code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # mRNA Expression Analysis using Hybridization Chain Reaction
# This code was used to analyze HCR results for putative BMPR1A targets identified by RNA-Seq.
#
# Required inputs for this script:
#
# 1. .csv file containing source data for each image (embryo) documenting the area, mean, intden, and raw intden for background, control, and experimental regions of interest (ROI)
#
# Script prepared by <NAME>, March 2021
# +
# Import data handling and analysis packages
import os
import glob
import pandas as pd
from scipy import stats
# Import plotting packages
import iqplot
import bokeh.io
from bokeh.io import output_file, show
from bokeh.layouts import column, row
bokeh.io.output_notebook()
# -
# ## Import source data
# +
source_data = pd.read_csv('Fig4_source_data.csv')
source_data.replace(to_replace=['tfap2b', 'id2', 'hes6', 'apod'],value=['TFAP2B', 'ID2', 'HES6-2', 'APOD'], inplace=True)
source_data.head()
# +
# Define control and experimental constructs
cntl_construct = 'RFP'
expt_construct = 'dnBMPR1A-FLAG'
# Get a list of experimental treatments in this dataframe
treatment_list = source_data.Treatment.unique()
treatment_list = treatment_list.tolist()
treatment_list
# -
# ## Isolate and analyze mean fluorescence intensity for each image
#
# This will determine the ratio of fluorescence intensities between control and experimental sides (Experimental/Control)
# +
# Get a list of target genes measured
target_list = source_data.Target.unique().tolist()
# Initialize for final dataframe collection
full_results = pd.DataFrame()
full_results_list = []
# Loop through target genes:
for target in target_list:
df_target = source_data.loc[source_data['Target'] == target][['Target','EmbID','Treatment',
'Somites','ROI','Area','Mean','IntDen']]
# Initialize for temporary dataframe collection
target_results = pd.DataFrame()
target_results_list = []
# Loop through embryos:
embryo_list = df_target.EmbID.unique().tolist()
for embryo in embryo_list:
df_embryo = df_target.loc[df_target['EmbID'] == embryo]
# Assemble output df from specific values in each embryo dataset
data = {'Target': [target, target], 'EmbID': [embryo, embryo]
,'Treatment': [df_embryo.tail(1)['Treatment'].values[0], df_embryo.tail(1)['Treatment'].values[0]]
,'Somites': [df_embryo.tail(1)['Somites'].values[0], df_embryo.tail(1)['Somites'].values[0]]
,'ROI': ['Cntl', 'Expt']
,'Mean': [float(df_embryo.loc[df_embryo['ROI'] == 'Cntl']['Mean']),
float(df_embryo.loc[df_embryo['ROI'] == 'Expt']['Mean'])]
}
embryo_results = pd.DataFrame(data)
target_results_list.append(embryo_results)
# Normalize mean levels within this target dataset to the mean of the control group
target_results = pd.concat(target_results_list, sort=False).reset_index().drop('index', axis=1)
cntl_mean = target_results.loc[target_results['ROI'] == 'Cntl']['Mean'].mean()
target_results['normMean'] = target_results['Mean']/cntl_mean
full_results_list.append(target_results)
# Assemble and view the final results
full_results = pd.concat(full_results_list,sort=False).reset_index().drop('index', axis=1)
full_results.head()
# -
# ## Parallel coordinate plots for single targets
#
# Displays Control and Experimental values, connected by a line to link measurements from same embryo
#
# Also perform two-tailed paired t-test for these values
# +
################### Isolate data for analysis ###################
# Annotate data further to plot
cntl_construct = 'RFP'
expt_construct = 'dnBMPR1A-FLAG'
# Gene to parse:
gene = ['ID2']
# Pull out only cells and treaments of interest, and rename ROIs with the appropriate constructs
df = full_results.loc[full_results['Target'].isin(gene)].copy()
df.replace(to_replace = {'Cntl': cntl_construct, 'Expt': expt_construct}, inplace=True)
################### Plot as strip plot ###################
# Plot as strip plot
p1 = iqplot.strip(data=df
,q='normMean', q_axis='y'
,cats=['ROI'], parcoord_column='EmbID'
,y_range=(0,2)
# ,frame_height = 300, frame_width = 200
,frame_height = 400, frame_width = 400
,y_axis_label=str('Normalized '+str(gene[0])+' expression')
,x_axis_label='Treatment'
,color_column='Somites'
,marker_kwargs=dict(size=10
# ,color='black'
)
,parcoord_kwargs=dict(line_width=1,color='gray')
# ,show_legend=True
,tooltips=[("Embryo", "@EmbID"), ]
)
# p1.axis.axis_label_text_font_style = 'bold italic'
p1.axis.axis_label_text_font_size = '14px'
p1.axis.major_label_text_font_size = '14px'
# p1.legend.location = "top_right"
show(row(p1))
################### Perform statistical analysis ###################
# Perform Paired t test
cntl = df.loc[df['ROI'] == cntl_construct]['Mean']
expt = df.loc[df['ROI'] == expt_construct]['Mean']
ttest = stats.ttest_rel(cntl,expt)
# Display test results
print('Paired t-test results: \n\t\t statistic = ' + str(ttest[0]) +
'\n\t\t p-value = ' + str(ttest[1]))
print('n = ' + str(len(cntl)) + ' embryos')
# -
# ## Assemble ratio dataframe (Experimental / Control measurements), then plot as a stripbox plot
# +
ratios_raw = full_results.copy()
ratios_raw['ExperimentID'] = ratios_raw['EmbID']+'_'+ratios_raw['Target']
expt_list = ratios_raw['ExperimentID'].unique().tolist()
ratio_results = pd.DataFrame()
list_ = []
for expt in expt_list:
expt_df = ratios_raw.loc[ratios_raw['ExperimentID'] == expt]
ratio_mean = (float(expt_df.loc[expt_df['ROI'] == 'Expt']['Mean'])
/float(expt_df.loc[expt_df['ROI'] == 'Cntl']['Mean']))
# Assemble output df
data = {'ExperimentID': [expt],
'ratioMean': [ratio_mean],
}
expt_results = pd.DataFrame(data)
list_.append(expt_results)
ratio_results = pd.concat(list_,sort=False).reset_index().drop('index', axis=1)
(ratio_results['Date'], ratio_results['Stains'], ratio_results['Embryo'], ratio_results['Target']
) = zip(*ratio_results['ExperimentID'].map(lambda x: x.split('_')))
ratio_results.head()
# +
# Choose targets to plot
targets = ['TFAP2B', 'ID2', 'HES6-2', 'APOD',]
data = ratio_results[ratio_results['Target'].isin(targets)]
# Build Stripbox plot
stripbox = iqplot.stripbox(
# Data to plot
data=data,
q='ratioMean', q_axis='y',
cats='Target',
# Plot details
jitter=True, jitter_kwargs=dict(width=0.3),
marker_kwargs=dict(alpha=0.8, size=8
# ,color='darkgray'
),
box_kwargs=dict(line_color='black', line_width=1.5),
whisker_kwargs=dict(line_color='black', line_width=1.5),
median_kwargs=dict(line_color='black', line_width=2),
top_level='box',
frame_width=350, frame_height=350,
# Plot customizations
order=targets,
y_range=(0,2.05),
y_axis_label='Relative HCR Intensity',
x_axis_label='Gene',
show_legend=False,
)
# Final customizations
stripbox.axis.axis_label_text_font_size = '16px'
stripbox.axis.major_label_text_font_size = '16px'
stripbox.axis.axis_label_text_font_style = 'bold'
stripbox.xaxis.major_label_text_font_style = 'italic'
# View plot
show(stripbox)
# -
| Fig4_HCR_Validation/2021_WholeMount_HCR_Intensity_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
cm_data = np.genfromtxt('scores/output.txt', dtype=str)
cm_data2 = np.genfromtxt('scores/output.txt', dtype=str)
fuse_df = fuse(f_list)
fuse_df.to_csv('merge_result.csv', sep=' ', header=False, index=False)
| tDCF_python_v1/FusionFunction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="qXyD34zG_QYr"
# # Artificial Neural Networks in Python using Tensorflow
#
# This notebook is a simple demonstration to how to build an Artificial Neuron Network using TensorFlow to a classification problem.
#
# **Problem Description**
#
# The goal of program is to build a classification model to predict if a certain client will leave the bank service in the next six months.
#
# **Dataset Description**
#
# The dataset is composed by 10000 instances (rows) and 14 features (columns). The features considered to build the model are:
#
# - RowNumber (This is not important to the model)
# - CustomerId (This is not important to the model)
# - Surname (This is not important to the model)
# - CreditScore (numerical variable)
# - Geography (categorical variable)
# - Gender (categorical variable)
# - Age (numerical variable)
# - Tenure (categorical variable)
# - Balance (numerical variable)
# - NumOfProducts (categorical variable)
# - HasCrCard (categorical variable)
# - EstimatedSalary (numerical variable)
# - Exited (target)
# -
# # Data preprocessing
# + [markdown] id="7ANWPtw_AFZu"
# ## Importing Libraries
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="ushgeC5iAKCu" outputId="56b2f19f-96fa-4456-a0c0-5e8650e3fa17"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import tensorflow as tf # We utilize it to build our ANN model.
tf.__version__
# + [markdown] id="SLTAs4t3AUg2"
# ## Importing Dataset
#
# + colab={"base_uri": "https://localhost:8080/"} id="OaApzIM5AXtU" outputId="aa17c74f-49f7-4e4b-c1f4-a04460d1e6af"
dataset = pd.read_csv('Churn_Modelling.csv')
# -
# ## Visualizing some informations from the dataset
dataset.shape
dataset.dtypes
dataset.isna().sum()
dataset.head(10)
dataset.describe()
# # Exploratory Data Analysis (EDA)
# ## Visualizing the categorical variables
# ### Target variable
dataset['Exited'].value_counts()
plt.figure(figsize=(15, 5))
plt.subplot(1, 2, 1)
sns.countplot(x=dataset['Exited'])
plt.subplot(1, 2, 2)
values = dataset.iloc[:, - 1].value_counts(normalize = True).values # to show the binirie values in parcentage
index = dataset.iloc[:, -1].value_counts(normalize = True).index
plt.pie(values, labels= index, autopct='%1.1f%%', colors=['b', 'tab:orange'])
plt.show()
# ### Others categorical variables
categorical_list = ['Geography', 'Gender', 'Tenure', 'NumOfProducts', 'HasCrCard','IsActiveMember', 'Exited']
data_cat = dataset[categorical_list]
fig = plt.figure(figsize=(15, 15))
plt.suptitle('Pie Chart Distribution', fontsize = 20)
for i in range(1, data_cat.shape[1]):
plt.subplot(2, 3, i)
f = plt.gca()
f.axes.get_yaxis().set_visible(False)
f.set_title(data_cat.columns.values[i - 1])
# Setting the biniries values
values = data_cat.iloc[:, i - 1].value_counts(normalize = True).values # to show the binirie values in parcentage
index = data_cat.iloc[:, i -1].value_counts(normalize = True).index
plt.pie(values, labels= index, autopct='%1.1f%%')
plt.axis('equal')
#fig.tight_layout(rect=[0, 0.03, 1, 0.95])
plt.show()
plt.figure(figsize=(15, 10))
for i in range(1, data_cat.shape[1]):
plt.subplot(3, 3, i)
sns.countplot(x=data_cat.iloc[: , i-1], hue=data_cat['Exited'])
plt.show()
# ## Visualizing the numerical variables
numerical_list = ['CreditScore', 'Age', 'Balance', 'EstimatedSalary', 'Exited']
data_num = dataset[numerical_list]
# ### Distribution of numerical variables
plt.figure(figsize=(25,15))
plt.suptitle('Histograms of numerical variables (mean values)', fontsize = 20)
for i in range(1, data_num.shape[1]):
plt.subplot(2, 2, i)
f = plt.gca()
sns.histplot(data_num.iloc[:, i-1], color = '#3F5D7D', kde= True)
plt.show()
plt.figure(figsize=(25,15))
plt.suptitle('Histograms of numerical variables (mean values)', fontsize = 20)
for i in range(1, data_num.shape[1]):
plt.subplot(2, 2, i)
f = plt.gca()
sns.histplot(data=data_num, x=data_num.iloc[:, i-1], hue='Exited', kde = True)
plt.show()
# ## Correlation and PairPlot (scatter)
# ### Correlation with the response variable
column_drop = ['RowNumber', 'CustomerId', 'Surname', 'Exited']
dataset.drop(columns=column_drop).corrwith(dataset.Exited).plot.bar(
figsize = (20, 10), title = "Correlation with Exited", fontsize = 15,
rot = 45, grid = True)
# ### Correlation Between the Variables
# +
column_drop = ['RowNumber', 'CustomerId', 'Surname']
## Correlation Matrix
sns.set(style="white")
# Compute the correlation matrix
corr = dataset.drop(columns=column_drop).corr()
# Generate a mask for the upper triangle
mask = np.zeros_like(corr, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(10, 20))
# Generate a custom diverging colormap
cmap = sns.diverging_palette(220, 10, as_cmap=True)
# Draw the heatmap with the mask and correct aspect ratio
sns.heatmap(corr, mask=mask, cmap=cmap, vmax=1, vmin=-1, center=0,
square=True, linewidths=.5, cbar_kws={"shrink": .5}, annot = True)
# -
# ### Pair plot for the numerical variables
sns.pairplot(data_num, hue = 'Exited', kind = 'scatter', corner=True, diag_kind='None')
# ## EDA conclusion
#
# **Target variable**
#
# Represented by 0 or 1 (stay/leave) shows a frequency of $79.6 \%$ for the customers which decided to stay in the bank against $20.4\%$ of customers that decided to leave the bank.
#
# **Categorical variables**
#
# The analysis of these variables shows that the most clients are French. The majority of clients are males. The frequency of these variables are approximately equals without a fact that requires attention and manipulation, it means feature engineering.
#
# **Numerical variables**
#
# The distribution of these variables are normal except for the salary. One interesting aspect is the age distribution for the clients which leaved the bank, the distribution shows that the almost clients have age among 40 and 50 years. This fact can be well visualized in the scatter plots.
#
# **Correlation**
#
# The correlation between the target and the independent variables are not so big, but considerable to build the model. The correlation among the independent variables are satisfactory, the almost of correlation show a low values.
# # Building the Artificial Neuron Network
# ## Data preprocessing
# ### Excluding not important columns
dataset = dataset.drop(columns=['RowNumber', 'CustomerId', 'Surname'])
# + [markdown] id="vAfvzqlxDigi"
# ### Encoding Categorical Data
# + colab={"base_uri": "https://localhost:8080/"} id="lNMBBSsVDlXx" outputId="eda5f00c-7fb2-4219-fc97-b1e11d759d08"
dataset['Gender'] = dataset['Gender'].astype('category').cat.codes
# + [markdown] id="KHsmjrNVExcN"
# #### One Hot Econding
# + colab={"base_uri": "https://localhost:8080/"} id="VPw3iUhJE2qS" outputId="ddcb325b-c92b-4154-9c13-a10ca1dfc320"
dataset = pd.get_dummies(dataset)
# -
dataset.head()
# ### Defining the independent and target variable
response = dataset['Exited']
i_var = dataset.drop(columns=['Exited'])
# + [markdown] id="WVPScdSdGeqx"
# ### Splitting the Dataset into train and test set
# + id="PKH9UuJRGkvg"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(i_var, response, test_size = 0.2, random_state = 0)
# + [markdown] id="rvmtD8VQHEQ3"
# ### Feature Scaling
# In almost ANN models we must to apply feature scale.
# + id="JO1fypeSHJbV"
from sklearn.preprocessing import StandardScaler
sc_X = StandardScaler()
X_train_bckp = pd.DataFrame(sc_X.fit_transform(X_train))
X_test_bckp = pd.DataFrame(sc_X.transform(X_test))
X_train_bckp.columns = X_train.columns.values
X_test_bckp.columns = X_test.columns.values
X_train_bckp.index = X_train.index.values
X_test_bckp.index = X_test.index.values
X_train = X_train_bckp
X_test = X_test_bckp
# + [markdown] id="RoNknmZsJoBy"
# ## The Artificial Neuron Network - ANN
#
# An Artificial Neuron Network is a technique that tries to reproduce brain functions. To build an ANN, we consider a set of input information about something, each of these information will be considered as a neuron, we call this set of neuron input layer. As we know, in the brain we have many neurons and the communication among them is made by synapses process. The neurons from the input layer will communicate with other set of neurons in a hidden layer by the synapses process. Once the communication between the neurons of input and hidden layer is made, the initial information is changed, in this moment, the neurons of the hidden layer will communicate with a others neurons, these neurons is considered as a output layer and it gives a response about something. For example, in a classification model, if we consider five independent variables, the input layer will be composed by five neurons, these neurons communicate with the hidden layer (the number of neurons must be chosen), the hidden layer communicate with the output layer, that provides the final response 0 or 1, if we have two class.
#
# **How do synapses work in an ANN?**
#
# The synapses process in an ANN is made by an activation function, this function transforms the input information according with an associated weight (this might be interpreted as an importance degree to each input variable). We have some kind of activation function as
# - Threshold function
# - Sigmoid function
# - Rectifier function
# - Hyperbolic tangent function.
# For this notebook we are interested in the rectifier function and sigmoid function.
#
# The rectifier function is defined as $\phi(x) = \max(x,0)$, if a certain values is less than 0 the function returns 0, otherwise the function returns the maximum value. The synapses or the communication is made according $\sum_{i=1}^{m}w_{i}x_{i}$, where $m$ is the number of input variable. We consider the rectifier function between the input layer and the hidden layers. The activation function that we consider to make the communication among the hidden and output layer is the sigmoid function. The sigmoid function returns the probability of occurrence to certain class, this function is defined as $\phi(x) = \frac{1}{1 + e^{-x}}$.
#
# **How do an ANN learn?**
#
# The learning process of an ANN starts with the input parameters in a input layer, this layer communicate with the hidden layer by synapses. The next step is the communication among the hidden layer with the output layer, to obtain a response. In this stage, the learning process is not ended. After the response, the ANN must calculate the loss function to measure the precision of the prediction. Once the loss function was calculated there is the retro-propagation process, this process tries to find the minimal of the loss function changing the associated weights to each neuron. After this step, the ANN remake the previous process of synapses among the layers. The number of retro-propagation is defined as epochs.
#
# **About loss function**
#
# One of the most used loss function is the cross entropy. To more details [see](https://towardsdatascience.com/cross-entropy-loss-function-f38c4ec8643e).
#
# **About the retro-propagation**
#
# Here, we consider the stochastic gradient descent. This method tries to minimize the loss function changing the weights of each neuron. In this process, if we have 128 instances, we choose a batch number as 32, it means that we calculate the response for the associated batch number and we returns to the begin to remake the same process to other 32 instances. To more details [see](https://en.wikipedia.org/wiki/Stochastic_gradient_descent#Adam).
#
# **Summarizing the ANN process**
#
# 1. Initializing the ANN
# 2. Build the input layer
# 3. Build the hidden layers
# 4. Build the output layer
# 5. Training the ANN
#
# 1. Compile
# 2. Train.
# + [markdown] id="Vk4TpIBUJsRp"
# ## Initializing the ANN
#
# The first step is to create an object to build a sequence layer. This sequence layer takes account the input layer (the parameters that we initialize the ANN), the hidden layers and output layer. To do it, we utilize the modulus Tensor Flow (version 2.0 or high) that allow us to call the Keras modulus.
# + id="qLSqUQRXJ0uS"
ann = tf.keras.models.Sequential()
# + [markdown] id="oagB1r01eJhp"
# ## Input layer
#
# In this step, we create the input layer, it means, we set all independent variables (considering these neurons as input neurons) . When the input values go to the first hidden layer we must to choose an activation function. For this case, we choose rectifier function (the weights to each input parameters are chosen by the ann object). Units is the number of input neurons, relu is the activation function.
# + id="9BxdU1ZsfukC"
ann.add(tf.keras.layers.Dense(units=X_train.shape[1], activation='relu'))
# + [markdown] id="t6AZbMdTinxv"
# ## Hidden layers
#
# Here, we set the hidden layers. We can put how much we want. For this problem, we consider just one hidden layer. The object to create the hidden layer is the same to the input layer, but we can change it according with the problem. The parameters are the same of the input layer.
# + id="343W94K3jFbE"
tf.keras.layers.Dropout(0.2) # To drop 20% of input neurons, to avoid overfit.
ann.add(tf.keras.layers.Dense(units=X_train.shape[1], activation='relu'))
# + [markdown] id="Fy_EkroCjJj0"
# ## Output layer
#
# The last layer is the output layer. We use the same object the we used to build the preceding layers, but here, we make some changes in the parameters. Like this problem has a binary response yes or no, the number of neuron corresponds to 1, but if the response gives more then two results (0, 1, 2, for example), we must to consider the correspondent number of responses. The second change is on the activation function. Here, we consider sigmoid activation function, for one simple reason, this gives to us the probability which will be interpreted as 0 or 1 according with the values. Likelihood less than $0.5$ is considered as 0, otherwise 1.
# + id="9RJRX5ODjQS7"
tf.keras.layers.Dropout(0.2)
ann.add(tf.keras.layers.Dense(units=1, activation='sigmoid'))
# + [markdown] id="g-v5KN4ClfZD"
# ## Learning process of the ANN
# + [markdown] id="sarW2BIVlkM7"
# ### Compiling the ANN
#
# Compile the ANN is one of most important step. We select a method to optimize our ANN, stochastic gradient descent, represented by adam. The lost function is also very import, because from this function we are able to improve the accuracy, precision and other relevant parameters. The lost function that we are going to use is Binary Cross Entropy (due to have a binary response). Finally, the metric which we choose is the accuracy. Beyond this metric, we have others important metrics as F1, precision and recall.
# + id="V_kuMk2PlqWK"
ann.compile(optimizer='adam', loss='binary_crossentropy', metrics=['accuracy',tf.keras.metrics.Recall(), tf.keras.metrics.Precision()])
# + [markdown] id="HceFPt3-my5m"
# ### Training the ANN
#
# Now we train the ANN. Here we have two important hype parameters. Batch size determines how many instances the stochastic gradient descent will consider to work in the minimizing process. Epochs indicates the number of retro-propagation we want to train the model.
# + colab={"base_uri": "https://localhost:8080/"} id="zn0oSK2Zm5Ig" outputId="8a4227c9-84d8-497e-d602-6f9e088cd2c1"
ann.fit(X_train, y_train, batch_size= 32, epochs= 100)
# + [markdown] id="PqFny2LvrAbP"
# ## Making a single prediction
#
# Here, we make a simple prediction. Remember, Geography and Gender was changed. We must take it account.
# + colab={"base_uri": "https://localhost:8080/"} id="GntqBk3nrTbh" outputId="13fec3a3-eb19-40f1-95ee-2502fdebb3b0"
print(ann.predict(sc_X.transform([[1, 0, 0, 600, 1, 1, 40, 60000, 2, 1, 1, 50000]])) > 0.5)
# + [markdown] id="izaEvAa9uC0z"
# ## Predicting the test results
# + colab={"base_uri": "https://localhost:8080/"} id="543-PUCbuFsT" outputId="20862437-cd09-4512-9b14-8f62cffd3932"
y_pred = ann.predict(X_test)
y_pred = (y_pred > 0.5) # Here we must put it, because we have as outcome the probability, but we want a binary response.
# + [markdown] id="AVftybBMuc2J"
# ## Making the confusion matrix and metrics scores
# + colab={"base_uri": "https://localhost:8080/"} id="6vppMTWRuiEv" outputId="62ab924f-8104-4580-9044-86f1f507d8c7"
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
plt.figure()
sns.heatmap(cm, annot=True)
plt.show()
# +
# Here, we consider the metrics class from TensorFlow library.
m1 = tf.keras.metrics.Accuracy() # Object
m2 = tf.keras.metrics.Recall()
m3 = tf.keras.metrics.Precision()
m1.update_state(y_test, y_pred) # Calculating the metric
m2.update_state(y_test, y_pred)
m3.update_state(y_test, y_pred)
print('Metric results for the test set\n')
print('Accuracy {:.2f}%'.format(m1.result().numpy()*100))
print('Recall {:.2f}%'.format(m2.result().numpy()*100))
print('Precision {:.2f}%'.format(m3.result().numpy()*100))
# +
score_train = ann.evaluate(X_train, y_train)
print('\n')
print('Metric results for the training set\n')
print('Accuracy {:.2f}%'.format(score_train[1]*100))
print('Recall {:.2f}%'.format(score_train[2]*100))
print('Precision {:.2f}%'.format(score_train[3]*100))
# -
# ## Metrics with cross validate
def build_classifier(optimizer='adam'):
classifier = tf.keras.models.Sequential()
classifier.add(tf.keras.layers.Dense(units=X_train.shape[1], activation='relu'))
classifier.add(tf.keras.layers.Dense(units=X_train.shape[1], activation='relu'))
classifier.add(tf.keras.layers.Dense(units=1, activation='sigmoid'))
classifier.compile(optimizer = optimizer, loss = 'binary_crossentropy', metrics = ['accuracy',tf.keras.metrics.Recall(), tf.keras.metrics.Precision()])
return classifier
# +
scoring = ['accuracy', 'recall', 'precision'] # List of metrics
from sklearn.model_selection import cross_validate
classifier = tf.keras.wrappers.scikit_learn.KerasClassifier(build_fn = build_classifier, batch_size = 32, epochs = 100)
accuracies = cross_validate(estimator = classifier, X = X_train, y = y_train,
cv = 10, n_jobs = -1, scoring = scoring)
# -
print('Metric results with cross validate cv=10')
print('\n')
print("Accuracy: {:.2f} %".format(accuracies['test_accuracy'].mean()*100))
print("Recall: {:.2f} %".format(accuracies['test_recall'].mean()*100))
print("Precision: {:.2f} %".format(accuracies['test_precision'].mean()*100))
# ## Boosting the model with GridSearchCV
# +
scoring = {'ACC' : 'accuracy', 'REC' : 'recall', 'PC' : 'precision'}
from sklearn.model_selection import GridSearchCV
classifier = tf.keras.wrappers.scikit_learn.KerasClassifier(build_fn = build_classifier)
parameters = {'batch_size': [8, 16, 32],
'epochs': [50, 100, 500],
'optimizer': ['adam', 'rmsprop']}
grid_search = GridSearchCV(estimator = classifier,
param_grid = parameters,
scoring = scoring,
refit = 'ACC',
cv = 10, n_jobs = -1)
# -
grid_search = grid_search.fit(X_train, y_train)
grid_search.best_params_
# ### Predicting new test results with the best parameters
boosted_predictions = grid_search.predict(X_test)
boosted_predictions = (boosted_predictions > 0.5)
print('Metrics results - Boosted')
print('\n')
print('Best Parameters')
print(grid_search.best_params_)
print('Confusion Matrix')
cmb2 = confusion_matrix(y_test, boosted_predictions)
plt.plot()
sns.heatmap(cmb2, annot=True)
plt.show()
print('Metrics Results')
print("- Accuracy: {:.2f} %".format(grid_search.cv_results_['mean_test_ACC'][grid_search.best_index_].mean()*100))
print("- Recall: {:.2f} %".format(grid_search.cv_results_['mean_test_REC'][grid_search.best_index_].mean()*100))
print("- Precision: {:.2f} %".format(grid_search.cv_results_['mean_test_PC'][grid_search.best_index_].mean()*100))
print('\n')
# + [markdown] id="1rPoWgp3BO3R"
# # Conclusion
#
# In this program, we shown a simple example how to build an ANN. The objective of build a good regression model was achieved, the model presents an accuracy of $85.97 \%$. Artificial Neuron Network has a has a wide applicability, we might also to build regression models. Whit the classification model, we are free to apply in many problems, for example image recognition. Methods like this might be helpful tool in decision-make about client polices.
| 1 - Artifical Neural Networks/Artifical_Neural_Networks_TensorFlow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8 (full)
# language: python
# name: python3-3.8-ufrc
# ---
# # Pandas
# ## Use the Pandas library to do statistics on tabular data.
# * Pandas is a widely-used Python library for statistics, particularly on tabular data.
# * Borrows many features from R's dataframes.
# * A 2-dimenstional table whose columns have names and potentially have different data types.
# * Load it with `import pandas`.
# * Read a Comma Separate Values (CSV) data file with `pandas.read_csv`.
# * Argument is the name of the file to be read.
# * Assign result to a variable to store the data that was read.
# +
import pandas
data = pandas.read_csv('data/gapminder_gdp_oceania.csv')
print(data)
# -
# * The columns in a dataframe are the observed variables, and the rows are the observations.
# * Pandas uses backslash `\` to show wrapped lines when output is too wide to fit the screen.
# #### File Not Found
# Our lessons store their `data` files in a data sub-directory, which is why the path to the file is `data/gapminder_gdp_oceania.csv`. If you forget to include `data/`, or if you include it but your copy of the file is somewhere else, you will get a runtime error that ends with a line like this:
# `OSError: File b'gapminder_gdp_oceania.csv' does not exist`
# ### Use `index_col` to specify that a column's values should be used as row headings.
# * Row headings are numbers (0 and 1 in this case).
# * Really want to index by country.
# * Pass the name of the column to `read_csv` as its `index_col` parameter to do this.
data = pandas.read_csv('data/gapminder_gdp_oceania.csv', index_col='country')
print(data)
# ## Use `DataFrame.info` to find out more about a dataframe.
data.info()
# * This is a `DataFrame`
# * Two rows named `'Australia'` and `'New Zealand'`
# * Twelve columns, each of which has two actual 64-bit floating point values.
# * We will talk later about null values, which are used to represent missing observations.
# * Uses 208 bytes of memory.
# ## The `DataFrame.columns` variable stores information about the dataframe's columns.
# * Note that this is data, *not* a method.
# * Like `math.pi`.
# * So do not use `()` to try to call it.
# * Called a *member variable*, or just *member*.
print(data.columns)
# ## Use `DataFrame.T` to transpose a dataframe.
# * Sometimes want to treat columns as rows and vice versa.
# * Transpose (written `.T`) doesn't copy the data, just changes the program's view of it.
# * Like `columns`, it is a member variable.
print(data.T)
# ## Use `DataFrame.describe` to get summary statistics about data.
# * `DataFrame.describe()` gets the summary statistics of only the columns that have numerical data. All other columns are ignored, unless you use the argument `include='all'`.
print(data.describe())
# * Not particularly useful with just two records, but very helpful when there are thousands.
# ## Questions
# #### Q1: Reading Other Data
# Read the data in `gapminder_gdp_americas.csv` (which should be the same directory as `gapminder_gdp_oceania.csv`) into the variable called `americas` and display its summary statistics.
# **Solution**
#
# Click on the '...' below to show the solution.
# + jupyter={"source_hidden": true}
# To read in a CSV, we use `pandas.read_csv`and pass the filename
# 'data/gapminder_gdp_americas.csv' to it. We also once again pass the column
# name 'country' to the parameter `index_col` in order to index by country:
americas = pandas.read_csv('data/gapminder_gdp_americas.csv', index_col='country')
# -
# #### Q2: Inspecting Data
# After reading the data for the AMericans, use `help(americas.head)` and `help(americas.tal)` to find out what `DataFrame.head` and `DataFrame.tail` do.
# 1. What method call will display the first three rows fo this data?
# 2. What method call will display the last three columns of this data? (Hint: you may need to change your view of the data)
# **Solution**
#
# Click on the '...' below to show the solution.
# + jupyter={"source_hidden": true}
# 1. We can check out the first five rows of `americas` by executing
# `americas.head()` (allowing us to view the head of the DataFrame). We can
# specify the number of rows we wish to see by specifying the parameter `n`
# in our call to `americas.head()`. To view the first three rows, execute:
americas.head(n=3)
# The output is then
continent gdpPercap_1952 gdpPercap_1957 gdpPercap_1962 \
country
Argentina Americas 5911.315053 6856.856212 7133.166023
Bolivia Americas 2677.326347 2127.686326 2180.972546
Brazil Americas 2108.944355 2487.365989 3336.585802`
gdpPercap_1967 gdpPercap_1972 gdpPercap_1977 gdpPercap_1982 \
country
Argentina 8052.953021 9443.038526 10079.026740 8997.897412
Bolivia 2586.886053 2980.331339 3548.097832 3156.510452
Brazil 3429.864357 4985.711467 6660.118654 7030.835878`
gdpPercap_1987 gdpPercap_1992 gdpPercap_1997 gdpPercap_2002 \
country
Argentina 9139.671389 9308.418710 10967.281950 8797.640716
Bolivia 2753.691490 2961.699694 3326.143191 3413.262690
Brazil 7807.095818 6950.283021 7957.980824 8131.212843`
gdpPercap_2007
country
Argentina 12779.379640
Bolivia 3822.137084
Brazil 9065.800825
# 2. To check out the last three rows of `americas`, we would use the command,
# `americas.tail(n=3)`, analogous to `head()` used above. However, here we want
# to look at the last three columns so we need to change our view and then use
# `tail()`. To do so, we create a new DataFrame in which rows and columns are
# switched
americas_flipped = americas.T
# We can then view the last three columns of `americas` by viewing the last
# three rows of `americas_flipped`:
americas_flipped.tail(n = 3)
# The output is then:
country Argentina Bolivia Brazil Canada Chile Colombia \
gdpPercap_1997 10967.3 3326.14 7957.98 28954.9 10118.1 6117.36
gdpPercap_2002 8797.64 3413.26 8131.21 33329 10778.8 5755.26
gdpPercap_2007 12779.4 3822.14 9065.8 36319.2 13171.6 7006.58
country Costa Rica Cuba Dominican Republic Ecuador ... \
gdpPercap_1997 6677.05 5431.99 3614.1 7429.46 ...
gdpPercap_2002 7723.45 6340.65 4563.81 5773.04 ...
gdpPercap_2007 9645.06 8948.1 6025.37 6873.26 ...
country Mexico Nicaragua Panama Paraguay Peru Puerto Rico \
gdpPercap_1997 9767.3 2253.02 7113.69 4247.4 5838.35 16999.4
gdpPercap_2002 10742.4 2474.55 7356.03 3783.67 5909.02 18855.6
gdpPercap_2007 11977.6 2749.32 9809.19 4172.84 7408.91 19328.7
country Trinidad and Tobago United States Uruguay Venezuela
gdpPercap_1997 8792.57 35767.4 9230.24 10165.5
gdpPercap_2002 11460.6 39097.1 7727 8605.05
gdpPercap_2007 18008.5 42951.7 10611.5 11415.8
Note: we could have done the above in a single line of code by 'chaining' the commands:
americas.T.tail(n=3)
# -
# #### Q3: Reading Files in Other Directories
# The data for your current project is stored in a file called `microbes.csv`, which is located in a folder called `field_data`. Your are doing analysis in a notebook called `analysis.ipynb` in a sibliong folder called `thesis`:
your home directory
+-- field data/
| +-- microbes.csv
+-- thesis/
+-- analysis.ipynb
# What value(s) should you pass to `read.csv` to read `microbes.csv` in `analysis.ipynb`?
# **Solution**
#
# Click on the '...' below to show the solution.
# + jupyter={"source_hidden": true}
# We need to specify the path to the file of interest in the call to
# `pandas.read_csv`. We first need to `jump` out of the folder `thesis` using
# `../` and then into the folder `field_data` using `field_data/`. Then we
# can specify the filename `microbes.csv`.
#
# The result is as follows:
data_microbes = pandas.read_csv('../field_data/microbes.csv')
# -
# #### Q4: Writing Data
# As well as the `read_csv` function for reafing data from a file, Pandas provides a `to_csv` function to write dataframes to files. Applying what you've learned about reading from files, write one of your dataframes to a file calles `processed.csv`. You can use `help` to get information on how to use `to_csv`.
# **Solution**
#
# Click on the '...' below to show the solution.
# + jupyter={"source_hidden": true}
# In order to write the DataFrame `americas` to a file called `processed.csv`,
# execute the following command:
americas.to_csv('processed.csv')
# For help on `to_csv`, you could execute, for example,
help(americas.to_csv)
# Note that `help(to_csv)` throws an error! This is a subtlety and is due to
# the fact that `to_csv` is NOT a function in and of itself and the actual
# call is `americas.to_csv`.
| _episodes_rapids/061_reading_tabular_panda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbgrader={}
# # Interact Exercise 3
# + [markdown] nbgrader={}
# ## Imports
# + nbgrader={}
# %matplotlib inline
from matplotlib import pyplot as plt
import numpy as np
# + nbgrader={}
from IPython.html.widgets import interact, interactive, fixed
from IPython.display import display
# + [markdown] nbgrader={}
# # Using interact for animation with data
# + [markdown] nbgrader={}
# A [*soliton*](http://en.wikipedia.org/wiki/Soliton) is a constant velocity wave that maintains its shape as it propagates. They arise from non-linear wave equations, such has the [Korteweg–de Vries](http://en.wikipedia.org/wiki/Korteweg%E2%80%93de_Vries_equation) equation, which has the following analytical solution:
#
# $$
# \phi(x,t) = \frac{1}{2} c \mathrm{sech}^2 \left[ \frac{\sqrt{c}}{2} \left(x - ct - a \right) \right]
# $$
#
# The constant `c` is the velocity and the constant `a` is the initial location of the soliton.
#
# Define `soliton(x, t, c, a)` function that computes the value of the soliton wave for the given arguments. Your function should work when the postion `x` *or* `t` are NumPy arrays, in which case it should return a NumPy array itself.
# + nbgrader={"checksum": "b95685e8808cf7e99f918ab07c87c11a", "solution": true}
def soliton(x, t, c, a):
"""Return phi(x, t) for a soliton wave with constants c and a."""
solt=0.5*c*(1/np.cosh(0.5*(c**.5)*(x-c*t-a))**2)
return np.array(solt)
# -
x=np.array([1,2,3,4,5])
t=np.array([6,7,8,9,10])
soliton(x,t,1,2)
# + deletable=false nbgrader={"checksum": "bcd15232a87c4354cbc68dcca28654ee", "grade": true, "grade_id": "interactex03a", "points": 2}
assert np.allclose(soliton(np.array([0]),0.0,1.0,0.0), np.array([0.5]))
# + [markdown] nbgrader={}
# To create an animation of a soliton propagating in time, we are going to precompute the soliton data and store it in a 2d array. To set this up, we create the following variables and arrays:
# + nbgrader={}
tmin = 0.0
tmax = 10.0
tpoints = 100
t = np.linspace(tmin, tmax, tpoints)
xmin = 0.0
xmax = 10.0
xpoints = 200
x = np.linspace(xmin, xmax, xpoints)
c = 1.0
a = 0.0
# + [markdown] nbgrader={}
# Compute a 2d NumPy array called `phi`:
#
# * It should have a dtype of `float`.
# * It should have a shape of `(xpoints, tpoints)`.
# * `phi[i,j]` should contain the value $\phi(x[i],t[j])$.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
phi=np.ndarray(shape=(xpoints,tpoints), dtype=float) #collaberated with <NAME>
for i in x:
for j in t:
phi[i,j]=soliton(x[i],t[j],c,a)
phi
# + deletable=false nbgrader={"checksum": "90baf1a97272cee6f5554e0104b50f47", "grade": true, "grade_id": "interactex03b", "points": 4}
assert phi.shape==(xpoints, tpoints)
assert phi.ndim==2
assert phi.dtype==np.dtype(float)
assert phi[0,0]==soliton(x[0],t[0],c,a)
# + [markdown] nbgrader={}
# Write a `plot_soliton_data(i)` function that plots the soliton wave $\phi(x, t[i])$. Customize your plot to make it effective and beautiful.
# + nbgrader={"checksum": "d857aa7adb31b1de9c4d53a7febb18d3", "solution": true}
def plot_soliton_data(i=0):
"""Plot the soliton data at t[i] versus x."""
plt.plot(soliton(x,t[i],c,a))
# + nbgrader={}
plot_soliton_data(0)
# + deletable=false nbgrader={"checksum": "a76632040b08c7c76c889e67ee93deb0", "grade": true, "grade_id": "interactex03c", "points": 2}
assert True # leave this for grading the plot_soliton_data function
# + [markdown] nbgrader={}
# Use `interact` to animate the `plot_soliton_data` function versus time.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
interact(plot_soliton_data,i=(0,100,10))
# + deletable=false nbgrader={"checksum": "ef5ed9fcab6418650cdf556757a4486a", "grade": true, "grade_id": "interactex03d", "points": 2}
assert True # leave this for grading the interact with plot_soliton_data cell
# -
| assignments/assignment05/InteractEx03.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Case Study
#
# ## 1. Objective
#
# 1. Predicting most probable policy to a new customer
# 2. Recommending alternate policy to existing customers
# 3. Factors affecting life time value
# 4. Understanding demographics and customer behaviour
#
#
# Let's start with first part
#
# ## 2. Section 1 - Data Extraction
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
sns.set()
#Reading the dataset in a dataframe using Pandas
df = pd.read_csv("policy_data.csv", index_col = 'Customer')
columns = ['State','Coverage','Education','Gender','Income','Location Code',
'Marital Status','Sales Channel','Vehicle Class','Vehicle Size']
df.drop(['EmploymentStatus'], axis = 1, inplace = True)
df.drop(['Customer Lifetime Value'], axis = 1, inplace = True)
df.head()
# -
df.corr()
# ## 3. Exploratory Data Analysis
# First let us see the distribution of existing customers
# 
# ## Selection of Policy based on customer characteristics
# 
#
# ## Selection of Policy based on vehicle
# 
#
# ## 4. Modeling
from sklearn.preprocessing import LabelEncoder
categorical_variables = df.dtypes[df.dtypes == 'object'].index
categorical_variables
# +
le = LabelEncoder()
for var in categorical_variables:
df[var] = le.fit_transform(df[var])
df.head()
# -
X = df.iloc[:, 1:]
y = df.iloc[:, 0]
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = .2, random_state = 0)
from sklearn.tree import DecisionTreeClassifier
decision = DecisionTreeClassifier()
decision = decision.fit(X_train, y_train)
y_pred = decision.predict(X_test)
from sklearn.metrics import accuracy_score
result = accuracy_score(y_test, y_pred) * 100
result
for df, importance in zip(columns, decision.feature_importances_):
print(df, importance * 100)
from sklearn.naive_bayes import GaussianNB
GNBClassifier = GaussianNB()
GNBClassifier = GNBClassifier.fit(X_train, y_train.ravel())
y_pred = GNBClassifier.predict(X_test)
result = accuracy_score(y_test, y_pred) * 100
result
from sklearn.neighbors import KNeighborsClassifier
classifier = KNeighborsClassifier()
knnClassifier = classifier.fit(X_train, y_train.ravel())
y_pred = knnClassifier.predict(X_test)
result = accuracy_score(y_test, y_pred)*100
result
from sklearn.ensemble import RandomForestClassifier
classifier = RandomForestClassifier(max_depth = 10, min_samples_leaf = 10, max_features= 'auto')
rfClassifier = classifier.fit(X_train, y_train.ravel())
y_pred = rfClassifier.predict(X_test)
result = accuracy_score(y_test, y_pred)*100
result
from sklearn.externals import joblib
joblib.dump(rfClassifier, 'model/nb.pkl')
# ## 5. Serving end points with Flask API
#
# On localhost 5000
#
# 
# 
# 
#
# ## Section 2 - CLV
# 2.1 - Overall
#
# 
#
# 2.2 - Based on Complaints
#
# 
#
# 2.3 - Based on Demographics
#
# 
#
#
# 2.4 - Based on Sales
#
# 
#
# 2.5 - Based on Personal Characteristic
#
# 
#
#
#
#
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
sns.set()
#Reading the dataset in a dataframe using Pandas
df = pd.read_csv("data.csv")
# -
#finding missing values
df.isnull().sum()
from sklearn.preprocessing import LabelEncoder
categorical_variables = df.dtypes[df.dtypes == 'object'].index
categorical_variables
# +
le = LabelEncoder()
for var in categorical_variables:
df[var] = le.fit_transform(df[var])
df.head()
# -
pd.cut(df['Customer Lifetime Value'], 8).head()
custom_bucket_array = np.linspace(0, 20, 9)
custom_bucket_array
df['Customer Lifetime Value'] = pd.cut(df['Customer Lifetime Value'], custom_bucket_array)
X = df.iloc[:, 1:]
y = df.iloc[:, 0]
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = .2, random_state = 0)
from sklearn.tree import DecisionTreeRegressor
decision = DecisionTreeRegressor(max_depth=2)
decision = decision.fit(X_train, y_train)
y_pred = decision.predict(X_test)
for df, importance in zip(df, decision.feature_importances_):
print(df, importance * 100)
# ## Section 3 - Demographic
| master.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# 
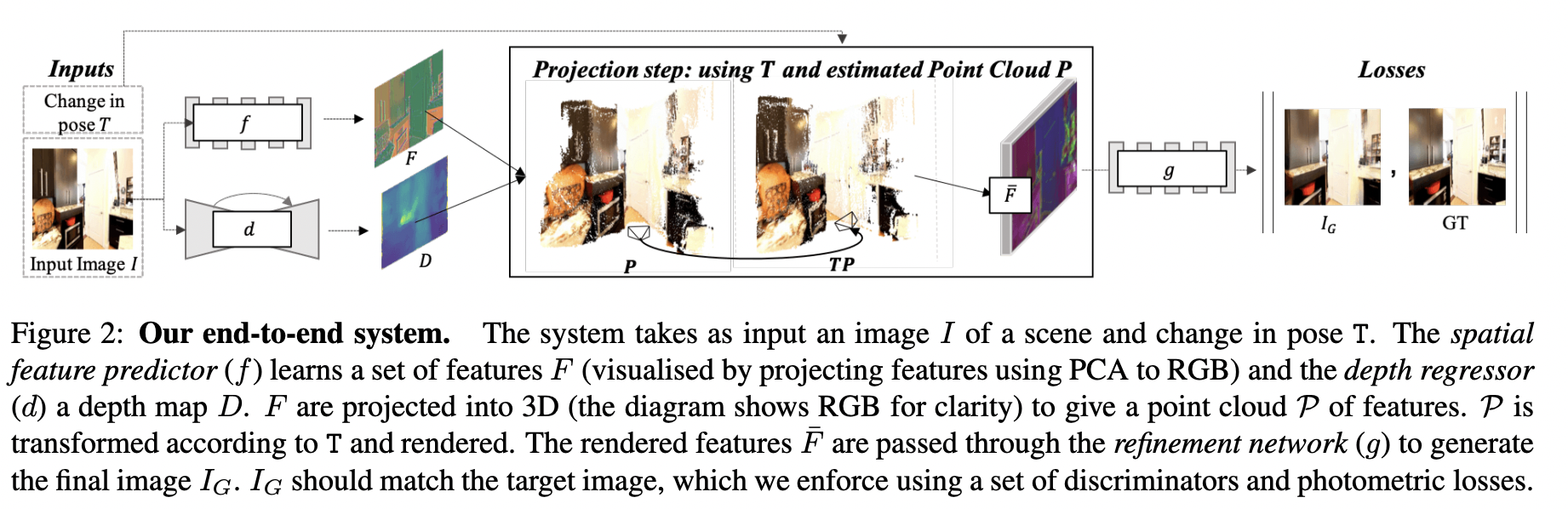
# In this homework, we will try to implement a `View Synthesis` model that allows us to generate new scene views based on a single image.
#
# The basic idea is to use `differentiable point cloud rendering`, which is used to convert a hidden 3D feature point cloud into a target view.
# The projected features are decoded by `refinement network` to inpaint missing regions and generate a realistic output image.
# ### Overall pipeline disribed below
# 
# # Data
# ## Download KITTI dataset
# +
# from gfile import download_file_from_google_drive
# download_file_from_google_drive(
# '1lqspXN10biBShBIVD0yvgnl1nIPPhRdC',
# 'kitti.zip'
# )
# +
# # !unzip kitti.zip
# -
# ## Dataset
# +
# %pylab inline
from tqdm import tqdm
from itertools import islice
from IPython.display import clear_output, HTML
from collections import defaultdict
from kitti import KITTIDataLoader
import torch
from torch import nn
from torch.utils.data import Subset, DataLoader
import torchvision
from pytorch3d.vis.plotly_vis import plot_scene
from pytorch3d.structures import Pointclouds
from pytorch3d.renderer import PerspectiveCameras, compositing, rasterize_points
# +
def split_RT(RT):
return RT[..., :3, :3], RT[..., :3, 3]
def renormalize_image(image):
return image * 0.5 + 0.5
# -
dataset = KITTIDataLoader('dataset_kitti')
# Each instance of dataset contain `source` and `target` images, `extrinsic` and `intrinsic` camera parameters for `source` and `targer` images.
#
# It is highly recommended to understand these concepts, e.g., here https://ksimek.github.io/2012/08/22/extrinsic/
images, cameras = dataset[0].values()
# +
plt.figure(figsize=(20, 10))
ax = plt.subplot(1, 2, 1)
ax.imshow(images[0].permute(1, 2, 0) * 0.5 + 0.5)
ax.set_title('Source Image Frame', fontsize=20)
ax.axis('off')
ax = plt.subplot(1, 2, 2)
ax.imshow(images[1].permute(1, 2, 0) * 0.5 + 0.5)
ax.set_title('Target Image Frame', fontsize=20)
ax.axis('off')
# +
source_camera = PerspectiveCameras(
R=split_RT(cameras[0]['P'])[0][None],
T=split_RT(cameras[0]['P'])[1][None],
K=torch.from_numpy(cameras[0]['K'])[None]
)
target_camera = PerspectiveCameras(
R=split_RT(cameras[1]['P'])[0][None],
T=split_RT(cameras[1]['P'])[1][None],
K=torch.from_numpy(cameras[1]['K'])[None]
)
plot_scene(
{
'scene': {
'source_camera': source_camera,
'target_camera': target_camera
}
},
)
# +
indexes = torch.randperm(len(dataset))
train_indexes = indexes[:-1000]
validation_indexes = indexes[-1000:]
train_dataset = Subset(dataset, train_indexes)
validation_dataset = Subset(dataset, validation_indexes)
train_dataloader = DataLoader(
train_dataset, batch_size=16, num_workers=6,
shuffle=True, drop_last=True, pin_memory=True
)
validation_dataloder = DataLoader(
validation_dataset, batch_size=10, num_workers=4,
pin_memory=True
)
# -
# ---
# # Models
#
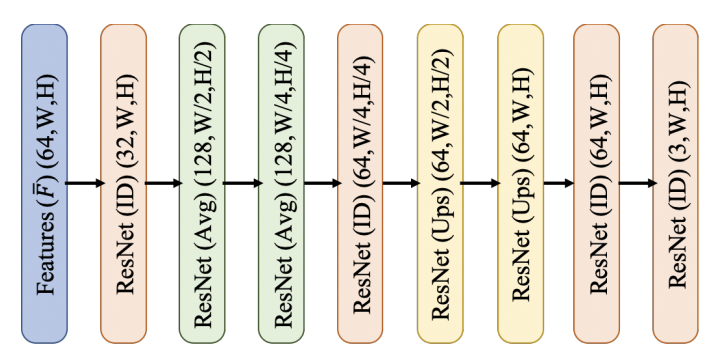
# So, we need to implement `Spatial Feature Predictor`, `Depth Regressor`, `Point Cloud Renderer` and `RefinementNetwork`.
# One of the main building blocks in these networks is `ResNetBlock`, but with some modifications:
#
# 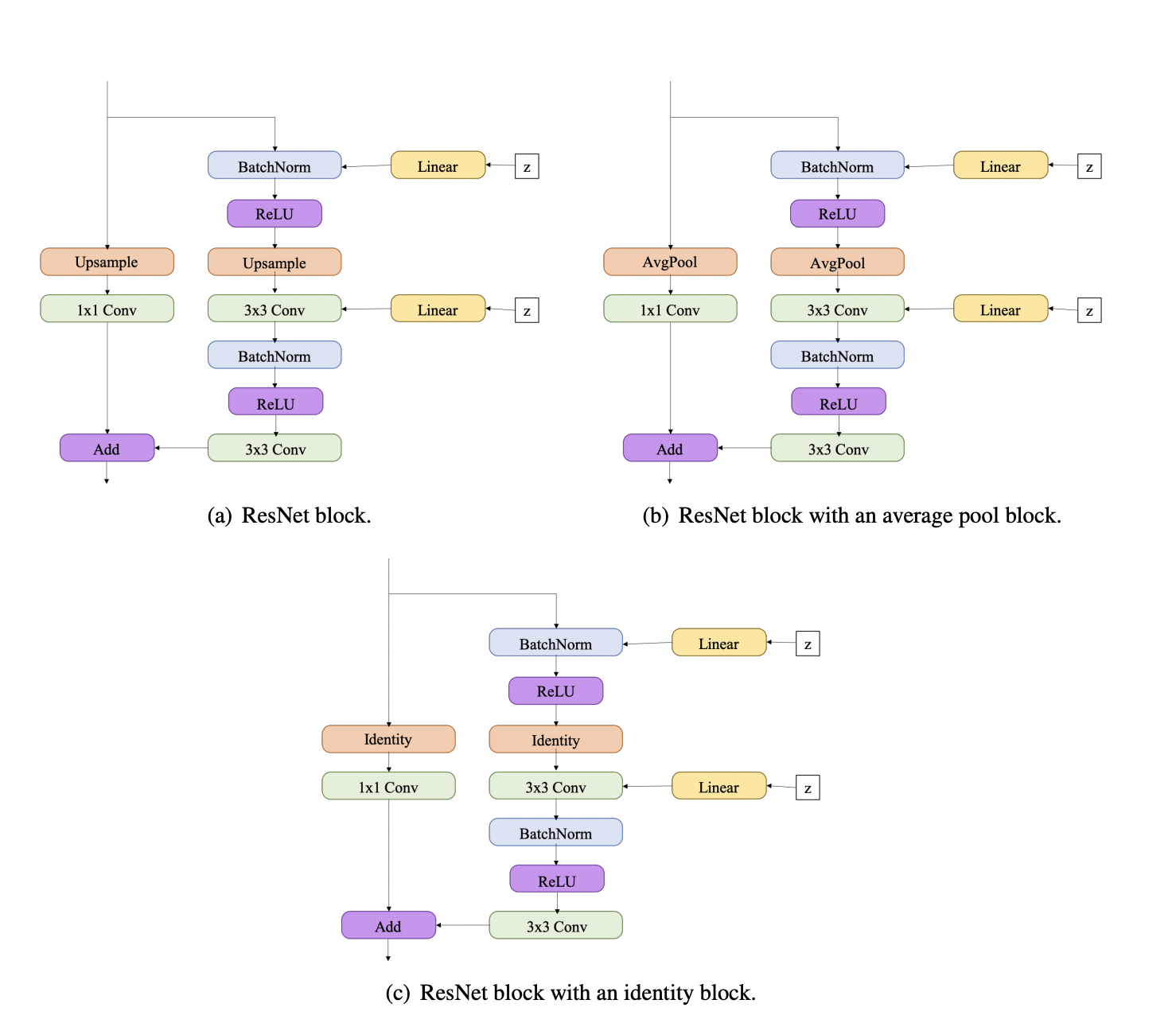
#
# So, let's implement it, but without the noise part `Linear + z` (let's omit it, since we do not use the adversarial criterion)
# + code_folding=[]
class ResNetBlock(nn.Module):
def __init__(
self,
in_channels: int,
out_channels: int,
stride: int = 1,
mode = 'identity'
):
super().__init__()
# TODO
def forward(self, input):
# TODO
# -
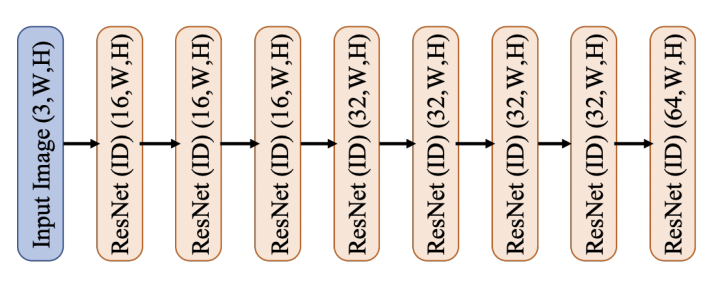
# ## Spatial Feature Predictor
#
# 
# + code_folding=[]
class SpatialFeatureNetwork(nn.Module):
def __init__(self, in_channels=3, out_channels=64):
super().__init__()
self.blocks = # TODO
def forward(self, input: torch.Tensor):
return self.blocks(input)
sf_net = SpatialFeatureNetwork()
# -
# ## Depth Regressor
#
# 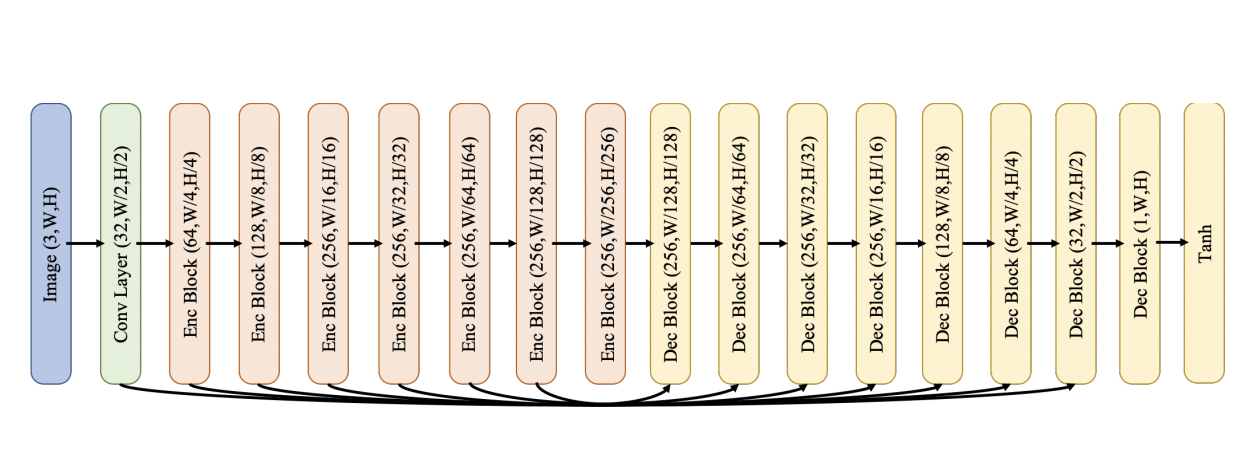
#
# An `Enc Block` consists of a sequence of Leaky ReLU, convolution (stride 2, padding 1, kernel size 4), and batch normalisation layers.
#
# A `Dec Block` consists of a sequence of ReLU, 2x bilinear upsampling, convolution (stride 1, padding 1, kernel size3), and batch normalisation layers (except for the final layer, which has no batch normalisation layer).
# + code_folding=[]
class Unet(nn.Module):
def __init__(
self,
num_filters=32,
channels_in=3,
channels_out=3
):
super(Unet, self).__init__()
# TODO
def forward(self, input):
# TODO
# -
# ## Refinement Network
#
# 
# + code_folding=[]
class RefinementNetwork(nn.Module):
def __init__(self, in_channels=64, out_channels=3):
super().__init__()
self.blocks = # TODO
def forward(self, input: torch.Tensor):
return self.blocks(input)
# -
# ## Auxiliary network
# + code_folding=[0]
class VGG19(nn.Module):
def __init__(self, requires_grad=False):
super().__init__()
vgg_pretrained_features = torchvision.models.vgg19(
pretrained=True
).features
self.slice1 = torch.nn.Sequential()
self.slice2 = torch.nn.Sequential()
self.slice3 = torch.nn.Sequential()
self.slice4 = torch.nn.Sequential()
self.slice5 = torch.nn.Sequential()
for x in range(2):
self.slice1.add_module(str(x), vgg_pretrained_features[x])
for x in range(2, 7):
self.slice2.add_module(str(x), vgg_pretrained_features[x])
for x in range(7, 12):
self.slice3.add_module(str(x), vgg_pretrained_features[x])
for x in range(12, 21):
self.slice4.add_module(str(x), vgg_pretrained_features[x])
for x in range(21, 30):
self.slice5.add_module(str(x), vgg_pretrained_features[x])
if not requires_grad:
for param in self.parameters():
param.requires_grad = False
def forward(self, X):
# Normalize the image so that it is in the appropriate range
h_relu1 = self.slice1(X)
h_relu2 = self.slice2(h_relu1)
h_relu3 = self.slice3(h_relu2)
h_relu4 = self.slice4(h_relu3)
h_relu5 = self.slice5(h_relu4)
out = [h_relu1, h_relu2, h_relu3, h_relu4, h_relu5]
return out
# -
# ---
# # Criterions & Metrics
# + code_folding=[0]
class PerceptualLoss(nn.Module):
def __init__(self):
super().__init__()
# Set to false so that this part of the network is frozen
self.model = VGG19(requires_grad=False)
self.criterion = nn.L1Loss()
self.weights = [1.0 / 32, 1.0 / 16, 1.0 / 8, 1.0 / 4, 1.0]
def forward(self, pred_img, gt_img):
gt_fs = self.model(gt_img)
pred_fs = self.model(pred_img)
# Collect the losses at multiple layers (need unsqueeze in
# order to concatenate these together)
loss = 0
for i in range(0, len(gt_fs)):
loss += self.weights[i] * self.criterion(pred_fs[i], gt_fs[i])
return loss
# + code_folding=[0]
def psnr(predicted_image, target_image):
batch_size = predicted_image.size(0)
mse_err = (
(predicted_image - target_image)
.pow(2).sum(dim=1)
.view(batch_size, -1).mean(dim=1)
)
psnr = 10 * (1 / mse_err).log10()
return psnr.mean()
# -
# ---
# # Point Cloud Renderer
#
# `Differential Rasterization` is a key component of our system. We will use the algorithm already implemented in `pytorch3d`.
#
# 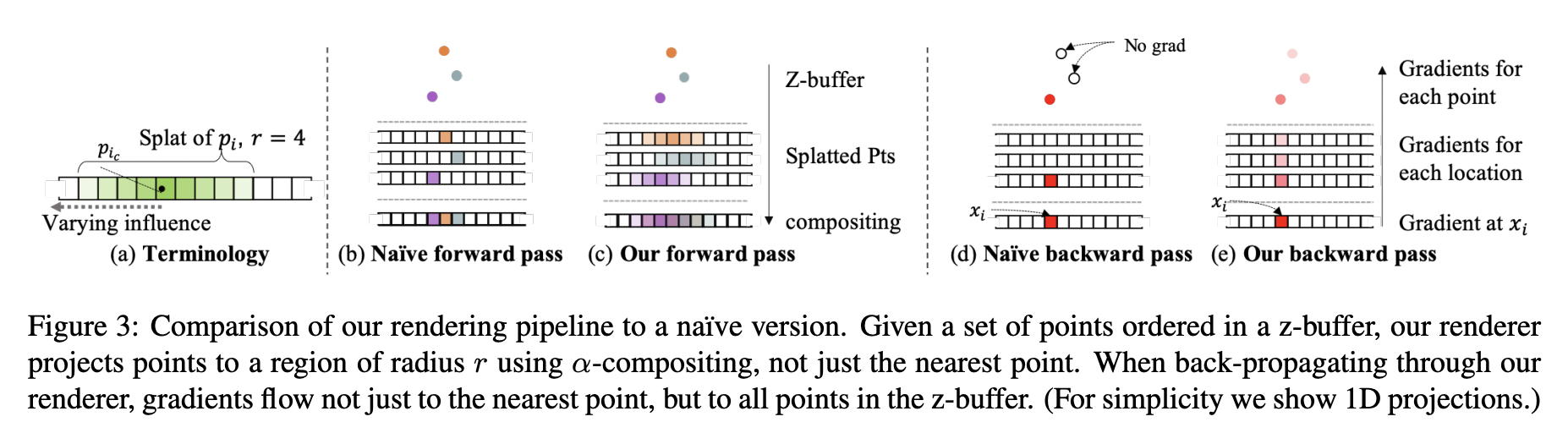
#
# For more details read (3.2) https://arxiv.org/pdf/1912.08804.pdf
# + code_folding=[]
class PointsRasterizerWithBlending(nn.Module):
"""
Rasterizes a set of points using a differentiable renderer.
"""
def __init__(self, radius=1.5, image_size=256, points_per_pixel=8):
super().__init__()
self.radius = radius
self.image_size = image_size
self.points_per_pixel = points_per_pixel
self.rad_pow = 2
self.tau = 1.0
def forward(self, point_cloud, spatial_features):
batch_size = spatial_features.size(0)
# Make sure these have been arranged in the same way
assert point_cloud.size(2) == 3
assert point_cloud.size(1) == spatial_features.size(2)
point_cloud[:, :, 1] = -point_cloud[:, :, 1]
point_cloud[:, :, 0] = -point_cloud[:, :, 0]
radius = float(self.radius) / float(self.image_size) * 2.0
point_cloud = Pointclouds(points=point_cloud, features=spatial_features.permute(0, 2, 1))
points_idx, _, dist = rasterize_points(
point_cloud, self.image_size, radius, self.points_per_pixel
)
dist = dist / pow(radius, self.rad_pow)
alphas = (
(1 - dist.clamp(max=1, min=1e-3).pow(0.5))
.pow(self.tau)
.permute(0, 3, 1, 2)
)
transformed_src_alphas = compositing.alpha_composite(
points_idx.permute(0, 3, 1, 2).long(),
alphas,
point_cloud.features_packed().permute(1, 0),
)
return transformed_src_alphas
# -
# And `PointsManipulator` do the following steps:
#
# 1) Create virtual image place in [normalized coordinate](https://pytorch3d.org/docs/cameras)
# 2) Move camera according to `regressed depth`
# 3) Rotate points according to target camera paramers
# 4) And finally render them with help of `PointsRasterizerWithBlending`
# + code_folding=[]
class PointsManipulator(nn.Module):
EPS = 1e-5
def __init__(self, image_size):
super().__init__()
# Assume that image plane is square
self.splatter = PointsRasterizerWithBlending(
radius=1.0,
image_size=image_size,
points_per_pixel=128,
)
xs = torch.linspace(0, image_size - 1, image_size) / \
float(image_size - 1) * 2 - 1
ys = torch.linspace(0, image_size - 1, image_size) / \
float(image_size - 1) * 2 - 1
xs = xs.view(1, 1, 1, image_size).repeat(1, 1, image_size, 1)
ys = ys.view(1, 1, image_size, 1).repeat(1, 1, 1, image_size)
xyzs = torch.cat(
(xs, -ys, -torch.ones(xs.size()), torch.ones(xs.size())), 1
).view(1, 4, -1)
self.register_buffer("xyzs", xyzs)
def project_pts(self, depth, K, K_inv, RT_cam1, RTinv_cam1, RT_cam2, RTinv_cam2):
# Project the world points into the new view
projected_coors = self.xyzs * depth
projected_coors[:, -1, :] = 1
# Transform into camera coordinate of the first view
cam1_X = K_inv.bmm(projected_coors)
# Transform into world coordinates
RT = RT_cam2.bmm(RTinv_cam1)
wrld_X = RT.bmm(cam1_X)
# And intrinsics
xy_proj = K.bmm(wrld_X)
# And finally we project to get the final result
mask = (xy_proj[:, 2:3, :].abs() < self.EPS).detach()
# Remove invalid zs that cause nans
zs = xy_proj[:, 2:3, :]
zs[mask] = self.EPS
sampler = torch.cat((xy_proj[:, 0:2, :] / -zs, xy_proj[:, 2:3, :]), 1)
sampler[mask.repeat(1, 3, 1)] = -10
# Flip the ys
sampler = sampler * torch.Tensor([1, -1, -1]).unsqueeze(0).unsqueeze(
2
).to(sampler.device)
return sampler
def forward_justpts(
self,
spatial_features, depth,
K, K_inv, RT_cam1, RTinv_cam1, RT_cam2, RTinv_cam2
):
# Now project these points into a new view
batch_size, c, w, h = spatial_features.size()
if len(depth.size()) > 3:
# reshape into the right positioning
depth = depth.view(batch_size, 1, -1)
spatial_features = spatial_features.view(batch_size, c, -1)
pointcloud = self.project_pts(
depth, K, K_inv, RT_cam1, RTinv_cam1, RT_cam2, RTinv_cam2
)
pointcloud = pointcloud.permute(0, 2, 1).contiguous()
result = self.splatter(pointcloud, spatial_features)
return result
# -
# ---
# # All together
class ViewSynthesisModel(nn.Module):
def __init__(self):
super().__init__()
self.spatial_feature_predictor = SpatialFeatureNetwork()
self.depth_regressor = Unet(channels_in=3, channels_out=1)
self.point_cloud_renderer = PointsManipulator(image_size=256)
self.refinement_network = RefinementNetwork()
# Special constant for KITTI dataset
self.z_min = 1.0
self.z_max = 50.0
def forward(self):
# TODO
# 1) Predict spatial feature for source image
# 2) Predict depth for source image (dont forget to renormalize depth with z_min/z_max)
# 3) Generate new features with `point_cloud_renderer`
# 4) And finnaly apply `refinement_network` to obtain new image
# 5) return new image, and depth of source image
# ---
# # Training
#
# In order for the work to be accepted, you must achieve a quality of ~0.5 (validation loss value) and visualize several samples as in the example
# +
device = torch.device('cuda:0')
model = ViewSynthesisModel().to(device)
optimizer = torch.optim.Adam(model.parameters(), 1e-4)
histoty = defaultdict(list)
l1_criterion = nn.L1Loss()
perceptual_criterion = PerceptualLoss().to(device)
# -
for epoch in range(10):
for i, batch in tqdm(enumerate(train_dataloader, 2), total=len(train_dataloader)):
source_image = batch["images"][0].to(device)
target_image = batch["images"][-1].to(device)
# TODO
generated_image, regressed_depth = model(...)
loss = l1_criterion(generated_image, target_image) \
+ 10 * perceptual_criterion(
renormalize_image(generated_image),
renormalize_image(target_image)
)
optimizer.zero_grad()
loss.backward()
optimizer.step()
histoty['train_loss'].append(loss.item())
for i, batch in tqdm(enumerate(validation_dataloder), total=len(validation_dataloder)):
source_image = batch["images"][0].to(device)
target_image = batch["images"][-1].to(device)
with torch.no_grad():
# TODO
generated_image, regressed_depth = model(...)
loss = l1_criterion(generated_image, target_image) \
+ 10 * perceptual_criterion(
renormalize_image(generated_image),
renormalize_image(target_image)
)
histoty['validation_loss'].append(loss.item())
clear_output()
fig = plt.figure(figsize=(30, 15), dpi=80)
ax1 = plt.subplot(3, 3, 1)
ax1.plot(histoty['train_loss'], label='Train')
ax1.set_xlabel('Iterations', fontsize=20)
ax1.set_ylabel(r'${L_1} + Perceptual$', fontsize=20)
ax1.legend()
ax1.grid()
ax2 = plt.subplot(3, 3, 4)
ax2.plot(histoty['validation_loss'], label='Validation')
ax2.set_xlabel('Iterations', fontsize=20)
ax2.set_ylabel(r'${L_1} + Perceptual$', fontsize=20)
ax2.legend()
ax2.grid()
for index, image in zip(
(2, 3, 5, 6),
(source_image, target_image, generated_image, regressed_depth)
):
ax = plt.subplot(3, 3, index)
im = ax.imshow(renormalize_image(image.detach().cpu()[0]).permute(1, 2, 0))
ax.axis('off')
plt.show()
# # Visualize
#
# Goes along depth and generate new views
RTs = []
for i in torch.linspace(0, 0.5, 40):
current_RT = torch.eye(4).unsqueeze(0)
current_RT[:, 2, 3] = i
RTs.append(current_RT.to(device))
identity_matrx = torch.eye(4).unsqueeze(0).to(device)
# +
random_instance_index = 245
with torch.no_grad():
images, cameras = validation_dataset[random_instance_index].values()
# Input values
input_img = images[0][None].cuda()
# Camera parameters
K = torch.from_numpy(cameras[0]["K"])[None].to(device)
K_inv = torch.from_numpy(cameras[0]["Kinv"])[None].to(device)
spatial_features = model.spatial_feature_predictor(input_img)
regressed_depth = torch.sigmoid(model.depth_regressor(input_img)) * \
(model.z_max - model.z_min) + model.z_min
new_images = []
for current_RT in RTs:
generated_features = model.point_cloud_renderer.forward_justpts(
spatial_features,
regressed_depth,
K,
K_inv,
identity_matrx,
identity_matrx,
current_RT,
None
)
generated_image = model.refinement_network(generated_features)
new_images.append(renormalize_image(generated_image.cpu()).clamp(0, 1).mul(255).to(torch.uint8))
# -
frames = torch.cat(new_images).permute(0, 2, 3, 1)
torchvision.io.write_video('video.mp4', frames, fps=20)
HTML("""
<video width="256" alt="test" controls>
<source src="video.mp4" type="video/mp4">
</video>
""")
# # Quality benchmark
# 
# 
# 
| homework04/homework-part2-new-view-synthesis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from keras.models import load_model, Model
from keras.layers import Dense, Flatten, MaxPooling2D
from keras.layers import Conv2D, Lambda, Input, Activation
from keras.layers import LSTM, TimeDistributed, Bidirectional, GRU
from keras.layers.merge import add, concatenate
from keras.optimizers import SGD
from keras import backend as K
from new_multi_gpu import *
from models_multi import *
# import warpctc_tensorflow
import tensorflow as tf
import random
import keras
import numpy as np
# class CRNN(object):
# """docstring for RNN"""
# def __init__(self, learning_rate = 0.001, output_dim = 63, gpu_count=2):
# conv_filters = 16
# kernel_size = (3, 3)
# pool_size = 2
# time_dense_size = 32
# rnn_size = 512
# img_h = 32
# act = 'relu'
# self.width = K.placeholder(name= 'width', ndim =0, dtype='int32')
# self.input_data = Input(name='the_input', shape=(None, img_h, 1), dtype='float32')
# self.inner = Conv2D(conv_filters, kernel_size, padding='same',
# activation=act, kernel_initializer='he_normal',
# name='conv1')(self.input_data)
# self.inner = MaxPooling2D(pool_size=(pool_size, pool_size), name='max1')(self.inner)
# self.inner = Conv2D(conv_filters, kernel_size, padding='same',
# activation=act, kernel_initializer='he_normal',
# name='conv2')(self.inner)
# self.inner = MaxPooling2D(pool_size=(pool_size, pool_size), name='max2')(self.inner)
# self.inner = Lambda(self.res, arguments={"last_dim": (img_h // (pool_size ** 2)) * conv_filters \
# , "width": self.width // 4})(self.inner)
# # cuts down input size going into RNN:
# self.inp = Dense(time_dense_size, activation=act, name='dense1')(self.inner)
# self.batch_norm = keras.layers.normalization.BatchNormalization()(self.inp)
# self.gru_1 = Bidirectional(GRU(rnn_size, return_sequences=True, kernel_initializer='he_normal',\
# name='gru1'),merge_mode="sum")(self.batch_norm)
# self.gru_2 = Bidirectional(GRU(rnn_size, return_sequences=True, kernel_initializer='he_normal',\
# name='gru2'),merge_mode="concat")(self.gru_1)
# self.gru_3 = Bidirectional(GRU(rnn_size, recurrent_dropout=0.5, return_sequences=True, \
# kernel_initializer='he_normal', name='gru3'),merge_mode="concat")(self.gru_2)
# self.gru_4 = Bidirectional(GRU(rnn_size, recurrent_dropout=0.5, return_sequences=True, \
# kernel_initializer='he_normal', name='gru4'),merge_mode="concat")(self.gru_3)
# self.y_pred = TimeDistributed(Dense(output_dim, kernel_initializer='he_normal', \
# name='dense2', activation='linear'))(self.gru_4)
# self.model = Model(inputs=self.input_data, outputs=self.y_pred)
# self.model = make_parallel(self.model, gpu_count)
# self.model.summary()
# self.output_ctc = self.model.outputs[0]
# self.out = K.function([self.input_data, self.width, K.learning_phase()], [self.y_pred])
# self.y_true = K.placeholder(name='y_true', ndim=1, dtype='int32')
# self.input_length = K.placeholder(name='input_length', ndim=1, dtype='int32')
# self.label_length = K.placeholder(name='label_length', ndim=1, dtype='int32')
# self.test = K.argmax(self.y_pred, axis=2)
# self.predict_step = K.function([self.input_data, self.width, K.learning_phase()], [self.test])
# def res (self, x, width, last_dim):
# return K.reshape(x, (-1, width, last_dim))
# -
from models_multi import CRNN
M = CRNN(1e-4, 219)
M.model.load_weights('crnn_219.h5')
model = M.model.get_layer('model_1')
model.summary()
model.layers.pop()
from keras.layers import TimeDistributed, Dense
from keras.models import Model
x = model.layers[-1].output
y_pred = TimeDistributed(Dense(219, kernel_initializer='he_normal', \
name='denseout', activation='linear'))(x)
new_model = Model(input=model.inputs, output=y_pred)
new_model.summary()
from new_multi_gpu import make_parallel
final_model = make_parallel(new_model, 2)
final_model.summary()
final_model.save_weights('crnn_219.h5')
# +
import matplotlib.pyplot as plt
import numpy as np
import os
from utils import pred, reshape
from scipy import ndimage
ims = os.listdir('./test/nhu cầu đầu vào')
im = './test/nhu cầu đầu vào/' + np.random.choice(ims)
# im = '/home/tailongnguyen/deep-anpr/output/0.png'
im = ndimage.imread(im)
plt.imshow(im, cmap ='gray')
plt.show()
im = np.expand_dims(reshape(im), axis = 0)
im.shape
pred(im, M, None, True)
# -
| ipython/Test-sentence.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from pycalphad import Database, Model, calculate, equilibrium, variables as v
from xarray import DataArray
# +
class PrecipitateModel(Model):
matrix_chempots = []
@property
def matrix_hyperplane(self):
return sum(self.moles(self.nonvacant_elements[i])*self.matrix_chempots[i]
for i in range(len(self.nonvacant_elements)))
@property
def GM(self):
return self.ast - self.matrix_hyperplane
class GibbsThompsonModel(Model):
"Spherical particle."
radius = 1e-6 # m
volume = 7.3e-6 # m^3/mol
interfacial_energy = 250e-3 # J/m^2
elastic_misfit_energy = 0 # J/mol
@property
def GM(self):
return self.ast + (2*self.interfacial_energy/self.radius + self.elastic_misfit_energy) * self.volume
# +
import numpy as np
def parallel_tangent(dbf, comps, matrix_phase, matrix_comp, precipitate_phase, temp):
conds = {v.N: 1, v.P: 1e5, v.T: temp}
conds.update(matrix_comp)
matrix_eq = equilibrium(dbf, comps, matrix_phase, conds)
# pycalphad currently doesn't have a way to turn global minimization off and directly specify starting points
if matrix_eq.isel(vertex=1).Phase.values.flatten() != ['']:
raise ValueError('Matrix phase has miscibility gap. This bug will be fixed in the future')
matrix_chempots = matrix_eq.MU.values.flatten()
# This part will not work until mass balance constraint can be relaxed
#precip = PrecipitateModel(dbf, comps, precipitate_phase)
#precip.matrix_chempots = matrix_chempots
#conds = {v.N: 1, v.P: 1e5, v.T: temp}
#df_eq = equilibrium(dbf, comps, precipitate_phase, conds, model=precip)
df_eq = calculate(dbf, comps, precipitate_phase, T=temp, N=1, P=1e5)
df_eq['GM'] = df_eq.X.values[0,0,0].dot(matrix_chempots) - df_eq.GM
selected_idx = df_eq.GM.argmax()
return matrix_eq.isel(vertex=0), df_eq.isel(points=selected_idx)
def nucleation_barrier(dbf, comps, matrix_phase, matrix_comp, precipitate_phase, temp,
interfacial_energy, precipitate_volume):
"Spherical precipitate."
