code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <small><i>This notebook was prepared by [<NAME>](http://donnemartin.com). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).</i></small>
# # Solution Notebook
# ## Problem: Find the start of a linked list loop.
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# ## Constraints
#
# * Is this a singly linked list?
# * Yes
# * Can we assume we are always passed a circular linked list?
# * No
# * Can we assume we already have a linked list class that can be used for this problem?
# * Yes
# ## Test Cases
#
# * Empty list -> None
# * Not a circular linked list -> None
# * One element
# * Two or more elements
# * Circular linked list general case
# ## Algorithm
#
# * Use two pointers i, j, initialized to the head
# * Increment i and j until they meet
# * j is incremented twice as fast as i
# * If j's next is None, we do not have a circular list
# * When i and j meet, move j to the head
# * Increment i and j one node at a time until they meet
# * Where they meet is the start of the loop
#
# Complexity:
# * Time: O(n)
# * Space: O(1)
# ## Code
# %run ../linked_list/linked_list.py
class MyLinkedList(LinkedList):
def find_loop_start(self):
if self.head is None or self.head.next is None:
return
i = self.head
j = self.head
i = i.next
j = j.next.next
# Increment i and j until they meet
# j is incremented twice as fast as i
while j != i:
i = i.next
if j is None or j.next is None:
return
j = j.next.next
# When i and j meet, move j to the head
j = self.head
# Increment i and j one node at a time until
# they meet, which is the start of the loop
while j != i:
i = i.next
j = j.next
return i.data
# ## Unit Test
# +
# %%writefile test_find_loop_start.py
from nose.tools import assert_equal
class TestFindLoopStart(object):
def test_find_loop_start(self):
print('Test: Empty list')
linked_list = MyLinkedList()
assert_equal(linked_list.find_loop_start(), None)
print('Test: Not a circular linked list: One element')
head = Node(1)
linked_list = MyLinkedList(head)
assert_equal(linked_list.find_loop_start(), None)
print('Test: Not a circular linked list: Two elements')
linked_list.append(2)
assert_equal(linked_list.find_loop_start(), None)
print('Test: Not a circular linked list: Three or more elements')
linked_list.append(3)
assert_equal(linked_list.find_loop_start(), None)
print('Test: General case: Circular linked list')
node10 = Node(10)
node9 = Node(9, node10)
node8 = Node(8, node9)
node7 = Node(7, node8)
node6 = Node(6, node7)
node5 = Node(5, node6)
node4 = Node(4, node5)
node3 = Node(3, node4)
node2 = Node(2, node3)
node1 = Node(1, node2)
node0 = Node(0, node1)
node10.next = node3
linked_list = MyLinkedList(node0)
assert_equal(linked_list.find_loop_start(), 3)
print('Success: test_find_loop_start')
def main():
test = TestFindLoopStart()
test.test_find_loop_start()
if __name__ == '__main__':
main()
# -
# %run -i test_find_loop_start.py
| linked_lists/find_loop_start/find_loop_start_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mentoria Evolution - Exercícios Python
# https://minerandodados.com.br
# * Para executar uma célula digite **Control + enter** ou clique em **Run**.
# * As celulas para rodar script Python devem ser do tipo code.
# * Crie células abaixo das celulas que foram escrito o enunciado das questões com as respostas.
# **Obs**: Caso de dúvidas, volte na aula anterior de Python. Não desista :)
# ## Exercícios de Fixação
# 1) Imprima a frase **"Eu sou Minerador <NAME>!"**
print ("Eu sou Minerador <NAME>")
# 2) Faça operações matemáticas de soma, subtração, multiplicação e divisão.
2 + 2
5 - 1
8 * 7
44 / 11
# 3) Imprima o tipo de um determinado do valor **10**.
type(10)
# 4) Converta o valor passado anteriormente para o tipo **float**
float(10)
# 5) Faça a conversão do valor float recebido anteriormente para o tipo **string**
str(10)
# 6) Utilizando variáveis faça:
# - a) Defina uma variável com o **seu nome** e imprima o seu valor.
# - b) Imprima o **tipo** dessa variável.
# - c) Substitua o valor da sua variável nome com a concatenação do seu valor e o seu sobrenome, depois, imprima o novo valor.
# - obs: O resultado dessa variável será Nome Sobrenome, exemplo, **<NAME>**.
# - d) Defina duas variaveis com os valores do seu email e idade com apenas uma linha de comando, depois, imprima seus valores.
nome = 'Cid'
print(nome)
type(nome)
nome = nome + ' Felipe'
print(nome)
nome, email = '<NAME>', '<EMAIL>'
print(nome)
print(email)
# 7) Utilizando funções de arredonamento faça:
# - a) Arredonde o valor 2.898
# - b) Arredonde o valor 2.4545455 e exiba apenas 4 cadas decimais
round(2.898)
round(2.4545455, 4)
# 9) Sobre os objetos Strings, faça:
# - a) Defina uma variavel com a string: "Python é uma maldição!" e imprima
# - b) Imprima o primeiro caracter da variavel.
# - c) Imprima os valores do primeiro caracter até o 6.
# - d) Faça a contagem de todos os caracteres utilizando uma única linha de comando.
# - e) Faça uma contagem de quantas vezes a palavra dados aparece na seguinte frase:
# - "dados é o novo petróleo, em Deus eu confio, para o resto me traga dados.."
# - Obs: Crie uma variável chamada "frase" com a frase acima.
# - f) Substitua o primeiro caracter da frase acima para maiúsculo
# - g) Quebre as palavras da frase separado por virgula, criando uma lista.
string = 'Python é uma maldição!'
print(string)
string[0]
string[0:6]
frase = "dados é o novo petróleo, em Deus eu confio, para o resto me traga dados.."
frase.count('dados')
frase.capitalize()
frase.split(',')
# - Ao concluir, salve seu notebook e envie suas respostas para **<EMAIL>**
| aula_03/mentoria_aula_03.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys; print(sys.version)
import os
import glob
import subprocess
import multiprocessing
import io
from collections import OrderedDict
import json
import numpy as np; print('numpy', np.__version__)
import pandas as pd; print('pandas',pd.__version__)
import allel; print('scikit-allel', allel.__version__)
import zarr; print('zarr', zarr.__version__)
import matplotlib as mpl
import statsmodels; print('statsmodels', statsmodels.__version__)
import statsmodels.api as sm
from IPython.display import display, HTML
# -
# %matplotlib notebook
mpl.rcParams['figure.facecolor'] = '#BBBBBB'
# name, fai, gtf, chroms
GENOMES = [
['AgamP4.11',
'/data/reference/Anopheles-gambiae-PEST_CHROMOSOMES_AgamP4.fa.fai',
'../datafiles/Anopheles-gambiae-PEST_BASEFEATURES_AgamP4.11.gtf',
['2R','2L','3R','3L','X']
],
['Aaegypti_L5.1',
'/data/reference/AaegL5.fa.fai',
'../datafiles/Aedes-aegypti-LVP_AGWG_BASEFEATURES_AaegL5.1.gtf',
['1','2','3']
],
]
# ## genome size
for genome_name, faifn, gtffn, chroms in GENOMES:
t = pd.read_csv(faifn, delimiter='\t', header=None,
names=['name','len','offst','linebases','linewidth'])
total_genome_len = t.loc[t['name'].isin(chroms),'len'].sum()
print(genome_name, faifn)
print(chroms)
# display(t)
print('total',total_genome_len)
# +
# TRANSFN = "../datafiles/transcript_list_{}".format(GENOME)
# tlist = pd.read_csv(TRANSFN, header=None, names=['transcript_id'])
# t = tlist['transcript_id'].str.rsplit('-', n=1, expand=True)
# t.columns = ['gene', 'splice_id']
# tlist = pd.concat((tlist,t), axis=1)
# -
for genome_name, faifn, gtffn, chroms in GENOMES:
print(genome_name, gtffn)
d = pd.read_csv(gtffn, sep='\t', comment='#', header=None,
names=['seqid',
'source',
'type',
'start',
'end',
'score',
'strand',
'phase',
'attributes'],
dtype={'seqid':str})
# total number of genes
t = d.loc[d['type']=='gene' ,:].copy(deep=True)
t['gene_id'] = t['attributes'].apply(
lambda x: dict([_.strip().split() for _ in
x.split(';') if _])['gene_id'].strip('"'))
print("total number of genes in gtf:", t['gene_id'].unique().shape[0])
# list of CDS
d = d.loc[d['type']=='CDS' ,:]
d['gene_id'] = d['attributes'].apply(
lambda x: dict([_.strip().split() for _ in
x.split(';') if _])['gene_id'].strip('"'))
d['transcript_id'] = d['attributes'].apply(
lambda x: dict([_.strip().split() for _ in
x.split(';') if _])['transcript_id'].strip('"'))
# filter to only CRHOMS
d = d.loc[d['seqid'].isin(chroms) ,:]
# cdslist = d
print('Total coding size (sum of CDS lens)',(d['end']-d['start']+1).sum())
# assert len(set(cdslist['transcript_id'])-set(tlist['transcript_id'])) == 0 # transcripts should be superset of CDS
# tcdslist = set(tlist['transcript_id']) & set(cdslist['transcript_id'])
# tcdsdf = tlist[tlist['transcript_id'].isin(tcdslist)]
# print('# trascripts with CDS:', len(tcdslist))
# num_coding_genes = tcdsdf['gene'].unique().shape[0]
# print('# genes with a transcript with a CDS:', num_coding_genes)
# ## Results summaries from json files
SETNAMES = [
'VGL-gam',
'Ag1000g-gam',
'VGL-col',
'Ag1000g-col',
'VGL-Aaeg',
]
dat_any = OrderedDict()
dat_onecall = OrderedDict()
for setname in SETNAMES:
try:
with open(setname+'_results.json','r') as fh:
dat_any[setname] = OrderedDict(json.load(fh))
except FileNotFoundError as e:
print(e)
pass
with open(setname+'_ignore-one-call_results.json','r') as fh:
dat_onecall[setname] = OrderedDict(json.load(fh))
# display(dat[setname])
# +
s = ''
for label,dat in [['any variation',dat_any],
['ignore single call variants',dat_onecall],
]:
s += '<h3>'+label+'</h3>\n'
s += '<table border="1">\n'
s += '<tr>\n'
s += '<th></th>\n'
for k,d in dat.items():
s += '<th>{:s}</th>\n'.format(k)
s += '</tr>\n'
s += '<tr>\n'
s += '<th>'+'N samples'+'</th>\n'
for k,d in dat.items():
s += '<td>{:d}</td>\n'.format(dat[k]['number of samples'])
s += '</tr>\n'
s += '<tr>\n'
s += '<th>'+'good unique targets'+'</th>\n'
for k,d in dat.items():
s += '<td>{:d}<br>{:.2f}% T<br>{:.2f}% P</td>\n'.format(
dat[k]['good unique targets'],
dat[k]['good unique targets % of total unique'],
dat[k]['good unique targets % of potential'])
s += '</tr>\n'
s += '<tr>\n'
s += '<th>'+'coding transcripts with good targets'+'</th>\n'
for k,d in dat.items():
s += '<td>{:d}<br>{:.2f}%</td>\n'.format(
dat[k]['coding transcripts w/ good targets'],
dat[k]['coding transcripts w/ good targets %'])
s += '</tr>\n'
s += '<tr>\n'
s += '<th>'+'coding genes with good targets'+'</th>\n'
for k,d in dat.items():
s += '<td>{:d}<br>{:.2f}%</td>\n'.format(
dat[k]['coding genes w/ good targets'],
dat[k]['coding genes w/ good targets %'])
s += '</tr>\n'
s += '</table>'
display(HTML(s))
# print(s)
# -
# ## filtering by probability of target being reference sequence
def good_counts(tout, num_total_unique_targets, num_coding_genes, min_refAF=None, min_p_ref=None):
"""return counts of good targets and genes derived from tout
min_refAF filter targets based on the minimum reference allele frequency for for all variants in target
thereby disregarding low frequency variants
...or...
min_p_ref filter targets based on probability of reference sequence for whole target
p_ref = the product of the reference freq for all variants in a target
~= probability of reference sequence for whole target
min_p_ref of 1 means any variants in a target will invalidate it
"""
results_dict = OrderedDict()
results_dict['total unique targets'] = num_total_unique_targets
results_dict['coding genes'] = num_coding_genes
t = tout[['gene','transcript_id','chrom','pos','nv','refAF','p_ref','Genomic location']]
if min_p_ref is not None:
good_targets = t[t['p_ref'] >= min_p_ref]
if min_refAF is not None:
good_targets = t[t['refAF'].apply(lambda x: min(x,default=1)) >= min_refAF]
# by target (not put in results_dict)
print('num potential targets (including non-unique):', t.shape[0])
print('num good targets (including non-unique):', good_targets.shape[0])
print("% good targets of potential (including non-unique): {:d}/{:d} = {:0.3f}%".format(
good_targets.shape[0], t.shape[0], 100*good_targets.shape[0]/t.shape[0]))
# by unique target
num_potential_unique_targets = t['Genomic location'].unique().shape[0]
results_dict['potential unique targets'] = num_potential_unique_targets
num_good_unique_targets = good_targets['Genomic location'].unique().shape[0]
results_dict['good unique targets'] = num_good_unique_targets
results_dict['good unique targets % of total unique'] = (100*
results_dict['good unique targets']/results_dict['total unique targets'])
results_dict['good unique targets % of potential'] = (100*
results_dict['good unique targets']/results_dict['potential unique targets'])
# by gene
num_coding_genes = results_dict['coding genes']
num_potential_genes = t['gene'].unique().shape[0]
num_good_genes = good_targets['gene'].unique().shape[0]
results_dict['coding genes w/ potential targets'] = num_potential_genes
results_dict['coding genes w/ potential targets %'] = (100*
num_potential_genes/num_coding_genes)
results_dict['coding genes w/ good targets'] = num_good_genes
results_dict['coding genes w/ good targets %'] = (100*
num_good_genes/num_coding_genes)
return results_dict
SETNAMES = [
'VGL-gam',
'Ag1000g-gam',
'VGL-col',
'Ag1000g-col',
'VGL-Aaeg',
]
# +
minRefAF_thresholds = [1, 0.995, 0.990, 0.985, 0.980, 0.5, 0]
saved_results_any = OrderedDict()
res_by_minRefAF = OrderedDict()
for setname in SETNAMES:#[1:2]:
print(setname)
# get the main target to variation results... takes a while to load
tout = pd.read_msgpack(setname+'_tout.msgpack')
# also need a few values from the 'any variation' results
with open(setname+'_results.json','r') as fh:
saved_results_any[setname] = OrderedDict(json.load(fh))
for minRefAF_threshold in minRefAF_thresholds:
if setname not in res_by_minRefAF:
res_by_minRefAF[setname] = OrderedDict()
res_by_minRefAF[setname][minRefAF_threshold] = good_counts(tout,
saved_results_any[setname]['unique location target sites'],
saved_results_any[setname]['coding genes'],
min_refAF=minRefAF_threshold)
# + run_control={"marked": true}
for k,v in res_by_minRefAF.items():
display(k)
for k2,v2 in v.items():
display(k2)
display(v2)
# -
# load the ignore-one-call results
res_onecall = OrderedDict()
for setname in SETNAMES:
with open(setname+'_ignore-one-call_results.json','r') as fh:
res_onecall[setname] = OrderedDict(json.load(fh))
res_onecall
# +
fig,axs = mpl.pyplot.subplots(2,1, squeeze=False, figsize=(6,6),
gridspec_kw={'height_ratios': [3, 1]})
barwidth = 0.25
ax = axs[0,0]
y = [res_by_minRefAF[s][0.99]['coding genes w/ good targets %'] for s in SETNAMES]
x = np.arange(0, len(y))
ax.bar(x+0*barwidth, y, width=barwidth*.9, label='allowing <1% variants')
y = [res_onecall[s]['coding genes w/ good targets %'] for s in SETNAMES]
ax.bar(x+1*barwidth, y, width=barwidth*.9, label='allowing 1-call variants')
y = [res_by_minRefAF[s][1]['coding genes w/ good targets %'] for s in SETNAMES]
ax.bar(x+2*barwidth, y, width=barwidth*.9, label='no detected variants')
ax.set_xticks(np.arange(0, len(y))+barwidth)
ax.set_xticklabels(SETNAMES)
ax.set_ylabel('% of genes with good targets')
ax.set_ylim((0,100))
ax_leg = axs[1,0]
lh,ll = ax.get_legend_handles_labels()
ax_leg.legend(lh,ll, loc='center')
ax_leg.set_axis_off()
# -
SETNAMES, LABELS = zip(*[
['VGL-gam', 'VGL\n'+r'$\mathit{An. gambiae}$'],
['Ag1000g-gam', 'Ag1000g\n'+r'$\mathit{An. gambiae}$'],
['VGL-col', 'VGL\n'+r'$\mathit{An. coluzzii}$'],
['Ag1000g-col', 'Ag1000g\n'+r'$\mathit{An. coluzzii}$'],
['VGL-Aaeg', 'VGL\n'+r'$\mathit{Ae. aegypti}$'],
])
# +
fig,axs = mpl.pyplot.subplots(2,1, squeeze=False, figsize=(6,6),
gridspec_kw={'height_ratios': [100, 1]})
barwidth = 1
ax = axs[0,0]
y1 = np.array([res_by_minRefAF[s][0.99]['coding genes w/ good targets %'] for s in SETNAMES])
y2 = np.array([res_onecall[s]['coding genes w/ good targets %'] for s in SETNAMES])
y3 = np.array([res_by_minRefAF[s][1]['coding genes w/ good targets %'] for s in SETNAMES])
x = np.arange(0, len(y1))
ax.bar(x, y1-y2, bottom=y2, width=barwidth*.8, label='allowing <1% variants', edgecolor='k', linewidth=0.5)
ax.bar(x, y2-y3, bottom=y3, width=barwidth*.8, label='allowing 1-call variants', edgecolor='k', linewidth=0.5)
ax.bar(x, y3, bottom=0, width=barwidth*.8, label='no detected variants', edgecolor='k', linewidth=0.5)
ax.set_xticks(np.arange(0, len(y)))#+barwidth)
ax.set_xticklabels([LABELS[i]+ #s.replace('-','\n')+
'\n$N='+str(res_onecall[s]['number of samples'])+'$' for i,s in enumerate(SETNAMES)])
# ax.set_xticklabels([s.replace('-','\n')+
# '\n$N='+str(res_onecall[s]['number of samples'])+'$' for s in SETNAMES])
ax.set_ylabel('% of genes with good targets')
ax.set_ylim((0,100))
ax_leg = axs[1,0]
lh,ll = ax.get_legend_handles_labels()
ax_leg.legend(lh,ll, loc='center')
ax_leg.set_axis_off()
fig.tight_layout()
fig.savefig('percent_good_genes_overview.svg')
fig.savefig('percent_good_genes_overview.png')
# -
# +
SETNAMES, LABELS = zip(*[
['VGL-gam', 'VGL\n'+r'$\mathit{An. gambiae}$'],
['Ag1000g-gam', 'Ag1000g\n'+r'$\mathit{An. gambiae}$'],
['VGL-col', 'VGL\n'+r'$\mathit{An. coluzzii}$'],
['Ag1000g-col', 'Ag1000g\n'+r'$\mathit{An. coluzzii}$'],
['VGL-Aaeg', 'VGL\n'+r'$\mathit{Ae. aegypti}$'],
])
SETNAMES, LABELS = zip(*[
['VGL-gam', 'VGL\n'+r'$\mathit{gambiae}$'],
['Ag1000g-gam', 'Ag1000g\n'+r'$\mathit{gambiae}$'],
['VGL-col', 'VGL\n'+r'$\mathit{coluzzii}$'],
['Ag1000g-col', 'Ag1000g\n'+r'$\mathit{coluzzii}$'],
['VGL-Aaeg', 'VGL\n'+r'$\mathit{aegypti}$'],
])
barwidth = .425
b2off = .3
fig,axs = mpl.pyplot.subplots(2,1, squeeze=False, figsize=(4,4),
gridspec_kw={'height_ratios': [100, 1]})
ax = axs[0,0]
y1 = np.array([res_by_minRefAF[s][0.99]['coding genes w/ good targets %'] for s in SETNAMES])
y2 = np.array([res_by_minRefAF[s][1]['coding genes w/ good targets %'] for s in SETNAMES])
x = np.arange(0, len(y1))
# ax.bar(x, y1-y2, bottom=y2, width=barwidth*.8, label='allowing <1% variants', edgecolor='k', linewidth=0.5)
ax.bar(x, y1, bottom=0, width=barwidth, label='allowing DRAs with <1% frequency', edgecolor='k', linewidth=0.5)
ax.bar(x+b2off, y2, bottom=0, width=barwidth, label='no detected DRAs', edgecolor='k', linewidth=0.5)
ax.set_xticks(np.arange(0, len(y))+b2off/2)
ax.set_xticklabels([LABELS[i]+ #s.replace('-','\n')+
'\n$N='+str(res_onecall[s]['number of samples'])+'$' for i,s in enumerate(SETNAMES)])
ax.tick_params(axis='both', labelsize=8)
# ax.set_xticklabels([s.replace('-','\n')+
# '\n$N='+str(res_onecall[s]['number of samples'])+'$' for s in SETNAMES])
ax.set_ylabel('% of genes with good targets')
ax.set_ylim((0,100))
ax_leg = axs[1,0]
lh,ll = ax.get_legend_handles_labels()
ax_leg.legend(lh,ll, loc='center')
ax_leg.set_axis_off()
fig.tight_layout()
fig.savefig('percent_good_genes_overview.svg')
fig.savefig('percent_good_genes_overview.png')
# -
# +
fig,axs = mpl.pyplot.subplots(2,1, squeeze=False, figsize=(6,6),
gridspec_kw={'height_ratios': [100, 1]})
barwidth = 1
ax = axs[0,0]
x = np.arange(0, len(y))
y1 = np.array([res_by_minRefAF[s][0.99]['good unique targets % of potential'] for s in SETNAMES])
y2 = np.array([res_onecall[s]['good unique targets % of potential'] for s in SETNAMES])
y3 = np.array([res_by_minRefAF[s][1]['good unique targets % of potential'] for s in SETNAMES])
# y1 = np.array([res_by_minRefAF[s][0.99]['coding genes w/ good targets %'] for s in SETNAMES])
# y2 = np.array([res_onecall[s]['coding genes w/ good targets %'] for s in SETNAMES])
# y3 = np.array([res_by_minRefAF[s][1]['coding genes w/ good targets %'] for s in SETNAMES])
ax.bar(x, y1-y2, bottom=y2, width=barwidth*.8, label='allowing <1% variants', edgecolor='k', linewidth=0.5)
ax.bar(x, y2-y3, bottom=y3, width=barwidth*.8, label='allowing 1-call variants', edgecolor='k', linewidth=0.5)
ax.bar(x, y3, bottom=0, width=barwidth*.8, label='no detected variants', edgecolor='k', linewidth=0.5)
ax.set_xticks(np.arange(0, len(x)))#+barwidth)
ax.set_xticklabels([s.replace('-','\n')+'\n$N='+str(res_onecall[s]['number of samples'])+'$' for s in SETNAMES])
ax.set_ylabel('% of good potential targets')
ax.set_ylim((0,100))
ax_leg = axs[1,0]
lh,ll = ax.get_legend_handles_labels()
ax_leg.legend(lh,ll, loc='center')
ax_leg.set_axis_off()
fig.tight_layout()
# fig.savefig('percent_good_genes_overview.svg')
# -
# +
n_samples = [res_onecall[s]['number of samples'] for s in SETNAMES]
gg = np.array([res_by_minRefAF[s][0.99]['coding genes w/ good targets %'] for s in SETNAMES])
# y2 = np.array([res_onecall[s]['coding genes w/ good targets %'] for s in SETNAMES])
# y3 = np.array([res_by_minRefAF[s][1]['coding genes w/ good targets %'] for s in SETNAMES])
model = sm.OLS(gg, sm.add_constant(n_samples)).fit()
display(model.summary())
fig, ax = mpl.pyplot.subplots(1,1)
ax.plot(n_samples, gg, '.')
# -
# ## targets per gene stats
# setname = 'VGL-gam'
setname = 'Ag1000g-gam'
tout = pd.read_msgpack(setname+'_tout.msgpack')
display(pd.Series(tout['gene'].value_counts()).describe())
setname = 'VGL-Aaeg'
tout = pd.read_msgpack(setname+'_tout.msgpack')
display(pd.Series(tout['gene'].value_counts()).describe())
1-(1-0.03)**47
| variants/results_summary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.chdir(r'/Users/rmccrickerd/desktop/jdheston')
import numpy as np
import pandas as pd
from jdheston import jdheston as jdh
from jdheston import utils as uts
from jdheston import config as cfg
from matplotlib import pyplot as plt
from scipy.stats import norm
from scipy.special import gamma
# import mpl
# # %matplotlib inline
nx = np.newaxis
cfg.config(scale=1.5,print_keys=False)
df = pd.read_excel('data/ccy-date.xlsx')
df
deltas = np.array([0.1,0.25,0.5,0.75,0.9])
expiries = np.array([1/12,3/12,6/12,1])
labels = ['1m','3m','6m','1y']
m,n = len(expiries),len(deltas)
mid_vols = np.zeros((m,n))
spreads = np.zeros((m,n))
mid_vols[:,0] = df.iloc[2:,9]
mid_vols[:,1] = df.iloc[2:,5]
mid_vols[:,2] = df.iloc[2:,1]
mid_vols[:,3] = df.iloc[2:,3]
mid_vols[:,4] = df.iloc[2:,7]
spreads[:,0] = df.iloc[2:,10]
spreads[:,1] = df.iloc[2:,6]
spreads[:,2] = df.iloc[2:,2]
spreads[:,3] = df.iloc[2:,4]
spreads[:,4] = df.iloc[2:,8]
plt.rcParams['figure.figsize'] = [4,8]
plt.rcParams['legend.loc'] = 'lower left'
plot,axes = plt.subplots(m)
# n = -1
for i in range(4):
# n += 1
axes[i].plot(deltas, mid_vols[i,:],'bo')
for j in range(n):
axes[i].plot([deltas[j],deltas[j]], [mid_vols[i,j] - spreads[i,j]/2,mid_vols[i,j] + spreads[i,j]/2],'b')
axes[i].plot(deltas, mid_vols[i,:] + spreads[i,:]/2,'b_')
axes[i].plot(deltas, mid_vols[i,:] - spreads[i,:]/2,'b_')
# axes[i].plot(k[n,:],100*BSV[n,:])
axes[i].set_ylabel(r'$\bar{\sigma}(\Delta,\mathrm{%s})$'%labels[i])
axes[i].set_ylim([5,15])
axes[i].set_xlim([0,1])
# axes[0,1].set_title(r'$\varepsilon=\mathrm{%s}.$'%TS)
# axes[0,1].set_title(r'$H = 0.05,\ \ \varepsilon=\mathrm{%s}.$'%TS)
# axes[1,1].legend([r'$\ \ \mathrm{rHeston}$',r'$\mathrm{qhrHeston\ (NIG)}$'])
axes[0].set_title(r'$\mathrm{GBPUSD 20190612}$')
axes[3].set_xlabel(r'$\Delta$')
plt.tight_layout()
# fname = r'LSR-comparison'
# plt.savefig(fname)
vols = mid_vols/100
vol_surface = jdh.vol_surface(expiries[:,nx], deltas[nx,:], vols)
vol_surface.fit_jheston()
sigma_max = np.max(vol_surface.jheston['sigma'])
sigma_max
T = np.array([1/12,3/12,6/12,1])[:,nx]
# M = ['1W','1M','3M','6M','9M','1Y']
Δ = np.linspace(5,95,19)[nx,:]/100
k = norm.ppf(Δ)*sigma_max*np.sqrt(T)
pd.DataFrame(k,index=T[:,0],columns=np.round(Δ[0,:],2))
vol_surface.jheston
sigma = np.array(vol_surface.jheston['sigma'])
sigma
average_vols = np.array(vol_surface.jheston['sigma'].copy())
forward_vols = average_vols.copy()
for i in np.arange(1,m):
forward_vols[i] = np.sqrt((average_vols[i]**2*T[i,0] - average_vols[i-1]**2*T[i-1,0])/(T[i,0] - T[i-1,0]))
forward_vols
# +
# jump fit
# jump_weight = 0.99
# rho = -0.35
# vee = 1.0
# +
# diffusion fit
# jump_weight = 0.01
# rho = -0.35
# vee = 2.5
# -
# jump-diffusion fit
jump_weight = 0.5
rho = -0.35
vee = 1.75
jump_sigma = jump_weight**0.5*forward_vols
diff_sigma = (1 - jump_weight)**0.5*forward_vols
jump_sigma, diff_sigma
times1 = np.array([0,1/12,3/12,6/12])
sigma1 = jump_sigma
rho1 = np.array([ +0.5])*np.ones_like(times1)
vee1 = np.array([1.0])*np.ones_like(times1)
epsilon1 = np.array([ 0])*np.ones_like(times1)
times2 = np.array([0,1/12,3/12,6/12])
sigma2 = diff_sigma
rho2 = np.array([-0.9])*np.ones_like(times2)
vee2 = np.array([2.5])*np.ones_like(times2)
epsilon2 = np.array([1])*np.ones_like(times2)
params1 = np.array([times1, sigma1, rho1, vee1, epsilon1]).T
params2 = np.array([times2, sigma2, rho2, vee2, epsilon2]).T
np.round(params1,3)
np.round(params2,3)
maturities = T
logstrikes = k
# call_prices = jdh.jdh_pricer(maturities, logstrikes, params1)
call_prices = jdh.jdh2f_pricer(maturities, logstrikes, [params1, params2])
implied_vols = jdh.surface(maturities, logstrikes, call_prices)
pd.DataFrame(implied_vols,index=T[:,0],columns=Δ[0,:])
# +
# plt.rcParams['figure.figsize'] = [2*1.618*2,2*3]
# plt.rcParams['legend.loc'] = 'lower left'
# -
plt.rcParams['figure.figsize'] = [3,3*9/16]
plot,axes = plt.subplots()
for i in range(len(T[:,0])):
axes.plot(k[i,:],100*implied_vols[i,:])
axes.set_xlabel(r'$k$')
axes.set_ylabel(r'$\bar{\sigma}(k,\tau)$')
# plt.savefig('temp')
implied_vars = implied_vols*np.sqrt(T)
Δ = norm.cdf(k/implied_vars + 0.5*implied_vars)
plt.rcParams['figure.figsize'] = [4,8]
plt.rcParams['legend.loc'] = 'upper right'
plot,axes = plt.subplots(m)
for i in range(len(T[:,0])):
axes[i].plot(deltas, mid_vols[i,:],'bo')
axes[i].plot(Δ[i,:],100*implied_vols[i,:])
axes[i].set_xlim([0,1])
axes[i].set_ylabel(r'$\bar{\sigma}(\Delta,\mathrm{%s})$'%labels[i])
axes[0].set_title(r'$\mathrm{GBPUSD 20190612}$')
axes[0].legend([r'$\mathrm{Market}$',r'$\mathrm{jdh2f\ model}$'])
axes[-1].set_xlabel(r'$\Delta$')
plt.tight_layout()
plt.savefig('plots/jump-diffusion-fit')
| notebooks/.ipynb_checkpoints/20190625-2F-jdheston-market-replication-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Mosaic Art
#
# We would like to use images as mosaic pieces to build a bigger image.
#
# Data originally comes from [here](https://www.kaggle.com/thedownhill/art-images-drawings-painting-sculpture-engraving).
# +
from os import listdir
from os.path import isfile, join
import numpy as np
import pandas as pd
from PIL import Image
from lets_plot import *
LetsPlot.setup_html()
# -
def get_grayscale_img(filepath):
if not isfile(filepath):
return None
img = Image.open(filepath)
if not img.mode in ['RGB', 'L']:
return None
return img.convert('L')
WIDTH, HEIGHT = 20, 20
IMAGES_PATH = 'images/sculpture'
data = []
for f in listdir(IMAGES_PATH):
img = get_grayscale_img(join(IMAGES_PATH, f))
img = img.resize((WIDTH, HEIGHT))
image_data = np.asarray(img).reshape(WIDTH * HEIGHT)
image_agg_data = np.array([np.round(image_data.mean()), np.round(image_data.std())])
data.append(np.concatenate((image_agg_data, image_data), axis=0).astype(int))
df = pd.DataFrame(data)
df = df.rename(columns={0: 'color', 1: 'std'})
df = df.sort_values(by=['color', 'std']).reset_index(drop=True)
ggplot() + \
geom_bar(aes(x='color', fill='color'), data=df, sampling=sampling_pick(n=256)) + \
scale_fill_gradient(low='black', high='white') + \
ggtitle('Gray Color Distribution')
for missing_color in set(range(256)) - set(df.color.unique()):
df = df.append([dict(color=missing_color)])
df = df.sort_values(by='color').reset_index(drop=True)
df = df.fillna(method='pad').fillna(method='backfill')
img = get_grayscale_img(join(IMAGES_PATH, 'i - 663.jpeg'))
img = img.resize((80, 80))
rows = ()
for row in np.asarray(img):
cols = ()
for color in row:
subimg = df[df.color == color].iloc[0, 2:].to_numpy().reshape(WIDTH, HEIGHT)
cols = cols + (subimg,)
rows = rows + (np.hstack(cols),)
image_data = np.vstack(rows)
ggplot() + \
geom_image(image_data=image_data) + \
ggsize(800, 800) + \
ggtitle('Mosaic Image') + \
theme_classic() + theme(axis='blank')
| source/examples/demo/mosaic_image.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="xLOXFOT5Q40E"
# ##### Copyright 2020 The TensorFlow Authors.
# + cellView="form" id="iiQkM5ZgQ8r2"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="UndbWF_UpN-X"
# # Noise
# + [markdown] id="i9Jcnb8bQQyd"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/quantum/tutorials/noise"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/quantum/blob/master/docs/tutorials/noise.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/quantum/blob/master/docs/tutorials/noise.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/quantum/docs/tutorials/noise.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="fHHaKIG06Iv_"
# Noise is present in modern day quantum computers. Qubits are susceptible to interference from the surrounding environment, imperfect fabrication, TLS and sometimes even [gamma rays](https://arxiv.org/abs/2104.05219). Until large scale error correction is reached, the algorithms of today must be able to remain functional in the presence of noise. This makes testing algorithms under noise an important step for validating quantum algorithms / models will function on the quantum computers of today.
#
# In this tutorial you will explore the basics of noisy circuit simulation in TFQ via the high level `tfq.layers` API.
#
# ## Setup
# + id="J2CRbYRqrLdt"
# !pip install tensorflow==2.4.1 tensorflow-quantum
# + id="QStNslxBwgte"
# !pip install -q git+https://github.com/tensorflow/docs
# + id="iRU07S4o8B52"
import random
import cirq
import sympy
import tensorflow_quantum as tfq
import tensorflow as tf
import numpy as np
# Plotting
import matplotlib.pyplot as plt
import tensorflow_docs as tfdocs
import tensorflow_docs.plots
# + [markdown] id="CVnAGxZyruv8"
# ## 1. Understanding quantum noise
#
# ### 1.1 Basic circuit noise
#
# Noise on a quantum computer impacts the bitstring samples you are able to measure from it. One intuitive way you can start to think about this is that a noisy quantum computer will "insert", "delete" or "replace" gates in random places like the diagram below:
#
# <img src="./images/noise_1.png" width=700>
#
# Building off of this intuition, when dealing with noise, you are no longer using a single pure state $|\psi \rangle$ but instead dealing with an *ensemble* of all possible noisy realizations of your desired circuit: $\rho = \sum_j p_j |\psi_j \rangle \langle \psi_j |$ . Where $p_j$ gives the probability that the system is in $|\psi_j \rangle$ .
#
# Revisiting the above picture, if we knew beforehand that 90% of the time our system executed perfectly, or errored 10% of the time with just this one mode of failure, then our ensemble would be:
#
# $\rho = 0.9 |\psi_\text{desired} \rangle \langle \psi_\text{desired}| + 0.1 |\psi_\text{noisy} \rangle \langle \psi_\text{noisy}| $
#
# If there was more than just one way that our circuit could error, then the ensemble $\rho$ would contain more than just two terms (one for each new noisy realization that could happen). $\rho$ is referred to as the [density matrix](https://en.wikipedia.org/wiki/Density_matrix) describing your noisy system.
#
# ### 1.2 Using channels to model circuit noise
#
# Unfortunately in practice it's nearly impossible to know all the ways your circuit might error and their exact probabilities. A simplifying assumption you can make is that after each operation in your circuit there is some kind of [channel](https://quantumai.google/cirq/noise) that roughly captures how that operation might error. You can quickly create a circuit with some noise:
# + id="Eu_vpHbfrQKQ"
def x_circuit(qubits):
"""Produces an X wall circuit on `qubits`."""
return cirq.Circuit(cirq.X.on_each(*qubits))
def make_noisy(circuit, p):
"""Add a depolarization channel to all qubits in `circuit` before measurement."""
return circuit + cirq.Circuit(cirq.depolarize(p).on_each(*circuit.all_qubits()))
my_qubits = cirq.GridQubit.rect(1, 2)
my_circuit = x_circuit(my_qubits)
my_noisy_circuit = make_noisy(my_circuit, 0.5)
my_circuit
# + id="1B7vmyPm_TQ7"
my_noisy_circuit
# + [markdown] id="EejhXc2e9Cl8"
# You can examine the noiseless density matrix $\rho$ with:
# + id="0QN9W69U8v_V"
rho = cirq.final_density_matrix(my_circuit)
np.round(rho, 3)
# + [markdown] id="RHHBeizr-DEo"
# And the noisy density matrix $\rho$ with:
#
# + id="zSD9H8SC9IJ1"
rho = cirq.final_density_matrix(my_noisy_circuit)
np.round(rho, 3)
# + [markdown] id="2YWiejLl-a0Z"
# Comparing the two different $ \rho $ 's you can see that the noise has impacted the amplitudes of the state (and consequently sampling probabilities). In the noiseless case you would always expect to sample the $ |11\rangle $ state. But in the noisy state there is now a nonzero probability of sampling $ |00\rangle $ or $ |01\rangle $ or $ |10\rangle $ as well:
# + id="Z4uj-Zs0AE3n"
"""Sample from my_noisy_circuit."""
def plot_samples(circuit):
samples = cirq.sample(circuit + cirq.measure(*circuit.all_qubits(), key='bits'), repetitions=1000)
freqs, _ = np.histogram(samples.data['bits'], bins=[i+0.01 for i in range(-1,2** len(my_qubits))])
plt.figure(figsize=(10,5))
plt.title('Noisy Circuit Sampling')
plt.xlabel('Bitstring')
plt.ylabel('Frequency')
plt.bar([i for i in range(2** len(my_qubits))], freqs, tick_label=['00','01','10','11'])
plot_samples(my_noisy_circuit)
# + [markdown] id="IpPh1Y0HEOWs"
# Without any noise you will always get $|11\rangle$:
# + id="NRCOhTVpEJzz"
"""Sample from my_circuit."""
plot_samples(my_circuit)
# + [markdown] id="EMbJBXAiT9GH"
# If you increase the noise a little further it will become harder and harder to distinguish the desired behavior (sampling $|11\rangle$ ) from the noise:
# + id="D2Fg-FUdUJQx"
my_really_noisy_circuit = make_noisy(my_circuit, 0.75)
plot_samples(my_really_noisy_circuit)
# + [markdown] id="oV-0WV5Z7FQ8"
# Note: Try experimenting with different channels in your circuit to generate noise. Common channels supported in both Cirq and TFQ can be found [here](https://github.com/quantumlib/Cirq/blob/master/cirq-core/cirq/ops/common_channels.py)
# + [markdown] id="atzsYj5qScn0"
# ## 2. Basic noise in TFQ
# With this understanding of how noise can impact circuit execution, you can explore how noise works in TFQ. TensorFlow Quantum uses monte-carlo / trajectory based simulation as an alternative to density matrix simulation. This is because the memory complexity of density matrix simulation limits large simulations to being <= 20 qubits with traditional full density matrix simulation methods. Monte-carlo / trajectory trades this cost in memory for additional cost in time. The `backend='noisy'` option available to all `tfq.layers.Sample`, `tfq.layers.SampledExpectation` and `tfq.layers.Expectation` (In the case of `Expectation` this does add a required `repetitions` parameter).
#
# ### 2.1 Noisy sampling in TFQ
# To recreate the above plots using TFQ and trajectory simulation you can use `tfq.layers.Sample`
# + id="byVI5nbNQ4_b"
"""Draw bitstring samples from `my_noisy_circuit`"""
bitstrings = tfq.layers.Sample(backend='noisy')(my_noisy_circuit, repetitions=1000)
# + id="ncl0ruCZrd2s"
numeric_values = np.einsum('ijk,k->ij', bitstrings.to_tensor().numpy(), [1, 2])[0]
freqs, _ = np.histogram(numeric_values, bins=[i+0.01 for i in range(-1,2** len(my_qubits))])
plt.figure(figsize=(10,5))
plt.title('Noisy Circuit Sampling')
plt.xlabel('Bitstring')
plt.ylabel('Frequency')
plt.bar([i for i in range(2** len(my_qubits))], freqs, tick_label=['00','01','10','11'])
# + [markdown] id="QfHq13RwuLlF"
# ### 2.2 Noisy sample based expectation
# To do noisy sample based expectation calculation you can use `tfq.layers.SampleExpectation`:
#
# + id="ep45G-09rfrA"
some_observables = [cirq.X(my_qubits[0]), cirq.Z(my_qubits[0]), 3.0 * cirq.Y(my_qubits[1]) + 1]
some_observables
# + [markdown] id="ur4iF_PGv0Xf"
# Compute the noiseless expectation estimates via sampling from the circuit:
# + id="jL6wJ3LCvNcn"
noiseless_sampled_expectation = tfq.layers.SampledExpectation(backend='noiseless')(
my_circuit, operators=some_observables, repetitions=10000
)
noiseless_sampled_expectation.numpy()
# + [markdown] id="c6hHgNtEv40i"
# Compare those with the noisy versions:
# + id="8U4Gm-LGvYqa"
noisy_sampled_expectation = tfq.layers.SampledExpectation(backend='noisy')(
[my_noisy_circuit, my_really_noisy_circuit], operators=some_observables, repetitions=10000
)
noisy_sampled_expectation.numpy()
# + [markdown] id="CqQ_2c7XwMku"
# You can see that the noise has particularly impacted the $\langle \psi | Z | \psi \rangle$ accuracy, with `my_really_noisy_circuit` concentrating very quickly towards 0.
#
# ### 2.3 Noisy analytic expectation calculation
# Doing noisy analytic expectation calculations is nearly identical to above:
#
#
# + id="pGXKlyCywAfj"
noiseless_analytic_expectation = tfq.layers.Expectation(backend='noiseless')(
my_circuit, operators=some_observables
)
noiseless_analytic_expectation.numpy()
# + id="6FUkJ7aOyTlI"
noisy_analytic_expectation = tfq.layers.Expectation(backend='noisy')(
[my_noisy_circuit, my_really_noisy_circuit], operators=some_observables, repetitions=10000
)
noisy_analytic_expectation.numpy()
# + [markdown] id="5KHvORT42XFV"
# ## 3. Hybrid models and quantum data noise
# Now that you have implemented some noisy circuit simulations in TFQ, you can experiment with how noise impacts quantum and hybrid quantum classical models, by comparing and contrasting their noisy vs noiseless performance. A good first check to see if a model or algorithm is robust to noise is to test under a circuit wide depolarizing model which looks something like this:
#
# <img src="./images/noise_2.png" width=500>
#
# Where each time slice of the circuit (sometimes referred to as moment) has a depolarizing channel appended after each gate operation in that time slice. The depolarizing channel with apply one of $\{X, Y, Z \}$ with probability $p$ or apply nothing (keep the original operation) with probability $1-p$.
#
# ### 3.1 Data
# For this example you can use some prepared circuits in the `tfq.datasets` module as training data:
# + id="_ZqVLEji2WUx"
qubits = cirq.GridQubit.rect(1, 8)
circuits, labels, pauli_sums, _ = tfq.datasets.xxz_chain(qubits, 'closed')
circuits[0]
# + [markdown] id="MFgNU_nBGeTm"
# Writing a small helper function will help to generate the data for the noisy vs noiseless case:
# + id="zkQofAqqGibQ"
def get_data(qubits, depolarize_p=0.):
"""Return quantum data circuits and labels in `tf.Tensor` form."""
circuits, labels, pauli_sums, _ = tfq.datasets.xxz_chain(qubits, 'closed')
if depolarize_p >= 1e-5:
circuits = [circuit.with_noise(cirq.depolarize(depolarize_p)) for circuit in circuits]
tmp = list(zip(circuits, labels))
random.shuffle(tmp)
circuits_tensor = tfq.convert_to_tensor([x[0] for x in tmp])
labels_tensor = tf.convert_to_tensor([x[1] for x in tmp])
return circuits_tensor, labels_tensor
# + [markdown] id="FtJrfsLCF9Z3"
# ### 3.2 Define a model circuit
# Now that you have quantum data in the form of circuits, you will need a circuit to model this data, like with the data you can write a helper function to generate this circuit optionally containing noise:
# + id="TwryFaFIG2Ya"
def modelling_circuit(qubits, depth, depolarize_p=0.):
"""A simple classifier circuit."""
dim = len(qubits)
ret = cirq.Circuit(cirq.H.on_each(*qubits))
for i in range(depth):
# Entangle layer.
ret += cirq.Circuit(cirq.CX(q1, q2) for (q1, q2) in zip(qubits[::2], qubits[1::2]))
ret += cirq.Circuit(cirq.CX(q1, q2) for (q1, q2) in zip(qubits[1::2], qubits[2::2]))
# Learnable rotation layer.
# i_params = sympy.symbols(f'layer-{i}-0:{dim}')
param = sympy.Symbol(f'layer-{i}')
single_qb = cirq.X
if i % 2 == 1:
single_qb = cirq.Y
ret += cirq.Circuit(single_qb(q) ** param for q in qubits)
if depolarize_p >= 1e-5:
ret = ret.with_noise(cirq.depolarize(depolarize_p))
return ret, [op(q) for q in qubits for op in [cirq.X, cirq.Y, cirq.Z]]
modelling_circuit(qubits, 3)[0]
# + [markdown] id="U-ZMaCpJI9TH"
# ### 3.3 Model building and training
# With your data and model circuit built, the final helper function you will need is one that can assemble both a noisy or a noiseless hybrid quantum `tf.keras.Model`:
# + id="r09CT5N9DWa_"
def build_keras_model(qubits, depolarize_p=0.):
"""Prepare a noisy hybrid quantum classical Keras model."""
spin_input = tf.keras.Input(shape=(), dtype=tf.dtypes.string)
circuit_and_readout = modelling_circuit(qubits, 4, depolarize_p)
if depolarize_p >= 1e-5:
quantum_model = tfq.layers.NoisyPQC(*circuit_and_readout, sample_based=False, repetitions=10)(spin_input)
else:
quantum_model = tfq.layers.PQC(*circuit_and_readout)(spin_input)
intermediate = tf.keras.layers.Dense(4, activation='sigmoid')(quantum_model)
post_process = tf.keras.layers.Dense(1)(intermediate)
return tf.keras.Model(inputs=[spin_input], outputs=[post_process])
# + [markdown] id="QbMtT7BZmhfm"
# ## 4. Compare performance
#
# ### 4.1 Noiseless baseline
#
# With your data generation and model building code, you can now compare and contrast model performance in the noiseless and noisy settings, first you can run a reference noiseless training:
# + id="QAgpq9c-EakW"
training_histories = dict()
depolarize_p = 0.
n_epochs = 50
phase_classifier = build_keras_model(qubits, depolarize_p)
phase_classifier.compile(optimizer=tf.keras.optimizers.Adam(learning_rate=0.02),
loss=tf.keras.losses.BinaryCrossentropy(from_logits=True),
metrics=['accuracy'])
# Show the keras plot of the model
tf.keras.utils.plot_model(phase_classifier, show_shapes=True, dpi=70)
# + id="9tKimWRMlVfL"
noiseless_data, noiseless_labels = get_data(qubits, depolarize_p)
training_histories['noiseless'] = phase_classifier.fit(x=noiseless_data,
y=noiseless_labels,
batch_size=16,
epochs=n_epochs,
validation_split=0.15,
verbose=1)
# + [markdown] id="A9oql6Synv3f"
# And explore the results and accuracy:
# + id="TG87YNUWKKLY"
loss_plotter = tfdocs.plots.HistoryPlotter(metric = 'loss', smoothing_std=10)
loss_plotter.plot(training_histories)
# + id="O2ZwM18YUxxm"
acc_plotter = tfdocs.plots.HistoryPlotter(metric = 'accuracy', smoothing_std=10)
acc_plotter.plot(training_histories)
# + [markdown] id="JlOwBxvSnzid"
# ### 4.2 Noisy comparison
# Now you can build a new model with noisy structure and compare to the above, the code is nearly identical:
# + id="0jy54uWpgwhi"
depolarize_p = 0.001
n_epochs = 50
noisy_phase_classifier = build_keras_model(qubits, depolarize_p)
noisy_phase_classifier.compile(optimizer=tf.keras.optimizers.Adam(learning_rate=0.02),
loss=tf.keras.losses.BinaryCrossentropy(from_logits=True),
metrics=['accuracy'])
# Show the keras plot of the model
tf.keras.utils.plot_model(noisy_phase_classifier, show_shapes=True, dpi=70)
# + [markdown] id="r-vYU6S3oN-J"
# Note: in the model diagram there is now a `tfq.layers.NoisyPQC` instead of a `tfq.layers.PQC` since the depolarization probability is no longer zero. Training will take significantly longer since noisy simulation is far more expensive than noiseless.
