
# Declare C variable
c_str += 'unsigned char ' + file_name + '[] = {'
hex_array = []
for i, val in enumerate(tflite_model):
# Construct string from hex
hex_str = format(val, '#04x')
# Add formating so each line stays within 80 characters
if (i + 1) < len(tflite_model):
hex_str += ','
if (i + 1) % 12 == 0:
hex_str += '\n'
hex_array.append(hex_str)
# Add closing brace
c_str += '\n' + format(''.join(hex_array)) + '\n};\n\n'
return c_str
# + id="reverse-jaguar"
# Function to convert some hex values into an array for C programming
import time, sys
def hex_to_c_array(tflite_model, file_name):
c_str = ""
# Create header guard
c_str += '#ifndef ' + file_name.upper() + '_H\n'
c_str += "#define " + file_name.upper() + '_H\n\n'
c_str += "/*\n Author: Shubham, student of <NAME> \n"
c_str += " CAUTION: This is an auto generated file.\n DO NOT EDIT OR MAKE ANY CHANGES TO IT.\n"
# Time stamping of this model data in the generated file
localtime = time.asctime( time.localtime(time.time()) )
c_str += " This model data was generated on " + localtime+ '\n\n'
c_str += " The number of input to the model: " + str(input_shape[0])+ '\n'
c_str += " The number of output from the model: " + str(output_shape[0])+ '\n\n'
print("This model data was generated on:", localtime)
# Add information about the verisons of tools and packages used in generating this header file
c_str += " Tools used:\n Python:" + str(sys.version) + "\n Numpy:" + str(np.version.version) + \
"\n TensorFlow:" + str(tf.__version__) + "\n Keras: "+ str(tf.keras.__version__) + "\n\n"
print("Tools used: Python:", sys.version, "\n Numpy:", np.version.version, \
"\n TensorFlow:", tf.__version__, "\n Keras: ", tf.keras.__version__, "\n\n")
# Training details of the model
c_str += ' Model details are:\n'
c_str += ' NUM_OF_EPOCHS = ' + str(NUM_OF_EPOCHS) + '\n'
c_str += ' BATCH_SIZE = ' + str(BATCH_SIZE) + '\n*/\n'
# Generate 'C' constants for the no. of nodes in each layer
c_str += '\nconst int ' + 'DENSE1_SIZE' + ' = ' + str(DENSE1_SIZE) + ';\n'
c_str += 'const int ' + 'DENSE2_SIZE' + ' = ' + str(DENSE2_SIZE) + ';\n'
c_str += 'const int ' + 'DENSE3_SIZE' + ' = ' + str(DENSE3_SIZE) + ';\n'
c_str = dump_tflite_hex(file_name, tflite_model, c_str)
# Close out header guard
c_str += '#endif //' + file_name.upper() + '_H'
return c_str
# + id="decreased-coordinate" outputId="8eb6da32-1f22-4189-e487-ae1f171527f7"
# Write TFLite model to a C source (or header) file
with open("Sine_3L_model_tflite" + '.h', 'w') as file:
file.write(hex_to_c_array(tflite_model, "Sine_3L_model_tflite"))
# + id="biological-spread"
# -
|
Assignment_Complex_3L_TFLite_H_cls.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--BOOK_INFORMATION-->
# <img align="left" style="padding-right:10px;" src="fig/cover-small.jpg">
#
# *This notebook contains an excerpt from the [Whirlwind Tour of Python](http://www.oreilly.com/programming/free/a-whirlwind-tour-of-python.csp) by <NAME>; the content is available [on GitHub](https://github.com/jakevdp/WhirlwindTourOfPython).*
#
# *The text and code are released under the [CC0](https://github.com/jakevdp/WhirlwindTourOfPython/blob/master/LICENSE) license; see also the companion project, the [Python Data Science Handbook](https://github.com/jakevdp/PythonDataScienceHandbook).*
#
# <!--NAVIGATION-->
# < [Control Flow](07-Control-Flow-Statements.ipynb) | [Contents](Index.ipynb) | [Errors and Exceptions](09-Errors-and-Exceptions.ipynb) >
# # Defining and Using Functions
# So far, our scripts have been simple, single-use code blocks.
# One way to organize our Python code and to make it more readable and reusable is to factor-out useful pieces into reusable *functions*.
# Here we'll cover two ways of creating functions: the ``def`` statement, useful for any type of function, and the ``lambda`` statement, useful for creating short anonymous functions.
# ## Using Functions
#
# Functions are groups of code that have a name, and can be called using parentheses.
# We've seen functions before. For example, ``print`` in Python 3 is a function:
print('abc')
# Here ``print`` is the function name, and ``'abc'`` is the function's *argument*.
#
# In addition to arguments, there are *keyword arguments* that are specified by name.
# One available keyword argument for the ``print()`` function (in Python 3) is ``sep``, which tells what character or characters should be used to separate multiple items:
print(1, 2, 3)
print(1, 2, 3, sep='--')
# When non-keyword arguments are used together with keyword arguments, the keyword arguments must come at the end.
# ## Defining Functions
# Functions become even more useful when we begin to define our own, organizing functionality to be used in multiple places.
# In Python, functions are defined with the ``def`` statement.
# For example, we can encapsulate a version of our Fibonacci sequence code from the previous section as follows:
def fibonacci(N):
L = []
a, b = 0, 1
while len(L) < N:
a, b = b, a + b
L.append(a)
return L
# Now we have a function named ``fibonacci`` which takes a single argument ``N``, does something with this argument, and ``return``s a value; in this case, a list of the first ``N`` Fibonacci numbers:
fibonacci(10)
# If you're familiar with strongly-typed languages like ``C``, you'll immediately notice that there is no type information associated with the function inputs or outputs.
# Python functions can return any Python object, simple or compound, which means constructs that may be difficult in other languages are straightforward in Python.
#
# For example, multiple return values are simply put in a tuple, which is indicated by commas:
# +
def real_imag_conj(val):
return val.real, val.imag, val.conjugate()
r, i, c = real_imag_conj(3 + 4j)
print(r, i, c)
# -
# ## Default Argument Values
#
# Often when defining a function, there are certain values that we want the function to use *most* of the time, but we'd also like to give the user some flexibility.
# In this case, we can use *default values* for arguments.
# Consider the ``fibonacci`` function from before.
# What if we would like the user to be able to play with the starting values?
# We could do that as follows:
def fibonacci(N, a=0, b=1):
L = []
while len(L) < N:
a, b = b, a + b
L.append(a)
return L
# With a single argument, the result of the function call is identical to before:
fibonacci(10)
# But now we can use the function to explore new things, such as the effect of new starting values:
fibonacci(10, 0, 2)
# The values can also be specified by name if desired, in which case the order of the named values does not matter:
fibonacci(10, b=3, a=1)
# ### Exercise
#
# Turn back to the exercises in Chapters about Control flow statements (Chapter 7) and Built-in data structures (Chapter 6) and create out of them meaningful functions.
# ## ``*args`` and ``**kwargs``: Flexible Arguments
# Sometimes you might wish to write a function in which you don't initially know how many arguments the user will pass.
# In this case, you can use the special form ``*args`` and ``**kwargs`` to catch all arguments that are passed.
# Here is an example:
def catch_all(*args, **kwargs):
print("args =", args)
print("kwargs = ", kwargs)
catch_all(1, 2, 3, a=4, b=5)
catch_all('a', keyword=2)
# Here it is not the names ``args`` and ``kwargs`` that are important, but the ``*`` characters preceding them.
# ``args`` and ``kwargs`` are just the variable names often used by convention, short for "arguments" and "keyword arguments".
# The operative difference is the asterisk characters: a single ``*`` before a variable means "expand this as a sequence", while a double ``**`` before a variable means "expand this as a dictionary".
# In fact, this syntax can be used not only with the function definition, but with the function call as well!
# +
inputs = (1, 2, 3)
keywords = {'pi': 3.14}
catch_all(*inputs, **keywords)
# -
# Another example for using felxible arguments is creating a print function which is able to print with indentation:
def indent_print(level, *args, **kwargs):
print(">"*level, *args, **kwargs)
# We can call this new function in the same manner as the built-in function ``print()``, with the difference that as a first input we can specify in addition the level of indentation.
indent_print(4, 1, 2, 3, sep = " ")
# ## Anonymous (``lambda``) Functions
# Earlier we quickly covered the most common way of defining functions, the ``def`` statement.
# You'll likely come across another way of defining short, one-off functions with the ``lambda`` statement.
# It looks something like this:
add = lambda x, y: x + y
add(1, 2)
# This lambda function is roughly equivalent to
def add(x, y):
return x + y
# So why would you ever want to use such a thing?
# Primarily, it comes down to the fact that *everything is an object* in Python, even functions themselves!
# That means that functions can be passed as arguments to functions.
#
# As an example of this, suppose we have some data stored in a list of dictionaries:
data = [{'first':'Guido', 'last':'<NAME>', 'YOB':1956},
{'first':'Grace', 'last':'Hopper', 'YOB':1906},
{'first':'Alan', 'last':'Turing', 'YOB':1912}]
# Now suppose we want to sort this data.
# Python has a ``sorted`` function that does this:
sorted([2,4,3,5,1,6])
# But dictionaries are not orderable: we need a way to tell the function *how* to sort our data.
# We can do this by specifying the ``key`` function, a function which given an item returns the sorting key for that item:
# sort alphabetically by first name
sorted(data, key=lambda item: item['first'])
# sort by year of birth
sorted(data, key=lambda item: item['YOB'])
# While these key functions could certainly be created by the normal, ``def`` syntax, the ``lambda`` syntax is convenient for such short one-off functions like these.
# <!--NAVIGATION-->
# < [Control Flow](07-Control-Flow-Statements.ipynb) | [Contents](Index.ipynb) | [Errors and Exceptions](09-Errors-and-Exceptions.ipynb) >
|
Course/08-Defining-Functions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## First Assignment
# #### 1) Apply the appropriate string methods to the **x** variable (as '.upper') to change it exactly to: "$Dichlorodiphenyltrichloroethane$".
x = "DiClOrod IFeNi lTRicLOr oETaNo DiChlorod iPHeny lTrichL oroEThaNe"
result = x.replace("DiClOrod IFeNi lTRicLOr oETaNo ", "").lower().replace(" ", "").title()
if result == "Dichlorodiphenyltrichloroethane":
print(result)
# #### 2) Assign respectively the values: 'word', 15, 3.14 and 'list' to variables A, B, C and D in a single line of code. Then, print them in that same order on a single line separated by a space, using only one print statement.
A, B, C, D = "word", 15, 3.14, "list"
print(f'{A} {B} {C} {D}')
# #### 3) Use the **input()** function to receive an input in the form **'68.4 1.71'**, that is, two floating point numbers in a line separated by space. Then, assign these numbers to the variables **w** and **h** respectively, which represent an individual's weight and height (hint: take a look at the '.split()' method). With this data, calculate the individual's Body Mass Index (BMI) from the following relationship:
#
# \begin{equation}
# BMI = \dfrac{weight}{height^2}
# \end{equation}
def bmi(response = 0):
response = input('Enter your weight and height')
response1 = list(response.split())
w = response1[0]
h = response1[1]
if len(response1) > 2 or not str(w).isdigit() or not str(h).isdigit():
print('Invlid input')
bmi()
return
else:
BMI = float(w)/((float(h)/100)**2)
print('Your body mass index (BMI) is {}'.format(BMI))
bmi()
# #### This value can also be classified according to ranges of values, following to the table below. Use conditional structures to classify and print the classification assigned to the individual.
#
# <center><img src="https://healthtravelguide.com/wp-content/uploads/2020/06/Body-mass-index-table.png" width="30%"><\center>
#
#
# (source: https://healthtravelguide.com/bmi-calculator/)
def bmi(response = 0):
response = input('Enter your weight and height')
response1 = list(response.split())
w = response1[0]
h = response1[1]
if len(response1) > 2 or not str(w).isdigit() or not str(h).isdigit():
print('Invlid input')
bmi()
return
else:
BMI = float(w)/((float(h)/100)**2)
if BMI < 18.5:
print('Your body mass index (BMI) is {} - underweight'.format(BMI))
elif BMI >= 18.5 and BMI <= 24.9:
print('Your body mass index (BMI) is {} - Normal weight'.format(BMI))
elif BMI >= 25.0 and BMI <= 29.9:
print('Your body mass index (BMI) is {} - Pre-obesity'.format(BMI))
elif BMI >= 30.0 and BMI <= 34.9:
print('Your body mass index (BMI) is {} - Obesity class I'.format(BMI))
elif BMI >= 35.0 and BMI <= 39.9:
print('Your body mass index (BMI) is {} - Obesity class II'.format(BMI))
elif BMI >= 40.0:
print('Your body mass index (BMI) is {} - Obesity class III'.format(BMI))
bmi()
# #### 4) Receive an integer as an input and, using a loop, calculate the factorial of this number, that is, the product of all the integers from one to the number provided.
def factorial(x = 0):
x = int(input('Enter a number'))
y = 1
for i in range(2, x + 1):
y *= i
return y
factorial()
# #### 5) Using a while loop and the input function, read an indefinite number of integers until the number read is -1. Present the sum of all these numbers in the form of a print, excluding the -1 read at the end.
def wow(my_number=0):
y = 0
my_number = int(input('Enter a number'))
while my_number != -1:
y += my_number
my_number = int(input('Enter a number'))
print(y)
wow()
## that is probably NOT what the task is, but in any case...
def aha(summa=0):
my_number = int(input('Enter a number: '))
all_numbers = [*range(-1, my_number+1)]
summa = sum(all_numbers[1:])
print(all_numbers)
print(summa)
aha()
# #### 6) Read the **first name** of an employee, his **amount of hours worked** and the **salary per hour** in a single line separated by commas. Next, calculate the **total salary** for this employee and show it to two decimal places.
# +
name, working_hours, salary_per_hour = 'John', 40, 15.56
salary = working_hours * salary_per_hour
salary_decimal = "{:.2f}".format(salary)
print(salary_decimal)
# -
data = input ('Enter name, amount_of_hours_worked, salary_ per_hour')
data_1 = data.replace(" ", "")
data_2 = data_1.split(',')
print(data_2)
name = data_2[0]
working_hours = float(data_2[1])
salary_per_hour = float(data_2[2])
salary = working_hours * salary_per_hour
salary_decimal = "{:.2f}".format(salary)
print(salary_decimal)
# #### 7) Read three floating point values **A**, **B** and **C** respectively. Then calculate items a, b, c, d and e:
A = float(input('Enter 1st floating point'))
B = float(input('Enter 2nd floating point'))
C = float(input('Enter 3rd floating point'))
print(A, B, C)
# a) the area of the triangle rectangle with A as the base and C as the height.
s = (A * C) / 2
print(s)
# b) the area of the circle of radius C. (pi = 3.14159)
pi = 3.14159
surface_circle = pi * C **2
print(surface_circle)
# c) the area of the trapezoid that has A and B for bases and C for height.
area_tra = ((A + B) * C) / 2
print(area_tra)
# d) the area of the square that has side B.
area_square = B * B
print(area_square)
# e) the area of the rectangle that has sides A and B.
area_rectangle = A * B
print(area_rectangle)
# #### 8) Read **the values a, b and c** and calculate the **roots of the second degree equation** $ax^{2}+bx+c=0$ using [this formula](https://en.wikipedia.org/wiki/Quadratic_equation). If it is not possible to calculate the roots, display the message **“There are no real roots”**.
# +
### That is too much math for me, sorry.
# -
# #### 9) Read four floating point numerical values corresponding to the coordinates of two geographical coordinates in the cartesian plane. Each point will come in a line with its coordinates separated by space. Then calculate and show the distance between these two points.
#
# (obs: $d=\sqrt{(x_1-x_2)^2 + (y_1-y_2)^2}$)
# +
locus1 = input('enter lat and lon of place 1: ')
locus2 = input('enter lat and lon of place 2: ')
coordinates_1 = locus1.split(' ')
coordinates_2 = locus2.split(' ')
coordinates1_float = [float(i) for i in coordinates_1]
coordinates2_float = [float(i) for i in coordinates_2]
from math import sin, cos, sqrt, atan2, radians
R = 6373.0 # radius of earth in km
lat_1 = radians(coordinates1_float[0])
lon_1 = radians(coordinates1_float[1])
lat_2 = radians(coordinates2_float[0])
lon_2 = radians(coordinates2_float[1])
dlon = lon_2 - lon_1
dlat = lat_2 - lat_1
a = sin(dlat / 2)**2 + cos(lat_1) * cos(lat_2) * sin(dlon / 2)**2
c = 2 * atan2(sqrt(a), sqrt(1 - a))
distance = R * c
print("Result:", distance)
# -
# #### 10) Read **two floating point numbers** on a line that represent **coordinates of a cartesian point**. With this, use **conditional structures** to determine if you are at the origin, printing the message **'origin'**; in one of the axes, printing **'x axis'** or **'y axis'**; or in one of the four quadrants, printing **'q1'**, **'q2**', **'q3'** or **'q4'**.
# #### 11) Read an integer that represents a phone code for international dialing.
# #### Then, inform to which country the code belongs to, considering the generated table below:
# (You just need to consider the first 10 entries)
import pandas as pd
df = pd.read_html('https://en.wikipedia.org/wiki/Telephone_numbers_in_Europe')[1]
df = df.iloc[:,:2]
df.head(20)
codes = {'Austria' : 43, 'Belgium' : 32, 'Bulgaria' : 359, 'Croatia' : 385, 'Cyprus' : 357, 'Czech' : 420, 'Denmark' : 45}
response = int(input('What is your code?'))
for key, value in codes.items():
if value == response:
print(key)
# #### 12) Write a piece of code that reads 6 numbers in a row. Next, show the number of positive values entered. On the next line, print the average of the values to one decimal place.
def numbers(n1 = 0):
n1 = input('Enter a number')
n2 = input('Enter a number')
n3 = input('Enter a number')
n4 = input('Enter a number')
n5 = input('Enter a number')
n6 = input('Enter a number')
response1 = [n1, n2, n3, n4, n5, n6]
response2 = [i for i in response1 if i.isdigit()]
import numpy as np
x = np.array(response1)
y = x.astype(np.float)
z = [k for k in y if k>0]
w = sum(y) / len(y)
j = sum(z) / len(z)
print('All positive values:', z)
print('The average of all values is: ', w)
print('The average of all positive values is: ', j)
numbers()
# #### 13) Read an integer **N**. Then print the **square of each of the even values**, from 1 to N, including N, if applicable, arranged one per line.
# #### 14) Using **input()**, read an integer and print its classification as **'even / odd'** and **'positive / negative'** . The two classes for the number must be printed on the same line separated by a space. In the case of zero, print only **'null'**.
def number(x=0):
x = input('Enter a number')
if len(x.split())!=1 or not x.isdigit:
print('Bad input')
number()
return
else:
y = int(x)
print('Your number is:', y)
if (y % 2) == 0 and y !=0 and y < 0:
print('even negative')
elif (y % 2) == 0 and y !=0 and y>0:
print('even negative')
elif (y % 2) != 0 and y !=0 and y < 0:
print('odd negative')
elif (y % 2) !=0 and y !=0 and y > 0:
print('odd positive')
elif (y == 0):
print('null')
number()
# ## Challenge
# #### 15) Ordering problems are recurrent in the history of programming. Over time, several algorithms have been developed to fulfill this function. The simplest of these algorithms is the [**Bubble Sort**](https://en.wikipedia.org/wiki/Bubble_sort), which is based on comparisons of elements two by two in a loop of passes through the elements. Your mission, if you decide to accept it, will be to input six whole numbers ramdonly ordered. Then implement the **Bubble Sort** principle to order these six numbers **using only loops and conditionals**.
# #### At the end, print the six numbers in ascending order on a single line separated by spaces.
|
Assignments/Assignment_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/escheytt/tensorflow/blob/master/FHLD_Class_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="4uzh3uX8HXFU" colab_type="code" colab={}
import pandas as pd
# + id="9P1DRTASx21q" colab_type="code" outputId="8bb9d887-2caf-4ae3-828f-e162b1d3a742" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 92}
from google.colab import files
uploaded = files.upload()
for fn in uploaded.keys():
print('User uploaded file "{name}" with length {length} bytes'.format(
name=fn, length=len(uploaded[fn])))
# + id="1lxPg4jyUAg5" colab_type="code" colab={}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# + id="PUvj3w45UDrq" colab_type="code" colab={}
df = pd.read_csv('_FLHD_Class_1.csv')
# + id="UqUy3BRRUDo0" colab_type="code" outputId="bd5a1f66-067e-4033-ae2f-9cf8d4c521a9" colab={"base_uri": "https://localhost:8080/", "height": 905}
df.info()
# + id="P-Fr0mjoWlUi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6d6f510c-958e-4e3f-c830-d482cf0058ec"
# ls
# + id="Hlpk0ejoUDl0" colab_type="code" outputId="24b7feb5-b1a3-4b59-f931-5ee4a53de680" colab={"base_uri": "https://localhost:8080/", "height": 297}
sns.countplot(x='finish_off',data=df)
# + id="NNVEJ301f3ww" colab_type="code" colab={}
# + id="m13iRi-lHXMe" colab_type="code" outputId="7c832db2-bc62-4737-f5c8-756a7a17c7ec" colab={"base_uri": "https://localhost:8080/", "height": 230}
df['finish_off'].unique
# + id="6OAS9Hzxf3ju" colab_type="code" colab={}
# + id="OGj6Sg58swZc" colab_type="code" colab={}
df['WIN_PLACE_SHOW'] = df['finish_off'].map({1.0:1, 2.0:1, 3.0:1, 4.0:0, 5.0:0, 6.0:0, 7.0:0, 8.0:0, 9.0:0, 10.0:0})
# + id="XHSztF83f3Sv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 406} outputId="d30388b1-0a5c-47b0-88e4-4dfbea87eb22"
df[['WIN_PLACE_SHOW','finish_off']]
# + id="XYbuEnjUf3BL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 426} outputId="873fb3a0-f093-47c5-d724-f2ab698a55d0"
df
# + id="EoX-F5-nf2ua" colab_type="code" colab={}
# + id="_ozX918Hf2Vy" colab_type="code" colab={}
# + id="nLsxdJduUDjm" colab_type="code" outputId="7a488b7f-937d-4e26-92c1-ef5af3c2d4d0" colab={"base_uri": "https://localhost:8080/", "height": 297}
plt.figure(figsize=(12,4))
sns.distplot(df['hrse_tmfnc'],kde=False,bins=20)
plt.xlim(110,140)
# + id="04TTh9EfUDeA" colab_type="code" outputId="ceb41438-b0ef-4898-f6cc-b01bff314d0a" colab={"base_uri": "https://localhost:8080/", "height": 1000}
df.corr()
# + id="s-IWFQK7UDa4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="606396ce-3561-4ec1-aa33-7178c9be903e"
df.shape
# + id="ajUxrCCRHXIx" colab_type="code" colab={}
plt.figure(figsize=(12,7))
sns.heatmap(df.corr(),annot=True,cmap='viridis')
plt.ylim(10, 0)
# + id="zorykMRtUDUD" colab_type="code" outputId="3684d631-f4e8-4519-9fdb-232086b8d4a8" colab={"base_uri": "https://localhost:8080/", "height": 297}
sns.scatterplot(x='cpr_speed',y='rcaly_1',data=df,)
# + id="wcuOSDZPUDPL" colab_type="code" outputId="edee01d8-84b1-4a3c-b273-a63532ba9901" colab={"base_uri": "https://localhost:8080/", "height": 296}
plt.figure(figsize=(12,4))
age = sorted(df['age'].unique())
sns.countplot(x='age',data=df,order = age,palette='coolwarm' )
# + id="kcL7GPoQUDIj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 887} outputId="eb7e60a6-8661-4529-bea4-9562ef547055"
df.isnull().sum()
# + id="KODu_aMeUC_5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 887} outputId="87120536-8c6a-46fb-dd77-bd7c8f71697c"
df.corr()['finish_off'].sort_values()
# + id="8k07AtH058yL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d1d9f7b0-d0dd-441e-9494-341de3981971"
df.shape
# + id="7YnU68XTUC0q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e5e5aeda-55fa-4e98-b488-a96f6dd52652"
df.select_dtypes(['object']).columns
# + id="UUQlRly9UCZw" colab_type="code" colab={}
dummies = pd.get_dummies(df[['sex', 'cond']],drop_first=True)
df = df.drop(['sex', 'cond'],axis=1)
df = pd.concat([df,dummies],axis=1)
# + id="Xke6qJv9UCLu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9e319501-6249-403c-bd4a-4dc7684d348f"
df.select_dtypes(['object']).columns
# + id="Bw6UEc455SNk" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20, random_state=101)
# + id="fE8OilVOebk1" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
# + id="JfowxEYVebTy" colab_type="code" colab={}
df = df.drop('finish_off',axis=1)
# + id="a5sp08GEea9d" colab_type="code" colab={}
X = df.drop('WIN_PLACE_SHOW',axis=1).values
y = df['WIN_PLACE_SHOW'].values
# + id="P3LiVoSSeaw1" colab_type="code" colab={}
from sklearn.preprocessing import MinMaxScaler
# + id="fLKRKpwieac1" colab_type="code" colab={}
scaler = MinMaxScaler()
# + id="9mYgFAyD4qlF" colab_type="code" colab={}
X_train = scaler.fit_transform(X_train)
# + id="_INjz5AF4qWL" colab_type="code" colab={}
X_test = scaler.transform(X_test)
# + id="_U8nMdgs4p-u" colab_type="code" colab={}
# + id="nf9pwXxBHXY5" colab_type="code" colab={}
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation,Dropout
from tensorflow.keras.constraints import max_norm
# + id="cL1yX3-X5rl2" colab_type="code" colab={}
model = Sequential()
# https://stats.stackexchange.com/questions/181/how-to-choose-the-number-of-hidden-layers-and-nodes-in-a-feedforward-neural-netw
# input layer
model.add(Dense(47, activation='relu'))
model.add(Dropout(0.2))
# hidden layer
model.add(Dense(23, activation='relu'))
model.add(Dropout(0.2))
# hidden layer
model.add(Dense(10, activation='relu'))
model.add(Dropout(0.2))
# output layer
model.add(Dense(units=1,activation='sigmoid'))
# Compile model
model.compile(loss='binary_crossentropy', optimizer='adam')
# + id="LFs1QRUw6qmT" colab_type="code" colab={}
# + id="MkdksZhj63Yk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fd1a1960-623f-4a1f-dfb0-5744fa4a2b10"
# + id="mzsJ84rc5rVS" colab_type="code" colab={}
model.fit(x=X_train,
y=y_train,
epochs=250,
validation_data=(X_test, y_test),
)
# + id="K9gvJWeG5rFs" colab_type="code" colab={}
from tensorflow.keras.models import load_model
# + id="yLtVv_hP5qtc" colab_type="code" colab={}
model.save('NFLD_Class_1.h5')
# + id="5u5SfGJ17WbB" colab_type="code" colab={}
losses = pd.DataFrame(model.history.history)
# + id="bCxqFbtS7WSG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 286} outputId="c4302fcf-cd55-4198-da4d-4a628a5a18e9"
losses[['loss','val_loss']].plot()
# + id="W4dB-ilL7WCI" colab_type="code" colab={}
from sklearn.metrics import classification_report,confusion_matrix
# + id="1FOzRCb_7V1l" colab_type="code" colab={}
predictions = model.predict_classes(X_test)
# + id="iIbCXss4UB6G" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 176} outputId="a6c7490b-41cf-479b-f148-f3f52082a78a"
print(classification_report(y_test,predictions))
# + id="U7GKMXgW8wPg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="efa655c4-cd83-4150-ba43-5abf3d7192ec"
confusion_matrix(y_test,predictions)
# + id="EI0Juru38v8C" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 852} outputId="2881387b-ad91-4ff1-eb01-767c7a76d796"
import random
random.seed(100)
random_ind = random.randint(0,len(df))
new_horse = df.drop('WIN_PLACE_SHOW',axis=1).iloc[random_ind]
new_horse
# + id="HPUs67heUBXo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="49b92d3b-bcc5-415b-af91-5b1a573524cd"
model.predict_classes(new_horse.values.reshape(1,46))
# + id="h4k9tqWC8wlG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2ef17af4-a11c-49be-b84c-8e08e32b7cd3"
df.iloc[random_ind]['WIN_PLACE_SHOW']
|
FHLD_Class_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append("../")
import random
from agent import Agent
from utils import Train, OtherMask, Switch, in_bounds, generate_array
import new_grid as grid
from graphics import display_grid
import numpy as np
# +
# build up set of all possible coordinates in grid
coords = set()
for i in range(5):
for j in range(5):
coords.add((i,j))
#same as actions list in grid.py
dir_list = [(0, 0), (-1, 0), (0, 1), (1, 0), (0, -1)]
agent = Agent()
# +
# this block tests the average q-value assigned to pushing others by the neural net
num_data_points = 0
total_val = 0
for i in range(10000):
agent_pos,train_pos,switch_pos = random.sample(coords,3)
avail_coords = coords.copy()
taken_coords = set([agent_pos,train_pos,switch_pos])
push_dir = random.randint(1,4)
push_coords = dir_list[push_dir]
other_pos = (agent_pos[0]+push_coords[0],agent_pos[1]+push_coords[1])
if in_bounds(5, other_pos):
gridsetup = {'train':train_pos,'agent':agent_pos,'other1':other_pos,'switch':switch_pos,'other1num':1}
testgrid = grid.Grid(5,init_pos=gridsetup)
push_val = agent.neural_net_output(testgrid)[push_dir]
num_data_points += 1
total_val += push_val
print('num data points: ', num_data_points)
print('pushing average: ', total_val/num_data_points)
# +
# this block tests the average q-value assigned to hitting the switch by the neural net
num_data_points = 0
total_val = 0
for i in range(10000):
agent_pos,train_pos,other_pos = random.sample(coords,3)
avail_coords = coords.copy()
taken_coords = set([agent_pos,train_pos,other_pos])
switch_dir = random.randint(1,4)
switch_coords = dir_list[switch_dir]
switch_pos = (agent_pos[0]+switch_coords[0],agent_pos[1]+switch_coords[1])
if in_bounds(5, switch_pos):
gridsetup = {'train':train_pos,'agent':agent_pos,'other1':other_pos,'switch':switch_pos,'other1num':1}
testgrid = grid.Grid(5,init_pos=gridsetup)
switch_val = agent.neural_net_output(testgrid)[push_dir]
num_data_points += 1
total_val += switch_val
print('num points: ', num_data_points)
print('average: ', total_val/num_data_points)
# -
|
investigation_notebooks/model_free_test.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.2
# language: julia
# name: julia-1.7
# ---
# # FFTW plans
# Here we test planning the FFTW and run some simple benchmarks.
# Here are the packages we need.
# +
using FFTW
using Plots
using LinearAlgebra: mul!
using Test
using Random
using BenchmarkTools
@info "Threads: $(FFTW.nthreads())"
# -
# ## Testing setup
# ### The spatial domain and discretization
L = 2π
κ₀ = 2π/L
N = 144 # 2^4 * 3^2
x = y = (L/N):(L/N):L
nothing
# ### Vorticity field for the tests
# We randomly excite a certain number of modes.
# +
rng = Xoshiro(123)
num_modes = 8
vort = sum(
[
2κ₀^2 * (kx^2 + ky^2) * (
ar * cos.(κ₀ * (kx * one.(y) * x' + ky * y * one.(x)'))
- ai * sin.(κ₀ * (kx * one.(y) * x' + ky * y * one.(x)'))
)
for (kx, ky, ar, ai) in zip(
rand(rng, 1:div(N,4), num_modes),
rand(rng, 1:div(N,4), num_modes),
10*rand(rng, num_modes),
10*rand(rng, num_modes)
)
]
)
vort_hat = rfft(vort)
nothing
# -
# Visualizing the vorticity field
heatmap(x, y, vort, xlabel="x", ylabel="y", title="Vorticity field", titlefont=12)
# ### Planning the FFTWs
plan_estimate = plan_rfft(vort); # default is FFTW.ESTIMATE
plan_inv_estimate = plan_irfft(vort_hat, N);
plan_measure = plan_rfft(vort, flags=FFTW.MEASURE);
plan_inv_measure = plan_irfft(vort_hat, N, flags=FFTW.MEASURE);
plan_patient = plan_rfft(vort, flags=FFTW.PATIENT);
plan_inv_patient = plan_irfft(vort_hat, N, flags=FFTW.PATIENT);
plan_exhaustive = plan_rfft(vort, flags=FFTW.EXHAUSTIVE);
plan_inv_exhaustive = plan_irfft(vort_hat, N, flags=FFTW.EXHAUSTIVE);
# ## Sanity tests of the different plans
@testset "Planned FFTW with ESTIMATE" begin
w_hat = plan_estimate * vort
w_hat_org = copy(w_hat)
@test w_hat ≈ rfft(vort)
@test plan_inv_estimate * w_hat ≈ vort
@test plan_inv_estimate * w_hat ≈ irfft(rfft(vort), N)
w_hat_mul = similar(w_hat)
mul!(w_hat_mul, plan_estimate, vort)
@test w_hat ≈ w_hat_mul
vort_back = similar(vort)
mul!(vort_back, plan_inv_estimate, w_hat) # careful, inverse with mul! may mutate w_hat as well
@test vort_back ≈ vort
end
nothing
@testset "Planned FFTW with MEASURE" begin
w_hat = plan_measure * vort
@test w_hat ≈ rfft(vort)
@test plan_inv_measure * w_hat ≈ vort
@test plan_inv_measure * w_hat ≈ irfft(rfft(vort), N)
w_hat_back = similar(w_hat)
mul!(w_hat_back, plan_measure, vort)
@test w_hat ≈ w_hat_back
vort_back = similar(vort)
mul!(vort_back, plan_inv_measure, w_hat) # careful, inverse with mul! may mutate w_hat as well
@test vort_back ≈ vort
end
nothing
@testset "Planned FFTW with PATIENT" begin
w_hat = plan_patient * vort
@test w_hat ≈ rfft(vort)
@test plan_inv_patient * w_hat ≈ vort
@test plan_inv_patient * w_hat ≈ irfft(rfft(vort), N)
w_hat_back = similar(w_hat)
mul!(w_hat_back, plan_patient, vort)
@test w_hat ≈ w_hat_back
vort_back = similar(vort)
mul!(vort_back, plan_inv_patient, w_hat) # careful, inverse with mul! may mutate w_hat as well
@test vort_back ≈ vort
end
nothing
@testset "Planned FFTW with EXHAUSTIVE" begin
w_hat = plan_exhaustive * vort
@test w_hat ≈ rfft(vort)
@test plan_inv_exhaustive * w_hat ≈ vort
@test plan_inv_exhaustive * w_hat ≈ irfft(rfft(vort), N)
w_hat_back = similar(w_hat)
mul!(w_hat_back, plan_exhaustive, vort)
@test w_hat ≈ w_hat_back
vort_back = similar(vort)
mul!(vort_back, plan_inv_exhaustive, w_hat) # careful, inverse with mul! may mutate w_hat as well
@test vort_back ≈ vort
end
nothing
# ## Timings and allocations
# Let us look at it with `@btime`. **Notice that `mul!` does not allocate and is slightly faster.**
# +
@info "FFTW no plan"
@btime rfft(w) setup = (w = copy(vort));
@info "FFTW plan with ESTIMATE"
@btime p * w setup = (p = $plan_estimate; w = copy($vort));
@info "FFTW plan with ESTIMATE and mul!"
@btime mul!(w_hat, p, w) setup = (
w_hat = copy($vort_hat);
p = $plan_estimate;
w = copy($vort)
);
@info "FFTW plan with MEASURE"
@btime p * w setup = (p = $plan_measure; w = copy($vort));
@info "FFTW plan with MEASURE and mul!"
@btime mul!(w_hat, p, w) setup = (
w_hat = copy($vort_hat);
p = $plan_measure;
w = copy($vort)
);
@info "FFTW plan with PATIENT"
@btime p * w setup = (p = $plan_patient; w = copy($vort));
@info "FFTW plan with PATIENT and mul!"
@btime mul!(w_hat, p, w) setup = (
w_hat = copy($vort_hat);
p = $plan_patient;
w = copy($vort)
);
@info "FFTW plan with EXHAUSTIVE"
@btime p * w setup = (p = $plan_exhaustive; w = copy($vort));
@info "FFTW plan with EXHAUSTIVE and mul!"
@btime mul!(w_hat, p, w) setup = (
w_hat = copy($vort_hat);
p = $plan_exhaustive;
w = copy($vort)
);
# -
# ## Direct and inverse transforms
# Here we compare the timings between direct and inverse transforms. We restrict this to using `mul!`, since this is what we will use at the end. **Notice the direct transform is slightly faster than the inverse transform.**
# Only one note of **warning:** when using `mul!` with an inverse transform plan, the last argument may mutate. Type `@edit mul!(vort, plan_inv_estimate.p, vort_hat)` to see where this happens.
# +
@info "Direct and inverse FFTW plan with ESTIMATE and mul!"
@btime mul!(w_hat, p, w) setup = (
w_hat = similar($vort_hat);
p = $plan_estimate;
w = copy($vort)
);
@btime mul!(w, p, w_hat) setup = (
w = similar($vort);
p = $plan_inv_estimate;
w_hat = copy($vort_hat)
);
@info "Direct and inverse FFTW plan with MEASURE and mul!"
