code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Machine-Learning-Tokyo/Intro-to-GANs/blob/master/more_advanced/Cond_DCGAN_Interpolation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="bOSDAVHnwhwY" colab_type="text"
# ## Conditional DCGAN Interpolation
# + id="z4ws72Zto309" colab_type="code" cellView="form" colab={}
#@title Imports
# %%capture
# !pip install pydrive
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from oauth2client.client import GoogleCredentials
from google.colab import auth
from IPython.display import HTML
from keras.models import load_model
# + id="euZxlD1d7E5x" colab_type="code" cellView="form" colab={}
#@title Get utility functions
# %%capture
# %cd /content
# %rm -r /content/23dbccd7e2008a51dde92c5889e4d940
# !git clone https://gist.github.com/dkatsios/23dbccd7e2008a51dde92c5889e4d940.git
# %cd 23dbccd7e2008a51dde92c5889e4d940
from interpolation_utils import *
# %cd /content
# + id="VyGKzNYqpU_o" colab_type="code" cellView="form" colab={}
#@title Get trained generator model
# %%capture
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
# %cd /content
downloaded = drive.CreateFile({'id': '1Fi0nLclyq7qjyjWZLpKH56z3Jh6gzhho'})
downloaded.GetContentFile('fashion_cond_dcgan_gen.h5')
generator = load_model('/content/fashion_cond_dcgan_gen.h5')
# + [markdown] id="ecn9BsS9CamN" colab_type="text"
# ### Generate images
# + id="5aFbMqxFwaqq" colab_type="code" colab={}
images_num = 80
samples_gen = samples_generator(generator, images_num)
# + id="c2qCzKJ3k64C" colab_type="code" outputId="cf2fd485-e5ce-4730-cc2d-2d3004dae9bb" colab={"base_uri": "https://localhost:8080/", "height": 595}
imgs, noise, classes = next(samples_gen)
plot_imgs(imgs, images_num)
# + [markdown] id="33TpuOq6Ce9U" colab_type="text"
# ### Interpolate between images
# + id="KSrwCXf5n9RM" colab_type="code" colab={}
im_1 = 3, 1
im_2 = 8, 3
# + id="Y-0fXB-Po6pA" colab_type="code" cellView="form" outputId="d9a160bb-acd0-403a-ec19-35fe0230cf68" colab={"base_uri": "https://localhost:8080/", "height": 233}
#@title Plot interpolation
HTML(plot_results(*plot_inter(get_inter_imgs(im_1, im_2, noise, classes, generator)), inches=2).to_jshtml())
|
more_advanced/Cond_DCGAN_Interpolation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ###Optimising Maintenance Schedule
# In this part, we aim to determine the shortest route that will allow the maintenance crew to fix all the various wells that are either in need of repair or not functional. Some factors which we may wish to consider:
#
# i) Can we assign a higher priority to wells which are not functional as opposed to those that are merely not working?
#
# ii) Can we take advantage of route information to concentrate on higher quality roads?
#
# Initially we will ignore differences in location, height etc. and assume that all points on the map are equally accessible. To calculate the pairwise distance between points we need to take account of the fact that the Earth is a sphere so we need to use the Haversine formula:
#
# $$\rm{haversin} \Big(\frac{d}{r}\Big) = \rm{haversin}(\phi_2 - \phi_1) + \rm{cos}(\phi_1)\,\,\rm{cos}(\phi_2)\,\,\rm{haversin}(\lambda_1 - \lambda_2)$$
#
# $$\rm{haversin}(\theta) = \rm{sin}^2\Big(\frac{\theta}{2}\Big) = \frac{1 - \rm{cos}(\theta)}{2} $$
#
# where $d$ is the distance between the two points, $r$ is the radius of the sphere (Earth), $\phi$ is the latitude and $\theta$ is the longitude. This can be rearranged to give the following formula as described in (<NAME>, "Virtues of the Haversine," Sky and Telescope, vol. 68, no. 2, 1984, p. 159):
#
# $$\rm{dlon} = \rm{lon2} - \rm{lon1}$$
# $$\rm{dlat} = \rm{lat2} - \rm{lat1}$$
# $$\rm{a} = (\rm{sin}(\frac{dlat}{2}))^2 + cos(lat1) \times cos(lat2) \times (sin(\frac{dlon}{2}))^2$$
# $$\rm{c} = 2 \times \rm{arctan}(\frac{\sqrt{a}}{\sqrt{1-a}})$$
# $$\rm{d} = \rm{R} \times \rm{c}$$
#
# %matplotlib inline
import numpy as np
import scipy as sp
import scipy.spatial
import matplotlib.pyplot as plt
import pandas as pd
import random
import math
import time
import seaborn as sns
from math import radians, sin, cos, asin, sqrt, pi, atan2
def getDistanceByHaversine(latitudes, longitudes):
'''Haversine formula - give coordinates as a 2D numpy array of
(lat_decimal,lon_decimal) pairs'''
# earth's mean radius = 6,371km
EARTHRADIUS = 6371.0
# create meshgrid:
lat, lon = np.meshgrid(latitudes, longitudes)
# convert to radians
lat *= np.pi / 180.0
lon *= np.pi / 180.0
# get transposed meshgrids for distances
lat_T = lat.T.copy()
lon_T = lon.T.copy()
dlon = lon_T - lon
dlat = lat_T - lat
a = (np.sin(dlat/2))**2 + np.cos(lat) * np.cos(lat_T) * (np.sin(dlon/2.0))**2
c = 2.0 * np.arctan2(np.sqrt(a), np.sqrt(1.0-a))
km = EARTHRADIUS * c
return km
# +
from pandas import Series, DataFrame, Panel
#import data and labels
train_file = pd.read_csv('Waterpump-training-values.csv')
train_labels = pd.read_csv('Waterpump-training-labels.csv')
train_file['status_group'] = train_labels['status_group']
train_file = train_file[train_file['longitude'] > 1]
train_file = train_file[train_file['latitude'] < 0]
features = ['longitude','latitude','status_group']
trainLoc = train_file[features]
# pandas barplot
trainLoc['status_group'].value_counts().plot(kind='bar');
# +
trainLoc = trainLoc[trainLoc['status_group'] != 'functional']
#trainLoc.head()
#subsetting data to just take 1% to make it easier to work with
np.random.seed(142)
test_idx = np.random.uniform(0, 1, len(trainLoc)) <= 0.01
trainLoc = trainLoc[test_idx==True]
# +
x = np.array(trainLoc['longitude'])#.tolist()
y = np.array(trainLoc['latitude'])#.tolist()
#insert coordinates for Dar Es Salaam (39.2833 E 6.8000S)
x = np.insert(x, 0, 39.2833, 0)
y = np.insert(y, 0, -6.8000, 0)
DarEs = np.array([39.2833, -6.800])
#A = numpy.array((x, y, z), dtype=float)
#tmpx, tmpy = np.meshgrid(x,y)
cities = np.array((x, y), dtype=float)
#cities = np.array([tmpx, tmpy])
cities = np.reshape(cities, (2,-1)).T
#print cities.shape
#print cities
plt.scatter(cities[:,0], cities[:,1])
plt.scatter(DarEs[0],DarEs[1], s=50, color='red') #Highlight Dar Es Salaam on Map as HQ
# -
cities[:5]
# +
# the distance between two cities on a sphere is found using the Haversine formula
def get_distance(city1, city2):
'''Haversine formula - give coordinates as a 2D numpy array of
(lat_decimal,lon_decimal) pairs'''
#print city1[:], city2[:]
# earth's mean radius = 6,371km
EARTHRADIUS = 6371.0
# create meshgrid:
lat0, lon0 = city1[1], city1[0]
lat1, lon1 = city2[1], city2[0]
# convert to radians
lat0 *= np.pi / 180.0
lon0 *= np.pi / 180.0
lat1 *= np.pi / 180.0
lon1 *= np.pi / 180.0
# get transposed meshgrids for distances
#lat1_T = lat1.T.copy()
#lon1_T = lon1.T.copy()
#dlon = lon_T - lon
#dlat = lat_T - lat
dlon = lon1 - lon0
dlat = lat1 - lat0
a = (np.sin(dlat/2))**2 + np.cos(lat0) * np.cos(lat1) * (np.sin(dlon/2.0))**2
c = 2.0 * np.arctan2(np.sqrt(a), np.sqrt(1.0-a))
km = EARTHRADIUS * c
return km
# the energy for the whole system corresponds to
# the total distance the salesman has to travel
def distance(pathcities):
distance = 0.
number_of_cities = pathcities.shape[0]
# loop over all cities
for j in xrange(number_of_cities):
if j == number_of_cities - 1: # FINAL POINT CONNECTS WITH THE FIRST ONE
distance += get_distance( pathcities[j,:], pathcities[0,:] )
else:
distance += get_distance( pathcities[j,:], pathcities[j+1,:] )
return distance
#get_distance(cities[0], cities[1])
# +
# create a new path by swapping the connection between
# n_swaps cities randomly
def changepath(inputcities, n_swaps):
indices = range(1,inputcities.shape[0]) #Don't include starting city in swaps so that HQ is always the same
cities = inputcities.copy()
for i in range(n_swaps):
swappedCities = swapindex(cities)
cities=swappedCities.copy()
return cities
def swapindex(cities):
indices = range(cities.shape[0]) #Don't include starting city in swaps so that HQ is always the same
# take two random indices to swap
c1 = np.random.choice(indices[1:])
c2 = np.random.choice(indices[1:])
while c2 == c1:
c2 = np.random.choice(indices[1:])
# remember the cities to swap
tmp1 = cities[c1,:]
tmp2 = cities[c2,:]
# do the actual swapping
changedCities = cities.copy()
changedCities[c1,:] = tmp2
changedCities[c2,:] = tmp1
return changedCities
# -
print DarEs[0], DarEs[1]
def plot_path(pcities):
plt.plot(pcities[:,0], pcities[:,1],'o')
plt.plot(pcities[:,0], pcities[:,1])
plt.scatter(DarEs[0],DarEs[1], s=250, color='red') #Highlight Dar Es Salaam on Map as HQ
plt.xlim(np.min(pcities[:,0])-1, np.max(pcities[:,0])+1)
plt.ylim(np.min(pcities[:,1])-1, np.max(pcities[:,1])+1)
# +
# function for simulated annealing
# pathcities: array with our cities represented by their coordinates
# init_temp: initial temperature
# thermostat: linear factor to decrease the temperature
# ftol, itol, otol: tolerance values for stopping
# reannealing: schedule for reheating
def simulated_annealing( pathcities, init_temp, thermostat, ftol, itol, otol, reannealing):
# ===============
# SET THESE FOR DIAGNOSTICS
# ================
m=100000
distsize=otol+1
dist=[]
temperature = init_temp
N = pathcities.shape[0]
# number of accepted steps
it = 0
# DISTANCE HERE IS OUR ENERGY
prev_E = distance(pathcities)
# number of iterations
atp=0
didPlot = False
while it >=0:
#while otol < itol:
## NUMBER OF CORNERS IS L
L = np.max((np.floor(np.sqrt(temperature)).astype(int),1))
#print "L", L
#L = 2
propose_path = changepath(pathcities, L)
new_E = distance(propose_path)
deltaE = new_E -prev_E
if new_E < prev_E:
pathcities = propose_path
#dist[it] =new_E
dist.append(new_E)
prev_E = new_E
it = it+1
didPlot = False
elif np.random.rand() < np.exp( -deltaE/temperature):
pathcities = propose_path
#dist[it] =new_E
dist.append(new_E)
prev_E = new_E
it = it+1
didPlot = False
atp =atp +1; # NUMBER OF ITERATIONS
# check if it is time to cool down
if it % reannealing == 0:
temperature = thermostat * temperature;
#temperature = temperature/log(it);
compl_temp=0;
#if we get too cold, reheat
if temperature < 0.01:
temperature = 1
if False: #some optional plotting
if (it % 100 == 0) and not didPlot:
display.clear_output()
plt.plot( dist, '-r')
display.display(plt.gcf())
print len(dist)
print raw_input('Iteration: ' + np.str(atp))
plt.close()
didPlot = True
if len(dist)>m and np.std(dist[-m:])/np.mean(dist[-m:]) < ftol:
print 'ftol'
break
if atp >itol:
print 'itol'
break
if len(dist)> 0 and dist[-1] <= otol:
print 'otol'
print dist[-1]
break
s = pathcities
return s, dist
# -
# +
# simulated_annealing( pathcities, init_temp, thermostat, ftol, itol, otol, reannealing):
pcities, distances = simulated_annealing(cities, 10.0, 0.8, 0.01, 20000, 0, 10)
print len(distances)
# -
plt.plot(distances)
# +
plt.subplot(2,1,1)
plot_path(cities)
plt.subplot(2,1,2)
plot_path(pcities)
# -
#print distances[-1]
print cities[0:2], pcities[0:2]
# ###Things to do:
# 1) Set <NAME> (39.2833 E 6.8000S) as starting/ending point. DONE
#
# 2) Set Maximum Distance that can be travelled in one day or try multiple maintenance crews. KIND OF DONE
#
# 3) Priortize pumps that have a small number of nearest neighbours. KIND OF DONE
#
# 4) Travelling Purchaser Problem where non-functioning pumps cost more to repair than 'needs repair'. NOT DONE
print pcities[pcities.shape[0]/2:pcities.shape[0]/2 + 10]
#create multiple maintenance crews by splitting annealed cities into 3, all leaving from same depot.
cities1 = pcities[:pcities.shape[0]/3+1]
cities2 = pcities[pcities.shape[0]/3+1:2*pcities.shape[0]/3+1]
cities3 = pcities[2*pcities.shape[0]/3+1:]
cities2 = np.insert(cities2, 0, [39.2833, -6.8000], 0)
cities3 = np.insert(cities3, 0, [39.2833, -6.8000], 0)
print cities1[0], cities2[0], cities3[0]
# +
pcities1, distances1 = simulated_annealing(cities1, 10.0, 0.8, 0.01, 20000, 0, 10)
pcities2, distances2 = simulated_annealing(cities2, 10.0, 0.8, 0.01, 20000, 0, 10)
pcities3, distances3 = simulated_annealing(cities3, 10.0, 0.8, 0.01, 20000, 0, 10)
print "1: ", len(distances1), "2: ", len(distances2), "3: ", len(distances3)
# +
#I would have expected to see greater segregation of cities into distinct regions but possibily density is too high
plt.subplot(4,1,1)
plot_path(pcities)
plt.subplot(4,1,2)
plot_path(pcities1)
plt.subplot(4,1,3)
plot_path(pcities2)
plt.subplot(4,1,4)
plot_path(pcities3)
# -
print distances[-1]
print distances1[-1] + distances2[-1] + distances3[-1]
# +
#attempt to make comparison between sim anneal and genetic algorithm
start_time = time.clock()
p_big_mutate = 0.05
p_small_mutate = 0.4
fitness_scale=-0.5
pop_size=100
generations = 10**4
std_big = 1
std_small= 0.05
#def ras_fitness(g):
# ans = 20+g[:,0]**2+g[:,1]**2-10.0*(np.cos(2*np.pi*g[:,0])+np.cos(2*np.pi*g[:,1]))
# return ans**fitness_scale
def distance(pathcities):
distance = 0.
number_of_cities = pathcities.shape[0]
# loop over all cities
for j in xrange(number_of_cities):
if j == number_of_cities - 1: # FINAL POINT CONNECTS WITH THE FIRST ONE
distance += get_distance( pathcities[j,:], pathcities[0,:] )
else:
distance += get_distance( pathcities[j,:], pathcities[j+1,:] )
return distance**fitness_scale
def transform(population_orig):
# select two individuals for recombination
population =population_orig.copy()
indices = range(pop_size)
np.random.shuffle(indices)
temp = population[indices[0],1]
population[indices[0],1] = population[indices[1],1]
population[indices[1],1] = temp
#perform mutation
for i in range(pop_size):
if np.random.rand() < p_big_mutate:
population[i,0] = population[i,0]+std_big*np.random.randn()
if np.random.rand()<p_small_mutate:
population[i,0] = population[i,0]+std_small*np.random.randn()
if np.random.rand()<p_big_mutate:
population[i,1] = population[i,1]+std_big*np.random.randn()
if np.random.rand()<p_small_mutate:
population[i,1] = population[i,1]+std_small*np.random.randn()
return population
#generates initial population
mean=[100,100]
cov=[[9,0],[0,9]]
#g_0 = np.random.multivariate_normal(mean,cov,pop_size)
g_0 = cities[:100]
generation_fitness = np.zeros(generations)
#put placeholder for optimal solution
optimal_sol = [-100,-100]
g_curr=g_0
for z in range(generations):
if not z==0:
g_curr = transform(g_curr)
fit_curr = distance(g_curr)
generation_fitness[z] = fit_curr.max()
if z==0:
optimal_sol = g_curr[np.argmax(fit_curr),:]
elif generation_fitness[z]>generation_fitness[z-1]:
optimal_sol = g_curr[np.argmax(fit_curr),:]
marg_fit = fit_curr.cumsum()/fit_curr.sum()
r=np.random.rand(pop_size)
counts=np.zeros(pop_size)
for i in range(pop_size):
counts[i] = np.sum(marg_fit<=r[i])
child_counts = counts
g_new = []
for i in range(pop_size):
g_new.append(g_curr[child_counts[i],:])
g_curr=np.array(g_new)
end_time = time.clock()
# +
def cartesian_matrix(coords):
'''create a distance matrix for the city coords
that uses straight line distance'''
matrix={}
for i,(x1,y1) in enumerate(coords):
for j,(x2,y2) in enumerate(coords):
dx,dy=x1-x2,y1-y2
dist=sqrt(dx*dx + dy*dy)
matrix[i,j]=dist
return matrix
matrix = cartesian_matrix(pcities)
#print matrix
# -
print optimal_sol
print end_time - start_time
plt.plot(generation_fitness)
plt.show()
# ###Prioritising remote pumps
# Here we try to develop a weighting to encourage the maintenance crews to visit the most remote pumps first. We identify remoteness by performing KDTree analysis of the average distance between the 5 nearest neighbours. We then add this as a weight to the distance function that the SA uses to optimise the routes. Currently the implementation means that the distance reported are no longer true but the order will be reflective of this new priority and leads to longer routes as the crews no longer move to the optimal step as determined by simple distance.
kdt = scipy.spatial.cKDTree(pcities1)
k = 5 # number of nearest neighbors
dists, neighs = kdt.query(pcities1, k+1)
avg_dists = np.mean(dists[:, 1:], axis=1)
# +
#avg_dists[:10]
#np.concatenate((a, b.T), axis=1)
avg_dists = avg_dists.reshape((pcities1.shape[0],1))
#avg_dists.shape
cities1 = np.concatenate((pcities1, avg_dists), axis=1)
print cities1.shape
# -
avg_dists.sort()
plt.hist(avg_dists)
# +
def get_distance(city1, city2):
'''Haversine formula - give coordinates as a 2D numpy array of
(lat_decimal,lon_decimal) pairs'''
#print city1[:], city2[:]
# earth's mean radius = 6,371km
EARTHRADIUS = 6371.0
# retrieve coords:
lat0, lon0 = city1[1], city1[0]
lat1, lon1 = city2[1], city2[0]
# convert to radians
lat0 *= np.pi / 180.0
lon0 *= np.pi / 180.0
lat1 *= np.pi / 180.0
lon1 *= np.pi / 180.0
# get transposed meshgrids for distances
#lat1_T = lat1.T.copy()
#lon1_T = lon1.T.copy()
#dlon = lon_T - lon
#dlat = lat_T - lat
dlon = lon1 - lon0
dlat = lat1 - lat0
a = (np.sin(dlat/2))**2 + np.cos(lat0) * np.cos(lat1) * (np.sin(dlon/2.0))**2
c = 2.0 * np.arctan2(np.sqrt(a), np.sqrt(1.0-a))
#add weight to encourage visits to remote pumps (i.e. higher avg_dist for kdt)
w = 1.0/(city1[2]*city2[2])
#print "w: ", w
km = EARTHRADIUS * c * w
return km
# the energy for the whole system corresponds to
# the total distance the salesman has to travel
def distance(pathcities):
distance = 0.
number_of_cities = pathcities.shape[0]
# loop over all cities
for j in xrange(number_of_cities):
if j == number_of_cities - 1: # FINAL POINT CONNECTS WITH THE FIRST ONE
distance += get_distance( pathcities[j,:], pathcities[0,:] )
else:
distance += get_distance( pathcities[j,:], pathcities[j+1,:] )
return distance
#get_distance(cities[0],cities[1])
# -
cities1.shape
# +
pcities4, distances4 = simulated_annealing(cities1, 10.0, 0.8, 0.01, 20000, 0, 10)
print len(distances4)
# -
plt.plot(distances)
#print distances[-1]
# +
plt.subplot(2,1,1)
plot_path(pcities1)
plt.subplot(2,1,2)
plot_path(pcities4)
# -
print "distance optimised cities: ", pcities1[:10], "remote prioritized cities: ", pcities4[:10]
|
Optimising_maintenance_route.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basics
#
# * YANG RFC: https://tools.ietf.org/html/rfc6020
# * YANG Stanards repo: https://github.com/YangModels/yang
# * Cisco Suuport for NetConf: https://developer.cisco.com/site/standard-network-devices/
#
# ## Install initial libraries
#
# * use `pip install ncclient pyang napalm` to install base libraries
# * `ncclient` connects remote hosts over netconf
# * `pyang` converts YANG XML payload into readable tree format
# * `napalm` conencts remote hosts for CLI configuration
import ncclient
import pyang
# ## Access details
#
# Developers and network engineers access the IOS XE on CSR Recommended Code Always On Sandbox directly using the following information from Cisco IOS XE Sandbox https://devnetsandbox.cisco.com/RM/Diagram/Index/27d9747a-db48-4565-8d44-df318fce37ad?diagramType=Topology
# * CSR1000V Host: ios-xe-mgmt.cisco.com
# * SSH Port: 8181
# * NETCONF Port: 10000
# * RESTCONF Ports: 9443 (HTTPS)
# * Username: developer
# * Password: <PASSWORD>
# ## import
from ncclient import manager
import pyang
# ## `get_capabilities()`
#
# This functions prints all the supported capabilities from a given device
def get_capabilities(m):
print('***Here are the Remote Devices Capabilities***')
for capability in m.server_capabilities:
print(capability.split('?')[0])
# ## `get_schema()`
#
# This function prints some common schema supported by a given device. It's a menu driven function to choose from the follwoing schema.
# * IOS-XE native
# * IOS-Interface
# * IETF-Interface
def get_schema(m):
bad_choice=False
print('Select a schema...')
print('\t [1] IOS-XE')
print('\t [2] IOS-interface')
print('\t [3] IETF-interface')
choice=int(input('Enter your choice...'))
if choice == 1:
s = 'Cisco-IOS-XE-native'
elif choice == 2:
s = 'Cisco-IOS-XE-interface-common'
elif choice == 3:
s = 'ietf-interfaces'
else:
bad_choice=True
if bad_choice:
schema = 'Invalid Choice...'
else:
schema = m.get_schema(s)
print(schema)
# ## `get_running(m, to_json)`
#
# This function prints the running config of a given device with session variable `m`. The `to_json` vatiable is a boolian, if set as True the outcome is in JSON, otherwise in XML
def get_running(m, to_json):
import xmltodict
import json
response = m.get_config('running')
if to_json:
print(json.dumps(
xmltodict.parse(
str(response) # parse() expects sring
),sort_keys=True, indent=4) # to beautify json
)
else:
print(response)
# ## `get_config(m, to_json)`
def get_config(m, to_json):
import xmltodict # to convert XML into dictionary
import json # to convert dictionary to JSON
print('******** \nOptions\n********')
print('\t [1] hostname')
print('\t [2] Username')
print('\t [3] Interface Config')
print('\t [4] Static Route')
print('\t [5] Routing protocol')
choice = int(input('Enter your choice... '))
if choice == 1: #hostname
_filter="""
<filter>
<native xmlns="http://cisco.com/ns/yang/Cisco-IOS-XE-native">
<hostname></hostname>
</native>
</filter>
"""
if choice == 2: #username
_filter="""
<filter>
<native xmlns="http://cisco.com/ns/yang/Cisco-IOS-XE-native">
<username></username>
</native>
</filter>
"""
if choice == 3: #interface
_filter="""
<filter>
<interfaces xmlns="urn:ietf:params:xml:ns:yang:ietf-interfaces">
<interface>
<name></name>
<description></description>
<enabled></enabled>
<ipv4 xmlns="urn:ietf:params:xml:ns:yang:ietf-ip">
<address>
<ip></ip>
<netmask></netmask>
</address>
</ipv4>
</interface>
</interfaces>
</filter>
"""
if choice == 4: #static route
_filter="""
<filter>
<interfaces xmlns="http://openconfig.net/yang/interfaces">
<ip><route></route></ip>
</native>
</filter>
"""
if choice == 5: # dynamic routes
_filter="""
<filter>
<routing xmlns="urn:ietf:params:xml:ns:yang:ietf-routing">
<routing-instance>
<name></name>
<description></description>
<routing-protocols>
<routing-protocol></routing-protocol>
</routing-protocols>
</routing-instance>
</routing>
</filter>
"""
response = m.get_config('running',_filter)
if to_json:
print(json.dumps(
xmltodict.parse(
str(response) # parse() expects sring
),sort_keys=True, indent=4) # to beautify json
)
else:
print(response)
def main():
#assign global variables
host='ios-xe-mgmt.cisco.com'
ssh_port=8181
nc_port=10000
rc_port=9443
uname='developer'
pward='C1sco12345'
print(f'Connecting... {host} ')
with manager.connect( host=host,
port=10000,
username=uname,
password=<PASSWORD>,
hostkey_verify=False,
device_params={'name': 'default'},
look_for_keys=False, allow_agent=False
) as m:
while True:
#os.system('clear')
print('[OK]')
print('OPTIONS')
print('\t [1] Get Capabilities')
print('\t [2] Get Schema')
print('\t [3] Get Running Cofig')
print('\t [4] Get Basic infor')
print('\t [0] Exit')
choice = int(input('Enter your choice...'))
if choice == 0:
break
elif choice == 1:
get_capabilities(m)
elif choice == 2:
get_schema(m)
elif choice == 3:
get_running(m, to_json=False)
elif choice == 4:
get_basic(m, to_json=True)
else:
print('Sorry invalid selection... ')
main()
# ## Accessing the device using Napalm over SSH
# +
import napalm as npm
#assign global variables
host='ios-xe-mgmt.cisco.com'
ssh_port=8181
nc_port=10000
rc_port=9443
uname='developer'
pward='C1sco12345'
driver = npm.get_network_driver('iosxr')
router = driver(hostname = 'ios-xe-mgmt.cisco.com',
username = 'developer',
password = '<PASSWORD>',
optional_args={"port": 8181}
)
print('Connecting...')
router.open()
print('[OK]')
router.close()
print('Connection Closed..')
# -
# ## Connecting Using Raw SSH
import paramiko
# +
#assign global variables
host='ios-xe-mgmt.cisco.com'
ssh_port=8181
nc_port=10000
rc_port=9443
uname='developer'
pword='<PASSWORD>'
ssh = paramiko.SSHClient()
ssh.set_missing_host_key_policy(paramiko.AutoAddPolicy())
print('Connecting using SSH...')
ssh.connect(host, ssh_port, uname, pword)
cmd = input('Enter a Command: ')
stdin, stdout, stderr = ssh.exec_command(cmd)
for line in stdout.readlines():
print(line)
|
ENCOR/Network Programmability .ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
print(pd.__version__)
df = pd.read_excel('data/src/sample.xlsx', index_col=0)
print(df)
print(type(df))
df_sheet_index = pd.read_excel('data/src/sample.xlsx', sheet_name=0, index_col=0)
print(df_sheet_index)
df_sheet_name = pd.read_excel('data/src/sample.xlsx', sheet_name='sheet2', index_col=0)
print(df_sheet_name)
df_sheet_multi = pd.read_excel('data/src/sample.xlsx', sheet_name=[0, 'sheet2'], index_col=0)
print(type(df_sheet_multi))
print(len(df_sheet_multi))
print(df_sheet_multi.keys())
print(df_sheet_multi[0])
print(type(df_sheet_multi[0]))
print(df_sheet_multi['sheet2'])
print(type(df_sheet_multi['sheet2']))
df_sheet_all = pd.read_excel('data/src/sample.xlsx', sheet_name=None, index_col=0)
print(type(df_sheet_all))
print(df_sheet_all.keys())
df_header_index = pd.read_excel('data/src/sample.xlsx', header=None, index_col=None)
print(df_header_index)
print(df_header_index.columns)
print(df_header_index.index)
df_default = pd.read_excel('data/src/sample.xlsx')
print(df_default)
print(df_default.columns)
print(df_default.index)
print(pd.read_excel('data/src/sample.xlsx', index_col=0))
df_use_skip = pd.read_excel('data/src/sample.xlsx', index_col=0,
usecols=[0, 1, 3], skiprows=[1], skipfooter=1)
print(df_use_skip)
|
notebook/pandas_read_excel.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# # Statistics Exercise
# ## Statistical Distributions. Properties of distributions. Applications of Probability and Statistics in Computer Science
# ### Problem 1. Plotting a Single Distribution. Digits in $\pi$ and $e$
# We expect that the decimal digits in $\pi$ and $e$ will be randomly distributed and there's no reason for any digit to dominate over others. Let's verify this.
#
# Using an algorithm, the first 10 004 digits of $\pi$ and $e$ were generated:
# $$
# \pi = 3.(141592 \dots 5678)5667
# $$
# $$
# e = 2.(718281 \dots 6788)5674
# $$
#
# The 10 000 digits in brackets were counted. You can see the results in `digits.dat`. Each column corresponds to one digit from 0 to 9. The first row is for $\pi$ and the second row is for $e$.
#
# How are these digits distributed? Are the two distributions different?
#
# **Note:** The dataset is **not properly formatted** to work easily. You can transpose it. Now, digit counts will be in rows and variables - in columns.
# ```python
# digits = pd.read_table("digits.dat", header = None).T
# ```
#
# You can also specify column names like this:
# ```python
# digits.columns = ["pi", "e"]
# ```
#
# Also note that **we are not creating the histogram of the distribution**. We already have the counts, we need to plot them. In a sense, the histogram has already been calculated.
#
# To do this, we can create a "bar chart" (using `plt.bar()`). We have to provide values for the x-axis and y-axis. For the x-axis, we have the numbers 0 through 9 (we can use the *index* of the dataset like this: `digits.index`). For the y-axis, we need to plot the digit counts directly.
#
# We can see that even the simplest datasets sometimes need a bit of preprocessing. This is always the case when we're working with data.
digits = pd.read_table('data\digits.dat', header = None)
digits = digits.T
digits.columns = ['pi', 'e']
digits
mean_count_pi = digits['pi'].mean()
mean_count_pi
def plot_digits(digits, column, number_name, color_bar, color_line):
mean_number = column.mean()
plt.bar(digits.index, column, color = color_bar)
plt.axhline(mean_number, color = color_line, label = 'Mean')
plt.xlabel('Digit')
plt.ylabel('Count')
plt.xticks(digits.index, digits.index)
plt.title(f'Distribution of digits in {number_name}')
plt.legend(loc = 'lower right')
# plt.show()
plot_digits(digits, digits['pi'], '$\pi$', 'skyblue', 'red')
plt.show()
plot_digits(digits, digits['e'], '$e$', 'skyblue', 'red')
plt.show()
# +
plt.bar(digits.index, digits.pi,label = "pi", color = "red")
plt.bar(digits.index, digits.e,label = "e")
plt.xlabel('Digit')
plt.ylabel('Count')
plt.xticks(digits.index, digits.index)
plt.title(f'Distribution of digits in $\pi$ and $e$')
plt.legend(loc = 'lower right')
plt.show()
# -
# Let's try something else. Scientists have measured the percentage of silica ($\text{SiO}_2$, sand / glass) for 22 meteors. You can find it in `silica.dat`. How are these distributed? What is a "typical" percentage? Is there such percentage at all?
#
# Print the mean, standard deviation (you can use the biased or unbiased formula), skewness and kurtosis of the distribution. What do these numbers tell you? How do they relate to the shape of the distribution? Can you characterize the distribution better? (An idea would be to characterize different parts of it on their own, as if they're different distributions.)
silica_data = pd.read_table('data/silica.dat', header = None)
silica_data.columns = ['context']
silica_data
# +
plt.hist(silica_data['context'], bins = 20)
plt.vlines(silica_data['context'].mean(),0,4,color = "red", linewidth = 2, linestyle = "dashed", label = "Mean")
plt.xticks(range(22, 36, 2), [str(num) + '%' for num in list(range(22, 36, 2))])
plt.yticks(range(5), range(5))
plt.xlabel('Silica')
plt.ylabel('Count')
plt.title(f'Silica content')
plt.legend(loc = 'upper left')
plt.show()
# -
silica_data = silica_data['context']
print('Mean: ', silica_data.mean())
print('Standard Deviation: ', silica_data.std())
print('Numpy Standard Deviation: ', np.std(silica_data, ddof = 1))
print('Skewness: ', silica_data.skew())
print('Kurtosis: ', silica_data.kurtosis())
# ### Problem 2. Categorical Variables. Comparing Categories
# In addition to numeric variables (like age and salary), in statistics we also use **categorical variables**. These are descriptions of quality (as opposed to quantity). Such variables can be gender, smoker / non-smoker, results of a medical study (healthy / not healthy), colors (red, green, blue), etc. To plot values of categories, we use *bar charts*. Since category names can be long, it's sometimes useful to plot the lines horizontally.
#
# <p style="color: #d9534f"><strong>There is a very significant difference between histograms and bar charts. Histograms are used to plot the frequency distribution of one numeric variable. Bar charts are used to plot categorical variables - how each value compares to other values.</strong></p>
#
# The dataset `budget.dat` contains the figures for the eight main items in the US budget for 1978 and 1979 in billions
# of dollars.
#
# Display the two budgets separately. Use `xlabel()` (or `ylabel()` if your plot is horizontal) to write the names of each category. You can use [this](https://matplotlib.org/examples/pylab_examples/barchart_demo.html) and [this](https://matplotlib.org/examples/pylab_examples/barchart_demo2.html) examples as a guide.
#
# Create another variable which shows the difference in budget $\Delta b = b_{1979} - b_{1978}$. Add this variable to the dataset (find out how). Plot it. How does the budget differ?
#
# Since the numbers are different, a better comparison will be if we convert them to percentages of the total budget. Create two more variables for 1978 and 1979 and add them to the dataset. Plot these now. Also plot the difference in percentage, like you did before.
budget = pd.read_table('data/budget.dat')
budget
def plot_budget(budget, year_str):
plt.barh(budget['Category'], budget[year_str])
plt.xlim(0, 120)
plt.title(f'US Budget {year_str}')
plt.xlabel('Budget [$10^7$ \$]')
plt.show()
plot_budget(budget, str(1978))
plot_budget(budget, str(1979))
# +
plt.barh(budget['Category'], budget['1979'], label = '1979', alpha = 0.7)
plt.barh(budget['Category'], budget['1978'], label = '1978', alpha = 0.9)
plt.title(f'US Budget 1978 vs. 1979')
plt.xlabel('Budget [$10^7$ \$]')
plt.legend()
plt.show()
# +
plt.barh(budget['Category'], budget['1979'] - budget['1978'])
plt.title(f'US Budget Difference 1978 vs. 1979')
plt.xlabel('Difference [%]')
plt.show()
# -
budget['1978_perc'] = budget['1978'] / budget['1978'].sum() * 100
budget['1979_perc'] = budget['1979'] / budget['1979'].sum() * 100
budget
# +
def plot_budget_perc_bar(budget, year_str):
plt.barh(budget['Category'], budget[year_str + '_perc'])
plt.xticks(range(0, 30, 5), [str(num) + '%' for num in list(range(0, 30, 5))])
plt.title(f'US Budget {year_str}')
plt.xlabel('Percentage data')
plt.show()
plot_budget_perc_bar(budget, str(1978))
plot_budget_perc_bar(budget, str(1979))
# +
def plot_budget_perc_pie(budget, year_str):
plt.pie(labels = budget['Category'], x = budget[year_str + '_perc'], autopct='%.2f%%')
plt.title(f'US Budget')
plt.show()
plot_budget_perc_pie(budget, str(1978))
plot_budget_perc_pie(budget, str(1979))
# +
plt.barh(budget['Category'], budget['1979_perc'] - budget['1978_perc'])
plt.axvline(0, color = 'red')
plt.xlim(-0.7, 0.7)
plt.title(f'US Budget Difference 1978 vs. 1979')
plt.xlabel('Difference [%]')
plt.show()
# +
index = np.arange(8)
bar_width = 0.35
plt.barh(index, budget['1978'], bar_width, color = "red", label = "1978")
plt.barh(index+bar_width, budget['1979'], bar_width, color = "green", label = "1979")
plt.barh(index+bar_width+bar_width, budget['1979'] - budget['1978'], bar_width, color = "skyblue", label = "1978 vs. 1979")
plt.title('US Budget')
plt.xlabel('Budget [$10^7$ \$]')
plt.yticks(index + bar_width / 2, budget['Category'])
plt.legend()
plt.show()
# +
def cm_to_inch(value):
return value/2.54
index = np.arange(8)
bar_width = 0.35
plt.figure(figsize=(cm_to_inch(25),cm_to_inch(15)))
plt.barh(index, budget['1978_perc'], bar_width, color = "red", label = "1978")
plt.barh(index+bar_width, budget['1979_perc'], bar_width, color = "green", label = "1979")
plt.barh(index+bar_width+bar_width, budget['1979_perc'] - budget['1978_perc'], bar_width, color = "skyblue", label = "1978 vs. 1979")
plt.axvline(0, color = '#f5e642')
plt.xlim(-25, 30, 5)
plt.title('US Budget')
plt.xlabel('Budget [$10^7$ \$]')
plt.yticks(index + bar_width / 2, budget['Category'])
plt.legend()
plt.show()
# -
# ### Problem 3. Correlations between Variables. Alcohol and Tobacco Usage
# The dataset `alcohol_tobacco.dat` shows the average weekly household spending, in British pounds, on tobacco products and alcoholic beverages for each of the 11 regions of Great Britain.
#
# Create a scatter plot. Print the correlation coefficient. You can use the **correlation matrix** (find out how).
#
# There's a major outlier. Which one is it?
#
# Remove the outlier from the dataset (find out how). Calculate the correlation coefficient once again. It should be much higher.
#
# This example is useful to show what an outlier is, and how an outlier can influence the results of an experiment.
#
# **Note:** Be careful with outliers. Sometimes they indicate human error (e.g. human height 1588 cm is obviously wrong) but sometimes they indicate important patterns in the data. Should you remove, replace, or leave them is a difficult question and should be answered separately for each dataset.
alcohol_tobacco_usage = pd.read_table('data/alcohol_tobacco.dat')
alcohol_tobacco_usage
alcohol_tobacco_usage.corr() # low correlation coefficient, probably there is outlier
# +
plt.scatter(alcohol_tobacco_usage['Alcohol'], alcohol_tobacco_usage['Tobacco'])
plt.title('Alcohol and Tobaco usage', pad = 20)
plt.xlabel('Alcohol')
plt.ylabel('Tobacco')
plt.show()
# -
alcohol_tobacco_usage[alcohol_tobacco_usage['Alcohol'] < 4.2]
alcohol_tobacco_usage_without_NI = alcohol_tobacco_usage[alcohol_tobacco_usage['Region'] != 'Northern Ireland']
alcohol_tobacco_usage_without_NI
alcohol_tobacco_usage_without_NI.corr()
# +
plt.scatter(alcohol_tobacco_usage_without_NI['Alcohol'], alcohol_tobacco_usage_without_NI['Tobacco'])
plt.title('Alcohol and Tobaco usage', pad = 20)
plt.xlabel('Alcohol')
plt.ylabel('Tobacco')
plt.show()
# -
# ### Problem 4. Simulation
# Another prediction technique based on statistics, is simulation. This means recreating a system's parameters and running the experiment on a computer instead of running it in real life. Simulation can give us many insights. It's useful for prediction, "what-if" analysis, etc. It's also very useful if we have very limited "real experimentation" resources and want to narrow down our possibilities.
#
# Let's see how we can simulate the profit of a grocery shop.
#
# The profit is dependent on the customers and what items they buy. Let's assume that the number of customers per months follows a normal distribution with mean 500 and standard deviation 20.
#
# $$ C \sim N(500, 20) $$
#
# In the shop, there are several items, each having a different popularity. The popularity represents the probability of buying each item.
#
# | Item | Price | Popularity |
# |--------------------|-------|------------|
# | Bread | 0.99 | 0.5 |
# | Milk | 2.89 | 0.15 |
# | Eggs, dozen | 2.00 | 0.2 |
# | Chicken fillet, kg | 6.39 | 0.15 |
#
# Each customer buys *exactly one* article at random. Each customer will generate an expected profit equal to $\text{price} . \text{popularity}$. Total profit: sum of all profits.
def get_customer_profit():
n = np.random.random()
if n <= 0.5:
return 0.99
elif n < 0.65:
return 2.89
elif n <= 0.85:
return 2
else:
return 6.39
days = 1000
def run_simulation():
profits = []
for day in range(days):
customers = np.floor(np.random.normal(500, 20))
profit = sum([get_customer_profit() for c in np.arange(customers)])
profits.append(profit)
return profits
profits = run_simulation()
plt.hist(profits, bins = 50)
plt.xlabel("Profit for " + str(days) + " days [$]")
plt.ylabel("Count")
plt.show()
# Now we can answer questions like:
# * What's the probability of profit less than \$1100?
# * What's the probability of profit between \$1300 and \$1400?
#
# We can also change our model. Let's suppose now that one customer can take 1, 2 or 3 items, with probabilities 0.5, 0.3 and 0.2 respectively. The picked items are independent. How does this change the distribution?
# +
def get_customer_profit_many_items(items = 1):
customer_sum = sum([get_customer_profit() for i in range(items)])
return customer_sum
def get_total_customer_profit():
n = np.random.random()
if n <= 0.5:
return get_customer_profit_many_items(1)
elif n <= 0.8:
return get_customer_profit_many_items(2)
else:
return get_customer_profit_many_items(3)
# -
def run_simulation_many_items():
days = 1000
profits_many_items = []
for day in range(days):
customers = np.floor(np.random.normal(500, 20))
profit = sum([get_total_customer_profit() for c in np.arange(customers)])
profits_many_items.append(profit)
return profits_many_items
profits_many_items = run_simulation_many_items()
plt.hist(profits_many_items, bins = 50)
plt.xlabel("Profit for " + str(days) + " days [$]")
plt.ylabel("Count")
plt.show()
plt.title("Comparison of profits: 1 vs 3 items")
plt.hist(profits, bins = 20)
plt.hist(profits_many_items, bins = 20)
plt.xlabel("Profit")
plt.ylabel("Count")
plt.show()
# ### ** Problem 5. Monte Carlo Simulation
# One common technique to apply simulations is called **Monte Carlo simulation**. It's similar to the simulation from the previous example. The main idea is to use random sampling to solve deterministic problems.
#
# Research what these simulations are. Give examples. Implement at least one case of a Monte Carlo simulation. You can use the following checklist to help with your research and work:
# * What is a simulation?
# * How is simulation used in science?
# * Why is a simulation useful?
# * How are statistics useful in simulation? How can we simulate unknown, random processes?
# * What is a Monte Carlo simulation (also known as "Monte Carlo method")?
# * A common use of Monte Carlo methods is numeric integration
# * Define the problem. Propose the solution. Implement it and test with some common functions
# * How does this method compare to other methods, e.g. the trapezoidal rule? Compare the performance (accuracy and time to execute) of both methods
# * Apply Monte Carlo simulation to a real-life system. There are many examples. You can see [Wikipedia](https://en.wikipedia.org/wiki/Monte_Carlo_method#Applications) or some other resource for inspiration.
# ### ** Problem 6. Probabilistic Data Structures
# A very interesting application of probability in computer science is a kind of data structures which have a probabilistic behaviour. Examples of these are **Bloom filter**, **Skip list**, **Count-min sketch** and **HyperLogLog**.
#
# Research how one of these structures works. Or write about many of them, if you wish. You can use the following checklist as a guide:
# * What is a data structure?
# * What is a probabilistic data structure?
# * Where does the probabilistic behaviour emerge?
# * What advantages do these structures provide?
# * For your chosen structure, how is it constructed?
# * What parts do you need? What are the details?
# * How does the structure work?
# * What operations can you do?
# * What are the typical probabilities associated with these operations?
# * Analyze the structure
# * Analyze the runtimes for all operations
# * Analyze the space usage
# * Compare to a similar, non-probabilistic data structure
# * What advantages does the new data structure have? What drawbacks do you need to be aware of?
# * Give at least one example where this structure is useful
# * E.g. Bloom filter - spell checkers
# * Analyze the use case
# * If possible, implement the use case
# * Display some metrics (e.g. % conserved space, % reduced time)
|
statistics/Statistics Exercise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# +
fc = 1e-5 # cutoff freq between 0 and 0.5
b = 5e-2 # transition band between 0 and 0.5
N = int(np.ceil((4 / b)))
if not N % 2: N += 1 # N has to be odd
n = np.arange(N)
# get sinc functon
sinc = np.sinc(2 * fc * (n - (N - 1) / 2))
# get hamming window
hamming_w = 0.54 - 0.46 * np.cos(2 * np.pi * (n / N))
# apply sinc to window
filt = sinc * hamming_w
filt = filt / np.sum(filt) # normalize
# +
res = 10000
def weierstrass(x, n, res=10000):
y = np.zeros(res)
for n in range(n):
y = y + np.cos(3**n*np.pi*x)/2**n
return y
x = np.linspace(-1, 1, res)
y = np.reshape(weierstrass(x, 500), (res,))
plt.figure(figsize=(10, 5))
plt.plot(x, y)
plt.show()
# -
plt.figure(figsize=(10, 5))
s = np.convolve(y, h, mode="same")
plt.plot(x, s)
plt.show()
|
diaries/ld3-audio-filter-design/hw-3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Getting started with TensorFlow
#
# **Learning Objectives**
# 1. Practice defining and performing basic operations on constant Tensors
# 1. Use Tensorflow's automatic differentiation capability
# 1. Learn how to train a linear regression from scratch with TensorFLow
#
# In this notebook, we will start by reviewing the main operations on Tensors in TensorFlow and understand how to manipulate TensorFlow Variables. We explain how these are compatible with python built-in list and numpy arrays.
#
# Then we will jump to the problem of training a linear regression from scratch with gradient descent. The first order of business will be to understand how to compute the gradients of a function (the loss here) with respect to some of its arguments (the model weights here). The TensorFlow construct allowing us to do that is `tf.GradientTape`, which we will describe.
#
# At last we will create a simple training loop to learn the weights of a 1-dim linear regression using synthetic data generated from a linear model.
#
# As a bonus exercise, we will do the same for data generated from a non linear model, forcing us to manual engineer non-linear features to improve our linear model performance.
import numpy as np
import tensorflow as tf
from matplotlib import pyplot as plt
print(tf.__version__)
# ## Operations on Tensors
# ### Variables and Constants
# Tensors in TensorFlow are either contant (`tf.constant`) or variables (`tf.Variable`).
# Constant values can not be changed, while variables values can be.
#
# The main difference is that instances of `tf.Variable` have methods allowing us to change
# their values while tensors constructed with `tf.constant` don't have these methods, and
# therefore their values can not be changed. When you want to change the value of a `tf.Variable`
# `x` use one of the following method:
#
# * `x.assign(new_value)`
# * `x.assign_add(value_to_be_added)`
# * `x.assign_sub(value_to_be_subtracted`
#
#
x = tf.constant([2, 3, 4])
x
x = tf.Variable(2.0, dtype=tf.float32, name="my_variable")
x.assign(45.8) # TODO 1
x
x.assign_add(4) # TODO 2
x
x.assign_sub(3) # TODO 3
x
# ### Point-wise operations
# Tensorflow offers similar point-wise tensor operations as numpy does:
#
# * `tf.add` allows to add the components of a tensor
# * `tf.multiply` allows us to multiply the components of a tensor
# * `tf.subtract` allow us to substract the components of a tensor
# * `tf.math.*` contains the usual math operations to be applied on the components of a tensor
# * and many more...
#
# Most of the standard aritmetic operations (`tf.add`, `tf.substrac`, etc.) are overloaded by the usual corresponding arithmetic symbols (`+`, `-`, etc.)
# +
a = tf.constant([5, 3, 8]) # TODO 1
b = tf.constant([3, -1, 2])
c = tf.add(a, b)
d = a + b
print("c:", c)
print("d:", d)
# +
a = tf.constant([5, 3, 8]) # TODO 2
b = tf.constant([3, -1, 2])
c = tf.multiply(a, b)
d = a * b
print("c:", c)
print("d:", d)
# +
# tf.math.exp expects floats so we need to explicitly give the type
a = tf.constant([5, 3, 8], dtype=tf.float32)
b = tf.math.exp(a)
print("b:", b)
# -
# ### NumPy Interoperability
#
# In addition to native TF tensors, tensorflow operations can take native python types and NumPy arrays as operands.
# native python list
a_py = [1, 2]
b_py = [3, 4]
tf.add(a_py, b_py) # TODO 1
# numpy arrays
a_np = np.array([1, 2])
b_np = np.array([3, 4])
tf.add(a_np, b_np) # TODO 2
# native TF tensor
a_tf = tf.constant([1, 2])
b_tf = tf.constant([3, 4])
tf.add(a_tf, b_tf) # TODO 3
# You can convert a native TF tensor to a NumPy array using .numpy()
a_tf.numpy()
# ## Linear Regression
#
# Now let's use low level tensorflow operations to implement linear regression.
#
# Later in the course you'll see abstracted ways to do this using high level TensorFlow.
# ### Toy Dataset
#
# We'll model the following function:
#
# \begin{equation}
# y= 2x + 10
# \end{equation}
# +
X = tf.constant(range(10), dtype=tf.float32)
Y = 2 * X + 10
print(f"X:{X}")
print(f"Y:{Y}")
# -
# Let's also create a test dataset to evaluate our models:
# +
X_test = tf.constant(range(10, 20), dtype=tf.float32)
Y_test = 2 * X_test + 10
print(f"X_test:{X_test}")
print(f"Y_test:{Y_test}")
# -
# #### Loss Function
# The simplest model we can build is a model that for each value of x returns the sample mean of the training set:
# +
y_mean = Y.numpy().mean()
def predict_mean(X):
y_hat = [y_mean] * len(X)
return y_hat
Y_hat = predict_mean(X_test)
# -
# Using mean squared error, our loss is:
# \begin{equation}
# MSE = \frac{1}{m}\sum_{i=1}^{m}(\hat{Y}_i-Y_i)^2
# \end{equation}
# For this simple model the loss is then:
errors = (Y_hat - Y) ** 2
loss = tf.reduce_mean(errors)
loss.numpy()
# This values for the MSE loss above will give us a baseline to compare how a more complex model is doing.
# Now, if $\hat{Y}$ represents the vector containing our model's predictions when we use a linear regression model
# \begin{equation}
# \hat{Y} = w_0X + w_1
# \end{equation}
#
# we can write a loss function taking as arguments the coefficients of the model:
def loss_mse(X, Y, w0, w1):
Y_hat = w0 * X + w1
errors = (Y_hat - Y) ** 2
return tf.reduce_mean(errors)
# ### Gradient Function
#
# To use gradient descent we need to take the partial derivatives of the loss function with respect to each of the weights. We could manually compute the derivatives, but with Tensorflow's automatic differentiation capabilities we don't have to!
#
# During gradient descent we think of the loss as a function of the parameters $w_0$ and $w_1$. Thus, we want to compute the partial derivative with respect to these variables.
#
# For that we need to wrap our loss computation within the context of `tf.GradientTape` instance which will reccord gradient information:
#
# ```python
# with tf.GradientTape() as tape:
# loss = # computation
# ```
#
# This will allow us to later compute the gradients of any tensor computed within the `tf.GradientTape` context with respect to instances of `tf.Variable`:
#
# ```python
# gradients = tape.gradient(loss, [w0, w1])
# ```
# We illustrate this procedure with by computing the loss gradients with respect to the model weights:
# TODO 1
def compute_gradients(X, Y, w0, w1):
with tf.GradientTape() as tape:
loss = loss_mse(X, Y, w0, w1)
return tape.gradient(loss, [w0, w1])
# +
w0 = tf.Variable(0.0)
w1 = tf.Variable(0.0)
dw0, dw1 = compute_gradients(X, Y, w0, w1)
# -
print("dw0:", dw0.numpy())
print("dw1", dw1.numpy())
# ### Training Loop
#
# Here we have a very simple training loop that converges. Note we are ignoring best practices like batching, creating a separate test set, and random weight initialization for the sake of simplicity.
# +
STEPS = 1000
LEARNING_RATE = 0.02
MSG = "STEP {step} - loss: {loss}, w0: {w0}, w1: {w1}\n"
w0 = tf.Variable(0.0)
w1 = tf.Variable(0.0)
for step in range(0, STEPS + 1):
dw0, dw1 = compute_gradients(X, Y, w0, w1)
w0.assign_sub(dw0 * LEARNING_RATE)
w1.assign_sub(dw1 * LEARNING_RATE)
if step % 100 == 0:
loss = loss_mse(X, Y, w0, w1)
print(MSG.format(step=step, loss=loss, w0=w0.numpy(), w1=w1.numpy()))
# -
# Now let's compare the test loss for this linear regression to the test loss from the baseline model that outputs always the mean of the training set:
loss = loss_mse(X_test, Y_test, w0, w1)
loss.numpy()
# This is indeed much better!
# ## Bonus
# Try modelling a non-linear function such as: $y=xe^{-x^2}$
X = tf.constant(np.linspace(0, 2, 1000), dtype=tf.float32)
Y = X * tf.exp(-(X**2))
# +
# %matplotlib inline
plt.plot(X, Y)
# -
def make_features(X):
f1 = tf.ones_like(X) # Bias.
f2 = X
f3 = tf.square(X)
f4 = tf.sqrt(X)
f5 = tf.exp(X)
return tf.stack([f1, f2, f3, f4, f5], axis=1)
def predict(X, W):
return tf.squeeze(X @ W, -1)
def loss_mse(X, Y, W):
Y_hat = predict(X, W)
errors = (Y_hat - Y) ** 2
return tf.reduce_mean(errors)
def compute_gradients(X, Y, W):
with tf.GradientTape() as tape:
loss = loss_mse(Xf, Y, W)
return tape.gradient(loss, W)
# +
STEPS = 2000
LEARNING_RATE = 0.02
Xf = make_features(X)
n_weights = Xf.shape[1]
W = tf.Variable(np.zeros((n_weights, 1)), dtype=tf.float32)
# For plotting
steps, losses = [], []
plt.figure()
for step in range(1, STEPS + 1):
dW = compute_gradients(X, Y, W)
W.assign_sub(dW * LEARNING_RATE)
if step % 100 == 0:
loss = loss_mse(Xf, Y, W)
steps.append(step)
losses.append(loss)
plt.clf()
plt.plot(steps, losses)
print(f"STEP: {STEPS} MSE: {loss_mse(Xf, Y, W)}")
plt.figure()
plt.plot(X, Y, label="actual")
plt.plot(X, predict(Xf, W), label="predicted")
plt.legend()
# -
# Copyright 2021 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
|
notebooks/introduction_to_tensorflow/solutions/1_core_tensorflow.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.9 64-bit (''base'': conda)'
# name: python379jvsc74a57bd077308f10e778eec9a13a21206849a589a8c13410aaae2405e5051cd249c67e86
# ---
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import os
from astropy.table import QTable
import astropy.units as u
from astropy.coordinates import SkyCoord, match_coordinates_sky
p1 = os.getcwd() + '/Catalogues/'
tab = QTable.read(p1 + 'HSC_I_COSMOS_1_model_psf_cat.fits')
#tab.columns
# Plotting
fig = plt.figure(figsize = (11,7))
ax = fig.add_subplot(111)
ax.set_xlabel(r'PSF Magnitude')
ax.set_ylabel(r"Mag aper $1''$-Mag aper $3''$")
ax.plot(tab['MAG_PSF'], tab['MAG_APER'][:,-1]-tab['MAG_APER'][:,4], '.', alpha=0.2)
ax.set_ylim([-0.25, 2])
ax.set_xlim([16,25])
plt.tight_layout()
tab2 = QTable.read(p1 + 'HSC_I_COSMOS_3_model_psf_cat.fits')
print(tab['MAG_APER'].shape)
print(tab2['MAG_APER'].shape)
# +
rad = 1.*u.arcsec
pos1 = SkyCoord(ra=tab['ALPHA_J2000'], dec=tab['DELTA_J2000'], unit='deg')
pos2 = SkyCoord(ra=tab2['ALPHA_J2000'], dec=tab2['DELTA_J2000'], unit='deg')
idx, d2d, _ = match_coordinates_sky(pos2, pos1, nthneighbor=1)
t1_match = tab[idx[d2d<rad]]
t2_match = tab2[d2d<rad]
# -
mag_diff = t1_match['MAG_PSF'] - t2_match['MAG_PSF']
print(np.mean(mag_diff))
# +
mask = np.where((t1_match['MAG_PSF'].value>18.5)&(t1_match['MAG_PSF'].value<21.5)&(t1_match['MAG_APER'][:,1].value-t1_match['MAG_APER'][:,4].value<0.55))[0]
print(np.mean(mag_diff[mask]))
# -
plt.hist(mag_diff[mask].value)
|
Extended_photo/p1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python для анализа данных
#
# ## Веб-скрэйпинг: переход по ссылкам, скачивание файлов
#
# *Автор: <NAME>, НИУ ВШЭ*
# На прошлом занятии мы с вами работали с табличными данными. Второй очень частой задачей для скрэйпинга является автоматический переход по ссылкам. Обычно мы встречаемся с двумя сценариями: переход по нумерованным страницам (обычно это выдача поиска или некоторый упорядоченный архив документов) и переход по определенным ссылкам на странице. Сегодня будем учиться делать и то, и то.
#
# Давайте начнем с учебного примера.
# # Задача
#
# По ссылке http://py4e-data.dr-chuck.net/known_by_Lilian.html список людей, которых знает Лилиан. Нужно найти ее 18-го друга (считаем с одного) и перейти по ссылке (там будет список людей, которых знает этот друг). Какой имя вы извлечете, если эту операцию повторить 7 раз? То есть нам нужно найти 18-го друга 6-го человека.
#
# Прежде всего изучите исходный код страницы. В каком теге лежат ссылки?
# +
# решение
import requests
from bs4 import BeautifulSoup
friends = requests.get('http://py4e-data.dr-chuck.net/known_by_Lilian.html').text
soup = BeautifulSoup(friends, 'lxml')
print(soup.find_all('a')[17]) # ссылки лежат в тэге 'a', находим 18-го друга
# -
# Обратите внимание, что нужная нам информация лежит в атрибуте href, достать текст, как мы делали раньше не поможет.
print(soup.find_all('a')[17].get('href')) # с помощью метода get достаем информацию из атрибут href
# Теперь осталось упаковать все в цикл.
# +
friends = requests.get('http://py4e-data.dr-chuck.net/known_by_Lilian.html').text
soup = BeautifulSoup(friends, 'lxml')
for ix in range(6):
link = soup.find_all('a')[17].get('href')
print(link)
soup = BeautifulSoup(requests.get(link).text, 'lxml')
print('Ответ: '+soup.find_all('a')[17].text)
|
lect7/BS_Links.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# KNN implementation in python
#
# Euclidean space is the length of a line segment between the two points. It can be calculated from the Cartesian coordinates of the points using the Pythagorean theorem
# +
import math
from collections import Counter
class KNN():
def __init__(self, k:int):
self.k = k
self.X = None
self.y = None
def fit(self, X, y):
self.X = X
self.y = y
def predit(self, X):
# store eucleandian distance to each of the training sample and its label
distance_label = []
for row, label in zip(self.X, self.y):
distance_label.append((self.euclidean_dist(row, X), label))
distance_label.sort()
labels = [label for _, label in distance_label][:self.k]
return Counter(labels).most_common(1)[0][0] # return the most frequent label
# return sum(labels)/self.k
def euclidean_dist(self, row1, row2):
res = 0
for x1, x2 in zip(row1, row2): # loop through each feature value for these two samples
res += (x1 - x2)**2
return math.sqrt(res)
# -
X = [[1,3,4,5,2], [2,3,4,3,2], [1,3,6,5,2], [6,3,9,5,2]]
y = [1,1,1,0]
m = KNN(5)
m.fit(X, y)
m.predit([1,1,1,1,1])
|
KNN implementation in python.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
import pickle
import plotly.express as px
import numpy as np
import os
# +
# load location index
with open("data/akl_loc_idx.pkl", 'rb') as f:
loc_idx = pickle.load(f) # datazone to point index
idx_loc = {v:k for k, v in loc_idx.items()} # point index to datazone
print(f" -- loaded location index with dimension {len(loc_idx)}")
# load time index
with open("data/akl_t_idx.pkl", 'rb') as f:
t_idx = pickle.load(f)
print(f" -- loaded time index with dimension {len(t_idx)}")
# load precomputed odt
with open("data/akl_odt.npy", 'rb') as f:
odt = np.load(f)
print(f" -- loaded odt cube with dimensions {odt.shape}")
# -
with open("data/akl_polygons.geojson", 'r') as f:
polys = json.load(f)
# add id field to each feature (requried by plotly even though it's supposed to work via subproperty ref.)
for f in polys["features"]:
f["id"] = f["properties"]["DZ2018"]
# +
# no longer necessary, we're keeping all properties
# # remove properties DZ2018 as it is now duplicated
# for f in polys["features"]:
# f["properties"] = f["properties"].pop("DZ2018", None)
# -
# combine polygons with duplicate feature ids
features = {}
for f in polys["features"]:
fid = f["id"]
if fid not in features:
features[fid] = f
else:
f_poly = f["geometry"]["coordinates"]
features[fid]["geometry"]["coordinates"] += f_poly
#print(fid, len(features[fid]["geometry"]["coordinates"]))
polys["features"] = list(features.values())
print(f"total features: {len(polys['features'])}")
# filter polygons by those that have population centroids
pids = [k for k, v in loc_idx.items()]
valid_features = [f for f in polys["features"] if f["id"] in pids]
polys["features"] = valid_features
print("total features:", len(polys["features"]))
# +
# this was an attempt to reduce the number of polygons displayed by filtering out those that have on valid joureny data
# it is no longer necessary becaue deck.gl is performant
filter_valid = False
if filter_valid:
# filter polygons by those that have valid journeys
jsum = np.sum(np.nansum(odt, axis=2), axis=0)
valid_idx = [i for i, v in enumerate(jsum) if v > 0]
valid_features = [f for f in polys["features"] if loc_idx[f["id"]] in valid_idx]
polys["features"] = valid_features
print(f"total features: {len(polys['features'])}")
# -
# write cube indices to data folder used by the web frontend
with open("data/../frontend/akl/akl_loc_idx.json", "w") as f:
json.dump(loc_idx, f)
with open("data/../frontend/akl/akl_idx_loc.json", "w") as f:
json.dump(idx_loc, f)
# write time index
#idx_t = {v:int(k.timestamp())*1000 for k,v in t_idx.items()} # milliseconds
idx_t = {v:str(k) for k,v in t_idx.items()}
with open("data/../frontend/akl/akl_idx_t.json", "w") as f:
json.dump(idx_t, f)
# write the cleaned polygon data to the frontend folder
# note the .json extension to avoid an additional webpack rule
with open("data/../frontend/akl/akl_polygons_id.json", "w") as f:
json.dump(polys, f)
# replace nan values in odt
nan_value = -1
odt_nan = np.nan_to_num(odt, nan=nan_value)
fmt = np.vectorize(lambda x: f"{x}")
# +
# save the outbound cube slices to the backend data dir
target_dir = "data/../backend/outbound/akl"
if not os.path.exists(target_dir):
os.makedirs(target_dir)
# save a separate dt_slice file for each location
for f in polys["features"]:
# origin id and odt index of origin
o = f["id"]
o_idx = loc_idx[o]
# get dt slice for the origin
dt_slice = odt_nan[o_idx, :, :]
dt_slice = fmt(dt_slice).astype(np.float).tolist()
# save the slice
path = os.path.join(target_dir, f"{o}.json")
with open(path, "w") as f:
json.dump(dt_slice, f)
# +
# save the inbound cube slices to the backend data dir
target_dir = "data/../backend/inbound/akl"
if not os.path.exists(target_dir):
os.makedirs(target_dir)
# save a separate dt_slice file for each location
for f in polys["features"]:
# destination id and odt index of destination
d = f["id"]
d_idx = loc_idx[d]
# get ot slice for the destination
ot_slice = odt_nan[:, d_idx, :]
ot_slice = fmt(ot_slice).astype(np.float).tolist()
# save the slice
path = os.path.join(target_dir, f"{d}.json")
with open(path, "w") as f:
json.dump(ot_slice, f)
# -
# data needs to be wgs84
locations = [f["id"] for f in polys["features"]]
fig = px.choropleth_mapbox(
geojson=polys,
featureidkey="id",
locations=locations,
center = {"lat": -36.8485, "lon": 174.7633},
mapbox_style="carto-positron",
zoom=12)
fig.update_layout(margin={"r":0,"t":0,"l":0,"b":0})
fig.show()
|
notebooks/preprocess-data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# default_exp Core
# -
# !which python
#hide
# %load_ext autoreload
# %autoreload 2
# # Core module
#
# > API details
# ### 1. parameters
from SEQLinkage.Main import *
# args = Args().parser.parse_args(['--fam','../sample_i/rare_positions/sample_i_coding.hg38_multianno.fam',
# '--vcf', '../sample_i/rare_positions/sample_i_coding.hg38_multianno.vcf.gz',
# '--blueprint','./data/vipgenemap.hg38.txt','-f','MERLIN',
# '--tempdir','./Tempdir_s1',
# '--build', 'hg38', '--freq', 'AF', '-K', '0.001', '--moi', 'AD', '-W', '0', '-M', '1',
# '--theta-max', '0.5', '--theta-inc', '0.05','--run-linkage', '--output', './testseqlink'])
#
# args = Args().parser.parse_args(['--fam','../sample_i/rare_positions/sample_i_coding.hg38_multianno.fam',
# '--vcf', '../sample_i/vcf/small_sample_i.vcf.gz',
# '--blueprint','./data/vipgenemap.hg38.txt','-f','MERLIN',
# '--tempdir','./Tempdir_s1',
# '--build', 'hg38', '--freq', 'AF', '-K', '0.001', '--moi', 'AD', '-W', '0', '-M', '1',
# '--theta-max', '0.5', '--theta-inc', '0.05','--run-linkage', '--output', './testseqlink'])
#
# args = Args().parser.parse_args(['--fam','./seqlinkage-example/seqlinkage-example.fam','--vcf','./seqlinkage-example/seqlinkage-example.vcf.gz','-f','MERLIN',
# '--tempdir','./seqlinkage-example/tmprst',
# '--build','hg19','--blueprint','./seqlinkage-example/twogenomap.txt','--freq','EVSEAAF','-K','0.001','--moi','AR','-W','0','-M','1',
# '--theta-max','0.5','--theta-inc','0.05','--run-linkage','--output','./seqlinkage-example/tsq20211130'])
# # ?shoud we set mle parameter as true?
# args = Args().parser.parse_args(['--fam','../MWE/sample2_uniq.fam',
# '--vcf', '../MWE/sample_ii_coding.hg38_multianno.vcf.gz',
# '--blueprint','../MWE/genemap.hg38.txt', '--chrom-prefix','1','-f','MERLIN',
# '--tempdir','./Tempdir',
# '--build', 'hg38', '--freq', 'AF', '-K', '0.001', '--moi', 'AD', '-W', '0', '-M', '1',
# '--theta-max', '0.5', '--theta-inc', '0.05','--run-linkage', '--output', './testseqlink1105'])
#
# args = Args().parser.parse_args('--bin 1 --fam ../seqlinkage-example/seqlinkage-example.fam --vcf ../seqlinkage-example/seqlinkage-example.vcf.gz -f MERLIN --blueprint ../data/genemap.txt --freq EVSEAAF'.split())
args = Args().parser.parse_args('--bin 1 --fam ../data/new_trim_ped_famless17_no:xx.fam --vcf ../data/first1000snp_full_samples.vcf.gz --anno ../data/first1000_chr1_multianno.csv --pop ../data/full_sample_fam_pop.txt -f MERLIN --blueprint ../data/genemap.hg38.txt --freq AF'.split())
# args = Args().parser.parse_args('--fam ../data/new_trim_ped_fam.fam --vcf ../data/first1000snp_full_samples.vcf.gz -f MERLIN --blueprint ../data/genemap.hg38.txt --freq AF'.split())
args = Args().parser.parse_args('--bin 1 --fam ../data/new_trim_ped_famless17_no:xx.fam --vcf /mnt/mfs/statgen/alzheimers-family/linkage_files/geno/full_sample/vcf/full_sample.vcf.gz --anno ../MWE/annotation/EFIGA_NIALOAD_chr22.hg38.hg38_multianno.csv --pop ../data/full_sample_fam_pop.txt -f MERLIN MEGA2 PLINK LINKAGE --build hg38 --freq AF -o data/mwechr19data -K 0.001 --moi AD -W 0 -M 1 --run-linkage -j 8'.split())
args
# ### 2.from Core import deepcopy
#export
from __future__ import print_function
from SEQLinkage.Utils import *
from SEQLinkage.Runner import *
from multiprocessing import Process, Queue
from collections import OrderedDict
import itertools
from copy import deepcopy
import sys, faulthandler, platform
import numpy as np
import pandas as pd
import pickle
import os
import time
from concurrent.futures import ProcessPoolExecutor, ThreadPoolExecutor
from itertools import repeat
if sys.version_info.major == 2:
from cstatgen import cstatgen_py2 as cstatgen
from cstatgen.egglib import Align
else:
from cstatgen import cstatgen_py3 as cstatgen
import egglib
from egglib import Align
# ### 3. RData class
#export
class RData(dict):
def __init__(self, vcf, tfam,anno_file=None,fam_pop_file=None,ind_sample_file=None,allele_freq_info = None):
# tfam.samples: a dict of {sid:[fid, pid, mid, sex, trait], ...}
# tfam.families: a dict of {fid:[s1, s2 ...], ...}
self.tfam = TFAMParser(tfam)
# name of allele frequency meta info
self.af_info = allele_freq_info
self.vs = self.load_vcf(vcf)
self.fam_pop = self.load_fam_info(fam_pop_file)
self.anno = self.load_anno(anno_file)
self.samples_vcf = self.vs.GetSampleNames()
self.samples_not_vcf = checkSamples(self.samples_vcf, self.tfam.samples.keys())[1]
# samples have to be in both vcf and tfam data
self.samples = OrderedDict([(k, self.tfam.samples[k]) for k in self.samples_vcf if k in self.tfam.samples])
# a dict of {fid:[member names], ...}
self.families = {k : [x for x in self.samples if x in self.tfam.families[k]] for k in self.tfam.families}
# a dict of {fid:[idx ...], ...}
self.famsampidx = {}
# a dict of {fid:[maf1, maf2 ...]}
self.maf = OrderedDict()
# finalized sub_regions that are compliant to all families
self.complied_markers = []
# finalized sub regions (variants)
self.combined_regions = []
self.coordinates_by_region = []
# RV varnames by family
self.varnames_by_fam = {}
self.patterns={}
self.gnomAD_estimate={'AFR':(1-0.4589)/(2*7652),'AMR':(1-0.4455)/(2*16791),'ASJ':(1-0.2357)/(2*4925),'EAS':(1-0.4735)/(2*8624),'FIN':(1-0.3048)/(2*11150),'NFE':(1-0.5729)/(2*55860),'OTH':(1-0.4386)/(2*2743),'SAS':(1-0.5624)/(2*15391)}
# reorder family samples based on order of VCF file
for k in list(self.families.keys()):
if len(self.families[k]) == 0:
# skip families having no samples in VCF file
del self.families[k]
else:
self.famsampidx[k] = [i for i, x in enumerate(self.samples_vcf) if x in self.families[k]]
# a dict of {fid:[idx ...], ...}
self.famvaridx = {}
self.famvarmafs = {}
self.wtvar = {}
self.freq_by_fam = {}
self.include_vars = []
self.total_varnames={}
self.total_mafs={}
self.wt_maf={}
self.freq = []
self.genotype_all={}
self.mle_mafs={}
self.missing_persons=[]
self.reset()
def load_vcf(self,vcf):
# load VCF file header
return cstatgen.VCFstream(vcf)
def load_anno(self,anno_file):
if anno_file is None:
return None
anno = pd.read_csv(anno_file)
anno.index = list(anno.Otherinfo1)
anno = anno[~anno.index.duplicated()]
tmp = anno[list(set(self.fam_pop.values()))]
tmp = tmp.replace('.',np.nan) #Fixme: missing mafs
tmp = tmp.replace(0,np.nan)
anno = pd.concat([anno[['Chr','Start']],tmp.astype(np.float64)],axis=1)
print('anno',anno.shape)
return anno
def load_fam_info(self,fam_pop_file):
if fam_pop_file is None:
return None
fam_pop = {}
with open(fam_pop_file) as f:
for line in f:
key, value = line.split()
if value == 'NA': #Fixme: deal with missing info
fam_pop[key]=self.af_info
else:
fam_pop[key] = value
return fam_pop
def load_ind_samples(self,ind_sample_file):
pass
def get_regions(self,step=1000):
'''separate chromosome to regions'''
regions=[]
chrom=self.anno.Chr.unique()[0]
for i,s in enumerate(self.anno.Start):
if i==0:
pre=None
cur=s
elif i%step==0:
pre=cur
cur=s
regions.append([str(chrom),str(pre),str(cur),'R'+str(pre)+'_'+str(cur),'.', '.', '.'])
if cur!=s:
pre=cur
cur=s
regions.append([str(chrom),str(pre),str(cur),'R'+str(pre)+'_'+str(cur),'.', '.', '.'])
return regions
def reset(self):
for item in self.tfam.samples: #for all samples in fam ( with or without vcfs)
self[item] = []
self.genotype_all[item] = []
self.variants = []
self.include_vars = []
self.total_varnames={}
self.total_mafs={}
self.chrom = None
for k in self.families.keys():
self.famvaridx[k] = []
self.famvarmafs[k] = []
self.maf = OrderedDict()
# superMarkerCount is the max num. of recombinant fragments among all fams
self.superMarkerCount = 0
self.complied_markers = []
self.combined_regions = []
self.coordinates_by_region = []
self.patterns={}
self.missing_persons=[]
self.gss = {} #test line
def getMidPosition(self):
if len(self.variants) == 0:
return None
return sum([x[1] for x in self.variants]) / len(self.variants)
def getFamVariants(self, fam, style = None):
if style is None:
return [item for idx, item in enumerate(self.variants) if idx in self.famvaridx[fam]]
elif style == "map":
names = []
pos = []
for idx in self.famvaridx[fam]:
names.append(self.variants[idx][0])
pos.append(self.variants[idx][1])
mafs = self.famvarmafs[fam]
return np.array(names), pos, np.array(mafs) #pos can't be array -> TypeError: in method 'HaplotypingEngine_Execute'
else:
raise ValueError("Unknown style '{}'".format(style))
def getFamSamples(self, fam):
nvar = len([item for idx, item in enumerate(self.variants) if idx in self.famvaridx[fam]])
output = [[]] * len(self.tfam.families[fam])
for idx, item in enumerate(self.tfam.sort_family(fam)):
# sample info, first 5 columns of ped
output[idx] = self.tfam.samples[item][:-1]
# sample genotypes
if item in self.samples:
output[idx].extend(self[item])
else:
output[idx].extend(["00"] * nvar)
return output
# ### 4.RegionExtractor class
class RegionExtractor:
'''Extract given genomic region from VCF
converting genotypes into dictionary of
genotype list'''
def __init__(self, filename, build = env.build, chr_prefix = None, allele_freq_info = None, include_vars_file=None):
self.vcf = cstatgen.VCFstream(filename)
self.chrom = self.startpos = self.endpos = self.name = None
self.chr_prefix = chr_prefix
# name of allele frequency meta info
self.af_info = allele_freq_info
self.xchecker = PseudoAutoRegion('X', build)
self.ychecker = PseudoAutoRegion('Y', build)
self.include_vars_file = include_vars_file
def apply(self, data):
# Clean up
data.reset()
data.chrom = self.chrom
self.vcf.Extract(self.chrom, self.startpos, self.endpos)
varIdx = 0
# for each variant site
while (self.vcf.Next()):
# skip tri-allelic sites
if not self.vcf.IsBiAllelic():
with env.triallelic_counter.get_lock():
env.triallelic_counter.value += 1
continue
if len(data.variants) > 0:
if self.vcf.GetPosition()==data.variants[-1][1]:
continue
# check if the line's sample number matches the entire VCF sample number
if not self.vcf.CountSampleGenotypes() == self.vcf.sampleCount:
raise ValueError('Genotype and sample mismatch for region {}: {:,d} vs {:,d}'.\
format(self.name, self.vcf.CountSampleGenotypes(), self.vcf.sampleCount))
# valid line found, get variant info
try:
if type(self.af_info) is list:
maf = []
large_maf = []
for pop_info in self.af_info:
large_maf.append(False)
try:
maf.append(float(self.vcf.GetInfo(pop_info)))
except ValueError:
maf.append(0.0)
for idx in range(len(maf)):
if maf[idx] > 0.5:
large_maf[idx]=True
maf[idx] = 1-maf[idx]
else:
large_maf=False
try:
maf = float(self.vcf.GetInfo(self.af_info)) if self.af_info else None
except ValueError:
maf = 0.0
if maf > 0.5:
large_maf=True
maf = 1 - maf
except Exception:
raise ValueError("VCF line {}:{} does not have valid allele frequency field {}!".\
format(self.vcf.GetChrom(), self.vcf.GetPosition(), self.af_info))
data.variants.append([self.vcf.GetChrom(), self.vcf.GetPosition(), self.name, maf])
# for each family assign member genotype if the site is non-trivial to the family
for k in data.families:
gs = self.vcf.GetGenotypes(data.famsampidx[k])
if len(data.freq_by_fam) > 0:
popidx=self.af_info.index(data.freq_by_fam[k])
if large_maf[popidx]:
tmpgs=[]
for tmpg in gs:
if tmpg=='00':
tmpgs.append(tmpg)
else:
tmpgs.append(''.join([str(3-int(tg)) for tg in tmpg]))
gs=tuple(tmpgs)
else:
if large_maf:
tmpgs=[]
for tmpg in gs:
if tmpg=='00':
tmpgs.append(tmpg)
else:
tmpgs.append(''.join([str(3-int(tg)) for tg in tmpg]))
gs=tuple(tmpgs)
for person, g in zip(data.families[k], gs):
data.genotype_all[person].append(g)
if len(set(''.join(gs))) <= 1:
# skip monomorphic gs
continue
else:
if len(set(''.join([x for x in gs if x != "00"]))) <= 1:
data.wtvar[k].append(varIdx)
# this variant is found in the family
data.famvaridx[k].append(varIdx)
for person, g in zip(data.families[k], gs):
data[person].append(g)
varIdx += 1
#
if varIdx == 0:
return 1
else:
if not self.include_vars_file is None:
with open(self.include_vars_file) as invar_fh:
for invar_line in invar_fh:
chrom, pos = invar_line.split()
for vidx,v in enumerate(data.variants):
if v[0] == chrom and v[1] == int(pos):
data.include_vars.append("{}".format(pos))
break
else:
data.include_vars = ["{}".format(item[1]) for item in data.variants]
with env.variants_counter.get_lock():
env.variants_counter.value += varIdx
return 0
def getRegion(self, region):
self.chrom, self.startpos, self.endpos, self.name = region[:4]
self.startpos = int(self.startpos)
self.endpos = int(self.endpos) + 1
if self.chrom in ['X','23']:
if self.xchecker.check(self.startpos) or self.xchecker.check(self.endpos):
self.chrom = 'XY'
if self.chrom in ['Y','24']:
if self.ychecker.check(self.startpos) or self.ychecker.check(self.endpos):
self.chrom = 'XY'
if self.chr_prefix and not self.chrom.startswith(self.chr_prefix):
self.chrom = self.chr_prefix + self.chrom
# ### 5.MarkerMaker class
class MarkerMaker:
def __init__(self, wsize, maf_cutoff = None,single_markers=False,recomb_max = 1,af_info=None,freq_by_fam=False,rsq=0.0,mle=False,rvhaplo=False,recomb_perfam=True):
self.missings = ("0", "0")
self.gtconv = {'1':0, '2':1}
self.recomb_max = recomb_max
self.haplotyper = cstatgen.HaplotypingEngine(verbose = env.debug)
self.af_info = af_info
self.freq_by_fam = freq_by_fam
self.rsq=rsq
self.mle=mle #use MLE estimate from families for MAF
self.count= not mle #count founder alleles to estimate MAF
self.rvhaplo=rvhaplo
self.recomb_perfam=recomb_perfam
if wsize == 0 or wsize >= 1:
self.r2 = None
else:
self.r2 = wsize
self.coder = cstatgen.HaplotypeCoder(wsize)
self.maf_cutoff = maf_cutoff
self.single_markers = single_markers
self.name = None
def apply(self, data):
# temp raw haplotype, maf and variant names data
haplotypes = OrderedDict()
mafs = {} ##Per fam per variant
uniq_vars = []
exclude_vars = []
varnames = {}
recombPos = {}
#try:
# haplotyping plus collect found allele counts
# and computer founder MAFS
self.__Haplotype(data, haplotypes, mafs, varnames,recombPos,uniq_vars,exclude_vars)
print('__Haplotype',haplotypes, mafs, varnames,recombPos,uniq_vars,exclude_vars)
self.haplotypes, self.mafs, self.varnames = haplotypes, mafs, varnames ###anno
if len(varnames):
if not any ([len(varnames[x]) - 1 for x in varnames]):
# all families have only one variant
self.__AssignSNVHaplotypes(data, haplotypes, mafs, varnames)
else:
# calculate LD clusters using founder haplotypes
clusters = self.__ClusterByLD(data, haplotypes, varnames)
# recoding the genotype of the region
self.__CodeHaplotypes(data, haplotypes, mafs, varnames, clusters)
#except Exception as e:
# if env.debug:
# raise
# return -1
self.__FormatHaplotypes(data,recombPos,varnames,uniq_vars)
return 0
def __getMLEfreq(self,data, markers_to_analyze, pos_all, families, rsq, output_log):
output_sample=[]
mle_mafs={}
if len(markers_to_analyze)==0:
return mle_mafs
for fam in families:
for person in data.tfam.sort_family(fam):
output_sample.append([])
last_ele=len(output_sample)-1
output_sample[last_ele] = data.tfam.samples[person][:-1]
if person in data.samples:
for marker in markers_to_analyze:
idx=int(marker.split('-')[0][1:])
output_sample[last_ele].append(data.genotype_all[person][idx])
else:
output_sample[last_ele].extend(["00"] * len(markers_to_analyze))
with stdoutRedirect(to = output_log):
af = self.haplotyper.Execute(data.chrom, markers_to_analyze, pos_all, output_sample, rsq, output_log,False)
with open(output_log) as mle_fh:
for line in mle_fh:
if line.startswith('V'):
tmp_eles = line.split(':')
if tmp_eles[0] not in mle_mafs:
freqs=tmp_eles[1].split()
mle_maf = float(freqs[1])
if mle_maf>0.5:
mle_mafs[tmp_eles[0]]=float("%.9f"%(1-mle_maf))
else:
#alt allele is more frequent
mle_mafs[tmp_eles[0]]=float("%.9f"%mle_maf)
marker_idx=int(tmp_eles[0].split('-')[0][1:])
for fam in families:
if marker_idx not in data.famvaridx[fam]:
continue
tmp_famvaridx=data.famvaridx[fam].index(marker_idx)
for person in data.families[fam]:
tmpg=data.genotype_all[person][marker_idx]
tmpg_switch=''.join([str(3-int(tg)) for tg in tmpg]) if tmpg!='00' else tmpg
data.genotype_all[person][marker_idx]=tmpg_switch
tmpg2=data[person][tmp_famvaridx]
tmpg_switch2=''.join([str(3-int(tg)) for tg in tmpg2]) if tmpg2!='00' else tmpg2
data[person][tmp_famvaridx]=tmpg_switch2
return mle_mafs
def __computefounderfreq(self,data, families):
#count founder alleles to estimate MAF
total_founder_alleles=0
tmp_haplotypes=OrderedDict()
tmp_mafs={}
for item in families:
tmp_haplotypes[item] = self.__PedToHaplotype(data.getFamSamples(item))
# count founder alleles
for hap in tmp_haplotypes[item]:
if not data.tfam.is_founder(hap[1]):
continue
total_founder_alleles+=1.0
for idxv, v in enumerate(data.getFamVariants(item,style="map")[0]):
if v not in tmp_mafs:
# [#alt, #haplotypes]
tmp_mafs[v] = [0, 0]
gt = hap[2 + idxv][1] if hap[2 + idxv][0].isupper() else hap[2 + idxv][0]
if not gt == "?":
#genotyped
tmp_mafs[v][0] += self.gtconv[gt]
else:
#genotype is missing
tmp_mafs[v][1] -= 1.0
#compute MAFs based on counts
for v in tmp_mafs:
if type(tmp_mafs[v]) is not list:
continue
tmp_mafs[v] = tmp_mafs[v][0] / (tmp_mafs[v][1]+total_founder_alleles) if tmp_mafs[v][1]+total_founder_alleles > 0 else 0.0
return tmp_mafs
def __Haplotype(self, data, haplotypes, mafs, varnames,recombPos,uniq_vars,exclude_vars):
'''genetic haplotyping. haplotypes stores per family data'''
# FIXME: it is SWIG's (2.0.12) fault not to properly destroy the object "Pedigree" in "Execute()"
# So there is a memory leak here which I tried to partially handle on C++
#
# Per family haplotyping
#
self.markers = ["V{}-{}".format(idx, item[1]) for idx, item in enumerate(data.variants)]
for item in data.families:
varnames[item], positions, vcf_mafs = data.getFamVariants(item, style = "map")
if len(varnames[item]) == 0:
for person in data.families[item]:
data[person] = self.missings
continue
if env.debug:
with env.lock:
sys.stderr.write('\n'.join(['\t'.join(x) for x in data.getFamSamples(item)]) + '\n\n')
# haplotyping
self.hap = {}
with env.lock:
if not env.prephased:
tmp_log_output=env.tmp_log + str(os.getpid())
#with stdoutRedirect(to = tmp_log_output + '.log'):
haplotypes[item] = self.haplotyper.Execute(data.chrom, varnames[item], sorted(positions),
data.getFamSamples(item), self.rsq, tmp_log_output)[0]
print('haplotyper execute',item,haplotypes[item])
self.hap[item] = haplotypes[item]
else:
haplotypes[item] = self.__PedToHaplotype(data.getFamSamples(item))
if len(haplotypes[item]) == 0:
# C++ haplotyping implementation failed
with env.chperror_counter.get_lock():
env.chperror_counter.value += 1
# either use privided MAF or computer MAF
if all(vcf_mafs):
for idx, v in enumerate(varnames[item]):
if v not in mafs:
mafs[v] = vcf_mafs[idx]
else:
# count founder alleles
for hap in haplotypes[item]:
if not data.tfam.is_founder(hap[1]):
continue
for idxv, v in enumerate(varnames[item]):
if v not in mafs:
# [#alt, #haplotypes]
mafs[v] = [0, 0]
gt = hap[2 + idxv][1] if hap[2 + idxv][0].isupper() else hap[2 + idxv][0]
if not gt == "?":
mafs[v][0] += self.gtconv[gt]
mafs[v][1] += 1.0
#
# Compute founder MAFs
#
for v in mafs:
if type(mafs[v]) is not list:
continue
mafs[v] = mafs[v][0] / mafs[v][1] if mafs[v][1] > 0 else 0.0
if env.debug:
with env.lock:
print("variant mafs = ", mafs, "\n", file = sys.stderr)
#
# Drop some variants if maf is greater than given threshold
#
if self.maf_cutoff is not None:
exclude_vars = []
for v in mafs.keys():
if mafs[v] > self.maf_cutoff:
exclude_vars.append(v)
for i in haplotypes.keys():
haplotypes[i] = listit(haplotypes[i])
for j in range(len(haplotypes[i])):
haplotypes[i][j] = haplotypes[i][j][:2] + \
[x for idx, x in enumerate(haplotypes[i][j][2:]) if varnames[i][idx] not in exclude_vars]
varnames[i] = [x for x in varnames[i] if x not in exclude_vars]
# handle trivial data
if len(varnames[i]) == 0:
for person in data.families[i]:
data[person] = self.missings
del varnames[i]
del haplotypes[i]
# count how many variants are removed
with env.commonvar_counter.get_lock():
env.commonvar_counter.value += len(exclude_vars)
def __ClusterByLD(self, data, haplotypes, varnames):
if self.r2 is None:
return None
# get founder haplotypes
founder_haplotypes = []
markers = sorted(set(itertools.chain(*varnames.values())), key = lambda x: int(x.split("-")[0][1:]))
for item in haplotypes:
for ihap, hap in enumerate(haplotypes[item]):
if not data.tfam.is_founder(hap[1]):
continue
gt = [hap[2 + varnames[item].index(v)] if v in varnames[item] else '?' for v in markers]
founder_haplotypes.append(("{}-{}".format(hap[1], ihap % 2), "".join([x[1] if x[0].isupper() else x[0] for x in gt])))
# calculate LD blocks, use r2 measure
ld = Align.create(founder_haplotypes).matrixLD(validCharacters="12")["r2"]
blocks = []
for j in ld:
block = [j]
for k in ld[j]:
if ld[j][k] > self.r2:
block.append(k)
if len(block) > 1:
blocks.append(block)
self.ld, self.blocks = ld, blocks
# get LD clusters
clusters = [[markers[idx] for idx in item] for item in list(connected_components(blocks))]
if env.debug:
with env.lock:
print("LD blocks: ", blocks, file = sys.stderr)
print("LD clusters: ", clusters, file = sys.stderr)
return clusters
def __CodeHaplotypes(self, data, haplotypes, mafs, varnames, clusters):
# apply CHP coding
for item in data.famvaridx:
if item not in haplotypes and data[data.families[item][0]] != ('0','0'):
# when only wild-type haplotypes are present in a family, still code them instead of ignoring the family
if self.freq_by_fam:
pop=data.freq_by_fam[item]
try:
varnames[item]=data.total_varnames[pop]
mafs[item]=data.total_mafs[pop]
except:
continue
else:
varnames[item]=data.total_varnames['pop']
mafs[item]=data.total_mafs
haplotypes[item]=[]
for person in data.families[item]:
tmp_person=[item, person]
if '00' in data[person]:
tmp_person+=['?:']*len(varnames[item])
else:
tmp_person+=['1:']*len(varnames[item])
haplotypes[item].append(tmp_person)
haplotypes[item].append(tmp_person)
elif item in haplotypes:
nonvar_hap_flag=False
#determine if wild-type haplotype is present in a family
for hap in haplotypes[item]:
tmp_genes=[]
for tmpa in hap[2:]:
if 'A' in tmpa or 'B' in tmpa:
tmp_genes.append(tmpa[1])
else:
tmp_genes.append(tmpa[0])
if set(tmp_genes)==set(['1']):
#non variant haplotype
nonvar_hap_flag=True
break
if not nonvar_hap_flag:
#if family don't have wild-type haplotype, add a fake one to ensure correct coding
var_num=len(varnames[item])
fake_person=[item, 'FAKEPERSON']+['1:']*var_num
haplotypes[item].append(fake_person)
for hidx,hap in enumerate(haplotypes[item]):
if hap[1] in data.missing_persons:
missing_person=[item,hap[1]]+['?:']*len(varnames[item])
haplotypes[item][hidx]=missing_person
if not clusters is None:
clusters_idx = [[[varnames[item].index(x) for x in y] for y in clusters] for item in haplotypes]
else:
clusters_idx = [[[]] for item in haplotypes]
if env.debug:
for item in haplotypes:
with env.lock:
print(varnames[item],file=sys.stderr)
print("hap{0}\t{1}\n".format(item,haplotypes[item]),file=sys.stderr)
self.coder.Execute(haplotypes.values(), [[mafs[item][v] for v in varnames[item]] for item in haplotypes], clusters_idx)
if env.debug:
with env.lock:
if clusters:
print("Family LD clusters: ", clusters_idx, "\n", file = sys.stderr)
self.coder.Print()
# line: [fid, sid, hap1, hap2]
for line in self.coder.GetHaplotypes():
if not line[1] in data:
# this sample is not in VCF file. Every variant site should be missing
# they have to be skipped for now
continue
data[line[1]] = (line[2].split(','), line[4].split(','))
#sub-region count for each sample individual
superMarkerCount=len(data[line[1]][0])
if line[0] not in data.patterns:
data.patterns[line[0]]=[[] for x in range(superMarkerCount)]
for t_Marker in range(superMarkerCount):
t_pat1=line[3].split(',')[t_Marker]
t_pat2=line[5].split(',')[t_Marker]
if t_pat1 not in data.patterns[line[0]][t_Marker]:
data.patterns[line[0]][t_Marker].append(t_pat1)
if t_pat2 not in data.patterns[line[0]][t_Marker]:
data.patterns[line[0]][t_Marker].append(t_pat2)
if len(data[line[1]][0]) > data.superMarkerCount:
data.superMarkerCount = len(data[line[1]][0])
# get MAF
for item in data.famvaridx:
if item not in haplotypes:
for person in data.families[item]:
data[person]=(['0']*data.superMarkerCount,['0']*data.superMarkerCount)
for item in haplotypes:
data.maf[item] = self.coder.GetAlleleFrequencies(item)
if not len(data.maf[item][0]):
continue
data.varnames_by_fam[item]=varnames[item]
wt_maf=0
if self.freq_by_fam:
try:
wt_maf=data.wt_maf[data.freq_by_fam[item]]
except:
pass
else:
wt_maf=data.wt_maf['pop']
tmp_data_maf=[]
for v in data.maf[item]:
if len(v)==1:
tmp_data_maf.append((v[0],1-v[0]))
else:
if np.sum(v)<1:
tmp_ratio=sum(v[1:])/(1-wt_maf)
tmp_list=[wt_maf]
if tmp_ratio==0:
tmp_list.append(1-wt_maf)
else:
for tmpv in v[1:]:
tmp_list.append(tmpv/tmp_ratio)
tmp_data_maf.append(tuple(tmp_list))
else:
tmp_data_maf.append(v)
data.maf[item]=tuple(tmp_data_maf)
if env.debug:
with env.lock:
print("marker freqs = ", data.maf, "\n", file = sys.stderr)
def __AssignSNVHaplotypes(self, data, haplotypes, mafs, varnames):
for item in haplotypes:
# each person's haplotype
data.varnames_by_fam[item]=varnames[item]
token = ''
for idx,line in enumerate(haplotypes[item]):
if line[1] in data.missing_persons:
data[line[1]]=('0','0')
else:
if not idx % 2:
token = line[2][1] if line[2][0].isupper() else line[2][0]
if token=='?':
token='0'
else:
tmp_token = line[2][1] if line[2][0].isupper() else line[2][0]
if tmp_token=='?':
tmp_token='0'
data[line[1]] = (token, tmp_token)
# get MAF
data.maf[item] = [(1 - mafs[item][varnames[item][0]], mafs[item][varnames[item][0]])]
data.maf[item] = tuple(tuple(np.array(v) / np.sum(v)) if np.sum(v) else v
for v in data.maf[item])
for item in data.famvaridx:
if item not in haplotypes and data[data.families[item][0]] != ('0','0'):
for person in data.families[item]:
if '00' in data[person]:
data[person]=('0','0')
else:
data[person]=('1','1')
t_maf=0
if self.freq_by_fam:
try:
t_maf=data.wt_maf[data.freq_by_fam[item]]
except:
for person in data.families[item]:
data[person]=('0','0')
else:
t_maf=data.wt_maf['pop']
data.maf[item]=((t_maf,1-t_maf),)
if env.debug:
with env.lock:
print("marker freqs = ", data.maf, "\n", file = sys.stderr)
def __FormatHaplotypes(self, data,recombPos,varnames,uniq_vars):
# Reformat sample genotypes
## Linhai Edit: Reformat to deal with recombination events in families
if self.recomb_perfam:
#code recombination per family basis, no need to consider overlap across families
for person in data:
if type(data[person]) is not tuple:
data[person] = self.missings
continue
diff = data.superMarkerCount - len(data[person][0])
data[person] = zip(*data[person])
if diff > 0:
data[person].extend([self.missings] * diff)
else:
#code recombination across families to generate sub-regions that extend across families
tmp_combined_recombPos={}
sorted_var = sorted(uniq_vars, key=lambda x: int(x.split('-')[0][1:]))
for fam in data.maf.keys():
if len(data.maf[fam])>1:
for pair in sorted(recombPos[fam].keys(), key=lambda x:(sorted_var.index(x[0]),sorted_var.index(x[1]))):
if pair[1] == varnames[fam][0]:
##remove recombination event if occurred at 1st RV
del recombPos[fam][pair]
continue
if fam not in tmp_combined_recombPos:
tmp_combined_recombPos[fam]=[pair]
else:
tmp_combined_recombPos[fam].append(pair)
tmp_all_recombs=[pair for pairs in tmp_combined_recombPos.values() for pair in pairs]
sorted_combined_recombPos=sorted(list(set(tmp_all_recombs)),key=lambda x:(sorted_var.index(x[0]),sorted_var.index(x[1])))
recomb_fams=tmp_combined_recombPos.keys()
##get sub-regions that applies to all families
for varidx,variant in enumerate(sorted_var):
included_fams=len(recomb_fams)
for recomb_region in sorted_combined_recombPos:
if varidx > sorted_var.index(recomb_region[0]) and varidx < sorted_var.index(recomb_region[1]):
##if the variant is in a recombination region
included_fams-=1
if included_fams==len(recomb_fams):
if data.combined_regions==[]:
data.combined_regions.append([variant])
else:
if sorted_var.index(data.combined_regions[-1][-1])==varidx-1:
neighbour_recomb_flag=False
for recomb_region in sorted_combined_recombPos:
recomb_idx=sorted_var.index(recomb_region[1])
if recomb_idx==varidx:
neighbour_recomb_flag=True
break
elif recomb_idx>varidx:
break
if neighbour_recomb_flag:
data.combined_regions.append([variant])
else:
data.combined_regions[-1].append(variant)
else:
data.combined_regions.append([variant])
##Get the markers in families compliant with the sub_regions
for sub_region in data.combined_regions:
markers={}
for fam in recomb_fams:
pidx=0
for pair in sorted(recombPos[fam].keys(), key=lambda x:(sorted_var.index(x[0]),sorted_var.index(x[1]))):
sub_region_start=sorted_var.index(sub_region[0])
sub_region_end=sorted_var.index(sub_region[-1])
recomb_start=sorted_var.index(pair[0])
recomb_end=sorted_var.index(pair[1])
if sub_region_end <= recomb_start:
markers[fam]=pidx
break
elif sub_region_end > recomb_start and sub_region_start>recomb_start and sub_region_end<recomb_end:
##within the recombination region
markers[fam]=None
break
pidx+=1
if fam not in markers:
markers[fam]=pidx
data.complied_markers.append(markers)
data.superMarkerCount=len(data.combined_regions)
#coordinates for sub_regions
data.coordinates_by_region=[(int(sub_region[0].split('-')[1])+int(sub_region[-1].split('-')[1]))/2 for sub_region in data.combined_regions]
for person in data:
if type(data[person]) is not tuple:
data[person] = self.missings
continue
diff = data.superMarkerCount - len(data[person][0])
data[person] = zip(*data[person])
if diff > 0:
if len(data[person]) == 1:
##only one whole region with no recombination
data[person].extend(data[person] * diff)
else:
famid=''
for fam in data.complied_markers[0].keys():
if person in data.families[fam]:
famid=fam
complied_data=[]
for marker in data.complied_markers:
complied_data.append(data[person][marker[famid]])
data[person]=complied_data
def __PedToHaplotype(self, ped):
'''convert prephased ped format to haplotype format.