# + id="210cLP5AoClJ"
noisy_data, noisy_labels = get_data(qubits, depolarize_p)
training_histories['noisy'] = noisy_phase_classifier.fit(x=noisy_data,
y=noisy_labels,
batch_size=16,
epochs=n_epochs,
validation_split=0.15,
verbose=1)
# + id="eQ8pknNdohzy"
loss_plotter.plot(training_histories)
# + id="nBtgnKWtuWRR"
acc_plotter.plot(training_histories)
# + [markdown] id="r86TeFxlubls"
# Success: The noisy model still managed to train under some mild depolarization noise. Try experimenting with different noise models to see how and when training might fail. Also look out for noisy functionality under `tfq.layers` and `tfq.noise`.
| docs/tutorials/noise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Plotting type of clouds symbols on geographical map
#
# This is a single cell notebook with example of advanced plotting of type of clouds symbols.
#
# List of all **msymbol** parameters you can find [in Magics documentation](https://confluence.ecmwf.int/display/MAGP/Symbol "Symbol parameters").
# More symbol plotting examples can be found in [Simple symbol plotting](../tutorials/Symbol_simple.ipynb "Symbol simple") and [Advanced symbol plotting](../tutorials/Advanced_simple.ipynb "Symbol advanced") notebook.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Installing Magics
# If you don't have Magics installed, run the next cell to install Magics using conda.
# -
# Install Magics in the current Jupyter kernel
import sys
# !conda install --yes --prefix {sys.prefix} Magics
# + slideshow={"slide_type": "slide"}
import Magics.macro as magics
# Loading data in geopoints format
tcc = magics.mgeo(geo_input_file_name = "../../data/ct.gpt") # Total cloud amount
#Coastlines
#Coastlines
coast = magics.mcoast(
map_coastline_colour = "RGB(0.8,0.8,0.8,0.5)",
map_coastline_resolution = "medium",
map_coastline_thickness = 1,
map_coastline_land_shade = "on",
map_coastline_land_shade_colour = "#FAF0E6",
map_coastline_sea_shade = "on",
map_coastline_sea_shade_colour = "#E0FFFF",
map_grid_line_style = "dash",
map_label_height = 0.35,
map_grid_colour = "RGB(0.8,0.8,0.8,0.5)")
#Geographical projection
central_europe = magics.mmap(
superpage_background_colour = "black",
subpage_map_library_area = "on",
subpage_map_area_name = "central_europe",
subpage_clipping = "on",
page_id_line = "off"
)
#Defining the symbols
tcc_symb = magics.msymb(
symbol_advanced_table_height_method = "calculate",
symbol_advanced_table_height_min_value = 1.5,
symbol_advanced_table_height_max_value = 1.5,
symbol_type = "marker",
legend = "on",
symbol_table_mode = "advanced",
symbol_marker_mode = "name",
symbol_advanced_table_selection_type = "list",
symbol_advanced_table_colour_method = "list",
symbol_advanced_table_colour_list = ['evergreen','evergreen','evergreen','#800000','navy',
'navy','navy','navy','#800000','red'],
symbol_advanced_table_level_list = [0.,1.,2.,3.,4.,5.,6.,7.,8.,9,10],
symbol_advanced_table_marker_name_list = ['C_0','C_1','C_2','C_3','C_4','C_5','C_6','C_7','C_8','C_9'])
# Defining the legend
symb_legend = magics.mlegend(
legend_user_lines = ["Ci","Cc", "Cs", "Ac", "As", "Ns", "Sc", "St", "Cu","Cb"],
legend_box_mode = "positional",
legend_text_composition = "user_text_only",
legend_text_colour = "charcoal",
legend_text_font_size = 0.5,
legend_column_count = 5,
legend_box_y_position = 16.5,
legend_box_x_position = 0.,
legend_box_x_length = 22.00,
legend_box_y_length = 3.30)
magics.plot(central_europe, coast, tcc, tcc_symb, symb_legend)
| visualisation/gallery/symbol_cloud_type.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Classification model to detect pneumonia
#
# ## Context
#
# This model detects pneumonia based on chest x-ray image. It is a convolutional neural networks with transfer learning using VGG16. The data was taken from Kaggle dataset [Chest X-Ray Images (Pneumonia)](https://www.kaggle.com/paultimothymooney/chest-xray-pneumonia/tasks?taskId=467).
#
# <img src='assets/intro_01.png' />
#
# *Illustrative examples of chest x-rays in patients with pneumonia. The normal chest X-ray (left panel) depicts clear lungs without any areas of abnormal opacification in the image. Bacterial pneumonia (middle) typically exhibits a focal lobar consolidation, in this case in the right upper lobe (white arrows), whereas viral pneumonia (right) manifests with a more diffuse ‘‘interstitial’’ pattern in both lungs* [(Kermany DS, Goldbaum M, Cai W, et al. (2018))]('http://www.cell.com/cell/fulltext/S0092-86741830154-5).
#
# Chest X-ray images (anterior-posterior) were selected from retrospective cohorts of pediatric patients of one to five years old from Guangzhou Women and Children’s Medical Center, Guangzhou. All chest X-ray imaging was performed as part of patients’ routine clinical care.
#
# For the analysis of chest x-ray images, all chest radiographs were initially screened for quality control by removing all low quality or unreadable scans. The diagnoses for the images were then graded by two expert physicians before being cleared for training the AI system. In order to account for any grading errors, the evaluation set was also checked by a third expert.
import numpy as np
import pandas as pd
import os
from glob import glob
# First, let's inspect the data that is available.
# +
# loading the directories
trai_dir = '../input/chest-xray-pneumonia/chest_xray/train/'
val_dir = '../input/chest-xray-pneumonia/chest_xray/val/'
test_dir = '../input/chest-xray-pneumonia/chest_xray/test/'
# getting the number of classes
folders = glob(trai_dir + '/*')
num_classes = len(folders)
class_labels=os.listdir(trai_dir)
print ('Total number of classes = ' + str(num_classes))
print('Class names: {0}'.format(class_labels))
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# getting number of files
train_files = np.array(glob(trai_dir+"*/*"))
val_files = np.array(glob(val_dir+"*/*"))
test_files = np.array(glob(test_dir+"*/*"))
# print number of images in each dataset
print('There are %d total train images.' % len(train_files))
print('There are %d total validation images.' % len(val_files))
print('There are %d total test images.' % len(test_files))
# +
train_normal = np.array(glob(trai_dir+"NORMAL/*"))
val_normal = np.array(glob(val_dir+"NORMAL/*"))
test_normal = np.array(glob(test_dir+"NORMAL/*"))
train_pneumonia = np.array(glob(trai_dir+"PNEUMONIA/*"))
val_pneumonia = np.array(glob(val_dir+"PNEUMONIA/*"))
test_pneumonia = np.array(glob(test_dir+"PNEUMONIA/*"))
print('There are %d total normal train images.' % len(train_normal))
print('There are %d total normal validation images.' % len(val_normal))
print('There are %d total normal test images.' % len(test_normal))
print('There are %d total pneumonia train images.' % len(train_pneumonia))
print('There are %d total pneumonia validation images.' % len(val_pneumonia))
print('There are %d total pneumonia test images.' % len(test_pneumonia))
# -
# Data distribution is very uneven. There is signinficantly less pneumonia images (3875) than normal ones (14341). Moreover, the validation set is particularly small, only 8 images for both normal and pneumonia classes.
#
# Let's display some sample images of healthy and infected lungs.
# +
import cv2
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
file_path = np.concatenate((train_normal[0:3],train_pneumonia[0:3]))
fig = plt.figure(figsize=(13, 9))
for i in range(len(file_path)):
ax = fig.add_subplot(2,3,i+1, xticks=[], yticks=[])
img=mpimg.imread(file_path[i])
ax.set_title(file_path[i].split('/')[-2])
imgplot = plt.imshow(img, cmap='gray', vmin=0, vmax=255)
plt.show()
# -
# ## Specify Data Loaders for the Dog Dataset
#
# Three separate data loaders for the training, validation and test datasets are created. Train and validation sets were resized to 256, randomly flipped and rotated up to 20 degrees. Next, all data sets have been cropped to 224 x 224 size images, similarly to other pretrained models available in torchvision.models module. Finally, the images were transformed to tensors and their channels were normalised with means of [0.485, 0.456, 0.406] and standard deviations of [0.229, 0.224, 0.225].
# +
import torch
import torchvision.transforms as transforms
from torchvision import datasets
# Create training and test dataloaders
# number of subprocesses to use for data loading
num_workers = 0
# how many samples per batch to load
batch_size = 32
# resize the picture
size = 256
data_transform_train = transforms.Compose([
transforms.RandomHorizontalFlip(), # randomly flip and rotate
transforms.RandomRotation(20),
transforms.Resize(size),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize((0.485, 0.456, 0.406),
(0.229, 0.224, 0.225))])
data_transform_test = transforms.Compose([
transforms.Resize(size),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize((0.485, 0.456, 0.406),
(0.229, 0.224, 0.225))])
dog_image_dir = '/data/dog_images'
# choose the training and test datasets
train_data = datasets.ImageFolder(trai_dir, transform=data_transform_train)
valid_data = datasets.ImageFolder(val_dir, transform=data_transform_train)
test_data = datasets.ImageFolder(test_dir, transform=data_transform_test)
# prepare data loaders
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size, shuffle=True, num_workers=num_workers)
valid_loader = torch.utils.data.DataLoader(valid_data, batch_size=batch_size, shuffle=True, num_workers=num_workers)
test_loader = torch.utils.data.DataLoader(test_data, batch_size=batch_size, shuffle=True, num_workers=num_workers)
data_loader = dict(train=train_loader, valid=valid_loader, test=test_loader)
# Let's verify that the data was loaded correctly by printing out data stats
print('Num training images: ', len(train_data))
print('Num validation images: ', len(valid_data))
print('Num test images: ', len(test_data))
# +
# Visualize some sample train data
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy() # convert images to numpy for display
# denormalize the image
def denormalise(image):
image = np.transpose(image, (1, 2, 0))
mean = [0.485, 0.456, 0.406]
std = [0.229, 0.224, 0.225]
image = (image * std + mean).clip(0, 1)
return image
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 9))
no_vis_imag = 14
for idx in np.arange(no_vis_imag):
ax = fig.add_subplot(2, no_vis_imag/2, idx+1, xticks=[], yticks=[])
plt.imshow(denormalise(images[idx]))
ax.set_title(class_labels[labels[idx]])
# -
# ## Create a CNN using Transfer Learning
#
# VGG16 is used with all the convolutional layers obtained from the pretrained model. The final fully-connected layer is replaced with a new classifier to match the two lung classes.
# +
import torchvision.models as models
#import torch.nn.functional as F
import torch.nn as nn
# Load the pretrained model from pytorch
VGG16 = models.vgg16(pretrained=True)
# Modify the last layer
n_inputs = VGG16.classifier[6].in_features
last_layer = nn.Linear(n_inputs, len(class_labels))
VGG16.classifier[6] = last_layer
# +
## Specify Loss Function and Optimizer
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(VGG16.parameters(), lr=0.01)
# check if CUDA is available
use_cuda = torch.cuda.is_available()
if use_cuda:
VGG16 = VGG16.cuda()
print('CUDA is available! Training on GPU ...')
else:
print('CUDA is not available. Training on CPU ...')
# +
#from PIL import ImageFile
#ImageFile.LOAD_TRUNCATED_IMAGES = True
def train(n_epochs, loaders, model, optimizer, criterion, use_cuda, save_path):
"""returns trained model with the possibility to resume analysis and load validation parameters"""
valid_loss_min = np.Inf # track change in validation loss
for epoch in range(1, n_epochs+1):
# initialize variables to monitor training and validation loss
train_loss = 0.0
valid_loss = 0.0
###################
# train the model #
###################
# set model to train mode by using dropout to prevent overfitting
model.train()
for batch_idx, (data, target) in enumerate(loaders['train']):
# move to GPU
if use_cuda:
data, target = data.cuda(), target.cuda()
# clear the gradients of all optimized variables
optimizer.zero_grad()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# update training loss
#train_loss += loss.item()*data.size(0)
train_loss = train_loss + ((1 / (batch_idx + 1)) * (loss.data - train_loss))
######################
# validate the model #
######################
model.eval()
for batch_idx, (data, target) in enumerate(loaders['valid']):
# move to GPU
if use_cuda:
data, target = data.cuda(), target.cuda()
## update the average validation loss
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# update average validation loss
valid_loss = valid_loss + ((1 / (batch_idx + 1)) * (loss.data - valid_loss))
# calculate average losses
train_loss = train_loss/len(train_loader.dataset)
valid_loss = valid_loss/len(valid_loader.dataset)
# print training/validation statistics
print('Epoch: {} \tTraining Loss: {:.6f} \tValidation Loss: {:.6f}'.format(
epoch, train_loss, valid_loss))
# save model if validation loss has decreased
if valid_loss <= valid_loss_min:
print('Validation loss decreased ({:.6f} --> {:.6f}). Saving model ...'.format(
valid_loss_min, valid_loss))
torch.save(model.state_dict(), save_path)
valid_loss_min = valid_loss
# return trained model
return model
# +
# train the model
n_epoch = 8
model = train(n_epoch, data_loader, VGG16, optimizer,
criterion, use_cuda, 'model.pt')
# load the model that got the best validation accuracy
model.load_state_dict(torch.load('model.pt'))
# -
# ## Test the Trained Network
# Test the trained model on previously unseen data.
# +
def test(loaders, model, criterion, use_cuda):
# monitor test loss and accuracy
test_loss = 0.
correct = 0.
total = 0.
model.eval()
#for batch_idx, (data, target) in enumerate(loaders['test']):
for batch_idx, (data, target) in enumerate(test_loader):
# move to GPU
if use_cuda:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# update average test loss
test_loss = test_loss + ((1 / (batch_idx + 1)) * (loss.data - test_loss))
# convert output probabilities to predicted class
pred = output.data.max(1, keepdim=True)[1]
# compare predictions to true label
correct += np.sum(np.squeeze(pred.eq(target.data.view_as(pred))).cpu().numpy())
total += data.size(0)
print('Test Loss: {:.6f}\n'.format(test_loss))
print('\nTest Accuracy: %2d%% (%2d/%2d)' % (
100. * correct / total, correct, total))
# call test function
test(data_loader, model, criterion, use_cuda)
# -
# ## Visualize Sample Test Results
#
# +
# obtain one batch of test images
dataiter = iter(test_loader)
images, labels = dataiter.next()
#images = images.numpy() # convert images to numpy for display
images_np = images.cpu().numpy() if not use_cuda else images.numpy()
# move model inputs to cuda, if GPU available
if use_cuda:
images = images.cuda()
# get sample outputs
output = model(images)
# convert output probabilities to predicted class
_, preds_tensor = torch.max(output, 1)
preds = np.squeeze(preds_tensor.numpy()) if not use_cuda else np.squeeze(preds_tensor.cpu().numpy())
#preds = np.squeeze(preds_tensor.cpu().numpy()) # if not use_cuda else np.squeeze(preds_tensor.cpu().numpy())
# plot the images in the batch, along with predicted and true labels
fig = plt.figure(figsize=(25, 4))
no_vis_imag = 16
for idx in np.arange(no_vis_imag):
ax = fig.add_subplot(2, no_vis_imag/2, idx+1, xticks=[], yticks=[])
plt.imshow(denormalise(images_np[idx]))
ax.set_title(class_labels[labels[idx]])
ax.set_title("{} ({})".format(class_labels[preds[idx]], class_labels[labels[idx]]),
color=("green" if preds[idx]==labels[idx].item() else "red"))
| .ipynb_checkpoints/SS-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from bokeh.io import output_file, show
from bokeh.plotting import figure
from bokeh.models import CustomJS, ColumnDataSource, HoverTool, NumeralTickFormatter
def candlestick_plot(df, name):
# Select the datetime format for the x axis depending on the timeframe
xaxis_dt_format = '%d %b %Y'
if df['Date'][0].hour > 0:
xaxis_dt_format = '%d %b %Y, %H:%M:%S'
fig = figure(sizing_mode='stretch_both',
tools="xpan,xwheel_zoom,reset,save",
active_drag='xpan',
active_scroll='xwheel_zoom',
x_axis_type='linear',
# x_range=Range1d(df.index[0], df.index[-1], bounds="auto"),
title=name
)
fig.yaxis[0].formatter = NumeralTickFormatter(format="$5.3f")
inc = df.Close > df.Open
dec = ~inc
# Colour scheme for increasing and descending candles
INCREASING_COLOR = '#17BECF'
DECREASING_COLOR = '#7F7F7F'
width = 0.5
inc_source = ColumnDataSource(data=dict(
x1=df.index[inc],
top1=df.Open[inc],
bottom1=df.Close[inc],
high1=df.High[inc],
low1=df.Low[inc],
Date1=df.Date[inc]
))
dec_source = ColumnDataSource(data=dict(
x2=df.index[dec],
top2=df.Open[dec],
bottom2=df.Close[dec],
high2=df.High[dec],
low2=df.Low[dec],
Date2=df.Date[dec]
))
# Plot candles
# High and low
fig.segment(x0='x1', y0='high1', x1='x1', y1='low1', source=inc_source, color=INCREASING_COLOR)
fig.segment(x0='x2', y0='high2', x1='x2', y1='low2', source=dec_source, color=DECREASING_COLOR)
# Open and close
r1 = fig.vbar(x='x1', width=width, top='top1', bottom='bottom1', source=inc_source,
fill_color=INCREASING_COLOR, line_color="black")
r2 = fig.vbar(x='x2', width=width, top='top2', bottom='bottom2', source=dec_source,
fill_color=DECREASING_COLOR, line_color="black")
# Add on extra lines (e.g. moving averages) here
# fig.line(df.index, <your data>)
# Add on a vertical line to indicate a trading signal here
# vline = Span(location=df.index[-<your index>, dimension='height',
# line_color="green", line_width=2)
# fig.renderers.extend([vline])
# Add date labels to x axis
fig.xaxis.major_label_overrides = {
i: date.strftime(xaxis_dt_format) for i, date in enumerate(pd.to_datetime(df["Date"]))
}
# Set up the hover tooltip to display some useful data
fig.add_tools(HoverTool(
renderers=[r1],
tooltips=[
("Open", "$@top1"),
("High", "$@high1"),
("Low", "$@low1"),
("Close", "$@bottom1"),
("Date", "@Date1{" + xaxis_dt_format + "}"),
],
formatters={
'Date1': 'datetime',
}))
fig.add_tools(HoverTool(
renderers=[r2],
tooltips=[
("Open", "$@top2"),
("High", "$@high2"),
("Low", "$@low2"),
("Close", "$@bottom2"),
("Date", "@Date2{" + xaxis_dt_format + "}")
],
formatters={
'Date2': 'datetime'
}))
# JavaScript callback function to automatically zoom the Y axis to
# view the data properly
source = ColumnDataSource({'Index': df.index, 'High': df.High, 'Low': df.Low})
callback = CustomJS(args={'y_range': fig.y_range, 'source': source}, code='''
clearTimeout(window._autoscale_timeout);
var Index = source.data.Index,
Low = source.data.Low,
High = source.data.High,
start = cb_obj.start,
end = cb_obj.end,
min = Infinity,
max = -Infinity;
for (var i=0; i < Index.length; ++i) {
if (start <= Index[i] && Index[i] <= end) {
max = Math.max(High[i], max);
min = Math.min(Low[i], min);
}
}
var pad = (max - min) * .05;
window._autoscale_timeout = setTimeout(function() {
y_range.start = min - pad;
y_range.end = max + pad;
});
''')
# Finalise the figure
fig.x_range.callback = callback
show(fig)
# Main function
if __name__ == '__main__':
# Read CSV
df = pd.read_csv("./BA_60min.csv").head(500)
# Reverse the order of the dataframe - comment this out if it flips your chart
df = df[::-1]
df.index = df.index[::-1]
# Trim off the unnecessary bit of the minute timeframe data - can be unnecessary
# depending on where you source your data
if '-04:00' in df['Date'][0]:
df['Date'] = df['Date'].str.slice(0, -6)
# Convert the dates column to datetime objects
df["Date"] = pd.to_datetime(df["Date"], format='%Y-%m-%d %H:%M:%S') # Adjust this
output_file("BA_1hour_plot0.html")
candlestick_plot(df, "BA Hourly")
# +
# The use ful code for out ploting of data
import pandas as pd
from bokeh.io import output_file, show
from bokeh.plotting import figure
from bokeh.models import CustomJS, ColumnDataSource, HoverTool, NumeralTickFormatter
def candlestick_plot(df, name):
# Select the datetime format for the x axis depending on the timeframe
xaxis_dt_format = '%d %b %Y'
if df['Date'][0].hour > 0:
xaxis_dt_format = '%d %b %Y, %H:%M:%S'
fig = figure(sizing_mode='stretch_both',
tools="xpan,xwheel_zoom,reset,save",
active_drag='xpan',
active_scroll='xwheel_zoom',
x_axis_type='linear',
# x_range=Range1d(df.index[0], df.index[-1], bounds="auto"),
title=name
)
fig.yaxis[0].formatter = NumeralTickFormatter(format="$5.3f")
inc = df.Close > df.Open
dec = ~inc
# Colour scheme for increasing and descending candles
INCREASING_COLOR = '#17BECF'
DECREASING_COLOR = '#7F7F7F'
width = 0.5
inc_source = ColumnDataSource(data=dict(
x1=df.index[inc],
top1=df.Open[inc],
bottom1=df.Close[inc],
high1=df.High[inc],
low1=df.Low[inc],
Date1=df.Date[inc]
))
dec_source = ColumnDataSource(data=dict(
x2=df.index[dec],
top2=df.Open[dec],
bottom2=df.Close[dec],
high2=df.High[dec],
low2=df.Low[dec],
Date2=df.Date[dec]
))
# Plot candles
# High and low
fig.segment(x0='x1', y0='high1', x1='x1', y1='low1', source=inc_source, color=INCREASING_COLOR)
fig.segment(x0='x2', y0='high2', x1='x2', y1='low2', source=dec_source, color=DECREASING_COLOR)
# Open and close
r1 = fig.vbar(x='x1', width=width, top='top1', bottom='bottom1', source=inc_source,
fill_color=INCREASING_COLOR, line_color="black")
r2 = fig.vbar(x='x2', width=width, top='top2', bottom='bottom2', source=dec_source,
fill_color=DECREASING_COLOR, line_color="black")
# Add on extra lines (e.g. moving averages) here
# fig.line(df.index, <your data>)
# Add on a vertical line to indicate a trading signal here
# vline = Span(location=df.index[-<your index>, dimension='height',
# line_color="green", line_width=2)
# fig.renderers.extend([vline])
# Add date labels to x axis
fig.xaxis.major_label_overrides = {
i: date.strftime(xaxis_dt_format) for i, date in enumerate(pd.to_datetime(df["Date"]))
}
# Set up the hover tooltip to display some useful data
fig.add_tools(HoverTool(
renderers=[r1],
tooltips=[
("Open", "$@top1"),
("High", "$@high1"),
("Low", "$@low1"),
("Close", "$@bottom1"),
("Date", "@Date1{" + xaxis_dt_format + "}"),
],
formatters={
'Date1': 'datetime',
}))
fig.add_tools(HoverTool(
renderers=[r2],
tooltips=[
("Open", "$@top2"),
("High", "$@high2"),
("Low", "$@low2"),
("Close", "$@bottom2"),
("Date", "@Date2{" + xaxis_dt_format + "}")
],
formatters={
'Date2': 'datetime'
}))
# JavaScript callback function to automatically zoom the Y axis to
# view the data properly
source = ColumnDataSource({'Index': df.index, 'High': df.High, 'Low': df.Low})
callback = CustomJS(args={'y_range': fig.y_range, 'source': source}, code='''
clearTimeout(window._autoscale_timeout);
var Index = source.data.Index,
Low = source.data.Low,
High = source.data.High,
start = cb_obj.start,
end = cb_obj.end,
min = Infinity,
max = -Infinity;
for (var i=0; i < Index.length; ++i) {
if (start <= Index[i] && Index[i] <= end) {
max = Math.max(High[i], max);
min = Math.min(Low[i], min);
}
}
var pad = (max - min) * .05;
window._autoscale_timeout = setTimeout(function() {
y_range.start = min - pad;
y_range.end = max + pad;
});
''')
# Finalise the figure
fig.x_range.callback = callback
show(fig)
# Main function
if __name__ == '__main__':
# Read CSV
df = pd.read_csv("data_prepd2.csv").head(500)
# Reverse the order of the dataframe - comment this out if it flips your chart
df = df[::-1]
df.index = df.index[::-1]
# Trim off the unnecessary bit of the minute timeframe data - can be unnecessary
# depending on where you source your data
if '-04:00' in df['Date'][0]:
df['Date'] = df['Date'].str.slice(0, -6)
# Convert the dates column to datetime objects
df["Date"] = pd.to_datetime(df["Date"], format='%Y-%d-%m %H:%M:%S') # Adjust this
output_file("BA_1hour_plot.html")
candlestick_plot(df, "BA Hourly")
# -
# +
# Renaming our coulmn as same name as Ploting code uses
df = pd.read_csv("Data_EURUSD.csv").head(500)
df['Local time']
df["Date"] = pd.to_datetime(df['Local time'])
df
# -
df1=df[['Date','Open','High','Low','Close']]
df1.to_csv('data_prepd2.csv')
| ver1_only_line_ plot matched to our data/Working_plot_code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
## Imports
import sys
import numpy as np
import matplotlib
from matplotlib import pyplot as plt
sys.path.append("../.")
import handybeam
import handybeam.world
import handybeam.tx_array_library
import handybeam.tx_array
import handybeam.visualise
import handybeam.samplers.hexagonal_sampler
from handybeam.solver import Solver
matplotlib.rcParams['figure.figsize'] = [20, 10]
# +
# Initialise the world
world = handybeam.world.World()
# Initialise a solver
solver = Solver(parent = world)
# Add a transmitter array to the world
world.tx_array = handybeam.tx_array_library.rectilinear(parent = world)
# Instruct the solver to solve for the activation coefficients
solver.single_focus_solver(x_focus = 0, y_focus = 0, z_focus = 200e-3)
# Set grid spacing per wavelength for rectilinear sampling grid.
grid_spacing_per_wavelength = 0.2
# Set grid extent around the origin.
grid_extent_around_origin = 0.1
# Specify the vector normal to the sampling grid
norm_vector = np.array((0,1,-1))
# Specify a vector that is orthogonal to this vector ( this will be parallel to one axis of the sampling grid)
par_vector = np.array((0,1,1))
# Add a rectilinear sampling grid to the world
hex_sampler = world.add_sampler(handybeam.samplers.hexagonal_sampler.HexagonalSampler( parent = world,
origin = np.array((0,0,200e-3)),
normal_vector = norm_vector,
parallel_vector = par_vector,
grid_spacing_per_wavelength = grid_spacing_per_wavelength,
grid_extent_around_origin = grid_extent_around_origin))
# Propagate the acoustic field
world.propagate()
# Visualise the result
hex_sampler.visualise_all_in_one()
# -
| demos/arbitrary_hexagonal_sampler_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <font color="blue">Projeto de Clustering</font>
# 
#
# ## Objetivo
#
# Criar um sistema de recomendação utilizando o algoritmo *não-supervisionado* KMeans do `Scikit-learn`.<br>
# Neste projeto será utilizado uma base de dados com músicas do serviço de *streaming* de áudio **Spotify**.
#
# ## Spotify
#
# É um serviço de *streaming* de áudio, disponível em praticamente todas as plataformas e está presente no mundo todo. Foi lançado oficialmente em 2008, na Suécia. Possui acordos com a *Universal Music*, *Sony BGM*, *EMI Music* e *Warner Music Group*.
#
# O usuário pode encontrar playlists e rádios, checar quais músicas estão fazendo sucesso entre os assinantes, criar coleções ou seguir as coleções de amigos e artistas. A plataforma conta com mais de 170 milhões de usuários e veio para o Brasil em 2014.
# ## Importando as bibliotecas
# +
# para identificar os arquivos em uma pasta
import glob
# para manipulação dos dados
import pandas as pd
import numpy as np
# para visualizações
import seaborn as sns
import matplotlib.pyplot as plt
# para pré-processamento
from sklearn.preprocessing import MinMaxScaler
# para machine learning do agrupamento
from sklearn.cluster import KMeans
# para ignorar eventuais warnings
import warnings
warnings.filterwarnings("ignore")
# algumas configurações do notebook
# %matplotlib inline
pd.options.display.max_columns = None
plt.style.use('ggplot')
# -
# ## importando a base de dados
# Vamos importar nossa base de dados, utilizando o método *glob* do framework de mesmo nome para buscar os arquivos *".csv"* que contem na pasta, no nosso caso temos 2 arquivos referentes aos anos de 2018 e 2019. Em seguida utilizaremos um *list comprehension* para fazer a leitura dos dados com o método do pandas, com isso podemos já inserir dentro de uma lista vazia criada, concatenando esses dois elementos da lista, pelas linhas, formando um arquivo único e por fim, visualizamos o resultado com as primeiras 5 linhas com o método *head()*.
#
# Temos 20 colunas ou *features* em nosso conjunto, vamos conhecer cada uma delas com o nosso **Dicionário de dados**:
#
# - **name:** nome da faixa
# - **album:** album que contém a faixa
# - **artist:** nome do artista
# - **release_date:** ano de lançamento da faixa
# - **length:** tempo de duração da música
# - **popularity:** quanto mais alto o valor, maior a popularidade
# - **track_id:** ID
# - **mode:** Modo indica a modalidade (maior ou menor) de uma faixa, o tipo de escala da qual seu conteúdo melódico é derivado
# - **acousticness:** quanto mais alto o valor, mais acústico é a música
# - **danceability:** quanto mais alto o valor, mais dançante é a música com base nos elementos musicais
# - **energy:** A energia da música, quanto mais alto o valor, mais enérgica é
# - **instrumentalness:** o valor de nível instrumental, se mais cantada ou sem vocal
# - **liveness:** presença de audiência, é a probabilidade da música ter tido público ou não
# - **valence:** quanto maior o valor, mais positiva (feliz, alegre, eufórico) a música é
# - **loudness:** quanto maior o valor, mais alto é a música
# - **speechiness:** quanto mais alto o valor, mais palavras cantadas a música possui
# - **tempo:** o tempo estimado em BPM (batida por minuto), relaciona com o ritmo derivando a duração média do tempo
# - **duration_ms:** a duração da faixa em milisegundos
# - **time_signature:** Uma estimativa de fórmula de compasso geral de uma faixa
# - **genre:** Genero do artista
# +
# identificando os arquivos .csv contidos na pasta
all_files = glob.glob("dados/*.csv")
# criando uma lista vazia para receber os dois arquivos
df_list = []
# fazendo um loop e armazenando os arquivos na pasta df_list
[df_list.append(pd.read_csv(filename, parse_dates = ['release_date'])) for filename in all_files]
# concatenando os dois arquivos para formar um
df = pd.concat(df_list, axis=0, ignore_index=True).drop('Unnamed: 0', axis=1)
# visualizando as primeiras 5 linhas
df.head()
# -
# Como extraimos também os *IDs* das músicas, que é a parte final da url, única para cada música, vamos juntar e formar a url completa.
# +
# definindo a url padrão
url_standard = 'https://open.spotify.com/track/'
# unindo com os IDs de cada música
df['url'] = url_standard + df['track_id']
# removendo a coluna dos IDs
df.drop(['track_id'], axis=1, inplace=True)
# visualizando as primeiras duas linhas
df.head(2)
# -
# Assim como as primeiras 5 linhas, vamos também visualizar as 5 últimas, com o método *tail()*
# visualizando as últimas 5 linhas
df.tail()
# Quais são as dimensões desse conjunto? O método `shape` pode nos ajudar a descobrir.
print(f"Quantidade de linhas: {df.shape[0]}")
print(f"Quantidade de colunas: {df.shape[1]}")
# ## Análise Exploratória dos dados
# Dados faltantes ou nulos, é uma característica normal para um conjunto de dados extraido no mundo real. Vamos dar uma olhada se esse conjunto possui algum.
# checando dados nulos
df.isnull().sum()
# Puxando os dados diretamente desta API, notamos que não foram trazidos nenhum dado faltante.
#
# Podemos notar que a maioria são dados numéricos, mas será que realmente estão nesse formato, vamos fazer um *check* utilizando o método *dtypes.
# checando os tipos de dados
df.dtypes
# Podemos ver que os formatos estão de acordo com o que deveriam ser.
#
# Vamos agora checar se há dados, ou melhor, linhas iguais ou duplicadas com o método *duplicated()*, caso encontramos vamos remover com o método *drop_duplicates* e checar as novas dimensões.
# checando dados duplicados
df.duplicated().sum()
# +
# removendo linhas duplicadas
df.drop_duplicates(inplace=True)
df = df.reset_index()
# -
# checando as novas dimensões
df.shape
# garantindo que não há mais dados duplicados
df.duplicated().sum()
# Vamos dar uma olhada em algumas estatísticas, para conhecermos mais nossos dados, e pra isso fazemos uso do método *describe()*.
# analisando estatísticas descritivas (numéricas)
df.describe()
# Como argumentos padrões do próprio método, foram retornadas somente das variáveis numéricas, mas também podemos fazer isso para as variáveis categóricas passando o tipo como argumento.
# analisando estatísticas descritivas (categóricas)
df.describe(include='O')
# Olhando somente para os números não são muito intuitivos, vamos então analisar graficamente, utilizado a biblioteca **seaborn**.<br>
# Primeiro, vamos criar um dataframe filtrando somente as variáveis de tipo numéricas, depois criaremos um loop para plotar os dados.
# +
# separando o conjunto de dados em tipos numéricos
df_num = df.select_dtypes(['float64', 'int64'])
# definindo a área de plotagem
plt.figure(figsize=(14,8))
# plotando os gráficos
for i in range(1, len(df_num.columns)):
ax = plt.subplot(3, 5, i)
sns.distplot(df_num[df_num.columns[i]])
ax.set_title(f'{df_num.columns[i]}')
ax.set_xlabel('')
# otimizando o espaçamento entre os gráficos
plt.tight_layout()
# -
# Podemos notar vários tipos de distribuição entre as variáveis e escalas diferentes. A princípio fizemos isso mais para saber como são as distribuições, não vamos analisar nada muito a fundo nesse momento.
#
# Vamos analisar também a correção entre essas variáveis, utilizando um `heatmap`.
# +
# definindo a área de plotagem
plt.figure(figsize=(14,8))
# plotando o gráfico
sns.heatmap(df_num.iloc[:, 1:].corr(), vmin=-1, vmax=1, annot=True).set_title('Correlação entre as variáveis');
# -
# Analisando as correlações, podemos notar que algumas variáveis se correlacionam fortemente, então não há problema em remover pelo menos uma delas, dependendo o caso o modelo pode ficar enviesado, vou optar em remover pelo menos uma, de cada duas correlactionadas fortemente.
df_num.drop(['length', 'loudness'], axis=1, inplace=True)
# ## Pré-processamento dos dados
# Para o pré-processamento dos dados, vamos utilizar o `MinMaxScaler` do **scikit-learn**. Vamos *normalizar* porque como vimos acima, nas distribuições dos dados, as escalas são muito diferentes o framework é sensível à isso, se não fizermos isso poderá dar um resultado bem diferente do esperado.
#
# O formato das distribuições continuarão as mesmas, porém estarão na mesma escala.
# +
# instanciando o tranformador
scaler = MinMaxScaler()
# treinando e transformando os dados
scaled = scaler.fit_transform(df_num)
# colocando os dados transformados em um dataframe
df_scaled = pd.DataFrame(scaled, columns=df_num.columns)
# olhando o resultado
df_scaled.head()
# -
# Vamos confirmar o que foi dito, sobre as distribuições e escalas.
# +
# definindo a área de plotagem
plt.figure(figsize=(14,8))
# plotando os gráficos
for i in range(1, len(df_scaled.columns)):
ax = plt.subplot(3, 5, i)
sns.distplot(df_scaled[df_scaled.columns[i]])
ax.set_title(f'{df_scaled.columns[i]}')
ax.set_xlabel('')
# otimizando o espaçamento entre os gráficos
plt.tight_layout()
# -
# Todas as escalas estão de 0 à 1.
# ## Machine Learning
#
# Agora vamos partir para o aprendizado do modelo, antes um breve introdução do algoritmo que será usado e como é seu funcionamento.
#
# ### Clustering
#
# **Clustering** é o conjunto de técnicas para análise de agrupamento de dados, que visa fazer agrupamentos automáticos de dados segundo o grau de semelhança. O algoritmo que vamos utilizar é o **K-Means**.
#
# #### K-Means
#
# Como funciona o algoritmo K-Means?<br>
# O K-Means agrupa os dados tentando separar as amostras em grupos de variancias iguais, minimizando um criterio conhecido como *inertia* ou *wcss (within-cluster sum-of-squares)*, em português, soma dos quadrados dentro do cluster, ou seja, minimizar essa soma dentro do cluster, quanto menor, melhor o agrupamento.
#
# Como definir a quantidade de grupos?<br>
# Uma técnica à se usar é a do **cotovelo**, com base na *inertia* ou *wcss*, onde definimos, basicamente, quando a diferença da *inertia* parar de ser significativa. Esse método compara a distância média de cada ponto até o centro do cluster para diferentes números de cluster.
#
# Além do método do cotovelo, para identificar o melhor número de clusters para nossos dados, podemos também utilizar inspeção visual, conhecimento prévio dos dados e do negócio e as vezes já temos até um número pré-definido, dependendo do objetivo.
#
# Exemplo de como funciona a técnica do cotovelo.
# 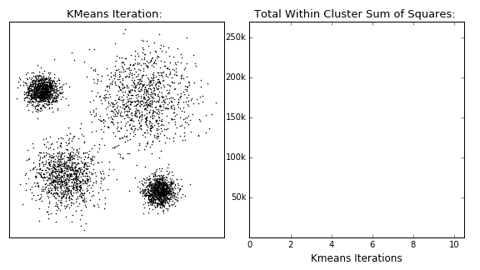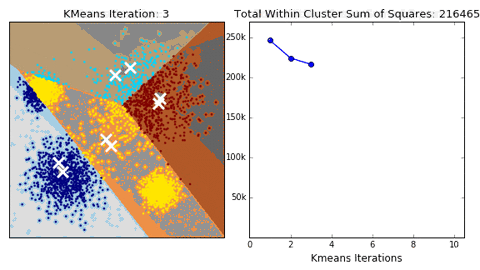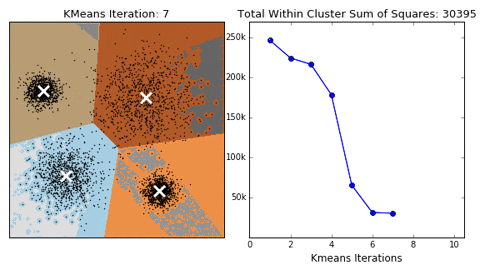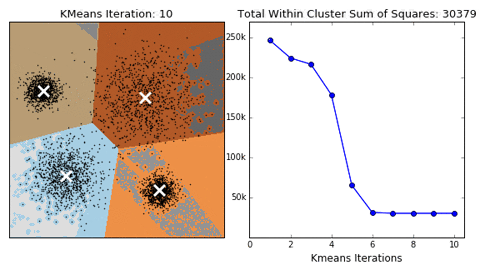
#
# Vamos começar!
# +
# criando uma lista vazia para inertia
wcss_sc = []
# criando o loop
for i in range(1, 50):
# instanciando o modelo
kmeans = KMeans(n_clusters=i, random_state=42)
# treinando o modelo
kmeans.fit(df_scaled.iloc[:, 1:])
# salvando os resultados
wcss_sc.append(kmeans.inertia_)
# -
# plotando o Elbow Method
plt.figure(figsize=(12,6))
plt.plot(range(1, 50), wcss_sc, 'o')
plt.plot(range(1, 50) , wcss_sc , '-' , alpha = 0.5)
plt.title('Elbow Method')
plt.xlabel('Number of Clusters')
plt.ylabel('WCSS')
# plt.savefig('Elbow_Method.png')
plt.show()
# Vamos fazer um teste com 20 grupos, pois podemos notar que a diferença não será mais muito significativa a partir desse valor. Claros que podemos testar com outros quantidades depois.
#
# Então, vamos instanciar o modelo que vamos usar, definindo o número correto de clusters, e vamos ver como os dados foram separados.
# +
# instanciando o modelo
kmeans = KMeans(n_clusters=20, random_state=42)
# treinando, excluindo a primeira linha que é o index
kmeans.fit(df_scaled.iloc[:, 1:])
# fazendo previsões
y_pred = kmeans.predict(df_scaled.iloc[:, 1:])
# +
# Visualizando os clusters em um dataframe
cluster_df = pd.DataFrame(y_pred, columns=['cluster'])
# visualizando a dimensão
cluster_df.shape
# -
# Vamos unir o dataframe original com os grupos que o algoritmo definiu para cada linha.
# +
# concatenando com o dataset original
df_new = pd.concat([df, cluster_df], axis=1)
# checando o dataframe
df_new.head()
# -
# olhando a nova dimensão
df_new.shape
# ## Analisando os clusters
#
# Vamos responder algumas perguntas, relacionadas ao resultado.
# ### Qual a média, desvio padrão, min e max dos elementos que compõe os clusters?
# agrupando por cluster e calculando a média
df_new.groupby('cluster')['name'].count().describe()
print(f"Temos em média {df_new.groupby('cluster')['name'].count().describe()['mean']:.2f} elementos por cluster, \
com um desvio padrão de {df_new.groupby('cluster')['name'].count().describe()['std']:.2f}.")
print(f"O Cluster com a menor quantidade de elementos possui {df_new.groupby('cluster')['name'].count().describe()['min']:.0f}\
e o maior possui {df_new.groupby('cluster')['name'].count().describe()['max']:.0f} elementos.")
# Podemos notar uma certa dispersão nos dados, isso pode ser devido à coleta de dados não balanceadas.
sns.distplot(df_new.groupby('cluster')['name'].count())
# Podemos notar que os grupos estão desbalanceados, precisaríamos levar em consideração as regras de negócios, para sabermos se os grupos criados estão adequados com base nas músicas, ritmos entre outros.
#
# Vamos dar uma olhada nos números de outros atributos.
# ### A popularidade entre os grupos seguem uma distribuição normal.
# agrupando por cluster e calculando a média
sns.distplot(df_new.groupby(['cluster'])['popularity'].mean())
df_new.groupby(['cluster'])['popularity'].mean().skew()
df_new.groupby(['cluster'])['popularity'].mean().kurtosis()
# Com os resultados de **skewness** e **kurtosis**, vemos que os dados não obedecem a uma distribuição normal, isso pode ser caracterizado por amostras de diferentes populações.
# ### Qual é o grupo com mais e menos elementos?
# checando o cluster com mais músicas
pd.DataFrame(df_new.cluster.value_counts()).reset_index()[:1].rename(columns={'index': 'cluster', 'cluster': 'qtd'})
# checando o cluster com menos músicas
pd.DataFrame(df_new.cluster.value_counts()).reset_index()[-1:].rename(columns={'index': 'cluster', 'cluster': 'qtd'})
# Vamos dar uma olhada nos dados dos grupos que contém o menor conjunto de elementos.
df_new[df_new.cluster == 12]
# Agora vamos salvar o dataframe com os respectivos grupos e para podemos analisar ou criar outras visualizações, com outros aplicativos como o *Power BI* ou *Tableau*, por exemplo.
#
# Além do dataframe, vamos salvar o nosso modelo com treinado para utilizarmos em outra aplicação.
# +
# # biblioteca que salvam os modelos
# import pickle
# # nome do modelo
# filename = 'model.pkl'
# # persistindo em disco local
# pickle.dump(kmeans, open(filename, 'wb'))
# # fazendo o download do modelo
# model_load = pickle.load(open("model.pkl", "rb"))
# # realizando as predições com o modelo treinado do disco
# model_load.predict(df_scaled.iloc[:, 1:])
# +
# Create a Pandas Excel writer using XlsxWriter as the engine.
writer = pd.ExcelWriter('grupo_musicas.xlsx', engine='xlsxwriter')
# Convert the dataframe to an XlsxWriter Excel object.
df_new.to_excel(writer, sheet_name='Sheet1', index=False)
# Close the Pandas Excel writer and output the Excel file.
writer.save()
# -
# ## Conclusão
# Criamos um framework para coleta dos dados direto da API do spotify e salvamos os dados em um arquivo *.csv*, fizemos algumas análises e pré-processamento dos dados.<br>
# Após criarmos um modelo de **clusterização**, o modelo fez as previsões para os grupos relacionados, colocamos todos em um dataframe e salvamos em um arquivo em excel.<br>
# Para os próximos passos, vamos criar uma *POC (Proof of Concept)* com o framework **streamlit** e publicar um dashboard criado com os dados em **Power BI**.
#
# link para dashboard do Power BI: https://bit.ly/3j2KCiU
#
# Há outros tipos de modelos para clusterização, mas utilizamos o KMeans por ser um dos mais utilizados e conseguir ter um bom resultado de forma fácil.
# ## Referencias
# https://developer.spotify.com/dashboard/applications/85bde5058f48488eb76c9a41fd7942eb<br>
# https://developer.spotify.com/documentation/web-api/reference/tracks/get-audio-features/<br>
# https://github.com/plamere/spotipy/tree/master/examples<br>
# https://morioh.com/p/31b8a607b2b0<br>
# https://medium.com/@maxtingle/getting-started-with-spotifys-api-spotipy-197c3dc6353b<br>
| Cluster spotify.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: fengine
# language: python
# name: fengine
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import OrdinalEncoder, StandardScaler
from feature_engine.wrappers import SklearnTransformerWrapper
from feature_engine.categorical_encoders import RareLabelCategoricalEncoder
# -
data = pd.read_csv('houseprice.csv')
data.head()
# +
# let's separate into training and testing set
X_train, X_test, y_train, y_test = train_test_split(
data.drop(['Id', 'SalePrice'], axis=1), data['SalePrice'], test_size=0.3, random_state=0)
X_train.shape, X_test.shape
# -
X_train[['LotFrontage', 'MasVnrArea']].isnull().mean()
# ## SimpleImputer
#
# ### Mean imputation
# +
imputer = SklearnTransformerWrapper(transformer = SimpleImputer(strategy='mean'),
variables = ['LotFrontage', 'MasVnrArea'])
imputer.fit(X_train)
# +
# we can find the mean values within the parameters of the
# simple imputer
imputer.transformer.statistics_
# +
X_train = imputer.transform(X_train)
X_test = imputer.transform(X_test)
X_train[['LotFrontage', 'MasVnrArea']].isnull().mean()
# -
# ### Frequent category imputation
cols = [c for c in data.columns if data[c].dtypes=='O' and data[c].isnull().sum()>0]
data[cols].head()
# +
imputer = SklearnTransformerWrapper(transformer = SimpleImputer(strategy='most_frequent'),
variables = cols)
imputer.fit(X_train)
# +
# we can find the most frequent values within the parameters of the
# simple imputer
imputer.transformer.statistics_
# +
X_train = imputer.transform(X_train)
X_test = imputer.transform(X_test)
X_train[cols].isnull().mean()
# -
# ## OrdinalEncoder
cols = ['Alley',
'MasVnrType',
'BsmtQual',
'BsmtCond',
'BsmtExposure',
'BsmtFinType1',
'BsmtFinType2',
'Electrical',
'FireplaceQu',
'GarageType',
'GarageFinish',
'GarageQual']
# +
# let's remove rare labels to avoid errors when encoding
rare_label_enc = RareLabelCategoricalEncoder(n_categories=2, variables=cols)
X_train = rare_label_enc.fit_transform(X_train)
X_test = rare_label_enc.transform(X_test)
# +
# now let's replace categories by integers
encoder = SklearnTransformerWrapper(transformer = OrdinalEncoder(),
variables = cols)
encoder.fit(X_train)
# +
# we can navigate to the parameters of the sklearn transformer
# like this:
encoder.transformer.categories_
# +
# encode categories
X_train = encoder.transform(X_train)
X_test = encoder.transform(X_test)
X_train[cols].isnull().mean()
# -
X_test[cols].head()
# ## Scaling
cols = [
'LotFrontage',
'MasVnrArea',
'Alley',
'MasVnrType',
'BsmtQual',
'BsmtCond',
'BsmtExposure',
'BsmtFinType1',
'BsmtFinType2',
'Electrical',
'FireplaceQu',
'GarageType',
'GarageFinish',
'GarageQual']
# +
# let's apply the standard scaler on the above variables
scaler = SklearnTransformerWrapper(transformer = StandardScaler(),
variables = cols)
scaler.fit(X_train)
# +
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
X_train[cols].isnull().mean()
# -
# mean values, learnt by the StandardScaler
scaler.transformer.mean_
# std values, learnt by the StandardScaler
scaler.transformer.scale_
# the mean of the scaled variables is 0
X_train[cols].mean()
# +
# the std of the scaled variables is ~1
X_train[cols].std()
# -
| examples/Sklearn-wrapper.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="exFeYM4KWlz9"
# ##### Copyright 2020 The TensorFlow Authors.
# + cellView="form" id="Oj6X6JHoWtVs"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="d5DZ2c-xfa9m"
# # TFF for Federated Learning Research: Model and Update Compression
#
# **NOTE**: This colab has been verified to work with the [latest released version](https://github.com/tensorflow/federated#compatibility) of the `tensorflow_federated` pip package, but the Tensorflow Federated project is still in pre-release development and may not work on `master`.
#
# In this tutorial, we use the [EMNIST](https://www.tensorflow.org/federated/api_docs/python/tff/simulation/datasets/emnist) dataset to demonstrate how to enable lossy compression algorithms to reduce communication cost in the Federated Averaging algorithm using the `tff.learning.build_federated_averaging_process` API and the [tensor_encoding](http://jakubkonecny.com/files/tensor_encoding.pdf) API. For more details on the Federated Averaging algorithm, see the paper [Communication-Efficient Learning of Deep Networks from Decentralized Data](https://arxiv.org/abs/1602.05629).