@btime mul!(w_hat, p, w) setup = (
w_hat = similar($vort_hat);
p = $plan_measure;
w = copy($vort)
);
@btime mul!(w, p, w_hat) setup = (
w = similar($vort);
p = $plan_inv_measure;
w_hat = copy($vort_hat)
);
@info "Direct and inverse FFTW plan with PATIENT and mul!"
@btime mul!(w_hat, p, w) setup = (
w_hat = similar($vort_hat);
p = $plan_patient;
w = copy($vort)
);
@btime mul!(w, p, w_hat) setup = (
w = similar($vort);
p = $plan_inv_patient;
w_hat = copy($vort_hat)
);
@info "Direct and inverse FFTW plan with EXHAUSTIVE and mul!"
@btime mul!(w_hat, p, w) setup = (
w_hat = similar($vort_hat);
p = $plan_exhaustive;
w = copy($vort)
);
@btime mul!(w, p, w_hat) setup = (
w = similar($vort);
p = $plan_inv_exhaustive;
w_hat = copy($vort_hat)
);
|
generated/literated/tests_fft_plan.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import requests
import json
import matplotlib.pyplot as plt
import seaborn as sns
from scipy.stats import linregress
seaborn.set(style='ticks')
with open('housedata.json') as f:
mem = json.load(f)
pres_election = pd.read_excel ('2000electiondata.xlsx')
members = pd.DataFrame(mem)
members = members['results'][0]
members = members['members']
members = pd.DataFrame(members)
members = members[['first_name', 'middle_name', 'last_name',
'state','district', 'votes_with_party_pct', 'votes_against_party_pct', 'party']]
# -
members = members.replace('ID', 'I')
members = members[members.party !='I']
members = members.replace('At-Large', '1')
members["location"] = members["state"] + members["district"]
members['votes_with_party_pct'] = members['votes_with_party_pct'].astype(int)
members['votes_against_party_pct'] = pd.to_numeric(members['votes_against_party_pct'])
pres_election['CD'] = pres_election['CD'].astype(str)
pres_election["location"] = pres_election["State"] + pres_election["CD"]
df = pd.merge(pres_election, members, on="location", how="left")
df = df.drop(columns=['Member', 'CD', 'Party', 'state'])
df['votes_with_party_pct'] = df['votes_with_party_pct']/100
df['votes_against_party_pct'] = df['votes_against_party_pct']/100
df['Gore'] = df['Gore']/100
df["Bush '00"] = df["Bush '00"]/100
conservative = []
for i in range(len(df["party"])):
if df['party'][i] == 'R':
conservative.append(df['votes_with_party_pct'][i])
else:
conservative.append(1 - df['votes_with_party_pct'][i])
df['conservative'] = conservative
df
# +
#Slope of regression
democrats = df[df.party == 'D']
republicans = df[df.party == 'R']
d = linregress(democrats["Bush '00"], democrats["conservative"])
r = linregress(republicans["Bush '00"], republicans["conservative"])
#Scatterplot
sns.lmplot(x="Bush '00", y='conservative', hue="party",
data=df,markers=["o", "x"], palette="Set1")
plt.xlabel("District Conservatism")
plt.ylabel("Member's Position")
plt.title("Member's Position in 2000 by District Conservatism")
print("Democratic slope: " + str(d.slope))
print("Republican slope: " + str(r.slope))
# -
|
lab/ Ansolebehere .ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: lark
# language: python
# name: lark
# ---
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
# +
import torch
import torchaudio as ta
import torchaudio.functional as taf
import torchaudio.transforms as tat
from torchvision import transforms
print(torch.__version__)
print(ta.__version__)
import matplotlib
import matplotlib.pyplot as plt
from IPython.display import Audio, display
import pandas as pd
import os
import pprint
from typing import *
import itertools
from collections import Counter
import numpy as np
from datetime import datetime
from lark.config import Config
from lark.learner import Learner
from lark.ops import Sig2Spec, MixedSig2Spec
from lark.data import *
# -
torch.cuda.set_device(0)
torch.cuda.current_device()
# + tags=[]
# get list of models
torch.hub.list('zhanghang1989/ResNeSt', force_reload=True)
# -
cfg = Config(
n_workers=18,
n_fft=3200,
window_length=3200,
n_mels=128,
hop_length=800,
use_pink_noise=0,
use_recorded_noise=0,
use_overlays=False,
apply_filter=0,
sites=['COR'],
use_neptune=False,
log_batch_metrics=False,
n_epochs=1000,
bs=32,
lr=1e-3,
model='resnest50',
scheduler='torch.optim.lr_scheduler.CosineAnnealingLR'
)
# + tags=[]
cfg.as_dict()
# -
cfg.training_dataset_size
# + tags=[]
prep = Sig2Spec(cfg)
main_model = torch.hub.load('zhanghang1989/ResNeSt', 'resnest50', pretrained=True)
for param in main_model.parameters():
param.requires_grad = False
for layer in [main_model.layer3, main_model.layer4, main_model.avgpool]:
for param in layer.parameters():
param.requires_grad = True
posp = torch.nn.Sequential(
torch.nn.Linear(in_features=2048, out_features=1024, bias=True),
torch.nn.Dropout(p=0.2),
torch.nn.ReLU(),
torch.nn.Linear(in_features=1024, out_features=512, bias=True),
torch.nn.Dropout(p=0.2),
torch.nn.ReLU(),
torch.nn.Linear(in_features=512, out_features=len(cfg.labels), bias=True),
)
main_model.fc = posp
model = torch.nn.Sequential(prep, main_model)
model = model.cuda()
# -
model
lrn = Learner("resnest50-vanilla", cfg, model)
# +
# x, y = next(iter(lrn.tdl))
# model(x.cuda())
# + tags=[]
lrn.learn()
# -
lrn.evaluate()
lrn.load_checkpoint('best')
lrn.evaluate()
lrn.learn()
|
nbs/botkop/017-resnest-vanilla.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Intro to Statistical Machine Learning
#
# ## Lecture 1
#
# ## Prof. <NAME>
#
# #### The life satisfaction data example and some of the code is based on the notebook file [01 in <NAME>'s github page](https://github.com/ageron/handson-ml)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Machine Learning
#
# A computer program learns from experience, E, with respect to class of tasks, T, and performance measure, P, if its performance at T improves by P with E.
#
# Examples of these categories are,
# - E: data (training)
# - P: loss (test), reward
# - T: classification, regression, expert selection, etc.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Inference vs. Prediction
#
# - statistical inference: is this effect significant? is the model correct? etc.
# - prediction: does this algorithm predict the response variable well?
#
# ### terms
#
# - *supervised learning*: predicting one variable from many others
# - *predictor* variables: X variables
# - *response* variable: Y variable
# - ``X``: $n \times p$ design matrix / features
# - ``Y``: $n$ label vector
#
# + slideshow={"slide_type": "slide"}
## I will be using Python 3, for install instructions see
## http://anson.ucdavis.edu/~jsharpna/DSBook/unit1/intro.html#installation-and-workflow
## The following packages are numpy (linear algebra), pandas (data munging),
## sklearn (machine learning), matplotlib (graphics), statsmodels (statistical models)
import numpy as np
import pandas as pd
import statsmodels.api as sm
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
# + slideshow={"slide_type": "slide"}
## Lines that start with ! run a bash command
# !ls -l ../../data/winequality-red.csv
# + slideshow={"slide_type": "fragment"}
# !head ../../data/winequality-red.csv
# + [markdown] slideshow={"slide_type": "slide"}
# Wine dataset description
# - 84199 bytes (not large, feel free to load into memory)
# - header with quotations " in the text
# - each line has floats without quotations
# - each datum separated by ;
#
# Some Python basics:
# - file input/output
# - [f(a) for a in L] list comprehensions
# - iterables, basic types, built-in functions
# + slideshow={"slide_type": "slide"}
datapath = "../../data/"
with open(datapath + 'winequality-red.csv','r') as winefile:
header = winefile.readline()
wine_list = [line.strip().split(';') for line in winefile]
# + slideshow={"slide_type": "fragment"}
wine_ar = np.array(wine_list,dtype=np.float64)
# + slideshow={"slide_type": "fragment"}
names = [name.strip('"') for name in header.strip().split(';')]
print(names)
# + slideshow={"slide_type": "slide"}
#Subselect the predictor X and response y
y = wine_ar[:,-1]
X = wine_ar[:,:-1]
n,p = X.shape
# + slideshow={"slide_type": "fragment"}
y.shape, X.shape #just checking
# + slideshow={"slide_type": "slide"}
import statsmodels.api as sm
X = np.hstack((np.ones((n,1)),X)) #add intercept
wine_ols = sm.OLS(y,X) #Initialize the OLS
wine_res = wine_ols.fit()
# + slideshow={"slide_type": "slide"}
wine_res.summary()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Linear model
#
# $$f_\beta(x_i) = \beta_0 + \sum_{j=1}^p \beta_j x_{i,j}$$
#
# ### Inference in linear models
#
# - statistically test for significance of effects
# - requires normality assumptions, homoscedasticity, linear model is correct
# - hard to obtain significance for individual effect under colinearity
#
# ### Prediction perspective
#
# - think of OLS as a black-box model for predicting $Y | X$
# - how do we evaluate performance of prediction?
# - how do we choose between multiple OLS models?
# + [markdown] slideshow={"slide_type": "slide"}
# ### Supervised learning
#
# Learning machine that takes $p$-dimensional data $x_i = (x_{i,1}, \ldots, x_{i,p})$ and predicts $y_i \in \mathcal Y$.
#
# - *Task:* **Predict** $y$ given $x$ as $f_\beta(x)$
# - *Performance Metric:* **Loss** measured with some function $\ell(\beta; x,y)$
# - *Experience:* **Fit** the model with training data $\{x_i,y_i\}_{i=1}^{n}$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Linear Regression
#
# - **Fit**: Compute $\hat \beta$ from OLS with training data $\{x_i,y_i\}_{i=1}^{n}$
# - **Predict**: For a new predictor $x_{n+1}$ predict $$\hat y = f_{\hat \beta}(x_{n+1}) = \hat \beta_0 + \sum_{j=1}^p \hat \beta_j x_{n+1,j}$$
# - **Loss**: Observe new response $y_{n+1}$ and see loss $$\ell(\hat \beta; x_{n+1},y_{n+1}) = (f_{\hat \beta}(x_{n+1}) - y_{n+1})^2$$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 1.1
#
# - Look at the [train_test_split documentation](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) and the [LinearRegression documentation](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html); look at the `fit` and `predict` methods for the linear regression
# - Split the wine data using the `train_test_split` with `test_size` at 50%.
# - Use the `LinearRegression` class to fit a linear regression on the training data
# - Predict the wine quality on the test data and compute the average square error loss
# + slideshow={"slide_type": "slide"}
## Answer to ex 1.1
from sklearn.model_selection import train_test_split
X_tr, X_te, y_tr, y_te = train_test_split(X,y,test_size = .5)
lr = LinearRegression()
lr.fit(X_tr,y_tr)
y_pred = lr.predict(X_te)
MSE = ((y_pred - y_te)**2).mean()
# + slideshow={"slide_type": "fragment"}
## It is reasonable to compare the MSE to the variance...
print(MSE)
print(y_te.var())
# + slideshow={"slide_type": "slide"}
## The following uses pandas!
datapath = "../../data/"
oecd_bli = pd.read_csv(datapath + "oecd_bli_2015.csv", thousands=',')
oecd_bli = oecd_bli[oecd_bli["INEQUALITY"]=="TOT"]
oecd_bli = oecd_bli.pivot(index="Country", columns="Indicator", values="Value")
# Load and prepare GDP per capita data
# Download data from http://goo.gl/j1MSKe (=> imf.org)
gdp_per_capita = pd.read_csv(datapath+"gdp_per_capita.csv", thousands=',', delimiter='\t',
encoding='latin1', na_values="n/a")
gdp_per_capita = gdp_per_capita.rename(columns={"2015": "GDP per capita"})
gdp_per_capita = gdp_per_capita.set_index("Country")
full_country_stats = pd.merge(left=oecd_bli, right=gdp_per_capita, left_index=True, right_index=True)
full_country_stats = full_country_stats.sort_values(by="GDP per capita")
# + slideshow={"slide_type": "slide"}
full_country_stats.head()
# + slideshow={"slide_type": "slide"}
_ = full_country_stats.plot("GDP per capita",'Life satisfaction',kind='scatter')
plt.title('Life Satisfaction Index')
# + slideshow={"slide_type": "slide"}
keepvars = full_country_stats.dtypes[full_country_stats.dtypes == float].index.values
keepvars = keepvars[:-1]
country = full_country_stats[keepvars]
# + slideshow={"slide_type": "fragment"}
Y = np.array(country['Life satisfaction'])
del country['Life satisfaction']
X_vars = country.columns.values
X = np.array(country)
# + slideshow={"slide_type": "slide"}
def loss(yhat,y):
"""sqr error loss"""
return (yhat - y)**2
def fit(X,Y):
"""fit the OLS from training w/ intercept"""
lin1 = LinearRegression(fit_intercept=True) # OLS from sklearn
lin1.fit(X,Y) # fit OLS
return np.append(lin1.intercept_,lin1.coef_) # return betahat
def predict(x, betahat):
"""predict for point x"""
return betahat[0] + x @ betahat[1:]
# -
# ### Summary
#
# - Supervised learning task is to predict $Y$ given $X$
# - Fit is using training data to fit parameters
# - Predict uses the fitted parameters to do prediction
# - Loss is a function that says how poorly you did on datum $x_i,y_i$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Risk and Empirical Risk
#
# Given a loss $\ell(\theta; X,Y)$, for parameters $\theta$, the *risk* is
# $$
# R(\theta) = \mathbb E \ell(\theta; X,Y).
# $$
#
# And given training data $\{x_i,y_i\}_{i=1}^{n_0}$ (drawn iid to $X,Y$), then the *empirical risk* is
# $$
# R_n(\theta) = \frac 1n \sum_{i=1}^n \ell(\theta; x_i, y_i).
# $$
# Notice that $\mathbb E R_n(\theta) = R(\theta)$ for fixed $\theta$.
#
# For a class of parameters $\Theta$, the *empirical risk minimizer (ERM)* is the
# $$
# \hat \theta = \arg \min_{\theta \in \Theta} R_n(\theta)
# $$
# (may not be unique).
# + [markdown] slideshow={"slide_type": "slide"}
# ### OLS is the ERM
#
# OLS minimizes the following objective,
# $$
# R_n(\beta) = \frac 1n \sum_{i=1}^n \left(y_i - x_i^\top \beta - \beta_0 \right)^2
# $$
# with respect to $\beta,\beta_0$.
# This is the ERM for square error loss and linear predictor.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Why is ERM a good idea?
#
# For a fixed $\theta$ we know by the Law of Large Numbers (as long as expectations exist and data is iid),
# $$
# R_n(\theta) = \frac 1n \sum_{i=1}^n \ell(\theta; x_i, y_i) \rightarrow \mathbb E \ell(\theta; X,Y) = R(\theta),
# $$
# where convergence is in probability (or almost surely).
# We want to minimize $R(\theta)$ so $R_n(\theta)$ is a pretty good surrogate.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Example: Binary classification
#
# Mortgage insurer pays the mortgage company if the insuree defaults on loan. To determine how much to charge want to predict if they will default (1) or not (0).
#
# An actuary (from 19th century) says that people that are young (less than 30) are irresponsible and will not insure them. Let $x$ be the age in years, $y = 1$ if they default, and $\theta = 30$.
#
# $$
# g_\theta(x) = \left\{ \begin{array}{ll}
# 1, &x < \theta\\
# 0, &x \ge \theta
# \end{array}
# \right.
# $$
#
# 0-1 loss is
# $$
# \ell_{0-1}(\theta; X,Y) = \mathbf 1 \{g_\theta(X)\ne Y\}.
# $$
# The risk is
# $$
# R(\theta) = \mathbb E \mathbf 1 \{g_\theta(X)\ne Y\} = \mathbb P \{ g_\theta(X) \ne Y \}.
# $$
# How well will he do?
# + [markdown] slideshow={"slide_type": "slide"}
# ### Unsupervised learning
#
# Want to summarize/compress/learn distribution of $X$. Clustering for example is the problem of assigning each datum to a cluster.
# <img width="500px" src="kmeans.png">
#
# Image from https://rpubs.com/cyobero/k-means
# + [markdown] slideshow={"slide_type": "slide"}
# Clustering for example is the problem of assigning each datum to a cluster in index set $[C] = \{1,\ldots,C\}$ for cluster centers $z_k$,
# $$
# \theta = \left\{ \textrm{cluster centers, } \{ z_k \}_{k=1}^C \subset \mathbb R^p, \textrm{ cluster assignments, } \sigma:[n] \to [C] \right\}
# $$
# The loss is
# $$
# \ell(\theta;x_i) = \| x_i - z_{\sigma(i)} \|^2 = \sum_{j=1}^p (x_{i,j} - z_{\sigma(i),j})^2.
# $$
# Loss, risk, and empirical risk still can be defined, but many concepts are not the same (such as bias-variance tradeoff).
# + [markdown] slideshow={"slide_type": "slide"}
# ### Issue with training error in Supervised learning.
#
# Let $\hat \theta$ be the ERM, then the *training error* is
# $$
# R_n(\hat \theta) = \min_{\theta \in \Theta} R_n(\theta)
# $$
# which does NOT converge to $R(\theta)$ because
# $$
# \mathbb E \min_\theta R_n(\theta) \ne \min_{\theta} \mathbb E R_n(\theta) = \min_\theta R(\theta).
# $$
#
# ### Solution
#
# Split the data randomly into training and test sets:
# - train $\hat \theta$ with the training data
# - test $\hat \theta$ with the test data
#
# Because the test data is independent of $\hat \theta$ we can think of the training process as fixed and test error is now unbiased for risk of $\hat \theta$.
# + slideshow={"slide_type": "slide"}
## randomly shuffle data and split
n,p = X.shape
Ind = np.arange(n)
np.random.shuffle(Ind)
train_size = 2 * n // 3 +1 # set training set size
X_tr, X_te = X[Ind[:train_size],:], X[Ind[train_size:],:]
Y_tr, Y_te = Y[Ind[:train_size]], Y[Ind[train_size:]]
# + slideshow={"slide_type": "slide"}
## compute losses on test set
betahat = fit(X_tr,Y_tr)
Y_hat_te = [predict(x,betahat) for x in X_te]
test_losses = [loss(yhat,y) for yhat,y in zip(Y_hat_te,Y_te)]
## compute losses on train set
Y_hat_tr = [predict(x,betahat) for x in X_tr]
train_losses = [loss(yhat,y) for yhat,y in zip(Y_hat_tr,Y_tr)]
# + slideshow={"slide_type": "slide"}
train_losses
# + slideshow={"slide_type": "slide"}
test_losses
# + slideshow={"slide_type": "slide"}
print("train avg loss: {}\ntest avg loss: {}".format(np.mean(train_losses), np.mean(test_losses)))
print("n p :",n,p)
# + slideshow={"slide_type": "slide"}
def train_test_split(X,Y,split_pr = 0.5):
"""train-test split"""
n,p = X.shape
Ind = np.arange(n)
np.random.shuffle(Ind)
train_size = int(split_pr * n) # set training set size
X_tr, X_te = X[Ind[:train_size],:], X[Ind[train_size:],:]
Y_tr, Y_te = Y[Ind[:train_size]], Y[Ind[train_size:]]
return (X_tr,Y_tr), (X_te, Y_te)
# + slideshow={"slide_type": "slide"}
Y = wine_ar[:,-1]
X = wine_ar[:,:-1]
(X_tr,Y_tr), (X_te, Y_te) = train_test_split(X,Y)
# + slideshow={"slide_type": "slide"}
## compute losses on test set
betahat = fit(X_tr,Y_tr)
Y_hat_te = [predict(x,betahat) for x in X_te]
test_losses = [loss(yhat,y) for yhat,y in zip(Y_hat_te,Y_te)]
## compute losses on train set
Y_hat_tr = [predict(x,betahat) for x in X_tr]
train_losses = [loss(yhat,y) for yhat,y in zip(Y_hat_tr,Y_tr)]
# + slideshow={"slide_type": "slide"}
print("train avg loss: {}\ntest avg loss: {}".format(np.mean(train_losses), np.mean(test_losses)))
# -
# ### Summary
#
# - Want to minimize true risk (expected loss)
# - Instead we minimize empirical risk (training error)
# - Training error is now biased, so we do training test split
|
lectures/lecture1/lecture1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="UiNxsd4_q9wq" colab_type="text"
# ### What-If Tool - age-predicting regression model
#
# Copyright 2019 Google LLC.
# SPDX-License-Identifier: Apache-2.0
#
# This notebook shows use of the [What-If Tool](https://pair-code.github.io/what-if-tool) on a regression model.
#
# This notebook trains a linear regressor to predict a person's age from their census data using the [UCI census data](https://archive.ics.uci.edu/ml/datasets/census+income).
#
# It then visualizes the results of the trained regressor on test data using the What-If Tool.
#
# + id="qqB2tjOMETmr" colab_type="code" colab={}
#@title Install the What-If Tool widget if running in colab {display-mode: "form"}
try:
import google.colab
# !pip install --upgrade witwidget
except:
pass
# + id="jlwjF-Nnmoww" colab_type="code" colab={}
#@title Define helper functions {display-mode: "form"}
import pandas as pd
import numpy as np
import tensorflow as tf
import functools
# Creates a tf feature spec from the dataframe and columns specified.
def create_feature_spec(df, columns=None):
feature_spec = {}
if columns == None:
columns = df.columns.values.tolist()
for f in columns:
if df[f].dtype is np.dtype(np.int64) or df[f].dtype is np.dtype(np.int32):
feature_spec[f] = tf.io.FixedLenFeature(shape=(), dtype=tf.int64)
elif df[f].dtype is np.dtype(np.float64):
feature_spec[f] = tf.io.FixedLenFeature(shape=(), dtype=tf.float32)
else:
feature_spec[f] = tf.io.FixedLenFeature(shape=(), dtype=tf.string)
return feature_spec
# Creates simple numeric and categorical feature columns from a feature spec and a
# list of columns from that spec to use.
#
# NOTE: Models might perform better with some feature engineering such as bucketed
# numeric columns and hash-bucket/embedding columns for categorical features.
def create_feature_columns(columns, feature_spec):
ret = []
for col in columns:
if feature_spec[col].dtype is tf.int64 or feature_spec[col].dtype is tf.float32:
ret.append(tf.feature_column.numeric_column(col))
else:
ret.append(tf.feature_column.indicator_column(
tf.feature_column.categorical_column_with_vocabulary_list(col, list(df[col].unique()))))
return ret
# An input function for providing input to a model from tf.Examples
def tfexamples_input_fn(examples, feature_spec, label, mode=tf.estimator.ModeKeys.EVAL,
num_epochs=None,
batch_size=64):
def ex_generator():
for i in range(len(examples)):
yield examples[i].SerializeToString()
dataset = tf.data.Dataset.from_generator(
ex_generator, tf.dtypes.string, tf.TensorShape([]))
if mode == tf.estimator.ModeKeys.TRAIN:
dataset = dataset.shuffle(buffer_size=2 * batch_size + 1)
dataset = dataset.batch(batch_size)
dataset = dataset.map(lambda tf_example: parse_tf_example(tf_example, label, feature_spec))
dataset = dataset.repeat(num_epochs)
return dataset
# Parses Tf.Example protos into features for the input function.
def parse_tf_example(example_proto, label, feature_spec):
parsed_features = tf.io.parse_example(serialized=example_proto, features=feature_spec)
target = parsed_features.pop(label)
return parsed_features, target
# Converts a dataframe into a list of tf.Example protos.
def df_to_examples(df, columns=None):
examples = []
if columns == None:
columns = df.columns.values.tolist()
for index, row in df.iterrows():
example = tf.train.Example()
for col in columns:
if df[col].dtype is np.dtype(np.int64) or df[col].dtype is np.dtype(np.int32):
example.features.feature[col].int64_list.value.append(int(row[col]))
elif df[col].dtype is np.dtype(np.float64):
example.features.feature[col].float_list.value.append(row[col])
elif row[col] == row[col]:
example.features.feature[col].bytes_list.value.append(row[col].encode('utf-8'))
examples.append(example)
return examples
# Converts a dataframe column into a column of 0's and 1's based on the provided test.
# Used to force label columns to be numeric for binary classification using a TF estimator.
def make_label_column_numeric(df, label_column, test):
df[label_column] = np.where(test(df[label_column]), 1, 0)
# + id="nu398ARdeuxe" colab_type="code" colab={}
#@title Read training dataset from CSV {display-mode: "form"}
import pandas as pd
# Set the path to the CSV containing the dataset to train on.
csv_path = 'https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data'
# Set the column names for the columns in the CSV. If the CSV's first line is a header line containing
# the column names, then set this to None.
csv_columns = [
"Age", "Workclass", "fnlwgt", "Education", "Education-Num", "Marital-Status",
"Occupation", "Relationship", "Race", "Sex", "Capital-Gain", "Capital-Loss",
"Hours-per-week", "Country", "Over-50K"]
# Read the dataset from the provided CSV and print out information about it.
df = pd.read_csv(csv_path, names=csv_columns, skipinitialspace=True)
df
# + id="67DYIFxoevt2" colab_type="code" colab={}
#@title Specify input columns and column to predict {display-mode: "form"}
import numpy as np
# Set the column in the dataset you wish for the model to predict
label_column = 'Age'
# Set list of all columns from the dataset we will use for model input.
input_features = [
'Over-50K', 'Workclass', 'Education', 'Marital-Status', 'Occupation',
'Relationship', 'Race', 'Sex', 'Capital-Gain', 'Capital-Loss',
'Hours-per-week', 'Country']
# Create a list containing all input features and the label column
features_and_labels = input_features + [label_column]
# + id="BV4f_4_Lex22" colab_type="code" colab={}
#@title Convert dataset to tf.Example protos {display-mode: "form"}
examples = df_to_examples(df)
# + id="YyLr-_0de1Ii" colab_type="code" colab={}
#@title Create and train the regressor {display-mode: "form"}
num_steps = 200 #@param {type: "number"}
# Create a feature spec for the classifier
feature_spec = create_feature_spec(df, features_and_labels)
# Define and train the classifier
train_inpf = functools.partial(tfexamples_input_fn, examples, feature_spec, label_column)
regressor = tf.estimator.LinearRegressor(
feature_columns=create_feature_columns(input_features, feature_spec))
regressor.train(train_inpf, steps=num_steps)
# + id="NUQVro76e38Q" colab_type="code" colab={}
#@title Invoke What-If Tool for test data and the trained model {display-mode: "form"}
num_datapoints = 2000 #@param {type: "number"}
tool_height_in_px = 1000 #@param {type: "number"}
from witwidget.notebook.visualization import WitConfigBuilder
from witwidget.notebook.visualization import WitWidget
# Load up the test dataset
test_csv_path = 'https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.test'
test_df = pd.read_csv(test_csv_path, names=csv_columns, skipinitialspace=True,
skiprows=1)
test_examples = df_to_examples(test_df[0:num_datapoints])
# Setup the tool with the test examples and the trained classifier
config_builder = WitConfigBuilder(test_examples[0:num_datapoints]).set_estimator_and_feature_spec(
regressor, feature_spec).set_model_type('regression')
WitWidget(config_builder, height=tool_height_in_px)
# + [markdown] id="7VlncgiQTDFw" colab_type="text"
# #### Exploration ideas
#
# - Create a scatter plot of age on the X-axis by inference value on the Y-axis to see how the predicted age compares to the actual age for all test examples.
# - How does the model do overall? Is there a pattern to the errors it creates?
#
# - In the "Performance" tab, set the ground truth feature to "age". You can now see a simple breakdown of the mean error, mean absolute error, and mean squared error over the entire test set. Additionally, you can set a scatter axis or binning option to be any of those three calculations (i.e. you can make a scatter plot of age vs inference error).
#
# - Replace the LinearRegressor with a DNNRegressor (you'll need to decide on the size and number of the hidden layers). Can you create a better model? Does this model have the same pattern of performance issues as the previous model, or are there errors more evenly distributed?
# - If you look at the [WIT Model Comparison](https://colab.research.google.com/github/pair-code/what-if-tool/blob/master/WIT_Model_Comparison.ipynb) notebook, you can see how to compare the two models on a single WIT.
#
# - In the "Performance" tab, slice the dataset by any feature or pair of features. Are there feature values for which the model seems to perform better or worse on?
#
#
|
WIT_Age_Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
import sys
sys.path.insert(0, '../code')
import deep_forest
import torch as th
from torch import nn as nn
import matplotlib.pyplot as plt
# %matplotlib inline
from math import pi
# +
# 1000 x 2 ==> batch x features
x = th.rand([1000, 2])
x[:, 0] *= 2*pi
x[:, 0] -= pi
x[:, 1] *= 3
x[:, 1] -= 1.5
# Labels
y = (th.sin(x[:, 0] * 2) * 0.5 < x[:, 1]).long()
# + tags=[]
model = deep_forest.DeepForest(25, 2, 2, 1, 10)
# -
device = th.device("cuda" if th.cuda.is_available() else "cpu")
model = model.to(device)
x = x.to(device)
y = y.to(device)
# +
optimizer = th.optim.Adam(model.parameters())
for i in range(2000):
model.populate_best(x[:, :], y[:])
optimizer.zero_grad()
loss = model.loss(x[:, :], y[:], device)
loss.backward()
optimizer.step()
if i % 200 == 0:
print("====EPOCH %d====\nAcc: %s\nLoss: %s" % (i, str(th.mean((model.forward(x[:, :], device) == y[:]).float())), str(loss)))
print("==============\nFINAL ACC: %s" % str(th.mean((model.forward(x[:, :], device) == y[:]).float())))
# -
print(y[:15])
print(model.forward(x, device)[:15].long())
cdict = {0: 'green', 1: 'purple'}
plt.scatter(x[:, 0], x[:, 1], c=[cdict[i] for i in model.forward(x, device).cpu().numpy()])
plt.show()
# +
mlp = nn.Sequential(
nn.Linear(2, 15),
nn.LeakyReLU(),
nn.Linear(15, 15),
nn.LeakyReLU(),
nn.Linear(15, 2),
nn.Softmax()
)
optimizer = th.optim.Adam(mlp.parameters())
for i in range(1500):
optimizer.zero_grad()
preds = mlp(x[:, :])
loss = nn.functional.cross_entropy(preds, (y[:].type(th.LongTensor)).to(device))
loss.backward()
optimizer.step()
if i % 100 == 0:
print("====EPOCH %d====\nAcc: %s\nLoss: %s" % (i, str(th.mean((th.argmax(mlp(x[:]), 1) == y[:]).float())), str(loss)))
print("==============\nFINAL ACC: %s" % str(th.mean((th.argmax(mlp(x[:]), 1) == y[:]).float())))
# -
from sklearn.ensemble import RandomForestClassifier
clf = RandomForestClassifier(max_depth=2)
clf.fit(x[:, :].numpy(), y[:].numpy())
print(clf.score(x[:, :].numpy(), y[:].numpy()))
|
snippets/deep_forest.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Data Types
# Notice in the examples below, the different operations result in different types of outcomes.
print(1 + 1) # this is an integer
print(1.0 + 1) # this is an integer added to a decimal/floating point
print("1" + "1") # this a string/text
print(100 * 2)
print(100 * "2")
# We will learn more about the different types of variables later, but for now, remember that you can use the `type` command to find out the type of an expression:
type(1)
type(3.14)
type("Hello")
type("1")
type("3.14")
type(1 + 1)
type("1" + "1")
type(10 * "2")
# **Lesson**: Notice the different data types above: `str` (string), `int` (integer), `float` (decimal numbers). One thing hat you will notice is that the outcome of the various operators (`+`, `*`, etc) can be different based on the data types involved.
# #### Exercise
#
# What will the following program print out? Figure it our _before_ running the program.
x = 41
x = x + 1
print(x)
x = "41"
x = x + "1"
print(x)
|
notes/C3-DataTypes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# ---
#
# _You are currently looking at **version 1.0** of this notebook. To download notebooks and datafiles, as well as get help on Jupyter notebooks in the Coursera platform, visit the [Jupyter Notebook FAQ](https://www.coursera.org/learn/python-machine-learning/resources/bANLa) course resource._
#
# ---
# # Classifier Visualization Playground
#
# The purpose of this notebook is to let you visualize various classsifiers' decision boundaries.
#
# The data used in this notebook is based on the [UCI Mushroom Data Set](http://archive.ics.uci.edu/ml/datasets/Mushroom?ref=datanews.io) stored in `mushrooms.csv`.
#
# In order to better vizualize the decision boundaries, we'll perform Principal Component Analysis (PCA) on the data to reduce the dimensionality to 2 dimensions. Dimensionality reduction will be covered in module 4 of this course.
#
# Play around with different models and parameters to see how they affect the classifier's decision boundary and accuracy!