Input: e.g. [['13346', '5888', '0', '0', '1', '11', '11', '11'], ['13346', '5856', '0', '0', '2', '12', '12', '12'], ['13346', '5920', '5888', '5856', '1', '12', '12', '12'], ['13346', '6589', '5888', '5856', '1', '11', '11', '11']]
Output: e.g. (('13346', '5856', '1:', '1:', '1:'), ('13346', '5856', '2:', '2:', '2:'), ('13346', '5888', '1:', '1:', '1:'), ('13346', '5888', '1:', '1:', '1:'), ('13346', '6589', '1:', '1|', '1|'), ('13346', '6589', '1:', '1|', '1|'), ('13346', '5920', '2:', '2|', '2|'), ('13346', '5920', '1:', '1|', '1|'))
'''
haps = []
for item in ped:
entry = [item[0], item[1]] + [x[0] + ':' if x[0] != '0' else '?:' for x in item[5:]]
haps.append(tuple(entry))
entry = [item[0], item[1]] + [x[1] + ':' if x[1] != '0' else '?:' for x in item[5:]]
haps.append(tuple(entry))
return tuple(haps)
def getRegion(self, region):
self.name = region[3]
# ### 6.LinkageWriter class
class LinkageWriter:
def __init__(self, num_missing_append = 0):
self.chrom = self.prev_chrom = self.name = self.distance = self.distance_avg = self.distance_m = self.distance_f = None
self.distance_by_region=[]
self.mid_position=None
self.reset()
self.missings = ["0", "0"]
self.num_missing = num_missing_append
def apply(self, data):
if self.chrom != self.prev_chrom:
if self.prev_chrom is None:
self.prev_chrom = self.chrom
else:
# new chrom entered,
# commit whatever is in buffer before accepting new data
self.commit()
# write tped output
position = str(data.getMidPosition())
if data.superMarkerCount <= 1:
# genotypes
gs = [data[s][0] for s in data.samples]
if len(set(gs)) == 1:
# everyone's genotype is the same (most likely missing or monomorphic)
return 2
self.tped += env.delimiter.join([self.chrom, self.name, self.distance, position] + \
list(itertools.chain(*gs)) + self.missings*self.num_missing) + "\n"
# freqs
for k in data.maf:
self.freq += env.delimiter.join([k, self.name] + map(str, data.maf[k][0])) + "\n"
else:
# have to expand each region into mutiple chunks to account for different recomb points
gs = zip(*[data[s] for s in data.samples])
# sub-chunk id
cid = 0
skipped_chunk = []
self.distance_by_region=[self.distance_converter(x,int(position)) for x in data.coordinates_by_region]
for idx, g in enumerate(gs):
if len(set(g)) == 1:
skipped_chunk.append(idx)
continue
cid += 1
self.tped += \
env.delimiter.join([self.chrom, '{}[{}]'.format(self.name, cid), self.distance_by_region[cid-1], position] + \
list(itertools.chain(*g)) + self.missings*self.num_missing) + "\n"
if cid == 0:
# everyone's genotype is the same (most likely missing or monomorphic)
return 2
# freqs
for k in data.maf:
cid = 0
for idx in range(data.superMarkerCount):
if idx in skipped_chunk:
continue
if not data.complied_markers:
#if recombination coded per family instead of across families
if idx >= len(data.maf[k]):
break
cid += 1
self.freq += env.delimiter.join([k, '{}[{}]'.format(self.name, cid)] + \
map(str, data.maf[k][idx])) + "\n"
else:
if len(data.maf[k])>1:
matched_idx=data.complied_markers[idx][k]
cid += 1
self.freq += env.delimiter.join([k, '{}[{}]'.format(self.name, cid)] + \
map(str, data.maf[k][matched_idx])) + "\n"
elif len(data.maf[k])==1:
cid += 1
self.freq += env.delimiter.join([k, '{}[{}]'.format(self.name, cid)] + \
map(str, data.maf[k][0])) + "\n"
if data.combined_regions:
self.chp += "CHP Super Marker positions: "+repr(data.combined_regions)+"\n"
for item in data.varnames_by_fam:
try:
pattern_txt=[tuple(sorted(data.patterns[item][tmarker],key=lambda x:x.count('2') )) for tmarker in range(len(data.patterns[item]))]
except:
pattern_txt=''
self.varfam += "{}\t{}\t{}\n".format(item,data.varnames_by_fam[item],pattern_txt)
if self.counter < env.batch:
self.counter += data.superMarkerCount
else:
self.commit()
return 0
def commit(self):
if self.tped:
with env.lock:
with open(os.path.join(env.tmp_cache, '{}.chr{}.tped'.format(env.output, self.prev_chrom)),
'a') as f:
f.write(self.tped)
if self.freq:
with env.lock:
with open(os.path.join(env.tmp_cache, '{}.chr{}.freq'.format(env.output, self.prev_chrom)),
'a') as f:
f.write(self.freq)
if self.chp:
with env.lock:
with open(os.path.join(env.tmp_cache, '{}.chr{}.chp'.format(env.output, self.prev_chrom)),
'a') as f:
f.write(self.chp)
if self.varfam:
with env.lock:
with open(os.path.join(env.tmp_cache, '{}.chr{}.var'.format(env.output, self.prev_chrom)),
'a') as f:
f.write(self.varfam)
self.reset()
def reset(self):
self.tped = ''
self.freq = ''
self.chp = ''
self.varfam = ''
self.counter = 0
self.prev_chrom = self.chrom
def distance_converter(self, x, mid_position):
delta=(x-mid_position)/1000000.0
distance='%.5f'%(float(self.distance_avg)+delta)
distance_m='%.5f'%(float(self.distance_m)+delta)
distance_f='%.5f'%(float(self.distance_f)+delta)
return ";".join([distance,distance_m,distance_f])
def getRegion(self, region):
self.chrom = region[0]
self.name, self.distance_avg, self.distance_m, self.distance_f = region[3:]
self.distance = ";".join([self.distance_avg, self.distance_m, self.distance_f])
# ### 7.EncoderWorker class
class EncoderWorker(Process):
def __init__(self, queue, length, data, extractor, coder, writer):
Process.__init__(self)
self.queue = queue
self.numGrps = float(length)
self.data = data
self.extractor = extractor
self.maker = coder
self.writer = writer
def report(self):
env.log('{:,d} units processed {{{:.2%}}} ...'.\
format(env.success_counter.value, env.total_counter.value / self.numGrps), flush = True)
def run(self):
while True:
try:
region = self.queue.pop(0) if isinstance(self.queue, list) else self.queue.get()
if region is None:
self.writer.commit()
self.report()
# total mendelian errors found
with env.mendelerror_counter.get_lock():
env.mendelerror_counter.value += self.maker.haplotyper.CountMendelianErrors()
# total recombination events found
with env.recomb_counter.get_lock():
env.recomb_counter.value += self.maker.coder.CountRecombinations()
break
else:
with env.total_counter.get_lock():
env.total_counter.value += 1
self.extractor.getRegion(region)
self.writer.getRegion(region)
self.maker.getRegion(region)
isSuccess = True
for m in [self.extractor, self.maker, self.writer]:
status = m.apply(self.data)
if status == -1:
with env.chperror_counter.get_lock():
# previous module failed
env.chperror_counter.value += 1
if status == 1:
with env.null_counter.get_lock():
env.null_counter.value += 1
if status == 2:
with env.trivial_counter.get_lock():
env.trivial_counter.value += 1
if status != 0:
isSuccess = False
break
if isSuccess:
with env.success_counter.get_lock():
env.success_counter.value += 1
if env.total_counter.value % (env.batch * env.jobs) == 0:
self.report()
except KeyboardInterrupt:
break
# # Old version
# +
class RData(dict):
def __init__(self, samples_vcf, tfam):
# tfam.samples: a dict of {sid:[fid, pid, mid, sex, trait], ...}
# tfam.families: a dict of {fid:[s1, s2 ...], ...}
self.tfam = tfam
# samples have to be in both vcf and tfam data
self.samples = OrderedDict([(k, tfam.samples[k]) for k in samples_vcf if k in tfam.samples])
# a dict of {fid:[member names], ...}
self.families = {k : [x for x in self.samples if x in tfam.families[k]] for k in tfam.families}
# a dict of {fid:[idx ...], ...}
self.famsampidx = {}
# a dict of {fid:[maf1, maf2 ...]}
self.maf = OrderedDict()
# reorder family samples based on order of VCF file
for k in self.families.keys():
if len(self.families[k]) == 0:
# skip families having no samples in VCF file
del self.families[k]
else:
self.famsampidx[k] = [i for i, x in enumerate(samples_vcf) if x in self.families[k]]
# a dict of {fid:[idx ...], ...}
self.famvaridx = {}
self.gss = {} #test line
self.reset()
def reset(self):
for item in self.samples:
self[item] = []
self.variants = []
self.chrom = None
for k in self.families:
self.famvaridx[k] = []
self.maf = OrderedDict()
# superMarkerCount is the max num. of recombinant fragments among all fams
self.superMarkerCount = 0
self.gss = {} #test line
def getMidPosition(self):
if len(self.variants) == 0:
return None
return sum([x[1] for x in self.variants]) / len(self.variants)
def getFamVariants(self, fam, style = None):
if style is None:
return [item for idx, item in enumerate(self.variants) if idx in self.famvaridx[fam]]
elif style == "map":
names = []
pos = []
mafs = []
for idx in self.famvaridx[fam]:
names.append("V{}-{}".format(idx, self.variants[idx][1]))
pos.append(self.variants[idx][1])
mafs.append(self.variants[idx][-1])
return names, pos, mafs
else:
raise ValueError("Unknown style '{}'".format(style))
def getFamSamples(self, fam):
nvar = len([item for idx, item in enumerate(self.variants) if idx in self.famvaridx[fam]])
output = [[]] * len(self.tfam.families[fam])
for idx, item in enumerate(self.tfam.sort_family(fam)):
# sample info, first 5 columns of ped
output[idx] = self.tfam.samples[item][:-1]
# sample genotypes
if item in self.samples:
output[idx].extend(self[item])
else:
output[idx].extend(["00"] * nvar)
return output
# -
# ### Old maker
# +
#export
class RegionExtractor:
'''Extract given genomic region from VCF
converting genotypes into dictionary of
genotype list'''
def __init__(self, filename, build = None, chr_prefix = None):
self.vcf = cstatgen.VCFstream(filename)
self.chrom = self.startpos = self.endpos = self.name = None
self.chr_prefix = chr_prefix
if build is None:
build = env.build
self.xchecker = PseudoAutoRegion('X', build)
self.ychecker = PseudoAutoRegion('Y', build)
def apply(self, data):
# Clean up
data.reset()
data.chrom = self.chrom
self.vcf.Extract(self.chrom, self.startpos, self.endpos)
if data.anno is None:
varIdx=self.extract_vcf(data)
else:
varIdx=self.extract_vcf_with_anno(data)
if varIdx == 0:
return 1
else:
with env.variants_counter.get_lock():
env.variants_counter.value += varIdx
return 0
def extract_vcf(self,data):
varIdx = 0
# for each variant site
while (self.vcf.Next()):
# check if the line's sample number matches the entire VCF sample number
if not self.vcf.CountSampleGenotypes() == self.vcf.sampleCount:
raise ValueError('Genotype and sample mismatch for region {}: {:,d} vs {:,d}'.\
format(self.name, self.vcf.CountSampleGenotypes(), self.vcf.sampleCount))
# skip tri-allelic sites
if not self.vcf.IsBiAllelic():
with env.triallelic_counter.get_lock():
env.triallelic_counter.value += 1
continue
# valid line found, get variant info
try:
maf = float(self.vcf.GetInfo(data.af_info))
except Exception as e:
raise ValueError("VCF line {}:{} does not have valid allele frequency field {}!".\
format(self.vcf.GetChrom(), self.vcf.GetPosition(), data.af_info))
# for each family assign member genotype if the site is non-trivial to the family
for k in data.families:
gs = self.vcf.GetGenotypes(data.famsampidx[k])
if len(set(''.join([x for x in gs if x != "00"]))) <= 1:
# skip monomorphic gs
continue
else:
# this variant is found in the family
data.famvaridx[k].append(varIdx)
data.famvarmafs[k].append(maf if maf < 0.5 else 1-maf)
for person, g in zip(data.families[k], gs):
data[person].append(g if maf<0.5 else self.reverse_genotypes(g))
data.variants.append([self.vcf.GetVariantID(), self.vcf.GetPosition(), self.name]) #remove maf
varIdx += 1
return varIdx
def extract_vcf_with_anno(self,data):
'''extract variants and annotation by region'''
if str(data.anno.Chr[0])!=self.chrom:
return 0
anno_idx = (data.anno.Start>=self.startpos) & (data.anno.Start<self.endpos)
if anno_idx.any()==False:
return 0
varmafs = data.anno[anno_idx]
varIdx = 0
i = -1
# for each variant site
while (self.vcf.Next()):
i += 1
# check if the line's sample number matches the entire VCF sample number
if not self.vcf.CountSampleGenotypes() == self.vcf.sampleCount:
raise ValueError('Genotype and sample mismatch for region {}: {:,d} vs {:,d}'.\
format(self.name, self.vcf.CountSampleGenotypes(), self.vcf.sampleCount))
# skip tri-allelic sites
if not self.vcf.IsBiAllelic():
with env.triallelic_counter.get_lock():
env.triallelic_counter.value += 1
continue
# valid line found, get variant info
try:
mafs=varmafs.loc[self.vcf.GetVariantID()][2:]
except:
print(self.vcf.GetVariantID(), 'is not in annotation')
continue
if mafs.any()==False:
continue
# for each family assign member genotype if the site is non-trivial to the family
for k in data.families:
gs = self.vcf.GetGenotypes(data.famsampidx[k])
if len(set(''.join([x for x in gs if x != "00"]))) <= 1:
# skip monomorphic gs
continue
else:
maf = mafs[data.fam_pop[k]]
if maf:
# this variant is found in the family
data.famvaridx[k].append(varIdx)
data.famvarmafs[k].append(maf if maf < 0.5 else 1-maf)
for person, g in zip(data.families[k], gs):
data[person].append(g if maf<0.5 else self.reverse_genotypes(g))
data.variants.append([self.vcf.GetVariantID(), self.vcf.GetPosition(), self.name]) #remove maf
#print(i,varmafs.shape,self.chrom, self.startpos, self.endpos, self.name,self.vcf.GetPosition())
varIdx += 1
return varIdx
def check_gs(self,gs):
'''skip monomorphic variants and singleton variants in a family'''
cg={'00':0,'11':0, '12':0, '22':0}
for i in gs:
cg[i]+=1
not00 = cg['11']+cg['12']+cg['22']
if cg['11']==not00 or cg['12']==not00 or cg['22']==not00:
#skip monomorphic variants
return False
if cg['12']+cg['22']<=1:
#skip sington variants
return False
return True
def reverse_genotypes(self,g):
''' 11->22,12,21,22->11 '''
if g=='11':
g='22'
elif g=='22':
g='11'
return g
def getRegion(self, region):
self.chrom, self.startpos, self.endpos, self.name = region[:4]
self.startpos = int(self.startpos)
self.endpos = int(self.endpos) + 1
if self.chrom in ['X','23']:
if self.xchecker.check(self.startpos) or self.xchecker.check(self.endpos):
self.chrom = 'XY'
if self.chrom in ['Y','24']:
if self.ychecker.check(self.startpos) or self.ychecker.check(self.endpos):
self.chrom = 'XY'
if self.chr_prefix and not self.chrom.startswith(self.chr_prefix):
self.chrom = self.chr_prefix + self.chrom
class MarkerMaker:
def __init__(self, wsize, maf_cutoff = None, recomb=False):
self.missings = ("0", "0")
self.gtconv = {'1':0, '2':1}
self.haplotyper = cstatgen.HaplotypingEngine(verbose = env.debug)
if wsize == 0 or wsize >= 1:
self.r2 = None
else:
self.r2 = wsize
self.coder = cstatgen.HaplotypeCoder(wsize)
self.maf_cutoff = maf_cutoff
self.rsq = 0.0
self.recomb = recomb
def getRegion(self, region):
self.name = region[3]
self.dtest = {}
self.dtest[self.name] = {}
self.dtest[self.name]['predata']={}
def apply(self, data):
#try:
# haplotyping plus collect found allele counts
# and computer founder MAFS
varnames,mafs,haplotypes=self.__Haplotype(data)
if len(varnames)==0:
return -1
if not any([len(varnames[x]) - 1 for x in varnames]):
# all families have only one variant
self.__AssignSNVHaplotypes(data, haplotypes, mafs, varnames)
else:
# calculate LD clusters using founder haplotypes
clusters = self.__ClusterByLD(data, haplotypes, varnames)
# recoding the genotype of the region
#env.dtest[self.name]['coder']['input'] = [data.copy(), haplotypes, mafs, varnames, clusters]
self.__CodeHaplotypes(data, haplotypes, mafs, varnames, clusters)
self.dtest[self.name]['maf']=data.maf
self.dtest[self.name]['hap']=self.haps
#env.dtest[self.name]['coder']['output'] = [self.coder.GetHaplotypes(),data.copy(),data.superMarkerCount,deepcopy(data.maf)]
#except Exception as e:
# return -1
self.__FormatHaplotypes(data)
#env.dtest[self.name]['format'] = data.copy()
return 0
def __Haplotype(self, data):
'''genetic haplotyping. haplotypes stores per family data'''
# FIXME: it is SWIG's (2.0.12) fault not to properly destroy the object "Pedigree" in "Execute()"
# So there is a memory leak here which I tried to partially handle on C++
#
# Per family haplotyping
#
items = get_family_with_var(data)
with ProcessPoolExecutor(max_workers = 8) as executor:
inputs = executor.map(phasing_haps,repeat(data.chrom),items,[data.getFamVariants(item, style = "map") for item in items],[data.getFamSamples(item) for item in items])
varnames,mafs,haplotypes = OrderedDict(),OrderedDict(),OrderedDict()
for item,item_varnames,item_mafs,item_haplotypes in inputs:
if len(item_haplotypes) == 0:
# C++ haplotyping implementation failed
with env.chperror_counter.get_lock():
env.chperror_counter.value += 1
env.log('{} family failed to phase haplotypes.'.format(item))
for person in data.families[item]:
data[person] = self.missings
continue
self.dtest[self.name]['predata'][item]=[item_varnames,item_mafs,item_haplotypes]
# Drop some variants if maf is greater than given threshold
if self.maf_cutoff is not None:
keep_idx = item_mafs<self.maf_cutoff
if not keep_idx.any():
for person in data.families[item]:
data[person] = self.missings
continue
item_mafs = item_mafs[keep_idx]
item_varnames = item_varnames[keep_idx]
item_haplotypes = item_haplotypes[:,np.concatenate(([True,True],keep_idx))]
varnames[item],mafs[item],haplotypes[item]= item_varnames,item_mafs,item_haplotypes
return varnames,mafs,haplotypes
def __ClusterByLD(self, data, haplotypes, varnames):
if self.r2 is None:
return None
# get founder haplotypes
founder_haplotypes = []
markers = sorted(set(itertools.chain(*varnames.values())), key = lambda x: int(x.split("-")[0][1:]))
for item in haplotypes:
for ihap, hap in enumerate(haplotypes[item]):
if not data.tfam.is_founder(hap[1]):
continue
gt = [hap[2 + list(varnames[item]).index(v)] if v in varnames[item] else '?' for v in markers]
founder_haplotypes.append(("{}-{}".format(hap[1], ihap % 2), "".join([x[1] if x[0].isupper() else x[0] for x in gt])))
# calculate LD blocks, use r2 measure
blocks = []
if sys.version_info.major == 2:
ld = Align.create(founder_haplotypes).matrixLD(validCharacters="12")["r2"]
for j in ld: #upper triangle
block = [j]
for k in ld[j]:
try:
if ld[j][k] > self.r2:
block.append(k)
except:
print('ld value',ld[j][k])
if len(block) > 1:
blocks.append(block)
else:
ldi,ld = egglib.stats.matrix_LD(Align.create(founder_haplotypes,egglib.Alphabet(cat='string',expl=['1','2'],miss='?')),('rsq'))
for j in range(len(ldi)): #lower triangle
block = [j]
for k in range(j+1,len(ldi)):
try:
if ld[k][j] > self.r2:
block.append(k)
except:
print('ld value',ld[k][j])
if len(block) > 1:
blocks.append(block)
# get LD clusters
clusters = [[markers[idx] for idx in item] for item in list(connected_components(blocks))]
#env.dtest[self.name]['ld'] = [ld,blocks,clusters]
if env.debug:
with env.lock:
print("LD blocks: ", blocks, file = sys.stderr)
print("LD clusters: ", clusters, file = sys.stderr)
return clusters
def __CodeHaplotypes(self, data, haplotypes, mafs, varnames, clusters):
# apply CHP coding
if clusters is not None:
clusters_idx = [[[list(varnames[item]).index(x) for x in y if x in varnames[item]] for y in clusters] for item in haplotypes]
else:
clusters_idx = [[[]] for item in haplotypes]
self.coder.Execute(list(haplotypes.values()), [mafs[item] for item in haplotypes], clusters_idx,self.recomb)
if env.debug:
with env.lock:
if clusters:
print("Family LD clusters: ", clusters_idx, "\n", file = sys.stderr)
self.coder.Print()
# line: [fid, sid, hap1, hap2]
self.haps = {}
for line in self.coder.GetHaplotypes():
#if not line[1] in data:
# this sample is not in VCF file. Every variant site should be missing
# they have to be skipped for now
# continue
data[line[1]] = (line[2].split(','), line[4].split(','))
self.haps[line[0]] = self.haps.get(line[0], line)
if len(data[line[1]][0]) > data.superMarkerCount:
data.superMarkerCount = len(data[line[1]][0])
# get MAF
for item in haplotypes:
data.maf[item] = self.coder.GetAlleleFrequencies(item)
data.maf[item] = tuple(tuple(np.array(v) / np.sum(v)) if np.sum(v) else v
for v in data.maf[item])
if env.debug:
with env.lock:
print("marker freqs = ", data.maf, "\n", file = sys.stderr)
def __AssignSNVHaplotypes(self, data, haplotypes, mafs, varnames):
print('SNVHap',self.name)
for item in haplotypes:
# each person's haplotype
token = ''
for idx, line in enumerate(haplotypes[item]):
if not idx % 2:
token = line[2][1] if line[2][0].isupper() else line[2][0]
else:
data[line[1]] = (token, line[2][1] if line[2][0].isupper() else line[2][0])
# get maf
data.maf[item] = [(1 - mafs[item][0], mafs[item][0])]
data.maf[item] = tuple(tuple(np.array(v) / np.sum(v)) if np.sum(v) else v
for v in data.maf[item])
if env.debug:
with env.lock:
print("marker freqs = ", data.maf, "\n", file = sys.stderr)
def __FormatHaplotypes(self, data):
# Reformat sample genotypes
for person in data:
if type(data[person]) is not tuple:
data[person] = self.missings
continue
diff = data.superMarkerCount - len(data[person][0])
data[person] = list(zip(*data[person]))
if diff > 0:
data[person].extend([self.missings] * diff)
def __PedToHaplotype(self, ped):
'''convert prephased ped format to haplotype format.
Input: e.g. [['13346', '5888', '0', '0', '1', '11', '11', '11'], ['13346', '5856', '0', '0', '2', '12', '12', '12'], ['13346', '5920', '5888', '5856', '1', '12', '12', '12'], ['13346', '6589', '5888', '5856', '1', '11', '11', '11']]
Output: e.g. (('13346', '5856', '1:', '1:', '1:'), ('13346', '5856', '2:', '2:', '2:'), ('13346', '5888', '1:', '1:', '1:'), ('13346', '5888', '1:', '1:', '1:'), ('13346', '6589', '1:', '1|', '1|'), ('13346', '6589', '1:', '1|', '1|'), ('13346', '5920', '2:', '2|', '2|'), ('13346', '5920', '1:', '1|', '1|'))
'''
haps = []
for item in ped:
entry = [item[0], item[1]] + [x[0] + ':' if x[0] != '0' else '?:' for x in item[5:]]
haps.append(tuple(entry))
entry = [item[0], item[1]] + [x[1] + ':' if x[1] != '0' else '?:' for x in item[5:]]
haps.append(tuple(entry))
return tuple(haps)
class LinkageWriter:
def __init__(self, num_missing_append = 0):
self.chrom = self.prev_chrom = self.name = self.distance = self.distance_avg = self.distance_m = self.distance_f = None
self.reset()
self.missings = ["0", "0"]
self.num_missing = num_missing_append
def apply(self, data):
if self.chrom != self.prev_chrom:
if self.prev_chrom is None:
self.prev_chrom = self.chrom
else:
# new chrom entered,
# commit whatever is in buffer before accepting new data
self.commit()
# write tped output
position = str(data.getMidPosition())
if data.superMarkerCount <= 1:
# genotypes
gs = [data[s][0] for s in data.tfam.samples]
if len(set(gs)) == 1:
# everyone's genotype is the same (most likely missing or monomorphic)
return 2
self.tped += env.delimiter.join([self.chrom, self.name, self.distance, position] + \
list(itertools.chain(*gs)) + self.missings*self.num_missing) + "\n"
# freqs
for k in data.maf:
self.freq += env.delimiter.join([k, self.name] + list(map(str, data.maf[k][0]))) + "\n"
else:
# have to expand each region into mutiple chunks to account for different recomb points
gs = list(zip(*[data[s] for s in data.tfam.samples]))
# sub-chunk id
cid = 0
skipped_chunk = []
for idx, g in enumerate(gs):
if len(set(g)) == 1:
skipped_chunk.append(idx)
continue
cid += 1
self.tped += \
env.delimiter.join([self.chrom, '{}[{}]'.format(self.name, cid), self.distance, position] + \
list(itertools.chain(*g)) + self.missings*self.num_missing) + "\n"
if cid == 0:
# everyone's genotype is the same (most likely missing or monomorphic)
return 2
# freqs
for k in data.maf:
cid = 0
for idx in range(data.superMarkerCount):
if idx in skipped_chunk:
continue
if idx >= len(data.maf[k]):
break
cid += 1
self.freq += env.delimiter.join([k, '{}[{}]'.format(self.name, cid)] + \
list(map(str, data.maf[k][idx]))) + "\n"
if self.counter < env.batch:
self.counter += data.superMarkerCount
else:
self.commit()
return 0
def commit(self):
if self.tped:
with env.lock:
with open(os.path.join(env.tmp_cache, '{}.chr{}.tped'.format(env.output, self.prev_chrom)),
'a') as f:
f.write(self.tped)
if self.freq:
with env.lock:
with open(os.path.join(env.tmp_cache, '{}.chr{}.freq'.format(env.output, self.prev_chrom)),
'a') as f:
f.write(self.freq)
self.reset()
def reset(self):
self.tped = ''
self.freq = ''
self.counter = 0
self.prev_chrom = self.chrom
def getRegion(self, region):
self.chrom = region[0]
self.name, self.distance_avg, self.distance_m, self.distance_f = region[3:]
self.distance = ";".join([self.distance_avg, self.distance_m, self.distance_f])
class EncoderWorker(Process):
def __init__(self, queue, length, data, extractor, coder, writer):
Process.__init__(self)
self.queue = queue
self.numGrps = float(length)
self.data = data
self.extractor = extractor
self.maker = coder
self.writer = writer
def report(self):
env.log('{:,d} units processed {{{:.2%}}} ...'.\
format(env.success_counter.value, env.total_counter.value / self.numGrps), flush = True)
def run(self):
while True:
try:
region = self.queue.pop(0) if isinstance(self.queue, list) else self.queue.get()
if region is None:
self.writer.commit()
self.report()
# total mendelian errors found
with env.mendelerror_counter.get_lock():
env.mendelerror_counter.value += self.maker.haplotyper.CountMendelianErrors()
# total recombination events found
with env.recomb_counter.get_lock():
env.recomb_counter.value += self.maker.coder.CountRecombinations()
break
else:
with env.total_counter.get_lock():
env.total_counter.value += 1
self.extractor.getRegion(region)
self.writer.getRegion(region)
self.maker.getRegion(region)
isSuccess = True
for m in [self.extractor, self.maker, self.writer]:
status = m.apply(self.data)
if status == -1:
with env.chperror_counter.get_lock():
# previous module failed
env.chperror_counter.value += 1
if status == 1:
with env.null_counter.get_lock():
env.null_counter.value += 1
if status == 2:
with env.trivial_counter.get_lock():
env.trivial_counter.value += 1
if status != 0:
isSuccess = False
break
if isSuccess:
with env.success_counter.get_lock():
env.success_counter.value += 1
if env.total_counter.value % (env.batch * env.jobs) == 0:
self.report()
except KeyboardInterrupt:
break
# -
# +
# export
def get_family_with_var(data):
items = []
for item,item_vars in data.famvaridx.items():
if len(item_vars) == 0: #no variants in the family
for person in data.families[item]:
data[person] = ("0", "0")
else:
items.append(item)
return items
haplotyper = cstatgen.HaplotypingEngine(verbose = env.debug)
def phasing_haps(chrom,item,fvar,fgeno):
item_varnames, positions, item_mafs = fvar
try:
item_haplotypes = haplotyper.Execute(chrom, item_varnames, positions, fgeno)[0]
except:
env.log("{} fail to phase haplotypes".format(item))
item_haplotypes = []
item_haplotypes = np.array(item_haplotypes)
return item,item_varnames,item_mafs,item_haplotypes
def run_each_region(regions,data,extractor,maker,writer):
'''get the haplotypes and allele frequency of variants in each region'''
results = {}
i=0
start = time.perf_counter()
for region in regions:
extractor.getRegion(region)
maker.getRegion(region)
writer.getRegion(region)
isSuccess = True
for m in [extractor, maker, writer]:
status = m.apply(data)
if status == -1:
with env.chperror_counter.get_lock():
# previous module failed
env.chperror_counter.value += 1
if status == 1:
with env.null_counter.get_lock():
env.null_counter.value += 1
if status == 2:
with env.trivial_counter.get_lock():
env.trivial_counter.value += 1
if status != 0:
isSuccess = False
break
if isSuccess:
with env.success_counter.get_lock():
env.success_counter.value += 1
results[region[3]]=maker.dtest[region[3]]
if len(results)==env.cache_size:
env.log('write to pickle: '+os.path.join(env.tmp_cache,env.output+str(i)+'.pickle')+',Gene number:'+str(len(results))+',Time:'+str((time.perf_counter()-start)/3600))
start = time.perf_counter()
with open(os.path.join(env.tmp_cache,env.output+str(i)+'.pickle'), 'wb') as handle:
pickle.dump(results, handle, protocol=pickle.HIGHEST_PROTOCOL)
results = {}
i +=1
env.log('write to pickle: '+os.path.join(env.tmp_cache,env.output+str(i)+'.pickle')+',Gene number:'+str(len(results))+',Time:'+str((time.perf_counter()-start)/3600))
with open(os.path.join(env.tmp_cache,env.output+str(i)+'.pickle'), 'wb') as handle:
pickle.dump(results, handle, protocol=pickle.HIGHEST_PROTOCOL)
results = {}
def run_each_region_genotypes(regions,data,extractor,maker,writer):
'''get the genotypes and allele frequency of variants in each region'''
results = {}
i=0
start = time.perf_counter()
for region in regions:
extractor.getRegion(region)
maker.getRegion(region)
writer.getRegion(region)
isSuccess = True
for m in [extractor]:
status = m.apply(data)
if status == -1:
with env.chperror_counter.get_lock():
# previous module failed
env.chperror_counter.value += 1
if status == 1:
with env.null_counter.get_lock():
env.null_counter.value += 1
if status == 2:
with env.trivial_counter.get_lock():
env.trivial_counter.value += 1
if status != 0:
isSuccess = False
break
if isSuccess:
with env.success_counter.get_lock():
env.success_counter.value += 1
#{'gene':{'predata':{'fam':[snp_ids,freq,genos]}}}
items = get_family_with_var(data)
predata={}
for item in items:
fvar=data.getFamVariants(item, style = "map")
fgeno=np.array(data.getFamSamples(item))
predata[item]=[fvar[0],fvar[2],fgeno]
results[region[3]]={'predata':predata}
if len(results)==25:
env.log('write to pickle: '+os.path.join(env.tmp_cache,env.output+str(i)+'.pickle')+',Gene number:'+str(len(results))+',Time:'+str((time.perf_counter()-start)/3600))
start = time.perf_counter()
with open(os.path.join(env.tmp_cache,env.output+str(i)+'.pickle'), 'wb') as handle:
pickle.dump(results, handle, protocol=pickle.HIGHEST_PROTOCOL)
results = {}
i +=1
env.log('write to pickle: '+os.path.join(env.tmp_cache,env.output+str(i)+'.pickle')+',Gene number:'+str(len(results))+',Time:'+str((time.perf_counter()-start)/3600))
with open(os.path.join(env.tmp_cache,env.output+str(i)+'.pickle'), 'wb') as handle:
pickle.dump(results, handle, protocol=pickle.HIGHEST_PROTOCOL)
results = {}
# -
def test(region,data,extractor,maker,writer):
extractor.getRegion(region)
maker.getRegion(region)
writer.getRegion(region)
status = extractor.apply(data)
if status == 1:
with env.null_counter.get_lock():
env.null_counter.value += 1
items = get_family_with_var(data)
print("Process parallel")
start = time.perf_counter()
with ProcessPoolExecutor(max_workers = 5) as executor:
maker_input = executor.map(phasing_haps,repeat(data.chrom),items,[data.getFamVariants(item, style = "map") for item in items],[data.getFamSamples(item) for item in items])
print(time.perf_counter()-start)
print("Thread parallel")
start = time.perf_counter()
with ThreadPoolExecutor() as executor:
maker_input = executor.map(phasing_haps,repeat(data.chrom),items,[data.getFamVariants(item, style = "map") for item in items],[data.getFamSamples(item) for item in items])
print(time.perf_counter()-start)
# # Main function
# +
checkParams(args)
if args.no_save:
cache = NoCache()
else:
cache = Cache(env.cache_dir, env.output, vars(args))
cache.setID('vcf')
# STEP 1: write encoded data to TPED format
if not args.vanilla and cache.check():
env.log('Loading regional marker data from archive ...')
cache.load(target_dir = env.tmp_dir, names = ['CACHE'])
env.success_counter.value = sum(map(fileLinesCount, glob.glob('{}/*.tped'.format(env.tmp_cache))))
env.batch = 10
else:
# load VCF file header
data = RData(args.vcf, args.tfam,args.anno,args.pop,allele_freq_info=args.freq)
vs = data.vs
samples_vcf = data.samples_vcf
if len(samples_vcf) == 0:
env.error("Fail to extract samples from [{}]".format(args.vcf), exit = True)
env.log('{:,d} samples found in [{}]'.format(len(samples_vcf), args.vcf))
samples_not_vcf = data.samples_not_vcf
if len(data.families) == 0:
env.error('No valid family to process. ' \
'Families have to be at least trio with at least one member in VCF file.', exit = True)
if len(data.samples) == 0:
env.error('No valid sample to process. ' \
'Samples have to be in families, and present in both TFAM and VCF files.', exit = True)
rewriteFamfile(os.path.join(env.tmp_cache, '{}.tfam'.format(env.output)),
data.tfam.samples, list(data.samples.keys()) + samples_not_vcf)
if args.single_markers:
regions = [(x[0], x[1], x[1], "{}:{}".format(x[0], x[1]), '.', '.', '.')
for x in data.vs.GetGenomeCoordinates()]
args.blueprint = None
elif not args.blueprint:
# load blueprint
try:
env.log('Loading marker map from [{}] ...'.format(args.blueprint))
with open(args.blueprint, 'r') as f:
regions = [x.strip().split() for x in f.readlines()]
except IOError:
env.error("Cannot load regional marker blueprint [{}]. ".format(args.blueprint), exit = True)
else:
env.log('separate chromosome to regions')
regions=data.get_regions(step=100)
env.log('{:,d} families with a total of {:,d} samples will be scanned for {:,d} pre-defined units'.\
format(len(data.families), len(data.samples), len(regions)))
env.jobs = max(min(args.jobs, len(regions)), 1)
regions.extend([None] * env.jobs)
queue = [] if env.jobs == 1 else Queue()
# -
# # Testing
extractor =RegionExtractor(args.vcf, build=env.build,chr_prefix = args.chr_prefix)
maker = MarkerMaker(args.bin, maf_cutoff = args.maf_cutoff,recomb=False)
writer = LinkageWriter(len(samples_not_vcf))
for j, region in enumerate(regions[:5]):#[:20]):
i = 0
#for region in rg:
extractor.getRegion(region)
maker.getRegion(region)
writer.getRegion(region)
isSuccess = True
for m in [extractor, maker, writer]:
status = m.apply(data)
print(data)
i+=1
if status == -1:
with env.chperror_counter.get_lock():
# previous module failed
env.chperror_counter.value += 1
if status == 1:
with env.null_counter.get_lock():
env.null_counter.value += 1
if status == 2:
with env.trivial_counter.get_lock():
env.trivial_counter.value += 1
if status != 0:
isSuccess = False
break
if isSuccess:
with env.success_counter.get_lock():
env.success_counter.value += 1
if j%1000==0:
print(j,i,len(data.variants),region)
env.dtest
env.dtest['PAPPA2']
env.dtest['MC1R']
env.dtest['WASH7P']
env.dtest['WASH7P']
a1 = {k:env.dtest[k] for k in ['MC1R','PAPPA2']}
# +
import pickle
with open('../data/dtestpy3_fixedcoder_seqtest_20220126.pickle', 'wb') as handle:
pickle.dump(a1, handle, protocol=pickle.HIGHEST_PROTOCOL)
#with open('dtestpy3_fixedcoder.pickle', 'rb') as handle:
# b = pickle.load(handle)
# -
a = {k:env.dtest[k] for k in ['MC1R','PAPPA2']}
# +
import pickle
with open('dtestpy3.pickle', 'wb') as handle:
pickle.dump(a, handle, protocol=pickle.HIGHEST_PROTOCOL)
#with open('dtestpy3.pickle', 'rb') as handle:
# b = pickle.load(handle)
# -
a['MC1R']
a['PAPPA2']
# ## One region test
for i,g in enumerate(regions):
if g[3]=='TPTEP1':
print(i)
region = regions[2]
region
extractor.getRegion(region)
maker.getRegion(region)
writer.getRegion(region)
extractor.apply(data)
items = get_family_with_var(data)
with open('../data/wg20220316/chr22test/TPTEP1genodata.pickle', 'wb') as handle:
pickle.dump({item:data.getFamSamples(item) for item in items}, handle, protocol=pickle.HIGHEST_PROTOCOL)
# ## concurrent.futures
import pickle
with open('../data/test_parallel.pickle', 'wb') as handle:
pickle.dump(data, handle, protocol=pickle.HIGHEST_PROTOCOL)
item = list(data.families.keys())[285]
item = '264'
item_varnames, positions, item_mafs = data.getFamVariants(item, style = "map")
item_varnames
tmp_log_output=env.tmp_log + str(os.getpid()) + '.log'
item_haplotypes = maker.haplotyper.Execute(data.chrom, item_varnames, positions, data.getFamSamples(item), 0, tmp_log_output)[0]
def test(item):
item_varnames, positions, item_mafs = data.getFamVariants(item, style = "map")
print(item,len(item_varnames))
tmp_log_output=env.tmp_log + str(os.getpid()) + '.log'
item_haplotypes = maker.haplotyper.Execute(data.chrom, item_varnames, positions, data.getFamSamples(item), 0, tmp_log_output)[0]
start = time.perf_counter()
count = 0
for item in data.families:
test(item)
print(time.perf_counter()-start,count)
from concurrent.futures import ProcessPoolExecutor
start = time.perf_counter()
with concurrent.futures.ProcessPoolExecutor() as executor:
results = executor.map(test,data.families.keys())
print(time.perf_counter()-start,count)
maker.rsq
maker.rsqply(data)
1
data
data.maf
haplotypes = OrderedDict()
mafs = {} ##Per fam per variant
varnames = {}
# maker._MarkerMaker__Haplotype(data, haplotypes, mafs, varnames)
import pandas as pd
new_trim_ped = pd.read_csv('../data/new_trim_ped.csv')
tmp = new_trim_ped.fid.value_counts()
tmp.index
tmp[tmp<=24]
#items = list(data.families.keys())
items = list(tmp[tmp<=24].index)
items.remove('10R_R114:0')
items.remove('10R_R57:0')
new_trim_ped[new_trim_ped.fid.isin(items[10:15])]
# ### 4.2 Debug of failing to phase haplotypes(10R_R114:0)
# #### remove the children without vcf
data.tfam.families['10R_R114:0'].remove('10R_R114_10')
data.tfam.families['10R_R114:0'].remove('10R_R114_11')
#data.tfam.families['10R_R114:0'].remove('10R_R114_6')
#data.tfam.families['10R_R114:0'].remove('10R_R114_8')
data.tfam.families_sorted['10R_R114:0'].remove('10R_R114_10')
data.tfam.families_sorted['10R_R114:0'].remove('10R_R114_11')
#data.tfam.families_sorted['10R_R114:0'].remove('10R_R114_6')
#data.tfam.families_sorted['10R_R114:0'].remove('10R_R114_8')
for item in ['10R_R114:0','10R_R57:0']:
varnames[item], positions, vcf_mafs = data.getFamVariants(item, style = "map")
if len(varnames[item]) == 0:
print('here')
for person in data.families[item]:
data[person] = maker.missings
print(pd.DataFrame(data.getFamSamples(item)))
# ### 4.3 write out the input of haplotyper
# #### 4.3.1 10R_R114:0 family
item = '10R_R114:0'
#for item in data.families:
print('running family',item)
varnames[item], positions, vcf_mafs = data.getFamVariants(item, style = "map")
a1 = [data.chrom, varnames[item], sorted(positions), data.getFamSamples(item), maker.rsq]
import pickle
with open('../data/test_haplotyper_10R_R114:0_removed2.pickle', 'wb') as handle:
pickle.dump(a1, handle, protocol=pickle.HIGHEST_PROTOCOL)
# #### 4.3.2 all families
varnames,positions,vcf_mafs,geno = {},{},{},{}
for item in data.families:
varnames[item], positions[item], vcf_mafs[item] = data.getFamVariants(item, style = "map")
geno[item] = data.getFamSamples(item)
a1 = [varnames, positions, vcf_mafs,geno]
import pickle
with open('../data/test_haplotyper_all_families.pickle', 'wb') as handle:
pickle.dump(a1, handle, protocol=pickle.HIGHEST_PROTOCOL)
len(varnames.keys())
item = '10R_R114:0'
#for item in data.families:
print('running family',item)
varnames[item], positions, vcf_mafs = data.getFamVariants(item, style = "map")
if len(varnames[item]) == 0:
print('here')
for person in data.families[item]:
data[person] = maker.missings
else:
print(varnames[item])
tmp_log_output=env.tmp_log + str(os.getpid())
maker.haplotyper.Execute(data.chrom, varnames[item], sorted(positions), data.getFamSamples(item), maker.rsq, tmp_log_output)
item
data.getFamSamples(item)
if all(vcf_mafs):
for idx, v in enumerate(varnames[item]):
if v not in mafs:
mafs[v] = vcf_mafs[idx]
if len(varnames):
if not any ([len(varnames[x]) - 1 for x in varnames]):
# all families have only one variant
self.__AssignSNVHaplotypes(data, haplotypes, mafs, varnames)
else:
# calculate LD clusters using founder haplotypes
#clusters = self.__ClusterByLD(data, haplotypes, varnames)
clusters=[]
#print('clusters:',clusters)
# recoding the genotype of the region
env.dtest[self.name]['coder']['input'] = [data.copy(), haplotypes, mafs, varnames, clusters]
self.__CodeHaplotypes(data, haplotypes, mafs, varnames, clusters)
env.dtest[self.name]['coder']['output'] = [self.coder.GetHaplotypes(),data.copy(),data.superMarkerCount,deepcopy(data.maf)]
#except Exception as e:
# return -1
self.__FormatHaplotypes(data)
self.markers = ["V{}-{}".format(idx, item[1]) for idx, item in enumerate(data.variants)]
for item in data.families:
varnames[item], positions, vcf_mafs = data.getFamVariants(item, style = "map")
if len(varnames[item]) == 0:
for person in data.families[item]:
data[person] = self.missings
continue
if env.debug:
with env.lock:
sys.stderr.write('\n'.join(['\t'.join(x) for x in data.getFamSamples(item)]) + '\n\n')
# haplotyping
with env.lock:
if not env.prephased:
#with stdoutRedirect(to = env.tmp_log + str(os.getpid()) + '.log'):
# haplotypes[item] = self.haplotyper.Execute(data.chrom, varnames[item],
# sorted(positions), data.getFamSamples(item))[0]
tmp_log_output=env.tmp_log + str(os.getpid())
#with stdoutRedirect(to = tmp_log_output + '.log'):
haplotypes[item] = self.haplotyper.Execute(data.chrom, varnames[item], sorted(positions),
data.getFamSamples(item), self.rsq, tmp_log_output)[0]
else:
haplotypes[item] = self.__PedToHaplotype(data.getFamSamples(item))
if len(haplotypes[item]) == 0:
# C++ haplotyping implementation failed
with env.chperror_counter.get_lock():
env.chperror_counter.value += 1
# either use privided MAF or computer MAF
if all(vcf_mafs):
for idx, v in enumerate(varnames[item]):
if v not in mafs:
mafs[v] = vcf_mafs[idx]
else:
# count founder alleles
for hap in haplotypes[item]:
if not data.tfam.is_founder(hap[1]):
continue
for idxv, v in enumerate(varnames[item]):
if v not in mafs:
# [#alt, #haplotypes]
mafs[v] = [0, 0]
gt = hap[2 + idxv][1] if hap[2 + idxv][0].isupper() else hap[2 + idxv][0]
if not gt == "?":
mafs[v][0] += self.gtconv[gt]
mafs[v][1] += 1.0
#
maker._MarkerMaker__Haplotype(data, haplotypes, mafs, varnames)
# ### Test3
if env.triallelic_counter.value:
env.log('{:,d} tri-allelic loci were ignored'.format(env.triallelic_counter.value))
if env.commonvar_counter.value:
env.log('{:,d} variants ignored due to having MAF > {} and other specified constraints'.\
format(env.commonvar_counter.value, args.maf_cutoff))
if env.null_counter.value:
env.log('{:,d} units ignored due to absence in VCF file'.format(env.null_counter.value))
if env.trivial_counter.value:
env.log('{:,d} units ignored due to absence of variation in samples'.format(env.trivial_counter.value))
fatal_errors = 0
try:
# Error msg from C++ extension
os.system("cat {}/*.* > {}".format(env.tmp_dir, env.tmp_log))
fatal_errors = wordCount(env.tmp_log)['fatal']
except KeyError:
pass
if env.chperror_counter.value:
env.error("{:,d} regional markers failed to be generated due to haplotyping failures!".\
format(env.chperror_counter.value))
if fatal_errors:
env.error("{:,d} or more regional markers failed to be generated due to runtime errors!".\
format(fatal_errors))
env.log('Archiving regional marker data to directory [{}]'.format(env.cache_dir))
cache.write(arcroot = 'CACHE', source_dir = env.tmp_cache)
env.jobs = args.jobs
env.tmp_cache
os.listdir(env.tmp_cache)
env.jobs
env.tmp_cache
# STEP 2: write to PLINK or mega2 format
tpeds = [os.path.join(env.tmp_cache, item) for item in os.listdir(env.tmp_cache) if item.startswith(env.output) and item.endswith('.tped')]
for fmt in args.format:
cache.setID(fmt)
if not args.vanilla and cache.check():
env.log('Loading {} data from archive ...'.format(fmt.upper()))
cache.load(target_dir = env.tmp_dir, names = [fmt.upper()])
else:
env.log('{:,d} units will be converted to {} format'.format(env.success_counter.value, fmt.upper()))
env.format_counter.value = 0
format(tpeds, os.path.join(env.tmp_cache, "{}.tfam".format(env.output)),
args.prevalence, args.wild_pen, args.muta_pen, fmt,
args.inherit_mode, args.theta_max, args.theta_inc)
env.log('{:,d} units successfully converted to {} format\n'.\
format(env.format_counter.value, fmt.upper()), flush = True)
if env.skipped_counter.value:
# FIXME: perhaps we need to rephrase this message?
env.log('{} region - family pairs skipped'.\
format(env.skipped_counter.value))
env.log('Archiving {} format to directory [{}]'.format(fmt.upper(), env.cache_dir))
cache.write(arcroot = fmt.upper(),
source_dir = os.path.join(env.tmp_dir, fmt.upper()), mode = 'a')
mkpath(env.outdir)
# ## 2. Testing run_linkage
args.run_linkage = True
cache.setID('analysis')
env.output
# ?cache.load
cache.cache_name
not args.vanilla and cache.check()
fmt = args.format[0]
args.blueprint
args.theta_inc
args.theta_max
args.output_limit
if args.run_linkage:
cache.setID('analysis')
if not args.vanilla and cache.check():
env.log('Loading linkage analysis result from archive ...'.format(fmt.upper()))
cache.load(target_dir = env.output, names = ['heatmap'])
else:
env.log('Running linkage analysis ...'.format(fmt.upper()))
run_linkage(args.blueprint, args.theta_inc, args.theta_max, args.output_limit)
env.log('Linkage analysis succesfully performed for {:,d} units\n'.\
format(env.run_counter.value, fmt.upper()), flush = True)
if env.makeped_counter.value:
env.log('{} "makeped" runtime errors occurred'.format(env.makeped_counter.value))
if env.pedcheck_counter.value:
env.log('{} "pedcheck" runtime errors occurred'.format(env.pedcheck_counter.value))
if env.unknown_counter.value:
env.log('{} "unknown" runtime errors occurred'.format(env.unknown_counter.value))
if env.mlink_counter.value:
env.log('{} "mlink" runtime errors occurred'.format(env.mlink_counter.value))
cache.write(arcroot = 'heatmap', source_dir = os.path.join(env.output, 'heatmap'), mode = 'a')
html(args.theta_inc, args.theta_max, args.output_limit)
else:
env.log('Saving data to [{}]'.format(os.path.abspath(env.output)))
cache.load(target_dir = env.output, names = [fmt.upper() for fmt in args.format])
env.log('Saving data to [{}]'.format(os.path.abspath(env.output)))
cache.load(target_dir = env.output, names = [fmt.upper() for fmt in args.format])
# ## Testing main
if args.no_save:
cache = NoCache()
else:
cache = Cache(env.cache_dir, env.output, vars(args))
cache.setID('vcf')
# STEP 1: write encoded data to TPED format
if not args.vanilla and cache.check():
env.log('Loading regional marker data from archive ...')
cache.load(target_dir = env.tmp_dir, names = ['CACHE'])
env.success_counter.value = sum(map(fileLinesCount, glob.glob('{}/*.tped'.format(env.tmp_cache))))
env.batch = 10
else:
# load VCF file header
checkVCFBundle(args.vcf)
cache.clear()
try:
vs = cstatgen.VCFstream(args.vcf)
except Exception as e:
env.error("{}".format(e), exit = True)
samples_vcf = vs.GetSampleNames()
if len(samples_vcf) == 0:
env.error("Fail to extract samples from [{}]".format(args.vcf), exit = True)
env.log('{:,d} samples found in [{}]'.format(len(samples_vcf), args.vcf))
samples_not_vcf = checkSamples(samples_vcf, getColumn(args.tfam, 2))[1]
# load sample info
data = RData(samples_vcf, TFAMParser(args.tfam))
if len(data.families) == 0:
env.error('No valid family to process. ' \
'Families have to be at least trio with at least one member in VCF file.', exit = True)
if len(data.samples) == 0:
env.error('No valid sample to process. ' \
'Samples have to be in families, and present in both TFAM and VCF files.', exit = True)
rewriteFamfile(os.path.join(env.tmp_cache, '{}.tfam'.format(env.output)),
data.tfam.samples, list(data.samples.keys()) + samples_not_vcf)
if args.single_markers:
regions = [(x[0], x[1], x[1], "{}:{}".format(x[0], x[1]), '.', '.', '.')
for x in vs.GetGenomeCoordinates()]
args.blueprint = None
else:
# load blueprint
try:
env.log('Loading marker map from [{}] ...'.format(args.blueprint))
with open(args.blueprint, 'r') as f:
regions = [x.strip().split() for x in f.readlines()]
except IOError:
env.error("Cannot load regional marker blueprint [{}]. ".format(args.blueprint), exit = True)
env.log('{:,d} families with a total of {:,d} samples will be scanned for {:,d} pre-defined units'.\
format(len(data.families), len(data.samples), len(regions)))
env.jobs = max(min(args.jobs, len(regions)), 1)
regions.extend([None] * env.jobs)
queue = Queue()
try:
faulthandler.enable(file=open(env.tmp_log + '.SEGV', 'w'))
for i in regions:
queue.put(i)
jobs = [EncoderWorker(
queue, len(regions), deepcopy(data),
RegionExtractor(args.vcf, chr_prefix = args.chr_prefix, allele_freq_info = args.freq),
MarkerMaker(args.bin, maf_cutoff = args.maf_cutoff),
LinkageWriter(len(samples_not_vcf))
) for i in range(env.jobs)]
for j in jobs:
j.start()
for j in jobs:
j.join()
faulthandler.disable()
except KeyboardInterrupt:
# FIXME: need to properly close all jobs
raise ValueError("Use 'killall {}' to properly terminate all processes!".format(env.prog))
else:
env.log('{:,d} units (from {:,d} variants) processed; '\
'{:,d} Mendelian inconsistencies and {:,d} recombination events handled\n'.\
format(env.success_counter.value,
env.variants_counter.value,
env.mendelerror_counter.value,
env.recomb_counter.value), flush = True)
if env.triallelic_counter.value:
env.log('{:,d} tri-allelic loci were ignored'.format(env.triallelic_counter.value))
if env.commonvar_counter.value:
env.log('{:,d} variants ignored due to having MAF > {}'.\
format(env.commonvar_counter.value, args.maf_cutoff))
if env.null_counter.value:
env.log('{:,d} units ignored due to absence in VCF file'.format(env.null_counter.value))
if env.trivial_counter.value:
env.log('{:,d} units ignored due to absence of variation in samples'.format(env.trivial_counter.value))
fatal_errors = 0
try:
# Error msg from C++ extension
os.system("cat {}/*.* > {}".format(env.tmp_dir, env.tmp_log))
fatal_errors = wordCount(env.tmp_log)['fatal']
except KeyError:
pass
if env.chperror_counter.value:
env.error("{:,d} regional markers failed to be generated due to haplotyping failures!".\
format(env.chperror_counter.value))
if fatal_errors:
env.error("{:,d} or more regional markers failed to be generated due to runtime errors!".\
format(fatal_errors))
env.log('Archiving regional marker data to directory [{}]'.format(env.cache_dir))
cache.write(arcroot = 'CACHE', source_dir = env.tmp_cache)
env.jobs = args.jobs
# STEP 2: write to PLINK or mega2 format
tpeds = [os.path.join(env.tmp_cache, item) for item in os.listdir(env.tmp_cache) if item.startswith(env.output) and item.endswith('.tped')]
for fmt in args.format:
print(fmt.lower())
cache.setID(fmt.lower())
if not args.vanilla and cache.check():
env.log('Loading {} data from archive ...'.format(fmt.upper()))
cache.load(target_dir = env.tmp_dir, names = [fmt.upper()])
else:
env.log('{:,d} units will be converted to {} format'.format(env.success_counter.value, fmt.upper()))
env.format_counter.value = 0
format(tpeds, os.path.join(env.tmp_cache, "{}.tfam".format(env.output)),
args.prevalence, args.wild_pen, args.muta_pen, fmt,
args.inherit_mode, args.theta_max, args.theta_inc)
env.log('{:,d} units successfully converted to {} format\n'.\
format(env.format_counter.value, fmt.upper()), flush = True)
if env.skipped_counter.value:
# FIXME: perhaps we need to rephrase this message?
env.log('{} region - family pairs skipped'.\
format(env.skipped_counter.value))
env.log('Archiving {} format to directory [{}]'.format(fmt.upper(), env.cache_dir))
cache.write(arcroot = fmt.upper(),
source_dir = os.path.join(env.tmp_dir, fmt.upper()), mode = 'a')
mkpath(env.outdir)
if args.run_linkage:
cache.setID('analysis')
if not args.vanilla and cache.check():
env.log('Loading linkage analysis result from archive ...'.format(fmt.upper()))
cache.load(target_dir = env.output, names = ['heatmap'])
else:
env.log('Running linkage analysis ...'.format(fmt.upper()))
run_linkage(args.blueprint, args.theta_inc, args.theta_max, args.output_limit)
env.log('Linkage analysis succesfully performed for {:,d} units\n'.\
format(env.run_counter.value, fmt.upper()), flush = True)
if env.makeped_counter.value:
env.log('{} "makeped" runtime errors occurred'.format(env.makeped_counter.value))
if env.pedcheck_counter.value:
env.log('{} "pedcheck" runtime errors occurred'.format(env.pedcheck_counter.value))
if env.unknown_counter.value:
env.log('{} "unknown" runtime errors occurred'.format(env.unknown_counter.value))
if env.mlink_counter.value:
env.log('{} "mlink" runtime errors occurred'.format(env.mlink_counter.value))
cache.write(arcroot = 'heatmap', source_dir = os.path.join(env.output, 'heatmap'), mode = 'a')
html(args.theta_inc, args.theta_max, args.output_limit)
else:
env.log('Saving data to [{}]'.format(os.path.abspath(env.output)))
cache.load(target_dir = env.output)
1
args.run_linkage = True
region = ['1', '69090', '70008', 'OR4F5', '4.866641545668504e-06', '6.181823219621424e-06', '3.6135725636621673e-06']
extractor.getRegion(region)
maker.getRegion(region)
writer.getRegion(region)
haplotypes = OrderedDict()
mafs = {} ##Per fam per variant
uniq_vars = []
exclude_vars = []
varnames = {}
recombPos = {}
extractor.apply(data)
maker._MarkerMaker__Haplotype(data, haplotypes, mafs, varnames,recombPos,uniq_vars,exclude_vars)
haplotypes['668']
maker._MarkerMaker__ClusterByLD(data, haplotypes, varnames)
maker._MarkerMaker__CodeHaplotypes(data, haplotypes, mafs, varnames, [])
clusters = []
if clusters is not None:
clusters_idx = [[[varnames[item].index(x) for x in y] for y in clusters] for item in haplotypes]
else:
clusters_idx = [[[]] for item in haplotypes]
maker.coder.Execute(haplotypes.values(), [[mafs[v] for v in varnames[item]] for item in haplotypes], clusters_idx)
maker.coder.Print()
maker.ld
[[mafs[item][v] for v in varnames[item]] for item in haplotypes]
varnames['668']
mafs
# ## Test clusterbyld
data.freq
data.variants
maker.apply(data)
haplotypes = OrderedDict()
mafs = {}
varnames = {}
maker._MarkerMaker__Haplotype(data, haplotypes, mafs, varnames)
type(data)
markers = ["V{}-{}".format(idx, item[1]) for idx, item in enumerate(data.variants)]
item = list(data.families.keys())[2]
item ='1036'
varnames = {}
varnames[item], positions, vcf_mafs = data.getFamVariants(item, style = "map")
varnames
item
varnames
tmp
maker.haplotyper.Execute(data.chrom, varnames[item], sorted(positions), data.getFamSamples(item))[0]
recombPos={}
varnames[item], positions, vcf_mafs = data.getFamVariants(item, style = "map")
recombPos[item]={}
var_for_haplotype=[]
positions_for_haplotype=[]
output_sample=[]
data.gnomAD_estimate.keys()
data.freq
positions
var_for_haplotype=varnames[item]
positions_for_haplotype=positions
item
famid =item
sorted_names = []
S_no_parents = filter(lambda x: True if data.tfam.is_founder(x) else False, data.tfam.families[famid])
graph = data.tfam.graph[famid].copy()
list(S_no_parents)
data.tfam.families[famid]
graph
while(S_no_parents):
n = S_no_parents.pop()
sorted_names.append(n)
if n not in graph:
continue
offsprings = graph.pop(n)
for m in offsprings:
father, mother = data.tfam.get_parents(m)
if father not in graph and mother not in graph:
S_no_parents.append(m)
if graph:
raise ValueError("There is a loop in the pedigree: {}\n".format(' '.join(graph.keys())))
else:
return sorted_names
data.tfam
#collect sample+genotypes
for person in data.tfam.sort_family(item):
output_sample.append([])
last_ele=len(output_sample)-1
output_sample[last_ele] = data.tfam.samples[person][:-1]
if person in data.samples:
for marker in var_for_haplotype:
idx=int(marker.split('-')[0][1:])
output_sample[last_ele].append(data.genotype_all[person][idx])
else:
output_sample[last_ele].extend(["00"] * len(var_for_haplotype))
len(data.tfam.sort_family(item))
set(output_sample[0][5:])
env.tmp_log
haplotypes = {}
tmp_log_output=env.tmp_log + str(os.getpid())
haplotypes[item] = maker.haplotyper.Execute(data.chrom, var_for_haplotype, positions_for_haplotype, output_sample, maker.rsq, tmp_log_output)[0]
1
var_for_haplotype
positions_for_haplotype
haplotypes
1+1
haplotypes
str(os.getpid())
var_for_haplotype
positions_for_haplotype
output_sample
maker.rsq
haplotypes['1']
#for item in haplotypes:
for hap_idx,haploid in enumerate(haplotypes[item]):
for vidx,var in enumerate(haploid[2:]):
if not var.endswith(':') and not var.endswith('|') and vidx!=0:
postvar_name=varnames[item][vidx]
prevar_name=varnames[item][vidx-1]
recomb_pair = (prevar_name,postvar_name)
print('run this')
try:
recombPos[item][recomb_pair].append(hap_idx)
except:
recombPos[item][recomb_pair]=[hap_idx]
haploid
var
haplotypes['1']
mafs
varnames
recombPos
uniq_vars
exclude_vars
maker.rsq
person
mafs
mafs = {}
# either use privided MAF or computer MAF
if all(vcf_mafs):
print('run this')
for idx, v in enumerate(varnames[item]):
if v not in mafs:
mafs[v] = vcf_mafs[idx]
else:
# count founder alleles
for hap in haplotypes[item]:
if not data.tfam.is_founder(hap[1]):
continue
for idxv, v in enumerate(varnames[item]):
if v not in mafs:
# [#alt, #haplotypes]
mafs[v] = [0, 0]
gt = hap[2 + idxv][1] if hap[2 + idxv][0].isupper() else hap[2 + idxv][0]
if not gt == "?":
mafs[v][0] += self.gtconv[gt]
mafs[v][1] += 1.0
mafs
vcf_mafs
type(mafs['V0-176659933'])
maker.maf_cutoff
exclude_vars = []
for v in mafs.keys():
if mafs[v] > maker.maf_cutoff:
exclude_vars.append(v)
for i in haplotypes.keys():
haplotypes[i] = listit(haplotypes[i])
for j in range(len(haplotypes[i])):
haplotypes[i][j] = haplotypes[i][j][:2] + \
[x for idx, x in enumerate(haplotypes[i][j][2:]) if varnames[i][idx] not in exclude_vars]
varnames[i] = [x for x in varnames[i] if x not in exclude_vars]
# handle trivial data
if len(varnames[i]) == 0:
for person in data.families[i]:
data[person] = self.missings
del varnames[i]
del haplotypes[i]
tmp_exclude_vars=exclude_vars
tmp_exclude_vars
recombPos
uniq_vars = []
i = '1'
for tmp_var in varnames[i]:
if tmp_var not in uniq_vars:
uniq_vars.append(tmp_var)
varnames[i] = [x for x in varnames[i] if x not in tmp_exclude_vars]
data
data.genotype_all
varnames
if len(varnames):
if not any ([len(varnames[x]) - 1 for x in varnames]):
# all families have only one variant
maker._MarkerMaker__AssignSNVHaplotypes(data, haplotypes, mafs, varnames)
else:
print('run this')
# calculate LD clusters using founder haplotypes
clusters = maker._MarkerMaker__ClusterByLD(data, haplotypes, varnames)
# recoding the genotype of the region
maker._MarkerMaker__CodeHaplotypes(data, haplotypes, mafs, varnames, clusters)
clusters
# ## def __ClusterByLD(self, data, haplotypes, varnames):
haplotypes
maker.r2
markers
# get founder haplotypes
gtt = []
founder_haplotypes = []
markers = sorted(set(itertools.chain(*varnames.values())), key = lambda x: int(x.split("-")[0][1:]))
for item in haplotypes:
for ihap, hap in enumerate(haplotypes[item]):
if not data.tfam.is_founder(hap[1]):
continue
gt = [hap[2 + varnames[item].index(v)] if v in varnames[item] else '?' for v in markers]
founder_haplotypes.append(("{}-{}".format(hap[1], ihap % 2), "".join([x[1] if x[0].isupper() else x[0] for x in gt])))
gtt.append(["{}-{}".format(hap[1], ihap % 2), [x[1] if x[0].isupper() else x[0] for x in gt]])
founder_haplotypes
egglib.stats.matrix_LD(Align.create(founder_haplotypes,egglib.Alphabet(cat='string',expl=['1','2'],miss='?')),('rsq'))
gtt
import egglib
gt
ldi,ld = egglib.stats.matrix_LD(Align.create(gtt,egglib.Alphabet(cat='char',expl=['1','2'],miss='?')),('rsq'))
blocks = []
for j in range(len(ldi)):
block = [j]
for k in range(j+1,len(ldi)):
if ld[k][j] > maker.r2:
block.append(k)
if len(block) > 1:
blocks.append(block)
clusters = [[markers[idx] for idx in item] for item in list(connected_components(blocks))]
clusters
varnames
# calculate LD blocks, use r2 measure
ld = Align.create(founder_haplotypes).matrixLD(validCharacters="12")["r2"]
blocks = []
for j in ld:
block = [j]
for k in ld[j]:
if ld[j][k] > maker.r2:
block.append(k)
if len(block) > 1:
blocks.append(block)
# get LD clusters
clusters = [[markers[idx] for idx in item] for item in list(connected_components(blocks))]
ld
block
list(connected_components([]))
list(connected_components(blocks))
clusters
# ## def __CodeHaplotypes(self, data, haplotypes, mafs, varnames, clusters):
# apply CHP coding
if clusters is not None:
clusters_idx = [[[varnames[item].index(x) for x in y] for y in clusters] for item in haplotypes]
else:
clusters_idx = [[[]] for item in haplotypes]
clusters_idx
maker.coder.Execute(haplotypes.values(), [[mafs[v] for v in varnames[item]] for item in haplotypes], clusters_idx)
data.superMarkerCount
# line: [fid, sid, hap1, hap2]
for line in maker.coder.GetHaplotypes():
print(line)
if not line[1] in data:
# this sample is not in VCF file. Every variant site should be missing
# they have to be skipped for now
continue
data[line[1]] = (line[2].split(','), line[3].split(','))
if len(data[line[1]][0]) > data.superMarkerCount:
data.superMarkerCount = len(data[line[1]][0])
data
data.superMarkerCount
# get MAF
for item in haplotypes:
data.maf[item] = maker.coder.GetAlleleFrequencies(item)
data.maf[item] = tuple(tuple(np.array(v) / np.sum(v)) if np.sum(v) else v
for v in data.maf[item])
data.maf
maker._MarkerMaker__FormatHaplotypes(data,recombPos,varnames,uniq_vars)
data
type(data['I:1'][0])
# apply CHP coding
for item in data.famvaridx:
if item not in haplotypes and data[data.families[item][0]] != ('0','0'):
# when only wild-type haplotypes are present in a family, still code them instead of ignoring the family
if self.freq_by_fam:
pop=data.freq_by_fam[item]
try:
varnames[item]=data.total_varnames[pop]
mafs[item]=data.total_mafs[pop]
except:
continue
else:
varnames[item]=data.total_varnames['pop']
mafs[item]=data.total_mafs
haplotypes[item]=[]
for person in data.families[item]:
tmp_person=[item, person]
if '00' in data[person]:
tmp_person+=['?:']*len(varnames[item])
else:
tmp_person+=['1:']*len(varnames[item])
haplotypes[item].append(tmp_person)
haplotypes[item].append(tmp_person)
elif item in haplotypes:
nonvar_hap_flag=False
#determine if wild-type haplotype is present in a family
for hap in haplotypes[item]:
tmp_genes=[]
for tmpa in hap[2:]:
if 'A' in tmpa or 'B' in tmpa:
tmp_genes.append(tmpa[1])
else:
tmp_genes.append(tmpa[0])
if set(tmp_genes)==set(['1']):
#non variant haplotype
nonvar_hap_flag=True
break
if not nonvar_hap_flag:
#if family don't have wild-type haplotype, add a fake one to ensure correct coding
var_num=len(varnames[item])
fake_person=[item, 'FAKEPERSON']+['1:']*var_num
haplotypes[item].append(fake_person)
for hidx,hap in enumerate(haplotypes[item]):
if hap[1] in data.missing_persons:
missing_person=[item,hap[1]]+['?:']*len(varnames[item])
haplotypes[item][hidx]=missing_person
if not clusters is None:
clusters_idx = [[[varnames[item].index(x) for x in y] for y in clusters] for item in haplotypes]
else:
clusters_idx = [[[]] for item in haplotypes]
if env.debug:
for item in haplotypes:
with env.lock:
print(varnames[item],file=sys.stderr)
print("hap{0}\t{1}\n".format(item,haplotypes[item]),file=sys.stderr)
self.coder.Execute(haplotypes.values(), [[mafs[item][v] for v in varnames[item]] for item in haplotypes], clusters_idx)
if env.debug:
with env.lock:
if clusters:
print("Family LD clusters: ", clusters_idx, "\n", file = sys.stderr)
self.coder.Print()
# line: [fid, sid, hap1, hap2]
for line in self.coder.GetHaplotypes():
if not line[1] in data:
# this sample is not in VCF file. Every variant site should be missing
# they have to be skipped for now
continue
data[line[1]] = (line[2].split(','), line[4].split(','))
#sub-region count for each sample individual
superMarkerCount=len(data[line[1]][0])
if line[0] not in data.patterns:
data.patterns[line[0]]=[[] for x in range(superMarkerCount)]
for t_Marker in range(superMarkerCount):
t_pat1=line[3].split(',')[t_Marker]
t_pat2=line[5].split(',')[t_Marker]
if t_pat1 not in data.patterns[line[0]][t_Marker]:
data.patterns[line[0]][t_Marker].append(t_pat1)
if t_pat2 not in data.patterns[line[0]][t_Marker]:
data.patterns[line[0]][t_Marker].append(t_pat2)
if len(data[line[1]][0]) > data.superMarkerCount:
data.superMarkerCount = len(data[line[1]][0])
# get MAF
for item in data.famvaridx:
if item not in haplotypes:
for person in data.families[item]:
data[person]=(['0']*data.superMarkerCount,['0']*data.superMarkerCount)
for item in haplotypes:
data.maf[item] = self.coder.GetAlleleFrequencies(item)
if not len(data.maf[item][0]):
continue
data.varnames_by_fam[item]=varnames[item]
wt_maf=0
if self.freq_by_fam:
try:
wt_maf=data.wt_maf[data.freq_by_fam[item]]
except:
pass
else:
wt_maf=data.wt_maf['pop']
tmp_data_maf=[]
for v in data.maf[item]:
if len(v)==1:
tmp_data_maf.append((v[0],1-v[0]))
else:
if np.sum(v)<1:
tmp_ratio=sum(v[1:])/(1-wt_maf)
tmp_list=[wt_maf]
if tmp_ratio==0:
tmp_list.append(1-wt_maf)
else:
for tmpv in v[1:]:
tmp_list.append(tmpv/tmp_ratio)
tmp_data_maf.append(tuple(tmp_list))
else:
tmp_data_maf.append(v)
data.maf[item]=tuple(tmp_data_maf)
if env.debug:
with env.lock:
print("marker freqs = ", data.maf, "\n", file = sys.stderr)
for item in data.families:
varnames[item], positions, vcf_mafs = data.getFamVariants(item, style = "map")
if len(varnames[item]) == 0:
for person in data.families[item]:
data[person] = self.missings
continue
if env.debug:
with env.lock:
sys.stderr.write('\n'.join(['\t'.join(x) for x in data.getFamSamples(item)]) + '\n\n')
# haplotyping
with env.lock:
if not env.prephased:
with stdoutRedirect(to = env.tmp_log + str(os.getpid()) + '.log'):
haplotypes[item] = self.haplotyper.Execute(data.chrom, varnames[item],
sorted(positions), data.getFamSamples(item))[0]
else:
haplotypes[item] = self.__PedToHaplotype(data.getFamSamples(item))
if len(haplotypes[item]) == 0:
# C++ haplotyping implementation failed
with env.chperror_counter.get_lock():
env.chperror_counter.value += 1
# either use privided MAF or computer MAF
if all(vcf_mafs):
for idx, v in enumerate(varnames[item]):
if v not in mafs:
mafs[v] = vcf_mafs[idx]
else:
# count founder alleles
for hap in haplotypes[item]:
if not data.tfam.is_founder(hap[1]):
continue
for idxv, v in enumerate(varnames[item]):
if v not in mafs:
# [#alt, #haplotypes]
mafs[v] = [0, 0]
gt = hap[2 + idxv][1] if hap[2 + idxv][0].isupper() else hap[2 + idxv][0]
if not gt == "?":
mafs[v][0] += self.gtconv[gt]
mafs[v][1] += 1.0
maker.haplotyper
tmp1
aa = []
for _ in range(10):
a = queue.get()
print(a)
tmp.getRegion(a)
tmp.apply(dd)
tmp1.apply(dd)
#tmp2.apply(dd)
if len(dd.variants) != 0:
aa.append(a)
data1 = deepcopy(data)
# ## _MarkerMaker__Haplotype
data = dd
haplotypes = OrderedDict()
mafs = {} ##Per fam per variant
uniq_vars = []
exclude_vars = []
varnames = {}
recombPos = {}
tmp1.markers
# + active=""
# def __Haplotype(self, data, haplotypes, mafs, varnames,recombPos,uniq_vars,exclude_vars):
# '''genetic haplotyping. haplotypes stores per family data'''
# # FIXME: it is SWIG's (2.0.12) fault not to properly destroy the object "Pedigree" in "Execute()"
# # So there is a memory leak here which I tried to partially handle on C++
# #
# # Per family haplotyping
# #
# -
tmp1.markers = ["V{}-{}".format(idx, item[1]) for idx, item in enumerate(data.variants)]
tmp_mafs = {}
if tmp1.freq_by_fam:
## if families are from different populations
## estimate MAF by different population
fam_to_analyze={}
for fam,pop in data.freq_by_fam.iteritems():
if pop not in fam_to_analyze:
fam_to_analyze[pop]=[fam]
else:
fam_to_analyze[pop].append(fam)
if tmp1.count:
## estimate MAF by counting founder alleles
if tmp1.freq_by_fam:
local_count_mafs={}
for pop in fam_to_analyze:
local_count_mafs[pop]=tmp1._MarkerMaker__computefounderfreq(data,fam_to_analyze[pop])
else:
local_count_mafs=tmp1._MarkerMaker__computefounderfreq(data,data.families.keys())
print('run here')
local_count_mafs
tmp1.mle = True
if tmp1.mle:
## estimate MLE allele frequency using all fam
local_mle_mafs={}
if tmp1.freq_by_fam:
for pop in fam_to_analyze:
local_mle_mafs[pop]={}
markers_to_analyze=[]
pos_all=[]
markers_analyzed={}
if pop not in data.mle_mafs:
data.mle_mafs[pop]={}
else:
for tmpv in data.mle_mafs[pop]:
markers_analyzed[tmpv.split('-')[-1]]=data.mle_mafs[pop][tmpv]
output_log=env.tmp_log+"AF_{}_{}.log".format(pop,tmp1.name)
popidx=tmp1.af_info.index(pop)
variants_in_fams=[]
for item in fam_to_analyze[pop]:
for tmpvar in data.getFamVariants(item):
if tmpvar not in variants_in_fams:
variants_in_fams.append(tmpvar)
variants_in_fams=sorted(variants_in_fams, key=lambda x: x[1])
for item in variants_in_fams:
idx=data.variants.index(item)
if item[-1][popidx]==0:
if str(item[1]) in markers_analyzed.keys():
#if variant has been analyzed
vname="V{}-{}".format(idx,item[1])
local_mle_mafs[pop][vname]=markers_analyzed[str(item[1])]
else:
#variant not analyzed before
markers_to_analyze.append("V{}-{}".format(idx,item[1]))
pos_all.append(item[1])
tmp_mle_mafs=tmp1._MarkerMaker__getMLEfreq(data, markers_to_analyze, pos_all, fam_to_analyze[pop], tmp1.rsq, output_log)
if len(tmp_mle_mafs) > 0:
for vname,vmaf in tmp_mle_mafs.iteritems():
data.mle_mafs[pop][vname]=vmaf
local_mle_mafs[pop][vname]=vmaf
else:
#Homogeneous families
markers_to_analyze=[]
pos_all=[]
markers_analyzed={}
for tmpv in data.mle_mafs:
markers_analyzed[tmpv.split('-')[-1]]=data.mle_mafs[tmpv]
variants_in_fams=[]
for item in data.families.keys():
var_per_fam=[tuple(tmpvar) for tmpvar in data.getFamVariants(item)]
variants_in_fams=list(set(var_per_fam+variants_in_fams))
variants_in_fams=[list(tmpvar) for tmpvar in sorted(variants_in_fams, key=lambda x: x[1])]
for item in variants_in_fams:
idx=data.variants.index(item)
if item[-1]==0 or tmp1.af_info is None:
if str(item[1]) in markers_analyzed.keys():
#if variant has been analyzed
vname="V{}-{}".format(idx,item[1])
local_mle_mafs[vname]=markers_analyzed[str(item[1])]
else:
#variant not analyzed before
markers_to_analyze.append("V{}-{}".format(idx,item[1]))
pos_all.append(item[1])
output_log=env.tmp_log+"AF_{}.log".format(tmp1.name)
tmp_mle_mafs=tmp1._MarkerMaker__getMLEfreq(data, markers_to_analyze, pos_all, data.families.keys(), tmp1.rsq, output_log)
if len(tmp_mle_mafs) > 0:
for vname, vmaf in tmp_mle_mafs.iteritems():
data.mle_mafs[vname]=vmaf
local_mle_mafs[vname]=vmaf
tmp_mle_mafs
varnames
data.families
data.families.keys()[0]
data.getFamVariants(data.families.keys()[1],style="map")
data.famvaridx
data.famsampidx
# +
gnomAD_pop=None
for item in data.families:
varnames[item], positions, vcf_mafs = data.getFamVariants(item, style = "map")
recombPos[item]={}
var_for_haplotype=[]
positions_for_haplotype=[]
output_sample=[]
if env.debug:
with env.lock:
sys.stderr.write('\n'+repr(varnames[item])+'\n')
sys.stderr.write('\n'.join(['\t'.join(x) for x in data.getFamSamples(item)]) + '\n\n')
# either use privided MAF or compute MAF
if tmp1.freq_by_fam:
mafs[item]={}
tfreq_fam=data.freq_by_fam[item]
for pop in data.gnomAD_estimate.keys():
if pop in tfreq_fam:
gnomAD_pop=pop
break
elif gnomAD_pop is None and data.freq is not None:
for pop in data.gnomAD_estimate.keys():
if pop in data.freq:
gnomAD_pop=pop
break
for idx, v in enumerate(varnames[item]):
tmp_maf_var=0
if tmp1.af_info is None:
#no vcf freq column specified
if v not in tmp_mafs:
if tmp1.mle:
#use MLE freq for all variants
tmp_mafs[v]=local_mle_mafs[v]
elif tmp1.count:
#estimate MAF based on founder counts if MLE not specified
tmp_mafs[v]=local_count_mafs[v]
tmp_maf_var=tmp_mafs[v]
elif not tmp1.af_info is None:
#if vcf freq column is specified
#use vcf_mafs if possible
if vcf_mafs[idx]:
tmp_maf_var=vcf_mafs[idx]
if tmp1.freq_by_fam:
mafs[item][v] = vcf_mafs[idx]
else:
if v not in tmp_mafs:
tmp_mafs[v] = vcf_mafs[idx]
else:
#if variants do not have valid vcf_mafs values if specified
if tmp1.freq_by_fam:
if gnomAD_pop is not None:
mafs[item][v]=data.gnomAD_estimate[gnomAD_pop]
elif tmp1.mle:
mafs[item][v]=local_mle_mafs[data.freq_by_fam[item]][v]
elif tmp1.count:
mafs[item][v]=local_count_mafs[data.freq_by_fam[item]][v]
tmp_maf_var=mafs[item][v]
else:
if v not in tmp_mafs:
if gnomAD_pop is not None:
tmp_mafs[v]=data.gnomAD_estimate[gnomAD_pop]
elif tmp1.mle:
tmp_mafs[v]=local_mle_mafs[v]
elif tmp1.count:
tmp_mafs[v]=local_count_mafs[v]
tmp_maf_var=tmp_mafs[v]
if tmp1.rvhaplo:
if tmp_maf_var<=tmp1.maf_cutoff:
var_for_haplotype.append(v)
positions_for_haplotype.append(positions[idx])
if not tmp1.rvhaplo:
var_for_haplotype=varnames[item]
positions_for_haplotype=positions
#collect sample+genotypes
for person in data.tfam.sort_family(item):
output_sample.append([])
last_ele=len(output_sample)-1
output_sample[last_ele] = data.tfam.samples[person][:-1]
if person in data.samples:
for marker in var_for_haplotype:
idx=int(marker.split('-')[0][1:])
output_sample[last_ele].append(data.genotype_all[person][idx])
else:
output_sample[last_ele].extend(["00"] * len(var_for_haplotype))
# haplotyping
if len(var_for_haplotype)==0:
varnames.pop(item,None)
#for person in data.families[item]:
# data[person] = tmp1.missings
continue
for person in output_sample:
if set(person[5:])==set(['00']):
data.missing_persons.append(person[1])
with env.lock:
if not env.prephased:
tmp_log_output=env.tmp_log + str(os.getpid())
with stdoutRedirect(to = tmp_log_output + '.log'):
haplotypes[item] = tmp1.haplotyper.Execute(data.chrom, var_for_haplotype, positions_for_haplotype, output_sample, tmp1.rsq, tmp_log_output)[0]
else:
haplotypes[item] = tmp1.__PedToHaplotype(data.getFamSamples(item))
if len(haplotypes[item]) == 0:
# C++ haplotyping implementation failed
with env.chperror_counter.get_lock():
env.chperror_counter.value += 1
varnames[item]=var_for_haplotype
for item in haplotypes:
for hap_idx,haploid in enumerate(haplotypes[item]):
for vidx,var in enumerate(haploid[2:]):
if not var.endswith(':') and not var.endswith('|') and vidx!=0:
postvar_name=varnames[item][vidx]
prevar_name=varnames[item][vidx-1]
recomb_pair = (prevar_name,postvar_name)
try:
recombPos[item][recomb_pair].append(hap_idx)
except:
recombPos[item][recomb_pair]=[hap_idx]
#
# Compute founder MAFs
#
if len(tmp_mafs) > 0:
if tmp1.freq_by_fam:
for pop in tmp_mafs:
for v in tmp_mafs[pop]:
if type(tmp_mafs[pop][v]) is list:
tmp_mafs[pop][v] = tmp_mafs[pop][v][0]/tmp_mafs[pop][v][1] if tmp_mafs[pop][v][1] >0 else 0.0
else:
for v in tmp_mafs:
if type(tmp_mafs[v]) is list:
tmp_mafs[v] = tmp_mafs[v][0]/tmp_mafs[v][1] if tmp_mafs[v][1] > 0 else 0.0
## Make mafs consistent in structure regardless of freq_by_fam
if tmp1.freq_by_fam:
for item in haplotypes:
popname=data.freq_by_fam[item]
if popname not in tmp_mafs:
continue
if item not in mafs:
mafs[item]=tmp_mafs[popname]
else:
for v in tmp_mafs[popname]:
if v not in mafs[item]:
mafs[item][v]=tmp_mafs[popname][v]
else:
for item in haplotypes:
mafs[item]=tmp_mafs
if env.debug:
with env.lock:
print("variant mafs = ", mafs, "\n", file = sys.stderr)
##
#
# Drop some variants if maf is greater than given threshold
#
if not tmp1.maf_cutoff is None or tmp1.single_markers:
if tmp1.freq_by_fam:
exclude_vars=[[] for x in range(len(data.freq))]
for i in haplotypes.keys():
if tmp1.freq_by_fam:
pop_idx=data.freq.index(data.freq_by_fam[i])
tmp_exclude_vars=exclude_vars[pop_idx]
else:
tmp_exclude_vars=exclude_vars
for v in mafs[i].keys():
if not tmp1.maf_cutoff is None:
if mafs[i][v] > tmp1.maf_cutoff and v not in tmp_exclude_vars or v.split('-')[-1] not in data.include_vars:
tmp_exclude_vars.append(v)
if tmp1.single_markers:
if v.split('-')[-1] not in data.include_vars:
tmp_exclude_vars.append(v)
haplotypes[i] = listit(haplotypes[i])
tmp_remain_vars=[x for x in varnames[i] if x not in tmp_exclude_vars]
recomb_remain_vars=[]
if len(tmp_remain_vars) == 0:
recombPos[i]={}
else:
if len(recombPos[i]) > 0:
#extend recombination signal to neighbouring RVs
#if the original variant is to be excluded
#Only allow a maximum of one recombination event between one pair of consecutive markers
for pair in recombPos[i].keys():
if pair[1] not in tmp_exclude_vars:
if tmp_remain_vars.index(pair[1])!=0 and pair[1] not in recomb_remain_vars:
recomb_remain_vars.append(pair[1])
else:
del recombPos[i][pair]
else:
if varnames[i].index(pair[1]) > varnames[i].index(tmp_remain_vars[-1]):
#last variant
del recombPos[i][pair]
continue
for tmp_idx in range(varnames[i].index(pair[1])+1,len(varnames[i])):
if varnames[i][tmp_idx] not in tmp_exclude_vars:
if tmp_remain_vars.index(varnames[i][tmp_idx])==0:
#delete recombination pair if the recombination was marked to the first remaining variant
del recombPos[i][pair]
break
for tmp_hap in recombPos[i][pair]:
tmp_var=haplotypes[i][tmp_hap][tmp_idx+2]
if tmp_var.endswith(':') or tmp_var.endswith('|'):
haplotypes[i][tmp_hap][tmp_idx+2]=tmp_var[:-1]+'/'
if varnames[i][tmp_idx] not in recomb_remain_vars:
recomb_remain_vars.append(varnames[i][tmp_idx])
else:
del recombPos[i][pair]
break
for j in range(len(haplotypes[i])):
haplotypes[i][j] = haplotypes[i][j][:2] + \
[x for idx, x in enumerate(haplotypes[i][j][2:]) if varnames[i][idx] not in tmp_exclude_vars]
for tmp_var in varnames[i]:
if tmp_var not in uniq_vars:
uniq_vars.append(tmp_var)
varnames[i] = [x for x in varnames[i] if x not in tmp_exclude_vars]
# handle trivial data
if len(varnames[i]) == 0:
del varnames[i]
del haplotypes[i]
if len(recombPos[i].keys())>tmp1.recomb_max:
#treat as missing if recombination events occurred more than speicified times
recombPos[i]={}
for person in data.families[i]:
data[person] = tmp1.missings
del varnames[i]
del haplotypes[i]
# count how many variants are removed
with env.commonvar_counter.get_lock():
if tmp1.freq_by_fam:
tmp_ex_vars=[tmp_var for tmp_vars in exclude_vars for tmp_var in tmp_vars]
env.commonvar_counter.value += len(set(tmp_ex_vars))
else:
env.commonvar_counter.value += len(exclude_vars)
# get total observed variants
if tmp1.freq_by_fam:
for item in varnames:
pop=data.freq_by_fam[item]
if pop not in data.total_mafs:
data.total_mafs[pop]={}
data.total_varnames[pop]=[]
for v in varnames[item]:
if v not in data.total_mafs[pop]:
data.total_varnames[pop].append(v)
data.total_mafs[pop][v]=mafs[item][v]
for pop in data.total_varnames:
data.total_varnames[pop]=sorted(data.total_varnames[pop], key=lambda x: int(x.split("-")[0][1:]))
data.wt_maf[pop]=1.0
for v,tmaf in data.total_mafs[pop].iteritems():
data.wt_maf[pop]*=(1-tmaf)
else:
data.total_varnames['pop']=[]
for item in varnames:
for v in varnames[item]:
if v not in data.total_mafs:
data.total_varnames['pop'].append(v)
data.total_mafs[v]=mafs[item][v]
data.wt_maf['pop']=1.0
for v,tmaf in data.total_mafs.iteritems():
data.wt_maf['pop']*=(1-tmaf)
data.total_varnames['pop']=sorted(data.total_varnames['pop'], key=lambda x: int(x.split("-")[0][1:]))
# -
data.total_varnames
??cstatgen.HaplotypingEngine()
data.wt_maf
aa = []
for _ in range(1000):
a = queue.get()
tmp.getRegion(a)
tmp.apply(dd)
if len(dd.variants) != 0:
aa.append(a)
len(aa)
tmp.getRegion(aa[0])
dd = deepcopy(data)
tmp.apply(dd)
tmp1.apply(dd)
tmp1._MarkerMaker__Haplotype
tmp1.freq_by_fam
tmp1.count
tmp1.mle
tmp1.rvhaplo
tmp1.apply(dd)
tmp2.apply(dd)
tmp1.freq_by_fam
tmp1.count
env.prephased
env.tmp_log
haplotypes = OrderedDict()
mafs = {} ##Per fam per variant
uniq_vars = []
exclude_vars = []
varnames = {}
recombPos = {}
tmp1._MarkerMaker__Haplotype(dd, haplotypes, mafs, varnames,recombPos,uniq_vars,exclude_vars)
tm
tmp1.recomb_perfam
tmp1.apply(dd)
tmp2.apply(dd)
data.variants
dd.include_vars
dd.variants
tmp1.getRegion(aa[0])
tmp1.apply(dd)
env.debug =True
data.variants
dd.chrom
dd.variants
tmp.vcf
??EncoderWorker
env.total_counter.value
jobs
for j in jobs:
j.start()
for j in jobs:
j.join()
faulthandler.disable()
try:
faulthandler.enable(file=open(env.tmp_log + '.SEGV', 'w'))
for i in regions:
if isinstance(queue, list):
queue.append(i)
else:
queue.put(i)
freq_by_fam_flag = False
if not args.freq_by_fam is None:
print('haha')
freq_by_fam_flag = True
with open(args.freq_by_fam) as freq_fh:
for freq_line in freq_fh:
tmp_eles=freq_line.split() #Fam and Population
data.freq_by_fam[tmp_eles[0]]=tmp_eles[1]
data.freq=sorted(list(set(data.freq_by_fam.values())))
else:
data.freq=args.freq
jobs = [EncoderWorker(
queue, len(regions), deepcopy(data),
RegionExtractor(args.vcf, chr_prefix = args.chr_prefix, allele_freq_info = data.freq, include_vars_file=args.include_vars),
MarkerMaker(args.bin, maf_cutoff = args.maf_cutoff,single_markers=args.single_markers,recomb_max=args.recomb_max,af_info=data.freq,freq_by_fam=freq_by_fam_flag,rsq=args.rsq,mle=args.mle, rvhaplo=args.rvhaplo, recomb_perfam=not args.recomb_cross_fam),
LinkageWriter(len(samples_not_vcf))
) for i in range(env.jobs)]
for j in jobs:
j.start()
for j in jobs:
j.join()
faulthandler.disable()
except KeyboardInterrupt:
# FIXME: need to properly close all jobs
raise ValueError("Use 'killall {}' to properly terminate all processes!".format(env.prog))
else:
env.log('{:,d} units (from {:,d} variants) processed; '\
'{:,d} Mendelian inconsistencies and {:,d} recombination events handled\n'.\
format(env.success_counter.value,
env.variants_counter.value,
env.mendelerror_counter.value,
env.recomb_counter.value), flush = True)
if env.triallelic_counter.value:
env.log('{:,d} tri-allelic loci were ignored'.format(env.triallelic_counter.value))
if env.commonvar_counter.value:
env.log('{:,d} variants ignored due to having MAF > {} and other specified constraints'.\
format(env.commonvar_counter.value, args.maf_cutoff))
if env.null_counter.value:
env.log('{:,d} units ignored due to absence in VCF file'.format(env.null_counter.value))
if env.trivial_counter.value:
env.log('{:,d} units ignored due to absence of variation in samples'.format(env.trivial_counter.value))
fatal_errors = 0
try:
# Error msg from C++ extension
os.system("cat {}/*.* > {}".format(env.tmp_dir, env.tmp_log))
fatal_errors = wordCount(env.tmp_log)['fatal']
except KeyError:
pass
if env.chperror_counter.value:
env.error("{:,d} regional markers failed to be generated due to haplotyping failures!".\
format(env.chperror_counter.value))
if fatal_errors:
env.error("{:,d} or more regional markers failed to be generated due to runtime errors!".\
format(fatal_errors))
env.log('Archiving regional marker data to directory [{}]'.format(env.cache_dir))
cache.write(arcroot = 'CACHE', source_dir = env.tmp_cache)
env.jobs = args.jobs
# STEP 2: write to PLINK or mega2 format
tpeds = [os.path.join(env.tmp_cache, item) for item in os.listdir(env.tmp_cache) if item.startswith(env.output) and item.endswith('.tped')]
for fmt in args.format:
cache.setID(fmt)
if not args.vanilla and cache.check():
env.log('Loading {} data from archive ...'.format(fmt.upper()))
cache.load(target_dir = env.tmp_dir, names = [fmt.upper()])
else:
env.log('{:,d} units will be converted to {} format'.format(env.success_counter.value, fmt.upper()))
env.format_counter.value = 0
format(tpeds, os.path.join(env.tmp_cache, "{}.tfam".format(env.output)),
args.prevalence, args.wild_pen, args.muta_pen, fmt,
args.inherit_mode, args.theta_max, args.theta_inc)
env.log('{:,d} units successfully converted to {} format\n'.\
format(env.format_counter.value, fmt.upper()), flush = True)
if env.skipped_counter.value:
# FIXME: perhaps we need to rephrase this message?
env.log('{} region - family pairs skipped'.\
format(env.skipped_counter.value))
env.log('Archiving {} format to directory [{}]'.format(fmt.upper(), env.cache_dir))
cache.write(arcroot = fmt.upper(),
source_dir = os.path.join(env.tmp_dir, fmt.upper()), mode = 'a')
mkpath(env.outdir)
if args.run_linkage:
cache.setID('analysis')
if not args.vanilla and cache.check():
env.log('Loading linkage analysis result from archive ...'.format(fmt.upper()))
cache.load(target_dir = env.output, names = ['heatmap'])
else:
env.log('Running linkage analysis ...'.format(fmt.upper()))
run_linkage(args.blueprint, args.theta_inc, args.theta_max, args.output_limit)
env.log('Linkage analysis succesfully performed for {:,d} units\n'.\
format(env.run_counter.value, fmt.upper()), flush = True)
if env.makeped_counter.value:
env.log('{} "makeped" runtime errors occurred'.format(env.makeped_counter.value))
if env.pedcheck_counter.value:
env.log('{} "pedcheck" runtime errors occurred'.format(env.pedcheck_counter.value))
if env.unknown_counter.value:
env.log('{} "unknown" runtime errors occurred'.format(env.unknown_counter.value))
if env.mlink_counter.value:
env.log('{} "mlink" runtime errors occurred'.format(env.mlink_counter.value))
cache.write(arcroot = 'heatmap', source_dir = os.path.join(env.output, 'heatmap'), mode = 'a')
html(args.theta_inc, args.theta_max, args.output_limit)
else:
env.log('Saving data to [{}]'.format(os.path.abspath(env.output)))
cache.load(target_dir = env.output, names = [fmt.upper() for fmt in args.format])
Args.get()
|
nbs/00_Core.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Continuous Control
# !pip -q install ./python
# !pip -q install tqdm
# +
from unityagents import UnityEnvironment
import numpy as np
from ddpg_agent import Agent
import matplotlib.pyplot as plt
# %matplotlib inline
from workspace_utils import active_session
# +
# select this option to load version 1 (with a single agent) of the environment
# env = UnityEnvironment(file_name='/data/Reacher_One_Linux_NoVis/Reacher_One_Linux_NoVis.x86_64')
# select this option to load version 2 (with 20 agents) of the environment
env = UnityEnvironment(file_name='/data/Reacher_Linux_NoVis/Reacher.x86_64')
# -
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
# +
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents
num_agents = len(env_info.agents)
print('Number of agents:', num_agents)
# size of each action
action_size = brain.vector_action_space_size
print('Size of each action:', action_size)
# examine the state space
states = env_info.vector_observations
state_size = states.shape[1]
print('There are {} agents. Each observes a state with length: {}'.format(states.shape[0], state_size))
print('The state for the first agent looks like:', states[0])
# -
agent = Agent(state_size=state_size, action_size=action_size, env=env, random_seed=1)
with active_session():
scores, average_scores = agent.train()
# Plot the scores
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(1, len(scores)+1), scores, label="Episode Scores")
plt.plot(np.arange(1, len(average_scores)+1), average_scores, label="Average Scores (over 100 episodes)")
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.legend()
plt.show()
env.close()
# Show the actor's architecture
print(agent.actor_local)
# Show the critic's architecture
print(agent.critic_local)
# # Project Report: Continuous Control
# For more details about how to execute the code, look at “Instructions” and “Files” sections in README.md.
#
# ## Learning Algorithm
# In this project, I used the Deep Deterministic Policy Gradients (DDPG) algorithm with 20 agents.
# The code itself is based on the DDPG with OpenAI Gym's BipedalWalker environment, from Udacity’s repo (https://github.com/udacity/deep-reinforcement-learning/tree/master/ddpg-bipedal).
# The agent consists of an Actor network, which takes the state as input and returns a deterministic action; and a Critic network which tries to approximate the value function and returns the action-value for the action taken by the actor.
#
# Furthermore, for each of the actor and critic there are actually two models, a main network and a target network. The target networks are used for calculating the error and are updated less often to avoid oscillations and make the learning more stable.
#
# The agent also makes use of experience replay (re-used from the code in a previous assignment), in order to mitigate issues with correlations between successive inputs.
#
# While the actor itself returns a deterministic action, some randomness is added to the output to encourage exploration. This random additive noise is generated through an Ornstein-Uhlenbeck process (also re-used from the code from Udacity repo), and is controlled via a decaying hyper-parameter epsilon, so as to encourage exploration initially and then rely more and more on exploitation as the models gets more training.
#
# As suggested in the “Benchmark Implementation” section of the course, I changed the code to update the local/main networks 10 times after every 20 timesteps, and used a “soft-update” for updating the target networks. However I did not have to clip the gradients.
#
# The training consist in running several episodes:
# 1) Starting from an initial state
# 2) Make the agent take an action (with additive noise for exploration)
# 3) Get the reward and next state for the taken action and add them to the replay buffer
# 4) After every 20 timesteps, for 10 times:
# - sample data from the replay buffer
# - perform a weight update for the local networks based on the expected and target action-values
# - Perform a “soft-update” is used to update the target network parameter every pre-defined number of steps.
# - When the requirement average score over 100 episodes at least 30 is satisfied, training is stopped and model weights are saved.
#
# ## Model Architecture
# The actor is a fully-connected network with 3 hidden layers and relu as an activation function, and batch normalization to normalize the input states and intermediate inputs to each layer. The normalization improves the learning because the entries in the 33-dimensional state vector have a different meaning and scale.
# The output is a 4-dimensional vector corresponding to the actions space size, with each action clipped to be in the range [-1, 1]
#
# The critic is a fully-connected network with 3 hidden layers and relu as an activation function, and batch normalization to normalize the input states and intermediate inputs to each layer.
# In addition to the state, the critic network also takes the action vector as an input. The resulting output is a scalar corresponding to the state-action value.
# See the above cells for the details.
#
# ## Hyperparameters
# The following were used:
# BUFFER_SIZE = int(1e6) # replay buffer size
# BATCH_SIZE = 128 # minibatch size
# GAMMA = 0.99 # discount factor
# TAU = 1e-3 # for soft update of target parameters
# LR_ACTOR = 1e-3 # learning rate of the actor
# LR_CRITIC = 1e-3 # learning rate of the critic
# WEIGHT_DECAY = 0 # L2 weight decay
# EPSILON_DECAY = 1e-6 # epsilon decay
# EPSILON = 1.0 # exploration actions space
# LEARN_INTERVAL = 20 # learning 10 times after every 20 timesteps
# LEARN_TIMES = 10
#
# As well as:
# fc1_units = fc2_units=512 # Size of hidden units for both actor and critic
#
# Furthermore, I used
# Adam optimizer as the optimizer
# Mean squared error (torch.nn.functional.mse_loss) as a loss function for the critic.
#
# The choice of the initial values for the hyperparameters was based on a previous assignment, and then verified that these values lead to good empirical results
#
# ## Plot of Rewards
# We see that after 125 episods, the agent reaches an average score of 30, the desired minimum, and the models’ weights are saved.
# See the figure above for the plot.
#
# ## Ideas for Future Work
#
# - I have actually started with 1-step TD actor-critic, but it did not work well and the reward was very low. My guess is that because of the online nature of the algorithm and no experience replay was used, the model was affected by correlations between consecutive inputs. Eventually DDPG worked better.
# - Another possibility for improvement is to further tune the hyperparameters. For this project I just selected them from previous Udacity assignments and got acceptable results. Doing the tuning may lead the model to fall in a better local minimum resulting in higher rewards, or use a different learning rate which could speed up the learning.
# - It’s worth trying other suggested algorithms like TRPO or D4PG
|
p2_continuous-control/Report.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import norm
import itertools
plt.rcParams["figure.figsize"] = (15, 15)
# %matplotlib inline
# -
# # Modeling the random walk
#
# We assume the following:
#
# 1) X and Y are independent
#
# 2) Each jump of the fire over given timestep t
#
# Our approach is to treat the spread of fire as a random walk on $x$ and $y$.
# For each step of the random walk, we'll treat the velocity as a random variable from a distribution $N(\mu, \sigma^2)$
#
# Our input data is a set of trajectories based off of the given tweets.
# Each trajectory gives us a set of jumps associated with the given fire of interest in our region.
#
# +
numtweets = 30
r = 50
variance = 5
binsize = 4
#x, y, and timestamp of each tweet
testx = np.linspace(0, r, numtweets) + np.random.normal(0,variance,numtweets)
testy = np.linspace(0, r, numtweets) + np.random.normal(0,variance,numtweets)
testt = np.arange(testx.shape[0])
bins = np.arange(min(testt), max(testt), binsize) #make some bins based on binsize
inds = np.digitize(testt, bins) #for each time in testt, assign it to a bin;
#the bin of element i in testt will be the bin for element i in testx and testy also
numbins = len(bins)
x_trajectories = []
y_trajectories = []
for i in range(numbins):
tempx = []
tempy = []
for j in range(len(testt)):
if inds[j] == i + 1: #since bin 0 is indexed as 1
tempx += [testx[j]] #if the jth element of testt is in bin i,
tempy += [testy[j]] #add testx[j] and testy[j] to their appropriate bins
x_trajectories += [tempx]
y_trajectories += [tempy]
combinatorial_x_trajectories = list(itertools.product(*x_trajectories))
combinatorial_y_trajectories = list(itertools.product(*y_trajectories))
x_distances = [np.diff(i) for i in combinatorial_x_trajectories]
y_distances = [np.diff(i) for i in combinatorial_y_trajectories]
x_distances = np.reshape(x_distances, -1)
y_distances = np.reshape(y_distances, -1)
vx = x_distances/binsize
vy = y_distances/binsize
# +
def MLE_mu(data):
n = len(data)
return 1/n * np.sum(data)
def MLE_sigma2(data):
n = len(data)
mu_hat = MLE_mu(data)
return 1/n * np.sum((data - mu_hat)**2)
# -
# Since the behavior of fire spread is uncertain, we assume each trajectory represents an equally likely path of the the fire.
# Based off of this, each link $A\to B$ gives us a "representative" sample of the underlying distribution of velocities, both of x and y.
#
# Therefore, the approach will to be to calculate the $v_x = d_x/t$ for each link of Tweets per trajectory, then to use these to calculate MLEs for the normal distribution governing the velocities of each hop.
#
# Once we have the normal distribution for these hops, we can use this to predict the probability that the fire has reached some point $A$ by some time $T$.
# There are two ways we can do this:
#
# 1) Set some timestep $t$ within the range of timesteps that we have seen in the data, and subdivide the desired segment into even chunks such that $\sum t_i = T$ and then add the normals such that the add up to the correct value.
# In this case, say we have $n$ chunks,
# Then, the probability that a fire has spread to at least to point $A$ is
# $$
# \begin{align}
# P\left(\sum_{i=1}^n x_i \geq A \right) &= P\left(\sum_{i=1}^n tv_i \geq A \right) \\
# &= P\left(\sum_{i=1}^n v_i \geq \frac{A}{t} \right)\\
# &= P\left(N(n\mu, n\sigma^2)\geq \frac{A}{t} \right)\\
# &= P\left(N(n\mu, n\sigma^2)\geq \frac{A}{T/n} \right)\\
# &= P\left(N(n\mu, n\sigma^2)\geq \frac{nA}{T} \right)\\
# &= P\left(N(\mu, \sigma^2)\geq \frac{A}{T} \right)
# \end{align}
# $$
#
# 2) Find the average velocity required to traverse the whole path in one go and find that probability.
# $$
# \begin{align}
# P(X \geq A) &= P\left(vT \geq A\right)\\
# &= P\left(v \geq \frac{A}{T}\right)\\
# &= P\left(N(\mu, \sigma^2) \geq \frac{A}{T}\right)\\
# \end{align}
# $$
#
# Let's apply these ideas below.
#
# First, calculate velocity components based off of the given data:
vx, vy
timesteps = np.diff(testt)
vx = np.diff(testx)/timesteps
vy = np.diff(testy)/timesteps
# Get MLE estimates for $v_x$, $v_y$
muhat_y, sigmahat_y = MLE_mu(vy), MLE_sigma2(vy)
muhat_x, sigmahat_x = MLE_mu(vx), MLE_sigma2(vx)
print("means, x and y: " , muhat_x, muhat_y)
print("variances, x and y: ", sigmahat_x, sigmahat_y)
vy_dist = norm(loc = muhat_y, scale = np.sqrt(sigmahat_y))
vx_dist = norm(loc = muhat_x, scale = np.sqrt(sigmahat_x))
predictx = [vx_dist.mean() * t for t in testt]
plt.scatter(testt, testx, label = "x test")
plt.scatter(testt, predictx, label = "x predict")
# plt.scatter(testt, testy, label = "y test")
plt.legend();
# +
trajectories = [
[t * vx_dist.rvs() for t in testt]
for i in range(5)
]
for t in trajectories:
plt.plot(testt, t)
# -
simulated_y_trajectories = []
for i in range(40):
simulated_position = [0]
last = 0
for t in testt[1:]:
last += vy_dist.rvs()
simulated_position += [last]
plt.plot(testt, simulated_position)
simulated_y_trajectories += [simulated_position]
simulated_x_trajectories = []
for i in range(40):
simulated_position = [0]
last = 0
for t in testt[1:]:
last += vx_dist.rvs()
simulated_position += [last]
plt.plot(testt, simulated_position)
simulated_x_trajectories += [simulated_position]
import seaborn as sns
simulated_x_trajectories_plot = np.reshape(simulated_x_trajectories, -1)
simulated_y_trajectories_plot = np.reshape(simulated_y_trajectories, -1)
plt.figure(figsize = (10, 10))
sns.kdeplot(simulated_x_trajectories_plot, simulated_y_trajectories_plot, shade = True)
for x_traj, y_traj in zip(simulated_x_trajectories, simulated_y_trajectories):
plt.plot(x_traj, y_traj)
plt.scatter(testx, testy, label = "test data", color = 'orange')
plt.legend();
# # What happens with multiple fires?
# Inclusion exclusion:
# $$
# \begin{align}
# P(fire @ A @ t) &= P(fire 1 @ A @ t) + P(fire 2 @ A @ t) - P(fire 1 \cap fire 2 @ A @ t)\\
# &= P(fire 1 @ A @ t) + P(fire 2 @ A @ t) - P(fire 1 @ A @ t)P(fire 1 @ A @ t)
# \end{align}
# $$
# for time $t$ and point of interest $A$
|
randomwalk.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Load packages
import pandas as pd
from sklearn.cross_validation import train_test_split
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.naive_bayes import MultinomialNB
from sklearn.metrics import classification_report
from sklearn.metrics import accuracy_score
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
from sklearn.metrics import f1_score
# +
# Load MS Excel file
dataset=pd.read_excel('matrix.xlsx')
dataset=dataset[['tag','Filename','MDA']]
# Split the dataset in train/test ratio: 0.20
train_set, test_set = train_test_split(dataset, test_size = 0.20)
# +
# Baseline all BUY
buy=[]
for i in range(0, len(test_set)):
buy.append(1)
allBuy = pd.Series(buy)
# Print accuracy, precision, recall, F measure
tab = pd.crosstab(test_set_pred, test_set.tag, rownames=['Predicted'], colnames=['Actual'], margins=True) # Print confusion matrix
print(tab)
classification_report(allBuy, test_set.tag)
a=accuracy_score(allBuy, test_set.tag)
p=precision_score(allBuy, test_set.tag)
r=recall_score(allBuy, test_set.tag)
f=f1_score(allBuy, test_set.tag)
print "Accuracy = ",a,"\nPrecision =",p,"\nRecall = ",r,"\nF-Score = ",f
# +
# Create the Count classifier
vectorizer = CountVectorizer(stop_words="english")
counts = vectorizer.fit_transform(train_set.MDA.values)
classifier = MultinomialNB(fit_prior="False")
# Train the classifier
classifier.fit(counts, train_set.tag)
# Test the classifier
predictions = classifier.predict(vectorizer.transform(test_set.MDA.values))
test_set_pred = pd.Series(predictions, index=test_set.index)
tab = pd.crosstab(test_set_pred, test_set.tag, rownames=['Predicted'], colnames=['Actual'], margins=True) # Print confusion matrix
print(tab)
# Print accuracy, precision, recall, F measure
classification_report(test_set_pred, test_set.tag)
a=accuracy_score(test_set_pred, test_set.tag)
p=precision_score(test_set_pred, test_set.tag)
r=recall_score(test_set_pred, test_set.tag)
f=f1_score(test_set_pred, test_set.tag)
print "Accuracy = ",a,"\nPrecision =",p,"\nRecall = ",r,"\nF-Score = ",f
# +
# Create the TFIDF classifier
vectorizer = TfidfVectorizer(stop_words="english")
counts = vectorizer.fit_transform(train_set.MDA.values)
classifier = MultinomialNB(fit_prior="False")
# Train the classifier
classifier.fit(counts, train_set.tag)
# Test the classifier
predictions = classifier.predict(vectorizer.transform(test_set.MDA.values))
test_set_pred = pd.Series(predictions, index=test_set.index)
tab = pd.crosstab(test_set_pred, test_set.tag, rownames=['Predicted'], colnames=['Actual'], margins=True) # Print confusion matrix
print(tab)
# Print accuracy, precision, recall, F measure
classification_report(test_set_pred, test_set.tag)
a=accuracy_score(test_set_pred, test_set.tag)
p=precision_score(test_set_pred, test_set.tag)
r=recall_score(test_set_pred, test_set.tag)
f=f1_score(test_set_pred, test_set.tag)
print "Accuracy = ",a,"\nPrecision =",p,"\nRecall = ",r,"\nF-Score = ",f
|
FinancialAnalystV4/BasicModels.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 ('my-env')
# language: python
# name: python3
# ---
# +
# Setup
import pandas as pd
import numpy as np
# -
# # Data
# # Import
# +
ROOT = "https://raw.githubusercontent.com/kirenz/modern-statistics/main/data/"
DATA = "loans.csv"
df = pd.read_csv(ROOT+DATA)
# -
# ## Overview
df
print(f"We have {len(df.index):,} observations and {len(df.columns)} columns in our dataset.")
df.info()
# ## Data Correction
df['state'] = df['state'].astype('category')
df['state']
# +
cat_convert = ['emp_title', 'homeownership', 'verification_income_joint']
for i in cat_convert:
df[i] = df[i].astype("category")
# -
df['metric'] = df['annual_income'] / df['total_credit_limit']
# ## Variable lists
#
# list of all numerical data
list_num = df.select_dtypes(include=[np.number]).columns.tolist()
list_num
# list of all categorical data
list_cat = df.select_dtypes(include=['category']).columns.tolist()
list_cat
# ## Prepare for Data Splitting
# +
# define outcome variable as y_label
y_label = 'interest_rate'
# select features
features = df.drop(columns=[y_label]).columns.tolist()
# create feature data for data splitting
X = df[features]
# list of numeric features
feat_num = X.select_dtypes(include=[np.number]).columns.tolist()
# list of categorical features
feat_cat = X.select_dtypes(include=['category']).columns.tolist()
# create response for data splitting
y = df[y_label]
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
# -
# ## Exploration data
# +
df_train = pd.DataFrame(X_train.copy()) #train_test_split-function gibt keine pandas-DataFrame zurück
df_train = df_train.join(pd.DataFrame(y_train))
df_train
# -
|
notebooks/case_loans.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Nirzu97/pyprobml/blob/transformer-torch/notebooks/transformers_torch.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="m6vJh-hiqW_w"
# # Transformers
#
# We show how to implement transformers.
# Based on sec 10.7 of http://d2l.ai/chapter_attention-mechanisms/transformer.html
#
#
#
# + id="Y5XhMfBtqSW4"
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import math
from IPython import display
import torch
from torch import nn
from torch.nn import functional as F
from torch.utils import data
import collections
import re
import random
import os
import requests
import zipfile
import hashlib
import time
np.random.seed(seed=1)
torch.manual_seed(1)
# !mkdir figures # for saving plots
# + [markdown] id="dnCwBw05qn6f"
# # Layers
# + id="8ubc9FVzqovL"
class PositionWiseFFN(nn.Module):
def __init__(self, ffn_num_input, ffn_num_hiddens, ffn_num_outputs,
**kwargs):
super(PositionWiseFFN, self).__init__(**kwargs)
self.dense1 = nn.Linear(ffn_num_input, ffn_num_hiddens)
self.relu = nn.ReLU()
self.dense2 = nn.Linear(ffn_num_hiddens, ffn_num_outputs)
def forward(self, X):
return self.dense2(self.relu(self.dense1(X)))
# + colab={"base_uri": "https://localhost:8080/"} id="TLehwIIwqpRU" outputId="a7ef9bb5-c7eb-4537-a9e0-c7bc88a2100b"
ffn = PositionWiseFFN(4, 4, 8) # batch 4 lengh 4 embed 8
ffn.eval()
Y = ffn(torch.ones((2, 3, 4)))
print(Y.shape)
# + id="D2A0wMZSq2qt"
class AddNorm(nn.Module):
def __init__(self, normalized_shape, dropout, **kwargs):
super(AddNorm, self).__init__(**kwargs)
self.dropout = nn.Dropout(dropout)
self.ln = nn.LayerNorm(normalized_shape)
def forward(self, X, Y):
return self.ln(self.dropout(Y) + X)
# + colab={"base_uri": "https://localhost:8080/"} id="VFitw8AFrEC3" outputId="6e780992-ce9a-45d2-f2ab-de28256390ff"
add_norm = AddNorm([3, 4], 0.5) # Normalized_shape is input.size()[1:]
add_norm.eval()
add_norm(torch.ones((2, 3, 4)), torch.ones((2, 3, 4))).shape
# + [markdown] id="hP7sUitWdzYH"
# ## Abstract base class
#
# + id="Kr1fXr8lclc3"
class Encoder(nn.Module):
"""The base encoder interface for the encoder-decoder architecture."""
def __init__(self, **kwargs):
super(Encoder, self).__init__(**kwargs)
def forward(self, X, *args):
raise NotImplementedError
# + id="zlBBCmCGc43K"
class Decoder(nn.Module):
"""The base decoder interface for the encoder-decoder architecture."""
def __init__(self, **kwargs):
super(Decoder, self).__init__(**kwargs)
def init_state(self, enc_outputs, *args):
raise NotImplementedError
def forward(self, X, state):
raise NotImplementedError
# + id="TLlbB1y1c68w"
class EncoderDecoder(nn.Module):
"""The base class for the encoder-decoder architecture."""
def __init__(self, encoder, decoder, **kwargs):
super(EncoderDecoder, self).__init__(**kwargs)
self.encoder = encoder
self.decoder = decoder
def forward(self, enc_X, dec_X, *args):
enc_outputs = self.encoder(enc_X, *args)
dec_state = self.decoder.init_state(enc_outputs, *args)
return self.decoder(dec_X, dec_state)
# + id="t2xSVsItUBsQ"
class AttentionDecoder(Decoder):
"""The base attention-based decoder interface."""
def __init__(self, **kwargs):
super(AttentionDecoder, self).__init__(**kwargs)
@property
def attention_weights(self):
raise NotImplementedError
# + [markdown] id="_iRpZ75OrIaa"
# # Encoder
# + id="qiKo7tphM-_M"
class DotProductAttention(nn.Module):
"""Scaled dot product attention."""
def __init__(self, dropout, **kwargs):
super(DotProductAttention, self).__init__(**kwargs)
self.dropout = nn.Dropout(dropout)
# Shape of `queries`: (`batch_size`, no. of queries, `d`)
# Shape of `keys`: (`batch_size`, no. of key-value pairs, `d`)
# Shape of `values`: (`batch_size`, no. of key-value pairs, value
# dimension)
# Shape of `valid_lens`: (`batch_size`,) or (`batch_size`, no. of queries)
def forward(self, queries, keys, values, valid_lens=None):
d = queries.shape[-1]
# Set `transpose_b=True` to swap the last two dimensions of `keys`
scores = torch.bmm(queries, keys.transpose(1, 2)) / math.sqrt(d)
self.attention_weights = masked_softmax(scores, valid_lens)
return torch.bmm(self.dropout(self.attention_weights), values)
class MultiHeadAttention(nn.Module):
def __init__(self, key_size, query_size, value_size, num_hiddens,
num_heads, dropout, bias=False, **kwargs):
super(MultiHeadAttention, self).__init__(**kwargs)
self.num_heads = num_heads
self.attention = DotProductAttention(dropout)
self.W_q = nn.Linear(query_size, num_hiddens, bias=bias)
self.W_k = nn.Linear(key_size, num_hiddens, bias=bias)
self.W_v = nn.Linear(value_size, num_hiddens, bias=bias)
self.W_o = nn.Linear(num_hiddens, num_hiddens, bias=bias)
def forward(self, queries, keys, values, valid_lens):
# Shape of `queries`, `keys`, or `values`:
# (`batch_size`, no. of queries or key-value pairs, `num_hiddens`)
# Shape of `valid_lens`:
# (`batch_size`,) or (`batch_size`, no. of queries)
# After transposing, shape of output `queries`, `keys`, or `values`:
# (`batch_size` * `num_heads`, no. of queries or key-value pairs,
# `num_hiddens` / `num_heads`)
queries = transpose_qkv(self.W_q(queries), self.num_heads)
keys = transpose_qkv(self.W_k(keys), self.num_heads)
values = transpose_qkv(self.W_v(values), self.num_heads)
if valid_lens is not None:
# On axis 0, copy the first item (scalar or vector) for
# `num_heads` times, then copy the next item, and so on
valid_lens = torch.repeat_interleave(valid_lens,
repeats=self.num_heads,
dim=0)
# Shape of `output`: (`batch_size` * `num_heads`, no. of queries,
# `num_hiddens` / `num_heads`)
output = self.attention(queries, keys, values, valid_lens)
# Shape of `output_concat`:
# (`batch_size`, no. of queries, `num_hiddens`)
output_concat = transpose_output(output, self.num_heads)
return self.W_o(output_concat)
# + id="bAhv_p0cNk3Z"
def masked_softmax(X, valid_lens):
"""Perform softmax operation by masking elements on the last axis."""
