code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import numpy as np
from sklearn.naive_bayes import MultinomialNB
from sklearn.metrics import confusion_matrix
from sklearn.svm import LinearSVC
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.model_selection import StratifiedKFold
import pandas as pd
fileLoc = 'trainDataMovieReviews.csv'
movieData = pd.read_csv(fileLoc)
corpus = []
for desc in movieData['review']:
corpus.append(desc)
#Stratified 10-cross fold validation with SVM and Multinomial NB
labels = movieData.iloc[:,-1]
labels = labels.tolist()
for i in range(len(labels)):
if (labels[i] == 'pos'):
labels[i] = 1
if (labels[i] == 'neg'):
labels[i] = -1
labels = np.array(labels)
kf = StratifiedKFold(n_splits=10)
totalsvm = 0 # Accuracy measure on 2000 files
totalNB = 0
totalMatSvm = np.zeros((2,2)); # Confusion matrix on 2000 files
totalMatNB = np.zeros((2,2));
for train_index, test_index in kf.split(corpus,labels):
X_train = [corpus[i] for i in train_index]
X_test = [corpus[i] for i in test_index]
y_train, y_test = labels[train_index], labels[test_index]
vectorizer = TfidfVectorizer(min_df=5, max_df = 0.6, sublinear_tf=True, use_idf=True,stop_words='english')
train_corpus_tf_idf = vectorizer.fit_transform(X_train)
test_corpus_tf_idf = vectorizer.transform(X_test)
model1 = LinearSVC()
model2 = MultinomialNB()
model1.fit(train_corpus_tf_idf,y_train)
model2.fit(train_corpus_tf_idf,y_train)
result1 = model1.predict(test_corpus_tf_idf)
result2 = model2.predict(test_corpus_tf_idf)
totalMatSvm = totalMatSvm + confusion_matrix(y_test, result1)
totalMatNB = totalMatNB + confusion_matrix(y_test, result2)
totalsvm = totalsvm+sum(y_test==result1)
totalNB = totalNB+sum(y_test==result2)
print(totalMatSvm, totalsvm/25000.0, totalMatNB, totalNB/25000.0)
# -
print(result1)
print(totalMatSvm)
print(result2)
print(totalMatNB)
pd.read_csv(testset)
| src/united.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 1. **히라가나/가타카나를 제거한 후에도 일본어 가사가 한글로 포함되어 있는 경우 전처리**
# <br> --> contains로 확인한뒤 행제거 반복 --> 현재 429곡 제거됨
# 2. **가사가 모두 영어, 중국어인 경우 전처리**
# <br> --> 가사에 한글이 하나도 들어가지 않는 행 제거 --> 현재 895곡 제거됨
import pandas as pd
import re
d1 = pd.read_csv('song_data_yewon_ver01.csv')
d1
d1.loc[d1['lyrics'].str.contains(r'(와타시|혼토|아노히|혼또|마센|에가이|히토츠|후타츠|마치노|몬다이|마에노|아메가)', regex=True)]
d2 = d1[d1.lyrics.str.contains(r'(와타시|혼토|아노히|혼또|마센|에가이|히토츠|후타츠|마치노|몬다이|마에노|아메가)') == False]
d2
d2.loc[d2['lyrics'].str.contains(r'(히카리|미라이|오나지|춋|카라다|큥|즛또|나캇|토나리|못또|뎅와|코이|히토리|맛스구|후타리|케시키|쟈나이|잇슌|이츠모|아타라|덴샤|즈쿠|에가오|소라오|난테|고멘네|아이시테|다키시|유메|잇탄다|소레|바쇼)', regex=True)]
d3 = d2[d2.lyrics.str.contains(r'(히카리|미라이|오나지|춋|카라다|큥|즛또|나캇|토나리|못또|뎅와|코이|히토리|맛스구|후타리|케시키|쟈나이|잇슌|이츠모|아타라|덴샤|즈쿠|에가오|소라오|난테|고멘네|아이시테|다키시|유메|잇탄다|소레|바쇼)') == False]
d3
d3.loc[d3['lyrics'].str.contains(r'(키미니|보쿠|세카이|도코데|즛토|소바니|바쇼|레루|스베테|탓테|싯테|요쿠)', regex=True)]
d4 = d3[d3.lyrics.str.contains(r'(키미니|보쿠|세카이|도코데|즛토|소바니|바쇼|레루|스베테|탓테|싯테|요쿠)') == False]
d4
# ---------------------------------여기까지 일본어전처리------------------429곡 제거---------------------------
d4.loc[d4['lyrics'].str.contains(r'[가-힣]+', regex=True)]
# 한글이 한글자라도 나오는 것만 저장합니다.
d5 = d4[d4.lyrics.str.contains(r'[가-힣]+') == True]
d5
d5.to_csv('song_data_yewon_ver02.csv', index=False)
| SongTidy/song_tidy_yewon_ver02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
from matplotlib import pyplot
import sys
from keras.datasets import cifar10
from keras.models import Sequential
from keras.utils import to_categorical
from keras.models import Sequential
from keras.layers import Conv2D
from keras.layers import MaxPooling2D
from keras.layers import Dense
from keras.layers import Flatten
from keras.optimizers import SGD
from keras.optimizers import Adam
# +
# load train and test dataset
def load_dataset ():
# load dataset
(X_train, y_train), (X_test, y_test) = cifar10.load_data ()
# one hot encode target values
y_train = to_categorical (y_train)
y_test = to_categorical (y_test)
return X_train, y_train, X_test, y_test
def get_larger_model():
model = Sequential()
model.add(Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same', input_shape=(32, 32, 3)))
model.add(Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
model.add(MaxPooling2D((2, 2)))
model.add(tf.keras.layers.Dropout(0.1))
model.add(Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
model.add(Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
model.add(MaxPooling2D((2, 2)))
model.add(tf.keras.layers.Dropout(0.2))
model.add(Conv2D(128, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
model.add(Conv2D(128, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
model.add(MaxPooling2D((2, 2)))
model.add(tf.keras.layers.Dropout(0.1))
model.add(Flatten())
model.add(Dense(128, activation='relu', kernel_initializer='he_uniform'))
model.add(tf.keras.layers.Dropout(0.2))
model.add(Dense(10, activation='softmax'))
# compile model
opt = SGD(lr=0.0001, momentum=0.9)
#model.compile(optimizer=opt, loss='categorical_crossentropy', metrics=['accuracy'])
model.compile(optimizer=opt, loss='categorical_crossentropy', metrics=['accuracy'])
model.summary ()
return model
def summarize_diagnostics (history):
# plot loss
pyplot.subplot (211)
pyplot.title ('Cross Entropy Loss')
pyplot.plot (history.history ['loss'], color='blue', label='train')
pyplot.plot (history.history ['val_loss'], color='orange', label='test')
# plot accuracy
pyplot.subplot (212)
pyplot.title ('Classification Accuracy')
pyplot.plot (history.history ['accuracy'], color='blue', label='train')
pyplot.plot (history.history ['val_accuracy'], color='orange', label='test')
# save plot to file
filename = sys.argv [0].split ('/') [-1]
pyplot.savefig (filename + '_plot.png')
pyplot.close ()
def main ():
EPOCH_BATCHES = 1
X_train, y_train, X_test, y_test = load_dataset ()
model = get_larger_model ()
print ('New model.')
for _ in range (EPOCH_BATCHES):
history = model.fit (X_train, y_train, epochs=2, batch_size=128, validation_split=0.2, verbose=1)
model.save ('./models/cnn.h5')
summarize_diagnostics (history)
# -
main ()
| src/specific_models/n-federated/clustering_metric_evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1. PubMed
#
# Search PubMed for papers
#
# https://www.ncbi.nlm.nih.gov/pubmed/
#
# https://www.ncbi.nlm.nih.gov/books/NBK25499/
# +
from collections import Counter
import os
from textwrap import wrap
from Bio import Entrez
from IPython.display import display
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import source.reuse as reuse
# +
ENTREZ_EMAIL = '<EMAIL>'
# base_query = '("Mathematical Concepts"[Mesh] OR "Operations Research"[Mesh] OR "Game Theory"[Mesh] OR "Markov Chains"[Mesh] OR "Heuristics"[Mesh] OR "robust optimization" OR "queuing systems" OR "operations research" OR "operational research" OR "markov decision process" OR "stochastic program" OR "stochastic processes" OR "combinatorial optimization" OR "discrete optimization" OR "approximation algorithms" OR "heuristics" OR "dynamic program" OR "dynamic programming" OR "linear program" OR "linear programming" OR "integer program" OR "integer programming" OR "mixed-integer program" OR "mixed-integer programming" OR "stochastic optimization" OR "convex optimization" OR "quadratic optimization" OR "quadratic program" OR "quadratic programming" OR "non-smooth optimization" OR "non-convex optimization" OR "multicriteria optimization" OR "goal programming" OR "queuing theory" OR "game theory" OR "tabu search" OR "genetic algorithm" OR "simulated annealing" OR "variable neighborhood search" OR "ant colony") AND ("Clinical Decision-Making"[Mesh] OR "Decision Support Techniques"[Mesh] OR "Decision Support Systems, Clinical"[Mesh] OR "Decision Making"[Mesh] OR "Decision Theory"[Mesh] OR "Clinical Decision Rules"[Mesh] OR "Decision Trees"[Mesh] OR "Cost-Benefit Analysis"[Mesh] OR "decision") AND ("Radiotherapy"[Mesh]) NOT "Radiotherapy Setup Errors"[Mesh] NOT "Uncertainty"[Mesh] NOT "Disaster Planning"[Mesh] NOT "Meta-Analysis" [Publication Type] NOT "Bionics"[Mesh] NOT "Quality Assurance, Health Care"[Mesh] NOT "Cells, Cultured"[Mesh] NOT "Survival Analysis"[Mesh] NOT "Quality Control"[Mesh] NOT "Retrospective Studies"[Mesh] NOT "Observational Study"[Publication Type] NOT "Phantoms, Imaging"[Mesh] NOT "Radiation Protection"[Mesh] NOT "Cryopreservation"[Mesh] NOT "Radiometry"[Mesh] NOT "Clinical Trial"[Publication Type] NOT "Software"[Mesh] NOT "Legal Case"[Publication Type] NOT "Anisotropy"[Mesh] NOT "Diagnosis, Differential"[Mesh] NOT "Patient Positioning"[Mesh] NOT "Radiology, Interventional"[Mesh] NOT "Algorithms"[Mesh]'
# extra_query = '("Mathematical Concepts"[Mesh] OR "Operations Research"[Mesh] OR "Game Theory"[Mesh] OR "Markov Chains"[Mesh] OR "Heuristics"[Mesh] OR "robust optimization" OR "queuing systems" OR "operations research" OR "operational research" OR "markov decision process" OR "stochastic program" OR "stochastic processes" OR "combinatorial optimization" OR "discrete optimization" OR "approximation algorithms" OR "heuristics" OR "dynamic program" OR "dynamic programming" OR "linear program" OR "linear programming" OR "integer program" OR "integer programming" OR "mixed-integer program" OR "mixed-integer programming" OR "stochastic optimization" OR "convex optimization" OR "quadratic optimization" OR "quadratic program" OR "quadratic programming" OR "non-smooth optimization" OR "non-convex optimization" OR "multicriteria optimization" OR "goal programming" OR "queuing theory" OR "game theory" OR "tabu search" OR "genetic algorithm" OR "simulated annealing" OR "variable neighborhood search" OR "ant colony") AND ("Clinical Decision-Making"[Mesh] OR "Decision Support Techniques"[Mesh] OR "Decision Support Systems, Clinical"[Mesh] OR "Decision Making"[Mesh] OR "Decision Theory"[Mesh] OR "Clinical Decision Rules"[Mesh] OR "Decision Trees"[Mesh] OR "Cost-Benefit Analysis"[Mesh] OR "decision") AND ("Radiotherapy"[Mesh] OR "radiotherapy") NOT "Radiotherapy Setup Errors"[Mesh] NOT "Uncertainty"[Mesh] NOT "Disaster Planning"[Mesh] NOT "Meta-Analysis" [Publication Type] NOT "Bionics"[Mesh] NOT "Quality Assurance, Health Care"[Mesh] NOT "Cells, Cultured"[Mesh] NOT "Survival Analysis"[Mesh] NOT "Quality Control"[Mesh] NOT "Retrospective Studies"[Mesh] NOT "Observational Study"[Publication Type] NOT "Phantoms, Imaging"[Mesh] NOT "Radiation Protection"[Mesh] NOT "Cryopreservation"[Mesh] NOT "Radiometry"[Mesh] NOT "Clinical Trial"[Publication Type] NOT "Software"[Mesh] NOT "Legal Case"[Publication Type] NOT "Anisotropy"[Mesh] NOT "Diagnosis, Differential"[Mesh] NOT "Patient Positioning"[Mesh] NOT "Radiology, Interventional"[Mesh] NOT "DNA"[Mesh] NOT "Radiotherapy Planning, Computer-Assisted"[Mesh] NOT "Machine Learning"[Mesh] NOT "Artificial Intelligence"[Mesh] NOT "Prognosis"[Mesh]'
base_query = '("Stochastic Processes"[Mesh] OR "Operations Research"[Mesh] OR "operations research" OR "operational research" OR "Markov Model" OR "Markov models" OR "Markov chain" OR "Markov chains" OR "Markov decision") AND ("Radiotherapy"[Mesh] OR "radiotherapy" OR "radiation therapy") AND ("Clinical Decision-Making"[Mesh] OR "Decision Support Techniques"[Mesh] OR "Decision Support Systems, Clinical"[Mesh] OR "Decision Making"[Mesh] OR "Decision Theory"[Mesh] OR "Clinical Decision Rules"[Mesh] OR "Decision Trees"[Mesh] OR "decision") NOT "Anisotropy"[Mesh] NOT "Artificial Intelligence"[Mesh] NOT "Bionics"[Mesh] NOT "Cells, Cultured"[Mesh] NOT "Clinical Trial"[Publication Type] NOT "Cryopreservation"[Mesh] NOT "Diagnosis, Differential"[Mesh] NOT "Disaster Planning"[Mesh] NOT "DNA"[Mesh] NOT "Legal Case"[Publication Type] NOT "Machine Learning"[Mesh] NOT "Meta-Analysis" [Publication Type] NOT "Observational Study"[Publication Type] NOT "Patient Positioning"[Mesh] NOT "Phantoms, Imaging"[Mesh] NOT "Radiation Protection"[Mesh] NOT "Radiology, Interventional"[Mesh] NOT "Radiometry"[Mesh] NOT "Radiotherapy Planning, Computer-Assisted"[Mesh] NOT "Radiotherapy Setup Errors"[Mesh] NOT "Retrospective Studies"[Mesh] NOT "Software"[Mesh] NOT "Survival Analysis"[Mesh] NOT "Quality Assurance, Health Care"[Mesh] NOT "Quality Control"[Mesh] AND "English" [LA] AND 2000:2022[dp]'
extra_query = '("Stochastic Processes"[Mesh] OR "Operations Research"[Mesh] OR "operations research" OR "operational research" OR "Markov Model" OR "Markov models" OR "Markov chain" OR "Markov chains") AND ("Radiotherapy"[Mesh] OR "radiotherapy" OR "radiation therapy") AND ("Clinical Decision-Making"[Mesh] OR "Decision Support Techniques"[Mesh] OR "Decision Support Systems, Clinical"[Mesh] OR "Decision Making"[Mesh] OR "Decision Theory"[Mesh] OR "Clinical Decision Rules"[Mesh] OR "Decision Trees"[Mesh] OR "decision") NOT "Anisotropy"[Mesh] NOT "Artificial Intelligence"[Mesh] NOT "Bionics"[Mesh] NOT "Cells, Cultured"[Mesh] NOT "Clinical Trial"[Publication Type] NOT "Cryopreservation"[Mesh] NOT "Diagnosis, Differential"[Mesh] NOT "Disaster Planning"[Mesh] NOT "DNA"[Mesh] NOT "Legal Case"[Publication Type] NOT "Machine Learning"[Mesh] NOT "Meta-Analysis" [Publication Type] NOT "Observational Study"[Publication Type] NOT "Patient Positioning"[Mesh] NOT "Prognosis"[Mesh] NOT "Phantoms, Imaging"[Mesh] NOT "Radiation Protection"[Mesh] NOT "Radiology, Interventional"[Mesh] NOT "Radiometry"[Mesh] NOT "Radiotherapy Planning, Computer-Assisted"[Mesh] NOT "Radiotherapy Setup Errors"[Mesh] NOT "Retrospective Studies"[Mesh] NOT "Software"[Mesh] NOT "Survival Analysis"[Mesh] NOT "Quality Assurance, Health Care"[Mesh] NOT "Quality Control"[Mesh] AND "English" [LA] AND 2000:2022[dp]'
# refined_query = '("Stochastic Processes"[Mesh] OR "Operations Research"[Mesh] OR "operations research" OR "operational research" OR "Markov Model" OR "Markov models" OR "Markov chain" OR "Markov chains") AND ("Radiotherapy"[Mesh] OR "radiotherapy" OR "radiation therapy") AND ("Clinical Decision-Making"[Mesh] OR "Decision Support Techniques"[Mesh] OR "Decision Support Systems, Clinical"[Mesh] OR "Decision Making"[Mesh] OR "Decision Theory"[Mesh] OR "Clinical Decision Rules"[Mesh] OR "Decision Trees"[Mesh] OR "decision") NOT "Anisotropy"[Mesh] NOT "Artificial Intelligence"[Mesh] NOT "Bionics"[Mesh] NOT "Cells, Cultured"[Mesh] NOT "Clinical Trial"[Publication Type] NOT "Cryopreservation"[Mesh] NOT "Diagnosis, Differential"[Mesh] NOT "Disaster Planning"[Mesh] NOT "DNA"[Mesh] NOT "Legal Case"[Publication Type] NOT "Machine Learning"[Mesh] NOT "Meta-Analysis" [Publication Type] NOT "Observational Study"[Publication Type] NOT "Patient Positioning"[Mesh] NOT "Prognosis"[Mesh] NOT "Phantoms, Imaging"[Mesh] NOT "Radiation Protection"[Mesh] NOT "Radiology, Interventional"[Mesh] NOT "Radiometry"[Mesh] NOT "Radiotherapy Planning, Computer-Assisted"[Mesh] NOT "Radiotherapy Setup Errors"[Mesh] NOT "Retrospective Studies"[Mesh] NOT "Software"[Mesh] NOT "Survival Analysis"[Mesh] NOT "Quality Assurance, Health Care"[Mesh] NOT "Quality Control"[Mesh] NOT "Uncertainty"[Mesh] AND "English" [LA] AND 2000:2022[dp]'
search_strings = [
base_query,
extra_query
# refined_query
]
# -
search_results_all = reuse.search_list(search_strings, ENTREZ_EMAIL, all=True)
# Display number of results
for ss in search_strings:
result = search_results_all[ss]
print(f'{ss}:\n - Count: {len(result.index)}')
# +
# Filter out to get the new URLs
all_urls = [list(search_results_all[k]['URL'].values) for k in search_results_all.keys()]
# If a new restriction is just added
all_urls = [i for sl in all_urls for i in sl]
new_urls = [l for l,c in dict(Counter(all_urls)).items() if c == 1]
new_pubs = search_results_all[list(search_results_all.keys())[0]]
# If a totally new search to add new publications
# new_urls = [l for l in all_urls[1] if l not in all_urls[0]]
# new_pubs = search_results_all[list(search_results_all.keys())[-1]]
new_pubs = new_pubs[new_pubs['URL'].isin(new_urls)]
# +
# Write the titles to files
write_dir = os.path.join('search_results', 'pubmed')
os.makedirs(write_dir, exist_ok=True)
new_pubs.to_csv(os.path.join(write_dir, 'new-publications_refined-revised.csv'), index=False)
# search_results_all[base_query].to_csv(os.path.join(write_dir, 'without-constraints-all.csv'), index=False)
# search_results_all[full_query].to_csv(os.path.join(write_dir, 'with-constraints-all.csv'), index=False)
# base_query_file = os.path.join(write_dir, 'without-constraints.txt')
# with open(base_query_file, 'w') as f:
# for line in search_results[base_query].paper_titles:
# f.write(line+'\n')
# full_query_file = os.path.join(write_dir, 'with-constraints.txt')
# with open(full_query_file, 'w') as f:
# for line in search_results[full_query].paper_titles:
# f.write(line+'\n')
# -
# Create a new one for each year
all_years = sorted(set(list(search_results_all[base_query]['Publication Year'])))
# all_years = sorted(set(list(search_results_all[full_query]['Publication Year'])))
for year in all_years:
year_df_without = search_results_all[base_query][search_results_all[base_query]['Publication Year'] == year]
# year_df_with = search_results_all[full_query][search_results_all[full_query]['Publication Year'] == year]
year_df_without.to_csv(os.path.join(write_dir, f'without-constraints-all_{year}.csv'), index=False)
# year_df_with.to_csv(os.path.join(write_dir, f'with-constraints-all_{year}.csv'), index=False)
# +
# Create a histogram of number of publications each year
all_years = [int(y) for y in sorted(set(list(search_results_all[base_query]['Publication Year'])))]
# all_years = [int(y) for y in sorted(set(list(search_results_all[full_query]['Publication Year'])))]
year_without = []
year_with = []
interval_years = range(min(all_years), max(all_years)+1)
for year in interval_years:
year_df_without = search_results_all[base_query][search_results_all[base_query]['Publication Year'] == str(year)]
# year_df_with = search_results_all[full_query][search_results_all[full_query]['Publication Year'] == str(year)]
for _ in range(len(year_df_without.index)):
year_without.append(year)
# for _ in range(len(year_df_with.index)):
# year_with.append(year)
plt.figure(figsize=(10,8))
h = plt.hist(year_without, facecolor='k', edgecolor='w', bins=np.arange(min(interval_years)-1, max(interval_years)+5)-0.5)
plt.xlabel('Year', fontsize=12)
plt.ylabel('Number of Publications', fontsize=12)
plt.ylim([0, 1.05*max(h[0])])
plt.title('\n'.join(wrap(search_strings[0], 140)), fontsize=8)
plt.annotate(f'N = {len(year_without)}', xy=(0.05,0.9), xytext=(0.05,0.9), xycoords='axes fraction', size=16)
plt.tight_layout()
plt.savefig(os.path.join(write_dir, 'without-constraints_histogram.jpg'))
plt.show()
# plt.figure(figsize=(10,6))
# plt.hist(year_with, facecolor='k', edgecolor='w', bins=np.arange(min(interval_years)-1, max(interval_years)+5)-0.5)
# plt.xlabel('Year', fontsize=12)
# plt.ylabel('Number of Publications', fontsize=12)
# plt.ylim([0, 1.05*max(h[0])])
# plt.title('\n'.join(wrap(search_strings[-1], 140)), fontsize=8)
# plt.savefig(os.path.join(write_dir, 'with-constraints_histogram.jpg'))
# plt.show()
# +
# Assign authors to articles to review
CANCER_AUTHORS = ['Lucas', 'Cem']
OPTIM_AUTHORS = ['Soheil', 'Mohammad', 'Aysenur']
ALL_AUTHORS = [*CANCER_AUTHORS, *OPTIM_AUTHORS]
# Create the dataframe from a subset of the original
assignment_file = new_pubs[['Title', 'URL']]
# assignment_file = search_results_all[base_query][['Title', 'URL']]
# assignment_file = search_results_all[full_query][['Title', 'URL']]
total_publications = len(assignment_file.index)
# Cycle through the authors to make it even
optim_index = 0
cancer_index = 0
reviewer1s = []
reviewer2s = []
for p in range(total_publications):
# Cycle through all authors
reviewer1s.append(ALL_AUTHORS[p%len(ALL_AUTHORS)])
# Match authors with their opposite domain
if reviewer1s[p] in CANCER_AUTHORS:
reviewer2s.append(OPTIM_AUTHORS[optim_index%len(OPTIM_AUTHORS)])
optim_index += 1
elif reviewer1s[p] in OPTIM_AUTHORS:
reviewer2s.append(CANCER_AUTHORS[cancer_index%len(CANCER_AUTHORS)])
cancer_index += 1
# Append to the dataframe
assignment_file['Reviewer1'] = reviewer1s
assignment_file['Decision1'] = total_publications*['']
assignment_file['Comments1'] = total_publications*['']
assignment_file['Reviewer2'] = reviewer2s
assignment_file['Decision2'] = total_publications*['']
assignment_file['Comments2'] = total_publications*['']
assignment_file = assignment_file.sort_values(by=['Reviewer1'])
assignment_file.to_csv(os.path.join(write_dir, f'new-publications_assignment-file.csv'), index=False)
# assignment_file.to_csv(os.path.join(write_dir, f'without-constraints_assignment-file.csv'), index=False)
# assignment_file.to_csv(os.path.join(write_dir, f'with-constraints_assignment-file.csv'), index=False)
# +
# Total
print('Total:\n')
print('Author | Reviewer 1 | Reviewer 2')
print('---------|------------|-----------')
for a in ALL_AUTHORS:
c1 = reviewer1s.count(a)
c2 = reviewer2s.count(a)
print(f'{a:8} | {c1:^10} | {c2:^10}')
# Combinations
print('\nCombinations:\n')
print(dict(Counter(tuple(sorted(tup)) for tup in list(zip(reviewer1s, reviewer2s)))))
# -
# # Read in results and filter by publication year
input_file = os.path.join('search_results', 'pubmed', 'review_filtered_04-23-2022.xlsx')
review_results = pd.read_excel(input_file)
# +
pmids = [int(r.split('/')[-2]) for r in review_results['URL'].to_list()]
Entrez.email = ENTREZ_EMAIL
all_years = []
for pmid in pmids:
handle = Entrez.efetch(db='pubmed', retmode='xml', id=pmid)
results = Entrez.read(handle)
try:
all_years.append(results['PubmedArticle'][0]['MedlineCitation']['DateCompleted']['Year'])
except TypeError:
print(results)
plt.hist(all_years)
# -
# Look at the effect of restricting the search by additional criteria.
# The differences show that many false positives, and a few true positives, are removed.
reuse.showdiff(search_results_all[search_strings[0]],
search_results_all[search_strings[1]])
# +
# Read in the labelled results for the general unconstrained search query
labelled_results = pd.read_csv(os.path.join(write_dir, 'without-constraints-inspected.tsv'), delimiter='\t', header=None)
false_positives = labelled_results.loc[labelled_results[1]=='F'][0].values
true_positives = labelled_results.loc[labelled_results[1]=='T'][0].values
print('Number of results found using the unconstrained search term:', len(labelled_results))
print('Number of false positives:', len(false_positives))
print('Number of true positives:', len(true_positives))
constrained_titles = search_results[search_strings[1]].paper_titles
print('\nCompare ^ true positives with:')
print('Number of results from the constrained search term:', len(constrained_titles))
missed_papers = set(true_positives) - set(constrained_titles)
print('Number of missed true positives:', len(missed_papers))
# +
# Take a look at some True positives missed by the constrained search term to figure out what else you can add.
# Write to a file to label comments.
write_dir = os.path.join('search_results', 'pubmed')
missed_papers_file = os.path.join(write_dir, 'missed-papers.tsv')
with open(missed_papers_file, 'w') as f:
for line in missed_papers:
f.write(line+'\n')
display(missed_papers)
# -
# # 2. Web of Science
# +
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
import re
import time
from IPython.display import display
base_mimic_query = '(mimic-ii OR mimic-iii OR mimicii OR mimiciii OR mimic-2 OR mimic-3 OR mimic2 OR mimic3)'
restriction_query = '(physionet OR icu OR “intensive care” OR “critical care”)'
def full_query(base_query, restriction_query):
return ' AND '.join([base_query, restriction_query])
full_mimic_query = full_query(base_mimic_query, restriction_query)
#base_search_url = 'https://apps.webofknowledge.com/WOS_GeneralSearch_input.do?product=WOS&search_mode=GeneralSearch&SID=2F46AeWkMQBRAZlzDWm&preferencesSaved='
base_search_url = 'https://apps.webofknowledge.com/WOS_GeneralSearch_input.do?product=WOS&search_mode=GeneralSearch&SID=1AnC2UMojuKrtrl7T5R&preferencesSaved='
all_titles = []
# +
# Get to the search page
driver = webdriver.Firefox()
driver.get(base_search_url)
# Input the query string
time.sleep(2.5)
searchbox = driver.find_element_by_id('value(input1)')
searchbox.send_keys(full_mimic_query)
# Search
time.sleep(1)
searchbutton = driver.find_element_by_css_selector('.standard-button.primary-button.large-search-button')
searchbutton.click()
# Get the total number of pages
npages = int(driver.find_element_by_id('pageCount.top').text)
# +
# Get the titles!!!
while True:
# Get the current page number
pagenum = int(driver.find_element_by_class_name('goToPageNumber-input').get_property('value'))
# Get the titles. This also captures the journals. So every second value is not a title.
elements = driver.find_elements_by_class_name('smallV110')
for e in elements[::2]:
all_titles.append(e.find_element_by_tag_name('value').text)
if pagenum < npages:
nextbutton = driver.find_element_by_class_name('paginationNext')
nextbutton.click()
else:
print('Got all paper titles!')
driver.close()
break
all_titles = set(all_titles)
#all_titles.remove('')
all_titles = [t.lower() for t in list(all_titles)]
# -
display(all_titles)
# +
# Write the titles to files
write_dir = os.path.join('search_results/wos')
full_query_file = os.path.join(write_dir, 'with-constraints.txt')
with open(full_query_file, 'w') as f:
for line in all_titles:
f.write(line+'\n')
# -
# # 3. SCOPUS
#
# Shit search
# # 4. IEEE
# +
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
import re
import time
from IPython.display import display
import os
base_mimic_query = '(mimic-ii OR mimic-iii OR mimicii OR mimiciii OR mimic-2 OR mimic-3 OR mimic2 OR mimic3)'
restriction_query = '(physionet OR icu OR “intensive care” OR “critical care”)'
def full_query(base_query, restriction_query):
return ' AND '.join([base_query, restriction_query])
full_mimic_query = full_query(base_mimic_query, restriction_query)
base_search_url = 'http://ieeexplore.ieee.org/search/advsearch.jsp?expression-builder'
all_titles = []
# +
# Get to the search page
driver = webdriver.Firefox()
driver.get(base_search_url)
# Input the query string
searchbox = driver.find_element_by_id('expression-textarea')
searchbox.send_keys(full_mimic_query)
# Select the 'full text and metadata' box
radiobutton = driver.find_element_by_id('Search_All_Text')
radiobutton.click()
# Search
time.sleep(1)
searchbutton = driver.find_element_by_class_name('stats-Adv_Command_search')
searchbutton.click()
# Get the total number of pages
#npages = int(driver.find_element_by_id('pageCount.top').text)
# +
# Get the titles!!!
while True:
# let the page load
time.sleep(2)
# Get scroll height
last_height = driver.execute_script("return document.body.scrollHeight")
while True:
driver.execute_script("window.scrollTo(0, document.body.scrollHeight);")
time.sleep(0.5)
new_height = driver.execute_script("return document.body.scrollHeight")
if new_height == last_height:
break
last_height = new_height
# Get the titles.
# They are in: <h2 class="result-item-title"><a class="ng-binding ng-scope">title</a></h2>
elements = driver.find_elements_by_class_name('result-item-title')
for e in elements:
# Text may appear with "[::sometext::]"
all_titles.append(e.find_element_by_tag_name('a').get_attribute('text').replace('[::', '').replace('::]', ''))
# New line separated journal info and such
#all_titles.append(e.text.split('\n')[0])
# Click next page if any
e = driver.find_element_by_class_name('next')
if 'disabled' in e.get_attribute('class'):
print('Got all paper titles!')
driver.close()
break
else:
nextbutton = driver.find_element_by_link_text('>')
nextbutton.click()
all_titles = set(all_titles)
all_titles = [t.lower() for t in list(all_titles)]
# -
print(len(all_titles))
display(all_titles)
# +
# Write the titles to files
write_dir = os.path.join('search_results/ieee')
full_query_file = os.path.join(write_dir, 'with-constraints.txt')
with open(full_query_file, 'w') as f:
for line in all_titles:
f.write(line+'\n')
# -
# # Combining Results - pubmed, wos, ieee
# +
result_dir = 'search_results'
combined_results = []
for service in ['pubmed', 'wos', 'ieee']:
# For pubmed, get the curated true positives from the unconstrained search instead
if service == 'pubmed':
df = pd.read_csv(os.path.join(result_dir, service, 'without-constraints-inspected.tsv'), delimiter='\t', header=None)
service_results = list(df.loc[df[1]=='T'][0].values)
# For other services, get the constrained search results
else:
with open(os.path.join(result_dir, service, 'with-constraints.txt')) as f:
service_results = f.readlines()
print('Number of results from service '+service+': '+str(len(service_results)))
combined_results = combined_results + [r.strip() for r in service_results]
print('\nTotal number of non-unique results: ', len(combined_results))
combined_results = sorted(list(set(combined_results)))
print('Total number of unique results: ', len(combined_results))
with open(os.path.join(result_dir, 'combined', 'with-constraints.txt'), 'w') as f:
for r in combined_results:
f.write(r+'\n')
# +
# may 21 2018
Number of results from service pubmed: 155
Number of results from service wos: 152
Number of results from service ieee: 322
Total number of non-unique results: 629
Total number of unique results: 456
# -
# # Attempting to parse GS automatically failed. Below is evidence of failure. Can ignore...
# # N. Search Google Scholar
#
# Packages found online:
# - https://github.com/ckreibich/scholar.py
# - https://github.com/venthur/gscholar
# - https://github.com/adeel/google-scholar-scraper
# - http://code.activestate.com/recipes/523047-search-google-scholar/
# - https://github.com/erdiaker/torrequest
# - https://github.com/NikolaiT/GoogleScraper
#
#
# - https://stackoverflow.com/questions/8049520/web-scraping-javascript-page-with-python
#
#
# Query: `("mimic ii" OR "mimic iii") AND ("database" OR "clinical" OR "waveform" OR ICU)`
#
# https://scholar.google.com/scholar?q=%28mimic-ii+OR+mimic-iii%29&btnG=&hl=en&as_sdt=1%2C22&as_vis=1
#
# https://scholar.google.com/scholar/help.html
#
#
# https://superuser.com/questions/565722/how-to-config-tor-to-use-a-http-socks-proxy
#
# ## Requirements
#
# 1. Browse with JS enabled. requests library uses http. Otherwise google will think (correctly) that you are a robot.
# 2. Change IP every time, or google will block.
# +
#from torrequest import TorRequest
from bs4 import BeautifulSoup
import urllib2
import getpass
import sys
import stem
import stem.connection
from stem.control import Controller
# -
# Show IP address
with TorRequest(proxy_port=9050, ctrl_port=9051, password=None) as tr:
response = tr.get('http://ipecho.net/plain')
print(response.text)
tr.reset_identity
# Show IP address
with TorRequest(proxy_port=9050, ctrl_port=9051, password=None) as tr:
response = tr.get('http://ipecho.net/plain')
print(response.text)
tr.reset_identity
# +
with TorRequest(proxy_port=9050, ctrl_port=9051, password=None) as tr:
# Specify HTTP verb and url.
resp = tr.get('https://scholar.google.com/scholar?q=%28mimic-ii+OR+mimic-iii%29&hl=en&as_sdt=1%2C22&as_vis=1&as_ylo=2017&as_yhi=2017')
print(resp.text)
# Change your Tor circuit,
# and likely your observed IP address.
tr.reset_identity()
# -
type(resp.text)
soup = BeautifulSoup(resp.text,'html.parser')
for anchor in soup.find_all('a'):
print(anchor.get('href', '/'))
# +
from bs4 import BeautifulSoup
import urllib2
webpage = urllib2.urlopen('http://en.wikipedia.org/wiki/Main_Page')
soup = BeautifulSoup(webpage,'html.parser')
for anchor in soup.find_all('a'):
print(anchor.get('href', '/'))
# -
with TorRequest() as tr:
response = tr.get('http://ipecho.net/plain')
print(response.text) # not your IP address
# +
with TorRequest(proxy_port=9050, ctrl_port=9051, password=<PASSWORD>) as tr:
# Specify HTTP verb and url.
resp = tr.get('https://scholar.google.com/scholar?q=%28mimic-ii+OR+mimic-iii%29&hl=en&as_sdt=1%2C22&as_vis=1&as_ylo=2017&as_yhi=2017')
print(resp.text)
# # Send data. Use basic authentication.
# resp = tr.post('https://api.example.com',
# data={'foo': 'bar'}, auth=('user', 'pass'))'
# print(resp.json)
# Change your Tor circuit,
# and likely your observed IP address.
tr.reset_identity()
# TorRequest object also exposes the underlying Stem controller
# and Requests session objects for more flexibility.
print(type(tr.ctrl)) # a stem.control.Controller object
tr.ctrl.signal('CLEARDNSCACHE') # see Stem docs for the full API
print(type(tr.session)) # a requests.Session object
c = cookielib.CookieJar()
tr.session.cookies.update(c) # see Requests docs for the full API
# -
scholar_url = 'https://scholar.google.com/scholar?as_vis=1&q=sepsis+mimic-iii&hl=en&as_sdt=1,22'
echo_ip_url = 'https://www.atagar.com/echo.php'
test_js_url = 'http://127.0.0.1:81/test-js.html'
# +
with TorRequest(proxy_port=9050, ctrl_port=9051, password=<PASSWORD>) as tr:
# Specify HTTP verb and url.
resp = tr.get('https://scholar.google.com/scholar?q=%28mimic-ii+OR+mimic-iii%29&hl=en&as_sdt=1%2C22&as_vis=1&as_ylo=2017&as_yhi=2017')
print(resp.text)
# # Send data. Use basic authentication.
# resp = tr.post('https://api.example.com',
# data={'foo': 'bar'}, auth=('user', 'pass'))'
# print(resp.json)
# Change your Tor circuit,
# and likely your observed IP address.
tr.reset_identity()
# -
# +
import io
import pycurl
import stem.process
from stem.util import term
SOCKS_PORT = 9000
def query(url):
"""
Uses pycurl to fetch a site using the proxy on the SOCKS_PORT.
"""
output = io.BytesIO()
query = pycurl.Curl()
query.setopt(pycurl.URL, url)
query.setopt(pycurl.PROXY, 'localhost')
query.setopt(pycurl.PROXYPORT, SOCKS_PORT)
query.setopt(pycurl.PROXYTYPE, pycurl.PROXYTYPE_SOCKS5_HOSTNAME)
query.setopt(pycurl.WRITEFUNCTION, output.write)
try:
query.perform()
return output.getvalue()
except pycurl.error as exc:
return "Unable to reach %s (%s)" % (url, exc)
# Start an instance of Tor configured to only exit through Russia. This prints
# Tor's bootstrap information as it starts. Note that this likely will not
# work if you have another Tor instance running.
def print_bootstrap_lines(line):
if "Bootstrapped " in line:
print(term.format(line, term.Color.BLUE))
print(term.format("Starting Tor:\n", term.Attr.BOLD))
tor_process = stem.process.launch_tor_with_config(
config = {
'SocksPort': str(SOCKS_PORT),
'ExitNodes': '{ru}',
},
init_msg_handler = print_bootstrap_lines,
)
print(term.format("\nChecking our endpoint:\n", term.Attr.BOLD))
print(term.format(query("https://www.atagar.com/echo.php"), term.Color.BLUE))
tor_process.kill() # stops tor
# -
q = query("https://www.atagar.com/echo.php")
import dryscrape
s = dryscrape.Session()
s.set_proxy(port=9050)
# +
import stem
from stem.control import Controller
from stem.process import launch_tor_with_config
import requests
import dryscrape
import time
class TorRequest(object):
def __init__(self,
proxy_port=9050,
ctrl_port=9051,
password=<PASSWORD>):
self.proxy_port = proxy_port
self.ctrl_port = ctrl_port
self._tor_proc = None
if not self._tor_process_exists():
self._tor_proc = self._launch_tor()
self.ctrl = Controller.from_port(port=self.ctrl_port)
self.ctrl.authenticate(password=password)
self.session = requests.Session()
self.session.proxies.update({
'http': 'socks5://localhost:%d' % self.proxy_port,
'https:': 'socks5://localhost:%d' % self.proxy_port,
})
def _tor_process_exists(self):
try:
ctrl = Controller.from_port(port=self.ctrl_port)
ctrl.close()
return True
except:
return False
def _launch_tor(self):
return launch_tor_with_config(
config={
'SocksPort': str(self.proxy_port),
'ControlPort': str(self.ctrl_port)
},
take_ownership=True)
def close(self):
try:
self.session.close()
except: pass
try:
self.ctrl.close()
except: pass
if self._tor_proc:
self._tor_proc.terminate()
def reset_identity_async(self):
self.ctrl.signal(stem.Signal.NEWNYM)
def reset_identity(self):
self.reset_identity_async()
time.sleep(self.ctrl.get_newnym_wait())
def get(self, *args, **kwargs):
return self.session.get(*args, **kwargs)
def post(self, *args, **kwargs):
return self.session.post(*args, **kwargs)
def put(self, *args, **kwargs):
return self.session.put(*args, **kwargs)
def patch(self, *args, **kwargs):
return self.session.patch(*args, **kwargs)
def delete(self, *args, **kwargs):
return self.session.delete(*args, **kwargs)
def __enter__(self):
return self
def __exit__(self, *args):
self.close()
# -
# Show IP address
with TorRequest(proxy_port=9050, ctrl_port=9051, password='16:<PASSWORD>') as tr:
response = tr.get('http://ipecho.net/plain')
print(response.text)
tr.reset_identity
# +
import dryscrape
import sys
from bs4 import BeautifulSoup
import time
scholar_url = 'https://scholar.google.com/scholar?as_vis=1&q=sepsis+mimic-iii&hl=en&as_sdt=1,22'
echo_ip_url = 'http://ipecho.net/plain'
test_js_url = 'http://1172.16.31.10:81/test-js.html'
if 'linux' in sys.platform:
# start xvfb in case no X is running. Make sure xvfb
# is installed, otherwise this won't work!
dryscrape.start_xvfb()
# +
s = dryscrape.Session()
s.visit(test_js_url)
s.body()
#s.visit('https://scholar.google.com/scholar?as_vis=1&q=sepsis+mimic-iii&hl=en&as_sdt=1,22')
# waiting for the first data row in a table to be present
# s.wait_for(lambda: s.at_css("tr.data-row0"))
# soup = BeautifulSoup(s.body(), 'lxml')
# +
s = dryscrape.Session()
s.set_proxy(host = "localhost", port = 8118)
#time.sleep(20)
s.visit(echo_ip_url)
#s.body()
# -
s.body()
| extract-citations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modeling and Simulation in Python
#
# Chapter 9
#
# Copyright 2017 <NAME>
#
# License: [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0)
#
# +
# Configure Jupyter to display the assigned value after an assignment
# %config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# import everything from SymPy.
from sympy import *
# Set up Jupyter notebook to display math.
init_printing()
# -
# The following displays SymPy expressions and provides the option of showing results in LaTeX format.
# +
from sympy.printing import latex
def show(expr, show_latex=False):
"""Display a SymPy expression.
expr: SymPy expression
show_latex: boolean
"""
if show_latex:
print(latex(expr))
return expr
# -
# ### Analysis with SymPy
# Create a symbol for time.
t = symbols('t')
s = symbols('s')
# If you combine symbols and numbers, you get symbolic expressions.
expr = t + 1
# The result is an `Add` object, which just represents the sum without trying to compute it.
type(expr)
# `subs` can be used to replace a symbol with a number, which allows the addition to proceed.
expr.subs(t, 2)
# `f` is a special class of symbol that represents a function.
f = Function('f')
# The type of `f` is `UndefinedFunction`
type(f)
# SymPy understands that `f(t)` means `f` evaluated at `t`, but it doesn't try to evaluate it yet.
f(t)
# `diff` returns a `Derivative` object that represents the time derivative of `f`
dfdt = diff(f(t), t)
type(dfdt)
# We need a symbol for `alpha`
alpha = symbols('alpha')
# Now we can write the differential equation for proportional growth.
eq1 = Eq(dfdt, alpha*f(t))
# And use `dsolve` to solve it. The result is the general solution.
solution_eq = dsolve(eq1)
# We can tell it's a general solution because it contains an unspecified constant, `C1`.
#
# In this example, finding the particular solution is easy: we just replace `C1` with `p_0`
C1, p_0 = symbols('C1 p_0')
particular = solution_eq.subs(C1, p_0)
# In the next example, we have to work a little harder to find the particular solution.
# ### Solving the quadratic growth equation
#
# We'll use the (r, K) parameterization, so we'll need two more symbols:
r, K = symbols('r K')
# Now we can write the differential equation.
eq2 = Eq(diff(f(t), t), r * f(t) * (1 - f(t)/K))
# And solve it.
solution_eq = dsolve(eq2)
# The result, `solution_eq`, contains `rhs`, which is the right-hand side of the solution.
general = solution_eq.rhs
# We can evaluate the right-hand side at $t=0$
at_0 = general.subs(t, 0)
# Now we want to find the value of `C1` that makes `f(0) = p_0`.
#
# So we'll create the equation `at_0 = p_0` and solve for `C1`. Because this is just an algebraic identity, not a differential equation, we use `solve`, not `dsolve`.
#
# The result from `solve` is a list of solutions. In this case, [we have reason to expect only one solution](https://en.wikipedia.org/wiki/Picard%E2%80%93Lindel%C3%B6f_theorem), but we still get a list, so we have to use the bracket operator, `[0]`, to select the first one.
solutions = solve(Eq(at_0, p_0), C1)
type(solutions), len(solutions)
value_of_C1 = solutions[0]
# Now in the general solution, we want to replace `C1` with the value of `C1` we just figured out.
particular = general.subs(C1, value_of_C1)
# The result is complicated, but SymPy provides a method that tries to simplify it.
particular = simplify(particular)
# Often simplicity is in the eye of the beholder, but that's about as simple as this expression gets.
#
# Just to double-check, we can evaluate it at `t=0` and confirm that we get `p_0`
particular.subs(t, 0)
# This solution is called the [logistic function](https://en.wikipedia.org/wiki/Population_growth#Logistic_equation).
#
# In some places you'll see it written in a different form:
#
# $f(t) = \frac{K}{1 + A e^{-rt}}$
#
# where $A = (K - p_0) / p_0$.
#
# We can use SymPy to confirm that these two forms are equivalent. First we represent the alternative version of the logistic function:
A = (K - p_0) / p_0
logistic = K / (1 + A * exp(-r*t))
# To see whether two expressions are equivalent, we can check whether their difference simplifies to 0.
simplify(particular - logistic)
# This test only works one way: if SymPy says the difference reduces to 0, the expressions are definitely equivalent (and not just numerically close).
#
# But if SymPy can't find a way to simplify the result to 0, that doesn't necessarily mean there isn't one. Testing whether two expressions are equivalent is a surprisingly hard problem; in fact, there is no algorithm that can solve it in general.
# ### Exercises
#
# **Exercise:** Solve the quadratic growth equation using the alternative parameterization
#
# $\frac{df(t)}{dt} = \alpha f(t) + \beta f^2(t) $
eq3 = Eq(dfdt, alpha*f(t) + beta*f(t)**2)
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# -
# **Exercise:** Use [WolframAlpha](https://www.wolframalpha.com/) to solve the quadratic growth model, using either or both forms of parameterization:
#
# df(t) / dt = alpha f(t) + beta f(t)^2
#
# or
#
# df(t) / dt = r f(t) (1 - f(t)/K)
#
# Find the general solution and also the particular solution where `f(0) = p_0`.
| code/chap09sympy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!DOCTYPE html>
# <html>
# <body>
# <div align="center">
# <h1>Road to AI Session 2</h1>
#
# <h3>Database,Graphs & Maths</h3>
# <h4>PANDAS</h4>
# </div>
# </body>
# </html>
# ## Importing the library
import numpy as np
import pandas as pd
print(pd.__version__)
# ## Reading the file
df = pd.read_csv('apy.csv')
# ## The Basics
df.head()
df.tail()
#We can see the dimensions of the dataframe using the the shape attribute
df.shape
#We can also extract all the column names as a list
df.columns.tolist()
#function to see statistics like mean, min, etc about each column of the dataset
df.describe()
## max() will show you the maximum values of all columns
df.max()
#get the max value for a particular column
df['Area'].max()
## find the mean of the Production score.
df['Production'].mean()
## function to identify the row index
df['Production'].argmax()
# **value_counts()** shows how many times each item appears in the column. This particular command shows the number of games in each season
df['Area'].value_counts()
# ## Acessing Values
## get attributes
df.iloc[[df['Production'].argmax()]]
df.iloc[[df['Production'].argmax()]]['Area']
# When you see data displayed in the above format, you're dealing with a Pandas **Series** object, not a dataframe object.
type(df.iloc[[df['Production'].argmax()]]['Area'])
type(df.iloc[[df['Production'].argmax()]])
# The other really important function in Pandas is the **loc** function. Contrary to iloc, which is an integer based indexing, loc is a "Purely label-location based indexer for selection by label". Since the table are ordered from 0 to 145288, iloc and loc are going to be pretty interchangable in this type of dataset
df.iloc[:3]
## loc is a "Purely label-location based indexer for selection by label"
df.loc[:3]
# Notice the slight difference in that iloc is exclusive of the second number, while loc is inclusive.
# Below is an example of how you can use loc to acheive the same task as we did previously with iloc
df.loc[df['Production'].argmax(), 'Area']
# A faster version uses the **at()** function. At() is really useful wheneever you know the row label and the column label of the particular value that you want to get.
df.at[df['Production'].argmax(), 'Area']
# # Sorting
## sort the dataframe in increasing order
df.sort_values('Area').head()
df.groupby('Area')
# # Filtering Rows Conditionally
df[df['Area'] > 500000]
df[(df['Area'] > 5000000) & (df['Area'] < 5555500)]
# # Grouping
## allows you to group entries by certain attributes
df.groupby('State_Name')['Area'].mean().head()
df.groupby('State_Name')['Area'].value_counts().head(9)
df.values
## Now, you can simply just access elements like you would in an array.
df.values[0][0]
# # Extracting Rows and Columns
# The bracket indexing operator is one way to extract certain columns from a dataframe.
df[['Production', 'Area']].head()
# Notice that you can acheive the same result by using the loc function. Loc is a veryyyy versatile function that can help you in a lot of accessing and extracting tasks.
df.loc[:, ['Production', 'Area']].head()
# Note the difference is the return types when you use brackets and when you use double brackets.
type(df['Production'])
type(df[['Production']])
# You've seen before that you can access columns through df['col name']. You can access rows by using slicing operations.
df[0:3]
# Here's an equivalent using iloc
df.iloc[0:3,:]
# # Data Cleaning
## if there are any missing values in the dataframe, and will then sum up the total for each column
df.isnull().sum()
# If you do end up having missing values in your datasets, be sure to get familiar with these two functions.
# * **dropna()** - This function allows you to drop all(or some) of the rows that have missing values.
# * **fillna()** - This function allows you replace the rows that have missing values with the value that you pass in.
# # Other Useful Functions
# * **drop()** - This function removes the column or row that you pass in (You also have the specify the axis).
# * **agg()** - The aggregate function lets you compute summary statistics about each group
# * **apply()** - Lets you apply a specific function to any/all elements in a Dataframe or Series
# * **get_dummies()** - Helpful for turning categorical data into one hot vectors.
# * **drop_duplicates()** - Lets you remove identical rows
# <!DOCTYPE html>
# <html>
# <body>
# <div align="center">
# <h1>THE END</h1>
#
# <h3>By <NAME> & <NAME></h3>
# </div>
# </body>
# </html>
| pandas/notebooks/pandas_classroom.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="http://cocl.us/pytorch_link_top">
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/Pytochtop.png" width="750" alt="IBM Product " />
# </a>
#
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/cc-logo-square.png" width="200" alt="cognitiveclass.ai logo" />
# <h1>Batch Normalization with the MNIST Dataset</h1>
# <h2>Table of Contents</h2>
# In this lab, you will build a Neural Network using Batch Normalization and compare it to a Neural Network that does not use Batch Normalization. You will use the MNIST dataset to test your network.
#
# <ul>
# <li><a href="#Train_Func">Neural Network Module and Training Function</a></li>
# <li><a href="#Makeup_Data">Load Data </a></li>
# <li><a href="#NN">Define Several Neural Networks, Criterion function, Optimizer</a></li>
# <li><a href="#Train">Train Neural Network using Batch Normalization and no Batch Normalization</a></li>
# <li><a href="#Result">Analyze Results</a></li>
# </ul>
# <p>Estimated Time Needed: <strong>25 min</strong></p>
# </div>
#
# <hr>
# <h2>Preparation</h2>
# We'll need the following libraries:
# +
# These are the libraries will be used for this lab.
# Using the following line code to install the torchvision library
# # !conda install -y torchvision
import torch
import torch.nn as nn
import torchvision.transforms as transforms
import torchvision.datasets as dsets
import torch.nn.functional as F
import matplotlib.pylab as plt
import numpy as np
torch.manual_seed(0)
# -
# <!--Empty Space for separating topics-->
# <h2 id="Train_Func">Neural Network Module and Training Function</h2>
# Define the neural network module or class
# Neural Network Module with two hidden layers using Batch Normalization
# +
# Define the Neural Network Model using Batch Normalization
class NetBatchNorm(nn.Module):
# Constructor
def __init__(self, in_size, n_hidden1, n_hidden2, out_size):
super(NetBatchNorm, self).__init__()
self.linear1 = nn.Linear(in_size, n_hidden1)
self.linear2 = nn.Linear(n_hidden1, n_hidden2)
self.linear3 = nn.Linear(n_hidden2, out_size)
self.bn1 = nn.BatchNorm1d(n_hidden1)
self.bn2 = nn.BatchNorm1d(n_hidden2)
# Prediction
def forward(self, x):
x = self.bn1(torch.sigmoid(self.linear1(x)))
x = self.bn2(torch.sigmoid(self.linear2(x)))
x = self.linear3(x)
return x
# Activations, to analyze results
def activation(self, x):
out = []
z1 = self.bn1(self.linear1(x))
out.append(z1.detach().numpy().reshape(-1))
a1 = torch.sigmoid(z1)
out.append(a1.detach().numpy().reshape(-1).reshape(-1))
z2 = self.bn2(self.linear2(a1))
out.append(z2.detach().numpy().reshape(-1))
a2 = torch.sigmoid(z2)
out.append(a2.detach().numpy().reshape(-1))
return out
# -
# Neural Network Module with two hidden layers with out Batch Normalization
# +
# Class Net for Neural Network Model
class Net(nn.Module):
# Constructor
def __init__(self, in_size, n_hidden1, n_hidden2, out_size):
super(Net, self).__init__()
self.linear1 = nn.Linear(in_size, n_hidden1)
self.linear2 = nn.Linear(n_hidden1, n_hidden2)
self.linear3 = nn.Linear(n_hidden2, out_size)
# Prediction
def forward(self, x):
x = torch.sigmoid(self.linear1(x))
x = torch.sigmoid(self.linear2(x))
x = self.linear3(x)
return x
# Activations, to analyze results
def activation(self, x):
out = []
z1 = self.linear1(x)
out.append(z1.detach().numpy().reshape(-1))
a1 = torch.sigmoid(z1)
out.append(a1.detach().numpy().reshape(-1).reshape(-1))
z2 = self.linear2(a1)
out.append(z2.detach().numpy().reshape(-1))
a2 = torch.sigmoid(z2)
out.append(a2.detach().numpy().reshape(-1))
return out
# -
# Define a function to train the model. In this case the function returns a Python dictionary to store the training loss and accuracy on the validation data
# +
# Define the function to train model
def train(model, criterion, train_loader, validation_loader, optimizer, epochs=100):
i = 0
useful_stuff = {'training_loss':[], 'validation_accuracy':[]}
for epoch in range(epochs):
for i, (x, y) in enumerate(train_loader):
model.train()
optimizer.zero_grad()
z = model(x.view(-1, 28 * 28))
loss = criterion(z, y)
loss.backward()
optimizer.step()
useful_stuff['training_loss'].append(loss.data.item())
correct = 0
for x, y in validation_loader:
model.eval()
yhat = model(x.view(-1, 28 * 28))
_, label = torch.max(yhat, 1)
correct += (label == y).sum().item()
accuracy = 100 * (correct / len(validation_dataset))
useful_stuff['validation_accuracy'].append(accuracy)
return useful_stuff
# -
# <!--Empty Space for separating topics-->
# <h2 id="Makeup_Data">Make Some Data</h2>
# Load the training dataset by setting the parameters <code>train </code> to <code>True</code> and convert it to a tensor by placing a transform object int the argument <code>transform</code>
# +
# load the train dataset
train_dataset = dsets.MNIST(root='./data', train=True, download=True, transform=transforms.ToTensor())
# -
# Load the validating dataset by setting the parameters train <code>False</code> and convert it to a tensor by placing a transform object into the argument <code>transform</code>
# +
# load the train dataset
validation_dataset = dsets.MNIST(root='./data', train=False, download=True, transform=transforms.ToTensor())
# -
# create the training-data loader and the validation-data loader object
# +
# Create Data Loader for both train and validating
train_loader = torch.utils.data.DataLoader(dataset=train_dataset, batch_size=2000, shuffle=True)
validation_loader = torch.utils.data.DataLoader(dataset=validation_dataset, batch_size=5000, shuffle=False)
# -
# <a id="ref3"></a>
# <h2 align=center>Define Neural Network, Criterion function, Optimizer and Train the Model </h2>
# Create the criterion function
# +
# Create the criterion function
criterion = nn.CrossEntropyLoss()
# -
# Variables for Neural Network Shape <code> hidden_dim</code> used for number of neurons in both hidden layers.
# +
# Set the parameters
input_dim = 28 * 28
hidden_dim = 100
output_dim = 10
# -
# <!--Empty Space for separating topics-->
# <h2 id="Train">Train Neural Network using Batch Normalization and no Batch Normalization </h2>
# Train Neural Network using Batch Normalization :
# +
# Create model, optimizer and train the model
model_norm = NetBatchNorm(input_dim, hidden_dim, hidden_dim, output_dim)
optimizer = torch.optim.Adam(model_norm.parameters(), lr = 0.1)
training_results_Norm=train(model_norm , criterion, train_loader, validation_loader, optimizer, epochs=5)
# -
# Train Neural Network with no Batch Normalization:
# +
# Create model without Batch Normalization, optimizer and train the model
model = Net(input_dim, hidden_dim, hidden_dim, output_dim)
optimizer = torch.optim.Adam(model.parameters(), lr = 0.1)
training_results = train(model, criterion, train_loader, validation_loader, optimizer, epochs=5)
# -
# <h2 id="Result">Analyze Results</h2>
# Compare the histograms of the activation for the first layer of the first sample, for both models.
model.eval()
model_norm.eval()
out=model.activation(validation_dataset[0][0].reshape(-1,28*28))
plt.hist(out[2],label='model with no batch normalization' )
out_norm=model_norm.activation(validation_dataset[0][0].reshape(-1,28*28))
plt.hist(out_norm[2],label='model with normalization')
plt.xlabel("activation ")
plt.legend()
plt.show()
# <!--Empty Space for separating topics-->
# We see the activations with Batch Normalization are zero centred and have a smaller variance.
# Compare the training loss for each iteration
# +
# Plot the diagram to show the loss
plt.plot(training_results['training_loss'], label='No Batch Normalization')
plt.plot(training_results_Norm['training_loss'], label='Batch Normalization')
plt.ylabel('Cost')
plt.xlabel('iterations ')
plt.legend()
plt.show()
# -
# Compare the validating accuracy for each iteration
# +
# Plot the diagram to show the accuracy
plt.plot(training_results['validation_accuracy'],label='No Batch Normalization')
plt.plot(training_results_Norm['validation_accuracy'],label='Batch Normalization')
plt.ylabel('validation accuracy')
plt.xlabel('epochs ')
plt.legend()
plt.show()
# -
# <!--Empty Space for separating topics-->
# <a href="http://cocl.us/pytorch_link_bottom">
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/notebook_bottom%20.png" width="750" alt="PyTorch Bottom" />
# </a>
# <h2>About the Authors:</h2>
#
# <a href="https://www.linkedin.com/in/joseph-s-50398b136/"><NAME></a> has a PhD in Electrical Engineering, his research focused on using machine learning, signal processing, and computer vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.
# Other contributors: <a href="https://www.linkedin.com/in/michelleccarey/"><NAME></a>, <a href="www.linkedin.com/in/jiahui-mavis-zhou-a4537814a"><NAME></a>
# <hr>
# Copyright © 2018 <a href="cognitiveclass.ai?utm_source=bducopyrightlink&utm_medium=dswb&utm_campaign=bdu">cognitiveclass.ai</a>. This notebook and its source code are released under the terms of the <a href="https://bigdatauniversity.com/mit-license/">MIT License</a>.
| IBM_AI/4_Pytorch/8.5.1BachNorm_v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
#Minecraft Turtle Example - Crazy Pattern
from mcpiext import minecraftturtle
from mcpi import minecraft, block
import random
#create connection to minecraft
mc = minecraft.Minecraft.create()
#get players position
pos = mc.player.getPos()
#create minecraft turtle
steve = minecraftturtle.MinecraftTurtle(mc, pos)
steve.penblock(block.WOOL.id, 11)
steve.speed(10)
for step in range(0,50):
steve.forward(50)
steve.right(123)
| classroom-code/turtle-examples/pattern.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.8 64-bit (''base'': conda)'
# name: python3
# ---
# # Assignment 4 (Oct 12)
#
# Today we will talk about **Monte Carlo (MC)** methods, techniques that involve random sampling. But before we dive into that,
# we will need to learn a little bit about random numbers in Python.
#
# We will go over:
#
# 1. Python Numpy Random Generator: `np.random.default_rng()`
# 2. Plot histogram
# 3. Monte Carlo -- Uncertainty Propagation
# 4. Monte Carlo -- Bootstrapping
#
# ## Readings (optional)
#
# If you find this week's material new or challenging, you may want to read through some or all the following resources while working on your assignment:
#
# - [SPIRL Ch. 3.6. Conditionals](https://cjtu.github.io/spirl/python_conditionals.html#conditionals)
# - [Numpy Random Generator](https://numpy.org/doc/stable/reference/random/generator.html)
#
import numpy as np
import matplotlib.pyplot as plt
# ## Random Number Generators (RNGs)
#
# To work with random numbers in Python, we can use the `random` module within `numpy`.
#
# ### Pseudo-random number generators (PRNG)
#
# It might sound easy to come up with a random number, but many surveys have found this to not be the case (e.g. [Asking 8500 students to pick a random number](https://www.reddit.com/r/dataisbeautiful/comments/acow6y/asking_over_8500_students_to_pick_a_random_number/)).
#
# Because computers are deterministic (a collection of switches that can be "on" or "off"), it is surprisingly hard to produce a set of numbers that are truly random. Luckily, we often don't need a *truly* random set of numbers. Usually when we say we want random numbers, we want a set of numbers that are:
#
# - not biased to any particular value
# - contain no repeated or recognizable patterns
#
# This is where a **pseudorandom number generator (PRNG)** can help.
#
# See what Wikipedia tell us about the PRNG:
#
# > A pseudorandom number generator (PRNG), also known as a deterministic random bit generator (DRBG), is an algorithm for generating a sequence of numbers whose properties approximate the properties of sequences of random numbers. The PRNG-generated sequence is not truly random, because it is completely determined by an initial value, called the PRNG's seed (which may include truly random values).
#
# The key "flaw" with a PRNG is that if you know a special value called the **seed**, you can regenerate the exact same sequence of random numbers again. But this ends up being a useful *feature* of PRNGs as we'll see later.
#
# Since all computer generated random number generators are PRNGs, we often just drop the "P" and simply call them **random number generators (RNGs)** (but now you know their secret).
#
# Read more about [Pseudorandom number generators on Wikipedia](https://en.wikipedia.org/wiki/Pseudorandom_number_generator).
#
# ### NumPy Random Module (`np.random`)
#
# In NumPy, there are two ways to generate random numbers:
#
# - Calling functions in `random` directly (**deprecated**): `np.random.func()`
# - Generating an `rng` object with `obj = np.random.default_rng()` and calling methods on it: `obj.method()`
#
# There are also different algorithms you can use to generate random numbers and if you mix and match RNG algorithms, you won't be guaranteed the same random numbers even if you know the **seed**. This is why random numbers in different programming languages won't necessarily be the same with the same seed (read more about [NumPy bit generators](https://numpy.org/doc/stable/reference/random/bit_generators/index.html)). For almost all applications, the `default_random` from NumPy is sufficient (see [NumPy simple random data](https://numpy.org/doc/stable/reference/random/generator.html#simple-random-data). Let's try it out!
# Initialize the random number generator object
rng = np.random.default_rng()
# help(rng.integers)
# Let's first try to generate random intergers with the `rng.integers()` function:
#
# ```
# integers(low, high=None, size=None, dtype=np.int64, endpoint=False)
#
# Return random integers from `low` (inclusive) to `high` (exclusive), or
# if endpoint=True, `low` (inclusive) to `high` (inclusive). Replaces
# `RandomState.randint` (with endpoint=False) and
# `RandomState.random_integers` (with endpoint=True)
#
# Return random integers from the "discrete uniform" distribution of
# the specified dtype. If `high` is None (the default), then results are
# from 0 to `low`.
# ...
# ```
# +
# Draw random intergers from the range `draw_range` `ndraws` times.
draw_range = (0, 10) # (low, high)
ndraws = 8 # how many to generate or "draw"
random_ints = rng.integers(*draw_range, ndraws)
print(random_ints)
# -
# Now use `for` loop to run it many times to see if any are duplicated:
rng = np.random.default_rng()
for i in range(10):
random_ints = rng.integers(*draw_range, ndraws)
print(f'Run {i}: ', random_ints)
# But remember: these are *pseudo*random numbers, meaning we can generate the same random sequence again if we know the **seed** value.
#
# This time, see what happens when we re-make the rng object with the same seed each time in the loop.
# +
draw_range = (0, 10)
ndraws = 10
seed = 100
for i in range(10):
rng = np.random.default_rng(seed=seed) # seed the default RNG
random_ints = rng.integers(*draw_range, ndraws)
print(f'run {i} give: ', random_ints)
# -
# ## Plot histogram with `plt.hist`
#
# We can verify how random our values are using a histogram.
#
# `help(plt.hist)`
#
# ```
# hist(x, bins=None, range=None, density=None, weights=None, cumulative=False, bottom=None, histtype='bar', align='mid', orientation='vertical', rwidth=None, log=False, color=None, label=None, stacked=False, normed=None, *, data=None, **kwargs)
# Plot a histogram.
# ...
# Returns
# -------
# n : array or list of arrays
# The values of the histogram bins. See *density* and *weights* for a
# description of the possible semantics. If input *x* is an array,
# then this is an array of length *nbins*. If input is a sequence of
# arrays ``[data1, data2,..]``, then this is a list of arrays with
# the values of the histograms for each of the arrays in the same
# order. The dtype of the array *n* (or of its element arrays) will
# always be float even if no weighting or normalization is used.
#
# bins : array
# The edges of the bins. Length nbins + 1 (nbins left edges and right
# edge of last bin). Always a single array even when multiple data
# sets are passed in.
#
# patches : list or list of lists
# Silent list of individual patches used to create the histogram
# or list of such list if multiple input datasets.
# ...
# ```
#
# We will also use a convenient helper function to take care of some of our plot formatting.
# If we want to apply the same format to each plot, we can make it a function!
def set_plot_axis_label(ax, xlabel, ylabel):
"""
Set formatting options on a matplotlib ax object.
Parameters
----------
ax : matplotlib.axes.Axes
The ax object to format.
xlabel : str
The x-axis label.
ylabel : str
The y-axis label.
"""
ax.tick_params(axis='both', which ='both', labelsize='small', right=True,
top=True, direction='in')
ax.set_xlabel(xlabel, size='medium', fontname='Helvetica')
ax.set_ylabel(ylabel, size='medium', fontname='Helvetica')
# +
# random number setup
draw_range = (0, 10)
ndraws = 10000 # Try doing different numbers of draws
# Do random draws
rng = np.random.default_rng()
random_ints = rng.integers(*draw_range, ndraws)
# Set up plot and plot the histogram
fig, axs = plt.subplots(1, 2, facecolor='white', figsize=(8, 3), dpi=150)
# Default hist
n, edges, _ = axs[0].hist(random_ints)
set_plot_axis_label(axs[0], 'Value', 'Count') # Our helper function
# Center the bins and show as a "step" function
n, edges, _ = axs[1].hist(random_ints, range=(-0.5, 9.5), bins=10,
histtype='step')
set_plot_axis_label(axs[1], 'Value', 'Count') # Our helper function
# add subticks for both axises
from matplotlib.ticker import AutoMinorLocator
for axi in range(2):
axs[axi].xaxis.set_minor_locator(AutoMinorLocator(2))
axs[axi].yaxis.set_minor_locator(AutoMinorLocator(5))
# -
# Let's see what our histogram actually gave us in the `n` and `edges` it returned.
print(f'Number in each bins: {n}')
print(f'Edges of each bin: {edges}')
# Now, let's overplot these data points back to the histogram.
#
# But, before that, we need to find out the center values of the each bins (we only have the left and right edges currently).
#
# ### [Short Quiz] Find the bin centers from `edges` array
#
# Try to write code to covert bin edges to bin centers (copying the following list is cheating! We want to do it in general).
#
# ```python
# edges = [-0.5 0.5 1.5 2.5 3.5 4.5 5.5 6.5 7.5 8.5 9.5]
# ```
# into
# ```python
# bin_center = [0, 1, 2, 3, 4, 5, 6, 7, 8, 9]
# ```
#
#
# put you code here, you have 5 mins
edges
# +
# Answer 1 -- using the for loop
bin_center = []
for i in range(len(edges)-1):
bin_center.append((edges[i] + edges[i+1]) / 2)
print(bin_center)
# -
# ### [Side note:] List comprehensions
#
# List comprehensions are a fancy way to make a list using a simple 1 line `for` loop.
#
# The basic syntax is square brackets `[]` with the familiar `for i in blah` inside.
#
# For example:
[i for i in range(10)]
# The `i` at the beginning is just our loop variable and indicates what we want Python to be put in the final list. So we can also do math or functions on that loop variable, as we would in a for loop.
# +
squared = [n**2 for n in range(10)]
# This is equivalent to:
squared2 = []
for n in range(10):
squared2.append(n**2)
print(squared)
print(squared2)
print(squared == squared2) # "==" tests for equality
# -
# Getting back to our problem of converting bin edges to bin centers, we can do this with a list comprehension!
#
# **Note:** This is about as complicated as a list comprehension should ever get. It is already a little tricky to read as is which can make bugs harder to spot. When in doubt, you can just use a traditional `for` loop and lay each step out so it's easier to understand later!
# +
# Answer 2 -- using the list comprehension
bin_center = [(edges[i] + edges[i+1]) / 2 for i in range(len(edges)-1)]
print(bin_center)
# -
# Finally, we have a 3rd solution which takes advantage of NumPy array indexing, slicing, and element-wise math. We call this **vectorization** and it is usually the most efficient way to solve a mathematical problem with code. It also often uses less code which can be good, since less code has less room for errors.
#
# Vectorizing code takes a little practice. The main idea is to think about arrays as collections of numbers we can do math on all at once.
# +
# Answer 3 -- array slicing (recommended)
bin_center = (edges[:-1] + edges[1:]) / 2
print(bin_center)
# -
# Let's break that example down to see what we did:
# +
print(edges[:-1]) # All edges except the last one
print(edges[1:]) # All edges except the first one
# Now we have 2 arrays of elements but they are offset by 1
# Now we want to take the avg of these adjacent elements to get the centers
print((edges[:-1] + edges[1:]) / 2) # Mean is just (prev_el + next_el) / 2
# -
# Whichever way you figured out how to compute the bin centers, we can now plot them on our histogram!
# +
# random number setup
draw_range = (0, 10)
ndraws = 1000
rng = np.random.default_rng()
random_ints = rng.integers(*draw_range,
ndraws)
# plotting
f = plt.figure(facecolor='white', figsize=(4,3), dpi=150 )
ax1 = f.subplots(1, 1)
# plot the histogram
n, edges, _ = ax1.hist(random_ints, range = (-0.5, 9.5), bins=10, histtype='step')
bin_center = (edges[:-1] + edges[1:])/2
ax1.plot(bin_center, n, '.', ms=3, c='tab:red')
set_plot_axis_label(ax1, 'Value', 'Count')
ax1.xaxis.set_minor_locator(AutoMinorLocator(2))
ax1.yaxis.set_minor_locator(AutoMinorLocator(5))
ax1.grid(lw=0.5)
# -
# ## NumPy Random - random draws from a distribution
#
# We can draw from a variety of statistical distributions using our `numpy.random.default_rng` object (see all of them at [NumPy Random Distributions](https://numpy.org/doc/stable/reference/random/generator.html#distributions).
#
# We'll see this in action with the normal (Gaussian) distribution.
#
# ```
# normal(...) method of numpy.random._generator.Generator instance
# normal(loc=0.0, scale=1.0, size=None)
#
# Draw random samples from a normal (Gaussian) distribution.
#
# Parameters
# ----------
# loc : float or array_like of floats
# Mean ("centre") of the distribution.
# scale : float or array_like of floats
# Standard deviation (spread or "width") of the distribution. Must be
# non-negative.
# size : int or tuple of ints, optional
# Output shape. If the given shape is, e.g., ``(m, n, k)``, then
# ``m * n * k`` samples are drawn. If size is ``None`` (default),
# a single value is returned if ``loc`` and ``scale`` are both scalars.
# Otherwise, ``np.broadcast(loc, scale).size`` samples are drawn.
# ...
# ```
#
# In the docstring for `rng.normal`, we can see it asks for a `loc` (mean) and `scale` (standard deviation) of the Gaussian distribution to randomly draw from.
# +
# random number setup
draw_mean_std = (10, 0.2) # (mean, stdev)
ndraws = 10
rng = np.random.default_rng()
draws = rng.normal(*draw_mean_std, ndraws)
print(draws)
# -
# Similarly to above, we can see that after many draws, our histogram begins to look like a typical Gaussian "bell curve" centered at the mean with standard deviation spread.
# +
# random number setup
draw_mean_std = (10, 0.2)
ndraws = 1000000 # <- try changing the number of draws
rng = np.random.default_rng()
draws = rng.normal(*draw_mean_std, ndraws)
# plotting
f = plt.figure(facecolor='white', figsize=(4, 3), dpi=150)
ax = f.subplots(1, 1)
n, edges, _ = ax.hist(draws, histtype='step', range=(9, 11), bins=100)
set_plot_axis_label(ax, 'Value', 'Count')
# ax.ticklabel_format(style='sci', axis='y', scilimits=(0,0))
# -
# ## Monte Carlo -- Uncertainty (error) Propagation
#
# Recall last week we had a simple measurement with some error associated with it.
#
# What happens when we cannot measure a quantity and its error directly but still want to estimate the errors on it?
#
# This is a job for **error propagation**!
#
# ### Error propagation (analytical)
#
# If you only feed your measurement data through simple functions, we can "propagate" that error with the following error propagation equations:
#
# 
#
# (from Clemson University)
#
# This is the "analytical" way to propagate errors since we have well defined functions and can plug values in to compute the final errors.
#
# For example, say we had a study where we measured a participant's height ($H_{\rm height}$) and then the length of their hair ($H_{\rm hair}$). We want to find what their "total height ($H_{\rm total}$)" is if they were to pull their hair straight up.
#
# $H_{\rm total} = H_{\rm height} + H_{\rm hair}$
#
# You might imagine that a single measurement will sometimes give biases values, so multiple measurements of both parameters ($H_{\rm height}$, $H_{\rm hair}$) are the way to go. The more measurements you get from both parameters the closer the mean value will be to the true value and the smaller the uncertainty (the standard deviation, std) will be.
#
# Now, suppose you done 10 measurements for both $H_{\rm height}$ and $H_{\rm hair}$. As good scientists, we want an accurate way to estimate the error of the $H_{\rm total}$. One way to do it will be just add up those 10 sets of $H_{\rm height}$ and $H_{\rm hair}$ values to get 10 $H_{\rm total}$, and take a the std. The other way will be using the first formula above in the table with the means and stds of $H_{\rm height}$ and $H_{\rm hair}$:
#
# $$\sigma_{H_{\rm total}} = \sqrt{\sigma_{H_{\rm height}}^2 + \sigma_{H_{\rm hair}}^2}$$
# +
# measurements
heights = np.array([1.93, 1.82, 2.08, 2.01, 1.46, 1.34, 1.75, 2.2 , 1.95])
hair_lengths = np.array([0.01, 0.56, 0.23, 0.27, 0.21, 0.1 , 0.04, 0.06, 0.15])
# Fist moethod:
htotal = heights + hair_lengths
htotal_avg = np.mean(htotal)
htotal_std = np.std(htotal)
print(f"This person's total height using method 1 is {htotal_avg:1.2f} +/- {htotal_std:1.3f} m")
# +
# Second method:
heights_avg = np.mean(heights)
hair_lengths_avg = np.mean(hair_lengths)
heights_std = np.std(heights)
hair_lengths_std = np.std(hair_lengths)
print(f"The H_height = {heights_avg:1.2f} +/- {heights_std:1.3f} m")
print(f"The H_hair = {hair_lengths_avg:1.2f} +/- {hair_lengths_std:1.3f} m")
htotal_avg = heights_avg + hair_lengths_avg
htotal_std = np.sqrt(heights_std**2 + hair_lengths_std**2)
print(f"This person's total height using method 2 is {htotal_avg:1.2f} +/- {htotal_std:1.3f} m")
# -
# Yay we got to report errors like good scientists! But we said this section was about randomness...
#
# Say now you only know the mean and std of $H_{\rm height}$ and $H_{\rm hair}$, is it still possible to get the uncertainty of $H_{\rm total}$ using the first method?? Want to avoid the fancy error propagation formulas... Here is when the MC error propagation come it!
#
# The idea here is that as long as our errors are normal (Gaussian), we can randomly draw $H_{\rm height}$ and $H_{\rm hair}$ values from a Gaussian distribution based on the mean and std of $H_{\rm height}$ and $H_{\rm hair}$. With these values, we can calculate the $\sigma_{H_{\rm total}}$ using the first method.
#
# We already know how to draw values from a Gaussian using our RNG so let's try it!
# +
# Start with the same data and add it, compute the mean as usual
heights = np.array([1.93, 1.82, 2.08, 2.01, 1.46, 1.34, 1.75, 2.2 , 1.95])
hair_lengths = np.array([0.01, 0.56, 0.23, 0.27, 0.21, 0.1 , 0.04, 0.06, 0.15])
h_totals = heights + hair_lengths
ndraws = 1000000
rng = np.random.default_rng(seed=100) # <- remove seed to see other random results
# Make our random arrays using the mean, standard dev of our measurements
g_height = rng.normal(np.mean(heights), np.std(heights), ndraws)
g_hair = rng.normal(np.mean(hair_lengths), np.std(hair_lengths), ndraws)
# Feed the gaussian samples through the sum
g_total_arr = g_height + g_hair
h_mean = np.mean(g_total_arr)
# Because the error propagation formula for mean is just a standard deviation
# we can estimate the error of the mean with the stdev of our total array
g_mean_err = np.std(g_total_arr)
# Now we report the same meanbut can use the gaussian standard dev as the err
print(f"Mean total height (N={ndraws}): {h_mean:.2f} +/- {g_mean_err:.3f}")
# -
# The larger the ndraws you use, the more stable the g_mean_err will be.
#
# We can try increasing the number of draws by hand to see when it levels out, but we have the power of code!
#
# Let's write up some code to see how many draws we need to get a stable g_mean_err.
#
# We can define stable as *changes by less than* $10^{-4}$ for now.
# +
# using for loop
rng = np.random.default_rng(seed=100)
g_mean_err_old = np.inf # Pick large value to start us off
for ndraws in np.logspace(1, 7, 10):
ndraws = int(ndraws)
g_height = rng.normal(np.mean(heights), np.std(heights), ndraws)
g_hair = rng.normal(np.mean(hair_lengths), np.std(hair_lengths), ndraws)
g_total_arr = g_height + g_hair
g_mean_err = np.std(g_total_arr)
delta_err = np.abs(g_mean_err_old - g_mean_err)
print(f'N={ndraws:.1e}: err={g_mean_err:.3f} (changed by {delta_err:.1e})')
if delta_err < 1e-4:
print('Changed by < 1e-4! Exiting loop...')
break
g_mean_err_old = g_mean_err
print(f'\n The uncertainty of the total height is about: {g_mean_err:1.3f} N={ndraws}')
# +
# using while loop
g_mean_err_old = np.inf # <- something big to start us off
delta_err = np.inf
ndraws_arr = np.logspace(1, 7, 10)
rng = np.random.default_rng(seed=100)
# Stop loop when the difference drops below 1e-4
i = 0
while delta_err >= 1e-4:
ndraws = int(ndraws_arr[i])
g_height = rng.normal(np.mean(heights), np.std(heights), ndraws)
g_hair = rng.normal(np.mean(hair_lengths), np.std(hair_lengths), ndraws)
g_total_arr = g_height + g_hair
g_mean_err = np.std(g_total_arr)
delta_err = np.abs(g_mean_err - g_mean_err_old)
print(f'N={ndraws:.1e}: err={g_mean_err:.3f} (changed by {delta_err:.1e})')
g_mean_err_old = g_mean_err
i += 1
# Now we can report our mean with confidence in our error precision
print(f'\n The uncertainty of the total height is about: {g_mean_err:1.3f} with N={ndraws}')
# -
# Now we have an "empirical" estimate of our final error which didn't need error propagation formulas!
#
# Monte Carlo error propagation is particularly useful when you need to do a lot of manipulations to your data and don't want to write out all the propagation formulas by hand. It is also useful when your analysis involves more complicated functions than just addition, subtraction, multiplication, division (e.g., exponentials, logarithms, other non-linear functions, etc).
# ## [Assignment 1]
#
# If you have something like
#
# $$\rho = \frac{m}{V} = \frac{m}{\pi r^2 (h/3)} = \frac{3m}{\pi r^2 h}$$
#
# How fast can you get the error of the density ($\rho$) propagated from errors of $m$, $r$, and $h$?
#
# | | units | values | uncertainty ($\sigma$) |
# |-----------------|:-----:|:------:|:----------------------:|
# | Mass (m) | g | 55.5 | 4.52 |
# | Cone radius (r) | cm | 14.2 | 1.11 |
# | Cone height (h) | cm | 9.9 | 0.59 |
#
# modified from [<NAME>'s youtube video](https://www.youtube.com/watch?v=V4U6RFI6HW8&t=298s)
#
# 1. Estimate the error of the density ($\rho$)
# 2. How many draws (`ndraws`) do we need to get a stable $\rho$ value? (follow the steps in the previous code cell)
#
# ## [Challenge!]
#
# 1. Use a more strict stable condition: `differ < 1e-6` must meet for 10 consecutive `ndraws`
# 2. Find a better/faster way to reach that stable values
# +
# [your code here]
# -
# ## Monte Carlo -- Bootstrapping
#
# We can also use Monte Carlo methods to "bootstrap" confidence intervals on our measured quantities. This is often useful when we have a small sample of measurements and don't know the errors involved. Using the scatter inherent in our data and our handy random number generator, we can still get a statistical measure of errors as confidence intervals, pulling the data up by its bootstraps.
#
# **Note:** The main assumption with bootstrapping is that all measurements are *iid normal*, meaning each observation was collected independently of other measurements and is expected to have normal (Gaussian) errors.
#
# Modified from [Introduction to Statistical Methodology, Second Edition Chaper 3](https://bookdown.org/dereksonderegger/570/3-confidence-intervals-via-bootstrapping.html)
#
# Let's read in an array of data that measures the mercury levels of several lakes in Florida.
# We'll do a deep dive on pandas soon! for now we're just using it to get data
import pandas as pd
df = pd.read_csv('https://www.lock5stat.com/datasets3e/FloridaLakes.csv')
avg_mercury = df['AvgMercury'].to_numpy()
print('N measurements:', len(avg_mercury))
# Let's make a histogram to see what we're looking at!
# Plot a histogram
fig, ax = plt.subplots(facecolor='white', figsize=(4,3), dpi=150)
set_plot_axis_label(ax, 'Avg mercury', 'Count')
ax.set_title('Mercury level in Florida Lakes', size='medium', fontname='Helvetica')
ax.hist(avg_mercury, histtype='step', range=(0, np.max(avg_mercury)), bins=10,
color='black')
plt.show()
# Hmm this histogram doesn't look very Gaussian... it seems a little skewed (not symmetrical).
#
# When we have a skewed distribution, the mean of the value is not always the best measure of the center of the values. Also, what would the standard deviation be? So far we've only seen symmetrical scatter in our data...
#
# We can still calculate them but they may not represent the data as nicely as the examples we've seen so far.
print(f'Mean mercury in Florida lakes is {np.mean(avg_mercury):1.2f}, +/- {np.std(avg_mercury):.3f}')
# When we have small numbers of observations that are skewed, we can't always be confident that the mean and standard deviation are good measures of the underlying distribution in our data. This is where **bootstrapping** becomes very useful!
#
# This time we will use `rng.choice()`.
#
# ```
# choice(a, size=None, replace=True, p=None, axis=0, shuffle=True)
#
# Generates a random sample from a given array
#
# Parameters
# ----------
# a : {array_like, int}
# If an ndarray, a random sample is generated from its elements.
# If an int, the random sample is generated from np.arange(a).
# size : {int, tuple[int]}, optional
# Output shape. If the given shape is, e.g., ``(m, n, k)``, then
# ``m * n * k`` samples are drawn from the 1-d `a`. If `a` has more
# than one dimension, the `size` shape will be inserted into the
# `axis` dimension, so the output ``ndim`` will be ``a.ndim - 1 +
# len(size)``. Default is None, in which case a single value is
# returned.
# replace : bool, optional
# Whether the sample is with or without replacement. Default is True,
# meaning that a value of ``a`` can be selected multiple times.
# p : 1-D array_like, optional
# The probabilities associated with each entry in a.
# If not given, the sample assumes a uniform distribution over all
# entries in ``a``.
# ...
# ```
#
# In **bootstrapping** we want to resample the same number of data points **with replacement**, meaning the same values can be drawn multiple times. This also means we need to set `replace=True` in our `choice()` method (but since this is the default we're ok not specifying it).
rng = np.random.default_rng()
ndraws = len(avg_mercury) # Take same number of draws as in the measurement
avg_mercury_resamp = rng.choice(avg_mercury, ndraws) # replace=True by default
print(avg_mercury_resamp)
# Below we'll plot the original mean as a vertical black line and the mean of the resampled array as a vertical red line using `ax.axvline()`.
# +
# Set up plot
fig, ax = plt.subplots(facecolor='white', figsize=(4,3), dpi=150 )
set_plot_axis_label(ax, 'Avg mercury', 'Count')
ax.set_title('Mercury level in Florida Lakes', size='medium', fontname='Helvetica')
# Plot original and mean
ax.hist(avg_mercury, histtype='step', range=(0, np.max(avg_mercury)), bins=10,
color='black')
ax.axvline(np.mean(avg_mercury), lw=1, color='black', ls='--')
# Plot resampled histogram and mean
avg_mercury_resamp = rng.choice(avg_mercury, len(avg_mercury))
ax.hist(avg_mercury_resamp, histtype='step', range=(0, np.max(avg_mercury)), bins=10,
color='tab:red')
ax.axvline(np.mean(avg_mercury_resamp), lw=1, color='tab:red', ls='--')
plt.show()
# -
# It might be surprising that the mean has changed! All we did was resample from the original data and take the mean. If we do this many time, we'll get a distribution of means which we can think of as a set of possible outcomes if we had taken the measurements again and again.
#
# Because we are reusing the same measurements to resample, this won't make our mean any more accurate (because we have no new data to go off of).
#
# What this *does* do is show us how much our mean would vary if we repeated our experiment many times with the same number of observations and similar scatter in the data... It's a new way to **bootstrap** uncertainty when we didn't have any info about the error arrays to do error propagation!
#
# Below, let's rum our resampling a bunch of times and see what values we get for the mean...
# +
# plotting
fig, ax = plt.subplots(facecolor='white', figsize=(4,3), dpi=150)
set_plot_axis_label(ax, 'Avg mercury', 'Count')
ax.set_title('Mercury level in Florida Lakes', size='medium', fontname='Helvetica')
ax.axvline(np.mean(avg_mercury), lw=1, color='black', ls='--')
# random draw result
sample_times = 100000
mean_collection = []
for _ in range(sample_times):
avg_mercury_resamp = rng.choice(avg_mercury, len(avg_mercury))
mean_collection.append(np.mean(avg_mercury_resamp))
ax.hist(mean_collection, histtype='step', color='tab:red', bins=50)
plt.show()
# -
# This looks like a normal (Gaussian) distribution, so we can talk about the dispersion of the mean in terms of the standard deviation!
#
# Now we can report the uncertainty of the mean on our original plot in a more satisfying way:
# +
# Set up plot
fig, ax = plt.subplots(facecolor='white', figsize=(4,3), dpi=150 )
set_plot_axis_label(ax, 'Avg mercury', 'Count')
ax.set_title('Mercury level in Florida Lakes', size='medium', fontname='Helvetica')
# Plot original and mean
ax.hist(avg_mercury, histtype='step', range=(0, np.max(avg_mercury)), bins=10,
color='black')
ax.axvline(np.mean(avg_mercury), lw=1, color='black', ls='--')
# Calculate the 95% confidence interval [2.5%, 97.5%]
pct95_low = np.percentile(mean_collection, 2.5)
pct95_upp = np.percentile(mean_collection, 97.5)
ax.axvline(pct95_low, lw=1, color='tab:blue', ls='--', label='95th pct')
ax.axvline(pct95_upp, lw=1, color='tab:blue', ls='--')
ax.legend()
print(f'Mean mercury in Florida lakes is {np.mean(avg_mercury):1.2f}', end='')
print(f' with a 95% confidence interval of [{pct95_low:.2f}, {pct95_upp:.2f}]')
plt.show()
# -
# Now using the distribution of the bootstrapped means, we can capture the scatter in our data with N=59 observations if we were to repeat the trial many times.
#
# Now we can say we are 95% confident that the true mean mercury in Florida lakes is within our 95% confidence interval, given the measurements we made.
# ## [Assignment 2] What's the value for $\pi$?
#
# In this assignment, you will need to use the MC method to estimate the values of $\pi$.
#
# Assuming you have a quarter circle with the radius of 1 and a square that share it's edge with the quarter circle's
# radius (see the plot below).
#
# 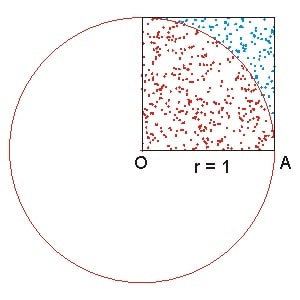
#
# Therefore, we know:
#
# $$
# Area_{\rm quarter\ circle} = \pi r^2/4 \\
# Area_{\rm square} = r^2
# $$
#
# The ratio of the two will be
#
# $$\frac{Area_{\rm quarter\ circle}}{Area_{\rm square}} = \frac{\pi r^2/4}{r^2} = \frac{\pi}{4}$$
#
# Rearange the equation and changes the area with the number of points we have:
#
# $$\pi = 4 \frac{Area_{\rm quarter\ circle}}{Area_{\rm square}} = 4 \frac{N_{\rm quarter\ circle}}{N_{\rm square}}$$
#
# where $N_{\rm quarter\ circle}$ is the number of points within the quarter circle and $N_{\rm square}$ is the number of points
# within the square.
#
# Tips:
# 1. You will need to generate two arrays with the random generator as the x and y axis of each points.
# 2. Calculate the number of points within the quarter circle and within the square
# 3. You get $\pi$ by dividing the two numbers and times 4
#
# **Show that your $\pi$ has $< 1e-5$ difference from `np.pi`**
# +
# [your code here]
| spirl/f21_a4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
import keras
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout
from keras.optimizers import RMSprop
batch_size = 128
num_classes = 10
epochs = 20
# +
# the data, shuffled and split between train and test sets
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train = x_train.reshape(60000, 784)
x_test = x_test.reshape(10000, 784)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# convert class vectors to binary class matrices
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# +
model = Sequential()
model.add(Dense(512, activation='relu', input_shape=(784,)))
model.add(Dropout(0.2))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(num_classes, activation='softmax'))
model.summary()
# +
model.compile(loss='categorical_crossentropy',
optimizer=RMSprop(),
metrics=['accuracy'])
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# -
| _posts/MLP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="CxSIyB92D3sk"
# ## Sentiment analysis using tweets in Croatian
# - tweets are translated to english
# - words in the tweet are represented by pre-trained word2vec embedding
# - mean vector of words in a tweet are used as input features
# - models are trained and tested
# + [markdown] colab_type="text" id="mHeYn87J_57O"
# #### Import libraries
# + colab={} colab_type="code" id="1yOgR_yc_XFW"
import pandas as pd # for data handling
import numpy as np # for linear algebra
import time # for timing
import matplotlib.pyplot as plt # for plotting
from gensim.models import KeyedVectors # for pre-trained embedding
from sklearn.model_selection import train_test_split # for reserving test data
# metrics for model evaluation
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
# sklearn classifiers
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from sklearn.neighbors import KNeighborsClassifier
from sklearn.linear_model import LogisticRegression
import tensorflow as tf # for neural networks
# + [markdown] colab_type="text" id="paG0XXPGEXoi"
# #### Load pre-trained word2vec model
# + colab={"base_uri": "https://localhost:8080/", "height": 136} colab_type="code" id="tUwxjhph_m0m" outputId="d2f7e50f-96ab-4ab7-ba1d-6d52640f537d"
# Retrieve embedding file using wget
# use this if embedding file is not available locally
URL = "https://s3.amazonaws.com/dl4j-distribution/" # source url
FILE = "GoogleNews-vectors-negative300.bin.gz" # source file name
SOURCE = URL+FILE # source for embedding file
DIR = "/root/input/" # directory
# ! wget -P "$DIR" -c "$SOURCE" # retrieve embedding file
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="m6N3F1qeAJi4" outputId="3a10090b-35ab-4bbe-d8c1-04ceae248c00"
# Load pre-trained word2vec model from embedding file
EMBEDDING_FILE = DIR + FILE
word2vec = KeyedVectors.load_word2vec_format(EMBEDDING_FILE, binary=True)
# Define vocabulary and embedding_size
vocabulary = set(word2vec.index2word) # set of words in vocabulary
embedding_size = word2vec.vector_size # dimension of word vector
print("Model contains %d words" %len(vocabulary))
print("Each word is represented by a %d dimensional vector" %embedding_size)
# + [markdown] colab_type="text" id="94u76zj_FGhs"
# #### Read data file with labeled tweets
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="HSOpVSuVELch" outputId="b4896d60-461f-4087-b6fc-c7bf1e434193"
DATAFILE = "croatian_cl.csv" # data file with labeled tweets
df = pd.read_csv(DATAFILE) # read file
# get tweets and targets
tweets, targets = df.en.values, df.target.values
print('Number of tweets = %d.' %len(tweets))
print('Number of labels = %d.' %len(targets))
# check for missing values and distinct tweets
print("Number of missing tweets: %d." %(np.sum(pd.isnull(tweets))))
print("Number of distinct tweets = %d" %(df.en.nunique()))
# + [markdown] colab_type="text" id="TF8ilPJkdWqm"
# #### Define function to obtain vector representation of a tweet
# + colab={} colab_type="code" id="cXZROtfLWkUI"
def mean_vector(tweet):
"""Returns mean of vector representation words in tweet.
Returns a vector of zeros if none of the words appear in vocabulary
See: https://github.com/USC-CSSL/DDR """
zero = np.zeros((embedding_size,), dtype="float32") # for null tweet
if pd.isnull(tweet): return zero # tweet missing
words = [w for w in tweet.split() if w in vocabulary] # valid words
if not words: return zero # no word in vocabulary
return np.mean([word2vec[w] for w in words], axis=0)
# + [markdown] colab_type="text" id="QW7R1PlpdiQX"
# #### Create a new dataframe with vector representations
# + colab={"base_uri": "https://localhost:8080/", "height": 270} colab_type="code" id="lFz8RnD8ZAMi" outputId="352fdb65-c2c4-4b3b-f632-469598a5a0e0"
cols = ['v_'+str(i) for i in range(embedding_size)] # column names
dfV = pd.DataFrame(list(map(mean_vector, tweets)), columns=cols) # new df
dfV.insert(0, 'target', df.target) # insert label for tweets
print('Representation has %d rows and %d columns' %dfV.shape)
dfV.to_csv("vectors_croatian.csv", index=False) # save as csv file
dfV.head() # display first 5 rows
# + [markdown] colab_type="text" id="9_9SuW-UnsJ6"
# #### Reserve testing data
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="ugSvYLenkMBG" outputId="4f1aae47-ffae-458b-e99a-0edc6c5bb87e"
features = list(dfV)[1:]
X_train, X_test, y_train, y_test = train_test_split(
dfV[features], dfV['target'], test_size=0.2, random_state=2019)
print("Training set contains %d examples" %len(X_train))
print("Test set contains %d examples" %len(X_test))
print("Number of input features = %d" %len(features))
# + colab={"base_uri": "https://localhost:8080/", "height": 187} colab_type="code" id="SAFqVS56o2Fp" outputId="94574b68-1de4-4236-d9d2-38dda3cab52d"
# Check distribution of target in training and validation data
def classDistribution(y_train, y_test):
"""Returns distribution of classes in training and test data"""
res, resCount = pd.DataFrame(), pd.DataFrame()
res['train'] = y_train.value_counts(normalize=True, sort=False)
res['test'] = y_test.value_counts(normalize=True, sort=False)
resCount['train'] = y_train.value_counts(normalize=False, sort=False)
resCount['test'] = y_test.value_counts(normalize=False, sort=False)
return res.transpose(), resCount.transpose()
dist, count = classDistribution(y_train, y_test)
print('\nClass distribution in training and test data:')
print(dist)
print('\nClass counts in training and test data:')
print(count)
# + [markdown] colab_type="text" id="M954Em8jrQu3"
# #### Train and test models
# + [markdown] colab_type="text" id="-CcNlZfu4meZ"
# ##### Specify function for evaluating trained model on test data
# + colab={} colab_type="code" id="PLWugQMMrU3D"
def evaluateModel(y_test, predicted):
"""evaluates trained model"""
acc = accuracy_score(y_test, predicted) # accuracy
print("\nAccuracy with validation data: %4.2f%%" %(100*acc))
print("\nClassification report:\n")
print(classification_report(y_test, predicted))
cm = confusion_matrix(y_test, predicted) # confusion_matrix
print("\nConfusion matrix:\n")
print(pd.DataFrame(cm))
return acc
# + [markdown] colab_type="text" id="Wb3x9JMEtBKZ"
# ##### Specify models to use
# + colab={} colab_type="code" id="m8QgKGxvyzEQ"
models = {} # dictionary of Scikit-Learn classifiers with non-default parameters
models['DT'] = DecisionTreeClassifier()
models['RF'] = RandomForestClassifier(n_estimators=100)
models['SVM'] = SVC(kernel='poly', gamma='scale')
models['KNN'] = KNeighborsClassifier(n_neighbors=3)
models['LRM'] = LogisticRegression(multi_class='auto', solver='lbfgs')
# + [markdown] colab_type="text" id="fVRhDTSb7Okf"
# ##### Train and test Scikit-Learn models
# + colab={"base_uri": "https://localhost:8080/", "height": 2652} colab_type="code" id="gnl-lpJmzJy2" outputId="156095d7-f15c-4173-9eb5-8ee73d4562d5"
result = []
for m in [m for m in models]:
model = models[m] # model to use
print("\nTraining classifier %s:\n%s" %(m, model))
st = time.time() # start time for training and testing
model.fit(X_train, y_train) # train model
predicted = model.predict(X_test) # predict test labels with trained model
t = time.time() - st # time to train and test model
print("Time to train and test classifier: %4.2f seconds" %(t))
acc = evaluateModel(y_test, predicted) # evaluate prediction accuracy
result.append([m, acc, t]) # record results
print(60*'=') # end training and testing for model
# + [markdown] colab_type="text" id="uIg1WZAm7nlq"
# ##### Use single-layered tensorflow neural network (ANN)
# + colab={} colab_type="code" id="Qs-jnuKa7lb4"
# Define single-layered tensorflow neural network (ANN)
def ann(**kwargs):
"""Returns trained single layered network"""
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(kwargs['nFeatures'],)),
tf.keras.layers.Dense(kwargs['nNeurons'], activation=tf.nn.relu),
tf.keras.layers.Dropout(kwargs['dropOutRate']),
tf.keras.layers.Dense(kwargs['nClasses'], activation=tf.nn.softmax)])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
return model
# + colab={} colab_type="code" id="Shr2DDhT79bK"
# Specify model parameters for ANN
nNeurons = 128 # number of neurons in hidden layer
dropOutRate = 0.2 # drop out rate
nFeatures= embedding_size # number of input features
nClasses = 2 # number of output classes
nEpochs = 10 # number of training epochs
# + colab={"base_uri": "https://localhost:8080/", "height": 1176} colab_type="code" id="_5ZCf1TG8OR3" outputId="1bcc617a-54e1-4375-a3e1-498da186f4a2"
# Train and test ANN model
m = 'ANN' # model name
model = ann(nNeurons=nNeurons, dropOutRate=dropOutRate,
nFeatures=nFeatures, nClasses=nClasses) # specify model
print(model.summary()) # display model summary
print("\nTraining classifier: %s" %m)
st = time.time() # start time for training and testing
hist = model.fit(X_train, y_train, epochs=nEpochs, validation_split=0.2) # train
predicted = model.predict(X_test) # predict test examples
predicted = np.argmax(predicted, axis=1) # most likely label
t = time.time() - st # time to train and test model
print("Time to train and test classifier: %4.2f seconds" %(t))
acc = evaluateModel(y_test, predicted) # evaluate prediction accuracy
result.append([m, acc, t]) # record results
print(60*'=') # end training and testing for model
# + colab={} colab_type="code" id="YBkPtpjJ90gm"
# plot training and test accuracy
def plotHistorty(model, history):
# plot history for accuracy
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title(model+' accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="3pFfYAoN-C23" outputId="9925921e-a705-46b0-f7e9-f48dcc361b05"
plotHistorty(m, hist)
# + [markdown] colab_type="text" id="WyvtBoPQAt57"
# ##### Show results
# + colab={"base_uri": "https://localhost:8080/", "height": 235} colab_type="code" id="DFXzyr7hAylF" outputId="81d4f8f7-4723-4032-e984-915fd3eb6b40"
pd.DataFrame(result, columns = ['model', 'accuracy', 'time']) # show results
| Croatian_SA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simulated annealing proof of concept
# +
# this imports the code written in the src directory
import sys
sys.path.insert(0, '../')
# imports the relevant custom code
from annealing import Annealer
from tours import DroneTour
# some useful numpy stuff used here
from numpy.random import rand
from numpy import array
# plotting fun!
from matplotlib.pyplot import subplots
# %matplotlib inline
# -
# Let $S = \{p_0, \dots, p_{n-1}\} \subseteq \mathbb{R}^2$ be a point set in the plane. A tour is simply an arbitrary sublist of the indices $\{0, \dots, n-1\}$ in arbitrary order. A $k$-tour is a list of $k$ such sublists with the property that each index in $\{0, \dots, n-1\}$ is represented in *exactly one* of the sublists.