# + [markdown] id="qrPTFv7ngz-P"
# ## Before we start
#
# Before we start, please run the following to make sure that your environment is
# correctly setup. If you don't see a greeting, please refer to the
# [Installation](../install.md) guide for instructions.
# + id="X_JnSqDxlw5T"
#@test {"skip": true}
# !pip install --quiet --upgrade tensorflow_federated_nightly
# !pip install --quiet --upgrade tensorflow-model-optimization
# !pip install --quiet --upgrade nest_asyncio
import nest_asyncio
nest_asyncio.apply()
# %load_ext tensorboard
# + id="ctxIBpYIl846"
import functools
import numpy as np
import tensorflow as tf
import tensorflow_federated as tff
from tensorflow_model_optimization.python.core.internal import tensor_encoding as te
# + [markdown] id="wj-O1cnxKHMw"
# Verify if TFF is working.
# + id="8VPepVmfdhHv"
@tff.federated_computation
def hello_world():
return 'Hello, World!'
hello_world()
# + [markdown] id="30Pln72ihL-z"
# ## Preparing the input data
# In this section we load and preprocess the EMNIST dataset included in TFF. Please check out [Federated Learning for Image Classification](https://www.tensorflow.org/federated/tutorials/federated_learning_for_image_classification#preparing_the_input_data) tutorial for more details about EMNIST dataset.
#
# + id="oTP2Dndbl2Oe"
# This value only applies to EMNIST dataset, consider choosing appropriate
# values if switching to other datasets.
MAX_CLIENT_DATASET_SIZE = 418
CLIENT_EPOCHS_PER_ROUND = 1
CLIENT_BATCH_SIZE = 20
TEST_BATCH_SIZE = 500
emnist_train, emnist_test = tff.simulation.datasets.emnist.load_data(
only_digits=True)
def reshape_emnist_element(element):
return (tf.expand_dims(element['pixels'], axis=-1), element['label'])
def preprocess_train_dataset(dataset):
"""Preprocessing function for the EMNIST training dataset."""
return (dataset
# Shuffle according to the largest client dataset
.shuffle(buffer_size=MAX_CLIENT_DATASET_SIZE)
# Repeat to do multiple local epochs
.repeat(CLIENT_EPOCHS_PER_ROUND)
# Batch to a fixed client batch size
.batch(CLIENT_BATCH_SIZE, drop_remainder=False)
# Preprocessing step
.map(reshape_emnist_element))
emnist_train = emnist_train.preprocess(preprocess_train_dataset)
# + [markdown] id="XUQA55yjhTGh"
# ## Defining a model
#
# Here we define a keras model based on the orginial FedAvg CNN, and then wrap the keras model in an instance of [tff.learning.Model](https://www.tensorflow.org/federated/api_docs/python/tff/learning/Model) so that it can be consumed by TFF.
#
# Note that we'll need a **function** which produces a model instead of simply a model directly. In addition, the function **cannot** just capture a pre-constructed model, it must create the model in the context that it is called. The reason is that TFF is designed to go to devices, and needs control over when resources are constructed so that they can be captured and packaged up.
# + id="f2dLONjFnE2E"
def create_original_fedavg_cnn_model(only_digits=True):
"""The CNN model used in https://arxiv.org/abs/1602.05629."""
data_format = 'channels_last'
max_pool = functools.partial(
tf.keras.layers.MaxPooling2D,
pool_size=(2, 2),
padding='same',
data_format=data_format)
conv2d = functools.partial(
tf.keras.layers.Conv2D,
kernel_size=5,
padding='same',
data_format=data_format,
activation=tf.nn.relu)
model = tf.keras.models.Sequential([
tf.keras.layers.InputLayer(input_shape=(28, 28, 1)),
conv2d(filters=32),
max_pool(),
conv2d(filters=64),
max_pool(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(512, activation=tf.nn.relu),
tf.keras.layers.Dense(10 if only_digits else 62),
tf.keras.layers.Softmax(),
])
return model
# Gets the type information of the input data. TFF is a strongly typed
# functional programming framework, and needs type information about inputs to
# the model.
input_spec = emnist_train.create_tf_dataset_for_client(
emnist_train.client_ids[0]).element_spec
def tff_model_fn():
keras_model = create_original_fedavg_cnn_model()
return tff.learning.from_keras_model(
keras_model=keras_model,
input_spec=input_spec,
loss=tf.keras.losses.SparseCategoricalCrossentropy(),
metrics=[tf.keras.metrics.SparseCategoricalAccuracy()])
# + [markdown] id="ipfUaPLEhYYj"
# ## Training the model and outputting training metrics
#
# Now we are ready to construct a Federated Averaging algorithm and train the defined model on EMNIST dataset.
#
# First we need to build a Federated Averaging algorithm using the [tff.learning.build_federated_averaging_process](https://www.tensorflow.org/federated/api_docs/python/tff/learning/build_federated_averaging_process) API.
# + id="SAsGGkL9nHEl"
federated_averaging = tff.learning.build_federated_averaging_process(
model_fn=tff_model_fn,
client_optimizer_fn=lambda: tf.keras.optimizers.SGD(learning_rate=0.02),
server_optimizer_fn=lambda: tf.keras.optimizers.SGD(learning_rate=1.0))
# + [markdown] id="Mn1FAPQ32FcV"
# Now let's run the Federated Averaging algorithm. The execution of a Federated Learning algorithm from the perspective of TFF looks like this:
#
# 1. Initialize the algorithm and get the inital server state. The server state contains necessary information to perform the algorithm. Recall, since TFF is functional, that this state includes both any optimizer state the algorithm uses (e.g. momentum terms) as well as the model parameters themselves--these will be passed as arguments and returned as results from TFF computations.
# 2. Execute the algorithm round by round. In each round, a new server state will be returned as the result of each client training the model on its data. Typically in one round:
# 1. Server broadcast the model to all the participating clients.
# 2. Each client perform work based on the model and its own data.
# 3. Server aggregates all the model to produce a sever state which contains a new model.
#
# For more details, please see [Custom Federated Algorithms, Part 2: Implementing Federated Averaging](https://www.tensorflow.org/federated/tutorials/custom_federated_algorithms_2) tutorial.
#
# Training metrics are written to the Tensorboard directory for displaying after the training.
# + cellView="form" id="t5n9fXsGOO6-"
#@title Load utility functions
def format_size(size):
"""A helper function for creating a human-readable size."""
size = float(size)
for unit in ['bit','Kibit','Mibit','Gibit']:
if size < 1024.0:
return "{size:3.2f}{unit}".format(size=size, unit=unit)
size /= 1024.0
return "{size:.2f}{unit}".format(size=size, unit='TiB')
def set_sizing_environment():
"""Creates an environment that contains sizing information."""
# Creates a sizing executor factory to output communication cost
# after the training finishes. Note that sizing executor only provides an
# estimate (not exact) of communication cost, and doesn't capture cases like
# compression of over-the-wire representations. However, it's perfect for
# demonstrating the effect of compression in this tutorial.
sizing_factory = tff.framework.sizing_executor_factory()
# TFF has a modular runtime you can configure yourself for various
# environments and purposes, and this example just shows how to configure one
# part of it to report the size of things.
context = tff.framework.ExecutionContext(executor_fn=sizing_factory)
tff.framework.set_default_context(context)
return sizing_factory
# + id="jvH6qIgynI8S"
def train(federated_averaging_process, num_rounds, num_clients_per_round, summary_writer):
"""Trains the federated averaging process and output metrics."""
# Create a environment to get communication cost.
environment = set_sizing_environment()
# Initialize the Federated Averaging algorithm to get the initial server state.
state = federated_averaging_process.initialize()
with summary_writer.as_default():
for round_num in range(num_rounds):
# Sample the clients parcitipated in this round.
sampled_clients = np.random.choice(
emnist_train.client_ids,
size=num_clients_per_round,
replace=False)
# Create a list of `tf.Dataset` instances from the data of sampled clients.
sampled_train_data = [
emnist_train.create_tf_dataset_for_client(client)
for client in sampled_clients
]
# Round one round of the algorithm based on the server state and client data
# and output the new state and metrics.
state, metrics = federated_averaging_process.next(state, sampled_train_data)
# For more about size_info, please see https://www.tensorflow.org/federated/api_docs/python/tff/framework/SizeInfo
size_info = environment.get_size_info()
broadcasted_bits = size_info.broadcast_bits[-1]
aggregated_bits = size_info.aggregate_bits[-1]
print('round {:2d}, metrics={}, broadcasted_bits={}, aggregated_bits={}'.format(round_num, metrics, format_size(broadcasted_bits), format_size(aggregated_bits)))
# Add metrics to Tensorboard.
for name, value in metrics['train'].items():
tf.summary.scalar(name, value, step=round_num)
# Add broadcasted and aggregated data size to Tensorboard.
tf.summary.scalar('cumulative_broadcasted_bits', broadcasted_bits, step=round_num)
tf.summary.scalar('cumulative_aggregated_bits', aggregated_bits, step=round_num)
summary_writer.flush()
# + id="xp3o3QcBlqY_"
# Clean the log directory to avoid conflicts.
# !rm -R /tmp/logs/scalars/*
# Set up the log directory and writer for Tensorboard.
logdir = "/tmp/logs/scalars/original/"
summary_writer = tf.summary.create_file_writer(logdir)
train(federated_averaging_process=federated_averaging, num_rounds=10,
num_clients_per_round=10, summary_writer=summary_writer)
# + [markdown] id="zwdpTySt7pGQ"
# Start TensorBoard with the root log directory specified above to display the training metrics. It can take a few seconds for the data to load. Except for Loss and Accuracy, we also output the amount of broadcasted and aggregated data. Broadcasted data refers to tensors the server pushes to each client while aggregated data refers to tensors each client returns to the server.
# + id="EJ9XQiL-7e1i"
#@test {"skip": true}
# %tensorboard --logdir /tmp/logs/scalars/ --port=0
# + [markdown] id="rY5tWN_5ht6-"
# ## Build a custom broadcast and aggregate function
#
# Now let's implement function to use lossy compression algorithms on broadcasted data and aggregated data using the [tensor_encoding](http://jakubkonecny.com/files/tensor_encoding.pdf) API.
#
# First, we define two functions:
# * `broadcast_encoder_fn` which creates an instance of [te.core.SimpleEncoder](https://github.com/tensorflow/model-optimization/blob/ee53c9a9ae2e18ac1e443842b0b96229f0afb6d6/tensorflow_model_optimization/python/core/internal/tensor_encoding/core/simple_encoder.py#L30) to encode tensors or variables in server to client communication (Broadcast data).
# * `mean_encoder_fn` which creates an instance of [te.core.GatherEncoder](https://github.com/tensorflow/model-optimization/blob/ee53c9a9ae2e18ac1e443842b0b96229f0afb6d6/tensorflow_model_optimization/python/core/internal/tensor_encoding/core/gather_encoder.py#L30) to encode tensors or variables in client to server communicaiton (Aggregation data).
#
# It is important to note that we do not apply a compression method to the entire model at once. Instead, we decide how (and whether) to compress each variable of the model independently. The reason is that generally, small variables such as biases are more sensitive to inaccuracy, and being relatively small, the potential communication savings are also relatively small. Hence we do not compress small variables by default. In this example, we apply uniform quantization to 8 bits (256 buckets) to every variable with more than 10000 elements, and only apply identity to other variables.
# + id="lkRHkZTTnKn2"
def broadcast_encoder_fn(value):
"""Function for building encoded broadcast."""
spec = tf.TensorSpec(value.shape, value.dtype)
if value.shape.num_elements() > 10000:
return te.encoders.as_simple_encoder(
te.encoders.uniform_quantization(bits=8), spec)
else:
return te.encoders.as_simple_encoder(te.encoders.identity(), spec)
def mean_encoder_fn(value):
"""Function for building encoded mean."""
spec = tf.TensorSpec(value.shape, value.dtype)
if value.shape.num_elements() > 10000:
return te.encoders.as_gather_encoder(
te.encoders.uniform_quantization(bits=8), spec)
else:
return te.encoders.as_gather_encoder(te.encoders.identity(), spec)
# + [markdown] id="82iYUklQKP2e"
# TFF provides APIs to convert the encoder function into a format that `tff.learning.build_federated_averaging_process` API can consume. By using the `tff.learning.framework.build_encoded_broadcast_from_model` and `tff.learning.framework.build_encoded_mean_from_model`, we can create two functions that can be passed into `broadcast_process` and `aggregation_process` agruments of `tff.learning.build_federated_averaging_process` to create a Federated Averaging algorithms with a lossy compression algorithm.
# + id="aqD61hqAGZiW"
encoded_broadcast_process = (
tff.learning.framework.build_encoded_broadcast_process_from_model(
tff_model_fn, broadcast_encoder_fn))
encoded_mean_process = (
tff.learning.framework.build_encoded_mean_process_from_model(
tff_model_fn, mean_encoder_fn))
federated_averaging_with_compression = tff.learning.build_federated_averaging_process(
tff_model_fn,
client_optimizer_fn=lambda: tf.keras.optimizers.SGD(learning_rate=0.02),
server_optimizer_fn=lambda: tf.keras.optimizers.SGD(learning_rate=1.0),
broadcast_process=encoded_broadcast_process,
aggregation_process=encoded_mean_process)
# + [markdown] id="v3-ADI0hjTqH"
# ## Training the model again
#
# Now let's run the new Federated Averaging algorithm.
# + id="0KM_THYdn1yH"
logdir_for_compression = "/tmp/logs/scalars/compression/"
summary_writer_for_compression = tf.summary.create_file_writer(
logdir_for_compression)
train(federated_averaging_process=federated_averaging_with_compression,
num_rounds=10,
num_clients_per_round=10,
summary_writer=summary_writer_for_compression)
# + [markdown] id="sE8Bnjel8TIA"
# Start TensorBoard again to compare the training metrics between two runs.
#
# As you can see in Tensorboard, there is a significant reduction between the `orginial` and `compression` curves in the `broadcasted_bits` and `aggregated_bits` plots while in the `loss` and `sparse_categorical_accuracy` plot the two curves are pretty similiar.
#
# In conclusion, we implemented a compression algorithm that can achieve similar performance as the orignial Federated Averaging algorithm while the comminucation cost is significently reduced.
# + id="K9M2_1re28ff"
#@test {"skip": true}
# %tensorboard --logdir /tmp/logs/scalars/ --port=0
# + [markdown] id="Jaz9_9H7NUMW"
# ## Exercises
#
# To implement a custom compression algorithm and apply it to the training loop,
# you can:
#
# 1. Implement a new compression algorithm as a subclass of
# [`EncodingStageInterface`](https://github.com/tensorflow/model-optimization/blob/ee53c9a9ae2e18ac1e443842b0b96229f0afb6d6/tensorflow_model_optimization/python/core/internal/tensor_encoding/core/encoding_stage.py#L75)
# or its more general variant,
# [`AdaptiveEncodingStageInterface`](https://github.com/tensorflow/model-optimization/blob/ee53c9a9ae2e18ac1e443842b0b96229f0afb6d6/tensorflow_model_optimization/python/core/internal/tensor_encoding/core/encoding_stage.py#L274)
# following
# [this example](https://github.com/google-research/federated/blob/master/compression/sparsity.py).
# 1. Construct your new
# [`Encoder`](https://github.com/tensorflow/model-optimization/blob/ee53c9a9ae2e18ac1e443842b0b96229f0afb6d6/tensorflow_model_optimization/python/core/internal/tensor_encoding/core/core_encoder.py#L38)
# and specialize it for
# [model broadcast](https://github.com/google-research/federated/blob/master/compression/run_experiment.py#L95)
# or
# [model update averaging](https://github.com/tensorflow/federated/blob/e67590f284b487c6b889c070a96c35b8e0341e3b/tensorflow_federated/python/research/compression/run_experiment.py#L95).
# 1. Use those objects to build the entire
# [training computation](https://github.com/tensorflow/federated/blob/e67590f284b487c6b889c070a96c35b8e0341e3b/tensorflow_federated/python/research/compression/run_experiment.py#L204).
#
# Potentially valuable open research questions include: non-uniform quantization, lossless compression such as huffman coding, and mechanisms for adapting compression based on the information from previous training rounds.
#
# Recommended reading materials:
# * [Expanding the Reach of Federated Learning by Reducing Client Resource Requirements](https://research.google/pubs/pub47774/)
# * [Federated Learning: Strategies for Improving Communication Efficiency](https://research.google/pubs/pub45648/)
# * _Section 3.5 Communication and Compression_ in [Advanced and Open Problems in Federated Learning](https://arxiv.org/abs/1912.04977)
| site/en-snapshot/federated/tutorials/tff_for_federated_learning_research_compression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from zipfile import ZipFile
import datetime
zip_file = 'my_files.zip'
with ZipFile(zip_file, 'r') as zf:
for detail in zf.infolist():
| Twitter Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# Python functions for audit
# Verify if the script actually does the right job and there is no manual error introduced
# -
import csv
import json
import JobsMapResultsFilesToContainerObjs as ImageMap
import importlib
importlib.reload(ImageMap)
jsonObj = json.load(open("../data/experiment2_gid_aid_features.json"))
gidSpeciesList = []
for gid in jsonObj.keys():
if jsonObj[gid] != None:
gidSpecies = {}
for dct in jsonObj[gid]:
for aid in dct.keys():
gidSpecies[gid] = gidSpecies.get(gid,[]) + [dct[aid][2][0]]
gidSpeciesList.append(gidSpecies)
for dct in gidSpeciesList:
for speciesLst in dct.values():
firstEle = speciesLst[0]
for ele in speciesLst:
if ele != firstEle:
print(dct.keys())
def extractImageFeaturesFromMap(gidAidMapFl,aidFtrMapFl,feature):
aidFeatureDict = ImageMap.genAidFeatureDictDict(aidFtrMapFl)
gidAidDict = ImageMap.genAidGidDictFromMap(gidAidMapFl)
gidFeatureLst = []
for gid in gidAidDict:
if gidAidJson[gid]!= None:
gidFtr = {}
for aid in gidAidJson[gid]:
gidFtr[gid] = gidFtr.get(gid,[]) + [aidFeatureDict[str(aid)][feature]]
gidFeatureLst.append(gidFtr)
return gidFeatureLst
# +
aidFeatureDict = ImageMap.genAidFeatureDictDict("../data/experiment2_aid_features.json")
gidAidJson = ImageMap.genAidGidDictFromMap("../data/experiment2_gid_aid_map.json")
featuresPerImg = ImageMap.extractImageFeaturesFromMap("../data/experiment2_gid_aid_map.json","../data/experiment2_aid_features.json","SPECIES")
# -
shareCountLogic = {}
for gid in featuresPerImg.keys():
numInds = len(featuresPerImg[ele])
isHomogeneous = True
firstEle = featuresPerImg[ele][0]
for species in featuresPerImg[ele]:
if species != firstEle:
isHomogeneous = False
if isHomogeneous:
countFor = firstEle
else:
countFor = None
shareCountLogic[gid] = [numInds,isHomogeneous,countFor]
list(filter(lambda x: not x[2],l))
| script/.ipynb_checkpoints/AuditFncs-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn.cluster import DBSCAN
from sklearn.datasets import make_blobs
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
X_blob, y_blob = make_blobs(n_samples=500, centers=4, n_features=2, random_state=800)
plt.scatter(X_blob[:,0], X_blob[:,1])
plt.show()
# +
def scratch_DBSCAN(x, eps, min_pts):
"""
param x (list of vectors): your dataset to be clustered
param eps (float): neigborhood radius threshold
param min_pts (int): minimum number of points threshold for a nieghborhood to be a cluster
"""
# Build a label holder that is comprised of all 0s
labels = [0]* x.shape[0]
# Arbitrary starting "current cluster" ID
C = 0
# For each point p in x...
# ('p' is the index of the datapoint, rather than the datapoint itself.)
for p in range(0, x.shape[0]):
# Only unvisited points can be evaluated as neighborhood centers
if not (labels[p] == 0):
continue
# Find all of p's neighbors.
neighbors = neighborhood_search(x, p, eps)
# If there are not enough neighbor points, then it is classified as noise (-1).
# Otherwise we can use this point as a neighborhood cluster
if len(neighbors) < min_pts:
labels[p] = -1
else:
C += 1
neighbor_cluster(x, labels, p, neighbors, C, eps, min_pts)
return labels
def neighbor_cluster(x, labels, p, neighbors, C, eps, min_pts):
# Assign the cluster label to original point
labels[p] = C
# Look at each neighbor of p (by index, not the points themselves) and evaluate
i = 0
while i < len(neighbors):
# Get the next point from the queue.
potential_neighbor_ix = neighbors[i]
# If potential_neighbor_ix is noise from previous runs, we can assign it to current cluster
if labels[potential_neighbor_ix] == -1:
labels[potential_neighbor_ix] = C
# Otherwise, if potential_neighbor_ix is unvisited, we can add it to current cluster
elif labels[potential_neighbor_ix] == 0:
labels[potential_neighbor_ix] = C
# Further find neighbors of potential neighbor
potential_neighbors_cluster = neighborhood_search(x, potential_neighbor_ix, eps)
if len(potential_neighbors_cluster) >= min_pts:
neighbors = neighbors + potential_neighbors_cluster
# Evaluate next neighbor
i += 1
def neighborhood_search(x, p, eps):
neighbors = []
# For each point in the dataset...
for potential_neighbor in range(0, x.shape[0]):
# If a nearby point falls below the neighborhood radius threshold, add to neighbors list
if np.linalg.norm(x[p] - x[potential_neighbor]) < eps:
neighbors.append(potential_neighbor)
return neighbors
# -
labels = scratch_DBSCAN(X_blob, 0.6, 5)
plt.scatter(X_blob[:,0], X_blob[:,1], c=labels)
plt.title("DBSCAN from Scratch Performance")
plt.show()
| Lesson03/Activity04/Activity04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import birdvoxpaint as bvp
import matplotlib
# %matplotlib inline
from matplotlib import pyplot as plt
import numpy as np
filename = '/beegfs/vl1019/BOGOTA_NFCs_2018/MonserrateNFCs/SWIFT_20181106_235545.wav'
# -
S_raw = bvp.transform(filename=filename,
frame_length=2048, hop_length=512,
n_mels=None, fmin=None, fmax=None,
indices=[bvp.indices.maximum_pcen],
segment_duration=10,
verbose=True, n_jobs=None)
plt.figure(figsize=(15, 5))
librosa.display.specshow(S[0, :, :], cmap='gray')
import os
os.listdir("/beegfs/vl1019/")
# +
import librosa
import numpy as np
S = S_raw*1 + 0
S[0, :, :] = (S[0, :, :]-2/3) * 2
S[1, :, :] = (S[1, :, :]-0.05) * 5
S[2, :, :] = (S[2, :, :]-0.2) * 3
S = np.clip(S, 0, 1)
sr = librosa.get_samplerate(filename)
bvp.display.specshow(
np.transpose(S_raw, (2, 1, 0)),
sr=sr, normalize=True
)
# -
import librosa.display
plt.figure(figsize=(15, 5))
librosa.display.specshow(S_raw[1, :, :])
# +
import numpy as np
import matplotlib
# %matplotlib inline
from matplotlib import pyplot as plt
plt.hist(np.ravel(S_raw[0, :, :]-0.6)*4, bins=np.linspace(0, 1, 1000));
# +
S = S_raw*1 + 0
S[0, :, :] = (S[0, :, :]-2/3) * 2
S[1, :, :] = (S[1, :, :]-0.05) * 5
plt.hist(np.ravel(S[0, :, :]), bins=np.linspace(0, 1, 1000));
# -
plt.hist(np.ravel(S[2, :, :]), bins=np.linspace(0, 1, 1000));
# +
import librosa
from librosa.display import specshow
import numpy as np
import os
import soundfile as sf
segment_duration=10
sr = librosa.get_samplerate(filename)
segment_length = segment_duration*sr
total_duration = len(sf.SoundFile(filename)) / sr
tick_duration = 5 * 60
xticks = np.arange(tick_duration, total_duration, tick_duration)
plt.figure(figsize=(15, 5))
specshow(S[0, :, :], x_axis='time', sr=sr, hop_length=segment_length)
plt.xticks(xticks)
plt.title(os.path.split(filename)[1]);
plt.xlabel("Time (hh:mm:ss)")
# -
S_rgb = bvp.transform(filename=filename,
frame_length=2048, hop_length=512,
n_mels=None, fmin=None, fmax=None,
indices=[bvp.indices.towsey_rgb],
segment_duration=10,
verbose=True, n_jobs=None)
joblib.cpu_count()
# +
# if n_jobs is None or -1, parallelize across all available CPUs
if not n_jobs or n_jobs < 0:
n_jobs = joblib.cpu_count()
# measure sample rate and segment length
sr = sf.SoundFile(filename).samplerate
segment_length = segment_duration * sr
n_frames_per_segment = int(segment_length/frame_length)
# adjust segment_duration so that it matches the unit roundoff
# of the Euclidean division above
segment_duration = n_frames_per_segment * frame_length
# create a librosa generator object to loop through blocks
librosa_generator = librosa.core.stream(
filename, n_frames_per_segment, frame_length, hop_length)
# measure duration of the recording
total_duration = librosa.get_duration(filename=filename)
n_segments = int(total_duration / segment_duration)
# contruct tqdm generator from librosa generator
# this allows to display a progress bar
tqdm_generator = tqdm.tqdm(librosa_generator, total=n_segments, disable=not verbose)
# define frequency slicing function
# this function reduces the STFT or melspectrogram to a
# specific subband [fmin, fmax], measured in Hertz.
slice_fun = util.freq_slice(fmin, fmax, sr, frame_length)
# define spectrogram function.
spec_fun = partial(spec,
n_fft=frame_length, hop_length=hop_length,
win_length=frame_length, n_mels=n_mels,
sr=sr, fmin=fmin, fmax=fmax,
_fft_slice=slice_fun)
# define a closure for computing acoustic indices of a segment y.
indices_fun = lambda y: [np.stack(
[acoustic_index(S) for acoustic_index in indices], axis=-1)
for S in [spec_fun(y)]][0]
# delay execution of the closure above
delayed_indices_fun = joblib.delayed(indices_fun)
# construct joblib generator from delayed joblib object.
joblib_generator = (delayed_indices_fun(y) for y in tqdm_generator)
# construct joblib Parallel object.
parallel_fun = joblib.Parallel(n_jobs=n_jobs)
# execute
S = np.stack(parallel_fun(joblib_generator))
# -
S.shape
total_duration
# +
# get blocks from file
orig_sr = librosa.get_samplerate(filename)
sr = sr or orig_sr
# see: https://librosa.github.io/librosa/_modules/librosa/core/audio.html#stream
# block_length is in units of `frames` so reverse calculation
block_length = max(segment_duration * orig_sr, frame_length) * n_blocks
block_n_frames = librosa.core.samples_to_frames(
block_length, hop_length)
block_length = librosa.core.frames_to_samples(
block_n_frames, hop_length)
# -
| notebooks/02 - birdvoxpaint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:.conda-python3] *
# language: python
# name: conda-env-.conda-python3-py
# ---
# ## Since iron & myelin IQR/std need to be computed from the raw data, we need to re-combine everything
import pandas as pd
import seaborn as sns
import numpy as np
import scipy
import os
import glob
import statsmodels.api as sm
from statsmodels.stats import outliers_influence
import matplotlib.pyplot as plt
# %matplotlib inline
# # Load data
# ### 1. Demographics (for sex/age)
demo = pd.read_csv('../data/source_data/ahead_demographics.csv', sep=';')
demo = demo.loc[pd.notnull(demo['ScanName'])]
demo['Segmentation'] = demo['ScanName'].apply(lambda x: 'sub-' + x[-3:])
demo.head()
# ### 2. Intracranial volume (ICV) estimates
icv = pd.read_csv('../data/source_data/ahead-qmr2-icv-statistics.csv')
del icv['Template']
del icv['Intensity']
icv['Segmentation'] = icv['Segmentation'].apply(lambda x: x.split('_')[0])
whole_brain_volume = icv.rename(columns={'Label_1': 'icv'})
# ### 3. Cortex statistics
# We need to merge five files:
# 1. source_data/qmri-cruise-cortex-final-statistics.csv, which contains the volumes and thickness, as well as an older version of the qMRI statistics (with a clipping issue)
# 2. source_data/qmri-cruise-cortex-r1hz/r2hz/qpd/qsm-wb2-statistics.csv, which contain the qMRI values (after clipping issue was resolved)
# +
def split_intensity_name(x):
if 'thick' in x:
return 'thickness'
if 'iron' in x:
return 'iron'
if 'myelin' in x:
return 'myelin'
else:
return x.split('_')[3].split('-')[1]
# old values: use to extract 'thickness' and 'seg' from intensity
cortical = pd.read_csv('../data/source_data/qmri2-cruise-cortex-final-statistics.csv')
cortical = cortical.loc[cortical['Segmentation'].apply(lambda x: 'sub-' in x and '.nii.gz' in x)]
cortical = cortical.loc[cortical['Intensity'].apply(lambda x: ('sub-' in x and '.nii.gz' in x) or (x=='-'))]
cortical['ROI'] = cortical['Segmentation'].apply(lambda x: x.split('_')[1])
cortical['Segmentation'] = cortical['Segmentation'].apply(lambda x: x.split('_')[0])
cortical.Intensity = cortical.Intensity.replace({'-': 'sub-xxx_ses-x_acq-xx_mod-seg_std_brain'}) # just a trick
cortical.Intensity = cortical.Intensity.apply(split_intensity_name)
# remove qMRI-values from first file
cortical = cortical.loc[cortical.Intensity.isin(['seg', 'thickness'])]
# get qMRi-values
#all_csvs = sorted(glob.glob('../data/source_data/qmri2-cruise-cortex-*wb2-statistics.csv'))
all_csvs = sorted(glob.glob('../data/source_data/qmri2-cruise-cortex-*qmri2fcm-statistics.csv'))
#all_csvs = [x for x in all_csvs if not 'iron' in x and not 'myelin' in x] # NB these csvs are created after notebook 2
qmri_values_cruise = []
for csv in all_csvs:
qmri_values_cruise.append(pd.read_csv(csv))
qmri_values_cruise = pd.concat(qmri_values_cruise)
qmri_values_cruise = qmri_values_cruise.loc[qmri_values_cruise['Segmentation'].apply(lambda x: 'sub-' in x and '.nii.gz' in x)]
qmri_values_cruise = qmri_values_cruise.loc[qmri_values_cruise['Intensity'].apply(lambda x: ('sub-' in x and '.nii.gz' in x) or (x=='-'))]
qmri_values_cruise['ROI'] = qmri_values_cruise['Segmentation'].apply(lambda x: x.split('_')[1])
qmri_values_cruise['Segmentation'] = qmri_values_cruise['Segmentation'].apply(lambda x: x.split('_')[0])
qmri_values_cruise.Intensity = qmri_values_cruise.Intensity.apply(split_intensity_name)
# merge, clean-up
cortical = pd.concat([cortical, qmri_values_cruise])
cortical.columns = ['Measure', 'Segmentation', 'Template', 'Intensity', 'GM', 'WM_and_subcortex', 'ROI']
cortical_wide = cortical.pivot_table(values='GM', index=['Measure', 'Segmentation', 'Template', 'Intensity'], columns='ROI').reset_index()
del cortical_wide['Template']
cortical_wide = cortical_wide.rename(columns={'cb': "Cerebellum", 'lcr': 'Cortex L', 'rcr': 'Cortex R'})
#cortical_wide.to_csv('interim_data/CRUISE_combined.csv')
# -
# ### 4. qMRI metrics subcortex
# +
all_csvs = sorted(glob.glob('../data/source_data/ahead-31struct-qmri2fcm-*-statistics.csv'))
#all_csvs = [x for x in all_csvs if not 'iron' in x and not 'myelin' in x] # NB these csvs are created after notebook 2
data = []
for csv in all_csvs:
this_csv = pd.read_csv(csv)
if 'seg' in csv:
this_csv = this_csv.loc[this_csv.Measure == 'Voxels']
tmp = this_csv.copy()
tmp.Measure = 'Volume'
# to volume in mm3
for x in np.arange(1, 32, dtype=int):
tmp['Label_'+str(x)] = tmp['Label_'+str(x)] * (0.64*0.64*0.7)
this_csv = pd.concat([this_csv, tmp])
this_csv['Intensity'] = csv.split('-')[-2]
data.append(this_csv)
data = pd.concat(data)
col_names = ['STR L', 'STR R',
'STN L', 'STN R',
'SN L', 'SN R',
'RN L', 'RN R',
'GPi L', 'GPi R',
'GPe L', 'GPe R',
'THA L', 'THA R',
'LV L', 'LV R',
'3V', '4V',
'AMG L', 'AMG R',
'ic L', 'ic R',
'VTA L', 'VTA R',
'fx',
'PAG L', 'PAG R',
'PPN L', 'PPN R',
'CL L', 'CL R']
# col_names_orig = list(pd.read_csv('./old_data_not_denoised/ahead-35struct-simple-statistics-edited.csv').columns[3:-1])
# col_names = [x for x in col_names_orig if not 'ICO ' in x and not 'SCO ' in x] # ICO & SCO were dropped due to poor segmentation performance
# +
data.columns = ['Measure', 'Segmentation', 'Template', 'Intensity'] + col_names
data['Segmentation'] = data['Segmentation'].apply(lambda x: x.split('_')[0])
data = data.loc[data['Segmentation'] != 'sub-070'] # subject was dropped due to data quality issues
del data['Template']
# merge subcortex data with demographics
data = pd.merge(data, demo[['Sexe', 'Age', 'Segmentation']], on='Segmentation')
data = data.sort_values(['Measure', 'Segmentation', 'Intensity'])
data.head()
# coerce to float
for col in col_names:
data[col] = pd.to_numeric(data[col], errors='coerce')
#data.to_csv('./final_data/AHEAD-wide-combined.csv')
# -
# # merge Ahead with cortex data
# +
data = pd.merge(data, cortical_wide, how='outer')
col_names += ['Cerebellum', 'Cortex L', 'Cortex R']
# correct qpd
# qpd_idx = data['Intensity'] == 'qpd'
# def mean_notnull(x):
# return x[x>0].mean()
# ## Correct QPD by referencing to (mean across ventricles) ventricle-value
# mean_qpd = data.loc[qpd_idx][['VENT R', 'VENT L', 'VENT 3', 'VENT 4']].apply(mean_notnull, 1)
# data.loc[qpd_idx, col_names] /= np.tile(mean_qpd[:,np.newaxis], (1, len(col_names)))
# data.loc[(qpd_idx)]
data.head()
#data.to_csv('./final_data/AHEAD_and_CRUISE-wide-combined-qpdcorrected.csv')
# -
# ## Merge with ICV
# +
# icv = pd.read_csv('./ahead-qmr2-icv-statistics.csv')
# del icv['Template']
# del icv['Intensity']
# icv['Segmentation'] = icv['Segmentation'].apply(lambda x: x.split('_')[0])
# whole_brain_volume = icv.rename(columns={'Label_1': 'icv'})
ahead = pd.merge(data, whole_brain_volume.loc[whole_brain_volume.Measure=='Volume',['Segmentation', 'icv']])
# +
## To long format
# Cast all columns except Measure, Intensity, Segmentation to float
structs = []
for col in ahead.columns:
if col in ['Measure', 'Intensity', 'Segmentation', 'Age', 'Sexe', 'icv']:
continue
ahead[col] = ahead[col].astype(float)
structs.append(col)
# Melt
ahead_long = ahead.melt(id_vars=['Intensity', 'Measure', "Segmentation", 'Age', 'Sexe', 'icv'],
value_vars=structs, value_name='Value', var_name='ROI')
ahead_long['ROI2'] = ahead_long['ROI'].str.split(' ').apply(lambda x: x[0])
# ahead_long.loc['ROI2'] = ahead_long['ROI'].str.split(' ').apply(lambda x: x[0])
ahead_long['hemisphere'] = ahead_long['ROI'].str.split(' ').apply(lambda x: x[1] if len(x)>1 else 'X')
ahead_long['tissue_type'] = ahead_long['ROI2'].map({'STR': 'GM',
'STN': 'GM',
'SN': 'GM',
'RN': 'GM',
'GPi': 'GM',
'GPe': 'GM',
'THA': 'GM',
'LV': 'CSF',
'AMG': 'GM',
'ic': 'WM',
'VTA': 'GM',
'fx': 'WM',
'PAG': 'GM',
'PPN': 'GM',
'ICO': 'GM',
'SCO': 'GM',
'CL': 'GM',
'Cortex': 'GM',
'Cerebellum': 'GM'})
ahead_long.head()
# -
# save
ahead_long.to_csv('../data/final_data/AHEAD_and_CRUISE_and_ICV-combined-long_incliron.csv', index=False)
# +
# # Restructure
# ahead_wide = ahead_long.pivot_table(values='Value', index=['Segmentation', 'Measure', 'Age', 'Sexe', 'ROI', 'ROI2', 'hemisphere', 'tissue_type', 'icv'], columns='Intensity')
# ahead_wide = ahead_wide.reset_index()
# # Calculate IQR relative to median values
# tmp = ahead_wide.copy()
# tmp2 = tmp.loc[tmp['Measure'] == 'IQR_intensity'].copy()
# cols = ['iron', 'myelin', 'qpd', 'qsm', 'r1hz', 'r2hz', 'thickness']
# tmp2[cols] = tmp2[cols] / np.abs(tmp.loc[tmp['Measure'] == 'Median_intensity', cols].values)
# tmp2['Measure'] = 'IQR_relative_to_median'
# ahead_wide = pd.concat([ahead_wide, tmp2])
# # long again
# ahead_long = ahead_wide.reset_index().melt(id_vars=['Measure', "Segmentation", 'Age', 'Sexe', 'ROI', 'ROI2', 'hemisphere', 'icv'],
# value_vars=['iron', 'myelin', 'r1hz', 'r2hz', 'qsm', 'thickness'], value_name='Value', var_name='Intensity')
# ahead_long
# +
# # save
# ahead_long.to_csv('../data/final_data/AHEAD_and_CRUISE_and_ICV-combined-long.csv', index=False)
# -
| 3. Combine & prep data 2 (post-iron myelin).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# #How to GitHub
# ##The Concept
# Most people who have spent time writing code with others have heard at least one of the following: "GitHub", "Git", "Bitbucket," but what are these tools? The answer to this question depends on who you ask and how they are using them.
# The short answer requires us to take a step back. GitHub is technically a website. GitHub is defined as a webased tool for using Git. Git is an open source software that allows multiple people to work on a single web page. In short, git is the engine of collaboration, and GitHub is the vehicle. Git is a command line tool that allows for a push and pull between local files and a web repository.
# But what makes GitHub any better than using something like Google Collab, or simply sharing files? The catch all phrase is version control. The idea of version control is that a single piece of software is being built by multiple people at multiple times. When working amongst multiple collaborators you need a good way to keep track of the changes that each person makes and the different phases of work being done. You can see what this looks like below
# https://www.nobledesktop.com/image/blog/git-branches-merge.png
# ##Basic Terms
# The basic elements of a Git workflow are
#
# <ol>
# <li>Repository</li>
# <li>Branch</li>
# <li>Commit</li>
# <li>Pull</li>
# <li>Merge</li>
# </ol>
#
# An interactive tutorial of these basics can be found here
# https://guides.github.com/activities/hello-world/
# ###Repository
# A Repository is like a folder that contains the elements of a project such as folders, script files, data sets, videos or images, and anything else a project might need. A README.md is almost always contained within a repository to describe the repository's elements.
# ###Branch
# A branch is like a version of a project. Every repository has a parent branch typically called, 'main.' You can create a new branch by making a copy of the main branch at any point in time. If someone else is working on the main branchIf someone else is working on a different branch while you were doing your work? You can pull in those changes too.? You can pull in those changes too. A synonym for branching is forking!
# ###Fork
# A fork is quite similar to a branch, but for personal use. A fork is a brand new project that clones another project at a specific point in time. It makes an entire copy of a code base that you can branch to make changes. Many public code repositories have you create a fork to make changes so there is no risk of editing the real project. When a fork is merged back into a project, git will pull two full copies of the code.
# ###Commit
# A commit is like savin the changes to a branch. In Git, all saved changes can be tracked and accessed. It is extremely important that commits are well documented so that when collaborators look at your commits, they understand why you made the changes you did.
# ###Pull
# A pull is called, "The heart of Github," by the team. When you submit a pull request, it is not pulling code from the repository to your local machine, it is submitting your collaboration to be pulled into a branch of the project. A pull request shows the differences between the branch you're submitting it to and the submission!
# ###Merge
# A merge brings a pull request back into the main branch!
# ###Other Questions
# There are many other nuances to the vocabulary surrounding Git and GitHub. This blog is an excellent resource for explaining in layman's terms, some of those distinctions.
#
# https://stackoverflow.com/questions/3329943/what-are-the-differences-between-git-branch-fork-fetch-merge-rebase-and-clon
# ##Use Cases
# Collaborators around the world use GitHub for all kinds of things!!
# ##Tools to Use
# There are two disctinctions for the phrase, "Tools to use." There are tools that allow you to use GitHub. There are also tools that function as an alternative to GitHub but allow you to use Git!
# Many IDE's have integrations for GitHub. These integrations allow you to push and pull changes seamlessly. One of the most common integrations is the Visual Studio Code integrations. Sublime and Slack are two additional common tools that integrate with GitHub. Here is some excellent documentation on the open source integration between github and VSCode. This integration creates a GUI that allows you to push specific pieces of work and evaluate differences.
# https://github.com/microsoft/vscode-docs
# Alternatives to GitHub with their own sets of integrations and are TaraVault and Bitbucket. There are additional open source tools but exploring the pros and cons of these tools is a long journey. Keep in mind the specs of your project if you find yourself needing to choose a way to collaborate on code. Researching the types of code you intend to write, what other types of files are required in your project, what security limitations you have, etc.
# #Learning to Use GitHub
# There are some excellent tutorials availible online! The internal resources from github are excellent interactive tutorials for using the free online resource! They can be found at
# https://guides.github.com/
| content/docs/KnowledgeObjects/Fall2021/GitHubIntro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Olfaction Model Demo
# This notebook illustrates how to run a Neurokernel-based model of part of the fruit fly's antennal lobe.
# ### Background
# The early olfactory system in *Drosophila* consists of two antennal lobes,
# one on each side of the fly brain. Each of these LPUs contain 49 glomeruli that
# differ in functionality, size, shape, and relative position. Each glomerulus
# receives axons from about 50 olfactory receptor neurons (ORNs) on each of the
# fly's two antennae that express the same odorant receptor. The axons of each ORN
# connect to the dendrites of 3 to 5 projection neurons (PNs) in the glomeruli.
# In addition to the PNs - which transmit olfactory information to the higher
# regions of the brain - the antennal lobes contain local neurons (LNs) whose
# connections are restricted to the lobes; inter-glomerular connectivity therefore
# subsumes synaptic connections between ORNs and PNs, ORNs and LNs, LNs and PNs, and feedback
# from PNs to LNs. The entire early olfactory system in *Drosophila*
# contains approximately 4000 neurons.
#
# The current model of the each antennal lobe comprises 49 glomerular channels
# with full intraglomerular connectivity in both hemispheres of the fly brain. The
# entire model comprises 2800 neurons, or 70% of the fly's entire antennal
# lobe. All neurons in the system are modeled using the Leaky Integrate-and-Fire
# (LIF) model and all synaptic currents elicited by spikes are modeled using alpha functions.
# Parameters for 24 of the glomerular channels are based upon currently available
# ORN type data [(Hallem et al., 2006)](#hallem_coding_2006); all other channels are configured with
# artificial parameters.
#
# A script that generates a [GEXF](http://gexf.net) file containing the antennal lobe model configuration is included in the ``examples/olfaction/data`` subdirectory of the Neurokernel source code.