# `X`: 3D tensor, `valid_lens`: 1D or 2D tensor
if valid_lens is None:
return nn.functional.softmax(X, dim=-1)
else:
shape = X.shape
if valid_lens.dim() == 1:
valid_lens = torch.repeat_interleave(valid_lens, shape[1])
else:
valid_lens = valid_lens.reshape(-1)
# On the last axis, replace masked elements with a very large negative
# value, whose exponentiation outputs 0
X = sequence_mask(X.reshape(-1, shape[-1]), valid_lens,
value=-1e6)
return nn.functional.softmax(X.reshape(shape), dim=-1)
def sequence_mask(X, valid_len, value=0):
"""Mask irrelevant entries in sequences."""
maxlen = X.size(1)
mask = torch.arange((maxlen), dtype=torch.float32,
device=X.device)[None, :] < valid_len[:, None]
X[~mask] = value
return X
def transpose_qkv(X, num_heads):
# Shape of input `X`:
# (`batch_size`, no. of queries or key-value pairs, `num_hiddens`).
# Shape of output `X`:
# (`batch_size`, no. of queries or key-value pairs, `num_heads`,
# `num_hiddens` / `num_heads`)
X = X.reshape(X.shape[0], X.shape[1], num_heads, -1)
# Shape of output `X`:
# (`batch_size`, `num_heads`, no. of queries or key-value pairs,
# `num_hiddens` / `num_heads`)
X = X.permute(0, 2, 1, 3)
# Shape of `output`:
# (`batch_size` * `num_heads`, no. of queries or key-value pairs,
# `num_hiddens` / `num_heads`)
return X.reshape(-1, X.shape[2], X.shape[3])
def transpose_output(X, num_heads):
"""Reverse the operation of `transpose_qkv`"""
X = X.reshape(-1, num_heads, X.shape[1], X.shape[2])
X = X.permute(0, 2, 1, 3)
return X.reshape(X.shape[0], X.shape[1], -1)
# + id="Zhnkr_RarJCh"
class PositionalEncoding(nn.Module):
def __init__(self, num_hiddens, dropout, max_len=1000):
super(PositionalEncoding, self).__init__()
self.dropout = nn.Dropout(dropout)
# Create a long enough `P`
self.P = torch.zeros((1, max_len, num_hiddens))
X = torch.arange(max_len, dtype=torch.float32).reshape(
-1, 1) / torch.pow(
10000,
torch.arange(0, num_hiddens, 2, dtype=torch.float32) /
num_hiddens)
self.P[:, :, 0::2] = torch.sin(X)
self.P[:, :, 1::2] = torch.cos(X)
def forward(self, X):
X = X + self.P[:, :X.shape[1], :].to(X.device)
return self.dropout(X)
class EncoderBlock(nn.Module):
def __init__(self, key_size, query_size, value_size, num_hiddens,
norm_shape, ffn_num_input, ffn_num_hiddens, num_heads,
dropout, use_bias=False, **kwargs):
super(EncoderBlock, self).__init__(**kwargs)
self.attention = MultiHeadAttention(key_size, query_size,
value_size, num_hiddens,
num_heads, dropout, use_bias)
self.addnorm1 = AddNorm(norm_shape, dropout)
self.ffn = PositionWiseFFN(ffn_num_input, ffn_num_hiddens,
num_hiddens)
self.addnorm2 = AddNorm(norm_shape, dropout)
def forward(self, X, valid_lens):
Y = self.addnorm1(X, self.attention(X, X, X, valid_lens))
return self.addnorm2(Y, self.ffn(Y))
# + colab={"base_uri": "https://localhost:8080/"} id="cgfrrCKJrJVp" outputId="17d0f000-cd7c-4f13-d69e-256fda3fd4b7"
X = torch.ones((2, 100, 24))
valid_lens = torch.tensor([3, 2])
encoder_blk = EncoderBlock(24, 24, 24, 24, [100, 24], 24, 48, 8, 0.5)
encoder_blk.eval()
encoder_blk(X, valid_lens).shape
# + id="9BHcHiBIrMZ_"
class TransformerEncoder(Encoder):
def __init__(self, vocab_size, key_size, query_size, value_size,
num_hiddens, norm_shape, ffn_num_input, ffn_num_hiddens,
num_heads, num_layers, dropout, use_bias=False, **kwargs):
super(TransformerEncoder, self).__init__(**kwargs)
self.num_hiddens = num_hiddens
self.embedding = nn.Embedding(vocab_size, num_hiddens)
self.pos_encoding = PositionalEncoding(num_hiddens, dropout)
self.blks = nn.Sequential()
for i in range(num_layers):
self.blks.add_module(
"block" + str(i),
EncoderBlock(key_size, query_size, value_size, num_hiddens,
norm_shape, ffn_num_input, ffn_num_hiddens,
num_heads, dropout, use_bias))
def forward(self, X, valid_lens, *args):
# Since positional encoding values are between -1 and 1, the embedding
# values are multiplied by the square root of the embedding dimension
# to rescale before they are summed up
X = self.pos_encoding(self.embedding(X) * math.sqrt(self.num_hiddens))
self.attention_weights = [None] * len(self.blks)
for i, blk in enumerate(self.blks):
X = blk(X, valid_lens)
self.attention_weights[
i] = blk.attention.attention.attention_weights
return X
# + [markdown] id="vn1g8puTrUdw"
# The shape of the transformer encoder output is (batch size, number of time steps, num_hiddens).
# + colab={"base_uri": "https://localhost:8080/"} id="O-u7PLrTrPsT" outputId="31e9d923-a95c-450e-f7cf-efee508bbd3c"
encoder = TransformerEncoder(200, 24, 24, 24, 24, [100, 24], 24, 48, 8, 2,
0.5)
encoder.eval()
encoder(torch.ones((2, 100), dtype=torch.long), valid_lens).shape
# + [markdown] id="l5JEeGFirWUZ"
# # Decoder
# + id="x1GX2mrhrWvJ"
class DecoderBlock(nn.Module):
# The `i`-th block in the decoder
def __init__(self, key_size, query_size, value_size, num_hiddens,
norm_shape, ffn_num_input, ffn_num_hiddens, num_heads,
dropout, i, **kwargs):
super(DecoderBlock, self).__init__(**kwargs)
self.i = i
self.attention1 = MultiHeadAttention(key_size, query_size,
value_size, num_hiddens,
num_heads, dropout)
self.addnorm1 = AddNorm(norm_shape, dropout)
self.attention2 = MultiHeadAttention(key_size, query_size,
value_size, num_hiddens,
num_heads, dropout)
self.addnorm2 = AddNorm(norm_shape, dropout)
self.ffn = PositionWiseFFN(ffn_num_input, ffn_num_hiddens,
num_hiddens)
self.addnorm3 = AddNorm(norm_shape, dropout)
def forward(self, X, state):
enc_outputs, enc_valid_lens = state[0], state[1]
# During training, all the tokens of any output sequence are processed
# at the same time, so `state[2][self.i]` is `None` as initialized.
# When decoding any output sequence token by token during prediction,
# `state[2][self.i]` contains representations of the decoded output at
# the `i`-th block up to the current time step
if state[2][self.i] is None:
key_values = X
else:
key_values = torch.cat((state[2][self.i], X), axis=1)
state[2][self.i] = key_values
if self.training:
batch_size, num_steps, _ = X.shape
# Shape of `dec_valid_lens`: (`batch_size`, `num_steps`), where
# every row is [1, 2, ..., `num_steps`]
dec_valid_lens = torch.arange(1, num_steps + 1,
device=X.device).repeat(
batch_size, 1)
else:
dec_valid_lens = None
# Self-attention
X2 = self.attention1(X, key_values, key_values, dec_valid_lens)
Y = self.addnorm1(X, X2)
# Encoder-decoder attention. Shape of `enc_outputs`:
# (`batch_size`, `num_steps`, `num_hiddens`)
Y2 = self.attention2(Y, enc_outputs, enc_outputs, enc_valid_lens)
Z = self.addnorm2(Y, Y2)
return self.addnorm3(Z, self.ffn(Z)), state
# + colab={"base_uri": "https://localhost:8080/"} id="3LQ9ei-ardtA" outputId="6d576a4a-189e-4894-81c7-da4c64eef744"
decoder_blk = DecoderBlock(24, 24, 24, 24, [100, 24], 24, 48, 8, 0.5, 0)
decoder_blk.eval()
X = torch.ones((2, 100, 24))
state = [encoder_blk(X, valid_lens), valid_lens, [None]]
decoder_blk(X, state)[0].shape
# + id="0eV-Q0NGrfwV"
class TransformerDecoder(AttentionDecoder):
def __init__(self, vocab_size, key_size, query_size, value_size,
num_hiddens, norm_shape, ffn_num_input, ffn_num_hiddens,
num_heads, num_layers, dropout, **kwargs):
super(TransformerDecoder, self).__init__(**kwargs)
self.num_hiddens = num_hiddens
self.num_layers = num_layers
self.embedding = nn.Embedding(vocab_size, num_hiddens)
self.pos_encoding = PositionalEncoding(num_hiddens, dropout)
self.blks = nn.Sequential()
for i in range(num_layers):
self.blks.add_module(
"block" + str(i),
DecoderBlock(key_size, query_size, value_size, num_hiddens,
norm_shape, ffn_num_input, ffn_num_hiddens,
num_heads, dropout, i))
self.dense = nn.Linear(num_hiddens, vocab_size)
def init_state(self, enc_outputs, enc_valid_lens, *args):
return [enc_outputs, enc_valid_lens, [None] * self.num_layers]
def forward(self, X, state):
X = self.pos_encoding(self.embedding(X) * math.sqrt(self.num_hiddens))
self._attention_weights = [[None] * len(self.blks) for _ in range(2)]
for i, blk in enumerate(self.blks):
X, state = blk(X, state)
# Decoder self-attention weights
self._attention_weights[0][
i] = blk.attention1.attention.attention_weights
# Encoder-decoder attention weights
self._attention_weights[1][
i] = blk.attention2.attention.attention_weights
return self.dense(X), state
@property
def attention_weights(self):
return self._attention_weights
# + [markdown] id="j4fhuNfGrrAF"
# # Full model
#
# + id="o_C6Ke82BM7V"
# Required functions for downloading data
def download(name, cache_dir=os.path.join('..', 'data')):
"""Download a file inserted into DATA_HUB, return the local filename."""
assert name in DATA_HUB, f"{name} does not exist in {DATA_HUB}."
url, sha1_hash = DATA_HUB[name]
os.makedirs(cache_dir, exist_ok=True)
fname = os.path.join(cache_dir, url.split('/')[-1])
if os.path.exists(fname):
sha1 = hashlib.sha1()
with open(fname, 'rb') as f:
while True:
data = f.read(1048576)
if not data:
break
sha1.update(data)
if sha1.hexdigest() == sha1_hash:
return fname # Hit cache
print(f'Downloading {fname} from {url}...')
r = requests.get(url, stream=True, verify=True)
with open(fname, 'wb') as f:
f.write(r.content)
return fname
def download_extract(name, folder=None):
"""Download and extract a zip/tar file."""
fname = download(name)
base_dir = os.path.dirname(fname)
data_dir, ext = os.path.splitext(fname)
if ext == '.zip':
fp = zipfile.ZipFile(fname, 'r')
elif ext in ('.tar', '.gz'):
fp = tarfile.open(fname, 'r')
else:
assert False, 'Only zip/tar files can be extracted.'
fp.extractall(base_dir)
return os.path.join(base_dir, folder) if folder else data_dir
# + id="-c0dzTV5AXXn"
def read_data_nmt():
"""Load the English-French dataset."""
data_dir = download_extract('fra-eng')
with open(os.path.join(data_dir, 'fra.txt'), 'r') as f:
return f.read()
def preprocess_nmt(text):
"""Preprocess the English-French dataset."""
def no_space(char, prev_char):
return char in set(',.!?') and prev_char != ' '
# Replace non-breaking space with space, and convert uppercase letters to
# lowercase ones
text = text.replace('\u202f', ' ').replace('\xa0', ' ').lower()
# Insert space between words and punctuation marks
out = [
' ' + char if i > 0 and no_space(char, text[i - 1]) else char
for i, char in enumerate(text)]
return ''.join(out)
def tokenize_nmt(text, num_examples=None):
"""Tokenize the English-French dataset."""
source, target = [], []
for i, line in enumerate(text.split('\n')):
if num_examples and i > num_examples:
break
parts = line.split('\t')
if len(parts) == 2:
source.append(parts[0].split(' '))
target.append(parts[1].split(' '))
return source, target
# + id="GE_0EjLhH1pR"
class Vocab:
"""Vocabulary for text."""
def __init__(self, tokens=None, min_freq=0, reserved_tokens=None):
if tokens is None:
tokens = []
if reserved_tokens is None:
reserved_tokens = []
# Sort according to frequencies
counter = count_corpus(tokens)
self.token_freqs = sorted(counter.items(), key=lambda x: x[1],
reverse=True)
# The index for the unknown token is 0
self.unk, uniq_tokens = 0, ['<unk>'] + reserved_tokens
uniq_tokens += [
token for token, freq in self.token_freqs
if freq >= min_freq and token not in uniq_tokens]
self.idx_to_token, self.token_to_idx = [], dict()
for token in uniq_tokens:
self.idx_to_token.append(token)
self.token_to_idx[token] = len(self.idx_to_token) - 1
def __len__(self):
return len(self.idx_to_token)
def __getitem__(self, tokens):
if not isinstance(tokens, (list, tuple)):
return self.token_to_idx.get(tokens, self.unk)
return [self.__getitem__(token) for token in tokens]
def to_tokens(self, indices):
if not isinstance(indices, (list, tuple)):
return self.idx_to_token[indices]
return [self.idx_to_token[index] for index in indices]
def count_corpus(tokens):
"""Count token frequencies."""
# Here `tokens` is a 1D list or 2D list
if len(tokens) == 0 or isinstance(tokens[0], list):
# Flatten a list of token lists into a list of tokens
tokens = [token for line in tokens for token in line]
return collections.Counter(tokens)
# + id="Gbs426U9IQU0"
reduce_sum = lambda x, *args, **kwargs: x.sum(*args, **kwargs)
astype = lambda x, *args, **kwargs: x.type(*args, **kwargs)
def build_array_nmt(lines, vocab, num_steps):
"""Transform text sequences of machine translation into minibatches."""
lines = [vocab[l] for l in lines]
lines = [l + [vocab['<eos>']] for l in lines]
array = torch.tensor([truncate_pad(
l, num_steps, vocab['<pad>']) for l in lines])
valid_len = reduce_sum(astype(array != vocab['<pad>'], torch.int32), 1)
return array, valid_len
# + id="3yYUa_itADJm"
def load_array(data_arrays, batch_size, is_train=True):
"""Construct a PyTorch data iterator."""
dataset = data.TensorDataset(*data_arrays)
return data.DataLoader(dataset, batch_size, shuffle=is_train)
def truncate_pad(line, num_steps, padding_token):
"""Truncate or pad sequences."""
if len(line) > num_steps:
return line[:num_steps] # Truncate
return line + [padding_token] * (num_steps - len(line))
def load_data_nmt(batch_size, num_steps, num_examples=600):
"""Return the iterator and the vocabularies of the translation dataset."""
text = preprocess_nmt(read_data_nmt())
source, target = tokenize_nmt(text, num_examples)
src_vocab = Vocab(source, min_freq=2,
reserved_tokens=['<pad>', '<bos>', '<eos>'])
tgt_vocab = Vocab(target, min_freq=2,
reserved_tokens=['<pad>', '<bos>', '<eos>'])
src_array, src_valid_len = build_array_nmt(source, src_vocab, num_steps)
tgt_array, tgt_valid_len = build_array_nmt(target, tgt_vocab, num_steps)
data_arrays = (src_array, src_valid_len, tgt_array, tgt_valid_len)
data_iter = load_array(data_arrays, batch_size)
return data_iter, src_vocab, tgt_vocab
# + [markdown] id="f1dieC4GqW68"
# # Data
#
# We use a english-french dataset. See [this colab](https://colab.research.google.com/github/probml/pyprobml/blob/master/notebooks/text_preproc_torch.ipynb#scrollTo=yDmK1xQ9T4IY) for details.
# + id="r-4oeZhRqZ2y" colab={"base_uri": "https://localhost:8080/"} outputId="028e5dd7-3bc2-496d-8338-b48441d09e4c"
DATA_HUB = dict()
DATA_URL = 'http://d2l-data.s3-accelerate.amazonaws.com/'
DATA_HUB['fra-eng'] = (DATA_URL + 'fra-eng.zip',
'94646ad1522d915e7b0f9296181140edcf86a4f5')
batch_size, num_steps = 64, 10
train_iter, src_vocab, tgt_vocab = load_data_nmt(batch_size, num_steps)
# + id="AbqbhodcrsGG"
num_hiddens, num_layers, dropout = 32, 2, 0.1
ffn_num_input, ffn_num_hiddens, num_heads = 32, 64, 4
key_size, query_size, value_size = 32, 32, 32
norm_shape = [32]
encoder = TransformerEncoder(len(src_vocab), key_size, query_size, value_size,
num_hiddens, norm_shape, ffn_num_input,
ffn_num_hiddens, num_heads, num_layers, dropout)
decoder = TransformerDecoder(len(tgt_vocab), key_size, query_size, value_size,
num_hiddens, norm_shape, ffn_num_input,
ffn_num_hiddens, num_heads, num_layers, dropout)
net = EncoderDecoder(encoder, decoder)
# + [markdown] id="THw6lR1bsHD-"
# # Training
# + id="0OJenK-mMh1m"
class Animator:
"""For plotting data in animation."""
def __init__(self, xlabel=None, ylabel=None, legend=None, xlim=None,
ylim=None, xscale='linear', yscale='linear',
fmts=('-', 'm--', 'g-.', 'r:'), nrows=1, ncols=1,
figsize=(3.5, 2.5)):
# Incrementally plot multiple lines
if legend is None:
legend = []
display.set_matplotlib_formats('svg')
self.fig, self.axes = plt.subplots(nrows, ncols, figsize=figsize)
if nrows * ncols == 1:
self.axes = [self.axes,]
# Use a lambda function to capture arguments
self.config_axes = lambda: set_axes(self.axes[
0], xlabel, ylabel, xlim, ylim, xscale, yscale, legend)
self.X, self.Y, self.fmts = None, None, fmts
def add(self, x, y):
# Add multiple data points into the figure
if not hasattr(y, "__len__"):
y = [y]
n = len(y)
if not hasattr(x, "__len__"):
x = [x] * n
if not self.X:
self.X = [[] for _ in range(n)]
if not self.Y:
self.Y = [[] for _ in range(n)]
for i, (a, b) in enumerate(zip(x, y)):
if a is not None and b is not None:
self.X[i].append(a)
self.Y[i].append(b)
self.axes[0].cla()
for x, y, fmt in zip(self.X, self.Y, self.fmts):
self.axes[0].plot(x, y, fmt)
self.config_axes()
display.display(self.fig)
display.clear_output(wait=True)
class Timer:
"""Record multiple running times."""
def __init__(self):
self.times = []
self.start()
def start(self):
"""Start the timer."""
self.tik = time.time()
def stop(self):
"""Stop the timer and record the time in a list."""
self.times.append(time.time() - self.tik)
return self.times[-1]
def avg(self):
"""Return the average time."""
return sum(self.times) / len(self.times)
def sum(self):
"""Return the sum of time."""
return sum(self.times)
def cumsum(self):
"""Return the accumulated time."""
return np.array(self.times).cumsum().tolist()
class Accumulator:
"""For accumulating sums over `n` variables."""
def __init__(self, n):
self.data = [0.0] * n
def add(self, *args):
self.data = [a + float(b) for a, b in zip(self.data, args)]
def reset(self):
self.data = [0.0] * len(self.data)
def __getitem__(self, idx):
return self.data[idx]
# + id="p2fqQdzkZvdF"
def set_axes(axes, xlabel, ylabel, xlim, ylim, xscale, yscale, legend):
"""Set the axes for matplotlib."""
axes.set_xlabel(xlabel)
axes.set_ylabel(ylabel)
axes.set_xscale(xscale)
axes.set_yscale(yscale)
axes.set_xlim(xlim)
axes.set_ylim(ylim)
if legend:
axes.legend(legend)
axes.grid()
def grad_clipping(net, theta):
"""Clip the gradient."""
if isinstance(net, nn.Module):
params = [p for p in net.parameters() if p.requires_grad]
else:
params = net.params
norm = torch.sqrt(sum(torch.sum((p.grad**2)) for p in params))
if norm > theta:
for param in params:
param.grad[:] *= theta / norm
def try_gpu(i=0):
"""Return gpu(i) if exists, otherwise return cpu()."""
if torch.cuda.device_count() >= i + 1:
return torch.device(f'cuda:{i}')
return torch.device('cpu')
# + id="OlHTsKhxg_RM"
class MaskedSoftmaxCELoss(nn.CrossEntropyLoss):
"""The softmax cross-entropy loss with masks."""
# `pred` shape: (`batch_size`, `num_steps`, `vocab_size`)
# `label` shape: (`batch_size`, `num_steps`)
# `valid_len` shape: (`batch_size`,)
def forward(self, pred, label, valid_len):
weights = torch.ones_like(label)
weights = sequence_mask(weights, valid_len)
self.reduction = 'none'
unweighted_loss = super(MaskedSoftmaxCELoss,
self).forward(pred.permute(0, 2, 1), label)
weighted_loss = (unweighted_loss * weights).mean(dim=1)
return weighted_loss
# + id="HORYDhVgpEvg"
def train_seq2seq(net, data_iter, lr, num_epochs, tgt_vocab, device):
"""Train a model for sequence to sequence."""
def xavier_init_weights(m):
if type(m) == nn.Linear:
nn.init.xavier_uniform_(m.weight)
if type(m) == nn.GRU:
for param in m._flat_weights_names:
if "weight" in param:
nn.init.xavier_uniform_(m._parameters[param])
net.apply(xavier_init_weights)
net.to(device)
optimizer = torch.optim.Adam(net.parameters(), lr=lr)
loss = MaskedSoftmaxCELoss()
net.train()
animator = Animator(xlabel='epoch', ylabel='loss',
xlim=[10, num_epochs])
for epoch in range(num_epochs):
timer = Timer()
metric = Accumulator(2) # Sum of training loss, no. of tokens
for batch in data_iter:
X, X_valid_len, Y, Y_valid_len = [x.to(device) for x in batch]
bos = torch.tensor([tgt_vocab['<bos>']] * Y.shape[0],
device=device).reshape(-1, 1)
dec_input = torch.cat([bos, Y[:, :-1]], 1) # Teacher forcing
Y_hat, _ = net(X, dec_input, X_valid_len)
l = loss(Y_hat, Y, Y_valid_len)
l.sum().backward() # Make the loss scalar for `backward`
grad_clipping(net, 1)
num_tokens = Y_valid_len.sum()
optimizer.step()
with torch.no_grad():
metric.add(l.sum(), num_tokens)
if (epoch + 1) % 10 == 0:
animator.add(epoch + 1, (metric[0] / metric[1],))
print(f'loss {metric[0] / metric[1]:.3f}, {metric[1] / timer.stop():.1f} '
f'tokens/sec on {str(device)}')
# + colab={"base_uri": "https://localhost:8080/", "height": 278} id="unvXgpuPr58M" outputId="e6008b4c-f5fd-4933-a88b-a380f6efd8bd"
lr, num_epochs, device = 0.005, 200, try_gpu()
train_seq2seq(net, train_iter, lr, num_epochs, tgt_vocab, device)
# + [markdown] id="XItVF3uzsdCM"
# # Evaluation
# + id="M2VB7oXlqPJD"
def predict_seq2seq(net, src_sentence, src_vocab, tgt_vocab, num_steps,
device, save_attention_weights=False):
"""Predict for sequence to sequence."""
# Set `net` to eval mode for inference
net.eval()
src_tokens = src_vocab[src_sentence.lower().split(' ')] + [
src_vocab['<eos>']]
enc_valid_len = torch.tensor([len(src_tokens)], device=device)
src_tokens = truncate_pad(src_tokens, num_steps, src_vocab['<pad>'])
# Add the batch axis
enc_X = torch.unsqueeze(
torch.tensor(src_tokens, dtype=torch.long, device=device), dim=0)
enc_outputs = net.encoder(enc_X, enc_valid_len)
dec_state = net.decoder.init_state(enc_outputs, enc_valid_len)
# Add the batch axis
dec_X = torch.unsqueeze(
torch.tensor([tgt_vocab['<bos>']], dtype=torch.long, device=device),
dim=0)
output_seq, attention_weight_seq = [], []
for _ in range(num_steps):
Y, dec_state = net.decoder(dec_X, dec_state)
# We use the token with the highest prediction likelihood as the input
# of the decoder at the next time step
dec_X = Y.argmax(dim=2)
pred = dec_X.squeeze(dim=0).type(torch.int32).item()
# Save attention weights (to be covered later)
if save_attention_weights:
attention_weight_seq.append(net.decoder.attention_weights)
# Once the end-of-sequence token is predicted, the generation of the
# output sequence is complete
if pred == tgt_vocab['<eos>']:
break
output_seq.append(pred)
return ' '.join(tgt_vocab.to_tokens(output_seq)), attention_weight_seq
# + id="qwybiVMkwPp8"
def bleu(pred_seq, label_seq, k):
"""Compute the BLEU."""
pred_tokens, label_tokens = pred_seq.split(' '), label_seq.split(' ')
len_pred, len_label = len(pred_tokens), len(label_tokens)
score = math.exp(min(0, 1 - len_label / len_pred))
for n in range(1, k + 1):
num_matches, label_subs = 0, collections.defaultdict(int)
for i in range(len_label - n + 1):
label_subs[''.join(label_tokens[i:i + n])] += 1
for i in range(len_pred - n + 1):
if label_subs[''.join(pred_tokens[i:i + n])] > 0:
num_matches += 1
label_subs[''.join(pred_tokens[i:i + n])] -= 1
score *= math.pow(num_matches / (len_pred - n + 1), math.pow(0.5, n))
return score
# + colab={"base_uri": "https://localhost:8080/"} id="JRbDnsZLsPdy" outputId="9502700f-8e64-4d2d-b9c3-5513f83cd5cc"
engs = ['go .', "i lost .", 'he\'s calm .', 'i\'m home .']
fras = ['va !', 'j\'ai perdu .', 'il est calme .', 'je suis chez moi .']
for eng, fra in zip(engs, fras):
translation, dec_attention_weight_seq = predict_seq2seq(
net, eng, src_vocab, tgt_vocab, num_steps, device, True)
print(f'{eng} => {translation}, ',
f'bleu {bleu(translation, fra, k=2):.3f}')
# + [markdown] id="_rxnfmBtsgnn"
# # Visualization of attention heatmaps
#
# We visualize the attention heatmaps for the last (english, french) pair, where the input has length 3 and the output has length 5.
#
# The shape of the encoder self-attention weights is (number of encoder layers, number of attention heads, num_steps or number of queries, num_steps or number of key-value pairs).
#
# + colab={"base_uri": "https://localhost:8080/"} id="OoNT_10Zs540" outputId="a90f0d49-ac4d-4575-d3c0-2364c7c025f2"
enc_attention_weights = torch.cat(net.encoder.attention_weights, 0).reshape(
(num_layers, num_heads, -1, num_steps))
enc_attention_weights.shape
# + [markdown] id="gS_fjqcItJBS"
# Encoder self-attention for each of the 2 encoder blocks.
# The input has length 4, so all keys are 0 after that.
# + id="kHyQ3ugTTX9q"
def show_heatmaps(matrices, xlabel, ylabel, titles=None, figsize=(2.5, 2.5),
cmap='Reds'):
display.set_matplotlib_formats('svg')
num_rows, num_cols = matrices.shape[0], matrices.shape[1]
fig, axes = plt.subplots(num_rows, num_cols, figsize=figsize,
sharex=True, sharey=True, squeeze=False)
for i, (row_axes, row_matrices) in enumerate(zip(axes, matrices)):
for j, (ax, matrix) in enumerate(zip(row_axes, row_matrices)):
pcm = ax.imshow(matrix.detach(), cmap=cmap)
if i == num_rows - 1:
ax.set_xlabel(xlabel)
if j == 0:
ax.set_ylabel(ylabel)
if titles:
ax.set_title(titles[j])
fig.colorbar(pcm, ax=axes, shrink=0.6)
# + colab={"base_uri": "https://localhost:8080/", "height": 329} id="smscWfpntKFt" outputId="a2a4a508-84e8-4a0a-a440-3b761ec8390e"
show_heatmaps(enc_attention_weights.cpu(), xlabel='Key positions',
ylabel='Query positions',
titles=['Head %d' % i
for i in range(1, 5)], figsize=(7, 3.5))
# + [markdown] id="gG6i2tQFtpSi"
# Next we visualize decoder attention heatmaps.
# + colab={"base_uri": "https://localhost:8080/"} id="0GEyFQ5ctKj6" outputId="7045ef48-f2cf-4372-8221-9cb9c5c2f66e"
dec_attention_weights_2d = [
head[0].tolist() for step in dec_attention_weight_seq for attn in step
for blk in attn for head in blk]
dec_attention_weights_filled = torch.tensor(
pd.DataFrame(dec_attention_weights_2d).fillna(0.0).values)
dec_attention_weights = dec_attention_weights_filled.reshape(
(-1, 2, num_layers, num_heads, num_steps))
dec_self_attention_weights, dec_inter_attention_weights = \
dec_attention_weights.permute(1, 2, 3, 0, 4)
dec_self_attention_weights.shape, dec_inter_attention_weights.shape
# + [markdown] id="O9Jb3D3Zt97P"
# Decoder self-attention.
# + colab={"base_uri": "https://localhost:8080/", "height": 329} id="RIK92fjltsav" outputId="65948cd7-ce29-47a0-9264-74eb01ad9d08"
# Plus one to include the beginning-of-sequence token
show_heatmaps(
dec_self_attention_weights[:, :, :, :len(translation.split()) + 1],
xlabel='Key positions', ylabel='Query positions',
titles=['Head %d' % i for i in range(1, 5)], figsize=(7, 3.5))
# + [markdown] id="cab19ZpWuI0A"
# Decoder encoder-attention.
# + colab={"base_uri": "https://localhost:8080/", "height": 298} id="vASzp_5tt_cH" outputId="e7839094-d57e-4fcb-f5b6-c6a5a6408261"
show_heatmaps(dec_inter_attention_weights, xlabel='Key positions',
ylabel='Query positions',
titles=['Head %d' % i
for i in range(1, 5)], figsize=(7, 3.5))
# + id="xdz_OJFPuKb-"
|
notebooks-d2l/transformers_torch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
def image_to_video(image_folder_path: str, fps, extension:str, video_name:str, output_format:str):
'''Convert all the images in a specified directory to vieo at a specified framerate
- image_folder_path is the directory of the images
- fps is the number of frames per seconds
- extension could be .jpeg, .tiff, .png etc depending on the image format
- video_name is the name of video file created
- output_format is the format of the created video, could be .mp4, .avi, .mov, etc
'''
import os
import moviepy.video.io.ImageSequenceClip
images = [image_folder_path+'/'+img for img in os.listdir(image_folder_path) if img.endswith(extension)]
movie_clip = moviepy.video.io.ImageSequenceClip.ImageSequenceClip(images, fps)
movie_clip.write_videofile(video_name+output_format)
if __name__ =='__main__':
image_to_video('/Users/...', 54, '.jpeg', 'Spectra_video', '.mov');
|
Research Data Analysis/Image Processing/Imagesequence_2_Video.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ibmfl-venv
# language: python
# name: ibmfl-venv
# ---
# ## Dashboard
#
# In this Notebook, we interact with the Experiment Manager to configure, setup and run experiments.
# ### Import dependencies, initialise configs
# + code_folding=[0]
## Imports and such
import sys
sys.path.append('../')
# %config Completer.use_jedi = False # to avoid autocomplete errors in Jupyter server
from ipywidgets import GridspecLayout, GridBox, Layout, Output
import dashboard_ui
dashboard_ui = dashboard_ui.DashboardUI()
# -
# ### Choose Model, Dataset and Fusion Algorithm
# #### Provide Data Handler
# - Only if you wish to use a Custom Dataset
# - Choose Yes in the `Custom Dataset?` option below
#
# Populate and then run the cell below to save the provided Data Handler class to file.
# + code_folding=[0]
# %%writefile custom_data_handler.py
### YOUR DATAHANDLER code goes below
import logging
import numpy as np
from ibmfl.data.data_handler import DataHandler
from ibmfl.util.datasets import load_mnist
logger = logging.getLogger(__name__)
class MnistKerasDataHandler(DataHandler):
"""
Data handler for MNIST dataset.
"""
def __init__(self, data_config=None, channels_first=False):
super().__init__()
self.file_name = None
if data_config is not None:
# Ensure your data files are either npz or csv
if 'npz_file' in data_config:
self.file_name = data_config['npz_file']
elif 'txt_file' in data_config:
self.file_name = data_config['txt_file']
self.channels_first = channels_first
# load the datasets
(self.x_train, self.y_train), (self.x_test, self.y_test) = self.load_dataset()
# pre-process the datasets
self.preprocess()
def get_data(self):
"""
Gets pre-process mnist training and testing data.
:return: the training and testing data.
:rtype: `tuple`
"""
return (self.x_train, self.y_train), (self.x_test, self.y_test)
def load_dataset(self, nb_points=500):
"""
Loads the training and testing datasets from a given local path. \
If no local path is provided, it will download the original MNIST \
dataset online, and reduce the dataset size to contain \
500 data points per training and testing dataset. \
Because this method \
is for testing it takes as input the number of datapoints, nb_points, \
to be included in the training and testing set.
:param nb_points: Number of data points to be included in each set if
no local dataset is provided.
:type nb_points: `int`
:return: training and testing datasets
:rtype: `tuple`
"""
if self.file_name is None:
(x_train, y_train), (x_test, y_test) = load_mnist()
# Reduce datapoints to make test faster
x_train = x_train[:nb_points]
y_train = y_train[:nb_points]
x_test = x_test[:nb_points]
y_test = y_test[:nb_points]
else:
try:
logger.info('Loaded training data from ' + str(self.file_name))
data_train = np.load(self.file_name)
x_train = data_train['x_train']
y_train = data_train['y_train']
x_test = data_train['x_test']
y_test = data_train['y_test']
except Exception:
raise IOError('Unable to load training data from path '
'provided in config file: ' +
self.file_name)
return (x_train, y_train), (x_test, y_test)
def preprocess(self):
"""
Preprocesses the training and testing dataset, \
e.g., reshape the images according to self.channels_first; \
convert the labels to binary class matrices.
:return: None
"""
num_classes = 10
img_rows, img_cols = 28, 28
if self.channels_first:
self.x_train = self.x_train.reshape(self.x_train.shape[0], 1, img_rows, img_cols)
self.x_test = self.x_test.reshape(self.x_test.shape[0], 1, img_rows, img_cols)
else:
self.x_train = self.x_train.reshape(self.x_train.shape[0], img_rows, img_cols, 1)
self.x_test = self.x_test.reshape(self.x_test.shape[0], img_rows, img_cols, 1)
# convert class vectors to binary class matrices
self.y_train = np.eye(num_classes)[self.y_train]
self.y_test = np.eye(num_classes)[self.y_test]
# + code_folding=[0]
## Model, Dataset and Fusion Algorithm
components = dashboard_ui.generate_model_dataset_fusion_ui()
# GridBox layout for UI
grid = GridspecLayout(2,2)
grid[0,:] = GridBox(children=list(components[:-4]),
layout=Layout(
width='100%',
grid_template_rows='auto auto',
grid_template_columns='48% 48%',
grid_template_areas='''
"model_header model_header"
"model_dr model_upload"
"dataset_header dataset_header"
"dataset dataset_spl"
"ppp ppp"
'''
# ,border='0.5px solid black'
))
grid[1,:] = GridBox(children=list(components[-4:]),
layout=Layout(
height='150px',
width='100%',
grid_template_rows='auto auto',
grid_template_columns='48% 48%',
grid_gap = '0px 0px',
grid_template_areas='''
"custom_data custom_data_html"
"fusion_dr metrics_choice"
'''
# , border='0.5px solid black'
))
# grid[2,:] = GridBox(children=list(components[-1:]),
# layout=Layout(
# height='55px',
# width='auto',
# grid_template_rows='100%',
# grid_template_columns='100%',
# grid_template_areas='''
# "fusion_dr"
# ''',
# border='0.5px solid black'
# ))
grid
# -
# ### Choose number of parties and hyperparameters
# Ensure you click `Confirm Hyperparameters` when done!
# + code_folding=[0]
## Parties and Hyperparameters
components = list(dashboard_ui.generate_parties_hyperparams_ui())
# GridBox layout for UI
grid = GridspecLayout(2,3)
grid[0,:] = GridBox(children=components[:-2],
layout = Layout(
width='100%',
grid_template_rows='auto auto',
grid_template_columns='48% 48%',
grid_template_areas='''
"header_parties header_parties"
"parties parties"
"header_hyperparams header_hyperparams"
''')
)
# Nested grid to vary spacing across various widgets
sub_grid_hyperparams = GridspecLayout(2,3)
sub_grid_hyperparams[0,:] = components[-1]
sub_grid_hyperparams[1,1] = components[-2]
grid[1, :] = sub_grid_hyperparams
party_hyperparam_ui = Output()
with party_hyperparam_ui:
display(grid)
party_hyperparam_ui
# -
# #### Provide Party specific data files
#
# - Only if you wish to use a Custom Dataset
# - Chose Yes in the `Custom Dataset?` option in Step 1.2 above
# + code_folding=[0]
## Upload party data files for each party:
if 'custom_data' in dashboard_ui.mgr.nb_config:
upload_boxes = dashboard_ui.generate_custom_party_data_ui()
for each in upload_boxes:
display(each)
# -
# ### Choose whether to run locally or on remote machines
# + code_folding=[0]
## Local or Remote run
components = dashboard_ui.generate_local_remote_ui()
# grid for displaying networking fields -- IP addr, port, ssh user, paths
partyDetails_grid = GridspecLayout(1,3)
partyDetails_grid[0, :] = components[1] # networking_deets_box
display(components[0])
partyDetails_grid
# -
# ### Generate and View Aggregator and Party Config
# + code_folding=[0]
## Generate Configs and Display them
components = dashboard_ui.generate_display_configs_ui()
# grid for displaying generated configurations
display_grid_1 = GridspecLayout(1,3)
display_grid_1[0, :] = components[1] # config_box
display_grid_1
# -
# ### Run the Experiment and Visualise Metrics
# If the configs above look alright, go ahead and run the cell below to run the experiment!
# + code_folding=[0]
## Run the experiment and see charts
import ibmfl_cli_automator.run as ibmfl_runner
from ipywidgets import Button, VBox, Output
exp_runner = ibmfl_runner.Runner()
monitoring_box = VBox()
no_plots_for_these = ['Federated Averaging', 'Gradient Averaging', 'Probabilistic Federated Neural Matching', 'Zeno', 'Shuffled Iterative Avg']
plot_button = Button(
description='Show Charts',
disabled=False,
button_style='warning', # 'success', 'info', 'warning', 'danger' or ''
tooltip='Displays the various plots for the experiment that ran',
layout = Layout(width='120px', height='40px', margin='5px 50px 5px 400px') ## margin to position button centrally
)
def invoke_runner():
monitoring_out = Output(layout={'border': '0.5px solid black'})
monitoring_box.children = [monitoring_out]
display(display_grid_2)
# some values needed by the Runner; there's only one trial for now
dashboard_ui.mgr.run_details['experiments'][0]['shuffle_party_machines'] = False
dashboard_ui.mgr.run_details['experiments'][0]['n_trials'] = 1
dashboard_ui.mgr.run_details['experiments'][0]['n_parties'] = dashboard_ui.mgr.nb_config['global']['num_parties']
dashboard_ui.mgr.run_details['experiments'][0]['n_rounds'] = dashboard_ui.mgr.nb_config['global']['rounds']
# values for postprocessing and showing default metrics
if dashboard_ui.mgr.nb_config['record_metrics']:
dashboard_ui.mgr.run_details['experiments'][0]['postproc_fn'] = {}
dashboard_ui.mgr.run_details['experiments'][0]['postproc_fn'] = 'gen_reward_vs_time_plots'
dashboard_ui.mgr.run_details['experiments'][0]['postproc_x_key'] = 'post_train:ts'
dashboard_ui.mgr.run_details['experiments'][0]['postproc_y_keys'] = ['post_train:eval:loss', 'post_train:eval:acc']#, 'post_train:eval:precision weighted', 'post_train:eval:recall weighted']
exp_machines = exp_runner.convert_machine_dict_from_nb_to_cli(dashboard_ui.mgr.run_details['machines'])
for exp_info in dashboard_ui.mgr.run_details['experiments']:
with open('{}/config_agg.yml'.format(dashboard_ui.mgr.nb_config['local_conf_dir']), 'r') as config_agg_file:
config_agg = config_agg_file.read()
config_parties = []
for pi in range(exp_info['n_parties']):
with open('{}/config_party{}.yml'.format(dashboard_ui.mgr.nb_config['local_conf_dir'], pi), 'r') as config_party_file:
config_parties += [config_party_file.read()]
with monitoring_out:
display(exp_runner.run_experiment(exp_info, dashboard_ui.mgr.run_details['machines'],
config_agg, config_parties, ui_mode='nb', ts=dashboard_ui.mgr.nb_config['timestamp_str']) \
or 'Finished!')
if dashboard_ui.mgr.nb_config['record_metrics']:
if 'Keras' in dashboard_ui.mgr.nb_config['model'] and dashboard_ui.mgr.nb_config['fusion'] not in no_plots_for_these:
# only some Keras models have plots currently
monitoring_box.children = monitoring_box.children + (plot_button,)
else:
with monitoring_out:
display('Plots for chosen model/fusion algorithm are not supported yet') # metrics processing not in place
else:
with monitoring_out:
display('No metrics were recorded, so no plots to show')
plots_box = VBox()
def get_plots(b):
b.disabled = True
plots_out = Output(layout={'border': '0.5px solid black'})
plots_box.children = [plots_out]
display(display_grid_3)
# generate the plot(s)
with plots_out:
display(exp_info = exp_runner.call_postproc_fn())
plot_button.on_click(get_plots)
# grid for displaying progress of running experiment
display_grid_2 = GridspecLayout(1,1)
display_grid_2[0, :] = monitoring_box
# grid for displaying charts from collected metrics
display_grid_3 = GridspecLayout(1,1)
display_grid_3[0, :] = plots_box
invoke_runner()
|
experiment_manager/Experiment_Manager_dashboard.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from scipy import stats
import statistics
import matplotlib
import matplotlib.pyplot as plt
from matplotlib.pyplot import figure
from IPython import display
import seaborn as sns
import csv
from statistics import mean
# -
df = pd.read_csv('startup.csv')
df.head()
df_dropped = df.dropna()
df_dropped.tail()
df_dropped['founded_at'].unique()
# ## Question1: what is the funding_total_usd for 2011? USD in K
df_dropped[df_dropped['founded_at']=='2011']['funding_total_usd']
# total funding for 2011
df_dropped[df_dropped['founded_at']=='2011']['funding_total_usd'].sum()
df_dropped['founded_at'].sort_values()
list(df_dropped['founded_at'].sort_values().unique())
df_dropped['founded_at'].value_counts()
df_dropped[df_dropped['founded_at'] == '2011']['funding_total_usd'].sum()
# ## Question2: what is the funding_total_usd for all available years?
# +
ls = []
for i in list(df_dropped['founded_at'].sort_values().unique()):
ls.append(df_dropped[df_dropped['founded_at'] == i]['funding_total_usd'].sum())
print(ls)
# -
plt.figure(figsize = (20,5), frameon = False)
plt.xticks(rotation=90)
plt.plot(list(df_dropped['founded_at'].sort_values().unique()), ls,'o')
print(df_dropped[df_dropped['founded_at'] == '2009']['funding_total_usd'].sum())
print(df_dropped[df_dropped['founded_at'] == '2007']['funding_total_usd'].sum())
print(df_dropped[(df_dropped['founded_at'] == '1953')]['funding_total_usd'].sum())
print(df_dropped[df_dropped['founded_at'] == '1953']['category_list'])
# ## Question 3: Answer Question 2 for Software Category
# +
ls_software = []
for i in list(df_dropped['founded_at'].sort_values().unique()):
ls_software.append(df_dropped[(df_dropped['founded_at'] == i) & (df_dropped['category_list'] == 'Software')]['funding_total_usd'].sum())
print(ls_software)
# -
plt.figure(figsize = (20,5), frameon = False)
plt.xticks(rotation=90)
plt.plot(list(df_dropped['founded_at'].sort_values().unique()), ls_software,'o')
# # Question 4: Answer Question 3 for Hardware Category
# +
ls_Hardware = []
for i in list(df_dropped['founded_at'].sort_values().unique()):
ls_Hardware.append(df_dropped[(df_dropped['founded_at'] == i) & (df_dropped['category_list'] == 'Hardware')]['funding_total_usd'].sum())
print(ls_Hardware)
# -
plt.figure(figsize = (20,5), frameon = False)
plt.xticks(rotation=90)
plt.plot(list(df_dropped['founded_at'].sort_values().unique()), ls_Hardware,'o')
# # Regardless of the year, how much fund does exist for Automation, Business, ...?
df_dropped.groupby('category_list')['funding_total_usd'].sum()
df_dropped.groupby('category_list')['funding_total_usd'].sum().plot(kind='bar')
print(sum(ls_software))
# # Question 5: What % of startups are software or Technology?
sample = df_dropped['category_list'].value_counts()
total = df_dropped['category_list'].value_counts().sum()
total
# +
c = df_dropped['category_list'].value_counts()
d = df_dropped['category_list'].value_counts().sum()
# -
def percentage (d, c):
return ((c/d)*100)
for each in c:
pc = percentage(d, each)
print (pc)
# # software funding in San Francisco
san_francisco = df_dropped.query('city == "San Francisco"')
san_francisco.head()
san_francisco.info()
# +
sf_software = []
for i in list(san_francisco['founded_at'].sort_values().unique()):
sf_software.append(san_francisco[(san_francisco['founded_at'] == i) & (san_francisco['category_list'] == 'Software')]['funding_total_usd'].sum())
print(sf_software)
# -
plt.figure(figsize = (20,5), frameon = False)
plt.xticks(rotation=90)
plt.plot(list(san_francisco['founded_at'].sort_values().unique()), sf_software,'o')
# # Pearson correlation between the number of investers and the funding total
# Null hypothesis: number of investers are not correlated with funding total.
X = df_dropped.Number_of_Investors.values
Y = df_dropped.funding_total_usd.values
def pearson_r(X, Y):
corr_mat=np.corrcoef(X,Y)
return corr_mat[0,1]
r_obs = pearson_r(X,Y)
print('Observed significance value=',r_obs)
# # Statistical analysis of total funding in software category.
software_funding = df_dropped.query('category_list == "Software"')
mean = np.mean(software_funding.funding_total_usd)
print(mean)
std_dev = np.std(software_funding.funding_total_usd)
print(std_dev)
mode = stats.mode(software_funding.funding_total_usd)
print(mode)
median = statistics.median(software_funding.funding_total_usd)
print(median)
software_funding.funding_total_usd.describe()
# # Analysis of the number of investors in startups
# matplotlib histogram
plt.hist(software_funding['Number_of_Investors'], color = 'blue', edgecolor = 'black',
bins = int(180/5))
# Density Plot and Histogram of all arrival delays
sns.distplot(software_funding['Number_of_Investors'], hist=True, kde=True,
bins=int(180/5), color = 'darkblue',
hist_kws={'edgecolor':'black'},
kde_kws={'linewidth': 4})
# # Showing the correlation between number_of_investors and funding_rounds.
corr = df_dropped.corr()
ax = sns.heatmap(
corr,
vmin=-1, vmax=1, center=0,
cmap=sns.diverging_palette(20, 220, n=200),
square=True
)
ax.set_xticklabels(
ax.get_xticklabels(),
rotation=90,
)
ax.set_yticklabels(
ax.get_yticklabels(),
rotation=0,
horizontalalignment='right'
);
# # Making an ECDF
x = np.sort(df_dropped['Number_of_Investors'])
y = np.arange(1, len(x)+1) /len(x)
_ = plt.plot(x,y, marker='.', linestyle='none')
_ = plt.xlabel('Number of investors')
_ = plt.ylabel('ECDF')
plt.margins(0.02)
plt.show()
# +
# Summary
# After years of growth since 2007, investment has continued to underperform.
# In 1983, massive funding was made in software industry.
# From 2010 to 2013, investment was declined significantly.
# In Hardware field, a biggest investment was made only in 1953. This might be due to the fact that IBM effectively created the computer market in 1953 with the IBM 650.
# For data format (Month-Date) in founded date, these data may belongs to recent years based on its low funding total in overall trend.
# Over 35% startups got 1~2 investors, and funding round and number of investors showed a mild correlation.
|
site/public/courses/DS-1.1/Notebooks/start_ups_EDA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using AHPTree from `pyanp` example 2
# ## First we import
import pandas as pd
import pyanp.ahptree as ahptree
# %pylab inline
from IPython.display import HTML
pylab.rcParams['figure.figsize'] = (12, 7.5)
# # Example model
# ## Read in the data and verify it looks right
excel_file = 'AHPTreeData2.xlsx'
ahp = ahptree.ahptree_fromdf(excel_file)
ahp
# ## Look at global priorities
gs = ahp.global_priority_table()
gs.plot(kind="bar")
gs
altscores = ahp.priority_table()
altscores.plot(kind="bar")
altscores
allincons = ahp.incond_std_table()
display(HTML(allincons.to_html().replace('\n','')))
allincons.plot(kind="bar")
plt.show()
|
examples/AHPTreeEx2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: analysis
# language: python
# name: analysis
# ---
# # Datasets from Scikit-Learn
import pandas as pd
import numpy as np
from sklearn import datasets
# ## Boston Dataset
data = datasets.load_boston()
df = pd.DataFrame(data= np.c_[data['data'], data['target']],columns= list(data['feature_names']) + ['target'])
df.info()
# ## Wine Dataset
data = datasets.load_wine()
df = pd.DataFrame(data= np.c_[data['data'], data['target']],columns= list(data['feature_names']) + ['target'])
df['target'] = df['target'].apply(lambda x: data['target_names'][int(x)])
df.info()
# ## Iris Dataset
data = datasets.load_iris()
df = pd.DataFrame(data= np.c_[data['data'], data['target']],columns= list(data['feature_names']) + ['target'])
df['target'] = df['target'].apply(lambda x: data['target_names'][int(x)])
df.info()
|
notebooks/datasets/datasets-scikit_learn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.2
# language: julia
# name: julia-1.5
# ---
# # Collocation
#
# This notebook solves the income fluctuations problem using collocation on the value function. Collocation is a dimensionality reduction technique which provides necessary efficiency gains when dealing with high-dimensional problems. With just ten asset grid points (on an unevenly spaced grid), we can obtain accurate solutions to the value function in less than one-tenth of the time relative to a "full" discretization of the state space (of course, this depends on the number of points, but compare this to the VFI notebook).
#
# The notebook heavily uses the powerful BasisMatrices.jl library and follows the example therein for the Aiyagari model. Take a look at the [examples](https://github.com/QuantEcon/BasisMatrices.jl/blob/master/demo/examples.ipynb) in the documentation to get a better idea of how BasisMatrices.jl constructs basis matrices and evaluates approximated functions.
# +
include("rowenhorst_tauchen.jl") # include discretization functions (make sure it is in your working directory)
using Parameters, Plots, LinearAlgebra, BasisMatrices, SparseArrays, QuantEcon, Optim
Household = @with_kw (apoints, # asset grid
amax, # asset max
bc, # borrowing constraint (must be weakly negative)
agrid, # asset grid
basis, # basis type
bs, # basis matrix, not expanded
expectation_basis, # expectation basis matrix
transition_matrix, # transition matrix
ygrid, # grid for income process
nodes, # tensor product of agrid, ygrid
beta = 0.98, # discount factor
alpha = 0.11, # capital share
deprec = 0.025, # depreciation rate
gamma = 1, # elasticity of substitution
phi = 0.8, # disutility factor
frisch = 0.5) # frisch elasticity for labor supply
# -
# ### Constructing grids
#
# The main difference relative to other notebooks (EGM, VFI) is that we need to do a bit more work in constructing the basis matrices (which we will use to approximate our desired function) and the grids.
# +
function construct_household(bc = 0, apoints = 10, amax = 70,
mean = 0, sd = 0.13, rho = 0.966, uncond_sd = sd/sqrt(1-rho^2), num_states = 7, order = 3, curve = 2)
agrid = range(bc^(1/curve), stop = amax^(1/curve), length = apoints).^curve
transition_matrix, ygrid = rowenhorst(mean, uncond_sd, rho, num_states)
a_basis = Basis(SplineParams(agrid, 0 , order))
y_basis = Basis(SplineParams(ygrid, 0 , 1))
basis = Basis(a_basis, y_basis)
s, (agrid, ygrid) = nodes(basis)
Ns, Na, Ny = size(s, 1), size(agrid, 1), size(ygrid, 1)
@assert Ny == num_states
# compute expectations matrix
bs = BasisMatrix(basis, Direct())
bs_expanded = convert(Expanded, bs).vals[1]
expectation_basis = kron(transition_matrix, sparse(I, Na, Na))*bs_expanded
hh = Household(apoints = apoints, amax = amax, bc = bc, agrid = agrid, basis = basis, bs = bs, expectation_basis = expectation_basis,
transition_matrix = transition_matrix, ygrid = ygrid, nodes = s)
return hh
end
hh = construct_household()
# -
# ### Approximating the value function
#
# We now use the basis we constructed to approximate the value function. Note, however, that the value function is not the only function we could approximate. We could have also approximated its derivative, or the consumption policy function itself and then used the endogenous grid method, or the envelope condition method (Maliar and Maliar, 2011). In many cases, this is desirable because it avoids root-finding. Indeed, this is what the endogenous grid method does indirectly -- it interpolates the current consumption function on the future grid. This is equivalent to forming a linear basis on the current grid, and then evaluating that function on the future asset grid.
# +
function utility(gamma, nodes, anext, r, w)
c = (1 + r).*nodes[:, 1] + w.*nodes[:, 2] - anext
u_mat = similar(c)
fill!(u_mat, 0)
for cc=1:length(c)
if c[cc] <= 0
u_mat[cc] = -Inf
else
u_mat[cc] = u(c[cc], gamma)
end
end
return u_mat
end
function utility_labor(hh, nodes, anext, r, w)
@unpack gamma, phi, frisch = hh
u_mat = similar(anext)
c(l, i) = max((1 + r).*nodes[i, 1] + w.*nodes[i, 2].*l - anext[i], 0)
for i in eachindex(anext)
res = optimize(l -> -u(c(l, i), gamma) + v(l, phi, frisch), 0.0, 1.0)
u_mat[i] = -res.minimum
end
return u_mat
end
function v(l, phi, frisch)
return phi*l^(1+frisch)/(1+frisch)
end
function u(c, gamma)
if gamma == 1
return log.(c)
else
return @. c^(1-gamma)/(1-gamma)
end
end
function value(hh; anext, r, w, ce, labor = true)
@unpack gamma, nodes, basis, beta = hh
# flow payoff
if labor
u = utility_labor(hh, nodes, anext, r, w)
else
u = utility(gamma, nodes, anext, r, w)
end
# Basis matrix for continuation value
ve = reshape(funeval(ce, basis, [anext nodes[:,2]]), (:,))
# Compute value
v1 = u + beta.*ve
return v1
end
value(hh, anext = ones(84).*10, r=1, w=1, ce = ones(84)*.100)
# -
function opt_value(hh, r, w, ce, c1)
@unpack nodes, bc, expectation_basis = hh
# solve maximization problem
lower_bound = zeros(size(nodes, 1)) .+ bc
upper_bound = (1 + r).*nodes[:, 1] + w.*nodes[:, 2]
f(anext) = value(hh, anext = anext, r = r, w = w, ce = ce)
ap, v1 = golden_method(f, lower_bound, upper_bound)
# compute expected value function
ve = expectation_basis*c1
return v1, ve, ap
end
opt_value(hh, 0.1, 0.2, zeros(84), zeros(84))
# +
function coeff(hh; r = 0.01, w = 0.5, ce, c1)
@unpack nodes, bs = hh
# Compute values
v1, ve, ap = opt_value(hh, r, w, ce, c1)
bs_expanded = convert(Expanded, bs).vals[1]
# Update coefficients
c1 = bs_expanded\v1
ce = bs_expanded\ve
return c1, ce
end
function vfi(hh; tol=1e-8, maxiter = 10000)
@unpack nodes = hh
ce_old = zeros(length(nodes[:,1]))
c1_old = zeros(length(nodes[:,1]))
for i=1:maxiter
c1, ce = coeff(hh, ce = ce_old, c1 = c1_old)
dc = norm([c1, ce] - [c1_old, ce_old], Inf)
if dc < tol
println("Converged in $(i) iterations.")
return c1, ce
else ce_old = ce; c1_old = c1;
end
end
error("no convergence!")
end
# +
c1, ce = vfi(hh)
r = 0.01; w = 0.5;
v1, _ , ap = opt_value(hh, r, w, ce, c1)
# +
# for the case without labor just set labor = false in the value function and set l = 1 below
l = similar(ap)
c(l, i) = max((1 + r).*hh.nodes[i, 1] + w.*hh.nodes[i, 2].*l - ap[i], 0)
for i in eachindex(ap)
res = optimize(l -> -u(c(l, i), hh.gamma) + v(l, hh.phi, hh.frisch), 0.0, 1.0)
l[i] = res.minimizer
end
consumption = (1 + r).*hh.nodes[:, 1] + w.*hh.nodes[:, 2].*l - ap
consumption = reshape(consumption, (:,length(hh.ygrid)))
plot(hh.agrid, consumption, legend = false, title = "consumption function for various incomes", xlabel = "assets" )
# -
# ### Final Remarks
#
# We can speed up the algorithm by using a form of the Howard improvement step. Because the derivative of the value function can be found in closed form with collocation, we can transform the iteration on the coefficients as a root finding problem using Newton-Raphson methods.
#
# Although collocation is a valuable dimensionality reduction technique, the state space still increases exponentially in the number of state variables. BasisMatrices.jl has the tools to construct Smolyak grids, which make the number of nodes grow at a polynomial rate. This is necessary when dealing with models with a very large state space, such as international business cycle models.
|
collocation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [tensorflow]
# language: python
# name: Python [tensorflow]
# ---
# + deletable=true editable=true
from monthly_totals import *
# + deletable=true editable=true
data = get_precip_data()
# + deletable=true editable=true
totals_pd, counts_pd = pivot_months_pandas(data)
# + deletable=true editable=true
totals_loop, counts_loop = pivot_months_loops(data)
# -
totals_loop # look how nicely Jupyter displays a DataFrame...
# + deletable=true editable=true
abs(totals_pd - totals_loop).values.sum() # the two totals should be the same
# + deletable=true editable=true
abs(counts_pd - counts_loop).values.sum() # ...counts too.
# + deletable=true editable=true
# %timeit pivot_months_pandas(data)
# + deletable=true editable=true
# %timeit pivot_months_loops(data)
|
ex1/code/timing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: env37
# language: python
# name: env37
# ---
#hide
import sys
sys.path.append('..')
import pandas as pd
# %load_ext autoreload
# %autoreload 2
from fastseq.all import *
from fastai2.basics import *
from fastseq.models.nbeats import *
from fastseq.data.external import *
# hide
import wandb
wandb.init(project='FastSeq',entity='TakoTabak',name='index_page')
from fastai2.callback.wandb import *
path = untar_data(URLs.m4_daily)
data = TSDataLoaders.from_folder(path,lookback=28, horizon = 14, step=3)
# items = dummy_data_generator(50, 10, nrows=1000)
# data = TSDataLoaders.from_items(items, horizon = 7)
data.show_batch()
# TODO make custom learner with custom model
learn = nbeats_learner(data,layers=[512, 512], stack_types=("trend","seasonality"), b_loss=.4, nb_blocks_per_stack=3,
loss_func=CombinedLoss(F.mse_loss, smape, ratio = {'smape':.05})
)
from fastai2.callback.all import *
learn.lr_find()
# +
# mname = 'M4-600rows-512*2-5blocksperstack'
# learn.add_cbs([WandbCallback(),SaveModelCallback(fname=mname)])
# -
learn.fit_one_cycle(3, 1e-4)
learn.recorder.plot_loss()
learn.show_results(2,max_n=9)
# ## Interperation
from fastai2.interpret import *
from fastseq.interpret import *
interp = NBeatsInterpretation.from_learner(learn)
interp.plot_top_losses(3)
|
nbs/archive/_train_m4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from ecdsa import SigningKey, SECP256k1
import sha3
def eth_gen():
keccak = sha3.keccak_256()
priv = SigningKey.generate(curve=SECP256k1)
pub = priv.get_verifying_key().to_string()
keccak.update(pub)
address = keccak.hexdigest()[24:]
return {"Address":('0x'+address),"private_key":priv.to_string().hex()}
#print("Private key:", priv.to_string().hex())
#print("Public key: ", pub.hex())
#print("Address: 0x" + address)
print(eth_gen())
# -
import time
while True:
for j in range(100):
filename = str('test\#'+str(j)+'.txt')
for i in range(100):
with open(filename,'a') as f:
eth = eth_gen()
f.write('{:0>5} {} {}\n'.format(i, eth['Address'],eth['private_key']))
time.sleep(2)
print('over')
time.sleep(400)
|
eth_addr_gen.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sai8e/StockMarketProject/blob/main/GettingStockData.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="-8CQeuP5ha7i"
# #Getting a single stock data and exporting it as CSV
#
#
# + id="Lm52y550VgGr"
# !pip install yfinance
# + id="Is7tTotnTO0G"
import yfinance as yf
import pandas as pd
import os
def writeToCvs(ticker, beginningDate, endDate):
dt = yf.download(ticker,beginningDate, endDate)
dt.to_csv(r'./stock/'+ticker+'.csv')
return dt
# Get the data for the stock
newpath = r'./stock'
if not os.path.exists(newpath):
os.makedirs(newpath)
IBM_data = writeToCvs('IBM', '1911-01-01','2021-10-13')
print(IBM_data)
# + id="MGoCRkRTgSD_" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="63cdccd6-fb0d-4217-e65e-a45d297cf803"
import matplotlib.pyplot as plt
# %matplotlib inline
IBM_data['Adj Close'].plot()
plt.show()
# + [markdown] id="U_iPi2vih6-8"
# # Multiple Tickers and exporting it as CSV
#
# + id="ZAUKw53nusU1"
# <NAME> 30
tickers = pd.read_html(
'https://en.wikipedia.org/wiki/Dow_Jones_Industrial_Average#Components')[1]['Symbol'].values.tolist()
print(tickers)
for ticker in tickers:
writeToCvs(ticker, '2019-1-1', '2021-1-1')
# S&P 500
tickers500 = pd.read_html(
'https://en.wikipedia.org/wiki/List_of_S%26P_500_companies')[0]['Symbol'].values.tolist()
for ticker in tickers500:
writeToCvs(ticker, '2019-1-1', '2021-1-1')
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="ytZkJWX8hOGw" outputId="5eb58bca-b8f0-4fe6-a643-6c63c6b78464"
import pandas as pd
import yfinance as yf
data = yf.download(tickers,'2019-1-1')['Close']
print(data.head())
((data.pct_change()+1).cumprod()).plot(figsize=(20, 15))
# data.plot(figsize=(20,15))
plt.legend()
plt.title("Returns", fontsize=16)
plt.ylabel('Cumulative Returns', fontsize=14)
plt.xlabel('Year', fontsize=14)
plt.grid(which="major", color='k', linestyle='-.', linewidth=0.5)
plt.show()
|
GettingStockData.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #!/usr/bin/env python
import matplotlib.pyplot as plt
import matplotlib.colors as colors
import matplotlib.tri as tri
from mpl_toolkits.axes_grid1.anchored_artists import AnchoredSizeBar
from matplotlib import ticker, cm
import numpy as np
from numpy import ma
import csv
# +
degree_sign= u'\N{DEGREE SIGN}'
CSV_FILE_PATH = '../../../../Data/ISER2021/Sunapee-20200715-path-1.csv'
#CSV_FILE_PATH2 = '../../../../Data/ISER2021/Sunapee-20200715-path-2.csv'
#CSV_FILE_PATH3 = '../../../../Data/ISER2021/Sunapee-20200715-path-3.csv'
with open(CSV_FILE_PATH, 'r') as csv_file:
reader = csv.reader(csv_file)
path1_list = np.array(list(reader))
'''
with open(CSV_FILE_PATH2, 'r') as csv_file:
reader = csv.reader(csv_file)
path2_list = np.array(list(reader))
with open(CSV_FILE_PATH3, 'r') as csv_file:
reader = csv.reader(csv_file)
path3_list = np.array(list(reader))
'''
#=============================== 07/15 ===============================
# one independent
z = path1_list[0:2127,29]
z = z.astype('float32')
# gps x,y
x = path1_list[0:2127,2]
x = x.astype('float32')
y = path1_list[0:2127,1]
y = y.astype('float32')
"""
# PATH 1
# temp 23
# chloro 29
z1 = path1_list[0:2126,29]
z1 = z1.astype('float32')
# gps x,y
x1 = path1_list[0:2126,2]
x1 = x1.astype('float32')
y1 = path1_list[0:2126,1]
y1 = y1.astype('float32')
## PATH 2
# chloro 29
z2 = path2_list[0:998,29]
z2 = z2.astype('float32')
# gps x,y
x2 = path2_list[0:998,2]
x2 = x2.astype('float32')
y2 = path2_list[0:998,1]
y2 = y2.astype('float32')
## PATH 3
# chloro 29
z3 = path3_list[0:597,29]
z3 = z3.astype('float32')
# gps x,y
x3 = path3_list[0:597,2]
x3 = x3.astype('float32')
y3 = path3_list[0:597,1]
y3 = y3.astype('float32')
x = np.concatenate([x1, x2, x3])
y = np.concatenate([y1, y2, y3])
z = np.concatenate([z1, z2, z3])
"""
#=====================================================================
f, ax = plt.subplots()
#ax.set_title('Catabot 07/15 Sunapee path 1: Temperature (' + degree_sign + 'C)')
ax.set_title('Catabot 07/15 Sunapee paths: Chlorophyll (RFU)')
vmax=0.20
vmin=0.02
levels = np.linspace(vmin,vmax, 50) # original 20
cs = ax.tricontourf(x,y,z, 10, norm=colors.SymLogNorm(linthresh=0.03, linscale=0.03), levels=levels,vmax=vmax,vmin=vmin)
#cs = ax.tricontourf(x,y,z, 20, vmin=24.35, vmax=26.94)
#cs = ax.tricontourf(x,y,z, 20)
cb_ticklabel = np.linspace(vmin, vmax, 10)
cb = f.colorbar(cs, ticks=cb_ticklabel, orientation='horizontal', format='%.2f')
ax.set_xlabel('Longitude')
plt.xlim([-72.038, -72.0325])
ax.set_xticks(np.arange(-72.0375, -72.0324, 0.001))
f.canvas.draw()
ax.set_xticklabels(['-72.0375', '-72.0365', '-72.0355', '-72.0345', '-72.0335', '-72.0325'])
ax.set_ylabel('Latitude')
plt.ylim([43.4095, 43.4120])
ax.set_yticks(np.arange(43.4095, 43.4121, 0.0005))
f.canvas.draw()
ax.set_yticklabels(['43.4100', '43.4105', '43.4110', '43.4115', '43.4120', '43.4125'])
# path 1,2,3
ax.plot(x,y,marker='o', color='k', markersize=0.1)
#ax.plot(x2,y2,marker='o', color='b', markersize=0.1)
#ax.plot(x3,y3,marker='o', color='r', markersize=0.1)
ax.set_aspect('equal')
plt.grid(True)
# summer buoy
ax.plot(np.array([-72.033128]), np.array([43.4096079]), color='k', marker='o', markersize=13)
ax.plot(np.array([-72.033128]), np.array([43.4096079]), color='yellow', marker='o', markersize=8)
# boathouse
ax.plot(np.array([-72.0369625]), np.array([43.4100466]), color='k', marker=(5,1), markersize=16)
ax.plot(np.array([-72.0369625]), np.array([43.4100466]), color='#FF4500', marker=(5,1), markersize=8)
# winter buoy
ax.plot(np.array([-72.0365116]), np.array([43.410345]), color='k', marker='o', markersize=13)
ax.plot(np.array([-72.0365116]), np.array([43.410345]), color='m', marker='o', markersize=8)
bar = AnchoredSizeBar(ax.transData, 0.00046, '40 m', 'upper right', pad=0.6, frameon=False)
ax.add_artist(bar)
plt.show()
f.savefig('0715-path-onlyone-path-black.png', bbox_inches ='tight')
# to save only color bar
#ax.remove()
#cb.ax.xaxis.set_ticks_position('top')
#cb.ax.xaxis.set_label_position('top')
#cb.ax.tick_params(labelsize=14)
#cb.set_label(label='Temperature (' + degree_sign + 'C)', size=14)
#plt.show()
# -
min(z)
max(z)
|
Jupyter_notebook/ISER2021/Path cover/Chlorophyll/.ipynb_checkpoints/20200720-Sunapee-path-cover-chloro-path-black-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.2
# language: julia
# name: julia-1.5
# ---
# ## Print
println("That's me or Julia Feels like Python?")
print("That's me or Julia Feels like Python?")
println("That's me or Julia Feels like Python?")
println("That's me or Julia Feels like Python?")
print("That's me or Julia Feels like Python?")
print("ThaZt's me or Julia Feels like Python?")
show("Julia got its own way too")
write(stdout, "Julia got its own way too")
x = 7310302560386184563
print(x)
write(stdout, x)
|
Learn Julia/03. Print.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://training.dwit.edu.np/frontend/images/computer-training-institute.png">
#
# # Data Science and Machine learning in Python
#
# ### Instructor: [<NAME>](https://www.kaggle.com/atishadhikari)
# <hr>
# ### Principle Component Analysis
import numpy as np
import matplotlib.pyplot as plt
# +
sigma = [
[2, 0.8],
[0.8, 0.6]
]
meu = [1.2, 2.2]
# -
data = np.random.multivariate_normal(meu, sigma, 100)
plt.scatter(data[:, 0], data[:, 1])
plt.show()
# <img src="https://dataconomy.com/wp-content/uploads/2016/01/pic2.png">
# Step1: Standardize
#
# Step 2: Compute eigen values and eigen vectors of sigma
var_covar = np.cov(data.T)
eig_val, eig_vec = np.linalg.eig(var_covar)
# Step 3: Select no of components to reduce data into (k) by looking into eigen values
#
# Step 4: Select k-number of max-eigen values
eig_val
2.93 / (2.93 + 0.24)
eig_vec = eig_vec.T
# Step 5: Select k corresponding eigen vectors of selected eigen-values
pca1 = eig_vec[0]
# Step 6: Project into new eigen vectors
# (Apply dot product keeping shape in mind)
reduced_data = data.dot(pca1)
plt.scatter(reduced_data, np.zeros(100))
plt.show()
# #### PCA on Iris Dataset
from sklearn.datasets import load_iris
iris = load_iris()
X = iris.data
y = iris.target
sigma = np.cov(X.T)
sigma #var-covar matrix
eig_val, eig_vec = np.linalg.eig(sigma)
eig_vec = eig_vec.T
eig_val
(4.22824 + 0.24267) / np.sum(eig_val) #varience preserved by 2-dimension
new_axis = eig_vec[[0,1]] #select corresponding eigen-vectors
new_axis.shape
X.shape
reduced = X.dot(new_axis.T) #dot product keeping shape in mind
plt.scatter(reduced[y==0, 0], reduced[y==0, 1], color="r")
plt.scatter(reduced[y==1, 0], reduced[y==1, 1], color="g")
plt.scatter(reduced[y==2, 0], reduced[y==2, 1], color="b")
plt.show()
from sklearn.decomposition import PCA
pca = PCA(n_components=2)
pca.fit(X)
reduced = pca.transform(X)
plt.scatter(reduced[y==0, 0], reduced[y==0, 1], color="r")
plt.scatter(reduced[y==1, 0], reduced[y==1, 1], color="g")
plt.scatter(reduced[y==2, 0], reduced[y==2, 1], color="b")
plt.show()
pca.explained_variance_ratio_
#Total information preserved
np.sum(pca.explained_variance_ratio_)
|
ML_10.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 style="font-size:42px; text-align:center; margin-bottom:30px;"><span style="color:SteelBlue">Module 2
# :</span> ABT Construction</h1>
# <hr>
# Welcome to the workbook for <span style="color:royalblue">Module 2: ABT Construction</span>!
#
# In this module, we're going to combine the **Data Cleaning** and **Feature Engineering** steps from Project 2.
#
# Remember, **better data beats better algorithms**.
#
#
# <br><hr id="toc">
#
# ### In this module...
#
# In this module, we'll cover the essential steps for building your analytical base table:
# 1. [Drop unwanted observations](#drop)
# 2. [Fix structural errors](#structural)
# 3. [Handle missing data](#missing-data)
# 4. [Engineer features](#engineer-features)
# 5. [Save the ABT](#save-abt)
#
# Finally, we'll save the ABT to a new file so we can use it in other modules.
#
# <br><hr>
# ### First, let's import libraries and load the dataset.
#
# In general, it's good practice to keep all of your library imports at the top of your notebook or program.
#
# We've provided comments for guidance.
# +
# print_function for compatibility with Python 3
from __future__ import print_function
print('Print function ready to serve.')
# NumPy for numerical computing
import numpy as np
# Pandas for DataFrames
import pandas as pd
pd.set_option('display.max_columns', 100)
# Matplotlib for visualization
from matplotlib import pyplot as plt
# display plots in the notebook
# %matplotlib inline
# Seaborn for easier visualization
import seaborn as sns
# -
# Next, let's import the dataset.
# * The file path is <code style="color:crimson">'project_files/employee_data.csv'</code>
# Load employee data from CSV
df = pd.read_csv('project_files/employee_data.csv')
# Now we're ready to jump into cleaning the data!
# <span id="drop"></span>
# # 1. Drop wanted observations
#
# The first step to data cleaning is removing samples from your dataset that you don't want to include in the model.
#
# <br>
# **First, <span style="color:royalblue">drop duplicates</span> from the dataset.**
# * Then, print the shape of the new dataframe.
# Drop duplicates
df = df.drop_duplicates()
print(df.shape)
# **Display all of the unique classes of the <code style="color:steelblue">'department'</code> feature**
# Unique classes of 'department'
df.department.unique()
# **Drop all observations that belong to the <span style="color:crimson">'temp'</span> department.**
# * **Hint:** This is the same as keeping all that don't belong to that department.
# * **Hint:** Remember to overwrite your original dataframe.
# * Then, print the shape of the new dataframe.
# Drop temporary workers
df = df[df.department != 'temp']
# <p style="text-align:center; margin: 40px 0 40px 0; font-weight:bold">
# <a href="#toc">Back to Contents</a>
# </p>
# <span id="structural"></span>
# # 2. Fix structural errors
#
# The next bucket under data cleaning involves fixing structural errors, which arise during measurement, data transfer, or other types of "poor housekeeping."
#
# <br>
# **Print the unique values of <code style="color:steelblue">'filed_complaint'</code> and <code style="color:steelblue">'recently_promoted'</code>.**
# Print unique values of 'filed_complaint'
print('Filed Complaint:' + str(df.filed_complaint.unique()))
# Print unique values of 'recently_promoted'
print('Recently Promoted:' + str(df.recently_promoted.unique()))
# **Fill missing <code style="color:steelblue">'filed_complaint'</code> and <code style="color:steelblue">'recently_promoted'</code> values with <code style="color:crimson">0</code>.**
# Missing filed_complaint values should be 0
df['filed_complaint'] = df.filed_complaint.fillna(0)
# Missing recently_promoted values should be 0
df['recently_promoted'] = df.recently_promoted.fillna(0)
# **Print the unique values of <code style="color:steelblue">'filed_complaint'</code> and <code style="color:steelblue">'recently_promoted'</code> again, just to confirm.**
# Print unique values of 'filed_complaint'
print('Filed Complaint: ', df.filed_complaint.unique())
# Print unique values of 'recently_promoted'
print('Recently Promoted: ', df.recently_promoted.unique())
# **Replace any instances of <code style="color:crimson">'information_technology'</code> with <code style="color:crimson">'IT'</code> instead.**
# * Remember to do it **inplace**.
# * Then, plot the **bar chart** for <code style="color:steelblue">'department'</code> to see its new distribution.
# 'information_technology' should be 'IT'
df.department.replace(['information_technology'], 'IT', inplace=True)
# Plot class distributions for 'department'
sns.countplot(y='department', data=df)
plt.show()
# <p style="text-align:center; margin: 40px 0 40px 0; font-weight:bold">
# <a href="#toc">Back to Contents</a>
# </p>
# <span id="missing-data"></span>
# # 3. Handle missing data
#
# Next, it's time to handle **missing data**.
#
# <br>
# **Display the <span style="color:royalblue">number of missing values</span> for each feature (both categorical and numeric).**
# Display number of missing values by feature
df.isnull().sum()
# **Label missing values in <code style="color:steelblue">'department'</code> as <code style="color:crimson">'Missing'</code>.**
# * By the way, the <code style="color:steelblue">.fillna()</code> function also has an <code style="color:steelblue">inplace=</code> argument, just like the <code style="color:steelblue">.replace()</code> function.
# * In the previous project, we just overwrote that column. This time, try using the <code style="color:steelblue">inplace=</code> argument instead.
# Fill missing values in department with 'Missing'
df['department'].fillna('Missing', inplace=True)
# **First, let's flag <code style="color:steelblue">'last_evaluation'</code> with an indicator variable of missingness.**
# * <code style="color:crimson">0</code> if not missing.
# * <code style="color:crimson">1</code> if missing.
#
# Let's name the new indicator variable <code style="color:steelblue">'last_evaluation_missing'</code>.
# * We can use the <code style="color:steelblue">.isnull()</code> function.
# * Also, remember to convert it with <code style="color:steelblue">.astype(int)</code>
# Indicator variable for missing last_evaluation
df['last_evaluation_missing'] = df.last_evaluation.isnull().astype(int)
# **Then, simply fill in the original missing value with <code style="color:crimson">0</code> just so your algorithms can run properly.**
# Fill missing values in last_evaluation with 0
df.last_evaluation.fillna(0, inplace=True)
# **Display the number of missing values for each feature (both categorical and numeric) again, just to confirm.**
# Display number of missing values by feature
df.isnull().sum()
# <p style="text-align:center; margin: 40px 0 40px 0; font-weight:bold">
# <a href="#toc">Back to Contents</a>
# </p>
# <span id="engineer-features"></span>
# # 4. Engineer features
#
# For this project, we're going to have an abbreviated version of feature engineering, since we've already covered many tactics in Project 2.
#
# <br>
# Do you remember the scatterplot of <code style="color:steelblue">'satisfaction'</code> and <code style="color:steelblue">'last_evaluation'</code> for employees who have <code style="color:crimson">'Left'</code>?
#
# **Let's reproduce it here, just so we have it in front of us.**
# Scatterplot of satisfaction vs. last_evaluation, only those who have left
sns.lmplot(x='satisfaction', y='last_evaluation', data=df[df.status=='Left'], fit_reg=False)
plt.show()
# These roughly translate to 3 **indicator features** we can engineer:
#
# * <code style="color:steelblue">'underperformer'</code> - last_evaluation < 0.6 and last_evaluation_missing == 0
# * <code style="color:steelblue">'unhappy'</code> - satisfaction_level < 0.2
# * <code style="color:steelblue">'overachiever'</code> - last_evaluation > 0.8 and satisfaction > 0.7
#
# <br>
# **Create those 3 indicator features.**
# * Use boolean masks.
# * **Important:** For <code style="color:steelblue">'underperformer'</code>, it's important to include <code style="color:steelblue">'last_evaluation_missing' == 0</code> to avoid those originally missing observations that we flagged and filled.
# +
# Create indicator features
df['underperformer'] = ((df.last_evaluation < 0.6) & (df.last_evaluation_missing == 0)).astype(int)
df['unhappy'] = ((df.satisfaction) < 0.2).astype(int)
df['overachiever'] = (((df.last_evaluation > 0.8) & (df.satisfaction > 0.7))).astype(int)
# -
# <br>
# **Next, run this code to check that you created the features correctly.**
# The proportion of observations belonging to each group
df[['underperformer', 'unhappy', 'overachiever']].mean()
# <p style="text-align:center; margin: 40px 0 40px 0; font-weight:bold">
# <a href="#toc">Back to Contents</a>
# </p>
# <span id="save-abt"></span>
# # 5. Save the ABT
#
# Finally, let's save the **analytical base table**.
#
# <br>
# **Convert <code style="color:steelblue">'status'</code> into an indicator variable.**
# * <code style="color:crimson">'Left'</code> should be <code style="color:crimson">1</code>
# * <code style="color:crimson">'Employed'</code> should be <code style="color:crimson">0</code>
# * There's also a really easy way you can use <code style="color:steelblue">pd.get_dummies()</code> here. See below.
# Convert status to an indicator variable
df['status'] = pd.get_dummies( df.status ).Left
# **To confirm we did that correctly, display the proportion of people in our dataset who left.**
# The proportion of observations who 'Left'
df.status.mean()
# **Overwrite your dataframe with a version that has <span style="color:royalblue">dummy variables</span> for the categorical features.**
# * Then, display the first 10 rows to confirm all of the changes we've made so far in this module.
# Create new dataframe with dummy features
df = pd.get_dummies(df, columns=['department', 'salary'])
# Display first 10 rows
df.head(10)
# **Save this dataframe as your <span style="color:royalblue">analytical base table</span> to use in later modules.**
# * Remember to set the argument <code style="color:steelblue">index=None</code> to save only the data.
# Save analytical base table
df.to_csv('analytical_base_table.csv', index=None)
# <p style="text-align:center; margin: 40px 0 40px 0; font-weight:bold">
# <a href="#toc">Back to Contents</a>
# </p>
# <br>
# ## Next Steps
#
# Congratulations for making through Project 3's ABT Construction module!
#
# As a reminder, here are a few things you did in this module:
# * You cleaned dropped irrelevant observations from the dataset.
# * You fixed various structural errors, such as wannabe indicator variables.
# * You handled missing data.
# * You engineered features by leveraging your exploratory analysis.
# * And you created dummy variables before saving the ABT.
#
# In the next module, <span style="color:royalblue">Module 3: Classification Algorithms</span>, we'll take a detour from the project and dive into some more machine learning concepts. We'll also introduce the classification algorithms we'll be using for this project.
#
# <p style="text-align:center; margin: 40px 0 40px 0; font-weight:bold">
# <a href="#toc">Back to Contents</a>
# </p>
|
.ipynb_checkpoints/Module 2 - ABT Construction-checkpoint.ipynb
|
# + [markdown] colab_type="text" id="q4WF3l23pumU"
# ##### Copyright 2018 The AdaNet Authors.
# + cellView="both" colab={} colab_type="code" id="Kic2quJWppmx"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="aL7SpaKdirqG"
# # Customizing AdaNet
#
# Often times, as a researcher or machine learning practitioner, you will have
# some prior knowledge about a dataset. Ideally you should be able to encode that
# knowledge into your machine learning algorithm. With `adanet`, you can do so by
# defining the *neural architecture search space* that the AdaNet algorithm should
# explore.
#
# In this tutorial, we will explore the flexibility of the `adanet` framework, and
# create a custom search space for an image-classificatio dataset using high-level
# TensorFlow libraries like `tf.layers`.
#
#
# + colab={"test": {"output": "ignore", "timeout": 900}} colab_type="code" id="x_3b6xx2s6B9"
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import functools
import adanet
from adanet.examples import simple_dnn
import tensorflow as tf
# The random seed to use.
RANDOM_SEED = 42
# + [markdown] colab_type="text" id="7gE5Mm9j2oYw"
# ## Fashion MNIST dataset
#
# In this example, we will use the Fashion MNIST dataset
# [[Xiao et al., 2017](https://arxiv.org/abs/1708.07747)] for classifying fashion
# apparel images into one of ten categories:
#
# 1. T-shirt/top
# 2. Trouser
# 3. Pullover
# 4. Dress
# 5. Coat
# 6. Sandal
# 7. Shirt
# 8. Sneaker
# 9. Bag
# 10. Ankle boot
#
# 
# + [markdown] colab_type="text" id="5_hRtdchqRZb"
# ## Download the data
#
# Conveniently, the data is available via Keras:
# + colab={"test": {"output": "ignore", "timeout": 900}} colab_type="code" id="uYklOnPJ4h7g"
(x_train, y_train), (x_test, y_test) = (
tf.keras.datasets.fashion_mnist.load_data())
# + [markdown] colab_type="text" id="tECo5dFd4QCa"
# ## Supply the data in TensorFlow
#
# Our first task is to supply the data in TensorFlow. Using the
# tf.estimator.Estimator covention, we will define a function that returns an
# `input_fn` which returns feature and label `Tensors`.
#
# We will also use the `tf.data.Dataset` API to feed the data into our models.
# + colab={"test": {"output": "ignore", "timeout": 900}} colab_type="code" id="gxTAoIXwsTH7"
FEATURES_KEY = "images"
def generator(images, labels):
"""Returns a generator that returns image-label pairs."""
def _gen():
for image, label in zip(images, labels):
yield image, label
return _gen
def preprocess_image(image, label):
"""Preprocesses an image for an `Estimator`."""
# First let's scale the pixel values to be between 0 and 1.
image = image / 255.
# Next we reshape the image so that we can apply a 2D convolution to it.
image = tf.reshape(image, [28, 28, 1])
# Finally the features need to be supplied as a dictionary.
features = {FEATURES_KEY: image}
return features, label
def input_fn(partition, training, batch_size):
"""Generate an input_fn for the Estimator."""
def _input_fn():
if partition == "train":
dataset = tf.data.Dataset.from_generator(
generator(x_train, y_train), (tf.float32, tf.int32), ((28, 28), ()))
else:
dataset = tf.data.Dataset.from_generator(
generator(x_test, y_test), (tf.float32, tf.int32), ((28, 28), ()))
# We call repeat after shuffling, rather than before, to prevent separate
# epochs from blending together.
if training:
dataset = dataset.shuffle(10 * batch_size, seed=RANDOM_SEED).repeat()
dataset = dataset.map(preprocess_image).batch(batch_size)
iterator = dataset.make_one_shot_iterator()
features, labels = iterator.get_next()
return features, labels
return _input_fn
# + [markdown] colab_type="text" id="vm9yudEv5lQZ"
# ## Establish baselines
#
# The next task should be to get somes baselines to see how our model performs on
# this dataset.
#
# Let's define some information to share with all our `tf.estimator.Estimators`:
# + colab={"test": {"output": "ignore", "timeout": 900}} colab_type="code" id="xNwSUWh-9_Ib"
# The number of classes.
NUM_CLASSES = 10
# We will average the losses in each mini-batch when computing gradients.
loss_reduction = tf.losses.Reduction.SUM_OVER_BATCH_SIZE
# A `Head` instance defines the loss function and metrics for `Estimators`.
head = tf.contrib.estimator.multi_class_head(
NUM_CLASSES, loss_reduction=loss_reduction)
# Some `Estimators` use feature columns for understanding their input features.
feature_columns = [
tf.feature_column.numeric_column(FEATURES_KEY, shape=[28, 28, 1])
]
# Estimator configuration.
config = tf.estimator.RunConfig(
save_checkpoints_steps=50000,
save_summary_steps=50000,
tf_random_seed=RANDOM_SEED)
# + [markdown] colab_type="text" id="QY0cv-ot-Gxs"
# Let's start simple, and train a linear model:
# + cellView="both" colab={"height": 53, "test": {"output": "ignore", "timeout": 900}} colab_type="code" executionInfo={"elapsed": 32813, "status": "ok", "timestamp": 1534440488365, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="s8wJKsi06blX" outputId="5acd325b-1de9-4c21-d825-8f9c10ffdd6c"
#@test {"skip": true}
#@title Parameters
LEARNING_RATE = 0.001 #@param {type:"number"}
TRAIN_STEPS = 5000 #@param {type:"integer"}
BATCH_SIZE = 64 #@param {type:"integer"}
estimator = tf.estimator.LinearClassifier(
feature_columns=feature_columns,
n_classes=NUM_CLASSES,
optimizer=tf.train.RMSPropOptimizer(learning_rate=LEARNING_RATE),
loss_reduction=loss_reduction,
config=config)
results, _ = tf.estimator.train_and_evaluate(
estimator,
train_spec=tf.estimator.TrainSpec(
input_fn=input_fn("train", training=True, batch_size=BATCH_SIZE),
max_steps=TRAIN_STEPS),
eval_spec=tf.estimator.EvalSpec(
input_fn=input_fn("test", training=False, batch_size=BATCH_SIZE),
steps=None))
print("Accuracy:", results["accuracy"])
print("Loss:", results["average_loss"])
# + [markdown] colab_type="text" id="a-1hE03c7_Yj"
# The linear model with default parameters achieves about **84.13% accuracy**.
#
# Let's see if we can do better with the `simple_dnn` AdaNet:
# + colab={"height": 53, "test": {"output": "ignore", "timeout": 900}} colab_type="code" executionInfo={"elapsed": 51048, "status": "ok", "timestamp": 1534440539502, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="9fAoRYd19eUs" outputId="0b5678f1-4d44-430b-d84d-d3315366d4b8"
#@test {"skip": true}
#@title Parameters
LEARNING_RATE = 0.003 #@param {type:"number"}
TRAIN_STEPS = 5000 #@param {type:"integer"}
BATCH_SIZE = 64 #@param {type:"integer"}
ADANET_ITERATIONS = 2 #@param {type:"integer"}
estimator = adanet.Estimator(
head=head,
subnetwork_generator=simple_dnn.Generator(
feature_columns=feature_columns,
optimizer=tf.train.RMSPropOptimizer(learning_rate=LEARNING_RATE),
seed=RANDOM_SEED),
max_iteration_steps=TRAIN_STEPS // ADANET_ITERATIONS,
evaluator=adanet.Evaluator(
input_fn=input_fn("train", training=False, batch_size=BATCH_SIZE),
steps=None),
config=config)
results, _ = tf.estimator.train_and_evaluate(
estimator,
train_spec=tf.estimator.TrainSpec(
input_fn=input_fn("train", training=True, batch_size=BATCH_SIZE),
max_steps=TRAIN_STEPS),
eval_spec=tf.estimator.EvalSpec(
input_fn=input_fn("test", training=False, batch_size=BATCH_SIZE),
steps=None))
print("Accuracy:", results["accuracy"])
print("Loss:", results["average_loss"])
# + [markdown] colab_type="text" id="ysWsJ3zXDwNx"
# The `simple_dnn` AdaNet model with default parameters achieves about **85.66%
# accuracy**.
#
# This improvement can be attributed to `simple_dnn` searching over
# fully-connected neural networks which have more expressive power than the linear
# model due to their non-linear activations.
#
# Fully-connected layers are permutation invariant to their inputs, meaning that
# if we consistently swapped two pixels before training, the final model would
# perform identically. However, there is spatial and locality information in
# images that we should try to capture. Applying a few convolutions to our inputs
# will allow us to do so, and that will require defining a custom
# `adanet.subnetwork.Builder` and `adanet.subnetwork.Generator`.
# + [markdown] colab_type="text" id="D3IE6-9vFVlg"
# ## Define a convolutional AdaNet model
#
# Creating a new search space for AdaNet to explore is straightforward. There are
# two abstract classes you need to extend:
#
# 1. `adanet.subnetwork.Builder`
# 2. `adanet.subnetwork.Generator`
#
# Similar to the tf.estimator.Estimator `model_fn`, `adanet.subnetwork.Builder`
# allows you to define your own TensorFlow graph for creating a neural network,
# and specify the training operations.
#
# Below we define one that applies a 2D convolution, max-pooling, and then a
# fully-connected layer to the images:
# + colab={"test": {"output": "ignore", "timeout": 900}} colab_type="code" id="IsYJ97tRwBkt"
class SimpleCNNBuilder(adanet.subnetwork.Builder):
"""Builds a CNN subnetwork for AdaNet."""
def __init__(self, learning_rate, max_iteration_steps, seed):
"""Initializes a `SimpleCNNBuilder`.
Args:
learning_rate: The float learning rate to use.
max_iteration_steps: The number of steps per iteration.
seed: The random seed.
Returns:
An instance of `SimpleCNNBuilder`.
"""
self._learning_rate = learning_rate
self._max_iteration_steps = max_iteration_steps
self._seed = seed
def build_subnetwork(self,
features,
logits_dimension,
training,
iteration_step,
summary,
previous_ensemble=None):
"""See `adanet.subnetwork.Builder`."""
images = features.values()[0]
kernel_initializer = tf.keras.initializers.he_normal(seed=self._seed)
x = tf.layers.conv2d(
images,
filters=16,
kernel_size=3,
padding="same",
activation="relu",
kernel_initializer=kernel_initializer)
x = tf.layers.max_pooling2d(x, pool_size=2, strides=2)
x = tf.layers.flatten(x)
x = tf.layers.dense(
x, units=64, activation="relu", kernel_initializer=kernel_initializer)
# The `Head` passed to adanet.Estimator will apply the softmax activation.
logits = tf.layers.dense(
x, units=10, activation=None, kernel_initializer=kernel_initializer)
# Use a constant complexity measure, since all subnetworks have the same
# architecture and hyperparameters.
complexity = tf.constant(1)
return adanet.Subnetwork(
last_layer=x,
logits=logits,
complexity=complexity,
persisted_tensors={})
def build_subnetwork_train_op(self,
subnetwork,
loss,
var_list,
labels,
iteration_step,
summary,
previous_ensemble=None):
"""See `adanet.subnetwork.Builder`."""
# Momentum optimizer with cosine learning rate decay works well with CNNs.
learning_rate = tf.train.cosine_decay(
learning_rate=self._learning_rate,
global_step=iteration_step,
decay_steps=self._max_iteration_steps)
optimizer = tf.train.MomentumOptimizer(learning_rate, .9)
# NOTE: The `adanet.Estimator` increments the global step.
return optimizer.minimize(loss=loss, var_list=var_list)
def build_mixture_weights_train_op(self, loss, var_list, logits, labels,
iteration_step, summary):
"""See `adanet.subnetwork.Builder`."""
return tf.no_op("mixture_weights_train_op")
@property
def name(self):
"""See `adanet.subnetwork.Builder`."""
return "simple_cnn"
# + [markdown] colab_type="text" id="OFamPrZHJ5ii"
# Next, we extend a `adanet.subnetwork.Generator`, which defines the search
# space of candidate `SimpleCNNBuilders` to consider including the final network.
# It can create one or more at each iteration with different parameters, and the
# AdaNet algorithm will select the candidate that best improves the overall neural
# network's `adanet_loss` on the training set.
#
# The one below is very simple: it always creates the same architecture, but gives
# it a different random seed at each iteration:
# + colab={"test": {"output": "ignore", "timeout": 900}} colab_type="code" id="-BAnb_XGwhRy"
class SimpleCNNGenerator(adanet.subnetwork.Generator):
"""Generates a `SimpleCNN` at each iteration.
"""
def __init__(self, learning_rate, max_iteration_steps, seed=None):
"""Initializes a `Generator` that builds `SimpleCNNs`.
Args:
learning_rate: The float learning rate to use.
max_iteration_steps: The number of steps per iteration.
seed: The random seed.
Returns:
An instance of `Generator`.
"""
self._seed = seed
self._dnn_builder_fn = functools.partial(
SimpleCNNBuilder,
learning_rate=learning_rate,
max_iteration_steps=max_iteration_steps)
def generate_candidates(self, previous_ensemble, iteration_number,
previous_ensemble_reports, all_reports):
"""See `adanet.subnetwork.Generator`."""
seed = self._seed
# Change the seed according to the iteration so that each subnetwork
# learns something different.
if seed is not None:
seed += iteration_number
return [self._dnn_builder_fn(seed=seed)]
# + [markdown] colab_type="text" id="8sdvharsLJ1T"
# With these defined, we pass them into a new `adanet.Estimator`:
# + colab={"height": 53, "test": {"output": "ignore", "timeout": 900}} colab_type="code" executionInfo={"elapsed": 74112, "status": "ok", "timestamp": 1534440614010, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 240} id="-Fhi1SjkzVBt" outputId="fd4b8c9c-4665-473b-ea7d-cd47c8680811"
#@title Parameters
LEARNING_RATE = 0.05 #@param {type:"number"}
TRAIN_STEPS = 5000 #@param {type:"integer"}
BATCH_SIZE = 64 #@param {type:"integer"}
ADANET_ITERATIONS = 2 #@param {type:"integer"}
max_iteration_steps = TRAIN_STEPS // ADANET_ITERATIONS
estimator = adanet.Estimator(
head=head,
subnetwork_generator=SimpleCNNGenerator(
learning_rate=LEARNING_RATE,
max_iteration_steps=max_iteration_steps,
seed=RANDOM_SEED),
max_iteration_steps=max_iteration_steps,
evaluator=adanet.Evaluator(
input_fn=input_fn("train", training=False, batch_size=BATCH_SIZE),
steps=None),
report_materializer=adanet.ReportMaterializer(
input_fn=input_fn("train", training=False, batch_size=BATCH_SIZE),
steps=None),
adanet_loss_decay=.99,
config=config)
results, _ = tf.estimator.train_and_evaluate(
estimator,
train_spec=tf.estimator.TrainSpec(
input_fn=input_fn("train", training=True, batch_size=BATCH_SIZE),
max_steps=TRAIN_STEPS),
eval_spec=tf.estimator.EvalSpec(
input_fn=input_fn("test", training=False, batch_size=BATCH_SIZE),
steps=None))
print("Accuracy:", results["accuracy"])
print("Loss:", results["average_loss"])
# + [markdown] colab_type="text" id="3wGtI-4_LRw1"
# Our `SimpleCNNGenerator` code achieves **90.41% accuracy**.
# + [markdown] colab_type="text" id="TKhCzP65hGyS"
# ## Conclusion and next steps
#
# In this tutorial, you learned how to customize `adanet` to encode your
# understanding of a particular dataset, and explore novel search spaces with
# AdaNet.
#
# One use-case that has worked for us at Google, has been to take a production
# model's TensorFlow code, convert it to into an `adanet.subnetwork.Builder`, and
# adaptively grow it into an ensemble. In many cases, this has given significant
# performance improvements.
#
# As an exercise, you can swap out the FASHION-MNIST with the MNIST handwritten
# digits dataset in this notebook using `tf.keras.datasets.mnist.load_data()`, and
# see how `SimpleCNN` performs.
|
adanet/examples/tutorials/customizing_adanet.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## _Inference after GNN Stage_
#
# **_Inference_** is done using callbacks defined in the **_LightningModules/GNN/Models/inference.py_**. The callbacks run during the **_test_step()_** _a.k.a_ model _**evalution**_.
#
# ### How to run _Inference_?
#
# 1. _`traintrack config/pipeline_quickstart.yaml`_. One can use `--inference` flag to run only the test_step() (Should work, but failed.)
# 2. _`infer.ipynb`_ notebook runs the _pl.Trainer().test()_
import sys, os, glob, yaml
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import pprint
import seaborn as sns
import trackml.dataset
import torch
from torch_geometric.data import Data
import itertools
device = 'cuda' if torch.cuda.is_available() else 'cpu'
os.environ['EXATRKX_DATA'] = os.path.abspath(os.curdir)
# + [markdown] tags=[]
# ## _Classifier Evaluation_
#
# Metrics to evaluate the GNN networks:
#
# [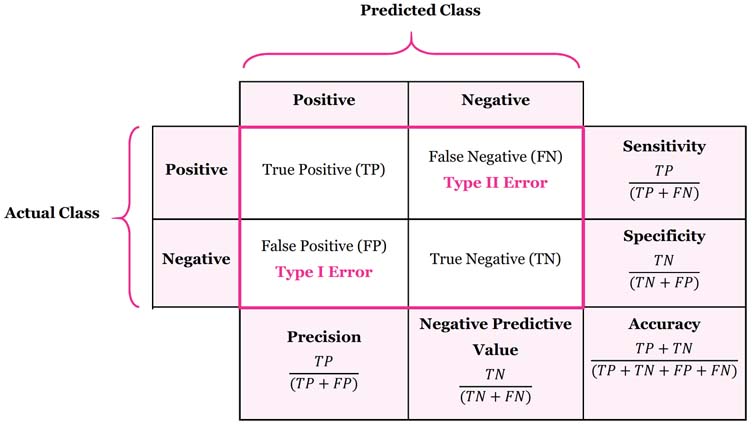](https://postimg.cc/z3zVdk3x)
#
#
#
# - Accuracy/ACC = $TP+TN/TP+TN+FP+FN$
# - sensitivity, recall, hit rate, or true positive rate ($TPR = 1 - FNR$)
# - specificity, selectivity or true negative rate ($TNR = 1 - FPR$)
# - miss rate or false negative rate ($FNR = 1 - TPR$)
# - fall-out or false positive rate ($FPR = 1 - TNR$)
# - F1-score = $2 \times (\text{PPV} \times \text{TPR})/(\text{PPV} + \text{TPR})$
# - Efficiency/Recall/Sensitivity/Hit Rate: $TPR = TP/(TP+FN)$
# - Purity/Precision/Positive Predictive Value: $PPV = TP/(TP+FP$
# - AUC-ROC Curve $\equiv$ FPR ($x-$axis) v.s. TPR ($y-$axis) plot
# - AUC-PRC Curve $\equiv$ TPR ($x-$axis) v.s. PPV ($y-$axis) plot
#
# -
import pytorch_lightning as pl
from LightningModules.GNN.Models.interaction_gnn import InteractionGNN
from LightningModules.GNN.Models.inference import GNNTelemetry, GNNMetrics
# ### _(+) - Test Dataset_
inputdir="run/gnn_evaluation/test"
testset = sorted(glob.glob(os.path.join(inputdir, "*")))
feature_data = torch.load(testset[0], map_location=device)
feature_data
# + [markdown] id="HfvKUCvjUlQA"
# ### _(+) - Load Model_
#
# - Load model checkpoint
# - Load saved config file
# -
ckpnt_path = "run/lightning_models/lightning_checkpoints/GNNStudy/version_1/checkpoints/last.ckpt"
# Model Checkpoint
checkpoint = torch.load(ckpnt_path, map_location=device)
# Checkpointing Hyperparameters
print(checkpoint["hyper_parameters"])
# Get the checkpoint config
config = checkpoint["hyper_parameters"]
# Modified defaults with custom settings
config["checkpoint_path"] = ckpnt_path
config["input_dir"] = "run/feature_store"
config["output_dir"] = "run/gnn_evaluation"
config["artifact_library"] = "lightning_models/lightning_checkpoints"
config["datatype_split"] = [90000, 5000, 5000]
config["callbacks"] = ['GNNTelemetry']
config["map_location"] = device
# Checkpointing Hyperparameters, with Corrections
print(checkpoint["hyper_parameters"])
# Init the InteractionGNN
model = InteractionGNN(config)
# 'config' has all needed info., just pass it as **kwargs
model = model.load_from_checkpoint(**config)
# Load Trainer
trainer = pl.Trainer(callbacks=[GNNMetrics()])
# Test Loop
results = trainer.test(model=model, verbose=True)
|
stt3_infer.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/xpertdesh/ml-class21/blob/main/labs/word_embeddings.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="L8bIz-dx6t3l" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="b1d74429-75ce-4ca4-cd17-61043021b2db"
import keras
keras.__version__
# + [markdown] id="fcwCWVoR6t3q"
# # Using word embeddings
#
#
# ## Meaning
# For hundreds of years linguists have been investigating how to encode the semantic information of a word. Some belief that there is a universal mental concept, for example `MOON`, with some universal representation in our brains that gets represented in English as *moon* in Uyghur as ئاي, as *Hanhepi wi* in Lakota, and as
# 月 in Japanese. It is also believed that concepts such as `MOON` and `SUN` share some common semantic features. Similar words would cluster together in semantic space. For example, moon, sun, Mars, Neptune, the Milky Way Galaxy seem to have some similarities and form a group as does a group we could label domestic animals that might include horses, dogs, and cats, which in turn are different from wild animals. So words like *dog*, *cat*, *horse*, have, as part of their semantic meaning, some feature that is the same among these words and that feature is not present in the representations for the words *moon* or *sun*. These semantic features impact how a word occurs in different contexts.
#
# For example, it would be rare for a verb to follow the word *the*:
#
# * *The says ...
# * *The haven't
# * *The see
#
# (Here the asterisk * represents a sentence most people would find odd). And again, this is just probability. Certainly, you can construct sentences that have these sequences that are perfectly fine (*The says who questioning method .*, *Just experimenting with the Says Who basics on myself on with clients*, *The haven't had time excuse*). But the likelihood of the word *says* following *the* is extremely rare.
#
# Similarly, sentences like
#
# * I fed my ___
# * I fed my dog
# * I fed my poodle
# * I fed my cat
# * I fed my horse
#
# occur much more frequently than
#
# * I fed my ___
# * I fed my moon
# * I fed my sun
# * I fed my Neptune
# * I fed my Milky Way Galaxy.
#
# So, we come to the quote by Firth:
#
# > You shall know a word by the company it keeps
#
# The idea is a simple one. If we find a bunch of words that can follow the sequence *I fed my* we can assume that they share some semantic feature. And, if we didn't know English we could still do this analysis. We wouldn't know that the feature is domestic animal but we would know that they share some feature, let's call it x29. This, as we talked about is a latent feature.
#
# Now, without going into any detail whatsoever, we can imagine giving an algorithm a gigaword corpus and telling it to come up with 100 semantic features that explain the distribution of the words in the corpus.
#
# That is what **word embeddings** are in their simplist form.
#
#
# ### Sparse vs. dense dimensions.
# In our previous work with text, we determined the size of our vocabulary, say 10,000 words, and created a vector where each column represented a different word in the vocabulary. So let's say column 1 was *a* and 2 *the*, and 7,253 *computer* and so on. If we encode a sentence that starts *the computer*, the word *the* would be represented by a '1' in column 1 and zeroes in the other 9,999 columns. The word *computer* would have a '1' in column 7,253 and zeroes in the other 9,999. That is an awful lot of zeroes, and this vector is called **sparse**. This looks like the first image below, where the blue square represents a '1' and the black represents all zeroes.