#
# An example of a 5-tour on a point set of 10 elements might be:
#
# [
# [1, 2, 8],
# [0],
# [3],
# [4, 9, 7],
# [6]
# ]
#
# The cost associated with any tour is the sum of the distances between the points in the tour:
#
# $$
# C(\mathit{tour}, S) = \sum_{t \in \mathit{tour}} \| p_t - p_{t-1} \|_2.
# $$
#
# (here $p_{n} = p_{0} $).
def plot_policies(tours, points, axis):
"""
:param tours: a given k-tour of the point set
:type tours: list[list]
:param points: a two dimemnsional point set
:type points: numpy.ndarray
:param axis: the object that gets the plot drawn on it.
:type axis: matplotlib.axes.Axes
:return: n/a (called for side effects)
:rtype: None
Given a set of tours and a two dimensional point set, plot the points, and draw the tours.
The axis that is specified is where the plot is actually drawn on.
"""
# plot the point set
axis.plot(*points.T, 'o')
# label each point 0, ..., points.shape[0] - 1
for i in range(points.shape[0]):
axis.annotate(str(i), points[i])
for tour in tours:
# for each tour, draw a polygon on the relevant points
axis.fill(*array([points[x] for x in tour]).T, fill=False)
# We can randomly generate two-dimensional points in the unit square $[0,1] \times [0,1]$ by use of the `numpy.random.rand` call.
# generate a two dimensional point set, uniformly distributed on the box [0,1] x [0,1]
points = rand(10, 2)
# As an example of how to use simulated annealing, we create a `KDroneTour` object from the generated point set and configure an `Annealer` object. From a random starting state, we perform simulated annealing until some kind of "convergence". We plot the result to show how it might work out here:
# +
# set up the simulated annealing with a k-drone tour
ktour = DroneTour(points, 5)
annealer = Annealer(ktour)
random_start = annealer.space.random_state()
# perform simulated annealing on the random starting k-tour and return the result.
annealed_tours, penalty = annealer.anneal(random_start);
# plot the results
fig, axes = subplots(2, 1, figsize=(15, 15))
plot_policies(random_start, points, axes[0])
plot_policies(annealed_tours, points, axes[1])
for ax in axes:
ax.set_aspect('equal')
ax.axis('off');
axes[0].set_title("Random start $k$-tour");
axes[1].set_title("Result of annealing $k$-tour");
| notebooks/Proof of concept for simulated annealing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ### Amazon Sentiment Data
import lxmls.readers.sentiment_reader as srs
from lxmls.deep_learning.utils import AmazonData
corpus = srs.SentimentCorpus("books")
data = AmazonData(corpus=corpus)
# ### Train Log Linear in Pytorch
# In order to learn the differences between a numpy and a Pytorch implementation, explore the reimplementation of Ex. 3.1 in Pytorch. Compare the content of each of the functions, in particular the `forward()` and `update()` methods. The comments indicated as IMPORTANT will highlight common sources of errors.
# +
from lxmls.deep_learning.utils import Model, glorot_weight_init
import numpy as np
import torch
from torch.autograd import Variable
class PytorchLogLinear(Model):
def __init__(self, **config):
# Initialize parameters
weight_shape = (config['input_size'], config['num_classes'])
# after Xavier Glorot et al
self.weight = glorot_weight_init(weight_shape, 'softmax')
self.bias = np.zeros((1, config['num_classes']))
self.learning_rate = config['learning_rate']
# IMPORTANT: Cast to pytorch format
self.weight = Variable(torch.from_numpy(self.weight).float(), requires_grad=True)
self.bias = Variable(torch.from_numpy(self.bias).float(), requires_grad=True)
# Instantiate softmax and negative logkelihood in log domain
self.logsoftmax = torch.nn.LogSoftmax(dim=1)
self.loss = torch.nn.NLLLoss()
def _log_forward(self, input=None):
"""Forward pass of the computation graph in logarithm domain (pytorch)"""
# IMPORTANT: Cast to pytorch format
input = Variable(torch.from_numpy(input).float(), requires_grad=False)
# Linear transformation
z = torch.matmul(input, torch.t(self.weight)) + self.bias
# Softmax implemented in log domain
log_tilde_z = self.logsoftmax(z)
# NOTE that this is a pytorch class!
return log_tilde_z
def predict(self, input=None):
"""Most probably class index"""
log_forward = self._log_forward(input).data.numpy()
return np.argmax(np.exp(log_forward), axis=1)
def update(self, input=None, output=None):
"""Stochastic Gradient Descent update"""
# IMPORTANT: Class indices need to be casted to LONG
true_class = Variable(torch.from_numpy(output).long(), requires_grad=False)
# Compute negative log-likelihood loss
loss = self.loss(self._log_forward(input), true_class)
# Use autograd to compute the backward pass.
loss.backward()
# SGD update
self.weight.data -= self.learning_rate * self.weight.grad.data
self.bias.data -= self.learning_rate * self.bias.grad.data
# Zero gradients
self.weight.grad.data.zero_()
self.bias.grad.data.zero_()
return loss.data.numpy()
# -
# Once you understand the model you can instantiate it and run it using the standard training loop we have used on previous exercises.
model = PytorchLogLinear(
input_size=corpus.nr_features,
num_classes=2,
learning_rate=0.05
)
# +
# Hyper-parameters
num_epochs = 10
batch_size = 30
# Get batch iterators for train and test
train_batches = data.batches('train', batch_size=batch_size)
test_set = data.batches('test', batch_size=None)[0]
# Epoch loop
for epoch in range(num_epochs):
# Batch loop
for batch in train_batches:
model.update(input=batch['input'], output=batch['output'])
# Prediction for this epoch
hat_y = model.predict(input=test_set['input'])
# Evaluation
accuracy = 100*np.mean(hat_y == test_set['output'])
# Inform user
print("Epoch %d: accuracy %2.2f %%" % (epoch+1, accuracy))
| labs/notebooks/non_linear_classifiers/exercise_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="NBQO0RjarlGP" colab_type="code" colab={}
import pandas as pd
from sklearn.tree import DecisionTreeClassifier
# + id="FXOW2dU1rlGY" colab_type="code" colab={}
df = pd.read_csv('winequality-white.csv', sep=';')
# + id="Pk6QNdRWrlGd" colab_type="code" colab={} outputId="afc7ee8f-9806-40b6-f1c2-678d48b02b57"
df.head()
# + id="WRejjTKGrlGh" colab_type="code" colab={} outputId="a089dfb8-14db-4e7a-c6fd-ac939154e9de"
print (df.shape)
df.describe()
# + id="Avgxe5PUrlGl" colab_type="code" colab={} outputId="3a58fe82-ba29-4f29-e2b2-e62f3ed10f3c"
df['quality'].unique()
# + id="TvXI9yFfrlGp" colab_type="code" colab={} outputId="2318d217-afce-46d0-b617-61511410df04"
df['quality'].value_counts()
# + [markdown] id="HHH82mJkrlGs" colab_type="text"
# We have to treating the problem as a classification problem, so we are going to use a decision tree to learn a classification model that predicts red wine quality based on the features. Since we have to predict the wine quality the attribute "quality" will become our label and the rest of the attributes will become the features.
#
# The target variable "quality" ranges from 3 to 9. We can notice that the most observations are in class 6. In class 9 we a few observations.
# + id="6xGUTggZrlGt" colab_type="code" colab={}
X = df.drop('quality', axis = 1)
# + id="grcpDcCKrlGw" colab_type="code" colab={}
Y = df.quality
# + id="IfNVoV-lrlGz" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.20, random_state=42)
# + id="R2C05TWqrlG2" colab_type="code" colab={} outputId="8cd38db3-21f2-496b-864f-9b6b5f1b1151"
from sklearn import preprocessing
X_train_scaled = preprocessing.scale(X_train)
print (X_train_scaled)
# + id="ZuMMemIdrlG5" colab_type="code" colab={} outputId="c59b3d26-c4f5-435a-e97a-b805e7d9c10c"
clf = DecisionTreeClassifier(criterion = 'entropy', splitter = 'best', max_depth = 15)
clf.fit(X_train, Y_train)
# + [markdown] id="y2AVRQyUrlG9" colab_type="text"
# We just stored label "quality" in Y, which is the common used to represent the labels in machine learning and in X the features. Next we split our dataset into test and train data.Training data is the data on which the machine learning programs learn to perform correlational tasks. Testing data is the data, whose outcome is already known (even the outcome of training data is known) and is used to determine the accuracy of the machine learning algorithm, based on the training data (how effectively the learning happened). We will be using train data to train our model for predicting the quality. Also we take the 20% of the original population and use it for testing.
# + id="XuZjKswprlG9" colab_type="code" colab={}
y_pred = clf.predict(X_test)
# + id="zsY2T430rlHA" colab_type="code" colab={} outputId="7beca415-562b-4db2-d6f6-ec7e11b70732"
from sklearn.metrics import accuracy_score
accuracy_score (Y_test, y_pred)
# + id="wDxYWk0_rlHE" colab_type="code" colab={}
import graphviz
dot_data = tree.export_graphviz(clf, out_file=None)
graph = graphviz.Source(dot_data)
graph
# + id="NQSWsHotrlHH" colab_type="code" colab={}
prominent_features = clf.feature_importances_
# + id="YHuJYuMdrlHJ" colab_type="code" colab={} outputId="3d03f6ef-7525-4856-bd3b-4579b2eeb07b"
for importance,feature in zip(prominent_features, X):
print ('{}: {}'.format(feature, importance))
# + [markdown] id="JaS0_miDrlHM" colab_type="text"
# We can observe that alcohol content and free sulfur dioxide play the two largest roles in the decision of classifier.
| Exercise 1 - Wine quality dataset/Decision_trees_white_wine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''base'': conda)'
# name: python3
# ---
# + [markdown] nteract={"transient": {"deleting": false}}
# ---
# author: <NAME> (<EMAIL>)
# ---
# + [markdown] tags=[] cell_id="00000-cf7aaffc-aa63-4477-a342-556a58ec1088" deepnote_cell_type="markdown"
# The solution below uses an example dataset about the teeth of 10 guinea pigs at three Vitamin C dosage levels (in mg) with two delivery methods (orange juice vs. ascorbic acid). (See how to quickly load some sample data.)
# + tags=[] cell_id="00001-963a7869-ed17-4fdb-8d6c-b2f1a721d5ab" deepnote_to_be_reexecuted=false source_hash="3efdab3d" execution_start=1626012552914 execution_millis=3 deepnote_cell_type="code"
from rdatasets import data
df = data('ToothGrowth')
# + [markdown] tags=[] cell_id="00002-03de9dff-1148-4c39-a7fc-80e86fcda6cb" deepnote_cell_type="markdown"
# If you wish to understand the distribution of a numeric variable (here "len") compared across different values of a categorical variable (here "supp"), you can construct a bivariate histogram. We use Seaborn and Matplotlib to do so.
# + tags=[] cell_id="00002-3b7a17d0-8d64-4841-aceb-36fbe1e9ac12" deepnote_to_be_reexecuted=false source_hash="f5a9e620" execution_start=1626012552958 execution_millis=1590 deepnote_cell_type="code"
import seaborn as sns
import matplotlib.pyplot as plt
sns.displot(df, x="len", col="supp", stat="density")
plt.show()
# + [markdown] tags=[] cell_id="00004-a7f93012-1e7c-48ea-9eb8-3f2802e5d2e6" deepnote_cell_type="markdown"
# To visualize the same information summarized using quartiles only, you can construct a bivariate box plot.
# + tags=[] cell_id="00003-4d8a8cc4-c009-4259-8d09-060fccb684d3" deepnote_to_be_reexecuted=false source_hash="632e8e8e" execution_start=1626012554593 execution_millis=157 deepnote_cell_type="code"
sns.boxplot(x="supp", y="len", data = df, order = ['OJ','VC'])
plt.show()
# + [markdown] tags=[] cell_id="00006-a4b0778e-bfc7-4120-86fa-ed8c607a3364" deepnote_cell_type="markdown"
# Even more simply, we may wish to plot just the means and 95% confidence intervals around the mean for the quantitative variable, for each of the values of the categorical variable. We do so with a point plot.
# + tags=[] cell_id="00006-ad976a11-1cef-4284-b4a1-3ba149d9fdfc" deepnote_to_be_reexecuted=false source_hash="30f30feb" execution_start=1626012602158 execution_millis=298 deepnote_cell_type="code"
sns.pointplot(x = 'supp', y = 'len', data = df,
ci = 95, # Which confidence interval? Here 95%.
capsize = 0.1) # Size of "cap" drawn on each confidence interval.
plt.show()
| database/tasks/How to create bivariate plots to compare groups/Python, using Matplotlib and Seaborn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="-nzEnXavJBmB"
# **Estimación puntual**
#
#
#
#
# + id="HHv8wwNTJCBW"
import numpy as np
import pandas as pd
import scipy.stats as stats
import matplotlib.pyplot as plt
import random
import math
# + id="SLRrHrJTJx0N" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="f86e8965-16b3-493b-c541-b8743df47a1f"
np.random.seed(2020)
population_ages_1 = stats.poisson.rvs(loc = 18, mu = 35, size = 1500000)
population_ages_2 = stats.poisson.rvs(loc = 18, mu = 10, size = 1000000)
population_ages = np.concatenate((population_ages_1, population_ages_2))
print(population_ages_1.mean())
print(population_ages_2.mean())
print(population_ages.mean())
# + id="TnXJN7unKs6t" colab={"base_uri": "https://localhost:8080/", "height": 641} outputId="095fc5a7-1329-4c6e-d169-3822754590d8"
pd.DataFrame(population_ages).hist(bins = 60, range = (17.5, 77.5), figsize = (10,10))
# + id="qh1C3KzNLUON" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="89dcc976-7643-488e-b29c-cf451bdfd72c"
stats.skew(population_ages)
# + id="IlcHDgyBLkGJ" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="17f86887-dd36-4873-cbed-2da864da6770"
stats.kurtosis(population_ages)
# + id="raYauBDiLySP" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9a3d3e3d-ec84-4f60-94ec-c575a58fee56"
np.random.seed(42)
sample_ages = np.random.choice(population_ages, 500)
print(sample_ages.mean())
# + id="iBL5JbfJMFpE" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7094db63-ed75-4765-e946-69971be891bb"
population_ages.mean() - sample_ages.mean()
# + id="B5vQhKnTMYAs"
population_races = (["blanca"]*1000000) + (["negra"]*500000) + (["hispana"]*500000) + (["asiatica"]*250000) + (["otros"]*250000)
# + id="qB-k9hzgM31A" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="8a06739d-7c77-42ae-cd92-7160e2873d4d"
for race in set(population_races):
print("Proporción de "+race)
print(population_races.count(race) / 2500000)
# + id="i7HIhCZPNXSI"
random.seed(31)
race_sample = random.sample(population_races, 1000)
# + id="P47SX_KNNhJO" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="14fc93b6-6526-43e3-939b-65b4135689ad"
for race in set(race_sample):
print("Proporción de "+race)
print(race_sample.count(race) / 1000)
# + id="VPkcGvDsN0zz" colab={"base_uri": "https://localhost:8080/", "height": 641} outputId="2b173f72-14d5-4696-803a-2a6ea2750f95"
pd.DataFrame(population_ages).hist(bins = 60, range = (17.5, 77.5), figsize = (10,10))
# + id="8z3MDtHAN2ZG" colab={"base_uri": "https://localhost:8080/", "height": 641} outputId="65d76530-91d2-4863-e888-e8d728d50174"
pd.DataFrame(sample_ages).hist(bins = 60, range = (17.5, 77.5), figsize = (10,10))
# + id="u8U6RJrQOJt0" colab={"base_uri": "https://localhost:8080/", "height": 554} outputId="673a588a-6e7f-41d1-bd78-993de0323036"
np.random.sample(1988)
point_estimates = []
for x in range(200):
sample = np.random.choice(population_ages, size = 500)
point_estimates.append(sample.mean())
pd.DataFrame(point_estimates).plot(kind = "density", figsize = (9,9), xlim = (40, 46) )
# + id="yudxlrFoPBtw" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a45c40c7-f923-430f-a5ff-724ea56e616a"
np.array(point_estimates).mean()
# + [markdown] id="wnwPnlqnS9n-"
# **Si conocemos la desviación típica**
# + id="IqFFnPcPP3Jj" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="32d75aad-5a43-46d4-9578-0b1f0b4a7594"
np.random.seed(10)
n = 1000
alpha = 0.05
sample = np.random.choice(population_ages, size = n)
sample_mean = sample.mean()
z_critical = stats.norm.ppf(q = 1-alpha/2)
sigma = population_ages.std()## sigma de la población
sample_error = z_critical * sigma / math.sqrt(n)
ci = (sample_mean - sample_error, sample_mean + sample_error)
ci
# + id="USs_XdPtRFbE"
np.random.seed(10)
n = 1000
alpha = 0.05
intervals = []
sample_means = []
z_critical = stats.norm.ppf(q = 1-alpha/2)
sigma = population_ages.std()## sigma de la población
sample_error = z_critical * sigma / math.sqrt(n)
for sample in range(100):
sample = np.random.choice(population_ages, size = n)
sample_mean = sample.mean()
sample_means.append(sample_mean)
ci = (sample_mean - sample_error, sample_mean + sample_error)
intervals.append(ci)
# + id="0sSpn89rRx8a" colab={"base_uri": "https://localhost:8080/", "height": 609} outputId="ba555650-d5ce-4b08-9d55-8edb5d49e317"
plt.figure(figsize=(10,10))
plt.errorbar(x = np.arange(0.1, 100, 1), y = sample_means, yerr=[(top-bottom)/2 for top, bottom in intervals], fmt='o')
plt.hlines(xmin = 0, xmax = 100, y = population_ages.mean(), linewidth=2.0, color="red")
# + [markdown] id="ixjOCYxoS3cB"
# **Si la desviación típica no es conocida...**
#
# + id="VLgeLSSwS1oa" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f24ee55c-b8a0-417d-fa91-7cab9a7851f5"
np.random.seed(10)
n = 25
alpha = 0.05
sample = np.random.choice(population_ages, size = n)
sample_mean = sample.mean()
t_critical = stats.t.ppf(q = 1-alpha/2, df = n-1)
sample_sd = sample.std(ddof=1)## desviación estándar de la muestra
sample_error = t_critical * sample_sd / math.sqrt(n)
ci = (sample_mean - sample_error, sample_mean + sample_error)
ci
# + id="BdA4_zsxTtMx" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="eb908ba9-e055-46de-c7c6-57038a9ead0d"
stats.t.ppf(q = 1-alpha, df = n-1) - stats.norm.ppf(1-alpha)
# + id="BQpn1PqIT2Af" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d18a8377-c9c8-4885-9fa2-b8ad42cfcef9"
stats.t.ppf(q = 1-alpha, df = 999) - stats.norm.ppf(1-alpha)
# + id="-vZU2n6yT-iY" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="938a4103-b7b3-44e7-8f4a-8a674890154c"
stats.t.interval(alpha = 0.95, df = 24, loc = sample_mean, scale = sample_sd/math.sqrt(n))
# + [markdown] id="cxpk5Etsdcx1"
# **Intervalo para la proporción poblacional**
# + id="dhoouJ5bUs7i" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9eac3581-7d91-4b9b-a6ac-e19c3ba640f2"
alpha = 0.05
n = 1000
z_critical = stats.norm.ppf(q=1-alpha/2)
p_hat = race_sample.count("blanca") / n
sample_error = z_critical * math.sqrt((p_hat*(1-p_hat)/n))
ci = (p_hat - sample_error, p_hat + sample_error)
ci
# + id="BmOc-Fy6VUoU" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="25161d22-48ee-45cb-8498-b9d74a353e02"
stats.norm.interval(alpha = 0.95, loc = p_hat, scale = math.sqrt(p_hat*(1-p_hat)/n))
# + [markdown] id="NDHY31KOdgvE"
# **Cómo interpretar el intervalo de confianza**
# + id="zFRnrkl-dllc"
shape, scale = 2.0, 2.0 #mean = 4, std = 2*sqrt(2)
s = np.random.gamma(shape, scale, 1000000)
mu = shape*scale
sigma = scale*np.sqrt(shape)
# + id="sK0HNKDme-a0" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="3ec6b587-8f75-43ce-cc6f-bc96a982d77e"
print(mu)
print(sigma)
# + id="KYgaW9P_eKg9" colab={"base_uri": "https://localhost:8080/", "height": 592} outputId="5ebb008b-1005-4156-fae0-11b689174451"
meansample = []
sample_size = 500
for i in range(0,50000):
sample = random.choices(s, k=sample_size)
meansample.append(sum(sample)/len(sample))
plt.figure(figsize=(20,10))
plt.hist(meansample, 200, density=True, color="lightblue")
plt.show()
# + id="RmOdXchUfLf_" colab={"base_uri": "https://localhost:8080/", "height": 592} outputId="5158ca00-30d1-481e-fe7d-73237cc7ae90"
plt.figure(figsize=(20,10))
plt.hist(meansample, 200, density=True, color="lightblue")
plt.plot([mu,mu], [0, 3.5], 'k-', lw=4, color='green')
plt.plot([mu-1.96*sigma/np.sqrt(sample_size), mu-1.96*sigma/np.sqrt(sample_size)], [0, 3.5], 'k-', lw=2, color="navy")
plt.plot([mu+1.96*sigma/np.sqrt(sample_size), mu+1.96*sigma/np.sqrt(sample_size)], [0, 3.5], 'k-', lw=2, color="navy")
plt.show()
# + id="ePtKCtKBgCl6" colab={"base_uri": "https://localhost:8080/", "height": 592} outputId="d0717c77-ec90-4b2f-ee91-98d2237423c3"
sample_data = np.random.choice(s, size = sample_size)
x_bar = sample_data.mean()
ss = sample_data.std()
plt.figure(figsize=(20,10))
plt.hist(meansample, 200, density=True, color="lightblue")
plt.plot([mu,mu], [0, 3.5], 'k-', lw=4, color='green')
plt.plot([mu-1.96*sigma/np.sqrt(sample_size), mu-1.96*sigma/np.sqrt(sample_size)], [0, 3.5], 'k-', lw=2, color="navy")
plt.plot([mu+1.96*sigma/np.sqrt(sample_size), mu+1.96*sigma/np.sqrt(sample_size)], [0, 3.5], 'k-', lw=2, color="navy")
plt.plot([x_bar, x_bar], [0,3.5], 'k-', lw=2, color="red")
plt.plot([x_bar-1.96*ss/np.sqrt(sample_size), x_bar-1.96*ss/np.sqrt(sample_size)], [0, 3.5], 'k-', lw=1, color="red")
plt.plot([x_bar+1.96*ss/np.sqrt(sample_size), x_bar+1.96*ss/np.sqrt(sample_size)], [0, 3.5], 'k-', lw=1, color="red")
plt.gca().add_patch(plt.Rectangle((x_bar-1.96*ss/np.sqrt(sample_size), 0), 2*(1.96*ss/np.sqrt(sample_size)), 3.5, fill=True, fc=(0.9, 0.1, 0.1, 0.15)))
plt.show()
# + id="JX6xBF3Di1az"
interval_list = []
z_critical = 1.96 #z_0.975
sample_size = 5000
c = 0
error = z_critical*sigma/np.sqrt(sample_size)
for i in range(0,100):
rs = random.choices(s, k=sample_size)
mean = np.mean(rs)
ub = mean + error
lb = mean - error
interval_list.append([lb, mean, ub])
if ub >= mu and lb <= mu:
c += 1
# + id="Pbj8UR89jpnE" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="61e41264-f9bd-41b9-f7fc-7cb07bf54e36"
c
# + id="pTrCmzAhjuSE" colab={"base_uri": "https://localhost:8080/", "height": 609} outputId="33ca8a33-15ff-4c7d-c14e-3937f57e0922"
print("Número de intervalos de confianza que contienen el valor real de mu: ",c)
plt.figure(figsize = (20, 10))
plt.boxplot(interval_list)
plt.plot([1,100], [mu, mu], 'k-', lw=2, color="red")
plt.show()
| inference-statistics/inference-statistics-course/notebooks/Intervalos_de_Confianza.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="iYkoOog-S7pV" papermill={"duration": 0.013762, "end_time": "2020-10-18T08:49:16.318900", "exception": false, "start_time": "2020-10-18T08:49:16.305138", "status": "completed"} tags=[]
# # Preparation
# + id="0-ecCOmQSbrr" outputId="588f32de-8b1d-4ac0-ca62-c73aef70e67c" papermill={"duration": 5.705715, "end_time": "2020-10-18T08:49:22.037698", "exception": false, "start_time": "2020-10-18T08:49:16.331983", "status": "completed"} tags=[]
#config
import warnings
warnings.filterwarnings("ignore")
import sys
sys.path.append('../input/iterative-stratification/iterative-stratification-master')
from iterstrat.ml_stratifiers import MultilabelStratifiedKFold
#Utils
import gc
import datetime
#Sklearn
from sklearn.model_selection import KFold, StratifiedKFold
from sklearn.metrics import log_loss
from scipy.optimize import minimize
from tqdm.notebook import tqdm
from time import time
#Tensorflow
import tensorflow as tf
tf.random.set_seed(93845)
import tensorflow.keras.backend as K
import tensorflow.keras.layers as L
import tensorflow.keras.models as M
from tensorflow.keras.callbacks import ReduceLROnPlateau, ModelCheckpoint, EarlyStopping
import tensorflow_addons as tfa
#Pandas and numpy
import pandas as pd
import numpy as np
np.random.seed(93845)
# + [markdown] id="fK9XL_AVVojN" papermill={"duration": 0.019067, "end_time": "2020-10-18T08:49:22.077151", "exception": false, "start_time": "2020-10-18T08:49:22.058084", "status": "completed"} tags=[]
# ## Configuration
# + id="IMYOKGdOVoGm" outputId="9d02acb0-0c88-4407-c454-d388b74c59db" papermill={"duration": 0.051436, "end_time": "2020-10-18T08:49:22.149778", "exception": false, "start_time": "2020-10-18T08:49:22.098342", "status": "completed"} tags=[]
MIXED_PRECISION = False
XLA_ACCELERATE = True
if MIXED_PRECISION:
from tensorflow.keras.mixed_precision import experimental as mixed_precision
if tpu: policy = tf.keras.mixed_precision.experimental.Policy('mixed_bfloat16')
else: policy = tf.keras.mixed_precision.experimental.Policy('mixed_float16')
mixed_precision.set_policy(policy)
print('Mixed precision enabled')
if XLA_ACCELERATE:
tf.config.optimizer.set_jit(True)
print('Accelerated Linear Algebra enabled')
# + [markdown] id="B9Jz2Xo7VruL" papermill={"duration": 0.021565, "end_time": "2020-10-18T08:49:22.193945", "exception": false, "start_time": "2020-10-18T08:49:22.172380", "status": "completed"} tags=[]
# ## Charge data
# + id="ZxX6gNhdTBJy" papermill={"duration": 5.852263, "end_time": "2020-10-18T08:49:28.067614", "exception": false, "start_time": "2020-10-18T08:49:22.215351", "status": "completed"} tags=[]
ss = pd.read_csv('../input/lish-moa/sample_submission.csv')
test_features = pd.read_csv('../input/lish-moa/test_features.csv')
train_features = pd.read_csv('../input/lish-moa/train_features.csv')
train_targets= pd.read_csv('../input/lish-moa/train_targets_scored.csv')
ss_2 = ss.copy()
ss_3 = ss.copy()
ss_blend = ss.copy()
cols = [c for c in ss.columns.values if c != 'sig_id']
# + [markdown] id="mRHrMDLzWa9Y" papermill={"duration": 0.01374, "end_time": "2020-10-18T08:49:28.095436", "exception": false, "start_time": "2020-10-18T08:49:28.081696", "status": "completed"} tags=[]
# # Data preparation
# + id="MDNOOpmdT6xN" papermill={"duration": 0.04819, "end_time": "2020-10-18T08:49:28.157894", "exception": false, "start_time": "2020-10-18T08:49:28.109704", "status": "completed"} tags=[]
def preprocess(df):
df.loc[:, 'cp_type'] = df.loc[:, 'cp_type'].map({'trt_cp': 0, 'ctl_vehicle': 1})
df.loc[:, 'cp_dose'] = df.loc[:, 'cp_dose'].map({'D1': 0, 'D2': 1})
del df['sig_id']
return df
def log_loss_metric(y_true, y_pred):
metrics = []
for _target in train_targets.columns:
metrics.append(log_loss(y_true.loc[:, _target], y_pred.loc[:, _target].astype(float), labels = [0,1]))
return np.mean(metrics)
train = preprocess(train_features)
test = preprocess(test_features)
del train_targets['sig_id']
# + [markdown] id="SoJw_fzNXjr2" papermill={"duration": 0.013565, "end_time": "2020-10-18T08:49:28.185423", "exception": false, "start_time": "2020-10-18T08:49:28.171858", "status": "completed"} tags=[]
# # Model function
# + id="RCPJf4DVXfqE" papermill={"duration": 0.037863, "end_time": "2020-10-18T08:49:28.237058", "exception": false, "start_time": "2020-10-18T08:49:28.199195", "status": "completed"} tags=[]
def create_model(num_columns, hidden_units, dropout_rate, learning_rate):
#First input
inp1 = tf.keras.layers.Input(shape = (num_columns, ))
x1 = tf.keras.layers.BatchNormalization()(inp1)
for i, units in enumerate(hidden_units[0]):
x1 = tfa.layers.WeightNormalization(tf.keras.layers.Dense(units, activation = 'elu'))(x1)
x1 = tf.keras.layers.Dropout(dropout_rate[0])(x1)
x1 = tf.keras.layers.BatchNormalization()(x1)
#Second input
inp2 = tf.keras.layers.Input(shape = (num_columns, ))
x2 = tf.keras.layers.BatchNormalization()(inp2)
for i, units in enumerate(hidden_units[1]):
x2 = tfa.layers.WeightNormalization(tf.keras.layers.Dense(units, activation = 'elu'))(x2)
x2 = tf.keras.layers.Dropout(dropout_rate[1])(x2)
x2 = tf.keras.layers.BatchNormalization()(x2)
#Third input
inp3 = tf.keras.layers.Input(shape = (num_columns, ))
x3 = tf.keras.layers.BatchNormalization()(inp3)
for i, units in enumerate(hidden_units[2]):
x3 = tfa.layers.WeightNormalization(tf.keras.layers.Dense(units, activation = 'elu'))(x3)
x3 = tf.keras.layers.Dropout(dropout_rate[1])(x3)
x3 = tf.keras.layers.BatchNormalization()(x3)
#Concatenate layer
x = tf.keras.layers.Concatenate()([x1,x2,x3])
x = tf.keras.layers.Dropout(dropout_rate[3])(x)
x = tf.keras.layers.BatchNormalization()(x)
#Final layers layer
for units in hidden_units[3]:
x = tfa.layers.WeightNormalization(tf.keras.layers.Dense(units, activation = 'elu'))(x)
x = tf.keras.layers.Dropout(dropout_rate[4])(x)
x = tf.keras.layers.BatchNormalization()(x)
out = tfa.layers.WeightNormalization(tf.keras.layers.Dense(206, activation = 'sigmoid'))(x)
model = tf.keras.models.Model(inputs = [inp1, inp2, inp3], outputs = out)
model.compile(optimizer = tfa.optimizers.Lookahead(tf.optimizers.Adam(learning_rate), sync_period= 10),
loss = 'binary_crossentropy')
return model
# + [markdown] id="_ri1UD6fzUXa" papermill={"duration": 0.01387, "end_time": "2020-10-18T08:49:28.264478", "exception": false, "start_time": "2020-10-18T08:49:28.250608", "status": "completed"} tags=[]
# ## Plot of the model
# + id="GzzxEUFly2x9" outputId="702f770c-d19f-4ed5-fd32-1a80f7e76346" papermill={"duration": 5.248937, "end_time": "2020-10-18T08:49:33.527257", "exception": false, "start_time": "2020-10-18T08:49:28.278320", "status": "completed"} tags=[]
hidden_units = [[2048, 512, 2048],
[512, 1024, 512],
[512, 1024, 2048, 1024, 512],
[1024, 1024]]
dropout_rate = [0.4, 0.3, 0.45, 0.3, 0.4]
size = int(np.ceil(0.8 * len(train.columns.values)))
model = create_model(size, hidden_units, dropout_rate, 1e-3)
tf.keras.utils.plot_model(model,
show_shapes = False,
show_layer_names = True,
rankdir = 'TB',
expand_nested = False,
dpi = 60)
# + [markdown] id="YrKctIuZ1KDH" papermill={"duration": 0.015367, "end_time": "2020-10-18T08:49:33.558442", "exception": false, "start_time": "2020-10-18T08:49:33.543075", "status": "completed"} tags=[]
# # Train the model
# + [markdown] id="mA9yW8vFIuZ-" papermill={"duration": 0.015645, "end_time": "2020-10-18T08:49:33.589805", "exception": false, "start_time": "2020-10-18T08:49:33.574160", "status": "completed"} tags=[]
# ## Top feats
#
# Already calculated by eli5
# + id="MTcMbn3lIpD_" outputId="95661649-a4eb-4257-af78-7bdab89e5a75" papermill={"duration": 0.071143, "end_time": "2020-10-18T08:49:33.676531", "exception": false, "start_time": "2020-10-18T08:49:33.605388", "status": "completed"} tags=[]
top_feats = [ 0, 1, 2, 3, 5, 6, 8, 9, 10, 11, 12, 14, 15,
16, 18, 19, 20, 21, 23, 24, 25, 27, 28, 29, 30, 31,
32, 33, 34, 35, 36, 37, 39, 40, 41, 42, 44, 45, 46,
48, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61,
63, 64, 65, 66, 68, 69, 70, 71, 72, 73, 74, 75, 76,
78, 79, 80, 81, 82, 83, 84, 86, 87, 88, 89, 90, 92,
93, 94, 95, 96, 97, 99, 100, 101, 103, 104, 105, 106, 107,
108, 109, 110, 111, 112, 113, 114, 115, 116, 117, 118, 119, 120,
121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 132, 133, 134,
135, 136, 137, 138, 139, 140, 141, 142, 143, 144, 145, 146, 147,
149, 150, 151, 152, 153, 154, 155, 157, 159, 160, 161, 163, 164,
165, 166, 167, 168, 169, 170, 172, 173, 175, 176, 177, 178, 180,
181, 182, 183, 184, 186, 187, 188, 189, 190, 191, 192, 193, 195,
197, 198, 199, 202, 203, 205, 206, 208, 209, 210, 211, 212, 213,
214, 215, 218, 219, 220, 221, 222, 224, 225, 227, 228, 229, 230,
231, 232, 233, 234, 236, 238, 239, 240, 241, 242, 243, 244, 245,
246, 248, 249, 250, 251, 253, 254, 255, 256, 257, 258, 259, 260,
261, 263, 265, 266, 268, 270, 271, 272, 273, 275, 276, 277, 279,
282, 283, 286, 287, 288, 289, 290, 294, 295, 296, 297, 299, 300,
301, 302, 303, 304, 305, 306, 308, 309, 310, 311, 312, 313, 315,
316, 317, 320, 321, 322, 324, 325, 326, 327, 328, 329, 330, 331,
332, 333, 334, 335, 338, 339, 340, 341, 343, 344, 345, 346, 347,
349, 350, 351, 352, 353, 355, 356, 357, 358, 359, 360, 361, 362,
363, 364, 365, 366, 368, 369, 370, 371, 372, 374, 375, 376, 377,
378, 379, 380, 382, 383, 384, 385, 386, 387, 388, 389, 390, 391,
392, 393, 394, 395, 397, 398, 399, 400, 401, 403, 405, 406, 407,
408, 410, 411, 412, 413, 414, 415, 417, 418, 419, 420, 421, 422,
423, 424, 425, 426, 427, 428, 429, 430, 431, 432, 433, 434, 435,
436, 437, 438, 441, 442, 443, 444, 445, 446, 447, 448, 449, 450,
452, 453, 454, 456, 457, 458, 459, 460, 461, 462, 463, 464, 465,
466, 468, 469, 471, 472, 473, 474, 475, 476, 477, 478, 479, 482,
483, 485, 486, 487, 488, 489, 491, 492, 494, 495, 496, 500, 501,
502, 503, 505, 506, 507, 509, 510, 511, 512, 513, 514, 516, 517,
518, 519, 521, 523, 525, 526, 527, 528, 529, 530, 531, 532, 533,
534, 536, 537, 538, 539, 540, 541, 542, 543, 544, 545, 546, 547,
549, 550, 553, 554, 555, 556, 557, 558, 559, 560, 561, 562, 563,
564, 565, 566, 567, 569, 570, 571, 572, 573, 574, 575, 577, 580,
581, 582, 583, 586, 587, 590, 591, 592, 593, 595, 596, 597, 598,
599, 600, 601, 602, 603, 605, 607, 608, 609, 611, 612, 613, 614,
615, 616, 617, 619, 622, 623, 625, 627, 630, 631, 632, 633, 634,
635, 637, 638, 639, 642, 643, 644, 645, 646, 647, 649, 650, 651,
652, 654, 655, 658, 659, 660, 661, 662, 663, 664, 666, 667, 668,
669, 670, 672, 674, 675, 676, 677, 678, 680, 681, 682, 684, 685,
686, 687, 688, 689, 691, 692, 694, 695, 696, 697, 699, 700, 701,
702, 703, 704, 705, 707, 708, 709, 711, 712, 713, 714, 715, 716,
717, 723, 725, 727, 728, 729, 730, 731, 732, 734, 736, 737, 738,
739, 740, 741, 742, 743, 744, 745, 746, 747, 748, 749, 750, 751,
752, 753, 754, 755, 756, 758, 759, 760, 761, 762, 763, 764, 765,
766, 767, 769, 770, 771, 772, 774, 775, 780, 781, 782, 783, 784,
785, 787, 788, 790, 793, 795, 797, 799, 800, 801, 805, 808, 809,
811, 812, 813, 816, 819, 820, 821, 822, 823, 825, 826, 827, 829,
831, 832, 833, 834, 835, 837, 838, 839, 840, 841, 842, 844, 845,
846, 847, 848, 850, 851, 852, 854, 855, 856, 858, 860, 861, 862,
864, 867, 868, 870, 871, 873, 874]
print(len(top_feats))
# + [markdown] id="ioNtWl0MI7Pb" papermill={"duration": 0.015843, "end_time": "2020-10-18T08:49:33.709438", "exception": false, "start_time": "2020-10-18T08:49:33.693595", "status": "completed"} tags=[]
# ## Training
# + [markdown] id="yZDeNK72baGh" papermill={"duration": 0.015707, "end_time": "2020-10-18T08:49:33.741222", "exception": false, "start_time": "2020-10-18T08:49:33.725515", "status": "completed"} tags=[]
# ### Personaized callback for time tracking on training
# + id="LNAJUU8sbZxF" papermill={"duration": 0.07527, "end_time": "2020-10-18T08:49:33.832770", "exception": false, "start_time": "2020-10-18T08:49:33.757500", "status": "completed"} tags=[]
from keras.callbacks import Callback
class TimeHistory(Callback):
def on_train_begin(self, logs={}):
self.times = []
def on_epoch_begin(self, epoch, logs={}):
self.epoch_time_start = time()
def on_epoch_end(self, epoch, logs={}):
self.times.append(time() - self.epoch_time_start)
# + id="3jaUXGvEzner" outputId="e4cdec25-1c67-488d-c234-4a4fc29c9332" papermill={"duration": 3285.083742, "end_time": "2020-10-18T09:44:18.933063", "exception": false, "start_time": "2020-10-18T08:49:33.849321", "status": "completed"} tags=[]
hidden_units = [[2048, 512, 2048],
[512, 1024, 512],
[512, 1024, 2048, 1024, 512],
[1024, 1024]]
dropout_rate = [0.4, 0.3, 0.45, 0.3, 0.4]
size = int(np.ceil(0.8 * len(top_feats)))
res = train_targets.copy()
ss.loc[:, train_targets.columns] = 0
res.loc[:, train_targets.columns] = 0
N_STARTS = 3
for seed in range(N_STARTS):
split_cols = []
for _ in range(len(hidden_units) - 1):
split_cols.append(np.random.choice(top_feats, size))
for n, (tr, te) in enumerate(MultilabelStratifiedKFold(n_splits = 5, random_state = seed, shuffle = True).split(train_targets,train_targets)):
start_time = time()
x_tr = [train.values[tr][:,split_cols[0]],
train.values[tr][:,split_cols[1]],
train.values[tr][:,split_cols[2]]]
x_val = [train.values[te][:, split_cols[0]],
train.values[te][:, split_cols[1]],
train.values[te][:, split_cols[2]]]
y_tr, y_val = train_targets.astype(float).values[tr], train_targets.astype(float).values[te]
x_tt = [test_features.values[:, split_cols[0]],
test_features.values[:, split_cols[1]],
test_features.values[:, split_cols[2]]]
#Model creation
model = create_model(size, hidden_units, dropout_rate, 1e-3)
#Model callbacks
rlr = ReduceLROnPlateau(monitor = 'val_loss', factor = 0.1, patience = 3, verbose = 0,
min_delta = 1e-4, min_lr = 1e-5, mode = 'min')
ckp = ModelCheckpoint(f'split_nn.hdf5', monitor = 'val_loss', verbose = 0,
save_best_only = True, save_weights_only = True, mode = 'min')
es = EarlyStopping(monitor = 'val_loss', min_delta = 1e-4, patience = 10, mode = 'min',
baseline = None, restore_best_weights = True, verbose = 0)
tm = TimeHistory()
#Model fit
history = model.fit(x_tr, y_tr,
validation_data = (x_val, y_val),
epochs = 100,
batch_size = 128,
callbacks = [rlr, ckp, es,tm],
verbose = 1)
hist = pd.DataFrame(history.history)
model.load_weights(f'split_nn.hdf5')
#Add predictions
ss.loc[:, train_targets.columns] += model.predict(x_tt, batch_size = 128) #Submision
res.loc[te, train_targets.columns] += model.predict(x_val, batch_size = 128) #Given data validation
#Print info
print(f'[{str(datetime.timedelta(seconds = time() - start_time))[2:7]}] Split NN: Seed {seed}, Fold {n}:', hist['val_loss'].min())
#Cleaning
K.clear_session()
del model, history, hist
x = gc.collect()
#Final media division
ss.loc[:, train_targets.columns] /= ((n + 1) * N_STARTS)
res.loc[:, train_targets.columns] /= N_STARTS
# + papermill={"duration": 14.956512, "end_time": "2020-10-18T09:44:48.960229", "exception": false, "start_time": "2020-10-18T09:44:34.003717", "status": "completed"} tags=[]
print(f'Split NN OOF Metric: {log_loss_metric(train_targets, res)}')
res.loc[train['cp_type'] == 1, train_targets.columns] = 0
ss.loc[test['cp_type'] == 1, train_targets.columns] = 0
print(f'Split NN OOF Metric with postprocessing: {log_loss_metric(train_targets, res)}')
# + papermill={"duration": 15.19356, "end_time": "2020-10-18T09:45:17.979426", "exception": false, "start_time": "2020-10-18T09:45:02.785866", "status": "completed"} tags=[]
ss.to_csv('submission.csv', index = False)
| notebooks/split-nn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:.conda-develop] *
# language: python
# name: conda-env-.conda-develop-py
# ---
# %% [markdown]
# ## Imports
# %%
# %load_ext autoreload
# %autoreload 2
import logging
import pprint
import tqdm.notebook as tqdm
import helpers.hdbg as dbg
import helpers.henv as env
import helpers.hprint as prnt
# %%
print(env.get_system_signature()[0])
prnt.config_notebook()
dbg.init_logger(verbosity=logging.INFO)
_LOG = logging.getLogger(__name__)
| dev_scripts/old/linter/test/Test_process_jupytext.test1_end_to_end/input/test_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (MXNet 1.6 Python 3.6 CPU Optimized)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-2:429704687514:image/mxnet-1.6-cpu-py36
# ---
# + [markdown] Collapsed="false"
# # AutoGluon Tabular with SageMaker
#
# [AutoGluon](https://github.com/awslabs/autogluon) automates machine learning tasks enabling you to easily achieve strong predictive performance in your applications. With just a few lines of code, you can train and deploy high-accuracy deep learning models on tabular, image, and text data.
# This notebook shows how to use AutoGluon-Tabular with Amazon SageMaker by creating custom containers.
# + [markdown] Collapsed="false"
# ## Prerequisites
#
# If using a SageMaker hosted notebook, select kernel `conda_mxnet_p36`.
# + Collapsed="false"
import subprocess
# Make sure docker compose is set up properly for local mode
subprocess.run("./setup.sh", shell=True)
# -
# For Studio
subprocess.run("apt-get update -y", shell=True)
subprocess.run("apt install unzip", shell=True)
subprocess.run("pip install ipywidgets", shell=True)
# + Collapsed="false"
import os
import sys
import boto3
import sagemaker
from time import sleep
from collections import Counter
import numpy as np
import pandas as pd
from sagemaker import get_execution_role, local, Model, utils, s3
from sagemaker.estimator import Estimator
from sagemaker.predictor import Predictor
from sagemaker.serializers import CSVSerializer
from sagemaker.deserializers import StringDeserializer
from sklearn.metrics import accuracy_score, classification_report
from IPython.core.display import display, HTML
from IPython.core.interactiveshell import InteractiveShell
# Print settings
InteractiveShell.ast_node_interactivity = "all"
pd.set_option('display.max_columns', 500)
pd.set_option('display.max_rows', 10)
# Account/s3 setup
session = sagemaker.Session()
local_session = local.LocalSession()
bucket = session.default_bucket()
prefix = 'sagemaker/autogluon-tabular'
region = session.boto_region_name
role = get_execution_role()
client = session.boto_session.client(
"sts", region_name=region, endpoint_url=utils.sts_regional_endpoint(region)
)
account = client.get_caller_identity()['Account']
registry_uri_training = sagemaker.image_uris.retrieve('mxnet', region, version= '1.7.0', py_version='py3', instance_type='ml.m5.2xlarge', image_scope='training')
registry_uri_inference = sagemaker.image_uris.retrieve('mxnet', region, version= '1.7.0', py_version='py3', instance_type='ml.m5.2xlarge', image_scope='inference')
ecr_uri_prefix = account +'.'+'.'.join(registry_uri_training.split('/')[0].split('.')[1:])
# + [markdown] Collapsed="false"
# ### Build docker images
# + [markdown] Collapsed="false"
# Build the training/inference image and push to ECR
# + Collapsed="false"
training_algorithm_name = 'autogluon-sagemaker-training'
inference_algorithm_name = 'autogluon-sagemaker-inference'
# -
# First, you may want to remove existing docker images to make a room to build autogluon containers.
subprocess.run("docker system prune -af", shell=True)
# + Collapsed="false"
subprocess.run(f"/bin/bash ./container-training/build_push_training.sh {account} {region} {training_algorithm_name} {ecr_uri_prefix} {registry_uri_training.split('/')[0].split('.')[0]} {registry_uri_training}", shell=True)
subprocess.run("docker system prune -af", shell=True)
# -
subprocess.run(f"/bin/bash ./container-inference/build_push_inference.sh {account} {region} {inference_algorithm_name} {ecr_uri_prefix} {registry_uri_training.split('/')[0].split('.')[0]} {registry_uri_inference}", shell=True)
subprocess.run("docker system prune -af", shell=True)
# ### Alternative way of building docker images using sm-docker
# The new Amazon SageMaker Studio Image Build convenience package allows data scientists and developers to easily build custom container images from your Studio notebooks via a new CLI.
# Newly built Docker images are tagged and pushed to Amazon ECR.
#
# To use the CLI, you need to ensure the Amazon SageMaker execution role used by your Studio notebook environment (or another AWS Identity and Access Management (IAM) role, if you prefer) has the required permissions to interact with the resources used by the CLI, including access to CodeBuild and Amazon ECR. Your role should have a trust policy with CodeBuild.
#
# You also need to make sure the appropriate permissions are included in your role to run the build in CodeBuild, create a repository in Amazon ECR, and push images to that repository.
#
# See also: https://aws.amazon.com/blogs/machine-learning/using-the-amazon-sagemaker-studio-image-build-cli-to-build-container-images-from-your-studio-notebooks/
# +
###subprocess.run("pip install sagemaker-studio-image-build", shell=True)
# +
'''
training_repo_name = training_algorithm_name + ':latest'
training_repo_name
!sm-docker build . --repository {training_repo_name} \
--file ./container-training/Dockerfile.training --build-arg REGISTRY_URI={registry_uri_training}
inference_repo_name = inference_algorithm_name + ':latest'
inference_repo_name
!sm-docker build . --repository {inference_repo_name} \
--file ./container-inference/Dockerfile.inference --build-arg REGISTRY_URI={registry_uri_inference}
'''
# + [markdown] Collapsed="false"
# ### Get the data
# + [markdown] Collapsed="false"
# In this example we'll use the direct-marketing dataset to build a binary classification model that predicts whether customers will accept or decline a marketing offer.
# First we'll download the data and split it into train and test sets. AutoGluon does not require a separate validation set (it uses bagged k-fold cross-validation).
# + Collapsed="false"
# Download and unzip the data
subprocess.run(f"aws s3 cp --region {region} s3://sagemaker-sample-data-{region}/autopilot/direct_marketing/bank-additional.zip .", shell=True)
subprocess.run("unzip -qq -o bank-additional.zip", shell=True)
subprocess.run("rm bank-additional.zip", shell=True)
local_data_path = './bank-additional/bank-additional-full.csv'
data = pd.read_csv(local_data_path)
# Split train/test data
train = data.sample(frac=0.7, random_state=42)
test = data.drop(train.index)
# Split test X/y
label = 'y'
y_test = test[label]
X_test = test.drop(columns=[label])
# + [markdown] Collapsed="false"
# ##### Check the data
# + Collapsed="false"
train.head(3)
train.shape
test.head(3)
test.shape
X_test.head(3)
X_test.shape
# + [markdown] Collapsed="false"
# Upload the data to s3
# + Collapsed="false"
train_file = 'train.csv'
train.to_csv(train_file,index=False)
train_s3_path = session.upload_data(train_file, key_prefix='{}/data'.format(prefix))
test_file = 'test.csv'
test.to_csv(test_file,index=False)
test_s3_path = session.upload_data(test_file, key_prefix='{}/data'.format(prefix))
X_test_file = 'X_test.csv'
X_test.to_csv(X_test_file,index=False)
X_test_s3_path = session.upload_data(X_test_file, key_prefix='{}/data'.format(prefix))
# + [markdown] Collapsed="false"
# ## Hyperparameter Selection
#
# The minimum required settings for training is just a target label, `init_args['label']`.
#
# Additional optional hyperparameters can be passed to the `autogluon.tabular.TabularPredictor.fit` function via `fit_args`.
#
# Below shows a more in depth example of AutoGluon-Tabular hyperparameters from the example [Predicting Columns in a Table - In Depth](https://auto.gluon.ai/stable/tutorials/tabular_prediction/tabular-indepth.html). Please see [fit parameters](https://auto.gluon.ai/stable/_modules/autogluon/tabular/predictor/predictor.html#TabularPredictor) for further information. Note that in order for hyperparameter ranges to work in SageMaker, values passed to the `fit_args['hyperparameters']` must be represented as strings.
#
# ```python
# nn_options = {
# 'num_epochs': "10",
# 'learning_rate': "ag.space.Real(1e-4, 1e-2, default=5e-4, log=True)",
# 'activation': "ag.space.Categorical('relu', 'softrelu', 'tanh')",
# 'layers': "ag.space.Categorical([100],[1000],[200,100],[300,200,100])",
# 'dropout_prob': "ag.space.Real(0.0, 0.5, default=0.1)"
# }
#
# gbm_options = {
# 'num_boost_round': "100",
# 'num_leaves': "ag.space.Int(lower=26, upper=66, default=36)"
# }
#
# model_hps = {'NN': nn_options, 'GBM': gbm_options}
#
# init_args = {
# 'eval_metric' : 'roc_auc'
# 'label': 'y'
# }
#
# fit_args = {
# 'presets': ['best_quality', 'optimize_for_deployment'],
# 'time_limits': 60*10,
# 'hyperparameters': model_hps,
# 'hyperparameter_tune': True,
# 'search_strategy': 'skopt'
# }
#
#
# hyperparameters = {
# 'fit_args': fit_args,
# 'feature_importance': True
# }
# ```
# **Note:** Your hyperparameter choices may affect the size of the model package, which could result in additional time taken to upload your model and complete training. Including `'optimize_for_deployment'` in the list of `fit_args['presets']` is recommended to greatly reduce upload times.
#
# <br>
# + Collapsed="false"
# Define required label and optional additional parameters
init_args = {
'label': 'y'
}
# Define additional parameters
fit_args = {
# Adding 'best_quality' to presets list will result in better performance (but longer runtime)
'presets': ['optimize_for_deployment'],
}
# Pass fit_args to SageMaker estimator hyperparameters
hyperparameters = {
'init_args': init_args,
'fit_args': fit_args,
'feature_importance': True
}
tags = [{
'Key' : 'AlgorithmName',
'Value' : 'AutoGluon-Tabular'
}]
# + [markdown] Collapsed="false"
# ## Train
#
# For local training set `train_instance_type` to `local` .
# For non-local training the recommended instance type is `ml.m5.2xlarge`.
#
# **Note:** Depending on how many underlying models are trained, `train_volume_size` may need to be increased so that they all fit on disk.
# + Collapsed="false"
# %%time
instance_type = 'ml.m5.2xlarge'
#instance_type = 'local'
ecr_image = f'{ecr_uri_prefix}/{training_algorithm_name}:latest'
estimator = Estimator(image_uri=ecr_image,
role=role,
instance_count=1,
instance_type=instance_type,
hyperparameters=hyperparameters,
volume_size=100,
tags=tags)
# Set inputs. Test data is optional, but requires a label column.
inputs = {'training': train_s3_path, 'testing': test_s3_path}
estimator.fit(inputs)
# -
# ### Review the performance of the trained model
# +
from utils.ag_utils import launch_viewer
launch_viewer(is_debug=False)
# + [markdown] Collapsed="false"
# ### Create Model
# + Collapsed="false"
# Create predictor object
class AutoGluonTabularPredictor(Predictor):
def __init__(self, *args, **kwargs):
super().__init__(*args,
serializer=CSVSerializer(),
deserializer=StringDeserializer(), **kwargs)
# + Collapsed="false"
ecr_image = f'{ecr_uri_prefix}/{inference_algorithm_name}:latest'
if instance_type == 'local':
model = estimator.create_model(image_uri=ecr_image, role=role)
else:
#model_uri = os.path.join(estimator.output_path, estimator._current_job_name, "output", "model.tar.gz")
model_uri = estimator.model_data
model = Model(ecr_image, model_data=model_uri, role=role, sagemaker_session=session, predictor_cls=AutoGluonTabularPredictor)
# + [markdown] Collapsed="false"
# ### Batch Transform
# + [markdown] Collapsed="false"
# For local mode, either `s3://<bucket>/<prefix>/output/` or `file:///<absolute_local_path>` can be used as outputs.
#
# By including the label column in the test data, you can also evaluate prediction performance (In this case, passing `test_s3_path` instead of `X_test_s3_path`).
# + Collapsed="false"
output_path = f's3://{bucket}/{prefix}/output/'
# output_path = f'file://{os.getcwd()}'
transformer = model.transformer(instance_count=1,
instance_type=instance_type,
strategy='MultiRecord',
max_payload=6,
max_concurrent_transforms=1,
output_path=output_path)
transformer.transform(test_s3_path, content_type='text/csv', split_type='Line')
transformer.wait()
# + [markdown] Collapsed="false"
# ### Endpoint
# + [markdown] Collapsed="false"
# ##### Deploy remote or local endpoint
# + Collapsed="false"
instance_type = 'ml.m5.2xlarge'
#instance_type = 'local'
predictor = model.deploy(initial_instance_count=1,
instance_type=instance_type)
# + [markdown] Collapsed="false"
# ##### Attach to endpoint (or reattach if kernel was restarted)
# + Collapsed="false"
# Select standard or local session based on instance_type
if instance_type == 'local':
sess = local_session
else:
sess = session
# Attach to endpoint
predictor = AutoGluonTabularPredictor(predictor.endpoint_name, sagemaker_session=sess)
# + [markdown] Collapsed="false"
# ##### Predict on unlabeled test data
# + Collapsed="false"
results = predictor.predict(X_test.to_csv(index=False)).splitlines()
# Check output
threshold = 0.5
y_results = np.array(['yes' if float(i.split(",")[1]) > threshold else 'no' for i in results])
print(Counter(y_results))
# + [markdown] Collapsed="false"
# ##### Predict on data that includes label column
# Prediction performance metrics will be printed to endpoint logs.
# + Collapsed="false"
results = predictor.predict(test.to_csv(index=False)).splitlines()
# Check output
threshold = 0.5
y_results = np.array(['yes' if float(i.split(",")[1]) > threshold else 'no' for i in results])
print(Counter(y_results))
# + [markdown] Collapsed="false"
# ##### Check that classification performance metrics match evaluation printed to endpoint logs as expected
# + Collapsed="false"
threshold = 0.5
y_results = np.array(['yes' if float(i.split(",")[1]) > threshold else 'no' for i in results])
print("accuracy: {}".format(accuracy_score(y_true=y_test, y_pred=y_results)))
print(classification_report(y_true=y_test, y_pred=y_results, digits=6))
# + [markdown] Collapsed="false"
# ##### Clean up endpoint
# + Collapsed="false"
predictor.delete_endpoint()
# -
# ## Explainability with Amazon SageMaker Clarify
#
# There are growing business needs and legislative regulations that require explainations of why a model made a certain decision. SHAP (SHapley Additive exPlanations) is an approach to explain the output of machine learning models. SHAP values represent a feature's contribution to a change in the model output. SageMaker Clarify uses SHAP to explain the contribution that each input feature makes to the final decision.
# ##### Set parameters for SHAP calculation
# +
seed = 0
num_rows = 500
#Write a csv file used by SageMaker Clarify
test_explainavility_file = 'test_explainavility.csv'
train.head(num_rows).to_csv(test_explainavility_file, index=False, header=False)
test_explainavility_s3_path = session.upload_data(test_explainavility_file, key_prefix='{}/data'.format(prefix))
# -
# ##### Specify computing resources
# +
from sagemaker import clarify
model_name = estimator.latest_training_job.job_name
container_def = model.prepare_container_def()
session.create_model(model_name,
role,
container_def)
clarify_processor = clarify.SageMakerClarifyProcessor(role=role,
instance_count=1,
instance_type='ml.c4.xlarge',
sagemaker_session=session)
model_config = clarify.ModelConfig(model_name=model_name,
instance_type='ml.c5.xlarge',
instance_count=1,
accept_type='text/csv')
# -
# ##### Run a SageMaker Clarify job
# +
shap_config = clarify.SHAPConfig(baseline=X_test.sample(15, random_state=seed).values.tolist(),
num_samples=100,
agg_method='mean_abs')
explainability_output_path = 's3://{}/{}/{}/clarify-explainability'.format(bucket, prefix, model_name)
explainability_data_config = clarify.DataConfig(s3_data_input_path=test_explainavility_s3_path,
s3_output_path=explainability_output_path,
label='y',
headers=train.columns.to_list(),
dataset_type='text/csv')
predictions_config = clarify.ModelPredictedLabelConfig(probability_threshold=0.5)
clarify_processor.run_explainability(data_config=explainability_data_config,
model_config=model_config,
explainability_config=shap_config)
# -
# ##### View the Explainability Report
# You can view the explainability report in Studio under the experiments tab. If you're not a Studio user yet, as with the Bias Report, you can access this report at the following S3 bucket.
subprocess.run(f"aws s3 cp {explainability_output_path} . --recursive", shell=True)
# Global explanatory methods allow understanding the model and its feature contributions in aggregate over multiple datapoints. Here we show an aggregate bar plot that plots the mean absolute SHAP value for each feature.
subprocess.run(f"{sys.executable} -m pip install shap", shell=True)
# ##### Compute global shap values out of out.csv
shap_values_ = pd.read_csv('explanations_shap/out.csv')
shap_values_.abs().mean().to_dict()
num_features = len(train.head(num_rows).drop(['y'], axis = 1).columns)
import shap
shap_values = [shap_values_.to_numpy()[:,:num_features], shap_values_.to_numpy()[:,num_features:]]
shap.summary_plot(shap_values,
plot_type='bar',
feature_names=train.head(num_rows).drop(['y'], axis = 1).columns.tolist())
# The detailed summary plot below can provide more context over the above bar chart. It tells which features are most important and, in addition, their range of effects over the dataset. The color allows us to match how changes in the value of a feature effect the change in prediction. The 'red' indicates higher value of the feature and 'blue' indicates lower (normalized over the features).
shap.summary_plot(shap_values_[shap_values_.columns[20:]].to_numpy(),
train.head(num_rows).drop(['y'], axis = 1))
| AutoGluon_Tabular_SageMaker.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "skip"} toc=true
# <h1>**Table of Contents**<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Feature-engineering---quantifying-access-to-facilities" data-toc-modified-id="Feature-engineering---quantifying-access-to-facilities-1">Feature engineering - quantifying access to facilities</a></span><ul class="toc-item"><li><span><a href="#Read-one-of-the-shortlisted-properties" data-toc-modified-id="Read-one-of-the-shortlisted-properties-1.1">Read one of the shortlisted properties</a></span></li><li><span><a href="#Create-symbols-for-facilities" data-toc-modified-id="Create-symbols-for-facilities-1.2">Create symbols for facilities</a></span><ul class="toc-item"><li><span><a href="#Get-5-mile-extent-around-the-property-of-interest" data-toc-modified-id="Get-5-mile-extent-around-the-property-of-interest-1.2.1">Get 5 mile extent around the property of interest</a></span></li><li><span><a href="#Plot-house-and-buffer-on-map" data-toc-modified-id="Plot-house-and-buffer-on-map-1.2.2">Plot house and buffer on map</a></span></li></ul></li><li><span><a href="#Geocode-for-facilities" data-toc-modified-id="Geocode-for-facilities-1.3">Geocode for facilities</a></span><ul class="toc-item"><li><span><a href="#Groceries" data-toc-modified-id="Groceries-1.3.1">Groceries</a></span></li><li><span><a href="#Restaurants" data-toc-modified-id="Restaurants-1.3.2">Restaurants</a></span></li><li><span><a href="#Hospitals" data-toc-modified-id="Hospitals-1.3.3">Hospitals</a></span></li><li><span><a href="#Coffee-shops" data-toc-modified-id="Coffee-shops-1.3.4">Coffee shops</a></span></li><li><span><a href="#Bars" data-toc-modified-id="Bars-1.3.5">Bars</a></span></li><li><span><a href="#Gas-stations" data-toc-modified-id="Gas-stations-1.3.6">Gas stations</a></span></li><li><span><a href="#Shops-and-service" data-toc-modified-id="Shops-and-service-1.3.7">Shops and service</a></span></li><li><span><a href="#Travel-and-transport" data-toc-modified-id="Travel-and-transport-1.3.8">Travel and transport</a></span></li><li><span><a href="#Parks-and-outdoors" data-toc-modified-id="Parks-and-outdoors-1.3.9">Parks and outdoors</a></span></li><li><span><a href="#Education" data-toc-modified-id="Education-1.3.10">Education</a></span></li><li><span><a href="#Present-the-results-in-a-table" data-toc-modified-id="Present-the-results-in-a-table-1.3.11">Present the results in a table</a></span></li><li><span><a href="#Find-duration-to-commute-to-work" data-toc-modified-id="Find-duration-to-commute-to-work-1.3.12">Find duration to commute to work</a></span></li></ul></li></ul></li></ul></div>
# + [markdown] slideshow={"slide_type": "slide"}
# # Feature engineering - quantifying access to facilities
# Often, when shortlisting facilities buyers look for access to facilities such as groceries, restaurants, schools, emergency and health care in thier neighborhood. In this notebook, we use the `geocoding` module to search for such facilities and build a table for each property.
# + slideshow={"slide_type": "skip"}
import pandas as pd
import matplotlib.pyplot as plt
from pprint import pprint
# %matplotlib inline
from arcgis.gis import GIS
from arcgis.geocoding import geocode
from arcgis.features import Feature, FeatureLayer, FeatureSet, GeoAccessor, GeoSeriesAccessor
from arcgis.features import SpatialDataFrame
from arcgis.geometry import Geometry, Point
from arcgis.geometry.functions import buffer
from arcgis.network import RouteLayer
# + [markdown] slideshow={"slide_type": "skip"}
# Connect to GIS
# + slideshow={"slide_type": "skip"}
gis = GIS(profile='')
# + [markdown] slideshow={"slide_type": "skip"}
# ## Read one of the shortlisted properties
# + slideshow={"slide_type": "skip"}
prop_list_df = pd.read_csv('resources/houses_for_sale_att_filtered.csv')
prop_list_df.shape
# + slideshow={"slide_type": "skip"}
prop_list_df = pd.DataFrame.spatial.from_xy(prop_list_df, 'LONGITUDE','LATITUDE')
type(prop_list_df)
# + slideshow={"slide_type": "skip"}
prop1 = prop_list_df[prop_list_df['MLS']==18389440]
prop1
# + [markdown] slideshow={"slide_type": "skip"}
# ## Create symbols for facilities
# Get your symbols using this online tool: [http://esri.github.io/arcgis-python-api/tools/symbol.html](http://esri.github.io/arcgis-python-api/tools/symbol.html)
# + slideshow={"slide_type": "skip"}
house_symbol = {"angle":0,"xoffset":0,"yoffset":0,"type":"esriPMS","url":"http://static.arcgis.com/images/Symbols/Shapes/RedStarLargeB.png","contentType":"image/png","width":24,"height":24}
grocery_symbol = symbol = {"angle":0,"xoffset":0,"yoffset":0,"type":"esriPMS","url":"http://static.arcgis.com/images/Symbols/PeoplePlaces/Shopping.png","contentType":"image/png","width":12,"height":12}
hospital_symbol = {"angle":0,"xoffset":0,"yoffset":0,"type":"esriPMS","url":"http://static.arcgis.com/images/Symbols/SafetyHealth/Hospital.png","contentType":"image/png","width":24,"height":24}
coffee_symbol = {"angle":0,"xoffset":0,"yoffset":0,"type":"esriPMS","url":"http://static.arcgis.com/images/Symbols/PeoplePlaces/Coffee.png","contentType":"image/png","width":12,"height":12}
restaurant_symbol = {"angle":0,"xoffset":0,"yoffset":0,"type":"esriPMS","url":"http://static.arcgis.com/images/Symbols/PeoplePlaces/Dining.png","contentType":"image/png","width":12,"height":12}
bar_symbol = {"angle":0,"xoffset":0,"yoffset":0,"type":"esriPMS","url":"http://static.arcgis.com/images/Symbols/PeoplePlaces/Bar.png","contentType":"image/png","width":12,"height":12}
gas_symbol = {"angle":0,"xoffset":0,"yoffset":0,"type":"esriPMS","url":"http://static.arcgis.com/images/Symbols/Transportation/esriBusinessMarker_72.png","contentType":"image/png","width":12,"height":12}
shops_service_symbol={"angle":0,"xoffset":0,"yoffset":0,"type":"esriPMS","url":"http://static.arcgis.com/images/Symbols/PeoplePlaces/esriBusinessMarker_58_Red.png","contentType":"image/png","width":10,"height":10}
transport_symbol = {"angle":0,"xoffset":0,"yoffset":0,"type":"esriPMS","url":"http://static.arcgis.com/images/Symbols/Transportation/esriDefaultMarker_195_White.png","contentType":"image/png","width":15,"height":15}
professional_symbol = {"angle":0,"xoffset":0,"yoffset":0,"type":"esriPMS","url":"http://static.arcgis.com/images/Symbols/PeoplePlaces/esriBusinessMarker_64_Yellow.png","contentType":"image/png","width":10,"height":10}
parks_symbol = {"angle":0,"xoffset":0,"yoffset":0,"type":"esriPMS","url":"http://static.arcgis.com/images/Symbols/OutdoorRecreation/RestArea.png","contentType":"image/png","width":10,"height":10}
education_symbol = {"angle":0,"xoffset":0,"yoffset":0,"type":"esriPMS","url":"http://static.arcgis.com/images/Symbols/PeoplePlaces/Note.png","contentType":"image/png","width":10,"height":10}
arts_symbol = {"angle":0,"xoffset":0,"yoffset":0,"type":"esriPMS","url":"http://static.arcgis.com/images/Symbols/PeoplePlaces/LiveShow.png","contentType":"image/png","width":12,"height":12}
destination_symbol = {"angle":0,"xoffset":0,"yoffset":12,"type":"esriPMS","url":"http://static.arcgis.com/images/Symbols/Basic/RedStickpin.png","contentType":"image/png","width":24,"height":24}
fill_symbol = {"type": "esriSFS","style": "esriSFSNull",
"outline":{"color": [255,0,0,255]}}
fill_symbol2 = {"type": "esriSFS","style": "esriSFSNull",
"outline":{"color": [0,0,0,255]}}
route_symbol = {"type": "esriSLS","style": "esriSLSSolid",
"color": [0, 120, 255, 255],"width": 1.5}
# + [markdown] slideshow={"slide_type": "skip"}
# ### Get 5 mile extent around the property of interest
# + slideshow={"slide_type": "skip"}
paddress = prop1.ADDRESS + ", " + prop1.CITY + ", " + prop1.STATE
prop_geom_fset = geocode(paddress.values[0], as_featureset=True)
# + [markdown] slideshow={"slide_type": "skip"}
# Create an envelope around the property using its extent
# + slideshow={"slide_type": "skip"}
prop_geom = prop_geom_fset.features[0]
prop_geom.geometry
# + slideshow={"slide_type": "skip"}
prop_geom = prop_geom_fset.features[0]
prop_buffer = buffer([prop_geom.geometry],
in_sr = 102100, buffer_sr=102100,
distances=0.05, unit=9001)[0]
prop_buffer_f = Feature(geometry=prop_buffer)
prop_buffer_fset = FeatureSet([prop_buffer_f])
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Plot house and buffer on map
# + slideshow={"slide_type": "fragment"}
pdx_map = gis.map('Portland, OR')
pdx_map.basemap='gray'
pdx_map
# -
# 
# + slideshow={"slide_type": "skip"}
pdx_map.draw(prop_buffer_fset, symbol=fill_symbol2)
pdx_map.draw(prop_geom_fset, symbol=house_symbol)
# + [markdown] slideshow={"slide_type": "skip"}
# ## Geocode for facilities
# We use the ArcGIS Geocoding service to search for facilities around this house
#
# ### Groceries
# -
neighborhood_data_dict = {}
# + slideshow={"slide_type": "skip"}
groceries = geocode('groceries', search_extent=prop_buffer.extent,
max_locations=20, as_featureset=True)
neighborhood_data_dict['groceries'] = []
for place in groceries:
popup={"title" : place.attributes['PlaceName'],
"content" : place.attributes['Place_addr']}
pdx_map.draw(place.geometry, symbol=grocery_symbol, popup=popup)
neighborhood_data_dict['groceries'].append(place.attributes['PlaceName'])
# + [markdown] slideshow={"slide_type": "subslide"}
# We will geocode for the following facilities within the said `5` mile buffer.
#
# Groceries
# Restaurants
# Hospitals
# Coffee shops
# Bars
# Gas stations
# Shops and service
# Travel and transport
# Parks and outdoors
# Education
# + [markdown] slideshow={"slide_type": "skip"}
# ### Restaurants
# + slideshow={"slide_type": "subslide"}
pdx_map2 = gis.map('Portland, OR')
pdx_map2.basemap='gray'
pdx_map2
# + [markdown] slideshow={"slide_type": "skip"}
# 
# + [markdown] slideshow={"slide_type": "skip"}
# 
# + [markdown] slideshow={"slide_type": "skip"}
# 
# + slideshow={"slide_type": "skip"}
pdx_map2.draw(prop_buffer_fset, symbol=fill_symbol2)
pdx_map2.draw(prop_geom_fset, symbol=house_symbol)
# + slideshow={"slide_type": "skip"}
restaurants = geocode('restaurant', search_extent=prop_buffer.extent, max_locations=200)
neighborhood_data_dict['restauruants'] = []
for place in restaurants:
popup={"title" : place['attributes']['PlaceName'],
"content" : place['attributes']['Place_addr']}
pdx_map2.draw(place['location'], symbol=restaurant_symbol, popup=popup)
neighborhood_data_dict['restauruants'].append(place['attributes']['PlaceName'])
# + [markdown] slideshow={"slide_type": "skip"}
# ### Hospitals
# + slideshow={"slide_type": "skip"}
hospitals = geocode('hospital', search_extent=prop_buffer.extent, max_locations=50)
neighborhood_data_dict['hospitals'] = []
for place in hospitals:
popup={"title" : place['attributes']['PlaceName'],
"content" : place['attributes']['Place_addr']}
pdx_map2.draw(place['location'], symbol=hospital_symbol, popup=popup)
neighborhood_data_dict['hospitals'].append(place['attributes']['PlaceName'])
# + [markdown] slideshow={"slide_type": "skip"}
# ### Coffee shops
# + slideshow={"slide_type": "skip"}
coffees = geocode('coffee', search_extent=prop_buffer.extent, max_locations=50)
neighborhood_data_dict['coffees'] = []
for place in coffees:
popup={"title" : place['attributes']['PlaceName'],
"content" : place['attributes']['Place_addr']}
pdx_map2.draw(place['location'], symbol=coffee_symbol, popup=popup)
neighborhood_data_dict['coffees'].append(place['attributes']['PlaceName'])
# + [markdown] slideshow={"slide_type": "skip"}
# ### Bars
# + slideshow={"slide_type": "skip"}
bars = geocode('bar', search_extent=prop_buffer.extent, max_locations=50)
neighborhood_data_dict['bars'] = []
for place in bars:
popup={"title" : place['attributes']['PlaceName'],
"content" : place['attributes']['Place_addr']}
pdx_map2.draw(place['location'], symbol=bar_symbol, popup=popup)
neighborhood_data_dict['bars'].append(place['attributes']['PlaceName'])
# + [markdown] slideshow={"slide_type": "skip"}
# ### Gas stations
# + slideshow={"slide_type": "skip"}
gas = geocode('gas station', search_extent=prop_buffer.extent, max_locations=50)
neighborhood_data_dict['gas'] = []
for place in gas:
popup={"title" : place['attributes']['PlaceName'],
"content" : place['attributes']['Place_addr']}
pdx_map2.draw(place['location'], symbol=gas_symbol, popup=popup)
neighborhood_data_dict['gas'].append(place['attributes']['PlaceName'])
# + [markdown] slideshow={"slide_type": "skip"}
# ### Shops and service
# + slideshow={"slide_type": "skip"}
shops_service = geocode("",category='shops and service', search_extent=prop_buffer.extent, max_locations=50)
neighborhood_data_dict['shops'] = []
for place in shops_service:
popup={"title" : place['attributes']['PlaceName'],
"content" : place['attributes']['Place_addr']}
pdx_map2.draw(place['location'], symbol=shops_service_symbol, popup=popup)
neighborhood_data_dict['shops'].append(place['attributes']['PlaceName'])
# + [markdown] slideshow={"slide_type": "skip"}
# ### Travel and transport
# + slideshow={"slide_type": "skip"}
transport = geocode("",category='travel and transport', search_extent=prop_buffer.extent, max_locations=50)
neighborhood_data_dict['transport'] = []
for place in transport:
popup={"title" : place['attributes']['PlaceName'],
"content" : place['attributes']['Place_addr']}
pdx_map2.draw(place['location'], symbol=transport_symbol, popup=popup)
neighborhood_data_dict['transport'].append(place['attributes']['PlaceName'])
# + [markdown] slideshow={"slide_type": "skip"}
# ### Parks and outdoors
# + slideshow={"slide_type": "skip"}
parks = geocode("",category='parks and outdoors', search_extent=prop_buffer.extent, max_locations=50)
neighborhood_data_dict['parks'] = []
for place in parks:
popup={"title" : place['attributes']['PlaceName'],
"content" : place['attributes']['Place_addr']}
pdx_map2.draw(place['location'], symbol=parks_symbol, popup=popup)
neighborhood_data_dict['parks'].append(place['attributes']['PlaceName'])
# + [markdown] slideshow={"slide_type": "skip"}
# ### Education
# + slideshow={"slide_type": "skip"}
education = geocode("",category='education', search_extent=prop_buffer.extent, max_locations=50)
neighborhood_data_dict['education'] = []
for place in education:
popup={"title" : place['attributes']['PlaceName'],
"content" : place['attributes']['Place_addr']}
pdx_map2.draw(place['location'], symbol=education_symbol, popup=popup)
neighborhood_data_dict['education'].append(place['attributes']['PlaceName'])
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Present the results in a table
# + slideshow={"slide_type": "fragment"}
neighborhood_df = pd.DataFrame.from_dict(neighborhood_data_dict, orient='index')
neighborhood_df = neighborhood_df.transpose()
neighborhood_df
# + slideshow={"slide_type": "subslide"}
neighborhood_df.count().plot(kind='bar')
plt.title('Facilities within 5 miles of {}'.format(prop1.ADDRESS.values[0]))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Find duration to commute to work
# + slideshow={"slide_type": "skip"}
route_service_url = gis.properties.helperServices.route.url
route_service = RouteLayer(route_service_url, gis=gis)
# + slideshow={"slide_type": "skip"}
stops = [paddress.values[0], '309 SW 6th Ave #600, Portland, OR 97204']
from arcgis.geocoding import geocode, batch_geocode
stops_geocoded = batch_geocode(stops)
stops_geocoded = [item['location'] for item in stops_geocoded]
stops_geocoded2 = '{},{};{},{}'.format(stops_geocoded[0]['x'],stops_geocoded[0]['y'],
stops_geocoded[1]['x'],stops_geocoded[1]['y'])
stops_geocoded2
# + slideshow={"slide_type": "skip"}
modes = route_service.retrieve_travel_modes()['supportedTravelModes']
for mode in modes:
print(mode['name'])
# + slideshow={"slide_type": "skip"}
route_service.properties.impedance
# + [markdown] slideshow={"slide_type": "skip"}
# Calculate time it takes to get to work. Set start time as `8:00 AM` on Mondays. ArcGIS routing service will use historic averages, so we provide this time as `8:00 AM, Monday, June 4 1990` in Unix epoch time. Read more about this [here](https://developers.arcgis.com/rest/network/api-reference/route-synchronous-service.htm#ESRI_SECTION3_72F22EAF69BF4F6CB6076B583CEB4074)
# + slideshow={"slide_type": "fragment"}
route_result = route_service.solve(stops_geocoded2, return_routes=True,
return_stops=True, return_directions=True,
impedance_attribute_name='TravelTime',
start_time=644511600000,
return_barriers=False, return_polygon_barriers=False,
return_polyline_barriers=False)
# + slideshow={"slide_type": "fragment"}
route_length = route_result['directions'][0]['summary']['totalLength']
route_duration = route_result['directions'][0]['summary']['totalTime']
route_duration_str = "{}m, {}s".format(int(route_duration),
round((route_duration %1)*60,2))
print("route length: {} miles, route duration: {}".format(round(route_length,3),
route_duration_str))
# + slideshow={"slide_type": "skip"}
route_features = route_result['routes']['features']
route_fset = FeatureSet(route_features)
stop_features = route_result['stops']['features']
stop_fset = FeatureSet(stop_features)
route_pop_up = {'title':'Name',
'content':'Total_Miles'}
pdx_map2.draw(route_fset, symbol=route_symbol, popup=route_pop_up)
# -
pdx_map2.draw(stop_fset, symbol=destination_symbol)
| talks/GeoDevPDX2018/03_feature-engineering-neighboring-facilities.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Building a neural network
# + [markdown] slideshow={"slide_type": "slide"}
# ### Neural networks in code
#
# Building a neural network from scratch in code is no mean feat. Some people make a habit out of it, sometimes going so far as to [live code a neural net library in one hour](https://www.youtube.com/watch?v=o64FV-ez6Gw).
#
# Fortunately, Python [is batteries-included](https://xkcd.com/353/) and has a wonderful community of ML practitioners. There are a number of wonderful, fully-developed and well-supported libraries out there for building and training neural networks.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Keras
#
# [Keras](https://keras.io/) is a high-level deep learning library, built to be simple, clear, and easy to use. It's my favourite of the neural network packages. With it, you can build and train a neural network, on a CPU or GPU, very quickly.
#
# The API that Keras provides is its true strength. The heavy lifting -- backpropagation and gradient descent -- are provided by other, lower-level frameworks, such as TensorFlow or Theano.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Let's dive straight into an example
# + slideshow={"slide_type": "skip"}
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# +
# Generate some data
x = np.linspace(-2, 2, 1000).reshape(-1, 1)
y = x**2 + np.random.randn(*x.shape) * 0.2
# Visualise it in a scatter plot
plt.scatter(x, y, alpha=0.3)
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's pull a few pieces from Keras.
# -
from keras.models import Sequential
from keras.layers import Dense
# + [markdown] slideshow={"slide_type": "subslide"}
# Next, let's put together a tiny neural net, with only one hidden layer.
# + slideshow={"slide_type": "skip"}
np.random.seed(3)
# +
# A "sequential" model -- a feed-forward neural network
model = Sequential()
# Let's add two fully-connected layers : one hidden, one output
model.add(Dense(2, activation="relu", input_dim=1))
model.add(Dense(1, activation="linear"))
# + [markdown] slideshow={"slide_type": "subslide"}
# We're done defining the net itself. We now need to provide some information about the loss function and gradient descent options.
# -
# We'll use vanilla stochastic gradient descent, and the mean squared error
model.compile(optimizer="sgd", loss="mse")
# + [markdown] slideshow={"slide_type": "subslide"}
# We're ready to train the net.
# -
history = model.fit(x, y, epochs=30, verbose=0)
# + [markdown] slideshow={"slide_type": "subslide"}
# How did we do ?
# + slideshow={"slide_type": "skip"}
def plot_network_results(x=x, y=y, model=model, history=history):
"""Plot the predictions and the loss history for the model."""
plt.figure(figsize=(15, 4))
plt.subplot(121)
plt.scatter(x, y, alpha=0.3)
plt.plot(x, model.predict(x), lw=5, c="k")
plt.legend(["Prediction", "Data"])
plt.subplot(122)
plt.plot(history.history["loss"], lw=5)
plt.title("Loss")
plt.xlabel("Epochs")
# -
plot_network_results()
# + [markdown] slideshow={"slide_type": "fragment"}
# A tiny neural network, with one hidden layer containing two units, was able to learn a good approximation of our function.
# + [markdown] slideshow={"slide_type": "subslide"}
# We say "tiny", but how small is it, really ?
# -
model.summary()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Predicting commute times
#
# Without further ado, let's train a larger network on some more realistic data.
# + slideshow={"slide_type": "subslide"}
import pandas as pd
# Let's import our data
training_data = pd.read_csv("../data/train_data_processed.csv", index_col=[0])
test_data = pd.read_csv("../data/test_data_processed.csv", index_col=[0])
# + slideshow={"slide_type": "subslide"}
training_data.head()
# -
X_train, y_train = training_data.drop(["commute_time"], axis=1), training_data["commute_time"]
X_test, y_test = test_data.drop(["commute_time"], axis=1), test_data["commute_time"]
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Neural networks appreciate scaled data
#
# Because most of the "interesting action" in an activation function occurs near zero, the neural net often learns best when it is fed data that is around zero mean, and scaled to reasonable variance.
#
# [We learn our scaling parameters](https://sebastianraschka.com/faq/docs/scale-training-test.html) from the training data, and apply them to the test data, to ensure our scaling is the same on both datasets.
# + slideshow={"slide_type": "subslide"}
from sklearn.preprocessing import StandardScaler
# Define a standard scaler and learn the parameters of the training data
scaler = StandardScaler()
scaler.fit(X_train)
# Rescale our data
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
# + slideshow={"slide_type": "skip"}
# A quick demonstration of scaling
def demonstrate_scaling():
"""Generate some fake data and rescaling it, plotting before and after."""
# First, some raw data
x_orig = np.random.normal(-4, 2, (10000, 1))
# Let's scale it !
scaler = StandardScaler()
x_scaled = scaler.fit_transform(x_orig)
# Plot the results
plt.hist(x_orig, bins=50, alpha=0.7, label="Original dataset")
plt.hist(x_scaled, bins=50, alpha=0.7, label="Scaled dataset")
plt.axvline(0, c="k", label=None)
plt.legend()
# + slideshow={"slide_type": "subslide"}
demonstrate_scaling()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Now let's build a neural net
# + slideshow={"slide_type": "skip"}
np.random.seed(3)
# + slideshow={"slide_type": "subslide"}
# A single hidden layer net, trained with stochastic gradient descent
model = Sequential()
model.add(Dense(32, activation="relu", input_dim=X_train.shape[1]))
model.add(Dense(1, activation="linear"))
model.compile(loss="mse", optimizer="sgd")
history = model.fit(X_train, y_train, epochs=15, verbose=0, validation_data=(X_test, y_test))
# + [markdown] slideshow={"slide_type": "subslide"}
# ### We're going to need some way to see how well we did
# + slideshow={"slide_type": "-"}
def evaluate_net(model, history, X_test=X_test, y_test=y_test):
"""Plot a net's training history and calculate its RMSE."""
plt.semilogy(history.history["loss"], lw=3)
plt.semilogy(history.history["val_loss"], lw=3, alpha=0.7)
plt.title("Final RMSE : {:.2f}".format((model.evaluate(X_test, y_test, verbose=0))**0.5))
plt.xlabel("Epochs")
plt.ylabel("MSE loss")
plt.legend(["Train", "Test"])
# + slideshow={"slide_type": "subslide"}
evaluate_net(model, history)
# + [markdown] slideshow={"slide_type": "slide"}
# ### We need to go deeper
#
# Well, at least most of us *want* to. However, before we add more layers, there are a few things we need to think about :
#
# - learning rate
# - minibatch size
# - number of epochs
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Learning rate
#
# If your learning rate is too high, you can overshoot in gradient descent, never finding the minimum, or worse, diverging completely. If your learning rate is too low, you will take a very long time training.
# + slideshow={"slide_type": "skip"}
from keras.optimizers import SGD
from keras.callbacks import TerminateOnNaN
def train_net(lr=0.01, batch_size=32, epochs=20, verbose=0, large=False):
"""Fit a neural network, returning the trained model and its history."""
# Fix the random seed
np.random.seed(3)
# Start with a model with a single 32-neuron hidden layer
model = Sequential()
model.add(Dense(32, activation="relu", input_dim=X_train.shape[1]))
# If we want a much larger net, throw in a couple of 256-neuron layers
if large:
model.add(Dense(256, activation="relu"))
model.add(Dense(256, activation="relu"))
# The output layer is a single linear unit
model.add(Dense(1, activation="linear"))
# Train the model using vanilla stochastic gradient descent
model.compile(loss="mse", optimizer=SGD(lr))
history = model.fit(
X_train, y_train,
epochs=epochs,
batch_size=int(batch_size),
validation_data=(X_test, y_test),
callbacks=[TerminateOnNaN()],
verbose=verbose
)
# In the case of first-epoch termination due to NaN,
# fill in the val_loss if it was never evaluated
if "val_loss" not in history.history:
history.history["val_loss"] = [np.nan]
return model, history
def compare_nets(first_model, first_history, second_model, second_history):
"""Plot, side by side, the losses of two neural networks with a shared y axis."""
# Create a figure that's about page width
plt.figure(figsize=(15, 5))
# Plot both models side by side
ax = plt.subplot(121)
evaluate_net(first_model, first_history)
plt.subplot(122, sharey=ax)
evaluate_net(second_model, second_history)
# Remove redundant white space
plt.tight_layout()
# + slideshow={"slide_type": "subslide"}
lr_high_model, lr_high_history = train_net(lr=0.1)
lr_low_model, lr_low_history = train_net(lr=1e-6)
compare_nets(lr_high_model, lr_high_history, lr_low_model, lr_low_history)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Minibatches
#
# A minibatch is like training on a small part of your data at a time : you forwards-propagate your minibatch, backpropagate your errors, and adjust your weights using gradient descent. Then you do the same on another minibatch. The advantage is that you update your weights a lot more often than in a single minibatch.
# + slideshow={"slide_type": "subslide"}
# Slow cell !
mb_large_model, mb_large_history = train_net(batch_size=len(X_train)/2)
mb_small_model, mb_small_history = train_net(batch_size=8)
compare_nets(mb_large_model, mb_large_history, mb_small_model, mb_small_history)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Number of epochs
#
# At the beginning, the more you train, the better your model gets. At some point, however, your model begins to overfit -- it stops learning the general function that maps your inputs to your outputs, and it begins learning the small intricacies of your training set. At this point, your training loss is still decreasing, but the test loss begins increasing.
# + slideshow={"slide_type": "subslide"}
# Warning : very slow cell !
long_train_model, long_train_history = train_net(epochs=500, lr=0.003, large=True)
evaluate_net(long_train_model, long_train_history)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### There are many other things to consider
#
# - depth of the neural net
# - number of neurons in each layer
# - initial distribution of weights
# - choice of optimiser and error metric
# - regularisation, dropout, and batch normalisation
# + [markdown] slideshow={"slide_type": "fragment"}
# So what might a slightly more beefy, carefully-built net look like ?
# + slideshow={"slide_type": "subslide"}
from keras.layers import BatchNormalization, Activation
np.random.seed(3)
# This time, we'll have three hidden layers with 32 neurons each
# We'll also batch-normalise, regularise, initialise our weights carefully
# Finally, we're using a more sophisticated grad descent algorithm : Adam
model = Sequential()
model.add(Dense(32, kernel_initializer="he_normal", kernel_regularizer="l2", input_dim=X_train.shape[1]))
model.add(BatchNormalization())
model.add(Activation("relu"))
model.add(Dense(32, kernel_initializer="he_normal", kernel_regularizer="l2"))
model.add(BatchNormalization())
model.add(Activation("relu"))
model.add(Dense(32, kernel_initializer="he_normal", kernel_regularizer="l2"))
model.add(BatchNormalization())
model.add(Activation("relu"))
model.add(Dense(1, kernel_initializer="he_normal", kernel_regularizer="l2", activation="linear"))
model.compile(loss="mse", optimizer="adam")
# + [markdown] slideshow={"slide_type": "skip"}
# This network has a lot more going on than our previous ones...
#
# #### Kernel initialisation
#
# If you recall, when we build a neural net, we pick the weights randomly, so that during training, we can take steps in a direction that'll improve the predictive power of the network. "Randomly" can mean many different things, however. Over time, people have discovered a number of different random selection methods that may help the net train faster. The [method used here](http://arxiv.org/abs/1502.01852) is one that's well-suited to the RELU activation function. Making a good choice in random weight initialisation can speed up training; making a poor choice can stop training completely.
#
# #### Batch normalisation
#
# Batch normalisation does the same thing as we did with our input data -- normalises it to zero mean, unit variance -- but at each layer, before the activation function. We do it here for the same reasons : because it can speed up training and help deeper nets converge faster.
#
# #### Regularisation
#
# We've added some L2 regularisation to our weights. Without going into much detail, regularisation keeps the net from overfitting and speeds up convergence by ensuring that the weights stay small. Gradient descent will more easily be able to influence the direction of a small weights vector, allowing us to explore more of the parameter space when training.
#
# #### Adam
#
# [Adam](https://arxiv.org/abs/1412.6980), or *adaptive momentum*, builds upon a number of enhancements to standard gradient descent by adding a momentum term that smoothes things out. In practice, Adam is often a great choice.