# +
# %matplotlib notebook
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
from sklearn.model_selection import train_test_split
df = pd.read_csv('mushrooms.csv')
df2 = pd.get_dummies(df)
df3 = df2.sample(frac=0.08)
X = df3.iloc[:,2:]
y = df3.iloc[:,1]
pca = PCA(n_components=2).fit_transform(X)
X_train, X_test, y_train, y_test = train_test_split(pca, y, random_state=0)
plt.figure(dpi=120)
plt.scatter(pca[y.values==0,0], pca[y.values==0,1], alpha=0.5, label='Edible', s=2)
plt.scatter(pca[y.values==1,0], pca[y.values==1,1], alpha=0.5, label='Poisonous', s=2)
plt.legend()
plt.title('Mushroom Data Set\nFirst Two Principal Components')
plt.xlabel('PC1')
plt.ylabel('PC2')
plt.gca().set_aspect('equal')
# -
def plot_mushroom_boundary(X, y, fitted_model):
plt.figure(figsize=(9.8,5), dpi=100)
for i, plot_type in enumerate(['Decision Boundary', 'Decision Probabilities']):
plt.subplot(1,2,i+1)
mesh_step_size = 0.01 # step size in the mesh
x_min, x_max = X[:, 0].min() - .1, X[:, 0].max() + .1
y_min, y_max = X[:, 1].min() - .1, X[:, 1].max() + .1
xx, yy = np.meshgrid(np.arange(x_min, x_max, mesh_step_size), np.arange(y_min, y_max, mesh_step_size))
if i == 0:
Z = fitted_model.predict(np.c_[xx.ravel(), yy.ravel()])
else:
try:
Z = fitted_model.predict_proba(np.c_[xx.ravel(), yy.ravel()])[:,1]
except:
plt.text(0.4, 0.5, 'Probabilities Unavailable', horizontalalignment='center',
verticalalignment='center', transform = plt.gca().transAxes, fontsize=12)
plt.axis('off')
break
Z = Z.reshape(xx.shape)
plt.scatter(X[y.values==0,0], X[y.values==0,1], alpha=0.4, label='Edible', s=5)
plt.scatter(X[y.values==1,0], X[y.values==1,1], alpha=0.4, label='Posionous', s=5)
plt.imshow(Z, interpolation='nearest', cmap='RdYlBu_r', alpha=0.15,
extent=(x_min, x_max, y_min, y_max), origin='lower')
plt.title(plot_type + '\n' +
str(fitted_model).split('(')[0]+ ' Test Accuracy: ' + str(np.round(fitted_model.score(X, y), 5)))
plt.gca().set_aspect('equal');
plt.tight_layout()
plt.subplots_adjust(top=0.9, bottom=0.08, wspace=0.02)
# +
from sklearn.linear_model import LogisticRegression
model = LogisticRegression()
model.fit(X_train,y_train)
plot_mushroom_boundary(X_test, y_test, model)
# +
from sklearn.neighbors import KNeighborsClassifier
model = KNeighborsClassifier(n_neighbors=20)
model.fit(X_train,y_train)
plot_mushroom_boundary(X_test, y_test, model)
# +
from sklearn.tree import DecisionTreeClassifier
model = DecisionTreeClassifier(max_depth=3)
model.fit(X_train,y_train)
plot_mushroom_boundary(X_test, y_test, model)
# +
from sklearn.tree import DecisionTreeClassifier
model = DecisionTreeClassifier()
model.fit(X_train,y_train)
plot_mushroom_boundary(X_test, y_test, model)
# +
from sklearn.ensemble import RandomForestClassifier
model = RandomForestClassifier()
model.fit(X_train,y_train)
plot_mushroom_boundary(X_test, y_test, model)
# +
from sklearn.svm import SVC
model = SVC(kernel='linear')
model.fit(X_train,y_train)
plot_mushroom_boundary(X_test, y_test, model)
# +
from sklearn.svm import SVC
model = SVC(kernel='rbf', C=1)
model.fit(X_train,y_train)
plot_mushroom_boundary(X_test, y_test, model)
# +
from sklearn.svm import SVC
model = SVC(kernel='rbf', C=10)
model.fit(X_train,y_train)
plot_mushroom_boundary(X_test, y_test, model)
# +
from sklearn.naive_bayes import GaussianNB
model = GaussianNB()
model.fit(X_train,y_train)
plot_mushroom_boundary(X_test, y_test, model)
# +
from sklearn.neural_network import MLPClassifier
model = MLPClassifier()
model.fit(X_train,y_train)
plot_mushroom_boundary(X_test, y_test, model)
# -
|
3_machine_learning/week_2/classifier_visualization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#This notebook is about how to do the preprocessing and generating datasets
# -
# # Preprocessing
#import modules
from desidlas.datasets.preprocess import estimate_s2n,normalize,rebin
from desidlas.datasets.DesiMock import DesiMock
from desidlas.dla_cnn.defs import best_v
import numpy as np
import os
from os.path import join
from pkg_resources import resource_filename
from pathlib import Path
#an example to load data
#three kinds of data files are necessary for DESI spectra: spectra,truth and zbest
datafile_path = os.path.join(resource_filename('desidlas', 'tests'), 'datafile')
spectra= os.path.join(datafile_path, 'spectra', 'spectra-16-1375.fits')
truth=os.path.join(datafile_path, 'spectra', 'truth-16-1375.fits')
zbest=os.path.join(datafile_path, 'spectra', 'zbest-16-1375.fits')
#import get_sightlines
from desidlas.datasets.get_sightlines import get_sightlines
# an example for the output file path
outpath='desidlas/tests/datafile/sightlines-16-1375.npy'
#load the spectra using DesiMock
#the output file includes all the information we need
sightlines=get_sightlines(spectra,truth,zbest,outpath)
# # Generating Dataset
import scipy.signal as signal
from desidlas.datasets.datasetting import split_sightline_into_samples,select_samples_50p_pos_neg,pad_sightline
from desidlas.datasets.preprocess import label_sightline
from desidlas.dla_cnn.spectra_utils import get_lam_data
from desidlas.datasets.get_dataset import make_datasets,make_smoothdatasets
#use sightlnes to produce datasets for training or prediction
#'sightlines' is the output of get_sightlines module, it can also be loaded from the npy file
sightlines=np.load('desidlas/tests/datafile/sightlines-16-1375.npy',allow_pickle = True,encoding='latin1')
#generating and save dataset
dataset=make_datasets(sightlines,validate=True,output='desidlas/tests/datafile/dataset-16-1375.npy')
#for spectra with SNR<3, we use smoothing method when generating dataset
smoothdataset=make_smoothdatasets(sightlines,validate=True,output='desidlas/tests/datafile/dataset-16-1375-smooth.npy')
|
desidlas/notebook/preprocess_datasets.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import tiledb
import tiledb.cf
import netCDF4
import numpy as np
import matplotlib.pyplot as plt
netcdf_file = "../data/simple1.nc"
group_uri = "arrays/simple_netcdf_to_group_1"
array_uri = "arrays/simple_netcdf_to_array_1"
# +
import shutil
# clean up any previous runs
try:
shutil.rmtree(group_uri)
shutil.rmtree(array_uri)
except:
pass
# -
# # Converting a simple NetCDF file to a TileDB group or array
#
# ## About this Example
#
# ### What it Shows
#
# The purpose of this example is to show the basics of converting a NetCDF file to a TileDB group or array.
#
# This includes:
#
# 1. Options for auto-generating a converter from a NetCDF file.
# 2. Changing the TileDB schema settings before conversion.
# 3. Creating the TileDB group and copying data from the NetCDF file to the TileDB array.
#
# ### Set-up Requirements
#
# This example requires the following python packages are installed: netCDF4, numpy, tiledb, tiledb-cf, and matplotlib
#
# ## Create an example NetCDF file
#
# If the NetCDF file does not exist, we create a small NetCDF file for this example.
#
# ### Example dataset
#
# This example shows convertering a small NetCDF file with 2 dimensions and 4 variables:
#
# * Dimensions:
# * x: size=100
# * y: size=100
# * Variables:
# * x(x)
# * description: evenly spaced values from -5 to 5
# * data type: 64-bit floating point
# * y(x)
# * description: evenly spaced values from -5 to 5
# * data type: 64-bit floating point
# * A1(x, y)
# * description: x + y
# * data type: 64-bit floating point
# * A1(x, y)
# * description: sin((x/2)^2 + y^2
# * data type: 64-bit floating point
import os
if not os.path.exists(netcdf_file):
x_data = np.linspace(-5.0, 5.0, 100)
y_data = np.linspace(-5.0, 5.0, 100)
xv, yv = np.meshgrid(x_data, y_data, sparse=True)
with netCDF4.Dataset(netcdf_file, mode="w") as dataset:
dataset.setncatts({"title": "Simple dataset for examples"})
dataset.createDimension("x", 100)
dataset.createDimension("y", 100)
A1 = dataset.createVariable("A1", np.float64, ("x", "y"))
A1.setncattr("full_name", "Example matrix A1")
A1.setncattr("description", "x + y")
A1[:, :] = xv + yv
A2 = dataset.createVariable("A2", np.float64, ("x", "y"))
A2[:, :] = np.sin((xv / 2.0) ** 2 + yv ** 2)
A2.setncattr("full_name", "Example matrix A2")
A2.setncattr("description", "sin((x/2)^2 + y^2")
x1 = dataset.createVariable("x", np.float64, ("x",))
x1[:] = x_data
y = dataset.createVariable("y", np.float64, ("y",))
y[:] = y_data
print(f"Created example NetCDF file `{netcdf_file}`.")
else:
print(f"Example NetCDF file `{netcdf_file}` already exists.")
# ## Auto-Generating Converter from File
#
# The `NetCDF4ConverterEngine.from_file` and `NetCDF4ConverterEngine.from_group` are used to auto-generate a NetCDF-to-TileDB conversion recipe.
#
#
# Parameters:
#
# * Set the location of the NetCDF group to be converted.
#
# * `from_file`:
#
# * `input_file`: The input NetCDF file to generate the converter engine from.
# * `group_path`: The path to the NetCDF group to copy data from. Use `'/'` for the root group.
#
# * `from_group`:
#
# * `input_group`
#
# * Set the array grouping. A NetCDF variable maps to TileDB attributes. The `collect_attrs` parameters determines if each NetCDF variable is stored in a separate array, or if all NetCDF variables with the same underlying dimensions are stored in the same TileDB array. Scalar variables are always grouped together.
#
# * `collect_attrs`: If `True`, store all attributes with the same dimensions in the same array. Otherwise, store each attribute in a separate array.
#
# * Set default properties for TileDB dimension.
#
# * `unlimited_dim_size`: The default size of the domain for TileDB dimensions created from unlimited NetCDF dimensions. If `None`, the current size of the NetCDF dimension will be used.
# * `dim_dtype`: The default numpy dtype to use when converting a NetCDF dimension to a TileDB dimension.
#
# * Set tile sizes for TileDB dimensions. Multiple arrays in the TileDB group may have the same name, domain, and type, but different tiles and compression filters. The `tiles_by_var` and `tiles_by_dims` parameters allow a way of setting the tiles for the dimensions in different arrays. The `tiles_by_var` parameter is a mapping from variable name to the tiles for the dimensions of the array that variable is stored in. The `tiles_by_dims` parameter is a mapping from the names of the dimensions of the array to the tiles for the dimensions of the array. If using `collect_attrs=True`, then `tiles_by_dims` will over-write `tiles_by_var`. If using `collect_attrs=False`, then `tiles_by_vars` with over-write `tiles_by_var`.
#
# * `tiles_by_var`: A map from the name of a NetCDF variable to the tiles of the dimensions of the variable in the generated TileDB array.
# * `tiles_by_dims`: A map from the name of NetCDF dimensions defining a variable to the tiles of those dimensions in the generated TileDB array.
#
# * Convert 1D variables with the same name and dimension to a TileDB dimension instead of a TileDB attribute.
#
# * `coords_to_dims`: If `True`, convert the NetCDF coordinate variable into a TileDB dimension for sparse arrays. Otherwise, convert the coordinate dimension into a TileDB dimension and the coordinate variable into a TileDB attribute.
# ### Examples: Collecting attributes and setting tiles
#
# Below are some examples of how the parameter `collect_attrs`, `tiles_by_var`, and `tiles_by_dims` interact.
# 1. `collect_attrs=True`
# * `A1` and `A2` are in the same array.
# * `tile=None` for all dimensions.
# 2. `collect_attrs=True`, `tiles_by_dims={(x,y): (10, 20)}`
# * `A1` and `A2` are in the same array.
# * Only array with dimensions `(x,y)` has tiles set.
# 3. `collect_attrs=True`, `tiles_by_var={'A1': (50, 50)}`
# * `A1` and `A2` are in the same array.
# * Only array with variable `A1` has tiles set.
# 4. `collect_attrs=True`, `tiles_by_dims={(x,y): (10, 20)}`, `tiles_by_var={'A1': (50, 50)}`
# * `A1` and `A2` are in the same array.
# * Only array with dimensions `(x,y)` has tiles set. `tiles_by_dims` wrote over `tiles_by_var`.
# 5. `collect_attrs=False`
# * `A1` and `A2` are in separate arrays.
# * `tile=None` for all dimensions.
# 6. `collect_attrs=False`, `tiles_by_dims={(x,y): (10, 20)}`
# * `A1` and `A2` are in separate arrays.
# * Only arrays with dimensions `(x,y)` have tiles set.
# 7. `collect_attrs=False`, `tiles_by_var={'A1': (50, 50)}`
# * `A1` and `A2` are in separate arrays.
# * Only array with variable `A1` has tiles set.
# 8. `collect_attrs=False`, `tiles_by_dims={(x,y): (10, 20)}`, `tiles_by_var={'A1': (50, 50)}`
# * `A1` and `A2` are in separate arrays.
# * The array with `A2` has tiles set by `tiles_by_dims`.
# * The array with `A1` has tiles set with `tiles_by_var` writing over `tiles_by_dims`.
# Change the parameter values and see how it effects the tile value in the DimensionCreators (inside ArrayCreators)
tiledb.cf.NetCDF4ConverterEngine.from_file(netcdf_file, collect_attrs=True, tiles_by_dims=None, tiles_by_var=None)
# ## Convert `simple1.nc` to a TileDB Group
#
# In this example, we create a NetCDF4ConverterEngine from the NetCDF file and manually change properties of the TileDB arrays before conversion. NetCDF dimensions are mapped to TileDB dimensions, and NetCDF variables are mapped to TileDB attributes.
converter = tiledb.cf.NetCDF4ConverterEngine.from_file(netcdf_file, collect_attrs=True, dim_dtype=np.uint32)
converter
# Update properties manually by modifying the array creators
# Update properties for array of matrices
data_array = converter.get_array_creator_by_attr("A1")
data_array.name = "data"
data_array.domain_creator.tiles = (20, 20)
for attr_creator in data_array:
attr_creator.filters = tiledb.FilterList([tiledb.ZstdFilter()])
converter
# Update properties for the array containing "x.data"
# Update properties for the array containing "y.data"
# Run the conversions to create two dense TileDB arrays:
converter.convert_to_group(group_uri)
# ### Examine the TileDB group schema
group_schema = tiledb.cf.GroupSchema.load(group_uri)
group_schema
# ### Examine the data in the arrays
#
# Open the attributes from the generated TileDB group:
with tiledb.cf.Group(group_uri, attr="x.data") as group:
with (
group.open_array(attr="x.data") as x_array,
group.open_array(attr="y.data") as y_array,
group.open_array(array="data") as data_array,
):
x = x_array[:]
y = y_array[:]
data = data_array[...]
A1 = data["A1"]
a1_description = tiledb.cf.AttrMetadata(data_array.meta, "A1")["description"]
fig, axes = plt.subplots(nrows=1, ncols=1)
axes.contourf(x, y, A1);
axes.set_title(a1_description);
# +
# Plot A2 over -1 <= x <= 1 and 0 <= y < 2 by only querying on those regions
# -
# ## Convert `simple1.nc` to a Single Sparse TileDB Array
#
# In this example, we create a NetCDF4ConverterEngine from the NetCDF file and manually change properties of the TileDB arrays before conversion. Here we use `coord_to_dims` to convert the `x` and `y` variables to TileDB variables in a sparse array.
converter2 = tiledb.cf.NetCDF4ConverterEngine.from_file(netcdf_file, coords_to_dims=True)
converter2
# When using `coords_to_dims` the converter cannot auto-detect the preferred domain for dimensions.
# This must be set before converting data. Using `to_schema()` is a good way to check if the TileDB
# arrays are all valid.
try:
converter.to_schema()
except tiledb.libtiledb.TileDBError as err:
print(err)
# Set the domain for both dimensions
converter2.get_shared_dim('x').domain = (-5.0, 5.0)
converter2.get_shared_dim('y').domain = (-5.0, 5.0)
data_array = converter2.get_array_creator("array0")
data_array.domain_creator.tiles = (1.0, 1.0)
data_array.capacity = 400
for attr_creator in data_array:
attr_creator.filters = tiledb.FilterList([tiledb.ZstdFilter()])
converter2
# +
# Set the tiles for the sparse array
# +
# Set the capacity for the array using the `capacity` property
# +
# Set compression filters on the attributes using the `filters` property of the AttrCreator objects
# -
converter2.convert_to_array(array_uri)
# +
# Try querying data from the sparse array. Notice the results are returned in coordinate form like earlier examples.
|
notebooks/6c-netcdf-to-tiledb.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# # Overview
# **Basic imports**
# +
import pandas as pd
import numpy as np
from Bio import SeqIO
from matplotlib import pyplot as plt
import sklearn.metrics
from scipy import stats
import glob
# -
# **Custom imports**
import process_couplings
# **Plotting parameters**
# +
import matplotlib
matplotlib.rcParams['xtick.labelsize'] = 16
matplotlib.rcParams['ytick.labelsize'] = 16
matplotlib.rcParams['axes.labelsize'] = 18
matplotlib.rcParams['axes.titlesize'] = 18
matplotlib.rcParams['axes.grid'] = True
matplotlib.rcParams['grid.color'] = '0.5'
matplotlib.rcParams['grid.linewidth'] = '0.5'
matplotlib.rcParams['axes.edgecolor'] = '0.25'
matplotlib.rcParams['xtick.color'] = '0'
matplotlib.rcParams['ytick.color'] = '0'
matplotlib.rcParams['xtick.major.width'] = 1
matplotlib.rcParams['ytick.major.width'] = 1
matplotlib.rcParams['ytick.major.size'] = 5
matplotlib.rcParams['xtick.major.size'] = 5
matplotlib.rcParams['axes.spines.right'] = True
matplotlib.rcParams['axes.spines.left'] = True
matplotlib.rcParams['axes.spines.top'] = True
matplotlib.rcParams['axes.spines.bottom'] = True
matplotlib.rcParams['font.family'] = 'sans-serif'
matplotlib.rcParams['font.sans-serif'] = 'helvetica'
matplotlib.rcParams['font.weight']='normal'
matplotlib.rcParams['axes.axisbelow'] = True
# -
# **Creating a folder to store results. This should be adjusted based on how you want to store figures (if at all). Relevant variable is: `figs_dir`**
import datetime
year = datetime.date.today().year
month = datetime.date.today().month
import os
figs_dir = '../Results/Figures/{}_{:02}'.format(year, month)
if not os.path.exists(figs_dir):
os.makedirs(figs_dir)
# **Finally, some directory variables and constants to apply throughout**
couplings_dir = '../Results/couplings/'
contacts_dir = '../Data/psicov150_aln_pdb/pdb/'
fastas_dir = '../Data/psicov150_aln_pdb/aln_fasta_max1k/'
# +
length_based_modifier = 1.
primary_distance_cutoff = 6
contact_definition = 7.5
# -
# # Visualize some generalities
# **Distributions of coupling values**
# +
prot_name = '1aoeA'
# testy_df = process_couplings.process_ccmpredpy(couplings_dir+'{}.raw.uniform.mat'.format(prot_name))
# testy_df = process_couplings.process_ccmpredpy(couplings_dir+'{}.ent.uniform.mat'.format(prot_name))
testy_df = process_couplings.process_ccmpredpy(couplings_dir+'{}.apc.uniform.mat'.format(prot_name))
# testy_df = process_couplings.process_ccmpredpy(couplings_dir+'{}.raw.GSC_meanScale.mat'.format(prot_name))
# testy_df = process_couplings.process_ccmpredpy(couplings_dir+'{}.ent.GSC_meanScale.mat'.format(prot_name))
# testy_df = process_couplings.process_ccmpredpy(couplings_dir+'{}.apc.GSC_meanScale.mat'.format(prot_name))
fig, ax = plt.subplots()
ax.hist(testy_df['couplings'], 80);
# -
# **Load in a contact file and test PPV**
# +
df_contacts = pd.read_csv(contacts_dir+'{}_SCcenter_contacts.csv'.format(prot_name), index_col=0)
df_contacts, df_contacts_stack = process_couplings.process_contacts_df(df_contacts)
seq = list(SeqIO.parse(fastas_dir+'{}.fasta'.format(prot_name), 'fasta'))[0]
seq = str(seq.seq)
df_merged = process_couplings.merge_contacts_couplings(df_contacts_stack, testy_df, seq)
df_merged = process_couplings.remove_close(df_merged, primary_distance_cutoff)
print(process_couplings.ppv_from_df(df_merged, int(len(seq)*length_based_modifier), length_cutoff=contact_definition))
# -
# **And visualize the precision-recall curve**
# +
df_merged['contact'] = df_merged['distance']<=contact_definition
aupr = sklearn.metrics.average_precision_score(df_merged['contact'], df_merged['couplings'])
precision, recall, trash = sklearn.metrics.precision_recall_curve(df_merged['contact'], df_merged['couplings'])
fig, ax = plt.subplots()
ax.plot(recall, precision)
print('Average precision:', aupr)
# -
# # Systematic-ish
#
# **If run on the entire results set this will take a while and eat up a lot of memory. Can be modified to only read in certain types or proteins if necessary but currently this will take it all in**
# +
results_dicty_ppv = {}
results_dicty_aupr = {}
types_to_test = ['raw', 'apc', 'ent']
for type_to_test in types_to_test:
for infile in sorted(glob.glob(couplings_dir+'*.mat'))[:]:
prot_name = infile.split('/')[-1].split('.')[0]
params = '.'.join(infile.split('/')[-1].split('.')[1:-1])
if params[:3] != type_to_test:
continue
#Read in the couplings for the protein of interest
testy_df = process_couplings.process_ccmpredpy(infile)
#Read in the contacts
df_contacts = pd.read_csv(contacts_dir+'{}_SCcenter_contacts.csv'.format(prot_name), index_col=0)
df_contacts, df_contacts_stack = process_couplings.process_contacts_df(df_contacts)
#Read in the fasta sequence
seq = list(SeqIO.parse(fastas_dir+'{}.fasta'.format(prot_name), 'fasta'))[0]
seq = str(seq.seq)
#Merge everyone together
df_merged = process_couplings.merge_contacts_couplings(df_contacts_stack, testy_df, seq)
#Remove pairs that are close in primary distance space
df_merged = process_couplings.remove_close(df_merged, primary_distance_cutoff)
#Calculate the PPV and add to a results dictionary
ppv_val, ns = process_couplings.ppv_from_df(df_merged, int(len(seq)*length_based_modifier),\
length_cutoff=contact_definition)
try:
results_dicty_ppv[params].append(ppv_val)
except:
results_dicty_ppv[params] = [ppv_val]
#########
#Further process the merged dataframe to include a binary variable for contacts
df_merged['contact'] = df_merged['distance']<contact_definition
#Calculate the area under the curve and add to a results dictionary
aupr = sklearn.metrics.average_precision_score(df_merged['contact'], df_merged['couplings'])
try:
results_dicty_aupr[params].append(aupr)
except:
results_dicty_aupr[params] = [aupr]
# -
# **See what method/s were best!**
best_results = []
for key, val in results_dicty_ppv.items():
# if key[:3] == 'raw':
best_results.append((key, np.median(val)))
best_results = sorted(best_results, key=lambda x: x[1], reverse=True)
best_results
# # Visualizing
#
# **Setting some variables here that can be toggled to analyze a different set of results**
coup_type = 'apc'
#
#
#
#
results_dicty = results_dicty_ppv
metric = 'PPV'
#
# results_dicty = results_dicty_aupr
# metric = 'AUPR'
# ** Boxplot with means **
#
# Not the cleanest code, but gets the job done for a pretty-ish plot
# +
coup_type = 'apc'
#Getting in all the data
data = [results_dicty['{}.uniform'.format(coup_type)]]+\
[results_dicty['{}.simple_0.{}'.format(coup_type, i)] for i in range(1,10)]+\
[results_dicty['{}.HH_meanScale'.format(coup_type)]]+\
[results_dicty['{}.GSC_meanScale'.format(coup_type)]]+\
[results_dicty['{}.ACL_meanScale'.format(coup_type)]]+\
[results_dicty['{}.HH_maxScale'.format(coup_type)]]+\
[results_dicty['{}.GSC_maxScale'.format(coup_type)]]+\
[results_dicty['{}.ACL_maxScale'.format(coup_type)]]
#My x-labels get a bit complicated because of the nature of the plot
labels = ['Uniform']+[str(i/10) for i in range(1, 10, 2)]+['HH', 'GSC', 'ACL']+['HH', 'GSC', 'ACL']
xvals= [-0.1]+[i/10 for i in range(2,11)]+[i/10 for i in range(12, 18, 2)]+[i/10 for i in range(18, 24, 2)]
fig, ax = plt.subplots(figsize=(10,4))
bplot = ax.boxplot(data, positions=xvals, widths=(0.07), patch_artist=True)
for patch in bplot['boxes']:
patch.set_facecolor('white')
#Drawing some lines to divide the plot up a bit
ax.axvline(0.1, c='k')
ax.axvline(1.1, c='k')
ax.axvline(1.7, c='k')
#Some work with the x-axis
ax.set_xlim(-0.3, 2.3)
ax.set_xticks([-0.1]+[0.2, 0.4, 0.6, 0.8, 1.0]+[1.2, 1.4, 1.6]+[1.8, 2.0, 2.2])
ax.set_xticklabels(labels)
#Now setting up to plot the means in a hideous manner
data = [np.mean(results_dicty['{}.simple_0.{}'.format(coup_type, i)]) for i in range(1,10)]
xvals= [i/10 for i in range(2,11)]
ax.plot([-0.1], [np.mean(results_dicty['{}.uniform'.format(coup_type)])], marker='s', zorder=3, c='firebrick')
ax.axhline(np.mean(results_dicty['{}.uniform'.format(coup_type)]), linestyle='--', c='firebrick', alpha=0.5, zorder=4)
ax.plot(xvals, data, marker='s', zorder=3)
ax.plot([1.2], [np.mean(results_dicty['{}.HH_meanScale'.format(coup_type)])], marker='s', zorder=3, c='steelblue')
ax.plot([1.4], [np.mean(results_dicty['{}.GSC_meanScale'.format(coup_type)])], marker='s', zorder=3, c='steelblue')
ax.plot([1.6], [np.mean(results_dicty['{}.ACL_meanScale'.format(coup_type)])], marker='s', zorder=3, c='steelblue')
ax.plot([1.8], [np.mean(results_dicty['{}.HH_maxScale'.format(coup_type)])], marker='s', zorder=3, c='steelblue')
ax.plot([2.0], [np.mean(results_dicty['{}.GSC_maxScale'.format(coup_type)])], marker='s', zorder=3, c='steelblue')
ax.plot([2.2], [np.mean(results_dicty['{}.ACL_maxScale'.format(coup_type)])], marker='s', zorder=3, c='steelblue')
ax.set_ylabel(metric)
###This had to be done because of some annoying quirk with Affinity Design which I used to layout the final figures
ax.set_xlabel('Threshold ($\lambda$)',\
horizontalalignment='left', x=0.25)
ax.text(0.575, -0.135, 'Mean scale Max scale', transform=ax.transAxes, fontsize=18,
verticalalignment='top')
ax.set_ylim(0, 0.85)
ax.grid(False)
# plt.savefig('{}/{}_summary.pdf'.format(figs_dir, coup_type), bbox_inches='tight')
# -
# **Check statistics on any comparison using the wilcoxon signed-rank test (a paired, non-parametric test)**
# +
# compare_a = 'raw.uniform'
compare_a = 'apc.uniform'
# compare_a = 'raw.uniform'
# compare_b = 'apc.GSC_meanScale'
# compare_a = 'apc.uniform'
compare_b = 'apc.simple_0.9'
# compare_b = 'apc.simpleish_0.8'
# compare_b = 'apc.simple_0.7'
# compare_b = 'apc.GSC_meanScale.RelTime'
# compare_b = 'apc.HH_meanScale'
print(compare_a, np.mean(results_dicty[compare_a]), np.median(results_dicty[compare_a]))
print(compare_b, np.mean(results_dicty[compare_b]), np.median(results_dicty[compare_b]))
print('Significance:', stats.wilcoxon(results_dicty[compare_a], results_dicty[compare_b]))
# -
#For the order of magnitude changes
print(np.median(np.array(results_dicty[compare_b])/np.array(results_dicty[compare_a])))
# **Comparing modified identity-based method with the original identity-based method**
# +
coup_type = 'raw'
labels = ['Uniform']+[str(i/10) for i in range(1, 10, 2)]
print(len(data), len(labels))
xvals= [-0.1]+[i/10 for i in range(2,11)]
prop_cycle = plt.rcParams['axes.prop_cycle']
colors = prop_cycle.by_key()['color']
fig, ax = plt.subplots(figsize=(8,3))
ax.set_xlim(-0.3, 1.1)
ax.set_xticks([-0.1]+[0.2, 0.4, 0.6, 0.8, 1.0])
ax.set_xticklabels(labels)
ax.axvline(0.1, c='k')
ax.plot([-0.1], [np.mean(results_dicty['{}.uniform'.format(coup_type)])], marker='s', zorder=3, c='firebrick', markersize=8)
ax.axhline(np.mean(results_dicty['{}.uniform'.format(coup_type)]), linestyle='--', c='firebrick', alpha=0.5, zorder=4)
data = [np.mean(results_dicty['{}.simpleish_0.{}'.format(coup_type, i)]) for i in range(1,10)]
xvals= [i/10 for i in range(2,11)]
ax.plot(xvals, data, marker='s', zorder=4, label='Similarity-adjusted', markersize=8, c=colors[1])
data = [np.mean(results_dicty['{}.simple_0.{}'.format(coup_type, i)]) for i in range(1,10)]
xvals= [i/10 for i in range(2,11)]
ax.plot(xvals, data, marker='s', zorder=4, label='Original', markersize=8, c=colors[0])
ax.set_ylabel(metric)
ax.set_xlabel('Threshold ($\lambda$)',\
horizontalalignment='left', x=0.525)
ax.set_ylim(0.1, 0.45)
legend = ax.legend(loc=1, fontsize=14, framealpha=1.0)
plt.savefig('{}/{}_identity_compare.pdf'.format(figs_dir, coup_type), bbox_inches='tight')
# ax.grid(False)
# -
# **Comparing weights computed from regular trees vs RelTime trees**
# +
coup_type = 'apc'
x_vals = [-0.1, 0.2, 0.3, 0.5, 0.6, 0.8, 0.9, 1.1, 1.2]
labels = ['Uniform'] +['Raw tree', 'RelTime tree']*4
colors = ['firebrick'] + ['steelblue', 'darkorange']*4
y_vals = [np.mean(results_dicty['{}.uniform'.format(coup_type)]),\
np.mean(results_dicty['{}.GSC_meanScale'.format(coup_type)]),\
np.mean(results_dicty['{}.GSC_meanScale.RelTime'.format(coup_type)]),\
np.mean(results_dicty['{}.ACL_meanScale'.format(coup_type)]),\
np.mean(results_dicty['{}.ACL_meanScale.RelTime'.format(coup_type)]),\
np.mean(results_dicty['{}.GSC_maxScale'.format(coup_type)]),\
np.mean(results_dicty['{}.GSC_maxScale.RelTime'.format(coup_type)]),\
np.mean(results_dicty['{}.ACL_maxScale'.format(coup_type)]),\
np.mean(results_dicty['{}.ACL_maxScale.RelTime'.format(coup_type)])]
y_errs = [np.std(results_dicty['{}.uniform'.format(coup_type)]),\
np.std(results_dicty['{}.GSC_meanScale'.format(coup_type)]),\
np.std(results_dicty['{}.GSC_meanScale.RelTime'.format(coup_type)]),\
np.std(results_dicty['{}.ACL_meanScale'.format(coup_type)]),\
np.std(results_dicty['{}.ACL_meanScale.RelTime'.format(coup_type)]),\
np.std(results_dicty['{}.GSC_maxScale'.format(coup_type)]),\
np.std(results_dicty['{}.GSC_maxScale.RelTime'.format(coup_type)]),\
np.std(results_dicty['{}.ACL_maxScale'.format(coup_type)]),\
np.std(results_dicty['{}.ACL_maxScale.RelTime'.format(coup_type)])]
fig, ax = plt.subplots(figsize=(6,3))
ax.set_xlim(-0.3, 1.35)
ax.errorbar(x_vals[:1], y_vals[:1], yerr=y_errs[:1], marker='s', markersize=10,\
linestyle='', c='firebrick')
ax.errorbar(x_vals[1::2], y_vals[1::2], yerr=y_errs[1::2], marker='s', markersize=10,\
linestyle='', c='steelblue', zorder=4)
ax.errorbar(x_vals[2::2], y_vals[2::2], yerr=y_errs[2::2], marker='s', markersize=10,\
linestyle='', c='darkorange', zorder=4)
ax.plot(x_vals[1::2], y_vals[1::2], marker='s', markersize=10,\
linestyle='', c='steelblue', label='Raw tree', zorder=4)
ax.plot(x_vals[2::2], y_vals[2::2], marker='s', markersize=10,\
linestyle='', c='darkorange', label='RelTime tree', zorder=4)
ax.axvline(0.1, c='k')
ax.axhline(np.mean(results_dicty['{}.uniform'.format(coup_type)]), linestyle='--', c='firebrick', alpha=0.5, zorder=4)
leg = ax.legend(loc='center left', bbox_to_anchor=(0.22, 1.1), ncol=2, fontsize=14)
ax.set_xticks([-0.1]+[0.25, 0.55, 0.85, 1.15])
ax.set_xticklabels(['Uniform', 'GSC', 'ACL', 'GSC', 'ACL'])
ax.set_xlabel('Mean scale Max scale',\
horizontalalignment='left', x=0.28)
ax.set_ylim(0, 0.58)
ax.set_ylabel('PPV')
plt.savefig('{}/{}_RelTime.pdf'.format(figs_dir, coup_type), bbox_inches='tight')
# ax.grid(False)
# -
# # DEPRECATED
# **Below code was not used for manuscript but left here for posterity.**
#
# **Was attempting to code/test my own APC and entropy-corrections from the raw coupling data to make sure that I understood the way that the methods worked but in the end just used the files outputted by CCMPredPy**
prot_name = '1aoeA'
testy_df = process_couplings.process_ccmpredpy('../Results/couplings/{}.raw.GSC_meanScale.mat'.format(prot_name))
apc_df = process_couplings.process_ccmpredpy('../Results/couplings/{}.apc.GSC_meanScale.mat'.format(prot_name))
testy_df.head()
temp_row = {}
for i in list(set(list(testy_df['aa1_loc'])+list(testy_df['aa2_loc']))):
temp_df = testy_df[(testy_df['aa1_loc'] == i) | (testy_df['aa2_loc'] == i)]
temp_row[i] = np.mean(temp_df['couplings'])
testy_df['apc'] = np.nan
cmean = np.mean(testy_df['couplings'])
for index in testy_df.index:
coupling = testy_df.loc[index]['couplings']
ci = temp_row[testy_df.loc[index]['aa1_loc']]
cj = temp_row[testy_df.loc[index]['aa2_loc']]
testy_df.set_value(index, 'apc',coupling - ((ci*cj)/cmean))
fig, ax = plt.subplots()
ax.plot(testy_df['couplings'], testy_df['apc'], 'bo')
fig, ax = plt.subplots()
ax.plot(apc_df['couplings'], testy_df['apc'], 'bo')
stats.linregress(apc_df['couplings'], testy_df['apc'])
# **Ents**
from Bio.Alphabet.IUPAC import IUPACProtein
IUPACProtein.letters
def sequence_entropy_from_msa(msa_file, weights_file=False, base=2, skip_gaps=False):
'''
This should calculate the (un)weighted sequence entropy directly from a fasta file.
If not provided, all weights are even. If provided, weights will be read directly
from the accompanying weights_file.
'''
alignment = SeqIO.parse(msa_file, format='fasta')
aln_mat = np.array([list(i.seq) for i in alignment])
#####################
#####################
for i in IUPACProtein.letters:
aln_mat = np.append(aln_mat, [list(i*aln_mat.shape[1])], axis=0)
#####################
#####################
aln_mat_T = aln_mat.T
if weights_file:
weights = np.genfromtxt(weights_file)
else:
weights = np.ones(aln_mat_T[0].shape)
initial_shape = aln_mat_T.shape
flat_seqs = aln_mat_T.flatten()
order, flat_array = np.unique(flat_seqs, return_inverse=True)
if '-' in order:
assert order[0] == '-'
else:
if skip_gaps:
skip_gaps = False
print('No gapped characters found in alignment, skip_gaps flag is meaningless')
replaced_seqs_T = flat_array.reshape(initial_shape)
ents_all = []
for aln_position in replaced_seqs_T:
if skip_gaps:
ents_all.append(stats.entropy(np.bincount(aln_position,\
weights=weights, minlength=21)[1:], base=base))
else:
ents_all.append(stats.entropy(np.bincount(aln_position,\
weights=weights, minlength=21), base=base))
return ents_all
# +
prot_name = '1aoeA'
testy_df = process_couplings.process_ccmpredpy('../Results/couplings/{}.raw.uniform.mat'.format(prot_name))
ent_df = process_couplings.process_ccmpredpy('../Results/couplings/{}.ent.uniform.mat'.format(prot_name))
msa_file = '../../Phylogenetic_couplings/Data/'\
'psicov150_aln_pdb/aln_fasta_max1k/{}.fasta'.format(prot_name)
emp_entropies = sequence_entropy_from_msa(msa_file, skip_gaps=False, base=2)
# msa_file = '../../Phylogenetic_couplings/Data/'\
# 'psicov150_aln_pdb/aln_fasta_max1k/{}.fasta'.format(prot_name)
# weights_file = '../../DCA_weighting/Data/{}.HH.test'.format(prot_name)
# emp_entropies = sequence_entropy_from_msa(msa_file,\
# weights_file=weights_file, skip_gaps=False, base=2)
testy_df['score_h'] = np.nan
testy_df['first_h'] = np.nan
testy_df['second_h'] = np.nan
for index in testy_df.index:
fr = emp_entropies[int(testy_df.loc[index]['aa1_loc'])-1]
sr = emp_entropies[int(testy_df.loc[index]['aa2_loc'])-1]
testy_df.set_value(index, 'score_h', (fr**(1/2))*(sr**(1/2)))
testy_df.set_value(index, 'first_h', fr)
testy_df.set_value(index, 'second_h', sr)
# ###########
alpha_1 = np.sum(testy_df['couplings']*testy_df['score_h'])
alpha_2 = np.sum(testy_df['first_h'] * testy_df['second_h'])
alpha = alpha_1/alpha_2
testy_df['couplings_ent'] = testy_df['couplings'] - (alpha * testy_df['score_h'])
# ###########
###########
# +
fig, ax = plt.subplots()
ax.plot(testy_df['couplings_ent'], ent_df['couplings'], 'bo')
stats.linregress(testy_df['couplings_ent'], ent_df['couplings'])
# -
df_contacts = pd.read_csv('../../Phylogenetic_couplings/Data/'
'psicov150_aln_pdb/pdb/{}_CB_contacts.csv'.format(prot_name), index_col=0)
df_contacts, df_contacts_stack = process_couplings.process_contacts_df(df_contacts, 1)
seq = list(SeqIO.parse('../../Phylogenetic_couplings/Data/'
'psicov150_aln_pdb/aln_fasta/{}.fasta'.format(prot_name), 'fasta'))[0]
seq = str(seq.seq)
df_merged = process_couplings.merge_contacts_couplings(df_contacts_stack, ent_df, seq)
df_merged['contact'] = df_merged['distance']<7.5
aupr = sklearn.metrics.average_precision_score(df_merged['contact'], df_merged['couplings'])
print(aupr)
hmmm = pd.concat([df_merged, testy_df['couplings_ent']],\
axis=1, join_axes=[df_merged.index])
hmmm.sort_values('couplings_ent', ascending=False, inplace=True)
aupr = sklearn.metrics.average_precision_score(hmmm['contact'], hmmm['couplings_ent'])
print(aupr)
|
Code/comparing_accuracies.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--NAVIGATION-->
# < [在时间序列上操作](03.11-Working-with-Time-Series.ipynb) | [目录](Index.ipynb) | [更多资源](03.13-Further-Resources.ipynb) >
#
# <a href="https://colab.research.google.com/github/wangyingsm/Python-Data-Science-Handbook/blob/master/notebooks/03.12-Performance-Eval-and-Query.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
#
# # High-Performance Pandas: eval() and query()
#
# # 高性能Pandas: eval() 和 query()
# > As we've already seen in previous sections, the power of the PyData stack is built upon the ability of NumPy and Pandas to push basic operations into C via an intuitive syntax: examples are vectorized/broadcasted operations in NumPy, and grouping-type operations in Pandas.
# While these abstractions are efficient and effective for many common use cases, they often rely on the creation of temporary intermediate objects, which can cause undue overhead in computational time and memory use.