#
#
# + [markdown] id="cH3xchp86t3r"
#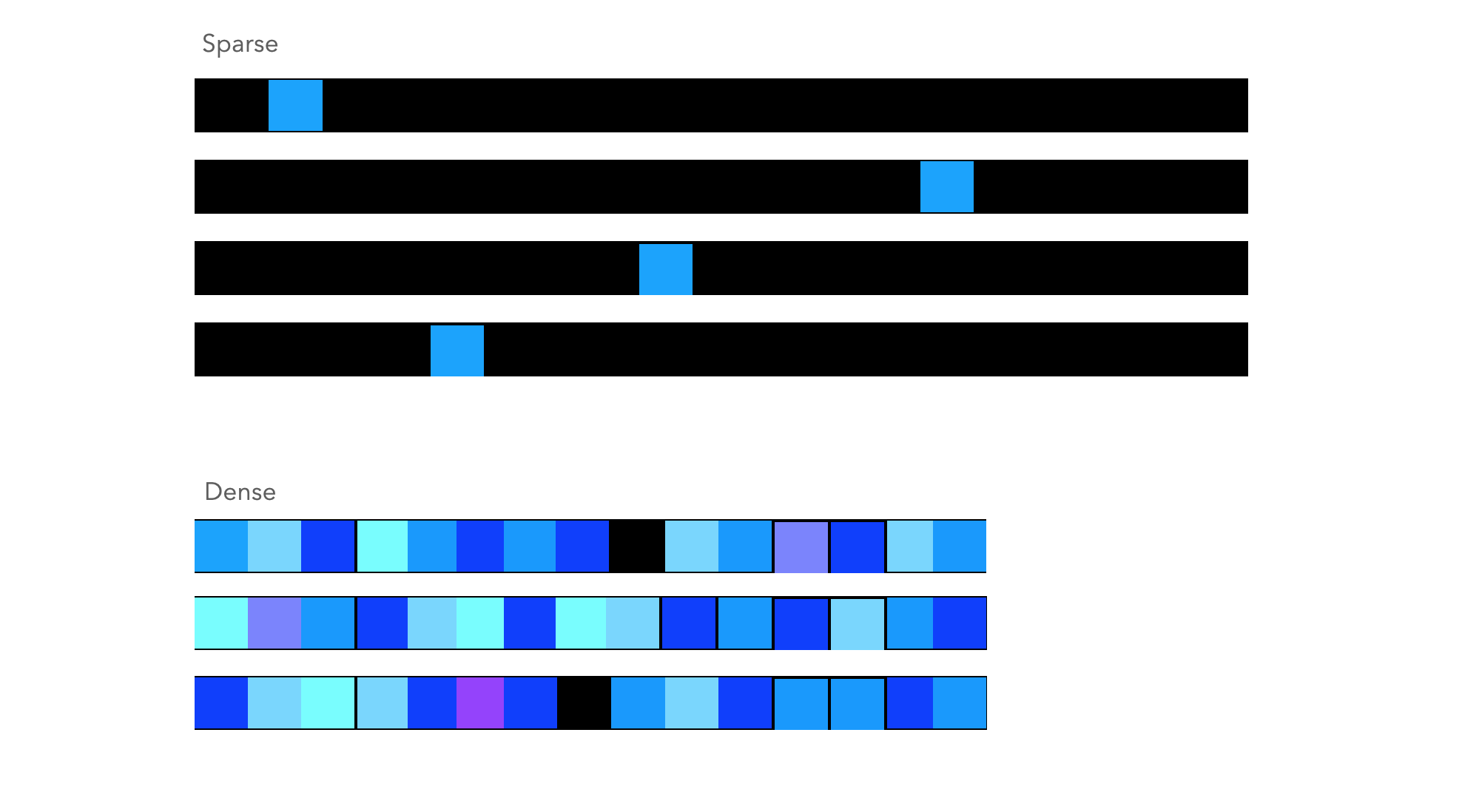 
# + [markdown] id="M3l4_QvP6t3r"
# With word embeddings we determine a feature size, 100, 200, 300 dimensions, and then each word is represented by a vector with values between 0 and 1. This is shown in the dense diagram above where the different colors represent different fractional values.
#
# In the one-hot-encoding method, each word was represented by a sparse vector the size of the vocabulary and vectors of 10,000 or 20,000 entries are not uncommon. In contrast, in the word embedding method, each word is represented by a dense vector of only 100 or 300 values. Thus, in the word embedding approach information is packed into a much smaller vector.
#
# In addition, in the one-hot encoding method there were no relationships between words. For example, *poodle* might be word 9,712 and *dog* 1,797, and they were treated completely separately. In the word embedding approach, the similarities of *poodle* and *dog* are represented within the word embeddings.
#
#
# #### Once again for emphasis
# While the vectors obtained through one-hot encoding are binary, sparse (mostly made of zeros) and very high-dimensional (same dimensionality as the
# number of words in the vocabulary), "word embeddings" are low-dimensional floating point vectors
#
#
# #### Note
# The following notebook is a remix of one by <NAME> (see the end of the notebook for more information)
#
#
# ### Obtaining word embeddings
# There are two ways to obtain word embeddings:
#
# * Learn word embeddings jointly with the main task you care about (e.g. document classification or sentiment prediction).
# In this setup, you would start with random word vectors, then learn your word vectors in the same way that you learn the weights of a neural network.
# * Load into your model word embeddings that were pre-computed using a different machine learning task than the one you are trying to solve.
# These are called "pre-trained word embeddings".
#
# Let's take a look at both.
# + [markdown] id="NbhGdm0e6t3s"
# ## Learning word embeddings with the `Embedding` layer
#
#
# The simplest way to associate a dense vector to a word would be to pick the vector at random. The problem with this approach is that the
# resulting embedding space would have no structure: for instance, the words "accurate" and "exact" may end up with completely different
# embeddings, even though they are interchangeable in most sentences. It would be very difficult for a deep neural network to make sense of
# such a noisy, unstructured embedding space.
#
# To get a bit more abstract: the geometric relationships between word vectors should reflect the semantic relationships between these words.
# Word embeddings are meant to map human language into a geometric space. For instance, in a reasonable embedding space, we would expect
# synonyms to be embedded into similar word vectors, and in general we would expect the geometric distance (e.g. L2 distance) between any two
# word vectors to relate to the semantic distance of the associated words (words meaning very different things would be embedded to points
# far away from each other, while related words would be closer). Even beyond mere distance, we may want specific __directions__ in the
# embedding space to be meaningful.
#
#
#
# In real-world word embedding spaces, common examples of meaningful geometric transformations are "gender vectors" and "plural vector". For
# instance, by adding a "female vector" to the vector "king", one obtain the vector "queen". By adding a "plural vector", one obtain "kings".
# Word embedding spaces typically feature thousands of such interpretable and potentially useful vectors.
#
# Is there some "ideal" word embedding space that would perfectly map human language and could be used for any natural language processing
# task? Possibly, but in any case, we have yet to compute anything of the sort. Also, there isn't such a thing as "human language", there are
# many different languages and they are not isomorphic, as a language is the reflection of a specific culture and a specific context. But more
# pragmatically, what makes a good word embedding space depends heavily on your task: the perfect word embedding space for an
# English-language movie review sentiment analysis model may look very different from the perfect embedding space for an English-language
# legal document classification model, because the importance of certain semantic relationships varies from task to task.
#
# It is thus reasonable to __learn__ a new embedding space with every new task. Thankfully, backpropagation makes this really easy, and Keras makes it
# even easier. It's just about learning the weights of a layer: the `Embedding` layer.
# + id="81dFNJsz6t3s"
from keras.layers import Embedding
# The Embedding layer takes at least two arguments:
# the number of possible tokens, here 1000 (1 + maximum word index),
# and the dimensionality of the embeddings, here 64.
embedding_layer = Embedding(1000, 64)
# + [markdown] id="OUqdtrTj6t3v"
#
# The `Embedding` layer is best understood as a dictionary mapping integer indices (which stand for specific words) to dense vectors. It takes
# as input integers, it looks up these integers into an internal dictionary, and it returns the associated vectors. It's effectively a dictionary lookup.
# + [markdown] id="-tPFE8U-6t3w"
#
# The `Embedding` layer takes as input a 2D tensor of integers, of shape `(samples, sequence_length)`, where each entry is a sequence of
# integers. So, for example, we may have a dataset of 10,000 tweets so that 10,000 is the `samples` and each sample consists of a sequence of integers representing the words in the tweet. It can embed sequences of variable lengths, so for instance we could feed into our embedding layer above batches that could have
# shapes `(32, 10)` (batch of 32 sequences of length 10) or `(64, 15)` (batch of 64 sequences of length 15). All sequences in a batch must
# have the same length, though (since we need to pack them into a single tensor), so sequences that are shorter than others should be padded
# with zeros, and sequences that are longer should be truncated.
#
# This layer returns a 3D floating point tensor, of shape `(samples, sequence_length, embedding_dimensionality)`.
#
# When you instantiate an `Embedding` layer, its weights (its internal dictionary of token vectors) are initially random, just like with any
# other layer. During training, these word vectors will be gradually adjusted via backpropagation, structuring the space into something that the
# downstream model can exploit. Once fully trained, your embedding space will show a lot of structure -- a kind of structure specialized for
# the specific problem you were training your model for.
#
# Let's apply this idea to the IMDB movie review sentiment prediction task that you are already familiar with. Let's quickly prepare
# the data. We will restrict the movie reviews to the top 20,000 most common words
# and cut the reviews after only 50 words. Our network will simply learn 50-dimensional embeddings for each of the 20,000 words, turn the
# input integer sequences (2D integer tensor) into embedded sequences (3D float tensor), flatten the tensor to 2D, and train a single `Dense`
# layer on top for classification.
# + id="h3ORYjSkkdo9" colab={"base_uri": "https://localhost:8080/"} outputId="c5551772-0e2f-40a6-992e-472eb8618fa7"
# !wget http://zacharski.org/files/courses/cs419/imdb.zip
# !unzip imdb.zip
# + id="Sc_HJW6lov9Q" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="a857c654-1024-4637-bf85-d937442e9170"
import pandas as pd
data = pd.read_csv('imdb.csv')
data
# + id="dU805Bd7pAOO" colab={"base_uri": "https://localhost:8080/"} outputId="2fc59ad3-f1af-4045-987e-f7a16d472865"
data_text = data.review
data_label = data.sentiment
data_label = data['sentiment'].apply(lambda x: 1 if x == 'positive' else 0)
data_label
# + [markdown] id="tvb9ry5-o_6u"
# ### Converting words to integers
# The first step we will do is convert words represented as strings to integers
#
# We will use the Keras Tokenizer and specify the size of the vocabulary.
#
# * num_words: the maximum number of words to keep, based on word frequency. Only the most common num_words-1 words will be kept.
#
# + id="DB6KIHrZpBOk"
from keras.preprocessing.text import Tokenizer
tokenizer = Tokenizer(num_words = 20000)
#build the word index
tokenizer.fit_on_texts(data_text)
# now convert the words to integers
data_sequences = tokenizer.texts_to_sequences(data_text)
data_sequences[0]
# + [markdown] id="y8zymRGLrOtA"
# Just for grins, let's convert an IMDB review back to text
#
#
# + id="b0SOKyfKrri7" colab={"base_uri": "https://localhost:8080/"} outputId="00db447f-5503-4e66-8a2d-62321b385cbf"
tokenizer.sequences_to_texts([data_sequences[0]])
# + [markdown] id="xAs2Yc6HvSya"
# ### Recap.
# We stated we wanted the vocabulary size to be 20,000. This means that if we have a sentence like *xenophobic manifestations moved online* and *xenophobic* was not among the 20,000 most frequent words in the corpus, our encoding of that sentence would skip that word. *xenophobic manifestations moved online* becomes *manifestations moved online*
#
# Now let's do a bit more preprocessing and truncate each review after 50 words (if the review is shorter than 50 words we will pad it with blank words):
# + id="lHpnZnf-6t3w" colab={"base_uri": "https://localhost:8080/"} outputId="238f79ea-40e4-4c1b-f7ad-f55e484f0ce9"
# Cut texts after this number of words
# (among top max_features most common words)
maxlen = 50
from keras import preprocessing
# This turns our lists of integers
# into a 2D integer tensor of shape `(samples, maxlen)`
data50 = preprocessing.sequence.pad_sequences(data_sequences, maxlen=maxlen)
data50[0]
# + [markdown] id="zhLEQdtOwrOC"
# # Now let's divide the data into training and testing
#
#
# + id="GeKwllN9w0dR" colab={"base_uri": "https://localhost:8080/"} outputId="0b9b60df-5463-4740-9991-919dad8e1a11"
from sklearn.model_selection import train_test_split
train50, test50, train_labels, test_labels = train_test_split(data50, data_label, test_size = 0.2, random_state=42)
test_labels
# + id="si1znRAtJj-b" colab={"base_uri": "https://localhost:8080/"} outputId="b0d08ec4-74d7-493b-bfdd-7c8d319d2c23"
test50.shape
# + id="_JBgVx0SZqls" outputId="2de80e3c-b03a-42e1-cdd9-bf6393a14613" colab={"base_uri": "https://localhost:8080/"}
train50.shape
# + [markdown] id="WYl2Y9WAxV94"
# # Time to build the model
#
# So far we have:
#
# * a 20,000 word vocabulary
# * each review is limited to 50 words
#
# And we want to create a word embedding with 50 features.
#
# (Those 50 are completely separate parameters) The 50 word limit is not related to the 50 features.)
#
# Let's create a network with an embedding layer
#
# + id="jDdDVHVb6t3z" colab={"base_uri": "https://localhost:8080/"} outputId="b726325f-b6eb-461a-e3d4-bf344157e263"
from keras.models import Sequential
from keras.layers import Flatten, Dense, Embedding
vocabulary_size = 20000
embedding_size = 50
model = Sequential()
# We specify the maximum input length to our Embedding layer
# so we can later flatten the embedded inputs
model.add(Embedding(vocabulary_size, embedding_size, input_length=maxlen))
# After the Embedding layer,
# our activations have shape `(samples, maxlen, embedding_size)`.
# We flatten the 3D tensor of embeddings
# into a 2D tensor of shape `(samples, maxlen * embedding_size)`
model.add(Flatten())
# We add the classifier on top
model.add(Dense(1, activation='sigmoid'))
model.compile(optimizer='rmsprop', loss='binary_crossentropy', metrics=['acc'])
model.summary()
# + [markdown] id="GyAB-38my7Vg"
# ## Training
# + id="1mIT1Lbvy9so" colab={"base_uri": "https://localhost:8080/"} outputId="d7709aa1-ddb4-48b3-ab64-fb29387dcf0b"
history = model.fit(
train50, train_labels,
steps_per_epoch=100,
epochs=10,
validation_split=0.2,
validation_steps=50)
# + [markdown] id="3B1mpi9B6t31"
# We get to a validation accuracy of ~82%, which is pretty good considering that we only look at the first 50 words in every review. But
# note that merely flattening the embedded sequences and training a single `Dense` layer on top leads to a model that treats each word in the
# input sequence separately, without considering inter-word relationships and structure sentence (e.g. it would likely treat both _"this movie
# is shit"_ and _"this movie is the shit"_ as being negative "reviews"). It would be much better to add recurrent layers or 1D convolutional
# layers on top of the embedded sequences to learn features that take into account each sequence as a whole.
# + [markdown] id="mOrcukRO6t31"
# ## Using pre-trained word embeddings
#
#
# Sometimes, we have so little training data available that we could never use the data alone to learn an appropriate task-specific embedding
# of your vocabulary. What to do then?
#
# Instead of learning word embeddings jointly with the problem we want to solve, we could be loading embedding vectors from a pre-computed
# embedding space known to be highly structured and to exhibit useful properties -- that captures generic aspects of language structure. The
# rationale behind using pre-trained word embeddings in natural language processing is very much the same as for using pre-trained convnets
# in image classification: we don't have enough data available to learn truly powerful features on our own, but we expect the features that
# we need to be fairly generic, i.e. common visual features or semantic features. In this case it makes sense to reuse features learned on a
# different problem.
#
# Such word embeddings are generally computed using word occurrence statistics (observations about what words co-occur in sentences or
# documents), using a variety of techniques, some involving neural networks, others not. The idea of a dense, low-dimensional embedding space
# for words, computed in an unsupervised way, was initially explored by Bengio et al. in the early 2000s, but it only started really taking
# off in research and industry applications after the release of one of the most famous and successful word embedding scheme: the Word2Vec
# algorithm, developed by Mikolov at Google in 2013. Word2Vec dimensions capture specific semantic properties, e.g. gender.
#
# There are various pre-computed databases of word embeddings that can download and start using in a Keras `Embedding` layer. Word2Vec is one
# of them. Another popular one is called "GloVe", developed by Stanford researchers in 2014. It stands for "Global Vectors for Word
# Representation", and it is an embedding technique based on factorizing a matrix of word co-occurrence statistics. Its developers have made
# available pre-computed embeddings for millions of English tokens, obtained from Wikipedia data or from Common Crawl data.
#
# Finally, there is FastText, also developed by Mikolov at Facebook. While state of the art, it is also the most resource intensive scheme.
#
# Let's take a look at how you can get started using GloVe embeddings in a Keras model. The same method will of course be valid for Word2Vec
# embeddings or any other word embedding database that you can download.
# + [markdown] id="QpnanKp26t32"
# ## Summary so far,
#
# We...
#
# 1. downloaded the text data which was in the form of a csv file
# 2. loaded the file into pandas.
# 3. divided the csv into the text columns and the labels columns
# 4. tokenized the text into sequences of integers
# 5. limited each text to 50 words
# 5. divided the sequences and labels into training and test data
#
# So, for example, the first entry of our data was
#
# + id="uqTEl1ys6t33" colab={"base_uri": "https://localhost:8080/"} outputId="619cadae-22d3-47e6-be14-e9553c151ae1"
train50[0]
# + [markdown] id="g7GNizUp6t38"
# ### Download the GloVe word embeddings
#
#
# Head to https://nlp.stanford.edu/projects/glove/ (where you can learn more about the GloVe algorithm), and download the pre-computed
# embeddings from 2014 English Wikipedia. It's a 822MB zip file named `glove.6B.zip`, containing 100-dimensional embedding vectors for
# 400,000 words (or non-word tokens). The downloading will take a bit of time, which might be an understatement.
#
# For convenience, I have made available a zip of the exact file we will need at http://zacharski.org/files/courses/cs419/glove.6B.100d.zip This is only 134MB
#
#
# Next, un-zip it.
# + id="GN0y6D-PBOcA" colab={"base_uri": "https://localhost:8080/"} outputId="4c7feb9a-48e9-440e-8652-64eb936f463d"
# !wget http://zacharski.org/files/courses/cs419/glove.6B.100d.zip
# !unzip glove.6B.100d.zip
# + [markdown] id="k6hBJa5p6t39"
# ### Pre-process the embeddings
#
#
# Let's parse the un-zipped file (it's a `txt` file) to build an index mapping words (as strings) to their vector representation (as number
# vectors).
# + id="zboejuFj6t39" colab={"base_uri": "https://localhost:8080/"} outputId="dba571dd-6211-41aa-c4b8-f47506977e94"
import numpy as np
embeddings_index = {}
f = open('glove.6B.100d.txt')
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
f.close()
print('Found %s word vectors.' % len(embeddings_index))
# + [markdown] id="QBlb3xFu6t3_"
# So this embedding file has a vocabulary size of 400,000 words and each word is associated with a 100 element vector. Let's not use all 400,000, but restrict our vocabulary to our original 20,000 (`max_words`).
#
# Now let's build an embedding matrix that we will be able to load into an `Embedding` layer. It must be a matrix of shape `(max_words,
# embedding_dim)`, where each entry `i` contains the `embedding_dim`-dimensional vector for the word of index `i` in our reference word index
# (built during tokenization). Note that the index `0` is not supposed to stand for any word or token -- it's a placeholder.
# + id="j4djtC3A6t3_"
embedding_dim = 100
max_words = 20000
word_index = tokenizer.word_index
embedding_matrix = np.zeros((max_words, embedding_dim))
for word, i in word_index.items():
embedding_vector = embeddings_index.get(word)
if i < max_words:
if embedding_vector is not None:
# Words not found in embedding index will be all-zeros.
embedding_matrix[i] = embedding_vector
# + id="XSLbfiv7cYfB" outputId="752b20f2-830c-404c-e37e-88d517248a6f" colab={"base_uri": "https://localhost:8080/"}
embedding_matrix.shape
# + [markdown] id="0EEPKbcd6t4C"
# ### Define a model
#
# We will be using the same model architecture as before:
# + id="NmRvo6sb6t4C" colab={"base_uri": "https://localhost:8080/"} outputId="01921803-3585-48c6-cb31-6b4151403495"
from keras.models import Sequential
from keras.layers import Embedding, Flatten, Dense
model = Sequential()
model.add(Embedding(max_words, embedding_dim, input_length=maxlen))
model.add(Flatten())
model.add(Dense(32, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.summary()
# + [markdown] id="lSVZCgJy6t4F"
# ### Load the GloVe embeddings in the model
#
#
# The `Embedding` layer has a single weight matrix: a 2D float matrix where each entry `i` is the word vector meant to be associated with
# index `i`. Simple enough. Let's just load the GloVe matrix we prepared into our `Embedding` layer, the first layer in our model:
# + id="7GUR2IMr6t4G"
model.layers[0].set_weights([embedding_matrix])
model.layers[0].trainable = False
# + [markdown] id="yyiOZNs76t4J"
#
# Additionally, we freeze the embedding layer (we set its `trainable` attribute to `False`), following the same rationale as what you are
# already familiar with in the context of pre-trained convnet features: when parts of a model are pre-trained (like our `Embedding` layer),
# and parts are randomly initialized (like our classifier), the pre-trained parts should not be updated during training to avoid forgetting
# what they already know. The large gradient update triggered by the randomly initialized layers would be very disruptive to the already
# learned features.
# + [markdown] id="taMHdF-a6t4J"
# ### Train and evaluate
#
# Let's compile our model and train it:
# + id="T-mHYyp86t4J" colab={"base_uri": "https://localhost:8080/"} outputId="aa9f07d2-8ce6-4865-e41d-dd7dda21b1b4"
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['acc'])
history = model.fit(train50, train_labels,
epochs=10,
batch_size=32,
validation_split=0.2,
validation_steps=50)
model.save_weights('pre_trained_glove_model.h5')
# + [markdown] id="212yP1DI6t4M"
# Let's plot its performance over time:
# + id="w4Eince06t4M" colab={"base_uri": "https://localhost:8080/", "height": 545} outputId="92b8c307-4a22-4764-b6ea-0913af2dea4e"
import matplotlib.pyplot as plt
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# + [markdown] id="ZB79P4nm6t4R"
#
# The model quickly starts overfitting, unsurprisingly given the small number of training samples. Validation accuracy has high variance for
# the same reason, but seems to reach high 50s.
#
# ## Let's Reflect
#
# ### Experiment 1
# We trained the model without loading the pre-trained word embeddings and without freezing the embedding layer. In that
# case, we would be learning a task-specific embedding of our input tokens, which is generally more powerful than pre-trained word embeddings
# when lots of data is available.
#
# ### Experiment 2
# However, we used a pre-existing word-embedding. This is especially useful when we have a limited amount of training data.
#
# ### Experiment 3
# Let's try a new approach. Suppose we loaded the GloVe pre-trained word embeddings but this time didn't freeeze the layer, meaning it would continue to refine the embeddings using our training data. How does that do?
#
# + id="UxHX4E_A6t4S"
from keras.models import Sequential
from keras.layers import Embedding, Flatten, Dense
model = Sequential()
model.add(Embedding(max_words, embedding_dim, input_length=maxlen))
model.add(Flatten())
model.add(Dense(32, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.layers[0].set_weights([embedding_matrix])
# Without freezing layer
model.layers[0].trainable = True
# + id="gJGU2CPq6t4U" colab={"base_uri": "https://localhost:8080/"} outputId="cbc5a1c4-c0eb-4851-cca7-7ea4c7148352"
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['acc'])
history = model.fit(train50, train_labels,
epochs=10,
batch_size=32,
validation_split=0.2,
validation_steps=50)
# + id="vO2jZN9zWaiL" colab={"base_uri": "https://localhost:8080/", "height": 545} outputId="0a513b0a-1d78-4cdb-8fc4-cb97616edd8e"
import matplotlib.pyplot as plt
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# + [markdown] id="ZHB5WNGFXeEV"
# It did better than freezing the layer
# + [markdown] id="3xN-GWKg6t4W"
# # You try - Reuters Dataset
#
# This is the same dataset we used in the previous notebook. The dataset consists of text from the Reuters newswire in 1987. We will use a subset of the data that contain texts from eight topics:
#
#
#
# Class | # train docs | # test docs | Total # docs
# :--- | --: | --: | --:
# acq | 1596 | 696 | 2292
# crude | 253 | 121 | 374
# earn | 2840 | 1083 | 3923
# grain |41 | 10 | 51
# interest | 190 | 81 | 271
# money-fx | 206 | 87 | 293
# ship | 108 | 36 | 144
# trade | 251 | 75 | 326
# Total | 5485 | 2189 | 7674
#
# The training dataset is http://zacharski.org/files/courses/cs419/r8-train-all-terms.txt
#
# The test dataset is http://zacharski.org/files/courses/cs419/r8-test-all-terms.txt
#
# Can you build a network that will classify texts into one of 8 categories?
#
# Make predictions on the test set to see how well it performs.
#
# Which is better
#
# 1. Computing our own embeddings
# 2. Using a pre-existing embeddings
#
# I think using pre-existing embeddings will do better but we shall see. The accuracy for the previous lab was 60%. I'm hoping this method will yield a 75% accuracy.
# + id="JVl6o8_96t4W"
import pandas as pd
train = pd.read_csv('http://zacharski.org/files/courses/cs419/r8-train-all-terms.txt', sep='\t')
test = pd.read_csv('http://zacharski.org/files/courses/cs419/r8-test-all-terms.txt', sep='\t')
# + id="dFfbhwzbZ9-7"
train_text = train.drop('class', axis=1).to_numpy().reshape(-1)
train_labels = train['class'].to_numpy().reshape(-1, 1)
test_text = test.drop('class', axis=1).to_numpy().reshape(-1)
test_labels = test['class'].to_numpy().reshape(-1, 1)
# + id="FTno_turaSC-" colab={"base_uri": "https://localhost:8080/"} outputId="bc3631bb-24b5-45f9-8f21-10ed0ed0f4f6"
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
y_train = le.fit_transform(train_labels)
y_test = le.fit_transform(test_labels)
from keras.utils import to_categorical
y_train = to_categorical(y_train)
y_test = to_categorical(y_test)
# + id="iEEtuODlbr-l" colab={"base_uri": "https://localhost:8080/"} outputId="2d72b949-45df-44be-a89e-17cfd38980d2"
from keras.preprocessing.text import Tokenizer
tokenizer = Tokenizer(num_words = 5000)
#build the word index
tokenizer.fit_on_texts(train_text)
tokenizer.fit_on_texts(test_text)
# now convert the words to integers
train_data_sequences = tokenizer.texts_to_sequences(train_text)
test_data_sequences = tokenizer.texts_to_sequences(test_text)
train_data_sequences[0]
# + id="lZtovUiHejhn" colab={"base_uri": "https://localhost:8080/"} outputId="cb0b7f9e-ec4e-47e2-a67b-b97f37073440"
tokenizer.sequences_to_texts([train_data_sequences[0]])
# + id="VsdNulcMewbR" colab={"base_uri": "https://localhost:8080/"} outputId="9d52450c-d6e2-473e-f933-10ab8f4b2e1c"
# Cut texts after this number of words
# (among top max_features most common words)
maxlen = 50
from keras import preprocessing
# This turns our lists of integers
# into a 2D integer tensor of shape `(samples, maxlen)`
X_train = preprocessing.sequence.pad_sequences(train_data_sequences, maxlen=maxlen)
X_test = preprocessing.sequence.pad_sequences(test_data_sequences, maxlen=maxlen)
X_train[0]
# + id="Z37_9G36li5M" outputId="b5408dd7-077b-45bb-92d5-023e7c0fa461" colab={"base_uri": "https://localhost:8080/"}
X_test.shape
# + id="WzIVv7P8fga5" colab={"base_uri": "https://localhost:8080/"} outputId="2efc4065-0a9b-46e3-9077-c3a7beaae540"
from keras.models import Sequential
from keras.layers import Flatten, Dense, Embedding
vocabulary_size = 5000
embedding_size = 50
# maxLen = 50
model = Sequential()
# We specify the maximum input length to our Embedding layer
# so we can later flatten the embedded inputs
model.add(Embedding(vocabulary_size, embedding_size, input_length=maxlen))
# After the Embedding layer,
# our activations have shape `(samples, maxlen, embedding_size)`.
# We flatten the 3D tensor of embeddings
# into a 2D tensor of shape `(samples, maxlen * embedding_size)`
model.add(Flatten())
# We add the classifier on top
model.add(Dense(8, activation='softmax'))
model.compile(optimizer='rmsprop', loss='binary_crossentropy', metrics=['acc'])
model.summary()
# + id="Chfb_CLMjAwb" colab={"base_uri": "https://localhost:8080/"} outputId="5d5180d5-c1a3-4fe0-c6f7-4253749d6283"
history = model.fit(
X_train, y_train,
steps_per_epoch=100,
epochs=10,
validation_split=0.2,
validation_steps=50)
# + [markdown] id="5bAshbzJjS0v"
# We get a good accuracy and a decent validation accuracy of ~85%. It's still overfitting but good overrall.
# + id="eMWJ9PoxjSAg" colab={"base_uri": "https://localhost:8080/", "height": 545} outputId="3017e4eb-d361-4fcf-90ff-aa888645246d"
import matplotlib.pyplot as plt
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training accuracy')
plt.plot(epochs, val_acc, 'b', label='Validation accuracy')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# + id="DJAJVWBRkLjy" colab={"base_uri": "https://localhost:8080/"} outputId="80c23c61-2239-4d3c-d49a-b50565ed7367"
scoreSeg = model.evaluate(X_test, y_test)
print("Accuracy: ", scoreSeg[1])
# + [markdown] id="Z14Y1N4LkZJ0"
# ##We get an accuracy of 88.57% for our own embeddings. Now, let's see how it performs with pre-existing embeddings.
# + id="MC0uNXaHmcqW"
embedding_dim = 100
max_words = 20000
word_index = tokenizer.word_index
embedding_matrix = np.zeros((max_words, embedding_dim))
for word, i in word_index.items():
embedding_vector = embeddings_index.get(word)
if i < max_words:
if embedding_vector is not None:
# Words not found in embedding index will be all-zeros.
embedding_matrix[i] = embedding_vector
# + id="aEfXQAO6nogm" outputId="dbc01fd6-2af7-428a-9a9f-f383f4326f56" colab={"base_uri": "https://localhost:8080/"}
embedding_matrix.shape
# + id="Up9hYk9MktP7" outputId="a01de817-0c0d-41cc-d33b-3bd355226e98" colab={"base_uri": "https://localhost:8080/"}
from keras.models import Sequential
from keras.layers import Flatten, Dense, Embedding
vocabulary_size = 20000
model = Sequential()
model.add(Embedding(vocabulary_size, embedding_dim, input_length=maxlen))
model.add(Flatten())
model.add(Dense(32, activation='relu'))
model.add(Dense(8, activation='softmax'))
model.layers[0].set_weights([embedding_matrix])
model.layers[0].trainable = False
model.compile(optimizer='rmsprop', loss='binary_crossentropy', metrics=['acc'])
model.summary()
# + id="RDzHp-bpl8WF" colab={"base_uri": "https://localhost:8080/"} outputId="f44848b3-bea0-4163-eb81-1b694e4b20ba"
history = model.fit(X_train, y_train,
epochs=10,
#steps_per_epoch=171,
#batch_size=32,
validation_split=0.2,
validation_steps=50)
# + [markdown] id="MpNYU7F_n6j5"
# So far, worse accuracy than the our own embeddings.
# + id="Povl5YJ2qYK7" outputId="2cca7ee9-c885-4235-f4a9-56e3477e7d0a" colab={"base_uri": "https://localhost:8080/"}
scoreSeg = model.evaluate(X_test, y_test)
print("Accuracy: ", scoreSeg[1])
# + [markdown] id="XReQFK8zqb4a"
# ## We get an accuracy of 85.15% which is pretty close to our own embeddings. Now, let's try unfreezing it
# + id="uKmFvOikq2mr" outputId="0408696e-a91e-4480-bac9-9dbe282d9009" colab={"base_uri": "https://localhost:8080/"}
from keras.models import Sequential
from keras.layers import Flatten, Dense, Embedding
vocabulary_size = 20000
model = Sequential()
model.add(Embedding(vocabulary_size, embedding_dim, input_length=maxlen))
model.add(Flatten())
model.add(Dense(32, activation='relu'))
model.add(Dense(8, activation='softmax'))
model.layers[0].set_weights([embedding_matrix])
model.layers[0].trainable = True
model.compile(optimizer='rmsprop', loss='binary_crossentropy', metrics=['acc'])
history = model.fit(X_train, y_train,
epochs=10,
#steps_per_epoch=171,
#batch_size=32,
validation_split=0.2,
validation_steps=50)
# + id="59pace_GrtOm" outputId="9e0ed863-4867-4ef6-cd5e-884ab7314952" colab={"base_uri": "https://localhost:8080/"}
scoreSeg = model.evaluate(X_test, y_test)
print("Accuracy: ", scoreSeg[1])
# + [markdown] id="sxCjhg7Grxqm"
# ## Unfreezing the pre-existing word embeddings did better than them being frozen. It reached a 87.30% accuracy, which is almost the same as our own embeddings.
# <br><br><br>
# + [markdown] id="Jzi9moNEswnt"
# ## Doing experiments with less words with pre-existing embeddings
# + id="X3seBOlmsU43" outputId="6cb644a7-414b-4ef2-ad47-763b1f2b7e8d" colab={"base_uri": "https://localhost:8080/"}
from keras.models import Sequential
from keras.layers import Flatten, Dense, Embedding
embedding_dim = 100
max_words = 5000
word_index = tokenizer.word_index
embedding_matrix = np.zeros((max_words, embedding_dim))
for word, i in word_index.items():
embedding_vector = embeddings_index.get(word)
if i < max_words:
if embedding_vector is not None:
# Words not found in embedding index will be all-zeros.
embedding_matrix[i] = embedding_vector
vocabulary_size = 5000
model = Sequential()
model.add(Embedding(vocabulary_size, embedding_dim, input_length=maxlen))
model.add(Flatten())
model.add(Dense(32, activation='relu'))
model.add(Dense(8, activation='softmax'))
model.layers[0].set_weights([embedding_matrix])
model.layers[0].trainable = False
model.compile(optimizer='rmsprop', loss='binary_crossentropy', metrics=['acc'])
history = model.fit(X_train, y_train,
epochs=10,
#steps_per_epoch=171,
#batch_size=32,
validation_split=0.2,
validation_steps=50)
# + id="Bocd20cbstSu" outputId="46d1ec19-5db1-4320-84ef-71803638ea97" colab={"base_uri": "https://localhost:8080/"}
scoreSeg = model.evaluate(X_test, y_test)
print("Accuracy: ", scoreSeg[1])
# + id="D9PHc34ls5Nt" outputId="de9f8158-7abc-4f6c-f759-f70291df06bb" colab={"base_uri": "https://localhost:8080/"}
from keras.models import Sequential
from keras.layers import Flatten, Dense, Embedding
embedding_dim = 100
max_words = 5000
word_index = tokenizer.word_index
embedding_matrix = np.zeros((max_words, embedding_dim))
for word, i in word_index.items():
embedding_vector = embeddings_index.get(word)
if i < max_words:
if embedding_vector is not None:
# Words not found in embedding index will be all-zeros.
embedding_matrix[i] = embedding_vector
vocabulary_size = 5000
model = Sequential()
model.add(Embedding(vocabulary_size, embedding_dim, input_length=maxlen))
model.add(Flatten())
model.add(Dense(32, activation='relu'))
model.add(Dense(8, activation='softmax'))
model.layers[0].set_weights([embedding_matrix])
model.layers[0].trainable = True
model.compile(optimizer='rmsprop', loss='binary_crossentropy', metrics=['acc'])
history = model.fit(X_train, y_train,
epochs=10,
#steps_per_epoch=171,
#batch_size=32,
validation_split=0.2,
validation_steps=50)
# + id="bnLPia8NtDjR" outputId="2529e05d-b8e5-4982-a0ba-070181fd6048" colab={"base_uri": "https://localhost:8080/"}
scoreSeg = model.evaluate(X_test, y_test)
print("Accuracy: ", scoreSeg[1])
# + [markdown] id="bnZRHQoUtHXP"
# ## So, it seems like the amount of words doesn't do much to the model
# <br><br><br>
# + [markdown] id="GEkeaxnyi2K0"
# #### MIT License
# Copyright (c) 2017 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
|
labs/word_embeddings.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img align="right" src="images/tf.png" width="128"/>
# <img align="right" src="images/logo.png" width="128"/>
# <img align="right" src="images/etcbc.png" width="128"/>
# <img align="right" src="images/dans.png" width="128"/>
#
# ---
#
# To get started: consult [start](start.ipynb)
#
# ---
#
# # Sharing data features
#
# ## Explore additional data
#
# Once you analyse a corpus, it is likely that you produce data that others can reuse.
# Maybe you have defined a set of proper name occurrences, or special numerals, or you have computed part-of-speech assignments.
#
# It is possible to turn these insights into *new features*, i.e. new `.tf` files with values assigned to specific nodes.
#
# ## Make your own data
#
# New data is a product of your own methods and computations in the first place.
# But how do you turn that data into new TF features?
# It turns out that the last step is not that difficult.
#
# If you can shape your data as a mapping (dictionary) from node numbers (integers) to values
# (strings or integers), then TF can turn that data into a feature file for you with one command.
#
# ## Share your new data
# You can then easily share your new features on GitHub, so that your colleagues everywhere
# can try it out for themselves.
#
# You can add such data on the fly, by passing a `mod={org}/{repo}/{path}` parameter,
# or a bunch of them separated by commas.
#
# If the data is there, it will be auto-downloaded and stored on your machine.
#
# Let's do it.
# %load_ext autoreload
# %autoreload 2
import os
from tf.app import use
# A = use('dss', hoist=globals())
A = use("dss:clone", checkout="clone", hoist=globals())
# # Making data
#
# We illustrate the data creation part by creating a new feature, `cert`.
# The idea is that we mark every consonant sign voor certainty.
#
# A certain consonant gets `cert=100`.
#
# If the consonant has the uncertain feature `unc`, then 10 times its value is subtracted from 100.
#
# If the consonant has the feature `rec`, it loses 45 points.
#
# Ancient removal `rem2` leads to minus 20, modern removal `rem` to minus 40.
#
# Ancient correction `cor2` leads to minus 12, modern correction `cor` to minus 18.
#
# Alternate marking `alt` leads to minus 25.
#
# The minimum is `1`.
#
# We extend the `cert` measure to words, fragments and scrolls by averaging over signs that
# have received a `cert` measure.
def measure(s):
c = 100
d = F.unc.v(s)
if d:
c -= 10 * d
d = F.rec.v(s)
if d:
c -= 45
d = F.rem.v(s)
if d == 1:
c -= 40
elif d == 2:
c -= 20
d = F.cor.v(s)
if d == 2 or d == 3:
c -= 12
elif d == 1:
c -= 18
d = F.alt.v(s)
if d:
c -= 25
if c < 1:
c = 1
return c
# +
CONS = "cons"
cert = {}
A.indent(reset=True)
for sc in F.otype.s("scroll"):
fN = 0
fSum = 0
for f in L.d(sc, otype="fragment"):
lN = 0
lSum = 0
for ln in L.d(f, otype="line"):
wN = 0
wSum = 0
for w in L.d(ln, otype="word"):
sN = 0
sSum = 0
for s in L.d(w, otype="sign"):
if F.type.v(s) != CONS:
continue
sCert = measure(s)
cert[s] = sCert
sN += 1
sSum += sCert
if sN:
wCert = int(round(sSum / sN))
cert[w] = wCert
wN += 1
wSum += wCert
if wN:
lCert = int(round(wSum / wN))
cert[ln] = lCert
lN += 1
lSum += lCert
if lN:
fCert = int(round(lSum / lN))
cert[f] = fCert
fN += 1
fSum += fCert
if fN:
scCert = int(round(fSum / fN))
cert[sc] = scCert
A.info(f"{len(cert)} certainties determined")
# -
# # Saving data
#
# The [documentation](https://annotation.github.io/text-fabric/tf/core/fabric.html#tf.core.fabric.FabricCore.save) explains how to save this data into a text-fabric
# data file.
#
# We choose a location where to save it, the `exercises` folder in the `dss` repository in the `dss` organization.
#
# In order to do this, we restart the TF api, but now with the desired output location in the `locations` parameter.
GITHUB = os.path.expanduser("~/github")
ORG = "etcbc"
REPO = "dss"
PATH = "exercises"
VERSION = A.version
# Note the version: we have built the version against a specific version of the data:
A.version
# Later on, we pass this version on, so that users of our data will get the shared data in exactly the same version as their core data.
# We have to specify a bit of metadata for this feature:
metaData = {
"cert": dict(
valueType="int",
description="measure of certainty of material, between 1 and 100 (most certain)",
creator="<NAME>",
),
}
# Now we can give the save command:
TF.save(
nodeFeatures=dict(cert=cert),
metaData=metaData,
location=f"{GITHUB}/{ORG}/{REPO}/{PATH}/tf",
module=VERSION,
)
# # Sharing data
#
# How to share your own data is explained in the
# [documentation](https://annotation.github.io/text-fabric/tf/about/datasharing.html).
#
# Here we show it step by step for the `cert` feature.
#
# If you commit your changes to the exercises repo, and have done a `git push origin master`,
# you already have shared your data!
#
# If you want to make a stable release, so that you can keep developing, while your users fall back
# on the stable data, you can make a new release.
#
# Go to the GitHub website for that, go to your repo, and click *Releases* and follow the nudges.
#
# If you want to make it even smoother for your users, you can zip the data and attach it as a binary to the release just created.
#
# We need to zip the data in exactly the right directory structure. Text-Fabric can do that for us:
# + language="sh"
#
# text-fabric-zip etcbc/dss/exercises/tf
# -
# All versions have been zipped, but it works OK if you only attach the newest version to the newest release.
#
# If a user asks for an older version in this release, the system can still find it.
# # Use the data
#
# We can use the data by calling it up when we say `use('dss', ...)`.
#
# Here is how:
#
# (use the line without `clone` if the data is really published,
# use the line with `clone` if you want to test your local copy of the feature).
# A = use('dss', hoist=globals(), mod='etcbc/dss/exercises/tf')
A = use(
"dss:clone", checkout="clone", hoist=globals(), mod="etcbc/dss/exercises/tf:clone"
)
# Above you see a new section in the feature list: **etcbc/dss/exercises/tf** with our foreign feature in it: `cert`.
#
# Now, suppose did not know much about this feature, then we would like to do a few basic checks:
F.cert.freqList()
# Which nodes have the lowest uncertainty?
{F.otype.v(n) for n in N.walk() if F.cert.v(n) and F.cert.v(n) < 10}
# Only signs are this uncertain.
#
# Let's look for pretty uncertain fragments:
results = A.search(
"""
fragment cert<50
"""
)
results = A.search(
"""
fragment cert<60
"""
)
A.table(results, start=1, end=20)
# Same for scrolls:
results = A.search(
"""
scroll cert<50
"""
)
results = A.search(
"""
scroll cert<60
"""
)
A.show(results)
# Lines with certainty of 50:
results = A.search(
"""
line cert<57
"""
)
A.show(results, start=100, end=102)
# With highlights and drilled down to sign level:
# +
highlights = {}
for s in F.otype.s("sign"):
if not F.cert.v(s):
continue
color = "lightsalmon" if F.cert.v(s) < 56 else "mediumaquamarine"
highlights[s] = color
# -
A.show(
results,
start=100,
end=102,
withNodes=True,
condensed=True,
highlights=highlights,
baseTypes="sign",
)
# # All together!
#
# If more researchers have shared data modules, you can draw them all in.
#
# Then you can design queries that use features from all these different sources.
#
# In that way, you build your own research on top of the work of others.
# Hover over the features to see where they come from, and you'll see they come from your local github repo.
# ---
#
# All chapters:
#
# * **[start](start.ipynb)** become an expert in creating pretty displays of your text structures
# * **[display](display.ipynb)** become an expert in creating pretty displays of your text structures
# * **[search](search.ipynb)** turbo charge your hand-coding with search templates
# * **[exportExcel](exportExcel.ipynb)** make tailor-made spreadsheets out of your results
# * **share** draw in other people's data and let them use yours
# * **[similarLines](similarLines.ipynb)** spot the similarities between lines
#
# ---
#
# See the [cookbook](cookbook) for recipes for small, concrete tasks.
#
# CC-BY <NAME>
|
dss/share.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="Ic4_occAAiAT"
# ##### Copyright 2018 The TensorFlow Authors.
# + cellView="form" colab_type="code" id="ioaprt5q5US7" colab={}
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + cellView="form" colab_type="code" id="yCl0eTNH5RS3" colab={}
#@title MIT License
#
# Copyright (c) 2017 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a
# # copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
# + [markdown] colab_type="text" id="ItXfxkxvosLH"
# # Classificação de texto com avaliações de filmes
# + [markdown] colab_type="text" id="hKY4XMc9o8iB"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/keras/basic_text_classification"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />Veja em TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/pt/tutorials/keras/basic_text_classification.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Execute em Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/pt/tutorials/keras/basic_text_classification.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />Veja a fonte em GitHub</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="Eg62Pmz3o83v"
#
# Este *notebook* classifica avaliações de filmes como **positiva** ou **negativa** usando o texto da avaliação. Isto é um exemplo de classificação *binária* —ou duas-classes—, um importante e bastante aplicado tipo de problema de aprendizado de máquina.
#
# Usaremos a base de dados [IMDB](https://www.tensorflow.org/api_docs/python/tf/keras/datasets/imdb) que contém avaliaçòes de mais de 50000 filmes do bando de dados [Internet Movie Database](https://www.imdb.com/). A base é dividida em 25000 avaliações para treinamento e 25000 para teste. Os conjuntos de treinamentos e testes são *balanceados*, ou seja, eles possuem a mesma quantidade de avaliações positivas e negativas.
#
# O notebook utiliza [tf.keras](https://www.tensorflow.org/guide/keras), uma API alto-nível para construir e treinar modelos com TensorFlow. Para mais tutoriais avançados de classificação de textos usando `tf.keras`, veja em [MLCC Text Classification Guide](https://developers.google.com/machine-learning/guides/text-classification/).
# + colab_type="code" id="JL-LtD0CBSZR" colab={}
# keras.datasets.imdb está quebrado em 1.13 e 1.14, pelo np 1.16.3
# !pip install tf_nightly
# + colab_type="code" id="2ew7HTbPpCJH" colab={}
from __future__ import absolute_import, division, print_function, unicode_literals
import tensorflow as tf
from tensorflow import keras
import numpy as np
print(tf.__version__)
# + [markdown] colab_type="text" id="iAsKG535pHep"
# ## Baixe a base de dados IMDB
#
# A base de dados vem empacotada com TensorFlow. Ele já vem pré-processado de forma que as avaliações (sequências de palavras) foi convertida em sequências de inteiros, onde cada inteiro representa uma palavra específica no dicionário.
#
# O código abaixo baixa a base de dados IMDB para a sua máquina (ou usa a cópia em *cache*, caso já tenha baixado):
# + colab_type="code" id="zXXx5Oc3pOmN" colab={}
imdb = keras.datasets.imdb
(train_data, train_labels), (test_data, test_labels) = imdb.load_data(num_words=10000)
# + [markdown] colab_type="text" id="odr-KlzO-lkL"
# O argumento `num_words=10000` mantém as 10000 palavras mais frequentes no conjunto de treinamento. As palavras mais raras são descartadas para preservar o tamanho dos dados de forma maleável.
# + [markdown] colab_type="text" id="l50X3GfjpU4r"
# ## Explore os dados
#
# Vamos parar um momento para entender o formato dos dados. O conjunto de dados vem pré-processado: cada exemplo é um *array* de inteiros representando as palavras da avaliação do filme. Cada *label* é um inteiro com valor ou de 0 ou 1, onde 0 é uma avaliação negativa e 1 é uma avaliação positiva.
# + colab_type="code" id="y8qCnve_-lkO" colab={}
print("Training entries: {}, labels: {}".format(len(train_data), len(train_labels)))
# + [markdown] colab_type="text" id="RnKvHWW4-lkW"
# O texto das avaliações foi convertido para inteiros, onde cada inteiro representa uma palavra específica no dicionário. Isso é como se parece a primeira revisão:
# + colab_type="code" id="QtTS4kpEpjbi" colab={}
print(train_data[0])
# + [markdown] colab_type="text" id="hIE4l_72x7DP"
# As avaliações dos filmes têm diferentes tamanhos. O código abaixo mostra o número de palavras da primeira e segunda avaliação. Sabendo que o número de entradas da rede neural tem que ser de mesmo também, temos que resolver isto mais tarde.
# + colab_type="code" id="X-6Ii9Pfx6Nr" colab={}
len(train_data[0]), len(train_data[1])
# + [markdown] colab_type="text" id="4wJg2FiYpuoX"
# ### Converta os inteiros de volta a palavras
#
# É util saber como converter inteiros de volta a texto. Aqui, criaremos uma função de ajuda para consultar um objeto *dictionary* que contenha inteiros mapeados em strings:
# + colab_type="code" id="tr5s_1alpzop" colab={}
# Um dicionário mapeando palavras em índices inteiros
word_index = imdb.get_word_index()
# Os primeiros índices são reservados
word_index = {k:(v+3) for k,v in word_index.items()}
word_index["<PAD>"] = 0
word_index["<START>"] = 1
word_index["<UNK>"] = 2 # unknown
word_index["<UNUSED>"] = 3
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
def decode_review(text):
return ' '.join([reverse_word_index.get(i, '?') for i in text])
# + [markdown] colab_type="text" id="U3CNRvEZVppl"
# Agora, podemos usar a função `decode_review` para mostrar o texto da primeira avaliação:
# + colab_type="code" id="s_OqxmH6-lkn" colab={}
decode_review(train_data[0])
# + [markdown] colab_type="text" id="lFP_XKVRp4_S"
# ## Prepare os dados
#
# As avaliações—o *arrays* de inteiros— deve ser convertida em tensores (*tensors*) antes de alimentar a rede neural. Essa conversão pode ser feita de duas formas:
#
# * Converter os arrays em vetores de 0s e 1s indicando a ocorrência da palavra, similar com *one-hot encoding*. Por exemplo, a sequência [3, 5] se tornaria um vetor de 10000 dimensões, onde todos seriam 0s, tirando 3 would become a 10,000-dimensional vector that is all zeros except for indices 3 and 5, which are ones. Then, make this the first layer in our network—a Dense layer—that can handle floating point vector data. This approach is memory intensive, though, requiring a `num_words * num_reviews` size matrix.
#
# * Alternatively, we can pad the arrays so they all have the same length, then create an integer tensor of shape `max_length * num_reviews`. We can use an embedding layer capable of handling this shape as the first layer in our network.
#
# In this tutorial, we will use the second approach.
#
# Since the movie reviews must be the same length, we will use the [pad_sequences](https://keras.io/preprocessing/sequence/#pad_sequences) function to standardize the lengths:
# + colab_type="code" id="2jQv-omsHurp" colab={}
train_data = keras.preprocessing.sequence.pad_sequences(train_data,
value=word_index["<PAD>"],
padding='post',
maxlen=256)
test_data = keras.preprocessing.sequence.pad_sequences(test_data,
value=word_index["<PAD>"],
padding='post',
maxlen=256)
# + [markdown] colab_type="text" id="VO5MBpyQdipD"
# Let's look at the length of the examples now:
# + colab_type="code" id="USSSBnkE-lky" colab={}
len(train_data[0]), len(train_data[1])
# + [markdown] colab_type="text" id="QJoxZGyfjT5V"
# And inspect the (now padded) first review:
# + colab_type="code" id="TG8X9cqi-lk9" colab={}
print(train_data[0])
# + [markdown] colab_type="text" id="LLC02j2g-llC"
# ## Construindo o modelo
#
# A rede neural é criada por camadas empilhadas —isso necessita duas decisões arquiteturais principais:
#
# * Quantas camadas serão usadas no modelo?
# * Quantas *hidden units* são usadas em cada camada?
#
# Neste exemplo, os dados de entrada são um *array* de palavras-índices. As *labels* para predizer são ou 0 ou 1. Vamos construir um modelo para este problema:
# + colab_type="code" id="xpKOoWgu-llD" colab={}
# O formato de entrada é a contagem vocabulário usados pelas avaliações dos filmes (10000 palavras)
vocab_size = 10000
model = keras.Sequential()
model.add(keras.layers.Embedding(vocab_size, 16))
model.add(keras.layers.GlobalAveragePooling1D())
model.add(keras.layers.Dense(16, activation=tf.nn.relu))
model.add(keras.layers.Dense(1, activation=tf.nn.sigmoid))
model.summary()
# + [markdown] colab_type="text" id="6PbKQ6mucuKL"
# As camadas são empilhadas sequencialmente para construir o classificador:
#
# 1. A primeira camada é uma camada `Embedding` layer (*`Embedding` layer*). Essa camada pega o vocabulário em inteiros e olha o vetor *embedding* em cada palavra-index. Esses vetores são aprendidos pelo modelo, ao longo do treinamento. Os vetores adicionam a dimensão ao *array* de saída. As dimensões resultantes são: `(batch, sequence, embedding)`.
# 2. Depois, uma camada `GlobalAveragePooling1D` retorna um vetor de saída com comprimento fixo para cada exemplo fazendo a média da sequência da dimensão. Isso permite o modelo de lidar com entradas de tamanhos diferentes da maneira mais simples possível.
# 3. Esse vetor de saída com tamanho fixo passa por uma camada *fully-connected* (`Dense`) layer com 16 *hidden units*.
# 4. A última camada é uma *densely connected* com um único nó de saída. Usando uma função de ativação `sigmoid`, esse valor é um float que varia entre 0 e 1, representando a probabilidade, ou nível de confiança.
# + [markdown] colab_type="text" id="0XMwnDOp-llH"
# ### Hidden units
#
# O modelo abaixo tem duas camadas intermediárias ou _"hidden"_ (hidden layers), entre a entrada e saída. O número de saídas (unidades— *units*—, nós ou neurônios) é a dimensão do espaço representacional para a camada. Em outras palavras, a quantidade de liberdade que a rede é permitida enquanto aprende uma representação interna.
#
# Se o modelo tem mais *hidden units* (um espaço representacional de maior dimensão), e/ou mais camadas, então a rede pode aprender representações mais complexas. Entretanto, isso faz com que a rede seja computacionamente mais custosa e pode levar o aprendizado de padrões não desejados— padrões que melhoram a performance com os dados de treinamento, mas não com os de teste. Isso se chama *overfitting*, e exploraremos mais tarde.
# + [markdown] colab_type="text" id="L4EqVWg4-llM"
# ### Função Loss e otimizadores (optimizer)
#
# O modelo precisa de uma função *loss* e um otimizador (*optimizer*) para treinamento. Já que é um problema de classificação binário e o modelo tem com saída uma probabilidade (uma única camada com ativação sigmoide), usaremos a função loss `binary_crossentropy`.
#
# Essa não é a única escolha de função loss, você poderia escolher, no lugar, a `mean_squared_error`. Mas, geralmente, `binary_crossentropy` é melhor para tratar probabilidades— ela mede a "distância" entre as distribuições de probabilidade, ou, no nosso caso, sobre a distribuição real e as previsões.
#
# Mais tarde, quando explorarmos problemas de regressão (como, predizer preço de uma casa), veremos como usar outra função loss chamada *mean squared error*.
#
# Agora, configure o modelo para usar o *optimizer* a função loss:
# + colab_type="code" id="Mr0GP-cQ-llN" colab={}
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['acc'])
# + [markdown] colab_type="text" id="hCWYwkug-llQ"
# ### Crie um conjunto de validação
#
# Quando treinando. queremos checar a acurácia do modelo com os dados que ele nunca viu. Crie uma conjunto de *validação* tirando 10000 exemplos do conjunto de treinamento original. (Por que não usar o de teste agora? Nosso objetivo é desenvolver e melhorar (tunar) nosso modelo usando somente os dados de treinamento, depois usar o de teste uma única vez para avaliar a acurácia).
# + colab_type="code" id="-NpcXY9--llS" colab={}
x_val = train_data[:10000]
partial_x_train = train_data[10000:]
y_val = train_labels[:10000]
partial_y_train = train_labels[10000:]
# + [markdown] colab_type="text" id="35jv_fzP-llU"
# ## Treine o modelo
#
# Treine o modelo em 40 *epochs* com *mini-batches* de 512 exemplos. Essas 40 iterações sobre todos os exemplos nos tensores `x_train` e `y_train`. Enquanto treina, monitore os valores do loss e da acurácia do modelo nos 10000 exemplos do conjunto de validação:
# + colab_type="code" id="tXSGrjWZ-llW" colab={}
history = model.fit(partial_x_train,
partial_y_train,
epochs=40,
batch_size=512,
validation_data=(x_val, y_val),
verbose=1)
# + [markdown] colab_type="text" id="9EEGuDVuzb5r"
# ## Avalie o modelo
#
# E vamos ver como o modelo se saiu. Dois valores serão retornados. Loss (um número que representa o nosso erro, valores mais baixos são melhores), e acurácia.
# + colab_type="code" id="zOMKywn4zReN" colab={}
results = model.evaluate(test_data, test_labels)
print(results)
# + [markdown] colab_type="text" id="z1iEXVTR0Z2t"
# Está é uma aproximação ingênua que conseguiu uma acurácia de 87%. Com mais abordagens avançadas, o modelo deve chegar em 95%.
# + [markdown] colab_type="text" id="5KggXVeL-llZ"
# ## Crie um gráfico de acurácia e loss por tempo
#
# `model.fit()` retorna um objeto `History` que contém um dicionário de tudo o que aconteceu durante o treinamento:
# + colab_type="code" id="VcvSXvhp-llb" colab={}
history_dict = history.history
history_dict.keys()
# + [markdown] colab_type="text" id="nRKsqL40-lle"
# Tem 4 entradas: uma para cada métrica monitorada durante a validação e treinamento. Podemos usá-las para plotar a comparação do loss de treinamento e validação, assim como a acurácia de treinamento e validação:
# + colab_type="code" id="nGoYf2Js-lle" colab={}
import matplotlib.pyplot as plt
acc = history_dict['acc']
val_acc = history_dict['val_acc']
loss = history_dict['loss']
val_loss = history_dict['val_loss']
epochs = range(1, len(acc) + 1)
# "bo" de "blue dot" ou "ponto azul"
plt.plot(epochs, loss, 'bo', label='Training loss')
# b de "solid blue line" "linha azul"
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# + colab_type="code" id="6hXx-xOv-llh" colab={}
plt.clf() # limpa a figura
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="oFEmZ5zq-llk"
# No gráfico, os pontos representam o loss e acurácia de treinamento, e as linhas são o loss e a acurácia de validação.
#
# Note que o loss de treinamento *diminui* a cada *epoch* e a acurácia *aumenta*. Isso é esperado quando usado um gradient descent optimization—ele deve minimizar a quantidade desejada a cada iteração.
#
# Esse não é o caso do loss e da acurácia de validação— eles parecem ter um pico depois de 20 epochs. Isso é um exemplo de *overfitting*: o modelo desempenha melhor nos dados de treinamento do que quando usado com dados nunca vistos. Depois desse ponto, o modelo otimiza além da conta e aprende uma representação *especifica* para os dados de treinamento e não *generaliza* para os dados de teste.
#
# Para esse caso particular, podemos prevenir o *overfitting* simplesmente parando o treinamento após mais ou menos 20 epochs. Depois, você verá como fazer isso automaticamente com um *callback*.
|
site/pt/tutorials/keras/basic_text_classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
data_src='/Users/josephthomas/Documents/Projects/Data Projects/covid_death_trends/data/'
df_2018=pd.read_csv(data_src+'raw/2018_raw.txt',delimiter="\t")
df_2019=pd.read_csv(data_src+'raw/2019_raw.txt',delimiter="\t")
df_2020=pd.read_csv(data_src+'raw/2020_raw.txt',delimiter="\t")
df_2021=pd.read_csv(data_src+'raw/2021_raw.txt',delimiter="\t")
df_series=pd.read_csv(data_src+'raw/Total_deaths_time.txt',delimiter="\t")
df=pd.read_csv(data_src+'raw/Total_state.txt',delimiter="\t")
df.head()
grouped=df[["Year","Month","Deaths"]].groupby(["Year","Month"]).sum().reset_index()
grouped["Month"]=grouped["Month"].apply(lambda x:x.split(".")[0])
grouped["Month"]=grouped["Month"].apply(lambda x:x.split(",")[0])
grouped["Year"]=grouped["Year"].apply(lambda x:x.split(" ")[0])
grouped["Year"]=pd.to_datetime(grouped.Year,format='%Y', errors='coerce').dt.year
grouped["Month"]=pd.to_datetime(grouped.Month,format='%b', errors='coerce').dt.month
df1=grouped[grouped["Year"]==2018].sort_values(by="Month")
df2=grouped[grouped["Year"]==2019].sort_values(by="Month")
df3=grouped[grouped["Year"]==2020].sort_values(by="Month")
df4=grouped[grouped["Year"]==2021].sort_values(by="Month")
plt.plot("Month","Deaths",data=df1,label="2018")
plt.plot("Month","Deaths",data=df2,label="2019")
plt.plot("Month","Deaths",data=df3,label="2020")
plt.plot("Month","Deaths",data=df4,label="2021")
plt.legend()
|
notebooks/0.1-josepht47-exploration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # flower REST API
#
# This document shows how to use the flower [REST API](https://github.com/mher/flower#api).
#
# We will use [requests](http://www.python-requests.org/en/latest/) for accessing the API. (See [here](http://www.python-requests.org/en/latest/user/install/) on how to install it.)
# # Code
# We'll use the following code throughout the documentation.
# ## tasks.py
# +
from celery import Celery
from time import sleep
celery = Celery()
celery.config_from_object({
'BROKER_URL': 'amqp://localhost',
'CELERY_RESULT_BACKEND': 'amqp://',
'CELERYD_POOL_RESTARTS': True, # Required for /worker/pool/restart API
})
@celery.task
def add(x, y):
return x + y
@celery.task
def sub(x, y):
sleep(30) # Simulate work
return x - y
# -
# ## Running
# You'll need a celery worker instance and a flower instance running. In one terminal window run
#
# celery worker --loglevel INFO -A proj -E --autoscale 10,3
#
# and in another terminal run
#
# celery flower -A proj
# # Tasks API
# The tasks API is *async*, meaning calls will return immediatly and you'll need to poll on task status.
# # Done once for the whole docs
# import requests, json
# api_root = 'http://localhost:5555/api'
# task_api = '{}/task'.format(api_root)
# ## async-apply
args = {'args': [1, 2]}
url = '{}/async-apply/tasks.add'.format(task_api)
print(url)
resp = requests.post(url, data=json.dumps(args))
reply = resp.json()
reply
# We can see that we created a new task and it's pending. Note that the API is *async*, meaning it won't wait until the task finish.
# ## apply
# For create task and wait results you can use 'apply' API.
args = {'args': [1, 2]}
url = '{}/apply/tasks.add'.format(task_api)
print(url)
resp = requests.post(url, data=json.dumps(args))
reply = resp.json()
reply
# ## result
# Gets the task result. This is *async* and will return immediatly even if the task didn't finish (with state 'PENDING')
url = '{}/result/{}'.format(task_api, reply['task-id'])
print(url)
resp = requests.get(url)
resp.json()
# ## revoke
# Revoke a running task.
# +
# Run a task
args = {'args': [1, 2]}
resp = requests.post('{}/async-apply/tasks.sub'.format(task_api), data=json.dumps(args))
reply = resp.json()
# Now revoke it
url = '{}/revoke/{}'.format(task_api, reply['task-id'])
print(url)
resp = requests.post(url, data='terminate=True')
resp.json()
# -
# ## rate-limit
# Update [rate limit](http://docs.celeryproject.org/en/latest/userguide/tasks.html#Task.rate_limit) for a task.
worker = 'miki-manjaro' # You'll need to get the worker name from the worker API (seel below)
url = '{}/rate-limit/{}'.format(task_api, worker)
print(url)
resp = requests.post(url, params={'taskname': 'tasks.add', 'ratelimit': '10'})
resp.json()
# ## timeout
# Set timeout (both [hard](http://docs.celeryproject.org/en/latest/userguide/tasks.html#Task.time_limit) and [soft](http://docs.celeryproject.org/en/latest/userguide/tasks.html#Task.soft_time_limit)) for a task.
url = '{}/timeout/{}'.format(task_api, worker)
print(url)
resp = requests.post(url, params={'taskname': 'tasks.add', 'hard': '3.14', 'soft': '3'}) # You can omit soft or hard
resp.json()
# # Worker API
# Once for the documentation
worker_api = '{}/worker'.format(api_root)
# ## workers
# List workers.
url = '{}/workers'.format(api_root) # Only one not under /worker
print(url)
resp = requests.get(url)
workers = resp.json()
workers
# ## pool/shutdown
# Shutdown a worker.
worker = workers.keys()[0]
url = '{}/shutdown/{}'.format(worker_api, worker)
print(url)
resp = requests.post(url)
resp.json()
# ## pool/restart
# Restart a worker pool, you need to have [CELERYD_POOL_RESTARTS](http://docs.celeryproject.org/en/latest/configuration.html#std:setting-CELERYD_POOL_RESTARTS) enabled in your configuration).
pool_api = '{}/pool'.format(worker_api)
url = '{}/restart/{}'.format(pool_api, worker)
print(url)
resp = requests.post(url)
resp.json()
# ## pool/grow
# Grows worker pool.
url = '{}/grow/{}'.format(pool_api, worker)
print(url)
resp = requests.post(url, params={'n': '10'})
resp.json()
# ## pool/shrink
# Shrink worker pool.
url = '{}/shrink/{}'.format(pool_api, worker)
print(url)
resp = requests.post(url, params={'n': '3'})
resp.json()
# ## pool/autoscale
# [Autoscale](http://docs.celeryproject.org/en/latest/userguide/workers.html#autoscaling) a pool.
url = '{}/autoscale/{}'.format(pool_api, worker)
print(url)
resp = requests.post(url, params={'min': '3', 'max': '10'})
resp.json()
# ## queue/add-consumer
# [Add a consumer](http://docs.celeryproject.org/en/latest/userguide/workers.html#std:control-add_consumer) to a queue.
queue_api = '{}/queue'.format(worker_api)
url = '{}/add-consumer/{}'.format(queue_api, worker)
print(url)
resp = requests.post(url, params={'queue': 'jokes'})
resp.json()
# ## queue/cancel-consumer
# [Cancel a consumer](http://docs.celeryproject.org/en/latest/userguide/workers.html#queues-cancelling-consumers) queue.
url = '{}/cancel-consumer/{}'.format(queue_api, worker)
print(url)
resp = requests.post(url, params={'queue': 'jokes'})
resp.json()
# # Queue API
#
# We assume that we've two queues; the default one 'celery' and 'all'
url = '{}/queues/length'.format(api_root)
print(url)
resp = requests.get(url)
resp.json()
|
docs/api.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LSTM for Part-of-Speech Tagging
#
# In this section, we will use an LSTM to predict part-of-speech tags for words. What exactly is part-of-speech tagging?
#
# Part of speech tagging is the process of determining the *category* of a word from the words in its surrounding context. You can think of part of speech tagging as a way to go from words to their [Mad Libs](https://en.wikipedia.org/wiki/Mad_Libs#Format) categories. Mad Libs are incomplete short stories that have many words replaced by blanks. Each blank has a specified word-category, such as `"noun"`, `"verb"`, `"adjective"`, and so on. One player asks another to fill in these blanks (prompted only by the word-category) until they have created a complete, silly story of their own. Here is an example of such categories:
#
# ```text
# Today, you'll be learning how to [verb]. It may be a [adjective] process, but I think it will be rewarding!
# If you want to take a break you should [verb] and treat yourself to some [plural noun].
# ```
# ... and a set of possible words that fall into those categories:
# ```text
# Today, you'll be learning how to code. It may be a challenging process, but I think it will be rewarding!
# If you want to take a break you should stretch and treat yourself to some puppies.
# ```
#
#
# ### Why Tag Speech?
#
# Tagging parts of speech is often used to help disambiguate natural language phrases because it can be done quickly and with high accuracy. It can help answer: what subject is someone talking about? Tagging can be used for many NLP tasks like creating new sentences using a sequence of tags that make sense together, filling in a Mad Libs style game, and determining correct pronunciation during speech synthesis. It is also used in information retrieval, and for word disambiguation (ex. determining when someone says *right* like the direction versus *right* like "that's right!").
#
# ---
#
# ### Preparing the Data
#
# Now, we know that neural networks do not do well with words as input and so our first step will be to prepare our training data and map each word to a numerical value.
#
# We start by creating a small set of training data, you can see that this is a few simple sentences broken down into a list of words and their corresponding word-tags. Note that the sentences are turned into lowercase words using `lower()` and then split into separate words using `split()`, which splits the sentence by whitespace characters.
#
# #### Words to indices
#
# Then, from this training data, we create a dictionary that maps each unique word in our vocabulary to a numerical value; a unique index `idx`. We do the same for each word-tag, for example: a noun will be represented by the number `1`.
# +
# import resources
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import matplotlib.pyplot as plt
# %matplotlib inline
# +
# training sentences and their corresponding word-tags
training_data = [
("The cat ate the cheese".lower().split(), ["DET", "NN", "V", "DET", "NN"]),
("She read that book".lower().split(), ["NN", "V", "DET", "NN"]),
("The dog loves art".lower().split(), ["DET", "NN", "V", "NN"]),
("The elephant answers the phone".lower().split(), ["DET", "NN", "V", "DET", "NN"])
]
# create a dictionary that maps words to indices
word2idx = {}
for sent, tags in training_data:
for word in sent:
if word not in word2idx:
word2idx[word] = len(word2idx)
# create a dictionary that maps tags to indices
tag2idx = {"DET": 0, "NN": 1, "V": 2}
# -
# Next, print out the created dictionary to see the words and their numerical values!
#
# You should see every word in our training set and its index value. Note that the word "the" only appears once because our vocabulary only includes *unique* words.
# print out the created dictionary
print(word2idx)
# +
import numpy as np
# a helper function for converting a sequence of words to a Tensor of numerical values
# will be used later in training
def prepare_sequence(seq, to_idx):
'''This function takes in a sequence of words and returns a
corresponding Tensor of numerical values (indices for each word).'''
idxs = [to_idx[w] for w in seq]
idxs = np.array(idxs)
return torch.from_numpy(idxs)
# -
# check out what prepare_sequence does for one of our training sentences:
example_input = prepare_sequence("The dog answers the phone".lower().split(), word2idx)
print(example_input)
# ---
# ## Creating the Model
#
# Our model will assume a few things:
# 1. Our input is broken down into a sequence of words, so a sentence will be [w1, w2, ...]
# 2. These words come from a larger list of words that we already know (a vocabulary)
# 3. We have a limited set of tags, `[NN, V, DET]`, which mean: a noun, a verb, and a determinant (words like "the" or "that"), respectively
# 4. We want to predict\* a tag for each input word
#
# \* To do the prediction, we will pass an LSTM over a test sentence and apply a softmax function to the hidden state of the LSTM; the result is a vector of tag scores from which we can get the predicted tag for a word based on the *maximum* value in this distribution of tag scores.
#
# Mathematically, we can represent any tag prediction $\hat{y}_i$ as:
#
# \begin{align}\hat{y}_i = \text{argmax}_j \ (\log \text{Softmax}(Ah_i + b))_j\end{align}
#
# Where $A$ is a learned weight and $b$, a learned bias term, and the hidden state at timestep $i$ is $h_i$.
#
#
# ### Word embeddings
#
# We know that an LSTM takes in an expected input size and hidden_dim, but sentences are rarely of a consistent size, so how can we define the input of our LSTM?
#
# Well, at the very start of this net, we'll create an `Embedding` layer that takes in the size of our vocabulary and returns a vector of a specified size, `embedding_dim`, for each word in an input sequence of words. It's important that this be the first layer in this net. You can read more about this embedding layer in [the PyTorch documentation](https://pytorch.org/tutorials/beginner/nlp/word_embeddings_tutorial.html#word-embeddings-in-pytorch).
#
# Pictured below is the expected architecture for this tagger model.
#
# <img src='images/speech_tagger.png' height=60% width=60% >
#
class LSTMTagger(nn.Module):
def __init__(self, embedding_dim, hidden_dim, vocab_size, tagset_size):
''' Initialize the layers of this model.'''
super(LSTMTagger, self).__init__()
self.hidden_dim = hidden_dim
# embedding layer that turns words into a vector of a specified size
self.word_embeddings = nn.Embedding(vocab_size, embedding_dim)
# the LSTM takes embedded word vectors (of a specified size) as inputs
# and outputs hidden states of size hidden_dim
self.lstm = nn.LSTM(embedding_dim, hidden_dim)
# the linear layer that maps the hidden state output dimension
# to the number of tags we want as output, tagset_size (in this case this is 3 tags)
self.hidden2tag = nn.Linear(hidden_dim, tagset_size)
# initialize the hidden state (see code below)
self.hidden = self.init_hidden()
def init_hidden(self):
''' At the start of training, we need to initialize a hidden state;
there will be none because the hidden state is formed based on perviously seen data.
So, this function defines a hidden state with all zeroes and of a specified size.'''
# The axes dimensions are (n_layers, batch_size, hidden_dim)
return (torch.zeros(1, 1, self.hidden_dim),
torch.zeros(1, 1, self.hidden_dim))
def forward(self, sentence):
''' Define the feedforward behavior of the model.'''
# create embedded word vectors for each word in a sentence
embeds = self.word_embeddings(sentence)
# get the output and hidden state by passing the lstm over our word embeddings
# the lstm takes in our embeddings and hiddent state
lstm_out, self.hidden = self.lstm(
embeds.view(len(sentence), 1, -1), self.hidden)
# get the scores for the most likely tag for a word
tag_outputs = self.hidden2tag(lstm_out.view(len(sentence), -1))
tag_scores = F.log_softmax(tag_outputs, dim=1)
return tag_scores
# ## Define how the model trains
#
# To train the model, we have to instantiate it and define the loss and optimizers that we want to use.
#
# First, we define the size of our word embeddings. The `EMBEDDING_DIM` defines the size of our word vectors for our simple vocabulary and training set; we will keep them small so we can see how the weights change as we train.
#
# **Note: the embedding dimension for a complex dataset will usually be much larger, around 64, 128, or 256 dimensional.**
#
#
# #### Loss and Optimization
#
# Since our LSTM outputs a series of tag scores with a softmax layer, we will use `NLLLoss`. In tandem with a softmax layer, NLL Loss creates the kind of cross entropy loss that we typically use for analyzing a distribution of class scores. We'll use standard gradient descent optimization, but you are encouraged to play around with other optimizers!
# +
# the embedding dimension defines the size of our word vectors
# for our simple vocabulary and training set, we will keep these small
EMBEDDING_DIM = 6
HIDDEN_DIM = 6
# instantiate our model
model = LSTMTagger(EMBEDDING_DIM, HIDDEN_DIM, len(word2idx), len(tag2idx))
# define our loss and optimizer
loss_function = nn.NLLLoss()
optimizer = optim.SGD(model.parameters(), lr=0.1)
# -
# Just to check that our model has learned something, let's first look at the scores for a sample test sentence *before* our model is trained. Note that the test sentence *must* be made of words from our vocabulary otherwise its words cannot be turned into indices.
#
# The scores should be Tensors of length 3 (for each of our tags) and there should be scores for each word in the input sentence.
#
# For the test sentence, "The cheese loves the elephant", we know that this has the tags (DET, NN, V, DET, NN) or `[0, 1, 2, 0, 1]`, but our network does not yet know this. In fact, in this case, our model starts out with a hidden state of all zeroes and so all the scores and the predicted tags should be low, random, and about what you'd expect for a network that is not yet trained!
# +
test_sentence = "The cheese loves the elephant".lower().split()
# see what the scores are before training
# element [i,j] of the output is the *score* for tag j for word i.
# to check the initial accuracy of our model, we don't need to train, so we use model.eval()
inputs = prepare_sequence(test_sentence, word2idx)
inputs = inputs.to(torch.int64)
tag_scores = model(inputs)
print(tag_scores)
# tag_scores outputs a vector of tag scores for each word in an inpit sentence
# to get the most likely tag index, we grab the index with the maximum score!
# recall that these numbers correspond to tag2idx = {"DET": 0, "NN": 1, "V": 2}
_, predicted_tags = torch.max(tag_scores, 1)
print('\n')
print('Predicted tags: \n',predicted_tags)
# -
# ---
# ## Train the Model
#
# Loop through all our training data for multiple epochs (again we are using a small epoch value for this simple training data). This loop:
#
# 1. Prepares our model for training by zero-ing the gradients
# 2. Initializes the hidden state of our LSTM
# 3. Prepares our data for training
# 4. Runs a forward pass on our inputs to get tag_scores
# 5. Calculates the loss between tag_scores and the true tag
# 6. Updates the weights of our model using backpropagation
#
# In this example, we are printing out the average epoch loss, every 20 epochs; you should see it decrease over time.
# +
# normally these epochs take a lot longer
# but with our toy data (only 3 sentences), we can do many epochs in a short time
n_epochs = 300
for epoch in range(n_epochs):
epoch_loss = 0.0
# get all sentences and corresponding tags in the training data
for sentence, tags in training_data:
# zero the gradients
model.zero_grad()
# zero the hidden state of the LSTM, this detaches it from its history
model.hidden = model.init_hidden()
# prepare the inputs for processing by out network,
# turn all sentences and targets into Tensors of numerical indices
sentence_in = prepare_sequence(sentence, word2idx)
targets = prepare_sequence(tags, tag2idx)
# forward pass to get tag scores
tag_scores = model(sentence_in.to(torch.int64))
# compute the loss, and gradients
loss = loss_function(tag_scores, targets.to(torch.int64))
epoch_loss += loss.item()
loss.backward()
# update the model parameters with optimizer.step()
optimizer.step()
# print out avg loss per 20 epochs
if(epoch%20 == 19):
print("Epoch: %d, loss: %1.5f" % (epoch+1, epoch_loss/len(training_data)))
# -
# ## Testing
#
# See how your model performs *after* training. Compare this output with the scores from before training, above.
#
# Again, for the test sentence, "The cheese loves the elephant", we know that this has the tags (DET, NN, V, DET, NN) or `[0, 1, 2, 0, 1]`. Let's see if our model has learned to find these tags!
# +
test_sentence = "The cheese loves the elephant".lower().split()
# see what the scores are after training
inputs = prepare_sequence(test_sentence, word2idx)
inputs = inputs.to(torch.int64)
tag_scores = model(inputs)
print(tag_scores)
# print the most likely tag index, by grabbing the index with the maximum score!
# recall that these numbers correspond to tag2idx = {"DET": 0, "NN": 1, "V": 2}
_, predicted_tags = torch.max(tag_scores, 1)
print('\n')
print('Predicted tags: \n',predicted_tags)
# -
# ## Great job!
#
# To improve this model, see if you can add sentences to this model and create a more robust speech tagger. Try to initialize the hidden state in a different way or play around with the optimizers and see if you can decrease model loss even faster.
|
CVND_Exercises/2_4_LSTMs/2. LSTM Training, Part of Speech Tagging.ipynb
|
# # Non-negative matrix factorization
# Reference: [Non-negative matrix factorization](http://scikit-learn.org/stable/modules/decomposition.html#non-negative-matrix-factorization-nmf-or-nnmf)
#
# NMF is an alternative approach to decomposition that assumes that the data and the components are non-negative. NMF can be plugged in instead of PCA or its variants, in the cases where the data matrix does not contain negative values. It finds a decomposition of samples X into two matrices W and H of non-negative elements, by optimizing for the squared Frobenius norm:
#
# $$
# \arg\min_{W,H} \frac{1}{2} ||X - WH||_{Fro}^{2}
# = \frac{1}{2} \sum_{i,j} (X_{ij} - {WH}_{ij})^{2}
# $$
#
# This norm is an obvious extension of the Euclidean norm to matrices. (Other optimization objectives have been suggested in the NMF literature, in particular Kullback-Leibler divergence, but these are not currently implemented.)
#
# Using: [sklearn.decomposition.NMF](http://scikit-learn.org/stable/modules/generated/sklearn.decomposition.NMF.html)
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import os, sys
import numpy as np
from skimage import io
import matplotlib.pylab as plt
sys.path += [os.path.abspath('.'), os.path.abspath('..')] # Add path to root
import notebooks.notebook_utils as uts
# ## load datset
# +
p_dataset = os.path.join(uts.DEFAULT_PATH, uts.SYNTH_DATASETS_FUZZY[0])
print ('loading dataset: ({}) exists -> {}'.format(os.path.exists(p_dataset), p_dataset))
p_atlas = os.path.join(uts.DEFAULT_PATH, 'dictionary/atlas.png')
atlas_gt = io.imread(p_atlas)
nb_patterns = len(np.unique(atlas_gt))
print ('loading ({}) <- {}'.format(os.path.exists(p_atlas), p_atlas))
plt.imshow(atlas_gt, interpolation='nearest')
_ = plt.title('Atlas; unique %i' % nb_patterns)
# -
list_imgs = uts.load_dataset(p_dataset)
print ('loaded # images: ', len(list_imgs))
img_shape = list_imgs[0].shape
print ('image shape:', img_shape)
# ## Pre-Processing
# +
X = np.array([im.ravel() for im in list_imgs]) # - 0.5
print ('input data shape:', X.shape)
plt.figure(figsize=(7, 3))
_= plt.imshow(X, aspect='auto'), plt.xlabel('features'), plt.ylabel('samples'), plt.colorbar()
# -
uts.show_sample_data_as_imgs(X, img_shape, nb_rows=1, nb_cols=5)
# ## Non-negative matrix factorization
# +
from sklearn.decomposition import NMF
nmf = NMF(n_components=nb_patterns, max_iter=99)
X_new = nmf.fit_transform(X[:1200,:])
print ('fitting parameters:', nmf.get_params())
print ('number of iteration:', nmf.n_iter_)
# -
# **show the estimated components - dictionary**
# +
comp = nmf.components_
coefs = np.sum(np.abs(X_new), axis=0)
print ('estimated component matrix:', comp.shape)
compSorted = [c[0] for c in sorted(zip(comp, coefs), key=lambda x: x[1], reverse=True) ]
uts.show_sample_data_as_imgs(np.array(compSorted), img_shape, nb_cols=nb_patterns, bool_clr=True)
# +
ptn_used = np.sum(np.abs(X_new), axis=0) > 0
atlas_ptns = comp[ptn_used, :].reshape((-1, ) + list_imgs[0].shape)
atlas_ptns = comp.reshape((-1, ) + list_imgs[0].shape)
atlas_estim = np.argmax(atlas_ptns, axis=0) + 1
atlas_sum = np.sum(np.abs(atlas_ptns), axis=0)
atlas_estim[atlas_sum < 1e-1] = 0
_ = plt.imshow(atlas_estim, interpolation='nearest'), plt.colorbar()
# -
l_idx = [1, 30, 40]
fig, axr = plt.subplots(len(l_idx), 2, figsize=(10, 2.5 * len(l_idx)))
for i, idx in enumerate(l_idx):
axr[i, 0].plot(atlas_ptns[:,:,idx].T), axr[i, 0].set_xlim([0, 63])
axr[i, 0].set_title('col: {}'.format(idx))
axr[i, 1].plot(np.abs(atlas_ptns[:,idx,:].T)), axr[i, 1].set_xlim([0, 63])
axr[i, 1].set_title('sum over comp.')
# **particular coding of each sample**
# +
plt.figure(figsize=(6, 3))
plt.imshow(X_new[:20,:], interpolation='nearest', aspect='auto'), plt.colorbar()
_= plt.xlabel('components'), plt.ylabel('samples')
coefs = np.sum(np.abs(X_new), axis=0)
# print 'used coeficients:', coefs.tolist()
# -
# **backword reconstruction from encoding and dictionary**
res = np.dot(X_new, comp)
print ('model applies by reverting the unmixing', res.shape)
uts.show_sample_data_as_imgs(res, img_shape, nb_rows=2, nb_cols=3, bool_clr=True)
|
notebooks/method_NMF.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Line-profiler
#
# This tutorial covers the basics of the [line_profiler](https://github.com/rkern/line_profiler) Python package. I'll cover what exactly line-profiler is (and why you should care) and how to use it.
#
#
# ## What is line-profiler
#
# As the name suggests line-profiler profiles the runtime of a Python script in a line-by-line manner. From the profiling you can identify **hotspots**; i.e., what lines of code take the longest time to run! While sometimes this may be of simple academic interest, alternative algorithms could be used and lead to significant time savings. If a loop is executed a million times and you save 10ms on each loop, congratulations your code now finishes three hours sooner!
#
# ## Installing line-profiler
#
# Provided you have a working Python distribution, installing should be as easy as
# ```
# $ pip install line_profiler
# ```
#
# If this fails, you can also clone the Github repository:
#
# ```
# $ git clone https://github.com/rkern/line_profiler.git
# ```
#
# And then install using
#
# ```
# $ python setup.py install
# ```
#
# remembering you may need the ```--user``` or ```--prefix=/directory/to/local/python/site-packages/``` argument if you're on a cluster such as ```g2```.
#
#
# ## Using line-profiler with Jupyter Notebook
#
# When using line-profiler there is a slight twist depending on whether you're running the Python script directly from the terminal (e.g., ``python my_script.py``) or from inside a Jupyter notebook. I'll go over both of these methods before showing some examples from my own code (which is all executed via command line).
#
# Loading line-profiler into your notebook is one easy command:
# %load_ext line_profiler
# Next we need some functions to test our run-time on.
#
# For the first example, say we are given particle positions ``x``, ``y`` and ``z`` and we wish to calculate the pair-wise distance between each particle. The pair-wise distance is a very useful property which allows us to compute further statistics such as the correlation function or power spectrum, and is necessary for things such as force calculations.
# +
from __future__ import print_function # Always do this >:(
from __future__ import division
import numpy as np
pos = np.random.random((1000,3)) # Generate a 3D vector of random points between 0 and 1.
print(pos[0])
print(pos[0,0])
# -
# First we'll do my favourite method; the crass, brute force method that involves nested ``for`` loops.
def pairwise_brute(pos):
npart = pos.shape[0]
ndim = pos.shape[1]
distance = np.empty((npart, npart), dtype = np.float64)
# Note that the distance between pair i-j is the same as j-i.
for i in range(npart):
for j in range(npart):
d = 0.0
for k in range(ndim):
d += pow(pos[i,k] - pos[j,k], 2) # Gets the (square) distance in one dimension between particles i and j.
distance[i,j] = np.sqrt(d)
assert(distance[i,i] == 0) # A particle should have zero distance from itself.
return distance
distance = pairwise_brute(pos)
print(distance[0,100]) # This is the distance between particle 0 and 100.
# Then we run ```line-profiler``` on our code. The argument ``-f`` specifies the function that we want to profile and we need to remember to provide the input arguments.
# %lprun -f pairwise_brute pairwise_brute(pos)
# + active=""
# Timer unit: 1e-06 s
#
# Total time: 9.24807 s
# File: <ipython-input-37-70d496d08d07>
# Function: pairwise_brute at line 1
#
# Line # Hits Time Per Hit % Time Line Contents
# ==============================================================
# 1 def pairwise_brute(pos):
# 2 1 3 3.0 0.0 npart = pos.shape[0]
# 3 1 1 1.0 0.0 ndim = pos.shape[1]
# 4 1 677 677.0 0.0 distance = np.empty((npart, npart), dtype = np.float64)
# 5
# 6 # Note that the distance between pair i-j is the same as j-i.
# 7 1001 722 0.7 0.0 for i in range(npart):
# 8 1001000 480533 0.5 5.2 for j in range(npart):
# 9 1000000 415336 0.4 4.5 d = 0.0
# 10 4000000 2215483 0.6 24.0 for k in range(ndim):
# 11 3000000 4031247 1.3 43.6 d += pow(pos[i,k] - pos[j,k], 2) # Gets the (square) distance in one dimension between particles i and j.
# 12 1000000 2102190 2.1 22.7 distance[i,j] = np.sqrt(d)
# 13 1000 1875 1.9 0.0 assert(distance[i,i] == 0) # A particle should have zero distance from itself.
# 14 1 1 1.0 0.0 return distance
# -
# Be careful that in the above table the **Time** column is expressed in units of $10^{-6}s$, i.e., $\mu s$. The **Time** column tells us how many $\mu s$ we spend on each line (total), the **Per Hit** column tells us the average amount of time spent on relative to the total amount of time spent in the function and finally the **% Time** column is the percentage of time spent on that line (relative to the total amount of time).
#
# From this table a number of things become apparent. Firstly, I enjoy writing long comments. Secondly, one of the most expensive line of code is line 13 where we perform our ``assert`` error check. This really highlights the power of a tool like line-profiler because whilst this line is expensive (taking $1.9\mu s$ per hit), the **relative** amount of time spent on this operation is small. The honour of most expensive operation goes to line 11 in which we spend $44\%$(!!) of our time on. This is a result of our nested ``for`` loops; we are hitting line 11 a grand total of $4031247$ times.
#
# Let's see if we can do better.
def pairwise_numpy(pos):
padded_pos = pos[:, None, :] # Pad the array to ensure the dimensionality will be correct
distance = np.sqrt(np.sum(np.square(padded_pos - pos), axis = -1)) # Perform Pythag and then sum to return to the original dimensions.
return distance
# %lprun -f pairwise_numpy pairwise_numpy(pos)
# + active=""
# Timer unit: 1e-06 s
#
# Total time: 0.050307 s
# File: <ipython-input-77-fb2fce4afd83>
# Function: pairwise_numpy at line 1
#
# Line # Hits Time Per Hit % Time Line Contents
# ==============================================================
# 1 def pairwise_numpy(pos):
# 2 1 8 8.0 0.0 padded_pos = pos[:, None, :] # Pad the array to ensure the dimensionality will be correct
# 3 1 50297 50297.0 100.0 distance = np.sqrt(np.sum(np.square(padded_pos - pos), axis = -1)) # Perform Pythag and then sum to return to the original dimensions.
# 4 1 2 2.0 0.0 return distance
# -
# Huzzah we have remarkable speed-up relative to the brute force method.
# **Beware:** When using Numpy broadcasting, temporary arrays are created potentially causing memory issues for (very) large arrays.
# ## Using line-profiler with terminal
#
# If you're like me and behind the times, you can also run line-profiler from the terminal. For this all you do is add the decorator ``@profile`` before the function you want to profile. Then you profile the results by
#
# ```
# $ kernprof -l my_script.py <Script Arguments>
# ```
#
# If you get the error '``bash: kernprof: command not found``' then fully specify the path to where you installed ``line_profiler``:
#
# ```
# $ python3 /home/jseiler/.local/lib/python3.5/site-packages/kernprof.py -l rewrite_files.py 15 15
# ```
#
# Once the profiling has completed you can view the results via
#
# ```
# $ python3 -m line_profiler my_script.py.lprof
# ```
#
# As an example from my own code (I've only included the lines with $>0.0$ in the **% Time** column):
#
# + active=""
# Timer unit: 1e-06 s
#
# Total time: 2114.36 s
# File: rewrite_files.py
# Function: link_fof_snapshot_full at line 322
#
# Line # Hits Time Per Hit % Time Line Contents
# ==============================================================
# 386 256 4468354 17454.5 0.2 with h5py.File(fname, 'r') as f:
# 390 256 6629947 25898.2 0.3 BoxSize = f['Header'].attrs['BoxSize'] # Size of the simulation box (in Mpc/h).
# 391 256 38943 152.1 0.0 Time = f['Header'].attrs['Time'] # Scale factor.
# 392 256 37268 145.6 0.0 Omega_m = f['Header'].attrs['Omega0'] # Matter density.
# 393 256 36343 142.0 0.0 Omega_l = f['Header'].attrs['OmegaLambda'] # Dark energy density.
# 394 256 36325 141.9 0.0 particle_mass = f['Header'].attrs['MassTable'] # Table of particle masses (length equal to TypeMax)
# 406 506 6359047 12567.3 0.3 snapshot_partid = particles['ParticleIDs']
# 407 506 1140906728 2254756.4 54.0 ids_found = (np.nonzero(np.in1d(snapshot_partid, fof_partid[type_idx], assume_unique = True)))[0] # np.in1d returns a True if the snapshot particle is found in the FoF list (False otherwise). np.nonzero then returns the indices of those values that have a 'True'.
# 424 228 16848850 73898.5 0.8 particle_position[type_idx].append(particles['Coordinates'][ids_found]) # For some super weird reason this needs 3 indices to properly reference. I.e. particle_position[1][0] will return all the coordinates for Particle Type 1.
# 425 228 17104189 75018.4 0.8 particle_velocity[type_idx].append(particles['Velocities'][ids_found])
# 426 228 1977059 8671.3 0.1 particle_id[type_idx].append(particles['ParticleIDs'][ids_found])
# 429 228 915102216 4013606.2 43.3 fof_partid_idx = np.int32(np.nonzero(np.in1d(fof_partid[type_idx], snapshot_partid, assume_unique = True))[0])
# 473 256 1306722 5104.4 0.1 print("Written Data to {0}.".format(fname))
# -
# From this we can see that a massive $97.3\%$ of time is spent doing two ```np.nonzero``` calculations. This really puts things into perspective: if I wanted to optimize my code these are **the** lines I would start with. It isn't even worth thinking about optimizing the other parts of my code when they form such a neglible percentage of time spent.
#
# ## Wrapping Up
#
# Hopefully this tutorial has given a basic demonstration how to use ```line_profiler```. More importantly, I hope it's outlined the use of such a tool; namely identifying hotspots within your code so you know **where** to start optimization. Blindly going through every line in your code and asking "Could I make this faster?" is a painful exercise and you often won't see a noticable speedup in your code.
#
# As always, the [Github page](https://github.com/rkern/line_profiler) for the package will contain a wealth of information on running the code and some tips and tricks.
|
code-review_archive/2017--2018/2017_12_07/line_profiler.py.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install html5lib beautifulsoup4
# !pip install selenium
from selenium import webdriver
from bs4 import BeautifulSoup
from time import sleep
import requests
import json
import re
import pandas as pd
from urllib import request
from selenium import webdriver
from selenium.webdriver.support.ui import WebDriverWait
from selenium.common.exceptions import NoSuchElementException
from selenium.webdriver.common.keys import Keys
idol_list = pd.read_csv('idol_list.csv', encoding = 'euc-kr')
idol_list.shape
idol_list.columns
artists = idol_list.artist
artists[1]
driver = webdriver.Chrome('C:/Users/pje17/Desktop/Lyrics/Crawlling/chromedriver')
driver.get("https://www.melon.com/search/total/index.htm?q=%EC%86%8C%EB%85%80%EC%8B%9C%EB%8C%80§ion=&searchGnbYn=Y&kkoSpl=N&kkoDpType=&ipath=srch_form")
artist_id = []
# +
# n=0
for artist in artists:
sleep(3)
search = driver.find_element_by_xpath('//*[@id="top_search"]')
search.send_keys(Keys.CONTROL + "a");
search.clear()
# WebDriverWait(driver, timeout)
search.send_keys(artist)
# 아티스트명 검색
search.send_keys(Keys.RETURN)
try:
# 아티스트 클릭
driver.find_element_by_xpath('//*[@id="conts"]/div[3]/div[1]/div[1]/div/a/strong').click()
url = driver.current_url
id = url.replace('https://www.melon.com/artist/timeline.htm?artistId=', '')
artist_id.append(id)
except NoSuchElementException:
artist_id.append('')
# print(artist_id[n])
# n+=1
# -
len(artist_id)
idol_id = artist_id.copy()
# +
idol_id = pd.DataFrame(idol_id)
idol_id.head()
# -
idol_id.columns = ['idol_id']
idol_id.head()
idol_list.head()
idol_list['idol_id'] = 0
len(idol_id)
idol_list.update(idol_id)
idol_list
idol_list.to_csv('idol_list_with_id.csv')
# ## id로 가사 크롤링
driver = webdriver.Chrome('C:/Users/pje17/Desktop/Lyrics/Crawlling/chromedriver')
driver.get("https://www.melon.com/artist/song.htm?artistId=780066#params%5BlistType%5D=A¶ms%5BorderBy%5D=ISSUE_DATE¶ms%5BartistId%5D=780066&po=pageObj&startIndex=1")
# +
# n=0
for artist in artists:
sleep(3)
search = driver.find_element_by_xpath('//*[@id="top_search"]')
search.send_keys(Keys.CONTROL + "a");
search.clear()
# WebDriverWait(driver, timeout)
search.send_keys(artist)
# 아티스트명 검색
search.send_keys(Keys.RETURN)
try:
# 아티스트 클릭
driver.find_element_by_xpath('//*[@id="conts"]/div[3]/div[1]/div[1]/div/a/strong').click()
url = driver.current_url
id = url.replace('https://www.melon.com/artist/timeline.htm?artistId=', '')
artist_id.append(id)
except NoSuchElementException:
artist_id.append('')
# print(artist_id[n])
# n+=1
|
Crawling/url_crawling/melon_urlid_crawling_SoHyun_ver01.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from __future__ import division
import pickle
import os
from sklearn import metrics
import numpy as np
import pandas as pd
from lentil import evaluate
from lentil import models
import mem
# -
from matplotlib import pyplot as plt
import seaborn as sns
# %matplotlib inline
import matplotlib as mpl
mpl.rc('savefig', dpi=300)
mpl.rc('text', usetex=True)
mpl.rc('text.latex', preamble='\usepackage{amsfonts}')
import matplotlib.lines as mlines
import logging
logging.getLogger().setLevel(logging.DEBUG)
with open(os.path.join('data', 'dutch_big_history.pkl'), 'rb') as f:
history = pickle.load(f)
# Build content features for `mnemosyne_v2`
with open(os.path.join('data', 'content_features.pkl'), 'rb') as f:
contents_of_item_id = pickle.load(f)
content_features = {k: (len(f) if f is not None else len(b)) for k, (b, f) in contents_of_item_id.iteritems()}
# Build content features for `dutch_big`
with open(os.path.join('data', 'original_of_module_id.pkl'), 'rb') as f:
original_of_module_id = pickle.load(f)
embedding_of_word = {}
with open(os.path.join('data', 'embeddings', 'cbow', 'size=50.embeddings'), 'rb') as f:
for line in f:
fields = line.strip().split(' ')
embedding_of_word[fields[0]] = np.array([float(x) for x in fields[1:]])
count_of_word = {}
with open(os.path.join('data', 'embeddings', 'cbow', 'unigram_frequencies'), 'rb') as f:
for line in f:
fields = line.strip().split(' ')
count_of_word[fields[0]] = int(fields[1])
total_count = sum(count_of_word.itervalues())
freq_of_word = {k: (v / total_count) for k, v in count_of_word.iteritems()}
content_features = {k: np.append(
embedding_of_word[original_of_module_id[k]],
[len(original_of_module_id[k]), freq_of_word[original_of_module_id[k]]]) \
for k in history.data['module_id'].unique()}
content_features = {k: np.array(
[len(original_of_module_id[k]), freq_of_word[original_of_module_id[k]]]) \
for k in history.data['module_id'].unique()}
content_features = {k: np.array(
[len(original_of_module_id[k])]) \
for k in history.data['module_id'].unique()}
# Setup the IRT benchmark models and memory models
# +
def build_1pl_irt_model(history, filtered_history, split_history=None):
model = models.OneParameterLogisticModel(
filtered_history, select_regularization_constant=True, name_of_user_id='user_id')
model.fit()
return model
def build_2pl_irt_model(history, filtered_history, split_history=None):
model = models.TwoParameterLogisticModel(
filtered_history, select_regularization_constant=True, name_of_user_id='user_id')
model.fit()
return model
def build_student_biased_coin_model(history, filtered_history, split_history=None):
model = models.StudentBiasedCoinModel(history, filtered_history, name_of_user_id='user_id')
model.fit()
return model
def build_assessment_biased_coin_model(history, filtered_history, split_history=None):
model = models.AssessmentBiasedCoinModel(history, filtered_history)
model.fit()
return model
def meta_build_efc_model(
strength_model='deck', using_delay=True,
using_global_difficulty=True, debug_mode_on=True, content_features=None,
coeffs_regularization_constant=1e-6, item_bias_regularization_constant=1e-6,
using_item_bias=True):
def build_efc_model(history, filtered_history, split_history=None):
model = mem.EFCModel(
filtered_history, strength_model=strength_model, using_delay=using_delay,
using_global_difficulty=using_global_difficulty, debug_mode_on=debug_mode_on,
content_features=content_features, using_item_bias=using_item_bias)
model.fit(
learning_rate=0.1,
#learning_rate=(1 if not using_global_difficulty else 0.1),
ftol=1e-6, max_iter=10000,
coeffs_regularization_constant=coeffs_regularization_constant,
item_bias_regularization_constant=item_bias_regularization_constant)
return model
return build_efc_model
def meta_build_logistic_regression_model(C=1.0):
def build_logistic_regression_model(history, filtered_history, split_history=None):
model = mem.LogisticRegressionModel(filtered_history)
model.fit(C=C)
return model
return build_logistic_regression_model
# -
model_builders = {
'1PL IRT' : build_1pl_irt_model,
'EFC I/-/-' : meta_build_efc_model(
strength_model='deck', using_delay=True, using_global_difficulty=False,
content_features=None, using_item_bias=True,
item_bias_regularization_constant=1e-3),
'EFC I/G/-' : meta_build_efc_model(
strength_model='deck', using_delay=True, using_global_difficulty=True,
content_features=None, using_item_bias=True,
item_bias_regularization_constant=1e-3, coeffs_regularization_constant=1e-3),
'EFC I/G/B' : meta_build_efc_model(
strength_model='deck', using_delay=True, using_global_difficulty=True,
content_features=content_features, using_item_bias=True,
item_bias_regularization_constant=1e-3, coeffs_regularization_constant=1e-3),
'EFC -/G/B' : meta_build_efc_model(
strength_model='deck', using_delay=True, using_global_difficulty=True,
content_features=content_features, using_item_bias=False,
coeffs_regularization_constant=1e-3),
'EFC -/-/B' : meta_build_efc_model(
strength_model='deck', using_delay=True, using_global_difficulty=False,
content_features=content_features, using_item_bias=False,
coeffs_regularization_constant=1e-3)
}
print "Number of models = %d" % (len(model_builders))
print '\n'.join(model_builders.keys())
# Perform the evaluations
results = evaluate.cross_validated_auc(
model_builders,
history,
num_folds=10,
random_truncations=True)
# dump results to file
with open(os.path.join('results', 'dutch_big_lesion_analysis.pkl'), 'wb') as f:
pickle.dump(results, f, pickle.HIGHEST_PROTOCOL)
# load results from file, replacing current results
with open(os.path.join('results', 'dutch_big_lesion_analysis.pkl'), 'rb') as f:
results = pickle.load(f)
df = history.data
# Compute validation AUCs for separate bins of data
def compute_auc(y_trues, probas_pred):
try:
y_trues, probas_pred = zip(*[(y, p) for y, p in zip(y_trues, probas_pred) if not np.isnan(p)])
fpr, tpr, thresholds = metrics.roc_curve(y_trues, probas_pred, pos_label=1)
return metrics.auc(fpr, tpr)
except:
return np.nan
ndata_in_logs = [df['module_id'].ix[idxes].value_counts() for idxes, y_trues, probas_pred in results.train_ixn_data]
ndata_of_val_ixns = [df['module_id'].ix[idxes].apply(lambda x: vc.get(x, 0)) for vc, (idxes, y_trues, probas_pred) in zip(ndata_in_logs, results.val_ixn_data)]
num_bins = 5
hist, bin_edges = np.histogram([y for x in ndata_of_val_ixns for y in x], bins=num_bins)
t = [(x+y)/2 for x, y in zip(bin_edges[:-1], bin_edges[1:])]
# +
model_names = [
'1PL IRT',
'EFC I/-/-',
'EFC I/G/-',
'EFC I/G/B',
'EFC -/G/B',
'EFC -/-/B']
model_labels = [
'1PL IRT',
r'$\gamma_i$',
r'$\gamma_i + \beta_0$',
r'$\gamma_i + \beta_0 + \vec{\beta}_{1:n} \cdot \vec{x}_i$',
r'$\beta_0 + \vec{\beta}_{1:n} \cdot \vec{x}_i$',
r'$\vec{\beta}_{1:n} \cdot \vec{x}_i$']
# +
plt.xlabel(r'$\log{(\theta_i)}$')
plt.boxplot([results.validation_aucs(m) for m in model_names])
plt.scatter(
range(1, len(model_names) + 1),
[results.test_auc(m) for m in model_names],
color='orange', s=100)
plt.xticks(
range(1, len(model_names) + 1),
model_labels, rotation=15)
plt.xlim([0.5, len(model_names) + .5])
orange_circle = mlines.Line2D([], [], color='orange', marker='o', label='Test')
red_line = mlines.Line2D([], [], color='red', marker='_', label='Validation')
plt.legend(handles=[red_line, orange_circle], loc='best')
plt.ylabel('AUC')
plt.savefig(os.path.join('figures', 'dutch_big', 'auc-box-plots-efc-cgi.pdf'), bbox_inches='tight')
plt.show()
# -
label_of_m = dict(zip(model_names, model_labels))
s_of_model = {}
for m in model_names:
s_of_model[m] = [[compute_auc(
[p for p, q in zip(y_trues, vf) if q>=x and (q<y or (bidx==len(bin_edges)-2 and q==y))],
[p for p, q in zip(probas_pred[m], vf) if q>=x and (q<y or (bidx==len(bin_edges)-2 and q==y))]) \
for (_, y_trues, probas_pred), vf in zip(results.val_ixn_data, ndata_of_val_ixns)] \
for bidx, (x, y) in enumerate(zip(bin_edges[:-1], bin_edges[1:]))]
# +
fig, ax1 = plt.subplots()
sns.set_style('dark')
ax2 = ax1.twinx()
ax2.bar(bin_edges[:-1], hist, [y-x for x, y in zip(bin_edges[:-1], bin_edges[1:])], color='gray', alpha=0.5, linewidth=0)
ax2.set_ylabel('Frequency (number of interactions)')
sns.set_style('darkgrid')
lines = []
for m, s1 in s_of_model.iteritems():
l1 = ax1.errorbar(
t, [np.nanmean(z) for z in s1], label='%s' % label_of_m[m],
yerr=[np.nanstd(z)/np.sqrt(len(z)) for z in s1])
lines.append(l1)
ax1.set_xlabel('Number of training logs for item')
ax1.set_ylabel('Validation AUC')
first_legend = plt.legend(handles=lines[:3], loc='lower center', bbox_to_anchor=(0.25, -0.4))
plt.gca().add_artist(first_legend)
plt.legend(handles=lines[3:], loc='lower center', bbox_to_anchor=(0.75, -0.4))
plt.savefig(os.path.join('figures', 'dutch_big', 'auc-vs-ndata.pdf'), bbox_inches='tight')
plt.show()
# -
|
nb/mnemosyne_evaluation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Here we look at the analytic expressions derived in the paper for the time-averaged $\cos^2{\theta}$ when $T = $ integer number of days
# +
from numpy import *
from PlotFuncs import *
import matplotlib.pyplot as plt
import matplotlib.cm as cm
from numpy import random
from tqdm import tqdm
from scipy.integrate import cumtrapz
from scipy.special import erf
from scipy.stats import percentileofscore
from scipy.optimize import fsolve
def FindPercentile(costh_samples,nbins=500):
h,ce = histogram(costh_samples,nbins)
cc = (ce[1:]+ce[0:-1])/2
P_costh = h/sum(h)
fs = lambda P0 : 0.05-sum(0.5*(1+erf(-P0*cc/sqrt(2)))*P_costh)
res = fsolve(fs,1.64*3)
cval_excl = 1.64/res
fs = lambda P0 : 0.05-sum(0.5*(1+erf((5-P0*cc)/sqrt(2)))*P_costh)
res = fsolve(fs,1.64*3+5)
cval_disc = (5+1.64)/res
return cval_excl,cval_disc
# +
# Random DP polarisations sampled isotropically
ngen = 100000
costh = 2*random.uniform(size=ngen)-1
th = arccos(costh)
cos2th = cos(2*th)
# Analytic results (see mathematica notebook for derivation)
costh_av_Z = lambda lat : (1/8)*(3+cos(2*th)-(1+3*cos(2*th))*cos(2*lat*pi/180))
costh_av_N = lambda lat : (1/8)*(3+cos(2*lat*pi/180)+(1+3*cos(2*lat*pi/180))*cos(2*th))
costh_av_W = lambda lat : (sin(th)**2)/2
# Loop over latitudes to show the dependence on lat
nloc = 1000
lats = linspace(-90,90,nloc)
val_excl_N = zeros(shape=nloc)
val_excl_Z = zeros(shape=nloc)
val_excl_W = zeros(shape=nloc)
val_disc_N = zeros(shape=nloc)
val_disc_Z = zeros(shape=nloc)
val_disc_W = zeros(shape=nloc)
# For each latitude calculate 5th percentile
# i.e. the value above which 95% of the distribution lies
for i in tqdm(range(0,nloc)):
Z = costh_av_Z(lats[i])
N = costh_av_N(lats[i])
W = costh_av_W(lats[i])
val_excl_N[i],val_disc_N[i] = FindPercentile(N)
val_excl_W[i],val_disc_W[i] = FindPercentile(W)
val_excl_Z[i],val_disc_Z[i] = FindPercentile(Z)
# +
import matplotlib.patheffects as pe
pek=[pe.Stroke(linewidth=6, foreground='k',alpha=1), pe.Normal()]
# Set up figure
fig,ax = MySquarePlot('Latitude [$^\circ$]',r'$\langle \cos^2{\theta}\rangle^{\rm disc.}_{T = 1{\rm day}}$',tfs=29)
# Plot lines
plt.plot(lats,val_disc_Z,'-',lw=4,color='firebrick',path_effects=pek)
plt.plot(lats,val_disc_W,'-',lw=4,color='steelblue',path_effects=pek)
plt.plot(lats,val_disc_N,'-',lw=4,color='SeaGreen',path_effects=pek)
# Location labels
Sydney = -33.86
LA = 34.0522
TelAviv = 32.0853
Stockholm = 59.3293
plt.text(Sydney+4.5,0.39,'Sydney',va='center',rotation_mode='anchor',rotation=-90)
plt.text(LA+3,0.37,'LA',va='center',rotation_mode='anchor',rotation=-90)
plt.text(TelAviv-3,0.33,'Tel Aviv',va='center',rotation_mode='anchor',rotation=90)
plt.text(Stockholm+4,0.395,'Stockholm',va='center',rotation_mode='anchor',rotation=-90)
# Plot location lines connected to the other lines
plt.plot([Sydney,Sydney],[0.4,val_disc_Z[argmin(abs(Sydney-lats))]],'k--',lw=2,zorder=-10)
plt.plot(Sydney,val_disc_Z[argmin(abs(Sydney-lats))],'o',color='k',markersize=10,mfc='none',mew=3)
plt.plot([TelAviv,TelAviv],[0.4,val_disc_Z[argmin(abs(TelAviv-lats))]],'k--',lw=2,zorder=-10)
plt.plot(TelAviv,val_disc_Z[argmin(abs(TelAviv-lats))],'o',color='k',markersize=10,mfc='none',mew=3)
plt.plot([LA,LA],[0.4,val_disc_Z[argmin(abs(LA-lats))]],'k--',lw=2,zorder=-10)
plt.plot(LA,val_disc_Z[argmin(abs(LA-lats))],'o',color='k',markersize=10,mfc='none',mew=3)
plt.plot([Stockholm,Stockholm],[0.4,val_disc_N[argmin(abs(Stockholm-lats))]],'k--',lw=2,zorder=-10)
plt.plot(Stockholm,val_disc_N[argmin(abs(Stockholm-lats))],'o',color='k',markersize=10,mfc='none',mew=3)
# Labels
plt.text(lats[150]*(1+0.08),val_disc_Z[150]*(1+0.2),r'{\bf Zenith-pointing}',color='firebrick',rotation=73.5,fontsize=28)
plt.text(lats[125],val_disc_W[120]*(1+0.135),r'{\bf West-pointing}',color='steelblue',rotation=0,fontsize=28)
plt.text(lats[100]*(1+0.06),val_disc_N[100]*(1+0.2),r'{\bf North-pointing}',color='SeaGreen',rotation=79,fontsize=28)
plt.gcf().text(0.0,0.5,'w',color='w',zorder=-1000) #<--- this is to stop the ylabel being partially cut off
plt.xlim([-90,90])
plt.ylim([0,0.4])
plt.title(r'{\bf Experiments sensitive to an axis}',fontsize=45,pad=20)
MySaveFig(fig,'LocationDependence')
# -
# # Now plot the values we just calculated
# +
import matplotlib.patheffects as pe
pek=[pe.Stroke(linewidth=6, foreground='k',alpha=1), pe.Normal()]
# Set up figure
fig,ax = MySquarePlot('Latitude [$^\circ$]',r'$\langle \cos^2{\theta}\rangle^{\rm excl.}_{T = 1{\rm day}}$',tfs=29)
# Plot lines
plt.plot(lats,val_excl_Z,'-',lw=4,color='firebrick',path_effects=pek)
plt.plot(lats,val_excl_W,'-',lw=4,color='steelblue',path_effects=pek)
plt.plot(lats,val_excl_N,'-',lw=4,color='SeaGreen',path_effects=pek)
# Location labels
Sydney = -33.86
LA = 34.0522
TelAviv = 32.0853
Stockholm = 59.3293
plt.text(Sydney+4.5,0.39,'Sydney',va='center',rotation_mode='anchor',rotation=-90)
plt.text(LA+3,0.37,'LA',va='center',rotation_mode='anchor',rotation=-90)
plt.text(TelAviv-3,0.33,'Tel Aviv',va='center',rotation_mode='anchor',rotation=90)
plt.text(Stockholm+3,0.397,'Stockholm',va='center',rotation_mode='anchor',rotation=-90,fontsize=24)
# Plot location lines connected to the other lines
plt.plot([Sydney,Sydney],[0.4,val_excl_Z[argmin(abs(Sydney-lats))]],'k--',lw=2,zorder=-10)
plt.plot(Sydney,val_excl_Z[argmin(abs(Sydney-lats))],'o',color='k',markersize=10,mfc='none',mew=3)
plt.plot([TelAviv,TelAviv],[0.4,val_excl_Z[argmin(abs(TelAviv-lats))]],'k--',lw=2,zorder=-10)
plt.plot(TelAviv,val_excl_Z[argmin(abs(TelAviv-lats))],'o',color='k',markersize=10,mfc='none',mew=3)
plt.plot([LA,LA],[0.4,val_excl_Z[argmin(abs(LA-lats))]],'k--',lw=2,zorder=-10)
plt.plot(LA,val_excl_Z[argmin(abs(LA-lats))],'o',color='k',markersize=10,mfc='none',mew=3)
plt.plot([Stockholm,Stockholm],[0.4,val_excl_N[argmin(abs(Stockholm-lats))]],'k--',lw=2,zorder=-10)
plt.plot(Stockholm,val_excl_N[argmin(abs(Stockholm-lats))],'o',color='k',markersize=10,mfc='none',mew=3)
# Labels
plt.text(lats[100]*(1+0.2),val_excl_Z[100]*(1-0.43),r'{\bf Zenith-pointing}',color='firebrick',rotation=73.5,fontsize=28)
plt.text(lats[380],val_excl_W[380]*(1-0.08),r'{\bf West-pointing}',color='steelblue',rotation=0,fontsize=28)
plt.text(lats[20]*(1+0.045),val_excl_N[20]*(1+0.11),r'{\bf North-pointing}',color='SeaGreen',rotation=68,fontsize=28)
plt.gcf().text(0.0,0.5,'w',color='w',zorder=-1000) #<--- this is to stop the ylabel being partially cut off
plt.xlim([-90,90])
plt.ylim([0,0.4])
plt.title(r'{\bf Experiments sensitive to an axis}',fontsize=45,pad=20)
MySaveFig(fig,'LocationDependence_excl')
# -
# # Do exactly the same but for the planar experiments
# +
costh_av_ZW = lambda lat : (1/8)*(5-cos(2*lat*pi/180)-(1+3*cos(2*lat*pi/180))*cos(2*th)) # North-facing
costh_av_ZN = lambda lat : (1/4)*(3+cos(2*th)) # West-facing
costh_av_NW = lambda lat : (1/8)*(5+cos(2*lat*pi/180)+(-1+3*cos(2*lat*pi/180))*cos(2*th)) # Zenith-facing
nloc = 1000
lats = linspace(-90,90,nloc)
val_excl_NW = zeros(shape=nloc)
val_excl_ZW = zeros(shape=nloc)
val_excl_ZN = zeros(shape=nloc)
val_disc_NW = zeros(shape=nloc)
val_disc_ZW = zeros(shape=nloc)
val_disc_ZN = zeros(shape=nloc)
for i in tqdm(range(0,nloc)):
NW = costh_av_NW(lats[i])
ZW = costh_av_ZW(lats[i])
ZN = costh_av_ZN(lats[i])
val_excl_NW[i],val_disc_NW[i] = FindPercentile(NW)
val_excl_ZW[i],val_disc_ZW[i] = FindPercentile(ZW)
val_excl_ZN[i],val_disc_ZN[i] = FindPercentile(ZN)
# +
ymax = 0.8
fig,ax = MySquarePlot('Latitude [$^\circ$]',r'$\langle \cos^2{\theta}\rangle^{\rm excl.}_{T = 1{\rm day}}$',tfs=29)
plt.plot(lats,val_excl_NW,'-',lw=4,color='firebrick',path_effects=pek)
plt.plot(lats,val_excl_ZN,'-',lw=4,color='steelblue',path_effects=pek,zorder=-1)
plt.plot(lats,val_excl_ZW,'-',lw=4,color='seagreen',path_effects=pek)
plt.text(Sydney+4,ymax-0.01,'Sydney',va='center',rotation_mode='anchor',rotation=-90,fontsize=28)
plt.text(LA+3,ymax*(1-0.08),'LA',va='center',rotation_mode='anchor',rotation=-90,fontsize=28)
plt.text(TelAviv-3,ymax-0.13,'Tel Aviv',va='center',rotation_mode='anchor',rotation=90,fontsize=28)
plt.text(Stockholm+3,ymax-0.01,'Stockholm',va='center',rotation_mode='anchor',rotation=-90,fontsize=22)
plt.plot([Sydney,Sydney],[ymax,val_excl_NW[argmin(abs(Sydney-lats))]],'k--',lw=2,zorder=-10)
plt.plot(Sydney,val_excl_NW[argmin(abs(Sydney-lats))],'o',color='k',markersize=10,mfc='none',mew=3)
plt.plot([TelAviv,TelAviv],[ymax,val_excl_NW[argmin(abs(TelAviv-lats))]],'k--',lw=2,zorder=-10)
plt.plot(TelAviv,val_excl_NW[argmin(abs(TelAviv-lats))],'o',color='k',markersize=10,mfc='none',mew=3)
plt.plot([LA,LA],[ymax,val_excl_NW[argmin(abs(LA-lats))]],'k--',lw=2,zorder=-10)
plt.plot(LA,val_excl_NW[argmin(abs(LA-lats))],'o',color='k',markersize=10,mfc='none',mew=3)
plt.plot([Stockholm,Stockholm],[ymax,val_excl_ZW[argmin(abs(Stockholm-lats))]],'k--',lw=2,zorder=-10)
plt.plot(Stockholm,val_excl_ZW[argmin(abs(Stockholm-lats))],'o',color='k',markersize=10,mfc='none',mew=3)
plt.text(lats[10],val_excl_NW[10]+0.04,r'{\bf Zenith-facing}',color='firebrick',rotation=59)
plt.text(lats[500],val_excl_ZN[500]-0.025,r'{\bf West-facing}',color='steelblue',rotation=0,ha='center',va='center')
plt.text(lats[360]-15,val_excl_ZW[360]-0.13,r'{\bf North-facing}',color='seagreen',rotation=-60)
plt.gcf().text(0.0,0.5,'w',color='w',zorder=-1000)
#plt.axhline(1/3,color='k',lw=2.5,alpha=0.4)
plt.xlim([-90,90])
plt.ylim([0,ymax])
plt.title(r'{\bf Experiments sensitive to a plane}',fontsize=45,pad=20)
MySaveFig(fig,'LocationDependence_2D_excl')
# +
ymax = 0.8
fig,ax = MySquarePlot('Latitude [$^\circ$]',r'$\langle \cos^2{\theta}\rangle^{\rm disc.}_{T = 1{\rm day}}$',tfs=29)
plt.plot(lats,val_disc_NW,'-',lw=4,color='firebrick',path_effects=pek)
plt.plot(lats,val_disc_ZN,'-',lw=4,color='steelblue',path_effects=pek,zorder=-1)
plt.plot(lats,val_disc_ZW,'-',lw=4,color='seagreen',path_effects=pek)
plt.text(Sydney+4,ymax-0.01,'Sydney',va='center',rotation_mode='anchor',rotation=-90,fontsize=28)
plt.text(LA+3,ymax*(1-0.08),'LA',va='center',rotation_mode='anchor',rotation=-90,fontsize=28)
plt.text(TelAviv-3,ymax-0.13,'Tel Aviv',va='center',rotation_mode='anchor',rotation=90,fontsize=28)
plt.text(Stockholm+3,ymax-0.01,'Stockholm',va='center',rotation_mode='anchor',rotation=-90,fontsize=22)
plt.plot([Sydney,Sydney],[ymax,val_disc_NW[argmin(abs(Sydney-lats))]],'k--',lw=2,zorder=-10)
plt.plot(Sydney,val_disc_NW[argmin(abs(Sydney-lats))],'o',color='k',markersize=10,mfc='none',mew=3)
plt.plot([TelAviv,TelAviv],[ymax,val_disc_NW[argmin(abs(TelAviv-lats))]],'k--',lw=2,zorder=-10)
plt.plot(TelAviv,val_disc_NW[argmin(abs(TelAviv-lats))],'o',color='k',markersize=10,mfc='none',mew=3)
plt.plot([LA,LA],[ymax,val_disc_NW[argmin(abs(LA-lats))]],'k--',lw=2,zorder=-10)
plt.plot(LA,val_disc_NW[argmin(abs(LA-lats))],'o',color='k',markersize=10,mfc='none',mew=3)
plt.plot([Stockholm,Stockholm],[ymax,val_disc_ZW[argmin(abs(Stockholm-lats))]],'k--',lw=2,zorder=-10)
plt.plot(Stockholm,val_disc_ZW[argmin(abs(Stockholm-lats))],'o',color='k',markersize=10,mfc='none',mew=3)
plt.text(lats[120]*(1+0.07),val_disc_NW[120]*(1+0.2),r'{\bf Zenith-facing}',color='firebrick',rotation=72)
plt.text(lats[500],val_disc_ZN[500]-0.025,r'{\bf West-facing}',color='steelblue',rotation=0,ha='center',va='center')
plt.text(lats[350]-12,val_disc_ZW[350],r'{\bf North-facing}',color='seagreen',rotation=-71)
plt.gcf().text(0.0,0.5,'w',color='w',zorder=-1000)
#plt.axhline(1/3,color='k',lw=2.5,alpha=0.4)
plt.xlim([-90,90])
plt.ylim([0,ymax])
plt.title(r'{\bf Experiments sensitive to a plane}',fontsize=45,pad=20)
MySaveFig(fig,'LocationDependence_2D')
# -
# ##
|
code/LocationDependence.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: python_venv1
# name: python_venv1
# ---
import cv2
import tensorflow as tf
from tensorflow.keras import models
face_cascade = cv2.CascadeClassifier('haarcascade_frontalface_default.xml')
# faces = face_cascade.detectMultiScale(image, scaleFactor=1.2, minNeighbors=5)
|
train_faces.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MongoDB initialisatie
#
# Met de onderstaande opdrachten vullen we de MongoDB database.
# Hiermee krijg je een goed-gedefinieerde beginsituatie.
# Dat kan handig zijn als je de nodige wijzigingen aangebracht hebt.
# + language="bash"
# mongoimport -d demo --drop -c contacts adressen.json
# -
# De onderstaande zoekopdracht toont alle documenten in de collection `adressen` van de database `demo`.
# + language="bash"
# mongosh demo --quiet
#
# db.contacts.find({})
# -
|
mongodb-init.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++11
// language: C++11
// name: xeus-cling-cpp11
// ---
// ## Constant References
// A simple demo that shows the usage of **constant references**
#include <iostream>
const std::string &ams562 = "ams562";
std::cout << "Welcome to " << ams562 << '\n';
// Bind a constant reference to a normal object is fine,
// but you cannot modify the object through the reference.
int a = 1;
const int &a_ref = a;
std::cout << "a_ref=" << a_ref << '\n';
// a_ref = 2; // error!