# + slideshow={"slide_type": "subslide"}
model.summary()
# + slideshow={"slide_type": "subslide"}
# Warning : slow cell !
history = model.fit(X_train, y_train, epochs=150, batch_size=64, verbose=0, validation_data=(X_test, y_test))
evaluate_net(model, history)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Too Long; Didn't Train
#
# - There's no **single correct architecture** for a problem or dataset
# - A lot of decisions can be made **scientifically**, but others are still **intuition**
# - Neural nets can be very **powerful** models, but are **slow** to train
# - They don't naturally allow much introspection
#
# I strongly recommend reading through [Neural Networks and Deep Learning](http://neuralnetworksanddeeplearning.com) by [<NAME>](http://michaelnielsen.org/). It's a wonderfully written text on neural nets, and it's freely available online.
| 3_Neural_networks/2_building_a_net.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:analysis]
# language: python
# name: conda-env-analysis-py
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# ## Read timings
steptimes = pd.read_csv('../data/restartable/AllStepTimes', sep=' ', names='case n time'.split())
fsizes = pd.read_csv('../data/restartable/WfuFileSizes', sep='\s+', names='case fsize'.split())
pred1 = pd.read_csv('../data/restartable/GO1_nfchl_cpu_to_wall.txt', sep=' ', names='case time pred1'.split())
pred1.loc[pred1.pred1 < 0, 'pred1'] = pred1[pred1.pred1 > 0].median()['pred1']
pred2 = pd.read_csv('../data/restartable/GO2_nfchl_cpu_to_wall.txt', sep=' ', names='case time pred2'.split())
pred2.loc[pred2.pred2 < 0, 'pred2'] = pred2[pred2.pred2 > 0].median()['pred2']
pred2.case = pred2.case.str.replace('.xyz2_ang', '')
q = pd.merge(steptimes, fsizes, how='left')
assert set(np.isnan(q.fsize.values)) == set((False,)) # fsize complete
qq = pd.merge(q, pred1, how='left', on='case')
#assert (qq[qq.n == 1].time_x == qq[qq.n == 1].time_y).values.all() # Heinis times equal to Max' times
q = qq
del q['time_y']
qq = pd.merge(q, pred2, how='left', on='case')
#assert (qq[qq.n == 2].time_x == qq[qq.n == 2].time).values.all() # Heinis times equal to Max' times
del qq['time']
qq.columns = 'case n time fsize step1 step2'.split()
df = qq.copy()
df = df.dropna(how='any')
# ## placement
# +
def simple(df, limit, diskMBps, jobQTime, jobOverhead):
""" Submit jobs as is, one job per molecule, restart until converged. Fallback to 1day queue if one step does not fit. """
fallback = 24*60*60
# random order
molecules = df.case.unique().copy()
np.random.shuffle(molecules)
# estimate packages
walltime = 0
cputime = 0
upgraded = 0
for molecule in molecules:
s = df.query('case == @molecule').sort_values('n')
# initialise walltime limit with target limit
moleculelimit = limit
remaining = moleculelimit
stepsinjob = 0
for idx, gostep in s.iterrows():
assert(remaining > 0)
if gostep.time <= remaining:
remaining -= gostep.time
stepsinjob += 1
else:
cputime += moleculelimit
walltime += moleculelimit + jobOverhead + jobQTime
remaining = moleculelimit - gostep.time
if gostep.n != 1:
remaining -= gostep.fsize / diskMBps
if remaining < 0:
assert (moleculelimit != fallback) # otherwise, 1day is too short for longest GO step
# run would not fit into current limit
remaining += (fallback - limit)
if stepsinjob > 1:
cputime += moleculelimit
walltime += moleculelimit + jobOverhead + jobQTime
moleculelimit = fallback
stepsinjob = 1
cputime += (moleculelimit - remaining)
walltime += (moleculelimit - remaining) + jobOverhead + jobQTime
if moleculelimit == fallback:
upgraded += 1
if upgraded / len(molecules) > 0.25:
raise ValueError('Too many upgrades: %d from %d' % (upgraded, len(molecules)))
return cputime, walltime
simple(df, 1*60*60, 100, 60*60, 30)
# +
def ml(df, limit, diskMBps, jobQTime, jobOverhead):
""" Use 2nd step pred to estimate continuation. """
fallback = 24*60*60
# order by largest first step
molecules = df.copy().sort_values('step1').case.unique()
# estimate packages
walltime = 0
cputime = 0
upgraded = 0
for molidx, molecule in enumerate(molecules):
s = df.query('case == @molecule').sort_values('n')
moleculelimit = limit
remaining = moleculelimit
for idx, gostep in s.iterrows():
assert(remaining >= 0)
if gostep.n == 1 and gostep.step1 > limit:
moleculelimit = fallback
remaining = moleculelimit
predtime = gostep.step2
if gostep.n == 1:
predtime = gostep.step1
if predtime > remaining:
# restart due to estimate
cputime += moleculelimit - remaining
walltime += (moleculelimit - remaining) + jobOverhead + jobQTime
# check whether upgrade is required
remaining = moleculelimit - gostep.fsize / diskMBps - predtime
if remaining < 0:
moleculelimit = fallback
remaining = moleculelimit - gostep.fsize / diskMBps - gostep.time
if remaining < 0:
assert (moleculelimit != fallback)
remaining += fallback - limit
cputime += moleculelimit
walltime += moleculelimit + jobOverhead + jobQTime
moleculelimit = fallback
continue
if gostep.time > remaining:
# restart due to limit
cputime += moleculelimit
walltime += moleculelimit + jobOverhead + jobQTime
remaining = moleculelimit - gostep.fsize / diskMBps - predtime
if remaining < 0:
assert(moleculelimit != fallback)
moleculelimit = fallback
remaining = moleculelimit - gostep.fsize / diskMBps - gostep.time
if remaining < 0:
assert(moleculelimit != fallback)
remaining += fallback - limit
cputime += moleculelimit
walltime += moleculelimit + jobOverhead + jobQTime
moleculelimit = fallback
continue
remaining -= gostep.time
cputime += (moleculelimit - remaining)
walltime += (moleculelimit - remaining) + jobOverhead + jobQTime
if moleculelimit == fallback:
upgraded += 1
if upgraded / len(molecules) > 0.25:
raise ValueError('Too many upgrades: %d from %d' % (upgraded, len(molecules)))
return cputime, walltime
ml(df, 1*60*60, 100, 60*60, 30)
# -
# ## visualisation
# +
ideal_cpu = df.time.sum()
limits = np.linspace(0.2, 12, 10)*60*60
DISK_SPEED = 1000
QUEUEING_TIME = 60*60
JOB_OVERHEAD = 30
visdata = []
methods = {'simple': simple, 'ml': ml}
for limit in limits:
for methodname, methodfunc in methods.items():
try:
q = methodfunc(df, limit, DISK_SPEED, QUEUEING_TIME, JOB_OVERHEAD)
except:
continue
visdata.append({'method': methodname, 'cputime': q[0], 'walltime': q[1], 'limit': limit})
visdata = pd.DataFrame(visdata)
# -
# medium disk
ideal_cpu = df.time.sum()
fig, axs = plt.subplots(1, 2, sharex=True, figsize=(10, 5))
cpuplot, wallplot = axs
for name, group in visdata.groupby('method'):
cpuplot.plot(group.limit/60/60, ((group.cputime/ideal_cpu)-1)*100, label=name)
relative = visdata.pivot(columns='method', index='limit', values='walltime').reset_index()
for method in sorted(methods.keys()):
wallplot.plot(relative.limit/60/60, ((relative[method] / relative['ml'])-1)*100, label=method)
cpuplot.legend()
wallplot.legend()
wallplot.set_ylabel('Wall Time overhead [%]')
cpuplot.set_ylabel('CPU Time overhead [%s]')
# ### Parallel workers
# +
class Worker(object):
""" One allocatable performing one job."""
def __init__(self, limit, fallback, diskMBps, jobQTime, jobOverhead, decider):
self._cputime = 0
self._walltime = 0
self._limit = limit
self._fallback = fallback
self._disk = diskMBps
self._jobQ = jobQTime
self._jobS = jobOverhead
self._decider = decider
def next_molecule_at(self):
return self._walltime + self._cputime
@staticmethod
def simple(gostep, queue, remaining, attempt):
if attempt == 1:
return queue
if attempt == 2:
return queue
if attempt == 3:
return 'long'
@staticmethod
def ml(gostep, queue, remaining, attempt):
if gostep.n == 1:
if gostep.step1 > remaining:
return 'long'
else:
if attempt == 1:
return queue
else:
return 'long'
assert (gostep.n != 1)
if gostep.step2 > remaining:
if attempt == 1:
return 'restart'
else:
return 'long'
else:
return queue
def _place_step(self, gostep, remaining, queue, restart, attempt):
planned_on = self._decider(gostep, queue, remaining, attempt)
if planned_on == 'restart':
restart = True
planned_on = queue
if planned_on != queue:
# decider wants to upgrade
restart = True
if restart:
# start a new job with this step
restart = False
self._walltime += self._jobQ
self._cputime += self._jobS
# IO component of restart
readwfntime = 0
if gostep.n != 1:
readwfntime = gostep.fsize / self._disk
self._cputime += readwfntime
queue = planned_on
if queue == 'short':
remaining = self._limit
else:
remaining = self._fallback
remaining -= readwfntime
if gostep.time < remaining:
remaining -= gostep.time
self._cputime += gostep.time
return True, remaining, queue, restart
else:
self._cputime += remaining
remaining = 0
restart = True
return False, remaining, queue, restart
def start_molecule(self, stepdata):
restart = True
queue = 'short'
remaining = self._limit
self._stepcount = 0
for _, gostep in stepdata.iterrows():
success, remaining, queue, restart = self._place_step(gostep, remaining, queue, restart, attempt=1)
if not success:
success, remaining, queue, restart = self._place_step(gostep, remaining, queue, restart, attempt=2)
if not success:
success, remaining, queue, restart = self._place_step(gostep, remaining, queue, restart, attempt=3)
assert(success)
return queue
def total_cputime(self):
return self._cputime
def do_run(hours, decider):
workers = [Worker(hours*60*60, 24*60*60, DISK_SPEED, QUEUEING_TIME, JOB_OVERHEAD, decider) for _ in range(NUMWORKERS)]
queues = []
for molecule in df.case.unique():
nextworker = min(workers, key=lambda x: x.next_molecule_at())
queues.append(nextworker.start_molecule(df.query('case == @molecule')))
#if len([_ for _ in queues if _ == 'long']) / len(queues) > 0.25:
# raise ValueError('Too many upgrades.')
walltime = max([_.next_molecule_at() for _ in workers])
cputime = sum([_.total_cputime() for _ in workers])
return cputime, walltime, len([_ for _ in queues if _ == 'long']), len([_ for _ in queues if _ == 'short'])
# +
ideal_cpu = df.time.sum()
limits = (1, 6, 12, 24)
DISK_SPEED = 1000
QUEUEING_TIME = 60*60
JOB_OVERHEAD = 30
NUMWORKERS = 10
visdata = []
methods = {'simple': Worker.simple, 'ml': Worker.ml}
for NUMWORKERS in (1, 4, 8, 16,):
for limit in limits:
for methodname, methodfunc in methods.items():
try:
q = do_run(limit, methodfunc)
except:
continue
visdata.append({'method': methodname, 'cputime': q[0], 'walltime': q[1], 'limit': limit, 'longs': q[2], 'shorts': q[3], 'workers': NUMWORKERS})
visdata = pd.DataFrame(visdata)
# -
ideal_cpu = df.time.sum()
fig, axs = plt.subplots(1, 2, sharex=True, figsize=(6, 3))
cpuplot, wallplot = axs
for name, group in visdata.query('workers == 4.').groupby('method'):
cpuplot.plot(group.limit, ((group.cputime/ideal_cpu)-1)*100, label={'ml': 'ML', 'simple': 'simple'}[name])
for workers in visdata.workers.unique():
relative = visdata.query('workers == @workers').pivot(columns='method', index='limit', values='walltime').reset_index()
if workers > 20:
continue
for method in sorted(methods.keys()):
if method == 'ml':
continue
if workers > 1:
pl = 's'
else:
pl = ''
wallplot.plot(relative.limit, (relative[method] / relative['ml']), label='%d core%s' % (workers, pl))
cpuplot.legend()
wallplot.legend()
wallplot.set_ylabel('Walltime / ML Walltime', fontsize=15)
cpuplot.set_ylabel('CPU Time Overhead [%]', fontsize=15)
cpuplot.set_xlabel('Walltime Limit [h]', fontsize=15)
wallplot.set_xlabel('Walltime Limit [h]', fontsize=15)
cpuplot.set_ylim(0,17)
wallplot.set_ylim(0.9,1.1)
wallplot.set_yticks([])
ax2 = wallplot.twinx()
ax2.set_ylim(0.9, 1.1)
ax2.set_yticks([0.9, 1.0, 1.1])
cpuplot.set_yticks([0, 5, 10, 15])
wallplot.tick_params('both', labelsize=15)
cpuplot.tick_params('both', labelsize=15)
ax2.tick_params('both', labelsize=15)
plt.tight_layout()
plt.savefig('gostepscheduling.pdf')
for name, group in visdata.query('workers == 4.').groupby('method'):
print (name)
print (group.cputime/ideal_cpu)
# ### scaling of estimate
def mlscaled(gostep, queue, remaining, attempt, scale):
if gostep.n == 1:
if gostep.step1*scale > remaining:
return 'long'
else:
if attempt == 1:
return queue
else:
return 'long'
assert (gostep.n != 1)
if gostep.step2*scale > remaining:
if attempt == 1:
return 'restart'
else:
return 'long'
else:
return queue
scaledata = []
NUMWORKERS = 4
for scale in np.arange(0.2, 5, 0.2):
try:
q = do_run(4, lambda g, q, r, a: mlscaled(g, q, r, a, scale))
except:
continue
scaledata.append({'scale': scale, 'cputime': q[0], 'walltime': q[1], 'longs': q[2], 'shorts': q[3]})
scaledata = pd.DataFrame(scaledata)
plt.plot(scaledata.scale, ((scaledata.cputime / scaledata.query('abs(scale-1.0) < 0.001').cputime.values[0])-1)*100, label='CPU')
plt.plot(scaledata.scale, ((scaledata.walltime / scaledata.query('abs(scale-1.0) < 0.001').walltime.values[0])-1)*100, label='WALL')
plt.axvline(x=1, color='lightgrey')
plt.xlabel('Scaling of time estimates')
plt.ylabel('% change relative to scale=1')
plt.legend()
scaledata.query('abs(scale-1.0) < 0.001')
| notebooks/InterruptionScheduling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/songqsh/MQP2019/blob/master/other/bsm_vanilla_pricing_v01.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="89t4J6Dmd-Eo"
# # BSM formula
#
# ## Abstract
#
# - create GBM class
# - define a method for BSM formula for a given option type
# + [markdown] colab_type="text" id="y5hyO8FseuLn"
# ## Analysis
#
# BS model assumes the distribution of stock as lognormal. In particular, it writes
# $$\ln \frac{S(T)}{S(0)} \sim \mathcal N((r - \frac 1 2 \sigma^2) T, \sigma^2 T)$$
# with respect to risk neutral measure. In the above, the parameters stand for
#
# * $S(0)$: The initial stock price
# * $S(T)$: The stock price at $T$
# * $r$: interest rate
# * $\sigma$: volatility
#
#
# + [markdown] colab_type="text" id="4BEWnmSve9oM"
#
# The call and put price with maturity $T$ and $K$ will be known as $C_0$ and $P_0$ given as below:
# $$C_0 = \mathbb E [e^{-rT} (S(T) - K)^+] = S_0 \Phi(d_1) - K e^{-rT} \Phi(d_2),$$
# and
# $$P_0 = \mathbb E [e^{-rT} (S(T) - K)^-] = K e^{-rT} \Phi(- d_2) - S_0 \Phi(- d_1),$$
# where $d_i$ are given as
# $$d_1 = ??,$$
# and
# $$d_2 = ??$$
#
# Put-call parity will be useful:
# $$C_0 - P_0 = S(0) - e^{-rT} K.$$
# + [markdown] colab_type="text" id="mewOxcQJfFnT"
# ## Code
# + colab={} colab_type="code" id="RXd_brmsfEs9"
import numpy as np
import scipy.stats as ss
# + [markdown] colab_type="text" id="M40EwMCkfS21"
# We reload the european option class created before.
# + colab={} colab_type="code" id="czvpqtvId_3D"
'''=========
option class init
=========='''
class VanillaOption:
def __init__(
self,
otype = 1, # 1: 'call'
# -1: 'put'
strike = 110.,
maturity = 1.,
market_price = 10.):
self.otype = otype
self.strike = strike
self.maturity = maturity
self.market_price = market_price #this will be used for calibration
def payoff(self, s): #s: excercise price
otype = self.otype
k = self.strike
maturity = self.maturity
return np.max([0, (s - k)*otype])
# + [markdown] colab_type="text" id="rdPRhkW0fhkn"
# Next, we create the gbm class, which is
# determined by three parameters. We shall initialize it
# as it is created.
# + colab={} colab_type="code" id="CQbFAFX-fYuw"
'''============
Gbm class inherited from sde_1d
============='''
class Gbm:
def __init__(self,
init_state = 100.,
drift_ratio = .0475,
vol_ratio = .2
):
self.init_state = init_state
self.drift_ratio = drift_ratio
self.vol_ratio = vol_ratio
# + [markdown] colab_type="text" id="6qcWtlDCgAO9"
# BSM formula is given by a method of Gbm class with an input of an option.
# + colab={} colab_type="code" id="KTFuh0GIfpOW"
'''========
Black-Scholes-Merton formula.
=========='''
def bsm_price(self, vanilla_option):
s0 = self.init_state
sigma = self.vol_ratio
r = self.drift_ratio
otype = vanilla_option.otype
k = vanilla_option.strike
maturity = vanilla_option.maturity
d1 = 0. #??
d2 = 0. #??
return (otype * s0 * ss.norm.cdf(otype * d1) #line break needs parenthesis
- otype * np.exp(-r * maturity) * k * ss.norm.cdf(otype * d2))
Gbm.bsm_price = bsm_price
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="iDswnsxjf_h5" outputId="40d37db8-a36c-415b-98d6-7059ba77d99d"
'''===============
Test bsm_price
================='''
gbm1 = Gbm()
option1 = VanillaOption()
print('>>>>>>>>>>call value is ' + str(gbm1.bsm_price(option1)))
option2 = VanillaOption(otype=-1)
print('>>>>>>>>>>put value is ' + str(gbm1.bsm_price(option2)))
# + colab={} colab_type="code" id="BrvYN7v0gWK5"
| other/bsm_vanilla_pricing_v01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import jsonlines
import re
import requests
import pyarabic.araby as araby
# ### Preprocessing
# +
with jsonlines.open("highlight_assault.jsonl", "r") as f:
assault = list(f.iter())
def actor_spans(entry):
chunks = []
for s in entry['spans']:
if s['label'] in ['SOURCE_ACTOR', 'TARGET_ACTOR']:
chunk = entry['text'][s['start']:s['end']+1]
chunks.append(chunk)
return chunks
actor_spans(assault[55])
all_chunks = []
for entry in assault:
try:
c = actor_spans(entry)
if c:
all_chunks.extend(c)
except Exception as e:
print(e)
len(all_chunks)
len(set(all_chunks))
# +
# Get data from udpipe
def udpipe(string) :
# Prepping String
words = re.findall(u'[\u0600-\u06FF]+', string) #getting arabic characters
data = ' '.join(words)
pipe_base_url = 'http://lindat.mff.cuni.cz/services/udpipe/api/process?tokenizer&tagger'
attributes = {}
attributes['model'] = 'arabic-ud-2.0-170801'
attributes['data'] = data
data = requests.get(pipe_base_url , attributes)
result = data.json()['result'].split('\n')
udpipe_results = [re.findall(u'[\u0600-\u06FF]+', i) for i in result] # cleaning
udpipe_results = [i for i in udpipe_results if i !=[]]
return udpipe_results
def udpipe_reconstruct(original_text) :
udpipe_results = udpipe(original_text)
for i in range(1, len(udpipe_results)):
if len(udpipe_results[i]) == 1 : # composite verbs get the next two words
original_text =original_text.replace(udpipe_results[i][0] , '{} {}'.format(udpipe_results[i+1][1],udpipe_results[i+2][1]))
else :
original_text =original_text.replace(udpipe_results[i][0] , udpipe_results[i][1])
original_text = araby.strip_tashkeel(original_text)
return original_text
def master_reconstruct_input(text_input , input_type):
# type 0: none-rule just do just do udpipe_reconstruction
# type 1 : fix the rule + do udpipe_reconstruction
if input_type == 0:
return udpipe_reconstruct(text_input)
else :
return udpipe_reconstruct(rule(text_input))
# -
chunks_unique = set(all_chunks)
len(chunks_unique)
list(chunks_unique)[:10]
chunks_unique_udpipe = []
for i in chunks_unique :
chunks_unique_udpipe.append(master_reconstruct_input(i ,0))
len(chunks_unique_udpipe)
list(chunks_unique_udpipe)[:10]
with open('actors_txt_file.txt', 'w') as outputf :
outputf.write('<Sentences>\n')
for i in range(len(chunks_unique_udpipe )):
outputf.write('<Sentence date = "20000715" id="{}" source = "afp" sentence = "True">\n'.format(i))
outputf.write('<Text>\n{}\n</Text>\n'.format(chunks_unique_udpipe[i]))
outputf.write('</Sentence>\n')
outputf.write('</Sentences>')
# ### proprocess the actor_txt_file.txt using UP preprocessing, then run the actor_code_extraction code to get the following results
#
# #### each element is actors_results is a list of 3 elements : Sentence (original Text) [0] , Matched Text[1], Code[2]
actors_results = []
with open('out_actors' , 'r') as inputf:
for i in inputf :
actors_results.append(i.replace('\n','').split(','))
actors_results[:10]
# +
#Filter stuff
extracted_actors_with_code = []
extracted_actors_with_out_code =[]
for i in actors_results :
if i[2] != '---' :
extracted_actors_with_code.append(i)
else :
extracted_actors_with_out_code.append(i)
print('Total Number of actors processed : {}'.format(len(actors_results)))
print('Extracted Actors with code : {} | {}%'.format(len(extracted_actors_with_code) , len(extracted_actors_with_code)/len(actors_results)))
print('Extracted Actors with out code : {} | {}%'.format(len(extracted_actors_with_out_code) , len(extracted_actors_with_out_code)/len(actors_results)))
# -
| scripts_and_code/stage3/.ipynb_checkpoints/actors_and_agents_matching-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Query Post-translational Modifications present in 3D Structures of the PDB.
#
# Post-translational modifications (PTMs) modulate protein function. By mapping the locations of modified amino acid residues onto 3D protein structures, insights into the effect of PTMs can be obtained.
#
# In this notebook PTMs present in PDB structures can be queried by residId, psimodId, uniprotId, and structureChainId. See examples below.
#
# Reference:
#
# BioJava-ModFinder: identification of protein modifications in 3D structures from the Protein Data Bank (2017) Bioinformatics 33: 2047–2049. [doi: doi.org/10.1093/bioinformatics/btx101](https://doi.org/10.1093/bioinformatics/btx101)
# + code_folding=[]
from pyspark.sql import SparkSession
from pyspark.sql.functions import collect_set, collect_list, col, concat_ws
from ipywidgets import interact, IntSlider, widgets
from IPython.display import display
from mmtfPyspark.datasets import pdbToUniProt, pdbPtmDataset
import py3Dmol
# -
spark = SparkSession.builder.appName("QueryPdbPTM").getOrCreate()
# ## Query PTMs by an Identifier
field = widgets.Dropdown(options=['residId', 'psimodId','uniProtId','structureChainId','all'],description='Select field:')
selection = widgets.Textarea(description='Enter id(s):', value='AA0151')
# Select query field and enter a comma separated list of ids. Examples:
#
# residId: AA0151 (N4-(N-acetylamino)glucosyl-L-asparagine)
#
# psimodId: MOD:00046, MOD:00047, MOD:00048 (O-phospho-L-serine, O-phospho-L-threonine, O4'-phospho-L-tyrosine)
#
# uniProtId: P13569
#
# structureChainId: 4ZTN.C
display(field)
display(selection)
# ## Create query string
query = field.value + " IN " + str(selection.value.split(',')).replace("[",'(').replace("]",')').replace(" ", "")
print("Query: " + query)
# ## Read dataset of PTMs present in PDB
db_ptm = pdbPtmDataset.get_ptm_dataset()
print("Total number of PTM records: ", db_ptm.count())
# Filter by PTM identifiers
# + code_folding=[0, 5]
if field.value in ['residId','psimodId']:
df = db_ptm.filter(query)
print("Filtered by query:", query)
print("Number of PTMs matching query:", df.count())
else:
df = db_ptm
df.limit(5).toPandas()
# -
# ## Get PDB to UniProt Residue Mappings
# + active=""
# Download PDB to UniProt mappings and filter out residues that were not observed in the 3D structure.
# -
up = pdbToUniProt.get_cached_residue_mappings().filter("pdbResNum IS NOT NULL")
print("Number of PDB to UniProt mappings:", up.count())
# Filter by UniProtID or structureChainIds
# + code_folding=[0]
if field.value in ['uniProtId','structureChainId']:
up = up.filter(query)
print("Filtered by query: ", query)
print("Number of records matching query:", up.count())
# -
# Find the intersection between the PTM dataset and PDB to UniProt mappings
df = df.withColumnRenamed("pdbResNum","resNum")
st = up.join(df, (up.structureChainId == df.pdbChainId) & (up.pdbResNum == df.resNum)).drop("pdbChainId").drop("resNum")
# Show some sample data
hits = st.count()
print("Hits:", hits)
fraction = min(10/hits, 1.0)
st.sample(False, fraction).toPandas().head()
# ## Aggregate PTM data by chain-level
# + code_folding=[0]
st = st.groupBy("structureChainId","uniprotId").agg(collect_list("pdbResNum").alias("pdbResNum"), \
collect_list("residId").alias("residId"), \
collect_list("psimodId").alias("psimodId"), \
collect_list("ccId").alias("ccId"))
# -
# Convert aggregated data to Pandas and display some results
pst = st.toPandas()
pst.head()
# Setup custom visualization
# + code_folding=[0] jupyter={"source_hidden": true}
def view_modifications(df, cutoff_distance, *args):
def view3d(show_labels=True,show_bio_assembly=False, show_surface=False, i=0):
pdb_id, chain_id = df.iloc[i]['structureChainId'].split('.')
res_num = df.iloc[i]['pdbResNum']
lab1 = df.iloc[i]['residId']
lab2 = df.iloc[i]['psimodId']
lab3 = df.iloc[i]['ccId']
# print header
print ("PDB Id: " + pdb_id + " chain Id: " + chain_id)
# print any specified additional columns from the dataframe
for a in args:
if df.iloc[i][a]:
print(a + ": " + df.iloc[i][a])
mod_res = {'chain': chain_id, 'resi': res_num}
# select neigboring residues by distance
surroundings = {'chain': chain_id, 'resi': res_num, 'byres': True, 'expand': cutoff_distance}
viewer = py3Dmol.view(query='pdb:' + pdb_id, options={'doAssembly': show_bio_assembly})
# polymer style
viewer.setStyle({'cartoon': {'color': 'spectrum', 'width': 0.6, 'opacity':0.8}})
# non-polymer style
viewer.setStyle({'hetflag': True}, {'stick':{'radius': 0.3, 'singleBond': False}})
# style for modifications
viewer.addStyle(surroundings,{'stick':{'colorscheme':'orangeCarbon', 'radius': 0.15}})
viewer.addStyle(mod_res, {'stick':{'colorscheme':'redCarbon', 'radius': 0.4}})
viewer.addStyle(mod_res, {'sphere':{'colorscheme':'gray', 'opacity': 0.7}})
# set residue labels
if show_labels:
for residue, l1, l2, l3 in zip(res_num, lab1, lab2, lab3):
viewer.addLabel(residue + ": " + l1 + " " + l2 + " " + l3, \
{'fontColor':'black', 'fontSize': 10, 'backgroundColor': 'lightgray'}, \
{'chain': chain_id, 'resi': residue})
viewer.zoomTo(surroundings)
if show_surface:
viewer.addSurface(py3Dmol.SES,{'opacity':0.8,'color':'lightblue'})
return viewer.show()
s_widget = IntSlider(min=0, max=len(df)-1, description='Structure', continuous_update=False)
return interact(view3d, show_labels=True, show_bio_assembly=False, show_surface=False, i=s_widget)
# -
# ## Visualize Results
# Residues with reported modifications are shown in an all atom prepresentation as red sticks with transparent spheres. Each modified residue position is labeled by the PDB residue number and the type of the modification. Residues surrounding modified residue (within 6 A) are highlighted as yellow sticks. Small molecules within the structure are rendered as gray sticks.
view_modifications(pst, 6, 'uniprotId');
# Most PTMs occur at the protein surface. To visualize the surface, check the show_surface checkbox above.
spark.stop()
| notebooks/QueryPdbPTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib as mpl
from utils import plot
import matplotlib.pyplot as plt
import numpy as np
import _pickle as pkl
import scipy.stats as stats
import tensorflow as tf
import time
from ig_attack import IntegratedGradientsAttack
from utils import dataReader, get_session, integrated_gradients, softmax
from model import Model
tf.logging.set_verbosity(tf.logging.ERROR)
# -
X, y = dataReader()
n = 50
original_label = y[n]
test_image = X[n]
plt.rcParams["figure.figsize"]=8,8
print("Image ID: {}, Image Label : {}".format(n, y[n]))
# %matplotlib inline
plt.imshow(X[n,:,:,0], cmap='gray')
# +
tf.reset_default_graph()
sess = get_session()
model = Model(create_saliency_op = 'ig')
model_dir = 'models/nat_trained'
saver = tf.train.Saver()
checkpoint = tf.train.latest_checkpoint(model_dir)
saver.restore(sess, checkpoint)
# +
k_top = 200 #Recommended for ImageNet
eval_k_top = 100
num_steps = 100 #Number of steps in Integrated Gradients Algorithm (refer to the original paper)
attack_method = 'topK'
epsilon = 0.3 #Maximum allowed perturbation for each pixel
attack_steps = 300
attack_times = 1
alpha = 0.01
attack_measure = "kendall"
reference_image = np.zeros((28,28,1)) #Our chosen reference(the mean image)
module = IntegratedGradientsAttack(sess = sess, test_image = test_image,
original_label = original_label, NET = model,
attack_method = attack_method, epsilon = epsilon,
k_top = k_top, eval_k_top = eval_k_top, num_steps = num_steps,
attack_iters = attack_steps,
attack_times = attack_times,
alpha = alpha,
attack_measure = attack_measure,
reference_image = reference_image,
same_label = True)
# +
output = module.iterative_attack_once()
print('''For maximum allowed perturbation size equal to {}, the resulting perturbation size was equal to {}'''.format(epsilon, np.max(np.abs(test_image - module.perturbed_image))))
print('''{} % of the {} most salient pixels in the original image are among {} most salient pixels of the
perturbed image'''.format(output[0]*100,eval_k_top,eval_k_top))
print("The Spearman rank correlation between salieny maps is equal to {}".format(output[1]))
print("The kendall rank correlation between salieny maps is equal to {}".format(output[2]))
# -
nat_output = sess.run(model.output_with_relu, feed_dict={model.input: [test_image]})
nat_pred = softmax(nat_output)
adv_output = sess.run(model.output_with_relu, feed_dict={model.input: [module.perturbed_image]})
adv_pred = softmax(adv_output)
print('original prediction: {}, confidence: {}'.format(np.argmax(nat_pred), np.max(nat_pred)))
print('perturbed prediction: {}, confidence: {}'.format(np.argmax(adv_pred), np.max(adv_pred)))
# +
original_IG = integrated_gradients(sess, reference_image, test_image, original_label, model, gradient_func='output_input_gradient', steps=num_steps)
mpl.rcParams["figure.figsize"]=8,8
plt.rc("text",usetex=False)
plt.rc("font",family="sans-serif",size=12)
saliency = np.sum(np.abs(original_IG),-1)
original_saliency = 28*28*saliency/np.sum(saliency)
plt.subplot(2,2,1)
plt.title("Original Image")
image = X[n,:,:,0]
plt.imshow(image, cmap='gray')
plt.subplot(2,2,2)
plt.title("Original Image Saliency Map")
plt.imshow(original_saliency, cmap="hot")
perturbed_IG = integrated_gradients(sess, reference_image, module.perturbed_image, original_label, model, gradient_func='output_input_gradient', steps=num_steps)
saliency = np.sum(np.abs(perturbed_IG),-1)
perturbed_saliency = 28*28*saliency/np.sum(saliency)
plt.subplot(2,2,3)
plt.title("Perturbed Image")
perturbed_image = (module.perturbed_image[:,:,0])
plt.imshow(perturbed_image, cmap='gray')
plt.subplot(2,2,4)
plt.title("Perturbed Image Saliency Map")
plt.imshow(perturbed_saliency, cmap="hot")
| MNIST/test_ig_attack.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Search in PTA
#
import json,csv,re
import unicodedata
import os,sys,glob
# MyCapytain == 2.0.9
from MyCapytain.resources.texts.local.capitains.cts import CapitainsCtsText
from MyCapytain.common.constants import Mimetypes, XPATH_NAMESPACES
from nltk.tokenize import RegexpTokenizer
import colored
from colored import stylize
import more_itertools as mit
# ## Functions
def convert_grcpta(files_path):
'''Read all greek files from files_path (without pta9999 = Bible) and convert to list of dictionaries (according subreferences)'''
xml_dir = os.path.expanduser(files_path)
xml_paths = glob.glob(xml_dir)
xml_paths = [path for path in sorted(xml_paths) if 'pta9999' not in path]
xml_paths = [path for path in sorted(xml_paths) if 'grc' in path]
pta_dict = []
for xml_path in xml_paths:
with open(xml_path, "r") as file_open:
plain_text = ""
_, xml_name = os.path.split(xml_path)
short_path = xml_path.split("/")
short_path = "/".join(short_path[8:])
urn = "".join(short_path[7:]).split(".xml")[0]
text = CapitainsCtsText(resource=file_open)
for ref in text.getReffs(level=len(text.citation)):
file_dict = {}
psg = text.getTextualNode(subreference=ref, simple=True)
psg.plaintext_string_join = ""
text_line = psg.export(Mimetypes.PLAINTEXT, exclude=["tei:note","tei:rdg"])
text_line = re.sub("\n","",text_line) # remove linebreaks
file_dict["id"] = urn+":"+ref
file_dict["text"] = text_line
pta_dict.append(file_dict)
return pta_dict
def tokenize_text(text):
'''Tokenize text by whitespace'''
word_breaks = RegexpTokenizer(r'\w+')
tokens = word_breaks.tokenize(text)
return tokens
# ### Search single word (regex)
def get_broader_context(files_path,urn,position,context_width):
'''Get broader context in text at position with context of context_width
Works with tokenized text, ngram = 1'''
texts = convert_grcpta(files_path)
entry = next((item for item in texts if item["id"] == urn), None)
mytext = tokenize_text(entry["text"])
context_before = [mytext[position-x] for x in range(context_width,0,-1)]
context_after = [mytext[position+x] for x in range(1,context_width+1)]
result = " ".join(context_before),'{:^10}'.format(stylize(mytext[position], colored.fg("blue")))," ".join(context_after)
result = " ".join(result)
return result
def search_single(tokens, search, context_width):
'''search single word, returns list of dictionaries with relative position of result and result'''
indices = [i for i, x in enumerate(tokens) if re.search(unicodedata.normalize("NFKC", search),x)]
count = 0
results = []
for entry in indices:
found = {}
count = count+1
try:
context_before = [tokens[entry-x] for x in range(context_width,0,-1)]
except:
context_before = []
try:
context_after = [tokens[entry+x] for x in range(1,context_width+1)]
except:
context_after = []
result = " ".join(context_before),'{:^10}'.format(stylize(tokens[entry], colored.fg("blue")))," ".join(context_after)
result = " ".join(result)
found["count"] = str(count)
found["pos"] = str(entry)
found["result"] = result
results.append(found)
return results
def search_word(word,context_width):
'''Search single word in texts, regex is allowed;
context at both sides = context_width'''
#texts = convert_grcpta(files_path)
for text in texts:
text_id = text["id"]
tokenized = text["text"]
tokens = tokenize_text(tokenized)
results = search_single(tokens, word, context_width)
for result in results:
print(stylize(text_id, colored.attr("bold"))+" (Ergebnis nr. "+result["count"]+" an Position "+result["pos"]+"): "+result["result"])
# ### Search list of words (regexes)
#
# Works also for single word, but output different from above
def generate_ngrams(words_list, n):
'''Generate ngrams of n length'''
ngrams_list = []
for num in range(0, len(words_list)):
ngram = ' '.join(words_list[num:num + n])
ngrams_list.append(ngram)
return ngrams_list
def search_words(list_of_words,distance):
'''Search a list of words in texts (regex is allowed) in files_path
within distance (ngram) number of words and give context of words at beginning and end'''
precompiled_list = [re.sub(" (\\\S\+ )+","|",x) for x in list_of_words] # case \S+ for words in between search
compiled_list = '(?:% s)' % '|'.join(precompiled_list)
#texts = convert_grcpta(files_path)
results = []
counted = 0
for text in texts:
text_id = text["id"]
ngramed = text["text"]
ngrams = generate_ngrams(tokenize_text(ngramed),distance)
numbers = len(ngrams)
res = [all([re.search(unicodedata.normalize("NFKC", k.lower()),s) for k in list_of_words]) for s in ngrams]
positions = [i for i in range(0, len(res)) if res[i]]
grouped_positions = [list(group) for group in mit.consecutive_groups(positions)]
found = {}
entry_results = []
for entry in grouped_positions:
# only the first entry to avoid overlap, alternative merge ngrams back to string
index = entry[0]
result_text = ngrams[index]
tokenized_result = tokenize_text(result_text)
emph_result = []
for word in tokenized_result:
# colorize search terms
if re.search(compiled_list, word):
emph_result.append(stylize(word, colored.attr("bold")))
else:
emph_result.append(word)
result_text = " ".join(emph_result)
try:
result_context_before = ngrams[index-distance]
except:
result_context_before = ""
try:
result_context_after = ngrams[index+distance]
except:
result_context_after = ""
result = "Position "+str(index)+"/"+str(numbers)+": "+result_context_before+" "+result_text+" "+result_context_after
entry_results.append(result)
counted = counted+1
found["id"] = text_id
found["results"] = entry_results
results.append(found)
print("Searched for "+" and ".join(list_of_words)+" within "+str(distance)+"-grams and found "+str(counted)+" results:")
for entry in results:
if entry["results"]:
print("===")
print(stylize(entry["id"], colored.attr("bold")))
for x in entry["results"]:
print("---")
print(x)
# # Search
#
# ## Examples for path
# - `~/Dokumente/projekte/First1KGreek/data/*/*/*.xml`
# - `~/Dokumente/projekte/pta_data/data/*/*/*.xml`
#
# ## Examples for search expressions
# - Search for words: `["[ἡἥ]λ[ίι].{1,2}$","οὐραν.*","καὶ"]`
# - Search for these consecutive words in distance of 2 words in between: `["καὶ \S+ \S+ οὐραν.*"]`
# ## Load corpus to be searched
#
# Needs to be done once
texts = convert_grcpta('~/Dokumente/projekte/pta_data/data/*/*/*.xml')
# ## Query
search_words(["[ἡἥ]λ[ίι].{1,2}$","οὐραν.*"],10)
search_word('~/Dokumente/projekte/pta_data/data/*/*/*.xml',"[ἡἥ]λ[ίι].{1,2}$", 10)
context = get_broader_context("pta0001.pta003.pta-grc1.xml",4881,100)
print(context)
| Search_PTA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#This notebook computes descriptive statistics and demographics of the COVID-19 Sounds dataset.
import pandas as pd
import numpy as np
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
pd.options.mode.chained_assignment = None
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_context("poster")
import matplotlib
# Say, "the default sans-serif font is Prox Nova"
matplotlib.rcParams['font.sans-serif'] = "Proxima Nova"
# Then, "ALWAYS use sans-serif fonts"
matplotlib.rcParams['font.family'] = "sans-serif"
# -
#data is separated by ":", the header/columns are in the first row, and row[1] is empty so we skip
web = pd.read_csv('web_data_processed.csv')
android = pd.read_csv('android_data_processed.csv')
ios = pd.read_csv('ios_data_processed.csv')
print ("Web data shape (users, variables):",web.shape)
print ("Android data shape (users, variables):",android.shape)
print ("IOS data shape (users, variables):",ios.shape)
#android, ios location is lat, lon, derived Country. Here we keep only the Country (last value after comma)
android['Location'] = android['Location'].str.rsplit(',').str[-1]
ios['Location'] = ios['Location'].str.rsplit(',').str[-1]
#TODO: convert web lat/lon to countries
data= pd.concat([web,android,ios],axis=0,sort=False,join='outer')
data = data.drop_duplicates()
data.shape
# ## Population-level statistics
#web data has miliseconds while android only seconds, therefore we strip the first 19 characters of the timestamp
data['date'] = pd.to_datetime(data['date_col'].astype(str).str[0:19], format ='%Y-%m-%d %H:%M:%S') #parse date (e.g. 2020-04-08-19_57_20_890867)
data.index = data['date']
#data=data[data.index > "2020-04-05"] #we launched the webform on the 6th of April 2020
del data['date_col'],data['date']# data["date"]
#find the first time that a user appeared
tmp = data.sort_values(['date']).drop_duplicates('Uid', keep='first')
# +
#cumulative hourly samples over time
series_samples = pd.Series(range(data.shape[0]), index=data.index) #a series with (range, index)
series_samples.resample('H').count().cumsum().plot(figsize=(10,5),lw=5, title="Evolution of reporting (cumulative, all platforms)", label='samples', color='goldenrod' ,) #resample hourly, apply cumulative sum, and plot
#cumulative hourly participants over time
series_users = pd.Series(range(tmp.shape[0]), index=tmp.index) #a series with (range, index)
series_users.resample('H').count().cumsum().plot(figsize=(10,5),lw=5, label='new users', color='cornflowerblue') #resample hourly, apply cumulative sum, and plot
#plt.savefig('evolution.png', bbox_inches='tight')
plt.legend()
plt.savefig('evolution.png', dpi=300, bbox_inches='tight')
# +
#data validation note: until 10/4/2020 (21:39pm) we were recording only yes/no, then was changed to never/last14/over14
data["Covid-Tested"][data["Covid-Tested"]=='never'] = 'Not tested'
data["Covid-Tested"][data["Covid-Tested"]=='neverThinkHadCOVIDNever'] = 'Not tested'
data["Covid-Tested"][data["Covid-Tested"]=='neverThinkHadCOVIDNow'] = 'Not tested (symptoms now)'
data["Covid-Tested"][data["Covid-Tested"]=='neverThinkHadCOVIDOver14'] = 'Not tested (symptoms past)'
data["Covid-Tested"][data["Covid-Tested"]=='neverThinkHadCOVIDLast14'] = 'Not tested (symptoms now)'
data["Covid-Tested"][data["Covid-Tested"]=='negativeNever'] = 'COVID-19 negative (always)'
data["Covid-Tested"][data["Covid-Tested"]=='negativeOver14'] = 'COVID-19 negative (past positive)'
data["Covid-Tested"][data["Covid-Tested"]=='negativeLast14'] = 'COVID-19 negative (recently positive)'
data["Covid-Tested"][data["Covid-Tested"]=='positiveOver14'] = 'COVID-19 positive (past)'
data["Covid-Tested"][data["Covid-Tested"]=='positiveLast14'] = 'COVID-19 positive'
data["Covid-Tested"][data["Covid-Tested"]=='last14'] = 'COVID-19 positive'
data["Covid-Tested"][data["Covid-Tested"]=='over14'] = 'COVID-19 positive (past)'
data["Covid-Tested"][data["Covid-Tested"]=='pnts'] = 'Prefer not to say'
data["Covid-Tested"][data["Covid-Tested"]=='ptns'] = 'Prefer not to say'
data["Covid-Tested"][data["Covid-Tested"]=='LocalizedStringKey(key: "pnts", hasFormatting: false, arguments: [])'] = 'Prefer not to say'
# -
data["Covid-Tested"].value_counts()
# +
from brokenaxes import brokenaxes
plt.figure(figsize=(15,8))
categs = data["Covid-Tested"].value_counts(normalize=False).index
y_pos = np.arange(len(categs))
#colors = ['silver','peachpuff','peachpuff','forestgreen','tomato', 'tomato', 'forestgreen', 'silver']
colors = ['silver','darkseagreen','peachpuff','darkseagreen',
'peachpuff', 'sandybrown', 'darkseagreen','silver', 'sandybrown']
bax = brokenaxes(xlims=((0, 13000), (32000, 35000)))
width = 0.85 # the width of the bars
bax.barh(y_pos, data["Covid-Tested"].value_counts().values, width, align='center', color=colors)
#this library has an issue with replacing ticks with your own labels
#this was so tricky to get it right (https://github.com/bendichter/brokenaxes/issues/17)
bax.set_yticklabels(categs)
bax.axs[0].set_yticks(y_pos)
#bax.invert_yaxis()
bax.set_xlabel('Samples', labelpad = 50)
plt.title("COVID-19 testing status")
plt.savefig('covid.png', dpi=300, bbox_inches='tight')
# -
# ## User-level statistics
#keep only one row per user (in order to calculate correct frequencies for age etc)
data = data.drop_duplicates(subset='Uid', keep="first")
data.shape
data.Age[data.Age=='16-19'] = '0-19'
data.Age[data.Age=='pnts'] = 'None'
data.Age.value_counts()
# +
plt.figure(figsize=(5,4)) #reordering the categories by reindexing the df
data.Age.value_counts().reindex(["0-19", "20-29", "30-39", "40-49", "50-59", "60-69", "70-79", "80-89","90-", "None"]).plot(kind='barh', rot=0) #value_counts(normalize=True) for percentages
plt.ylabel('Age')
plt.xlabel('Users')
plt.title("Age distribution")
plt.savefig('age.png', dpi=300, bbox_inches='tight')
# -
data.Sex[data.Sex=='ptns'] = 'Other/ \n None'
data.Sex[data.Sex=='pnts'] = 'Other/ \n None'
data.Sex[data.Sex=='Other'] = 'Other/ \n None'
data.Sex.value_counts(normalize=True)#.sum()
# +
plt.figure(figsize=(5,5))
plt.title("Sex distribution")
my_circle=plt.Circle( (0,0), 0.7, color='white')
plt.text(-0.5,0.2,'62%',rotation=0, size=20)
plt.text(0.1,-0.4,'36%',rotation=0, size=20)
plt.pie(data.Sex.value_counts().values, labels=data.Sex.value_counts().index.values, wedgeprops = { 'linewidth' : 1, 'edgecolor' : 'white' })
p=plt.gcf()
p.gca().add_artist(my_circle)
plt.savefig('sex.png', dpi=300, bbox_inches='tight')
plt.show()
# -
data.Smoking[data.Smoking=='ex'] = 'ex-smoker'
data.Smoking[data.Smoking=='1to10'] = '1-10 cigs'
data.Smoking[data.Smoking=='11to20'] = '11-20 cigs'
data.Smoking[data.Smoking=='ltOnce'] = '<1 cig'
data.Smoking[data.Smoking=='pnts'] = 'None'
data.Smoking[data.Smoking=='ptns'] = 'None'
data.Smoking[data.Smoking=='21+'] = '21+ cigs'
data.Smoking[data.Smoking=='ecig'] = 'e-cig'
data.Smoking.value_counts()
# +
plt.figure(figsize=(5,4))
data.Smoking.value_counts().plot(kind='barh')
plt.xlabel('Users')
plt.title("Smoking status")
plt.savefig('smoking.png', dpi=300, bbox_inches='tight')
# -
data.Location[data.Location=='unavailable'] = 'None'
data.Location[data.Location=='United Kingdom'] = 'UK'
data.Location[data.Location=='Argentina'] = 'Arg.'
data.Location[data.Location=='Portugal'] = 'Port.'
data.Location[data.Location=='Switzerland'] = 'Switz.'
data.Location[data.Location=='Sweden'] = 'Swed.'
data.Location[data.Location=='Russian Federation'] = 'Russia'
data.Location[data.Location==' Islamic Republic of'] = 'Iran' #the name Iran was lost because it was split by comma when we did str.rsplit
#countries with >60 users
top_countries = data.Location.value_counts().loc[lambda x : x>190]
data.Location.value_counts().sum()
top_countries
# +
#libraries
import squarify # pip install squarify (algorithm for treemap)
from matplotlib import cm
#14 colors
cs=cm.tab20(np.arange(14))
plt.figure(figsize=(12,5))
# Change color
squarify.plot(sizes=top_countries.values[1:], label=top_countries.index[1:], color=cs,alpha=.4 )
plt.axis('off')
plt.savefig('countries.png', dpi=300, bbox_inches='tight')
plt.show()
| Descriptive statistics/statistics_all_platforms.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <H1><center>pymgal example</center></H1>
import pymgal # import the model -- install requirements, numpy, astropy, pyfits, scipy
import numpy as np
import matplotlib
# %matplotlib inline
import pylab as pl
# ## 1. import SSP models
# + active=""
# Load simple stellar population models.
#
# model = SSP_model(model_file, IMF="chab", metal=[], is_ised=False, is_fits=False,
# is_ascii=False, has_masses=False, units='a', age_units='gyrs', nsample=None)
#
# model_file : File name for the SSP models. Only "name" + _ + "model",
# such as c09_exp, or bc03_ssp. The whole name of the model
# must be name_model_z_XXX_IMF.model (or .ised, .txt, .fits)
#
# IMF : Name of the Initial mass function. Default: "chab"
# Specify one of these IMF names: "chab", "krou" or "salp"
#
# metal : The metallicity of the model. Specify at here if you only
# want one or several metallicity included. Default: [],
# all metallicity models are included to do interpolation.
#
# nsample : The frequency sample points. Default: None, uses the model
# frequences. Otherwise, the interpolated frequency will be used.
# For the popurse of reducing required memory.
# Int number or list, numpy array. If int number, frequency will be
# assigned equally between min and max of model frequency.
# If list, or numpy array, it will be directly interpolated with
# the model frequency.
# Note, must be consistent with the units.
# -
mm=pymgal.SSP_models("bc03_ssp", metal=[0.008],nsample=1000)
# ## 2. load simulation data
# + active=""
# load analysing data from simulation snapshots (gadget format), yt, or raw data.
#
# data = load_data(snapname='', snapshot=False, yt_data=None,
# datafile=None, center=None, radius=None)
#
# -
simd=pymgal.load_data(snapname="/home/weiguang/Downloads/snap_127",snapshot=True,
center=[500000,500000,500000], radius=800)
# ### define projection direction and pixel size
simd.rotate_grid(axis='z',nx=128)
# ## 3. load filters
filters = pymgal.filters(f_name="sloan_r")
filters.add_filter("sloan_g")
filters.filter_order
# ## 4. Dust function before get seds
# + active=""
# Currently pymgal only has charlot_fall (2000) dust law.
# -
dustf=pymgal.dusts.charlot_fall()
# ## 5. Now get seds
seds=mm.get_seds(simd,dust_func=dustf)
seds_nodust = mm.get_seds(simd)
# ## calculate mag
mag = filters.calc_mag(mm.vs['0.008'],seds,z=0.1)
mag_nodust = filters.calc_mag(mm.vs['0.008'],seds_nodust,z=0.1)
print(mag['sloan_r'].shape, mag_nodust['sloan_r'].shape)
pl.figure(figsize=(9.3, 9))
pl.imshow(mag['sloan_r'].T,origin='lower')
pl.colorbar()
pl.figure(figsize=(9.3, 9))
pl.imshow(mag_nodust['sloan_r'].T,origin='lower')
pl.colorbar()
pl.figure(figsize=(8, 8))
pl.plot(simd.S_pos[:,0],simd.S_pos[:,1],'r,')
| doc/Test-pymgal.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **This notebook is an exercise in the [SQL](https://www.kaggle.com/learn/intro-to-sql) course. You can reference the tutorial at [this link](https://www.kaggle.com/dansbecker/group-by-having-count).**
#
# ---
#
# # Introduction
#
# Queries with **GROUP BY** can be powerful. There are many small things that can trip you up (like the order of the clauses), but it will start to feel natural once you've done it a few times. Here, you'll write queries using **GROUP BY** to answer questions from the Hacker News dataset.
#
# Before you get started, run the following cell to set everything up:
# Set up feedback system
from learntools.core import binder
binder.bind(globals())
from learntools.sql.ex3 import *
print("Setup Complete")
# The code cell below fetches the `comments` table from the `hacker_news` dataset. We also preview the first five rows of the table.
# +
from google.cloud import bigquery
# Create a "Client" object
client = bigquery.Client()
# Construct a reference to the "hacker_news" dataset
dataset_ref = client.dataset("hacker_news", project="bigquery-public-data")
# API request - fetch the dataset
dataset = client.get_dataset(dataset_ref)
# Construct a reference to the "comments" table
table_ref = dataset_ref.table("comments")
# API request - fetch the table
table = client.get_table(table_ref)
# Preview the first five lines of the "comments" table
client.list_rows(table, max_results=5).to_dataframe()
# -
# # Exercises
#
# ### 1) Prolific commenters
#
# Hacker News would like to send awards to everyone who has written more than 10,000 posts. Write a query that returns all authors with more than 10,000 posts as well as their post counts. Call the column with post counts `NumPosts`.
#
# In case sample query is helpful, here is a query you saw in the tutorial to answer a similar question:
# ```
# query = """
# SELECT parent, COUNT(1) AS NumPosts
# FROM `bigquery-public-data.hacker_news.comments`
# GROUP BY parent
# HAVING COUNT(1) > 10
# """
# ```
# +
# Query to select prolific commenters and post counts
prolific_commenters_query ="""
SELECT author, COUNT(1) AS NumPosts
FROM `bigquery-public-data.hacker_news.comments`
GROUP BY author
HAVING COUNT(1) > 10000
""" # Your code goes here
# Set up the query (cancel the query if it would use too much of
# your quota, with the limit set to 1 GB)
safe_config = bigquery.QueryJobConfig(maximum_bytes_billed=10**10)
query_job = client.query(prolific_commenters_query, job_config=safe_config)
# API request - run the query, and return a pandas DataFrame
prolific_commenters = query_job.to_dataframe()
# View top few rows of results
print(prolific_commenters.head())
# Check your answer
q_1.check()
# -
# For the solution, uncomment the line below.
q_1.solution()
# ### 2) Deleted comments
#
# How many comments have been deleted? (If a comment was deleted, the `deleted` column in the comments table will have the value `True`.)
# +
# Write your query here and figure out the answer
deleted_posts_query ="""SELECT COUNT(1) AS num_deleted_posts
FROM `bigquery-public-data.hacker_news.comments`
WHERE deleted = True
"""
query_job = client.query(deleted_posts_query)
# API request - run the query, and return a pandas DataFrame
deleted_posts = query_job.to_dataframe()
print(deleted_posts)
# +
num_deleted_posts = query_job.to_dataframe() # Put your answer here
# View results
print(deleted_posts)
# Check your answer
q_2.check()
# -
# For the solution, uncomment the line below.
q_2.solution()
# # Keep Going
# **[Click here](https://www.kaggle.com/dansbecker/order-by)** to move on and learn about the **ORDER BY** clause.
# ---
#
#
#
#
# *Have questions or comments? Visit the [Learn Discussion forum](https://www.kaggle.com/learn-forum/161314) to chat with other Learners.*
| exercise-group-by-having-count.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_tensorflow2_p36)
# language: python
# name: conda_tensorflow2_p36
# ---
# +
import pandas as pd
import numpy as np
np.set_printoptions(precision=6, suppress=True)
from sklearn.model_selection import train_test_split
from tqdm import tqdm
import tensorflow as tf
from tensorflow.keras import *
import tensorflow_addons as tfa
tf.__version__
# -
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
from matplotlib.ticker import (LinearLocator, MultipleLocator, FormatStrFormatter)
from matplotlib.dates import MONDAY
from matplotlib.dates import MonthLocator, WeekdayLocator, DateFormatter
from matplotlib import gridspec
from pandas.plotting import register_matplotlib_converters
register_matplotlib_converters()
# %matplotlib inline
# +
plt.rcParams['figure.figsize'] = ((8/2.54), (6/2.54))
plt.rcParams["font.family"] = "Arial"
plt.rcParams["mathtext.default"] = "rm"
plt.rcParams.update({'font.size': 11})
MARKER_SIZE = 15
cmap_m = ["#f4a6ad", "#f6957e", "#fccfa2", "#8de7be", "#86d6f2", "#24a9e4", "#b586e0", "#d7f293"]
cmap = ["#e94d5b", "#ef4d28", "#f9a54f", "#25b575", "#1bb1e7", "#1477a2", "#a662e5", "#c2f442"]
plt.rcParams['axes.spines.top'] = False
# plt.rcParams['axes.edgecolor'] =
plt.rcParams['axes.linewidth'] = 1
plt.rcParams['lines.linewidth'] = 1.5
plt.rcParams['xtick.major.width'] = 1
plt.rcParams['xtick.minor.width'] = 1
plt.rcParams['ytick.major.width'] = 1
plt.rcParams['ytick.minor.width'] = 1
# -
tf.config.list_physical_devices('GPU')
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
try:
for gpu in gpus:
tf.config.experimental.set_memory_growth(gpu, True)
logical_gpus = tf.config.experimental.list_logical_devices('GPU')
print(len(gpus), "Physical GPUs,", len(logical_gpus), "Logical GPUs")
except RuntimeError as e:
print(e)
strategy = tf.distribute.MirroredStrategy()
# # Hyperparameters
PRE_TRAINED = './models/Transformer.h5'
TRAINING_EPOCHS = 200
LEARNING_RATE = 0.001
EPSILON = 1e-06
BATCH_SIZE = 16
# # Data loading
l = np.load('./results/2020_W/fw_ct_dataset.npz', allow_pickle=True)
data_indices = l['data_indices']
input_data = l['input_data']
output_label = l['output_label']
INPUT_MAXS = l['INPUT_MAXS']
INPUT_MINS = l['INPUT_MINS']
OUTPUT_MAX = l['OUTPUT_MAX']
OUTPUT_MIN = l['OUTPUT_MIN']
input_data = input_data.astype('float32')
output_label = output_label.astype('float32')
print(input_data.shape)
print(output_label.shape)
print(INPUT_MAXS)
print(INPUT_MINS)
print(OUTPUT_MAX)
print(OUTPUT_MIN)
N_TRAIN = int(input_data.shape[0]*.09)
N_DEV = int(input_data.shape[0]/3)
TRAIN_INDEX = [_ for _ in range(N_TRAIN)] + \
[_ for _ in range(N_DEV, N_DEV+N_TRAIN)] + \
[_ for _ in range(N_DEV*2, N_DEV*2+N_TRAIN)]
TEST_INDEX = [_ for _ in range(input_data.shape[0]) if _ not in TRAIN_INDEX]
train_input = input_data[TRAIN_INDEX, ...]
train_label = output_label[TRAIN_INDEX, ...]
train_indices = data_indices[TRAIN_INDEX]
test_input = input_data[TEST_INDEX, ...]
test_label = output_label[TEST_INDEX, ...]
test_indices = data_indices[TEST_INDEX]
train_indices, val_indices, train_input, val_input, train_label, val_label = train_test_split(train_indices, train_input, train_label, test_size=0.3, shuffle=True, random_state=3101)
print(f'number of data set: {input_data.shape[0]}')
print(f'number of training set: {train_input.shape[0]}')
print(f'number of validation set: {val_input.shape[0]}')
print(f'number of test set: {test_input.shape[0]}')
with strategy.scope():
train_dataset = tf.data.Dataset.from_tensor_slices((train_input, train_label))
train_dataset = train_dataset.cache().shuffle(BATCH_SIZE*10).batch(BATCH_SIZE, drop_remainder=False)
val_dataset = tf.data.Dataset.from_tensor_slices((val_input, val_label))
val_dataset = val_dataset.cache().shuffle(BATCH_SIZE*10).batch(BATCH_SIZE, drop_remainder=False)
# # Model construction
class EmbeddingLayer(layers.Layer):
def __init__(self, num_nodes):
super(EmbeddingLayer, self).__init__()
self.n = num_nodes
self.dense = layers.Dense(self.n)
self.norm = layers.LayerNormalization(epsilon=1e-6)
def call(self, inp, is_train=True, **kwargs):
inp = self.dense(inp)
inp = self.norm(inp, training=is_train)
return inp
class EncoderBlock(layers.Layer):
def __init__(self, num_nodes, num_heads):
super(EncoderBlock, self).__init__()
self.n = num_nodes
self.h = num_heads
self.d = self.n // self.h
self.wq = layers.Dense(self.n)
self.wk = layers.Dense(self.n)
self.wv = layers.Dense(self.n)
self.dropout = layers.Dropout(0.1)
self.norm1 = layers.LayerNormalization(epsilon=1e-6)
self.dense1 = layers.Dense(self.n, activation=tf.nn.relu)
self.dense2 = layers.Dense(self.n)
self.norm2 = layers.LayerNormalization(epsilon=1e-6)
def head_maker(self, x, axis_1=2, axis_2=0):
x = tf.concat(tf.split(x, self.h, axis=axis_1), axis=axis_2)
return x
def call(self, inp, is_train=True, **kwargs):
Q = self.head_maker(self.wq(inp))
K = self.head_maker(self.wk(inp))
V = self.head_maker(self.wv(inp))
oup = tf.matmul(Q, tf.transpose(K, (0, 2, 1)))
oup = oup / tf.math.sqrt(tf.cast(K.shape[-1], tf.float32))
oup = tf.nn.softmax(oup)
oup = self.dropout(oup, training=is_train)
oup = tf.matmul(oup, V)
oup = self.head_maker(oup, 0, 2)
oup += inp
oup = self.norm1(oup, training=is_train)
oup_ffnn = self.dense1(oup)
oup_ffnn = self.dense2(oup_ffnn)
oup += oup_ffnn
oup = self.norm2(oup, training=is_train)
return oup
class TransformerLike(Model):
def __init__(self, num_nodes, num_heads, num_layers):
super(TransformerLike, self).__init__()
self.n = num_nodes
self.h = num_heads
self.l = num_layers
self.emb = EmbeddingLayer(self.n)
self.encs = [EncoderBlock(self.n, self.h) for _ in range(self.l)]
self.dense1 = layers.Dense(self.n, activation=tf.nn.relu)
self.dense2 = layers.Dense(self.n, activation=tf.nn.relu)
self.dense3 = layers.Dense(self.n, activation=tf.nn.relu)
self.flatten = layers.Flatten()
self.outdense1 = layers.Dense(32)
self.outdense2 = layers.Dense(1)
def call(self, inp, is_train=True, **kwargs):
inp = self.emb(inp)
for i in range(self.l):
inp = self.encs[i](inp, training=is_train)
inp = self.dense1(inp)
inp = self.dense2(inp)
inp = self.dense3(inp)
inp = self.outdense1(self.flatten(inp))
inp = self.outdense2(inp)
return inp
with strategy.scope():
model = TransformerLike(128, 4, 2)
with strategy.scope():
opt = optimizers.Adam(learning_rate=LEARNING_RATE, epsilon=EPSILON)
model.compile(optimizer=opt, loss='mae')
model.predict(val_dataset)
model.load_weights(PRE_TRAINED)
pred_output = model.predict(test_input)
pred_output = pred_output*(OUTPUT_MAX - OUTPUT_MIN) + OUTPUT_MIN
test_label = test_label*(OUTPUT_MAX - OUTPUT_MIN) + OUTPUT_MIN
# +
fig = plt.figure(figsize=((8.5/2.54*2), (6/2.54*2)))
ax0 = plt.subplot()
ax0.spines['right'].set_visible(False)
ax0.spines['left'].set_position(('outward', 5))
ax0.spines['bottom'].set_position(('outward', 5))
ax0.plot(test_label, pred_output, 'o', ms=5, mec='k', c=cmap[0])
fig.tight_layout()
# -
pred_df = pd.DataFrame(test_label, index=test_indices[:, 0], columns=['label'])
pred_df['pred'] = pred_output
# +
fig = plt.figure(figsize=((8.5/2.54*2), (6/2.54*2)))
ax0 = plt.subplot()
ax0.spines['right'].set_visible(False)
ax0.spines['left'].set_position(('outward', 5))
ax0.spines['bottom'].set_position(('outward', 5))
ax0.plot(pred_df.index, pred_df['label'], '-o', ms=5, mec='k', c=cmap[4])
ax0.plot(pred_df.index, pred_df['pred'], 'o', ms=5, mec='k', c=cmap[0])
fig.tight_layout()
# -
pred_df.to_csv('./results/model_output/transformer_pre.csv')
| 8-3_Transformer_pre.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# #Hi
#
import multiprocessing
from multiprocessing.managers import BaseManager
from queue import Queue
class QueueManager(BaseManager): pass
queue = Queue()
QueueManager.register('get_queue', callable=lambda:queue)
m = QueueManager(address=('0.0.0.0', 410), authkey=b'ADHM!@#')
s = m.get_server()
s.serve_forever()
# !curl ifconfig.me
from multiprocessing.managers import BaseManager
class QueueManager(BaseManager): pass
QueueManager.register('get_queue')
m = QueueManager(address=('172.16.31.10', 410), authkey=b'ADHM!@#')
m.connect()
queue = m.get_queue()
queue.get()
| literature_review.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://bit.ly/2VnXWr2" width="100" align="left">
#
# # The Snail and the Well
#
# A snail falls at the bottom of a 125 cm well. Each day the snail rises 30 cm. But at night, while sleeping, slides 20 cm because the walls are wet. How many days does it take for the snail to escape the well?
#
# **Hint**: The snail gets out of the well when it surpasses the 125cm of height.
#
# ## Tools
#
# 1. Loop: **while**
# 2. Conditional statements: **if-else**
# 3. Function: **print()**
#
# ## Tasks
#
# #### 1. Assign the challenge data to variables with representative names: `well_height`, `daily_distance`, `nightly_distance` and `snail_position`.
well_height = 125
daily_distance = 30
nightly_distance = 20
snail_position = 0
# #### 2. Create a variable `days` to keep count of the days that pass until the snail escapes the well.
days = 0
#
# #### 3. Find the solution to the challenge using the variables defined above.
# +
while well_height > snail_position:
snail_position = snail_position + daily_distance
days +=1
if well_height > snail_position:
snail_position = snail_position - nightly_distance
print('snail position', snail_position, 'cm', 'days', days)
else: break
print('snail position:', snail_position, 'cm', 'days', days)
# -
# #### 4. Print the solution.
print('left, after %s days' %days)
# ## Bonus
# The distance traveled by the snail each day is now defined by a list.
# ```
# advance_cm = [30, 21, 33, 77, 44, 45, 23, 45, 12, 34, 55]
# ```
# On the first day, the snail rises 30cm but during the night it slides 20cm. On the second day, the snail rises 21cm but during the night it slides 20cm, and so on.
#
# #### 1. How many days does it take for the snail to escape the well?
# Follow the same guidelines as in the previous challenge.
#
# **Hint**: Remember that the snail gets out of the well when it surpasses the 125cm of height.
advance_cm = [30, 21, 33, 77, 44, 45, 23, 45, 12, 34, 55]
well_height = 125
daily_distance = advance_cm
nightly_distance = 20
snail_position = 0
days = 0
daily_displacement = []
while well_height > snail_position:
for i in daily_distance :
snail_position = snail_position + i
daily_displacement.append(i-nightly_distance)
days += 1
if well_height > snail_position:
snail_position = snail_position - nightly_distance
print('snail position', snail_position, 'cm', 'days', days)
else: break
print('snail position:', snail_position, 'cm', 'days', days)
# #### 2. What is its maximum displacement in one day? And its minimum? Calculate the displacement using only the travel distance of the days used to get out of the well.
# **Hint**: Remember that displacement means the total distance risen taking into account that the snail slides at night.
import statistics
print(max(daily_displacement))
print(min(daily_displacement))
# #### 3. What is its average progress? Take into account the snail slides at night.
print(sum(daily_displacement)/len(daily_displacement))
print(statistics.mean(daily_displacement))
# #### 4. What is the standard deviation of its displacement? Take into account the snail slides at night.
print(statistics.stdev(daily_displacement))
| Python/1.-Snail-and-Well/snail-and-well.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Figuring out how to train SLAYER model using VAE method developed with Decolle and torchneuromorphic
# ### The problem:
# Training digit classifier(0-9) on a subset(1000 training and 100 testing) of NMNIST digit spikes recorded using DVS camera. For full training, use standard nmnist dataloaders
# ## Load proper paths for SLAYER Pytorch source modules¶
# +
import sys, os
CURRENT_TEST_DIR = os.getcwd()
sys.path.append(CURRENT_TEST_DIR + "slayerPytorch/src")
# -
os.environ["CUDA_VISIBLE_DEVICES"] = "0"
# ## Load required modules
# * SLAYER modules are available as `snn`.{`layer`, `params`, `loss`, `predict`, `io`}
# * SLAYER-Loihi module implements `spikeLayer` (defines custom Loihi neuron behaviour) and `quantizeWeights` (defines weight quantization scheme)
# * Optimizer implements custom NADAM optimizer
from torchneuromorphic import transforms
import importlib
from tqdm import tqdm
from torch import nn
from datetime import datetime
import numpy as np
import matplotlib.pyplot as plt
import torch
from torch.utils.data import Dataset, DataLoader
import slayerSNN as snn
import slayerSNN.auto
from slayerSNN import loihi as spikeLayer
from slayerSNN import quantize as quantizeParams
from slayerSNN import learningStats as learningStats
from slayerSNN import optimizer as optimizer
# from slayerLoihi import spikeLayer
# from quantizeParams import quantizeWeights
# from learningStats import learningStats
# import optimizer
import zipfile
from torchvision.utils import make_grid
# ## Define function for converting events to time surfaces
# +
def generate_process_target(aug_epoch=0):
tau1 = 1/(1-0.97)
tau2 = 1/(1-0.92)
print(transforms.ToCountFrame)
t1 = transforms.ExpFilterEvents(tau=tau2, length = int(6*tau2), tpad=int(6*tau2), device='cuda' )
t2 = transforms.ExpFilterEvents(tau=tau1, length = int(6*tau1), tpad=int(6*tau1), device='cuda' )
if aug_epoch<1:
filter_data = transforms.Compose([t1, t2, transforms.Rescale(50.)])
else:
filter_data = transforms.Compose([t1, t2, transforms.Rescale(50.), transforms.Jitter()])
def process_target(data, aug_epoch=0):
l = data.shape[1]
if aug_epoch > 0:
jitter_data = transforms.Compose([filter_data, transforms.Jitter(xs=1,ys=1,th=5)])
return jitter_data(data)[:,l]
return filter_data(data)[:,l]
return filter_data, process_target
filter_data, process_target = generate_process_target()
# -
# ## To use torchneuromorphic data with SLAYER
# SLAYER formats torch.Size([12, 2, 34, 34, 350]) i.e. (batch,p,x,y,t)
#
# This is different from torchneuromorphic format which uses torch.Size([12, 350, 2, 32, 32]) (batch,p,t,x,y) format (not sure if the order of x and y is correct)
#
# Therefore, to use torchneuromorphic with SLAYER, need to transpose along the correct dimensions to make the data usable by SLAYER type models
# ## Dataset Definition
# The dataset definition follows standard PyTorch dataset definition. Internally, it utilizes `snn.io` modules to read spikes and returns the spike in correct tensor format (`CHWT`).
# * `datasetPath`: the path where the spike files are stored.
# * `sampleFile`: the file that contains a list of sample indices and its corresponding clases.
# * `samplingTime`: the sampling time (in ms) to bin the spikes.
# * `sampleLength`: the length of the sample (in ms)
#
# Note: This is a simple dataset class. A dataset that utilizes the folder hierarchy or xml list is easy to create just like any other PyTorch dataset definition.
# Define the cuda device to run the code on.
device = torch.device('cuda')
# +
root_nmnist = 'data/dvsgesture/dvs_gestures.hdf5'
dataset = importlib.import_module('torchneuromorphic.dvs_gestures.dvsgestures_dataloaders')
try:
create_data = dataset.create_data
except AttributeError:
create_data = dataset.create_dataloader
# -
train_dl, test_dl = create_data(root=root_nmnist,
chunk_size_train=200,#1450,
chunk_size_test=200,#1450,
batch_size=100,#params['batch_size'],
dt=1000,#params['deltat'],
num_workers=12,
ds=4,
time_shuffle=True)#params['num_dl_workers'])
# +
data_batch, target_batch = next(iter(train_dl))
data_batch = data_batch[target_batch[:,-1,:].argmax(1)!=10]
data_batch = torch.Tensor(data_batch).to(device)
target_batch = torch.Tensor(target_batch).to(device)
# -
print(data_batch.shape)
print(target_batch.shape)
def transpose_torchneuromorphic_to_SLAYER(data_batch):
data_batch = torch.transpose(data_batch, 1,2)
data_batch = torch.transpose(data_batch,2,3)
data_batch = torch.transpose(data_batch,3,4)
# it looks like torchneuromorphic does y,x so switch them
data_batch = torch.transpose(data_batch,2,3)
return data_batch
data_batch_transposed = transpose_torchneuromorphic_to_SLAYER(data_batch)
def batch_one_hot(targets, num_classes=10):
one_hot = torch.zeros((targets.shape[0],num_classes))
#print("targets shape", targets.shape)
for i in range(targets.shape[0]):
one_hot[i][targets[i]] = 1
return one_hot
def three_unsqueeze(target_batch):
# use torch.unsqueeze to make slayer compatible with current error calculation.
# will probably want to use different error calculation method to conform to what I did in decolle
# but let's just get this working for now
target_batch = torch.unsqueeze(target_batch,-1)
target_batch = torch.unsqueeze(target_batch,-1)
target_batch = torch.unsqueeze(target_batch,-1)
return target_batch
# ## Describe the network
# Network is a standard VAE using an encoder and then a decoder.
# Because the goal is to put it on the Loihi, everything will need to use
# SLAYER with its Loihi simulator.
# In the decolle implementation we did not use spiking encoder head or classifiers,
# so I'm really hoping this will work ok using spikes...
# +
netDesc = {
'simulation' : {'Ts': 1, 'tSample': 300},#1450},
'neuron' : {
'type' : 'LOIHI',
'vThMant' : 80,#40,#80,
'vDecay' : 128,
'iDecay' : 1024,
'refDelay' : 1,
'wgtExp' : 0,
'tauRho' : 1,
'scaleRho' : 1024,#1024, # sumit said adjusting this could help with vanishing gradients. Loihi uses 128-1024
},
# 'layer' : [
# {'dim' : '32x32x2'},
# {'dim' : 512},
# {'dim' : 512},
# {'dim' : 10},
# ],
'training' : {
'error' : {
'type' : 'NumSpikes',
'tgtSpikeRegion': {'start': 0, 'stop': 1450}, # only valid for NumSpikes and ProbSpikes
'tgtSpikeCount': {'true': 180, 'false': 30}, # only valid for NumSpikes
}
}
}
netParams = snn.params(dict=netDesc)
# + tags=[]
headNetDesc = {
'simulation' : {'Ts': 1, 'tSample': 300},#1450},
'neuron' : {
'type' : 'LOIHI',
'vThMant' : 80,#40,#80,
'vDecay' : 3072,#4096,
'iDecay' : 1024,
'refDelay' : 1,
'wgtExp' : 0,
'tauRho' : 1,
'scaleRho' : 1024,#1024, # sumit said adjusting this could help with vanishing gradients. Loihi uses 128-1024
},
# 'layer' : [
# {'dim' : '32x32x2'},
# {'dim' : 512},
# {'dim' : 512},
# {'dim' : 10},
# ],
'training' : {
'error' : {
'type' : 'NumSpikes',
'tgtSpikeRegion': {'start': 0, 'stop': 1450}, # only valid for NumSpikes and ProbSpikes
'tgtSpikeCount': {'true': 180, 'false': 30}, # only valid for NumSpikes
}
}
}
headNetParams = snn.params(dict=headNetDesc)
# -
class Encoder(torch.nn.Module):
def __init__(self, netParams):
# since gradients are having trouble flowing through net
# maybe too big?
# try reducing layers and see if that works or something
super(Encoder, self).__init__()
# initialize slayer
slayer = spikeLayer(netParams['neuron'], netParams['simulation'])
self.slayer = slayer
# define network functions
# Encoder
self.conv1 = slayer.conv(2, 16, 5, padding=2, weightScale=10) # in, out, kernel, padding.
self.conv2 = slayer.conv(16, 32, 3, padding=1, weightScale=50)
self.pool1 = slayer.pool(1)
self.pool2 = slayer.pool(2)
self.pool3 = slayer.pool(2)
self.fc1 = slayer.dense((8*8*32), 512)
self.fc2 = slayer.dense(512, 128)
self.drop = slayer.dropout(0.1)
def forward(self, spikeInput):
# because of torchneuromorphic, don't need first pooling layer, probably
spike = self.slayer.spikeLoihi(self.pool1(spikeInput )) # 32, 32, 2
# with torch.no_grad():
# print(torch.sum(spike))
spike = self.slayer.delayShift(spike, 1)
#spike = self.drop(spike)
spike = self.slayer.spikeLoihi(self.conv1(spike)) # 32, 32, 16
# with torch.no_grad():
# print(torch.sum(spike))
spike = self.slayer.delayShift(spike, 1)
spike = self.slayer.spikeLoihi(self.pool2(spike)) # 16, 16, 16
# with torch.no_grad():
# print(torch.sum(spike))
spike = self.slayer.delayShift(spike, 1)
spike = self.drop(spike)
spike = self.slayer.spikeLoihi(self.conv2(spike)) # 16, 16, 32
# with torch.no_grad():
# print(torch.sum(spike))
spike = self.slayer.delayShift(spike, 1)
spike = self.slayer.spikeLoihi(self.pool3(spike)) # 8, 8, 32
spike = spike.reshape((spike.shape[0], -1, 1, 1, spike.shape[-1]))
# with torch.no_grad():
# print(torch.sum(spike))
spike = self.slayer.delayShift(spike, 1)
spike = self.drop(spike)
spike = self.slayer.spikeLoihi(self.fc1 (spike)) # 512
# with torch.no_grad():
# print(torch.sum(spike))
spike = self.slayer.delayShift(spike, 1)
spike = self.slayer.spikeLoihi(self.fc2 (spike)) # 11
# with torch.no_grad():
# print(torch.sum(spike))
spike = self.slayer.delayShift(spike, 1)
return spike
# mean and variance layer (encoder head)
# for beta=0 set leak term to max (voltage decay)
# try fp logvar
class EncoderHead(torch.nn.Module):
def __init__(self, netParams):
# since gradients are having trouble flowing through net
# maybe too big?
# try reducing layers and see if that works or something
super(EncoderHead, self).__init__()
# initialize slayer
slayer = spikeLayer(netParams['neuron'], netParams['simulation'])
self.slayer = slayer
self.fc_mu = slayer.dense(128, 100)
self.fc_logvar = slayer.dense(128,100)
def forward(self, spikeInput):
# put spiking encoder head, that will give mu logvar
# as spike rates. THese will be trained. Then the reparameterized
# z can be clustered, hopefully, on loihi. hell yeah let's gooooo
mu_spike = self.slayer.spikeLoihi( self.fc_mu(spikeInput))
with torch.no_grad():
print(torch.sum(mu_spike))
#mu_spike = self.slayer.delayShift(mu_spike, 1)
logvar_spike = self.slayer.spikeLoihi( self.fc_logvar(spikeInput))
#logvar_spike = self.slayer.delayShift(logvar_spike, 1)
with torch.no_grad():
print(torch.sum(logvar_spike))
return mu_spike , logvar_spike
# evaluate viability of using voltage for disentanglement
def get_voltage(self, spikeInput):
spike, current, mu_voltage = self.slayer.spikeLoihiFull(self.fc_mu(spikeInput))
spike, current, logvar_voltage = self.slayer.spikeLoihiFull(self.fc_logvar(spikeInput))
return mu_voltage, logvar_voltage
# meta sugur figures: log normal, get rid of labels for figs
# +
# For the rest of the VAE parts, in decolle I defined these parts using pytorch and they were not spiking
# this can be fine for initial testing, but if I want to put the encoder on loihi then I need
# to have the encoder_head and excitation classifier to be spiking
# not really sure how this would even work, it's kind of a glaring flaw that I somehow didn't see until now
# I guess I can see what kind of output a SLAYER encoder_head would produce, but then reparameterizing is a mystery
# or just use something that converts ANN to SNN and hope it's okay...
# +
class Reshape(nn.Module):
def __init__(self, *args):
super(Reshape, self).__init__()
self.shape = args
def forward(self, x):
return x.view(self.shape)
class VAE(torch.nn.Module):
def __init__(self, netParams,headParams,error,out_features=128,dimz=100,ngf=16,num_classes=10):
super(VAE, self).__init__()
self.dimz=dimz
self.error = error
self.encoder = Encoder(netParams)
self.encoder_head = EncoderHead(headParams)
# # non spiking encoder_head. This won't work on Loihi.
# self.encoder_head = nn.ModuleDict({'mu':nn.Linear(out_features, dimz),
# 'logvar':nn.Linear(out_features, dimz)})
# initialize encoder_head
# for l in self.encoder_head:
# #print(self.encoder_head[l])
# if isinstance(self.encoder_head[l], nn.Linear):
# torch.nn.init.normal_(self.encoder_head[l].weight)
#print("init")
# this is a pytorch decoder, non spiking. It's ok if this part is non spiking
# autograd should handle it
self.decoder = nn.Sequential(
nn.Linear(dimz, out_features),
Reshape(-1,out_features,1,1),
nn.ConvTranspose2d(out_features, ngf * 8, 4, 2, 0, bias=False),
nn.ReLU(True),
nn.ConvTranspose2d(ngf * 8, ngf * 4, 4, 2, 1, bias=False),
nn.ReLU(True),
nn.ConvTranspose2d(ngf * 4, ngf * 2, 4, 2, 1, bias=False),
nn.ReLU(True),
nn.ConvTranspose2d(ngf * 2, 2, 4, 2, 1, bias=False),
nn.ReLU())
#self.init_parameters()
from collections import OrderedDict
layer_size = 100
layer_size2 = 100 #300
layer_size3 = 100 #400
self.cls_sq = nn.Sequential(
OrderedDict([
('lin1', nn.Linear(num_classes,layer_size)),
('norm1', nn.BatchNorm1d(layer_size)),
('relu1', nn.LeakyReLU(negative_slope=0.2,inplace=True)),
('lin2', nn.Linear(layer_size,layer_size2)),
('norm2', nn.BatchNorm1d(layer_size2)),
('relu2', nn.LeakyReLU(negative_slope=0.2,inplace=True)),
('lin4', nn.Linear(layer_size3, num_classes))
])
)
# init model weights
for l in self.cls_sq:
if isinstance(l, nn.Linear):
torch.nn.init.kaiming_uniform_(l.weight, nonlinearity='leaky_relu')
# def init_parameters(self):
# self.encoder_head['logvar'].weight.data[:] *= 1e-16
# self.encoder_head['logvar'].bias.data[:] *= 1e-16
# return
def encode(self, x):
spike = self.encoder(x)
# spike = error.slayerPSP(spike)
# kernel = self.encoder.slayer.srmKernel
# maxKernel = torch.max(kernel).detach()
# spike = spike/maxKernel
# input into encoder_head
mu_spike, logvar_spike = self.encoder_head(spike)
mu_spike = self.error.slayerPSP(mu_spike) # post synaptic potential
logvar_spike = self.error.slayerPSP(logvar_spike)
kernel = self.encoder_head.slayer.srmKernel
maxKernel = torch.max(kernel).detach()
mu_spike = mu_spike/maxKernel
logvar_spike = logvar_spike/maxKernel
#print(mu_spike)
mu_spike = torch.squeeze(mu_spike,2)
mu_spike = torch.squeeze(mu_spike,2)
#mu_voltage = torch.mean(mu_voltage,2)
mu_spike = torch.transpose(mu_spike,0,2)
mu_spike = torch.transpose(mu_spike,1,2)
mu_spike = mu_spike[-1]
logvar_spike = torch.squeeze(logvar_spike,2)
logvar_spike = torch.squeeze(logvar_spike,2)
#logvar_spike = torch.mean(logvar_spike,2)
logvar_spike = torch.transpose(logvar_spike,0,2)
logvar_spike = torch.transpose(logvar_spike,1,2)
logvar_spike = logvar_spike[-1]
#print(type(mu))
#rint(mu_voltage
#h1 = torch.nn.functional.leaky_relu(spike)
# need to get membrane potential from mu and logvar instead of spieks
return mu_spike, logvar_spike
# see if voltage is usable for encoding. it might not be...
def encode_voltage(self, x):
spike = self.encoder(x)
mu_voltage, logvar_voltage = self.encoder_head.get_voltage(spike)
mu_voltage = torch.squeeze(mu_voltage,2)
mu_voltage = torch.squeeze(mu_voltage,2)
#mu_voltage = torch.mean(mu_voltage,2)
mu_voltage = torch.transpose(mu_voltage,0,2)
mu_voltage = torch.transpose(mu_voltage,1,2)
mu_voltage = mu_voltage[-1]
logvar_voltage = torch.squeeze(logvar_voltage,2)
logvar_voltage = torch.squeeze(logvar_voltage,2)
#logvar_spike = torch.mean(logvar_spike,2)
logvar_voltage = torch.transpose(logvar_voltage,0,2)
logvar_voltage = torch.transpose(logvar_voltage,1,2)
logvar_voltage = logvar_voltage[-1]
return mu_voltage, logvar_voltage
# for reparameterizing mu and logvar output by encoder_head
def reparameterize(self, mu, logvar):
std = torch.exp(0.5*logvar)
eps = torch.randn_like(std)
return mu + eps*std
def decode(self, z):
return self.decoder(z)
def excite_z(self,z,num_classes=10):
exc_z = torch.zeros((z.shape[0],num_classes))
for i in range(z.shape[0]):
exc_z[i] = z[i,:num_classes]#[t[i]]
return exc_z
def forward(self, x):
mu, logvar = self.encode(x)
# print(mu) # to see what mu is like
#logvar = torch.zeros(mu.shape).cuda()
#logvar = torch.ones(mu.shape).cuda()
#with torch.no_grad():
# print(mu)
# print(logvar)
z = self.reparameterize(mu, logvar)
clas = self.cls_sq(self.excite_z(z,10).cuda())
return self.decode(z), mu, logvar, clas
def z_voltage(self, x):
mu_voltage, logvar_voltage = self.encode_voltage(x)
z = mu_voltage #self.reparameterize(mu_voltage, logvar_voltage)
clas = self.cls_sq(self.excite_z(z,10).cuda())
return self.decode(z), mu_voltage, logvar_voltage, clas
# +
#print(net.encoder.slayer.srmKernel)
# -
class InhibNet(nn.Module):
def __init__(self, dimz, num_classes, hidden_layers):
super(InhibNet, self).__init__()
input_size = dimz-num_classes
output_size = num_classes
self.num_classes = num_classes
self.model = nn.Sequential(hidden_layers)
# init model weights
for l in self.model:
if isinstance(l, nn.Linear):
torch.nn.init.kaiming_uniform_(l.weight, nonlinearity='leaky_relu')
def forward(self, x):
x=self.model(x)
return x
def excite_z(self,z):
exc_z = torch.zeros((z.shape[0],self.num_classes))
for i in range(z.shape[0]):
exc_z[i] = z[i,:self.num_classes]#[t[i]]
return exc_z
def inhibit_z(self,z):
inhib_z = torch.zeros((z.shape[0], z.shape[1]-self.num_classes))
for i in range(z.shape[0]):
inhib_z[i] = z[i,self.num_classes:]
return inhib_z
out_features = 128
dimz = 100
ngf = 16
from collections import OrderedDict
layer_size = 100
inhib_layers = OrderedDict([
('lin1', nn.Linear(dimz-10,layer_size)),
('norm1', nn.BatchNorm1d(layer_size)),
('relu1', nn.LeakyReLU(negative_slope=0.2,inplace=True)),
('lin2', nn.Linear(layer_size,layer_size)),
('norm2', nn.BatchNorm1d(layer_size)),
('relu2', nn.LeakyReLU(negative_slope=0.2,inplace=True)),
#('droput',nn.Dropout(0.05)),
('lin3', nn.Linear(layer_size, 10))#,
#('soft', nn.LogSoftmax(dim=1))#nn.LogSoftmax(dim=1))
])
class InhibNet(nn.Module):
def __init__(self, dimz, num_classes, hidden_layers):
super(InhibNet, self).__init__()
input_size = dimz-num_classes
output_size = num_classes
self.num_classes = num_classes
self.model = nn.Sequential(hidden_layers)
# init model weights
for l in self.model:
if isinstance(l, nn.Linear):
torch.nn.init.kaiming_uniform_(l.weight, nonlinearity='leaky_relu')
def forward(self, x):
x=self.model(x)
return x
def excite_z(self,z):
exc_z = torch.zeros((z.shape[0],self.num_classes))
for i in range(z.shape[0]):
exc_z[i] = z[i,:self.num_classes]#[t[i]]
return exc_z
def inhibit_z(self,z):
inhib_z = torch.zeros((z.shape[0], z.shape[1]-self.num_classes))
for i in range(z.shape[0]):
inhib_z[i] = z[i,self.num_classes:]
return inhib_z
from collections import OrderedDict
layer_size = 100
inhib_layers = OrderedDict([
('lin1', nn.Linear(dimz-10,layer_size)),
('norm1', nn.BatchNorm1d(layer_size)),
('relu1', nn.LeakyReLU(negative_slope=0.2,inplace=True)),
('lin2', nn.Linear(layer_size,layer_size)),
('norm2', nn.BatchNorm1d(layer_size)),
('relu2', nn.LeakyReLU(negative_slope=0.2,inplace=True)),
#('droput',nn.Dropout(0.05)),
('lin3', nn.Linear(layer_size, 10))#,
#('soft', nn.LogSoftmax(dim=1))#nn.LogSoftmax(dim=1))
])
inhib = InhibNet(dimz,10,inhib_layers).cuda()
def vae_loss_fn(recon_x, x, mu, logvar, vae_beta = 4.0):
# not sure how well this will work with SLAYER, if at all, or if special modifications will need to be made
llhood = torch.nn.functional.mse_loss(recon_x, x)
#negKLD = -0.5 * torch.sum(1 + logvar - mu.pow(2) - logvar.exp())
kld_loss = torch.mean(-0.5 * torch.sum(1 + logvar - mu ** 2 - logvar.exp(), dim = 1), dim = 0)/len(train_dl)
#print(llhood,kld_loss)
return llhood + vae_beta*kld_loss
def guided_vae_loss_fn(recon_x, x, mu, logvar, excite_loss, inhib_loss, vae_beta = 4.0):
llhood = torch.nn.functional.mse_loss(recon_x, x)
#negKLD = -0.5 * torch.sum(1 + logvar - mu.pow(2) - logvar.exp())
kld_loss = torch.mean(-0.5 * torch.sum(1 + logvar - mu ** 2 - logvar.exp(), dim = 1), dim = 0)/len(train_dl)
return llhood+vae_beta*kld_loss+(excite_loss-inhib_loss)
criterion = nn.MultiLabelSoftMarginLoss(reduction='sum')
# ## Initialize the network
# * Define the device to run the code on.
# * Create network instance.
# * Create loss instance.
# * Define optimizer module.
# * Define training and testing dataloader.
# * Cereate instance for learningStats.
from decolle.utils import MultiOpt
from itertools import chain
# Create snn loss instance.
error = snn.loss(netParams, snn.loihi).to(device)
headError = snn.loss(headNetParams, snn.loihi).to(device)
# Create network instance.
#net = snn.auto.loihi.Network(netParams).to(device)
net = VAE(netParams,headNetParams,headError).to(device)
# +
# Define optimizer module.
opt1 = torch.optim.Adamax(net.encoder.parameters(), lr = 0.000001,betas=[0,0.95],eps=1e-4) #snn.utils.optim.Nadam(net.encoder.parameters(), lr = 0.0003, amsgrad = True) # lr=0.0003
opt2 = torch.optim.Adam(chain(*[net.encoder_head.parameters(),net.decoder.parameters()]), lr=0.000001) #snn.utils.optim.Nadam(chain(*[net.encoder_head.parameters(),net.decoder.parameters()]), lr=0.0003) # lr=0.0003
opt_excititory = torch.optim.Adam(net.cls_sq.parameters(), lr=0.000001) # lr=0.0001
opt_inhibitory = torch.optim.Adam(inhib.model.parameters(), lr=0.000001) # lr=0.0001
opt = MultiOpt(opt1, opt2, opt_excititory, opt_inhibitory)
# Learning stats instance.
stats = snn.utils.stats()
# -
# ## Train the network
# * Best network is stored for inferencing later
#
# ## Guided VAE
from utils import save_checkpoint, load_model_from_checkpoint
starting_epoch=0
# ### Training Loop
# for loading a model from checkpoint
starting_epoch = load_model_from_checkpoint('loihi_sim_model', net, opt, inhib, n_checkpoint=-1)
# +
# Guided VAE training with SLAYER
# The decolle vae quickly achieves decent accuracy(50% after 5 epochs) with decent reconstructions
# the slayer vae seems to be quickly converging to 16% accuracy and has uniform blob reconstructions
# the architecture should be the same between both. Is it the neuron and surrogate gradient differences?
# or some parameter needs changing? maybe network is not good for slayer, idk
# or maybe too much noise in the data?
losses = []
for epoch in range(5001):
tSt = datetime.now()
print(epoch)
# Training loop.
# use torchneuromophic dataloaders for training
sum_vae_loss = 0
for x,t in tqdm(iter(train_dl)):
net.train()
inhib.train()
# Reset gradients to zero.
opt.zero_grad()
new_t = t[t[:,-1,:].argmax(1)!=10]
# if new_t.shape[0] < 2:
# #print(new_t.shape)
# continue
new_t = new_t[:,-1,:].argmax(1)
#print(new_t.shape)
x = x[t[:,-1,:].argmax(1)!=10]
#print("t shape", new_t.shape)
x_c = x.to(device)
# get time surface for reconstruction
frames = process_target(x_c)#torch.transpose(x_c,3,4)) # make sure the image is not sideways
# process target and input to make SLAYER compatible
x_c = transpose_torchneuromorphic_to_SLAYER(x_c)
hot_ts = batch_one_hot(new_t, num_classes=10)
# Forward pass of the network.
y,mu,logvar,clas = net(x_c)
#print(y.shape)
#print(frames.shape)
loss = vae_loss_fn(y, frames, mu, logvar, vae_beta=1.2)
clas_loss = criterion(clas, hot_ts.to(device))
vae_loss = loss
loss += clas_loss
loss.backward()
# print("net grads first pass")
with torch.no_grad():
print("encoder")
print(net.encoder.conv1.weight.grad.abs().mean())
print(net.encoder.conv2.weight.grad.abs().mean())
# print(net.encoder.conv3.weight.grad.abs().mean())
# #print(net.encoder.conv4.weight.grad.abs().mean())
print(net.encoder.fc1.weight.grad.abs().mean())
# print(net.encoder.mu.weight.grad.abs().mean())
# print(net.encoder.logvar.weight.grad.abs().mean())
#print(net.encoder.fc2.weight.grad.abs().mean())
print("encoder_head")
print(net.encoder_head.fc_mu.weight.grad.abs().mean())
print(net.encoder_head.fc_logvar.weight.grad.abs().mean())
print("classifier last")
print(net.cls_sq.lin4.weight.grad.abs().mean())
opt.step()
opt.zero_grad()
z = net.reparameterize(mu,logvar).detach()
inhib_z = inhib.inhibit_z(z)
excite_output = inhib.model(inhib_z.to(device))
loss = criterion(excite_output, hot_ts.to(device))
excite_loss = loss
loss.backward()
opt.step()
opt.zero_grad()
#mu, logvar = net.encode(x_c)
mu, logvar = net.encode(x_c)
#logvar = torch.zeros(mu.shape).cuda()
z = net.reparameterize(mu, logvar)
inhib_z = inhib.inhibit_z(z)
inhib_output= inhib.model(inhib_z.to(device))
inhib_hot_ts = torch.empty_like(hot_ts).fill_(0.5)
loss = criterion(inhib_output, inhib_hot_ts.to(device))
inhib_loss = loss
loss.backward()
opt.step()
# Gather the training stats.
# stats.training.correctSamples += torch.sum( snn.predict.getClass(output) == label ).data.item()
# stats.training.numSamples += len(label)
# Calculate loss.
#loss = error.numSpikes(output, t) # spikeLoss.py
# Gather training loss stats.
stats.training.lossSum += loss.cpu().data.item()
# Display training stats. (Suitable for normal python implementation)
# if i%10 == 0: stats.print(epoch, i, (datetime.now() - tSt).total_seconds())
sum_vae_loss+=vae_loss.detach().cpu()
print(sum_vae_loss/12)
losses.append(sum_vae_loss/12)
# save the model for future use/training
#if epoch%10==0: # start saving every 10 bc memory
save_checkpoint(epoch+starting_epoch+1, 'Trained2/guided_vae', net, opt, inhib)
# save the weights to look at them and make sure they are quantizing or not or whatever
# for name, param in net.named_parameters():
# print(name, param.shape)
# np.save(f'Trained2/grads/{name}.npy',param.detach().cpu().numpy())
# np.save(f'Trained2/grads/{l.name}.npy',param.weight.grad.detach().cpu().numpy())
# # Update testing stats.
# if epoch%10==0: stats.print(epoch, timeElapsed=(datetime.now() - tSt).total_seconds())
# stats.update()
# if stats.training.bestLoss is True: torch.save(net.state_dict(), 'Trained/nmnistNet.pt')
# +
print(losses)
save_checkpoint(epoch+starting_epoch+1, 'Trained2/guided_vae', net, opt, inhib)
starting_epoch = epoch+starting_epoch
# +
#starting_epoch = load_model_from_checkpoint('Trained2/guided_vae', net, opt, inhib, n_checkpoint=-1)
# + [markdown] tags=[]
# ### Plot reconstructions, check accuracy, tsne projections for model evaluation
# -
orig = process_target(data_batch).detach().cpu().view(*[[-1]+[2,32,32]])[:,0:1]
print(orig.shape)
figure2 = plt.figure(99)
plt.imshow(make_grid(orig, scale_each=True, normalize=True).transpose(0,2).numpy())
recon_batch,mu,logvar,clas = net(data_batch_transposed.to(device))
# try voltage, it might go terribly
recon_batch,mu,logvar,clas = net.z_voltage(data_batch_transposed.to(device))
recon_batch_c = recon_batch.detach().cpu()
figure = plt.figure()
img = recon_batch_c.view(*[[-1]+[2,32,32]])[:,0:1]
print(img.shape)
plt.imshow(make_grid(img, scale_each=True, normalize=True).transpose(0,2).numpy())
# # TSNE Projections to Evaluate Disentanglement
def get_latent_train(dl, net, iterations=1):
#all_d = []
lats = []
tgts = []
for x,t in tqdm(iter(dl)):#,l,u in tqdm(iter(dl)):
new_t = t[t[:,-1,:].argmax(1)!=10]
# if new_t.shape[0] < 2:
# #print(new_t.shape)
# continue
new_t = new_t[:,-1,:].argmax(1)
#print(new_t.shape)
x = x[t[:,-1,:].argmax(1)!=10]
#print(new_t.shape)
# process target and input to make SLAYER compatible
x = transpose_torchneuromorphic_to_SLAYER(x)
with torch.no_grad():
mu, logvar = net.encode(x.to(device))
#logvar = torch.ones(mu.shape).to(device)
lat = net.reparameterize(mu,logvar).detach().cpu().numpy()
lats += lat.tolist()
tgts += new_t.tolist()
#all_d += process_target(x).tolist()
return np.array(lats), np.array(tgts)#[:,-1,:].argmax(1)
def get_latent_voltage(dl, net):
lats = []
tgts = []
for x,t in tqdm(iter(dl)):#,l,u in tqdm(iter(dl)):
new_t = t[t[:,-1,:].argmax(1)!=10]
# if new_t.shape[0] < 2:
# #print(new_t.shape)
# continue
new_t = new_t[:,-1,:].argmax(1)
#print(new_t.shape)
x = x[t[:,-1,:].argmax(1)!=10]
#print(new_t.shape)
# process target and input to make SLAYER compatible
x = transpose_torchneuromorphic_to_SLAYER(x)
with torch.no_grad():
mu, logvar = net.encode_voltage(x.to(device))
#logvar = torch.ones(mu.shape).to(device)
lat = mu.detach().cpu().numpy() #net.reparameterize(mu,logvar).detach().cpu().numpy()
lats += lat.tolist()
tgts += new_t.tolist()
#all_d += process_target(x).tolist()
return np.array(lats), np.array(tgts)#[:,-1,:].argmax(1)
# + tags=[]
def tsne_project(lats, tgts, net, do_plot = True):
from sklearn.manifold import TSNE
#print(lats.shape)
lat_tsne = TSNE(n_components=2).fit_transform(lats)
inhib_tsne = TSNE(n_components=2).fit_transform(net.inhibit_z(torch.from_numpy(lats)).numpy())
exc_tsne = TSNE(n_components=2).fit_transform(net.excite_z(torch.from_numpy(lats)).numpy())
#tgts = tgts[:10]
if do_plot:
fig = plt.figure(figsize=(16,10))
fig4 = plt.figure(figsize=(16,10))
fig5 = plt.figure(figsize=(16,10))
ax = fig.add_subplot()
ax4 = fig4.add_subplot()
ax5 = fig5.add_subplot()
for i in range(10):#1):
idx = tgts==i
ax.scatter(lat_tsne[idx,0],lat_tsne[idx,1], label = dataset.mapping[i])
ax4.scatter(exc_tsne[idx,0],exc_tsne[idx,1], label = dataset.mapping[i])
ax5.scatter(inhib_tsne[idx,0],inhib_tsne[idx,1], label = dataset.mapping[i])
ax.legend()
ax4.legend()
ax5.legend()
return lat_tsne, fig, fig4, fig5
else:
return lat_tsne
# -
def eval_accuracy(lats, tgts, excite, net):
correct_count, all_count = 0, 0
zs = excite.excite_z(torch.from_numpy(lats))
net.eval()
print(zs.shape)
for i in range(len(tgts)):
net.eval()
with torch.no_grad():
logps = net(torch.unsqueeze(zs[i],0).to(device)) # was net.model(...)
ps = torch.exp(logps.cuda())
probab = list(ps.cpu().numpy()[0])
pred_label = probab.index(max(probab))
true_label = tgts[i]#.numpy()[i]
#print(true_label)
if(true_label == pred_label):
correct_count += 1
all_count += 1
return correct_count/all_count
lats, tgts = get_latent_train(train_dl, net, iterations=1)
print(lats.shape)
lats_test, tgts_test = get_latent_train(test_dl, net, iterations=1)
train_acc = eval_accuracy(lats, tgts, inhib, net.cls_sq)
print(train_acc)
test_acc = eval_accuracy(lats_test, tgts_test, inhib, net.cls_sq)
print(test_acc)
print(dataset.mapping)
lat_tsne, fig, fig4, fig5 = tsne_project(lats, tgts, inhib)
lat_tsne, fig, fig4, fig5 = tsne_project(lats_test, tgts_test, inhib)
lats, tgts = get_latent_voltage(train_dl, net)
print(lats.shape)
lats_test, tgts_test = get_latent_voltage(test_dl, net)
train_acc = eval_accuracy(lats, tgts, inhib, net.cls_sq)
print(train_acc)
test_acc = eval_accuracy(lats_test, tgts_test, inhib, net.cls_sq)
print(test_acc)
print(lats)
# +
#print(np.clip(lats,a_min=-99999,a_max=99999))
# +
# note to self: need to clamp values after using reparamaterized voltage values
# could also try non reparameterized values
# +
#lats = np.clip(lats,a_min=-999999,a_max=999999)
# -
lat_tsne, fig, fig4, fig5 = tsne_project(lats, tgts, inhib)
# + [markdown] tags=[]
# ## Plot the Results
# +
# Plot the results.
# Learning loss
plt.figure(1)
plt.semilogy(stats.training.lossLog, label='Training')
plt.semilogy(stats.testing .lossLog, label='Testing')
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.legend()
# Learning accuracy
plt.figure(2)
plt.plot(stats.training.accuracyLog, label='Training')
plt.plot(stats.testing .accuracyLog, label='Testing')
plt.xlabel('Epoch')
plt.ylabel('Accuracy')
plt.legend()
plt.show()
# -
# ## Save training statistics
stats.save('model_weights/')
# ## Generate Loihi parameters
# Generate Loihi parameters i.e. weight files.
# +
#net.load_state_dict(torch.load('/nmnistNet.pt'))
conv1Encoder = quantizeParams.apply(net.encoder.conv1.weight, 2).flatten().cpu().data.numpy()
conv2Encoder = quantizeParams.apply(net.encoder.conv2.weight, 2).flatten().cpu().data.numpy()
pool1Encoder = quantizeParams.apply(net.encoder.pool1.weight, 2).flatten().cpu().data.numpy()
pool2Encoder = quantizeParams.apply(net.encoder.pool2.weight, 2).flatten().cpu().data.numpy()
pool3Encoder = quantizeParams.apply(net.encoder.pool3.weight, 2).flatten().cpu().data.numpy()
fc1Encoder = quantizeParams.apply(net.encoder.fc1.weight, 2).flatten().cpu().data.numpy()
fc2Encoder = quantizeParams.apply(net.encoder.fc2.weight, 2).flatten().cpu().data.numpy()
muHead = quantizeParams.apply(net.encoder_head.fc_mu.weight, 2).flatten().cpu().data.numpy()
logvarHead = quantizeParams.apply(net.encoder_head.fc_logvar.weight, 2).flatten().cpu().data.numpy()
np.save('model_weights/VAEConv1.npy', conv1Encoder)
np.save('model_weights/VAEConv2.npy', conv2Encoder)
np.save('model_weights/VAEpool1.npy', pool1Encoder)
np.save('model_weights/VAEpool2.npy', pool2Encoder)
np.save('model_weights/VAEpool3.npy', pool3Encoder)
np.save('model_weights/VAEfc1.npy', fc1Encoder)
np.save('model_weights/VAEfc2.npy', fc2Encoder)
np.save('model_weights/Headfc_mu.npy', muHead)
np.save('model_weights/Headfc_logvar.npy', logvarHead)
plt.figure(11)
plt.hist(conv1Encoder, 256)
plt.title('conv1 weights')
plt.figure(12)
plt.hist(conv2Encoder, 256)
plt.title('conv2 weights')
plt.show()
# -
| slayer_loihi_example/Loihi_Simulator_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PS Clock Control
# This notebook demonstrates how to use `Clocks` class to control the PL clocks.
#
# By default, there are at most 4 PL clocks enabled in the system. They all can be reprogrammed to valid clock rates.
#
# Whenever the overlay is downloaded, the required clocks will also be configured.
#
# References:
#
# https://www.xilinx.com/support/documentation/user_guides/ug585-Zynq-7000-TRM.pdf
# +
import os, warnings
from pynq import PL
from pynq import Overlay
if not os.path.exists(PL.bitfile_name):
warnings.warn('There is no overlay loaded after boot.', UserWarning)
# -
# **Note**: If you see a warning message in the above cell, it means that no overlay
# has been loaded after boot, hence the PL server is not aware of the
# current status of the PL. In that case you won't be able to run this notebook
# until you manually load an overlay at least once using:
#
# ```python
# from pynq import Overlay
# ol = Overlay('your_overlay.bit')
# ```
#
# If you do not see any warning message, you can safely proceed.
#
# ### Show All Clocks
#
# The following example shows all the current clock rates on the board.
# +
from pynq import Clocks
print(f'CPU: {Clocks.cpu_mhz:.6f}MHz')
print(f'FCLK0: {Clocks.fclk0_mhz:.6f}MHz')
print(f'FCLK1: {Clocks.fclk1_mhz:.6f}MHz')
print(f'FCLK2: {Clocks.fclk2_mhz:.6f}MHz')
print(f'FCLK3: {Clocks.fclk3_mhz:.6f}MHz')
# -
# ### Set Clock Rates
# The easiest way is to set the attributes directly. Random clock rates are used in the following examples; the clock manager will set the clock rates with best effort.