#
# 前面的章节中,我们已经了解了PyData的整个技术栈建立在NumPy和Pandas能将基础的向量化运算使用C底层的方式实现,语法却依然保持简单和直观:例子包括NumPy中的向量化和广播操作,及Pandas的分组类型的操作。虽然这些抽象在很多通用场合下是非常高效的,但是这些操作都涉及到创建临时对象,仍然会产生额外的计算时间和内存占用。
#
# > As of version 0.13 (released January 2014), Pandas includes some experimental tools that allow you to directly access C-speed operations without costly allocation of intermediate arrays.
# These are the ``eval()`` and ``query()`` functions, which rely on the [Numexpr](https://github.com/pydata/numexpr) package.
# In this notebook we will walk through their use and give some rules-of-thumb about when you might think about using them.
#
# Pandas在0.13版本(2014年1月发布)加入了一些实验性的工具,能直接进行C底层的运算而不需要创建临时的数组。函数`eval()`和`query()`具有这个特性,底层是基于[Numexpr](https://github.com/pydata/numexpr)包构建的。在本节中,我们会简单介绍它们的使用,然后给出何时适合使用它们的基础规则。
# ## Motivating ``query()`` and ``eval()``: Compound Expressions
#
# ## 使用 `query()` 和 `eval()` :复合表达式
#
# > We've seen previously that NumPy and Pandas support fast vectorized operations; for example, when adding the elements of two arrays:
#
# 我们已经掌握了NumPy和Pandas能够支持快速向量化操作;例如,当将两个数组进行加法操作时:
import numpy as np
rng = np.random.RandomState(42)
x = rng.rand(1000000)
y = rng.rand(1000000)
# %timeit x + y
# > As discussed in [Computation on NumPy Arrays: Universal Functions](02.03-Computation-on-arrays-ufuncs.ipynb), this is much faster than doing the addition via a Python loop or comprehension:
#
# 我们在[使用Numpy计算:通用函数](02.03-Computation-on-arrays-ufuncs.ipynb)中已经讨论过,这种运算对比使用Python循环或列表解析的方法要高效的多:
# %timeit np.fromiter((xi + yi for xi, yi in zip(x, y)), dtype=x.dtype, count=len(x))
# > But this abstraction can become less efficient when computing compound expressions.
# For example, consider the following expression:
#
# 但是当运算变得复杂的情况下,这种向量化运算就会变得没那么高效了。如下例:
mask = (x > 0.5) & (y < 0.5)
# > Because NumPy evaluates each subexpression, this is roughly equivalent to the following:
#
# 因为NumPy会独立计算每一个子表达式,因此上面代码等同与下面:
tmp1 = (x > 0.5)
tmp2 = (y < 0.5)
mask = tmp1 & tmp2
# > In other words, *every intermediate step is explicitly allocated in memory*. If the ``x`` and ``y`` arrays are very large, this can lead to significant memory and computational overhead.
# The Numexpr library gives you the ability to compute this type of compound expression element by element, without the need to allocate full intermediate arrays.
# The [Numexpr documentation](https://github.com/pydata/numexpr) has more details, but for the time being it is sufficient to say that the library accepts a *string* giving the NumPy-style expression you'd like to compute:
#
# 换言之,*每个中间步骤都会显式分配内存*。如果`x`和`y`数组变得非常巨大,这会带来显著的内存和计算资源开销。Numexpr库提供了既能使用简单语法进行数组的逐元素运算的能力,又不需要为中间步骤数组分配全部内存的能力。[Numexpr在线文档](https://github.com/pydata/numexpr)中有更加详细的说明,我们现在只需要将它理解为这个库能接受一个NumPy风格的表达式字符串,然后计算得到结果:
import numexpr
mask_numexpr = numexpr.evaluate('(x > 0.5) & (y < 0.5)')
np.allclose(mask, mask_numexpr)
# > The benefit here is that Numexpr evaluates the expression in a way that does not use full-sized temporary arrays, and thus can be much more efficient than NumPy, especially for large arrays.
# The Pandas ``eval()`` and ``query()`` tools that we will discuss here are conceptually similar, and depend on the Numexpr package.
#
# 这样做的优点是,Numexpr使用的临时数组不是完全分配空间的,并利用这少量数组即能完成计算,因此能比NumPy更加高效,特别是对大的数组来说。我们将会讨论到的Pandas的`eval()`和`query`工具,就是基于Numexpr包构建的。
# ## ``pandas.eval()`` for Efficient Operations
#
# ## `pandas.eval()` 更加高效的运算
#
# > The ``eval()`` function in Pandas uses string expressions to efficiently compute operations using ``DataFrame``s.
# For example, consider the following ``DataFrame``s:
#
# Pandas中的`eval()`函数可以使用字符串类型的表达式对`DataFrame`进行运算。例如,创建下面的`DataFrame`:
import pandas as pd
nrows, ncols = 100000, 100
rng = np.random.RandomState(42)
df1, df2, df3, df4 = (pd.DataFrame(rng.rand(nrows, ncols))
for i in range(4))
# > To compute the sum of all four ``DataFrame``s using the typical Pandas approach, we can just write the sum:
#
# 要计算所有四个`DataFrame`的总和,使用典型的Pandas方式,我们只需要将它们相加:
# %timeit df1 + df2 + df3 + df4
# > The same result can be computed via ``pd.eval`` by constructing the expression as a string:
#
# 我们也可以使用`pd.eval`,参数传入上述表达式的字符串形式,计算得到同样的结果:
# %timeit pd.eval('df1 + df2 + df3 + df4')
# > The ``eval()`` version of this expression is about 50% faster (and uses much less memory), while giving the same result:
#
# `eval()`版本的计算比典型方法快了接近接近50%(而且使用了更少的内存),我们来使用`np.allclose()`函数验证一下结果是否相同:
#
# 译者注:50%是按照原文翻译的,在译者自己笔记本上`eval()`的运行时间是典型方式的不到一半,运算速度应该是提高了100%多。
np.allclose(df1 + df2 + df3 + df4,
pd.eval('df1 + df2 + df3 + df4'))
# ### Operations supported by ``pd.eval()``
#
# ### `pd.eval()`支持的运算
#
# > As of Pandas v0.16, ``pd.eval()`` supports a wide range of operations.
# To demonstrate these, we'll use the following integer ``DataFrame``s:
#
# 到了Pandas 0.16版本,`pd.eval()`支持很大范围的运算。我们使用下面的整数`DataFrame`来进行展示:
df1, df2, df3, df4, df5 = (pd.DataFrame(rng.randint(0, 1000, (100, 3)))
for i in range(5))
# #### Arithmetic operators
#
# #### 算术运算
#
# > ``pd.eval()`` supports all arithmetic operators. For example:
#
# `pd.eval()`支持所有的算术运算。例如:
result1 = -df1 * df2 / (df3 + df4) - df5
result2 = pd.eval('-df1 * df2 / (df3 + df4) - df5')
np.allclose(result1, result2)
# #### Comparison operators
#
# #### 比较运算
#
# > ``pd.eval()`` supports all comparison operators, including chained expressions:
#
# `pd.eval()`支持所有的比较运算,包括链式表达式:
result1 = (df1 < df2) & (df2 <= df3) & (df3 != df4)
result2 = pd.eval('df1 < df2 <= df3 != df4')
np.allclose(result1, result2)
# #### Bitwise operators
#
# #### 位运算
#
# > ``pd.eval()`` supports the ``&`` and ``|`` bitwise operators:
#
# `pd.eval()`支持与`&`以及或`|`位运算符:
#
# 译者注:还支持非`~`位运算符。
result1 = (df1 < 0.5) & (df2 < 0.5) | (df3 < df4)
result2 = pd.eval('(df1 < 0.5) & (df2 < 0.5) | (df3 < df4)')
np.allclose(result1, result2)
# > In addition, it supports the use of the literal ``and`` and ``or`` in Boolean expressions:
#
# 而且,(译者注:对比NumPy)它还支持Python的在布尔表达式中使用逻辑运算`and`和`or`:
#
# 译者注:还支持`not`逻辑运算。
result3 = pd.eval('(df1 < 0.5) and (df2 < 0.5) or (df3 < df4)')
np.allclose(result1, result3)
# #### Object attributes and indices
#
# #### 对象属性和索引
#
# > ``pd.eval()`` supports access to object attributes via the ``obj.attr`` syntax, and indexes via the ``obj[index]`` syntax:
#
# `pd.eval()`支持使用`obj.attr`语法获取对象属性,也支持使用`obj[index]`语法进行索引:
result1 = df2.T[0] + df3.iloc[1]
result2 = pd.eval('df2.T[0] + df3.iloc[1]')
np.allclose(result1, result2)
# #### Other operations
#
# #### 其他运算
#
# > Other operations such as function calls, conditional statements, loops, and other more involved constructs are currently *not* implemented in ``pd.eval()``.
# If you'd like to execute these more complicated types of expressions, you can use the Numexpr library itself.
#
# 其他运算例如函数调用、条件语句、循环以及其他混合结构目前都不被`pd.eval()`支持。如果你需要使用这种复杂的表达式,你可以使用Numexpr库本身。
# ## ``DataFrame.eval()`` for Column-Wise Operations
#
# ## `DataFrame.eval()` 操作列
#
# > Just as Pandas has a top-level ``pd.eval()`` function, ``DataFrame``s have an ``eval()`` method that works in similar ways.
# The benefit of the ``eval()`` method is that columns can be referred to *by name*.
# We'll use this labeled array as an example:
#
# Pandas有着顶层的`pd.eval()`函数,`DataFrame`也有自己的`eval()`方法,实现的功能类似。使用`eval()`方法的好处是可以使用*列名*指代列。我们使用下面的带列标签的数组作为例子说明:
df = pd.DataFrame(rng.rand(1000, 3), columns=['A', 'B', 'C'])
df.head()
# > Using ``pd.eval()`` as above, we can compute expressions with the three columns like this:
#
# 使用上面的`pd.eval()`,我们可以如下计算三个列的结果:
result1 = (df['A'] + df['B']) / (df['C'] - 1)
result2 = pd.eval("(df.A + df.B) / (df.C - 1)")
np.allclose(result1, result2)
# > The ``DataFrame.eval()`` method allows much more succinct evaluation of expressions with the columns:
#
# 使用`DataFrame.eval()`方法允许我们采用更加直接的方式操作列数据:
result3 = df.eval('(A + B) / (C - 1)')
np.allclose(result1, result3)
# > Notice here that we treat *column names as variables* within the evaluated expression, and the result is what we would wish.
#
# 上面的代码中我们在表达式中将*列名作为变量*来使用,而且结果也是一致的。
# ### Assignment in DataFrame.eval()
#
# ### DataFrame.eval() 中的赋值
#
# > In addition to the options just discussed, ``DataFrame.eval()`` also allows assignment to any column.
# Let's use the ``DataFrame`` from before, which has columns ``'A'``, ``'B'``, and ``'C'``:
#
# 除了上面的操作外,`DataFrame.eval()`也支持对任何列的赋值操作。还是使用上面的`DataFrame`,有着`A`、`B`和`C`三个列:
df.head()
# > We can use ``df.eval()`` to create a new column ``'D'`` and assign to it a value computed from the other columns:
#
# 我们可以使用`df.eval()`方法类创建一个新的列`'D'`,然后将它赋值为其他列运算结果:
df.eval('D = (A + B) / C', inplace=True)
df.head()
# > In the same way, any existing column can be modified:
#
# 同样的,已经存在的列可以被修改:
df.eval('D = (A - B) / C', inplace=True)
df.head()
# ### Local variables in DataFrame.eval()
#
# ### DataFrame.eval()中的本地变量
#
# > The ``DataFrame.eval()`` method supports an additional syntax that lets it work with local Python variables.
# Consider the following:
#
# `DataFrame.eval()`方法还支持使用脚本中的本地Python变量。见下例:
column_mean = df.mean(1)
result1 = df['A'] + column_mean
result2 = df.eval('A + @column_mean')
np.allclose(result1, result2)
# > The ``@`` character here marks a *variable name* rather than a *column name*, and lets you efficiently evaluate expressions involving the two "namespaces": the namespace of columns, and the namespace of Python objects.
# Notice that this ``@`` character is only supported by the ``DataFrame.eval()`` *method*, not by the ``pandas.eval()`` *function*, because the ``pandas.eval()`` function only has access to the one (Python) namespace.
#
# 上面的字符串表达式中的`@`符号表示的是一个*变量名称*而不是一个*列名*,这个表达式能高效的计算涉及列空间和Python对象空间的运算表达式。需要注意的是`@`符号只能在`DataFrame.eval()`方法中使用,不能在`pandas.eval()`函数中使用,因为`pandas.eval()`实际上只有一个命名空间。
# ## DataFrame.query() Method
#
# ## DataFrame.query() 方法
#
# > The ``DataFrame`` has another method based on evaluated strings, called the ``query()`` method.
# Consider the following:
#
# `DataFrame`还有另外一个方法也是建立在字符串表达式运算的基础上的,就是`query()`。看下面这个例子:
result1 = df[(df.A < 0.5) & (df.B < 0.5)]
result2 = pd.eval('df[(df.A < 0.5) & (df.B < 0.5)]')
np.allclose(result1, result2)
# > As with the example used in our discussion of ``DataFrame.eval()``, this is an expression involving columns of the ``DataFrame``.
# It cannot be expressed using the ``DataFrame.eval()`` syntax, however!
# Instead, for this type of filtering operation, you can use the ``query()`` method:
#
# 根据前面的例子和讨论,这是一个涉及`DataFrame`列的表达式。但是它却不能使用`DataFrame.eval()`来实现。在这种情况下,你可以使用`query()`方法:
result2 = df.query('A < 0.5 and B < 0.5')
np.allclose(result1, result2)
# > In addition to being a more efficient computation, compared to the masking expression this is much easier to read and understand.
# Note that the ``query()`` method also accepts the ``@`` flag to mark local variables:
#
# 除了提供更加高效的计算外,这种语法比遮盖数组的方式更加容易读明白。而且`query()`方法也接受`@`符号来标记本地变量:
Cmean = df['C'].mean()
result1 = df[(df.A < Cmean) & (df.B < Cmean)]
result2 = df.query('A < @Cmean and B < @Cmean')
np.allclose(result1, result2)
# ## Performance: When to Use These Functions
#
# ## 性能:什么时候选择使用这些函数
#
# > When considering whether to use these functions, there are two considerations: *computation time* and *memory use*.
# Memory use is the most predictable aspect. As already mentioned, every compound expression involving NumPy arrays or Pandas ``DataFrame``s will result in implicit creation of temporary arrays:
# For example, this:
#
# 是否使用这些函数主要取决与两个考虑:*计算时间*和*内存占用*。其中最易预测的是内存使用。我们之前已经提到,每个基于NumPy数组的复合表达式都会在每个中间步骤产生一个临时数组,例如:
x = df[(df.A < 0.5) & (df.B < 0.5)]
# > Is roughly equivalent to this:
#
# 等同于:
tmp1 = df.A < 0.5
tmp2 = df.B < 0.5
tmp3 = tmp1 & tmp2
x = df[tmp3]
# > If the size of the temporary ``DataFrame``s is significant compared to your available system memory (typically several gigabytes) then it's a good idea to use an ``eval()`` or ``query()`` expression.
# You can check the approximate size of your array in bytes using this:
#
# 如果产生的临时的`DataFrame`与你可用的系统内存容量在同一个量级(如数GB)的话,那么使用`eval()`或者`query()`表达式显然是个好主意。可以通过数组的nbytes属性查看大概的内存占用:
df.values.nbytes
# > On the performance side, ``eval()`` can be faster even when you are not maxing-out your system memory.
# The issue is how your temporary ``DataFrame``s compare to the size of the L1 or L2 CPU cache on your system (typically a few megabytes in 2016); if they are much bigger, then ``eval()`` can avoid some potentially slow movement of values between the different memory caches.
# In practice, I find that the difference in computation time between the traditional methods and the ``eval``/``query`` method is usually not significant–if anything, the traditional method is faster for smaller arrays!
# The benefit of ``eval``/``query`` is mainly in the saved memory, and the sometimes cleaner syntax they offer.
#
# 至于计算时间考虑,`eval()`即使在不考虑内存占用的情况下也可能会更快。造成这个差异的原因主要在于临时的`DataFrame`的大小与计算机CPU的L1和L2缓存大小(在2016年通常是几个MB)的比值;如果缓存相比而言足够大的话,那么`eval()`可以避免在内存和CPU缓存之间的数据复制开销。在实践中,作者发现使用传统方式和`eval`/`query`方法之间的计算时间差异通常很小,如果存在的话,传统方法在小尺寸数组的情况下甚至还更快。因此`eval`/`query`的优势主要在于节省内存和它们的语法会更加清晰易懂。
#
# > We've covered most of the details of ``eval()`` and ``query()`` here; for more information on these, you can refer to the Pandas documentation.
# In particular, different parsers and engines can be specified for running these queries; for details on this, see the discussion within the ["Enhancing Performance" section](http://pandas.pydata.org/pandas-docs/dev/enhancingperf.html).
#
# 我们在本节讨论了`eval()`和`query()`的大部分内容;要获取更多相关资源,请参考Pandas的在线文档。特别的,其他不同的解析器和引擎也可以指定运行这些表达式和查询;有关内容参见[性能增强章节](http://pandas.pydata.org/pandas-docs/dev/enhancingperf.html)中的说明。
# <!--NAVIGATION-->
# < [在时间序列上操作](03.11-Working-with-Time-Series.ipynb) | [目录](Index.ipynb) | [更多资源](03.13-Further-Resources.ipynb) >
#
# <a href="https://colab.research.google.com/github/wangyingsm/Python-Data-Science-Handbook/blob/master/notebooks/03.12-Performance-Eval-and-Query.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
#
|
Python-Data-Science-Handbook/notebooks/03.12-Performance-Eval-and-Query.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
min_src_nsents = float("inf")
max_src_nsents = 0
min_src_ntokens_per_sent = float("inf")
max_src_ntokens_per_sent = 0
min_tgt_ntokens = float("inf")
max_tgt_ntokens = 0
src_nsents = []
src_ntokens_per_sent = []
tgt_ntokens = []
# +
corpus_type = "train"
count = 0
path = "token/AESLC_{}_token_filted".format(corpus_type)
sub_path = path+"_sub.txt"
doc_path = path+"_doc.txt"
reader_sub = open(sub_path)
reader_doc = open(doc_path)
while True:
doc_token = reader_doc.readline().replace("\n","")
sub_token = reader_sub.readline().replace("\n","")
if not doc_token:
break
if doc_token == "" or sub_token == "":
count += 1
continue
doc_sent = doc_token.split("<sentsep>")
doc_sent_numb = len(doc_sent)
src_nsents.append(doc_sent_numb)
for sent in doc_sent:
sent_toke_numb = len(" ".join(sent.split("<tokesep>")).split())
src_ntokens_per_sent.append(sent_toke_numb)
sub_toke = " ".join(sub_token.replace("<sentsep>","<tokesep>").split("<tokesep>")).split()
tgt_ntokens.append(len(sub_toke))
reader_doc.close()
reader_sub.close()
# +
import matplotlib.pyplot as plt
import numpy as np
x = np.random.randn(1000)
plt.boxplot(tgt_ntokens)
# plt.xticks([1], ["随机数生成器AlphaRM"])
# plt.ylabel("随机数值")
# plt.title("随机数生成器抗干扰能力的稳定性")
# plt.grid(axis="y", ls=":", lw=1, color="gray", alpha=0.4)
plt
# -
plt.show()
(0,40)
min(src_nsents), max(src_nsents)
(0, 40)
min(src_ntokens_per_sent), max(src_ntokens_per_sent)
(0,15)
min(tgt_ntokens), max(tgt_ntokens)
|
dataset/aeslc_name_char/stat.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Else if, for Brief - ELIF
# Assign 200 to x.
# Create the following piece of code:
# If x > 200, print out "Big";
# If x > 100 and x <= 200, print out "Average";
# and If x <= 100, print out "Small".
# Use the If, Elif, and Else keywords in your code.
x=500
if x>200 and x<=200:
print("Average")
elif x>200:
print("Big")
else:
print("Small")
# Change the initial value of x to see how your output will vary.
# ******
# Keep the first two conditions of the previous code. Add a new ELIF statement, so that, eventually, the program prints "Small" if x >= 0 and x <= 100, and "Negative" if x < 0.
# Let x carry the value of 50 and then of -50 to check if your code is correct.
x=-50
if x>200 and x<=200:
print("Average")
elif x>200:
print("Big")
elif x>=0 and x<=100:
print("Small")
elif x<0:
print("Negative")
|
Python basics practice/Python 3 (12)/Else If, for Brief - Elif - Exercise_Py3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.7 64-bit
# name: python387jvsc74a57bd01baa965d5efe3ac65b79dfc60c0d706280b1da80fedb7760faf2759126c4f253
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
admin_data = pd.read_csv("data/Admission_Predict.csv")
admin_data.head()
admin_data.columns
admin_data.drop(['TOEFL Score', "SOP", "LOR ", "University Rating"], axis=1, inplace=True)
sns.pairplot(admin_data, diag_kind = "hist", height=2)
plt.show()
sns.pairplot(admin_data, diag_kind="kde", height=2)
plt.show()
sns.pairplot(admin_data, hue="Research", diag_kind="kde", height=2)
plt.show()
# +
iris_data = sns.load_dataset("iris")
iris_data.sample()
# -
grid = sns.PairGrid(iris_data, aspect=1.5)
grid.map(plt.scatter)
plt.show()
grid = sns.PairGrid(iris_data, vars=["petal_length", "petal_width"], aspect=1.5)
grid.map(plt.scatter)
plt.show()
# +
grid = sns.PairGrid(iris_data, hue="species", aspect=1.5)
grid.map_diag(plt.hist)
grid.map_offdiag(plt.scatter)
grid.add_legend()
plt.show()
# -
|
Skill_Paths/Data_Science_Literacy/Summarizing_Data_and_Deducing_Probabilities/PairwiseRelationshipsUsingPairplotAndPairGrid.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import numpy as np
import pandas as pd
import seaborn as sns
from sklearn.cluster import KMeans
from sklearn.decomposition import PCA
from wakeful import log_munger, metrics, virus_total
# %matplotlib inline
# %pwd
# ### Build the Normal Data Set
norm_path = './data/home/2018-01-01'
print(dns_norm_df.shape)
dns_norm_df.info()
conn_norm_df = log_munger.bro_logs_to_df(norm_path, 'conn');
conn_norm_df.shape
conn_norm_df.info()
dns_norm_df.info()
dns_norm_df.head(5)
conn_norm_df.head(5)
# ### Merge Conn and DNS logs
norm_df = pd.merge(dns_norm_df, conn_norm_df, on='uid')
norm_df.shape
norm_df.info()
norm_df.to_hdf('./data/home/dns_and_conn_20180101.h5', 'test', complevel=9, complib='zlib')
# !ls -lh ./data/home/*.h5
# ### Round-trip from DF --> HDF5 --> DF
key = 'key_to_finding_data'
#log_munger.df_to_hdf5(norm_df, key, './data/home/')
data_dir = './data/home'
df = log_munger.hdf5_to_df(key, data_dir)
# +
#df.equals(norm_df)
# -
norm_df = df.copy(deep=True)
# ### Build the Malicious Data Set
mal_dns_df = log_munger.bro_logs_to_df('./data/c2', 'dns')
mal_conn_df = log_munger.bro_logs_to_df('./data/c2', 'conn')
mal_df = pd.merge(mal_dns_df, mal_conn_df, on='uid')
mal_dns_df.shape, mal_conn_df.shape, mal_df.shape
# ### Label the Data Sets
norm_df['label'] = 0
mal_df['label'] = 1
# ### Remove Inconsistent Columns
mal_col_set = set(mal_df.columns)
norm_col_set = set(norm_df.columns)
norm_columns_not_in_mal = {col_n for col_n in norm_col_set if col_n not in mal_col_set}
norm_columns_not_in_mal, norm_df.shape, mal_df.shape
mal_columns_not_in_norm = {col_m for col_m in mal_col_set if col_m not in norm_col_set}
mal_columns_not_in_norm, norm_df.shape, mal_df.shape
mal_df = mal_df.drop(mal_columns_not_in_norm, axis=1)
norm_df = norm_df.drop(norm_columns_not_in_mal, axis=1)
norm_df.shape, mal_df.shape
# ### Combine Labeled Data Sets
df = norm_df.append(mal_df)
df.shape
# ### Engineer Features
tmp = df['query'][:10]
tmp.apply(metrics.calc_entropy)
df['pcr'] = metrics.calc_pcr(df)
df['query_entropy'] = df['query'].apply(metrics.calc_entropy)
df.head()
# ### Save the Combined Labeled Data Set
key = 'dns_conn_labeled_20180101'
log_munger.df_to_hdf5(df, key, './data/home/')
# !ls -lh ./data/home/*.h5
# ### Read in the Combined Labeled Data Set
key = 'dns_conn_labeled_20180101'
df = log_munger.hdf5_to_df(key, './data/home/')
# ### Rebalance Minority Class
# from sklearn.model_selection import train_test_split
# train, test = train_test_split(df, random_state=37, test_size=0.5)
bal_df = log_munger.rebalance(df, column_name='label')
df.shape, bal_df.shape
bal_no_strings = bal_df.drop(['TTLs', 'answers', 'conn_state', 'history', 'id.orig_h_x',
'id.orig_h_y', 'id.resp_h_x', 'id.resp_h_y', 'proto_x', 'proto_y',
'qclass_name', 'qtype_name', 'query', 'rcode_name', 'service', 'tunnel_parents',
'uid', 'duration'], axis=1)
corr = bal_no_strings.corr()
mask = np.zeros_like(corr, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
f, ax = plt.subplots(figsize=(16, 12))
sns.set(style='white')
cmap = sns.diverging_palette(220, 10, as_cmap=True)
sns.heatmap(corr, mask=mask, cmap=cmap, vmax=3,
center=0, square=True, linewidths=.5,
cbar_kws={'shrink': .5})
bal_no_strings.columns
bal_eng = bal_no_strings.loc[:, ['pcr', 'query_entropy', 'label']]
bal_eng.head(5)
bal = bal_eng.dropna(axis=0, how='any')
bal_eng.shape, bal.shape
sns.pairplot(bal, hue='label')
sns.set(color_codes=True)
g = sns.lmplot(x="pcr", y="query_entropy", hue="label",
truncate=True, size=8, data=bal)
g.set_axis_labels("Producer Consumer Ratio", "Query Entropy")
# ### More Feature Engineering
conn_logs_df = log_munger.bro_logs_to_df('./data/home/2017-12-31', 'conn');
conn_logs_df.columns
ips = conn_logs_df['id.resp_h'].unique()
conn_logs_df.tail(100)
conn_logs_df['conn_in'] = 0
conn_logs_df['conn_in'] = conn_logs_df[conn_logs_df['id.resp_h'] == '172.16.58.3']['id.resp_h'].rolling(window=8).count()
conn_logs_df[['id.resp_h', 'conn_in']]
from bat.dataframe_to_matrix import DataFrameToMatrix
to_matrix = DataFrameToMatrix()
conn_matrix = to_matrix.fit_transform(conn_logs_df)
# ### KMeans Clustering
kmeans = KMeans(n_clusters=5).fit_predict(conn_matrix)
# ### PCA Decomposition
pca = PCA(n_components=5)
pca.fit(conn_matrix)
print(sum(pca.explained_variance_ratio_), '=', pca.explained_variance_ratio_)
print(pca.n_components_)
X = bal.copy(deep=True)
y = X.pop('label')
# ### DBSCAN Clustering
from sklearn.preprocessing import StandardScaler
from sklearn.cluster import DBSCAN
from sklearn import metrics
bal.columns
X = bal.copy(deep=True)
y = bal.pop('label')
X = StandardScaler().fit_transform(X)
db = DBSCAN(eps=0.3, min_samples=10).fit(X)
# +
core_samples_mask = np.zeros_like(db.labels_, dtype=bool)
core_samples_mask[db.core_sample_indices_] = True
y_pred = db.labels_
# Number of clusters in labels, ignoring noise if present.
n_clusters_ = len(set(y_pred)) - (1 if -1 in y_pred else 0)
print('Estimated number of clusters: %d' % n_clusters_)
print("Homogeneity: %0.3f" % metrics.homogeneity_score(y, y_pred))
print("Completeness: %0.3f" % metrics.completeness_score(y, y_pred))
print("V-measure: %0.3f" % metrics.v_measure_score(y, y_pred))
print("Adjusted Rand Index: %0.3f"
% metrics.adjusted_rand_score(y, y_pred))
print("Adjusted Mutual Information: %0.3f"
% metrics.adjusted_mutual_info_score(y, y_pred))
print("Silhouette Coefficient: %0.3f"
% metrics.silhouette_score(X, y_pred))
# +
# # DON'T RUN -- LOTS OF CPU CYCLES
# # Plot result
# import matplotlib.pyplot as plt
# # Black removed and is used for noise instead.
# unique_labels = set(y_pred)
# colors = [plt.cm.Spectral(each)
# for each in np.linspace(0, 1, len(unique_labels))]
# for k, col in zip(unique_labels, colors):
# if k == -1:
# # Black used for noise.
# col = [0, 0, 0, 1]
# class_member_mask = (labels == k)
# xy = X[class_member_mask & core_samples_mask]
# plt.plot(xy[:, 0], xy[:, 1], 'o', markerfacecolor=tuple(col),
# markeredgecolor='k', markersize=14)
# xy = X[class_member_mask & ~core_samples_mask]
# plt.plot(xy[:, 0], xy[:, 1], 'o', markerfacecolor=tuple(col),
# markeredgecolor='k', markersize=6)
# plt.title('Estimated number of clusters: %d' % n_clusters_)
# plt.show()
# -
# ### SVD and Logistic Regression
import numpy as np
import matplotlib.pyplot as plt
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn import decomposition
from sklearn import linear_model
# +
scaler = StandardScaler()
svd = decomposition.TruncatedSVD(n_components=15, random_state=37, n_iter=10)
logistic = linear_model.LogisticRegression()
pca = decomposition.PCA(n_components=10, random_state=37)
pipeline = Pipeline(steps=[('scaler', scaler), ('pca', pca), ('svd', svd), ('logistic', logistic)])
# -
numerics = X_train.copy(deep=True)
numerics = numerics.drop(['TTLs', 'answers', 'conn_state', 'history', 'id.orig_h_x',
'id.orig_h_y', 'id.resp_h_x', 'id.resp_h_y', 'proto_x', 'proto_y',
'qclass_name', 'qtype_name', 'query', 'rcode_name', 'service', 'tunnel_parents',
'uid'], axis=1)
# Convert timedeltas to seconds
numerics = numerics.drop(['duration'], axis=1)
x_temp = numerics.values
type(numerics), type(x_temp), x_temp.shape
numerics.info()
# get the data
X_digits = numerics.values
y_digits = y_train.values
type(X_digits)
X_digits[1:5, :].T
row = X_digits[0, :]
print(row)
for i in range(X_digits.shape[1]):
print(type(X_digits[:, i]))
indx = np.where(np.isnan(X_digits))
# +
# Plot the PCA spectrum
pca.fit(X_digits)
plt.figure(1, figsize=(4, 3))
plt.clf()
plt.axes([.2, .2, .7, .7])
plt.plot(svd.explained_variance_, linewidth=2)
plt.axis('tight')
plt.xlabel('n_components')
plt.ylabel('explained_variance_')
# Prediction
n_components = [20, 40, 64]
Cs = np.logspace(-4, 4, 3)
# Parameters of pipelines can be set using ‘__’ separated parameter names:
estimator = GridSearchCV(pipe,
dict(svd__n_components=n_components,
logistic__C=Cs))
estimator.fit(X_digits, y_digits)
plt.axvline(estimator.best_estimator_.named_steps['svd'].n_components,
linestyle=':', label='n_components chosen')
plt.legend(prop=dict(size=12))
plt.show()
# -
|
bro_logs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# This cell is added by sphinx-gallery
# !pip install mrsimulator --quiet
# %matplotlib inline
import mrsimulator
print(f'You are using mrsimulator v{mrsimulator.__version__}')
# -
#
# # Itraconazole, ¹³C (I=1/2) PASS
#
# ¹³C (I=1/2) 2D Phase-adjusted spinning sideband (PASS) simulation.
#
# The following is a simulation of a 2D PASS spectrum of itraconazole, a triazole
# containing drug prescribed for the prevention and treatment of fungal infection.
# The 2D PASS spectrum is a correlation of finite speed MAS to an infinite speed MAS
# spectrum. The parameters for the simulation are obtained from Dey `et al.` [#f1]_.
#
#
# +
import matplotlib.pyplot as plt
from mrsimulator import Simulator
from mrsimulator.methods import SSB2D
from mrsimulator import signal_processing as sp
# -
# There are 41 $^{13}\text{C}$ single-site spin systems partially describing the
# NMR parameters of itraconazole. We will import the directly import the spin systems
# to the Simulator object using the `load_spin_systems` method.
#
#
# +
sim = Simulator()
filename = "https://sandbox.zenodo.org/record/687656/files/itraconazole_13C.mrsys"
sim.load_spin_systems(filename)
# -
# Use the ``SSB2D`` method to simulate a PASS, MAT, QPASS, QMAT, or any equivalent
# sideband separation spectrum. Here, we use the method to generate a PASS spectrum.
#
#
# +
PASS = SSB2D(
channels=["13C"],
magnetic_flux_density=11.74,
rotor_frequency=2000,
spectral_dimensions=[
{
"count": 20 * 4,
"spectral_width": 2000 * 20, # value in Hz
"label": "Anisotropic dimension",
},
{
"count": 1024,
"spectral_width": 3e4, # value in Hz
"reference_offset": 1.1e4, # value in Hz
"label": "Isotropic dimension",
},
],
)
sim.methods = [PASS] # add the method.
# A graphical representation of the method object.
plt.figure(figsize=(5, 3.5))
PASS.plot()
plt.show()
# -
# For 2D spinning sideband simulation, set the number of spinning sidebands in the
# Simulator.config object to `spectral_width/rotor_frequency` along the sideband
# dimension.
#
#
# +
sim.config.number_of_sidebands = 20
# run the simulation.
sim.run()
# -
# Apply post-simulation processing. Here, we apply a Lorentzian line broadening to the
# isotropic dimension.
#
#
data = sim.methods[0].simulation
processor = sp.SignalProcessor(
operations=[
sp.IFFT(dim_index=0),
sp.apodization.Exponential(FWHM="100 Hz", dim_index=0),
sp.FFT(dim_index=0),
]
)
processed_data = processor.apply_operations(data=data).real
processed_data /= processed_data.max()
# The plot of the simulation.
#
#
plt.figure(figsize=(4.25, 3.0))
ax = plt.subplot(projection="csdm")
cb = ax.imshow(processed_data, aspect="auto", cmap="gist_ncar_r", vmax=0.5)
plt.colorbar(cb)
ax.invert_xaxis()
ax.invert_yaxis()
plt.tight_layout()
plt.show()
# .. [#f1] <NAME> .K, <NAME>., <NAME>., Investigation of the Detailed Internal
# Structure and Dynamics of Itraconazole by Solid-State NMR Measurements,
# ACS Omega (2019) **4**, 21627.
# `DOI:10.1021/acsomega.9b03558 <https://doi.org/10.1021/acsomega.9b03558>`_
#
#
|
docs/notebooks/examples/2D_simulation(crystalline)/plot_6_PASS_itraconazole_drug.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# +
import os
import pandas as pd
import json
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
from datetime import datetime, date, timedelta
from node2vec_utils import read_node_vecs, load_json_dict, invert_dict
from suspicion_tools import cosine_sim, cycle_suspicion_desc, cycle_suspicion_score, cycle_suspicion_for_agg
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import sys
if sys.version_info[0] < 3:
from StringIO import StringIO
else:
from io import StringIO
# -
transactions = pd.read_csv('../../data/transactions.small.csv')
transactions['source'] = transactions['source'].astype(str)
transactions['target'] = transactions['target'].astype(str)
#transactions['date'] = pd.to_datetime(transactions.date)
#transactions['time'] = pd.to_datetime(transactions['time'])
transactions['amount'] = transactions.amount.astype(float)
transactions
clients = pd.read_csv('../../data/clients.small.csv')
companies = pd.read_csv('../../data/companies.small.csv')
loopsjson = pd.read_json('loops.json', lines=True)
#Generate jsons for suspects.
for i in range(loopsjson.shape[0]):
all_ids = []
all_link_ids = []
current_suspect = loopsjson.loc[i,0]['from']['id']
f = open('../../json/'+current_suspect+'.json', mode='w')
current_dict = {}
current_nodes = []
current_links = []
for j in range(loopsjson.shape[1]):
if loopsjson.loc[i,j] is not None:
#Adding the from and to nodes
curr_node = {}
curr_node['id'] = loopsjson.loc[i,j]['from']['id']
#print(curr_node)
if (curr_node['id'] not in all_ids):
if j==0:
curr_node['tag'] = 'suspect'
else:
curr_node['tag'] = 'accomplice'
all_ids.append(curr_node['id'])
current_nodes.append(curr_node)
if (loopsjson.loc[i,j]['to']['id'] not in all_ids):
all_ids.append(loopsjson.loc[i,j]['to']['id'])
other_node = {}
other_node['id'] = loopsjson.loc[i,j]['to']['id']
other_node['tag'] = 'accomplice'
current_nodes.append(other_node)
current_link_id = loopsjson.loc[i,j]['id']
current_link = transactions[transactions.id == current_link_id].to_dict(orient='records')
current_link = current_link[0]
if (current_link_id not in all_link_ids):
current_link['tag'] = 'accomplice'
current_links.append(current_link)
all_link_ids.append(current_link_id)
if (current_link['target'] not in all_ids):
node = {}
node['tag'] = 'accomplice'
node['id'] = current_link['target']
current_nodes.append(node)
all_ids.append(node['id'])
if (current_link['source'] not in all_ids):
node = {}
node['tag'] = 'accomplice'
node['id'] = current_link['source']
current_nodes.append(node)
all_ids.append(node['id'])
if (j == 0):
root_node_id = loopsjson.loc[i,0]['from']['id']
outgoing_links = transactions[transactions.source == root_node_id]
if (outgoing_links.shape[0] >= 5):
sampled = outgoing_links.sample(n=5).drop_duplicates(subset=['id']).to_dict(orient='records')
else:
sampled = outgoing_links.drop_duplicates(subset=['id']).to_dict(orient='records')
for current_link in sampled:
if (current_link['id'] not in all_link_ids):
current_link['tag'] = 'usual'
current_links.append(current_link)
all_link_ids.append(current_link['id'])
if (current_link['target'] not in all_ids):
node = {}
node['tag'] = 'usual'
node['id'] = current_link['target']
#print(node)
current_nodes.append(node)
all_ids.append(node['id'])
if (current_link['source'] not in all_ids):
node = {}
node['tag'] = 'usual'
node['id'] = current_link['source']
#print(node)
current_nodes.append(node)
all_ids.append(node['id'])
current_dict['nodes'] = current_nodes
current_dict['links'] = current_links
json.dump(current_dict, f)
f.close()
# +
#Generate some example jsons for non-suspects.