# ### Executing the Model
# Assuming that the Neurokernel source has been cloned to `~/neurokernel`, we first generate an input signal of duration 1.0 seconds and construct the LPU configuration:
# %cd -q ~/neurokernel/examples/olfaction/data
# %run gen_olf_input.py
# %run create_olf_gexf.py
# Next, we identify the indices of the olfactory sensory neurons (OSNs) and projection neurons (PNs) associated with a specific glomerulus; in this case, we focus on glomerulus DA1:
# +
import re
import networkx as nx
import neurokernel.tools.graph
g = nx.read_gexf('antennallobe.gexf.gz')
df_node, df_edge = neurokernel.tools.graph.graph_to_df(g)
glom_name = 'DA1'
osn_ind = sorted(list(set([ind[0] for ind in \
df_edge[df_edge.name.str.contains('.*-%s_.*' % glom_name)].index])))
pn_ind = sorted(list(set([ind[1] for ind in \
df_edge[df_edge.name.str.contains('.*-%s_.*' % glom_name)].index])))
# Get OSN and PN label indices:
osn_ind_labels = [int(re.search('osn_.*_(\d+)', name).group(1)) \
for name in df_node.ix[osn_ind].name]
pn_ind_labels = [int(re.search('.*_pn_(\d+)', name).group(1)) \
for name in df_node.ix[pn_ind].name]
# -
# We now execute the model:
# %cd -q ~/neurokernel/examples/olfaction
# %run olfaction_demo.py
# Next, we display the input odorant concentration profile and the spikes produced by the 25 OSNs and 3 PNs associated with glomerulus DA1 in the model:
# +
# %matplotlib inline
import h5py
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import numpy as np
fmt = lambda x, pos: '%2.2f' % (float(x)/1e4)
with h5py.File('./data/olfactory_input.h5', 'r') as fi, \
h5py.File('olfactory_output_spike.h5','r') as fo:
data_i = fi['array'].value
data_o = fo['array'].value
mpl.rcParams['figure.dpi'] = 120
mpl.rcParams['figure.figsize'] = (12,9)
raster = lambda data: plt.eventplot([np.nonzero(data[i, :])[0] for i in xrange(data.shape[0])],
colors = [(0, 0, 0)],
lineoffsets = np.arange(data.shape[0]),
linelengths = np.ones(data.shape[0])/2.0)
f = plt.figure()
plt.subplot(311)
ax = plt.gca()
ax.xaxis.set_major_formatter(ticker.FuncFormatter(fmt))
plt.plot(data_i[:10000, 0]);
ax.set_ylim(np.min(data_i)-1, np.max(data_i)+1)
ax.set_xlim(0, 10000)
plt.title('Input Stimulus'); plt.ylabel('Concentration')
plt.subplot(312)
raster(data_o.T[osn_ind, :])
plt.title('Spikes Generated by OSNs'); plt.ylabel('OSN #');
ax = plt.gca()
ax.set_ylim(np.min(osn_ind_labels), np.max(osn_ind_labels))
ax.xaxis.set_major_formatter(ticker.FuncFormatter(fmt))
ax.yaxis.set_major_locator(ticker.MultipleLocator(base=5.0))
plt.subplot(313)
raster(data_o.T[pn_ind, :])
plt.title('Spikes Generated by PNs'); plt.ylabel('PN #');
ax = plt.gca()
ax.set_ylim(np.min(pn_ind_labels)-0.5, np.max(pn_ind_labels)+0.5)
ax.xaxis.set_major_formatter(ticker.FuncFormatter(fmt))
ax.yaxis.set_major_locator(ticker.MultipleLocator(base=1.0))
plt.xlabel('time (s)')
plt.subplots_adjust()
f.savefig('olfactory_output.png')
# -
# ### Acknowledgements
# The olfactory model demonstrated in this notebook was developed by <NAME>.
# ### References
# <a name="hallem_coding_2006"></a><NAME>. and <NAME>. (2006), Coding of odors by a receptor repertoire, Cell, 125, 1, 143–160, doi:10.1016/j.cell.2006.01.050
| notebooks/olfaction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import dagstermill
# + tags=["parameters"]
from dagster import ModeDefinition, ResourceDefinition
from collections import namedtuple
url = 'postgresql://{username}:{password}@{hostname}:5432/{db_name}'.format(
username='test', password='<PASSWORD>', hostname='localhost', db_name='test'
)
DbInfo = namedtuple('DbInfo', 'url')
context = dagstermill.get_context(
mode_def=ModeDefinition(
resource_defs={'db_info': ResourceDefinition(lambda _: DbInfo(url))}
)
)
table_name = 'average_sfo_outbound_avg_delays_by_destination'
# -
db_url = context.resources.db_info.url
# +
import os
import sqlalchemy as sa
import matplotlib.pyplot as plt
import pandas as pd
from dagster.utils import mkdir_p
# -
engine = sa.create_engine(db_url)
from matplotlib.backends.backend_pdf import PdfPages
plots_path = os.path.join(os.getcwd(), 'plots')
mkdir_p(plots_path)
pdf_path = os.path.join(plots_path, 'sfo_delays_by_destination.pdf')
pp = PdfPages(pdf_path)
delays = pd.read_sql('select * from {table_name}'.format(table_name=table_name), engine)
delays.head()
plt.hist(delays['arrival_delay'], bins=100)
plt.title('Flight Delays (Origin SFO)')
plt.xlabel('Delay at Arrival (Minutes)')
plt.ylabel('Number of Flights')
pp.savefig()
plt.scatter(delays['departure_delay'], delays['arrival_delay'], alpha=.05)
plt.plot([-100,1400], [-100,1400], 'k:', alpha=0.75)
plt.title('Flight Delays (Origin SFO)')
plt.xlabel('Delay at Departure (Minutes)')
plt.ylabel('Delay at Arrival (Minutes)')
pp.savefig()
(delays['departure_delay'] - delays['arrival_delay']).describe()
# +
departure_delays_by_destination = delays.groupby('destination').mean()['departure_delay']
arrival_delays_by_destination = delays.groupby('destination').mean()['arrival_delay']
n_flights_by_destination = delays.groupby('destination').count()['origin']
import math
fig, ax = plt.subplots(figsize=(10,10))
# ax.scatter(
# departure_delays_by_destination,
# arrival_delays_by_destination,
# s=n_flights_by_destination.map(math.sqrt),
# alpha=0.5
# )
plt.xlim(0, 25)
plt.ylim(-5, 20)
plt.plot([-5,20], [-5,20], 'k-', alpha=0.5)
for i, destination in enumerate(n_flights_by_destination.index):
departure_delay = departure_delays_by_destination[i]
arrival_delay = arrival_delays_by_destination[i]
n_flights = n_flights_by_destination[i]
if (departure_delay > 0 and departure_delay < 25 and arrival_delay > -25 and departure_delay < 25 and n_flights > 500):
arrow_width = math.sqrt(n_flights/15000)
arrow_head_width = 2 * arrow_width
arrow_head_length = arrow_width
improved = (arrival_delay - departure_delay) < 0
annotation_y = arrival_delay + 0.3 if not improved else (arrival_delay - 0.3)
ax.arrow(
departure_delay,
departure_delay,
0,
arrival_delay - departure_delay,
width=arrow_width,
head_length=arrow_head_length,
head_width=arrow_head_width,
alpha=0.5,
length_includes_head=True)
ax.annotate(
destination,
(departure_delay, annotation_y),
horizontalalignment='center',
verticalalignment=('top' if improved else 'bottom'))
plt.title('Flight Delays (SFO to Destinations with > 500 Q2 Departures')
plt.xlabel('Average Delay at Departure by Destination (Minutes)')
plt.ylabel('Average Delay at Arrival by Destination (Minutes)')
pp.savefig()
# -
pp.close()
from dagster import LocalFileHandle
dagstermill.yield_result(LocalFileHandle(pdf_path))
| examples/dagster_examples/airline_demo/notebooks/SFO_Delays_by_Destination.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib notebook
import matplotlib.pyplot as plt
import numpy as np
import glob
import os
from fits_align.ident import make_transforms
from astropy.io import fits
from astropy import wcs
from astropy.units import degree
from astropy.coordinates import SkyCoord
indir = '/Users/egomez/Downloads/lcogtdata-20200116-213/'
img_list = glob.glob('/Users/egomez/Downloads/lcogtdata-20200116-213/*.fz')
# Make a large ndarray of all the data, sorted by flux
def data_sort(image):
with fits.open(image) as hdul:
b = np.sort(hdul[2].data, order=['flux'])
w = wcs.WCS(hdul[1].header)
ra, dec = w.wcs_pix2world(b['x'], b['y'], 1)
med = np.median(b['flux'])
a = b[b['flux']>0.5*med][::-1]
counts = a['flux']
countserr = a['fluxerr']
ref_a = np.array([a['flux'],a['x'], a['y']])
return ref_a
# Align the arrays
ref_image = img_list[0]
images_to_align = img_list[1:]
identifications = make_transforms(ref_image, images_to_align)
# +
# dt = [('flux',tmpa.dtype), ('x', tmpa.dtype)]
# assert tmpa.flags['C_CONTIGUOUS']
# phot_ref = tmpa.ravel().view(dt)
# phot_ref.sort(order=['flux','x'])
# -
photarray = []
old = []
for id in identifications:
a = data_sort(id.ukn.filepath)
coords = [a[1],a[2]]
inv = id.trans.inverse()
(matrix, offset) = inv.matrixform()
xy = np.dot(matrix,coords) - offset.reshape(2,1)
# ra, dec = w.wcs_pix2world(xy[0], xy[1], 1)
tmpa = np.array([a[0],xy[0],xy[1]])
photarray.append(tmpa)
a = data_sort(ref_image)
b = photarray[10].copy()
m = min(a.shape, b.shape)[1]
pa_align = []
# Check if some of the coordinates are switch (because the fluxes were close)
m_tot = 10000
for b in photarray:
b_t = b.T
m = min(a.shape, b.shape)[1]
m_tot = m if m < m_tot else m_tot
c = abs(a.T[:m]-b.T[:m])
for ind in range(2,m-1):
if c[ind,1]> 3:
for ia in range(0,4):
if np.abs(a.T[ind]-b_t[ind-ia])[1] < 3 and np.abs(a.T[ind]-b_t[ind-ia])[2] < 3:
b_t[[ind,ind-ia]] = b_t[[ind-ia,ind]]
continue
pa_align.append(b_t)
# How close are the 2 arrays to matching
# Make a map of the images to use by comparing the coordinate columns in each frame with the reference frame. Remove images which have less than 70 matching stars because these probably have things wrong with them.
cl = np.ones((m_tot, 3), dtype=bool)
count =0
phot = [a]
for b in pa_align:
if np.sum(np.isclose(a[1,:m_tot], b.T[1,:m_tot], atol=5)) < 70:
continue
count +=1
phot.append(b.T)
cl *= np.isclose(a.T[:m_tot], b[:m_tot], atol=5)
print(f"{np.sum(cl.T[1])}:{np.sum(cl.T[2])} comparison stars")
print(f"{count/len(pa_align):.2f} % of files used")
# for i in range(0,len(photarray)):
# plt.scatter(photarray[i],marker='.',c='blue',alpha=0.005)
plt.scatter(phot_ref,phot_ref,marker='.',c='red',alpha=0.5)
| docs/Catalogue Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import sys
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import json
from os.path import expanduser
import pickle
import random
import time
import tensorflow as tf
from tensorflow import keras
from keras.models import Sequential
from keras.layers import Dense, BatchNormalization, Dropout
from tensorflow.keras import layers
from tensorflow.keras.layers.experimental import preprocessing
# +
run_name = 'P3856_YHE211_1_Slot1-1_1_5104'
experiment_name = 'P3856_YHE211'
EXPERIMENT_DIR = '/media/big-ssd/experiments/{}'.format(experiment_name)
# RESULTS_DIR = '/media/big-ssd/results-P3856'
# IDENTS_PASEF_DIR = '{}/P3856-results-cs-true-fmdw-true-2021-05-20-02-44-34/identifications-pasef'.format(RESULTS_DIR)
IDENTS_PASEF_DIR = '{}/identifications-pasef'.format(EXPERIMENT_DIR)
IDENTS_PASEF_FILE = '{}/exp-{}-identifications-pasef-recalibrated.feather'.format(IDENTS_PASEF_DIR, experiment_name)
FEATURES_PASEF_DIR = '{}/features-pasef'.format(EXPERIMENT_DIR)
FEATURES_PASEF_FILE = '{}/exp-{}-run-{}-features-pasef-dedup.feather'.format(FEATURES_PASEF_DIR, experiment_name, run_name)
# -
# define a straight line to exclude the charge-1 cloud
def scan_coords_for_single_charge_region(mz_lower, mz_upper):
scan_for_mz_lower = max(int(-1 * ((1.2 * mz_lower) - 1252)), 0)
scan_for_mz_upper = max(int(-1 * ((1.2 * mz_upper) - 1252)), 0)
return {'scan_for_mz_lower':scan_for_mz_lower, 'scan_for_mz_upper':scan_for_mz_upper}
input_names = ['deconvolution_score','coelution_coefficient','mobility_coefficient','isotope_count']
# #### identified features
MAXIMUM_Q_VALUE = 0.01
# load the features identified
idents_df = pd.read_feather(IDENTS_PASEF_FILE)
idents_df = idents_df[(idents_df.run_name == run_name) & (idents_df['percolator q-value'] <= MAXIMUM_Q_VALUE)].copy()
idents_df['excluded'] = idents_df.apply(lambda row: row.scan_apex < scan_coords_for_single_charge_region(row.monoisotopic_mz, row.monoisotopic_mz)['scan_for_mz_lower'], axis=1)
idents_df = idents_df[(idents_df.excluded == False)]
print('{} identifications'.format(len(idents_df)))
# #### detected features
# load the features detected by PASEF
features_df = pd.read_feather(FEATURES_PASEF_FILE)
features_df['excluded'] = features_df.apply(lambda row: row.scan_apex < scan_coords_for_single_charge_region(row.monoisotopic_mz, row.monoisotopic_mz)['scan_for_mz_lower'], axis=1)
features_df = features_df[(features_df.excluded == False)]
print('{} features detected'.format(len(features_df)))
sets_d = {'detected':set(), 'identified':set()}
sets_d['detected'] = set(features_df.feature_id.tolist())
sets_d['identified'] = set(idents_df.feature_id.tolist()) - (set(idents_df.feature_id.tolist()) - set(features_df.feature_id.tolist()))
# +
import matplotlib.pyplot as plt
from matplotlib_venn import venn2
f, ax1 = plt.subplots()
f.set_figheight(10)
f.set_figwidth(15)
plt.margins(0.06)
plt.title('features detected and identified\nexperiment {}, run {}'.format(experiment_name, run_name))
venn2([sets_d['detected'],sets_d['identified']], ('detected','identified'))
plt.show()
# -
# #### build the training set
# features detected but not identified
features_not_identified = sets_d['detected'] - sets_d['identified']
features_identified = sets_d['identified']
len(features_not_identified), len(features_identified)
features_not_identified_df = features_df[features_df.feature_id.isin(features_not_identified)]
features_identified_df = features_df[features_df.feature_id.isin(features_identified)]
SAMPLE_SIZE = 8500
# +
features_not_identified_df = features_not_identified_df.sample(n=SAMPLE_SIZE)
features_not_identified_df['category'] = 0
features_identified_df = features_identified_df.sample(n=SAMPLE_SIZE)
features_identified_df['category'] = 1
# -
combined_df = pd.concat([features_not_identified_df, features_identified_df], axis=0, sort=False, ignore_index=True)
combined_df.isna().sum()
combined_df.fillna(0, inplace=True)
feature_ids_l = combined_df.feature_id.tolist()
train_proportion = 0.8
val_proportion = 0.1
train_n = round(len(feature_ids_l) * train_proportion)
val_n = round(len(feature_ids_l) * val_proportion)
train_set = random.sample(feature_ids_l, train_n)
val_test_set = list(set(feature_ids_l) - set(train_set))
val_set = random.sample(val_test_set, val_n)
test_set = list(set(val_test_set) - set(val_set))
train_df = combined_df[combined_df.feature_id.isin(train_set)]
valid_df = combined_df[combined_df.feature_id.isin(val_set)]
test_df = combined_df[combined_df.feature_id.isin(test_set)]
train_df.to_pickle('{}/train_df.pkl'.format(expanduser('~')))
valid_df.to_pickle('{}/valid_df.pkl'.format(expanduser('~')))
test_df.to_pickle('{}/test_df.pkl'.format(expanduser('~')))
print('train: {} ({}%), validation: {} ({}%), test: {} ({}%)'.format(len(train_df), round(len(train_df)/len(combined_df)*100), len(test_df), round(len(test_df)/len(combined_df)*100), len(valid_df), round(len(valid_df)/len(combined_df)*100)))
X_train = train_df[input_names].to_numpy()
y_train = train_df[['category']].to_numpy()[:,0]
X_valid = valid_df[input_names].to_numpy()
y_valid = valid_df[['category']].to_numpy()[:,0]
X_test = test_df[input_names].to_numpy()
y_test = test_df[['category']].to_numpy()[:,0]
# define the keras model
model = Sequential()
model.add(BatchNormalization(input_shape=(4,)))
model.add(Dense(200, activation='relu'))
model.add(Dropout(0.4))
model.add(BatchNormalization())
model.add(Dense(200, activation='relu'))
model.add(Dropout(0.4))
model.add(BatchNormalization())
model.add(Dense(200, activation='relu'))
model.add(Dropout(0.4))
model.add(BatchNormalization())
model.add(Dense(1, activation='sigmoid'))
# compile the keras model
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['binary_accuracy'])
model.summary()
# for visualisation
tf.keras.models.save_model(model, "{}/model.h5".format(expanduser('~')))
# fit the keras model on the dataset
history = model.fit(X_train,
y_train,
validation_data=(X_valid, y_valid),
epochs=4000,
batch_size=512,
verbose=0)
history_df = pd.DataFrame(history.history)
history_df.loc[5:, ['loss', 'val_loss']].plot()
history_df.loc[5:, ['binary_accuracy', 'val_binary_accuracy']].plot()
print(("best validation loss: {:0.4f}"+"\nbest validation accuracy: {:0.4f}").format(history_df['val_loss'].min(), history_df['val_binary_accuracy'].max()))
# #### evaluate the model on the test data
results = model.evaluate(X_test, y_test, batch_size=128)
print("test loss, test acc:", results)
# #### use the model to classify the 3DID features
experiment_name = 'P3856'
EXPERIMENT_DIR = '/media/big-ssd/experiments/{}'.format(experiment_name)
MODEL_DIR = '{}/features-3did-classifier'.format(EXPERIMENT_DIR)
# #### save the model
model.save(MODEL_DIR)
| notebooks/papers/3did/train the feature classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="CxmDMK4yupqg"
# # Object Detection
#
# + [markdown] id="Sy553YSVmYiK"
# This lab is similar to the previous lab, except now instead of printing out the bounding box coordinates, you can visualize these bounding boxes on top of the image!
# + [markdown] id="v4XGxDrCkeip"
# ## Setup
#
# + id="6cPY9Ou4sWs_"
# For running inference on the TF-Hub module.
import tensorflow as tf
import tensorflow_hub as hub
# For downloading the image.
import matplotlib.pyplot as plt
import tempfile
from six.moves.urllib.request import urlopen
from six import BytesIO
# For drawing onto the image.
import numpy as np
from PIL import Image
from PIL import ImageColor
from PIL import ImageDraw
from PIL import ImageFont
from PIL import ImageOps
# For measuring the inference time.
import time
# Print Tensorflow version
print(tf.__version__)
# Check available GPU devices.
print("The following GPU devices are available: %s" % tf.test.gpu_device_name())
# + [markdown] id="t-VdfLbC1w51"
# ### Select and load the model
# As in the previous lab, you can choose an object detection module. Here are two that we've selected for you:
# * [ssd + mobilenet V2](https://tfhub.dev/tensorflow/ssd_mobilenet_v2/2) small and fast.
# * [FasterRCNN + InceptionResNet V2](https://tfhub.dev/google/faster_rcnn/openimages_v4/inception_resnet_v2/1): high accuracy
# + id="uazJ5ASc2_QE"
# you can switch the commented lines here to pick the other model
# ssd mobilenet version 2
module_handle = "https://tfhub.dev/google/openimages_v4/ssd/mobilenet_v2/1"
# You can choose inception resnet version 2 instead
#module_handle = "https://tfhub.dev/google/faster_rcnn/openimages_v4/inception_resnet_v2/1"
# + [markdown] id="xRVwVr40grw6"
# #### Load the model
#
# Next, you'll load the model specified by the `module_handle`.
# - This will take a few minutes to load the model.
# + id="MsFSuo2Wgrw6"
model = hub.load(module_handle)
# + [markdown] id="0gXxP2xYgrw6"
# #### Choose the default signature
#
# As before, you can check the available signatures using `.signature.keys()`
# + id="VGkmPO1lgrw6"
# take a look at the available signatures for this particular model
model.signatures.keys()
# + [markdown] id="VdwgkV7igrw6"
# Please choose the 'default' signature for your object detector.
# + id="IJ6plWWUgrw6"
detector = model.signatures['default']
# + [markdown] id="o_aN2zXggrw6"
# ### download_and_resize_image
#
# As you saw in the previous lab, this function downloads an image specified by a given "url", pre-processes it, and then saves it to disk.
# - What new compared to the previous lab is that you an display the image if you set the parameter `display=True`.
# + id="gjmOWdPpgrw6"
def display_image(image):
"""
Displays an image inside the notebook.
This is used by download_and_resize_image()
"""
fig = plt.figure(figsize=(20, 15))
plt.grid(False)
plt.imshow(image)
def download_and_resize_image(url, new_width=256, new_height=256, display=False):
'''
Fetches an image online, resizes it and saves it locally.
Args:
url (string) -- link to the image
new_width (int) -- size in pixels used for resizing the width of the image
new_height (int) -- size in pixels used for resizing the length of the image
Returns:
(string) -- path to the saved image
'''
# create a temporary file ending with ".jpg"
_, filename = tempfile.mkstemp(suffix=".jpg")
# opens the given URL
response = urlopen(url)
# reads the image fetched from the URL
image_data = response.read()
# puts the image data in memory buffer
image_data = BytesIO(image_data)
# opens the image
pil_image = Image.open(image_data)
# resizes the image. will crop if aspect ratio is different.
pil_image = ImageOps.fit(pil_image, (new_width, new_height), Image.ANTIALIAS)
# converts to the RGB colorspace
pil_image_rgb = pil_image.convert("RGB")
# saves the image to the temporary file created earlier
pil_image_rgb.save(filename, format="JPEG", quality=90)
print("Image downloaded to %s." % filename)
if display:
display_image(pil_image)
return filename
# + [markdown] id="2m5qn23ogrw6"
# ### Select and load an image
# Load a public image from Open Images v4, save locally, and display.
# + id="ntMGNQldgrw6"
# By <NAME>, Source: https://commons.wikimedia.org/wiki/File:Naxos_Taverna.jpg
image_url = "https://upload.wikimedia.org/wikipedia/commons/6/60/Naxos_Taverna.jpg" #@param
downloaded_image_path = download_and_resize_image(image_url, 1280, 856, True)
# + [markdown] id="nA<KEY>"
# ### Draw bounding boxes
#
# To build on what you saw in the previous lab, you can now visualize the predicted bounding boxes, overlaid on top of the image.
# - You can use `draw_boxes` to do this. It will use `draw_bounding_box_on_image` to draw the bounding boxes.
# + id="J5rUpPPqgrw7"
def draw_bounding_box_on_image(image,
ymin,
xmin,
ymax,
xmax,
color,
font,
thickness=4,
display_str_list=()):
"""
Adds a bounding box to an image.
Args:
image -- the image object
ymin -- bounding box coordinate
xmin -- bounding box coordinate
ymax -- bounding box coordinate
xmax -- bounding box coordinate
color -- color for the bounding box edges
font -- font for class label
thickness -- edge thickness of the bounding box
display_str_list -- class labels for each object detected
Returns:
No return. The function modifies the `image` argument
that gets passed into this function
"""
draw = ImageDraw.Draw(image)
im_width, im_height = image.size
# scale the bounding box coordinates to the height and width of the image
(left, right, top, bottom) = (xmin * im_width, xmax * im_width,
ymin * im_height, ymax * im_height)
# define the four edges of the detection box
draw.line([(left, top), (left, bottom), (right, bottom), (right, top),
(left, top)],
width=thickness,
fill=color)
# If the total height of the display strings added to the top of the bounding
# box exceeds the top of the image, stack the strings below the bounding box
# instead of above.
display_str_heights = [font.getsize(ds)[1] for ds in display_str_list]
# Each display_str has a top and bottom margin of 0.05x.
total_display_str_height = (1 + 2 * 0.05) * sum(display_str_heights)
if top > total_display_str_height:
text_bottom = top
else:
text_bottom = top + total_display_str_height
# Reverse list and print from bottom to top.
for display_str in display_str_list[::-1]:
text_width, text_height = font.getsize(display_str)
margin = np.ceil(0.05 * text_height)
draw.rectangle([(left, text_bottom - text_height - 2 * margin),
(left + text_width, text_bottom)],
fill=color)
draw.text((left + margin, text_bottom - text_height - margin),
display_str,
fill="black",
font=font)
text_bottom -= text_height - 2 * margin
def draw_boxes(image, boxes, class_names, scores, max_boxes=10, min_score=0.1):
"""
Overlay labeled boxes on an image with formatted scores and label names.
Args:
image -- the image as a numpy array
boxes -- list of detection boxes
class_names -- list of classes for each detected object
scores -- numbers showing the model's confidence in detecting that object
max_boxes -- maximum detection boxes to overlay on the image (default is 10)
min_score -- minimum score required to display a bounding box
Returns:
image -- the image after detection boxes and classes are overlaid on the original image.
"""
colors = list(ImageColor.colormap.values())
try:
font = ImageFont.truetype("/usr/share/fonts/truetype/liberation/LiberationSansNarrow-Regular.ttf",
25)
except IOError:
print("Font not found, using default font.")
font = ImageFont.load_default()
for i in range(min(boxes.shape[0], max_boxes)):
# only display detection boxes that have the minimum score or higher
if scores[i] >= min_score:
ymin, xmin, ymax, xmax = tuple(boxes[i])
display_str = "{}: {}%".format(class_names[i].decode("ascii"),
int(100 * scores[i]))
color = colors[hash(class_names[i]) % len(colors)]
image_pil = Image.fromarray(np.uint8(image)).convert("RGB")
# draw one bounding box and overlay the class labels onto the image
draw_bounding_box_on_image(image_pil,
ymin,
xmin,
ymax,
xmax,
color,
font,
display_str_list=[display_str])
np.copyto(image, np.array(image_pil))
return image
# + [markdown] id="r5rVOWRSgrw7"
# ### run_detector
#
# This function will take in the object detection model `detector` and the path to a sample image, then use this model to detect objects.
# - This time, run_dtector also calls `draw_boxes` to draw the predicted bounding boxes.
# + id="twlQG6TPgrw7"
def load_img(path):
'''
Loads a JPEG image and converts it to a tensor.
Args:
path (string) -- path to a locally saved JPEG image
Returns:
(tensor) -- an image tensor
'''
# read the file
img = tf.io.read_file(path)
# convert to a tensor
img = tf.image.decode_jpeg(img, channels=3)
return img
def run_detector(detector, path):
'''
Runs inference on a local file using an object detection model.
Args:
detector (model) -- an object detection model loaded from TF Hub
path (string) -- path to an image saved locally
'''
# load an image tensor from a local file path
img = load_img(path)
# add a batch dimension in front of the tensor
converted_img = tf.image.convert_image_dtype(img, tf.float32)[tf.newaxis, ...]
# run inference using the model
start_time = time.time()
result = detector(converted_img)
end_time = time.time()
# save the results in a dictionary
result = {key:value.numpy() for key,value in result.items()}
# print results
print("Found %d objects." % len(result["detection_scores"]))
print("Inference time: ", end_time-start_time)
# draw predicted boxes over the image
image_with_boxes = draw_boxes(
img.numpy(), result["detection_boxes"],
result["detection_class_entities"], result["detection_scores"])
# display the image
display_image(image_with_boxes)
# + [markdown] id="TyB3hroOgrw7"
# ### Run the detector on your selected image!
# + id="vchaUW1XDodD"
run_detector(detector, downloaded_image_path)
# + [markdown] id="WUUY3nfRX7VF"
# ### Run the detector on more images
# Perform inference on some additional images of your choice and check how long inference takes.
# + id="rubdr2JXfsa1"
image_urls = [
# Source: https://commons.wikimedia.org/wiki/File:The_Coleoptera_of_the_British_islands_(Plate_125)_(8592917784).jpg
"https://upload.wikimedia.org/wikipedia/commons/1/1b/The_Coleoptera_of_the_British_islands_%28Plate_125%29_%288592917784%29.jpg",
# By <NAME>, Source: https://commons.wikimedia.org/wiki/File:Biblioteca_Maim%C3%B3nides,_Campus_Universitario_de_Rabanales_007.jpg
"https://upload.wikimedia.org/wikipedia/commons/thumb/0/0d/Biblioteca_Maim%C3%B3nides%2C_Campus_Universitario_de_Rabanales_007.jpg/1024px-Biblioteca_Maim%C3%B3nides%2C_Campus_Universitario_de_Rabanales_007.jpg",
# Source: https://commons.wikimedia.org/wiki/File:The_smaller_British_birds_(8053836633).jpg
"https://upload.wikimedia.org/wikipedia/commons/0/09/The_smaller_British_birds_%288053836633%29.jpg",
]
def detect_img(image_url):
start_time = time.time()
image_path = download_and_resize_image(image_url, 640, 480)
run_detector(detector, image_path)
end_time = time.time()
print("Inference time:",end_time-start_time)
# + id="otPnrxMKIrj5"
detect_img(image_urls[0])
# + id="H5F7DkD5NtOx"
detect_img(image_urls[1])
# + id="DZ18R7dWNyoU"
detect_img(image_urls[2])
| Advanced Computer Vision with TensorFlow/Week 2 - Object Detection/Copy of C3_W2_Lab_2_Object_Detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Slicing the data
# ## 1. Introduction
#
# The most important number in the COMPAS data is the seed. The seed represents the unique identifier to a specific system in a simulation. Therefore the properties of a single system can be recovered by looking at seeds in different files.
#
# Here we introduce the basics of manipulating the data using the seeds. We provide an example on how we get the initial parameters of systems that ended up forming double compact objects (DCOs).
#
# Naively, we might try to use loops with conditions to extract systems of interest to a list. However, this can potentially be computationally expensive.
#
# Here we present a method to more efficiently `slice` the data using boolean masks. These are slightly more involved but are computationally quick and use intuitive logic.
#
# If you do not already have a ``COMPAS_Output.h5`` file ready, you can download some data from [compas.science](https://compas.science/)
# ***Note:* These cells may take a long time if you test them on large datasets.**
# ### 1.1 Paths
#
pathToData = '../COMPAS_Output.h5'
# ### 1.2 Imports
#python libraries
import numpy as np # for handling arrays
import h5py as h5 # for reading the COMPAS data
import time # for finding computation time
Data = h5.File(pathToData)
print(list(Data.keys()))
# The print statement shows the different files that are combined in your HDF5 file.
#
# The system seed links information contained within (e.g.) the BSE_Supernovae file to information in the BSE_System_Parameters file.
# ## 2. Finding the initial total masses of the DCOs
#
def calculateTotalMassesNaive(pathData=None):
Data = h5.File(pathToData)
totalMasses = []
#for syntax see section 1
seedsDCOs = Data['BSE_Double_Compact_Objects']['SEED'][()]
#get info from ZAMS
seedsSystems = Data['BSE_System_Parameters']['SEED'][()]
M1ZAMSs = Data['BSE_System_Parameters']['Mass@ZAMS(1)'][()]
M2ZAMSs = Data['BSE_System_Parameters']['Mass@ZAMS(2)'][()]
for seedDCO in seedsDCOs:
for nrseed in range(len(seedsSystems)):
seedSystem = seedsSystems[nrseed]
if seedSystem == seedDCO:
M1 = M1ZAMSs[nrseed]
M2 = M2ZAMSs[nrseed]
Mtot = M1 + M2
totalMasses.append(Mtot)
Data.close()
return totalMasses
# +
# calculate function run time
start = time.time()
MtotOld = calculateTotalMassesNaive(pathData=pathToData)
end = time.time()
timeDiffNaive = end-start
print('%s seconds, using for loops.' %(timeDiffNaive))
# -
# ## 3. Optimizing the above loop using built-in NumPy routines
# `NumPy` is a comprehensive library of mathematical and array computing tools, underpinned by highly optimised C-code that provides fast vectorization, indexing, and broadcasting functions. Visit [numpy.org](https://numpy.org/) for detailed information regarding ``NumPy``.
#
# The `NumPy` library provides tools that allow the user to bypass computationally heavy loops. For example, we can speed up the calculation of the element-wise sum of two arrays with:
# +
M1ZAMS = Data['BSE_System_Parameters']['Mass@ZAMS(1)'][()]
M2ZAMS = Data['BSE_System_Parameters']['Mass@ZAMS(2)'][()]
mTotalAllSystems = np.add(M1ZAMS, M2ZAMS)
# -
# ## 4. Using boolean masks in a single file
# There is a useful trick for when you want only those elements which satisfy a specific condition. Where previously we put the condition in an if statement nested within a for loop, now we will use an array of booleans to mask out the undesired elements.
#
# The boolean array will have the same length as the input array, with:
# +
# Create a boolean array from the total mass array which is True
# if the total mass of the corrresponding system is less than 40.
maskMtot = (mTotalAllSystems <= 40)
# -
# Crucially, you can apply this mask to all other columns in the same file because, by construction, they all have the same length.
# seeds of systems with total mass below 40
seeds = Data['SystemParameters']['SEED'][()]
seedsMtotBelow40 = seeds[maskMtot]
# Note that this works because the order of the two columns (seeds and total masses) are the same. For example, the total mass of the third system entry corresponds to the seed at the third system entry.
# ## 5. Using seeds as masks between files
# ### 5.1 Example 1
#
# Before we continue it is useful to understand how the COMPAS printing works.
#
# Each simulated system will be initialized only once and so will have only one line in the `BSE_System_Parameters` file. However, lines in the `BSE_Common_Envelopes` file are created whenever a system goes through a common envelope (CE) event, which might happen multiple times for a single system, or potentially not at all. Similarly, in the `BSE_Supernovae` file, you will find at most two lines per system, but possibly none. `BSE_Double_Compact_Object` file lines are printed only when both remnants are either neutron stars or black holes (but disrupted systems are also included), which happens at most once per system.
#
# For this reason, it is in general not the case that the system on line $n$ of one file corresponds to the system on line $n$ of another file.
#
# In order to match systems across files, we need to extract the seeds of desired systems from one file, and apply them as a mask in the other file.
# +
# example mock data from two files
SystemSeeds = np.array([1, 2, 3, 4 ])
SystemMass1 = np.array([1, 20, 5, 45 ])
DCOSeeds = np.array([ 2, 4 ])
# Calculate mask for which elements of SystemSeeds are found in DCOSeeds - see numpy.in1d documentation for details
mask = np.in1d(SystemSeeds, DCOSeeds)
print(mask)
print(SystemSeeds[mask])
print(SystemMass1[mask])
# -
# ### 5.2 Optimized loop
def calculateTotalMassesOptimized(pathData=None):
Data = h5.File(pathToData)
totalMasses = []
#for syntax see section 1 with basic syntax
seedsDCOs = Data['DoubleCompactObjects']['SEED'][()]
#get info from ZAMS
seedsSystems = Data['BSE_System_Parameters']['SEED'][()]
M1ZAMSs = Data['BSE_System_Parameters']['Mass@ZAMS(1)'][()]
M2ZAMSs = Data['BSE_System_Parameters']['Mass@ZAMS(2)'][()]
MZAMStotal = np.add(M1ZAMS, M2ZAMS)
maskSeedsBecameDCO = np.in1d(seedsSystems, seedsDCOs)
totalMassZAMSDCO = MZAMStotal[maskSeedsBecameDCO]
Data.close()
return totalMassZAMSDCO
# +
# calculate function run time
start = time.time()
MtotNew = calculateTotalMassesNaive(pathData=pathToData)
end = time.time()
timeDiffOptimized = end-start
# calculate number of Double Compact Objects
nrDCOs = len(Data['DoubleCompactObjects']['SEED'][()])
print('Compare')
print('%s seconds, using Optimizations.' %(timeDiffOptimized))
print('%s seconds, using For Loops.' %(timeDiffNaive))
print('Using %s DCO systems' %(nrDCOs))
# -
# *Note:* The time difference will depend on the number of systems under investigation, as well as the number of bypassed loops.
# test that the two arrays are in fact identical
print(np.array_equal(MtotOld, MtotNew))
# Note that the above loop can easily be expanded with more conditions.
#
# If you do not want all the DCO initial total masses but only of the double neutron stars, then you just need to apply another mask to the `seedsDCOs`.
def calculateTotalMassesDNS(pathToData=None):
Data = h5.File(pathToData)
totalMasses = []
#for syntax see section 1 with basic syntax
seedsDCOs = Data['BSE_Double_Compact_Objects']['SEED'][()]
type1 = Data['BSE_Double_Compact_Objects']['Stellar_Type(1)'][()]
type2 = Data['BSE_Double_Compact_Objects']['Stellar_Type(2)'][()]
maskDNS = (type1 == 13) & (type2 == 13)
seedsDNS = seedsDCOs[maskDNS]
#get info from ZAMS
seedsSystems = Data['BSE_System_Parameters']['SEED'][()]
M1ZAMSs = Data['BSE_System_Parameters']['Mass@ZAMS(1)'][()]
M2ZAMSs = Data['BSE_System_Parameters']['Mass@ZAMS(2)'][()]
MZAMStotal = np.add(M1ZAMS, M2ZAMS)
maskSeedsBecameDNS = np.in1d(seedsSystems, seedsDNS)
totalMassZAMSDNS = MZAMStotal[maskSeedsBecameDNS]
Data.close()
return totalMassZAMSDNS
# +
# calculate function run time
start = time.time()
MtotDNS = calculateTotalMassesDNS(pathToData=pathToData)
end = time.time()
timeDiffDNS = end-start
# calculate number of DNS systems
nrDNSs = len(MtotDNS)
print('%s seconds for all %s DNS systems.' %(timeDiffDNS, nrDNSs))
# -
# ### 5.3 Example 2
#
# The previous example uses the fact that both ``BSE_System_Parameters`` and ``BSE_Double_Compact_Objects`` only contain at most one line per system. However, as mentioned above, events such as supernovae or common envelopes might happen multiple times to a given system, and as a result there would be multiple occurences of a given seed in the relevant file.
#
# To account for this, we will need to modify the previous method. Consider again the 4 seeds of the previous example. Both 2 and 4 formed a DCO, and hence both stars in these binaries went SN. Seeds 1 and 3 are low mass stars hence they did not go SN. (Note that we do not specify the companion masses for any of these systems, but for simplicity we assume that the companions to 1 and 3 are also sufficiently low mass to not produce a supernova). The ``BSE_Supernovae`` file contains one line per SN and therefore seeds 2 and 4 appear twice each.
#
# Imagine you want the primary masses of systems that experienced at any point a core collapse supernova (CCSN). We'll reuse our mock data, with additional information about the types of SN which occured in each star. Here, PPISN refers to Pulsational Pair Instability Supernovae.
# +
# example mock data from above
SystemSeeds = np.array([1, 2, 3, 4 ])
SystemMass1 = np.array([1, 20, 5, 45 ])
DCOSeeds = np.array([ 2, 4 ])
SNSeeds = np.array([ 2, 2, 4, 4 ])
SNTypes = np.array(['CCSN', 'CCSN', 'CCSN', 'PPISN' ])
# get seeds which had a CCSN
maskCCSN = SNTypes == 'CCSN'
seedsCCSN = SNSeeds[maskCCSN]
print('CCSN seeds =%s' %(seedsCCSN))
#compare which element of 1-d array are in other
#this because in
seedsCCSN = np.unique(seedsCCSN)
# in this particular case, it is not necessary to reduce seedsCCSN to it's unique entries.
# the numpy.in1d function will work with duplicate seeds, but we include it explicitly here
# as other more complicated scenarios might rely on unique sets of seeds
mask = np.in1d(SystemSeeds, seedsCCSN)
print(SystemMass1[mask])
# -
# Always remember to close your data file
Data.close()
| docs/online-docs/notebooks/pythonBasics/1_slicingData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pandas for Data Exploration
# - All the material in this notebook is from Dr.Angela of [100 Days of Code](https://www.udemy.com/course/100-days-of-code)
# ## Upload the Data and Read the .csv File
#
# Then import pandas into your notebook and read the .csv file.
# +
## Upload the Data and Read the .csv File
import pandas as pd
df = pd.read_csv('salaries_by_college_major.csv')
# -
# Now take a look at the Pandas dataframe we've just created with .head(). This will show us the first 5 rows of our dataframe.
# Get the head of the data set
df.head()
# Get the shape of the data set
df.shape
# Show the title of the columns
df.columns
# ## Missing Values and Junk Data
# Before we can proceed with our analysis we should try and figure out if there are any missing or junk data in our dataframe. That way we can avoid problems later on. In this case, we're going to look for NaN (Not A Number) values in our dataframe. NAN values are blank cells or cells that contain strings instead of numbers. Use the .isna() method and see if you can spot if there's a problem somewhere.
df.isna()
df.tail()
clean_df = df.dropna()
clean_df.tail()
# ## Accessing Columns and Individual Cells in a Dataframe
# ### 1.Find college major with highest starting salaries:
#
# To access a particular column from data frame we can use the square bracket notation, like so :
clean_df['Starting Median Salary']
clean_df['Starting Median Salary'].max()
clean_df['Starting Median Salary'].idxmax()
clean_df['Starting Median Salary'].loc[43]
# * Note that the .loc[id] is return the value of the column with index. The structure is the 2 dimension array.
# * Translate go to 'Undergraduate Major' column and get the value with index 43
clean_df['Undergraduate Major'].loc[43]
clean_df['Undergraduate Major'][43]
# ### 2.The Highest Mid-Career Salary
print(clean_df['Mid-Career Median Salary'].max())
print(f"Index for the mad mid career salary: {clean_df['Mid-Career Median Salary'].idxmax()}")
clean_df['Undergraduate Major'][8]
# ### 3.The Lowest Starting and Mid-Career Salary
# Lowest Starting Salary of College Degere is:
print(f"Lowest Starting Salary of a college degree is"\
f" {clean_df['Undergraduate Major'].loc[clean_df['Starting Median Salary'].idxmin()]}"\
f" with a salaries of {clean_df['Starting Median Salary'].min()} per year")
# Lowest Starting Salary of Mid-Career Salary is
print(f"The Lowest mid-career is"\
f" {clean_df['Undergraduate Major'].loc[clean_df['Mid-Career Median Salary'].idxmin()]}"\
f" with a salaries of {clean_df['Mid-Career Median Salary'].min()} per year")
# ## Sorting Values & Adding Columns: Majors with the Most Potential vs Lowest Risk
# ### Lowest Risk Majors
# A low-risk major is a degree where there is a small difference between the lowest and highest salaries. In other words, if the difference between the 10th percentile and the 90th percentile earnings of your major is small, then you can be more certain about your salary after you graduate.
#
# How would we calculate the difference between the earnings of the 10th and 90th percentile? Well, Pandas allows us to do simple arithmetic with entire columns, so all we need to do is take the difference between the two columns:
clean_df['Mid-Career 90th Percentile Salary'] - clean_df['Mid-Career 10th Percentile Salary']
# We can also use ```.subtract()``` methods
spread_col = clean_df['Mid-Career 90th Percentile Salary'].subtract(clean_df['Mid-Career 10th Percentile Salary'])
# Insert a column into a data frame
clean_df.insert(1, 'Spread', spread_col)
clean_df.head()
# ### Sorting by the Lowest Spread
# To see which degrees have the smallest spread , we can use the ```.sort_values() ``` method. And since we are interested in only seeing the name of the degree and the major, we can pass a list of these column names to look at the ``` .head() ``` of these two columns exclusively.
low_risk = clean_df.sort_values('Spread')
low_risk[['Undergraduate Major', 'Spread']].head()
# Does ```.sort_values()``` sort in ascending or descending order? To find out, check out the Pandas documentation: https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.sort_values.html
# ### Degrees with the highest potential
# the top 5 degrees with the highest values in the 90th percentile
highest_potential = clean_df.sort_values('Mid-Career 90th Percentile Salary', ascending=False)
highest_potential[['Undergraduate Major', 'Mid-Career 90th Percentile Salary']].head()
highest_spread = clean_df.sort_values('Spread', ascending=False)
highest_spread[['Undergraduate Major', 'Spread']].head()
# ### Grouping and Pivoting Data with Pandas
# Often times you will want to sum rows that belong to a particular category. For example, which category of degrees has the highest average salary? Is it STEM, Business or HASS (Humanities, Arts, and Social Science)?
#
# To answer this question we need to learn to use the .groupby() method. This allows us to manipulate data similar to a Microsoft Excel Pivot Table.
#
# We have three categories in the 'Group' column: STEM, HASS and Business. Let's count how many majors we have in each category:
# ## Reference:
# All the material in this notebook is from Dr.Angela of [100 Days of Code](https://www.udemy.com/course/100-days-of-code). Retrieved from https://www.udemy.com/course/100-days-of-code
| Python_Pandas/.ipynb_checkpoints/pandas-introduction-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
# Import relevant packages
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
from collections import Counter
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import matplotlib.pyplot as plt
import methods_py as m
import re
import sys,time
# reload to make sure we have the latest version of method.py loaded
import importlib
importlib.reload(m)
# NLTK package for stopwords
import nltk
nltk.download(['punkt', 'wordnet','stopwords'])
from nltk.corpus import stopwords
stop_words = set(stopwords.words('english'))
from nltk.tokenize import word_tokenize
from nltk.stem import WordNetLemmatizer
# sklearn
from sklearn.pipeline import Pipeline
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
from sklearn.tree import DecisionTreeClassifier
from sklearn.svm import LinearSVC
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.metrics import classification_report, plot_precision_recall_curve, roc_auc_score
from sklearn.multiclass import OneVsRestClassifier
from sklearn.naive_bayes import MultinomialNB
# -
# References Credits: https://towardsdatascience.com/multi-label-text-classification-with-scikit-learn-30714b7819c5
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# Read in the train data
df = pd.read_csv('/kaggle/input/jigsaw-toxic-comment-classification-challenge/train.csv.zip')
# display the first 5 rows
df.head()
# -
# ## 1. Explore the dataframe
# call custom method to explore the dataframe
m.explore_df(df)
# list of columns for the labelled comment type
df.drop(['id','comment_text'],axis=1).columns
# +
# adding some of the ad-hoc words coming up as frequent and I want to exclude those
stop_words.update(["u","go","as","like","wikipedia","jim","hi","get"])
# tokeinze method to remove punctuations, lemmatize and remove stop words
def tokenize(text):
'''
INPUT
text- the text that needs to be tokenized
OUTPUT
tokens - a list of tokenized words after cleaning up the input text
This function :
1. converts text to all lower case and removes punctuations
2. tokenize entire text into words
2. lemmatize each word using WordNetLemmatizer
3. remove all stop words from text as per english corpus of stop words
'''
# normalize case and remove punctuation
text = re.sub(r"[^a-zA-Z0-9]", " ", text.lower())
# tokenize text
tokens = word_tokenize(text)
# initiate lemmatizer
lemmatizer = WordNetLemmatizer()
# lemmatize and remove stop words
tokens = [lemmatizer.lemmatize(word) for word in tokens if not word in stop_words]
return tokens
# +
print(" ###### WARNING : Comments below contain strong profanity and abusive words ####### \n")
# utilizing counter to get the most common 10 words coming up in each category
for category in list(df.drop(['id','comment_text'],axis=1).columns):
print("10 most frequent words in "+category+" comments")
print(dict(Counter(tokenize(" ".join(df[df[category]==1].comment_text))).most_common(10)).keys())
print("\n ********************* \n")
# -
# access custom method to explore dataframe
m.explore_null(df)
# +
# Lets explore the number of comments for each category
# This gives us a quick view of how the dataset is populated
df_toxic = df.drop(['id', 'comment_text'], axis=1)
counts = []
categories = list(df_toxic.columns.values)
for i in categories:
counts.append((i, df_toxic[i].sum()))
df_stats = pd.DataFrame(counts, columns=['category', 'number_of_comments'])
df_stats.plot(x='category', y='number_of_comments', kind='barh', legend=False, grid=True, figsize=(8, 5))
plt.title("Number of comments per category")
plt.ylabel('# of Occurrences', fontsize=12)
plt.xlabel('category', fontsize=12)
# +
# How many comments have multi labels?
df.drop(['id','comment_text'],axis=1).sum(axis=1).value_counts().plot(kind='bar')
plt.ylabel('# of Occurrences', fontsize=12)
plt.xlabel('# of categories', fontsize=12)
# +
# % of comments which are labeled
round((sum(df.drop(['id','comment_text'],axis=1).sum(axis=1) > 0)/len(df))*100,2)
# -
# Observation : This is an imbalanced set with almost 90% of comments not classified into any one of the 6 categories
# +
#The distribution of the number of words in comment texts.
lens = df.comment_text.str.len()
lens.hist(bins = np.arange(0,5000,50))
plt.ylabel('# of occurences', fontsize=12)
plt.xlabel('# of words', fontsize=12)
# -
# ## 2. Modeling
#
# #### We evaluate four different classification models : LinearSVC, NaiveBayes, Decision Tree Classifer and Logistic Regression
# +
# Credits: https://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html
# https://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.TfidfTransformer.html
# https://scikit-learn.org/stable/modules/multiclass.html
def build_model(model_type, c):
'''
INPUT
model type - which model we want to build between Linear SVC, NaiveBayes and Logistic Regression
OUTPUT
pipeline- a pipeline built to vectorize, tokenize, transform and classify text data
This function :
1. builds a pipeline of countvectorizer, tfidf transformer, and
2. a random forest multi output classifier
'''
#check the model_type provided by user
if model_type == 'linear_svc':
pipeline = Pipeline([
('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer()),
('clf', OneVsRestClassifier(LinearSVC(C=c), n_jobs=3))
])
#check the model_type provided by user
elif model_type== 'naivebayes':
pipeline = Pipeline([
('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer()),
('clf', OneVsRestClassifier(MultinomialNB(fit_prior=True, class_prior=None)))
])
#check the model_type provided by user
elif model_type== 'logistic_reg':
pipeline = Pipeline([
('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer()),
('clf', OneVsRestClassifier(LogisticRegression(solver='sag'), n_jobs=1))
])
#check the model_type provided by user
elif model_type== 'random_forest':
pipeline = Pipeline([
('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer()),
('clf', OneVsRestClassifier(RandomForestClassifier(max_depth=2, random_state=0), n_jobs=1))
])
#check the model_type provided by user
elif model_type== 'decision_tree':
pipeline = Pipeline([
('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer()),
('clf', OneVsRestClassifier(DecisionTreeClassifier(random_state=0), n_jobs=1))
])
# return the built pipeline
return pipeline
# -
# ## 3. Predict and evaluate models
# Credits: https://machinelearningmastery.com/roc-curves-and-precision-recall-curves-for-imbalanced-classification/
#
# We consider using the ROC AUC score for evaluation the model as it is an imbalanced dataset with around 90% of comments with no labels i.e. clean comments
#
# ***Although widely used, the ROC AUC is not without problems.***
#
# For imbalanced classification with a severe skew and few examples of the minority class, the ROC AUC can be misleading. This is because a small number of correct or incorrect predictions can result in a large change in the ROC Curve or ROC AUC score.
#
# A common alternative is the precision-recall curve and area under curve (Precision-Recall Curves and AUC)
#
def evaluate_model(model, X_test, Y_test):
'''
INPUT
model : the model that needs to be evaluated
X_test : validation data set i.e. messages in this case
Y_test : the output data for X_test validation set i.e. 36 categories values
category_names : the 36 category names
OUTPUT
classification report for the model based on predictions, gives
the recall, precision and f1 score
This function :
1. utilizes the input model to make predictions
2. compares the predictions to the test data to provide a classification report
'''
y_pred = model.predict(X_test)
# ROC AUC Score
return round(roc_auc_score(Y_test, y_pred),3)
# +
# we will be using each one of the category below to utilize them as the Predicted value (Y)
categories = ['toxic', 'severe_toxic', 'obscene', 'threat', 'insult', 'identity_hate']
# we will utilize the comment text column to predict Y
X= df['comment_text']
# -
# We first try working with a Naive Bayes Model to baseline our score
# +
model_type = 'naivebayes'
print("Training with a "+model_type+" model")
model_dict_nb= dict()
for category in categories:
print(" ---------------------------------------- ")
#print(" Working on evaluating ROC AUC score for "+category+" comments ............")
# the category becomes our predicted variable
Y= df[category]
# split the data set to a train and test set
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.2)
print('Building model for ',category,' comments')
model = build_model(model_type,1)
print('Training model...')
start_time = time.time()
model.fit(X_train, Y_train)
elapsed_time = round((time.time() - start_time)/60 , 2)
print('Training time...{} minutes'.format(elapsed_time))
print('Evaluating model...')
print("ROC AUC Score: ",evaluate_model(model, X_test, Y_test))
#save the model into a dictionary
model_dict_nb[model] = category
print("Model saved for ",category," comments")
# -
# Logistic model
# +
model_type = 'logistic_reg'
print("Training with a "+model_type+" model")
model_dict_lr= dict()
for category in categories:
print(" ---------------------------------------- ")
#print(" Working on evaluating ROC AUC score for "+category+" comments ............")
# the category becomes our predicted variable
Y= df[category]
# split the data set to a train and test set
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.2)
print('Building model for ',category,' comments')
model = build_model(model_type,1)
print('Training model...')
start_time = time.time()
model.fit(X_train, Y_train)
elapsed_time = round((time.time() - start_time)/60 , 2)
print('Training time...{} minutes'.format(elapsed_time))
print('Evaluating model...')
print("ROC AUC Score: ",evaluate_model(model, X_test, Y_test))
#save the model into a dictionary
model_dict_lr[model] = category
print("Model saved for ",category," comments")
# -
# Lets see if a decision tree classifier helps us get a better score
# +
model_type = 'decision_tree'
print("Training with a "+model_type+" model")
model_dict_dt= dict()
for category in categories:
print(" ---------------------------------------- ")
#print(" Working on evaluating ROC AUC score for "+category+" comments ............")
# the category becomes our predicted variable
Y= df[category]
# split the data set to a train and test set
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.2)
print('Building model for ',category,' comments')
model = build_model(model_type,1)
print('Training model...')
start_time = time.time()
model.fit(X_train, Y_train)
elapsed_time = round((time.time() - start_time)/60 , 2)
print('Training time...{} minutes'.format(elapsed_time))
print('Evaluating model...')
print("ROC AUC Score: ",evaluate_model(model, X_test, Y_test))
#save the model into a dictionary
model_dict_dt[model] = category
print("Model saved for ",category," comments")
# -
# ### Model Observation
# Very interesting to see that for toxic comments,
# the Linear SVC is able to generate ROC AUC Score of 0.843 in 2.45 minutes
# vs
# Decision tree classifier generating ROC AUC Score of 0.835 in 13.6 minutes
#
# IMPORTANT NOTE: More complex algorithms wont necessarily end up with same efficiency and may be even lesser score
#
# NEXT STEP:
# 1. Why is Linear SVC performing better score and faster than Decision tree ?
# 2. Why does the Random forest classifier does worse than Decision tree?
# 3. Why logistic regression and NaiveBayes are doing worse than Linear SVC
# 4. Are these specific to text classification?
# Linear SVC Model
# Credits: https://medium.com/all-things-ai/in-depth-parameter-tuning-for-svc-758215394769
# +
model_type = 'linear_svc'
print("Training with a "+model_type+" model")
model_dict_svc= dict()
for category in categories:
print(" ---------------------------------------- ")
#print(" Working on evaluating ROC AUC score for "+category+" comments ............")