matrix_eq, precip_eq = parallel_tangent(dbf, comps, matrix_phase, matrix_comp, precipitate_phase, temp)
precip_driving_force = float(precip_eq.GM.values) # J/mol
elastic_misfit_energy = 0 # J/m^3
barrier = 16./3 * np.pi * interfacial_energy **3 / (precip_driving_force + elastic_misfit_energy) ** 2 # J/mol
critical_radius = 2 * interfacial_energy / ((precip_driving_force / precipitate_volume) + elastic_misfit_energy) # m
print(temp, critical_radius)
if critical_radius < 0:
barrier = np.inf
cluster_area = 4*np.pi*critical_radius**2
cluster_volume = (4./3) * np.pi * critical_radius**3
return barrier, precip_driving_force, cluster_area, cluster_volume, matrix_eq, precip_eq
def growth_rate(dbf, comps, matrix_phase, matrix_comp, precipitate_phase, temp, particle_radius):
conds = {v.N: 1, v.P: 1e5, v.T: temp}
conds.update(matrix_comp)
# Fictive; Could retrieve from TDB in principle
mobility = 1e-7 * np.exp(-14e4/(8.3145*temp)) # m^2 / s
mobilities = np.eye(len(matrix_comp)+1) * mobility
matrix_ff_eq, precip_eq = parallel_tangent(dbf, comps, matrix_phase, matrix_comp, precipitate_phase, temp)
precip_driving_force = float(precip_eq.GM.values) # J/mol
interfacial_energy = 250e-3 # J/m^2
precipitate_volume = 7.3e-6 # m^3/mol
elastic_misfit_energy = 0 # J/m^3
# Spherical particle
critical_radius = 2 * interfacial_energy / ((precip_driving_force / precipitate_volume) + elastic_misfit_energy) # m
# XXX: Should really be done with global min off, fixed-phase conditions, etc.
# As written, this will break with miscibility gaps
particle_mod = GibbsThompsonModel(dbf, comps, precipitate_phase)
particle_mod.radius = particle_radius
interface_eq = equilibrium(dbf, comps, [matrix_phase, precipitate_phase], conds,
model={precipitate_phase: particle_mod})
matrix_idx = np.nonzero((interface_eq.Phase==matrix_phase).values.flatten())[0]
if len(matrix_idx) > 1:
raise ValueError('Matrix phase has miscibility gap')
elif len(matrix_idx) == 0:
# Matrix is metastable at this composition; massive transformation kinetics?
print(interface_eq)
raise ValueError('Matrix phase is not stable')
else:
matrix_idx = matrix_idx[0]
matrix_interface_eq = interface_eq.isel(vertex=matrix_idx)
precip_idx = np.nonzero((interface_eq.Phase==precipitate_phase).values.flatten())[0]
if len(precip_idx) > 1:
raise ValueError('Precipitate phase has miscibility gap')
elif len(precip_idx) == 0:
precip_conc = np.zeros(len(interface_eq.component))
# Precipitate is metastable at this radius (it will start to dissolve)
# Compute equilibrium for precipitate by itself at parallel tangent composition
pt_comp = {v.X(str(comp)): precip_eq.X.values.flatten()[idx] for idx, comp in enumerate(precip_eq.component.values[:-1])}
conds = {v.N: 1, v.P: 1e5, v.T: temp}
conds.update(pt_comp)
precip_interface_eq = equilibrium(dbf, comps, precipitate_phase, conds,
model={precipitate_phase: particle_mod})
precip_interface_eq = precip_interface_eq.isel(vertex=0)
else:
precip_idx = precip_idx[0]
precip_interface_eq = interface_eq.isel(vertex=precip_idx)
eta_geo_factor = 1.0
# Growth equation from Eq. 12, <NAME> and <NAME>, Calphad, 2019
velocity = 2*interfacial_energy * precipitate_volume * (1./critical_radius - 1./particle_radius)
x_int_precip = precip_interface_eq.X.values.flatten()
x_int_matrix = matrix_interface_eq.X.values.flatten()
denominator = 0
for i in range(mobilities.shape[0]-1):
for j in range(mobilities.shape[1]-1):
denominator += (x_int_precip[i] - x_int_matrix[i]) * (x_int_precip[j] - x_int_matrix[j]) / mobilities[i,j]
velocity /= denominator
return velocity
# +
dbf = Database('Al-Cu-Zr_Zhou.tdb')
comps = ['AL', 'CU', 'VA']
matrix_compositions = np.linspace(1e-4, 1-1e-4, num=30)
particle_radii = np.logspace(-10, -3, num=60)
velocities = np.zeros((matrix_compositions.shape[0], particle_radii.shape[0]))
for mc_idx in range(velocities.shape[0]):
matrix_comp = {v.X('CU'): matrix_compositions[mc_idx]}
for r_idx in range(velocities.shape[1]):
print(mc_idx, r_idx)
velocities[mc_idx, r_idx] = growth_rate(dbf, comps, 'BCC_A2', matrix_comp, 'ETA2', 800, particle_radii[r_idx])
# -
from pycalphad import calculate
import matplotlib.pyplot as plt
X, Y = np.meshgrid(matrix_compositions, np.log(particle_radii))
plt.contourf(X, Y, velocities.T, levels=5)
# +
import matplotlib.pyplot as plt
plt.plot(matrix_compositions, velocities[:,55])
# -
| GrowthRate-Interp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NYX Taxi trip predicton
import tensorflow
import pandas as pd
df1 = pd.read_csv('NYCTaxiFares.csv')
df = pd.read_csv('taxifare.csv')
df.head()
df1.head()
df.shape
# checking features
df.info()
# We are performing feature engineering with respect to date time
import datetime
pd.to_datetime(df['pickup_datetime'])
df['pickup_datetime'] = pd.to_datetime(df['pickup_datetime'])-datetime.timedelta(hours=4)
df.info()
df.head()
df['Year'] = df['pickup_datetime'].dt.year
df['Month'] = df['pickup_datetime'].dt.month
df['Days'] = df['pickup_datetime'].dt.day
df['Hours'] = df['pickup_datetime'].dt.hour
df['Minutes'] = df['pickup_datetime'].dt.minute
df.shape
df.head()
import numpy as np
df['mornornight'] = np.where(df['Hours']<12,0,1)
# Adding one hot encoding kind of structure where if less than 12 hours "0" shows morning else "1" is night
df.head()
df.drop('pickup_datetime',axis=1,inplace=True)
df.head()
df['fare_class'].unique()
from sklearn.metrics.pairwise import haversine_distances
from math import radians
newdelhi=[28.6139,77.2090]
bangalore=[12.9716,77.5946]
newdelhi_in_radians = [radians(_) for _ in newdelhi]
bangalore_in_radians = [radians(_) for _ in bangalore]
result = haversine_distances([newdelhi_in_radians,bangalore_in_radians])
result
##Just multiplying result by radius of earth
result*6371
import numpy as np
def haversine(df):
"""
Calculate the great circle distance in kilometers between two points
on the earth (specified in decimal degrees)
"""
# convert decimal degrees to radians
lat1 = np.radians(df["pickup_latitude"])
lat2 = np.radians(df["pickup_longitude"])
# haversine formula
dlat =np.radians(df["dropoff_latitude"]-df["pickup_latitude"])
dlon = np.radians(df["dropoff_longitude"]-df["pickup_longitude"])
a = np.sin(dlat/2)**2 + np.cos(lat1) * np.cos(lat2) * np.sin(dlon/2)**2
c = 2 * np.arctan2(np.sqrt(a),np.sqrt(1-a))
r = 6371 # Radius of earth in kilometers. Use 3956 for miles. Determines return value units.
return c * r
df['Total_distance']=haversine(df)
df.head()
df.drop(["dropoff_latitude","dropoff_longitude","pickup_latitude","pickup_longitude"],axis=1,inplace=True)
df.head()
# # Storing Data in MongoDB
import json
records = json.loads(df.T.to_json()).values()
# !pip install pymongo
import pymongo
client = pymongo.MongoClient('mongodb://localhost:27017/')
db = client["newyorktaxi"]
col = db["rides"]
# + active=""
# records
# -
col.insert_many(records)
# # Use of Regression model
X = df.iloc[:,1:]
Y = df.iloc[:,0]
#feature importance
from sklearn.ensemble import ExtraTreesRegressor
import matplotlib.pyplot as plt
model = ExtraTreesRegressor()
model.fit(X,Y)
### Plot graph of feature importances for better visualisation
ft_importance = pd.Series(model.feature_importances_, index=X.columns)
ft_importance.nlargest(7).plot(kind='barh')
plt.show()
X.head()
Y.head()
from sklearn.model_selection import train_test_split
X_train,X_test,Y_train,Y_test=train_test_split(X,Y,test_size=0.3,random_state=100)
pip install xgboost
import xgboost
regressor = xgboost.XGBRegressor()
regressor.fit(X_train,Y_train)
y_pred = regressor.predict(X_test)
import matplotlib.pyplot as plt
import seaborn as sns
sns.displot(Y_test-y_pred)
##If you get this kind of gausian distribution between the y test and y pred it indicates that the model is pretty
##good
plt.scatter(Y_test,y_pred)
from sklearn import metrics
print('R-Square :',np.sqrt(metrics.r2_score(Y_test,y_pred)))
print('MAE:',metrics.mean_absolute_error(Y_test,y_pred))
print('MSE:',metrics.mean_squared_error(Y_test,y_pred))
print('RMSE:',np.sqrt(metrics.mean_squared_error(Y_test,y_pred)))
# # Hyperparameter Tuning
from sklearn.model_selection import RandomizedSearchCV
n_estimators = [int(x) for x in np.linspace(start=100,stop=1200,num=12)]
print(n_estimators)
# +
#Randomized Search CV
#Number of trees in Xgboost
n_estimators = [int(x) for x in np.linspace(start=100,stop=1200,num=12)]
#Various learning rates
learning_rate=['0.05','0.1','0.2','0.3','0.4','0.5','0.6']
#Max number of levels in tree
max_depth = [int(x) for x in np.linspace(5,30,num=6)]
# minimum child weight parameter
min_child_weight=[3,4,5,6,7]
#subsample param values
subsample=[0.7,0.6,0.5,0.4]
# -
#Random Grid
random_grid = {'n_estimators':n_estimators,
'learning_rate':learning_rate,
'max_depth':max_depth,
'min_child_weight':min_child_weight,
'subsample':subsample}
print(random_grid)
# Use random grid to search for best param and search across 100 different combinations,
regressor = xgboost.XGBRegressor()
#Random search of parameters using 3 fold cross validation, search across 100 diff. combos
xg_random = RandomizedSearchCV(estimator=regressor,param_distributions=random_grid,scoring='neg_mean_squared_error',n_iter=3,cv=5,verbose=2,random_state=42,n_jobs=1)
xg_random.fit(X_train,Y_train)
xg_random.best_params_
y_pred = xg_random.predict(X_test)
sns.distplot(Y_test-y_pred)
plt.scatter(Y_test,y_pred)
from sklearn import metrics
print('R-Square :',np.sqrt(metrics.r2_score(Y_test,y_pred)))
print('MAE:',metrics.mean_absolute_error(Y_test,y_pred))
print('MSE:',metrics.mean_squared_error(Y_test,y_pred))
print('RMSE:',np.sqrt(metrics.mean_squared_error(Y_test,y_pred)))
| NYtaxipricepred.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import os
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ## Read Data
items = pd.read_csv('items.csv')
shops = pd.read_csv('shops.csv')
item_cats = pd.read_csv('item_categories.csv')
sales = pd.read_csv('sales_train.csv')
test = pd.read_csv('test.csv')
items.head(5)
shops.head(5)
item_cats.head(5)
sales.head(5)
test.head(5)
# ## Exploratory Data Analysis
from datetime import datetime
sales['year'] = pd.to_datetime(sales['date']).dt.strftime('%Y')
sales['month'] = sales.date.apply(lambda x: datetime.strptime(x,'%d.%m.%Y').strftime('%m'))
sales.head(2)
grouped = pd.DataFrame(sales.groupby(['year','month'])['item_cnt_day'].sum().reset_index())
sns.pointplot(x='month', y='item_cnt_day', hue='year', data=grouped)
grouped_price = pd.DataFrame(sales.groupby(['year','month'])['item_price'].mean().reset_index())
sns.pointplot(x='month', y='item_price', hue='year', data=grouped_price)
ts=sales.groupby(["date_block_num"])["item_cnt_day"].sum()
ts.astype('float')
plt.figure(figsize=(16,8))
plt.title('Total Sales of the whole time period')
plt.xlabel('Time')
plt.ylabel('Sales')
plt.plot(ts);
sns.jointplot(x="item_cnt_day", y="item_price", data=sales, height=8)
plt.show()
sales.item_cnt_day.hist(bins=100)
sales.item_cnt_day.describe()
# ## Data Cleaning
print('Data set size before remove item price 0 cleaning:', sales.shape)
sales = sales.query('item_price > 0')
print('Data set size after remove item price 0 cleaning:', sales.shape)
print('Data set size before filter valid:', sales.shape)
sales = sales[sales['shop_id'].isin(test['shop_id'].unique())]
sales = sales[sales['item_id'].isin(test['item_id'].unique())]
print('Data set size after filter valid:', sales.shape)
print('Data set size before remove outliers:', sales.shape)
sales = sales.query('item_cnt_day >= 0 and item_cnt_day <= 125 and item_price < 75000')
print('Data set size after remove outliers:', sales.shape)
# +
sns.jointplot(x="item_cnt_day", y="item_price", data=sales, height=8)
plt.show()
cleaned = pd.DataFrame(sales.groupby(['year','month'])['item_cnt_day'].sum().reset_index())
sns.pointplot(x='month', y='item_cnt_day', hue='year', data=cleaned)
# -
# ## Feature Selection
monthly_sales=sales.groupby(["date_block_num","shop_id","item_id"])[
"date_block_num","date","item_price","item_cnt_day"].agg({"date_block_num":'mean',"date":["min",'max'],"item_price":"mean","item_cnt_day":"sum"})
monthly_sales.head(5)
sales_data_flat = monthly_sales.item_cnt_day.apply(list).reset_index()
sales_data_flat = pd.merge(test,sales_data_flat,on = ['item_id','shop_id'],how = 'left')
sales_data_flat.fillna(0,inplace = True)
sales_data_flat.drop(['shop_id','item_id'],inplace = True, axis = 1)
sales_data_flat.head(20)
pivoted_sales = sales_data_flat.pivot_table(index='ID', columns='date_block_num',fill_value = 0,aggfunc='sum' )
pivoted_sales.head(20)
# ## Split train and test set
# +
X_train = pivoted_sales.values[:,:-1]
y_train = pivoted_sales.values[:,-1:]
X_test = pivoted_sales.values[:,1:]
print(X_train.shape,y_train.shape,X_test.shape)
# -
# ## Create XGBoost model
# +
from xgboost import XGBRegressor
from xgboost import plot_importance
model = XGBRegressor(
max_depth=10,
n_estimators=1000,
min_child_weight=0.5,
colsample_bytree=0.8,
subsample=1,
eta=0.3,
seed=42)
# -
model.fit(X_train, y_train, eval_metric="rmse", eval_set=[(X_train, y_train)], verbose=True, early_stopping_rounds = 10)
plot_importance(model)
# ## Predict Test
y_pred = model.predict(X_test)
y_pred = y_pred.clip(0,20)
y_pred.ravel()
# ## Create Submission
submission_df = pd.DataFrame({'ID': test['ID'], 'item_cnt_month': y_pred.ravel()})
submission_df.head()
submission_df['item_cnt_month'].max()
submission_df.to_csv('submission.csv', index = False)
| TimeSeriesUsingXGB.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.2
# language: julia
# name: julia-1.6
# ---
using Revise
using LinearAlgebra
using TransportBasedInference
import TransportBasedInference: optimize
using Statistics
using Test
using ForwardDiff
using QuadGK
using SpecialFunctions
using BenchmarkTools
using Optim
# +
Nx = 2
Ne = 100
X = Matrix([ 0.096932076386797 -0.062113573952594
1.504907056818036 -1.270761688825393
0.122588515870727 0.138604054973047
0.095040305032907 -0.097989669097228
0.056360334325948 -0.098106117790437
0.034757567319988 -0.152864122871058
0.260831995265119 -0.564465060524812
0.191690179163063 0.097123146831467
0.131835643270215 -0.303532750891873
0.121404994711150 -0.414223062306829
-0.046027217149733 0.267918560101165
0.222891154723706 0.054734106232808
0.264238681579693 -0.561044970267127
0.021345749974379 0.023020769850203
0.395423632217738 -0.619284581169702
0.231607199656522 0.319089487692097
0.389505867820041 0.572610562725449
-0.146845693861564 -0.007959126279965
0.138489687840940 0.330481872227252
1.617286792209003 -1.272765331253319
0.013078993247482 0.006243503157725
0.304266481947197 -0.508127858160603
-0.134224572142272 -0.057784550290700
0.058245459428341 -0.388163133932979
0.143773871017238 -0.569911233902741
-0.020852128497605 0.159888469685458
0.042924979621615 -0.285726823344180
0.566514904142701 -0.815010017214964
0.022782788493659 -0.468191560991914
-0.072005082647238 -0.139317736738881
-0.088861617714439 -0.042323531263036
0.146019714866274 -0.441835803838725
0.062827433058116 0.255887684932700
0.024887528661752 -0.036316627235643
0.079288422911736 0.009625876863572
0.124929172288805 -0.196711126223702
0.886085483369468 1.012837297288688
-0.062868708331948 0.065147939699519
0.117047537973873 -0.301180703268764
0.147117619773736 0.466610279392604
0.690523534483231 0.722717906410571
-0.110919157441807 0.000008657527041
2.055034349742512 1.423365847218695
0.091130503033388 -0.192933472923564
0.075510323340039 0.075087088602955
-0.181121769738736 -0.158594842114751
-0.080866378396288 -0.081393537160189
0.184561993178819 0.409693133491638
-0.032018126656783 0.040851821749292
0.702808256935611 0.978185567019995
-0.029041940950512 0.097268795862442
1.406479923500753 -1.166340130567743
-0.109758856235453 -0.297180107753266
0.236494055349260 0.548223372779986
-0.108550945852467 -0.097740533128956
0.908726493797006 -0.957752761353643
0.178092997560103 -0.359297653971968
1.103683368372646 -1.064482193617671
1.907065203740822 1.349542162065178
0.523722913693736 0.725695151794695
0.261131020580618 0.576363120484564
0.118044539009197 0.196304662470752
0.289261334786348 0.399639383890177
0.902906400981006 -0.957301599045371
-0.054657884786803 -0.292760625531884
-0.021735291600425 0.029650166664389
0.065200888050752 -0.295894159582647
1.486186253485923 -1.217814186300608
0.889545420155124 0.939789761164950
-0.174386606410644 -0.092037014472893
-0.065037226616579 0.009771974040525
0.074486430996400 -0.287910597788305
0.174336742307535 -0.400464726305446
0.096781997529499 -0.153178844921250
0.796408810115516 0.881930856163525
0.005874471676882 0.067750993468743
0.156654113730103 -0.239182272065197
0.333688106104332 -0.629954291766549
0.086388606764696 0.305488995071947
0.211268899950691 -0.299878322704640
0.104223240397571 0.199354790284364
0.336858958710283 -0.620166113933000
0.145071750152222 0.250136305618056
0.032242317206686 -0.233223578816564
0.064558616046395 -0.007577632839606
-0.055022872335109 -0.190212128003969
-0.169436992810515 -0.206948615170099
0.150088939478557 0.090560438773547
0.256256842094403 0.598874371523965
0.340882741111244 -0.516594535669757
0.278186325120137 -0.547968005821968
0.645979568807173 -0.827959899268083
0.436535833804569 0.689957746461832
0.268437499571141 0.341281325011944
0.120485843238972 -0.301999984465851
0.160365386980321 -0.202012022307596
0.154560496835611 0.244912011403144
0.117966782622546 0.342990099492354
0.280465408470057 -0.526206432878627
1.174002195932550 1.136790584581798]')
L = LinearTransform(X)
X = transform(L, X)
# -
m = 100
S = HermiteMap(m, X; diag = true, factor = 1.0, α = 0.0, b = "CstProHermiteBasis");
S = optimize(S, X, 15; maxterms = 15, withconstant = false, withqr = true, verbose = false,
maxpatience = 30, start = 1, hessprecond = true)
getidx(S[2])
getcoeff(S[2])
#
# https://math.stackexchange.com/questions/2753316/generating-a-random-tridiagonal-symmetric-positive-definite-matrix
function triposdef(N)
# The function generates a random tridiagonal symmetric
# positive definite N by N matrix;
b = randn(N-1);
a = [abs.(b); 0] + [0; abs.(b)] + abs.(randn(N));
return SymTridiagonal(a, b)
end
# +
Nx = 10
Ne = 5000
Σ = Matrix(triposdef(10))
L = cholesky(Σ)
πX = MvNormal(zeros(Nx), Σ)
# +
X = rand(πX, Ne)
# X = randn(Nx, Ne).^2 + randn(Nx, Ne) .+ randn(Nx)
# -
m = 30
S = HermiteMap(m, X; diag = true, α = 1.0e-6);
S = optimize(S, X, "kfolds"; withqr = true, verbose = true, hessprecond = true)
getidx(S[10])
plot(S)
stor = Storage(S[10].I.f, X);
q = QRscaling(stor)
# +
P = zeros(ncoeff(S[10]), ncoeff(S[10]))
precond!(P, getcoeff(S[10]), stor, S[10], X)
Pqr = zeros(ncoeff(S[10]), ncoeff(S[10]))
qrprecond!(Pqr, q.U*getcoeff(S[10]), q, stor, S[10], X)
# -
heatmap(abs.(Pqr), yflip = true, ratio = 1.0)
cond(P)
cond(Pqr)
F = evaluate(S, X)
Ωhat = zeros(Nx, Nx)
cache = zeros(1, Nx, Nx)
for i=1:Ne
hess_x_log_pdf!(cache, S, X[:,i:i])
Ωhat .+= copy(cache[1,:,:]).^2
end
rmul!(Ωhat, 1/Ne)
# +
plt = plot(size = (800, 800))
heatmap!(plt,log10.(Ωhat), ratio = 1, yflip = true,
colorbar = true, color = :plasma, clim = (1.5, Inf), colorbar_title = L"\log_{10}(\hat{\Omega})",
xlabel = "Index", ylabel = "Index", background_color_inside = palette(:plasma)[1],
yticks = (reverse(collect(0:10:Nx))))
plt
# +
plt = plot(size = (800, 800))
heatmap!(plt,log10.(abs.(Ωhat - Σ.^2)), ratio = 1, yflip = true,
colorbar = true, color = :plasma, clim = (1.5, Inf), colorbar_title = L"\log_{10}(\hat{\Omega})",
xlabel = "Index", ylabel = "Index", background_color_inside = palette(:plasma)[1],
yticks = (reverse(collect(0:10:Nx))))
plt
# -
| notebooks/Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install image_slicer
# from image_slicer import slice
# slice('ScanImage1.tif', 2048)
from image_slicer import slice
slice('Equalsize.tif', 1024)
| images/.ipynb_checkpoints/ImageSlicing-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Reducing WFM data
#
# This notebook aims to illustrate how to work with the wavelength frame multiplication submodule `wfm`.
#
# We will create a beamline that resembles the ODIN instrument beamline,
# generate some fake neutron data,
# and then show how to convert the neutron arrival times at the detector to neutron time-of-flight,
# from which a wavelength can then be computed (or process also commonly known as 'stitching').
import numpy as np
import matplotlib.pyplot as plt
plt.ioff() # Turn of auto-showing of figures
import scipp as sc
from scipp import constants
import scippneutron as scn
import ess.wfm as wfm
import ess.choppers as ch
np.random.seed(1) # Fixed for reproducibility
# ## Create beamline components
#
# We first create all the components necessary to a beamline to run in WFM mode
# (see [Introduction to WFM](introduction-to-wfm.ipynb) for the meanings of the different symbols).
# The beamline will contain
#
# - a neutron source, located at the origin ($x = y = z = 0$)
# - a pulse with a defined length ($2860 ~\mu s$) and $t_0$ ($130 ~\mu s$)
# - a single pixel detector, located at $z = 60$ m
# - two WFM choppers, located at $z = 6.775$ m and $z = 7.225$ m, each with 6 frame windows/openings
#
# The `wfm` module provides a helper function to quickly create such a beamline.
# It returns a `dict` of coordinates, that can then be subsequently added to a data container.
coords = wfm.make_fake_beamline(nframes=6)
coords
# ## Generate some fake data
#
# Next, we will generate some fake imaging data (no scattering will be considered),
# that is supposed to mimic a spectrum with a Bragg edge located at $4\unicode{x212B}$.
# We start with describing a function which will act as our underlying distribution
x = np.linspace(1, 10.0, 100000)
a = 20.0
b = 4.0
y1 = 0.7 / (np.exp(-a * (x - b)) + 1.0)
y2 = 1.4-0.2*x
y = y1 + y2
fig1, ax1 = plt.subplots()
ax1.plot(x, y)
ax1.set_xlabel("Wavelength [angstroms]")
fig1
# We then proceed to generate two sets of 1,000,000 events:
# - one for the `sample` using the distribution defined above
# - and one for the `vanadium` which will be just a flat random distribution
#
# For the events in both `sample` and `vanadium`,
# we define a wavelength for the neutrons as well as a birth time,
# which will be a random time between the pulse $t_0$ and the end of the useable pulse $t_0$ + pulse_length.
nevents = 1_000_000
events = {
"sample": {
"wavelengths": sc.array(
dims=["event"],
values=np.random.choice(x, size=nevents, p=y/np.sum(y)),
unit="angstrom"),
"birth_times": sc.array(
dims=["event"],
values=np.random.random(nevents) * coords["source_pulse_length"].value,
unit="us") + coords["source_pulse_t_0"]
},
"vanadium": {
"wavelengths": sc.array(
dims=["event"],
values=np.random.random(nevents) * 9.0 + 1.0,
unit="angstrom"),
"birth_times": sc.array(
dims=["event"],
values=np.random.random(nevents) * coords["source_pulse_length"].value,
unit="us") + coords["source_pulse_t_0"]
}
}
# We can then take a quick look at our fake data by histogramming the events
# Histogram and plot the event data
bins = np.linspace(1.0, 10.0, 129)
fig2, ax2 = plt.subplots()
for key in events:
h = ax2.hist(events[key]["wavelengths"].values, bins=128, alpha=0.5, label=key)
ax2.set_xlabel("Wavelength [angstroms]")
ax2.set_ylabel("Counts")
ax2.legend()
fig2
# We can also verify that the birth times fall within the expected range:
for key, item in events.items():
print(key)
print(sc.min(item["birth_times"]))
print(sc.max(item["birth_times"]))
# We can then compute the arrival times of the events at the detector pixel
# The ratio of neutron mass to the Planck constant
alpha = sc.to_unit(constants.m_n / constants.h, 'us/m/angstrom')
# The distance between the source and the detector
dz = sc.norm(coords['position'] - coords['source_position'])
for key, item in events.items():
item["arrival_times"] = alpha * dz * item["wavelengths"] + item["birth_times"]
events["sample"]["arrival_times"]
# ## Visualize the beamline's chopper cascade
#
# We first attach the beamline geometry to a Dataset
ds = sc.Dataset(coords=coords)
ds
# The `wfm.plot` submodule provides a useful tool to visualise the chopper cascade as a time-distance diagram.
# This is achieved by calling
wfm.plot.time_distance_diagram(ds)
# This shows the 6 frames, generated by the WFM choppers,
# as well as their predicted time boundaries at the position of the detector.
#
# Each frame has a time window during which neutrons are allowed to pass through,
# as well as minimum and maximum allowed wavelengths.
#
# This information is obtained from the beamline geometry by calling
frames = wfm.get_frames(ds)
frames
# ## Discard neutrons that do not make it through the chopper windows
#
# Once we have the parameters of the 6 wavelength frames,
# we need to run through all our generated neutrons and filter out all the neutrons with invalid flight paths,
# i.e. the ones that do not make it through both chopper openings in a given frame.
# +
for item in events.values():
item["valid_indices"] = []
near_wfm_chopper = ds.coords["chopper_wfm_1"].value
far_wfm_chopper = ds.coords["chopper_wfm_2"].value
near_time_open = ch.time_open(near_wfm_chopper)
near_time_close = ch.time_closed(near_wfm_chopper)
far_time_open = ch.time_open(far_wfm_chopper)
far_time_close = ch.time_closed(far_wfm_chopper)
for item in events.values():
# Compute event arrival times at wfm choppers 1 and 2
slopes = 1.0 / (alpha * item["wavelengths"])
intercepts = -slopes * item["birth_times"]
times_at_wfm1 = (sc.norm(near_wfm_chopper["position"].data) - intercepts) / slopes
times_at_wfm2 = (sc.norm(far_wfm_chopper["position"].data) - intercepts) / slopes
# Create a mask to see if neutrons go through one of the openings
mask = sc.zeros(dims=times_at_wfm1.dims, shape=times_at_wfm1.shape, dtype=bool)
for i in range(len(frames["time_min"])):
mask |= ((times_at_wfm1 >= near_time_open["frame", i]) &
(times_at_wfm1 <= near_time_close["frame", i]) &
(item["wavelengths"] >= frames["wavelength_min"]["frame", i]).data &
(item["wavelengths"] <= frames["wavelength_max"]["frame", i]).data)
item["valid_indices"] = np.ravel(np.where(mask.values))
# -
# ## Create a realistic Dataset
#
# We now create a dataset that contains:
# - the beamline geometry
# - the time coordinate
# - the histogrammed events
# +
for item in events.values():
item["valid_times"] = item["arrival_times"].values[item["valid_indices"]]
tmin = min([item["valid_times"].min() for item in events.values()])
tmax = max([item["valid_times"].max() for item in events.values()])
dt = 0.1 * (tmax - tmin)
time_coord = sc.linspace(dim='time',
start=tmin - dt,
stop=tmax + dt,
num=257,
unit=events["sample"]["arrival_times"].unit)
# Histogram the data
for key, item in events.items():
da = sc.DataArray(
data=sc.ones(dims=['time'], shape=[len(item["valid_times"])],
unit=sc.units.counts, with_variances=True),
coords={
'time': sc.array(dims=['time'], values=item["valid_times"], unit=sc.units.us)})
ds[key] = sc.histogram(da, bins=time_coord)
ds
# -
ds.plot()
# ## Stitch the frames
#
# Wave-frame multiplication consists of making 6 new pulses from the original pulse.
# This implies that the WFM choppers are acting as a source chopper.
# Hence, to compute a wavelength from a time and a distance between source and detector,
# the location of the source must now be at the position of the WFM choppers,
# or more exactly at the mid-point between the two WFM choppers.
#
# The stitching operation equates to converting the `time` dimension to `time-of-flight`,
# by subtracting from each frame a time shift equal to the mid-point between the two WFM choppers.
#
# This is performed with the `stitch` function in the `wfm` module:
stitched = wfm.stitch(frames=frames,
data=ds,
dim='time',
bins=257)
stitched
stitched.plot()
# For diagnostic purposes,
# it can be useful to visualize the individual frames before and after the stitching process.
# The `wfm.plot` module provides two helper functions to do just this:
wfm.plot.frames_before_stitching(data=ds['sample'], frames=frames, dim='time')
wfm.plot.frames_after_stitching(data=ds['sample'], frames=frames, dim='time')
# ## Convert to wavelength
#
# Now that the data coordinate is time-of-flight (`tof`),
# we can use `scippneutron` to perform the unit conversion from `tof` to `wavelength`.
from scippneutron.tof.conversions import beamline, elastic
graph = {**beamline(scatter=False), **elastic("tof")}
converted = stitched.transform_coords("wavelength", graph=graph)
converted.plot()
# ## Normalization
#
# Normalization is performed simply by dividing the counts of the `sample` run by the counts of the `vanadium` run.
normalized = converted['sample'] / converted['vanadium']
normalized.plot()
# ## Comparing to the raw wavelengths
#
# The final step is a sanity check to verify that the wavelength-dependent data obtained from the stitching process
# agrees (to within the beamline resolution) with the original wavelength distribution that was generated at
# the start of the workflow.
#
# For this, we simply histogram the raw neutron events using the same bins as the `normalized` data,
# filtering out the neutrons with invalid flight paths.
for item in events.values():
item["wavelength_counts"], _ = np.histogram(
item["wavelengths"].values[item["valid_indices"]],
bins=normalized.coords['wavelength'].values)
# We then normalize the `sample` by the `vanadium` run,
# and plot the resulting spectrum alongside the one obtained from the stitching.
# +
original = sc.DataArray(
data=sc.array(dims=['wavelength'],
values=events["sample"]["wavelength_counts"] /
events["vanadium"]["wavelength_counts"]),
coords = {"wavelength": normalized.coords['wavelength']})
sc.plot({"stitched": normalized, "original": original})
# -
# We can see that the counts in the `stitched` data agree very well with the original data.
# There is some smoothing of the data seen in the `stitched` result,
# and this is expected because of the resolution limitations of the beamline due to its long source pulse.
# This smoothing (or smearing) would, however, be much stronger if WFM choppers were not used.
#
# ## Without WFM choppers
#
# In this section, we compare the results obtained above to a beamline that does not have a WFM chopper system.
# We make a new set of events,
# where the number of events is equal to the number of neutrons that make it through the chopper cascade in the previous case.
nevents_no_wfm = len(events["sample"]["valid_times"])
events_no_wfm = {
"sample": {
"wavelengths": sc.array(
dims=["event"],
values=np.random.choice(x, size=nevents_no_wfm, p=y/np.sum(y)),
unit="angstrom"),
"birth_times": sc.array(
dims=["event"],
values=np.random.random(nevents_no_wfm) * coords["source_pulse_length"].value,
unit="us") + coords["source_pulse_t_0"]
},
"vanadium": {
"wavelengths": sc.array(
dims=["event"],
values=np.random.random(nevents_no_wfm) * 9.0 + 1.0,
unit="angstrom"),
"birth_times": sc.array(
dims=["event"],
values=np.random.random(nevents_no_wfm) * coords["source_pulse_length"].value,
unit="us") + coords["source_pulse_t_0"]
}
}
for key, item in events_no_wfm.items():
item["arrival_times"] = alpha * dz * item["wavelengths"] + item["birth_times"]
events_no_wfm["sample"]["arrival_times"]
# We then histogram these events to create a new Dataset.
# Because we are no longer make new pulses with the WFM choppers,
# the event time-of-flight is simply the arrival time of the event at the detector.
# +
tmin = min([item["arrival_times"].values.min() for item in events_no_wfm.values()])
tmax = max([item["arrival_times"].values.max() for item in events_no_wfm.values()])
dt = 0.1 * (tmax - tmin)
time_coord_no_wfm = sc.linspace(dim='tof',
start=tmin - dt,
stop=tmax + dt,
num=257,
unit=events_no_wfm["sample"]["arrival_times"].unit)
ds_no_wfm = sc.Dataset(coords=coords)
# Histogram the data
for key, item in events_no_wfm.items():
da = sc.DataArray(
data=sc.ones(dims=['tof'], shape=[len(item["arrival_times"])],
unit=sc.units.counts, with_variances=True),
coords={
'tof': sc.array(dims=['tof'], values=item["arrival_times"].values, unit=sc.units.us)})
ds_no_wfm[key] = sc.histogram(da, bins=time_coord_no_wfm)
ds_no_wfm
# -
sc.plot(ds_no_wfm)
# We then perform the standard unit conversion and normalization
converted_no_wfm = ds_no_wfm.transform_coords("wavelength", graph=graph)
normalized_no_wfm = converted_no_wfm['sample'] / converted_no_wfm['vanadium']
normalized_no_wfm.plot()
# In the same manner and in the previous section, we compare to the real neutron wavelengths
for item in events_no_wfm.values():
item["wavelength_counts"], _ = np.histogram(
item["wavelengths"].values,
bins=normalized_no_wfm.coords['wavelength'].values)
# +
original_no_wfm = sc.DataArray(
data=sc.array(dims=['wavelength'],
values=events_no_wfm["sample"]["wavelength_counts"] /
events_no_wfm["vanadium"]["wavelength_counts"]),
coords = {"wavelength": normalized_no_wfm.coords['wavelength']})
w_min = 2.0 * sc.units.angstrom
w_max = 5.5 * sc.units.angstrom
sc.plot({"without WFM": normalized_no_wfm['wavelength', w_min:w_max],
"original": original_no_wfm['wavelength', w_min:w_max]},
errorbars=False)
# -
# We can see that there is a significant shift between the calculated wavelength of the Bragg edge around $4\unicode{x212B}$
# and the original underlying wavelengths.
# In comparison, the same plot for the WFM run yields a much better agreement
sc.plot({"stitched": normalized['wavelength', w_min:w_max],
"original": original['wavelength', w_min:w_max]},
errorbars=False)
# ## Working in event mode
#
# It is also possible to work with WFM data in event mode.
# The `stitch` utility will accept both histogrammed and binned (event) data.
#
# We first create a new dataset, with the same events as in the first example,
# but this time we bin the data with `sc.bin` instead of using `sc.histogram`,
# so we can retain the raw events.
# +
for item in events.values():
item["valid_times"] = item["arrival_times"].values[item["valid_indices"]]
tmin = min([item["valid_times"].min() for item in events.values()])
tmax = max([item["valid_times"].max() for item in events.values()])
dt = 0.1 * (tmax - tmin)
time_coord = sc.linspace(dim='time',
start=tmin - dt,
stop=tmax + dt,
num=257,
unit=events["sample"]["arrival_times"].unit)
ds_event = sc.Dataset(coords=coords)
# Bin the data
for key, item in events.items():
da = sc.DataArray(
data=sc.ones(dims=['event'], shape=[len(item["valid_times"])], unit=sc.units.counts,
with_variances=True),
coords={
'time': sc.array(dims=['event'], values=item["valid_times"], unit=sc.units.us)})
ds_event[key] = sc.bin(da, edges=[time_coord])
ds_event
# -
# The underlying events can be inspected by using the `.bins.constituents['data']` property of our objects:
ds_event["sample"].bins.constituents['data']
# We can visualize this to make sure it looks the same as the histogrammed case above:
sc.plot(ds_event.bins.sum())
# As explained above, the `stitch` routine accepts both histogrammed and binned (event) data.
# So stitching the binned data works in the exact same way as above, namely
stitched_event = wfm.stitch(frames=frames,
data=ds_event,
dim='time')
stitched_event
# The `stitch` function will return a data structure with a single bin in the `'tof'` dimension.
# Visualizing this data is therefore slightly more tricky,
# because the data needs to be histogrammed using a finer binning before a useful plot can be made.
sc.plot(sc.histogram(stitched_event,
bins=sc.linspace(
dim='tof',
start=stitched_event.coords['tof']['tof', 0].value,
stop=stitched_event.coords['tof']['tof', -1].value,
num=257,
unit=stitched_event.coords['tof'].unit)))
# At this point, it may be useful to compare the results of the two different stitching operations.
rebinned = sc.bin(stitched_event["sample"], edges=[stitched["sample"].coords['tof']])
sc.plot({"events": rebinned.bins.sum(), "histogram": stitched["sample"]}, errorbars=False)
# We note that histogramming the data early introduces some smoothing in the data.
#
# We can of course continue in event mode and perform the unit conversion and normalization to the Vanadium.
converted_event = stitched_event.transform_coords("wavelength", graph=graph)
# Normalizing binned data is done using the sc.lookup helper
hist = sc.histogram(
converted_event["vanadium"], bins=converted["sample"].coords['wavelength'])
normalized_event = converted_event["sample"].bins / sc.lookup(func=hist, dim='wavelength')
# Finally, we compare the end result with the original wavelengths, and see that the agreement is once again good.
to_plot = sc.bin(normalized_event,
edges=[converted["sample"].coords['wavelength']]).bins.sum()
sc.plot({"stitched_event": to_plot, "original": original})
# We can also compare directly to the histogrammed version,
# to see that both methods remain in agreement to a high degree.
sc.plot({"stitched": normalized['wavelength', w_min:w_max],
"original": original['wavelength', w_min:w_max],
"stitched_event": to_plot['wavelength', w_min:w_max]})
| docs/techniques/wfm/reducing-wfm-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
from collections import deque
from functools import partial
from itertools import cycle
import pandas as pd
import datashader as ds
import datashader.transfer_functions as tf
from datashader.colors import viridis
from streamz import Stream
# -
def taxi_trips_stream(source='data/nyc_taxi.csv', frequency='T'):
"""Generate dataframes grouped by given frequency"""
def get_group(resampler, key):
try:
df = resampler.get_group(key)
df.reset_index(drop=True)
except KeyError:
df = pd.DataFrame()
return df
df = pd.read_csv(source,
infer_datetime_format=True,
parse_dates=['tpep_pickup_datetime', 'tpep_pickup_datetime'])
df = df.set_index('tpep_pickup_datetime', drop=True)
df = df.sort_index()
r = df.resample(frequency)
chunks = [get_group(r, g) for g in sorted(r.groups)]
indices = cycle(range(len(chunks)))
while True:
yield chunks[next(indices)]
# ### Create streams
# Given a stream of dataframes representing NYC taxi data, we create four streams: two streams are sliding window aggregations over some time period, while two other streams track the cumulative average for a particular value. The pipeline visualization below shows each step that makes up each stream.
#
# For each aggregation stream, the steps are 1) aggregate each dataframe using a Datashader reduction, 2) keep sliding window of those aggregations, and 3) combine sliding window collection into image. The first stream creates a two-day sliding window aggregation, while the second stream creates a 1-week sliding window aggregation.
#
# For each cumulative average stream, we track the cumulative sum of each value along with the number of cumulative data points.
#
# We use the primitives given in the `streamz` library to accomplish this. `aggregated_sliding_window_image_queue` creates each aggregation stream. `cumulative_mean_queue` creates each cumulative average stream, but this will likely be replaced by a native `streamz.StreamingDataFrame` container when ready. Each stream will place its final result into a double-ended queue, which is used to keep a history of previous results. By default, we only keep the most recent.
def aggregate_df(df, cvs, x, y, agg=None):
return df.index.min(), df.index.max(), cvs.points(df, x, y, agg)
def aggregate_images(iterable, cmap):
name = "{:.10} - {:.10}".format(str(iterable[0][0]), str(iterable[-1][1]))
total = sum([item[2] for item in iterable])
return tf.shade(total, cmap=cmap, name=name)
def aggregated_sliding_window_image_queue(source, agg1, agg2, window=1, history=1):
q = deque(maxlen=history)
s = source.map(agg1).sliding_window(window)
s.map(agg2).sink(q.append)
return q
def cumulative_mean_queue(source, column, history=1):
def accumulator(acc, df):
n, total, oldest = acc
if not oldest:
oldest = df.index.min()
return n + 1, total + df[column].sum(), oldest, df.index.max()
def merge(value):
n, total, oldest, latest = value
return oldest, latest, total / n
q = deque(maxlen=history)
source.accumulate(accumulator, start=(0, 0, None)).map(merge).sink(q.append)
return q
def show_queue(q, column):
pd.options.display.float_format = '{:.2f}'.format
return pd.DataFrame(list(q), columns=['start', 'end', column])
x_range = (-8243204.0, -8226511.0)
y_range = (4968192.0, 4982886.0)
cvs = ds.Canvas(plot_width=800, plot_height=600, x_range=x_range, y_range=y_range)
# +
# Helper functions for useful aggregations
min_amount = partial(aggregate_df, cvs, x='pickup_x', y='pickup_y', agg=ds.min('total_amount'))
max_amount = partial(aggregate_df, cvs, x='pickup_x', y='pickup_y', agg=ds.max('total_amount'))
mean_amount = partial(aggregate_df, cvs, x='pickup_x', y='pickup_y', agg=ds.mean('total_amount'))
sum_amount = partial(aggregate_df, cvs, x='pickup_x', y='pickup_y', agg=ds.sum('total_amount'))
max_passengers = partial(aggregate_df, cvs, x='pickup_x', y='pickup_y', agg=ds.max('passenger_count'))
sum_passengers = partial(aggregate_df, cvs, x='pickup_x', y='pickup_y', agg=ds.sum('passenger_count'))
sum_pickups = partial(aggregate_df, cvs, x='pickup_x', y='pickup_y', agg=ds.count())
reduce_viridis = partial(aggregate_images, cmap=viridis)
# +
source = Stream()
q_days = aggregated_sliding_window_image_queue(source, window=2, history=6, agg1=max_amount, agg2=reduce_viridis)
q_week = aggregated_sliding_window_image_queue(source, window=7, agg1=max_amount, agg2=reduce_viridis)
q_avg_passengers = cumulative_mean_queue(source, 'passenger_count', history=7)
q_avg_amount = cumulative_mean_queue(source, 'total_amount', history=7)
# -
source.visualize()
# ### Simplifying stream creation
# As you can see in the previous section, there are a few areas to improve upon:
#
# - less code/boilerplate
# - hide individual steps seen in stream diagram
# - encapsulate separate stream construction methods into helper classes
# - separate stream creation and stream sink
# - allow for partial results from sliding windows (not currently supported by `streamz`)
# - output results into other collections besides queues
#
# By subclassing `streamz.Stream`, we've accomplished the above without sacrificing readability.
class SlidingWindowImageAggregate(Stream):
def __init__(self, source, canvas, x, y, agg, n=7, cmap=None, bgcolor='black'):
# Set internal streamz instance variables to control names in diagram
self.n = n
def aggregate_df(df):
return df.index.min(), df.index.max(), canvas.points(df, x, y, agg)
def aggregate_images(iterable):
name = "{:.10} - {:.10}".format(str(iterable[0][0]), str(iterable[-1][1]))
total = sum([item[2] for item in iterable])
return tf.set_background(tf.shade(total, cmap, name=name), color=bgcolor)
self.cache = deque(maxlen=n)
self.agg1 = aggregate_df
self.agg2 = aggregate_images
Stream.__init__(self, source)
def update(self, x, who=None):
self.cache.append(self.agg1(x))
return self.emit(self.agg2(tuple(self.cache)))
class CumulativeMean(Stream):
def __init__(self, source, column):
# Set internal streamz instance variables to control names in diagram
self.str_list = ['column']
self.column = column
self.count = 0
self.total = 0
self.oldest = None
Stream.__init__(self, source)
def update(self, x, who=None):
if not self.oldest:
self.oldest = x.index.min()
self.count, self.total = self.count + 1, self.total + x[self.column].sum()
return self.emit((self.oldest, x.index.max(), self.total / self.count))
# +
source = Stream()
cvs = ds.Canvas(plot_width=800, plot_height=600, x_range=x_range, y_range=y_range)
q_days = deque(maxlen=6)
s_days = SlidingWindowImageAggregate(source, cvs, 'pickup_x', 'pickup_y', ds.max('total_amount'), n=2, cmap=viridis)
s_days.sink(q_days.append)
q_week = deque(maxlen=1)
s_week = SlidingWindowImageAggregate(source, cvs, 'pickup_x', 'pickup_y', ds.max('total_amount'), n=7, cmap=viridis)
s_week.sink(q_week.append)
q_avg_passengers = deque(maxlen=7)
s_avg_passengers = CumulativeMean(source, 'passenger_count')
s_avg_passengers.sink(q_avg_passengers.append)
q_avg_amount = deque(maxlen=7)
s_avg_amount = CumulativeMean(source, 'total_amount')
s_avg_amount.sink(q_avg_amount.append)
# -
source.visualize()
# ### Push data through streams
# We initially push 3 days worth of dataframes through the streams to view partial results.
trips_per_day = taxi_trips_stream(frequency='D')
for i in range(3):
source.emit(next(trips_per_day))
tf.Images(*list(q_week))
for i in range(4):
source.emit(next(trips_per_day))
tf.Images(*list(q_week))
# #### Cumulative average of passengers (ordered by oldest first)
show_queue(q_avg_passengers, 'cumulative average passengers')
# #### Cumulative average of total fare (ordered by oldest first)
show_queue(q_avg_amount, 'cumulative average total fare')
# #### History of 2-day aggregations (ordered by oldest first)
tf.Images(*list(q_days))
# #### Current 1-week aggregation
tf.Images(*list(q_week))
# Now we get the next day's worth of data and see how the streams have updated.
source.emit(next(trips_per_day))
# #### Cumulative average of passengers (ordered by oldest first)
show_queue(q_avg_passengers, 'cumulative average passengers')
# #### Cumulative average of total fare (ordered by oldest first)
show_queue(q_avg_amount, 'cumulative average total fare')
# #### History of 2-day aggregations (ordered by oldest first)
tf.Images(*list(q_days))
# #### Current 1-week aggregation
tf.Images(*list(q_week))
| datashader-work/datashader-examples/streaming-aggregation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import itk
import matplotlib.pyplot as plt
import icon_registration.test_utils
import numpy as np
logo_path = str(icon_registration.test_utils.TEST_DATA_DIR / "itkLogo.jpg")
logo_rgb = itk.imread(logo_path)
logo = itk.image_from_array(itk.array_from_image(logo_rgb)[:, :, 0])
logo.SetSpacing(logo_rgb.GetSpacing())
logo.SetDirection(itk.matrix_from_array(np.sqrt(.5) * np.array([[0, -1], [1, 1]])))
logo_no_spacing = itk.image_from_array(itk.array_from_image(logo))
def itk_show(im):
print(im.GetSpacing())
print(itk.template(im))
plt.imshow(itk.array_from_image(im))
plt.show()
itk_show(logo)
itk_show(logo_no_spacing)
from icon_registration.itk_wrapper import resampling_transform
transform = resampling_transform(logo, [200, 100])
resampler = itk.LinearInterpolateImageFunction.New(logo)
out = itk.resample_image_filter(
logo,
transform=transform,
interpolator = resampler,
size=[200, 100],
)
itk_show(out)
invTrans = transform.GetInverseTransform()
resampler = itk.LinearInterpolateImageFunction.New(out)
out2 = itk.resample_image_filter(
out,
transform=invTrans,
interpolator=resampler,
size=itk.size(logo),
output_spacing=itk.spacing(logo),
output_direction=logo.GetDirection())
itk_show(out2)
itk.template(logo)[0][itk.template(logo)[1]]
logo.GetLargestPossibleRegion().GetSize()[0]
ct = itk.CompositeTransform[itk.D, 3].New()
ct.PrependTransform()
itk.CenteredTransformInitializer.GetTypes()
t3 = itk.VersorRigid3DTransform[itk.D].New()
t3.GetVersor()
t3.GetInverseTransform().SetMatrix
| notebooks/ITK registration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import seaborn as sns
import folium
import pandas as pd
import geopandas as gpd
import geopandas
import geoplot
data=pd.read_csv('/Users/jeffreyng/Downloads/Data/Housing_Loss_Data/orange_fl_processed_2017_to_2019_20210916.csv')
result=data.set_index('census_tract_GEOID')
data.columns
map_data=result.reset_index()[['census_tract_GEOID', 'housing-loss-index']]
data1=pd.read_csv('/Users/jeffreyng/Downloads/Data/American_Community_Survey_Data/orange_acs5-2019_census.csv')
df = gpd.read_file('/Users/jeffreyng/Downloads/Data/Housing_Loss_Data/orange_fl_2010_tracts_formatted.geojson')
df
df.geometry
df=df.geometry
import geoplot
import geoplot.crs as gcrs
map_data=map_data.reset_index()
df=df.reset_index()
import geoplot as gplt
import matplotlib.pyplot as plt
map_data.census_tract_GEOID = map_data.census_tract_GEOID.astype(str).astype(int)
df.census_tract_GEOID= df.census_tract_GEOID.astype(str).astype(int)
fig, ax = plt.subplots(1, 1, figsize=(16, 12))
fullData = map_data.merge(df, left_on=['census_tract_GEOID'], right_on=['census_tract_GEOID'])
fullData.head(2)
# Set up the color sheme:
import mapclassify as mc
scheme = mc.Quantiles(fullData['housing-loss-index'], k=10)
gdf = geopandas.GeoDataFrame(fullData, crs="EPSG:4326")
# florida= geopandas.GeoDataFrame('/Users/jeffreyng/Downloads/Data_04/US_STATE.shp')
florida=gpd.read_file('/Users/jeffreyng/Downloads/Data_04/US_STATE.shp')
florida.geometry
orange=gpd.read_file('/Users/jeffreyng/Downloads/Data_04/US_COUNTY.shp')
gdf= geopandas.GeoDataFrame(orange)
orange1=gdf[gdf.STATE=='Florida']
gdf=geopandas.GeoDataFrame(orange1)
gdf=gdf.geometry
sys=geopandas.GeoDataFrame(orange)
sys1= sys[sys.COUNTY=='Orange County']
sys1.loc[2617]
gdf.drop(2617)
# +
counties=gpd.GeoSeries.plot(gdf.drop(2617), alpha=.7)
# +
# Map
# Initialize the figure
import matplotlib.pyplot as plt
fig, ax = plt.subplots(1, 1, figsize=(16, 12))
# Set up the color sheme:
import mapclassify as mc
scheme = mc.Quantiles(gdf['housing-loss-index'], k=10)
gplt.choropleth(gdf,
hue="housing-loss-index",
linewidth=.1,
scheme=scheme, cmap='inferno_r',
legend=True,
edgecolor='black',
ax=ax);
ax.set_title('Housing Loss Index in Orange County, FL', fontsize=13);
# -
data1.GEOID
data1['index'][:][0]
results=data1.set_index('GEOID')
result
result['median-household-income', 'single-parent-household', 'english-fluency', 'housing-loss-index', 'level-of-education',
]
total.columns[0:40]
# +
# -
result=result.reset_index('census_tract_GEOID')
results=results.reset_index('GEOID')
total= pd.merge(result, results, left_index=True, right_index=True)
total.set_index('census_tract_GEOID')
df
income= total[['census_tract_GEOID', 'median-household-income']]
# total=total.reset_index()
# df= df.reset_index()
income.census_tract_GEOID = income.census_tract_GEOID.astype(str).astype(int)
df.census_tract_GEOID = df.census_tract_GEOID.astype(str).astype(int)
income_map= pd.merge(df, income, on='census_tract_GEOID', how='inner',)
income_map
fullData.drop('index', inplace=True, axis=1)
fullData.dropna(inplace=True)
fullData
# +
fig, ax = plt.subplots(1, 1, figsize=(16, 12))
housing_loss=geopandas.GeoDataFrame(fullData)
scheme = mc.Quantiles(housing_loss['housing-loss-index'], k=10)
gplt.choropleth(housing_loss,
hue="housing-loss-index",
linewidth=.1,
scheme=scheme, cmap='inferno_r',
legend=True,
edgecolor='black',
ax=ax)
ax.set_title('Housing Loss Index in Orange County, FL', fontsize=13);
# +
fig, ax = plt.subplots(1, 1, figsize=(16, 12))
sys=geopandas.GeoDataFrame(income_map)
import matplotlib.pyplot as plt
# Set up the color sheme:
import mapclassify as mc
scheme = mc.Quantiles(sys['median-household-income'], k=10)
gplt.choropleth(sys,
hue="median-household-income",
linewidth=.1,
scheme=scheme, cmap='viridis_r',
legend=True,
edgecolor='black',
ax=ax);
ax.set_title('Median Household Income in Orange County, FL', fontsize=13)
# -
results.rename(mapper=dict_mapper, inplace=True)
data2=pd.read_csv('/Users/jeffreyng/Downloads/Data/American_Community_Survey_Data/acs5_variable_dict_2014_2019.csv')
list_vc=data2.variable_code
list_label=data2.label
dict_mapper=dict(zip(list_vc, list_label))
dict_mapper
df
results.columns
unemployed=results[['GEOID','DP03_0009PE']].rename(columns={'GEOID': 'census_tract_GEOID'})
unemployed.census_tract_GEOID = unemployed.census_tract_GEOID.astype(str).astype(int)
df.census_tract_GEOID = df.census_tract_GEOID.astype(str).astype(int)
unemployed_map= pd.merge(df, unemployed, on='census_tract_GEOID', how='inner',)
unemployed_map
# +
fig, ax = plt.subplots(1, 1, figsize=(16, 12))
sys=geopandas.GeoDataFrame(unemployed_map)
scheme = mc.Quantiles(sys['DP03_0009PE'], k=8)
gplt.choropleth(sys,
hue="DP03_0009PE",
linewidth=.1,
scheme=scheme, cmap='plasma_r',
legend=True,
edgecolor='black',
ax=ax);
ax.set_title('Unemployment Rates in Orange County, FL', fontsize=13)
# -
| code/jeffng/DataKind.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Load GLEIF Data from public repository on AWS (s3://gleif/)
# Copyright (C) 2021 OS-Climate
#
# Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
# +
# pip install boto3
# pip install python-dotenv
# pip install trino
# pip install pandas
# pip install anytree
# pip install sqlalchemy
# pip install sqlalchemy-trino
# pip install osc-ingest-tools
# -
from dotenv import dotenv_values, load_dotenv
import os
import pathlib
import boto3
from botocore import UNSIGNED
from botocore.client import Config
import pandas as pd
import trino
import io
import zipfile
import anytree
import osc_ingest_trino
# Use unsigned access to public S3 resource for GLEIF bucket
s3_resource = boto3.resource(
service_name="s3",
config=Config(signature_version=UNSIGNED),
)
bucket = s3_resource.Bucket("gleif")
# Retrieve latest LEI data set
LEI_data_sets = []
for obj in bucket.objects.filter(Prefix="data/csv/rr/"):
LEI_data_sets.append(obj.key)
print(f"Fetched data sets: {len(LEI_data_sets)}")
current_rr_data_set = LEI_data_sets[len(LEI_data_sets) - 1]
date_str_data, time_str_data = tuple(current_rr_data_set.split("/")[-1].split("-")[:2])
# +
from botocore import UNSIGNED
from botocore.config import Config
client = boto3.client(
"s3", config=Config(signature_version=UNSIGNED)
) # low-level functional API
obj = client.get_object(Bucket="gleif", Key=current_rr_data_set)
list_hierarchy_relevant_columns = [
"Relationship.RelationshipType",
"Relationship.StartNode.NodeID",
"Relationship.EndNode.NodeID",
"Relationship.RelationshipStatus",
]
pd_rr_data_set = pd.read_csv(
obj["Body"], usecols=list_hierarchy_relevant_columns, compression="gzip"
).convert_dtypes()
# -
# # Definition of necessary data processing functions
# +
def _create_issuer_lei_mapping(
pd_input: pd.DataFrame, relationship_type: str, end_node_key: str
) -> pd.DataFrame:
df_mask = (pd_input["Relationship.RelationshipStatus"] == "ACTIVE") & (
pd_input["Relationship.RelationshipType"] == relationship_type
)
dict_key_renaming = {
"Relationship.StartNode.NodeID": "DIRECT_ISSUER_LEI",
"Relationship.EndNode.NodeID": end_node_key,
}
return pd_input.rename(columns=dict_key_renaming)[dict_key_renaming.values()][
df_mask
]
def create_ultimate_issuer_lei_mapping(pd_input: pd.DataFrame) -> pd.DataFrame:
return _create_issuer_lei_mapping(
pd_input,
relationship_type="IS_ULTIMATELY_CONSOLIDATED_BY",
end_node_key="ULTIMATE_PARENT_ISSUER_LEI",
)
def create_direct_parent_issuer_lei_mapping(pd_input: pd.DataFrame) -> pd.DataFrame:
return _create_issuer_lei_mapping(
pd_input,
relationship_type="IS_DIRECTLY_CONSOLIDATED_BY",
end_node_key="DIRECT_PARENT_ISSUER_LEI",
)
def add_hierarchy_level_to_nodes(
list_unassigned_relationships: list, hierarchy_level: int, dict_nodes: dict
) -> bool:
"""
Recursive function, which matches the most recent version of the tree with a list of currently
unassigned directly consolidated relationships.
Loops through the hierarchy levels, takes the most recent version of the tree and checks if in the tree there are potential parents
of directly consolidated entities in the current list of unassigned directly consolidated
-> If a parent is found, the relationship is appended to the tree and popped from the list of unassigned relationships
"""
list_remaining_unassigned_relationships = []
amount_initially_unassigned_entities = len(list_unassigned_relationships)
print(f"Hierarchy_level {hierarchy_level}")
for unassigned_relationship_tuple in list_unassigned_relationships:
# Check if there direct parent issuer is already allocated in the tree structure,
# otherwise try again in the next call of the function
if (
unassigned_relationship_tuple[1] in dict_nodes.keys()
and unassigned_relationship_tuple[0] != unassigned_relationship_tuple[1]
):
parent_node = dict_nodes.get(unassigned_relationship_tuple[1])
if (
unassigned_relationship_tuple[0] in dict_nodes.keys()
): # LEI was already assigned
try:
# Overwrite the previous parent attribute in the node with a new one
dict_nodes.get(
unassigned_relationship_tuple[0]
).parent = parent_node
except:
print(
f"Unable to change parent for {unassigned_relationship_tuple[0]}"
)
list_remaining_unassigned_relationships.append(
unassigned_relationship_tuple
)
else:
# assign new parent child relationship in the tree structure
dict_nodes[unassigned_relationship_tuple[0]] = anytree.Node(
unassigned_relationship_tuple[0], parent=parent_node
)
else:
list_remaining_unassigned_relationships.append(
unassigned_relationship_tuple
)
# Terminate iff there was no mutation in the tree structure since the last function call,
# call function recursively otherwise
if amount_initially_unassigned_entities == len(
list_remaining_unassigned_relationships
):
print(
f"Remaining {len(list_remaining_unassigned_relationships)} could not be assigned to the tree"
)
return True
else:
add_hierarchy_level_to_nodes(
list_unassigned_relationships=list_remaining_unassigned_relationships,
hierarchy_level=hierarchy_level + 1,
dict_nodes=dict_nodes,
)
def create_company_hierarchy_tree_structure(pd_input: pd.DataFrame):
list_tuple_ultimate_issuer_relationship = create_ultimate_issuer_lei_mapping(
pd_rr_data_set
).to_records(index=False)
list_tuple_direct_parent_issuer_relationship = (
create_direct_parent_issuer_lei_mapping(pd_rr_data_set).to_records(index=False)
)
list_direct_issuer_LEI, list_ultimate_issuer_LEI = zip(
*list_tuple_ultimate_issuer_relationship
)
root = anytree.Node("root")
tmp_dict_nodes = (
{}
) # this dict is used as a performant hashmap lookup table to the nodes to avoid
# using the native search function of anytree which loops through the entire tree
for ultimate_issuer_lei in set(list_ultimate_issuer_LEI):
tmp_dict_nodes[ultimate_issuer_lei] = anytree.Node(
ultimate_issuer_lei, parent=root
)
# Some relationships in the GLEIF data are exclusively entered as an ultimate issuer relationship and not also
# as a directly consolidated relationship. To build a complete dataset and also include those we take the following approach:
# initially we assume all ultimate issuer relationships also as directly consolidated relationships. In a subsequent
# process step this assumption is checked:
# we overwrite the initially assumed direct relationship to the ultimate issuer with the more specific data from
# the direct issuer relationship dataset:
# - Case1: There is no entry in the directly consolidated issuer relationship,
# meaning the assumed relationship was correct
# - Case2: There is an entry in the directly consolidated issuer relationship: We delete the initially assumed
# entry and replace it with the more specific one
for relationship_tuple in list_tuple_ultimate_issuer_relationship:
parent_node = tmp_dict_nodes.get(relationship_tuple[1])
tmp_dict_nodes[relationship_tuple[0]] = anytree.Node(
relationship_tuple[0], parent=parent_node
)
add_hierarchy_level_to_nodes(
list_unassigned_relationships=list_tuple_direct_parent_issuer_relationship,
hierarchy_level=1,
dict_nodes=tmp_dict_nodes,
)
return root
# +
def rename_recursive_key_dicts(
dict_to_rename: dict, dict_of_key_old_to_new, new_name_dict_holder_key: str
) -> None:
for old_key, new_key in dict_of_key_old_to_new.items():
if old_key in dict_to_rename.keys():
dict_to_rename[new_key] = dict_to_rename.pop(old_key)
# after renaming
if new_name_dict_holder_key in dict_to_rename.keys():
list_dict_holder = dict_to_rename.get(new_name_dict_holder_key)
for held_dict in list_dict_holder:
rename_recursive_key_dicts(
dict_to_rename=held_dict,
dict_of_key_old_to_new=dict_of_key_old_to_new,
new_name_dict_holder_key=new_name_dict_holder_key,
)
def add_recursive_key_to_dict(
dict_to_add_to: dict, key_to_add: str,
value_to_add: int, name_dict_holder_key: str
):
"""
Increments the value of key_to_add by one
"""
dict_to_add_to[key_to_add] = value_to_add
if name_dict_holder_key in dict_to_add_to.keys():
list_dict_holder = dict_to_add_to.get(name_dict_holder_key)
for held_dict in list_dict_holder:
add_recursive_key_to_dict(
dict_to_add_to=held_dict,
key_to_add=key_to_add,
value_to_add=value_to_add + 1,
name_dict_holder_key=name_dict_holder_key,
)
# -
# # Creating data set for ultimate issuer LEI mapping
# +
df_ultimate_issuer_relationship_LEI = create_ultimate_issuer_lei_mapping(pd_rr_data_set)
str_timestamp = f'{date_str_data[:4]}-{date_str_data[4:6]}-{date_str_data[6:]}T{time_str_data[:2]}'
data_time_stamp = pd.Timestamp(str_timestamp, tz='UTC')
df_ultimate_issuer_relationship_LEI["time_stamp"] = data_time_stamp
df_ultimate_issuer_relationship_LEI = df_ultimate_issuer_relationship_LEI.convert_dtypes()
df_ultimate_issuer_relationship_LEI = osc_ingest_trino.enforce_sql_column_names(
df_ultimate_issuer_relationship_LEI)
# -
# # Creating data set for ISIN to LEI mapping
# So far there is no access to the data via the GLEIF AWS bucket / this will hopefully change in the future
# +
url_gleif_isin_LEI = "https://isinmapping.gleif.org/api/v2/isin-lei/3381/download"
tmp_file_name = "isin-lei-20211028T070142.zip"
df_mapping_LEI_ISIN = pd.read_csv(url_gleif_isin_LEI, compression="zip")
str_timestamp = f'{date_str_data[:4]}-{date_str_data[4:6]}-{date_str_data[6:]}T{time_str_data[:2]}'
data_time_stamp = pd.Timestamp(str_timestamp, tz='UTC')
df_mapping_LEI_ISIN["time_stamp"] = data_time_stamp
df_mapping_LEI_ISIN = df_mapping_LEI_ISIN.convert_dtypes()
df_mapping_LEI_ISIN = osc_ingest_trino.enforce_sql_column_names(
df_mapping_LEI_ISIN)
# -
# # Creating data set for company hierarchy tree
# +
anytree_root = create_company_hierarchy_tree_structure(pd_rr_data_set)
from anytree.exporter import DictExporter
exporter = DictExporter()
dict_anytree_formating = exporter.export(anytree_root)
# Renaming for better readability
dict_key_renaming = {"name": "entity_LEI", "children": "entity_children"}
rename_recursive_key_dicts(
dict_to_rename=dict_anytree_formating,
dict_of_key_old_to_new=dict_key_renaming,
new_name_dict_holder_key="entity_children",
)
add_recursive_key_to_dict(
dict_to_add_to=dict_anytree_formating,
key_to_add="entity_hierarchy_level",
value_to_add=-1,
name_dict_holder_key="entity_children",
)
list_hierarchy_items = dict_anytree_formating.get("entity_children")
import json
json_hierarchy_file = f"{date_str_data}_{time_str_data}_company_hierarchy_tree.json"
with open(f"/tmp/{json_hierarchy_file}", "w") as file:
json.dump(list_hierarchy_items, file)
# -
# # Create an S3 client to upload the data to a S3 bucket
# +
dotenv_dir = os.environ.get(
"CREDENTIAL_DOTENV_DIR", os.environ.get("PWD", "/opt/app-root/src")
)
dotenv_path = pathlib.Path(dotenv_dir) / "credentials.env"
if os.path.exists(dotenv_path):
load_dotenv(dotenv_path=dotenv_path, override=True)
# +
# Create an S3 client to upload the files
s3 = boto3.client(
service_name="s3",
endpoint_url=os.environ["S3_DEV_ENDPOINT"],
aws_access_key_id=os.environ["S3_DEV_ACCESS_KEY"],
aws_secret_access_key=os.environ["S3_DEV_SECRET_KEY"],
)
# This affects both upload location and SQL schema/catalog information
gleif_schema = "gleif"
# -
# # Upload data set ultimate issuer LEI mapping to S3
# +
df_ultimate_issuer_relationship_LEI.info(verbose=True)
tmp_file_name = f"{date_str_data}_{time_str_data}_ultimate_issuer_LEI_mapping.parquet"
df_ultimate_issuer_relationship_LEI.to_parquet(f"/tmp/{tmp_file_name}", index=False)
s3.upload_file(
Bucket=os.environ["S3_DEV_BUCKET"],
Key=f"trino/{gleif_schema}/date={date_str_data}/time={time_str_data}/ultimateissuer/{tmp_file_name}",
Filename=f"/tmp/{tmp_file_name}",
)
# -
# # Upload data set ISIN LEI mapping to S3
# +
df_mapping_LEI_ISIN.info(verbose=True)
tmp_file_name = f"{date_str_data}_{time_str_data}_ISIN_LEI_mapping.parquet"
df_mapping_LEI_ISIN.to_parquet(f"/tmp/{tmp_file_name}", index=False)
s3.upload_file(
Bucket=os.environ["S3_DEV_BUCKET"],
Key=f"trino/{gleif_schema}/date={date_str_data}/time={time_str_data}/isinlei/{tmp_file_name}",
Filename=f"/tmp/{tmp_file_name}",
)
# -
# # Upload data set company hierarchy tree to S3
s3.upload_file(
Bucket=os.environ["S3_DEV_BUCKET"],
Key=f"trino/{gleif_schema}/date={date_str_data}/time={time_str_data}/companyhierarchy/{json_hierarchy_file}",
Filename=f"/tmp/{json_hierarchy_file}",
)
# # Create Schema/Table and upload .parquet files to DB via trino
# +
conn = trino.dbapi.connect(
host=os.environ["TRINO_HOST"],
port=int(os.environ["TRINO_PORT"]),
user=os.environ["TRINO_USER"],
http_scheme="https",
auth=trino.auth.JWTAuthentication(os.environ["TRINO_PASSWD"]),
verify=True,
catalog='osc_datacommons_dev',
)
cur = conn.cursor()
# Define tablenames to use
tablename_isin_lei = "gleif_isin_lei"
tablename_direct_issuer_ultimate_issuer = "gleif_direct_issuer_ultimate_issuer"
# -
# Show available schemas to ensure trino connection is set correctly
cur.execute("show schemas")
cur.fetchall()
# Only permitted with admin status
cur.execute(f"create schema if not exists {gleif_schema}")
cur.fetchall()
# Dropping tables for ISIN/LEI mapping and Direct Issuer/Ultimate Issuer in case they exist already
cur.execute(f"drop table if exists {gleif_schema}.{tablename_isin_lei}")
cur.fetchall()
cur.execute(
f"drop table if exists {gleif_schema}.{tablename_direct_issuer_ultimate_issuer}"
)
cur.fetchall()
# Create table for ISIN LEI mapping
# +
column_schema = osc_ingest_trino.create_table_schema_pairs(df_mapping_LEI_ISIN)
tabledef = f"""create table if not exists {gleif_schema}.{tablename_isin_lei}(
{column_schema}
) with (
format = 'parquet',
external_location = 's3a://{os.environ["S3_DEV_BUCKET"]}/trino/{gleif_schema}/date={date_str_data}/time={time_str_data}/isinlei/'
)"""
print(tabledef)
# tables created externally may not show up immediately in cloud-beaver
cur.execute(tabledef)
cur.fetchall()
# -
# Create table for issuer/ultimate issuer mapping
# +
column_schema = osc_ingest_trino.create_table_schema_pairs(df_ultimate_issuer_relationship_LEI)
tabledef = f"""create table if not exists {gleif_schema}.{tablename_direct_issuer_ultimate_issuer}(
{column_schema}
) with (
format = 'parquet',
external_location = 's3a://{os.environ["S3_DEV_BUCKET"]}/trino/{gleif_schema}/date={date_str_data}/time={time_str_data}/ultimateissuer/'
)"""
print(tabledef)
# tables created externally may not show up immediately in cloud-beaver
cur.execute(tabledef)
cur.fetchall()
# -
# Testing connection
# +
import trino
from sqlalchemy.engine import create_engine
sqlstring = "trino://{user}@{host}:{port}/".format(
user=os.environ["TRINO_USER"],
host=os.environ["TRINO_HOST"],
port=os.environ["TRINO_PORT"],
)
sqlargs = {
"auth": trino.auth.JWTAuthentication(os.environ["TRINO_PASSWD"]),
"http_scheme": "https",
"catalog": "osc_datacommons_dev",
}
engine = create_engine(sqlstring, connect_args=sqlargs)
print("connecting with engine " + str(engine))
connection = engine.connect()
df = pd.read_sql("show catalogs", engine).convert_dtypes()
df
# -
engine.execute(f"show tables in {gleif_schema}").fetchall()
engine.execute(f"select count (*) from {gleif_schema}.gleif_direct_issuer_ultimate_issuer").fetchall()
# +
sqlquery = (
f"select * from {gleif_schema}.{tablename_direct_issuer_ultimate_issuer}"
)
df = pd.read_sql(sqlquery, engine).convert_dtypes()
df
# -
engine.execute(f"select * from {gleif_schema}.{tablename_direct_issuer_ultimate_issuer} where direct_issuer_lei='1HNPXZSMMB7HMBMVBS46'").fetchall()
engine.execute(f"select * from {gleif_schema}.{tablename_direct_issuer_ultimate_issuer} where direct_issuer_lei='8YQ2GSDWYZXO2EDN3511'").fetchall()
len(df_mapping_LEI_ISIN)
| notebooks/gleif_ingestion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data preprocessing
# ## Load data
# +
import gzip
interactions = {}
data = []
# Load data
org_id = '9606' # Change to 9606 for Human
with gzip.open(f'data/{org_id}.protein.links.v11.0.txt.gz', 'rt') as f:
next(f) # Skip header
for line in f:
p1, p2, score = line.strip().split()
if float(score) < 700: # Filter high confidence interactions
continue
if p1 not in interactions:
interactions[p1] = set()
if p2 not in interactions:
interactions[p2] = set()
if p2 not in interactions[p1]:
interactions[p1].add(p2)
interactions[p2].add(p1)
data.append((p1, p2))
print('Total number of interactions:', len(data))
print('Total number of proteins:', len(interactions.keys()))
# -
# ## Split training, validation and testing data
#
# +
import numpy as np
import math
np.random.seed(seed=0) # Fix random seed for reproducibility
np.random.shuffle(data)
train_n = int(math.ceil(len(data) * 0.8))
valid_n = int(math.ceil(train_n * 0.8))
train_data = data[:valid_n]
valid_data = data[valid_n:train_n]
test_data = data[train_n:]
print('Number of training interactions:', len(train_data))
print('Number of validation interactions:', len(valid_data))
print('Number of testing interactions:', len(test_data))
# -
# ## Save the data
# +
def save(filename, data):
with open(filename, 'w') as f:
for p1, p2 in data:
f.write(f'{p1}\t{p2}\n')
f.write(f'{p2}\t{p1}\n')
save(f'data/train/{org_id}.protein.links.v11.0.txt', train_data)
save(f'data/valid/{org_id}.protein.links.v11.0.txt', valid_data)
save(f'data/test/{org_id}.protein.links.v11.0.txt', test_data)
# -
# ## Generate negative interactions
import random
proteins =set ()
negatives = []
for (p1,p2) in data:
proteins.add(p1)
proteins.add(p2)
while len(negatives)<len(data):
s = random.sample(proteins, 2)
prot1= s[0]
prot2= s[1]
if (prot1,prot2) in negatives or (prot2,prot1) in negatives :
continue
if prot1 not in interactions[prot2]:
negatives.append((prot1, prot2))
print('Total number of negative interactions:', len(negatives))
# Split negative data
neg_train_data = negatives[:valid_n]
neg_valid_data = negatives[valid_n:train_n]
neg_test_data = negatives[train_n:]
print('Number of negative training interactions:', len(neg_train_data))
print('Number of negative validation interactions:', len(neg_valid_data))
print('Number of negative testing interactions:', len(neg_test_data))
# Save negative data
save(f'data/train/{org_id}.negative_interactions.txt', neg_train_data)
save(f'data/valid/{org_id}.negative_interactions.txt', neg_valid_data)
save(f'data/test/{org_id}.negative_interactions.txt', neg_test_data)
# ## Preprocess GO annotations
# ### Load id mapping between annotation database and StringDB IDs
# +
mapping = {}
source = {'4932': 'SGD_ID', '9606': 'Ensembl_UniProt_AC'} # mapping source
with gzip.open(f'data/{org_id}.protein.aliases.v11.0.txt.gz', 'rt') as f:
next(f) # Skip header
for line in f:
string_id, p_id, sources = line.strip().split('\t')
if source[org_id] not in sources.split():
continue
if p_id not in mapping:
mapping[p_id] = set()
mapping[p_id].add(string_id)
print('Loaded mappings', len(mapping))
# -
# ### Load annotations
# +
gaf_files = {'4932': 'sgd.gaf.gz', '9606': 'goa_human.gaf.gz'}
annotations = set()
with gzip.open(f'data/{gaf_files[org_id]}', 'rt') as f:
for line in f:
if line.startswith('!'): # Skip header
continue
it = line.strip().split('\t')
p_id = it[1]
go_id = it[4]
if it[6] == 'IEA' or it[6] == 'ND': # Ignore predicted or no data annotations
continue
if p_id not in mapping: # Not in StringDB
continue
s_ids = mapping[p_id]
for s_id in s_ids:
annotations.add((s_id, go_id))
print('Number of annotations:', len(annotations))
# Save annotations
with open(f'data/train/{org_id}.annotation.txt', 'w') as f:
for p_id, go_id in annotations:
f.write(f'{p_id}\t{go_id}\n')
# -
# ## Generate Plain Training Data
# +
import os
tdf = open(f'data/train/{org_id}.plain.nt', 'w')
# Load GO
with open('data/go.obo') as f:
tid = ''
for line in f:
line = line.strip()
if line.startswith('id:'):
tid = line[4:]
if not tid.startswith('GO:'):
continue
if line.startswith('is_a:'):
tid2 = line[6:].split(' ! ')[0]
tdf.write(f'<http://{tid}> <http://is_a> <http://{tid2}> .\n')
if line.startswith('relationship:'):
it = line[14:].split(' ! ')[0].split()
tdf.write(f'<http://{tid}> <http://{it[0]}> <http://{it[1]}> .\n')
# Load interactions
with open(f'data/train/{org_id}.protein.links.v11.0.txt') as f:
for line in f:
it = line.strip().split()
tdf.write(f'<http://{it[0]}> <http://interacts> <http://{it[1]}> .\n')
# Load annotations
with open(f'data/train/{org_id}.annotation.txt') as f:
for line in f:
it = line.strip().split()
tdf.write(f'<http://{it[0]}> <http://hasFunction> <http://{it[1]}> .\n')
tdf.close()
if not os.path.exists('data/transe'):
os.makedirs('data/transe')
# ! wc -l 'data/train/{org_id}.plain.nt'
# -
# ## Generate Classes Training Data for ELEmbeddings
if not os.path.exists('data/elembeddings'):
os.makedirs('data/elembeddings')
# ! groovy el-embeddings/GenerateTrainingDataClasses -i 'data/train/{org_id}.protein.links.v11.0.txt' -a 'data/train/{org_id}.annotation.txt' -o 'data/train/{org_id}.classes.owl'
# ### Normalize training data classes into four normal forms
# +
# ! groovy -cp el-embeddings/jar/jcel.jar el-embeddings/Normalizer.groovy -i 'data/train/{org_id}.classes.owl' -o 'data/train/{org_id}.classes-normalized.owl'
# -
# ## Generate RDF Representation of ELEmbeddings training data
# +
# ! rapper 'data/train/{org_id}.classes.owl' -o ntriples > 'data/train/{org_id}.classes-rdf.nt'
# -
# ## Generate Onto/OPA2Vec-compatible associations
import re
# generate OPA2VEC compatible associations
with open(f'data/train/{org_id}.OPA_associations.txt', 'w') as f:
for p_id, go_id in annotations:
go_num = re.split ('[A-Z]|:+',go_id)
f.write (str(p_id)+" \t"+"<http://purl.obolibrary.org/obo/GO_"+str(go_num[3])+">\n")
| data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import dataset
import pandas as pd
import numpy as np
df = pd.read_csv('cbb.csv')
# # Inspect dataset
## view the first 5 rows of the dataset
df.head()
df.info()
df.describe()
## size of dataset
df.shape
## any NA values
df.isnull().sum()
# #### From this, I can conclude/confirm that there are only two columns that contain NA values. In the case of this dataset and for the purposes of this project, these NA values actually tell us something: that "(1757-340)= 1417" teams have never made it to the March Madness tournament.
#
# ### Inspecting the target column, "SEED"
df['SEED'].values
df['SEED'].value_counts
print(np.min(df['SEED']))
print(np.max(df['SEED']))
# #### This informs me that the highest SEED number is 16, for each basketball tournament. (Refer to *00_dataset-variables.ipynb* for more information on "SEED")
df['SEED'].fillna(value=0, inplace = True)
df['SEED'].isnull().sum()
# #### The null values in the dataset have been converted to 0s.
df['YEAR'].unique()
# #### The dataset spans 5 years.
| 10-import_and_inspect.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Short training sequences
# In this notebook I will try to text whether sequences trained for very short periods of work well in a couple of domains.