#
# If the desired frequency and the closest possible clock rate differs more than 1%, a warning will be raised.
# +
Clocks.fclk0_mhz = 27.123456
Clocks.fclk1_mhz = 31.436546
Clocks.fclk2_mhz = 14.597643
Clocks.fclk3_mhz = 0.251954
print(f'CPU: {Clocks.cpu_mhz:.6f}MHz')
print(f'FCLK0: {Clocks.fclk0_mhz:.6f}MHz')
print(f'FCLK1: {Clocks.fclk1_mhz:.6f}MHz')
print(f'FCLK2: {Clocks.fclk2_mhz:.6f}MHz')
print(f'FCLK3: {Clocks.fclk3_mhz:.6f}MHz')
# -
# ### Reset Clock Rates
# Recover the original clock rates. This can be done by simply reloading the overlay
# (overlay will be downloaded automatically after instantiation).
# +
_ = Overlay(PL.bitfile_name)
print(f'FCLK0: {Clocks.fclk0_mhz:.6f}MHz')
print(f'FCLK1: {Clocks.fclk1_mhz:.6f}MHz')
print(f'FCLK2: {Clocks.fclk2_mhz:.6f}MHz')
print(f'FCLK3: {Clocks.fclk3_mhz:.6f}MHz')
| pynq/notebooks/common/zynq_clocks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Kanghee-Lee/Mask-RCNN_TF/blob/master/balloon_data.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="apdJS7A7N5hQ" colab_type="code" colab={}
# + id="gEYl7jp3OKD0" colab_type="code" colab={}
from google.colab import drive
drive.mount('/content/drive')
# + id="QUpCHFg1QlTo" colab_type="code" colab={}
'''
!unzip -uq "/content/drive/My Drive/논문구현/Mask-RCNN/FastMaskRCNN" -d "/content/FastMaskRCNN"
!cd 'FastMaskRCNN/libs/datasets/pycocotools/' && make
!unzip -uq "/content/drive/My Drive/논문구현/Mask-RCNN/balloon_dataset" -d "/content/FastMaskRCNN/data"
!wget http://download.tensorflow.org/models/resnet_v1_50_2016_08_28.tar.gz
!mkdir "/content/FastMaskRCNN/data/pretrained_models"
!tar -zxvf "/content/resnet_v1_50_2016_08_28.tar.gz" -C "/content/FastMaskRCNN/data/pretrained_models"
!cd 'FastMaskRCNN/libs/' && make
!python "/content/FastMaskRCNN/train/train.py"
'''
# + id="YesgKc90TdPo" colab_type="code" colab={}
# !git clone https://github.com/matterport/Mask_RCNN.git
# !wget https://github.com/matterport/Mask_RCNN/releases/download/v2.1/balloon_dataset.zip
# !wget https://github.com/matterport/Mask_RCNN/releases/download/v2.1/mask_rcnn_balloon.h5 -P Mask_RCNN/
# !mkdir -p "/content/Mask_RCNN/datasets/"
# !unzip -uq "balloon_dataset.zip" -d "/content/Mask_RCNN/datasets/"
# + id="NIkd9cTJTl00" colab_type="code" colab={}
import os
import cv2
import sys
import itertools
import math
import logging
import json
import re
import random
from collections import OrderedDict
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import matplotlib.lines as lines
from matplotlib.patches import Polygon
# Root directory of the project
ROOT_DIR = os.path.abspath("/content/Mask_RCNN")
print(ROOT_DIR)
# Import Mask RCNN
sys.path.append(ROOT_DIR) # To find local version of the library
from mrcnn import utils
from mrcnn import visualize
from mrcnn.visualize import display_images
import mrcnn.model as modellib
from mrcnn.model import log
from samples.balloon import balloon
# %matplotlib inline
# + id="jjXDvFf6DK63" colab_type="code" colab={}
config = balloon.BalloonConfig()
BALLOON_DIR = os.path.join(ROOT_DIR, "datasets/balloon")
# !cd 'Mask_RCNN/datasets/balloon' && ls
# + id="1JEX9QNMDUM5" colab_type="code" colab={}
# Load dataset
# Get the dataset from the releases page
# https://github.com/matterport/Mask_RCNN/releases
# #!cd '/Mask_RCNN/datasets/balloon/train/' && !ls
dataset = balloon.BalloonDataset()
dataset.load_balloon(BALLOON_DIR, "train")
# Must call before using the dataset
dataset.prepare()
print("Image Count: {}".format(len(dataset.image_ids)))
print("Class Count: {}".format(dataset.num_classes))
for i, info in enumerate(dataset.class_info):
print("{:3}. {:50}".format(i, info['name']))
# + id="fNRBVOBVD4-l" colab_type="code" colab={}
# Load and display random samples
image_ids = np.random.choice(dataset.image_ids, 4)
for image_id in image_ids:
image = dataset.load_image(image_id)
mask, class_ids = dataset.load_mask(image_id)
visualize.display_top_masks(image, mask, class_ids, dataset.class_names)
# + id="wZPD40o3FMWF" colab_type="code" colab={}
# Load random image and mask.
image_id = random.choice(dataset.image_ids)
image = dataset.load_image(image_id)
print(image.shape)
mask, class_ids = dataset.load_mask(image_id)
print(mask)
print(np.where(True==mask))
print(class_ids)
print('#'*40)
# Compute Bounding box
bbox = utils.extract_bboxes(mask)
print(bbox)
norm=utils.norm_boxes(bbox, [1365, 2048])
print(norm)
# Display image and additional stats
print("image_id ", image_id, dataset.image_reference(image_id))
log("image", image)
log("mask", mask)
log("class_ids", class_ids)
log("bbox", bbox)
# Display image and instances
visualize.display_instances(image, bbox, mask, class_ids, dataset.class_names)
# + id="E9UvxriFGvPM" colab_type="code" colab={}
import tensorflow as tf
import numpy as np
'''
i=tf.keras.layers.Input(shape=(1,))
b=tf.keras.layers.Dense(32)(i)
m=tf.keras.models.Model(inputs=i, outputs=b)
aa=np.array([[1, ]])
sess=tf.Session()
print(m([aa]))
print(m(aa).eval(session=sess))
print(m)
'''
import keras.layers as KL
print('#'*20)
input_image = KL.Input(
shape=[None, None, 3], name="input_image")
print('#'*20)
input_image_meta = KL.Input(shape=[12,],
name="input_image_meta")
print('#'*20)
input_rpn_match = KL.Input(
shape=[None, 1], name="input_rpn_match", dtype=tf.int32)
input_rpn_bbox = KL.Input(
shape=[None, 4], name="input_rpn_bbox", dtype=tf.float32)
print('#'*20)
# Detection GT (class IDs, bounding boxes, and masks)
# 1. GT Class IDs (zero padded)
input_gt_class_ids = KL.Input(
shape=[None], name="input_gt_class_ids", dtype=tf.int32)
# 2. GT Boxes in pixels (zero padded)
# [batch, MAX_GT_INSTANCES, (y1, x1, y2, x2)] in image coordinates
input_gt_boxes = KL.Input(
shape=[None, 4], name="input_gt_boxes", dtype=tf.float32)
# Normalize coordinates
gt_boxes = KL.Lambda(lambda x: norm_boxes_graph(
x, K.shape(input_image)[1:3]))(input_gt_boxes)
# 3. GT Masks (zero padded)
# [batch, height, width, MAX_GT_INSTANCES]
print('#'*20)
input_gt_masks = KL.Input(
shape=[config.MINI_MASK_SHAPE[0],
config.MINI_MASK_SHAPE[1], None],
name="input_gt_masks", dtype=bool)
class BatchNorm(KL.BatchNormalization):
"""Extends the Keras BatchNormalization class to allow a central place
to make changes if needed.
Batch normalization has a negative effect on training if batches are small
so this layer is often frozen (via setting in Config class) and functions
as linear layer.
"""
def call(self, inputs, training=None):
"""
Note about training values:
None: Train BN layers. This is the normal mode
False: Freeze BN layers. Good when batch size is small
True: (don't use). Set layer in training mode even when making inferences
"""
return super(self.__class__, self).call(inputs, training=training)
def identity_block(input_tensor, kernel_size, filters, stage, block,
use_bias=True, train_bn=True):
"""The identity_block is the block that has no conv layer at shortcut
# Arguments
input_tensor: input tensor
kernel_size: default 3, the kernel size of middle conv layer at main path
filters: list of integers, the nb_filters of 3 conv layer at main path
stage: integer, current stage label, used for generating layer names
block: 'a','b'..., current block label, used for generating layer names
use_bias: Boolean. To use or not use a bias in conv layers.
train_bn: Boolean. Train or freeze Batch Norm layers
"""
nb_filter1, nb_filter2, nb_filter3 = filters
conv_name_base = 'res' + str(stage) + block + '_branch'
bn_name_base = 'bn' + str(stage) + block + '_branch'
x = KL.Conv2D(nb_filter1, (1, 1), name=conv_name_base + '2a',
use_bias=use_bias)(input_tensor)
x = BatchNorm(name=bn_name_base + '2a')(x, training=train_bn)
x = KL.Activation('relu')(x)
x = KL.Conv2D(nb_filter2, (kernel_size, kernel_size), padding='same',
name=conv_name_base + '2b', use_bias=use_bias)(x)
x = BatchNorm(name=bn_name_base + '2b')(x, training=train_bn)
x = KL.Activation('relu')(x)
x = KL.Conv2D(nb_filter3, (1, 1), name=conv_name_base + '2c',
use_bias=use_bias)(x)
x = BatchNorm(name=bn_name_base + '2c')(x, training=train_bn)
x = KL.Add()([x, input_tensor])
x = KL.Activation('relu', name='res' + str(stage) + block + '_out')(x)
return x
def conv_block(input_tensor, kernel_size, filters, stage, block,
strides=(2, 2), use_bias=True, train_bn=True):
"""conv_block is the block that has a conv layer at shortcut
# Arguments
input_tensor: input tensor
kernel_size: default 3, the kernel size of middle conv layer at main path
filters: list of integers, the nb_filters of 3 conv layer at main path
stage: integer, current stage label, used for generating layer names
block: 'a','b'..., current block label, used for generating layer names
use_bias: Boolean. To use or not use a bias in conv layers.
train_bn: Boolean. Train or freeze Batch Norm layers
Note that from stage 3, the first conv layer at main path is with subsample=(2,2)
And the shortcut should have subsample=(2,2) as well
"""
nb_filter1, nb_filter2, nb_filter3 = filters
conv_name_base = 'res' + str(stage) + block + '_branch'
bn_name_base = 'bn' + str(stage) + block + '_branch'
x = KL.Conv2D(nb_filter1, (1, 1), strides=strides,
name=conv_name_base + '2a', use_bias=use_bias)(input_tensor)
x = BatchNorm(name=bn_name_base + '2a')(x, training=train_bn)
x = KL.Activation('relu')(x)
x = KL.Conv2D(nb_filter2, (kernel_size, kernel_size), padding='same',
name=conv_name_base + '2b', use_bias=use_bias)(x)
x = BatchNorm(name=bn_name_base + '2b')(x, training=train_bn)
x = KL.Activation('relu')(x)
x = KL.Conv2D(nb_filter3, (1, 1), name=conv_name_base +
'2c', use_bias=use_bias)(x)
x = BatchNorm(name=bn_name_base + '2c')(x, training=train_bn)
shortcut = KL.Conv2D(nb_filter3, (1, 1), strides=strides,
name=conv_name_base + '1', use_bias=use_bias)(input_tensor)
shortcut = BatchNorm(name=bn_name_base + '1')(shortcut, training=train_bn)
x = KL.Add()([x, shortcut])
x = KL.Activation('relu', name='res' + str(stage) + block + '_out')(x)
return x
def resnet_graph(input_image, architecture, stage5=False, train_bn=True):
"""Build a ResNet graph.
architecture: Can be resnet50 or resnet101
stage5: Boolean. If False, stage5 of the network is not created
train_bn: Boolean. Train or freeze Batch Norm layers
"""
print('#'*20)
assert architecture in ["resnet50", "resnet101"]
# Stage 1
x = KL.ZeroPadding2D((3, 3))(input_image)
x = KL.Conv2D(64, (7, 7), strides=(2, 2), name='conv1', use_bias=True)(x)
x = BatchNorm(name='bn_conv1')(x, training=train_bn)
x = KL.Activation('relu')(x)
C1 = x = KL.MaxPooling2D((3, 3), strides=(2, 2), padding="same")(x)
# Stage 2
x = conv_block(x, 3, [64, 64, 256], stage=2, block='a', strides=(1, 1), train_bn=train_bn)
x = identity_block(x, 3, [64, 64, 256], stage=2, block='b', train_bn=train_bn)
C2 = x = identity_block(x, 3, [64, 64, 256], stage=2, block='c', train_bn=train_bn)
# Stage 3
x = conv_block(x, 3, [128, 128, 512], stage=3, block='a', train_bn=train_bn)
x = identity_block(x, 3, [128, 128, 512], stage=3, block='b', train_bn=train_bn)
x = identity_block(x, 3, [128, 128, 512], stage=3, block='c', train_bn=train_bn)
C3 = x = identity_block(x, 3, [128, 128, 512], stage=3, block='d', train_bn=train_bn)
# Stage 4
x = conv_block(x, 3, [256, 256, 1024], stage=4, block='a', train_bn=train_bn)
block_count = {"resnet50": 5, "resnet101": 22}[architecture]
for i in range(block_count):
x = identity_block(x, 3, [256, 256, 1024], stage=4, block=chr(98 + i), train_bn=train_bn)
C4 = x
# Stage 5
if stage5:
x = conv_block(x, 3, [512, 512, 2048], stage=5, block='a', train_bn=train_bn)
x = identity_block(x, 3, [512, 512, 2048], stage=5, block='b', train_bn=train_bn)
C5 = x = identity_block(x, 3, [512, 512, 2048], stage=5, block='c', train_bn=train_bn)
else:
C5 = None
return [C1, C2, C3, C4, C5]
_, C2, C3, C4, C5 = resnet_graph(input_image, config.BACKBONE,
stage5=True, train_bn=config.TRAIN_BN)
# Top-down Layers
# TODO: add assert to varify feature map sizes match what's in config
P5 = KL.Conv2D(config.TOP_DOWN_PYRAMID_SIZE, (1, 1), name='fpn_c5p5')(C5)
P4 = KL.Add(name="fpn_p4add")([
KL.UpSampling2D(size=(2, 2), name="fpn_p5upsampled")(P5),
KL.Conv2D(config.TOP_DOWN_PYRAMID_SIZE, (1, 1), name='fpn_c4p4')(C4)])
P3 = KL.Add(name="fpn_p3add")([
KL.UpSampling2D(size=(2, 2), name="fpn_p4upsampled")(P4),
KL.Conv2D(config.TOP_DOWN_PYRAMID_SIZE, (1, 1), name='fpn_c3p3')(C3)])
P2 = KL.Add(name="fpn_p2add")([
KL.UpSampling2D(size=(2, 2), name="fpn_p3upsampled")(P3),
KL.Conv2D(config.TOP_DOWN_PYRAMID_SIZE, (1, 1), name='fpn_c2p2')(C2)])
# Attach 3x3 conv to all P layers to get the final feature maps.
P2 = KL.Conv2D(config.TOP_DOWN_PYRAMID_SIZE, (3, 3), padding="SAME", name="fpn_p2")(P2)
P3 = KL.Conv2D(config.TOP_DOWN_PYRAMID_SIZE, (3, 3), padding="SAME", name="fpn_p3")(P3)
P4 = KL.Conv2D(config.TOP_DOWN_PYRAMID_SIZE, (3, 3), padding="SAME", name="fpn_p4")(P4)
P5 = KL.Conv2D(config.TOP_DOWN_PYRAMID_SIZE, (3, 3), padding="SAME", name="fpn_p5")(P5)
# P6 is used for the 5th anchor scale in RPN. Generated by
# subsampling from P5 with stride of 2.
P6 = KL.MaxPooling2D(pool_size=(1, 1), strides=2, name="fpn_p6")(P5)
# Note that P6 is used in RPN, but not in the classifier heads.
rpn_feature_maps = [P2, P3, P4, P5, P6]
mrcnn_feature_maps = [P2, P3, P4, P5]
print('#'*20)
anchors = self.get_anchors(config.IMAGE_SHAPE)
# Duplicate across the batch dimension because Keras requires it
# TODO: can this be optimized to avoid duplicating the anchors?
anchors = np.broadcast_to(anchors, (config.BATCH_SIZE,) + anchors.shape)
# A hack to get around Keras's bad support for constants
anchors = KL.Lambda(lambda x: tf.Variable(anchors), name="anchors")(input_image)
# RPN Model
rpn = build_rpn_model(1,
3, 256)
# Loop through pyramid layers
layer_outputs = [] # list of lists
for p in rpn_feature_maps:
layer_outputs.append(rpn([p]))
# Concatenate layer outputs
# Convert from list of lists of level outputs to list of lists
# of outputs across levels.
# e.g. [[a1, b1, c1], [a2, b2, c2]] => [[a1, a2], [b1, b2], [c1, c2]]
output_names = ["rpn_class_logits", "rpn_class", "rpn_bbox"]
outputs = list(zip(*layer_outputs))
outputs = [KL.Concatenate(axis=1, name=n)(list(o))
for o, n in zip(outputs, output_names)]
| demo/balloon_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from textblob import TextBlob
y="Son of a bitch"
b=y.split()
print(b)
profanity = ['fuck', 'shit', 'bitch', 'motherfucker', 'fucker', 'slut', 'whore', 'cock', 'dick']
for i in b:
for j in profanity:
if i==j:
s=1
print(s)
r=TextBlob(x).sentiment.polarity
if (s==1) and (r<0):
print("Your status is negative and profane")
elif (s==1) and (r>=0):
print("Your status contains profanity")
else:
print ("Good to go")
z = TextBlob("Son of a bitch")
z.sentiment.polarity
| bumblebee/sentiment_analysis/notebook/TextBlob.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Adam
# Presented during ML reading group, 2019-11-19.
#
# Author: <NAME>, <EMAIL>
# The [Adam paper](https://arxiv.org/pdf/1412.6980)
#
# The Adam method combines the advantages of two other methods: AdaGrad (Duchi et al., 2011) and RMSProp (Tieleman & Hinton, 2012):
# - it works well with sparse gradients (AdaGrad)
# - it works well with on-line and non-stationary settings (RMSProp)
#
# The Adam method computes the adaptive learning rates from estimates of first and second moments of gradients. The first and second moments of gradients are estimated using exponentially moving averages with hyper-parameters $\beta_{1}, \beta_{2} \in [0, 1)$ decay rates of these moving averages.
#
# The advantages of the Adam method are:
# - the magnitudes of parameter updates are invariant to rescaling of the gradient
# - its stepsize are approximately bounded by the stepsize hyperparameter
# - it does not require a stationary objective
# - it works with sparse gradients
# - it naturally performs a form of stepsize annealing
#
#
#
#
#
#
#
# ## The algorithm
#
# ***
#
# **Algorithm 1:**
#
# $g_{t}^2$ indicates the elementwise square of $g_{t}$
#
# Good default settings for the tested machine learning problems are $\alpha = 0.001$, $\beta_{1} = 0.9$, $\beta_{2} = 0.999$ and $\epsilon = 10^{-8} $
#
# All operations on vectors are element-wise.
# With $ \beta_{1}^t$ and $\beta_{2}^t$ we denote $\beta_{1}$ and $\beta_{2}$ to the power $t$
#
# ***
# **Require:** $\alpha$ Stepsize
# **Require:** $\beta_{1}, \beta_{2} \in [0, 1)$ : Exponential decay rates for the moment estimates
# **Require** $f(\theta)$ : Stochastic objective function with parameters $\theta$
# **Require** $\theta_0$ : Initial parameter vector
# $\;\;\;\;$ $m_{0} \leftarrow 0$ (Initialize 1st moment vector)
# $\;\;\;\;$ $v_{0} \leftarrow 0$ (Initialize 2nd moment vector)
# $\;\;\;\;$ $t \leftarrow 0$ (Initialize timestep)
# $\;\;\;\;$ **while** $\theta_{t}$ not converged **do**:
# $\;\;\;\;\;\;\;\;$ $t \leftarrow t+1$
# $\;\;\;\;\;\;\;\;$ $g_{t} \leftarrow \nabla(f(\theta_{t-1}))$ (Get gradients w.r.t. stochastic objective at timestep $t$)
# $\;\;\;\;\;\;\;\;$ $m_{t} \leftarrow \beta_{1} \cdot m_{t-1}+(1-\beta_{1}) \cdot g_{t}$ (Update biased first moment estimate)
# $\;\;\;\;\;\;\;\;$ $v_{t} \leftarrow \beta_{2} \cdot v_{t-1}+(1-\beta_{2}) \cdot g_{t}^2$ (Update biased second raw moment estimate)
# $\;\;\;\;\;\;\;\;$ $\hat{m_{t}} \leftarrow \dfrac{m_{t}}{(1-\beta_{1}^t)}$ (Compute bias-corrected first moment estimate)
# $\;\;\;\;\;\;\;\;$ $\hat{v_{t}} \leftarrow \dfrac{v_{t}}{(1-\beta_{2}^t)} $ (Compute bias-corrected second raw moment estimate)
# $\;\;\;\;\;\;\;\;$ $\theta_{t} \leftarrow \theta_{t-1} - \dfrac{\alpha \cdot \hat{m_{t}}}{\sqrt{\hat{v_{t}}}+\varepsilon } $ (Update parameters)
# $\;\;\;\;$**end while**
# $\;\;\;\;$**return** $\theta_{t}$ (Resulting parameters)
#
# ***
# * Adam uses estimations of first and second moments of gradient to adapt the learning rate for each weight of the neural network.
# * The algorithm updates exponential moving averages of the gradient ($m_{t}$) and the squared gradient
# ($v_{t}$) where the hyper-parameters $\beta_{1}, \beta_{2} \in [0, 1)$ control the exponential decay rates of these moving
# averages.
# * The moving averages themselves are estimates of the 1st moment (the mean) and the 2nd raw moment (the uncentered variance) of the gradient.
# * However, these moving averages are initialized as (vectors of) 0’s, leading to moment estimates that are biased towards zero, especially during the initial timesteps, and especially when the decay rates are small (i.e. the $\beta_{s}$ are close to 1). The good news is that this initialization bias can be easily counteracted, resulting in bias-corrected estimates $\hat{m_{t}}$ and $\hat{v_{t}}$
#
# * The efficiency of the algorithm 1 can be improved with the following rule:
#
# $ \alpha_{t} = \alpha \cdot \dfrac{\sqrt{1-\beta_{2}^t}} {1-\beta_{1}^t}$ and $\theta_{t} \leftarrow \theta_{t-1} - \dfrac{\alpha_{t} \cdot m_{t}}{\sqrt{v_{t}}+\hat{\varepsilon }} $
#
#
#
# ## The update rule
#
# * Assuming $\varepsilon = 0$, the effective step taken in parameter space at timestep $t$ is $\Delta_{t} = \alpha \cdot \dfrac{\hat{m_{t}}} {\sqrt{\hat{v_{t}}}}$
# * The effective step size has two bounds:
# * In the most severe case of sparsity: when a gradient has been zero at all timesteps except at the current timestep
# * $\mid \Delta_{t} \mid \le \dfrac{\alpha \cdot (1-\beta_{1})}{\sqrt{1-\beta_{2}}}$, in the case $(1-\beta_{1}) \gt \sqrt{1-\beta_{2}}$
# * Otherwise
# * $\mid \Delta_{t} \mid \le \alpha$
#
# * In more common scenarios, we will have $\mid \dfrac{ \hat{m_{t}} } { \sqrt{\hat{v_{t}}} } \mid \; \approx 1$ since $\mid \dfrac{ E[g_{t}] } { E[g_{t}^2] } \mid \le 1$
# * The effective magnitude of the steps taken in parameter space at each timestep are approximately bounded by the stepsize setting $\alpha$, i.e., $|\Delta_{t}| \le \alpha$
#
#
# * With a slight abuse of terminology, we will call the ratio $\dfrac{ \hat{m_{t}} } { \sqrt{\hat{v_{t}}} }$ the signal-to-noise ratio (SNR).
# * With a smaller SNR the effective stepsize $\Delta{t}$ will be closer to zero. This is a desirable property, since a smaller SNR means that there is greater uncertainty about whether the direction of $\hat{m_{t}}$ corresponds to the direction of the true gradient. For example, the SNR value typically becomes closer to 0 towards an optimum, leading to smaller effective steps in parameter space: a form of automatic annealing.
#
#
# * The effective stepsize $\Delta_{t}$ is also invariant to the scale of the gradients; rescaling the gradients $g$ with factor $c$ will scale $\hat{m_{t}}$ with a factor $c$ and $\hat{v_{t}}$ with a factor $c^2$, which cancel out: $ \dfrac{(c \cdot \hat{m_{t}})} {\sqrt{c^2 \cdot \hat{v_{t}}}} = \dfrac{\hat{m_{t}}}{\sqrt{\hat{v_{t}}}} $
# ## Initialization bias correction
#
# Adam uses estimations of first and second moments of gradient to adapt the learning rate for each weight of the neural network. The algorithm updates exponential moving averages of the gradient ($m_{t}$) and the squared gradient
# ($v_{t}$) where the hyper-parameters $\beta_{1}, \beta_{2} \in [0, 1)$ control the exponential decay rates of these moving averages. The moving averages themselves are estimates of the 1st moment (the mean) and the 2nd raw moment (the uncentered variance) of the gradient.
# Since $m_{t}$ and $v_{t}$ are the estimates of the first moment and the second moment of the gradient, respectively, and the first moment and second moment are the estimates used to adapt the learning rate for each weight of the neural network, we want to ensure that both sets of estimators estimate the same expected value, hence the following equalities must be true:
#
# $$E[m_{t}] = E[g{t}]$$
# $$E[v_{t}] = E[g{t}^2]$$
#
# Expected values of the estimators should equal the parameter we're trying to estimate, as it happens. If these properties would held true, it means we have **unbiased estimators**.
#
#
# Looking at some values of $m$:
# $m_{0} = 0$
# $m_{1} = \beta_{1} \cdot m_{0} + (1- \beta_{1}) \cdot g_{1} = (1- \beta_{1}) \cdot g_{1}$
# $m_{2} = \beta_{1} \cdot m_{1} + (1- \beta_{1}) \cdot g_{2} = \beta_{1} \cdot (1- \beta_{1}) \cdot g_{1} + (1- \beta_{1}) \cdot g_{2} $
# $m_{3} = \beta_{1} \cdot m_{2} + (1- \beta_{1}) \cdot g_{3} = \beta_{1} ^ 2 \cdot (1- \beta_{1}) \cdot g_{1} + \beta_{1} \cdot (1- \beta_{1}) \cdot g_{2} + (1- \beta_{1}) \cdot g_{3}$
#
# we can rewrite the formula for our moving average:
#
# $$ m_{t} = (1-\beta_{1}) \cdot \sum_{i=0}^{t}{\beta_{1} ^ {t-i} \cdot g_{i} }$$
#
# Now, we can take a look at the expected value of $m_{t}$, to see how it relates to the true first moment, so we can correct for the discrepancy of the two :
#
# $$ E[m_{t}] = E[(1-\beta_{1}) \cdot \sum_{i=0}^{t}{\beta_{1} ^ {t-i} \cdot g_{i} }]$$
#
# $$ E[m_{t}] = E[g_{t}](1-\beta_{1}) \cdot \sum_{i=0}^{t}{\beta_{1} ^ {t-i} } + C$$
#
# $$ E[m_{t}] = E[g_{t}](1-\beta_{1}^t) + C$$
#
# Since we have a biased estimator $E[m_{t}]$, we have to correct it, so that the expected value is the one we want. This step is usually referred to as bias correction. The final formulas for our estimator will be as follows:
#
# $$ \hat{m_{t}} = \dfrac{m_{t}}{1-\beta_{1}^t}$$
# $$ \hat{v_{t}} = \dfrac{v_{t}}{1-\beta_{2}^t}$$
# ## Example
# +
# #%matplotlib notebook
# %matplotlib inline
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
print(f'Numpy version: {np.__version__}')
# -
# ### Generate data
# +
from scipy.sparse import random #to generate sparse data
np.random.seed(10) # for reproducibility
m_data = 100
n_data = 4 #number of features of the data
_scales = np.array([1,10, 10,1 ]) # play with these...
_parameters = np.array([3, 0.5, 1, 7])
def gen_data(m, n, scales, parameters, add_noise=True):
# Adam as Adagrad is designed especially for sparse data.
# produce: X, a 2d tensor with m lines and n columns
# and X[:, k] uniformly distributed in [-scale_k, scale_k] with the first and the last column containing sparse data
#(approx 75% of the elements are 0)
#
# To generate a sparse data matrix with m rows and n columns
# and random values use S = random(m, n, density=0.25).A, where density = density of the data. S will be the
# resulting matrix
# more information at https://docs.scipy.org/doc/scipy/reference/generated/scipy.sparse.random.html
#
# To obtain X - generate a random matrix with X[:, k] uniformly distributed in [-scale_k, scale_k]
# set X[:, 0] and X[:, -1] to 0 and add matrix S with the sparse data.
#
# let y be X@parameters.T + epsilon, with epsilon ~ N(0, 1); y is a vector with m elements
# parameters - the ideal weights, used to produce output values y
#
X = np.random.rand(m,n) *2*scales - scales
X[:, 0] = 0
X[:, -1] = 0
S = random(m, n, density=0.25).A
X = X + S
y = X@parameters.T + np.random.randn(m)
y = np.reshape(y, (-1, 1))
return X, y
# -
X, y = gen_data(m_data, n_data, _scales, _parameters)
print(X)
print(y)
# ### Define error function, gradient, inference
def model_estimate(X, w):
'''Computes the linear regression estimation on the dataset X, using coefficients w
:param X: 2d tensor with m_data lines and n_data columns
:param w: a 1d tensor with n_data coefficients (no intercept)
:return: a 1d tensor with m_data elements y_hat = w @X.T
'''
w = w.reshape(-1, 1)
y_hat = X@w
return y_hat
def J(X, y, w):
"""Computes the mean squared error of model. See the picture from last week's sheet.
:param X: input values, of shape m_data x n_data
:param y: ground truth, column vector with m_data values
:param w: column with n_data coefficients for the linear form
:return: a scalar value >= 0
:use the same formula as in the exercise from last week
"""
w = w.reshape(-1, 1)
expr = (X@w - y)
err = np.asscalar(1.0/(2 * X.shape[0]) * expr.T @ expr)
return err
def gradient(X, y, w):
'''Commputes the gradients to be used for gradient descent.
:param X: 2d tensor with training data
:param y: 1d tensor with y.shape[0] == W.shape[0]
:param w: 1d tensor with current values of the coefficients
:return: gradients to be used for gradient descent.
:use the same formula as in the exercise from last week
'''
n = len(y)
w = w.reshape(-1, 1)
grad = 1.0 / n * X.T @ (X@w - y)
return grad## implement
# ### Momentum algorithm
#The function from last week for comparison
def gd_with_momentum(X, y, w_init, eta=1e-1, gamma = 0.9, thresh = 0.001):
"""Applies gradient descent with momentum coefficient
:params: as in gd_no_momentum
:param gamma: momentum coefficient
:param thresh: the threshold for gradient norm (to stop iterations)
:return: the list of succesive errors and the found w* vector
"""
w = w_init.reshape(-1, 1)
w_err=[]
delta = np.zeros_like(w)
while True:
grad = gradient(X, y, w)
err = J(X, y, w)
w_err.append(err)
w_nou = w + gamma * delta - eta * grad
delta = w_nou - w
w = w_nou
if np.linalg.norm(grad) < thresh :
break;
return w_err, w
w_init = np.array([0, 0, 0, 0])
errors_momentum, w_best = gd_with_momentum(X, y, w_init,0.0001, 0.9)
print(f'Momentum: How many iterations were made: {len(errors_momentum)}')
w_best
fig, axes = plt.subplots()
axes.plot(list(range(len(errors_momentum))), errors_momentum)
axes.set_xlabel('Epochs')
axes.set_ylabel('Error')
axes.set_title('Optimization with momentum')
# ### Apply AdaGrad and report resulting $\eta$'s
def ada_grad(X, y, w_init, eta_init=1e-1, eps = 0.001, thresh = 0.001):
'''Iterates with gradient descent. algorithm
:param X: 2d tensor with data
:param y: 1d tensor, ground truth
:param w_init: 1d tensor with the X.shape[1] initial coefficients
:param eta_init: the initial learning rate hyperparameter
:param eps: the epsilon value from the AdaGrad formula
:param thresh: the threshold for gradient norm (to stop iterations)
:return: the list of succesive errors w_err, the found w - the estimated feature vector
:and rates the learning rates after the final iteration
'''
w = w_init.reshape(-1, 1)
w_err=[]
cum_sq_grad = np.zeros(w.shape)
rates = np.zeros(w.shape)+eta_init
while True:
grad = gradient(X, y, w)
err = J(X, y, w)
w_err.append(err)
w = w - rates * grad
cum_sq_grad += grad**2
rates = eta_init/np.sqrt(eps+cum_sq_grad)
if np.linalg.norm(grad) < thresh :
break;
return w_err, w, rates
w_init = np.array([0,0,0,0])
adaGerr, w_ada_best, rates = ada_grad(X, y, w_init)
print(rates)
print(f'AdaGrad: How many iterations were made: {len(adaGerr)}')
w_ada_best
fig, axes = plt.subplots()
axes.plot(list(range(len(adaGerr))),adaGerr)
axes.set_xlabel('Epochs')
axes.set_ylabel('Error')
axes.set_title('Optimization with AdaGrad')
# ### Apply Adam and report resulting parameters
def adam(X, y, w_init, step_size = 0.001, beta_1=0.9, beta_2=0.999, eps = 1e-8, thresh = 0.001):
'''Iterates with gradient descent. algorithm
:param X: 2d tensor with data
:param y: 1d tensor, ground truth
:param w_init: 1d tensor with the X.shape[1] initial coefficients
:param step_size: the step size hyperparameter
:param beta_1: Exponential decay rate for the 1st moment estimate (mean)
:param beta_1: Exponential decay rate for the 2nd moment estimate (uncentered variance)
:param eps: the epsilon value from the Adam formula (avoid division by zero)
:param thresh: the threshold for gradient norm (to stop iterations)
:return: the list of succesive errors w_err, the found w - the estimated feature vector
:and rates the learning rates after the final iteration
'''
w = w_init.reshape(-1, 1)
w_err=[]
t = 0
m = np.zeros(w.shape)
v = np.zeros(w.shape)
while True:
t += 1
grad = gradient(X, y, w)
err = J(X, y, w)
w_err.append(err)
#Update biased first moment estimate
#Update biased second raw moment estimate
#Compute bias-corrected first moment estimate)
#Compute bias-corrected second raw moment estimate)
#Update parameters
w = w + delta_w
if np.linalg.norm(grad) < thresh :
break;
return w_err, w, delta_w
w_init = np.array([0,0,0,0])
adamErr, w_adam_best, delta_w = adam(X, y, w_init)
print(delta_w)
print(f'Adam: How many iterations were made: {len(adamErr)}')
w_adam_best
fig, axes = plt.subplots()
axes.plot(list(range(len(adamErr))),adamErr)
axes.set_xlabel('Epochs')
axes.set_ylabel('Error')
axes.set_title('Optimization with Adam')
# ## Adamax algorithm
# ***
#
# **Algorithm 2:** AdaMax, a variant of Adam based on the infinity norm.
# Good default settings for the tested machine learning problems are $\alpha = 0.002, \beta_{1} = 0.9$ and
# $\beta_{2} = 0.999$. With $\beta_{1}^t$ we denote $\beta_{1}$ to the power $t$. Here, $(\dfrac{\alpha} {(1 - \beta_{1}^t)}
# )$ is the learning rate with the bias-correction term for the first moment. All operations on vectors are element-wise.
#
# ***
# **Require:** $\alpha$ Stepsize
# **Require:** $\beta_{1}, \beta_{2} \in [0, 1)$ : Exponential decay rates for the moment estimates
# **Require** $f(\theta)$ : Stochastic objective function with parameters $\theta$
# **Require** $\theta_0$ : Initial parameter vector
# $\;\;\;\;$ $m_{0} \leftarrow 0$ (Initialize 1st moment vector)
# $\;\;\;\;$ $u_{0} \leftarrow 0$ (Initialize the exponentially weighted infinity norm)
# $\;\;\;\;$ $t \leftarrow 0$ (Initialize timestep)
# $\;\;\;\;$ **while** $\theta_{0}$ not converged **do**:
# $\;\;\;\;\;\;\;\;$ $t \leftarrow t+1$
# $\;\;\;\;\;\;\;\;$ $g_{t} \leftarrow \nabla(f(\theta_{t-1}))$ (Get gradients w.r.t. stochastic objective at timestep $t$)
# $\;\;\;\;\;\;\;\;$ $m_{t} \leftarrow \beta_{1} \cdot m_{t-1}+(1-\beta_{1}) \cdot g_{t}$ (Update biased first moment estimate)
# $\;\;\;\;\;\;\;\;$ $u_{t} \leftarrow max(\beta_{2} \cdot u_{t-1}, \mid g_{t} \mid )$ (Update the exponentially weighted infinity norm)
# $\;\;\;\;\;\;\;\;$ $\theta_{t} \leftarrow \theta_{t-1} - \dfrac{\alpha} {(1 - \beta_{1}^t)} \cdot \dfrac{m_{t}}{u_{t}} $ (Update parameters)
# $\;\;\;\;$**end while**
# $\;\;\;\;$**return** $\theta_{t}$ (Resulting parameters)
#
# ***
# ### Apply Adamax and report resulting parameters
def adamax(X, y, w_init, step_size = 0.001, beta_1=0.9, beta_2=0.999, eps = 1e-8, thresh = 0.001):
'''Iterates with gradient descent. algorithm
:param X: 2d tensor with data
:param y: 1d tensor, ground truth
:param w_init: 1d tensor with the X.shape[1] initial coefficients
:param step_size: the step size hyperparameter
:param beta_1: Exponential decay rate for the 1st moment estimate (mean)
:param beta_1: Exponential decay rate for the 2nd moment estimate (uncentered variance)
:param eps: the epsilon value from the Adam formula (avoid division by zero)
:param thresh: the threshold for gradient norm (to stop iterations)
:return: the list of succesive errors w_err, the found w - the estimated feature vector
:and rates the learning rates after the final iteration
'''
w = w_init.reshape(-1, 1)
w_err=[]
t = 0
m = np.zeros_like(w)
u = np.zeros_like(w)
while True:
t += 1
grad = gradient(X, y, w)
err = J(X, y, w)
w_err.append(err)
#Update biased first moment estimate
#Compute bias-corrected first moment estimate)
#Update biased second raw moment estimate
#Update parameters
w = w + delta_w
if np.linalg.norm(grad) < thresh :
break;
return w_err, w, delta_w
w_init = np.array([0,0,0,0])
adamaxErr, w_adamax_best, delta_w = adamax(X, y, w_init)
print(delta_w)
print(f'AdaMax: How many iterations were made: {len(adamaxErr)}')
w_adamax_best
fig, axes = plt.subplots()
axes.plot(list(range(len(adamaxErr))),adamaxErr)
axes.set_xlabel('Epochs')
axes.set_ylabel('Error')
axes.set_title('Optimization with Adamax')
# # Bibliography
# * <NAME>, <NAME>, [ADAM: A METHOD FOR STOCHASTIC OPTIMIZATION](https://arxiv.org/pdf/1412.6980)
# * <NAME>, [An overview of gradient descent optimization algorithms](https://ruder.io/optimizing-gradient-descent/)
# * <NAME>, [Adam — latest trends in deep learning optimization](https://towardsdatascience.com/adam-latest-trends-in-deep-learning-optimization-6be9a291375c)
| Presentations/2019/11.November/19/Adam_Worksheet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="Vtr5GXBYQviz"
# Mount my google drive, where I stored the dataset.
# + colab_type="code" id="w12ajwYkRYMj" colab={}
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="ICzn7m-J8OVy" colab_type="text"
# **Download dependencies**
# + colab_type="code" executionInfo={"elapsed": 7733, "status": "ok", "timestamp": 1577541425786, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16049478229633186858"}, "user_tz": -120} id="-00MlLK5bd_o" outputId="d36dccf0-dfc8-4b18-e216-0c185b5f17dc" colab={"base_uri": "https://localhost:8080/", "height": 358}
# !pip3 install sklearn matplotlib GPUtil
# + id="urxkHeuu90mV" colab_type="code" colab={} outputId="5e5f049e-ea0c-428f-d3bb-41e5af83acbf"
# !pip3 install torch torchvision
# + [markdown] colab_type="text" id="xTU7V2ervf9J"
# **Download Data**
# + [markdown] colab_type="text" id="1lZ8z6MIZag1"
# In order to acquire the dataset please navigate to:
#
# https://ieee-dataport.org/documents/cervigram-image-dataset
#
# Unzip the dataset into the folder "dataset".
#
# For your environment, please adjust the paths accordingly.
# + colab_type="code" executionInfo={"elapsed": 15250, "status": "ok", "timestamp": 1577541443401, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16049478229633186858"}, "user_tz": -120} id="30WlKBSGZfpq" outputId="16fb86f9-b408-47ef-c947-8f954539995b" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !rm -vrf "dataset"
# !mkdir "dataset"
# # !cp -r "/content/drive/My Drive/Studiu doctorat leziuni cervicale/cervigram-image-dataset-v2.zip" "dataset/cervigram-image-dataset-v2.zip"
# !cp -r "cervigram-image-dataset-v2.zip" "dataset/cervigram-image-dataset-v2.zip"
# !unzip "dataset/cervigram-image-dataset-v2.zip" -d "dataset"
# + [markdown] id="pzmtIBtraodf" colab_type="text"
# **Constants**
# + [markdown] id="IPVLzc_lHTQe" colab_type="text"
# For your environment, please modify the paths accordingly.
# + colab_type="code" id="g1YBANp9XJeA" colab={}
# TRAIN_PATH = '/content/dataset/data/train/'
# TEST_PATH = '/content/dataset/data/test/'
TRAIN_PATH = 'dataset/data/train/'
TEST_PATH = 'dataset/data/test/'
CROP_SIZE = 260
IMAGE_SIZE = 224
BATCH_SIZE = 100
# + [markdown] colab_type="text" id="B_koVi65ZlHP"
# **Imports**
# + colab_type="code" id="qyNuLlPNsdl1" colab={}
import torch as t
import torchvision as tv
import numpy as np
import PIL as pil
import matplotlib.pyplot as plt
from torchvision.datasets import ImageFolder
from torch.utils.data import DataLoader
from torch.nn import Linear, BCEWithLogitsLoss
import sklearn as sk
import sklearn.metrics
from os import listdir
import time
import random
import GPUtil
# + [markdown] colab_type="text" id="6Iq1zdisXRnn"
# **Memory Stats**
# + colab_type="code" executionInfo={"elapsed": 6912, "status": "ok", "timestamp": 1577541807375, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16049478229633186858"}, "user_tz": -120} id="_temaFmzXRHm" outputId="9c0793b5-4cb2-4502-a744-a42d85e12bcc" colab={"base_uri": "https://localhost:8080/", "height": 33}
import GPUtil
def memory_stats():
for gpu in GPUtil.getGPUs():
print("GPU RAM Free: {0:.0f}MB | Used: {1:.0f}MB | Util {2:3.0f}% | Total {3:.0f}MB".format(gpu.memoryFree, gpu.memoryUsed, gpu.memoryUtil*100, gpu.memoryTotal))
memory_stats()
# + [markdown] colab_type="text" id="r3KfXoIH187Z"
# **Deterministic Measurements**
# + [markdown] colab_type="text" id="bQ3rHF5K2BpX"
# This statements help making the experiments reproducible by fixing the random seeds. Despite fixing the random seeds, experiments are usually not reproducible using different PyTorch releases, commits, platforms or between CPU and GPU executions. Please find more details in the PyTorch documentation:
#
# https://pytorch.org/docs/stable/notes/randomness.html
# + colab_type="code" id="PQtgx_3H2MEx" colab={}
SEED = 0
t.manual_seed(SEED)
t.cuda.manual_seed(SEED)
t.backends.cudnn.deterministic = True
t.backends.cudnn.benchmark = False
np.random.seed(SEED)
random.seed(SEED)
# + [markdown] colab_type="text" id="ugovPCwFvBc4"
# **Loading Data**
# + [markdown] colab_type="text" id="8_PMRn9BcaPO"
# The dataset is structured in multiple small folders of 7 images each. This generator iterates through the folders and returns the category and 7 paths: one for each image in the folder. The paths are ordered; the order is important since each folder contains 3 types of images, first 5 are with acetic acid solution and the last two are through a green lens and having iodine solution(a solution of a dark red color).
# + colab_type="code" id="qU2qOUVWZZNl" colab={}
def sortByLastDigits(elem):
chars = [c for c in elem if c.isdigit()]
return 0 if len(chars) == 0 else int(''.join(chars))
def getImagesPaths(root_path):
for class_folder in [root_path + f for f in listdir(root_path)]:
category = int(class_folder[-1])
for case_folder in listdir(class_folder):
case_folder_path = class_folder + '/' + case_folder + '/'
img_files = [case_folder_path + file_name for file_name in listdir(case_folder_path)]
yield category, sorted(img_files, key = sortByLastDigits)
# + [markdown] colab_type="text" id="MSrKoW7TdRFr"
# We define 3 datasets, which load 3 kinds of images: natural images, images taken through a green lens and images where the doctor applied iodine solution (which gives a dark red color). Each dataset has dynamic and static transformations which could be applied to the data. The static transformations are applied on the initialization of the dataset, while the dynamic ones are applied when loading each batch of data.
# + colab_type="code" id="EmgWK_7-doGb" colab={}
class SimpleImagesDataset(t.utils.data.Dataset):
def __init__(self, root_path, transforms_x_static = None, transforms_x_dynamic = None, transforms_y_static = None, transforms_y_dynamic = None):
self.dataset = []
self.transforms_x = transforms_x_dynamic
self.transforms_y = transforms_y_dynamic
for category, img_files in getImagesPaths(root_path):
for i in range(5):
img = pil.Image.open(img_files[i])
if transforms_x_static != None:
img = transforms_x_static(img)
if transforms_y_static != None:
category = transforms_y_static(category)
self.dataset.append((img, category))
def __getitem__(self, i):
x, y = self.dataset[i]
if self.transforms_x != None:
x = self.transforms_x(x)
if self.transforms_y != None:
y = self.transforms_y(y)
return x, y
def __len__(self):
return len(self.dataset)
class GreenLensImagesDataset(SimpleImagesDataset):
def __init__(self, root_path, transforms_x_static = None, transforms_x_dynamic = None, transforms_y_static = None, transforms_y_dynamic = None):
self.dataset = []
self.transforms_x = transforms_x_dynamic
self.transforms_y = transforms_y_dynamic
for category, img_files in getImagesPaths(root_path):
# Only the green lens image
img = pil.Image.open(img_files[-2])
if transforms_x_static != None:
img = transforms_x_static(img)
if transforms_y_static != None:
category = transforms_y_static(category)
self.dataset.append((img, category))
class RedImagesDataset(SimpleImagesDataset):
def __init__(self, root_path, transforms_x_static = None, transforms_x_dynamic = None, transforms_y_static = None, transforms_y_dynamic = None):
self.dataset = []
self.transforms_x = transforms_x_dynamic
self.transforms_y = transforms_y_dynamic
for category, img_files in getImagesPaths(root_path):
# Only the green lens image
img = pil.Image.open(img_files[-1])
if transforms_x_static != None:
img = transforms_x_static(img)
if transforms_y_static != None:
category = transforms_y_static(category)
self.dataset.append((img, category))
# + [markdown] colab_type="text" id="vRqFvAHwvVTl"
# **Preprocess Data**
# + [markdown] colab_type="text" id="UOI_JvcCe0cR"
# Convert pytorch tensor to numpy array.
# + colab_type="code" id="OKxTUFnkezdb" colab={}
def to_numpy(x):
return x.cpu().detach().numpy()
# + [markdown] colab_type="text" id="TmnrMN5BmHRg"
# Data transformations for the test and training sets.
# + colab_type="code" id="_hBCnG-fvwjB" colab={}
norm_mean = [0.485, 0.456, 0.406]
norm_std = [0.229, 0.224, 0.225]
transforms_train = tv.transforms.Compose([
tv.transforms.RandomAffine(degrees = 45, translate = None, scale = (1., 2.), shear = 30),
# tv.transforms.CenterCrop(CROP_SIZE),
tv.transforms.Resize(IMAGE_SIZE),
tv.transforms.RandomHorizontalFlip(),
tv.transforms.ToTensor(),
tv.transforms.Lambda(lambda t: t.cuda()),
tv.transforms.Normalize(mean=norm_mean, std=norm_std)
])
transforms_test = tv.transforms.Compose([
# tv.transforms.CenterCrop(CROP_SIZE),
tv.transforms.Resize(IMAGE_SIZE),
tv.transforms.ToTensor(),
tv.transforms.Normalize(mean=norm_mean, std=norm_std)
])
y_transform = tv.transforms.Lambda(lambda y: t.tensor(y, dtype=t.long, device = 'cuda:0'))
# + [markdown] colab_type="text" id="5ky8gX31q-NW"
# Initialize pytorch datasets and loaders for training and test.
# + colab_type="code" id="2hqOyTVGnjjy" colab={}
def create_loaders(dataset_class):
dataset_train = dataset_class(TRAIN_PATH, transforms_x_dynamic = transforms_train, transforms_y_dynamic = y_transform)
dataset_test = dataset_class(TEST_PATH, transforms_x_static = transforms_test,
transforms_x_dynamic = tv.transforms.Lambda(lambda t: t.cuda()), transforms_y_dynamic = y_transform)
loader_train = DataLoader(dataset_train, BATCH_SIZE, shuffle = True, num_workers = 0)
loader_test = DataLoader(dataset_test, BATCH_SIZE, shuffle = False, num_workers = 0)
return loader_train, loader_test, len(dataset_train), len(dataset_test)
# + colab_type="code" id="Un_LZV7KqFCn" colab={}
loader_train_simple_img, loader_test_simple_img, len_train, len_test = create_loaders(SimpleImagesDataset)
# + [markdown] colab_type="text" id="yIcoJBpn0jZ8"
# **Visualize Data**
# + [markdown] colab_type="text" id="ARRQsL2drbzv"
# Load a few images so that we can see the effects of the data augmentation on the training set.
# + colab_type="code" id="mDi25EAxrSal" colab={}
def plot_one_prediction(x, label, pred):
x, label, pred = to_numpy(x), to_numpy(label), to_numpy(pred)
x = np.transpose(x, [1, 2, 0])
if x.shape[-1] == 1:
x = x.squeeze()
x = x * np.array(norm_std) + np.array(norm_mean)
plt.title(label, color = 'green' if label == pred else 'red')
plt.imshow(x)
def plot_predictions(imgs, labels, preds):
fig = plt.figure(figsize = (20, 5))
for i in range(20):
fig.add_subplot(2, 10, i + 1, xticks = [], yticks = [])
plot_one_prediction(imgs[i], labels[i], preds[i])
# + colab_type="code" id="Q0jkeNIVrVf3" colab={}
# x, y = next(iter(loader_train_simple_img))
# plot_predictions(x, y, y)
# + [markdown] colab_type="text" id="teYfBi5v0yuj"
# **Model**
# + [markdown] colab_type="text" id="TFAXychCsIFp"
# Define a few models to experiment with.
# + colab_type="code" id="3rQm0JTXsLOO" colab={}
def get_mobilenet_v2():
model = t.hub.load('pytorch/vision', 'mobilenet_v2', pretrained=True)
model.classifier[1] = Linear(in_features=1280, out_features=4, bias=True)
model = model.cuda()
return model
def get_vgg_19():
model = tv.models.vgg19(pretrained = True)
model = model.cuda()
model.classifier[6].out_features = 4
return model
def get_res_next_101():
model = t.hub.load('facebookresearch/WSL-Images', 'resnext101_32x8d_wsl')
model.fc.out_features = 4
model = model.cuda()
return model
def get_resnet_18():
model = tv.models.resnet18(pretrained = True)
model.fc.out_features = 4
model = model.cuda()
return model
def get_dense_net():
model = tv.models.densenet121(pretrained = True)
model.classifier.out_features = 4
model = model.cuda()
return model
class MobileNetV2_FullConv(t.nn.Module):
def __init__(self):
super().__init__()
self.cnn = get_mobilenet_v2().features
self.cnn[18] = t.nn.Sequential(
tv.models.mobilenet.ConvBNReLU(320, 32, kernel_size=1),
t.nn.Dropout2d(p = .7)
)
self.fc = t.nn.Linear(32, 4)
def forward(self, x):
x = self.cnn(x)
x = x.mean([2, 3])
x = self.fc(x);
return x
# + colab_type="code" executionInfo={"elapsed": 1674, "status": "ok", "timestamp": 1577543132882, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16049478229633186858"}, "user_tz": -120} id="DRM168aEuTid" outputId="e4af1f49-815b-41d0-de5e-f4285de26e93" colab={"base_uri": "https://localhost:8080/", "height": 1000}
model_simple = t.nn.DataParallel(get_res_next_101())
# + [markdown] colab_type="text" id="eVNbPOX31_rS"
# **Train & Evaluate**
# + [markdown] colab_type="text" id="6G1HQmJ1vJt8"
# Timer utility function. This is used to measure the execution speed.
# + colab_type="code" id="77HDFtMd1Ra_" colab={}
time_start = 0
def timer_start():
global time_start
time_start = time.time()
def timer_end():
return time.time() - time_start
# + [markdown] colab_type="text" id="2ZE3UqwMvbfM"
# This function trains the network and evaluates it at the same time. It outputs the metrics recorded during the training for both train and test. We are measuring accuracy and the loss. The function also saves a checkpoint of the model every time the accuracy is improved. In the end we will have a checkpoint of the model which gave the best accuracy.
# + colab_type="code" id="FwUA4abu2pRe" colab={}
def train_eval(optimizer, model, loader_train, loader_test, chekpoint_name, epochs):
metrics = {
'losses_train': [],
'losses_test': [],
'acc_train': [],
'acc_test': [],
'prec_train': [],
'prec_test': [],
'rec_train': [],
'rec_test': [],
'f_score_train': [],
'f_score_test': []
}
best_acc = 0
loss_fn = t.nn.CrossEntropyLoss()
try:
for epoch in range(epochs):
timer_start()
train_epoch_loss, train_epoch_acc, train_epoch_precision, train_epoch_recall, train_epoch_f_score = 0, 0, 0, 0, 0
test_epoch_loss, test_epoch_acc, test_epoch_precision, test_epoch_recall, test_epoch_f_score = 0, 0, 0, 0, 0
# Train
model.train()
for x, y in loader_train:
y_pred = model.forward(x)
loss = loss_fn(y_pred, y)
loss.backward()
optimizer.step()
# memory_stats()
optimizer.zero_grad()
y_pred, y = to_numpy(y_pred), to_numpy(y)
pred = y_pred.argmax(axis = 1)
ratio = len(y) / len_train
train_epoch_loss += (loss.item() * ratio)
train_epoch_acc += (sk.metrics.accuracy_score(y, pred) * ratio)
precision, recall, f_score, _ = sk.metrics.precision_recall_fscore_support(y, pred, average = 'macro')
train_epoch_precision += (precision * ratio)
train_epoch_recall += (recall * ratio)
train_epoch_f_score += (f_score * ratio)
metrics['losses_train'].append(train_epoch_loss)
metrics['acc_train'].append(train_epoch_acc)
metrics['prec_train'].append(train_epoch_precision)
metrics['rec_train'].append(train_epoch_recall)
metrics['f_score_train'].append(train_epoch_f_score)
# Evaluate
model.eval()
with t.no_grad():
for x, y in loader_test:
y_pred = model.forward(x)
loss = loss_fn(y_pred, y)
y_pred, y = to_numpy(y_pred), to_numpy(y)
pred = y_pred.argmax(axis = 1)
ratio = len(y) / len_test
test_epoch_loss += (loss * ratio)
test_epoch_acc += (sk.metrics.accuracy_score(y, pred) * ratio )
precision, recall, f_score, _ = sk.metrics.precision_recall_fscore_support(y, pred, average = 'macro')
test_epoch_precision += (precision * ratio)
test_epoch_recall += (recall * ratio)
test_epoch_f_score += (f_score * ratio)
metrics['losses_test'].append(test_epoch_loss)
metrics['acc_test'].append(test_epoch_acc)
metrics['prec_test'].append(test_epoch_precision)
metrics['rec_test'].append(test_epoch_recall)
metrics['f_score_test'].append(test_epoch_f_score)
if metrics['acc_test'][-1] > best_acc:
best_acc = metrics['acc_test'][-1]
t.save({'model': model.state_dict()}, 'checkpint {}.tar'.format(chekpoint_name))
print('Epoch {} acc {} prec {} rec {} f {} minutes {}'.format(
epoch + 1, metrics['acc_test'][-1], metrics['prec_test'][-1], metrics['rec_test'][-1], metrics['f_score_test'][-1], timer_end() / 60))
except KeyboardInterrupt as e:
print(e)
print('Ended training')
return metrics
# + [markdown] colab_type="text" id="BPznZeFDxhtA"
# Plot a metric for both train and test.
# + colab_type="code" id="4QjYASRPRnIr" colab={}
def plot_train_test(train, test, title, y_title):
plt.plot(range(len(train)), train, label = 'train')
plt.plot(range(len(test)), test, label = 'test')
plt.xlabel('Epochs')
plt.ylabel(y_title)
plt.title(title)
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="vVNLAgGOsbPB"
# Plot precision - recall curve
# + colab_type="code" id="WhtUUcgjsanm" colab={}
def plot_precision_recall(metrics):
plt.scatter(metrics['prec_train'], metrics['rec_train'], label = 'train')
plt.scatter(metrics['prec_test'], metrics['rec_test'], label = 'test')
plt.legend()
plt.title('Precision-Recall')
plt.xlabel('Precision')
plt.ylabel('Recall')
# + [markdown] colab_type="text" id="T8adJjmiCHs6"
# Train a model for several epochs. The steps_learning parameter is a list of tuples. Each tuple specifies the steps and the learning rate.
# + colab_type="code" id="1jO6UBhaP926" colab={}
def do_train(model, loader_train, loader_test, checkpoint_name, steps_learning):
for steps, learn_rate in steps_learning:
metrics = train_eval(t.optim.Adam(model.parameters(), lr = learn_rate, weight_decay = 0), model, loader_train, loader_test, checkpoint_name, steps)
print('Best test accuracy :', max(metrics['acc_test']))
plot_train_test(metrics['losses_train'], metrics['losses_test'], 'Loss (lr = {})'.format(learn_rate))
plot_train_test(metrics['acc_train'], metrics['acc_test'], 'Accuracy (lr = {})'.format(learn_rate))
# + [markdown] colab_type="text" id="Q91nZ33rCryC"
# Perform actual training.
# + colab_type="code" id="BA2tEyFmCybQ" colab={}
def do_train(model, loader_train, loader_test, checkpoint_name, steps_learning):
t.cuda.empty_cache()
for steps, learn_rate in steps_learning:
metrics = train_eval(t.optim.Adam(model.parameters(), lr = learn_rate, weight_decay = 0), model, loader_train, loader_test, checkpoint_name, steps)
index_max = np.array(metrics['acc_test']).argmax()
print('Best test accuracy :', metrics['acc_test'][index_max])
print('Corresponding precision :', metrics['prec_test'][index_max])
print('Corresponding recall :', metrics['rec_test'][index_max])
print('Corresponding f1 score :', metrics['f_score_test'][index_max])
plot_train_test(metrics['losses_train'], metrics['losses_test'], 'Loss (lr = {})'.format(learn_rate), 'Loss')
plot_train_test(metrics['acc_train'], metrics['acc_test'], 'Accuracy (lr = {})'.format(learn_rate), 'Accuracy')
plot_train_test(metrics['prec_train'], metrics['prec_test'], 'Precision (lr = {})'.format(learn_rate), 'Precision')
plot_train_test(metrics['rec_train'], metrics['rec_test'], 'Recall (lr = {})'.format(learn_rate), 'Recall')
plot_train_test(metrics['f_score_train'], metrics['f_score_test'], 'F1 Score (lr = {})'.format(learn_rate), 'F1 Score')
plot_precision_recall(metrics)
# + colab_type="code" executionInfo={"elapsed": 3635, "status": "error", "timestamp": 1577541623118, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16049478229633186858"}, "user_tz": -120} id="oY7_WOnucOJD" outputId="0eb5fa19-4762-4746-d738-db189a1c7790" colab={"base_uri": "https://localhost:8080/", "height": 296}
do_train(model_simple, loader_train_simple_img, loader_test_simple_img, 'simple_1', [(50, 1e-4)])
# + colab_type="code" id="H-ulUIj-Ertw" colab={}
# checkpoint = t.load('/content/checkpint simple_1.tar')
# model_simple.load_state_dict(checkpoint['model'])
| Mobilenetv2 Tuning/ResNext101 Baseline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/adgsenpai/PickNPayScrapping/blob/main/Scrapping_Pick_N_Pay_Products.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="CZHRRc8QERX5"
# 
# # Scrapping Pick N Pay Products to a .CSV File
#
# ### a notebook by <NAME>
# + [markdown] id="loYqMSo1FMG1"
# #### Introduction
#
# why did i do this? First of all my project idea is to take the nth food products and yth type of food products and generate a recipe/instructions for these items in an OpenAI.
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="2friJrQZ9JHy" outputId="3278f110-1a77-44c3-92b7-855e42b61ea9"
# Importing of Modules
import requests # this is to make https GET requests
import pandas as pd # This is used for a DataFrame
from bs4 import BeautifulSoup as bs #This is a module to parse HTML bodies
#global variable to init our prod and imglinks
prod = []
imglink = []
# this function parses the items
def ReturnItems(url,pagenumber):
items = []
for x in range(pagenumber+1):
if url[len(url)-1] == "#":
response= requests.get(url)
else:
response= requests.get(url+str(x))
soup = bs(response.content,"html.parser")
products = soup.findAll("div",{"id":"productCarouselItemContainer_000000000000127152_EA"})
items = items + soup.findAll("img")
return items
#all bad data might have other bad data but can fix it manually by human intervention
garbagedata = ['https://cdn-prd-02.pnp.co.za/sys-master/images/h34/h8b/10423060725790/app-logos.png','https://cdn-prd-02.pnp.co.za/sys-master/images/h26/h64/10206546133022/pick-n-pay-header2.png','/pnpstorefront/_ui/responsive/theme-blue/images/Icon-favourites-list-grey.svg','app-logos.png','https://cdn-prd-02.pnp.co.za/sys-master/images/h2c/h9c/10269822877726/footer-pnp-white.png','footer-pnp-white.png','pick-n-pay-header2.png']
#Get links from here https://www.pnp.co.za/pnpstorefront/pnp/en/All-Products/c/pnpbase for categories (we only looking for food products/alcohol) - some manual intervention went here
items = ReturnItems('https://www.pnp.co.za/pnpstorefront/pnp/en/All-Products/Fresh-Fruit-%26-Vegetables/c/fresh-fruit-and-vegetables-423144840?q=%3Arelevance&pageSize=72&page=',2)
items += ReturnItems('https://www.pnp.co.za/pnpstorefront/pnp/en/All-Products/Bakery/c/bakery-423144840?q=%3Arelevance&pageSize=72&page=',1)
items += ReturnItems('https://www.pnp.co.za/pnpstorefront/pnp/en/All-Products/Milk%2C-Dairy-%26-Eggs/c/milk-dairy-and-eggs-423144840?q=%3Arelevance&pageSize=72&page=',4)
items += ReturnItems('https://www.pnp.co.za/pnpstorefront/pnp/en/All-Products/Meat%2C-Poultry-%26-Seafood/c/meat-poultry-and-seafood-423144840?pageSize=72&q=%3Arelevance&show=Page#',0)
items += ReturnItems('https://www.pnp.co.za/pnpstorefront/pnp/en/All-Products/Ready-Meals-%26-Desserts/c/ready-meals-and-desserts-423144840?pageSize=72&q=%3Arelevance&show=Page#',0)
items += ReturnItems('https://www.pnp.co.za/pnpstorefront/pnp/en/All-Products/Frozen-Food/c/frozen-food-423144840?q=%3Arelevance&pageSize=72&page=',3)
items += ReturnItems('https://www.pnp.co.za/pnpstorefront/pnp/en/All-Products/Chocolates%2C-Chips-%26-Snacks/c/chocolates-chips-and-snacks-423144840?q=%3Arelevance&pageSize=18&page=',8)
items += ReturnItems('https://www.pnp.co.za/pnpstorefront/pnp/en/All-Products/Food-Cupboard/c/food-cupboard-423144840?q=%3Arelevance&pageSize=72&page=',28)
items += ReturnItems('https://www.pnp.co.za/pnpstorefront/pnp/en/All-Products/Beverages/c/beverages-423144840?q=%3Arelevance&pageSize=72&page=',13)
items += ReturnItems('https://www.pnp.co.za/pnpstorefront/pnp/en/All-Products/Wine/c/wine-423144840?q=%3Arelevance&pageSize=72&page=',14)
items += ReturnItems('https://www.pnp.co.za/pnpstorefront/pnp/en/All-Products/Spirits/c/spirits-423144840?q=%3Arelevance&pageSize=72&page=',5)
# this is a for loop to append the metadata into the lists
for img in items:
if img.has_attr('title'):
if img['title'] in garbagedata:
pass
else:
prod.append(img['title'])
if img.has_attr('src'):
if img['src'] in garbagedata:
pass
else:
imglink.append(img['src'])
#Our Dataframe to display our stuff
df = pd.DataFrame(list(zip(prod, imglink)),
columns =['Product', 'ImageLink'])
df
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="fx7E3_q9RbTN" outputId="915a3bb8-90fb-401b-b1ac-0bf8c0441380"
# Viewing top 5 items in dataset
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="hgGInu_pRhpG" outputId="e8f0cafd-62bd-4125-a494-9d5607c93f4b"
# Saving our dataset to a .csv
try:
df.to_csv('pnpfoodprod.csv')
print('file saved ...')
except Exception as e:
print('failed: ',e)
# + colab={"base_uri": "https://localhost:8080/"} id="Wov1gacKRykk" outputId="038833dd-590a-4370-cf68-c7ad23a8cc85"
# to dict can be used as a RestAPI maybe - if im free could put this in a RestAPI on my website ADGSTUDIOS.co.za
df.to_dict()
# + [markdown] id="hpvLvsAySC6o"
# ##### Copyright (c) ADGSTUDIOS 2021 All Rights Reserved
| Scrapping_Pick_N_Pay_Products.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import pyarrow.parquet as pq
df = pq.read_table(source="ml(1).parquet").to_pandas()
pd.set_option('display.max_columns', 500)
df
df[["mean_flag","counts","month_mean"]][600:620]
df['created_date_year'] = pd.DatetimeIndex(df['created_date']).year
df['created_date_month'] = pd.DatetimeIndex(df['created_date']).month
df['created_date_day'] = pd.DatetimeIndex(df['created_date']).day
df['created_date_dow'] = pd.DatetimeIndex(df['created_date']).dayofweek
# Creamos la variable created_date_woy (numero de la semana del año de la fecha created_date)
df['created_date_woy'] = pd.DatetimeIndex(df['created_date']).week
def festivo(date):
'''
Recorre el arreglo de dias festivos e indica si un dia es festivos
Funcion auxiliar a create_feature_table
'''
import datetime
h = ['2010-01-01', '2010-12-31', '2010-01-18', '2010-02-15', '2010-05-31', '2010-07-04', '2010-07-05', '2010-09-06', '2010-10-11', '2010-11-11', '2010-11-25', '2010-12-25', '2010-12-24', '2011-01-01', '2010-12-31', '2011-01-17', '2011-02-21', '2011-05-30', '2011-07-04', '2011-09-05', '2011-10-10', '2011-11-11', '2011-11-24', '2011-12-25', '2011-12-26', '2012-01-01', '2012-01-02', '2012-01-16', '2012-02-20', '2012-05-28', '2012-07-04', '2012-09-03', '2012-10-08', '2012-11-11', '2012-11-12', '2012-11-22', '2012-12-25', '2013-01-01', '2013-01-21', '2013-02-18', '2013-05-27', '2013-07-04', '2013-09-02', '2013-10-14', '2013-11-11', '2013-11-28', '2013-12-25', '2014-01-01', '2014-01-20', '2014-02-17', '2014-05-26', '2014-07-04', '2014-09-01', '2014-10-13', '2014-11-11', '2014-11-27', '2014-12-25', '2015-01-01', '2015-01-19', '2015-02-16', '2015-05-25', '2015-07-04', '2015-07-03', '2015-09-07', '2015-10-12', '2015-11-11', '2015-11-26', '2015-12-25', '2016-01-01', '2016-01-18', '2016-02-15', '2016-05-30', '2016-07-04', '2016-09-05', '2016-10-10', '2016-11-11', '2016-11-24', '2016-12-25', '2016-12-26', '2017-01-01', '2017-01-02', '2017-01-16', '2017-02-20', '2017-05-29', '2017-07-04', '2017-09-04', '2017-10-09', '2017-11-11', '2017-11-10', '2017-11-23', '2017-12-25', '2018-01-01', '2018-01-15', '2018-02-19', '2018-05-28', '2018-07-04', '2018-09-03', '2018-10-08', '2018-11-11', '2018-11-12', '2018-11-22', '2018-12-25', '2019-01-01', '2019-01-21', '2019-02-18', '2019-05-27', '2019-07-04', '2019-09-02', '2019-10-14', '2019-11-11', '2019-11-28', '2019-12-25', '2020-01-01', '2020-01-20', '2020-02-17', '2020-05-25', '2020-07-04', '2020-07-03', '2020-09-07', '2020-10-12', '2020-11-11', '2020-11-26', '2020-12-25']
h=pd.to_datetime(h).date
for festive in h:
if(date==festive):
return 1
return 0
df["date_holiday"]=df["created_date"].apply(festivo)
history_days=10
#fecha de inicio
for i in range(1,history_days):
for j in range(len(df["counts"])):
if(j==0):
var_name = f"number_cases_{i}_days_ago"
df[var_name]=0
if(j<i):
df[var_name][j]=0
else:
df[var_name][j]=df["counts"][(j-i)]
means=df.loc[:,['created_date_month','counts']]
means=means.groupby(['created_date_month'],as_index=False).mean()
means.columns=['created_date_month','month_mean']
import numpy as np
df=pd.merge(df,means,how='left',on=["created_date_month"])
df["flagt"]=0
index=df["flagt"].where(df["counts"]>df["month_mean"],1)
#df["flagt"][index[0]]=1
df["f"]= df["counts"] - df["month_mean"]
# +
def flags(x):
if(x<0):
return -1
else:
return 0
df["f"]=df["f"].apply(flags)
# -
df
dummies=pd.get_dummies(df["created_date_month"],prefix='y')
df=pd.concat([df,dummies],axis=1)
df
df=df.drop(["created_date_year","created_date_month","created_date_day","created_date_dow","created_date_woy"],axis=1)
percentiles_classes = [.1, .3, .6]
y = np.hstack([[ii] * int(100 * perc) for ii, perc in enumerate(percentiles_classes)])
list(enumerate(percentiles_classes))
groups = np.hstack([[ii] * 10 for ii in range(10)])
groups
# # Modelado
df
y=df["mean_flag"]
df2=df.drop(columns=["mean_flag","created_date","counts"])
df2
from sklearn.model_selection import TimeSeriesSplit
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
#primero separamos los datos de entrenamiento y prueba
x_train, x_test, y_train, y_test= train_test_split(df2,y,test_size=0.8)
#separamos los primeros 70% de los datos para entrenar
X_train = df2[:int(df2.shape[0]*0.7)].to_numpy()
X_test = df2[int(df2.shape[0]*0.7):].to_numpy()
y_train = y[:int(df2.shape[0]*0.7)].to_numpy()
y_test = y[int(df2.shape[0]*0.7):].to_numpy()
X_train.shape
# +
#partimos los datos con temporal cv
tscv=TimeSeriesSplit(n_splits=5)
for tr_index, val_index in tscv.split(X_train):
X_tr, X_val=X_train[tr_index], X_train[val_index]
y_tr, y_val = y_train[tr_index], y_train[val_index]
# -
model=RandomForestClassifier(max_depth=10,criterion='gini',n_estimators=100,n_jobs=-1)
model.fit(X_tr,y_tr)
#PRedecimos sobre datos de validacion
preds=model.predict(X_val)
model.score(X_train,y_train)
model.score(X_val,y_val)
model.score(X_test,y_test)
1-sum(abs(preds-y_val))/len(preds)
#guarda pickle
import pickle
modelo=pickle.dumps(model)
#load pickle
desde_pickle=pickle.loads(modelo)
#usa el pickle
desde_pickle.score(X_test,y_test)
| scripts/model/model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from pathlib import Path
import pickle
path = Path('/home/gaurav/PycharmProjects/nlp-for-panjabi/datasets-preparation/panjabi-wikipedia-dataset')
p = path.glob('panjabi-wikipedia-articles/*')
files = [x for x in p if x.is_file()]
len(files)
files[0]
text = ''
for file in files:
with open(file, 'rb') as f:
text += pickle.load(f)
len(text)
with open('panjabi.txt', 'w') as f:
f.write(text)
with open('panjabi.txt', 'r') as f:
t = f.read()
len(t)
| dataset-preparation/collect_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: mypydev
# language: python
# name: mypydev
# ---
from splinter import Browser
from bs4 import BeautifulSoup
from webdriver_manager.chrome import ChromeDriverManager
executable_path = {'executable_path': ChromeDriverManager().install()}
browser = Browser('chrome', **executable_path, headless=False)
url = 'http://books.toscrape.com/'
browser.visit(url)
# +
html = browser.html
soup = BeautifulSoup(html, 'html.parser')
sidebar = soup.find('ul', class_='nav-list')
categories = sidebar.find_all('li')
category_list = []
url_list = []
book_url_list = []
for category in categories:
title = category.text.strip()
category_list.append(title)
book_url = category.find('a')['href']
url_list.append(book_url)
book_url_list = ['http://books.toscrape.com/' + url for url in url_list]
titles_and_urls = zip(category_list, book_url_list)
try:
for title_url in titles_and_urls:
browser.links.find_by_partial_text('next').click()
except ElementDoesNotExist:
print("Scraping Complete")
# -
book_url_list
browser.quit()
| 01-Lesson-Plans/12-Web-Scraping-and-Document-Databases/2/Activities/08-Stu_Splinter/Solved/Stu_Splinter_Advanced.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Тестирование на нормальном распределении
# ## 1. Генерация данных
# +
import numpy as np
import matplotlib.pyplot as plt
import sys
sys.path.append('../metric')
sys.path.append('../util')
import nn
import metrics
import loss
import train_set
# -
n = 5
size = 30
mus= np.array([[-1, 1], [-1, 0], [0, -1], [1, 0], [1, 1]])
sigmas = np.array([[0.5, 0.5], [0.5, 0.5],
[0.5, 0.5], [0.5, 0.5], [0.5, 0.5]])
train_xs, test_xs, train_ys, test_ys = train_set.gauss_cls(n, size, 0.3,
mus, sigmas)
train_len = np.size(train_xs, axis=0)
test_len = np.size(test_xs, axis=0)
# ## 2. kNN
# ### 2.1 Классификация
# +
k = 20
color = ['r', 'g', 'b', 'y', 'm']
classes = np.zeros(test_len, dtype=np.int)
for i in range(test_len):
cls, results = nn.knn(n, k,
train_xs, train_ys,
test_xs[i, :],
metrics.euclidean)
classes[i] = cls
def plot(train_xs, train_ys,
test_xs, test_ys,
classes, color):
colors = [color[train_ys[i]] for i in range(train_len)]
plt.scatter(train_xs[:, 0], train_xs[:, 1], c=colors, marker='+')
plt.scatter(test_xs[:, 0],
test_xs[:, 1],
c=[color[cls] for cls in classes],
marker='^')
plt.scatter(test_xs[:, 0],
test_xs[:, 1],
c=[color[cls] for cls in test_ys],
marker='*')
plot(train_xs, train_ys, test_xs, test_ys, classes, color)
# -
loss.empirical_risk_cls(classes, test_ys)
# ### 2.2 Поиск оптимального k
# +
def a(n, k, xs, ys, u):
return nn.knn(n, k, xs, ys, u, metrics.euclidean)
opt_k, ks = nn.leave_one_out(n, train_len, a, train_xs, train_ys)
# -
opt_k
plt.plot(np.arange(0, np.size(ks, axis=0), 1), ks)
# +
classes = np.zeros(test_len, dtype=np.int)
for i in range(test_len):
cls, results = nn.knn(n, opt_k,
train_xs, train_ys,
test_xs[i, :],
metrics.euclidean)
classes[i] = cls
plot(train_xs, train_ys, test_xs, test_ys, classes, color)
# -
loss.empirical_risk_cls(classes, test_ys)
# ## 3. Метод парзеновского окна
# ### 3.1 Классификация
# +
k = 20
classes = np.zeros(test_len, dtype=np.int)
for i in range(test_len):
cls, results = nn.knn_parzen(n, k,
train_xs, train_ys,
test_xs[i, :],
metrics.euclidean)
classes[i] = cls
plot(train_xs, train_ys, test_xs, test_ys, classes, color)
# -
loss.empirical_risk_cls(classes, test_ys)
# ### 3.2 Выбор оптимального k
# +
def a(n, k, xs, ys, u):
return nn.knn_parzen(n, k, xs, ys, u, metrics.euclidean)
opt_k, ks = nn.leave_one_out(n, train_len - 1, a, train_xs, train_ys)
# -
opt_k
plt.plot(np.arange(0, np.size(ks, axis=0), 1), ks)
classes = np.zeros(test_len, dtype=np.int)
for i in range(test_len):
cls, results = nn.knn_parzen(n, opt_k,
train_xs, train_ys,
test_xs[i, :],
metrics.euclidean)
classes[i] = cls
plot(train_xs, train_ys, test_xs, test_ys, classes, color)
loss.empirical_risk_cls(classes, test_ys)
# ## 4. Отбор эталонов
# ### 4.1 Отступы
# +
def a(u):
return nn.knn_parzen(n, opt_k, train_xs, train_ys, u, metrics.euclidean)
margins = nn.margin(n, a, train_xs, train_ys)
margins.sort()
# -
plt.plot(np.arange(np.size(margins, axis=0)), margins)
plt.grid()
# ### 4.2 Сжатие обучающего множества алгоритмом STOLP
# +
def a(u, k, xs, ys):
return nn.knn_parzen(n, k, xs, ys, u, metrics.euclidean)
compr_train_xs, compr_train_ys = \
nn.stolp(n, opt_k, 0, 0.15 * train_len, a, train_xs, train_ys)
# -
# Размеры обучающего множества до и после сжатия:
train_len, np.size(compr_train_xs, axis=0)
# Классификация и эмпирический риск:
classes = np.zeros(test_len, dtype=np.int)
for i in range(test_len):
cls, results = nn.knn_parzen(n, n + 10,
compr_train_xs, compr_train_ys,
test_xs[i, :],
metrics.euclidean)
classes[i] = cls
plot(train_xs, train_ys, test_xs, test_ys, classes, color)
loss.empirical_risk_cls(classes, test_ys)
| test/kNN-gauss-test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (taxparams-dev)
# language: python
# name: taxparams-dev
# ---
# +
import taxcalc
import taxparams
# -
recs = taxcalc.Records.cps_constructor()
class TaxParams(taxparams.TaxParams):
"""
TaxParams class that implements the methods taxcalc.Calculator
expects it to have. Note that you need to change these lines of
code on Tax-Calculator:
https://github.com/PSLmodels/Tax-Calculator/blob/2.5.0/taxcalc/calculator.py#L96-L99
"""
def set_year(self, year):
self.set_state(year=year)
for name in self._data:
arr = getattr(self, name)
setattr(self, name, arr[0])
@property
def current_year(self):
return self.label_grid["year"][0]
@property
def start_year(self):
return self._stateless_label_grid["year"][0]
@property
def end_year(self):
return self._stateless_label_grid["year"][-1]
@property
def parameter_warnings(self):
return self.errors
calc1 = taxcalc.Calculator(policy=TaxParams(), records=recs)
calc1.advance_to_year(2020)
calc1.calc_all()
# +
params2 = TaxParams()
params2.adjust(
{
"EITC_c": [
{"year": 2020, "EIC": "0kids", "value": 10000},
{"year": 2020, "EIC": "1kid", "value": 10001},
{"year": 2020, "EIC": "2kids", "value": 10002},
{"year": 2020, "EIC": "3+kids", "value": 10003},
]
}
)
calc2 = taxcalc.Calculator(policy=params2, records=recs)
calc2.advance_to_year(2020)
calc2.calc_all()
# -
calc2.difference_table(calc1, "weighted_deciles", "combined")
| TaxParams w taxcalc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''CSE499'': conda)'
# name: python3
# ---
# + papermill={"duration": 2.431982, "end_time": "2021-08-21T14:55:20.463183", "exception": false, "start_time": "2021-08-21T14:55:18.031201", "status": "completed"} tags=[]
import os
import gc
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from pytorch_tabnet.tab_model import TabNetClassifier
import torch
from sklearn.impute import SimpleImputer
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
# + papermill={"duration": 0.018291, "end_time": "2021-08-21T14:55:20.492942", "exception": false, "start_time": "2021-08-21T14:55:20.474651", "status": "completed"} tags=[]
DATA_DIRECTORY = ""
# + papermill={"duration": 52.375964, "end_time": "2021-08-21T14:56:12.880392", "exception": false, "start_time": "2021-08-21T14:55:20.504428", "status": "completed"} tags=[]
train = pd.read_csv(os.path.join(DATA_DIRECTORY, 'train.csv'))
test = pd.read_csv(os.path.join(DATA_DIRECTORY, 'test.csv'))
labels = pd.read_csv(os.path.join(DATA_DIRECTORY, 'labels.csv'))
# -
test_id = test['SK_ID_CURR']
# + papermill={"duration": 39.722848, "end_time": "2021-08-21T14:56:52.614854", "exception": false, "start_time": "2021-08-21T14:56:12.892006", "status": "completed"} tags=[]
imputer = SimpleImputer(strategy = 'median')
imputer.fit(train)
train = imputer.transform(train)
test = imputer.transform(test)
# + papermill={"duration": 0.2603, "end_time": "2021-08-21T14:56:52.886786", "exception": true, "start_time": "2021-08-21T14:56:52.626486", "status": "failed"} tags=[]
target = labels.to_numpy()
del labels
gc.collect()
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
x_train, x_val, y_train, y_val = train_test_split(train, target, test_size=0.30, random_state=8)
# -
def accuracy_score(y_true, y_pred):
y_pred = np.concatenate(tuple(y_pred))
y_true = np.concatenate(tuple([t for t in y] for y in y_true)).reshape(
y_pred.shape
)
return (y_true == y_pred).sum() / float(len(y_true))
# +
from sklearn.model_selection import ParameterGrid
from sklearn.metrics import roc_auc_score
# Function that instantiates a tabnet model.
def create_tabnet(n_d=32, n_steps=5, lr=0.02, gamma=1.5,
n_independent=2, n_shared=2, lambda_sparse=1e-4,
momentum=0.3, clip_value=2.):
return TabNetClassifier(
n_d=n_d, n_a=n_d, n_steps=n_steps,
gamma=gamma, n_independent=n_independent, n_shared=n_shared,
lambda_sparse=lambda_sparse, momentum=momentum, clip_value=clip_value,
optimizer_fn=torch.optim.Adam,
scheduler_params = {"gamma": 0.95,
"step_size": 20},
scheduler_fn=torch.optim.lr_scheduler.StepLR, epsilon=1e-15, verbose = 0
)
# Generate the parameter grid.
param_grid = dict(n_d = [8, 16, 32, 64],
n_steps = [3, 4, 5],
gamma = [1, 1.5, 2],
lambda_sparse = [1e-2, 1e-3, 1e-4],
momentum = [0.3, 0.4, 0.5],
n_shared = [2],
n_independent = [2],
clip_value = [2.],
)
grid = ParameterGrid(param_grid)
search_results = pd.DataFrame()
for params in grid:
params['n_a'] = params['n_d'] # n_a=n_d always per the paper
tabnet = create_tabnet()
tabnet.set_params(**params)
tabnet.fit(
x_train,y_train[:,0],
eval_set=[(x_train, y_train[:,0]), (x_val, y_val[:,0])],
eval_name=['train', 'valid'],
eval_metric=['auc'],
max_epochs=1000 , patience=50,
batch_size=256, virtual_batch_size=128,
num_workers=0,
weights=1,
drop_last=False
)
y_prob = tabnet.predict_proba(x_val)
auc = roc_auc_score(y_val, y_prob[:, 1])
score = max(2*auc - 1, 0.)
# score = accuracy_score(y_val[:, 0], y_prob[: 1])
results = pd.DataFrame([params])
results['score'] = np.round(score, 3)
search_results = search_results.append(results)
# -
search_results.to_csv(os.path.join(DATA_DIRECTORY, 'search_results.csv'), index=False)
| notebooks/CSE499B/sakib/home-credit-loan-tabnet copy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fast Audio Clip Dataset for PyTorch
# - Reads WAV audio at ~10M samples/sec (from an SSD; no resampling)
# - Splits audio files into short clips (very fast)
# - Supports resampling and downmixing to mono
# - Supports WAV, FLAC, OGG audio
# %load_ext autoreload
# %autoreload 2
# ## Import dependencies
# +
from time import time
from tqdm.auto import tqdm
from pathlib import Path
from IPython.display import display, Audio
from torch.utils.data import DataLoader
from beatbrain.datasets.audio import AudioClipDataset
# -
# ## Define constants
AUDIO_DIR = Path("../data/edm/wav/")
MAX_SEGMENT_LENGTH = 5
MIN_SEGMENT_LENGTH = 5
SAMPLE_RATE = 22050
MONO = True
# ## Create dataset
dataset = AudioClipDataset(AUDIO_DIR, max_segment_length=MAX_SEGMENT_LENGTH, min_segment_length=MIN_SEGMENT_LENGTH, sample_rate=SAMPLE_RATE, mono=MONO)
# ## Preview audio clip
audio, sr = dataset[0]
display(Audio(audio, rate=sr))
# ## Benchmark read performance
dataloader = DataLoader(dataset, batch_size=32, shuffle=True, pin_memory=True)
start = time()
for audio, sr in tqdm(dataloader):
pass
elapsed = time() - start
print(f"Total time: {elapsed:.2f}s")
print(f"{len(dataset) * SAMPLE_RATE / (elapsed * 1e6):.5f} M samples/sec")
| notebooks/audio_clip_dataset_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={} tags=[]
# <img width="10%" alt="Naas" src="https://landen.imgix.net/jtci2pxwjczr/assets/5ice39g4.png?w=160"/>
# + [markdown] papermill={} tags=[]
# # Github - Create issue
# <a href="https://app.naas.ai/user-redirect/naas/downloader?url=https://raw.githubusercontent.com/jupyter-naas/awesome-notebooks/master/Github/Github_Create_issue.ipynb" target="_parent"><img src="https://naasai-public.s3.eu-west-3.amazonaws.com/open_in_naas.svg"/></a>
# + [markdown] papermill={} tags=[]
# **Tags:** #github #productivity #code
# + [markdown] papermill={} tags=[]
# ## Input
# + [markdown] papermill={} tags=[]
# ### Import library
# + papermill={} tags=[]
from github import Github
# + [markdown] papermill={} tags=[]
# ### Enter repository path and token
# + papermill={} tags=[]
repo_name = "**********" # Repository path
git_key = "**********" # Settings/Developer settings
assignee = "**********" # Asignee name (optional) or put ""
issue_title = "This is a another issue" # Issue title
issue_description = "This is another issue body created using api" # Issue description
# + [markdown] papermill={} tags=[]
# ## Model
# + [markdown] papermill={} tags=[]
# ### Establishing connection
# + papermill={} tags=[]
g = Github(git_key)
# + [markdown] papermill={} tags=[]
# ## Output
# + [markdown] papermill={} tags=[]
# ### Creating github issue with assignee
# + papermill={} tags=[]
repo = g.get_repo(repo_name)
repo.create_issue(title = issue_title, body = issue_description, assignee = assignee)
| Github/Github_Create_issue.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using Pre-Training Models to Classify Objects in Photographs
# # Sources
# * [How to Use The Pre-Trained VGG Model to Classify Objects in Photographs](https://machinelearningmastery.com/use-pre-trained-vgg-model-classify-objects-photographs/)
| notebooks/Object Detection & Classification/Using Pre-Training Models to Classify Objects in Photographs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Emission Intensity
#
# This notebook shows how to create a chart similar to the infographics [Emission intensities of economies](https://faktaoklimatu.cz/infografiky/emisni-intenzity):
#
# 
#
# The chart is very complex – it shows:
# - greenhouse gases emissions per GDP along x-axis (g CO2eq / \$) – this is called **emission intensity of the economy**
# - GDP per capita along y-axis (\$ per capita, expressed in constant international 2011 dollars)
# - the product of x-value and y-value is emissions per capita – so countries with the same level of emissions per capita are along the same hyperbola (expressed in tonnes of CO2eq per capita per year)
# - the sizes of bubbles correspond to total population, color coding shows different continents
#
# More detailed description (in Czech only) can be found on the page showing this infographic.
# ## Load prerequisities
# It is necessary to install all the required packages before the first use (and it has to be done only once for all notebooks). Then we can import the libraries (and set up better plotting settings via matplotlibrc file).
# !pip install -r requirements.txt
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
mpl.rc_file_defaults()
import world_bank_data as wb
# ## Prepare the dataset
#
# The chart combines three variables:
# - population
# - sizes of economics (GDP)
# - greenhouse gases emissions
#
# In addition, we need labels for all countries and information in what regions the countries are located. The most recent greenhouse gas emissions data are available for 2015, so we need all data sources only for this year.
#
# The final dataset is stored under `data/world-emissions-2015-regions.csv`, so you can skip the preparation and go straight to plotting if you wish.
# ### Population data
#
# The population data is loaded from World Bank via their [python library](https://pypi.org/project/world-bank-data/). The population data is stored under `SP.POP.TOTL` identifier. The World Bank uses ISO 3166 alpha-3 as country identifiers.
pop_raw = wb.get_series('SP.POP.TOTL', id_or_value='id')
pop = pop_raw.reset_index().query('Year == "2015"')[['Country', 'SP.POP.TOTL']].rename(columns={'Country': 'code', 'SP.POP.TOTL': 'pop'})
pop = pop[np.isfinite(pop['pop'])] # there are two countries with no population data
pop['pop'] = np.int_(pop['pop'])
pop = pop.sort_values('code').reset_index(drop=True)
pop.head()
# ### GDP data
#
# GDP data is loaded from World Bank too. We use `NY.GDP.MKTP.PP.KD` series, containing GDP PPP, constant 2017 international \$.
gdp_raw = wb.get_series('NY.GDP.MKTP.PP.KD', id_or_value='id')
gdp = gdp_raw.reset_index().query('Year == "2015"')[['Country', 'NY.GDP.MKTP.PP.KD']] \
.rename(columns={'Country': 'code', 'NY.GDP.MKTP.PP.KD': 'gdp'}) \
.sort_values('code') \
.reset_index(drop=True)
gdp.head()
# The World Bank changed this particular series recently. It used to contain data expressed in constant 2011 international \\$ and our infographic is based on this older version. The comparison of different countries is slightly changed with the updated GDP data (overall results are preserved of course). If you would like to get identical chart to our infographic, you can load the GDP data in constant 2011 international \\$ from `data/world-bank/NY.GDP.MKTP.PP.KD.2011.csv`.
# +
# gdp = pd.read_csv('../data/world-bank/NY.GDP.MKTP.PP.KD.2011.csv', skiprows=3)[['Country Code', '2015']] \
# .rename(columns={'Country Code': 'code', '2015': 'gdp'})
# -
# ### Greenhouse gases emissions data
#
# The World Bank publishes emissions data too, in series `EN.ATM.GHGT.KT.CE`. Unfortunately the series does not contain newer data than 2012, so it is better to use a specialized EDGAR database provided by Joint Research Centre of European Commission. EDGAR stands for Emissions Database for Global Atmospheric Research and the newest data is published under [EDGAR v5.0](https://edgar.jrc.ec.europa.eu/overview.php?v=50_GHG). The link is frequently unavailable, so we downloaded the emission data files into `data\edgar\v5.0`. EDGAR also provides more [detailed description](https://edgar.jrc.ec.europa.eu/overview.php?v=50_readme) of the dataset.
#
# We will use three XLS files from `data\edgar\v5.0`:
# - `v50_CO2_excl_short-cycle_org_C_1970_2018.xls` – CO2 emissions, excluding short cycles, for 1970–2018
# - `v50_CH4_1970_2015.xls` – CH4 emissions for 1970–2015
# - `v50_N2O_1970_2015.xls` – NO2 emissions for 1970–2015
#
# Conversion to CO2 equivalent is based on [IPCC AR5](https://www.ipcc.ch/assessment-report/ar5/), with coefficients 28 for CH4 and 265 for NO2. The total emissions do not include some marginal greenhouse gases as these are not provided in EDGAR. The values are in Gg (Gigagrams), i.e. thousand tonnes.
# +
ghgs = ['CO2', 'CH4', 'N2O']
edgar_files = ['CO2_excl_short-cycle_org_C', 'CH4', 'N2O']
edgar = None
for gas in ghgs:
ef = 'CO2_excl_short-cycle_org_C' if gas == 'CO2' else gas
ey = 2018 if gas == 'CO2' else 2015
filename = f'../data/edgar/v5.0/v50_{ef}_1970_{ey}.xls'
frame = pd.read_excel(filename, sheet_name='TOTALS BY COUNTRY', header=9)
frame = frame[['ISO_A3'] + list(range(1970, ey + 1))].rename(columns={'ISO_A3': 'code'}).set_index('code')
frame.columns = frame.columns.rename('year')
frame = frame.unstack().rename(gas).reset_index()
frame = frame[~frame['code'].isin(['SEA', 'AIR'])]
if edgar is None:
edgar = frame
else:
edgar = pd.merge(edgar, frame, how='outer')
# -
edgar.head()
ghg = edgar.query('year == 2015').drop(columns=['year']).sort_values('code').reset_index(drop=True)
ghg['ghg'] = ghg['CO2'] + 28 * ghg['CH4'] + 265 * ghg['N2O']
ghg.head()
# ### Regional data
#
# As we frequently needed to aggregate countries to larger regions, we created our regional classification that allows us to group countries easily based on ISO 3166 codes. The regional classification is stored in `data/regions.csv` and we will use `region_B_en` and `continent_en` in this notebook (note that the continents actually contain Russia as a standalone category, as it overlaps both Europe and Asia).
regions = pd.read_csv('../data/regions.csv', keep_default_na=False)[['code', 'region_B_en', 'continent_en']] \
.rename(columns={'region_B_en': 'region', 'continent_en': 'continent'}) \
.query('region != ""')
regions.head()
# ### Merge all datasets
#
# The last step is to merge all the prepared pieces together and aggregate at regional levels.
countries = pd.merge(regions, pop)
countries = pd.merge(countries, gdp)
countries = pd.merge(countries, ghg[['code', 'ghg']])
# This dataset is stored under `data/world-emissions-2015.csv`.
# +
# countries.to_csv('../data/world-emissions-2015.csv', index=False)
# -
countries.head()
# Some countries might have missing data for GDP or for greenhouse gases emissions. Check out how large part of data it is and then filter them out.
print('Overall population is', countries['pop'].sum())
print('After NaNs removed, population is', countries.dropna()['pop'].sum())
countries = countries.dropna().reset_index(drop=True)
# So the missing data is related with slightly more than 100 million people, that is less than 2% of population. It is acceptable to remove this part of data.