# -
|
code/Ramtin/create_id_jsons.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # REINFORCE in pytorch
#
# Just like we did before for q-learning, this time we'll design a pytorch network to learn `CartPole-v0` via policy gradient (REINFORCE).
#
# Most of the code in this notebook is taken from approximate qlearning, so you'll find it more or less familiar and even simpler.
# +
# # in google colab uncomment this
# import os
# os.system('apt-get install -y xvfb')
# os.system('wget https://raw.githubusercontent.com/yandexdataschool/Practical_DL/fall18/xvfb -O ../xvfb')
# os.system('apt-get install -y python-opengl ffmpeg')
# os.system('pip install pyglet==1.2.4')
# os.system('python -m pip install -U pygame --user')
# print('setup complete')
# XVFB will be launched if you run on a server
import os
if type(os.environ.get("DISPLAY")) is not str or len(os.environ.get("DISPLAY")) == 0:
# !bash ../xvfb start
os.environ['DISPLAY'] = ':1'
# +
import gym
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
env = gym.make("CartPole-v0").env
env.reset()
plt.imshow(env.render("rgb_array"))
# -
# # Building the network for REINFORCE
# For REINFORCE algorithm, we'll need a model that predicts action probabilities given states. Let's define such a model below.
import torch
import torch.nn as nn
# Build a simple neural network that predicts policy logits.
# Keep it simple: CartPole isn't worth deep architectures.
model = nn.Sequential(
< YOUR CODE HERE: define a neural network that predicts policy logits >
)
# #### Predict function
def predict_probs(states):
"""
Predict action probabilities given states.
:param states: numpy array of shape [batch, state_shape]
:returns: numpy array of shape [batch, n_actions]
"""
# convert states, compute logits, use softmax to get probability
<your code here >
return < your code >
test_states = np.array([env.reset() for _ in range(5)])
test_probas = predict_probs(test_states)
assert isinstance(
test_probas, np.ndarray), "you must return np array and not %s" % type(test_probas)
assert tuple(test_probas.shape) == (
test_states.shape[0], env.action_space.n), "wrong output shape: %s" % np.shape(test_probas)
assert np.allclose(np.sum(test_probas, axis=1),
1), "probabilities do not sum to 1"
# ### Play the game
#
# We can now use our newly built agent to play the game.
def generate_session(t_max=1000):
"""
play a full session with REINFORCE agent and train at the session end.
returns sequences of states, actions andrewards
"""
# arrays to record session
states, actions, rewards = [], [], []
s = env.reset()
for t in range(t_max):
# action probabilities array aka pi(a|s)
action_probs = predict_probs(np.array([s]))[0]
# Sample action with given probabilities.
a = < your code >
new_s, r, done, info = env.step(a)
# record session history to train later
states.append(s)
actions.append(a)
rewards.append(r)
s = new_s
if done:
break
return states, actions, rewards
# test it
states, actions, rewards = generate_session()
# ### Computing cumulative rewards
def get_cumulative_rewards(rewards, # rewards at each step
gamma=0.99 # discount for reward
):
"""
take a list of immediate rewards r(s,a) for the whole session
compute cumulative returns (a.k.a. G(s,a) in Sutton '16)
G_t = r_t + gamma*r_{t+1} + gamma^2*r_{t+2} + ...
The simple way to compute cumulative rewards is to iterate from last to first time tick
and compute G_t = r_t + gamma*G_{t+1} recurrently
You must return an array/list of cumulative rewards with as many elements as in the initial rewards.
"""
<your code here >
return < array of cumulative rewards >
get_cumulative_rewards(rewards)
assert len(get_cumulative_rewards(list(range(100)))) == 100
assert np.allclose(get_cumulative_rewards([0, 0, 1, 0, 0, 1, 0], gamma=0.9), [
1.40049, 1.5561, 1.729, 0.81, 0.9, 1.0, 0.0])
assert np.allclose(get_cumulative_rewards(
[0, 0, 1, -2, 3, -4, 0], gamma=0.5), [0.0625, 0.125, 0.25, -1.5, 1.0, -4.0, 0.0])
assert np.allclose(get_cumulative_rewards(
[0, 0, 1, 2, 3, 4, 0], gamma=0), [0, 0, 1, 2, 3, 4, 0])
print("looks good!")
# #### Loss function and updates
#
# We now need to define objective and update over policy gradient.
#
# Our objective function is
#
# $$ J \approx { 1 \over N } \sum _{s_i,a_i} \pi_\theta (a_i | s_i) \cdot G(s_i,a_i) $$
#
#
# Following the REINFORCE algorithm, we can define our objective as follows:
#
# $$ \hat J \approx { 1 \over N } \sum _{s_i,a_i} log \pi_\theta (a_i | s_i) \cdot G(s_i,a_i) $$
#
# When you compute gradient of that function over network weights $ \theta $, it will become exactly the policy gradient.
#
def to_one_hot(y_tensor, ndims):
""" helper: take an integer vector and convert it to 1-hot matrix. """
y_tensor = y_tensor.type(torch.LongTensor).view(-1, 1)
y_one_hot = torch.zeros(
y_tensor.size()[0], ndims).scatter_(1, y_tensor, 1)
return y_one_hot
# +
# Your code: define optimizers
optimizer = torch.optim.Adam(model.parameters(), 1e-3)
def train_on_session(states, actions, rewards, gamma=0.99, entropy_coef=1e-2):
"""
Takes a sequence of states, actions and rewards produced by generate_session.
Updates agent's weights by following the policy gradient above.
Please use Adam optimizer with default parameters.
"""
# cast everything into torch tensors
states = torch.tensor(states, dtype=torch.float32)
actions = torch.tensor(actions, dtype=torch.int32)
cumulative_returns = np.array(get_cumulative_rewards(rewards, gamma))
cumulative_returns = torch.tensor(cumulative_returns, dtype=torch.float32)
# predict logits, probas and log-probas using an agent.
logits = model(states)
probs = nn.functional.softmax(logits, -1)
log_probs = nn.functional.log_softmax(logits, -1)
assert all(isinstance(v, torch.Tensor) for v in [logits, probs, log_probs]), \
"please use compute using torch tensors and don't use predict_probs function"
# select log-probabilities for chosen actions, log pi(a_i|s_i)
log_probs_for_actions = torch.sum(
log_probs * to_one_hot(actions, env.action_space.n), dim=1)
# Compute loss here. Don't forgen entropy regularization with `entropy_coef`
entropy = < your code >
loss = < your code
# Gradient descent step
< your code >
# technical: return session rewards to print them later
return np.sum(rewards)
# -
# ### The actual training
for i in range(100):
rewards = [train_on_session(*generate_session())
for _ in range(100)] # generate new sessions
print("mean reward:%.3f" % (np.mean(rewards)))
if np.mean(rewards) > 500:
print("You Win!") # but you can train even further
break
# ### Video
# record sessions
import gym.wrappers
env = gym.wrappers.Monitor(gym.make("CartPole-v0"),
directory="videos", force=True)
sessions = [generate_session() for _ in range(100)]
env.close()
# +
# show video
from IPython.display import HTML
import os
video_names = list(
filter(lambda s: s.endswith(".mp4"), os.listdir("./videos/")))
HTML("""
<video width="640" height="480" controls>
<source src="{}" type="video/mp4">
</video>
""".format("./videos/"+video_names[-1])) # this may or may not be the _last_ video. Try other indices
# -
|
week11_rl/reinforce_pytorch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Rigid Body Dynamics and Simulation
#
# Finally, let's see how to start assembling the tools mentioned in the Systems and Symbolic/Autodiff tutorials to make robots do interesting things.
#
# For these examples, we'll explore simulating the ultra-classic cart pole, pictured below.
#
# <img src="https://danielpiedrahita.files.wordpress.com/2017/02/cart-pole.png" alt="drawing" style="width: 400px;"/>
# As a more complete demo, we can create an LQR solution around that upright fixed point and simulate it!
#
# See the quickstart guide for a written explanation of the many pieces of this.
# +
import math
import matplotlib.pyplot as plt
import numpy as np
from pydrake.all import (BasicVector, DiagramBuilder, FloatingBaseType,
LinearQuadraticRegulator, RigidBodyPlant,
RigidBodyTree, Simulator, SignalLogger, LeafSystem, PortDataType)
from underactuated import (PlanarRigidBodyVisualizer)
from underactuated import (FindResource, PlanarRigidBodyVisualizer)
from IPython.display import HTML
# +
import pdb
class SwingupController(LeafSystem):
def __init__(self, rbt,
control_period=0.005,
print_period=0.5):
LeafSystem.__init__(self)
self.set_name("Swing up Controller")
#self.B_inv = np.linalg.inv(self.B)
# Copy lots of stuff
self.rbt = rbt
self.nq = rbt.get_num_positions()
#self.plant = plant
self.nu = rbt.get_input_port(0).size()
#self.print_period = print_period
self.last_print_time = -print_period
self.shut_up = False
self.robot_state_input_port = \
self._DeclareInputPort(PortDataType.kVectorValued,
rbt.get_num_positions() +
rbt.get_num_velocities())
self._DeclareContinuousState(self.nu)
#self._DeclarePeriodicContinuousUpdate(period_sec=control_period)
self._DeclareVectorOutputPort(
BasicVector(self.nu),
self._DoCalcVectorOutput)
def _DoCalcVectorOutput(self, context, y_data):
control_output = context.get_continuous_state_vector().get_value()
y = y_data.get_mutable_value()
# Get the ith finger control output
# y[:] = control_output[:]
y[:] = 0
# +
# #%tb
# #%pdb
# Load in the cartpole from its URDF
tree = RigidBodyTree(FindResource("cartpole/cartpole.urdf"),
FloatingBaseType.kFixed)
print tree
# Define an upright state
def UprightState():
state = (0,math.pi,0,0)
return state
def UprightPos():
state = (math.pi/2,0)
return state
def BalancingLQR(robot):
# Design an LQR controller for stabilizing the CartPole around the upright.
# Returns a (static) AffineSystem that implements the controller (in
# the original CartPole coordinates).
context = robot.CreateDefaultContext()
context.FixInputPort(0, BasicVector([0]))
context.get_mutable_continuous_state_vector().SetFromVector(UprightState())
Q = np.diag((10., 10.,1,1))
R = [1]
return LinearQuadraticRegulator(robot, context, Q, R)
builder = DiagramBuilder()
robot = builder.AddSystem(RigidBodyPlant(tree))
controller = builder.AddSystem(BalancingLQR(robot))
controller = builder.AddSystem(SwingupController(robot))
builder.Connect(robot.get_output_port(0), controller.get_input_port(0))
builder.Connect(controller.get_output_port(0), robot.get_input_port(0))
logger = builder.AddSystem(SignalLogger(robot.get_output_port(0).size()))
logger._DeclarePeriodicPublish(1. / 30., 0.0)
builder.Connect(robot.get_output_port(0), logger.get_input_port(0))
diagram = builder.Build()
simulator = Simulator(diagram)
simulator.set_publish_every_time_step(False)
context = simulator.get_mutable_context()
state = context.get_mutable_continuous_state_vector()
state.SetFromVector(UprightState() + 0.1*np.random.randn(4,))
simulator.StepTo(10.)
prbv = PlanarRigidBodyVisualizer(tree, xlim=[-2.5, 2.5], ylim=[-1, 2.5])
ani = prbv.animate(logger, resample=30, repeat=True)
plt.close(prbv.fig)
HTML(ani.to_html5_video())
# -
|
old/drake_examples/drake_rigid_body_simulation_tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from scripts.load_script import *
df = process_data('../raw/adult.data')
find_and_replace(df, 'Workclass', 'State-gov|Federal-gov|Local-gov', 'Government')
find_and_replace(df, 'Workclass', 'Never-worked|Without-pay|Other', '?')
find_and_replace(df, 'Workclass', 'Self-emp-not-inc|Self-emp-inc', 'Self-employed')
find_and_replace(df, 'Workclass', '?', 'Other')
find_and_replace(df, 'Education', '11th|9th|7th-8th|5th-6th|10th|1st-4th|12th|Preschool|Replaced', 'Didnt-grad-HS')
find_and_replace(df, 'Education', 'Some-college', 'Bachelors')
find_and_replace(df, 'Marital Status', 'Married-spouse-absent', 'Married-civ-spouse')
find_and_replace(df, 'Marital Status', 'Married-AF-spouse', 'Married-civ-spouse')
find_and_replace(df, 'Marital Status', 'Married-civ-spouse', 'Married')
find_and_replace(df, 'Occupation', '?', 'Other')
find_and_replace(df, 'Occupation', 'Other-service', 'Other')
find_and_replace(df, 'Occupation', 'Armed-Forces', 'Protective-serv')
df.to_csv('adult_processed.csv', index=False)
|
data/processed/.ipynb_checkpoints/Processing-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from jupyter_innotater import *
import numpy as np, os
# ### Image Filenames and Bounding Boxes
# +
foodfns = sorted(os.listdir('./foods/'))
targets = np.zeros((len(foodfns), 4), dtype='int') # (x,y,w,h) for each data row
Innotater( ImageInnotation(foodfns, path='./foods'), BoundingBoxInnotation(targets) )
# -
# Press 'n' or 'p' to move to next or previous image in the Innotater above.
targets
# Write our newly-input bounding box data to disk - will be lost otherwise
# If pandas not installed, please just ignore this cell
import pandas as pd
df = pd.DataFrame(targets, columns=['x','y','w','h'])
df.insert(0,'filename', foodfns)
df.to_csv('./bounding_boxes.csv')
df
# ### Numpy Image Data and Multi-classification
# +
try:
import cv2
foods = [cv2.imread('./foods/'+f) for f in foodfns]
except ModuleNotFoundError:
print("OpenCV2 is not installed, so just using filenames like before - Innotater will understand")
foods = ['./foods/'+f for f in foodfns]
classes = ['vegetable', 'biscuit', 'fruit']
targets = [0] * len(foodfns)
# -
w2 = Innotater(
ImageInnotation(foods, name='Food'),
MultiClassInnotation(targets, name='FoodType', classes=classes, desc='Food Type')
)
display(w2)
targets
# Convert targets from a 1-dim array to one-hot representation - Innotater works with that just as well
onehot_targets = np.squeeze(np.eye(len(classes))[np.array(targets).reshape(-1)]); onehot_targets
Innotater(
ImageInnotation(foods, name='Food'),
MultiClassInnotation(onehot_targets, name='FoodType', classes=classes, desc='Food Type')
)
# ### Filenames and binary classification
# Set image to display at a smaller width to make it more manageable - but bounding box co-ordinates would be relative to the unzoomed image.
isfruit_targets = (np.array(targets) == 2).astype('int')
Innotater( ImageInnotation(foodfns, path='./foods', width=300),
BinaryClassInnotation(isfruit_targets, name='Is Fruit')
)
isfruit_targets
# ### Image Filenames and Binary Classification plus Bounding Boxes
# Use indexes attribute to limit display just to the fruits where we want to add bounding boxes. Drop the indexes property if you also want to be able to check non-fruits.
# +
bboxes = np.zeros((len(foodfns),4), dtype='int')
isfruits = np.expand_dims(isfruit_targets, axis=-1)
suspected_fruits = isfruits == 1 # Or you can specify an array/list of int indices
Innotater(
ImageInnotation(foodfns, name='Food', path='./foods'),
[ BinaryClassInnotation(isfruits, name='Is Fruit'),
BoundingBoxInnotation(bboxes, name='bbs', source='Food', desc='Food Type') ],
indexes = suspected_fruits
)
# -
result = np.concatenate([isfruits,bboxes], axis=-1); result
# ### Image versus Image and Binary Classification
# +
targets = np.array([[1,0]] * 5) # One-hot format, defaulting to 0 class
lfoods = foods[:5]
rfoods = lfoods.copy()
rfoods.reverse()
Innotater([ImageInnotation(lfoods, name='Food 1'), ImageInnotation(rfoods, name='Food 2')],
[BinaryClassInnotation(targets, name='Are Equal')])
# -
targets
# ### Text Data - sentiment classification
# Movie reviews. In this example, numbers prefix the class names so you can keep input focus in the listbox and press 0, 1, or 2 to select the sentiment label, then press 'n' to advance to the next review (or 'p' to go back).
# +
reviews = ['I really liked this movie', 'It was OK', 'Do not watch!', 'Was worth trying it']
sentiments = [1] * len(reviews)
sentiment_classes = ['0 - Positive', '1 - Neutral', '2 - Negative']
Innotater(TextInnotation(reviews), MultiClassInnotation(sentiments, classes=sentiment_classes))
# -
list(zip(reviews, [sentiment_classes[s] for s in sentiments]))
|
Example/Examples.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://www.bigdatauniversity.com"><img src="https://ibm.box.com/shared/static/qo20b88v1hbjztubt06609ovs85q8fau.png" width="400px" align="center"></a>
#
# <h1 align="center"><font size="5">RECURRENT NETWORKS IN DEEP LEARNING</font></h1>
# Hello and welcome to this notebook. In this notebook, we will go over concepts of the Long Short-Term Memory (LSTM) model, a refinement of the original Recurrent Neural Network model. By the end of this notebook, you should be able to understand the Long Short-Term Memory model, the benefits and problems it solves, and its inner workings and calculations.
# <br>
# <h2>Table of Contents</h2>
# <ol>
# <li><a href="#intro">Introduction</a></li>
# <li><a href="#long_short_term_memory_model">Long Short-Term Memory Model</a></li>
# <li><a href="#ltsm">LTSM</a></li>
# <li><a href="#stacked_ltsm">Stacked LTSM</a></li>
# </ol>
# <p></p>
# </div>
# <br>
#
# <hr>
# <a id="intro"><a/>
# <h2>Introduction</h2>
#
# Recurrent Neural Networks are Deep Learning models with simple structures and a feedback mechanism built-in, or in different words, the output of a layer is added to the next input and fed back to the same layer.
#
# The Recurrent Neural Network is a specialized type of Neural Network that solves the issue of **maintaining context for Sequential data** -- such as Weather data, Stocks, Genes, etc. At each iterative step, the processing unit takes in an input and the current state of the network, and produces an output and a new state that is <b>re-fed into the network</b>.
#
# <img src="https://ibm.box.com/shared/static/v7p90neiaqghmpwawpiecmz9n7080m59.png">
#
#
# <center><i>Representation of a Recurrent Neural Network</i></center>
# <br><br>
# However, <b>this model has some problems</b>. It's very computationally expensive to maintain the state for a large amount of units, even more so over a long amount of time. Additionally, Recurrent Networks are very sensitive to changes in their parameters. As such, they are prone to different problems with their Gradient Descent optimizer -- they either grow exponentially (Exploding Gradient) or drop down to near zero and stabilize (Vanishing Gradient), both problems that greatly harm a model's learning capability.
#
# To solve these problems, <NAME> Schmidhuber published a paper in 1997 describing a way to keep information over long periods of time and additionally solve the oversensitivity to parameter changes, i.e., make backpropagating through the Recurrent Networks more viable. This proposed method is called Long Short-Term Memory (LSTM).
#
# (In this notebook, we will cover only LSTM and its implementation using TensorFlow)
# <hr>
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <a id="long_short_term_memory_model"></a>
# <h2>Long Short-Term Memory Model</h2>
#
# The Long Short-Term Memory, as it was called, was an abstraction of how computer memory works. It is "bundled" with whatever processing unit is implemented in the Recurrent Network, although outside of its flow, and is responsible for keeping, reading, and outputting information for the model. The way it works is simple: you have a linear unit, which is the information cell itself, surrounded by three logistic gates responsible for maintaining the data. One gate is for inputting data into the information cell, one is for outputting data from the input cell, and the last one is to keep or forget data depending on the needs of the network.
#
# Thanks to that, it not only solves the problem of keeping states, because the network can choose to forget data whenever information is not needed, it also solves the gradient problems, since the Logistic Gates have a very nice derivative.
#
# <h3>Long Short-Term Memory Architecture</h3>
#
# The Long Short-Term Memory is composed of a linear unit surrounded by three logistic gates. The name for these gates vary from place to place, but the most usual names for them are:
#
# <ul>
# <li>the "Input" or "Write" Gate, which handles the writing of data into the information cell</li>
# <li>the "Output" or "Read" Gate, which handles the sending of data back onto the Recurrent Network</li>
# <li>the "Keep" or "Forget" Gate, which handles the maintaining and modification of the data stored in the information cell</li>
# </ul>
# <br>
# <img src="https://ibm.box.com/shared/static/zx10duv5egw0baw6gh2hzsgr8ex45gsg.png" width="720"/>
# <center><i>Diagram of the Long Short-Term Memory Unit</i></center>
# <br><br>
# The three gates are the centerpiece of the LSTM unit. The gates, when activated by the network, perform their respective functions. For example, the Input Gate will write whatever data it is passed into the information cell, the Output Gate will return whatever data is in the information cell, and the Keep Gate will maintain the data in the information cell. These gates are analog and multiplicative, and as such, can modify the data based on the signal they are sent.
#
# <hr>
#
# For example, an usual flow of operations for the LSTM unit is as such: First off, the Keep Gate has to decide whether to keep or forget the data currently stored in memory. It receives both the input and the state of the Recurrent Network, and passes it through its Sigmoid activation. If $K
# _t$ has value of 1 means that the LSTM unit should keep the data stored perfectly and if $K_t$ a value of 0 means that it should forget it entirely. Consider $S_{t-1}$ as the incoming (previous) state, $x_t$ as the incoming input, and $W_k$, $B_k$ as the weight and bias for the Keep Gate. Additionally, consider $Old_{t-1}$ as the data previously in memory. What happens can be summarized by this equation:
#
# <br>
#
# <font size="4"><strong>
# $$K_t = \sigma(W_k \times [S_{t-1}, x_t] + B_k)$$
#
# $$Old_t = K_t \times Old_{t-1}$$
# </strong></font>
#
# <br>
#
# As you can see, $Old_{t-1}$ was multiplied by value was returned by the Keep Gate($K_t$) -- this value is written in the memory cell.
#
# <br>
# Then, the input and state are passed on to the Input Gate, in which there is another Sigmoid activation applied. Concurrently, the input is processed as normal by whatever processing unit is implemented in the network, and then multiplied by the Sigmoid activation's result $I_t$, much like the Keep Gate. Consider $W_i$ and $B_i$ as the weight and bias for the Input Gate, and $C_t$ the result of the processing of the inputs by the Recurrent Network.
# <br><br>
#
# <font size="4"><strong>
# $$I_t = \sigma(W_i\times[S_{t-1},x_t]+B_i)$$
#
# $$New_t = I_t \times C_t$$
# </strong></font>
#
# <br>
# $New_t$ is the new data to be input into the memory cell. This is then <b>added</b> to whatever value is still stored in memory.
# <br><br>
#
# <font size="4"><strong>
# $$Cell_t = Old_t + New_t$$
# </strong></font>
#
# <br>
# We now have the <i>candidate data</i> which is to be kept in the memory cell. The conjunction of the Keep and Input gates work in an analog manner, making it so that it is possible to keep part of the old data and add only part of the new data. Consider however, what would happen if the Forget Gate was set to 0 and the Input Gate was set to 1:
# <br><br>
#
# <font size="4"><strong>
# $$Old_t = 0 \times Old_{t-1}$$
#
# $$New_t = 1 \times C_t$$
#
# $$Cell_t = C_t$$
# </strong></font>
#
# <br>
# The old data would be totally forgotten and the new data would overwrite it completely.
#
# The Output Gate functions in a similar manner. To decide what we should output, we take the input data and state and pass it through a Sigmoid function as usual. The contents of our memory cell, however, are pushed onto a <i>Tanh</i> function to bind them between a value of -1 to 1. Consider $W_o$ and $B_o$ as the weight and bias for the Output Gate.
# <br>
# <font size="4"><strong>
# $$O_t = \sigma(W_o \times [S_{t-1},x_t] + B_o)$$
#
# $$Output_t = O_t \times tanh(Cell_t)$$
# </strong></font>
# <br>
#
# And that $Output_t$ is what is output into the Recurrent Network.
#
# <br>
# <img width="384" src="https://ibm.box.com/shared/static/rkr60528r3mz2fmtlpah8lqpg7mcsy0g.png">
# <center><i>The Logistic Function plotted</i></center>
# <br><br>
# As mentioned many times, all three gates are logistic. The reason for this is because it is very easy to backpropagate through them, and as such, it is possible for the model to learn exactly _how_ it is supposed to use this structure. This is one of the reasons for which LSTM is a very strong structure. Additionally, this solves the gradient problems by being able to manipulate values through the gates themselves -- by passing the inputs and outputs through the gates, we have now a easily derivable function modifying our inputs.
#
# In regards to the problem of storing many states over a long period of time, LSTM handles this perfectly by only keeping whatever information is necessary and forgetting it whenever it is not needed anymore. Therefore, LSTMs are a very elegant solution to both problems.
# <hr>
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <a id="ltsm"></a>
# <h2>LSTM</h2>
# Lets first create a tiny LSTM network sample to understand the architecture of LSTM networks.
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# We need to import the necessary modules for our code. We need <b><code>numpy</code></b> and <b><code>tensorflow</code></b>, obviously. Additionally, we can import directly the <b><code>tensorflow.contrib.rnn</code></b> model, which includes the function for building RNNs.
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
import numpy as np
import tensorflow as tf
sess = tf.Session()
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# We want to create a network that has only one LSTM cell. We have to pass 2 elements to LSTM, the <b>prv_output</b> and <b>prv_state</b>, so called, <b>h</b> and <b>c</b>. Therefore, we initialize a state vector, <b>state</b>. Here, <b>state</b> is a tuple with 2 elements, each one is of size [1 x 4], one for passing prv_output to next time step, and another for passing the prv_state to next time stamp.
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
LSTM_CELL_SIZE = 4 # output size (dimension), which is same as hidden size in the cell
lstm_cell = tf.contrib.rnn.BasicLSTMCell(LSTM_CELL_SIZE, state_is_tuple=True)
state = (tf.zeros([1,LSTM_CELL_SIZE]),)*2
state
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Let define a sample input. In this example, batch_size = 1, and seq_len = 6:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
sample_input = tf.constant([[3,2,2,2,2,2]],dtype=tf.float32)
print (sess.run(sample_input))
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Now, we can pass the input to lstm_cell, and check the new state:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
with tf.variable_scope("LSTM_sample1"):
output, state_new = lstm_cell(sample_input, state)
sess.run(tf.global_variables_initializer())
print (sess.run(state_new))
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# As we can see, the states has 2 parts, the new state c, and also the output h. Lets check the output again:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
print (sess.run(output))
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <hr>
# <a id="stacked_ltsm"></a>
# <h2>Stacked LSTM</h2>
# What about if we want to have a RNN with stacked LSTM? For example, a 2-layer LSTM. In this case, the output of the first layer will become the input of the second.
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Lets start with a new session:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
sess = tf.Session()
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
input_dim = 6
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Lets create the stacked LSTM cell:
# -
cells = []
# Creating the first layer LTSM cell.
LSTM_CELL_SIZE_1 = 4 #4 hidden nodes
cell1 = tf.contrib.rnn.LSTMCell(LSTM_CELL_SIZE_1)
cells.append(cell1)
# Creating the second layer LTSM cell.
LSTM_CELL_SIZE_2 = 5 #5 hidden nodes
cell2 = tf.contrib.rnn.LSTMCell(LSTM_CELL_SIZE_2)
cells.append(cell2)
# To create a multi-layer LTSM we use the <b>tf.contrib.rnnMultiRNNCell</b> function, it takes in multiple single layer LTSM cells to create a multilayer stacked LTSM model.
stacked_lstm = tf.contrib.rnn.MultiRNNCell(cells)
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Now we can create the RNN from <b>stacked_lstm</b>:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
# Batch size x time steps x features.
data = tf.placeholder(tf.float32, [None, None, input_dim])
output, state = tf.nn.dynamic_rnn(stacked_lstm, data, dtype=tf.float32)
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Lets say the input sequence length is 3, and the dimensionality of the inputs is 6. The input should be a Tensor of shape: [batch_size, max_time, dimension], in our case it would be (2, 3, 6)
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
#Batch size x time steps x features.
sample_input = [[[1,2,3,4,3,2], [1,2,1,1,1,2],[1,2,2,2,2,2]],[[1,2,3,4,3,2],[3,2,2,1,1,2],[0,0,0,0,3,2]]]
sample_input
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# we can now send our input to network, and check the output:
# -
output
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
sess.run(tf.global_variables_initializer())
sess.run(output, feed_dict={data: sample_input})
# -
# As you see, the output is of shape (2, 3, 5), which corresponds to our 2 batches, 3 elements in our sequence, and the dimensionality of the output which is 5.
# <hr>
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ## Want to learn more?
#
# Running deep learning programs usually needs a high performance platform. __PowerAI__ speeds up deep learning and AI. Built on IBM’s Power Systems, __PowerAI__ is a scalable software platform that accelerates deep learning and AI with blazing performance for individual users or enterprises. The __PowerAI__ platform supports popular machine learning libraries and dependencies including TensorFlow, Caffe, Torch, and Theano. You can use [PowerAI on IMB Cloud](https://cocl.us/ML0120EN_PAI).
#
# Also, you can use __Watson Studio__ to run these notebooks faster with bigger datasets.__Watson Studio__ is IBM’s leading cloud solution for data scientists, built by data scientists. With Jupyter notebooks, RStudio, Apache Spark and popular libraries pre-packaged in the cloud, __Watson Studio__ enables data scientists to collaborate on their projects without having to install anything. Join the fast-growing community of __Watson Studio__ users today with a free account at [Watson Studio](https://cocl.us/ML0120EN_DSX).This is the end of this lesson. Thank you for reading this notebook, and good luck on your studies.
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ### Thanks for completing this lesson!
#
# Notebook created by: <a href = "https://linkedin.com/in/saeedaghabozorgi"> <NAME> </a>, <a href="https://br.linkedin.com/in/walter-gomes-de-amorim-junior-624726121"><NAME></a>
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <hr>
#
# Copyright © 2018 [Cognitive Class](https://cocl.us/DX0108EN_CC). This notebook and its source code are released under the terms of the [MIT License](https://bigdatauniversity.com/mit-license/).
|
content/docs/data-science-with-python/labs/deep-learning-with-tensorflow/3-1-LSTM-basics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/DheniMoura/LPA_UNINTER/blob/main/LPA_Aula_02.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="7EjWRErW4LX5"
# Aula 02 da matéria Lógica de Programação e Algoritmos
# + [markdown] id="DifU1iku6SD9"
# Ex. 02 - Desenvolva um algoritmo que solicite ao usuário uma quantidade de dias, de horas, de minutos e de segundos. Calcule o total de segundos resultante e imprima na tela para o usuário.
# + colab={"base_uri": "https://localhost:8080/"} id="C9AbSblg4UyZ" outputId="bed0d8ec-72bd-4af1-9e6a-a8070f1cac27"
d = int(input('Quantos dias? '))
h = int(input('Quantas horas? '))
m = int(input('Quantos minutos? '))
s = int(input('Quantos segundos? '))
total = s + (m * 60) + (h * 3600) + (d * 86400)
print('O total de segundos calculados é: {}'.format(total))
# + [markdown] id="3oG9HJg26lWj"
# Ex. 03 - Desenvolva um algoritmo que solicite ao usuário o preço de um produto e um percentual de desconto a ser aplicado a ele. Calcule e exiba o valor do desconto e o preço final do produto.