#
# * Learning long list of sequences.
# * Disambugation.
# +
import pprint
import subprocess
import sys
sys.path.append('../')
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
import matplotlib.gridspec as gridspec
from mpl_toolkits.axes_grid1 import make_axes_locatable
import seaborn as sns
# %matplotlib inline
np.set_printoptions(suppress=True, precision=2)
sns.set(font_scale=2.0)
# -
# #### Git machinery
run_old_version = False
if run_old_version:
hash_when_file_was_written = '063dfb2c68ee9f1a3f22e951cfe291a03543e9bd'
hash_at_the_moment = subprocess.check_output(["git", 'rev-parse', 'HEAD']).strip()
print('Actual hash', hash_at_the_moment)
print('Hash of the commit used to run the simulation', hash_when_file_was_written)
subprocess.call(['git', 'checkout', hash_when_file_was_written])
# #### Load libraries
from network import Protocol, BCPNNFast, NetworkManager
from plotting_functions import plot_winning_pattern
from analysis_functions import calculate_recall_success
from analysis_functions import calculate_timings
# ## Simple example
# +
# Network parameters
minicolumns = 40
hypercolums = 4
number_of_patterns = 20
patterns_indexes = [i for i in range(number_of_patterns)]
# Training parameters
dt = 0.001
training_time = 0.020
inter_pulse_interval = 0.0
epochs = 3
# Build the network, manager and protocol
nn = BCPNNFast(hypercolumns=hypercolums, minicolumns=minicolumns)
manager = NetworkManager(nn, dt=dt, values_to_save=['o'])
simple_protocol = Protocol()
simple_protocol.simple_protocol(patterns_indexes=patterns_indexes, training_time=training_time,
inter_sequence_interval=1.0, epochs=epochs)
# -
# Train
manager.run_network_protocol(protocol=simple_protocol, verbose=False)
# Recall
T_recall = 5
T_cue = training_time
manager.run_network_recall(T_recall=T_recall, T_cue=0.020, I_cue=0)
# Plot
plot_winning_pattern(manager);
# +
timings = calculate_timings(manager=manager, remove=0.100)
timings = [timings[index] for index in patterns_indexes]
pattern_number = [time[0] for time in timings]
recall_times = [time[1] for time in timings]
# +
import seaborn as sns
sns.set(font_scale=2.0)
fig = plt.figure(figsize=(16, 12))
ax = fig.add_subplot(111)
ax.plot(pattern_number, recall_times, '*-', markersize=12)
ax.set_title('Recall duration as a function of the position in the chain')
ax.set_xlabel('Pattern Number')
ax.set_ylabel('Recall duration')
ax.set_ylim([0, 1.1]);
# -
# # Long sequence probing at different points
# +
# Network parameters
minicolumns = 80
hypercolums = 4
number_of_patterns = 60
patterns_indexes = [i for i in range(number_of_patterns)]
# Training parameters
dt = 0.001
training_time = 0.020
inter_pulse_interval = 0.0
epochs = 3
# Build the network, manager and protocol
nn = BCPNNFast(hypercolumns=hypercolums, minicolumns=minicolumns)
manager = NetworkManager(nn, dt=dt, values_to_save=['o'])
simple_protocol = Protocol()
simple_protocol.simple_protocol(patterns_indexes=patterns_indexes, training_time=training_time,
inter_sequence_interval=1.0, epochs=epochs)
# -
# Train
manager.run_network_protocol(protocol=simple_protocol, verbose=False)
# Recall
successes = []
T_cue = training_time
I_cue_range = np.arange(0, number_of_patterns - 1, 5)
for I_cue in I_cue_range:
recall_patterns_indexes = [i for i in range(I_cue, number_of_patterns)]
T_recall = len(recall_patterns_indexes) * 9 * training_time
print(I_cue)
print(T_recall)
success = calculate_recall_success(manager, T_recall=T_recall, T_cue=0.100, I_cue=I_cue, n=10,
patterns_indexes=recall_patterns_indexes)
print(success)
timings = calculate_timings(manager=manager, remove=0.010)
pprint.pprint(timings)
successes.append(success)
recall_patterns_indexes
# +
fig = plt.figure(figsize=(16, 12))
ax = fig.add_subplot(111)
ax.plot(I_cue_range, successes, '*-', markersize=13)
ax.set_title('Recall as a function of probe position')
ax.set_xlabel('Probe position')
ax.set_ylabel('Recall success')
ax.set_ylim([0, 110]);
# -
# #### Git reload
if run_old_version:
subprocess.call(['git', 'checkout', 'master'])
| notebooks/2017-04-13(Short Training sequences).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: torch
# language: python
# name: torch
# ---
import os
import re
import json
import copy
import nltk
import numpy as np
from tqdm.notebook import tqdm
from bs4 import BeautifulSoup
'''
Change this to the root directory of your patent collection.
You will need to make sure that each folder contains the <Patent_ID>.xml of the patent document
'''
xml_root = 'Table_files/Tables'
def parse_table(table):
def remove_spaces(cell):
return re.sub('\s+', '', cell)
rows = table.find_all('row')
extracted_table = []
meta_data = {}
meta_data['tid'] = table['id']
chemistry_list = []
for i, r in enumerate(rows):
extracted_row = []
for j, entry in enumerate(r.find_all('entry')):
if entry.find_all('chemistry') != 0:
for chemistry in entry.find_all('chemistry'):
chemistry_list.append((i, j, str(chemistry)))
if entry.has_attr('namest'):
# print(entry)
try:
if entry['namest'].startswith('col'):
start = int(entry['namest'][3:])
end = int(entry['nameend'][3:])
elif entry['namest'] == 'offset':
start = len(extracted_row)
end = int(entry['nameend'])
else:
start = int(entry['namest'])
end = int(entry['nameend'])
except:
print(table)
span = end - start + 1
extracted_row.append(remove_spaces(entry.get_text()))
for i in range(1, span):
extracted_row.append('')
else:
extracted_row.append(remove_spaces(entry.get_text()))
extracted_table.append(extracted_row)
meta_data['chemistry'] = chemistry_list
if len(table.find_all('title')) != 0:
titles = [i.get_text() for i in table.find_all('title')]
meta_data['titles'] = titles
if len(table.find_all('thead')) != 0:
header_list = []
t = table.find_all('table')[0]
childs = t.findChildren()
child_names = [c.name for c in childs]
row_count = 0
in_thead = 0
start = 0
end = 0
for i, c in enumerate(child_names):
if c == 'row':
row_count += 1
# if start of thead element
if c == 'thead':
start = row_count
in_thead = 1
# if thead ends or table ends
if (c == 'tbody' or i == len(child_names) - 1) and in_thead == 1:
end = row_count
header_list.append((start, end))
in_thead = 0
start, end = 0, 0
meta_data['thead'] = header_list
return extracted_table, meta_data
def extract_all(root_dir):
tables = []
xml_list = os.listdir(root_dir)
for pid in tqdm(xml_list):
xml_path = os.path.join(root_dir, pid, pid+'.xml')
if not os.path.exists(xml_path):
continue
ts = BeautifulSoup('\n'.join(open(xml_path, 'r').read().split('\n')[1:]), 'xml').find_all('tables')
for t in ts:
text, meta_data = parse_table(t)
tables.append({
'pid': pid,
'tid': meta_data['tid'],
'meta_data': meta_data,
'data': text,
})
return tables
tables = extract_all(xml_root)
json.dump(tables, open('extracted_tables.json', 'w+'))
| extract_tables_from_xmls.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import re
import csv
with open("uniprot6239reviewedfastasonly.fasta") as uniprot:
uniprot = uniprot.readlines()
counter = 0
gene_ontology=None
filename=None
protein_name=None
gene_name=None
gene_ontology=[]
#print(re.findall(">.*", uniprot))
for i in uniprot[0:]:
if i.startswith(">") is True: #Finds the fasta header
filename =re.findall(">sp\|(.*)\|", i)[0] #regex: everything in between the |
protein_name =re.findall(">sp.*L (.*) OS", i)[0]# regex: From L to the naming of the organism(i.e. the protein name)
#
try:#
gene_name= re.findall(">sp.*GN=(.*) PE", i)[0] #Try assign gene names to variable
except:
gene_name=None #Not unifrom dataset
gene_ontology.append([filename, protein_name, gene_name])
#print(i)
def protein_table_maker(fasta):
#Takes a fasta from uniprot and tries to return a table with the filename, protein name, and gene name
filename =re.findall(">sp\|(.*)\|", fasta)[0]
protein_name =re.findall(">sp.*L (.*) OS", fasta)[0]
try:
gene_name= re.findall(">sp.*GN=(.*) PE", fasta)[0]
except:
gene_name=""
gene_name_length=len(gene_name)
if gene_name in protein_name and gene_name_length!=0:
# Many of these fasta files have the gen name inside of them so here I'm searching for a gene name in the
# protein name and extracting it if it exists in two places
subtractor =len(gene_name)
length =len(protein_name)
#old_protein_name = protein_name
#print(protein_name)
protein_name = protein_name[0:length-subtractor]
#print(gene_name)
return(filename+"|"+ protein_name+"|"+ gene_name)
for i in uniprot[:]:
if i.startswith(">") is True:
handle = open("protein_names.csv", "a")
protein_writer = csv.writer(handle, delimiter=",")
row = protein_table_maker(i)
separated_row = row.split("|")
protein_writer.writerow(separated_row)
handle.close()
one = "abc"
two = "abc dce the care"
type(one in two)
type(len(one)==3)
# cd proteome_analysis/
# ls
| Format_proteins_ontology.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Notebook for Codementor Machine Learning Class 2
# ## U.S. Dept. of Education College Scorecard
# ### Topics
# * Data Science career discussion
# * Incorporate insights from data characterization (Class 1)
# * Principle Components Analysis (PCA)
# * K-means clustering on transformed (PCA) data
# * Provide a prototype useful to an engineer
#
# ### Career Discussion
#
# * Now that we've examined and characterized our client's data, we'll need to quickly create and deliver a prototype product. Start quickly, fail fast, fail often. (Engineering wisdom.) We accept that all models are wrong, but some are useful. Each time we create a product, we test and determine its value. We envision a better product, throw away the old one, and build the new one.
#
# * In general, your clients will not be interested in your notebooks, your prototypes, your graphs, your studies, etc. (Your data science colleagues will appreciate these things, but your clients/management want results.) It is extremely important to your success as a data scientist to find out exactly what your client wants. It is your job to provide it.
#
# * For our purposes, let's assume we are working with our client's engineer who will provide us with an input query in JSON (JavaScript Object Notation). Example: { "home_state": "NJ", "major": "engineering", "math": 600, "verbal": 620, "writing": 580, "out_of_state": false, "budget": 30000}. We are to provide useful information about recommended academic colleges and universities in return, in JSON.
#
# * We have not yet developed an engine which uses _all_ of the input data. However, in this case we feel it is important to provide something quickly. This way our engineer can get to work. Then, we can continue to develop the engine and provide updates. We'll also get feedback. This way, we can change or modify our course if necessary.
# ### 1. Load and prepare raw data
# %matplotlib inline
# allows plotting in cells, we'll use later on.
import pylab
pylab.rcParams['figure.figsize'] = (10, 6) # set a larger figure size
import sqlite3
conn = sqlite3.connect('C:/Users/peter/Documents/Codementor/MLClass/data/college/database.sqlite')
import pandas
query = """
SELECT UNITID,
INSTNM,
CONTROL as type,
COSTT4_A AvgYrCostAcademic,
COSTT4_P AvgYrCostProgram,
md_earn_wne_p10 medEarn10yrs,
md_earn_wne_p6 medEarn6yrs,
Year,
UGDS NumStudents,
SATMTMID Math,
SATVRMID Verbal,
SATWRMID Writing,
STABBR State,
GRAD_DEBT_MDN DebtAtGrad,
C150_4 completionRate
FROM Scorecard
WHERE Year='{0}'
"""
# I found that 2011 is the latest year with earnings data!
# Plan: use 2013 cost and 2011 earnings.
# Earnings have been approximately flat in adjusted dollars for the past decade;
# so we'll just need to adjust 2011 dollars to 2013 dollars at some point
df2013 = pandas.read_sql(query.format(2013), conn)
df2011 = pandas.read_sql(query.format(2011), conn)
# #### Index by the unique identifier (but remember, it is not a requirement that indices have unique values for each row)
df2011.set_index(['UNITID'], inplace=True)
df2013.set_index(['UNITID'], inplace=True)
# #### Create the numeric earnings column (year 2011 data) and add our computed SAT column (year 2013 data).
df2011['earn10'] = pandas.to_numeric(df2011['medEarn10yrs'], errors='coerce')
df2013['sat'] = df2013['Math'] + df2013['Verbal']
# Using Pandas.concat, let's tack on the 2011 earnings data to our 2013 dataframe. This is done by matching UNITIDs. In SQL, this would be an inner join and we'd have to explicitly state the join variable (UNITID) in each table. In Pandas, it's assumed we'll join on the DataFrame/Series index (and we are free to state otherwise). The Pandas concat function handles the join.
dfearn = df2011['earn10'].dropna()
dfearn.head() # note the Series retains the UNITID index
type(dfearn)
# before
df2013.columns
df_joined = pandas.concat([df2013, dfearn], axis=1, join='inner') # equivalent to sql inner join on UNITID
# +
# after
df_joined.columns
# -
# #### Raw data prep complete: df_joined
df_joined.head()
# ### 2. Principle Components Analysis (PCA)
#
# Reference: http://scikit-learn.org/stable/modules/decomposition.html#pca
# PCA is a _dimensionality reduction_ technique. Dimensionality reduction means we have data that are redundant/related in some way, and we want to simplify things. In our case, we have already seen our variables are correlated. High incoming SAT scores are related to higher graduation rates. So let's see if we can distill our 4 variables down to one or two.
# From Wikipedia: "PCA is mathematically defined as an orthogonal linear transformation that transforms the data to a new coordinate system such that the greatest variance by some projection of the data comes to lie on the first coordinate (called the first principal component), the second greatest variance on the second coordinate, and so on."
# #### Select our 4 numeric variables
dfc = df_joined[['sat', 'AvgYrCostAcademic', 'earn10', 'completionRate']].dropna()
# #### Scale, to put all variables on equal footing, numerically
from sklearn.preprocessing import scale
dfscale = pandas.DataFrame(scale(dfc), index=dfc.index).rename(columns={0:'sat_scaled', 1:'cost', 2:'earn', 3:'gradrate'})
# #### Apply PCA to our scaled data
from sklearn.decomposition import PCA
pca = PCA(n_components=4)
pca_array = pca.fit_transform(dfscale[[u'sat_scaled', u'cost', u'earn', u'gradrate']])
dfscale.head()
type(pca_array)
pca_array[0:5]
# Let's see how well the PCA went:
pca.explained_variance_ratio_
# the transformation matrix it came up with, for linear algebra fans:
pca.components_
# We have a happy case where we can pack most of the useful information (71 + 14 = 85%) into the first two PCA variables. This means we can cluster using only the first two PCA variables and visualize it completely in a 2-D chart.
# Let's have a look at our transformed variables, which we assigned to `pca_array`:
pca_array[0:20]
# Let's look at our transformed data alongside the original data:
dfscale['PCA0'] = pca_array[:,0]
dfscale['PCA1'] = pca_array[:,1]
dfscale['PCA2'] = pca_array[:,2]
dfscale['PCA3'] = pca_array[:,3]
dfscale.head(10) # shows data before PCA (left 4 columns) and after PCA (right 4 columns)
dfscale.describe()
# ### 3. Clustering (via Kmeans)
# Create an instance of the clustering object
from sklearn.cluster import KMeans
km = KMeans(init='k-means++', n_clusters=7)
# Cluster using just PCA0 and PCA1. Store the cluster numbers in our dataframe.
# +
dfscale['KM_PCA_cluster'] = km.fit_predict(dfscale[['PCA0','PCA1']])
# -
# For purposes of comparision, cluster will all the original data:
dfscale['KM_cluster'] = km.fit_predict(dfscale[['sat_scaled', 'cost', 'earn', 'gradrate']])
dfscale.head()
# Let's have a look! First, clustering on PCA0 and PCA1 only:
# +
import matplotlib.pyplot as plt
groups = dfscale.groupby('KM_PCA_cluster')
fig, ax = plt.subplots()
ax.margins(0.05) # Optional, just adds 5% padding to the autoscaling
for name, group in groups:
ax.plot(group.PCA0, group.PCA1, marker='o', linestyle='', ms=5, label=name)
ax.legend()
ax.set_xlabel("PCA0")
ax.set_ylabel("PCA1")
ax.text(-2, 3, 'PCA Cluster Visualization')
plt.show()
# -
groups
groups.median().sort_values(by='sat_scaled') # bottom cluster is the low-cost group!
# Let's see how clustering with the raw data compared to clustering PCA-transformed variables.
groups_raw = dfscale.groupby('KM_cluster')
groups_raw.median().sort_values(by='sat_scaled')
# The median values of the clusters are nearly the same. Not much difference, so far. What about visualization? Well, the first problem is: what do we graph on each axis?
from pandas.tools.plotting import scatter_matrix
scatter_matrix(dfscale[['sat_scaled', 'cost', 'earn', 'gradrate']])
# Let's try one that already shows distinct groups, such as cost vs. gradrate.
# +
groups = dfscale.groupby('KM_cluster')
fig, ax = plt.subplots()
ax.margins(0.05)
for name, group in groups:
ax.plot(group.gradrate, group.cost, marker='o', linestyle='', ms=5, label=name)
ax.legend()
ax.set_xlabel("gradrate")
ax.set_ylabel("cost")
ax.text(-2, 3, 'Cluster Visualization without PCA')
plt.show()
# -
# Not too bad-- but still a bit of a mess compared to our PCA. There, the clustering algorithm ran on two dimensional data. Here, it made 4-dimensional clusters. If you feel like you should be able peek your head around the side of the graph to get a better view of the clusters, PCA is exactly that; a transformation of variables. It transforms in such as way as to pack as much information as possible into the first PCA variable, then the second, and so on. That is why PCA is referred to as a "dimensionality reduction" technique. You can also think of clustering as a dimensionality reduction technque, where we map down to one categorical dimension (cluster number).
# ### 4. Prototype construction using the clustered data
# Back to business-- my plan for a prototype product-- (yours could be different, and better). Let's take the SAT score and align it to the nearest cluster median. We'll return three schools from that cluster within the budget. Then we'll take three schools from the cluster above ("reach schools") and below ("safe schools"). If we're in the top or bottom cluster to begin with, we'll return 3 additional schools from the matching cluster. So that we can easily get the cluster above and below, let's also re-map the cluster numbers.
# #### Re-map cluster numbers. Right now the numbers don't mean anything.
# Let's make cluster 0 correspond to the lowest median SAT scores,
# cluster 1 to the next highest, etc., cluster 6 to the highest.
# Save the new meaningful cluster number in our dataframe as `cluster ordinal`.
groups = dfscale.groupby('KM_PCA_cluster')
cluster_number_map = groups.median().sort_values(by='sat_scaled').index # cluster numbers sorted from lowest to highest SAT
cluster_number_map
d = { cluster_number_map[i]: i for i in range(7) }
d
d[6]
d[2]
cluster_ord = dfscale.KM_PCA_cluster.map(lambda x: d[x])
cluster_ord.head(10)
cluster_ord = dfscale.KM_PCA_cluster.map(lambda x: d[x])
dfscale['cluster_ordinal'] = cluster_ord
dfscale.head()
groups = dfscale.groupby('KM_PCA_cluster')
groups.median().sort_values(by='sat_scaled')
# #### Side lesson: What's a lambda function??
def add_three(x):
return x + 3
add_three(10)
def foo(some_fun):
print "you passed", some_fun
print "I will now put 10 in that function"
print some_fun(10)
foo(add_three)
foo(lambda x: x + 3)
foo(lambda x: 2*x)
# #### *end of side lesson on lambda functions*
# Design decision: let's work with unscaled cost and SAT scores. This way we do not have to scale incoming data. Since we've retained the school IDs, we simply need to bring our cluster labels back to the raw dataframe. Let's see how this would work in the notebook, then code up the prototype as a python command-line program.
df_joined.head()
dfscale.head()
df_out = pandas.concat([dfscale, df_joined], axis=1, join='inner')
df_out.head()
df_out.columns
out_groups = df_out[['cluster_ordinal','sat','AvgYrCostAcademic','earn10', 'completionRate']].groupby('cluster_ordinal')
df_cluster_lookup = out_groups.median()
df_cluster_lookup
# There is a handy pandas function for our purposes called 'argsort'. We can use it to return the closest clusters to our query SAT score, in order.
query_sat = 1100
(df_cluster_lookup.sat - query_sat).abs()
(df_cluster_lookup.sat - query_sat).abs().argsort()
(df_cluster_lookup.sat - query_sat).abs().argsort()[0]
# +
df_cluster_lookup.iloc[(df_cluster_lookup.sat - query_sat).abs().argsort()[:1]]
# -
# I still like having ordinal labels for the clusters-- it clearly validates our argsort technique. Now, let's get to picking out some schools, given a cluster and a budget.
# +
query_sat = 1100
query_maxcost = 40000
query_sat_add_max = 10
query_cluster = 3 # got this with argsort above
c1 = df_out['cluster_ordinal'] == query_cluster
c2 = df_out['AvgYrCostAcademic'] <= query_maxcost
c3 = df_out['sat'] <= query_sat + query_sat_add_max
condition = c1 & c2 & c3
df_out.loc[condition].sort_values(by='sat', ascending=False)[0:3]
# -
# Really we just need the UNITIDS
df_out.loc[condition].sort_values(by='sat', ascending=False)[0:3].index
# +
picks = df_out.loc[condition].sort_values(by='sat', ascending=False)[0:5].index
df_response = df_joined.loc[picks,
['INSTNM', 'type', 'NumStudents', 'AvgYrCostAcademic', 'sat', 'earn10']]
df_response
# -
df_response.to_json(orient='records')
df_response.to_json()
# +
# pandas.DataFrame.to_json?
# -
| college-prototype.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Working with the github API
#
# In this notebook, we work explore the github API by makig some example requests. The full documentation on the github API is available here: https://developer.github.com/v3/
# For all of our calls, we will be using the Python requests library. Documentation on requests is available here: http://docs.python-requests.org/en/master/
# +
# import the requests library
import requests
# import the json parsing library
import json
# -
# base URL for all github API requests
base_url = 'https://api.github.com'
# make a basic GET request to the 'users' collection
rsp = requests.get('{}/users'.format(base_url))
# check the status code:
rsp.status_code
# pull json out of the response:
result = rsp.json()
# what kind of thingy is result?
type(result)
# oh, interesting; it's just a python list. let's check how many user records were returned:
len(result)
# let's look at the first record:
result[0]
# and what kind of thingy is that?
type(result[0])
# oh cool, a python dictionary. we can access the data under specific keys like so:
result[0]['url']
# so now let's look ourselves:
my_github_account = 'joestubbs'
rsp2 = requests.get('{}/users/{}'.format(base_url, my_github_account))
# did that work?
rsp2.status_code
# cool. what's in there?
rsp2.json()
# ok, more detail that the summary in the listing. similarly, we can pull out specific info:
rsp2.json()['name']
# let's look at a subcollection -- the followers i have:
rsp3 = requests.get('{}/users/{}/followers'.format(base_url, my_github_account))
rsp3.status_code
rsp3.json()[0]
| notebooks/REST APIs - the github API.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
1 + 1
2 * 2
1 - 1
10 ^ 10
10 ** 10
"cats"
"dogs"
x = "person1"
y = "person2"
z = [x,y]
z
x = "person3"
z
z = "cats" + dfaf
my_list = [1,2,3,4,5,6,7,8,9,10]
len(my_list)
subset = my_list[2:7]
subset
odds = [1,3,5,7]
odds.append(11)
odds
odds += [13]
odds
del(odds[0])
print("Hello World")
first_dict = {}
type(first_dict)
first_dict["x"] = "some_value"
first_dict["x"]
first_dict["y"] = 1001
first_dict
new_dict = {1:"a", 2:"b", 3:"c"}
new_dict[2]
True
False
type(True)
1 == 1
2 == 1
1 < 10
1 > 10
1 <= 10
# +
x = 15
if x < 10:
print("This workshop is okay")
print("Hello world")
result = 5
# -
result
for value in range(0, 10):
print(value)
# # for loops
#
#
# +
string = "hello"
string_list = []
for ii in string:
string_list += ii
print(string_list)
# -
def fahr_to_kelvin(temp):
return ((temp - 32) * (5/9)) + 273.15
fahr_to_kelvin(95)
# +
names = ["brandon", "joseph", "jeramiah"]
def hello_name(name):
print("Hello %s" % name)
for name in names:
hello_name(name)
# -
| python_part1_follow_through.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# name: python3
# ---
# # Collectl Log Analysis
# ## Functionalities
# - Plot CPU utilization graphs.
# - Plot memory utilization graphs.
# - Plot disk I/O utilization graphs.
#
# ## Input
# Log files are read from a directory in `../data`. This directory is assumed to have the following structure:
# ```
# logs/
# [node-1]/
# collectl.tar.gz
# ...
# [node-n]/
# collectl.tar.gz
# ```
# A tarball `collectl.tar.gz` contains log files. The log file extension identifies the type of resource monitored:
# - `.cpu.gz`: CPU monitoring log file.
# - `.numa.gz`: memory monitoring log file.
# - `.dsk.gz`: disk I/O monitoring log file.
# ## Notebook Configuration
# +
########## GENERAL
# Name of the directory in `../data`
EXPERIMENT_DIRNAME = "BuzzBlogBenchmark_2021-11-11-16-06-13"
########## CPU
# Analyzed metric (options: "user", "nice", "system", "wait", "irq", "soft",
# "steal", "idle", "total", "guest", "guest_n", "intrpt")
COLLECTL_CPU_METRIC = "total"
# List of core numbers to be analyzed
COLLECTL_CPU_CORES = range(0, 8)
########## MEMORY
# Analyzed metric (options: "used", "free", "slab", "mapped", "anon", "anonh", "inactive", "hits")
COLLECTL_MEM_METRIC = "free"
########## DISK I/O
# Analyzed metric (options: "reads", "rmerge", "rkbytes", "waitr", "writes", "wmerge", "wkbytes", "waitw", "request",
# "quelen", "wait", "svctim", "util")
COLLECTL_DSK_METRIC = "quelen"
# -
# ## Notebook Setup
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import os
import pandas as pd
import sys
import warnings
warnings.filterwarnings("ignore")
sys.path.append(os.path.abspath(os.path.join("..")))
from parsers.collectl_parser import CollectlParser
from utils.utils import *
# -
# ## Log Parsing
# Build data frames
cpu = pd.concat([
pd.DataFrame.from_dict(CollectlParser(logfile, "cpu").parse()).assign(node_name=node_name)
for node_name, logfile in get_collectl_cpu_logfiles(EXPERIMENT_DIRNAME)
], ignore_index=True)
mem = pd.concat([
pd.DataFrame.from_dict(CollectlParser(logfile, "mem").parse()).assign(node_name=node_name)
for node_name, logfile in get_collectl_mem_logfiles(EXPERIMENT_DIRNAME)
], ignore_index=True)
dsk = pd.concat([
pd.DataFrame.from_dict(CollectlParser(logfile, "dsk").parse()).assign(node_name=node_name)
for node_name, logfile in get_collectl_dsk_logfiles(EXPERIMENT_DIRNAME)
], ignore_index=True)
# Filter data frames
start_time = get_experiment_start_time(EXPERIMENT_DIRNAME)
cpu = cpu[(cpu["timestamp"] > start_time) & (cpu["hw_metric"] == COLLECTL_CPU_METRIC) &
(cpu["hw_no"].isin(COLLECTL_CPU_CORES))]
mem = mem[(mem["timestamp"] > start_time) & (mem["hw_metric"] == COLLECTL_MEM_METRIC)]
dsk = dsk[(dsk["timestamp"] > start_time) & (dsk["hw_metric"] == COLLECTL_DSK_METRIC)]
# (Re) Build columns
cpu["timestamp"] = cpu.apply(lambda r: (r["timestamp"] - start_time).total_seconds(), axis=1)
mem["timestamp"] = mem.apply(lambda r: (r["timestamp"] - start_time).total_seconds(), axis=1)
dsk["timestamp"] = dsk.apply(lambda r: (r["timestamp"] - start_time).total_seconds(), axis=1)
# (Re) Create index
cpu.set_index("timestamp", inplace=True)
mem.set_index("timestamp", inplace=True)
dsk.set_index("timestamp", inplace=True)
# Get values
node_names = get_node_names(EXPERIMENT_DIRNAME)
# ## CPU Monitoring
# +
########## LOCAL CONFIG
# Minimum time (in seconds)
MIN_TIME = None
# Maximum time (in seconds)
MAX_TIME = None
# Plot CPU utilization
fig = plt.figure(figsize=(24, len(node_names) * 12))
for (i, node_name) in enumerate(node_names):
df = cpu[(cpu["node_name"] == node_name)]
if MIN_TIME:
df = df[(df["timestamp"] >= MIN_TIME)]
if MAX_TIME:
df = df[(df["timestamp"] <= MAX_TIME)]
df = df.groupby(["timestamp", "hw_no"])["value"].mean()
df = df.unstack()
ax = fig.add_subplot(len(node_names), 1, i + 1)
ax.set_xlim((df.index.min(), df.index.max()))
ax.set_ylim((0, 100))
ax.grid(alpha=0.75)
df.plot(ax=ax, kind="line", title="%s - CPU Utilization" % node_name, xlabel="Time (seconds)",
ylabel="%s (%%)" % COLLECTL_CPU_METRIC, grid=True, legend=False, yticks=range(0, 101, 10))
# -
# ## Memory Monitoring
# +
########## LOCAL CONFIG
# Minimum time (in seconds)
MIN_TIME = None
# Maximum time (in seconds)
MAX_TIME = None
# Plot memory utilization
fig = plt.figure(figsize=(24, len(node_names) * 12))
for (i, node_name) in enumerate(node_names):
df = mem[(mem["node_name"] == node_name)]
if MIN_TIME:
df = df[(df["timestamp"] >= MIN_TIME)]
if MAX_TIME:
df = df[(df["timestamp"] <= MAX_TIME)]
df = df.groupby(["timestamp", "hw_no"])["value"].mean()
df = df.unstack()
ax = fig.add_subplot(len(node_names), 1, i + 1)
ax.set_xlim((df.index.min(), df.index.max()))
ax.set_ylim((0, df.values.max()))
ax.grid(alpha=0.75)
df.plot(ax=ax, kind="line", title="%s - Mem Utilization" % node_name, xlabel="Time (seconds)",
ylabel="%s" % COLLECTL_MEM_METRIC, grid=True)
# -
# ## Disk Monitoring
# +
########## LOCAL CONFIG
# Minimum time (in seconds)
MIN_TIME = None
# Maximum time (in seconds)
MAX_TIME = None
# Plot disk I/O utilization
fig = plt.figure(figsize=(24, len(node_names) * 12))
for (i, node_name) in enumerate(node_names):
df = dsk[(dsk["node_name"] == node_name)]
if MIN_TIME:
df = df[(df["timestamp"] >= MIN_TIME)]
if MAX_TIME:
df = df[(df["timestamp"] <= MAX_TIME)]
df = df.groupby(["timestamp", "hw_no"])["value"].mean()
df = df.unstack()
ax = fig.add_subplot(len(node_names), 1, i + 1)
ax.set_xlim((df.index.min(), df.index.max()))
ax.set_ylim((0, df.values.max()))
ax.grid(alpha=0.75)
df.plot(ax=ax, kind="line", title="%s - Disk I/O Utilization" % node_name, xlabel="Time (seconds)",
ylabel="%s" % COLLECTL_DSK_METRIC, grid=True)
| analysis/notebooks/CollectlLogAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
import os
from functools import partial
sys.path.append('../')
from dataloader.MiddleburyLoader import myImageFloder, disparity_loader
from notebooks.notebook_utils import visualize_sample, download_sample, get_lidar_train_list
# -
train_list = get_lidar_train_list()
def walk_through_dataset(dataset_list):
for example in dataset_list:
print(example)
folders = example.split('/')[-3:]
folder = '/'.join(folders)
out_folder = '../data/lidar/{}'.format(folder)
download_sample(example, out_folder)
all_left_img = [os.path.join(out_folder, 'im0.png')]
all_right_img = [os.path.join(out_folder, 'im1.png')]
all_left_disp = [os.path.join(out_folder, 'disp0GT.pfm')]
all_right_disp = [os.path.join(out_folder, 'disp1GT.pfm')]
rand_scale = [0.5, 1.2]
loader_lidar = myImageFloder(all_left_img, all_right_img,
all_left_disp, right_disparity=all_right_disp,
rand_scale=rand_scale, rand_bright=[0.8, 1,2],
order=0, flip_disp_ud=True, occlusion_size=[10, 25])
for (img_l, img_r, disp_l) in loader_lidar:
visualize_sample(img_l, img_r, disp_l)
yield
train_iter = walk_through_dataset(train_list)
next(train_iter)
| notebooks/lidar_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
import numpy as np
from matplotlib import pyplot as plt
import cv2
from scipy.ndimage.morphology import distance_transform_edt
example = np.zeros((128, 128))
example[40:60, 40:70] = 255
plt.imshow(distance_transform_edt(np.abs(255 - example)))
plt.show()
ex = torch.from_numpy(distance_transform_edt(example)).float()
ex2 = torch.from_numpy(distance_transform_edt(np.abs(255 - example))).float()
masks = torch.nn.Conv2d(1, 4, kernel_size=3, bias=False).float()
weights = torch.tensor(
[
[[1, 1, 1], [0, 0, 0], [-1, -1, -1]],
[[-1, -1, -1], [0, 0, 0], [1, 1, 1]],
[[1, 0, -1], [1, 0, -1], [1, 0, -1]],
[[-1, 0, 1], [-1, 0, 1], [-1, 0, 1]],
[[1, 1, 0], [1, 0, -1], [0, -1, -1]],
[[-1, -1, 0], [-1, 0, 1], [0, 1, 1]],
[[0, 1, 1], [-1, 0, 1], [-1, -1, 0]],
[[0, -1, -1], [1, 0, -1], [1, 1, 0]],
]
).unsqueeze(1).float()
masks.weight = torch.nn.Parameter(weights)
# +
output = masks(ex.unsqueeze(0).unsqueeze(1))
output2 = masks(ex2.unsqueeze(0).unsqueeze(1))
output = torch.cat([torch.zeros(output.shape[0], 1, output.shape[2], output.shape[3], device=output.device) + 0.1, output + output2], dim=1)
output2 = torch.cat([torch.zeros(output.shape[0], 1, output.shape[2], output.shape[3], device=output.device) + 0.1, output2], dim=1)
for idx in range(4):
plt.imshow(output[0, idx, :, :].detach().numpy())
plt.show()
map_ = torch.argmax(output[0, :, :, :], dim=0).detach().numpy()
map_2 = torch.argmax(output2[0, :, :, :], dim=0).detach().numpy()
plt.imshow(map_)
plt.show()
# -
map_[45, 35:45]
| notebooks/direction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/eitanebench/linearSolver/blob/master/LinearSystemsSolver.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="-IW_jl1RHozX" colab_type="text"
# Created by: <NAME>
# Student for Material Engineering
# This is my code for solving linear systems of equations.
# Enjoy.
# + id="UgiWW9y-byqW" colab_type="code" colab={}
#first import numpy so we can work with arrays
import numpy as np
# + id="9Q57ltJJMFy1" colab_type="code" colab={}
#lets create the array from user inputs
def list2array(length):
#length = int(input("How many elements are in the Array?"))
X =np.zeros((length,length))
for n in range(length):
A2 = []
for k in range(length):
A = float(input("Write the elements one at a time then press enter.\nFor example: 10"))
A2.append(A)
X[n] = A2
return X
# + id="lGOjA4cWRJ6n" colab_type="code" colab={}
#this is the solutions vector
def list2vector(length):
X = []
for i in range(length):
a = float(input("add solution vector elements 1by1."))
X.append(a)
return np.array(X)
# + id="mnoPPpO7GrO5" colab_type="code" colab={}
# finally the 2 previouse functions compiled into one to solve your problem
def linear_system_solver_app():
length = int(input("what is the length of your matrix?"))
A = list2array(length)
B = list2vector(length)
a = np.array([0])
if type(A) == type(a) and np.linalg.det(A)!=0:
answers = np.linalg.solve(A,B)
else:
return print("Your Matrix is linear dependant.\nThere is either no solution or infinite solutions")
print("The solutions is " +str(answers))
return answers
# + id="-4e7n0Q_UUAM" colab_type="code" colab={}
linear_system_solver_app()
| LinearSystemsSolver.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={} tags=["awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb", "awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb"]
# <img width="10%" alt="Naas" src="https://landen.imgix.net/jtci2pxwjczr/assets/5ice39g4.png?w=160"/>
# + [markdown] papermill={} tags=["awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb", "awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb"]
# # Google Search - Get LinkedIn company url from name
# <a href="https://app.naas.ai/user-redirect/naas/downloader?url=https://raw.githubusercontent.com/jupyter-naas/awesome-notebooks/master/Google%20Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb" target="_parent"><img src="https://naasai-public.s3.eu-west-3.amazonaws.com/open_in_naas.svg"/></a>
# + [markdown] papermill={} tags=["awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb", "awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb"]
# **Tags:** #googlesearch #snippet #operations #url
# + [markdown] papermill={} tags=["naas", "awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb", "awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb"]
# **Author:** [<NAME>](https://www.linkedin.com/in/ACoAABCNSioBW3YZHc2lBHVG0E_TXYWitQkmwog/)
# + [markdown] papermill={} tags=["awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb", "awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb"]
# ## Input
# + [markdown] papermill={} tags=["awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb", "awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb"]
# ### Import library
# + papermill={} tags=["awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb", "awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb"]
try:
from googlesearch import search
except:
# !pip install google
from googlesearch import search
import re
# + [markdown] papermill={} tags=["awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb", "awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb"]
# ### Variables
# + papermill={} tags=["awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb", "awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb"]
company = "Tesla"
# + [markdown] papermill={} tags=["awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb", "awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb"]
# ## Model
# + [markdown] papermill={} tags=["awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb", "awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb"]
# ### Get LinkedIn url
# + papermill={} tags=["awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb", "awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb"]
def get_linkedin_url(company):
# Init linkedinbio
linkedinbio = None
# Create query
query = f"{company}+Linkedin"
print("Google query: ", query)
# Search in Google
for i in search(query, tld="com", num=10, stop=10, pause=2):
pattern = "https:\/\/.+.linkedin.com\/company\/.([^?])+"
result = re.search(pattern, i)
# Return value if result is not None
if result != None:
linkedinbio = result.group(0).replace(" ", "")
return linkedinbio
return linkedinbio
linkedin_url = get_linkedin_url(company)
# + [markdown] papermill={} tags=["awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb", "awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb"]
# ## Output
# + [markdown] papermill={} tags=["awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb", "awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb"]
# ### Display the result of the search
# + papermill={} tags=["awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb", "awesome-notebooks/Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb"]
linkedin_url
| Google Search/Google_Search_Get_LinkedIn_company_url_from_name.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7
# language: python
# name: python3
# ---
# +
import numpy as np
import scipy.signal
import matplotlib.pyplot as plt
"""
PointEnv from rllab
The goal is to control an agent and get it to the target located at (0,0).
At each timestep the agent gets its current location (x,y) as observation,
takes an action (dx,dy), and is transitioned to (x+dx, y+dy).
"""
class PointEnv():
def reset(self):
self._state = np.random.uniform(-1, 1, size=(2,))
state = np.copy(self._state)
return state
def step(self, action):
action = np.clip(action, -1, 1)
self._state = self._state + 0.1*action
x, y = self._state
reward = -(x**2 + y**2)**0.5 - 0.02*np.sum(action**2)
done = abs(x) < 0.01 and abs(y) < 0.01
next_state = np.copy(self._state)
return next_state, reward, done
class Gauss_Policy():
def __init__(self):
self.action_dim = 2
self.theta = 0.5 * np.ones(4)
# theta here is a length 4 array instead of a matrix for ease of processing
# Think of treating theta as a 2x2 matrix and then flatenning it, which gives us:
# action[0] = state[0]*[theta[0], theta[1]]
# action[1] = state[1]*[theta[2], theta[3]]
def get_action_and_grad(self, state):
# Exercise I.1:
mean_act = np.array([np.dot(self.theta[:2], state), np.dot(self.theta[2:], state)])
sampled_act = mean_act + 0.5 * np.random.randn(self.action_dim)
grad_log_pi = np.ravel([state[0] * (sampled_act - mean_act), state[1] * (sampled_act - mean_act)])
# end
return sampled_act, grad_log_pi
# This function collects some trajectories, given a policy
def gather_paths(env, policy, num_paths, max_ts=100):
paths = []
for i in range(num_paths):
ts = 0
states = []
act = []
grads = []
rwd = []
done = False
s = env.reset()
while not done and ts<max_ts:
a, grad_a = policy.get_action_and_grad(s)
next_s, r, done = env.step(a)
states += [s]
act += [a]
rwd += [r]
grads += [grad_a]
s = next_s
ts += 1
path = {'states': np.array(states),
'actions': np.array(act),
'grad_log_pi': np.array(grads),
'rwd': np.array(rwd)}
paths += [path]
return paths
def baseline(paths):
path_features = []
for path in paths:
s = path["states"]
l = len(path["rwd"])
al = np.arange(l).reshape(-1, 1) / 100.0
path_features += [np.concatenate([s, s ** 2, al, al ** 2, al ** 3, np.ones((l, 1))], axis=1)]
ft = np.concatenate([el for el in path_features])
targets = np.concatenate([el['returns'] for el in paths])
# Exercise I.2(a): Compute the regression coefficents
coeffs = np.linalg.lstsq(ft, targets)[0]
# Exercise I.2(b): Calculate the values for each state
for i, path in enumerate(paths):
path['value'] = np.dot(path_features[i], coeffs)
def process_paths(paths, discount_rate=1):
grads = []
for path in paths:
# Exercise 1.3a: Implement the discounted return
path['returns'] = scipy.signal.lfilter([1], [1, float(-discount_rate)], path['rwd'][::-1], axis=0)[::-1]
# End
baseline(paths)
for path in paths:
#path['value'] = np.zeros(len(path['value']))
path['adv'] = path['returns'] - path['value']
rets_for_grads = np.atleast_2d(path['adv']).T
rets_for_grads = np.repeat(rets_for_grads, path['grad_log_pi'].shape[1], axis=1)
path['grads'] = path['grad_log_pi']*rets_for_grads
grads += [np.sum(path['grads'], axis=0)]
grads = np.sum(grads, axis=0)/len(paths)
return grads
env = PointEnv()
alpha = 0.05
num_itr = 1000
runs = 2
rwd = np.zeros((num_itr, runs))
for st in range(runs):
policy = Gauss_Policy()
# print(st)
for i in range(num_itr):
paths = gather_paths(env, policy, num_paths=5)
rwd[i, st] = np.mean([np.sum(path['rwd']) for path in paths])
grads = process_paths(paths, discount_rate=0.995)
policy.theta += alpha * grads
mean_rwd = np.mean(rwd, axis=1)
sd_rwd = np.std(rwd, axis=1) / np.sqrt(10)
plt.plot(mean_rwd)
plt.fill_between(np.arange(len(mean_rwd)), mean_rwd + sd_rwd, mean_rwd - sd_rwd, alpha=0.3)
plt.ylim([-500, 0])
plt.xlim([0, num_itr])
plt.show()
# -
policy.theta
| Machine Learning Summer School 2019 (Moscow, Russia)/tutorials/reinforcement_learning3/part_i_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Inputs
print('Enter any character:')
i = input()
i
# Input() function always takes input in string format.
int(i)
type(int(i))
print('Enter any number')
i=int(input())
print(i)
i=int(input('Enter Number: '))
print(i,type(i))
i=float(input('Enter number: '))
i
# ### List comprehension
# .split() is very important function and works on a string.
my_string='Himanshu'
my_string.split('a')
my_str='I am Himanshu and you are in bootcamp'
my_str.split(' ')
l=input("Enter number: ")
l.split(' ')
l=[]
for i in range(4):
i=int(input('Enter number: '))
l.append(i)
print(l)
l=[int(i) for i in input('Enter number: ').split()]
print(l)
l=input('Enter number: ').split()
print(l)
# ### Input Dictionary
#1st way
d={}
for i in range(1,4):
k=input('Enter key: ')
l=int(input('Enter numeric val: '))
d.update({k:l})
print(d)
i=[int(i) for i in input('Enter number: ').split()]
j=[int(i) for i in input('Enter number: ').split()]
k=zip(i,j)
d=dict(k)
print(d)
d=eval(input('Enter number: '))
print(d)
| 07-Inputs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Saya sangat jarang nulis di blog tentang COVID-19. Terakhir ngepost tentang stats covid kayaknya waktu bahas soal [kematian](https://www.krisna.or.id/post/covdeath/). Hari ini mau coba lihat lagi statistik COVID-19 di Indonesia, terutama belakangan ini karena lagi ramai lagi soal undertesting.
#
# <blockquote class="twitter-tweet"><p lang="in" dir="ltr">Penambahan kasus Covid-19 harian cenderung menurun. Hal ini terjadi seiring dengan turunnya jumlah pemeriksaan secara signifikan. Masih terlalu dini untuk menyimpulkan bahwa gelombang Covid-19 telah terkendali. <a href="https://twitter.com/hashtag/Humaniora?src=hash&ref_src=twsrc%5Etfw">#Humaniora</a> <a href="https://twitter.com/hashtag/AdadiKompas?src=hash&ref_src=twsrc%5Etfw">#AdadiKompas</a> <a href="https://twitter.com/aik_arif?ref_src=twsrc%5Etfw">@aik_arif</a> <a href="https://t.co/eYvloMmpIC">https://t.co/eYvloMmpIC</a></p>— <NAME> (@hariankompas) <a href="https://twitter.com/hariankompas/status/1417642983227236357?ref_src=twsrc%5Etfw">July 21, 2021</a></blockquote> <script async src="https://platform.twitter.com/widgets.js" charset="utf-8"></script>
#
# Tentu saja andalan saya adalah [Our World in Data](https://ourworldindata.org/coronavirus-source-data#deaths-and-cases-our-data-source) [^1] yang datanya [bisa diakses siapa saja](https://github.com/owid/covid-19-data/tree/master/public/data) dengan mudah.
#
# [^1]: <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME> and <NAME> (2020) - "Coronavirus Pandemic (COVID-19)". Published online at OurWorldInData.org. Retrieved from: 'https://ourworldindata.org/coronavirus' [Online Resource]
#
# Kita tarik dulu datanya dari internet dan menampilkan 10 baris teratas.
url='https://covid.ourworldindata.org/data/owid-covid-data.csv' # simpan url
df=pd.read_csv(url, parse_dates=['date']) # download dari url. parse_dates untuk menjadikan kolom date jadi tipe waktu
df.head(6) # menampilkan 10 baris paling atas
# Berhubung saya jarang banget ngeliatin data COVID-19, saya ga hapal apa aja variabel yang dikumpulkan oleh Ritchie dkk. Panggil dulu semua kolomnya dengan `df.columns`.
df.columns # untuk panggil list dari nama-nama variabel
# Wah gilaaa banyak banget ya nama variabelnya. Cape banget pasti ngumpulin ini semua. Hebat emang Hannah Ritchie dkk. Oke deh sekarang coba kita ngelihat jumlah kasus baru. ada setidaknya tiga nama variabel yang bisa diambil, yaitu `new_cases` dan `new_cases_smoothed`. Kalau dari namanya sih ketaker ya kalo yang smoothed itu kayaknya moving average, alias diambil alusnya dari data `new_cases` yang bisa jadi sangat volatil. Sering terjadi di data harian gini karena, misalnya, setiap hari senin selalu membludak, sementara sabtu minggu selalu sepi. Apa malah kebalik, sabtu minggu malah rame karena orang libur jadi bisa datang ke tempat testing. Yg jelas ada *pattern* mingguan yang bikin data harian jadi volatil. Nah kita ambil dua-duanya yok!
#
# Kalau kamu cuma tertarik ambil data Indonesia, maka jangan lupa diambil yang Indonesia aja.
indo=df[["iso_code","date","new_cases","new_cases_smoothed"]].query('iso_code == "IDN"')
# Saatnya diplot!
sns.lineplot(data=indo,x='date',y='new_cases')
sns.lineplot(data=indo,x='date',y='new_cases_smoothed')
plt.xticks(rotation=45)
# kayaknya new_cases_smoothed ini adalah 7-day rollong average. Seperti yang saya tulis sebelumnya, data harian biasanya punya tren mingguan. Makanya dibuat *rolling average* dalam 7 hari adalah hal yang umum dilakukan. Yok kita cek! Saya pake rolling averagenya panda yang [saya contek di sini](https://datavizpyr.com/how-to-make-a-time-series-plot-with-rolling-average-in-python/) dengan tambahan shift(-3) untuk mundurin NaN 3 hari ke depan dan 3 hari ke belakang.
indo['cases_7day_ave'] = indo.new_cases.rolling(7).mean().shift(-3)
indo.head(10)
# Seperti bisa dilihat di atas, sepertinya bener `new_cases_smoothed` adalah 7-day average soalnya ada 6 NaN berturut-turut. Kayaknya bedanya adalah sama si Ritchie dkk nggak di-shift. ya gpp kita tes plot aja yuk.
sns.lineplot(data=indo,x='date',y='new_cases')
sns.lineplot(data=indo,x='date',y='new_cases_smoothed')
sns.lineplot(data=indo,x='date',y='cases_7day_ave')
plt.xticks(rotation=45)
plt.legend(['new cases','new cases smoothed','7-day average bikinan sendiri'])
plt.ylabel('kasus')
plt.xlabel('tanggal')
# berhubung satu setengah tahun adalah waktu yang panjang (gila lama juga ya covid ga beres-beres), maka kita potong aja deh tahun 2020. Kalo dilihat di atas sih sepertinya first wave yang sebenarnya malah baru keliatna di awal-awal 2021 ya. Oh iya jangan lupa Indonesia juga undertesting kalau dilihat positive rate. Anyway, ayo kita pangkas lagi data kita di `indo` dengan membuah tahun 2020. Oh iya satu lagi, kita pakai aja `new_cases_smoothed` yang dari aslinya ga usah kita buat *rolling average* sendiri.
indo2=indo.query('date>20210101') # ambil hanya setelah 1 Januari 2021
# lalu kita plot persis seperti di atas
sns.lineplot(data=indo2,x='date',y='new_cases')
sns.lineplot(data=indo2,x='date',y='new_cases_smoothed')
plt.xticks(rotation=45)
plt.legend(['new cases','new cases smoothed','7-day average bikinan sendiri'])
plt.ylabel('kasus')
plt.xlabel('tanggal')
# Hmm keliatannya sih emang new cases nya cenderung turun lagi ya. Turunnya dalem juga, bahkan setelah di-*smooth*. Coba sekarang kita bandingkan dengan testing. Jumlah tes baru juga dicatat oleh Our World in Data. mantap banget.
#
# Sekarang kita ulangi ambil datanya, kali ini ambil lebih banyak variabel dari data asli Our World in Data. Sama kita plot tahun 2021 aja.
indo=df[["iso_code","date","new_tests","new_tests_smoothed",
"new_cases","new_cases_smoothed","positive_rate"]].query('iso_code == "IDN"')
indo2=indo.query('date>20210101')
fig, axes = plt.subplots(1, 2, figsize=(18, 10))
fig.suptitle('Data tes baru dan positive rate Indonesia')
sns.lineplot(ax=axes[0],data=indo2,x='date',y='new_tests')
sns.lineplot(ax=axes[0],data=indo2,x='date',y='new_tests_smoothed')
axes[0].tick_params(labelrotation=45)
axes[0].legend(['new tests','new tests smoothed'])
axes[0].set_ylabel('tes baru')
axes[0].set_xlabel('tanggal')
axes[0].set_title('new cases')
sns.lineplot(ax=axes[1],data=indo2,x='date',y='positive_rate')
plt.xticks(rotation=45)
plt.ylabel('0-1')
plt.xlabel('tanggal')
axes[1].set_title('positive rate')
# Memang benar tes-nya berkurang. Tapi di saat yang sama, positive rate juga berkurang di akhir-akhir. Kalau misalnya tes-nya dikurangi tapi kasus di lapangan tetap tinggi, mestinya sih positive rate naik ya. Kecuali ya tes-nya disasar ke orang-orang yang cenderung keliatan ga bergejala.
#
# Sebenernya ini bisa ketawan kalo ternyata yang masuk rumah sakit banyak atau tingkat kematian masih tinggi. Sayangnya data Indonesia untuk hospital dan ICU admissions nggak ada.
df.query('iso_code=="IDN"')[['weekly_icu_admissions','weekly_hosp_admissions']]
# Sementara itu, jika dilihat plot kematian (*Dear God, bless all the lost souls and those who they left*), sepertinya belum ada tanda-tanda berkurang.
indo=df[["iso_code","date","new_deaths","new_deaths_smoothed"]].query('iso_code == "IDN"')
indo2=indo.query('date>20210101')
sns.lineplot(data=indo2,x='date',y='new_deaths')
sns.lineplot(data=indo2,x='date',y='new_deaths_smoothed')
plt.xticks(rotation=45)
plt.legend(['kematian baru','kematian baru rerata bergerak 7 hari'])
plt.ylabel('kasus')
plt.xlabel('tanggal')
# Jika dilihat dari tingkat kematian, sepertinya pandemi masih belum selesai, sayangnya. Perlu dicatat bahwa mungkin tingkat kematian akan turun seiring dengan turunnya kasus, karena bisa jadi tingkat kematian agak mundur dibandingkan tingkat ketahuan positif. Namun demikian, jika tes-nya bermasalah, ada kemungkinan pencatatan kematian juga bermasalah. Sepertinya sih memang kita tidak bisa bereaksi berlebihan, baik ketika kasus naik maupun kasus turun, selama integritas datanya masih dapat dipertanyakan.
#
# Berarti apakah kita bisa bergantung pada vaksinasi? Kalau dilihat dari grafik di atas, kasus baru harian dan positive rate meningkat sejak bulan Juni. Apakah bulan Juni adalah bulan-bulan di mana mulai masuk varian delta? Ada keramaian apa bulan itu? Mobilitas tinggi apa yang dibiarkan di bulan sekitar itu? Mungkin Pemerintah bertaruh membiarkan mobilitas karena program vaksinasi sudah dimulai ya? Kalau begitu, saya akan akhiri postingan kali ini dengan data vaksinasi, dengan komparasi negara-negara lain.
#
# <iframe src="https://ourworldindata.org/grapher/daily-covid-vaccination-doses-per-capita?country=BRA~CHN~IND~LKA~TUR~OWID_WRL~IDN" loading="lazy" style="width: 100%; height: 600px; border: 0px none;"></iframe>
| content/id/post/covidcases/.ipynb_checkpoints/ourworldindata-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#As seen from the results below, this table is unwieldy and difficult to use for our requirments.
#Therefore, this script/notebook contains scripts to convert the source data into a form we like and as described in our reports.
#The data we get from this notebook is used to populate our database
# -
import pandas as pd
import datetime
import numpy as np
# +
# Helper Functions
def dayofweek(day, month, year):
i = "%s-%s-%s" % (year, month, day)
df = pd.Timestamp(i)
return (df.dayofweek+1)%7+1
months = {"January": 1, "February": 2, "March": 3, "April": 4, "May": 5, "June": 6, "July": 7, "August": 8, "September": 9, "October": 10, "November": 11, "December": 12}
daysofweek = {"Sunday": 1, "Monday": 2, "Tuesday": 3, "Wednesday": 4, "Thursday": 5, "Friday": 6, "Saturday": 7}
def ageOut(age):
if ("unknown" == age):
return np.NaN
i = age.split(" ")
if ("over" == i[0].lower()):
return int(i[1])
else:
i = age.split(" ")
return int((int(i[0]) + int(i[2])) / 2)
# -
# Setup the crime table
crime = pd.read_csv('sourcedata/major_crime.csv')
crime = crime[crime['occurrenceday'].notna()]
crime = crime[crime['reportedday'].notna()]
print(crime.columns.tolist())
# Setup the bicycle table
print(bicycleThefts.columns.tolist())
bicycleThefts = pd.read_csv('sourcedata/Bicycle_Thefts.csv')
# Setup the Traffic Table
trafficall = pd.read_csv('sourcedata/Aggressive_Driving.csv')
trafficall.columns.tolist()
Alcohol_Related = pd.read_csv('sourcedata/Alcohol_Related.csv')
Alcohol_Related.columns.tolist()
Automobile = pd.read_csv('sourcedata/Automobile.csv')
Automobile.columns.tolist()
Cyclists = pd.read_csv('sourcedata/Cyclists.csv')
Cyclists.columns.tolist()
Emergency_vehicles = pd.read_csv('sourcedata/Emergency_vehicles.csv')
Emergency_vehicles.columns.tolist()
Passenger = pd.read_csv('sourcedata/Passenger.csv')
Passenger.columns.tolist()
Pedestrians = pd.read_csv('sourcedata/Pedestrians.csv')
Pedestrians.columns.tolist()
Physical_Medical_Disability = pd.read_csv('sourcedata/Physical_Medical_Disability.csv')
Physical_Medical_Disability.columns.tolist()
Red_Light = pd.read_csv('sourcedata/Red_Light.csv')
Red_Light.columns.tolist()
Speeding = pd.read_csv('sourcedata/Speeding.csv')
Speeding.columns.tolist()
Truck = pd.read_csv('sourcedata/Truck.csv')
Truck.columns.tolist()
TTC_Municipal_Vehicle = pd.read_csv('sourcedata/TTC_Municipal_Vehicle.csv')
TTC_Municipal_Vehicle.columns.tolist()
trafficall = trafficall.append(Alcohol_Related)
trafficall = trafficall.append(Automobile)
trafficall = trafficall.append(Cyclists)
trafficall = trafficall.append(Emergency_vehicles)
trafficall = trafficall.append(Passenger,sort=True)
trafficall = trafficall.append(Pedestrians,sort=True)
trafficall = trafficall.append(Physical_Medical_Disability,sort=True)
trafficall = trafficall.append(Red_Light,sort=True)
trafficall = trafficall.append(Speeding,sort=True)
trafficall = trafficall.append(Truck,sort=True)
trafficall = trafficall.append(TTC_Municipal_Vehicle,sort=True)
trafficall.drop_duplicates(inplace=True)
trafficall.to_csv("trafficAll2.csv")
trafficall = trafficall[['ACCNUM', 'DATE', 'ROAD_CLASS', 'TRAFFCTL', 'VISIBILITY', 'RDSFCOND', 'Hood_ID','LATITUDE', 'LONGITUDE' , 'INVAGE', 'INVTYPE', 'INJURY', 'VEHTYPE', 'MANOEUVER']]
trafficall.rename(columns={'ROAD_CLASS': 'classification','TRAFFCTL':'traffic_control_type', 'VISIBILITY':'visibility', 'RDSFCOND':'surface_condition', 'ACCNUM':"accident_id", 'Hood_ID': 'hood_id', 'LATITUDE': 'latitude', 'LONGITUDE':'longitude', 'INVTYPE': 'involvement_type', 'INVAGE': 'age', 'INJURY': 'injury', 'VEHTYPE': 'vehicle_type', 'MANOEUVER': 'action_taken'}, inplace=True)
print(trafficall.columns.tolist())
# Fix the Accident ID
fix = trafficall[['accident_id']]
fix.drop_duplicates(subset = 'accident_id', keep = 'first', inplace = True)
fix.reset_index(drop = True, inplace = True)
fix = fix.rename_axis('accident_id2').reset_index()
fix['accident_id2'] += 1
trafficall = trafficall.merge(fix, on=['accident_id'])
trafficall = trafficall[['accident_id2', 'DATE', 'classification', 'traffic_control_type', 'visibility', 'surface_condition', 'hood_id', 'latitude', 'longitude', 'age', 'involvement_type', 'injury', 'vehicle_type', 'action_taken']]
trafficall.rename(columns={'accident_id2': 'accident_id'}, inplace=True)
print(trafficall)
# +
# Incident Time
# Get the time from Bicycle Thefts in the right form
bicycleThefts.rename(columns={"Occurrence_Time": "hour", "Occurrence_Day": "day", "Occurrence_Month": "month", "Occurrence_Year": "year"}, inplace=True)
bicycleThefts['hour'] = [i.split(':')[0] for i in bicycleThefts['hour']]
bicycleThefts['dayofweek'] = bicycleThefts.apply(lambda row: dayofweek(row.day, row.month, row.year), axis = 1)
bicycleTheft_Time = bicycleThefts[["hour", "day", "month", "year", "dayofweek"]]
bicycleTheft_Time.drop_duplicates(inplace=True)
#print(bicycleTheft_Time)
# Get the time from regular crimes in the right form
crime['occurrencemonth'] = crime.apply(lambda row: months[row.occurrencemonth], axis = 1)
crime['occurrencedayofweek'] = crime['occurrencedayofweek'].str.strip()
crime['occurrencedayofweek'] = crime.apply(lambda row: daysofweek[row.occurrencedayofweek], axis = 1)
crime[['occurrenceday', 'occurrencemonth', 'occurrenceyear', 'occurrencedayofweek']] = crime[['occurrenceday', 'occurrencemonth', 'occurrenceyear', 'occurrencedayofweek']].astype(int)
crime['reportedmonth'] = crime.apply(lambda row: months[row.reportedmonth], axis = 1)
crime['reporteddayofweek'] = crime['reporteddayofweek'].str.strip()
crime['reporteddayofweek'] = crime.apply(lambda row: daysofweek[row.reporteddayofweek], axis = 1)
crime[['reportedday', 'reportedmonth', 'reportedyear', 'reporteddayofweek']] = crime[['reportedday', 'reportedmonth', 'reportedyear', 'reporteddayofweek']].astype(int)
crime_Time = crime[[ "occurrencehour", "occurrenceday","occurrencemonth", "occurrenceyear", "occurrencedayofweek"]]
crime_Time_2 = crime[["reportedhour", "reportedday","reportedmonth", "reportedyear", "reporteddayofweek"]]
crime_Time_2.rename(columns={"reportedhour":"occurrencehour","reportedday":"occurrenceday","reportedmonth":"occurrencemonth","reportedyear":"occurrenceyear","reporteddayofweek":"occurrencedayofweek"}, inplace=True)
crime_Time = crime_Time.append(crime_Time_2)
crime_Time.rename(columns={"occurrencehour":"hour","occurrenceday":"day","occurrencemonth":"month","occurrenceyear":"year","occurrencedayofweek":"dayofweek"}, inplace=True)
crime_Time = crime_Time.drop_duplicates()
crime_Time.drop_duplicates(inplace=True)
#print(crime_Time)
# Get the time from traffic crimes in the right form
trafficall['hour'] = trafficall.apply(lambda row: row.DATE[11:13], axis = 1)
trafficall['day'] = trafficall.apply(lambda row: row.DATE[8:10], axis = 1)
trafficall['month'] = trafficall.apply(lambda row: row.DATE[5:7], axis = 1)
trafficall['year'] = trafficall.apply(lambda row: row.DATE[0:4], axis = 1)
trafficall['dayofweek'] = trafficall.apply(lambda row: dayofweek(row.day, row.month, row.year), axis = 1)
trafficall[['hour', 'day', 'month', 'year','dayofweek']] = trafficall[['hour', 'day', 'month', 'year','dayofweek']].astype(int)
traffic_Time = trafficall[['hour', 'day', 'month', 'year','dayofweek']]
traffic_Time.drop_duplicates(inplace=True)
#print(traffic_Time)
# Output the time tables
IncidentTime = crime_Time.append(bicycleTheft_Time)
IncidentTime = IncidentTime.append(traffic_Time)
IncidentTime[['hour', 'day', 'month', 'year','dayofweek']] = IncidentTime[['hour', 'day', 'month', 'year','dayofweek']].astype(int)
IncidentTime.drop_duplicates(inplace = True)
IncidentTime.reset_index(drop = True, inplace = True)
IncidentTime = IncidentTime.rename_axis('time_id').reset_index()
IncidentTime['time_id'] += 1
IncidentTime.to_csv("IncidentTime.csv", index = False)
print(IncidentTime)
# -
# Output the road condition tables
RoadCondition = trafficall[['classification', 'traffic_control_type', 'visibility', 'surface_condition']]
RoadCondition.drop_duplicates(inplace = True)
RoadCondition.reset_index(drop = True, inplace = True)
RoadCondition = RoadCondition.rename_axis('road_condition_id').reset_index()
RoadCondition['road_condition_id'] += 1
RoadCondition.to_csv("RoadCondition.csv", index = False)
print(RoadCondition)
# Output the traffic event tables
TrafficEvent = trafficall.drop_duplicates(subset = 'accident_id', keep = 'first')
TrafficEvent = TrafficEvent.merge(IncidentTime, on=['hour', 'day', 'month', 'year','dayofweek'])
TrafficEvent = TrafficEvent.merge(RoadCondition, on=['classification', 'traffic_control_type', 'visibility', 'surface_condition'])
TrafficEvent = TrafficEvent[['accident_id', 'time_id', 'road_condition_id', 'hood_id', 'latitude', 'longitude']]
TrafficEvent.drop_duplicates(inplace = True)
TrafficEvent.to_csv("TrafficEvent.csv", index = False)
print(TrafficEvent)
# Output the traffic event tables
InvolvedPerson = trafficall[['accident_id', 'involvement_type', 'age', 'injury', 'vehicle_type', 'action_taken']]
InvolvedPerson['age'] = InvolvedPerson.apply(lambda row: ageOut(row.age), axis = 1)
InvolvedPerson.drop_duplicates(inplace = True)
InvolvedPerson.reset_index(drop = True, inplace = True)
InvolvedPerson = InvolvedPerson.rename_axis('person_id').reset_index()
InvolvedPerson['person_id'] += 1
InvolvedPerson = InvolvedPerson[['accident_id', 'person_id', 'involvement_type', 'age', 'injury', 'vehicle_type', 'action_taken']]
InvolvedPerson.to_csv("InvolvedPerson.csv", index = False)
print(InvolvedPerson)
# At Vikram's Request, see DataParserV3 for his code
bicycleThefts.to_csv("bicyclethefts_inter.csv")
trafficall.to_csv("trafficall_inter.csv")
crime.to_csv("crime_inter.csv")
| milestones/Milestone_2/data/scripts/DataParserV2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python для анализа данных
#
# *<NAME>, НИУ ВШЭ*
#
#
# ## Библиотека pandas. Работа с датами.
#
# Очень часто мы сталкиваемся с переменными, обозначающими время. У работы с ними есть некоторые особенности, которые стоит вынести в отдельный небольшой блокнот. Здесь мы рассмотрим пример на наборе данных о встречах с НЛО.
import pandas as pd
data = pd.read_csv('ufo.csv')
data.head()
# Посмотрим, что лежит в колонке Time.
data['Time'][0]
type(data['Time'][0])
# Можно такую историю распарсить вручную, а можно воспользоваться встроенным модулем datetime.
# трансформация в дату и время
#
# https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.to_datetime.html
data['Time'] = pd.to_datetime(data['Time'])
type(data['Time'][0])
# Видим, что поменялся тип данных в переменной. Теперь посмотрим, что с этим можно делать.
data.head() # видим, что и вид данных поменялся
# Так как это теперь объект timestamp, в его атрибутах зашиты элементы даты и мы можем к ним обращаться через ключевые слова.
data['Time'][0].year
data['Time'][0].month
data['Time'][0].day
data['Time'][0].hour
data['Time'][0].minute
data['Time'][0].second
data['Time'][0].week # номер недели
data['Time'][0].day_name()
# Pandas способен распарсить очень много вариантов написания даты. А если он ломается всегда можно указать ему формат, в котором она записана. Можно составлять дату из нескольких колонок (посмотрим в другом примере).
#
# Подробнее смотрим документацию.
#
# https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.to_datetime.html
#
# Давайте, теперь попробуем использовать нашу новую переменную для аггрегации и фильтрации данных.
#
# У модуля datetime есть атрибут, который превращает нашу переменную в объект Series, заполненную значениями, которые мы указали.
pd.to_datetime("4th of July, 2015")
# И даже так.
pd.to_datetime([pd.datetime(2015, 7, 3), '4th of July, 2015', '2015-Jul-6', '07-07-2015', '20150708'])
pd.to_datetime('13000501', format='%Y%m%d', errors = "ignore")
# Вернемся к тарелочкам.
data['Time'].dt.year.head() # вызвали колонку, обратились к атбриту .dt, выбрали в нем year, получили колонку заполненную годами.
data['Time'].dt.day_name().head()
# А вот такой объект уже можно использовать для группировки. Давайте посчитаем, сколько НЛО видели в разные годы.
# %matplotlib inline
data.groupby(data['Time'].dt.year).size().plot()
data.groupby(data['Time'].dt.day_name()).size().plot(kind = 'bar')
data.groupby(data['Time'].dt.hour).size().plot(kind = 'bar')
# Посмотрим, какие есть города в нашем наборе данных
data['City'].value_counts()[data['City'].value_counts() > 50] # выберем только те, где НЛО видели больше 50 раз
# **В какой день недели чаще всего видели НЛО в Miami?**
data[data['City'] == 'Miami']['Time'].dt.day_name().value_counts().head(1)
data[data['City'] == 'Miami']['Time'].dt.day_name().mode()
# **Какой формы НЛО видели в 1956?**
data[data['Time'].dt.year == 1956].groupby('Shape Reported').size()
data[data['Time'].dt.year == 2000].groupby('Shape Reported').size()
# **В какие месяцы видели НЛО в разные годы?**
data['month'] = data.Time.dt.month
data['Month'] = data.Time.dt.month
data['Year'] = data.Time.dt.year
data.head()
data.groupby([data.Year, data.Month]).size()
data.month
# # какие формы чаще в каком году?
# +
data['Year'] = data.Time.dt.year
data_counts = data[['Year', 'Shape Reported']]
simple_query = '''
Select Year, "Shape Reported" from
(SELECT
Year,
"Shape Reported",
count(*) as cnt
FROM data_counts
Group by Year)
group by Year
Having cnt == MAX(cnt)
'''
tst = ps.sqldf(simple_query, locals())
pd.set_option('display.max_rows', 100)
tst.head(100)
# -
# Так же колонку timestamp можно сделать индексом.
data[data.Year == 1942]
data.index = data['Time']
data.head()
# И индексировать теперь можно поэлементно.
data.loc["2000-06-01"]
data.loc["June 1 2000"] # то же самое
# Можем выбрать целый месяц в году.
data.loc["June 2000"].head()
data.loc["1965"].head() # Или целый год
# На этом пока все, с timestamp обязательно столкнемся в будущем, особенно когда будем смотреть, как работать с временными рядами.
| 01 python/lecture 15 materials/2020_DPO_15_2_Pandas_Datetime.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Animation of the PSF of dancing interferometers
#
# _<NAME>_ Leiden Observatory <EMAIL>
#
# Based on hcipy tutorials https://docs.hcipy.org/0.3.1/tutorials/index.html and examples, and http://louistiao.me/posts/notebooks/embedding-matplotlib-animations-in-jupyter-as-interactive-javascript-widgets/
#
# +
#pip install progressbar2
# +
#pip install ffpmeg
# +
import numpy as np
import matplotlib.pyplot as plt
from hcipy import *
from scenes import *
from progressbar import progressbar
anim_version='v1'
# -
from matplotlib import animation, rc
#from IPython.display import HTML
# +
# each Dish now has a Stage with each variable!
class Dish:
def __init__(self, x=0.0, y=0.0, d=0.1, ampl=1.0):
'Dish - a single element in a telescope array at position (x,y) with diameter d and electric field amplitud ampl'
self.x = x
self.y = y
self.d = d
self.ampl = ampl
self.xsc = Stage()
self.ysc = Stage()
self.dsc = Stage()
self.asc = Stage()
def set_polar(self, r, theta):
'set position of Dish with polar coordinates'
self.x = r*np.cos(theta)
self.y = r*np.sin(theta)
def get_polar(self):
r = np.sqrt(self.x*self.x + self.y*self.y)
theta = np.atan2(self.y, self.x)
return (r, theta)
class TelArray:
def __init__(self):
self.dishes = []
def show_array(self):
fig, ax = plt.subplots(figsize=(6,6))
ax.set_xlim(-0.6,0.6)
ax.set_ylim(-0.6,0.6)
c = plt.Circle((0.0, 0.0), 0.5, fc=None, fill=False, linestyle='--', color='black', zorder=-5)
ax.add_artist(c)
for i, (a) in enumerate(self.dishes):
draw_circle = plt.Circle((a.x,a.y), a.d/2)
ax.add_artist(draw_circle)
ax.text(a.x, a.y, i, fontsize=16, color='white', weight='bold', horizontalalignment='center', verticalalignment='center')
plt.title('Telescope array')
plt.show()
def add(self, Dish):
'add a Dish to the Array'
self.dishes.append(Dish)
return len(self.dishes)
def add_linear(self, n=5, length=1., diam=0.1, ampl=1.0):
'a linear array of n Dishes with centre of first dish to last dish of length, diameter diam, transmission ampl'
xpos = np.linspace(-length/2., length/2., n)
for x in xpos:
dish = Dish(x, 0., diam, ampl)
self.add(dish)
return len(self.dishes)
def add_middle(self, n=5, diam=0.1, ampl=1.0):
'add n Dishes at the origin diameter diam, transmission ampl'
for x in xpos:
dish = Dish(0, 0., diam, ampl)
self.add(dish)
return len(self.dishes)
def make_grid(self):
'returns a HCIpy Grid containing all the dishes in TelArray'
# this seems very daft - surely there's a pythonic way of getting a quantity of a list of Objects?
xd = []
yd = []
diam = []
ampl = []
for dish in self.dishes:
xd.append(dish.x)
yd.append(dish.y)
diam.append(dish.d)
ampl.append(dish.ampl)
print(xd)
interf_array = UnstructuredCoords((xd,yd))
grid = Grid(interf_array)
# currently we keep all dishes the same diameter - the first Diah
dish = circular_aperture(diam[0]) # this is a Field generator
interferom = make_segmented_aperture(dish, grid, np.array(ampl))
return interferom
def longest_time(self):
'''calculates the longest time of all the Stages listed in the TelArray'''
longest_time = 0.
# loop over all the Dishes in the TelArray
for di in self.dishes:
# loop through the Stages in the Dish object
for st in (di.xsc, di.ysc, di.dsc, di.asc):
# find longest time of all the Stages
if (st.total_time() > longest_time):
longest_time = st.total_time()
return longest_time
def t(self, t):
'given a TelArray, makes an HCIpy Grid at SINGLE time t'
print(0)
# WARNING: should barf if number of Dishes is not the same as 4* number of Scenes in the Play
xd = []
yd = []
diam = []
ampl = []
for dish in self.dishes:
xd.append(dish.xsc.t(t))
yd.append(dish.ysc.t(t))
diam.append(dish.dsc.t(t))
ampl.append(dish.asc.t(t))
print(xd)
interf_array = UnstructuredCoords((xd,yd))
grid = Grid(interf_array)
# currently we keep all dishes the same diameter - the first Diah
dish = circular_aperture(diam[0]) # this is a Field generator
interferom = make_segmented_aperture(dish, grid, np.array(ampl))
return interferom
def tidyup(self):
# loop through all Scenes in your Dishes in your TelArray, bring them up to the latest time value
return 0
def tidyup(self):
'''adds constant value Acts to brings up all the other Stages to the longest time'''
# print('longest time is {}'.format(self.longest_time()))
longest_time = self.longest_time()
# loop through all the Dishes
for di in self.dishes:
# loop over all the Stages in each Dish object
for s in (di.xsc, di.ysc, di.dsc, di.asc):
stage_time = s.total_time()
if (stage_time < longest_time):
# print("adding a time delta to this one")
dt = longest_time - stage_time
last_value = s.t(stage_time)
s.add(Act(last_value,last_value,dt))
# you can now build up your animation by adding TelArray[0].xsc.Add(asdf)
# and then run tidyup on your Telarray
arr = TelArray()
print(arr.add_linear(4,1-0.2,0.2))
arr.show_array()
# start with two dishes
interfer = arr.make_grid()
pupil_grid = make_pupil_grid(1024,1.5)
inter_pupil = evaluate_supersampled(interfer, pupil_grid, 8)
imshow_field(inter_pupil)
# -
# +
ndish = 5
ddish = .1
dish_x = np.linspace(-0.5+(ddish/2.), 0.5-(ddish/2.), ndish)
dish_y = np.zeros_like(dish_x)
interf_array = UnstructuredCoords((dish_x,dish_y))
grid = Grid(interf_array)
dish = circular_aperture(ddish) # this is a Field generator
interferom = make_segmented_aperture(dish, grid, np.array([1.0]))
pupil_grid = make_pupil_grid(1024,1.5)
inter_pupil = evaluate_supersampled(interferom, pupil_grid, 8)
imshow_field(inter_pupil)
# +
focal_grid = make_focal_grid(8, 14)
prop = FraunhoferPropagator(inter_pupil, focal_grid)
wf = Wavefront(inter_pupil)
focal_image = prop(wf).intensity
imshow_field(np.log10(focal_image / focal_image.max()), vmin=-5, cmap='inferno')
# -
# ## HCIpy setup
# +
ddish = Stage()
ddish.add(Act(1.0, 0.05, 5, 'sig',10))
ddish.add(Act(0.05, 0.1, 5, 'sig',10))
ddish.add(Act(0.1, 0.2, 5, 'sig',10))
print(ddish.total_time())
t = np.arange(0,ddish.total_time(),0.05)
plt.plot(t, ddish.t(t))
# +
total_time_animation = ddish.total_time() # seconds
frame_rate = 10 # frames per second
total_frames = total_time_animation * frame_rate # seconds
supersampling = 4
plt.figure(figsize=(8,4))
anim = FFMpegWriter('interferometers_{}.mp4'.format(anim_version), framerate=frame_rate)
times = np.arange(0,total_time_animation, 1./frame_rate)
dd = ddish.t(times)
ndish = 5
pupil_grid = make_pupil_grid(1024,1.5)
focal_grid = make_focal_grid(8, 14)
for t in progressbar(range(np.int(total_frames))):
dish = circular_aperture(dd[t]) # this is a Field generator
dish_x = np.linspace(-0.5+(dd[t]/2.), 0.5-(dd[t]/2.), ndish)
dish_y = np.zeros_like(dish_x)
interf_array = UnstructuredCoords((dish_x,dish_y))
grid = Grid(interf_array)
interferom = make_segmented_aperture(dish, grid, np.array([1.0]))
inter_pupil = evaluate_supersampled(interferom, pupil_grid, supersampling)
plt.clf()
# plt.suptitle('Timestep %d / %d' % (t, total_frames))
plt.subplot(1,2,1)
imshow_field(inter_pupil, cmap='gray')
plt.title('Aperture')
plt.text(0.05,0.05,anim_version, fontsize=12, color='white',
horizontalalignment='left', verticalalignment='bottom',
transform=plt.gca().transAxes)
prop = FraunhoferPropagator(inter_pupil, focal_grid)
wf = Wavefront(inter_pupil)
focal_image = prop(wf).intensity
plt.subplot(1,2,2)
imshow_field(np.log10(focal_image / focal_image.max()), vmin=-5, cmap='inferno')
#plt.title('frame {} at time {:.2f}'.format(timestep, timestep/frame_rate))
plt.title('PSF')
plt.text(0.95,0.05,"@mattkenworthy", fontsize=12, color='white',
horizontalalignment='right', verticalalignment='bottom',
transform=plt.gca().transAxes)
anim.add_frame()
plt.close()
anim.close()
# Show created animation
anim
# -
# !ffmpeg -i interferometers_v1.mp4 -vf "fps=10,scale=400:-1:flags=lanczos" -c:v pam -f image2pipe - | convert -delay 10 - -loop 0 -layers optimize interferometers_v1.gif
| anim_interferometer_dance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bayesian Regression
#
# Regression is one of the most common and basic supervised learning tasks in machine learning. Suppose we're given a dataset $\mathcal{D}$ of the form
#
# $$ \mathcal{D} = \{ (X_i, y_i) \} \qquad \text{for}\qquad i=1,2,...,N$$
#
# The goal of linear regression is to fit a function to the data of the form:
#
# $$ y = w X + b + \epsilon $$
#
# where $w$ and $b$ are learnable parameters and $\epsilon$ represents observation noise. Specifically $w$ is a matrix of weights and $b$ is a bias vector.
#
# Let's first implement linear regression in PyTorch and learn point estimates for the parameters $w$ and $b$. Then we'll see how to incorporate uncertainty into our estimates by using Pyro to implement Bayesian linear regression.
# ## Setup
# As always, let's begin by importing the modules we'll need.
# +
import os
import numpy as np
import torch
import torch.nn as nn
import pyro
from pyro.distributions import Normal
from pyro.infer import SVI, Trace_ELBO
from pyro.optim import Adam
# for CI testing
smoke_test = ('CI' in os.environ)
pyro.enable_validation(True)
# -
# ## Data
# We'll generate a toy dataset with one feature and $w = 3$ and $b = 1$ as follows:
# +
N = 100 # size of toy data
def build_linear_dataset(N, p=1, noise_std=0.01):
X = np.random.rand(N, p)
# w = 3
w = 3 * np.ones(p)
# b = 1
y = np.matmul(X, w) + np.repeat(1, N) + np.random.normal(0, noise_std, size=N)
y = y.reshape(N, 1)
X, y = torch.tensor(X).type(torch.Tensor), torch.tensor(y).type(torch.Tensor)
data = torch.cat((X, y), 1)
assert data.shape == (N, p + 1)
return data
# -
# Note that we generate the data with a fixed observation noise $\sigma = 0.1$.
#
# ## Regression
# Now let's define our regression model. We'll use PyTorch's `nn.Module` for this. Our input $X$ is a matrix of size $N \times p$ and our output $y$ is a vector of size $p \times 1$. The function `nn.Linear(p, 1)` defines a linear transformation of the form $Xw + b$ where $w$ is the weight matrix and $b$ is the additive bias. As you can see, we can easily make this a logistic regression by adding a non-linearity in the `forward()` method.
# +
class RegressionModel(nn.Module):
def __init__(self, p):
# p = number of features
super(RegressionModel, self).__init__()
self.linear = nn.Linear(p, 1)
def forward(self, x):
return self.linear(x)
regression_model = RegressionModel(1)
# -
# ## Training
# We will use the mean squared error (MSE) as our loss and Adam as our optimizer. We would like to optimize the parameters of the `regression_model` neural net above. We will use a somewhat large learning rate of `0.01` and run for 500 iterations.
# +
loss_fn = torch.nn.MSELoss(size_average=False)
optim = torch.optim.Adam(regression_model.parameters(), lr=0.05)
num_iterations = 1000 if not smoke_test else 2
def main():
data = build_linear_dataset(N)
x_data = data[:, :-1]
y_data = data[:, -1]
for j in range(num_iterations):
# run the model forward on the data
y_pred = regression_model(x_data).squeeze(-1)
# calculate the mse loss
loss = loss_fn(y_pred, y_data)
# initialize gradients to zero
optim.zero_grad()
# backpropagate
loss.backward()
# take a gradient step
optim.step()
if (j + 1) % 50 == 0:
print("[iteration %04d] loss: %.4f" % (j + 1, loss.item()))
# Inspect learned parameters
print("Learned parameters:")
for name, param in regression_model.named_parameters():
print("%s: %.3f" % (name, param.data.numpy()))
if __name__ == '__main__':
main()
# -
# **Sample Output**:
# ```
# [iteration 0400] loss: 0.0105
# [iteration 0450] loss: 0.0096
# [iteration 0500] loss: 0.0095
# [iteration 0550] loss: 0.0095
# [iteration 0600] loss: 0.0095
# [iteration 0650] loss: 0.0095
# [iteration 0700] loss: 0.0095
# [iteration 0750] loss: 0.0095
# [iteration 0800] loss: 0.0095
# [iteration 0850] loss: 0.0095
# [iteration 0900] loss: 0.0095
# [iteration 0950] loss: 0.0095
# [iteration 1000] loss: 0.0095
# Learned parameters:
# linear.weight: 3.004
# linear.bias: 0.997
# ```
# Not too bad - you can see that the regressor learned parameters that were pretty close to the ground truth of $w = 3, b = 1$. But how confident should we be in these point estimates?