// instead, do this
a = 2;
std::cout << "a_ref=" << a_ref << '\n';
// Binding a reference to another reference is allowed
// int &a_ref2 = a_ref; // error! why?
const int &a_ref2 = a_ref;
a = 3;
std::cout << "a_ref=" << a_ref << ", a_ref2=" << a_ref2 << '\n';
|
notebooks/2/const_ref.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.display import HTML
css_file = './custom.css'
HTML(open(css_file, "r").read())
# # Logistic Regression
#
# © 2018 <NAME>
# ## 1. Definition
#
# Logistic Regression is used for ***classification***, even though its called ***regression***.
#
# Therefore, it works on ***categorical labels***, namely, 0 and 1 for ***binary classification***.
#
# The ***Logistic Regression*** is a model that makes predictions in the [0, 1] interval, denoting ***probabilities***. Labels of the ***negative class*** are associated with 0, as labels of the ***positive class*** are associated with 1. So, the output is the ***probability of being a sample of the positive class***.
#
# Why is it called ***regression*** then? It actually fits a ***linear regression*** on the features and ***squishes*** the outputs using a ***Logistic / Sigmoid*** function.
#
# $$
# \hat{p} = \sigma(z) = \frac{1}{1 + e^{-z}} = \frac{1}{1 + e^{-(b + w_1x_1 + w_2x_2 + \dots + w_nx_n)}}
# $$
#
# 
# <center>Source: Wikipedia</center>
#
# Since its output is a ***probability***, we need to ***threshold*** it to get the predicted class. The default threshold is 0.5:
#
# $$
# \hat{y} =
# \begin{cases} 0 &\mbox{if } \hat{p} \lt 0.5 \\
# 1 & \mbox{if } \hat{p} \geq 0.5
# \end{cases}
# $$
#
# It is possible to ***change the threshold*** to achieve different goals, as reducing ***false positive*** or ***false negatives*** for instance - the next lesson on evaluation metrics will cover this topic in more depth.
#
# You can observe this behavior on the ***interactive example*** below. The sliders allow you to control the ***bias*** and ***coefficient*** of the simple linear regression that is going to be ***squished*** by the ***sigmoid function***.
from intuitiveml.supervised.classification.LogisticRegression import *
plotLogistic.plot_sigmoid_curve(x=np.linspace(-3, 3, 100))
# ### 1.1 Loss Function
#
# How do we train the model? Differently from a linear regression, the ***Logistic Regression*** uses ***binary cross entropy*** (also called ***log loss***) as its loss function.
#
# What does it mean? It takes the ***log*** of the probability of ***correctly classifying*** a sample as positive or negative and then average it over all samples. For a single instance:
#
# $$
# loss =
# \begin{cases} -log(\hat{p}) &\mbox{if } y = 1 \\
# -log(1-\hat{p}) & \mbox{if } y = 0
# \end{cases}
# $$
#
# And, for all $m$ instances:
#
# $$
# J={-\frac{1}{m}\sum_{i=1}^{m}{y^{(i)} \cdot log(\hat{p}(y^{(i)})) + (1-y^{(i)}) \cdot log(1-\hat{p}(y^{(i)}))}}
# $$
#
# I've written a very thorough explanation of this loss function on Towards Data Science: [Understanding binary cross-entropy / log loss: a visual explanation](https://towardsdatascience.com/understanding-binary-cross-entropy-log-loss-a-visual-explanation-a3ac6025181a)
# ## 2. Experiment
#
# Time to try it yourself!
#
# You have 8 data points, either ***green (positive)*** or ***red (negative)***.
#
# There is only ***one feature***, represented on the horizontal axis. The ***y axis*** is the probability output of your ***Logistic Regression***.
#
# You want to start training your logistic regression, so you need to find both the ***bias*** (intercept) $b$ and the ***single weight*** $w_1$ that minimize the ***log loss***.
#
# The sliders below allow you to change both values, and you can observe the effect they have on the distribution of losses (on the upper right plot), as well as the ***log loss***.
#
# Use the slider to play with different configurations and answer the ***questions*** below.
mylr = plotLogistic(x=(-1, 0), n_samples=8, betas=(2, 1))
vb1 = VBox(build_figure_fit(mylr))
vb1.layout.align_items = 'center'
vb1
# #### Questions
#
# 1. What happens to the probabilities as you increase $w_1$? What about the losses?
#
#
#
# 2. What happens to the probabilities as you increase $b$? What about the losses?
#
#
#
#
# 3. Try to ***minimize*** the log loss.
#
#
#
# ## 3. Scikit-Learn
#
# [Logistic Regression](https://scikit-learn.org/stable/modules/linear_model.html#logistic-regression)
#
# Please check <NAME>'s "Hand-On Machine Learning with Scikit-Learn and Tensorflow" notebook on Linear Models [here](http://nbviewer.jupyter.org/github/ageron/handson-ml/blob/master/04_training_linear_models.ipynb).
# ## 4. More Resources
#
# [InfoGraphic](https://github.com/Avik-Jain/100-Days-Of-ML-Code/blob/master/Info-graphs/Day%204.jpg)
# ## 5. Keras
#
# Just like we did with Linear Regression, we can also build a simple one neuron network to train a ***Logistic Regression***. The model has ***two*** differences:
#
# 1. It has a ***sigmoid activation*** (instead of linear)
# 2. It uses ***binary cross-entropy*** as loss (instead of MSE)
#
# Effectively, it computes:
#
# $$
# z = \sigma(b + w_1x)
# $$
#
# If you compare the ***weights*** of a model trained using Keras versus a model trained using Scikit-Learn, you'll see they are somewhat different.
#
# This is due to the fact that Scikit-Learn uses a different optimizer and uses regularization by default. This is ***not*** the case of the simple neural network we built using Keras.
#
# ```python
# from keras.layers import Dense
# from keras.models import Sequential
# from keras.optimizers import SGD
#
# model = Sequential()
# model.add(Dense(input_dim=1, units=1, activation='sigmoid', kernel_initializer='glorot_uniform'))
# model.compile(loss='binary_crossentropy', optimizer=SGD(lr=0.1))
# model.fit(m.x, m.y, epochs=100)
# ```
import warnings
warnings.filterwarnings("ignore")
from keras.layers import Dense
from keras.models import Sequential
from keras.optimizers import SGD
model = Sequential()
model.add(Dense(input_dim=1, units=1, activation='sigmoid', kernel_initializer='glorot_uniform'))
model.compile(loss='binary_crossentropy', optimizer=SGD(lr=0.1))
model.fit(mylr.x, mylr.y, epochs=100, verbose=False)
model.summary()
# ```python
# model.get_weights()
# ```
print(model.get_weights())
# #### This material is copyright <NAME> and made available under the Creative Commons Attribution (CC-BY) license ([link](https://creativecommons.org/licenses/by/4.0/)).
#
# #### Code is also made available under the MIT License ([link](https://opensource.org/licenses/MIT)).
from IPython.display import HTML
HTML('''<script>
function code_toggle() {
if (code_shown){
$('div.input').hide('500');
$('#toggleButton').val('Show Code')
} else {
$('div.input').show('500');
$('#toggleButton').val('Hide Code')
}
code_shown = !code_shown
}
$( document ).ready(function(){
code_shown=false;
$('div.input').hide()
});
</script>
<form action="javascript:code_toggle()"><input type="submit" id="toggleButton" value="Show Code"></form>''')
|
11. Logistic Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/markbriers/data-science-jupyter/blob/main/week2_notes.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="qNIPLENaVzvu"
# # Python fundamentals 1 (Week 2)
# + [markdown] id="5VntuezuaDbf"
# ## Module: Learning outcomes
#
# * Describe the six stages of a data processing pipeline (using CRISP-DM)
#
# * Demonstrate an understanding of the python programming language through the production of elementary data analysis programme
#
# * Analyse at least three different data sources by applying at least one python data processing library to extract and explore pertinent features
#
# * Be able to design a set of data requirements for a specified business problem
#
# * Describe and apply (using the python programming language) the main approaches to supervised learning for a given classification problem
#
# * Understand the use cases of Big Data technology (in particular Spark)
#
# * Produce a report including appropriate data visualisations covering the analysis of a business problem using a data science based approach
# + [markdown] id="EBoNhH2zabbZ"
# ## Week 2: Learning outcome
#
# * At the end of week 2, you will have a foundational level knowledge of Python. You will be able to write code and text using Markdown.
# + [markdown] id="DTJ-cX2Rat5B"
# ## An introduction to Markdown
# + [markdown] id="E3mGzyvZXOFv"
# Let's start with the text interface, which allows Markdown to be written. Markdown is a lightweight markup language (similar to HTML for webpages) that allows us to write text and embed images and mathematics. This allows for interactive visualisations, allowing for better reproducibility, sharing, and decision making.
# + [markdown] id="rMYj0H7zXlrX"
# Example:
# + [markdown] id="SDmq3Et1Xo3J"
# # Title 1
# + [markdown] id="dG-PDjEvYeMS"
# ## Title 2
# + [markdown] id="1i1esfmwYikA"
# ### Title 3
# + [markdown] id="wE0LDedgYn8k"
# This is a line of text.
# + [markdown] id="x_FUn9ivYpVT"
# * This is a bullet
# * This is a second bullet
# * This is a bullet with _italic_ text
# * This is a bullet with **bold** text
# + [markdown] id="EUXCaIYdYsdx"
# This is an equation written in LaTex:
# \begin{equation}
# f(x) = x^2
# \end{equation}
# + [markdown] id="YyfnmOMlYy0T"
# This is an equation $x^2$ written inline.
# + [markdown] id="4csP65qNZpl9"
# Further details on Markdown and LaTex can be found here:
# Mandatory exercises
# - [ ] Read this reference: https://colab.research.google.com/notebooks/markdown_guide.ipynb
#
# Optional exercises
# - [ ] Use this cheat sheet when writing Markdown for your own work: https://medium.com/ibm-data-science-experience/markdown-for-jupyter-notebooks-cheatsheet-386c05aeebed
# - [ ] Read this reference: https://www.math.ubc.ca/~pwalls/math-python/jupyter/latex/
# + [markdown] id="aEVGIE86b_uk"
# ## Mathematical foundations
# + [markdown] id="Qhg6yw59cE-z"
# ### Notation
#
# * $4\times4\times4 = 4^3 = 64$
# * $x\times x\times x = x^3$ (e.g. when $x=4$, $x^3 = 64$)
# * In the expression above, $x$ is a _variable_
# + colab={"base_uri": "https://localhost:8080/"} id="ab0anNz1cnsw" outputId="26498082-e3e2-41cc-b236-0e7d7a572435"
4 * 4 * 4
# + [markdown] id="c8XS_JlNc0Vx"
# This can be equivalently written as:
# + colab={"base_uri": "https://localhost:8080/"} id="8ecHJeH0c4gS" outputId="022cfbbd-4f97-4b78-d3a3-e0ca7404ad43"
4 ** 3
# + [markdown] id="KfHOUQdYdT9k"
# We can introduce variables, such as $x = 4$
# + id="50b1FzLFdlxU"
x = 4
# + [markdown] id="CZO6IRtYdoOt"
# We can then manipulate variable x, rather than specific values.
# + colab={"base_uri": "https://localhost:8080/"} id="gLKoYU4dd2PD" outputId="6fb5deff-b532-40a5-b82a-5a531036bc4f"
x * x * x
# + id="Rdm2JdZRd7Wo"
y = 2
# + colab={"base_uri": "https://localhost:8080/"} id="-h4Ka42teJ9x" outputId="0b6a9b9c-6fa6-4694-8124-6ff56b4a5759"
x ** y
# + [markdown] id="0QQdZKmReMDr"
# We can then change the values of x and y, to maximise code reuse.
# + [markdown] id="tDfM05RCciUG"
# * We could generalise further and say $f(x) = x^2$, where "$f$ is a function of the variable $x$" (e.g. $f(4) = 16$)
# * A function is a relation between sets that associates to every element of a first set exactly one element of the second set
# * A set is a well defined collection of distinct items. e.g. $\{apple,orange,banana\}$
#
# Let's look at how we do this in Python.
# + [markdown] id="UUTR-fbxiUqY"
# In Python, a function is defined using the def keyword:
# + id="B3CMEGQ-iTWO" colab={"base_uri": "https://localhost:8080/"} outputId="22d40f93-77bf-4c0a-e0b8-31dc82b49edb"
def print_hello():
print("Hello world")
# this is a comment
x = 4
print(x)
# + id="vyjNXOK8yo9h"
def print_hello_user(user):
print("Hello "+user)
# + colab={"base_uri": "https://localhost:8080/"} id="5z7qXxEqiX1b" outputId="cc11a421-6126-469e-bfff-9e9d6ea09016"
print_hello_user("Mark")
# + colab={"base_uri": "https://localhost:8080/"} id="zuDOGyk0xLHU" outputId="b5b27238-8078-4425-892f-df478fcafb1e"
print_hello()
# + [markdown] id="aO1ZICojioed"
# We can pass variable arguments to functions:
# + id="U0_LkJHechDN"
def square(x):
return x ** 2
# + [markdown] id="fP1Cx_KJe-Zd"
# We can use this function several times, in order to reduce the amount of code that we need to write and to maximise code sharing.
# + colab={"base_uri": "https://localhost:8080/"} id="rtZ4wYDafhyi" outputId="4c19797a-098b-4917-ed1a-cc82e86f508b"
x = 3
print(x)
print(x == 4)
# + colab={"base_uri": "https://localhost:8080/"} id="qT3tOWsgfj9h" outputId="1fed6837-08e5-4e32-ea75-3a2a98615fd3"
square(5)
# + [markdown] id="DWqFm9ytixeB"
# We can also define recursive functions (e.g. factorial(5) = 5! = 5 x 4 x 3 x 2 x 1)
# + id="hteT6gC4iyy3"
def factorial(n):
if n == 1:
return n
else:
return n*factorial(n-1)
# + colab={"base_uri": "https://localhost:8080/"} id="stEl9x3ri-68" outputId="f7d7ed01-5910-4fed-ccd9-473449f5e881"
factorial(5)
# + colab={"base_uri": "https://localhost:8080/"} id="sIQ74Fp4i-uA" outputId="adb9e582-125b-4fab-d2dd-6ebd90c8bcc0"
factorial(3)
# + [markdown] id="Z9TZ8s_Sgmow"
# We can perform operations with different Python native datatypes (Boolean, Integer, Float, List, String).
#
# Booleans are either assigned to True or False. Python expects an expression to evaluate to a boolean value. These are called Boolean contexts.
# + colab={"base_uri": "https://localhost:8080/"} id="f3Ri3XZcgsif" outputId="e7d41cd0-52ac-41c8-feca-00bbac451eb1"
x > 10
# + colab={"base_uri": "https://localhost:8080/"} id="8YLz8hnhgx1c" outputId="97065786-fb0b-48d8-95f6-5e2b2917dda3"
x == 2
# + colab={"base_uri": "https://localhost:8080/"} id="jmDh5vCIhBIP" outputId="ba82131e-00ca-47a6-ba61-d95ee66efcba"
if x < 4:
print("x<4")
else:
print("x>=4")
# + id="qdznU3ihhk1r"
x = 10
# + [markdown] id="LFHIFcj7h3R1"
# A float (floating point real number) is defined with a decimal, or explicitly:
# + id="XTupgEkoh8YJ"
pi = 3.14
# + colab={"base_uri": "https://localhost:8080/"} id="aNtY8EhDiBvP" outputId="6bb8530a-7b8d-41e5-8e16-69fd0e6c60d2"
print(pi)
# + colab={"base_uri": "https://localhost:8080/"} id="SWfLQu2biGV_" outputId="fb87d252-e9d9-45e7-ea9e-c9f4598da8b1"
float(2)
# + [markdown] id="CF<KEY>"
# ## Libraries
# + [markdown] id="PZrXqu6SjeDl"
# The power of Python (with respect to data science) is its extensibility. We can import libraries and use functions and data types from such libraries. NumPy (https://numpy.org/) is the basis of many data science libraries. We import it as follows:
# + id="OfpgjGVzfpi9"
import numpy as np
# + [markdown] id="sSjenVGpjo7U"
# We can use numpy to define _vectors_ and _matrices_, and to perform mathematical operations:
# + id="VVzcuFj6jgMo"
# Create a vector as a row
vector_row = np.array([1, 2, 3])
# + colab={"base_uri": "https://localhost:8080/"} id="WEWgseyyj77q" outputId="79c9aaf8-2c2b-444e-ab91-55852f8e2aa2"
print(vector_row)
# + id="6woweiySj9Tn"
# Create a vector as a column
vector_column = np.array([[1],
[2],
[3]])
# + colab={"base_uri": "https://localhost:8080/"} id="FPHLZhDG3m1Q" outputId="2d2c0416-270f-4370-ebb2-290bc1ca24c7"
print(vector_column)
# + [markdown] id="vkJp7RMJkCka"
# Let's call the _linspace_ function, in order to produce a list of values that are equally spaced:
# + colab={"base_uri": "https://localhost:8080/"} id="i9JZy1Mpj_iF" outputId="c553bd90-b28e-4a15-a190-89ed24b3fedf"
# Build array/vector:
x = np.linspace(-np.pi, np.pi, 10)
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="l7nVfI10kI2t" outputId="4e2a0e14-1ea9-48ff-86d8-a59db19c2eb4"
print(x[0]) # first element
print(x[2]) # third element
print(x[-1]) # last element
print(x[-2]) # second to last element
# + colab={"base_uri": "https://localhost:8080/"} id="T6sb5zw_kUnY" outputId="2e44873c-2f11-4702-dd20-224ad7830845"
print(x[1:4]) # second to fourth element. Element 5 is not included
print(x[0:-1:2]) # every other element
print(x[:]) # print the whole vector
print(x[-1:0:-1]) # reverse the vector!
# + [markdown] id="89z9r27pk9jK"
# Consider if we want the part of the vector where x > 2:
# + colab={"base_uri": "https://localhost:8080/"} id="Z7nmO6GCklm-" outputId="010fd59d-4eb7-40ff-d616-43b322f8060f"
print(x > 2)
y = x[x > 2]
print(y)
y[0]
# + [markdown] id="fo29SrBYmhwU"
# In the above, we have been programmatically manipulating arrays. A mathematical representation of an array is known as a vector.
# * (Imprecisely) a *vector* is an ordered set of elements, $\vec{x}=(x_{1}~x_{2}~\ldots~x_{n})\in\mathbb{R}^n$
# + [markdown] id="7ob7qB3ImxfX"
# * Let:
# \begin{eqnarray}
# \vec{a} = \left[\begin{matrix}
# a_{11} \\
# a_{21} \\
# a_{31} \\
# \end{matrix}\right], & ~~~ &
# \vec{b} = \left[\begin{matrix}
# b_{11} \\
# b_{21} \\
# b_{31} \\
# \end{matrix}\right]
# \end{eqnarray}
# + [markdown] id="iZEBaFWLnY-R"
# * Then:
# \begin{equation}
# \vec{a}+\vec{b} = \left[\begin{matrix}
# a_{11}+b_{11} \\
# a_{21}+b_{21} \\
# a_{31}+b_{31} \\
# \end{matrix}\right]
# \end{equation}
# + [markdown] id="9VNHL-GPncAD"
# We will return to this when we delve deeper into modelling, later in the course.
# + [markdown] id="PrXph2zinqRC"
# ## Matrix
#
# * A matrix is a two-dimensional representation of numbers (a grid)
# * In data science, we will use linear representations to map our input to our output space, to store neural network weights, while features are stored as vector inputs
# * This allows us to have compact mathematical representations of the models, and to exploit computational resources (such as GPUs) in order to perform extremely fast processing
# * Example:
# \begin{equation}
# A =
# \left[\begin{matrix}
# 11 & 14 & 18 & 9 \\
# 32 & 14 & 24 & 17 \\
# 10 & 7 & 6 & 28 \\
# \end{matrix}\right]
# \end{equation}
# * We can define operations between matrices, or between matrices and vectors
# + id="ekNNa8zanbu7"
# Create a matrix
matrix = np.array([[11, 14, 18, 9],
[32, 14, 24, 17],
[10, 7, 6, 28]])
# + colab={"base_uri": "https://localhost:8080/"} id="SMbD1KClk_ji" outputId="bdde018d-3c4a-4f41-f6a8-ce8aea6178c2"
matrix[0,2]
# + colab={"base_uri": "https://localhost:8080/"} id="6HabVpRIn93K" outputId="b0268ef5-e2a7-488d-cd63-e0785abf44a9"
matrix.size
# + colab={"base_uri": "https://localhost:8080/"} id="Ccp-du1woAOi" outputId="daeb62a6-bf43-4f6d-d7c1-f0f0f9fedf5e"
matrix.shape
# + colab={"base_uri": "https://localhost:8080/"} id="ngV2eOoJoCv3" outputId="9d2786e0-2b21-4bfb-c900-5849d0bec6a9"
matrix.T
# + [markdown] id="2XiQAFj8oKUM"
# We can perform algebraic operations on matrices:
# + colab={"base_uri": "https://localhost:8080/"} id="usxBDzbsoEoA" outputId="d21364d7-984d-4b3d-aedc-98291d606ae2"
# Create matrix
matrix_a = np.array([[1, 1, 1],
[1, 1, 1],
[1, 1, 2]])
# Create matrix
matrix_b = np.array([[1, 3, 1],
[1, 3, 1],
[1, 3, 8]])
# Add two matrices
np.add(matrix_a, matrix_b)
# + [markdown] id="hwa3oXsxoX8R"
# Or equivalently:
# + colab={"base_uri": "https://localhost:8080/"} id="Ah81qW5BoQa_" outputId="ee8b95a7-84a5-40fe-8d52-864bd36d9098"
matrix_a + matrix_b
# + colab={"base_uri": "https://localhost:8080/"} id="U-UJIiYyoZzF" outputId="c8892695-af36-4c3a-9383-6d890b0ba06f"
matrix_a - matrix_b
# + [markdown] id="J5IZwPwdoleT"
# ## Exercises
# + [markdown] id="sXzyTw-oon9Y"
# Mandatory exercises:
# - [ ] Introduce a function that computes $x+2$ for any value of $x$. Call the function addTwo.
# - [ ] Replicate all of the code in this notebook.
#
#
# Advanced (optional) exercises (for students with existing Python knowledge):
# - [ ] By reading the NumPy documentation, create a 3-dimensional array (a _tensor_)
# - [ ] Create your own simple Python class for a Matrix, that is, an object based representation of a two-dimensional array
|
week2_notes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: exercise
# language: python
# name: exercise
# ---
# +
from Exercise import Exercise, MarkdownBlock
from config import URL, TOKEN
import json
import numpy as np
import sympy as sp
Exercise.URL = URL
Exercise.TOKEN = TOKEN
# +
from sympy import Rational, Symbol, latex, UnevaluatedExpr
u = lambda x : UnevaluatedExpr(x)
# Helper functions
def explain_add(a, b):
assert(np.shape(a) == np.shape(b))
rows, columns = np.shape(a)
return sp.Matrix([[Symbol(f"({latex(u(a[i,j]))} + {latex(u(b[i,j]))})") for j in range(columns)] for i in range(rows)])
def symbolic_matrix(character, rows, columns):
return sp.Matrix([[Symbol(f"{{{character}}}_{{{i+1}, {j+1}}}") for j in range(columns)] for i in range(rows)])
def explain_multiply(a, b):
# #rows in b == #columns in a
assert(np.shape(a)[1] == np.shape(b)[0])
rows = np.shape(a)[0]
columns = np.shape(b)[1]
result = np.empty(shape=(rows, columns), dtype=object)
for i in range(rows):
row = a[i,:]
for j in range(columns):
column = b[:,j]
zipped = zip(row, column)
mapped = list(map(lambda t: f"{latex(u(t[0]))} \cdot {latex(u(t[1]))}", zipped))
s = Symbol("")
result[i, j] = Symbol(" + ".join(mapped), evaluate=False)
return sp.Matrix(result)
# TODO: fixme for >= 3 dim matrices
# TODO: print intermediate steps at each return
def explain_det(a):
# square matrix
assert(np.shape(a)[0] == np.shape(a)[1])
if np.shape(a)[0] == 1:
return a[0,0]
if np.shape(a)[0] == 2:
return f"{a[0,0]} \cdot {a[1,1]} - {a[0,1]} \cdot {a[0,1]}"
else:
return f"{a[0,0]} \cdot {explain_det(a[1:,1:])} - {a[0,1]} \cdot {[[a[1,0], a[1,2]], [a[2,0], a[2,2]]]} + {a[0,2]} \cdot {[[a[1,0], a[1,2]], [a[1,1], a[2,1]]]}"
# A = symbolic_matrix("a", 3, 3)
# e = Exercise(f"${explain_det(A)}$")
# e.display()
# +
def generator():
length = np.random.randint(1, 7)
v1 = sp.randMatrix(r=length, c=1, min=0, max=10)
v2 = sp.randMatrix(r=length, c=1, min=0, max=10)
s = "What is $@v1 + @v2$?"
params = {}
params["v1"] = v1
params["v2"] = v2
e = Exercise(MarkdownBlock(s, params))
params["v3"] = explain_add(v1,v2)
params["v4"] = v1 + v2
s1 = "Yes, $@v1 + @v2 = @v3 = @v4$!"
e.add_answer(v1 + v2, True, MarkdownBlock(s1, params))
a = symbolic_matrix("a", length, 1)
b = symbolic_matrix("b", length, 1)
ab = explain_add(a, b)
default_feedback = "Remember the definition of matrix addition: $@a + @b = @ab$"
e.add_default_feedback(MarkdownBlock(default_feedback, dict(a=a, b=b, ab=ab)))
return e
Exercise.write_multiple(generator, 100, "vector_add_2")
# +
s = "What is $@a @b$?"
rows = np.random.randint(1, 4)
columns = np.random.randint(1, 4)
params = {}
params["a"] = sp.randMatrix(r=rows, c=columns, min=0, max=10)
params["b"] = sp.randMatrix(r=columns, c=rows+2, min=0, max=10)
ans = params["a"] * params["b"]
e = Exercise(MarkdownBlock(s, params))
e.add_answer(ans, True, "That's right!")
e.play()
# +
def generator():
s = "What is $@a @b$?"
rows = np.random.randint(1, 4)
columns = np.random.randint(1, 4)
params = {}
params["a"] = sp.randMatrix(r=rows, c=columns, min=0, max=10)
params["b"] = sp.randMatrix(r=columns, c=rows+2, min=0, max=10)
ans = params["a"] * params["b"]
e = Exercise(MarkdownBlock(s, params))
e.add_answer(ans, True, "That's right!")
return e
Exercise.write_multiple(generator, 10, "mat_multiply")
# +
def generator():
s = "What is $@a \cdot @b$?"
rows = np.random.randint(1, 4)
columns = np.random.randint(1, 4)
params = {}
params["a"] = sp.Matrix(np.random.randint(5, size=rows*columns).reshape((rows,columns)))
params["b"] = sp.Matrix(np.random.randint(5, size=(2+rows)*columns).reshape((columns,rows+2)))
e = Exercise(MarkdownBlock(s, params))
ans = params["a"] * params["b"]
e.add_answer(params["a"] * params["b"], True, "That's right!")
params = {}
params["x"] = symbolic_matrix("a", rows, columns)
params["y"] = symbolic_matrix("b", columns, rows+2)
params["z"] = explain_multiply(params["x"], params["y"])
f = """Remember the definition of matrix multiplication: \n $@x \cdot @y = @z$"""
e.add_default_feedback(MarkdownBlock(f, params))
return e
Exercise.write_multiple(generator, 10, "mat_multiply")
# +
def explain_multiply(a, b):
# #rows in b == #columns in a
assert(np.shape(a)[1] == np.shape(b)[0])
rows = np.shape(a)[0]
columns = np.shape(b)[1]
result = np.empty(shape=(rows, columns), dtype=object)
for i in range(rows):
row = a[i,:]
for j in range(columns):
column = b[:,j]
zipped = zip(row, column)
mapped = list(map(lambda t: f"{latex(u(t[0]))} \cdot {latex(u(t[1]))}", zipped))
s = Symbol("")
result[i, j] = Symbol(" + ".join(mapped), evaluate=False)
return sp.Matrix(result)
# explain_multiply(np.arange(4).reshape((2,2)), np.arange(4).reshape((2,2)))
# -
v = symbolic_matrix("a", 3, 1)
display(3*u(v))
display(3*v)
# # TODO: meaningfull, contextualized exercises for each of these:
# - scalar def.
# - vector def.
# - matrix def.
# - vector indexing
# - matrix indexing
# - total values in a matrix
# - vector dimensions
# - matrix dimensions
# - special matrices
# - transpose
# - operations (+, *), linear combinations
# - dot product, cross product, norm
# - cosine similarity
# ## Matrix Transpose
# +
m = "Determine $@a^\intercal$?"
a = np.arange(6).reshape((2, 3))
params = {}
params["a"] = sp.Matrix(a)
e = Exercise(MarkdownBlock(m, params))
e.display()
e.add_answer(params["a"].T, True, "Correct!")
e.write()
e.publish()
# -
# ## Matrix Inverse
# +
m = "Determine $@a^{-1}$?"
a = np.arange(4).reshape((2, 2))
params = {}
params["a"] = sp.Matrix(a)
e = Exercise(MarkdownBlock(m, params))
e.display()
e.add_answer(params["a"].inv(), True, "Correct!")
e.write()
e.publish()
# +
m = "Given $A = @a$, what is $det(A)$?"
a = np.arange(4).reshape((2, 2))
params = {}
params["a"] = sp.Matrix(a)
e = Exercise(MarkdownBlock(m, params))
e.display()
e.add_answer(params["a"].det(), True, "Correct!")
e.write()
e.publish()
# -
|
step-by-step-explain.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
"""
Important Stuff
"""
# Webcam index, this should be 0 for most laptops
DEVICE = 0
# The object detection network weights file
WEIGHTS='weights/squirrel_detector.hdf5'
"""
User Interface Stuff (You can ignore this)
"""
import IPython
import ipywidgets as widgets
from IPython.display import display
ipython = IPython.get_ipython()
style = {'description_width': 'initial'}
w_image = widgets.Image(width=224, height=224, format='png',
layout=widgets.Layout(width='100%'))
w_heatmap = widgets.Checkbox(
value=True,
description='Overlay heatmap on image',
style=style
)
w_bbox = widgets.Checkbox(
value=True,
description='Show Bounding Boxes (BBoxes)',
style=style
)
w_bbox_thresh = widgets.FloatSlider(min=0, max=1, value=0.99, step=0.01,
description='BBox Confidence Threshold',
layout=widgets.Layout(width='100%'),
style=style)
w_merge = widgets.Checkbox(
value=True,
description='Merge close BBoxes',
style=style
)
w_merge_thresh = widgets.FloatSlider(min=0, max=1, value=0.75, step=0.01,
description='BBox Merge Threshold',
layout=widgets.Layout(width='100%'),
style=style)
vbox = widgets.VBox([w_image, w_heatmap, w_bbox, w_bbox_thresh, w_merge, w_merge_thresh])
display(vbox)
# + pycharm={"name": "#%%\n"}
# Load the model
from model import MobileDetectNetModel
import numpy as np
import cv2
keras_model = MobileDetectNetModel.complete_model()
keras_model.load_weights(WEIGHTS, by_name=True)
tf_engine = keras_model.tf_engine()
# + pycharm={"name": "#%%\n"}
cap = cv2.VideoCapture(0)
try:
while True:
# Have to call this to get update values from sliders / dropdowns
ipython.kernel.do_one_iteration()
# Read the frame from the camera
ret, frame = cap.read()
img_original = frame
img_draw = img_original.copy()
img_resize = cv2.resize(img_original, (224, 224))
scale_width = img_original.shape[1] / 224
scale_height = img_original.shape[0] / 224
"""
The network expects the image to be scaled between -1 and 1,
but most images are scaled between 0 and 255 normally.
We divide by 127.5 to scale between 0 and 2, and subtract one to
be between -1 and 1
"""
img_input = (img_resize / 127.5) - 1
"""
The neural network expects a "batch" of images as an input
This converts our single image with a shape of (224, 224, 3) to (1, 224, 224, 3)
The 1 at the beginning is called the batch dimension
"""
batch = np.expand_dims(img_input, axis=0)
# Do the actual inference
bboxes, classes = tf_engine.infer(batch)
rectangles = []
for y in range(0, 7):
for x in range(0, 7):
if classes[0, y, x, 0] >= w_bbox_thresh.value:
rect = [
int(bboxes[0, int(y), int(x), 0] * 224),
int(bboxes[0, int(y), int(x), 1] * 224),
int(bboxes[0, int(y), int(x), 2] * 224),
int(bboxes[0, int(y), int(x), 3] * 224)]
rectangles.append(rect)
if w_heatmap.value:
cls_img = cv2.resize((classes[0]*255).astype(np.uint8), (img_draw.shape[1], img_draw.shape[0]), interpolation=cv2.INTER_AREA)
cls_cmap = cv2.applyColorMap(cls_img, cv2.COLORMAP_JET)
cls_add = (img_draw).astype(np.float32) + (np.expand_dims(cls_img, axis=-1)*cls_cmap).astype(np.float32)
img_draw = (255*(cls_add / np.max(cls_add))).astype(np.uint8)
if w_merge.value:
rectangles, merges = cv2.groupRectangles(rectangles, 1, eps=w_bbox_thresh.value)
if w_bbox.value:
for rect in rectangles:
cv2.rectangle(img_draw,
(int(rect[0]*scale_width), int(rect[1]*scale_height)),
(int(rect[2]*scale_width), int(rect[3]*scale_height)),
(0, 255, 0), 5)
# Visualization Code
result, img_png = cv2.imencode('.png', img_draw)
w_image.value = img_png.tobytes()
except KeyboardInterrupt:
pass
finally:
cap.release()
# -
|
realtime.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #List
L =[1,2,3,4,5,6,121,43]
print(L)
# .Indexing of list
L[0],L[3],L[-1],L[-3]
# * For the length of list
len(L)
# * To add the element from last
L.append(22)
L
# To find sum of the elements
sum(L)
# To find maximum and minimum of the list
max(L),min(L)
# # ? To create a list of heterogeneous element types
L =[1,2,3,4,5.0,"Banana",True,False]
type(L[2]),type(L[5])
type(L[4]) ,type(L[-1])
# # To create a list of list
L =[[1,2,3],[4,5,6],[7,8,9]]
# To find list inside a list
L[0][2]
# .To add two lists in case of set
L1 = [1,2,3] ; L2 = [2,4,6]
L1+L2, set (L1+L2)
# To add element fromend of the list
L = [1,2,3,4,9,0,6]
L.append(45)
L
# To insert (29) on first index
L = [1,2,3,4,9,0,6]
L.insert(1,29)
L
# To remove an element (2)
L = [1,2,3,4,9,0,6]
L.remove(2)
L
# To remove the element from specific index
L = [1,2,3,4,9,0,6]
L.pop(-2)
L
# To sort the list
L = [1,2,3,4,9,0,6]
L.sort()
L
# To reverse the list
L = [1,2,3,4,9,0,6]
L.reverse()
L
# List comprehension
L = [x for x in range(50)]
print(L)
L = [x for x in range(50) if x%2==0]
print(L)
import random as rn
rn.randint(0,50)
import random as rn
R = [rn.randint(0,50) for k in range (100)]
print(R)
R = [rn.choice(['A' ,'T','G','C']) for i in range (200)]
print(R)
# # Tuples
# To define a tuple from scratch
t =(2,3,4,6)
# Find type
type(t)
t[2]
L =[(1,2),(2,3),(3,4)]
L[0][0]
# Create a list of tuple
L =[(1,2),("a","b"),(True,False)]
L
# # Dictionary
# ## Initiate dictionary
D = dict()
DD = {}
# Create a dictionary from scratch
D ={"fruits": 'banana',
"Vegetable": 'radish',
"rice": '3.00',
"water": '100'}
# What are keys?
D.keys()
D.values()
|
List tuple dictionary.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import mlflow
import mlflow.sklearn
from gensim.utils import simple_preprocess
from sklearn.model_selection import train_test_split
from gensim.models.doc2vec import Doc2Vec, TaggedDocument
import gensim
from gensim import corpora
import nltk.stem
nltk.download('rslp')
from gensim.utils import simple_preprocess
from gensim.parsing.preprocessing import remove_stopwords
import nltk
nltk.download('stopwords')
mlflow.set_tracking_uri("http://mlflow_server:5000")
from sklearn.model_selection import train_test_split
mlflow.set_experiment("Doc2Vec-Decision-Tree")
from sklearn.tree import DecisionTreeClassifier
#importing dataset
df_train = pd.read_csv(str( 'sample_products.csv'),sep=',')
df_test = pd.read_csv(str( 'test_products.csv'), sep=',')
mlflow.sklearn.autolog(log_models=True,log_model_signatures=True)
# -
# 1o passo Removal of Stop Words
# 2o passo Tokenization
# 3o passo Stemming
#
# concatening title and tags
df_copy = df_train.copy()
df_copy["text"] = df_copy["concatenated_tags"] + " " + df_copy["query"]+ " " + df_copy["title"]
df_copy = df_copy[df_copy["concatenated_tags"].notnull()]
# +
# tokenization
# Tokenize the text column to get the new column 'tokenized_text'
df_copy['tokenized_text'] = [simple_preprocess(line, deacc=True) for line in df_copy['text']]
print(df_copy['tokenized_text'].head(10))
# +
# Removal of Stop Words
stopwords = nltk.corpus.stopwords.words('portuguese')
# Exclude stopwords with Python's list comprehension and pandas.DataFrame.apply.
df_copy['tokens'] = df_copy['text'].apply(lambda x: ' '.join([word for word in x.split() if word not in (stopwords)]))
print(df_copy['tokens'].head(10))
# tokenization
# Tokenize the text column to get the new column 'tokenized_text'
df_copy['tokenized_text'] = [simple_preprocess(line, deacc=True) for line in df_copy['tokens']]
print(df_copy['tokenized_text'].head(10))
# +
# Stemming
stemmer = nltk.stem.RSLPStemmer()
# Get the stemmed_tokens
df_copy['stemmed_tokens'] = [[stemmer.stem(word) for word in tokens] for tokens in df_copy['tokenized_text']]
df_copy['stemmed_tokens'].head(10)
# +
# building dictionaries
# Build the dictionary
mydict = corpora.Dictionary(df_copy['stemmed_tokens'])
print("Total unique words:")
print(len(mydict.token2id))
print("\nSample data from dictionary:")
i = 0
# Print top 4 (word, id) tuples
for key in mydict.token2id.keys():
print("Word: {} - ID: {} ".format(key, mydict.token2id[key]))
if i == 3:
break
i += 1
# +
#Generating Bow Vectors
vocab_len = len(mydict)
print("Example of how the BOW words")
arr = []
for line in df_copy['stemmed_tokens']:
print("Doc2Bow Line:")
print(mydict.doc2bow(line))
for word in line:
arr.append(mydict.token2id[word])
print("Actual line:")
print(line)
print("(Word, count) Tuples:")
print([(mydict[id], count) for id, count in mydict.doc2bow(line) ])
print("Sparse bow vector for the line")
print(gensim.matutils.corpus2csc([mydict.doc2bow(line)],num_terms=vocab_len).toarray()[:,0])
break
print("Sorted word id list")
print(sorted(arr))
df_copy = df_copy.fillna(0)
print(df_copy.info())
#Create column for each category
df_one = pd.get_dummies(df_copy.category)
print(df_one.head())
df_copy = pd.concat([df_copy, df_one], axis=1)
# +
# Train Test Split Function
top_data_df_small = df_copy
def split_train_test(top_data_df_small,category, test_size=0.3, shuffle_state=True):
X_train, X_test, Y_train, Y_test = train_test_split(top_data_df_small[['product_id', 'seller_id','search_page','position', 'creation_date', 'price','weight','express_delivery','minimum_quantity','view_counts','order_counts', 'stemmed_tokens']],
top_data_df_small[category],
shuffle=shuffle_state,
test_size=test_size,
random_state=15)
print("Value counts for Train set")
print(Y_train.value_counts())
print("Value counts for Test set")
print(Y_test.value_counts())
print(type(X_train))
print(type(Y_train))
X_train = X_train.reset_index()
X_test = X_test.reset_index()
Y_train = Y_train.to_frame()
Y_train = Y_train.reset_index()
Y_test = Y_test.to_frame()
Y_test = Y_test.reset_index()
print(X_train.head())
return X_train, X_test, Y_train, Y_test
# Call the train_test_split
X_train, X_test, Y_train, Y_test = split_train_test(top_data_df_small,category='category')
# +
# TaggedDocuments are tuple of stemmed_tokens and class lable, example is printed (scroll to the right to see label)
documents = [TaggedDocument(doc, [i]) for i, doc in enumerate(top_data_df_small['stemmed_tokens'])]
print(documents[1])
# Initialize the model
doc2vec_model = Doc2Vec(documents, vector_size=vocab_len, window=3, min_count=1, workers=4)
# Sample vector for the stemmed tokens
vector = doc2vec_model.infer_vector(top_data_df_small['stemmed_tokens'][0])
# Printing sample vector
print(len(vector))
print("Top 10 values in Doc2Vec inferred vector:")
print(vector[:10])
# -
# Store the vectors for train data in
doc2vec_filename = 'train_review_doc2vec.csv'
with open(doc2vec_filename, 'w+') as doc2vec_file:
for index, row in X_train.iterrows():
model_vector = doc2vec_model.infer_vector(row['stemmed_tokens'])
if index == 0:
header = ",".join(str(ele) for ele in range(vocab_len))
doc2vec_file.write(header)
doc2vec_file.write("\n")
line1 = ",".join( [str(vector_element) for vector_element in model_vector] )
doc2vec_file.write(line1)
doc2vec_file.write('\n')
# +
# Load from the filename
doc2vec_df = pd.read_csv(doc2vec_filename)
clf_decision_doc2vec = DecisionTreeClassifier()
categories = df_copy['category'].unique().tolist()
mydict.save_as_text("my_dict_Doc2Vec.txt", sort_by_word=True)
# Fit the models
for category in categories:
X_train, X_test, Y_train, Y_test = split_train_test(top_data_df_small,category=category)
with mlflow.start_run(run_name='Doc2Vec_categorizer_'+category) as run:
clf_decision_doc2vec.fit(doc2vec_df, Y_train[category])
#print("Logged data and model in run {}".format(run.info.run_id))
mlflow.sklearn.log_model(
sk_model=clf_decision_doc2vec,
artifact_path="sklearn-model",
registered_model_name="Doc2Vec-DecisionTreeClass-"+category
)
mlflow.log_artifact("my_dict_Doc2Vec.txt")
|
training/notebook_Doc2vec.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://apssdc.in"><img src="https://camo.githubusercontent.com/e7501c5948d48f88dad8ab2ab6bd448e1cfd6c79/68747470733a2f2f64726976652e676f6f676c652e636f6d2f75633f6578706f72743d646f776e6c6f61642669643d3135414b51365f2d42697857344b366d4c36525070684635454b58715946327a6a" width="900" align="center"></a>
#
# <h1><center>Python Basics</center></h1>
# ## Today Objectives
#
#
# - Ouput & Input
# - Variables in Python
# - Type Conversions in Python
# - Operators in Python
# - Conditional Statements
print("Hello World")
print("Todays date is",29)
print("Todays date is",29, sep = '-')
print(1,2,3,4,5,6, sep = '\t')
print(1,2,3,4,5,6, sep = 'abc')
print("Hello World")
print("Hello World")
print("Hello World3")
print("Hello World", end = '\t')
print("Hello World")
print("Hello World3")
# ### Variables
#
# It is the named memory location which holds some value
#
#
# ### Properties of declaring a variable
#
# - It can contain AZaz09_
# - It shouldn't start with a number
# - No special characters should included in variable name except `_`
# - Keywords are not allowed
# - Case Sensitive
a = 10
A = 5
a = 55
_a = 2256
# 1a = 66
# a-b = 23+66
_ = 625
a9 = 'Python'
a9_ = 'APSSDC'
# ### Data Types in Python
#
#
# - Primary Data Types -> int, float, complex, string, boolean
# - Secondary Data Types -> Containers -> List, Tuple, Dictoinary, set
a = 55
b = 55.66
c = 5 + 6j
d = True
e = 'a+5/.,;'
f = ['python', 65, 5+6j, 55.665]
g = ('python', 65, 5+6j, 55.665)
h = {'Name': 'Python'}
i = {1,2,3,4}
print(type(a))
# +
a = 55.66
print(type(a))
# -
print(type(c), type(i))
a = input()
abc = 556.6
print(type(a))
b = input()
print(b, type(b))
# ### Type Conversions
# +
a = int(input())
print(a, type(a))
# +
a = int(input())
print(a, type(a))
# +
a = float(input())
print(a, type(a))
# -
# ### Number System
#
# - Decimal -> 10 -> 0-9
# - Binary -> 2 -> 0,1
# - Octal -> 8 -> 0-7
# - Hexadecimal -> 16 -> 0-F
# +
a = 55
print(bin(a), oct(a), hex(a))
# -
print(int('110111', 2), int('67', 8), int('37', 16))
print(ord('a'))
print(chr(97))
# ### Operators in Python
#
#
# - Arthematic Operator -> +, -, *, /, %, //, **
# - Logical -> and, or, not
# - Comparision/relational -> <, >, <=, >=, ==, !=
# - Bitwise Operator -> &(and), |(or), ~(Not), ^(XOR), >>(right shift), <<(left shift)
# - Assignment Operator -> +=, -=, /=, %=, //=, **=
# - Identity operators -> is, not is
# - Membership operators -> in, not in
# +
a = 4
b = 10
print(a+b, a-b, a*b, a/b, a % b, a//b, a ** 2, sep = '->')
# -
# ### and (inp1 * inp2)
#
# |inp1| inp2|inp1 and inp2|
# |----|----|----|
# |T|T|T|
# |T|F|F|
# |F|T|F|
# |F|F|F|
#
# ### or (inp1 + inp2)
#
# |inp1| inp2|inp1 or inp2|
# |----|----|----|
# |T|T|T|
# |T|F|T|
# |F|T|T|
# |F|F|F|
#
# ### not
#
# |inp1|not inp1 |
# |----|----|
# |F|T|
# |T|F|
#
# +
a = 1
b = 0
c = '0'
print(a and b, a or b, not a, a and c, type(a and c), not c, sep = '->')
# -
print(c and a, a or c)
# +
a = 5
b = 6
c = 0
print(a and b)
# -
print(a and c, c and a)
# +
a = 5
b = 6
print(a < b, a>b, a == b)
# -
# ### Bitwise Operators
#
#
# &, |, ~, ^
#
# ```
# a = 4
# b = 10
#
#
#
# 0000 -> 0 -> and
# 1110 -> 14 -> or
#
# ~a
# 4 -> 0100
# 0001
# -0101 -> *5
#
# ```
# +
a = 4
b = 10
print(a & b, a | b, ~a)
# -
# ###
#
# XOR -> A'B + AB'
#
# |inp1| inp2|inp1 and inp2|
# |----|----|----|
# |T|T|F|
# |T|F|T|
# |F|T|T|
# |F|F|F|
#
# ```
# a = 4 -> 0100
# b = 10 -> 1010
# _____
# 1110 -> 14
# ```
print(a ^ b)
# ### Left Shift and right shift
#
#
#
# a = 4 -> 0100
#
#
# a << 2 -> 010000 -> 16
#
# a << 2 -> 0001 -> 1
# +
a = 4
print( a << 2, a >> 2)
# +
a = 5
a = a + 10
print(a)
a += 10
print(a)
a -= 5
print(a)
a **= 2
print(a)
# -
# ### Membership Operators
# +
a = 'Python'
b = 'Py'
c = 'py'
print(a in b, a in c, a not in c)
# -
# ### Identity Operators
a = 5
b = 5.5
c = 5
d = [1,2,3]
e = [1,2,3]
f = d
print(f)
print(a == c, d == e, d == f)
print(id(a))
print(id(c))
print(id(d))
print(id(e))
print(id(f))
# +
d.append('Python')
print(d)
print(f)
# -
print(d,e,f)
print(d is e, d is f, e is f, d is not e)
|
Notebooks/Day06_Python_Basics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# numpy is great for doing vector arithmetic.
# numpy stands for numeric pyhton.
# numpy array can not contain elements with different data types.
# If you try to build such a list, some of the elements' types are changed to end up with a homogeneous list. This is known as type coercion.
#
import numpy as np
x = [range(100)]
print (type(x))
print (x)
np_x = np.array(x)
print(np_x)
np_greater_than_25 = np_x > 25
print (np_greater_than_25)
np_2d_array = np.array([[1,2,3,4,5],
[6,7,8,9,10]])
np_2d_array
np_2d_array.shape
np_3d_array = np.array([[[1,2,3,4,5],
[6,7,8,9,10],
[11,12,13,14,15]]])
np_3d_array.shape
np_3d_array
np_3d_array[0][0]
np_3d_array[0][0][0]
np_3d_array[0][1]
|
numpy_practice.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf2.3
# language: python
# name: tf2.3
# ---
# # Solution8:實作Efficientnet網路架構
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/taipeitechmmslab/MMSLAB-TF2/blob/master/Exercise/Solution8.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/taipeitechmmslab/MMSLAB-TF2/blob/master/Exercise/Solution8.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# ### Import必要套件
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
# ---
# ## Keras Applications
# ### 創建EfficientNetB0網路架構
# - 輸入大小(預設):(224, 224, 3)
# - 權重(預設):`imagenet`
# - 輸出類別(預設):1000個類別
#
# keras.applications API: https://www.tensorflow.org/api_docs/python/tf/keras/applications
#
# ※注意: EfficientNet支援`TensorFlow 2.3+`
model = tf.keras.applications.EfficientNetB0(include_top=True, weights='imagenet')
# 透過`model.summary`可以察看網路模型的每一層資訊:
model.summary()
# 將網路模型儲存到TensorBoard上:
model_tb = tf.keras.callbacks.TensorBoard(log_dir='lab8-logs-efficientnetB0-keras')
model_tb.set_model(model)
# ### 資料前處理和輸出解碼
# Import資料預處理和輸出解碼的函式:
from tensorflow.keras.applications.efficientnet import preprocess_input
from tensorflow.keras.applications.efficientnet import decode_predictions
# ### 預測輸出結果
# 創建影像讀取的函式:讀取影像,並將影像大小縮放大224x224x3的尺寸。
def read_img(img_path, resize=(224,224)):
img_string = tf.io.read_file(img_path) # 讀取檔案
img_decode = tf.image.decode_image(img_string) # 將檔案以影像格式來解碼
img_decode = tf.image.resize(img_decode, resize) # 將影像resize到網路輸入大小
# 將影像格式增加到4維(batch, height, width, channels),模型預測要求格式
img_decode = tf.expand_dims(img_decode, axis=0)
return img_decode
# 從資料夾中讀取一張影像(elephant.jpg)作為測試:
img_path = '../image/elephant.jpg'
img = read_img(img_path) # 透過剛創建的函式讀取影像
plt.imshow(tf.cast(img, tf.uint8)[0]) # 透過matplotlib顯示圖片需將影像轉為Integers
# 預測結果:
img = preprocess_input(img) # 影像前處理
preds = model.predict(img) # 預測圖片
print("Predicted:", decode_predictions(preds, top=3)[0]) # 輸出預測最高的三個類別
# ---
# ## TensorFlow Hub
#
# Install:
# ```
# pip install tensorflow-hub
# ```
#
# Search:
# https://tfhub.dev/
import tensorflow as tf
import tensorflow_hub as hub
# ### 創建EfficientNet B0模型
#
# Model:
# https://tfhub.dev/tensorflow/efficientnet/b0/classification/1
#
# num_classes = 1001 classes of the classification from the original training
#
# Image:height x width = 224 x 224 pixels, 3 RGB color values in the range 0~1
#
# labels file: https://storage.googleapis.com/download.tensorflow.org/data/ImageNetLabels.txt
# +
# EfficientNet B0預訓練模型的URL
module_url = "https://tfhub.dev/tensorflow/efficientnet/b0/classification/1"
# 創建一個Sequential Model,網路模型裡面包含了EfficientNet B0網路層
model = tf.keras.Sequential([
# hub.KerasLayer將載入的EfficientNet B0模型封裝成網路層(Keras Layer)
hub.KerasLayer(module_url,
name='EfficientNetB0') # 網路層名稱
])
model.build([None, 224, 224, 3])
# -
model.summary()
# ### 資料前處理和輸出解碼
# 創建資料前處理函式:
def read_img(img_path, resize=(224,224)):
img_string = tf.io.read_file(img_path) # 讀取檔案
img_decode = tf.image.decode_image(img_string) # 將檔案以影像格式來解碼
img_decode = tf.image.convert_image_dtype(img_decode, tf.float32)
img_decode = tf.image.resize(img_decode, resize) # 將影像resize到網路輸入大小
# 將影像格式增加到4維(batch, height, width, channels),模型預測要求格式
img_decode = tf.expand_dims(img_decode, axis=0) #
return img_decode
# 創建輸出解碼器:
# +
# 下載ImageNet 的標籤檔
labels_path = tf.keras.utils.get_file('ImageNetLabels.txt', 'https://storage.googleapis.com/download.tensorflow.org/data/ImageNetLabels.txt')
# 讀取標籤檔中的數據
with open(labels_path) as file:
lines = file.read().splitlines()
print(lines) # 顯示讀取的標籤
imagenet_labels = np.array(lines) # 將標籤轉成numpy array做為網路輸出的解碼器
# -
# ### 預測輸出結果
# 從資料夾中讀取一張影像(elephant.jpg)作為測試:
img_path = '../image/elephant.jpg'
img = read_img(img_path) # 透過剛創建的函式讀取影像
plt.imshow(img[0])
# 預測結果:
preds = model.predict(img) # 預測圖片
index = np.argmax(preds) # 取得預測結果最大的Index
print("Predicted:", imagenet_labels[index]) # 透過解碼器將輸出轉成標籤
# 顯示最好的三個預測:
# 取得預測結果最大的三個indexs
top3_indexs = np.argsort(preds)[0, ::-1][:3]
print("Predicted:", imagenet_labels[top3_indexs+1]) # 透過解碼器將輸出轉成標籤
|
Exercise/Solution8.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _kg_hide-input=false _kg_hide-output=true _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 56.552131, "end_time": "2021-06-14T20:00:56.448284", "exception": false, "start_time": "2021-06-14T19:59:59.896153", "status": "completed"} tags=[]
# !pip install keras-tcn tensorflow==2.5.0
# + papermill={"duration": 2.765742, "end_time": "2021-06-14T20:00:59.383374", "exception": false, "start_time": "2021-06-14T20:00:56.617632", "status": "completed"} tags=[]
# data setup code
import numpy as np # linear algebra
np.set_printoptions(precision=3, suppress=True) # improve printing
from scipy.signal import lfilter
import scipy
import tensorflow as tf
from tensorflow.keras import layers, models, losses, metrics, Input, utils
import tensorflow_addons as tfa
from tcn import TCN, tcn_full_summary
import os
from random import choice
import re
from scipy.fft import fft
file_name_regex = re.compile("([I\d]{3})_SIG_II\.npy")
# file loader
def load_files(path):
data = {}
for entry in os.scandir(path):
if entry.is_dir():
for file in os.scandir(entry.path):
match = file_name_regex.match(file.name)
if match and file.is_file():
data[match.groups()[0]] = np.load(file.path)
return data
# + papermill={"duration": 0.1736, "end_time": "2021-06-14T20:00:59.719837", "exception": false, "start_time": "2021-06-14T20:00:59.546237", "status": "completed"} tags=[]
print(scipy.__version__, np.__version__, tf.__version__)
# + papermill={"duration": 0.299321, "end_time": "2021-06-14T20:01:00.177946", "exception": false, "start_time": "2021-06-14T20:00:59.878625", "status": "completed"} tags=[]
try:
# detect and init the TPU
tpu = tf.distribute.cluster_resolver.TPUClusterResolver.connect()
# instantiate a distribution strategy
strategy = tf.distribute.experimental.TPUStrategy(tpu)
except ValueError:
print("No TPU detected; Defaulting to CPU or GPU training.")
print("GPUs available: ", tf.config.list_physical_devices('GPU'))
strategy = tf.distribute.get_strategy()
# + papermill={"duration": 3.9335, "end_time": "2021-06-14T20:01:04.333809", "exception": false, "start_time": "2021-06-14T20:01:00.400309", "status": "completed"} tags=[]
# actually load the data
data = load_files("../input/ecg-lead-2-dataset-physionet-open-access/db_npy")
# + papermill={"duration": 0.168644, "end_time": "2021-06-14T20:01:04.662192", "exception": false, "start_time": "2021-06-14T20:01:04.493548", "status": "completed"} tags=[]
# select the person you want the model to learn to recognise:
target = choice(list(data.keys()))
# you can also define it manually:
# target = "100"
print(target)
# + papermill={"duration": 5.834116, "end_time": "2021-06-14T20:01:10.654899", "exception": false, "start_time": "2021-06-14T20:01:04.820783", "status": "completed"} tags=[]
# prepare training and validation data
training_data, validation_data, training_labels, validation_labels = ([], [], [], [])
length = len(data.keys())
def moving_average(ar, N):
return lfilter(np.ones(N)/N, [1], ar)
for index, (label, array) in enumerate(data.items()):
# cut the samples at 30 minutes - makes dividing the data easier
if array.size>230400:
array = array[:230400]
noise = np.random.normal(0,1,230400)
array = array+noise
norm = np.linalg.norm(array)
array = array/norm
array = moving_average(array, 5)
split = np.array_split(array, 40)
checks = [choice(list(data.keys())) if choice([True,False]) else label for i in range(len(split))]
training_data.extend([np.reshape(fft(arr), (45, 128)) for arr in split[:20]])
training_labels.extend([np.insert(np.zeros(length-1), index, 1)] * 20)
validation_data.extend([np.reshape(fft(arr), (45, 128)) for arr in split[20:]])
validation_labels.extend([np.insert(np.zeros(length-1), index, 1)]*len(checks[20:]))
# + papermill={"duration": 11.005554, "end_time": "2021-06-14T20:01:21.985212", "exception": false, "start_time": "2021-06-14T20:01:10.979658", "status": "completed"} tags=[]
training_dataset = tf.data.Dataset.from_tensor_slices((training_data, training_labels))
validation_dataset = tf.data.Dataset.from_tensor_slices((validation_data, validation_labels))
# + papermill={"duration": 0.178394, "end_time": "2021-06-14T20:01:22.328049", "exception": false, "start_time": "2021-06-14T20:01:22.149655", "status": "completed"} tags=[]
training_dataset
# + papermill={"duration": 0.176715, "end_time": "2021-06-14T20:01:22.668824", "exception": false, "start_time": "2021-06-14T20:01:22.492109", "status": "completed"} tags=[]
# shuffle and batch the dataset
BATCH_SIZE = 64 * strategy.num_replicas_in_sync
SHUFFLE_BUFFER_SIZE = 1000
training_dataset = training_dataset.shuffle(SHUFFLE_BUFFER_SIZE).batch(BATCH_SIZE)
validation_dataset = validation_dataset.batch(BATCH_SIZE)
# + papermill={"duration": 0.993236, "end_time": "2021-06-14T20:01:23.824983", "exception": false, "start_time": "2021-06-14T20:01:22.831747", "status": "completed"} tags=[]
# prepare
with strategy.scope():
used_metrics = [
metrics.TruePositives(name='tp'),
metrics.FalsePositives(name="fp"),
metrics.TrueNegatives(name='tn'),
metrics.FalseNegatives(name='fn'),
metrics.CategoricalAccuracy(name='accuracy'),
]
checkpoint_callback = tf.keras.callbacks.ModelCheckpoint(filepath="./checkpoints/checkpoint.ckpt",
save_weights_only=True,
save_best_only=True,
monitor='val_accuracy',
verbose=1)
model = models.Sequential([
TCN(input_shape=(45,128), kernel_size=3, use_skip_connections=True, nb_filters=64, dilations=[1,2,4,8], return_sequences=True, use_batch_norm=True, dropout_rate=0.05),
TCN(kernel_size=3, use_skip_connections=True, nb_filters=16, dilations=[1,2,4,8], use_batch_norm=True, dropout_rate=0.05),
layers.Dense(32, activation="linear"),
layers.Dense(96, activation="linear"),
layers.Dense(length, activation="softmax")
])
model.compile(loss="categorical_crossentropy", optimizer="adam", metrics=used_metrics)
# + papermill={"duration": 0.621521, "end_time": "2021-06-14T20:01:24.609072", "exception": false, "start_time": "2021-06-14T20:01:23.987551", "status": "completed"} tags=[]
utils.plot_model(model, show_shapes=True)
# + papermill={"duration": 2104.590333, "end_time": "2021-06-14T20:36:29.363330", "exception": false, "start_time": "2021-06-14T20:01:24.772997", "status": "completed"} tags=[]
epochs = 200
history = model.fit(
training_dataset,
validation_data=validation_dataset,
epochs=epochs,
callbacks=[checkpoint_callback]
)
# + papermill={"duration": 3.574304, "end_time": "2021-06-14T20:36:36.541535", "exception": false, "start_time": "2021-06-14T20:36:32.967231", "status": "completed"} tags=[]
model.load_weights("./checkpoints/checkpoint.ckpt")
# + papermill={"duration": 7.233456, "end_time": "2021-06-14T20:36:47.219487", "exception": false, "start_time": "2021-06-14T20:36:39.986031", "status": "completed"} tags=[]
# evaluate
loss, *validation_metrics = model.evaluate(validation_dataset)
print("Loss: ", loss)
print("Accuracy: ", validation_metrics)
# + papermill={"duration": 6.226871, "end_time": "2021-06-14T20:36:56.960354", "exception": false, "start_time": "2021-06-14T20:36:50.733483", "status": "completed"} tags=[]
prediction_test = model.predict(validation_dataset)
# + papermill={"duration": 3.71935, "end_time": "2021-06-14T20:37:04.152464", "exception": false, "start_time": "2021-06-14T20:37:00.433114", "status": "completed"} tags=[]
#for element in prediction_test:
# print(element)
print(element[0])
# + papermill={"duration": 3.440987, "end_time": "2021-06-14T20:37:11.394466", "exception": false, "start_time": "2021-06-14T20:37:07.953479", "status": "completed"} tags=[]
# + papermill={"duration": 16.51308, "end_time": "2021-06-14T20:37:31.773911", "exception": false, "start_time": "2021-06-14T20:37:15.260831", "status": "completed"} tags=[]
# save the model:
model.save_weights('./checkpoints/final_checkpoint.ckpt')
model.save_weights('./checkpoints/final_checkpoint.h5')
model.save('./checkpoints/model.h5')
tf.saved_model.save(model, "./model")
# + papermill={"duration": 9.852756, "end_time": "2021-06-14T20:37:45.072239", "exception": false, "start_time": "2021-06-14T20:37:35.219483", "status": "completed"} tags=[]
# convert to tflite
converter = tf.lite.TFLiteConverter.from_saved_model("./model")
tflite_model = converter.convert()
with open('./model.tflite', 'wb') as f:
f.write(tflite_model)
# + papermill={"duration": 3.70924, "end_time": "2021-06-14T20:37:52.211934", "exception": false, "start_time": "2021-06-14T20:37:48.502694", "status": "completed"} tags=[]
|
software/prototyp.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="rX8mhOLljYeM"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab_type="code" id="BZSlp3DAjdYf" colab={}
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="3wF5wszaj97Y"
# # TensorFlow 2 quickstart for experts
# + [markdown] colab_type="text" id="DUNzJc4jTj6G"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/quickstart/advanced"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/quickstart/advanced.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/quickstart/advanced.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/quickstart/advanced.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="hiH7AC-NTniF"
# This is a [Google Colaboratory](https://colab.research.google.com/notebooks/welcome.ipynb) notebook file. Python programs are run directly in the browser—a great way to learn and use TensorFlow. To follow this tutorial, run the notebook in Google Colab by clicking the button at the top of this page.
#
# 1. In Colab, connect to a Python runtime: At the top-right of the menu bar, select *CONNECT*.
# 2. Run all the notebook code cells: Select *Runtime* > *Run all*.
# + [markdown] colab_type="text" id="eOsVdx6GGHmU"
# Download and install TensorFlow 2. Import TensorFlow into your program:
#
# Note: Upgrade `pip` to install the TensorFlow 2 package. See the [install guide](https://www.tensorflow.org/install) for details.
# + colab_type="code" id="ioLbtB3uGKPX" colab={}
try:
# # %tensorflow_version only exists in Colab.
# %tensorflow_version 2.x
except Exception:
pass
# + [markdown] colab_type="text" id="QS7DDTiZGRTo"
# Import TensorFlow into your program:
# + colab_type="code" id="0trJmd6DjqBZ" colab={}
from __future__ import absolute_import, division, print_function, unicode_literals
import tensorflow as tf
from tensorflow.keras.layers import Dense, Flatten, Conv2D
from tensorflow.keras import Model
# + [markdown] colab_type="text" id="7NAbSZiaoJ4z"
# Load and prepare the [MNIST dataset](http://yann.lecun.com/exdb/mnist/).
# + colab_type="code" id="JqFRS6K07jJs" colab={}
mnist = tf.keras.datasets.mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
# Add a channels dimension
x_train = x_train[..., tf.newaxis]
x_test = x_test[..., tf.newaxis]
# + [markdown] colab_type="text" id="k1Evqx0S22r_"
# Use `tf.data` to batch and shuffle the dataset:
# + colab_type="code" id="8Iu_quO024c2" colab={}
train_ds = tf.data.Dataset.from_tensor_slices(
(x_train, y_train)).shuffle(10000).batch(32)
test_ds = tf.data.Dataset.from_tensor_slices((x_test, y_test)).batch(32)
# + [markdown] colab_type="text" id="BPZ68wASog_I"
# Build the `tf.keras` model using the Keras [model subclassing API](https://www.tensorflow.org/guide/keras#model_subclassing):
# + colab_type="code" id="h3IKyzTCDNGo" colab={}
class MyModel(Model):
def __init__(self):
super(MyModel, self).__init__()
self.conv1 = Conv2D(32, 3, activation='relu')
self.flatten = Flatten()
self.d1 = Dense(128, activation='relu')
self.d2 = Dense(10)
def call(self, x):
x = self.conv1(x)
x = self.flatten(x)
x = self.d1(x)
return self.d2(x)
# Create an instance of the model
model = MyModel()
# + [markdown] colab_type="text" id="uGih-c2LgbJu"
# Choose an optimizer and loss function for training:
# + colab_type="code" id="u48C9WQ774n4" colab={}
loss_object = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True)
optimizer = tf.keras.optimizers.Adam()
# + [markdown] colab_type="text" id="JB6A1vcigsIe"
# Select metrics to measure the loss and the accuracy of the model. These metrics accumulate the values over epochs and then print the overall result.
# + colab_type="code" id="N0MqHFb4F_qn" colab={}
train_loss = tf.keras.metrics.Mean(name='train_loss')
train_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(name='train_accuracy')
test_loss = tf.keras.metrics.Mean(name='test_loss')
test_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(name='test_accuracy')
# + [markdown] colab_type="text" id="ix4mEL65on-w"
# Use `tf.GradientTape` to train the model:
# + colab_type="code" id="OZACiVqA8KQV" colab={}
@tf.function
def train_step(images, labels):
with tf.GradientTape() as tape:
# training=True is only needed if there are layers with different
# behavior during training versus inference (e.g. Dropout).
predictions = model(images, training=True)
loss = loss_object(labels, predictions)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
train_loss(loss)
train_accuracy(labels, predictions)
# + [markdown] colab_type="text" id="Z8YT7UmFgpjV"
# Test the model:
# + colab_type="code" id="xIKdEzHAJGt7" colab={}
@tf.function
def test_step(images, labels):
# training=False is only needed if there are layers with different
# behavior during training versus inference (e.g. Dropout).
predictions = model(images, training=False)
t_loss = loss_object(labels, predictions)
test_loss(t_loss)
test_accuracy(labels, predictions)
# + colab_type="code" id="i-2pkctU_Ci7" colab={}
EPOCHS = 5
for epoch in range(EPOCHS):
# Reset the metrics at the start of the next epoch
train_loss.reset_states()
train_accuracy.reset_states()
test_loss.reset_states()
test_accuracy.reset_states()
for images, labels in train_ds:
train_step(images, labels)
for test_images, test_labels in test_ds:
test_step(test_images, test_labels)
template = 'Epoch {}, Loss: {}, Accuracy: {}, Test Loss: {}, Test Accuracy: {}'
print(template.format(epoch+1,
train_loss.result(),
train_accuracy.result()*100,
test_loss.result(),
test_accuracy.result()*100))
# + [markdown] colab_type="text" id="T4JfEh7kvx6m"
# The image classifier is now trained to ~98% accuracy on this dataset. To learn more, read the [TensorFlow tutorials](https://www.tensorflow.org/tutorials).
|
site/en/tutorials/quickstart/advanced.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + nbsphinx="hidden"
import open3d as o3d
import numpy as np
import os
import sys
# monkey patches visualization and provides helpers to load geometries
sys.path.append('..')
import open3d_tutorial as o3dtut
# change to True if you want to interact with the visualization windows
o3dtut.interactive = not "CI" in os.environ
# -
# # File IO
# This tutorial shows how basic data structures are read and written by Open3D.
# ## Point cloud
# The code below reads and writes a point cloud.
print("Testing IO for point cloud ...")
pcd = o3d.io.read_point_cloud("../../test_data/fragment.pcd")
print(pcd)
o3d.io.write_point_cloud("copy_of_fragment.pcd", pcd)
# By default, Open3D tries to infer the file type by the filename extension. The following point cloud file types are supported:
#
# Format | Description
# ---------|---------------
# `xyz` | Each line contains `[x, y, z]`, where `x`, `y`, `z` are the 3D coordinates
# `xyzn` | Each line contains `[x, y, z, nx, ny, nz]`, where `nx`, `ny`, `nz` are the normals
# `xyzrgb` | Each line contains `[x, y, z, r, g, b]`, where `r`, `g`, `b` are in floats of range `[0, 1]`
# `pts` | The first line is an integer representing the number of points. The subsequent lines follow one of these formats: `[x, y, z, i, r, g, b]`, `[x, y, z, r, g, b]`, `[x, y, z, i]` or `[x, y, z]`, where `x`, `y`, `z`, `i` are of type `double` and `r`, `g`, `b` are of type `uint8`
# `ply` | See [Polygon File Format](http://paulbourke.net/dataformats/ply), the ply file can contain both point cloud and mesh data
# `pcd` | See [Point Cloud Data](http://pointclouds.org/documentation/tutorials/pcd_file_format.html)
#
# It’s also possible to specify the file type explicitly. In this case, the file extension will be ignored.
pcd = o3d.io.read_point_cloud("../../test_data/my_points.txt", format='xyz')
# ## Mesh
# The code below reads and writes a mesh.
print("Testing IO for meshes ...")
mesh = o3d.io.read_triangle_mesh("../../test_data/knot.ply")
print(mesh)
o3d.io.write_triangle_mesh("copy_of_knot.ply", mesh)
# Compared to the point cloud data structure, a mesh has triangles that define the 3D surface.
#
# By default, Open3D tries to infer the file type by the filename extension. The following mesh file types are supported:
#
# Format | Description
# ----------------|---------------
# `ply` | See [Polygon File Format](http://paulbourke.net/dataformats/ply/), the ply file can contain both point cloud and mesh data
# `stl` | See [StereoLithography](http://www.fabbers.com/tech/STL_Format)
# `obj` | See [Object Files](http://paulbourke.net/dataformats/obj/)
# `off` | See [Object File Format](http://www.geomview.org/docs/html/OFF.html)
# `gltf`/`glb` | See [GL Transmission Format](https://github.com/KhronosGroup/glTF/tree/master/specification/2.0)
# ## Image
# The code below reads and writes an image.
print("Testing IO for images ...")
img = o3d.io.read_image("../../test_data/lena_color.jpg")
print(img)
o3d.io.write_image("copy_of_lena_color.jpg", img)
# The size of the image is readily displayed using `print(img)`.
#
# Both `jpg` and `png` image files are supported.
|
examples/python/geometry/file_io.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/probml/probml-notebooks/blob/main/notebooks/Superimport.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="_Qd2diMQZlQL"
# # Superimport demo
#
# The [superimport library](https://github.com/probml/superimport), written by [<NAME>](https://github.com/mjsML), takes care of installing missing python packages for you. All you have to do is type `pip install superimport` (once per colab session), and then add `import superimport` to the top of any of your python files; then, when you run those files, superimport will read the source code, figure out any missing dependencies, install them for you automagically, and then run the rest of your code as usual. We illustrate this below.
#
#
# + id="1G16DyRzi6kK" colab={"base_uri": "https://localhost:8080/"} outputId="b5baeea5-861e-4436-d126-0d6275a254b4"
# !pip install superimport -qqq
# !pip install deimport -qqq
# + id="iiC72KXh5jae"
import superimport
def try_deimport():
try:
from deimport.deimport import deimport
deimport(superimport,verbose=False)
except Exception as e:
print(e)
# + [markdown] id="V_82RC0lahoP"
# # An example with PgmPy
#
# Colab has most popular ML packages already installed. However, there are a few missing ones, such as [PgmPy](https://github.com/pgmpy/pgmpy). Below we create a short file, called `test.py`, that relies on that missing library. We then show what happens if we try to run the script without first installing the library.
# + colab={"base_uri": "https://localhost:8080/"} id="UuJYZu_1kP1B" outputId="1e0b7fc1-98a6-4d54-a92c-ecc378ea54cb"
# %%file test.py
import pgmpy
import numpy
import matplotlib
print('pgmpy ', pgmpy.__version__)
# + [markdown] id="mkwHEjfwknJt"
# Without importing superimport, if you have a missing package your script will fail.
# + colab={"base_uri": "https://localhost:8080/", "height": 215} id="6T6BDgaRkSdM" outputId="437179d6-4d6b-4f40-d570-1283f5108549"
# %run test.py
# + [markdown] id="cZd4ITLYa33l"
#
#
# Now we add one new line to our file: `import superimport`
# + colab={"base_uri": "https://localhost:8080/"} id="fMPOfeIHi9i3" outputId="b78a00c1-cd7b-41da-9287-cd86ea719b4e"
# %%file test.py
import superimport
import pgmpy
import numpy
import matplotlib
print('pgmpy ', pgmpy.__version__)
# + [markdown] id="aIFQaI6pkxIL"
# We can now successfully the script, and it will install any missing packages.
#
#
# Note, however, that we have to deimport the `superimport` symbol before running any code that uses superimport, to force the package to be reloaded (and hence re-executed), otherwise colab will use the cached version (if available) of superimport, which may be stale.
# + colab={"base_uri": "https://localhost:8080/"} id="tL4_358-jJD8" outputId="a8fe754e-af56-400a-c663-39695ad4862b"
try_deimport()
# %run -n test.py
# + [markdown] id="-IDlmbuCgDMx"
# # An example with NumPyro
#
# This time we make a demo that uses numpyro, that is not installed in colab by default.
# + id="SuSqhaqEgQ3z" colab={"base_uri": "https://localhost:8080/"} outputId="682be66a-a277-44e9-b9b9-23c06c9975d1"
# %%file test.py
import superimport
import numpyro
print('numpyro version ', numpyro.__version__)
# + id="yHihouB-gUJK" colab={"base_uri": "https://localhost:8080/"} outputId="8787fa90-4332-42fd-da3c-6c4ce3657df0"
try_deimport()
# %run -n test.py
# + [markdown] id="KJSga2iNeauy"
# # An example with Pyro
#
# This time we make a demo that uses pyro, that is not installed in colab by default. Furthermore, its package name (pyro-ppl) does not match its import name (pyro).
# + id="Sy7eFOQxfQB6" colab={"base_uri": "https://localhost:8080/"} outputId="b16bde2c-d2ef-4cb2-9292-e2f99f70a3d9"
# %%file test.py
import superimport
import pyro
print('pyro version ', pyro.__version__)
# + id="wMgsJq1ieoeH" colab={"base_uri": "https://localhost:8080/"} outputId="9fb807c0-999f-41b8-aaae-4efdde5fe571"
try_deimport()
# %run -n test.py
# + [markdown] id="KFogkyP8gWZ5"
# # An example from the book
# + id="4DvNikcygYlC"
# !git clone --depth 1 https://github.com/probml/pyprobml /pyprobml &> /dev/null
# %cd -q /pyprobml/scripts
# + colab={"base_uri": "https://localhost:8080/", "height": 550} id="aOl69rgCgvVX" outputId="139d0c93-446e-4c9e-a264-11cd353fa1fe"
try_deimport()
# %run -n linreg_residuals_plot.py
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="HpDO7eE1gz-Y" outputId="3e2299dc-01c9-4738-b301-66a05458d3c5"
try_deimport()
# %run -n linreg_poly_vs_degree.py
# + colab={"base_uri": "https://localhost:8080/", "height": 553} id="UnKMPSBOh16k" outputId="5b77a9d2-c1cb-47a3-ea47-c899aea0f19f"
try_deimport()
# %run -n iris_kmeans.py
# + [markdown] id="gERNSvUpcru-"
# # Sharp edges
#
# * There are some packages whose install names differ from their import names (eg we type `pip install pyro-ppl` but `import pyro`). There is a [public mapping file](https://github.com/bndr/pipreqs/blob/master/pipreqs/mapping) stored by pipreqs. However, this is missing some entries (such as pyro). These must be manually added to the [mapping2 file](https://github.com/probml/superimport/blob/main/superimport/mapping2). If your favorite package is missing, open a PR on the superimport repo.
#
# * There are some packages that do not list of all of their requirements.txt (eg GPyOpt depends on matplotlib, but does not mention this). If this 'hidden requirement' is missing, superimport cannot find it either. If it is not already installed in colab, then your script will fail, even with superimport.
# + id="GUWz9Gr-d5SO"
|
notebooks/Superimport.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="JSjG64ra4aFu"
# from google.colab import drive
# drive.mount('/content/drive')
# + id="V8-7SARDZErK"
import torch.nn as nn
import torch.nn.functional as F
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import torch
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
from matplotlib import pyplot as plt
import copy
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
# + id="acRFqJNrZErV" colab={"base_uri": "https://localhost:8080/", "height": 104, "referenced_widgets": ["72cca7ed617b4edfa64e583525e736d2", "245d08d165d94bde923f1f8e7481addd", "90a27c83898046dc8a961614126b2464", "e47b49518d924f3eb0dc11e5dd3efe46", "6457c156a43e4d8d9fd97e05724196f5", "703d6c8eaaa54a03b4347ed2277e5d1a", "61ac81042200465fb8a64b7f401564dc", "8abfbccad4724360ba4dfe26318221e0"]} outputId="50f42aa1-4c2d-43d2-ce96-ce0c9f4d7a86"
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=transform)
# + id="gh5DXuAV1tp5"
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle=False)
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
foreground_classes = {'plane', 'car', 'bird'}
background_classes = {'cat', 'deer', 'dog', 'frog', 'horse','ship', 'truck'}
fg1,fg2,fg3 = 0,1,2
# + id="V_JUhwCeZErk" colab={"base_uri": "https://localhost:8080/"} outputId="2bfc06d9-a432-42fe-aad4-c30eba4d3359"
dataiter = iter(trainloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=10
for i in range(5000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
# + id="uW9MkktGysAp"
def create_mosaic_img(bg_idx,fg_idx,fg):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]].type("torch.DoubleTensor"))
j+=1
else:
image_list.append(foreground_data[fg_idx].type("torch.DoubleTensor"))
label = foreground_label[fg_idx]- fg1 # minus 7 because our fore ground classes are 7,8,9 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
# + id="lWxkp87fNwnM"
desired_num = 30000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(desired_num):
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
# + id="AJuGak6_zXgx"
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label, fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx], self.fore_idx[idx]
batch = 250
msd = MosaicDataset(mosaic_list_of_images, mosaic_label , fore_idx)
train_loader = DataLoader( msd,batch_size= batch ,shuffle=True)
# + id="SadRzWBBZEsP"
class Focus(nn.Module):
def __init__(self):
super(Focus, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=6, kernel_size=3, padding=0)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(in_channels=6, out_channels=6, kernel_size=3, padding=0)
# self.conv3 = nn.Conv2d(in_channels=12, out_channels=32, kernel_size=3, padding=0)
self.fc1 = nn.Linear(1014, 512)
self.fc2 = nn.Linear(512, 64)
# self.fc3 = nn.Linear(512, 64)
# self.fc4 = nn.Linear(64, 10)
self.fc3 = nn.Linear(64,1)
def forward(self,z): #y is avg image #z batch of list of 9 images
y = torch.zeros([batch,3, 32,32], dtype=torch.float64)
x = torch.zeros([batch,9],dtype=torch.float64)
y = y.to("cuda")
x = x.to("cuda")
for i in range(9):
x[:,i] = self.helper(z[:,i])[:,0]
x = F.softmax(x,dim=1)
x1 = x[:,0]
torch.mul(x1[:,None,None,None],z[:,0])
for i in range(9):
x1 = x[:,i]
y = y + torch.mul(x1[:,None,None,None],z[:,i])
return x, y
def helper(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = (F.relu(self.conv2(x)))
# print(x.shape)
# x = (F.relu(self.conv3(x)))
x = x.view(x.size(0), -1)
# print(x.shape)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
# x = F.relu(self.fc3(x))
# x = F.relu(self.fc4(x))
x = self.fc3(x)
return x
# + id="1GvXR1zV5n4w"
focus_net = Focus().double()
focus_net = focus_net.to("cuda")
# + id="LYdCXceZzSk9"
class Classification(nn.Module):
def __init__(self):
super(Classification, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=6, kernel_size=3, padding=0)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(in_channels=6, out_channels=12, kernel_size=3, padding=0)
# self.conv3 = nn.Conv2d(in_channels=12, out_channels=20, kernel_size=3, padding=0)
self.fc1 = nn.Linear(2028, 1024)
self.fc2 = nn.Linear(1024, 64)
# self.fc3 = nn.Linear(512, 64)
# self.fc4 = nn.Linear(64, 10)
self.fc3 = nn.Linear(64,3)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = (F.relu(self.conv2(x)))
# print(x.shape)
# x = (F.relu(self.conv3(x)))
x = x.view(x.size(0), -1)
# print(x.shape)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
# x = F.relu(self.fc3(x))
# x = F.relu(self.fc4(x))
x = self.fc3(x)
return x
# + id="uPYplUGazU9I"
classify = Classification().double()
classify = classify.to("cuda")
# + id="l789TLMP9zJX"
test_images =[] #list of mosaic images, each mosaic image is saved as laist of 9 images
fore_idx_test =[] #list of indexes at which foreground image is present in a mosaic image
test_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(10000):
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx_test.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
test_images.append(image_list)
test_label.append(label)
# + id="bBzV9dKS5po7"
test_data = MosaicDataset(test_images,test_label,fore_idx_test)
test_loader = DataLoader( test_data,batch_size= batch ,shuffle=False)
# + id="n5g3geNJ5zEu"
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer_classify = optim.Adam(classify.parameters(), lr=0.001)#, betas=(0.9, 0.999), eps=1e-08, weight_decay=0, amsgrad=False)
optimizer_focus = optim.Adam(focus_net.parameters(), lr=0.001)#, betas=(0.9, 0.999), eps=1e-08, weight_decay=0, amsgrad=False)
# + id="Nvr_dqW5ReUP"
col1=[]
col2=[]
col3=[]
col4=[]
col5=[]
col6=[]
col7=[]
col8=[]
col9=[]
col10=[]
col11=[]
col12=[]
col13=[]
# + id="iwk82DG9RfYe" colab={"base_uri": "https://localhost:8080/"} outputId="1c6de816-f392-4803-db0e-135832139c5e"
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
count += 1
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
print(count)
print("="*100)
col1.append(0)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
# + id="XYGo-DLbR1ZV" colab={"base_uri": "https://localhost:8080/"} outputId="c3040698-ce09-454e-a481-a894f66b1c9c"
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
# + id="tFfAJZkcZEsY" colab={"base_uri": "https://localhost:8080/"} outputId="315da31e-3295-42b7-fa53-63069e2714f7"
nos_epochs = 200
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for epoch in range(nos_epochs): # loop over the dataset multiple times
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
running_loss = 0.0
epoch_loss = []
cnt=0
iteration = desired_num // batch
#training data set
for i, data in enumerate(train_loader):
inputs , labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
# zero the parameter gradients
optimizer_focus.zero_grad()
optimizer_classify.zero_grad()
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
# print(outputs)
# print(outputs.shape,labels.shape , torch.argmax(outputs, dim=1))
loss = criterion(outputs, labels)
loss.backward()
optimizer_focus.step()
optimizer_classify.step()
running_loss += loss.item()
mini = 60
if cnt % mini == mini-1: # print every 40 mini-batches
print('[%d, %5d] loss: %.3f' %(epoch + 1, cnt + 1, running_loss / mini))
epoch_loss.append(running_loss/mini)
running_loss = 0.0
cnt=cnt+1
if epoch % 5 == 0:
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
if(np.mean(epoch_loss) <= 0.005):
break;
if epoch % 5 == 0:
# focus_net.eval()
# classify.eval()
col1.append(epoch+1)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
#************************************************************************
#testing data set
with torch.no_grad():
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
print('Finished Training')
# + id="n0zuujPPzLHq"
# torch.save(focus_net.state_dict(),"/content/drive/My Drive/Research/Cheating_data/16_experiments_on_cnn_3layers/"+name+"_focus_net.pt")
# + id="WIAJ3UZN8rPE"
# torch.save(classify.state_dict(),"/content/drive/My Drive/Research/Cheating_data/16_experiments_on_cnn_3layers/"+name+"_classify.pt")
# + id="2LgQKXW-8MH-"
columns = ["epochs", "argmax > 0.5" ,"argmax < 0.5", "focus_true_pred_true", "focus_false_pred_true", "focus_true_pred_false", "focus_false_pred_false" ]
# + id="tSKphM888Y5o"
df_train = pd.DataFrame()
df_test = pd.DataFrame()
# + id="FrWoEGXZ8cBO"
df_train[columns[0]] = col1
df_train[columns[1]] = col2
df_train[columns[2]] = col3
df_train[columns[3]] = col4
df_train[columns[4]] = col5
df_train[columns[5]] = col6
df_train[columns[6]] = col7
df_test[columns[0]] = col1
df_test[columns[1]] = col8
df_test[columns[2]] = col9
df_test[columns[3]] = col10
df_test[columns[4]] = col11
df_test[columns[5]] = col12
df_test[columns[6]] = col13
# + id="RGJoMFcK8eTe" colab={"base_uri": "https://localhost:8080/", "height": 583} outputId="738c0640-df22-49bd-eee7-c1376464882e"
df_train
# + id="Ei9HVQBZ8gn4" colab={"base_uri": "https://localhost:8080/", "height": 573} outputId="789d1897-20ba-4baa-93f8-3905c497e2d0"
# plt.figure(12,12)
plt.plot(col1,col2, label='argmax > 0.5')
plt.plot(col1,col3, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.title("On Training set")
plt.show()
plt.plot(col1,col4, label ="focus_true_pred_true ")
plt.plot(col1,col5, label ="focus_false_pred_true ")
plt.plot(col1,col6, label ="focus_true_pred_false ")
plt.plot(col1,col7, label ="focus_false_pred_false ")
plt.title("On Training set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.savefig("train_ftpt.pdf", bbox_inches='tight')
plt.show()
# + id="_QKYVO8i8ivA" colab={"base_uri": "https://localhost:8080/", "height": 583} outputId="3f665667-766c-4a80-c225-fbdc2252ce82"
df_test
# + id="aRlpgnjy8k1n" colab={"base_uri": "https://localhost:8080/", "height": 573} outputId="c4e17e4b-8368-44df-8426-4839834605d9"
# plt.figure(12,12)
plt.plot(col1,col8, label='argmax > 0.5')
plt.plot(col1,col9, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.title("On Testing set")
plt.show()
plt.plot(col1,col10, label ="focus_true_pred_true ")
plt.plot(col1,col11, label ="focus_false_pred_true ")
plt.plot(col1,col12, label ="focus_true_pred_false ")
plt.plot(col1,col13, label ="focus_false_pred_false ")
plt.title("On Testing set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.savefig("test_ftpt.pdf", bbox_inches='tight')
plt.show()
# + id="TTmdoz9L--GB" colab={"base_uri": "https://localhost:8080/"} outputId="1e89480a-e72f-44b3-a10e-d20e752dac58"
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
# + id="40RP4DzU_A2C" colab={"base_uri": "https://localhost:8080/"} outputId="7fd4b73c-255d-425d-a82f-f86f55a0d7e3"
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
# + id="iJEMJnUI9FP2" colab={"base_uri": "https://localhost:8080/"} outputId="f7723f64-9f9f-431b-91a4-d7f8beef1aa3"
correct = 0
total = 0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
# + id="an7qmNLB-Ilb" colab={"base_uri": "https://localhost:8080/"} outputId="c405d5b8-2ba2-4c87-9bfb-b91f94cfd8cd"
correct = 0
total = 0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
# + id="BXCzL7CJYegv" colab={"base_uri": "https://localhost:8080/"} outputId="3cbce00b-6a5f-45ee-f5e8-c6ff5463fa0f"
max_alpha =[]
alpha_ftpt=[]
argmax_more_than_half=0
argmax_less_than_half=0
for i, data in enumerate(test_loader):
inputs, labels,_ = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
alphas, avg = focus_net(inputs)
outputs = classify(avg)
mx,_ = torch.max(alphas,1)
max_alpha.append(mx.cpu().detach().numpy())
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if (focus == fore_idx[j] and predicted[j] == labels[j]):
alpha_ftpt.append(alphas[j][focus].item())
max_alpha = np.concatenate(max_alpha,axis=0)
print(max_alpha.shape)
# + id="AjoipAyjYgEI" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="369ca4f8-22bd-4620-c1af-a86bbed3e658"
plt.figure(figsize=(6,6))
_,bins,_ = plt.hist(max_alpha,bins=50,color ="c")
plt.title("alpha values histogram")
plt.savefig("alpha_hist.pdf")
# + id="H4I73-wzYmwv" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="16a48921-9d4e-4e3a-aed9-dc83ebf1494c"
plt.figure(figsize=(6,6))
_,bins,_ = plt.hist(np.array(alpha_ftpt),bins=50,color ="c")
plt.title("alpha values in ftpt")
plt.savefig("alpha_hist_ftpt.pdf")
# + id="_VlCBPbwYzsM"
|
14_complexity_comparison_cifar_mosaic/2_focus_6_6_classify_6_12/focus_6_6_classify_6_12_r4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Functions Exercises
import unittest
# #### Sum of List
# - function name is `sum_of_list`
# - input arguments are:
# - list of numbers : `nums`
# - return is:
# - number: sum of all elements in `nums`
# +
# A:
# -
# #### Multiply Elements of List
# - function name is `multiply_list`
# - input arguments are:
# - list of numbers : `nums`
# - return is:
# - number: product of all elements in `nums`
# +
# A:
# -
# #### Reverse a String
# - function name is `reverse_string`
# - input arguments are:
# - string : `string_to_reverse`
# - return is:
# - string : `reversed_string`, the reversed result of input `string_to_reverse`
#
# Note: Do not use `[::-1]` indexing to solve this problem, use loops.
# +
# A:
# -
# #### Max of Three
# - function name is `max_of_three`
# - input arguments are:
# - int : `x`
# - int : `y`
# - int : `z`
# - return is:
# - int: max value of `x`, `y`, and `z`
# +
# A:
# -
# #### Unique Elements Only
# - function name is `make_unique`
# - input arguments are:
# - list (mixed types)
# - return is:
# - list (mixed types)
#
# Note: Do not use `set` to solve this problem.
# +
# A:
# -
# #### Is It a [Palindrome?](https://en.wikipedia.org/wiki/Palindrome)
# - function name is `is_word_palindrome`
# - input arguments are:
# - string
# - return is:
# - boolean (True if input string is Palindrome, else False)
#
# Note: Do not use [::-1] indexing to solve this problem.
# Note 2: Whitespace should be considered as a character against which to evaluate symmetry.
# +
# A:
# -
# #### Unit Tests
# Run the cell below to determine if your functions were correctly written.
#
# You will see the following if all your functions were correctly written:
# ```
# ......
# Currently testing 'test_is_word_palindrome'...
# Currently testing 'test_make_unique'...
# Currently testing 'test_max_of_three'...
# Currently testing 'test_multiply_list'...
# Currently testing 'test_reverse_string'...
# Currently testing 'test_sum_of_list'...
#
# ----------------------------------------------------------------------
# Ran 6 tests in 0.008s
#
# OK
# ```
#
#
# If you have an error in your function, you will see an output similar to the following:
# ```
# ..F...
# Currently testing 'test_is_word_palindrome'...
# Currently testing 'test_make_unique'...
# Currently testing 'test_max_of_three'...
# Currently testing 'test_multiply_list'...
# Currently testing 'test_reverse_string'...
# Currently testing 'test_sum_of_list'...
#
# ======================================================================
# FAIL: test_max_of_three (__main__.TestAllFunctions)
# ----------------------------------------------------------------------
# Traceback (most recent call last):
# File "<ipython-input-10-e39454a386b1>", line 17, in test_max_of_three
# self.assertEqual(max_of_three(3, 6, -5), 6)
# AssertionError: -5 != 6
#
# ----------------------------------------------------------------------
# Ran 6 tests in 0.021s
#
# FAILED (failures=1)
# ```
#
# The error we've encountered in the above output occurred at test 3 of 6 (note how the `F` occurrs at the 3rd dot of 6 dots along the top of the output). Our function `test_max_of_three` returned a value of `-5` but the test we used expected a value of `6`. We will need to revisit our function `test_max_of_three` to ensure it returns a value of `6` when passed values of `3`, `6`, and `-5`.