# the category becomes our predicted variable
Y= df[category]
# split the data set to a train and test set
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.2)
tmp_score = 0
for C in [0.1,1,10]:
print('Building model for ',category,' comments for C=',C,'...')
model = build_model(model_type,C)
print('Training model...')
start_time = time.time()
model.fit(X_train, Y_train)
elapsed_time = round((time.time() - start_time)/60 , 2)
print('Training time...{} minutes'.format(elapsed_time))
print('Evaluating model...')
print("ROC AUC Score: ",evaluate_model(model, X_test, Y_test))
if evaluate_model(model, X_test, Y_test) > tmp_score:
#assign the latest evaluation score to tmp_score
tmp_score = evaluate_model(model, X_test, Y_test)
#assign this model to the tmp_model
tmp_model = model
#save the model into a dictionary
model_dict_svc[tmp_model] = category
print("Model saved for ",category," comments with the highest score of ",tmp_score)
# -
# Based on our comparisons between NaiveBayes , Logistic Regression, Decision Tree classifier and Linear SVC Model
#
# we consider utilizing the Linear SVC Model for our classification
#
# Comparing classifiers : https://www.cnblogs.com/yymn/p/4518016.html
# ## Testing some unseen user provided input
#
# #### ---- User Warning: The comments below contain profanity and abusive language ------
# +
comment_list= \
["==shame on you all!!!== \n\n You want to speak about gays and not about romanians...","what the hell is wrong with u","I love being here","I am going to kill this bastard",
"fuck this entire post","you are an ignoratn bastard","are you mad","you gay idiot",
"nigger go die somehere","get your fagget ass out","you are such a weiner pussy","like the new ideas","good things are done"]
for comment in comment_list:
# new list
classification_list= list()
# switch to signify clean vs toxic comments
switch=1
# put the comment into a list form
inp = [comment]
# print the comment
print("\n User Comment : ",comment,"\n")
for model in model_dict_svc.keys():
if model.predict(inp):
classification_list.append(model_dict_svc[model])
switch=0
if switch:
print("CLASSIFICATION: The comment is clean and does not contain any toxicity")
else:
print("CLASSIFICATION: ", classification_list)
# -
#set option to view the entire comment
pd.set_option('display.max_colwidth',None)
#Read in the train data
df_test = pd.read_csv('/kaggle/input/jigsaw-toxic-comment-classification-challenge/test.csv.zip')
#first 5 rows
df_test.head()
# +
# to measure the time elapsed for producing the classification
start_time = time.time()
def classify_toxicity(text):
class_list = list()
for model in model_dict_svc.keys():
if model.predict([text]):
class_list.append(model_dict_svc[model])
return class_list
df_test['classification'] = df_test['comment_text'].apply(lambda x: classify_toxicity(x))
elapsed_time = round((time.time() - start_time)/60 , 2)
print('Time taken for Classification ...{} minutes'.format(elapsed_time))
#final classification stored in a "classification" column with all labels applicable
df_test.head(20)
# -
# ## Conclusion
# **Overall,the LinearSVC model is able to perform most efficiently in terms of training time and the ROC AUC Score.**
#
# One of the reasons LinearSVC works better than Logistic Regression because LinearSVC tries to finds the “best” margin (distance between the line and the support vectors) that separates the classes and this reduces the risk of error on the data, while logistic regression does not, instead it can have different decision boundaries with different weights that are near the optimal point.
#
# SVM works well with unstructured and semi-structured data like text and images while logistic regression works with already identified independent variables. SVM is based on geometrical properties of the data while logistic regression is based on statistical approaches
| Toxic-comment-classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Understanding TF-IDF
#
# In this section we will work with texts and derive weighted metrics based on words (or terms) frequencies within these texts. More precisely, we will look at the _TF-IDF_ metric, which stands for _Term Frequency-Inverse Document Frequency_, to produce our metrics which will allow us to measure and evaluate how important certain words are in documents that are part of our IMDb corpus. The "texts" or "documents" we will look at are 'plot' descriptions in the IMDb dataset.
# ## Loading the IMDb dataset
#
# Load the IMDb dataset and look closely at the 'Plot' column
# +
import pandas as pd
df_imdb = pd.read_csv('data/imdb.csv', sep=',')
# df_imdb['Plot']
# -
# ## Create a data structure
#
# We need a custom data structure to carry out our TF-IDF calculations. Create a python dictionary having for keys the indices of the dataframe above and for value another dictionary with 'plot' as an entry for each row in the dataframe.
plot_dict = {}
for index, plot in df_imdb['Plot'].items():
if type(plot) != str:
continue
plot_dict[index] = {'plot': plot}
# ## Tokenize and filter
#
# Now that we have the plot of each IMDb entry in our dictionary, it is time to tokenize each plot's text and clean it up. Do we need punctuations as part of our tokens? Are there "stop words" we could get rid off? Please complete the following tokenizer function utilising the spacy library (which you used in Data Mining - also, remember to uncomment the first line if you are using spacy for the first time). When this is done, augment your custom dictionary with the plot's tokens for each entry.
# +
# # !python3 -m spacy download en_core_web_sm
import spacy
nlp = spacy.load("en_core_web_sm")
def split_and_stop(text):
# tokenize the text with spacy
tokens = nlp(text.lower())
return [token.text for token in tokens if not token.is_punct and not token.is_stop]
# -
for index, v in plot_dict.items():
v['tokens'] = split_and_stop(v['plot'])
#this may take some time... so we are going to print something to keep up with progress
print('.', end='')
# +
# for k in list(plot_dict.keys())[:5]:
# print("------")
# print(plot_dict[k]['plot'])
# print(plot_dict[k]['tokens'])
# -
# ## Understanding Term Frequency (TF)
#
# $$
# tf(t, d) = \frac{n_{t}} {\sum_{k} n_{k}}
# $$
#
# _Term Frequency_ is a normalised metric that measures how frequent a certain term $t$ is in a given document $d$. In the formula above ${n_{t}}$ stands for the number of times the term $t$ occur in document $d$ while $\sum_{k} n_{k}$ is the sum of all terms in the document (its length in other words). Note that term $t$ can potentially occur many times in $d$ hence the need to normalise the metric over the sum of all terms. Below is a function definition `calculate_tf` which takes as input the `tokens` of a certain document $d$ and counts the number of occurences of each terms in the document and calculate their normalised frequency.
def calculate_tf(tokens):
unique_tokens = set(tokens)
term_count = dict.fromkeys(unique_tokens, 0)
term_frequency = dict.fromkeys(unique_tokens, 0)
N = float(len(tokens))
for term in tokens:
term_count[term] += 1
term_frequency[term] += 1 / N
return term_count, term_frequency
# Considering the function `calculate_tf` above, augment your custom dictionary with both the `term_count` and normalised `term_frequency` given the respective plot's `tokens` you previously computed.
for index, v in plot_dict.items():
t_count, t_frequency = calculate_tf(v['tokens'])
v['count'] = t_count
v['tf'] = t_frequency
# +
# for k in list(plot_dict.keys())[:5]:
# print("------")
# print(plot_dict[k])
# -
# ## Understanding Inverse Document Frequency (IDF)
#
# $$
# idf(t, D) = \log\frac{|D|}{|{d_{i} \in D : t \in d_{i}}|}
# $$
#
# _Inverse Document Frequency_ is a metric that measures of important a term $t$ is in a given corpus (or collection) $D$ of documents $d_{i}$. While _Term Frequency_ measures the frequency of a term $t$ in a single document $d$, here _IDF_ consider frequency of a term $t$ over the whole corpus $D$ as to derive a weight on the statistical significance of term $t$ overall. The idea here is that common words which occur in many documents ("man" or a stop word like "it" for example) hold little importance overall as they are redundant. What _IDF_ does is to give more weight to words that are uncommon overall yet possibly significant for certain documents. This is the reason why the metric takes the $\log$ of the fraction $\frac{|D|}{|{d_{i} \in D : t \in d_{i}}|}$ where $|D|$ is the number of documents in corpus $D$ and $|{d_{i} \in D : t \in d_{i}}|$ is the number of times a term $t$ appears in a document in the corpus.
# The first thing we need to do to calculate _IDF_ is to establish the overall vocabulary of the entire corpus. What are all the unique words (or terms) in all of our plots? How many unique words do we have? Consider the following `bag_of_words` python set and fill it with all the unique terms present in our plots.
# +
# Vocabulary -> bag of words
bag_of_words = set()
for index, v in plot_dict.items():
bag_of_words = bag_of_words.union(set(v['tokens']))
len(bag_of_words)
# -
# Now, remember we calculated a `term_count` for each term in each document when we calculated the _TF_ with `calculate_tf` above? We need to use this pre-calculated informatin here to derive $|{d_{i} \in D : t \in d_{i}}|$ which is the number of times a term $t$ appears in a document in the corpus. Make a list of each `term_count` you recorded in your custom dicitonary as to use it to computer _IDF_ below.
list_all_documents_count = [v['count'] for index, v in plot_dict.items()]
# Here is function defintion `calculate_idf` that computes the _IDF_ of all the terms in our corpus. It takes a list of `term_count` as `documents_count_list` and a overall vocabulary as `bag_of_words`. Can you make sense of the function in light of the $idf(t, D)$ formula above?
# +
import math
def calculate_idf(documents_count_list, bag_of_words):
idf = dict.fromkeys(bag_of_words, 0)
D = len(documents_count_list)
for d in documents_count_list:
for term, count in d.items():
if count > 0:
idf[term] += 1
for term, document_count in idf.items():
idf[term] = math.log(D / float(document_count))
return idf
# -
# Lets calculate the _IDF_ then using the function above. What are the highest weight? What are the lowest weight?
# +
idf = calculate_idf(list_all_documents_count, bag_of_words)
# helper to visualise the terms in the IDF , sorted according to their score
sorted_idf = {k: v for k, v in sorted(idf.items(), key=lambda item: item[1])}
# sorted_idf = {k: v for k, v in sorted(idf.items(), key=lambda item: item[1], reverse=True)}
for k in list(sorted_idf.keys())[:50]:
print(f'{k} - {sorted_idf[k]}')
# -
# ## Putting it together: TF-IDF
#
# $$
# tf-idf(t, d, D) = tf(t, d) \cdot idf(t, D)
# $$
#
# Putting _TF_ and _IDF_ together is quite simple. Since _IDF_ is a weight for each term in the corpus, simply multiply the terms' weight value to all the _TF_ we already have calculated. Here is a function `calculate_tf_idf` that does just that!
def calculate_tf_idf(tf, idf):
tf_idf = dict.fromkeys(tf.keys(), 0)
for term, frequency in tf.items():
tf_idf[term] = frequency * idf[term]
return tf_idf
# With the function above, calculate the _TF-IDF_ of all plots in your custom dictionary and record the results in the dictionary itself.
for index, v in plot_dict.items():
tf_idf = calculate_tf_idf(v['tf'], idf)
v['tf_idf'] = tf_idf
# What is the difference between _TF_ and _IDF_ for a given plot?
for k in list(plot_dict.keys())[:5]:
print("------")
print(plot_dict[k]['tf'])
print(plot_dict[k]['tf_idf'])
# ## Save the data
#
# Save your custom dictionary you have constructed above in a json file.
# +
import json
with open('data/IFIDF_IMDb_plots.json', 'w') as fp:
json.dump(plot_dict, fp, indent=2)
| Solutions/Week 03/01-understanding-TF-IDF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:carnd-term1]
# language: python
# name: conda-env-carnd-term1-py
# ---
# **Vehicle Detection Project**
#
# The goals / steps of this project are the following:
#
# * Perform a Histogram of Oriented Gradients (HOG) feature extraction on a labeled training set of images and train a classifier Linear SVM classifier
# * Optionally, you can also apply a color transform and append binned color features, as well as histograms of color, to your HOG feature vector.
# * Note: for those first two steps don't forget to normalize your features and randomize a selection for training and testing.
# * Implement a sliding-window technique and use your trained classifier to search for vehicles in images.
# * Run your pipeline on a video stream (start with the test_video.mp4 and later implement on full project_video.mp4) and create a heat map of recurring detections frame by frame to reject outliers and follow detected vehicles.
# * Estimate a bounding box for vehicles detected.
# +
import numpy as np
import cv2
import matplotlib.pyplot as plt
import matplotlib.image as img
import glob
# -
#utility function to fetch the image path+names for cars and non cars respectively.
def get_image_names():
non_vehicles1=np.array(glob.glob('TrainingData/non-vehicles/non-vehicles/Extras/ex*.png'))
non_vehicles2=np.array(glob.glob('TrainingData/non-vehicles/non-vehicles/GTI/im*.png'))
non_vehicles=np.append(non_vehicles1,non_vehicles2)
vehicles=np.array(glob.glob('TrainingData/vehicles/vehicles/*/*.png'))
return non_vehicles,vehicles
# ### Visualizing Training Data
# So in the training set we have *8968 Non Vehicle Images* and *8792 vehicle Images*
# +
data=get_image_names()
print('non_vehicle images=',len(data[0]),'and vehhile images=',len(data[1]))
# -
def load_images():
non_vehicle,vehicle=get_image_names()
cars=[]
non_cars=[]
for name in vehicle:
cars.append(cv2.imread(name))
for name in non_vehicle:
non_cars.append(cv2.imread(name))
return cars,non_cars
# ### Training Data Shape
# Each training image has 64x64x3 shape.
cars,non_cars=load_images()
print(cars[0].shape)
# #### Visualizing Images
# Below is an example of Car and Non Car Image
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(10,5))
ax1.imshow(cv2.cvtColor(cars[0],cv2.COLOR_BGR2RGB))
ax1.set_title('Car Image', fontsize=30)
ax2.imshow(cv2.cvtColor(non_cars[0],cv2.COLOR_BGR2RGB))
ax2.set_title('Non car Image', fontsize=30)
# ## HOG Features
# To detect the vehicles I used Histogram of Oriented Gradients as one of the feature. I took HOG on the 'YCrCb' color space and to be more specific I used 'Cr' color channel to extract HOG features. I tried different color spaces and different color channels while going through the classroom quizes, however while trying different combinations and found that the classifier accuracy is best if I use color channel 'Cr' for the hog features.
#
# Function below takes image and color space name as input, orientation and other parameters are optional. However during the training I used ``pix_per_cell=16``
# ``orient=9`` ``Color_space=YCrCb`` ``cells_per_block=2`` and ``Channel=1``
#
# I used this configurations because I realized that the accuracy of classifier is above 95% if I am feeding it data, taken out of hog with this configuration. The feature vector length is *576* if I use this configuration.
from skimage.feature import hog
def get_hog_features(image,cspace, orient=9, pix_per_cell=8, cell_per_block=2, vis=True,
feature_vec=True,channel=0):
if cspace != 'BGR':
if cspace == 'HSV':
feature_image = cv2.cvtColor(image, cv2.COLOR_BGR2HSV)
elif cspace == 'LUV':
feature_image = cv2.cvtColor(image, cv2.COLOR_BGR2LUV)
elif cspace == 'HLS':
feature_image = cv2.cvtColor(image, cv2.COLOR_BGR2HLS)
elif cspace == 'YUV':
feature_image = cv2.cvtColor(image, cv2.COLOR_BGR2YUV)
elif cspace == 'YCrCb':
feature_image = cv2.cvtColor(image, cv2.COLOR_BGR2YCrCb)
elif cspace == 'RGB':
feature_image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
else: feature_image = np.copy(image)
return_list = hog(feature_image[:,:,channel], orientations=orient, pixels_per_cell=(pix_per_cell, pix_per_cell),
cells_per_block=(cell_per_block, cell_per_block),
block_norm= 'L2-Hys', transform_sqrt=False,
visualise= vis, feature_vector= feature_vec)
# name returns explicitly
hog_features = return_list[0]
if vis:
hog_image = return_list[1]
return hog_features, hog_image
else:
return hog_features
# ### Output of HOG
# Below is the example of HOG output
# +
hog_features,hog_image=get_hog_features(cars[1],'YCrCb',channel=1,pix_per_cell=16)
print('shape of hog features ',hog_features.shape)
plt.imshow(hog_image,cmap='gray')
# -
# ## Spatial Binning
# I used Spatial Binning to extract more features from the image. So in Spatial Binning we take the raw pixel values from the image. The basic concept here is; in images, even if we decrease the size of the image within certain range, it still retains most of its information.
#
# So here input image was 64x64 image which I resized as 16x16 image and then I used it as feature vector for the classifier along with HOG feature vector.
# I used ``ravel()`` function to convert the 2D array to vector.
#
# I used 'YUV' color space for spatial binning, the below function takes the image input and convert it to the given Color space. After few observations it was clear 'YUV' gives good result in our case, this can be seen in the sample outputs below:
def bin_spatial(image, cspace='BGR', size=(16, 16)):
# Convert image to new color space (if specified)
if cspace != 'BGR':
if cspace == 'HSV':
feature_image = cv2.cvtColor(image, cv2.COLOR_BGR2HSV)
elif cspace == 'LUV':
feature_image = cv2.cvtColor(image, cv2.COLOR_BGR2LUV)
elif cspace == 'HLS':
feature_image = cv2.cvtColor(image, cv2.COLOR_BGR2HLS)
elif cspace == 'YUV':
feature_image = cv2.cvtColor(image, cv2.COLOR_BGR2YUV)
elif cspace == 'YCrCb':
feature_image = cv2.cvtColor(image, cv2.COLOR_BGR2YCrCb)
elif cspace == 'RGB':
feature_image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
else: feature_image = np.copy(image)
# Use cv2.resize().ravel() to create the feature vector
small_img=cv2.resize(feature_image,size) # Remove this line!
# Return the feature vector
features=small_img.ravel()
return features
# #### Spatial Binning output for Car Images
plt.plot(bin_spatial(cars[0],'YUV'))
# #### Spatial Binning output for Non Car Images
plt.plot(bin_spatial(non_cars[0],'YUV'))
# ## Color Histogram
# I also used Color Histograms to fetch features out of an image. As the name implies we take an image and based on the given color channel and bin size specifications we calculate the histogram for each given channel and bin size and then append them together to form a feature vector.
#
# I used HLS color space and 'S' color channel for the color histogram feature vector. After doing some experimentaion I found that Saturation can be a reliable feature to identify the Vehicles.
#
# I used ``Number of bins=32`` ``color space=HLS`` and ``bins range=0-256``
#
# Below is the sample output of color histogram for a given image and given color space(HLS in our case).
def color_hist(image, nbins=32, channel=None,bins_range=(0, 256),cspace='BGR',v=False):
# Compute the histogram of the RGB channels separately
if cspace != 'BGR':
if cspace == 'HSV':
feature_image = cv2.cvtColor(image, cv2.COLOR_BGR2HSV)
elif cspace == 'LUV':
feature_image = cv2.cvtColor(image, cv2.COLOR_BGR2LUV)
elif cspace == 'HLS':
feature_image = cv2.cvtColor(image, cv2.COLOR_BGR2HLS)
elif cspace == 'YUV':
feature_image = cv2.cvtColor(image, cv2.COLOR_BGR2YUV)
elif cspace == 'YCrCb':
feature_image = cv2.cvtColor(image, cv2.COLOR_BGR2YCrCb)
elif cspace == 'RGB':
feature_image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
else: feature_image = np.copy(image)
if(channel==None):
first_hist = np.histogram(feature_image[:,:,0],bins=nbins,range=bins_range)
second_hist = np.histogram(feature_image[:,:,1],bins=nbins,range=bins_range)
third_hist = np.histogram(feature_image[:,:,2],bins=nbins,range=bins_range)
bin_edges=first_hist[1]
bin_centers = (bin_edges[1:]+bin_edges[0:len(bin_edges)-1])/2
if(v):
return first_hist, second_hist, third_hist,bin_centers
else:
hist_features = np.concatenate((first_hist[0], second_hist[0], third_hist[0]))
# Return the individual histograms, bin_centers and feature vector
return hist_features
else:
first_hist = np.histogram(feature_image[:,:,channel],bins=nbins,range=bins_range)
bin_edges=first_hist[1]
# Generating bin centers
bin_centers = (bin_edges[1:]+bin_edges[0:len(bin_edges)-1])/2
# Concatenate the histograms into a single feature vector
# hist_features = np.concatenate((rhist[0],ghist[0],bhist[0]))
# Return the individual histograms, bin_centers and feature vector
if(v):
return first_hist,bin_centers
return first_hist[0]
# #### Output of Color Histogram function
histogram=color_hist(cars[0],cspace='HLS',v=True)
fig = plt.figure(figsize=(12,3))
plt.subplot(131)
plt.bar(histogram[3], histogram[0][0])
plt.xlim(0, 256)
plt.title('H Histogram')
plt.subplot(132)
plt.bar(histogram[3], histogram[1][0])
plt.xlim(0, 256)
plt.title('L Histogram')
plt.subplot(133)
plt.bar(histogram[3], histogram[2][0])
plt.xlim(0, 256)
plt.title('S Histogram')
fig.tight_layout()
histogram=color_hist(cars[0],cspace='YUV',channel=1,v=True)
fig = plt.figure(figsize=(24,6))
plt.subplot(131)
plt.bar(histogram[1], histogram[0][0])
plt.xlim(0, 256)
plt.title('S Histogram')
# ## Classifier
# I used Support Vector Machine as my classifier, I choose this because it has simple implementaion and training time for this classifier is also considerably small while compared with Neural Networks and other classifiers.
#
# Initially I was using 'linear' kernel, but even after acheiving 96% test accuracy with the linear kernel there were too many false postive detections. Then I thought of increasing the size of feature vector or to use Radial Basis function('rbf') as kernel. However I used 'rbf' kernel since it gave 99% test accuracy and the number of false positive detection also decreased drastically.
# +
from sklearn.svm import LinearSVC
from sklearn.svm import SVC
from sklearn.preprocessing import StandardScaler
def train_model(X_train,y_train):
svc=SVC(kernel='rbf')
svc.fit(X_train,y_train)
return svc
# -
# ## Extract Features
# Function ``extract_fetures()`` is used to fetch feature vector from each image during the training phase of the classifier. This function simply extracts the feature vector for each image and it dumps these features into a pickle file, later we use these features to traing our classifier.
import pickle
import time
from sklearn.preprocessing import StandardScaler
from sklearn.cross_validation import train_test_split
def extract_features():
cars,non_cars=load_images()
cars_features=[]
non_cars_features=[]
for car in cars:
color_hist_features1=color_hist(car,cspace='HLS',channel=2)
#color_hist_features2=color_hist(car,cspace='YUV',channel=1)
hog_features=get_hog_features(car,'YCrCb',channel=1,pix_per_cell=16)[0]
spatial_features=bin_spatial(car,'YUV')
temp=np.array([])
temp=np.append(temp,color_hist_features1)
#temp=np.append(temp,color_hist_features2)
temp=np.append(temp,hog_features)
temp=np.append(temp,spatial_features)
cars_features.append(temp)
for non_car in non_cars:
color_hist_features1=color_hist(non_car,cspace='HLS',channel=2)
#color_hist_features2=color_hist(non_car,cspace='YUV',channel=1)
hog_features=get_hog_features(non_car,'YCrCb',channel=1,pix_per_cell=16)[0]
spatial_features=bin_spatial(non_car,'YUV')
temp=np.array([])
temp=np.append(temp,color_hist_features1)
#temp=np.append(temp,color_hist_features2)
temp=np.append(temp,hog_features)
temp=np.append(temp,spatial_features)
non_cars_features.append(temp)
file=open('data.pkl','wb')
obj1=['cars',cars_features]
obj2=['non_cars',non_cars_features]
pickle.dump(obj1, file)
pickle.dump(obj2, file)
file.close()
# ## Train Model and save
# Function ``train_and_save()`` uses the features received from function ``extract_features`` to train the classifier and later it save it to a pickle file.
#
# I have used ``StandardScaler()`` to scale all the features in the feature vector for all the images, it is important since if there is so much variation among the values of the features then there are chances that the classifier gets biased towards the higher value features. I had to save the scaler once I fetch it, since same scaler shall be used to make predictions which was used to scale the input during the training.
#
# Length of the feature vector is 1124
# +
def train_and_save(flag_extract_features=False):
if(flag_extract_features):
extract_features()
pickle_in = open("data.pkl","rb")
example_dict = pickle.load(pickle_in)
cars_features=example_dict[1]
example_dict = pickle.load(pickle_in)
non_cars_features=example_dict[1]
pickle_in.close()
print('Length of feature vector=',cars_features[0].shape[0])
X = np.vstack((cars_features, non_cars_features)).astype(np.float64)
# Define the labels vector
y = np.hstack((np.ones(len(cars_features)), np.zeros(len(non_cars_features))))
# Split up data into randomized training and test sets
rand_state = np.random.randint(0, 100)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=rand_state)
X_scaler = StandardScaler().fit(X_train)
X_train = X_scaler.transform(X_train)
X_test = X_scaler.transform(X_test)
t=time.time()
clf=train_model(X_train,y_train)
t2 = time.time()
print(round(t2-t, 2), 'Seconds to train SVC...')
# Check the score of the SVC
print('Test Accuracy of SVC = ', round(clf.score(X_test, y_test), 4))
file=open('classifier.pkl','wb')
obj1=['model',clf]
obj2=['scaler',X_scaler]
pickle.dump(obj1,file)
pickle.dump(obj2,file)
file.close()
return clf,X_scaler
# -
train_and_save()
# ## Sliding Window
# Once I was done training the classifier, next challenge was how to find the vehicles in a given image. Well, I used sliding window approach to find vehices in an image. In this we use different sized windows and move them accross the image, fetch feature vector for that window and feed those features to our trained classifier, if classifier predicts that Yes! it is a vehicle then mark that window.
#
# It was challenging to fit the good window size for the sliding window, after experimenting different combinations for the sliding window size I finally used two window sizes:
# 1. 50x50 window for y=400 to y=500 since near the horizon the cars will be far and small in size, in this case overlap is 50% for both x and y.
# 2. 80x100 window for y=500 to y=650 since in this region cars will appear larger in size, in this case overlap is 70% for both x and y.
#
# I have used different sized windows because vehicles in differnt regions of the image appears different, i.e. vehicles near the car appears bigger and far from the car appears smaller. I tried different overlaping factors, use of small overlaping factor worked well, if the window size is small too, for large windows overlaping factor should also be large. I realized the overlaping factor also depends on what threshold you use during the heatmap implementaion.
def slide_window(img,window_list, x_start_stop=[None, None], y_start_stop=[None, None],
xy_window=(100, 70), xy_overlap=(0.8, 0.8)):
# If x and/or y start/stop positions not defined, set to image size
if x_start_stop[0] == None:
x_start_stop[0] = 0
if x_start_stop[1] == None:
x_start_stop[1] = img.shape[1]
if y_start_stop[0] == None:
y_start_stop[0] = 0
if y_start_stop[1] == None:
y_start_stop[1] = img.shape[0]
# Compute the span of the region to be searched
xspan = x_start_stop[1] - x_start_stop[0]
yspan = y_start_stop[1] - y_start_stop[0]
# Compute the number of pixels per step in x/y
nx_pix_per_step = np.int(xy_window[0]*(1 - xy_overlap[0]))
ny_pix_per_step = np.int(xy_window[1]*(1 - xy_overlap[1]))
# Compute the number of windows in x/y
nx_buffer = np.int(xy_window[0]*(xy_overlap[0]))
ny_buffer = np.int(xy_window[1]*(xy_overlap[1]))
nx_windows = np.int((xspan-nx_buffer)/nx_pix_per_step)
ny_windows = np.int((yspan-ny_buffer)/ny_pix_per_step)
# Initialize a list to append window positions to
# Loop through finding x and y window positions
# Note: you could vectorize this step, but in practice
# you'll be considering windows one by one with your
# classifier, so looping makes sense
for ys in range(ny_windows):
for xs in range(nx_windows):
# Calculate window position
startx = xs*nx_pix_per_step + x_start_stop[0]
endx = startx + xy_window[0]
starty = ys*ny_pix_per_step + y_start_stop[0]
endy = starty + xy_window[1]
# Append window position to list
window_list.append(((startx, starty), (endx, endy)))
# Return the list of windows
return window_list
def search_windows(image, windows, clf,scaler):
#1) Create an empty list to receive positive detection windows
on_windows = []
#2) Iterate over all windows in the list
for window in windows:
#3) Extract the test window from original image
test_img = cv2.resize(image[window[0][1]:window[1][1], window[0][0]:window[1][0]], (64, 64))
#4) Extract features for that window using single_img_features()
test_features=[]
color_hist_features1=color_hist(test_img,cspace='HLS',channel=2)
#color_hist_features2=color_hist(test_img,cspace='YUV',channel=1)
hog_features=get_hog_features(test_img,'YCrCb',channel=1,pix_per_cell=16)[0]
spatial_features=bin_spatial(test_img,'YUV')
temp=np.array([])
temp=np.append(temp,color_hist_features1)
#temp=np.append(temp,color_hist_features2)
temp=np.append(temp,hog_features)
temp=np.append(temp,spatial_features)
test_features.append(temp)
#print(test_features)
#5) Scale extracted features to be fed to classifier
#scaler=StandardScaler().fit(test_features)
features = scaler.transform(np.array(test_features).reshape(1, -1))
#print(features)
#6) Predict using your classifier
prediction = clf.predict(features)
#7) If positive (prediction == 1) then save the window
#print(prediction)
if prediction == 1:
on_windows.append(window)
#8) Return windows for positive detections
return on_windows
def draw_boxes(img, bboxes, color=(0, 0, 255), thick=6):
# Make a copy of the image
imcopy = np.copy(img)
# Iterate through the bounding boxes
for bbox in bboxes:
# Draw a rectangle given bbox coordinates
cv2.rectangle(imcopy, bbox[0], bbox[1], color, thick)
# Return the image copy with boxes drawn
return imcopy
# ## Heatmap
# Since window size is small, hence our classfier predicts 1(Vehicle) for most of the windows that contain some part of the vehicle in it because of this for a vehicle we have different windows marked. But at last we want to show only one bounding box for a vehicle. To overcome this problem we use heatmap.
#
# ``add_heat`` function is used to find out which part of image was considered how many times by the classifier that this part has a vehicle. i.e. if the value of a pixel is 10 it means that pixel was included 10 times in such windows for which the prediction was 1.
#
# Once we have a heatmap now we can apply thresholding to it so that we have only those regions that have higher probabilities that there is a vehicle in the region.
#
# #### Label
# Label the obtained detection areas with the ``label()`` function of the scipy.ndimage.measurements package. In this step we outline the boundaries of labels that is, we label each cluster of windows as one car, so in this step we simply get the bounding box of that cluster(vehicle).
#
# #### False Positive Filtering
# To filter false positives I ignored all the windows which has dimensions smaller than 30x30, using this I was able to filter out most of the false positives in my output.
# +
from scipy.ndimage.measurements import label
def add_heat(heatmap, bbox_list):
# Iterate through list of bboxes
for box in bbox_list:
# Add += 1 for all pixels inside each bbox
# Assuming each "box" takes the form ((x1, y1), (x2, y2))
heatmap[box[0][1]:box[1][1], box[0][0]:box[1][0]] += 1
# Return updated heatmap
return heatmap# Iterate through list of bboxes
def apply_threshold(heatmap, threshold):
# Zero out pixels below the threshold
heatmap[heatmap <= threshold] = 0
# Return thresholded map
return heatmap
def draw_labeled_bboxes(img, labels):
# Iterate through all detected cars
for car_number in range(1, labels[1]+1):
# Find pixels with each car_number label value
nonzero = (labels[0] == car_number).nonzero()
# Identify x and y values of those pixels
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
# Define a bounding box based on min/max x and y
bbox = ((np.min(nonzerox), np.min(nonzeroy)), (np.max(nonzerox), np.max(nonzeroy)))
# False Positve Filtering
if((np.absolute(bbox[0][0]-bbox[1][0])>30) & ( np.absolute(bbox[0][1]-bbox[1][1])>30)):
cv2.rectangle(img, bbox[0], bbox[1], (0,0,255), 6)
# Return the image
return img
# -
# #### Output after using heatmap
# +
test_imagee=cv2.imread('./test_images/test1.jpg')
windows=[]
windows=slide_window(image,windows,x_start_stop=[200, None], y_start_stop=[400, 500],xy_window=(50,50),xy_overlap=(0.5,0.5))
windows=slide_window(image,windows,x_start_stop=[200, None], y_start_stop=[400, 656],xy_window=(100,80),xy_overlap=(0.7,0.7))
#windows=slide_window(test_imagee,windows,x_start_stop=[200, None], y_start_stop=[500, 650],xy_window=(128,128),xy_overlap=(0.6,0.6))
pickle_input = open("classifier.pkl","rb")
example_dict = pickle.load(pickle_input)
clf1=example_dict[1]
example_dict = pickle.load(pickle_input)
scaler1=example_dict[1]
#clf,scaler=train_and_save()
pickle_input.close()
on_windows=search_windows(test_imagee, windows, clf1,scaler1)
heat=np.zeros_like(test_imagee[:,:,0]).astype(np.float)
heatmap=add_heat(heat,on_windows)
th=apply_threshold(heatmap,0.7)
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(15,8))
ax1.imshow(heatmap,cmap='hot')
ax1.set_title('HeatMap', fontsize=20)
ax2.imshow(th,cmap='hot')
ax2.set_title('heatmap with threshold', fontsize=20)
# -
# ## Pipeline
#
# I have used a class named ``vehicle_detection`` to keep data from the previous frames. A vehicle will not move more than few pixels in any direction hence we can use data collected from previous frames so that there is small variation in the window size in consecutive frames.
#
# The pipeline performs few steps during execution:
# 1. It takes an image as an input and converts it from RGB to BGR color space.
# 2. It calls `slide_window()` function to get different windows.
# 3. It loads the trained classifier and scaler from the pickle file.
# 4. It calls `search_window()` and provides image and windows from step 2 to the function, this function fetches features for that window and feeds them to classifier to get the predicted value.
# 5. It calls the heatmap fucntion to get only a bounding box for each image in an image.
# 6. It keeps the running average of the heatmap values for previous 18 frames, later I used the mean of those values.
# 9. Draw the bounding box and return the image
class vehicle_detection:
heatmap_average=np.array([])
def pipeline(self,image):
windows=[]
image=cv2.cvtColor(image,cv2.COLOR_RGB2BGR)
windows=slide_window(test_imagee,windows,x_start_stop=[200, None], y_start_stop=[400, 500],xy_window=(50,50),xy_overlap=(0.5,0.5))
windows=slide_window(test_imagee,windows,x_start_stop=[200, None], y_start_stop=[400, 656],xy_window=(100,80),xy_overlap=(0.7,0.7))
pickle_in = open("classifier.pkl","rb")
example_dict = pickle.load(pickle_in)
clf=example_dict[1]
example_dict = pickle.load(pickle_in)
scaler=example_dict[1]
#clf,scaler=train_and_save()
pickle_in.close()
on_windows=search_windows(image, windows, clf,scaler)
#output=draw_boxes(image,on_windows)
heat=np.zeros_like(image[:,:,0]).astype(np.float)
heatmap=add_heat(heat,on_windows)
self.heatmap_average=np.append(self.heatmap_average,heatmap)
if(len(self.heatmap_average)>18*len(np.array(heatmap).ravel())):
self.heatmap_average=self.heatmap_average[len(np.array(heatmap).ravel()):]
#print(len(self.heatmap_average),len(np.array(heatmap).ravel()))
heatmap=np.mean((self.heatmap_average.reshape(-1,len(np.array(heatmap).ravel()))),axis=0)
heatmap=heatmap.reshape(-1,image.shape[1])
#print(heatmap.shape)
heatmap=apply_threshold(heatmap,0.7)
labels = label(heatmap)
output = draw_labeled_bboxes(np.copy(image), labels)
return cv2.cvtColor(output,cv2.COLOR_BGR2RGB)
# #### FInal Output for one frame
test_imagee=img.imread('./test_images/test1.jpg')
detection=vehicle_detection()
plt.imshow(detection.pipeline(test_imagee))
from moviepy.editor import VideoFileClip
from IPython.display import HTML
white_output = 'project_video_Submission_final.mp4'
detection=vehicle_detection()
clip1 = VideoFileClip("project_video.mp4")
white_clip = clip1.fl_image(detection.pipeline) #NOTE: this function expects color images!!
# %time white_clip.write_videofile(white_output, audio=False)
# ## Challenges
# I faced following challenges while working on the project.
# 1. First challenge was to choose what features to use, it took a lot of experimentation to select the appropriate features.
# 2. Second was which classifier should I use, I chose SVM as classifier and 'linear' as my kernel but even after achieving 95% of test accuracy with linear kernel lots of false postive detections were made. I later used 'rbf' kernel to overcome the problem of false detection.
# 3. What size of windows should I choose was the most challenging part, It took lots of effort and testing to come up with the window size that would work fine for our problem. I chose different sized windows for vehicle detection.
# 4. After detection bounding boxes were wobbling too much frame to frame, to overcome this I tried keeping a running average of heatmap from previous frames. It worked out really well for this problem.
| Vehicle_Detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Interpreting Coefficients
#
# It is important that not only can you fit complex linear models, but that you then know which variables you can interpret.
#
# In this notebook, you will fit a few different models and use the quizzes below to match the appropriate interpretations to your coefficients when possible.
#
# In some cases, the coefficients of your linear regression models wouldn't be kept due to the lack of significance. But that is not the aim of this notebook - **this notebook is strictly to assure you are comfortable with how to interpret coefficients when they are interpretable at all**.
# +
import numpy as np
import pandas as pd
import statsmodels.api as sm;
df = pd.read_csv('./house_prices.csv')
df.head()
# -
# We will be fitting a number of different models to this dataset throughout this notebook. For each model, there is a quiz question that will allow you to match the interpretations of the model coefficients to the corresponding values. If there is no 'nice' interpretation, this is also an option!
#
# ### Model 1
#
# `1.` For the first model, fit a model to predict `price` using `neighborhood`, `style`, and the `area` of the home. Use the output to match the correct values to the corresponding interpretation in quiz 1 below. Don't forget an intercept! You will also need to build your dummy variables, and don't forget to drop one of the columns when you are fitting your linear model. It may be easiest to connect your interpretations to the values in the first quiz by creating the baselines as neighborhood C and home style **lodge**.
dummies1 = pd.get_dummies(df['neighborhood'])
dummies2 = pd.get_dummies(df['style'])
dumm_df = df.join(dummies1)
new_df = dumm_df.join(dummies2)
new_df.head()
new_df['intercept'] = 1
l_m = sm.OLS(new_df['price'], new_df[['intercept', 'A','B', 'area', 'ranch', 'victorian']])
model = l_m.fit()
model.summary()
# ### Model 2
#
# `2.` Now let's try a second model for predicting price. This time, use `area` and `area squared` to predict price. Also use the `style` of the home, but not `neighborhood` this time. You will again need to use your dummy variables, and add an intercept to the model. Use the results of your model to answer quiz questions 2 and 3.
new_df['area_squared'] = new_df['area']*new_df['area']
new_df.head()
new_df['intercept'] = 1
lm = sm.OLS(new_df['price'], new_df[['intercept', 'area', 'area_squared', 'victorian', 'ranch']])
model1 = lm.fit()
model1.summary()
| Practical_statistics/Regression/Interpreting Model Coefficients.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # 07 - Deep Learning
# + [markdown] slideshow={"slide_type": "slide"}
# ## 06 - RNN Introduction
# -
# 
# Picture by [<NAME>](https://unsplash.com/photos/lxujDxNigL4)
# + [markdown] slideshow={"slide_type": "skip"}
# ___
# + [markdown] slideshow={"slide_type": "skip"}
# Today is about another widely used kind of neural networks: the Recurrent Neural Networks. They are used in many modern applications requiring to handle sequences of information, such as language translation or speech recognition.
# -
# # I. A Sequential Problem
# ## I.1. What is sequential information?
# Let's consider a typical problem: you have an image of a ball in motion, like the following, can you predict where the ball will go next?
#
# 
# Unless you know newtonian physics better than any current scientist, you can not predict where the ball will go.
#
# But now assume that you have some previous positions of the ball as well, like in the following image, can you predict the direction?
#
# 
# Sure you can, because you have a sequential information: not just a snapshot a given time, but **several snapshots** at different times.
# ## I.2. Examples of sequential problems
# You are playing everyday with sequential information, without even noticing it!
#
# For example, audio signals (music, speech, any sound...) are sequential information.
#
# 
# An audio signal is just a long list of numbers, they represent a sequence of amplitude.
# Another good example is what you are just reading... Yeah, **a sentence is a sequence of words**!
# Thus, there are numbers of sequential problems:
# * Language translation
# * speech recognition
# * music generation
# * sentiment classification
# * video activity
# * ...
# ## I.3. Limitations of classical approaches
# Let's take a simple case. We have the following problem:
#
# 
#
# How would you solve it using machine learning and NLP?
# **Idea 1**: a window of few previous words and one hot encoding:
# 
#
#
# This is a good idea, but what if we have long term dependencies, like in a sentence like the following:
# > "I used to live in China when I was a kid, even though I'm French. That's why I speak fluent ..."
#
# Would that that on this kind of sentence? Wouldn't it predict that the guy speaks French while it is meant to predict Chinese?
# **Idea 2**: a BOW (or TF-IDF):
# 
#
# BOW and TF-IDF are really powerful features. But they **do not preserve order** of the sequence. For a BOW, the two following sentences are the same:
#
# > "The food is **good**, **not bad** at all!"
#
# > "The food is **bad**, **not good** at all!"
#
# This is a serious issue when dealing with sentiment analysis!
# + [markdown] slideshow={"slide_type": "slide"}
# # II. RNN Basics
# -
# ## II.1. Requirements to handle sequential information
# To properly handle sequential information, here is what we would need to do:
# - handle variable length sequences
# - track long-term dependencies
# - keep information about order
# - share parameters across the sequence
#
# Well, **Recurrent Neural Networks (RNN)** can do most of it, let's see how!
# ## II.2. Representation
# Up to now, you used to see diagrams where neural networks are going from left (with the features) to right (with the prediction).
#
# 
# When using recurrent neural networks, we will change this representation to the following:
# 
# Where in blue are the input features, in green here are the layers of the neural network, and in purple the prediction.
# ## II.3. Types of RNN
# Depending on the problem to solve, many types of RNN can be used. Indeed, a sentiment analysis (one ouput) or a language translation (multiple output) have different requirements.
# 
# So to summarize:
# - A one-to-one is a MLP as you already know it
# - A many-to-many could be a translation model: it inputs a sequence of words in english and outputs a sequence of words in french
# - A many-to-one could be a sentiment analysis model: it inputs a sequence of words and outputs a review
# ## II.4. Hidden state
# Before digging into the details of the computation, one more concept to add is the **hidden state**.
#
# 
#
# Actually, as you will see in the next section, a RNN has a two step computation. First, you compute a **hidden state** $h_t$ using both:
# - the input features $x_t$
# - the previous hidden state $h_{t-1}$
#
# After that only, you compute the prediction $\hat{y}$ using this hidden state $h_t$.
# # III. RNN Computation
# ## III.1. Step-by-step computation
# So, how does a RNN computes predictions?
#
# We will consider $x_1$, $x_2$, $x_3$... $x_t$ to be the words number 1, 2, 3... t of a sentence in english.
#
# The target $y_1$, $y_2$, $y_3$... $y_t$ are the words number 1, 2, 3... t of the same sentence in french.
#
# So our neural network will look like this:
# 
# In a RNN, actually the same weights are shared for all steps of the sequence:
#
# 
#
# The weights $W_{xh}$ and $W_{hh}$ will allow to compute the hidden state as a perceptron would do:
#
# $$
# \large h_t = g(W_{hh} h_{t-1} + W_{xh} X_t + b)
# $$
#
# This is just like a perceptron, or a classical neural network, where $W_{xh} X_t$ is a weighted sum of features $X_t$, and $g$ is just an activation function.
# Then the second step will be to compute the predictions $\hat{y}$.
#
# 
#
# This is where the weights $W_{hy}$ appear. Using those weights, this will again work just like a perceptron:
#
# $$
# \large \hat{y_t} = g(W_{hy} h_{t} + b)
# $$
# ## III.2. Loss computation (optional)
# You might be wondering... how do we compute the loss in such a complicated network? And thus, what do we minimize with gradient descent?
# Each time step is a regular neural network (a MLP). So, we can compute a loss for each time step, right?
#
# 
# So that at each step *t*, we end up with a loss $L_t$. Then, sum them and you will have the global loss!
# 
# # IV. Application to movie review analysis
# ## IV.1. Preprocessing the data
# The IMDB movie review dataset is a dataset containing review for movies, as well as an associated label 0 (negative review) or 1 (positive review).
#
# Let's load it.
# +
import numpy as np
from tensorflow.keras import datasets
imdb = datasets.imdb
(X_train, y_train), (X_test, y_test) = imdb.load_data(num_words=10000)
X_train.shape, y_train.shape
# -
print('possible labels:', np.unique(y_train))
# The train set is composed of 25000 samples. In each training sample, there is a list of numbers, corresponding to the output of a BOW with 10000 words:
print(X_train[0])
# So the training set actually is a sequence of words, encoded into numbers. In the first exercise, you will be able to find the correspondences between those numbers and the corresponding words.
#
#
# The dataset is already preprocessed, which is highly convenient. But still, the sequences may not have the same length:
print('length of sequence 0:', len(X_train[0]))
print('length of sequence 1:', len(X_train[1]))
# We will have to pad our training set. This is exactly what we did with images in CNN: we will add zeros so that all the sequences have the same length.
# +
from tensorflow.keras.preprocessing import sequence
X_train = sequence.pad_sequences(X_train,
value=0,
padding='post', # to add zeros at the end
maxlen=256) # the length we want
X_test = sequence.pad_sequences(X_test,
value=0,
padding='post', # to add zeros at the end
maxlen=256) # the length we want
# -
print('length of sequence 0:', len(X_train[0]))
print('length of sequence 1:', len(X_train[1]))
# To do this padding properly, we would first have to check the distribution of the length of our sequences.
# Indeed, if most sequences have a length below 80, it makes no sense to keep a length of 256. On the other hand, if most sequences are 800 words long, a 256 padding will lose most of the information.
#
# You will implement this in the exercises.
#
# Now we are ready to build our first RNN.
# ## IV.2. Building a RNN
# Now we will build a RNN to predict the labels. We can build a RNN with two layers and 8 units each. Finally we will add a sigmoid dense layer that computes the final prediction.