#
# [Bayesian modeling](http://mlg.eng.cam.ac.uk/zoubin/papers/NatureReprint15.pdf) offers a systematic framework for reasoning about model uncertainty. Instead of just learning point estimates, we're going to learn a _distribution_ over values of the parameters $w$ and $b$ that are consistent with the observed data.
# ## Bayesian Regression
#
# In order to make our linear regression Bayesian, we need to put priors on the parameters $w$ and $b$. These are distributions that represent our prior belief about reasonable values for $w$ and $b$ (before observing any data).
#
# ### `random_module()`
#
# In order to do this, we'll 'lift' the parameters $w$ and $b$ to random variables. We can do this in Pyro via `random_module()`, which effectively takes a given `nn.Module` and turns it into a distribution over the same module; in our case, this will be a distribution over regressors. Specifically, each parameter in the original regression model is sampled from the provided prior. This allows us to repurpose vanilla regression models for use in the Bayesian setting. For example:
loc = torch.zeros(1, 1)
scale = torch.ones(1, 1)
# define a unit normal prior
prior = Normal(loc, scale)
# overload the parameters in the regression module with samples from the prior
lifted_module = pyro.random_module("regression_module", regression_model, prior)
# sample a regressor from the prior
sampled_reg_model = lifted_module()
# ### Model
#
# We now have all the ingredients needed to specify our model. First we define priors over $w$ and $b$. Because we're uncertain about the parameters a priori, we'll use relatively wide priors $\mathcal{N}(\mu = 0, \sigma = 10)$. Then we wrap `regression_model` with `random_module` and sample an instance of the regressor, `lifted_reg_model`. We then run the regressor forward on the inputs `x_data`. Finally we use the `obs` argument to the `pyro.sample` statement to condition on the observed data `y_data`. Note that we use the same fixed observation noise that was used to generate the data.
def model(data):
# Create unit normal priors over the parameters
loc, scale = torch.zeros(1, 1), 10 * torch.ones(1, 1)
bias_loc, bias_scale = torch.zeros(1), 10 * torch.ones(1)
w_prior = Normal(loc, scale).independent(1)
b_prior = Normal(bias_loc, bias_scale).independent(1)
priors = {'linear.weight': w_prior, 'linear.bias': b_prior}
# lift module parameters to random variables sampled from the priors
lifted_module = pyro.random_module("module", regression_model, priors)
# sample a regressor (which also samples w and b)
lifted_reg_model = lifted_module()
with pyro.plate("map", N):
x_data = data[:, :-1]
y_data = data[:, -1]
# run the regressor forward conditioned on data
prediction_mean = lifted_reg_model(x_data).squeeze(-1)
# condition on the observed data
pyro.sample("obs",
Normal(prediction_mean, 0.1 * torch.ones(data.size(0))),
obs=y_data)
# ### Guide
#
# In order to do inference we're going to need a guide, i.e. a parameterized family of distributions over $w$ and $b$. Writing down a guide will proceed in close analogy to the construction of our model, with the key difference that the guide parameters need to be trainable. To do this we register the guide parameters in the ParamStore using `pyro.param()`.
# +
softplus = torch.nn.Softplus()
def guide(data):
# define our variational parameters
w_loc = torch.randn(1, 1)
# note that we initialize our scales to be pretty narrow
w_log_sig = torch.tensor(-3.0 * torch.ones(1, 1) + 0.05 * torch.randn(1, 1))
b_loc = torch.randn(1)
b_log_sig = torch.tensor(-3.0 * torch.ones(1) + 0.05 * torch.randn(1))
# register learnable params in the param store
mw_param = pyro.param("guide_mean_weight", w_loc)
sw_param = softplus(pyro.param("guide_log_scale_weight", w_log_sig))
mb_param = pyro.param("guide_mean_bias", b_loc)
sb_param = softplus(pyro.param("guide_log_scale_bias", b_log_sig))
# guide distributions for w and b
w_dist = Normal(mw_param, sw_param).independent(1)
b_dist = Normal(mb_param, sb_param).independent(1)
dists = {'linear.weight': w_dist, 'linear.bias': b_dist}
# overload the parameters in the module with random samples
# from the guide distributions
lifted_module = pyro.random_module("module", regression_model, dists)
# sample a regressor (which also samples w and b)
return lifted_module()
# -
# Note that we choose Gaussians for both guide distributions. Also, to ensure positivity, we pass each log scale through a `softplus()` transformation (an alternative to ensure positivity would be an `exp()`-transformation).
#
# ## Inference
#
# To do inference we'll use stochastic variational inference (SVI) (for an introduction to SVI, see [SVI Part I](svi_part_i.ipynb)). Just like in the non-Bayesian linear regression, each iteration of our training loop will take a gradient step, with the difference that in this case, we'll use the ELBO objective instead of the MSE loss by constructing a `Trace_ELBO` object that we pass to `SVI`.
optim = Adam({"lr": 0.05})
svi = SVI(model, guide, optim, loss=Trace_ELBO())
# Here `Adam` is a thin wrapper around `torch.optim.Adam` (see [here](svi_part_i.ipynb#Optimizers) for a discussion). The complete training loop is as follows:
# +
def main():
pyro.clear_param_store()
data = build_linear_dataset(N)
for j in range(num_iterations):
# calculate the loss and take a gradient step
loss = svi.step(data)
if j % 100 == 0:
print("[iteration %04d] loss: %.4f" % (j + 1, loss / float(N)))
if __name__ == '__main__':
main()
# -
# To take an ELBO gradient step we simply call the `step` method of `SVI`. Notice that the `data` argument we pass to `step` will be passed to both `model()` and `guide()`.
#
# ## Validating Results
# Let's compare the variational parameters we learned to our previous result:
for name in pyro.get_param_store().get_all_param_names():
print("[%s]: %.3f" % (name, pyro.param(name).data.numpy()))
# **Sample Output**:
# ```
# [guide_log_scale_weight]: -3.217
# [guide_log_scale_bias]: -3.164
# [guide_mean_weight]: 2.966
# [guide_mean_bias]: 0.941
# ```
# As you can see, the means of our parameter estimates are pretty close to the values we previously learned. Now, however, instead of just point estimates, the parameters `guide_log_scale_weight` and `guide_log_scale_bias` provide us with uncertainty estimates. (Note that the scales are in log-space here, so the more negative the value, the narrower the width).
# Finally, let's evaluate our model by checking its predictive accuracy on new test data. This is known as _point evaluation_. We'll sample 20 neural nets from our posterior, run them on the test data, then average across their predictions and calculate the MSE of the predicted values compared to the ground truth.
X = np.linspace(6, 7, num=20)
y = 3 * X + 1
X, y = X.reshape((20, 1)), y.reshape((20, 1))
x_data, y_data = torch.tensor(X).type(torch.Tensor), torch.tensor(y).type(torch.Tensor)
loss = nn.MSELoss()
y_preds = torch.zeros(20, 1)
for i in range(20):
# guide does not require the data
sampled_reg_model = guide(None)
# run the regression model and add prediction to total
y_preds = y_preds + sampled_reg_model(x_data)
# take the average of the predictions
y_preds = y_preds / 20
print("Loss: ", loss(y_preds, y_data).item())
# **Sample Output**:
# ```
# Loss: 0.00025596367777325213
# ```
# See the full code on [Github](https://github.com/uber/pyro/blob/dev/examples/bayesian_regression.py).
| tutorial/source/bayesian_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Puzzle 1: Find the oxygen system using a droid and intcode
# ### status: 0 --> wall; 1 --> moved 1 step; 2 --> moved 1 step, found oxygen system; movements: north (1), south (2), west (3), east (4)
import numpy as np
import matplotlib.pyplot as plt
import networkx as nx
from copy import deepcopy
# ## Load input
with open('./input15.txt', 'r') as file:
software = file.readlines()
# #### Convert data to list of integers
software = list(map(int, software[0].split(',')))
# ## Calculation
class intcode_computer(object):
def __init__(self, data):
self.data = data
self.data += [0]*100000
self.inputs = []
self.i = 0
self.relative_base = 0
self.done = False
def step(self, input):
self.run = True
self.out = []
input = [input]
while self.run is True:
add = (5-len(str(self.data[self.i])))*'0'
self.data[self.i] = '{0}{1}'.format(add, str(self.data[self.i]))
optcode = self.data[self.i][-2:]
mode1 = self.data[self.i][-3]
mode2 = self.data[self.i][-4]
mode3 = self.data[self.i][-5]
if mode1 == '0' and optcode != '99':
param1 = int(self.data[self.data[self.i+1]])
elif mode1 == '1' and optcode != '99':
param1 = int(self.data[self.i+1])
elif mode1 == '2' and optcode != '99':
param1 = int(self.data[self.data[self.i+1]+self.relative_base])
if optcode == '01':
if mode2 == '0':
param2 = int(self.data[self.data[self.i+2]])
elif mode2 == '1':
param2 = int(self.data[self.i+2])
elif mode2 == '2':
param2 = int(self.data[self.data[self.i+2]+self.relative_base])
if mode3 == '0' or mode3 == '1':
self.data[self.data[self.i+3]] = param1 + param2
elif mode3 == '2':
self.data[self.data[self.i+3]+self.relative_base] = param1 + param2
self.i += 4
if optcode == '02':
if mode2 == '0':
param2 = int(self.data[self.data[self.i+2]])
elif mode2 == '1':
param2 = int(self.data[self.i+2])
elif mode2 == '2':
param2 = int(self.data[self.data[self.i+2]+self.relative_base])
if mode3 == '0' or mode3 == '1':
self.data[self.data[self.i+3]] = param1 * param2
elif mode3 == '2':
self.data[self.data[self.i+3]+self.relative_base] = param1 * param2
self.i += 4
if optcode == '03':
if len(input) > 0:
if mode1 == '0' or mode1 == '1':
self.data[self.data[self.i+1]] = input[0]
elif mode1 == '2':
self.data[self.data[self.i+1]+self.relative_base] = input[0]
del input[0]
self.i += 2
else:
self.run = False
# print('Waiting for input.')
break
if optcode == '04':
if mode1 == '0':
self.out.append(self.data[self.data[self.i+1]])
# print(self.out)
if mode1 == '1':
self.out.append(self.data[self.i+1])
# print(self.out)
elif mode1 == '2':
self.out.append(self.data[self.data[self.i+1]+self.relative_base])
# print(self.out)
self.i += 2
if optcode == '05':
if mode2 == '0':
param2 = int(self.data[self.data[self.i+2]])
elif mode2 == '1':
param2 = int(self.data[self.i+2])
elif mode2 == '2':
param2 = int(self.data[self.data[self.i+2]+self.relative_base])
if param1 != 0:
self.i = param2
else:
self.i += 3
if optcode == '06':
if mode2 == '0':
param2 = int(self.data[self.data[self.i+2]])
elif mode2 == '1':
param2 = int(self.data[self.i+2])
elif mode2 == '2':
param2 = int(self.data[self.data[self.i+2]+self.relative_base])
if param1 == 0:
self.i = param2
else:
self.i += 3
if optcode == '07':
if mode2 == '0':
param2 = int(self.data[self.data[self.i+2]])
elif mode2 == '1':
param2 = int(self.data[self.i+2])
elif mode2 == '2':
param2 = int(self.data[self.data[self.i+2]+self.relative_base])
if param1 < param2:
if mode3 == '0' or mode3 == '1':
self.data[self.data[self.i+3]] = 1
elif mode3 =='2':
self.data[self.data[self.i+3]+self.relative_base] = 1
else:
if mode3 == '0' or mode3 == '1':
self.data[self.data[self.i+3]] = 0
elif mode3 == '2':
self.data[self.data[self.i+3]+self.relative_base] = 0
self.i += 4
if optcode == '08':
if mode2 == '0':
param2 = int(self.data[self.data[self.i+2]])
elif mode2 == '1':
param2 = int(self.data[self.i+2])
if mode2 == '2':
param2 = int(self.data[self.data[self.i+2]+self.relative_base])
if param1 == param2:
if mode3 == '0' or mode3 == '1':
self.data[self.data[self.i+3]] = 1
elif mode3 == '2':
self.data[self.data[self.i+3]+self.relative_base] = 1
else:
if mode3 == '0' or mode3 == '1':
self.data[self.data[self.i+3]] = 0
elif mode3 == '2':
self.data[self.data[self.i+3]+self.relative_base] = 0
self.i += 4
if optcode == '09':
self.relative_base += param1
self.i += 2
if optcode == '99':
self.run = False
self.done = True
self.i += 1
return self.out
class repair_droid(object):
def __init__(self):
self.moveset = {'1': (0, -1), '2': (0, 1), '3': (-1, 0), '4': (1, 0)}
self.status = {'0': 'w', '1': 'm', '2': 'o'}
self.control = 0
self.state = (0, 0)
self.environment = nx.Graph()
self.environment.add_node(self.state)
self.oxygen_state = ()
def step(self, new_status, action):
current_status = self.status['{0}'.format(new_status[0])]
if current_status == 'w':
pass
elif current_status =='m':
new_state = (self.state[0] + self.moveset['{0}'.format(action)][0],
self.state[1] + self.moveset['{0}'.format(action)][1])
self.environment.add_edge(self.state, new_state)
self.state = new_state
elif current_status == 'o':
new_state = (self.state[0] + self.moveset['{0}'.format(action)][0],
self.state[1] + self.moveset['{0}'.format(action)][1])
self.environment.add_edge(self.state, new_state)
self.state = new_state
self.oxygen_state = new_state
return current_status
def path_to_oxygen(self):
path = nx.shortest_path(self.environment, (0, 0), self.oxygen_state)
path_length = len(path)-1
return path, path_length
def flood_with_oxygen(self):
time = nx.eccentricity(self.environment, v=self.oxygen_state)
return time
# +
computer = intcode_computer(deepcopy(software))
droid = repair_droid()
status = False
while status != 'o':
random_action = np.random.randint(1, 5)
output = computer.step(random_action)
status = droid.step(output, random_action)
path, path_length = droid.path_to_oxygen()
print('The shortest way to the path requires {0} steps.'.format(path_length))
# -
# # Puzzle 2: Time required to fill map with oxygen
time = droid.flood_with_oxygen()
print('{0} minutes are required to fill the map with oxygen.'.format(time))
| day15/Script15.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="HhR5048dZ3e1" colab_type="text"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab_type="code" id="f0A2utIXbPc5" colab={}
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="1qF0JETfbfIR" colab_type="text"
# # Mandelbrot set
# + [markdown] id="p8Z8Pb5nbtZ3" colab_type="text"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/non-ml/mandelbrot"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/non-ml/mandelbrot.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/non-ml/mandelbrot.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] id="lqPLlJWqcSFZ" colab_type="text"
# Visualizing the [Mandelbrot set](https://en.wikipedia.org/wiki/Mandelbrot_set) doesn't have anything to do with machine learning, but it makes for a fun example of how one can use TensorFlow for general mathematics. This is actually a pretty naive implementation of the visualization, but it makes the point. (We may end up providing a more elaborate implementation down the line to produce more truly beautiful images.)
# + [markdown] id="80RrFh7EcnLT" colab_type="text"
# ## Basic setup
#
# You'll need a few imports to get started.
# + id="xc-QSV_SdEG4" colab_type="code" colab={}
from __future__ import absolute_import, division, print_function, unicode_literals
# Import libraries for simulation
import tensorflow as tf
import numpy as np
# Imports for visualization
import PIL.Image
from io import BytesIO
from IPython.display import clear_output, Image, display
# + [markdown] id="mP5YEOuTieH0" colab_type="text"
#
# Now you'll define a function to actually display the image once you have iteration counts.
# + id="_q_HC5cGhX4h" colab_type="code" colab={}
def DisplayFractal(a, fmt='jpeg'):
"""Display an array of iteration counts as a
colorful picture of a fractal."""
a_cyclic = (6.28*a/20.0).reshape(list(a.shape)+[1])
img = np.concatenate([10+20*np.cos(a_cyclic),
30+50*np.sin(a_cyclic),
155-80*np.cos(a_cyclic)], 2)
img[a==a.max()] = 0
a = img
a = np.uint8(np.clip(a, 0, 255))
f = BytesIO()
PIL.Image.fromarray(a).save(f, fmt)
display(Image(data=f.getvalue()))
# + [markdown] id="xEptO88QikEM" colab_type="text"
# # Session and variable initialization
#
# For playing around like this, an interactive session is often used, but a regular session would work as well.
# + id="8_yDY6Uih7bD" colab_type="code" colab={}
sess = tf.InteractiveSession()
# + [markdown] id="_NFwmNL5iqBd" colab_type="text"
#
# It's handy that you can freely mix NumPy and TensorFlow.
# + id="fHu_sT7chbg_" colab_type="code" colab={}
# Use NumPy to create a 2D array of complex numbers
Y, X = np.mgrid[-1.3:1.3:0.005, -2:1:0.005]
Z = X+1j*Y
# + [markdown] id="u7SsqtHqivVW" colab_type="text"
#
# Now you define and initialize TensorFlow tensors.
# + id="UpGYdAWQhhCN" colab_type="code" colab={}
xs = tf.constant(Z.astype(np.complex64))
zs = tf.Variable(xs)
ns = tf.Variable(tf.zeros_like(xs, tf.float32))
# + [markdown] id="gqvhBLXbi4al" colab_type="text"
#
# TensorFlow requires that you explicitly initialize variables before using them.
# + id="RmjN39LHhob2" colab_type="code" colab={}
tf.global_variables_initializer().run()
# + [markdown] id="ao_esnw4jAJp" colab_type="text"
# # Defining and running the computation
#
# Now you specify more of the computation...
# + id="ZMup0FHjiGEx" colab_type="code" colab={}
# Compute the new values of z: z^2 + x
zs_ = zs*zs + xs
# Have we diverged with this new value?
not_diverged = tf.abs(zs_) < 4
# Operation to update the zs and the iteration count.
#
# Note: We keep computing zs after they diverge! This
# is very wasteful! There are better, if a little
# less simple, ways to do this.
#
step = tf.group(
zs.assign(zs_),
ns.assign_add(tf.cast(not_diverged, tf.float32))
)
# + [markdown] id="9qqqbNu7jCrj" colab_type="text"
#
# ... and run it for a couple hundred steps
# + id="twC_FiUSiN8s" colab_type="code" colab={}
for i in range(200): step.run()
# + [markdown] id="vfoDAWtijLKd" colab_type="text"
#
# Let's see what you've got.
# + id="8qqfdbuOiV90" colab_type="code" colab={}
DisplayFractal(ns.eval())
# + [markdown] id="vB-3S5cFjVYQ" colab_type="text"
# Not bad!
| site/en/tutorials/non-ml/mandelbrot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="../img/logo_white_bkg_small.png" align="right" />
#
# # Automate it All! - Answers
# This worksheet covers concepts relating to automating a machine learning model using the techniques we learned. It should take no more than 20-30 minutes to complete. Please raise your hand if you get stuck.
# Load Libraries - Make sure to run this cell!
import pandas as pd
from sklearn.model_selection import train_test_split
import scikitplot as skplt
from sklearn.metrics import classification_report
import matplotlib.pyplot as plt
import lime
from tpot import TPOTClassifier
# %matplotlib inline
# ## Step One: Import the Data
# In this example, we're going to use the dataset we used in worksheet 5.3. Run the following code to read in the data, extract the features and target vector.
df = pd.read_csv('../data/dga_features_final_df.csv')
target = df_final['isDGA']
feature_matrix = df_final.drop(['isDGA'], axis=1)
# Next, perform the test/train split in the conventional manner.
feature_matrix_train, feature_matrix_test, target_train, target_test = train_test_split(feature_matrix,
target,
test_size=0.25)
# ## Step Two: Run the Optimizer
# In the next step, use TPOT to create a classification pipeline using the DGA data set that we have been using. The `TPOTClassifier()` has many configuration options and in the interest of time, please set the following variables when you instantiate the classifier.
#
# * `max_eval_time_mins`: In the interests of time, set this to 15 or 20.
# * `verbosity`: Set to 1 or 2 so you can see what TPOT is doing.
#
#
# **Note: This step will take some time, so you might want to get some coffee or a snack when it is running.** While this is running take a look at the other configuration options available here: http://epistasislab.github.io/tpot/api/.
# Your code here...
optimizer = TPOTClassifier(n_jobs=-1, verbosity=3)
optimizer.fit(feature_matrix_train, target_train)
# ## Step Three: Evaluate the Performance
# Now that you have a trained model, the next step is to evaluate the performance and see how TPOT did in comparison with earlier models we created. Use the techniques you've learned to evaluate the performance of your model. Specifically, print out the `classification report` and a confusion matrix.
#
# Unfortunately, Yellowbrick will not work in this instance, however, you can generate a similar visual confusion matrix with the following code:
#
# ```
# import scikitplot as skplt
# skplt.metrics.confusion_matrix(optimized_preds, target_test)
#
# ```
#
# What is the accuracy of your model? Is it significantly better than what you did in earlier labs?
predictions = optimizer.predict(feature_matrix_test)
print(classification_report(predictions, target_test))
skplt.metrics.plot_confusion_matrix(optimized_preds, target_test)
# ## Step 4: Export your Pipeline
# If you are happy with the results from `TPOT` you can export the pipeline as python code. The final step in this lab is to export the pipeline as a file called `automate_ml.py` and examine it. What model and preprocessing steps did TPOT find? Was this a surprise?
optimizer.export('automate_ml.py')
| answers/Worksheet 5.4 - Automate it All! - Answers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 多维标度法(MDS)
# 如,给定城市间距,要求画出地理地图
# +
# 课本例10.20实现
import numpy as np
D = np.array(
[[0, 1, np.sqrt(3), 2, np.sqrt(3), 1, 1],
[0, 0, 1, np.sqrt(3), 2, np.sqrt(3), 1],
[0, 0, 0, 1, np.sqrt(3), 2, 1],
[0, 0, 0, 0, 1, np.sqrt(3), 1],
[0, 0, 0, 0, 0, 1, 1],
[0, 0, 0, 0, 0, 0, 1],
[0, 0, 0, 0, 0, 0, 0]],
dtype=np.float)
D = D + D.T
A = -D*D/2
H = np.eye(A.shape[0]) - np.ones(A.shape)/A.shape[0]
B = np.matmul(H, np.matmul(A, H))
eigenvalues, eigenvectors = np.linalg.eig(B)
print('特征值: ', eigenvalues)
print('特征向量: ', eigenvectors)
# 取最大两个
trans = eigenvectors[[1,3]]
coordinate = np.matmul(D, trans.T)
print('变换到二维坐标: ', coordinate)
| .ipynb_checkpoints/10.7 多维标度法-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://maltem.com/wp-content/uploads/2020/04/LOGO_MALTEM.png" style="float: left; margin: 20px; height: 55px">
#
# <br>
# <br>
# <br>
# <br>
#
#
# # Intro to Decision Trees
# Authors: <NAME>, <NAME>
#
# <img src="../images/sms1.png" align="left">
# ### Learning Objectives
#
# - Understand the intuition behind decision trees.
# - Calculate Gini.
# - Describe how decision trees use Gini to make decisions.
# - Fit, generate predictions from, and evaluate decision tree models.
# - Interpret and tune `max_depth`, `min_samples_split`, `min_samples_leaf`, `ccp_alpha`.
# - Visualize a decision tree.
# ## What will we get for dinner?
#
# |$Y = $ Food|$X_1 = $ Weather|$X_2 = $ Day|
# |:---------:|:--------------:|:----------:|
# | Indian | Rainy | Weekday |
# | Sushi | Sunny | Weekday |
# | Indian | Rainy | Weekend |
# | Mexican | Sunny | Weekend |
# | Indian | Rainy | Weekday |
# | Mexican | Sunny | Weekend |
# <details><summary>It's a rainy day. Based on our past orders, what do you think we'll order?</summary>
#
# - Indian food.
# - In 100% of past cases where the weather is rainy, we've eaten Indian food!
#
# |$Y = $ Food|$X_1 = $ Weather|$X_2 = $ Day|
# |:---------:|:--------------:|:----------:|
# | Indian | Rainy | Weekday |
# | Indian | Rainy | Weekend |
# | Indian | Rainy | Weekday |
#
# </details>
# <details><summary>It's a sunny day. Based on our past orders, what do you think we'll order?</summary>
#
# - Either Sushi or Mexican food... but we can't say with certainty whether we'd eat sushi or Mexican food.
# - Based on our past orders, we eat sushi on 1/3 of sunny days and we eat Mexican food on 2/3 of sunny days.
# - If I **had** to make a guess here, I'd probably predict Mexican food, but we may want to use additional information to be certain.
#
# |$Y = $ Food|$X_1 = $ Weather|$X_2 = $ Day|
# |:---------:|:--------------:|:----------:|
# | Sushi | Sunny | Weekday |
# | Mexican | Sunny | Weekend |
# | Mexican | Sunny | Weekend |
#
# </details>
# <details><summary>It's a sunny day that also happens to be a weekend. Based on our past orders, what do you think we'll order?</summary>
#
# - Mexican food.
# - In 100% of past cases where the weather is sunny and where it's a weekend, we've eaten Mexican food!
#
# |$Y = $ Food|$X_1 = $ Weather|$X_2 = $ Day|
# |:---------:|:--------------:|:----------:|
# | Mexican | Sunny | Weekend |
# | Mexican | Sunny | Weekend |
#
# </details>
# # Decision Trees: Overview
#
# A decision tree:
# - takes a dataset consisting of $X$ and $Y$ data,
# - finds rules based on our $X$ data that partitions (splits) our data into smaller datasets such that
# - by the bottom of the tree, the values $Y$ in each "leaf node" are as "pure" as possible.
#
# We frequently see decision trees represented by a graph.
#
# <img src="../images/order_food_dt.png" alt="order_food" width="750"/>
#
# - (This image was created using [Draw.io](https://www.draw.io/).)
#
# ### Terminology
# Decision trees look like upside down trees.
# - What we see on top is known as the "root node," through which all of our observations are passed.
# - At each internal split, our dataset is partitioned.
# - A "parent" node is split into two or more "child" nodes.
# - At each of the "leaf nodes" (colored orange), we contain a subset of records that are as pure as possible.
# - In this food example, each leaf node is perfectly pure. Once we get to a leaf node, every observation in that leaf node has the exact same value of $Y$!
# - There are ways to quantify the idea of "purity" here so that we can let our computer do most of the tree-building (model-fitting) process... we'll come back to this later.
#
# Decision trees are also called "**Classification and Regression Trees**," sometimes abbreviated "**CART**."
# - [DecisionTreeClassifier Documentation](https://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeClassifier.html)
# - [DecisionTreeRegressor Documentation](https://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeRegressor.html)
# ## 20 Questions
#
# If you aren't familiar with the game [20 Questions](https://en.wikipedia.org/wiki/Twenty_Questions), it's a game with two players (or teams).
# - Player A thinks of an item but doesn't say what the item is.
# - Player B then attempts to guess what the item is by asking a series of 20 questions with a yes or no answer.
# - If player B correctly guesses the item, then player B wins!
# - If player B does not correctly guess the item, then player A wins!
#
# Let's play a quick game of "5 Questions" to get a feel for it.
#
# ---
#
#
# #### Decision trees operate in a fashion that's pretty similar to 20 Questions.
# - Decisions are made in a sequential fashion. Once you know a piece of information, you use that piece of information when asking future questions.
# - Example: If you know that the item you're trying to guess is a person, then you can use that information to ask better subsequent questions.
# - It's possible to get lucky by making very specific guesses early, but it's pretty unlikely that this is a winning strategy.
# - Example: If you asked, "Is it an airplane? Is it a boat? Is it a car?" as your first three questions, it's not very likely that you'll win the game.
#
# When fitting a decision tree, we're effectively getting our computer to play a game of 20 Questions. We give the computer some data and it figures out the best $X$ variable to split on at the right time.
# - Above, our "what food should we order?" decision tree first asked what the weather was, **then** asked whether it was a weekday or weekend.
# - If we had asked "is it a weekday or weekend" first, we'd have ended up with a slightly more complicated decision tree.
#
# Just like with all of our models, in order for the computer to learn which $X$ variable to split on and when, the computer needs a loss function to quantify how good a particular split is. This is where the idea of **purity** comes into play.
# ## Purity in Decision Trees
#
# When quantifying how "pure" a node is, we want to see what the distribution of $Y$ is in each node, then summarize this distribution with a number.
#
# <img src="./images/order_food_dt.png" alt="order_food" width="750"/>
#
# - For continuous $Y$ (i.e. using a decision tree to predict income), the default option is mean squared error.
# - This is the `criterion = 'mse'` argument in `DecisionTreeRegressor`.
# - When the decision tree is figuring out which split to make at a given node, it picks the split that maximizes the drop in MSE from the parent node to the child node.
#
# - For discrete $Y$, the default option is the Gini impurity. *(Bonus: This is not quite the same thing as the [Gini coefficient](https://en.wikipedia.org/wiki/Gini_coefficient).)*
#
# ---
#
# The **Gini impurity** is the probability a randomly chosen class will be mislabeled if it was randomly labeled.
#
# Suppose $p_i$ is the probability that class $i$ would be chosen uniformly at random. Then:
#
# $$
# \begin{eqnarray*}
# \text{Gini impurity} &=& \sum_{i=1}^{classes} p_i(1 - p_i) \\
# &=& 1 - \sum_{i=1}^{classes} p_i^2. \\
# \text{Gini impurity (2 classes)} &=& 1 - p_1^2 - p_2^2. \\
# \text{Gini impurity (3 classes)} &=& 1 - p_1^2 - p_2^2 - p_3^2. \\
# \end{eqnarray*}
# $$
# Define Gini function, called gini.
def gini(obs):
# Create a list to store my squared class probabilities.
gini_sum = []
# Iterate through each class.
for class_i in set(obs):
# Calculate observed probability of class i.
prob = (obs.count(class_i) / len(obs))
# Square the probability and append it to gini_sum.
gini_sum.append(prob ** 2)
# Return Gini impurity.
return 1 - sum(gini_sum)
# Create our y variable from our "where should we eat" dataframe.
y = ['Indian', 'Sushi', 'Indian', 'Mexican', 'Indian', 'Mexican']
# <details><summary>This is a classification problem. How many classes do we have, and what are they?</summary>
#
# - 3 classes: Indian, Sushi, Mexican.
# </details>
# Check to see if your Gini function is correct on the
# "where should we eat" data. (Should get 0.6111.)
gini(y)
# ### Gini Practice
#
# $$\text{Gini impurity} = 1 - \sum_{i=1}^{classes} p_i^2$$
# <details><summary>What is the Gini impurity of a node when every item is from the same class?</summary>
#
# - Our Gini impurity is zero.
#
# $$
# \begin{eqnarray*}
# \text{Gini impurity} &=& 1 - \sum_{i=1}^{classes} p_i^2 \\
# &=& 1 - p_1^2 \\
# &=& 1 - 1^2 \\
# &=& 1 - 1 \\
# &=& 0
# \end{eqnarray*}
# $$
# </details>
# What is Gini when every item is from the same class?
gini(['Indian', 'Indian', 'Indian'])
# <details><summary>What is the Gini impurity of a node when we have two classes, each with two items?</summary>
#
# - Our Gini impurity is 0.5.
#
# $$
# \begin{eqnarray*}
# \text{Gini impurity} &=& 1 - \sum_{i=1}^{classes} p_i^2 \\
# &=& 1 - p_1^2 - p_2^2 \\
# &=& 1 - \left(\frac{1}{2}\right)^2 - \left(\frac{1}{2}\right)^2 \\
# &=& 1 - \frac{1}{4} - \frac{1}{4} \\
# &=& \frac{1}{2}
# \end{eqnarray*}
# $$
# </details>
# What is Gini when we have two classes, each with two items?
gini(['Indian', 'Indian', 'Mexican', 'Mexican'])
# <details><summary>What is the Gini impurity of a node when we have two classes, each with three items?</summary>
#
# - Our Gini impurity is 0.5.
#
# $$
# \begin{eqnarray*}
# \text{Gini impurity} &=& 1 - \sum_{i=1}^{classes} p_i^2 \\
# &=& 1 - p_1^2 - p_2^2 \\
# &=& 1 - \left(\frac{1}{2}\right)^2 - \left(\frac{1}{2}\right)^2 \\
# &=& 1 - \frac{1}{4} - \frac{1}{4} \\
# &=& \frac{1}{2}
# \end{eqnarray*}
# $$
# </details>
# What is Gini when we have two classes, each with three items?
gini(['Indian', 'Indian', 'Mexican', 'Mexican', 'Indian', 'Mexican'])
# <details><summary>What is the Gini impurity of a node when we have three classes, each with two items?</summary>
#
# - Our Gini impurity is 0.6667.
#
# $$
# \begin{eqnarray*}
# \text{Gini impurity} &=& 1 - \sum_{i=1}^{classes} p_i^2 \\
# &=& 1 - p_1^2 - p_2^2 - p_3^2 \\
# &=& 1 - \left(\frac{1}{3}\right)^2 - \left(\frac{1}{3}\right)^2 - \left(\frac{1}{3}\right)^2 \\
# &=& 1 - \frac{1}{9} - \frac{1}{9} - \frac{1}{9} \\
# &=& 1 - \frac{1}{3} \\
# &=& \frac{2}{3}
# \end{eqnarray*}
# $$
# What is Gini when we have three classes, each with two items?
gini(['Indian', 'Indian', 'Mexican', 'Mexican', 'Sushi', 'Sushi'])
# <details><summary>Summary of Gini Impurity Scores</summary>
#
# - A Gini score of 0 means all of our observations are from the same class!
# - In the binary case, Gini impurity ranges from 0 to 0.5.
# - If we have three classes, Gini impurity ranges from 0 to 0.66667.
# - If we have $k$ classes, Gini impurity ranges from 0 to $1-\frac{1}{k}$.
# </details>
# +
# Import libraries.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# Set figure size.
plt.figure(figsize = (12,8))
# Generate x values (for percentage of obs. in class A).
percent_in_class_A = np.linspace(0, 1, 200)
percent_in_class_B = 1 - percent_in_class_A
# Calculate Gini values.
gini_values = 1 - np.square(percent_in_class_A) - np.square(percent_in_class_B)
# Plot line.
plt.plot(percent_in_class_A,
gini_values)
# Establish title, axes, and labels.
plt.title('Gini Score in Binary Classification', fontsize = 24)
plt.xlabel('Percent of Observation in Class A', fontsize = 20)
plt.ylabel('Gini Score', fontsize = 20, rotation = 0, ha = 'right')
plt.xticks(fontsize = 18)
plt.yticks(fontsize = 18);
# -
# ### So how does a decision tree use Gini to decide which variable to split on?
#
# - At any node, consider the subset of our dataframe that exists at that node.
# - Iterate through each variable that could potentially split the data.
# - Calculate the Gini impurity for every possible split.
# - Select the variable that decreases Gini impurity the most from the parent node to the child node.
#
# One consequence of this is that a decision tree is fit using a **greedy** algorithm. Simply put, a decision tree makes the best short-term decision by optimizing at each node individually. _This might mean that our tree isn't optimal (in the number of nodes) in the long run!_
# ## Building a Decision Tree
# +
# Read in Titanic data.
titanic = pd.read_csv('../datasets/titanic_clean.csv')
# Change sex to int.
titanic['Sex'] = titanic['Sex'].map({'male':0,
'female':1})
# Create embarked_S column.
titanic['Embarked_s'] = titanic['Embarked'].map({'S':1,
'C':0,
'Q':0})
# Create embarked_C column.
titanic['Embarked_c'] = titanic['Embarked'].map({'S':0,
'C':1,
'Q':0})
# Conduct train/test split.
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(titanic.drop(['Survived','PassengerId','Name','Embarked'], axis=1),
titanic['Survived'],
test_size = 0.3,
random_state = 42)
# -
# Check out first five rows of X_train.
X_train.head()
y_train.head()
# Import model.
from sklearn.tree import DecisionTreeClassifier
# Instantiate model with random_state = 42.
dt = DecisionTreeClassifier(random_state = 42)
# Fit model.
dt.fit(X_train, y_train)
# Evaluate model.
print(f'Score on training set: {dt.score(X_train, y_train)}')
print(f'Score on testing set: {dt.score(X_test, y_test)}')
# <details><summary>What conclusion would you make here?</summary>
#
# - Our model is **very** overfit to the data.
# </details>
# When fitting a decision tree, your model will always grow until it nearly perfectly predicts every observation!
# - This is like playing a game of 20 questions, but instead calling it "Infinite Questions." You're always going to be able to win!
# <details><summary>Intuitively, what might you try to do to solve this problem?</summary>
#
# - As with all models, try to gather more data.
# - As with all models, remove some features.
# - Is there a way for us to stop our model from growing? (Yes!)
# </details>
# ### Hyperparameters of Decision Trees
# There are four hyperparameters of decision trees that we may commonly tune in order to prevent overfitting.
#
# - `max_depth`: The maximum depth of the tree.
# - By default, the nodes are expanded until all leaves are pure (or some other argument limits the growth of the tree).
# - In the 20 questions analogy, this is like "How many questions we can ask?"
#
#
# - `min_samples_split`: The minimum number of samples required to split an internal node.
# - By default, the minimum number of samples required to split is 2. That is, if there are two or more observations in a node and if we haven't already achieved maximum purity, we can split it!
#
#
# - `min_samples_leaf`: The minimum number of samples required to be in a leaf node (a terminal node at the end of the tree).
# - By default, the minimum number of samples required in a leaf node is 1. (This should ring alarm bells - it's very possible that we'll overfit our model to the data!)
#
#
# - `ccp_alpha`: A [complexity parameter](https://scikit-learn.org/stable/modules/tree.html#minimal-cost-complexity-pruning) similar to $\alpha$ in regularization. As `ccp_alpha` increases, we regularize more.
# - By default, this value is 0.
#
# [Source: Documentation](https://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeClassifier.html).
# +
# Instantiate model with:
# - a maximum depth of 5.
# - at least 7 samples required in order to split an internal node.
# - at least 3 samples in each leaf node.
# - a cost complexity of 0.01.
# - random state of 42.
dt = DecisionTreeClassifier(max_depth = 5,
min_samples_split = 7,
min_samples_leaf = 3,
ccp_alpha = 0.01,
random_state = 42)
# -
# Fit model.
dt.fit(X_train, y_train)
# Evaluate model.
print(f'Score on training set: {dt.score(X_train, y_train)}')
print(f'Score on testing set: {dt.score(X_test, y_test)}')
# #### Let's GridSearch to try to find a better tree.
#
# - Check [2, 3, 5, 7] for `max_depth`.
# - Check [5, 10, 15, 20] for `min_samples_split`.
# - Check [2, 3, 4, 5, 6] for `min_samples_leaf`.
# - Check [0, 0.001, 0.01, 0.1, 1, 10] for `ccp_alpha`.
# - Run 5-fold cross-validation.
# <details><summary>How many models are being fit here?</summary>
#
# - 4 * 4 * 5 * 6 * 5 = 2400 models.
# </details>
from sklearn.model_selection import GridSearchCV
grid = GridSearchCV(estimator = DecisionTreeClassifier(),
param_grid = {'max_depth': [2, 3, 5, 7],
'min_samples_split': [5, 10, 15, 20],
'min_samples_leaf': [2, 3, 4, 5, 6],
'ccp_alpha': [0, 0.001, 0.01, 0.1, 1, 10]},
cv = 5,
verbose = 1)
# +
import time
# Start our timer.
t0 = time.time()
# Let's GridSearch over the above parameters on our training data.
grid.fit(X_train, y_train)
# Stop our timer and print the result.
print(time.time() - t0)
# -
# What is our best decision tree?
grid.best_estimator_
# What was the cross-validated score of the above decision tree?
grid.best_score_
# Evaluate model.
print(f'Score on training set: {grid.score(X_train, y_train)}')
print(f'Score on testing set: {grid.score(X_test, y_test)}')
# Generate predictions on test set.
preds = grid.predict(X_test)
# Import confusion_matrix.
from sklearn.metrics import confusion_matrix
# +
# Generate confusion matrix.
tn, fp, fn, tp = confusion_matrix(y_test,
preds).ravel()
print(confusion_matrix(y_test,
preds))
# +
# Calculate sensitivity.
sens = tp / (tp + fn)
print(f'Sensitivity: {round(sens, 4)}')
# +
# Calculate specificity.
spec = tn / (tn + fp)
print(f'Specificity: {round(spec, 4)}')
# -
# ## Visualizing the Output of Decision Trees
#
# One advantage to using a decision tree is that you can easily visualize them in `sklearn`. The two functions used to do this are `plot_tree` and `export_text`.
# - [`plot_tree` documentation](https://scikit-learn.org/stable/modules/generated/sklearn.tree.plot_tree.html#sklearn.tree.plot_tree)
# - [`export_text` documentation](https://scikit-learn.org/stable/modules/generated/sklearn.tree.export_text.html)
# +
# Import plot_tree from sklearn.tree module.
from sklearn.tree import plot_tree
# Establish size of figure.
plt.figure(figsize = (50, 30))
# Plot our tree.
plot_tree(grid.best_estimator_,
feature_names = X_train.columns,
class_names = ['Dead', 'Survived'],
filled = True);
# +
# Import export_text from sklearn.tree module.
from sklearn.tree import export_text
# Print out tree in plaintext.
print(export_text(grid.best_estimator_,
feature_names=list(X_train.columns)))
# -
# As with all visualizations, just because we _can_ doesn't mean that we _should_. If our depth is much larger than 2 or 3, the tree may be unreadable.
#
# While these visuals may be helpful to us, it may be helpful to clean it up before presenting it to stakeholders.
# ## Why use a decision tree?
#
#
# ### 1. We don't have to scale our data.
# The scale of our inputs don't affect decision trees.
#
# ### 2. Decision trees don't make assumptions about how our data is distributed.
# Is our data heavily skewed or not normally distributed? Decision trees are nonparametric, meaning we don't make assumptions about how our data or errors are distributed.
#
# ### 3. Easy to interpret.
# The output of a decision tree is easy to interpret and thus are relatable to non-technical people. (We'll talk about `feature_importance` later.)
#
# ### 4. Speed.
# Decision trees fit very quickly!
# ## Why not use a decision tree?
#
#
# ### 1. Decision trees can very easily overfit.
# Decision trees often suffer from high error due to variance, so we need to take special care to avoid this. (There are lots of algorithms designed to do exactly this!)
#
# ### 2. Decision trees are locally optimal.
# Because we're making the best decision at each node (greedy), we might end up with a worse solution in the long run.
#
# ### 3. Decision trees don't work well with unbalanced data.
# We often will bias our results toward the majority class. We need to take steps to avoid this as well! (Check out the `class_weight` parameter if you're interested.)
# ## Interview Question
# <details><summary>If you're comparing decision trees and logistic regression, what are the pros and cons of each?</summary>
#
# (Answers may vary; this is not an exhaustive list!)
# - **Interpretability**: The coefficients in a logistic regression model are interpretable. (They represent the change in log-odds caused by the input variables.) However, this is complicated and not easy for non-technical audiences. Decision trees are interpretable; it is easy to explain to show a picture of a decision tree to a client or boss and get them to understand how predictions are made.
# - **Performance**: Decision trees have a tendency to easily overfit, while logistic regression models usually do not overfit as easily.
# - **Assumptions**: Decision trees have no assumptions about how data are distributed; logistic regression does make assumptions about how data are distributed.
# - **Frequency**: Logistic regression is more commonly than decision trees.
# - **Y variable**: Decision trees can handle regression and classification problems; logistic regression is only really used for classification problems.
# </details>
| Notebook/Lesson-decision-trees/solution-code/solution-code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py35]
# language: python
# name: conda-env-py35-py
# ---
# ### Fun with Mazes
#
# Simple maze representation and solvers.
#
# See [The LeetCode example problem](https://leetcode.com/problems/trim-a-binary-search-tree/description/)
# +
debugging = False
debugging = True
logging = True
def dprint(f, *args):
if debugging:
print((' DBG:' + f).format(*args))
def log(f, *args):
if logging:
print((f).format(*args))
def logError(f, *args):
if logging:
print(('*** ERROR:' + f).format(*args))
def className(instance):
return type(instance).__name__
# -
class Maze(object):
""" Representation of a maze with narrow walls """
class MazeCell(object):
def __init__():
west_wall = False # A wall is present (on the west side)
north_wall = False # A wall is present (above to the north)
state = None # Nothing interesting has happened in this cell.
pass
m = Maze()
# +
class MazeCell(object):
def __init__(cell):
cell.blocked = False
cell.count = 0
cell.visited = 0
class BlockMaze(object):
""" Representation of a maze with walls that take up entire cells """
def __init__(self):
self.maze = []
def loadMaze(self, mrows):
assert len(mrows) > 0 and isinstance(mrows, list)
for s in mrows:
row = []
for c in s:
cell = MazeCell()
cell.blocked = True if c == 'X' else False
row.append(cell)
self.maze.append(row)
def printMaze(self):
rc = len(self.maze)
cc = len(self.maze[0])
def pe(v):
print(v,end='')
def printTopBot():
pe('+-')
for c in range(cc): pe('---')
print('-+')
printTopBot()
for row in self.maze:
pe('| ')
for c in row:
txt = '(X)' if c.blocked else ' '
pe(txt)
print(' |')
printTopBot()
# -
x = BlockMaze()
x.loadMaze([" X X ",
" X X ",
" X ",
" X ",
" X X ",
" X X ",
" X X X ",
" X X "])
x.printMaze()
x
| maze.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SimpleQA
#
# ## References
# [Network Structure explanation](http://edu.dataguru.cn/thread-643001-1-1.html)
#
# [Dataset](https://github.com/shuzi/insuranceQA)
import tensorflow as tf
import numpy
def Model():
def __init__(seqLength,batchSize=5,vecDim,hlSize)
with tf.name_scope("ModelScope"):
self.inputQ=tf.placeholder(dtype=tf.float32,shape=[batchSize,seqLength,vecDim],name="Q")
self.inputA=tf.placeholder(dtype=tf.float32,shape=[batchSize,seqLength,vecDim],name="A")
W_HL=tf.get_variable(dtype=tf.float32,shape=[vecDim,hlSize])
Q=tf.matmul(self.inputQ,W_HL)#[batchSize,seqLength,hlSize]
A=tf.matmul(self.inputQ,W_HL)#[batchSize,seqLength,hlSize]
#Add a dimension for channel
#[batch, in_height, in_width, in_channels]
Q=tf.reshape(tensor=Q,shape=[batchSize,seqLength,hlSize,1])
A=tf.reshape(tensor=A,shape=[batchSize,seqLength,hlSize,1])
tf.nn.conv2d()
| SimpleQA/SimpleQA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python Wikipedia Library User Guide
# ## <NAME>
# ## Table of Contents
# - [Search](#search)
# - [Summary](#summary)
# - [Page](#page)
# - [Additional](#additional)
# import libraries
import wikipedia as wiki
# ## Search <a class="anchor" id="search"></a>
# .search() method finds article titles containing a term
print(wiki.search("Bill"))
# limit results of search
print(wiki.search("Bill", results = 2))
# increase results of search
print(wiki.search("Bill", results = 100))
# may need to use GeoData extension to work
# https://www.mediawiki.org/wiki/Extension:GeoData
print(wiki.geosearch(38.9, 77.0, results = 10, radius = 10000))
# ## Summary <a class="anchor" id="summary"></a>
# .summary() method shows summary of requested article
print(wiki.summary('Kill Bill'))
# limit summary by number of sentences
print(wiki.summary('Tiger Woods', sentences = 3))
# summary will raise 'DisambiguationError' if:
# requested page doesn't exist or,
# request is disambiguous
print(wiki.summary('key'))
# to get summary of a specific 'key' article
print(wiki.summary('key (cryptography)'))
# ## Page <a class="anchor" id="Page"></a>
# wiki.page() to create a WikipediaPage object
# contains contents, categories, coordinates, images, links,
# and other metadata from article page
wiki.page('Animal Crossing')
# wiki.page().content to get all plain text of page
# excludes images, tables, links, etc.
print(wiki.page('Animal Crossing').content)
# wiki.page().url to get page URL
print(wiki.page('Animal Crossing').url)
# wiki.page().references to get URLS to external links
print(wiki.page('Animal Crossing').references)
# wiki.page().title to get title of page
print(wiki.page('Animal Crossing New Leaf').title)
# wiki.page().categories to get list of categories of page
print(wiki.page('Animal Crossing New Leaf').categories)
# wiki.page().links to get alphabetical list of links within article page
print(wiki.page('Animal Crossing New Leaf').links)
# get coordinates of article when relevant
print(wiki.page('White House').coordinates)
# use wiki.page(lat, long) to bring up articles related to the location
print(wiki.page(37.787, -122.4))
# wiki.page().images[] to pull image urls from wiki page
print(wiki.page('Animal Crossing').images[0])
# wiki.page().html() to get full article page in html
print(wiki.page('Animal Crossing').html())
# wiki.page().section() returns plain text of a section of page
print(wiki.page('Animal Crossing').section('Characters'))
# unique ID for the current version of the page
print(wiki.page('Animal Crossing').revision_id)
# ISSUE this should return a list of sections on the page
wiki.page('Animal Crossing').sections
# ## Additional <a class="anchor" id="additional"></a>
# returns a random list of wikipedia articles
# doesn't get Category, Usertalk, or other meta-wikipedia pages
# max ten
print(wiki.random(pages = 5))
# wiki.set_lang to use other language article
# uses standard prefix codes:
# https://en.wikipedia.org/wiki/List_of_ISO_639-1_codes
wiki.set_lang('fr')
print(wiki.summary('Animal Crossing', sentences = 2))
# reset language
wiki.set_lang('en')
print(wiki.summary('Animal Crossing', sentences = 2))
# use try and except to handle DisambiguationError possibilities
try:
print(wiki.summary('Mercury'))
except wiki.exceptions.DisambiguationError as e:
print(e.options)
# ### Exceptions
# - `wiki.exceptions.DisambiguationError` - raised when page resolves to a disambiguation page
# - `wiki.exceptions.HTTPTimeoutError` - raised when a request to the Mediawiki servers times out
# - `wiki.exceptions.PageError` - raised when no Wikipedia matched a query
# - `wiki.exceptions.RedirectError` - raised when a page title unexpectedly resolves to a redirect
# - `wiki.exceptions.WikipediaException` - base exception class
# opens link to donation page on wikipedia
wiki.donate()
| Python_Wikipedia_Library_User_Guide.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_mxnet_p36)
# language: python
# name: conda_mxnet_p36
# ---
# %matplotlib inline
from __future__ import print_function
import numpy as np
import mxnet as mx
import matplotlib.pyplot as plt
from mxnet import nd, autograd, gluon, optimizer
from mxnet.image import color_normalize
mx.random.seed(1)
mx.__version__
# ## Set the context
ctx = mx.gpu()
# ## Load our data
# +
#Binary Classifier
num_outputs = 1
batch_size = 64
train_data = mx.io.ImageRecordIter(path_imgrec='lego_train.rec',
min_img_size=512,
data_shape=(3, 512, 512),
rand_crop=False,
shuffle=True,
batch_size=batch_size,
max_rotation=15,
rand_mirror=True)
test_data = mx.io.ImageRecordIter(path_imgrec='lego_val.rec',
min_img_size=512,
data_shape=(3, 512, 512),
batch_size=batch_size)
train_data.reset()
test_data.reset()
# -
batch = train_data.next()
data = batch.data[0]
print(data.shape)
for i in range(4):
plt.subplot(1,4,i+1)
plt.imshow(data[i].asnumpy().astype(np.uint8).transpose((1,2,0)))
# ## Define a convolutional neural network
#
# Again, a few lines here is all we need in order to change the model. Let's add a couple of convolutional layers using gluon.nn.
#
#
net = gluon.nn.Sequential()
with net.name_scope():
# First convolutional layer
net.add(gluon.nn.Conv2D(channels=96, kernel_size=11, strides=(4,4), activation='relu'))
net.add(gluon.nn.MaxPool2D(pool_size=3, strides=2))
# Second convolutional layer
net.add(gluon.nn.Conv2D(channels=192, kernel_size=5, activation='relu'))
net.add(gluon.nn.MaxPool2D(pool_size=3, strides=(2,2)))
# Third convolutional layer
net.add(gluon.nn.Conv2D(channels=384, kernel_size=3, activation='relu'))
# Fourth convolutional layer
net.add(gluon.nn.Conv2D(channels=384, kernel_size=3, activation='relu'))
# Fifth convolutional layer
net.add(gluon.nn.Conv2D(channels=256, kernel_size=3, activation='relu'))
net.add(gluon.nn.MaxPool2D(pool_size=3, strides=2))
# Flatten and apply fullly connected layers
net.add(gluon.nn.Flatten())
net.add(gluon.nn.Dense(4096, activation="relu"))
net.add(gluon.nn.Dense(4096, activation="relu"))
net.add(gluon.nn.Dense(num_outputs, activation="sigmoid"))
# ## Parameter Initialization
net.collect_params().initialize(mx.init.Xavier(magnitude=2.24), ctx=ctx)
# ## Loss Function
sigmoid_binary_cross_entropy = gluon.loss.SigmoidBinaryCrossEntropyLoss(from_sigmoid=True)
logisitc_loss = gluon.loss.LogisticLoss(label_format='binary')
# ## Optimizer
trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': 0.0001})
# ## Accuracy Evaluation
# +
metric = mx.metric.create(['rmse'])
def evaluate(net, data_iter, ctx):
data_iter.reset()
for batch in data_iter:
data = gluon.utils.split_and_load(batch.data[0], ctx_list=ctx, batch_axis=0)
label = gluon.utils.split_and_load(batch.label[0], ctx_list=ctx, batch_axis=0)
outputs = []
for x in data:
outputs.append(net(x))
metric.update(label, outputs)
out = metric.get()
metric.reset()
return out
# -
# ## Training Loop
# +
epochs = 100
smoothing_constant = .01
moving_loss = 0
train_data.reset()
for e in range(epochs):
train_data.reset()
for i, batch in enumerate(train_data):
data = gluon.utils.split_and_load(nd.array(batch.data[0]), ctx_list=[ctx])
label = gluon.utils.split_and_load(nd.array(batch.label[0]), ctx_list=[ctx])
with autograd.record():
for x, y in zip(data, label):
output = net(x)
loss = sigmoid_binary_cross_entropy(output, y)
loss.backward()
trainer.step(64)
##########################
# Keep a moving average of the losses
##########################
curr_loss = nd.mean(loss).asscalar()
moving_loss = (curr_loss if ((i == 0) and (e == 0))
else (1 - smoothing_constant) * moving_loss + smoothing_constant * curr_loss)
test_accuracy = evaluate(net, test_data, [ctx])
train_accuracy = evaluate(net, train_data, [ctx])
print("Epoch %s. Loss: %s, Train_acc %s, Test_acc %s" % (e, moving_loss, train_accuracy, test_accuracy))
# -
# ## Time to Test!
# +
import cv2
def get_image(url, show=False):
# download and show the image
fname = mx.test_utils.download(url)
img = cv2.cvtColor(cv2.imread(fname), cv2.COLOR_BGR2RGB)
if img is None:
return None
if show:
plt.imshow(img)
plt.axis('off')
# convert into format (batch, RGB, width, height)
img = cv2.resize(img, (512, 512))
img = np.swapaxes(img, 0, 2)
img = np.swapaxes(img, 1, 2)
img = img[np.newaxis, :]
return img
# -
brick = get_image('https://27gen.files.wordpress.com/2013/06/lego-red-brick.jpg', True)
with mx.Context(ctx):
print(net(nd.array(brick)))
minifigure = get_image('https://i.ebayimg.com/images/g/cOQAAOxyA9dSZp5~/s-l1600.jpg', True)
with mx.Context(ctx):
print(net(nd.array(minifigure)))
| basic_convnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import scipy.stats as scs
import matplotlib.pyplot as plt
import pandas as pd
# %matplotlib inline
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = 'all'
#
# - Find the fuel cell's rack ID, which is its X coordinate plus 10.
# - Begin with a power level of the rack ID times the Y coordinate.
# - Increase the power level by the value of the grid serial number (your puzzle input).
# - Set the power level to itself multiplied by the rack ID.
# - Keep only the hundreds digit of the power level (so 12345 becomes 3; numbers with no hundreds digit become 0).
# - Subtract 5 from the power level.
# For example, to find the power level of the fuel cell at 3,5 in a grid with serial number 8:
#
# - The rack ID is 3 + 10 = 13.
# - The power level starts at 13 * 5 = 65.
# - Adding the serial number produces 65 + 8 = 73.
# - Multiplying by the rack ID produces 73 * 13 = 949.
# - The hundreds digit of 949 is 9.
# - Subtracting 5 produces 9 - 5 = 4.
# - So, the power level of this fuel cell is 4.
#
# grid serial number = 8979
def construct_power_grid(grid_serial_number):
"""
"""
grid = np.zeros((300, 300))
for x_idx in range(300):
x = x_idx + 1
rack_id = x + 10
for y_idx in range(300):
y = y_idx + 1
power_level = rack_id * y
power_level += grid_serial_number
power_level = power_level * rack_id
power_level = int(str(power_level)[-3])
power_level -= 5
grid[y_idx, x_idx] = power_level
return grid
# +
def find_highest_3x3(grid):
"""
"""
y_s = 0
y_e = 3
x_s = 0
x_e = 3
highest_sum = 0
top_left = 0, 0
for y_i in range(298):
for x_i in range(298):
three_grid = grid[y_s+y_i:y_e+y_i, x_s+x_i:x_e+x_i]
three_sum = three_grid.sum()
if three_sum > highest_sum:
highest_sum = three_sum
top_left = (x_s+x_i+1, y_s+y_i+1)
return top_left
# -
find_highest_3x3(grid)
grid = construct_power_grid(8979)
def find_highest_square(grid):
"""
"""
highest_sum = 0
top_left = 0, 0
top_dim = 0
for dim in range(1, 301):
if dim % 10 == 0:
print("Calculating {} dimensions".format(dim))
for y_i in range(301-dim):
for x_i in range(301-dim):
square = grid[y_i:y_i+dim, x_i:x_i+dim]
square_sum = square.sum()
if square_sum > highest_sum:
highest_sum = square_sum
top_left = (x_i+1, y_i+1)
top_dim = dim
return top_left, top_dim
find_highest_square(grid)
| day_11.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Especificação
#
# O objetivo deste projeto é construir um classificador de Spam usando o algoritmo de classificação Naive Bayes.
#
# O modelo de documento que usaremos aqui é um modelo de saco de palavras(bag of words). Usaremos o modelo bag of words:
#
# - Com base na frequência de palavras (frequência de ocorrência de palavra no documento, o que tornará os atributos de entrada contínuos)
# ### Modelo Bag of Words
#
# Um saco de palavras (bag of words) é uma representação de um texto como um agrupamento de palavras, sem qualquer consideração da sua estrutura gramatical ou da ordem das palavras. É simplesmente um histograma sobre as palavras da língua, e cada documento é representado como um vetor sobre estas palavras. As entradas neste vetor simplesmente correspondem à presença ou à ausência da palavra correspondente.
# A seguir criaremos uma função para ler o conjunto de dados, verificar a frequência de palavras e então ajustar o modelo.
#Importando as bibliotecas
import os
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.metrics import confusion_matrix
from sklearn.naive_bayes import MultinomialNB
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.metrics import accuracy_score, precision_score
sns.set()
warnings.filterwarnings('ignore')
# Exemplo simples.
#Cria uma matix cujas linhas são frases, alocadas em uma única coluna
documents = ['Hello, how are you!',
'Win money, win from home.',
'Call me now.',
'Hello, Call hello you tomorrow?']
# Para obter a frequência de cada palavra em cada frase (linha), será utilizado a função CountVectorizer() do scikit learning.
# +
count_vector = CountVectorizer() #set the variable
count_vector.fit(documents) #fit the function
count_vector.get_feature_names() #get the outputs
doc_array = count_vector.transform(documents).toarray()
doc_array
frequency_matrix = pd.DataFrame(doc_array,
columns = count_vector.get_feature_names()
)
frequency_matrix
# -
# ### PART 1: DATA PREPROCESSING
#
#
# #### Preparando o conjunto de dados
# Temos que definir onde estão os diretórios do conjunto de treino e teste.
# Em seguida temos que ler todos os e mails e armazená-los em uma lista, será armazenado também em outra lista o valor do e mail (spam ou notspam) e se pertence ao conjunto de trino ou de teste.
files_path='Dados\\train' # Local onde se encontram os dados
traintest=[]
documents=[]
my_label=[]
dirs = os.listdir(files_path)
for class_dir_name in dirs:
print ("Processando datasset de teino {}. Aguarde...".format(class_dir_name))
for f in os.listdir(os.path.join(files_path, class_dir_name)):
document = os.path.join(files_path, class_dir_name, f)
with open(document, 'r', encoding = "latin1") as file:
words = file.read()
# .split()
documents.append(words)
my_label.append(class_dir_name)
traintest.append('train')
print ("O processo foi finalizado com sucesso ...")
files_path='Dados\\test' # Local onde se encontram os dados
dirs = os.listdir(files_path)
for class_dir_name in dirs:
print ("Processando {}. Seja paciente e aguarde...".format(class_dir_name))
for f in os.listdir(os.path.join(files_path, class_dir_name)):
document = os.path.join(files_path, class_dir_name, f)
with open(document, 'r', encoding = "latin1") as file:
words = file.read()
# .split()
documents.append(words)
my_label.append(class_dir_name)
traintest.append('test')
print ("O processo foi finalizado com sucesso ...")
# A seguir será criado um dataframe contendo a classificação na primera colulna e o e-mail na segunda. Iremos também substituir as strings notspam e spam por 0 e 1 respectivamente
Dict = {'Dividir':traintest,'labels': my_label, 'mensagem': documents}
df=pd.DataFrame(Dict)
df['labels'] = df.labels.map({'notspam':0, 'spam':1})
df.head()
# Preparando os dados com a função CountVectorizer
count_vector = CountVectorizer() #set the variable
dataset_transformado = count_vector.fit_transform(documents)
# #### Dividindo o datasset em trinamento e teste.
# indices do datassest de traino e de teste
slice_indices=np.arange(df.shape[0])
slice_train=slice_indices[df.Dividir=='train']
slice_test=slice_indices[df.Dividir=='test']
X_train=dataset_transformado[slice_train,:]
X_test=dataset_transformado[slice_test,:]
y_train=df.labels[df.Dividir=='train']
y_test=df.labels[df.Dividir=='test']
# ### PART 2: AJUSTANDO O MODELO
naive_bayes = MultinomialNB() #call the method
naive_bayes.fit(X_train, y_train) #train the classifier on the training set
predictions = naive_bayes.predict(X_train) #predic using the model on the testing set
print('accuracy score: {}'.format(accuracy_score(y_train,predictions)))
print('precision score: {}'.format(precision_score(y_train,predictions)))
# ### PART 3: AVALIANDO O MODELO
# Para avaliar o modelo utilizaremos uma Validação cruzada k-Fold.
# +
# Aplicando validação cruzada de 5 folds
mnb = MultinomialNB()
scores = cross_val_score(mnb, X_train, y_train, cv = 5, scoring='accuracy')
print('Cross-validation scores:{}'.format(scores))
# +
# cross_val_score?
# +
# computar Pontuação média de validação cruzada
print('Pontuação média de validação cruzada: {:.4f}'.format(scores.mean()))
# -
# ### PART 4: FAZENDO PREVISÕES
# Com o modelo ajustado, é necessário testar sua capacidade de predizer valores de mensagens que ele ainda não tenha visto. Assim, ireos carregar o arquivo de teste e utilizar o modelo ajustado para predizer os valores dos e-mails. E então comparar com seu real valor.
# #### Predizendo os valores dos dados de teste.
predictions = naive_bayes.predict(X_test) #predic using the model on the testing set
print('accuracy score: {}'.format(accuracy_score(y_test,predictions)))
print('precision score: {}'.format(precision_score(y_test,predictions)))
#confusion_matrix
cnf_matrix = confusion_matrix(y_test,predictions)
p = sns.heatmap(pd.DataFrame(cnf_matrix), annot=True, cmap="YlGnBu" ,fmt='g',yticklabels=['notspam','spam'],xticklabels=['notspam','spam'])
plt.title('Confusion matrix', y=1.1)
plt.ylabel('True label')
plt.xlabel('Predicted label')
p=sns.heatmap(cnf_matrix/np.sum(cnf_matrix), annot=True, fmt='.2%', cmap="YlGnBu" ,yticklabels=['notspam','spam'],xticklabels=['notspam','spam'])
plt.title('Confusion matrix', y=1.1)
plt.ylabel('True label')
plt.xlabel('Predicted label')
| Classificador_Completo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import spacy
import re
from scipy import stats
from gensim.models import Phrases, LdaModel, CoherenceModel, Word2Vec
from gensim.models.word2vec import LineSentence
from gensim.corpora import Dictionary, MmCorpus
from gensim.test.utils import datapath
import pyLDAvis
import pyLDAvis.gensim
from ast import literal_eval
# -
# Import model from disc
lda_model = LdaModel.load("../models/ldatrain")
dct = Dictionary.load_from_text("../models/ldatrain_dct")
# Test that import was successful
for topic in range(2):
print([(a, round(b*100, 2)) for (a, b) in lda_model.show_topic(topic, topn=5)])
print("\n")
data = pd.read_csv("../data/interim/trigrams3.csv")
data = data.drop(['Unnamed: 0'], axis=1)
data['TRIGRAMS'] = data['TRIGRAMS'].apply(literal_eval)
data.head()
def remove_numbers(corpus):
regexp = re.compile(r'^[0-9]*$')
return [[word for word in line if not regexp.search(word)] for line in corpus]
full_corpus = data['TRIGRAMS'].tolist()
full_corpus = remove_numbers(full_corpus)
full_dct = dct
full_corpus_nums = [full_dct.doc2bow(text) for text in full_corpus]
# Test that topics have actually been assigned to documents
lda_model.get_document_topics(full_corpus_nums[2])
# +
# Write a function to identify the topic most strongly associated with each document
def get_best_topic(document, model):
best_topic = None
best_prob = 0
candidates = model.get_document_topics(document)
for (topic, prob) in candidates:
if prob > best_prob:
best_topic = topic
best_prob = prob
return best_topic
# -
# Test the function
get_best_topic(full_corpus_nums[2], lda_model)
get_best_topic(full_corpus_nums[1828], lda_model)
# +
# %%time
# Assign each document in the corpus to a particular topic
data.loc[:, 'TOPIC'] = [get_best_topic(full_corpus_nums[row], lda_model)
for row in range(len(data))]
# -
data.head()
train = data.sample(frac=0.8, random_state=42)
test = data.drop(train.index)
train.head()
test.head()
train_means = train.groupby(['TOPIC']).mean()
train_means
test_means = test.groupby(['TOPIC']).mean()
test_means
compare_means = pd.DataFrame({"Train": train_means['FINE'], "Test": test_means['FINE']})
compare_means
# Display average fines by topic for training set vs. test set
compare_means.index=['Life Support', 'Escape', 'Sores', 'Administration', 'Abuse', 'Theft', 'Diabetes', 'Rashes']
pd.options.display.float_format = '${:,.0f}'.format
compare_means.sort_values('Train', ascending=False)
# Average fines are very tightly correlated bewteen topics, r=0.96
stats.pearsonr(compare_means['Train'], compare_means['Test'])
| notebooks/08 Topic Prediction with Train and Test Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !ls converted_trt_graph
# +
import tensorflow as tf
from tensorflow.keras.models import save_model, Sequential
model_path = "./converted_trt_graph/"
model = tf.keras.models.load_model(model_path)
save_model(model,model_path + "\new_model.h5", save_format='h5')
| tf2_pb_to_h5_converter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: titanic
# language: python
# name: titanic
# ---
import pandas as pd
import numpy as np
import os
raw_data_path = os.path.join (os.path.pardir,'data','raw')
train_file_path = os.path.join (raw_data_path,'train.csv')
test_file_path = os.path.join (raw_data_path,'test.csv')
train_df = pd.read_csv (train_file_path, index_col = 'PassengerId')
test_df = pd.read_csv (test_file_path, index_col = 'PassengerId')
type (train_df)
train_df.info ()
test_df.info ()
test_df ['Survived'] = -888
df = pd.concat ((train_df, test_df), axis = 0)
df.info ()
df.head (10)
df.tail (10)
df.Name
df ['Name']
df [['Name','Age']]
df.loc [5:10,]
df.loc [5:10,'Age': 'Pclass']
df.iloc [5:10, 3:8]
male_passenger = df.loc [df.Sex == 'male',:]
print (len(male_passenger))
first_class_male_passenger = df.loc [(df.Sex == 'male') & (df.Pclass == 1),:]
print (len (first_class_male_passenger))
df.describe ()
print (df.Fare.mean())
print (df.Fare.quantile (.25))
# %matplotlib inline
df.Fare.plot (kind = 'box')
df.describe (include = 'all')
df.Sex.value_counts (normalize = True)
df [df.Survived != -888].Survived.value_counts (normalize = True)
df.Pclass.value_counts().plot (kind = 'bar', rot = 0, title = 'Class wise passenger',color = 'c');
| notebooks/exploring-processing-data.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.4
# language: julia
# name: julia-1.5
# ---
using BenchmarkTools, SpecialFunctions
using LinearAlgebra, Statistics
using Parameters
using Plots
using Optim #https://github.com/JuliaNLSolvers/Optim.jl
using Random, Distributions
gr(fmt=:png); # setting for easier display in jupyter notebooks
# **Introduction**
#
# I'm teaching myself ``julia``. The simplest way for me todo so is to work with a model that I know extremely well---the Ricardian model of Eaton and Kortum (2002). So below I will outline the economic environment and then use the properties of ``julia`` to map the fundamental, mathematical environment into a computational/quantitative outcome. Outside of performance properties, the more I reflect on this, this is a huge appeal of the ``julia`` language. We can express the code and design it in a way that most closely corresponds with the math/model. In contrast, in for example MATLAB, where the map from white board to computation is big.
# ### EK (2002) Ricardian Model
#
# **Countries.** To keep things simple, let's first work with a simple two country set up---a home and a foreign country, $h$, $f$.
#
# **Individual Goods Production.**
# Within a country, there is a continuum of intermediate goods indexed by $\omega \in [0, 1]$. As in the Ricardian model of \citet{dornbusch1977comparative} and \citet{eaton2002technology}, intermediate goods are not nationally differentiated. What this means is that intermediate $\omega$ produced in one country is a perfect substitute for the same intermediate $\omega$ produced by another country.
#
# Competitive firms produce intermediate goods with linear production technologies:
#
# \begin{align}
# q(\omega) = z(\omega) \ell,
# \end{align}
#
# where $z$ is the productivity level of firms and $\ell$ is the number of efficiency units of labor.
#
# To ship a technology to another country, iceberg costs are incurred. I will write this as $\tau_{hf}$ which is the cost the home country must incur to buy from the foreign country. The notation here will correspond with the row, column format of a matrix, so for a given row all the trade costs across the different columns are the ones that they face to buy from different countries. Then the usual normalization here is that $\tau_{hh} = 1$. Again, on the computer, this means that the diagonal entries are one.
#
# **The Aggregator.** In this model, intermediate goods are aggregated by a competitive final-goods producer. In Julia, what would be interesting is to make the aggregator generic and let the user specify how this works. But for know, we will follow the canonical situation with a standard CES production function:
#
# \begin{align}
# Q = \left[ \int_0^1 q(\omega)^{\frac{\sigma - 1}{\sigma}} d\omega \right]^{\frac{\sigma}{\sigma - 1}},
# \label{eq:ces}
# \end{align}
#
# where $q(\omega)$ is the quantity of individual intermediate goods $\omega$ demanded by the final-goods firm, and $\sigma$ is elasticity of substitution across variety. I want to talk through some properties/definitions related to this aggregator.
#
# - The aggregator in (\ref{eq:ces}) gives rise to the following the demand curve (in quantities) for an individual variety:
#
# \begin{align}
# q(\omega) & = \left(\frac{p_h(\omega)}{P_h}\right)^{-\sigma}Q.
# \label{eq:demand_curve}
# \end{align}
#
# where $Q$ is the aggregate demand for the final good; $P_h$ is the CES price index, defined below; And $p_h(\omega)$ is the price paid by the home country for good $\omega$ (which is not the same necessarily as the price offered by a home country supplier).
#
# - The demand curve can also be expressed in shares, not quantities. So
#
# \begin{align}
# \frac{p_h(\omega)q(\omega)}{P_h Q} & = \left(\frac{p_h(\omega)}{P_h}\right)^{1-\sigma}.
# \label{eq:demand_curve_share}
# \end{align}
#
# which says that the value of of good $\omega$ relative to total expenditures $P_h Q$ is given by the formula above. This helps as we can compute expenditure patterns while being agnostic about the actual size of the economy, i.e. $Q$. One useful relationship that builds on this is to condition on the source and aggregate to arrive at the share of goods purchased from another country. How does this work?
#
# \begin{align}
# \pi_{h,f} = \int_{p_h = p_{h,f}} \left(\frac{p_h(\omega)}{P_h}\right)^{1-\sigma} d\omega
# \end{align}
#
# Where the notation here $\pi_{h,f}$ is the share of goods that the home country sources from the foreign country. This is equal do the integral over all goods purchased from them, hence, the conditioning where $p_h = p_{h,f}$ and then the inside of the brackets is the same as above. In the computer, the code will be setup with the same row, column notation. So fix a row, then as you go across the different columns you'll view the share of expenditures the, say home country, coming from other countries.
#
# A small side note: If you stare at this enough (and know the liturature), than you can start to see a gravity like relationship here without even imposing the Frechet assumption. The key issue is the conditional distribution of goods prices and how it relates to the aggregate, home CES price index discussed next.
#
# - The **CES Price Index** is given by
#
# \begin{align}
# P_h = \left [ \int_{0}^{1} p_h(\omega)^{1-\sigma} d\omega \right]^{\frac{1}{1-\sigma}}
# \label{eq:ces_price_index}
# \end{align}
#
# which takes all the prices at the micro-level and is like a harmonic mean of these prices. Also note how one can substitute \ref{eq:ces_price_index} into the share formula to get an expression of expenditures shares purely in terms of micro-level prices.
# **The Technology Distribution**
#
# This is the key component that makes Eaton and Kortum fly. First, let's be a bit more general here. One could simply think of each countries **individual productivity** draws being characterized by some distribution:
#
# \begin{align}
# Z_h \sim F_{h}(z)
# \end{align}
#
# where $F_{h}$ is a well defined cumulative probability distribution with the associated pdf $f_h(z)$. What this means is for each $\omega$ variety, in say the home country, the technology/productivity to produce that good is a random $z$ which is characterized by the distribution above. As I'll talk about below, what this allows one to think about is (i) if you hand me a distribution from which I can simulate from (ii) then with assumptions on market structure I can construct prices and (iii) with prices I can construct trade flows.
#
# Now Eaton and Kortum make a very specific distributional assumption. They use a Type II extreme value distribution also known as a Frechet:
#
# \begin{align}
# Z_i \sim \exp\left(-T_i z^{-\theta}\right)
# \end{align}
#
# where the distribution is characterized by a centering parameter $T_i$ which is allowed to vary across countries and a shape parameter $\theta$ which is assumed to be constant across countries. These assumptions are critical to the closed form characterizations in Eaton and Kortum. However, return to the discussion above. On the computer there are only two limiting factors (i) can you simulate from the distribution and (ii) the empirical question of how to discipline the distribution from which you are simulating.
#
# **Optimization**
#
# I'm going to walk through a couple of observations about how this economy works.
#
# The first observation is that within a country, the key equilibrium price is the wage rate $w$. This wage is the same independent of the labor market / goods-producing sector / etc. This derives from the (within-country) free mobility of labor assumption (which I did not talk about). In some other notebooks, I'll relax this. But for now, labor is freely mobile and now we just need to find the wage so that labor demand equals labor supply.
#
# The second observation is that given competitive firms and that they face the wage rate $w$, we know that their marginal cost (and hence price at which they are willing to sell the good domestically) must be:
#
# \begin{align}
# p_{h,h}(\omega) = \frac{w_{h}}{z_{h}(\omega)},
# \label{eq:ek_wage}
# \end{align}
#
# and then the price at which they would be willing to export the good, say from home to foreign is just adjusted by the ice-berg trade costs so
#
# \begin{align}
# p_{f,h}(\omega) = \frac{\tau_{f,h} w_{h}}{z_{h}(\omega)}.
# \end{align}
#
# The third question is who buys what from whom? This aspect of the model is that really is just an application of the min operator. So buy the good from the country with the lowest price:
#
# \begin{align}
# p_{h}(\omega) = \min\left\{ \frac{w_{h}}{z_{h}(\omega)}, \ \frac{\tau_{h,f} w_{f}}{z_{f}(\omega)} \right\}
# \end{align}
#
# where the thing on the left is the domestic price, and then the thing on the right is on the world price (inclusive of the trade cost).
# ### Equilibrium
#
# I'm going to present this in a way slightly different than usual. So I want to think of the wage as clearing the labor market. This implies that goods demand will equal goods supply which will reduce to balanced trade. The idea here is to build up into some later notebooks where we can think of more sophisticated labor market settings with elastic labor supply, mobility/migration frictions similar to my work with Spencer.
#
# **Labor Demand** so the key idea here is the the CES demand curve also is a labor demand curve when combined with the production function. First, consider the simple case of a closed economy. Here we have:
#
# \begin{align}
# \ell(\omega) & = \frac{1}{z(\omega)}\left(\frac{w / z(\omega)}{P_h}\right)^{-\sigma}Q.
# \label{eq:labor_demand_curve}
# \end{align}
#
# where I substituted in the price equaling marginal costs, which after pulling things out we have that
#
# \begin{align}
# \ell_h(\omega) & = z_h(\omega)^{\sigma - 1}\left(\frac{w_h}{P_h} \right)^{-\sigma}Q.
# \label{eq:demand_curve2}
# \end{align}
#
# which has the inuitive idea that (i) if productivity is high or aggregate demand is high you need **more workers** and (ii) if the wage is high you demand less workers. Now let's extend this to the open economy setting. So labor demand is
#
# \begin{align}
# \ell_h(\omega) & = \underbrace{\frac{1}{z_h(\omega)}\left(\frac{w_h / z_h(\omega)}{P_h}\right)^{-\sigma}Q_h}_{\mbox{domestic demand}} \ + \ \underbrace{\frac{1}{z_h(\omega)}\left(\frac{\tau_{f,h} w_h / z_h(\omega)}{P_f}\right)^{-\sigma}Q_f}_{\mbox{foreign demand}}
# \end{align}
#
# where again, I substituted in the price for good $\omega$ in the domestic demand curve and (if purchased) into the foreign demand curve. Note that this is a generic representation in the sense that domestic demand and foreign demand might be zero. In this case, the good is imported from abroad. Or it might be only domesticly demanded (which in this cases the good is non traded) and then it's both bought at home and sent abroad (so that good is exported).
#
# Then one way to think about an equilibrium is we need to find a wage vector $\mathbb{w}$ such labor supply equals labor demand. So
#
# \begin{align}
# L_h^{D} = \int \ell_h(\omega) d\omega = \int_{p_h = p_{h,h}} \frac{1}{z(\omega)}\left(\frac{w_h / z(\omega)}{P_h}\right)^{-\sigma}Q_h + \int_{p_f = p_{f,h}} \frac{\tau_{f,h}}{z(\omega)}\left(\frac{\tau_{f,h} w_h / z(\omega)}{P_f}\right)^{-\sigma}Q_f
# \end{align}
#
# and we would require that $w$ be such that $L_i^{D}(\mathbb{w}) = L^{S}$.
#
# **Goods Demand and Goods Supply** This is an alternative way to find the equillibrium wage vector via the goods market. Now there are in principal two ways to go about this. One is to work from the income side. We know that total income (and spending) in the country is
#
# \begin{align}
# L_h w_h
# \end{align}
#
# which because all this income is only spent on goods, this is the same as goods demand. Then we know that income recived is related to spending and the expenditure shares. So working from the perspective of the home country, we have:
#
# \begin{align}
# \sum_{i} L_i w_i \pi_{i,h} (\mathbb{w})
# \end{align}
#
# which is the sum of each country's expenditure on goods from home. In other words, these are the payments for all the production in the home country. Now this sum has a nice matrix representation.If you define $L_i w_i = Y_i$ which is the typical element in the vector $\mathbb{Y}$, the vector of income for the production of goods across all countries is:
#
# \begin{align}
# \mathbb{Y}'(\mathbb{w}) \times \mathbb{\pi}(\mathbb{w})
# \end{align}
#
# which must equall total spending which is $\mathbb{Y}'(\mathbb{w})$ in equillibrium.
# ### Computing
#
# Given everything laid out, some wages $w_i$ and then primitives regarding technologies $T$, $\theta$, $\sigma$ and $\tau$ we can compute an equillibrium and the objects associated with it, i.e. the pattern of trade, the price index, quantities, etc. Now to compute the equillibrium, we do need to computationally compute the integrals above. I'm going to explore two approaches:
#
# ##### Simmulation
# One approach is to evaluate the integrals by simulation: the idea will be to think of the product space $\omega$ as an integer on the real line which indexes the goods location. That is $\omega = 1,2,...$ Then for each of those products, each country is randomly assigned a productivity from the distribution. Then the number of goods, so to speak, is set to be large and thus will approximate the integrals above. So what this means is, for example, the expenditure share of goods that country $h$ buys from country $f$ is:
#
# \begin{align}
# \pi_{h,f} = (1 / N) \sum_{\omega = 1}^N \mathbb{1}\{p_h = p_{h,f}\} \left(\frac{p_h(\omega)}{P_h}\right)^{1-\sigma}
# \end{align}
#
# Note that the probability country $h$ sources from country $f$ is
# \begin{align}
# \hat \pi_{h,f} = (1 / N) \sum_{\omega = 1}^N \mathbb{1}\{p_h = p_{h,f}\}
# \end{align}
#
# where it turns out that the sourcing probability happens to correspond with the expenditure share, so $\pi_{h,f} = \hat \pi_{h,f}$. This latter point is very subtle, special to the Frechet distribution + CES. One way to see what is going on is to note that if (this is super loose, but in the discretized case I think you can see this easier) the expected valye of imported $p_{h,f}$ goods is the same as the aggregate price index, then things cancel and one could see the correspondence.