# +
class TestAllFunctions(unittest.TestCase):
def setUp(self):
print("Currently testing \'{0}\'...".format(self._testMethodName))
def test_sum_of_list(self):
self.assertEqual(sum_of_list((8, 2, 3, 0, 7)), 20)
self.assertEqual(sum_of_list([-8, 2, 3, 0, 7]), 4)
def test_multiply_list(self):
self.assertEqual(multiply_list([2, 2, 4]), 16)
def test_reverse_string(self):
self.assertEqual(reverse_string('greg'), 'gerg')
def test_max_of_three(self):
self.assertEqual(max_of_three(3, 6, -5), 6)
self.assertEqual(max_of_three(5, 5, -5), 5)
def test_make_unique(self):
# note that set is used here for order invariance of the list
self.assertEqual(set(make_unique([1,1,1,'greg',2,2,3,3,3,3,'greg'])), set([1,2,3,'greg']))
def test_is_word_palindrome(self):
self.assertFalse(is_word_palindrome('greg'))
self.assertTrue(is_word_palindrome('gohangasalamiimalasagnahog'))
unittest.main(argv=['first-arg-is-ignored'], exit=False);
|
functions/functions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="_bl3voWCkk3h"
# !wget https://zenodo.org/record/3824876/files/SignalTrain_LA2A_Dataset_1.1.tgz?download=1
# + id="4ngnQna5wVUd"
# !tar -xvf SignalTrain_LA2A_Dataset_1.1.tgz?download=1
# + id="4oTslF_uMi7j"
# !ls
# + id="020lsO59kn0X"
from google.colab import drive
drive.mount('/content/drive')
# + id="sf2KwNnqNOzK"
# !mv SignalTrain_LA2A_Dataset_1.1/ "/content/drive/My Drive"
# + id="d7wFbvG3TWjU"
# !mv ssh.tar.gz "/content/drive/My Drive"
# + id="B_KkMs1qTi_i"
# !rm -rf /root/.ssh
# !mkdir /root/.ssh
# !tar -xvzf "/content/drive/My Drive/ssh.tar.gz"
# !cp ssh-colab/* /root/.ssh && rm -rf ssh-colab && rm -rf ssh.tar.gz
# #!chmod 700 /root/.ssh$
# !touch /root/.ssh/known_hosts
# !ssh-keyscan github.com >> /root/.ssh/known_hosts
# !chmod 644 /root/.ssh/known_hosts
# !chmod 600 /root/.ssh/id_rsa_colab
# !ssh -T git@github.com
# + [markdown] id="oeHvtEXRSV5d"
# # Imports
# + id="N88_6TctYA8G"
# !pip install git+ssh://git@github.com/csteinmetz1/auraloss.git
# + id="oGRzWf1PUs74"
# !pip install pytorch_lightning
# !pip install torchaudio
# + id="SNHKahKCSWAy"
import os
import sys
import glob
import torch
import auraloss # here is our package!
import torchaudio
import numpy as np
import torchsummary
from google.colab import output
import pytorch_lightning as pl
from argparse import ArgumentParser
torchaudio.set_audio_backend("sox_io")
# + [markdown] id="StL1vcYmRnr2"
# # Dataset
# + id="uhMSoTGkCBpq"
# first we will load the full dataset onto the local disk (this takes about 20 min)
# !mkdir "/content/data"
# !rsync -aP "/content/drive/My Drive/SignalTrain_LA2A_Dataset_1.1.zip/" "/content/data/"
# !unzip "/content/data/SignalTrain_LA2A_Dataset_1.1.zip"
# + id="a4jNI_eRM--R"
class SignalTrainLA2ADataset(torch.utils.data.Dataset):
""" SignalTrain LA2A dataset. Source: [10.5281/zenodo.3824876](https://zenodo.org/record/3824876)."""
def __init__(self, root_dir, subset="train", length=16384, preload=False):
"""
Args:
root_dir (str): Path to the root directory of the SignalTrain dataset.
subset (str, optional): Pull data either from "train", "val", or "test" subsets. (Default: "train")
length (int, optional): Number of samples in the returned examples. (Default: 40)
preload (bool, optional): Read in all data into RAM during init. (Default: False)
"""
self.root_dir = root_dir
self.subset = subset
self.length = length
self.preload = preload
# get all the target files files in the directory first
self.target_files = glob.glob(os.path.join(self.root_dir, self.subset.capitalize(), "target_*.wav"))
self.input_files = glob.glob(os.path.join(self.root_dir, self.subset.capitalize(), "input_*.wav"))
self.params = [(float(f.split("__")[1].replace(".wav","")), float(f.split("__")[2].replace(".wav",""))) for f in self.target_files]
self.examples = []
self.audio_files = []
self.hours = 0 # total number of hours of data in the subset
# ensure that the sets are ordered correctlty
self.target_files.sort()
self.input_files.sort()
# loop over files to count total length
for idx, (tfile, ifile, params) in enumerate(zip(self.target_files, self.input_files, self.params)):
print(os.path.basename(tfile), os.path.basename(ifile))
md = torchaudio.info(tfile)
self.hours += (md.num_frames / md.sample_rate) / 3600
num_frames = md.num_frames
if self.preload:
output.clear('status_text')
with output.use_tags('status_text'):
print(f"* Pre-loading... {idx+1:3d}/{len(self.target_files):3d} ...")
input, sr = torchaudio.load(ifile, normalize=False)
target, sr = torchaudio.load(tfile, normalize=False)
#input /= ((2**31) - 1) # apply float32 normalization
#target /= ((2**31) - 1)
input = input.half()
target = target.half()
self.audio_files.append({"target" : target, "input" : input})
num_frames = input.shape[-1]
if self.subset == "train":
if idx > 25: break
if self.subset == "val":
if idx > 1: break
# create one entry for each patch
for n in range((num_frames // self.length) - 1):
offset = int(n * self.length)
self.examples.append({"idx": idx,
"target_file" : tfile,
"input_file" : ifile,
"params" : params,
"offset": offset,
"frames" : num_frames})
# we then want to get the input files
print(f"Located {len(self.examples)} examples totaling {self.hours:0.1f} hr in the {self.subset} subset.")
def __len__(self):
return len(self.examples)
def __getitem__(self, idx):
if self.preload:
audio_idx = self.examples[idx]["idx"]
offset = self.examples[idx]["offset"]
input = self.audio_files[audio_idx]["input"][:,offset:offset+self.length]
target = self.audio_files[audio_idx]["target"][:,offset:offset+self.length]
else:
offset = self.examples[idx]["offset"]
input, sr = torchaudio.load(self.examples[idx]["input"],
num_frames=self.length,
frame_offset=offset,
normalize=False)
target, sr = torchaudio.load(self.examples[idx]["target"],
num_frames=self.length,
frame_offset=offset,
normalize=False)
# apply float32 normalization
input /= ((2**31) - 1)
target /= ((2**31) - 1)
# at random with p=0.5 flip the phase
if np.random.rand() > 0.5:
input *= -1
target *= -1
# then get the tuple of parameters
params = torch.tensor(self.examples[idx]["params"]).unsqueeze(0)
params[:,1] /= 100
return input, target, params
# + [markdown] id="bhn_QYoXRsAu"
# # Model
# + id="qj9pVigeRrVG"
def center_crop(x, shape):
start = (x.shape[-1]-shape[-1])//2
stop = start + shape[-1]
return x[...,start:stop]
class FiLM(torch.nn.Module):
def __init__(self, num_features, cond_dim):
super(FiLM, self).__init__()
self.num_features = num_features
self.bn = torch.nn.BatchNorm1d(num_features, affine=False)
self.adaptor = torch.nn.Linear(cond_dim, num_features * 2)
def forward(self, x, cond):
cond = self.adaptor(cond)
g, b = torch.chunk(cond, 2, dim=-1)
g = g.permute(0,2,1)
b = b.permute(0,2,1)
x = self.bn(x) # apply BatchNorm without affine
x = (x * g) + b # then apply conditional affine
return x
class TCNBlock(torch.nn.Module):
def __init__(self, in_ch, out_ch, kernel_size=3, padding=0, dilation=1, depthwise=False, conditional=False):
super(TCNBlock, self).__init__()
self.in_ch = in_ch
self.out_ch = out_ch
self.kernel_size = kernel_size
self.padding = padding
self.dilation = dilation
self.depthwise = depthwise
self.conditional = conditional
groups = out_ch if depthwise and (in_ch % out_ch == 0) else 1
self.conv1 = torch.nn.Conv1d(in_ch,
out_ch,
kernel_size=kernel_size,
padding=padding,
dilation=dilation,
groups=groups,
bias=False)
self.conv2 = torch.nn.Conv1d(out_ch,
out_ch,
kernel_size=kernel_size,
padding=padding,
dilation=1,
groups=groups,
bias=False)
if depthwise:
self.conv1b = torch.nn.Conv1d(out_ch, out_ch, kernel_size=1)
self.conv2b = torch.nn.Conv1d(out_ch, out_ch, kernel_size=1)
self.bn1 = torch.nn.BatchNorm1d(in_ch)
if conditional:
self.film = FiLM(out_ch, 64)
else:
self.bn2 = torch.nn.BatchNorm1d(out_ch)
self.relu1 = torch.nn.LeakyReLU()
self.relu2 = torch.nn.LeakyReLU()
self.res = torch.nn.Conv1d(in_ch, out_ch, kernel_size=1, bias=False)
def forward(self, x, p=None):
x_in = x
x = self.bn1(x)
x = self.relu1(x)
x = self.conv1(x)
if self.depthwise: # apply pointwise conv
x = self.conv1b(x)
if p is not None: # apply FiLM conditioning
x = self.film(x, p)
else:
x = self.bn2(x)
x = self.relu2(x)
x = self.conv2(x)
if self.depthwise:
x = self.conv2b(x)
x_res = self.res(x_in)
x = x + center_crop(x_res, x.shape)
return x
class TCNModel(pl.LightningModule):
""" Temporal convolutional network with conditioning module.
Params:
nparams (int): Number of conditioning parameters.
ninputs (int): Number of input channels (mono = 1, stereo 2). Default: 1
ninputs (int): Number of output channels (mono = 1, stereo 2). Default: 1
nblocks (int): Number of total TCN blocks. Default: 10
kernel_size (int): Width of the convolutional kernels. Default: 3
dialation_growth (int): Compute the dilation factor at each block as dilation_growth ** (n % stack_size). Default: 1
channel_growth (int): Compute the output channels at each black as in_ch * channel_growth. Default: 2
channel_width (int): When channel_growth = 1 all blocks use convolutions with this many channels. Default: 64
stack_size (int): Number of blocks that constitute a single stack of blocks. Default: 10
depthwise (bool): Use depthwise-separable convolutions to reduce the total number of parameters. Default: False
"""
def __init__(self,
nparams,
ninputs=1,
noutputs=1,
nblocks=10,
kernel_size=3,
dilation_growth=1,
channel_growth=1,
channel_width=64,
stack_size=10,
depthwise=False,
**kwargs):
super(TCNModel, self).__init__()
self.save_hyperparameters()
# setup loss functions
self.l1 = torch.nn.L1Loss()
self.esr = auraloss.time.ESRLoss()
self.dc = auraloss.time.DCLoss()
self.logcosh = auraloss.time.LogCoshLoss()
self.stft = auraloss.freq.STFTLoss()
self.mrstft = auraloss.freq.MultiResolutionSTFTLoss()
self.rrstft = auraloss.freq.RandomResolutionSTFTLoss()
if nparams > 0:
self.gen = torch.nn.Sequential(
torch.nn.Linear(nparams, 16),
torch.nn.PReLU(),
torch.nn.Linear(16, 32),
torch.nn.PReLU(),
torch.nn.Linear(32, 64),
torch.nn.PReLU()
)
self.blocks = torch.nn.ModuleList()
for n in range(nblocks):
in_ch = out_ch if n > 0 else ninputs
out_ch = in_ch * channel_growth if channel_growth > 1 else channel_width
dilation = dilation_growth ** (n % stack_size)
self.blocks.append(TCNBlock(in_ch,
out_ch,
kernel_size=kernel_size,
dilation=dilation,
depthwise=self.hparams.depthwise,
conditional=True if nparams > 0 else False))
self.output = torch.nn.Conv1d(out_ch, noutputs, kernel_size=1)
def forward(self, x, p=None):
# if parameters present,
# compute global conditioning
if p is not None:
cond = self.gen(p)
else:
cond = None
# iterate over blocks passing conditioning
for block in self.blocks:
x = block(x, cond)
return self.output(x)
def compute_receptive_field(self):
""" Compute the receptive field in samples."""
rf = self.hparams.kernel_size
for n in range(1,self.hparams.nblocks):
dilation = self.hparams.dilation_growth ** (n % self.hparams.stack_size)
rf = rf + ((self.hparams.kernel_size-1) * dilation)
rf = rf + ((self.hparams.kernel_size-1) * 1)
return rf
def training_step(self, batch, batch_idx):
input, target, params = batch
# pass the input thrgouh the mode
pred = self(input, params)
# crop the target signal
target = center_crop(target, pred.shape)
# compute the error using appropriate loss
if self.hparams.train_loss == "l1":
loss = self.l1(pred, target)
elif self.hparams.train_loss == "esr+dc":
loss = self.esr(pred, target) + self.dc(pred, target)
elif self.hparams.train_loss == "logcosh":
loss = self.logcosh(pred, target)
elif self.hparams.train_loss == "stft":
loss = torch.stack(self.stft(pred, target),dim=0).sum()
elif self.hparams.train_loss == "mrstft":
loss = torch.stack(self.mrstft(pred, target),dim=0).sum()
elif self.hparams.train_loss == "rrstft":
loss = torch.stack(self.rrstft(pred, target),dim=0).sum()
else:
raise NotImplementedError(f"Invalid loss fn: {self.hparams.train_loss}")
self.log('train_loss',
loss,
on_step=True,
on_epoch=True,
prog_bar=True,
logger=True)
return loss
def validation_step(self, batch, batch_idx):
input, target, params = batch
# pass the input thrgouh the mode
pred = self(input, params)
# crop the target signal
target = center_crop(target, pred.shape)
# compute the validation error using all losses
l1_loss = self.l1(pred, target)
esr_loss = self.esr(pred, target)
dc_loss = self.dc(pred, target)
logcosh_loss = self.logcosh(pred, target)
stft_loss = torch.stack(self.stft(pred, target),dim=0).sum()
mrstft_loss = torch.stack(self.mrstft(pred, target),dim=0).sum()
rrstft_loss = torch.stack(self.rrstft(pred, target),dim=0).sum()
aggregate_loss = l1_loss + \
esr_loss + \
dc_loss + \
logcosh_loss + \
mrstft_loss + \
stft_loss + \
rrstft_loss
self.log('val_loss', aggregate_loss)
self.log('val_loss/L1', l1_loss)
self.log('val_loss/ESR', esr_loss)
self.log('val_loss/DC', dc_loss)
self.log('val_loss/LogCosh', logcosh_loss)
self.log('val_loss/STFT', stft_loss)
self.log('val_loss/MRSTFT', mrstft_loss)
self.log('val_loss/RRSTFT', rrstft_loss)
# move tensors to cpu for logging
outputs = {
"input" : input.cpu().numpy(),
"target": target.cpu().numpy(),
"pred" : pred.cpu().numpy()}
return outputs
def validation_epoch_end(self, validation_step_outputs):
# flatten the output validation step dicts to a single dict
outputs = res = {k: v for d in validation_step_outputs for k, v in d.items()}
i = outputs["input"][0].squeeze()
c = outputs["target"][0].squeeze()
p = outputs["pred"][0].squeeze()
# log audio examples
self.logger.experiment.add_audio("input", i, self.global_step, sample_rate=self.hparams.sample_rate)
self.logger.experiment.add_audio("target", c, self.global_step, sample_rate=self.hparams.sample_rate)
self.logger.experiment.add_audio("pred", p, self.global_step, sample_rate=self.hparams.sample_rate)
def configure_optimizers(self):
return torch.optim.Adam(self.parameters(), lr=self.hparams.lr)
# add any model hyperparameters here
@staticmethod
def add_model_specific_args(parent_parser):
parser = ArgumentParser(parents=[parent_parser], add_help=False)
# --- model related ---
parser.add_argument('--ninputs', type=int, default=1)
parser.add_argument('--noutputs', type=int, default=1)
parser.add_argument('--nblocks', type=int, default=10)
parser.add_argument('--kernel_size', type=int, default=3)
parser.add_argument('--dilation_growth', type=int, default=1)
parser.add_argument('--channel_growth', type=int, default=1)
parser.add_argument('--channel_width', type=int, default=64)
parser.add_argument('--stack_size', type=int, default=10)
parser.add_argument('--depthwise', default=False, action='store_true')
# --- training related ---
parser.add_argument('--lr', type=float, default=1e-3)
parser.add_argument('--train_loss', type=str, default="l1")
return parser
# + [markdown] id="<KEY>"
# # Training
# + id="6PBu7O7_R0fp"
# add PROGRAM level args
#root_dir = '/content/drive/My Drive/SignalTrain_LA2A_Dataset_1.1'
root_dir = '/content/data/SignalTrain_LA2A_Dataset_1.1'
sample_rate = 44100
shuffle = True
train_subset = "train"
val_subset = "val"
train_length = 16384
eval_length = 262144
batch_size = 128
num_workers = 0
preload = False
precision = 16
# init the trainer and model
trainer = pl.Trainer(gpus=1, precision=precision)
# setup the dataloaders
train_dataset = SignalTrainLA2ADataset(root_dir,
subset=train_subset,
length=train_length,
preload=preload)
train_dataloader = torch.utils.data.DataLoader(train_dataset,
shuffle=shuffle,
batch_size=batch_size,
num_workers=num_workers)
val_dataset = SignalTrainLA2ADataset(root_dir,
subset=val_subset,
length=eval_length,
preload=preload)
val_dataloader = torch.utils.data.DataLoader(val_dataset,
shuffle=shuffle,
batch_size=batch_size,
num_workers=num_workers)
dict_args = {
"nparams" : 2,
"ninputs" : 1,
"noutputs" : 1,
"nblocks" : 10,
"kernel_size": 3,
"dilation_growth" : 1,
"channel_growth" : 1,
"channel_width" : 64,
"stack_size" : 10,
"depthwise" : False,
"lr" : 0.001,
"sample_rate" : sample_rate,
"train_loss" : "mrstft"
}
model = TCNModel(**dict_args)
# find proper learning rate
#trainer.tune(model, train_dataloader)
#torchsummary.summary(model, [(1,eval_length), (1,2)])
device = "cuda:0"
model.stft.to(device)
# train!
trainer.fit(model, train_dataloader, val_dataloader)
# + id="BF-sjlZbbALa"
# Start tensorboard.
# %reload_ext tensorboard
# %tensorboard --logdir lightning_logs/
# + id="9sp_005lpYl3"
|
examples/auraloss.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PyRosetta.notebooks]
# language: python
# name: conda-env-PyRosetta.notebooks-py
# ---
# Before you turn this problem in, make sure everything runs as expected. First, **restart the kernel** (in the menubar, select Kernel$\rightarrow$Restart) and then **run all cells** (in the menubar, select Cell$\rightarrow$Run All).
#
# Make sure you fill in any place that says `YOUR CODE HERE` or "YOUR ANSWER HERE", as well as your name and collaborators below:
NAME = ""
COLLABORATORS = ""
# ---
# <!--NOTEBOOK_HEADER-->
# *This notebook contains material from [PyRosetta](https://RosettaCommons.github.io/PyRosetta.notebooks);
# content is available [on Github](https://github.com/RosettaCommons/PyRosetta.notebooks.git).*
# <!--NAVIGATION-->
# < [Example of Using PyRosetta with GNU Parallel](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/16.03-GNU-Parallel-Via-Slurm.ipynb) | [Contents](toc.ipynb) | [Index](index.ipynb) | [Part I: Parallelized Global Ligand Docking with `pyrosetta.distributed`](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/16.05-Ligand-Docking-dask.ipynb) ><p><a href="https://colab.research.google.com/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/16.04-dask.delayed-Via-Slurm.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a>
# # Examples Using the `dask` Module
# ### We can make use of the `dask` library to parallelize code
# *Note:* This Jupyter notebook uses parallelization and is **not** meant to be executed within a Google Colab environment.
# *Note:* This Jupyter notebook requires the PyRosetta distributed layer which is obtained by building PyRosetta with the `--serialization` flag or installing PyRosetta from the RosettaCommons conda channel
#
# **Please see Chapter 16.00 for setup instructions**
# +
import dask
import dask.array as da
import graphviz
import logging
logging.basicConfig(level=logging.INFO)
import numpy as np
import os
import pyrosetta
import pyrosetta.distributed
import pyrosetta.distributed.dask
import pyrosetta.distributed.io as io
import random
import sys
from dask.distributed import Client, LocalCluster, progress
from dask_jobqueue import SLURMCluster
from IPython.display import Image
if 'google.colab' in sys.modules:
print("This Jupyter notebook uses parallelization and is therefore not set up for the Google Colab environment.")
sys.exit(0)
# -
# Initialize PyRosetta within this Jupyter notebook using custom command line PyRosetta flags:
flags = """-out:level 100
-ignore_unrecognized_res 1
-ignore_waters 0
-detect_disulf 0 # Do not automatically detect disulfides
""" # These can be unformatted for user convenience, but no spaces in file paths!
pyrosetta.distributed.init(flags)
# If you are running this example on a high-performance computing (HPC) cluster with SLURM scheduling, use the `SLURMCluster` class described below. For more information, visit https://jobqueue.dask.org/en/latest/generated/dask_jobqueue.SLURMCluster.html. **Note**: If you are running this example on a HPC cluster with a job scheduler other than SLURM, `dask_jobqueue` also works with other job schedulers: http://jobqueue.dask.org/en/latest/api.html
#
# The `SLURMCluster` class in the `dask_jobqueue` module is very useful! In this case, we are requesting four workers using `cluster.scale(4)`, and specifying each worker to have:
# - one thread per worker with `cores=1`
# - one process per worker with `processes=1`
# - one CPU per task per worker with `job_cpu=1`
# - a total of 4GB memory per worker with `memory="4GB"`
# - itself run on the "short" queue/partition on the SLURM scheduler with `queue="short"`
# - a maximum job walltime of 3 hours using `walltime="03:00:00"`
# - output dask files directed to `local_directory`
# - output SLURM log files directed to file path and file name (and any other SLURM commands) with the `job_extra` option
# - pre-initialization with the same custom command line PyRosetta flags used in this Jupyter notebook, using the `extra=pyrosetta.distributed.dask.worker_extra(init_flags=flags)` option
#
#
if not os.getenv("DEBUG"):
scratch_dir = os.path.join("/net/scratch", os.environ["USER"])
cluster = SLURMCluster(
cores=1,
processes=1,
job_cpu=1,
memory="4GB",
queue="short",
walltime="02:59:00",
local_directory=scratch_dir,
job_extra=["-o {}".format(os.path.join(scratch_dir, "slurm-%j.out"))],
extra=pyrosetta.distributed.dask.worker_extra(init_flags=flags)
)
cluster.scale(4)
client = Client(cluster)
else:
cluster = None
client = None
# **Note**: The actual sbatch script submitted to the Slurm scheduler under the hood was:
if not os.getenv("DEBUG"):
print(cluster.job_script())
# Otherwise, if you are running this example locally on your laptop, you can still spawn workers and take advantage of the `dask` module:
# +
# cluster = LocalCluster(n_workers=1, threads_per_worker=1)
# client = Client(cluster)
# -
# Open the `dask` dashboard, which shows diagnostic information about the current state of your cluster and helps track progress, identify performance issues, and debug failures:
client
# ### Consider the following example that runs within this Jupyter notebook kernel just fine but could be parallelized:
# +
def inc(x):
return x + 1
def double(x):
return x + 2
def add(x, y):
return x + y
# +
output = []
for x in range(10):
a = inc(x)
b = double(x)
c = add(a, b)
output.append(c)
total = sum(output)
print(total)
# -
# With a slight modification, we can parallelize it on the HPC cluster using the `dask` module
# +
output = []
for x in range(10):
a = dask.delayed(inc)(x)
b = dask.delayed(double)(x)
c = dask.delayed(add)(a, b)
output.append(c)
delayed = dask.delayed(sum)(output)
print(delayed)
# -
# We used the `dask.delayed` function to wrap the function calls that we want to turn into tasks. None of the `inc`, `double`, `add`, or `sum` calls have happened yet. Instead, the object total is a `Delayed` object that contains a task graph of the entire computation to be executed.
#
# Let's visualize the task graph to see clear opportunities for parallel execution.
if not os.getenv("DEBUG"):
delayed.visualize()
# We can now compute this lazy result to execute the graph in parallel:
if not os.getenv("DEBUG"):
total = delayed.compute()
print(total)
# We can also use `dask.delayed` as a python function decorator for identical performance
# +
@dask.delayed
def inc(x):
return x + 1
@dask.delayed
def double(x):
return x + 2
@dask.delayed
def add(x, y):
return x + y
output = []
for x in range(10):
a = inc(x)
b = double(x)
c = add(a, b)
output.append(c)
total = dask.delayed(sum)(output).compute()
print(total)
# -
# We can also use the `dask.array` library, which implements a subset of the NumPy ndarray interface using blocked algorithms, cutting up the large array into many parallelizable small arrays.
#
# See `dask.array` documentation: http://docs.dask.org/en/latest/array.html, along with that of `dask.bag`, `dask.dataframe`, `dask.delayed`, `Futures`, etc.
if not os.getenv("DEBUG"):
x = da.random.random((10000, 10000, 10), chunks=(1000, 1000, 5))
y = da.random.random((10000, 10000, 10), chunks=(1000, 1000, 5))
z = (da.arcsin(x) + da.arccos(y)).sum(axis=(1, 2))
z.compute()
# The dask dashboard allows visualizing parallel computation, including progress bars for tasks. Here is a snapshot of the dask dashboard while executing the previous cell:
Image(filename="inputs/dask_dashboard_example.png")
# For more info on interpreting the dask dashboard, see: https://distributed.dask.org/en/latest/web.html
# # Example Using `dask.delayed` with PyRosetta
# Let's look at a simple example of sending PyRosetta jobs to the `dask-worker`, and the `dask-worker` sending the results back to this Jupyter Notebook.
#
# We will use the crystal structure of the *de novo* mini protein gEHEE_06 from PDB ID 5JG9
# +
@dask.delayed
def mutate(ppose, target, new_res):
import pyrosetta
pose = io.to_pose(ppose)
mutate = pyrosetta.rosetta.protocols.simple_moves.MutateResidue(target=target, new_res=new_res)
mutate.apply(pose)
return io.to_packed(pose)
@dask.delayed
def refine(ppose):
import pyrosetta
pose = io.to_pose(ppose)
scorefxn = pyrosetta.create_score_function("ref2015_cart")
mm = pyrosetta.rosetta.core.kinematics.MoveMap()
mm.set_bb(True)
mm.set_chi(True)
min_mover = pyrosetta.rosetta.protocols.minimization_packing.MinMover()
min_mover.set_movemap(mm)
min_mover.score_function(scorefxn)
min_mover.min_type("lbfgs_armijo_nonmonotone")
min_mover.cartesian(True)
min_mover.tolerance(0.01)
min_mover.max_iter(200)
min_mover.apply(pose)
return io.to_packed(pose)
@dask.delayed
def score(ppose):
import pyrosetta
pose = io.to_pose(ppose)
scorefxn = pyrosetta.create_score_function("ref2015")
total_score = scorefxn(pose)
return pose, total_score
if not os.getenv("DEBUG"):
pose = pyrosetta.io.pose_from_file("inputs/5JG9.clean.pdb")
keep_chA = pyrosetta.rosetta.protocols.grafting.simple_movers.KeepRegionMover(
res_start=str(pose.chain_begin(1)), res_end=str(pose.chain_end(1))
)
keep_chA.apply(pose)
#kwargs = {"extra_options": pyrosetta.distributed._normflags(flags)}
output = []
for target in random.sample(range(1, pose.size() + 1), 10):
if pose.sequence()[target - 1] != "C":
for new_res in ["ALA", "TRP"]:
a = mutate(io.to_packed(pose), target, new_res)
b = refine(a)
c = score(b)
output.append((target, new_res, c[0], c[1]))
delayed_obj = dask.delayed(np.argmin)([x[-1] for x in output])
delayed_obj.visualize()
# -
print(output)
if not os.getenv("DEBUG"):
delayed_result = delayed_obj.persist()
progress(delayed_result)
# The dask progress bar allows visualizing parallelization directly within the Jupyter notebook. Here is a snapshot of the dask progress bar while executing the previous cell:
Image(filename="inputs/dask_progress_bar_example.png")
if not os.getenv("DEBUG"):
result = delayed_result.compute()
print("The mutation with the lowest energy is residue {0} at position {1}".format(output[result][1], output[result][0]))
# *Note*: For best practices while using `dask.delayed`, see: http://docs.dask.org/en/latest/delayed-best-practices.html
# <!--NAVIGATION-->
# < [Example of Using PyRosetta with GNU Parallel](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/16.03-GNU-Parallel-Via-Slurm.ipynb) | [Contents](toc.ipynb) | [Index](index.ipynb) | [Part I: Parallelized Global Ligand Docking with `pyrosetta.distributed`](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/16.05-Ligand-Docking-dask.ipynb) ><p><a href="https://colab.research.google.com/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/16.04-dask.delayed-Via-Slurm.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a>
|
student-notebooks/16.04-dask.delayed-Via-Slurm.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <font size=6>
# <b>Analyze_Text.ipynb:</b> Analyze Text with Pandas and Watson Natural Language Understanding
# </font>
# # Introduction
#
# This notebook shows how the open source library [Text Extensions for Pandas](https://github.com/CODAIT/text-extensions-for-pandas) lets you use use [Pandas](https://pandas.pydata.org/) DataFrames and the [Watson Natural Language Understanding](https://www.ibm.com/cloud/watson-natural-language-understanding) service to analyze natural language text.
#
# We start out with an excerpt from the [plot synopsis from the Wikipedia page
# for *Monty Python and the Holy Grail*](https://en.wikipedia.org/wiki/Monty_Python_and_the_Holy_Grail#Plot).
# We pass this example document to the Watson Natural Language
# Understanding (NLU) service. Then we use Text Extensions for Pandas to convert the output of the
# Watson NLU service to Pandas DataFrames. Next, we perform an example analysis task both with
# and without Pandas to show how Pandas makes analyzing NLP information easier. Finally, we
# walk through all the different DataFrames that Text Extensions for Pandas can extract from
# the output of Watson Natural Language Understanding.
# # Environment Setup
#
# This notebook requires a Python 3.7 or later environment with the following packages:
# * The dependencies listed in the ["requirements.txt" file for Text Extensions for Pandas](https://github.com/CODAIT/text-extensions-for-pandas/blob/master/requirements.txt)
# * The "[ibm-watson](https://pypi.org/project/ibm-watson/)" package, available via `pip install ibm-watson`
# * `text_extensions_for_pandas`
#
# You can satisfy the dependency on `text_extensions_for_pandas` in either of two ways:
#
# * Run `pip install text_extensions_for_pandas` before running this notebook. This command adds the library to your Python environment.
# * Run this notebook out of your local copy of the Text Extensions for Pandas project's [source tree](https://github.com/CODAIT/text-extensions-for-pandas). In this case, the notebook will use the version of Text Extensions for Pandas in your local source tree **if the package is not installed in your Python environment**.
# +
# Core Python libraries
import json
import os
import sys
import pandas as pd
from typing import *
# IBM Watson libraries
import ibm_watson
import ibm_watson.natural_language_understanding_v1 as nlu
import ibm_cloud_sdk_core
# And of course we need the text_extensions_for_pandas library itself.
try:
import text_extensions_for_pandas as tp
except ModuleNotFoundError as e:
# If we're running from within the project source tree and the parent Python
# environment doesn't have the text_extensions_for_pandas package, use the
# version in the local source tree.
if not os.getcwd().endswith("notebooks"):
raise e
if ".." not in sys.path:
sys.path.insert(0, "..")
import text_extensions_for_pandas as tp
# -
# # Set up the Watson Natural Language Understanding Service
#
# In this part of the notebook, we will use the Watson Natural Language Understanding (NLU) service to extract key features from our example document.
#
# You can create an instance of Watson NLU on the IBM Cloud for free by navigating to [this page](https://www.ibm.com/cloud/watson-natural-language-understanding) and clicking on the button marked "Get started free". You can also install your own instance of Watson NLU on [OpenShift](https://www.openshift.com/) by using [IBM Watson Natural Language Understanding for IBM Cloud Pak for Data](
# https://catalog.redhat.com/software/operators/detail/5e9873e13f398525a0ceafe5).
#
# You'll need two pieces of information to access your instance of Watson NLU: An **API key** and a **service URL**. If you're using Watson NLU on the IBM Cloud, you can find your API key and service URL in the IBM Cloud web UI. Navigate to the [resource list](https://cloud.ibm.com/resources) and click on your instance of Natural Language Understanding to open the management UI for your service. Then click on the "Manage" tab to show a page with your API key and service URL.
#
# The cell that follows assumes that you are using the environment variables `IBM_API_KEY` and `IBM_SERVICE_URL` to store your credentials. If you're running this notebook in Jupyter on your laptop, you can set these environment variables while starting up `jupyter notebook` or `jupyter lab`. For example:
# ``` console
# IBM_API_KEY='<my API key>' \
# IBM_SERVICE_URL='<my service URL>' \
# jupyter lab
# ```
#
# Alternately, you can uncomment the first two lines of code below to set the `IBM_API_KEY` and `IBM_SERVICE_URL` environment variables directly.
# **Be careful not to store your API key in any publicly-accessible location!**
# +
# If you need to embed your credentials inline, uncomment the following two lines and
# paste your credentials in the indicated locations.
# os.environ["IBM_API_KEY"] = "<API key goes here>"
# os.environ["IBM_SERVICE_URL"] = "<Service URL goes here>"
# Retrieve the API key for your Watson NLU service instance
if "IBM_API_KEY" not in os.environ:
raise ValueError("Expected Watson NLU api key in the environment variable 'IBM_API_KEY'")
api_key = os.environ.get("IBM_API_KEY")
# Retrieve the service URL for your Watson NLU service instance
if "IBM_SERVICE_URL" not in os.environ:
raise ValueError("Expected Watson NLU service URL in the environment variable 'IBM_SERVICE_URL'")
service_url = os.environ.get("IBM_SERVICE_URL")
# -
# # Connect to the Watson Natural Language Understanding Python API
#
# This notebook uses the IBM Watson Python SDK to perform authentication on the IBM Cloud via the
# `IAMAuthenticator` class. See [the IBM Watson Python SDK documentation](https://github.com/watson-developer-cloud/python-sdk#iam) for more information.
#
# We start by using the API key and service URL from the previous cell to create an instance of the
# Python API for Watson NLU.
natural_language_understanding = ibm_watson.NaturalLanguageUnderstandingV1(
version="2019-07-12",
authenticator=ibm_cloud_sdk_core.authenticators.IAMAuthenticator(api_key)
)
natural_language_understanding.set_service_url(service_url)
natural_language_understanding
# # Pass a Document through the Watson NLU Service
#
# Once you've opened a connection to the Watson NLU service, you can pass documents through
# the service by invoking the [`analyze()` method](https://cloud.ibm.com/apidocs/natural-language-understanding?code=python#analyze).
#
# The [example document](https://raw.githubusercontent.com/CODAIT/text-extensions-for-pandas/master/resources/holy_grail_short.txt) that we use here is an excerpt from
# the plot summary for *Monty Python and the Holy Grail*, drawn from the [Wikipedia entry](https://en.wikipedia.org/wiki/Monty_Python_and_the_Holy_Grail) for that movie.
#
# Let's show what the raw text looks like:
# +
from IPython.core.display import display, HTML
doc_file = "../resources/holy_grail_short.txt"
with open(doc_file, "r") as f:
doc_text = f.read()
display(HTML(f"<b>Document Text:</b><blockquote>{doc_text}</blockquote>"))
# -
# In the code below, we instruct Watson Natural Language Understanding to perform five different kinds of analysis on the example document:
# * entities (with sentiment)
# * keywords (with sentiment and emotion)
# * relations
# * semantic_roles
# * syntax (with sentences, tokens, and part of speech)
#
# See [the Watson NLU documentation](https://cloud.ibm.com/apidocs/natural-language-understanding?code=python#text-analytics-features) for a full description of the types of analysis that NLU can perform.
# Make the request
response = natural_language_understanding.analyze(
text=doc_text,
# TODO: Use this URL once we've pushed the shortened document to Github
#url="https://raw.githubusercontent.com/CODAIT/text-extensions-for-pandas/master/resources/holy_grail_short.txt",
return_analyzed_text=True,
features=nlu.Features(
entities=nlu.EntitiesOptions(sentiment=True, mentions=True),
keywords=nlu.KeywordsOptions(sentiment=True, emotion=True),
relations=nlu.RelationsOptions(),
semantic_roles=nlu.SemanticRolesOptions(),
syntax=nlu.SyntaxOptions(sentences=True,
tokens=nlu.SyntaxOptionsTokens(lemma=True, part_of_speech=True))
)).get_result()
# The response from the `analyze()` method is a Python dictionary. The dictionary contains an entry
# for each pass of analysis requested, plus some additional entries with metadata about the API request
# itself. Here's a list of the keys in `response`:
response.keys()
# # Perform an Example Task
#
# Let's use the information that Watson Natural Language Understanding has extracted from our example document to perform an example task: *Find all the pronouns in each sentence, broken down by sentence.*
#
# This task could serve as first step to a number of more complex tasks, such as
# resolving anaphora (for example, associating "King Arthur" with "his" in the phrase "King Arthur and his squire, Patsy") or analyzing the relationship between sentiment and the gender of pronouns.
#
# We'll start by doing this task using straight Python code that operates directly over the output of Watson NLU's `analyze()` method. Then we'll redo the task using Pandas DataFrames and Text Extensions for Pandas. This exercise will show how Pandas DataFrames can represent the intermediate data structures of an NLP application in a way that is both easier to understand and easier to manipulate with less code.
#
# Let's begin.
# ## Perform the Task Without Using Pandas
#
# All the information that we need to perform our task is in the "syntax" section of the response
# we captured above from Watson NLU's `analyze()` method. Syntax analysis captures a large amount
# of information, so the "syntax" section of the response is very verbose.
#
# For reference, here's the text of our example document again:
#
#
display(HTML(f"<b>Document Text:</b><blockquote>{doc_text}</blockquote>"))
# And here's the output of Watson NLU's syntax analysis, converted to a string:
response["syntax"]
# Buried in the above data structure is all the information we need to perform our example task:
# * The location of every token in the document.
# * The part of speech of every token in the document.
# * The location of every sentence in the document.
#
# The Python code in the next cell uses this information to construct a list of pronouns
# in each sentence in the document.
# +
import collections
# Create a data structure to hold a mapping from sentence identifier
# to a list of pronouns. This step requires defining sentence ids.
def sentence_id(sentence_record: Dict[str, Any]):
return tuple(sentence_record["location"])
pronouns_by_sentence_id = collections.defaultdict(list)
# Pass 1: Use nested for loops to identify pronouns and match them with
# their containing sentences.
# Running time: O(num_tokens * num_sentences), i.e. O(document_size^2)
for t in response["syntax"]["tokens"]:
pos_str = t["part_of_speech"] # Decode numeric POS enum
if pos_str == "PRON":
found_sentence = False
for s in response["syntax"]["sentences"]:
if (t["location"][0] >= s["location"][0]
and t["location"][1] <= s["location"][1]):
found_sentence = True
pronouns_by_sentence_id[sentence_id(s)].append(t)
if not found_sentence:
raise ValueError(f"Token {t} is not in any sentence")
pass # Make JupyterLab syntax highlighting happy
# Pass 2: Translate sentence identifiers to full sentence metadata.
sentence_id_to_sentence = {sentence_id(s): s
for s in response["syntax"]["sentences"]}
result = [
{
"sentence": sentence_id_to_sentence[key],
"pronouns": pronouns
}
for key, pronouns in pronouns_by_sentence_id.items()
]
result
# -
# The code above is quite complex given the simplicity of the task. You would need to stare at the previous cell for a few minutes to convince yourself that the algorithm is correct. This implementation also has scalability issues: The worst-case running time of the nested for loops section is proportional to the square of the document length.
#
# We can do better.
# ## Repeat the Example Task Using Pandas
#
# Let's revisit the example task we just performed in the previous cell. Again, the task is: *Find all the pronouns in each sentence, broken down by sentence.* This time around, let's perform this task using Pandas.
#
# Text Extensions for Pandas includes a function `parse_response()` that turns the output of Watson NLU's `analyze()` function into a dictionary of Pandas DataFrames. Let's run our response object through that conversion.
dfs = tp.io.watson.nlu.parse_response(response)
dfs.keys()
# The output of each analysis pass that Watson NLU performed is now a DataFrame.
# Let's look at the DataFrame for the "syntax" pass:
syntax_df = dfs["syntax"]
syntax_df
# The DataFrame has one row for every token in the document. Each row has information on
# the span of the token, its part of speech, its lemmatized form, and the span of the
# containing sentence.
#
# Let's use this DataFrame to perform our example task a second time.
pronouns_by_sentence = syntax_df[syntax_df["part_of_speech"] == "PRON"][["sentence", "span"]]
pronouns_by_sentence
# That's it. With the DataFrame version of this data, we can perform our example task with **one line of code**.
#
# Specifically, we use a Pandas selection condition to filter out the tokens that aren't pronouns, and then then we
# project down to the columns containing sentence and token spans. The result is another DataFrame that
# we can display directly in our Jupyter notebook.
# # How it Works
#
#
# Let's take a moment to drill into the internals of the DataFrames we just used.
# For reference, here are the first three rows of the syntax analysis DataFrame:
syntax_df.head(3)
# And here is that DataFrame's data type information:
syntax_df.dtypes
# Two of the columns in this DataFrame — "span" and "sentence" — contain
# extension types from the Text Extensions for Pandas library. Let's look first at the "span"
# column.
#
# The "span" column is stored internally using the class `SpanArray` from
# Text Extensions for Pandas.
# `SpanArray` is a subclass of
# [`ExtensionArray`](
# https://pandas.pydata.org/docs/reference/api/pandas.api.extensions.ExtensionArray.html),
# the base class for custom 1-D array types in Pandas.
#
# You can use the property [`pandas.Series.array`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.array.html) to access the `ExtensionArray` behind any Pandas extension type:
print(syntax_df["span"].array)
# Internally, a `SpanArray` is stored as Numpy arrays of begin and end offsets, plus a Python string
# containing the target text. You can access this internal data as properties if your application needs that
# information:
syntax_df["span"].array.begin[:10], syntax_df["span"].array.end[:10]
# You can also convert an individual element of the array into a Python object of type `Span`:
span_obj = syntax_df["span"].array[0]
print(f"\"{span_obj}\" is an object of type {type(span_obj)}")
# Or you can convert the entire array (or a slice of it) into Python objects, one object per span:
syntax_df["span"].iloc[:10].to_numpy()
# A `SpanArray` can also render itself using [Jupyter Notebook callbacks](https://ipython.readthedocs.io/en/stable/config/integrating.html). To
# see the HTML representation of the `SpanArray`, pass the array object
# to Jupyter's [`display()`](https://ipython.readthedocs.io/en/stable/api/generated/IPython.display.html#IPython.display.display)
# function; or make that object be the last line of the cell, as in the following example:
# Show the first 10 tokens in context
syntax_df["span"].iloc[:10].array
# Let's take another look at our DataFrame of syntax information:
syntax_df.head(3)
# The "sentence" column is backed by an object of type `TokenSpanArray`.
# `TokenSpanArray`, another extension type from Text Extensions for Pandas,
# is a version of `SpanArray` for representing a set of spans that are
# constrained to begin and end on token boundaries. In addition to all the
# functionality of a `SpanArray`, a `TokenSpanArray` encodes additional
# information about the relationships between its spans and a tokenization
# of the document.
#
# Here are the distinct elements of the "sentence" column rendered as HTML:
syntax_df["sentence"].unique()
# As the table in the previous cell's output shows, each span in the `TokenSpanArray` has begin and end offsets in terms
# of both characters and tokens. Internally, the `TokenSpanArray` is stored as follows:
# * A Numpy array of begin offsets, measured in tokens
# * A Numpy array of end offsets in tokens
# * A reference to a `SpanArray` of spans representing the tokens
#
# The `TokenSpanArray` object computes the character offsets and covered text of its spans on demand.
#
# Applications can access the internals of a `TokenSpanArray` via the properties `begin_token`, `end_token`, and `tokens`:
token_span_array = syntax_df["sentence"].unique()
print(f"""
Offset information (stored in the TokenSpanArray):
`begin_token` property: {token_span_array.begin_token}
`end_token` property: {token_span_array.end_token}
Token information (`tokens` property, shared among mulitple TokenSpanArrays):
{token_span_array.tokens}
""")
# The extension types in Text Extensions for Pandas support the full set of Pandas array operations. For example, we can build up a DataFrame of the spans of all sentences in the document by applying `pandas.DataFrame.drop_duplicates()` to the `sentence` column:
syntax_df[["sentence"]].drop_duplicates()
# # A More Complex Example
#
# Now that we've had an introduction to the Text Extensions for Pandas span types, let's take another
# look at the DataFrame that our "find pronouns by sentence" code produced:
pronouns_by_sentence
# This DataFrame contains two columns backed by Text Extensions for Pandas span types:
pronouns_by_sentence.dtypes
# That means that we can use the full power of Pandas' high-level operations on this DataFrame.
# Let's use the output of our earlier task to build up a more complex task:
# *Highlight all pronouns in sentences containing the word "Arthur"*
mask = pronouns_by_sentence["sentence"].map(lambda s: s.covered_text).str.contains("Arthur")
pronouns_by_sentence["span"][mask].values
# Here's another variation: *Pair each instance of the word "Arthur" with the pronouns that occur in the same sentence.*
(
syntax_df[syntax_df["span"].array.covered_text == "Arthur"] # Find instances of "Arthur"
.merge(pronouns_by_sentence, on="sentence") # Match with pronouns in the same sentence
.rename(columns={"span_x": "arthur_span", "span_y": "pronoun_span"})
[["arthur_span", "pronoun_span", "sentence"]] # Reorder columns
)
# # Other Outputs of Watson NLU as DataFrames
#
# The examples so far have used the DataFrame representation of Watson Natural Language Understanding's syntax analysis.
# In addition to syntax analysis, Watson NLU can perform several other types of analysis. Let's take a look at the
# DataFrames that Text Extensions for Pandas can produce from the output of Watson NLU.
#
# We'll start by revisiting the results of our earlier code that ran
# ```python
# dfs = tp.io.watson.nlu.parse_response(response)
# ```
# over the `response` object that the Watson NLU's Python API returned. `dfs` is a dictionary of DataFrames.
dfs.keys()
# The "syntax" element of `dfs` contains the syntax analysis DataFrame that we showed earlier.
# Let's take a look at the other elements.
# The "entities" element of `dfs` contains the named entities that Watson Natural Language
# Understanding found in the document.
dfs["entities"].head()
# The "entity_mentions" element of `dfs` contains the locations of individual mentions of
# entities from the "entities" DataFrame.
dfs["entity_mentions"].head()
# Note that the DataFrame under "entitiy_mentions" may contain multiple mentions of the same
# name:
arthur_mentions = dfs["entity_mentions"][dfs["entity_mentions"]["text"] == "Arthur"]
arthur_mentions
# The "type" and "text" columns of the "entity_mentions" DataFrame refer back to the
# "entities" DataFrame columns of the same names.
# You can combine the global and local information about entities into a single DataFrame
# using Pandas' `DataFrame.merge()` method:
arthur_mentions.merge(dfs["entities"], on=["type", "text"], suffixes=["_mention", "_entity"])
dfs["keywords"].head()
dfs["relations"].head()
dfs["semantic_roles"].head()
|
notebooks/Analyze_Text.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pickle, sys
from tqdm import tqdm
import torch, torchvision
from datasets import Fashion200k
from img_text_composition_models import TIRG
# -
embed_dim = 512
dataset_path = '/Users/bo/Downloads/200k/'
model_path = '/Users/bo/Downloads/checkpoint_fashion200k.pth'
text_path = '/Users/bo/Downloads/texts.pkl'
# +
trainset = Fashion200k(path=dataset_path,
split='train',
transform=torchvision.transforms.Compose([
torchvision.transforms.Resize(224),
torchvision.transforms.CenterCrop(224),
torchvision.transforms.ToTensor(),
torchvision.transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])
]))
testset = Fashion200k(path=dataset_path,
split='test',
transform=torchvision.transforms.Compose([
torchvision.transforms.Resize(224),
torchvision.transforms.CenterCrop(224),
torchvision.transforms.ToTensor(),
torchvision.transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])
]))
texts = trainset.get_all_texts()
# +
__copyright__ = "Copyright (c) 2020 Jina AI Limited. All rights reserved."
__license__ = "Apache-2.0"
import os
import sys
import pickle
from typing import List
import numpy as np
from jina.executors.devices import TorchDevice
from jina.excepts import PretrainedModelFileDoesNotExist
from jina.executors.decorators import batching_multi_input, as_ndarray
from jina.executors.encoders.multimodal import BaseMultiModalEncoder
# sys.path.append(".")
from img_text_composition_models import TIRG
class TirgMultiModalEncoder(TorchDevice, BaseMultiModalEncoder):
def __init__(self, model_path: str,
texts_path: str,
positional_modality: List[str] = ['visual', 'textual'],
channel_axis: int = -1,
*args, **kwargs):
"""
:param model_path: the path where the model is stored.
"""
super().__init__(*args, **kwargs)
self.model_path = model_path
self.texts_path = texts_path
self.positional_modality = positional_modality
self.channel_axis = channel_axis
# axis 0 is the batch
self._default_channel_axis = 1
def post_init(self):
super().post_init()
import torch
if self.model_path and os.path.exists(self.model_path):
with open (self.texts_path, 'rb') as fp:
texts = pickle.load(fp)
self.model = TIRG(texts, 512)
model_sd = torch.load(self.model_path, map_location=torch.device('cpu'))
self.model.load_state_dict(model_sd['model_state_dict'])
self.model.eval()
self.to_device(self.model)
else:
raise PretrainedModelFileDoesNotExist(f'model {self.model_path} does not exist')
def _get_features(self, data):
visual_data = data[(self.positional_modality.index('image'))]
if self.channel_axis != self._default_channel_axis:
visual_data = np.moveaxis(visual_data, self.channel_axis, self._default_channel_axis)
textual_data = data[(self.positional_modality.index('text'))]
visual_data = torch.stack(visual_data).float()
if self.on_gpu:
visual_data = visual_data.cuda()
textual_data = textual_data.cuda()
img_features = self.model.extract_img_feature(visual_data)
text_features = self.model.extract_text_feature(textual_data)
return self.model.compose_img_text_features(img_features, text_features)
@batching_multi_input
@as_ndarray
def encode(self, *data: 'np.ndarray', **kwargs) -> 'np.ndarray':
import torch
feature = self._get_features(*data).detach()
if self.on_gpu:
feature = feature.cpu()
feature = feature.numpy()
return feature
# -
encoder = TirgMultiModalEncoder(model_path, text_path, positional_modality = ['image', 'text'], channel_axis=1)
# ## Example, encode one single image
data = []
data.append([trainset.get_img(0)]) # image at position 0
data.append([texts[0]]) # text at position 1
encoded_multimodal = encoder.encode(data)
encoded_multimodal.shape
# ## Encode a batch of images
batch_size=32
fashion_200k_loader = trainset.get_loader(batch_size=batch_size)
for batch in fashion_200k_loader:
# use multimodal encoder
data = []
assert len(batch) == batch_size
batch_of_imgs = [item['source_img_data'] for item in batch]
batch_of_text = [item['source_caption'] for item in batch]
data.append(batch_of_imgs)
data.append(batch_of_text)
assert len(data) == 2
encoded_batch = encoder.encode(data)
assert len(encoded_batch) == batch_size
break
# ## Test
#
# 1. Ensure encoder works the same as the original model at instance level
# 2. Ensure encoder works the same as the original model at batch level
# Initialize the model
model = TIRG(texts, embed_dim)
model_sd = torch.load(model_path, map_location=torch.device('cpu'))
model.load_state_dict(model_sd['model_state_dict'])
_ = model.eval()
# Ensure encoded result is correct at instance level
for i in range(10):
print(f"testing multimodal encoder with img {i}")
# extract feature via jina encoder
data = []
data.append([trainset.get_img(i)]) # visual at position 0
data.append([texts[i]]) # textual at position 1
encoded = encoder.encode(data)
# extract image text feature
text_feature = model.extract_text_feature([texts[i]])
img = [trainset.get_img(i)]
img = torch.stack(img).float()
img_feature = model.extract_img_feature(img)
extracted = model.compose_img_text_features(img_feature, text_feature).cpu().detach().numpy()
assert encoded.shape == extracted.shape
assert encoded.all() == extracted.all()
# Ensure encoded result is correct at batch level
batch_size=64
fashion_200k_loader = trainset.get_loader(batch_size=batch_size)
for batch in tqdm(fashion_200k_loader):
print(f"testing multimodal encoder with batch size {batch_size}")
# use multimodal encoder
data = []
assert len(batch) == batch_size
batch_of_imgs = [item['source_img_data'] for item in batch]
batch_of_text = [item['source_caption'] for item in batch]
data.append(batch_of_imgs)
data.append(batch_of_text)
assert len(data) == 2
encoded_batch = encoder.encode(data)
# use the original model
batch_of_text_features = model.extract_text_feature(batch_of_text)
batch_of_imgs = torch.stack(batch_of_imgs).float()
batch_of_img_features = model.extract_img_feature(batch_of_imgs)
extracted_batch = model.compose_img_text_features(batch_of_img_features, batch_of_text_features).cpu().detach().numpy()
assert len(extracted_batch) == batch_size
assert extracted_batch.all() == encoded_batch.all()
break
|
multimodal-search-tirg/TirgMultimodalEncoder/multimodal-encoder.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# #### this script calculates morgan similarity features
# #### test compounds are compared to themselves and a number of other known odorants
import pandas as pd
import numpy as np
from rdkit import Chem
from rdkit.Chem import AllChem
from rdkit import DataStructs
from rdkit.ML.Descriptors import MoleculeDescriptors
# +
# get train, leaderboard and test CIDs
with open('CIDs.txt') as f:
content = f.readlines()
CIDs = list(content)
CIDs = [int(x) for x in CIDs]
# get smiles
smiles = pd.read_csv('all_smiles.csv', index_col=0) # load smiles if the file already exists
# +
# function to calculate the similarity features from Morgan fingerprints
# creates the fingerprints and calculates similarities
# Inputs:
# list of ids
# Morgan radius
# Returns:
# feature vector with size of len(cids) x number of features
def calulate_similarities(ids, radius):
ms = [Chem.MolFromSmiles(x) for x in smiles.smiles]
fps = [AllChem.GetMorganFingerprint(x,radius) for x in ms]
all_features =[]
for idx, cid in enumerate(ids):
ms_sample = Chem.MolFromSmiles(smiles.loc[cid].smiles)
fp_sample = AllChem.GetMorganFingerprint(ms_sample,radius)
features = [cid]
for fp in fps:
features.append(DataStructs.DiceSimilarity(fp,fp_sample))
all_features.append(features)
all_features = pd.DataFrame(all_features)
all_features = all_features.set_index(0)
all_features.columns = smiles.index
return all_features
# +
# get the similarity features
features_sim = calulate_similarities(CIDs, 5)
print(features_sim.shape)
# -
features_sim.head()
# save it
features_sim.to_csv('morgan_sim.csv')
|
opc_python/hulab/similarity_feature_calculation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import cvxpy as cvx
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from sklearn import linear_model as lm
# %matplotlib inline
# -
# # Exploratory data analysis
# Read in the *Engel food expenditure* data from the R package [`quantreg`](https://cran.r-project.org/package=quantreg).
engel = pd.read_csv('https://vincentarelbundock.github.io/Rdatasets/csv/quantreg/engel.csv', index_col=0)
engel.head()
# ## Scatter plot of income vs food expenditure
engel.plot.scatter(x='income', y='foodexp', color='darkblue')
# # Modelling
# Define model matrix and outcome.
X = np.hstack((
np.ones(engel.shape[0])[:, np.newaxis],
engel['income'][:, np.newaxis]
))
y = engel['foodexp'].values
X[:5, :]
y[:5]
# Define a variable `betas` for the regression coefficients and a parameter `tau` representing the quantile.
betas = cvx.Variable(X.shape[1])
tau = cvx.Parameter()
# Define variables `u` and `v` for the positive and negative deviations.
u = cvx.Variable(X.shape[0], nonneg=True)
v = cvx.Variable(X.shape[0], nonneg=True)
# Define the objective function and constraints.
objective = cvx.sum(tau * u) + cvx.sum((1-tau) * v)
constraints = [cvx.matmul(X, betas) + u - v == y]
# Set a value for $\tau$.
tau.value = 0.5
# Define and solve the quantile regression problem.
problem = cvx.Problem(cvx.Minimize(objective), constraints)
problem.solve()
# Extract the regression coefficients.
betas.value
# Define a new range of values to predict over.
n_pred = 100
X_pred = np.hstack((
np.ones(n_pred)[:, np.newaxis],
np.linspace(min(engel['income']), max(engel['income']), n_pred)[:, np.newaxis]
))
X_pred[:5, :]
y_pred = X_pred @ betas.value
y_pred[:5]
# Plot the original data and quantile regression line.
engel.plot.scatter(x='income', y='foodexp', color='darkblue')
plt.plot(X_pred[:, 1], y_pred, color='darkgreen')
# Compare with the linear regression (OLS) line.
betas_lr = lm.LinearRegression(fit_intercept=False).fit(X, y).coef_
engel.plot.scatter(x='income', y='foodexp', color='darkblue')
plt.plot(X_pred[:, 1], y_pred, color='darkgreen')
plt.plot(X_pred[:, 1], X_pred @ betas_lr, color='red')
|
notebooks/03_Quantile_Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <span style="font-family:Papyrus; font-size:3em;">Computational Experiments</span>
#
# <span style="font-family:Papyrus; font-size:2em;">Two Factors in Combination (TFC) Design</span>
# This lecture continues the analysis of the Wolf model of glycolytic oscillations to consider the effects of two parameters in combination.
# # Programming Preliminaries
IS_COLAB = False
# Code installs. Only needed once.
if IS_COLAB:
# !pip install matplotlib
# !pip install numpy
# !pip install pandas
# !pip install scipy
# !pip install tellurium
# !pip install seaborn
# Constants for standalone notebook
if not IS_COLAB:
DIR = "/home/ubuntu/advancing-biomedical-models/common" # Insert your local directory path
else:
from google.colab import drive
drive.mount('/content/drive')
DIR = "/content/drive/MyDrive/Winter 2021/common"
import sys
sys.path.insert(0, DIR)
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
import numpy as np
import pandas as pd
from scipy import fftpack
import tellurium as te
import seaborn as sns
import util_doe1 # Various functions from lecture 5
import wolf_model as wm
# # Design
# A TFC design is used to identify interactions between factors.
# This means that there is an experiment for every combination of levels of both factors.
# That is, there are $n_1 \times n_2$ experiments if there are $n_i$ levels of the two factors.
# ## Challenge: Reducing combinatorics
# A first challenge with a TFC is the number of constants that must be estimated.
# For example, the Wolf model has 16 parameters, which means that there are 120 interactions, each of which need to be calculated at several levels.
# With 10 levels for each interaction, this results in $10 \times 10 \times 120 = 12,000$ constants to calculate.
# ### Question
# 1. What are some approaches to reducing the number of constants to estimate in a TFC design?
# 1. What interaction terms should be estimated for the Wolf model?
# ## Challenge: Visualization
# ### Question
# 1. How should we present the responses that clearly presents which interaction terms are displayed, their values, and patterns in the responses?
# # Generalize runExperiments
# Generalize ``runExperiments`` so that it calculates responses for all combinations of levels of two parameters.
# +
def runExperimentsTwoParameters(parameter1, parameter2, percent1s, percent2s, isRelative=True):
"""
Runs experiments for changes in multipleparameters of the model at different percent changes in the parameter value (levels).
Parameter
---------
parameter1: str
parameter2: str
percent1s: list-int
percent2s: list-int
isRelative: bool
True: values are percent changes relative to baseline
False: absolute value
Returns
-------
frequencyDF: DataFrame, amplitude: DataFrame
index: tuple of percent changes in parameter values
columns: molecule
value: depends on isRelative
index.name: string of tuple (rowParameter, columnParameter)
"""
INDEX1 = "index1"
INDEX2 = "index2"
# Calculate the baseline values
baseFrequencySer, baseAmplitudeSer = util_doe1.runExperiment({})
#
def calcResponseSer(ser, isFrequency=True):
"""
Calculates the relative response.
Parameters
----------
ser: pd.Series
index: molecule
value: absolute respoinse
isFrequency: bool
if True, frequency response; else, amplitude response
Returns
-------
pd.Series
"""
if not isRelative:
return ser
if isFrequency:
baseSer = baseFrequencySer
else:
baseSer = baseAmplitudeSer
resultSer = 100*(ser - baseSer)/baseSer
return resultSer
#
def calcLevelDF(isFrequency=None):
"""
Calculates the dataframe of levels dataframe.
Parameter
--------
isFrequency: bool
If True, frequency response. Otherwise, amplitude response
Returns
-------
pd.DataFrame
index: tuple-int
levels of parameters
columns: str
molecule
values: response
"""
if isFrequency is None:
raise ValueError("Must specify isFrequency!")
sers = [] # Collection of experiment results
index1 = []
index2 = []
for percent1 in percent1s:
for percent2 in percent2s:
#indices.append("%d_%d" % (percent1, percent2))
index1.append(percent1)
index2.append(percent2)
parameterDct = {parameter1: percent1, parameter2: percent2}
frequencySer, amplitudeSer = util_doe1.runExperiment(parameterDct)
if isFrequency:
ser = frequencySer
else:
ser = amplitudeSer
adjSer = calcResponseSer(ser, isFrequency=isFrequency)
sers.append(pd.DataFrame(adjSer).transpose())
resultDF = pd.concat(sers)
resultDF[INDEX1] = index1
resultDF[INDEX2] = index2
resultDF = resultDF.set_index([index1, index2])
for column in [INDEX1, INDEX2]:
del resultDF[column]
resultDF.index.name = str((parameter1, parameter2))
return resultDF
#
frequencyDF = calcLevelDF(isFrequency=True)
amplitudeDF = calcLevelDF(isFrequency=False)
return frequencyDF, amplitudeDF
# Tests
percents = [-7, -5, -2, 0, 2, 5, 7]
fDF, aDF = runExperimentsTwoParameters("J1_Ki", "J1_k1", percents, percents)
assert(aDF.loc[(percents[-1], percents[-1]), "Glucose"] < 0 )
assert(fDF.loc[(percents[0], percents[0]), "Glucose"] < 0 )
# -
# Test Data
TEST_FDF = fDF
TEST_ADF = aDF
# ## Question
# 1. Display the amplitude response if there is no change in either J1_Ki or J1_k1.
# 1. Display the amplitude responses for J1_Ki is 0% and all changes in J1_k1.
# ## Solution
# +
# Solution 1
print("** Solution to 1:\n")
print(aDF.loc[(0,0), :])
# Solution 2
indices = [(0, p) for p in percents]
print("\n\n** Solution to 2:\n")
print(aDF.loc[indices, :])
# -
# # Visualizing results of an Experiment
# We will visualize the TFC reponses using a heatmap. The horizontal and vertical axes
# will be percent changes in parameter values; cells will be colored with the change in magnitude.
# ## Example of a heatmap in python
# The example below generates a heatmap from a pandas DataFrame.
# The horizontal axis are the column names; the vertical axis are the values of the index of the DataFrame.
# Values are encoded as a heat representation from 0 to 1.0.
# DataFrame
df = pd.DataFrame({"a": np.random.random(5), "b": np.random.random(5)})
df.index = [10*v for v in range(len(df))]
df
# Generating the heatmap
_ = sns.heatmap(df, cmap='seismic', vmin=0, vmax=1)
# ## Heat map for two parameters in combination
# ### Question
# 1. Generate a heatmap for the results of experiments varying the parameters "J1_k1" and "J1_Ki" in combination
# for levels -10, -5, 0, 5, 10.
# The names of molecules should be the rows.
# ### Solution
percents = [-10, -5, 0, 5, 10]
fDF, aDF = runExperimentsTwoParameters("J1_k1", "J1_Ki", percents, percents)
_ = sns.heatmap(aDF.transpose(), cmap='seismic', vmin=-100, vmax=100)
# ### Question
# 1. Create a function that transforms the frequency (or amplitude) DataFrame so that rows are values of parameter 1, columns are values of parameter 2,
# and cells are values of a designated molecule.
# ### Solution
# +
def pivotResponse(responseDF, molecule):
"""
Creates a DataFrame in which values of the parameters are rows and columns.
Parameters
----------
responseDF: DataFrame
columns: molecule names
rows: tuple of percent changes in parameters
values: response
index.name: string of tuple (rowParameter, columnParameter)
molecule: str
Returns
-------
DataFrame
columns: values of columnParameter
rows: values of rowParameter
cells: response for molecule
"""
LEVEL0 = "level_0"
LEVEL1 = "level_1"
df = responseDF.reset_index()
resultDF = pd.pivot_table(df, values=molecule, index=LEVEL0, columns=LEVEL1)
parameters = eval(responseDF.index.name)
resultDF.index.name = parameters[0]
resultDF.columns.name = parameters[1]
return resultDF
# Tests
df = pivotResponse(TEST_FDF, "Glucose")
#pd.pivot_table(df, values="Glucose", index="level_0", columns="level_1")
assert(len(df.index) == len(df.columns))
# -
# ### Question
# 1. Write a function that constructs a heatmap that displays interaction results for a single molecule (e.g., Glucose) and has rows that are changes to one parameter and columns that are changes to the other parameter. The function should return the axis.
# ### Solution
# +
def makeHeatmap(responseDF, molecule, isColorbar=True, ax=None, vmin=-100, vmax=100, cbar_ax=None):
"""
Creates a heatmap showing the interactions of response values for two parameters.
Parameters
----------
reponseDF: DataFrame
index: tuples of percent changes in parameter values
columns: molecule
values: response
molecule: str
isColorbar: bool
show the colorbar
vmin: float
minimum value in color bar
vmax: float
maximum value in color bar
cbar_ax: Axes
"""
df = pivotResponse(responseDF, molecule)
df = df.sort_index(ascending=False) # Show large values on top
if ax is None:
ax = sns.heatmap(df, cmap='seismic', vmin=vmin, vmax=vmax, cbar=isColorbar, cbar_ax=cbar_ax)
else:
_ = sns.heatmap(df, cmap='seismic', vmin=vmin, vmax=vmax, cbar=isColorbar, ax=ax, cbar_ax=cbar_ax)
return ax
# Tests
_ = makeHeatmap(TEST_FDF, "ATP")
# -
# # Running a Study
# ## Question
# 1. Write a function that generates an upper triangular matrix of plots labelled with the interactions between parameter pairs.
# ## Solution
# +
def runStudyTFC(molecule, parameters, percents, isRelative=True, isFrequency=True, figsize=(20, 10)):
"""
Creates an upper triangular plot of the interactions between parameter pairs in combinations.
Parameters
----------
molecule: str
parameters: list-str
percents: list-int
isRelative: bool
Results are percent change w.r.t. base
isFrequency: bool
Results are for frequencies
figisze: tuple-int
Size of figures
"""
numParameter = len(parameters)
fig = plt.figure(figsize=figsize)
gs = fig.add_gridspec(numParameter, numParameter)
cbar_ax = fig.add_axes([.91, .3, .03, .4])
for rowIdx in range(numParameter):
parmRowidx = rowIdx
for colIdx in range(numParameter-rowIdx-1):
parmColidx = numParameter - colIdx - 1
frequencyDF, amplitudeDF = runExperimentsTwoParameters(
parameters[parmRowidx], parameters[parmColidx],
percents, percents, isRelative=isRelative)
if isFrequency:
responseDF = frequencyDF
else:
responseDF = amplitudeDF
ax = plt.subplot(gs[rowIdx, colIdx])
# Label the parameters for each column
if rowIdx == 0:
ax.text(1, -0.4, parameters[parmColidx], fontsize=util_doe1.FONTSIZE)
# Only display colorbar for the last plot
if (rowIdx == numParameter - 2):
_ = makeHeatmap(responseDF, molecule, ax=ax, isColorbar=True, cbar_ax=cbar_ax)
else:
_ = makeHeatmap(responseDF, molecule, ax=ax, isColorbar=False)
ax.set_xlabel("")
# Only display ylabel for left-most plot
if colIdx == 0:
ax.set_ylabel(parameters[parmRowidx], fontsize=util_doe1.FONTSIZE)
else:
ax.set_ylabel("")
ax.set_yticklabels([])
# Only show x-tics for bottom-most plot
if colIdx != (numParameter - rowIdx - 2):
ax.set_xticklabels([])
if isFrequency:
responseType = "Frequency"
else:
responseType = "Amplitude"
if isRelative:
metricType = "relative"
else:
metricType = "absolute"
suptitle = "%s: %s response (%s)" % (molecule, responseType, metricType)
plt.suptitle(suptitle, fontsize=2*util_doe1.FONTSIZE)
# Smoke test
runStudyTFC("ATP", wm.WOLF_PRMS[0:3], [-5, 0, 5], isRelative=True, isFrequency=True)
# -
# ## Question
# 1. Analyze realtive frequencies for Glucose.
# ## Solution
# Analysis for frequency with relative (precentage) responses
percents = 3*(np.array(range(7)) - 3)
for molecule in wm.WOLF_MOL[0:1]:
for isFrequency in [True, False]:
runStudyTFC(molecule, wm.WOLF_PRMS[0:4], percents, isRelative=True, isFrequency=isFrequency)
# ## Question
# 1. Which combination of parameters for which response suggest a strong interaction between parameter pairs?# Analysis for frequency with relative (precentage) responses
|
archived_lectures/Winter 2021/lecture_6/6_computational_experiments_TFC.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Video Game Sales
#
# `<NAME> (<EMAIL>)`
#
# `<NAME> (<EMAIL>)`
# +
from IPython.display import display
import numpy as np
import pandas as pd
import warnings
warnings.filterwarnings('ignore')
from sklearn.neural_network import MLPClassifier
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.metrics import accuracy_score
from sklearn.preprocessing import OneHotEncoder, LabelEncoder
# -
df = pd.read_csv('VideoGamesSalesWithRatings.csv')
df.head()
df.info()
df
minor = df['Year_of_Release'].min()
major = df['Year_of_Release'].max()
print("Menor ano da base: %d"%(minor))
display(df[df['Year_of_Release'] == minor])
print("Maior ano da base: %d"%(major))
display(df[df['Year_of_Release'] == major])
df.drop(labels=5936,inplace=True)
major = df['Year_of_Release'].max()
print("Maior ano da base: %d"%(major))
display(df[df['Year_of_Release'] == major])
# O jogo `Imagine: Makeup Artist` possui vendas espalhadas pelo mundo todo mesmo antes de seu lançamento, isso nos levou a pensar que fosse um tipo de pre-venda, porém após algumas pesquisas constatamos que o jogo foi lançado em 2009 e não possuia nenhuma previsão de lançamento para 2020, por isso tudo indica que seja um dado ruidoso, nos levando a questionar se os outros valores referentes a ele estariam realmente corretos. Optamos então por deletar esse valor do nosso dataset.
df.drop(labels=[14086,16222,16385],inplace=True)
major = df['Year_of_Release'].max()
print("Maior ano da base: %d"%(major))
display(df[df['Year_of_Release'] == major])
# Os jogos `Phantasy Star Online 2 Episode 4: Deluxe Package` de PS4 e PSV e `Brothers Conflict: Precious Baby` foram lançados em 2016 apenas no Japão e tinham como 2017 o ano de lançamento no resto do mundo. Porém, como a base data de 22 de Dezembro de 2016, não foi computado as vendas no resto do mundo, nem nota da crítica e etc. Sendo assim, optamos por deletar esses valores do nosso dataset.
df.columns
# +
qtdUserScoreTbd = len(list(filter(lambda x : x is True, [x == 'tbd' for x in df['User_Score']])))
display("Quantidade de valores \"tbd\" na coluna User_Score: %d"%(qtdUserScoreTbd))
df['User_Score'].replace('tbd', np.nan, inplace=True)
qtdUserScoreNan = len(list(filter(lambda x : x is True, [x is np.nan for x in df['User_Score']])))
display("Quantidade de valores \"NaN\" na coluna User_Score após a converção: %d"%(qtdUserScoreNan))
df['User_Score'] = df['User_Score'].apply(lambda x : float(x))
df.info()
# -
# A coluna `User_Score` possuia valores igual a `tbd` que no Metacritic significa "to be determined". Ou seja, é quando não houve o número mínimo de 4 avaliações para que o Metacritic pode-se considerar a avaliação média dos usuários. Sendo assim, resolvemos considerar que os valores iguais a `tbd` equivalem a `NaN`, visto que os valores correspondentes a `User_count` nesses registros está como `NaN`, o que implíca dizer que não houve usuários avaliadores.
#
# Após a substituição de valores, convertemos o tipo da coluna `User_Score` de categórico para numérico
# +
qtdAnosNan = len(df[np.isnan(df['Year_of_Release'])])
display("Quantidade de valores \"NaN\" na coluna Year_of_Release: %d"%(qtdAnosNan))
qtdNASalesNan = len(df[np.isnan(df['NA_Sales'])])
display("Quantidade de valores \"NaN\" na coluna NA_Sales: %d"%(qtdNASalesNan))
qtdEUSalesNan = len(df[np.isnan(df['EU_Sales'])])
display("Quantidade de valores \"NaN\" na coluna EU_Sales: %d"%(qtdEUSalesNan))
qtdJPSalesNan = len(df[np.isnan(df['JP_Sales'])])
display("Quantidade de valores \"NaN\" na coluna JP_Sales: %d"%(qtdJPSalesNan))
qtdOtherSalesNan = len(df[np.isnan(df['Other_Sales'])])
display("Quantidade de valores \"NaN\" na coluna Other_Sales: %d"%(qtdOtherSalesNan))
qtdGlobalSalesNan = len(df[np.isnan(df['Global_Sales'])])
display("Quantidade de valores \"NaN\" na coluna Global_Sales: %d"%(qtdGlobalSalesNan))
qtdCriticScoreNan = len(df[np.isnan(df['Critic_Score'])])
display("Quantidade de valores \"NaN\" na coluna Critic_Score: %d"%(qtdCriticScoreNan))
qtdCriticCountNan = len(df[np.isnan(df['Critic_Count'])])
display("Quantidade de valores \"NaN\" na coluna Critic_Count: %d"%(qtdCriticCountNan))
qtdUserScoreNan = len(df[np.isnan(df['User_Score'])])
display("Quantidade de valores \"NaN\" na coluna User_Score: %d"%(qtdUserScoreNan))
qtdUserCountNan = len(df[np.isnan(df['User_Count'])])
display("Quantidade de valores \"NaN\" na coluna User_Count: %d"%(qtdUserCountNan))
# +
qtdNameNan = len(list(filter(lambda x : x is True, [x is np.nan for x in df['Name']])))
display("Quantidade de valores \"NaN\" na coluna Name: %d"%(qtdNameNan))
qtdPlatformNan = len(list(filter(lambda x : x is True, [x is np.nan for x in df['Platform']])))
display("Quantidade de valores \"NaN\" na coluna Platform: %d"%(qtdPlatformNan))
qtdGenreNan = len(list(filter(lambda x : x is True, [x is np.nan for x in df['Genre']])))
display("Quantidade de valores \"NaN\" na coluna Genre: %d"%(qtdGenreNan))
qtdPublisherNan = len(list(filter(lambda x : x is True, [x is np.nan for x in df['Publisher']])))
display("Quantidade de valores \"NaN\" na coluna Publisher: %d"%(qtdPublisherNan))
qtdDeveloperNan = len(list(filter(lambda x : x is True, [x is np.nan for x in df['Developer']])))
display("Quantidade de valores \"NaN\" na coluna Developer: %d"%(qtdDeveloperNan))
qtdRatingNan = len(list(filter(lambda x : x is True, [x is np.nan for x in df['Rating']])))
display("Quantidade de valores \"NaN\" na coluna Rating: %d"%(qtdRatingNan))
# +
df = df.dropna(subset = ['Name', 'Genre', 'Publisher', 'Year_of_Release'])
qtdNameNan = len(list(filter(lambda x : x is True, [x is np.nan for x in df['Name']])))
qtdGenreNan = len(list(filter(lambda x : x is True, [x is np.nan for x in df['Genre']])))
qtdPublisherNan = len(list(filter(lambda x : x is True, [x is np.nan for x in df['Publisher']])))
qtdAnosNan = len(df[np.isnan(df['Year_of_Release'])])
display("Quantidade após drop desses valores \"NaN\" na coluna Name: %d"%(qtdNameNan))
display("Quantidade após drop desses valores \"NaN\" na coluna Genre: %d"%(qtdGenreNan))
display("Quantidade após drop desses valores \"NaN\" na coluna Publisher: %d"%(qtdPublisherNan))
display("Quantidade após drop desses valores \"NaN\" na coluna Year_of_Release: %d"%(qtdAnosNan))
# +
qtdCriticScoreNan = len(df[np.isnan(df['Critic_Score'])])
display("Quantidade de valores \"NaN\" na coluna Critic_Score: %d"%(qtdCriticScoreNan))
qtdCriticCountNan = len(df[np.isnan(df['Critic_Count'])])
display("Quantidade de valores \"NaN\" na coluna Critic_Count: %d"%(qtdCriticCountNan))
qtdUserScoreNan = len(df[np.isnan(df['User_Score'])])
display("Quantidade de valores \"NaN\" na coluna User_Score: %d"%(qtdUserScoreNan))
qtdUserCountNan = len(df[np.isnan(df['User_Count'])])
display("Quantidade de valores \"NaN\" na coluna User_Count: %d"%(qtdUserCountNan))
df['Critic_Score'].replace(np.nan, 0, inplace=True)
df['Critic_Count'].replace(np.nan, 0, inplace=True)
df['User_Score'].replace(np.nan, 0, inplace=True)
df['User_Count'].replace(np.nan, 0, inplace=True)
display("Após substituição dos valores \"NaN\" para \"0\" temos:")
qtdCriticScoreZero = len(list(filter(lambda x : x is True, [x == 0 for x in df['Critic_Score']])))
display("Quantidade de valores \"0\" na coluna Critic_Score: %d"%(qtdCriticScoreZero))
qtdCriticCountZero = len(list(filter(lambda x : x is True, [x == 0 for x in df['Critic_Count']])))
display("Quantidade de valores \"0\" na coluna Critic_Count: %d"%(qtdCriticCountZero))
qtdUserScoreZero = len(list(filter(lambda x : x is True, [x == 0 for x in df['User_Score']])))
display("Quantidade de valores \"0\" na coluna User_Score: %d"%(qtdUserScoreZero))
qtdUserCountZero = len(list(filter(lambda x : x is True, [x == 0 for x in df['User_Count']])))
display("Quantidade de valores \"0\" na coluna User_Count: %d"%(qtdUserCountZero))
# +
qtdDeveloperNan = len(list(filter(lambda x : x is True, [x is np.nan for x in df['Developer']])))
display("Quantidade de valores \"NaN\" na coluna Developer: %d"%(qtdDeveloperNan))
def teste(x):
if((x['Developer'] is np.nan) and (x['Publisher'] is not np.nan)):
x['Developer'] = x['Publisher']
return x
df[['Developer', 'Publisher']] = df[['Developer', 'Publisher']].apply(teste, axis=1)
qtdDeveloperNan = len(list(filter(lambda x : x is True, [x is np.nan for x in df['Developer']])))
display("Quantidade de valores \"NaN\" na coluna Developer após substituição: %d"%(qtdDeveloperNan))
qtdRatingNan = len(list(filter(lambda x : x is True, [x is np.nan for x in df['Rating']])))
display("Quantidade de valores \"NaN\" na coluna Rating: %d"%(qtdRatingNan))
df['Rating'].replace(np.nan, 'E', inplace=True)
qtdRatingNan = len(list(filter(lambda x : x is True, [x is np.nan for x in df['Rating']])))
display("Quantidade de valores \"NaN\" na coluna Rating após substituição: %d"%(qtdRatingNan))
# -
# ### Tratamento de valores "NaN"
#
# Testamos se havia valores `NaN`em todas as colunas do dataframe e vimos que as seguintes colunas possuiam as seguintes quantidades desses valores ruidosos:
# - Year_of_Release: 269
# - Critic_Score: 8578
# - Critic_Count: 8578
# - User_Score: 9125
# - User_Count: 9125
# - Name: 2
# - Genre: 2
# - Publisher: 54
# - Developer: 6620
# - Rating: 6766
#
# Como as colunas `Year_of_Release`, `Name`, `Genre` e `Publisher` possuem poucos valores ruidosos comparado ao total de dados do dataframe (16.715 linhas), optamos por dropar esses registros da base.
#
# Já as colunas `Critic_Score`, `Critic_Count`, `User_Score` e `User_Count` possuiam mais de 8000 registros ruidosos. Sendo assim optamos por substituir os valores `NaN` por `0`, pois consideramos que `Critic_Count` e `User_Count` com valores `NaN` significa que nenhum usuário avaliou aquele jogo e por isso o `Critic_Score` e `User_Score` possuiriam nota 0.
#
# Além disso, como `Rating` é uma coluna de valores categóricos e possuia mais de 6000 registros ruidosos. Neste caso, consideramos que um registro de rating `NaN` seria equivalente ao rating `E`.
#
# Por fim, na coluna `Developer` consideramos que os registros que possuiam `NaN` como valor seria substituido pelo valor da `Publisher`.
df['Year_of_Release'] = df['Year_of_Release'].apply(lambda x : int(x))
df.drop(labels=['Critic_Count', 'User_Count'], axis=1, inplace=True)
df
# Convertemos os valores da coluna `Year_of_Release` de float para int.
# Consideramos que as colunas `Critic_Count` e `User_Count` são irrelevantes dentre as hipóteses levantadas, e por isso decidimos dropálas da base.
# ## Normalização
#
# Normalizamos os valores de vendas de acordo com o maior valor de cada coluna em específica `value/maxValue`.
max_NA_Sales = df['NA_Sales'].max()
print(max_NA_Sales)
max_EU_Sales = df['EU_Sales'].max()
print(max_EU_Sales)
max_JP_Sales = df['JP_Sales'].max()
print(max_JP_Sales)
max_Other_Sales = df['Other_Sales'].max()
print(max_Other_Sales)
max_Global_Sales = df['Global_Sales'].max()
print(max_Global_Sales)
for i in df.columns[5:]:
if(i == 'NA_Sales'):
df[i] = df[i].apply(lambda x : x/max_NA_Sales)
elif(i == 'JP_Sales'):
df[i] = df[i].apply(lambda x : x/max_JP_Sales)
elif(i == 'EU_Sales'):
df[i] = df[i].apply(lambda x : x/max_EU_Sales)
elif(i == 'Other_Sales'):
df[i] = df[i].apply(lambda x : x/max_Other_Sales)
elif(i == 'Global_Sales'):
df[i] = df[i].apply(lambda x : x/max_Global_Sales)
df
print("qtd possíveis valores para plataforma: {}".format(len(df['Platform'].value_counts())))
print("qtd possíveis valores para genero: {}".format(len(df['Genre'].value_counts())))
print("qtd possíveis valores para publisher: {}".format(len(df['Publisher'].value_counts())))
print("qtd possíveis valores para developer: {}".format(len(df['Developer'].value_counts())))
print("qtd possíveis valores para rating: {}".format(len(df['Rating'].value_counts())))
# +
# gen_le = LabelEncoder()
# gen_labels = gen_le.fit_transform(df['Genre'])
# df['Genre_Labels'] = gen_labels
# plat_le = LabelEncoder()
# platform_labels = plat_le.fit_transform(df['Platform'])
# df['Platform_Labels'] = platform_labels
# rat_le = LabelEncoder()
# rating_labels = rat_le.fit_transform(df['Rating'])
# df['Rating_Labels'] = rating_labels
# df_sub = df[['Name', 'Genre', 'Genre_Labels',
# 'Platform', 'Platform_Labels','Rating','Rating_Labels']]
# df_sub
df = df.join(pd.get_dummies(df.Platform), how='outer', lsuffix='_left', rsuffix='_right')
df = df.join(pd.get_dummies(df.Genre), how='outer', lsuffix='_left', rsuffix='_right')
df = df.join(pd.get_dummies(df.Publisher), how='outer', lsuffix='_left', rsuffix='_right')
df = df.join(pd.get_dummies(df.Rating), how='outer', lsuffix='_left', rsuffix='_right')
display(df.head())
display(len(df))
display(len(df.columns))
# +
# gen_ohe = OneHotEncoder()
# gen_feature_arr = gen_ohe.fit_transform(df[['Genre_Labels']]).toarray()
# gen_feature_labels = list(gen_le.classes_)
# gen_features = pd.DataFrame(gen_feature_arr, columns=gen_feature_labels)
# plat_ohe = OneHotEncoder()
# plat_feature_arr = plat_ohe.fit_transform(df[['Platform_Labels']]).toarray()
# plat_feature_labels = list(plat_le.classes_)
# plat_features = pd.DataFrame(plat_feature_arr, columns=plat_feature_labels)
# rat_ohe = OneHotEncoder()
# rat_feature_arr = rat_ohe.fit_transform(df[['Rating_Labels']]).toarray()
# rat_feature_labels = list(rat_le.classes_)
# rat_features = pd.DataFrame(rat_feature_arr, columns=rat_feature_labels)
# df = pd.concat([df.reset_index(drop=True), gen_features.reset_index(drop=True),plat_features.reset_index(drop=True), rat_features.reset_index(drop=True)], axis=1)
# df
# -
df.to_csv(r'vgsalesPP2.csv')
|
PreProcessamento.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
from tensorforce.environments import Environment
from tensorforce.agents import Agent
from tensorforce.execution import Runner
# + pycharm={"name": "#%%\n"}
train_env_kwargs = {'filename':'sp500.csv',
'date_from':'2008-01-01',
'date_to':'2017-12-31',
'investment':1000000,
'risk_free_rate': 0.5, # approx US Treasury Note return
'sample_size':100,
#'report_point':252,
'random_sample':True,
'reward_function':'portfolio_value'}
train_portfolio = Environment.create(
environment='PortfolioTF',
**train_env_kwargs
)
_ = train_portfolio.reset()
total_timesteps = 1 * (len(train_portfolio._environment.data.date.unique())-1)
# + pycharm={"name": "#%%\n"}
vanilla = Agent.create(
agent='reinforce',
environment=train_portfolio,
max_episode_timesteps=total_timesteps,
network=dict(type='auto',
rnn=10 ),
batch_size=5
)
# + pycharm={"name": "#%%\n"}
runner = Runner(
agent=vanilla,
environment=train_portfolio
)
# + pycharm={"name": "#%%\n"}
print(runner.agent.get_architecture())
# + pycharm={"name": "#%%\n"}
runner.agent.get_specification()
# + pycharm={"name": "#%%\n"}
runner.run(num_episodes=10)
# + pycharm={"name": "#%%\n"}
runner.run(num_episodes=10, evaluation=True)
# + pycharm={"name": "#%%\n"}
timesteps = len(train_portfolio._environment.data.date.unique())-1
random = Agent.create(
agent='random',
environment=train_portfolio,
max_episode_timesteps=timesteps,
)
# + pycharm={"name": "#%%\n"}
randomrunner = Runner(
agent=random,
environment=train_portfolio
)
# + pycharm={"name": "#%%\n"}
randomrunner.run(num_episodes=10)
# + pycharm={"name": "#%%\n"}
total_timesteps = 1000 * (len(train_portfolio._environment.data.date.unique())-1)
a2cagent = Agent.create(
agent='a2c',
environment=train_portfolio,
max_episode_timesteps=total_timesteps,
network=dict(type='auto',
rnn=252 ),
critic_optimizer=dict(optimizer='adam'),
batch_size=5
)
# + pycharm={"name": "#%%\n"}
a2crunner = Runner(
agent=a2cagent,
environment=train_portfolio
)
# + pycharm={"name": "#%%\n"}
print(a2crunner.agent.get_architecture())
# + pycharm={"name": "#%%\n"}
a2crunner.agent.get_specification()
# + pycharm={"name": "#%%\n"}
a2crunner.run(num_episodes=1000)
# + pycharm={"name": "#%%\n"}
a2crunner.run(num_episodes=1000, evaluation=True, save_best_agent='.' )
##%
# + pycharm={"name": "#%%\n"}
test_env_kwargs = {'filename':'sp500.csv',
'date_from':'2017-01-01',
'date_to':'2019-12-31',
'investment':1000000,
'risk_free_rate': 0.5, # approx US Treasury Note return
'sample_size':100,
#'report_point':252,
'random_sample':False,
'reward_function':'portfolio_value'}
test_portfolio = Environment.create(
environment='PortfolioTF',
**test_env_kwargs
)
_ = test_portfolio.reset()
test_timesteps = 1 * (len(test_portfolio._environment.data.date.unique())-1)
# + pycharm={"name": "#%%\n"}
testa2c=Agent.load(directory='.',filename='best-model',environment=test_portfolio)
# + pycharm={"name": "#%%\n"}
test2crunner = Runner(
agent=testa2c,
environment=test_portfolio
)
# + pycharm={"name": "#%%\n"}
print(test2crunner.agent.get_architecture())
test2crunner.agent.get_specification()
# + pycharm={"name": "#%%\n", "is_executing": true}
test2crunner.run(num_episodes=10)
# + pycharm={"name": "#%%\n"}
a2crunner.run(num_episodes=1000, evaluation=True )
|
Tensorflow/TFAzure.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.3
# language: julia
# name: julia-1.5
# ---
# # A Short introduction to Julia
# ## Variables, Numbers, and Strings
# ### Variables and numerical types
# Variables do not need to be declared
x = 3
# but the value they are *bound to* has a type
typeof(x)
# We can re-assign ```x``` to another value of a different type
x = 3.4
typeof(x)
# In addition to ```Int64``` (integers) and ```Float64``` (double-precision floating point), we also have others numeric types, like rational and complex numbers:
x = 3//4
typeof(x)
x = 4.1 + 3.4im
typeof(x)
# The common operations works on all numerical types:
3//4 + 7//9
(4.2 + 3.4im)*(7 - 2im)
# Exponentiation works with ```^```:
2^10
# If we need integer division we can use $\div$
13 ÷ 5
# The reminder is, as in many other languages, ```%```:
13 % 5
# ### Characters and Strings
# The ```Char``` type is a single UTF-8 character represented in the code by enclosing it in single quotes
'a'
'🐱'
# A string is a (finite) sequence of characters and they are represented in the code by enclosing it in either double quotes or triple double quotes. Strings can span multiple lines
"Hello World"
"""Hello
World"""
"""Hello "Cruel" World"""
# Strings can be concatenated with ```*``` (and not ```+``` like in other languages)
"Hello" * " " * "World"
# Strings can be interpolated by using ```$```:
x = 4
"The value of x is $x"
"The result of x + 3 is $(x + 3)"
# Other kinds of Strings literals exists (e.g., binary, regular expressions):
b"binary values" # An array of unsigned 8 bits integers
r"[a-z]+\s*" # A regular expression
# ### Symbols
# A special kind of type is the ```Symbol``` type:
Symbol("hello")
typeof(:foo)
# Symbols are *interned strings*. You can use them, for example, as keywords (no need to use a String), but they have many other uses.
#
# Symbols should be familiar to people programming in Lisp.
# ## Array, Dictionaries, and Tuples
# ### Arrays
# Arrays can be defined similary to languages like Python by encosing the values in square brackets
v = [2,4,6,8]
# A one dimensional array of ```Int64```.
#
# We can obtain the type of the elements contained in an array with ```eltype```
eltype(v)
# and the length of the array by using ```length```
length(v)
# Arrays can be of mixed types (might not be the best for efficiency)
v2 = [1, 3.4, "Hello"]
# In Julia, array are by default indexed from ```1``` and not from ```0```
v[1]
v[length(v)]
# We can actually use any starting index with the package ```OffsetArrays.jl```
#
# We can also retrieve the first and last element of the array using ```begin``` and ```end``` as indexes:
v[begin]
v[end]
# #### Slices and viewes
#
# As with many modern languages, arrays can be sliced using the syntax ```start_index:end_index```
v[2:3]
# Slices can be assigned to variables. In that case a *copy* is made
w = v[2:3]
w[1] = 0
v
# If we want to refer to the same underlying memory we use ```view```
z = view(v,2:3)
z[1] = 0
v
# #### Broadcasting
# We may want to assign the same value to an entire slice of an array.
#
# However, this will give an error:
v[1:3] = 4
# We can prepend ```.``` to assignements (and any function call) to "broadcast" a value:
v[1:3] .= 4
v
v
# #### Multidimensional arrays
#
# Julia supports multi-dimensional array.
#
# We can write them remembering the following rules:
#
# - Arguments separated by semicolons ```;``` or newlines are vertically concatenated
# - Arguments separated by spaces or tabs are horizontally concatenated
M = [0 1 0 0
1 0 0 1
0 0 1 1]
# We can obtain the dimensions of a multi-dimensional array using the ```ndims``` function
ndims(M)
# And the length of a specific dimension using the ```size``` function
size(M)
# Indexing is made by using all indexes, separated by commas, inside square brackets:
M[1,2]
# Notice that multidimensional arrays and arrays-of-arrays are different!
M2 = [[0,1,0,0], [1,0,0,1], [0,0,1,1]]
# As before, we can slice the arrays. This time along multiple dimensions:
M[1:2,2:4]
M[2:3,:]
M[:,2:4]
# As before, we can create views of the array
A = view(M, 2:3, 2:4)
A[1,1] = 99
M
# And the shape of an array can be changed via the function ```reshape```:
reshape(M, 4, 3)
reshape(M, 2, 3, 2)
zeros(2,4,3)
# #### Array Comprehension
#
# As with list comprehension in languages like Python, Julia also supports array comprehension
squares = [i^2 for i in 1:10]
mult_table = [i * j for i in 1:5, j in 1:10]
# ### Dictionaries
#
# Dictionaries represent maps from a set $K$ of keys to a set $V$ of values
d = Dict('a' => 1, 2 => "text", :foo => :bar)
# Dictionaries can be indexed similarly to arrays
d['a']
# Assignment for non existing keys is possible and will add the key to the dictionary
d['z'] = 26
d
# The keys and values of a dictionary can be obtained by using the functions ```keys``` and ```values```, respectively:
keys(d)
values(d)
# ### Tuples
#
# Tuples are fixed-length containers that cannot be modified (i.e., they are immutable).
tuple = (1.2, 2, "hello")
typeof(tuple)
# Tuples are indexed starting from $1$, like arrays
tuple[1]
# For tuples consisting of only one element remember the comma at the end:
small_tuple = (3,)
# When created, tuples can also be *named*
named_tuple = (a = 1.2, b = 2, c = "hello")
# The elements of the tuple can now be accessed using the names (in addition to the indexes) of the fields
named_tuple.b
# When assigning values *from* a tuple, Julia supports some destructuring
x, y, z = tuple
y
# This is useful when a function returns multiple outputs as a tuple
# ## Functions
# ### Named functions
# Functions can be defined via the ```function``` keyword and terminated by ```end```:
function add_one_v1(x)
return x + 1
end
# The ```return``` keyword is not essential, the last value in the function is the one returned
function add_one_v2(x)
x + 1
end
# Simple functions can be defined in a single line without using the ```function``` keyword
add_one_v3(x) = x + 1
# Functions can also return multiple values as a tuple
function add_and_mul(x, y)
x + y, x * y
end
a, m = add_and_mul(3, 4)
# #### Keyword arguments
# Keyword arguments can be added after a semicolon in the argument list with a default value (in this case they are not mandatory) or without (in this case they are mandatory)
function add_constant(x; constant = 0)
x + constant
end
add_constant(3)
# Functions can have a variable number of arguments. You can use ```...``` following a variable name to collect the remaining arguments in an array.
function varargs(x...)
sum(x)
end
varargs(1,2,3)
# ### Anonymous (lambda) functions
# Sometimes it is useful to define a function without giving it a name. We can use anonymous (lambda) functions
(x,y) -> x^2 + y^2
# Functions can be assigned to variables and called like "normal" functions
ff = (x,y) -> x^2 + y^2
ff(2, 3)
typeof(ff)
# ### Broadcasting functions
# Function call is among the operations that can be braodcasted using ```.```:
ff([1,2,3], [4,5,6]) # This gives an error
ff.([1,2,3], [4,5,6])
# Since also the normal arithmetical operations are functions, they too can be broadcasted:
[1,2,3,4] .+ 1
# ## Selection and Iteration
# ### Selection: ```if```, ```if```..```else```
# Selection is performed, as usual, with the ```if``` construct
x = 2
if x < 3
println("x is less than 3")
end
# Multiple conditions can be checked by combining them in a C-like way
if x < 3 && x % 2 == 0
println("x even and less than 3")
end
# As usual, an ```if``` statement can include an ```else``` part
if x < 3
println("x is less than 3")
else
println("x is at least 3")
end
# Multiple ```if``` statements can be nested with ```elseif```
if x < 3
println("x is less than 3")
elseif x == 3
println("x is exactly 3")
else
println("x is more than 3")
end
# ### Iteration: ```for``` and ```while```
# Iteration has the usual ```for``` and ```while``` constructs
x = 1
while x ≤ 3
println(x)
x += 1
end
# The ```for``` construct operates similarly to the ```for``` in Python. This means that we usually have ```for variable in container```.
for v in 1:5
println(v)
end
# It is actually possible to use $\in$ instead of ```in```
for v ∈ [1,2,3]
println(v)
end
# ### Functional constructs
# Some common operations can better be expressed as functional constructs (e.g., ```map```, ```filter```) or array comprehension instead of using loops directly
# ```map``` applies a _unary_ function to all the elements of a container
map(x -> x^2 + x, [1,2,3,4,5])
# ```reduce``` applies a _binary_ function to all elements of a container "merging" them in a single value
v = [4,5,6,7,3]
reduce(+, v)
# Some kinds of reductions are so common that they have special functions defined
minimum(v)
maximum(v)
sum(v)
prod(v)
# Another common operation is to filter a collection taking only the elements respecting a certain condition
filter(x -> x ≥ 5, v)
# Many complex operations can be carried by combining filtering, mapping, and reducting.
#
# For example, let us compute the sum of the square of all the positive values inside a vector
v = rand(10) .- 0.5
v1 = filter(x -> x ≥ 0, v)
v2 = map(x -> x^2, v1)
v3 = sum(v2)
# This application of functions where the output of one goes as input to the next is so common that a special idiom using ```|>``` exists. Similar operators exist, for example, in R and Clojure.
filter(x -> x ≥ 0, v) |> x -> x.^2 |> sum
# In general the form ```x |> f``` applies ```f``` to ```x```, so ```x |> f |> g |> h``` is ```h(g(f(x)))```
# ## Types
# Until now we have not investigated how types can be useful for functions. We will start by defining how type annotations works
# Some types: ```Float64```, ```Rational```, ```Int64```, ```Number```, ```Real```, etc. The types have a hierarchy: for example ```Number``` includes all numeric types, while ```Real``` all real numbers, and so on.
function f_only_reals(x::Real, y::Real)
x + y
end
# Now a call like this works:
f_only_reals(3.5,4.0)
# While using a string will give an error
f_only_reals("hello", "world") # This gives an error
# We can also annotate variables to force them to only contains values of a specific type
function g(x, y)
v::Array{Int64,1} = [x,y]
sum(v)
end
g(1,4)
# Notice that some types have a parameter enclosed in curly brackets, like ```Array{Int64,1}``` or ```Rational{Int64}```.
#
# They are type parameters, they have some similarities to templates in C++ or generics in Java.
# It is possible to define union of types. For example if a parameter can be either a ```Float64``` or ```Int32``` we can define:
Union{Float64, Int32}
# A useful type to define missing values is the ```Missing``` type, whose only value is ```missing```.
#
# For example to define a function that accepts a one-dimensional array of ```Float64``` values or missing values we can define it using a union.
function f_missing(v::Array{Union{Float64,Missing},1})
filter(x -> !ismissing(x), v) |> sum
end
f_missing([3.5, missing, 5, 8, missing, -3])
missing + 4
# ### Structures
# We can define new (product) types by defining structures:
struct MyPair
first
second
end
# We have now defined a new type (```MyComplex```) that can be instantiated and used as a "normal" type:
c = MyPair(3, 5)
typeof(c)
# We can access the different fields by using the dot notation:
c.first
# Notice that we **cannot** modify a struct, since by default they are _immutable_:
c.second = 4 # this gives an error
# If we really need a _mutable_ structure we must define it so
mutable struct MyMutablePair
first
second
end
p = MyMutablePair(4, 7)
p.second = 8
p
# We can define a "template" structure by using a type parameter
struct MySameTypePair{T}
first::T
second::T
end
MySameTypePair(3,4)
MySameTypePair{String}("Hello", "World")
MySameTypePair(2, "hello") # This gives an error
# Fields of a structure can be typed independently.
struct MyPairV2
first::String
second::Float64
end
# ## Multiple Dispatch
# Why are types useful?
#
# - They provide a way to give an error if the type in the annotation is not respected
# - They can be used for dispatch! (see [The Unreasonable Effectiveness of Multiple Dispatch by <NAME>](https://www.youtube.com/watch?v=kc9HwsxE1OY))
# Until now we have defined a function only once, we might want to define it multiple times depending on the types of the parameters
# +
function h(x::Float64, y::Float64)
println("x and y are float")
end
function h(x::Float64, y::Int64)
println("x is float and y is integer")
end
function h(x::Int64, y::Float64)
println("x is integer and y is float")
end
function h(x::Int64, y::Int64)
println("x and y are integer")
end
# -
h(3.4, 2)
h(3, 2.7)
# We can see how many _methods_ every function has:
methods(h)
# We can see it even for built-in functions
methods(+)
# Why is multiple dispatch useful?
#
# We can write a function using, for example, ```+``` and the correct implementation will be the one used.
#
# For example we can write a function working with matrix multiplication and the correct implementation (depending if the matrix is, for example, diagonal, tridiagonal, etc) will be used.
# ## Environments and Packages
# It is essential to be able to create multiple environments, each one with a different set of packages installed and to make the environment reproducible across multiple machines.
#
# We can either work directly with the ```Pkg``` package (you can import it with ```using Pkg```) or on the REPL by going into the package management with ```]```
# - ```activate``` allows to create a (or activate an existing) new self-contained project. All installs will be local to the project. The ```Manifest.toml``` and ```Project.toml``` files contains the information about the installed packages and their versions
# - ```add package_name``` install the package, resolving the dependencies. For example, ```add IJulia``` to have a notebook interface.
# - ```rm package_name``` to remove a package from the current project
# - ```update``` to update the packages
# All code that we will see in the course can be installed as packages or it is in the base library.
# ## Macros
# Macros are a tool for _metaprogramming_. That is, for writing code that writes code. They are well known in the Lisp word and are gaining popularity in other languages.
#
# Recall that the arguments of a function are evaluated before being passed to a function.
#
# Arguments to a macro are not evaluated. The macro receives the (representation of the) code and can manipulate it returning new code.
#
# Macro should not be used too estensively, but they are a powerful tool. In Julia they always have ```@``` before their name
# #### Some useful macros
# The ```@time``` macro allows to find the amount of time spent in executing a piece of code
@time map(x->x^2, rand(1000));
# It is possible to explore which method of a function Julia is going to call using the ```@which``` macro
@which 3 + 5
# It is actually possible to observe which is the code generate by Julia with a collection of macros ```@code_lowered```, ```@code_typed```, ```@code_llvm```, ```@code_native```
@code_lowered 3 + 5
@code_typed 3 + 5
@code_llvm 3 + 5
@code_native 3 + 5
# We can see how a macro acts on the code via ```@macroexpand```
@macroexpand @timed 3 + 5
# Macros are one of the features that allows Julia to be flexible.
# #### Code representation
# Can can be forced not be evaluated by putting it in a ```quote``` block or by using the ```:``` operator
# +
code1 = quote
x = 4
y = 6
x + y
end
code2 = :(3 + 5)
code1
# -
# We can manipulate code as data (a concept that should be familiar to programmers in the lisp family of languages)
typeof(code2)
Meta.show_sexpr(code2)
code2.head
code2.args
# The code starts with a function call (```:call```) where the function is ```+``` and the arguments are ```3``` and ```5```.
#
# We can modify the code directly
code2.args[1] = :* # We modify the function from + to *
code2
eval(code2)
# ## Linear Algebra
# Let see some common linear algebra operations.
#
# We generate some random matrices and vectors
A = rand(3, 3) # Random 3 by 3 matrix
b = rand(3) # random vector of three elements
# The ```\``` operator solves the linear system $Ax = b$.
x = A\b
# Matrix multiplication, addition, etc. are already possible via multiple dispatch using the same operators used for numbers
A*b
A^3
b * transpose(b)
# Notice that all operation like ```reduce```, ```prod```, ```sum``` works on arrays of matrices as well.
sum([rand(2,2) for i in 1:5])
prod([rand(2,2) for i in 1:5])
# We can use more linear algebra functions by importing the module ```LinearAlgebra```
using LinearAlgebra
# We have now access to code for findig eigenvalues and eigenvectors
# +
A = [1 0 2
3 4 8
0 0 7]
eigen(A)
# -
# Or to compute, for example, the SVD
svd(A)
# But how many methods do we have to compute the SVD?
methods(svd)
# Let us create a _diagonal_ matrix:
D = Diagonal([1.,5.,9.])