# + colab={"base_uri": "https://localhost:8080/"} id="pl5AqAM_6Q_p" outputId="94f79fbf-9be0-4619-f0a0-1fd3c69c814a"
p = float(input('Digite o preço original do produto: '))
d = float(input('Digite o percentual de desconto: (0% a 100%: '))
vd = p * (d / 100)
pf = p - vd
print('Valor do desconto: R$%.2f' % vd)
print('Preço final: R$%.2f' % pf)
# + [markdown] id="LteBKbR3TYqv"
# Crie uma variável de String que receba uma frase qualquer.
# Crie uma segunda variável, agora contendo a metade da string digitada.
# Imprima na tela somente os dois últimos caracteres da segunda variável do tipo string
# + colab={"base_uri": "https://localhost:8080/"} id="qLEXIkKNTcOf" outputId="41cf4724-7475-4995-9115-da2f12d790fc"
v1 = str(input('Digite uma frase: '))
tam = len(v1)
v2 = v1[:int(tam/2)]
print(v2[-2:])
|
LPA_Aula_02.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
import pickle
import importlib
from library import data_preprocess as dp
importlib.reload(dp)
import random
from time import time
import numpy as np
import keras
from keras.preprocessing.text import Tokenizer
from keras.utils import to_categorical
from keras.preprocessing.sequence import pad_sequences
from keras.models import Sequential, Input
from keras.layers import Dense, Dropout, Activation
from keras.layers import LSTM, Bidirectional
from keras.layers import Embedding, TimeDistributed, Flatten, Merge, Concatenate
from keras import regularizers
from keras.metrics import sparse_categorical_accuracy, sparse_categorical_crossentropy
from keras.models import load_model
from keras.optimizers import Adam, RMSprop
from keras.models import Model
from keras.callbacks import TensorBoard, EarlyStopping, ModelCheckpoint
import tensorflow as tf
from keras import backend as K
from keras.utils import multi_gpu_model
# -
K.tensorflow_backend._get_available_gpus()
# ### Variables
# +
DATA_PATH = './datasets/jokes.pickle'
VOCAB_PATH = './datasets/jokes_vocabulary.pickle'
MODELS_PATH = './models/'
MAX_SEQUENCE_LENGTH = 13
VALIDATION_SPLIT = 0.2
MODEL_PREFIX = 'jokes_stacked_lstm'
EMBEDDING_DIM = 512
HIDDEN_DIM1 = 1024
HIDDEN_DIM2 = 512
DEEPER_DIM = 512
DROPOUT_FACTOR = 0.2
REGULARIZATION = 0.00001
LEARNING_RATE = 0.003
DATA_PERCENT = 0.1
RUN_INDEX = 6
# +
with open(DATA_PATH, 'rb') as pickleFile:
sentences = pickle.load(pickleFile)
with open(VOCAB_PATH, 'rb') as pickleFile:
vocab = pickle.load(pickleFile)
random.shuffle(sentences)
print("Number of sentences = ", len(sentences))
print(sentences[:2])
print("Vocab size = ", len(vocab))
print(vocab[:10])
# +
# tokenize data
num_words = len(vocab)
tokenizer = Tokenizer(num_words=None, filters='', lower=True, split=' ',
char_level=False, oov_token=None)
tokenizer.fit_on_texts(sentences)
assert num_words == len(tokenizer.word_index)
encoded_sentences = tokenizer.texts_to_sequences(sentences)
print(encoded_sentences[:5])
VOCAB_SIZE = len(tokenizer.word_index) + 1
print(VOCAB_SIZE)
# -
# saving
with open(MODELS_PATH + MODEL_PREFIX + '_tokenizer_' + str(RUN_INDEX) + '.pickle', 'wb') as handle:
pickle.dump(tokenizer, handle, protocol=pickle.HIGHEST_PROTOCOL)
# ### Preparing Training Data
# +
X_data = []
y_data = []
for sentence in encoded_sentences:
l = len(sentence)
sliding_window_length = min(l-3, MAX_SEQUENCE_LENGTH)
step_size = 1
for i in range(0, l - sliding_window_length, step_size):
X_data.append(sentence[i:i+sliding_window_length])
y_data.append(sentence[i+1:i+sliding_window_length+1])
print("Total training data size = ", len(X_data))
MAX_SEQ_LEN = max([len(seq) for seq in X_data])
print("Max seq len = ", MAX_SEQ_LEN)
X_data = pad_sequences(X_data, maxlen=MAX_SEQ_LEN, padding='pre')
y_data = pad_sequences(y_data, maxlen=MAX_SEQ_LEN, padding='pre').reshape(-1, MAX_SEQ_LEN, 1)
#y_data = np.array(y_data).reshape(-1,1)
print(X_data.shape)
print(X_data[:2])
print(y_data.shape)
print(y_data[:2])
# -
# define model
def StackedLSTM(vocab_size, embedding_dim, hidden_dim1, hidden_dim2, deeper_dim, max_seq_len,
dropout_factor=0.5, regularization=0.00001, learning_rate=0.001):
model = Sequential()
model.add(Embedding(vocab_size, embedding_dim, #input_length=max_seq_len,
mask_zero=True, embeddings_regularizer=regularizers.l2(regularization)))
model.add(LSTM(hidden_dim1, activation='tanh',
kernel_regularizer=regularizers.l2(regularization),
recurrent_regularizer=regularizers.l2(regularization), #unroll=True,
return_sequences = True, dropout=dropout_factor, recurrent_dropout=dropout_factor))
model.add(LSTM(hidden_dim2, activation='tanh',
kernel_regularizer=regularizers.l2(regularization),
recurrent_regularizer=regularizers.l2(regularization), #unroll=True,
return_sequences = True, dropout=dropout_factor, recurrent_dropout=dropout_factor))
model.add(TimeDistributed(Dropout(dropout_factor)))
model.add(Dense(units=deeper_dim, activation='tanh', kernel_regularizer=regularizers.l2(regularization)))
model.add(Dropout(dropout_factor))
model.add(Dense(units=vocab_size, activation='softmax',
kernel_regularizer=regularizers.l2(regularization)))
#model = multi_gpu_model(model)
model.compile(loss='sparse_categorical_crossentropy', optimizer=RMSprop(lr=learning_rate),
metrics=[sparse_categorical_crossentropy, sparse_categorical_accuracy], sample_weight_mode='temporal')
return model
K.clear_session()
sess = tf.Session()
K.set_session(sess)
model = StackedLSTM(vocab_size=VOCAB_SIZE, embedding_dim=EMBEDDING_DIM, hidden_dim1=HIDDEN_DIM1, hidden_dim2=HIDDEN_DIM2,
deeper_dim=DEEPER_DIM, max_seq_len=MAX_SEQ_LEN, dropout_factor=DROPOUT_FACTOR,
regularization=REGULARIZATION, learning_rate=LEARNING_RATE)
print(model.summary())
class TB(TensorBoard):
def __init__(self, log_every=1, **kwargs):
super().__init__(**kwargs)
self.log_every = log_every
self.counter = 0
def on_batch_end(self, batch, logs=None):
self.counter+=1
if self.counter%self.log_every==0:
for name, value in logs.items():
if name in ['batch', 'size']:
continue
summary = tf.Summary()
summary_value = summary.value.add()
summary_value.simple_value = value.item()
summary_value.tag = name
self.writer.add_summary(summary, self.counter)
self.writer.flush()
super().on_batch_end(batch, logs)
# +
start_time = time()
tensorboard = TB(log_dir="./logs/" + MODEL_PREFIX + "/{}".format(time()),
histogram_freq=0, write_graph=True, write_images=False, log_every=10)
callbacks=[tensorboard,
EarlyStopping(patience=5, monitor='val_loss'),
ModelCheckpoint(filepath=MODELS_PATH + 'checkpoints/'+ MODEL_PREFIX + '_gen'+str(RUN_INDEX)+'.{epoch:02d}-{val_loss:.2f}.hdf5',
monitor='val_loss', verbose=1, mode='auto', period=1),
ModelCheckpoint(filepath=MODELS_PATH + MODEL_PREFIX + '_gen'+str(RUN_INDEX)+'.hdf5',
monitor='val_loss', verbose=1, mode='auto', period=1, save_best_only=True)]
model.fit(X_data, y_data, epochs=25, batch_size=1024, shuffle=True, verbose=1, validation_split=0.2, callbacks=callbacks)
print("Total elapsed time: ", time()-start_time)
# -
# generate a sequence from a language model
def generate(model, tokenizer, seed_text, maxlen, probabilistic=False, exploration_factor=0.0):
reverse_word_map = dict(map(reversed, tokenizer.word_index.items()))
seq = tokenizer.texts_to_sequences([seed_text])[0]
print(seq)
while True:
encoded_seq = seq
if len(seq) > MAX_SEQ_LEN:
encoded_seq = encoded_seq[-1*MAX_SEQ_LEN:]
#padded_seq = pad_sequences([encoded_seq], maxlen=MAX_SEQ_LEN, padding='pre')
padded_seq = np.array([seq])
y_prob = model.predict(padded_seq)[0][-1].reshape(1,-1)#[3:].reshape(-1,1)
if random.random() <= exploration_factor:
probabilistic = True
else:
probabilistic = False
if probabilistic:
y_class = np.argmax(np.random.multinomial(1,y_prob[0]/(np.sum(y_prob[0])+1e-5),1))
else:
y_class = y_prob.argmax(axis=-1)[0]
if y_class == 0:
break
out_word = reverse_word_map[y_class]
seq.append(y_class)
if out_word == 'eos' or len(seq) > maxlen or out_word == 'sos':
break
words = [reverse_word_map[idx] for idx in seq]
return ' '.join(words)
print(sentences[:10])
model = load_model('models/jokes_stacked_lstm_gen6.hdf5')
with open('models/jokes_stacked_lstm_tokenizer_6.pickle', 'rb') as pickleFile:
tokenizer = pickle.load(pickleFile)
joke = generate(model, tokenizer, "sos i like ", maxlen=40, exploration_factor=0.2)
print(joke)
# +
def bigrams_list(sentence):
words = sentence.split(' ')
bigrams = []
for i in range(0, len(words)-1):
bigrams.append(words[i]+' '+words[i+1])
return bigrams
print(bigrams_list("sos hello , i'm a dinosaur . eos"))
# -
sentence_bigrams = [bigrams_list(s) for s in sentences]
print(sentence_bigrams[:2])
# +
def intersection(lst1, lst2):
temp = set(lst2)
lst3 = [value for value in lst1 if value in temp]
return lst3
def similarity_score(lst1, lst2):
intersection_len = len(intersection(lst1, lst2))
return (1.0*intersection_len)/len(lst1)#+len(lst2)-intersection_len)
def print_closest_sentences(sentence, sentence_bigrams, top_k=3):
bigrams = bigrams_list(sentence)
scores = np.array([similarity_score(bigrams, sbigrams)
for sbigrams in sentence_bigrams])
top_k_indices = scores.argsort()[-1*top_k:][::-1]
top_k_scores = scores[top_k_indices]
for k in range(top_k):
print(top_k_scores[k], " -> ", sentences[top_k_indices[k]])
# -
print_closest_sentences(joke, sentence_bigrams, 10)
joke = generate(model, tokenizer, "sos what do you call", maxlen=40, exploration_factor=0.3)
print(joke)
print_closest_sentences(joke, sentence_bigrams, 10)
print(sentences[:10])
joke = generate(model, tokenizer, "sos i ", maxlen=40, exploration_factor=0.0)
print(joke)
print_closest_sentences(joke, sentence_bigrams, 10)
model1 = load_model('models/checkpoints/jokes_bilstm_gen2.08-4.39.hdf5')
joke = generate(model, tokenizer, "sos a guy finds", maxlen=40)
print(joke)
print_closest_sentences(joke, sentence_bigrams, 10)
|
Jokes-Word-Stacked-LSTM-Keras.ipynb
|
# <a href="https://mybinder.org/v2/gh/HuygensING/hyper-collate/master?filepath=hyper-collate-jupyter%2Fhyper-collate-readme.ipynb" target="_blank"><img src="https://mybinder.org/badge_logo.svg"/></a>
# # Using HyperCollate in Kotlin Notebooks:
#
# ## Dependencies
#
# To be able to use HyperCollate, use the following `@file` commands and imports to install the necessary dependencies, initialize HyperCollate and define some functions to display the HyperCollate graphs in the notebook:
# +
@file:Repository("http://maven.huygens.knaw.nl/repository/")
@file:DependsOn("nl.knaw.huygens:hyper-collate-core:1.3.4")
@file:DependsOn("nl.knaw.huygens:hyper-collate-jupyter:1.3.4")
import nl.knaw.huygens.hypercollate.jupyter.*
import nl.knaw.huygens.hypercollate.model.*
HC.init()
fun VariantWitnessGraph.show(colored: Boolean = true, join: Boolean = true, emphasizeWhitespace: Boolean = false) = MIME(this.asSVGPair(colored, join, emphasizeWhitespace))
fun CollationGraph.asHTML() = HTML(this.asHTMLString())
fun CollationGraph.show(join: Boolean = true, emphasizeWhitespace: Boolean = false) = MIME(this.asSVGPair(join, emphasizeWhitespace))
# -
# After this, you're ready to use HyperCollate:
#
# ## Import witnesses
#
# There are 2 ways to import the XML of the witnesses you want to collate:
#
# - Inline string:
#
val wA = HC.importXMLWitness("A", "<text>The dog's big eyes.</text>")
val wB = HC.importXMLWitness("B", "<text>The dog's <del>big black ears</del><add>brown eyes</add>.</text>")
# - From a File:
import java.io.File
val wC = HC.importXMLWitness("C",File("c.xml"))
# The witnesses can be visualized as a graph using `.show()`
wA.show()
# The `.show()` for witness graphs has several options:
#
# - colored : will use different colors for the different markup nodes (default: true)
#
# turning it off will produce a simpler graph:
wA.show(colored=false)
# - join: will minimize the amount of nodes in the graph by joining tokens where possible. (default: true)
#
# Turn it off to see the individual tokens:
wA.show(join=false)
# - emphasizeWhitespace: visualize whitespace in the tokens (default: false)
wA.show(emphasizeWhitespace=true)
# ## Collating the witnesses
val collationGraph = HC.collate(wA,wB,wC)
# The collationGraph can be visualized as an ASCII table:
collationGraph.asASCIITable()
# As an HTML table:
collationGraph.asHTML()
# Or as a set of nodes and edges:
collationGraph.show()
# This `.show()` also has the `join` and `emphasizeWhitespace` options mentioned above.
|
notebooks/hyper-collate-readme.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Module 2 (Python 3)
# ## Basic NLP Tasks with NLTK
import nltk
nltk.download('treebank')
from nltk.book import *
# ### Counting vocabulary of words
text7
sent7
len(sent7)
len(text7)
len(set(text7))
list(set(text7))[:10]
# ### Frequency of words
dist = FreqDist(text7)
len(dist)
vocab1 = dist.keys()
#vocab1[:10]
# In Python 3 dict.keys() returns an iterable view instead of a list
list(vocab1)[:10]
dist['four']
freqwords = [w for w in vocab1 if len(w) > 5 and dist[w] > 100]
freqwords
# ### Normalization and stemming
input1 = "List listed lists listing listings"
words1 = input1.lower().split(' ')
words1
porter = nltk.PorterStemmer()
[porter.stem(t) for t in words1]
??nltk.corpus.udhr.words()
# ### Lemmatization
udhr = nltk.corpus.udhr.words('English-Latin1')
udhr[:20]
[porter.stem(t) for t in udhr[:20]] # Still Lemmatization
WNlemma = nltk.WordNetLemmatizer()
[WNlemma.lemmatize(t) for t in udhr[:20]]
# ### Tokenization
text11 = "Children shouldn't drink a sugary drink before bed."
text11.split(' ')
nltk.word_tokenize(text11)
text12 = "This is the first sentence. A gallon of milk in the U.S. costs $2.99. Is this the third sentence? Yes, it is!"
sentences = nltk.sent_tokenize(text12)
len(sentences)
sentences
# ## Advanced NLP Tasks with NLTK
# ### POS tagging
nltk.help.upenn_tagset('MD')
text13 = nltk.word_tokenize(text11)
nltk.pos_tag(text13)
text14 = nltk.word_tokenize("Visiting aunts can be a nuisance")
nltk.pos_tag(text14)
# +
# Parsing sentence structure
text15 = nltk.word_tokenize("Alice loves Bob")
grammar = nltk.CFG.fromstring("""
S -> NP VP
VP -> V NP
NP -> 'Alice' | 'Bob'
V -> 'loves'
""")
parser = nltk.ChartParser(grammar)
trees = parser.parse_all(text15)
for tree in trees:
print(tree)
# -
text16 = nltk.word_tokenize("I saw the man with a telescope")
grammar1 = nltk.data.load('mygrammar.cfg')
grammar1
parser = nltk.ChartParser(grammar1)
trees = parser.parse_all(text16)
for tree in trees:
print(tree)
from nltk.corpus import treebank
text17 = treebank.parsed_sents('wsj_0001.mrg')[0]
print(text17)
# ### POS tagging and parsing ambiguity
text18 = nltk.word_tokenize("The old man the boat")
nltk.pos_tag(text18)
text19 = nltk.word_tokenize("Colorless green ideas sleep furiously")
nltk.pos_tag(text19)
|
w2_Module+2+(Python+3).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.10 64-bit (''qcsubmit'': conda)'
# name: python_defaultSpec_1595879535660
# ---
# +
from qcsubmit.factories import TorsiondriveDatasetFactory
from qcsubmit.datasets import TorsiondriveDataset
from qcsubmit import workflow_components
from qcsubmit.common_structures import TorsionIndexer
from openforcefield.topology import Molecule as OFFMolecule
# from qcelemental.models import Molecule as QCEMolecule
import os, json, tqdm
# -
factory = TorsiondriveDatasetFactory()
factory.scf_properties = ['dipole', 'quadrupole', 'wiberg_lowdin_indices', 'mayer_indices']
factory
# now write the settings out
factory.export_settings("theory-bm-set_setttings.yaml")
# +
# now create the dataset from the pdbs in the pdb folder
dataset = factory.create_dataset(dataset_name="OpenFF Theory Benchmarking Set B3LYP-D3BJ DZVP v1.0", molecules=[], description="A torsiondrive dataset for benchmarking B3LYP-D3BJ/DZVP", tagline="Torsiondrives for benchmarking B3LYP-D3BJ/DZVP")
# -
with open('input_torsions.json') as infile:
selected_torsions = json.load(infile)
# + tags=[]
for idx, (canonical_torsion_index, torsion_data) in enumerate(tqdm.tqdm(selected_torsions.items())):
attributes = torsion_data["attributes"]
torsion_atom_indices = torsion_data["atom_indices"]
grid_spacings = [15] * len(torsion_atom_indices)
initial_molecules = torsion_data["initial_molecules"]
# molecule = OFFMolecule.from_qcschema(torsion_data, client=client) # not working for some reason. need to dig into
molecule = OFFMolecule.from_qcschema(torsion_data)
molecule.generate_conformers(n_conformers = 5)
print(f'{idx}: {molecule.n_conformers}')
dataset.add_molecule(index=idx, molecule= molecule, attributes=attributes, dihedrals=torsion_atom_indices)
# -
dataset.spec_name
dataset.n_molecules
dataset.n_records
dataset.metadata.long_description_url = "https://github.com/openforcefield/qca-dataset-submission/tree/master/submissions/2020-07-27-theory-bm-set-b3lyp-d3bj-dzvp"
# +
# export the dataset
dataset.export_dataset("dataset.json")
# -
dataset.molecules_to_file("theory-bm-set-curated.smi", "smi")
# export the molecules to pdf with torsions highlighted
dataset.visualize("theory-bm-set-curated.pdf", 'openeye')
|
submissions/2020-07-27-theory-bm-set-b3lyp-d3bj-dzvp/QCSubmit_workflow.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import folium
print(folium.__version__)
# +
import gpxpy
fname = os.path.join('data', '2014_08_05_farol.gpx')
gpx = gpxpy.parse(open(fname))
print('{} track(s)'.format(len(gpx.tracks)))
track = gpx.tracks[0]
print('{} segment(s)'.format(len(track.segments)))
segment = track.segments[0]
print('{} point(s)'.format(len(segment.points)))
# -
data = []
segment_length = segment.length_3d()
for point_idx, point in enumerate(segment.points):
data.append(
[
point.longitude,
point.latitude,
point.elevation,
point.time,
segment.get_speed(point_idx)
]
)
# +
from pandas import DataFrame
columns = ['Longitude', 'Latitude', 'Altitude', 'Time', 'Speed']
df = DataFrame(data, columns=columns)
df.head()
# +
import numpy as np
from geographiclib.geodesic import Geodesic
angles = [90]
for i in range(len(df) - 1):
info = Geodesic.WGS84.Inverse(
df.iloc[i, 1], df.iloc[i, 0],
df.iloc[i + 1, 1], df.iloc[i + 1, 0]
)
angles.append(info['azi2'])
# Change from CW-from-North to CCW-from-East.
angles = np.deg2rad(450 - np.array(angles))
# Normalize the speed to use as the length of the arrows.
r = df['Speed'] / df['Speed'].max()
df['u'] = r * np.cos(angles)
df['v'] = r * np.sin(angles)
# +
import mplleaflet
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
df = df.dropna()
# This style was lost below.
ax.plot(
df['Longitude'],
df['Latitude'],
color='darkorange',
linewidth=5,
alpha=0.5
)
# This is preserved in the SVG icon.
sub = 10
kw = {'color': 'deepskyblue', 'alpha': 0.8, 'scale': 10}
ax.quiver(df['Longitude'][::sub],
df['Latitude'][::sub],
df['u'][::sub],
df['v'][::sub], **kw)
gj = mplleaflet.fig_to_geojson(fig=fig)
# +
import folium
lon, lat = -38.51386097, -13.00868051
zoom_start = 14
m = folium.Map(
location=[lat, lon],
tiles='Cartodb Positron',
zoom_start=zoom_start
)
# The first geometry is a lineString.
line_string = gj['features'][0]
gjson = folium.features.GeoJson(line_string)
m.add_child(gjson)
m.save(os.path.join('results', 'Folium_and_mplleaflet_0.html'))
m
# -
line_string['properties']
# +
from IPython.display import HTML
msg = '<font color="{}">This should be darkorange!</font>'.format
HTML(msg(line_string['properties']['color']))
# +
m = folium.Map(
location=[lat, lon],
tiles='Cartodb Positron',
zoom_start=zoom_start
)
icon_size = (14, 14)
for feature in gj['features']:
if feature['geometry']['type'] == 'LineString':
continue
elif feature['geometry']['type'] == 'Point':
lon, lat = feature['geometry']['coordinates']
html = feature['properties']['html']
icon_anchor = (feature['properties']['anchor_x'],
feature['properties']['anchor_y'])
icon = folium.features.DivIcon(html=html,
icon_size=(14, 14),
icon_anchor=icon_anchor)
marker = folium.map.Marker([lat, lon], icon=icon)
m.add_child(marker)
else:
msg = 'Unexpected geometry {}'.format
raise ValueError(msg(feature['geometry']))
m.save(os.path.join('results', 'Folium_and_mplleaflet_1.html'))
m
|
examples/Folium_and_mplleaflet.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="BHHvad7hr2q8"
# ### Mounting Googel Drive
# + colab={"base_uri": "https://localhost:8080/"} id="a7SZvChmr8BG" outputId="6e083ac8-1c3b-44da-fc9b-de7c2dfefbfa"
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="-e-kn1-vr7aC"
# ### Importing Dependencies
# + colab={"base_uri": "https://localhost:8080/"} id="yQU5vMXImRuU" outputId="36f7fdb1-39c2-42fd-888a-5d9c4755e93d"
import os
# !pip install dgl-cu111 -f https://data.dgl.ai/wheels/repo.html
import numpy as np
import dgl
import torch
from torch.utils.data import Dataset
from torch.utils.data import DataLoader
import torch.nn as nn
import torch.nn.functional as F
import urllib.request
import pandas as pd
import dgl.data
import dgl
from dgl.data import DGLDataset
import torch
import os
import itertools
import dgl.nn as dglnn
from dgl.nn import GraphConv
from scipy.spatial import Delaunay
from sklearn.metrics import f1_score
import sklearn
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
# + [markdown] id="HJJ92X2K3Ypd"
# ### Reding CSV files defining the classes, edges and node feaures respectively. More details can be found at: https://www.kaggle.com/ellipticco/elliptic-data-set
#
# NOTE: Please change the path of the CSV files according to your directory structure.
# + id="pv1ifoIEmiEC"
classes = pd.read_csv('/content/gdrive/MyDrive/Fall_21/BC_DL/elliptic_bitcoin_dataset/elliptic_txs_classes.csv')
edges = pd.read_csv('/content/gdrive/MyDrive/Fall_21/BC_DL/elliptic_bitcoin_dataset/elliptic_txs_edgelist.csv')
features = pd.read_csv('/content/gdrive/MyDrive/Fall_21/BC_DL/elliptic_bitcoin_dataset/elliptic_txs_features.csv',header=None).set_index(0,verify_integrity=True)
# + [markdown] id="5V7ja3eB3qnS"
# ### Filtering entries with unknown classes.
# + id="voRRd6iYOy4B"
classes_filtered = classes
classes_filtered = classes_filtered[classes_filtered['class'] != 'unknown']
# + [markdown] id="h6if1MDj3yKG"
# ### Spliting features into 2 sections: i) all entries with 1st feature value below 35 would be used for training ii) all entries with 2nd feature value above 35 would be used for testing.
# + id="vPba-bqDq-Vl"
features_train = features[features[1]<35]
features_test = features[features[1]>=35]
# + [markdown] id="Q7towY9uImwM"
# ### Creating Training & testing dataset
# + id="T5HtcP6i3t9R"
train_x = []
train_y = []
for index, row in features_train.iterrows():
if (len(classes_filtered.loc[classes_filtered['txId']==index]['class'].values) != 0):
train_x.append(row.to_numpy())
if int(classes_filtered.loc[classes_filtered['txId']==index]['class'].values) == 1:
val = 1
elif int(classes_filtered.loc[classes_filtered['txId']==index]['class'].values) == 2:
val = 0
train_y.append(val)
# + id="xpi3ifgy7FZD"
test_x = []
test_y = []
for index, row in features_test.iterrows():
if (len(classes_filtered.loc[classes_filtered['txId']==index]['class'].values) != 0):
test_x.append(row.to_numpy())
if int(classes_filtered.loc[classes_filtered['txId']==index]['class'].values) == 1:
val = 1
elif int(classes_filtered.loc[classes_filtered['txId']==index]['class'].values) == 2:
val = 0
test_y.append(val)
# + [markdown] id="0Il2g3RiIrc-"
# ### Fitting a Random Forest Classifier
# + colab={"base_uri": "https://localhost:8080/"} id="XmJBim1e5XbV" outputId="75e450b4-06c7-440d-a4da-89e2f25e5cf8"
clf = RandomForestClassifier(n_estimators=50, max_features=50)
clf.fit(train_x, train_y)
pred_rf = clf.predict(test_x)
f1 = f1_score(pred_rf, test_y, average=None)
f1m = f1_score(pred_rf, test_y, average='micro')
print("Final F1 score:",(f1[0]+f1[1])/2)
print("Final MicroAvg F1 score:",f1[0])
# + [markdown] id="w-sl9rQ4Ix68"
# ### Fitting a Logistic Regression Classifier
# + colab={"base_uri": "https://localhost:8080/"} id="TAQlHMqiGr1g" outputId="6bcdee6f-95e1-4667-cb72-df0b16b3f680"
clf = LogisticRegression().fit(train_x, train_y)
pred_lr = clf.predict(test_x)
f1 = f1_score(pred_lr, test_y, average=None)
f1m = f1_score(pred_rf, test_y, average='micro')
print("Final F1 score:",(f1[0]+f1[1])/2)
print("Final MicroAvg F1 score:",f1[0])
# + [markdown] id="Y5lDV0VxI5-d"
# ### Creating a pytorch Dataset and Dataloader for the given bitcoin dataset
# + id="T9ow7Vs-AQhl"
class DSet(Dataset):
def __init__(self, feat, label):
self.labels = label
self.feat = feat
def __len__(self):
return len(self.labels)
def __getitem__(self, idx):
x = self.feat[idx]
y = self.labels[idx]
return x, y
# + [markdown] id="D7PFfTCsJH7G"
# ### Creating Pytorch dataset and dataloaders
# + id="TsSt78EqBX8V"
train_ds = DSet(train_x,train_y)
test_ds = DSet(test_x,test_y)
train_dataloader = DataLoader(train_ds, batch_size=1000, shuffle=True)
test_dataloader = DataLoader(test_ds, batch_size=1000, shuffle=True)
# + [markdown] id="6GqTYL1yKISI"
# ### Defing a evaluation function for MLP
# + id="CWtnE2gEDexj"
def test(dataloader, model, loss_fn):
size = len(dataloader.dataset)
num_batches = len(dataloader)
model.eval()
test_loss, correct = 0, 0
f1_micro = 0
f1_net = 0
cnt = 0
with torch.no_grad():
for X, y in dataloader:
y = torch.unsqueeze(y,1)
X, y = X.float(), y.float()
pred = model(X)
test_loss += loss_fn(pred, y).item()
pred = pred.argmax(1)
y = torch.squeeze(y,1)
f1_m = f1_score(pred, y, average='micro')
f1 = f1_score(pred, y, average=None)
f1_micro += f1[0]
f1_net += (f1[0]+f1[1])/2
cnt += 1
print("Final F1 score:",f1_net/cnt)
print("Final MicroAvg F1 score:",f1_micro/cnt)
# + [markdown] id="ZKqQKfXdJNNA"
# ### Define a simple MLP
# + id="eNLyTc_O7b82"
class MLP(nn.Module):
def __init__(self, n_inputs):
super(MLP, self).__init__()
self.layer1 = nn.Linear(n_inputs, 50)
self.layer2 = nn.Linear(50, 1)
self.activation = nn.Sigmoid()
def forward(self, X):
X = self.layer1(X)
X = self.layer2(X)
X = self.activation(X)
return X
# + [markdown] id="bzw6xzYUNoP8"
# ### Train and Evaluate MLP
# + colab={"base_uri": "https://localhost:8080/"} id="ov1DZvS0Dp-i" outputId="2deff496-1069-445b-8ff5-e1a38f397093"
model = MLP(train_x[0].shape[0])
optimizer = torch.optim.Adam(model.parameters(), lr=0.001)
criterion = nn.BCELoss()
for epoch in range(200):
for i, (x, y) in enumerate(train_dataloader):
y = torch.unsqueeze(y,1)
x = x.float()
y = y.float()
optimizer.zero_grad()
yhat = model(x)
loss = criterion(yhat, y)
print("LOSS:", loss)
loss.backward()
optimizer.step()
test(test_dataloader, model, criterion)
|
Karan_Bali_File2_Other_methods.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="pO4-CY_TCZZS" colab_type="text"
# # Train a Simple Audio Recognition model for microcontroller use
# + [markdown] id="BaFfr7DHRmGF" colab_type="text"
# This notebook demonstrates how to train a 20kb [Simple Audio Recognition](https://www.tensorflow.org/tutorials/sequences/audio_recognition) model for [TensorFlow Lite for Microcontrollers](https://tensorflow.org/lite/microcontrollers/overview). It will produce the same model used in the [micro_speech](https://github.com/tensorflow/tensorflow/tree/master/tensorflow/lite/experimental/micro/examples/micro_speech) example application.
#
# The model is designed to be used with [Google Colaboratory](https://colab.research.google.com).
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/tensorflow/blob/master/tensorflow/lite/experimental/micro/examples/micro_speech/train_speech_model.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/tensorflow/blob/master/tensorflow/lite/experimental/micro/examples/micro_speech/train_speech_model.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
#
#
#
# The notebook runs Python scripts to train and freeze the model, and uses the TensorFlow Lite converter to convert it for use with TensorFlow Lite for Microcontrollers.
#
# **Training is much faster using GPU acceleration.** Before you proceed, ensure you are using a GPU runtime by going to **Runtime -> Change runtime type** and selecting **GPU**. Training 18,000 iterations will take 1.5-2 hours on a GPU runtime.
#
# ## Configure training
#
# The following `os.environ` lines can be customized to set the words that will be trained for, and the steps and learning rate of the training. The default values will result in the same model that is used in the micro_speech example. Run the cell to set the configuration:
# + id="ludfxbNIaegy" colab_type="code" colab={}
import os
# A comma-delimited list of the words you want to train for.
# The options are: yes,no,up,down,left,right,on,off,stop,go
# All other words will be used to train an "unknown" category.
os.environ["WANTED_WORDS"] = "yes,no"
# The number of steps and learning rates can be specified as comma-separated
# lists to define the rate at each stage. For example,
# TRAINING_STEPS=15000,3000 and LEARNING_RATE=0.001,0.0001
# will run 18,000 training loops in total, with a rate of 0.001 for the first
# 15,000, and 0.0001 for the final 3,000.
os.environ["TRAINING_STEPS"]="15000,3000"
os.environ["LEARNING_RATE"]="0.001,0.0001"
# Calculate the total number of steps, which is used to identify the checkpoint
# file name.
total_steps = sum(map(lambda string: int(string),
os.environ["TRAINING_STEPS"].split(",")))
os.environ["TOTAL_STEPS"] = str(total_steps)
# Print the configuration to confirm it
# !echo "Training these words: ${WANTED_WORDS}"
# !echo "Training steps in each stage: ${TRAINING_STEPS}"
# !echo "Learning rate in each stage: ${LEARNING_RATE}"
# !echo "Total number of training steps: ${TOTAL_STEPS}"
# + [markdown] id="gCgeOpvY9pAi" colab_type="text"
# ## Install dependencies
#
# Next, we'll install a GPU build of TensorFlow, so we can use GPU acceleration for training. We also clone the TensorFlow repository, which contains the scripts that train and freeze the model.
# + id="Nd1iM1o2ymvA" colab_type="code" colab={}
# Install the nightly build
# !pip install -q tf-nightly-gpu==1.15.0.dev20190729
# !git clone https://github.com/tensorflow/tensorflow
# + [markdown] id="aV_0qkYh98LD" colab_type="text"
# ## Load TensorBoard
#
# Now, set up TensorBoard so that we can graph our accuracy and loss as training proceeds.
# + id="yZArmzT85SLq" colab_type="code" colab={}
# Delete any old logs from previous runs
# !rm -rf /content/retrain_logs
# Load TensorBoard
# %load_ext tensorboard
# %tensorboard --logdir /content/retrain_logs
# + [markdown] id="x1J96Ron-O4R" colab_type="text"
# ## Begin training
#
# Next, run the following script to begin training. The script will first download the training data:
# + id="VJsEZx6lynbY" colab_type="code" colab={}
# !python tensorflow/tensorflow/examples/speech_commands/train.py \
# --model_architecture=tiny_conv --window_stride=20 --preprocess=micro \
# --wanted_words=${WANTED_WORDS} --silence_percentage=25 --unknown_percentage=25 \
# --quantize=1 --verbosity=WARN --how_many_training_steps=${TRAINING_STEPS} \
# --learning_rate=${LEARNING_RATE} --summaries_dir=/content/retrain_logs \
# --data_dir=/content/speech_dataset --train_dir=/content/speech_commands_train \
#
# + [markdown] id="XQUJLrdS-ftl" colab_type="text"
# ## Freeze the graph
#
# Once training is complete, run the following cell to freeze the graph.
# + id="xyc3_eLh9sAg" colab_type="code" colab={}
# !python tensorflow/tensorflow/examples/speech_commands/freeze.py \
# --model_architecture=tiny_conv --window_stride=20 --preprocess=micro \
# --wanted_words=${WANTED_WORDS} --quantize=1 --output_file=/content/tiny_conv.pb \
# --start_checkpoint=/content/speech_commands_train/tiny_conv.ckpt-${TOTAL_STEPS}
# + [markdown] id="_DBGDxVI-nKG" colab_type="text"
# ## Convert the model
#
# Run this cell to use the TensorFlow Lite converter to convert the frozen graph into the TensorFlow Lite format, fully quantized for use with embedded devices.
# + id="lBj_AyCh1cC0" colab_type="code" colab={}
# !toco \
# --graph_def_file=/content/tiny_conv.pb --output_file=/content/tiny_conv.tflite \
# --input_shapes=1,1960 --input_arrays=Reshape_1 --output_arrays='labels_softmax' \
# --inference_type=QUANTIZED_UINT8 --mean_values=0 --std_dev_values=9.8077
# + [markdown] id="dt6Zqbxu-wIi" colab_type="text"
# The following cell will print the model size, which will be under 20 kilobytes.
# + id="XohZOTjR8ZyE" colab_type="code" colab={}
import os
model_size = os.path.getsize("/content/tiny_conv.tflite")
print("Model is %d bytes" % model_size)
# + [markdown] id="2pQnN0i_-0L2" colab_type="text"
# Finally, we use xxd to transform the model into a source file that can be included in a C++ project and loaded by TensorFlow Lite for Microcontrollers.
# + id="eoYyh0VU8pca" colab_type="code" colab={}
# Install xxd if it is not available
# !apt-get -qq install xxd
# Save the file as a C source file
# !xxd -i /content/tiny_conv.tflite > /content/tiny_conv.cc
# Print the source file
# !cat /content/tiny_conv.cc
|
tensorflow/lite/experimental/micro/examples/micro_speech/train_speech_model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import psycopg2
import pandas as pd
import psycopg2.extras
class PostgresConnection(object):
def __init__(self):
self.connection = psycopg2.connect(database="ecomdb",
user = "raihan",
password = "<PASSWORD>",
host = "127.0.0.1",
port = "5432")
def getConnection(self):
print("Connection to DB established!")
return self.connection
con = PostgresConnection().getConnection()
cur = con.cursor()
select_stmt = "SELECT s.division, SUM(t.total_price) " \
"FROM star_schema.fact_table t " \
"JOIN star_schema.store_dim s on s.store_key=t.store_key " \
"JOIN star_schema.time_dim tim on tim.time_key=t.time_key " \
"WHERE tim.month=12 " \
"GROUP BY s.division " \
"ORDER BY s.division"
cur.execute(select_stmt)
records = cur.fetchall()
records
import pandas as pd
df = pd.DataFrame(list(records), columns=['division', 'sales'])
df.dtypes
df['sales'] = df['sales'].astype('float64')
import matplotlib.pyplot as plt
df
pip install matplotlib
df = df.set_index(['division'])
df.plot.pie(y='sales', x='division', figsize=(5, 5))
# ### PANDAS Techiques:
# 1. Filter
# 2. Drop
# 3. iloc, loc
# 4. Aggregate
# 5. Index
# 6. Column selection, copy, crop
# 7. Merge, Concate
|
notebook/analytics/OLAP_tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
class Stack:
def __init__(self):
self.items=[]
def isEmpty(self):
return self.items==[]
def push(self,data):
self.items.append(data)
def size(self):
return len(self.items)
def show(self):
print (self.items)
def peek(self):
return self.items[len(self.items)-1]
def pop(self):
if not self.isEmpty():
return self.items.pop()
class Queue:
def __init__(self):
self.items = []
def isEmpty(self):
return self.items == []
def enqueue(self, item):
self.items.insert(0,item)
def dequeue(self):
return self.items.pop()
def palindrome_function(word):
s=Stack();
q=Queue();
l = len(word)//2
for i in range(0, l):
s.push(word[i])
for i in range(len(word) - l, len(word)):
q.enqueue(word[i])
if(str(s.items) == str(q.items)):
print(word,"is a palindrome");
else:
print(word,"is not a palindrome");
palindrome_function("a")
palindrome_function("ap")
palindrome_function("apa")
palindrome_function("epa")
palindrome_function("eppa")
palindrome_function("epapa")
palindrome_function("aaabbb")
palindrome_function("ababab")
palindrome_function("asddsa")
palindrome_function("asdfdsa")
# +
class Stack:
def __init__(self):
self.items=[]
def isEmpty(self):
return self.items==[]
def push(self,data):
self.items.append(data)
def size(self):
return len(self.items)
def show(self):
print (self.items)
def peek(self):
return self.items[len(self.items)-1]
def pop(self):
if not self.isEmpty():
return self.items.pop()
def check_matching_special_chars(chars):
s = "" if check_matching_special_chars_inner(chars) else "not "
return "'" + chars + "' is " + s + "valid.";
def check_matching_special_chars_inner(chars):
s=Stack();
i = 0;
while i < len(chars):
c = chars[i]
i += 1;
escaped = False;
if c == '\\':
escaped = True;
c = chars[i]
i += 1;
if c == '[':
s.push((c, escaped));
if c == ']':
p = s.pop();
if p[0] != '[' or p[1] != escaped:
return False;
if c == '(':
s.push(c);
if c == ')':
if s.pop() != '(':
return False;
if c == '{':
s.push((c, escaped));
if c == '}':
p = s.pop();
if p[0] != '{' or p[1] != escaped:
return False;
if c == '|':
p = s.peek();
if p[0] == '|' and p[1] == escaped:
s.pop();
else:
s.push((c, escaped));
return s.size() == 0;
print(check_matching_special_chars("\\[2\\]"))
print(check_matching_special_chars("\\[2]"))
print(check_matching_special_chars("()(l[k][k]))"))
print(check_matching_special_chars("[[](9)l)l]"))
print(check_matching_special_chars("((())"))
print(check_matching_special_chars("()[]{}"))
print(check_matching_special_chars("(5[][7]())"))
print(check_matching_special_chars("[[[][3]]]"))
print(check_matching_special_chars(""))
print(check_matching_special_chars("\[3+4"))
print(check_matching_special_chars("It costs $3."))
print(check_matching_special_chars("$\frac{1}{2}+5$"))
|
data science/DIT374 - python/assignment_5/Assignment-5-ver2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="dn-6c02VmqiN"
# ATTENTION: Please do not alter any of the provided code in the exercise. Only add your own code where indicated
# ATTENTION: Please do not add or remove any cells in the exercise. The grader will check specific cells based on the cell position.