#
# Now, to compute the expenditure shares, we must compute the CES price index. This is computed as:
#
# \begin{align}
# P_h = \left [ (1 / N) \sum_{\omega = 1}^N p_h(\omega)^{1-\sigma} \right]^{\frac{1}{1-\sigma}}
# \end{align}
#
# And labor demand is computed as:
#
# \begin{align}
# L^D = (1 / N) \sum_{\omega = 1}^N \mathbb{1}\{p_h = p_{h,h}\} \frac{1}{z(\omega)}\left(\frac{w_h / z_h(\omega)}{P_h}\right)^{-\sigma}Q_h +
# (1 / N) \sum_{\omega = 1}^N \mathbb{1}\{p_f = p_{f,h}\} \frac{\tau_{f,h}}{z(\omega)}\left(\frac{\tau_{f,h} w_h / z_f(\omega)}{P_f}\right)^{-\sigma}Q_f
# \end{align}
#
# - The CES function ``ces_shares`` that takes prices and sourcing decisions to compute trade shares.
#
# - A function that computes potential prices based on marginal costs, ``marginal_costs``
#
# - A function that constructs a vector of productivity for each good, across all countries, given the Frechet distribution, this is the ``make_productivity`` function. Note, future, it would be nice if the distribution type could be directly passed into this and also appropriately indexed. Maybe with a dictionary?
# +
# this creates a tuple with the named parameters
# then with the @unpack command (from parameters)
# you can simply grab from it what you need...
# so pp = ekparams(θ = 5.0) creates the parameter structure
# but with theta = 5. if not specified, below are the defalut values.
# for example @unpack θ, T, σ = pp; only pulls out
# the theta, T, sigma from the
ekparams = @with_kw (θ = 4.0,
T = [1.0 2.5],
τ_matrix = [1.0 1.5; 1.5 1.0],
L = [1.0 1.0],
ngoods = 10000,
ncntry = ndims(τ_matrix),
σ = 2.0,
seed = 3281978
)
# +
# This is me trying multiple dispatch
# so the ces function will do different things depending upon
# the arguments passed. The frist one deliivers the shares
# but if aggregate demand is passed (Q), then you get out the
# real quantities.
function ces(p, σ)
ngoods = size(p)[1]
return ( sum( p.^(one(σ)-σ) ) / ngoods ).^( one(σ) ./ (one(σ) - σ) )
# So if it's just small p, then kick back the price index
# I'm using the one(sigma) to make sure type stable. maybe
# this might be doing ints on floats here...need to think more
# about this, use promotions?
end
function ces(p, P, σ)
return (p ./ P).^(one(σ) - σ)
# So if it has the big P, it return the expenditure share
end
function ces(p, P, Q, σ)
return ((p ./ P).^(- σ) ).*Q
# return the quantity if big Q is included
end
# +
function ces_shares(p, source_country::Array{CartesianIndex{2},2}, σ = 2.0)
# this will take in (i) prices **purchased at** and
# a cartesian index that can determine country supplier
ngoods = length(p) # number of goods
ncntry = ndims(source_country) # number of coutnries
price_index = ces(p, σ) # CES price index
# now construct the shares...
trade_shares = zeros(ncntry)
source_prob = zeros(ncntry)
for cntry = 1:ncntry
# This is the loop that conditions on the source country
# first fix a country...
supplier = [ zzz[2] == cntry for zzz in source_country ]
# source_country is a 2dim caartesian index which
# says for each good (first dimension), which country is the low cost
# supplier (second country).
#
# this is then a simple inline expression saying, for each
# good (so every entry along the first dimension), which values in the second
# dimension zzz[2] are equall to the country in question "cntry"
# the result is supplier which is a vector of logicals, with true (one)
# values indicating that "cntry: is the low cost supplier
#
# To map to the math above, this is compute the indicator functions
# for each good, from the perspective of each country.
source_prob[cntry] = (1 / ngoods) * sum(supplier)
# This will compute the simple probability of being the low cost
# supplier, by summing across the indicator/bools
# In the Frechet model = trade share. Alternative distributional
# assumptions will deliver something different.
trade_shares[cntry] = (1 / ngoods) * sum(ces(p[supplier], price_index, σ))
# then this is just the ces demand curve described above.
# sum across p's, conditional on it being supplied,
# then divide by number of goods
end
return price_index, trade_shares, source_prob
end
# +
function marginal_cost(z, wage, τ)
return (τ.*wage)./z
end
# +
function make_productity(model_params)
@unpack θ, T, ncntry, ngoods, seed = model_params;
z = zeros(ngoods, ncntry)
for cntry = 1:ncntry
dist = Frechet(θ, T[cntry]^(one(θ)/θ))
# Ideally, it would be cool to potentially be able to pass as an argument
# the distibution for each country or something like that.
z[:,cntry] = rand(MersenneTwister(seed + cntry), dist, ngoods, 1);
#I tried the rand! command (if modifies the argument),
# it did not work...seemed to rewrite everytime
# this thing looped through.
# MersenneTwister is the seed setter for the random number generator...
# works simmilar to matlab...takes an integer and then specifies
end
return z
end
# +
function low_cost_supplier(z, w, τ)
return p, source_country = findmin(marginal_cost(z, w, τ); dims = 2);
end
# +
function equillibrium_evaluate(w, z, model_params)
@unpack ncntry, ngoods, τ_matrix, L = model_params;
price_index = similar(w)
trade_shares = zeros(ncntry,ncntry)
source_prob = similar(trade_shares)
source_country = Array{CartesianIndex{2}}(undef, size(z))
# this I thought I knew what I was doing, not sure now
for cntry = 1:ncntry
p, sc = low_cost_supplier(z, w, τ_matrix[cntry,:]');
# find the min over marginal costs
# note that the tau_matrix returns a vector, not a row vector as I was anticipating
# so take the transpose to make it work.
(price_index[cntry], trade_shares[cntry,:], source_prob[cntry,:]) = ces_shares(p, sc);
# then this is set up so the row is the buyer...then you go across the columns
# and that is share of goods the home country buys from that country.
source_country[:, cntry] = sc ;
end
value_expenditure = (L.*w)*trade_shares ;
# Need to spell this out in markdown
# Note this is a matrix operation. Lw is N x 1, trades shares is N x N
# so this gives a N x 1 vector of expendeture.
value_production = L.*w ;
return norm(value_production - value_expenditure), trade_shares, price_index, source_country, source_prob
end
##########################################################################################################
# here again, I'm going to try and explit multiple dispatch. So if I pass an additional variable then it
# will compute the equillibrium via labor demand
##########################################################################################################
function equillibrium_evaluate(w, z, model_params, ld_flag)
@unpack ncntry, ngoods, τ_matrix, L = model_params;
price_index = similar(w)
agg_l_demand = similar(w)
trade_shares = zeros(ncntry,ncntry)
source_prob = similar(trade_shares)
source_country = Array{CartesianIndex{2}}(undef, size(z))
# this I thought I knew what I was doing, not sure now
for cntry = 1:ncntry
p, sc = low_cost_supplier(z, w, τ_matrix[cntry,:]');
# find the min over marginal costs
# note that the tau_matrix returns a vector, not a row vector as I was anticipating
# so take the transpose to make it work.
(price_index[cntry], trade_shares[cntry,:], source_prob[cntry,:]) = ces_shares(p, sc);
# then this is set up so the row is the buyer...then you go across the columns
# and that is share of goods the home country buys from that country.
source_country[:, cntry] = sc ;
end
endogenous_var = @with_kw (source_country = source_country,
z = z,
w = w,
price_index = price_index,
Q = L.*w ./ price_index)
for cntry = 1:ncntry
agg_l_demand[cntry] = labor_demand(endogenous_var(), cntry, model_params)[1];
end
return norm(agg_l_demand - L), trade_shares, price_index, source_country, source_prob
end
# +
function labor_demand(endogenous_var, country, model_params)
@unpack source_country, z, w, price_index, Q = endogenous_var;
@unpack τ_matrix, σ = model_params;
destinations = Array{Bool}(undef, size(z))
ncntry = size(z)[2]
ngoods = size(z)[1]
for cntry = 1:ncntry
destinations[:, cntry] = [ zzz[2] == country for zzz in source_country[:, cntry] ]
# this is different than the ces demand line. it says for a given country, go through
# each country low cost supplier and find the places where that given country is the
# low cost supplier.
# Then the destination country is a matrix of bools so indicating if country
# is the low cost supplier to those different (columns) destiation.
# really this is like from the exporter side....
end
mc = marginal_cost(z[:, country], w[country], τ_matrix[:, country]')
q_demand = ces(mc, price_index, Q, σ)
# here I'm using the multiple dispatch of ces...when I pass
# the z argument to it, it returns labor demand
l_demand = q_demand ./ ( z[:, country] ./ τ_matrix[:, country]' ) ./ ngoods
agg_l_demand = sum(sum(l_demand.*destinations, dims = 2))
return agg_l_demand, l_demand.*destinations, destinations
end
# +
model_params = ekparams(ngoods = 100000, θ = 4.0)
z = make_productity(model_params)
ld_flag = true
f(x) = equillibrium_evaluate(([1.0 x]), z, model_params)[1]
# this is like creating an inline function in matlab. much easier
# just create f(x) where x is the variable of interest.
# again, the ld_flag if present uses the labor demand condition
# This is an example of univarite, between bounds optimization
result = optimize(f, (0.05), (5.0), show_trace = true,
show_every = 10 )
# -
println(result.minimizer)
println(result.minimizer)
# +
w = [1.0 result.minimizer]
obj = equillibrium_evaluate(w, z, model_params)
println("expenditure shares: ", obj[2])
println("sourcing probabilities: ", obj[5])
# +
@unpack τ_matrix = model_params;
p, sc = low_cost_supplier(z, w, τ_matrix[1,:]');
mc = marginal_cost(z, w, τ_matrix[1,:]');
histogram(p, bins = 100, alpha = 0.5, color = "dark blue", label = "traded prices", xlabel = "prices/marginal costs")
histogram!(mc[:,1], bins = 100, alpha = 0.5, color = "red", label = "domestic marginal costs")
# Note there is something up with the bins here. Eg. it is overriding my stuff...
# not sure why.
# +
obj = equillibrium_evaluate(w, z, model_params)
@unpack L = model_params;
endogenous_var = @with_kw (source_country = obj[4],
z = z,
w = w,
price_index = obj[3],
Q = L.*w ./ price_index,
)
agg_l_demand, ld, srcprb = labor_demand(endogenous_var(), 1, model_params);
lall = sum(ld[ld[:,1] .> 0.0, :], dims = 2)
lexp = sum(ld[ld[:,2] .> 0.0, :], dims = 2)
histogram(log.(lall), bins = 100, alpha = 0.25, color = "dark blue", xlabel = "log labor demand", label = "all goods")
histogram!(log.(lexp), bins = 100, alpha = 0.5, color = "red", label = "exported goods")
# same deal and then the histogram looks messed up as the binning is
# different
# -
# +
ql = sum(sum(ld, dims = 2));
print("Labor Demand ", string(ql)[1:5])
# -
agg_l_demand
@btime f(result.minimizer)
@btime f(result.minimizer)
@btime labor_demand(endogenous_var(), 1, model_params)
@btime labor_demand(endogenous_var(), 1, model_params)
@code_warntype equillibrium_evaluate(w, z, model_params)
@btime equillibrium_evaluate(([1.0 result.minimizer]), z, L, tau_matrix)
| ek-model.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 9.0
# language: sage
# name: sagemath
# ---
# # Actividad 3. Método de Gauss
#
# Este notebook servira para corroborar los ejercicios de la _Actividad 3_ de la _Unidad 2_ del curso de _Álgera Lineal_ de la _UnADM_.
#
# El uso de este material, en actividades realcionadas con la _UnADM_, debe regirse por el [código de ética](https://www.unadmexico.mx/images/descargables/codigo_de_etica_de_estudiantes_de_la_unadm.pdf) de la institución. Para cualquier otro proposito favor de seguir los lineamientos expresados en el archivo [readme.md](../../../readme.md) de este repositorio repositorio.
# +
# Naive Gaussian reduction
def shown( obj ):
"""Funcion que permite establecer la forma de visualizacion de la funcion
siguiente
"""
# Mostrar la representacion latex de la matriz
#print(latex(obj))
# Mostrar la matriz
print(obj)
def gauss_method(M,rescale_leading_entry=False):
"""Describe the reduction to echelon form of the given matrix of rationals.
M matrix of rationals e.g., M = matrix(QQ, [[..], [..], ..])
rescale_leading_entry=False boolean make the leading entries to 1's
Returns: None. Side effect: M is reduced. Note: this is echelon form,
not reduced echelon form; this routine does not end the same way as does
M.echelon_form().
"""
num_rows=M.nrows()
num_cols=M.ncols()
shown(M)
col = 0 # all cols before this are already done
for row in range(0,num_rows):
# ?Need to swap in a nonzero entry from below
while (col < num_cols
and M[row][col] == 0):
for i in M.nonzero_positions_in_column(col):
if i > row:
print(" intercambiamos la fila",row+1," con la fila",i+1)
M.swap_rows(row,i)
shown(M)
break
else:
col += 1
if col >= num_cols:
break
# Now guaranteed M[row][col] != 0
if (rescale_leading_entry and M[row][col] != 1):
print(" multiplicamos ",1/M[row][col]," veces la fila ",row+1)
M.rescale_row(row,1/M[row][col])
shown(M)
change_flag=False
for changed_row in range(row+1,num_rows):
if M[changed_row][col] != 0:
change_flag=True
factor=-1*M[changed_row][col]/M[row][col]
print(" hacemos que la fila", changed_row+1,
"sea",factor,
"veces la fila",row+1,
"mas la fila",changed_row+1)
M.add_multiple_of_row(changed_row,row,factor)
if change_flag:
shown(M)
col +=1
# Fuente: https://ask.sagemath.org/question/8840/how-to-show-the-steps-of-gauss-method/
# -
def comprueba(M):
gauss_method(M)
print("\n\n y al final vemos que las soluciones son ")
dim = 3
# Solucione con matriz inversa
A = M[:, :dim]
b = M[:, dim]
sol = (~A)*b
print(sol)
# ### Test
#
# Vamos a probar la funcionalidad que usaremos durante este ejercicio. Primero la funcion anterior, para corroborar el paso en Gauss de matriz triangular invertida, para luego resolver con inversa.
#
# El ejemplo fue tomado de https://www.mathwords.com/g/gaussian_elimination.htm
# Eliminacion gaussiana sobre matriz
M = matrix(QQ, [[1,1,1,3],
[2,3,7,0],
[1,3,-2,17]
])
gauss_method(M)
# +
# Correccion de metodo
M[2] = (0,0,1, -2)
dim = 3
# Solucione con matriz inversa
A = M[:, :dim]
b = M[:, dim]
sol = (~A)*b
print(sol)
# -
# ## Sistema 1
M = matrix(QQ, [[2,7,6,48],
[4,5,9,24],
[3,1,-2,14]
])
comprueba(M)
# ## Sistema 2
M = matrix(QQ, [[1,12, 3,19],
[0, 5, 6,20],
[0, 7, 2, 1]
])
comprueba(M)
# ## Sistema 3
M = matrix(QQ, [[1,-2, 4, 7],
[0, 2,-8, 6],
[0, 5, 7,21]
])
comprueba(M)
# ### Correción de ejercicio
#
# El profesor realizo algunas correciones al ejercicio; al final estos son los sistemas que se deban resolver para la tarea.
# #### Sistema 2
M = matrix(QQ, [[1,12, 3,19],
[4, 5, 6,24],
[3, 7, 2, 4]
])
comprueba(M)
M = matrix(QQ, [[1,-2, 4, 7],
[4, 2,-8,10],
[2, 5, 7,23]
])
comprueba(M)
| B1-1/BALI/Actividades/BALI_U2_A3_BERC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
# reading csv file
athlete_events = pd.read_csv('../CSV for ML models/athlete_events.csv')
athlete_events.head()
filter_data = athlete_events[["Sex", "Age", "Height", "Weight", "Team", "Year", "Season", "Sport", "Event", "Medal"]]
filter_data.head()
# get the data for winter Olympic
winter_data = filter_data[filter_data["Season"] == "Winter"]
winter_data.head()
# +
# get the data for summer Olympic
summer_data = filter_data[(filter_data["Season"] == "Summer")]
summer_data.head()
# -
# print out every sport in the summer Olympic
summer_data["Sport"].unique()
summer_sports = summer_data["Sport"].unique()
# +
# For loop to train the Logistic Regression model and get the testing score for male athletes in every Summer Olympic Sport
for sport in summer_sports:
try:
athlete_m = summer_data[(summer_data["Sport"] == f'{sport}') & (summer_data["Sex"] == "M")]
athlete_m = athlete_m[["Age", "Height", "Weight", "Medal"]]
athlete_m = athlete_m.dropna(subset=['Height', 'Weight']).reset_index(drop = True)
# One-hot encoding
athlete_m = athlete_m.replace("Gold", 1)
athlete_m = athlete_m.replace("Silver", 1)
athlete_m = athlete_m.replace("Bronze", 1)
athlete_m["Medal"] = athlete_m["Medal"].fillna(0)
athlete_m = athlete_m.dropna()
athlete_m["Medal"].unique()
# Assign X (data) and y (target)
X1 = athlete_m[['Height', "Weight", "Age"]]
print(X1.shape)
y1 = athlete_m['Medal']
print(y1.shape)
print(sport)
# Split our data into training and testing
X1_train, X1_test, y1_train, y1_test = train_test_split(X1, y1, random_state=42)
# Create a Logistic Regression Model
classifier = LogisticRegression()
#Fit (train) or model using the training data
classifier.fit(X1_train, y1_train)
#Validate the model using the test data
print(f"Training Data Score: {classifier.score(X1_train, y1_train)}")
print(f"Testing Data Score: {classifier.score(X1_test, y1_test)}")
# Predict the testing data point
predictions = classifier.predict(X1_test)
pd.DataFrame({"Prediction": predictions, "Actual": y1_test})
print("----------------------------------------------------")
except:
print("An exception occurred")
print("----------------------------------------------------")
# +
# For loop to train the Logistic Regression model and get the testing score for female athletes in every Summer Olympic Sport
for sport in summer_sports:
try:
athlete_f = summer_data[(summer_data["Sport"] == f'{sport}') & (summer_data["Sex"] == "F")]
athlete_f = athlete_f[["Age", "Height", "Weight", "Medal"]]
athlete_f = athlete_f.dropna(subset=['Height', 'Weight']).reset_index(drop = True)
# One-hot encoding
athlete_f = athlete_f.replace("Gold", 1)
athlete_f = athlete_f.replace("Silver", 1)
athlete_f = athlete_f.replace("Bronze", 1)
athlete_f["Medal"] = athlete_f["Medal"].fillna(0)
athlete_f = athlete_f.dropna()
athlete_f["Medal"].unique()
# Assign X (data) and y (target)
X = athlete_f[['Height', "Weight", "Age"]]
print(X.shape)
y = athlete_f['Medal']
print(y.shape)
print(sport)
# Split our data into training and testing
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# Create a Logistic Regression Model
classifier = LogisticRegression()
classifier
# Fit (train) or model using the training data
classifier.fit(X_train, y_train)
# Validate the model using the test data
print(f"Training Data Score: {classifier.score(X_train, y_train)}")
print(f"Testing Data Score: {classifier.score(X_test, y_test)}")
# Predict the testing data point
predictions = classifier.predict(X_test)
pd.DataFrame({"Prediction": predictions, "Actual": y_test})
print("----------------------------------------------------")
except:
print("An exception occurred")
print("----------------------------------------------------")
# -
# print out every sport in the winter Olympic
winter_data['Sport'].unique()
winter_sports = winter_data['Sport'].unique()
# For loop to train the Logistic Regression model and get the testing score for male athletes in every Winter Olympic Sport
for sport in winter_sports:
try:
athlete_m = winter_data[(winter_data["Sport"] == f'{sport}') & (winter_data["Sex"] == "M")]
athlete_m = athlete_m[["Age", "Height", "Weight", "Medal"]]
athlete_m = athlete_m.dropna(subset=['Height', 'Weight']).reset_index(drop = True)
# One-hot encoding
athlete_m = athlete_m.replace("Gold", 1)
athlete_m = athlete_m.replace("Silver", 1)
athlete_m = athlete_m.replace("Bronze", 1)
athlete_m["Medal"] = athlete_m["Medal"].fillna(0)
athlete_m = athlete_m.dropna()
athlete_m["Medal"].unique()
# Assign X (data) and y (target)
X = athlete_m[['Height', "Weight", "Age"]]
print(X.shape)
y = athlete_m['Medal']
print(y.shape)
print(sport)
# Split our data into training and testing
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# Create a Logistic Regression Model
classifier = LogisticRegression()
classifier
# Fit (train) or model using the training data
classifier.fit(X_train, y_train)
# Validate the model using the test data
print(f"Training Data Score: {classifier.score(X_train, y_train)}")
print(f"Testing Data Score: {classifier.score(X_test, y_test)}")
# Predict the testing data point
#predictions = classifier.predict(X_test)
#pd.DataFrame({"Prediction": predictions, "Actual": y_test})
print("----------------------------------------------------")
except:
print("An exception occurred")
print("----------------------------------------------------")
# For loop to train the Logistic Regression model and get the testing score for female athletes in every Winter Olympic Sport
for sport in winter_sports:
try:
athlete_f = winter_data[(winter_data["Sport"] == f'{sport}') & (winter_data["Sex"] == "F")]
athlete_f = athlete_f[["Age", "Height", "Weight", "Medal"]]
athlete_f = athlete_f.dropna(subset=['Height', 'Weight']).reset_index(drop = True)
# One-hot encoding
athlete_f = athlete_f.replace("Gold", 1)
athlete_f = athlete_f.replace("Silver", 1)
athlete_f = athlete_f.replace("Bronze", 1)
athlete_f["Medal"] = athlete_f["Medal"].fillna(0)
athlete_f = athlete_f.dropna()
athlete_f["Medal"].unique()
# Assign X (data) and y (target)
X = athlete_f[['Height', "Weight", "Age"]]
print(X.shape)
y = athlete_f['Medal']
print(y.shape)
print(sport)
# Split our data into training and testing
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# Create a Logistic Regression Model
classifier = LogisticRegression()
classifier
# Fit (train) or model using the training data
classifier.fit(X_train, y_train)
# Validate the model using the test data
print(f"Training Data Score: {classifier.score(X_train, y_train)}")
print(f"Testing Data Score: {classifier.score(X_test, y_test)}")
# Predict the testing data point
#predictions = classifier.predict(X_test)
#pd.DataFrame({"Prediction": predictions, "Actual": y_test})
print("----------------------------------------------------")
except:
print("An exception occurred")
print("----------------------------------------------------")
# ## Logistic Regression for male athletes in Gymnastics
Gymnastics_M = summer_data[(summer_data["Sport"] == "Gymnastics") & (summer_data["Sex"] == "M")]
Gymnastics_M = Gymnastics_M[["Age", "Height", "Weight", "Medal"]]
Gymnastics_M = Gymnastics_M.dropna(subset=['Height', 'Weight']).reset_index(drop = True)
Gymnastics_M = Gymnastics_M.replace("Gold", 1)
Gymnastics_M = Gymnastics_M.replace("Silver", 1)
Gymnastics_M = Gymnastics_M.replace("Bronze", 1)
Gymnastics_M["Medal"] = Gymnastics_M["Medal"].fillna(0)
Gymnastics_M = Gymnastics_M.dropna()
# +
# Visualizing athlete with medal vs athlete without medal
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure(1, figsize=(5, 5))
axes = Axes3D(fig, elev=20, azim=45)
axes.scatter(Gymnastics_M['Age'], Gymnastics_M['Height'], Gymnastics_M['Weight'], c=Gymnastics_M['Medal'], cmap=plt.cm.get_cmap("Spectral"))
plt.show()
# +
a = summer_data[(summer_data["Sport"] == "Gymnastics") & (summer_data["Sex"] == "M")]
a = a[["Age", "Height", "Weight", "Medal"]].dropna().reset_index(drop = True)
a = a.replace("Gold", 1)
a = a.replace("Silver", 2)
a = a.replace("Bronze", 3)
# +
# Visualizing Gold, Silver, Bronze medalists data
fig = plt.figure(1, figsize=(5, 5))
axes = Axes3D(fig, elev=20, azim=45)
axes.scatter(a['Age'], a['Height'], a['Weight'], c=a['Medal'], cmap=plt.cm.get_cmap("Spectral"))
plt.show()
# -
Gymnastics_M["Medal"].unique()
# +
# Assign X (data) and y (target)
X = Gymnastics_M[['Height', "Weight", "Age"]]
print(X.shape)
y = Gymnastics_M['Medal']
print(y.shape)
# +
# Split our data into training and testing
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# +
# Create a Logistic Regression Model
classifier = LogisticRegression()
classifier
classifier.fit(X_train, y_train)
# -
# Validate the model using the test data
print(f"Training Data Score: {classifier.score(X_train, y_train)}")
print(f"Testing Data Score: {classifier.score(X_test, y_test)}")
# +
# Predict the testing data point
predictions = classifier.predict(X_test)
pd.DataFrame({"Prediction": predictions, "Actual": y_test})
| ML Models/LogisticRegression_olympic medalists.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import random
sample_data = pd.read_csv("./data/blogdata.csv").transpose()
sample_data.head()
# ## Cicle Names and Pincode data
circle_pincode_df = pd.read_csv("./data/all_india_PO_list_without_APS_offices_ver2_lat_long.csv")
circle_pincode_df.head()
# #### Column Names
column_names = list(sample_data.loc['Customer Engagement Variables'])
column_names.insert(0, "Customer ID")
column_names
# ### Function to get a list of random values
def get_random_data(entity_categories):
entity_list = []
for _ in range(10000):
r = random.randint(0, (len(entity_categories)-1))
entity_list.append(entity_categories[r])
return entity_list
# #### 01 - Customer ID
customer_id = list(range(10000))
# #### 02 - Circle Names & 05 - Pincode
circle_names = []
pincodes = []
while len(circle_names) < 10000:
r = random.randint(0, 154797)
if circle_pincode_df.iloc[r]['Taluk'] is not np.nan:
circle_names.append(circle_pincode_df.iloc[r]['Taluk'])
pincodes.append(circle_pincode_df.iloc[r]['pincode'])
# #### 03 - Age
age_categories = ['20-25', '25-30', '30-35', '35-40', '40-45', '45-50', '50-55', '55-60', '60-65', '65-70']
age = get_random_data(age_categories)
# #### 04 - Gender
gender_categories = ['M', 'F']
gender = get_random_data(gender_categories)
count = 0
for g in gender:
if g == "F":
count+=1
count
# #### 06 - Age On Network
age_on_network_slabs = ['0-2 Months', '2-4 Months', '4-6 Months', '6-8 Months', '8-10 Months', '10-12 Months',
'12-14 Months', '14-16 Months', '16-18 Months', '18-20 Months', '20-22 Months', '22-24 Months']
age_on_network = get_random_data(age_on_network_slabs)
# #### 07 - Connection Type
connection_type_categories = ['Prepaid', 'Postpaid']
connection_type = get_random_data(connection_type_categories)
# #### 08 - Voice Usage
voice_usage_slabs = ['1-50', '51-100', '101-150', '151-200', '201-250', '250-300', '301-500', '501-750', '751-1000', '1001-1500', '1501-2000']
voice_usage = get_random_data(voice_usage_slabs)
# #### 09 - Data Usage
data_usage_slabs = ['0GB-0.5GB', '0.5GB-1GB', '1GB-1.5GB', '1.5GB-2GB', '2GB-2.5GB', '2.5GB-5GB',
'5GB-7.5GB', '7.5GB-10GB', '10GB-15GB', '>15GB']
data_usage = get_random_data(data_usage_slabs)
# #### 10 - SMS Usage
sms_usage_slabs = ['1-50', '51-100', '101-150', '151-200', '201-250', '250-300', '301-500', '501-750', '751-1000', '>1000']
sms_usage = get_random_data(sms_usage_slabs)
# #### 11 - ARPU
average_revenue_slabs = ['1-50', '51-100', '101-150', '151-200', '201-250', '250-300', '301-500', '501-750', '751-1000', '>1000']
arpu = get_random_data(average_revenue_slabs)
# #### 12 - ISD Usage
isd_usage_categories = ['Yes', 'No']
isd_usage = get_random_data(isd_usage_categories)
# #### 13 - IR Usage
ir_usage_categories = ['Yes', 'No']
ir_usage = get_random_data(ir_usage_categories)
# #### 14 - Amazon Prime
amazon_prime_categories = ['Yes', 'No']
amazon_prime = get_random_data(amazon_prime_categories)
# #### 15 - Netflix
netflix_categories = ['Yes', 'No']
netflix = get_random_data(netflix_categories)
# #### 16 - Sender ID
sender_id_categories = ['Healthcare', 'Entertainment', 'Banking Services', 'Food', 'Shopping', 'Work', 'Lifestyle', 'Travel', '']
sender_id = get_random_data(sender_id_categories)
# #### 17 - Value Added Services
vas_subscription_categories = ['Yes', 'No']
vas_subscription = get_random_data(vas_subscription_categories)
# #### 18 - VIL App Web
vil_app_web_categories = ['Yes', 'No']
vil_app_web = get_random_data(vil_app_web_categories)
# #### 19 - Active User (Web/App)
active_user_categories = ['Yes', 'No']
active_user = get_random_data(active_user_categories)
# #### 20 - Recharge Own Asset
recharge_own_asset_categories = ['Yes', 'No']
recharge_own_asset = get_random_data(recharge_own_asset_categories)
# #### 21 - Recharge Aggregator (Value)
recharge_aggregator_value_categories = ['1-50', '51-100', '101-150', '151-200', '201-250', '250-300', '301-500', '501-750', '751-1000', '>1000']
recharge_aggregator_value = get_random_data(recharge_aggregator_value_categories)
# #### 22 - Recharge Aggregator (Volume)
recharge_aggregator_volume_categories = list(range(10))
recharge_aggregator_volume = get_random_data(recharge_aggregator_volume_categories)
# #### 23 - Recharge Aggregator (Partner Category)
recharge_aggregator_partner_categories = ['Payment Banking', 'Wallet']
recharge_aggregator_partner_category = get_random_data(recharge_aggregator_partner_categories)
# #### 24 - Bill Payment Own Asset
bill_payment_own_asset_slabs = ['Yes', 'No']
bill_payment_own_asset = get_random_data(bill_payment_own_asset_slabs)
# #### 25 - Bill Payment Aggregator (Value)
bill_payment_aggregator_value_slabs = ['1-50', '51-100', '101-150', '151-200', '201-250', '250-300', '301-500', '501-750', '751-1000', '>1000']
bill_payment_aggregator_value = get_random_data(bill_payment_aggregator_value_slabs)
# #### 26 - Bill Payment Aggregator (Volume)
bill_payment_aggregator_volume_slabs = list(range(10))
bill_payment_aggregator_volume = get_random_data(bill_payment_aggregator_volume_slabs)
# #### 27 - Bill Payment Aggregator (Partner Name)
bill_payment_aggregator_partner_name_slabs = ['Payment Banking', 'Wallet']
bill_payment_aggregator_partner_name = get_random_data(bill_payment_aggregator_partner_name_slabs)
# #### 28 - Content App Stream
content_app_stream_slabs = ['Yes', 'No']
content_app_stream = get_random_data(content_app_stream_slabs)
# #### 29 - Content App Active User
content_app_active_user_slabs = ['Yes', 'No']
content_app_active_user = get_random_data(content_app_active_user_slabs)
# #### 30 - Content App (Movie/Live TV/Shows)
content_app_categories = ['Movies', 'Live TV', 'Shows']
content_app = get_random_data(content_app_categories)
# #### 31 - Content Genre
content_genre_slabs = ['Action', 'Comedy', 'Romance', 'Drama', 'Thriller', 'Sci-fi', 'Horror', 'Fantasy']
content_genre = get_random_data(content_genre_slabs)
# #### 32 - Content Language
content_language_slabs = ['Hindi', 'English']
content_language = get_random_data(content_language_slabs)
# #### 33 - Credit Limit
credit_limit_slabs = ['500', '1000', '1500', '2000', '2500', '3000', '4000', '5000', '6000', '7000', '8000', '9000', '10000']
credit_limit = get_random_data(credit_limit_slabs)
# #### 34 - Category DND Flag
category_dnd_slabs = ['Yes', 'No']
category_dnd = get_random_data(category_dnd_slabs)
# #### 35 - Brand Identifier
brands = ['Vodafone', 'Idea']
brand_identifier = get_random_data(brands)
# # Making a Dataframe
practice_dict = {'Customer ID': customer_id, 'Circle Name': circle_names, 'Age':age, 'Gender': gender, 'Pincode': pincodes,
'Age On Network': age_on_network, 'Connection Type (Prepaid/Postpaid)': connection_type,
'Voice Usage (Slab)': voice_usage, 'Data Usage (Slab)': data_usage, 'Sms Usage (Slab)': sms_usage,
'ARPU': arpu, 'ISD Usage': isd_usage, 'IR Usage': ir_usage, 'Amazon Prime User': amazon_prime,
'Netflix User': netflix, 'Sender ID': sender_id, 'VAS Subscription': vas_subscription,
'VIL App Web (Login/Registration)': vil_app_web, 'Active User (App / Web)': active_user,
'Recharge Own Asset': recharge_own_asset, 'Recharge Aggregator (Value)': recharge_aggregator_value,
'Recharge Aggregator (Volume)': recharge_aggregator_volume, 'Recharge Aggregator (Partner Category)': recharge_aggregator_partner_category,
'Bill Payment Own Asset': bill_payment_own_asset, 'Bill Payment Aggregator (Value)': bill_payment_aggregator_value,
'Bill Payment Aggregator (Volume)': bill_payment_aggregator_volume, 'Bill Payment Aggregator (Partner Name)': bill_payment_aggregator_partner_name,
'Content App Stream': content_app_stream, 'Content App Active User': content_app_active_user,
'Content App (Movie/Live TV/Shows)': content_app, 'Content Genre': content_genre, 'Content Language': content_language,
'Credit Limit': credit_limit, 'Category DND Flag': category_dnd, 'Brand Identifier': brand_identifier}
practice_df = pd.DataFrame.from_dict(practice_dict)
practice_df.head(15)
# ## Saving to a csv file
practice_df.to_csv('./data/practice.csv', )
| CustomDash-master/data preparation/dataset preparation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
people = {
"first": ["Corey", 'Jane', 'John'],
"last": ["Schafer", 'Doe', 'Doe'],
"email": ["<EMAIL>", '<EMAIL>', '<EMAIL>']
}
df = pd.DataFrame(people)
df
# -
df.columns
df.columns = ['first_name','last_name','email']
df.columns
df
df.columns = [x.upper() for x in df.columns]
df
df.columns = df.columns.str.replace('_',' ')
df
df.columns = [x.lower() for x in df.columns]
df.columns = df.columns.str.replace(' ','_')
df
df.rename(columns = {'first_name':'first','last_name':'last'},inplace=True)
df
df.loc[2]
df.loc[2] = ['John','Smith','<EMAIL>']
df
df.loc[2,['last','email']] = ['Doe','<EMAIL>']
df
df.loc[2,'last'] = 'Smith'
df
df.at[2,'last'] = 'Doe'
df
filt = (df['email'] == '<EMAIL>')
df[filt]
df[filt]['last']
df.loc[filt,'last'] = 'Smith'
df
df['email'].str.lower()
df['email'] = df['email'].str.lower()
df
df['email'].apply(len) # function
def update_email(email):
return email.upper()
df['email'].apply(update_email)
df['email'] = df['email'].apply(update_email)
df
df['email'] = df['email'].apply(lambda x : x.lower())
df
df['email'].apply(len)
df.apply(len) # length function to each series
len(df['email'])
df.apply(len,axis='columns')
df.apply(pd.Series.min)
df.apply(lambda x: x.min())
df.applymap(len)
df.applymap(str.lower)
df['first'].map({'Corey':'Chris','Jane':'Mary'}) # map all cell
df['first'].replace({'Corey':'Chris','Jane':'Mary'})
df['first'] = df['first'].replace({'Corey':'Chris','Jane':'Mary'})
df
df = pd.read_csv('survey_results_public.csv',index_col='Respondent')
schema_df = pd.read_csv('survey_results_schema.csv',index_col='Column')
pd.set_option('display.max_columns',10)
pd.set_option('display.max_rows',10)
df.head()
df.rename(columns={'Student':'People'})
df.rename(columns={'Student':'People'},inplace=True)
df
df['People']
df['Hobby']
df['Hobby'].map({'Yes':True,'No':False})
df['Hobby'] = df['Hobby'].map({'Yes':True,'No':False})
df
| 05-Machine-Learning-Code/数据分析工具/Pandas/.ipynb_checkpoints/5_update_rows_columns-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <NAME> --- October 2020
import torch
import torch.nn as nn
import numpy as np
import random
import gym
import matplotlib.pyplot as plt
import seaborn as sns
import math
import torch.optim as optim
from gym import spaces, logger
from gym.utils import seeding
from torch.utils.tensorboard import SummaryWriter
import datetime
import time
import pickle
class BanditEnv(gym.Env):
'''
Toy env to test your implementation
The state is fixed (bandit setup)
Action space: gym.spaces.Discrete(10)
Note that the action takes integer values
'''
def __init__(self):
self.action_space = gym.spaces.Discrete(10)
self.observation_space = gym.spaces.Box(low=np.array([-1]), high=np.array([1]), dtype=np.float32)
def reset(self):
return np.array([0])
def step(self, action):
assert int(action) in self.action_space
done = True
s = np.array([0])
r = -(action - 7)**2
info = {}
return s, r, done, info
class Policy(nn.Module):
def __init__(self, num_inputs, num_actions):
super(Policy, self).__init__()
self.layers = nn.Sequential(
nn.Linear(num_inputs, 16),
nn.ReLU(),
nn.Linear(16, 16),
nn.ReLU(),
nn.Linear(16, 16),
nn.ReLU(),
nn.Linear(16, num_actions),
nn.Softmax(dim=0)
)
self.layers.apply(self.init_weights)
self.num_inputs = num_inputs
self.num_actions = num_actions
def init_weights(self, m):
if type(m) == nn.Linear:
torch.nn.init.xavier_uniform_(m.weight.data)
def forward(self, x):
return self.layers(torch.from_numpy(np.asarray(x)).float())
def act(self, state):
actions = list(range(int(self.num_actions)))
probabilities = self.forward(state)
act = np.random.choice(actions, p=probabilities.detach().numpy())
return act
class ValueFunction(nn.Module):
def __init__(self, num_inputs):
super(ValueFunction, self).__init__()
self.layers = nn.Sequential(
nn.Linear(num_inputs, 32),
nn.ReLU(),
nn.Linear(32, 32),
nn.ReLU(),
nn.Linear(32, 1)
)
self.num_inputs = num_inputs
def forward(self, x):
return self.layers(torch.from_numpy(np.asarray(x)).float())
# # HW3-P2 "REINFORCE" on BanditEnv
# +
NUM_ITERATIONS = 15000
NUM_EVALUATION_ROLLOUTS = 100
EVALUATION_PERIOD = 100
GAMMA = 0.99
LEARNING_RATE = .0001
# +
random.seed(17)
np.random.seed(17)
torch.manual_seed(17)
env = BanditEnv()
env.action_space.seed(17)
env.seed(17)
# +
performance = np.zeros((NUM_EVALUATION_ROLLOUTS,math.ceil(NUM_ITERATIONS/EVALUATION_PERIOD)))
pi = Policy(env.observation_space.shape[0], env.action_space.n)
optimizer = optim.Adam(pi.parameters(),lr=LEARNING_RATE)
now = datetime.datetime.now()
writer = SummaryWriter("runs/scalar-{}".format(now.strftime("%Y%m%d-%H%M%S")))
# 1 iteration == deploying a fixed stationary stochastic policy in the environment for 1 rollout until
# termination and then using the collected data from that rollout to perform 1 policy gradient update
for iteration in range(NUM_ITERATIONS):
history = []
done = False
state = env.reset()
while not done:
action = pi.act(state)
next_state, reward, done, info = env.step(action)
history.append([state, action, reward, next_state, done])
state = next_state
# Sweep backwards through the collected data and use sample_return to accumulate the rewards
# to form the Monte Carlo estimates of the returns
sample_return = 0
loss = 0
optimizer.zero_grad()
for entry in reversed(history):
state, action, reward, next_state, done = entry
sample_return = reward + (GAMMA * sample_return)
loss -= torch.log(pi(state)[action])*(sample_return) / len(history)
# Take a policy gradient step
loss.backward()
optimizer.step()
# Evaluate current policy occasionaly
if iteration % EVALUATION_PERIOD == 0:
evaluation_returns = np.zeros(NUM_EVALUATION_ROLLOUTS)
for test_index in range(NUM_EVALUATION_ROLLOUTS):
done = False
evaluation_return = 0
state = env.reset()
while not done:
action = pi.act(state)
next_state, reward, done, info = env.step(action)
state = next_state
evaluation_return += reward # Instructor response on Piazza post @457 asks to evaluate without discounting.
evaluation_returns[test_index] = evaluation_return
writer.add_scalar('Expected Return (BanditEnv)', np.mean(evaluation_returns), iteration)
writer.flush()
performance[:,int(iteration/EVALUATION_PERIOD)] = evaluation_returns
print('Iteration: ', iteration,'\t', 'Evaluation: ', np.mean(evaluation_returns))
writer.close()
# -
sns.lineplot(x=list(range(0,NUM_ITERATIONS,EVALUATION_PERIOD))*performance.shape[0], y=performance.flatten(),ci='sd')
plt.title('REINFORCE; BanditsEnv;\nMean and S.D.')
plt.xlabel('Iteration')
plt.ylabel('Evaluation Return')
# # HW3-P2 "REINFORCE" on LunarLandar-v2
# +
NUM_ITERATIONS = 60000
NUM_EVALUATION_ROLLOUTS = 100
EVALUATION_PERIOD = 100
GAMMA = 0.99
LEARNING_RATE = .0001
# +
random.seed(17)
np.random.seed(17)
torch.manual_seed(17)
env = gym.make('LunarLander-v2')
env.action_space.seed(17)
env.seed(17)
# +
performance = np.zeros((NUM_EVALUATION_ROLLOUTS,math.ceil(NUM_ITERATIONS/EVALUATION_PERIOD)))
pi = Policy(env.observation_space.shape[0], env.action_space.n)
optimizer = optim.Adam(pi.parameters(),lr=LEARNING_RATE)
now = datetime.datetime.now()
writer = SummaryWriter("runs/scalar-{}".format(now.strftime("%Y%m%d-%H%M%S")))
# 1 iteration == deploying a fixed stationary stochastic policy in the environment for 1 rollout until
# termination and then using the collected data from that rollout to perform 1 policy gradient update
for iteration in range(NUM_ITERATIONS):
history = []
done = False
state = env.reset()
while not done:
action = pi.act(state)
next_state, reward, done, info = env.step(action)
history.append([state, action, reward, next_state, done])
state = next_state
# Sweep backwards through the collected data and use sample_return to accumulate the rewards
# to form the Monte Carlo estimates of the returns
sample_return = 0
loss = 0
optimizer.zero_grad()
for entry in reversed(history):
state, action, reward, next_state, done = entry
sample_return = reward + (GAMMA * sample_return)
loss -= torch.log(pi(state)[action])*(sample_return) / len(history)
# Take a policy gradient step
loss.backward()
optimizer.step()
# Evaluate current policy occasionaly
if iteration % EVALUATION_PERIOD == 0:
evaluation_returns = np.zeros(NUM_EVALUATION_ROLLOUTS)
for test_index in range(NUM_EVALUATION_ROLLOUTS):
done = False
evaluation_return = 0
state = env.reset()
while not done:
action = pi.act(state)
next_state, reward, done, info = env.step(action)
state = next_state
evaluation_return += reward # Instructor response on Piazza post @457 asks to evaluate without discounting.
evaluation_returns[test_index] = evaluation_return
writer.add_scalar('Expected Return (Lunar Lander) (Extended Run)', np.mean(evaluation_returns), iteration)
writer.flush()
performance[:,int(iteration/EVALUATION_PERIOD)] = evaluation_returns
print('Iteration: ', iteration,'\t', 'Evaluation: ', np.mean(evaluation_returns))
writer.close()
# -
sns.lineplot(x=list(range(0,NUM_ITERATIONS,EVALUATION_PERIOD))*performance.shape[0], y=performance.flatten(),ci='sd')
plt.title('REINFORCE; LunarLander-v2;\nMean and S.D.')
plt.xlabel('Iteration')
plt.ylabel('Evaluation Return')
plt.savefig('REINFORCE_LL.png',dpi=300)
# # HW3-P2 "REINFORCE with Value-Function Baseline" on BanditEnv
# +
NUM_ITERATIONS = 15000
NUM_EVALUATION_ROLLOUTS = 100
EVALUATION_PERIOD = 100
GAMMA = 0.99
LEARNING_RATE = .0001
LEARNING_RATE_V = .0001
# +
random.seed(17)
np.random.seed(17)
torch.manual_seed(17)
env = BanditEnv()
env.action_space.seed(17)
env.seed(17)
# +
performance_b = np.zeros((NUM_EVALUATION_ROLLOUTS,math.ceil(NUM_ITERATIONS/EVALUATION_PERIOD)))
pi_b = Policy(env.observation_space.shape[0], env.action_space.n)
optimizer = optim.Adam(pi_b.parameters(),lr=LEARNING_RATE)
v = ValueFunction(env.observation_space.shape[0])
optimizer_v = optim.Adam(v.parameters(),lr=LEARNING_RATE_V)
now = datetime.datetime.now()
writer = SummaryWriter("runs/scalar-{}".format(now.strftime("%Y%m%d-%H%M%S")))
# 1 iteration == deploying a fixed stationary stochastic policy in the environment for 1 rollout until
# termination and then using the collected data from that rollout to perform 1 policy gradient update
for iteration in range(NUM_ITERATIONS):
history = []
done = False
state = env.reset()
while not done:
action = pi_b.act(state)
next_state, reward, done, info = env.step(action)
history.append([state, action, reward, next_state, done])
state = next_state
# Sweep backwards through the collected data and use sample_return to accumulate the rewards
# to form the Monte Carlo estimates of the returns
sample_return = 0
loss = 0
optimizer.zero_grad()
# Update Value-function first
for entry in reversed(history):
state, action, reward, next_state, done = entry
sample_return = reward + (GAMMA * sample_return)
optimizer_v.zero_grad()
loss_v = (sample_return - v(state))**2
loss_v.backward()
optimizer_v.step()
# Calculate loss with value function baseline
for entry in reversed(history):
state, action, reward, next_state, done = entry
sample_return = reward + (GAMMA * sample_return)
loss -= torch.log(pi_b(state)[action])*(sample_return - v(state)) / len(history)
# Take a policy gradient step
loss.backward()
optimizer.step()
# Evaluate current policy occasionaly
if iteration % EVALUATION_PERIOD == 0:
evaluation_returns = np.zeros(NUM_EVALUATION_ROLLOUTS)
for test_index in range(NUM_EVALUATION_ROLLOUTS):
done = False
evaluation_return = 0
state = env.reset()
while not done:
action = pi_b.act(state)
next_state, reward, done, info = env.step(action)
state = next_state
evaluation_return += reward # Instructor response on Piazza post @457 asks to evaluate without discounting.
evaluation_returns[test_index] = evaluation_return
writer.add_scalar('Expected Return (BanditEnv; RwB)', np.mean(evaluation_returns), iteration)
writer.flush()
performance_b[:,int(iteration/EVALUATION_PERIOD)] = evaluation_returns
print('Iteration: ', iteration,'\t', 'Evaluation: ', np.mean(evaluation_returns))
writer.close()
# -
sns.lineplot(x=list(range(0,NUM_ITERATIONS,EVALUATION_PERIOD))*performance_b.shape[0], y=performance_b.flatten(),ci='sd')
plt.title('RwB; BanditsEnv;\nMean and S.D.')
plt.xlabel('Iteration')
plt.ylabel('Evaluation Return')
# # HW3-P2 "REINFORCE with Value-Function Baseline" on LunarLander-v2
# +
NUM_ITERATIONS = 60000
NUM_EVALUATION_ROLLOUTS = 100
EVALUATION_PERIOD = 100
GAMMA = 0.99
LEARNING_RATE = .0001
LEARNING_RATE_V = .0001
# +
random.seed(17)
np.random.seed(17)
torch.manual_seed(17)
env = gym.make('LunarLander-v2')
env.action_space.seed(17)
env.seed(17)
# +
performance_b = np.zeros((NUM_EVALUATION_ROLLOUTS,math.ceil(NUM_ITERATIONS/EVALUATION_PERIOD)))
pi_b = Policy(env.observation_space.shape[0], env.action_space.n)
optimizer = optim.Adam(pi_b.parameters(),lr=LEARNING_RATE)
v = ValueFunction(env.observation_space.shape[0])
optimizer_v = optim.Adam(v.parameters(),lr=LEARNING_RATE_V)
now = datetime.datetime.now()
writer = SummaryWriter("runs/scalar-{}".format(now.strftime("%Y%m%d-%H%M%S")))
# 1 iteration == deploying a fixed stationary stochastic policy in the environment for 1 rollout until
# termination and then using the collected data from that rollout to perform 1 policy gradient update
for iteration in range(NUM_ITERATIONS):
history = []
done = False
state = env.reset()
while not done:
action = pi_b.act(state)
next_state, reward, done, info = env.step(action)
history.append([state, action, reward, next_state, done])
state = next_state
# Sweep backwards through the collected data and use sample_return to accumulate the rewards
# to form the Monte Carlo estimates of the returns
sample_return = 0
loss = 0
optimizer.zero_grad()
for entry in reversed(history):
state, action, reward, next_state, done = entry
sample_return = reward + (GAMMA * sample_return)
loss_v += (sample_return - v(state))**2
loss_v.backward()
optimizer_v.step()
for entry in reversed(history):
state, action, reward, next_state, done = entry
sample_return = reward + (GAMMA * sample_return)
loss -= torch.log(pi_b(state)[action])*(sample_return - v(state)) / len(history)
# Take a policy gradient step
loss.backward()
optimizer.step()
# Evaluate current policy occasionaly
if iteration % EVALUATION_PERIOD == 0:
evaluation_returns = np.zeros(NUM_EVALUATION_ROLLOUTS)
for test_index in range(NUM_EVALUATION_ROLLOUTS):
done = False
evaluation_return = 0
state = env.reset()
while not done:
action = pi_b.act(state)
next_state, reward, done, info = env.step(action)
state = next_state
evaluation_return += reward # Instructor response on Piazza post @457 asks to evaluate without discounting.
evaluation_returns[test_index] = evaluation_return
writer.add_scalar('Expected Return (Lunar Lander; RwB)', np.mean(evaluation_returns), iteration)
writer.flush()
performance_b[:,int(iteration/EVALUATION_PERIOD)] = evaluation_returns
print('Iteration: ', iteration,'\t', 'Evaluation: ', np.mean(evaluation_returns))
writer.close()
# -
sns.lineplot(x=list(range(0,NUM_ITERATIONS,EVALUATION_PERIOD))*performance_b.shape[0], y=performance_b.flatten(),ci='sd')
plt.title('RwB; LunarLander-v2;\nMean and S.D.')
plt.xlabel('Iteration')
plt.ylabel('Evaluation Return')
| examples/Problem_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import xarray as xr
import pandas as pd
import numpy as np
from cast_to_xarray import *
# +
winklerpd1 = pd.read_csv('../O2calib/Cruise1Winkler.csv')
winkler1 = winklerpd1.set_index('Station').to_xarray()
winklerpd2 = pd.read_csv('../O2calib/Cruise2Winkler.csv')
winkler2 = winklerpd2.set_index('Station').to_xarray()
# -
winkler1
# match the CTD station name with the winkler station name
CTD1_RACE = cast_to_xarray('data_cruise1/TheRace_loop_filter_teos10_bin.cnv', 'Race')
CTD1_5 = cast_to_xarray('data_cruise1/Station5_loop_filter_teos10_bin.cnv', '5')
CTD1_WF = cast_to_xarray('data_cruise1/StationWF_loop_filter_teos10_bin.cnv', 'WF')
CTD1_4 = cast_to_xarray('data_cruise1/Station4_TSwift_loop_filter_teos10_bin.cnv', '4')
# +
#CTD1_RACE
# -
# combine all the casts into one xarray dataset
ctdsection = xr.concat([CTD1_RACE, CTD1_5, CTD1_WF, CTD1_4],"station")
ctdsection
# +
# the first station is the race
stnname = 'Race'
# get the winkler depths
deptharray = winkler1.sel(Station=stnname).Depth.values
# get the corresponding CTD data
stnsel = ctdsection.sel(station=stnname)
CTDval = stnsel.sel(depth=deptharray, method='nearest').oxygen.values
# +
# do this in a loop
# make an array that combines the data
stnnames = ['Race', '5', 'WF', '4']
CTD_QC=[]
for stnname in stnnames:
# get the winkler depths
deptharray = winkler1.sel(Station=stnname).Depth.values
# get the corresponding CTD data
stnsel = ctdsection.sel(station=stnname)
for i in range(len(deptharray)):
CTDval = stnsel.sel(depth=deptharray, method='nearest').oxygen.values[i]
CTD_QC.append(CTDval)
# -
#CTDval
CTD_QC
Wval=winkler1.Oxygen.values
plt.scatter(CTD_QC, Wval)
# +
correlation_matrix = np.corrcoef(CTD_QC, Wval)
correlation_xy = correlation_matrix[0,1]
r_squared = correlation_xy**2
r_squared
# -
| oxygen_QC_combined.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <small><small><i>
# All the IPython Notebooks in this lecture series by Dr. <NAME> are available @ **[GitHub](https://github.com/milaan9/02_Python_Datatypes/tree/main/006_Python_Sets_Methods)**
# </i></small></small>
# # Python Set `intersection()`
#
# The **`intersection()`** method returns a new set with elements that are common to all sets.
#
# The intersection of two or more sets is the set of elements that are common to all sets. For example:
# ```python
# A = {1, 2, 3, 4}
# B = {2, 3, 4, 9}
# C = {2, 4, 9 10}
#
# Then,
# A∩B = B∩A ={2, 3, 4}
# A∩C = C∩A ={2, 4}
# B∩C = C∩B ={2, 4, 9}
#
# A∩B∩C = {2, 4}
# ```
# <div>
# <img src="img/intersection.png" width="250"/>
# </div>
# **Syntax**:
#
# ```python
# A.intersection(*other_sets)
# ```
# ## `intersection()` Parameters
#
# The **`intersection()`** method allows arbitrary number of arguments (sets).
#
# >**Note:** __`*`__ is not part of the syntax. It is used to indicate that the method allows arbitrary number of arguments.
# ## Return Value from `intersection()`
#
# The **`intersection()`** method returns the intersection of set A with all the sets (passed as argument).
#
# If the argument is not passed to **`intersection()`**, it returns a shallow copy of the set **`(A)`**.
# +
# Example 1: How intersection() works?
A = {2, 3, 5, 4}
B = {2, 5, 100}
C = {2, 3, 8, 9, 10}
print(B.intersection(A))
print(B.intersection(C))
print(A.intersection(C))
print(C.intersection(A, B))
# +
# Example 2:
A = {100, 7, 8}
B = {200, 4, 5}
C = {300, 2, 3}
D = {100, 200, 300}
print(A.intersection(D))
print(B.intersection(D))
print(C.intersection(D))
print(A.intersection(B, C, D))
# -
# You can also find the intersection of sets using **`&`** operator.
# +
# Example 3: Set Intersection Using & operator
A = {100, 7, 8}
B = {200, 4, 5}
C = {300, 2, 3, 7}
D = {100, 200, 300}
print(A & C)
print(A & D)
print(A & C & D)
print(A & B & C & D)
# -
| 006_Python_Sets_Methods/008_Python_Set_intersection().ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jonkrohn/ML-foundations/blob/master/notebooks/5-probability.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="aTOLgsbN69-P"
# # Probability & Information Theory
# + [markdown] id="yqUB9FTRAxd-"
# This class, *Probability & Information Theory*, introduces the mathematical fields that enable us to quantify uncertainty as well as to make predictions despite uncertainty. These fields are essential because machine learning algorithms are both trained by imperfect data and deployed into noisy, real-world scenarios they haven’t encountered before.
#
# Through the measured exposition of theory paired with interactive examples, you’ll develop a working understanding of variables, probability distributions, metrics for assessing distributions, and graphical models. You’ll also learn how to use information theory to measure how much meaningful signal there is within some given data. The content covered in this class is itself foundational for several other classes in the *Machine Learning Foundations* series, especially *Intro to Statistics* and *Optimization*.
# + [markdown] id="d4tBvI88BheF"
# Over the course of studying this topic, you'll:
#
# * Develop an understanding of what’s going on beneath the hood of predictive statistical models and machine learning algorithms, including those used for deep learning.
# * Understand the appropriate variable type and probability distribution for representing a given class of data, as well as the standard techniques for assessing the relationships between distributions.
# * Apply information theory to quantify the proportion of valuable signal that’s present amongst the noise of a given probability distribution.
#
# + [markdown] id="Z68nQ0ekCYhF"
# **Note that this Jupyter notebook is not intended to stand alone. It is the companion code to a lecture or to videos from <NAME>'s [Machine Learning Foundations](https://github.com/jonkrohn/ML-foundations) series, which offer detail on the following:**
#
# *Segment 1: Introduction to Probability*
# * What Probability Theory Is
# * A Brief History: Frequentists vs Bayesians
# * Applications of Probability to Machine Learning
# * Random Variables
# * Discrete vs Continuous Variables
# * Probability Mass and Probability Density Functions
# * Expected Value
# * Measures of Central Tendency: Mean, Median, and Mode
# * Quantiles: Quartiles, Deciles, and Percentiles
# * The Box-and-Whisker Plot
# * Measures of Dispersion: Variance, Standard Deviation, and Standard Error
# * Measures of Relatedness: Covariance and Correlation
# * Marginal and Conditional Probabilities
# * Independence and Conditional Independence
#
# *Segment 2: Distributions in Machine Learning*
# * Uniform
# * Gaussian: Normal and Standard Normal
# * The Central Limit Theorem
# * Log-Normal
# * Exponential and Laplace
# * Binomial and Multinomial
# * Poisson
# * Mixture Distributions
# * Preprocessing Data for Model Input
#
# *Segment 3: Information Theory*
# * What Information Theory Is
# * Self-Information
# * Nats, Bits and Shannons
# * Shannon and Differential Entropy
# * Kullback-Leibler Divergence
# * Cross-Entropy
# + [markdown] id="HDE74CXX5ChI"
# ## Segment 1: Introduction to Probability
# + id="kzRpCm2a5ChJ"
import numpy as np
import scipy.stats as st
import matplotlib.pyplot as plt
import seaborn as sns
# + [markdown] id="U_680ypO5ChJ"
# ### What Probability Theory Is
# + [markdown] id="uqzuaq1oD1XV"
# #### Events and Sample Spaces
# + [markdown] id="fwEAqIqR5ChJ"
# Let's assume we have a fair coin, which is equally likely to come up heads (H) or tails (T).
# + [markdown] id="kFl3g0GW5ChK"
# In instances like this, where the two outcomes are equally likely, we can use probability theory to express the likelihood of a particular **event** by comparing it with the **sample space** (the set of all possible outcomes; can be denoted as $\Omega$):
# + [markdown] id="X82RIGz15ChK"
# $$ P(\text{event}) = \frac{\text{# of outcomes of event}}{\text{# of outcomes in }\Omega} $$
# + [markdown] id="VuVWT8NI5ChK"
# If we're only flipping the coin once, then there are only two possible outcomes in the sample space $\Omega$: it will either be H or T (using set notation, we could write this as $\Omega$ = {H, T}).
# + [markdown] id="9W5MsKt15ChK"
# Therefore: $$ P(H) = \frac{1}{2} = 0.5 $$
# + [markdown] id="V_bzjMvs5ChK"
# Equally: $$ P(T) = \frac{1}{2} = 0.5 $$
# + [markdown] id="PfHEQzcd5ChL"
# As a separate example, consider drawing a single card from a standard deck of 52 playing cards. In this case, the number of possible outcomes in the sample space $\Omega$ is 52.
# + [markdown] id="qtAa3aRW5ChL"
# There is only one ace of spades in the deck, so the probability of drawing it is: $$ P(\text{ace of spades}) = \frac{1}{52} \approx 0.019 $$
# + [markdown] id="81-uQ4ug5ChL"
# In contrast there are four aces, so the probability of drawing an ace is: $$ P(\text{ace}) = \frac{4}{52} \approx 0.077 $$
# + [markdown] id="uqco79Cd5ChL"
# Some additional examples:
# $$ P(\text{spade}) = \frac{13}{52} = 0.25 $$
# $$ P(\text{ace OR spade}) = \frac{16}{52} \approx 0.307 $$
# $$ P(\text{card}) = \frac{52}{52} = 1 $$
# $$ P(\text{turnip}) = \frac{0}{52} = 0 $$
# + [markdown] id="snINg1tz5ChL"
# #### Multiple Independent Observations
# + [markdown] id="gIPR6og95ChL"
# Let's return to coin flipping to illustrate situations where we have an event consisting of multiple independent observations. For example, the probability of throwing two consecutive heads is: $$ P(\text{HH}) = \frac{1}{4} = 0.25 $$ ...because there is one HH event in the sample set of four possible events ($\Omega$ = {HH, HT, TH, TT}).
# + [markdown] id="rT11_oHg5ChM"
# Likewise, the probability of throwing *three* consecutive heads is: $$ P(\text{HHH}) = \frac{1}{8} = 0.125 $$ ...because there is one HHH event in the sample set of eight possible events ($\Omega$ = {HHH, HHT, HTH, THH, HTT, THT, TTH, TTT}).
# + [markdown] id="ru44PKUe5ChM"
# As final examples, the probability of throwing exactly two heads in three tosses is $ P = \frac{3}{8} = 0.375 $ while the probability of throwing at least two heads in three tosses is $ P = \frac{4}{8} = 0.5 $.
# + [markdown] id="vh3WnRhm5ChM"
# #### Combining Probabilities
# + [markdown] id="YInbLu1i5ChM"
# In order to combine probabilities, we can multiply them. So the probability of throwing five consecutive heads, for example, is the product of probabilities we've already calculated: $$ P(\text{HHHHH}) = P(\text{HH}) \times P(\text{HHH}) = \frac{1}{4} \times \frac{1}{8} = \frac{1}{32} \approx 0.031 $$
# + [markdown] id="w-wlHpI05ChM"
# #### Combinatorics
# + [markdown] id="1ckSVU3p5ChM"
# *Combinatorics* is a field of mathematics devoted to counting that can be helpful to studying probabilities. We can use **factorials** (e.g., $4! = 4 \times 3 \times 2 \times 1 = 24$), which feature prominently in combinatorics, to calculate probabilities instead of painstakingly determining all of the members of the sample space $\Omega$ and counting subsets within $\Omega$.
# + [markdown] id="zYL4ODP75ChN"
# More specifically, we can calculate the number of outcomes of an event using the "number of combinations" equation: $$ {n \choose k} = \frac{n!}{k!(n - k)!} $$
# + [markdown] id="kQhpdNEL5ChN"
# The left-hand side of the equation is read "$n$ choose $k$" and is most quickly understood via an example: If we have three coin flips, $n = 3$, and if we're interested in the number of ways to get two head flips (or two tail flips, for that matter), $k = 2$. We would read this as "3 choose 2" and calculate it as:
# $$ {n \choose k} = {3 \choose 2} = \frac{3!}{2!(3 - 2)!} = \frac{3!}{(2!)(1!)} = \frac{3 \times 2 \times 1}{(2 \times 1)(1)} = \frac{6}{(2)(1)} = \frac{6}{2} = 3 $$
# + [markdown] id="PoPfNW275ChN"
# This provide us with the numerator for event-probability equation from above: $$ P(\text{event}) = \frac{\text{# of outcomes of event}}{\text{# of outcomes in }\Omega} $$
# + [markdown] id="ThoyCpl35ChO"
# In the case of coin-flipping (or any binary process with equally probable outcomes), the denominator can be calculated with $2^n$ (where $n$ is again the number of coin flips), so: $$ \frac{\text{# of outcomes of event}}{\text{# of outcomes in }\Omega} = \frac{3}{2^n} = \frac{3}{2^3} = \frac{3}{8} = 0.375 $$
# + [markdown] id="QKRugWrQ5ChO"
# **Exercises**:
#
# 1. What is the probability of drawing the ace of spades twice in a row? (Assume that any card drawn on the first draw will be put back in the deck before the second draw.)
# 2. You draw a card from a deck of cards. After replacing the drawn card back in the deck and shuffling thoroughly, what is the probability of drawing the same card again?
# 3. Use $n \choose k$ to calculate the probability of throwing three heads in five coin tosses.
# 4. Create a Python method that solves exercise 3 and incorporates the $n \choose k$ formula $\frac{n!}{k!(n - k)!}$. With the method in hand, calculate the probability of -- in five tosses -- throwing each of zero, one, two, three, four, and five heads.
# + [markdown] id="X1d1mBjR5ChO"
# **Spoiler alert**: Solutions are below so scroll carefully...
# + id="_ZzjXjHy5ChO"
# + id="sbq5esRv5ChO"
# + id="ru1t8_LRajrP"
# + id="1UGIpLpbajZf"
# + id="oPtm03U95ChO"
# + [markdown] id="U7IgNJwV5ChO"
# **Solutions**:
# + [markdown] id="qiSjQ1mR5ChP"
# 1. $$ P(\text{ace of spades}) \times P(\text{ace of spades}) = \left(\frac{1}{52}\right)^2 = \frac{1}{2704} = 0.00037 = 0.037\% $$
# + [markdown] id="R3aUyZvi5ChP"
# 2. $$ P(\text{any card}) = \frac{52}{52} = 1 $$
# $$ P(\text{same card as first draw}) = \frac{1}{52} \approx 0.019 $$
# $$ P(\text{any card})P(\text{same card as first draw}) = (1)(\frac{1}{52}) = \frac{1}{52} \approx 0.019$$
# + [markdown] id="S8tA4XlQ5ChP"
# 3. $$ {n \choose k} = {5 \choose 3} = \frac{5!}{3!(5 - 3)!} = \frac{5!}{(3!)(2!)} = \frac{5 \times 4 \times 3 \times 2 \times 1}{(3 \times 2 \times 1)(2 \times 1)} = \frac{120}{(6)(2)} = \frac{120}{12} = 10 $$
# + [markdown] id="aS4bc83U5ChP"
# $$P = \frac{10}{2^n} = \frac{10}{2^5} = \frac{10}{32} = 0.3125 $$
# + id="0_3pRRTX5ChP"
from math import factorial
# + id="MgdCyK805ChP"
def coinflip_prob(n, k):
n_choose_k = factorial(n)/(factorial(k)*factorial(n-k))
return n_choose_k/2**n
# + id="4N3_Ebkum1Vh" outputId="89b59a2b-715c-4b69-cdfe-948e695918b7" colab={"base_uri": "https://localhost:8080/"}
coinflip_prob(5, 3)
# + id="6UO8q5N45ChQ" colab={"base_uri": "https://localhost:8080/"} outputId="0268c14d-c28f-42cf-cbd2-090c4702c411"
[coinflip_prob(5, h) for h in range(6)]
# + [markdown] id="gjSkHJ8r5ChQ"
# #### The Law of Large Numbers
# + [markdown] id="U9NDLpfo5ChR"
# While a fair coin should land heads up 50% of the time, as we've seen above, with small sample sizes, there is a non-trivial possibility that in a given experiment we could flip heads on all of the tosses. For example, we've calculated that there's a 3.1% chance that we'll get heads on every toss in a small five-toss experiment.
# + [markdown] id="7rfVQfP55ChR"
# The **law of large numbers** states that the more experiments we run, the closer we will tend to get to the expected probability.
# + [markdown] id="7A3Va-dB5ChR"
# Let's run some code to examine this in practice. To start, we'll create a vector of exponentially increasing $n$umbers of coin tosses per experiment:
# + id="ymsouctT5ChR"
ns = np.array([2, 4, 8, 16, 32, 64, 128, 256, 512, 1024, 2048, 4096])
# + id="hgmoAn2g5ChR"
np.random.seed(42) # for reproducibility
# + [markdown] id="hKtZzZKK5ChR"
# We will discuss the `binomial()` method in more detail in *Segment 2*. For now it suffices to think of its two arguments as *number of coin flips in experiment* and *probability of heads*, while it returns the number of flips that are heads in the experiment.
# + id="xRQh-0iG5ChS" colab={"base_uri": "https://localhost:8080/"} outputId="0d7a21c0-b715-4ffa-b893-2854bdd0cbab"
np.random.binomial(1, 0.5)
# + id="OwfWWhFm5ChS" colab={"base_uri": "https://localhost:8080/"} outputId="e6146ce6-748d-4012-b3d7-1eb8b9048508"
heads_count = [np.random.binomial(n, 0.5) for n in ns]
heads_count
# + id="rL4PbNmY5ChS" colab={"base_uri": "https://localhost:8080/"} outputId="ad56af0f-9bd3-4031-af21-02d087787e71"
proportion_heads = heads_count/ns
proportion_heads
# + id="vlz7a5DU5ChS" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="8e306028-59a4-4d48-8528-5d07b39981ec"
fig, ax = plt.subplots()
plt.xlabel('Number of coin flips in experiment')
plt.ylabel('Proportion of flips that are heads')
plt.axhline(0.5, color='orange')
_ = ax.scatter(ns, proportion_heads)
# + [markdown] id="RvrI0woM5ChS"
# It is important at this juncture to address the **gambler's fallacy**. It is a common misconception that the law of large numbers dictates that if, say, five heads have been flipped in a row, then the probability of tails is higher on the sixth flip. In fact, probability theory holds that each coin flip is completely independent of all others. Thus, every single flip of a fair coin has a 50% chance of being heads, no matter what happened on preceding flips.
# + [markdown] id="j2-wA_6E5ChT"
# (To capitalize on this misunderstanding, roulette tables at casinos often have prominent displays tracking the history of red versus black even though there's a 47.4% chance of each on every spin of the roulette wheel no matter what happened on preceding spins. Gamblers will note, say, five reds in a row at a given table and flock to it to bet on black.)
# + [markdown] id="hKciO43C5ChT"
# #### Statistics
# + [markdown] id="WqitDIkk5ChT"
# The field of statistics applies probability theory to make inferences with a quantifiable degree of confidence. For example, let's say we ran the five-coin-flip experiment 1000 times with a fair coin:
# + id="Nvc9rCnq5ChT"
n_experiments = 1000
heads_count = np.random.binomial(5, 0.5, n_experiments)
# + [markdown] id="siW-zTqm5ChT"
# We can plot the results of our experiment to create a **probability distribution**:
# + id="FRiZSpwy5ChT"
heads, event_count = np.unique(heads_count, return_counts=True)
# + id="8TEmy-_c5ChT" colab={"base_uri": "https://localhost:8080/"} outputId="dc23d929-72c2-4371-9236-324642aa8891"
heads
# + id="Yn5wlP-A5ChU" colab={"base_uri": "https://localhost:8080/"} outputId="2fc0aaf2-8a08-4798-d5d1-1ec67637c442"
event_count
# + id="LaBGlw-95ChU" outputId="402835b3-8666-4656-ffd9-fe1d2a20e331" colab={"base_uri": "https://localhost:8080/"}
event_proba = event_count/n_experiments
event_proba
# + id="ecU6IMfE5ChU" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="f4a46c76-f3b0-4ea3-c235-8de91742f7b9"
plt.bar(heads, event_proba, color='mediumpurple')
plt.xlabel('Heads flips (out of 5 tosses)')
_ = plt.ylabel('Event probability')
# + [markdown] id="KwhO9Zn75ChU"
# Let's say we'd like to now test a coin as to whether it's fair or not. We flip it five times and all five times it comes up heads. Does this imply it's not a fair coin? Statistics arms us with tools to use the probability distribution above to answer this question with a particular degree of confidence.
# + [markdown] id="AENw26_O5ChU"
# **Return to slides here.**
# + [markdown] id="LMC_Cvs85ChV"
# ### Expected Value
# + id="XOyKNFZw5ChV" colab={"base_uri": "https://localhost:8080/"} outputId="8c944f76-706b-4d23-ce7a-5b31736d2305"
P = [coinflip_prob(5, x) for x in range(6)]
P
# + [markdown] id="XCCclUvR5ChV"
# From the slides:
# $$ \mathbb{E} = \sum_x xP(x) $$
# + id="37gjROzg5ChV" colab={"base_uri": "https://localhost:8080/"} outputId="fef29d34-1027-4db5-c717-52ac399679bf"
E = sum([P[x]*x for x in range(6)])
E
# + [markdown] id="xOq8r0Ud5ChV"
# **Exercises**:
#
# 1. Assuming heads = 1 and tails = 0, what’s the expected value of the toss of a fair coin?
# 2. ...and of two tosses of a fair coin?
# 3. You are one of a thousand people to purchase a lottery ticket. The ticket costs \\$10 and pays out \\$5000. What value should you expect from this transaction?
#
# + [markdown] id="uziVFQUR6A4x"
# **Spoiler Alert**: Solutions below
# + id="vsl_1I3n5ChV"
# + id="sZLZtHsN5ChV"
# + id="q2Th7Qb-5ChW"
# + [markdown] id="ZYyeFfVW5ChW"
# **Solutions**:
#
# 1. (½)0 + (½)1 = 0 + ½ = ½
# 2. (¼)0 + (½)1 + (¼)2 = 0 + ½ + ½ = 1
#
# 3.
# (1/1000)(\\$5000-\\$10) + (999/1000)(-\\$10) \
# = (1/1000)(\\$4990) + (999/1000)(-\\$10) \
# = \\$4.99 + (\\$-9.99) \
# = -\\$5
#
# + [markdown] id="IwCzvbE05ChW"
# ### Measures of Central Tendency
# + [markdown] id="Nz_RAtEf5ChW"
# Measures of central tendency provide a summary statistic on the center of a given distribution, a.k.a., the "average" value of the distribution.
# + [markdown] id="HrAt0oVb5ChW"
# #### Mean
# + [markdown] id="pCKs9uFF5ChW"
# The most common measure of central tendency, synonomous with the term "average", is the **mean**, often symbolized with $\mu$ (population) or $\bar{x}$ (sample):
# + [markdown] id="sBIObqs05ChW"
# $$ \bar{x} = \frac{\sum_{i=1}^n x_i}{n} $$
# + [markdown] id="-O0hmQam5ChX"
# Expected value is in fact the long-term *mean* of some function (i.e., $\mu = \mathbb{E}$). Let's calculate how close the sample mean, $\bar{x}$, of our five-coin-flip experiments comes to the expected value, $\mathbb{E} = 2.5$.
# + id="fJY0mexL5ChX" colab={"base_uri": "https://localhost:8080/"} outputId="7d1c0f46-d7a2-4d91-e332-a889125c3cc0"
len(heads_count)
# + id="sU3-c-EJ5ChX" colab={"base_uri": "https://localhost:8080/"} outputId="e4218377-3664-4b4c-bd39-ea4f666c0316"
heads_count[0:20]
# + id="pA0ai4I45ChX" colab={"base_uri": "https://localhost:8080/"} outputId="7f1de0ca-22ac-40d6-8104-058eba801b75"
sum(heads_count)/len(heads_count)
# + [markdown] id="fN9wBEb55ChX"
# Unsurprisingly, NumPy comes with a built-in function:
# + id="OXRi9fki-5CB" outputId="bb58234f-053b-4018-a41f-d66bc95262b9" colab={"base_uri": "https://localhost:8080/"}
np.mean(heads_count)
# + id="4c6p-UVK-aok" outputId="bbd05a09-8f61-4139-f955-265501674459" colab={"base_uri": "https://localhost:8080/"}
heads_count.mean() # fun!
# + [markdown] id="vD8bvjix5ChY"
# Pretty close! The law of large numbers implies that as we increase the number of experiments (e.g., to a million -- you're welcome to try it), we'll converge on $\mathbb{E}=2.5$.
# + [markdown] id="Q1qtow1O5ChY"
# #### Median
# + [markdown] id="4TZHQsES5ChY"
# The second most common measure of central tendency is the **median**, the midpoint value in the distribution:
# + id="Ff3aQam65ChY"
heads_count.sort()
# + id="0ECSOjaA5ChY" colab={"base_uri": "https://localhost:8080/"} outputId="806d020a-98b3-4fc5-8e4c-079bb985e32b"
heads_count[0:20]
# + id="ySeyJ81I5ChY" colab={"base_uri": "https://localhost:8080/"} outputId="6d05fde2-4ecc-4ca3-ae0b-7827f8d36bfd"
heads_count[-20:]
# + id="SdeMf4RR5ChZ" colab={"base_uri": "https://localhost:8080/"} outputId="59906afc-e8c8-4be9-ee6d-30456acbae30"
len(heads_count)
# + [markdown] id="WWfqQ4rI5ChZ"
# With an odd number of values in a distribution, we consider the single midpoint to be the median, e.g., the midpoint of the sorted array `[2, 4, 6, 9, 10]` is six.
#
# With an even number, we take the mean of the two values that straddle the midpoint:
# + id="xQPz0RT45ChZ" colab={"base_uri": "https://localhost:8080/"} outputId="381670c8-24b9-49e1-abbd-118a0079798d"
heads_count[499]
# + id="RavYR6JX5ChZ" colab={"base_uri": "https://localhost:8080/"} outputId="320112e3-3485-4bed-d1fc-8e6fde3c074c"
heads_count[500]
# + [markdown] id="o7CLGD4z5ChZ"
# ...which in this case is obviously three:
# + id="MZ6cEUJ25ChZ" colab={"base_uri": "https://localhost:8080/"} outputId="fd6ffaa8-ad24-40f9-a034-dbd665005943"
(3+3)/2
# + [markdown] id="udtzZf_X5Cha"
# Built-in method:
# + id="dxxtSiCp5Cha" colab={"base_uri": "https://localhost:8080/"} outputId="97319c5a-7613-48f0-ce05-e9709163ec92"
np.median(heads_count)
# + [markdown] id="k3Fv1AMu5Cha"
# #### Mode
# + [markdown] id="RiUTOi3U5Cha"
# The **mode** is the final common measure of central tendency. It is simply the value in the distribution that occurs most frequently.
#
# As is clear in the most recently output chart, in the case of our five-coin-toss experiment, the mode is three.
# + [markdown] id="ukKeQScV5Cha"
# Method available in the SciPy `stats` module:
# + id="FPrHiYh65Cha" colab={"base_uri": "https://localhost:8080/"} outputId="a621ecd5-897f-4692-b26e-d81319074b50"
st.mode(heads_count)
# + id="PFUn0iJX5Chb" colab={"base_uri": "https://localhost:8080/"} outputId="b0ef3e17-382b-4b73-896b-ea4675f8e5f1"
st.mode(heads_count)[0][0]
# + [markdown] id="36qtYyry5Chb"
# With small sample sizes, the mean typically provides the most accurate measure of central tendency.
#
# With larger sample sizes, the mean, median, and mode will tend to coincide, as long as the distribution isn't skewed:
# + id="NcU6qKo35Chb"
x = st.skewnorm.rvs(0, size=1000) # first argument is "skewness"; 0 has no skew
# + id="c13nv_wO5Chb" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="ca2f764e-de6c-467f-a9fc-f3519305c27f"
fig, ax = plt.subplots()
plt.axvline(x = np.mean(x), color='orange')
plt.axvline(x = np.median(x), color='green')
_ = plt.hist(x, color = 'lightgray')
# Note: Mode typically only applies to PMFs; this is a PDF
# + [markdown] id="EdRT_Lme5Chb"
# Skewed distributions, in contrast, drag the mean away from the center and toward the tail:
# + id="85C_EFZZ5Chb"
x = st.skewnorm.rvs(10, size=1000)
# + id="pT0zxSPd5Chb" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="70df9666-57fa-496a-b1ff-1b7031c6455f"
fig, ax = plt.subplots()
plt.axvline(x = np.mean(x), color='orange')
_ = plt.hist(x, color = 'lightgray')
# + [markdown] id="-uDxp8YT5Chc"
# The mode is least impacted by skew, but is only applicable to discrete distributions. For continuous distributions with skew (e.g., salary data), median is typically the choice measure of central tendency:
# + id="zzrBO1fg5Chc" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="a37b0035-69cf-4501-8748-b4f0c77a3e1d"
fig, ax = plt.subplots()
plt.axvline(x = np.mean(x), color='orange')
plt.axvline(x = np.median(x), color='green')
_ = plt.hist(x, color = 'lightgray')
# + [markdown] id="RRr8oPbC5Chc"
# ### Quantiles
# + [markdown] id="Sorh-6kS5Chc"
# The median, which divides a distribution in two at its midpoint, is the most well-known example of a quantile:
# + id="CZfX5qP95Chc" colab={"base_uri": "https://localhost:8080/"} outputId="c04b6c11-c8db-483c-92f0-ac654b0a5e0a"
np.median(x)
# + id="YqPFRjcO5Chc" colab={"base_uri": "https://localhost:8080/"} outputId="51ec4f9e-7f32-496e-e3ff-3a72b96abccf"
np.quantile(x, 0.5)
# + [markdown] id="HpK3wcmy5Chd"
# Generally speaking, quantiles divide up distributions and the most common are:
#
# * Percentiles,
# * Quartiles, and
# * Deciles.
# + [markdown] id="kIB-rEJQ5Chd"
# **Percentiles** divide the distribution at any point out of one hundred ("pour cent" is French for "out of a hundred"). For example, if we'd like to identify the threshold for the top 5% of a distribution, we'd cut it at the 95th percentile. Or, for the top 1%, we'd cut at the 99th percentile.