# We can still call the ```svd``` function and the correct implementation will be called
svd(D)
@which svd(D)
# We have many different kinds of matrices
T = LowerTriangular(A)
svd(T)
@which svd(T)
# This allows to write the same code and have it compile down to the correct (and optimized) implementation
# ### The end
# The following is valid Julia code:
for 🌙 in '🌑':'🌘'
print(🌙)
end
# ## Extras
# #### Custom ```+``` for string concatenation
# Let us add ```+``` as string concatenation. We start by importing ```Base:+```
import Base:+
# We define how ```+``` should work with strings
function +(x::String, y::String)
x * y
end
"Hello " + "World"
# And now functions written **before** we wrote our ```+``` implementation will make use of it.
sum(["Hello", " ", "world"])
# #### A ```@unless``` macro
# We want to write a macro that execute a piece of code _unless_ a certain condition is satisfied
macro unless(test, body)
quote
if !$test
$body
end
end
end
x = 4
@unless x ∈ 1:5 println("Foo")
@unless x > 10 println("Bar")
@macroexpand @unless x ∈ 1:5 println("Foo")
# #### Push!, Sort!, and functions modifying their arguments
# Some functions modify their arguments. By convention such functions are denoted by a ```!``` at the end of the name.
#
# For example ```push!``` and ```pop!``` can be used to add and remove elements at the end of an array
v = [1,2,3,4]
push!(v, 5)
v
pop!(v)
v
# Some functions have two versions. Like ```sort```, returning a sorted copy of the original array, and ```sort!```, sorting the array passed as argument directly.
w = rand(10)
sort(w)
w
sort!(w)
w
|
Lecture 1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] hide_input=false
# ## 마무리
# ### 머신 러닝 문제 접근 방법
# ### 의사 결정 참여
# -
# ### 프로토타입에서 제품까지
# ### 제품 시스템 테스트
# ### 나만의 예측기 만들기
# +
from sklearn.base import BaseEstimator, TransformerMixin
class MyTransformer(BaseEstimator, TransformerMixin):
def __init__(self, first_paramter=1, second_parameter=2):
# __init__ 메소드에 필요한 모든 매개변수를 나열합니다
self.first_paramter = 1
self.second_parameter = 2
def fit(self, X, y=None):
# fit 메소드는 X와 y 매개변수만을 갖습니다
# 비지도 학습 모델이더라도 y 매개변수를 받도록 해야합니다!
# 모델 학습 시작
print("모델 학습을 시작합니다")
# 객체 자신인 self를 반환합니다
return self
def transform(self, X):
# transform 메소드는 X 매개변수만을 받습니다
# X를 변환합니다
X_transformed = X + 1
return X_transformed
# -
# ### 더 배울 것들
# #### 이론
# #### 다른 머신 러닝 프레임워크, 패키지
# #### 랭킹, 추천 시스템과 그외 다른 알고리즘
# #### 확률 모델렝, 추론, 확률적 프로그래밍
# #### 신경망
# #### Scaling to larger datasets
# #### Honing your skills
# #### Conclusion
|
Python/scikit-learn/08-conclusion.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp layers
# default_cls_lvl 3
# -
#export
from local.core.all import *
from local.torch_imports import *
from local.torch_core import *
from local.test import *
from torch.nn.utils import weight_norm, spectral_norm
from local.notebook.showdoc import *
# # Layers
# > Custom fastai layers and basic functions to grab them.
# ## Basic manipulations and resize
#export
class Identity(Module):
"Do nothing at all"
def forward(self,x): return x
# export
class Lambda(Module):
"An easy way to create a pytorch layer for a simple `func`"
def __init__(self, func): self.func=func
def forward(self, x): return self.func(x)
def __repr__(self): return f'{self.__class__.__name__}({self.func})'
# > Warning: In the tests below, we use lambda functions for convenience, but you shouldn't do this when building a real modules as it would make models that won't pickle (so you won't be able to save/export them).
tst = Lambda(lambda x:x+2)
x = torch.randn(10,20)
test_eq(tst(x), x+2)
# export
class PartialLambda(Lambda):
"Layer that applies `partial(func, **kwargs)`"
def __init__(self, func, **kwargs):
super().__init__(partial(func, **kwargs))
self.repr = f'{func.__name__}, {kwargs}'
def forward(self, x): return self.func(x)
def __repr__(self): return f'{self.__class__.__name__}({self.repr})'
def test_func(a,b=2): return a+b
tst = PartialLambda(test_func, b=5)
test_eq(tst(x), x+5)
# export
class View(Module):
"Reshape `x` to `size`"
def __init__(self, *size): self.size = size
def forward(self, x): return x.view(self.size)
tst = View(10,5,4)
test_eq(tst(x).shape, [10,5,4])
# export
class ResizeBatch(Module):
"Reshape `x` to `size`, keeping batch dim the same size"
def __init__(self, *size): self.size = size
def forward(self, x): return x.view((x.size(0),) + self.size)
tst = ResizeBatch(5,4)
test_eq(tst(x).shape, [10,5,4])
# export
class Flatten(Module):
"Flatten `x` to a single dimension, often used at the end of a model. `full` for rank-1 tensor"
def __init__(self, full=False): self.full = full
def forward(self, x): return x.view(-1) if self.full else x.view(x.size(0), -1)
tst = Flatten()
x = torch.randn(10,5,4)
test_eq(tst(x).shape, [10,20])
tst = Flatten(full=True)
test_eq(tst(x).shape, [200])
# export
class Debugger(nn.Module):
"A module to debug inside a model."
def forward(self,x):
set_trace()
return x
# export
def sigmoid_range(x, low, high):
"Sigmoid function with range `(low, high)`"
return torch.sigmoid(x) * (high - low) + low
test = tensor([-10.,0.,10.])
assert torch.allclose(sigmoid_range(test, -1, 2), tensor([-1.,0.5, 2.]), atol=1e-4, rtol=1e-4)
assert torch.allclose(sigmoid_range(test, -5, -1), tensor([-5.,-3.,-1.]), atol=1e-4, rtol=1e-4)
assert torch.allclose(sigmoid_range(test, 2, 4), tensor([2., 3., 4.]), atol=1e-4, rtol=1e-4)
# export
class SigmoidRange(Module):
"Sigmoid module with range `(low, high)`"
def __init__(self, low, high): self.low,self.high = low,high
def forward(self, x): return sigmoid_range(x, self.low, self.high)
tst = SigmoidRange(-1, 2)
assert torch.allclose(tst(test), tensor([-1.,0.5, 2.]), atol=1e-4, rtol=1e-4)
# ## Pooling layers
# export
class AdaptiveConcatPool2d(nn.Module):
"Layer that concats `AdaptiveAvgPool2d` and `AdaptiveMaxPool2d`"
def __init__(self, size=None):
super().__init__()
self.size = size or 1
self.ap = nn.AdaptiveAvgPool2d(self.size)
self.mp = nn.AdaptiveMaxPool2d(self.size)
def forward(self, x): return torch.cat([self.mp(x), self.ap(x)], 1)
# If the input is `bs x nf x h x h`, the output will be `bs x 2*nf x 1 x 1` if no size is passed or `bs x 2*nf x size x size`
tst = AdaptiveConcatPool2d()
x = torch.randn(10,5,4,4)
test_eq(tst(x).shape, [10,10,1,1])
max1 = torch.max(x, dim=2, keepdim=True)[0]
maxp = torch.max(max1, dim=3, keepdim=True)[0]
test_eq(tst(x)[:,:5], maxp)
test_eq(tst(x)[:,5:], x.mean(dim=[2,3], keepdim=True))
tst = AdaptiveConcatPool2d(2)
test_eq(tst(x).shape, [10,10,2,2])
# export
mk_class('PoolType', **{o:o for o in 'Avg Max Cat'.split()})
# export
_all_ = ['PoolType']
#export
def pool_layer(pool_type):
return nn.AdaptiveAvgPool2d if pool_type=='Avg' else nn.AdaptiveMaxPool2d if pool_type=='Max' else AdaptiveConcatPool2d
# export
class PoolFlatten(nn.Sequential):
"Combine `nn.AdaptiveAvgPool2d` and `Flatten`."
def __init__(self, pool_type=PoolType.Avg): super().__init__(pool_layer(pool_type)(1), Flatten())
tst = PoolFlatten()
test_eq(tst(x).shape, [10,5])
test_eq(tst(x), x.mean(dim=[2,3]))
# ## BatchNorm layers
# export
NormType = Enum('NormType', 'Batch BatchZero Weight Spectral')
#export
def BatchNorm(nf, norm_type=NormType.Batch, ndim=2, **kwargs):
"BatchNorm layer with `nf` features and `ndim` initialized depending on `norm_type`."
assert 1 <= ndim <= 3
bn = getattr(nn, f"BatchNorm{ndim}d")(nf, **kwargs)
bn.bias.data.fill_(1e-3)
bn.weight.data.fill_(0. if norm_type==NormType.BatchZero else 1.)
return bn
# `kwargs` are passed to `nn.BatchNorm` and can be `eps`, `momentum`, `affine` and `track_running_stats`.
tst = BatchNorm(15)
assert isinstance(tst, nn.BatchNorm2d)
test_eq(tst.weight, torch.ones(15))
tst = BatchNorm(15, norm_type=NormType.BatchZero)
test_eq(tst.weight, torch.zeros(15))
tst = BatchNorm(15, ndim=1)
assert isinstance(tst, nn.BatchNorm1d)
tst = BatchNorm(15, ndim=3)
assert isinstance(tst, nn.BatchNorm3d)
# export
class BatchNorm1dFlat(nn.BatchNorm1d):
"`nn.BatchNorm1d`, but first flattens leading dimensions"
def forward(self, x):
if x.dim()==2: return super().forward(x)
*f,l = x.shape
x = x.contiguous().view(-1,l)
return super().forward(x).view(*f,l)
tst = BatchNorm1dFlat(15)
x = torch.randn(32, 64, 15)
y = tst(x)
mean = x.mean(dim=[0,1])
test_close(tst.running_mean, 0*0.9 + mean*0.1)
var = (x-mean).pow(2).mean(dim=[0,1])
test_close(tst.running_var, 1*0.9 + var*0.1, eps=1e-4)
test_close(y, (x-mean)/torch.sqrt(var+1e-5) * tst.weight + tst.bias, eps=1e-4)
# export
class LinBnDrop(nn.Sequential):
"Module grouping `BatchNorm1d`, `Dropout` and `Linear` layers"
def __init__(self, n_in, n_out, bn=True, p=0., act=None, lin_first=False):
layers = [BatchNorm(n_out if lin_first else n_in, ndim=1)] if bn else []
if p != 0: layers.append(nn.Dropout(p))
lin = [nn.Linear(n_in, n_out, bias=not bn)]
if act is not None: lin.append(act)
layers = lin+layers if lin_first else layers+lin
super().__init__(*layers)
# The `BatchNorm` layer is skipped if `bn=False`, as is the dropout if `p=0.`. Optionally, you can add an activation for after the linear laeyr with `act`.
# +
tst = LinBnDrop(10, 20)
mods = list(tst.children())
test_eq(len(mods), 2)
assert isinstance(mods[0], nn.BatchNorm1d)
assert isinstance(mods[1], nn.Linear)
tst = LinBnDrop(10, 20, p=0.1)
mods = list(tst.children())
test_eq(len(mods), 3)
assert isinstance(mods[0], nn.BatchNorm1d)
assert isinstance(mods[1], nn.Dropout)
assert isinstance(mods[2], nn.Linear)
tst = LinBnDrop(10, 20, act=nn.ReLU(), lin_first=True)
mods = list(tst.children())
test_eq(len(mods), 3)
assert isinstance(mods[0], nn.Linear)
assert isinstance(mods[1], nn.ReLU)
assert isinstance(mods[2], nn.BatchNorm1d)
tst = LinBnDrop(10, 20, bn=False)
mods = list(tst.children())
test_eq(len(mods), 1)
assert isinstance(mods[0], nn.Linear)
# -
# ## Convolutions
#export
def init_default(m, func=nn.init.kaiming_normal_):
"Initialize `m` weights with `func` and set `bias` to 0."
if func and hasattr(m, 'weight'): func(m.weight)
with torch.no_grad():
if getattr(m, 'bias', None) is not None: m.bias.fill_(0.)
return m
#export
def _conv_func(ndim=2, transpose=False):
"Return the proper conv `ndim` function, potentially `transposed`."
assert 1 <= ndim <=3
return getattr(nn, f'Conv{"Transpose" if transpose else ""}{ndim}d')
#hide
test_eq(_conv_func(ndim=1),torch.nn.modules.conv.Conv1d)
test_eq(_conv_func(ndim=2),torch.nn.modules.conv.Conv2d)
test_eq(_conv_func(ndim=3),torch.nn.modules.conv.Conv3d)
test_eq(_conv_func(ndim=1, transpose=True),torch.nn.modules.conv.ConvTranspose1d)
test_eq(_conv_func(ndim=2, transpose=True),torch.nn.modules.conv.ConvTranspose2d)
test_eq(_conv_func(ndim=3, transpose=True),torch.nn.modules.conv.ConvTranspose3d)
# export
defaults.activation=nn.ReLU
# export
class ConvLayer(nn.Sequential):
"Create a sequence of convolutional (`ni` to `nf`), ReLU (if `use_activ`) and `norm_type` layers."
def __init__(self, ni, nf, ks=3, stride=1, padding=None, bias=None, ndim=2, norm_type=NormType.Batch, bn_1st=True,
act_cls=defaults.activation, transpose=False, init=nn.init.kaiming_normal_, xtra=None, **kwargs):
if padding is None: padding = ((ks-1)//2 if not transpose else 0)
bn = norm_type in (NormType.Batch, NormType.BatchZero)
if bias is None: bias = not bn
conv_func = _conv_func(ndim, transpose=transpose)
conv = init_default(conv_func(ni, nf, kernel_size=ks, bias=bias, stride=stride, padding=padding, **kwargs), init)
if norm_type==NormType.Weight: conv = weight_norm(conv)
elif norm_type==NormType.Spectral: conv = spectral_norm(conv)
layers = [conv]
act_bn = []
if act_cls is not None: act_bn.append(act_cls())
if bn: act_bn.append(BatchNorm(nf, norm_type=norm_type, ndim=ndim))
if bn_1st: act_bn.reverse()
layers += act_bn
if xtra: layers.append(xtra)
super().__init__(*layers)
# The convolution uses `ks` (kernel size) `stride`, `padding` and `bias`. `padding` will default to the appropriate value (`(ks-1)//2` if it's not a transposed conv) and `bias` will default to `True` the `norm_type` is `Spectral` or `Weight`, `False` if it's `Batch` or `BatchZero`. Note that if you don't want any normalization, you should pass `norm_type=None`.
#
# This defines a conv layer with `ndim` (1,2 or 3) that will be a ConvTranspose if `transpose=True`. `act_cls` is the class of the activation function to use (instantiated inside). Pass `act=None` if you don't want an activation function. If you quickly want to change your default activation, you can change the value of `defaults.activation`.
#
# `init` is used to initialize the weights (the bias are initiliazed to 0) and `xtra` is an optional layer to add at the end.
tst = ConvLayer(16, 32)
mods = list(tst.children())
test_eq(len(mods), 3)
test_eq(mods[1].weight, torch.ones(32))
test_eq(mods[0].padding, (1,1))
x = torch.randn(64, 16, 8, 8)#.cuda()
# +
# tst = tst.cuda()
# -
#Padding is selected to make the shape the same if stride=1
test_eq(tst(x).shape, [64,32,8,8])
#Padding is selected to make the shape half if stride=2
tst = ConvLayer(16, 32, stride=2)
test_eq(tst(x).shape, [64,32,4,4])
#But you can always pass your own padding if you want
tst = ConvLayer(16, 32, padding=0)
test_eq(tst(x).shape, [64,32,6,6])
#No bias by default for Batch NormType
assert mods[0].bias is None
#But can be overriden with `bias=True`
tst = ConvLayer(16, 32, bias=True)
test_eq(list(tst.children())[0].bias, torch.zeros(32))
#For no norm, or spectral/weight, bias is True by default
for t in [None, NormType.Spectral, NormType.Weight]:
tst = ConvLayer(16, 32, norm_type=t)
test_eq(list(tst.children())[0].bias, torch.zeros(32))
#Various n_dim/tranpose
tst = ConvLayer(16, 32, ndim=3)
assert isinstance(list(tst.children())[0], nn.Conv3d)
tst = ConvLayer(16, 32, ndim=1, transpose=True)
assert isinstance(list(tst.children())[0], nn.ConvTranspose1d)
#No activation/leaky
tst = ConvLayer(16, 32, ndim=3, act_cls=None)
mods = list(tst.children())
test_eq(len(mods), 2)
tst = ConvLayer(16, 32, ndim=3, act_cls=partial(nn.LeakyReLU, negative_slope=0.1))
mods = list(tst.children())
test_eq(len(mods), 3)
assert isinstance(mods[2], nn.LeakyReLU)
nn.MaxPool2d
#export
def AdaptiveAvgPool(sz=1, ndim=2):
"nn.AdaptiveAvgPool layer for `ndim`"
assert 1 <= ndim <= 3
return getattr(nn, f"AdaptiveAvgPool{ndim}d")(sz)
#export
def MaxPool(ks=2, stride=None, padding=0, ndim=2):
"nn.MaxPool layer for `ndim`"
assert 1 <= ndim <= 3
return getattr(nn, f"MaxPool{ndim}d")(ks, stride=stride, padding=padding)
#export
def AvgPool(ks=2, stride=None, padding=0, ndim=2, ceil_mode=False):
"nn.AvgPool layer for `ndim`"
assert 1 <= ndim <= 3
return getattr(nn, f"AvgPool{ndim}d")(ks, stride=stride, padding=padding, ceil_mode=ceil_mode)
# ## fastai loss functions
# The following class if the base class to warp a loss function it provides several added functionality:
# - it flattens the tensors before trying to take the losses since it's more convenient (with a potential tranpose to put `axis` at the end)
# - it has a potential `activation` method that tells the library if there is an activation fused in the loss (useful for inference and methods such as `Learner.get_preds` or `Learner.predict`)
# - it has a potential `decodes` method that is used on predictions in inference (for instance, an argmax in classification)
F.binary_cross_entropy_with_logits(torch.randn(4,5), torch.randint(0, 2, (4,5)).float(), reduction='none')
# export
@funcs_kwargs
class BaseLoss():
"Same as `loss_cls`, but flattens input and target."
activation=decodes=noops
_methods = "activation decodes".split()
def __init__(self, loss_cls, *args, axis=-1, flatten=True, floatify=False, is_2d=True, **kwargs):
store_attr(self, "axis,flatten,floatify,is_2d")
self.func = loss_cls(*args,**kwargs)
functools.update_wrapper(self, self.func)
def __repr__(self): return f"FlattenedLoss of {self.func}"
@property
def reduction(self): return self.func.reduction
@reduction.setter
def reduction(self, v): self.func.reduction = v
def __call__(self, inp, targ, **kwargs):
inp = inp .transpose(self.axis,-1).contiguous()
targ = targ.transpose(self.axis,-1).contiguous()
if self.floatify and targ.dtype!=torch.float16: targ = targ.float()
if targ.dtype in [torch.int8, torch.int16, torch.int32]: targ = targ.long()
if self.flatten: inp = inp.view(-1,inp.shape[-1]) if self.is_2d else inp.view(-1)
return self.func.__call__(inp, targ.view(-1) if self.flatten else targ, **kwargs)
# The `args` and `kwargs` will be passed to `loss_cls` during the initialization to instantiate a loss function. `axis` is put at the end for losses like softmax that are often performed on the last axis. If `floatify=True` the targs will be converted to float (usefull for losses that only accept float targets like `BCEWithLogitsLoss`) and `is_2d` determines if we flatten while keeping the first dimension (batch size) or completely flatten the input. We want the first for losses like Cross Entropy, and the second for pretty much anything else.
# export
@delegates(keep=True)
class CrossEntropyLossFlat(BaseLoss):
"Same as `nn.CrossEntropyLoss`, but flattens input and target."
y_int = True
def __init__(self, *args, axis=-1, **kwargs): super().__init__(nn.CrossEntropyLoss, *args, axis=axis, **kwargs)
def decodes(self, x): return x.argmax(dim=self.axis)
def activation(self, x): return F.softmax(x, dim=self.axis)
# +
tst = CrossEntropyLossFlat()
output = torch.randn(32, 5, 10)
target = torch.randint(0, 10, (32,5))
#nn.CrossEntropy would fail with those two tensors, but not our flattened version.
_ = tst(output, target)
test_fail(lambda x: nn.CrossEntropyLoss()(output,target))
#Associated activation is softmax
test_eq(tst.activation(output), F.softmax(output, dim=-1))
#This loss function has a decodes which is argmax
test_eq(tst.decodes(output), output.argmax(dim=-1))
# +
#In a segmentation task, we want to take the softmax over the channel dimension
tst = CrossEntropyLossFlat(axis=1)
output = torch.randn(32, 5, 128, 128)
target = torch.randint(0, 5, (32, 128, 128))
_ = tst(output, target)
test_eq(tst.activation(output), F.softmax(output, dim=1))
test_eq(tst.decodes(output), output.argmax(dim=1))
# -
# export
@delegates(keep=True)
class BCEWithLogitsLossFlat(BaseLoss):
"Same as `nn.CrossEntropyLoss`, but flattens input and target."
def __init__(self, *args, axis=-1, floatify=True, thresh=0.5, **kwargs):
super().__init__(nn.BCEWithLogitsLoss, *args, axis=axis, floatify=floatify, is_2d=False, **kwargs)
self.thresh = thresh
def decodes(self, x): return x>self.thresh
def activation(self, x): return torch.sigmoid(x)
# +
tst = BCEWithLogitsLossFlat()
output = torch.randn(32, 5, 10)
target = torch.randn(32, 5, 10)
#nn.BCEWithLogitsLoss would fail with those two tensors, but not our flattened version.
_ = tst(output, target)
test_fail(lambda x: nn.BCEWithLogitsLoss()(output,target))
output = torch.randn(32, 5)
target = torch.randint(0,2,(32, 5))
#nn.BCEWithLogitsLoss would fail with int targets but not our flattened version.
_ = tst(output, target)
test_fail(lambda x: nn.BCEWithLogitsLoss()(output,target))
#Associated activation is sigmoid
test_eq(tst.activation(output), torch.sigmoid(output))
# -
# export
def BCELossFlat(*args, axis=-1, floatify=True, **kwargs):
"Same as `nn.BCELoss`, but flattens input and target."
return BaseLoss(nn.BCELoss, *args, axis=axis, floatify=floatify, is_2d=False, **kwargs)
tst = BCELossFlat()
output = torch.sigmoid(torch.randn(32, 5, 10))
target = torch.randint(0,2,(32, 5, 10))
_ = tst(output, target)
test_fail(lambda x: nn.BCELoss()(output,target))
# export
def MSELossFlat(*args, axis=-1, floatify=True, **kwargs):
"Same as `nn.MSELoss`, but flattens input and target."
return BaseLoss(nn.MSELoss, *args, axis=axis, floatify=floatify, is_2d=False, **kwargs)
tst = MSELossFlat()
output = torch.sigmoid(torch.randn(32, 5, 10))
target = torch.randint(0,2,(32, 5, 10))
_ = tst(output, target)
test_fail(lambda x: nn.MSELoss()(output,target))
#hide
#cuda
#Test losses work in half precision
output = torch.sigmoid(torch.randn(32, 5, 10)).half().cuda()
target = torch.randint(0,2,(32, 5, 10)).half().cuda()
for tst in [BCELossFlat(), MSELossFlat()]: _ = tst(output, target)
#export
class LabelSmoothingCrossEntropy(Module):
y_int = True
def __init__(self, eps:float=0.1, reduction='mean'): self.eps,self.reduction = eps,reduction
def forward(self, output, target):
c = output.size()[-1]
log_preds = F.log_softmax(output, dim=-1)
if self.reduction=='sum': loss = -log_preds.sum()
else:
loss = -log_preds.sum(dim=-1)
if self.reduction=='mean': loss = loss.mean()
return loss*self.eps/c + (1-self.eps) * F.nll_loss(log_preds, target.long(), reduction=self.reduction)
def activation(self, out): return F.softmax(out, dim=-1)
def decodes(self, out): return out.argmax(dim=-1)
# On top of the formula we define:
# - a `reduction` attribute, that will be used when we call `Learner.get_preds`
# - an `activation` function that represents the activation fused in the loss (since we use cross entropy behind the scenes). It will be applied to the output of the model when calling `Learner.get_preds` or `Learner.predict`
# - a `decodes` function that converts the output of the model to a format similar to the target (here indices). This is used in `Learner.predict` and `Learner.show_results` to decode the predictions
# ## Embeddings
# export
def trunc_normal_(x, mean=0., std=1.):
"Truncated normal initialization (approximation)"
# From https://discuss.pytorch.org/t/implementing-truncated-normal-initializer/4778/12
return x.normal_().fmod_(2).mul_(std).add_(mean)
# export
class Embedding(nn.Embedding):
"Embedding layer with truncated normal initialization"
def __init__(self, ni, nf):
super().__init__(ni, nf)
trunc_normal_(self.weight.data, std=0.01)
# Truncated normal initialization bounds the distribution to avoid large value. For a given standard deviation `std`, the bounds are roughly `-std`, `std`.
tst = Embedding(10, 30)
assert tst.weight.min() > -0.02
assert tst.weight.max() < 0.02
test_close(tst.weight.mean(), 0, 1e-2)
test_close(tst.weight.std(), 0.01, 0.1)
# ## Self attention
# export
class SelfAttention(nn.Module):
"Self attention layer for `n_channels`."
def __init__(self, n_channels):
super().__init__()
self.query,self.key,self.value = [self._conv(n_channels, c) for c in (n_channels//8,n_channels//8,n_channels)]
self.gamma = nn.Parameter(tensor([0.]))
def _conv(self,n_in,n_out):
return ConvLayer(n_in, n_out, ks=1, ndim=1, norm_type=NormType.Spectral, act_cls=None, bias=False)
def forward(self, x):
#Notation from the paper.
size = x.size()
x = x.view(*size[:2],-1)
f,g,h = self.query(x),self.key(x),self.value(x)
beta = F.softmax(torch.bmm(f.transpose(1,2), g), dim=1)
o = self.gamma * torch.bmm(h, beta) + x
return o.view(*size).contiguous()
# Self-attention layer as introduced in [Self-Attention Generative Adversarial Networks](https://arxiv.org/abs/1805.08318).
#
# Initially, no change is done to the input. This is controlled by a trainable parameter named `gamma` as we return `x + gamma * out`.
tst = SelfAttention(16)
x = torch.randn(32, 16, 8, 8)
test_eq(tst(x),x)
# Then during training `gamma` will probably change since it's a trainable parameter. Let's see what's hapenning when it gets a nonzero value.
tst.gamma.data.fill_(1.)
y = tst(x)
test_eq(y.shape, [32,16,8,8])
# The attention mechanism requires three matrix multiplications (here represented by 1x1 convs). The multiplications are done on the channel level (the second dimension in our tensor) and we flatten the feature map (which is 8x8 here). As in the paper, we note `f`, `g` and `h` the results of those multiplications.
q,k,v = tst.query[0].weight.data,tst.key[0].weight.data,tst.value[0].weight.data
test_eq([q.shape, k.shape, v.shape], [[2, 16, 1], [2, 16, 1], [16, 16, 1]])
f,g,h = map(lambda m: x.view(32, 16, 64).transpose(1,2) @ m.squeeze().t(), [q,k,v])
test_eq([f.shape, g.shape, h.shape], [[32,64,2], [32,64,2], [32,64,16]])
# The key part of the attention layer is to compute attention weights for each of our location in the feature map (here 8x8 = 64). Those are positive numbers that sum to 1 and tell the model to pay attention to this or that part of the picture. We make the product of `f` and the transpose of `g` (to get something of size bs by 64 by 64) then apply a softmax on the first dimension (to get the positive numbers that sum up to 1). The result can then be multiplied with `h` transposed to get an output of size bs by channels by 64, which we can then be viewed as an output the same size as the original input.
#
# The final result is then `x + gamma * out` as we saw before.
beta = F.softmax(torch.bmm(f, g.transpose(1,2)), dim=1)
test_eq(beta.shape, [32, 64, 64])
out = torch.bmm(h.transpose(1,2), beta)
test_eq(out.shape, [32, 16, 64])
test_close(y, x + out.view(32, 16, 8, 8), eps=1e-4)
# export
class PooledSelfAttention2d(nn.Module):
"Pooled self attention layer for 2d."
def __init__(self, n_channels):
super().__init__()
self.n_channels = n_channels
self.query,self.key,self.value = [self._conv(n_channels, c) for c in (n_channels//8,n_channels//8,n_channels//2)]
self.out = self._conv(n_channels//2, n_channels)
self.gamma = nn.Parameter(tensor([0.]))
def _conv(self,n_in,n_out):
return ConvLayer(n_in, n_out, ks=1, norm_type=NormType.Spectral, act_cls=None, bias=False)
def forward(self, x):
n_ftrs = x.shape[2]*x.shape[3]
f = self.query(x).view(-1, self.n_channels//8, n_ftrs)
g = F.max_pool2d(self.key(x), [2,2]).view(-1, self.n_channels//8, n_ftrs//4)
h = F.max_pool2d(self.value(x), [2,2]).view(-1, self.n_channels//2, n_ftrs//4)
beta = F.softmax(torch.bmm(f.transpose(1, 2), g), -1)
o = self.out(torch.bmm(h, beta.transpose(1,2)).view(-1, self.n_channels//2, x.shape[2], x.shape[3]))
return self.gamma * o + x
# Self-attention layer used in the [Big GAN paper](https://arxiv.org/abs/1809.11096).
#
# It uses the same attention as in `SelfAttention` but adds a max pooling of stride 2 before computing the matrices `g` and `h`: the attention is ported on one of the 2x2 max-pooled window, not the whole feature map. There is also a final matrix product added at the end to the output, before retuning `gamma * out + x`.
#export
def _conv1d_spect(ni:int, no:int, ks:int=1, stride:int=1, padding:int=0, bias:bool=False):
"Create and initialize a `nn.Conv1d` layer with spectral normalization."
conv = nn.Conv1d(ni, no, ks, stride=stride, padding=padding, bias=bias)
nn.init.kaiming_normal_(conv.weight)
if bias: conv.bias.data.zero_()
return spectral_norm(conv)
#export
class SimpleSelfAttention(Module):
def __init__(self, n_in:int, ks=1, sym=False):
self.sym,self.n_in = sym,n_in
self.conv = _conv1d_spect(n_in, n_in, ks, padding=ks//2, bias=False)
self.gamma = nn.Parameter(tensor([0.]))
def forward(self,x):
if self.sym:
c = self.conv.weight.view(self.n_in,self.n_in)
c = (c + c.t())/2
self.conv.weight = c.view(self.n_in,self.n_in,1)
size = x.size()
x = x.view(*size[:2],-1)
convx = self.conv(x)
xxT = torch.bmm(x,x.permute(0,2,1).contiguous())
o = torch.bmm(xxT, convx)
o = self.gamma * o + x
return o.view(*size).contiguous()
# ## PixelShuffle
# PixelShuffle introduced in [this article](https://arxiv.org/pdf/1609.05158.pdf) to avoid checkerboard artifacts when upsampling images. If we want an output with `ch_out` filters, we use a convolution with `ch_out * (r**2)` filters, where `r` is the upsampling factor. Then we reorganize those filters like in the picture below:
#
# <img src="https://github.com/fastai/fastai_dev/blob/master/dev/images/pixelshuffle.png?raw=1" alt="Pixelshuffle" style="width: 100%; height: auto;"/>
# export
def icnr_init(x, scale=2, init=nn.init.kaiming_normal_):
"ICNR init of `x`, with `scale` and `init` function"
ni,nf,h,w = x.shape
ni2 = int(ni/(scale**2))
k = init(x.new_zeros([ni2,nf,h,w])).transpose(0, 1)
k = k.contiguous().view(ni2, nf, -1)
k = k.repeat(1, 1, scale**2)
return k.contiguous().view([nf,ni,h,w]).transpose(0, 1)
# ICNR init was introduced in [this article](https://arxiv.org/abs/1707.02937). It suggests to initialize the convolution that will be used in PixelShuffle so that each of the `r**2` channels get the same weight (so that in the picture above, the 9 colors in a 3 by 3 window are initially the same).
#
# > Note: This is done on the first dimension because PyTorch stores the weights of a convolutional layer in this format: `ch_out x ch_in x ks x ks`.
tst = torch.randn(16*4, 32, 1, 1)
tst = icnr_init(tst)
for i in range(0,16*4,4):
test_eq(tst[i],tst[i+1])
test_eq(tst[i],tst[i+2])
test_eq(tst[i],tst[i+3])
# export
class PixelShuffle_ICNR(nn.Sequential):
"Upsample by `scale` from `ni` filters to `nf` (default `ni`), using `nn.PixelShuffle`."
def __init__(self, ni, nf=None, scale=2, blur=False, norm_type=NormType.Weight, act_cls=defaults.activation):
super().__init__()
nf = ifnone(nf, ni)
layers = [ConvLayer(ni, nf*(scale**2), ks=1, norm_type=norm_type, act_cls=act_cls),
nn.PixelShuffle(scale)]
layers[0][0].weight.data.copy_(icnr_init(layers[0][0].weight.data))
if blur: layers += [nn.ReplicationPad2d((1,0,1,0)), nn.AvgPool2d(2, stride=1)]
super().__init__(*layers)
# The convolutional layer is initialized with `icnr_init` and passed `act_cls` and `norm_type` (the default of weight normalization seemed to be what's best for super-resolution problems, in our experiments).
#
# The `blur` option comes from [Super-Resolution using Convolutional Neural Networks without Any Checkerboard Artifacts](https://arxiv.org/abs/1806.02658) where the authors add a little bit of blur to completely get rid of checkerboard artifacts.
psfl = PixelShuffle_ICNR(16, norm_type=None) #Deactivate weight norm as it changes the weight
x = torch.randn(64, 16, 8, 8)
y = psfl(x)
test_eq(y.shape, [64, 16, 16, 16])
#ICNR init makes every 2x2 window (stride 2) have the same elements
for i in range(0,16,2):
for j in range(0,16,2):
test_eq(y[:,:,i,j],y[:,:,i+1,j])
test_eq(y[:,:,i,j],y[:,:,i ,j+1])
test_eq(y[:,:,i,j],y[:,:,i+1,j+1])
# ## Sequential extensions
# export
class SequentialEx(Module):
"Like `nn.Sequential`, but with ModuleList semantics, and can access module input"
def __init__(self, *layers): self.layers = nn.ModuleList(layers)
def forward(self, x):
res = x
for l in self.layers:
res.orig = x
nres = l(res)
# We have to remove res.orig to avoid hanging refs and therefore memory leaks
res.orig = None
res = nres
return res
def __getitem__(self,i): return self.layers[i]
def append(self,l): return self.layers.append(l)
def extend(self,l): return self.layers.extend(l)
def insert(self,i,l): return self.layers.insert(i,l)
# This is useful to write layers that require to remember the input (like a resnet block) in a sequential way.
# export
class MergeLayer(Module):
"Merge a shortcut with the result of the module by adding them or concatenating them if `dense=True`."
def __init__(self, dense:bool=False): self.dense=dense
def forward(self, x): return torch.cat([x,x.orig], dim=1) if self.dense else (x+x.orig)
res_block = SequentialEx(ConvLayer(16, 16), ConvLayer(16,16))
res_block.append(MergeLayer()) # just to test append - normally it would be in init params
x = torch.randn(32, 16, 8, 8)
y = res_block(x)
test_eq(y.shape, [32, 16, 8, 8])
test_eq(y, x + res_block[1](res_block[0](x)))
# ## Concat
# Equivalent to keras.layers.Concatenate, it will concat the outputs of a ModuleList over a given dimesion (default the filter dimesion)
#export
class Cat(nn.ModuleList):
"Concatenate layers outputs over a given dim"
def __init__(self, layers, dim=1):
self.dim=dim
super().__init__(layers)
def forward(self, x): return torch.cat([l(x) for l in self], dim=self.dim)
layers = [ConvLayer(2,4), ConvLayer(2,4), ConvLayer(2,4)]
x = torch.rand(1,2,8,8)
cat = Cat(layers)
test_eq(cat(x).shape, [1,12,8,8])
test_eq(cat(x), torch.cat([l(x) for l in layers], dim=1))
# ## Ready-to-go models
# export
class SimpleCNN(nn.Sequential):
"Create a simple CNN with `filters`."
def __init__(self, filters, kernel_szs=None, strides=None, bn=True):
nl = len(filters)-1
kernel_szs = ifnone(kernel_szs, [3]*nl)
strides = ifnone(strides , [2]*nl)
layers = [ConvLayer(filters[i], filters[i+1], kernel_szs[i], stride=strides[i],
norm_type=(NormType.Batch if bn and i<nl-1 else None)) for i in range(nl)]
layers.append(PoolFlatten())
super().__init__(*layers)
# The model is a succession of convolutional layers from `(filters[0],filters[1])` to `(filters[n-2],filters[n-1])` (if `n` is the length of the `filters` list) followed by a `PoolFlatten`. `kernel_szs` and `strides` defaults to a list of 3s and a list of 2s. If `bn=True` the convolutional layers are successions of conv-relu-batchnorm, otherwise conv-relu.
tst = SimpleCNN([8,16,32])
mods = list(tst.children())
test_eq(len(mods), 3)
test_eq([[m[0].in_channels, m[0].out_channels] for m in mods[:2]], [[8,16], [16,32]])
# Test kernel sizes
tst = SimpleCNN([8,16,32], kernel_szs=[1,3])
mods = list(tst.children())
test_eq([m[0].kernel_size for m in mods[:2]], [(1,1), (3,3)])
# Test strides
tst = SimpleCNN([8,16,32], strides=[1,2])
mods = list(tst.children())
test_eq([m[0].stride for m in mods[:2]], [(1,1),(2,2)])
# export
class ResBlock(nn.Module):
"Resnet block from `ni` to `nh` with `stride`"
@delegates(ConvLayer.__init__)
def __init__(self, expansion, ni, nh, stride=1, sa=False, sym=False,
norm_type=NormType.Batch, act_cls=defaults.activation, ndim=2, **kwargs):
super().__init__()
norm2 = NormType.BatchZero if norm_type==NormType.Batch else norm_type
nf,ni = nh*expansion,ni*expansion
layers = [ConvLayer(ni, nh, 3, stride=stride, norm_type=norm_type, act_cls=act_cls, ndim=ndim, **kwargs),
ConvLayer(nh, nf, 3, norm_type=norm2, act_cls=None, ndim=ndim, **kwargs)
] if expansion == 1 else [
ConvLayer(ni, nh, 1, norm_type=norm_type, act_cls=act_cls, ndim=ndim, **kwargs),
ConvLayer(nh, nh, 3, stride=stride, norm_type=norm_type, act_cls=act_cls, ndim=ndim, **kwargs),
ConvLayer(nh, nf, 1, norm_type=norm2, act_cls=None, ndim=ndim, **kwargs)
]
self.convs = nn.Sequential(*layers)
self.sa = SimpleSelfAttention(nf,ks=1,sym=sym) if sa else noop
self.idconv = noop if ni==nf else ConvLayer(ni, nf, 1, act_cls=None, ndim=ndim, **kwargs)
self.pool = noop if stride==1 else AvgPool(2, ndim=ndim, ceil_mode=True)
self.act = defaults.activation(inplace=True) if act_cls is defaults.activation else act_cls()
def forward(self, x): return self.act(self.sa(self.convs(x)) + self.idconv(self.pool(x)))
# This is a resnet block (normal or bottleneck depending on `expansion`, 1 for the normal block and 4 for the traditional bottleneck) that implements the tweaks from [Bag of Tricks for Image Classification with Convolutional Neural Networks](https://arxiv.org/abs/1812.01187). In particular, the last batchnorm layer (if that is the selected `norm_type`) is initialized with a weight (or gamma) of zero to facilitate the flow from the beginning to the end of the network.
#
# The `kwargs` are passed to `ConvLayer` along with `norm_type`.
# ## Swish and Mish
# +
#export
from torch.jit import script
@script
def _swish_jit_fwd(x): return x.mul(torch.sigmoid(x))
@script
def _swish_jit_bwd(x, grad_output):
x_sigmoid = torch.sigmoid(x)
return grad_output * (x_sigmoid * (1 + x * (1 - x_sigmoid)))
class _SwishJitAutoFn(torch.autograd.Function):
@staticmethod
def forward(ctx, x):
ctx.save_for_backward(x)
return _swish_jit_fwd(x)
@staticmethod
def backward(ctx, grad_output):
x = ctx.saved_variables[0]
return _swish_jit_bwd(x, grad_output)
# -
#export
def swish(x, inplace=False): return _SwishJitAutoFn.apply(x)
#export
class Swish(Module):
def forward(self, x): return _SwishJitAutoFn.apply(x)
# +
#export
@script
def _mish_jit_fwd(x): return x.mul(torch.tanh(F.softplus(x)))
@script
def _mish_jit_bwd(x, grad_output):
x_sigmoid = torch.sigmoid(x)
x_tanh_sp = F.softplus(x).tanh()
return grad_output.mul(x_tanh_sp + x * x_sigmoid * (1 - x_tanh_sp * x_tanh_sp))
class MishJitAutoFn(torch.autograd.Function):
@staticmethod
def forward(ctx, x):
ctx.save_for_backward(x)
return _mish_jit_fwd(x)
@staticmethod
def backward(ctx, grad_output):
x = ctx.saved_variables[0]
return _mish_jit_bwd(x, grad_output)
# -
#export
def mish(x): return MishJitAutoFn.apply(x)
#export
class MishJit(Module):
def forward(self, x): return MishJitAutoFn.apply(x)
# ## Helper functions for submodules
# It's easy to get the list of all parameters of a given model. For when you want all submodules (like linear/conv layers) without forgetting lone parameters, the following class wraps those in fake modules.
# export
class ParameterModule(Module):
"Register a lone parameter `p` in a module."
def __init__(self, p): self.val = p
def forward(self, x): return x
# export
def children_and_parameters(m):
"Return the children of `m` and its direct parameters not registered in modules."
children = list(m.children())
children_p = sum([[id(p) for p in c.parameters()] for c in m.children()],[])
for p in m.parameters():
if id(p) not in children_p: children.append(ParameterModule(p))
return children
# +
# export
class TstModule(Module):
def __init__(self): self.a,self.lin = nn.Parameter(torch.randn(1)),nn.Linear(5,10)
tst = TstModule()
children = children_and_parameters(tst)
test_eq(len(children), 2)
test_eq(children[0], tst.lin)
assert isinstance(children[1], ParameterModule)
test_eq(children[1].val, tst.a)
# +
#export
def _has_children(m:nn.Module):
try: next(m.children())
except StopIteration: return False
return True
nn.Module.has_children = property(_has_children)
# -
class A(Module): pass
assert not A().has_children
assert TstModule().has_children
# export
def flatten_model(m):
"Return the list of all submodules and parameters of `m`"
return sum(map(flatten_model,children_and_parameters(m)),[]) if m.has_children else [m]
tst = nn.Sequential(TstModule(), TstModule())
children = flatten_model(tst)
test_eq(len(children), 4)
assert isinstance(children[1], ParameterModule)
assert isinstance(children[3], ParameterModule)
#export
class NoneReduce():
"A context manager to evaluate `loss_func` with none reduce."
def __init__(self, loss_func): self.loss_func,self.old_red = loss_func,None
def __enter__(self):
if hasattr(self.loss_func, 'reduction'):
self.old_red = self.loss_func.reduction
self.loss_func.reduction = 'none'
return self.loss_func
else: return partial(self.loss_func, reduction='none')
def __exit__(self, type, value, traceback):
if self.old_red is not None: self.loss_func.reduction = self.old_red
# +
x,y = torch.randn(5),torch.randn(5)
loss_fn = nn.MSELoss()
with NoneReduce(loss_fn) as loss_func:
loss = loss_func(x,y)
test_eq(loss.shape, [5])
test_eq(loss_fn.reduction, 'mean')
loss_fn = F.mse_loss
with NoneReduce(loss_fn) as loss_func:
loss = loss_func(x,y)
test_eq(loss.shape, [5])
test_eq(loss_fn, F.mse_loss)
# -
#export
def in_channels(m):
"Return the shape of the first weight layer in `m`."
for l in flatten_model(m):
if hasattr(l, 'weight'): return l.weight.shape[1]
raise Exception('No weight layer')
test_eq(in_channels(nn.Sequential(nn.Conv2d(5,4,3), nn.Conv2d(4,3,3))), 5)
test_eq(in_channels(nn.Sequential(nn.AvgPool2d(4), nn.Conv2d(4,3,3))), 4)
test_fail(lambda : in_channels(nn.Sequential(nn.AvgPool2d(4))))
# ## Export -
#hide
from local.notebook.export import *
notebook2script(all_fs=True)
|
dev/03a_layers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1A.e - TD noté 2015 rattrapage (énoncé, écrit et oral)
#
# Questions posées à l'oral autour du jeu 2048 et d'un exercice Google Jam sur le position de carreaux dans un plus grand carré : [Problem D. Cut Tiles](https://code.google.com/codejam/contest/3214486/dashboard#s=p3).
from jyquickhelper import add_notebook_menu
add_notebook_menu()
# ## Question 1
#
# On s'intéresse à une phase du jeu [2048](http://gabrielecirulli.github.io/2048/). On part d'une grille :
mat = [[2,0,0,4],[0,2,8,2],[0,2,4,2],[2,2,8,0],]
for m in mat:
print(m)
# On veut écrire une fonction qui calcule l'état du jeu après la pression de la touche ``bas``.
# ## Question 2
#
# Modifier la fonction pour gérer les quatre directions.
# ## Question 3
#
# Remplir deux cases vides choisies aléatoirement avec deux chiffres 2.
# ## Question 4
#
# Proposer des solutions pour le problème [Problem D. Cut Tiles](https://code.google.com/codejam/contest/3214486/dashboard#s=p3). On n'implémentera pas ces solutions.
|
_doc/notebooks/exams/td_note_2015_rattrapage_enonce.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basics: overfitting a MLP on CIFAR10
#
# Training loop over CIFAR10 (40,000 train images, 10,000 test images). What happens if you
# - switch the training to a GPU? Is it faster?
# - Remove the `ReLU()`?
# - Increase the learning rate?
# - Stack more layers?
# - Perform more epochs?
#
# Can you completely overfit the training set (i.e. get 100% accuracy?)
#
# This code is highly non-modulable. Create functions for each specific task.
# (hint: see [this](https://github.com/pytorch/examples/blob/master/mnist/main.py))
#
# Your training went well. Good. Why not save the weights of the network (`net.state_dict()`) using `torch.save()`?
# +
import torch
import torchvision
import torch.nn as nn
import torchvision.transforms as t
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# model.to(device)
# define network structure
net = nn.Sequential(
nn.Conv2d(in_channels=3, out_channels=64, kernel_size=5, stride=1, padding=0, dilation=0, bias=True),
nn.MaxPool2d(2), nn.ReLU(),
nn.Conv2d(in_channels=64, out_channels=16, kernel_size=3, stride=1, padding=0, dilation=0, bias=True),
nn.MaxPool2d(2), nn.ReLU(),
view
nn.Linear(16*6*6, 256),
nn.ReLU(),
nn.Dropout(p=0.5, inplace=False),
nn.Linear(256, 64),
nn.ReLU(),
nn.Dropout(p=0.5, inplace=False),
nn.Linear(64, 10),
nn.Softmax(dim=1)
).to(device)
criterion = nn.CrossEntropyLoss(reduction = 'mean')
optimizer = torch.optim.SGD(net.parameters(), lr=0.02, momentum=0.9)
# optimizer = torch.optim.Adamax(net.parameters(), lr=0.01, betas=[0.4,0.2], eps=1e-6, weight_decay=0.9)
# optimizer = torch.optim.Adagrad(net.parameters(), lr=0.01, lr_decay=0, weight_decay=0, initial_accumulator_value=0, eps=1e-10)
# optimizer = torch.optim.Adam(net.parameters(), lr=0.001, betas=(0.9, 0.999), eps=1e-08, weight_decay=0, amsgrad=False)
# load data
to_tensor = t.ToTensor()
normalize = t.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))
flatten = t.Lambda(lambda x:x.view(-1))
transform_list = t.Compose([to_tensor, normalize]) #, flatten])
train_set = torchvision.datasets.CIFAR10(root='.', train=True, transform=transform_list, download=True)
test_set = torchvision.datasets.CIFAR10(root='.', train=False, transform=transform_list, download=True)
train_loader = torch.utils.data.DataLoader(train_set, batch_size=64)
test_loader = torch.utils.data.DataLoader(test_set, batch_size=64)
# +
# === Train === ###
net.train()
# train loop
for epoch in range(100):
train_correct = 0
train_loss = 0
print('Epoch {}'.format(epoch))
# loop per epoch
for i, (batch, targets) in enumerate(train_loader):
batch = batch.to(device)
targets = targets.to(device)
output = net(batch)
loss = criterion(output, targets)
optimizer.zero_grad()
loss.backward()
optimizer.step()
pred = output.max(1, keepdim=True)[1]
train_correct += pred.eq(targets.view_as(pred)).sum().item()
train_loss += loss
if i % 1000 == 10: print('Train loss {:.4f}, Train accuracy {:.2f}%'.format(
train_loss / ((i+1) * 64), 100 * train_correct / ((i+1) * 64)))
print('End of training.\n')
# === Test === ###
test_correct = 0
net.eval()
# loop, over whole test set
for i, (batch, targets) in enumerate(test_loader):
batch = batch.to(device)
targets = targets.to(device)
output = net(batch)
pred = output.max(1, keepdim=True)[1]
test_correct += pred.eq(targets.view_as(pred)).sum().item()
print('End of testing. Test accuracy {:.2f}%'.format(
100 * test_correct / (len(test_loader) * 64)))
# -
print(torch.cuda.current_device())
# ## Autograd tips and tricks
# Pointers are everywhere!
# +
net = nn.Linear(2, 2)
w = net.weight
print(w)
x = torch.rand(1, 2)
y = net(x).sum()
y.backward()
net.weight.data -= 0.01 * net.weight.grad # <--- What is this?
print(w)
# +
net = nn.Linear(2, 2)
w = net.weight.clone()
print(w)
x = torch.rand(1, 2)
y = net(x).sum()
y.backward()
net.weight.data -= 0.01 * net.weight.grad # <--- What is this?
print(w)
# -
# Sharing weights
# +
net = nn.Sequential(nn.Linear(2, 2), nn.Linear(2, 2))
net[0].weight = net[1].weight # weight sharing
x = torch.rand(1, 2)
y = net(x).sum()
y.backward()
print(net[0].weight.grad)
print(net[1].weight.grad)
|
Module5/Stacking_layers_MLP_CIFAR10.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pytorch
# language: python
# name: pytorch
# ---
# + tags=[]
import numpy as np
import torch
from torchvision import datasets, transforms
from torch.utils.data import DataLoader
from torchsummary import summary
from torch import nn
import torch.nn
from torch.functional import F
from tqdm import tqdm
import torch.optim as optim
import matplotlib.pyplot as plt
import os
os.chdir('d:\Python Projects\EVA')
# + tags=[]
train_transforms = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,),(0.3081,))
])
test_transforms = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,),(0.3081,))
])
# -
train = datasets.MNIST('./data', train=True, download=True, transform=train_transforms)
test = datasets.MNIST('./data',train=False, download=True, transform=test_transforms)
# + tags=[]
#
SEED = 1
device = 'cuda' if torch.cuda else 'cpu'
print(device)
# for Reproducable results
torch.manual_seed(SEED)
#Setting seed if CUDA
if torch.cuda.is_available():
torch.cuda.manual_seed(SEED)
# Dataloader Arguments
dataloader_args = dict(shuffle=True, num_workers=4, batch_size=128, pin_memory=True) if torch.cuda.is_available() else dict(shuffle=True, batch_size=64)
train_dataloader = DataLoader(train,**dataloader_args)
test_dataloader = DataLoader(test, **dataloader_args)
# + tags=[]
train_data = train.train_data
train_data = train.transform(train_data.numpy())
print(' - Shape ', train.train_data.cpu().numpy().shape)
print(' - Max ', torch.max(train_data))
print(' - Min ', torch.min(train_data))
print(' - Mean ',torch.mean(train_data))
print(' - Std dev ', torch.std(train_data))
print(' - Var ', torch.var(train_data))
# Displaying images
dataiter = iter(train_dataloader)
images, labels = dataiter.next()
plt.imshow(images[0].numpy().squeeze(), cmap='gray_r')
# +
#Plot bunch of images
figure = plt.figure()
NUM_IMAGES = 60
for index in range(1,NUM_IMAGES+1):
plt.subplot(6,10,index)
plt.axis('off')
plt.imshow(images[index].numpy().squeeze(), cmap='gray_r')
# + [markdown] tags=[]
# # The Model
# +
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Sequential(
nn.Conv2d(1, 10, 3,padding=1), # 28
nn.ReLU(),
# nn.Dropout(0.10),
nn.BatchNorm2d(10),
)
self.conv2 = nn.Sequential(
nn.Conv2d(10, 16, 3, padding=1), # 28
nn.ReLU(),
nn.BatchNorm2d(16),
# nn.Dropout(0.10),
nn.MaxPool2d(2,2),
)
self.conv3 = nn.Sequential(
nn.Conv2d(16, 10, 3, padding=1), # 14
nn.ReLU(),
# nn.Dropout(0.10),
nn.BatchNorm2d(10),
)
self.conv4 = nn.Sequential(
nn.Conv2d(10, 16, 3, padding=1), # 14
nn.ReLU(),
nn.BatchNorm2d(16),
# nn.Dropout(0.10),
nn.MaxPool2d(2,2), # 7
)
self.conv5 = nn.Sequential(
nn.Conv2d(16, 10, 3), # 5
nn.ReLU(),
# nn.Dropout(0.10),
nn.BatchNorm2d(10),
)
self.conv6 = nn.Sequential(
nn.Conv2d(10, 10, 3), #3
nn.ReLU(),
nn.BatchNorm2d(10),
)
self.conv7 = nn.Sequential(
nn.Conv2d(10, 10, 3), #1
)
def forward(self,x):
x = self.conv1(x)
x = self.conv2(x)
x = self.conv3(x)
x = self.conv4(x)
x = self.conv5(x)
x = self.conv6(x)
x = self.conv7(x)
x = x.view(-1,10)
return F.log_softmax(x, dim=-1)
# + tags=[]
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
model = Net().to(device)
summary(model, input_size=(1, 28, 28))
# + tags=[]
train_acc = []
test_acc = []
train_losses = []
test_losses = []
def train(model, device, train_dataloader, optimizer, epochs):
model.train()
pbar = tqdm(train_dataloader)
correct = 0
processed = 0
for batch_idx, (data, target) in enumerate(pbar):
data, target = data.to(device), target.to(device)
optimizer.zero_grad()
y_pred = model(data)
loss = F.nll_loss(y_pred, target)
train_losses.append(loss)
loss.backward()
optimizer.step()
pred = y_pred.argmax(dim=1, keepdim=True)
correct += pred.eq(target.view_as(pred)).sum().item()
processed += len(data)
pbar.set_description(desc=f'Loss={loss.item()} Batch_id={batch_idx} Accuracy={100*correct/processed:0.2f}')
train_acc.append(100*correct/processed)
def test(model, device, test_dataloader):
model.eval()
test_loss = 0
correct = 0
with torch.no_grad():
for data, target in test_dataloader:
data, target = data.to(device), target.to(device)
output = model(data)
pred = output.argmax(dim=1, keepdim=True)
test_loss += F.nll_loss(output, target, reduction='sum').item()
correct += pred.eq(target.view_as(pred)).sum().item()
test_loss /= len(test_dataloader.dataset)
test_losses.append(test_loss)
print('\nTest set: Average loss: {:.4f}, Accuracy: {}/{} ({:.2f}%)\n'.format(
test_loss, correct, len(test_dataloader.dataset),
100. * correct / len(test_dataloader.dataset)))
test_acc.append(100. * correct / len(test_dataloader.dataset))
# + tags=[]
optimizer = optim.SGD(model.parameters(), lr=0.01, momentum=0.9)
EPOCHS = 15
for epoch in range(EPOCHS):
print('EPOCH - ', epoch)
train(model, device, train_dataloader, optimizer, epoch)
test(model, device, test_dataloader)
# -
fig, axs = plt.subplots(2,2, figsize=(15,10))
axs[0,0].plot(train_losses)
axs[0,0].set_title('train_losses')
axs[0,1].plot(train_acc)
axs[0,1].set_title('Training Accuracy')
axs[1,0].plot(test_losses)
axs[1,0].set_title('Test Losses')
axs[1,1].plot(test_acc)
axs[1,1].set_title('Test Accuracy')
|
Assignment-5/Martyrs/Toppings - OnlyNorm_7k.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# language: python
# name: python38564bitf6f61b6d739443d992db3bfd3acfcce2
# ---
# # Direct Methods
#
# Direct Methods allow to obtain the exact solution in finitely many operations.
#
# The problem is with scale.
# For example, the well known Gauss elimination algorithm has $O(n^3)$ arithmetic complexity [<sup>1</sup>](#fn1).
# The real world problems may have a very large number of equations, like $(10^6 - 10^8)$.
# Think of a 3D, uniformly discretized mesh block like 1000x1000x1000 = $10^9$.
#
# Last but not least, the nonlinear systems of equations are always solved by iterative algorithms.
#
# ---
# <span id="fn1"> [Gauss algorithm](https://en.wikipedia.org/wiki/Gaussian_elimination#Computational_efficiency) </span>
# # Iterative methods
#
# ## Intuitive Example - direct iteration method
#
# Suppose, that we would like to solve a simple equation
#
# $$ 10x = 500 $$
#
# Let us rewrite this equation into two equivalent forms:
#
# $$ 6x = 500 - 4x \hspace{5em} and \hspace{5em} 4x = 500 - 6x $$
#
# Each form can be adopted to generate an iterative algorithm (let's call them version A and B).
#
# $$ x_A^{k+1} = (500 - 4x_A^k)/6 \hspace{5em} and \hspace{5em} x_B^{k+1} = (500 - 6x_B^k)/4 $$
#
#
# ### Task
#
# Write both versions of the iterative algorithm and try to solve the equations.
#
# Start with $ x^{(0)} = 37 $
# +
import numpy as np #here we load numpy
import matplotlib.pyplot as plt
import time, sys #and load some utilities
from numpy import array, zeros, diag, diagflat, dot
np.set_printoptions(precision=3, suppress=True)
# +
x_IC = 37
N = 50 # number of iterations
x_A = np.zeros(N)
x_A[0] = x_IC
x_B = np.zeros(N)
x_B[0] = x_IC
for i in range(N-1):
x_A[i+1] = (500 - 4*x_A[i])/6
x_B[i+1] = (500 - 6*x_B[i])/4
# iterations = np.arange(N)
# plt.plot(iterations, x_A)
# plt.plot(iterations, x_B)
# -
# ### Task
#
# Modify the algorithm to solve $ 10x = 500 + sin(x) $
#
# ### Question
#
# How an **intuitive** convergence criterion can be formulated?
# ### Answer
#
# The convergence is related to the fact the bigger part of the unknown $ x $ was used to form a new iteration.
# # System of linear Equations
#
# Now, we will focus on iterative methods for solving system of linear equations:
#
# $$
# \mathbb{A} \boldsymbol{x} = \boldsymbol{b}
# $$
#
# ## Jacobi iterative Method
#
# Let us decompose the $\mathbb{A}$ matrix into a Lower triangular matrix, Diagonal and Upper triangular matrix:
#
# $$
# \mathbb{A} = \mathbb{L} + \mathbb{D} + \mathbb{U}
# $$
#
#
# $$
# \underbrace{
# \begin{bmatrix}
# 1 & 2 & 3 \\
# 4 & 5 & 6 \\
# 7 & 8 & 9 \\
# \end{bmatrix}
# }_{\mathbb{A}}
# =
# \underbrace{
# \begin{bmatrix}
# 0 & 0 & 0 \\
# 4 & 0 & 0 \\
# 7 & 8 & 0 \\
# \end{bmatrix}
# }_{\mathbb{L}}
# +
# \underbrace{
# \begin{bmatrix}
# 1 & 0 & 0 \\
# 0 & 5 & 0 \\
# 0 & 0 & 9 \\
# \end{bmatrix}
# }_{\mathbb{D}}
# +
# \underbrace{
# \begin{bmatrix}
# 0 & 2 & 3 \\
# 0 & 0 & 6 \\
# 0 & 0 & 0 \\
# \end{bmatrix}
# }_{\mathbb{U}}
# $$
#
#
# +
A = [[1,2,3],[4,5,6],[7,8,9]]
L = np.tril(A,-1)
D = diagflat(diag(A)) # diag(A) - diagonal vector, diagflat(vector) - matrix form
U = np.triu(A,1)
print(f"L = \n{L}")
print(f"D = \n{D}")
print(f"U = \n{U}")
print(f"D^-1 = \n{np.linalg.inv(D)}")
# -
# The system $ \mathbb{A} \boldsymbol{x} = \boldsymbol{b}$, can be rewrtitten as:
#
# $$
# \mathbb{D} \boldsymbol{x} = -(\mathbb{L} + \mathbb{U})\boldsymbol{x} + \boldsymbol{b}
# $$
#
# Next, the iterative approach can be proposed as
#
# $$
# \mathbb{D} \boldsymbol{x}^{k+1} = -(\mathbb{L} + \mathbb{U})\boldsymbol{x}^k + \boldsymbol{b}
# $$
#
# or in an equivalent form:
#
# $$
# \boldsymbol{x}^{k+1} =
# \mathbb{D}^{-1} \left(\boldsymbol{b} - (\mathbb{L} + \mathbb{U})\boldsymbol{x}^k \right)
# $$
#
# Using index notation:
#
# $$
# x_{i}^{k+1}=\frac{1}{a_{i i}}\left(b_{i}-\sum_{j=1}^{i-1} a_{i j} x_{j}^{k}-\sum_{j=i+1}^{n} a_{i j} x_{j}^{k}\right), i=1, \ldots, n
# $$
#
# +
# Let us define the study case:
# An ugly Matrix
# A = np.array([[ 4, -1, -6, 0],
# [-5, -4, 10, 8],
# [ 0, 9, 4, -2],
# [ 1, 0, -7, 5]])
# b = np.array([2, 21, -12, -6])
# A nice Matrix
A = np.array([[10., -1., 2., 0.],
[-1., 11., -1., 3.],
[2., -1., 10., -1.],
[0., 3., -1., 8.]])
b = np.array([6.0, 25.0, -11.0, 15.0])
initial_guess = np.zeros(4)
alpha = 0.5 # Relaxation factor
N = 25 # number of iterations
iterations = np.arange(N)
# +
def jacobi(A,b,N,x0):
# https://www.quantstart.com/articles/Jacobi-Method-in-Python-and-NumPy/
d = diag(A) # this is a vector
LU = A - diagflat(d) # diagflat() puts the values of the vector on the diagonal
x = np.copy(x0)
residuals_norm = np.arange(N)
# Iterate for N times
for i in range(N):
residual = b - A@x
residuals_norm[i] = np.linalg.norm(residual)
x = (b - LU@x) / d
return x, residuals_norm
x_jacobi, residuals_norm_jacobi = jacobi(A,b,N,initial_guess)
print(f"jacobi: {x_jacobi}")
x_np = np.linalg.solve(A, b)
print(f"np.linalg.solve(A, b): {x_np}")
plt.rcParams.update({'font.size': 16})
figure, axis = plt.subplots(1, 1, figsize=(8, 6))
# plt.subplots_adjust(hspace=1)
axis.set_title('Residuals')
axis.plot(iterations, residuals_norm_jacobi, label=r'residuals_norm_jacobi')
axis.set_yscale('log')
axis.set_xlabel('iterations')
axis.set_ylabel('residuals_norm')
axis.legend(loc="upper right")
# -
#
# ## Gauss-Seidel Method
#
# Observe, that variables ($x$) with lower indices ($j < i$) are already known at $k+1$ iteration, thus they can be inserted 'on the fly' to enhance the convergence
#
# $$
# (\mathbb{L} + \mathbb{D}) \boldsymbol{x} = -\mathbb{U}\boldsymbol{x} + \boldsymbol{b}
# $$
#
# Next, the iterative approach can be proposed as
#
# $$
# (\mathbb{L} + \mathbb{D}) \boldsymbol{x}^{k+1} = -\mathbb{U}\boldsymbol{x}^k + \boldsymbol{b} \Leftrightarrow \\
# \Leftrightarrow \boldsymbol{x}^{k+1} = \mathbb{D}^{-1} \left(-\mathbb{L} \boldsymbol{x}^{k+1} -\mathbb{U}\boldsymbol{x}^k + \boldsymbol{b} \right)
# $$
#
# Using index notation:
# $$
# x_{i}^{k+1}=\frac{1}{a_{i i}}\left(b_{i}-\sum_{j=1}^{i-1} a_{i j} x_{j}^{k+1}-\sum_{j=i+1}^{n} a_{i j} x_{j}^{k}\right), i=1, \ldots, n
# $$
#
#
# + tags=[]
def gauss_seidel(A, b, N, x0):
# https://en.wikipedia.org/wiki/Gauss%E2%80%93Seidel_method
x = np.copy(x0)
d = diag(A)
L = np.tril(A,-1)
# D = diagflat(diag(A))
U = np.triu(A,1)
residuals_norm = np.arange(N)
for t in range(N):
residual = b - A@x
residuals_norm[t] = np.linalg.norm(residual)
for i in range(len(x)):
x[i] = (-L[i,:] @ x - U[i,:] @ x + b[i])/d[i]
return x, residuals_norm
x_GS, residuals_norm_GS = gauss_seidel(A,b,N,initial_guess)
print(f"gauss_seidel:\t\t{x_GS}")
x_np = np.linalg.solve(A, b)
print(f"np.linalg.solve(A, b): {x_np}")
plt.rcParams.update({'font.size': 16})
figure, axis = plt.subplots(1, 1, figsize=(8, 6))
axis.set_title('Residualse')
axis.plot(iterations, residuals_norm_GS, label=r'residuals_norm_GS')
axis.set_yscale('log')
axis.set_xlabel('iterations')
axis.set_ylabel('residuals_norm')
axis.legend(loc="upper right")
# -
# ## Errors and Residuals
#
# Let us denote the exact solution by $ \boldsymbol{x} $.
#
# ### Definitions
# The error is defined as:
#
# $$
# \boldsymbol{e}^k = \boldsymbol{x} - \boldsymbol{x}^k
# $$
#
# The residuum is defined as:
#
# $$
# \boldsymbol{r}^k = \boldsymbol{b} - \boldsymbol{b}^k = \boldsymbol{b} - \mathbb{A} \boldsymbol{x}^k = \mathbb{A}(\boldsymbol{x} - \boldsymbol{x}^k) = \mathbb{A} \boldsymbol{e}^k
#
# $$
#
# Consider an iterative method:
#
# $$
# \boldsymbol{x}^{k+1} = \boldsymbol{x}^k + \boldsymbol{p}^k
# $$
#
# where $ \boldsymbol{p}$ is a correction between iterations.
#
# Observe that if you calculate the error... and add it to the $\boldsymbol{x}^k$ you should get an exact solution :)
#
# Let us state for a while that $ \boldsymbol{p} = \boldsymbol{e} = \mathbb{A}^{-1}\boldsymbol{r}$
#
# $$
# \boldsymbol{x}^{k+1} = \boldsymbol{x}^k + \underbrace{\mathbb{A}^{-1}\boldsymbol{r}^k}_{\boldsymbol{p}^k} = \boldsymbol{x}^k + \mathbb{A}^{-1}\underbrace{(\boldsymbol{b} - \mathbb{A}\boldsymbol{x}^k)}_{\boldsymbol{r}^k} \\
# = \boldsymbol{x}^k - \mathbb{A}^{-1}\mathbb{A}\boldsymbol{x}^k + \mathbb{A}^{-1}\boldsymbol{b} \\
# = \mathbb{A}^{-1}\boldsymbol{b}
# $$
#
# The problem is, the we do not know $\mathbb{A}^{-1}$, and we do not want to calculate it as it is cumbersome.
# Instead, we would like to approximate it with another, **similar ** matrix, which has better numerical properties (is easy to invert).
# Such a matrix is called **preconditioner**.
#
#
# ## Jacobi method - revisited
#
# Notice, that in the case Jacobi method, the $\mathbb{A}$ matrix is approximated by its diagonal $\mathbb{D}$:
#
# $$
# \boldsymbol{x}^{k+1} = \boldsymbol{x}^k + \boldsymbol{p}^k = \boldsymbol{x}^k + \underbrace{\mathbb{D}^{-1}}_{\sim \mathbb{A}^{-1}}\boldsymbol{r}^k = \boldsymbol{x}^k + \underbrace{\mathbb{D}^{-1} (\boldsymbol{b} - \mathbb{A}\boldsymbol{x}^k)}_{\boldsymbol{p}^k}
# $$
#
# Which is equivalant to the formula presented before:
#
# $$
# \boldsymbol{x}^{k+1} = \underbrace{\boldsymbol{x}^k - \mathbb{D}^{-1} \mathbb{D} \boldsymbol{x}^k}_{=0} +
# \underbrace{\mathbb{D}^{-1} \left(\boldsymbol{b} - (\mathbb{L} + \mathbb{U})\boldsymbol{x}^k \right)}_{\boldsymbol{\hat{p}}^k}
# $$
#
# ## Relaxation
#
# Another 'trick', which helps to converge an iterative method, is to introduce a relaxation parameter, $\alpha$.
#
# $$
# \boldsymbol{x}^{k+1}
# = (1 - \alpha) \boldsymbol{x}^k + \alpha \boldsymbol{\hat{p}}^k
# = \boldsymbol{x}^k + \alpha \underbrace{(\boldsymbol{\hat{p}}^k - \boldsymbol{x}^k)}_{\boldsymbol{p}^k} \\
# $$
#
# From the implementation point of view, it may be more convienient to use $\boldsymbol{\hat{p}} = \boldsymbol{p}^k + \boldsymbol{x}^k$.
#
# ### Task
#
# Introduce the relaxation to the Jacobi and Gauss-Seidel method.
#
# * Jacobi:
# $$
# \boldsymbol{\hat{p}}^k = \mathbb{D}^{-1} \left(\boldsymbol{b} - (\mathbb{L} + \mathbb{U})\boldsymbol{x}^k \right)
# $$
#
# * Gauss-Seidel:
#
# $$
# \boldsymbol{\hat{p}}^k = \mathbb{D}^{-1} \left(-\mathbb{L} \boldsymbol{x}^{k+1} -\mathbb{U}\boldsymbol{x}^k + \boldsymbol{b} \right)
# $$
#
#
# +
def jacobi_with_relaxation(A,b,N,x0,alpha):
x = np.copy(x0)
d = diag(A)
residuals_norm = np.arange(N)
for i in range(N):
residual = b - A@x
residuals_norm[i] = np.linalg.norm(residual)
p = residual/d
x = x + alpha*p
return x, residuals_norm
def gauss_seidel_SOR(A,b,N,x0, alpha=1):
# https://en.wikipedia.org/wiki/Successive_over-relaxation
x = np.copy(x0)
d = diag(A)
L = np.tril(A,-1)
# D = diagflat(diag(A))
U = np.triu(A,1)
residuals_norm = np.arange(N)
for t in range(N):
residual = b - A@x
residuals_norm[t] = np.linalg.norm(residual)
for i in range(len(x)):
x[i] = (1. - alpha)*x[i] + alpha*(-L[i,:] @ x - U[i,:] @ x + b[i])/d[i]
return x, residuals_norm
x_jacobi_with_relaxation, residuals_norm_jacobi_with_relaxation = jacobi_with_relaxation(A,b,N,initial_guess,alpha)
print(f"jacobi_relaxation {alpha}: {x_jacobi_with_relaxation}")
x_GS_SOR, residuals_norm_GS_SOR = gauss_seidel_SOR(A,b,N,initial_guess,alpha)
print(f"gauss_seidel_SOR {alpha}: {x_GS_SOR}")
x_np = np.linalg.solve(A, b)
print(f"np.linalg.solve(A, b): {x_np}")
plt.rcParams.update({'font.size': 16})
figure, axis = plt.subplots(1, 1, figsize=(8, 6))
# plt.subplots_adjust(hspace=1)
axis.set_title('Residualse')
axis.plot(iterations, residuals_norm_jacobi_with_relaxation, label=r'residuals_norm_jacobi_with_relaxation')
axis.plot(iterations, residuals_norm_GS_SOR,label=r'gauss_seidel_SOR')
axis.set_yscale('log')
axis.set_xlabel('iterations')
axis.set_ylabel('residuals_norm')
axis.legend(loc="upper right")
# -
# ## Optimal relaxation rate
#
# It is immediately clear, that convergence rate depends on the $\alpha$ coefficient.
# It would be beneficial, to tune the coefficient in each iteration:
#
# $$
# \mathbf{x}^{k+1} = \mathbf{x}^{k} + \alpha^k \mathbf{p}^k
# $$
#
# How the residual evolve in each step?
# Let us multiply the formula above by $-\mathbf{A}$, next add $\mathbf{b}$ and use the definition of the residual.
#
# $$
# \mathbf{r}^{k+1} = \mathbf{r}^{k} - \alpha^k \mathbf{A} \mathbf{p}^k
# $$
#
# Square of the norm of the residual is equal to:
#
# $$
# \|\mathbf{r}^{k+1}\| = (\mathbf{r}^{k+1})^T \mathbf{r}^{k+1} =
# (\mathbf{r}^{k} - \alpha^k \mathbf{A} \mathbf{p}^k)^T (\mathbf{r}^{k} - \alpha^k \mathbf{A} \mathbf{p}^k) =
# $$
#
# $$
# (\mathbf{r}^{k})^T \mathbf{r}^{k} -
# 2 \alpha^k (\mathbf{r}^{k})^T \mathbf{A} \mathbf{p}^k +
# (\alpha^k)^2(\mathbf{A}\mathbf{p}^k)^T \mathbf{A}\mathbf{p}^k
# $$
#
# Observe, that the square of the norm of the residual is a quadratic function of $\alpha^k$.
# This function has a minimum, because the coefficient in front of $(\alpha^k)^2$ is positive.
# Let us calculate the derivative with respect to $\alpha^k$:
#
# $$
# \frac{d}{d\alpha^k} \left( \|\mathbf{r}^{k+1}\| \right) =
# -2(\mathbf{r}^{k})^T \mathbf{A} \mathbf{p}^k +
# 2 \alpha^k (\mathbf{A} \mathbf{p}^k)^T \mathbf{A} \mathbf{p}^k
# $$
#
# The minimum is when $\frac{d}{d\alpha^k} \left( \|\mathbf{r}^{k+1}\| \right) = 0$, which corresponds with
#
# $$
# \alpha^k = \frac{(\mathbf{r}^{k})^T \mathbf{A} \mathbf{p}^k}{(\mathbf{A} \mathbf{p}^k)^T \mathbf{A} \mathbf{p}^k}
# $$
#
# Scheme with such coefficient,$\alpha^k$, is called Minimal Residual Method --- **MINRES** (**metoda najmniejszych residuów**).
#
# +
def gauss_seidel_MINRES(A,b,N,x0):
x = np.copy(x0)
d = diag(A)
L = np.tril(A,-1)
D = diagflat(diag(A))
U = np.triu(A,1)
p = np.zeros(len(x))
residuals_norm = list()
convergence_criteria = 1e-6
iteration = 0
residual = b - A@x
while np.linalg.norm(residual) > convergence_criteria and iteration < N:
# while iteration < N: this fails when residuals --> 0
residual = b - A@x
residuals_norm.append(np.linalg.norm(residual))
for i in range(len(x)):
phat = (-L[i,:] @ x - U[i,:] @ x + b[i])/d[i]
# x[i] = (1. - alpha)*x[i] + alpha*phat
## OR
p[i] = phat - x[i]
alpha = residual.transpose() @ (A @ p)
alpha /= (A @ p).transpose() @ (A @ p)
x = x + alpha *p
iteration +=1
return x, residuals_norm
x_gauss_seidel_MINRES, residuals_norm_GS_MINRES = gauss_seidel_MINRES(A,b,N,initial_guess)
print(f"gauss_seidel_MINRES: {x_gauss_seidel_MINRES}")
x_np = np.linalg.solve(A, b)
print(f"np.linalg.solve(A, b): {x_np}")
# If the Hessian is positive definite
# (equivalently, has all eigenvalues positive) at a, then f=f(x)=Ax attains a local minimum at f(x=a).
def is_pos_def(x):
return np.all(np.linalg.eigvals(x) > 0)
if not is_pos_def(A):
print("A is not positive definite.")
plt.rcParams.update({'font.size': 16})
figure, axis = plt.subplots(1, 1, figsize=(8, 6))
# plt.subplots_adjust(hspace=1)
axis.set_title('Residuals')
axis.plot(iterations[:len(residuals_norm_GS_MINRES)], residuals_norm_GS_MINRES, label=r'residuals_norm_GS_MINRES')
axis.set_yscale('log')
axis.set_xlabel('iterations')
axis.set_ylabel('residual')
axis.legend(loc="upper right")
# -
# ## Questions:
#
# 1. How much memory is requirement to store $\boldsymbol{x}^{k+1}$ and $\boldsymbol{x}^k$ between iteration in Jacobi and Gauss-Seidel method?
#
# 2. How to parallelize the Jacobi and Gauss-Seidel method?
#
# 3. Why the Jacobi algorithm fails in case of the *ugly* matrix?
#
# 4a. Why the Minimal Residual Method (+Gauss-Seidel) fails in case of the *ugly* matrix?
#
# 4b. (extra?) Why the Minimal Residual Method converges slower the SOR with lucky $\alpha = 0.5$
#
# 5. What is the difference between the *ugly* and the *nice* matrix?
#
# 6. How the numerical routine like `solve(A,b)` the picks up the proper method?
#
# 7. Are there any special algorithm to store sparse matrices?
# ## Answers
#
# 1a. In Jacobi method both the $\boldsymbol{x}^{k+1}$ and $\boldsymbol{x}^k$ solution vectors have to be stored until the end of iteration. As a result, the required amount of memory is $memory = 2*len(x)*sizeof(float/double)$.
#
# 1b. In the Gauss-Seidel method just one solution vector, $x$, has to be stored as the new iteration ovewrites the previos one in an 'in place' manner.
#
# 2. The Gauss-Seidel method is sequential, thus it can not be parallelized.
#
# 3. The Jacobi Method fails in case of the ugly matrix, because it is not diagonally dominant.
#
# 4a. The Minimal Residual Method (+Gauss-Seidel) fails in case of the ugly matrix, because it is not positive definite.
#
# The optimal relaxation coefficient $\frac{d}{d\alpha^k} \left( \|\mathbf{r}^{k+1}\| \right) = 0$ fails to minimize the residuum. There is a saddle point!
#
# 4b. It seems that the reason is in not so lucky preconditioner? But at least it shall converge, while guessing $\alpha$ for large system maybe tedious...
#
# 5. The *nice* matrix is diagonally dominant and positive definite.
#
# 6. The numerical routine like `solve(A,b)` checks properties of the A matrix, the picks up the best method.
#
# 7. To store a sparse matrix in an efficient way, only the nonzero elements are saved.
#
# 8. (Extra). The MINRES methods is a variant of the conjugate gradient method that avoids the LU decomposition and does not suffer from breakdown. MINRES minimizes the residual in the 2-norm.
# ## References:
#
# 1. "<NAME> dla studentów technik informacyjnych", <NAME>, Warszawa 2013
# 2. "Computational Fluid Dynamics - lecture notes", <NAME>, Warszawa 2014
# <https://www.meil.pw.edu.pl/za/content/download/29896/156221/file/ComputationalFluidDynamics_20140910.pdf>
# 3. <NAME> -laboratorium 3, <NAME>,
# <http://ccfd.github.io/courses/metnum_lab3.html>
# 4. <https://www.quantstart.com/articles/Jacobi-Method-in-Python-and-NumPy/>
# 5. <https://en.wikipedia.org/wiki/Successive_over-relaxation>
#
# ## Historical notes
#
# * Human Computers: <https://en.wikipedia.org/wiki/Computer_(job_description)>
#
# * "Pan raczy żartować, panie Feynman!" <NAME> (Original title: „Surely You're Joking, Mr. Feynman!”: Adventures of a Curious Character, Princeton University Press, USA 1985)
#
|
lab3_iterative_solvers_diffusion_and_laplace_eq/.ipynb_checkpoints/SOLUTION_iterative_solvers-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import bib_parse
cited = bib_parse.bib_to_df('../bibtex/2017-02-01-times_cited_descending.bib')
newest = bib_parse.bib_to_df('../bibtex/2017-02-01-publication_date_newest.bib')
oldest = bib_parse.bib_to_df('../bibtex/2017-02-01-publication_date_oldest.bib')
c180 = bib_parse.bib_to_df('../bibtex/2017-02-01-useage_count_180.bib')
c2013 = bib_parse.bib_to_df('../bibtex/2017-02-01-useage_count_2013.bib')
cited.columns
of_interest = pd.merge(left=cited, right=pd.DataFrame(c2013['unique-id']), on='unique-id', how='inner')
of_interest[['title', 'author', 'times-cited', 'year', 'journal',
'usage-count-since-2013', 'usage-count-(last-180-days)',
'web-of-science-categories']].\
sort_values('year', ascending=False)
|
src/2017-02-01-initial_papers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Importing pandas module
import pandas as pd
# Importing numpy module
import numpy as np
# Importing train_test_split method from sklearn.model_selection
from sklearn.model_selection import train_test_split
# Importing preprocsessing method from sklearn
from sklearn import preprocessing
# Importing RandomForest module from sklearn.ensemble
from sklearn.ensemble import RandomForestRegressor
# Importing cross-validation pipeline
from sklearn.pipeline import make_pipeline
from sklearn.model_selection import GridSearchCV
# Importing evaluation metrics
from sklearn.metrics import mean_squared_error, r2_score
# Importing module to save sklearn models
from sklearn.externals import joblib
# Loading wine data using url
dataset_url = 'http://mlr.cs.umass.edu/ml/machine-learning-databases/wine-quality/winequality-red.csv'
data = pd.read_csv(dataset_url)
# printing sample data
data.head()
# Cleaning the data
data = pd.read_csv(dataset_url, sep=';')
data.head()
# shape of the data
data.shape
# describing the data
data.describe()
# Features of the data
data.columns
# target feature of the data
y = data.quality
# input features of the data
x = data.drop('quality', axis = 1)
# spliting data into train and test sets
x_train, x_test, y_train, y_test = train_test_split(x,y,test_size = 0.2, random_state = 123, stratify = y)
# Scaling the trained data
x_train_scaled = preprocessing.scale(x_train)
print(x_train_scaled)
# verifying the scaled data
print(x_train_scaled.mean(axis = 0))
print(x_train_scaled.std(axis = 0))
# Fitting the transformer API
scaler = preprocessing.StandardScaler().fit(x_train)
print(scaler)
# Applying Transformer to trained data
x_train_scaled = scaler.transform(x_train)
print(x_train_scaled.mean(axis = 0))
print(x_train_scaled.std(axis = 0))
# Applying Transformer to test data
x_test_scaled = scaler.transform(x_test)
print(x_test_scaled.mean(axis = 0))
print(x_test_scaled.std(axis = 0))
# Creating the pipeline
pipeline = make_pipeline(preprocessing.StandardScaler(), RandomForestRegressor(n_estimators = 100))
print(pipeline)
# printing tunable hyper parameters
print(pipeline.get_params())
# Tuning the hyperparameters
hyperparameters = {'randomforestregressor__max_features':['auto','sqrt','log2'],
'randomforestregressor__max_depth':[None,5,3,1]}
print(hyperparameters)
# Cross validation with pipeline
clf = GridSearchCV(pipeline, hyperparameters, cv = 10)
clf.fit(x_train, y_train)
# printing the best parameters
print(clf.best_params_)
# retraining the model
print(clf.refit)
# Predicting new set of data
y_pred = clf.predict(x_test)
print(y_pred)
# Evaluating model performance
print(r2_score(y_test,y_pred))
print(mean_squared_error(y_test,y_pred))
# Saving the model
joblib.dump(clf,'rf_regressor.pkl')
|
Wine Quality Prediction (Random Forest).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import seaborn as sb
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn import metrics
from sklearn.neighbors import KNeighborsClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
df=pd.read_csv("F:\\ML_DATASETS\\Iris_data_sample.csv")
df1=pd.read_csv("F:\\ML_DATASETS\\Iris_data_sample.csv",index_col=0) # index_col is used to remove first column
df1
df1.head()
# # UNIVARIATE ANALYSIS
df1_setosa=df1.loc[df1['Species']=='Iris-setosa']
df1_virginica=df1.loc[df1['Species']=='Iris-virginica']
df1_versicolor=df1.loc[df1['Species']=='Iris-versicolor']
plt.plot(df1_setosa['SepalLengthCm'],np.zeros_like(df1_setosa['SepalLengthCm']))
plt.plot(df1_virginica['SepalLengthCm'],np.zeros_like(df1_virginica['SepalLengthCm']))
plt.plot(df1_versicolor['SepalLengthCm'],np.zeros_like(df1_versicolor['SepalLengthCm']))
plt.xlabel('Petal Length')
plt.show()
sb.FacetGrid(df1,hue="Species",height=5).map(plt.scatter,"SepalLengthCm","SepalWidthCm").add_legend() # addlegend-shows the species in right of graph
plt.show()
# # MULTIVARIATE ANALYSIS
sb.pairplot(df1,hue="Species",height=5)
df1.shape
df1['Species'].value_counts()
df1.isnull() # to check null values
# # Training the model using knnearest classification and modeling with scikit learn
x=df1.iloc[:,:-1] #data
y=df1.iloc[:,4] #target
X_train,X_test,y_train,y_test=train_test_split(x,y,random_state=0)
knn=KNeighborsClassifier(n_neighbors=1)
knn.fit(X_train,y_train)
x_new=np.array([[5,2.9,1,0.3]])
print("X_new.shape:",x_new.shape)
prediction=knn.predict(x_new)
print("prediction: ",prediction)
#print("Predicted target name:",df1[y])
# # Evaluating the model whether the prediction is correct or not
y_pred=knn.predict(X_test)
print('Test set prediction : ',y_pred)
print("Test set score:",np.mean(y_pred==y_test))
print("Test set score:",knn.score(X_test,y_test))
|
Iris_Dataset.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Customer Survival Analysis
# ## Theory
# If time to event has the probability density function $f(t)$ and cumulative distribution function $F(t)$, then the probability of surviving at least to time $t$ is: $Pr(T>t)=S(t)=1-F(t)$.
#
# Cumulative hazard at time t is defined as $H(t)=-ln(S(t))$ and instantaneous hazard at time $t$ is $h(t)=\frac{dH(t)}{dt}$. The instantateous hazard can also be written as $h(t)=\frac{f(t)}{S(t)}$
#
# The likelihood function for survival analysis is described as:
#
# $$ l(\beta) = \prod_{n=1}^{n} h(t_{i})^{d_{i}} S(t_{i}) $$
# where $d_i$ is the censoring variable that equals to 1 if the event is observed for individual $i$ and 0 if the event is not observed (censored) for individual $i$, $h(t_i)$ is the hazard for individual $i$ at time $t$, $H(t_i)$ is the cumulative hazard for individual $i$ at time $t$, and $S(t_i)$ is the survival probability for individual $i$ at time $t$. Note that when $d_i=0$, the contribution of the $i$'th individual to the likelihood function is just its survival probability until time $t$: S(t). If the individual has the event, the contribution to the likelihood function is given by the density function $f(t)=h(t)S(t)$.
#
# The log of likelihood is:
#
# $$ logl(\beta) = \sum_{i=1}^n d_i log(h(t_i)) - H(t_i) $$
# where $log$ is the natural logarithm.
# ## Importing Libraries
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from scipy.stats import norm
import statsmodels.api as st
from sklearn.preprocessing import LabelEncoder
labelencoder = LabelEncoder()
#Lifelines is a survival analysis package
from lifelines import KaplanMeierFitter
from lifelines.statistics import multivariate_logrank_test
from lifelines.statistics import logrank_test
from lifelines import CoxPHFitter
# -
# ## Data Preparation
df = pd.read_csv("C:/Data/Telco-Customer-Churn.csv")
df.head()
df.info()
# Here, Churn is an event which indicates whether customer exited or not. Tenure shows how long customer remained in our service. Both of these variables are very important for customer survival analysis.
df.Churn = labelencoder.fit_transform(df.Churn)
df.Churn.value_counts()
eventvar = df['Churn']
timevar = df['tenure']
# For the analysis, I will need to create dummy variables for all categorical variables.
# +
categorical = ['gender', 'SeniorCitizen', 'Partner', 'Dependents', 'PhoneService', 'MultipleLines',
'InternetService', 'OnlineSecurity', 'OnlineBackup', 'DeviceProtection',
'TechSupport', 'StreamingTV', 'StreamingMovies', 'Contract',
'PaperlessBilling', 'PaymentMethod']
survivaldata = pd.get_dummies(df, columns = categorical, drop_first= True)
survivaldata.head()
# -
# We need to drop variables such as customerID, tenure, Churn as they are not needed in survival data. Also, we need to add constant for survival analysis.
survivaldata.drop(['customerID', 'tenure', 'Churn'], axis = 1, inplace= True)
survivaldata = st.add_constant(survivaldata, prepend=False)
survivaldata.head()
# ## Survival Analysis
# ### Kaplan-Meier Curve
# The Kaplan-Meier method calculates the probability of survival at time 𝑡 as:
#
# $$ S(t) = \prod_{i=1}^{t-1} (1 - \frac{d_i}{n_i}) $$
#
# where,
# - 𝑆(𝑡) is the probability of survival until time 𝑡,
# - $𝑑_𝑖$ is the number of units that experienced the event at time 𝑡,
# - $𝑛_𝑖$ is the number of units at risk of experiencing the event at time 𝑡.
#
# $𝑛_𝑖$ decreases with time, as units experience the event or are censored. $\frac{d_i}{n_i}$ is the probability of experiencing the event at time 𝑖 and $(1− \frac{d_i}{n_i})$ is the probability of surviving at time 𝑖.
#
# Note that this method does not use any parameters, it only depends on the data on time and censoring.
#Create a KaplanMeier object, imported from lifelines
kmf = KaplanMeierFitter()
#Calculate the K-M curve for all groups
kmf.fit(timevar,event_observed = eventvar,label = "All Customers")
#Plot the curve and assign labels
kmf.plot()
plt.ylabel('Probability of Customer Survival')
plt.xlabel('Tenure')
plt.title('Kaplan-Meier Curve');
# As we can see, there is a sudden drop in the starting which says that after one tenure only customers starts churning rapidly and after that churning rate decreases. To deal with that we can consider giving more discounts on long-term plans and make more customers to subscribe for long term plans.
# ### Log-Rank Test
# We can use non-parametric method log-rank test to compare survival curves between different groups. The log-rank test assumes that the hazards of the groups are proportional. Under the null hypothesis, the probability of event across the groups is the same for all time points.
#
# To test the null hypothesis, the log-rank test calculates the difference between the observed number of events and the number of events expected in each group proportional to the size of the groups at each time point an event is observed. The log-rank test statistic for group $j$ $(k_{j})$ follows a $\chi^2$ distribution and is calculated as:
#
# $$k_{j} = \frac{(O_{j}-E_{j})^{2}}{var(O_{j}-E_{j})}$$
#
# $O_{j}-E_{j}$ is calculated as:
#
# $$O_{j}-E_{j} = \sum_{i}(o_{ij}-e_{ij})$$
#
# and $var(O_{j}-E_{j})$ is:
#
# $$var(O_{j}-E_{j}) = o_{i}\frac{n_{ij}}{n_{i}}\Big(1-\frac{n_{ij}}{n_{i}}\Big)\frac{(n_{i}-o_{i})}{(n_{i}-1)}$$
#
# $o_{ij}$ is the observed number of events in group $j$ at time $i$ and $e_{ij}$ is the expected number of events in group $j$ at time $i$, which is calculated as $e_{ij} = \frac{n_{ij}}{n_i}{o_{i}}$. Note that $\frac{n_{ij}}{n_i}$ is the proportion of units in group $j$ at risk of event at time $i$ ($n_{ij}$) to the number of units in all groups at risk of event at time $i$ ($n_{i}$) and ${o_{i}}$ is the observed number of events in all groups at time $i$.
# When comparing multiple groups, we first calculate the pairwise log-rank test scores between each of the $k-1$ groups, and write them as a vector of log-rank statistics, $\bf{Z}$ which has $k - 1$ elements. We can leave any of one of the statistics out, because the $k$ covariances are linearly dependent on each other (the sum of log-rank statistics is 0, $\sum k_{j}=0$.
#
# The test statistic for the hypothesis that there is no difference in survival times of $k$ groups is calculated as:
#
# $$logrankstatistic = \bf{Z} {\sum}^{-1} \bf{Z}'$$
#
# which has a $\chi^2$ distribution, where ${\sum}^{-1}$ is the inverse of the $k-1$ by $k-1$ variance-covariance matrix of $\bf{Z}$, which has variance of $k_{j}$ on its diagonal elements and $covar(k_{jg})$ on its off-diagonal elements.
#
# The variance of observed number of events in group $j$ is calculated as $var(O_{j}-E_{j})$ as demonstrated above. The covariance between the observed number of events in group $j$ and $g$ is calculated as:
#
# $$covar(k_{jg})=o_{i}\frac{(n_{ij}n_{ig})}{(n_{i}n_{i})}\frac{(n_{i}-o_{i})}{(n_{i}-1)}$$
#
# Note that rejecting the null hypothesis means that the survival times of groups do not come from the same distribution, it does not specify which group's survival time is different. The following plots and test statistics compare the groups in the dataset in terms of the different explanatory variables. Astatistically significant log-rank test statistic indicates that we can reject the null hypothesis that time to survival in all groups come from the same distribution.
# ### Gender
# +
male = (survivaldata['gender_Male'] == 1)
female = (survivaldata['gender_Male'] == 0)
plt.figure()
ax = plt.subplot(1,1,1)
kmf.fit(timevar[male],event_observed = eventvar[male],label = "Male")
plot1 = kmf.plot(ax = ax)
kmf.fit(timevar[female],event_observed = eventvar[female],label = "Female")
plot2 = kmf.plot(ax = plot1)
plt.title('Survival of customers: Gender')
plt.xlabel('Tenure')
plt.ylabel('Survival Probability')
plt.yticks(np.linspace(0,1,11))
groups = logrank_test(timevar[male], timevar[female], event_observed_A=eventvar[male], event_observed_B=eventvar[female])
groups.print_summary()
# -
# ### Senior Citizen
# +
SeniorCitizen = (survivaldata['SeniorCitizen_1'] == 1)
no_SeniorCitizen = (survivaldata['SeniorCitizen_1'] == 0)
plt.figure()
ax = plt.subplot(1,1,1)
kmf.fit(timevar[SeniorCitizen],event_observed = eventvar[SeniorCitizen],label = "Senior Citizen")
plot1 = kmf.plot(ax = ax)
kmf.fit(timevar[no_SeniorCitizen],event_observed = eventvar[no_SeniorCitizen],label = "Not a Senior Citizen")
plot2 = kmf.plot(ax = plot1)
plt.title('Survival of customers: Senior Citizen')
plt.xlabel('Tenure')
plt.ylabel('Survival Probability')
plt.yticks(np.linspace(0,1,11))
groups = logrank_test(timevar[SeniorCitizen], timevar[no_SeniorCitizen], event_observed_A=eventvar[SeniorCitizen], event_observed_B=eventvar[no_SeniorCitizen])
groups.print_summary()
# -
# ### Partner
# +
partner = (survivaldata['Partner_Yes'] == 1)
no_partner = (survivaldata['Partner_Yes'] == 0)
plt.figure()
ax = plt.subplot(1,1,1)
kmf.fit(timevar[partner],event_observed = eventvar[partner],label = "Has partner")
plot1 = kmf.plot(ax = ax)
kmf.fit(timevar[no_partner],event_observed = eventvar[no_partner],label = "Does not have a partner")
plot2 = kmf.plot(ax = plot1)
plt.title('Survival of customers: Partner')
plt.xlabel('Tenure')
plt.ylabel('Survival Probability')
plt.yticks(np.linspace(0,1,11))
groups = logrank_test(timevar[partner], timevar[no_partner], event_observed_A=eventvar[partner], event_observed_B=eventvar[no_partner])
groups.print_summary()
# -
# ### Dependents
# +
Dependents = (survivaldata['Dependents_Yes'] == 1)
no_Dependents = (survivaldata['Dependents_Yes'] == 0)
plt.figure()
ax = plt.subplot(1,1,1)
kmf.fit(timevar[Dependents],event_observed = eventvar[Dependents],label = "Has dependents")
plot1 = kmf.plot(ax = ax)
kmf.fit(timevar[no_Dependents],event_observed = eventvar[no_Dependents],label = "Does not have dependents")
plot2 = kmf.plot(ax = plot1)
plt.title('Survival of customers: Dependents')
plt.xlabel('Tenure')
plt.ylabel('Survival Probability')
plt.yticks(np.linspace(0,1,11))
groups = logrank_test(timevar[Dependents], timevar[no_Dependents], event_observed_A=eventvar[Dependents], event_observed_B=eventvar[no_Dependents])
groups.print_summary()
# -
# ### PhoneService
# +
PhoneService = (survivaldata['PhoneService_Yes'] == 1)
no_PhoneService = (survivaldata['PhoneService_Yes'] == 0)
plt.figure()
ax = plt.subplot(1,1,1)
kmf.fit(timevar[PhoneService],event_observed = eventvar[PhoneService],label = "Has a phone service")
plot1 = kmf.plot(ax = ax)
kmf.fit(timevar[no_PhoneService],event_observed = eventvar[no_PhoneService],label = "Does not have a phone service")
plot2 = kmf.plot(ax = plot1)
plt.title('Survival of customers: Phone Service')
plt.xlabel('Tenure')
plt.ylabel('Survival Probability')
plt.yticks(np.linspace(0,1,11))
groups = logrank_test(timevar[PhoneService], timevar[no_PhoneService], event_observed_A=eventvar[PhoneService], event_observed_B=eventvar[no_PhoneService])
groups.print_summary()
# -
# ### MultipleLines
# +
no_phone = (survivaldata['MultipleLines_No phone service'] == 1)
multiLines = (survivaldata['MultipleLines_Yes'] == 1)
no_multiLines = ((survivaldata['MultipleLines_Yes'] == 0) & (survivaldata['MultipleLines_No phone service'] == 0))
plt.figure()
ax = plt.subplot(1,1,1)
kmf.fit(timevar[no_phone],event_observed = eventvar[no_phone],label = "No Phone Service")
plot1 = kmf.plot(ax = ax)
kmf.fit(timevar[multiLines],event_observed = eventvar[multiLines],label = "Multiple Lines")
plot2 = kmf.plot(ax = plot1)
kmf.fit(timevar[no_multiLines],event_observed = eventvar[no_multiLines],label = "Single Line")
plot3 = kmf.plot(ax = plot2)
plt.title('Survival of customers: Mutliple Lines')
plt.xlabel('Tenure')
plt.ylabel('Survival Probability')
plt.yticks(np.linspace(0,1,11))
twoplusgroups_logrank = multivariate_logrank_test(df['tenure'], df['MultipleLines'], df['Churn'], alpha = 0.95)
twoplusgroups_logrank.print_summary()
# -
# ### Internet Service
# +
Fiber_optic = (survivaldata['InternetService_Fiber optic'] == 1)
No_Service = (survivaldata['InternetService_No'] == 1)
DSL = ((survivaldata['InternetService_Fiber optic'] == 0) & (survivaldata['InternetService_No'] == 0))
plt.figure()
ax = plt.subplot(1,1,1)
kmf.fit(timevar[Fiber_optic],event_observed = eventvar[Fiber_optic],label = "Fiber optic")
plot1 = kmf.plot(ax = ax)
kmf.fit(timevar[No_Service],event_observed = eventvar[No_Service],label = "No Service")
plot2 = kmf.plot(ax = plot1)
kmf.fit(timevar[DSL],event_observed = eventvar[DSL],label = "DSL")
plot3 = kmf.plot(ax = plot2)
plt.title('Survival of customers: Internet Service')
plt.xlabel('Tenure')
plt.ylabel('Survival Probability')
plt.yticks(np.linspace(0,1,11))
twoplusgroups_logrank = multivariate_logrank_test(df['tenure'], df['InternetService'], df['Churn'], alpha = 0.95)
twoplusgroups_logrank.print_summary()
# -
# ### Online Security
# +
no_internetService = (survivaldata['OnlineSecurity_No internet service'] == 1)
onlineSecurity = (survivaldata['OnlineSecurity_Yes'] == 1)
no_onlineSecurity = ((survivaldata['OnlineSecurity_No internet service'] == 0) & (survivaldata['OnlineSecurity_Yes'] == 0))
plt.figure()
ax = plt.subplot(1,1,1)
kmf.fit(timevar[no_internetService],event_observed = eventvar[no_internetService],label = "No Internet Service")
plot1 = kmf.plot(ax = ax)
kmf.fit(timevar[onlineSecurity],event_observed = eventvar[onlineSecurity],label = "Online Security")
plot2 = kmf.plot(ax = plot1)
kmf.fit(timevar[no_onlineSecurity],event_observed = eventvar[no_onlineSecurity],label = "No online Security")
plot3 = kmf.plot(ax = plot2)
plt.title('Survival of customers: Online Security')
plt.xlabel('Tenure')
plt.ylabel('Survival Probability')
plt.yticks(np.linspace(0,1,11))
twoplusgroups_logrank = multivariate_logrank_test(df['tenure'], df['OnlineSecurity'], df['Churn'], alpha = 0.95)
twoplusgroups_logrank.print_summary()
# -
# ### Online Backup
# +
no_internetService = (survivaldata['OnlineBackup_No internet service'] == 1)
onlineBackup = (survivaldata['OnlineBackup_Yes'] == 1)
no_onlineBackup = ((survivaldata['OnlineBackup_No internet service'] == 0) & (survivaldata['OnlineBackup_Yes'] == 0))
plt.figure()
ax = plt.subplot(1,1,1)
kmf.fit(timevar[no_internetService],event_observed = eventvar[no_internetService],label = "No Internet Service")
plot1 = kmf.plot(ax = ax)
kmf.fit(timevar[onlineBackup],event_observed = eventvar[onlineBackup],label = "Online Backup")
plot2 = kmf.plot(ax = plot1)
kmf.fit(timevar[no_onlineBackup],event_observed = eventvar[no_onlineBackup],label = "No online Backup")
plot3 = kmf.plot(ax = plot2)
plt.title('Survival of customers: Online Backup')
plt.xlabel('Tenure')
plt.ylabel('Survival Probability')
plt.yticks(np.linspace(0,1,11))
twoplusgroups_logrank = multivariate_logrank_test(df['tenure'], df['OnlineBackup'], df['Churn'], alpha = 0.95)
twoplusgroups_logrank.print_summary()
# -
# ### Device Protection
# +
no_internetService = (survivaldata['DeviceProtection_No internet service'] == 1)
DeviceProtection = (survivaldata['DeviceProtection_Yes'] == 1)
no_DeviceProtection = ((survivaldata['DeviceProtection_No internet service'] == 0) & (survivaldata['DeviceProtection_Yes'] == 0))
plt.figure()
ax = plt.subplot(1,1,1)
kmf.fit(timevar[no_internetService],event_observed = eventvar[no_internetService],label = "No Internet Service")
plot1 = kmf.plot(ax = ax)
kmf.fit(timevar[DeviceProtection],event_observed = eventvar[DeviceProtection],label = "Device Protection")
plot2 = kmf.plot(ax = plot1)
kmf.fit(timevar[no_DeviceProtection],event_observed = eventvar[no_DeviceProtection],label = "No Device Protection")
plot3 = kmf.plot(ax = plot2)
plt.title('Survival of customers: Device Protection')
plt.xlabel('Tenure')
plt.ylabel('Survival Probability')
plt.yticks(np.linspace(0,1,11))
twoplusgroups_logrank = multivariate_logrank_test(df['tenure'], df['DeviceProtection'], df['Churn'], alpha = 0.95)
twoplusgroups_logrank.print_summary()
# -
# ### Tech Support
# +
no_internetService = (survivaldata['TechSupport_No internet service'] == 1)
TechSupport = (survivaldata['TechSupport_Yes'] == 1)
no_TechSupport = ((survivaldata['TechSupport_No internet service'] == 0) & (survivaldata['TechSupport_Yes'] == 0))
plt.figure()
ax = plt.subplot(1,1,1)
kmf.fit(timevar[no_internetService],event_observed = eventvar[no_internetService],label = "No Internet Service")
plot1 = kmf.plot(ax = ax)
kmf.fit(timevar[TechSupport],event_observed = eventvar[TechSupport],label = "Tech Support")
plot2 = kmf.plot(ax = plot1)
kmf.fit(timevar[no_TechSupport],event_observed = eventvar[no_TechSupport],label = "No Tech Support")
plot3 = kmf.plot(ax = plot2)
plt.title('Survival of customers: Tech Support')
plt.xlabel('Tenure')
plt.ylabel('Survival Probability')
plt.yticks(np.linspace(0,1,11))
twoplusgroups_logrank = multivariate_logrank_test(df['tenure'], df['TechSupport'], df['Churn'], alpha = 0.95)
twoplusgroups_logrank.print_summary()
# -
# ### Streaming TV
# +
no_internetService = (survivaldata['StreamingTV_No internet service'] == 1)
StreamingTV = (survivaldata['StreamingTV_Yes'] == 1)
no_StreamingTV = ((survivaldata['StreamingTV_No internet service'] == 0) & (survivaldata['StreamingTV_Yes'] == 0))
plt.figure()
ax = plt.subplot(1,1,1)
kmf.fit(timevar[no_internetService],event_observed = eventvar[no_internetService],label = "No Internet Service")
plot1 = kmf.plot(ax = ax)
kmf.fit(timevar[StreamingTV],event_observed = eventvar[StreamingTV],label = "Streaming TV")
plot2 = kmf.plot(ax = plot1)
kmf.fit(timevar[no_StreamingTV],event_observed = eventvar[no_StreamingTV],label = "No Streaming TV")
plot3 = kmf.plot(ax = plot2)
plt.title('Survival of customers: Streaming TV')
plt.xlabel('Tenure')
plt.ylabel('Survival Probability')
plt.yticks(np.linspace(0,1,11))
twoplusgroups_logrank = multivariate_logrank_test(df['tenure'], df['StreamingTV'], df['Churn'], alpha = 0.95)
twoplusgroups_logrank.print_summary()
# -
# ### Streaming Movies
# +
no_internetService = (survivaldata['StreamingMovies_No internet service'] == 1)
StreamingMovies = (survivaldata['StreamingMovies_Yes'] == 1)
no_StreamingMovies = ((survivaldata['StreamingMovies_No internet service'] == 0) & (survivaldata['StreamingMovies_Yes'] == 0))
plt.figure()
ax = plt.subplot(1,1,1)
kmf.fit(timevar[no_internetService],event_observed = eventvar[no_internetService],label = "No Internet Service")
plot1 = kmf.plot(ax = ax)
kmf.fit(timevar[StreamingMovies],event_observed = eventvar[StreamingMovies],label = "Streaming Movies")
plot2 = kmf.plot(ax = plot1)
kmf.fit(timevar[no_StreamingMovies],event_observed = eventvar[no_StreamingMovies],label = "No Streaming Movies")
plot3 = kmf.plot(ax = plot2)
plt.title('Survival of customers: Streaming Movies')
plt.xlabel('Tenure')
plt.ylabel('Survival Probability')
plt.yticks(np.linspace(0,1,11))
twoplusgroups_logrank = multivariate_logrank_test(df['tenure'], df['StreamingMovies'], df['Churn'], alpha = 0.95)
twoplusgroups_logrank.print_summary()
# -
# ### Contract
# +
Contract_One_year = (survivaldata['Contract_One year'] == 1)
Contract_Two_year = (survivaldata['Contract_Two year'] == 1)
Contract_month_to_month = ((survivaldata['Contract_One year'] == 0) & (survivaldata['Contract_Two year'] == 0))
plt.figure()
ax = plt.subplot(1,1,1)
kmf.fit(timevar[Contract_One_year],event_observed = eventvar[Contract_One_year],label = "One year Contract")
plot1 = kmf.plot(ax = ax)
kmf.fit(timevar[Contract_Two_year],event_observed = eventvar[Contract_Two_year],label = "Two year Contract")
plot2 = kmf.plot(ax = plot1)
kmf.fit(timevar[Contract_month_to_month],event_observed = eventvar[Contract_month_to_month],label = "Month to month Contract")
plot3 = kmf.plot(ax = plot2)
plt.title('Survival of customers: Contract')
plt.xlabel('Tenure')
plt.ylabel('Survival Probability')
plt.yticks(np.linspace(0,1,11))
twoplusgroups_logrank = multivariate_logrank_test(df['tenure'], df['Contract'], df['Churn'], alpha = 0.95)
twoplusgroups_logrank.print_summary()
# -
# ### Payment Method
# +
automatic_Credit_Card = (survivaldata['PaymentMethod_Credit card (automatic)'] == 1)
electronic_check = (survivaldata['PaymentMethod_Electronic check'] == 1)
mailed_check = (survivaldata['PaymentMethod_Mailed check'] == 1)
automatic_Bank_Transfer = ((survivaldata['PaymentMethod_Credit card (automatic)'] == 0) & (survivaldata['PaymentMethod_Electronic check'] == 0) & (survivaldata['PaymentMethod_Mailed check'] == 0))
plt.figure()
ax = plt.subplot(1,1,1)
kmf.fit(timevar[automatic_Credit_Card],event_observed = eventvar[automatic_Credit_Card],label = "Automatic Credit card Payment")
plot1 = kmf.plot(ax = ax)
kmf.fit(timevar[electronic_check],event_observed = eventvar[electronic_check],label = "Electronic Check")
plot2 = kmf.plot(ax = plot1)
kmf.fit(timevar[mailed_check],event_observed = eventvar[mailed_check],label = "Mailed_check")
plot3 = kmf.plot(ax = plot2)
kmf.fit(timevar[automatic_Bank_Transfer],event_observed = eventvar[automatic_Bank_Transfer],label = "Automatic Bank Transfer")
plot4 = kmf.plot(ax = plot3)
plt.title('Survival of customers: PaymentMethod')
plt.xlabel('Tenure')
plt.ylabel('Survival Probability')
plt.yticks(np.linspace(0,1,11))
twoplusgroups_logrank = multivariate_logrank_test(df['tenure'], df['PaymentMethod'], df['Churn'], alpha = 0.95)
twoplusgroups_logrank.print_summary()
# -
# ### Paperless Billing
# +
PaperlessBilling = (survivaldata['PaperlessBilling_Yes'] == 1)
no_PaperlessBilling = (survivaldata['PaperlessBilling_Yes'] == 0)
plt.figure()
ax = plt.subplot(1,1,1)
kmf.fit(timevar[PaperlessBilling],event_observed = eventvar[PaperlessBilling],label = "Paperless Billing")
plot1 = kmf.plot(ax = ax)
kmf.fit(timevar[no_PhoneService],event_observed = eventvar[no_PhoneService],label = "No Paperless Billing")
plot2 = kmf.plot(ax = plot1)
plt.title('Survival of customers: Paperless Billing')
plt.xlabel('Tenure')
plt.ylabel('Survival Probability')
plt.yticks(np.linspace(0,1,11))
groups = logrank_test(timevar[PaperlessBilling], timevar[no_PaperlessBilling], event_observed_A=eventvar[PaperlessBilling], event_observed_B=eventvar[no_PaperlessBilling])
groups.print_summary()
# -
# ## Survival Regression
def datapreparation(filepath):
df = pd.read_csv(filepath)
df.drop(["customerID"], inplace = True, axis = 1)
df.TotalCharges = df.TotalCharges.replace(" ",np.nan)
df.TotalCharges.fillna(0, inplace = True)
df.TotalCharges = df.TotalCharges.astype(float)
cols1 = ['Partner', 'Dependents', 'PaperlessBilling', 'Churn', 'PhoneService']
for col in cols1:
df[col] = df[col].apply(lambda x: 0 if x == "No" else 1)
df.gender = df.gender.apply(lambda x: 0 if x == "Male" else 1)
df.MultipleLines = df.MultipleLines.map({'No phone service': 0, 'No': 0, 'Yes': 1})
cols2 = ['OnlineSecurity', 'OnlineBackup', 'DeviceProtection', 'TechSupport', 'StreamingTV', 'StreamingMovies']
for col in cols2:
df[col] = df[col].map({'No internet service': 0, 'No': 0, 'Yes': 1})
df = pd.get_dummies(df, columns=['InternetService', 'Contract', 'PaymentMethod'], drop_first=True)
return df
regression_df = datapreparation("C:/Data/Telco-Customer-Churn.csv")
regression_df.head()
# #### Survival Regression Ananlysis using Cox Proportional Hazard model
# +
cph = CoxPHFitter()
cph.fit(regression_df, duration_col='tenure', event_col='Churn')
cph.print_summary()
# -
cph.score_
fig, ax = plt.subplots(figsize = (10,7))
cph.plot(ax = ax);
test_id = regression_df.sample(1)
fig, ax = plt.subplots()
cph.predict_cumulative_hazard(test_id).plot(ax = ax, color = 'red')
plt.axvline(x=test_id.tenure.values[0], color = 'blue', linestyle='--')
plt.legend(labels=['Hazard','Current Position'])
ax.set_xlabel('Tenure', size = 10)
ax.set_ylabel('Cumulative Hazard', size = 10)
ax.set_title('Cumulative Hazard Over Time');
fig, ax = plt.subplots()
cph.predict_survival_function(test_id).plot(ax = ax, color = 'red')
plt.axvline(x=test_id.tenure.values[0], color = 'blue', linestyle='--')
plt.legend(labels=['Survival Function','Current Position'])
ax.set_xlabel('Tenure', size = 10)
ax.set_ylabel('Survival Probability', size = 10)
ax.set_title('Survival Probability Over Time');
# Saving the model
import pickle
pickle.dump(cph, open('survivemodel.pkl','wb'))
# ## Customer Lifetime Value
# To calculate customer lifetime value, I would multiply the Monthly charges the customer is paying to Telcom and the expected life time of the customer.