# we will merge continents back after the aggregation
continents = countries[['region', 'continent']].drop_duplicates().reset_index(drop=True)
df = countries.groupby('region')[['pop', 'gdp', 'ghg']].sum().reset_index()
df = pd.merge(df, continents)
# make the units more sane - and create per capita and per gdp columns
df['ghg'] = df['ghg'] / 1e3 # in million tonnes (rather than in gigagrams, i.e. thousand tonnes)
df['gdp_per_capita'] = df['gdp'] / df['pop']
df['ghg_per_capita'] = 1e6 * df['ghg'] / df['pop'] # in tonnes per capita
df['ghg_per_gdp'] = 1e12 * df['ghg'] / df['gdp'] # in grams per dollar
# This dataset is stored under `world-emissions-2015-regions.csv`.
# +
# df.to_csv('../data/world-emissions-2015-regions.csv', index=False)
# -
# ## Plot the results
#
# We can just load the final dataset, so it's possible to skip its creation.
df = pd.read_csv('../data/world-emissions-2015-regions.csv')
df.head()
# Calculation of world averages (for gdp / capita, ghg / capita and ghg / gdp, together with appropriate unit conversions):
avg_gdp_per_capita = df['gdp'].sum() / df['pop'].sum()
avg_ghg_per_capita = 1e6 * df['ghg'].sum() / df['pop'].sum()
avg_ghg_per_gdp = 1e12 * df['ghg'].sum() / df['gdp'].sum()
# We need to define color schema for different continents:
color_schema = {
'Asia': '#c32b2a',
'North America': '#2d2e73',
'Europe': '#562D84',
'South and Latin America': '#4591CE',
'Africa': '#CE8529',
'Russia': '#49BFB5',
'Australia and New Zealand': '#0E9487'
}
# As the chart contains a huge amount of data, we increase the size of the chart and decrease font size (though it will be cluttered still – there is certainly value in postprocessing by a graphic designer!).
plt.rcParams['figure.figsize'] = 12, 6
plt.rcParams['font.size'] = 8
# +
fig, ax = plt.subplots()
# normalization of bubble sizes
norm = plt.Normalize(df['pop'].min(), df['pop'].max())
# iterate over continents and plot the data with the right colors
for continent, cdf in df.groupby('continent'):
sns.scatterplot('ghg_per_gdp', 'gdp_per_capita', data=cdf, color=color_schema[continent], label=continent, size='pop',
sizes=(10, 800), size_norm=norm, legend=False, ec=None)
# country labels
for i, row in cdf.iterrows():
plt.text(row['ghg_per_gdp'], row['gdp_per_capita'] + 100 + np.sqrt(row['pop']) / 15, row['region'],
color=color_schema[continent], ha='center', va='bottom')
# set the plot limits
xmax = df['ghg_per_gdp'].max() * 1.1
ymax = df['gdp_per_capita'].max() * 1.1
ax.set(xlim=(0, xmax), ylim=(0, ymax))
# add hyperbolas
xs = np.linspace(10, xmax, 200)
for i in list(range(2, 41, 2)):
ys = 1e6 * i / xs
sns.lineplot(xs, ys, lw=0.6, alpha=0.15, color='black')
plt.text(1e6 * i / 6.25e4, 6.25e4, str(i), color='black', ha='center', va='center')
# plot the world averages
ax.axvline(avg_ghg_per_gdp, color='black', alpha=0.4, linestyle=':')
ax.axhline(avg_gdp_per_capita, color='black', alpha=0.4, linestyle=':')
# add titles and descriptions
plt.text(1e6 / 6e4 - 3, 6.45e4, 'Levels of emissions per capita (t CO2 eq)', color='black', ha='left', va='center')
ax.set(xlabel='Emissions per GDP (g CO2eq / $)', ylabel='GDP per capita ($)',
title='Emissions per GDP and per capita')
plt.show()
# -
# Matplotlib also allows to save the chart as PDF or SVG. These vector formats can then be loaded into a specialized graphic software such as Illustrator or Inkscape to turn them into a nice, proper visualization that can be found on our website.
| notebooks/emission-intensity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from __future__ import print_function
import os
import pandas as pd
import numpy as np
# %matplotlib inline
from matplotlib import pyplot as plt
from pandas.tools.plotting import autocorrelation_plot
from statsmodels.graphics.tsaplots import plot_acf
#Read data from Excel file
daily_temp = pd.read_excel('datasets/mean-daily-temperature-fisher-river.xlsx')
#Display first 20 rows of the DataFrame
daily_temp.head(10)
#Make formatted date as the row index of the dataset and drop the Date column
daily_temp.index = daily_temp['Date'].map(lambda date: pd.to_datetime(date, '%Y-%m-%d'))
daily_temp.drop('Date', axis=1, inplace=True)
#Re-display the first 10 rows of the modified DataFrame
daily_temp.head(10)
fig = plt.figure(figsize=(5.5, 5.5))
ax = fig.add_subplot(1,1,1)
ax.set_title('Daily temperatures of Fisher River, TX, US')
daily_temp.plot(ax=ax)
plt.savefig('plots/ch2/B07887_02_08.png', format='png', dpi=300)
#Calculate monthly mean temperature
montly_resample = daily_temp['Mean_Temperature'].resample('M')
monthly_mean_temp = montly_resample.mean()
print('Shape of monthly mean temperature dataset:', monthly_mean_temp.shape)
monthly_mean_temp.head(10)
#Plot the monthly mean temparature
fig = plt.figure(figsize=(5.5, 5.5))
ax = fig.add_subplot(1,1,1)
ax.set_title('Monthly mean temperatures of Fisher River, TX, US')
monthly_mean_temp.plot(ax=ax)
plt.savefig('plots/ch2/B07887_02_09.png', format='png', dpi=300)
#Plot ACF of the monthly mean temparature using pandas.tools.plotting.autocorrelation_plot
fig = plt.figure(figsize=(5.5, 5.5))
ax = fig.add_subplot(1,1,1)
ax.set_title('ACF of monthly mean temperatures of Fisher River, TX, US')
#autocorrelation_plot(monthly_mean_temp, ax=ax)
plot_acf(monthly_mean_temp,lags=30,
title='ACF of monthly mean temperatures of Fisher River, TX, US',
ax=ax
)
#plt.savefig('plots/ch2/B07887_02_10.png', format='png', dpi=300)
plot_acf(monthly_mean_temp,lags=20, title='ACF of monthly mean temperatures of Fisher River, TX, US')
plt.acorr(monthly_mean_temp, maxlags=20)
#Take seasonal differences with a period of 12 months on monthly mean temperatures
seasonal_diff = monthly_mean_temp.diff(12)
seasonal_diff = seasonal_diff[12:]
#Plot the seasonal differences
fig = plt.figure(figsize=(5.5, 5.5))
ax = fig.add_subplot(1,1,1)
ax.set_title('Seasonal differences')
seasonal_diff.plot(ax=ax)
plt.savefig('plots/ch2/B07887_02_11.png', format='png', dpi=300)
#Plot the seasonal differences
fig = plt.figure(figsize=(5.5, 5.5))
ax = fig.add_subplot(1,1,1)
ax.set_title('ACF of Seasonal differences')
autocorrelation_plot(seasonal_diff, ax=ax, )
plt.savefig('plots/ch2/B07887_02_12.png', format='png', dpi=300)
#Perform Ljung-Box test on monthly mean temperature to get the p-values
#We will use lags of upto 10
_, _, _, pval_monthly_mean = stattools.acf(monthly_mean_temp, unbiased=True,
nlags=10, qstat=True, alpha=0.05)
print('Null hypothesis is rejected for lags:', np.where(pval_monthly_mean<=0.05))
#Perform Ljung-Box test on monthly mean temperature to get the p-values
#We will use lags of upto 10
_, _, _, pval_seasonal_diff = stattools.acf(seasonal_diff, unbiased=True,
nlags=10, qstat=True, alpha=0.05)
print('Null hypothesis is rejected for lags:', np.where(pval_seasonal_diff<=0.05))
acf , confint, qstat , pval_monthly_mean = stattools.acf(monthly_mean_temp, unbiased=True,
nlags=10, qstat=True, alpha=0.05)
confint
| time series regression/autocorelation, mov avg etc/Seasonal_Differencing-AutoCorr_Plot_Correction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: py35-paddle1.2.0
# ---
# + jupyter={"outputs_hidden": false}
#解压数据
# !unzip -d data/data117383/train_img/ -q data/data117383/GAMMA.zip
# + jupyter={"outputs_hidden": false}
# # !python train.py
# !python predect.py --model_id=186 --gpu=True
# + jupyter={"outputs_hidden": false}
# 如果需要进行持久化安装, 需要使用持久化路径, 如下方代码示例:
# If a persistence installation is required,
# you need to use the persistence path as the following:
# !mkdir /home/aistudio/external-libraries
# !pip install beautifulsoup4 -t /home/aistudio/external-libraries
# + jupyter={"outputs_hidden": false}
# 同时添加如下代码, 这样每次环境(kernel)启动的时候只要运行下方代码即可:
# Also add the following code,
# so that every time the environment (kernel) starts,
# just run the following code:
import sys
sys.path.append('/home/aistudio/external-libraries')
# -
# 请点击[此处](https://ai.baidu.com/docs#/AIStudio_Project_Notebook/a38e5576)查看本环境基本用法. <br>
# Please click [here ](https://ai.baidu.com/docs#/AIStudio_Project_Notebook/a38e5576) for more detailed instructions.
| main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.4 64-bit (conda)
# name: python394jvsc74a57bd0613fe122287fdb1a4092b1ec324ab5e18de9ec977608057b646301948d1df577
# ---
# # Neural Network for Fashion MNIST dataset
import numpy as np
from numpy.random import default_rng
import matplotlib.pyplot as plt
from sklearn.neural_network import MLPClassifier
from sklearn.model_selection import train_test_split, GridSearchCV, cross_val_score
from sklearn.metrics import f1_score
from sklearn.metrics import plot_confusion_matrix
# read csv and save output as numpy array
df_train = np.loadtxt('fashion-mnist_train.csv', delimiter=',', skiprows=1)
df_val = np.loadtxt('fashion-mnist_test.csv', delimiter=',', skiprows=1)
# Split features and target arrays
X = df_train[:, 1:]
y = df_train[:, 0]
# random permutation of indexes
# use 1st 10000 rows for training and evaluation
rng = np.random.default_rng(seed=42)
r = rng.choice(len(df_train), len(df_train), replace=False)
X_train, y_train = X[r[:10000], :], y[r[:10000]]
# scale the train data in ranage [0,1] for better performance
X_train = X_train/255
# verify that dataset is properly balanced
from collections import Counter
sorted(Counter(y_train).items())
# ### Plot 16 random samples from the training set with the corresponding labels.
# +
# show 1st 16 pictures
# https://scikit-learn.org/stable/auto_examples/ensemble/plot_voting_decision_regions.html
fig, axs = plt.subplots(4, 4, sharex='col', sharey='row', figsize=(8, 8))
for i, ax in enumerate(axs.flat):
image = X_train[i].reshape(28,28)
ax.imshow(image, cmap=plt.cm.binary)
ax.set_title(y_train[i])
# -
# ### Task 2. Train a multilayer perceptron with hyperparameters tweaking
# Define baseline model
clf = MLPClassifier(random_state=1, early_stopping=True, validation_fraction=0.2)
clf.fit(X_train, y_train.ravel())
# Scoring
clf_score = cross_val_score(clf, X_train, y_train, cv=3, scoring='f1_micro')
print(f'Model = baseline MLPClassifier, f1_score={round(clf_score.mean(), 6)}, deviation={round(clf_score.std(), 6)}')
# Baseline model does provide good f1 score of 85% <br> Let us try to improve it by some parameter tuning
# We start with hidden layers tuning
# + tags=[]
parameters = {'hidden_layer_sizes': [[256, 256, 128], [256, 128, 128], [256, 128, 128, 128]]
} # use simple network with 3 hidden layers
clf = MLPClassifier(random_state=1, early_stopping=True, validation_fraction=0.2)
clf_grid = GridSearchCV(clf, parameters, verbose=3, cv=3, scoring='f1_micro')
clf_grid.fit(X_train, y_train.ravel())
print(f'The most optimal value for {clf_grid.best_params_} gives an F1 score of {round(clf_grid.best_score_, 5)}')
# -
# We got an improved score comparing to a baseline model using different set on neurons in each layer. By default, MLClassifier uses 3 layers with 100 neurons in each. I have tested 3 alternatives and by using 3 layers in 256, 128 and 128 neurons respectively we were able to slightly increase the f1 score
# Let us try further model score increase by tuning "batch size" and "learning rate" parameters using the tuned hidden layer parameter
# +
parameters = {'batch_size': [200, 600, 1000],
'learning_rate_init': [0.001, 0.01, 0.1]
} # use simple network with 3 hidden layers
clf = MLPClassifier(random_state=1, early_stopping=True, validation_fraction=0.2, hidden_layer_sizes=[256, 128, 128])
clf_grid = GridSearchCV(clf, parameters, verbose=3, cv=3, scoring='f1_micro')
clf_grid.fit(X_train, y_train.ravel())
print(f'The most optimal value for {clf_grid.best_params_} gives an F1 score of {round(clf_grid.best_score_, 5)}')
# -
# Seems like that by using default parameters we get the highest score from the parameters set, so we keep the default ones.
# We will try to optimize the model by tuning regularization parameter "alpha" and different solvers for weight optimization
# +
parameters = {'alpha': [0.0001, 0.001, 0.01],
'solver': ['sgd', 'adam']
} # use simple network with 3 hidden layers
clf = MLPClassifier(random_state=1, early_stopping=True, validation_fraction=0.2, hidden_layer_sizes=[256, 128, 128])
clf_grid = GridSearchCV(clf, parameters, verbose=3, cv=3, scoring='f1_micro')
clf_grid.fit(X_train, y_train.ravel())
print(f'The most optimal value for {clf_grid.best_params_} gives an F1 score of {round(clf_grid.best_score_, 5)}')
# -
# Again, default parameters show the highest score, so no additional updates done here
# Finally, we will test different activation funcitons
# +
parameters = {'activation': ['logistic', 'tanh', 'relu'] # default is "relu"
}
clf = MLPClassifier(random_state=1, early_stopping=True, validation_fraction=0.2, hidden_layer_sizes=[256, 128, 128])
clf_grid = GridSearchCV(clf, parameters, verbose=3, cv=3, scoring='f1_micro')
clf_grid.fit(X_train, y_train.ravel())
print(f'The most optimal value for {clf_grid.best_params_} gives an F1 score of {round(clf_grid.best_score_, 5)}')
# -
# We will keep the default activation function ("relu")
# Let us calculate the score for using the complete dataset fro training and validation
# Split validation dataset
X_val = df_val[:, 1:]
y_val = df_val[:, 0]
clf = MLPClassifier(random_state=1, early_stopping=True, validation_fraction=0.2, hidden_layer_sizes=[256, 128, 128])
clf.fit(X, y)
# +
# Predictions
train_predict = clf.predict(X)
test_predict = clf.predict(X_val)
# Evaluate the model
print('Model performance on training set:')
score = f1_score(train_predict, y, average='micro')
print(f'Training F1 score: {score}')
print('Model performance on testing set:')
score = f1_score(test_predict, y_val, average='micro')
print(f'Test F1 score: {score}')
# -
# Quite good f1 score achieved with over 92% on training dataset and 89% on a validation dataset
# ### Confusion matrix
# +
# https://scikit-learn.org/stable/auto_examples/model_selection/plot_confusion_matrix.html#sphx-glr-auto-examples-model-selection-plot-confusion-matrix-py
# +
fig, ax = plt.subplots(figsize=(8, 8))
disp = plot_confusion_matrix(clf, X_val, y_val, cmap=plt.cm.Blues, ax=ax, normalize='true')
disp.ax_.set_title('Confusion matrix')
# -
# Most categories are well predicted, however we do have particluaryt one category with low prediciton score. Let us evaluate it further
# +
# Define lables for categories https://www.kaggle.com/zalando-research/fashionmnist:
labels = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat', 'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
# re-cfreate confusion matrix
fig, ax = plt.subplots(figsize=(8, 8))
disp = plot_confusion_matrix(clf, X_val, y_val, cmap=plt.cm.Blues, ax=ax, normalize='true', display_labels=labels, xticks_rotation='vertical')
disp.ax_.set_title('Confusion matrix')
# -
# Plot shows that category "shirt" is often mixed with categories "T-shirt/top", "Pullover" or "Coat". tThis makes sense as all the categories represent similar types of garments <br>
#
# From another sie categories as "Ancle boot" and "Trouser" differ quite much from any other categories and therefore deliver very good predicton score.
| Jupyter_Notebook/Supervised_Learning/mlp_fashion_mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#bring the dependencies
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from scipy.interpolate import Rbf, InterpolatedUnivariateSpline
# %matplotlib inline
#import a spreadsheet file containing sieving data of a coarse sand and gravel sample (one phi interval)
df = pd.read_csv(r'C:\pathtoyourfile\filename.csv')
df.head(10) #This is just to show an example
# +
#Make a cumulative freq. graph of grain size using phi values for the x axis
fig,ax = plt.subplots(figsize=(8, 6))
x=df['phi'].values
y=df['cumul_wt_percent'].values
# Hide the right and top lines of the default box
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
#PLot the dots and line and assign ticks and labels
plt.plot(x, y, marker='o')
lines = plt.gca().lines[0].get_xydata() #Create an array of the points along the line
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.xlabel(r'$\phi$', fontsize=16)
plt.ylabel('Cumul. Freq.\n$\\regular_{wt. percent}$', fontsize=18) #r'$\alpha > \beta$'
#Add the grid lines and show the plot
ax.grid(True, linestyle='--')
plt.show()
#fig.savefig('cumFq.svg', bbox_inches = 'tight', format='svg')
print(lines)
# +
#Make a cumulative freq. graph of grain size using size categories for the x axis
fig, ax = plt.subplots(figsize=(12, 10))
ax.set_yticks( np.linspace(0, 100, 11 ) )
x=df['phi'].values
y=df['cumul_wt_percent'].values
# Hide the right and top lines of the default box
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
#PLot the dots and line and assign ticks and labels
plt.plot(x, y, marker='o')
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.xlabel('Size class', fontsize=18)
plt.ylabel('Cumul. Freq.\n$\\regular_{wt. percent}$', fontsize=18) #r'$\alpha > \beta$'
xticks = [ -3, -2, -1, 0, 1, 2, 3, 4, 5 ]
xlbls = ['large pebble \n($\phi$= -3)', 'pebble \n($\phi$= -2)', 'granule \n($\phi$= -1)', 'v. coarse sand \n($\phi$= 0)', 'coarse sand \n($\phi$= 1)', 'medium sand \n($\phi$= 2)', 'fine sand \n($\phi$= 3)', 'v fine sand \n($\phi$= 4)', 'silt and clay \n($\phi$= 5)']
ax.set_xticks( xticks )
ax.set_xticklabels( xlbls )
ax.set_xlim(-3, 5.5)
# vertical alignment of xtick labels
va = [ 0, -.06, 0, -.06, 0, -.06, 0, -.06 ]
for t, y in zip( ax.get_xticklabels( ), va ):
t.set_y( y )
#Add the grid lines and show the plot
ax.grid(True, linestyle='--')
plt.show()
#fig.savefig('cumFq2.svg', bbox_inches = 'tight', format='svg')
# +
#Same graph but this time with a RBF-linear combo interpolation
x=df['phi'].values
y=df['cumul_wt_percent'].values
xi = np.linspace(-2.0, 3.0, 40)
xj=np.linspace(3.0, 5.0, 10) #That one is for the short linear line at the right extremity
xk=np.linspace(-3.0, -2.0, 10) #That one is for the short linear line at the left extremity
fig = plt.figure(figsize=(12.5,8))
ax = fig.add_subplot(1,1,1) # row-col-num
ax.set_xlim((-3.0, 5.5,))
# Hide the right and top lines of the default box
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
#PLot the dots and line and assign ticks and labels
plt.plot(x, y, 'bo')
plt.xlabel('Size class', fontsize=18)
plt.ylabel('Cumul. Freq.\n$\\regular_{wt. percent}$', fontsize=18) #r'$\alpha > \beta$'
# use RBF method
rbf = Rbf(x, y, function='thin_plate')
fi = rbf(xi)
rbf=Rbf(x, y, function='linear')
fj=rbf(xj)
rbf=Rbf(x, y, function='linear')
fk=rbf(xk)
plt.plot(xi, fi, 'g')
plt.plot(xj, fj, 'g')
plt.plot(xk, fk, 'g')
#location of the ticks and their labels
xticks = [ -3.5, -2.5, -1.5, -0.5, 0.5, 1.5, 2.5, 3.5, 4.5 ]
xticks_minor = [ -3, -2, -1, 0, 1, 2, 3, 4 ]
xlbls = ['large pebble \n($\phi$= <-3)', 'pebble \n($\phi$= -2 to -3)', 'granule \n($\phi$= -1 to -2)',
'v. coarse sand \n($\phi$= 0 to -1)', 'coarse sand \n($\phi$= 0 to 1)', 'medium sand \n($\phi$= 1 to 2)', 'fine sand \n($\phi$= 2 to 3)', 'v fine sand \n($\phi$= 3 to 4)', 'silt and clay \n($\phi$= >4)']
ax.set_xticks( xticks )
ax.set_xticks( xticks_minor, minor=True )
ax.set_xticklabels( xlbls, fontsize=12 )
ax.tick_params( axis='x', which='minor', direction='out', length=30 )
#ax.tick_params( axis='x', which='major', bottom='off', top='off' ) optional to show ticks at the top
# vertical alignment of xtick labels
va = [ 0, -.07, 0, -.07, 0, -.07, 0, -.07 ]
for t, y in zip( ax.get_xticklabels( ), va ):
t.set_y( y )
#Add the grid lines
ax.grid(True, which='minor', axis='x', linestyle='--')
ax.grid(True, which='major', axis='y', linestyle='--')
#fig.savefig('cumFq2.svg', bbox_inches = 'tight', format='svg')
# +
#to get the default black frame
plt.style.use('default')
fig,ax = plt.subplots(figsize=(8,6))
# Hide the right and top spines
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
x_mid=df['phi_mid'].values
#Make the bar plot
plt.bar(x_mid, df['wt_percent'], color=('b'), alpha=0.5, edgecolor='black', linewidth=1.2, width=1.0) #alpha is for transparency
plt.xticks(x_mid, fontsize=14)
plt.yticks(fontsize=14)
plt.xlabel(r'$\phi$', fontsize=16)
plt.ylabel('Freq. (wt. %)', fontsize=16)
#Add the points
plt.scatter(x_mid, df['wt_percent'], c='red', marker='o')
fig.savefig('Hist_new.svg', bbox_inches = 'tight', format='svg')
| Particle_Size_Sand_Gravel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# What is Hypothesis Testing?
#
# A statistical hypothesis is an assumption about a population parameter. This assumption may or may not be true. Hypothesis testing refers to the formal procedures used by statisticians to accept or reject statistical hypotheses.
#
# Statistical Hypotheses
# The best way to determine whether a statistical hypothesis is true would be to examine the entire population. Since that is often impractical, researchers typically examine a random sample from the population. If sample data are not consistent with the statistical hypothesis, the hypothesis is rejected.
#
# There are two types of statistical hypotheses.
#
# Null hypothesis. The null hypothesis, denoted by H0, is usually the hypothesis that sample observations result purely from chance.
#
# Alternative hypothesis. The alternative hypothesis, denoted by H1 or Ha, is the hypothesis that sample observations are influenced by some non-random cause.
# For example, suppose we wanted to determine whether a coin was fair and balanced. A null hypothesis might be that half the flips would result in Heads and half, in Tails. The alternative hypothesis might be that the number of Heads and Tails would be very different. Symbolically, these hypotheses would be expressed as
#
# H0: P = 0.5
# Ha: P ≠ 0.5
#
# Suppose we flipped the coin 50 times, resulting in 40 Heads and 10 Tails. Given this result, we would be inclined to reject the null hypothesis. We would conclude, based on the evidence, that the coin was probably not fair and balanced.
#
# Can We Accept the Null Hypothesis?
# Some researchers say that a hypothesis test can have one of two outcomes: you accept the null hypothesis or you reject the null hypothesis. Many statisticians, however, take issue with the notion of "accepting the null hypothesis." Instead, they say: you reject the null hypothesis or you fail to reject the null hypothesis.
#
# Why the distinction between "acceptance" and "failure to reject?" Acceptance implies that the null hypothesis is true. Failure to reject implies that the data are not sufficiently persuasive for us to prefer the alternative hypothesis over the null hypothesis.
#
#
# Hypothesis Tests
# Statisticians follow a formal process to determine whether to reject a null hypothesis, based on sample data. This process, called hypothesis testing, consists of four steps.
#
# State the hypotheses. This involves stating the null and alternative hypotheses. The hypotheses are stated in such a way that they are mutually exclusive. That is, if one is true, the other must be false.
#
# Formulate an analysis plan. The analysis plan describes how to use sample data to evaluate the null hypothesis. The evaluation often focuses around a single test statistic.
#
# Analyze sample data. Find the value of the test statistic (mean score, proportion, t statistic, z-score, etc.) described in the analysis plan.
#
# Interpret results. Apply the decision rule described in the analysis plan. If the value of the test statistic is unlikely, based on the null hypothesis, reject the null hypothesis.
#
# Decision Errors
# Two types of errors can result from a hypothesis test.
#
# Type I error. A Type I error occurs when the researcher rejects a null hypothesis when it is true. The probability of committing a Type I error is called the significance level. This probability is also called alpha, and is often denoted by α.
#
# Type II error. A Type II error occurs when the researcher fails to reject a null hypothesis that is false. The probability of committing a Type II error is called Beta, and is often denoted by β. The probability of not committing a Type II error is called the Power of the test.
# Decision Rules
# The analysis plan includes decision rules for rejecting the null hypothesis. In practice, statisticians describe these decision rules in two ways - with reference to a P-value or with reference to a region of acceptance.
#
# P-value. The strength of evidence in support of a null hypothesis is measured by the P-value. Suppose the test statistic is equal to S. The P-value is the probability of observing a test statistic as extreme as S, assuming the null hypotheis is true. If the P-value is less than the significance level, we reject the null hypothesis.
#
# Region of acceptance. The region of acceptance is a range of values. If the test statistic falls within the region of acceptance, the null hypothesis is not rejected. The region of acceptance is defined so that the chance of making a Type I error is equal to the significance level.
#
# The set of values outside the region of acceptance is called the region of rejection. If the test statistic falls within the region of rejection, the null hypothesis is rejected. In such cases, we say that the hypothesis has been rejected at the α level of significance.
#
# These approaches are equivalent. Some statistics texts use the P-value approach; others use the region of acceptance approach. In subsequent lessons, this tutorial will present examples that illustrate each approach.
#
# One-Tailed and Two-Tailed Tests
# A test of a statistical hypothesis, where the region of rejection is on only one side of the sampling distribution, is called a one-tailed test. For example, suppose the null hypothesis states that the mean is less than or equal to 10. The alternative hypothesis would be that the mean is greater than 10. The region of rejection would consist of a range of numbers located on the right side of sampling distribution; that is, a set of numbers greater than 10.
#
# A test of a statistical hypothesis, where the region of rejection is on both sides of the sampling distribution, is called a two-tailed test. For example, suppose the null hypothesis states that the mean is equal to 10. The alternative hypothesis would be that the mean is less than 10 or greater than 10. The region of rejection would consist of a range of numbers located on both sides of sampling distribution; that is, the region of rejection would consist partly of numbers that were less than 10 and partly of numbers that were greater than 10.
| stats/hypothesis_testing/hypothesis_testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''py38'': conda)'
# name: python3
# ---
import numpy as np
import torch
from tqdm import tqdm
from skimage.metrics import structural_similarity as ssim
from torch.autograd import Variable
from matplotlib.pyplot import imshow
import matplotlib.pyplot as plt
from torchvision.datasets import EMNIST
from time import time
import sklearn.preprocessing
import numpy as np
import robust_onlinehd
from GenAttack import GenAttack
scaler = sklearn.preprocessing.Normalizer()
torch.manual_seed(54)
# +
# loads simple mnist dataset
def load():
temp = EMNIST('./data/EMNIST', split = 'letters', train = True, download = True)
x = temp.data.unsqueeze(3).numpy().transpose((0,2,1,3))
y = temp.targets.numpy() - 1
temp = EMNIST('./data/EMNIST', split = 'letters', train = False, download = True)
x_test = temp.data.unsqueeze(3).numpy().transpose((0,2,1,3))
y_test = temp.targets.numpy() - 1
# changes data to pytorch's tensors
x = torch.from_numpy(x).float()
y = torch.from_numpy(y).long().squeeze()
x_test = torch.from_numpy(x_test).float()
y_test = torch.from_numpy(y_test).long().squeeze()
if len(x.shape) == 3:
x = x.unsqueeze(3)
x_test = x_test.unsqueeze(3)
return x, x_test, y, y_test
print('Loading...')
x, x_test, y, y_test = load()
# -
#criterias = [(0, 100, 0), (100, 150, 125), (150, 200, 175), (200, 256, 255)]
#criterias = [(0, 50, 0), (50, 100, 75), (100, 125, 124), (125, 150, 149), (150, 175, 174), (175, 200, 199), (200, 225, 224), (225, 256, 255)]
#criterias = []
kernel_size = 3
#kernel_size = 1
classes = y.unique().size(0)
features = x.size(1) * x.size(2)
model = robust_onlinehd.OnlineHD(kernel_size, scaler, classes, features, dim = 10000)
# +
model.set_criterias(x, 8)
if torch.cuda.is_available():
#x = x.cuda()
#y = y.cuda()
#x_test = x_test.cuda()
#y_test = y_test.cuda()
model = model.to('cuda')
print('Using GPU!')
print('Training...')
t = time()
model = model.fit(x, y, bootstrap=.3, lr=0.095, epochs=300, batch_size=8196)
t = time() - t
print('Validating...')
yhat = model(x).cpu()
yhat_test = model(x_test).cpu()
acc = (y == yhat).float().mean()
acc_test = (y_test == yhat_test).float().mean()
print(f'{acc = :6f}')
print(f'{acc_test = :6f}')
print(f'{t = :6f}')
# -
preds = model(x_test).cpu().numpy()
#preds = model(x).cpu().numpy()
targets = torch.randint(0, 10, preds.shape)
for i in tqdm(range(len(preds))):
while targets[i] == preds[i]:
targets[i] = torch.randint(0,10, (1,)).item()
unif = torch.ones(targets.shape[0])
while True:
indices = unif.multinomial(100)
for idx in indices:
if targets[idx] == y_test[idx]:
break
if idx == indices[-1] and targets[idx] != y_test[idx]:
break
else:
indices = unif.multinomial(100)
attacker = GenAttack(model, classes, 28 * 28, scaler, 0.6, 'cuda')
N = 8 # size of population to evolve
G = 5000 # number of generations to evolve through
p = torch.FloatTensor([0.9]) # the parameter for Bernoulli distribution used in mutation
alpha = torch.FloatTensor([1.0]) # the parameter controlling mutation amount (step-size in the original paper)
delta = torch.FloatTensor([0.9]) # the parametr controlling mutation amount (norm threshold in the original paper)
pops = []
results = []
# +
t = time()
for i in tqdm(indices):
temp = attacker.attack(x_test[i], targets[i], delta, alpha, p, N, G)
pops.append(temp[0].numpy())
results.append(temp[1])
t = time() - t
print(f'{t = :6f}')
# -
pops = np.array(pops)
sample_preds = preds[indices]
new_preds = []
for i in range(100):
new_preds.append(model(torch.tensor(pops[i])).cpu().numpy())
success = 0
success_idx = []
for i in range(100):
if targets[indices[i]].item() in new_preds[i]:
success_idx.append((indices[i].item(), (i, np.where(new_preds[i] == targets[indices[i]].item())[0][0])))
success += 1
print(success)
cache = {
'indices' : indices,
'sample_preds' : sample_preds,
'pops' : np.array(pops),
'hyper_parameter' : [N, G, p, alpha, delta],
'success_idx' : success_idx,
'model' : model,
'scaler' : model.scaler,
'targets' : targets,
'results' : results
}
torch.save(cache, 'robust_onlinehd_emnist.pt')
model(torch.tensor(pops[0]))
labels = {
0 : 'a',
1 : 'b',
2 : 'c',
3 : 'd',
4 : 'e',
5 : 'f',
6 : 'g',
7 : 'h',
8 : 'i',
9 : 'j',
10 : 'k',
11 : 'l',
12 : 'm',
13 : 'n',
14 : 'o',
15 : 'p',
16 : 'q',
17 : 'r',
18 : 's',
19 : 't',
20 : 'u',
21 : 'v',
22 : 'w',
23 : 'x',
24 : 'y',
25 : 'z'
}
# +
origin_idx, (new_idx, new_idx_idx) = success_idx[torch.randint(0, len(success_idx), (1,)).item()]
f, axes = plt.subplots(1, 2)
axes[0].imshow(x_test[origin_idx], cmap=plt.gray())
_ = axes[0].set_title('Properly classified : %s' % labels[sample_preds[new_idx].item()])
axes[1].imshow(pops[new_idx][new_idx_idx].astype(np.int32))
_ = axes[1].set_title('Misclassified : %s' % labels[new_preds[new_idx][new_idx_idx]])
# -
# temp = torch.load('robust_onlinehd_emnist.pt')
# model = temp['model']
# pops = temp['pops']
# targets = temp['targets']
# indices = temp['indices']
| test_emnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # RQ3: Reskilling intensity
# <NAME> | 15.06.2021
#
# ## Core Analysis Goal(s)
# 1. How close are occupations likely less demanded in a green transition to those being more demanded? (~ reskilling intensity)
# 2. Further disaggregate the analysis to the ISCO-08 1-digit level groups?
#
# ## Key Insight(s)
# 1. Ranking of average similarities is surprising. Only looking at occupation similarity, brown occupations are on average closer to green occupations than neutral occupations.
# - Neutral & Green
# - Brown & Green
# - Brown & Neutral
# 2. Cross-pair differences are statistically significant at p < 0.001
# 3. Between-pair differences not significant at p < 0.001 are neutral-green and green-brown
# 4. Results are not robust to changing the similarity matrix (upeksha vs kanders).
# using Upeksha's matrix, green-brown is closest, followed by green-neutral and neutral-brown
# + pycharm={"name": "#%%\n"}
import os
import sys
import logging
from pathlib import Path
import numpy as np
import scipy as sp
import statsmodels.api as sm
from statsmodels.formula.api import ols
# %load_ext autoreload
# %autoreload 2
import matplotlib as mpl
import matplotlib.pyplot as plt
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import seaborn as sns
sns.set_context("paper")
sns.set(rc={'figure.figsize': (16, 9.)})
sns.set_style("ticks")
import pandas as pd
pd.set_option("display.max_rows", 120)
pd.set_option("display.max_columns", 120)
from tqdm import tqdm
import scipy
logging.basicConfig(level=logging.INFO, stream=sys.stdout)
# -
# Define directory structure
# + pycharm={"name": "#%%\n"}
# project directory
abspath = os.path.abspath('')
project_dir = str(Path(abspath).parents[0])
# sub-directories
data_raw = os.path.join(project_dir, "data", "raw")
data_interim = os.path.join(project_dir, "data", "interim")
data_processed = os.path.join(project_dir, "data", "processed")
figure_dir = os.path.join(project_dir, "reports", "figures")
# -
# ESCO data set components
# + pycharm={"name": "#%%\n"}
occ = pd.read_csv(os.path.join(data_raw, "esco", "v1.0.3", "occupations_en.csv"))
skill_groups = pd.read_csv(os.path.join(data_raw, "esco", "v1.0.3", "skillGroups_en.csv"))
skills = pd.read_csv(os.path.join(data_raw, "esco", "v1.0.3", "skills_en.csv"))
occ_skills_mapping = pd.read_csv(os.path.join(data_raw, "esco", "v1.0.3", "occupationSkillRelations.csv"))
# + pycharm={"name": "#%%\n"}
df_metadata = pd.read_csv(
os.path.join(data_interim, "ESCO_ONET_METADATA.csv"),
index_col=0
)
df_metadata.greenness_vona_2018_v2 = df_metadata.greenness_vona_2018_v2.fillna(0)
# convert job zone to categorical var
#df_metadata.job_zone = pd.Categorical(df_metadata.job_zone, ordered=True)
df_metadata.isco_level_1 = pd.Categorical(df_metadata.isco_level_1, ordered=False)
df_metadata.isco_level_2 = pd.Categorical(df_metadata.isco_level_2, ordered=False)
# -
# #### Remove military occupations from df_metadata
# 21 occupations in total, all coded as neutral
#
# ISCO codes (n=21)
# - 110: n=12
# - 210: n=4
# - 310: n=5
# + pycharm={"name": "#%%\n"}
cond = (df_metadata.isco_level_4 == 110) | (df_metadata.isco_level_4 == 210) | (df_metadata.isco_level_4 == 310)
cond.name = "military"
df_cond = cond.reset_index()
df_cond.to_csv(os.path.join(data_interim, "esco_military_occupations.csv"),)
# for re-indexing the sim matrices
non_military_indices = df_cond[~df_cond.military].index.values
df_metadata = df_metadata.loc[~cond]
df_metadata = df_metadata.reset_index(drop=True)
# + pycharm={"name": "#%%\n"}
df_metadata
# + pycharm={"name": "#%%\n"}
# nesta report
sim_kanders = np.load(
os.path.join(data_raw, "mcc_data", "processed", "sim_matrices", "OccupationSimilarity_Combined.npy")
)
np.fill_diagonal(sim_kanders, 0)
# + pycharm={"name": "#%%\n"}
xn, yn = np.meshgrid(non_military_indices, non_military_indices)
sim_kanders_nm = sim_kanders[xn, yn]
np.save(
file=os.path.join(data_raw, "mcc_data", "processed", "sim_matrices", "OccupationSimilarity_Combined_no_military.npy"),
arr=sim_kanders_nm
)
# + pycharm={"name": "#%%\n"}
# + pycharm={"name": "#%%\n"}
# brown occupations
df_brown = pd.read_csv(
os.path.join(data_interim, "occupations_brown_vona_esco.csv"),
index_col=0
)
# + pycharm={"name": "#%%\n"}
# add brown occupation classification
df_brown["is_brown"] = np.ones(df_brown.shape[0], dtype=bool)
# merge
df_metadata = pd.merge(
df_metadata,
df_brown[["concept_uri", "is_brown"]],
how="left",
on="concept_uri"
)
df_metadata.is_brown = df_metadata.is_brown.fillna(False)
df_metadata["is_green"] = df_metadata.greenness_vona_2018_v2 > 0
df_metadata["is_neutral"] = (df_metadata.is_green == False) & (df_metadata.is_brown == False)
# drop duplicates
df_metadata = df_metadata.drop_duplicates(subset=["concept_uri"])
# 8ung: there are 30 occupations that have been matched to both brown and green
# the neutral occupations are fine! have to make decision on where to map the
# ambiguous cases
query = (df_metadata.is_brown == True) & (df_metadata.is_green == True)
df_metadata.query("is_brown == True & is_green == True").to_csv(
os.path.join(data_interim, "ESCO_ONET_METADATA_gbn_ambiguous_cases.csv")
)
# --> define ambiguous cases as brown as a first solution
df_metadata.loc[query, "is_green"] = False
df_metadata = df_metadata.reset_index(drop=True)
# + pycharm={"name": "#%%\n"}
query = (df_metadata.is_brown == True) & (df_metadata.is_green == True)
df_metadata.loc[query]
# + pycharm={"name": "#%%\n"}
df_metadata.to_csv(
os.path.join(data_interim, "ESCO_ONET_METADATA_gbn.csv")
)
# + [markdown] pycharm={"name": "#%% md\n"}
# #### Number of green/brown/neutral occupations per ISCO 1-digit group
# + pycharm={"name": "#%%\n"}
isco_lvl1_mapping = {
1: "Managers",
2: "Professionals",
3: "Technicians and associate professionals",
4: "Clerical support workers",
5: "Service and sales workers",
6: "Skilled agricultural, forestry and fishery workers",
7: "Craft and related trades workers",
8: "Plant and machine operators and assemblers",
9: "Elementary occupations"
}
# + pycharm={"name": "#%%\n"}
# table for overleaf document
cols = ["preferred_label", "isco_level_1", "is_brown", "is_green", "is_neutral"]
df_sub = df_metadata[cols]
#df_sub["isco_level_1_txt"] = df_sub.isco_level_1.replace(isco_lvl1_mapping)
df_sub_counts = df_sub.groupby("isco_level_1").sum().reset_index()
df_sub_counts["isco_level_1_txt"] = [isco_lvl1_mapping[val] for val in df_sub_counts.isco_level_1.values]
count_sums = df_sub_counts[["is_brown", "is_green", "is_neutral"]].sum(axis=1)
col_order = ["isco_level_1", "isco_level_1_txt", "is_brown"]
count_sums
df_sub_counts["is_brown_pc"] = (df_sub_counts["is_brown"] / count_sums) * 100
df_sub_counts["is_green_pc"] = (df_sub_counts["is_green"] / count_sums) * 100
df_sub_counts["is_neutral_pc"] = (df_sub_counts["is_neutral"] / count_sums) * 100
# store
df_sub_counts.to_excel(
os.path.join(data_processed, "occupation_group_by_isco_lvl1.xlsx")
)
# -
# ## Iterate over all occupations and compute avg reskilling intensities
# + pycharm={"name": "#%%\n"}
ids_brown = df_metadata.loc[df_metadata.is_brown == True, :].index.values
ids_green = df_metadata.loc[df_metadata.is_green == True, :].index.values
ids_neutral = df_metadata.loc[df_metadata.is_neutral == True, :].index.values
len(ids_neutral) * len(ids_neutral)
# + [markdown] pycharm={"name": "#%% md\n"}
# #### Case 1: Brown - Green
# + pycharm={"name": "#%%\n"}
sim_matrix = sim_kanders_nm
# + pycharm={"name": "#%%\n"}
sim_brown_green = []
for b in tqdm(ids_brown):
for g in ids_green:
sim_brown_green.append(sim_matrix[b, g])
np.mean(sim_brown_green)
sim_green_brown = []
for b in tqdm(ids_brown):
for g in ids_green:
sim_green_brown.append(sim_matrix[g, b])
np.mean(sim_green_brown)
# + [markdown] pycharm={"name": "#%% md\n"}
# #### Case 2: Brown - Neutral
# + pycharm={"name": "#%%\n"}
sim_brown_neutral = []
for b in tqdm(ids_brown):
for n in ids_neutral:
sim_brown_neutral.append(sim_matrix[b, n])
np.mean(sim_brown_neutral)
sim_neutral_brown = []
for b in tqdm(ids_brown):
for n in ids_neutral:
sim_neutral_brown.append(sim_matrix[n, b])
np.mean(sim_neutral_brown)
# -
# #### Case 3: Neutral - Green
# + pycharm={"name": "#%%\n"}
sim_neutral_green = []
for n in tqdm(ids_neutral):
for g in ids_green:
sim_neutral_green.append(sim_matrix[n, g])
np.mean(sim_neutral_green)
sim_green_neutral = []
for n in tqdm(ids_neutral):
for g in ids_green:
sim_green_neutral.append(sim_matrix[g, n])
np.mean(sim_green_neutral)
# -
# #### Case 4: all occupations
# + pycharm={"name": "#%%\n"}
ids_all = df_metadata.index.values
sim_all = []
for i in tqdm(ids_all):
for j in ids_all:
sim_all.append(sim_matrix[i, j])
# + pycharm={"name": "#%%\n"}
np.mean(sim_all)
np.median(sim_all)
# -
# #### Combine in single df
# + pycharm={"name": "#%%\n"}
def pad(seq, target_length, padding=None):
length = len(seq)
seq.extend([padding] * (target_length - length))
return seq
# + pycharm={"name": "#%%\n"}
df = pd.DataFrame(index=range(len(sim_neutral_green)))
df["sim_green_brown"] = pad(sim_green_brown, len(sim_neutral_green), np.nan)
df["sim_brown_green"] = pad(sim_brown_green, len(sim_neutral_green), np.nan)
df["sim_brown_neutral"] = pad(sim_brown_neutral, len(sim_neutral_green), np.nan)
df["sim_neutral_brown"] = pad(sim_neutral_brown, len(sim_neutral_green), np.nan)
df["sim_neutral_green"] = pad(sim_neutral_green, len(sim_neutral_green), np.nan)
df["sim_green_neutral"] = pad(sim_green_neutral, len(sim_neutral_green), np.nan)
# + pycharm={"name": "#%%\n"}
df2 = pd.DataFrame(index=range(len(sim_all)))
df2["sim_all"] = pad(sim_all, len(sim_all), np.nan)
df2["sim_green_brown"] = pad(sim_green_brown, len(sim_all), np.nan)
df2["sim_brown_green"] = pad(sim_brown_green, len(sim_all), np.nan)
df2["sim_brown_neutral"] = pad(sim_brown_neutral, len(sim_all), np.nan)
df2["sim_neutral_brown"] = pad(sim_neutral_brown, len(sim_all), np.nan)
df2["sim_neutral_green"] = pad(sim_neutral_green, len(sim_all), np.nan)
df2["sim_green_neutral"] = pad(sim_green_neutral, len(sim_all), np.nan)
# + [markdown] pycharm={"name": "#%% md\n"}
# #### Visualisations
# + pycharm={"name": "#%%\n"}
df.median().sort_values(ascending=False)
# + pycharm={"name": "#%%\n"}
df2.median().sort_values(ascending=False)
# + pycharm={"name": "#%%\n"}
scipy.stats.shapiro(
df.sim_neutral_green.dropna().values
)
# + pycharm={"name": "#%%\n"}
scipy.stats.probplot(df2.sim_all.dropna().values, dist="norm", plot=plt)
# + pycharm={"name": "#%%\n"}
df.mean().sort_values(ascending=False)
# + pycharm={"name": "#%%\n"}
vline_colors = ["#4a1486", "#807dba", "#f16913", "#8c2d04", "#4292c6", "#084594"]
alpha = 0.5
ax = sns.histplot(
data=df,
stat="count",
palette=vline_colors,
alpha=alpha,
)
for val, color in zip(df.median().values, vline_colors):
ax.axvline(val, color=color)
# indicate overall median
ax.axvline(df2.sim_all.median(), color="black", linestyle="--", label="sim_all")
ax.set_xlim(0, 0.5)
ax.set_xlabel("Occupation similarity [-]")
sns.despine()
plt.tight_layout()
plt.savefig(
os.path.join(figure_dir, "RQ3", "avg_sims_histogram_v2_appendix.png"),
bbox_inches="tight",
dpi=300
)
# + [markdown] pycharm={"name": "#%% md\n"}
# vline_colors = ["grey", "#807dba", "#4a1486", "#f16913", "#8c2d04", "#4292c6", "#084594"]
# alpha = 0.5
#
# ax = sns.histplot(
# data=df2,
# stat="count",
# palette=vline_colors,
# alpha=alpha
# )
#
# for val, color in zip(df.median().values, vline_colors):
# ax.axvline(val, color=color, alpha=alpha)
#
# ax.set_xlim(0, 0.6)
# ax.set_xlabel("Occupation similarity [-]")
#
# sns.despine()
# plt.tight_layout()
#
# plt.savefig(
# os.path.join(figure_dir, "RQ3", "avg_sims_histogram_v3.png"),
# bbox_inches="tight",
# dpi=150
# )
# + pycharm={"name": "#%%\n"}
col_sel = ["sim_brown_green", "sim_brown_neutral", "sim_neutral_green"]
col_rename = {
"sim_brown_green":"Brown to Green",
"sim_brown_neutral": "Brown to Neutral",
"sim_neutral_green": "Neutral to Green"
}
# ["#807dba", "#4a1486", "#f16913", "#8c2d04", "#4292c6", "#084594"]
vline_colors = ["#807dba", "#f16913", "#4292c6"]
alpha = 0.7
ax = sns.histplot(
data=df[col_sel].rename(columns=col_rename),
stat="count",
palette=vline_colors,
alpha=alpha,
)
for val, color in zip(df[col_sel].median().values, vline_colors):
ax.axvline(val, color=color)
# indicate overall median
ax.axvline(df2.sim_all.median(), color="black", linestyle="--", label="sim_all")
ax.set_xlim(0, 0.5)
ax.set_xlabel("Occupation similarity [-]")
sns.despine()
plt.tight_layout()
plt.savefig(
os.path.join(figure_dir, "RQ3", "avg_sims_histogram_v2_main.png"),
bbox_inches="tight",
dpi=300
)
# + pycharm={"name": "#%%\n"}
# + [markdown] pycharm={"name": "#%% md\n"}
# #### Difference of central tendencies and post-hoc tests
# + pycharm={"name": "#%%\n"}
# ANOVA
scipy.stats.f_oneway(
df.sim_brown_green.dropna().values,
df.sim_brown_neutral.dropna().values,
)
# + pycharm={"name": "#%%\n"}
# Tukey-HSD
import statsmodels.stats.multicomp as mc
comp = mc.MultiComparison(
df.melt().dropna().value,
df.melt().dropna().variable
)
post_hoc_res = comp.tukeyhsd(alpha=0.001)
post_hoc_res.summary()
# + pycharm={"name": "#%%\n"}
# Kruskall-Wallis H-test
# Appendix
scipy.stats.kruskal(
df.sim_brown_green.dropna().values,
df.sim_green_brown.dropna().values,
df.sim_brown_neutral.dropna().values,
df.sim_neutral_brown.dropna().values,
df.sim_green_neutral.dropna().values,
df.sim_neutral_green.dropna().values,
)
# Main
scipy.stats.kruskal(
df.sim_brown_green.dropna().values,
df.sim_brown_neutral.dropna().values,
df.sim_neutral_green.dropna().values,
)
# + pycharm={"name": "#%%\n"}
import scikit_posthocs as sp
dfm = df.melt(var_name="groups", value_name="values").dropna(subset=["values"])
sp.posthoc_dunn(
dfm,
val_col="values",
group_col="groups",
p_adjust="bonferroni"
)
# + pycharm={"name": "#%%\n"}
# Appendix
df2m = df2.melt(var_name="groups", value_name="values").dropna(subset=["values"])
sp.posthoc_dunn(
df2m,
val_col="values",
group_col="groups",
p_adjust="bonferroni"
)
# + pycharm={"name": "#%%\n"}
delete = ["sim_green_brown", "sim_neutral_brown", "sim_green_neutral"]
df3m = df2.drop(columns=delete).melt(var_name="groups", value_name="values").dropna(subset=["values"])
# Main
sp.posthoc_dunn(
df3m,
val_col="values",
group_col="groups",
p_adjust="bonferroni"
)
# + [markdown] pycharm={"name": "#%% md\n"}
# #### Repeat analysis at the level of ISCO-08 1-digit groups (within-groups)
# - brown-green vs. brown-neutral (no reverse directions)
# - look at within-group similarities
# + pycharm={"name": "#%%\n"}
results = {}
for grp, df_isco1 in df_metadata.groupby("isco_level_1"):
#print(grp, df.shape)
# get ids of groups in sub-df
ids_brown = df_isco1.loc[df_isco1.is_brown == True, :].index.values
ids_green = df_isco1.loc[df_isco1.is_green == True, :].index.values
ids_neutral = df_isco1.loc[df_isco1.is_neutral == True, :].index.values
ids_all = df_isco1.index.values
# brown - green
sim_brown_green = []
for b in ids_brown:
for g in ids_green:
sim_brown_green.append(sim_matrix[b, g])
sim_brown_neutral = []
for b in ids_brown:
for n in ids_neutral:
sim_brown_neutral.append(sim_matrix[b, n])
# all
sim_all = []
for i in ids_all:
for j in ids_all:
sim_all.append(sim_matrix[i, j])
# Kruskall-Wallis H-test
H, p_bg_bn = scipy.stats.kruskal(sim_brown_green, sim_brown_neutral)
H_bg, p_bg_all = scipy.stats.kruskal(sim_brown_green, sim_all)
H_bn, p_bn_all = scipy.stats.kruskal(sim_brown_neutral, sim_all)
# store results
results[grp] = [
np.nanmedian(sim_brown_green),
np.nanmedian(sim_brown_neutral),
np.nanmedian(sim_all),
p_bg_bn,
p_bg_all,
p_bn_all
]
# + pycharm={"name": "#%%\n"}
results
# + pycharm={"name": "#%%\n"}
df_results = pd.DataFrame.from_dict(
results, orient="index", columns=[
"$M_{bg}$", "$M_{bn}$", "$M_{all}$",
"$p_{bg-bn}$", "$p_{bg-all}$", "$p_{bn-all}$"
]
)
df_results = df_results.reset_index().rename(columns={"index": "ISCO"})
df_results["Major group"] = [isco_lvl1_mapping[val] for val in df_results.ISCO.values]
col_order = [
'ISCO', 'Major group', "$M_{bg}$", "$M_{bn}$", "$M_{all}$",
"$p_{bg-bn}$", "$p_{bg-all}$", "$p_{bn-all}$"
]
df_results = df_results[col_order]
# store
df_results.to_csv(
os.path.join(data_processed, "gbn_median_sims_by_isco_lvl1_v2.csv"),
float_format='%.3f',
index=False
)
df_results
# -
# #### Repeat analysis at the level of ISCO-08 1-digit groups (cross-group)
# - brown-green vs. brown-neutral (no reverse directions)
# - look at cross-group similarities (heatmaps)
# + pycharm={"name": "#%%\n"}
df_metadata.loc[df_metadata.isco_level_1 != 1]
df_metadata.query("isco_level_1 != 1")
# + pycharm={"name": "#%%\n"}
results_cross_group = []
# loop over source group
for grp_source, df_source in df_metadata.groupby("isco_level_1"):
#df_metadata_sub = df_metadata.loc[df_metadata.isco_level_1 != grp]
# get ids of groups in source sub-df
ids_brown_source = df_source.loc[df_source.is_brown == True, :].index.values
ids_green_source = df_source.loc[df_source.is_green == True, :].index.values
ids_neutral_source = df_source.loc[df_source.is_neutral == True, :].index.values
for grp_target, df_target in df_metadata.groupby("isco_level_1"):
# get ids of groups in target sub-df
ids_brown_target = df_target.loc[df_target.is_brown == True, :].index.values
ids_green_target = df_target.loc[df_target.is_green == True, :].index.values
ids_neutral_target = df_target.loc[df_target.is_neutral == True, :].index.values
# brown - green
sim_brown_green = []
for b in ids_brown_source:
for g in ids_green_target:
sim_brown_green.append(sim_matrix[b, g])
# brown - neutral
sim_brown_neutral = []
for b in ids_brown_source:
for n in ids_neutral_target:
sim_brown_neutral.append(sim_matrix[b, n])
# store
results_cross_group.append(
[grp_source,
grp_target,
np.nanmedian(sim_brown_green),
np.nanmedian(sim_brown_neutral)
]
)
# + pycharm={"name": "#%%\n"}
mat_results_bg = np.zeros((9, 9))
mat_results_bn = np.zeros((9, 9))
for l in results_cross_group:
x = l[0]
y = l[1]
bg = l[2]
bn = l[3]
mat_results_bg[x-1, y-1] = bg
mat_results_bn[x-1, y-1] = bn
# + pycharm={"name": "#%%\n"}
high = "#08306b"
medium = "#2171b5"
low = "#6baed6"
isco_lvl1_skill_level_colors = {
0: high,
1: high,
2: high,
3: medium,
4: medium,
5: medium,
6: medium,
7: medium,
8: low
}
# + pycharm={"name": "#%%\n"}
fig, (ax1, ax2) = plt.subplots(ncols=2, sharex=True, sharey=True)
# Brown - Green
df_results_bg = pd.DataFrame(
mat_results_bg,
index=list(isco_lvl1_mapping.values()),
columns=list(isco_lvl1_mapping.values())
)
sns.heatmap(
data=df_results_bg,
cmap=plt.get_cmap("Blues", 14),
vmin=0,
vmax=0.35,
annot=True,
square=True,
cbar=False,
ax=ax1,
fmt=".2f"
)
ax1.set_title("Brown-Green")
#plt.tight_layout()
#plt.savefig(
# os.path.join(figure_dir, "RQ3", "avg_sims_heatmap_bg.png"),
# bbox_inches="tight",
# dpi=150
#)
# Brown - Neutral
df_results_bn = pd.DataFrame(
mat_results_bn,
index=list(isco_lvl1_mapping.values()),
columns=list(isco_lvl1_mapping.values())
)
sns.heatmap(
data=df_results_bn,
cmap=plt.get_cmap("Blues", 14),
vmin=0,
vmax=0.35,
annot=True,
square=True,
cbar=False,
ax=ax2,
fmt=".2f"
)
ax2.set_title("Brown-Neutral")
# color tick labels
for k, v in isco_lvl1_skill_level_colors.items():
ax1.get_xticklabels()[k].set_color(v)
ax1.get_yticklabels()[k].set_color(v)
ax2.get_xticklabels()[k].set_color(v)
plt.tight_layout()
plt.savefig(
os.path.join(figure_dir, "RQ3", "avg_sims_heatmap_combined.png"),
bbox_inches="tight",
dpi=150
)
# + pycharm={"name": "#%%\n"}
df_results_delta = df_results_bg - df_results_bn
ax = sns.heatmap(
data=df_results_delta,
cmap=plt.get_cmap("PiYG"),
center=0,
annot=True,
square=True,
cbar=False,
fmt=".2f"
)
# color tick labels
for k, v in isco_lvl1_skill_level_colors.items():
ax.get_xticklabels()[k].set_color(v)
ax.get_yticklabels()[k].set_color(v)
ax.set_title("Brown-Green minus Brown-Neutral")
plt.tight_layout()
plt.savefig(
os.path.join(figure_dir, "RQ3", "avg_sims_heatmap_bg_minus_bn.png"),
bbox_inches="tight",
dpi=150
)
# + pycharm={"name": "#%%\n"}
| notebooks/16-fz-reskilling-intensity-analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:whwtimeseries]
# language: python
# name: conda-env-whwtimeseries-py
# ---
# # Extract SWE data from SnowModel output and export to NetCDF
# 3/18-16/2020. Emilio
# ## From Nina
#
# Possible tracking parameters:
#
# 1. Total daily water volume by watershed
# - could be aggregated to monthly resolution for GRACE
# 2. Total daily change in water volume by watershed classified by:
# - Elevation
# - Aspect
# - Slope
# - Landcover
# 3. PDFs and CDFs of SWE change by watershed
# 4. Watershed hypsometry
# 5. Watershed SWE hypsometry
# 6. Percent snow covered area by watershed
# - Could compare to MODIS-derived daily snow covered area
# 7. Considering there may not be a ‘best’ calibration/assimilation run, with different parameters performing better at certain SNOTEL stations, we could select the top 3-5 combinations of parameters to create a SWE ensemble. Visual would be a time series of the uncertainty in daily SWE based off an ensemble of parameters or assimilation combinations
# ## MinIO
# - https://docs.min.io/docs/minio-quickstart-guide.html
# - https://docs.min.io/docs/minio-docker-quickstart-guide
# - https://blog.alexellis.io/meet-minio/
# - See the other links I entered into RedNotebook, 3/10/2020
# ### References for NetCDF CF/ACDD conventions
# - https://www.nodc.noaa.gov/data/formats/netcdf/v2.0/
# - https://www.nodc.noaa.gov/data/formats/netcdf/v2.0/grid.cdl
#
# ### Strategies, TO-DOs
# - **IMMEDIATE:** Something is wrong! SWE values are coming out as spanning both negative and positive values. But that doesn't make sense (negative depths?), and more importantly the values in Nina's plots are all positive. Could this be related to byte order (big/small endian)? **Also** need to confirm more directly that the map plots are correct and not transposed or wrong in other ways
# - https://en.wikipedia.org/wiki/Endianness. It looks like most common platforms these days are little endian
# - https://www.gamedev.net/forums/topic/688000-safe-to-say-all-are-in-little-endian-nowadays/5339764/
# - https://www.bignerdranch.com/blog/its-the-endian-of-the-world-as-we-know-it/
# - Plan to create a set of netcdf files segmented by time, say, one per year, with the file name encoding the time slice. eg, `swed_gdat_2014.nc`.
# - That series files can then be opened with `xarray.open_mfdataset`, which uses lazy loading (!).
# - Alternatively, later, it may be more practical to read the binary data in the same time segments, and write to zarr instead if such writing allows for concatenation along the time dimension
# - DONE. Add dimension variables (time, x, y), with corresponding attributes.
# - Add more global attributes, including ones matching with CF conventions.
# - Add more variable attributes, including CF standard name
# +
import os
from collections import OrderedDict
import numpy as np
import pandas as pd
import xarray as xr
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ## Parameters defining the XYT dimensions, file name, etc
# +
##USER INPUTS## - most of this can be read directly from the .ctl file or the .par file
#-> need to read in text file
# model filename
inFile = 'swed.gdat'
# start date
st = "2014-10-01"
# end date
ed = "2019-09-29"
# number of timesteps in model run
num_mod_timesteps = 365 * 5
# from .ctl file
nx = 1382 # number of cells in the x dimension?
ny = 2476 # number of cells in the y dimension?
xll = 487200
yll = 4690100
clsz = 100 # cellsize?
# -
#Build lat/lon array
lon = np.arange(xll, xll+nx*clsz, clsz)
lat = np.arange(yll, yll+ny*clsz, clsz)
len(lon), len(lat), len(lon)*len(lat)
# ## Check file size vs dimensions
filesize_bytes = os.path.getsize(inFile)
# file size in bytes and GB's (using bit shift)
filesize_bytes, filesize_bytes >> 30
# confirm that dimensions match file size
# 32 bit (4 byte) floating point data
4 * num_mod_timesteps * (len(lon)*len(lat)) == filesize_bytes
# ## Read only a small portion of the file, for testing
# I can't read it all at once b/c it's larger than my available laptop memory (23 GB vs 15.4 GB)
# **Notes about `np.fromfile`:**
#
# - Do not rely on the combination of `tofile` and `fromfile` for data storage, as the binary files generated are not platform independent. In particular, no byte-order or data-type information is saved. Data can be stored in the platform independent ``.npy`` format using `save` and `load` instead.
# - `count : int.` Number of items to read. ``-1`` means all items (i.e., the complete file).
nxy = len(lon)*len(lat)
nts = 10 # number of time steps to read
# +
# open grads model output file into a numpy array
with open(inFile, 'rb') as gradsfile:
numpy_data = np.fromfile(inFile, dtype='float32', count=nts*nxy)
numpy_data = np.reshape(numpy_data, (nts, ny, nx))
numpy_data.shape
# -
# NOTE: "lat" and "lon" actually look like Northing and Easting. Per Nina's notebook:
# ```python
# mod_proj = 'epsg:32612' # UTM zone 12N
# ```
# +
# Test byte swapping (changing endian order)
# numpy_data = numpy_data.byteswap(False)
# NOPE, didn't make things better
# -
# ## Create xarray DataArray, including some variable attributes
time = pd.date_range(st, periods=nts) # periods=num_mod_timesteps
time
# convert to xarray DataArray
swe = xr.DataArray(
numpy_data,
dims=('time', 'y', 'x'),
coords={'time': time, 'y': lat, 'x': lon}
)
# DataArray at this point has no attributes at all. Add attributes like name, units, etc.
swe.attrs['long_name'] = 'Snow Water Equivalent'
swe.attrs['standard_name'] = 'lwe_thickness_of_surface_snow_amount'
swe.attrs['units'] = 'meters'
fig, ax = plt.subplots(1, 2, figsize=(14, 6))
swe[1,:,:].plot(ax=ax[0])
swe[-1,:,:].plot(ax=ax[1]);
# ## Create xarray Dataset
# +
d = OrderedDict()
d['time'] = ('time', time)
d['x'] = ('x', lon)
d['y'] = ('y', lat)
d['swe'] = swe
ds = xr.Dataset(d)
# -
# Add global and variable attributes
# +
# Switch to using a dictionary (OrderdDict) to populate attributes
ds.attrs['description'] = "SnowModel model run, SWE variable only"
ds.attrs['CRS'] = "UTM Zone 12N, EPSG:32612"
ds.time.attrs['standard_name'] = "time"
ds.time.attrs['axis'] = "T"
ds.x.attrs['long_name'] = "Easting"
ds.x.attrs['units'] = "meters"
ds.x.attrs['axis'] = "X"
ds.y.attrs['long_name'] = "Northing"
ds.y.attrs['units'] = "meters"
ds.y.attrs['axis'] = "Y"
# -
ds
ds.swe.dims
ds.swe.attrs
# ### Export to netcdf
ds.to_netcdf('swed_gdat.nc', format='NETCDF4', engine='netcdf4')
# Verify that the netcdf file looks ok
ds_read = xr.open_dataset('swed_gdat.nc')
ds_read
# !ncdump -h 'swed_gdat.nc'
| awszarr/Read_SnowModeloutput_saveto_netcdf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# * 请在环境变量中设置`DB_URI`指向数据库
# +
# %matplotlib inline
import os
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from alphamind.api import *
from PyFin.api import *
plt.style.use('ggplot')
# +
"""
Back test parameter settings
"""
start_date = '2020-01-02'
end_date = '2020-02-21'
frequency = '10b'
ref_dates = makeSchedule(start_date, end_date, frequency, 'china.sse')
industry_lower = 1.0
industry_upper = 1.0
method = 'risk_neutral'
neutralize_risk = industry_styles
industry_name = 'sw'
industry_level = 1
benchmark_total_lower = 0.8
benchmark_total_upper = 1.0
horizon = map_freq(frequency)
weight_gap = 0.01
benchmark_code = 300
universe_name = 'hs300'
universe = Universe(universe_name)
executor = NaiveExecutor()
data_source = os.environ['DB_URI']
engine = SqlEngine(data_source)
# +
"""
Constraints settings
"""
industry_names = industry_list(industry_name, industry_level)
constraint_risk = ['SIZE', 'SIZENL', 'BETA'] + industry_names[:-1]
total_risk_names = constraint_risk + ['benchmark', 'total']
b_type = []
l_val = []
u_val = []
for name in total_risk_names:
if name == 'benchmark':
b_type.append(BoundaryType.RELATIVE)
l_val.append(benchmark_total_lower)
u_val.append(benchmark_total_upper)
elif name in {'SIZE', 'SIZENL', 'BETA'}:
b_type.append(BoundaryType.ABSOLUTE)
l_val.append(0.0)
u_val.append(0.0)
else:
b_type.append(BoundaryType.RELATIVE)
l_val.append(industry_lower)
u_val.append(industry_upper)
bounds = create_box_bounds(total_risk_names, b_type, l_val, u_val)
# +
def factor_analysis(engine, factor_name, universe, benchmark_code, positive):
"""
Data phase
"""
index_return = engine.fetch_dx_return_index_range(benchmark_code, start_date, end_date, horizon=horizon,
offset=1).set_index('trade_date')
codes_return = engine.fetch_dx_return_range(universe,
dates=ref_dates,
horizon=horizon,
offset=1,
benchmark=benchmark_code)
return_groups = codes_return.groupby('trade_date')
"""
Model phase: we need 1 constant linear model and one linear regression model
"""
industry_total = engine.fetch_industry_matrix_range(universe, dates=ref_dates, category=industry_name, level=industry_level)
industry_groups = industry_total.groupby('trade_date')
alpha_name = [str(factor_name) + '_' + ('pos' if positive else 'neg')]
simple_expression = CSRes(LAST(factor_name), 'EARNYILD') if positive else -CSRes(LAST(factor_name), 'EARNYILD')
const_features = {alpha_name[0]: simple_expression}
const_weights = {alpha_name[0]: 1.}
const_model = ConstLinearModel(features=alpha_name,
weights=const_weights)
const_model_factor_data = engine.fetch_data_range(universe,
factors=const_features,
dates=ref_dates,
benchmark=benchmark_code)['factor'].dropna()
rets = []
turn_overs = []
leverags = []
ics = []
index_dates = []
factor_groups = const_model_factor_data.groupby('trade_date')
for i, value in enumerate(factor_groups):
date = value[0]
data = value[1]
index_dates.append(date)
industry_matrix = industry_groups.get_group(date)
total_data = data.fillna(data[alpha_name].median())
total_data = pd.merge(total_data, industry_matrix, on=['code'])
alpha_logger.info('{0}: {1}'.format(date, len(total_data)))
risk_exp = total_data[neutralize_risk].values.astype(float)
benchmark_w = total_data.weight.values
is_in_benchmark = (benchmark_w > 0.).astype(float).reshape(-1, 1)
constraint_exp = total_data[constraint_risk].values
risk_exp_expand = np.concatenate((constraint_exp,
is_in_benchmark,
np.ones_like(is_in_benchmark)), axis=1).astype(float)
total_risk_exp = pd.DataFrame(risk_exp_expand, columns=total_risk_names)
constraints = LinearConstraints(bounds, total_risk_exp, benchmark_w)
lbound = np.maximum(0., benchmark_w - weight_gap)
ubound = weight_gap + benchmark_w
factor_values = factor_processing(total_data[alpha_name].values,
pre_process=[winsorize_normal, standardize],
risk_factors=risk_exp,
post_process=[winsorize_normal, standardize])
# const linear model
er = const_model.predict(pd.DataFrame(data={alpha_name[0]: factor_values.flatten()}))
alpha_logger.info('{0} full re-balance'.format(date))
target_pos, _ = er_portfolio_analysis(er,
total_data.industry_name.values,
None,
constraints,
False,
benchmark_w,
method=method,
lbound=lbound,
ubound=ubound)
target_pos['code'] = total_data['code'].values
turn_over, executed_pos = executor.execute(target_pos=target_pos)
dx_returns = return_groups.get_group(date)
result = pd.merge(executed_pos, total_data[['code', 'weight']], on=['code'], how='inner')
result = pd.merge(result, dx_returns, on=['code'])
leverage = result.weight_x.abs().sum()
excess_return = np.exp(result.dx.values) - 1. - index_return.loc[date, 'dx']
raw_weight = result.weight_x.values
activate_weight = raw_weight - result.weight_y.values
ret = raw_weight @ excess_return
risk_adjusted_ic = np.corrcoef(excess_return, activate_weight)[0, 1]
rets.append(np.log(1. + ret))
ics.append(risk_adjusted_ic)
executor.set_current(executed_pos)
turn_overs.append(turn_over)
leverags.append(leverage)
alpha_logger.info('{0} is finished'.format(date))
ret_df = pd.DataFrame({'returns': rets, 'turn_over': turn_overs, 'IC': ics, 'leverage': leverags}, index=index_dates)
ret_df.loc[advanceDateByCalendar('china.sse', ref_dates[-1], frequency)] = 0.
ret_df = ret_df.shift(1)
ret_df.iloc[0] = 0.
ret_df['tc_cost'] = ret_df.turn_over * 0.002
return alpha_name[0], ret_df
def worker_func_positive(factor_name):
from alphamind.api import SqlEngine
engine = SqlEngine(data_source)
return factor_analysis(engine, factor_name, universe, benchmark_code, positive=True)
def worker_func_negative(factor_name):
from alphamind.api import SqlEngine
engine = SqlEngine(data_source)
return factor_analysis(engine, factor_name, universe, benchmark_code, positive=False)
# -
factors = ["EMA5D", "EMV6D"]
# +
# # %%time
res1 = [worker_func_positive(factor) for factor in factors]
res2 = [worker_func_negative(factor) for factor in factors]
factor_df = pd.DataFrame()
ic_df = pd.DataFrame()
for f_name, res in res1:
factor_df[f_name] = res['returns']
ic_df[f_name] = res['IC']
for f_name, res in res2:
factor_df[f_name] = res['returns']
ic_df[f_name] = res['IC']
# +
factor_res = factor_df.agg(['mean', 'std']).T
factor_res['t.'] = factor_res['mean'] / factor_res['std'] * np.sqrt(len(factor_df))
ic_res = ic_df.agg(['mean', 'std']).T
ic_res['t.'] = ic_res['mean'] / ic_res['std'] * np.sqrt(len(ic_df))
# -
with pd.ExcelWriter(f'{universe_name}_{benchmark_code}.xlsx', engine='xlsxwriter') as writer:
factor_df.to_excel(writer, sheet_name='ret')
ic_df.to_excel(writer, sheet_name='ic')
factor_res.to_excel(writer, sheet_name='ret_stat')
ic_res.to_excel(writer, sheet_name='ic_stat')
| notebooks/Example 1 - Factor IC analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 5
# #### Name : <NAME>¶
# #### ID : PY04233
# #### Date : 11/12/2019
# #### Subject : Python Onlin
# # Q1
# +
def factorial(num):
if num == 1:
return 1
else:
return num * factorial(num - 1)
factorial(5)
# -
# # Q2
def lengthOfString(string):
dict1 = {"Upper" : 0, "Lower" : 0}
for char in string:
if char.isupper():
dict1["Upper"] += 1
elif char.islower():
dict1["Lower"] += 1
else:
pass
print(f"Result : {dict1}")
string = str(input("Enter String : "))
lengthOfString(string)
# # Q3
def even(list1):
for i in list1:
if i % 2 == 0:
print(f"Even : {i}")
list1 = [ 1, 3, 5, 2, 4, 27, 88, 964]
even(list1)
# # Q4
def pallendrom(string2):
string2 = string2.lower()
if string2 == string2[::-1]:
print(f"{string2} is Pallendrom")
else:
print(f"{string2} is not Pallendrom")
pallendrom("maDam")
# # Q5
def prime(num):
if num > 1:
for i in range(2, num):
if num % i == 0:
print(f"{num} is not prime")
print(f"{i} times {num/i} is {num} ")
break
else:
print(f"{num} is prime")
break
else:
print(f"{num} culd not be a prime")
prime(11)
# # Q6
# +
def items(*opt):
print(opt)
items("Masala", "Meddison", "Watch")
# -
| Assignments/Certified Python By Saylani/Assignment 5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sigeisler/robustness_of_gnns_at_scale/blob/notebook/notebooks/Quick_start_robustness_gnns_at_scale.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ixdVBV5PugOh"
# # Robustness of Graph Neural Networks at Scale - Quick Start
#
# This notebook can be run in google colab and serves as a quick introduction to the [Robustness of Graph Neural Networks at Scale](https://github.com/sigeisler/robustness_of_gnns_at_scale) repository.
#
# ## 0. Setup
#
# First, let's get the code and install requirements.
#
# + id="srl7gR-Gy5gX"
# clone package repository
# !git clone https://github.com/sigeisler/robustness_of_gnns_at_scale.git
# navigate to the repository
# %cd robustness_of_gnns_at_scale
# install package requirements
# !pip install -r requirements.txt
# !pip install -r requirements-dev.txt
# install package
# # !python setup.py install
# !pip install --use-feature=in-tree-build .
# build kernels
# !pip install --use-feature=in-tree-build ./kernels
# + [markdown] id="FQScUvGr-b08"
# ### Imports
# + id="7ztTeoK3-Vqh"
from matplotlib import pyplot as plt
from experiments import (
experiment_train,
experiment_local_attack_direct,
experiment_global_attack_direct
)
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# + [markdown] id="Rb2y6lUbdYi_"
# ## 1. Training
#
# For the training and evaluation code we decided to provide Sacred experiments which make it very easy to run the same code from the command line or on your cluster. To train or attack the models you can use the `script_execute_experiment` script and simply specify the respective configuration or execute the experiment directly by passing the desired configuration in [experiments/experiment_train.py](https://github.com/sigeisler/robustness_of_gnns_at_scale/blob/main/experiments/experiment_train.py#L74).
#
# In the example below, we train a `GCN` on `Cora ML`.
# + id="6mga1GGvdbLp" colab={"base_uri": "https://localhost:8080/", "height": 717, "referenced_widgets": ["9294fcc4c6e04d57a05867d8264ee17e", "d889fb0f161f4d56bab40c559d2fd258", "08e10fade58341be954e200140312400", "6c3e126c918b4a76b9f9a1f00cbf3449", "91c8ae4a4d2d42eba3a29d769b1a008e", "4d6a0605ef7e421f9c0ff128a79e7b4a", "78e019c77f344834931ec885b6680a67", "c29741c70fb54c4aa84ae95a9566c6d9"]} outputId="9f1f1819-8905-41a6-af57-70a65104ab07"
train_statistics = experiment_train.run(
data_dir = './data',
dataset = 'cora_ml',
model_params = dict(
label="Vanilla GCN",
model="GCN",
do_cache_adj_prep=True,
n_filters=64,
dropout=0.5,
svd_params=None,
jaccard_params=None,
gdc_params={"alpha": 0.15, "k": 64}),
train_params = dict(
lr=1e-2,
weight_decay=1e-3,
patience=300,
max_epochs=3000),
binary_attr = False,
make_undirected = True,
seed=0,
artifact_dir = 'cache',
model_storage_type = 'demo',
ppr_cache_params = dict(),
device = 0,
data_device = 0,
display_steps = 100,
debug_level = "info"
)
# plot train and val loss curves
fig, ax = plt.subplots()
color = plt.rcParams['axes.prop_cycle'].by_key()['color'][0]
ax.set_xlabel('Epoch $t$')
ax.set_ylabel("Loss")
ax.plot(train_statistics['trace_train'], color=color, label='Train')
color = plt.rcParams['axes.prop_cycle'].by_key()['color'][1]
ax.plot(train_statistics['trace_val'], color=color, label='Val')
ax.legend()
plt.gcf().show()
# + colab={"base_uri": "https://localhost:8080/"} id="Diozgu7Vivj7" outputId="f6d7227f-fa9c-4e4a-d591-bb2a70ca8755"
clean_acc = train_statistics["accuracy"]
print(f'Accuracy of the model: {100*clean_acc:.2f}%')
# + [markdown] id="JzCocLocdb2y"
# ## 2. Evaluation
#
# For evaluation, we use the locally stored models. Similarly to training, we provide a script that runs the attacks for different seeds for all pretrained models. For all experiments, please check out the [config](https://github.com/sigeisler/robustness_of_gnns_at_scale/tree/main/config) folder.
#
# ### 2.1 Local PR-BCD Attack
# We provide an example for a `local PR-BCD` attack on the `Vanilla GCN` model trained previously by passing the desired configuration in [experiments/experiment_local_attack_direct.py](https://github.com/sigeisler/robustness_of_gnns_at_scale/blob/main/experiments/experiment_local_attack_direct.py#L68).
# + colab={"base_uri": "https://localhost:8080/"} id="A-n0IWrJC1QQ" outputId="94944c64-dcac-46b8-c2a2-d33ea4e3b43a"
local_prbcd_statistics = experiment_local_attack_direct.run(
data_dir = './data',
dataset = 'cora_ml',
attack = 'LocalPRBCD',
attack_params = dict(
ppr_cache_params = dict(
data_artifact_dir = 'cache',
data_storage_type = 'ppr'),
epochs = 500,
fine_tune_epochs = 100,
search_space_size = 10_000,
ppr_recalc_at_end = True,
loss_type = 'Margin',
lr_factor = 0.05),
nodes = None,
nodes_topk = 5,
seed=0,
epsilons = [1],
min_node_degree = None,
binary_attr = False,
make_undirected = True,
artifact_dir = 'cache',
model_label = 'Vanilla GCN',
model_storage_type = 'demo',
device = 0,
data_device = 0,
debug_level = "info"
)
# + colab={"base_uri": "https://localhost:8080/"} id="Saio7dZwlulQ" outputId="87df7551-21ec-4acb-fe03-96b52175dac4"
n_change = 0
for node in local_prbcd_statistics['results']:
if node['margin'] < 0:
n_change += 1
flipped_ratio = n_change / len(local_prbcd_statistics['results'])
print(f'Percentage of changed node predictions : {100*flipped_ratio:.2f}%')
# + [markdown] id="UMtpxI6cCkwz"
# ### 2.2 Global PR-BCD Attack
#
# Now let's do the same with a non-local `PR-BCD` attack. For this, we use [experiments/experiment_global_attack_direct.py](https://github.com/sigeisler/robustness_of_gnns_at_scale/blob/main/experiments/experiment_global_attack_direct.py#L63).
# + colab={"base_uri": "https://localhost:8080/"} id="NdZaHYI8Had1" outputId="a7ae6789-4241-432f-ea5d-8a4dd9f268c1"
global_prbcd_statistics = experiment_global_attack_direct.run(
data_dir = './data',
dataset = 'cora_ml',
attack = 'PRBCD',
attack_params = dict(
epochs=500,
fine_tune_epochs=100,
keep_heuristic="WeightOnly",
search_space_size=100_000,
do_synchronize=True,
loss_type="tanhMargin",
),
epsilons = [0.25],
binary_attr = False,
make_undirected = True,
seed=0,
artifact_dir = 'cache',
pert_adj_storage_type = 'evasion_global_adj',
pert_attr_storage_type = 'evasion_global_attr',
model_label = 'Vanilla GCN',
model_storage_type = 'demo',
device = 0,
data_device = 0,
debug_level = "info"
)
print(global_prbcd_statistics)
# + colab={"base_uri": "https://localhost:8080/"} id="UQwlL5mnkXHO" outputId="03150aca-0629-443b-c494-a01bdd06b653"
perturbed_acc = global_prbcd_statistics["results"][0]['accuracy']
print(f'Clean accuracy: {100*clean_acc:.2f}%; Accuracy after global PRBCD attack: {100*perturbed_acc:.2f}%')
| notebooks/Quick_start_robustness_gnns_at_scale.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/dimi-fn/Various-Data-Science-Scripts/blob/main/special_matrices.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="FC4zCCrz9j9z"
# Writing a function that will test if a 4x4 matrix is singular before calculating it.
#
# Note: A matrix A is singular if an inverse matrix A-1 exists.
#
# The function goes through the matrix replacing each row so that it can be turned into the echelor form. A matrix is in elechon form if it has the shape resulting from a Gaussian elimination [reference](https://en.wikipedia.org/wiki/Row_echelon_form)
# + id="mMh73IfY9lD0"
import numpy as np
# + id="UdWNJzJ39tf8"
# Creating a function that will go through the matrix replacing each row in order to turn it into the echelon form.
def is_Singular(a) :
# making matrix b as a copy of a (since its values are going to be altered)
b = np.array(a, dtype=np.float_)
try:
fix_row_zero(b)
fix_row_one(b)
fix_row_two(b)
fix_row_three(b)
except matrix_is_Singular:
return True
return False
# + id="mXeXtE2t9v-9"
# handling errors
class matrix_is_Singular(Exception): pass
# + id="GdY7VgbZ9xVL"
# In the first row a[0], the 1st element a[0,0] should be equal to 1
# In the end, the row is divided by the value of a[0, 0]
def fix_row_zero(a) :
# check if a[0,0] is equal to zero
# if a[0,0]=0, one of the lower rows will be added to the first one before the division
if a[0,0] == 0 :
a[0] = a[0] + a[1]
if a[0,0] == 0 :
a[0] = a[0] + a[2]
if a[0,0] == 0 :
a[0] = a[0] + a[3]
if a[0,0] == 0 :
raise matrix_is_Singular()
# divide the 1st row by the value of the 1st element a[0, 0]
a[0] = a[0] / a[0,0]
return a
# + id="MaQ1BKhf9zGH"
# 2nd row
def fix_row_one(a):
# setting the sub-diagonal elements to zero; in the 2nd row there is only the a[1,0] element
a[1] = a[1] - a[1,0] * a[0]
# check if the sub-diagonal element a[1,1] is equal to zero
if a[1,1] == 0 :
a[1] = a[1] + a[2]
a[1] = a[1] - a[1,0] * a[0]
if a[1,1] == 0 :
a[1] = a[1] + a[3]
a[1] = a[1] - a[1,0] * a[0]
if a[1,1] == 0 :
raise matrix_is_Singular()
# divide the 2nd row by the value of a[1, 1]
a[1] = a[1] / a[1,1]
return a
# + id="gCmfmUE290zb"
# 3rd row
def fix_row_two(a) :
# setting the sub-diagonal elements to zero, i.e. a[2,0] and a[2,1]
a[2] = a[2] - a[2,0] * a[0]
a[2] = a[2] - a[2,1] * a[1]
# check if the sub-diagonal elements a[2,0] and a[2,1] are equal to zero
if a[2,2] == 0 :
a[2] = a[2] + a[3]
a[2] = a[2] - a[2,0] * a[0]
a[2] = a[2] - a[2,1] * a[1]
a[2] = a[2] + a[3]
a[2] = a[2] - a[1,0] * a[0]
if a[2,2] == 0 :
raise matrix_is_Singular()
# setting the diagonal element to one by dividing the 3rd row by the value of a[2,2]
a[2] = a[2] / a[2,2]
return a
# + id="ZC2bdmja92sM"
# 4th row
def fix_row_three(a) :
# setting the sub-diagonal elements to zero, i.e. elements a[3,0], a[3,1], and a[3,2]
a[3] = a[3] - a[3,0] * a[0]
a[3] = a[3] - a[3,1] * a[1]
a[3] = a[3] - a[3,2] * a[2]
if a[3,3] == 0 :
raise matrix_is_Singular()
# Transform the row to set the diagonal element to one.
# setting the diagonal element a[3,3] to one by dividing the 4th row by the value of a[3,3]
a[3] = a[3] / a[3,3]
return a
| Maths - Statistics/Mathematics_for_ML/special_matrices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Full packet capture
#
# ## /Me
#
# > https://github.com/markuskont/Talsec-meetup
stuff = [
"IDS",
"PCAP",
"coding",
"teaching",
"hunting",
"devops",
"logging",
"/^.*data.*$/"
]
speciality = "spec==[{}]".format(",".join(stuff[:-1]))
speciality = " || ".join([speciality, "spec=={}".format(stuff[-1])])
print(speciality)
# ## /Owl
# > Moloch is a large scale, open source, indexed packet capture and search system.
#
# * https://molo.ch/
# * https://github.com/aol/moloch
# * https://github.com/ccdcoe/CDMCS/tree/master/Moloch
# ## Architecture
# ### Simple
# +
from graphviz import Digraph, Source
dot = Digraph(comment='moloch-general')
dot.attr(compound='true')
dot.format = 'png'
dot.landscape = True
dot.node("m", "Mirror")
dot.node("c", "Capture")
dot.node("v", "Viewer")
dot.node("w", "WISE")
dot.node("e", "Elastic")
dot.node("f", "PCAP file")
dot.node("z", "...")
dot.edge("m", "c")
dot.edge("c", "f")
dot.edge("c", "e")
dot.edge("e", "v")
dot.edge("f", "v")
dot.edge("z", "w")
dot.edge("w", "c")
dot.edge("w", "v")
Source(dot)
# -
# ### General
# +
from graphviz import Digraph, Source
dot = Digraph(comment='moloch-general')
dot.attr(compound='true')
dot.format = 'png'
dot.landscape = True
with dot.subgraph(name="cluster_network") as net:
net.node("m", "Mirror")
with dot.subgraph(name="cluster_moloch") as molo:
molo.attr(label="Moloch")
molo.node("c", "Capture")
molo.node("v", "Viewer")
molo.node("w", "Wise")
with dot.subgraph(name="cluster_elastic") as elastic:
elastic.attr(label="Elastic")
elastic.node("ep", "Proxy")
elastic.node("em", "Master")
elastic.node("ed", "Data")
elastic.edge("ed", "ep")
elastic.edge("em", "ep")
elastic.edge("em", "ed")
with dot.subgraph(name="cluster_filesystem") as fs:
fs.attr(label="Filesystem")
fs.node("d", "Pcap files")
fs.node("g", "GeoIP database")
fs.node("f", "File")
with dot.subgraph(name="cluster_datasource") as ds:
ds.attr(label="External")
ds.node("et", "Emergingthreats")
ds.node("r", "Redis")
ds.node("z", "...")
with dot.subgraph(name="cluster_IDS") as suri:
suri.attr(label="IDS")
suri.node("s", "Suricata")
suri.node("j", "eve.json")
suri.edge("s", "j")
dot.edge("g", "c")
dot.edge("c", "d")
dot.edge("c", "ep")
dot.edge("ep", "v")
dot.edge("d", "v")
dot.edge("w", "c")
dot.edge("w", "v")
dot.edge("r", "w")
dot.edge("f", "w")
dot.edge("et", "w")
dot.edge("z", "w")
dot.edge("j", "c")
dot.edge("m", "c")
dot.edge("m", "s")
Source(dot)
# -
# ### Mine (2019)
#
# * https://gist.github.com/markuskont/734a9ec946bf40801494f14b368a0668
# * https://github.com/ccdcoe/frankenstack/blob/master/states/yellow/data/elastic.sls
# * https://github.com/ccdcoe/frankenstack/blob/master/pillar/worker.sls
# * https://github.com/markuskont/moloch/blob/custom/wisePlugins/wiseService/source.ls19.js
# +
from graphviz import Digraph, Source
dot = Digraph(comment='moloch-general')
dot.attr(compound='true')
dot.attr(ratio="compress")
dot.format = 'png'
dot.landscape = False
with dot.subgraph(name="cluster_net") as net:
net.attr(label="Lab network")
net.node("erspan", "ERSPAN")
net.node("gigamon", "Packet broker")
net.edge("erspan", "gigamon")
with dot.subgraph(name="cluster_server1") as s:
s.attr(label="Server 1 - Golden")
s.node("s1cap", "Intel X710")
s.node("disk", "RAID 10")
s.node("oride", "override.ini")
s.node("syslog", "Rsyslog")
s.node("af", "Suricata Alert File")
s.edge("syslog", "af")
s.node("tgrf1", "Telegraf")
s.edge("s1cap", "tgrf1")
with s.subgraph(name="cluster_Moloch") as molo:
molo.attr(label="Moloch")
molo.node("c", "Capture")
molo.node("v", "Viewer")
molo.node("w", "WISE")
molo.node("sls19", "source.ls19.js")
s.edge("c", "disk")
s.edge("disk", "v")
s.edge("oride", "c")
s.edge("af", "c")
molo.edge("w", "c")
molo.edge("w", "v")
molo.edge("sls19", "w")
s.edge("s1cap", "c")
with dot.subgraph(name="cluster_server2") as s:
s.attr(label="Server 2 - Experimental")
s.node("s2cap", "Intel X710")
s.node("meer", "Suricata")
s.edge("s2cap", "meer")
s.node("disk2", "RAID 0")
s.edge("meer", "disk2")
s.node("syslog2", "Rsyslog")
s.edge("meer", "syslog2")
s.node("elastic2", "Elastic")
s.node("tgrf2", "Telegraf")
s.edge("s2cap", "tgrf2")
s.edge("meer", "tgrf2")
with dot.subgraph(name="cluster_mon_vm") as mon:
mon.attr(label="Monitoring VM")
mon.node("go", "Asset API")
mon.node("syslogC", "Central Rsyslog")
mon.node("flux", "InfluxDB")
mon.node("grfna", "Grafana")
mon.node("elasticC", "Elastic")
mon.edge("grfna", "flux")
mon.edge("syslogC", "elasticC")
with dot.subgraph(name="cluster_dswarm") as dsrm:
dsrm.attr(label="Docker Swarm")
for i in [0,1,2]:
with dsrm.subgraph(name="cluster_vm_hot_{}".format(i)) as host:
host.attr(label="Elastic HOT VM-{}".format(i))
host.node("em-{}".format(i), "Master-{}".format(i))
host.node("ed-{}".format(i), "Data-{}".format(i))
host.node("ep-{}".format(i), "Proxy-{}".format(i))
host.edge("ed-{}".format(i), "ep-{}".format(i))
host.edge("em-{}".format(i), "ep-{}".format(i))
host.edge("em-{}".format(i), "ed-{}".format(i))
dot.edge("c", "ep-{}".format(i))
dot.edge("ep-{}".format(i), "v")
for i in [5,6,7]:
with dsrm.subgraph(name="cluster_vm_cold_{}".format(i)) as host:
host.attr(label="Elastic COLD VM-{}".format(i))
host.node("em-{}".format(i), "Master-{}".format(i))
host.node("ed-{}".format(i), "Data-{}".format(i))
host.node("ep-{}".format(i), "Proxy-{}".format(i))
host.edge("ed-{}".format(i), "ep-{}".format(i))
host.edge("em-{}".format(i), "ep-{}".format(i))
host.edge("em-{}".format(i), "ed-{}".format(i))
dot.node("score", "Scoring server")
dot.node("collab", "WIKI")
dot.node("vsphere", "Vsphere")
dot.node("ps1", "Powershell")
dot.edge("vsphere", "ps1")
dot.edge("ps1", "go")
dot.edge("collab", "go")
dot.edge("score", "go")
dot.edge("go", "sls19")
dot.edge("syslog2", "syslog")
dot.edge("gigamon", "s1cap")
dot.edge("gigamon", "s2cap")
dot.edge("syslog", "syslogC")
dot.edge("syslog2", "syslogC")
dot.edge("tgrf1", "flux")
dot.edge("tgrf2", "flux")
Source(dot)
# -
# ## Stats
#
# ```
# ➜ ls19-moloch find 2019/ -type f -name '*.pcap' | wc -l
# 984
# ```
# ```
# ➜ ls19-moloch du -hs 2019/*
# 4.0T 2019/09
# 4.7T 2019/10
# 2.9T 2019/11
# ```
# ```
# ➜ ls19-moloch du -hs $DATA/ls19/moloch/elastic/docker-volume
# 956G $DATA/ls19/moloch/elastic/docker-volume
# ```
#
# ```
# ➜ ls19-moloch curl -ss localhost:9255/_cat/indices | grep open | grep sessions2 | egrep "190409|190410|190411" | sort -h
# green open sessions2-190409h00 OVRxicRBTPSlSYDvqfMGJg 5 0 12302252 0 14.2gb 14.2gb
# green open sessions2-190409h01 SbCpm0_YTx-lGwH-lwZtgw 5 0 12906843 0 15gb 15gb
# green open sessions2-190409h02 kHuKvQYDSXiDupsygdOjvw 5 0 12931933 0 15gb 15gb
# green open sessions2-190409h03 DUhy8HquSK6llnXno00YSg 5 0 12915792 0 15.1gb 15.1gb
# green open sessions2-190409h04 QVbw-0EEQW-birmT3j2xyQ 5 0 13077484 0 15.3gb 15.3gb
# green open sessions2-190409h05 Aq9eNYbcTk-gviUy8PJkHg 5 0 13479043 0 15.9gb 15.9gb
# green open sessions2-190409h06 OH58gsSFRsOgsnWk8ljQjw 5 0 15924732 1 19.3gb 19.3gb
# green open sessions2-190409h07 kf-YeQWaQCKjQSibED1ZKw 5 0 21196529 0 25.4gb 25.4gb
# green open sessions2-190409h08 TUbq23q5TtiQbVyyCKOnpw 5 0 21417591 1 26gb 26gb
# green open sessions2-190409h09 1Y-aAl_VTa2fMyQEQEv7DQ 5 0 22544788 0 28.4gb 28.4gb
# green open sessions2-190409h10 D_-EeHXiRJW59YYmUg7vHw 5 0 23838898 2 27.8gb 27.8gb
# green open sessions2-190409h11 -jyLNcYjT2GAujuftrqpLg 5 0 26553756 0 30.9gb 30.9gb
# green open sessions2-190409h12 1kxjtQNITp6YEB2i1A125g 5 0 22696376 0 26.5gb 26.5gb
# green open sessions2-190409h13 R8iXgdU_TC--Icdy7RsXHA 5 0 21163617 2 27.8gb 27.8gb
# green open sessions2-190409h14 wuqxd2heTMeA8NaRZMqCHA 5 0 15927452 0 19.5gb 19.5gb
# green open sessions2-190409h15 yCS2JXWyT_S9afYdG0rcsA 5 0 3383608 0 3.3gb 3.3gb
# green open sessions2-190409h16 x2LSNzcbTNK8OLVecFy5pg 5 0 5701935 0 7gb 7gb
# green open sessions2-190409h17 3H8Q5x5PQD6C_A3_LUMGVA 5 0 9677885 0 11.8gb 11.8gb
# green open sessions2-190409h18 v90fVz7_Sz-xs9hsdWW7FA 5 0 10173588 0 11.4gb 11.4gb
# green open sessions2-190409h19 _SC6qAYgTau1n_qN1MO89A 5 0 7354839 0 7.9gb 7.9gb
# green open sessions2-190409h20 mH8C4ekQRXaHup2u-uFnXg 5 0 9671269 0 11.2gb 11.2gb
# green open sessions2-190409h21 _KAZV-iGREyCxGSWXxco5g 5 0 10560991 0 12.6gb 12.6gb
# green open sessions2-190409h22 Ws3PW1rBSkqioPLfjYpJog 5 0 10462608 0 12.3gb 12.3gb
# green open sessions2-190409h23 DsHSqT4lR9O0pUH49i_Mqw 5 0 11027245 0 13gb 13gb
# green open sessions2-190410h00 9X6HyeLJRqWrZowwhM1clA 5 0 10446490 0 12.4gb 12.4gb
# green open sessions2-190410h01 _hM1waSsRGOuEHX91svZ6w 5 0 10348412 0 12.3gb 12.3gb
# green open sessions2-190410h02 eufIexQxT8usg8Woic2ZCQ 5 0 10536682 0 12.6gb 12.6gb
# green open sessions2-190410h03 tLYsjM06SvulmA1wkZpbgg 5 0 10481627 0 12.4gb 12.4gb
# green open sessions2-190410h04 ZIotJF6GT3yDZzVT-jkdqA 5 0 10607329 0 13gb 13gb
# green open sessions2-190410h05 N6F76JTgQa2kkSZfY6R0bA 5 0 11618714 0 14.1gb 14.1gb
# green open sessions2-190410h06 uc11PcM_RqiXTbnK9ABLGQ 5 0 17067197 1 25.8gb 25.8gb
# green open sessions2-190410h07 n9xeMdJEQX6MCf6tKexUYg 5 0 20652101 0 26.4gb 26.4gb
# green open sessions2-190410h08 H-twZvhmT_al9opxXpEmOg 5 0 19042897 0 25.4gb 25.4gb
# green open sessions2-190410h09 hZTy4SKTQi2ih6JZklWAqg 5 0 19801400 0 25.7gb 25.7gb
# green open sessions2-190410h10 na-tVioqSQeFNThD-AlAlg 5 0 20628343 37 26.4gb 26.4gb
# green open sessions2-190410h11 DEdpgeaeRzyJ6dqwwfaeyA 5 0 20639794 52 24.9gb 24.9gb
# green open sessions2-190410h12 l0T9gpHuSZesDvXtiRc0IQ 5 0 21995421 89 27.1gb 27.1gb
# green open sessions2-190410h13 OEssatZFR9m7pgnf2Ht9KA 5 0 20636339 62 26.9gb 26.9gb
# green open sessions2-190410h14 W7O7BeagT2CEmwFoCirOMg 5 0 16939975 0 19.1gb 19.1gb
# green open sessions2-190410h15 dxv4y3ZYQxauG9gycDXdyA 5 0 25505640 0 20.2gb 20.2gb
# green open sessions2-190410h16 arlxCdJ6TNu4zDZDsU2Ctw 5 0 17553667 0 15.1gb 15.1gb
# green open sessions2-190410h17 W_KPaA5NSzuUe-xmmYwDEw 5 0 13599611 0 13gb 13gb
# green open sessions2-190410h18 PmdYgQmnSoO5uo4gIb2uOg 5 0 25547795 0 20.6gb 20.6gb
# green open sessions2-190410h19 JbEJk9B6RyGCy5ki2q3hNA 5 0 11592538 0 11.1gb 11.1gb
# green open sessions2-190410h20 hyGeHPTlR7649RPWseZZ0Q 5 0 11095865 0 11.1gb 11.1gb
# green open sessions2-190410h21 VIR-Yy2XSoqND4sYp0JoVA 5 0 11278815 0 11.2gb 11.2gb
# green open sessions2-190410h22 YymDM6DKSICss_QFPWETGQ 5 0 10743235 0 10.8gb 10.8gb
# green open sessions2-190410h23 8gNnqL6aS4yBuL8onlVlpw 5 0 10122448 0 10.3gb 10.3gb
# green open sessions2-190411h00 XNpOg_waQHGoFpBKVMB24g 5 0 9682173 0 9.9gb 9.9gb
# green open sessions2-190411h01 EEO_KKtLQdeZNCF_AU73NA 5 0 9718077 0 10gb 10gb
# green open sessions2-190411h02 7BttoZ-bRjC38xwGIpW3JA 5 0 10576311 0 10.7gb 10.7gb
# green open sessions2-190411h03 5MurV4_0SFeuQsRpCynvrQ 5 0 10659113 0 10.7gb 10.7gb
# green open sessions2-190411h04 Etd83aM0R8ul-mOv-a9ajQ 5 0 10785198 0 10.8gb 10.8gb
# green open sessions2-190411h05 xzeS4ay-TpGG0INGhACg9g 5 0 12309578 0 13.6gb 13.6gb
# green open sessions2-190411h06 65TvKFIBSaON7ejJ25WfSA 5 0 19400827 3 23.7gb 23.7gb
# green open sessions2-190411h07 ZGPUO4heT32Xs3wNpYG3Vg 5 0 20396488 3 25.7gb 25.7gb
# green open sessions2-190411h08 AFkt2-ItR_asYXzJx9ThsA 5 0 22451368 0 27.1gb 27.1gb
# green open sessions2-190411h09 AWEwej_tR0-pZyNeVN9VaQ 5 0 22499734 0 26.2gb 26.2gb
# green open sessions2-190411h10 nsaeI6EyREu-hrz0bAfuFA 5 0 21557422 0 26.1gb 26.1gb
# green open sessions2-190411h11 YYcL9TuEQDaCyo_OdCgXtg 5 0 20941641 0 24.8gb 24.8gb
# green open sessions2-190411h12 igozBC31TN65IiP7jRYYBQ 5 0 20182659 1 25.4gb 25.4gb
# green open sessions2-190411h13 IQJcyRsERbaNVUv2iJ6E4w 5 0 20135921 0 23.6gb 23.6gb
# green open sessions2-190411h14 _BYF3V5MTiOH_uK8MxVjbQ 5 0 645930 0 847.9mb 847.9mb
# ```
# ## Performance optimizations
#
# * https://github.com/ccdcoe/CDMCS/tree/master/Moloch/tuning
# * https://github.com/pevma/SEPTun
# * https://github.com/pevma/SEPTun-Mark-II
| 000.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Quick Draw venv
# language: python
# name: venv
# ---
# +
data = []
with open('train.log', 'r+') as log_file:
data = log_file.read().split('\n')
import re
def clean_num(digit):
result = re.sub('[^0-9 | ^\.]','', digit)
return float(result)
dis_loss = []
dis_acc = []
gen_loss = []
gen_acc = []
for line in data:
line = line.split(" ")
if len(line) < 6: continue
epoch = line[0]
dis_l, dis_a = clean_num(line[3]), clean_num(line[5])
dis_loss.append(dis_l)
dis_acc.append(dis_a)
gen_l, gen_a = clean_num(line[-3]), clean_num(line[-1])
gen_loss.append(gen_l)
gen_acc.append(gen_a)
# +
import numpy as np
import matplotlib.pyplot as plt
plt.subplot(2, 1, 2)
plt.plot(dis_loss)
plt.ylabel("Loss")
plt.xlabel("Epoch")
plt.title("Discriminator Training Loss")
plt.show()
plt.subplot(2, 1, 2)
plt.plot(dis_acc)
plt.ylabel("Accuracy")
plt.xlabel("Epoch")
plt.title("Discriminator Training Accuracy")
plt.show()
plt.subplot(2, 1, 2)
plt.plot(gen_loss)
plt.ylabel("Loss")
plt.xlabel("Epoch")
plt.title("Generator Training Loss")
plt.show()
plt.subplot(2, 1, 2)
plt.plot(gen_acc)
plt.ylabel("Accuracy")
plt.xlabel("Epoch")
plt.title("Generator Training Accuracy")
plt.show()
#plt.savefig("dis_train")
# +
import pandas as pd
import numpy as np
classes = ['saxophone',
'raccoon',
'piano',
'panda',
'leg',
'headphones',
'ceiling_fan',
'bed',
'basket',
'aircraft_carrier']
channels = 1
img_size = 28
n_classes = 10
latent_dim = 100
batch_size = 32
learning_rate = .0002
b1 = .5
b2 = .999
sample_interval = 400
n_epochs = 2
img_shape = (channels, img_size, img_size)
df = pd.DataFrame([], columns=['Image', 'Label'])
for i, label in enumerate(classes):
data = np.load('../data/%s.npy' % label) / 255
data = np.reshape(data, [data.shape[0], img_size, img_size, 1])
df2 = pd.DataFrame([(row, i) for row in data], columns=['Image', 'Label'])
df = df.append(df2)
df.sample(frac=1)
# -
real_imgs = df.sample(200)['Image']
# +
import matplotlib.pyplot as plt
plt.figure(figsize=(5,5))
for k in range(16):
plt.subplot(4, 4, k+1)
plt.imshow(real_imgs.iloc[k].reshape((img_size, img_size)), cmap='gray')
plt.axis('off')
plt.tight_layout()
plt.show()
| src/notebooks/Plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas as pd
import numpy as np
import os
import math
import graphlab
import graphlab as gl
import graphlab.aggregate as agg
'''钢炮'''
path = '/home/zongyi/bimbo_data/'
train = gl.SFrame.read_csv(path + 'train_fs_w9.csv', verbose=False)
town = gl.SFrame.read_csv(path + 'towns.csv', verbose=False)
town = town['Agencia_ID','Producto_ID','tcc']
train = train.join(town, on=['Agencia_ID','Producto_ID'], how='left')
# train = train.fillna('t_c',1)
train = train.fillna('tcc',0)
# train = train.fillna('tp_sum',0)
del town
# +
# relag_train = gl.SFrame.read_csv(path + 're_lag_train.csv', verbose=False)
# train = train.join(relag_train, on=['Cliente_ID','Producto_ID','Semana'], how='left')
# train = train.fillna('re_lag1',0)
# train = train.fillna('re_lag2',0)
# train = train.fillna('re_lag3',0)
# train = train.fillna('re_lag4',0)
# train = train.fillna('re_lag5',0)
# del relag_train
# +
# pd = gl.SFrame.read_csv(path + 'products.csv', verbose=False)
# train = train.join(pd, on=['Producto_ID'], how='left')
# train = train.fillna('prom',0)
# train = train.fillna('weight',0)
# train = train.fillna('pieces',1)
# train = train.fillna('w_per_piece',0)
# train = train.fillna('healthy',0)
# train = train.fillna('drink',0)
# del train['brand']
# del train['NombreProducto']
# del pd
# +
# client = gl.SFrame.read_csv(path + 'clients.csv', verbose=False)
# train = train.join(client, on=['Cliente_ID'], how='left')
# del client
# -
del train['prior_sum']
del train['lag_sum']
# del train['week_times']
# del train['Semana']
del train['n_t']
print train.column_names()
print len(train.column_names())
# +
# Make a train-test split
# train_data, test_data = train.random_split(0.999)
# Create a model.
model = gl.boosted_trees_regression.create(train, target='Demada_log',
step_size=0.1,
max_iterations=1500,
max_depth = 10,
metric='rmse',
random_seed=998,
column_subsample=0.5,
row_subsample=0.85,
validation_set=None,
model_checkpoint_path=path,
model_checkpoint_interval=1500)
# +
# '''resume_from_checkpoint'''
# train_data, test_data = train.random_split(0.999)
# model = gl.boosted_trees_regression.create(train_data, target='Demada_log',
# step_size=0.1,
# max_iterations=1000,
# max_depth = 10,
# metric='rmse',
# random_seed=461,
# column_subsample=0.75,
# row_subsample=0.85,
# validation_set=test_data,
# resume_from_checkpoint=path+'model_checkpoint_1000_w8',
# model_checkpoint_path=path,
# model_checkpoint_interval=1000)
# -
'''feature important'''
w = model.get_feature_importance()
w = w.add_row_number()
w
# +
from IPython.core.pylabtools import figsize
import numpy as np
from matplotlib import pyplot as plt
import seaborn as sns
sns.set_style('darkgrid', {'grid.color': '.8','grid.linestyle': u'--'})
# %matplotlib inline
figsize(12, 6)
plt.bar(w['id'], w['count'], tick_label=w['name'])
plt.xticks(rotation=45)
# +
from IPython.core.pylabtools import figsize
import numpy as np
from matplotlib import pyplot as plt
import seaborn as sns
sns.set_style('darkgrid', {'grid.color': '.8','grid.linestyle': u'--'})
# %matplotlib inline
figsize(16, 6)
plt.scatter(model.progress['Iteration'], model.progress['Training-rmse'],alpha=.5)
plt.ylim(.4,.5)
# plt.xticks(rotation=45)
# +
# Save predictions to an SArray
# predictions = model.predict(train)
# Evaluate the model and save the results into a dictionary
# results = model.evaluate(train)
# print results
# -
# ## predict
test = gl.SFrame.read_csv(path + 'test_fs_w9.csv', verbose=False)
# test = test.join(town, on=['Agencia_ID','Producto_ID'], how='left')
# del test['Town']
# test = test.fillna('t_c',1)
# test = test.fillna('tcc',0)
# test = test.fillna('tp_sum',0)
# +
del test['Canal_ID']
del test['lag_sum']
del test['prior_sum']
del test['n_t']
del test['prom']
del test['brand']
del test['healthy']
del test['drink']
# -
print test.column_names()
print len(test.column_names())
'''Add feature to week 11'''
def feature_w11(test, lag_sum=0, prior_sum=0):
test_full = test.copy()
ids = test['id']
del test['id']
del test['Semana']
demand_log = model.predict(test)
sub1 = gl.SFrame({'id':ids,'Demanda_uni_equil':demand_log})
test_full = test_full.join(sub1,on=['id'],how='left')
lag11 = test_full.groupby(key_columns=['Semana','Cliente_ID','Producto_ID'], operations={'lag11':agg.MEAN('Demanda_uni_equil')})
lag11['Semana'] = lag11['Semana'].apply(lambda x: x+1)
test_full = test_full.join(lag11,on=['Semana','Cliente_ID','Producto_ID'],how='left')
test_full = test_full.fillna('lag11',0)
test_full['lag1'] = test_full['lag1'] + test_full['lag11']
if lag_sum == 1:
test_full['lag_sum'] = test_full['lag_sum'] + test_full['lag11']
if prior_sum == 1:
lag_sum11 = test_full.groupby(key_columns=['Semana','Cliente_ID','Producto_ID'], operations={'lag_sum11':agg.SUM('Demanda_uni_equil')})
lag_sum11['Semana'] = lag_sum11['Semana'].apply(lambda x: x+1)
test_full = test_full.join(lag_sum11,on=['Semana','Cliente_ID','Producto_ID'],how='left')
test_full = test_full.fillna('lag_sum11',0)
test_full['prior_sum'] = test_full['prior_sum'] + test_full['lag_sum11']
del test_full['lag_sum11']
del test_full['lag11']
del test_full['Demanda_uni_equil']
return test_full
# +
test_full = feature_w11(test, lag_sum=0, prior_sum=0)
ids = test_full['id']
del test_full['id']
del test_full['Semana']
demand_log = model.predict(test_full)
sub = gl.SFrame({'id':ids,'Demanda_uni_equil':demand_log})
# -
import math
sub['Demanda_uni_equil'] = sub['Demanda_uni_equil'].apply(lambda x: math.expm1(max(0, x)))
file_name = 'w9'+'_f'+str(model.num_features)+'_n'+str(model.max_iterations)+'_c'+str(model.column_subsample)
sub.save(path + file_name,format='csv')
sub
| Bimbo/GBRT_w9_fs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install pandas
# !pip install seaborn
# !pip install matplotlib
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
happiness = pd.read_csv("happiness.csv", index_col=1)
happiness.drop("Unnamed: 0", axis="columns", inplace=True)
happiness.sort_values("Overall.rank", ascending=True, inplace=True)
sns.set(font_scale=1)
score_20 = sns.barplot(x=happiness["Country.or.region"][:20], y=happiness["Score"][:20])
score_20.set_xticklabels(score_20.get_xticklabels(), rotation=90)
plt.show()
sns.set(font_scale=.4)
score = sns.barplot(x=happiness["Country.or.region"], y=happiness["Score"])
score.set_xticklabels(score.get_xticklabels(), rotation=90)
plt.show()
gdp = sns.lmplot(x="GDP.per.capita", y="Score", data=happiness)
total = sns.pairplot(happiness)
gdp = sns.lmplot(x="Social.support", y="Score", data=happiness)
gdp = sns.lmplot(x="Healthy.life.expectancy", y="Score", data=happiness)
gdp = sns.lmplot(x="GDP.per.capita", y="Social.support", data=happiness)
gdp = sns.lmplot(x="Social.support", y="Healthy.life.expectancy", data=happiness)
gdp = sns.lmplot(x="GDP.per.capita", y="Healthy.life.expectancy", data=happiness)
# Can do some interesting things with grouping. If I had more time, I would probably add a column that categorized the "Country.or.region" into regions such as "North America, South America, Asia, Europe, etc". They I could play with the hue function and show each as a seperate color on the graphs. This might give some insight into how different parts of the world scored.
| Visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
#
# Signal-space separation (SSS) and Maxwell filtering
# ===================================================
#
# This tutorial covers reducing environmental noise and compensating for head
# movement with SSS and Maxwell filtering.
# :depth: 2
#
# As usual we'll start by importing the modules we need, loading some
# `example data <sample-dataset>`, and cropping it to save on memory:
#
# +
import os
import mne
sample_data_folder = mne.datasets.sample.data_path()
sample_data_raw_file = os.path.join(sample_data_folder, 'MEG', 'sample',
'sample_audvis_raw.fif')
raw = mne.io.read_raw_fif(sample_data_raw_file, verbose=False)
raw.crop(tmax=60).load_data()
# -
# Background on SSS and Maxwell filtering
# ^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
#
# Signal-space separation (SSS) [1]_ [2]_ is a technique based on the physics
# of electromagnetic fields. SSS separates the measured signal into components
# attributable to sources *inside* the measurement volume of the sensor array
# (the *internal components*), and components attributable to sources *outside*
# the measurement volume (the *external components*). The internal and external
# components are linearly independent, so it is possible to simply discard the
# external components to reduce environmental noise. *Maxwell filtering* is a
# related procedure that omits the higher-order components of the internal
# subspace, which are dominated by sensor noise. Typically, Maxwell filtering
# and SSS are performed together (in MNE-Python they are implemented together
# in a single function).
#
# Like `SSP <tut-artifact-ssp>`, SSS is a form of projection. Whereas SSP
# empirically determines a noise subspace based on data (empty-room recordings,
# EOG or ECG activity, etc) and projects the measurements onto a subspace
# orthogonal to the noise, SSS mathematically constructs the external and
# internal subspaces from `spherical harmonics`_ and reconstructs the sensor
# signals using only the internal subspace (i.e., does an oblique projection).
#
# <div class="alert alert-danger"><h4>Warning</h4><p>Maxwell filtering was originally developed for Elekta Neuromag® systems,
# and should be considered *experimental* for non-Neuromag data. See the
# Notes section of the :func:`~mne.preprocessing.maxwell_filter` docstring
# for details.</p></div>
#
# The MNE-Python implementation of SSS / Maxwell filtering currently provides
# the following features:
#
# - Bad channel reconstruction
# - Cross-talk cancellation
# - Fine calibration correction
# - tSSS
# - Coordinate frame translation
# - Regularization of internal components using information theory
# - Raw movement compensation (using head positions estimated by MaxFilter)
# - cHPI subtraction (see :func:`mne.chpi.filter_chpi`)
# - Handling of 3D (in addition to 1D) fine calibration files
# - Epoch-based movement compensation as described in [1]_ through
# :func:`mne.epochs.average_movements`
# - **Experimental** processing of data from (un-compensated) non-Elekta
# systems
#
#
# Using SSS and Maxwell filtering in MNE-Python
# ^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
#
# For optimal use of SSS with data from Elekta Neuromag® systems, you should
# provide the path to the fine calibration file (which encodes site-specific
# information about sensor orientation and calibration) as well as a crosstalk
# compensation file (which reduces interference between Elekta's co-located
# magnetometer and paired gradiometer sensor units).
#
#
fine_cal_file = os.path.join(sample_data_folder, 'SSS', 'sss_cal_mgh.dat')
crosstalk_file = os.path.join(sample_data_folder, 'SSS', 'ct_sparse_mgh.fif')
# Before we perform SSS we'll set a couple additional bad channels — ``MEG
# 2313`` has some DC jumps and ``MEG 1032`` has some large-ish low-frequency
# drifts. After that, performing SSS and Maxwell filtering is done with a
# single call to :func:`~mne.preprocessing.maxwell_filter`, with the crosstalk
# and fine calibration filenames provided (if available):
#
#
raw.info['bads'].extend(['MEG 1032', 'MEG 2313'])
raw_sss = mne.preprocessing.maxwell_filter(raw, cross_talk=crosstalk_file,
calibration=fine_cal_file)
# <div class="alert alert-danger"><h4>Warning</h4><p>Automatic bad channel detection is not currently implemented. It is
# critical to mark bad channels in ``raw.info['bads']`` *before* calling
# :func:`~mne.preprocessing.maxwell_filter` in order to prevent bad
# channel noise from spreading.</p></div>
#
# To see the effect, we can plot the data before and after SSS / Maxwell
# filtering.
#
#
raw.pick(['meg']).plot(duration=2, butterfly=True)
raw_sss.pick(['meg']).plot(duration=2, butterfly=True)
# Notice that channels marked as "bad" have been effectively repaired by SSS,
# eliminating the need to perform `interpolation <tut-bad-channels>`.
# The heartbeat artifact has also been substantially reduced.
#
# The :func:`~mne.preprocessing.maxwell_filter` function has parameters
# ``int_order`` and ``ext_order`` for setting the order of the spherical
# harmonic expansion of the interior and exterior components; the default
# values are appropriate for most use cases. Additional parameters include
# ``coord_frame`` and ``origin`` for controlling the coordinate frame ("head"
# or "meg") and the origin of the sphere; the defaults are appropriate for most
# studies that include digitization of the scalp surface / electrodes. See the
# documentation of :func:`~mne.preprocessing.maxwell_filter` for details.
#
#
# Spatiotemporal SSS (tSSS)
# ^^^^^^^^^^^^^^^^^^^^^^^^^
#
# An assumption of SSS is that the measurement volume (the spherical shell
# where the sensors are physically located) is free of electromagnetic sources.
# The thickness of this source-free measurement shell should be 4-8 cm for SSS
# to perform optimally. In practice, there may be sources falling within that
# measurement volume; these can often be mitigated by using Spatiotemporal
# Signal Space Separation (tSSS) [2]_. tSSS works by looking for temporal
# correlation between components of the internal and external subspaces, and
# projecting out any components that are common to the internal and external
# subspaces. The projection is done in an analogous way to
# `SSP <tut-artifact-ssp>`, except that the noise vector is computed
# across time points instead of across sensors.
#
# To use tSSS in MNE-Python, pass a time (in seconds) to the parameter
# ``st_duration`` of :func:`~mne.preprocessing.maxwell_filter`. This will
# determine the "chunk duration" over which to compute the temporal projection.
# The chunk duration effectively acts as a high-pass filter with a cutoff
# frequency of $\frac{1}{\mathtt{st\_duration}}~\mathrm{Hz}$; this
# effective high-pass has an important consequence:
#
# - In general, larger values of ``st_duration`` are better (provided that your
# computer has sufficient memory) because larger values of ``st_duration``
# will have a smaller effect on the signal.
#
# If the chunk duration does not evenly divide your data length, the final
# (shorter) chunk will be added to the prior chunk before filtering, leading
# to slightly different effective filtering for the combined chunk (the
# effective cutoff frequency differing at most by a factor of 2). If you need
# to ensure identical processing of all analyzed chunks, either:
#
# - choose a chunk duration that evenly divides your data length (only
# recommended if analyzing a single subject or run), or
#
# - include at least ``2 * st_duration`` of post-experiment recording time at
# the end of the :class:`~mne.io.Raw` object, so that the data you intend to
# further analyze is guaranteed not to be in the final or penultimate chunks.
#
# Additional parameters affecting tSSS include ``st_correlation`` (to set the
# correlation value above which correlated internal and external components
# will be projected out) and ``st_only`` (to apply only the temporal projection
# without also performing SSS and Maxwell filtering). See the docstring of
# :func:`~mne.preprocessing.maxwell_filter` for details.
#
#
# Movement compensation
# ^^^^^^^^^^^^^^^^^^^^^
#
# If you have information about subject head position relative to the sensors
# (i.e., continuous head position indicator coils, or :term:`cHPI <hpi>`), SSS
# can take that into account when projecting sensor data onto the internal
# subspace. Head position data is loaded with the
# :func:`~mne.chpi.read_head_pos` function. The `example data
# <sample-dataset>` doesn't include cHPI, so here we'll load a :file:`.pos`
# file used for testing, just to demonstrate:
#
#
head_pos_file = os.path.join(mne.datasets.testing.data_path(), 'SSS',
'test_move_anon_raw.pos')
head_pos = mne.chpi.read_head_pos(head_pos_file)
mne.viz.plot_head_positions(head_pos, mode='traces')
# The cHPI data file could also be passed as the ``head_pos`` parameter of
# :func:`~mne.preprocessing.maxwell_filter`. Not only would this account for
# movement within a given recording session, but also would effectively
# normalize head position across different measurement sessions and subjects.
# See `here <example-movement-comp>` for an extended example of applying
# movement compensation during Maxwell filtering / SSS. Another option is to
# apply movement compensation when averaging epochs into an
# :class:`~mne.Evoked` instance, using the :func:`mne.epochs.average_movements`
# function.
#
# Each of these approaches requires time-varying estimates of head position,
# which is obtained from MaxFilter using the ``-headpos`` and ``-hp``
# arguments (see the MaxFilter manual for details).
#
#
# Caveats to using SSS / Maxwell filtering
# ^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
#
# 1. There are patents related to the Maxwell filtering algorithm, which may
# legally preclude using it in commercial applications. More details are
# provided in the documentation of
# :func:`~mne.preprocessing.maxwell_filter`.
#
# 2. SSS works best when both magnetometers and gradiometers are present, and
# is most effective when gradiometers are planar (due to the need for very
# accurate sensor geometry and fine calibration information). Thus its
# performance is dependent on the MEG system used to collect the data.
#
#
# References
# ^^^^^^^^^^
#
# .. [1] <NAME> and <NAME>. (2005). Presentation of electromagnetic
# multichannel data: The signal space separation method. *J Appl Phys*
# 97, 124905 1-10. https://doi.org/10.1063/1.1935742
#
# .. [2] <NAME> and <NAME>. (2006). Spatiotemporal signal space separation
# method for rejecting nearby interference in MEG measurements. *Phys
# Med Biol* 51, 1759-1768. https://doi.org/10.1088/0031-9155/51/7/008
#
#
# .. LINKS
#
#
#
| dev/_downloads/243172b1ef6a2d804d3245b8c0a927ef/plot_60_maxwell_filtering_sss.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bayesian Optimization
#
# [Bayesian optimization](https://en.wikipedia.org/wiki/Bayesian_optimization) is a powerful strategy for minimizing (or maximizing) objective functions that are costly to evaluate. It is an important component of [automated machine learning](https://en.wikipedia.org/wiki/Automated_machine_learning) toolboxes such as [auto-sklearn](https://automl.github.io/auto-sklearn/stable/), [auto-weka](http://www.cs.ubc.ca/labs/beta/Projects/autoweka/), and [scikit-optimize](https://scikit-optimize.github.io/), where Bayesian optimization is used to select model hyperparameters. Bayesian optimization is used for a wide range of other applications as well; as cataloged in the review [2], these include interactive user-interfaces, robotics, environmental monitoring, information extraction, combinatorial optimization, sensor networks, adaptive Monte Carlo, experimental design, and reinforcement learning.
#
# ## Problem Setup
#
# We are given a minimization problem
#
# $$ x^* = \text{arg}\min \ f(x), $$
#
# where $f$ is a fixed objective function that we can evaluate pointwise.
# Here we assume that we do _not_ have access to the gradient of $f$. We also
# allow for the possibility that evaluations of $f$ are noisy.
#
# To solve the minimization problem, we will construct a sequence of points $\{x_n\}$ that converge to $x^*$. Since we implicitly assume that we have a fixed budget (say 100 evaluations), we do not expect to find the exact minumum $x^*$: the goal is to get the best approximate solution we can given the allocated budget.
#
# The Bayesian optimization strategy works as follows:
#
# 1. Place a prior on the objective function $f$. Each time we evaluate $f$ at a new point $x_n$, we update our model for $f(x)$. This model serves as a surrogate objective function and reflects our beliefs about $f$ (in particular it reflects our beliefs about where we expect $f(x)$ to be close to $f(x^*)$). Since we are being Bayesian, our beliefs are encoded in a posterior that allows us to systematically reason about the uncertainty of our model predictions.
#
# 2. Use the posterior to derive an "acquisition" function $\alpha(x)$ that is easy to evaluate and differentiate (so that optimizing $\alpha(x)$ is easy). In contrast to $f(x)$, we will generally evaluate $\alpha(x)$ at many points $x$, since doing so will be cheap.
#
# 3. Repeat until convergence:
#
# + Use the acquisition function to derive the next query point according to
# $$ x_{n+1} = \text{arg}\min \ \alpha(x). $$
#
# + Evaluate $f(x_{n+1})$ and update the posterior.
#
# A good acquisition function should make use of the uncertainty encoded in the posterior to encourage a balance between exploration—querying points where we know little about $f$—and exploitation—querying points in regions we have good reason to think $x^*$ may lie. As the iterative procedure progresses our model for $f$ evolves and so does the acquisition function. If our model is good and we've chosen a reasonable acquisition function, we expect that the acquisition function will guide the query points $x_n$ towards $x^*$.
#
# In this tutorial, our model for $f$ will be a Gaussian process. In particular we will see how to use the [Gaussian Process module](http://docs.pyro.ai/en/0.3.1/contrib.gp.html) in Pyro to implement a simple Bayesian optimization procedure.
# +
import matplotlib.gridspec as gridspec
import matplotlib.pyplot as plt
import torch
import torch.autograd as autograd
import torch.optim as optim
from torch.distributions import constraints, transform_to
import pyro
import pyro.contrib.gp as gp
assert pyro.__version__.startswith('0.3.1')
pyro.enable_validation(True) # can help with debugging
pyro.set_rng_seed(1)
# -
# ## Define an objective function
#
# For the purposes of demonstration, the objective function we are going to consider is the [Forrester et al. (2008) function](https://www.sfu.ca/~ssurjano/forretal08.html):
#
# $$f(x) = (6x-2)^2 \sin(12x-4), \quad x\in [0, 1].$$
#
# This function has both a local minimum and a global minimum. The global minimum is at $x^* = 0.75725$.
def f(x):
return (6 * x - 2)**2 * torch.sin(12 * x - 4)
# Let's begin by plotting $f$.
x = torch.linspace(0, 1)
plt.figure(figsize=(8, 4))
plt.plot(x.numpy(), f(x).numpy())
plt.show()
# ## Setting a Gaussian Process prior
# [Gaussian processes](https://en.wikipedia.org/wiki/Gaussian_process) are a popular choice for a function priors due to their power and flexibility. The core of a Gaussian Process is its covariance function $k$, which governs the similarity of $f(x)$ for pairs of input points. Here we will use a Gaussian Process as our prior for the objective function $f$. Given inputs $X$ and the corresponding noisy observations $y$, the model takes the form
#
# $$f\sim\mathrm{MultivariateNormal}(0,k(X,X)),$$
#
# $$y\sim f+\epsilon,$$
#
# where $\epsilon$ is i.i.d. Gaussian noise and $k(X,X)$ is a covariance matrix whose entries are given by $k(x,x^\prime)$ for each pair of inputs $(x,x^\prime)$.
#
# We choose the [Matern](https://en.wikipedia.org/wiki/Mat%C3%A9rn_covariance_function) kernel with $\nu = \frac{5}{2}$ (as suggested in reference [1]). Note that the popular [RBF](https://en.wikipedia.org/wiki/Radial_basis_function_kernel) kernel, which is used in many regression tasks, results in a function prior whose samples are infinitely differentiable; this is probably an unrealistic assumption for most 'black-box' objective functions.
# initialize the model with four input points: 0.0, 0.33, 0.66, 1.0
X = torch.tensor([0.0, 0.33, 0.66, 1.0])
y = f(X)
gpmodel = gp.models.GPRegression(X, y, gp.kernels.Matern52(input_dim=1),
noise=torch.tensor(0.1), jitter=1.0e-4)
# The following helper function `update_posterior` will take care of updating our `gpmodel` each time we evaluate $f$ at a new value $x$.
def update_posterior(x_new):
y = f(x_new) # evaluate f at new point.
X = torch.cat([gpmodel.X, x_new]) # incorporate new evaluation
y = torch.cat([gpmodel.y, y])
gpmodel.set_data(X, y)
# optimize the GP hyperparameters using Adam with lr=0.001
optimizer = torch.optim.Adam(gpmodel.parameters(), lr=0.001)
gp.util.train(gpmodel, optimizer)
# ## Define an acquisition function
# There are many reasonable options for the acquisition function (see references [1] and [2] for a list of popular choices and a discussion of their properties). Here we will use one that is 'simple to implement and interpret,' namely the 'Lower Confidence Bound' acquisition function.
# It is given by
#
# $$
# \alpha(x) = \mu(x) - \kappa \sigma(x)
# $$
#
# where $\mu(x)$ and $\sigma(x)$ are the mean and square root variance of the posterior at the point $x$, and the arbitrary constant $\kappa>0$ controls the trade-off between exploitation and exploration. This acquisition function will be minimized for choices of $x$ where either: i) $\mu(x)$ is small (exploitation); or ii) where $\sigma(x)$ is large (exploration). A large value of $\kappa$ means that we place more weight on exploration because we prefer candidates $x$ in areas of high uncertainty. A small value of $\kappa$ encourages exploitation because we prefer candidates $x$ that minimize $\mu(x)$, which is the mean of our surrogate objective function. We will use $\kappa=2$.
def lower_confidence_bound(x, kappa=2):
mu, variance = gpmodel(x, full_cov=False, noiseless=False)
sigma = variance.sqrt()
return mu - kappa * sigma
# The final component we need is a way to find (approximate) minimizing points $x_{\rm min}$ of the acquisition function. There are several ways to proceed, including gradient-based and non-gradient-based techniques. Here we will follow the gradient-based approach. One of the possible drawbacks of gradient descent methods is that the minimization algorithm can get stuck at a local minimum. In this tutorial, we adopt a (very) simple approach to address this issue:
#
# - First, we seed our minimization algorithm with 5 different values: i) one is chosen to be $x_{n-1}$, i.e. the candidate $x$ used in the previous step; and ii) four are chosen uniformly at random from the domain of the objective function.
# - We then run the minimization algorithm to approximate convergence for each seed value.
# - Finally, from the five candidate $x$s identified by the minimization algorithm, we select the one that minimizes the acquisition function.
#
# Please refer to reference [2] for a more detailed discussion of this problem in Bayesian Optimization.
def find_a_candidate(x_init, lower_bound=0, upper_bound=1):
# transform x to an unconstrained domain
constraint = constraints.interval(lower_bound, upper_bound)
unconstrained_x_init = transform_to(constraint).inv(x_init)
unconstrained_x = unconstrained_x_init.clone().detach().requires_grad_(True)
minimizer = optim.LBFGS([unconstrained_x])
def closure():
minimizer.zero_grad()
x = transform_to(constraint)(unconstrained_x)
y = lower_confidence_bound(x)
autograd.backward(unconstrained_x, autograd.grad(y, unconstrained_x))
return y
minimizer.step(closure)
# after finding a candidate in the unconstrained domain,
# convert it back to original domain.
x = transform_to(constraint)(unconstrained_x)
return x.detach()
# ## The inner loop of Bayesian Optimization
#
# With the various helper functions defined above, we can now encapsulate the main logic of a single step of Bayesian Optimization in the function `next_x`:
def next_x(lower_bound=0, upper_bound=1, num_candidates=5):
candidates = []
values = []
x_init = gpmodel.X[-1:]
for i in range(num_candidates):
x = find_a_candidate(x_init, lower_bound, upper_bound)
y = lower_confidence_bound(x)
candidates.append(x)
values.append(y)
x_init = x.new_empty(1).uniform_(lower_bound, upper_bound)
argmin = torch.min(torch.cat(values), dim=0)[1].item()
return candidates[argmin]
# ## Running the algorithm
# To illustrate how Bayesian Optimization works, we make a convenient plotting function that will help us visualize our algorithm's progress.
def plot(gs, xmin, xlabel=None, with_title=True):
xlabel = "xmin" if xlabel is None else "x{}".format(xlabel)
Xnew = torch.linspace(-0.1, 1.1)
ax1 = plt.subplot(gs[0])
ax1.plot(gpmodel.X.numpy(), gpmodel.y.numpy(), "kx") # plot all observed data
with torch.no_grad():
loc, var = gpmodel(Xnew, full_cov=False, noiseless=False)
sd = var.sqrt()
ax1.plot(Xnew.numpy(), loc.numpy(), "r", lw=2) # plot predictive mean
ax1.fill_between(Xnew.numpy(), loc.numpy() - 2*sd.numpy(), loc.numpy() + 2*sd.numpy(),
color="C0", alpha=0.3) # plot uncertainty intervals
ax1.set_xlim(-0.1, 1.1)
ax1.set_title("Find {}".format(xlabel))
if with_title:
ax1.set_ylabel("Gaussian Process Regression")
ax2 = plt.subplot(gs[1])
with torch.no_grad():
# plot the acquisition function
ax2.plot(Xnew.numpy(), lower_confidence_bound(Xnew).numpy())
# plot the new candidate point
ax2.plot(xmin.numpy(), lower_confidence_bound(xmin).numpy(), "^", markersize=10,
label="{} = {:.5f}".format(xlabel, xmin.item()))
ax2.set_xlim(-0.1, 1.1)
if with_title:
ax2.set_ylabel("Acquisition Function")
ax2.legend(loc=1)
# Our surrogate model `gpmodel` already has 4 function evaluations at its disposal; however, we have yet to optimize the GP hyperparameters. So we do that first. Then in a loop we call the `next_x` and `update_posterior` functions repeatedly. The following plot illustrates how Gaussian Process posteriors and the corresponding acquisition functions change at each step in the algorith. Note how query points are chosen both for exploration and exploitation.
plt.figure(figsize=(12, 30))
outer_gs = gridspec.GridSpec(5, 2)
optimizer = torch.optim.Adam(gpmodel.parameters(), lr=0.001)
gp.util.train(gpmodel, optimizer)
for i in range(8):
xmin = next_x()
gs = gridspec.GridSpecFromSubplotSpec(2, 1, subplot_spec=outer_gs[i])
plot(gs, xmin, xlabel=i+1, with_title=(i % 2 == 0))
update_posterior(xmin)
plt.show()
# Because we have assumed that our observations contain noise, it is improbable that we will find the exact minimizer of the function $f$. Still, with a relatively small budget of evaluations (12) we see that the algorithm has converged to very close to the global minimum at $x^* = 0.75725$.
#
# While this tutorial is only intended to be a brief introduction to Bayesian Optimization, we hope that we have been able to convey the basic underlying ideas. Consider watching the lecture by Nando de Freitas [3] for an excellent exposition of the basic theory. Finally, the reference paper [2] gives a review of recent research on Bayesian Optimization, together with many discussions about important technical details.
# ## References
#
# [1] `Practical bayesian optimization of machine learning algorithms`,<br />
# <NAME>, <NAME>, and <NAME>
#
# [2] `Taking the human out of the loop: A review of bayesian optimization`,<br />
# <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>
#
# [3] [Machine learning - Bayesian optimization and multi-armed bandits](https://www.youtube.com/watch?v=vz3D36VXefI)
| tutorial/source/bo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from keras.models import Sequential
from keras.layers import Dense, Activation
from keras.datasets import boston_housing
(X_train, Y_train), (X_test, Y_test) = boston_housing.load_data()
# +
nFeatures = X_train.shape[1]
model = Sequential()
model.add(Dense(1, input_shape=(nFeatures,), kernel_initializer='uniform'))
model.add(Activation('linear'))
model.compile(optimizer='rmsprop', loss='mse', metrics=['mse', 'mae'])
model.fit(X_train, Y_train, batch_size=4, epochs=1000)
# +
model.summary()
model.evaluate(X_test, Y_test, verbose=True)
# -
Y_pred = model.predict(X_test)
print Y_test[:5]
print Y_pred[:5,0]
| BookSrc/learnOpenCV/keras-linear-regression/intro-to-keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Finite Elements Lab 2 Worksheet
from IPython.core.display import HTML
css_file = 'https://raw.githubusercontent.com/ngcm/training-public/master/ipython_notebook_styles/ngcmstyle.css'
HTML(url=css_file)
# ## 2d problem
# We've looked at the problem of finding the static temperature distribution in a bar. Now let's move on to finding the temperature distribution of a plate of length $1$ on each side. The temperature $T(x, y) = T(x_1, x_2)$ satisfies
#
# $$
# \nabla^2 T + f({\bf x}) = \left( \partial_{xx} + \partial_{yy} \right) T + f({\bf x}) = 0.
# $$
#
# We'll fix the temperature to be zero at the right edge, $T(1, y) = 0$. We'll allow heat to flow out of the other edges, giving the boundary conditions on all edges as
#
# $$
# \begin{align}
# \partial_x T(0, y) &= 0, & T(1, y) &=0, \\
# \partial_y T(x, 0) &= 0, & \partial_y T(x, 1) &=0.
# \end{align}
# $$
# Once again we want to write down the weak form by integrating by parts. To do that we rely on the divergence theorem,
#
# $$
# \int_{\Omega} \text{d}\Omega \, \nabla_i \phi = \int_{\Gamma} \text{d}\Gamma \, \phi n_i.
# $$
#
# Here $\Omega$ is the domain (which in our problem is the plate, $x, y \in [0, 1]$) and $\Gamma$ its boundary (in our problem the four lines $x=0, 1$ and $y=0, 1$), whilst ${\bf n}$ is the (inward-pointing) normal vector to the boundary.
#
# We then multiply the strong form of the static heat equation by a *weight function* $w(x, y)$ and integrate by parts, using the divergence theorem, to remove the second derivative. To enforce the boundary conditions effectively we again choose the weight function to vanish where the value of the temperature is explicitly given, i.e. $w(1, y) = 0$. That is, we split the boundary $\Gamma$ into a piece $\Gamma_D$ where the boundary conditions are in Dirichlet form (the value $T$ is given) and a piece $\Gamma_N$ where the boundary conditions are in Neumann form (the value of the normal derivative $n_i \nabla_i T$ is given). We then enforce that on $\Gamma_D$ the weight function vanishes.
#
# For our problem, this gives
#
# $$
# \int_{\Omega} \text{d} \Omega \, \nabla_i w \nabla_i T = \int_{\Omega} \text{d} \Omega \, w f.
# $$
#
# Re-writing for our explicit domain and our Cartesian coordinates we get
#
# $$
# \int_0^1 \text{d} y \, \int_0^1 \text{d} x \, \left( \partial_x w \partial_x T + \partial_y w \partial_y T \right) = \int_0^1 \text{d} y \, \int_0^1 \text{d} x \, w(x, y) f(x, y).
# $$
#
# This should be compared to the one dimensional case
#
# $$
# \int_0^1 \text{d}x \, \partial_x w(x) \partial_x T(x) = \int_0^1 \text{d}x \, w(x) f(x).
# $$
# We can now envisage using the same steps as the one dimensional case. Split the domain into elements, represent all functions in terms of known *shape functions* on each element, assemble the problems in each element to a single matrix problem, and then solve the matrix problem.
# ### Elements
# Here we will use triangular elements. As a simple example we'll split the plate into two triangles.
# %matplotlib inline
import numpy
from matplotlib import pyplot
from matplotlib import rcParams
rcParams['font.family'] = 'serif'
rcParams['font.size'] = 16
rcParams['figure.figsize'] = (12,6)
nodes = numpy.array([[0.0, 0.0], [1.0, 0.0], [0.0, 1.0], [1.0, 1.0]])
IEN = numpy.array([[0, 1, 2],
[1, 3, 2]])
pyplot.figure()
pyplot.axis('equal')
pyplot.triplot(nodes[:,0], nodes[:,1], triangles=IEN, lw=2)
pyplot.plot(nodes[:,0], nodes[:,1], 'ro')
for e in range(nodes.shape[1]):
barycentre = numpy.mean(nodes[IEN[e,:],:], axis=0)
pyplot.text(barycentre[0], barycentre[1], "{}".format(e),
bbox=dict(facecolor='red', alpha=0.5))
for n in range(3):
pyplot.text(nodes[IEN[e,n],0]-0.07*(-1)**e,nodes[IEN[e,n],1]+0.07, r"${}_{{{}}}$".format(n,e),
bbox=dict(facecolor='blue', alpha=0.25 + 0.5*e))
for n in range(nodes.shape[0]):
pyplot.text(nodes[n,0]-0.07, nodes[n,1]-0.07, "{}".format(n),
bbox=dict(facecolor='green', alpha=0.3))
pyplot.xlim(-0.2, 1.2)
pyplot.ylim(-0.2, 1.2)
pyplot.xlabel(r"$x$")
pyplot.ylabel(r"$y$");
# What we're doing here is
#
# 1. Providing a list of nodes by their global coordinates.
# 2. Providing the *element node array* `IEN` which says how the elements are linked to the nodes.
#
# We have that for element $e$ and *local* node number $a = 0, 1, 2$ the global node number is $A = IEN(e, a)$. This notation is sufficiently conventional that `matplotlib` recognizes it with its `triplot`/`tripcolor`/`trisurf` functions.
#
# It is convention that the nodes are ordered in the anti-clockwise direction as the local number goes from $0$ to $2$.
#
# The plot shows the
#
# * element numbers in the red boxes
# * the *global* node numbers in the green boxes
# * the *local* element numbers in the blue boxes (the subscript shows the element number).
# We will need one final array, which is the $ID$ or *destination* array. This links the *global* node number to the *global* equation number in the final linear system. As the order of the equations in a linear system doesn't matter, this essentially encodes whether a node should have any equation in the linear system. Any node on $\Gamma_D$, where the value of the temperature is given, should not have an equation. In the example above the right edge is fixed, so nodes $1$ and $3$ lie on $\Gamma_D$ and should not have an equation. Thus in our case we have
ID = numpy.array([0,-1,1,-1])
# In the one dimensional case we used the *location matrix* or $LM$ array to link local node numbers in elements to equations. With the $IED$ and $ID$ arrays the $LM$ matrix is strictly redundant, as $LM(a, e) = ID(IEN(e, a))$. However, it's still standard to construct it:
LM = numpy.zeros_like(IEN.T)
for e in range(IEN.shape[0]):
for a in range(IEN.shape[1]):
LM[a,e] = ID[IEN[e,a]]
LM
# ### Function representation and shape functions
# We're going to want to write our unknown functions $T, w$ in terms of shape functions. These are easiest to write down for a single reference element, in the same way as we did for the one dimensional case where our reference element used the coordinates $\xi$. In two dimensions we'll use the reference coordinates $\xi_0, \xi_1$, and the standard "unit" triangle:
corners = numpy.array([[0.0, 0.0], [1.0, 0.0], [0.0, 1.0], [0.0, 0.0]])
pyplot.plot(corners[:,0],corners[:,1],linewidth=2)
pyplot.xlabel(r"$\xi_0$")
pyplot.ylabel(r"$\xi_1$")
pyplot.axis('equal')
pyplot.ylim(-0.1,1.1);
# The shape functions on this triangle are
#
# \begin{align}
# N_0(\xi_0, \xi_1) &= 1 - \xi_0 - \xi_1, \\
# N_1(\xi_0, \xi_1) &= \xi_0, \\
# N_2(\xi_0, \xi_1) &= \xi_1.
# \end{align}
#
# The derivatives are all either $0$ or $\pm 1$.
# As soon as we have the shape functions, our weak form becomes
#
# $$
# \sum_A T_A \int_{\Omega} \text{d}\Omega \, \left( \partial_{x} N_A (x, y) \partial_{x} N_B(x, y) + \partial_{y} N_A(x, y) \partial_{y} N_B(x, y) \right) = \int_{\Omega} \text{d}\Omega \, N_B(x, y) f(x, y).
# $$
#
# If we restrict to a single element the weak form becomes
#
# $$
# \sum_A T_A \int_{\triangle} \text{d}\triangle \, \left( \partial_{x} N_A (x, y) \partial_{x} N_B(x, y) + \partial_{y} N_A(x, y) \partial_{y} N_B(x, y) \right) = \int_{\triangle} \text{d}\triangle \, N_B(x, y) f(x, y).
# $$
# We need to map the triangle and its $(x, y) = {\bf x}$ coordinates to the reference triangle and its $(\xi_0, \xi_1) = {\bf \xi}$ coordinates. We also need to work out the integrals that appear in the weak form. We need the transformation formula
#
# $$
# \int_{\triangle} \text{d}\triangle \, \phi(x, y) = \int_0^1 \text{d}\xi_1 \, \int_0^{1-\xi_1} \text{d}\xi_0 \, \phi \left( x(\xi_0, \xi_1), y(\xi_0, \xi_1) \right) j(\xi_0, \xi_1),
# $$
#
# where the *Jacobian matrix* $J$ is
#
# $$
# J = \left[ \frac{\partial {\bf x}}{\partial {\bf \xi}} \right] = \begin{pmatrix} \partial_{\xi_0} x & \partial_{\xi_1} x \\ \partial_{\xi_0} y & \partial_{\xi_1} y \end{pmatrix}
# $$
#
# and hence the *Jacobian determinant* $j$ is
#
# $$
# j = \det{J} = \det \left[ \frac{\partial {\bf x}}{\partial {\bf \xi}} \right] = \det \begin{pmatrix} \partial_{\xi_0} x & \partial_{\xi_1} x \\ \partial_{\xi_0} y & \partial_{\xi_1} y \end{pmatrix}.
# $$
#
# We will also need the Jacobian matrix when writing the derivatives of the shape functions in terms of the coordinates on the reference triangle, i.e.
#
# $$
# \begin{pmatrix} \partial_x N_A & \partial_y N_A \end{pmatrix} = \begin{pmatrix} \partial_{\xi_0} N_A & \partial_{\xi_1} N_A \end{pmatrix} J^{-1} .
# $$
#
# The integral over the reference triangle can be directly approximated using, for example, Gauss quadrature. To second order we have
#
# $$
# \int_0^1 \text{d}\xi_1 \, \int_0^{1-\xi_1} \text{d}\xi_0 \, \psi \left( x(\xi_0, \xi_1), y(\xi_0, \xi_1) \right) \simeq \frac{1}{6} \sum_{j = 1}^{3} \psi \left( x((\xi_0)_j, (\xi_1)_j), y((\xi_0)_j, (\xi_1)_j) \right)
# $$
#
# where
#
# $$
# \begin{align}
# (\xi_0)_1 &= \frac{1}{6}, & (\xi_1)_1 &= \frac{1}{6}, \\
# (\xi_0)_2 &= \frac{4}{6}, & (\xi_1)_2 &= \frac{1}{6}, \\
# (\xi_0)_3 &= \frac{1}{6}, & (\xi_1)_3 &= \frac{4}{6}.
# \end{align}
# $$
#
# Finally, we need to map from the coordinates ${\bf \xi}$ to the coordinates ${\bf x}$. This is straightforward if we think of writing each component $(x, y)$ in terms of the shape functions. So for element $e$ with node locations $(x^e_a, y^e_a)$ for local node number $a = 0, 1, 2$ we have
#
# $$
# \begin{align}
# x &= x^e_0 N_0(\xi_0, \xi_1) + x^e_1 N_1(\xi_0, \xi_1) + x^e_2 N_2(\xi_0, \xi_1), \\
# y &= y^e_0 N_0(\xi_0, \xi_1) + y^e_1 N_1(\xi_0, \xi_1) + y^e_2 N_2(\xi_0, \xi_1).
# \end{align}
# $$
# ### Tasks
# 1. Write a function that, given ${\bf \xi}$, returns that shape functions at that location.
# 2. Write a function that, given ${\bf \xi}$, returns the derivatives of the shape functions at that location.
# 3. Write a function that, given the (global) locations ${\bf x}$ of the nodes of a triangular element and the local coordinates ${\bf \xi}$ within the element returns the corresponding global coordinates.
# 5. Write a function that, given the (global) locations ${\bf x}$ of the nodes of a triangular element and the local coordinates ${\bf \xi}$, returns the Jacobian matrix at that location.
# 6. Write a function that, given the (global) locations ${\bf x}$ of the nodes of a triangular element and the local coordinates ${\bf \xi}$, returns the determinant of the Jacobian matrix at that location.
# 4. Write a function that, given the (global) locations ${\bf x}$ of the nodes of a triangular element and the local coordinates ${\bf \xi}$ within the element returns the derivatives $\partial_{\bf x} N_a = J^{-1} \partial_{\bf \xi} N_a$.
# 7. Write a function that, given a function $\psi({\bf \xi})$, returns the quadrature of $\psi$ over the reference triangle.
# 8. Write a function that, given the (global) locations of the nodes of a triangular element and a function $\phi(x, y)$, returns the quadrature of $\phi$ over the element.
# 9. Test all of the above by integrating simple functions (eg $1, \xi, \eta, x, y$) over the elements above.
# ### More tasks
# 1. Write a function to compute the coefficients of the stiffness matrix for a single element,
# $$
# k^e_{ab} = \int_{\triangle^e} \text{d}\triangle^e \, \left( \partial_{x} N_a (x, y) \partial_{x} N_b(x, y) + \partial_{y} N_a(x, y) \partial_{y} N_b(x, y) \right).
# $$
# 2. Write a function to compute the coefficients of the force vector for a single element,
# $$
# f^e_b = \int_{\triangle^e} \text{d}\triangle^e \, N_b(x, y) f(x, y).
# $$
# ### Algorithm
# This gives our full algorithm:
#
# 1. Set number of elements $N_{\text{elements}}$.
# 2. Set node locations ${\bf x}_A, A = 0, \dots, N_{\text{nodes}}$. Note that there is no longer a direct connection between the number of nodes and elements.
# 3. Set up the $IEN$ and $ID$ arrays linking elements to nodes and elements to equation numbers. From these set the location matrix $LM$. Work out the required number of equations $N_{\text{equations}}$ (the maximum of the $ID$ array plus $1$).
# 4. Set up arrays of zeros for the global stiffness matrix (size $N_{\text{equations}} \times N_{\text{equations}}$) and force vector (size $N_{\text{equations}}$).
# 5. For each element:
#
# 1. Form the element stiffness matrix $k^e_{ab}$.
# 2. Form the element force vector $f^e_a$.
# 3. Add the contributions to the global stiffness matrix and force vector
#
# 6. Solve $K {\bf T} = {\bf F}$.
# ### Algorithm tasks
# 1. Write a function that given a list of nodes and the $IEN$ and $ID$ arrays and returns the solution ${\bf T}$.
# 2. Test on the system $f(x, y) = 1$ with exact solution $T = (1-x^2)/2$.
# 3. For a more complex case with the same boundary conditions try
# $$
# f(x, y) = x^2 (x - 1) \left( y^2 + 4 y (y - 1) + (y - 1)^2 \right) + (3 x - 1) y^2 (y - 1)^2
# $$
# with exact solution
# $$
# T(x, y) = \tfrac{1}{2} x^2 (1 - x) y^2 (1 - y)^2.
# $$
# A useful function is a grid generator or mesher. Good meshers are generally hard: here is a very simple one for this specific problem.
def generate_2d_grid(Nx):
"""
Generate a triangular grid covering the plate math:`[0,1]^2` with Nx (pairs of) triangles in each dimension.
Parameters
----------
Nx : int
Number of triangles in any one dimension (so the total number on the plate is math:`2 Nx^2`)
Returns
-------
nodes : array of float
Array of (x,y) coordinates of nodes
IEN : array of int
Array linking elements to nodes
ID : array of int
Array linking nodes to equations
"""
Nnodes = Nx+1
x = numpy.linspace(0, 1, Nnodes)
y = numpy.linspace(0, 1, Nnodes)
X, Y = numpy.meshgrid(x,y)
nodes = numpy.zeros((Nnodes**2,2))
nodes[:,0] = X.ravel()
nodes[:,1] = Y.ravel()
ID = numpy.zeros(len(nodes), dtype=numpy.int)
n_eq = 0
for nID in range(len(nodes)):
if nID % Nnodes == Nx:
ID[nID] = -1
else:
ID[nID] = n_eq
n_eq += 1
IEN = numpy.zeros((2*Nx**2,3), dtype=numpy.int)
for i in range(Nx):
for j in range(Nx):
IEN[2*i+2*j*Nx , :] = i+j*Nnodes, i+1+j*Nnodes, i+(j+1)*Nnodes
IEN[2*i+1+2*j*Nx, :] = i+1+j*Nnodes, i+1+(j+1)*Nnodes, i+(j+1)*Nnodes
return nodes, IEN, ID
| FEEG6016 Simulation and Modelling/08-Finite-Elements-Lab-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Find the Repos Available in your Database, and What Repository Groups They Are In
# ## Connect to your database
# +
import psycopg2
import pandas as pd
import sqlalchemy as salc
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import warnings
import datetime
import json
warnings.filterwarnings('ignore')
with open("config.json") as config_file:
config = json.load(config_file)
database_connection_string = 'postgres+psycopg2://{}:{}@{}:{}/{}'.format(config['user'], config['password'], config['host'], config['port'], config['database'])
dbschema='augur_data'
engine = salc.create_engine(
database_connection_string,
connect_args={'options': '-csearch_path={}'.format(dbschema)})
# -
# ### Retrieve Available Respositories
# +
repolist = pd.DataFrame()
repo_query = salc.sql.text(f"""
SELECT a.rg_name,
a.repo_group_id,
b.repo_name,
b.repo_id,
b.forked_from,
b.repo_archived
FROM
repo_groups a,
repo b
WHERE
a.repo_group_id = b.repo_group_id
ORDER BY
rg_name,
repo_name;
""")
repolist = pd.read_sql(repo_query, con=engine)
display(repolist)
repolist.dtypes
# -
# ### Create a Simpler List for quickly Identifying repo_group_id's and repo_id's for other queries
# +
repolist = pd.DataFrame()
repo_query = salc.sql.text(f"""
SELECT b.repo_id,
a.repo_group_id,
b.repo_name,
a.rg_name
FROM
repo_groups a,
repo b
WHERE
a.repo_group_id = b.repo_group_id
ORDER BY
rg_name,
repo_name;
""")
repolist = pd.read_sql(repo_query, con=engine)
display(repolist)
repolist.dtypes
# -
| START_HERE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
x = np.linspace(0,5,11)
y = x ** 2
x
y
# FUNCTIONAL METHOD -> BASIC METHOD
plt.plot(x,y)
plt.show() # Required if you are not in the jupyter notbook
plt.plot(x,y)
plt.xlabel('X Label')
plt.ylabel('Y Label')
plt.title('Title')
plt.show()
# +
# Plotting multiple charts
plt.subplot(1,2,1) # num of rows, columns, plot you're refering to
plt.plot(x,y,'r')
plt.subplot(1,2,2)
plt.plot(y,x, 'b')
plt.show()
# +
# OBJECT ORIENTED METHOD
fig = plt.figure()
axes = fig.add_axes([0.1, 0.1, 0.8, 0.8]) # left, botton, width and height -> range from 0 to 1
# Now plot something on the axes
axes.plot(x, y)
# +
# Multiple axes on the same fig
fig1 = plt.figure()
axes1 = fig1.add_axes([0.1, 0.1, 0.8, 0.8])
axes2 = fig1.add_axes([0.2, 0.5, 0.4, 0.3])
axes1.plot(x,y)
axes1.set_xlabel('X1')
axes1.set_ylabel('Y1')
axes1.set_title('Title 1')
axes2.plot(y,x, 'r')
axes2.set_xlabel('X2')
axes2.set_ylabel('Y2')
axes2.set_title('Title 2')
# +
# Part 2
# +
# Use the subplot call to automatically create n plots. There is some
# unpacking that happens and one of those is an array of axes you can reference.
fig,axes = plt.subplots(nrows=1,ncols=2)
for current_ax in axes:
current_ax.plot(x,y)
# +
fig1,ax1 = plt.subplots(nrows=1,ncols=2)
ax1[0].plot(x,y)
ax1[0].set_title('Title 0')
ax1[1].plot(x,y)
ax1[1].set_title('Title 1')
# -
# ## Figure Size and DPI
# +
# fit = plt.figure(figsize=(3,2), dpi=100)
fig = plt.figure(figsize=(8,2))
ax = fig.add_axes([0,0,1,1])
ax.plot(x,y)
# +
fig,axes = plt.subplots(nrows=2, ncols=1, figsize=(8,2))
axes[0].plot(x,y)
axes[1].plot(y,x)
plt.tight_layout()
# +
# Save a figure
# png,jpeg,pdf,xvg
fig.savefig('my_pic.png', dpi=200)
# +
# Quick way of setting titles
fig = plt.figure(figsize=(8,4))
ax = fig.add_axes([0,0,1,1])
ax.plot(x,x**2)
ax.plot(x,x**3)
ax.set_title('Title')
ax.set_xlabel('X Label')
ax.set_ylabel('Y Label')
# +
# Setting a legend
fig = plt.figure(figsize=(8,4))
ax = fig.add_axes([0,0,1,1])
ax.plot(x,x**2, label='X Squared')
ax.plot(x,x**3, label='X Cubed')
ax.legend(loc=0)
# -
| python-datasci-bootcamp/05-Data-Visualization-with-Matplotlib/t_matplotlib_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import scipy
import seaborn as sns
import matplotlib.pyplot as plt
import os
from functools import reduce
from statsmodels.tsa.stattools import coint
from sklearn import mixture as mix
from pykalman import KalmanFilter
import statsmodels.api as sm
import ffn
sns.set(style='white')
# Retrieve intraday price data and combine them into a DataFrame.
# 1. Load downloaded prices from folder into a list of dataframes.
folder_path = 'STATICS/PRICE'
file_names = ['ryaay.csv','rya.csv']
tickers = [name.split('.')[0] for name in file_names]
df_list = [pd.read_csv(os.path.join('STATICS/PRICE', name)) for name in file_names]
# +
# 2. Replace the closing price column name by the ticker.
for i in range(len(df_list)):
df_list[i].rename(columns={'close': tickers[i]}, inplace=True)
# 3. Merge all price dataframes
df = reduce(lambda x, y: pd.merge(x, y, on='date'), df_list)
df.set_index('date',inplace=True)
df.index = pd.to_datetime(df.index)
df.head()
# -
df.describe()
df.plot()
returns = df.pct_change()
returns.plot()
# +
sns.distplot(returns.iloc[:,0:1])
log_ret_RYAAY = np.log(df['ryaay']) - np.log(df['ryaay'].shift(1))
log_ret_RYA = np.log(df['rya']) - np.log(df['rya'].shift(1))
#Plot using Seaborn's jointplot function
sns.jointplot(log_ret_RYAAY, log_ret_RYA, kind='reg', size=12)
# -
score, pvalue, _ = coint(df['ryaay'],df['rya'])
print(pvalue)
print(df['ryaay'].corr(df['rya']))
kf = KalmanFilter(transition_matrices = [1],
observation_matrices = [1],
initial_state_mean = 0,
initial_state_covariance = 1,
observation_covariance=1,
transition_covariance=.01)
x_state_means, _ = kf.filter(df['rya'].values)
y_state_means, _ = kf.filter(df['ryaay'].values)
state_means = pd.Series(state_means.flatten(), index=x.index)
# +
def KalmanFilterAverage(x):
# Construct a Kalman filter
kf = KalmanFilter(transition_matrices = [1],
observation_matrices = [1],
initial_state_mean = 0,
initial_state_covariance = 1,
observation_covariance=1,
transition_covariance=.01)
# Use the observed values of the price to get a rolling mean
state_means, _ = kf.filter(x.values)
state_means = pd.Series(state_means.flatten(), index=x.index)
return state_means
# Kalman filter regression
def KalmanFilterRegression(x,y):
delta = 1e-3
trans_cov = delta / (1 - delta) * np.eye(2) # How much random walk wiggles
obs_mat = np.expand_dims(np.vstack([[x], [np.ones(len(x))]]).T, axis=1)
kf = KalmanFilter(n_dim_obs=1, n_dim_state=2, # y is 1-dimensional, (alpha, beta) is 2-dimensional
initial_state_mean=[0,0],
initial_state_covariance=np.ones((2, 2)),
transition_matrices=np.eye(2),
observation_matrices=obs_mat,
observation_covariance=2,
transition_covariance=trans_cov)
# Use the observations y to get running estimates and errors for the state parameters
state_means, state_covs = kf.filter(y.values)
return state_means
def half_life(spread):
spread_lag = spread.shift(1)
spread_lag.iloc[0] = spread_lag.iloc[1]
spread_ret = spread - spread_lag
spread_ret.iloc[0] = spread_ret.iloc[1]
spread_lag2 = sm.add_constant(spread_lag)
model = sm.OLS(spread_ret,spread_lag2)
res = model.fit()
halflife = int(round(-np.log(2) / res.params[1],0))
if halflife <= 0:
halflife = 1
return halflife
# -
def backtest(s1, s2, x, y):
#############################################################
# INPUT:
# s1: the symbol of contract one
# s2: the symbol of contract two
# x: the price series of contract one
# y: the price series of contract two
# OUTPUT:
# df1['cum rets']: cumulative returns in pandas data frame
# sharpe: sharpe ratio
# CAGR: CAGR
# run regression to find hedge ratio and then create spread series
df1 = pd.DataFrame({'y':y,'x':x})
state_means = KalmanFilterRegression(KalmanFilterAverage(x),KalmanFilterAverage(y))
df1['hr'] = - state_means[:,0]
df1['hr'] = df1['hr'].round(3)
df1['spread'] = df1.y + (df1.x * df1.hr)
# calculate half life
halflife = half_life(df1['spread'])
# calculate z-score with window = half life period
meanSpread = df1.spread.rolling(window=halflife).mean()
stdSpread = df1.spread.rolling(window=halflife).std()
df1['zScore'] = ((df1.spread-meanSpread)/stdSpread)#.shift(1)
##############################################################
# trading logic
entryZscore = 2
exitZscore = 0
#set up num units long
df1['long entry'] = ((df1.zScore < - entryZscore) & ( df1.zScore.shift(1) > - entryZscore))
df1['long exit'] = ((df1.zScore > - exitZscore) & (df1.zScore.shift(1) < - exitZscore))
df1['num units long'] = np.nan
df1.loc[df1['long entry'],'num units long'] = 1
df1.loc[df1['long exit'],'num units long'] = 0
df1['num units long'][0] = 0
df1['num units long'] = df1['num units long'].fillna(method='pad') #set up num units short
df1['short entry'] = ((df1.zScore > entryZscore) & ( df1.zScore.shift(1) < entryZscore))
df1['short exit'] = ((df1.zScore < exitZscore) & (df1.zScore.shift(1) > exitZscore))
df1.loc[df1['short entry'],'num units short'] = -1
df1.loc[df1['short exit'],'num units short'] = 0
df1['num units short'][0] = 0
df1['num units short'] = df1['num units short'].fillna(method='pad')
df1['numUnits'] = df1['num units long'] + df1['num units short']
# Boolean whether transaction occurred
df1['transaction'] = df1.numUnits.shift(1) != df1.numUnits
df1['positionSwitch'] = (df1.numUnits.shift(1) == (-df1.numUnits)) & df1['transaction']
# Cost of transaction
df1['tradecosts'] = (df1['transaction'] *1 + df1['positionSwitch']*1 ) * 0.0063
# Save hr during holding period
df1['hr_memory'] = np.nan
df1['hr_memory'][df1['transaction'] & df1['numUnits'] != 0] = df1.hr[df1['transaction'] & df1['numUnits'] != 0]
df1['hr_memory'].fillna(method='ffill',inplace=True)
# Save investment amount during holding period
df1['invest_memory'] = np.nan
df1['invest_memory'][df1['transaction'] & df1['numUnits'] != 0] = ((df1['x'] * abs(df1['hr'])) + df1['y'])[df1['transaction'] & df1['numUnits'] != 0]
df1['invest_memory'].fillna(method='ffill',inplace=True)
df1['spreadmemory'] = df1.y + (df1.x * df1.hr_memory)
df1['spread pct ch'] = (df1['spreadmemory'] - df1['spreadmemory'].shift(1)) / df1['invest_memory']
df1['port rets'] = df1['spread pct ch'] * df1['numUnits'].shift(1) - (df1['tradecosts'] /df1['invest_memory'])
#Account for the position switch
df1['port rets'][df1['positionSwitch']] = (((df1.y + (df1.x * df1.hr_memory.shift(1))\
- df1['spreadmemory'].shift(1)) / df1['invest_memory'].shift(1))\
* df1['numUnits'].shift(1) - (df1['tradecosts'] /df1['invest_memory'].shift(1)))[df1['positionSwitch']]
df1['cum rets'] = df1['port rets'].cumsum()
df1['cum rets'] = df1['cum rets'] + 1
name = "bt"+ s1 + "-" + s2 + ".csv"
df1.to_csv(name)
return df1
# +
import warnings
warnings.filterwarnings('ignore')
out = backtest('rya','ryaay',df['rya'],df['ryaay'])
# -
out['cum rets'][0] = 1
perf =out['cum rets'].calc_stats()
perf.set_riskfree_rate(0.0016)
perf.display()
def plot_signals(d, fromDate, toDate):
#idx = pd.date_range(fromDate,toDate, freq="1min")
d = d[fromDate:toDate]
#d = d.reindex(idx, fill_value= np.nan)
d.index = d.index.map(str)
# Plot the prices and buy and sell signals from z score
S = d.y - d.x * 5
S.plot(color='b')
buyS = 0*S.copy()
sellS = 0*S.copy()
exitL = 0*S.copy()
exitS = 0*S.copy()
longentry = d['long entry'] * d.transaction * (d.numUnits == 1)
longexit = d['long exit'] * d.transaction * (d.numUnits.shift(1) == 1)
shortentry = d['short entry'] * d.transaction * (d.numUnits == -1)
shortexit = d['short exit'] * d.transaction * (d.numUnits.shift(1) == -1)
buyS[longentry] = S[longentry]
sellS[shortentry] = S[shortentry]
exitL[longexit] = S[longexit]
exitS[shortexit] = S[shortexit]
buyS.plot(color='g', linestyle='None', marker='o')
sellS.plot(color='r', linestyle='None', marker='o')
exitL.plot(color='g', linestyle='None', marker='x')
exitS.plot(color='r', linestyle='None', marker='x')
x1,x2,y1,y2 = plt.axis()
plt.axis((x1,x2,S.min(),S.max()))
plt.legend(['LOP Spread', 'Enter Long', 'Enter Short','Exit Long', 'Exit Short'])
plt.xticks(rotation=45, ha="right")
plt.show()
print('{} percent return in time window'.format(round(d['port rets'].sum() *100,2)))
# +
plot_signals(out,"2020-03-18","2020-04-25")
plt.plot(out["2018-03-24":"2020-04-25"]['cum rets'])
# +
#Get spread
S = out.y - out.x * out.hr
#Fit gaussian
unsup = mix.GaussianMixture(n_components=4,
covariance_type="spherical",
n_init=100,
random_state=42)
unsup.fit(S.values.reshape(-1, 1))
# Predict
regime = unsup.predict(S.values.reshape(-1, 1))
S = S.to_frame()
S['Return']= np.log(S/S.shift(1))
Regimes=pd.DataFrame(regime,columns=['Regime'],index=S.index).join(S, how='inner')\
.assign(market_cu_return=S.Return.cumsum())\
.reset_index(drop=False)\
.rename(columns={'index':'date'})
fig = sns.FacetGrid(data=Regimes,hue='Regime',hue_order=[0,1,2,3],aspect=2,size= 4)
fig.map(plt.scatter,'date','market_cu_return', s=4).add_legend()
plt.show()
# -
for i in [0,1,2,3]:
print('Mean for regime %i: '%i,unsup.means_[i][0])
print('Co-Variancefor regime %i: '%i,(unsup.covariances_[i]))
| .ipynb_checkpoints/Benchmark-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 21741, "status": "ok", "timestamp": 1609942401542, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="e4abR9zSaWNk" outputId="94ee80ef-152f-452f-d852-06919b9942a4"
# Mount Google Drive
from google.colab import drive # import drive from google colab
ROOT = "/content/drive" # default location for the drive
print(ROOT) # print content of ROOT (Optional)
drive.mount(ROOT) # we mount the google drive at /content/drive
# + executionInfo={"elapsed": 4192, "status": "ok", "timestamp": 1610025781584, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="gk5AKGKcYGOo"
# !pip install pennylane
from IPython.display import clear_output
clear_output()
# + id="GigSJusGbx1b"
import os
def restart_runtime():
os.kill(os.getpid(), 9)
restart_runtime()
# + executionInfo={"elapsed": 857, "status": "ok", "timestamp": 1610025787385, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="HoLmJLkIX810"
# # %matplotlib inline
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1 import make_axes_locatable
import numpy as np
import tensorflow as tf
from tensorflow.keras.utils import to_categorical
# + [markdown] id="vZFNOwFXoY8N"
# # Loading Raw Data
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2726, "status": "ok", "timestamp": 1610025792089, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="IvdFsGCVof9g" outputId="c1e94fa1-a11f-4bff-c7b7-975b8b1b79fb"
(x_train, y_train), (x_test, y_test) = tf.keras.datasets.mnist.load_data()
x_train = x_train[:, 0:27, 0:27]
x_test = x_test[:, 0:27, 0:27]
# + executionInfo={"elapsed": 1251, "status": "ok", "timestamp": 1610025792677, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="c6zvGFvIoxAN"
x_train_flatten = x_train.reshape(x_train.shape[0], x_train.shape[1]*x_train.shape[2])/255.0
x_test_flatten = x_test.reshape(x_test.shape[0], x_test.shape[1]*x_test.shape[2])/255.0
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 948, "status": "ok", "timestamp": 1610025792678, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="Rmj1dzaso00h" outputId="f59eea77-95bb-4b6a-b0ba-14a7e5760b11"
print(x_train_flatten.shape, y_train.shape)
print(x_test_flatten.shape, y_test.shape)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 945, "status": "ok", "timestamp": 1610025793507, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="d10VoIC6o5_I" outputId="774bafb6-5cbe-42b3-98d0-643e7e943f91"
x_train_0 = x_train_flatten[y_train == 0]
x_train_1 = x_train_flatten[y_train == 1]
x_train_2 = x_train_flatten[y_train == 2]
x_train_3 = x_train_flatten[y_train == 3]
x_train_4 = x_train_flatten[y_train == 4]
x_train_5 = x_train_flatten[y_train == 5]
x_train_6 = x_train_flatten[y_train == 6]
x_train_7 = x_train_flatten[y_train == 7]
x_train_8 = x_train_flatten[y_train == 8]
x_train_9 = x_train_flatten[y_train == 9]
x_train_list = [x_train_0, x_train_1, x_train_2, x_train_3, x_train_4, x_train_5, x_train_6, x_train_7, x_train_8, x_train_9]
print(x_train_0.shape)
print(x_train_1.shape)
print(x_train_2.shape)
print(x_train_3.shape)
print(x_train_4.shape)
print(x_train_5.shape)
print(x_train_6.shape)
print(x_train_7.shape)
print(x_train_8.shape)
print(x_train_9.shape)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 901, "status": "ok", "timestamp": 1610025796926, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="snFw4LqepFOl" outputId="d62bdedd-8ad4-4da9-c7e8-75b8a0a9ba7b"
x_test_0 = x_test_flatten[y_test == 0]
x_test_1 = x_test_flatten[y_test == 1]
x_test_2 = x_test_flatten[y_test == 2]
x_test_3 = x_test_flatten[y_test == 3]
x_test_4 = x_test_flatten[y_test == 4]
x_test_5 = x_test_flatten[y_test == 5]
x_test_6 = x_test_flatten[y_test == 6]
x_test_7 = x_test_flatten[y_test == 7]
x_test_8 = x_test_flatten[y_test == 8]
x_test_9 = x_test_flatten[y_test == 9]
x_test_list = [x_test_0, x_test_1, x_test_2, x_test_3, x_test_4, x_test_5, x_test_6, x_test_7, x_test_8, x_test_9]
print(x_test_0.shape)
print(x_test_1.shape)
print(x_test_2.shape)
print(x_test_3.shape)
print(x_test_4.shape)
print(x_test_5.shape)
print(x_test_6.shape)
print(x_test_7.shape)
print(x_test_8.shape)
print(x_test_9.shape)
# + [markdown] id="SAxUS6Lhp95g"
# # Selecting the dataset
#
# Output: X_train, Y_train, X_test, Y_test
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 810, "status": "ok", "timestamp": 1609950462215, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}, "user_tz": -420} id="f--pX5Oto_XB" outputId="cc074972-59f4-4d74-cd69-7745260a1dd4"
n_train_sample_per_class = 200
n_class = 4
X_train = x_train_list[0][:n_train_sample_per_class, :]
Y_train = np.zeros((X_train.shape[0]*n_class,), dtype=int)
for i in range(n_class-1):
X_train = np.concatenate((X_train, x_train_list[i+1][:n_train_sample_per_class, :]), axis=0)
Y_train[(i+1)*n_train_sample_per_class:(i+2)*n_train_sample_per_class] = i+1
X_train.shape, Y_train.shape
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1252, "status": "ok", "timestamp": 1609950468260, "user": {"displayName": "<NAME>", "photoUrl": "<KEY>", "userId": "03770692095188133952"}, "user_tz": -420} id="W_SHH9e3rqwG" outputId="2f38a646-7a1a-4e68-828f-bce0c97bfe70"
n_test_sample_per_class = int(2.5*n_train_sample_per_class)
X_test = x_test_list[0][:n_test_sample_per_class, :]
Y_test = np.zeros((X_test.shape[0]*n_class,), dtype=int)
for i in range(n_class-1):
X_test = np.concatenate((X_test, x_test_list[i+1][:n_test_sample_per_class, :]), axis=0)
Y_test[(i+1)*n_test_sample_per_class:(i+2)*n_test_sample_per_class] = i+1
X_test.shape, Y_test.shape
# -
# # Dataset Preprocessing
# +
X_train = X_train.reshape(X_train.shape[0], 27, 27, 1)
X_test = X_test.reshape(X_test.shape[0], 27, 27, 1)
X_train.shape, X_test.shape
# -
class_label = np.loadtxt('./tetra_class_label.txt')
# +
Y_train = to_categorical(Y_train)
Y_test = to_categorical(Y_test)
# for i in range(n_class):
# Y_train[Y_train[:, i] == 1.] = class_label[i]
# for i in range(n_class):
# Y_test[Y_test[:, i] == 1.] = class_label[i]
# -
Y_train = np.concatenate((Y_train, Y_train), axis=1)
Y_test = np.concatenate((Y_test, Y_test), axis=1)
Y_train.shape, Y_test.shape
# # Quantum
# +
# session_conf = tf.compat.v1.ConfigProto(intra_op_parallelism_threads=16, inter_op_parallelism_threads=16)
# tf.compat.v1.set_random_seed(1)
# sess = tf.compat.v1.Session(graph=tf.compat.v1.get_default_graph(), config=session_conf)
# tf.compat.v1.keras.backend.set_session(sess)
# +
import pennylane as qml
from pennylane import numpy as np
from pennylane.optimize import AdamOptimizer, GradientDescentOptimizer
qml.enable_tape()
# Set a random seed
np.random.seed(2020)
# -
# Define output labels as quantum state vectors
def density_matrix(state):
"""Calculates the density matrix representation of a state.
Args:
state (array[complex]): array representing a quantum state vector
Returns:
dm: (array[complex]): array representing the density matrix
"""
return np.outer(state, np.conj(state))
state_labels = np.loadtxt('./tetra_states.txt', dtype=np.complex_)
state_labels = np.concatenate((state_labels, state_labels), axis=0)
# +
# my_bucket = "amazon-braket-0f5d17943f73" # the name of the bucket
# my_prefix = "Tugas_Akhir" # the name of the folder in the bucket
# s3_folder = (my_bucket, my_prefix)
# device_arn = "arn:aws:braket:::device/quantum-simulator/amazon/sv1"
# +
n_qubits = int(2*n_class)
dev = qml.device("default.qubit", wires=n_qubits)
#dev = qml.device('cirq.simulator', wires=n_qubits)
#interface="tf", grad_method="backprop"
@qml.qnode(dev)
def qcircuit(params, inputs):
"""A variational quantum circuit representing the DRC.
Args:
params (array[float]): array of parameters
inputs = [x, y]
x (array[float]): 1-d input vector
y (array[float]): single output state density matrix
Returns:
float: fidelity between output state and input
"""
# layer iteration
for l in range(len(params[0])):
# qubit iteration
for q in range(n_class):
# first qubit
qml.Rot(*(params[0][l][q][0:3] * inputs[0:3] + params[1][l][q][0:3]), wires=q)
qml.Rot((params[0][l][q][3] * inputs[3] + params[1][l][q][3]), params[1][l][q][4], params[1][l][q][5], wires=q)
# second qubit
qml.Rot(*(params[0][l][q+n_class][0:3] * inputs[0:3] + params[1][l][q+n_class][0:3]), wires=(q+n_class))
qml.Rot((params[0][l][q+n_class][3] * inputs[3] + params[1][l][q+n_class][3]), params[1][l][q+n_class][4], params[1][l][q+n_class][5], wires=(q+n_class))
# entangling layer
qml.CZ(wires=[q, (q+n_class)])
return [qml.expval(qml.Hermitian(density_matrix(state_labels[i]), wires=[i])) for i in range(n_qubits)]
# -
class class_weights(tf.keras.layers.Layer):
def __init__(self):
super(class_weights, self).__init__()
w_init = tf.random_normal_initializer()
self.w = tf.Variable(
initial_value=w_init(shape=(1, 2*n_class), dtype="float32"),
trainable=True,
)
def call(self, inputs):
return (inputs * self.w)
# +
X = tf.keras.Input(shape=(27,27,1))
conv_layer_1 = tf.keras.layers.Conv2D(filters=1, kernel_size=[3,3], strides=[2,2], name='Conv_Layer_1', activation='relu')(X)
conv_layer_2 = tf.keras.layers.Conv2D(filters=1, kernel_size=[3,3], strides=[2,2], name='Conv_Layer_2', activation='relu')(conv_layer_1)
conv_layer_3 = tf.keras.layers.Conv2D(filters=1, kernel_size=[3,3], strides=[3,3], name='Conv_Layer_3', activation='relu')(conv_layer_2)
#max__pool_layer = tf.keras.layers.MaxPooling2D(pool_size=(2, 2), strides=None, name='Max_Pool_Layer')(conv_layer_2)
reshapor_layer = tf.keras.layers.Reshape((4,), name='Reshapor_Layer')(conv_layer_3)
qlayer = qml.qnn.KerasLayer(qcircuit, {"params": (2, 2, 2*n_class, 6)}, output_dim=2*n_class, name='Quantum_Layer')(reshapor_layer)
class_weights_layer = class_weights()(qlayer)
model = tf.keras.Model(inputs=X, outputs=class_weights_layer, name='Conv DRC')
# -
model(X_train[0:32])
model.summary()
# +
import keras.backend as K
# def custom_loss(y_true, y_pred):
# return K.sum(((y_true.shape[1]-2)*y_true+1)*K.square(y_true-y_pred))/len(y_true)
def custom_loss(y_true, y_pred):
return K.sum(K.square(y_true-y_pred))/(2*len(y_true))
# -
opt = tf.keras.optimizers.Adam(learning_rate=0.1)
model.compile(opt, loss=custom_loss, metrics=["accuracy"])
model.fit(X_train, Y_train, epochs=50, batch_size=32, validation_data=(X_test, Y_test), verbose=1)
predict_test = model.predict(X_test)
| PennyLane/Data Reuploading Classifier/Conv + DRC Keras MNIST 4 class - 2 qubit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""Intermolecular Interactions and Symmetry-Adapted Perturbation Theory"""
__authors__ = "<NAME>"
__email__ = ["<EMAIL>"]
__copyright__ = "(c) 2008-2020, The Psi4Education Developers"
__license__ = "BSD-3-Clause"
__date__ = "2020-07-16"
# -
# This lab activity is designed to teach students about weak intermolecular interactions, and the calculation and interpretation of the interaction energy between two molecules. The interaction energy can be broken down into physically meaningful contributions (electrostatics, induction, dispersion, and exchange) using symmetry-adapted perturbation theory (SAPT). In this exercise, we will calculate complete interaction energies and their SAPT decomposition using the procedures from the Psi4 software package, processing and analyzing the data with NumPy and Matplotlib.
#
# Prerequisite knowledge: the Hartree-Fock method, molecular orbitals, electron correlation and the MP2 theory. The lab also assumes all the standard Python prerequisites of all Psi4Education labs.
#
# Learning Objectives:
# 1. Recognize and appreciate the ubiquity and diversity of intermolecular interactions.
# 2. Compare and contrast the supermolecular and perturbative methods of calculating interaction energy.
# 3. Analyze and interpret the electrostatic, induction, dispersion, and exchange SAPT contributions at different intermolecular separations.
#
# Author: <NAME>, Auburn University (<EMAIL>; ORCID: 0000-0002-4468-207X)
#
# Copyright: Psi4Education Project, 2020
#
# # Weak intermolecular interactions
#
# In this activity, you will examine some properties of weak interactions between molecules. As the molecular subunits are not connected by any covalent (or ionic) bonds, we often use the term *noncovalent interactions*. Suppose we want to calculate the interaction energy between molecule A and molecule B for a certain geometry of the A-B complex (obviously, this interaction energy depends on how far apart the molecules are and how they are oriented). The simplest way of doing so is by subtraction (in the so-called *supermolecular approach*):
#
# \begin{equation}
# E_{\rm int}=E_{\rm A-B}-E_{\rm A}-E_{\rm B}
# \end{equation}
#
# where $E_{\rm X}$ is the total energy of system X, computed using our favorite electronic structure theory and basis set. A negative value of $E_{\rm int}$ means that A and B have a lower energy when they are together than when they are apart, so they do form a weakly bound complex that might be stable at least at very low temperatures. A positive value of $E_{\rm int}$ means that the A-B complex is unbound - it is energetically favorable for A and B to go their separate ways.
#
# Let's consider a simple example of two interacting helium atoms and calculate $E_{\rm int}$ at a few different interatomic distances $R$. You will use Psi4 to calculate the total energies that you need to perform subtraction. When you do so for a couple different $R$, you will be able to sketch the *potential energy curve* - the graph of $E_{\rm int}(R)$ as a function of $R$.
#
# OK, but how should you pick the electronic structure method to calculate $E_{\rm A-B}$, $E_{\rm A}$, and $E_{\rm B}$? Let's start with the simplest choice and try out the Hartree-Fock (HF) method. In case HF is not accurate enough, we will also try the coupled-cluster method with single, double, and perturbative triple excitations - CCSD(T). If you haven't heard about CCSD(T) before, let's just state that it is **(1)** usually very accurate (it's even called the *gold standard* of electronic structure theory) and **(2)** very expensive for larger molecules. For the basis set, let's pick the augmented correlation consistent triple-zeta (aug-cc-pVTZ) basis of Dunning which should be quite OK for both HF and CCSD(T).
#
# +
# A simple Psi4 input script to compute the potential energy curve for two helium atoms
# %matplotlib notebook
import time
import numpy as np
import scipy
from scipy.optimize import *
np.set_printoptions(precision=5, linewidth=200, threshold=2000, suppress=True)
import psi4
import matplotlib.pyplot as plt
# Set Psi4 & NumPy Memory Options
psi4.set_memory('2 GB')
psi4.core.set_output_file('output.dat', False)
numpy_memory = 2
psi4.set_options({'basis': 'aug-cc-pVTZ',
'e_convergence': 1e-10,
'd_convergence': 1e-10,
'INTS_TOLERANCE': 1e-15})
# -
# We need to collect some data points to graph the function $E_{\rm int}(R)$. Therefore, we set up a list of distances $R$ for which we will run the calculations (we go with 11 of them). For each distance, we need to remember three values ($E_{\rm A-B}$, $E_{\rm A}$, and $E_{\rm B}$). For this purpose, we will prepare two $11\times 3$ NumPy arrays to hold the HF and CCSD(T) results.
#
# +
distances = [4.0,4.5,5.0,5.3,5.6,6.0,6.5,7.0,8.0,9.0,10.0]
ehf = np.zeros((11,3))
eccsdt = np.zeros((11,3))
# -
# We are almost ready to crunch some numbers! One question though: how are we going to tell Psi4 whether we want $E_{\rm A-B}$, $E_{\rm A}$, or $E_{\rm B}$?
# We need to define three different geometries. The $E_{\rm A-B}$ one has two helium atoms $R$ atomic units from each other - we can place one atom at $(0,0,0)$ and the other at $(0,0,R)$. The other two geometries involve one actual helium atom, with a nucleus and two electrons, and one *ghost atom* in place of the other one. A ghost atom does not have a nucleus or electrons, but it does carry the same basis functions as an actual atom - we need to calculate all energies in the same basis set, with functions centered at both $(0,0,0)$ and $(0,0,R)$, to prevent the so-called *basis set superposition error*. In Psi4, the syntax `Gh(X)` denotes a ghost atom where basis functions for atom type X are located.
#
# Using ghost atoms, we can now easily define geometries for the $E_{\rm A}$ and $E_{\rm B}$ calculations.
#
# +
for i in range(len(distances)):
dimer = psi4.geometry("""
He 0.0 0.0 0.0
--
He 0.0 0.0 """+str(distances[i])+"""
units bohr
symmetry c1
""")
psi4.energy('ccsd(t)') #HF will be calculated along the way
ehf[i,0] = psi4.variable('HF TOTAL ENERGY')
eccsdt[i,0] = psi4.variable('CCSD(T) TOTAL ENERGY')
psi4.core.clean()
monomerA = psi4.geometry("""
He 0.0 0.0 0.0
--
Gh(He) 0.0 0.0 """+str(distances[i])+"""
units bohr
symmetry c1
""")
psi4.energy('ccsd(t)') #HF will be calculated along the way
ehf[i,1] = psi4.variable('HF TOTAL ENERGY')
eccsdt[i,1] = psi4.variable('CCSD(T) TOTAL ENERGY')
psi4.core.clean()
monomerB = psi4.geometry("""
Gh(He) 0.0 0.0 0.0
--
He 0.0 0.0 """+str(distances[i])+"""
units bohr
symmetry c1
""")
psi4.energy('ccsd(t)') #HF will be calculated along the way
ehf[i,2] = psi4.variable('HF TOTAL ENERGY')
eccsdt[i,2] = psi4.variable('CCSD(T) TOTAL ENERGY')
psi4.core.clean()
# -
# We have completed the $E_{\rm A-B}$, $E_{\rm A}$, or $E_{\rm B}$ calculations for all 11 distances $R$ (it didn't take that long, did it?). We will now perform the subtraction to form NumPy arrays with $E_{\rm int}(R)$ values for each method, converted from atomic units (hartrees) to kcal/mol, and graph the resulting potential energy curves using the matplotlib library.
#
# +
#COMPLETE the two lines below to generate interaction energies. Convert them from atomic units to kcal/mol.
einthf =
eintccsdt =
print ('HF PEC',einthf)
print ('CCSD(T) PEC',eintccsdt)
plt.plot(distances,einthf,'r+',linestyle='-',label='HF')
plt.plot(distances,eintccsdt,'bo',linestyle='-',label='CCSD(T)')
plt.hlines(0.0,4.0,10.0)
plt.legend(loc='upper right')
plt.show()
# -
# *Questions*
# 1. Which curve makes more physical sense?
# 2. Why does helium form a liquid at very low temperatures?
# 3. You learned in freshman chemistry that two helium atoms do not form a molecule because there are two electrons on a bonding orbital and two electrons on an antibonding orbital. How does this information relate to the behavior of HF (which does assume a molecular orbital for every electron) and CCSD(T) (which goes beyond the molecular orbital picture)?
# 4. When you increase the size of the interacting molecules, the CCSD(T) method quickly gets much more expensive and your calculation might take weeks instead of seconds. It gets especially expensive for the calculation of $E_{\rm A-B}$ because A-B has more electrons than either A or B. Your friend suggests to use CCSD(T) only for the easier terms $E_{\rm A}$ and $E_{\rm B}$ and subtract them from $E_{\rm A-B}$ calculated with a different, cheaper method such as HF. Why is this a really bad idea?
#
# *To answer the questions above, please double click this Markdown cell to edit it. When you are done entering your answers, run this cell as if it was a code cell, and your Markdown source will be recompiled.*
#
# A nice feature of the supermolecular approach is that it is very easy to use - you just need to run three standard energy calculations, and modern quantum chemistry codes such as Psi4 give you a lot of methods to choose from. However, the accuracy of subtraction hinges on error cancellation, and we have to be careful to ensure that the errors do cancel between $E_{\rm A-B}$ and $E_{\rm A}+E_{\rm B}$. Another drawback of the supermolecular approach is that it is not particularly rich in physical insight. All that we get is a single number $E_{\rm int}$ that tells us very little about the underlying physics of the interaction. Therefore, one may want to find an alternative approach where $E_{\rm int}$ is computed directly, without subtraction, and it is obtained as a sum of distinct, physically meaningful terms. Symmetry-adapted perturbation theory (SAPT) is such an alternative approach.
#
# # Symmetry-Adapted Perturbation Theory (SAPT)
#
# SAPT is a perturbation theory aimed specifically at calculating the interaction energy between two molecules. Contrary to the supermolecular approach, SAPT obtains the interaction energy directly - no subtraction of similar terms is needed. Moreover, the result is obtained as a sum of separate corrections accounting for the electrostatic, induction, dispersion, and exchange contributions to interaction energy, so the SAPT decomposition facilitates the understanding and physical interpretation of results.
# - *Electrostatic energy* arises from the Coulomb interaction between charge densities of isolated molecules.
# - *Induction energy* is the energetic effect of mutual polarization between the two molecules.
# - *Dispersion energy* is a consequence of intermolecular electron correlation, usually explained in terms of correlated fluctuations of electron density on both molecules.
# - *Exchange energy* is a short-range repulsive effect that is a consequence of the Pauli exclusion principle.
#
# In this activity, we will explore the simplest level of the SAPT theory called SAPT0 (see [Parker:2014] for the definitions of different levels of SAPT). A particular SAPT correction $E^{(nk)}$ corresponds to effects that are of $n$th order in the intermolecular interaction and $k$th order in the intramolecular electron correlation. In SAPT0, intramolecular correlation is neglected, and intermolecular interaction is included through second order:
#
# \begin{equation}
# E_{\rm int}^{\rm SAPT0}=E^{(10)}_{\rm elst}+E^{(10)}_{\rm exch}+E^{(20)}_{\rm ind,resp}+E^{(20)}_{\rm exch-ind,resp}+E^{(20)}_{\rm disp}+E^{(20)}_{\rm exch-disp}+\delta E^{(2)}_{\rm HF}
# \end{equation}
#
# In this equation, the consecutive corrections account for the electrostatic, first-order exchange, induction, exchange induction, dispersion, and exchange dispersion effects, respectively. The additional subscript ''resp'' denotes that these corrections are computed including response effects - the HF orbitals of each molecule are relaxed in the electric field generated by the other molecule. The last term $\delta E^{(2)}_{\rm HF}$ approximates third- and higher-order induction and exchange induction effects and is taken from a supermolecular HF calculation.
#
# Sticking to our example of two helium atoms, let's now calculate the SAPT0 interaction energy contributions using Psi4. In the results that follow, we will group $E^{(20)}_{\rm ind,resp}$, $E^{(20)}_{\rm exch-ind,resp}$, and $\delta E^{(2)}_{\rm HF}$ to define the total induction effect (including its exchange quenching), and group $E^{(20)}_{\rm disp}$ with $E^{(20)}_{\rm exch-disp}$ to define the total dispersion effect.
#
# +
distances = [4.0,4.5,5.0,5.3,5.6,6.0,6.5,7.0,8.0,9.0,10.0]
eelst = np.zeros((11))
eexch = np.zeros((11))
eind = np.zeros((11))
edisp = np.zeros((11))
esapt = np.zeros((11))
for i in range(len(distances)):
dimer = psi4.geometry("""
He 0.0 0.0 0.0
--
He 0.0 0.0 """+str(distances[i])+"""
units bohr
symmetry c1
""")
psi4.energy('sapt0')
eelst[i] = psi4.variable('SAPT ELST ENERGY') * 627.509
eexch[i] = psi4.variable('SAPT EXCH ENERGY') * 627.509
eind[i] = psi4.variable('SAPT IND ENERGY') * 627.509
edisp[i] = psi4.variable('SAPT DISP ENERGY') * 627.509
esapt[i] = psi4.variable('SAPT TOTAL ENERGY') * 627.509
psi4.core.clean()
plt.close()
plt.ylim(-0.2,0.4)
plt.plot(distances,eelst,'r+',linestyle='-',label='SAPT0 elst')
plt.plot(distances,eexch,'bo',linestyle='-',label='SAPT0 exch')
plt.plot(distances,eind,'g^',linestyle='-',label='SAPT0 ind')
plt.plot(distances,edisp,'mx',linestyle='-',label='SAPT0 disp')
plt.plot(distances,esapt,'k*',linestyle='-',label='SAPT0 total')
plt.hlines(0.0,4.0,10.0)
plt.legend(loc='upper right')
plt.show()
# -
# *Questions*
# 1. What is the origin of attraction between two helium atoms?
# 2. For the interaction of two helium atoms, which SAPT terms are *long-range* (vanish with distance like some inverse power of $R$) and which are *short-range* (vanish exponentially with $R$ just like the overlap of molecular orbitals)?
# 3. The dispersion energy decays at large $R$ like $R^{-n}$. Find the value of $n$ by fitting a function to the five largest-$R$ results. You can use `scipy.optimize.curve_fit` to perform the fitting, but you have to define the appropriate function first.
# Does the optimal exponent $n$ obtained by your fit agree with what you know about van der Waals dispersion forces? Is the graph of dispersion energy shaped like the $R^{-n}$ graph for large $R$? What about intermediate $R$?
#
# *Do you know how to calculate $R^{-n}$ if you have an array with $R$ values? If not, look it up in the NumPy documentation!*
#
# +
#COMPLETE the definition of function f below.
def f
ndisp = scipy.optimize.curve_fit(f,distances[-5:],edisp[-5:])
print ("Optimal dispersion exponent:",ndisp[0][0])
# -
# # Interaction between two water molecules
#
# For the next part, you will perform the same analysis and obtain the supermolecular and SAPT0 data for the interaction of two water molecules. We now have many more degrees of freedom: in addition to the intermolecular distance $R$, we can change the relative orientation of two molecules, or even their internal geometries (O-H bond lengths and H-O-H angles). In this way, the potential energy curve becomes a multidimensional *potential energy surface*. It is hard to graph functions of more than two variables, so we will stick to the distance dependence of the interaction energies. Therefore, we will assume one particular orientation of two water molecules (a hydrogen-bonded one) and vary the intermolecular distance $R$ while keeping the orientation, and molecular geometries, constant. The geometry of the A-B complex has been defined for you, but you have to request all the necessary Psi4 calculations and extract the numbers that you need. To save time, we will downgrade the basis set to aug-cc-pVDZ and use MP2 (an approximate method that captures most of electron correlation) in place of CCSD(T).
#
# *Hints:* To prepare the geometries for the individual water molecules A and B, copy and paste the A-B geometry, but use the Gh(O2)... syntax to define the appropriate ghost atoms. Remember to run `psi4.core.clean()` after each calculation.
#
# +
distances_h2o = [2.7,3.0,3.5,4.0,4.5,5.0,6.0,7.0,8.0,9.0]
ehf_h2o = np.zeros((10,3))
emp2_h2o = np.zeros((10,3))
psi4.set_options({'basis': 'aug-cc-pVDZ'})
for i in range(len(distances_h2o)):
dimer = psi4.geometry("""
O1
H1 O1 0.96
H2 O1 0.96 H1 104.5
--
O2 O1 """+str(distances_h2o[i])+""" H1 5.0 H2 0.0
X O2 1.0 O1 120.0 H2 180.0
H3 O2 0.96 X 52.25 O1 90.0
H4 O2 0.96 X 52.25 O1 -90.0
units angstrom
symmetry c1
""")
#COMPLETE the MP2 energy calculations for A-B, A, and B, and prepare the data for the graph.
#Copy and paste the A-B geometry, but use the Gh(O2)... syntax to define the appropriate ghost atoms for the A and B calculations.
#Remember to run psi4.core.clean() after each calculation.
print ('HF PEC',einthf_h2o)
print ('MP2 PEC',eintmp2_h2o)
plt.close()
plt.plot(distances_h2o,einthf_h2o,'r+',linestyle='-',label='HF')
plt.plot(distances_h2o,eintmp2_h2o,'bo',linestyle='-',label='MP2')
plt.hlines(0.0,2.5,9.0)
plt.legend(loc='upper right')
plt.show()
# +
eelst_h2o = np.zeros((10))
eexch_h2o = np.zeros((10))
eind_h2o = np.zeros((10))
edisp_h2o = np.zeros((10))
esapt_h2o = np.zeros((10))
#COMPLETE the SAPT calculations for 10 distances to prepare the data for the graph.
plt.close()
plt.ylim(-10.0,10.0)
plt.plot(distances_h2o,eelst_h2o,'r+',linestyle='-',label='SAPT0 elst')
plt.plot(distances_h2o,eexch_h2o,'bo',linestyle='-',label='SAPT0 exch')
plt.plot(distances_h2o,eind_h2o,'g^',linestyle='-',label='SAPT0 ind')
plt.plot(distances_h2o,edisp_h2o,'mx',linestyle='-',label='SAPT0 disp')
plt.plot(distances_h2o,esapt_h2o,'k*',linestyle='-',label='SAPT0 total')
plt.hlines(0.0,2.5,9.0)
plt.legend(loc='upper right')
plt.show()
# -
# Before we proceed any further, let us check one thing about your first MP2 water-water interaction energy calculation, the one that produced `eintmp2_h2o[0]`. Here's the geometry of that complex again:
#
#all x,y,z in Angstroms
atomtypes = ["O1","H1","H2","O2","H3","H4"]
coordinates = np.array([[0.116724185090, 1.383860971547, 0.000000000000],
[0.116724185090, 0.423860971547, 0.000000000000],
[-0.812697549673, 1.624225775439, 0.000000000000],
[-0.118596320329, -1.305864713301, 0.000000000000],
[0.362842754701, -1.642971982825, -0.759061990794],
[0.362842754701, -1.642971982825, 0.759061990794]])
# First, write the code to compute the four O-H bond lengths and two H-O-H bond angles in the two molecules. *(Hint: if the angles look weird, maybe they are still in radians - don't forget to convert them to degrees.)* Are the two water molecules identical?
#
# Then, check the values of the MP2 energy for these two molecules (the numbers $E_{\rm A}$ and $E_{\rm B}$ that you subtracted to get the interaction energy). If the molecules are the same, why are the MP2 energies close but not the same?
#
# *Hints:* The most elegant way to write this code is to define functions `distance(point1,point2)` for the distance between two points $(x_1,y_1,z_1)$ and $(x_2,y_2,z_2)$, and `angle(vec1,vec2)` for the angle between two vectors $(x_{v1},y_{v1},z_{v1})$ and $(x_{v2},y_{v2},z_{v2})$. Recall that the cosine of this angle is related to the dot product $(x_{v1},y_{v1},z_{v1})\cdot(x_{v2},y_{v2},z_{v2})$. If needed, check the documentation on how to calculate the dot product of two NumPy vectors.
#
# When you are parsing the NumPy array with the coordinates, remember that `coordinates[k,:]` is the vector of $(x,y,z)$ values for atom number $k$, $k=0,1,2,\ldots,N_{\rm atoms}-1$.
#
# +
#COMPLETE the distance and angle calculations below.
ro1h1 =
ro1h2 =
ro2h3 =
ro2h4 =
ah1o1h2 =
ah3o2h4 =
print ('O-H distances: %5.3f %5.3f %5.3f %5.3f' % (ro1h1,ro1h2,ro2h3,ro2h4))
print ('H-O-H angles: %6.2f %6.2f' % (ah1o1h2,ah3o2h4))
print ('MP2 energy of molecule 1: %18.12f hartrees' % emp2_h2o[0,1])
print ('MP2 energy of molecule 2: %18.12f hartrees' % emp2_h2o[0,2])
# -
# We can now proceed with the analysis of the SAPT0 energy components for the complex of two water molecules. *Please edit this Markdown cell to write your answers.*
# 1. Which of the four SAPT terms are long-range, and which are short-range this time?
# 2. For the terms that are long-range and decay with $R$ like $R^{-n}$, estimate $n$ by fitting a proper function to the 5 data points with the largest $R$, just like you did for the two interacting helium atoms (using `scipy.optimize.curve_fit`). How would you explain the power $n$ that you obtained for the electrostatic energy?
#
#COMPLETE the optimizations below.
nelst_h2o =
nind_h2o =
ndisp_h2o =
print ("Optimal electrostatics exponent:",nelst_h2o[0][0])
print ("Optimal induction exponent:",nind_h2o[0][0])
print ("Optimal dispersion exponent:",ndisp_h2o[0][0])
# The water molecules are polar - each one has a nonzero dipole moment, and at large distances we expect the electrostatic energy to be dominated by the dipole-dipole interaction (at short distances, when the orbitals of two molecules overlap, the multipole approximation is not valid and the electrostatic energy contains the short-range *charge penetration* effects). Let's check if this is indeed the case. In preparation for this, we first find the HF dipole moment vector for each water molecule.
#
# +
waterA = psi4.geometry("""
O 0.116724185090 1.383860971547 0.000000000000
H 0.116724185090 0.423860971547 0.000000000000
H -0.812697549673 1.624225775439 0.000000000000
units angstrom
noreorient
nocom
symmetry c1
""")
comA = waterA.center_of_mass()
comA = np.array([comA[0],comA[1],comA[2]])
E, wfn = psi4.energy('HF',return_wfn=True)
dipoleA = np.array([psi4.variable('SCF DIPOLE X'),psi4.variable('SCF DIPOLE Y'),
psi4.variable('SCF DIPOLE Z')])*0.393456 # conversion from Debye to a.u.
psi4.core.clean()
print("COM A in a.u.",comA)
print("Dipole A in a.u.",dipoleA)
waterB = psi4.geometry("""
O -0.118596320329 -1.305864713301 0.000000000000
H 0.362842754701 -1.642971982825 -0.759061990794
H 0.362842754701 -1.642971982825 0.759061990794
units angstrom
noreorient
nocom
symmetry c1
""")
comB = waterB.center_of_mass()
comB = np.array([comB[0],comB[1],comB[2]])
E, wfn = psi4.energy('HF',return_wfn=True)
dipoleB = np.array([psi4.variable('SCF DIPOLE X'),psi4.variable('SCF DIPOLE Y'),
psi4.variable('SCF DIPOLE Z')])*0.393456 # conversion from Debye to a.u.
psi4.core.clean()
print("COM B in a.u.",comB)
print("Dipole B in a.u.",dipoleB)
comA_to_comB = comB - comA
print("Vector from COMA to COMB:",comA_to_comB)
# -
# Our goal now is to plot the electrostatic energy from SAPT against the interaction energy between two dipoles $\boldsymbol{\mu_A}$ and $\boldsymbol{\mu_B}$:
#
# \begin{equation}
# E_{\rm dipole-dipole}=\frac{\boldsymbol{\mu_A}\cdot\boldsymbol{\mu_B}}{R^3}-\frac{3(\boldsymbol{\mu_A}\cdot{\mathbf R})(\boldsymbol{\mu_B}\cdot{\mathbf R})}{R^5}
# \end{equation}
#
# Program this formula in the `dipole_dipole` function below, taking ${\mathbf R}$, $\boldsymbol{\mu_A}$, and $\boldsymbol{\mu_B}$ in atomic units and calculating the dipole-dipole interaction energy, also in atomic units (which we will later convert to kcal/mol).
# With your new function, we can populate the `edipdip` array of dipole-dipole interaction energies for all intermolecular separations, and plot these energies alongside the actual electrostatic energy data from SAPT.
#
# Note that ${\mathbf R}$ is the vector from the center of mass of molecule A to the center of mass of molecule B. For the shortest intermolecular distance, the atomic coordinates are listed in the code above, so `R = comA_to_comB`. For any other distance, we obtained the geometry of the complex by shifting one water molecule away from the other along the O-O direction, so we need to shift the center of mass of the second molecule in the same way.
#
# +
#the geometries are related to each other by a shift of 1 molecule along the O-O vector:
OA_to_OB = (np.array([-0.118596320329,-1.305864713301,0.000000000000])-np.array(
[0.116724185090,1.383860971547,0.000000000000]))/0.529177249
OA_to_OB_unit = OA_to_OB/np.sqrt(np.sum(OA_to_OB*OA_to_OB))
print("Vector from OA to OB:",OA_to_OB,OA_to_OB_unit)
def dipole_dipole(R,dipA,dipB):
#COMPLETE the definition of the dipole-dipole energy. All your data are in atomic units.
edipdip = []
for i in range(len(distances_h2o)):
shiftlength = (distances_h2o[i]-distances_h2o[0])/0.529177249
R = comA_to_comB + shiftlength*OA_to_OB_unit
edipdip.append(dipole_dipole(R,dipoleA,dipoleB)*627.509)
edipdip = np.array(edipdip)
print (edipdip)
plt.close()
plt.ylim(-10.0,10.0)
plt.plot(distances_h2o,eelst_h2o,'r+',linestyle='-',label='SAPT0 elst')
plt.plot(distances_h2o,edipdip,'bo',linestyle='-',label='dipole-dipole')
plt.hlines(0.0,2.5,9.0)
plt.legend(loc='upper right')
plt.show()
# -
# We clearly have a favorable dipole-dipole interaction, which results in negative (attractive) electrostatic energy. This is how the origins of hydrogen bonding might have been explained to you in your freshman chemistry class: two polar molecules have nonzero dipole moments and the dipole-dipole interaction can be strongly attractive. However, your SAPT components show you that it's not a complete explanation: the two water molecules are bound not only by electrostatics, but by two other SAPT components as well. Can you quantify the relative (percentage) contributions of electrostatics, induction, and dispersion to the overall interaction energy at the van der Waals minimum? This minimum is the second point on your curve, so, for example, `esapt_h2o[1]` is the total SAPT interaction energy.
#
# +
#now let's examine the SAPT0 contributions at the van der Waals minimum, which is the 2nd point on the curve
#COMPLETE the calculation of percentages.
percent_elst =
percent_ind =
percent_disp =
print ('At the van der Waals minimum, electrostatics, induction, and dispersion')
print (' contribute %5.1f, %5.1f, and %5.1f percent of interaction energy, respectively.'
% (percent_elst,percent_ind,percent_disp))
# -
# You have now completed some SAPT calculations and analyzed the meaning of different corrections. Can you complete the table below to indicate whether different SAPT corrections can be positive (repulsive), negative (attractive), or both, and why?
#
#Type in your answers below.
#COMPLETE this table. Do not remove the comment (#) signs.
#
#SAPT term Positive/Negative/Both? Why?
#Electrostatics
#Exchange
#Induction
#Dispersion
# # Ternary diagrams
#
# Higher levels of SAPT calculations can give very accurate interaction energies, but are more computationally expensive than SAPT0. SAPT0 is normally sufficient for qualitative accuracy and basic understanding of the interaction physics. One important use of SAPT0 is to *classify different intermolecular complexes according to the type of interaction*, and a nice way to display the results of this classification is provided by a *ternary diagram*.