# + id="3T9AE9aS5Chd" colab={"base_uri": "https://localhost:8080/"} outputId="3fc34ef5-479f-4b81-bd93-60bba04eea1e"
p = np.percentile(x, [95, 99])
p
# + id="A_guMMCh5Chd" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="785ea9b7-6bc3-4c92-9f58-e7714f113289"
fig, ax = plt.subplots()
plt.axvline(x = p[0], color='orange')
plt.axvline(x = p[1], color='red')
_ = plt.hist(x, color = 'lightgray')
# + [markdown] id="atT5Wbtn5Chd"
# Note that the Numpy `quantile()` method is identical to the `percentile()` method except we pass proportions (ranging from zero to one) into the former and percentages (ranging from zero to 100) into the latter:
# + id="xEQo8y7u5Chd" colab={"base_uri": "https://localhost:8080/"} outputId="2c98e4b2-f92e-4eb8-a4bc-6d00a937b798"
np.quantile(x, [.95, .99])
# + [markdown] id="vDhFcNWj5Che"
# **Quartiles**, as their name suggests, are quantiles that divide a distribution into quarters by splitting the distribution at the 25th percentile, the median (a.k.a. the 50th percentile), and the 75th percentile:
# + id="miEwAsQV5Che" colab={"base_uri": "https://localhost:8080/"} outputId="e15cdd18-b73c-44c7-cc19-7407f349e01b"
q = np.percentile(x, [25, 50, 75])
q
# + id="kQvjcq3O5Che" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="6e68df01-f07c-4c34-94e4-6f8dd13b8d56"
fig, ax = plt.subplots()
plt.axvline(x = q[0], color='cornflowerblue')
plt.axvline(x = q[1], color='green')
plt.axvline(x = q[2], color='cornflowerblue')
_ = plt.hist(x, color = 'lightgray')
# + [markdown] id="JWVX933i5Che"
# Finally, **deciles** (from Latin *decimus*, meaning "tenth") divide a distribution into ten evenly-sized segments:
# + id="SD59HmMa5Che" colab={"base_uri": "https://localhost:8080/"} outputId="5016eada-cc0f-4122-9b35-cd5a956a544c"
[i for i in range(10, 100, 10)]
# + id="pVuo_iTo5Chf" colab={"base_uri": "https://localhost:8080/"} outputId="a93fbff4-7765-4bcd-a060-4850965cfd38"
d = np.percentile(x, range(10, 100, 10))
d
# + id="wex6mxXx5Chf" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="1f3ddb15-52f7-42d5-e694-0d91e8263a4d"
fig, ax = plt.subplots()
[plt.axvline(x = d_i, color='cornflowerblue') for d_i in d]
_ = plt.hist(x, color = 'lightgray')
# + [markdown] id="ul0eYeis5Chf"
# As examples, we can refer to the bottom 10% as the bottom decile or the first decile, while the top 10% is the top decile or tenth decile.
# + [markdown] id="hW2cvXR45Chf"
# ### The Box-and-Whisker Plot
# + id="hstuGQgw5Chf"
sns.set(style='whitegrid')
# + id="KTJsZfB25Chg" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="a6ae3824-2de8-4cec-aeaa-65d96971fdd2"
_ = sns.boxplot(x=x)
# + [markdown] id="LYTGM6aL5Chg"
# Box edges and median are determined by quartiles:
# + id="cwobiHMQ5Chg" colab={"base_uri": "https://localhost:8080/"} outputId="c4dbfa40-c81d-4009-acca-056a0c5746bb"
q
# + [markdown] id="87xWa_O25Chg"
# Box edges define the **inter-quartile range** (IQR):
# + id="geDTr1LS5Chg" colab={"base_uri": "https://localhost:8080/"} outputId="87fcd2c7-de15-46c4-a165-4cfd8e8ad056"
r = 1.194 - 0.325
r
# + [markdown] id="AoLruNpq5Chh"
# Whisker lengths are determined by furthest data points within $1.5 \times \text{IQR}$ of the box edges.
# + [markdown] id="or6R8lpL5Chh"
# In this case, the lower whisker could stretch as far down (to the left in the plot) as:
# + id="PgFCmy2g5Chh" colab={"base_uri": "https://localhost:8080/"} outputId="2b624400-47e2-4cfd-a7f1-d234a83ace60"
0.325 - 1.5*r
# + [markdown] id="1jGgu10C5Chh"
# The lowest value is inside of that so the whisker is plotted where that lowest value is:
# + id="DoTmuS6h5Chh" colab={"base_uri": "https://localhost:8080/"} outputId="8d8d5497-6860-4ae9-fb8c-ac42b9c9c142"
np.min(x)
# + [markdown] id="VAqf6jZ25Chi"
# The upper whisker could stretch as far up (to the right in the plot) as:
# + id="owW6yEe75Chi" colab={"base_uri": "https://localhost:8080/"} outputId="37467a24-1eb5-4247-df4f-fbc3a15c9fd1"
1.194 + 1.5*r
# + [markdown] id="wdC_HHjO5Chi"
# There are several (eleven) values beyond this threshold in the distribution. These values are considered **outliers** and are plotted as individual points:
# + id="MB1QCond5Chi"
x.sort()
# + id="mrWngeiJ5Chi" colab={"base_uri": "https://localhost:8080/"} outputId="79f82b6c-5eed-4ff3-84df-4c0f704625bf"
x[-15:]
# + [markdown] id="myU38cQq5Chi"
# The twelth-highest value, 2.435, is the largest within the upper whisker's maximum reach and so the upper whisker is plotted there.
# + [markdown] id="plWdKLkK5Chj"
# Useful for examining distributions on their own, box-and-whisker plots are especially helpful for comparing distributions:
# + id="hU6gylwx5Chj" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="a1926c5b-a4a7-452d-e4e1-03a213d2add9"
iris = sns.load_dataset('iris')
iris
# + id="vKqPo2sn5Chj" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="a5013887-d7a1-490d-acc9-d4707ffcdea3"
_ = sns.boxplot(data=iris)
# + id="lYtj4rTU5Chj" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="d9f21c47-8ce3-422c-ae60-0c3f1465317e"
tips = sns.load_dataset('tips')
tips
# + id="kwt0xVEP5Chj" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="f9d281b3-e200-4b62-d1d0-e33c8899fe50"
_ = sns.boxplot(x='day', y='total_bill', hue='smoker', data=tips)
# + [markdown] id="quNYihVQ5Chk"
# ### Measures of Dispersion
# + [markdown] id="MKaq4xJ25Chk"
# IQR is a relatively rare measure of the dispersion of values around the center of a distribution. The most widely-used are:
#
# * Variance,
# * Standard deviation, and
# * Standard error.
# + [markdown] id="Ti6GdZeZ5Chk"
# **Variance** (denoted with $\sigma^2$) can be written using expected-value notation, but it's easier to understand without it:
# $$ \sigma^2 = \frac{\sum_{i=1}^n (x_i-\bar{x})^2}{n} $$
# + [markdown] id="B1lo6mxN5Chk"
# (The astute follower of the *Machine Learning Foundations* series may notice that this formula is the same as mean-squared-error cost, except $x_i$ is being compared to the mean $\mu$ instead of a predicted value $\hat{x}_i$.)
# + [markdown] id="wiOq636c5Chk"
# (Technically speaking, we should divide by $n$-1 with a sample of data, but with the large datasets typical of machine learning, it's a negligible difference. If $n$ were equal to a small number like 8 then it would matter.)
# + [markdown] id="pD27D-0g5Chk"
# (Also technically speaking, the variance of a sample is typically denoted with $s^2$ as opposed to the Greek $\sigma^2$, akin to how $\bar{x}$ denotes the mean of a sample while the Greek $\mu$ is reserved for population mean.)
# + [markdown] id="4G-ogAFP5Chk"
# As an example let's calculate the variance of the PDF `x` from earlier:
# + id="AGO5TXzt5Chk" colab={"base_uri": "https://localhost:8080/"} outputId="367f5a61-db75-40fe-85bd-87299ccc6bfb"
xbar = np.mean(x)
xbar
# + id="u-fEygHR5Chl"
squared_differences = [(x_i - xbar)**2 for x_i in x]
# + id="qdvZ5_fg5Chl" colab={"base_uri": "https://localhost:8080/"} outputId="d806aaa1-07fb-4e5a-c34e-d64e08ce183a"
squared_differences[0:10]
# + id="jduq8CVv5Chl" colab={"base_uri": "https://localhost:8080/"} outputId="4501df61-bbdb-4d9a-f132-c6fa84fb9220"
sigma_squared = sum(squared_differences)/len(x)
sigma_squared
# + [markdown] id="mnZoMJAj5Chl"
# Of course there's a built-in NumPy method:
# + id="pIci-1It5Chl" colab={"base_uri": "https://localhost:8080/"} outputId="e86df4d3-47e2-460e-e64d-ae662eda8aff"
np.var(x)
# + [markdown] id="TChK1Qii5Chl"
# #### Standard Deviation
# + [markdown] id="Ei-js4cF5Chm"
# A straightforward derivative of variance is **standard deviation** (denoted with $\sigma$), which is convenient because its units are on the same scale as the values in the distribution:
# $$ \sigma = \sqrt{\sigma^2} $$
# + id="ginsMxgi5Chm" colab={"base_uri": "https://localhost:8080/"} outputId="cbb90697-f761-4430-91dd-bf6d305fc57b"
sigma = sigma_squared**(1/2)
sigma
# + id="mT0pi9Qx5Chm" colab={"base_uri": "https://localhost:8080/"} outputId="4560e541-2537-4c35-f9ff-17e384c2d2ba"
np.std(x)
# + id="k7a_hKT75Chm" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="1280c67d-9818-45f8-a4a6-07ca6d6d4d55"
fig, ax = plt.subplots()
plt.axvline(x = xbar, color='orange')
plt.axvline(x = xbar+sigma, color='olivedrab')
plt.axvline(x = xbar-sigma, color='olivedrab')
_ = plt.hist(x, color = 'lightgray')
# + [markdown] id="TghPdBIE5Chm"
# #### Standard Error
# + [markdown] id="vuIykMWN5Chm"
# A further derivation of standard deviation is **standard error**, which is denoted with $\sigma_\bar{x}$:
# $$ \sigma_\bar{x} = \frac{\sigma}{\sqrt{n}} $$
# + id="6C2xB1UU5Chn" colab={"base_uri": "https://localhost:8080/"} outputId="7ba71331-7221-4685-d1b5-770dd1032064"
sigma/(len(x))**(1/2)
# + id="Itaub6WD5Chn" colab={"base_uri": "https://localhost:8080/"} outputId="8aa80640-7752-48bb-e7bc-5f7a3f908667"
st.sem(x) # defaults to 1 degree of freedom (n-1), which can be ignored with the larger data sets of ML
# + id="YAjsX8WB5Chn" colab={"base_uri": "https://localhost:8080/"} outputId="f6611684-66ae-4140-ccf7-0aac92d1eece"
st.sem(x, ddof=0) # 0 degrees of freedom (n)
# + [markdown] id="MuPQwbJn5Chn"
# Standard error enables us to compare whether the means of two distributions differ *significantly*, a focus of *Intro to Stats*.
# + [markdown] id="XwRxQf9E5Chn"
# ### Measures of Relatedness
# + [markdown] id="ttvO-Q_B5Chn"
# If we have two vectors of the same length, $x$ and $y$, where each element of $x$ is paired with the corresponding element of $y$, **covariance** provides a measure of how related the variables are to each other:
# $$ \text{cov}(x, y) = \frac{\sum_{i=1}^n (x_i - \bar{x})(y_i - \bar{y}) }{n} $$
# + id="oP01sWnf5Cho" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="5ee9566e-57d0-4981-9d58-fe8279742196"
iris
# + id="t9rSqTHC5Cho"
x = iris.sepal_length
y = iris.petal_length
# + id="hM-_V5jF5Cho" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="fd6e46d1-6683-4b61-86f7-1a4901c509c1"
_ = sns.scatterplot(x=x, y=y)
# + id="rfOYkAzt5Cho"
n = len(x)
# + id="-VIXsv3j5Cho"
xbar = sum(x)/n
ybar = sum(y)/n
# + id="15WoxxTH5Cho"
product = []
for i in range(n):
product.append((x[i]-xbar)*(y[i]-ybar))
# + id="gygz20ti5Cho" colab={"base_uri": "https://localhost:8080/"} outputId="3682a178-2bd6-421c-9042-8abd80480527"
cov = sum(product)/n
cov
# + [markdown] id="YEc4sv0v5Chp"
# The NumPy `cov()` method returns a **covariance matrix**, which is a $2 \times 2$ matrix because $x$ and $y$ together describe a two-dimensional space:
# + id="6n9HymBG5Chp" colab={"base_uri": "https://localhost:8080/"} outputId="5a106bb0-03b1-4f36-8826-c9e157b8f831"
np.cov(x, y, ddof=0) # again, defaults to ddof=1
# + [markdown] id="yBXI9B2-5Chp"
# The diagonal elements of the covariance matrix represent the variance of $x$ and $y$, respectively:
# + id="uKhq8mSA5Chp" colab={"base_uri": "https://localhost:8080/"} outputId="11186075-417b-46bd-eddd-e78a70c5bcc2"
np.var(x)
# + id="NrtQqVjr5Chp" colab={"base_uri": "https://localhost:8080/"} outputId="97318707-9cc4-47ad-99fd-fccbdec37587"
np.var(y)
# + [markdown] id="1y2wfLeD5Chq"
# If $x$ and $y$ are inversely related, their covariance is negative. The less related they are, the closer their covariance is to zero:
# + id="epIqQ-p25Chq" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="768406cd-f6f4-4f2d-ed53-c8bf7c909cbd"
_ = sns.scatterplot(x=iris.sepal_length, y=iris.sepal_width)
# + id="QoIKgBHf5Chq" colab={"base_uri": "https://localhost:8080/"} outputId="d194e10c-27c3-4c1b-c725-d924711c4fd6"
np.cov(iris.sepal_length, iris.sepal_width, ddof=0)
# + [markdown] id="t7GPaB_m5Chq"
# #### Correlation
# + [markdown] id="uFDuVBQE5Chq"
# A drawback of covariance is that it confounds the relative scale of two variables with a measure of the variables' relatedness. **Correlation** builds on covariance and overcomes this drawback via rescaling, thereby measuring relatedness exclusively. Correlation is much more common because of this difference.
#
# The correlation coefficient (developed by <NAME> in the 20th c. though known in the 19th c.) is often denoted with $r$ or $\rho$ and is defined by:
# $$ \rho_{x,y} = \frac{\text{cov}(x,y)}{\sigma_x \sigma_y} $$
# + id="j2FfBUs_5Chq"
cov = -0.04215111
sigma_sq_x = 0.68112222
sigma_sq_y = 0.18871289
# + id="r6fRzR015Chr"
sigma_x = sigma_sq_x**(1/2)
sigma_y = sigma_sq_y**(1/2)
# + id="O3fHMlFY5Chr" colab={"base_uri": "https://localhost:8080/"} outputId="5ec80b5b-4ae4-4d67-9203-1c9f7646e78f"
cov / (sigma_x * sigma_y)
# + id="LYHyhLnC5Chr" colab={"base_uri": "https://localhost:8080/"} outputId="46657183-79de-4b0e-f631-9bbb28575469"
st.pearsonr(iris.sepal_length, iris.sepal_width)
# + [markdown] id="Jp8yMtiM5Chr"
# The second value output of `pearsonr` is a measure of statistical significance, which we'll detail in *Intro to Stats*.
# + [markdown] id="J3HRJ9S7klIp"
# $\rho$ has a range of -1 to 1, with values closer to zero indicating less correlation:
# + id="6vF_9wgj5Chr" colab={"base_uri": "https://localhost:8080/"} outputId="a195fafa-b7de-40ad-ac42-9c2015ca438a"
st.pearsonr(iris.sepal_length, iris.sepal_width)[0]
# + [markdown] id="dUIa7O-6lXy8"
# The closer $\rho$ is to 1 or -1, the stronger the positive or negative correlation, respectively:
# + id="d9KqkNOr5Chr" colab={"base_uri": "https://localhost:8080/"} outputId="a88d431f-600d-492a-a0a9-31e62a733b3b"
st.pearsonr(iris.sepal_length, iris.petal_length)[0]
# + [markdown] id="negAO_iA5Chs"
# N.B.: Covariance and correlation only account for linear relationships. Two variables could be non-linearly related to each other and these metrics could come out as zero.
# + [markdown] id="elRFzq495Chs"
# **Return to slides here.**
# + [markdown] id="fwp0wDiy5Chs"
# ## Segment 2: Distributions in Machine Learning
# + [markdown] id="UtNPNOzj5Cht"
# ### Uniform
# + [markdown] id="mkg5ZlcY5Cht"
# The uniform distribution is about as straightforward as they come. It has constant probabilities across the entire range of values in its domain:
# + id="m3kOc_-t5Cht"
u = np.random.uniform(size=10000)
# + id="Nfqw-vNJ5Cht"
sns.set_style('ticks')
# + id="yFF75RyzQCTu" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="7832b245-e4c4-4c98-d3d5-e0ab2d3d9de9"
_ = sns.displot(u)
# + [markdown] id="iAccWery5Cht"
# Real-world examples include:
#
# * Dice rolling (PMF)
# * Card drawing (PMF)
# * Model hyperparameters
# * Emission of radioactive particles
# * Economic demand
# * Analog-to-digital signal quantization errors
# + [markdown] id="q0cQIl945Cht"
# ### Gaussian
# + [markdown] id="-rW0khrV5Cht"
# <NAME> (early 19th c. German mathematician and scientist) is the namesake of over a hundred concepts across mathematics, physics, and astronomy. One of those concepts is the Gaussian distribution, also known as the "bell curve" (though several distributions are bell-shaped) or **normal distribution**:
# + id="zhcadaJu5Chu"
x = np.random.normal(size=10000)
# + id="1cQ6-LIw5Chu" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="d284c154-ba0e-4c36-9b87-d5353e7ce8a9"
_ = sns.displot(x)
# + id="sFpUGuMX5Chu" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="ca95419f-1f4c-47c2-9a3b-27657cc2cd43"
_ = sns.displot(x, kde=True)
# + [markdown] id="fCrRRX1F5Chu"
# When the normal distribution has a mean ($\mu$) of zero and standard deviation ($\sigma$) of one, as it does by default with the NumPy `normal()` method...
# + id="DMa4Z2h35Chu" colab={"base_uri": "https://localhost:8080/"} outputId="d464e02b-27a1-41ae-c998-66dd874a00d2"
np.mean(x)
# + id="k8QSIvEJ5Chu" colab={"base_uri": "https://localhost:8080/"} outputId="77b47571-090a-48b1-94d4-afb3e292941d"
np.std(x)
# + [markdown] id="iD-WgAZd5Chv"
# ...it is a **standard normal distribution** (a.k.a., standard Gaussian distribution). Normal distributions can be denoted with the *math calligraphy* font as $\mathcal{N}(\mu, \sigma^2)$, thus the standard normal distribution can be denoted as $\mathcal{N}(0, 1)$.
#
# (N.B.: $\sigma^2 = \sigma$ in this case because $1^2 = 1$.)
# + [markdown] id="3FelqdY05Chv"
# Normal distributions are by far the most common distribution in statistics and machine learning. They are typically the default option, particularly if you have limited information about the random process you're modeling, because:
#
# 1. Normal distributions assume the greatest possible uncertainty about the random variable they represent (relative to any other distribution of equivalent variance). Details of this are beyond the scope of this tutorial.
# 2. Simple and very complex random processes alike are, under all common conditions, normally distributed when we sample values from the process. Since we sample data for statistical and machine learning models alike, this so-called **central limit theorem** (covered next) is a critically important concept.
# + [markdown] id="WE8R8Y385Chv"
# Real-world examples include:
#
# * Probability of heads across *n* tosses (PMF; $n = 5$ earlier, $n = 100$ below)
# * Sum of di(c)e rolls (PMF)
# * Height of adult women
# * Height of adult men
# * Education test scores, e.g., SAT
# + id="iLylH-4m5Chv"
n_experiments = 10000
heads_count = [np.random.binomial(100, 0.5) for i in range (n_experiments)]
heads, event_count = np.unique(heads_count, return_counts=True)
event_proba = event_count/n_experiments
# + id="1OAA4vlZ5Chv" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="630c7d28-1e0f-4028-f56e-2a0d3267c11b"
plt.bar(heads, event_proba)
_ = plt.xlabel('Heads flips (out of 100 tosses)')
# + [markdown] id="HOY28bQX5Chv"
# ### The Central Limit Theorem
# + [markdown] id="9UIXSsAh5Chv"
# To develop a functional understanding of the CLT, let's sample some values from our normal distribution:
# + id="GJkIY5N75Chw" colab={"base_uri": "https://localhost:8080/"} outputId="ab039c5e-9083-48f1-dfeb-c7b5e6971629"
x_sample = np.random.choice(x, size=10, replace=False)
x_sample
# + [markdown] id="SrS19VdK5Chw"
# The mean of a sample isn't always going to be close to zero with such a small sample:
# + id="2B3kjm-H5Chw" colab={"base_uri": "https://localhost:8080/"} outputId="51bae542-ee5e-4c06-bd28-17830bcd5ca0"
np.mean(x_sample)
# + [markdown] id="nOlR7gor5Chw"
# Let's define a function for generating **sampling distributions** of the mean of a given input distribution:
# + id="YiPPxQLF5Chw"
def sample_mean_calculator(input_dist, sample_size, n_samples):
sample_means = []
for i in range(n_samples):
sample = np.random.choice(input_dist, size=sample_size, replace=False)
sample_means.append(sample.mean())
return sample_means
# + id="0FAWIZvn5Chw" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="800adc78-a094-4d6f-d132-86b1de881e1b"
sns.displot(sample_mean_calculator(x, 10, 10), color='green')
_ = plt.xlim(-1.5, 1.5)
# + [markdown] id="5KWmR4HE5Chx"
# The more samples we take, the more likely that the sampling distribution of the means will be normally distributed:
# + id="_8ZtuXJu5Chx" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="3f623974-5a47-4a50-c5fd-1204e8fd4f80"
sns.displot(sample_mean_calculator(x, 10, 1000), color='green', kde=True)
_ = plt.xlim(-1.5, 1.5)
# + [markdown] id="jgeYnkEz5Chx"
# The larger the sample, the tighter the sample means will tend to be around the population mean:
# + id="d1plzceb5Chx" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="61cda73a-ec69-4978-bb14-9eb0f6d6baa4"
sns.displot(sample_mean_calculator(x, 100, 1000), color='green', kde=True)
_ = plt.xlim(-1.5, 1.5)
# + id="ITeo-nOc5Chx" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="d7fdb42e-d921-460d-f4d8-be8ec512b817"
sns.displot(sample_mean_calculator(x, 1000, 1000), color='green', kde=True)
_ = plt.xlim(-1.5, 1.5)
# + [markdown] id="Jti_SH175Chx"
# #### Sampling from a skewed distribution
# + id="T-cjJ4K65Chx"
s = st.skewnorm.rvs(10, size=10000)
# + id="bYcXrbZ45Chx" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="04af0856-6666-4570-a80c-870d3f161902"
_ = sns.displot(s, kde=True)
# + id="X5asvK2p5Chy" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="0651daa8-b5e0-418d-cd5a-ff96e7cb0c48"
_ = sns.displot(sample_mean_calculator(s, 10, 1000), color='green', kde=True)
# + id="o5-tyS_N5Chy" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="a1a1a4ac-a1db-42e8-989b-67c2f18f1dbb"
_ = sns.displot(sample_mean_calculator(s, 1000, 1000), color='green', kde=True)
# + [markdown] id="VD1FyOP-5Chy"
# #### Sampling from a multimodal distribution
# + id="qefz7pqq5Chy"
m = np.concatenate((np.random.normal(size=5000), np.random.normal(loc = 4.0, size=5000)))
# + id="dUVIgXcq5Chy" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="23c9d418-605f-4823-bf21-86c5030cc936"
_ = sns.displot(m, kde=True)
# + id="RooddaTD5Chy" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="78bd7195-678a-4485-a5d5-a78649793113"
_ = sns.displot(sample_mean_calculator(m, 1000, 1000), color='green', kde=True)
# + [markdown] id="5LjEuVP85Chz"
# #### Sampling from uniform
# + [markdown] id="IwjCW3vY5Chz"
# Even sampling from the highly non-normal uniform distribution, the sampling distribution comes out normal:
# + id="9yf6jQQW5Chz" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="21a4ec75-0720-4d65-eeeb-bd9fc6524be2"
_ = sns.displot(u)
# + id="AKEuTLxd5Chz" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="e4f4a983-1354-410a-bd22-dc5ed03384d8"
_ = sns.displot(sample_mean_calculator(u, 1000, 1000), color='green', kde=True)
# + [markdown] id="3rU0m--S5Chz"
# Therefore, with large enough sample sizes, we can assume the sampling distribution of the means will be normally distributed, allowing us to apply statistical and ML models that are configured for normally distributed noise, which is often the default assumption.
#
# As an example, the "*t*-test" (covered in *Intro to Stats*) allows us to infer whether two samples come from different populations (say, an experimental group that receives a treatment and a control group that receives a placebo). Thanks to the CLT, we can use this test even if we have no idea what the underlying distributions of the populations being tested are, which may be the case more frequently than not.
# + [markdown] id="UHfmjg3I5Chz"
# (Despite being associated with such a key concept as the CLT, the name "normal distribution" wasn't originally intended to imply that other distributions are "abnormal". It's a historical quirk related to Gauss describing orthogonal (technically a.k.a. "normal") equations associated with applied uses of the distribution.)
# + [markdown] id="I7HILoi55Chz"
# (Finally, you may wonder what the purpose of sampling means is! Well, when we gather data from the real world we are nearly always sampling a subset of all the available data produced by a given random process. And, once we've collected a sample of data, the aspect of it we're often most interested in above all else is its mean.)
# + [markdown] id="cY6d8KLa5Chz"
# ### Log-Normal Distribution
# + [markdown] id="JhgvQLTF5Ch0"
# The natural logarithm of the **log-normal** distribution is normally distributed:
# + id="nwf4rsyP5Ch0"
x = np.random.lognormal(size=10000) # defaults to standard normal mu=0, sigma=1
# + id="SwTmkQTu5Ch0" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="09bafaac-725d-4eeb-afec-1a8ad6f3330d"
_ = sns.displot(x, kde=True)
# + id="WPw_gPIY5Ch0" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="f313ba45-0385-4dcf-e689-c78449d6ffb3"
_ = sns.displot(np.log(x), color='brown', kde=True)
# + [markdown] id="DaSbudQP5Ch0"
# Real-world examples:
#
# * Income
# * Length of comments in online forums
# * Duration of chess games or Rubik's Cube solves
# * Size of publicly-available video files
# * Number of hospitalized cases in an epidemic where public interventions are involved
# + [markdown] id="pj-m50lH5Ch0"
# ### Exponential Distribution
# + [markdown] id="oBeXEmHF5Ch0"
# Relatively squished up against zero and then decreases exponentially toward higher values. Log-normal distribution, in contrast, could take negative values and both increases and decreases.
# + id="tQy3DRwB5Ch0"
x = np.random.exponential(scale=4, size=10000) # "scale" parameter defaults to one
# + id="sTFF5gku5Ch0" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="2e382b46-5b9a-472a-d932-b71ee512b062"
_ = sns.displot(x)
# + [markdown] id="NMC_Fxai5Ch1"
# Its logarithm has a skewed distribution:
# + id="dV1cUVv65Ch1" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="5e7df796-0faa-4bca-8f36-bad2ba49a226"
_ = sns.displot(np.log(x), color='brown', kde=True)
# + [markdown] id="5T79UJ9c5Ch1"
# Real-world examples:
#
# * Time between requests to access Wikipedia pages
# * Used frequently in deep learning
# + [markdown] id="xwnjFNt25Ch1"
# ### Laplace Distribution
# + [markdown] id="tpfYwTti5Ch1"
# Named after <NAME>, whom we mentioned (and pictured) earlier as a key figure in the development of Bayesian statistics.
# + id="o-sFY5c_5Ch1"
x = np.random.laplace(size=10000)
# + [markdown] id="IGOrnTD25Ch1"
# Alternatively referred to as the "double exponential distribution":
# + id="J9xS2v3K5Ch1" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="fc0ee1ee-1517-4faa-d3df-ec90e0ae0338"
_ = sns.displot(x, kde=True)
# + [markdown] id="_Yq0Lj3h5Ch2"
# In addition to the scale parameter of the exponential function, it has a location parameter that allows it to be centered on any value:
# + id="HkgA0XMm5Ch2" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="05b90767-3626-4f4e-8ae4-674f11abf043"
_ = sns.displot(np.random.laplace(loc=25, size=10000), kde=True)
# + [markdown] id="KfY4anzI5Ch2"
# Real-world examples:
#
# * Extreme weather events, e.g., maximum rainfall in a day
# * Many machine learning applications; wherever an acute peak of probability is desired
# + [markdown] id="9Dvi1l8o5Ch2"
# ### Binomial Distribution
# + [markdown] id="Yco-2t4s5Ch2"
# All distributions so far have been for continuous variables (PDFs). This one is discrete (PMF).
#
# We've already been using it for coin flips; it's used for binary (0 or 1) outcome.
#
# Its parameters are:
#
# * *n*: number of trials
# * *p*: probability of outcome of 1
# * *size*: number of experiments with *n* trials each
# + id="LmaDYQm75Ch2"
n = 5
n_experiments = 1000
# + [markdown] id="vfSCVQer5Ch2"
# #### Fair coin (p = 0.5)
# + id="YnyWevsS5Ch2"
heads_count = np.random.binomial(n, 0.5, n_experiments)
heads, event_count = np.unique(heads_count, return_counts=True)
event_proba = event_count/n_experiments
# + id="5i0myZqv5Ch2" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="cddef694-fb5c-4238-d31a-fbb7d305ae83"
plt.bar(heads, event_proba, color='mediumpurple')
plt.xlabel('Heads flips (out of 5 tosses)')
_ = plt.ylabel('Event probability')
# + [markdown] id="i5qIJ8-m5Ch3"
# #### Weighted coin (p = 0.8)
# + id="eT0s3yiI5Ch3"
heads_count = np.random.binomial(n, 0.8, n_experiments)
heads, event_count = np.unique(heads_count, return_counts=True)
event_proba = event_count/n_experiments
# + id="SB2qD95l5Ch3" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="440cb648-14d2-4257-e7da-2235e5d743f4"
plt.bar(heads, event_proba, color='mediumpurple')
plt.xlabel('Heads flips (out of 5 tosses)')
_ = plt.ylabel('Event probability')
# + [markdown] id="cjCnz1zD5Ch3"
# Technically, binomial distributions are created by sampling $n>1$ "Bernoulli trials". The **Bernoulli** distribution is equivalent to the binomial distribution where $n=1$:
# + id="9j9WlO9v5Ch3" colab={"base_uri": "https://localhost:8080/"} outputId="6e31e058-53cd-4be5-9295-80a0a7cac225"
np.random.binomial(1, 0.5)
# + [markdown] id="xuv_176n5Ch3"
# Real-world examples:
#
# * Making it to work on time
# * Candidate being invited to interview
# * Epidemiology: probability of death or catching disease
# + [markdown] id="dXZYgPdw5Ch4"
# ### Multinomial Distribution
# + [markdown] id="aVhAIJvE5Ch4"
# Generalization of the binomial distribution to discrete random variables with more than two possible outcomes, e.g., the roll of a die:
# + id="bvnRByqA5Ch4"
n = 1000
# + id="tYPHmsMT5Ch4" colab={"base_uri": "https://localhost:8080/"} outputId="91569fbb-8cb7-4fcf-c3dd-cc68821bbd1a"
rolls = np.random.multinomial(n, [1/6.]*6)
rolls
# + id="dB9d5Zo_5Ch4"
event_proba = rolls/n
# + id="04T9DyGf5Ch4" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="bdeafc1f-706e-44a9-91dd-f0d5ca9ecb5e"
plt.bar(range(1, 7), event_proba, color='mediumpurple')
plt.xlabel('Die roll')
_ = plt.ylabel('Event probability')
# + [markdown] id="aWl-KRXL5Ch4"
# As in the binomial case, multinomial distributions are created by sampling $n>1$ multinoulli distributions, where the multinoulli distribution is equivalent to the multinomial distribution when $n=1$:
# + id="MrwYus375Ch4" colab={"base_uri": "https://localhost:8080/"} outputId="ac085f89-8800-410c-c6aa-fdf21ca12b60"
np.random.multinomial(1, [1/6.]*6)
# + [markdown] id="Bs2JiMaN5Ch5"
# Since multinomial distributions can represent any categorical variable (e.g., burger, hot dog, pizza; 52 playing cards; NFL teams), not just integer values, we can't always calculate an expected value.
# + [markdown] id="dKoLXZ5I5Ch5"
# ### Poisson Distribution
# + [markdown] id="Z3EQEF9u5Ch5"
# **Poisson** is a third widely-used distribution for discrete random variables and it's the final distribution we'll cover.
# + [markdown] id="C_9PDyAf5Ch5"
# It's named after French mathematician <NAME> and is used for count data, e.g.:
#
# * Number of cars that drive by in a minute
# * Number of guests at restaurant in an evening
# * Number of new hires in a month
# + [markdown] id="CcX5Mtbl5Ch5"
# First argument to NumPy `poisson()` method is $\lambda$, which must be greater than zero and guides peak of probability distribution. E.g., sampling from Poisson with $\lambda=5$ will tend to draw samples near 5.
# + id="zi31GP2P5Ch5"
lam=5
# + [markdown] id="d45vG8QF5Ch5"
# Second argument is number of samples to draw:
# + id="Y3VB3Y1P5Ch5"
n=1000
# + id="WxtctajF5Ch5"
samples = np.random.poisson(lam, n)
# + id="2Ayn0Vfb5Ch6" colab={"base_uri": "https://localhost:8080/"} outputId="575dbc9f-d0ea-4c5c-86ae-56eba6af78e8"
samples[0:20]
# + id="F2ys381P5Ch6"
x, x_count = np.unique(samples, return_counts=True)
# + id="_JSi8Jtc5Ch6" colab={"base_uri": "https://localhost:8080/"} outputId="901f0c66-3c4b-4f2c-d579-25060e2c1aca"
x
# + id="APZbiuhN5Ch6" colab={"base_uri": "https://localhost:8080/"} outputId="155d6e3a-67a9-4596-be7e-2a9dad9e576c"
x_count
# + id="BqXl7-hB5Ch6"
Px = x_count/n
# + id="mwzYi1QW5Ch7" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="ff75fe4f-1cc4-467b-e07c-c5f6f7d35d20"
plt.bar(x, Px, color='mediumpurple')
plt.title('PMF of Poisson with lambda = {}'.format(lam))
plt.xlabel('x')
_ = plt.ylabel('P(x)')
# + [markdown] id="Ekuyn4Zx5Ch7"
# ### Mixture Distributions
# + [markdown] id="gHHStGFb5Ch7"
# Adapted from [Stack Overflow post](https://stackoverflow.com/questions/47759577/creating-a-mixture-of-probability-distributions-for-sampling):
# + [markdown] id="gD4x2RJu5Ch7"
# Multinomial distribution (driven by `np.random.choice()`) with probabilities set by `coefficients` array determines which of the three distributions to sample from.
# + id="v4-2kJVr5Ch7" colab={"base_uri": "https://localhost:8080/", "height": 0} outputId="96d4bb9f-d7f6-4538-8cf3-f0a3b4014de5"
distributions = [
{"type": np.random.normal, "kwargs": {"loc": -3, "scale": 2}},
{"type": np.random.uniform, "kwargs": {"low": 4, "high": 6}},
{"type": np.random.normal, "kwargs": {"loc": 2, "scale": 1}},
]
coefficients = np.array([0.5, 0.2, 0.3])
coefficients /= coefficients.sum() # in case these did not add up to 1
sample_size = 10000
num_distr = len(distributions)
data = np.zeros((sample_size, num_distr))
for idx, distr in enumerate(distributions):
data[:, idx] = distr["type"](size=(sample_size,), **distr["kwargs"])
random_idx = np.random.choice(np.arange(num_distr), size=(sample_size,), p=coefficients)
sample = data[np.arange(sample_size), random_idx]
_ = sns.displot(sample, bins=100, kde=True)
# + [markdown] id="zctZVQbB5Ch7"
# **Gaussian mixture model** (GMM) is common type of mixture distribution, wherein all of the component distributions are normal.
# + [markdown] id="Lvje2Own5Ch8"
# **Return to slides here.**
# + [markdown] id="tAvcSl185Ch8"
# ## Segment 3: Information Theory
# + [markdown] id="CIj2L7gD5Ch8"
# From the slides, the **self-information** formula is:
# $$ I(x)=-\text{log}P(x) $$
# + id="xcm3rorj5Ch8"
def self_info(my_p):
return -1*np.log(my_p)
# + id="d2PfIyUV5Ch8" colab={"base_uri": "https://localhost:8080/"} outputId="d974f9b1-ea4b-4944-e236-d2d40dd08437"
self_info(1)
# + id="W8xI5mfL5Ch8" colab={"base_uri": "https://localhost:8080/"} outputId="db33af67-02e0-44e4-dd4f-702148be8bf9"
self_info(0.1)
# + id="sF-OyYxq5Ch8" colab={"base_uri": "https://localhost:8080/"} outputId="88397ccb-6684-4f5d-e26f-5e53a2be4e0a"
self_info(0.01)
# + id="82cIxElx5Ch8" colab={"base_uri": "https://localhost:8080/"} outputId="700be077-6353-4309-ab71-bbfee6deb9e1"
self_info(0.5)
# + id="PycwWSm_5Ch9" colab={"base_uri": "https://localhost:8080/"} outputId="9714e9d8-26b3-451c-da65-625515840564"
self_info(0.5) + self_info(0.5)
# + [markdown] id="xRSf8Q7M5Ch9"
# Depending on what logarithm base we use, the units of self-information vary. Most frequently, the units are either:
#
# * **nats**:
# * Natural logarithm, as above with `np.log()`
# * Typical in ML
# * **bits**:
# * Base-2 logarithm
# * A.k.a. **shannons**
# * Typical in computer science
#
# So, the self-information of $P(x) = 0.1$ is ~2.3 nats.
# + [markdown] id="892saRjz5Ch9"
# ### Shannon and Differential Entropy
# + [markdown] id="gy2ByHs35Ch9"
# To quantify uncertainty about a probability distribution (as opposed to a single event), we can use **Shannon entropy**, which is denoted $H(x)$ or $H(P)$:
# $$ H(x) = \mathbb{E}_{\text{x}\sim P}[I(x)] \\
# = -\mathbb{E}_{\text{x}\sim P}[\text{log}P(x)] $$
#
# Conveys the expected informational content associated with an event drawn from the distribution:
# * **Low entropy**:
# * Distribution is ~deterministic and outcomes are ~certain
# * E.g., weighted coin always lands heads up ($P(1)=1$)
# * **High entropy**:
# * Outcomes are uncertain
# * Uniform distribution can be highest entropy
# * E.g., coin is as likely to be heads as tails ($P(0)=P(1)=0.5$)
# + [markdown] id="1h_CUdNh5Ch9"
# As an example, Shannon entropy for a binary random variable (e.g., coin flip) is:
# $$ (p-1)\text{log}(1-p)-p \text{log}p $$
# + id="MQAUp05w5Ch9"
def binary_entropy(my_p):
return (my_p-1)*np.log(1-my_p) - my_p*np.log(my_p)
# + [markdown] id="og4zT2kS5Ch-"
# Will throw `nan` with $p=0$ (always tails) or $p=1$ (always heads), but we can get close:
# + id="TohThggW5Ch-" colab={"base_uri": "https://localhost:8080/"} outputId="2ad506a7-ccf6-4de2-88c8-03e5288b47d6"
binary_entropy(0.00001)
# + id="Pg8tz7f95Ch-" colab={"base_uri": "https://localhost:8080/"} outputId="83186813-2f40-4354-a840-55e583d4f19e"
binary_entropy(0.99999)
# + id="9cJU5CP-5Ch-" colab={"base_uri": "https://localhost:8080/"} outputId="ec5d1860-79f7-4a59-deac-787cfbced659"
binary_entropy(0.9)
# + id="zASdy_6X5Ch-" colab={"base_uri": "https://localhost:8080/"} outputId="51969bf0-71a9-4e80-d1c6-57202664d986"
binary_entropy(0.5)
# + id="hEFpmgMF5Ch-"
p = np.linspace(0.001, 0.999, 1000) # start, finish, n points
# + id="63lu2Yda5Ch-"
H = binary_entropy(p)
# + id="g7oaT7kb5Ch_" colab={"base_uri": "https://localhost:8080/", "height": 301} outputId="92b73bf3-2416-4562-89ce-d71d6d539eaf"
fig, ax = plt.subplots()
plt.title('Shannon entropy of Bernoulli trial')
plt.xlabel('p')
plt.ylabel('H (nats)')
_ = ax.plot(p,H)
# + [markdown] id="Gg4IxIJ25Ch_"
# **Differential entropy**: simply the term for Shannon entropy if distribution is PDF
# + [markdown] id="uVpOXC3K5Ch_"
# ### Kullback-Leibler Divergence
# + [markdown] id="b-qE4xHD5Ch_"
# **KL divergence** enables us to quantify the relative Shannon (or differential) entropy of two probability distributions that are over the same random variable x.
#
# For example, if we have one probability distribution described by $P(x)$ and another by $Q(x)$, their KL divergence (denoted $D_\text{KL}$) is:
# $$ D_\text{KL}(P||Q) = \mathbb{E}_{\text{x} \sim P}[\text{log}P(x) - \text{log}Q(x)] $$
# + [markdown] id="0IGVYCUx5Ch_"
# ### Cross-Entropy
# + [markdown] id="TaFwwQIC5Ch_"
# Cross-entropy is a concept derived from KL divergence. Its detail is beyond the scope of this series except to mention that it provides us with the **cross-entropy cost** function.
#
# This cost function is ubiquitous in neural networks as it's the cost function of choice for predicting discrete, categorical outcomes. E.g., for a binary classifier, the cross-entropy cost ($C$) is:
#
# $$ C = -(y \cdot \text{log}(\hat{y}) + (1-y) \cdot \text{log}(1-\hat{y})) $$
# + id="vSUxkRMX5Ch_"
def cross_entropy(y, a):
return -1*(y*np.log(a) + (1-y)*np.log(1-a))
# + id="0RNuubWo5Ch_" colab={"base_uri": "https://localhost:8080/"} outputId="e9111867-88ca-437a-9d5f-c4b6c8713023"
cross_entropy(1, 0.9997)
# + id="rRiZkkxX5CiA" colab={"base_uri": "https://localhost:8080/"} outputId="32bc007f-297e-4bb1-ca8d-b732f2239e27"
cross_entropy(1, 0.9)
# + id="kvQ-JUk45CiA" colab={"base_uri": "https://localhost:8080/"} outputId="f8549a6e-a604-4ad7-e75c-d6d1401230f6"
cross_entropy(1, 0.6)
# + id="__hYqZuT5CiA" colab={"base_uri": "https://localhost:8080/"} outputId="5c3848d5-a16d-48e3-91c0-720584a58c4d"
cross_entropy(1, 0.3)
| notebooks/5-probability.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from __future__ import absolute_import, division, print_function
# # Github
# https://github.com/jbwhit/OSCON-2015/commit/6750b962606db27f69162b802b5de4f84ac916d5
# ## A few Python Basics
# Create a [list]
days = ['Monday', # multiple lines
'Tuesday', # acceptable
'Wednesday',
'Thursday',
'Friday',
'Saturday',
'Sunday', # trailing comma is fine!
]
days
# Simple for-loop
for day in days:
print(day)
# Double for-loop
for day in days:
for letter in day:
print(letter)
print(days)
print(*days)
# Double for-loop
for day in days:
for letter in day:
print(letter)
print()
for day in days:
for letter in day:
print(letter.lower())
# ## List Comprehensions
length_of_days = [len(day) for day in days]
length_of_days
letters = [letter for day in days
for letter in day]
print(letters)
letters = [letter for day in days for letter in day]
print(letters)
[num for num in xrange(10) if num % 2]
[num for num in xrange(10) if num % 2 else "doesn't work"]
[num if num % 2 else "works" for num in xrange(10)]
[num for num in xrange(10)]
sorted_letters = sorted([x.lower() for x in letters])
print(sorted_letters)
unique_sorted_letters = sorted(set(sorted_letters))
print("There are", len(unique_sorted_letters), "unique letters in the days of the week.")
print("They are:", ''.join(unique_sorted_letters))
print("They are:", '; '.join(unique_sorted_letters))
def first_three(input_string):
"""Takes an input string and returns the first 3 characters."""
return input_string[:3]
import numpy as np
# tab
np.linspace()
[first_three(day) for day in days]
def last_N(input_string, number=2):
"""Takes an input string and returns the last N characters."""
return input_string[-number:]
[last_N(day, 4) for day in days if len(day) > 6]
# +
from math import pi
print([str(round(pi, i)) for i in xrange(2, 9)])
# -
list_of_lists = [[i, round(pi, i)] for i in xrange(2, 9)]
print(list_of_lists)
for sublist in list_of_lists:
print(sublist)
# +
# Let this be a warning to you!
# If you see python code like the following in your work:
for x in range(len(list_of_lists)):
print("Decimals:", list_of_lists[x][0], "expression:", list_of_lists[x][1])
# +
print(list_of_lists)
# Change it to look more like this:
for decimal, rounded_pi in list_of_lists:
print("Decimals:", decimal, "expression:", rounded_pi)
# +
# enumerate if you really need the index
for index, day in enumerate(days):
print(index, day)
# -
# ## Dictionaries
#
# Python dictionaries are awesome. They are [hash tables](https://en.wikipedia.org/wiki/Hash_table) and have a lot of neat CS properties. Learn and use them well.
from IPython.display import IFrame, HTML
HTML('<iframe src=https://en.wikipedia.org/wiki/Hash_table width=100% height=550></iframe>')
fellows = ["Jonathan", "Alice", "Bob"]
universities = ["UCSD", "UCSD", "Vanderbilt"]
for x, y in zip(fellows, universities):
print(x, y)
# Don't do this
{x: y for x, y in zip(fellows, universities)}
# Doesn't work like you might expect
{zip(fellows, universities)}
dict(zip(fellows, universities))
fellows
fellow_dict = {fellow.lower(): university
for fellow, university in zip(fellows, universities)}
fellow_dict
fellow_dict['bob']
rounded_pi = {i:round(pi, i) for i in xrange(2, 9)}
rounded_pi[5]
sum([i ** 2 for i in range(10)])
sum(i ** 2 for i in range(10))
huh = (i ** 2 for i in range(10))
huh.next()
# ## Participate in StackOverflow
#
# An example: http://stackoverflow.com/questions/6605006/convert-pdf-to-image-with-high-resolution
| notebooks/07-Some_basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: mosquito-networking
# language: python
# name: mosquito-networking
# ---
# # Trying out a full pytorch experiment, with tensorboard, // processing, etc
# +
# OPTIONAL: Load the "autoreload" extension so that code can change
# #%load_ext autoreload
# OPTIONAL: always reload modules so that as you change code in src, it gets loaded
# #%autoreload 2
import numpy as np
import pandas as pd
from src.data import make_dataset
from src.data import read_dataset
from src.data import util
from torchsummary import summary
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
# -
df = make_dataset.main(reduce_mem_usage=False)
sns.countplot(df["label"])
# ## Convert data to pytorch types
data_vector = util.get_train_test(df)
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
x_train, x_test, y_train, y_test = util.transform_torch(data_vector, device=device)
print(x_train.shape, x_test.shape)
# -
# ## Basic 1D convolutional network
# [1D conv in Pytorch](https://pytorch.org/docs/stable/nn.html#torch.nn.Conv1d)
#
# In the simplest case, the output value of the layer with input size
#
# $$ (N, C_{\text{in}}, L) $$ and output $$ (N, C_{\text{out}}, L_{\text{out}}) $$ can be
#
# $$ (N, C_{\text{in}}, L) $$ and output $$ (N, C_{\text{out}}, L_{\text{out}}) $$ can be
# precisely described as:
#
# $$
# \text{out}(N_i, C_{\text{out}_j}) = \text{bias}(C_{\text{out}_j}) +
# \sum_{k = 0}^{C_{in} - 1} \text{weight}(C_{\text{out}_j}, k)
# \star \text{input}(N_i, k)
# $$
#
# where $$ \star $$ is the valid "cross-correlation" operator,
# N is a batch size, C denotes a number of channels,
# L is a length of signal sequence.
# # Create Model
# + code_folding=[]
class BasicMosquitoNet(nn.Module):
"""A basic 1D conv net.
We use 1D convolution, followed by max pool, 1D convolution, max pool, FC, FC.
"""
def __init__(self, conv1_out=100, kernel_1=6, stride_1=3,
conv2_out=10, kernel_2=4, stride_2=2):
"""
conv1: (22050 - 6)/3 + 1 = 7349
max_pool_1 = floor((Lin + −dilation×(kernel_size−1)−1)/stride_2) + 1
= floor(7349-2 /2) + 1 = 3673 + 1 = 3674
conv2 = (3674 - 4)/2 + 1 = 1836
max_pool_2 = floor(1836-2 /2) + 1 = 918
"""
super(BasicMosquitoNet, self).__init__()
self.conv1 = nn.Conv1d(in_channels=1, out_channels=conv1_out,
kernel_size=kernel_1, stride=stride_1)
self.conv2 = nn.Conv1d(in_channels=conv1_out, out_channels=conv2_out,
kernel_size=kernel_2, stride=stride_2)
self.fc1 = nn.Linear(918*conv2_out, 1)
#self.fc1 = nn.Linear(918*conv2_out, 120)
#self.fc2 = nn.Linear(120, 84)
#self.fc3 = nn.Linear(84, 2)
def forward(self, x):
"""
In the forward function we accept a Tensor of input data and we must return
a Tensor of output data.
"""
# Max pooling over a (2, 2) window
x = F.max_pool1d(F.relu(self.conv1(x)), 2)
# If the size is a square you can only specify a single number
x = F.max_pool1d(F.relu(self.conv2(x)), 2)
x = x.view(-1, self.num_flat_features(x))
x = self.fc1(x)
"""
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
"""
# We use BCEWithLogitsLoss instead of applying sigmoid here
# It is better computationally
return x
def num_flat_features(self, x):
size = x.size()[1:] # all dimensions except the batch dimension
num_features = 1
for s in size:
num_features *= s
return num_features
# -
# create your optimizer
net = BasicMosquitoNet()
criterion = nn.BCEWithLogitsLoss()
optimizer = optim.SGD(net.parameters(), lr=0.01, momentum=0.9)
summary(net, input_size=x_train.shape[1:])
# # Experiment params
# +
# Parameters
params = {'batch_size': 32,
'shuffle': True,
'num_workers': 0}
max_epochs = 1
# version = !python3 --version
version = version[0].split(".")[1]
if int(version) < 7 and params["num_workers"]:
print("WARNING\n"*10)
print("Parallel execution only works for python3.7 or above!")
print("Running in parallel with other versions is not guaranted to work")
print("See https://discuss.pytorch.org/t/valueerror-signal-number-32-out-of-range-when-loading-data-with-num-worker-0/39615/2")
# -
# # Start tensorboard
# Run in our terminal:
#
# `cd notebooks`
#
# `tensorboard --logdir runs`
# +
from torch.utils.tensorboard import SummaryWriter
# default `log_dir` is "runs" - we'll be more specific here
writer = SummaryWriter()
# -
# # Create data generator from dataset for batch processing
# +
# Generators
training_set = read_dataset.MosquitoDataset(x_train, y_train)
training_generator = torch.utils.data.DataLoader(training_set, **params)
testing_set = read_dataset.MosquitoDataset(x_test, y_test)
testing_generator = torch.utils.data.DataLoader(testing_set, **params)
# -
# Simple train function
def train(net, optimizer):
# Loop over epochs
last_test_loss = 0
for epoch in range(max_epochs):
# Training
for idx, (local_batch, local_labels) in enumerate(training_generator):
optimizer.zero_grad() # zero the gradient buffers
output = net(local_batch)
loss = criterion(output, local_labels)
loss.backward()
optimizer.step() # Does the update
writer.add_scalar("Train Loss Batch", loss.data.item(), idx)
# Validation
with torch.set_grad_enabled(False):
# Transfer to GPU
#local_batch, local_labels = local_batch.to(device), local_labels.to(device)
cumulative_test_loss = 0
for idx, (local_batch, local_labels) in enumerate(training_generator):
output = net(local_batch)
loss = criterion(output, local_labels)
writer.add_scalar("Test Loss Batch", loss.data.item(), idx)
cumulative_test_loss += loss.data.item()
cumulative_test_loss /= idx
last_test_loss = cumulative_test_loss
writer.add_scalar("Test Loss Epoch", loss.data.item(), idx)
return last_test_loss
writer.close()
# %%time
train(net, optimizer)
# The first Net had a random loss from the second batch onwards with regards to the training - Let's try something really small first
#
#
# - - -
#
# <br>
# <br>
# <br>
# <br>
# <br>
# <br>
# <br>
# <br>
# <br>
# <br>
# <br>
# <br>
# new writer
writer = SummaryWriter()
# create your optimizer
net = BasicMosquitoNet(conv1_out=32, conv2_out=4)
optimizer = optim.SGD(net.parameters(), lr=0.01, momentum=0.9)
summary(net, input_size=x_train.shape[1:])
# %%time
train(net, optimizer)
# Save the model
torch.save(net.state_dict(), "../models/0.6-BrunoGomesCoelho-test-experiment.pt")
| notebooks/0.6-BrunoGomesCoelho-test-experiment.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# **Notas para contenedor de docker:**
# Comando de docker para ejecución de la nota de forma local:
#
# nota: cambiar `dir_montar` por la ruta de directorio que se desea mapear a `/datos` dentro del contenedor de docker.
#
# ```
# dir_montar=<ruta completa de mi máquina a mi directorio>#aquí colocar la ruta al directorio a montar, por ejemplo:
# #dir_montar=/Users/erick/midirectorio.
# ```
#
# Ejecutar:
#
# ```
# $docker run --rm -v $dir_montar:/datos --name jupyterlab_prope_r_kernel_tidyverse -p 8888:8888 -d palmoreck/jupyterlab_prope_r_kernel_tidyverse:2.1.4
#
# ```
# Ir a `localhost:8888` y escribir el password para jupyterlab: `<PASSWORD>`
#
# Detener el contenedor de docker:
#
# ```
# docker stop jupyterlab_prope_r_kernel_tidyverse
# ```
#
# Documentación de la imagen de docker `palmoreck/jupyterlab_prope_r_kernel_tidyverse:2.1.4` en [liga](https://github.com/palmoreck/dockerfiles/tree/master/jupyterlab/prope_r_kernel_tidyverse).
# ---
# Para ejecución de la nota usar:
#
# [docker](https://www.docker.com/) (instalación de forma **local** con [Get docker](https://docs.docker.com/install/)) y ejecutar comandos que están al inicio de la nota de forma **local**.
#
# O bien dar click en alguno de los botones siguientes:
# [](https://mybinder.org/v2/gh/palmoreck/dockerfiles-for-binder/jupyterlab_prope_r_kernel_tidyerse?urlpath=lab/tree/Propedeutico/R/clases/2_probabilidad/7_variables_aleatorias.ipynb) esta opción crea una máquina individual en un servidor de Google, clona el repositorio y permite la ejecución de los notebooks de jupyter.
# [](https://repl.it/languages/Rlang) esta opción no clona el repositorio, no ejecuta los notebooks de jupyter pero permite ejecución de instrucciones de Python de forma colaborativa con [repl.it](https://repl.it/). Al dar click se crearán nuevos ***repl*** debajo de sus users de ***repl.it***.
#
# ### Lo siguiente está basado los libros:
#
# ### * <NAME>, Pensando Antes de Actuar: Fundamentos de Elección Racional, 2009.
#
# ### * <NAME>, Introduction to Probability and Statistics Using R, 2014.
# ### El libro de <NAME> tiene github: [jkerns/IPSUR](https://github.com/gjkerns/IPSUR)
# **Notas:**
#
# * Se utilizará el paquete [prob](https://cran.r-project.org/web/packages/prob/index.html) de R para los experimentos descritos en la nota y aunque con funciones nativas de R se pueden crear los experimentos, se le da preferencia a mostrar cómo en R se tienen paquetes para muchas aplicaciones.
#
# * En algunas líneas no es necesario colocar `print` y sólo se ha realizado para mostrar los resultados de las funciones en un formato similar al de R pues la nota se escribió con jupyterlab y R.
#
# * Cuidado al utilizar las funciones del paquete `prob` para construir espacios de probabilidad grandes como lanzar un dado 9 veces... (tal experimento tiene 10 millones de posibles resultados)
options(repr.plot.width=5, repr.plot.height=5) #esta línea sólo se ejecuta para jupyterlab con R
library(prob)
library(ggplot2)
# # Variables aleatorias
# En este tema nos interesa asociar un *número* con el experimento aleatorio, aún más, que el número esté asociado con cada resultado del experimento aleatorio de modo que al realizar el experimento y observar el resultado $r$ en $S$, el espacio de resultados, se calcule el número $X$. Esto es, para cada resultado (outcome) $r$ en $S$ se asocia el número $X(r)=x$.
# Una variable aleatoria $X$ es una **función** $X: S \rightarrow \mathbb{R}$ que asocia cada resultado (outcome) $r \in S$ de un experimento aleatorio, exactamente un número $X(r)=x$.
# **Obs:** obsérvese que $X$ es aleatoria pues se define en términos del experimento aleatorio, por lo que $X$ toma valores númericos, cada uno con cierta probabilidad.
# **Obs2:** típicamente se utiliza las letras mayúsculas para denotar a las variables aleatorias (funciones) y sus valores observados (los números que pueden tomar) con letras minúsculas.
# Así como $S$ es el espacio de resultados posibles del experimento, el **soporte de una variable aleatoria** es el conjunto de posibles valores que la variable aleatoria puede tomar. Si $X$ es variable aleatoria su soporte se denota: $S_X$.
# ## Ejemplos
# 1) **Experimento:** lanzar dos veces una moneda.
#
# Se tiene: $S=${AA, AS, SA, SS}. Defínase $X$ la variable aleatoria como: $X$ cuenta el número de águilas. Entonces $X(\{AA\}) = 2$, $X(\{AS\})=1$. Y podemos realizar una tabla con las posibilidades:
#
# |$r \in S$| AA|AS|SA|SS|
# |:-----------:|:-----------:|:-----------:|:-----------:|:-----------:
# |$X(r) = x$|2|1|1|0|
# Se tiene que $S_X=\{0,1,2\}$ (conjunto de valores de $X$).
# 2) **Experimento:** lanzar una moneda hasta observar un águila.
#
# Se tiene: $S=\{A, SA, SSA, SSSA, \dots\}$. Defínase $Y$ la variable aleatoria como: $Y$ cuenta el número de soles antes del primer águila. Entonces $S_Y=\{0, 1, 2, \dots\}$
# 3) Considérese el experimento de lanzar una moneda al aire y defínase la variable aleatoria $Z$ como: $Z$ el tiempo en segundos que le toma a la moneda caer al suelo. Entonces $S$ es el conjunto de números positivos y de hecho $S_Z$ también lo es.
# **Obs:** Obsérvese en los ejemplos anteriores que los soportes de las variables $X, Y, Z$ son distintos. De hecho: $S_X$ es un conjunto finito, $S_Y$ es un conjunto infinito pero contable y $S_Z$ es un conjunto infinito no contable. Variables aleatorias con soportes similares a los de $X$ y $Y$ se les nombra **discretas** y similares al soporte de $Z$ se les nombra **continuas**.
# ## Distribución de probabilidad de una variable aleatoria
# Supongamos que $X$ es una variable aleatoria tal que dado un conjunto de resultados inciertos $\{r_1,\dots, r_n\}$ ésta toma valores $X(r_1)=x_1,\dots, X(r_n)=x_n$. Nótese que éstos no necesariamente son distintos. La probabilidad de que la variable aleatoria tome el valor $x_i$ se denota por: $P(X = x_i)$.
# A la regla de correspondencia que asigna a cada valor $x_i$ su probabilidad $P(X = x_i),$ se le llama **distribución de probabilidad** de la variable aleatoria $X$. Si denotamos a esta regla por $f$ se tiene que: $f(x_i) = P(X = x_i)$.
# **Notas:**
#
# * A veces también se utiliza la notación: $f_X(x_i)$.
#
# * Obsérvese que en estricto sentido se debería escribir: $f(x_i) = P(X(r_i) = x_i)$ pero por simplicidad en la escritura y notación se utiliza $f(x_i) = P(X = x_i)$.
# ### Ejemplos
# 1) Supongamos que un volado ofrece una ganancia de $\$ 100$ si sale águila y una pérdida de $\$ 60$ si se sale sol. La variable aleatoria $X$ es la función dada por: $X(A) = 100$, $X(S) = -60$. La distribución de probabilidad $f$ es: $$f(100) = P(X = 100) = P(\{A\}) = \frac{1}{2}$$
# $$f(-60) = P(X = -60) = P(\{S\}) = \frac{1}{2}$$
# 2) Supóngase que el evento aleatorio es: $E=$ {el día de hoy llueve} y defínase $X$ variable aleatoria que representa un número que asocia mi satisfacción con la situación "llevar paraguas" y toma valores: $X(E) = 5, X(E^c) = -1$, esto es: llevo paraguas y si llueve me produce una satisfacción de $5$ puntos y si llevo paraguas y no llueve la inconveniencia me causa un disgusto de -1 punto.
#
#
# También defínase $Y$ como la variable aleatoria que representa un número que asocia mi satisfacción con la situación "no llevar paraguas" y toma valores: $Y(E) = -6$, $Y(E^c)=4$.
#
#
# Las distribuciones de probabilidad de $X$ y $Y$ están dadas por la siguiente tabla y supóngase que $f$ es la distribución de probabilidad $X$ y $g$ es la de $Y$:
#
# |Evento|Valor de $X$|Valor de $Y$|P(Variable aleatoria = valor)|
# |:----:|:----:|:----:|:----:
# |llueve|5|-6|f(5) = P(X=5) = g(-6) = P(Y=-6) = .45|
# |no llueve|-1|4|f(-1) = P(X = -1) = g(4) = P(Y = 4) = .55|
# 3) Pensemos en una urna con $10$ canicas rojas, $2$ amarillas y $3$ azules. Se nos ofrecen los siguientes premios: perder $\$100$ si sacamos una canica roja, ganar $\$150$ si sacamos un canica azul o amarilla. La variable aleatoria $Y$ es $Y(\{roja\}) = -100, Y (\{azul\}) = Y (\{amarilla\}) = 150$. La distribución de probabilidad de $f$ es:
#
# $$f(-100) = P(Y = -100) = P(\{roja\}) = \frac{10}{15}$$
# $$f(150) = P(Y = 150) = P(\{azul\} \cup \{amarilla\}) = \frac{3}{15} + \frac{2}{15}$$
# ## Representación de la distribución de probabilidad de una variable aleatoria
# La distribución de probabilidad puede representarse visualmente en un diagrama al que llamamos **histograma de probabilidad**. Un histograma es simplemente una gráfica de barras. Para este caso, en el eje horizontal se representan los valores que puede tomar la variable aleatoria y en el eje vertical la probabilidad asociada a cada valor. Por ejemplo, para el caso del ejemplo $3)$ anterior:
df = data.frame(evento = c('no llueve', 'llueve'), X = c(-1, 5), probabilidad=c(.55,.45))
df
ggplot(data = df, aes(x=X, y=probabilidad, fill=evento)) +
geom_col(width=1) +
ggtitle('Histograma') #aes help us to map variables
#in the data frame to objects
#in the graph
# La base de los rectángulos, por convención mide una unidad observemos que la suma de las áreas de los rectángulos siempre es $1$ pues ésta es simplemente la suma de las probabilidades de todos los valores posibles que toma la variable aleatoria, concretamente: $$\displaystyle \sum_{i=1}^nf(x_i) = 1.$$
# ## Transformaciones de variables aleatorias
# ### ¿Cómo hacerlo en R?
# Podemos usar la función [addrv](https://www.rdocumentation.org/packages/prob/versions/1.0-1/topics/addrv) del paquete [prob](https://cran.r-project.org/web/packages/prob/index.html) y se tienen $2$ opciones (la función `addrv` se aplica únicamente a **data frames**).
args(addrv)
# **Primera opción**
# La idea es escribir una fórmula que defina la variable aleatoria dentro de la función `addrv`. El resultado se añadirá como columna a un *data frame*.
# #### Ejemplo
S <- rolldie(3, nsides = 4, makespace = TRUE) #roll die three times.
#die has 4 sides
head(S)
nrow(S) #or: NROW(S)
# Defínase la variable aleatoria: $U = X1-X2+X3$. Entonces:
S <- addrv(S, U = X1-X2+X3)
print(head(S))
ncol(S)
# Se pueden responder preguntas del tipo:
Prob(S, U > 6)
# **Segunda opción**
# Usar una función de R o hecha por users como argumento a la función `addrv`. Por ejemplo, defínanse $V=\max(X1, X2, X3)$ y $W=X1+X2+X3$, entonces:
# +
S <- addrv(S, FUN = max,
invars = c("X1","X2","X3"),
name = "V") #max is defined in R
S <- addrv(S, FUN = sum, invars = c("X1","X2","X3"),
name = "W") #sum is defined in R
# -
print(head(S))
# **Obs:** obsérvese que se utilizó el argumento `invars` para especificar a qué variables se les debe aplicar la función definida en el argumento `FUN` (si no se especifica, `FUN` se aplica a todas las columnas diferentes de `probs`). También obsérvese que tiene un argumento `name` para nombrar a la nueva variable.
# ### Distribución marginal de variables aleatorias
# En los ejemplos anteriores se observa que añadir variables aleatorias al espacio de probabilidad (data frame al que se le añade la columna `probs`) con `addrv` resultan en renglones de la variable aleatoria en cuestión repetidos por ejemplo:
#
# |row|W|probs|
# |---|---|---|
# |2|4|0.015625|
# |5|4|0.015625|
# Se puede utilizar la función [marginal](https://www.rdocumentation.org/packages/prob/versions/1.0-1/topics/marginal) para agregar los renglones del espacio de resultados por los valores de la variable aleatoria y acumular la probabilidad asociada con los valores distintos de ésta:
marginal(S, vars = "V") #equivalently: sum(S[S$V==1,]$probs)
#for V=1
# Esto se conoce como la distribución de probabilidad marginal de $V$ o sólo distribución marginal.
# También es posible calcular la distribución de probabilidad conjunta entre $V$ y $W$:
marginal(S, vars = c("V", "W")) #equivalently: sum(S[S$V==3 & S$W==7,]$probs)
#for V=3 y W=7
# ## Valor esperado de variables aleatorias
# Se quiere hacer una rifa para recaudar fondos para alguna buena causa. Hay un premio único de $\$10,000$ y los boletos de la rifa cuestan $\$100$. Sea $n$ el número de boletos que se venden, de manera que la probabilidad de ganar el premio es de $\frac{1}{n}$. De esta forma, la compradora de un boleto tiene como valor esperado para esta rifa: $$\frac{1}{n}(10000-100) + \left(1-\frac{1}{n}\right)(-100)$$.
# y la organizadora de la misma tiene la ganancia segura de: $100n-10000$. Evidentemente la rifa no debe realizarse si no se venden al menos $100$ boletos, ya que la organizadora perdería dinero. La siguiente tabla muestra el valor esperado de la rifa para la compradora del boleto y la ganancia de la organizadora para algunos valores de $n$
#
# |n|Valor esperado (compradora)|Ganancia (organizadora)|
# |:----:|:----:|:----:|
# |100|0|0|
# |150|$-33.\bar{3}$|5000|
# |200|-50|10000|
# |500|-80|40000|
df_compradora <- data.frame(n=c(100,150,200,500), valor_esperado=c(0,-100/3,-50,-80))
ggplot(data=df_compradora, aes(x=n, y=valor_esperado)) +
geom_line(colour='coral3') +
ggtitle('Compradora')
df_organizadora <- data.frame(n=c(100,150,200,500),ganancia=c(0,5000,10000,40000))
ggplot(data=df_organizadora, aes(x=n,y=ganancia)) +
geom_line(colour='steelblue') +
ggtitle('Organizadora') +
theme(plot.title = element_text(size=10, hjust = 0.5)) #ajustar el título
# Supongamos que la variable aleatoria $X$ toma valores $x_1, \dots, x_n$ y sean $f(x_1),\dots,f(x_n)$ los valores de la distribución de probabilidad asociada. Definimos el valor esperado o esperanza de $X$ como: $$E(X) = \displaystyle \sum_{i=1}^nx_iP(X=x_i) = \sum_{i=1}^nx_if(x_i).$$
# **Obs:** obsérvese que si todos los valores son equiprobables, esto es: $P(X=x_1) = P(X = x_2) = \cdots = P(X = x_n) = \frac{1}{n}$, el valor esperado de $X$ es: $$E(X) = \frac{1}{n}\displaystyle\sum_{i=1}^nx_i.$$
# Así, el valor esperado no es más que un **promedio ponderado por la probabilidad de obtener $x_i$** como resultado.
# ### Ejemplos
# 1) Considérese la situación del ejemplo anterior del evento "el día de hoy llueve" y las variables aleatorias $X$ y $Y$ que representan un número que asocia mi satisfacción con la situación "llevar paraguas" o "no llevar paraguas" respectivamente y toman valores: $X(E) = 5, X(E^c) = -1, Y(E) = -6, Y(E^c)=4$ con probabilidades dadas por la tabla:
# |Evento|Valor de X|Valor de Y|P(Variable aleatoria = valor)|
# |:----:|:----:|:----:|:----:|
# |llueve|5|-6|f(5) = P(X=5) = g(-6) = P(Y=-6) = .45|
# |no llueve|-1|4|f(-1) = P(X = -1) = g(4) = P(Y = 4) = .55|
# Entonces los valores esperados de $X$ y $Y$ son: $$E(X) = 5*.45 - 1*.55 = 1.7$$
# $$E(Y) = -6*.45+4*.55 = -0.5$$
# Interpretemos esta esperanza como el valor que nos proporcionan las acciones de llevar o no el paraguas. De acuerdo a los valores obtenidos, $E(X) > E(Y)$, equivalentemente, la acción de “llevar el paraguas” nos lleva a una consecuencia con mayor valor esperado. Si somos racionales, elegiremos llevar el paraguas.
# ## El caso continuo
# Existen datos en la naturaleza que no podemos contar sino más bien **medir**, por ejemplo distancias, tiempo o peso de una persona. Para este caso las variables aleatorias pueden tomar valores en un continuo como un intervalo de la recta real o un cuadrado en dos dimensiones o un cubo en tres dimensiones. No tiene sentido definir medidas de probabilidad para eventos de la forma {mi peso es igual a 65 kg} o {a las 15:04 lleguen por mi} pues si recordamos que la distribución de probabilidad de una variable aleatoria se representa con el área de un rectángulo con base igual a $1$ y $X$ mide mi peso en kg entonces que $X$ tome un valor en particular generaría una línea que no tiene área. Para estos casos consideramos eventos de la forma {mi peso se encuentre en un intervalo entre 63 y 70 kg} o {lleguen por mí entre 15:00 y 15:15}.
# Así para el caso continuo, no se utilizan técnicas de conteo (principio multiplicativo, permutaciones o combinaciones) pues implican una forma de contar el número de elementos en algún conjunto; es indispensable para este caso utilizar **técnicas de medida** como longitud de un intervalo, área de un cuadrado o volumen de un cubo.
# ### Ejemplo:
# 1) Imaginemos un juego de dardos, en el cual el tirador es malísimo y los lanza sin ton ni son. El tablero es un cuadro formado por nueve cuadrados más pequeños, como se muestra a continuación:
# <img src="https://dl.dropboxusercontent.com/s/6067r9ief3xsv76/var_aleatorias_2.jpg?dl=0" heigth="200" width="200">
#
# La probabilidad de que el dardo caiga en algún color puede calcularse como la proporción del área que dicho color representa como parte del área total. Entonces: $$P(\{blanco\}) = \frac{4}{9}, P(\{negro\}) = \frac{3}{9}, P(\{gris\}) = \frac{2}{9}.$$
# Nótese que la probabilidad de que el dardo caiga en un punto particular es una probabilidad nula pues un punto carece de área. La solución sería pensar en una pequeña superficie alrededor del punto y calcular su área.
# 2) Pensemos en el siguiente juego de azar: se toma el intervalo $[0,1]$ de la recta numérica, es decir, todos los números (reales) entre el $0$ y el $1$ y se elige aleatoriamente un número del intervalo. Podemos pensar a este juego como una ruleta continua y asociarle una variable aletoria $X$ que representa al número obtenido. Todos los números son equiprobables de manera que la probabilidad de que $X$ caiga en el intervalo $[0,\frac{1}{2}]$ es: $$P(0 \leq X \leq \frac{1}{2}) = \frac{1}{2}.$$
#
# La razón de esta estimación es que $[0,\frac{1}{2}]$ mide la mitad del intervalo $[0,1]$, en donde medida se refiere a la longitud. Asimismo, $$P(\frac{1}{3} \leq X < 1) = \frac{2}{3},$$ ya que la longitud del intervalo $(\frac{1}{3},1)$ es $\frac{2}{3}$ partes del intervalo $[0,1]$.
# 3) Supongamos que ahora la ruleta continua consiste en elegir numeros en el intervalo $[0,4]$ y $X$ es la variable aleatoria que representa al número resultante. Entonces, $$P(0 \leq X \leq 3) = \frac{3}{4}$$ en donde podemos pensar a este número como $\frac{\text{longitud de } (0,3)}{\text{longitud de } [0,4]}$.
# Nótese que en los ejemplos $2$ y $3$ anteriores la medida o longitud del intervalo $(a, b)$ es la misma que la del intervalo cerrado $[a, b]$. En general, la longitud de cualquiera de los intervalos $(a, b), [a, b], (a, b]$ y $[a, b)$ es de $b-a$.
# Si pensamos en el intervalo original $[a,b]$ como nuestro espacio muestral y tomamos $c$ y $d$ tales que $a \leq c \leq d \leq b$, entonces, la probabilidad del evento “el número cae entre $c$ y $d$” es, $$P(c < X < d) = \frac{\text{longitud de } [c,d]}{\text{longitud de } [a,b]} = \frac{d-c}{b-a}.$$
# En el caso continuo, **la longitud o medida del intervalo toma el lugar del número de elementos** en el caso discreto. Evidentemente, en el caso del tablero de dardos descrito arriba medimos áreas en lugar de medidas lineales de longitud.
# ### ¿Distribución de probabilidad de una variable aleatoria continua?
# Si $X$ es una variable aleatoria continua, no podemos definir la distribución de probabilidad como en el caso discreto ya que no podemos hablar de la probabilidad que $X$ tome un valor específico. Existe, sin embargo, una densidad de probabilidad, que es una curva tal que el área debajo de la curva, entre a y b, representa a la probabilidad de que $X$ tome valores entre $a$ y $b$, es decir, $P (a < X < b)$. Visualmente tenemos:
# <img src="https://dl.dropboxusercontent.com/s/ncls12iyzrrxs01/var_aleatorias_1.jpg?dl=0" heigth="500" width="500">
# Como en el caso discreto,la curva es también el conjunto de puntos $(x,f(x))$ para la función de densidad $f$. La diferencia es que en el caso continuo, $f(a)\neq P(X = a)$.
# En el caso discreto la suma de las probabilidades de todos los valores que toma la variable aleatoria es la unidad. Aquí se tiene el resultado análogo: el área total bajo la función de densidad que representa la probabilidad de que la variable $X$ tome cualquier valor (real), es la unidad, es decir, $$P(-\infty < X < \infty) = 1.$$
# Por ejemplo:
set.seed(2000)
n <- 1000
df <- data.frame(x=rnorm(n))
y <- dnorm(df$x)
df$y <- y
gf <- ggplot(data = df, aes(x=x, y=y)) + geom_point(size=.01) + ggtitle('rnorm') +
theme(plot.title = element_text(size=10, hjust = 0.5))
gf
gf + geom_smooth(formula='y~x', method='loess', colour='red', size=.5) + #lowess: locally weighted
#regression
geom_text(x=0,y=0.1,label='area=1', size=5)
# Para el cálculo del valor esperado de una variable aleatoria continua es necesario utilizar el **cálculo integral** para evaluarlo. Sin embargo, el valor esperado tiene una misma interpretación: representa un promedio ponderado de los valores que toma la variable aleatoria. La notación que se utiliza es la misma que antes, es decir, si $X$ es una variable aleatoria discreta o continua, su valor esperado se representa por $E(X)$. En el caso continuo se tiene: $$E(X)=\displaystyle \int_{X \in S}xf(x)dx.$$
# si $\displaystyle \int_{X \in S}|x|f(x)dx < \infty$ (es finita).
# ## (Tarea) Ejercicios
# **Nota: Varios de los siguientes ejercicios tienen su solución en las referencias dadas al inicio de la nota. Se sugiere resolverlos por su cuenta y posteriormente compararlos con sus respectivas soluciones.**
# 1) Considérese lanzar dos monedas distintas. Sea $X$ la variable aleatoria que cuenta el número de águilas que aparecen. Calcula la probabilidad de $P(X=0), P(X=1), P(X=2)$.
# 2) Considérese el sorteo de Melate (elegir una combinación de seis números diferentes entre el $1$ y el $56$ sin orden y sin reemplazo) "simplificado" con una bolsa acumulada de $N$ y la variable aleatoria $X$ que toma dos valores: $N$ si se elige la combinación ganadora y $0$ en cualquier otro caso. ¿Cuál es la distribución de probabilidad de $X$?
#
# (es simplificado pues en realidad la bolsa acumulada se reparte entre el número de personas ganadoras y varios premios menores, según el número de cifras acertadas entre dos y seis.)
# 3) Calcular el valor esperado de los ejemplos $1)$ y $3)$ de la sección "Distribución de probabilidad de una variable aleatoria" y responder: ¿Si nos ofrecieran jugar uno de estos dos juegos cuál deberíamos elegir?. Nótese que ninguno de los premios o pérdidas es igual al valor esperado, éste simplemente nos da una idea del promedio de los premios y pérdidas que resultarían si el juego se repitiese un gran número de veces.
#
# 4) Los juegos de azar en los casinos pueden representarse por medio de variables aleatorias. Consideremos una versión simplificada de una máquina de palanca: existen dos figuras, digamos un cuadrado y un triángulo, que pueden aparecer en tres columnas de una cinta circular. El jugador baja la palanca, las cintas giran y al detenerse cada columna muestra un cuadrado o un triángulo. Las $8$ posibilidades equiprobables son:
#
# $$\begin{array}{ccc} \blacksquare &\blacksquare& \blacksquare \\ \blacksquare & \blacksquare& \blacktriangle \\ \blacksquare & \blacktriangle & \blacksquare \\ \blacksquare & \blacktriangle & \blacktriangle \\ \blacktriangle & \blacksquare & \blacksquare \\ \blacktriangle & \blacksquare & \blacktriangle \\ \blacktriangle & \blacktriangle & \blacksquare \\ \blacktriangle & \blacktriangle & \blacktriangle \end{array}$$
#
# Supongamos que para jugar se introduce una ficha de $\$300$ y los premios asociados son: $\$500$ si salen tres figuras iguales y $\$0$ de cualquier otra forma. Sea $X$ la variable aleatoria que mide la ganancia o pérdida de jugar en la máquina. Calcular $E(X)$.