# +
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import SimpleRNN, Dense, Embedding
def my_RNN():
model = Sequential()
# The input_dim is the number of different words we have in our corpus: here 10000
# The input_length is the length of our sequences: here 256 thanks to padding
model.add(Embedding(input_dim=10000, output_dim=32, input_length=256))
# We add two layers of RNN
model.add(SimpleRNN(units=8, return_sequences=True))
model.add(SimpleRNN(units=8, return_sequences=False))
# Finally we add a sigmoid
model.add(Dense(units=1, activation='sigmoid'))
return model
# -
# Several things can be noticed here:
# - We have an `Embedding` layer: it converts our BOW input into Word Embedding like numbers, so here basically each word is transformed into a 32 features array
# - We have stacked two `RNN` layers: this does not mean we are doing two RNN, this just means our neural network contains two layers
# - This first `RNN` has `return_sequences=True` and the second one has `return_sequences=False`: indeed the sequence is needed when another layer of `RNN` is added. The thumb rule for *many-to-one* is: `return_sequences=True` when there is another `RNN` layer, `return_sequences=False` otherwise
# For our example, we will take only one layer:
def my_RNN():
model = Sequential()
# The input_dim is the number of different words we have in our corpus: here 10000
# The input_length is the length of our sequences: here 256 thanks to padding
model.add(Embedding(input_dim=10000, output_dim=32, input_length=256))
# We add one layers of RNN
model.add(SimpleRNN(units=32, return_sequences=False))
# Finally we add a sigmoid
model.add(Dense(units=1, activation='sigmoid'))
return model
# As you already know, we can then compile the model:
# +
model = my_RNN()
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['accuracy'])
# -
# Finally, let's train our RNN:
model.fit(x=X_train, y=y_train, validation_data=(X_test, y_test), epochs=10, batch_size=128)
# +
from sklearn.metrics import accuracy_score
print('accuracy on train with NN:', model.evaluate(X_train, y_train)[1])
print('accuracy on test with NN:', model.evaluate(X_test, y_test)[1])
# -
# The accuracy is quite high for such a simple RNN: it took only a few lines of code and a couple of minutes to reach a 78 % accuracy!
| chapters/05-rnn-intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="euM-0r-85VDa" colab_type="text"
# # Install TensorFlow.
# * Install TensorFlow-1.14.0 with CUDA-10 support.
# * Install Keras-2.2.4, which compatible with TensorFlow-1.14.0.
# + id="iyh4xR_l5PV9" colab_type="code" colab={}
# %tensorflow_version 1.x
# !pip uninstall tensorflow -y
# !pip uninstall tensorflow-gpu -y
# !pip install --upgrade tensorflow-gpu==1.14.0
# !pip install --upgrade keras==2.2.4
# + [markdown] id="VaPC9vH86lww" colab_type="text"
# # Use installed TensorFlow version by running Runtime -> Restart runtime.
# + [markdown] id="vyy3gyE-6UMu" colab_type="text"
# # Install keras-contrib module.
# + id="1TlaM-SmI6OQ" colab_type="code" colab={}
# %tensorflow_version 1.x
# !sudo pip install git+https://www.github.com/keras-team/keras-contrib.git
# + [markdown] id="4OnZ0jJf685g" colab_type="text"
# # Import python modules.
# + id="ABMgyjrMXqyz" colab_type="code" colab={}
# %tensorflow_version 1.x
from keras.models import Model
from keras.models import Input
from keras.layers import Conv2D
# + id="KFjoDCm5xY99" colab_type="code" colab={}
from keras.layers import LeakyReLU
from keras_contrib.layers.normalization.instancenormalization import InstanceNormalization
# + [markdown] id="Zv8Fyp9YCsxR" colab_type="text"
# # Create the weight initializer.
# * Gaussian distribution - N(0.0, 0.02).
# + id="-qiZIkQyCmTL" colab_type="code" colab={}
from keras.initializers import RandomNormal
initializer = RandomNormal(mean=0.0, stddev=0.02)
# + [markdown] id="HMDcrWS1D0io" colab_type="text"
# # Create the optimizer.
# * Adam optimizer.
# * Learning rate = 0.0002 - First 100 epochs.
# * Learning rate is decreased from 0.0002 to 0.0 in 100 epochs.
# + id="ICG_1Z4C_XTb" colab_type="code" colab={}
from keras.optimizers import Adam
optimizer = Adam(lr=0.0002, beta_1=0.5)
# + [markdown] id="B2-lA-rS7OJf" colab_type="text"
# # Create the discriminator model.
# * Domain-A -> Discriminator-A -> [Real/Fake]
# * Domain-B -> Generator-A -> Discriminator-A -> [Real/Fake]
# * Domain-B -> Discriminator-B -> [Real/Fake]
# * Domain-A -> Generator-B -> Discriminator-B -> [Real/Fake]
#
#
#
#
# + id="2qgtOP7hhAIy" colab_type="code" colab={}
def create_discriminator(image_shape):
# Source image input.
input_image = Input(shape=image_shape)
# Create layer C64.
layer = Conv2D(64, (4,4), strides=(2,2), padding='same', kernel_initializer=initializer)(input_image)
layer = LeakyReLU(alpha=0.2)(layer)
# Create layer C128.
layer = Conv2D(128, (4,4), strides=(2,2), padding='same', kernel_initializer=initializer)(layer)
layer = InstanceNormalization(axis=-1)(layer)
layer = LeakyReLU(alpha=0.2)(layer)
# Create layer C256.
layer = Conv2D(256, (4,4), strides=(2,2), padding='same', kernel_initializer=initializer)(layer)
layer = InstanceNormalization(axis=-1)(layer)
layer = LeakyReLU(alpha=0.2)(layer)
# Create layer C512.
layer = Conv2D(512, (4,4), strides=(2,2), padding='same', kernel_initializer=initializer)(layer)
layer = InstanceNormalization(axis=-1)(layer)
layer = LeakyReLU(alpha=0.2)(layer)
# Create second last layer.
layer = Conv2D(512, (4,4), padding='same', kernel_initializer=initializer)(layer)
layer = InstanceNormalization(axis=-1)(layer)
layer = LeakyReLU(alpha=0.2)(layer)
# Create the discriminator output layer.
model_output = Conv2D(1, (4,4), strides=(1,1), padding='same', kernel_initializer=initializer)(layer)
# Create the discriminator model.
discriminator_model = Model(input_image, model_output)
# Compile the discriminator model.
# Loss - MSE - Weighted by 1/2 = 0.5
discriminator_model.compile(loss='mse', optimizer=optimizer, loss_weights=[0.5])
return( discriminator_model )
# + id="mu2mXqNg_Ev5" colab_type="code" colab={}
# Define input image shape.
image_shape = (256, 256, 3)
# Create the discriminator model.
model = create_discriminator(image_shape)
# Show the model summary.
model.summary()
# + [markdown] id="S5jmWbSGCVFi" colab_type="text"
# # Define the ResNet block.
# + id="B6juNSDJxfLV" colab_type="code" colab={}
from keras.layers import Activation
from keras.layers import Concatenate
# + id="F-mr8-H8goah" colab_type="code" colab={}
def create_resnet_block(number_of_filters, input_layer):
# Create first convolutional layer.
block_output = Conv2D(number_of_filters, (3,3), padding='same', kernel_initializer=initializer)(input_layer)
block_output = InstanceNormalization(axis=-1)(block_output)
block_output = Activation('relu')(block_output)
# Create second convolutional layer.
block_output = Conv2D(number_of_filters, (3,3), padding='same', kernel_initializer=initializer)(block_output)
block_output = InstanceNormalization(axis=-1)(block_output)
# Concatenate and merge channel-wise with input layer.
block_output = Concatenate()([block_output, input_layer])
return(block_output)
# + [markdown] id="Nz2ZfUYXvfoS" colab_type="text"
# # Create the generator model.
# * Domain-B -> Generator-A -> Domain-A
# * Domain-A -> Generator-B -> Domain-B
# + id="XGltPiYnxQSP" colab_type="code" colab={}
from keras.layers import Conv2DTranspose
# + id="5EjM_RxSjUnf" colab_type="code" colab={}
def create_generator(image_shape=(256,256,3), resnet_blocks=9):
# Create input image.
input_image = Input(shape=image_shape)
######################################################################
# c7s1-k - Conv2D-InstanceNormalization-ReLU block.
# - Conv2D 7×7 with k filters and stride 1.
# dk - Conv2D-InstanceNormalization-ReLU block.
# - Conv2D 3×3 with k filters and stride 2.
# Rk - Residual block
# - 2 3×3 Conv2D layers with k filters on both layers.
# uk - Fractional-strided Conv2D-InstanceNormalization-ReLU block.
# - Conv2D 3×3 with k filters and stride 1/2.
######################################################################
# Create c7s1-64 block.
layer = Conv2D(64, (7,7), strides=(1,1), padding='same', kernel_initializer=initializer)(input_image)
layer = InstanceNormalization(axis=-1)(layer)
layer = Activation('relu')(layer)
# Create d128 block.
layer = Conv2D(128, (3,3), strides=(2,2), padding='same', kernel_initializer=initializer)(layer)
layer = InstanceNormalization(axis=-1)(layer)
layer = Activation('relu')(layer)
# Create d256 block.
layer = Conv2D(256, (3,3), strides=(2,2), padding='same', kernel_initializer=initializer)(layer)
layer = InstanceNormalization(axis=-1)(layer)
layer = Activation('relu')(layer)
# Create R256 blocks.
for _ in range(resnet_blocks):
layer = create_resnet_block(256, layer)
# Create u128 block.
layer = Conv2DTranspose(128, (3,3), strides=(2,2), padding='same', kernel_initializer=initializer)(layer)
layer = InstanceNormalization(axis=-1)(layer)
layer = Activation('relu')(layer)
# Create u64 block.
layer = Conv2DTranspose(64, (3,3), strides=(2,2), padding='same', kernel_initializer=initializer)(layer)
layer = InstanceNormalization(axis=-1)(layer)
layer = Activation('relu')(layer)
# Create c7s1-3 block.
layer = Conv2D(3, (7,7), strides=(1,1), padding='same', kernel_initializer=initializer)(layer)
layer = InstanceNormalization(axis=-1)(layer)
output_image = Activation('tanh')(layer)
# Create the generator model.
generator_model = Model(input_image, output_image)
return( generator_model)
# + id="hgknbgQa1mvb" colab_type="code" colab={}
# Define input image shape.
image_shape = (256, 256, 3)
# Create the generator model.
model = create_generator(image_shape)
# Show the model summary.
model.summary()
# + [markdown] id="epcrBYet7ecf" colab_type="text"
# # Create a composite model for updating the generators using adversarial and cycle loss.
# + id="5FDHXNdVTqUb" colab_type="code" colab={}
def create_composite_model(generator_A_to_B, discriminator_2, generator_B_to_A, image_shape):
# Generator for domain A to B is trainable.
generator_A_to_B.trainable = True
# Discriminator for domain B is NOT trainable.
discriminator_2.trainable = False
# Generator for domain B to A is NOT trainable.
generator_B_to_A.trainable = False
# Define domain B discriminator output.
generator_input = Input(shape=image_shape)
generator_A2B_output = generator_A_to_B(generator_input)
discriminator_B_output = discriminator_2(generator_A2B_output)
# Define identity output.
identity_input = Input(shape=image_shape)
identity_output = generator_A_to_B(identity_input)
# Define forward cycle output.
forward_cycle_output = generator_B_to_A(generator_A2B_output)
# Define backward cycle output.
generator_B2A_output = generator_B_to_A(identity_input)
backward_cycle_output = generator_A_to_B(generator_B2A_output)
# Create the composite model.
composite_model = Model([generator_input, identity_input],
[discriminator_B_output, identity_output,
forward_cycle_output, backward_cycle_output])
# Compile the composite model with weighted loss using least squares loss and L1 loss.
composite_model.compile(loss=['mse', 'mae', 'mae', 'mae'], loss_weights=[1, 5, 10, 10], optimizer=optimizer)
return(composite_model)
# + id="5cAoon0TTmnn" colab_type="code" colab={}
# select a batch of random samples, returns images and target
def generate_real_samples(dataset, n_samples, patch_shape):
# choose random instances
ix = np.random.randint(0, dataset.shape[0], n_samples)
# retrieve selected images
X = dataset[ix]
# generate 'real' class labels (1)
y = np.ones((n_samples, patch_shape, patch_shape, 1))
return X, y
# + id="HOFtaJzKLgmg" colab_type="code" colab={}
# generate a batch of images, returns images and targets
def generate_fake_samples(g_model, dataset, patch_shape):
# generate fake instance
X = g_model.predict(dataset)
# create 'fake' class labels (0)
y = np.zeros((len(X), patch_shape, patch_shape, 1))
return X, y
# + id="GH1hkjwfLvEs" colab_type="code" colab={}
# update image pool for fake images
def update_image_pool(pool, images, max_size=50):
selected = list()
for image in images:
if len(pool) < max_size:
# stock the pool
pool.append(image)
selected.append(image)
elif np.random.random() < 0.5:
# use image, but don't add it to the pool
selected.append(image)
else:
# replace an existing image and use replaced image
ix = np.random.randint(0, len(pool))
selected.append(pool[ix])
pool[ix] = image
return np.asarray(selected)
# + id="-wJA3MkaTtC2" colab_type="code" colab={}
# train cyclegan models
def train(d_model_A, d_model_B, g_model_AtoB, g_model_BtoA, c_model_AtoB, c_model_BtoA, dataset, number_of_epochs=100, batch_size=1):
# determine the output square shape of the discriminator
n_patch = d_model_A.output_shape[1]
# unpack dataset
trainA, trainB = dataset
# prepare image pool for fakes
poolA, poolB = list(), list()
# calculate the number of batches per training epoch
bat_per_epo = int(len(trainA) / batch_size)
# calculate the number of training iterations
n_steps = bat_per_epo * number_of_epochs
# manually enumerate epochs
for i in range(n_steps):
# select a batch of real samples
X_realA, y_realA = generate_real_samples(trainA, batch_size, n_patch)
X_realB, y_realB = generate_real_samples(trainB, batch_size, n_patch)
# generate a batch of fake samples
X_fakeA, y_fakeA = generate_fake_samples(g_model_BtoA, X_realB, n_patch)
X_fakeB, y_fakeB = generate_fake_samples(g_model_AtoB, X_realA, n_patch)
# update fakes from pool
X_fakeA = update_image_pool(poolA, X_fakeA)
X_fakeB = update_image_pool(poolB, X_fakeB)
# update generator B->A via adversarial and cycle loss
g_loss2, _, _, _, _ = c_model_BtoA.train_on_batch([X_realB, X_realA], [y_realA, X_realA, X_realB, X_realA])
# update discriminator for A -> [real/fake]
dA_loss1 = d_model_A.train_on_batch(X_realA, y_realA)
dA_loss2 = d_model_A.train_on_batch(X_fakeA, y_fakeA)
# update generator A->B via adversarial and cycle loss
g_loss1, _, _, _, _ = c_model_AtoB.train_on_batch([X_realA, X_realB], [y_realB, X_realB, X_realA, X_realB])
# update discriminator for B -> [real/fake]
dB_loss1 = d_model_B.train_on_batch(X_realB, y_realB)
dB_loss2 = d_model_B.train_on_batch(X_fakeB, y_fakeB)
# summarize performance
print('>%d, dA[%.3f,%.3f] dB[%.3f,%.3f] g[%.3f,%.3f]' % (i+1, dA_loss1,dA_loss2, dB_loss1,dB_loss2, g_loss1,g_loss2))
# + id="UgZmJ2SvQ9yC" colab_type="code" colab={}
import os
import keras
dataset_url = 'https://people.eecs.berkeley.edu/~taesung_park/CycleGAN/datasets/horse2zebra.zip'
zip_file = keras.utils.get_file(origin=dataset_url,
fname="horse2zebra.zip",
extract=True)
#"ae_photos" "apple2orange" "summer2winter_yosemite" "horse2zebra" "monet2photo" "cezanne2photo" "ukiyoe2photo" "vangogh2photo" "maps" "cityscapes" "facades" "iphone2dslr_flower" "ae_photos"
base_dir = os.path.join(os.path.dirname(zip_file), 'horse2zebra')
# + id="RPf9e048SOmL" colab_type="code" colab={}
# !ls -al /root/.keras/datasets/horse2zebra
# + id="naz1Qlt8V_E-" colab_type="code" colab={}
import cv2
from glob import glob
import numpy as np
def load_data(domain, is_testing=False):
data_type = "train%s" % domain if not is_testing else "test%s" % domain
path = glob('%s/%s/*' % (base_dir,data_type))
imgs = []
for img_path in path:
img = cv2.imread(img_path, cv2.IMREAD_COLOR).astype(np.float)
if not is_testing:
img = cv2.resize(img, (256, 256))
if np.random.random() > 0.5:
img = np.fliplr(img)
else:
img = cv2.resize(img, (256, 256))
imgs.append(img)
imgs = np.array(imgs)/127.5 - 1.
return imgs
# + id="GG3bpSI6aow9" colab_type="code" colab={}
train_horses = load_data('A', is_testing=False)
train_zebras = load_data('B', is_testing=False)
print(len(train_horses))
print(len(train_zebras))
# + id="8Zvw0pfxveMa" colab_type="code" colab={}
test_horses = load_data('A', is_testing=True)
test_zebras = load_data('B', is_testing=True)
print(len(test_horses))
print(len(test_zebras))
# + id="iYn0YzWtTw5V" colab_type="code" colab={}
# load a dataset as a list of two numpy arrays
dataset = [train_horses, train_zebras]
# input shape
image_shape = (256,256,3)
# generator: A -> B
g_model_AtoB = create_generator(image_shape)
# generator: B -> A
g_model_BtoA = create_generator(image_shape)
# discriminator: A -> [real/fake]
d_model_A = create_discriminator(image_shape)
# discriminator: B -> [real/fake]
d_model_B = create_discriminator(image_shape)
# composite: A -> B -> [real/fake, A]
c_model_AtoB = create_composite_model(g_model_AtoB, d_model_B, g_model_BtoA, image_shape)
# composite: B -> A -> [real/fake, B]
c_model_BtoA = create_composite_model(g_model_BtoA, d_model_A, g_model_AtoB, image_shape)
number_of_epochs = 1
batch_size = 1
# train models
train(d_model_A, d_model_B, g_model_AtoB, g_model_BtoA, c_model_AtoB, c_model_BtoA, dataset, number_of_epochs, batch_size)
# + id="F1hwO97CdOZY" colab_type="code" colab={}
g_model_AtoB.save_weights('g_model_AtoB.h5')
g_model_BtoA.save_weights('g_model_BtoA.h5')
# + id="STdqk09LvFCG" colab_type="code" colab={}
output_images = g_model_AtoB.predict(test_horses)
for input_image, output_image in zip(test_horses, output_images):
input_image = (input_image + 1) * 127.5
output_image = (output_image + 1) * 127.5
cv2.imwrite('input_image.png', input_image)
cv2.imwrite('output_image.png', output_image)
| GAN/CycleGAN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
width=0.01
x = np.arange(0, 1, width)
y = x**2
plt.plot(x, y)
plt.xlim(0, 1)
plt.ylim(0, 1)
plt.show()
# -
data = tuple(np.arange(0, 1, width))
y_pos = np.arange(len(data))
plt.bar(y_pos, y, align='center', width=1, alpha=0.5)
plt.plot(y_pos, y)
plt.xlim(0, 1)
plt.ylim(0, 1)
plt.xticks(y_pos, data)
#
# $$
# \begin{align*}
# I &=\int_{0}^{1} f(x) dx = \int_{0}^{1}x^2 dx \\
# &=\tfrac{1}{3} x^3
# \Big|_0^1=\tfrac{1}{3} 1^3 -0=\tfrac{1}{3}
# \end{align*}
# $$
# +
from scipy.integrate import quad
def integrand(x, a, b):
return a*x**2 + b
a = 1
b = 0
I = quad(integrand, 0, 1, args=(a,b))
I
# +
cintegral = width * sum(y)
print(cintegral,(1/3)-cintegral)
# -
| student_pyw3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: dviz
# language: python
# name: dviz
# ---
# # W3 Lab: Perception
#
# In this lab, we will learn basic usage of `pandas` library and then perform a small experiment to test the Stevens' power law.
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# ## Vega datasets
#
# Before going into the perception experiment, let's first talk about some handy datasets that you can play with.
#
# It's nice to have clean datasets handy to practice data visualization. There is a nice small package called [`vega-datasets`](https://github.com/altair-viz/vega_datasets), from the [altair project](https://github.com/altair-viz).
#
# You can install the package by running
#
# $ pip install vega-datasets
#
# or
#
# $ pip3 install vega-datasets
#
# Once you install the package, you can import and see the list of datasets:
# +
from vega_datasets import data
data.list_datasets()
# -
# or you can work with only smaller, local datasets.
from vega_datasets import local_data
local_data.list_datasets()
# Ah, we have the `anscombe` data here! Let's see the description of the dataset.
local_data.anscombe.description
# ## Anscombe's quartet dataset
#
# How does the actual data look like? Very conveniently, calling the dataset returns a Pandas dataframe for you.
df = local_data.anscombe()
df.head()
# **Q1: can you draw a scatterplot of the dataset "I"?** You can filter the dataframe based on the `Series` column and use `plot` function that you used for the Snow's map.
# TODO: put your code here
# ## Some histograms with pandas
# Let's look at a slightly more complicated dataset.
car_df = local_data.cars()
car_df.head()
# Pandas provides useful summary functions. It identifies numerical data columns and provides you with a table of summary statistics.
car_df.describe()
# If you ask to draw a histogram, you get all of them. :)
car_df.hist()
# Well this is too small. You can check out [the documentation](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.hist.html) and change the size of the figure.
#
# **Q2: by consulting the documentation, can you make the figure larger so that we can see all the labels clearly? And then make the layout 2 x 3 not 3 x 2, then change the number of bins to 20?**
# TODO: put your code here
# ## Stevens’ power-law and your own psychophysics experiment!
# Let's do an experiment! The procedure is as follows:
#
# 1. Generate a random number between \[1, 10\];
# 1. Use a horizontal bar to represent the number, i.e., the length of the bar is equal to the number;
# 1. Guess the length of the bar by comparing it to two other bars with length 1 and 10 respectively;
# 1. Store your guess (perceived length) and actual length to two separate lists;
# 1. Repeat the above steps many times;
# 1. Check whether Steven's power-law holds.
#
# First, let's define the length of a short and a long bar. We also create two empty lists to store perceived and actual length.
# +
import random
import time
import numpy as np
l_short_bar = 1
l_long_bar = 10
perceived_length_list = []
actual_length_list = []
# -
# ### Perception of length
#
# Let's run the experiment.
#
# The [**`random`**](https://docs.python.org/3.6/library/random.html) module in Python provides various random number generators, and the [**`random.uniform(a,b)`**](https://docs.python.org/3.6/library/random.html#random.uniform) function returns a floating point number in \[a,b\].
#
# We can plot horizontal bars using the [**`pyplot.barh()`**](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.barh) function. Using this function, we can produce a bar graph that looks like this:
# +
mystery_length = random.uniform(1, 10) # generate a number between 1 and 10. this is the *actual* length.
plt.barh(np.arange(3), [l_short_bar, mystery_length, l_long_bar], align='center')
plt.yticks(np.arange(3), ('1', '?', '10'))
plt.xticks([]) # no hint!
# -
# Btw, `np.arange` is used to create a simple integer list `[0, 1, 2]`.
np.arange(3)
# Now let's define a function to perform the experiment once. When you run this function, it picks a random number between 1.0 and 10.0 and show the bar chart. Then it asks you to input your estimate of the length of the middle bar. It then saves that number to the `perceived_length_list` and the actual answer to the `actual_length_list`.
def run_exp_once():
mystery_length = random.uniform(1, 10) # generate a number between 1 and 10.
plt.barh(np.arange(3), [l_short_bar, mystery_length, l_long_bar], height=0.5, align='center')
plt.yticks(np.arange(3), ('1', '?', '10'))
plt.xticks([]) # no hint!
plt.show()
perceived_length_list.append( float(input()) )
actual_length_list.append(mystery_length)
run_exp_once()
# Now, run the experiment many times to gather your data. Check the two lists to make sure that you have the proper dataset. The length of the two lists should be the same.
# TODO: Run your experiment many times here
# ### Plotting the result
#
# Now we can draw the scatter plot of perceived and actual length. The `matplotlib`'s [**`scatter()`**](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.scatter) function will do this. This is the backend of the pandas' scatterplot. Here is an example of how to use `scatter`:
plt.scatter(x=[1,5,10], y=[1,10, 5])
# **Q3: Now plot your result using the `scatter()` function. You should also use `plt.title()`, `plt.xlabel()`, and `plt.ylabel()` to label your axes and the plot itself.**
# TODO: put your code here
# After plotting, let's fit the relation between actual and perceived lengths using a polynomial function. We can easily do it using [**`curve_fit(f, x, y)`**](http://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.curve_fit.html) in Scipy, which is to fit $x$ and $y$ using the function `f`. In our case, $f = a*x^b +c$. For instance, we can check whether this works by creating a fake dataset that follows the exact form:
# +
from scipy.optimize import curve_fit
def func(x, a, b, c):
return a * np.power(x, b) + c
x = np.arange(20) # [0,1,2,3, ..., 19]
y = np.power(x, 2) # [0,1,4,9, ... ]
popt, pcov = curve_fit(func, x, y)
print('{:.2f} x^{:.2f} + {:.2f}'.format(*popt))
# -
# **Q4: Now fit your data!** Do you see roughly linear relationship between the actual and the perceived lengths? It's ok if you don't!
# TODO: your code here
# ### Perception of area
#
# Similar to the above experiment, we now represent a random number as a circle, and the area of the circle is equal to the number.
#
# First, calculate the radius of a circle from its area and then plot using the **`Circle()`** function. `plt.Circle((0,0), r)` will plot a circle centered at (0,0) with radius `r`.
# +
n1 = 0.005
n2 = 0.05
radius1 = np.sqrt(n1/np.pi) # area = pi * r * r
radius2 = np.sqrt(n2/np.pi)
random_radius = np.sqrt(n1*random.uniform(1,10)/np.pi)
plt.axis('equal')
plt.axis('off')
circ1 = plt.Circle( (0,0), radius1, clip_on=False )
circ2 = plt.Circle( (4*radius2,0), radius2, clip_on=False )
rand_circ = plt.Circle((2*radius2,0), random_radius, clip_on=False )
plt.gca().add_artist(circ1)
plt.gca().add_artist(circ2)
plt.gca().add_artist(rand_circ)
# -
# Let's have two lists for this experiment.
perceived_area_list = []
actual_area_list = []
# And define a function for the experiment.
def run_area_exp_once(n1=0.005, n2=0.05):
radius1 = np.sqrt(n1/np.pi) # area = pi * r * r
radius2 = np.sqrt(n2/np.pi)
mystery_number = random.uniform(1,10)
random_radius = np.sqrt(n1*mystery_number/math.pi)
plt.axis('equal')
plt.axis('off')
circ1 = plt.Circle( (0,0), radius1, clip_on=False )
circ2 = plt.Circle( (4*radius2,0), radius2, clip_on=False )
rand_circ = plt.Circle((2*radius2,0), random_radius, clip_on=False )
plt.gca().add_artist(circ1)
plt.gca().add_artist(circ2)
plt.gca().add_artist(rand_circ)
plt.show()
perceived_area_list.append( float(input()) )
actual_area_list.append(mystery_number)
# **Q5: Now you can run the experiment many times, plot the result, and fit a power-law curve to test the Stevens' power-law!**
# +
# TODO: put your code here. You can use multiple cells.
# -
# What is your result? How are the exponents different from each other? Have you observed a result consistent with the Stevens' power-law?
| m04-perception/lab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Let's Create a Simple Blockchain
# At the very core, we want to create a linked list. Each block points to the fingerprint of the previous block. Additionally, we include a checksum of the content of each block (i.e., fingerprint).
# <img src="include/block_diagram.png" alt="block diagram/" style="width: 300px;"/>
# ## *<font color=" #6495ED">Exercise</font>*
#
# - Define a class called "block" to represent the structure above
# SOLUTION
test_block = block(b"1", b"DUMMY DATA")
test_block
# ## Genesis Block
#
# As we can now we can represent each block, and we can point to the previous block. Any modification to any block will propagate to the other blocks. If the data in any block changes, the hash changes, and this would reflect in the consequent blocks.
#
# BUT, where does the first block come from? There is no previous block to the first block.
#
# We should define one block as the **genesis block** where every other block agrees upon.
# ## *<font color=" #6495ED">Exercise</font>*
#
# - Define your genesis block, with following parameters:
# - previous hash: "0000000000000000000000000000000000000000000000000000000000000000
# - nonce: b"FFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFFF"
# - data: "PyCon 2018 Genesis Block"
# SOLUTION
# ## Putting the chain into blockchain
# In our simple blockchain, we will use a *list* to store each block of our blockchain.
# ## *<font color=" #6495ED">Exercise</font>*
# - Define a class called "blockchain", define a the following functions
# - "\__init__": initialize a empty list
# - "add_blocks": simply append a block to the list, (block is passed as an argument)
# - "blockchain_valid": to check if the blockchain is valid, for now just return True
# SOLUTION
test_blockchain = blockchain()
test_blockchain
test_blockchain.blocks
test_blockchain.add_block(genesis_block)
test_blockchain
test_blockchain.blocks
# ## Blockchain validity
# Now we want to check if our blockchain is valid. In our example, we want to check if the hash of each block is correct. First we need to create a hash of the block header + data and check if it is correct. Furthermore, we need to check if the previous hash in each block is correct.
# ## *<font color=" #6495ED">Exercise</font>*
# - Implement the "blockchain_valid", according to the description
# +
# SOLUTION
# -
# Good Genesis Block
test_blockchain = blockchain()
test_blockchain.add_block(genesis_block)
print(test_blockchain.blockchain_valid()) # this should be True
# Bad Genesis Block
test_blockchain = blockchain()
test_blockchain.add_block(block(b"1", b"2"))
print(test_blockchain.blockchain_valid(), "--Bad Genesis Block--") # this should be False
# +
# Blocks are correct
test_blockchain = blockchain()
test_blockchain.add_block(genesis_block)
print(test_blockchain.blockchain_valid())
print(genesis_block)
test_block = block(genesis_block.hash, b"DUMMY DATA")
print(test_block)
test_blockchain.add_block(test_block)
test_blockchain.blockchain_valid()
# -
test_block2 = block(test_block.hash, b"DUMMY DATA2")
print(test_block2)
test_blockchain.add_block(test_block2)
test_blockchain.blockchain_valid()
print(test_blockchain)
# ## Proof of Work
#
# Well, now we have a basic blockchain working. However, there is one small problem, everyone can create blocks and chain them together. There is not much effort needed to create new blocks, hash them and add them to the ***list***.
#
# ### Hashing to the rescue
#
# Why not make the creation of valid blocks hard. Then, we only accepts the blocks that have certain amount of work and effort done on them as valid. Each block hash should start with certain amount of zeros. We have set the number of zeros dynamically to be adapt with new technology and also number of ***block truthfulness seekers*** (miners).
# ## *<font color=" #6495ED">Exercise</font>*
# - Let's put this in code. Set the number of leading zeros to 3.
# SOLUTION
# Blocks are correct
test_blockchain = blockchain()
test_blockchain.add_block(genesis_block)
test_blockchain
test_block = block(genesis_block.hash, b"DUMMY DATA")
test_block
test_block.find_nonce(3)
test_block
test_blockchain.add_block(test_block)
test_blockchain.blockchain_valid()
test_block2 = block(test_block.hash, b"DUMMY DATA2")
print(test_block2, "\n")
test_block2.find_nonce(3)
print(test_block2)
test_blockchain.add_block(test_block2)
test_blockchain.blockchain_valid()
test_blockchain
# ## The tale of two blockchains
#
# Once again, we have something working. But there is one more problem, if you have to ***branches*** of the blockchain that are ***valid***, how to decide which one to use. We only want to have one true blockchain branch.
#
# Why not go with the blockchain that has the most effort and work done on it then? This translates into selecting the blockchain with the longest chain length. Easy!
# +
test_blockchain1 = blockchain()
test_blockchain2 = blockchain()
test_blockchain1.add_block(genesis_block)
test_blockchain2.add_block(genesis_block)
print(test_blockchain1)
print()
print(test_blockchain2)
# -
# ## *<font color=" #6495ED">Exercise</font>*
# - Set the difficulty to 3 and start creating two branches of the blockchain for 10 second each.
# SOLUTION
mine_blocks(test_blockchain1)
mine_blocks(test_blockchain2)
len(test_blockchain1.blocks)
len(test_blockchain2.blocks)
# Congratulations! You just created a simple blockchain, with mining, dynamic difficulty level, proof of work, and longest chain consensus algorithm. Now, what does it take to make our blockchain into a cryptocurrency?
# ## Cryptocurrency
# To transfer our correct blockchain into a cryptocurrency, instead of storing random data, we need to store transaction data.
#
# - From: Alice
# - To: Bob
# - Amount: 10$
# ## *<font color=" #6495ED">Exercise</font>*
# - If we just store the transactions, what stops anyone to put wrong transactions there?
# - Bob can just create a message indicating Alice-> Bob 100000..0000$
# ## Aysmmetric Encryption System and Signatures
#
# Fortunately, it's not too hard to mitigate against this.
#
# - Instead of Alice, Bob, X let's use $H(pub(Alice))$, $H(pub(Bob))$, $H(pub(X))$. $ADDR_X=H(pub(X))$
# - Now just sign the hash of a transaction with your private key
# - $S_X(H(ADDR_X -> ADDR_Y: 10\$))$, S_X, signing using private key of X
#
#TX data from, to, amount
tx_data = "{0}======={1}======={2}"
# +
from cryptography.hazmat.backends import default_backend
from cryptography.hazmat.primitives import hashes
from cryptography.hazmat.primitives.asymmetric import ec
from cryptography.hazmat.primitives.serialization import PublicFormat
from cryptography.hazmat.primitives.serialization import Encoding
from cryptography.hazmat.primitives import serialization
import base64
def hash_pub_key(private_key):
digest = hashes.Hash(hashes.SHA256(), backend=default_backend())
digest.update(private_key.public_key().public_bytes(Encoding.DER, PublicFormat.SubjectPublicKeyInfo))
return base64.b16encode(digest.finalize())
def sign_tx(tx, private_key):
return private_key.sign(tx, ec.ECDSA(hashes.SHA256()))
def serialize_pubkey(publickey):
serialized_public = publickey.public_bytes(
encoding=Encoding.PEM,
format=PublicFormat.SubjectPublicKeyInfo)
return serialized_public
def parse_serialized_pubkey(serialized_pubkey):
loaded_public_key = serialization.load_pem_public_key(serialized_pubkey, backend=default_backend())
return loaded_public_key
# -
# ## *<font color=" #6495ED">Exercise</font>*
# - Define three private keys (Alice, Bob, Carol) and H(pubkey) for each
# SOLUTION
# ## Creating a transaction + signature
# Alice -> Bob, 10$
tx_data_b = str.encode(tx_data.format(hash_pub_key(private_keys["Alice"]),
hash_pub_key(private_keys["Bob"]),
"10"))
serialized_pubkey = serialize_pubkey(private_keys["Alice"].public_key())
siganture = sign_tx(tx_data_b, private_keys["Alice"])
# transaction data
tx_b = tx_data_b+b'======='+serialized_pubkey+b'======='+siganture
# Bob -> Carol, 15$
tx_data_b = str.encode(tx_data.format(hash_pub_key(private_keys["Bob"]),
hash_pub_key(private_keys["Carol"]),
"15"))
serialized_pubkey = serialize_pubkey(private_keys["Bob"].public_key())
siganture = sign_tx(tx_data_b, private_keys["Bob"])
# transaction data
tx_b2 = tx_data_b+b'======='+serialized_pubkey+b'======='+siganture
# Carol -> Alice, 20$
tx_data_b = str.encode(tx_data.format(hash_pub_key(private_keys["Carol"]),
hash_pub_key(private_keys["Alice"]),
"20"))
serialized_pubkey = serialize_pubkey(private_keys["Carol"].public_key())
siganture = sign_tx(tx_data_b, private_keys["Carol"])
# transaction data
tx_b3 = tx_data_b+b'======='+serialized_pubkey+b'======='+siganture
# ## *<font color=" #6495ED">Exercise</font>*
# - Let's add 1 transactions to each block
# - Check if the signature is correct
# SOLUTION
# Blocks are correct
test_blockchain = blockchain()
test_blockchain.add_block(genesis_block)
# +
test_block = block(genesis_block.hash, tx_b)
test_block.find_nonce(3)
test_blockchain.add_block(test_block)
test_block = block(test_block.hash, tx_b2)
test_block.find_nonce(3)
test_blockchain.add_block(test_block)
test_block = block(test_block.hash, tx_b3)
test_block.find_nonce(3)
test_blockchain.add_block(test_block)
test_blockchain.blockchain_valid()
# -
test_blockchain
for _block in test_blockchain.blocks[1:]:
verify_tx_signatures(_block.data)
# ## *<font color=" #6495ED">Exercise</font>*
# - How to make sure the money is not already spent? (double spending)
| 3. Blockchain.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python
# name: conda-env-python-py
# ---
# <h3 align=center>List to Set </h3>
# Cast the following list to a set:
['A','B','C','A','B','C']
# <h3 align=center>Add an Element to the Set</h3>
# Add the string <code>'D'</code> to the set S
S={'A','B','C'}
S.add('D')
print(S)
# <h3 align=center>Intersection</h3>
# Find the intersection of set <code>A</code> and <code>B</code>
# + jupyter={"outputs_hidden": false}
A={1,2,3,4,5}
B={1,3,9, 12}
print(B in A)
# -
# <hr>
# <small>Copyright © 2018 IBM Cognitive Class. This notebook and its source code are released under the terms of the [MIT License](https://cognitiveclass.ai/mit-license/).</small>
| 2.2_notebook_quizz_sets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Una persona que va de compras a la tienda “Enano, S.A.”, decide llevar un control
#sobre lo que va comprando, para saber la cantidad de dinero que tendrá que pagar al llegar a
#la caja. La tienda tiene una promoción del 20% de descuento sobre aquellos artículos cuya
#etiqueta sea roja. Determinar la cantidad de dinero que esta persona deberá pagar.
precio_total = 0
while True:
print 'Cual es el valor del producto?'
precio = float(input())
print 'Que tipo de etiqueta tiene?(roja/blanca)'
tipo_etiqueta = str(raw_input())
if tipo_etiqueta == 'Roja' or tipo_etiqueta == 'roja':
precio = precio * 0.8
precio_total = precio_total + precio
print 'Desea ingresar otro producto?'
opcion = str(raw_input())
if opcion == "Si" or opcion == "si":
continue
else:
break
print 'El costo total de los productos a comprar es: ' + str(precio_total) + ' pesos:'
| 3.algoritmo_ciclo_mientras/Ejercicio_8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
import pickle
import json
import pandas as pd
import numpy as np
from sklearn.metrics import classification_report, accuracy_score
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
import torch.nn as nn
from matplotlib import pyplot as plt
import spacy
import re
nlp = spacy.load("en_core_web_sm")
# +
dirname = '../Dataset_v3/chunks/CharlesDickens/ATaleOfTwoCities_chunkByLength_256.csv'
charles1 = pd.read_csv(dirname)
print("Charles ATaleOfTwoCities_chunkByLength_256: {0} ".format(charles1.shape))
dirname = '../Dataset_v3/chunks/CharlesDickens/BarnabyRudge_chunkByLength_256.csv'
charles2 = pd.read_csv(dirname)
print("charles BarnabyRudge_chunkByLength_256: {0}" .format(charles2.shape))
dirname = '../Dataset_v3/chunks/CharlesDickens/DavidCopperfield_chunkByLength_256.csv'
charles3 = pd.read_csv(dirname)
print("charles DavidCopperfield_chunkByLength_256: {0}" .format(charles3.shape))
dirname = '../Dataset_v3/chunks/CharlesDickens/DombeyAndSon_chunkByLength_256.csv'
charles4 = pd.read_csv(dirname)
print("charles DombeyAndSon_chunkByLength_256: {0}" .format(charles4.shape))
dirname = '../Dataset_v3/chunks/CharlesDickens/GreatExpectations_chunkByLength_256.csv'
charles5 = pd.read_csv(dirname)
print("charles GreatExpectations_chunkByLength_256: {0}" .format(charles5.shape))
dirname = '../Dataset_v3/chunks/CharlesDickens/LittleDorrit_chunkByLength_256.csv'
charles6 = pd.read_csv(dirname)
print("charles LittleDorrit_chunkByLength_256: {0}" .format(charles6.shape))
dirname = '../Dataset_v3/chunks/CharlesDickens/OliverTwist_chunkByLength_256.csv'
charles7 = pd.read_csv(dirname)
print("charles OliverTwist_chunkByLength_256: {0}" .format(charles7.shape))
dirname = '../Dataset_v3/chunks/CharlesDickens/OurMutualFriend_chunkByLength_256.csv'
charles8 = pd.read_csv(dirname)
print("charles OurMutualFriend_chunkByLength_256: {0}" .format(charles8.shape))
dirname = '../Dataset_v3/chunks/CharlesDickens/TheChimes_chunkByLength_256.csv'
charles9 = pd.read_csv(dirname)
print("charles TheChimes_chunkByLength_256: {0}" .format(charles9.shape))
dirname = '../Dataset_v3/chunks/CharlesDickens/TheLifeAndAdventuresOfMartinChuzzlewit_chunkByLength_256.csv'
charles10 = pd.read_csv(dirname)
print("charles TheLifeAndAdventuresOfMartinChuzzlewit_chunkByLength_256: {0}" .format(charles10.shape))
dirname = '../Dataset_v3/chunks/CharlesDickens/TheLifeAndAdventuresOfNicholasNickleby_chunkByLength_256.csv'
charles11 = pd.read_csv(dirname)
print("charles TheLifeAndAdventuresOfNicholasNickleby_chunkByLength_256: {0}" .format(charles11.shape))
dirname = '../Dataset_v3/chunks/CharlesDickens/TheMysteryOfEdwinDrood_chunkByLength_256.csv'
charles12 = pd.read_csv(dirname)
print("charles TheMysteryOfEdwinDrood_chunkByLength_256: {0}" .format(charles12.shape))
dirname = '../Dataset_v3/chunks/CharlesDickens/TheOldCuriosityShop_chunkByLength_256.csv'
charles13 = pd.read_csv(dirname)
print("charles TheOldCuriosityShop_chunkByLength_256: {0}" .format(charles13.shape))
dirname = '../Dataset_v3/chunks/CharlesDickens/ThePickwickPapers_chunkByLength_256.csv'
charles14 = pd.read_csv(dirname)
print("charles ThePickwickPapers_chunkByLength_256: {0}" .format(charles14.shape))
# +
dirname = '../Dataset_v3/chunks/Fitzgerald/The_Great_Gatsby_chunkByLength_256.csv'
Fitzgerald1 = pd.read_csv(dirname)
print("Fitzgerald The_Great_Gatsby: {0}".format(Fitzgerald1.shape))
dirname = '../Dataset_v3/chunks/Fitzgerald/TheBeautifulAndDamned_chunkByLength_256.csv'
Fitzgerald2 = pd.read_csv(dirname)
print("Fitzgerald TheBeautifulAndDamned_chunkByLength_256: {0}".format(Fitzgerald2.shape))
dirname = '../Dataset_v3/chunks/Fitzgerald/ThisSideOfParadise_chunkByLength_256.csv'
Fitzgerald3 = pd.read_csv(dirname)
print("Fitzgerald ThisSideOfParadise_chunkByLength_256: {0}".format(Fitzgerald3.shape))
dirname = '../Dataset_v3/chunks/Fitzgerald/FlappersAndPhilosophers_chunkByLength_256.csv'
Fitzgerald4 = pd.read_csv(dirname)
print("Fitzgerald FlappersAndPhilosophers_chunkByLength_256: {0}".format(Fitzgerald4.shape))
dirname = '../Dataset_v3/chunks/Fitzgerald/TalesOfTheJazzAge_chunkByLength_256.csv'
Fitzgerald5 = pd.read_csv(dirname)
print("Fitzgerald TalesOfTheJazzAge_chunkByLength_256: {0}".format(Fitzgerald5.shape))
# +
dirname = '../Dataset_v3/chunks/Jane/Pride_and_Prejudice_Jane_Austen_chunkByLength_256.csv'
Jane1 = pd.read_csv(dirname)
print("Jane Pride_and_Prejudice_Jane_Austen_chunkByLength_256: {0}".format(Jane1.shape))
dirname = '../Dataset_v3/chunks/Jane/Emma_Jane_Austen_chunkByLength_256.csv'
Jane2 = pd.read_csv(dirname)
print("Jane Emma_Jane_Austen_chunkByLength_256: {0}".format(Jane2.shape))
dirname = '../Dataset_v3/chunks/Jane/MansfieldPark_chunkByLength_256.csv'
Jane3 = pd.read_csv(dirname)
print("Jane MansfieldPark_chunkByLength_256: {0}".format(Jane3.shape))
dirname = '../Dataset_v3/chunks/Jane/NorthangerAbbey_chunkByLength_256.csv'
Jane4 = pd.read_csv(dirname)
print("Jane NorthangerAbbey_chunkByLength_256: {0}".format(Jane4.shape))
dirname = '../Dataset_v3/chunks/Jane/Persuasion_chunkByLength_256.csv'
Jane5 = pd.read_csv(dirname)
print("Jane Persuasion_chunkByLength_256: {0}".format(Jane5.shape))
dirname = '../Dataset_v3/chunks/Jane/SenseAndSensibility_chunkByLength_256.csv'
Jane6 = pd.read_csv(dirname)
print("Jane SenseAndSensibility_chunkByLength_256: {0}".format(Jane6.shape))
# +
dirname = '../Dataset_v3/chunks/Mark/TheAdventuresOfTomSawyer_chunkByLength_256.csv'
Mark1 = pd.read_csv(dirname)
print("Mark TheAdventuresOfTomSawyer: {0}".format(Mark1.shape))
dirname = '../Dataset_v3/chunks/Mark/ThePrinceAndThePauper_chunkByLength_256.csv'
Mark2 = pd.read_csv(dirname)
print("Mark ThePrinceAndThePauper_chunkByLength_256: {0}".format(Mark2.shape))
dirname = '../Dataset_v3/chunks/Mark/AConnecticutYankeeInKingArthursCourt_chunkByLength_256.csv'
Mark3 = pd.read_csv(dirname)
print("Mark AConnecticutYankeeInKingArthursCourt_chunkByLength_256: {0}".format(Mark3.shape))
dirname = '../Dataset_v3/chunks/Mark/LifeOnTheMississippi_chunkByLength_256.csv'
Mark4 = pd.read_csv(dirname)
print("Mark LifeOnTheMississippi_chunkByLength_256: {0}".format(Mark4.shape))
dirname = '../Dataset_v3/chunks/Mark/TheMysteriousStranger_chunkByLength_256.csv'
Mark5 = pd.read_csv(dirname)
print("Mark TheMysteriousStranger_chunkByLength_256: {0}".format(Mark5.shape))
dirname = '../Dataset_v3/chunks/Mark/TheTragedyOfPuddnheadWilson_chunkByLength_256.csv'
Mark6 = pd.read_csv(dirname)
print("Mark TheTragedyOfPuddnheadWilson_chunkByLength_256: {0}".format(Mark6.shape))
# +
dirname = '../Dataset_v3/chunks/Various/AlicesAdventuresInWonderlandByLewisCarroll_chunkByLength_256.csv'
Various1 = pd.read_csv(dirname)
print("Various AlicesAdventuresInWonderlandByLewisCarroll_chunkByLength_256: {0}".format(Various1.shape))
dirname = '../Dataset_v3/chunks/Various/AModestProposalByJonathanSwift_chunkByLength_256.csv'
Various2 = pd.read_csv(dirname)
print("Various AModestProposalByJonathanSwift_chunkByLength_256: {0}".format(Various2.shape))
dirname = '../Dataset_v3/chunks/Various/FrankensteinOrTheModernPrometheusByMaryWollstonecraftShelley_chunkByLength_256.csv'
Various3 = pd.read_csv(dirname)
print("Various FrankensteinOrTheModernPrometheusByMaryWollstonecraftShelley_chunkByLength_256: {0}".format(Various3.shape))
dirname = '../Dataset_v3/chunks/Various/MetamorphosisByFranzKafka_chunkByLength_256.csv'
Various4 = pd.read_csv(dirname)
print("Various MetamorphosisByFranzKafka_chunkByLength_256: {0}".format(Various4.shape))
dirname = '../Dataset_v3/chunks/Various/MobyDickOrTheWhalebyHermanMelville_chunkByLength_256.csv'
Various5 = pd.read_csv(dirname)
print("Various MobyDickOrTheWhalebyHermanMelville_chunkByLength_256: {0}".format(Various5.shape))
dirname = '../Dataset_v3/chunks/Various/TheAdventuresofSherlockHolmesbyArthurConanDoyle_chunkByLength_256.csv'
Various6 = pd.read_csv(dirname)
print("Various TheAdventuresofSherlockHolmesbyArthurConanDoyle_chunkByLength_256: {0}".format(Various6.shape))
dirname = '../Dataset_v3/chunks/Various/TheImportanceOfBeingEarnestATrivialComedyForSeriousPeopleByOscarWilde_chunkByLength_256.csv'
Various7 = pd.read_csv(dirname)
print("Various TheImportanceOfBeingEarnestATrivialComedyForSeriousPeopleByOscarWilde_chunkByLength_256: {0}".format(Various7.shape))
dirname = '../Dataset_v3/chunks/Various/TheScarletLetterbyNathanielHawthorne_chunkByLength_256.csv'
Various8 = pd.read_csv(dirname)
print("Various TheScarletLetterbyNathanielHawthorne_chunkByLength_256: {0}".format(Various8.shape))
dirname = '../Dataset_v3/chunks/Various/TheStrangeCaseofDrJekyllandMrHydebyRobertLouisStevenson_chunkByLength_256.csv'
Various9 = pd.read_csv(dirname)
print("Various TheStrangeCaseofDrJekyllandMrHydebyRobertLouisStevenson_chunkByLength_256: {0}".format(Various9.shape))
dirname = '../Dataset_v3/chunks/Various/TheYellowWallpaperbyCharlottePerkinsGilman_chunkByLength_256.csv'
Various10 = pd.read_csv(dirname)
print("Various TheYellowWallpaperbyCharlottePerkinsGilman_chunkByLength_256: {0}".format(Various10.shape))
# -
charleslist = [charles1, charles2, charles3, charles4, charles5, charles6, charles7,
charles8, charles9, charles10, charles11,charles12,charles13,charles14]
marklist = [Mark1, Mark2, Mark3, Mark4, Mark5, Mark6]
variouslsit = [Various1, Various2, Various3, Various4, Various5, Various6, Various7, Various8,
Various9, Various10]
janelist = [Jane1, Jane2, Jane3, Jane4, Jane5, Jane6]
fitzgeraldlist = [Fitzgerald1, Fitzgerald2, Fitzgerald3, Fitzgerald4,Fitzgerald5]
# +
# 14240 <NAME>
# 2043 <NAME>
# 2405 various
# 3033 <NAME>
# 1625 Fitzgerald
totcount = 0
for df in fitzgeraldlist:
totcount += df.shape[0]
totcount
# -
# * <NAME>: Mark + Various
# * Fitzgerald: Fitzgerald + Various
# * Charles: Charles + all
# +
df_charles = pd.concat(charleslist,ignore_index=True)
df_charles['Label'] = "1"
df_charles = df_charles[['Text','Label']]
df_others = pd.concat(marklist +variouslsit+ janelist +fitzgeraldlist,ignore_index=True)
df_others['Label'] = "0"
df_others = df_others[['Text','Label']]
df_charles_f = pd.concat([df_charles,df_others],ignore_index=True)
print(df_charles_f.shape[0])
df_charles_f.Label.describe()
# +
df_mark = pd.concat(marklist,ignore_index=True)
df_mark['Label'] = "1"
df_mark = df_mark[['Text','Label']]
df_others = pd.concat(variouslsit,ignore_index=True)
df_others['Label'] = "0"
df_others = df_others[['Text','Label']]
df_mark_f = pd.concat([df_mark,df_others],ignore_index=True)
print(df_mark_f.shape[0])
df_mark_f.Label.describe()
# +
df_fitzgerald = pd.concat(fitzgeraldlist,ignore_index=True)
df_fitzgerald['Label'] = "1"
df_fitzgerald = df_fitzgerald[['Text','Label']]
df_others = pd.concat(variouslsit,ignore_index=True)
df_others['Label'] = "0"
df_others = df_others[['Text','Label']]
df_fitzgerald_f = pd.concat([df_fitzgerald,df_others],ignore_index=True)
print(df_fitzgerald_f.shape[0])
df_fitzgerald_f.Label.describe()
# -
# apply on the df['Text']
# retain all columns and add addtional columns spacy_words
def NER_replace(df):
df['spacy_words'] = df['Text'].apply(lambda x: list(nlp(x).ents))
for index, row in df.iterrows():
thistext = row.Text
for ent in row.spacy_words:
if ent.label_ in ['PERSON','FAC','GPE','LOC','ORG']:
thistext = thistext.replace(ent.text,ent.label_)
df.at[index, 'Text'] = thistext
return df
df_fitzgerald_f = NER_replace(df_fitzgerald_f)
df_mark_f = NER_replace(df_mark_f)
df_charles_f = NER_replace(df_charles_f)
df_fitzgerald_f.Text[0]
for ent in df_fitzgerald_f.spacy_words[0]:
if ent.label_ in ['PERSON','FAC','GPE','LOC','ORG']:
print(ent.text, ent.start_char, ent.end_char,
ent.label_)
df_fitzgerald_f.to_csv("../Dataset_v3/deidentified/Fitzgerald_256_di.csv")
df_mark_f.to_csv("../Dataset_v3/deidentified/Mark_256_di.csv")
df_charles_f.to_csv("../Dataset_v3/deidentified/Charles_256_di.csv")
| Preprocessing/.ipynb_checkpoints/SpacyCleaning-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from neuronet import ActivePerceptron, Perceptron, PseudoRandomNeuroNet
from deductor_parser import DeductorParser
# from PRNN import NeuroNet
from random import random
# -
parser = DeductorParser('test.ded')
# +
# parser.getdocumentsxml().keys()
# -
parser.setdocument('Текстовый файл (C:\\Users\\Aleksandr\\Desktop\\PyOptimizer\\test.csv)')
# +
# parser.getneuronetsxml().keys()
# +
parser.setneuronet('Нейросеть [2 x 10 x 10 x 1]', parse=True)
nw_active = ActivePerceptron()
nw_active.loader(parser.neurodata)
# -
nw = Perceptron()
nw.loader(parser.neurodata)
nw_active(10, 10)
nw(10, 10)
# %timeit nw_active(random() * 20 - 10, random() * 20 - 10)
# %timeit nw(random() * 20 - 10, random() * 20 - 10)
nrs = NeuroNet(2, 1, 20, 10)
nrs.setinnormal([(-10, 10, -1, 1), (-10, 10, -1, 1)])
nrs.setdiscretout([False] * 1)
nrs.setoutnormal([(-20, 120, 0, 1)] * 1)
nrs.setdelta(0.01)
nrs.setcountlearn(10)
nrs.is_stabilization = False
nrs.reinit()
nrs.setspeed(0.5)
nrs.gety([random() * 20 - 10, random() * 20 - 10])
# %timeit nrs.gety([random() * 20 - 10, random() * 20 - 10])
nrs_stab = NeuroNet(2, 1, 20, 10)
nrs_stab.setinnormal([(-10, 10, -1, 1), (-10, 10, -1, 1)])
nrs_stab.setdiscretout([False] * 1)
nrs_stab.setoutnormal([(-20, 120, 0, 1)] * 1)
nrs_stab.setdelta(0.01)
nrs.setcountlearn(10)
nrs_stab.is_stabilization = True
nrs_stab.reinit()
nrs_stab.setspeed(0.5)
nrs_stab.gety([random() * 20 - 10, random() * 20 - 10])
# %timeit nrs_stab.gety([random() * 20 - 10, random() * 20 - 10])
# %timeit x = random() * 20 - 10; y = random() * 20 - 10; z = x ** 2 + y ** 2 + x + y; nrs_stab.learn([x, y, z])
# %timeit x = random() * 20 - 10; y = random() * 20 - 10; z = x ** 2 + y ** 2 + x + y; nw.learn_bprop([[[x, y], [z]]])
import hashlib
x = '15'
hashlib.md5(b'x').hexdigest()
# %timeit nw_new.gety([random() * 20 - 10, random() * 20 - 10])
# %timeit x = random() * 20 - 10; y = random() * 20 - 10; z = x ** 2 + y ** 2 + x + y; nw_new.learn_bprop([[[x, y], [z]]])
# +
from neuronet import ActivePerceptron, Perceptron, PseudoRandomNeuroNet
from deductor_parser import DeductorParser
# from PRNN import NeuroNet
from random import random
# -
parser = DeductorParser('Тест.ded')
parser.documents
parser.setdocument('Текстовый файл (C:\\Users\\Aleksandr\\Desktop\\PyOptimizer\\test.csv)')
parser.neuronets
parser.setneuronet('Нейросеть [4 x 10 x 4]', parse=True)
parser.neurodata
nw_active = ActivePerceptron()
nw = Perceptron()
nw.loader(parser.neurodata)
nw([0] * 4)
# %timeit nw([0] * 4)
nw_active.loader(nw.neurodata)
nw_active([0] * 4)
# %timeit nw_active([0] * 4)
| test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Plotting Figure 2
# This script is used to show multi-model ensemble mean change in UHWs
import xarray as xr
import datetime
import pandas as pd
import numpy as np
#from mpl_toolkits.basemap import Basemap
import matplotlib.pyplot as plt
import matplotlib.ticker as mticker
import time
import gc
import util
import cartopy
import cartopy.crs as ccrs
import cartopy.feature as cfeature
from cartopy.mpl.gridliner import LONGITUDE_FORMATTER, LATITUDE_FORMATTER
from cartopy.mpl.ticker import LongitudeFormatter, LatitudeFormatter
# import cartopy.io.shapereader as shapereader
# ## Step 1: Load 2006 and 2061 data
# +
# Load CMIP
CMIP={}
path="../data/uhws/UHWs_CMIP/"
CMIP["diff_raw"]={}
CMIP["diff_stat"]={}
for year in ["2006","2061"]:
CMIP[year]={}
for vari in ["frequency","totaldays","intensity"]:
CMIP[year][vari]=pd.read_csv(path+year+"_"+vari+".csv").set_index(["lat","lon"])
CMIP[year]["duration"]=CMIP[year]["totaldays"]/CMIP[year]["frequency"]
for vari in ["frequency","totaldays","intensity","duration"]:
CMIP["diff_raw"][vari] = CMIP["2061"][vari]-CMIP["2006"][vari]
CMIP["diff_stat"][vari+"_diff_mean"] = CMIP["diff_raw"][vari].mean(axis=1)
CMIP["diff_stat"][vari+"_diff_std"] = CMIP["diff_raw"][vari].std(axis=1)
#CMIP["diff_stat"][vari+"_diff_0.025"] = CMIP["diff_raw"][vari].quantile(0.025,axis=1)
CMIP["diff_stat"][vari+"_diff_SNR"] = CMIP["diff_stat"][vari+"_diff_mean"].abs()/CMIP["diff_stat"][vari+"_diff_std"]
"""
for keys in CMIP["diff_stat"]:
print(keys)
print(CMIP["diff_stat"][keys].describe())
"""
# Load CESM
CESM={}
path="../data/uhws/UHWs_CESM/"
CESM["diff_raw"]={}
CESM["diff_stat"]={}
for year in ["2006","2061"]:
CESM[year]={}
for vari in ["frequency","totaldays","intensity"]:
CESM[year][vari]=pd.read_csv(path+year+"_"+vari+".csv").set_index(["lat","lon"])
CESM[year]["duration"]=CESM[year]["totaldays"]/CESM[year]["frequency"]
for vari in ["frequency","totaldays","intensity","duration"]:
CESM["diff_raw"][vari] = CESM["2061"][vari]-CESM["2006"][vari]
CESM["diff_stat"][vari+"_diff_mean"] = CESM["diff_raw"][vari].mean(axis=1)
CESM["diff_stat"][vari+"_diff_std"] = CESM["diff_raw"][vari].std(axis=1)
#CESM["diff_stat"][vari+"_diff_0.025"] = CESM["diff_raw"][vari].quantile(0.025,axis=1)
CESM["diff_stat"][vari+"_diff_SNR"] = CESM["diff_stat"][vari+"_diff_mean"].abs()/CESM["diff_stat"][vari+"_diff_std"]
"""
for keys in CESM["diff_stat"]:
print(keys)
print(CESM["diff_stat"][keys].describe())
"""
# merge CMIP with CESM member 002
merge_ens={}
for vari in ["frequency","totaldays","intensity","duration"]:
merge_ens[vari]=pd.concat([CMIP["diff_raw"][vari].reset_index(),
CESM["diff_raw"][vari]["002_max"].reset_index()["002_max"]],
axis=1).set_index(["lat","lon"])
merge_ens[vari+"_diff_mean"]=merge_ens[vari].mean(axis=1)
merge_ens[vari+"_diff_std"]=merge_ens[vari].std(axis=1)
merge_ens[vari+"_diff_SNR"]=merge_ens[vari+"_diff_mean"].abs()\
/merge_ens[vari+"_diff_std"]
#list(merge_ens)
# -
# ## Step 2: Plotting
# +
def setup_globe(ax):
ax.set_extent([-180,180,-60,75],crs=ccrs.PlateCarree())
ax.coastlines(zorder=5)
# ax.add_feature(cartopy.feature.LAND, facecolor='lightgray',zorder=0)
ax.add_feature(cartopy.feature.LAKES, facecolor='none',edgecolor='black',
linewidth=0.5,zorder=5)
ax.add_feature(cartopy.feature.BORDERS,
facecolor='none',
edgecolor='black',
linewidth=0.5,zorder=5)
def setup_colorbar_text(ax,p,title_text,var_text,cbar_text):
ax.set_xlabel("")
ax.set_ylabel("")
cbar = plt.colorbar(p, ax=ax,
orientation="vertical",
fraction=0.20,
shrink=0.80,
pad=0.02,
aspect=30,
extend="both")
cbar.ax.set_title(cbar_text)
g = ax.gridlines(color='grey', linestyle='--', draw_labels=False,zorder=4)
g.xlocator = mticker.FixedLocator([-90, 0, 90])
lon_formatter = LongitudeFormatter()#zero_direction_label=True)
lat_formatter = LatitudeFormatter()
ax.xaxis.set_major_formatter(lon_formatter)
ax.yaxis.set_major_formatter(lat_formatter)
# props = dict(boxstyle='round', facecolor='lightcyan')
# # place a text box in upper left in axes coords
# ax.text(0.03, 0.20, var_text, transform=ax.transAxes, fontsize=12,
# verticalalignment='top', bbox=props,zorder=6)
ax.set_title(title_text, fontweight="bold", loc="left")
def plot_scatter(ax,df,SNR,vmin,vmax,cmap):
df_temp = df.copy()
df_temp["lon_new"] = np.where(df_temp['lon'] <= 180,
df_temp['lon'],
df_temp['lon']-360)
p = ax.scatter(df_temp["lon"],df_temp["lat"],c=df_temp[0],
s=0.5,
vmin=vmin,
vmax=vmax,
cmap=cmap,
zorder=3,rasterized=True)
df_SNR = SNR[SNR[0]>2.0].copy()
df_SNR["lon_new"] = np.where(df_SNR['lon'] <= 180,
df_SNR['lon'],
df_SNR['lon']-360)
ax.scatter(df_SNR["lon"],df_SNR["lat"],
s=0.05,c="k",marker=".",
zorder=3,rasterized=True)
return p
def plot_map(df,SNR,i,vmin,vmax,cmap,title_text,var_text,cbar_text):
ax = plt.subplot(2,2,i+1,projection=ccrs.PlateCarree())
setup_globe(ax)
if i==0:
ax.set_yticks(np.array([-60,-30,0,30,60]), crs=ccrs.PlateCarree())
elif i==1:
pass
elif i==2:
ax.set_xticks(np.linspace(-90, 90, 3), crs=ccrs.PlateCarree())
ax.set_yticks(np.array([-60,-30,0,30,60]), crs=ccrs.PlateCarree())
elif i==3:
ax.set_xticks(np.linspace(-90, 90, 3), crs=ccrs.PlateCarree())
p = plot_scatter(ax,df,SNR,vmin,vmax,cmap)
setup_colorbar_text(ax,p,title_text,var_text,cbar_text)
# +
fig = plt.figure(figsize=(12,5))
cmap="RdYlBu_r"
#cmap = "rainbow"
var_list=["intensity","frequency","duration","totaldays"]
max_val={"frequency":10,
"totaldays":90,
"intensity":3,
"duration":7}
min_val={"frequency":0,
"totaldays":0,
"intensity":-0.35,
"duration":-0.3}
# var_text_dict={"frequency":"Frequency\n(events/year)",
# "totaldays":"Total days\n(days/year)",
# "intensity":"Intensity\n(K)",
# "duration":"Duration\n(days/event)"}
var_text_dict={"frequency":"Frequency",
"totaldays":"Total days",
"intensity":"Intensity",
"duration":"Duration"}
title_text_dict={"frequency":"b",
"totaldays":"d",
"intensity":"a",
"duration":"c"}
cbar_text_dict={"frequency":"Frequency\n[events/year]",
"totaldays":"Total days\n[days/year]",
"intensity":"Intensity\n[K]",
"duration":"Duration\n[days/event]"}
for i in range(4):
val = var_list[i]
print(val)
df = merge_ens[val+"_diff_mean"]
SNR = merge_ens[val+"_diff_SNR"].reset_index()[merge_ens["intensity_diff_mean"].reset_index()[0]>1.5]
display(df.describe())
plot_map(df.reset_index(),SNR,i,min_val[val], max_val[val],cmap,
title_text_dict[val],var_text_dict[val],cbar_text_dict[val])
print("##########################################")
plt.tight_layout()
#plt.subplots_adjust(wspace=0, hspace=0)
plt.savefig("../figures/uhws.pdf",dpi=188)
plt.show()
| 5_event_analysis/fig2_uhws.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Inheritance with the Gaussian Class
#
# To give another example of inheritance, take a look at the code in this Jupyter notebook. The Gaussian distribution code is refactored into a generic Distribution class and a Gaussian distribution class. Read through the code in this Jupyter notebook to see how the code works.