#
# The relative importance of attractive electrostatic, induction, and dispersion contributions to a SAPT interaction energy for a particular structure can be marked as a point inside a triangle, with the distance to each vertex of the triangle depicting the relative contribution of a given type (the more dominant a given contribution is, the closer the point lies to the corresponding vertex). If the electrostatic contribution is repulsive, we can display the relative magnitudes of electrostatic, induction, and dispersion terms in the same way, but we need the second triangle (the left one). The combination of two triangles forms the complete diagram and we can mark lots of different points corresponding to different complexes and geometries.
#
# Let's now mark all your systems on a ternary diagram, in blue for two helium atoms and in red for two water molecules. What kinds of interaction are represented? Compare your diagram with the one pictured below, prepared for 2510 different geometries of the complex of two water molecules, with all kinds of intermolecular distances and orientations (this graph is taken from [Smith:2016]). What conclusions can you draw about the interaction of two water molecules at *any* orientation?
#
# +
def ternary(sapt, title='', labeled=True, view=True, saveas=None, relpath=False, graphicsformat=['pdf']):
#Adapted from the QCDB ternary diagram code by <NAME>
"""Takes array of arrays *sapt* in form [elst, indc, disp] and builds formatted
two-triangle ternary diagrams. Either fully-readable or dotsonly depending
on *labeled*.
"""
from matplotlib.path import Path
import matplotlib.patches as patches
# initialize plot
plt.close()
fig, ax = plt.subplots(figsize=(6, 3.6))
plt.xlim([-0.75, 1.25])
plt.ylim([-0.18, 1.02])
plt.xticks([])
plt.yticks([])
ax.set_aspect('equal')
if labeled:
# form and color ternary triangles
codes = [Path.MOVETO, Path.LINETO, Path.LINETO, Path.CLOSEPOLY]
pathPos = Path([(0., 0.), (1., 0.), (0.5, 0.866), (0., 0.)], codes)
pathNeg = Path([(0., 0.), (-0.5, 0.866), (0.5, 0.866), (0., 0.)], codes)
ax.add_patch(patches.PathPatch(pathPos, facecolor='white', lw=2))
ax.add_patch(patches.PathPatch(pathNeg, facecolor='#fff5ee', lw=2))
# label corners
ax.text(1.0,
-0.15,
u'Elst (−)',
verticalalignment='bottom',
horizontalalignment='center',
family='Times New Roman',
weight='bold',
fontsize=18)
ax.text(0.5,
0.9,
u'Ind (−)',
verticalalignment='bottom',
horizontalalignment='center',
family='Times New Roman',
weight='bold',
fontsize=18)
ax.text(0.0,
-0.15,
u'Disp (−)',
verticalalignment='bottom',
horizontalalignment='center',
family='Times New Roman',
weight='bold',
fontsize=18)
ax.text(-0.5,
0.9,
u'Elst (+)',
verticalalignment='bottom',
horizontalalignment='center',
family='Times New Roman',
weight='bold',
fontsize=18)
xvals = []
yvals = []
cvals = []
geomindex = 0 # first 11 points are He-He, the next 10 are H2O-H2O
for sys in sapt:
[elst, indc, disp] = sys
# calc ternary posn and color
Ftop = abs(indc) / (abs(elst) + abs(indc) + abs(disp))
Fright = abs(elst) / (abs(elst) + abs(indc) + abs(disp))
xdot = 0.5 * Ftop + Fright
ydot = 0.866 * Ftop
if geomindex <= 10:
cdot = 'b'
else:
cdot = 'r'
if elst > 0.:
xdot = 0.5 * (Ftop - Fright)
ydot = 0.866 * (Ftop + Fright)
#print elst, indc, disp, '', xdot, ydot, cdot
xvals.append(xdot)
yvals.append(ydot)
cvals.append(cdot)
geomindex += 1
sc = ax.scatter(xvals, yvals, c=cvals, s=15, marker="o",
edgecolor='none', vmin=0, vmax=1, zorder=10)
# remove figure outline
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['bottom'].set_visible(False)
ax.spines['left'].set_visible(False)
# save and show
plt.show()
return 1
sapt = []
for i in range(11):
sapt.append([eelst[i],eind[i],edisp[i]])
for i in range(10):
sapt.append([eelst_h2o[i],eind_h2o[i],edisp_h2o[i]])
idummy = ternary(sapt)
from IPython.display import Image
Image(filename='water2510.png')
# -
# # Some further reading:
#
# 1. How is the calculation of SAPT corrections actually programmed? The Psi4NumPy projects has some tutorials on this topic: https://github.com/psi4/psi4numpy/tree/master/Tutorials/07_Symmetry_Adapted_Perturbation_Theory
# 2. A classic (but recently updated) book on the theory of interactions between molecules: "The Theory of Intermolecular Forces"
# > [[Stone:2013](https://www.worldcat.org/title/theory-of-intermolecular-forces/oclc/915959704)] A. Stone, Oxford University Press, 2013
# 3. The classic review paper on SAPT: "Perturbation Theory Approach to Intermolecular Potential Energy Surfaces of van der Waals Complexes"
# > [[Jeziorski:1994](http://pubs.acs.org/doi/abs/10.1021/cr00031a008)] <NAME>, <NAME>, and <NAME>, *Chem. Rev.* **94**, 1887 (1994)
# 4. A brand new (as of 2020) review of SAPT, describing new developments and inprovements to the theory: "Recent developments in symmetry‐adapted perturbation theory"
# > [[Patkowski:2020](https://onlinelibrary.wiley.com/doi/abs/10.1002/wcms.1452)] <NAME>, *WIREs Comput. Mol. Sci.* **10**, e1452 (2020)
# 5. The definitions and practical comparison of different levels of SAPT: "Levels of symmetry adapted perturbation theory (SAPT). I. Efficiency and performance for interaction energies"
# > [[Parker:2014](http://aip.scitation.org/doi/10.1063/1.4867135)] <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>, *J. Chem. Phys.* **140**, 094106 (2014)
# 6. An example study making use of the SAPT0 classification of interaction types, with lots of ternary diagrams in the paper and in the supporting information: "Revised Damping Parameters for the D3 Dispersion Correction to Density Functional Theory"
# > [[Smith:2016](https://pubs.acs.org/doi/abs/10.1021/acs.jpclett.6b00780)] <NAME>, <NAME>, <NAME>, and <NAME>, *J. Phys. Chem. Lett.* **7**, 2197 (2016).
#
| labs/Symmetry_Adapted_Perturbation_Theory/sapt0_student.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import requests
from bs4 import BeautifulSoup
import pandas as pd
import re
import os
pd.set_option('display.max_rows', None)
URL = "https://www.oryxspioenkop.com/2022/02/attack-on-europe-documenting-equipment.html"
page = requests.get(URL)
soup = BeautifulSoup(page.content, "html.parser")
results = soup.find(id="post-body-8087922975012177708")
all_h3 = soup.find_all('h3')
#equipment_types_auto = pd.DataFrame()
equipment_types_auto_tmp = []
equipment_types_auto = []
# +
for h3 in all_h3:
if not 'Ukraine' in h3.get_text() and not 'Russia' in h3.get_text():
equipment_type = h3.get_text().partition("(")[0]
equipment_type = re.sub(r"\s$", "", equipment_type)
equipment_types_auto_tmp.append([equipment_type])
for i in equipment_types_auto_tmp:
if i not in equipment_types_auto:
equipment_types_auto.append(i)
equipment_types_auto = [val for sublist in equipment_types_auto for val in sublist]
equipment_types_auto.remove('')
# -
equipment_types_auto
equipment_subtypes_auto_tmp = []
equipment_subtypes_auto = []
all_li = soup.find_all('li', attrs={'class': None})
# +
for li in all_li:
equipment_subtype = re.search(r'(.*):', li.get_text())
if equipment_subtype is not None:
equipment_subtype = equipment_subtype.group(0)
equipment_subtype = re.sub("^ \d+", "", equipment_subtype)
equipment_subtype = re.sub(":", "", equipment_subtype)
equipment_subtype = re.sub(r"^\s", "", equipment_subtype)
equipment_subtypes_auto_tmp.append([equipment_subtype])
for i in equipment_subtypes_auto_tmp:
if i[0] not in equipment_subtypes_auto:
equipment_subtypes_auto.append(i[0])
# -
equipment_subtypes_auto
status_types_auto_tmp = []
status_types_auto = []
all_a = soup.find_all('a')
# +
for a in all_a:
status = re.search(r"\((.*)\)", a.get_text())
if status is not None:
status = status.group(0)
status = re.sub("\(", "", status)
status = re.sub("\)", "", status)
status = re.search(r"([^\,]+$)", status)
status = status.group(0)
status = re.sub(r"^\s", "", status)
status_types_auto_tmp.append([status])
for i in status_types_auto_tmp:
if i[0] not in status_types_auto:
status_types_auto.append(i[0])
# -
status_types_auto
# +
for h3 in all_h3:
if h3.get_text().count('Ukraine') == 1:
all_russian = h3.find_all_previous()
all_ukraine = h3.find_all_next()
all_russian_all_ukraine = [all_russian, all_ukraine]
# -
list_tmp = []
# +
country = 'RUS'
for section in all_russian_all_ukraine:
if all_russian_all_ukraine.index(section) == 1:
country = 'UKR'
for element in section:
if element.name == 'h3':
for equipment_type_i in equipment_types_auto:
equipment_type = element.get_text().partition("(")[0]
equipment_type = re.sub(r"\s$", "", equipment_type)
if equipment_type_i == equipment_type:
current_type = equipment_type_i
#if equipment_type in element.get_text():
# current_type = equipment_type
ul = element.nextSibling.nextSibling
try:
li_list = ul.find_all('li')
except:
pass
for li in li_list:
li_a_list = li.find_all('a')
for equipment_subtype_i in equipment_subtypes_auto:
equipment_subtype = re.search(r'(.*):', li.get_text())
if equipment_subtype is not None:
equipment_subtype = equipment_subtype.group(0)
equipment_subtype = re.sub("^ \d+", "", equipment_subtype)
equipment_subtype = re.sub(":", "", equipment_subtype)
equipment_subtype = re.sub(r"^\s", "", equipment_subtype)
if equipment_subtype_i == equipment_subtype:
current_subtype = equipment_subtype_i
for status_i in status_types_auto:
for single_report in li_a_list:
current_a_text = single_report.get_text()
current_a_text = re.sub("\(", "", current_a_text)
current_a_text = re.sub("\)", "", current_a_text)
current_a_text = re.search(r"([^\,]+$)", current_a_text)
current_a_text = current_a_text.group(0)
current_a_text = re.sub(r"^\s", "", current_a_text)
if status_i == current_a_text:
list_tmp.append([country, current_type, current_subtype, current_a_text, single_report['href']])
# -
df = pd.DataFrame(list_tmp, columns=['country', 'equipment_type', 'equipment_subtype', 'satus', 'source'])
df
cwd = os.getcwd()
path = cwd + '/export.csv'
df.to_csv(path)
| .ipynb_checkpoints/notebook-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import gym
from gym import logger as gymlogger
gymlogger.set_level(40) # error only
import pybulletgym # register PyBullet enviroments with open ai gym
import numpy as np
from gym import wrappers
import torch
import torch.nn as nn
import torch.autograd as autograd
# +
#define the simulated environment we are going to use
env = gym.make("InvertedPendulumMuJoCoEnv-v0")
#check out the pacman action space!
print(env.action_space)
# should return a state vector if everything worked (value doesn't matter)
env.reset()
# +
env = gym.make("Walker2DMuJoCoEnv-v0")
print(env.action_space)
env.reset()
# +
env = gym.make("HalfCheetahMuJoCoEnv-v0")
print(env.action_space)
env.reset()
# -
env = gym.make("HalfCheetahMuJoCoEnv-v0")
env = wrappers.Monitor(env, './HalfCheetahVideo/')
env.reset()
for _ in range(1000):
env.render()
env.step(env.action_space.sample()) # take a random action
env.close()
| Environment_Setup/Environment_Check.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.4
# language: julia
# name: julia-1.6
# ---
# # Nonlinear Online Estimation
#
# In this demo we consider a model for the spead of a virus in a population. We are interested in estimating the reproduction rate from daily observations of the number of infected individuals. The reproduction rate indicates how many others are (on average) infected by one infected individual per time unit.
using ForneyLab
using PyPlot
# ## Generate Data
#
# We start by generating a toy dataset for a virus with a reproduction rate $a$. Here, $y$ represents the measured number of infected individuals, and $x$ a latent state.
# +
# Generate toy dataset
T = 15
x_0_data = 0.6
a_data = 1.2
x_data = Vector{Float64}(undef, T)
y_data = Vector{Float64}(undef, T)
x_t_min_data = x_0_data
for t=1:T
x_data[t] = a_data*x_t_min_data
y_data[t] = ceil(x_data[t])
x_t_min_data = x_data[t]
end
;
# -
# Inspect data
bar(1:T, y_data)
grid("on")
xlabel("t [days]")
ylabel("y [infected]")
;
# ## Define Model
#
# We define a state-space model, where we specify the state transition by a `Nonlinear` node. We may optionally define an inverse function for each of the arguments of the forward transition `g`. It is important that the function arguments, and the order in which these functions are passed to the `Nonlinear` node specification, follow the order: `out`, `in1`, `in2`, etc. In our specific case, we have `x_t`, `x_t_min`, `a`, with corresponding functions `g`, `g_inv_x_t_min`, `g_inv_a`, respectively. If an inverse function is unavailable, `nothing` may be substituted. In this case, ForneyLab will attempt to compute the backward message by an RTS-smoothing procedure.
# +
fg = FactorGraph()
# Specify forward transition
g(x_t_min::Float64, a::Float64) = a*x_t_min
# Specify inverse functions (optional)
# g_inv_x_t_min(x_t::Float64, a::Float64) = x_t/a
# g_inv_a(x_t::Float64, x_t_min::Float64) = x_t/x_t_min
@RV x_t_min ~ GaussianMeanVariance(placeholder(:m_x_t_min), placeholder(:v_x_t_min))
@RV a ~ GaussianMeanVariance(placeholder(:m_a_t_min), placeholder(:v_a_t_min))
@RV x_t ~ Nonlinear{Unscented}(x_t_min, a; g=g) # Inverse functions (optional): g_inv=[g_inv_x_t_min, g_inv_a]
@RV y_t ~ GaussianMeanVariance(x_t, 0.2)
placeholder(y_t, :y_t)
;
# -
# ## Derive Algorithm
#
# The message passing algorithm, including a procedure for evaluating the free energy, is automatically derived.
algo = messagePassingAlgorithm([x_t, a], free_energy=true)
code = algorithmSourceCode(algo, free_energy=true)
;
# +
# println(code)
# -
eval(Meta.parse(code))
;
# ## Execute Algorithm
#
# We execute the algorithm in an online fashion, where after each timestep the posteriors for the state and reproduction rate are used as priors for the next timestep.
# +
# Prior statistics
m_x_0 = 0.6
v_x_0 = 0.5
m_a_0 = 1.0
v_a_0 = 0.5
F = Vector{Float64}(undef, T)
m_x = Vector{Float64}(undef, T)
v_x = Vector{Float64}(undef, T)
m_a = Vector{Float64}(undef, T)
v_a = Vector{Float64}(undef, T)
# Execute online algorithm
m_x_t_min = m_x_0
v_x_t_min = v_x_0
m_a_t_min = m_a_0
v_a_t_min = v_a_0
for t=1:T
data = Dict(:y_t => y_data[t],
:m_x_t_min => m_x_t_min,
:v_x_t_min => v_x_t_min,
:m_a_t_min => m_a_t_min,
:v_a_t_min => v_a_t_min)
marginals = step!(data) # Infer posteriors
F[t] = freeEnergy(data, marginals) # Evaluate free energy
# Extract posterior statistics
(m_x_t, v_x_t) = ForneyLab.unsafeMeanCov(marginals[:x_t])
(m_a_t, v_a_t) = ForneyLab.unsafeMeanCov(marginals[:a])
# Reset for next step
m_x_t_min = m_x[t] = m_x_t
v_x_t_min = v_x[t] = v_x_t
m_a_t_min = m_a[t] = m_a_t
v_a_t_min = v_a[t] = v_a_t
end
# -
# ## Inspect Results
# +
subplot(3,1,1)
bar(1:T, y_data)
plot(1:T, m_x, color="black")
fill_between(1:T, m_x.-sqrt.(v_x), m_x.+sqrt.(v_x), color="black", alpha=0.3)
grid("on")
xlim(1,T)
ylabel("y [infected]")
subplot(3,1,2)
plot(1:T, m_a, color="black")
fill_between(1:T, m_a.-sqrt.(v_a), m_a.+sqrt.(v_a), color="black", alpha=0.3)
plot(1:T, a_data*ones(T), color="black", linestyle="dashed")
grid("on")
xlim(1,T)
ylabel("a [repr. rate]")
subplot(3,1,3)
plot(1:T, F, color="black")
grid("on")
xlim(1,T)
xlabel("t [days]")
ylabel("F [free energy]")
;
# -
| demo/nonlinear_online_estimation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.10 64-bit (''nma'': conda)'
# name: python3710jvsc74a57bd03e19903e646247cead5404f55ff575624523d45cf244c3f93aaf5fa10367032a
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W3D1_BayesianDecisions/student/W3D1_Tutorial1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# -
# # Neuromatch Academy: Week 3, Day 1, Tutorial 1
# # Bayes with a binary hidden state
#
# __Content creators:__ [insert your name here]
#
# __Content reviewers:__
# # Tutorial Objectives
# This is the first in a series of two core tutorials on Bayesian statistics. In these tutorials, we will explore the fundemental concepts of the Bayesian approach from two perspectives. This tutorial will work through an example of Bayesian inference and decision making using a binary hidden state. The second main tutorial extends these concepts to a continuous hidden state. In the next days, each of these basic ideas will be extended--first through time as we consider what happens when we infere a hidden state using multiple observations and when the hidden state changes across time. In the third day, we will introduce the notion of how to use inference and decisions to select actions for optimal control. For this tutorial, you will be introduced to our binary state fishing problem!
#
# This notebook will introduce the fundamental building blocks for Bayesian statistics:
#
# 1. How do we use probability distributions to represent hidden states?
# 2. How does marginalization work and how can we use it?
# 3. How do we combine new information with our prior knowledge?
# 4. How do we combine the possible loss (or gain) for making a decision with our probabilitic knowledge?
#
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 517} outputId="f2f0a0ff-8ebf-486d-8a45-c3e012298e7d"
#@title Video 1: Introduction to Bayesian Statistics
from IPython.display import YouTubeVideo
video = YouTubeVideo(id='JiEIn9QsrFg', width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
# -
# ## Setup
# Please execute the cells below to initialize the notebook environment.
# +
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import patches
from matplotlib import transforms
from matplotlib import gridspec
from scipy.optimize import fsolve
from collections import namedtuple
# + cellView="form"
#@title Figure Settings
import ipywidgets as widgets # interactive display
from ipywidgets import GridspecLayout
from IPython.display import clear_output
# %config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
import warnings
warnings.filterwarnings("ignore")
# + cellView="form"
# @title Plotting Functions
def plot_joint_probs(P, ):
assert np.all(P >= 0), "probabilities should be >= 0"
# normalize if not
P = P / np.sum(P)
marginal_y = np.sum(P,axis=1)
marginal_x = np.sum(P,axis=0)
# definitions for the axes
left, width = 0.1, 0.65
bottom, height = 0.1, 0.65
spacing = 0.005
# start with a square Figure
fig = plt.figure(figsize=(5, 5))
joint_prob = [left, bottom, width, height]
rect_histx = [left, bottom + height + spacing, width, 0.2]
rect_histy = [left + width + spacing, bottom, 0.2, height]
rect_x_cmap = plt.cm.Blues
rect_y_cmap = plt.cm.Reds
# Show joint probs and marginals
ax = fig.add_axes(joint_prob)
ax_x = fig.add_axes(rect_histx, sharex=ax)
ax_y = fig.add_axes(rect_histy, sharey=ax)
# Show joint probs and marginals
ax.matshow(P,vmin=0., vmax=1., cmap='Greys')
ax_x.bar(0, marginal_x[0], facecolor=rect_x_cmap(marginal_x[0]))
ax_x.bar(1, marginal_x[1], facecolor=rect_x_cmap(marginal_x[1]))
ax_y.barh(0, marginal_y[0], facecolor=rect_y_cmap(marginal_y[0]))
ax_y.barh(1, marginal_y[1], facecolor=rect_y_cmap(marginal_y[1]))
# set limits
ax_x.set_ylim([0,1])
ax_y.set_xlim([0,1])
# show values
ind = np.arange(2)
x,y = np.meshgrid(ind,ind)
for i,j in zip(x.flatten(), y.flatten()):
c = f"{P[i,j]:.2f}"
ax.text(j,i, c, va='center', ha='center', color='black')
for i in ind:
v = marginal_x[i]
c = f"{v:.2f}"
ax_x.text(i, v +0.1, c, va='center', ha='center', color='black')
v = marginal_y[i]
c = f"{v:.2f}"
ax_y.text(v+0.2, i, c, va='center', ha='center', color='black')
# set up labels
ax.xaxis.tick_bottom()
ax.yaxis.tick_left()
ax.set_xticks([0,1])
ax.set_yticks([0,1])
ax.set_xticklabels(['Silver','Gold'])
ax.set_yticklabels(['Small', 'Large'])
ax.set_xlabel('color')
ax.set_ylabel('size')
ax_x.axis('off')
ax_y.axis('off')
return fig
# test
# P = np.random.rand(2,2)
# P = np.asarray([[0.9, 0.8], [0.4, 0.1]])
# P = P / np.sum(P)
# fig = plot_joint_probs(P)
# plt.show(fig)
# plt.close(fig)
# fig = plot_prior_likelihood(0.5, 0.3)
# plt.show(fig)
# plt.close(fig)
def plot_prior_likelihood_posterior(prior, likelihood, posterior):
# definitions for the axes
left, width = 0.05, 0.3
bottom, height = 0.05, 0.9
padding = 0.1
small_width = 0.1
left_space = left + small_width + padding
added_space = padding + width
fig = plt.figure(figsize=(10, 4))
rect_prior = [left, bottom, small_width, height]
rect_likelihood = [left_space , bottom , width, height]
rect_posterior = [left_space + added_space, bottom , width, height]
ax_prior = fig.add_axes(rect_prior)
ax_likelihood = fig.add_axes(rect_likelihood, sharey=ax_prior)
ax_posterior = fig.add_axes(rect_posterior, sharey = ax_prior)
rect_colormap = plt.cm.Blues
# Show posterior probs and marginals
ax_prior.barh(0, prior[0], facecolor = rect_colormap(prior[0, 0]))
ax_prior.barh(1, prior[1], facecolor = rect_colormap(prior[1, 0]))
ax_likelihood.matshow(likelihood, vmin=0., vmax=1., cmap='Reds')
ax_posterior.matshow(posterior, vmin=0., vmax=1., cmap='Greens')
# Probabilities plot details
ax_prior.set(xlim = [1, 0], yticks = [0, 1], yticklabels = ['left', 'right'],
ylabel = 'state (s)', title = "Prior p(s)")
ax_prior.axis('off')
# Likelihood plot details
ax_likelihood.set(xticks = [0, 1], xticklabels = ['fish', 'no fish'],
yticks = [0, 1], yticklabels = ['left', 'right'],
ylabel = 'state (s)', xlabel = 'measurement (m)',
title = 'Likelihood p(m (right) | s)')
ax_likelihood.xaxis.set_ticks_position('bottom')
ax_likelihood.spines['left'].set_visible(False)
ax_likelihood.spines['bottom'].set_visible(False)
# Posterior plot details
ax_posterior.set(xticks = [0, 1], xticklabels = ['fish', 'no fish'],
yticks = [0, 1], yticklabels = ['left', 'right'],
ylabel = 'state (s)', xlabel = 'measurement (m)',
title = 'Posterior p(s | m)')
ax_posterior.xaxis.set_ticks_position('bottom')
ax_posterior.spines['left'].set_visible(False)
ax_posterior.spines['bottom'].set_visible(False)
# show values
ind = np.arange(2)
x,y = np.meshgrid(ind,ind)
for i,j in zip(x.flatten(), y.flatten()):
c = f"{posterior[i,j]:.2f}"
ax_posterior.text(j,i, c, va='center', ha='center', color='black')
for i,j in zip(x.flatten(), y.flatten()):
c = f"{likelihood[i,j]:.2f}"
ax_likelihood.text(j,i, c, va='center', ha='center', color='black')
for i in ind:
v = prior[i, 0]
c = f"{v:.2f}"
ax_prior.text(v+0.2, i, c, va='center', ha='center', color='black')
def plot_prior_likelihood(ps, p_a_s1, p_a_s0, measurement):
likelihood = np.asarray([[p_a_s1, 1-p_a_s1],[p_a_s0, 1-p_a_s0]])
assert 0.0 <= ps <= 1.0
prior = np.asarray([ps, 1 - ps])
if measurement:
posterior = likelihood[:, 0] * prior
else:
posterior = (likelihood[:, 1] * prior).reshape(-1)
posterior /= np.sum(posterior)
# definitions for the axes
left, width = 0.05, 0.3
bottom, height = 0.05, 0.9
padding = 0.1
small_width = 0.22
left_space = left + small_width + padding
small_padding = 0.05
fig = plt.figure(figsize=(10, 4))
rect_prior = [left, bottom, small_width, height]
rect_likelihood = [left_space , bottom , width, height]
rect_posterior = [left_space + width + small_padding, bottom , small_width, height]
ax_prior = fig.add_axes(rect_prior)
ax_likelihood = fig.add_axes(rect_likelihood, sharey=ax_prior)
ax_posterior = fig.add_axes(rect_posterior, sharey=ax_prior)
prior_colormap = plt.cm.Blues
posterior_colormap = plt.cm.Greens
# Show posterior probs and marginals
ax_prior.barh(0, prior[0], facecolor = prior_colormap(prior[0]))
ax_prior.barh(1, prior[1], facecolor = prior_colormap(prior[1]))
ax_likelihood.matshow(likelihood, vmin=0., vmax=1., cmap='Reds')
# ax_posterior.matshow(posterior, vmin=0., vmax=1., cmap='')
ax_posterior.barh(0, posterior[0], facecolor = posterior_colormap(posterior[0]))
ax_posterior.barh(1, posterior[1], facecolor = posterior_colormap(posterior[1]))
# Probabilities plot details
ax_prior.set(xlim = [1, 0], yticks = [0, 1], yticklabels = ['left', 'right'],
ylabel = 'state (s)', title = "Prior p(s)")
ax_prior.axis('off')
# Likelihood plot details
ax_likelihood.set(xticks = [0, 1], xticklabels = ['fish', 'no fish'],
yticks = [0, 1], yticklabels = ['left', 'right'],
ylabel = 'state (s)', xlabel = 'measurement (m)',
title = 'Likelihood p(m | s)')
ax_likelihood.xaxis.set_ticks_position('bottom')
ax_likelihood.spines['left'].set_visible(False)
ax_likelihood.spines['bottom'].set_visible(False)
# Posterior plot details
ax_posterior.set(xlim = [0, 1], yticks = [0, 1], yticklabels = ['left', 'right'],
ylabel = 'state (s)', title = "Posterior p(s | m)")
ax_posterior.axis('off')
# ax_posterior.set(xticks = [0, 1], xticklabels = ['fish', 'no fish'],
# yticks = [0, 1], yticklabels = ['left', 'right'],
# ylabel = 'state (s)', xlabel = 'measurement (m)',
# title = 'Posterior p(s | m)')
# ax_posterior.xaxis.set_ticks_position('bottom')
# ax_posterior.spines['left'].set_visible(False)
# ax_posterior.spines['bottom'].set_visible(False)
# show values
ind = np.arange(2)
x,y = np.meshgrid(ind,ind)
# for i,j in zip(x.flatten(), y.flatten()):
# c = f"{posterior[i,j]:.2f}"
# ax_posterior.text(j,i, c, va='center', ha='center', color='black')
for i in ind:
v = posterior[i]
c = f"{v:.2f}"
ax_posterior.text(v+0.2, i, c, va='center', ha='center', color='black')
for i,j in zip(x.flatten(), y.flatten()):
c = f"{likelihood[i,j]:.2f}"
ax_likelihood.text(j,i, c, va='center', ha='center', color='black')
for i in ind:
v = prior[i]
c = f"{v:.2f}"
ax_prior.text(v+0.2, i, c, va='center', ha='center', color='black')
return fig
# fig = plot_prior_likelihood(0.5, 0.3)
# plt.show(fig)
# plt.close(fig)
from matplotlib import colors
def plot_utility(ps):
prior = np.asarray([ps, 1 - ps])
utility = np.array([[2, -3], [-2, 1]])
expected = prior @ utility
# definitions for the axes
left, width = 0.05, 0.16
bottom, height = 0.05, 0.9
padding = 0.04
small_width = 0.1
left_space = left + small_width + padding
added_space = padding + width
fig = plt.figure(figsize=(17, 3))
rect_prior = [left, bottom, small_width, height]
rect_utility = [left + added_space , bottom , width, height]
rect_expected = [left + 2* added_space, bottom , width, height]
ax_prior = fig.add_axes(rect_prior)
ax_utility = fig.add_axes(rect_utility, sharey=ax_prior)
ax_expected = fig.add_axes(rect_expected)
rect_colormap = plt.cm.Blues
# Data of plots
ax_prior.barh(0, prior[0], facecolor = rect_colormap(prior[0]))
ax_prior.barh(1, prior[1], facecolor = rect_colormap(prior[1]))
ax_utility.matshow(utility, cmap='cool')
norm = colors.Normalize(vmin=-3, vmax=3)
ax_expected.bar(0, expected[0], facecolor = rect_colormap(norm(expected[0])))
ax_expected.bar(1, expected[1], facecolor = rect_colormap(norm(expected[1])))
# Probabilities plot details
ax_prior.set(xlim = [1, 0], yticks = [0, 1], yticklabels = ['left', 'right'],
ylabel = 'state (s)', title = "Probability of state")
ax_prior.axis('off')
# Utility plot details
ax_utility.set(xticks = [0, 1], xticklabels = ['left', 'right'],
yticks = [0, 1], yticklabels = ['left', 'right'],
ylabel = 'state (s)', xlabel = 'action (a)',
title = 'Utility')
ax_utility.xaxis.set_ticks_position('bottom')
ax_utility.spines['left'].set_visible(False)
ax_utility.spines['bottom'].set_visible(False)
# Expected utility plot details
ax_expected.set(title = 'Expected utility', ylim = [-3, 3],
xticks = [0, 1], xticklabels = ['left', 'right'],
xlabel = 'action (a)',
yticks = [])
ax_expected.xaxis.set_ticks_position('bottom')
ax_expected.spines['left'].set_visible(False)
ax_expected.spines['bottom'].set_visible(False)
# show values
ind = np.arange(2)
x,y = np.meshgrid(ind,ind)
for i,j in zip(x.flatten(), y.flatten()):
c = f"{utility[i,j]:.2f}"
ax_utility.text(j,i, c, va='center', ha='center', color='black')
for i in ind:
v = prior[i]
c = f"{v:.2f}"
ax_prior.text(v+0.2, i, c, va='center', ha='center', color='black')
for i in ind:
v = expected[i]
c = f"{v:.2f}"
ax_expected.text(i, 2.5, c, va='center', ha='center', color='black')
return fig
def plot_prior_likelihood_utility(ps, p_a_s1, p_a_s0,measurement):
assert 0.0 <= ps <= 1.0
assert 0.0 <= p_a_s1 <= 1.0
assert 0.0 <= p_a_s0 <= 1.0
prior = np.asarray([ps, 1 - ps])
likelihood = np.asarray([[p_a_s1, 1-p_a_s1],[p_a_s0, 1-p_a_s0]])
utility = np.array([[2.0, -3.0], [-2.0, 1.0]])
# expected = np.zeros_like(utility)
if measurement:
posterior = likelihood[:, 0] * prior
else:
posterior = (likelihood[:, 1] * prior).reshape(-1)
posterior /= np.sum(posterior)
# expected[:, 0] = utility[:, 0] * posterior
# expected[:, 1] = utility[:, 1] * posterior
expected = posterior @ utility
# definitions for the axes
left, width = 0.05, 0.15
bottom, height = 0.05, 0.9
padding = 0.05
small_width = 0.1
large_padding = 0.07
left_space = left + small_width + large_padding
fig = plt.figure(figsize=(17, 4))
rect_prior = [left, bottom+0.05, small_width, height-0.1]
rect_likelihood = [left_space, bottom , width, height]
rect_posterior = [left_space + padding + width - 0.02, bottom+0.05 , small_width, height-0.1]
rect_utility = [left_space + padding + width + padding + small_width, bottom , width, height]
rect_expected = [left_space + padding + width + padding + small_width + padding + width, bottom+0.05 , width, height-0.1]
ax_likelihood = fig.add_axes(rect_likelihood)
ax_prior = fig.add_axes(rect_prior, sharey=ax_likelihood)
ax_posterior = fig.add_axes(rect_posterior, sharey=ax_likelihood)
ax_utility = fig.add_axes(rect_utility, sharey=ax_posterior)
ax_expected = fig.add_axes(rect_expected)
prior_colormap = plt.cm.Blues
posterior_colormap = plt.cm.Greens
expected_colormap = plt.cm.Wistia
# Show posterior probs and marginals
ax_prior.barh(0, prior[0], facecolor = prior_colormap(prior[0]))
ax_prior.barh(1, prior[1], facecolor = prior_colormap(prior[1]))
ax_likelihood.matshow(likelihood, vmin=0., vmax=1., cmap='Reds')
ax_posterior.barh(0, posterior[0], facecolor = posterior_colormap(posterior[0]))
ax_posterior.barh(1, posterior[1], facecolor = posterior_colormap(posterior[1]))
ax_utility.matshow(utility, vmin=0., vmax=1., cmap='cool')
# ax_expected.matshow(expected, vmin=0., vmax=1., cmap='Wistia')
ax_expected.bar(0, expected[0], facecolor = expected_colormap(expected[0]))
ax_expected.bar(1, expected[1], facecolor = expected_colormap(expected[1]))
# Probabilities plot details
ax_prior.set(xlim = [1, 0], yticks = [0, 1], yticklabels = ['left', 'right'],
ylabel = 'state (s)', title = "Prior p(s)")
ax_prior.axis('off')
# Likelihood plot details
ax_likelihood.set(xticks = [0, 1], xticklabels = ['fish', 'no fish'],
yticks = [0, 1], yticklabels = ['left', 'right'],
ylabel = 'state (s)', xlabel = 'measurement (m)',
title = 'Likelihood p(m | s)')
ax_likelihood.xaxis.set_ticks_position('bottom')
ax_likelihood.spines['left'].set_visible(False)
ax_likelihood.spines['bottom'].set_visible(False)
# Posterior plot details
ax_posterior.set(xlim = [0, 1], yticks = [0, 1], yticklabels = ['left', 'right'],
ylabel = 'state (s)', title = "Posterior p(s | m)")
ax_posterior.axis('off')
# Utility plot details
ax_utility.set(xticks = [0, 1], xticklabels = ['left', 'right'],
xlabel = 'action (a)',
title = 'Utility')
ax_utility.xaxis.set_ticks_position('bottom')
ax_utility.spines['left'].set_visible(False)
ax_utility.spines['bottom'].set_visible(False)
# Expected Utility plot details
ax_expected.set(ylim = [-2, 2], xticks = [0, 1], xticklabels = ['left', 'right'],
xlabel = 'action (a)', title = 'Expected utility', yticks=[])
# ax_expected.axis('off')
ax_expected.spines['left'].set_visible(False)
# ax_expected.set(xticks = [0, 1], xticklabels = ['left', 'right'],
# xlabel = 'action (a)',
# title = 'Expected utility')
# ax_expected.xaxis.set_ticks_position('bottom')
# ax_expected.spines['left'].set_visible(False)
# ax_expected.spines['bottom'].set_visible(False)
# show values
ind = np.arange(2)
x,y = np.meshgrid(ind,ind)
for i in ind:
v = posterior[i]
c = f"{v:.2f}"
ax_posterior.text(v+0.2, i, c, va='center', ha='center', color='black')
for i,j in zip(x.flatten(), y.flatten()):
c = f"{likelihood[i,j]:.2f}"
ax_likelihood.text(j,i, c, va='center', ha='center', color='black')
for i,j in zip(x.flatten(), y.flatten()):
c = f"{utility[i,j]:.2f}"
ax_utility.text(j,i, c, va='center', ha='center', color='black')
# for i,j in zip(x.flatten(), y.flatten()):
# c = f"{expected[i,j]:.2f}"
# ax_expected.text(j,i, c, va='center', ha='center', color='black')
for i in ind:
v = prior[i]
c = f"{v:.2f}"
ax_prior.text(v+0.2, i, c, va='center', ha='center', color='black')
for i in ind:
v = expected[i]
c = f"{v:.2f}"
ax_expected.text(i, v, c, va='center', ha='center', color='black')
# # show values
# ind = np.arange(2)
# x,y = np.meshgrid(ind,ind)
# for i,j in zip(x.flatten(), y.flatten()):
# c = f"{P[i,j]:.2f}"
# ax.text(j,i, c, va='center', ha='center', color='white')
# for i in ind:
# v = marginal_x[i]
# c = f"{v:.2f}"
# ax_x.text(i, v +0.2, c, va='center', ha='center', color='black')
# v = marginal_y[i]
# c = f"{v:.2f}"
# ax_y.text(v+0.2, i, c, va='center', ha='center', color='black')
return fig
# + cellView="form"
# @title Helper Functions
def compute_marginal(px, py, cor):
# calculate 2x2 joint probabilities given marginals p(x=1), p(y=1) and correlation
p11 = px*py + cor*np.sqrt(px*py*(1-px)*(1-py))
p01 = px - p11
p10 = py - p11
p00 = 1.0 - p11 - p01 - p10
return np.asarray([[p00, p01], [p10, p11]])
# test
# print(compute_marginal(0.4, 0.6, -0.8))
def compute_cor_range(px,py):
# Calculate the allowed range of correlation values given marginals p(x=1) and p(y=1)
def p11(corr):
return px*py + corr*np.sqrt(px*py*(1-px)*(1-py))
def p01(corr):
return px - p11(corr)
def p10(corr):
return py - p11(corr)
def p00(corr):
return 1.0 - p11(corr) - p01(corr) - p10(corr)
Cmax = min(fsolve(p01, 0.0), fsolve(p10, 0.0))
Cmin = max(fsolve(p11, 0.0), fsolve(p00, 0.0))
return Cmin, Cmax
# -
# ---
# # Section 1: <NAME>'
#
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 517} outputId="2209f6d0-af50-45f2-fc41-3117db3246dd"
#@title Video 2: <NAME>'
from IPython.display import YouTubeVideo
video = YouTubeVideo(id='McALsTzb494', width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
# -
# You were just introduced to the **binary hidden state problem** we are going to explore. You need to decide which side to fish on. We know fish like to school together. On different days the school of fish is either on the left or right side, but we don’t know what the case is today. We will represent our knowledge probabilistically, asking how to make a decision (where to decide the fish are or where to fish) and what to expect in terms of gains or losses. In the next two sections we will consider just the probability of where the fish might be and what you gain or lose by choosing where to fish.
#
# Remember, you can either think of your self as a scientist conducting an experiment or as a brain trying to make a decision. The Bayesian approach is the same!
#
# ---
# # Section 2: Deciding where to fish
#
#
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 517} outputId="5fe78511-352c-4122-80d3-6d8ad232cc95"
#@title Video 3: Utility
from IPython.display import YouTubeVideo
video = YouTubeVideo(id='xvIVZrqF_5s', width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
# -
# You know the probability that the school of fish is on the left side of the dock today, $P(s = left)$. You also know the probability that it is on the right, $P(s = right)$, because these two probabilities must add up to 1. You need to decide where to fish. It may seem obvious - you could just fish on the side where the probability of the fish being is higher! Unfortunately, decisions and actions are always a little more complicated. Deciding to fish may be influenced by more than just the probability of the school of fish being there as we saw by the potential issues of submarines and sunburn.
#
# We quantify these factors numerically using **utility**, which describes the consequences of your actions: how much value you gain (or if negative, lose) given the state of the world ($s$) and the action you take ($a$). In our example, our utility can be summarized as:
#
# | Utility: U(s,a) | a = left | a = right |
# | ----------------- |----------|----------|
# | s = Left | 2 | -3 |
# | s = right | -2 | 1 |
#
# To use utility to choose an action, we calculate the **expected utility** of that action by weighing these utilities with the probability of that state occuring. This allows us to choose actions by taking probabilities of events into account: we don't care if the outcome of an action-state pair is a loss if the probability of that state is very low. We can formalize this as:
#
# $$\text{Expected utility of action a} = \sum_{s}U(s,a)P(s) $$
#
# In other words, the expected utility of an action a is the sum over possible states of the utility of that action and state times the probability of that state.
#
# ## Interactive Demo 2: Exploring the decision
#
# Let's start to get a sense of how all this works.
#
# Take a look at the interactive demo below. You can change the probability that the school of fish is on the left side ($p(s = left)$ using the slider. You will see the utility matrix and the corresponding expected utility of each action.
#
# First, make sure you understand how the expected utility of each action is being computed from the probabilities and the utility values. In the initial state: the probability of the fish being on the left is 0.9 and on the right is 0.1. The expected utility of the action of fishing on the left is then $U(s = left,a = left)p(s = left) + U(s = right,a = left)p(s = right) = 2(0.9) + -2(0.1) = 1.6$.
#
# For each of these scenarios, think and discuss first. Then use the demo to try out each and see if your action would have been correct (that is, if the expected value of that action is the highest).
#
#
# 1. You just arrived at the dock for the first time and have no sense of where the fish might be. So you guess that the probability of the school being on the left side is 0.5 (so the probability on the right side is also 0.5). Which side would you choose to fish on given our utility values?
# 2. You think that the probability of the school being on the left side is very low (0.1) and correspondingly high on the right side (0.9). Which side would you choose to fish on given our utility values?
# 3. What would you choose if the probability of the school being on the left side is slightly lower than on the right side (0. 4 vs 0.6)?
# + cellView="form"
# @markdown Execute this cell to use the widget
ps_widget = widgets.FloatSlider(0.9, description='p(s = left)', min=0.0, max=1.0, step=0.01)
@widgets.interact(
ps = ps_widget,
)
def make_utility_plot(ps):
fig = plot_utility(ps)
plt.show(fig)
plt.close(fig)
return None
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D1_BayesianDecisions/solutions/W3D1_Tutorial1_Solution_459cbf35.py)
#
#
# -
# In this section, you have seen that both the utility of various state and action pairs and our knowledge of the probability of each state affects your decision. Importantly, we want our knowledge of the probability of each state to be as accurate as possible!
#
# So how do we know these probabilities? We may have prior knowledge from years of fishing at the same dock. Over those years, we may have learned that the fish are more likely to be on the left side for example. We want to make sure this knowledge is as accurate as possible though. To do this, we want to collect more data, or take some more measurements! For the next few sections, we will focus on making our knowledge of the probability as accurate as possible, before coming back to using utility to make decisions.
# ---
# # Section 3: Likelihood of the fish being on either side
#
#
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 517} outputId="ce7995af-5602-4cb3-b403-3f759d1bb0af"
#@title Video 4: Likelihood
from IPython.display import YouTubeVideo
video = YouTubeVideo(id='l4m0JzMWGio', width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
# -
# First, we'll think about what it means to take a measurement (also often called an observation or just data) and what it tells you about what the hidden state may be. Specifically, we'll be looking at the **likelihood**, which is the probability of your measurement ($m$) given the hidden state ($s$): $P(m | s)$. Remember that in this case, the hidden state is which side of the dock the school of fish is on.
#
# We will watch someone fish (for let's say 10 minutes) and our measurement is whether they catch a fish or not. We know something about what catching a fish means for the likelihood of the fish being on one side or the other.
# ## Think! 3: Guessing the location of the fish
#
# Let's say we go to different dock from the one in the video. Here, there are different probabilities of catching fish given the state of the world. In this case, if they fish on the side of the dock where the fish are, they have a 70% chance of catching a fish. Otherwise, they catch a fish with only 20% probability.
#
# The fisherperson is fishing on the left side.
#
# 1) Figure out each of the following:
# - probability of catching a fish given that the school of fish is on the left side, $P(m = catch\text{ } fish | s = left )$
# - probability of not catching a fish given that the school of fish is on the left side, $P(m = no \text{ } fish | s = left)$
# - probability of catching a fish given that the school of fish is on the right side, $P(m = catch \text{ } fish | s = right)$
# - probability of not catching a fish given that the school of fish is on the right side, $P(m = no \text{ } fish | s = right)$
#
# 2) If the fisherperson catches a fish, which side would you guess the school is on? Why?
#
# 3) If the fisherperson does not catch a fish, which side would you guess the school is on? Why?
#
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D1_BayesianDecisions/solutions/W3D1_Tutorial1_Solution_2284aaf4.py)
#
#
# -
# In the prior exercise, you guessed where the school of fish was based on the measurement you took (watching someone fish). You did this by choosing the state (side of school) that maximized the probability of the measurement. In other words, you estimated the state by maximizing the likelihood (had the highest probability of measurement given state $P(m|s$)). This is called maximum likelihood estimation (MLE) and you've encountered it before during this course, in W1D3!
#
# What if you had been going to this river for years and you knew that the fish were almost always on the left side? This would probably affect how you make your estimate - you would rely less on the single new measurement and more on your prior knowledge. This is the idea behind Bayesian inference, as we will see later in this tutorial!
# ---
# # Section 4: Correlation and marginalization
#
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 517} outputId="e4c1eecf-211f-4836-e4ed-aa45c607d0cd"
#@title Video 5: Correlation and marginalization
from IPython.display import YouTubeVideo
video = YouTubeVideo(id='vsDjtWi-BVo', width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
# -
# In this section, we are going to take a step back for a bit and think more generally about the amount of information shared between two random variables. We want to know how much information you gain when you observe one variable (take a measurement) if you know something about another. We will see that the fundamental concept is the same if we think about two attributes, for example the size and color of the fish, or the prior information and the likelihood.
# ## Math Exercise 4: Computing marginal likelihoods
#
# To understand the information between two variables, let's first consider the size and color of the fish.
#
# | P(X, Y) | Y = silver | Y = gold |
# | ----------------- |----------|----------|
# | X = small | 0.4 | 0.2 |
# | X = large | 0.1 | 0.3 |
#
# The table above shows us the **joint probabilities**: the probability of both specific attributes occuring together. For example, the probability of a fish being small and silver ($P(X = small, Y = silver$) is 0.4.
#
# We want to know what the probability of a fish being small regardless of color. Since the fish are either silver or gold, this would be the probability of a fish being small and silver plus the probability of a fish being small and gold. This is an example of marginalizing, or averaging out, the variable we are not interested in across the rows or columns.. In math speak: $P(X = small) = \sum_y{P(X = small, Y)}$. This gives us a **marginal probability**, a probability of a variable outcome (in this case size), regardless of the other variables (in this case color).
#
# Please complete the following math problems to further practice thinking through probabilities:
#
# 1. Calculate the probability of a fish being silver.
# 2. Calculate the probability of a fish being small, large, silver, or gold.
# 3. Calculate the probability of a fish being small OR gold. (Hint: $P(A\ \textrm{or}\ B) = P(A) + P(B) - P(A\ \textrm{and}\ B)$)
#
#
#
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D1_BayesianDecisions/solutions/W3D1_Tutorial1_Solution_65e69cd1.py)
#
#
# -
# ## Think! 4: Covarying probability distributions
#
# The relationship between the marginal probabilities and the joint probabilities is determined by the correlation between the two random variables - a normalized measure of how much the variables covary. We can also think of this as gaining some information about one of the variables when we observe a measurement from the other. We will think about this more formally in Tutorial 2.
#
# Here, we want to think about how the correlation between size and color of these fish changes how much information we gain about one attribute based on the other. See Bonus Section 1 for the formula for correlation.
#
# Use the widget below and answer the following questions:
#
# 1. When the correlation is zero, $\rho = 0$, what does the distribution of size tell you about color?
# 2. Set $\rho$ to something small. As you change the probability of golden fish, what happens to the ratio of size probabilities? Set $\rho$ larger (can be negative). Can you explain the pattern of changes in the probabilities of size as you change the probability of golden fish?
# 3. Set the probability of golden fish and of large fish to around 65%. As the correlation goes towards 1, how often will you see silver large fish?
# 4. What is increasing the (absolute) correlation telling you about how likely you are to see one of the properties if you see a fish with the other?
#
# + cellView="form"
# @markdown Execute this cell to enable the widget
style = {'description_width': 'initial'}
gs = GridspecLayout(2,2)
cor_widget = widgets.FloatSlider(0.0, description='ρ', min=-1, max=1, step=0.01)
px_widget = widgets.FloatSlider(0.5, description='p(color=golden)', min=0.01, max=0.99, step=0.01, style=style)
py_widget = widgets.FloatSlider(0.5, description='p(size=large)', min=0.01, max=0.99, step=0.01, style=style)
gs[0,0] = cor_widget
gs[0,1] = px_widget
gs[1,0] = py_widget
@widgets.interact(
px=px_widget,
py=py_widget,
cor=cor_widget,
)
def make_corr_plot(px, py, cor):
Cmin, Cmax = compute_cor_range(px, py) #allow correlation values
cor_widget.min, cor_widget.max = Cmin+0.01, Cmax-0.01
if cor_widget.value > Cmax:
cor_widget.value = Cmax
if cor_widget.value < Cmin:
cor_widget.value = Cmin
cor = cor_widget.value
P = compute_marginal(px,py,cor)
# print(P)
fig = plot_joint_probs(P)
plt.show(fig)
plt.close(fig)
return None
# gs[1,1] = make_corr_plot()
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D1_BayesianDecisions/solutions/W3D1_Tutorial1_Solution_9727d8f3.py)
#
#
# -
# We have just seen how two random variables can be more or less independent. The more correlated, the less independent, and the more shared information. We also learned that we can marginalize to determine the marginal likelihood of a hidden state or to find the marginal probability distribution of two random variables. We are going to now complete our journey towards being fully Bayesian!
# ---
# # Section 5: Bayes' Rule and the Posterior
# Marginalization is going to be used to combine our prior knowlege, which we call the **prior**, and our new information from a measurement, the **likelihood**. Only in this case, the information we gain about the hidden state we are interested in, where the fish are, is based on the relationship between the probabilities of the measurement and our prior.
#
# We can now calculate the full posterior distribution for the hidden state ($s$) using Bayes' Rule. As we've seen, the posterior is proportional the the prior times the likelihood. This means that the posterior probability of the hidden state ($s$) given a measurement ($m$) is proportional to the likelihood of the measurement given the state times the prior probability of that state (the marginal likelihood):
#
# $$ P(s | m) \propto P(m | s) P(s) $$
#
# We say proportional to instead of equal because we need to normalize to produce a full probability distribution:
#
# $$ P(s | m) = \frac{P(m | s) P(s)}{P(m)} $$
#
# Normalizing by this $P(m)$ means that our posterior is a complete probability distribution that sums or integrates to 1 appropriately. We now can use this new, complete probability distribution for any future inference or decisions we like! In fact, as we will see tomorrow, we can use it as a new prior! Finally, we often call this probability distribution our beliefs over the hidden states, to emphasize that it is our subjective knowlege about the hidden state.
#
# For many complicated cases, like those we might be using to model behavioral or brain inferences, the normalization term can be intractable or extremely complex to calculate. We can be careful to choose probability distributions were we can analytically calculate the posterior probability or numerical approximation is reliable. Better yet, we sometimes don't need to bother with this normalization! The normalization term, $P(m)$, is the probability of the measurement. This does not depend on state so is essentially a constant we can often ignore. We can compare the unnormalized posterior distribution values for different states because how they relate to each other is unchanged when divided by the same constant. We will see how to do this to compare evidence for different hypotheses tomorrow. (It's also used to compare the likelihood of models fit using maximum likelihood estimation, as you did in W1D5.)
#
# In this relatively simple example, we can compute the marginal probability $P(m)$ easily by using:
# $$P(m) = \sum_s P(m | s) P(s)$$
# We can then normalize so that we deal with the full posterior distribution.
#
# ## Math Exercise 5: Calculating a posterior probability
#
# Our prior is $p(s = left) = 0.3$ and $p(s = right) = 0.7$. In the video, we learned that the chance of catching a fish given they fish on the same side as the school was 50%. Otherwise, it was 10%. We observe a person fishing on the left side. Our likelihood is:
#
#
# | Likelihood: p(m \| s) | m = catch fish | m = no fish |
# | ----------------- |----------|----------|
# | s = left | 0.5 | 0.5 |
# | s = right | 0.1 | 0.9 |
#
#
# Calculate the posterior probability (on paper) that:
#
# 1. The school is on the left if the fisherperson catches a fish: $p(s = left | m = catch fish)$ (hint: normalize by compute $p(m = catch fish)$)
# 2. The school is on the right if the fisherperson does not catch a fish: $p(s = right | m = no fish)$
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D1_BayesianDecisions/solutions/W3D1_Tutorial1_Solution_46a5b352.py)
#
#
# -
# ## Coding Exercise 5: Computing Posteriors
#
# Let's implement our above math to be able to compute posteriors for different priors and likelihood.s
#
# As before, our prior is $p(s = left) = 0.3$ and $p(s = right) = 0.7$. In the video, we learned that the chance of catching a fish given they fish on the same side as the school was 50%. Otherwise, it was 10%. We observe a person fishing on the left side. Our likelihood is:
#
#
# | Likelihood: p(m \| s) | m = catch fish | m = no fish |
# | ----------------- |----------|----------|
# | s = left | 0.5 | 0.5 |
# | s = right | 0.1 | 0.9 |
#
#
# We want our full posterior to take the same 2 by 2 form. Make sure the outputs match your math answers!
#
#
# +
def compute_posterior(likelihood, prior):
""" Use Bayes' Rule to compute posterior from likelihood and prior
Args:
likelihood (ndarray): i x j array with likelihood probabilities where i is
number of state options, j is number of measurement options
prior (ndarray): i x 1 array with prior probability of each state
Returns:
ndarray: i x j array with posterior probabilities where i is
number of state options, j is number of measurement options
"""
#################################################
## TODO for students ##
# Fill out function and remove
raise NotImplementedError("Student exercise: implement compute_posterior")
#################################################
# Compute unnormalized posterior (likelihood times prior)
posterior = ... # first row is s = left, second row is s = right
# Compute p(m)
p_m = np.sum(posterior, axis = 0)
# Normalize posterior (divide elements by p_m)
posterior /= ...
return posterior
# Make prior
prior = np.array([0.3, 0.7]).reshape((2, 1)) # first row is s = left, second row is s = right
# Make likelihood
likelihood = np.array([[0.5, 0.5], [0.1, 0.9]]) # first row is s = left, second row is s = right
# Compute posterior
posterior = compute_posterior(likelihood, prior)
# Visualize
with plt.xkcd():
plot_prior_likelihood_posterior(prior, likelihood, posterior)
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D1_BayesianDecisions/solutions/W3D1_Tutorial1_Solution_042b00b7.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=669 height=314 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W3D1_BayesianDecisions/static/W3D1_Tutorial1_Solution_042b00b7_0.png>
#
#
# -
# ## Interactive Demo 5: What affects the posterior?
#
# Now that we can understand the implementation of *Bayes rule*, let's vary the parameters of the prior and likelihood to see how changing the prior and likelihood affect the posterior.
#
# In the demo below, you can change the prior by playing with the slider for $p( s = left)$. You can also change the likelihood by changing the probability of catching a fish given that the school is on the left and the probability of catching a fish given that the school is on the right. The fisherperson you are observing is fishing on the left.
#
#
# 1. Keeping the likelihood constant, when does the prior have the strongest influence over the posterior? Meaning, when does the posterior look most like the prior no matter whether a fish was caught or not?
# 2. Keeping the likelihood constant, when does the prior exert the weakest influence? Meaning, when does the posterior look least like the prior and depend most on whether a fish was caught or not?
# 3. Set the prior probability of the state = left to 0.6 and play with the likelihood. When does the likelihood exert the most influence over the posterior?
# + cellView="form"
# @markdown Execute this cell to enable the widget
style = {'description_width': 'initial'}
ps_widget = widgets.FloatSlider(0.3, description='p(s = left)',
min=0.01, max=0.99, step=0.01)
p_a_s1_widget = widgets.FloatSlider(0.5, description='p(fish | s = left)',
min=0.01, max=0.99, step=0.01, style=style)
p_a_s0_widget = widgets.FloatSlider(0.1, description='p(fish | s = right)',
min=0.01, max=0.99, step=0.01, style=style)
observed_widget = widgets.Checkbox(value=False, description='Observed fish (m)',
disabled=False, indent=False, layout={'width': 'max-content'})
@widgets.interact(
ps=ps_widget,
p_a_s1=p_a_s1_widget,
p_a_s0=p_a_s0_widget,
m_right=observed_widget
)
def make_prior_likelihood_plot(ps,p_a_s1,p_a_s0,m_right):
fig = plot_prior_likelihood(ps,p_a_s1,p_a_s0,m_right)
plt.show(fig)
plt.close(fig)
return None
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D1_BayesianDecisions/solutions/W3D1_Tutorial1_Solution_a004d68d.py)
#
#
# -
# # Section 6: Making Bayesian fishing decisions
#
# We will explore how to consider the expected utility of an action based on our belief (the posterior distribution) about where we think the fish are. Now we have all the components of a Bayesian decision: our prior information, the likelihood given a measurement, the posterior distribution (belief) and our utility (the gains and losses). This allows us to consider the relationship between the true value of the hidden state, $s$, and what we *expect* to get if we take action, $a$, based on our belief!
#
# Let's use the following widget to think about the relationship between these probability distributions and utility function.
# ## Think! 6: What is more important, the probabilities or the utilities?
#
# We are now going to put everything we've learned together to gain some intuitions for how each of the elements that goes into a Bayesian decision comes together. Remember, the common assumption in neuroscience, psychology, economics, ecology, etc. is that we (humans and animals) are tying to maximize our expected utility.
#
# 1. Can you find a situation where the expected utility is the same for both actions?
# 2. What is more important for determining the expected utility: the prior or a new measurement (the likelihood)?
# 3. Why is this a normative model?
# 4. Can you think of ways in which this model would need to be extended to describe human or animal behavior?
# + cellView="form"
# @markdown Execute this cell to enable the widget
style = {'description_width': 'initial'}
ps_widget = widgets.FloatSlider(0.3, description='p(s)',
min=0.01, max=0.99, step=0.01)
p_a_s1_widget = widgets.FloatSlider(0.5, description='p(fish | s = left)',
min=0.01, max=0.99, step=0.01, style=style)
p_a_s0_widget = widgets.FloatSlider(0.1, description='p(fish | s = right)',
min=0.01, max=0.99, step=0.01, style=style)
observed_widget = widgets.Checkbox(value=False, description='Observed fish (m)',
disabled=False, indent=False, layout={'width': 'max-content'})
@widgets.interact(
ps=ps_widget,
p_a_s1=p_a_s1_widget,
p_a_s0=p_a_s0_widget,
m_right=observed_widget
)
def make_prior_likelihood_utility_plot(ps, p_a_s1, p_a_s0,m_right):
fig = plot_prior_likelihood_utility(ps, p_a_s1, p_a_s0,m_right)
plt.show(fig)
plt.close(fig)
return None
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D1_BayesianDecisions/solutions/W3D1_Tutorial1_Solution_3a230382.py)
#
#
# -
# ---
# # Summary
#
# In this tutorial, you learned about combining prior information with new measurements to update your knowledge using Bayes Rulem, in the context of a fishing problem.
#
# Specifically, we covered:
#
# * That the likelihood is the probability of the measurement given some hidden state
#
# * That how the prior and likelihood interact to create the posterior, the probability of the hidden state given a measurement, depends on how they covary
#
# * That utility is the gain from each action and state pair, and the expected utility for an action is the sum of the utility for all state pairs, weighted by the probability of that state happening. You can then choose the action with highest expected utility.
#
# ---
# # Bonus
# ## Bonus Section 1: Correlation Formula
# To understand the way we calculate the correlation, we need to review the definition of covariance and correlation.
#
# Covariance:
#
# $$
# cov(X,Y) = \sigma_{XY} = E[(X - \mu_{x})(Y - \mu_{y})] = E[X]E[Y] - \mu_{x}\mu_{y}
# $$
#
# Correlation:
#
# $$
# \rho_{XY} = \frac{cov(Y,Y)}{\sqrt{V(X)V(Y)}} = \frac{\sigma_{XY}}{\sigma_{X}\sigma_{Y}}
# $$
| tutorials/W3D1_BayesianDecisions/student/W3D1_Tutorial1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# +
#使用字典创建一个DataFrame。
d = {'国家': ['中国', '美国', '日本'],
'人口': [14.22, 3.18, 1.29]}
df = pd.DataFrame(d)
df
# +
#利用索引,添加一行数据。
df.loc[3] = {'国家': '俄罗斯', '人口': 1.4}
df
# +
#创建另一个DataFrame。
df2 = pd.DataFrame({'国家': ['英国', '德国'],
'人口': [0.66, 0.82]})
#利用DataFrame的内建函数append,追加一个DataFrame(多条数据)。
new_df = df.append(df2)
new_df
# +
#使用pandas的concat功能,拼接两个DataFrame,同时重置索引。
new_df = pd.concat([df, df2], ignore_index=True)
new_df
| Chapter_4/Section_4.4.1.1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/123Edward/CPEN-2IA-ECE-2-1/blob/main/Midterm_Exam.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="v6UuWs-G3DCv"
# #Midterm Exam
# + [markdown] id="ayYJRbUxCL9a"
# ##Problem Statement 1
# + colab={"base_uri": "https://localhost:8080/"} id="Rl07AbQt3ce_" outputId="a881d566-57f7-4ce2-9b6f-ef5e7b475dae"
a="<NAME>"
b="202012046"
c="20 years old"
d="March 09, 2001"
e="Purok 21, Dap-dap West, Tagaytay City, Cavite"
f="Programming Logic and Design"
g="1.6"
print(a)
print(b)
print(c)
print(d)
print(e)
print(f)
print(g)
# + [markdown] id="9VOVdla1CS4a"
# ##Problem Statement 2
# + colab={"base_uri": "https://localhost:8080/"} id="DAY7Xe0L6qdh" outputId="269fbe5b-084d-4297-d206-04aeaeb1568e"
n=4
answ= "Y"
a=(2<n) and (n<6)
b=(2<n) or (n==6)
c= not (2<n) or (n==6)
d= not (n<6)
e=(answ=="Y") or (answ=="y")
f=(answ=="Y") and (answ=="y")
g= not (answ=="y")
h= ((2<n) and (n==5+1)) or (answ=="no")
i=((n==2) and (n==7)) or (answ=="Y")
j=(n==2) and ((n==7) or (answ=="Y"))
print(a)
print(b)
print(c)
print(d)
print(e)
print(f)
print(g)
print(h)
print(i)
print(j)
# + [markdown] id="ND26LqAACX5z"
# ##Problem Statement 3
# + colab={"base_uri": "https://localhost:8080/"} id="Vcgq4HVgCcML" outputId="a90a9de1-0a83-49da-95f2-bc7fa763c461"
x=2
y=-3
w=7
z=-10
print(x/y)
print(w/y/x)
print(z/y%x)
print(x%-y*w)
print(x%y)
print(z%w-y/x*5+5)
print(9-x%(2+y))
print(z//w)
print((2+y)**2)
print(w/x*2)
| Midterm_Exam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: python-zqp4N2IQ
# language: python
# name: python-zqp4n2iq
# ---
# # Multiples of 3 and 5
# ### Problem 1
#
# If we list all the natural numbers below 10 that are multiples of 3 or 5, we get 3, 5, 6 and 9. The sum of these multiples is 23.
#
# Find the sum of all the multiples of 3 or 5 below 1000.
# +
# sample
s = 0
for i in range(1, 10):
if i % 3 == 0 or i % 5 == 0:
s += i
s
# +
# solution
s = 0
for i in range(1, 1000):
if i % 3 == 0 or i % 5 == 0:
s += i
s
| pe-solution/src/main/python/PEP_001 Multiples of 3 and 5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from pycalphad import equilibrium
from pycalphad.core.equilibrium import _compute_constraints
from pycalphad.core.custom_ufuncify import UfuncifyCodeWrapper, ufuncify
from sympy.printing.ccode import CCodePrinter, ccode
from pycalphad import Database, Model
from sympy.printing import ccode
import pycalphad.variables as v
db_alfe = Database('alfe_sei.TDB')
my_phases_alfe = ['LIQUID', 'HCP_A3', 'AL5FE2', 'AL2FE', 'AL5FE4', 'FCC_A1', 'B2_BCC', 'AL13FE4']
temp = 600
# %time eq= equilibrium(db_alfe, ['AL', 'FE', 'VA'], my_phases_alfe, {v.X('AL'): (0,1,0.05), v.T: 600, v.P: 101325})
# +
# %matplotlib inline
import matplotlib.pyplot as plt
from pycalphad.plot.utils import phase_legend
temp = 600
plt.gca().set_xlim((0,1))
plt.gca().set_title('Chemical potential of Fe vs X(AL), {} K'.format(temp))
plt.gca().set_xlabel('X(AL)')
plt.gca().set_ylabel('MU(FE)')
phase_handles, phasemap = phase_legend(my_phases_alfe)
phasecolors = [phasemap[str(p)] for p in eq.Phase.sel(T=temp, vertex=0).values[0] if p != '']
plt.scatter(eq.X.sel(T=temp, component='AL', vertex=0), eq.MU.sel(T=temp, component='FE'), color=phasecolors)
phasecolors = [phasemap[str(p)] for p in eq.Phase.sel(T=temp, vertex=1).values[0] if p != '']
plt.scatter(eq.X.sel(T=temp, component='AL', vertex=1), eq.MU.sel(T=temp, component='FE'), color=phasecolors)
plt.gca().legend(phase_handles, my_phases_alfe, loc='lower left')
# -
| Equilibrium2Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def free_colors(v, u_colors, col_graph):
n_colors = []
for n in col_graph[v]["neighbors"]:
c_n = col_graph[n]["color"]
if c_n != 0:
n_colors.append(c_n)
f_colors = [c for c in u_colors if c not in n_colors]
return f_colors
def graph_coloring_seq(graph, seq):
# Cor igual a 0 representa vértice ainda não colorido
col_graph = {v: {"neighbors": graph[v], "color": 0} for v in graph.keys()}
# Começando com a cor 1
used_colors = [1]
for v in seq:
print("Fazendo v:", v)
f_colors = free_colors(v, used_colors, col_graph)
if len(f_colors) != 0:
print("Pode usar a cor:", min(f_colors))
col_graph[v]["color"] = min(f_colors)
else:
print("Tem que criar uma nova:", max(used_colors)+1)
new_color = max(used_colors) + 1
col_graph[v]["color"] = new_color
used_colors.append(new_color)
print()
return {v: col_graph[v]["color"] for v in seq}
exemplo = {
"S": {"I", "H", "C", "L"},
"I": {"S", "L", "M"},
"H": {"S", "L", "G", "A"},
"L": {"G", "P", "H", "S", "I", "M"},
"P": {"G", "L"},
"G": {"A", "P", "H", "L"},
"C": {"S", "A"},
"A": {"C", "H", "G", "M"},
"M": {"I", "L", "G", "A"}
}
seq_ex = ["G", "L", "H", "P", "M", "A", "I", "S", "C"]
graph_coloring_seq(exemplo, seq_ex)
| notebooks/e2/greedy_graph_coloring.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import utils
# +
import pandas as pd
import jinja2
from collections import OrderedDict
from json import dumps
from IPython.display import display, Javascript, HTML
# -
graph = utils.pickle_from_file('../../data/yago_hierarchy.pickle')
sorted_nodes = sorted(graph.nodes())
nodes = [{
'name': ' '.join(str(x).replace('wordnet_', '').split('_')[:-1]),
'root': len(graph.predecessors(x))
} for x in sorted_nodes]
links = [{
'source': sorted_nodes.index(source),
'target': sorted_nodes.index(target)
} for source, target in graph.edges()]
# + language="javascript"
# require.config({
# paths: {
# d3: '//cdnjs.cloudflare.com/ajax/libs/d3/3.4.8/d3.min'
# }
# });
# -
d3_template = jinja2.Template(
"""
// Based on http://bl.ocks.org/mbostock/3885304
require(["d3"], function(d3) {
var graph = {
'nodes': {{ nodes }},
'links': {{ links }}
};
var zoom = d3.behavior.zoom()
.scaleExtent([-10, 10])
.on("zoom", zoomed);
d3.select("#chart_d3 svg").remove();
var width = 960,
height = 500;
var force = d3.layout.force()
.size([width, height])
.charge(-200)
.linkDistance(50)
.on("tick", tick);
var drag = force.drag()
.on("dragstart", dragstart);
var svg = d3.select("#chart_d3").append("svg")
.attr("width", width)
.attr("height", height)
.call(zoom);
var container = svg.append("g");
// build the arrow.
container.append("svg:defs").selectAll("marker")
.data(["end"]) // Different link/path types can be defined here
.enter().append("svg:marker") // This section adds in the arrows
.attr("id", String)
.attr("viewBox", "0 -5 10 10")
.attr("refX", 15)
.attr("refY", -1.5)
.attr("markerWidth", 3)
.attr("markerHeight", 3)
.attr("orient", "auto")
.append("svg:path")
.attr("d", "M0,-5L10,0L0,5");
var link = container.selectAll(".link"),
node = container.selectAll(".node");
force.nodes(graph.nodes)
.links(graph.links)
.start();
link = link.data(graph.links)
.enter().append("line")
.attr("class", "link")
.attr("marker-end", "url(#end)");;
node = node.data(graph.nodes)
.enter().append("g")
.attr("class", "node")
.classed("root-node", function(d) {return d.root == 0})
.call(force.drag);
node.append("text")
.attr("dx", -10)
.attr("dy", ".18em")
.text(function(d) { return d.name });
function tick() {
link.attr("x1", function(d) { return d.source.x; })
.attr("y1", function(d) { return d.source.y; })
.attr("x2", function(d) { return d.target.x; })
.attr("y2", function(d) { return d.target.y; });
node.attr("transform", function(d) { return "translate(" + d.x + "," + d.y + ")"; });
}
function dblclick(d) {
d3.select(this).classed("fixed", d.fixed = false);
}
function dragstart(d) {
d3.select(this).classed("fixed", d.fixed = true);
}
function zoomed() {
container.attr("transform", "translate(" + d3.event.translate + ")scale(" + d3.event.scale + ")");
}
});
"""
)
display(HTML("""
<style>
.link {
stroke: #ccc;
stroke-width: 1.5px;
}
.node {
cursor: move;
fill: #000;
}
.node.root-node {
fill: blue;
}
.node.fixed {
fill: #f00;
}
</style>
<div id="chart_d3"/>
"""))
display(Javascript(d3_template.render(nodes=nodes, links=links)))
| yago_scripts/YAGO (Sub)graph visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import pickle
from spread import *
EXP_RES_FILE = "data500network-200peers.pickle"
with open(EXP_RES_FILE, 'rb') as f:
networks, results, scenarios = pickle.load(f)
print("loaded")
# -
def small_desc(s: SpreadScenario) -> str:
if "random picker" in s.description:
picker = "random"
elif "reliability favor picker" in s.description:
picker = "rel exp weight"
elif "first picker" in s.description:
picker = "rel first"
return f"({s.spread_n_peers}, {s.spread_tick_interval}, {picker})"
def ticks_to_time(ticks):
secs = ticks * 0.5
if secs < 60:
return f"{secs}s"
else:
return f"{round(secs/60, 2)}min"
# +
from dataclasses import dataclass
import pandas as pd
@dataclass
class AggregatedColumn:
success_all: bool = True
success_ratio: float = 0.0
success_ratio_good_peers: float = 0.0
repeated_msgs_per_tick: int = 0
repeated_msgs_per_tick_good_peers: int = 0
repeated_msgs_ratio: float = 0.0
repeated_msgs_ratio_good_peers: float = 0.0
avg_end_tick: float = 0
avg_end_tick_good_peers: float = 0.0
worst_case_end_tick: int = 0
worst_case_end_tick_good_peers: int = 0
def set_no_val(col: AggregatedColumn):
col.repeated_msgs_per_tick = -1
col.repeated_msgs_ratio = -1
col.repeated_msgs_ratio_good_peers = -1
col.avg_end_tick = -1
col.avg_end_tick_good_peers = -1
def df_of(key: float) -> pd.DataFrame:
networks_results = results[key]
columns = [AggregatedColumn() for _ in scenarios]
for net_result in networks_results:
for i, result in enumerate(net_result.scenario_results):
if result.end_tick == 9999:
columns[i].success_all = False
if result.end_tick > columns[i].worst_case_end_tick:
columns[i].worst_case_end_tick = result.end_tick
if result.end_tick_good_peers > columns[i].worst_case_end_tick_good_peers:
columns[i].worst_case_end_tick_good_peers = result.end_tick_good_peers
if result.success_good_peers:
columns[i].success_ratio_good_peers += 1
if not result.success:
continue
columns[i].success_ratio += 1
columns[i].repeated_msgs_per_tick += result.repeated_msgs / result.end_tick
columns[i].repeated_msgs_per_tick_good_peers += result.repeated_msgs_good_peers / result.end_tick
columns[i].repeated_msgs_ratio += result.repeated_msgs / (len(result.spread_edges) + result.repeated_msgs)
columns[i].repeated_msgs_ratio_good_peers += result.repeated_msgs_good_peers / (result.repeated_msgs_good_peers + result.good_peers_notified)
columns[i].avg_end_tick += result.end_tick
columns[i].avg_end_tick_good_peers += result.end_tick_good_peers
for col in columns:
if col.success_ratio:
col.repeated_msgs_per_tick /= col.success_ratio
col.repeated_msgs_per_tick_good_peers /= col.success_ratio
col.repeated_msgs_ratio /= col.success_ratio
col.repeated_msgs_ratio_good_peers /= col.success_ratio
col.avg_end_tick /= col.success_ratio
col.avg_end_tick_good_peers /= col.success_ratio
else:
set_no_val(col)
col.success_ratio_good_peers /= len(networks_results)
col.success_ratio /= len(networks_results)
df = pd.DataFrame(columns) \
.round(2) \
.rename({i : small_desc(s) for (i,s) in enumerate(scenarios)}) \
return df
# +
pd.set_option('display.max_rows', None)
df0 = df_of(0.0)
df0 = df0[df0["success_all"]]
df0 = df0.drop(columns=["success_all", "success_ratio", "success_ratio_good_peers", "repeated_msgs_per_tick_good_peers", "repeated_msgs_ratio_good_peers", "avg_end_tick_good_peers", "worst_case_end_tick_good_peers"])
df0["repeated_msgs_ratio"].describe()
# -
x = df0.sort_values("repeated_msgs_per_tick", ascending=True).head(5)
x["avg_end_tick"] = x["avg_end_tick"].apply(ticks_to_time)
x["worst_case_end_tick"] = x["worst_case_end_tick"].apply(ticks_to_time)
x
x = df0.sort_values("repeated_msgs_per_tick", ascending=True).tail(5)
x["avg_end_tick"] = x["avg_end_tick"].apply(ticks_to_time)
x["worst_case_end_tick"] = x["worst_case_end_tick"].apply(ticks_to_time)
x
x = df0.sort_values("avg_end_tick", ascending=True).head(5)
x["avg_end_tick"] = x["avg_end_tick"].apply(ticks_to_time)
x["worst_case_end_tick"] = x["worst_case_end_tick"].apply(ticks_to_time)
x
x = df0.sort_values("worst_case_end_tick", ascending=True).head(5)
x["avg_end_tick"] = x["avg_end_tick"].apply(ticks_to_time)
x["worst_case_end_tick"] = x["worst_case_end_tick"].apply(ticks_to_time)
x
# +
# Malicious environments analysis in cells below:
# +
from dataclasses import dataclass
import pandas as pd
@dataclass
class AggregatedColumn:
success_all: bool = True
success_ratio: float = 0.0
success_ratio_good_peers: float = 0.0
repeated_msgs_per_tick: int = 0
repeated_msgs_per_tick_good_peers: int = 0
repeated_msgs_ratio: float = 0.0
repeated_msgs_ratio_good_peers: float = 0.0
avg_end_tick: float = 0
avg_end_tick_good_peers: float = 0.0
worst_case_end_tick: int = 0
worst_case_end_tick_good_peers: int = 0
def set_no_val(col: AggregatedColumn):
col.repeated_msgs_per_tick = -1
col.repeated_msgs_ratio = -1
col.repeated_msgs_ratio_good_peers = -1
col.avg_end_tick = -1
col.avg_end_tick_good_peers = -1
def df_of(key: float) -> pd.DataFrame:
networks_results = results[key]
columns = [AggregatedColumn() for _ in scenarios]
for net_result in networks_results:
for i, result in enumerate(net_result.scenario_results):
if result.end_tick == 9999:
columns[i].success_all = False
if result.end_tick > columns[i].worst_case_end_tick:
columns[i].worst_case_end_tick = result.end_tick
if result.end_tick_good_peers > columns[i].worst_case_end_tick_good_peers:
columns[i].worst_case_end_tick_good_peers = result.end_tick_good_peers
if result.success_good_peers:
columns[i].success_ratio_good_peers += 1
if result.success:
columns[i].success_ratio += 1
# if not result.success:
# continue
# columns[i].success_ratio += 1
columns[i].repeated_msgs_per_tick += result.repeated_msgs / result.end_tick
columns[i].repeated_msgs_per_tick_good_peers += result.repeated_msgs_good_peers / result.end_tick
columns[i].repeated_msgs_ratio += result.repeated_msgs / (len(result.spread_edges) + result.repeated_msgs)
if result.good_peers_notified != 0:
columns[i].repeated_msgs_ratio_good_peers += result.repeated_msgs_good_peers / (result.repeated_msgs_good_peers + result.good_peers_notified)
if result.success:
columns[i].avg_end_tick += result.end_tick
if result.success_good_peers:
columns[i].avg_end_tick_good_peers += result.end_tick_good_peers
for col in columns:
col.avg_end_tick /= col.success_ratio
col.avg_end_tick_good_peers /= col.success_ratio_good_peers
col.repeated_msgs_per_tick /= len(networks_results)
col.repeated_msgs_per_tick_good_peers /= len(networks_results)
col.repeated_msgs_ratio /= len(networks_results)
col.repeated_msgs_ratio_good_peers /= len(networks_results)
col.success_ratio_good_peers /= len(networks_results)
col.success_ratio /= len(networks_results)
df = pd.DataFrame(columns) \
.round(2) \
.rename({i : small_desc(s) for (i,s) in enumerate(scenarios)}) \
return df
# -
df25 = df_of(0.25)
df25[df25["success_ratio_good_peers"] == 1.0]["repeated_msgs_ratio_good_peers"].describe()
x = df25[df25["success_ratio_good_peers"] == 1.0].sort_values("repeated_msgs_per_tick_good_peers", ascending=True).head(5).drop(columns=["success_all", "success_ratio", "success_ratio_good_peers"])
x["avg_end_tick"] = x["avg_end_tick"].apply(ticks_to_time)
x["avg_end_tick_good_peers"] = x["avg_end_tick_good_peers"].apply(ticks_to_time)
x["worst_case_end_tick"] = x["worst_case_end_tick"].apply(ticks_to_time)
x["worst_case_end_tick_good_peers"] = x["worst_case_end_tick_good_peers"].apply(ticks_to_time)
x
df50 = df_of(0.50)
df50[df50["success_ratio_good_peers"] == 0.99]["repeated_msgs_ratio_good_peers"].describe()
x = df50[df50["success_ratio_good_peers"] == 0.99].sort_values("repeated_msgs_per_tick_good_peers", ascending=True).head(5).drop(columns=["success_all", "success_ratio", "success_ratio_good_peers"])
x["avg_end_tick"] = x["avg_end_tick"].apply(ticks_to_time)
x["avg_end_tick_good_peers"] = x["avg_end_tick_good_peers"].apply(ticks_to_time)
x["worst_case_end_tick"] = x["worst_case_end_tick"].apply(ticks_to_time)
x["worst_case_end_tick_good_peers"] = x["worst_case_end_tick_good_peers"].apply(ticks_to_time)
x
df75 = df_of(0.75)
df75[df75["success_ratio_good_peers"] == 0.99]["repeated_msgs_ratio_good_peers"].describe()
x = df75[df75["success_ratio_good_peers"] == 0.99].sort_values("repeated_msgs_per_tick_good_peers", ascending=True).head(5).drop(columns=["success_all", "success_ratio", "success_ratio_good_peers"])
x["avg_end_tick"] = x["avg_end_tick"].apply(ticks_to_time)
x["avg_end_tick_good_peers"] = x["avg_end_tick_good_peers"].apply(ticks_to_time)
x["worst_case_end_tick"] = x["worst_case_end_tick"].apply(ticks_to_time)
x["worst_case_end_tick_good_peers"] = x["worst_case_end_tick_good_peers"].apply(ticks_to_time)
x
# todo check the "fastest" strategies using "avg_end_tick_good_peers" and "worst_case_end_tick_good_peers" columns
x = df25[df25["success_ratio_good_peers"] == 1.0].sort_values("avg_end_tick_good_peers", ascending=True).head(5).drop(columns=["success_all", "success_ratio", "success_ratio_good_peers"])
x["avg_end_tick"] = x["avg_end_tick"].apply(ticks_to_time)
x["avg_end_tick_good_peers"] = x["avg_end_tick_good_peers"].apply(ticks_to_time)
x["worst_case_end_tick"] = x["worst_case_end_tick"].apply(ticks_to_time)
x["worst_case_end_tick_good_peers"] = x["worst_case_end_tick_good_peers"].apply(ticks_to_time)
x
df25[df25["success_ratio_good_peers"] == 1.0].sort_values("worst_case_end_tick_good_peers", ascending=True).head(5)
# todo check the "fastest" strategies using "avg_end_tick_good_peers" and "worst_case_end_tick_good_peers" columns
x = df50[df50["success_ratio_good_peers"] == 0.99].sort_values("avg_end_tick_good_peers", ascending=True).head(5).drop(columns=["success_all", "success_ratio", "success_ratio_good_peers"])
x["avg_end_tick"] = x["avg_end_tick"].apply(ticks_to_time)
x["avg_end_tick_good_peers"] = x["avg_end_tick_good_peers"].apply(ticks_to_time)
x["worst_case_end_tick"] = x["worst_case_end_tick"].apply(ticks_to_time)
x["worst_case_end_tick_good_peers"] = x["worst_case_end_tick_good_peers"].apply(ticks_to_time)
x
df50[df50["success_ratio_good_peers"] == 0.99].sort_values("worst_case_end_tick_good_peers", ascending=True).head(5)
# todo check the "fastest" strategies using "avg_end_tick_good_peers" and "worst_case_end_tick_good_peers" columns
x = df75[df75["success_ratio_good_peers"] == 0.99].sort_values("avg_end_tick_good_peers", ascending=True).head(5).drop(columns=["success_all", "success_ratio", "success_ratio_good_peers"])
x["avg_end_tick"] = x["avg_end_tick"].apply(ticks_to_time)
x["avg_end_tick_good_peers"] = x["avg_end_tick_good_peers"].apply(ticks_to_time)
x["worst_case_end_tick"] = x["worst_case_end_tick"].apply(ticks_to_time)
x["worst_case_end_tick_good_peers"] = x["worst_case_end_tick_good_peers"].apply(ticks_to_time)
x
# todo check the "fastest" strategies using "avg_end_tick_good_peers" and "worst_case_end_tick_good_peers" columns
df75[df75["success_ratio_good_peers"] == 0.99].sort_values("worst_case_end_tick_good_peers", ascending=True).head(5)
| experiments/spread/evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#This is the python code for visdualizing the tropomi data
# -
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# +
#Code to load the tropomi data
# -
import netCDF4 as nc4
# +
#Visualizing the data
# -
# %matplotlib inline
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm
from mpl_toolkits.basemap import Basemap
from datetime import datetime
from pytropomi.downs5p import downs5p
beginPosition = datetime(2019, 7, 7, 0)
endPosition = datetime(2019, 7, 7, 23)
pro = 'L2_NO2___'
longitude = 120
latitude = 30
downs5p(producttype=pro, longitude=longitude, latitude=latitude, processingmode='Near real time',
beginPosition=beginPosition, endPosition=endPosition)
# +
#! -*- coding: utf-8 -*-
from pytropomi.downs5p import downs5p
def test_lonlat(products, longitude, latitude, beginPosition, savepath=None):
for pro in products:
downs5p(producttype=pro, longitude=longitude, latitude=latitude, processingmode='Near real time',
beginPosition=beginPosition, savepath=savepath)
def test_polygon1(products, polygon, area, savepath=None):
for pro in products:
downs5p(producttype=pro, polygon=polygon, area=area, processingmode='Near real time',
savepath=savepath)
def test_polygon2(products, polygon, area, beginPosition, endPosition, savepath=None):
for pro in products:
downs5p(producttype=pro, polygon=polygon, area=area, processingmode='Near real time',
beginPosition=beginPosition, endPosition=endPosition, savepath=savepath)
def test_polygon3(products, polygon, beginPosition, endPosition, savepath=None):
for pro in products:
downs5p(producttype=pro, polygon=polygon, processingmode='Near real time',
beginPosition=beginPosition, endPosition=endPosition, savepath=savepath)
def test_polygon4(products, polygon, beginPosition, savepath=None):
for pro in products:
downs5p(producttype=pro, polygon=polygon, processingmode='Near real time',
beginPosition=beginPosition, savepath=savepath)
def test_polygon5(products, polygon, endPosition, savepath=None):
for pro in products:
downs5p(producttype=pro, polygon=polygon, processingmode='Near real time',
endPosition=endPosition, savepath=savepath)
def test_polygon6(products, polygon, endPosition, savepath):
for pro in products:
downs5p(producttype=pro, polygon=polygon, processingmode='Near real time',
endPosition=endPosition, savepath=savepath)
def test_polygon7(products, polygon, endPosition, savepath):
for pro in products:
downs5p(producttype=pro, polygon=polygon, processingmode='Offline',
endPosition=endPosition, savepath=savepath)
def test_polygon8(products, polygon, endPosition, savepath):
for pro in products:
downs5p(producttype=pro, polygon=polygon, processingmode='Reprocessing',
endPosition=endPosition, savepath=savepath)
if __name__=='__main__':
import os
from datetime import datetime
from shapely.geometry import Polygon
products = ['L2__AER_AI']
polygon = Polygon([(100, 20), (105, 25), (110, 30), (115, 35), (120, 30), (125, 25), (130, 20), (120, 15)])
area = 20
longitude = 121
latitude = 32
beginPosition = datetime(2019, 7, 7)
endPosition = datetime(2019, 7, 8)
savepath = './test'
test_polygon1(products, polygon, area, savepath=None)
# +
lon_0 = lons.mean()
lat_0 = lats.mean()
m = Basemap(width=5000000,height=3500000,
resolution='l',projection='stere',\
lat_ts=40,lat_0=lat_0,lon_0=lon_0)
xi, yi = m(lons, lats)
# Plot Data
cs = m.pcolor(xi,yi,np.squeeze(no2),norm=LogNorm(), cmap='jet')
# Add Grid Lines
m.drawparallels(np.arange(-80., 81., 10.), labels=[1,0,0,0], fontsize=10)
m.drawmeridians(np.arange(-180., 181., 10.), labels=[0,0,0,1], fontsize=10)
# Add Coastlines, States, and Country Boundaries
m.drawcoastlines()
m.drawstates()
m.drawcountries()
# Add Colorbar
cbar = m.colorbar(cs, location='bottom', pad="10%")
cbar.set_label(no2_units)
# Add Title
plt.title('NO2 in atmosphere')
plt.show()
# -
from netCDF4 import Dataset
import numpy as np
my_example_nc_file = 'DATA/TROPOMI_AER_AI/S5P_NRTI_L2__AER_AI_20210213T040730_20210213T041230_17291_01_010400_20210213T044042.nc'
fh = Dataset(my_example_nc_file, mode='r')
print (fh)
# +
print (fh.groups)
print (fh.groups['PRODUCT'])
# -
print (fh.groups['PRODUCT'].variables.keys())
print (fh.groups['PRODUCT'].variables['aerosol_index_354_388'])
# +
lons = fh.groups['PRODUCT'].variables['longitude'][:][0,:,:]
lats = fh.groups['PRODUCT'].variables['latitude'][:][0,:,:]
ai = fh.groups['PRODUCT'].variables['aerosol_index_354_388'][0,:,:]
print (lons.shape)
print (lats.shape)
print (no2.shape)
ai_units = fh.groups['PRODUCT'].variables['aerosol_index_354_388'].units
# -
# %matplotlib inline
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm
from mpl_toolkits.basemap import Basemap
lon_0 = lons.mean()
lon_0
# +
lon_0 = lons.mean()
lat_0 = lats.mean()
m = Basemap(width=5000000,height=3500000,
resolution='l',projection='stere',\
lat_ts=40,lat_0=lat_0,lon_0=lon_0)
xi, yi = m(lons, lats)
# Plot Data
cs = m.pcolor(xi,yi,np.squeeze(ai),norm=LogNorm(), cmap='jet')
# Add Grid Lines
m.drawparallels(np.arange(-80., 81., 10.), labels=[1,0,0,0], fontsize=10)
m.drawmeridians(np.arange(-180., 181., 10.), labels=[0,0,0,1], fontsize=10)
# Add Coastlines, States, and Country Boundaries
m.drawcoastlines()
m.drawstates()
m.drawcountries()
# Add Colorbar
cbar = m.colorbar(cs, location='bottom', pad="10%")
cbar.set_label(no2_units)
# Add Title
plt.title('Aerosol Index in the atmosphere')
plt.show()
# -
import geopandas as gpd
from bokeh.io import output_notebook, show, output_file
from bokeh.plotting import figure
from bokeh.models import GeoJSONDataSource, LinearColorMapper, ColorBar
from bokeh.palettes import brewer
from bokeh.plotting import figure, output_file, show
output_file('my_first_graph.html')
x = [1, 3, 5, 7]
y = [2, 4, 6, 8]
p.legend.click_policy='hide'
show(p)
# +
from bokeh.io import output_file, show
from bokeh.models import ColumnDataSource, GMapOptions
from bokeh.plotting import gmap
output_file("gmap.html")
map_options = GMapOptions(lat=lats.mean(), lng=lons.mean(), map_type="roadmap", zoom=2)
# For GMaps to function, Google requires you obtain and enable an API key:
#
# https://developers.google.com/maps/documentation/javascript/get-api-key
#
# Replace the value below with your personal API key:
p = gmap("<KEY>", map_options, title="Austin")
source = ColumnDataSource(
data=dict(lat=lats,
lon=lons)
)
p.circle(x="lon", y="lat", size=2, fill_color="blue", fill_alpha=0.2, source=source)
show(p)
# +
from bokeh.io import output_file, show
from bokeh.models import GeoJSONDataSource
from bokeh.plotting import figure
from bokeh.sampledata.sample_geojson import geojson
import json
output_file("geojson.html")
data = json.loads(geojson)
for i in range(len(data['features'])):
data['features'][i]['properties']['Color'] = ['blue', 'red'][i%2]
geo_source = GeoJSONDataSource(geojson=json.dumps(data))
TOOLTIPS = [
('Organisation', '@OrganisationName')
]
p = figure(background_fill_color="lightgrey", tooltips=TOOLTIPS)
p.circle(x='x', y='y', size=15, color='Color', alpha=0.7, source=geo_source)
show(p)
# +
from datetime import datetime, timedelta
import numpy as np
import pys5p
#from pys5p.lib.plotlib import FIGinfo
#from pys5p.s5p_msm import S5Pmsm
#from pys5p.s5p_plot import
# -
pys5p
| src/Plotting TROPOMI data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# ## 1.2. Preprocessing weather indexes
library("tidyverse")
library("weathercan")
library("lubridate") # yday(), ymd()
library("Amelia")
# Load the data frame processed in previous file `1.1_preprocessing-1.ipynb`.
data_df <- read.csv('output/balanced_potato_df.csv')
# ### Compute 5 years (mean or total) weather indices
#
# Because the process is long, you can load `data_stations.csv` file and make a left join with `data_df` using "Annee", "LatDD", "LonDD", "DatePlantation", and "DateRecolte" as common features.
data_stations = read_csv("output/weather_stations.csv")
# Custom fonctions
# shannon diversity index
SDI_f <- function(x) {
p <- x/sum(x, na.rm = TRUE)
SDI <- sum(p * log(p), na.rm = TRUE) / log(length(x))
return(SDI)
}
# growing degree-days
GDD_f <- function(x, delim = 5) {
sum(x[x >= delim], na.rm = TRUE)
}
# Create a data frame for computations
data_df$DatePlantation <- ymd(data_df$DatePlantation)
data_df$DateRecolte <- ymd(data_df$DateRecolte)
# If you've loaded the `weather_stations.csv`, from here, you coud jump to the fith code chaine from the end (insertion into the main table).
# + active=""
#
# + active=""
#
# -
data_stations <- data_df %>%
distinct(Annee, LatDD, LonDD, DatePlantation, DateRecolte) %>%
filter(!is.na(Annee))
glimpse(data_stations)
# Impute missing dates if year is recorded
# Check missing dates
data_stations %>%
filter(is.na(DatePlantation))
# Check missing dates
data_stations %>%
filter(is.na(DateRecolte))
# +
data_stations_imp <- data_stations %>%
mutate(DatePlantation_yd = yday(DatePlantation), # yday = year day
DateRecolte_yd = yday(DateRecolte))
data_stations_imp <- amelia(x = data_stations_imp %>%
select(Annee, LatDD, LonDD, DatePlantation_yd,
DateRecolte_yd), m=1)$imputations[[1]]
# +
data_stations_imp$Annee <- as.Date(paste0(data_stations_imp$Annee, "-01-01"))
data_stations$DatePlantation <- data_stations_imp$Annee + data_stations_imp$DatePlantation_yd - 1
data_stations$DateRecolte <- data_stations_imp$Annee + data_stations_imp$DateRecolte_yd - 1
# -
data_stations$DatePlantation <- as.Date(data_stations$DatePlantation)
data_stations$DateRecolte <- as.Date(data_stations$DateRecolte)
# Load data from Environnement Canada (weathercan)
# +
year_step <- 5 # start 5 years preceeding trial year
data_stations$station_id <- NA # identify weather stations
station_weather <- list() # list of weather stations containning climate data
for (i in 1:nrow(data_stations)) {
print(paste(i, "/", nrow(data_stations), '...'))
# identify the nearest station. Search for stations by name or location
all_stations <- stations_search(coords = c(data_stations$LatDD[i],
data_stations$LonDD[i]),
interval = "day",
dist = 500)
annee <- data_stations$Annee[i]
# Available station for nearest year
closest_station <- all_stations %>%
filter(start <= annee - year_step) %>%
filter(end >= annee) %>%
slice(which.min(distance))
data_stations$station_id[i] <- closest_station$station_id %>%
as.character() %>%
as.numeric()
print(paste("Station id:", data_stations$station_id[i]))
# # Download weather data from Environment Canada of the station identified
station_weather[[i]] <- weather_dl(station_ids = data_stations$station_id[i],
start = as.Date(paste0(annee - year_step, "-01-01")),
end = as.Date(paste0(annee, "-01-01")),
interval = "day")
print(paste(i, "/", nrow(data_stations)))
}
# name of the list elements
names(station_weather) <- as.character(data_stations$station_id)
# -
# Compute indexes for each station
# +
# Initialize columns to record indexes
data_stations$temp_moy_5years <- NA
data_stations$prec_tot_5years <- NA
data_stations$sdi_5years <- NA
data_stations$gdd_5years <- NA
for (i in 1:nrow(data_stations)) {
starts <- data_stations$DatePlantation[i] - (year_step:1)*365.25 # beginings of seasons
ends <- data_stations$DateRecolte[i] - (year_step:1)*365.25 # ends of seasons, for previous years
# Create vectors to record indices from previous years
temp_moy_j <- rep(NA, length(starts))
prectot_j <- temp_moy_j
sdi_j <- temp_moy_j
gdd_j <- temp_moy_j
for (j in 1:length(starts)) {
# Filter the downloaded weather data tables to keep only data measured between the season dates
season <- station_weather[[i]] %>%
filter(date >= starts[j]) %>%
filter(date <= ends[j])
temp_moy_j[j] <- mean(season$mean_temp, na.rm = TRUE) # average temperature of the season
prectot_j[j] <- sum(season$total_precip, na.rm = TRUE) # total precipitations of the season
sdi_j[j] <- SDI_f(season$total_precip) # season SDI
gdd_j[j] <- GDD_f(season$mean_temp, delim = 5) # season GDD
}
data_stations$temp_moy_5years[i] <- mean(temp_moy_j, na.rm = TRUE) # average temperature
data_stations$prec_tot_5years[i] <- mean(prectot_j, na.rm = TRUE) # average total rainfalls
data_stations$sdi_5years[i] <- mean(sdi_j, na.rm = TRUE) # average SDI
data_stations$gdd_5years[i] <- mean(gdd_j, na.rm = TRUE) # average GDD
}
# -
write_csv(x = data_stations, path = "output/weather_stations.csv") # a backup
# + active=""
#
# + active=""
#
# -
# If you've loaded the `weather_stations.csv`, continue from here.
# Insert into the main table
weath_col <- c(
'Annee', 'LatDD', 'LonDD', 'DatePlantation', 'DateRecolte',
'temp_moy_5years', 'prec_tot_5years', 'sdi_5years', 'gdd_5years'
)
data.frame(weath_col, weath_col %in% colnames(data_df))
data_df <- left_join(data_df, data_stations[weath_col],
by = c("Annee", "LatDD", "LonDD", "DatePlantation", "DateRecolte"))
wcol <- c('temp_moy_5years', 'prec_tot_5years', 'sdi_5years', 'gdd_5years')
data.frame(wcol, wcol %in% colnames(data_df))
write_csv(data_df, "output/pr_potato_df.csv") # backup
# + active=""
#
| 1.2_preprocessing-weather-indexes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="i5OW4csoRjXr"
# # Part 2 - Regression with PyTorch
# + id="0WcjqDImRf87"
import os
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from matplotlib import pyplot as plt
# To disable all warnings
import warnings
warnings.filterwarnings("ignore")
# + id="TegKk2P5M3xK"
print(torch.__version__)
# + id="jNYWWGqo0Fkm"
# Device to perform training. Default to 'cpu'
device = 'cuda'
# + [markdown] id="L7yS0sZ8Ls8H"
# ## Prepare Dummy Data
# + id="ZFOUrJxwXsJi"
X = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10]
y = [4, 8, 12, 16, 20, 24, 28, 32, 36, 40]
X_tensor = torch.tensor(X).float().to(device)
y_tensor = torch.tensor(y).float().to(device)
# + [markdown] id="-mtvBedpLz31"
# ## Hyperparameter
# + id="-vKKszIyLXYT"
learning_rate = 0.1
momentum = 0.5
n_epochs = 100
# + [markdown] id="EDZsbPpHLwea"
# ## Define model
# + id="HQrG4ScyYtoo"
# EXERCISE: Create you model here
class Net(nn.Module):
pass
# + id="Wnef-6bHg_9M"
# EXERCISE: Build model instance. Set training to GPU
model = None
# + id="PuaOBb6ML6HJ"
# EXERCISE: Define training optimizer
optimizer = None
# + [markdown] id="QLEf2nTQL_qg"
# ## Train model
# + id="MQttClGbaOrh"
loss_values = []
for epoch in range(1, n_epochs + 1):
print("Epoch {:02d}".format(epoch))
log_interval = 10
model.train()
running_loss = 0.0
for batch_idx, (data, target) in enumerate(zip(X_tensor, y_tensor)):
# EXERCISE: Build your training loop Here
pass
# + id="ei2pQJASbCUI"
model_path = os.path.join("model.pth")
torch.save(model.state_dict(), model_path)
# + id="rb3pEeYRtUZr"
ax = plt.plot(loss_values)
plt.title("Training Loss")
# + [markdown] id="nCTYWVsRIOV4"
# ## Test Model
# + id="9be2NOFXgUYD"
# EXERCISE: Prepare test data. Test the model with input 11
test_data = None
# + id="MwuHUDwDc3sz"
# Set model in inference mode
# Will change behavior of certain layers from training to inference. eg: dropout, batchnorm
model.eval()
with torch.no_grad():
prediction = model(test_data)
# + id="zouaLkfxII7e"
print(prediction)
# + [markdown] id="5IKIRVSRK21S"
# ### Extra: Tensor to numpy
# + id="BxKdZ8goI34I"
prediction_np = prediction.cpu().detach().numpy()
prediction_np
# + id="jNDM581YK804"
# Get value only
prediction_np[0]
# + [markdown] id="dzO1wXW6OBBJ"
# ### Extra: Let's view the trained weight
# + id="g83ZNJCDLKuQ"
model.state_dict()
| notebooks/Day_3/Regression_Exercise/Ex-Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# default_exp indexers.notelist.parser
# export
import bs4
import random
from pyintegrators.indexers.notelist.util import *
from pyintegrators.indexers.notelist.notelist import *
from pyintegrators.data.schema import *
from pyintegrators.data.basic import *
from pyintegrators.imports import *
#hide
from nbdev.showdoc import *
# # HTMLListParser
# This parsers takes Note objects with a .content field containing the html content of the note as input, and parsers the lists from the html content.