# Los siguientes ejercicios realizarlos en `R` y con `ggplot2`:
# 5) Encontrar los histogramas de probabilidad para las distribuciones asociadas a las siguientes variables aleatorias:
#
# a) El número de águilas que aparecen cuando lanzamos tres monedas al aire.
#
# b) La suma de los números de las caras cuando se lanzan dos dados.
#
# 6) Se venden 8000 boletos para una rifa de $\$5000.00$ y cada boleto cuesta $\$2.00$.
#
# a) Encontrar la ganancia esperada del comprador de un boleto.
#
# b) Hacer la gráfica de la ganancia esperada que tiene una compradora en términos de un número de boletos $n$.
#
# c) ¿Cuál debería de ser el premio mínimo para que se pudiese garantizar “salir a mano” al comprar todos los boletos.
| R/clases/2_probabilidad/7_variables_aleatorias.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Canada Population Vs GDP DataFrame
# ### In This notebook we will import Canada's GDP and Population data from world bank and Visualize it
import os
os.getcwd()
# +
from pyspark.sql import Row
import matplotlib.pyplot as plt
import numpy as np
import matplotlib.pyplot as plt
import pylab as P
from pyspark import SparkContext
from pyspark.sql import SparkSession
from pyspark.sql.types import *
spark = SparkSession.builder.master("local").appName("Canada").getOrCreate()
get_ipython().magic('matplotlib inline')
plt.rcdefaults()
# -
static = spark.read.format("csv").option("header","true").option("inferSchema","true").load("/home/jovyan/GDP_EX.csv")
static.show(20)
static.createOrReplaceTempView("canada")
GDP = spark.sql("SELECT GDP FROM canada")
GDP.describe().show()
GDPList = GDP.rdd.map(lambda p: p.GDP).collect()
plt.hist(GDPList)
plt.title("GDP Distribution\n")
plt.xlabel("GDP")
plt.ylabel("freq")
plt.show(block=False)
emissions = spark.sql("SELECT greenhouse_emissions FROM canada")
emissions.describe().show()
EList = emissions.rdd.map(lambda p: p.greenhouse_emissions).collect()
plt.hist(EList)
plt.title("GDP Distribution\n")
plt.xlabel("GDP")
plt.ylabel("freq")
plt.show(block=False)
from scipy.stats import gaussian_kde
density = gaussian_kde(EList)
xAxisValues = np.linspace(0,100,1000) # Use the range of ages from 0 to 100 and the number of data points
density.covariance_factor = lambda : .5
density._compute_covariance()
plt.title("GDP density\n")
plt.xlabel("GDP")
plt.ylabel("Density")
plt.plot(xAxisValues, density(xAxisValues))
plt.show(block=False)
plt.subplot(121)
plt.hist(EList)
plt.title("Emmisions distribution\n")
plt.xlabel("Age")
plt.ylabel("Number of users")
plt.subplot(122)
plt.title("Summary of distribution\n")
plt.xlabel("Age")
plt.boxplot(EList, vert=False)
plt.show(block=False)
Top10 = spark.sql("SELECT Male_population, SUM(GDP_per_capita) as GDP_per_capita FROM canada GROUP BY Male_population LIMIT 10")
Top10.show()
Male = spark.sql('SELECT Male_population FROM canada')
Male.describe().show()
MList = Male.rdd.map(lambda p: p.Male_population).collect()
Female = spark.sql('SELECT female_population FROM canada')
FList = Female.rdd.map(lambda p: p.female_population).collect()
plt.scatter(MList, FList, marker='o');
TS = spark.sql('SELECT YEAR, Male_population, female_population, total_population from canada ORDER BY YEAR')
TS.show(3)
series = TS.toPandas()
series.plot(figsize=(20,10), linewidth=5, fontsize=20)
plt.xlabel('YEAR', fontsize=20);
plt.ylabel('Population in Millions', fontsize=20)
plt.title('Time Series Graph', fontsize=40)
percapita = spark.sql('SELECT (GDP_per_capita*10000) as percapita from canada ORDER BY YEAR')
percapita.show(3)
perList = percapita.rdd.map(lambda p: p.percapita).collect()
# +
rng = np.random.RandomState(0)
x = rng.randn(28)
y = rng.randn(28)
colors = rng.rand(28)
sizes = 1000 * rng.rand(28)
plt.scatter(GDPList, perList, c=colors, s=sizes, alpha=0.3, cmap='viridis')
plt.xlabel('GDP', fontsize=20);
plt.ylabel('GDP Per Capita', fontsize=20)
plt.title('Scatter Plot', fontsize=30)
plt.colorbar();
# -
| notebooks/Canada_Viz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Loan Prediction 04 - Data Imputation With Random Forest
# Let us try to improve the previous results by imputing missing data with a Random Forest.
#
# But first, we will remove rows with more than one missing value.
# +
import sys
sys.path.append('utils')
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import math
plt.style.use('seaborn')
from missingpy import MissForest
from sklearn.preprocessing import OrdinalEncoder
import dataframe_utils
import preprocess_utils
# -
# ### Loading original dataset
df_import = pd.read_csv('dataset/train_loan.csv')
df_import.drop(columns=['Loan_ID'],inplace = True)
df_import.shape
# ### Counting missing values by column
df_import.isnull().sum()
# ### Rows with one or no missing value
null_rows = df_import.isnull().sum(axis = 1)
df_rows = df_import.loc[(null_rows == 1) | (null_rows == 0),:]
df_rows
# ## Replacing missing values with MissForest
# Let us prepare the dataset in order to execute the MissForest algorithm
ordinal_encoder = OrdinalEncoder()
ordinal_encoder.categories_ = np.load('saves/variable_encoder_categories.npy', allow_pickle= True)
categorical_columns = ['Gender', 'Married', 'Dependents', 'Education', 'Self_Employed', 'Property_Area','Credit_History','Loan_Amount_Term','Loan_Status']
df_encoded_nans = preprocess_utils.encode_with_nan(df_rows, categorical_columns, ordinal_encoder)
df_encoded_nans
# Although Credit_History and Loan_Amount_Term are represented as numerical values, we will set them as categorical variables.
#
# That is because these variables have a categorical behavior, as shown below.
dataframe_utils.show_column_options(df_import[['Credit_History','Loan_Amount_Term']])
# +
X = df_encoded_nans.copy().drop(columns = ['Loan_Status'])
y = df_encoded_nans.copy()[['Loan_Status']]
categorical_index = [0,1,2,3,4,8,9,10]
imputer = MissForest(oob_score=True, random_state = 0, class_weight = 'balanced')
imputer.fit(X,y,cat_vars = categorical_index)
# -
X_filled = imputer.transform(X)
df_fill = pd.DataFrame(X_filled,columns = X.columns)
df_fill
df_fill['Loan_Status'] = y.values.ravel()
# ## Pre processing after Imputation
# ### Calculating Base_Loan_Installment and Remaining_Income
# +
base_loan_installment = df_fill['LoanAmount'] * 1000 / df_fill['Loan_Amount_Term']
total_income = df_fill['ApplicantIncome'] + df_fill['CoapplicantIncome']
remaining_income = (total_income - base_loan_installment) / total_income
df_fill['Base_Loan_Installment'] = base_loan_installment
df_fill['Remaining_Income'] = remaining_income
# -
# ### Removing outliers
numerical_columns = ['ApplicantIncome', 'CoapplicantIncome', 'LoanAmount','Base_Loan_Installment','Remaining_Income']
df_fill_no_outlier = preprocess_utils.remove_outliers(df_fill,numerical_columns,threshold = 3)
# ### Envoding Loan_Status
df_fill_no_outlier.loc[df_fill_no_outlier['Loan_Status'] == 'Y','Loan_Status'] = 1
df_fill_no_outlier.loc[df_fill_no_outlier['Loan_Status'] == 'N','Loan_Status'] = 0
df_fill_no_outlier.to_csv('dataset/train_rf_imputed.csv',index = False)
| 04_miss_forest_imputation_pre_processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Computer Lab 1: Part 2. Naive Models
#
# In this practical will apply our knowledge in
#
# * Creating baseline naive forecasts
# * Performing a train-test split
# * Using forecast error metrics MAE and MAPE to select the best method
# # Standard Imports
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import sys
# -
# # Install forecast-tools
# install forecast-tools if running in Google Colab
# if you are not using hsma4_forecast conda env then you need to pip install it!
if 'google.colab' in sys.modules:
# !pip install forecast-tools
# # forecast-tools imports
# +
# baseline forecast methods
from forecast_tools.baseline import (Naive1,
SNaive,
Drift,
Average,
baseline_estimators)
from forecast_tools.metrics import (mean_absolute_percentage_error,
mean_absolute_error)
# -
# # Helper functions
def preds_as_series(data, preds):
'''
Helper function for plotting predictions.
Converts a numpy array of predictions to a
pandas.DataFrame with datetimeindex
Parameters
-----
data - arraylike - the training data
preds - numpy.array, vector of predictions
Returns:
-------
pandas.DataFrame
'''
start = pd.date_range(start=data.index.max(), periods=2,
freq=data.index.freq).max()
idx = pd.date_range(start=start, periods=len(preds), freq=data.index.freq)
return pd.DataFrame(preds, index=idx)
# ## Exercise 1: Using Naive1 to forecast monthly outpatient appointments.
# **Step 1: Import monthly outpatient appointments time series**
#
# This can be found in **"data/out_appoints_mth.csv"**
# or https://raw.githubusercontent.com/hsma4/module_9_a/main/data/out_appoints_mth.csv
#
# * Hint: this is monthly data. You can use the monthly Start ('MS') frequency
# your code here ...
url = 'https://raw.githubusercontent.com/hsma4/module_9_a/main/data/' \
+ 'out_appoints_mth.csv'
appoints = pd.read_csv(url,
index_col='date', parse_dates=True, dayfirst=True)
appoints.index.freq = 'MS'
appoints.info()
# **Step 2 Plot the data**
# +
# your code here ...
# -
_ = appoints.plot(figsize=(12,4))
# **Step 3: Create and fit Naive1 forecast model**
#
# * Hint: you want to fit `appoints['out_apts']`
# +
# your code here ...
# -
nf1 = Naive1()
nf1.fit(appoints['out_apts'])
# **Step 4: Plot the Naive1 fitted values**
#
# All the baseline models have fitted values. These are the in-sample prediction i.e. the predictions of the training data.
#
# Once you have created and fitted a Naive1 model you can access the fitted values using the `.fittedvalues` property. This returns a `DataFrame`.
#
# Plot the fitted values against the observed data.
# +
# your code here ...
# -
ax = appoints.plot(figsize=(12,4))
_ = nf1.fittedvalues.plot(ax=ax, color='green', linestyle='--')
# **Step 5: Forecast the next 6 months**
#
# After you have created a forecast plot the predictions.
#
# * Hint: use the `pred_as_series()` method to plot the predictions. See the lecture notes for exampes of how to use it.
# +
# your code here ...
# -
preds = nf1.predict(horizon=6)
preds = preds_as_series(appoints, preds)
ax = appoints.plot(figsize=(12,4), marker='o')
preds.plot(ax=ax, marker='o')
_ = ax.legend(['training', 'forecast'])
# ## Exercise 2. Choose the best baseline forecast method for ED reattendances
# **Step 1: Import emergency department reattendance data.**
#
# This is a time series from a hospital that measures the number of patients per month that have reattended an ED within 7 days of a previous attendance.
#
# This can be found in **"ed_reattend.csv"**:
# https://raw.githubusercontent.com/hsma4/module_9_a/main/data/ed_reattend.csv
#
# * Hint 1: The format of the 'date' column is in UK standard dd/mm/yyyy. You will need to set the `dayfirst=True` of `pd.read_csv()` to make sure pandas interprets the dates correctly.
#
# * Hint 2: The data is monthly and the dates are all the first day of the month. This is called monthly start and its shorthand is 'MS'
# your code here ...
url = 'https://raw.githubusercontent.com/hsma4/module_9_a/main/data/' \
+ 'ed_reattend.csv'
reattends = pd.read_csv(url,
index_col='date', parse_dates=True, dayfirst=True)
reattends.index.freq = 'MS'
# **Step 2: Perform a calender adjustment**
# +
# your code here ...
# -
reattend_rate = reattends['reattends'] / reattends.index.days_in_month
# **Step 3: Perform a train-test split**
#
# Create a train test split where you holdback the final 6 months of the data.
#
# Remember to work with the calender adjusted data.
#
# * Hint: The test set is the last 6 rows in your pandas DataFrame
# +
#your code here ...
# -
# train test split
train = reattend_rate.iloc[:-6]
test = reattend_rate.iloc[-6:]
train.shape
test.shape
# **Step 4: Plot the TRAINING data**
#
# Remember don't look at the test data just yet. You don't want to bias your model selection process.
# +
# your code here ...
# -
ax = train.plot(figsize=(12,4), color='red', marker='o', legend=False)
_ = ax.set_ylabel('mean reattends within 7 days')
# **Step 5: Create and fit Naive1, and SNaive baseline models**
#
# * Hint: Fit the TRAINING data.
# +
# your code here ...
# -
model_1 = Naive1()
model_2 = SNaive(12)
model_1.fit(train)
model_2.fit(train)
# **Step 6: Use each model to predict 6 months ahead**
#
# * Hint. You need to store the prediction results so that later on you can calculate the forecast error.
# +
# your code here ...
# -
preds_1 = model_1.predict(horizon=6)
preds_2 = model_2.predict(horizon=6)
# **Step 7: Calculate the mean absolute error of each forecast method**
#
# Based on the results which method would you choose?
# +
# your code here ...
# +
mae_1 = mean_absolute_error(y_true=test, y_pred=preds_1)
mae_2 = mean_absolute_error(y_true=test, y_pred=preds_2)
print(mae_1)
print(mae_2)
# -
# **Step 8: Calculate the out of sample MAPE of each forecast method**
#
# Would you still choose the same forecasting method?
#
# Is it useful to calculate both metrics?
# +
mape_1 = mean_absolute_percentage_error(y_true=test, y_pred=preds_1)
mape_2 = mean_absolute_percentage_error(y_true=test, y_pred=preds_2)
print(mape_1)
print(mape_2)
| exercises/Practical_2_SOLUTIONS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:nlp]
# language: python
# name: conda-env-nlp-py
# ---
#imports
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import pickle
import itertools
import spacy
import nltk
# %matplotlib inline
from sklearn.model_selection import train_test_split
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import GridSearchCV
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer, TfidfTransformer
from sklearn.metrics import accuracy_score, confusion_matrix, classification_report
from sklearn.pipeline import Pipeline
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.naive_bayes import MultinomialNB
from sklearn.svm import SVC, LinearSVC
from sklearn.ensemble import AdaBoostClassifier
from sklearn.ensemble import RandomForestClassifier
# dataframe display options
pd.set_option('display.max_colwidth', -1)
pd.set_option('display.max_rows', 200)
# + [markdown] slideshow={"slide_type": "slide"}
# # Problem Definition
# <img src='../images/form2.png'>
# -
# > The Objective of the project is to use Natural Language Processing on two fields entered by the user ("event title" and "additional info") to guide the user into selecting the correct category.
# + **"Event type" (category)** is a key field used by the company to match events and the capabilities of their pool of photographers.
# + [markdown] slideshow={"slide_type": "slide"}
# # Problem Definition
#
# Classification task: Predicting discrete-valued quantity $y$
# # + Multi-class $ y \in \{1,2,3 \ldots k\}$
#
#
# # Data Acquisition
#
# <img src='../images/data.png'>
# -
# In order to validate the possibility of a project I scraped information on 100 events. Links to the events were provided by the company.
# +
def search_titles(df, expression):
categories = df[df['category'].str.contains(expression, regex=True) == True]['category']
idxs = df[df['category'].str.contains(expression, regex=True) == True].index
print(categories)
return idxs
def update_category(df, indices, category_title):
for i in indices:
df.loc[i, 'category'] = category_title
# -
b = search_titles(g, r'baptism')
update_category(g, b, 'life celebration')
# ## After Categories were regrouped
# + slideshow={"slide_type": "slide"}
g['category'].value_counts()[:20][::-1].plot(kind='barh');
plt.title("Top 20 categories");
# -
## Pickle data after cleanup
with open('events.pickle', 'wb') as f:
pickle.dump(g, f)
reset -fs
# # Loading events dataframe from pickled object
with open('events.pickle', 'rb') as f:
g = pickle.load(f)
categories = g.category.unique()
print("Number of categories: {}".format(len(categories)))
g['category'].value_counts()[:20][::-1].plot(kind='barh');
plt.title("Top Categories");
g['category'].value_counts()[:20]
g.tail()
# # EDA
length = g[['title', 'additional_info', 'category']].copy()
length['title'][0]
# ### How many characters are in "title" and "additional_info"?
length['title_length'] = length['title'].map(lambda text: len(str(text)))
length.head()
length['additional_info_length'] = length['additional_info'].map(lambda text: len(str(text)))
length.head()
length.title_length.plot(bins=20, kind='hist');
plt.title("Title - Character length");
length.additional_info_length.plot(bins=15, kind='hist');
plt.title("Additional Info - Character length");
# > We can see that "additional info" is an optional field and that overall length of each of the samples is small.
length[length["category"] == 'kids birthday']["title_length"].plot(bins=20, kind='hist');
plt.title("KIDS BIRTHDAY - Title - Character length");
| notebooks/DSCI 6003-6004 - Introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.10.1 64-bit (''motorcycles-prices'': conda)'
# language: python
# name: python3
# ---
# !pip install opendatasets
import opendatasets as op
import os
dataset = 'https://www.kaggle.com/ropali/used-bike-price-in-india'
data_dir = '../data/raw'
op.download(dataset, data_dir=data_dir)
os.listdir(data_dir)
| notebooks/0.0-motorcycles-data-collect.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
BASE_URL = 'http://localhost:8000'
AUTH = ('admin', 'admin')
TOKEN = '8bd704f41e0def27e0406e5ae8ec479ce575aa8f'
URI_PREFIX = 'https://rdmorganiser.github.io/terms/'
from rdmo_client import Client
client = Client(BASE_URL, auth=AUTH)
# -
client.list_catalogs(key='catalog')
client.list_sections(catalog=1)
client.list_questions(is_collection=False)
client.list_questions(widget_type='radio')
| notebooks/questions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import numpy as np
import pandas as pd
from skimage import filters
import matplotlib.pyplot as plt
with np.load('dataset/kay_images.npz') as dobj:
data = dict(**dobj)
images = np.zeros(data["stimuli"].shape)
for i in range(0,1750):
images[i] = cv2.normalize(data["stimuli"][i], None, 0, 255, cv2.NORM_MINMAX, cv2.CV_8U)
thresholds = np.array([filters.threshold_otsu(image) for image in images])
luminance = np.array([images[i][images[i]>thresholds[i]].mean() for i in range(0,1750)])
luminance.shape
| .ipynb_checkpoints/luminance_calculation-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# <div class="contentcontainer med left" style="margin-left: -50px;">
# <dl class="dl-horizontal">
# <dt>Title</dt> <dd> Histogram Element</dd>
# <dt>Dependencies</dt> <dd>Matplotlib</dd>
# <dt>Backends</dt> <dd><a href='./Histogram.ipynb'>Matplotlib</a></dd> <dd><a href='../bokeh/Histogram.ipynb'>Bokeh</a></dd>
# </dl>
# </div>
import numpy as np
import holoviews as hv
hv.extension('matplotlib')
# ``Histogram``s partition the `x` axis into discrete (but not necessarily regular) bins, showing counts in each as a bar. A ``Histogram`` accepts the output of ``np.histogram`` as input, which consists of a tuple of the histogram values with a shape of ``N`` and bin edges with a shape of ``N+1``. As a simple example we will generate a histogram of a normal distribution with 20 bins.
np.random.seed(1)
data = np.random.randn(10000)
frequencies, edges = np.histogram(data, 20)
print('Values: %s, Edges: %s' % (frequencies.shape[0], edges.shape[0]))
hv.Histogram(frequencies, edges)
# The ``Histogram`` Element will also expand evenly sampled bin centers, therefore we can easily cast between a linearly sampled Curve or Scatter and a Histogram.
xs = np.linspace(0, np.pi*2)
ys = np.sin(xs)
curve = hv.Curve((xs, ys))
curve + hv.Histogram(curve)
# The ``.hist`` method is an easy way to compute a histogram from an existing Element:
# +
points = hv.Points(np.random.randn(100,2))
points.hist(dimension=['x','y'])
# -
# The ``.hist`` method is just a convenient wrapper around the ``histogram`` operation that computes a histogram from an Element, and then adjoins the resulting histogram to the main plot. You can also do this process manually; here we create an additional set of ``Points``, compute a ``Histogram`` for the 'x' and 'y' dimension on each, and then overlay them and adjoin to the plot.
# +
# %%opts Histogram (alpha=0.3)
from holoviews.operation import histogram
points2 = hv.Points(np.random.randn(100,2)*2+1).redim.range(x=(-5, 5), y=(-5, 5))
xhist, yhist = (histogram(points2, bin_range=(-5, 5), dimension=dim) *
histogram(points, bin_range=(-5, 5), dimension=dim)
for dim in 'xy')
(points2 * points) << yhist(plot=dict(width=125)) << xhist(plot=dict(height=125))
| examples/reference/elements/matplotlib/Histogram.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.1 64-bit
# name: python38164bit8a7f52ad8ff0431c85e03ecdbd324643
# ---
# +
# !pip install numpy
# !pip install scipy
# !pip install matplotlib
#Import the neccesary packages
import numpy as np
import pandas as pd
import pydst
import matplotlib.pyplot as plt
#DATACLEANING#
#We use pydst to use an API to Denmark's statistics
Dst = pydst.Dst(lang='en')
Dst.get_data(table_id='INDKP101')
Vars = Dst.get_variables(table_id='INDKP101')
#To find the variables we need, we inspect the table that we have imported:
Vars.values
#After picking out values, we can get our data:
Everything = Dst.get_data(table_id = 'INDKP101', variables={'OMRÅDE':['000','01','02','03','04','05','06','07','08','09','10','11'], 'KOEN':['M','K'], 'TID':['*'], 'ENHED':['116'], 'INDKOMSTTYPE':['100']}).rename(columns={'OMRÅDE':'Municipality'})
#Changing the index to Municipality:
New_index = Everything.set_index('Municipality')
#Picking out the neccesary variables:
Sortet = New_index[['TID','KOEN','INDHOLD']].rename(columns={'KOEN':'Gender', 'TID':'Year', 'INDHOLD':'disposable income'})
#Making a table for each gender:
Men = Sortet[Sortet['Gender']=='Men'].sort_values(['Municipality','Year']).rename(columns={'disposable income':'disposable_income_men'})
Women = Sortet[Sortet['Gender']=='Women'].sort_values(['Municipality', 'Year']).rename(columns={'disposable income':'disposable_income_women'})
#We don't want year to appear twice when we concat:
Women_without_year = Women[['Gender', 'disposable_income_women']]
#Concatenate the two tables:
Concatenated_table = pd.concat([Men, Women_without_year], axis=1)
#Removing the gender nicer look:
Final_table = Concatenated_table[['Year','disposable_income_men','disposable_income_women']]
#DATACLEANING COMPLETE#
#APPLYING METHODS#
#Creates a function with provides the difference between the genders in %:
def f(x):
"""Gives the procentual difference between the genders"""
return round((x['disposable_income_men']/x['disposable_income_women']-1)*100, 2)
#Applying the function to the end of the table:
Final_table['Difference in %']=Final_table.apply(f, axis=1)
#We now wish to create a individual table for each province, and do it like this:
#We start by finding the unique values in the table AKA all the provinces
unik = Final_table.index.unique()
#Making an empty dictionary which will contain our unique values with their seperate table later
d = {}
#Filling the empty dictionary
for i in unik:
d.update( {i : Final_table.loc[i]})
#We can now plot the difference between men and women in a graph like this:
def Difference(Region):
#Simply plotting the difference against years to see the evolution
plt.plot(d[Region]['Year'],d[Region]['Difference in %'])
plt.xlabel('Year')
plt.ylabel('Difference in %')
plt.title(f'Difference in disposable_income for {str(Region)}')
plt.axis([1986,2018,9,32.5])
plt.grid(True)
return plt.show()
#To compare the genders visually, we create two normal distributions:
def normal(Region):
#Making subplots to be shown in the same figure:
plt.subplot(2,1,1)
#Creating the normal distribution for the men:
s = np.random.normal(d[Region]['disposable_income_men'].mean(), d[Region]['disposable_income_men'].std(), 10000)
count, bins, ignored = plt.hist(s, 30, density=True)
#Plotting the distribution:
plt.plot(bins, 1/(d[Region]['disposable_income_men'].std() *np.sqrt(2*np.pi)) * np.exp(-(bins-d[Region]['disposable_income_men'].mean())**2 / (2 * d[Region]['disposable_income_men'].std()**2)), linewidth = 4)
#Some formal stuff
plt.title(f'Men in {str(Region)}')
plt.xlabel('Disposable income')
plt.axis([0,300000,0,0.000011])
#The other subplot:
plt.subplot(2,1,2)
#Creating the normal distribution for the women:
s = np.random.normal(d[Region]['disposable_income_women'].mean(), d[Region]['disposable_income_women'].std(), 10000)
count, bins, ignored = plt.hist(s, 30, density=True)
#Plotting the distribution:
plt.plot(bins, 1/(d[Region]['disposable_income_women'].std() *np.sqrt(2*np.pi)) * np.exp(-(bins-d[Region]['disposable_income_women'].mean())**2 / (2 * d[Region]['disposable_income_women'].std()**2)), linewidth = 4)
#Formal figure stuff again
plt.title(f'Women in {str(Region)}')
plt.xlabel('Disposable income')
plt.axis([0,300000,0,0.000011])
plt.subplots_adjust(top=2, bottom=0, left=0, right=1, hspace=0.2)
return plt.show(), print('For men, the mean is ','{0:.0f}'.format(d[Region]['disposable_income_men'].mean()), 'and the standard deviation is ','{0:.0f}'.format(d[Region]['disposable_income_men'].std())), print('For women, the mean is ','{0:.0f}'.format(d[Region]['disposable_income_women'].mean()), 'and the standard deviation is ','{0:.0f}'.format(d[Region]['disposable_income_women'].std()))
#Graph that shows the growth in disposable income over the years
def growth(Region):
plt.plot(d[Region]['Year'], d[Region]['disposable_income_men'], label = 'Men')
plt.plot(d[Region]['Year'], d[Region]['disposable_income_women'], label = 'Women')
plt.ylabel('Disposable income')
plt.gca().legend(('Men', 'Women'))
plt.title(f'{str(Region)}')
return plt.show()
# -
| Project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from tensorflow.python.summary.summary_iterator import summary_iterator
it = summary_iterator("../runs/Jan01_15-45-00_DESKTOP-I8HN3PB_pvc1_shallownet/events.out.tfevents.1609533900.DESKTOP-I8HN3PB.3117.0")
# +
import tensorflow as tf
from tensorflow.python.summary.summary_iterator import summary_iterator
from tensorboard.plugins.hparams import plugin_data_pb2
si = summary_iterator(
"../runs/Jan01_15-45-00_DESKTOP-I8HN3PB_pvc1_shallownet/1609533900.105029/events.out.tfevents.1609533900.DESKTOP-I8HN3PB.3117.1"
)
count = 0
for event in si:
for value in event.summary.value:
count += 1
proto_bytes = value.metadata.plugin_data.content
if len(proto_bytes) > 0:
print('Found event!')
plugin_data = plugin_data_pb2.HParamsPluginData.FromString(proto_bytes)
print(plugin_data)
if plugin_data.HasField("experiment"):
print(
"Got experiment metadata with %d hparams and %d metrics"
% (
len(plugin_data.experiment.hparam_infos),
len(plugin_data.experiment.metric_infos),
),
)
elif plugin_data.HasField("session_start_info"):
print(
"Got session start info with concrete hparam values: %r"
% (dict(plugin_data.session_start_info.hparams),)
)
# -
event
# +
import tensorflow as tf
from tensorflow.python.summary.summary_iterator import summary_iterator
from tensorboard.plugins.hparams import plugin_data_pb2
si = summary_iterator(
"../runs/Jan06_16-36-46_DESKTOP-I8HN3PBshallownet_symmetric_repeat/events.out.tfevents.1609969007.DESKTOP-I8HN3PB.11783.0"
)
count = 0
for event in si:
for value in event.summary.value:
if value.tag == 'Tune/corr/mean':
print(value)
# +
import torch
filename = '../models/shallownet_symmetric_repeat/model.ckpt-1120000-2021-01-07 04-20-03.771753.pt'
data = torch.load(filename)
# -
import matplotlib.pyplot as plt
import numpy as np
_ = plt.plot((np.arange(-8, 1)) / 30, data['wt'].cpu().detach().numpy())
plt.xlabel('delay')
| scripts/retired/Read tensorboard hparams.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### **Packing and Unpacking**
# We have list, tuple, dictionary, and set which can store multiple elements in Python.
# > Packing refers to collecting several values into a single variable.
# As a complement, Unpacking refers to an operation that consists of assigning an iterable values to a tuple.
# #### **Tuple Packing and Unpacking**
# * Unpacking of values into varaibles.
a,b,c = 1,2,3
a,b,c
# > When we put tuples on both sides of an assignment operator, a tuple unpacking operation takes place.
# > The number of variables and values must match. Else we will get `ValueError`
a,b,c = 1,2
# Unpacking a string
a,b,c = "123"
a,b,c
# Unpacking list
a,b,c = [1,2,3]
a,b,c
# range
a,b,c = range(3)
a,b,c
# -----------
#
# #### *** Operator**
# > The `*` operator is known, in this context as tuple unpacking operator.
# > It extends unpacking functionality to allow us to collect or pack multiple values in a single variable.
# Packing
*a, = 1,2,3,4
a
# Packing the trailing values in b. This can be interchanged.
# a should get value. its mandatory
a, *b = 1,2,3,4
a,b
# Packing no values in a (a defaults to []) because b, c are mandatory
*a, b, c = 1,2
a,b,c
# * We can't use the unpacking operator`*` to pack multiple values into one variable without adding a trailing comma`,` to the variable on the left side of the assignment.
# * `,` makes an assignment to an iterable target list, in which a starred target is valid syntax.
*a = range(10)
*a, = range(10)
a
# * We cannot use more than one * in an assignment.
*a, *b, = range(10)
# ----------
# #### **Ignoring unwanted values**
#
# `_` is a dummy variable.
a,b,*_ = 2,3,10,40
a,b,_
print(_)
# * **Returning multiple values from a Function** : We can return several values from a function seperated by commas.
# +
def GetNumbers(n):
return n+1,n+2,n+3
print(GetNumbers(2))
# -
# #### **Dictionary Packing and Unpacking**
# > `**` is called Dictionary unpacking operator.
my_dict = {"Name":"Gagana","Subject":"Python"}
Marks = {"Python":95,"Math": 90}
# > Merging multiple dictionary into one final dictionary.
Combine = {**my_dict,**Marks}
Combine
# > If the dictionaries we're trying to merge have common keys, then the values of the right-most dictionary will override the values of the left-most dictionary
# #### **Unpacking in `for` Loops**
Students = [("Sam",10),("James",20),("Justin",60)]
# This is using indexing
for student in Students:
print("Student Name : {0}, Marks scored : {1} ".format(student[0],student[1]))
# Unpacking
for name, marks in Students:
print("Student Name : {0}, Marks scored : {1} ".format(name,marks))
# #### **Passing Multiple values to a function** :
# In Functions chapter we have come accross Arbitrary argument passing using tuple (*)
# **Q**. How a dictionary is passed ?
# +
def Report(**kwargs):
print(type(kwargs))
for key, value in kwargs.items():
print("Student Name : {0}, Marks scored : {1} ".format(key,value))
Report(Sam = 10,James = 20,Justin = 60)
# -
# ----
#
# **Q**. When we try to pass duplicate keys?
# +
def Report(**kwargs):
print(type(kwargs))
for key, value in kwargs.items():
print("Student Name : {0}, Marks scored : {1} ".format(key,value))
Report(Sam = 10,James = 20,Justin = 60, Sam = 20)
# -
# --------------
| Basics/PackingUnpacking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.8 64-bit (conda)
# name: python388jvsc74a57bd083ad9dc287f1bd68e1373a062e5fec25449c786b53be0804b995b765c2d61fc9
# ---
# ----
# # Introduction to Data Science and Systems
#
# ## Self-study: Arrays, `numpy` and vectorisation
#
#
# ##### University of Glasgow - material prepared by <NAME>* (adapted to IDSS by BSJ).
#
#
# $$\newcommand{\vec}[1]{{\bf #1}}
# \newcommand{\real}{\mathbb{R}}
# \newcommand{\expect}[1]{\mathbb{E}[#1]}
# \DeclareMathOperator*{\argmin}{arg\,min}
# %\vec{x}
# %\real
# $$
#
# ----
# # Content:
#
#
#
# ## [1: Why use arrays](idss_selfstudy_numerical_i_ch_1.ipynb)
# * what vectorized computation is
# * what numerical arrays are and what they are useful for
# * the general categories of array operations
# * how images and sounds map onto arrays
#
# ## [2: Typing and shapes of arrays](idss_selfstudy_numerical_i_ch_2.ipynb)
# * the naming of different types of arrays (vector, matrix, tensor)
# * what shape and dtype are
# * what axes of an array are and how they are named (row, column, etc.)
#
# ## [3: Creating, indexing, slicing, joining and rotating](idss_selfstudy_numerical_i_ch_3.ipynb)
# * creating new arrays
# * slicing and indexing operations and their syntax
# * how to rotate, flip and transpose arrays
# * how to split and join arrays and the rules governing this
# * boolean arrays and fancy indexing
# * swapping, adding dimensions, reshaping and adding dimensions
#
# ## [4: Arithmetic, broadcasting and aggregation](idss_selfstudy_numerical_i_ch_4.ipynb)
# * scalar and elementwise arithmetic on arrays
# * broadcasting rules
# * basic aggregation operations like summation, mean, cumulative sum
# * sorting and selection like argmax, argsort, find
#
# ## [5: Nummerical aspects](idss_selfstudy_numerical_i_ch_5.ipynb)
# * how IEEE 754 `float32` and `float64` numbers are represented
# * how infinity and NaN are represented, how they occur and how they are used
# * what roundoff error is and how it tends to be caused
# * how to compare floating point numbers
# * what machine epsilon is and how it is defined
#
#
# ## [6: Vectorisation](idss_selfstudy_numerical_i_ch_6.ipynb)
# * how to vectorise basic algorithms like summations and elementwise operations
# * how to mask elements in vectorised operations
# * how to write simple equations as vectorised operations
#
#
# ## [Appendix: Numpy Reference](idss_selfstudy_numerical_i_ch_numpyreference.ipynb)
# - a list of highly releveant `numpy` functions/features
#
# ---
# ## Extra resources for this self-study:
# * [From Python to Numpy](http://www.labri.fr/perso/nrougier/from-python-to-numpy/)
# * [100 numpy exercises](http://www.labri.fr/perso/nrougier/teaching/numpy.100/index.html)
# * [NumPy tutorial](http://scipy.github.io/old-wiki/pages/Tentative_NumPy_Tutorial)
# * [Introduction to NumPy](https://jakevdp.github.io/PythonDataScienceHandbook/02.00-introduction-to-numpy.html)
# * [Linear algebra cheat sheet](https://medium.com/towards-data-science/linear-algebra-cheat-sheet-for-deep-learning-cd67aba4526c#.739w4i3m1) *not actually linear algebra!*
| Self-Study/idss_selfstudy_numerical_i_ch_0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from statistics import mean
import numpy as np
import matplotlib.pyplot as plt
xs = np.array([1, 2, 3, 4, 5, 6], dtype=np.float64)
ys = np.array([5, 4, 6, 5, 6, 7], dtype=np.float64)
plt.scatter(xs,ys)
plt.show()
# -
def best_fit_slope(xs, ys):
numerator = np.mean(xs) * np.mean(ys) - np.mean(xs * ys)
denominator = np.mean(xs)**2 - np.mean(xs**2)
return numerator / denominator
# +
m = best_fit_slope(xs, ys)
print(m)
# -
def best_fit_intercept(xs, ys, m):
return np.mean(ys) - m * np.mean(xs)
# +
b = best_fit_intercept(xs, ys, m)
print(b)
# +
regression_line = [(m * x) + b for x in xs]
from matplotlib import style
style.use('fivethirtyeight')
# -
predict_x = 8
predict_y = (m * predict_x) + b
plt.scatter(xs, ys)
plt.scatter(predict_x, predict_y)
plt.plot(xs, regression_line)
plt.show()
# +
# Calculate how good our fit line is with squared error
# +
def squared_error(ys_original, ys_line):
return sum((ys_line - ys_original)**2)
def coefficient_of_determination(ys_original, ys_line):
ys_mean_line = [np.mean(ys_original) for y in ys_original]
squared_err_regr = squared_error(ys_original, ys_line)
squared_err_y_mean = squared_error(ys_original, ys_mean_line)
return 1 - (squared_err_regr) / (squared_err_y_mean)
# -
r_squared = coefficient_of_determination(ys, regression_line)
print(r_squared)
print([np.mean(ys) for y in ys])
# +
# testing
import random
import numpy as np
# size tells how many points to generate
###
# variance determines how scattered our data becomes
# it could make our dataset non-linear if set too low
###
# step is how much to add to our value
# corelation is either positive or negative
def create_dataset(size, variance, step = 2, correlation = False):
val = 1
ys = []
for i in range(size):
y = val + random.randrange(-variance, variance)
ys.append(y)
if correlation == 'pos':
val += step
elif correlation == 'neg':
val -= step
xs = [i for i in range(len(ys))]
# return xs and ys
return np.array(xs, dtype=np.float64), np.array(ys, dtype=np.float64)
# -
xs, ys = create_dataset(40, 40, 2, correlation='pos')
# +
m = best_fit_slope(xs, ys)
b = best_fit_intercept(xs, ys, m)
regression_line = [(m * x) + b for x in xs]
predict_x = 8
predict_y = (m * predict_x) + b
r_squared = coefficient_of_determination(ys, regression_line)
print(r_squared)
# -
plt.scatter(xs, ys)
plt.scatter(predict_x, predict_y)
plt.plot(xs, regression_line)
plt.show()
| machine-learning-python/Regression_self-made.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GradCAM: Continuation (Part 2) - Lecture Notebook
# In the previous lecture notebook (GradCAM Part 1) we explored what Grad-Cam is and why it is useful. We also looked at how we can compute the activations of a particular layer using Keras API. In this notebook we will check the other element that Grad-CAM requires, the gradients of the model's output with respect to our desired layer's output. This is the "Grad" portion of Grad-CAM.
#
# Let's dive into it!
import keras
from keras import backend as K
from util import *
# The `load_C3M3_model()` function has been taken care of and as last time, its internals are out of the scope of this notebook.
# Load the model we used last time
model = load_C3M3_model()
# Kindly recall from the previous notebook (GradCAM Part 1) that our model has 428 layers.
# We are now interested in getting the gradients when the model outputs a specific class. For this we will use Keras backend's `gradients(..)` function. This function requires two arguments:
#
# - Loss (scalar tensor)
# - List of variables
#
# Since we want the gradients with respect to the output, we can use our model's output tensor:
# +
# Save model's output in a variable
y = model.output
# Print model's output
y
# -
# However this is not a scalar (aka rank-0) tensor because it has axes. To transform this tensor into a scalar we can slice it like this:
y = y[0]
y
# It is still *not* a scalar tensor so we will have to slice it again:
y = y[0]
y
# Now it is a scalar tensor!
#
# The above slicing could be done in a single statement like this:
#
# ```python
# y = y[0,0]
# ```
#
# But the explicit version of it was shown for visibility purposes.
#
# The first argument required by `gradients(..)` function is the loss, which we will like to get the gradient of, and the second is a list of parameters to compute the gradient with respect to. Since we are interested in getting the gradient of the output of the model with respect to the output of the last convolutional layer we need to specify the layer as we did in the previous notebook:
# +
# Save the desired layer in a variable
layer = model.get_layer("conv5_block16_concat")
# Compute gradient of model's output with respect to last conv layer's output
gradients = K.gradients(y, layer.output)
# Print gradients list
gradients
# -
# Notice that the gradients function returns a list of placeholder tensors. To get the actual placeholder we will get the first element of this list:
# +
# Get first (and only) element in the list
gradients = gradients[0]
# Print tensor placeholder
gradients
# -
# As with the activations of the last convolutional layer in the previous notebook, we still need a function that uses this placeholder to compute the actual values for an input image. This can be done in the same manner as before. Remember this **function expects its arguments as lists or tuples**:
# +
# Instantiate the function to compute the gradients
gradients_function = K.function([model.input], [gradients])
# Print the gradients function
gradients_function
# -
# Now that we have the function for computing the gradients, let's test it out on a particular image. Don't worry about the code to load the image, this has been taken care of for you, you should only care that an image ready to be processed will be saved in the x variable:
# +
# Load dataframe that contains information about the dataset of images
df = pd.read_csv("nih_new/train-small.csv")
# Path to the actual image
im_path = 'nih_new/images-small/00000599_000.png'
# Load the image and save it to a variable
x = load_image(im_path, df, preprocess=False)
# Display the image
plt.imshow(x, cmap = 'gray')
plt.show()
# -
# We should normalize this image before going forward, this has also been taken care of:
# +
# Calculate mean and standard deviation of a batch of images
mean, std = get_mean_std_per_batch(df)
# Normalize image
x = load_image_normalize(im_path, mean, std)
# -
# Now we have everything we need to compute the actual values of the gradients. In this case we should also **provide the input as a list or tuple**:
# Run the function on the image and save it in a variable
actual_gradients = gradients_function([x])
# An important intermediary step is to trim the batch dimension which can be done like this:
# Remove batch dimension
actual_gradients = actual_gradients[0][0, :]
# +
# Print shape of the gradients array
print(f"Gradients of model's output with respect to output of last convolutional layer have shape: {actual_gradients.shape}")
# Print gradients array
actual_gradients
# -
# Looks like everything worked out nicely! You will still have to wait for the assignment to see how these elements are used by Grad-CAM to get visual interpretations. Before you go you should know that there is a shortcut for these calculations by getting both elements from a single Keras function:
# +
# Save multi-input Keras function in a variable
activations_and_gradients_function = K.function([model.input], [layer.output, gradients])
# Run the function on our image
act_x, grad_x = activations_and_gradients_function([x])
# Remove batch dimension for both arrays
act_x = act_x[0, :]
grad_x = grad_x[0, :]
# +
# Print actual activations
print(act_x)
# Print actual gradients
print(grad_x)
# -
# **Congratulations on finishing this lecture notebook!** Hopefully you will now have a better understanding of how to leverage Keras's API power for computing gradients. Keep it up!
| andrew_ng/machine_learning/medicine/concepts/course_3/week_3/AI4M_C3_M3_lecture_notebook_gradcam_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### 14. World Coordinates Explanation
# This notebook shows to importance of using the world coordinates of images when registering them.
#
import itk
from itkwidgets import compare, checkerboard, view
import numpy as np
# +
# Import Images with world coordinates
fixed_image = itk.imread('data/CT_3D_lung_fixed.mha', itk.F)
moving_image = itk.imread('data/CT_3D_lung_moving.mha', itk.F)
# Recast Image to numpy, then to itk to replace original world coordinates with itk default once.
fixed_image_np = np.asarray(fixed_image).astype(np.float32)
fixed_image_np = itk.image_view_from_array(fixed_image_np)
moving_image_np = np.asarray(moving_image).astype(np.float32)
moving_image_np = itk.image_view_from_array(moving_image_np)
# Registration with original itk image
result_image, result_transform_parameters = itk.elastix_registration_method(
fixed_image, moving_image)
# Registration with recasted numpy image with default world coordinates.
result_image_np, result_transform_parameters = itk.elastix_registration_method(
fixed_image_np, moving_image_np)
# -
# Compare result images with itk widgets, images do not occupy same fysical space.
compare(result_image, result_image_np)
# Set origin and spacing equal for pixel-wise image comparison
result_image_no_wc = np.asarray(result_image).astype(np.float32)
result_image_np_no_wc = np.asarray(result_image_np).astype(np.float32)
# Compare result images, images now occupy same fysical space, but still have (smaller) differences.
compare(result_image_no_wc, result_image_np_no_wc)
| examples/ITK_Example14_WorldCoordinatesExplanation_Napari.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # K-Nearest Neighbor
# *Complete and hand in this completed worksheet (including its outputs and any supporting code outside of the worksheet) with your assignment submission. Please check the pdf file for more details.*
#
# In this exercise you will:
#
# - implement the **K-Nearest Neighbor** algorithm
# - play with the hyperparameter K
# - try KNN on **real-world data**, i.e. the CAPTCHA of a website in ZJU
#
# Please note that **YOU CANNOT USE ANY MACHINE LEARNING PACKAGE SUCH AS SKLEARN** for any homework, unless you are asked.
# +
# some basic imports
import scipy.io as sio
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# # %matplotlib notebook
# %load_ext autoreload
# %autoreload 2
# -
# ### KNN experiment with hyperparameter K
# +
from mkdata import mkdata
from knn_plot import knn_plot
[X, y] = mkdata()
K = [1, 10, 100]
for k in K:
knn_plot(X, y, k)
# -
# ### Now Let's hack the website http://jwbinfosys.zju.edu.cn/default2.aspx
from hack import hack
from extract_image import extract_image
from show_image import show_image
import urllib.request
test_img = './CheckCode.aspx' # change it yourself
urllib.request.urlretrieve("http://jwbinfosys.zju.edu.cn/CheckCode.aspx", 'data/tmp.jpg')
hack("data/tmp.jpg")
| hw3/knn/knn_exp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# Some functions are from or based on work done by <NAME> in the below notebook
# https://github.com/KevinLiao159/MyDataSciencePortfolio/blob/master/movie_recommender/movie_recommendation_using_ALS.ipynb
# Intialization
import os
import time
import warnings
warnings.filterwarnings("ignore", message="numpy.dtype size changed")
# spark imports
from pyspark.sql import SparkSession
from pyspark.sql.functions import UserDefinedFunction, explode, desc
from pyspark.sql.types import StringType, ArrayType
from pyspark.ml.recommendation import ALS
from pyspark.ml.linalg import Vectors
from pyspark.ml.feature import VectorAssembler
from pyspark.ml.regression import GBTRegressor, GBTRegressionModel
# data science imports
import numpy as np
import pandas as pd
from sklearn.preprocessing import MultiLabelBinarizer
# -
data_path = 'hdfs:///user/andrew/'
# +
# %%time
# Read in data through spark since the data is sored in hadoop and format the columns
# Convert to pandas dataframes for easier and faster manipulation
from pyspark.sql.types import *
from pyspark.sql import SQLContext, Row
from pyspark.sql.functions import *
sqlContext = SQLContext(sc)
movies = sqlContext.read.parquet(data_path + 'movie_20m_metadata_OHE_subset')
movies_df = movies.toPandas()
movies_df = movies_df.set_index(movies_df.item_id) # set index so no sorting errors occur
# movies_gp = sqlContext.read.parquet('hdfs:///user/andrew/movie_genre_and_people_metadata_ohe_subset')
movies_gp = movies.drop('title', 'imdb_id', 'imdb_rating', 'imdb_votes', 'metascore', 'runtime', 'year')
ratings = sqlContext.read.parquet(data_path + 'ratings_20m')
ratings = ratings.drop('timestamp')
ratings = ratings.withColumn("userId", ratings["userId"].cast("int"))
ratings = ratings.withColumn("rating", ratings["rating"] * 2) #Multiply by 2 so that values are whole numbers -> values 1 to 10
ratings = ratings.select('userId', 'movieId', 'rating').toDF('user_id', 'item_id', 'label')
# -
# User input function - takes user input data, strpis it down, and calls other functions on that data
# Takes in user age, gender, occupation (of 20 options - may drop this), list of favorite movies
# All movies in the list of favorite movies will be rated 5 stars
def new_user_input(fav_movies, all_ratings, movies, spark_context,
sqlContext = None, num_recs = 10, movies_gp = None,
movies_df = None):
# age should be an integer in 1 - 100
# gender should be M or F
# fav_movies should be in the form of ["Iron Man", "The Shawshank Redemption", "<NAME>"]
# If there are multiple versions of the movie and the user wishes for one other than the most recent one, they
# should specify with a year in parenthesis, like "<NAME> (1993)"
# Occupation...
# collect favorite movie ids
print 'Collecting favorite movie IDs'
movieIds = get_movieId(movies_df, fav_movies)
print 'Favorite movies in the available set'
print movies_df[['item_id', 'title', 'year']].loc[movieIds]
print 'Adding ratings to full set'
# add new user movie ratings to all ratings dataframe
# all_ratings_updated, user_ratings, user_ratings_binary = add_new_user_to_data(all_ratings, movieIds, spark_context)
all_ratings_updated, new_user_ratings = add_new_user_to_data(all_ratings, movieIds, spark_context)
del all_ratings
print 'Creating prediction set'
# get all unrated movies for user (unnecessary in Spark 2.2+, instead use the recommendForAllUsers(num_to_rec) method)
all_user_unrated = get_inference_data(all_ratings_updated, movieIds)
print 'Training ALS model'
# train ALS model, then predict movie ratings
als = ALS(seed = 42, regParam = 0.1, maxIter = 15, rank = 12, # coldStartStrategy = 'drop', # drops userIds/movieIds from the validation set or test set so that NaNs are not returned
userCol = "user_id", itemCol = "item_id", ratingCol = "label")
als_model = als.fit(all_ratings_updated)
del all_ratings_updated
print 'Making Predictions'
# keep top 30 predictions
full_predictions_sorted = als_model.transform(all_user_unrated).sort(desc('prediction'))
als_top_n_predictions = full_predictions_sorted.take(num_recs)
# extract movie ids
als_top_n_ids = [r[1] for r in als_top_n_predictions]
als_movie_recs = movies.filter(movies.item_id.isin(als_top_n_ids)).select('title', 'year')
print ''
print 'ALS Recommendations'
print als_movie_recs.toPandas()
# format data for prediction using GBTs
# create user_id x item_id matrix need to get data in the form of user_id, item_id, label, then pivot
# filter movies_gp dataframe by the movieIds. pivot new_user_ratings into a vector,
# then multiply by the filtered movies_gp dataframe; divide by binarized user ratings;
# this should now be a vector of user preferences.
# join a OHE age, gender, and possibly occupation, to the user preferences
user_summary = get_user_preferences(user_ratings = new_user_ratings, movieIds = movieIds,
movies_gp = movies_gp, sqlContext = sqlContext)
# Extract movie ids from the top 5*num_recs for Gradient Boosted Trees prediction
als_top_3xn_predictions = full_predictions_sorted.take(3*num_recs)
als_top_3xn_ids = [r[1] for r in als_top_3xn_predictions]
all_user_unrated_top_3xn = all_user_unrated.filter(all_user_unrated.item_id.isin(als_top_3xn_ids))
top_3xn_movies_metadata = movies.filter(movies.item_id.isin(als_top_3xn_ids))
# lastly, replicate the user pref rows for each rated movieId, then join with the filtered movies dataframe
# (MAKE SURE ALL COLUMNS ARE ORDERED AND NAMED CORRECTLY)
unrated_with_movie_metadata = all_user_unrated_top_3xn \
.join(top_3xn_movies_metadata, on = 'item_id', how = 'left')
unrated_with_full_metadata = unrated_with_movie_metadata \
.join(user_summary, on = 'user_id', how = 'left') \
.drop('user_id', 'title', 'imdb_id')
# the GBT model takes in the rows as vectors, so the columns must be converted to the feature space
unrated_with_full_metadata_rdd = unrated_with_full_metadata.rdd.map(lambda x: (x[0], Vectors.dense(x[1:])))
unrated_metadata_features = sqlContext.createDataFrame(unrated_with_full_metadata_rdd, schema = ['item_id', 'features'])
# import the GBT model, in this case a GBTRegressionModel with tree depth of 10
GBTRegD10Model = GBTRegressionModel.load(data_path + 'GBTRegD10Model_20m')
# use pre-trained GBT model to predict movie ratings
gbtr_preds = GBTRegD10Model.transform(unrated_metadata_features)
# sort by predicted rating, and keep top recommend top n
gbtr_top_n_predictions = gbtr_preds.sort(desc('prediction')).take(num_recs)
# extract movie ids
gbtr_top_n_ids = [r[0] for r in gbtr_top_n_predictions]
gbtr_movie_recs = movies.filter(movies.item_id.isin(gbtr_top_n_ids)).select('title', 'year')
print ''
print 'GBTR Recommendations'
print gbtr_movie_recs.toPandas()
def get_movieId(movies_df, fav_movie_list):
"""
return all movieId(s) of user's favorite movies
Parameters
----------
df_movies: spark Dataframe, movies data
fav_movie_list: list, user's list of favorite movies
Return
------
movieId_list: list of movieId(s)
"""
movieId_list = []
for movie in fav_movie_list:
if movie[0:4] == 'The ':
movie = movie[4:]
elif movie[0:3] == 'An ':
movie = movie[3:]
elif movie[0:3] == 'La ':
movie = movie[3:]
elif movie[0:2] == 'A ':
movie = movie[3:]
if movie[-6:-5] == '(':
year = int(movie[-5:-1])
movie = movie[0:-7]
movieIds = movies_df.item_id[(movies_df.title.str.contains(movie)) & (movies_df.year == year)]
movieId_list.extend(movieIds)
elif len(movie.split(' ')) == 1:
movieIds = movies_df.item_id[movies_df.title == movie]
movieId_list.extend(movieIds)
else:
movieIds = movies_df.item_id[movies_df.title.str.contains(movie)]
movieId_list.extend(movieIds)
return movieId_list
def add_new_user_to_data(train_data, movieIds, spark_context):
"""
add new rows with new user, user's movie and ratings to
existing train data
Parameters
----------
train_data: Spark DataFrame, ratings data
movieIds: spark DataFrame, single column of movieId(s)
spark_context: Spark Context object
Return
------
new train data with the new user's rows
"""
# get new user id
new_id = train_data.agg({"user_id": "max"}).collect()[0][0] + 1
# get max rating
max_rating = train_data.agg({"label": "max"}).collect()[0][0]
# create new user sdf for max rating
user_rows_max = [(new_id, movieId, max_rating) for movieId in movieIds]
new_sdf_max = spark_context.parallelize(user_rows_max).toDF(['user_id', 'item_id', 'label'])
# return new train data
return train_data.union(new_sdf_max), new_sdf_max # , new_sdf_binary
def get_inference_data(train_data, movieIds):
"""
return a rdd with the userid and all movies (except ones in movieId_list)
Parameters
----------
train_data: spark RDD, ratings data
df_movies: spark Dataframe, movies data
movieId_list: list, list of movieId(s)
Return
------
inference data: Spark RDD
"""
# get new user id
new_id = train_data.agg({"user_id": "max"}).collect()[0][0]
distinct_unrated_items = ratings.select('item_id').distinct().filter(~col('item_id').isin(movieIds))
user_unrated = distinct_unrated_items.withColumn('user_id', lit(new_id)).select('user_id', 'item_id')
return user_unrated
def get_user_preferences(user_ratings, movieIds, movies_gp, sqlContext):
#new_user_ratings
# pivoted_user_ratings = user_ratings.groupBy('user_id').pivot('item_id').agg(avg('label'))
# pivoted_new_user_ratings_binary = user_ratings_binary.groupBy('user_id').pivot('item_id').agg(avg('label')).drop('user_id')
pivoted_user_ratings_df = user_ratings.toPandas() \
.pivot(index='user_id',
columns='item_id',
values='label') \
.fillna(0)
pivoted_user_ratings_df_binary = pivoted_user_ratings_df / pivoted_user_ratings_df
movies_gp_filtered = movies_gp.filter(col('item_id').isin(movieIds))
movies_gp_filtered_df = movies_gp_filtered.toPandas()
# movies_gp_filtered_df.item_id = movies_gp_filtered_df.item_id.astype(str) only necessary when pivot was done on spark df
movies_gp_filtered_df = movies_gp_filtered_df.set_index('item_id')
user_summary_total = pivoted_user_ratings_df.dot(movies_gp_filtered_df)
user_summary_count = pivoted_user_ratings_df_binary.dot(movies_gp_filtered_df)
user_summary_avg = (user_summary_total / user_summary_count).fillna(0)
user_summary_avg = user_summary_avg.add_suffix('_avg_rating').reset_index()
sorted_columns = list(user_summary_avg.columns.sort_values())
user_summary_sdf = sqlContext.createDataFrame(user_summary_avg[sorted_columns])
return user_summary_sdf
# ### Step by Step Walkthrough of Main Function (to show runtime)
# %%time
fav_movies = ['Tinker Tailor Soldier Spy', 'Shawshank Redemption', 'Lord of the Rings']
# collect favorite movie ids
print 'Collecting favorite movie IDs'
movieIds = get_movieId(movies_df, fav_movies)
if movies_df is not None:
print 'Favorite movies in the available set'
print movies_df[['item_id', 'title', 'year']].loc[movieIds]
# %%time
print 'Adding ratings to full set'
# add new user movie ratings to all ratings dataframe
# all_ratings_updated, user_ratings, user_ratings_binary = add_new_user_to_data(all_ratings, movieIds, spark_context)
all_ratings_updated, user_ratings = add_new_user_to_data(ratings, movieIds, sc)
# %%time
print 'Creating prediction set'
# get all unrated movies for user (unnecessary in Spark 2.2+, instead use the recommendForAllUsers(num_to_rec) method)
all_user_unrated = get_inference_data(all_ratings_updated, movieIds)
# %%time
print 'Training ALS model'
# train ALS model, then predict movie ratings
als = ALS(seed = 42, regParam = 0.1, maxIter = 15, rank = 12,
userCol = "user_id", itemCol = "item_id", ratingCol = "label")
als_model = als.fit(all_ratings_updated)
del all_ratings_updated
# +
# %%time
print 'Making Predictions'
# keep top 15 predictions
num_recs = 15
full_predictions = als_model.transform(all_user_unrated)
als_top_n_predictions = full_predictions.sort(desc('prediction')).take(num_recs)
# extract movie ids
als_top_n_ids = [r[1] for r in als_top_n_predictions]
als_movie_recs = movies.filter(movies.item_id.isin(als_top_n_ids)).select('title', 'year')
print 'ALS Recommendations'
print als_movie_recs.toPandas()
# -
# %%time
# import GBT model input data format
# Create user_id x item_id matrix need to get data in the form of user_id, item_id, label, then pivot
# filter movies_gp dataframe by the movieIds. pivot new_user_ratings into a vector,
# then multiply by the filtered movies_gp dataframe; divide by binarized user ratings;
# this should now be a vector of user preferences.
# join a OHE age, gender, and possibly occupation, to the user preferences
user_summary_sdf = get_user_preferences(user_ratings = user_ratings, movieIds = movieIds,
movies_gp = movies_gp, sqlContext = sqlContext)
# %%time
als_top_3xn_predictions = full_predictions.sort(desc('prediction')).take(3*num_recs)
als_top_3xn_ids = [r[1] for r in als_top_3xn_predictions]
all_user_unrated_top_3xn = all_user_unrated.filter(all_user_unrated.item_id.isin(als_top_3xn_ids)) #looks good
top_3xn_movies_metadata = movies.filter(movies.item_id.isin(als_top_3xn_ids)) #looks good
# %%time
# lastly, replicate the user pref rows for each rated movieId, then join with the filtered movies dataframe
# (MAKE SURE ALL COLUMNS ARE ORDERED AND NAMED CORRECTLY)
unrated_with_movie_metadata = all_user_unrated_top_3xn \
.join(top_3xn_movies_metadata, on = 'item_id', how = 'left') #looks good
unrated_with_full_metadata = unrated_with_movie_metadata \
.join(user_summary_sdf, on = 'user_id', how = 'left') \
.drop('user_id', 'title', 'imdb_id') #looks good
# +
# %%time
# convert predictors to "features" and it is ready for prediction.
# features_cols = list(unrated_with_full_metadata.columns)
# features_cols.remove('item_id')
# vecAssembler = VectorAssembler(inputCols = features_cols, outputCol="features")
# unrated_metadata_features = vecAssembler.transform(unrated_with_full_metadata)
unrated_with_full_metadata_rdd = unrated_with_full_metadata.rdd.map(lambda x: (x[0], Vectors.dense(x[1:])))
unrated_metadata_features = sqlContext.createDataFrame(unrated_with_full_metadata_rdd, schema = ['item_id', 'features'])
#looks good
# -
# %%time
GBTRegD10Model = GBTRegressionModel.load(data_path + 'GBTRegD10Model_20m')
gbtr_preds = GBTRegD10Model.transform(unrated_metadata_features)
# +
# %%time
gbtr_top_n_predictions = gbtr_preds.sort(desc('prediction')).take(num_recs)
# extract movie ids
gbtr_top_n_ids = [r[0] for r in gbtr_top_n_predictions]
gbtr_movie_recs = movies.filter(movies.item_id.isin(gbtr_top_n_ids)).select('title', 'year')
print 'GBTR Recommendations'
print gbtr_movie_recs.toPandas()
# -
# ### Full Function Recommendation Examples
# %%time
fav_movies = ['Iron Man', 'Tinker Tailor Soldier Spy', 'Shawshank Redemption', 'Lord of the Rings (2002)', '<NAME>',
'The Family Stone', 'Shaun of the Dead', 'Up', 'A View to a Kill']
new_user_input(fav_movies = fav_movies, all_ratings = ratings,
movies = movies, spark_context = sc, sqlContext = sqlContext,
num_recs = 10, movies_gp = movies_gp, movies_df = movies_df)
# %%time
fav_movies = ['Tinker Tailor Soldier Spy', 'Shawshank Redemption', 'Lord of the Rings']
new_user_input(fav_movies = fav_movies, all_ratings = ratings,
movies = movies, spark_context = sc, sqlContext = sqlContext,
num_recs = 10, movies_gp = movies_gp, movies_df = movies_df)
# %%time
fav_movies = ['Frozen', 'Tangled', 'Oceans Eleven', 'Toy Story', 'The Princess Bride',
'The Incredibles', 'Castle in the Sky', 'Monsters, Inc']
new_user_input(fav_movies = fav_movies, all_ratings = ratings,
movies = movies, spark_context = sc, sqlContext = sqlContext,
num_recs = 10, movies_gp = movies_gp, movies_df = movies_df)
# %%time
fav_movies = ['The Sound of Music', 'Blackhawk Down', 'Pearl Harbor', 'Toy Story', 'The Princess Bride',
'Foreign Student', 'Star Wars', 'The Shining', 'Rear Window', 'Groundhog Day', 'Ghostbusters',
'Robin Hood (1993)', 'Die Hard']
new_user_input(fav_movies = fav_movies, all_ratings = ratings,
movies = movies, spark_context = sc, sqlContext = sqlContext,
num_recs = 10, movies_gp = movies_gp, movies_df = movies_df)
| Recommendation_Comparison/Mini_Movie_Recommender_ALS_plus_Trees_20m.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
cov_df = pd.read_csv('../cerebra/coverageBatch.csv')
cov_df
for i in range(0, len(cov_df.index)):
currRow = cov_df.iloc[i]
chrom_ = currRow['chrom']
start_ = currRow['start_pos']
end_ = currRow['end_pos']
outfile_ = currRow['outfile']
currRow
| py_notebooks/scratch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Script
# - **Input:** Airbnb - listings, reviews, single location.
# - **Output:** Creating entities and nodes to be loaded into Neo4j using both the listings and review data.
#
# - 5.5k listings | 250k reviews
# ### References
#
# - https://neo4j.com/developer/guide-import-csv/
# ### Import libraries, raw data
import pandas as pd
import os
import numpy as np
import warnings
warnings.filterwarnings('ignore')
# ls
root='./Data/raw/'
processor = './Data/processing/'
listings = pd.read_csv(root+'listings.csv.gz')
## These are the columns that will be used in the dialog generation.
listingTemplate = pd.read_csv(processor+'listing_Template.csv')
listing_processed = pd.read_csv(processor+'Processed_Airbnb/listings_text_processed.csv')
listings=listing_processed
listing_reviews = pd.read_csv(processor+'Processed_Airbnb/listing_with_reviews.csv')
listing_reviews = listing_reviews.set_index('listing_id').T.to_dict('list')
# ### Compute unique entities, create Neo4j nodes & edge map.
listings = listings[listingTemplate.columns]
listings.dropna()
listings.info()
len(listings)
### Add Geo-location, check more efficient way to add location.
listings['City'] = 'Amsterdam'
listings['Country'] = 'Netherlands'
listings[listings['neighbourhood_cleansed'].isna()]
listings['neighbourhood'] = listings['neighbourhood'].fillna(listings['neighbourhood_cleansed'])
listings['neighbourhood'] = listings['neighbourhood'].str.strip()
listings['State'] = listings['neighbourhood']
# #### Add all unique entities
# https://towardsdatascience.com/dealing-with-list-values-in-pandas-dataframes-a177e534f173
def to_1D(series):
return pd.Series([x for _list in series for x in _list])
listings["amenities"] = listings["amenities"].apply(eval)
amenities = to_1D(listings["amenities"]).unique()
# +
#[_id,_labels,name]
_id=[]
_labels=[]
name=[]
_start = []
_end = []
_rating = []
_relationship=[]
_listing_url=[]
_picture_url=[]
_host_identity_verified=[]
_accomodates=[]
_bedrooms=[]
_bathrooms=[]
_beds=[]
_price=[]
_review_scores_rating=[]
def convertList2Dict(lst,prev_index,label):
dict_ret = dict()
for index,value in enumerate(lst):
rec_index = index + prev_index
dict_ret[value] = int(rec_index)
##
_id.append(int(rec_index))
_labels.append(label)
name.append(value)
_start.append(np.NaN)
_end.append(np.NaN)
_relationship.append(np.NaN)
_rating.append(np.NaN)
_listing_url.append(np.NaN)
_picture_url.append(np.NaN)
_host_identity_verified.append(np.NaN)
_accomodates.append(np.NaN)
_bedrooms.append(np.NaN)
_bathrooms.append(np.NaN)
_beds.append(np.NaN)
_price.append(np.NaN)
_review_scores_rating.append(np.NaN)
prev_index = int(rec_index)
return dict_ret, prev_index
# -
City = listings["City"].dropna().unique()
State = listings["State"].dropna().unique()
Country = listings["Country"].dropna().unique()
Listing_Text = listings["Listing_Text"].dropna().unique()
Review_Text = listings["Review_Text"].dropna().unique()
# listings["price"].min()
# listings["price"].max()
# listings["price"] = listings["price"].replace('[\$,]', '', regex=True).astype(float)
# listings["price"] = pd.cut(listings['price'], [0, 250, 500,750,999], labels=['low', 'average', 'high','expensive'])
price = listings["price"].dropna().unique()
listings["review_scores_rating"].dropna()
listings["review_scores_rating"] = pd.cut(listings['review_scores_rating'], [0, 1, 2, 3,4,5], labels=['horrible', 'bad', 'average','good','very good'])
rating = listings["review_scores_rating"].unique()
listings["review_scores_accuracy"].dropna()
listings["review_scores_accuracy"] = pd.cut(listings['review_scores_accuracy'], [0, 1, 2, 3,4,5], labels=['horrible', 'bad', 'average','good','very good'])
acc_rating = listings["review_scores_accuracy"].unique()
listings["review_scores_cleanliness"].dropna()
listings["review_scores_cleanliness"] = pd.cut(listings['review_scores_cleanliness'], [0, 1, 2, 3,4,5], labels=['horrible', 'bad', 'average','good','very good'])
clean_rating = listings["review_scores_cleanliness"].unique()
listings["review_scores_checkin"].dropna()
listings["review_scores_checkin"] = pd.cut(listings['review_scores_checkin'], [0, 1, 2, 3,4,5], labels=['horrible', 'bad', 'average','good','smooth'])
checkin_rating = listings["review_scores_checkin"].unique()
listings["review_scores_communication"].dropna()
listings["review_scores_communication"] = pd.cut(listings['review_scores_communication'], [0, 1, 2, 3,4,5], labels=['horrible', 'bad', 'average','good','very good'])
communication_rating = listings["review_scores_communication"].unique()
listings["review_scores_location"].dropna()
listings["review_scores_location"] = pd.cut(listings['review_scores_location'], [0, 1, 2, 3,4,5], labels=['horrible', 'bad', 'average','good','very good'])
location_rating = listings["review_scores_location"].unique()
host_identity_verified = listings["host_identity_verified"].dropna().unique()
property_type = listings["property_type"].dropna().unique()
room_type = listings["room_type"].dropna().unique()
accommodates = listings["accommodates"].dropna().unique().astype(float)
bathrooms = listings["bathrooms_text"].dropna().unique()
bedrooms = listings["bedrooms"].dropna().unique()
beds = listings["beds"].dropna().unique()
accommodates=np.delete(accommodates,np.where(accommodates == 0))
amenities_dict, last_index = convertList2Dict(amenities,0,':Amenities')
City_dict, last_index = convertList2Dict(City,(last_index+1),':City')
State_dict, last_index = convertList2Dict(State,(last_index+1),':State')
Country_dict, last_index = convertList2Dict(Country,(last_index+1),':Country')
price_dict, last_index = convertList2Dict(price,(last_index+1),':price')
property_type_dict, last_index = convertList2Dict(property_type,(last_index+1),':property_type')
room_type_dict, last_index = convertList2Dict(room_type,(last_index+1),':room_type')
accommodates_dict, last_index = convertList2Dict(accommodates,(last_index+1),':accommodates')
bathrooms_dict, last_index = convertList2Dict(bathrooms,(last_index+1),':bathrooms')
bedrooms_dict, last_index = convertList2Dict(bedrooms,(last_index+1),':bedrooms')
beds_dict, last_index = convertList2Dict(beds,(last_index+1),':beds')
listingText_dict, last_index = convertList2Dict(Listing_Text,(last_index+1),':Listing_Text')
reviewText_dict, last_index = convertList2Dict(Review_Text,(last_index+1),':Review_Text')
import json
dictionary = {
"amenities":amenities_dict,
"City": City_dict,
"State":State_dict,
"Country":Country_dict,
"price":price_dict,
"property_type":property_type_dict,
"room_type":room_type_dict,
"accommodates":accommodates_dict,
"bathrooms":bathrooms_dict,
"bedrooms":bedrooms_dict,
"beds":beds_dict,
"Listing_Text":listingText_dict,
"Review_Text":reviewText_dict
}
with open(processor+'Processed_Airbnb/listings_entities_filter.json', 'w', encoding='utf-8') as f:
f.write(json.dumps(dictionary, ensure_ascii=False))
df = pd.read_json(processor+'Processed_Airbnb/listings_entities_filter.json')
df.to_csv(processor+'Processed_Airbnb/listings_entities_filter.csv')
with open(processor+'Processed_Airbnb/listings_info_filter.json', 'w', encoding='utf-8') as f:
f.write(listings.to_json(orient = 'records'))
reviews0 = pd.read_csv(processor+'Processed_Airbnb/ratings_filter.csv')
reviews0.info()
reviews=reviews0.dropna()
reviews = reviews[['listing_id','id','reviewer_id','comments','rating']]
# #### Add users
reviewer = reviews['reviewer_id'].unique()
for user in reviewer:
if user>1:
last_index = last_index+1
_id.append(last_index)
_labels.append(':User')
name.append(int(user))
_start.append(np.NaN)
_end.append(np.NaN)
_relationship.append(np.NaN)
_rating.append(np.NaN)
_listing_url.append(np.NaN)
_picture_url.append(np.NaN)
_host_identity_verified.append(np.NaN)
_accomodates.append(np.NaN)
_bedrooms.append(np.NaN)
_bathrooms.append(np.NaN)
_beds.append(np.NaN)
_price.append(np.NaN)
_review_scores_rating.append(np.NaN)
# #### Add listings
for index in listings.index:
last_index = last_index+1
_id.append(last_index)
_labels.append(':Listing')
name.append(listings['id'][index])
_start.append(np.NaN)
_end.append(np.NaN)
_relationship.append(np.NaN)
_rating.append(np.NaN)
_listing_url.append('https://www.airbnb.com/rooms/'+str(listings['id'][index]))
_picture_url.append(listings['picture_url'][index])
_host_identity_verified.append(listings['host_identity_verified'][index])
_accomodates.append(listings['accommodates'][index])
_bedrooms.append(listings['bedrooms'][index])
_bathrooms.append(listings['bathrooms_text'][index])
_beds.append(listings['beds'][index])
_price.append(listings['price'][index])
_review_scores_rating.append(listings['review_scores_rating'][index])
# +
#form a list of (u_id,l_id)
data = {'_id':_id,
'_labels':_labels,
'name':name,
'_start':_start,
'_end':_end,
'_type':_relationship,
'rating':_rating,
'url':_listing_url,
'picture_url':_picture_url,
'host_identity_verified':_host_identity_verified,
'accomodates':_accomodates,
'bedrooms':_bedrooms,
'bathrooms':_bathrooms,
'beds':_beds,
'price':_price,
'review_scores_rating':_review_scores_rating
}
#Create DataFrame
neo4J_format_df = pd.DataFrame(data)
neo4J_format_df.to_csv(processor+'Neo4j/neo4J_nodes.csv',index=False)
# -
# #### Create edge maps
# +
# Store listing ids
# Store user ids
neoNodes = pd.read_csv(processor+'Neo4j/neo4J_nodes.csv')
# +
neoListings = neoNodes[neoNodes['_labels']==':Listing']
neoAmenities = neoNodes[neoNodes['_labels']==':Amenities']
neoCity = neoNodes[neoNodes['_labels']==':City']
neoState = neoNodes[neoNodes['_labels']==':State']
neoCountry = neoNodes[neoNodes['_labels']==':Country']
neoProperty_type = neoNodes[neoNodes['_labels']==':property_type']
neoRoom_type = neoNodes[neoNodes['_labels']==':room_type']
neoUser = neoNodes[neoNodes['_labels']==':User']
neoListingText = neoNodes[neoNodes['_labels']==':Listing_Text']
neoReviewText = neoNodes[neoNodes['_labels']==':Review_Text']
print('Unique listings:'+str(len(neoListings)))
print('Unique Amenities:'+str(len(neoAmenities)))
print('Unique City:'+str(len(neoCity)))
print('Unique State:'+str(len(neoState)))
print('Unique Country:'+str(len(neoCountry)))
print('Unique property_type:'+str(len(neoProperty_type)))
print('Unique room_type:'+str(len(neoRoom_type)))
print('Unique User:'+str(len(neoUser)))
print('Unique listings:'+str(len(neoListingText)))
print('Unique listings:'+str(len(neoReviewText)))
# +
neoAmenities1 = neoAmenities[['_id','name','_labels']]
neoAmenities1['_labels'] = 'Amenity; Amenity'
neoAmenities1 = neoAmenities.rename(columns={"_id": "id:ID(Amenity)", "_labels": ":LABEL"})
neoAmenities1.to_csv(processor+"Neo4j/Amenity.csv",index=False)
neoCity1 = neoCity[['_id','name','_labels']]
neoCity1['_labels'] = 'City; City'
neoCity1 = neoCity1.rename(columns={"_id": "id:ID(City)", "_labels": ":LABEL"})
neoCity1.to_csv(processor+"Neo4j/City.csv",index=False)
neoState1 = neoState[['_id','name','_labels']]
neoState1['_labels'] = 'State; State'
neoState1 = neoState1.rename(columns={"_id": "id:ID(State)", "_labels": ":LABEL"})
neoState1.to_csv(processor+"Neo4j/State.csv",index=False)
neoCountry1 = neoCountry[['_id','name','_labels']]
neoCountry1['_labels'] = 'Country; Country'
neoCountry1 = neoCountry1.rename(columns={"_id": "id:ID(Country)", "_labels": ":LABEL"})
neoCountry1.to_csv(processor+"Neo4j/Country.csv",index=False)
neoProperty_type1 = neoProperty_type[['_id','name','_labels']]
neoProperty_type1['_labels'] = 'Property_type; Property_type'
neoProperty_type1 = neoProperty_type1.rename(columns={"_id": "id:ID(property_type)", "_labels": ":LABEL"})
neoProperty_type1.to_csv(processor+"Neo4j/Property_type.csv",index=False)
neoRoom_type1 = neoRoom_type[['_id','name','_labels']]
neoRoom_type1['_labels'] = 'Room_type; Room_type'
neoRoom_type1 = neoRoom_type1.rename(columns={"_id": "id:ID(room_type)", "_labels": ":LABEL"})
neoRoom_type1.to_csv(processor+"Neo4j/Room_type.csv",index=False)
neoListingText1 = neoListingText[['_id','name','_labels']]
neoListingText1['_labels'] = 'Listing_Text; Listing_Text'
neoListingText1 = neoListingText1.rename(columns={"_id": "id:ID(Listing_Text)", "_labels": ":LABEL"})
neoListingText1.to_csv(processor+"Neo4j/Listing_Text.csv",index=False)
neoReviewText1 = neoReviewText[['_id','name','_labels']]
neoReviewText1['_labels'] = 'Listing_Text; Listing_Text'
neoReviewText1 = neoReviewText1.rename(columns={"_id": "id:ID(Listing_Text)", "_labels": ":LABEL"})
neoReviewText1.to_csv(processor+"Neo4j/Listing_Text.csv",index=False)
# -
neoUser1 = neoUser[['_id','name','_labels']]
neoUser1['_labels'] = 'User; User'
neoUser1 = neoUser1.rename(columns={"_id": "id:ID(User)", "_labels": ":LABEL"})
neoUser1.to_csv(processor+"Neo4j/User.csv",index=False)
neoListings1 = neoListings[['_id','name','_labels','url','picture_url','host_identity_verified','accomodates','bedrooms','bathrooms','beds','price','review_scores_rating']]
neoListings1['_labels'] = 'Listing; Listing'
neoListings1 = neoListings1.rename(columns={"_id": "id:ID(Listing)", "_labels": ":LABEL"})
neoListings1.to_csv(processor+"Neo4j/Listings.csv",index=False)
# ### Define headers
# +
amenities_header = pd.DataFrame(columns=[':END_ID(Amenity)',':START_ID(Listing)'])
amenities_header.to_csv(processor+"Neo4j/amenity_header.csv",index=False)
city_header = pd.DataFrame(columns=[':END_ID(City)',':START_ID(Listing)'])
city_header.to_csv(processor+"Neo4j/city_header.csv",index=False)
state_header = pd.DataFrame(columns=[':END_ID(State)',':START_ID(Listing)'])
state_header.to_csv(processor+"Neo4j/state_header.csv",index=False)
country_header = pd.DataFrame(columns=[':END_ID(Country)',':START_ID(Listing)'])
country_header.to_csv(processor+"Neo4j/country_header.csv",index=False)
property_type_header = pd.DataFrame(columns=[':END_ID(property_type)',':START_ID(Listing)'])
property_type_header.to_csv(processor+"Neo4j/property_type_header.csv",index=False)
room_type_header = pd.DataFrame(columns=[':END_ID(room_type)',':START_ID(Listing)'])
room_type_header.to_csv(processor+"Neo4j/room_type_header.csv",index=False)
listing_text_header = pd.DataFrame(columns=[':END_ID(Listing_Text)',':START_ID(Listing)'])
listing_text_header.to_csv(processor+"Neo4j/listing_text_header.csv",index=False)
user_rating_header = pd.DataFrame(columns=[':END_ID(Listing)','RATED',':START_ID(User)'])
user_rating_header.to_csv(processor+"Neo4j/user_rating_header.csv",index=False)
# -
# ### Construct edges
# +
def UserListingRating(neoListings,neoUser,reviews):
neoListings['listing_id']=neoListings['name']
neoListings['listing_id'] = (neoListings['listing_id']).astype('str')
neoListings = neoListings[['listing_id','_id']]
trans_df = neoListings.set_index("listing_id").T
neoListingsDict = trans_df.to_dict("records")
reviews=reviews.replace({"listing_id": neoListingsDict[0]})
reviews['_end'] = reviews['listing_id']
neoUser['reviewer_id']= neoUser['name']
neoUser['reviewer_id'] = (neoUser['reviewer_id']).astype('int')
neoUser = neoUser[['reviewer_id','_id']]
trans_df = neoUser.set_index("reviewer_id").T
neoUserDict = trans_df.to_dict("records")
# print(neoUserDict)
reviews=reviews.replace({"reviewer_id": neoUserDict[0]})
# print(reviews)
reviews['_start'] = (reviews['reviewer_id']).astype('int')
reviews['_end'] = (reviews['listing_id']).astype('int')
reviews['_comments'] = (reviews['comments'])
reviews['_type'] = 'RATED'
reviews = reviews[['_end','rating','_start','_comments']]
reviews.to_csv(processor+'Neo4j/user_rating_review.csv',index=False,header=False)
reviews = reviews[['_end','rating','_start']]
reviews.to_csv(processor+'Neo4j/user_rating.csv',index=False,header=False)
UserListingRating(neoListings,neoUser,reviews)
# +
def ListingEdges(neoListings,neoEdge,listings,edge,type,id_col):
neoListings['listing_id']=neoListings['name']
neoListings['listing_id'] = (neoListings['listing_id'])
neoListings = neoListings[['listing_id','_id']]
trans_df = neoListings.set_index("listing_id").T
neoListingsDict = trans_df.to_dict("records")
listings=listings.replace({"id": neoListingsDict[0]})
listings['_start'] = listings['id']
neoEdge[edge]= neoEdge['name']
neoEdge[edge] = (neoEdge[edge]).astype(str)
neoEdge = neoEdge[[edge,'_id']]
trans_df = neoEdge.set_index(edge).T
neoEdgeDict = trans_df.to_dict("records")
listings=listings.replace({edge: neoEdgeDict[0]})
listings['_end'] = listings[edge]
listings['_type'] = type
header_list = ['_id','_labels','name', '_start','_end','_type','rating','url','picture_url','host_identity_verified','accomodates','bedrooms','bathrooms','beds','price','review_scores_rating']
listings = listings.reindex(columns = header_list)
listings = listings[['_end','_start']]
listings.to_csv(processor+'Neo4j/'+ edge +'_Listing.csv',index=False,header=False)
col_name = 'City'
listingCity = listings[['id',col_name]]
ListingEdges(neoListings,neoCity,listingCity,col_name,'IN_CITY','id:ID(City)')
col_name = 'State'
listingState = listings[['id',col_name]]
ListingEdges(neoListings,neoState,listingState,col_name,'IN_STATE','id:ID(State)')
col_name = 'Country'
listingCountry = listings[['id',col_name]]
ListingEdges(neoListings,neoCountry,listingCountry,col_name,'IN_COUNTRY','id:ID(Country)')
col_name = 'property_type'
listingPropType = listings[['id',col_name]]
ListingEdges(neoListings,neoProperty_type,listingPropType,col_name,'HAS_PROPERTY_TYPE','id:ID(property_type)')
col_name = 'room_type'
listingRoomType = listings[['id',col_name]]
ListingEdges(neoListings,neoRoom_type,listingRoomType,col_name,'HAS_ROOM_TYPE','id:ID(room_type)')
col_name = 'Listing_Text'
listingText = listings[['id',col_name]]
ListingEdges(neoListings,neoListingText,listingText,col_name,'HAS_TEXT','id:ID(Listing_Text)')
col_name = 'Review_Text'
reviewText = listings[['id',col_name]]
ListingEdges(neoListings,neoReviewText,reviewText,col_name,'HAS_REVIEW','id:ID(Review_Text)')
# -
def ListingAmenities(listings):
ListingID=[]
AmenityID=[]
for index in listings.index:
try:
listing_id = int(listings['id'][index])
if(listing_id>=1):
try:
#nodes
amenities = listings['amenities'][index]
for every_amenity in amenities:
ListingID.append(listing_id)
AmenityID.append(every_amenity)
except:
print('No amenties found')
except:
print('listing id null')
data={'id':ListingID,'Amenity':AmenityID}
listingsAmenity = pd.DataFrame(data)
return listingsAmenity
listingsAmenity=ListingAmenities(listings)
col_name = 'Amenity'
listingAmenities = listingsAmenity[['id',col_name]]
ListingEdges(neoListings,neoAmenities,listingAmenities,col_name,'HAS_AMENITY','aId:ID(Amenity)')
listingsAmenity
| 2d_Neo4jData_Gen.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jagatabhay/TSAI/blob/master/S15/DenseDepth.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="hm_cgXOfKV1u" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 407} outputId="327f2ceb-402f-4f6c-edb5-edabca39e228"
# !nvidia-smi
# + id="bH3mkE_mKGJc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 129} outputId="2363c4ad-706d-4727-9cd9-6d450268a026"
from google.colab import drive
drive.mount('/content/gdrive')
# + id="ahkR4C5dEnR0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 165} outputId="cf4e0c42-52d2-4dc4-ab17-dc42b6adb47a"
# !git clone https://github.com/jagatabhay/Deep-Learning.git
# + id="fFQgwMlNExak" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 240} outputId="d1554570-9e9a-49bb-84ba-2ab4120fc5d5"
# !wget https://s3-eu-west-1.amazonaws.com/densedepth/nyu.h5 -O ./Deep-Learning/depth_estimation/nyu.h5
# + id="u3BJP9KpK1HO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="65db1c95-7e82-4d73-b796-cafbf01d63a5"
import shutil
shutil.move('gdrive/My Drive/nyu_data.zip','Deep-Learning/depth_estimation')
# + id="AiJKd6uLE9Gr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="91a9d967-bf63-4bf0-941e-8ceec2241d92"
# cd Deep-Learning/depth_estimation
# + id="g2lou8HPh5tQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 148} outputId="ba07b4bc-24a3-4bac-9d96-82e0e4ece2ae"
# ls
# + id="09go-bvHiXg7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="f8d364de-e0c6-4a07-eaea-8a1714c45dbc"
# !python train.py --data nyu --bs 4 --full --dnetVersion small
# + id="642aaQnGfdNP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 146} outputId="34da56<PASSWORD>"
# !python test.py
# + id="HjzqM74-FfyL" colab_type="code" colab={}
'''from matplotlib import pyplot as plt
from skimage import io
plt.figure(figsize=(20,20))
plt.imshow( io.imread('results.png') )'''
| S15/DenseDepth.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/rwfchan/ResumeScanner/blob/main/MySQLSearchQueryBeta.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="1AUI-9ya0H1o"
# 1st parameter: folder path
# 2nd parameter: job title query
# 3rd parameter: degree query
# 4th parameter: major query
# 5th parameter: skill query
resumeFilter("", "", "", "", "")
# + id="C4uiE0uQM8Bw"
# 1st parameter: folder path
# 2nd parameter: partial query call to user's MySQL database
# "[column_name1] LIKE '%[keyword1]%' AND '%[keyword2]%' AND [column_name2] LIKE '%[keyword3]%'..."
# where column_namex can be ["Name, Organization, Position, Email, Phone, Skill, SchoolName, SchoolDegree"]
resumeQuery("", "")
# + id="pUwIZpJwy_cm"
import mysql.connector
def resumeFilter(path, jobs = None, degrees = None, majors = None , skills = None):
try:
# need to delete the database information when move to beta version
mydb = mysql.connector.connect(host="Enter Host Address", user="Enter Username", passwd="Enter Password", database="Enter Database Name")
print("Connection to MySQL DB successful")
except Error as e:
print(f"The error '{e}' occurred")
mycursor = mydb.cursor()
job_list = []
degree = []
skill_list = []
major_list = []
if jobs:
for job in jobs.split(' '):
job_list.append(job)
if majors:
for major in majors.split(' '):
major_list.append(major)
if degrees:
degrees = degrees.lower()
if degrees[0] == 'b':
degree = ['BS','B S', 'bachelor']
elif degrees[0] == 'm':
degree = ['master', 'M S', 'MS']
elif degrees[0] == 'p':
degree = ['PhD', 'Ph D', 'Philosophy']
else:
print('no such degree')
degree = []
if skills:
skill_list = skills.split(',')
# need to change the database and table name
query = "SELECT * FROM csv_output.resume_scanner_beta WHERE "
if job_list:
job_query = "("
for job in job_list:
job_query += "Position LIKE '%" + job + "%' AND "
job_query = job_query[:-5] + ")"
query += job_query
if job_list and degree:
query += " AND "
if degree:
degree_query = "("
for degree_title in degree:
degree_query += "SchoolDegree LIKE '%" + degree_title + "%' OR "
degree_query = degree_query[:-4] + ")"
query += degree_query
if (job_list or degree) and skill_list:
query += " AND "
if skill_list:
skill_query = "("
for skill in skill_list:
skill_query += "Skill LIKE '%" + skill + "%' AND "
skill_query = skill_query[:-5] + ")"
query += skill_query
if (job_list or degree or skill_list) and major_list:
query += " AND "
if major_list:
major_query = "("
for major in major_list:
major_query += "SchoolDegree LIKE '%" + major + "%' AND "
major_query = major_query[:-5] + ")"
query += major_query
import pandas as pd
print("Reading BLOB data from resume_scanner_beta table")
try:
mycursor.execute(query)
record = mycursor.fetchall()
print(record)
directory = 'output'
dir_path = os.path.join(path,directory)
if not os.path.isdir(dir_path):
os.mkdir(dir_path)
for row in record:
image = row[10]
print("Storing employee image and bio-data on disk \n")
new_path = dir_path + "/" + row[0][2:-2] + ".jpg"
write_file(image, new_path)
except mysql.connector.Error as error:
print("Failed to read BLOB data from MySQL table {}".format(error))
finally:
if (mydb.is_connected()):
mycursor.close()
mydb.close()
print("MySQL connection is closed")
# + id="H4TBNeV7MxJ1"
# SQL injection
# input: folder path, sql query
# output: the jpg file
def resumeQuery(path, user_query):
try:
# need to delete the database information when move to beta version
mydb = mysql.connector.connect(host="Enter Host Address", user="Enter Username", passwd="<PASSWORD>", database="Enter Database Name")
print("Connection to MySQL DB successful")
except Error as e:
print(f"The error '{e}' occurred")
mycursor = mydb.cursor()
# need to change the database and table name
query = "SELECT * FROM csv_output.resume_scanner_beta WHERE "
final_query = query + user_query
import pandas as pd
print("Reading BLOB data from resume_scanner_beta table")
try:
mycursor.execute(final_query)
record = mycursor.fetchall()
print(record)
directory = 'output'
dir_path = os.path.join(path,directory)
if not os.path.isdir(dir_path):
os.mkdir(dir_path)
for row in record:
image = row[10]
print("Storing employee image and bio-data on disk \n")
new_path = dir_path + "/" + row[0][2:-2] + ".jpg"
write_file(image, new_path)
except mysql.connector.Error as error:
print("Failed to read BLOB data from MySQL table {}".format(error))
finally:
if (mydb.is_connected()):
mycursor.close()
mydb.close()
print("MySQL connection is closed")
# + id="jurtslKNIdRP"
pip install mysql-connector-python
# + id="mwz-_1pIvdbc"
def write_file(data, filename):
# Convert binary data to proper format and write it on Hard Disk
with open(filename, 'wb') as file:
file.write(data)
| MySQLSearchQueryBeta.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.1
# language: julia
# name: julia-1.6
# ---
function interiorPoints(C)
rayMatrix=Array(Polymake.common.primitive(C.RAYS))
l=size(rayMatrix,1)
dim=size(rayMatrix,2)
if rank(rayMatrix)<l
error("Input cone is not simplicial.")
end
subsets=collect(powerset([1:l;]))
vertices=[]
for elt in subsets
vert=zeros(Polymake.Rational,1,dim)
for i in 1:l
if i in elt
vert+=rayMatrix[[i],:]
end
end
append!(vertices,[vert])
end
V=vcat(vertices...)