#
# The Distribution class takes care of the initialization and the read_data_file method. Then the rest of the Gaussian code is in the Gaussian class. You'll later use this Distribution class in an exercise at the end of the lesson.
#
# Run the code in each cell of this Jupyter notebook. This is a code demonstration, so you do not need to write any code.
class Distribution:
def __init__(self, mu=0, sigma=1):
""" Generic distribution class for calculating and
visualizing a probability distribution.
Attributes:
mean (float) representing the mean value of the distribution
stdev (float) representing the standard deviation of the distribution
data_list (list of floats) a list of floats extracted from the data file
"""
self.mean = mu
self.stdev = sigma
self.data = []
def read_data_file(self, file_name):
"""Function to read in data from a txt file. The txt file should have
one number (float) per line. The numbers are stored in the data attribute.
Args:
file_name (string): name of a file to read from
Returns:
None
"""
with open(file_name) as file:
data_list = []
line = file.readline()
while line:
data_list.append(int(line))
line = file.readline()
file.close()
self.data = data_list
# +
import math
import matplotlib.pyplot as plt
class Gaussian(Distribution):
""" Gaussian distribution class for calculating and
visualizing a Gaussian distribution.
Attributes:
mean (float) representing the mean value of the distribution
stdev (float) representing the standard deviation of the distribution
data_list (list of floats) a list of floats extracted from the data file
"""
def __init__(self, mu=0, sigma=1):
Distribution.__init__(self, mu, sigma)
def calculate_mean(self):
"""Function to calculate the mean of the data set.
Args:
None
Returns:
float: mean of the data set
"""
avg = 1.0 * sum(self.data) / len(self.data)
self.mean = avg
return self.mean
def calculate_stdev(self, sample=True):
"""Function to calculate the standard deviation of the data set.
Args:
sample (bool): whether the data represents a sample or population
Returns:
float: standard deviation of the data set
"""
if sample:
n = len(self.data) - 1
else:
n = len(self.data)
mean = self.calculate_mean()
sigma = 0
for d in self.data:
sigma += (d - mean) ** 2
sigma = math.sqrt(sigma / n)
self.stdev = sigma
return self.stdev
def plot_histogram(self):
"""Function to output a histogram of the instance variable data using
matplotlib pyplot library.
Args:
None
Returns:
None
"""
plt.hist(self.data)
plt.title('Histogram of Data')
plt.xlabel('data')
plt.ylabel('count')
def pdf(self, x):
"""Probability density function calculator for the gaussian distribution.
Args:
x (float): point for calculating the probability density function
Returns:
float: probability density function output
"""
return (1.0 / (self.stdev * math.sqrt(2*math.pi))) * math.exp(-0.5*((x - self.mean) / self.stdev) ** 2)
def plot_histogram_pdf(self, n_spaces = 50):
"""Function to plot the normalized histogram of the data and a plot of the
probability density function along the same range
Args:
n_spaces (int): number of data points
Returns:
list: x values for the pdf plot
list: y values for the pdf plot
"""
mu = self.mean
sigma = self.stdev
min_range = min(self.data)
max_range = max(self.data)
# calculates the interval between x values
interval = 1.0 * (max_range - min_range) / n_spaces
x = []
y = []
# calculate the x values to visualize
for i in range(n_spaces):
tmp = min_range + interval*i
x.append(tmp)
y.append(self.pdf(tmp))
# make the plots
fig, axes = plt.subplots(2,sharex=True)
fig.subplots_adjust(hspace=.5)
axes[0].hist(self.data, density=True)
axes[0].set_title('Normed Histogram of Data')
axes[0].set_ylabel('Density')
axes[1].plot(x, y)
axes[1].set_title('Normal Distribution for \n Sample Mean and Sample Standard Deviation')
axes[0].set_ylabel('Density')
plt.show()
return x, y
def __add__(self, other):
"""Function to add together two Gaussian distributions
Args:
other (Gaussian): Gaussian instance
Returns:
Gaussian: Gaussian distribution
"""
result = Gaussian()
result.mean = self.mean + other.mean
result.stdev = math.sqrt(self.stdev ** 2 + other.stdev ** 2)
return result
def __repr__(self):
"""Function to output the characteristics of the Gaussian instance
Args:
None
Returns:
string: characteristics of the Gaussian
"""
return "mean {}, standard deviation {}".format(self.mean, self.stdev)
# +
# initialize two gaussian distributions
gaussian_one = Gaussian(25, 3)
gaussian_two = Gaussian(30, 2)
# initialize a third gaussian distribution reading in a data efile
gaussian_three = Gaussian()
gaussian_three.read_data_file('numbers.txt')
gaussian_three.calculate_mean()
gaussian_three.calculate_stdev()
# +
# print out the mean and standard deviations
print(gaussian_one.mean)
print(gaussian_two.mean)
print(gaussian_one.stdev)
print(gaussian_two.stdev)
print(gaussian_three.mean)
print(gaussian_three.stdev)
# -
# plot histogram of gaussian three
gaussian_three.plot_histogram_pdf()
# add gaussian_one and gaussian_two together
gaussian_one + gaussian_two
| inheritance_probability_distribution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sklearn, XGBoost
# ## sklearn.ensemble.RandomForestClassifier
# +
from sklearn import ensemble , cross_validation, learning_curve, metrics
import numpy as np
import pandas as pd
import xgboost as xgb
# -
# %pylab inline
# ### Данные
# Задача на kaggle: https://www.kaggle.com/c/bioresponse
#
# Данные: https://www.kaggle.com/c/bioresponse/data
#
# По данным характеристикам молекулы требуется определить, будет ли дан биологический ответ (biological response).
#
# Признаки нормализаваны.
#
# Для демонстрации используется обучающая выборка из исходных данных train.csv, файл с данными прилагается.
bioresponce = pd.read_csv('bioresponse.csv', header=0, sep=',')
bioresponce.head()
bioresponce_target = bioresponce.Activity.values
bioresponce_data = bioresponce.iloc[:, 1:]
# ### Модель RandomForestClassifier
# #### Зависимость качества от количесвта деревьев
n_trees = [1] + range(10, 55, 5)
# %%time
scoring = []
for n_tree in n_trees:
estimator = ensemble.RandomForestClassifier(n_estimators = n_tree, min_samples_split=5, random_state=1)
score = cross_validation.cross_val_score(estimator, bioresponce_data, bioresponce_target,
scoring = 'accuracy', cv = 3)
scoring.append(score)
scoring = np.asmatrix(scoring)
scoring
pylab.plot(n_trees, scoring.mean(axis = 1), marker='.', label='RandomForest')
pylab.grid(True)
pylab.xlabel('n_trees')
pylab.ylabel('score')
pylab.title('Accuracy score')
pylab.legend(loc='lower right')
# #### Кривые обучения для деревьев большей глубины
# %%time
xgb_scoring = []
for n_tree in n_trees:
estimator = xgb.XGBClassifier(learning_rate=0.1, max_depth=5, n_estimators=n_tree, min_child_weight=3)
score = cross_validation.cross_val_score(estimator, bioresponce_data, bioresponce_target,
scoring = 'accuracy', cv = 3)
xgb_scoring.append(score)
xgb_scoring = np.asmatrix(xgb_scoring)
xgb_scoring
pylab.plot(n_trees, scoring.mean(axis = 1), marker='.', label='RandomForest')
pylab.plot(n_trees, xgb_scoring.mean(axis = 1), marker='.', label='XGBoost')
pylab.grid(True)
pylab.xlabel('n_trees')
pylab.ylabel('score')
pylab.title('Accuracy score')
pylab.legend(loc='lower right')
# #### **Если Вас заинтересовал xgboost:**
# python api: http://xgboost.readthedocs.org/en/latest/python/python_api.html
#
# установка: http://xgboost.readthedocs.io/en/latest/build.html
| Yandex data science/2/Week 4/sklearn.rf_vs_gb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="ALXIo7pg34kU" executionInfo={"status": "ok", "timestamp": 1632266951021, "user_tz": -420, "elapsed": 3644, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "15709245763635183075"}} outputId="f7459ecd-de88-4e16-9919-60b5b063e6f8"
# Download all packages
from google.colab import files, output, drive
# !rm -rf darknet
# !git clone https://github.com/AlexeyAB/darknet
output.clear()
print("All packages downloaded!")
# + id="32Fb_5n4s9FP" executionInfo={"status": "ok", "timestamp": 1632266951024, "user_tz": -420, "elapsed": 17, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "15709245763635183075"}}
# Load all packages
import os
import shutil
# + colab={"base_uri": "https://localhost:8080/"} id="X5yVsh7rVWAY" executionInfo={"status": "ok", "timestamp": 1632267055639, "user_tz": -420, "elapsed": 104632, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "15709245763635183075"}} outputId="c712669b-acee-4dd2-e43a-cb7b49736b23"
# Set up Yolov4
# %cd darknet
# !sed -i 's/OPENCV=0/OPENCV=1/' Makefile
# !sed -i 's/GPU=0/GPU=1/' Makefile
# !sed -i 's/CUDNN=0/CUDNN=1/' Makefile
# !sed -i 's/CUDNN_HALF=0/CUDNN_HALF=1/' Makefile
# !make
# !rm yolov4.conv.137
# !wget https://github.com/AlexeyAB/darknet/releases/download/darknet_yolo_v3_optimal/yolov4.conv.137
output.clear()
print("Yolov4 Ready!")
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": "OK"}}, "base_uri": "https://localhost:8080/", "height": 35} id="2JGXKGhpYKp1" executionInfo={"status": "ok", "timestamp": 1632267081400, "user_tz": -420, "elapsed": 25797, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "15709245763635183075"}} outputId="43d78bff-8419-42d2-cdcc-df05be451791"
# Connect colab to kaggle
# %cd ../
# !rm kaggle.json
files.upload()
# !mkdir -p ~/.kaggle
# !cp kaggle.json ~/.kaggle/
# !ls ~/.kaggle
# !chmod 600 /root/.kaggle/kaggle.json
output.clear()
print("Colab connected to Kaggle!")
# + colab={"base_uri": "https://localhost:8080/"} id="fbeS81h_hKvW" executionInfo={"status": "ok", "timestamp": 1632267361331, "user_tz": -420, "elapsed": 618, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "15709245763635183075"}} outputId="09f8a3bf-4bc4-4aa1-f96e-8045354e15fe"
# Mount colab to google drive
drive.mount('/content/gdrive')
output.clear()
print("Colab connected to Drive!")
# + colab={"base_uri": "https://localhost:8080/"} id="cVKCadIrYc3x" executionInfo={"status": "ok", "timestamp": 1632267164193, "user_tz": -420, "elapsed": 53935, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "15709245763635183075"}} outputId="333a2efa-65ad-4787-9940-caef3be9a752"
# Setup datasets for Weapons
# %cd darknet
# !rm weapons-in-images-segmented-videos.zip
# !kaggle datasets download -d jubaerad/weapons-in-images-segmented-videos
# !rm -rf Weapons-in-Images
# !unzip weapons-in-images-segmented-videos.zip
# !rm weapons-in-images-segmented-videos.zip
# !rm -rf "Weapon in Images (Segmented Video)"
# !rm -r data/*
# !mkdir -p data/{obj,test}
directory_weapons = "./Weapons-in-Images/Weapons-in-Images"
all_data_weapons = os.listdir(directory_weapons)
for data in all_data_weapons:
real_data, extension_data = data.split(".")
if extension_data == "jpg" and f"{real_data}.txt" not in all_data_weapons:
os.remove(f"./Weapons-in-Images/Weapons-in-Images/{data}")
all_data_weapons = os.listdir(directory_weapons)
all_data_weapons.sort()
train_composition = int(((len(all_data_weapons) / 2) * 0.8) * 2)
test_composition = len(all_data_weapons) - train_composition
count = 0
for data in all_data_weapons:
if count < test_composition:
shutil.move(os.path.join(directory_weapons, data), "data/test/")
elif count < train_composition:
shutil.move(os.path.join(directory_weapons, data), "data/obj/")
count += 1
# !rm -rf Weapons-in-Images
output.clear()
print("Datasets ready!")
# + colab={"base_uri": "https://localhost:8080/"} id="i_8GK_lbM9lG" executionInfo={"status": "ok", "timestamp": 1632267164197, "user_tz": -420, "elapsed": 65, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "15709245763635183075"}} outputId="c250c6b1-d18e-45cb-b81a-57f2fe63e179"
# Create train.txt
image_files = []
os.chdir(os.path.join("data", "obj"))
for filename in os.listdir(os.getcwd()):
if filename.endswith(".jpg"):
image_files.append("data/obj/" + filename)
os.chdir("..")
with open("train.txt", "w") as outfile:
for image in image_files:
outfile.write(image)
outfile.write("\n")
outfile.close()
os.chdir("..")
output.clear()
print("train.txt created!")
# + colab={"base_uri": "https://localhost:8080/"} id="mirIHYJCNmZv" executionInfo={"status": "ok", "timestamp": 1632267164200, "user_tz": -420, "elapsed": 61, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "15709245763635183075"}} outputId="2d07c07c-d965-4de7-d1f6-02dac1ddb6ac"
# Create test.txt
image_files = []
os.chdir(os.path.join("data", "test"))
for filename in os.listdir(os.getcwd()):
if filename.endswith(".jpg"):
image_files.append("data/test/" + filename)
os.chdir("..")
with open("test.txt", "w") as outfile:
for image in image_files:
outfile.write(image)
outfile.write("\n")
outfile.close()
os.chdir("..")
output.clear()
print("test.txt created!")
# + colab={"base_uri": "https://localhost:8080/", "height": 227} id="VgmW1Jt5kjip" executionInfo={"status": "error", "timestamp": 1632267164208, "user_tz": -420, "elapsed": 62, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "15709245763635183075"}} outputId="a2a0a52b-54a3-4e48-86b9-5800ab23d146"
# Set yolov4 config and backup dataset
!>data/obj.data
!>data/obj.names
output.clear()
print("Yolov4 configured and backup dataset!")
"""
- MAKE SURE SET CONFIG FOR CUSTOM yolov4-custom.cfg
- MAKE SURE SET YOUR DATASET obj.data AND obj.names
"""
assert False
# + colab={"base_uri": "https://localhost:8080/"} id="oJ8Bh2ddOOie" executionInfo={"status": "ok", "timestamp": 1632269899106, "user_tz": -420, "elapsed": 2466632, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "15709245763635183075"}} outputId="1f1bebc1-b5bd-4e64-896c-fc29ce7d71b4"
# Train model
# !./darknet detector train \
# data/obj.data \
# cfg/yolov4-custom.cfg \
# ./yolov4.conv.137 \
# -dont_show
output.clear()
print("Train done!")
# + id="dGjPDcyyPfGF" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632270370478, "user_tz": -420, "elapsed": 147464, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "15709245763635183075"}} outputId="f00937ec-0bf6-4d51-a97d-79fbfaa6bbbc"
# Checking Mean Average Precision Each Weights
print("============================")
print("LAST WEIGHTS")
# !./darknet detector map \
# data/obj.data \
# cfg/yolov4-custom.cfg \
# /content/gdrive/MyDrive/"Week 3"/Task/google_colab_data/backup/yolov4-custom_last.weights
print("============================")
print("1000 MAX BATCHERS")
# !./darknet detector map \
# data/obj.data \
# cfg/yolov4-custom.cfg \
# /content/gdrive/MyDrive/"Week 3"/Task/google_colab_data/backup/yolov4-custom_1000.weights
# + id="LZy3c0DexRjX" executionInfo={"status": "aborted", "timestamp": 1632267164207, "user_tz": -420, "elapsed": 49, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "15709245763635183075"}}
| jupyter_notebook/train_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="9vC5WqnedfxY"
# You are hired by a venture capitalist to predict the profit of a startup. Fo that you have to deal with a dataset which contains the details of 50 startups and predicts the profit of a new Startup based on certain features. Based on your decision and prediction, whether one should invest in a particular startup or not.
#
# Dataset contains the following fields:
# R&D Spend - Total amount of money spent on Research and Development Administration - Total amount of money spent on Administration
# Marketing Spend - Total amount of money spent on Markeing
# State - The state where the startup operates
# Profit - Profit earned by startup
#
# You have to perform following task before applying machine learning algorithms:
# 1) Handle missing values
# 2) Prepare data for training and testing
# 3) Apply Decision Tree algorithm to train the model
# 4) Apply Random Forest Regressor algorithm to train the model
# 5) Compare the accuracy with Linear Regression too.
# + id="6lR8tV62dfxY"
#import required libraries
import numpy as np
import pandas as pd
# + id="6Dpu0nXadfxY"
#loading data file
data = pd.read_csv('50_Startups.csv')
# + id="oiebWqQ_dfxY" outputId="89749d5d-302c-4701-91b3-e9bd310e7e2b"
#displays number of columns and records/rows in dataset
#check if there is any missing data or not
data.info()
# + id="Lo44s7DndfxZ" outputId="c5767812-1bf4-414b-f01b-980851504143"
data.describe()
# + id="G5SysJQPdfxa"
features = data.iloc[:,:-1].values
label = data.iloc[:,[-1]].values
# + id="rHpEErZ9dfxa" outputId="ae4c1d2d-d773-4703-9135-7fc41d89fa7d"
features
# + id="b_wVlq18dfxa" outputId="0f3024bc-bcfb-44b1-de59-8615d6f5ffc0"
#convert the categorical features to numerical features as
#sklearn works only with numpy array
#Instead of label enconding and then onehotencoding,
#newer version directly works with onehotencoding using ColumnTransformer
from sklearn.preprocessing import OneHotEncoder
from sklearn.compose import ColumnTransformer
transformer = ColumnTransformer(transformers=[
("OneHot", # Just a name
OneHotEncoder(), # The transformer class
[3] # The column(s) to be applied on.
)
],
remainder='passthrough' # donot apply anything to the remaining columns
)
features = transformer.fit_transform(features.tolist())
features
# + id="68pre3y0dfxa"
#converting an object to normal array
features = features.astype(float)
# + id="YPYY8Shidfxa" outputId="d6a212b0-befa-4696-9cff-e492ede3011a"
features
# + id="HS085G0Udfxa"
#sampling the dataset
#normally 20% dataset is used for testing and 80% is used for training --> test_size=0.2 means 20%
#Training set will be used to train the model
#Create Training and Testing sets
# Testing set will be used to test our model
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test = train_test_split(features,
label,
test_size=0.2,
random_state=1)
# + id="tyilNWsOdfxb" outputId="787102a5-98ec-4ac3-e874-abaa09cf93ad"
#Create our model using Linear Regression
from sklearn.tree import DecisionTreeRegressor
DTR = DecisionTreeRegressor(max_depth=3)
DTR.fit(X_train,y_train)
# + id="5h2aNur1dfxb" outputId="fe951170-1f9d-4e30-eab7-584377de5113"
#checking score of training as well as testing
print(DTR.score(X_train,y_train))
print(DTR.score(X_test,y_test))
# + [markdown] id="AcjPwerwdfxb"
# Model is not generalized yet. You may check for other values of max_depth.
# max_depth generally is odd values. You may check for 5,7,9... And check for training and testing score and choose the model with generalized score.
# + id="E0RRKdn9dfxb" outputId="20049b71-941d-4137-a9c2-941775188d76"
from sklearn.ensemble import RandomForestRegressor
RF=RandomForestRegressor(n_estimators=3)
RF.fit(X_train,y_train.ravel())
print(RF.score(X_train,y_train))
print(RF.score(X_test,y_test))
# + id="ZuaMtn7Idfxb" outputId="12930ce0-cfbf-4c65-b537-754d6d5f19a4"
from sklearn.ensemble import RandomForestRegressor
for i in range(4,10):
RF=RandomForestRegressor(n_estimators=3)
RF.fit(X_train,y_train.ravel())
print("n_estimator = ",i)
print("Training Score =",RF.score(X_train,y_train))
print("TEsting Score = ",RF.score(X_test,y_test))
# + [markdown] id="aQwKRpJ4dfxb"
# You can conclude that with n_estimator 8, generalized model canbe derived where testing score is more than training score.
# -
label2 = data.iloc[-1:-1].values
label2
label = data.iloc[:,[-1]].values
label
| Greycampus/DecisionTree_RandomForest_Regression (1)-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="dZBAW1TiUikU"
# # !pip install -q condacolab -q
# import condacolab
# condacolab.install()
# # !conda install geopandas
# + id="ZXxqPj6y9ogi"
# # !pip install awscli
# # !pip install cloudpathlib
# # !pip install geopandas
# # !pip install rasterio
# # !pip install pyhdf
# # !pip install cloudpathlib[s3]
# # !pip install rtree
# # !pip install pqdm
# + id="89c1PKIjCD_r"
import os
import re
import random
import pickle
from pathlib import Path
os.chdir('/content/drive/MyDrive/datadriven/airathon')
DATA_PATH = Path.cwd() / 'data'
RAW = DATA_PATH / 'raw'
PROCESSED = DATA_PATH / 'processed3'
from utils0 import *
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
from osgeo import gdal
import geopandas as gpd
from pyhdf.SD import SD, SDC, SDS
from typing import Dict, List, Union
import pyproj
from pyproj import CRS, Proj
from pqdm.processes import pqdm
# import multiprocessing
# n_cpus = multiprocessing.cpu_count()
mpl.rcParams['figure.dpi'] = 100
# + id="9WH5hXF8Hf73"
satellite = pd.read_csv(RAW / 'pm25_satellite_metadata.csv', parse_dates=['time_start', 'time_end'], index_col=0)
grid = pd.read_csv(RAW / 'grid_metadata.csv', index_col=0)
train_labels = pd.read_csv(RAW / 'train_labels.csv', parse_dates=["datetime"])
train_labels.rename(columns={'value': 'pm25'}, inplace=True)
submission = pd.read_csv(RAW / 'submission_format.csv', parse_dates=['datetime'])
submission.value = submission.value.replace(0, np.NaN)
# + colab={"base_uri": "https://localhost:8080/", "height": 412, "referenced_widgets": ["9ff7d1e808694472b39ea317a48e611e", "b4e9b87e6ecb4ea28f3a53af608a519d", "cedb5f87254e4dafba296e48c624f844", "d2295ad0ba624658bffa706c7e342a79", "0dbe3b87234c4f8ab4550248325c35a9", "d3fc7fbc269943d580d21dc85be74798", "3a196efc40ca47ca99355427253fc131", "8a5289d4e54d4aaea0319ccf1651325d", "<KEY>", "30ce826003f345dfb9ca8814a9fb56e6", "<KEY>", "29a8b88ceef94b42b1994b1bb88f626f", "2099e9c299ea4dff86547974d6b05ec4", "721d4db863914a49b06f94459a242d7c", "4de1530ebd7b4835b91af48c84614744", "<KEY>", "a05acf45526f4c649a149df60e43e279", "<KEY>", "2b78a6d405c249c588563703c623850c", "<KEY>", "<KEY>", "b582d7b58d544ad5be3ee9ae3a0ce103", "c5d7afbb20e241cab97b46636766dac0", "5fa8fe4bb9e24f8c9261120aec833ad8", "37ca3f0a04cb4e4f9fd0d6c6edcca9a4", "fc44e9e793474243ad38581a4f061ef9", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "4c7814c240b949d992f91f5a4d23e3e4", "<KEY>", "<KEY>", "7fa34089693544e7b6ba2f1fdf7f1065", "<KEY>", "945e8d24cbeb459ea3961eff66191809", "<KEY>", "01542d11d8e4442db25a2b01269fce8b", "<KEY>", "<KEY>", "9d665c8adcdc4e959c49f229ed71216d", "<KEY>", "2b20849d554a4d7dbe35db845d76c5dd", "6e7a24023390444f84341ad6fbe08d5f", "a90927f981fe4bd5be96d1c97a5d9fd7", "ff0734755e494f048b83c566af0b6224", "18836e661ca8405faac7c95120c4acb2", "<KEY>", "<KEY>", "<KEY>", "5dc12518512a4443be57fddee5e26767", "<KEY>", "<KEY>", "<KEY>", "be7d754760f644f2930a118818e6888f", "d695a04407484a9685dd1b6494672db0", "3216ce1f707043e6ae913b873d2e16e7", "<KEY>", "<KEY>", "3a732c2f15eb4a06983f4dde21e50fe5", "<KEY>", "12cd259d0eb541cdbceee082c4e70926", "<KEY>", "<KEY>", "<KEY>", "0e237b2193e54baa9e79f7d061541d8e", "<KEY>", "d41f101334074629a2d1891fe7e1624c", "<KEY>", "4a3c5341e9154095aba45ef9799cefaf", "<KEY>", "a2d73966d0c040f4914a9f01b168a9b4", "182016fedabd4dbe833b6745c3dbd616", "e232af979f3a4e9ba88889ce9c86b24a", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "9c50ad1fcc034eb88c4ae7793b3a2b94", "662b9572d2ef40129cd74e292e5b067e", "<KEY>", "3afa082349ba41689b2691b556412545", "2188c10ea49942d28785a68951273868", "<KEY>", "<KEY>", "<KEY>", "6fdd964f097840f5bc5ad6871ef0ea2f", "<KEY>", "569f03537bc345c7a2d137ab85e8f52d", "e6afceeeebe8434aa0bf98bd75e1b8b5", "063ed2ce87634a068d596a593e92308f", "<KEY>", "9e46065eb3904b51a05bba51604cc8cc", "<KEY>", "f813482c2f4f404a9347289dd0fe7c8a", "92957e2115ed4db6b4fe8f7e71ecf608", "<KEY>", "ed4cb51028054ae3b45617c80558695e", "525a6988c2434e60bb286934ec374ab2"]} id="Q9aptUaULby9" outputId="c012c9fe-c587-454c-8c67-403ab15df151"
train_maiac = satellite[(satellite["product"] == "maiac") & (satellite["split"] == "train")].copy()
datasets = ['Optical_Depth_047', 'Optical_Depth_055', 'AOD_Uncertainty', 'Column_WV']
train_dict = {}
locations = [['la', 'Los Angeles (SoCAB)'], ['dl', 'Delhi'], ['tpe', 'Taipei']]
for loc in locations:
print(f'Processing: {loc[1]}\n')
train_dict[loc[0]] = {}
satellite_subset = train_maiac[train_maiac.location == loc[0]].copy()
file_paths = list(satellite_subset.us_url)
grid_subset = grid[grid.location == loc[1]].copy()
assert grid_subset.index.isin(train_labels.grid_id).all()
polys = gpd.GeoSeries.from_wkt(grid_subset.wkt, crs=wgs84_crs)
polys.name = "geometry"
polys_gDF = gpd.GeoDataFrame(polys)
train_dict[loc[0]]['train'] = parallel_process(file_paths, polys_gDF, datasets)
# + id="XyeJ4GLNb5aP"
for loc in train_dict:
train_df = train_dict[loc]['train']
train_dict[loc]['features'] = calculate_features(feature_df=train_df, labels_df=train_labels, datasets=datasets, type_='train')
train_dict[loc]['features'].to_csv(str(PROCESSED / f'train_{loc}.csv'), index=False)
with open(str(PROCESSED / 'train_dict.pickle'), 'wb') as handle:
pickle.dump(train_dict, handle, protocol=pickle.HIGHEST_PROTOCOL)
# + colab={"base_uri": "https://localhost:8080/", "height": 412, "referenced_widgets": ["1d786d0890b3442fb6fcaa95161465fd", "d3e49ec2613b4c3c98bac193be72ee32", "c22fdc7ffbad4018b4da26fea19ce6ab", "0dca8a6cec094f2282f47d55e0955e9a", "ed38127b2e64488c824ca4ce26a66549", "80a4fb38c73249cd9d8ff29af6027558", "d6d7bf8d6cf842e49a5dcece6c0eff59", "70c0347ac55846878e789ad0857d4dc0", "ce8d3edcb8544d69a315b0231e8bd53f", "e1f9d5c57d004618aee7fa32ababca04", "b71c6991edbb4e48b8816204f896d4ee", "2779848da6ca4d9585986aab643a4963", "c40f05a69b9b491fb5be27bc9f58fa31", "d7b707afa3cb4d13bfe4ea92ffbc51c8", "5376001ce11349e69b963a41e2806d75", "<KEY>", "ee240139057943eb9b3260739098c3e0", "5f42f55a6c2744e494c5a2028d87a143", "<KEY>", "<KEY>", "7cde21e3df4b4253a7265962e6e8e465", "<KEY>", "1352074ec81f45148f303ad871239daa", "<KEY>", "<KEY>", "30fc3e39622f445fa76e440d0314be18", "d35a35a181eb4b6389e42dbd018108b1", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "844b098cec8e46bd8e9210454d5c5a67", "36026cbacc0742e1a032455b5ec3d82d", "<KEY>", "<KEY>", "e0fd64701e054e0d8aa5481ce2ec972e", "8d406f301948434398d5f3b13ee43d3a", "f47288379a2b428d9c3c3eec6361679a", "<KEY>", "<KEY>", "<KEY>", "15b29002e80a4e7884a4bc3621cc6dd6", "<KEY>", "cf9fb10a0f08419ca49b2e0f7edaf4cb", "ccd60ceab500461aad35a747f2b6eeed", "<KEY>", "<KEY>", "575056c510244680bad5a955d3fcc419", "ed31fae853c8462a81c63860c11a4489", "ae0070f452dc48329ab8da2e2ace084f", "<KEY>", "<KEY>", "db06e23a66fe49f6b25e80d23ec3bc82", "<KEY>", "92ae239c9c994c58a6bec17ccadcfa42", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "588fa9bee7f2426e914c8db9b2fa9012", "<KEY>", "c853f8b0e168455aa35ad25bb3e10500", "5e4d4a6dc36d4266afe6818ff56bf51a", "38821bb6723d44c6afecc8d0a5d78b71", "<KEY>", "<KEY>", "cf95e7d80cf04790a5f6500f2f27b709", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "1813311a1ceb432fb4c75992313ec1ad", "684c1e9dab2644a29aa8b24b02ec6c7a", "3fc9549cc1614a9285d6fe08a0eb2127", "<KEY>", "152a77db19324867a20b5a94186be7f1", "<KEY>", "ebd58587b0e24f38a3da63eb01328b34", "0f598fb13ce7402f940c42fede1c5c2b", "ad240f30da274fd9abb318694aac1222", "17cddce2650d41628dfd31746ad648fa", "<KEY>", "5f43bb4c5596475a8bc3a50d5f56be1c", "<KEY>", "00b5668bec064f9e955515710f6e9a26", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "c23f7971e4d6432cab6c2aff68fd8954", "ef620651d1de48f7b2021e4b5527f32a", "56c6c2daae5c4a64ade6969fcccab487", "<KEY>", "52ee20e44d2c49deb16e7ff39adf4830", "<KEY>", "<KEY>", "d92dc8eefd754c1a9f4e6ecc21a6d765"]} id="T74D4u6hXJZ1" outputId="fdaeef2a-c7e9-4e0b-b952-a361c99c892a"
test_maiac = satellite[(satellite["product"] == "maiac") & (satellite["split"] == "test")].copy()
grid_test = grid[grid.index.isin(submission.grid_id)]
test_dict = {}
locations = [['la', 'Los Angeles (SoCAB)'], ['dl', 'Delhi'], ['tpe', 'Taipei']]
for loc in locations:
print(f'Processing: {loc[1]}\n')
test_dict[loc[0]] = {}
satellite_subset = test_maiac[test_maiac.location == loc[0]].copy()
file_paths = list(satellite_subset.us_url)
grid_subset = grid_test[grid_test.location == loc[1]].copy()
assert grid_subset.index.isin(submission.grid_id).all()
polys = gpd.GeoSeries.from_wkt(grid_subset.wkt, crs=wgs84_crs)
polys.name = "geometry"
polys_gDF = gpd.GeoDataFrame(polys)
test_dict[loc[0]]['test'] = parallel_process(file_paths, polys_gDF, datasets)
# + id="kbxCfGLDgCB5"
cl = ['Optical_Depth_047_mean', 'Optical_Depth_047_min', 'Optical_Depth_047_max',
'Optical_Depth_055_mean', 'Optical_Depth_055_min', 'Optical_Depth_055_max',
'AOD_Uncertainty_mean', 'AOD_Uncertainty_min', 'AOD_Uncertainty_max',
'Column_WV_mean', 'Column_WV_min', 'Column_WV_max']
for loc in locations:
test_df = test_dict[loc[0]]['test']
submission_subset = submission[submission.grid_id.isin(grid[grid.location==loc[1]].index)].copy()
test_dict[loc[0]]['features'] = calculate_features(feature_df=test_df, labels_df=submission_subset, datasets=datasets, type_='test')
for c in cl:
if c.split('_')[-1] == 'mean':
replace_val = train_dict[loc[0]]['features'][c].mean()
elif c.split('_')[-1] == 'min':
replace_val = train_dict[loc[0]]['features'][c].min()
elif c.split('_')[-1] == 'max':
replace_val = train_dict[loc[0]]['features'][c].max()
else:
raise Exception('Incorrect parsing')
test_dict[loc[0]]['features'][c] = test_dict[loc[0]]['features'][c].fillna(replace_val)
test_dict[loc[0]]['features'].to_csv(str(PROCESSED / f'test_{loc[0]}.csv'), index=False)
with open(str(PROCESSED / 'test_dict.pickle'), 'wb') as handle:
pickle.dump(test_dict, handle, protocol=pickle.HIGHEST_PROTOCOL)
# + id="Wiyo4rIxAvse"
| src/models/preprocess_4maiac.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.10 64-bit (conda)
# metadata:
# interpreter:
# hash: e59a34f0b7d7ce4897ba06866367588679c8b5eae666dd4da2c81117d1a6e756
# name: python3
# ---
# ## My reading notes of [Bayesian Statistics The Fun Way](https://nostarch.com/learnbayes).
# ### Chapter 4 Creating a Binomial Probability Distribution
#
# Binomial Probability Distribution is used to calculate the probability of a certain number of successful outcomes, given a number of trials and the probality of the successful outcome.
#
# B(k; n, p)
#
# - k = The number of outcomes we care about
# - n = The number of trials
# - p = The probability of the event happening
#
# So you have to know the probability of the event happening in order to get this calculated, which is not usually the case in real life.
#
# The code below is coming from example at [scipy.stats.binom](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.binom.html).
# +
from scipy.stats import binom
import matplotlib.pyplot as plt
import numpy as np
fig, ax = plt.subplots(1, 1)
num_of_trials = 10
prob = 1 / 6
# x = np.arange(
# binom.ppf( 0.01, num_of_trials, prob ),
# binom.ppf( 0.99, num_of_trials, prob )
# )
x = np.arange(
start = 0,
stop = 11,
step = 1
)
ax.plot(
x,
binom.pmf( x, num_of_trials, prob ),
'bo',
ms=2,
label='Binomial Probablity Mass Function'
)
ax.vlines(x, 0, binom.pmf( x, num_of_trials, prob ), colors='b', lw=5, alpha=0.3)
# -
# ### Chapter 5 The Beta Distribution
#
# Probability, Statistics, and Inference
#
# - Probability: The chance of the event we care about happening. In real life you don't know this and want to figure it out.
# - Statistics: Data you observe. You make best of it and try to find probability out.
# - Inference: The process of figuring probability out from statistics.
#
#
# Usually you don't know the probability, but have just some data, a series of observations, or statistics. Starting from there you want to find out the probability.
#
# You can draw a beta distribution with [scipy.stats.beta](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.beta.html). Also you can calculate an integral with [scipy.integrate](https://docs.scipy.org/doc/scipy/reference/tutorial/integrate.html).