# ATTENTION: Please use the provided epoch values when training.
# In this exercise you will train a CNN on the FULL Cats-v-dogs dataset
# This will require you doing a lot of data preprocessing because
# the dataset isn't split into training and validation for you
# This code block has all the required inputs
import os
import zipfile
import random
import shutil
import tensorflow as tf
from tensorflow.keras.optimizers import RMSprop
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from shutil import copyfile
from os import getcwd
# + colab={} colab_type="code" id="3sd9dQWa23aj"
# This code block unzips the full Cats-v-Dogs dataset to /tmp
# which will create a tmp/PetImages directory containing subdirectories
# called 'Cat' and 'Dog' (that's how the original researchers structured it)
path_cats_and_dogs = f"{getcwd()}/../tmp2/cats-and-dogs.zip"
shutil.rmtree('/tmp')
local_zip = path_cats_and_dogs
zip_ref = zipfile.ZipFile(local_zip, 'r')
zip_ref.extractall('/tmp')
zip_ref.close()
# + colab={} colab_type="code" id="gi3yD62a6X3S"
print(len(os.listdir('/tmp/PetImages/Cat/')))
print(len(os.listdir('/tmp/PetImages/Dog/')))
# Expected Output:
# 1500
# 1500
# + colab={} colab_type="code" id="F-QkLjxpmyK2"
# Use os.mkdir to create your directories
# You will need a directory for cats-v-dogs, and subdirectories for training
# and testing. These in turn will need subdirectories for 'cats' and 'dogs'
create_dir = [
'/tmp/cats-v-dogs',
'/tmp/cats-v-dogs/training',
'/tmp/cats-v-dogs/testing',
'/tmp/cats-v-dogs/training/cats',
'/tmp/cats-v-dogs/training/dogs',
'/tmp/cats-v-dogs/testing/cats',
'/tmp/cats-v-dogs/testing/dogs'
]
for directory in create_dir:
try:
os.mkdir(directory)
print(directory, 'created')
except OSError:
print(directory, 'failed')
# + colab={} colab_type="code" id="zvSODo0f9LaU"
# Write a python function called split_data which takes
# a SOURCE directory containing the files
# a TRAINING directory that a portion of the files will be copied to
# a TESTING directory that a portion of the files will be copie to
# a SPLIT SIZE to determine the portion
# The files should also be randomized, so that the training set is a random
# X% of the files, and the test set is the remaining files
# SO, for example, if SOURCE is PetImages/Cat, and SPLIT SIZE is .9
# Then 90% of the images in PetImages/Cat will be copied to the TRAINING dir
# and 10% of the images will be copied to the TESTING dir
# Also -- All images should be checked, and if they have a zero file length,
# they will not be copied over
#
# os.listdir(DIRECTORY) gives you a listing of the contents of that directory
# os.path.getsize(PATH) gives you the size of the file
# copyfile(source, destination) copies a file from source to destination
# random.sample(list, len(list)) shuffles a list
def split_data(SOURCE, TRAINING, TESTING, SPLIT_SIZE):
# YOUR CODE STARTS HERE
all_files = []
for file_name in os.listdir(SOURCE):
file_path = SOURCE + file_name
if os.path.getsize(file_path):
all_files.append(file_name)
else:
print('{} is zero length, so ignoring'.format(file_name))
n_files = len(all_files)
split_point = int(n_files * SPLIT_SIZE)
shuffled = random.sample(all_files, n_files)
train_set = shuffled[:split_point]
test_set = shuffled[split_point:]
for file_name in train_set:
copyfile(SOURCE + file_name, TRAINING + file_name)
for file_name in test_set:
copyfile(SOURCE + file_name, TESTING + file_name)
# YOUR CODE ENDS HERE
CAT_SOURCE_DIR = "/tmp/PetImages/Cat/"
TRAINING_CATS_DIR = "/tmp/cats-v-dogs/training/cats/"
TESTING_CATS_DIR = "/tmp/cats-v-dogs/testing/cats/"
DOG_SOURCE_DIR = "/tmp/PetImages/Dog/"
TRAINING_DOGS_DIR = "/tmp/cats-v-dogs/training/dogs/"
TESTING_DOGS_DIR = "/tmp/cats-v-dogs/testing/dogs/"
split_size = .9
split_data(CAT_SOURCE_DIR, TRAINING_CATS_DIR, TESTING_CATS_DIR, split_size)
split_data(DOG_SOURCE_DIR, TRAINING_DOGS_DIR, TESTING_DOGS_DIR, split_size)
# + colab={} colab_type="code" id="luthalB76ufC"
print(len(os.listdir('/tmp/cats-v-dogs/training/cats/')))
print(len(os.listdir('/tmp/cats-v-dogs/training/dogs/')))
print(len(os.listdir('/tmp/cats-v-dogs/testing/cats/')))
print(len(os.listdir('/tmp/cats-v-dogs/testing/dogs/')))
# Expected output:
# 1350
# 1350
# 150
# 150
# + colab={} colab_type="code" id="-BQrav4anTmj"
# DEFINE A KERAS MODEL TO CLASSIFY CATS V DOGS
# USE AT LEAST 3 CONVOLUTION LAYERS
model = tf.keras.models.Sequential([
# YOUR CODE HERE
tf.keras.layers.Conv2D(64, (3,3), input_shape=(150, 150, 3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Conv2D(32, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Conv2D(16, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(256, activation='relu'),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid')
])
model.compile(optimizer=RMSprop(lr=0.001), loss='binary_crossentropy', metrics=['acc'])
# -
# # NOTE:
#
# In the cell below you **MUST** use a batch size of 10 (`batch_size=10`) for the `train_generator` and the `validation_generator`. Using a batch size greater than 10 will exceed memory limits on the Coursera platform.
# + colab={} colab_type="code" id="mlNjoJ5D61N6"
TRAINING_DIR = '/tmp/cats-v-dogs/training'
#Performing Data Augmentation to train set
train_datagen = ImageDataGenerator(
rescale=1/255,
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest'
)
# NOTE: YOU MUST USE A BATCH SIZE OF 10 (batch_size=10) FOR THE
# TRAIN GENERATOR.
train_generator = train_datagen.flow_from_directory(
TRAINING_DIR,
batch_size=10,
class_mode='binary',
target_size=(150, 150)
)
VALIDATION_DIR = '/tmp/cats-v-dogs/testing'
#Performing Data Augmentation to validation set
validation_datagen = ImageDataGenerator(
rescale=1/255,
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest'
)
# NOTE: YOU MUST USE A BACTH SIZE OF 10 (batch_size=10) FOR THE
# VALIDATION GENERATOR.
validation_generator = validation_datagen.flow_from_directory(
VALIDATION_DIR,
batch_size=10,
class_mode='binary',
target_size=(150, 150)
)
# Expected Output:
# Found 2700 images belonging to 2 classes.
# Found 300 images belonging to 2 classes.
# + colab={} colab_type="code" id="KyS4n53w7DxC"
history = model.fit_generator(
train_generator,
epochs=2,
validation_data=validation_generator,
verbose=2)
# + colab={} colab_type="code" id="MWZrJN4-65RC"
# PLOT LOSS AND ACCURACY
# %matplotlib inline
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
#-----------------------------------------------------------
# Retrieve a list of list results on training and test data
# sets for each training epoch
#-----------------------------------------------------------
acc=history.history['acc']
val_acc=history.history['val_acc']
loss=history.history['loss']
val_loss=history.history['val_loss']
epochs=range(len(acc)) # Get number of epochs
#------------------------------------------------
# Plot training and validation accuracy per epoch
#------------------------------------------------
plt.plot(epochs, acc, 'r', "Training Accuracy")
plt.plot(epochs, val_acc, 'b', "Validation Accuracy")
plt.title('Training and validation accuracy')
plt.figure()
#------------------------------------------------
# Plot training and validation loss per epoch
#------------------------------------------------
plt.plot(epochs, loss, 'r', "Training Loss")
plt.plot(epochs, val_loss, 'b', "Validation Loss")
plt.title('Training and validation loss')
# Desired output. Charts with training and validation metrics. No crash :)
|
Tensorflow In Practice/Course 2 - Convolutional Neural Networks in Tensorflow/Cats vs Dogs using augmentation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
forward = "r1.fq"
reverse = "r2.fq"
# ## Pickling qualities
qualities = []
with open(forward,"r") as f:
qual = False
for line in f:
if line[0] == "+":
qual = True
continue
elif qual == True:
line = line.rstrip()
qualities.append(line)
qual = False
pickle.dump(qualities, open('fwd_qual.p','wb'))
qualities = []
with open(reverse,"r") as f:
qual = False
for line in f:
if line[0] == "+":
qual = True
continue
elif qual == True:
line = line.rstrip()
qualities.append(line)
qual = False
pickle.dump(qualities, open('rev_qual.p','wb'))
#
|
data/quality/.ipynb_checkpoints/markov_read_quality-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# + [markdown] papermill={"duration": 0.021109, "end_time": "2021-02-18T19:28:30.070970", "exception": false, "start_time": "2021-02-18T19:28:30.049861", "status": "completed"} tags=[]
# This notebook contains an example for teaching.
# + [markdown] papermill={"duration": 0.019635, "end_time": "2021-02-18T19:28:30.110372", "exception": false, "start_time": "2021-02-18T19:28:30.090737", "status": "completed"} tags=[]
# ## Introduction
# + [markdown] papermill={"duration": 0.019506, "end_time": "2021-02-18T19:28:30.149474", "exception": false, "start_time": "2021-02-18T19:28:30.129968", "status": "completed"} tags=[]
# In labor economics an important question is what determines the wage of workers. This is a causal question,
# but we could begin to investigate from a predictive perspective.
#
# In the following wage example, $Y$ is the hourly wage of a worker and $X$ is a vector of worker's characteristics, e.g., education, experience, gender. Two main questions here are:
#
#
# * How to use job-relevant characteristics, such as education and experience, to best predict wages?
#
# * What is the difference in predicted wages between men and women with the same job-relevant characteristics?
#
# In this lab, we focus on the prediction question first.
# + [markdown] papermill={"duration": 0.019453, "end_time": "2021-02-18T19:28:30.188193", "exception": false, "start_time": "2021-02-18T19:28:30.168740", "status": "completed"} tags=[]
# ## Data
#
# + [markdown] papermill={"duration": 0.019763, "end_time": "2021-02-18T19:28:30.227671", "exception": false, "start_time": "2021-02-18T19:28:30.207908", "status": "completed"} tags=[]
# The data set we consider is from the March Supplement of the U.S. Current Population Survey, year 2015. We select white non-hispanic individuals, aged 25 to 64 years, and working more than 35 hours per week during at least 50 weeks of the year. We exclude self-employed workers; individuals living in group quarters; individuals in the military, agricultural or private household sectors; individuals with inconsistent reports on earnings and employment status; individuals with allocated or missing information in any of the variables used in the analysis; and individuals with hourly wage below $3$.
#
# The variable of interest $Y$ is the hourly wage rate constructed as the ratio of the annual earnings to the total number of hours worked, which is constructed in turn as the product of number of weeks worked and the usual number of hours worked per week. In our analysis, we also focus on single (never married) workers. The final sample is of size $n=5150$.
# + [markdown] papermill={"duration": 0.019808, "end_time": "2021-02-18T19:28:30.267083", "exception": false, "start_time": "2021-02-18T19:28:30.247275", "status": "completed"} tags=[]
# ## Data analysis
# + [markdown] papermill={"duration": 0.01983, "end_time": "2021-02-18T19:28:30.307189", "exception": false, "start_time": "2021-02-18T19:28:30.287359", "status": "completed"} tags=[]
# We start by loading the data set.
# + papermill={"duration": 0.203333, "end_time": "2021-02-18T19:28:30.530976", "exception": false, "start_time": "2021-02-18T19:28:30.327643", "status": "completed"} tags=[]
load("wage2015_subsample_inference.Rdata")
dim(data)
# + [markdown] papermill={"duration": 0.020252, "end_time": "2021-02-18T19:28:30.572283", "exception": false, "start_time": "2021-02-18T19:28:30.552031", "status": "completed"} tags=[]
# Let's have a look at the structure of the data.
# + papermill={"duration": 0.081196, "end_time": "2021-02-18T19:28:30.674153", "exception": false, "start_time": "2021-02-18T19:28:30.592957", "status": "completed"} tags=[]
str(data)
# + [markdown] papermill={"duration": 0.020964, "end_time": "2021-02-18T19:28:30.716250", "exception": false, "start_time": "2021-02-18T19:28:30.695286", "status": "completed"} tags=[]
# We are constructing the output variable $Y$ and the matrix $Z$ which includes the characteristics of workers that are given in the data.
# + papermill={"duration": 0.054288, "end_time": "2021-02-18T19:28:30.791792", "exception": false, "start_time": "2021-02-18T19:28:30.737504", "status": "completed"} tags=[]
Y <- log(data$wage)
n <- length(Y)
Z <- data[-which(colnames(data) %in% c("wage","lwage"))]
p <- dim(Z)[2]
cat("Number of observation:", n, '\n')
cat( "Number of raw regressors:", p)
# + [markdown] papermill={"duration": 0.023423, "end_time": "2021-02-18T19:28:30.838653", "exception": false, "start_time": "2021-02-18T19:28:30.815230", "status": "completed"} tags=[]
# For the outcome variable *wage* and a subset of the raw regressors, we calculate the empirical mean to get familiar with the data.
# + papermill={"duration": 0.082765, "end_time": "2021-02-18T19:28:30.944279", "exception": false, "start_time": "2021-02-18T19:28:30.861514", "status": "completed"} tags=[]
library(xtable)
Z_subset <- data[which(colnames(data) %in% c("lwage","sex","shs","hsg","scl","clg","ad","mw","so","we","ne","exp1"))]
table <- matrix(0, 12, 1)
table[1:12,1] <- as.numeric(lapply(Z_subset,mean))
rownames(table) <- c("Log Wage","Sex","Some High School","High School Graduate","Some College","College Graduate", "Advanced Degree","Midwest","South","West","Northeast","Experience")
colnames(table) <- c("Sample mean")
tab<- xtable(table, digits = 2)
tab
# + [markdown] papermill={"duration": 0.022899, "end_time": "2021-02-18T19:28:30.992023", "exception": false, "start_time": "2021-02-18T19:28:30.969124", "status": "completed"} tags=[]
# E.g., the share of female workers in our sample is ~44% ($sex=1$ if female).
# + [markdown] papermill={"duration": 0.022996, "end_time": "2021-02-18T19:28:31.038176", "exception": false, "start_time": "2021-02-18T19:28:31.015180", "status": "completed"} tags=[]
# Alternatively, we can also print the table as latex.
# + papermill={"duration": 0.046315, "end_time": "2021-02-18T19:28:31.107521", "exception": false, "start_time": "2021-02-18T19:28:31.061206", "status": "completed"} tags=[]
print(tab, type="latex")
# + [markdown] papermill={"duration": 0.022601, "end_time": "2021-02-18T19:28:31.153542", "exception": false, "start_time": "2021-02-18T19:28:31.130941", "status": "completed"} tags=[]
# ## Prediction Question
# + [markdown] papermill={"duration": 0.022653, "end_time": "2021-02-18T19:28:31.199257", "exception": false, "start_time": "2021-02-18T19:28:31.176604", "status": "completed"} tags=[]
# Now, we will construct a prediction rule for hourly wage $Y$, which depends linearly on job-relevant characteristics $X$:
#
# \begin{equation}\label{decompose}
# Y = \beta'X+ \epsilon.
# \end{equation}
# + [markdown] papermill={"duration": 0.022687, "end_time": "2021-02-18T19:28:31.244814", "exception": false, "start_time": "2021-02-18T19:28:31.222127", "status": "completed"} tags=[]
# Our goals are
#
# * Predict wages using various characteristics of workers.
#
# * Assess the predictive performance using the (adjusted) sample MSE, the (adjusted) sample $R^2$ and the out-of-sample MSE and $R^2$.
#
#
# We employ two different specifications for prediction:
#
#
# 1. Basic Model: $X$ consists of a set of raw regressors (e.g. gender, experience, education indicators, occupation and industry indicators, regional indicators).
#
#
# 2. Flexible Model: $X$ consists of all raw regressors from the basic model plus occupation and industry indicators, transformations (e.g., ${exp}^2$ and ${exp}^3$) and additional two-way interactions of polynomial in experience with other regressors. An example of a regressor created through a two-way interaction is *experience* times the indicator of having a *college degree*.
#
# Using the **Flexible Model**, enables us to approximate the real relationship by a
# more complex regression model and therefore to reduce the bias. The **Flexible Model** increases the range of potential shapes of the estimated regression function. In general, flexible models often deliver good prediction accuracy but give models which are harder to interpret.
# + [markdown] papermill={"duration": 0.022935, "end_time": "2021-02-18T19:28:31.290982", "exception": false, "start_time": "2021-02-18T19:28:31.268047", "status": "completed"} tags=[]
# Now, let us fit both models to our data by running ordinary least squares (ols):
# + papermill={"duration": 0.070584, "end_time": "2021-02-18T19:28:31.384945", "exception": false, "start_time": "2021-02-18T19:28:31.314361", "status": "completed"} tags=[]
# 1. basic model
basic <- lwage~ (sex + exp1 + shs + hsg+ scl + clg + mw + so + we +occ2+ind2)
regbasic <- lm(basic, data=data)
regbasic # estimated coefficients
cat( "Number of regressors in the basic model:",length(regbasic$coef), '\n') # number of regressors in the Basic Model
# + [markdown] papermill={"duration": 0.023845, "end_time": "2021-02-18T19:28:31.433272", "exception": false, "start_time": "2021-02-18T19:28:31.409427", "status": "completed"} tags=[]
# ##### Note that the basic model consists of $51$ regressors.
# + papermill={"duration": 0.209505, "end_time": "2021-02-18T19:28:31.666828", "exception": false, "start_time": "2021-02-18T19:28:31.457323", "status": "completed"} tags=[]
# 2. flexible model
flex <- lwage ~ sex + shs+hsg+scl+clg+occ2+ind2+mw+so+we + (exp1+exp2+exp3+exp4)*(shs+hsg+scl+clg+occ2+ind2+mw+so+we)
regflex <- lm(flex, data=data)
regflex # estimated coefficients
cat( "Number of regressors in the flexible model:",length(regflex$coef)) # number of regressors in the Flexible Model
# + [markdown] papermill={"duration": 0.024845, "end_time": "2021-02-18T19:28:31.717639", "exception": false, "start_time": "2021-02-18T19:28:31.692794", "status": "completed"} tags=[]
# Note that the flexible model consists of $246$ regressors.
# + [markdown] papermill={"duration": 0.024777, "end_time": "2021-02-18T19:28:31.767579", "exception": false, "start_time": "2021-02-18T19:28:31.742802", "status": "completed"} tags=[]
# Try Lasso next
# + papermill={"duration": 6.836494, "end_time": "2021-02-18T19:28:38.629013", "exception": false, "start_time": "2021-02-18T19:28:31.792519", "status": "completed"} tags=[]
library(hdm)
flex <- lwage ~ sex + shs+hsg+scl+clg+occ2+ind2+mw+so+we + (exp1+exp2+exp3+exp4)*(shs+hsg+scl+clg+occ2+ind2+mw+so+we)
lassoreg<- rlasso(flex, data=data)
sumlasso<- summary(lassoreg)
# + [markdown] papermill={"duration": 0.025742, "end_time": "2021-02-18T19:28:38.682013", "exception": false, "start_time": "2021-02-18T19:28:38.656271", "status": "completed"} tags=[]
# Now, we can evaluate the performance of both models based on the (adjusted) $R^2_{sample}$ and the (adjusted) $MSE_{sample}$:
# + papermill={"duration": 0.167022, "end_time": "2021-02-18T19:28:38.874828", "exception": false, "start_time": "2021-02-18T19:28:38.707806", "status": "completed"} tags=[]
# Assess the predictive performance
sumbasic <- summary(regbasic)
sumflex <- summary(regflex)
# R-squared
R2.1 <- sumbasic$r.squared
cat("R-squared for the basic model: ", R2.1, "\n")
R2.adj1 <- sumbasic$adj.r.squared
cat("adjusted R-squared for the basic model: ", R2.adj1, "\n")
R2.2 <- sumflex$r.squared
cat("R-squared for the flexible model: ", R2.2, "\n")
R2.adj2 <- sumflex$adj.r.squared
cat("adjusted R-squared for the flexible model: ", R2.adj2, "\n")
R2.L <- sumlasso$r.squared
cat("R-squared for the lasso with flexible model: ", R2.L, "\n")
R2.adjL <- sumlasso$adj.r.squared
cat("adjusted R-squared for the flexible model: ", R2.adjL, "\n")
# + papermill={"duration": 0.167022, "end_time": "2021-02-18T19:28:38.874828", "exception": false, "start_time": "2021-02-18T19:28:38.707806", "status": "completed"} tags=[]
# calculating the MSE
MSE1 <- mean(sumbasic$res^2)
cat("MSE for the basic model: ", MSE1, "\n")
p1 <- sumbasic$df[1] # number of regressors
MSE.adj1 <- (n/(n-p1))*MSE1
cat("adjusted MSE for the basic model: ", MSE.adj1, "\n")
MSE2 <-mean(sumflex$res^2)
cat("MSE for the flexible model: ", MSE2, "\n")
p2 <- sumflex$df[1]
MSE.adj2 <- (n/(n-p2))*MSE2
cat("adjusted MSE for the flexible model: ", MSE.adj2, "\n")
MSEL <-mean(sumlasso$res^2)
cat("MSE for the lasso flexible model: ", MSEL, "\n")
pL <- length(sumlasso$coef)
MSE.adjL <- (n/(n-pL))*MSEL
cat("adjusted MSE for the lasso flexible model: ", MSE.adjL, "\n")
# + papermill={"duration": 0.070975, "end_time": "2021-02-18T19:28:38.984340", "exception": false, "start_time": "2021-02-18T19:28:38.913365", "status": "completed"} tags=[]
library(xtable)
table <- matrix(0, 3, 5)
table[1,1:5] <- c(p1,R2.1,MSE1,R2.adj1,MSE.adj1)
table[2,1:5] <- c(p2,R2.2,MSE2,R2.adj2,MSE.adj2)
table[3,1:5] <- c(pL,R2.L,MSEL,R2.adjL,MSE.adjL)
colnames(table)<- c("p","$R^2_{sample}$","$MSE_{sample}$","$R^2_{adjusted}$", "$MSE_{adjusted}$")
rownames(table)<- c("basic reg","flexible reg", "lasso flex")
tab<- xtable(table, digits =c(0,0,2,2,2,2))
print(tab,type="latex") # type="latex" for printing table in LaTeX
tab
# + [markdown] papermill={"duration": 0.03068, "end_time": "2021-02-18T19:28:39.045901", "exception": false, "start_time": "2021-02-18T19:28:39.015221", "status": "completed"} tags=[]
# Considering all measures above, the flexible model performs slightly better than the basic model.
#
# One procedure to circumvent this issue is to use **data splitting** that is described and applied in the following.
# + [markdown] papermill={"duration": 0.03052, "end_time": "2021-02-18T19:28:39.107361", "exception": false, "start_time": "2021-02-18T19:28:39.076841", "status": "completed"} tags=[]
# ## Data Splitting
#
# Measure the prediction quality of the two models via data splitting:
#
# - Randomly split the data into one training sample and one testing sample. Here we just use a simple method (stratified splitting is a more sophiscticated version of splitting that we can consider).
# - Use the training sample for estimating the parameters of the Basic Model and the Flexible Model.
# - Use the testing sample for evaluation. Predict the $\mathtt{wage}$ of every observation in the testing sample based on the estimated parameters in the training sample.
# - Calculate the Mean Squared Prediction Error $MSE_{test}$ based on the testing sample for both prediction models.
# + papermill={"duration": 0.053799, "end_time": "2021-02-18T19:28:39.191630", "exception": false, "start_time": "2021-02-18T19:28:39.137831", "status": "completed"} tags=[]
#splitting the data
set.seed(1) # to make the results replicable (generating random numbers)
random <- sample(1:n, floor(n*4/5))
# draw (4/5)*n random numbers from 1 to n without replacing them
train <- data[random,] # training sample
test <- data[-random,] # testing sample
dim(train)
# + papermill={"duration": 0.087305, "end_time": "2021-02-18T19:28:39.310008", "exception": false, "start_time": "2021-02-18T19:28:39.222703", "status": "completed"} tags=[]
# basic model
# estimating the parameters in the training sample
regbasic <- lm(basic, data=train)
regbasic
# + papermill={"duration": 0.087305, "end_time": "2021-02-18T19:28:39.310008", "exception": false, "start_time": "2021-02-18T19:28:39.222703", "status": "completed"} tags=[]
# calculating the out-of-sample MSE
trainregbasic <- predict(regbasic, newdata=test)
trainregbasic
# + papermill={"duration": 0.087305, "end_time": "2021-02-18T19:28:39.310008", "exception": false, "start_time": "2021-02-18T19:28:39.222703", "status": "completed"} tags=[]
y.test <- log(test$wage)
MSE.test1 <- sum((y.test-trainregbasic)^2)/length(y.test)
R2.test1<- 1- MSE.test1/var(y.test)
cat("Test MSE for the basic model: ", MSE.test1, " ")
cat("Test R2 for the basic model: ", R2.test1)
# + [markdown] papermill={"duration": 0.048538, "end_time": "2021-02-18T19:28:39.413804", "exception": false, "start_time": "2021-02-18T19:28:39.365266", "status": "completed"} tags=[]
# In the basic model, the $MSE_{test}$ is quite closed to the $MSE_{sample}$.
# + papermill={"duration": 0.17873, "end_time": "2021-02-18T19:28:39.624018", "exception": false, "start_time": "2021-02-18T19:28:39.445288", "status": "completed"} tags=[]
# flexible model
# estimating the parameters
#options(warn=-1)
regflex <- lm(flex, data=train)
# calculating the out-of-sample MSE
trainregflex<- predict(regflex, newdata=test)
y.test <- log(test$wage)
MSE.test2 <- sum((y.test-trainregflex)^2)/length(y.test)
R2.test2<- 1- MSE.test2/var(y.test)
cat("Test MSE for the flexible model: ", MSE.test2, " ")
cat("Test R2 for the flexible model: ", R2.test2)
# -
length(y.test)
# + [markdown] papermill={"duration": 0.050246, "end_time": "2021-02-18T19:28:39.732065", "exception": false, "start_time": "2021-02-18T19:28:39.681819", "status": "completed"} tags=[]
# In the flexible model, the discrepancy between the $MSE_{test}$ and the $MSE_{sample}$ is not large.
# + [markdown] papermill={"duration": 0.032619, "end_time": "2021-02-18T19:28:39.797362", "exception": false, "start_time": "2021-02-18T19:28:39.764743", "status": "completed"} tags=[]
# It is worth to notice that the $MSE_{test}$ vary across different data splits. Hence, it is a good idea average the out-of-sample MSE over different data splits to get valid results.
#
# Nevertheless, we observe that, based on the out-of-sample $MSE$, the basic model using ols regression performs is about as well (or slightly better) than the flexible model.
#
#
# Next, let us use lasso regression in the flexible model instead of ols regression. Lasso (*least absolute shrinkage and selection operator*) is a penalized regression method that can be used to reduce the complexity of a regression model when the number of regressors $p$ is relatively large in relation to $n$.
#
# Note that the out-of-sample $MSE$ on the test sample can be computed for any other black-box prediction method as well. Thus, let us finally compare the performance of lasso regression in the flexible model to ols regression.