VH=hcat(ones(Polymake.Rational,size(V,1)),V)
P=Polymake.polytope.Polytope(POINTS=VH)
#print(P.POINTS)
if size(P.INTERIOR_LATTICE_POINTS,1)==0
return nothing
end
intPoints=Array(P.INTERIOR_LATTICE_POINTS)[:,2:(dim+1)]
validPoints=[]
#return intPoints
for i in 1:size(intPoints,1)
point=intPoints[i,:]
if gcd(point)==1
append!(validPoints,[point])
end
end
return Array(vcat(validPoints...))
end
C=Polymake.polytope.Cone(RAYS=[1 2;2 1])
A=interiorPoints(C)
| BerghAlgorithms/src/OldFiles/interiorPoints.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Getting Started with ctapipe
#
# This hands-on was presented at the Paris CTA Consoritum meeting (<NAME>)
# ## Part 1: load and loop over data
from ctapipe.io import event_source
from ctapipe import utils
from matplotlib import pyplot as plt
# %matplotlib inline
path = utils.get_dataset_path("gamma_test_large.simtel.gz")
source = event_source(path, max_events=4)
for event in source:
print(event.count, event.r0.event_id, event.mc.energy)
event
event.r0
for event in source:
print(event.count, event.r0.tels_with_data)
event.r0.tel[2]
r0tel = event.r0.tel[2]
r0tel.waveform
r0tel.waveform.shape
# note that this is ($N_{channels}$, $N_{pixels}$, $N_{samples}$)
plt.pcolormesh(r0tel.waveform[0])
plt.plot(r0tel.waveform[0,10])
# +
from ipywidgets import interact
@interact
def view_waveform(chan=0, pix_id=200):
plt.plot(r0tel.waveform[chan, pix_id])
# -
# try making this compare 2 waveforms
# ## Part 2: Explore the instrument description
# This is all well and good, but we don't really know what camera or telescope this is... how do we get instrumental description info?
#
# Currently this is returned *inside* the event (it will soon change to be separate in next version or so)
subarray = event.inst.subarray # soon EventSource will give you event, subarray separate
subarray
subarray.peek()
subarray.to_table()
subarray.tel[2]
subarray.tel[2].camera
subarray.tel[2].optics
tel = subarray.tel[2]
tel.camera
tel.optics
tel.camera.pix_x
tel.camera.to_table()
tel.optics.mirror_area
from ctapipe.visualization import CameraDisplay
disp = CameraDisplay(tel.camera)
disp = CameraDisplay(tel.camera)
disp.image = r0tel.waveform[0,:,10] # display channel 0, sample 0 (try others like 10)
# ** aside: ** show demo using a CameraDisplay in interactive mode in ipython rather than notebook
# ## Part 3: Apply some calibration and trace integration
from ctapipe.calib import CameraCalibrator
calib = CameraCalibrator()
for event in source:
calib.calibrate(event) # fills in r1, dl0, and dl1
print(event.dl1.tel.keys())
event.dl1.tel[2]
dl1tel = event.dl1.tel[2]
dl1tel.image.shape # note this will be gain-selected in next version, so will be just 1D array of 1855
dl1tel.pulse_time
CameraDisplay(tel.camera, image=dl1tel.image[0])
CameraDisplay(tel.camera, image=dl1tel.pulse_time[0])
# Now for Hillas Parameters
from ctapipe.image import hillas_parameters, tailcuts_clean
image = dl1tel.image[0]
mask = tailcuts_clean(tel.camera, image, picture_thresh=10, boundary_thresh=5)
mask
CameraDisplay(tel.camera, image=mask)
cleaned = image.copy()
cleaned[~mask] = 0
disp = CameraDisplay(tel.camera, image=cleaned)
disp.cmap = plt.cm.coolwarm
disp.add_colorbar()
plt.xlim(-1.0,0)
plt.ylim(0,1.0)
params = hillas_parameters(tel.camera, cleaned)
print(params)
disp = CameraDisplay(tel.camera, image=cleaned)
disp.cmap = plt.cm.coolwarm
disp.add_colorbar()
plt.xlim(-1.0,0)
plt.ylim(0,1.0)
disp.overlay_moments(params, color='white', lw=2)
# ## Part 4: Let's put it all together:
# - loop over events, selecting only telescopes of the same type (e.g. LST:LSTCam)
# - for each event, apply calibration/trace integration
# - calculate Hillas parameters
# - write out all hillas paremeters to a file that can be loaded with Pandas
# first let's select only those telescopes with LST:LSTCam
subarray.telescope_types
subarray.get_tel_ids_for_type("LST:LSTCam")
# Now let's write out program
data = utils.get_dataset_path("gamma_test_large.simtel.gz")
source = event_source(data, allowed_tels=[1,2,3,4], max_events=10) # remove the max_events limit to get more stats
for event in source:
calib.calibrate(event)
for tel_id, tel_data in event.dl1.tel.items():
tel = event.inst.subarray.tel[tel_id]
mask = tailcuts_clean(tel.camera, tel_data.image[0])
params = hillas_parameters(tel.camera[mask], tel_data.image[0][mask])
from ctapipe.io import HDF5TableWriter
with HDF5TableWriter(filename='hillas.h5', group_name='dl1', overwrite=True) as writer:
for event in source:
calib.calibrate(event)
for tel_id, tel_data in event.dl1.tel.items():
tel = event.inst.subarray.tel[tel_id]
mask = tailcuts_clean(tel.camera, tel_data.image[0])
params = hillas_parameters(tel.camera[mask], tel_data.image[0][mask])
writer.write("hillas", params)
# ### We can now load in the file we created and plot it
# !ls *.h5
# +
import pandas as pd
hillas = pd.read_hdf("hillas.h5", key='/dl1/hillas')
hillas
# -
_ = hillas.hist(figsize=(8,8))
# If you do this yourself, loop over more events to get better statistics
| docs/tutorials/ctapipe_handson.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:azureml_py36_tensorflow]
# language: python
# name: conda-env-azureml_py36_tensorflow-py
# ---
# +
import numpy as np
import pandas as pd
import os
from pathlib import Path
import glob
import json
import tensorflow as tf
import tensorflow_datasets as tfds
import os
import nltk
import cv2
import matplotlib.pyplot as plt
import random
from wordcloud import WordCloud
import seaborn as sns
#Check GPU is available for training or not Or whether the tensorflow version can utilize gpu
physical_devices = tf.config.list_physical_devices('GPU')
print("Number of GPUs :", len(physical_devices))
print("Tensorflow GPU :",tf.test.is_built_with_cuda())
if len(physical_devices)>0:
device="/GPU:0"
else:
device="/CPU:0"
# -
def load_data(N, mode='train'):
with open(f'clevr/CLEVR_v1/questions/CLEVR_{mode}_questions.json') as f:
data = json.load(f)
data_set_size = len(data['questions'])
records = []
for k in np.random.uniform(0, data_set_size, N).astype(int):
i = data['questions'][k]
temp=[]
for path in glob.glob(f'clevr/CLEVR_v1/images/{mode}/'+i['image_filename']):
temp.append(path)
temp.append(i['question'])
temp.append(i['answer'])
records.append(temp)
labels=['Path','Question','Answer']
return pd.DataFrame.from_records(records, columns=labels)
train_df = load_data(70000, 'train')
val_df = load_data(15000, 'val')
train_df.head()
val_df.head()
# +
def tokenize_qs(questions, tokenizer):
res = set()
for question in questions:
res.update(tokenizer.tokenize(question))
return res
tokenizer = tfds.deprecated.text.Tokenizer()
answer_set = set(np.append(train_df['Answer'].unique(),val_df['Answer'].unique()))
question_set = set(np.append(train_df['Question'].unique(),val_df['Question'].unique()))
vocab_set = tokenize_qs(question_set, tokenizer) | answer_set
encoder=tfds.deprecated.text.TokenTextEncoder(vocab_set)
answer_encoder=tfds.deprecated.text.TokenTextEncoder(answer_set)
# +
BATCH_SIZE=32
IMG_SIZE=(128,128)
def encode_fn(text):
return np.array(encoder.encode(text.numpy()))
def answer_encode_fn(text):
return np.array(answer_encoder.encode(text.numpy()))
def preprocess(ip,ans):
img,ques=ip
img=tf.io.read_file(img)
img=tf.image.decode_jpeg(img,channels=3)
img=tf.image.resize(img,IMG_SIZE)
img=tf.math.divide(img, 255)#
ques=tf.py_function(encode_fn,inp=[ques],Tout=tf.int32)
paddings = [[0, 50-tf.shape(ques)[0]]]
ques = tf.pad(ques, paddings, 'CONSTANT', constant_values=0)
ques.set_shape([50])
ans=tf.py_function(answer_encode_fn,inp=[ans],Tout=tf.int32)
ans.set_shape([1])
return (img,ques),ans
def create_pipeline(dataframe):
raw_df=tf.data.Dataset.from_tensor_slices(((dataframe['Path'],dataframe['Question']),dataframe['Answer']))
df=raw_df.map(preprocess)
df=df.batch(BATCH_SIZE)
return df
train_dataset=create_pipeline(train_df)
validation_dataset=create_pipeline(val_df)
# +
from tensorflow.keras.layers import Conv2D, MaxPooling2D, Flatten
from tensorflow.keras.layers import Input, LSTM, Embedding, Dense
from tensorflow.keras.models import Model, Sequential
class CNN_plus_LSTM(tf.keras.Model):
def __init__(self):
super(CNN_plus_LSTM, self).__init__()
self.conv2d_padding1 = tf.keras.layers.Conv2D(64, kernel_size=(3, 3), padding='same', activation='relu', input_shape=(224, 224, 3))
self.conv2d1 = tf.keras.layers.Conv2D(64, kernel_size=(3, 3), activation='relu')
self.max_pooling1 = tf.keras.layers.MaxPooling2D((2, 2))
self.conv2d_padding2 = tf.keras.layers.Conv2D(128, kernel_size=(3, 3), padding='same', activation='relu', input_shape=(224, 224, 3))
self.conv2d2 = tf.keras.layers.Conv2D(128, kernel_size=(3, 3), activation='relu')
self.max_pooling2 = tf.keras.layers.MaxPooling2D((2, 2))
self.conv2d_padding3 = tf.keras.layers.Conv2D(256, kernel_size=(3, 3), padding='same', activation='relu', input_shape=(224, 224, 3))
self.conv2d3 = tf.keras.layers.Conv2D(256, kernel_size=(3, 3), activation='relu')
self.conv2d4 = tf.keras.layers.Conv2D(256, kernel_size=(3, 3), activation='relu')
self.max_pooling3 = tf.keras.layers.MaxPooling2D((2, 2))
self.flatten = Flatten()
self.embeding = Embedding(input_dim=len(vocab_set)+1, output_dim=256, input_length=100)
self.lstm = LSTM(256)
self.conc = tf.keras.layers.Concatenate()
self.fc = Dense(len(answer_set) + 1, activation='softmax')
def call(self, inputs):
#Image input
x1 = self.conv2d_padding1(inputs[0])
x1 = self.conv2d1(x1)
x1 = self.max_pooling1(x1)
x1 = self.conv2d_padding2(x1)
x1 = self.conv2d2(x1)
x1 = self.max_pooling2(x1)
x1 = self.conv2d_padding3(x1)
x1 = self.conv2d3(x1)
x1 = self.conv2d4(x1)
x1 = self.max_pooling3(x1)
x1 = self.flatten(x1)
#Question input
x2 = self.embeding(inputs[1])
x2 = self.lstm(x2)
#Unite
x = self.conc([x1,x2])
return self.fc(x)
def model(self, im_shape, vocab_size):
x1, x2 = Input(shape=im_shape), Input(shape=(vocab_size,))
return Model(inputs=[x1, x2], outputs=self.call([x1, x2]))
# -
model = CNN_plus_LSTM()
model = model.model((IMG_SIZE[0], IMG_SIZE[1], 3), 50)
model.compile('adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
model.summary()
tf.keras.utils.plot_model(model, show_shapes=True)
epochs=50
with tf.device(device):
history = model.fit(train_dataset,
validation_data=validation_dataset,
epochs=epochs)
losses = pd.DataFrame(history.history)
losses[['val_accuracy', 'accuracy']].plot()
losses[['val_loss', 'loss']].plot()
# +
import datetime
date_str = datetime.datetime.now().strftime("%Y-%m-%d %H:%M:%S")
val_accuracy = losses[['val_accuracy']].values[-1][0]
date_str = datetime.datetime.now().strftime("%Y-%m-%d_%H:%M:%S")
checkpoint_path = f"models/cnn_plus_lstm-{date_str}-{val_accuracy}-{epochs}"
model.save(checkpoint_path)
# +
# checkpoint_path = 'models/cnn_plus_lstm-2021-05-21_01:17:49-0.45873332023620605-50'
# model = tf.keras.models.load_model(checkpoint_path)
# +
# predictions = model.predict(validation_dataset)
# -
| CNN_plus_LSTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Learning Objectives:
#
# 1. Reading files
# 2. Exploring the read dataframe
# 3. Checking the dataframe info
# 4. Merging the two dataframes into one
# 5. Defining questions for the analysis
# 6. Cleaning Steps
import numpy as np
import pandas as pd
# ## Reading the files
# +
movies_df = pd.read_csv("../data/movies.csv")
credits_df = pd.read_csv("https://raw.githubusercontent.com/harshitcodes/tmdb_movie_data_analysis/master/tmdb-5000-movie-dataset/tmdb_5000_credits.csv")
# -
# ## Exploring the read dataframe
#
# * Look at samples rows
# * Columns and shape of the dataframe
# * Check if you can merge the files
# * Understanding the type of questions that we can answer using the data.
# * Define the cleaning steps - using pandas
# * Start looking for answers
movies_df.head()
movies_df.sample(5)
movies_df.shape
credits_df.head()
# ## Checking the dataframe info
movies_df.info()
credits_df.info()
movies_df.head()
credits_df.head()
# ## Merging the two dataframes
movies_df = pd.merge(movies_df, credits_df, left_on='id', right_on='movie_id')
movies_df.head()
# ## Define questions for the analysis
#
# 1. Which are the top 5 most expensive movies?
# 2. Top profitable movies? Comparision between min and max profits.
# 3. Most talked about movies.
# 4. Average runtime of movies.
# 5. Movies which are rated above 7 by critics.
# 6. Which year did we have the most profitable movies?
# ## Cleaning Steps
#
# 1. Need to remove redundant columns
# 2. Remove duplicate rows
# 3. Some movies in the data have zero budget/ zero revenue, that us there valueis not recorded.
# 4. Change the data types of columns wherever required.
# 5. Replacing zero with NAN in runtime column.
# 6. Changing the format of budget and revenue.
# 7. Cleaning or flattening the genres, cast and other columns that contain JSON data.
#
| module_4_pandas/exploratory_analysis_part1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#data format library
import h5py
#numpy
import numpy as np
import numpy.ma as ma
import matplotlib.pyplot as plt
# # %matplotlib notebook
from sklearn.cluster import KMeans
import sys
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.colors as colors
import os
from scipy.integrate import odeint
#change path to where 'manuscript_data' is saved
dir_path = '/home/antonio/Repositories/manuscript_data/'
sys.path.append(dir_path+'/code/utils/')
import operator_calculations as op_calc
import stats
plt.rc('text', usetex=True)
plt.rc('font',size=14)
# +
k_B_T = 0.5
f = h5py.File(dir_path+'DoubleWell/simulations/simulation_k_B_T_{}.h5'.format(k_B_T),'r')
sim = np.array(f['simulation'])
T = np.array(f['MetaData/T'])[0]
discard_t = np.array(f['MetaData/discarded_t'])[0]
dt = np.array(f['MetaData/integration_step'])[0]
print(sim.shape)
f.close()
print(dt,discard_t)
f = h5py.File(dir_path+'DoubleWell/embedding/phspace_k_B_T_{}.h5'.format(k_B_T),'r')
# print(list(f.keys()))
traj_matrix = np.array(f['traj_matrix'])
K_star = np.array(f['K_star'],dtype=int)[0]
m_star = np.array(f['m_star'],dtype=int)[0]
phspace = np.array(f['phspace'])
print(traj_matrix.shape)
f.close()
f = h5py.File(dir_path+'DoubleWell/tscales_compute_1000_clusters.h5','r')
print(list(f.keys()))
T_range = np.array(list(f.keys()),dtype='float')
ts_traj_T = []
for T in T_range:
ts_traj_delay = np.array(f[str(T)]['ts_traj_delay'])
delay_range = np.array(f[str(T)]['delay_range'])
length = int(np.array(f[str(T)]['seq_length'])[0])
ts_traj_T.append(ts_traj_delay)
f.close()
print(length*dt)
ts_traj_delay = ts_traj_T[0]
# -
n_clusters=1000
f = h5py.File(dir_path+'DoubleWell/symbol_sequences/labels_phspace_k_B_T_{}_nseeds_{}.h5'.format(k_B_T,n_clusters),'r')
labels_traj = np.array(f['labels_traj'],dtype=int)
centers_traj = np.array(f['centers_traj'])
labels_phspace = np.array(f['labels_phspace'],dtype=int)
centers_phspace = np.array(f['centers_phspace'])
f.close()
# # Projected Boltzmann distribution
# +
def Boltzmann_dist(x,y,k_B_T,gamma=1):
beta = 1/k_B_T
return np.exp(-beta*((x**2-1)**2+gamma*y**2/2))
from sklearn.linear_model import LinearRegression
X = sim[int(K_star/2):-int(K_star/2)-1,:]
y = phspace[:,:2]
reg = LinearRegression().fit(X, y)
R2 = reg.score(X, y)
m = reg.coef_
b = reg.intercept_
centers_proj = centers_phspace.dot(np.linalg.pinv(m))
P_B_phspace = np.array([Boltzmann_dist(x_,y_,k_B_T) for x_,y_ in centers_proj])
P_B_phspace = P_B_phspace/P_B_phspace.sum()
# -
# # operator time scales
# +
mean = np.mean(ts_traj_delay[:,:,0],axis=0)
cil = np.percentile(ts_traj_delay[:,:,0],2.5,axis=0)
ciu = np.percentile(ts_traj_delay[:,:,0],97.5,axis=0)
mean_1 = np.mean(ts_traj_delay[:,:,1],axis=0)
cil_1 = np.percentile(ts_traj_delay[:,:,1],2.5,axis=0)
ciu_1 = np.percentile(ts_traj_delay[:,:,1],97.5,axis=0)
# +
kd=199
delay = int(delay_range[kd])
print(delay*dt)
P = op_calc.transition_matrix(labels_phspace,delay)
prob = op_calc.stationary_distribution(P)
R = op_calc.get_reversible_transition_matrix(P)
eigvals,eigvecs = op_calc.sorted_spectrum(R,k=10)
PF_eigvecs = op_calc.sorted_spectrum(R.T,k=10)[1]
# -
sizes = np.array([np.var(phspace[labels_phspace==label,:2] - centers_phspace[label]) for label in np.unique(labels_phspace)])
p_phspace = prob/sizes
p_phspace = p_phspace/p_phspace.sum()
plt.figure()
plt.title('$t_1 = {:.3f}\,({:.3f},{:.3f}) \, s$'.format(mean[kd],cil[kd],ciu[kd]))
plt.tricontourf(centers_phspace[:,0],centers_phspace[:,1],eigvecs[:,1].real,cmap='seismic',levels=np.linspace(-.05,.05,1000))
plt.xlim(-0.0004,0.0004)
plt.ylim(-0.001,0.001)
plt.colorbar(ticks = np.linspace(-0.05,0.05,9))
# plt.savefig('Koopman_eigfun_DW_k_B_T_{}_delay_Fig.pdf'.format(k_B_T))
plt.show()
# # Fig4B_simulation
# +
idx = np.argmin(np.linalg.norm(centers_phspace-np.array([0.00025,0]),axis=1))
rho_0 = np.zeros(n_clusters)
rho_0[idx] = 1
n_iters= int(10000/delay)
new_rho = np.zeros((n_iters,n_clusters))
new_rho[0] = rho_0
for k in range(n_iters-1):
new_rho[k+1,:] = (P.T).dot(new_rho[k])
# -
plt.scatter(centers_phspace[:,0],centers_phspace[:,1],c=new_rho[0]/sizes)
plt.xlim(-0.0006,0.0006)
plt.ylim(-0.001,0.001)
plt.show()
k=1
fig = plt.figure(figsize=(5,5))
ax = fig.add_subplot(111, projection='3d',)
ax.view_init(elev=15., azim=53)
p = new_rho[k,:]/sizes
p = p/p.sum()
# ax.plot_trisurf(centers[:,0],centers[:,1],p,alpha=.5,cmap='jet')
ax.scatter(centers_phspace[:,0],centers_phspace[:,1],p,cmap='jet',c=p,s=10)#, linewidth=0, edgecolor='none', antialiased=False)
ax.plot_trisurf(centers_phspace[:,0],centers_phspace[:,1],P_B_phspace/P_B_phspace.sum(),alpha=.2)#,cmap='jet',c=prob,s=10)#, linewidth=0, edgecolor='none', antialiased=False)
# plt.axis('off')
ax.zaxis.set_rotate_label(False)
ax.set_xlabel(r'$x$',fontsize=12)
ax.set_ylabel(r'$y$',fontsize=12)
ax.set_zlabel(r'$\rho$',fontsize=12,rotation=0)
# ax.set_zlim(0,0.08)
ax.set_xlim(-4e-4,4e-4)
ax.set_ylim(-9e-4,8e-4)
ax.set_zlim(0,6e-3)
ax.text2D(-.05, .05, "t = {:.02f} s".format(k*delay*dt), color='k',fontsize=12)
plt.tight_layout()
# plt.savefig('img_{:05d}_k_B_T_{}.pdf'.format(k,k_B_T))
plt.show()
# +
k=2
fig = plt.figure(figsize=(5,5))
ax = fig.add_subplot(111, projection='3d',)
ax.view_init(elev=15., azim=53)
p = new_rho[k,:]/sizes
p = p/p.sum()
ax.scatter(centers_phspace[:,0],centers_phspace[:,1],p,cmap='jet',c=p,s=10)#, linewidth=0, edgecolor='none', antialiased=False)
ax.plot_trisurf(centers_phspace[:,0],centers_phspace[:,1],P_B_phspace/P_B_phspace.sum(),alpha=.2)#,cmap='jet',c=prob,s=10)#, linewidth=0, edgecolor='none', antialiased=False)
ax.zaxis.set_rotate_label(False)
ax.set_xlabel(r'$x$',fontsize=12)
ax.set_ylabel(r'$y$',fontsize=12)
ax.set_zlabel(r'$\rho$',fontsize=12,rotation=0)
ax.set_xlim(-4e-4,4e-4)
ax.set_ylim(-9e-4,8e-4)
ax.set_zlim(0,4e-3)
ax.text2D(-.05, .05, "t = {:.02f} s".format(k*delay*dt), color='k',fontsize=12)
plt.tight_layout()
# plt.savefig('img_{:05d}_k_B_T_{}.pdf'.format(k,k_B_T))
plt.show()
# +
k=4
fig = plt.figure(figsize=(5,5))
ax = fig.add_subplot(111, projection='3d',)
ax.view_init(elev=15., azim=53)
p = new_rho[k,:]/sizes
p = p/p.sum()
ax.scatter(centers_phspace[:,0],centers_phspace[:,1],p,cmap='jet',c=p,s=10)#, linewidth=0, edgecolor='none', antialiased=False)
ax.plot_trisurf(centers_phspace[:,0],centers_phspace[:,1],P_B_phspace/P_B_phspace.sum(),alpha=.2)#,cmap='jet',c=prob,s=10)#, linewidth=0, edgecolor='none', antialiased=False)
ax.zaxis.set_rotate_label(False)
ax.set_xlabel(r'$x$',fontsize=12)
ax.set_ylabel(r'$y$',fontsize=12)
ax.set_zlabel(r'$\rho$',fontsize=12,rotation=0)
ax.set_xlim(-4e-4,4e-4)
ax.set_ylim(-9e-4,8e-4)
ax.set_zlim(0,3e-3)
ax.text2D(-.05, .05, "t = {:.02f} s".format(k*delay*dt), color='k',fontsize=12)
plt.tight_layout()
# plt.savefig('img_{:05d}_k_B_T_{}.pdf'.format(k,k_B_T))
plt.show()
# +
t_range = np.arange(0,200)
plt.figure(figsize=(5,5))
plt.plot(t_range,np.exp(-(1/mean[kd])*t_range))
plt.fill_between(t_range,np.exp(-(1/cil[kd])*t_range),np.exp(-(1/ciu[kd])*t_range),alpha=.5)
plt.xlim(0,90)
plt.ylim(0,1)
plt.text(30,0.9,'$t_1 = {:.3f}\,({:.3f},{:.3f}) \, s$'.format(mean[kd],cil[kd],ciu[kd]),fontsize=12)
# plt.savefig('eigfun_decay_k_B_T_{}.pdf'.format(k_B_T))
plt.show()
# -
# # Hopping rates
# +
from sklearn.linear_model import LinearRegression
def decay_rate(lifetimes,xrange=[2,500],plot=False,min_counts=10):
x,y = stats.cumulative_dist(lifetimes,xrange)
y = 1-np.array([np.mean(y[x==x_unique]) for x_unique in np.unique(x)])
counts = np.array([np.sum(x==x_unique) for x_unique in np.unique(x)])
sel = counts>min_counts
logy = np.log(y[sel])
x = np.sort(np.unique(x))[sel].reshape(-1,1)
w = counts[sel]
y = y[sel]
reg = LinearRegression().fit(x, logy)#,sample_weight=w)
R2 = reg.score(x, logy)
m = reg.coef_
b = reg.intercept_
if plot==True:
plt.title('m = {:.2f}; R2 = {:.5f}'.format(-1/m[0],R2))
plt.scatter(x,y)
plt.plot(x,np.exp(m*x+b))
plt.yscale('log')
plt.show()
return np.abs(m[0])
# +
# participation ratio is probably not a good idea either!!!
delay_idx_T = np.zeros(len(T_range),dtype=int)
rate_mix_T = np.zeros((len(T_range),3))
delay_mix_T = np.zeros((len(T_range),3))
for kt,T in enumerate(T_range):
ts_traj_delay = ts_traj_T[kt]
mean = np.mean(1/ts_traj_delay[:,:,0],axis=0)
cil = np.percentile(1/ts_traj_delay[:,:,0],0.5,axis=0)
ciu = np.percentile(1/ts_traj_delay[:,:,0],99.5,axis=0)
std = np.std(1/ts_traj_delay[:,:,0],axis=0)/2
kd = np.argmax(mean)
print(kd,delay_range[kd]*dt)
rate_mix_T[kt,:] = np.array([mean[kd],cil[kd],ciu[kd]])
delay_idx_T[kt] = delay_range[kd]
mean = np.mean(ts_traj_delay[:,:,0],axis=0)
cil = np.percentile(ts_traj_delay[:,:,0],2.5,axis=0)
ciu = np.percentile(ts_traj_delay[:,:,0],97.5,axis=0)
delay_mix_T[kt,:] = np.array([mean[kd],cil[kd],ciu[kd]])
# -
def obtain_tscales_T(kt,k_B_T,delay,length,plot=False):
f = h5py.File(dir_path+'DoubleWell/symbol_sequences/labels_phspace_k_B_T_{}_nseeds_1000.h5'.format(k_B_T),'r')
labels = ma.array(f['labels_traj'],dtype=int)
centers = np.array(f['centers_traj'])
f.close()
f = h5py.File(dir_path+'DoubleWell/simulations/simulation_k_B_T_{}.h5'.format(k_B_T),'r')
sim = np.array(f['simulation'])[int(K_star/2):-int(K_star/2)-1]
f.close()
print(labels.shape,sim.shape)
print(k_B_T,delay)
lcs,P = op_calc.transition_matrix(labels,delay,return_connected=True)
inv_measure = op_calc.stationary_distribution(P)
final_labels = op_calc.get_connected_labels(labels,lcs)
n_modes=2
R = op_calc.get_reversible_transition_matrix(P)
eigvals,eigvecs = op_calc.sorted_spectrum(R,k=n_modes)
eigfunctions = eigvecs.real/np.linalg.norm(eigvecs.real,axis=0)
phi2 = eigfunctions[:,1]
#label according to potential
cluster_traj = ma.zeros(labels.shape,dtype=int)
cluster_traj[sim[:,0]>0] = 1
split_trajs = op_calc.get_split_trajs(cluster_traj,int(length))
data_decay_rates = []
data_decay_times = []
for traj in split_trajs:
data_lifetimes = stats.state_lifetime(ma.masked_invalid(traj),dt)
data_decay_rate = decay_rate(np.hstack(data_lifetimes),xrange=[2,500],plot=plot,min_counts=1)
data_decay_times.append(1/data_decay_rate)
data_decay_rates.append(data_decay_rate)
#save optimal partition results
c_range,rho_sets,_,_ = op_calc.optimal_partition(phi2,inv_measure,R,return_rho=True)
return c_range,rho_sets,data_decay_times,data_decay_rates
# +
Tmin=0
scaled_T = (T_range - Tmin) / T_range.ptp()
colors_ = plt.cm.Reds(scaled_T)
# +
nseqs = len(op_calc.get_split_trajs(labels_phspace,length))
tscales_T = np.zeros((len(T_range),nseqs))
decay_rates_T = np.zeros((len(T_range),nseqs))
plt.figure(figsize=(5,5))
crange_T = []
rho_sets_T = []
for kt,k_B_T in enumerate(T_range):
delay=delay_idx_T[kt]
print(delay*dt)
c_range,rho_sets,data_tscales,data_decay_rates = obtain_tscales_T(kt,k_B_T,delay,length)
tscales_T[kt] = np.array(data_tscales)
decay_rates_T[kt] = np.array(data_decay_rates)
crange_T.append(c_range)
rho_sets_T.append(rho_sets)
print(np.mean(data_tscales),np.mean(data_decay_rates))
# -
plt.figure(figsize=(5,5))
for kt,k_B_T in enumerate(T_range):
c_range = crange_T[kt]
rho_sets = rho_sets_T[kt]
plt.plot(c_range,rho_sets[:,0],lw=2,c=colors_[kt])
plt.plot(c_range,rho_sets[:,1],lw=2,c=colors_[kt])
rho_c = np.min(rho_sets,axis=1)
plt.plot(c_range,rho_c,c='gray',ls='--')
plt.ylim(0,1)
plt.xlim(-0.05,0.05)
plt.axvline(0,ls='--',c='k')
plt.xlabel(r'$\phi_2$',fontsize=15)
plt.ylabel(r'$\rho$',fontsize=15)
plt.xticks(fontsize=12)
plt.tight_layout()
# plt.savefig('coherence_DW.pdf')
# plt.xlim(-0.04,0.04)
plt.show()
decay_rates_T_ci = np.vstack([np.mean(decay_rates_T,axis=1),np.percentile(decay_rates_T,2.5,axis=1),np.percentile(decay_rates_T,97.5,axis=1)]).T
tscales_T_ci = np.vstack([np.mean(tscales_T,axis=1),np.percentile(tscales_T,2.5,axis=1),np.percentile(tscales_T,97.5,axis=1)]).T
# +
plt.figure(figsize=(5,5))
plt.plot(np.arange(0,.51,.02),np.arange(0,.51,.02),ls='--',c='k')
for kt in np.arange(len(T_range)):
cil_PF = rate_mix_T[kt,0]-rate_mix_T[kt,1]
ciu_PF = rate_mix_T[kt,2]-rate_mix_T[kt,0]
cil_data = decay_rates_T_ci[kt,0]-decay_rates_T_ci[kt,1]
ciu_data = decay_rates_T_ci[kt,2]-decay_rates_T_ci[kt,0]
plt.errorbar(decay_rates_T_ci[kt,0],.5*rate_mix_T[kt,0],
yerr = [[.5*cil_PF],[.5*ciu_PF]],xerr = [[cil_data],[ciu_data]], c=colors_[kt],capsize=3,fmt='o',ms=5)
plt.xticks(fontsize=12)
plt.yticks(fontsize=12)
plt.xlabel('hopping rate $(s^{-1})$ ',fontsize=14)
plt.ylabel('$|\Lambda_2|/2\, (s^{-1})$ ',fontsize=14)
plt.xlim(0,0.42)
plt.ylim(0,0.42)
# plt.savefig('hopping_rate_diagonal_idx_{}.pdf'.format(idx))
plt.show()
# -
half_tscales_T_ci = np.vstack([np.mean(tscales_T*.5,axis=1),np.percentile(tscales_T*.5,2.5,axis=1),np.percentile(tscales_T*.5,97.5,axis=1)]).T
y_operator = []
y_data = []
for kt in range(len(T_range)):
ts_traj_delay = ts_traj_T[kt] #operator
mean = np.mean(ts_traj_delay[:,:,0],axis=0) #operator
cil = np.percentile(ts_traj_delay[:,:,0],2.5,axis=0)
ciu = np.percentile(ts_traj_delay[:,:,0],97.5,axis=0)
y_operator.append([mean,cil,ciu])
y_data.append(tscales_T_ci[kt]/2)
# +
plt.figure(figsize=(5,5))
for kt in range(len(T_range)):
ts_traj_delay = ts_traj_T[kt]
mean = np.mean(1/ts_traj_delay[:,:,0],axis=0)
# kd = np.where(np.diff(mean)<np.min(std))[0][1]
kd = np.argmax(mean)
print(kd,delay_range[kd]*dt)
mean = np.mean(ts_traj_delay[:,:,0],axis=0)
cil = np.percentile(ts_traj_delay[:,:,0],2.5,axis=0)
ciu = np.percentile(ts_traj_delay[:,:,0],97.5,axis=0)
print(np.array([mean[kd],cil[kd],ciu[kd]])*2)
plt.plot(delay_range*dt,mean,c=colors_[kt])
plt.fill_between(delay_range*dt,cil,ciu,alpha=.5,color=colors_[kt])
plt.plot(delay_range*dt,np.ones(len(mean))*half_tscales_T_ci[kt,0],c='gray',ls='--')
plt.fill_between(delay_range*dt,np.ones(len(mean))*half_tscales_T_ci[kt,1],np.ones(len(mean))*half_tscales_T_ci[kt,2],color='gray',alpha=.2)
plt.ylim(0,30.5)
plt.xlim(0,35)
plt.xlabel(r'$\tau$',fontsize=24)
plt.ylabel(r'$t_{imp}$',fontsize=25)
plt.tight_layout()
# plt.savefig('timp_decay_rate.pdf')
plt.show()
# +
from mpl_toolkits.mplot3d import Axes3D
x = delay_range*dt
zmax=25.5
xmax=40.5
sel_x = x<xmax
offset=.5
plt.figure(figsize=(5,5))
ax = plt.subplot(projection='3d')
angles = [20,-65]
ax.view_init(angles[0],angles[1])
for kt in np.arange(len(T_range)):
y = np.zeros(x.size)-kt*offset
z_operator = y_operator[kt]
z_data = y_data[kt]
sel_zo = np.logical_and(x<xmax,z_operator[2]<zmax)
ax.plot(x[sel_zo],y[sel_zo],z_operator[0][sel_zo],c=colors_[kt])
ax.add_collection3d(plt.fill_between(x[sel_zo],z_operator[1][sel_zo],z_operator[2][sel_zo],alpha=.5,color=colors_[kt]),zs = -kt*offset,zdir='y')
ax.plot(x[sel_x],y[sel_x],np.ones(len(mean))[sel_x]*z_data[0],c='gray',ls='--')
ax.add_collection3d(plt.fill_between(x[sel_x],np.ones(len(mean))[sel_x]*z_data[1],np.ones(len(mean))[sel_x]*z_data[2],color='gray',alpha=.3),zs = -kt*offset,zdir='y')
ax.set_zlim3d(0,zmax)
ax.set_xlim3d(0,xmax)
# plt.xticks(np.arange(0,41,10))
# plt.yticks(np.arange(0.5,))
# plt.savefig('tscales_DW_3d_{}_{}.pdf'.format(angles[0],angles[1]))
plt.show()
# +
from mpl_toolkits.mplot3d import Axes3D
x = delay_range*dt
zmax=25
xmax=45
sel_x = x<xmax
offset=.5
plt.figure(figsize=(5,5))
ax = plt.subplot(projection='3d')
angles = [20,-75]
ax.view_init(angles[0],angles[1])
ax.plot(delay_idx_T*dt,np.linspace(-4,0,len(T_range))[::-1],np.zeros(len(T_range)),c='k',lw=3)
for kt in np.arange(len(T_range)):
y = np.zeros(x.size)-kt*offset
z_operator = y_operator[kt]
z_data = y_data[kt]
sel_zo = np.logical_and(x<xmax,z_operator[2]<zmax)
ax.plot(x[sel_zo],y[sel_zo],z_operator[0][sel_zo],c=colors_[kt])
ax.add_collection3d(plt.fill_between(x[sel_zo],z_operator[1][sel_zo],z_operator[2][sel_zo],alpha=.5,color=colors_[kt]),zs = -kt*offset,zdir='y')
ax.plot(x[sel_x],y[sel_x],np.ones(len(mean))[sel_x]*z_data[0],c='gray',ls='--')
ax.add_collection3d(plt.fill_between(x[sel_x],np.ones(len(mean))[sel_x]*z_data[1],np.ones(len(mean))[sel_x]*z_data[2],color='gray',alpha=.3),zs = -kt*offset,zdir='y')
ax.set_zlim3d(0,zmax)
ax.set_xlim3d(0,xmax)
# plt.savefig('tscales_DW_3d_{}_{}.pdf'.format(angles[0],angles[1]))
plt.show()
# -
# # Sup Fig eigfuns
# +
# kd=14
for kt,k_B_T in enumerate(T_range):
delay=delay_idx_T[kt]
f = h5py.File(dir_path+'DoubleWell/symbol_sequences/labels_phspace_k_B_T_{}_nseeds_1000.h5'.format(k_B_T),'r')
labels = ma.array(f['labels_phspace'],dtype=int)
centers = np.array(f['centers_phspace'])
f.close()
lcs,P = op_calc.transition_matrix(labels,delay,return_connected=True)
inv_measure = op_calc.stationary_distribution(P)
final_labels = op_calc.get_connected_labels(labels,lcs)
n_modes=2
R = op_calc.get_reversible_transition_matrix(P)
eigvals,eigvecs = op_calc.sorted_spectrum(R,k=n_modes)
eigfunctions = eigvecs.real/np.linalg.norm(eigvecs.real,axis=0)
phi2 = eigfunctions[:,1]
plt.figure()
cmax = np.max(np.abs(eigvecs[:,1].real))
plt.tricontourf(centers[:,0],centers[:,1],eigvecs[:,1].real,cmap='seismic',levels=np.linspace(-1.1*cmax,1.1*cmax,1000))
plt.xlim(-0.0005,0.0005)
plt.ylim(-0.001,0.001)
plt.colorbar(ticks = np.linspace(-0.05,0.05,9))
# plt.savefig('Koopman_eigfun_DW_k_B_T_{}.pdf'.format(k_B_T))
plt.show()
# -
# # SVD subfigure
u,s,v = np.linalg.svd(traj_matrix,full_matrices=False)
bootstrap_s = []
for k in range(100):
random_indices = np.random.randint(0,len(traj_matrix),len(traj_matrix))
_,s_,_ = np.linalg.svd(traj_matrix[random_indices],full_matrices=False)
bootstrap_s.append(s_)
print(k)
bootstrap_s = np.array(bootstrap_s)
cum_sum_boot = (np.cumsum(bootstrap_s**2,axis=1).T/np.sum(bootstrap_s**2,axis=1)).T
std = np.std(cum_sum_boot,axis=0)
plt.bar(range(len(s)),np.cumsum(s**2)/np.sum(s**2),yerr = std,capsize=5)
plt.ylim(0.95,1)
plt.plot(range(len(s)),np.cumsum(s**2)/np.sum(s**2),c='k')
plt.xticks(np.arange(7),np.arange(1,8))
# plt.xlabel('# svd modes')
# plt.ylabel('cumsum(s^2)')
# plt.savefig('svd_spectrum.pdf')
plt.show()
plt.imshow(-v[:2,:].T,cmap='seismic',extent=[-0.5,1.5,-0.5,6.5])
plt.xticks([0,1],['$u_1$','$u_2$'],fontsize=12)
plt.yticks(np.arange(0,7),np.arange(-3,4)[::-1])
plt.colorbar()
# plt.savefig('svd_modes.pdf')
plt.show()
| DoubleWell/Fig3_B,C,D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#snake game
import turtle
import time
import random
delay = 0.2
score = 0
high_score = 0
#screen setup
wn = turtle.Screen()
wn.title("Snake Game By @DeependraYadav")
wn.bgcolor("green")
wn.setup(width=600, height=600)
wn.tracer(0)
#snake head
head = turtle.Turtle()
head.speed(0)
head.shape("square")
head.color("black")
head.penup()
head.goto(0, 0)
head.direction = "stop"
#snake food
food = turtle.Turtle()
food.speed(0)
food.shape("circle")
food.color("red")
food.penup()
food.goto(0, 100)
segments = []
pen = turtle.Turtle()
pen.speed(0)
pen.shape("square")
pen.color("white")
pen.penup()
pen.hideturtle()
pen.goto(0, 260)
pen.write("Score: 0 High Score: 0", align="center", font=("Courier", 24, "normal"))
#functions
def go_up():
if head.direction != "down":
head.direction = "up"
def go_down():
if head.direction != "up":
head.direction = "down"
def go_left():
if head.direction != "right":
head.direction = "left"
def go_right():
if head.direction != "left":
head.direction = "right"
def move():
if head.direction == "up":
y = head.ycor()
head.sety(y + 20)
if head.direction == "down":
y = head.ycor()
head.sety(y - 20)
if head.direction == "right":
x = head.xcor()
head.setx(x + 20)
if head.direction == "left":
x = head.xcor()
head.setx(x - 20)
#keyboard binding
wn.listen()
wn.onkeypress(go_up, "w")
wn.onkeypress(go_down, "s")
wn.onkeypress(go_left, "a")
wn.onkeypress(go_right, "d")
#main game loop
while True:
wn.update()
if head.xcor()>290 or head.xcor()<-290 or head.ycor()>290 or head.ycor()<-290 :
time.sleep(1)
head.goto(0,0)
head.direction="stop"
#hide the segments list
for segment in segments:
segment.goto(1000,1000)
#clear the segment
segments.clear()
score=0
pen.clear()
pen.write("Score: {} High Score: {}".format(score,high_score), align="center", font=("Courier", 24, "normal"))
if head.distance(food) < 20:
x = random.randint(-290, 290)
y = random.randint(-290, 290)
food.goto(x, y)
new_segment = turtle.Turtle()
new_segment.speed(0)
new_segment.color("grey")
new_segment.shape("square")
new_segment.penup()
segments.append(new_segment)
score+= 10
if score > high_score:
high_score= score
pen.clear()
pen.write("Score: {} High Score: {}".format(score,high_score), align="center", font=("Courier", 24, "normal"))
for index in range(len(segments) - 1, 0, -1):
x = segments[index - 1].xcor()
y = segments[index - 1].ycor()
segments[index].goto(x, y)
if len(segments) > 0:
x = head.xcor()
y = head.ycor()
segments[0].goto(x, y)
move()
for segment in segments :
if segment.distance(head)<20 :
time.sleep(1)
head.goto(0,0)
head.direction = "stop"
#hide the segments
for segment in segments :
segment.goto(1000,1000)
#clear the segments
segments.clear()
score=0
pen.clear()
pen.write("Score: {} High Score: {}".format(score,high_score), align="center", font=("Courier", 24, "normal"))
time.sleep(delay)
wn.mainloop()
# -
| Snake Game.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import tensorflow as tf
with tf.name_scope('hidden') as scope:
a = tf.constant(5, name='alpha')
W = tf.Variable(tf.random_uniform([1, 2], -1.0, 1.0), name='weights')
b = tf.Variable(tf.zeros([1]), name='biases')
# -
# Train the model, and also write summaries.
# Every 10th step, measure test-set accuracy, and write test summaries
# All other steps, run train_step on training data, & add training summaries
def feed_dict(train):
"""Make a TensorFlow feed_dict: maps data onto Tensor placeholders."""
if train or FLAGS.fake_data:
xs, ys = mnist.train.next_batch(100, fake_data=FLAGS.fake_data)
k = FLAGS.dropout
else:
xs, ys = mnist.test.images, mnist.test.labels
k = 1.0
return {x: xs, y_: ys, keep_prob: k}
for i in range(FLAGS.max_steps):
if i % 10 == 0: # Record summaries and test-set accuracy
summary, acc = sess.run([merged, accuracy], feed_dict=feed_dict(False))
test_writer.add_summary(summary, i)
print('Accuracy at step %s: %s' % (i, acc))
else: # Record train set summaries, and train
if i % 100 == 99: # Record execution stats
run_options = tf.RunOptions(trace_level=tf.RunOptions.FULL_TRACE)
run_metadata = tf.RunMetadata()
summary, _ = sess.run([merged, train_step],
feed_dict=feed_dict(True),
options=run_options,
run_metadata=run_metadata)
train_writer.add_run_metadata(run_metadata, 'step%d' % i)
train_writer.add_summary(summary, i)
print('Adding run metadata for', i)
else: # Record a summary
summary, _ = sess.run([merged, train_step], feed_dict=feed_dict(True))
train_writer.add_summary(summary, i)
| crackingcode/day6/cc_tf_day6_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import Linear_Regression as lr # Custom created module implements Linear regression using Gradient Descent
# # Single variable data
# Read Data
df = pd.read_csv('RegData.csv', header=None)
df.head()
# Prepare data as numpy arrays
X = np.atleast_2d(df[0].to_numpy()).T
y = df[1].to_numpy()
y = np.atleast_2d(y).T
# Initialize object from class Linear regression and solve
# Vanilla Batch GD
lr_batch_GD = lr.Linear_Regression(X, y)
theta = lr_batch_GD.fit(solver="batch", alpha=1e-3)
lr_batch_GD.show_summary()
lr_batch_GD.plot_LR_2D()
lr_batch_GD.plot_MSE()
# # Multivariable data
# Multivarialble LR
df_mv = pd.read_csv('resources/MultipleLR.csv', header=None)
df_mv.head()
# Prepare data as numpy arrays
X = df_mv.iloc[:, 0:-1].to_numpy()
y = df_mv.iloc[:, -1].to_numpy()
y = np.atleast_2d(y).T
lr_batch_multiV = lr.Linear_Regression(X, y)
theta = lr_batch_multiV.fit(alpha=1e-4)
lr_batch_multiV.show_summary()
lr_batch_multiV.plot_MSE()
| Optimization in ML/Gradient_descent/Practical_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pattern matching in spaCy
#
# Looking at the token matcher and phrase matcher as well as some tricks to speed up the matcher.
#
# This notebook is based on https://spacy.io/usage/rule-based-matching, which also goes into the dependency matcher and the entity ruler.
import spacy
from spacy.matcher import Matcher
from spacy.matcher import PhraseMatcher
nlp = spacy.load("en_core_web_sm")
# ## Matching on tokens
#
# Here we define patterns by using a dicitonary for each token. The following patterns matches 'iPhone X':
#
#
# ```json
# [{"TEXT": "iPhone"}, {"TEXT": "X"}]),
# ```
#
# Instead of accessing the text you can access many of the features on a token, the following matches "2018 FIFA World Cup":
#
# ```json
# [{"IS_DIGIT": True}, {"LOWER": "fifa"}, {"LOWER": "world"}, {"LOWER": "cup"}]
# ```
#
# You can access parts of speech and lemmas:
#
# ```json
# [{"LEMMA": "love", "POS": "VERB"}, {"POS": "NOUN"}]
# ```
#
# And use some Kleene operators (possible values are "!", "?", "*" and "+", where "!" is negation, as in, no match):
#
# ```json
# [{"LEMMA": "buy"}, {"POS": "DET", "OP": "?"}, {"POS": "NOUN"}]
# ```
# +
class SimpleMatcher(object):
"""A simple matcher that you give a list of named pattern."""
def __init__(self, patterns):
self.matcher = Matcher(nlp.vocab)
for pattern_name, pattern in patterns:
self.matcher.add(pattern_name, [pattern])
def run(self, sentence):
doc = nlp(sentence)
spacy.displacy.render(doc, options={"word_spacing": 30, "distance": 100})
matches = self.matcher(doc)
for match_id, start, end in matches:
pattern_name = self.matcher.vocab.strings[match_id]
print(f"{pattern_name:12} {start:2} {end:2} [{doc[start:end]}]")
patterns = [
("ROOT", [{"DEP": "ROOT"}]),
("IPHONE", [{"TEXT": "iPhone"},
{"TEXT": "X"}]),
("WORLD_CUP", [{"IS_DIGIT": True},
{"LOWER": "fifa"},
{"LOWER": "world"},
{"LOWER": "cup"},
{"IS_PUNCT": True}]),
("LOVE_THING", [{"LEMMA": "love", "POS": "VERB"},
{"POS": "NOUN"}]),
("BUY_STUFF", [{"LEMMA": "buy"},
{"POS": "DET", "OP": "?"},
{"POS": "NOUN"}])
]
matcher = SimpleMatcher(patterns)
# -
matcher.run("Upcoming iPhone X release date leaked")
matcher.run("2018 FIFA World Cup: France won")
matcher.run("I loved dogs but now I love cats more.")
matcher.run("I bought a smartphone. Now I am buying apps.")
# ## Matching on phrases
#
# https://spacy.io/usage/rule-based-matching#phrasematcher
#
# If you need to match large terminology lists, you can also use the PhraseMatcher and create Doc objects instead of token patterns, which is much more efficient overall. The Doc patterns can contain single or multiple tokens.
# +
# With attr="LOWER" you do a full lowercase match, without it we match on the text by default
# Note that you cannot lower case the "D.C" part because it changes the tokenization, so using
# lower is of limited value.
pmatcher = PhraseMatcher(nlp.vocab, attr="LOWER")
terms = ["barack obama", "<NAME>", "washington, D.C."]
# Only run nlp.make_doc to speed things up
patterns = [nlp.make_doc(text) for text in terms]
pmatcher.add("TerminologyList", patterns)
# same here, use make_doc if you do not need anything beyond tokenization
doc = nlp.make_doc(
"German Chancellor <NAME> and US President <NAME> "
"converse in the Oval Office inside the White House in Washington, D.C.")
matches = pmatcher(doc)
for match_id, start, end in matches:
span = doc[start:end]
print(start, end, span.text)
# -
# ### Some speed bench marks
#
# Using `make_doc()` gives a speed boost. For an additional boost, you can also use the `nlp.tokenizer.pipe()` method, which will process the texts as a stream. This boost is not as spectacular as bypassing the entire piple line. In one run using the code below the elapsed time (wall time) went from 1.24s to 3.58 ms to 2.70 ms, actual values will vary each time you run it.
LOTS_OF_TERMS = ["<NAME>", "<NAME>", "Washington, D.C."] * 100
# %%time
patterns = [nlp(term) for term in LOTS_OF_TERMS]
# %%time
patterns = [nlp.make_doc(term) for term in LOTS_OF_TERMS]
# %%time
patterns = list(nlp.tokenizer.pipe(LOTS_OF_TERMS))
| code/week4-spacy/spacy-matching.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py37torch190
# language: python
# name: py37torch190
# ---
# + [markdown] jupyter={"outputs_hidden": true} tags=[]
# ## **Stage 2** on **miniImageNet**:Ablation Studies results
# #### Note: This scripts shows the results of our baseline, which is SEGA **without semantic using** and just with AttentionBasedBlock from DynamicFSL(Gidaris&Komodakis, CVPR 2018)
# +
import sys
import torch
sys.path.append("..")
from traincode import train_stage1,train_stage2
from testcode import test
from args_config_train import argparse_config_train
from args_config_test import argparse_config_test
# -
# 5W1S
# + tags=[]
opt = argparse_config_train() # refer to args_config_train.py for details of auguments
opt.save_path = './experiments/miniimagenet_stage2_5W1S_transfer_base_weight'
opt.embnet_pretrainedandfix = True
opt.pretrian_embnet_path = './experiments/miniimagenet_stage1/best_model.pth'
opt.gpu = '0'
opt.network = 'ResNet12'
opt.head = 'SEGA'
opt.weight_generator_type = 'transfer_base_weight'
# opt.semantic_path = '/data/FSLDatasets/MiniImagenet/label2vec_glove_miniimagenet.npy'
opt.nKall = 64
opt.nKbase = 64
opt.train_query = 6
opt.nTestBase = 5*opt.train_query
opt.epoch_size = 600
opt.dataset = 'miniImageNet'
opt.avg_pool = True
opt.nfeat = 640
opt.val_episode = 1000
opt.num_epoch = 20
opt.episodes_per_batch = 8
opt.milestones = [5,10,15,20]
opt.lambdalr = [1.0,0.5,0.1,0.05]
opt.train_way, opt.train_shot = opt.test_way, opt.val_shot = [5,1]
train_stage2(opt)
torch.cuda.empty_cache()
# + tags=[]
opt = argparse_config_test() # refer to args_config_test.py for details of auguments
opt.load = './experiments/miniimagenet_stage2_5W1S_transfer_base_weight/best_model.pth'
opt.gpu = '0'
opt.network = 'ResNet12'
opt.head = 'SEGA'
opt.weight_generator_type = 'transfer_base_weight'
# opt.semantic_path = '/data/FSLDatasets/MiniImagenet/label2vec_glove_miniimagenet.npy'
opt.nKall = 64
opt.dataset = 'miniImageNet'
opt.avg_pool = True
opt.nfeat = 640
opt.way, opt.shot = [5,1]
opt.query=50
opt.episode=5000
test(opt)
torch.cuda.empty_cache()
# -
# 5W5S
# + tags=[]
opt = argparse_config_train() # refer to args_config_train.py for details of auguments
opt.save_path = './experiments/miniimagenet_stage2_5W5S_transfer_base_weight'
opt.embnet_pretrainedandfix = True
opt.pretrian_embnet_path = './experiments/miniimagenet_stage1/best_model.pth'
opt.gpu = '0'
opt.network = 'ResNet12'
opt.head = 'SEGA'
opt.weight_generator_type = 'transfer_base_weight'
# opt.semantic_path = '/data/FSLDatasets/MiniImagenet/label2vec_glove_miniimagenet.npy'
opt.nKall = 64
opt.nKbase = 64
opt.train_query = 6
opt.nTestBase = 5*opt.train_query
opt.epoch_size = 600
opt.dataset = 'miniImageNet'
opt.avg_pool = True
opt.nfeat = 640
opt.val_episode = 1000
opt.num_epoch = 20
opt.episodes_per_batch = 8
opt.milestones = [5,10,15,20]
opt.lambdalr = [1.0,0.5,0.1,0.05]
opt.train_way, opt.train_shot = opt.test_way, opt.val_shot = [5,5]
train_stage2(opt)
torch.cuda.empty_cache()
# + tags=[]
opt = argparse_config_test() # refer to args_config_test.py for details of auguments
opt.load = './experiments/miniimagenet_stage2_5W5S_transfer_base_weight/best_model.pth'
opt.gpu = '0'
opt.network = 'ResNet12'
opt.head = 'SEGA'
opt.weight_generator_type = 'transfer_base_weight'
# opt.semantic_path = '/data/FSLDatasets/MiniImagenet/label2vec_glove_miniimagenet.npy'
opt.nKall = 64
opt.dataset = 'miniImageNet'
opt.avg_pool = True
opt.nfeat = 640
opt.way, opt.shot = [5,5]
opt.query=50
opt.episode=5000
test(opt)
torch.cuda.empty_cache()
# -
| scripts/01_miniimagenet_stage2_the_ablation_studies.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyCharm (pycon2015_tutorial322)
# language: python
# name: pycharm-cef1f773
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Imports
# + pycharm={"name": "#%%\n"}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from IPython.core.display import display
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Load Data
# + pycharm={"name": "#%%\n"}
df = pd.read_csv('../Data/combined_data_cleaned.csv')
# + pycharm={"name": "#%%\n"}
df.head(2)
# + pycharm={"name": "#%%\n"}
df.info()
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Explore Data
# + [markdown] pycharm={"name": "#%% md\n"}
# #### Numeric Data
# Overall rating, vote, and price.
# + pycharm={"name": "#%%\n"}
df.describe()
# + pycharm={"name": "#%%\n"}
# overall rating
labels, counts = np.unique(df.overall, return_counts=True)
plt.bar(labels, counts, align='center')
plt.xlabel('Rating')
plt.ylabel('Count')
plt.show()
# + [markdown] pycharm={"name": "#%% md\n"}
# The data is heavily skewed towards 5-star reviews.
# + pycharm={"name": "#%%\n"}
# Some of the prices appear to be pretty high for groceries; but apparently valid.
df[df['price'] > 950].sort_values('price', ascending=False).head(2)
# + pycharm={"name": "#%%\n"}
df.groupby('main_cat')[['price']].agg(['mean', 'count'])
# + pycharm={"name": "#%%\n"}
# Recommendation system may get skewed using price because of the non normal distribution.
df.price.hist(bins=30)
plt.xlabel('Prices')
plt.ylabel('Item Count')
plt.title('Distribution of prices')
plt.show()
# + pycharm={"name": "#%%\n"}
# Very skewed. May need to narrow the price range in the future.
df.price[df['price'] < 100].describe()
# + pycharm={"name": "#%%\n"}
df.price[df['price'] < 100].hist(bins=30)
# + pycharm={"name": "#%%\n"}
# vote has a wide standard deviation and is also highly skewed.
df.vote.hist(bins=30)
plt.xlabel('Votes')
plt.ylabel('Item Count')
plt.title('Distribution of votes')
plt.show()
# + pycharm={"name": "#%%\n"}
df.vote[df['vote'] < 25].hist(bins=30)
# -
# #### Boolean Data
# + pycharm={"name": "#%%\n"}
labels, counts = np.unique(df.verified, return_counts=True)
plt.bar(labels, counts, align='center')
plt.xticks([0, 1])
plt.xlabel('Rating')
plt.ylabel('Count')
plt.show()
# + [markdown] pycharm={"name": "#%% md\n"}
# #### Heat Map
# None of the features are correlated.
# + pycharm={"name": "#%%\n"}
sns.heatmap(round(df.corr(), 2), annot=True)
plt.title('Heat Map')
plt.show()
# + [markdown] pycharm={"name": "#%% md\n"}
# #### Analyze Categories/Objects
# + pycharm={"name": "#%%\n"}
df_objects = df.select_dtypes(include='object')
dfo_unique = pd.DataFrame(df_objects.nunique()).reset_index()
dfo_unique.columns = ['Features', 'Number of Categories']
dfo_unique.sort_values(by='Number of Categories', ascending=False)
dfo_unique
# + [markdown] pycharm={"name": "#%% md\n"}
# Except for the main_cat, there are too many unique categories for several traditional types of exploratory data analysis to be helpful.
# For example, one hot encoding using pd.get_dummies(df) runs into memory errors even when the biggest features are dropped from the analysis.
# Therefore, in later notebooks will use options created specifically for recommendation systems.
# + pycharm={"name": "#%%\n"}
for col in df_objects.columns:
counts = df[col].value_counts()
print(f'Most common {col}:\n{counts.head(3)}\n')
# + [markdown] pycharm={"name": "#%% md\n"}
# Possible issue #1: The most common details feature is empty.
# Earlier, when cleaning the data there were checks for empty lists; however, not for empty dictionaries.
# However, after further analysis the details feature is still useful because there are more than a million rows and only about 24k are empty.
# -
# Possible issue #2: The most common information in the summary column might be a duplicate of the overall column.
# This could cause problems because the overall column is the target feature.
# + pycharm={"name": "#%%\n"}
df['summary'].value_counts()
# + pycharm={"name": "#%%\n"}
df['overall'].value_counts()
# + pycharm={"name": "#%%\n"}
# Yes, these columns do have a lot of overlap. Therefore, dropping summary.
df.drop('summary', axis=1, inplace=True)
# + [markdown] pycharm={"name": "#%% md\n"}
# #### Grouping by target
#
# Grouping by 'overall' rating to show the average highest and lowest product ratings.
# + pycharm={"name": "#%%\n"}
non_target_features = df.columns.drop('overall')
for feature in non_target_features:
overall_grouping_avg = df.groupby(feature)[['overall']].mean()
display(overall_grouping_avg.sort_values('overall', ascending=False))
# + [markdown] pycharm={"name": "#%% md\n"}
# The above also shows that most of the features have at least some distribution of the product's rating.
# The main exception is the verified boolean.
# + pycharm={"name": "#%%\n"}
df.info()
# + [markdown] pycharm={"name": "#%% md\n"}
# Usually, would add default values for nulls in several of the columns either here, or earlier in the process.
# However, for the recommendation systems it is expected that a significant amount of the data could be null.
# Therefore, leaving this as is for now.
# -
# ## Save Data
# + pycharm={"name": "#%%\n"}
df.to_csv('../Data/eda_data.csv', index=False)
# -
# ## Summary
# - There are only 3 numeric (overall, price, and vote) and 1 boolean (verified) features in the data set.
# - All of these are skewed toward one value or range of values.
# - Overall ratings are heavily skewed towards 5.
# - Both lower values for price and votes are most frequent.
# - Majority of reviews are verified.
# - The majority of the features are categories/objects/text.
# - Dropped the summary feature because it contained a lot of duplicate data with the target feature of overall rating.
# - Most of the features have at least some distribution of the product's rating. The main exception is the verified boolean.
# - There are so many categories it wasn't possible to do one hot encoding because of memory issues.
# - However, this didn't matter because will use processes designed specifically for recommendation systems to overcome these limitations.
| Notebooks/Exploratory_Data_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: deepTFM
# language: python
# name: deeptfm
# ---
# ### SAME EXPERIMENT ON COLAB: https://colab.research.google.com/drive/1aDHkE02n-ahCBoQxcTCp4AldgTpQ13r3#scrollTo=Fg7NQp8Hw8LG
# +
import torch
import torchvision
import shutil
batch_size= 32
train_loader = torch.utils.data.DataLoader(
torchvision.datasets.MNIST('../datasets/mnist', train=True, download=True,
transform=torchvision.transforms.Compose([
torchvision.transforms.Resize([32, 32]),
torchvision.transforms.ToTensor(),
#torchvision.transforms.Normalize(
# (0.5,), (0.5,))
])),
batch_size=batch_size, shuffle=True)
test_loader = torch.utils.data.DataLoader(
torchvision.datasets.MNIST('../datasets/mnist', train=False, download=True,
transform=torchvision.transforms.Compose([
torchvision.transforms.Resize([32, 32]),
torchvision.transforms.ToTensor(),
#torchvision.transforms.Normalize(
# (0.5,), (0.5,))
])),
batch_size=16, shuffle=False, drop_last= True)
# +
import torch
from torch.nn import functional as F
device_ = 'cuda'
for X, _ in train_loader:
#X= torch.randint(0, 256, (1,1,32,32))/255
X= X.to(device_)
Ht = torch.sigmoid(torch.randn((1, 1, 32, 32))).to(device_)
kernel = torch.sigmoid(torch.randn((1, 1, 5, 5))).to(device_)
A1= F.conv2d(Ht, kernel, padding= 2)*X
yt= F.conv2d(A1, kernel, padding= 2)
print(f'X range : {X.min()}, {X.max()}')
print(f'Ht range : {Ht.min()}, {Ht.max()}')
print(f'kernel range : {kernel.min()}, {kernel.max()}')
print(f'A1 range (conv(Ht, kernel)) : {A1.min()}, {A1.max()}')
print(f'yt range (conv(A1, kernel)) : {yt.min()}, {yt.max()}')
break
# -
# !/nvcc --version
# !cat /usr/include/cudnn.h | grep CUDNN_MAJOR -A 2
# !nvidia-smi
torch.__version__
| aim2/support_notebooks/old/Error_PyTorch_CUDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import os
# # Change images to numbers
# +
path = 'images/month2/'
dir_contents = np.array(os.listdir(path))
############################################################
# Make sure we don't trip over ourselves renaming stuff
############################################################
if ('1.jpg' in dir_contents):
switch = 'a'
else:
switch = ''
############################################################
# Rename Files
############################################################
for i in np.arange(0,len(dir_contents)):
os.rename(path+str(dir_contents[i]),path+str(i)+switch+'.jpg')
len(dir_contents)
# -
# # Create HTML page for month
# +
import math
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import os
#groups = ['pst','month1','month2','month3','month4']
groups = ['month5']
# +
#Select group
os.chdir('/home/pj/pj_blog/pics/')
f = open('example_pics.html','r')
text = f.read()
f.close()
# -
for i in np.arange(0,len(groups)):
group = groups[i]
base_img_folder = '/home/pj/pj_blog/images/'+group+'/'
albums = os.listdir(base_img_folder)
newline = '\t\t\t\t\t\t\t\t\t\t\t\t<div class="col-4"><span class="image fit"><img src="pathzzz" alt="" class="rotateimg0" /></span></div>\n'
big0 = text.split('<!-- Content -->')[0]
big2 = text.split('<!-- Content -->')[2]
first = text.split('<!-- Content -->')[1].split(newline)[0]
last = text.split('<!-- Content -->')[1].split(newline)[1]
if group[0] == 'm':
big0 = big0.replace('Month # - - - - - (Mon 20 - Mon 20)', 'Month '+group[-1])
else:
big0 = big0.replace('Month # - - - - - (Mon 20 - Mon 20)', group)
whole = big0
for sub_album in albums:
images = os.listdir(base_img_folder+sub_album+'/')
listr = ''
for img in images:
listr += (newline.replace('pathzzz',base_img_folder+sub_album+'/'+img))
first1 = first.replace('Subheading',sub_album)
whole += (first1+listr+last)
whole += big2
os.chdir('/home/pj/pj_blog/pics/')
f2 = open(group+'_pics.html','w')
f2.write(whole)
f2.close()
base_img_folder
'./images/'+grou
big0.replace('>Month # - - - - - (Mon 20 - Mon 20)', 'Month '+group[-1])
# +
whole = big0
for sub_album in albums:
images = os.listdir(base_img_folder+sub_album+'/')
listr = ''
for img in images:
listr += (newline.replace('pathzzz',base_img_folder+sub_album+'/'+img))
first1 = first.replace('Subheading',sub_album)
whole += (first1+listr+last)
whole += big2
# -
os.chdir('/home/pj/pj_blog/pics/')
f2 = open(group+'_pics.html','w')
f2.write(whole)
f2.close()
| Change Image Names.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="84UkIMuNKYII"
# # Naive Bayes for Text Classification
#
# - **source:**
# https://towardsdatascience.com/algorithms-for-text-classification-part-1-naive-bayes-3ff1d116fdd8
#
# - **original:** https://colab.research.google.com/drive/1bEfgKPZQerDQgDv5kEkyVfStXZuAVAwP#scrollTo=G5M3NkK6SnPF
# + colab={} colab_type="code" id="g5s_dj3WLkJk"
import pandas
import numpy
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.naive_bayes import MultinomialNB
from sklearn.metrics import roc_auc_score
from sklearn.preprocessing import LabelEncoder
# + colab={} colab_type="code" id="C79Ne8GgLAZW"
url = "https://raw.githubusercontent.com/shantanuo/naive_bayes_for_text_classification/master/spam.csv"
data = pandas.read_csv(url, encoding="latin-1")
data.head()
# + colab={} colab_type="code" id="0YfxfrCsLpnG"
for i in range(2, 5):
column_name = 'Unnamed: ' + str(i)
data = data.drop(column_name, axis=1)
data.head()
# + colab={} colab_type="code" id="AkeKaMslMjG2"
data.columns = ['Target', 'Text']
data['Target'] = LabelEncoder().fit_transform(data['Target'])
data.head()
# + colab={} colab_type="code" id="mAvJCjKLPpRl"
train, test = train_test_split(data, test_size=0.2, random_state=0)
train['Text'].head()
# + colab={} colab_type="code" id="Cilb4GHTRZAI"
countVectorizer = CountVectorizer()
X_train_vectorized = countVectorizer.fit_transform(train['Text'])
ndf = pandas.SparseDataFrame(
X_train_vectorized.toarray(), columns=countVectorizer.get_feature_names()
)
ndf.iloc[0][ndf.iloc[0] > 0]
# + colab={} colab_type="code" id="EI9M7_nbSJH5"
naive_bayes = MultinomialNB(alpha=0.1)
naive_bayes.fit(X_train_vectorized, train['Target'])
# + colab={} colab_type="code" id="Acw93lI8dRbp"
y = naive_bayes.predict(countVectorizer.transform(test['Text']))
# + colab={} colab_type="code" id="XQUWHUuvdu0f"
predictedData = {'Target': y, 'Text': test['Text']}
predictedDataFrame = pandas.DataFrame(predictedData)
predictedDataFrame.head(10)
# + colab={} colab_type="code" id="q4nOoW4OejM7"
predictedDataFrame['Target'] = predictedDataFrame['Target'].replace(0, 'ham')
predictedDataFrame['Target'] = predictedDataFrame['Target'].replace(1, 'spam')
predictedDataFrame.head(10)
# + colab={} colab_type="code" id="Um_7XmcHhGX5"
predictedDataFrame[predictedDataFrame.Target == 'ham'].head(10)
# + colab={} colab_type="code" id="1ZTwogi9APRV"
predictedDataFrame[predictedDataFrame.Target == 'spam'].head(10)
# + colab={} colab_type="code" id="OelDVODPhmwi"
roc_auc_score(test['Target'], y)
# + colab={} colab_type="code" id="4rZiPKfSnEOw"
my_url = "https://raw.githubusercontent.com/hentai-lab/Machine-Learning/master/docs/my_own_spam_data.csv"
my_data = pandas.read_csv(my_url, encoding="latin-1")
my_data['Target'] = LabelEncoder().fit_transform(my_data['Target'])
my_data = my_data.sample(frac=1)
my_data
# + colab={} colab_type="code" id="CZ9SEIDR0i9O"
my_y = naive_bayes.predict(countVectorizer.transform(my_data['Text']))
# + colab={} colab_type="code" id="2IfUyHsJ0rW8"
my_predictedData = {'Target': my_y, 'Text': my_data['Text']}
my_predictedDataFrame = pandas.DataFrame(my_predictedData)
my_predictedDataFrame
# + colab={} colab_type="code" id="dCZJeRc10_FF"
my_predictedDataFrame['Target'] = my_predictedDataFrame['Target'].replace(0, 'ham')
my_predictedDataFrame['Target'] = my_predictedDataFrame['Target'].replace(1, 'spam')
my_predictedDataFrame
# + colab={} colab_type="code" id="xJ4NfvLi1RW7"
roc_auc_score(my_data['Target'], my_y)
# + colab={} colab_type="code" id="lmGHvGC4_mOY"
my_X_train_vectorized = countVectorizer.fit_transform(my_data['Text'])
ndf = pandas.SparseDataFrame(
my_X_train_vectorized.toarray(), columns=countVectorizer.get_feature_names()
)
ndf.iloc[0][ndf.iloc[0] > 0]
naive_bayes.fit(my_X_train_vectorized, my_data['Target'])
# + colab={} colab_type="code" id="J5JF<PASSWORD>"
input_text = [input('Enter your message: ')]
input_target = input('Is it spam or not?: ')
if input_target == 'no' or input_target == 'No' or input_target == 'NO':
input_target = 0
elif input_target == 'yes' or input_target == 'Yes' or input_target == 'YES' or input_target == 'yEs' or input_target == 'yeS':
input_target = 1
another_y = naive_bayes.predict(countVectorizer.transform(input_text))
another_predictedData = [['User', input_target, input_text[0]], ['Naive Bayes', another_y, input_text[0]]]
another_predictedData = pandas.DataFrame(another_predictedData, columns=['Source', 'Target', 'Text'])
another_predictedData['Target'] = another_predictedData['Target'].replace(0, 'ham')
another_predictedData['Target'] = another_predictedData['Target'].replace(1, 'spam')
another_predictedData
| notebooks/IA/Naive_Bayes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.12 ('nanodet')
# language: python
# name: python3
# ---
# +
import os
import time
import cv2
from nanodet.data.transform import Pipeline
from nanodet.util import cfg, load_config
from nanodet.util.path import mkdir
image_ext = [".jpg", ".jpeg", ".webp", ".bmp", ".png"]
video_ext = ["mp4", "mov", "avi", "mkv"]
def get_image_list(path):
image_names = []
for maindir, subdir, file_name_list in os.walk(path):
for filename in file_name_list:
apath = os.path.join(maindir, filename)
ext = os.path.splitext(apath)[1]
if ext in image_ext:
image_names.append(apath)
return image_names
config = "/home/tao/Github/nanodet_custom/config/gray_stereo.yml"
path = "/home/tao/Pictures/real_data_stereo_ann/data"
save_result = True
local_rank = 0
load_config(cfg, config)
pipeline = Pipeline(cfg.data.train.pipeline, cfg.data.train.keep_ratio)
current_time = time.localtime()
if os.path.isdir(path):
files = get_image_list(path)
else:
files = [path]
files.sort()
for image_name in files:
img_info = {"id": 0}
if isinstance(image_name, str):
img_info["file_name"] = os.path.basename(image_name)
img = cv2.imread(image_name)
else:
img_info["file_name"] = None
height, width = img.shape[:2]
img_info["height"] = height
img_info["width"] = width
meta = dict(img_info=img_info, raw_img=img, np_img=img, img=img)
meta = pipeline(None, meta, cfg.data.train.input_size)
if save_result:
save_folder = os.path.join(
cfg.save_dir, time.strftime("%Y_%m_%d_%H_%M_%S", current_time)
)
mkdir(local_rank, save_folder)
save_file_name = os.path.join(save_folder, os.path.basename(image_name))
result_image = meta["img"]
cv2.imwrite(save_file_name, result_image)
print(result_image.shape)
| demo/test_pipeline.ipynb |