# export
class HTMLListParser():
'''Extracts lists from HTML data, generated by an HTML text editor like evernote'''
def __init__(self):
self.one_line_list_pa = ["buy", "read", "watch"]
words = ["do", "read", "watch", "buy", "listen"]
prefixes = ["to", "to-", "to ", ""]
self.single_item_list_patterns = [prefix+word for word in words for prefix in prefixes]
def get_lists(self, note):
"""Extracts lists from a note"""
text = note.content
parsed = bs4.BeautifulSoup(text, 'html.parser')
note.content=str(parsed)
uls = get_toplevel_elements(text, "ul", parsed=parsed)
ols = get_toplevel_elements(text, "ol", parsed=parsed)
html_lists = [ULNoteList.from_data(title=None, content=str(x),
textContent=remove_html(str(x)), note=note, span=get_span(note, x, parsed))
for x in uls + ols]
unformatted_lists = self.get_unformatted_lists(note, text, parsed)
all_lists = html_lists + unformatted_lists
for l in all_lists:
note.add_edge("noteList", l)
return all_lists
def get_single_line_list(self, elem):
"""Get single list lists. An example could be: '<strong>read</strong>: great book title'"""
ps = ["read", "buy", "watch", "do"]
pat = "|".join([f"(<strong>|<em>|<u>)?{p}(</strong>|</em>|</u>)?:? ?" for p in ps])
match = re.search(pat, str(elem), re.IGNORECASE)
if match is None: return None, None
cleaned_elem = remove_html(str(elem))
cleaned_title = remove_html(match.group()) if match is not None else None
if len(cleaned_elem) > len(cleaned_title) + 2:
title = match.group()
content = str(elem)[len(title):]
return title, content
else:
return None, None
def get_unformatted_lists(self, note, txt, parsed):
"""retrieve lists without <ul></ul> tags. We have two options:
1) multiline lists prefixed with a title keyword (e.g. "Buy:" "Read:")
2) single element single line lists"""
toplevel_div = get_toplevel_elements(txt, "div")[0]
ls = []
for elem in toplevel_div.children:
if elem.name == "div" and not is_newline_div(elem):
children = get_children(elem)
for i, child in enumerate(children):
# this extracts the lists that have a title and are not on a single line
if div_is_unstructured_list_title(child):
title = child
successors = list(children)[i+1:]
if len(successors) == 0:
continue
items = [x for x in find_till_double_br(successors) if not is_newline(str(x))]
items_str = [str(x) for x in items]
items_span = [get_span(note, x, parsed) for x in items_str]
span1 = get_span(note, title, parsed)
span2 = get_span(note, items[-1], parsed)
span = Span.from_data(startIdx=span1.startIdx, endIdx=span2.endIdx)
html_content = "".join(items_str)
l = INoteList.from_data(note=note, title=title, content=str(html_content), itemSpan=items_span, span=span)
ls.append(l)
else:
title, html_content = self.get_single_line_list(child)
if title is not None:
span = get_span(note, child, parsed)
itemSpan = [Span.from_data(startIdx=span.startIdx + len(str(title)), endIdx=span.endIdx)]
l = INoteList.from_data(note=note, title=title, content=str(html_content), itemSpan=itemSpan, span=span)
ls.append(l)
return ls
show_doc(HTMLListParser.get_lists)
show_doc(HTMLListParser.get_unformatted_lists)
show_doc(HTMLListParser.get_single_line_list)
# # Usage
# Lets see how this works for an example note. We start with a note that was imported from evernote as example and show its content.
evernote_file = PYI_TESTDATA / "notes" / "evernote" / "evernote-test-note-1.html"
txt = read_file(evernote_file)
note = INote.from_data(content=txt)
note.show()
#hide
from IPython.core.display import display, HTML
# Which corresponds to this when rendered
display(HTML(note.content))
# We can parse these using the `HTMLListParser`
parser = HTMLListParser()
lists = parser.get_lists(note)
lists
# # Export -
# hide
from nbdev.export import *
notebook2script()
| nbs/indexers.NoteListIndexer.Parser.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # Ibis for SQL Programmers
#
# Ibis provides a full-featured replacement for SQL
# `SELECT` queries, but expressed with Python code that is:
#
# - Type-checked and validated as you go. No more debugging cryptic
# database errors; Ibis catches your mistakes right away.
# - Easier to write. Pythonic function calls with tab completion in
# IPython.
# - More composable. Break complex queries down into easier-to-digest
# pieces
# - Easier to reuse. Mix and match Ibis snippets to create expressions
# tailored for your analysis.
#
# We intend for all `SELECT` queries to be fully portable to Ibis.
# Coverage of other DDL statements (e.g. `CREATE TABLE` or `INSERT`) may
# vary from engine to engine.
#
# This document will use the Impala SQL compiler (i.e.
# `ibis.impala.compile`) for convenience, but the code here is portable to
# whichever system you are using Ibis with.
#
# > **Note**: If you find any SQL idioms or use cases in your work that
# > are not represented here, please reach out so we can add more to this
# > guide!
# -
import ibis
ibis.options.sql.default_limit = None
# ## Projections: select/add/remove columns
#
# All tables in Ibis are immutable. To select a subset of a table's
# columns, or to add new columns, you must produce a new table by means of
# a *projection*.
t = ibis.table([('one', 'string'), ('two', 'float'), ('three', 'int32')], 'my_data')
t
# In SQL, you might write something like:
#
# ```sql
# SELECT two, one
# FROM my_data
# ```
#
# In Ibis, this is
proj = t['two', 'one']
# or
proj = t.projection(['two', 'one'])
# This generates the expected SQL:
print(ibis.impala.compile(proj))
# What about adding new columns? To form a valid projection, all column
# expressions must be **named**. Let's look at the SQL:
#
# ``` sql
# SELECT two, one, three * 2 AS new_col
# FROM my_data
# ```
#
# The last expression is written:
new_col = (t.three * 2).name('new_col')
# Now, we have:
proj = t['two', 'one', new_col]
print(ibis.impala.compile(proj))
# ### `mutate`: Add or modify columns easily
#
# Since adding new columns or modifying existing columns is so common,
# there is a convenience method `mutate`:
mutated = t.mutate(new_col=t.three * 2)
# Notice that using the `name` was not necessary here because we're using
# Python keywords to provide the name. Indeed:
print(ibis.impala.compile(mutated))
# If you modify an existing column with `mutate` it will list out all the
# other columns:
mutated = t.mutate(two=t.two * 2)
print(ibis.impala.compile(mutated))
# ### `SELECT *` equivalent
#
# Especially in combination with relational joins, it's convenient to be
# able to select all columns in a table using the `SELECT *` construct. To
# do this, use the table expression itself in a projection:
proj = t[t]
print(ibis.impala.compile(proj))
# This is how `mutate` is implemented. The example above
# `t.mutate(new_col=t.three * 2)` can be written as a normal projection:
proj = t[t, new_col]
print(ibis.impala.compile(proj))
# Let's consider a table we might wish to join with `t`:
t2 = ibis.table([('key', 'string'), ('value', 'float')], 'dim_table')
# Now let's take the SQL:
#
# ``` sql
# SELECT t0.*, t0.two - t1.value AS diff
# FROM my_data t0
# INNER JOIN dim_table t1
# ON t0.one = t1.key
# ```
#
# To write this with Ibis, it is:
diff = (t.two - t2.value).name('diff')
joined = t.join(t2, t.one == t2.key)[t, diff]
# And verify the generated SQL:
print(ibis.impala.compile(joined))
# ### Using functions in projections
#
# If you pass a function instead of a string or Ibis expression in any
# projection context, it will be invoked with the "parent" table as its
# argument. This can help significantly when [composing complex operations.
# Consider this SQL:
#
# ``` sql
# SELECT one, avg(abs(the_sum)) AS mad
# FROM (
# SELECT one, three, sum(two) AS the_sum
# FROM my_data
# GROUP BY 1, 2
# ) t0
# GROUP BY 1
# ```
#
# This can be written as one chained expression:
expr = (
t.group_by(['one', 'three'])
.aggregate(the_sum=t.two.sum()) .group_by('one')
.aggregate(mad=lambda x: x.the_sum.abs().mean())
)
# Indeed:
print(ibis.impala.compile(expr))
# ## Filtering / `WHERE`
#
# You can add filter clauses to a table expression either by indexing with
# `[]` (like pandas) or use the `filter` method:
filtered = t[t.two > 0]
print(ibis.impala.compile(filtered))
# `filter` can take a list of expressions, which must all be satisfied for
# a row to be included in the result:
filtered = t.filter([t.two > 0, t.one.isin(['A', 'B'])])
print(ibis.impala.compile(filtered))
# To compose boolean expressions with `AND` or `OR`, use the respective
# `&` and `|` operators:
cond = (t.two < 0) | ((t.two > 0) | t.one.isin(['A', 'B']))
filtered = t[cond]
print(ibis.impala.compile(filtered))
# ## Aggregation / `GROUP BY`
#
# To aggregate a table, you need:
#
# - Zero or more grouping expressions (these can be column names)
# - One or more aggregation expressions
#
# Let's look at the `aggregate` method on tables:
stats = [t.two.sum().name('total_two'),
t.three.mean().name('avg_three')]
agged = t.aggregate(stats)
# If you don't use any group expressions, the result will have a single
# row with your statistics of interest:
agged.schema()
print(ibis.impala.compile(agged))
# To add groupings, use either the `by` argument of `aggregate` or use the
# `group_by` construct:
agged2 = t.aggregate(stats, by='one')
agged3 =t.group_by('one').aggregate(stats)
print(ibis.impala.compile(agged3))
# ### Non-trivial grouping keys
#
# You can use any expression (or function, like in projections) deriving
# from the table you are aggregating. The only constraint is that the
# expressions must be named. Let's look at an example:
events = ibis.table([('ts', 'timestamp'),
('event_type', 'int32'), ('session_id', 'int64')],
name='web_events')
# Suppose we wanted to total up event types by year and month:
# +
keys = [events.ts.year().name('year'),
events.ts.month().name('month')]
sessions = events.session_id.nunique()
stats = (events.group_by(keys)
.aggregate(total=events.count(), sessions=sessions))
# -
# Now we have:
print(ibis.impala.compile(stats))
# ### Aggregates considering table subsets
#
# In analytics is it common to compare statistics from different subsets
# of a table. Let's consider a dataset containing people's name, age,
# gender, and nationality:
pop = ibis.table([('name', 'string'),
('country', 'string'), ('gender', 'string'), ('age', 'int16')],
name='population')
# Now, suppose you wanted to know for each country:
#
# - Average overall age
# - Average male age
# - Average female age
# - Total number of persons
#
# In SQL, you may write:
#
# ``` sql
# SELECT country,
# count(*) AS num_persons,
# AVG(age) AS avg_age
# AVG(CASE WHEN gender = 'M'
# THEN age
# ELSE NULL
# END) AS avg_male,
# AVG(CASE WHEN gender = 'F'
# THEN age
# ELSE NULL
# END) AS avg_female,
# FROM population
# GROUP BY 1
# ```
#
# Ibis makes this much simpler by giving you `where` option in aggregation
# functions:
expr = pop.group_by('country').aggregate(
num_persons=pop.count(), avg_age=pop.age.mean(),
avg_male=pop.age.mean(where=pop.gender == 'M'),
avg_female=pop.age.mean(where=pop.gender == 'F')
)
# This indeed generates the correct SQL. Note that SQL engines handle
# `NULL` values differently in aggregation functions, but Ibis will write
# the SQL expression that is correct for your query engine.
print(ibis.impala.compile(expr))
# ### `count(*)` convenience: `size()`
#
# Computing group frequencies is so common that, like pandas, we have a
# method `size` that is a shortcut for the `count(*)` idiom:
freqs = events.group_by(keys).size()
print(ibis.impala.compile(freqs))
# ### Frequency table convenience: `value_counts`
#
# Consider the SQL idiom:
#
# ``` sql
# SELECT some_column_expression, count(*)
# FROM table
# GROUP BY 1
# ```
#
# This is so common that, like pandas, there is a generic array method
# `value_counts` which does this for us:
expr = events.ts.year().value_counts()
print(ibis.impala.compile(expr))
# ### `HAVING` clause
#
# The SQL `HAVING` clause enables you to filter the results of an
# aggregation based on some group-wise condition holding true. For
# example, suppose we wanted to limit our analysis to groups containing at
# least 1000 observations:
#
# ``` sql
# SELECT one, sum(two) AS total
# FROM my_data
# GROUP BY 1
# HAVING count(*) >= 1000
# ```
#
# With Ibis, you can do:
expr = (t.group_by('one')
.having(t.count() >= 1000) .aggregate(t.two.sum().name('total')))
print(ibis.impala.compile(expr))
# ## Sorting / `ORDER BY`
#
# To sort a table, use the `sort_by` method along with either column names
# or expressions that indicate the sorting keys:
# +
sorted = events.sort_by([events.ts.year(), events.ts.month()])
print(ibis.impala.compile(sorted))
# -
# The default for sorting is in ascending order. To reverse the sort
# direction of any key, either wrap it in `ibis.desc` or pass a tuple with
# `False` as the second value:
# +
sorted = events.sort_by([
ibis.desc('event_type'),
(events.ts.month(), False)
]) .limit(100)
print(ibis.impala.compile(sorted))
# -
# ## `LIMIT` and `OFFSET`
#
# This one is easy. The table `limit` function truncates a table to the
# indicates number of rows. So if you only want the first 1000 rows (which
# may not be deterministic depending on the SQL engine), you can do:
limited = t.limit(1000)
print(ibis.impala.compile(limited))
# The `offset` option in `limit` skips rows. So if you wanted rows 11
# through 20, you could do:
limited = t.limit(10, offset=10)
print(ibis.impala.compile(limited))
# ## Common column expressions
#
# See the full `API documentation <api>`{.interpreted-text role="ref"} for
# all of the available value methods and tools for creating value
# expressions. We mention a few common ones here as they relate to common
# SQL queries.
#
# ### Type casts
#
# Ibis's type system is independent of any SQL system. You cast Ibis
# expressions from one Ibis type to another. For example:
# +
expr = t.mutate(date=t.one.cast('timestamp'),
four=t.three.cast('float'))
print(ibis.impala.compile(expr))
# -
# ### `CASE` statements
#
# SQL dialects typically support one or more kind of `CASE` statements.
# The first is the *simple case* that compares against exact values of an
# expression.
#
# ``` sql
# CASE expr
# WHEN value_1 THEN result_1
# WHEN value_2 THEN result_2
# ELSE default
# END
# ```
#
# Value expressions in Ibis have a `case` method that allows us to emulate
# these semantics:
# +
case = (t.one.cast('timestamp')
.year()
.case()
.when(2015, 'This year')
.when(2014, 'Last year')
.else_('Earlier')
.end())
expr = t.mutate(year_group=case)
print(ibis.impala.compile(expr))
# -
# The more general case is that of an arbitrary list of boolean
# expressions and result values:
#
# ``` sql
# CASE
# WHEN boolean_expr1 THEN result_1
# WHEN boolean_expr2 THEN result_2
# WHEN boolean_expr3 THEN result_3
# ELSE default
# END
# ```
#
# To do this, use `ibis.case`:
# +
case = (ibis.case()
.when(t.two < 0, t.three * 2)
.when(t.two > 1, t.three)
.else_(t.two)
.end())
expr = t.mutate(cond_value=case)
print(ibis.impala.compile(expr))
# -
# There are several places where Ibis builds cases for you in a simplified
# way. One example is the `ifelse` function:
switch = (t.two < 0).ifelse('Negative', 'Non-Negative')
expr = t.mutate(group=switch)
print(ibis.impala.compile(expr))
# ### Using `NULL` in expressions
#
# To use `NULL` in an expression, either use the special `ibis.NA` value
# or `ibis.null()`:
pos_two = (t.two > 0).ifelse(t.two, ibis.NA)
expr = t.mutate(two_positive=pos_two)
print(ibis.impala.compile(expr))
# ### Set membership: `IN` / `NOT IN`
#
# Let's look again at the population dataset. Suppose you wanted to
# combine the United States and Canada data into a "North America"
# category. Here would be some SQL to do it:
#
# ``` sql
# CASE
# WHEN upper(country) IN ('UNITED STATES', 'CANADA')
# THEN 'North America'
# ELSE country
# END AS refined_group
# ```
#
# The Ibis equivalent of `IN` is the `isin` method. So we can write:
# +
refined = (pop.country.upper()
.isin(['UNITED STATES', 'CANADA'])
.ifelse('North America', pop.country))
expr = pop.mutate(refined_group=refined)
print(ibis.impala.compile(expr))
# -
# The opposite of `isin` is `notin`.
#
# ### Constant and literal expressions
#
# Consider a SQL expression like:
#
# ``` sql
# 'foo' IN (column1, column2)
# ```
#
# which is equivalent to
#
# ``` sql
# column1 = 'foo' OR column2 = 'foo'
# ```
#
# To build expressions off constant values, you must first convert the
# value (whether a Python string or number) to an Ibis expression using
# `ibis.literal`:
# +
t3 = ibis.table([('column1', 'string'),
('column2', 'string'), ('column3', 'float')],
'data')
value = ibis.literal('foo')
# -
# Once you've done this, you can use the literal expression like any
# other array or scalar expression:
# +
has_foo = value.isin([t3.column1, t3.column2])
expr = t3.mutate(has_foo=has_foo)
print(ibis.impala.compile(expr))
# -
# In many other situations, you can use constants without having to use
# `ibis.literal`. For example, we could add a column containing nothing
# but the number 5 like so:
expr = t3.mutate(number5=5)
print(ibis.impala.compile(expr))
# ### `IS NULL` and `IS NOT NULL`
#
# These are simple: use the `isnull` and `notnull` functions respectively,
# which yield boolean arrays:
# +
indic = t.two.isnull().ifelse('valid', 'invalid')
expr = t.mutate(is_valid=indic)
print(ibis.impala.compile(expr))
agged = (expr[expr.one.notnull()]
.group_by('is_valid')
.aggregate(three_count=lambda t: t.three.notnull().sum()))
print(ibis.impala.compile(agged))
# -
# ### `BETWEEN`
#
# The `between` method on arrays and scalars compiles to the SQL `BETWEEN`
# keyword. The result of `between` is boolean and can be used with any
# other boolean expression:
expr = t[t.two.between(10, 50) & t.one.notnull()]
print(ibis.impala.compile(expr))
# ## Joins
#
# Ibis supports several kinds of joins between table expressions:
#
# - `inner_join`: maps to SQL `INNER JOIN`
# - `cross_join`: a cartesian product join with no keys. Equivalent to
# `inner_join` with no join predicates
# - `left_join`: maps to SQL `LEFT OUTER JOIN`
# - `outer_join`: maps to SQL `FULL OUTER JOIN`
# - `semi_join`: maps to SQL `LEFT SEMI JOIN`. May or may not be an
# explicit join type in your query engine.
# - `anti_join`: maps to SQL `LEFT ANTI JOIN`. May or may not be an
# explicit join type in your query engine.
#
# The `join` table method is by default the same as `inner_join`.
#
# Let's look at a couple example tables to see how joins work in Ibis:
# +
t1 = ibis.table([('value1', 'float'),
('key1', 'string'), ('key2', 'string')],
'table1')
t2 = ibis.table([('value2', 'float'), ('key3', 'string'), ('key4', 'string')],
'table2')
# -
# Let's join on one key:
joined = t1.left_join(t2, t1.key1 == t2.key3)
# The immediate result of a join does not yet have a set schema. That is
# determined by the next action that you take. There's several ways
# forward from here that address the spectrum of SQL use cases.
#
# ### Join + projection
#
# Consider the SQL:
#
# ``` sql
# SELECT t0.*, t1.value2
# FROM table1 t0
# LEFT OUTER JOIN table2 t1
# ON t0.key1 = t1.key3
# ```
#
# After one or more joins, you can reference any of the joined tables in a
# projection immediately after:
expr = joined[t1, t2.value2]
print(ibis.impala.compile(expr))
# If you need to compute an expression that involves both tables, you can
# do that also:
expr = joined[t1.key1, (t1.value1 - t2.value2).name('diff')]
print(ibis.impala.compile(expr))
# ### Join + aggregation
#
# You can directly aggregate a join without need for projection, which
# also allows you to form statistics that reference any of the joined
# tables.
#
# Consider this SQL:
#
# ``` sql
# SELECT t0.key1, avg(t0.value1 - t1.value2) AS avg_diff
# FROM table1 t0
# LEFT OUTER JOIN table2 t1
# ON t0.key1 = t1.key3
# GROUP BY 1
# ```
#
# As you would hope, the code is as follows:
avg_diff = (t1.value1 - t2.value2).mean()
expr = (t1.left_join(t2, t1.key1 == t2.key3)
.group_by(t1.key1)
.aggregate(avg_diff=avg_diff))
print(ibis.impala.compile(expr))
# ### Join with `SELECT *`
#
# If you try to compile or execute a join that has not been projected or
# aggregated, it will be *fully materialized*:
joined = t1.left_join(t2, t1.key1 == t2.key3)
print(ibis.impala.compile(joined))
# ### Multiple joins
#
# You can join multiple tables together in succession without needing to
# address any of the above concerns.
# +
t3 = ibis.table([('value3', 'float'),
('key5', 'string')], 'table3')
total = (t1.value1 + t2.value2 + t3.value3).sum()
expr = (t1.join(t2, [t1.key1 == t2.key3, t1.key2 == t2.key4])
.join(t3, t1.key1 == t3.key5)
.group_by([t2.key4, t3.key5])
.aggregate(total=total))
print(ibis.impala.compile(expr))
# -
# ### Self joins
#
# What about when you need to join a table on itself? For example:
#
# ``` sql
# SELECT t0.one, avg(t0.two - t1.three) AS metric
# FROM my_data t0
# INNER JOIN my_data t1
# ON t0.one = t1.one
# GROUP BY 1
# ```
#
# The table `view` method enables you to form a *self-reference* that is
# referentially distinct in expressions. Now you can proceed normally:
# +
t_view = t.view()
stat = (t.two - t_view.three).mean()
expr = (t.join(t_view, t.three.cast('string') == t_view.one)
.group_by(t.one)
.aggregate(metric=stat))
print(ibis.impala.compile(expr))
# -
# ### Overlapping join keys
#
# In many cases the columns being joined between two tables or table
# expressions have the same name. Consider this example:
# +
t4 = ibis.table([('key1', 'string'),
('key2', 'string'),
('key3', 'string'),
('value1', 'float')], 'table4')
t5 = ibis.table([('key1', 'string'),
('key2', 'string'),
('key3', 'string'),
('value2', 'float')], 'table5')
# -
# In these case, we can specify a list of common join keys:
joined = t4.join(t5, ['key1', 'key2', 'key3'])
expr = joined[t4, t5.value2]
print(ibis.impala.compile(expr))
# You can mix the overlapping key names with other expressions:
joined = t4.join(t5, ['key1', 'key2', t4.key3.left(4) == t4.key3.left(4)])
expr = joined[t4, t5.value2]
print(ibis.impala.compile(expr))
# ### Non-equality join predicates
#
# You can join tables with boolean clauses that are not equality. Some
# query engines support these efficiently, some inefficiently, or some not
# at all. In the latter case, these conditions get moved by Ibis into the
# `WHERE` part of the `SELECT` query.
expr = (t1.join(t2, t1.value1 < t2.value2)
.group_by(t1.key1) .size())
print(ibis.impala.compile(expr))
# ### Other ways to specify join keys
#
# You can also pass a list of column names instead of forming boolean
# expressions:
joined = t1.join(t2, [('key1', 'key3'),
('key2', 'key4')])
# ## Subqueries
#
# Ibis creates inline views and nested subqueries automatically. This
# section concerns more complex subqueries involving foreign references
# and other advanced relational algebra.
#
# ### Correlated `EXISTS` / `NOT EXISTS` filters
#
# The SQL `EXISTS` and `NOT EXISTS` constructs are typically used for
# efficient filtering in large many-to-many relationships.
#
# Let's consider a web dataset involving website session / usage data and
# purchases:
# +
events = ibis.table([('session_id', 'int64'),
('user_id', 'int64'),
('event_type', 'int32'),
('ts', 'timestamp')], 'events')
purchases = ibis.table([('item_id', 'int64'),
('user_id', 'int64'),
('price', 'float'),
('ts', 'timestamp')], 'purchases')
# -
# Now, the key `user_id` appears with high frequency in both tables. But
# let's say you want to limit your analysis of the `events` table to only
# sessions by users who have made a purchase.
#
# In SQL, you can do this using the somewhat esoteric `EXISTS` construct:
#
# ``` sql
# SELECT t0.*
# FROM events t0
# WHERE EXISTS (
# SELECT 1
# FROM purchases t1
# WHERE t0.user_id = t1.user_id
# )
# ```
#
# To describe this operation in Ibis, you compare the `user_id` columns
# and use the `any` reduction:
cond = (events.user_id == purchases.user_id).any()
# This can now be used to filter `events`:
expr = events[cond]
print(ibis.impala.compile(expr))
# If you negate the condition, it will instead give you only event data
# from user *that have not made a purchase*:
expr = events[-cond]
print(ibis.impala.compile(expr))
# ### Subqueries with `IN` / `NOT IN`
#
# Subquery filters with `IN` (and `NOT IN`) are functionally similar to
# `EXISTS` subqueries. Let's look at some SQL:
#
# ``` sql
# SELECT *
# FROM events
# WHERE user_id IN (
# SELECT user_id
# FROM purchases
# )
# ```
#
# This is almost semantically the same as the `EXISTS` example. Indeed,
# you can write with Ibis:
cond = events.user_id.isin(purchases.user_id)
expr = events[cond]
print(ibis.impala.compile(expr))
# Depending on the query engine, the query planner/optimizer will often
# rewrite `IN` or `EXISTS` subqueries into the same set of relational
# algebra operations.
#
# ### Comparison with scalar aggregates
#
# Sometime you want to compare a value with an unconditional aggregate
# value from a different table. Take the SQL:
#
# ``` sql
# SELECT *
# FROM table1
# WHERE value1 > (
# SELECT max(value2)
# FROM table2
# )
# ```
#
# With Ibis, the code is simpler and more pandas-like:
expr = t1[t1.value1 > t2.value2.max()]
print(ibis.impala.compile(expr))
# ### Conditional aggregates
#
# Suppose you want to compare a value with the aggregate value for some
# common group values between two tables. Here's some SQL:
#
# ``` sql
# SELECT *
# FROM table1 t0
# WHERE value1 > (
# SELECT avg(value2)
# FROM table2 t1
# WHERE t0.key1 = t1.key3
# )
# ```
#
# This query computes the average for each distinct value of `key3` and
# uses the corresponding average for the comparison, rather than the
# whole-table average as above.
#
# With Ibis, the code is similar, but you add the correlated filter to the
# average statistic:
stat = t2[t1.key1 == t2.key3].value2.mean()
expr = t1[t1.value1 > stat]
print(ibis.impala.compile(expr))
# ## `DISTINCT` expressions
#
# In SQL, the `DISTINCT` keyword is used in a couple of ways:
#
# - Deduplicating identical rows in some `SELECT` statement
# - Aggregating on the distinct values of some column expression
#
# Ibis supports both use cases. So let's have a look. The first case is
# the simplest: call `distinct` on a table expression. First, here's the
# SQL:
#
# ``` sql
# SELECT DISTINCT *
# FROM table1
# ```
#
# And the Ibis Python code:
expr = t1.distinct()
print(ibis.impala.compile(expr))
# For distinct aggregates, the most common case is `COUNT(DISTINCT ...)`,
# which computes the number of unique values in an expression. So if
# we're looking at the `events` table, let's compute the number of
# distinct `event_type` values for each `user_id`. First, the SQL:
#
# ``` sql
# SELECT user_id, COUNT(DISTINCT event_type) AS unique_events
# FROM events
# GROUP BY 1
# ```
#
# In Ibis this is:
metric = events.event_type.nunique()
expr =(events.group_by('user_id')
.aggregate(unique_events=metric))
print(ibis.impala.compile(expr))
# ## Window functions
#
# Window functions in SQL allow you to write expressions that involve
# possibly-ordered groups of a table. Each window function involves one of
# the following:
#
# - An analytic function. Most aggregate functions are valid analytic
# functions, and there are additional ones such as `LEAD`, `LAG`,
# `NTILE`, and others.
# - A `PARTITION BY` clause. This may be omitted.
# - An `ORDER BY` clause. This may be omitted for many functions.
# - A window frame clause. The default is to use the entire partition.
#
# So you may see SQL like:
#
# ``` sql
# AVG(value) OVER (PARTITION BY key1)
# ```
#
# Or simply
#
# ``` sql
# AVG(value) OVER ()
# ```
#
# Ibis will automatically write window clauses when you use aggregate
# functions in a non-aggregate context. Suppose you wanted to subtract the
# mean of a column from itself:
expr = t.mutate(two_demean=t.two - t.two.mean())
print(ibis.impala.compile(expr))
# If you use `mutate` in conjunction with `group_by`, it will add a
# `PARTITION BY` to the `OVER` specification:
# +
expr = (t.group_by('one')
.mutate(two_demean=t.two - t.two.mean()))
print(ibis.impala.compile(expr))
# -
# For functions like `LAG` that require an ordering, we can add an
# `order_by` call:
# +
expr = (t.group_by('one')
.order_by(t.two) .mutate(two_first_diff=t.two - t.two.lag()))
print(ibis.impala.compile(expr))
# -
# For more precision, you can create a `Window` object that also includes
# a window frame clause:
w = ibis.window(group_by='one', preceding=5, following=5)
expr = t.mutate(group_demeaned=t.two - t.two.mean().over(w))
print(ibis.impala.compile(expr))
# ## Top-K operations
#
# A common SQL idiom is the "top-K" or "top-N" operation: subsetting a
# dimension by aggregate statistics:
#
# ``` sql
# SELECT key1, count(*) AS `count`
# FROM table1
# GROUP BY 1
# ORDER BY `count` DESC
# LIMIT 10
# ```
#
# Ibis has a special analytic expression `topk`:
expr = t1.key1.topk(10)
# This can be evaluated directly, yielding the above query:
print(ibis.impala.compile(expr))
# You can also use `expr` as a filter:
expr2 = t1[expr]
print(ibis.impala.compile(expr2))
# ## Date / time data
#
# See `Timestamp methods <api.timestamp>`{.interpreted-text role="ref"}
# for a table of available date/time methods.
#
# For example, we can do:
# +
expr = events.mutate(year=events.ts.year(),
month=events.ts.month())
print(ibis.impala.compile(expr))
# -
# ### Casting to date / time types
#
# In many cases, you can convert string values to datetime / timestamp
# with `strings.cast('timestamp')`, but you may have to do some more
# reconnaissance into the data if this does not work.
#
# ### Intervals
#
# Ibis has a set of interval APIs that allow you to do date/time
# arithmetic. For example:
expr = events[events.ts > (ibis.now() - ibis.interval(years=1))]
print(ibis.impala.compile(expr))
# The implementation of each timedelta offset will depend on the query
# engine.
#
# ## Buckets and histograms
#
# To appear.
#
# ## Unions
#
# SQL dialects often support two kinds of `UNION` operations:
#
# - `UNION`: the combination of *distinct* rows from each table.
# - `UNION ALL`: the combination of all rows from each table, whether or
# not they are distinct.
#
# The Ibis `union` function by distinct is a `UNION ALL`, and you can set
# `distinct=True` to get the normal `UNION` behavior:
# +
expr1 = t1.limit(10)
expr2 = t1.limit(10, offset=10)
expr = expr1.union(expr2)
print(ibis.impala.compile(expr))
# -
# ## Esoterica
#
# This area will be the spillover for miscellaneous SQL concepts and how
# queries featuring them can be ported to Ibis.
#
# ### Common table expressions (CTEs)
#
# The simplest SQL CTE is a SQL statement that is used multiple times in a
# `SELECT` query, which can be "factored" out using the `WITH` keyword:
#
# ``` sql
# WITH t0 AS (
# SELECT region, kind, sum(amount) AS total
# FROM purchases
# GROUP BY 1, 2
# )
# SELECT t0.region, t0.total - t1.total
# FROM t0
# INNER JOIN t0 t1
# ON t0.region = t1.region
# WHERE t0.kind = 'foo' AND t1.kind = 'bar'
# ```
#
# Explicit CTEs are not necessary with Ibis. Let's look at an example
# involving joining an aggregated table on itself after filtering:
# +
purchases = ibis.table([('region', 'string'),
('kind', 'string'),
('user', 'int64'),
('amount', 'float')], 'purchases')
metric = purchases.amount.sum().name('total')
agged = (purchases.group_by(['region', 'kind'])
.aggregate(metric))
left = agged[agged.kind == 'foo']
right = agged[agged.kind == 'bar']
result = (left.join(right, left.region == right.region)
[left.region, (left.total - right.total).name('diff')])
# -
# Ibis automatically creates a CTE for `agged`:
print(ibis.impala.compile(result))
| docs/ibis-for-sql-programmers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import operator as op
from functools import reduce
import numpy as np
pd.set_option("display.max_rows", 101)
# 1. Import confirmed, recovered and deaths data set for a given country
confirmed = pd.read_csv('DATA/COVID-19/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_confirmed_global.csv')
recovered = pd.read_csv('DATA/COVID-19/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_recovered_global.csv')
deaths = pd.read_csv('DATA/COVID-19/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_deaths_global.csv')
# 2. Pre-process the datasets
def reformat(df, country, col_name):
df = df[df['Country/Region'] == country]
index = df.iloc[0].name
df = df.T
df = df.rename({index: col_name}, axis = 1)
# get rid of columns that do not represent the number of recovered/confirmed/deaths
df = df.iloc[4:,:]
return df
#df = df[df[col_name] > 0]
# +
recovered = reformat(recovered, 'Switzerland', 'Recovered cases')
confirmed = reformat(confirmed, 'Switzerland', 'Confirmed cases')
deaths = reformat(deaths, 'Switzerland', 'Deaths cases')
recovered.tail()
# -
# 3. Create new data frame recovered per day so we know how many people recovered from the epidemic on a given day
#
# **problem** : Here, I see that recovered_daily contains a negative value, therefore recovered is not fully incremental. ??? Change dataset?
# +
recovered_daily = recovered.diff()
recovered_daily.iloc[0,:] = 0
deaths_daily = deaths.diff()
deaths_daily.iloc[0] = 0
# -
# 4. Generate dataset infected representing the number of infected individuals at a given time t (not the new number of infected, number of actively infected individuals)
#
# $$\text{confirmed}(t)=\text{confirmed}(t)-\sum_{i=\text{day}_0}^{t}\text{recovered_daily(i)}-\sum_{i=\text{day}_0}^{t}\text{deaths_daily}(i)$$
# +
infected_ = []
days_ = []
# i: date (time t)
# row: row
acc_deaths = 0
acc_recovered = 0
for i, row in confirmed.iterrows():
acc_recovered += recovered_daily.loc[i]['Recovered cases']
acc_deaths += deaths_daily.loc[i]['Deaths cases']
diff = row['Confirmed cases'] - acc_recovered - acc_deaths
infected_.append(diff)
days_.append(i)
# -
# Careful, infected represents active number of infected, not new number of infected
# +
infected = pd.DataFrame({'Date': days_,'Infected Cases': infected_})
recovered = recovered.reset_index().rename({'index':'Date'}, axis = 1)
# -
recovered.plot()
ax = pd.DataFrame(recovered['Recovered cases'].diff()).iloc[55:,:].plot.hist(bins=70,figsize=(10,7))
ax.set(xlabel="Number of individuals recovered")
#pd.DataFrame(recovered['Recovered cases'].diff()).iloc[55:,:].plot.hist(xlabel="Number of Recovered")
#(title="Frequency per number of daily recovered counted since 55th day (March 17th)")#.groupby()
# in order to see zeros from when people started recovering
ax = pd.DataFrame(recovered['Recovered cases'].diff()).iloc[55:,:].plot(figsize=(10,7))
ax.set(xlabel='Day')
# Now we can start working on predicting our gammas.
# Recovered is cumulative, as expected (absorbing state in our SIR model) and infected represents the number of infected individual at a certain date
# +
data = recovered.merge(infected, on='Date')
data['Date'] = pd.to_datetime(data['Date'])
#df['DOB'] = pd.to_datetime(df.DOB)
data = data.set_index('Date')
data['Recovered Daily'] = recovered_daily
data['Deaths Daily'] = deaths_daily
data = data[['Infected Cases', 'Recovered Daily', 'Deaths Daily']]
data.head()
# -
# ## Create X and y to export data
data = data.reset_index()
dataframe = pd.concat([data['Date'],data['Recovered Daily'], data['Deaths Daily'], data['Infected Cases'].shift(7),
data['Infected Cases'].shift(8),data['Infected Cases'].shift(9),
data['Infected Cases'].shift(10),data['Infected Cases'].shift(11),
data['Infected Cases'].shift(12),data['Infected Cases'].shift(13),
data['Infected Cases'].shift(14),data['Infected Cases'].shift(15),
data['Infected Cases'].shift(16),data['Infected Cases'].shift(17),
data['Infected Cases'].shift(18),data['Infected Cases'].shift(19),
data['Infected Cases'].shift(20),data['Infected Cases'].shift(21)], axis=1)
dataframe
# To go on I think it would be better to have a larger data set
# +
# create lagged dataset
dataframe.columns = ['Date', 'Recovered Daily', 'Deaths Daily','Infected cases t-7','Infected cases t-8','Infected cases t-9',
'Infected cases t-10','Infected cases t-11','Infected cases t-12','Infected cases t-13',
'Infected cases t-14','Infected cases t-15','Infected cases t-16','Infected cases t-17',
'Infected cases t-18','Infected cases t-19','Infected cases t-20','Infected cases t-21']
# Keep data such that we have value t-21 at any given day t
dataframe = dataframe[~np.isnan(dataframe['Infected cases t-21'])]
# -
dataframe = dataframe.set_index('Date')
# ### Delete first 15 days (noise data)
dataframe_new = dataframe.iloc[15:,:].copy()
# +
y_recovered = dataframe_new['Recovered Daily']
y_deaths = dataframe_new['Deaths Daily']
X = dataframe_new.iloc[:,2:]
# -
# ### Apply changes on Dataframe when recovered value equals 0
df = dataframe_new.copy()
df['Recovered Daily'].reset_index()
# +
# don't forget to round up
recovered = df['Recovered Daily'].reset_index()
#for i, el in recovered.iterrows():
#for i in range(recovered.shape[0]-1):
i = 0
# ignore last two elements from list as they are equal to zero so we cannot smooth them
while(i < recovered.shape[0]-2):
if (i > 24) & (recovered.iloc[i]['Recovered Daily'] == 0):
#print('hey')
j = 0
denom = 1
acc = 0
while(recovered.iloc[i+j]['Recovered Daily'] == 0):
#print('i:', i, 'j:', j)
j+=1
denom+=1
div = np.ceil(recovered.iloc[i+j]['Recovered Daily']/denom)
#print(div, denom)
recovered.iloc[i:i+j+1,1] = div
i +=j
else:
i+=1
#mean = round(recovered.iloc[i+1,1]/2)
#print(i,mean)
#recovered.iloc[i,1] = mean
#recovered.iloc[i+1, 1] = mean
#acc.append(mean)
#acc.append(mean)
# -
recovered
y_recovered_smoothed = recovered.set_index('Date').rename({'Recovered Daily': 'Recovered Daily Smoothed'}, axis=1)
data = pd.concat([X, y_recovered, y_deaths, y_recovered_smoothed], axis=1)
data.to_csv('data.csv')
| .ipynb_checkpoints/0. Data Cleaning-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="mt9dL5dIir8X"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab_type="code" id="ufPx7EiCiqgR" colab={}
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" deletable=true editable=true id="ucMoYase6URl"
# # Загрузка изображений
# + [markdown] colab_type="text" id="_Wwu5SXZmEkB"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/load_data/images"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />Смотрите на TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ru/tutorials/load_data/images.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Запустите в Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ru/tutorials/load_data/images.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />Изучайте код на GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ru/tutorials/load_data/images.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Скачайте ноутбук</a>
# </td>
# </table>
# + [markdown] id="3W5HaXTbpf4H" colab_type="text"
# Note: Вся информация в этом разделе переведена с помощью русскоговорящего Tensorflow сообщества на общественных началах. Поскольку этот перевод не является официальным, мы не гарантируем что он на 100% аккуратен и соответствует [официальной документации на английском языке](https://www.tensorflow.org/?hl=en). Если у вас есть предложение как исправить этот перевод, мы будем очень рады увидеть pull request в [tensorflow/docs](https://github.com/tensorflow/docs) репозиторий GitHub. Если вы хотите помочь сделать документацию по Tensorflow лучше (сделать сам перевод или проверить перевод подготовленный кем-то другим), напишите нам на [<EMAIL> list](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-ru).
# + [markdown] colab_type="text" id="Oxw4WahM7DU9"
# В этом руководстве приведен простой пример загрузки датасета изображений с использованием `tf.data`.
#
# Набор данных, используемый в этом примере, распространяется в виде папок изображений, с одним классом изображений в каждой папке.
# + [markdown] colab_type="text" deletable=true editable=true id="hoQQiZDB6URn"
# ## Setup
# + colab_type="code" id="QGXxBuPyKJw1" colab={}
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# # %tensorflow_version существует только в Colab.
# %tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
# + colab_type="code" id="KT6CcaqgQewg" colab={}
AUTOTUNE = tf.data.experimental.AUTOTUNE
# + [markdown] colab_type="text" id="rxndJHNC8YPM"
# ## Скачайте и проверьте набор данных
# + [markdown] colab_type="text" deletable=true editable=true id="wO0InzL66URu"
# ### Получите изображения
#
# Перед тем как начать любое обучение вам нужен набор изображений для обучения нейронной сети новым классам которые вы хотите распознавать. Вы уже создали архив распространяемых по свободной лицензии фото цветов для первоначального использования:
# + colab_type="code" id="rN-Pc6Zd6awg" colab={}
import pathlib
data_root_orig = tf.keras.utils.get_file(origin='https://storage.googleapis.com/download.tensorflow.org/example_images/flower_photos.tgz',
fname='flower_photos', untar=True)
data_root = pathlib.Path(data_root_orig)
print(data_root)
# + [markdown] colab_type="text" id="rFkFK74oO--g"
# После скачивания 218MB, вам теперь доступна копия изображений цветов:
# + colab_type="code" id="7onR_lWE7Njj" colab={}
for item in data_root.iterdir():
print(item)
# + colab_type="code" id="4yYX3ZRqGOuq" colab={}
import random
all_image_paths = list(data_root.glob('*/*'))
all_image_paths = [str(path) for path in all_image_paths]
random.shuffle(all_image_paths)
image_count = len(all_image_paths)
image_count
# + colab_type="code" id="t_BbYnLjbltQ" colab={}
all_image_paths[:10]
# + [markdown] colab_type="text" id="vkM-IpB-6URx"
# ### Просмотрите изображения
# Сейчас давайте быстро просмотрим парочку изображений, чтобы вы знали с чем имеете дело:
# + colab_type="code" id="wNGateQJ6UR1" colab={}
import os
attributions = (data_root/"LICENSE.txt").open(encoding='utf-8').readlines()[4:]
attributions = [line.split(' CC-BY') for line in attributions]
attributions = dict(attributions)
# + colab_type="code" id="jgowG2xu88Io" colab={}
import IPython.display as display
def caption_image(image_path):
image_rel = pathlib.Path(image_path).relative_to(data_root)
return "Image (CC BY 2.0) " + ' - '.join(attributions[str(image_rel)].split(' - ')[:-1])
# + colab_type="code" id="YIjLi-nX0txI" colab={}
for n in range(3):
image_path = random.choice(all_image_paths)
display.display(display.Image(image_path))
print(caption_image(image_path))
print()
# + [markdown] colab_type="text" id="OaNOr-co3WKk"
# ### Определите метку для каждого изображения
# + [markdown] colab_type="text" id="-weOQpDw2Jnu"
# Выведите на экран все доступные метки:
# + colab_type="code" id="ssUZ7Qh96UR3" colab={}
label_names = sorted(item.name for item in data_root.glob('*/') if item.is_dir())
label_names
# + [markdown] colab_type="text" id="9l_JEBql2OzS"
# Присвойте индекс каждой метке:
# + colab_type="code" id="Y8pCV46CzPlp" colab={}
label_to_index = dict((name, index) for index, name in enumerate(label_names))
label_to_index
# + [markdown] colab_type="text" id="VkXsHg162T9F"
# Создайте список всех файлов и индексов их меток:
# + colab_type="code" id="q62i1RBP4Q02" colab={}
all_image_labels = [label_to_index[pathlib.Path(path).parent.name]
for path in all_image_paths]
print("First 10 labels indices: ", all_image_labels[:10])
# + [markdown] colab_type="text" id="i5L09icm9iph"
# ### Загрузите и отформатируйте изображения
# + [markdown] colab_type="text" id="SbqqRUS79ooq"
# TensorFlow включает все инструменты которые вам могут понадобиться для загрузки и обработки изображений:
# + colab_type="code" id="jQZdySHvksOu" colab={}
img_path = all_image_paths[0]
img_path
# + [markdown] colab_type="text" id="2t2h2XCcmK1Y"
# Вот сырые данные:
# + colab_type="code" id="LJfkyC_Qkt7A" colab={}
img_raw = tf.io.read_file(img_path)
print(repr(img_raw)[:100]+"...")
# + [markdown] colab_type="text" id="opN8AVc8mSbz"
# Преобразуйте ее в тензор изображения:
# + colab_type="code" id="Tm0tdrlfk0Bb" colab={}
img_tensor = tf.image.decode_image(img_raw)
print(img_tensor.shape)
print(img_tensor.dtype)
# + [markdown] colab_type="text" id="3k-Of2Tfmbeq"
# Измените ее размер для вашей модели:
# + colab_type="code" id="XFpz-3_vlJgp" colab={}
img_final = tf.image.resize(img_tensor, [192, 192])
img_final = img_final/255.0
print(img_final.shape)
print(img_final.numpy().min())
print(img_final.numpy().max())
# + [markdown] colab_type="text" id="aCsAa4Psl4AQ"
# Оберните их в простые функции для будущего использования.
# + colab_type="code" id="HmUiZJNU73vA" colab={}
def preprocess_image(image):
image = tf.image.decode_jpeg(image, channels=3)
image = tf.image.resize(image, [192, 192])
image /= 255.0 # normalize to [0,1] range
return image
# + colab_type="code" id="einETrJnO-em" colab={}
def load_and_preprocess_image(path):
image = tf.io.read_file(path)
return preprocess_image(image)
# + colab_type="code" id="3brWQcdtz78y" colab={}
import matplotlib.pyplot as plt
image_path = all_image_paths[0]
label = all_image_labels[0]
plt.imshow(load_and_preprocess_image(img_path))
plt.grid(False)
plt.xlabel(caption_image(img_path))
plt.title(label_names[label].title())
print()
# + [markdown] colab_type="text" id="n2TCr1TQ8pA3"
# ## Постройте `tf.data.Dataset`
# + [markdown] colab_type="text" id="6H9Z5Mq63nSH"
# ### Датасет изображений
# + [markdown] colab_type="text" id="GN-s04s-6Luq"
# Простейший способ построения `tf.data.Dataset` это использование метода `from_tensor_slices`.
#
# Нарезка массива строк, приводит к датасету строк:
# + colab_type="code" id="6oRPG3Jz3ie_" colab={}
path_ds = tf.data.Dataset.from_tensor_slices(all_image_paths)
# + [markdown] colab_type="text" id="uML4JeMmIAvO"
# Параметры `shapes` и `types` описывают содержимое каждого элемента датасета. В этом случае у нас множество скалярных двоичных строк
# + colab_type="code" id="mIsNflFbIK34" colab={}
print(path_ds)
# + [markdown] colab_type="text" id="ZjyGcM8OwBJ2"
# Сейчас создадим новый датасет который загружает и форматирует изображения на лету пройдясь с `preprocess_image` по датасету с путями к файлам.
# + colab_type="code" id="D1iba6f4khu-" colab={}
image_ds = path_ds.map(load_and_preprocess_image, num_parallel_calls=AUTOTUNE)
# + colab_type="code" id="JLUPs2a-lEEJ" colab={}
import matplotlib.pyplot as plt
plt.figure(figsize=(8,8))
for n, image in enumerate(image_ds.take(4)):
plt.subplot(2,2,n+1)
plt.imshow(image)
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.xlabel(caption_image(all_image_paths[n]))
plt.show()
# + [markdown] colab_type="text" id="P6FNqPbxkbdx"
# ### Датасет пар `(image, label)`
# + [markdown] colab_type="text" id="YgvrWLKG67-x"
# Используя тот же метод `from_tensor_slices` вы можете собрать датасет меток:
# + colab_type="code" id="AgBsAiV06udj" colab={}
label_ds = tf.data.Dataset.from_tensor_slices(tf.cast(all_image_labels, tf.int64))
# + colab_type="code" id="HEsk5nN0vyeX" colab={}
for label in label_ds.take(10):
print(label_names[label.numpy()])
# + [markdown] colab_type="text" id="jHjgrEeTxyYz"
# Поскольку датасеты следуют в одном и том же порядке, вы можете просто собрать их вместе при помощи функции zip в набор данных пары `(image, label)`:
# + colab_type="code" id="AOEWNMdQwsbN" colab={}
image_label_ds = tf.data.Dataset.zip((image_ds, label_ds))
# + [markdown] colab_type="text" id="yA2F09SJLMuM"
# `Shapes` и `types` нового датасета это кортежи размерностей и типов описывающие каждое поле:
# + colab_type="code" id="DuVYNinrLL-N" colab={}
print(image_label_ds)
# + [markdown] colab_type="text" id="2WYMikoPWOQX"
# Примечание. Если у вас есть такие массивы, как «all_image_labels» и «all_image_paths», альтернативой «tf.data.dataset.Dataset.zip» является срез (slice) пары массивов.
# + colab_type="code" id="HOFwZI-2WhzV" colab={}
ds = tf.data.Dataset.from_tensor_slices((all_image_paths, all_image_labels))
# Кортежи распаковываются в позиционные аргументы отображаемой функции
def load_and_preprocess_from_path_label(path, label):
return load_and_preprocess_image(path), label
image_label_ds = ds.map(load_and_preprocess_from_path_label)
image_label_ds
# + [markdown] colab_type="text" id="vYGCgJuR_9Qp"
# ### Базовые способы обучения
# + [markdown] colab_type="text" id="wwZavzgsIytz"
# Для обучения модели на этом датасете, вам необходимо, чтобы данные:
#
# * Были хорошо перемешаны.
# * Были упакованы.
# * Повторялись вечно.
# * Пакеты должны быть доступны как можно скорее.
#
# Эти свойства могут быть легко добавлены с помощью `tf.data` api.
# + colab_type="code" id="uZmZJx8ePw_5" colab={}
BATCH_SIZE = 32
# Установка размера буфера перемешивания, равного набору данных, гарантирует
# полное перемешивание данных.
ds = image_label_ds.shuffle(buffer_size=image_count)
ds = ds.repeat()
ds = ds.batch(BATCH_SIZE)
# `prefetch` позволяет датасету извлекать пакеты в фоновом режиме, во время обучения модели.
ds = ds.prefetch(buffer_size=AUTOTUNE)
ds
# + [markdown] colab_type="text" id="6JsM-xHiFCuW"
# Здесь необходимо отметить несколько вещей:
#
# 1. Важна последовательность действий.
#
# * `.shuffle` после `.repeat` перемешает элементы вне границ эпох (некоторые элементы будут увидены дважды в то время как другие ни разу).
# * `.shuffle` после `.batch` перемешает порядок пакетов, но не перемешает элементы внутри пакета.
#
# 1. Используйте `buffer_size` такого же размера как и датасет для полного перемешивания. Вплоть до размера набора данных, большие значения обеспечивают лучшую рандомизацию, но используют больше памяти.
#
# 1. Из буфера перемешивания не выбрасываются элементы пока он не заполнится. Поэтому большой размер `buffer_size` может стать причиной задержки при запуске `Dataset`.
#
# 1. Перемешанный датасет не сообщает об окончании, пока буфер перемешивания полностью не опустеет. `Dataset` перезапускается с помощью` .repeat`, вызывая еще одно ожидание заполнения буфера перемешивания.
#
# Последний пункт может быть решен использованием метода `tf.data.Dataset.apply` вместе со слитой функцией `tf.data.experimental.shuffle_and_repeat`:
# + colab_type="code" id="Ocr6PybXNDoO" colab={}
ds = image_label_ds.apply(
tf.data.experimental.shuffle_and_repeat(buffer_size=image_count))
ds = ds.batch(BATCH_SIZE)
ds = ds.prefetch(buffer_size=AUTOTUNE)
ds
# + [markdown] colab_type="text" id="GBBZMSuAmQVL"
# ### Передайте датасет в модель
#
# Получите копию MobileNet v2 из `tf.keras.applications`.
#
# Она будет использована для простого примера передачи обучения (transfer learning).
#
# Установите веса MobileNet необучаемыми:
# + colab_type="code" id="KbJrXn9omO_g" colab={}
mobile_net = tf.keras.applications.MobileNetV2(input_shape=(192, 192, 3), include_top=False)
mobile_net.trainable=False
# + [markdown] colab_type="text" id="Y7NVWiLF3Vbf"
# Эта модель предполагает нормализацию входных данных в диапазоне `[-1,1]`:
#
# ```
# help(keras_applications.mobilenet_v2.preprocess_input)
# ```
#
# <pre>
# ...
# Эта функция применяет препроцессинг "Inception" который преобразует
# RGB значения из [0, 255] в [-1, 1]
# ...
# </pre>
# + [markdown] colab_type="text" id="CboYya6LmdQI"
# Перед передачей входных данных в модель MobilNet, вам нужно конвертировать их из диапазона `[0,1]` в `[-1,1]`:
# + colab_type="code" id="SNOkHUGv3FYq" colab={}
def change_range(image,label):
return 2*image-1, label
keras_ds = ds.map(change_range)
# + [markdown] colab_type="text" id="QDzZ3Nye5Rpv"
# MobileNet возвращает `6x6` сетку признаков для каждого изображения.
#
# Передайте ей пакет изображений чтобы увидеть:
# + colab_type="code" id="OzAhGkEK6WuE" colab={}
# Датасету может понадобиться несколько секунд для старта пока заполняется буфер перемешивания.
image_batch, label_batch = next(iter(keras_ds))
# + colab_type="code" id="LcFdiWpO5WbV" colab={}
feature_map_batch = mobile_net(image_batch)
print(feature_map_batch.shape)
# + [markdown] colab_type="text" id="vrbjEvaC5XmU"
# Постройте модель обернутую вокруг MobileNet и используйте `tf.keras.layers.GlobalAveragePooling2D` для усреднения по этим размерностям пространства перед выходным слоем `tf.keras.layers.Dense`:
# + colab_type="code" id="X0ooIU9fNjPJ" colab={}
model = tf.keras.Sequential([
mobile_net,
tf.keras.layers.GlobalAveragePooling2D(),
tf.keras.layers.Dense(len(label_names), activation = 'softmax')])
# + [markdown] colab_type="text" id="foQYUJs97V4V"
# Сейчас он производит выходные данные ожидаемых размеров:
# + colab_type="code" id="1nwYxvpj7ZEf" colab={}
logit_batch = model(image_batch).numpy()
print("min logit:", logit_batch.min())
print("max logit:", logit_batch.max())
print()
print("Shape:", logit_batch.shape)
# + [markdown] colab_type="text" id="pFc4I_J2nNOJ"
# Скомпилируйте модель чтобы описать процесс обучения:
# + colab_type="code" id="ZWGqLEWYRNvv" colab={}
model.compile(optimizer=tf.keras.optimizers.Adam(),
loss='sparse_categorical_crossentropy',
metrics=["accuracy"])
# + [markdown] colab_type="text" id="tF1mO6haBOSd"
# Есть две переменные для обучения - Dense `weights` и `bias`:
# + colab_type="code" id="pPQ5yqyKBJMm" colab={}
len(model.trainable_variables)
# + colab_type="code" id="kug5Wg66UJjl" colab={}
model.summary()
# + [markdown] colab_type="text" id="f_glpYZ-nYC_"
# Вы готовы обучать модель.
#
# Отметим, что для демонстрационных целей вы запустите только 3 шага на эпоху, но обычно вам нужно указать действительное число шагов, как определено ниже, перед передачей их в `model.fit()`:
# + colab_type="code" id="AnXPRNWoTypI" colab={}
steps_per_epoch=tf.math.ceil(len(all_image_paths)/BATCH_SIZE).numpy()
steps_per_epoch
# + colab_type="code" id="q_8sabaaSGAp" colab={}
model.fit(ds, epochs=1, steps_per_epoch=3)
# + [markdown] colab_type="text" id="UMVnoBcG_NlQ"
# ## Производительность
#
# Примечание: Этот раздел лишь показывает пару простых приемов которые могут помочь производительности. Для более глубокого изучения см. [Производительность входного конвейера](https://www.tensorflow.org/guide/performance/datasets).
#
# Простые конвейеры, использованные выше, прочитывают каждый файл отдельно во время каждой эпохи. Это подходит для локального обучения на CPU, может быть недостаточно для обучения на GPU и абсолютно неприемлемо для любого вида распределенного обучения.
# + [markdown] colab_type="text" id="oNmQqgGhLWie"
# Чтобы исследовать производительность наших датасетов, сперва постройте простую функцию:
# + colab_type="code" id="_gFVe1rp_MYr" colab={}
import time
default_timeit_steps = 2*steps_per_epoch+1
def timeit(ds, steps=default_timeit_steps):
overall_start = time.time()
# Выберем один пакет для передачи в пайплайн (заполнение буфера перемешивания),
# перед запуском таймера
it = iter(ds.take(steps+1))
next(it)
start = time.time()
for i,(images,labels) in enumerate(it):
if i%10 == 0:
print('.',end='')
print()
end = time.time()
duration = end-start
print("{} batches: {} s".format(steps, duration))
print("{:0.5f} Images/s".format(BATCH_SIZE*steps/duration))
print("Total time: {}s".format(end-overall_start))
# + [markdown] colab_type="text" id="TYiOr4vdLcNX"
# Производительность данного датасета равна:
# + colab_type="code" id="ZDxLwMJOReVe" colab={}
ds = image_label_ds.apply(
tf.data.experimental.shuffle_and_repeat(buffer_size=image_count))
ds = ds.batch(BATCH_SIZE).prefetch(buffer_size=AUTOTUNE)
ds
# + colab_type="code" id="IjouTJadRxyp" colab={}
timeit(ds)
# + [markdown] colab_type="text" id="HsLlXMO7EWBR"
# ### Кеш
# + [markdown] colab_type="text" id="lV1NOn2zE2lR"
# Используйте tf.data.Dataset.cache, чтобы с легкостью кэшировать вычисления от эпохи к эпохе. Это особенно эффективно если данные помещаются в память.
#
# Здесь изображения кэшируются после предварительной обработки (перекодирования и изменения размера):
# + colab_type="code" id="qj_U09xpDvOg" colab={}
ds = image_label_ds.cache()
ds = ds.apply(
tf.data.experimental.shuffle_and_repeat(buffer_size=image_count))
ds = ds.batch(BATCH_SIZE).prefetch(buffer_size=AUTOTUNE)
ds
# + colab_type="code" id="rdxpvQ7VEo3y" colab={}
timeit(ds)
# + [markdown] colab_type="text" id="usIv7MqqZQps"
# Одним из недостатков использования кэша памяти является то, что кэш должен перестраиваться при каждом запуске, давая одинаковую начальную задержку при каждом запуске датасета:
# + colab_type="code" id="eKX6ergKb_xd" colab={}
timeit(ds)
# + [markdown] colab_type="text" id="jUzpG4lYNkN-"
# Если данные не помещаются в памяти, используйте файл кэша:
# + colab_type="code" id="vIvF8K4GMq0g" colab={}
ds = image_label_ds.cache(filename='./cache.tf-data')
ds = ds.apply(
tf.data.experimental.shuffle_and_repeat(buffer_size=image_count))
ds = ds.batch(BATCH_SIZE).prefetch(1)
ds
# + colab_type="code" id="eTIj6IOmM4yA" colab={}
timeit(ds)
# + [markdown] colab_type="text" id="qqo3dyB0Z4t2"
# Также у файла кеша есть преимущество использования быстрого перезапуска датасета без перестраивания кеша. Обратите внимание, насколько быстрее это происходит во второй раз:
# + colab_type="code" id="hZhVdR8MbaUj" colab={}
timeit(ds)
# + [markdown] colab_type="text" id="WqOVlf8tFrDU"
# ### Файл TFRecord
# + [markdown] colab_type="text" id="y1llOTwWFzmR"
# #### Необработанные данные - изображения
#
# TFRecord файлы - это простой формат для хранения двоичных блобов (blob). Упаковывая несколько примеров в один файл, TensorFlow может читать несколько элементов за раз, что особенно важно для производительности особенно при использовании удаленного сервиса хранения, такого как GCS.
#
# Сперва построим файл TFRecord из необработанных данных изображений:
# + colab_type="code" id="EqtARqKuHQLu" colab={}
image_ds = tf.data.Dataset.from_tensor_slices(all_image_paths).map(tf.io.read_file)
tfrec = tf.data.experimental.TFRecordWriter('images.tfrec')
tfrec.write(image_ds)
# + [markdown] colab_type="text" id="flR2GXWFKcO1"
# Затем построим датасет, который прочитывает файл TFRecord и обрабатывает изображения с использованием функции `preprocess_image`, которую вы задали ранее:
# + colab_type="code" id="j9PVUL2SFufn" colab={}
image_ds = tf.data.TFRecordDataset('images.tfrec').map(preprocess_image)
# + [markdown] colab_type="text" id="cRp1eZDRKzyN"
# Объедините этот датасет с датасетом меток, который вы определили ранее, чтобы получить пару из `(image,label)`:
# + colab_type="code" id="7XI_nDU2KuhS" colab={}
ds = tf.data.Dataset.zip((image_ds, label_ds))
ds = ds.apply(
tf.data.experimental.shuffle_and_repeat(buffer_size=image_count))
ds=ds.batch(BATCH_SIZE).prefetch(AUTOTUNE)
ds
# + colab_type="code" id="3ReSapoPK22E" colab={}
timeit(ds)
# + [markdown] colab_type="text" id="wb7VyoKNOMms"
# Это медленнее `cache` версии, поскольку обработанные изображения не кешируются.
# + [markdown] colab_type="text" id="NF9W-CTKkM-f"
# #### Сериализованные тензоры
# + [markdown] colab_type="text" id="J9HzljSPkxt0"
# Чтобы сохранить некоторый препроцессинг в файл TFRecord сперва, как и ранее, создайте датасет обработанных изображений:
# + colab_type="code" id="OzS0Azukkjyw" colab={}
paths_ds = tf.data.Dataset.from_tensor_slices(all_image_paths)
image_ds = paths_ds.map(load_and_preprocess_image)
image_ds
# + [markdown] colab_type="text" id="onWOwLpYlzJQ"
# Сейчас вместо датасета строк `.jpeg`, у вас датасет тензоров.
#
# Чтобы сериализовать это в файл TFRecord сперва сконвертируйте датасет тензоров в датасет строк:
# + colab_type="code" id="xxZSwnRllyf0" colab={}
ds = image_ds.map(tf.io.serialize_tensor)
ds
# + colab_type="code" id="w9N6hJWAkKPC" colab={}
tfrec = tf.data.experimental.TFRecordWriter('images.tfrec')
tfrec.write(ds)
# + [markdown] colab_type="text" id="OlFc9dJSmcx0"
# С кешированным препроцессингом данные могут быть выгружены из TFrecord файла очень эффективно - не забудьте только десериализовать тензор перед использованием:
# + colab_type="code" id="BsqFyTBFmSCZ" colab={}
ds = tf.data.TFRecordDataset('images.tfrec')
def parse(x):
result = tf.io.parse_tensor(x, out_type=tf.float32)
result = tf.reshape(result, [192, 192, 3])
return result
ds = ds.map(parse, num_parallel_calls=AUTOTUNE)
ds
# + [markdown] colab_type="text" id="OPs_sLV9pQg5"
# Сейчас добавьте метки и примените те же стандартные операции, что и ранее:
# + colab_type="code" id="XYxBwaLYnGop" colab={}
ds = tf.data.Dataset.zip((ds, label_ds))
ds = ds.apply(
tf.data.experimental.shuffle_and_repeat(buffer_size=image_count))
ds=ds.batch(BATCH_SIZE).prefetch(AUTOTUNE)
ds
# + colab_type="code" id="W8X6RmGan1-P" colab={}
timeit(ds)
| site/ru/tutorials/load_data/images.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
import numpy as np
import pandas as pd
from collections import Counter, defaultdict
import csv
import math
import operator
import data_processing_training
coupon_Id = "51800000050"
df_coupon = pd.read_csv('coupon.csv', dtype={'COUPON_UPC': str, 'CAMPAIGN': str, 'PRODUCT_ID': str})
campaigns = data_processing_training.get_campaigns_for_coupon(coupon_Id, df_coupon)
product_list = data_processing_training.get_products_for_coupon(coupon_Id, df_coupon)
del df_coupon
df_campaign_table = pd.read_csv('campaign_table.csv', dtype={'household_key': str, 'CAMPAIGN': str})
df_campaign_desc = pd.read_csv('campaign_desc.csv', dtype={'CAMPAIGN': str})
hh_start_dates = data_processing_training.get_households_for_campaigns(campaigns, df_campaign_table, df_campaign_desc)
len(hh_start_dates)
del df_campaign_table
hh_start_dates.head()
hh_start_dates.drop(columns=['DESCRIPTION_x', 'DESCRIPTION_y'], inplace=True)
hh_start_dates.head()
df_transactions = pd.read_csv('transaction_data.csv', dtype={'BASKET_ID': str, 'PRODUCT_ID': str, 'household_key': str, 'DAY': str})
len(df_transactions['BASKET_ID'].unique())
len(df_transactions['household_key'].unique())
hh_list = hh_start_dates['household_key'].unique()
transactions_hh = df_transactions[df_transactions['household_key'].isin(hh_list)]
len(transactions_hh)
len(df_transactions)
df_transactions = df_transactions.merge(hh_start_dates, on='household_key', how='left')
len(df_transactions)
df_transactions.head()
len(df_transactions[df_transactions['START_DAY'].isnull()])
#trans_merge['START_DAY'].fillna(10000, inplace=True)
#trans_merge = trans_merge[trans_merge['DAY'].astype(float) < trans_merge['START_DAY']]
len(df_transactions[df_transactions['START_DAY'].isnull()])
len(df_transactions[df_transactions['household_key'] == "2375"])
transactions_hh = df_transactions[~df_transactions['household_key'].isin(hh_list)]
transactions_hh.head()
trans_merge_2375 = df_transactions[df_transactions['household_key'] == "2375"]
trans_merge_2375.head()
len(df_transactions[df_transactions['household_key'] == "1364"])
len(trans_merge)
df_transactions['CUSTOMER_PAID'] = df_transactions['SALES_VALUE'] + df_transactions['COUPON_DISC']
len(df_transactions[df_transactions['BASKET_ID'] == "26984851472"])
df_transactions[df_transactions['BASKET_ID'] == "26984851472"]
df_grouped_basket = df_transactions.groupby(['household_key', 'BASKET_ID', 'DAY'])
df_grouped_basket_count = df_grouped_basket.size().reset_index()
df_grouped_basket_count.head()
df_grouped_basket_count.columns = ['household_key', 'BASKET_ID', 'DAY', 'PROD_PURCHASE_COUNT']
df_grouped_basket_count[df_grouped_basket_count['BASKET_ID'] == "26984851472"]
df_grouped_basket_sum = df_grouped_basket.sum().reset_index()
df_grouped_basket_sum.drop(['RETAIL_DISC', 'TRANS_TIME', 'COUPON_MATCH_DISC', 'START_DAY', 'END_DAY'], axis=1, inplace=True)
df_grouped_basket_sum[df_grouped_basket_sum['BASKET_ID'] == "26984851472"]
df_grouped_basket = df_grouped_basket.apply(
lambda x : 1 if len(set(x.PRODUCT_ID.tolist()) & set(product_list)) > 0 else 0
).reset_index().rename(columns={0:"label"})
df_grouped_basket[df_grouped_basket['BASKET_ID'] == "26984851472"]
df_grouped_basket_merge = df_grouped_basket_sum.merge(df_grouped_basket, on=["household_key", "BASKET_ID"]).reset_index(drop=True)
df_grouped_basket_merge[df_grouped_basket_merge['BASKET_ID'] == "26984851472"]
df_grouped_basket_merge = df_grouped_basket_merge.merge(df_grouped_basket_count, on=["household_key", "BASKET_ID"]).reset_index(drop=True)
df_grouped_basket_merge[df_grouped_basket_merge['BASKET_ID'] == "26984851472"]
df_grouped_basket_merge = df_grouped_basket_merge.drop(['DAY_x'], axis=1, errors="ignore")
df_grouped_basket_merge.rename(columns={'DAY_y': 'DAY'}, inplace=True)
df_grouped_basket_merge[df_grouped_basket_merge['BASKET_ID'] == "26984851472"]
#merging demographics
df_demographic = pd.read_csv('hh_demographic.csv', dtype={'household_key': str})
df_grouped_basket_merge = df_grouped_basket_merge.merge(df_demographic, on="household_key", how="left").reset_index(drop=True)
df_grouped_basket_merge[df_grouped_basket_merge['BASKET_ID'] == "26984851472"]
df_grouped_basket_merge.head()
| data_processing_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import keras
import json
from datetime import datetime
import numpy as np
import pandas as pd
from keras.preprocessing.sequence import pad_sequences
from keras.preprocessing.text import Tokenizer
import pickle
with open('pad_encode_text.pk', 'rb') as f:
pad_encode_text = pickle.load(f)
with open('pad_decode_text.pk', 'rb') as f:
pad_decode_text = pickle.load(f)
with open('tokenizer.pk', 'rb') as f:
tokenizer = pickle.load(f)
# all kinds parameters
min_length = 2
max_length = 20
VOC_SIZE = 10000
EMBEDDING_DIM = 100
HIDDEN_DIM = 256
batch_size = 64
epochs = 100
fit_size = 20000
# read embedding file
embeddings_index = {}
with open("./glove/glove.6B.100d.txt", 'r', encoding="utf8") as f:
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
# generate embedding matrix
embedding_matrix = np.zeros((VOC_SIZE + 1, EMBEDDING_DIM))
count = 0
for i in range(1, VOC_SIZE):
embedding_vector = embeddings_index.get(tokenizer.index_word[i])
if embedding_vector is not None:
count += 1
embedding_matrix[i] = embedding_vector
from keras.layers import Embedding
from keras.models import Model
from keras.layers import Input, LSTM, Dense
from keras.layers import TimeDistributed
from keras.layers import concatenate
from keras_self_attention import SeqSelfAttention
embedding_layer = Embedding(VOC_SIZE + 1,
EMBEDDING_DIM,
weights=[embedding_matrix],
input_length=max_length,
trainable=False)
# +
encoder_inputs = Input(shape=(max_length, ))
encoder_embedding = embedding_layer(encoder_inputs)
encoder_LSTM = LSTM(HIDDEN_DIM, return_sequences = True, return_state = True)
encoder_outputs, state_h, state_c = encoder_LSTM(encoder_embedding)
encoder_states = [state_h, state_c]
encoder_att = SeqSelfAttention(attention_type=SeqSelfAttention.ATTENTION_TYPE_MUL,
kernel_regularizer=keras.regularizers.l2(1e-4),
bias_regularizer=keras.regularizers.l1(1e-4),
attention_regularizer_weight=1e-4,
name='Attention')(encoder_outputs)
decoder_inputs = Input(shape=(max_length, ))
decoder_embedding = embedding_layer(decoder_inputs)
decoder_LSTM = LSTM(HIDDEN_DIM, return_sequences=True, return_state=True)
decoder_outputs, _, _, = decoder_LSTM(decoder_embedding, initial_state=encoder_states)
concate = concatenate([encoder_att, decoder_outputs], axis=-1)
dense = Dense(VOC_SIZE+1, activation='softmax')
outputs = dense(concate)
# -
model = Model([encoder_inputs, decoder_inputs], [outputs])
model.compile(optimizer='rmsprop', loss='categorical_crossentropy',
metrics=['accuracy'])
model.summary()
for epoch in range(epochs):
for i in range(int(pad_decode_text.shape[0]/fit_size)+1):
pad_encode_text_sample = pad_encode_text[i*fit_size:(i+1)*fit_size]
pad_decode_text_sample = pad_decode_text[i*fit_size:(i+1)*fit_size]
one_hot_target_text = np.zeros((len(pad_decode_text_sample), max_length, VOC_SIZE+1), dtype='bool')
for k, seqs in enumerate(pad_decode_text_sample):
for j, seq in enumerate(seqs):
if j > 0:
one_hot_target_text[k][j-1][seq] = 1
model.fit([pad_encode_text_sample, pad_decode_text_sample], one_hot_target_text,
batch_size=batch_size,
epochs=1,
validation_split=0.1)
model.save('s2s_attention_voc10000.h5')
| seq2seq_attention.ipynb |
# # Putting it all together (PyTorch)
# Install the Transformers and Datasets libraries to run this notebook.
# !pip install datasets transformers[sentencepiece]
# +
from transformers import AutoTokenizer
checkpoint = "distilbert-base-uncased-finetuned-sst-2-english"
tokenizer = AutoTokenizer.from_pretrained(checkpoint)
sequence = "I've been waiting for a HuggingFace course my whole life."
model_inputs = tokenizer(sequence)
# +
sequence = "I've been waiting for a HuggingFace course my whole life."
model_inputs = tokenizer(sequence)
# +
sequences = [
"I've been waiting for a HuggingFace course my whole life.",
"So have I!"
]
model_inputs = tokenizer(sequences)
# +
# Will pad the sequences up to the maximum sequence length
model_inputs = tokenizer(sequences, padding="longest")
# Will pad the sequences up to the model max length
# (512 for BERT or DistilBERT)
model_inputs = tokenizer(sequences, padding="max_length")
# Will pad the sequences up to the specified max length
model_inputs = tokenizer(sequences, padding="max_length", max_length=8)
# +
sequences = [
"I've been waiting for a HuggingFace course my whole life.",
"So have I!"
]
# Will truncate the sequences that are longer than the model max length
# (512 for BERT or DistilBERT)
model_inputs = tokenizer(sequences, truncation=True)
# Will truncate the sequences that are longer than the specified max length
model_inputs = tokenizer(sequences, max_length=8, truncation=True)
# +
sequences = [
"I've been waiting for a HuggingFace course my whole life.",
"So have I!"
]
# Returns PyTorch tensors
model_inputs = tokenizer(sequences, padding=True, return_tensors="pt")
# Returns TensorFlow tensors
model_inputs = tokenizer(sequences, padding=True, return_tensors="tf")
# Returns NumPy arrays
model_inputs = tokenizer(sequences, padding=True, return_tensors="np")
# +
sequence = "I've been waiting for a HuggingFace course my whole life."
model_inputs = tokenizer(sequence)
print(model_inputs["input_ids"])
tokens = tokenizer.tokenize(sequence)
ids = tokenizer.convert_tokens_to_ids(tokens)
print(ids)
# -
print(tokenizer.decode(model_inputs["input_ids"]))
print(tokenizer.decode(ids))
# +
import torch
from transformers import AutoTokenizer, AutoModelForSequenceClassification
checkpoint = "distilbert-base-uncased-finetuned-sst-2-english"
tokenizer = AutoTokenizer.from_pretrained(checkpoint)
model = AutoModelForSequenceClassification.from_pretrained(checkpoint)
sequences = [
"I've been waiting for a HuggingFace course my whole life.",
"So have I!"
]
tokens = tokenizer(sequences, padding=True, truncation=True, return_tensors="pt")
output = model(**tokens)
| course/chapter2/section6_pt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''base'': conda)'
# name: python3
# ---
# + [markdown] nteract={"transient": {"deleting": false}}
# ---
# author: <NAME> (<EMAIL>)
# ---
# + [markdown] cell_id="00000-bfbf62b3-38d2-465b-98fc-1c2e2a429dff" deepnote_cell_type="markdown" tags=[]
# The solution below uses an example dataset about the teeth of 10 guinea pigs at three Vitamin C dosage levels (in mg) with two delivery methods (orange juice vs. ascorbic acid). (See how to quickly load some sample data.)
# + cell_id="00001-663e0c93-5200-4700-9ef5-b6776235aa87" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=8 execution_start=1626633316972 source_hash="3efdab3d" tags=[]
from rdatasets import data
df = data('ToothGrowth')
# + [markdown] cell_id="00002-242fa0db-f6bd-4b8b-9994-93ae8a22fc40" deepnote_cell_type="markdown" tags=[]
# In this dataset, there are only two treatments (orange juice and ascorbic acid, in the variable `supp`). We can therefore perrform a two-sample $t$ test. But first we must filter the outcome variable `len` (tooth length) based on `supp`.
# + cell_id="00003-e88454b4-571c-4052-b76b-619675200cb1" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=810 execution_start=1626633316991 source_hash="d43df127" tags=[]
subjects_receiving_oj = df[df['supp']=='OJ']['len']
subjects_receiving_vc = df[df['supp']=='VC']['len']
import scipy.stats as stats
stats.ttest_ind( subjects_receiving_oj, subjects_receiving_vc, equal_var=False )
# + [markdown] cell_id="00003-9da37a6f-6bb5-435c-9c4f-7100f469dbed" deepnote_cell_type="markdown" tags=[]
# At the 5% significance level, we see that the length of the tooth does not differ between the two delivery methods. We assume that the model assumptions are met, but do not check that here.
#
# If there are multiple levels (two or more), you can apply the parametric ANOVA test which in this case will provide a similar $p$ value.
# + cell_id="00003-000f6552-92f3-428c-8418-874f7845f321" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=475 execution_start=1626633317799 source_hash="e74ff0ff" tags=[]
from statsmodels.formula.api import ols
model = ols('len ~ supp', data = df).fit()
import statsmodels.api as sm
sm.stats.anova_lm(model, typ=1)
# + [markdown] cell_id="00004-fd34f9b0-df8e-4434-a3de-1ecb876ab312" deepnote_cell_type="markdown" tags=[]
# We see the $p$ value in the final column is very similar.
#
# However, if the assumptions of ANOVA are not met, we can utilize a nonparametric approach via the Kruskal-Wallis Test. We use the filtered variables defined above and import the `kruskal` function from SciPy.
# + cell_id="00006-e5f5e711-02e3-4fb4-8097-b2d593416703" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=7 execution_start=1626633318283 source_hash="651fe4ce" tags=[]
from scipy.stats import kruskal
kruskal( subjects_receiving_oj, subjects_receiving_vc )
# + [markdown] cell_id="00007-dafc5d87-ac98-4796-b532-bbcf8566ba5a" deepnote_cell_type="markdown" tags=[]
# Similar to the previous results, the length of the tooth does not differ between the delivery methods at the 5% significance level.
| database/tasks/How to test for a treatment effect in a single factor design/Python, using SciPy and statsmodels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:MS-ML-P3]
# language: python
# name: conda-env-MS-ML-P3-py
# ---
# <center><img src='img/ms_logo.jpeg' height=40% width=40%></center>
#
#
# <center><h1>Building a Model to Predict Survival for Titanic Passengers</h1></center>
#
#
# **Welcome to _DS2: Introduction to Machine Learning_**! This course will be all about _predictive analytics_--that is, using data and algorithms to make accurate predictions. For our introductory exercise for this course, we're going to focus on the one of the areas where machine learning really shines--**_Classification_**. We're going to examine the data and build a simple model to predict whether or not a passenger survived the Titanic disaster. Here's the catch: before we use any machine learning, we're going to build a classifier by hand to gain an intuition about how classification actually works.
# <br>
# <br>
# <center><h2>The Gameplan</h2></center>
#
# We're going to start by building the simplest model possible, and then slowly add complexity as we notice patterns that can make our classifier more accurate.
#
# Recall that we've investigated this dataset before, in DS1. We're going to use our _Data Analysis_ and _Visualization_ skills from DS1 to investigate our dataset and see if we can find some patterns that we can use in our prediction algorithm. In order to successfully build a prediction algorithm, we'll use the following process:
#
# **1. Load and explore the data.**
# --We'll begin by reading our data into a dataframe, and then visualizing our data to see if we can find certain groups that had higher survival rates than others. At this step, we'll also remove the `Survived` column from the dataframe and store it in a separate variable.
#
# **2.Write a prediction function.**
# <br>
# -- We'll write a function that takes in a dataframe and predicts 0 (died) or 1(survived) for each passenger based on whatever we decide is important. This function should output a vector containing only 0's and 1's, where the first element is the prediction for the first passenger in the dataframe, the 2nd element is the prediction for the second passenger, etc.
#
# **3. Write an evaluation function.**
# <br>
# -- In order to evaluate how accurate our prediction function is, we'll need to track how it does. To do this, we'll create a _confusion matrix_. This matrix will exist as a dictionary that tracks the number of _True Positives_, _True Negatives_, _False Positives_, and _False Negatives_ our algorithm makes--don't worry if you haven't seen these terms before. We'll define them in a later section.
#
# **4. Tweak our prediction function until we're happy!**
# --once we've built out the functions that underpin our predictive algorithm, we'll tweak them until we hit our desired accuracy metric. In this case, **_we'll shoot for an accuracy of at least 80%._**
# <br>
# <br>
# <center>Let's get started!</center>
#Import everything needed for the project.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# <center><h2>Step 1: Load and Explore the Data</h2></center>
#
# In this section, we'll:
#
# 1. Read the data from `titanic.csv` and store it in a dataframe (you'll find this file in the `/datasets` folder).
# 2. Remove the `Survived` column from the dataframe and store it as a Pandas Series in a variable.
# 3. Create a general purpose function that visualizes survivors vs deaths in any data frame passed in.
# 4. Clean our dataframe (remove unnecessary columns, deal with null values, etc).
# 5. Explore our data and figure out which groups are most likely to survive.
#
#
# NOTE: There are many ways to successfully visualize survival rates across the different features. The most inuitive way to visualize survival rates as a stacked bar chart, where 'survived' and 'dead' are different colors on the same bar. For an easy explanation of how to make these bar charts, see [this Stack Overflow question](https://stackoverflow.com/questions/41622054/stacked-histogram-of-grouped-values-in-pandas).
# +
# Read in the titanic.csv dataset from the /datasets folder.
raw_df = None
# Store the survived column in the labels variable, and then drop the column from the data frame.
labels = None
#Don't forget to remove these columns from the dataframe!
columns_to_remove = ['PassengerId', 'Name', 'Ticket', 'Cabin']
# -
# Next, we'll create a function that allows us to quickly visualize the survival rates of any dataframe of passengers. This way, we can iterate quickly by slicing our dataframe and visualizing the survival rate to see if we can find any patterns that will be useful to us.
#
# As an example, if we wanted to visualize the survival rates of men versus women, we would create a dataframe object that contains only the information that matters to us, and then pass it into this function. When completed, this function should output a histogram plot that looks like the ones seen in the Stack Overflow link listed above.
# Create a function used to visualize survival rates for the data frame passed in
def visualize_survival_rates(dataframe, xlabel=None, ylabel="Count"):
"""
Inputs: dataframe--a pandas dataframe object consisting of the things you want visualized.
labels--a pandas series object that tells us whether each passenger died (0) or survived(1)
Outputs: A 2 color histogram that visualizes the survival rate of passengers based on the values contained
within the dataframe. For instance, if we pass in a visualization
NOTE: You should rely on the dataframe's .hist() method to do most of the heavy lifting for visualizations.
Any slicing of the dataframe should be done BEFORE you call this function. For instance, if you want to visualize
survival rates of men under 30 vs women under 30, you should create a dataframe containing only these rows and
columns before passing it into this function, rather than passing in the full original dataframe. This will
allow you to keep the logic in this function simple.
"""
pass
# <center><h3>Building a Prediction Function</h3></center>
#
# Next, we'll write a prediction function. We'll use basic control flow to examine each row in the data set and make a prediction based on whatever we think is important. If you explored the data set, you may have stumbled upon a few interesting discoveries, such as:
#
# * Women were more likely to survive than men.
# * Rich people were more likely to survive than poor people.
# * Young people were more likely to survive than others.
#
# (NOTE: We made these up--don't automatically assume they're true without investigating first!)
#
# These may seem obvious, but don't discount their usefulness! We can use these facts to build a prediction function that has decent accuracy! For instance, let's pretend that we found that 80% of all women survived. Knowing this, if we then tell our algorithm to predict than all female passengers survived, we'll be right 80% of the time for female passengers!
#
# Complete the following prediction function. It should take in a dataframe of titanic passengers. Based on the things you think are important (just use a bunch of nested control flow statements), you'll output a 1 if you think this passenger survived, or a if you think they died.
#
# The function should output an array where the first item is the prediction for the first row in the dataframe, the 2nd item in the array is the prediction for the seconf row in the dataframe, etc.
def predict_survival(dataframe):
predictions = []
# WRITE YOUR PREDICTION CODE BELOW!
return predictions
# <center><h3>Evaluating Your Predictions</h3></center>
#
# Great! Now we've evaluated our data and made a bunch of predictions--but predictions are only interesting if they're accurate. In order to do this, we're going to create a **_Confusion Matrix_** to track what we got right and wrong (and _how_ we were right and wrong).
#
# There are 4 different possible outcomes for each prediction:
#
# 1. **True Positive** -- You predicted they survived (1), and they actually survived (1).
# 2. **True Negative** -- You predicted they died (0), and they actually died (0).
# 3. **False Positive** -- You predicted they survived (1), and they actually died (0).
# 4. **False Negative** -- You predicted they died (0), and they actually survived (1).
#
# We're going to write a function that takes in our predictions and the actual labels (the "Survived" column we removed from the actual data frame), and determines which possible outcome we had for each prediction. We will keep track of how many times each outcome happened by incrementing a counter for each in our _Confusion Matrix_ dictionary.
#
# +
def create_confusion_matrix(predictions, labels):
confusion_matrix = {"TP": 0, "TN": 0, "FP": 0, "FN": 0}
# Recall each index in both 'predictions' and 'labels' are referring to the corresponding row.
# E.G. predictions[0] and label [0] both refer to row 0 in the dataframe that was passed into the
# prediction function.
#TODO: Create the confusion matrix by comparing the values in predictions to the corresponding values in labels.
# Use the definitions in the text above to determine which item in the dictionary you should increment.
return confusion_matrix
def get_accuracy(confusion_matrix):
# Create a function that returns the accuracy score for your classifier.
# The formula for accuracy = TP + TN / TP + TN + FP + FN
pass
# -
# <center><h3>Where to Go From Here</h3></center>
#
# Now that you have a way to evaluate your predictions, modify your prediction function until you can achieve an evaluation score above 80%!
| site/public/courses/DS-2.1/Assignments/00_Titanic_Survival_Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: py35-paddle1.2.0
# ---
# ## 准备工作
# ### 安装PaddleSeg以及更新依赖
# > 这里使用的PaddleSeg的2.3版本
# 解压PaddleSeg套件
# !unzip -oq /home/aistudio/data/data114849/PaddleSeg-release-2.3.zip
# 为方便后期操作将文件夹改名
# !mv PaddleSeg-release-2.3 PaddleSeg
# +
# 安装依赖,AIStudio实验环境已有相关依赖
# #!pip install -r PaddleSeg/requirements.txt
# -
# ### 解压数据集
# !unzip -oq /home/aistudio/data/data77571/train_and_label.zip -d data/
# !unzip -oq /home/aistudio/data/data77571/img_test.zip -d data/
# ### 生成训练列表文件
# > 这里直接按比例分割,没有用shuffle
# +
import os
img_train_dir = os.listdir('./data/img_train')
train_val_ratio = 0.9 #划分训练集和验证集的比例
train_img_num = int(len(img_train_dir)*train_val_ratio)
val_img_num = len(img_train_dir) - train_img_num
train_list_origin = img_train_dir[:train_img_num]
val_list_origin = img_train_dir[train_img_num:]
print(len(train_list_origin),len(val_list_origin)) #输出训练集和验证集数量
#生成trainlist
with open('train_list.txt','w') as f:
for i in range(len(train_list_origin)):
train_img_name = 'img_train/' + train_list_origin[i]
train_lab_name = 'lab_train/' + train_list_origin[i].split('.')[0] + '.png'
f.write(train_img_name + ' ' + train_lab_name + '\n')
#生成vallist
with open('val_list.txt','w') as f:
for i in range(len(val_list_origin)):
val_img_name = 'img_train/' + val_list_origin[i]
val_lab_name = 'lab_train/' + val_list_origin[i].split('.')[0] + '.png'
f.write(val_img_name + ' ' + val_lab_name + '\n')
# -
# ## 模型选择和实践
# ### Swin Transformer简介
# > Swin Transformer是ViT的一大进步,建议参考李沐大神读论文的方法啃原论文
# [Swin Transformer: Hierarchical Vision Transformer using Shifted Windows](https://arxiv.org/pdf/2103.14030.pdf)
#
# 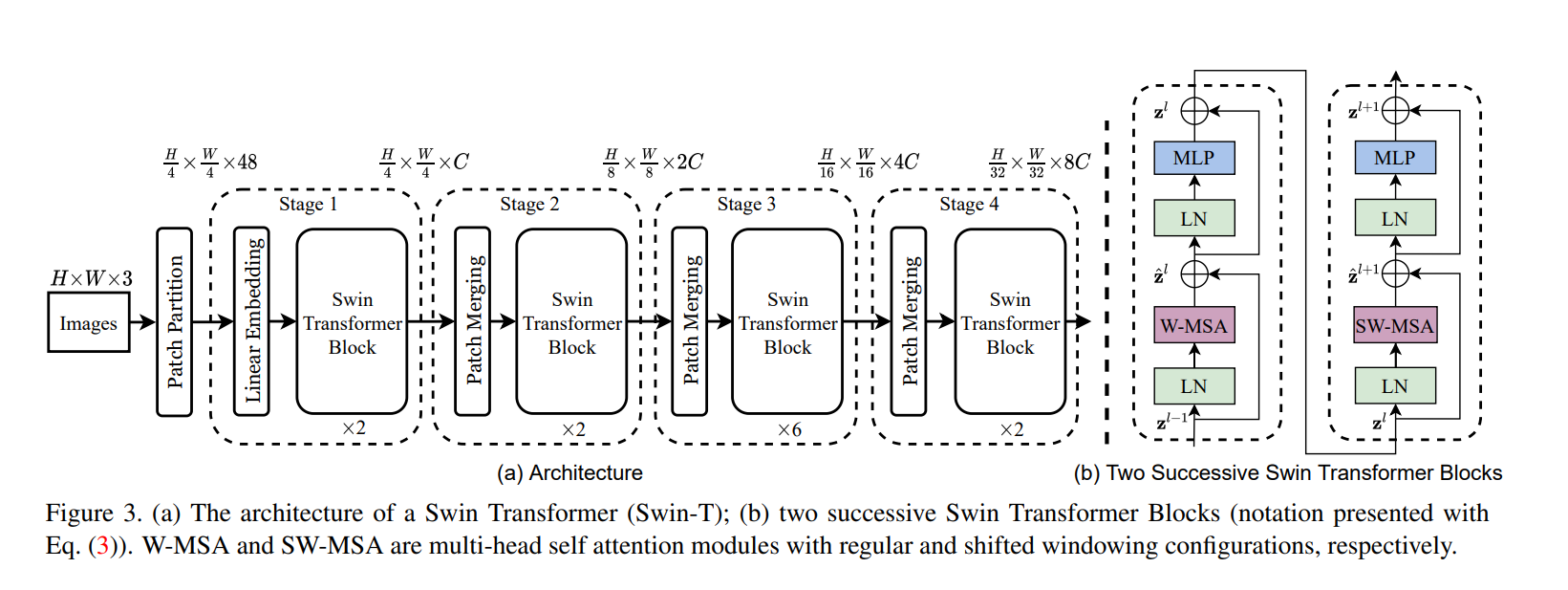
#
#
# ### 模型训练
# !python PaddleSeg/train.py \
# --config swin.yml \
# --do_eval \
# --use_vdl \
# --save_interval 4000 \
# --save_dir output
# ### 模型预测
# !python PaddleSeg/predict.py \
# --config swin0.yml \
# --model_path output/best_model/model.pdparams \
# --image_path data/img_testA \
# --save_dir ./result
# > 前往output\home\aistudio\result\pseudo_color_prediction打包预测结果
# ## 改进想法
# 1. 面对数据集不均衡现象采用欠采样或过采样方法重新处理数据集
# 1. 对欠采样的数据集使用不同的数据增强方法
# 1. Loss使用LovaszSoftmaxLoss
# ## 参考链接
# * [2020 CCF BDCI 地块分割Top1方案 & 语义分割trick整理](https://zhuanlan.zhihu.com/p/346862877)
#
# * [飞桨常规赛:遥感影像地块分割-7月第3名方案(增强修改版)](https://aistudio.baidu.com/aistudio/projectdetail/1789075)
#
# * [飞桨常规赛:遥感影像地块分割8月第1名方案](https://aistudio.baidu.com/aistudio/projectdetail/2284066)
| 2562409.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modeling
#
# This notebook applies a rebalancing algorithm to the target dataset. It then applies a series of classification models to the transformed dataset, prints the scores, and presents the results graphically.
# +
import psycopg2
import pandas as pd
import numpy as np
import os
import seaborn as sns
import matplotlib.pyplot as plt
sns.set(style="ticks", color_codes=True)
import matplotlib
import warnings
import time
import pickle
import yellowbrick as yb
from sklearn.preprocessing import OneHotEncoder, LabelEncoder
from sklearn.metrics import f1_score, precision_score, recall_score, accuracy_score, classification_report
from sklearn.pipeline import Pipeline
from sklearn.linear_model import LinearRegression, Lasso, Ridge
from sklearn.metrics import r2_score
from sklearn.svm import LinearSVC, NuSVC, SVC
from sklearn.neighbors import KNeighborsClassifier
from sklearn.preprocessing import OneHotEncoder, LabelEncoder
from sklearn.linear_model import LogisticRegressionCV, LogisticRegression, SGDClassifier
from sklearn.ensemble import BaggingClassifier, ExtraTreesClassifier, RandomForestClassifier
from sklearn.model_selection import train_test_split, GridSearchCV, KFold
from yellowbrick.classifier import ClassificationReport
warnings.simplefilter(action='ignore', category=FutureWarning)
# -
# ### Import modeling dataset
conn = psycopg2.connect(
host = 'project.cgxhdwn5zb5t.us-east-1.rds.amazonaws.com',
port = 5432,
user = 'postgres',
password = '<PASSWORD>',
database = 'postgres')
cursor = conn.cursor()
DEC2FLOAT = psycopg2.extensions.new_type(
psycopg2.extensions.DECIMAL.values,
'DEC2FLOAT',
lambda value, curs: float(value) if value is not None else None)
psycopg2.extensions.register_type(DEC2FLOAT)
# +
cursor.execute('Select * from "ahs_household_class"')
rows = cursor.fetchall()
col_names = []
for elt in cursor.description:
col_names.append(elt[0])
df = pd.DataFrame(data=rows, columns=col_names )
# -
df.head()
path = os.path.join(os.getcwd(), 'data', 'working')
df = pd.read_csv(os.path.join(path, 'AHS Household Class.csv'))
df.shape
# # Linear Models
df = pd.read_csv(r"C:\Users\Michael\Workspace\First-Home-Recommender\data\working\AHS Household Reg.csv")
df.shape
y = df['RATINGHS']
X = df.drop('RATINGHS', axis=1)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
# +
models = {
'Linear Regression':LinearRegression(),
'Lasso Regression':Lasso(),
'Ridge Regression':Ridge()
}
for title, model in models.items():
m = model
m.fit(X_train, y_train)
y_pred = m.predict(X_test)
#print(r2_score(y_test, y_pred))
print('{}: {}'.format(title, r2_score(y_test, y_pred)))
# -
# ### Class Rebalancing
X = df.drop(['Unnamed: 0','RATINGHS'], axis=1)
y = df['RATINGHS']
# +
from sklearn.preprocessing import LabelEncoder
encoder = LabelEncoder().fit(y)
y = encoder.transform(y)
encoder = LabelEncoder().fit(df['HINCP'])
X['HINCP'] = encoder.transform(df['HINCP'])
encoder = LabelEncoder().fit(df['FINCP'])
X['FINCP'] = encoder.transform(df['FINCP'])
# -
# Create the train and test data
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
pd.Series(y_train).value_counts().plot.bar()
from imblearn.over_sampling import SMOTE
sm = SMOTE(random_state = 33)
X_sm, y_sm = sm.fit_sample(X_train, y_train)
X_sm, y_sm = sm.fit_sample(X_sm, y_sm.ravel())
pd.Series(y_sm).value_counts().plot.bar()
# # Modeling
#
# ### ExtraTreesClassifier
X_train, X_test, y_train, y_test = train_test_split(X_sm, y_sm, test_size=0.2, random_state=42)
model = ExtraTreesClassifier()
model.fit(X_train, y_train)
expected = y_test
predicted = model.predict(X_test)
print("{}: {}".format(model.__class__.__name__, f1_score(expected, predicted, average='micro')))
def fit_and_evaluate(X, y, model, label, **kwargs):
"""
Because of the Scikit-Learn API, we can create a function to
do all of the fit and evaluate work on our behalf!
"""
start = time.time() # Start the clock!
scores = {'precision':[], 'recall':[], 'accuracy':[], 'f1':[]}
kf = KFold(n_splits = 12, shuffle=True)
for train, test in kf.split(X, y):
X_train, X_test = X.iloc[train], X.iloc[test]
y_train, y_test = y[train], y[test]
estimator = model(**kwargs)
estimator.fit(X_train, y_train)
expected = y_test
predicted = estimator.predict(X_test)
# Append our scores to the tracker
scores['precision'].append(precision_score(expected, predicted, average="weighted"))
scores['recall'].append(recall_score(expected, predicted, average="weighted"))
scores['accuracy'].append(accuracy_score(expected, predicted))
scores['f1'].append(f1_score(expected, predicted, average="weighted"))
# Report
print("Build and Validation of {} took {:0.3f} seconds".format(label, time.time()-start))
print("Validation scores are as follows:\n")
print(pd.DataFrame(scores).mean())
# Write official estimator to disk
estimator = model(**kwargs)
estimator.fit(X, y)
outpath = label.lower().replace(" ", "-") + ".pickle"
with open(outpath, 'wb') as f:
pickle.dump(estimator, f)
print("\nFitted model written to:\n{}".format(os.path.abspath(outpath)))
fit_and_evaluate(X, y, ExtraTreesClassifier, "ExtraTreeClassifier", n_estimators=100)
# # Gridsearch
#
# This section uses GridSearchCV to hyperparameter tune on ExtraTreeClassifier
model.get_params()
# +
param_grid = {'n_estimators': np.arange(1,255),
'max_features': ['auto','sqrt','log2'],
'max_depth': [100],
'n_jobs': [1]
}
grid = GridSearchCV(ExtraTreesClassifier(), param_grid, refit=True, verbose=3)
grid.fit(X_train, y_train)
# -
print('Best Score: {}'.format(grid.best_score_))
print('Best Estimator: {}'.format(grid.best_estimator_))
grid_predictions = grid.predict(X_test)
print(classification_report(y_test, grid_predictions))
# +
def visualize_model(X, y, estimator):
"""
Test various estimators.
"""
y = LabelEncoder().fit_transform(y)
model = Pipeline([
('one_hot_encoder', OneHotEncoder()),
('estimator', estimator)
])
# Instantiate the classification model and visualizer
visualizer = ClassificationReport(
model, classes=['extremely satisfied','satisfied','not satisfied ','very satisfied'],
cmap="Reds", size=(600, 360)
)
visualizer.fit(X, y)
visualizer.score(X, y)
visualizer.show()
for model in models:
visualize_model(X_sm, y_sm.ravel(), model)
# -
# ### Confusion Matrixs
#
# +
from yellowbrick.classifier import ConfusionMatrix
model = ExtraTreesClassifier(n_estimators=100)
cm = ConfusionMatrix(model, classes=['extremely satisfied','satisfied','not satisfied ','very satisfied'])
# Fit fits the passed model. This is unnecessary if you pass the visualizer a pre-fitted model
cm.fit(X_sm, y_sm.ravel())
cm.score(X_sm, y_sm.ravel())
cm.show()
# +
model = RandomForestClassifier(n_estimators=100)
cm = ConfusionMatrix(model, classes=['extremely satisfied','satisfied','not satisfied ','very satisfied'])
# Fit fits the passed model. This is unnecessary if you pass the visualizer a pre-fitted model
cm.fit(X_sm, y_sm.ravel())
cm.score(X_sm, y_sm.ravel())
cm.show()
# -
| Archive/Modeling_Mike.ipynb |