#
# I utilize the survival function of a customer to calculate its expected life time. I would like to be little bit conservative and consider the customer is churned when the survival probability of him is 10%.
def LTV(info):
life = cph.predict_survival_function(info).reset_index()
life.columns = ['Tenure', 'Probability']
max_life = life.Tenure[life.Probability > 0.1].max()
LTV = max_life * info['MonthlyCharges'].values[0]
return LTV
print('LTV of a testid is:', LTV(test_id), 'dollars.')
|
Customers Survival Analysis orignal.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import requests
# ## INPUT
#
# Here, to use a file for reading words, comment the input line, and remove comment # from the rest of the lines
words = input('Enter word to get suggestions for!\n')
# f = open('./../DATASET/SearchSuggestionKeywords.txt', 'r')
# words = []
# for w in f:
# words.append(w.replace('\n', '').replace(' ', '+'))
# +
suggestions = []
for w in words:
url = 'https://completion.amazon.co.uk/api/2017/suggestions?session-id=258-8222884-0745536&customer-id=A1DWR5F96BKA68&request-id=5HNHF7Y14AJTCZTPAGHA&page-type=Gateway&lop=en_IN&site-variant=desktop&client-info=amazon-search-ui&mid=A21TJRUUN4KGV&alias=aps&b2b=0&fresh=0&ks=80&prefix='+w+'&event=onKeyPress&limit=11&fb=1&suggestion-type=KEYWORD&suggestion-type=WIDGET&_=1596303646507'
source = requests.get(url).json()
sugg = {}
idx = 1
for s in source['suggestions']:
sugg[idx]= s['value']
idx +=1
print(sugg)
suggestions.append({w:sugg})
# -
# ## Output
#
# Suggestions is now a list of dictionaries, where the key is the word used and the value is a dictionary containing its respective search suggestions.
print(suggestions)
|
SCRIPTS/SearchSuggestions.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Q#
# language: qsharp
# name: iqsharp
# ---
# # Ripple Carry Adder Kata
#
# The **Ripple Carry Adder** quantum kata is a series of exercises designed
# to get you familiar with ripple carry addition on a quantum computer.
#
# * The simplest quantum adder, covered in part I, closely mirrors its classical counterpart,
# using the same basic components and the same algorithm.
# * Part II explores building an in-place adder.
# * A more complex version of an in-place adder covered in part III of the kata uses a different algorithm
# to reduce the number of ancillary qubits needed.
# * Finally, part IV covers building an in-place quantum subtractor.
#
# It is recommended to complete the [BasicGates kata](./../BasicGates/BasicGates.ipynb) before this one to get familiar with the basic gates used in quantum computing. The list of basic gates available in Q# can be found at [Microsoft.Quantum.Intrinsic](https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic). For the syntax of flow control statements in Q#, see [the Q# documentation](https://docs.microsoft.com/quantum/language/statements#control-flow).
#
# Each task is wrapped in one operation preceded by the description of the task.
# Your goal is to fill in the blank (marked with // ... comments)
# with some Q# code that solves the task. To verify your answer, run the cell using Ctrl/⌘+Enter.
#
# Within each section, tasks are given in approximate order of increasing difficulty; harder ones are marked with asterisks.
# To begin, first prepare this notebook for execution (if you skip the first step, you'll get "Syntax does not match any known patterns" error when you try to execute Q# code in the next cells; if you skip the second step, you'll get "Invalid kata name" error):
%package Microsoft.Quantum.Katas::0.9.1909.3002
# > The package versions in the output of the cell above should always match. If you are running the Notebooks locally and the versions do not match, please install the IQ# version that matches the version of the `Microsoft.Quantum.Katas` package.
# > <details>
# > <summary><u>How to install the right IQ# version</u></summary>
# > For example, if the version of `Microsoft.Quantum.Katas` package above is 0.1.2.3, the installation steps are as follows:
# >
# > 1. Stop the kernel.
# > 2. Uninstall the existing version of IQ#:
# > dotnet tool uninstall microsoft.quantum.iqsharp -g
# > 3. Install the matching version:
# > dotnet tool install microsoft.quantum.iqsharp -g --version 0.1.2.3
# > 4. Reinstall the kernel:
# > dotnet iqsharp install
# > 5. Restart the Notebook.
# > </details>
%workspace reload
# ## Part I. Simple Adder Outputting to Empty Qubits
#
#
# ### Theory
#
# * [Classical binary adder on Wikipedia](https://en.wikipedia.org/wiki/Adder_(electronics)).
# * Part 2 of the [paper on quantum binary addition](https://arxiv.org/pdf/quant-ph/0008033.pdf) by <NAME> explains how to adapt the classical adder to a quantum environment.
# ### Task 1.1. Summation of two bits
#
# **Inputs:**
#
# 1. qubit `a` in an arbitrary state $|\phi\rangle$,
# 2. qubit `b` in an arbitrary state $|\psi\rangle$,
# 3. qubit `sum` in state $|0\rangle$.
#
# **Goal:** Transform the `sum` qubit into the lowest bit of the binary sum of $\phi$ and $\psi$.
#
# * $|0\rangle + |0\rangle \to |0\rangle$
# * $|0\rangle + |1\rangle \to |1\rangle$
# * $|1\rangle + |0\rangle \to |1\rangle$
# * $|1\rangle + |1\rangle \to |0\rangle$
#
# Any superposition should map appropriately.
#
# **Example:** (Recall that $|+\rangle = \frac{1}{\sqrt{2}}(|0\rangle + |1\rangle)$, $|-\rangle = \frac{1}{\sqrt{2}}(|0\rangle - |1\rangle)$)
#
# $|+\rangle \otimes |-\rangle \otimes |0\rangle \to \frac{1}{2}(|000\rangle + |101\rangle - |011\rangle - |110\rangle)$
# +
%kata T11_LowestBitSum_Test
operation LowestBitSum (a : Qubit, b : Qubit, sum : Qubit) : Unit is Adj {
// ...
}
# -
# ### Task 1.2. Carry of two bits
#
# **Inputs:**
#
# 1. qubit `a` in an arbitrary state $|\phi\rangle$,
# 2. qubit `b` in an arbitrary state $|\psi\rangle$,
# 3. qubit `carry` in state $|0\rangle$.
#
# **Goal:** Set the `carry` qubit to $|1\rangle$ if the binary sum of $\phi$ and $\psi$ produces a carry.
#
# * $|0\rangle$ and $|0\rangle \to |0\rangle$
# * $|0\rangle$ and $|1\rangle \to |0\rangle$
# * $|1\rangle$ and $|0\rangle \to |0\rangle$
# * $|1\rangle$ and $|1\rangle \to |1\rangle$
#
# Any superposition should map appropriately.
#
# **Example:**
#
# $|+\rangle \otimes |-\rangle \otimes |0\rangle \to \frac{1}{2}(|000\rangle + |100\rangle - |010\rangle - |111\rangle)$
# +
%kata T12_LowestBitCarry_Test
operation LowestBitCarry (a : Qubit, b : Qubit, carry : Qubit) : Unit is Adj {
// ...
}
# -
# ### Task 1.3. One-bit adder
#
# **Inputs:**
#
# 1. qubit `a` in an arbitrary state $|\phi\rangle$,
# 2. qubit `b` in an arbitrary state $|\psi\rangle$,
# 3. two qubits `sum` and `carry` in state $|0\rangle$.
#
# **Goals:**
#
# * Transform the `sum` qubit into the lowest bit of the binary sum of $\phi$ and $\psi$.
# * Transform the `carry` qubit into the carry bit produced by that sum.
# +
%kata T13_OneBitAdder_Test
operation OneBitAdder (a : Qubit, b : Qubit, sum : Qubit, carry : Qubit) : Unit is Adj {
// ...
}
# -
# ### Task 1.4. Summation of 3 bits
#
# **Inputs:**
#
# 1. qubit `a` in an arbitrary state $|\phi\rangle$,
# 2. qubit `b` in an arbitrary state $|\psi\rangle$,
# 3. qubit `carryin` in an arbitrary state $|\omega\rangle$,
# 4. qubit `carryout` in state $|0\rangle$.
#
# **Goal:** Transform the `sum` qubit into the lowest bit of the binary sum of $\phi$, $\psi$ and $\omega$.
# +
%kata T14_HighBitSum_Test
operation HighBitSum (a : Qubit, b : Qubit, carryin : Qubit, sum : Qubit) : Unit is Adj {
// ...
}
# -
# ### Task 1.5. Carry of 3 bits
#
# **Inputs:**
#
# 1. qubit `a` in an arbitrary state $|\phi\rangle$,
# 2. qubit `b` in an arbitrary state $|\psi\rangle$,
# 3. qubit `carryin` in an arbitrary state $|\omega\rangle$,
# 4. qubit `carryout` in state $|0\rangle$.
#
# **Goal:** Transform the `carryout` qubit into the carry bit produced by the sum of $\phi$, $\psi$ and $\omega$.
# +
%kata T15_HighBitCarry_Test
operation HighBitCarry (a : Qubit, b : Qubit, carryin : Qubit, carryout : Qubit) : Unit is Adj {
// ...
}
# -
# ### Task 1.6. Two-bit adder
#
# **Inputs:**
#
# 1. two-qubit register `a` in an arbitrary state $|\phi\rangle$,
# 2. two-qubit register `b` in an arbitrary state $|\psi\rangle$,
# 3. two-qubit register `sum` in state $|00\rangle$,
# 4. qubit `carry` in state $|0\rangle$.
#
# **Goals:**
#
# * Transform the `sum` register into the binary sum (little-endian) of $\phi$ and $\psi$.
# * Transform the `carry` qubit into the carry bit produced by that sum.
#
# > All registers in this kata are in **little-endian** order.
# > This means that they have the least significant bit first, followed by the next least significant, and so on.
# >
# > In this exercise, for example, $1$ would be represented as $|10\rangle$, and $2$ as $|01\rangle$.
# >
# > The sum of $|10\rangle$ and $|11\rangle$ would be $|001\rangle$, with the last qubit being the carry qubit.
#
# <br/>
# <details>
# <summary>Need a hint? Click here</summary>
# Don't forget that you can allocate extra qubits.
# </details>
# +
%kata T16_TwoBitAdder_Test
operation TwoBitAdder (a : Qubit[], b : Qubit[], sum : Qubit[], carry : Qubit) : Unit is Adj {
// ...
}
# -
# ### Task 1.7. N-bit adder
#
# **Inputs:**
#
# 1. $N$-qubit register `a` in an arbitrary state $|\phi\rangle$,
# 2. $N$-qubit register `b` in an arbitrary state $|\psi\rangle$,
# 3. $N$-qubit register `sum` in state $|0...0\rangle$,
# 4. qubit `carry` in state $|0\rangle$.
#
# **Goals:**
#
# * Transform the `sum` register into the binary sum (little-engian) of $\phi$ and $\psi$.
# * Transform the `carry` qubit into the carry bit produced by that sum.
#
# **Challenge:**
#
# Can you do this without allocating extra qubits?
# +
%kata T17_ArbitraryAdder_Test
operation ArbitraryAdder (a : Qubit[], b : Qubit[], sum : Qubit[], carry : Qubit) : Unit is Adj {
// ...
}
# -
# ## Part II. Simple In-place Adder
#
# The adder from the previous section requires empty qubits to accept the result.
# This section adapts the previous adder to calculate the sum in-place,
# that is, to reuse one of the numerical inputs for storing the output.
# ### Task 2.1. In-place summation of two bits
#
# **Inputs:**
#
# 1. qubit `a` in an arbitrary state $|\phi\rangle$,
# 2. qubit `b` in an arbitrary state $|\psi\rangle$.
#
# **Goals:**
#
# * Transform qubit `b` into the lowest bit of the sum of $\phi$ and $\psi$.
# * Leave qubit `a` unchanged.
# +
%kata T21_LowestBitSumInPlace_Test
operation LowestBitSumInPlace (a : Qubit, b : Qubit) : Unit is Adj {
// ...
}
# -
# > Can we re-use one of the input bits to calculate the carry in-place as well? Why or why not?
# ### Task 2.2. In-place one-bit adder
#
# **Inputs:**
#
# 1. qubit `a` in an arbitrary state $|\phi\rangle$,
# 2. qubit `b` in an arbitrary state $|\psi\rangle$,
# 3. qubit `carry` in state $|0\rangle$.
#
# **Goals:**
#
# * Transform the `carry` qubit into the carry bit from the addition of $\phi$ and $\psi$.
# * Transform qubit `b` into the lowest bit of $\phi + \psi$.
# * Leave qubit `a` unchanged.
#
# <br/>
# <details>
# <summary>Need a hint? Click here</summary>
# Think very carefully about the order in which you apply the operations.
# </details>
# +
%kata T22_OneBitAdderInPlace_Test
operation OneBitAdderInPlace (a : Qubit, b : Qubit, carry : Qubit) : Unit is Adj {
// ...
}
# -
# ### Task 2.3. In-place summation of three bits
#
# **Inputs:**
#
# 1. qubit `a` in an arbitrary state $|\phi\rangle$,
# 2. qubit `b` in an arbitrary state $|\psi\rangle$,
# 3. qubit `carryin` in an arbitrary state $|\omega\rangle$.
#
# **Goals:**
#
# * Transform qubit `b` into the lowest bit from the sum $\phi + \psi + \omega$.
# * Leave qubits `a` and `carryin` unchanged.
# +
%kata T23_HighBitSumInPlace_Test
operation HighBitSumInPlace (a : Qubit, b : Qubit, carryin : Qubit) : Unit is Adj {
// ...
}
# -
# ### Task 2.4. In-place two-bit adder
#
# **Inputs:**
#
# 1. two-qubit register `a` in an arbitrary state $|\phi\rangle$,
# 2. two-qubit register `b` in an arbitrary state $|\psi\rangle$,
# 3. qubit `carry` in state $|0\rangle$.
#
# **Goals:**
#
# * Transform register `b` into the state $|\phi + \psi\rangle$.
# * Transform the `carry` qubit into the carry bit from the addition.
# * Leave register `a` unchanged.
# +
%kata T24_TwoBitAdderInPlace_Test
operation TwoBitAdderInPlace (a : Qubit[], b : Qubit[], carry : Qubit) : Unit is Adj {
// ...
}
# -
# ### Task 2.5. In-place N-bit adder
#
# **Inputs:**
#
# 1. $N$-qubit register `a` in an arbitrary state $|\phi\rangle$,
# 2. $N$-qubit register `b` in an arbitrary state $|\psi\rangle$,
# 3. qubit `carry` in state $|0\rangle$.
#
# **Goals:**
#
# * Transform register `b` into the state $|\phi + \psi\rangle$.
# * Transform the `carry` qubit into the carry bit from the addition.
# * Leave register `a` unchanged.
# +
%kata T25_ArbitraryAdderInPlace_Test
operation ArbitraryAdderInPlace (a : Qubit[], b : Qubit[], carry : Qubit) : Unit is Adj {
// ...
}
# -
# ## Part III*. Improved In-place Adder
#
# The in-place adder doesn't require quite as many qubits for the inputs and outputs, but it still requires an array of extra ("ancillary") qubits to perform the calculation.
#
# A relatively recent algorithm allows you to perform the same calculation using only one additional qubit.
#
# ### Theory
#
# * [Paper on improved ripple carry addition](https://arxiv.org/pdf/quant-ph/0410184.pdf) by <NAME>, <NAME>, <NAME>, and <NAME>.
# ### Task 3.1. Majority gate
#
# **Inputs:**
#
# 1. qubit `a` in an arbitrary state $|\phi\rangle$,
# 2. qubit `b` in an arbitrary state $|\psi\rangle$,
# 3. qubit `c` in an arbitrary state $|\omega\rangle$.
#
# **Goal:** Construct the "in-place majority" gate:
#
# * Transform qubit `a` into the carry bit from the sum $\phi + \psi + \omega$.
# * Transform qubit `b` into $|\phi + \psi\rangle$.
# * Transform qubit `c` into $|\phi + \omega\rangle$.
# +
%kata T31_Majority_Test
operation Majority (a : Qubit, b : Qubit, c : Qubit) : Unit is Adj {
// ...
}
# -
# ### Task 3.2. UnMajority and Add gate
#
# **Inputs:**
#
# 1. qubit `a` storing the carry bit from the sum $\phi + \psi + \omega$,
# 2. qubit `b` in state $|\phi + \psi\rangle$,
# 3. qubit `c` in state $|\phi + \omega\rangle$.
#
# **Goal:** Construct the "un-majority and add", or "UMA" gate:
#
# * Restore qubit `a` into state $|\phi\rangle$.
# * Transform qubit `b` into state $|\phi + \psi + \omega\rangle$.
# * Restore qubit `c` into state $|\omega\rangle$.
# +
%kata T32_UnMajorityAdd_Test
operation UnMajorityAdd (a : Qubit, b : Qubit, c : Qubit) : Unit is Adj {
// ...
}
# -
# ### Task 3.3. One-bit Majority-UMA adder
#
# **Inputs:**
#
# 1. qubit `a` in an arbitrary state $|\phi\rangle$,
# 2. qubit `b` in an arbitrary state $|\psi\rangle$,
# 3. qubit `carry` in state $|0\rangle$.
#
# **Goal:** Construct a one-bit binary adder from task 2.2 using Majority and UMA gates.
#
# <br/>
# <details>
# <summary>Need a hint? Click here</summary>
# Allocate an extra qubit to hold the carry bit used in Majority and UMA gates during the computation. It's less efficient here, but it will be helpful for the next tasks.
# </details>
# +
%kata T33_OneBitMajUmaAdder_Test
operation OneBitMajUmaAdder (a : Qubit, b : Qubit, carry : Qubit) : Unit is Adj {
// ...
}
# -
# ### Task 3.4. Two-bit Majority-UMA adder
#
# **Inputs:**
#
# 1. two-qubit register `a` in an arbitrary state $|\phi\rangle$,
# 2. two-qubit register `b` in an arbitrary state $|\psi\rangle$,
# 3. qubit `carry` in state $|0\rangle$.
#
# **Goal:** Construct a two-bit binary adder from task 2.4 using Majority and UMA gates.
#
# <br/>
# <details>
# <summary>Need a hint? Click here</summary>
# Think carefully about which qubits you need to pass to the two gates.
# </details>
# +
%kata T34_TwoBitMajUmaAdder_Test
operation TwoBitMajUmaAdder (a : Qubit[], b : Qubit[], carry : Qubit) : Unit is Adj {
// ...
}
# -
# ### Task 3.5. N-bit Majority-UMA adder
#
# **Inputs:**
#
# 1. $N$-qubit register `a` in an arbitrary state $|\phi\rangle$,
# 2. $N$-qubit register `b` in an arbitrary state $|\psi\rangle$,
# 3. qubit `carry` in state $|0\rangle$.
#
# **Goal:** Construct an N-bit binary adder from task 2.5 using only one ancillary qubit.
# +
%kata T35_ArbitraryMajUmaAdder_Test
operation ArbitraryMajUmaAdder (a : Qubit[], b : Qubit[], carry : Qubit) : Unit is Adj {
// ...
}
# -
# ## Part IV*. In-place Subtractor
#
# Subtracting a number is the same operation as adding a negative number.
# As such, it's pretty easy to adapt the adder we just built to act as a subtractor.
# ### Task 4.1. N-bit Subtractor
#
# **Inputs:**
#
# 1. $N$-qubit register `a` in an arbitrary state $|\phi\rangle$,
# 2. $N$-qubit register `b` in an arbitrary state $|\psi\rangle$,
# 3. qubit `borrow` in state $|0\rangle$.
#
# **Goal:** Construct an N-bit binary subtractor.
#
# * Transform register `b` into the state $|\psi - \phi\rangle$.
# * Set qubit `borrow` to $|1\rangle$ if that subtraction requires a borrow.
# * Leave register `a` unchanged.
#
# <br/>
# <details>
# <summary>Need a hint? Click here</summary>
# Use the adder you already built. Experiment with inverting registers before and after the addition.
# </details>
# +
%kata T41_Subtractor_Test
operation Subtractor (a : Qubit[], b : Qubit[], borrow : Qubit) : Unit is Adj {
// ...
}
|
RippleCarryAdder/RippleCarryAdder.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This code works till numpy version 1.19.5
# Please look for a solution if you want it to work wwith numpy version 1.20
import pandas as pd
import tensorflow as tf
df = pd.read_csv('fake-news/train.csv')
df.head()
#check if the gpu is accessible here or not
tf.test.is_gpu_available(cuda_only=True)
# checking for nan values
df.isnull().sum()
# dropping the nan values
df.dropna()
#Storing the dependent variables
X = df.drop(["label"],1)
# Storing the independent variable
y= df["label"]
y.value_counts()
X.shape, y.shape
from tensorflow.keras.layers import Embedding
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.models import Sequential
from tensorflow.keras.preprocessing.text import one_hot
from tensorflow.keras.layers import LSTM
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import Dropout
# Now we set the vocabulary size
voc_size = 5000
# # OneHot Representation
messages=X.copy()
messages['title'][1]
# We are resetting indexes here because of the dropped nan values above
messages.reset_index(inplace=True)
import nltk
# re = regular expression
import re
# stopwords is used so as to remove the not so important words
from nltk.corpus import stopwords
nltk.download('stopwords')
# # Dataset Preprocessing and cleaning
# +
from nltk.stem.porter import PorterStemmer
ps = PorterStemmer()
corpus = []
for i in range(0, len(messages)):
print(i)
# substituting everything with a blank space
review = re.sub('[^a-zA-Z]', ' ', str(messages['title'][i]))
review = review.lower()
review = review.split()
review = [ps.stem(word) for word in review if not word in stopwords.words('english')]
review = ' '.join(review)
corpus.append(review)
# -
corpus
# Now we assign the words in the corpus some index
onehot_repr=[one_hot(words,voc_size)for words in corpus]
onehot_repr
# # Embedding Representation
sent_length=20
embedded_docs=pad_sequences(onehot_repr,padding='pre',maxlen=sent_length)
print(embedded_docs)
embedded_docs[0]
embedded_docs
# # Model Creation
## Now we start creating the model for it
embedding_vector_features=40
model=Sequential()
model.add(Embedding(voc_size,embedding_vector_features,input_length=sent_length))
# model.add(Dropout(0.3))
model.add(LSTM(100))
# model.add(Dropout(0.3))
model.add(Dense(1,activation='sigmoid'))
model.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
print(model.summary())
len(embedded_docs),y.shape
import numpy as np
X_final=np.array(embedded_docs)
y_final=np.array(y)
X_final.shape,y_final.shape
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X_final, y_final, test_size=0.33, random_state=42)
# # Model Training
### Finally Training
model.fit(X_train,y_train,validation_data=(X_test,y_test),epochs=10,batch_size=64)
# # Adding Dropout
from tensorflow.keras.layers import Dropout
# setting seed to not get different values every time
import tensorflow
tensorflow.random.set_seed(42)
## Creating model
embedding_vector_features=40
model=Sequential()
model.add(Embedding(voc_size,embedding_vector_features,input_length=sent_length))
model.add(Dropout(0.3))
model.add(LSTM(100))
model.add(Dropout(0.3))
model.add(Dense(1,activation='sigmoid'))
model.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
# # Performance Metrics And Accuracy
y_pred=model.predict_classes(X_test)
from sklearn.metrics import confusion_matrix
confusion_matrix(y_test,y_pred)
from sklearn.metrics import accuracy_score
accuracy_score(y_test,y_pred)
import numpy
numpy.__version__
|
Fake News Classifier using LSTM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# ## Train a model with bike rental data using XGBoost algorithm
# ### Model is trained with XGBoost installed in notebook instance
# ### In the later examples, we will train using SageMaker's XGBoost algorithm
# Install xgboost in notebook instance.
#### Command to install xgboost
# !pip install xgboost==1.2
# +
import sys
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.metrics import mean_squared_error, mean_absolute_error
# XGBoost
import xgboost as xgb
# -
column_list_file = 'bike_train_column_list.txt'
train_file = 'bike_train.csv'
validation_file = 'bike_validation.csv'
test_file = 'bike_test.csv'
columns = ''
with open(column_list_file,'r') as f:
columns = f.read().split(',')
columns
# Specify the column names as the file does not have column header
df_train = pd.read_csv(train_file,names=columns)
df_validation = pd.read_csv(validation_file,names=columns)
df_train.head()
df_validation.head()
# +
X_train = df_train.iloc[:,1:] # Features: 1st column onwards
y_train = df_train.iloc[:,0].ravel() # Target: 0th column
X_validation = df_validation.iloc[:,1:]
y_validation = df_validation.iloc[:,0].ravel()
# -
# XGBoost Training Parameter Reference:
# https://github.com/dmlc/xgboost/blob/master/doc/parameter.md
#regressor = xgb.XGBRegressor(max_depth=5,eta=0.1,subsample=0.7,num_round=150)
regressor = xgb.XGBRegressor(max_depth=5,n_estimators=150)
regressor
regressor.fit(X_train,y_train, eval_set = [(X_train, y_train), (X_validation, y_validation)])
eval_result = regressor.evals_result()
training_rounds = range(len(eval_result['validation_0']['rmse']))
print(training_rounds)
plt.scatter(x=training_rounds,y=eval_result['validation_0']['rmse'],label='Training Error')
plt.scatter(x=training_rounds,y=eval_result['validation_1']['rmse'],label='Validation Error')
plt.grid(True)
plt.xlabel('Iteration')
plt.ylabel('RMSE')
plt.title('Training Vs Validation Error')
plt.legend()
plt.show()
xgb.plot_importance(regressor)
plt.show()
# Verify Quality using Validation dataset
# Compare actual vs predicted performance with dataset not seen by the model before
df = pd.read_csv(validation_file,names=columns)
df.head()
df.shape
X_test = df.iloc[:,1:]
print(X_test[:5])
result = regressor.predict(X_test)
result[:5]
df['count_predicted'] = result
df.head()
# Negative Values are predicted
df['count_predicted'].describe()
df[df['count_predicted'] < 0]
df['count_predicted'].hist()
plt.title('Predicted Count Histogram')
plt.show()
def adjust_count(x):
if x < 0:
return 0
else:
return x
df['count_predicted'] = df['count_predicted'].map(adjust_count)
df[df['count_predicted'] < 0]
# Actual Vs Predicted
plt.plot(df['count'], label='Actual')
plt.plot(df['count_predicted'],label='Predicted')
plt.xlabel('Sample')
plt.ylabel('Count')
plt.xlim([100,150])
plt.title('Validation Dataset - Predicted Vs. Actual')
plt.legend()
plt.show()
# +
# Over prediction and Under Prediction needs to be balanced
# Training Data Residuals
residuals = (df['count'] - df['count_predicted'])
plt.hist(residuals)
plt.grid(True)
plt.xlabel('Actual - Predicted')
plt.ylabel('Count')
plt.title('Residuals Distribution')
plt.axvline(color='r')
plt.show()
# -
value_counts = (residuals > 0).value_counts(sort=False)
print(' Under Estimation: {0:0.2f}'.format(value_counts[True]/len(residuals)))
print(' Over Estimation: {0:0.2f}'.format(value_counts[False]/len(residuals)))
print("RMSE: {0:0.2f}".format(mean_squared_error(df['count'],df['count_predicted'])**.5))
# +
# RMSlE - Root Mean Squared Log Error
# RMSLE Metric is used by Kaggle for this competition
# RMSE Cost Function - Magnitude of difference matters
# RMSLE cost function - "Only Percentage difference matters"
# Reference:<NAME>, <NAME>
# https://www.slideshare.net/KhorSoonHin/rmsle-cost-function
def compute_rmsle(y_true, y_pred):
if type(y_true) != np.ndarray:
y_true = np.array(y_true)
if type(y_pred) != np.ndarray:
y_pred = np.array(y_pred)
return(np.average((np.log1p(y_pred) - np.log1p(y_true))**2)**.5)
# -
print('RMSLE')
print(compute_rmsle(100,50),
compute_rmsle(1000,500),
compute_rmsle(10000,5000))
print('RMSLE')
print(compute_rmsle(100,25),
compute_rmsle(1000,250),
compute_rmsle(10000,2500))
print('RMSE')
print(mean_squared_error([100],[50])**.5,
mean_squared_error([1000],[500])**.5,
mean_squared_error([10000],[5000])**.5)
print('RMSE')
print(mean_squared_error([100],[25])**.5,
mean_squared_error([1000],[250])**.5,
mean_squared_error([10000],[2500])**.5)
print("RMSLE: {0}".format(compute_rmsle(df['count'],df['count_predicted'])))
# Prepare Data for Submission to Kaggle
df_test = pd.read_csv(test_file,parse_dates=['datetime'])
df_test.head()
X_test = df_test.iloc[:,1:] # Exclude datetime for prediction
X_test.head()
result = regressor.predict(X_test)
result[:5]
df_test["count"] = result
df_test.head()
df_test[df_test["count"] < 0]
df_test["count"] = df_test["count"].map(adjust_count)
df_test[['datetime','count']].to_csv('predicted_count.csv',index=False)
# +
# RMSLE (Kaggle) Score
# Test 1: 0.62
# -
|
xgboost/BikeSharingRegression/bikerental_xgboost_localmode_rev1.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.5
# language: julia
# name: julia-1.0
# ---
# # 1D Spin Chain Part 2
# ### <NAME>
#
# ### Category: Grad
#
# ### 1D Spin Chain Series
# * [Jacobi Transformation](../Numerics_Prog/Jacobi-Transformation.ipynb)
# * [1D Spin Chain Prerequisites](../Graduate/1D-Spin-Chain-Prerequisites.ipynb)
# * [1D Spin Chain Pt. 2](../Graduate/1D-Spin-Chain-2.ipynb)
#
# Check out part 1 for the theoretical background. Today is all programming.
# Remember to keep a reasonable number of spins, $n$. The number of states goes as $2^n$, and the size of the Hamiltonian will go as $2^n \times 2^n$. A 10-spin chain will have over a million entries, before taking into account any memory reducing tricks.
#
# Here's the details of the largest spin-chains that fit on my machine which has 16GB of RAM and a 3.2 GHz Intel i5 processor:
#
# | n | m_z | t (min) | Memory (Gb) |
# |---|-----|---------|-------------|
# |16 | 8 | 1.5 | 2 |
# |18 | 9 | ? | >16 |
# |18 | 8 | 32.1 | 15.5 |
#
# I have included a file in this directory, ED.jl, that is just the necessary executable parts of this Jupyter notebook. For large $n$, I recommend running ED.jl.
using LinearAlgebra
# Here, we input one parameter `n`, the number of spins in our chain.
#
# The program automatically calculates the parameter `nstates`.
n=4
nstates=2^n
# Now, let's write out all of our possible states in the $S^z$ basis.
psi=convert.(Int8, collect(0:(nstates-1)) )
for p in psi
println(bitstring(p)[end-n:end],' ',p)
end
# As in Part 1, we will be using the powers of 2 to compute magnetization, and masks to flip spins. To not have to calculate them each time, we just store them in memory.
# +
powers2=collect(0:(n-1));
powers2=convert.(Int8, 2.0 .^powers2 );
mask=[0;powers2]+[powers2;0];
mask=convert.(Int8, [mask[2:end-1];[1+2^(n-1)]])
for m in mask
println(bitstring(m)[end-n:end])
end
# -
# In Part 1, I used the number of up-spins as a proxy for magnetization. Here, we need the <i>actual</i> magnetization, not a proxy. An up-spin is $+1/2$ and a down-spin is $-1/2$. We modify our magnetization by
#
# \begin{equation}
# m=\frac{1}{2}\left(n_{\uparrow}-n_{\downarrow} \right)
# = \frac{1}{2} \left( n_{\uparrow}- \left( n - n_{\uparrow} \right) \right),
# \end{equation}
#
# \begin{equation}
# m = n_{\uparrow} - \frac{n}{2}.
# \end{equation}
m=zeros(Int8,length(psi))
for i in 1:nstates
m[i]=sum((psi[i].&powers2)./(powers2))
end
m=m.-n/2
# ## Grouped by Magnetization
#
# Now that we have the magnetizations corresponding to each state, we perform some trickery that allows us to reduce the difficulty of our problem dramatically.
#
# Magnetization is a conserved quantity. By Noether's theorem, we know that the Hamiltonian is not going to mix states of different magnetizations. We only deal with one magnetization at a time, which is a much smaller problem.
# The possible values for magnetization
ma=collect(0:1:n).-n/2
# Now let's just pick out a single magnetization quantum number $mz$ and only work with that matrix for the rest of the post.
# +
# The magnetic quantum number
mz=3
# An array of states with the correct magnetization
psi_mz=psi[m.==ma[mz]]
[psi_mz bitstring.(psi_mz) m[psi_mz.+1]]
# -
# And now creating the matrix.
# +
dim=length(psi_mz)
M=ma[mz]*(ma[mz]+1)*Matrix{Float64}(I,dim,dim)
#M=zeros(Float64,dim,dim); use this for XY model
# -
# Though we have significantly reduced the size of the matrix by restricting to one magnetization, we no longer have our states in `1,2,3,4...` order. Their position in an array no longer corresponds to their value. Therefore, we need a function to determine their index once we know their value.
#
# We can find the index of the flipped state multiple different ways, but the simplest is by the Midpoint method. We split the interval in half, and see if the value we are looking for is higher or lower than the middle point. Then we get a new interval.
function findstate(state,set)
#Lower bound of interval
imin=1
#Upper bound of interval
imax=length(set)
# checking if the lower bound is what we are looking for
if set[imin] == state
return imin
end
# checking if the upper bound is what we are looking for
if set[imax] == state
return imax
end
# Initializing variables
# looking to see if we've found it yet
found=false
# how many times we've gone around the while loop
count=0
while found==false && count < length(set)
count+=1
tester=floor(Int,imin+(imax-imin)/2)
if state < set[tester]
imax=tester-1
elseif state > set[tester]
imin=tester+1
else
found=true
return tester
end
end
if found == false
println("findstate never found a match")
println("Are you sure the state is in that Array?")
end
return 0
end
# Now time to generate the matrix.
#
# For each state and for each pair of adjacent spins within that state, we apply the operator that flips adjacent spins, `$mask`. Sometimes the adjacent spins will take on the same value, `00` or `11`. In this circumstance, the off-diagonal part of the Hamiltonian would not act on those spins. The state generated by the operator would have a different magnetization, and we can neglect that pair.
#
# If the new state produced by this process has the same magnetization, we know that the flip exists in the Hamiltonian, and add the entry to the matrix.
#
# In this algorithm, we do end up going over each pair twice, but I have yet to figure out how to take advantage of the degeneracy to cut the calculation in half. Let me know if you have a better way to write this.
# +
mp=sum(psi_mz[1].&powers2./powers2)
for ii in 1:length(psi_mz)
p=psi_mz[ii]
for jj in 1:n
flipped=p⊻mask[jj]
if sum((flipped.&powers2)./powers2) == mp
tester=findstate(flipped,psi_mz)
M[ii,tester]=.5
M[tester,ii]=.5
println(bitstring(p)[end-n:end],'\t',bitstring(flipped)[end-n:end])
end
end
end
# -
M
F=eigen(M)
display(F.values)
display(F.vectors)
# Now we have eigenvalues and eigenvectors! You just solved the Heisenburg Spin Chain!
#
# In my next post, I will analyze what this tells us about the system and what we can do with the information.
# @inproceedings{sandvik2010computational,
# title={Computational studies of quantum spin systems},
# author={Sandvik, <NAME>},
# booktitle={AIP Conference Proceedings},
# volume={1297},
# number={1},
# pages={135--338},
# year={2010},
# organization={AIP}
# }
|
Graduate/1D-Spin-Chain-2.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# name: ir
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/daeun-j/leaf/blob/master/FA.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="xjPcIeI3LPsj"
library(tidyverse)
library(psych)
library(lavaan)
library(semPlot)
# + id="MIoy3rJQLaWh"
# load data
data <- read.csv(file = "aging SCT only(-2SD)_200612_v1.csv", header = TRUE)
data %>% str()
# + [markdown] id="5Jn5UpTdMYPN"
# One initial step with any factor analysis is to check the factorability of the data.
# + id="skc6fxRrMa_3"
KMO(select(data,SCT_A2_C:SCT_P_NC))
# + id="FdWaIxxeNBti"
cortest.bartlett(select(data,SCT_A2_C:SCT_P_NC), n = 233)
# + [markdown] id="btOZ3ZhdNMEr"
# Proceed with EFA.
# # EFA
# With fa() function from psych, we determine the number of likely factors in the data.
#
# + id="bkxQ8lFMNVdH"
efa <- fa(select(data,SCT_A2_C:SCT_P_NC), nfactors = 3,
rotate = "none", fm = "ml")
efa
# + [markdown] id="aJUb8MfRO52e"
# Two approaches to determining number of factors are a screeplot and parallel analysis. First, the screeplot of eigenvalues:
#
# + id="-EGo-5EnO7HT"
plot(efa$e.values)
# + [markdown] id="pRWVSOowPDSr"
# It seems at least 2 - 3 factors.
# + id="VXcTOsBdPM_-"
fa.parallel(select(data,SCT_A2_C:SCT_P_NC), fa = "fa", fm = "ml")
# + [markdown] id="4B6_VFxCPbi0"
# This indicates that we have 2 factors which are above the level of random noise. 2 seems like a good number to go with.
#
# Now we fit a 2-factor solution and rotate it - we’ll use an oblique rotation that allows the factors to correlate.
# + id="zl8Z7Iq2PgAR"
efa_with2 <- fa(select(data,SCT_A2_C:SCT_P_NC), nfactors = 2, rotate = "oblimin", fm = "ml")
efa_with2
# + [markdown] id="xXckcpqkRGzy"
# #CFA
# Now we’ll use the same data to run a CFA. We’ll be aiming for a 2 factor solution based on the theory behind the various measures in this dataset.
#
# + [markdown] id="V1GuHcwzRn4t"
# Step 1: Specify the Model
# + id="d2d-uLGERX5i"
modelOne <- "y =~ SCT_A2_C+SCT_A3_NC+SCT_P_C"
# + [markdown] id="2N7yAJVWRqN2"
# Step 2: Estimate (fit) the model
# + id="tstJTA-GRttQ"
modelOne.fit <- cfa(model = modelOne, data = data)
# + [markdown] id="tcME2dunZ3-y"
# Step 3: Model Fit and Step 4: Interpret
# + id="gXug6HeTZ4t3"
lavaan::summary(modelOne.fit, estimates = T, standardized = T, fit.measures = T)
|
FA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from twitter_credentials import credentials
import tweepy
auth = tweepy.OAuthHandler(credentials["TWITTER_CONSUMER_KEY"],
credentials["TWITTER_CONSUMER_KEY_SECRET"])
auth.set_access_token(credentials["TWITTER_ACCESS_TOKEN"],
credentials["TWITTER_ACCESS_TOKEN_SECRET"])
api=tweepy.API(auth)
# +
def processTweet(tweet):
tweet_dict = {
'datetime': tweet.created_at,
'tweet': tweet.text,
'entities': tweet.entities
}
if tweet.coordinates:
tweet_dict['coordinates'] = tweet.coordinates
if tweet.geo:
tweet_dict['geo'] = tweet.geo
return tweet_dict
# -
cursor = tweepy.Cursor(api.search, q='vote', since_id='1059596220564422659', max_id='1059958608442535936').items(2000)
# get a list of dictionaries
tweets = [ processTweet(tweet) for tweet in cursor ]
print(tweets)
# +
import json
from pymongo import MongoClient
# This connects us to the "legistlation" database, and the "collection" [think table] news
# in that database
# These don't already exist, we're making them as we're connecting to them
# If we wanted to connect to Mongo on AWS, this is where we'd set that info
client = MongoClient()
db = client.legistlation
news_collection = db.news
|
project_4/.ipynb_checkpoints/Project_4_code_2-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # OpenStreetMap Data Case Study
#
# ## Map Area and Source
#
# The map area covers Singapore, SG. I live here.
#
# Map Source: [Mapzen Metro Extract](https://mapzen.com/data/metro-extracts/metro/singapore/)
#
# ## Problems Encountered In the Map
# A first-pass ETL of `singapore.osm` into a SQLite3 database via `data.py`, several problems were identified and addressed in order:
# - *Lorong*, the local equivalent of *street* or *avenue* has been expressed inconsistently - some abbreviated (i.e. `Lor`), some in full (i.e. '`Lorong`')
# - *Lorong*s not in consistent order (e.g. some `Toa Payoh Lorong 1`, some `Lorong 1 Geylang`)
# - Invalid postcodes (e.g. `#B1-42`, `Johor Bahru`, `<different>`)
# - Improperly formed postcodes (e.g. `S 278989`, `Singapore 408564`)
# - Non-local 5-digit postcodes (e.g. `18953`)
#
# ### Detecting and Fixing Lorongs
#
# The preferred way of referring *Lorong*-style streets is having *Lorong* first, as opposed to regular street names in English (e.g Pickering Street). A number or name follows *Lorong*:
# - Lorong 1 Geylang
# - Lorong Chuan
# - Lorong Ah Soo
#
# Inconsistent abbreviations were found via a regex audit:
# ```python
# def audit_lorong_type(lorong_types, street_name):
# """ Process addr:street tag
# Args:
# lorong_types - Dictionary of keys that are Lorong-street types
# street_name - Street name; 'v' attribute of addr:street
# """
# lorong_type_re = re.compile(r'(\blor\b)|(\blorong\b)', re.IGNORECASE)
#
# match_result = lorong_type_re.search(street_name)
# if match_result:
# lorong_type = match_result.group()
# lorong_types[lorong_type].add(street_name)
# ```
# and fixed via a simple search/replace function, shown together with the subsequent fix.
#
# __*Lorongs* First!__
#
# If, for example '*Lorong* 1 Geylang' is inverted, i.e 'Geylang *Lorong* 1' - this is still perfectly understandable, however, examples like 'Chuan *Lorong*' is unheard of in local vernacular. Hence, having *Lorong* in front is consistent.
#
# One caveat: if a *Lorong*-style street name begins with a digit, e.g. '2 Lorong Napiri', it would be better practice to ignore it, as it does not refer to a street, so swapping 'Lorong Napiri 2' will not make sense. The code below takes care of the street name cleanup.
# ```python
# def clean_lorong(sname):
# """ Cleans lorong-style street names
# Args:
# sname - Street name; 'v' attribute of addr:street
# Returns:
# Cleaned street name
# """
# verbose = True
#
# # first find and replace Lor with Lorong
# result = re.sub(r'\bLor\b', 'Lorong', sname)
# # if sname != result: print("original: {}, cleaned: {}".format(sname, result))
#
# # then swap order of Lorong if applicable
# idx_l = result.find('Lorong')
# # if street name does not begin with lorong & a digit, swap lorong
# if (idx_l > 0) and (result[0] not in set('123456890')):
# newname = result[idx_l:]+ " " + result[:idx_l]
# if verbose: print("swap: {} -> {}".format(result, newname))
# return newname
# else:
# return result
# ```
#
# ### Detecting and Fixing Postcodes
# Singapore postal codes are strictly 6 digits in length, with the first 2 digits denoting the postal sector. A simple regular expression can be used to check for non-compliance:
# ```python
# def audit_postcode_type(postcode_types, postcode):
# """ Process addr:postcode tag
# Args:
# postcode_types - Dictionary of keys not compliant to Singapore postcode format
# postcode - Postcode; 'v' attribute of addr:postcode
# """
# postcode_type_re = re.compile(r'^\d{6}$')
#
# match_result = postcode_type_re.search(postcode)
# if not match_result:
# postcode_types[postcode].add(postcode)
# ```
# The non-compliant postal codes fell into 3 categories - either invalid, malformed or non-Singapore codes. The Mapzen metro extract boundaries extend past Singapore's borders, into both neighbouring Malaysia as well as Indonesia. This accounts for the number of 5-digit postal codes in the data set, which are simply ignored in this analysis.
#
# Malformed codes can be corrected by stripping all other characters and returning the 6-digit postal code.
#
# To resolve the other invalid codes it was possible to either update the postal code to a valid code that does not exist<sup>[1]</sup> (i.e. `000000`), or simply drop the tag. The first option of updating the postal code was chosen, to preserve the tag counts/statistics for later analysis.
# ```python
# def clean_postcode(postcode):
# """ Cleans postcode
# Args:
# postcode - Postcode; 'v' attribute of addr:postcode
# Returns:
# Cleaned postcode
# """
# verbose = True
# pattern = re.compile(r'^\d{5,6}$') # valid codes are 5 or 6-digits
#
# match_result = pattern.search(postcode)
# if not match_result: # process non-compliant codes
# result = re.sub(r'[^\d]', '', postcode) # clean up malformed codes
# else: # do not process compliant codes
# return postcode
#
# if len(result) != 6: # if not compliant after clean up, set default postcode
# result = "000000"
#
# if verbose: print("original: {}, cleaned: {}".format(postcode, result))
# return result
# ```
#
# ## Overview of the Data & Additional Queries
# An overview of the data sources is provided in this section. The source OSM XML file was downloaded on **26 Aug 2017** from a standard Mapzen Metro Extract.
# ### Map Data Quality
# The overall data quality with regards to street names is remarkably consistent and clean. This could be due to the fact that Singapore has strict guidelines on how streets are named<sup>[2]</sup>.
# ### Database and Source Files
#
# File | Size | Type
# --- | --- | ---
# singapore.osm | 329M | OpenStreetMap XML
# singapore.sqlite | 230M | SQLite3 DB
# nodes.csv | 119M | Nodes CSV File
# nodes_tags.csv | 4.9M | Node Tags CSV File
# ways.csv | 14M | Ways CSV File
# ways_nodes.csv | 44M | Way Nodes CSV File
# ways_tags.csv | 22M | Way Node Tags CSV File
#
# ### Nodes & Ways
# The number of nodes extracted from `singapore.osm` are as follows:
# ```sql
# sqlite> select count(*) as n from nodes;
# n
# --------------------
# 1513629
# ```
# The number of ways extracted from `singapore.osm` are as follows:
# ```sql
# sqlite> select count(*) as n from ways;
# n
# --------------------
# 237163
# ```
# ### Number of Unique Users
# ```sql
# sqlite> select count(*) from
# (select u.user from
# (select user from nodes union all select user from ways) as u
# group by u.user) as c;
# count(*)
# -------------------------
# 2144
#
# ```
# ### Changesets
# A changeset can be open for up to 24 hours, and have maximum of 10,000 entries. The 10-largest changesets are:
# ```sql
# sqlite> select u.changeset, count(*) as n from
# ...> (select changeset from nodes union all select changeset from ways) as u
# ...> group by u.changeset order by n desc limit 10;
# changeset n
# -------------------- ----------
# 35657677 9875
# 29647823 9870
# 25268928 9009
# 40738823 8938
# 15807798 8717
# 19508643 8682
# 27022028 7786
# 24098829 7544
# 19934418 7039
# 40236725 5949
# ```
# The size of changesets probably reveals that the users of these changesets are bots.
# ### Which users made the 10-largest changesets?
# ```sql
# sqlite> select j.user from
# (select user, changeset from nodes
# union all
# select user, changeset from ways) as j
# join
# (select u.changeset, count(*) as n from
# (select changeset from nodes
# union all
# select changeset from ways) as u
# group by u.changeset order by n desc limit 10) as k
# on j.changeset = k.changeset
# group by j.user;
# user
# --------------------
# JaLooNz
# mdk
# ```
# The top 10 largest changesets were made by only 2 users! This lends more evidence that the these users contributing to the largest changesets are bots. Another check for largest contributors corroborates this as well:
# ```sql
# sqlite> select u.user, count(*) as n from
# (select user from nodes union all select user from ways) as u
# group by u.user order by n desc limit 10;
# user n
# -------------------- ----------
# JaLooNz 405158
# berjaya 117460
# rene78 77593
# cboothroyd 72280
# lmum 50780
# kingrollo 39068
# Luis36995 38823
# ridixcr 38240
# <NAME> 37160
# calfarome 32946
# ```
#
# ## Suggestions for Improvement
# OSM provides GPS longitude and latitude data - but does it also record altitude data? OSM documentation<sup>[3]</sup> on this shows that it could potentially be stored in `ele=` tags. A search on the full Singapore dataset reveals that there are very few such tags compared to the circa 1.5 million nodes in the dataset:
# ```sql
# sqlite> select count(*) from (select * from nodes_tags union all select * from ways_tags) as u where key = 'ele';
# 54
# ```
# The OSM documentation also reveals the difficulty in recording altitude data:
# - Altitude information from consumer-grade GPS devices is often not accurate enough
# - Lack of definitive reference altitudes in Singapore (and around the world) for other readings to compare accurately from
# - Pressure/barometric-based altimeters are unstable (dependent on temperature and other factors) requiring frequent calibration
# - Many reference systems/elevation models of altitude measurement (e.g. Nullebene, Geoid, Referenzellipsoid) requires a significant amount of skill and knowledge from the contributor
# - Measuring depth is yet another complication
#
# A possible solution to this, is to involve big data. While GPS readings from a single contributor may not offer the desired accuracy, GPS tracks from multiple contributors could be taken instead. Building 3-dimensional location heatmaps of a location to achieve decent accuracy. This new data would need to be processed with big data methods because:
# - High data volume (fine-grained crowd-sourced data)
# - High data veracity (expect noise, bias in the contributor data)
# - Moderate data variety (Elevation data could be in the form of GPS, barometric readings, different systems, different formats; raw data could even come from pictures/videos with GPS tags, etc)
#
# Further down the data pipeline, more advanced statistical or machine-learning methods would need to be applied to clean and analyse the data.
#
# As availability and accuracy of the resulting altitude data is dependent on contributor data, the limitations on this approach would be obvious - areas with fewer contributors will have less reliable altitude information. In the future, devices or methods to obtain accurate elevation data may be available and reliance on contributer data would be reduced.
#
# However, I still think it is a good starting point sufficient for many uses - sufficiently accurate elevation data for hiking, biking, driving.
#
# ## Additional Ideas
#
# ### Most Common Speed Limit on Asphalt Roads
# The official speed limit in Singapore is 50 kilometers per hour on most roads. The query returns 60 instead. Analysing the reasons may require understanding the nature of the roads (e.g. geographical features, location) or source of `maxspeed`.
# ```sql
# sqlite> select key, value, count(*) as n from ways_tags where id in
# (select id from ways_tags where value = 'asphalt') and key = 'maxspeed'
# group by key, value order by n desc limit 5;
# key value n
# ------------------------- ---------- ----------
# maxspeed 60 3136
# maxspeed 50 1875
# maxspeed 70 1336
# maxspeed 90 668
# maxspeed 40 224
# ```
# ### Common Cafes
# The most common cafe - Starbucks, appears in multiple rows below, due to the different naming conventions of the company. Cleaning data from any `name=` fields will be daunting, as there is considerable variety of correct/valid names with little constraints. Improving the query may require tools that SQL may find challenging.
# ```sql
# sqlite> select value, count(*) as n from nodes_tags where id in
# (select id from nodes_tags where value = 'cafe') and key = 'name'
# group by value order by n desc limit 10;
# value n
# ------------------------- ----------
# Starbucks 58
# The Coffee Bean & Tea Lea 13
# Toast Box 8
# Starbucks Coffee 6
# Coffee Bean & Tea Leaf 4
# Coffee Shop 4
# The Coffee Bean and Tea L 4
# Killiney Kopitiam 3
# Koi Cafe 3
# Lola's Cafe 3
# ```
# ### Unicode Data
# Some unicode tags in Chinese language were discovered, which mostly consisted of road names. This raises an interesting point - these unicode data will have to be handled separately in the data audit and cleaning process, because Singapore is a multi-cultural/multi-language society. Inclusions of other language data will increase in future.
# ```sql
# sqlite> select u.value, count(*) as n from
# (select key, value from nodes_tags union all
# select key, value from ways_tags) as u
# where key = 'zh' group by u.value order by n desc limit 10;
# value n
# ------------------------- ----------
# 南北大道 64
# 第二通道高速公路 43
# 新柔长堤 35
# 麦士威路 15
# 克罗士街上段 13
# 尼路 11
# 奎因街 9
# 桥南路 9
# 迪沙鲁路 7
# 笨珍路 6
# ```
# ## Conclusion
# This report covered the most pertinent problems encountered when auditing and cleaning the data set and provided a brief overview of the map data including users, nodes and ways as well as some interesting queries.
#
# Street-level data was generally clean but at the same time, scope for further improvements can be made in the areas highlighted from the preceding section.
#
# [1]:https://www.ura.gov.sg/realEstateIIWeb/resources/misc/list_of_postal_districts.htm
# [2]:https://www.ura.gov.sg/uol/guidelines/development-control/street-naming
# [3]:http://wiki.openstreetmap.org/wiki/Altitude
|
osm-case-study.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:torch] *
# language: python
# name: conda-env-torch-py
# ---
import numpy as np
import cv2
import matplotlib.pyplot as plt
import scipy.io
import glob
import os
from math import sqrt
import json
import torch
import torch.nn as nn
import torchvision
from torchvision.transforms import transforms
from torch.utils.data import DataLoader
from torch.optim import Adam
from torch.autograd import Variable
import pathlib
from PIL import Image
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print(device)
batch_size = 16
lr = 0.001
epochs = 50
training_dir = 'training_vacancy/'
testing_dir = 'testing_vacancy/'
train_count=len(glob.glob(training_dir+'/**/*.jpg'))
test_count=len(glob.glob(testing_dir+'/**/*.jpg'))
print('train : {}, test : {}'.format(train_count,test_count))
transformer = transforms.Compose([
transforms.RandomHorizontalFlip(p = 0.5),
transforms.ToTensor(),
transforms.Normalize((0.5,0.5,0.5),(0.5,0.5,0.5))
])
train_loader = DataLoader(
torchvision.datasets.ImageFolder(training_dir,transform = transformer),
batch_size = 512,
shuffle = True
)
test_loader=DataLoader(
torchvision.datasets.ImageFolder(testing_dir,transform=transformer),
batch_size=32, shuffle=True
)
classes = ['vacant','non-vacant','parking']
class VPSNet(nn.Module):
def __init__(self,num_classes = 3):
super(VPSNet,self).__init__()
self.model = nn.Sequential(
nn.Conv2d(3, 40, kernel_size=(3, 9), stride=(1, 2)),
nn.ReLU(inplace=True),
nn.MaxPool2d(kernel_size=(3, 3), stride=2),
nn.Conv2d(40, 80, kernel_size=(3, 5), padding=(1, 0)),
nn.ReLU(inplace=True),
nn.MaxPool2d(kernel_size=3, stride=2, padding=(1, 0)),
nn.Conv2d(80, 120, kernel_size=3, padding=1),
nn.ReLU(inplace=True),
nn.Conv2d(120, 160, kernel_size=3, padding=1),
nn.ReLU(inplace=True),
nn.MaxPool2d(kernel_size=3, stride=2),
nn.Flatten(),
nn.Dropout(),
nn.Linear(160 * 5 * 5, 512),
nn.ReLU(inplace=True),
nn.Linear(512, num_classes)
)
def forward(self,x):
x = self.model(x)
return x
model = VPSNet(num_classes=3).to(device)
optimizer = Adam(model.parameters(),lr = lr,betas=(0.9,0.999),eps = 1e-08,weight_decay=0.0001)
loss_fn = nn.CrossEntropyLoss()
for epoch in range(epochs):
model.train()
train_accuracy = 0.0
train_loss = 0.0
for i,(images,labels) in enumerate(train_loader):
images = Variable(images.cuda())
labels = Variable(labels.cuda())
optimizer.zero_grad()
loss = loss_fn
optimizer.zero_grad()
outputs = model(images)
loss = loss_fn(outputs,labels)
loss.backward()
optimizer.step()
train_loss += loss.cpu().data*images.size(0)
_,prediction = torch.max(outputs.data,1)
train_accuracy += int(torch.sum(prediction == labels.data))
train_accuracy = train_accuracy/train_count
train_loss = train_loss/train_count
model.eval()
test_accuracy=0.0
for i, (images,labels) in enumerate(test_loader):
print(images.shape)
images=Variable(images.cuda())
labels=Variable(labels.cuda())
outputs=model(images)
_,prediction=torch.max(outputs.data,1)
test_accuracy+=int(torch.sum(prediction==labels.data))
test_accuracy=test_accuracy/test_count
print('epoch : {} , train loss: {} , train accuracy : {}, test accuracy : {}'.format(epoch,train_loss,train_accuracy,test_accuracy))
#save or load model
#save
path = 'model_save/98_test.pth'
#path = 'model_save/98_test.pt'
#torch.save(model.state_dict(),path)
#model.save(path)
#load
model = VPSNet()
model.load_state_dict(torch.load(path))
model.cuda()
#model = torch.load(path)
# +
def image_loader(path):
loader = transforms.Compose([transforms.ToTensor(),transforms.Normalize((0.5,0.5,0.5),(0.5,0.5,0.5))])
image = Image.open(path)
image = loader(image).float()
image = Variable(image)
image = image.unsqueeze(0)
return image.cuda()
def pred_image(model,img_name):
img = image_loader(img_name)
prediction = model(img)
prediction = torch.argmax(prediction.data).cpu().item()
classification = 'vacant'
if (prediction == 2):
classification = 'vacant'
elif (prediction == 0):
classification = 'non-vacant'
elif (prediction == 1):
classification = 'parking'
return classification
# -
#visualize output
model.eval()
sample_id = 177
testing_dir = 'testing_vacancy/'
test_class = 'parking/'
for file in glob.glob(testing_dir+test_class+'/*.jpg'):
classification = pred_image(model,file)
print(classification)
|
vacancy_model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example: shaping the model domain in- and outside
import pychan3d
# In this example we create a full lattice network and carve different shape inside and on the outside of the domain
# before calculating the steady state flow solution.
# +
# We will create a lattice network with a spacing of 10m in each direction
dx, dy, dz = 10., 10., 10.
# and 30 channels in the X-direction (length), 10 channels in the Y- and Z-directions (width and heigth)
nx, ny, nz = 30, 10, 10
# So the resulting total dimensions are 300m x 100m x 100m
# We choose the mean log conductance and a standard deviation for the log conductances which are assumed to be
# lognormally distributed
logC, sigma = -6., 1.
# We create the Lattice network, and use an offset to have the center of the box at coordinates (0., 0., 0.)
LN = pychan3d.LatticeNetwork(nx, ny, nz, dx, dy, dz, logC, logC, logC, sigma, sigma, sigma,
offset=[-150., -50., -50.], seed=987654321)
LN.export2vtk('example2_convex_full_network')
# -
# Now, we carve a cylindrical tunnel at the center of the network, the point on the axis of the cylinder and on the
# 'bottom' surface has coordinates (-51,0,0), the point on the axis and on the 'top' surface has coordinates (51,0,0),
# the cylinder has radius of 2.5m, and we discard the points located inside the cylinder.
# This cylinder can be visualized in Paraview by loading the companion file 'example2_cylindrical_cavity.vtp'.
# We also collect the indices of the nodes located on the cylinder into an array (nodes1).
nodes1 = pychan3d.carve_cylinder(LN, [-51., 0., 0.], [51., 0., 0.], radius=2.5, carve_in=True, create_vtk_output=True)
# We assign fixed head boundary conditions (0m) for all the nodes on the tunnel wall
for n in nodes1:
LN.hbnds[n] = 0.
LN.export2vtk('example2_cylinder_carved_in')
# Now, we trim the outer part of the domain to a cylindrical shape with a radius of 48m.
nodes2 = pychan3d.carve_cylinder(LN, [-151., 0., 0.], [151., 0., 0.], radius=48., carve_in=False, create_vtk_output=True)
# We assign a fixed head boundary condition at the nodes located on the outer surface of the domain (10m)
for n in nodes2:
LN.hbnds[n] = 10.
LN.export2vtk('example2_cylinder_carved_out')
# Inside, the domain we will also carve a convex cavity defined by its corner points, here a cubic box (but it could be
# any convex shape defined a minimum of 4 non-coplanar points).
# This box can be visualized in Paraview by loading the companion file 'example2_convex_cavity.vtp'.
points = [[34., -16., 16.], [34., -16., -16.], [34., 16., 16.], [34., 16., -16.],
[66., 16., 16.], [66., 16., -16.], [66., -16., 16.], [66., -16., -16.]]
nodes3 = pychan3d.carve_convex(LN, points, carve_in=True, create_vtk_output=True)
for n in nodes3: # and we assign fixed head boundary conditions
LN.hbnds[n] = 0.
LN.export2vtk('example2_convex_carved_in')
# We also carve a spherical cavity centered at (-51,0,0) of radius 21m.
# This sphere can be visualized in Paraview by loading the companion file 'example2_spherical_cavity.vtp'.
nodes4 = pychan3d.carve_sphere(LN, [-51., 0., 0.], 21., carve_in=True, create_vtk_output=True)
for n in nodes4: # and we assign fixed head boundary conditions
LN.hbnds[n] = 0.
LN.export2vtk('example2_sphere_carved_in')
# finally we solve the steady state flow problem using the direct solver
LN.solve_steady_state_flow_scipy_direct()
# and export the results to a vtk file for visualization in paraview
LN.export2vtk('example2_flow_data')
# you can compare your result with the provided file given that you did not change the seed value on line16
|
examples/example2/Example2_notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Numbers
#
# The main [numeric types](https://docs.python.org/3/library/stdtypes.html#numeric-types-int-float-complex) in Python are integers, floating point numbers and complex numbers. The syntax for arithmetic operators are: addition `+`, subtraction `-`, multiplication `*`, division `/` and exponentiation `**`.
# ## Integers
#
# Add integers:
8 + 12
# Subtract integers:
2019 - 21
# Multiply integers:
45 * 11
# Divide integers (and notice that division of integers *always* returns a float):
100 / 4
# Compute powers of integers:
2**10
# Use the built-in function `type()` to verify the type of a Python object:
type(42)
# ## Floating Point Numbers
#
# A floating point number (or float) is a real number written in decimal form. Python stores floats and integers in different ways and if we combine integers and floats using arithmetic operations the result is always a float.
#
# Approximate $\sqrt{2} \,$:
2**0.5
# Approximate $2 \pi$:
2 * 3.14159
# Use scientific notation to create $0.00001$:
1e-5
# Again, use the `type()` function to verify the type of a number:
type(42)
type(42.0)
# ## Complex Numbers
#
# Use the built-in function `complex()` to create a complex number in Python or use the letter `j` for $j = \sqrt{-1}$. The built-in function `complex()` takes 2 parameters defining the real and imaginary part of the complex number.
#
# Create the complex number $1 + j$:
complex(1,1)
# Add complex numbers:
(1 + 2j) + (2 - 3j)
# Multiply complex numbers:
(2 - 1j) * (5 + 2j)
# Use the `type()` function to verify the type of a number:
type(2 - 7j)
# <!--
#
# ### Complex Methods
#
# The complex datatype has a few methods. For example, we can access the real and imaginary parts of a complex number:
#
# ```python
# z.real
# ```
#
# ```nohighlight
# 1.0
# ```
#
# ```python
# z.imag
# ```
#
# ```nohighlight
# 1.0
# ```
#
# ```python
# print(z)
# ```
#
# ```nohighlight
# (1+1j)
# ```
#
# The conjugate of a complex number $z = a + b\sqrt{-1}$ is $\overline{z} = a - b\sqrt{-1}$.
#
# ```python
# z.conjugate()
# ```
#
# ```nohighlight
# (1-1j)
# ```
#
# The modulus of a complex number $z = a + b\sqrt{-1}$ is $|z| = \sqrt{a^2 + b^2}$ which is computed by the builtin function `abs` (which is the absolute value when applied to integers and floats).
#
# ```python
# print(z)
# abs(z)
# ```
#
# ```nohighlight
# (1+1j)
# 1.4142135623730951
# ```
#
# ```python
# (1**2 + 1**2)**(0.5)
# ```
#
# ```nohighlight
# 1.4142135623730951
# ```
# -->
# ## Arithmetic Operators
#
# The syntax for arithmetic operators in Python are:
#
# | Operator | Description |
# | :---: | :---: |
# | `+` | addition |
# | `-` | subtraction |
# | `*` | multiplication |
# | `/` | division |
# | `**` | exponentiation |
# | `%` | remainder (or modulo) |
# | `//` | integer division |
#
# Notice that division of integers always returns a float:
4 / 3
# Even if the mathematical result is an integer:
4 / 2
# Use parentheses to group combinations of arithmetic operations:
5 * (4 + 3) - 2
# An integer power of an integer is again an integer:
2**4
# An exponent involving a float is a float:
9**0.5
# The remainder operator computes the remainder of division of integers:
11 % 4
# Integer division is:
11 // 4
# ## Examples
#
# ### Taylor Approximation
#
# The [Taylor series](https://en.wikipedia.org/wiki/Taylor_series) of the exponential function $e^x$ is given by
#
# $$
# e^x = \sum_{k=0}^{\infty} \frac{x^k}{k!}
# $$
#
# Compute the Taylor polynomial of degree 5 evaluated at $x = 1$ to find an approximation of $e$
#
# $$
# e \approx \frac{1}{0!} + \frac{1}{1!} + \frac{1}{2!} + \frac{1}{3!} + \frac{1}{4!} + \frac{1}{5!}
# $$
1 + 1 + 1/2 + 1/(3*2) + 1/(4*3*2) + 1/(5*4*3*2)
# ### Ramanujan's $\pi$ Formula
#
# [Srinivasa Ramanujan](https://en.wikipedia.org/wiki/Srinivasa_Ramanujan#Mathematical_achievements) discovered the following beautiful (and very rapidly converging) series representation of $\pi$
#
# $$
# \frac{1}{\pi} = \frac{2 \sqrt{2}}{99^2} \sum_{k = 0}^{\infty} \frac{(4k)!}{k!^4} \frac{1103 + 26390k}{396^{4k}}
# $$
#
# Let's find an approximation of $\pi$ by computing the *reciprocal* of the sum of the first 3 terms of the series:
#
# $$
# \pi \approx \frac{99^2}{2 \sqrt{2}} \frac{1}{\left( 1103 + 4! \frac{1103 + 26390}{396^{4}} + \frac{8!}{2^4} \frac{1103 + 26390(2)}{396^{8}} \right)}
# $$
99**2 / (2 * 2**0.5) / (1103 + 4*3*2 * (26390 + 1103) / 396**4
+ 8*7*6*5*4*3*2 / 2**4 * (26390*2 + 1103) / 396**8)
# These are exactly the first 16 digits of [$\pi$](https://en.wikipedia.org/wiki/Pi).
# ## Exercises
#
# 1. The [Taylor series](https://en.wikipedia.org/wiki/Taylor_series) of $\cos x$ is given by
#
# $$
# \cos x = \sum_{k=0}^{\infty} (-1)^k \frac{x^{2k}}{(2k)!}
# $$
#
# Compute the Taylor polynomial of degree 6 evaluated at $x=2$:
#
# $$
# \cos(2) \approx 1 - \frac{2^2}{2!} + \frac{2^4}{4!} - \frac{2^6}{6!}
# $$
#
# 2. The [Riemann zeta function](https://en.wikipedia.org/wiki/Riemann_zeta_function) is the infinite series
#
# $$
# \zeta(s) = \sum_{n=1}^{\infty} \frac{1}{n^s}
# $$
#
# and is intimately related to prime numbers by the [Euler product formula](https://en.wikipedia.org/wiki/Riemann_zeta_function#Euler_product_formula)
#
# $$
# \sum_{n=1}^{\infty} \frac{1}{n^s} = \prod_p \left( \frac{1}{1 - p^{-s}} \right)
# $$
#
# where the product is over all primes $p = 2,3,5,7,11,13,\dots$
#
# Compute the 5th partial sum for $s=2$
#
# $$
# 1 + \frac{1}{2^2} + \frac{1}{3^2} + \frac{1}{4^2} + \frac{1}{5^2}
# $$
#
# Compute the 5th partial product for $s=2$
#
# $$
# \left( \frac{1}{1 - 2^{-2}} \right) \left( \frac{1}{1 - 3^{-2}} \right) \left( \frac{1}{1 - 5^{-2}} \right) \left( \frac{1}{1 - 7^{-2}} \right) \left( \frac{1}{1 - 11^{-2}} \right)
# $$
#
# Given [Euler's special value formula](https://en.wikipedia.org/wiki/Basel_problem)
#
# $$
# \zeta(2) = \frac{\pi^2}{6}
# $$
#
# which converges more quickly: the infinite series or product?
#
# 3. The [continued fraction](https://en.wikipedia.org/wiki/Continued_fraction#Square_roots) for $\sqrt{2}$ is given by
#
# $$
# \sqrt{2} = 1 + \frac{1}{2 + \frac{1}{2 + \frac{1}{2 + \frac{1}{2 + \ddots}}}}
# $$
#
# Compute the following (partial) continued fraction to approximate $\sqrt{2}$
#
# $$
# \sqrt{2} \approx 1 + \frac{1}{2 + \frac{1}{2 + \frac{1}{2 + \frac{1}{2}}}}
# $$
|
computing-primer/python/numbers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Part 3
#
# How much data would be needed to use pytorch ann to get feedback?
#
# ## Approach
#
# Transfer learning with fastai
#
# ## References
# https://nlp.fast.ai/
# http://docs.fast.ai/text.html#text
|
Part_3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Overview of Notebooks
#
# These notebooks are included to illustrate a hypothetical Machine Learning project created following best practices.
#
# The goal of this ML project is to predict the heat capacity of inorganic materials given the chemical composition and condition (the measurement temperature).
# We will use both classical ML models as well as neural networks.
#
# To do this, we must:
# 1. Clean and process our dataset, removing obviously erroneous or empty values.
# 1. Partition our data into train, validation, and test splits.
# 1. Featurize our data, turning the chemical formulae into CBFVs.
# 1. Train models on our data and assess the predictive power of the models.
# 1. Compare the performance of the models fairly and reproducibly.
# 1. Visualize the prediction results of the models.
# 1. Share our models and enable others to reproduce your work and aid collaboration.
#
#
# If you require more information about how to use Jupyter notebooks, you can consult:
# * The main README file inside this repository: https://github.com/anthony-wang/BestPractices/blob/master/README.md
# * The official Jupyter Notebook documentation: https://jupyter-notebook.readthedocs.io/en/stable/notebook.html
#
#
# To read the main publication for which these notebooks are made, please see:
#
# Wang, <NAME>; Murdock, <NAME>.; <NAME>.; Oliynyk, <NAME>.; <NAME>; <NAME>; Persson, <NAME>.; <NAME>., [Machine Learning for Materials Scientists: An Introductory Guide toward Best Practices](https://doi.org/10.1021/acs.chemmater.0c01907), *Chemistry of Materials* **Just Accepted Manuscript**, 2020. DOI: [10.1021/acs.chemmater.0c01907](https://doi.org/10.1021/acs.chemmater.0c01907)
#
# Please also consider citing the work if you choose to adopt or adapt the methods and concepts shown in these notebooks or in the publication:
#
# ```bibtex
# @article{Wang2020bestpractices,
# author = {Wang, <NAME> and Murdock, <NAME>. and <NAME>. and Oliynyk, <NAME> Gurlo, <NAME>, <NAME> Sparks, <NAME>.},
# date = {2020},
# title = {Machine Learning for Materials Scientists: An Introductory Guide toward Best Practices},
# issn = {0897-4756},
# journal = {Chemistry of Materials},
# url = {https://doi.org/10.1021/acs.chemmater.0c01907},
# doi = {10.1021/acs.chemmater.0c01907}
# }
# ```
#
|
notebooks/0a-overview.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Images as Grids of Pixels
# ### Import resources
# +
import numpy as np
import matplotlib.image as mpimg # for reading in images
import matplotlib.pyplot as plt
import cv2 # computer vision library
# %matplotlib qt
# -
# ### Read in and display the image
# +
# Read in the image
image = mpimg.imread('images/waymo_car.jpg')
# Print out the image dimensions
print('Image dimensions:', image.shape)
# Change from color to grayscale
gray_image = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
plt.imshow(gray_image, cmap='gray')
# +
# Prints specific grayscale pixel values
# What is the pixel value at x = 400 and y = 300 (on the body of the car)?
x = 400
y = 300
print(gray_image[y,x])
# +
# Finds the maximum and minimum grayscale values in this image
max_val = np.amax(gray_image)
min_val = np.amin(gray_image)
print('Max: ', max_val)
print('Min: ', min_val)
# +
# Create a 5x5 image using just grayscale, numerical values
tiny_image = np.array([[0, 20, 30, 150, 120],
[200, 200, 250, 70, 3],
[50, 180, 85, 40, 90],
[240, 100, 50, 255, 10],
[30, 0, 75, 190, 220]])
# To show the pixel grid, use matshow
plt.matshow(tiny_image, cmap='gray')
## TODO: See if you can draw a tiny smiley face or something else!
## You can change the values in the array above to do this
# -
|
06_Computer_Vision/01_Image_Representation_and_Classification/01_Images_as_numerical_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="8nsx2WMTeKoc" colab_type="text"
#
# # Introduction
#
# ### - This notebook explores a novel convolutional network architechture as discussed in the following research paper to build a classification system for better assistance in diagonosing Acute Lymphoblastic Leukemia in blood cells.
# **Research Paper : http://www.ijcte.org/vol10/1198-H0012.pdf**
#
# ### - The dataset has been taken from : https://homes.di.unimi.it/scotti/all/
# * Here, ALL_IDB2 version of the dataset has been used
#
# * This dataset is completely balanced with equal number of samples in both the classes.
#
#
# ### - Data augmentation ensures that data is large enough and model extracts features efficiently without overfitting and therefore we have analysed two types of data augmentation techniques in this notebook
# * Techniques used in the research paper discussing the following parameters:
#
# 1. Grayscaling of image
# 2. Horizontal reflection
# 3. Vertical reflection
# 4. Gaussian Blurring
# 5. Histogram Equalization
# 6. Rotation
# 7. Translation
# 8. Shearing
#
# (Using these methods, the dataset size increased from 260 images to 2340 images)
#
# * Keras in-built ImageDataGenerator
#
#
# **The dataset was split into 80% and 20% for training and testing respectively.**
#
# ### - The results of our present analysis is:
#
# | Data Augmentation | Accuracy | Precision | Recall | ROC |
# |---|---|---|---|--|
# | Used in paper | 91% | 0.93 | 0.88 | 0.97 |
# | Keras ImageDataGenerator | 76% | 0.74 | 0.79 | 0.82 |
#
#
# **This result has been recorded for maximum number of epochs that model could be trained for without overfitting**
#
# **We can infer that the augmentation used in the paper outperforms the in-built augmentation technique used by Keras**
#
#
# ### Below is the detailed code implementation of this research paper
# + id="o3wkJHL6gKTI" colab_type="code" outputId="c0520ea5-b8b0-4e3a-98de-da5a1659ac11" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 75}
from google.colab import files
files.upload()
# + id="tWyAsmJZlvMW" colab_type="code" colab={}
import zipfile
zip_ref = zipfile.ZipFile('ALL_Data2 (1).zip', 'r')
zip_ref.extractall()
zip_ref.close()
# + id="HwetK3C2pT-C" colab_type="code" outputId="181628e0-a3f3-4b89-a11c-36998809c976" colab={"base_uri": "https://localhost:8080/", "height": 34}
# !ls
# + id="CimI3AcDR3kl" colab_type="code" outputId="16ab3d0d-c47f-4e1a-863d-240f3455d6b3" colab={"base_uri": "https://localhost:8080/", "height": 232}
# !pip install keras_metrics
# + [markdown] id="AfZJS4lHhUXE" colab_type="text"
# ## **Loading requires packages**
# + id="vQ8BE0xmnLD6" colab_type="code" outputId="e8e6a233-273a-40ca-a0eb-876908cd46ef" colab={"base_uri": "https://localhost:8080/", "height": 34}
from pathlib import Path
import glob
import random
import cv2
from numpy.random import seed
from tensorflow import set_random_seed
import pandas as pd
import numpy as np
from sklearn.utils import shuffle
from sklearn.model_selection import train_test_split
from scipy import ndimage
from skimage import exposure
import skimage
from skimage import io
from skimage import transform as tm
import seaborn as sns
import tensorflow as tf
import tensorflow_hub as hub
import keras
from keras.utils import np_utils
from keras.models import Model,Sequential
from keras.layers import Dense,Flatten,Activation
from keras.callbacks import ModelCheckpoint, EarlyStopping
from keras.preprocessing.image import ImageDataGenerator
from keras.layers import Activation, Convolution2D, Dropout, Conv2D
from keras.layers import AveragePooling2D, BatchNormalization
from keras.layers import GlobalAveragePooling2D
from keras.layers import Input,GaussianNoise
from keras.layers import MaxPooling2D
from keras.layers import SeparableConv2D
from keras import layers
from keras.regularizers import l2
import keras_metrics
import matplotlib.pyplot as plt
from keras.applications.vgg16 import VGG16,preprocess_input
from keras.applications.xception import Xception,preprocess_input
from keras.applications.inception_v3 import InceptionV3
from keras.optimizers import Adam,RMSprop,SGD
from sklearn.metrics import confusion_matrix,precision_score,recall_score
from sklearn.metrics import roc_auc_score
from keras import backend as K
# %matplotlib inline
# + id="tH2z2eQB6ZpX" colab_type="code" colab={}
# for consistemt results across multiple executions
seed(3)
set_random_seed(3)
# + [markdown] id="ZpdCO-UDjGnU" colab_type="text"
# ### Reading data and inserting into a dataframe
# + id="DJwo22WNpQjI" colab_type="code" outputId="41f73e88-9898-47ea-8f04-6840905c7f9a" colab={"base_uri": "https://localhost:8080/", "height": 189}
images_dir = Path('img')
images = images_dir.glob("*.tif")
train_data = []
counter = 0
for img in images:
counter += 1
if counter <= 130:
train_data.append((img,1))
else:
train_data.append((img,0))
train_data = pd.DataFrame(train_data,columns=['image','label'],index = None)
train_data = train_data.sample(frac=1.).reset_index(drop=True)
train_data.tail()
# + id="s6_eBeXbKyKj" colab_type="code" outputId="97ad4b62-f830-49d4-f197-71f7bd7df7ee" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(train_data)
# + [markdown] id="zewhaj6Ul0Y_" colab_type="text"
# ## Data Exploration and Augmentation as presented in the paper
#
# ### 8 augmentation techniques have been used here
# 1. Grayscaling of image
# 2. Horizontal reflection
# 3. Vertical reflection
# 4. Gaussian Blurring
# 5. Histogram Equalization
# 6. Rotation
# 7. Translation
# 8. Shearing
# + id="f71MR6OdWyJO" colab_type="code" colab={}
# histogram equalization function
def hist(img):
img_to_yuv = cv2.cvtColor(img,cv2.COLOR_BGR2YUV)
img_to_yuv[:,:,0] = cv2.equalizeHist(img_to_yuv[:,:,0])
hist_equalization_result = cv2.cvtColor(img_to_yuv, cv2.COLOR_YUV2BGR)
return hist_equalization_result
# + id="81rPbbtegU2h" colab_type="code" colab={}
# function to perform rotation on an image
def rotation(img):
rows,cols = img.shape[0],img.shape[1]
randDeg = random.randint(-180, 180)
matrix = cv2.getRotationMatrix2D((cols/2, rows/2), randDeg, 0.70)
rotated = cv2.warpAffine(img, matrix, (rows, cols), borderMode=cv2.BORDER_CONSTANT,
borderValue=(144, 159, 162))
return rotated
# + id="Urz26j6qZJFG" colab_type="code" colab={}
# function to perform shearing of an image
def shear(img):
# Create Afine transform
afine_tf = tm.AffineTransform(shear=0.5)
# Apply transform to image data
modified = tm.warp(img, inverse_map=afine_tf)
return modified
# + id="sYS6DhKxKrvJ" colab_type="code" colab={}
def aug_method(dataframe,dim,method):
if method == 'paper':
n = len(dataframe)
data = np.zeros((n*9,dim,dim,3),dtype = np.float32)
labels = np.zeros((n*9,2),dtype = np.float32)
count = 0
for j in range(0,n):
img_name = dataframe.iloc[j]['image']
label = dataframe.iloc[j]['label']
encoded_label = np_utils.to_categorical(label, num_classes=2)
img = cv2.imread(str(img_name))
img = cv2.resize(img, (dim,dim))
if img.shape[2]==1:
img = np.dstack([img, img, img])
orig_img = img.astype(np.float32)/255.
data[count] = orig_img
labels[count] = encoded_label
aug_img1 = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
aug_img2 = cv2.flip(img, 0)
aug_img3 = cv2.flip(img,1)
aug_img4 = ndimage.gaussian_filter(img, sigma= 5.11)
aug_img5 = hist(img)
aug_img6 = rotation(img)
aug_img7 = cv2.warpAffine(img, np.float32([[1, 0, 84], [0, 1, 56]]), (img.shape[0], img.shape[1]),
borderMode=cv2.BORDER_CONSTANT, borderValue=(144, 159, 162))
aug_img8 = shear(img)
aug_img1 = np.dstack([aug_img1, aug_img1, aug_img1])
aug_img1 = aug_img1.astype(np.float32)/255.
aug_img2 = aug_img2.astype(np.float32)/255.
aug_img3 = aug_img3.astype(np.float32)/255.
aug_img4 = aug_img4.astype(np.float32)/255.
aug_img5 = aug_img5.astype(np.float32)/255.
aug_img6 = aug_img6.astype(np.float32)/255.
aug_img7 = aug_img7.astype(np.float32)/255.
aug_img8 = aug_img8.astype(np.float32)/255.
data[count+1] = aug_img1
labels[count+1] = encoded_label
data[count+2] = aug_img2
labels[count+2] = encoded_label
data[count+3] = aug_img3
labels[count+3] = encoded_label
data[count+4] = aug_img4
labels[count+4] = encoded_label
data[count+5] = aug_img5
labels[count+5] = encoded_label
data[count+6] = aug_img5
labels[count+6] = encoded_label
data[count+7] = aug_img5
labels[count+7] = encoded_label
data[count+8] = aug_img5
labels[count+8] = encoded_label
count +=9
elif method == 'keras':
n = len(dataframe)
data = np.zeros((n,dim,dim,3),dtype = np.float32)
labels = np.zeros((n,2),dtype = np.float32)
count = 0
for j in range(0,n):
img_name = dataframe.iloc[j]['image']
label = dataframe.iloc[j]['label']
encoded_label = np_utils.to_categorical(label, num_classes=2)
img = cv2.imread(str(img_name))
img = cv2.resize(img, (dim,dim))
if img.shape[2]==1:
img = np.dstack([img, img, img])
orig_img = img.astype(np.float32)/255.
data[count] = orig_img
labels[count] = encoded_label
count +=1
return data,labels
# + id="MllJOUPlItHB" colab_type="code" colab={}
data,labels = aug_method(train_data,dim=100,method='paper')
# + id="Tn-7I1xAIrqS" colab_type="code" outputId="30286592-fe1f-4250-b1a2-ed4c1cb2dd05" colab={"base_uri": "https://localhost:8080/", "height": 34}
data.shape
# + id="HBBN9vhqI0JB" colab_type="code" outputId="dde7aefa-2391-40f2-f906-293d86fd2393" colab={"base_uri": "https://localhost:8080/", "height": 34}
labels.shape
# + id="IomV1Dx-I6xD" colab_type="code" outputId="4e6d1d34-667f-4561-ce3a-634246bd64a0" colab={"base_uri": "https://localhost:8080/", "height": 34}
data = np.asarray(data)
labels = np.asarray(labels)
Data,Label = shuffle(data,labels, random_state=3)
data_list = [Data,Label]
type(data_list)
# + id="nLuFjhqsDsAd" colab_type="code" colab={}
y = np.argmax(Label, axis=-1)
# + [markdown] id="viodvfyvmDSS" colab_type="text"
# ## **Visualizing dataset images**
# + id="aB2BiOAmJg5t" colab_type="code" outputId="c0f7da67-4014-41ef-f088-27971be6b1da" colab={"base_uri": "https://localhost:8080/", "height": 444}
f, ax = plt.subplots(4,5, figsize=(30,7))
for i in range(0,20):
ax[i//5, i%5].imshow(Data[i])
if y[i]==1:
ax[i//5, i%5].set_title("Non-ALL")
else:
ax[i//5, i%5].set_title("ALL")
# + [markdown] id="qh2eNGw2U9jK" colab_type="text"
# ### **Splitting into training and test set**
# + id="vVhGNDOp4SpL" colab_type="code" colab={}
(X, y) = (data_list[0],data_list[1])
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=3)
# + [markdown] id="qrAQNIXMpPv7" colab_type="text"
#
#
# ###**The following model was used in the paper**
# Additionaly three dropout layers with different dropout rates have been used to reduce overfitting
# + id="X6-Em20CpBof" colab_type="code" colab={}
model = Sequential()
model.add(Conv2D(16,(5,5),padding='valid',input_shape = X_train.shape[1:]))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2,2),strides=2,padding = 'valid'))
model.add(Dropout(0.4))
model.add(Conv2D(32,(5,5),padding='valid'))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2,2),strides=2,padding = 'valid'))
model.add(Dropout(0.6))
model.add(Conv2D(64,(5,5),padding='valid'))
model.add(Activation('relu'))
model.add(Dropout(0.8))
model.add(Flatten())
model.add(Dense(2,activation = 'softmax'))
# + id="ZnjtIKJqt28j" colab_type="code" outputId="a231148f-178d-485c-c6ac-52e5d1178a29" colab={"base_uri": "https://localhost:8080/", "height": 610}
model.summary()
# + [markdown] id="I60o6qEQleL4" colab_type="text"
# ### Model compilation and fitting
# + id="-Tw9CHEPMz1K" colab_type="code" outputId="70bd06e5-82e9-43fb-a86c-253a95b1804d" colab={"base_uri": "https://localhost:8080/", "height": 162}
batch_size = 100
epochs= 300
optimizer = keras.optimizers.rmsprop(lr = 0.0001, decay = 1e-6)
model.compile(loss = 'binary_crossentropy',optimizer = optimizer, metrics = ['accuracy',keras_metrics.precision(), keras_metrics.recall()])
# + id="_zPlEYfsAGvn" colab_type="code" outputId="89545161-d508-4c16-f555-c4a13ee4ef1d" colab={"base_uri": "https://localhost:8080/", "height": 1000}
history = model.fit(X_train,y_train,steps_per_epoch = int(len(X_train)/batch_size),epochs=epochs)
history
# + id="FbqWD-vXGM6s" colab_type="code" outputId="2f50e8ea-1770-4e46-9e81-1eb7434dbfb9" colab={"base_uri": "https://localhost:8080/", "height": 88}
score = model.evaluate(X_test,y_test,verbose=0)
score
# + id="32dFEYLzJ379" colab_type="code" outputId="69363066-b006-483c-acb9-c845f833195a" colab={"base_uri": "https://localhost:8080/", "height": 34}
y_pred = model.predict_proba(X_test)
roc_auc_score(y_test, y_pred)
# + [markdown] id="iquoDxPyscPR" colab_type="text"
# **Result for 300 epochs**
#
# **1.Accuracy -91%**
#
# **2.Precision -0.93**
#
# **3.Recall -0.88**
#
# **4. AUC score -0.97**
#
# The model stops learning after 300 epochs
#
#
# + [markdown] id="_XwTprDWlM83" colab_type="text"
# ### Visualizing accuracy and loss
# + id="ObK148TAr20z" colab_type="code" outputId="1ec034a7-1ae9-43c6-f51a-98d014ff673b" colab={"base_uri": "https://localhost:8080/", "height": 500}
acc = history.history['acc']
loss = history.history['loss']
plt.figure(figsize=(8, 8))
plt.subplot(2, 1, 1)
plt.plot(acc, label='Training Accuracy')
plt.legend(loc='lower right')
plt.ylabel('Accuracy')
plt.ylim([min(plt.ylim()),1])
plt.title('Training Accuracy')
plt.subplot(2, 1, 2)
plt.plot(loss, label='Training Loss')
plt.legend(loc='upper right')
plt.ylabel('Cross Entropy')
plt.ylim([0,max(plt.ylim())])
plt.title('Training Loss')
plt.show()
# + [markdown] id="rQF_2YQGhXTk" colab_type="text"
# ## Preparing data in order to augment it with Keras Data Generator
# + id="H1NXbrCl0aPX" colab_type="code" colab={}
data,labels = aug_method(train_data,dim=100,method='keras')
# + id="xziUW9yO2ZTh" colab_type="code" outputId="9c2a77bc-d99e-471a-fe36-dd009aaf774c" colab={"base_uri": "https://localhost:8080/", "height": 34}
data.shape
# + id="UVt7rens5mzG" colab_type="code" colab={}
data = np.asarray(data)
labels = np.asarray(labels)
# + id="doSrgp8l3ZTh" colab_type="code" outputId="3862d4c3-eebb-479c-f769-303a279cd580" colab={"base_uri": "https://localhost:8080/", "height": 34}
Data,Label = shuffle(data,labels, random_state=3)
data_list = [Data,Label]
type(data_list)
# + [markdown] id="CtTTdgx7h3_k" colab_type="text"
# ### Train-test split
# + id="yastyD1tKLVz" colab_type="code" colab={}
(X, y) = (data_list[0],data_list[1])
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=3)
# + id="LdaPsrkPQd2Z" colab_type="code" colab={}
# custom functions for histogram equalization , blurring and gray-scale to be fed into keras ImageDataGenerator as pre-processing functions
def AHE(img):
modified = exposure.equalize_adapthist(img, clip_limit=0.03)
return modified
def blur(img):
modified = ndimage.gaussian_filter(img, sigma= 5.11)
return modified
def gray(img):
aug_img = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
modified = np.dstack([aug_img, aug_img, aug_img])
return modified
# + [markdown] id="SOWAJexSlY7Q" colab_type="text"
# ### **Using Keras data augmentation**
# + id="Z6K0lCTkEIzA" colab_type="code" outputId="948d0902-7fa2-4c6e-9a60-ec368df02c83" colab={"base_uri": "https://localhost:8080/", "height": 34}
i = random.randrange(0,3)
if i == 0:
data_generator = ImageDataGenerator(width_shift_range=0.1,height_shift_range=0.1,rotation_range=180,zoom_range=0.1,horizontal_flip = True , vertical_flip = True,shear_range = 0.5,preprocessing_function = AHE)
elif i == 1:
data_generator = ImageDataGenerator(width_shift_range=0.1,height_shift_range=0.1,rotation_range=180,zoom_range=0.1,horizontal_flip = True , vertical_flip = True,shear_range = 0.5,preprocessing_function = blur)
elif i == 2:
data_generator = ImageDataGenerator(width_shift_range=0.1,height_shift_range=0.1,rotation_range=180,zoom_range=0.1,horizontal_flip = True , vertical_flip = True,shear_range = 0.5,preprocessing_function = gray)
print(i)
# + id="wlvF8hyDSunk" colab_type="code" outputId="9d8f4d9a-2436-47a0-e797-2ca173467e59" colab={"base_uri": "https://localhost:8080/", "height": 52}
model = Sequential()
model.add(Conv2D(16,(5,5),padding='valid',input_shape = X_train.shape[1:]))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2,2),strides=2,padding = 'valid'))
model.add(Dropout(0.4))
model.add(Conv2D(32,(5,5),padding='valid'))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2,2),strides=2,padding = 'valid'))
model.add(Dropout(0.6))
model.add(Conv2D(64,(5,5),padding='valid'))
model.add(Activation('relu'))
model.add(Dropout(0.8))
model.add(Flatten())
model.add(Dense(2,activation = 'softmax'))
# + [markdown] id="-rEgByMyljQZ" colab_type="text"
# ### Model compilation and fitting
# + id="SqI8aIEv4U3h" colab_type="code" outputId="dc32379e-3021-436e-d0c1-11514277bdee" colab={"base_uri": "https://localhost:8080/", "height": 1000}
batch_size = 8
epochs = 100
optimizer = keras.optimizers.rmsprop(lr = 0.0001, decay = 1e-6)
model.compile(loss = 'binary_crossentropy',optimizer = optimizer, metrics = ['accuracy',keras_metrics.precision(), keras_metrics.recall()])
history = model.fit_generator(data_generator.flow(X_train,y_train,batch_size = batch_size),steps_per_epoch = int(len(X_train)/batch_size),epochs=epochs)
history
# + id="FxiMOam1_zxI" colab_type="code" outputId="d81c0e72-3dd7-4465-c98f-f9319d057fe5" colab={"base_uri": "https://localhost:8080/", "height": 88}
score = model.evaluate(X_test,y_test,verbose=0)
score
# + id="cPffn6fO_7Bx" colab_type="code" outputId="3421d7a6-0a3a-4dbf-8b72-9fb10887c0bd" colab={"base_uri": "https://localhost:8080/", "height": 34}
y_pred = model.predict_proba(X_test)
roc_auc_score(y_test, y_pred)
# + [markdown] id="c_cozl43HeyM" colab_type="text"
# **Result for 200 epochs**
#
# **1.Accuracy -76%**
#
# **2.Precision -0.74**
#
# **3.Recall -0.79**
#
# **4. AUC score -0.82**
#
# The model stopped learning after 200 epochs
# + [markdown] id="4hfMo8pBlF0S" colab_type="text"
# ### Visualizing accuracy and loss
# + id="FhCRkSZzjapI" colab_type="code" outputId="aa45b99c-6c4e-40f2-b12b-fee494451be1" colab={"base_uri": "https://localhost:8080/", "height": 498}
acc = history.history['acc']
loss = history.history['loss']
plt.figure(figsize=(8, 8))
plt.subplot(2, 1, 1)
plt.plot(acc, label='Training Accuracy')
plt.legend(loc='lower right')
plt.ylabel('Accuracy')
plt.ylim([min(plt.ylim()),1])
plt.title('Training Accuracy')
plt.subplot(2, 1, 2)
plt.plot(loss, label='Training Loss')
plt.legend(loc='upper right')
plt.ylabel('Cross Entropy')
plt.ylim([0,max(plt.ylim())])
plt.title('Training Loss')
plt.show()
|
Python/_Keras/NonQuantisedCode/ALL_Keras.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tensorflow]
# language: python
# name: conda-env-tensorflow-py
# ---
# # XGBoost Article
#
# The data here is taken form the Data Hackathon3.x - http://datahack.analyticsvidhya.com/contest/data-hackathon-3x
# ## Import Libraries:
# +
import os
import pandas as pd
import numpy as np
import xgboost as xgb
from xgboost.sklearn import XGBClassifier
from sklearn import cross_validation, metrics
from sklearn.grid_search import GridSearchCV
from sklearn.model_selection import train_test_split
import matplotlib.pylab as plt
# %matplotlib inline
from matplotlib.pylab import rcParams
rcParams['figure.figsize'] = 12, 4
# -
# ## Load Data:
#
# The data has gone through following pre-processing:
# 1. City variable dropped because of too many categories
# 2. DOB converted to Age | DOB dropped
# 3. EMI_Loan_Submitted_Missing created which is 1 if EMI_Loan_Submitted was missing else 0 | EMI_Loan_Submitted dropped
# 4. EmployerName dropped because of too many categories
# 5. Existing_EMI imputed with 0 (median) - 111 values were missing
# 6. Interest_Rate_Missing created which is 1 if Interest_Rate was missing else 0 | Interest_Rate dropped
# 7. Lead_Creation_Date dropped because made little intuitive impact on outcome
# 8. Loan_Amount_Applied, Loan_Tenure_Applied imputed with missing
# 9. Loan_Amount_Submitted_Missing created which is 1 if Loan_Amount_Submitted was missing else 0 | Loan_Amount_Submitted dropped
# 10. Loan_Tenure_Submitted_Missing created which is 1 if Loan_Tenure_Submitted was missing else 0 | Loan_Tenure_Submitted dropped
# 11. LoggedIn, Salary_Account removed
# 12. Processing_Fee_Missing created which is 1 if Processing_Fee was missing else 0 | Processing_Fee dropped
# 13. Source - top 2 kept as is and all others combined into different category
# 14. Numerical and One-Hot-Coding performed
path = "./data/allstate"
inputFilePath = os.path.join(path, "train.csv.zip")
df = pd.read_csv(inputFilePath, compression="zip", header=0)
msk = np.random.rand(len(df)) < 0.8
train = df[msk]
test = df[~msk]
train.shape, test.shape
target='loss'
IDcol = 'id'
train[target].value_counts()
# ## Define a function for modeling and cross-validation
#
# This function will do the following:
# 1. fit the model
# 2. determine training accuracy
# 3. determine training AUC
# 4. determine testing AUC
# 5. update n_estimators with cv function of xgboost package
# 6. plot Feature Importance
# +
test_results = pd.read_csv('test_results.csv')
def modelfit(alg, dtrain, dtest, predictors,useTrainCV=True, cv_folds=5, early_stopping_rounds=50):
if useTrainCV:
xgb_param = alg.get_xgb_params()
xgtrain = xgb.DMatrix(dtrain[predictors].values, label=dtrain[target].values)
xgtest = xgb.DMatrix(dtest[predictors].values)r
cvresult = xgb.cv(xgb_param, xgtrain, num_boost_round=alg.get_params()['n_estimators'], nfold=cv_folds,
metrics='auc', early_stopping_rounds=early_stopping_rounds, show_progress=False)
alg.set_params(n_estimators=cvresult.shape[0])
#Fit the algorithm on the data
alg.fit(dtrain[predictors], dtrain[target],eval_metric='auc')
#Predict training set:
dtrain_predictions = alg.predict(dtrain[predictors])
dtrain_predprob = alg.predict_proba(dtrain[predictors])[:,1]
#Print model report:
print "\nModel Report"
print "Accuracy : %.4g" % metrics.accuracy_score(dtrain[target].values, dtrain_predictions)
print "AUC Score (Train): %f" % metrics.roc_auc_score(dtrain[target], dtrain_predprob)
# Predict on testing data:
dtest['predprob'] = alg.predict_proba(dtest[predictors])[:,1]
results = test_results.merge(dtest[['ID','predprob']], on='ID')
print 'AUC Score (Test): %f' % metrics.roc_auc_score(results[target], results['predprob'])
feat_imp = pd.Series(alg.booster().get_fscore()).sort_values(ascending=False)
feat_imp.plot(kind='bar', title='Feature Importances')
plt.ylabel('Feature Importance Score')
# -
# ## Step 1- Find the number of estimators for a high learning rate
predictors = [x for x in train.columns if x not in [target, IDcol]]
xgb1 = XGBRegresor(
learning_rate =0.1,
n_estimators=1000,
max_depth=5,
min_child_weight=1,
gamma=0,
subsample=0.8,
colsample_bytree=0.8,
objective= 'binary:logistic',
nthread=4,
scale_pos_weight=1,
seed=27)
modelfit(xgb1, train, test, predictors)
#Grid seach on subsample and max_features
#Choose all predictors except target & IDcols
param_test1 = {
'max_depth':range(3,10,2),
'min_child_weight':range(1,6,2)
}
gsearch1 = GridSearchCV(estimator = XGBClassifier( learning_rate =0.1, n_estimators=140, max_depth=5,
min_child_weight=1, gamma=0, subsample=0.8, colsample_bytree=0.8,
objective= 'binary:logistic', nthread=4, scale_pos_weight=1, seed=27),
param_grid = param_test1, scoring='roc_auc',n_jobs=4,iid=False, cv=5)
gsearch1.fit(train[predictors],train[target])
gsearch1.grid_scores_, gsearch1.best_params_, gsearch1.best_score_
#Grid seach on subsample and max_features
#Choose all predictors except target & IDcols
param_test2 = {
'max_depth':[4,5,6],
'min_child_weight':[4,5,6]
}
gsearch2 = GridSearchCV(estimator = XGBClassifier( learning_rate=0.1, n_estimators=140, max_depth=5,
min_child_weight=2, gamma=0, subsample=0.8, colsample_bytree=0.8,
objective= 'binary:logistic', nthread=4, scale_pos_weight=1,seed=27),
param_grid = param_test2, scoring='roc_auc',n_jobs=4,iid=False, cv=5)
gsearch2.fit(train[predictors],train[target])
gsearch2.grid_scores_, gsearch2.best_params_, gsearch2.best_score_
#Grid seach on subsample and max_features
#Choose all predictors except target & IDcols
param_test2b = {
'min_child_weight':[6,8,10,12]
}
gsearch2b = GridSearchCV(estimator = XGBClassifier( learning_rate=0.1, n_estimators=140, max_depth=4,
min_child_weight=2, gamma=0, subsample=0.8, colsample_bytree=0.8,
objective= 'binary:logistic', nthread=4, scale_pos_weight=1,seed=27),
param_grid = param_test2b, scoring='roc_auc',n_jobs=4,iid=False, cv=5)
gsearch2b.fit(train[predictors],train[target])
gsearch2b.grid_scores_, gsearch2b.best_params_, gsearch2b.best_score_
#Grid seach on subsample and max_features
#Choose all predictors except target & IDcols
param_test3 = {
'gamma':[i/10.0 for i in range(0,5)]
}
gsearch3 = GridSearchCV(estimator = XGBClassifier( learning_rate =0.1, n_estimators=140, max_depth=4,
min_child_weight=6, gamma=0, subsample=0.8, colsample_bytree=0.8,
objective= 'binary:logistic', nthread=4, scale_pos_weight=1,seed=27),
param_grid = param_test3, scoring='roc_auc',n_jobs=4,iid=False, cv=5)
gsearch3.fit(train[predictors],train[target])
gsearch3.grid_scores_, gsearch3.best_params_, gsearch3.best_score_
predictors = [x for x in train.columns if x not in [target, IDcol]]
xgb2 = XGBClassifier(
learning_rate =0.1,
n_estimators=1000,
max_depth=4,
min_child_weight=6,
gamma=0,
subsample=0.8,
colsample_bytree=0.8,
objective= 'binary:logistic',
nthread=4,
scale_pos_weight=1,
seed=27)
modelfit(xgb2, train, test, predictors)
# Tune subsample and colsample_bytree
#Grid seach on subsample and max_features
#Choose all predictors except target & IDcols
param_test4 = {
'subsample':[i/10.0 for i in range(6,10)],
'colsample_bytree':[i/10.0 for i in range(6,10)]
}
gsearch4 = GridSearchCV(estimator = XGBClassifier( learning_rate =0.1, n_estimators=177, max_depth=4,
min_child_weight=6, gamma=0, subsample=0.8, colsample_bytree=0.8,
objective= 'binary:logistic', nthread=4, scale_pos_weight=1,seed=27),
param_grid = param_test4, scoring='roc_auc',n_jobs=4,iid=False, cv=5)
gsearch4.fit(train[predictors],train[target])
gsearch4.grid_scores_, gsearch4.best_params_, gsearch4.best_score_
# tune subsample:
#Grid seach on subsample and max_features
#Choose all predictors except target & IDcols
param_test5 = {
'subsample':[i/100.0 for i in range(75,90,5)],
'colsample_bytree':[i/100.0 for i in range(75,90,5)]
}
gsearch5 = GridSearchCV(estimator = XGBClassifier( learning_rate =0.1, n_estimators=177, max_depth=4,
min_child_weight=6, gamma=0, subsample=0.8, colsample_bytree=0.8,
objective= 'binary:logistic', nthread=4, scale_pos_weight=1,seed=27),
param_grid = param_test5, scoring='roc_auc',n_jobs=4,iid=False, cv=5)
gsearch5.fit(train[predictors],train[target])
gsearch5.grid_scores_, gsearch5.best_params_, gsearch5.best_score_
# Got the same value as assument and no change requried.
# Try regularization:
#Grid seach on subsample and max_features
#Choose all predictors except target & IDcols
param_test6 = {
'reg_alpha':[1e-5, 1e-2, 0.1, 1, 100]
}
gsearch6 = GridSearchCV(estimator = XGBClassifier( learning_rate =0.1, n_estimators=177, max_depth=4,
min_child_weight=6, gamma=0.1, subsample=0.8, colsample_bytree=0.8,
objective= 'binary:logistic', nthread=4, scale_pos_weight=1,seed=27),
param_grid = param_test6, scoring='roc_auc',n_jobs=4,iid=False, cv=5)
gsearch6.fit(train[predictors],train[target])
gsearch6.grid_scores_, gsearch6.best_params_, gsearch6.best_score_
#Grid seach on subsample and max_features
#Choose all predictors except target & IDcols
param_test7 = {
'reg_alpha':[0, 0.001, 0.005, 0.01, 0.05]
}
gsearch7 = GridSearchCV(estimator = XGBClassifier( learning_rate =0.1, n_estimators=177, max_depth=4,
min_child_weight=6, gamma=0.1, subsample=0.8, colsample_bytree=0.8,
objective= 'binary:logistic', nthread=4, scale_pos_weight=1,seed=27),
param_grid = param_test7, scoring='roc_auc',n_jobs=4,iid=False, cv=5)
gsearch7.fit(train[predictors],train[target])
gsearch7.grid_scores_, gsearch7.best_params_, gsearch7.best_score_
xgb3 = XGBClassifier(
learning_rate =0.1,
n_estimators=1000,
max_depth=4,
min_child_weight=6,
gamma=0,
subsample=0.8,
colsample_bytree=0.8,
reg_alpha=0.005,
objective= 'binary:logistic',
nthread=4,
scale_pos_weight=1,
seed=27)
modelfit(xgb3, train, test, predictors)
xgb4 = XGBClassifier(
learning_rate =0.01,
n_estimators=5000,
max_depth=4,
min_child_weight=6,
gamma=0,
subsample=0.8,
colsample_bytree=0.8,
reg_alpha=0.005,
objective= 'binary:logistic',
nthread=4,
scale_pos_weight=1,
seed=27)
modelfit(xgb4, train, test, predictors)
|
XGBoostmodels.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from scipy import ndimage, misc
import matplotlib.pyplot as plt
import cv2
# +
fig = plt.figure()
plt.gray() # show the filtered result in grayscale
ax1 = fig.add_subplot(121) # left side
ax2 = fig.add_subplot(122) # right side
image = cv2.imread('dir_x.png', 0).astype('float')
# IMAGE HAS GRADIENT ONLY IN X DIRECTION --> axis = 0
result = ndimage.sobel(image, axis=0)
ax1.imshow(image)
ax2.imshow(result)
plt.show()
# +
fig = plt.figure()
plt.gray() # show the filtered result in grayscale
ax1 = fig.add_subplot(121) # left side
ax2 = fig.add_subplot(122) # right side
image = cv2.imread('dir_y.png', 0).astype('float')
# IMAGE HAS GRADIENT ONLY IN X DIRECTION --> axis = 1
result = ndimage.sobel(image, axis=1)
ax1.imshow(image)
ax2.imshow(result)
plt.show()
# -
|
exercise_3/Sobel filter_image gradient_axis0vs1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/BigDanVader/COVID_Scrapper/blob/main/HW5.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="E24jzURMokC2" outputId="ce6f0d81-7a95-425a-b753-1392648cf086"
# !pip install -U spacy
# !pip install newsapi-python
import en_core_web_lg
from newsapi import NewsApiClient
import pickle
import pandas as pd
from collections import Counter
import token
nlp = en_core_web_lg.load()
newsapi = NewsApiClient (api_key='8586c8f99df3450f82ad10220d27e4aa')
articles = []
for pagina in range(1, 6):
temp = newsapi.get_everything(q='coronavirus', language='en', from_param='2021-02-24', to='2021-03-23', sort_by='relevancy', page=pagina)
articles.append(temp)
filename = 'articlesCOVID.pckl'
pickle.dump(articles, open(filename, 'wb'))
filename = 'articlesCOVID.pckl'
loaded_model = pickle.load(open(filename, 'rb'))
filepath = '/articlesCOVID.pckl'
pickle.dump(loaded_model, open(filepath, 'wb'))
dados = []
for i, article in enumerate(articles):
for x in article['articles']:
title = x['title']
date = x['publishedAt']
description = x['description']
content = x['content']
dados.append({'title':title, 'date':date, 'desc':description, 'content':content})
df = pd.DataFrame(dados)
df = df.dropna()
df.head()
def get_keywords_eng(record):
doc = nlp(content)
pos_tag = ['NOUN', 'VERB', 'PROPN']
result = []
for token in doc:
if (token.text in nlp.Defaults.stop_words or token.is_punct ):
continue
if (token.pos_ in pos_tag):
result.append(token.text)
return result
results = []
for content in df.content.values:
results.append([('#' + x[0]) for x in Counter(get_keywords_eng(content)).most_common(5)])
df['keywords'] = results
df.to_csv(r'/output.csv')
|
HW5.ipynb
|