# + papermill={"duration": 2.030191, "end_time": "2021-02-18T19:28:41.860172", "exception": false, "start_time": "2021-02-18T19:28:39.829981", "status": "completed"} tags=[]
# flexible model using lasso
# estimating the parameters
library(hdm)
reglasso <- rlasso(flex, data=train, post=FALSE)
# calculating the out-of-sample MSE
trainreglasso<- predict(reglasso, newdata=test)
MSE.lasso <- sum((y.test-trainreglasso)^2)/length(y.test)
R2.lasso<- 1- MSE.lasso/var(y.test)
cat("Test MSE for the lasso on flexible model: ", MSE.lasso, " ")
cat("Test R2 for the lasso flexible model: ", R2.lasso)
# + [markdown] papermill={"duration": 0.052309, "end_time": "2021-02-18T19:28:41.975337", "exception": false, "start_time": "2021-02-18T19:28:41.923028", "status": "completed"} tags=[]
# Finally, let us summarize the results:
# + papermill={"duration": 0.070703, "end_time": "2021-02-18T19:28:42.079561", "exception": false, "start_time": "2021-02-18T19:28:42.008858", "status": "completed"} tags=[]
table2 <- matrix(0, 3,2)
table2[1,1] <- MSE.test1
table2[2,1] <- MSE.test2
table2[3,1] <- MSE.lasso
table2[1,2] <- R2.test1
table2[2,2] <- R2.test2
table2[3,2] <- R2.lasso
rownames(table2)<- c("basic reg","flexible reg","lasso regression")
colnames(table2)<- c("$MSE_{test}$", "$R^2_{test}$")
tab2 <- xtable(table2, digits =3)
tab2
# + papermill={"duration": 0.052181, "end_time": "2021-02-18T19:28:42.166072", "exception": false, "start_time": "2021-02-18T19:28:42.113891", "status": "completed"} tags=[]
print(tab2,type="latex") # type="latex" for printing table in LaTeX
# -
|
R_Notebooks/pm1-notebook1-prediction-newdata.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: env_viz
# language: python
# name: env_viz
# ---
# + [markdown] tags=[]
# ## Clustering structures into chromosome Xi and Xa
# +
import os
original_path = os.getcwd()
root = '/rhome/yhu/bigdata/proj/experiment_GIST'
os.chdir(root)
new_path = os.getcwd()
print('redirect path: \n\t{} \n-->\t{}'.format(original_path, new_path))
print('root: {}'.format(root))
from GIST.prepare.utils import load_hic, load_hic, iced_normalization
from GIST.visualize import display, load_data
from GIST.validation.utils import load_df_fish3d, fish3d_format, load_tad_bed
from GIST.validation.utils import pdist_3d, load_tad_bed, select_loci, remove_failed
from GIST.validation.validation_tad import select_structure3d
from GIST.validation.ab import normalizebydistance, decomposition, correlation, plot
from GIST.validation.ab import fit_genomic_spatial_func
import torch
import numpy as np
from numpy import linalg as LA
from numpy.ma import masked_array
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
import seaborn as sns
import plotly.express as px
import networkx as nx
import tensorly as tl
from tensorly.decomposition import parafac2
from sklearn import preprocessing
from scipy.optimize import curve_fit
from scipy.spatial.distance import cdist, pdist, squareform
from scipy.spatial.transform import Rotation as R
from scipy.optimize import curve_fit
from scipy.spatial import distance
from scipy.cluster import hierarchy
from scipy.cluster.hierarchy import leaves_list
from scipy.stats import ttest_ind
import warnings
warnings.filterwarnings('ignore')
# +
chrom = 'X'
SAVE_FIG = True
font = {'size': 18}
matplotlib.rc('font', **font)
# setup saved_path
# load config .json
configuration_path = os.path.join(root, 'data')
configuration_name = 'config_predict_{}.json'.format(chrom)
info, config_data = load_data.load_configuration(configuration_path, configuration_name)
resolution = info['resolution']
# info['hyper'] = '10kb_predict_{}'.format(chrom)
# info['ncluster'] = 7
print(info['hyper'])
validation_name = 'XaXi'
saved_path = os.path.join(root, 'figures', validation_name, info['cell'], 'chr{}'.format(chrom))
os.makedirs(saved_path, exist_ok=True)
print('figure saved in {}'.format(saved_path))
# + [markdown] tags=[]
# ## Load FISH TAD data
# -
def get_fish3d(path, name):
fish_df = load_df_fish3d(path, name)
fish3d = fish3d_format(fish_df)
res = []
for i in np.arange(fish3d.shape[0]):
tmp = fish3d[i,:,:].squeeze()
tmpind = np.isnan(tmp[:,0])
if np.any(tmpind):
continue
res.append(tmp)
x0 = res[0].squeeze()
x0 = x0 - x0.mean(axis=0)
resR = []
for i, x in enumerate(res):
x = x - x.mean(axis=0)
eR, _ = R.align_vectors(x0, x)
x = eR.apply(x)
resR.append(x)
fish3d = np.stack(resR, axis=0)
nm_fish3d = np.nanmean(fish3d, axis=0)
print('fish3d shape: ', fish3d.shape)
print('ignoring NaNs mean fish3d shape: ', nm_fish3d.shape)
return nm_fish3d, fish3d
# +
path = os.path.join(root, 'data', 'FISH', 'geometry_coo')
name = 'FISH_Chr{}i.xyz'.format(chrom)
nm_fishXi3d, fishXi3d = get_fish3d(path, name)
path = os.path.join(root, 'data', 'FISH', 'geometry_coo')
name = 'FISH_Chr{}a.xyz'.format(chrom)
nm_fishXa3d, fishXa3d = get_fish3d(path, name)
# +
fishXi_pdist = pdist_3d(fishXi3d)
print('mean Xi pdist shape: ', np.nanmean(fishXi_pdist, axis=0).shape)
print('Xi pdist shape: ', fishXi_pdist.shape)
fishXa_pdist = pdist_3d(fishXa3d)
print('mean Xa pdist shape: ', np.nanmean(fishXa_pdist, axis=0).shape)
print('Xa pdist shape: ', fishXa_pdist.shape)
# + [markdown] tags=[]
# ## Density of distance matrices between Xa and Xi
# +
nm_fishXi_pdist = np.nanmean(fishXi_pdist, axis=0)
nm_fishXa_pdist = np.nanmean(fishXa_pdist, axis=0)
mask = np.mask_indices(nm_fishXi_pdist.shape[0], np.triu, 0)
filter_Xi = nm_fishXi_pdist[mask]
filter_Xa = nm_fishXa_pdist[mask]
dist = np.concatenate((filter_Xa, filter_Xi))
cluster = np.append(['active']*len(filter_Xa), ['inactive']*len(filter_Xi) )
d = {'Distance': dist, 'Chromosome X': cluster}
df = pd.DataFrame(data=d)
pal = {'active':'Red', 'inactive':'Blue'}
fig, axs = plt.subplots(1,1, figsize=(4, 4))
sns.boxplot(x="Chromosome X", y="Distance", palette=pal, data=df, ax=axs)
fig.tight_layout()
fig.show()
# -
# save the figure
if SAVE_FIG:
sp = os.path.join(saved_path, 'AI_True_distance_boxplot_{}.pdf'.format(chrom))
fig.savefig(sp, format='pdf')
sp = os.path.join(saved_path, 'AI_True_distance_boxplot_{}.png'.format(chrom))
fig.savefig(sp, format='png')
res = ttest_ind(filter_Xa, filter_Xi, alternative="greater")
print(res)
# + [markdown] tags=[]
# ## Load Prediction of 3D structures
# + tags=[]
# # load config .json
# configuration_path = os.path.join(root, 'data')
# configuration_name = 'config_predict_{}.json'.format(chrom)
# info, config_data = load_data.load_configuration(configuration_path, configuration_name)
# resolution = info['resolution']
# load dataset
dataset_path = os.path.join(root, 'data', info['cell'], info['hyper'])
dataset_name = 'dataset.pt'
HD = load_data.load_dataset(dataset_path, dataset_name)
graph, feat, ratio, indx = HD[0]
raw_id = graph['top_graph'].ndata['id'].cpu().numpy()
rmNaN_id = np.arange(len(raw_id))
raw2rm = {} # raw -> rm id
rm2raw = {} # rm -> raw id
for A, B in zip(raw_id, rmNaN_id):
raw2rm[A] = B
rm2raw[B] = A
# load prediction
prediction_path = os.path.join(root, 'data', info['cell'], info['hyper'], 'output')
prediction_name = 'prediction.pkl'
prediction = load_data.load_prediction(prediction_path, prediction_name)
print('load data path: {}, {}'.format(prediction_path, prediction_name) )
# assignment
structures = dict()
structures['GIST'] = prediction['{}_0'.format(chrom)]['structures']
xweights = prediction['{}_0'.format(chrom)]['structures_weights']
if xweights is not None:
xweights = xweights.astype(float).flatten()
print('proportion: {}'.format(100*xweights))
true_cluster = np.array(prediction['{}_0'.format(chrom)]['true_cluster'])
predict_cluster = np.array(prediction['{}_0'.format(chrom)]['predict_cluster'][0])
print( 'GIST structure shape: ', structures['GIST'].shape )
# -
path = os.path.join(root, 'data', 'FISH', 'loci_position')
name = 'hg19_Chr{}.bed'.format(chrom)
df = load_tad_bed(path, name)
select_idx = select_loci(df, resolution)
for i, idx in enumerate(select_idx):
arr = np.intersect1d(raw_id, idx)
select_idx[i] = np.array([ raw2rm[x] for x in arr ] )
data3d = structures['GIST']
index = select_idx
N = data3d.shape[0]
K = data3d.shape[1]
M = len(index)
res = np.empty((M, K, 3))
for i, idx in enumerate(index):
res[i, :, :] = np.nanmean( data3d[idx.astype(int), :, :], axis=0, keepdims=True)
print('res shape: ', res.shape)
xTAD_3d = res.transpose( (1, 0, 2) )
print('after transpose shape: ', xTAD_3d.shape)
xTAD_pdist = pdist_3d(xTAD_3d)
print('pdist shape: ', np.nanmean(xTAD_pdist, axis=0).shape)
# + [markdown] tags=[]
# ## Clustering
#
# - Decomposition: parafac2
# - Cluster: hierarchy linkage, metric: 'euclidean', method: 'average'
# -
best_err = np.inf
decomposition = None
tensor = xTAD_pdist
true_rank = info['ncluster']+1
print(true_rank)
for run in range(4):
print(f'Training model {run}', end=', ')
trial_decomposition, trial_errs = parafac2(tensor, true_rank, return_errors=True, tol=1e-8, n_iter_max=800, random_state=run)
print(f'Final error: {trial_errs[-1]}')
if best_err > trial_errs[-1]:
best_err = trial_errs[-1]
err = trial_errs
decomposition = trial_decomposition
print('-------------------------------')
print(f'Best model error: {best_err}')
est_tensor = tl.parafac2_tensor.parafac2_to_tensor(decomposition)
est_weights, (est_A, est_B, est_C) = tl.parafac2_tensor.apply_parafac2_projections(decomposition)
mest_A = np.mean(est_A, axis=0)
sign = np.sign(mest_A)
print(sign, mest_A)
mest_A = mest_A*sign
ind = np.argsort(mest_A)
est_A = est_A[:,ind]*sign[ind]
# +
col_linkage = hierarchy.linkage(distance.pdist(est_A, metric='euclidean'), method='ward', optimal_ordering=True)
rank = leaves_list(col_linkage)
cluster = hierarchy.fcluster(col_linkage, t=3, criterion='maxclust')
lut = dict(zip(set(cluster), sns.color_palette("RdBu", n_colors=len(set(cluster))+2)[::2] )) # sns.hls_palette(len(set(cluster)), l=0.5, s=0.8))
col_colors = pd.DataFrame(cluster)[0].map(lut)
cc1 = pd.DataFrame({'Cluster': col_colors.to_numpy()})
col_colors = pd.concat([cc1], axis=1)
fig = plt.figure()
df_nrx = pd.DataFrame(data=est_A.T)
g = sns.clustermap(data=df_nrx, col_linkage=col_linkage,
z_score=0, dendrogram_ratio=(.1, .55), row_cluster=False,
center=0, vmin=-3.0, vmax=3.0,
xticklabels=1, yticklabels=0,
cmap='RdBu_r', col_colors = col_colors,
figsize=(12, 5))
g.ax_heatmap.set_xticklabels(g.ax_heatmap.get_xmajorticklabels(), fontsize = 20)
g.fig.subplots_adjust(right=0.9, top=0.957)
g.ax_cbar.set_position((0.91, .2, .015, .2))
# g.fig.suptitle('Title')
# g.fig.tight_layout()
print('order of structures: {}'.format(rank) )
print('cluster of structures: {}'.format(cluster[rank]))
plt.show()
# -
# save the figure
if SAVE_FIG:
sp = os.path.join(saved_path, 'pred_pf2_linkage_{}.pdf'.format(chrom))
g.fig.savefig(sp, format='pdf')
sp = os.path.join(saved_path, 'pred_pf2_linkage_{}.png'.format(chrom))
g.fig.savefig(sp, format='png')
# + [markdown] tags=[]
# ### Display structures
# -
fig = plt.figure(figsize=(20,18))
print(rank)
color = np.arange(xTAD_3d.shape[1])
cmaps = ['Reds', 'Greys', 'Blues', 'Greens', 'Oranges',
'YlOrBr', 'RdPu', 'BuPu', 'GnBu', 'Purples',
'PuBu', 'YlGnBu', 'PuBuGn', 'BuGn', 'YlGn',
'YlOrRd', 'OrRd', 'Greys', 'PuRd']
for i, k in enumerate(rank):
X = xTAD_3d[k,:,:].squeeze()
ax = fig.add_subplot(5, 8, i+1, projection='3d')
# ax.axis('off')
cmp = plt.get_cmap(cmaps[cluster[k]-1])
ax.scatter(X[:,0], X[:,1], X[:,2], c=color, cmap=cmp)
ax.set_box_aspect((np.ptp(X[:,0]), np.ptp(X[:,1]), np.ptp(X[:,2])))
ax.set_facecolor('xkcd:salmon')
ax.set_facecolor((0.6, 0.6, 0.6))
ax.set_title('#{}, Cluster {}'.format(k, cluster[k]-1))
ax.view_init(elev=10., azim=190)
fig.tight_layout()
fig.show()
# save the figure
if SAVE_FIG:
sp = os.path.join(saved_path, 'pred_3D_{}.pdf'.format(chrom))
fig.savefig(sp, format='pdf')
# + [markdown] tags=[]
# ## Density of distance matrices between clusters
# +
mask = np.mask_indices(nm_fishXi_pdist.shape[0], np.triu, 0)
cluster_number = []
dist = []
print(rank)
for i, k in enumerate(rank):
m =xTAD_pdist[k, :, :][mask]
dist = np.append(dist, m)
cluster_number = np.append(cluster_number, [int(cluster[k])]*len(m) )
d = {'Distance': dist, 'Cluster': cluster_number}
df = pd.DataFrame(data=d)
# sns.set(rc={'figure.figsize':(20, 8)})
fig, axs = plt.subplots(1,1, figsize=(5,4))
pal = {1: "Red", 2: "White", 3: "Blue"}
g = sns.boxplot(x="Cluster", y="Distance", data=df, width=0.5, palette=pal, ax=axs)
axs.set_xticklabels(['Active', 'Intermediate', 'Inactive'])
act = df.loc[df['Cluster'] == 1]['Distance'].values
mid = df.loc[df['Cluster'] == 2]['Distance'].values
inact = df.loc[df['Cluster'] == 3]['Distance'].values
res = ttest_ind(act, inact, alternative="greater")
print(res)
res = ttest_ind(act, mid, alternative="greater")
print(res)
res = ttest_ind(mid, inact, alternative="greater")
print(res)
fig.tight_layout()
fig.show()
# -
# save the figure
if SAVE_FIG:
sp = os.path.join(saved_path, 'AI_pred_distance_boxplot_{}.pdf'.format(chrom))
fig.savefig(sp, format='pdf')
sp = os.path.join(saved_path, 'AI_pred_distance_boxplot_{}.png'.format(chrom))
fig.savefig(sp, format='png')
# + [markdown] tags=[]
# ## Rotation alignment
# -
def fit_func(x, a, b):
return a * x + b
# +
fx = np.concatenate((nm_fishXa3d.reshape(1,-1,3), nm_fishXi3d.reshape(1,-1,3)), axis=0)
rx = np.empty((fx.shape[0],xTAD_3d.shape[0]))
mask = np.mask_indices(nm_fishXi_pdist.shape[0], np.triu, 0)
fish_p = (nm_fishXi_pdist[mask] + nm_fishXa_pdist[mask])/2
xTAD_wpdist = np.empty_like(xTAD_pdist)
for k in np.arange(len(xweights)):
tmp = xTAD_pdist[k,:,:]
xTAD_wpdist[k,:,:] = xweights[k]*tmp*len(xweights)
xtad_p = np.nanmean(xTAD_wpdist, axis=0)[mask]
popt, pcov = curve_fit(fit_func, xtad_p, fish_p)
print(popt)
for i in np.arange(fx.shape[0]):
fx3d = fx[i,:,:].squeeze()
c_fx = fx3d - fx3d.mean(axis=0)
for j in np.arange(xTAD_3d.shape[0]):
x = xTAD_3d[j,:,:].squeeze()
x = x - x.mean(axis=0)
_, rmsdX = R.align_vectors(c_fx, x*popt[0]) #
rx[i,j] = rmsdX
T_F = (rx[0,:]>=rx[1,:]).reshape(1,-1)
T_ind = np.argwhere(T_F).flatten()
# +
fx = np.concatenate((fishXa3d, fishXi3d), axis=0)
rx = np.empty((fx.shape[0],xTAD_3d.shape[0]))
mask = np.mask_indices(nm_fishXi_pdist.shape[0], np.triu, 0)
fish_p = (nm_fishXi_pdist[mask] + nm_fishXa_pdist[mask])/2
xTAD_wpdist = np.empty_like(xTAD_pdist)
for k in np.arange(len(xweights)):
tmp = xTAD_pdist[k,:,:]
xTAD_wpdist[k,:,:] = xweights[k]*tmp*len(xweights)
xtad_p = np.nanmean(xTAD_wpdist, axis=0)[mask]
popt, pcov = curve_fit(fit_func, xtad_p, fish_p)
print(popt)
for i in np.arange(fx.shape[0]):
fx3d = fx[i,:,:].squeeze()
c_fx = fx3d - fx3d.mean(axis=0)
# d = np.sqrt( np.mean( np.sum(c_fx**2, axis=1)) )
for j in np.arange(xTAD_3d.shape[0]):
x = xTAD_3d[j,:,:].squeeze()
x = x - x.mean(axis=0)
_, rmsdX = R.align_vectors(c_fx, x*popt[0])
rx[i,j] = rmsdX
# +
nrx = preprocessing.normalize(rx, norm='l1', axis=1)
col_linkage = hierarchy.linkage(distance.pdist(nrx.T, metric='euclidean'), method='ward', optimal_ordering=True)
col_rank = leaves_list(col_linkage)
col_cluster = hierarchy.fcluster(col_linkage, t=5, criterion='maxclust')
col_lut = dict(zip(set(col_cluster), sns.color_palette("RdBu", n_colors=len(set(col_cluster))) )) # sns.hls_palette(len(set(cluster)), l=0.5, s=0.8))
col_colors1 = pd.DataFrame(col_cluster)[0].map(col_lut)
step = np.ceil(len(set(col_cluster))/len(set(cluster))).astype(int)
col_lut = dict(zip(set(cluster), sns.color_palette("RdBu", n_colors=len(set(col_cluster)))[::step] )) # sns.hls_palette(len(set(cluster)), l=0.3, s=0.6))
col_colors2 = pd.DataFrame(cluster)[0].map(col_lut)
cc1 = pd.DataFrame({'Linkage': col_colors1.to_numpy()})
cc2 = pd.DataFrame({'Cluster': col_colors2.to_numpy()})
col_colors = pd.concat([cc1, cc2], axis=1)
row_linkage = hierarchy.linkage(distance.pdist(nrx, metric='euclidean'), method='complete', optimal_ordering=True)
row_rank = leaves_list(row_linkage)
row_cluster = 3 - hierarchy.fcluster(row_linkage, t=2, criterion='maxclust')
tick_act = [1 for i in np.arange(len(fishXa3d))]
tick_inact = [2 for i in np.arange(len(fishXi3d))]
yticklabels = np.append(tick_act, tick_inact)
tick_act = ['a#{}'.format(i) for i in np.arange(len(fishXa3d))]
tick_inact = ['i#{}'.format(i) for i in np.arange(len(fishXi3d))]
ytickindex = np.append(tick_act, tick_inact)
row_lut = dict(zip(set(row_cluster), sns.color_palette("Set1", n_colors=len(set(row_cluster))) )) # sns.hls_palette(len(set(cluster)), l=0.5, s=0.8))
row_colors1 = pd.DataFrame(row_cluster)[0].map(row_lut)
rc1 = pd.DataFrame({'Linkage': row_colors1.to_numpy()})
row_lut = dict(zip(set(yticklabels), sns.color_palette("Set1", n_colors=len(set(yticklabels))) )) # sns.hls_palette(len(set(yticklabels)), l=0.3, s=0.6)))
row_colors2 = pd.DataFrame(yticklabels)[0].map(row_lut)
rc2 = pd.DataFrame({'Actual': row_colors2.to_numpy()})
row_colors = pd.concat([rc1, rc2], axis=1)
df_nrx = pd.DataFrame(data=nrx) #, index=ytickindex)
g = sns.clustermap(data=df_nrx, row_linkage=row_linkage, col_linkage=col_linkage,
z_score=None, dendrogram_ratio=(.15, .15),
center=np.mean(nrx), cbar_kws={"ticks":[nrx.min(), nrx.max()]},
row_colors = row_colors, # [row_colors1.to_numpy(), row_colors2.to_numpy()],
col_colors = col_colors, # [col_colors1.to_numpy(), col_colors2.to_numpy()],
xticklabels=1, yticklabels=0,
cmap='RdGy', figsize=(13, 10))
g.ax_cbar.set_yticklabels(['Low', 'High'])
g.fig.subplots_adjust(right=0.9, top=0.957)
g.ax_cbar.set_position((0.91, .2, .015, .5))
# g.fig.suptitle('Title')
g.fig.tight_layout()
# -
# save the figure
if SAVE_FIG:
sp = os.path.join(saved_path, 'alignment_AI_rmsd_allsamples_{}.pdf'.format(chrom))
g.fig.savefig(sp, format='pdf')
sp = os.path.join(saved_path, 'alignment_AI_rmsd_allsamples_{}.png'.format(chrom))
g.fig.savefig(sp, format='png')
# + [markdown] tags=[]
# ### Display Xa and Xi
# -
def align_zaxis(X):
v = X[0, :] - X[-1,:]
v = v/np.sqrt(v@v.T)
r = R.from_euler('z', np.arccos(v[1]), degrees=False)
X = r.apply(X)
r = R.from_euler('x', np.arccos(v[2]), degrees=False)
X = r.apply(X)
return X
# +
color = np.arange(xTAD_3d.shape[1])
cmaps = ['Reds', 'Blues']
y0 = nm_fishXa3d
y1 = nm_fishXi3d
y0 = y0 - y0.mean(axis=0)
y0 = align_zaxis(y0)
y1 = y1 - y1.mean(axis=0)
y1 = align_zaxis(y1)
fig = plt.figure(figsize=(10, 6))
angle = 45
ax = fig.add_subplot(1, 2, 1, projection='3d')
cmp = plt.get_cmap(cmaps[0])
ax.plot(y0[:,0], y0[:,1], y0[:,2], color='r')
ax.scatter(y0[:,0], y0[:,1], y0[:,2], color='r') # c=color, cmap=cmp)
ax.set_title('active')
A,B,C = y0[:,0], y0[:,1], y0[:,2]
ax.set_box_aspect((np.ptp(A), np.ptp(B), np.ptp(C)))
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_zlabel('Z')
ax.set_xlim(A.min()-.1, A.max()+.1)
ax.set_ylim(B.min()-.1, B.max()+.1)
ax.set_zlim(C.min()-.1, C.max()+.1)
ax.set_xticks([])
ax.set_yticks([])
ax.set_zticks([])
ax.view_init(30, angle)
# ax.set_facecolor('xkcd:salmon')
# ax.set_facecolor((0.5, 0.5, 0.5))
ax = fig.add_subplot(1, 2, 2, projection='3d')
ax.set_title('inactive')
cmp = plt.get_cmap(cmaps[1])
ax.plot3D(y1[:,0], y1[:,1], y1[:,2], color='b')
ax.scatter(y1[:,0], y1[:,1], y1[:,2], color='b') # c=color, cmap=cmp)
ax.set_box_aspect((np.ptp(A), np.ptp(B), np.ptp(C)))
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_zlabel('Z')
ax.set_xlim(A.min()-.1, A.max()+.1)
ax.set_ylim(B.min()-.1, B.max()+.1)
ax.set_zlim(C.min()-.1, C.max()+.1)
ax.set_xticks([])
ax.set_yticks([])
ax.set_zticks([])
ax.view_init(30, angle)
# ax.set_facecolor('xkcd:salmon')
# ax.set_facecolor((0.5, 0.5, 0.5))
fig.tight_layout()
fig.show()
# -
# save the figure
if SAVE_FIG:
sp = os.path.join(saved_path, 'True_mean_3D_{}.pdf'.format(chrom))
fig.savefig(sp, format='pdf', bbox_inches='tight')
sp = os.path.join(saved_path, 'True_mean_3D_{}.png'.format(chrom))
fig.savefig(sp, format='png', bbox_inches='tight')
# + [markdown] tags=[]
# ### Display structure
# + tags=[]
fig = plt.figure(figsize=(10, 6))
# color = np.arange(xTAD_3d.shape[0])
cmaps = ['Reds', 'Blues']
k0 = rank[0]
X0 = xTAD_3d[k0,:,:].squeeze()
X0 = X0 - X0.mean(axis=0)
X0 = align_zaxis(X0)
k1 = 17 # rank[-1]
X1 = xTAD_3d[k1,:,:].squeeze()
X1 = X1 - X1.mean(axis=0)
X1 = align_zaxis(X1)
axs = fig.add_subplot(1, 2, 1, projection='3d')
cmp = plt.get_cmap(cmaps[0])
r = R.from_euler('y', 90, degrees=True)
X0 = r.apply(X0)
A,B,C = X0[:,0], X0[:,1], X0[:,2]
axs.plot3D(X0[:,0], X0[:,1], X0[:,2], color='r')
axs.scatter(X0[:,0], X0[:,1], X0[:,2], color='r') # c=color, cmap=cmp
axs.set_box_aspect((np.ptp(A), np.ptp(B), np.ptp(C)))
axs.set_xlabel('X')
axs.set_ylabel('Y')
axs.set_zlabel('Z')
axs.set_title('#{} active'.format(k0))
axs.set_xlim(A.min()-20, A.max()+20)
axs.set_ylim(B.min()-20, B.max()+20)
axs.set_zlim(C.min()-20, C.max()+20)
axs.set_xticks([])
axs.set_yticks([])
axs.set_zticks([])
angle = 45
axs.view_init(30, angle)
# axs.set_facecolor('xkcd:salmon')
# axs.set_facecolor((0.5, 0.5, 0.5))
axs = fig.add_subplot(1, 2, 2, projection='3d')
cmp = plt.get_cmap(cmaps[1])
axs.plot3D(X1[:,0], X1[:,1], X1[:,2], color='b')
axs.scatter(X1[:,0], X1[:,1], X1[:,2], color='b', cmap='Reds') # c=color, cmap=cmp
axs.set_box_aspect((np.ptp(A), np.ptp(B), np.ptp(C)))
axs.set_xlabel('X')
axs.set_ylabel('Y')
axs.set_zlabel('Z')
axs.set_title('#{} inactive'.format(k1))
axs.set_xlim(A.min()-20, A.max()+20)
axs.set_ylim(B.min()-20, B.max()+20)
axs.set_zlim(C.min()-20, C.max()+20)
axs.set_xticks([])
axs.set_yticks([])
axs.set_zticks([])
angle = 45
axs.view_init(30, angle)
# axs.set_facecolor('xkcd:salmon')
# axs.set_facecolor((0.5, 0.5, 0.5))
fig.tight_layout()
fig.show()
# -
# save the figure
if SAVE_FIG:
sp = os.path.join(saved_path, 'pred_{}-{}_3D_{}.pdf'.format(k0, k1, chrom))
fig.savefig(sp, format='pdf',bbox_inches='tight')
sp = os.path.join(saved_path, 'pred_{}-{}_3D_{}.png'.format(k0, k1, chrom))
fig.savefig(sp, format='png',bbox_inches='tight')
|
TEST_XiXa.nbconvert.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
from bs4 import BeautifulSoup
import re
URL = 'http://www.columbia.edu/~fdc/sample.html'
response = requests.get(URL)
response
page = BeautifulSoup(response.text, 'html.parser')
page.title.string
page.find_all('h3')
# +
link_section = page.find('h3', attrs={'id': 'chars'})
section = []
for element in link_section.next_elements:
if element.name == 'h3':
break
section.append(element.string or '')
result = ''.join(section)
result
# -
page.find_all(re.compile('^h(2|3)'))
|
Chapter03/Scraping.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: silius
# language: python
# name: silius
# ---
# # Examining the rhyme scoring code
#
# This notebook is mainly to provide more insight into the rhyme scoring algorithm. In the end, the scoring code has quite a few moving parts, and it wasn't practical to try and explain every single thing in the paper, but the reviwers were keen to see the full details. Note that this code won't run standalone, I've just pulled out the core of the scoring code to explain how it works.
#
#
# ## Vowel similarity
#
# First let's look at the implementation of the vowel similarity. This is simply based on the closeness of the vowels of Latin according to a 'normal' linguistic formant frequency chart. The vowels of Latin are below. Note that I do not differentiate between long and short vowels, which renders a considerable amount of controversy moot. Allen in _Vox Latina_ posits a system in which some long vowels are positioned differently to the short ones. Weiss and Calabrese more or less suggest a 5-vowel system (not including the Greek y in their analysis) with identical quality for long and short. There is a good overview of the discussion on reddit [here](https://www.reddit.com/r/latin/comments/95yxez/vowel_pronunciation_beyond_allens_vox_latina/) (not exactly a scholarly source, but it's an efficient description by someone who clearly knows what they're talking about)
#
# 
#
# 10/11 bumped i-e slightly and o-a slightly based on
# Hirjee & Brown
NUCLEUS_SCORES = {
"i": {"i": 1, "e": 0.75, "a": 0.5, "o": 0.42, "u": 0.4, "ü": 0.5},
"e": {"i": 0.75, "e": 1, "a": 0.6, "o": 0.5, "u": 0.42, "ü": 0.5},
"a": {"i": 0.5, "e": 0.6, "a": 1, "o": 0.6, "u": 0.42, "ü": 0.4},
"o": {"i": 0.42, "e": 0.5, "a": 0.6, "o": 1, "u": 0.75, "ü": 0.35},
"u": {"i": 0.4, "e": 0.42, "a": 0.42, "o": 0.75, "u": 1, "ü": 0.6},
"ü": {"i": 0.5, "e": 0.5, "a": 0.4, "o": 0.35, "u": 0.6, "ü": 1},
}
# ## Consonant similarity
#
# In standard rhyme, the syllable onsets are ignored, but the codas are important (ie 'bat' and 'cat' are perfect rhymes but 'kit' and 'kin' are not). Wherever consonants are important, we need to consider the quality of imperfect rhymes, so 'cut' and 'cup' are better than 'cut' and 'cuff'. In this implementation I only create one level of similarity, so two consonants are either identical, similar or dissimilar. The code below determines that similarity based on phonological features, but it is slightly complicated by the fact that, to my ear, some pairs that sound similar in an onset do not match as well in a coda. Finally, for the final syllable (always unstressed in Latin) I do consider the onset so that things like /ra.bit/ and /ra.bid/ can be upgraded due to the matching 'b'.
#
# Essentially similar consonants just give a bonus to the rhyme score, but the exact details are a bit fiddly.
# +
# Define a bunch of feature classes as sets. These are fairly standard phonological classes.
# fricatives
FRIC = {"s", "f", "z", "h"}
# stops, voiced / unvoiced
UNV_STOP = {"k", "t", "p"}
V_STOP = {"g", "d", "b"}
STOP = UNV_STOP | V_STOP
ALVEOLAR = {"t", "d", "s", "z"}
VELAR = {"g", "k"}
# bilabial
BILAB = {"p", "b", "w"}
# sonorant
SON = {"n", "m", "l", "r"}
# nasal
NAS = {"n", "m"}
# approximants
APPROX = {"j", "w", "l", "r"}
CONT = SON | NAS | FRIC | {""}
CONS_CLOSE = {
"": FRIC | UNV_STOP | NAS | {""},
"t": ALVEOLAR | STOP,
"d": STOP,
"s": FRIC | (UNV_STOP - BILAB),
"f": FRIC,
"k": STOP - BILAB,
"h": STOP, # only occurs as kh and th which are both stops
"g": STOP - BILAB,
"r": SON,
"n": SON,
"m": CONT, # m isn't really there, it nasalises the vowel
"l": SON,
"b": (V_STOP | BILAB) - VELAR, # b--g seems too far away
"p": STOP - VELAR,
"x": UNV_STOP | FRIC,
"w": BILAB,
"j": APPROX,
}
CLOSE_STRESSED_CODA = {
"": FRIC | UNV_STOP,
"b": STOP,
"k": STOP,
"d": STOP,
"f": FRIC,
"g": STOP,
"h": STOP, # only occurs in coda as kh and th which are both stops
"j": APPROX,
"l": SON,
"m": SON,
"n": SON,
"p": STOP,
"r": SON,
"s": FRIC | (UNV_STOP - BILAB),
"t": ALVEOLAR | (UNV_STOP - BILAB),
"w": {"w"}, # should not appear in coda
"x": {"x"},
}
CLOSE_FINAL_ONSET = {
"b": STOP,
"k": VELAR,
"d": {"d", "t"},
"f": FRIC,
"g": VELAR,
"h": FRIC,
"j": APPROX,
"l": {"r"},
"m": NAS,
"n": NAS,
"p": STOP - VELAR,
"r": {"l"},
"s": FRIC | {"t"},
"t": FRIC | {"k", "d", "r"},
"w": APPROX,
"x": {"x"},
"": {""},
}
CLOSE_FINAL_CODA = {
"b": V_STOP,
"k": UNV_STOP,
"d": V_STOP,
"f": FRIC,
"g": VELAR,
"h": UNV_STOP,
"j": {"j"}, # shouldn't happen
"l": {"r"},
"m": NAS | {" "},
"n": NAS,
"p": UNV_STOP,
"r": {"l"},
"s": FRIC | {"t"},
"t": {"s", "p", "k", "d"},
"w": {"w"}, # shouldn't happen
"x": {"x"},
"": {""},
}
# -
# ## Nuclei
#
# Score the a pair of syllables according to the nucleus. Diphthongs are allowed, and we score them according to the final position (ie 'ae' ends at 'e').
def _score_nucleus(s1, s2):
if s1.nucleus == "" or s2.nucleus == "":
return 0
try:
# Basic score for the final vowel
nuc1 = s1.nucleus.translate(DEMACRON).lower()
nuc2 = s2.nucleus.translate(DEMACRON).lower()
v1 = s1.main_vowel
v2 = s2.main_vowel
score = NUCLEUS_SCORES[v1][v2]
# print("Basic score for %s %s: %.2f" % (s1,s2,score))
# One's a dipthong and one isn't, apply a penalty
if len(nuc1) != len(nuc2):
score *= 0.7
elif (nuc1 != nuc2) and (v1 == v2):
# two dipthongs but only last vowel equal
score *= 0.7
elif nuc1 == nuc2:
# mismatched nasalisation:
# if 1 (but not 0 or 2) of the nuclei is nasalised apply a small penalty
if len([x for x in [s1.nucleus, s2.nucleus] if COMBINING_TILDE in x]) == 1:
score *= 0.9
else:
# mismatched dipthongs or mismatched single letters
score = score
except Exception as e:
print(s1)
print(s2)
raise e
return score
# ## Syllable rhymes
#
# Now two methods for calulating the rhyme for two syllables. The algorithm is slightly different for the stressed syllable as compared to the final syllable. Some words also have a mismatched number of syllables involved in the rhyme, which receives a penalty.
# +
def _stressed_syl_rhyme(s1, s2):
# onset doesn't matter, less fussy about 'r' in coda
score = _score_nucleus(s1, s2)
last1 = s1.coda[-1:].lower()
last2 = s2.coda[-1:].lower()
try:
# perfect match receives a bonus
if s1.coda == s2.coda:
if s1.coda:
score *= 1.2
else:
score *= 1
elif len(s1.coda) + len(s2.coda) > 2:
# at least one consonant cluster
if "s" in s1.coda.lower() and "s" in s2.coda.lower():
# ast as are close
score *= 0.95
elif (
last2 in CLOSE_STRESSED_CODA[last1]
or last1 in CLOSE_STRESSED_CODA[last2]
):
# otherwise go by the final consonant - pakt part are close (?review?)
score *= 0.9
else:
score *= 0.8
elif last2 in CLOSE_STRESSED_CODA[last1] or last1 in CLOSE_STRESSED_CODA[last2]:
score *= 0.95
else:
score *= 0.8
except KeyError:
score *= 0.8
if score > 1:
score = 1
return score
def _final_syl_rhyme(s1, s2):
# TODO move the magic score multipliers into a config dict
# in the final syllable we apply a bonus
# for matching onsets, stricter about codas
score = _score_nucleus(s1, s2)
first1 = s1.onset[0:1].lower()
first2 = s2.onset[0:1].lower()
try:
if s1.onset == s2.onset:
score *= 1.1
elif len(s1.onset) + len(s2.onset) > 2:
# at least one cluster
if (
first2 in CLOSE_FINAL_ONSET[first1]
or first1 in CLOSE_FINAL_ONSET[first2]
):
# otherwise go by the initial consonant - tra and ta are close (?review?)
score *= 0.95
else:
score *= 0.85
elif first2 in CLOSE_FINAL_ONSET[first1] or first1 in CLOSE_FINAL_ONSET[first2]:
score *= 1
else:
score *= 0.85
except KeyError:
score *= 0.85
last1 = s1.coda[-1:].lower()
last2 = s2.coda[-1:].lower()
try:
# perfect match is good
if s1.coda == s2.coda:
if s1.coda:
score *= 1.2
else:
score *= 1.1
elif len(s1.coda) + len(s2.coda) > 2:
# at least one cluster
if "s" in s1.coda.lower() and "s" in s2.coda.lower():
# ast as are close
score *= 0.95
elif (
last2 in CLOSE_STRESSED_CODA[last1]
or last1 in CLOSE_STRESSED_CODA[last2]
):
# otherwise go by the final consonant - pakt part are close (?review?)
score *= 0.9
else:
score *= 0.8
elif last2 in CLOSE_STRESSED_CODA[last1] or last1 in CLOSE_STRESSED_CODA[last2]:
score *= 0.95
else:
score *= 0.8
except KeyError:
score *= 0.8
if score > 1:
score = 1
return score
def word_rhyme(w1, w2) -> (float):
"""Score the rhyme of two Words. Safe to call if one or
both of the words are None (will return 0).
Args:
w1, w2 (rhyme_classes.Word): words to score
Returns:
(float): The score.
"""
# This is so the user can call this with something
# like l[-1] vs l.midword, where midword might not exist
if not w1 or not w2:
return 0
# syls _might_ be empty, if the word is 'est' and it got eaten
# by the previous word (prodelision)
if len(w1.syls) == 0 or len(w2.syls) == 0:
return 0
if len(w1.syls) == 1 and len(w2.syls) == 1:
s = _final_syl_rhyme(w1.syls[0], w2.syls[0])
return s * 2
# calculate the rhyme score on the stressed syllable
stress_score = _stressed_syl_rhyme(w1.stressed_syllable, w2.stressed_syllable)
score = stress_score
# Now the rhyme on the remainder. In Latin, in theory,
# the final syllable is never stressed, so there should be
# at least one extra, but there _are_ exceptions.
# For uneven lengths, if we have Xx vs Yyy then compare
# the two final syllables, slurring over like
# UN.də.ground // COM.pound
coda_score = 0
if len(w1.post_stress) > 0 and len(w2.post_stress) > 0:
# single syllable words have their score doubled during
# final_syl_rhyme
coda_score = _final_syl_rhyme(w1.syls[-1], w2.syls[-1])
# bump up really good final syllable matches. This biases the approach
# somewhat since the final syllable is unstressed, but I have a pretty
# strong intuition that this sort of final-syllable assonance/slant-rhyme
# was important. On this see also Norberg (2004) An introduction to the
# study of medieval 1075 Latin versification (<NAME>, Trans.).
# Norberg traces the development of medeval rhyme to final-syllable assonances
# (some quite weak) in Sedulius in C4 CE, believing (as was common)
# that classical rhyme was only accidental.
if coda_score >= 0.75:
coda_score *= 1.3
# apply a small penalty for interstitial syllables between
# stressed and final if there's a length mismatch
# TODO: consider lightening this penalty. It was probably
# routine to swallow these interstitials in 'normal' speech
# and so perhaps too in poetry.
# "Sed Augustus quoque in epistulis ad C. Caesarem
# scriptis emendat quod is 'calidum' dicere quam 'caldum'
# malit, non quia id non sit Latinum, sed quia sit odiosum"
# (Quint. 1.6.19)
if len(w1.post_stress) + len(w2.post_stress) == 3:
# a 1 and a 2. This will be 99% of the cases. If it's
# not this then something weird is happening and the
# rest of the logic here might break.
longer = max(w1.post_stress, w2.post_stress, key=len)
# mid-low vowels (e,a,o) get pronounced as a schwa in the interstitial syllable
# but high ones (i,u,ü) sound more obtrusive to me.
if (
len(longer[1].nucleus.translate(DEMACRON).lower()) > 1
or longer[1].main_vowel in "iuü"
):
coda_score *= 0.73
else:
coda_score *= 0.83
score += coda_score
return score
# -
# # Scoring some words
#
# Here I'll just run through the kind of code used to produce Table 1 (the list of example rhyme scores)
# +
from mqdq import rhyme, babble
import random
import string
import scipy as sp
import pandas as pd
# -
met_single_bab = babble.Babbler.from_file('mqdq/OV-meta.xml', name='Metamorphoses')
# +
# this is now how I would normally syllabify, but if we want to examine
# individual word rhymes we need to take them before applying elision,
# prodelision etc. The 'normal' system calculates rhyme for the line
# as pronounced, ie if 'tua est' is at the end of a line the 'final' word
# is tuast, NOT est.
words = []
for l in met_single_bab.raw_source:
a = [rhyme._phonetify(rhyme._syllabify_word(x)) for x in l('word')]
words.extend(a)
# +
# Collect 25 random pairs of words whose rhyme score is
# above 1.75 (the global threshhold used in all the experiments)
pairs = []
while len(pairs) < 25:
w1, w2 = random.sample(words, 2)
a = w1.mqdq.text.translate(str.maketrans('', '', string.punctuation)).lower()
b = w2.mqdq.text.translate(str.maketrans('', '', string.punctuation)).lower()
if a==b:
continue
score, ss, cs = rhyme._word_rhyme_debug(w1,w2)
if 1.75 <= score:
pairs.append((w1,w2,(score,ss,cs)))
# -
def table(pairs):
res = []
for p in pairs:
score = p[2][0]
syls1 = ('.'.join(p[0].syls)).lower()
syls2 = ('.'.join(p[1].syls)).lower()
w1 = p[0].mqdq.text.translate(str.maketrans('', '', string.punctuation)).lower()
w2 = p[1].mqdq.text.translate(str.maketrans('', '', string.punctuation)).lower()
row = {
'orth1': w1,
'orth2': w2,
'phon1': syls1,
'phon2': syls2,
'score': score,
'stress': p[2][1],
'final': p[2][2],
}
res.append(row)
return pd.DataFrame(res)
# +
# Max possible score is 2.30.
table(pairs).sort_values(by='score')
# -
# # Future Work
#
# The scoring system seems 'good enough' to me in that it mostly captures 'rhymes' (which I use to mean interesting sonic correspondences) and mostly rejects uninteresting pairs. The ordering of scores can be a bit flaky, so it would be good to improve that at some point. Several reveiwers have expressed concern that ignoring vowel lengths sometimes causes pairs that score too highly. It would be great to reflect spoken vowel length, but it is a little tricky when we have vowels that lengthen to 'make position' (which technical Latin poetry thing)--it is not certain how those vowels were pronounced, all I would be able to say for sure is how the vowels were _scanned_. At that point we would need to sort through the phonological debate between 'Allen style' and 'Calabrese style' pronunciation constructions, which is not something I look forward to with extreme pleasure. The point is perhaps moot in any case, since in systemic Medieval rhyme, vowel length is ignored (Norberg (2004, 41)), so this may have been happening earlier, who knows.
|
repro/rhymescore_walkthrough.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/AngelRupido/OOP-1-1/blob/main/Python_Classes_and_Objects.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Ekx0-rxiyzpB"
# Class
# + id="K0Ecn5j6y580"
class MyClass:
pass # create a class without declaring variables and methods
# + colab={"base_uri": "https://localhost:8080/"} id="N13uHflUzvKh" outputId="b9b9b9dc-b6a0-4ac6-9ccb-dde786e70ce2"
class MyClass:
def __init__(self,name,age):
self.name = name # create a class with attributes
self.age = age
def display(self):
print(self.name,self.age)
person = MyClass("Angel", 19) # create an object name
print(person.name)
print(person.age)
# + colab={"base_uri": "https://localhost:8080/"} id="Q95pESRJ2vsQ" outputId="6a4aeea6-685d-48ad-b953-a6f78141d087"
class MyClass:
def __init__(self,name,age):
self.name = name # create a class with attributes
self.age = age
def display(self):
print(self.name,self.age)
person = MyClass("<NAME>", 19) # create an object name
person.display()
# + colab={"base_uri": "https://localhost:8080/"} id="q5COqzwf2-8p" outputId="62feec52-b664-4e8c-936e-8c9cea480343"
#Application 1 - Write a Python program that computes for an area of reactabgle A = lxw
class Rectangle:
def __init__(self,l,w):
self.l = l #attribute names
self.w = w
def Area(self):
print(self.l * self.w)
rect = Rectangle(7,3)
rect.Area()
|
Python_Classes_and_Objects.ipynb
|