# +
# let's draw the beta distribution
from scipy.stats import beta
import matplotlib.pyplot as plt
import numpy as np
fig, ax = plt.subplots(1, 1)
# alpha is the number of positive outcomes
a = 14
# beta is the number of negative outcomes
b = 27
x = np.linspace(
beta.ppf(0.001, a, b),
beta.ppf(0.999, a, b),
100
)
ax.plot(x,
beta.pdf(x, a, b),
'g-',
lw = 2,
alpha = 0.2,
label = 'Beta Probablity Density Function'
)
# +
# you can calcualte integral with scipy.integrate
import scipy.integrate as integrate
result = integrate.quad(lambda p: beta.pdf(p, a, b), 0, 0.5)
result # returns (0.9807613458579021, 2.7538776087617885e-12)
| notes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Dependencies
import random
import json
import requests
# Let's get the JSON for 100 posts sequentially.
url = "http://jsonplaceholder.typicode.com/posts/"
# Create an empty list to store the responses
response_json = []
# Create random indices representing
# a user's choice of posts
indices = random.sample(list(range(1, 100)), 10)
indices
# Make a request for each of the indices
for x in range(len(indices)):
print(f"Making request number: {x} for ID: {indices[x]}")
# Get one of the posts
post_response = requests.get(url + str(indices[x]))
# Save post's JSON
response_json.append(post_response.json())
# Now we have 10 post objects,
# which we got by making 100 requests to the API.
print(f"We have {len(response_json)} posts!")
response_json
| 1/Activities/09-Ins_IterativeRequests/Solved/Ins_IterativeRequests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import numpy as np
import matplotlib.pyplot as plt
from simtk import unit
from simtk import openmm as omm
from simtk.openmm import app
import molsysmt as msm
from tqdm import tqdm
# # Alanine dipeptide in explicit solvent
# ## With OpenMM from scratch
from molecular_systems import files
pdbfile_path = files.alanine_dipeptide['octahedral_14.pdb']
msm.info(pdbfile_path)
view = msm.view(pdbfile_path, surface=True)
view
# +
temperature = 300.0*unit.kelvin
collisions_rate = 1.0/unit.picoseconds
total_time = 50.0*unit.picoseconds
saving_timestep = 1.0*unit.picoseconds
integration_timestep = 2.0*unit.femtoseconds
steps_per_cicle = round(saving_timestep/integration_timestep)
n_steps = round(total_time/integration_timestep)
n_cicles = round(n_steps/steps_per_cicle)
pdbfile = app.PDBFile(pdbfile_path)
topology = pdbfile.topology
positions = pdbfile.positions
forcefield = app.ForceField('amber14-all.xml', 'amber14/tip3p.xml')
system = forcefield.createSystem(topology,
nonbondedMethod=app.PME, nonbondedCutoff=9.0*unit.angstroms,
switchDistance=7.5*unit.angstroms,
rigidWater=True, constraints=app.HBonds, hydrogenMass=None)
forces = {ii.__class__.__name__ : ii for ii in system.getForces()}
forces['NonbondedForce'].setUseDispersionCorrection(True)
forces['NonbondedForce'].setEwaldErrorTolerance(1.0e-5)
integrator = omm.LangevinIntegrator(temperature, collisions_rate, integration_timestep)
integrator.setConstraintTolerance(0.00001)
platform = omm.Platform.getPlatformByName('CUDA')
simulation_properties = {}
simulation_properties['CudaPrecision']='mixed'
# Context.
context = omm.Context(system, integrator, platform, simulation_properties)
context.setPositions(positions)
# Energy Minimization
Potential_Energy = context.getState(getEnergy=True).getPotentialEnergy()
print('Before energy minimization: {}'.format(Potential_Energy))
omm.LocalEnergyMinimizer_minimize(context)
Potential_Energy = context.getState(getEnergy=True).getPotentialEnergy()
print('After energy minimization: {}'.format(Potential_Energy))
# +
# Initial velocities
n_atoms = topology.getNumAtoms()
velocities = np.zeros([n_atoms, 3], np.float32) * unit.nanometers/unit.picosecond
context.setVelocities(velocities)
# Reporter arrays: time, position, velocity, kinetic_energy, potential_energy
time = np.zeros([n_cicles], np.float32) * unit.picoseconds
trajectory = np.zeros([n_cicles, n_atoms, 3], np.float32) * unit.nanometers
velocity = np.zeros([n_cicles, n_atoms, 3], np.float32) * unit.nanometers/unit.picosecond
kinetic_energy = np.zeros([n_cicles], np.float32) * unit.kilocalories_per_mole
potential_energy = np.zeros([n_cicles], np.float32) * unit.kilocalories_per_mole
# Initial context in reporters
state = context.getState(getPositions=True, getVelocities=True, getEnergy=True)
time[0] = state.getTime()
trajectory[0] = state.getPositions()
velocity[0] = state.getVelocities()
kinetic_energy[0] = state.getKineticEnergy()
potential_energy[0] = state.getPotentialEnergy()
# Integration loop saving every cicle steps
for ii in tqdm(range(1, n_cicles)):
context.getIntegrator().step(steps_per_cicle)
state = context.getState(getPositions=True, getVelocities=True, getEnergy=True)
time[ii] = state.getTime()
trajectory[ii] = state.getPositions()
velocity[ii] = state.getVelocities()
kinetic_energy[ii] = state.getKineticEnergy()
potential_energy[ii] = state.getPotentialEnergy()
# -
trajectory.shape
print('The trajectory occupies {} gigabytes.'.format(trajectory.nbytes / 1024**3))
plt.plot(time, potential_energy)
plt.show()
_, _, phi_values, psi_values = msm.ramachandran_angles([topology, trajectory])
plt.scatter(phi_values, psi_values)
plt.show()
# ## With MolSysMT
# ## With this library
from molecular_systems import AlanineDipeptideExplicitSolvent
from molecular_systems.tools.md import langevin_NVT, energy_minimization
# +
dialanine=AlanineDipeptideExplicitSolvent(forcefield='AMBER14')
energy_minimization(dialanine)
time, trajectory, velocity, kin_energy, pot_energy = langevin_NVT(dialanine,
temperature = 300 * unit.kelvin,
friction = 1.0 / unit.picoseconds,
saving_timestep = 1.0 * unit.picoseconds,
total_time = 50.0 * unit.picoseconds)
# -
_, _, phi_values, psi_values = msm.ramachandran_angles([dialanine.topology, trajectory])
plt.scatter(phi_values, psi_values)
view = msm.view([dialanine.topology, trajectory], surface=False)
view
| docs/contents/alanine_tetrapeptide/Alanine_Tetrapeptide_Explicit_Solvent.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] colab_type="text" id="hRWOI1nxutyx"
# # Overview
#
# This CodeLab demonstrates how to build a LSTM model for MNIST recognition using Keras, and how to convert it to TensorFlow Lite.
#
# The CodeLab is very similar to the `tf.lite.experimental.nn.TFLiteLSTMCell`
# [CodeLab](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/lite/experimental/examples/lstm/TensorFlowLite_LSTM_Keras_Tutorial.ipynb). However, with the control flow support in the experimental new converter, we can define the model with control flow directly without refactoring the code.
#
# Also note: We're not trying to build the model to be a real world application, but only demonstrate how to use TensorFlow Lite. You can a build a much better model using CNN models. For a more canonical lstm codelab, please see [here](https://github.com/keras-team/keras/blob/master/examples/imdb_lstm.py).
# + [markdown] colab_type="text" id="wZCbNdY7MNSP"
# # Step 0: Prerequisites
#
# It's recommended to try this feature with the newest TensorFlow nightly pip build.
# + colab={} colab_type="code" id="6Zk2sUHUm5td"
# !pip install tf-nightly --upgrade
# + [markdown] colab_type="text" id="R3Ku1Lx9vvfX"
#
# ## Step 1: Build the MNIST LSTM model.
#
#
#
# + colab={} colab_type="code" id="yQpmCIqJPetJ"
import numpy as np
import tensorflow as tf
# + colab={} colab_type="code" id="wiYZoDlC5SEJ"
model = tf.keras.models.Sequential([
tf.keras.layers.Input(shape=(28, 28), name='input'),
tf.keras.layers.LSTM(20),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(10, activation=tf.nn.softmax, name='output')
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
model.summary()
# + [markdown] colab_type="text" id="Ff6X9gg_wk7K"
# ## Step 2: Train & Evaluate the model.
# We will train the model using MNIST data.
# + colab={} colab_type="code" id="23W41fiRPOmh"
# Load MNIST dataset.
(x_train, y_train), (x_test, y_test) = tf.keras.datasets.mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
x_train = x_train.astype(np.float32)
x_test = x_test.astype(np.float32)
# Change this to True if you want to test the flow rapidly.
# Train with a small dataset and only 1 epoch. The model will work poorly
# but this provides a fast way to test if the conversion works end to end.
_FAST_TRAINING = False
_EPOCHS = 5
if _FAST_TRAINING:
_EPOCHS = 1
_TRAINING_DATA_COUNT = 1000
x_train = x_train[:_TRAINING_DATA_COUNT]
y_train = y_train[:_TRAINING_DATA_COUNT]
model.fit(x_train, y_train, epochs=_EPOCHS)
model.evaluate(x_test, y_test, verbose=0)
# + [markdown] colab_type="text" id="NtPJGiIQw0nM"
# ## Step 3: Convert the Keras model to TensorFlow Lite model.
#
# Note here: we just convert to TensorFlow Lite model as usual.
# + colab={} colab_type="code" id="Tbuu_8PFz-x_"
converter = tf.lite.TFLiteConverter.from_keras_model(model)
# Note: It will NOT work without enabling the experimental converter!
# `experimental_new_converter` flag.
converter.experimental_new_converter = True
tflite_model = converter.convert()
# + [markdown] colab_type="text" id="5rHrZkIuxxar"
# ## Step 4: Check the converted TensorFlow Lite model.
#
# Now load the TensorFlow Lite model and use the TensorFlow Lite python interpreter to verify the results.
# + colab={} colab_type="code" id="8lao097MnFf2"
# Run the model with TensorFlow to get expected results.
expected = model.predict(x_test[0:1])
# Run the model with TensorFlow Lite
interpreter = tf.lite.Interpreter(model_content=tflite_model)
interpreter.allocate_tensors()
input_details = interpreter.get_input_details()
output_details = interpreter.get_output_details()
interpreter.set_tensor(input_details[0]["index"], x_test[0:1, :, :])
interpreter.invoke()
result = interpreter.get_tensor(output_details[0]["index"])
# Assert if the result of TFLite model is consistent with the TF model.
np.testing.assert_almost_equal(expected, result)
print("Done. The result of TensorFlow matches the result of TensorFlow Lite.")
# + colab={} colab_type="code" id="DWhGUkIs71Qu"
| tensorflow/lite/examples/experimental_new_converter/keras_lstm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### SageMaker Batch Transform outline
#
# 1. Download or otherwise retrieve the data.
# 2. Process / Prepare the data.
# 3. Upload the processed data to S3.
# - Save data locally
# - Upload to S3
# 4. Train a chosen model.
# - Set up the training job: https://docs.aws.amazon.com/sagemaker/latest/dg/API_CreateTrainingJob.html
# - Execute training job
# - Build model
# 5. Test the trained model (typically using a batch transform job).
# - Set up batch transform job
# - Execute batch transform job
# 6. Deploy the trained model.
# 7. Use the deployed model.
#
# #### SageMaker model components
#
# In SageMaker, a model is a collection of information that describes how to perform inference. For the most part, this comprises two very important pieces.
#
# The first is the **container** that holds the model inference functionality. For different types of models this code may be different but for simpler models and models provided by Amazon this is typically the same container that was used to train the model.
#
# The second is the **model artifacts**. These are the pieces of data that were created during the training process. For example, if we were fitting a linear model then the coefficients that were fit would be saved as model artifacts.
# When a model is fit using SageMaker, the process is as follows.
#
# First, a compute instance (basically a server somewhere) is started up with the properties that we specified.
#
# Next, when the compute instance is ready, the code, in the form of a container, that is used to fit the model is loaded and executed. When this code is executed, it is provided access to the training (and possibly validation) data stored on S3.
#
# Once the compute instance has finished fitting the model, the resulting model artifacts are stored on S3 and the compute instance is shut down.
#
# ### Sentiment analysis app deployment
#
# <img src="part-6_images/deployment_prod.png" alt="Deployment schema" style="width: 500px;"/>
#
# The way data flows through the app is as follows.
# - The user enters a review on our website.
# - Next, our website sends that data off to an endpoint, created using API Gateway.
# - Our endpoint acts as an interface to our Lambda function so our user data gets sent to the Lambda function.
# - Our Lambda function processes the user data and sends it off to the deployed model's endpoint.
# - The deployed model perform inference on the processed data and returns the inference results to the Lambda function.
# - The Lambda function returns the results to the original caller using the endpoint constructed using API Gateway.
# - Lastly, the website receives the inference results and displays those results to the user.
#
| part-6/SageMaker_Notes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# change color of some values in a dataframe based on a condition
# set_table_styles : mouse:hover
# .hide_index() hides the index of dataframe
# CSS styles for dataframe
import pandas as pd
df = pd.read_csv('datasets/diabetes.csv', nrows=10)
df.head()
df.info()
def color_heigh_red(value):
color = 'red' if (value > 30 or value < 18.5) else 'green'
return 'color: %s' % color
# using subset=[] we can pass the list of columns
df.style.applymap(color_heigh_red, subset=['BMI'])
# +
#highlight max
# -
def highlight_max(s):
is_max = s == s.max()
return ['background-color: pink' if v else '' for v in is_max]
df.style.apply(highlight_max).hide_index()
df.style.set_table_styles(
[{'selector': 'tr:hover',
'props': [('background-color', 'pink')]}]
)
df.style.set_table_styles(
[{'selector': 'tr:nth-of-type(even)',
'props': [('background', '#eee')]},
{'selector': 'tr:nth-of-type(odd)',
'props': [('background', '#ccc')]},
{'selector': 'th',
'props': [('background', 'red'),
('color', 'white'),
('font-family', 'Tahoma')]},
{'selector': 'td',
'props': [('font-family', 'Tahoma')]},
]
).hide_index()
| 01-Highlight_values_in_dataframe.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
#focus on spatial features, such as shot_distance
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# -
data = pd.read_csv('data.csv')
print data.shape
data.info()
#target variable, 5000 missing as test dataset
print data['shot_made_flag'].value_counts()
print data['shot_made_flag'].isnull().sum()
# Useless features: game_event_id, game_id, lat, lon, team_id, team_name
# ## 'action_type' and 'combined_shot_type'
# Both 'action_type' and 'combined_shot_type' seem to be relevant features
data['action_type'].unique()
data.groupby('action_type')['shot_made_flag'].mean().plot(kind='bar',figsize=(12,6))
data['combined_shot_type'].value_counts()
data.groupby('combined_shot_type')['shot_made_flag'].mean().plot(kind='bar',figsize=(12,6))
# ## Shot type
#
# two-point vs three-point, seems like relevant features
pd.crosstab(data['shot_type'],data['shot_made_flag'])
data.groupby('shot_type')['shot_made_flag'].mean()
# ## shot_distance
#
# seem like relevant features, may overlap with shot_zone_area/basic/range
plt.figure(figsize=(12,6))
sns.distplot(data[data['shot_made_flag']==1.0]['shot_distance'],label='shots')
sns.distplot(data[data['shot_made_flag']==0.0]['shot_distance'],label='missed')
plt.legend()
sns.violinplot(x='shot_made_flag',y='shot_distance',data=data)
# ## shot_zone_area/basic/range
#
# Correlate with shot_percentage, but highly correlate with shot_distance, maybe create 'backcourt' as a feature
data.groupby('shot_zone_area')['shot_made_flag'].mean()
sns.factorplot(x="shot_zone_area", y="shot_distance", hue="shot_made_flag",data=data, \
kind="violin",size=6, aspect=2)
data.groupby('shot_zone_basic')['shot_made_flag'].mean()
sns.factorplot(x="shot_zone_basic", y="shot_distance", hue="shot_made_flag",data=data, \
kind="violin",size=6, aspect=2)
data.groupby('shot_zone_range')['shot_made_flag'].mean()
sns.factorplot(x="shot_zone_range", y="shot_distance", hue="shot_made_flag",data=data, \
kind="violin",size=4, aspect=2)
# ## loc_x and loc_y
#
# Take absolute for loc_x; both loc_x and loc_y seem to correlate with shot_distance
data['loc_x'].describe()
plt.figure(figsize=(12,6))
sns.distplot(data[data['shot_made_flag']==1.0]['loc_x'],label='shots')
sns.distplot(data[data['shot_made_flag']==0.0]['loc_x'],label='missed')
plt.legend()
data['abs_loc_x'] = data['loc_x'].apply(lambda x: abs(x))
data['abs_loc_x'].describe()
plt.figure(figsize=(12,6))
sns.distplot(data[data['shot_made_flag']==1.0]['abs_loc_x'],label='shots')
sns.distplot(data[data['shot_made_flag']==0.0]['abs_loc_x'],label='missed')
plt.legend()
sns.lmplot(x='abs_loc_x',y='shot_distance',hue='shot_made_flag',data=data)
data['loc_y'].describe()
plt.figure(figsize=(12,6))
sns.distplot(data[data['shot_made_flag']==1.0]['loc_y'],label='shots')
sns.distplot(data[data['shot_made_flag']==0.0]['loc_y'],label='missed')
plt.xlim([-100,400])
plt.legend()
sns.lmplot(x='loc_y',y='shot_distance',hue='shot_made_flag',data=data)
# ## Modify base modeling
# Use both 'roc_auc' and 'accuracy' as metrics for cross-validation
# <br>
# Add 'shot_zone_basic/area/range' but no improvement
# <br>
# Add 'backcourt?' and no improvement
# <br>
# Add 'loc_y' and 'abs_loc_x' and some improvement
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.cross_validation import KFold, train_test_split,cross_val_score
from sklearn import metrics
#input df with both train and test
def base_feature(data):
#at home or not
data['home?'] = data['matchup'].apply(lambda x: x.split('@')[0].strip()=='LAL')
#2-point or 3-point shot
data['shot_type?'] = data['shot_type']=='2PT Field Goal'
#playoff or not
data['playoffs?'] = data['playoffs']==1
return data
#encode categorical variable into sorted integer, and plotting
def sort_encode(df, field):
ct = pd.crosstab(df.shot_made_flag, df[field]).apply(lambda x:x/x.sum(), axis=0)
temp = list(zip(ct.values[1, :], ct.columns))
temp.sort()
new_map = {}
for index, (acc, old_number) in enumerate(temp):
new_map[old_number] = index
new_field = field + '_sort_enumerated'
df[new_field] = df[field].map(new_map)
get_acc(df, new_field)
#plot one col in df against shot_made_flag percentage
def get_acc(df, col):
ct = pd.crosstab(df.shot_made_flag, df[col]).apply(lambda x:x/x.sum(), axis=0)
x, y = ct.columns, ct.values[1, :]
plt.figure(figsize=(7, 5))
plt.plot(x, y)
plt.xlabel(col)
plt.ylabel('% shots made')
# +
#base modeling with randomforest
def test_accuracy(data): #input data with cols_use and 'shot_made_flag'
clf = RandomForestClassifier(n_estimators=100,n_jobs=-1,max_depth=7) #specify tree and depth
return cross_val_score(clf, data.drop('shot_made_flag', 1), data.shot_made_flag,scoring='accuracy', cv=10)
# another test function with roc_auc as metrics instead of accuracy
def test_auc(data): #input data with cols_use and 'shot_made_flag'
clf = RandomForestClassifier(n_estimators=100,n_jobs=-1,max_depth=7) #specify tree and depth
return cross_val_score(clf, data.drop('shot_made_flag', 1), data.shot_made_flag,\
scoring='roc_auc', cv=10)
# -
data = base_feature(data)
data.columns
action_map = {action: i for i, action in enumerate(data.action_type.unique())}
data['action_type_enumerated'] = data.action_type.map(action_map)
#add column 'action_type_enumerated_sort_enumerated'
sort_encode(data, 'action_type_enumerated')
cols_use=['action_type_enumerated_sort_enumerated',
'playoffs?','home?', 'shot_type?', 'shot_distance','shot_made_flag']
df = data[cols_use]
df = df.dropna()
print test_auc(df).mean()
print test_accuracy(df).mean()
#add shot_zone_basic', slight improvement
sort_encode(data,'shot_zone_basic')
cols_use1=['action_type_enumerated_sort_enumerated','shot_zone_basic_sort_enumerated',
'playoffs?','home?', 'shot_type?', 'shot_distance','shot_made_flag']
df = data[cols_use1]
df = df.dropna()
print test_auc(df).mean()
print test_accuracy(df).mean()
#add 'shot_zone_area', slight improvement
sort_encode(data,'shot_zone_area')
cols_use2=['action_type_enumerated_sort_enumerated','shot_zone_area_sort_enumerated',
'playoffs?','home?', 'shot_type?', 'shot_distance','shot_made_flag']
df = data[cols_use2]
df = df.dropna()
print test_auc(df).mean()
print test_accuracy(df).mean()
#add 'shot_zone_range', no change
sort_encode(data,'shot_zone_range')
cols_use3=['action_type_enumerated_sort_enumerated','shot_zone_range_sort_enumerated',
'playoffs?','home?', 'shot_type?', 'shot_distance','shot_made_flag']
df = data[cols_use3]
df = df.dropna()
print test_auc(df).mean()
print test_accuracy(df).mean()
data['backcourt?'] = data['shot_zone_basic']=='Backcourt'
data['backcourt?'].value_counts()
#add 'backcourt?' as a feature but slight decrease in auc
cols_use4=['action_type_enumerated_sort_enumerated','backcourt?',
'playoffs?','home?', 'shot_type?', 'shot_distance','shot_made_flag']
df = data[cols_use4]
df = df.dropna()
print test_auc(df).mean()
print test_accuracy(df).mean()
#add loc_y as a feature, slight improve in both auc and accuracy
cols_use4=['action_type_enumerated_sort_enumerated','loc_y',
'playoffs?','home?', 'shot_type?', 'shot_distance','shot_made_flag']
df = data[cols_use4]
df = df.dropna()
print test_auc(df).mean()
print test_accuracy(df).mean()
#add abs_loc_x as a feature, more improve in accuracy than auc
data['abs_loc_x'] = data['loc_x'].apply(lambda x: abs(x))
cols_use4=['action_type_enumerated_sort_enumerated','abs_loc_x',
'playoffs?','home?', 'shot_type?', 'shot_distance','shot_made_flag']
df = data[cols_use4]
df = df.dropna()
print test_auc(df).mean()
print test_accuracy(df).mean()
#add both loc_y and abs_loc_x as features
cols_use5=['action_type_enumerated_sort_enumerated','abs_loc_x','loc_y',
'playoffs?','home?', 'shot_type?', 'shot_distance','shot_made_flag']
df = data[cols_use5]
df = df.dropna()
print test_auc(df).mean()
print test_accuracy(df).mean()
| kobe_data_exploration1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NOAA-CIRES Reanalysis temperature data
# The NOAA-CIRES [Twentieth Century Reanalysis (V2)](https://psl.noaa.gov/data/gridded/data.20thC_ReanV2.html) project provides objectively analyzed four-dimensional (latitude, longitude, height, time) weather data and their uncertainty.
# Data are available from 1871 January 1 through 2012 December 31,
# and are presented with 6-hr, daily, and monthly means
# on a 2.5-degree latitude x 2.5-degree longitude global grid
# at multiple pressure levels in the atmosphere, from the surface to 10 mb.
# The [GitHub repository](https://github.com/csdms/reccs-2021/) for this workshop includes a Twentieth Century Reanalysis (V2) dataset that contains daily mean 500 mb [geopotential height](https://en.wikipedia.org/wiki/Geopotential_height) data for a single day in 2010.
# The goal of this exercise is to read these data into this notebook and display them.
# The data are in [NetCDF](https://en.wikipedia.org/wiki/NetCDF), a data storage format that's ubiquitous in the geosciences. Over the years, there have been several libraries developed in Python to read and write NetCDF files, including:
#
# * scipy.io.netcdf
# * netcdf4
# * xarray
#
# The last, [*xarray*]((http://xarray.pydata.org/en/stable/)), is the current best choice for working with NetCDF.
# (It's really amazing.)
# However, this example is old--I think I wrote it first in 2011--and rather than rewrite it to use *xarray*, I kept the original library.
# I think this shows the power of a well-written Python library:
# it continues to work even after it's been superseded by a newer technology.
# Start by importing libraries used in this example. There are several.
import time
import calendar
import math
import numpy as np
from scipy.io import netcdf
from matplotlib import pyplot as plt
from mpl_toolkits.basemap import Basemap
# Because there's a bit of work required to read, prepare, and plot the data, I've broken the work into steps, each with a function to do the work of the step.
#
# The first is a function called *read*:
def read(reanalysis_file):
""" Reads data from a NOAA-CIRES 20th Century Reanalysis V2 file.
The file is NetCDF. It contains global 2.5-deg daily mean 500 mb heights.
Parameters
----------
reanalysis_file : str
The path to a 20CRv2 file.
Returns
-------
dict
A dictionary of data read from the file.
"""
try:
f = netcdf.netcdf_file(reanalysis_file, 'r', mmap=False)
except IOError:
print('File "' + reanalysis_file + '" cannot be read.')
return
data = {
'file':f.filename,
'level':f.variables['level'][0],
'time':f.variables['time'],
'hgt':f.variables['hgt'],
'lat':f.variables['lat'],
'lon':f.variables['lon']
}
f.close()
return data
# Let's identify the data file and use the *read* function to load its contents.
rean_file = "../data/X174.29.255.181.65.14.23.9.nc"
rean_data = read(rean_file)
rean_data
# **Discussion:** What happened here?
# +
# Take a quick look at the file contents with a built-in NetCDF tool.
# # !ncdump -h "../data/X174.172.16.58.3.14.23.9.nc"
# -
# To prepare the data for plotting, use another function, called *prep*:
def prep(rean_data):
"""Prepares 20th Century Reanalysis V2 for display.
Converts data read from a NOAA-CIRES 20th Century Reanalysis V2
file into more convenient formats. Returns a dict containing lon, lat
and hgt as NumPy arrays, and time as list of struct_time tuples.
Parameters
----------
rean_data : dict
A dict of data as returned from the `read` function.
Returns
-------
dict
A dict containing lon, lat and hgt as NumPy arrays, and time as list
of struct_time tuples.
"""
# Make a dict for storing results.
data = {
'lat' : rean_data['lat'].data,
'lon' : rean_data['lon'].data
}
# Apply scale_factor and add_offset properties to hgt variable.
# Add to data dict.
data['hgt'] = rean_data['hgt'].data * rean_data['hgt'].scale_factor \
+ rean_data['hgt'].add_offset
# Convert time variable (which is in hours since 0001-01-01) into
# calendar dates. Add to data dict.
start_time = '0001-01-01' # from rean_data['time'].units
start_time_cal = time.strptime(start_time, '%Y-%m-%d')
start_time_sec = calendar.timegm(start_time_cal)
sec_in_hour = 60.0*60.0
time_in_sec = rean_data['time'].data*sec_in_hour + start_time_sec
time_in_struct_time = [time.gmtime(i) for i in time_in_sec]
data['time'] = [time.strftime('%Y-%m-%d', j) for j in time_in_struct_time]
return data
# Pass the data read from the reanalysis file through the *prep* function:
prep_data = prep(rean_data)
prep_data
# **Discussion:** What happened here?
# The data are now ready to be plotted.
#
# To perform this task, use a third function, *view*:
def view(prep_data, dayofyear=46, show=False, outfile='gph.png'):
"""Draws a contour plot of the mean 500 mb geopotential surface.
Plot is for a specified day of the year with data from a NOAA-CIRES
20th Century Reanalysis file. The plot can be saved to a PNG file.
Parameters
----------
prep_data : dict
A dict of data returned from the `prep` function.
day_of_year: int, optional
An ordinal date.
show : bool, optional
Set this flag to display plot on screen; otherwise, write to file.
outfile : str, optional
Name of PNG file.
"""
# Set up map projection.
map = Basemap(projection='ortho',
lon_0=-105,
lat_0=60,
resolution='l')
map.drawcoastlines()
map.drawmapboundary()
map.drawparallels(range(-90, 120, 30))
map.drawmeridians(range(0, 420, 60))
# Transform lat/lon into map coordinates (meters).
x, y = map(*np.meshgrid(prep_data['lon'], prep_data['lat']))
# Extract a single day of heights.
hgt = prep_data['hgt'][dayofyear, 0, :, :]
# Set up consistent contour levels so the colorbar doesn't change.
delta = 100
hgt_min = math.floor(prep_data['hgt'].min()/delta)*delta
hgt_max = math.ceil(prep_data['hgt'].max()/delta)*delta
clevels = np.arange(hgt_min, hgt_max, delta)
# Draw contours of gph and annotate.
c = map.contourf(x, y, hgt, levels=clevels, cmap=plt.cm.RdYlBu_r)
cb = map.colorbar(c, 'right', size="3%", pad='5%')
cb.set_label('Geopotential Height (m)')
plt.title('500 mb Geopotential Heights : ' + prep_data['time'][dayofyear])
plt.text(0.5*max(plt.axis()), -0.1*max(plt.axis()),
'Data: NOAA-CIRES 20th Century Reanalysis, Version 2',
fontsize=10,
verticalalignment='bottom',
horizontalalignment='center')
# Either show plot or save it to a PNG file.
if show is True:
plt.show()
else:
plt.savefig(outfile, dpi=96)
plt.close()
return
# Pass the prepared data into the *view* function to produce a plot.
view(prep_data, show=True)
# ## Summary
# That's it! Go forth and try new things in Python.
| notebooks/6_noaa_cires_reanalysis_temperature_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SQL for Analyst 3 - Aggregation
# +
# set MySQL URL
user = "dz3vg"
password = ""
host = "localhost"
port = 3306
dbname = "dataapplab_db"
with open("MySQL.key", "r") as file:
password = file.read()
connection_string = f"mysql+mysqlconnector://{user}:{password}@{host}:{port}/{dbname}"
# connect to MySQL server
# %load_ext sql
# %sql $connection_string
# clean password
password = ""
connection_string = ""
# -
# %sql use dalba;
# + language="sql"
#
# select
# count(product_id),
# count(distinct product_id),
# count(distinct price),
# count(distinct sales_status)
# from
# products
# ;
# + language="sql"
#
# select
# min(price) as min_msrp,
# max(price) as max_msrp,
# sum(price) as total_msrp,
# count(product_id) as num_products
# from
# products
# ;
# + language="sql"
#
# select
# any_value(product_name),
# sales_status,
# count(distinct product_id),
# sum(price)
# from
# products
# group by sales_status
# ;
# + language="sql"
#
# select
# group_concat(distinct product_name separator " ~~~ "),
# sales_status,
# count(distinct product_id),
# sum(price)
# from
# products
# group by sales_status
# ;
# + language="sql"
#
# select
# sales_status,
# price,
# group_concat(distinct product_name
# order by price asc
# separator " >>> ") as product_names,
# count(*)
# from
# products
# group by sales_status, price
# ;
# + language="sql"
#
# select
# create_date,
# sum(price)
# from
# products
# group by create_date
# ;
# -
| Data Scientist Bootcamp/MySQL for Analyst/SQL-for-Analyst-3-Aggregation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import json
import pandas as pd
import datetime
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
matplotlib.rcParams['figure.figsize'] = [12, 8]
# +
raw_0 = open('raw_0.txt', 'r')
raw_1 = open('raw_1.txt', 'w')
for line in raw_0:
try:
line = line.replace('"', '|').replace("'", '"').strip()
message = json.loads(line)
msg = ['date', 'subject']
for header in message:
if header['name'] == 'Subject':
msg[1] = header['value'].replace("Noto", "").replace("[", "").replace("]", "").replace(",", " ").strip()
if header['name'] == 'Date':
msg[0] = header['value'].split(",")[-1].strip()
raw_1.write("{}\n".format(",".join(msg)))
print(msg)
except json.JSONDecodeError as e:
continue
raw_0.close()
raw_1.close()
# +
# Split into two files, add 'date,word'
# Clean timezone information
# TODO: modify original data collection script...
raw_1_lucas = open('raw_1_lucas.txt', 'r')
raw_1_charlie = open('raw_1_charlie.txt', 'r')
lucas = pd.read_csv(raw_1_lucas).iloc[::-1].reset_index()
charlie = pd.read_csv(raw_1_charlie).iloc[::-1].reset_index()
lucas.date = pd.to_datetime(lucas.date)
charlie.date = pd.to_datetime(charlie.date)
lucas["months"] = [int(i.days) / 30 for i in lucas.date - datetime.datetime(2016, 2, 12)]
charlie["months"] = [int(i.days) / 30 for i in charlie.date - datetime.datetime(2017, 9, 8)]
lucas["wc"] = lucas.index + 1
charlie["wc"] = charlie.index + 1
# -
lucas
# +
fig, ax = plt.subplots()
plt.plot(lucas.months, lucas.wc, label="Lucas")
plt.plot(charlie.months, charlie.wc, label="Charlie")
plt.title("New words over time", fontsize=20)
plt.xlabel("Age (months)")
plt.ylabel("Number of words")
plt.xticks(np.arange(13, 32, step=1))
plt.legend()
plt.grid(b=True, which='major', axis='both')
ax.yaxis.set_major_formatter(matplotlib.ticker.StrMethodFormatter('{x:,.0f}'))
# -
| words.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Magic Commands
# Magic commands (those that start with `%`) are commands that modify a configuration of Jupyter Notebooks. A number of magic commands are available by default (see list [here](http://ipython.readthedocs.io/en/stable/interactive/magics.html))--and many more can be added with extensions. The magic command added in this section allows `matplotlib` to display our plots directly on the browser instead of having to save them on a local file.
# %matplotlib inline
# # Activity 3.01: Optimizing a deep learning model
# In this activity we optimize our deep learning model. We aim achieve greater performance than our model `bitcoin_lstm_v0`, which is off at about **8.4%** from the real Bitcoin prices. We explore the following topics in this notebook:
#
# * Experimenting with different layers and the number of nodes
# * Grid search strategy for epoch and activation functions
import math
import numpy as np
import pandas as pd
import seaborn as sb
import matplotlib.pyplot as plt
from datetime import datetime, timedelta
from keras.models import load_model
from keras.models import Sequential
from keras.layers.recurrent import LSTM
from keras.callbacks import TensorBoard
from keras.layers.core import Dense, Activation, Dropout, ActivityRegularization
from scripts.utilities import (create_groups, split_lstm_input,
train_model, plot_two_series, rmse,
mape, denormalize)
plt.style.use('seaborn-white')
np.random.seed(0)
# ### Load Data
# We will load our same train and testing set from previous activitites.
train = pd.read_csv('data/train_dataset.csv')
test = pd.read_csv('data/test_dataset.csv')
train_data = create_groups(
train['close_point_relative_normalization'].values)
test_data = create_groups(
test['close_point_relative_normalization'].values)
X_train, Y_train = split_lstm_input(train_data)
# ### Reference Model
# For reference, let's load data for `v0` of our model and train it alongside future modifications.
model_v0 = load_model('bitcoin_lstm_v0.h5')
# %%time
train_model(model=model_v0, X=X_train, Y=Y_train, epochs=100, version=0, run_number=0)
# ### Adding Layers and Nodes
# We can modify our model to include other layers now. When using LSTM cells, one typically adds other LSTM layer in a sequence. In our case the layer will have the same number of neurons as the original layer.
#
# In order for this to work, however, we need to modify the parameter `return_sequences` to `True` on the first LSTM layer. We do this because the first layer expects a sequence of data with the same as input that the of the first layer. When this parameter is set to `False` the LSTM layer outputs the predicted parameters in a different incompatible output.
period_length = 7
number_of_periods = 76
batch_size=1
# +
model_v1 = Sequential()
model_v1.add(LSTM(
units=period_length,
batch_input_shape=(batch_size, number_of_periods, period_length),
input_shape=(number_of_periods, period_length),
return_sequences=True, stateful=False))
#
# Add new LSTM layer to this network here.
#
model_v1.add(Dense(units=period_length))
model_v1.add(Activation("linear"))
model_v1.compile(loss="mse", optimizer="rmsprop")
# -
# %%time
train_model(model=model_v1, X=X_train, Y=Y_train, epochs=100, version=1, run_number=0)
# ### Epochs
# Epochs are the number of times the network adjust its weights in response to data passing through and its loss function. Running a model for more epochs can allow it to learn more from data, but you also run the risk of overfitting.
#
# When training a model, prefer to increase the epochs exponentially until the loss function starts to plateau. In the case of the `bitcoin_lstm_v0` model, its loss function plateaus at about 100 epochs. If one attempts to train it at 10^3 epochs, the model barely gains any improvements.
#
# Change the number of epochs below
# to a higher number (try 10**3) and
# evaluate the results on TensorBoard.
#
number_of_epochs = 300
# +
model_v2 = Sequential()
model_v2.add(LSTM(
units=period_length,
batch_input_shape=(batch_size, number_of_periods, period_length),
input_shape=(number_of_periods, period_length),
return_sequences=True, stateful=False))
model_v2.add(LSTM(
units=period_length,
batch_input_shape=(batch_size, number_of_periods, period_length),
input_shape=(number_of_periods, period_length),
return_sequences=False, stateful=False))
model_v2.add(Dense(units=period_length))
model_v2.add(Activation("linear"))
model_v2.compile(loss="mse", optimizer="rmsprop")
# -
# %%time
train_model(model=model_v2, X=X_train, Y=Y_train, epochs=number_of_epochs,
version=2, run_number=0)
# ### Activation Functions
# Due to its non-linear properties and efficient computation, we will use the `relu` function as this network's activation function.
#
# Instead of using a ReLU, visit
# the Keras official documentation (https://keras.io/activations/)
# and choose a different function to try (maybe "tanh").
#
activation_function = "linear"
# +
model_v3 = Sequential()
model_v3.add(LSTM(
units=period_length,
batch_input_shape=(batch_size, number_of_periods, period_length),
input_shape=(number_of_periods, period_length),
return_sequences=True, stateful=False))
model_v3.add(LSTM(
units=period_length,
batch_input_shape=(batch_size, number_of_periods, period_length),
input_shape=(number_of_periods, period_length),
return_sequences=False, stateful=False))
model_v3.add(Dense(units=period_length))
model_v3.add(Activation(activation_function))
model_v3.compile(loss="mse", optimizer="rmsprop")
# -
# %%time
train_model(model=model_v3, X=X_train, Y=Y_train, epochs=300,
version=3, run_number=0)
# ### Regularization Strategies
# In this section we implement a `Dropout()` regularization strategy.
# +
model_v3 = Sequential()
model_v3.add(LSTM(
units=period_length,
batch_input_shape=(batch_size, number_of_periods, period_length),
input_shape=(number_of_periods, period_length),
return_sequences=True, stateful=False))
#
# Implement a Dropout() here.
#
model_v3.add(LSTM(
units=period_length,
batch_input_shape=(batch_size, number_of_periods, period_length),
input_shape=(number_of_periods, period_length),
return_sequences=False, stateful=False))
#
# Implement a Dropout() here too.
#
model_v3.add(Dense(units=period_length))
model_v3.add(Activation(activation_function))
model_v3.compile(loss="mse", optimizer="rmsprop")
# -
# %%time
train_model(model=model_v3, X=X_train, Y=Y_train, epochs=600,
version=3, run_number=0)
# ## Evaluate Models
# After creating the model versions above, we now have to evaluate which model is performing best in our test data. In order to do that we will use three metrics: MSE, RMSE, and MAPE. MSE is used to compare the error rates of the model on each predicted week. RMSE and MAPE are computed for making the model results easier to interpret.
combined_set = np.concatenate((train_data, test_data), axis=1)
def evaluate_model(model, kind='series'):
"""
Uses Keras model.evaluate() method to compute
the MSE for all future weeks in period.
Parameters
----------
model: Keras trained model
kind: str, default 'series'
Kind of evaluation to perform. If 'series',
then the model will perform an evaluation
over the complete series.
Returns
-------
evaluated_weeks: list
List of MSE values for each evaluated
test week.
"""
if kind == 'series':
predicted_weeks = []
for i in range(0, test_data.shape[1]):
input_series = combined_set[0:,i:i+76]
predicted_weeks.append(model.predict(input_series))
predicted_days = []
for week in predicted_weeks:
predicted_days += list(week[0])
return predicted_days
else:
evaluated_weeks = []
for i in range(0, test_data.shape[1]):
input_series = combined_set[0:,i:i+77]
X_test = input_series[0:,:-1].reshape(1, input_series.shape[1] - 1, 7)
Y_test = input_series[0:,-1:][0]
result = model.evaluate(x=X_test, y=Y_test, verbose=0)
evaluated_weeks.append(result)
return evaluated_weeks
def plot_weekly_mse(series, model_name, color):
ax = pd.Series(series).plot(drawstyle="steps-post",
figsize=(14,4),
linewidth=2,
color=color,
grid=True,
label=model_name,
alpha=0.7,
title='Mean Squared Error (MSE) for Test Data (all models)'.format(
model_name))
y = [i for i in range(0, len(series))]
yint = range(min(y), math.ceil(max(y))+1)
plt.xticks(yint)
ax.set_xlabel("Predicted Week")
ax.set_ylabel("MSE")
return ax
# Let's plot the weekly MSE.
def plot_weekly_predictions(predicted_days, name, display_plot=True,
variable='close'):
combined = pd.concat([train, test])
last_day = datetime.strptime(train['date'].max(), '%Y-%m-%d')
list_of_days = []
for days in range(1, len(predicted_days) + 1):
D = (last_day + timedelta(days=days)).strftime('%Y-%m-%d')
list_of_days.append(D)
predicted = pd.DataFrame({
'date': list_of_days,
'close_point_relative_normalization': predicted_days
})
combined['date'] = combined['date'].apply(
lambda x: datetime.strptime(x, '%Y-%m-%d'))
predicted['date'] = predicted['date'].apply(lambda x: datetime.strptime(x, '%Y-%m-%d'))
observed = combined[combined['date'] > train['date'].max()]
predicted['iso_week'] = predicted['date'].apply(
lambda x: x.strftime('%Y-%U'))
predicted_close = predicted.groupby('iso_week').apply(
lambda x: denormalize(observed, x))
plot_two_series(observed, predicted_close,
variable=variable,
title='{}: Predictions per Week'.format(name))
print('RMSE: {:.4f}'.format(
rmse(observed[variable][:-3],
predicted_close[variable])))
print('MAPE: {:.1f}%'.format(
mape(observed[variable][:-3],
predicted_close[variable])))
# Finally, let's evaluate each one the models trained in this activity in sequence.
models = [model_v0, model_v1, model_v2, model_v3]
for i, M in enumerate(models):
predicted_days = evaluate_model(M, kind='other')
plot_weekly_predictions(predicted_days, 'model_v{}'.format(i), display_plot=False)
# Our first model outperformed all the other models. Take the opportunity and teak the values for the optimization techniques above and attempt to beat the performance of that model!
| Chapter03/Activity3.01/.ipynb_checkpoints/Activity3.01_Optimizing_a_deep_learning_model-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Davidips/daa_2021_1/blob/master/2Diciembre.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="k4ACJ_fmAcRV"
def fnRecInfinita():
print("Hola")
fnRecInfinita()
# + colab={"base_uri": "https://localhost:8080/"} id="Pm2_ONJeArmE" outputId="13445110-3f4f-4249-9960-85e0dac9c2b5"
def fnRec(x):
if x == 0:
print("stop")
else:
fnRec(x-1)
print(x)
def main():
print("inicio del programa")
fnRec(5)
print("fin del programa ")
main()
# + colab={"base_uri": "https://localhost:8080/"} id="Isx--VrNBa_a" outputId="527dfde9-6ef3-425a-b2b4-ee013f21d374"
def printRev(x):
if x > 0:
printRev(x-1)
print(x)
printRev(3)
# + colab={"base_uri": "https://localhost:8080/"} id="FWfH4R-vIzPW" outputId="c649b805-2315-4d4a-a234-9195e032d13f"
def fibonacci(n):
if n==1 or n==0:
return n
else:
return (fibonacci(n-1) + fibonacci(n-2))
print(fibonacci(8))
| 2Diciembre.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simple convolution neural network MNIST digits classification with Keras and Tensorflow
# +
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# Keras (from TensorFlow) imports for the dataset and building NN
from tensorflow.keras import Sequential
from tensorflow.keras.layers import Dense, Dropout, Activation, Conv2D, MaxPooling2D, Flatten
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.datasets import mnist
from tensorflow.keras.models import load_model
from tensorflow.keras.losses import categorical_crossentropy
from tensorflow.keras.optimizers import Adadelta
from tensorflow.keras.backend import image_data_format
from tensorflow.keras.regularizers import l2
# -
# load train/test datasets
(X_train, y_train), (X_test, y_test) = mnist.load_data()
# Draw first several figures
fig = plt.figure()
for i in range(9):
plt.subplot(3,3,i+1)
plt.tight_layout()
plt.imshow(X_train[i], cmap='gray', interpolation='none')
plt.title("Digit: {}".format(y_train[i]))
plt.xticks([])
plt.yticks([])
# Draw pixel distribution of the first train digit
fig = plt.figure()
plt.subplot(2,1,1)
plt.imshow(X_train[0], cmap='gray', interpolation='none')
plt.title("Digit: {}".format(y_train[0]))
plt.xticks([])
plt.yticks([])
plt.subplot(2,1,2)
plt.hist(X_train[0].reshape(784))
plt.title("Pixel Value Distribution")
# +
# input image dimensions
img_rows, img_cols = 28, 28
# get input shape
if image_data_format() == 'channels_first':
X_train = X_train.reshape(X_train.shape[0], 1, img_rows, img_cols)
X_test = X_test.reshape(X_test.shape[0], 1, img_rows, img_cols)
input_shape = (1, img_rows, img_cols)
else:
X_train = X_train.reshape(X_train.shape[0], img_rows, img_cols, 1)
X_test = X_test.reshape(X_test.shape[0], img_rows, img_cols, 1)
input_shape = (img_rows, img_cols, 1)
# normalize train/test data
X_train = X_train.astype('float32')
X_test = X_test.astype('float32')
X_train /= 255
X_test /= 255
print('X_train:', X_train.shape)
print(X_train.shape[0], 'train samples')
print(X_test.shape[0], 'test samples')
# -
# convert train/test output data into categories
num_classes = 10
Y_train = to_categorical(y_train, num_classes)
Y_test = to_categorical(y_test, num_classes)
print('Y_train:', Y_train.shape)
print('Y_test:', Y_test.shape)
# +
# more info: https://keras.io/examples/mnist_cnn/
# https://github.com/AmmirMahdi/mnist-with-Keras---Conv2D/blob/master/Deep_Learning_mnist_with_Conv2D.ipynb
model = Sequential()
model.add(Conv2D(32, (3, 3), padding='same', activation='relu', input_shape=input_shape, kernel_regularizer=l2(0.01)))
model.add(MaxPooling2D((2, 2)))
model.add(Dropout(0.15))
#model.add(Conv2D(64, (3, 3), activation='relu'))
#model.add(MaxPooling2D((2, 2)))
#model.add(Dropout(0.15))
model.add(Flatten())
model.add(Dense(512, activation='relu', kernel_regularizer=l2(0.01)))
model.add(Dropout(0.15))
model.add(Dense(512, activation='relu', kernel_regularizer=l2(0.01)))
model.add(Dropout(0.15))
model.add(Dense(10, activation='softmax'))
model.compile(loss=categorical_crossentropy,
optimizer=Adadelta(),
metrics=['accuracy'])
# +
# train NN
batch_size = 128
# adapt number of iterations
epochs = 20
history = model.fit(X_train, Y_train,
batch_size=128,
epochs=epochs,
verbose=1,
validation_data=(X_test, Y_test))
# +
# evaluate NN model on test data
score = model.evaluate(X_test, Y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# +
# save data (optionally)
import os
save_dir = "results/"
if not os.path.exists(save_dir):
os.makedirs(save_dir)
# saving the model
model_name = 'keras_mnist_with_cnn_v1.h5'
model_path = os.path.join(save_dir, model_name)
model.save(model_path)
print('Saved trained model at %s ' % model_path)
# +
# restore data from file (optional)
mnist_model = load_model(model_path)
loss_and_metrics = mnist_model.evaluate(X_test, Y_test, verbose=2)
print("Test Loss", loss_and_metrics[0])
print("Test Accuracy", loss_and_metrics[1])
# +
# see which we predicted correctly and which not
predicted_classes = mnist_model.predict_classes(X_test)
print("predicted_classes:", predicted_classes.shape)
correct_indices = np.nonzero(predicted_classes == y_test)[0]
incorrect_indices = np.nonzero(predicted_classes != y_test)[0]
print(len(correct_indices)," classified correctly")
print(len(incorrect_indices)," classified incorrectly")
# +
# show 9 correctly and 9 incorrectly classified digits
plt.rcParams['figure.figsize'] = (7,14)
figure_evaluation = plt.figure()
# plot 9 correct predictions
for i, correct in enumerate(correct_indices[:9]):
plt.subplot(6,3,i+1)
plt.imshow(X_test[correct].reshape(28,28), cmap='gray', interpolation='none')
plt.title(
"Predicted: {}, Truth: {}".format(predicted_classes[correct],
y_test[correct]))
plt.xticks([])
plt.yticks([])
# plot 9 incorrect predictions
for i, incorrect in enumerate(incorrect_indices[:9]):
plt.subplot(6,3,i+10)
plt.imshow(X_test[incorrect].reshape(28,28), cmap='gray', interpolation='none')
plt.title(
"Predicted {}, Truth: {}".format(predicted_classes[incorrect],
y_test[incorrect]))
plt.xticks([])
plt.yticks([])
# -
| ml_mnist_digits_classification_cnn_with_keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
text = "The agent's phone number is 404-555-1234. Call soon!"
'phone' in text
import re
pattern = 'phone'
re.search(pattern,text)
pattern = 'NOT IN TEXT'
re.search(pattern,text)
pattern = 'phone'
match = re.search(pattern,text)
match
match.span()
match.start()
match.end()
text = "My phone number is 408-555-7777"
phone = re.search(r'\d\d\d-\d\d\d-\d\d\d\d', text)
phone
phone.group()
# +
# if we have a lot of digits so do not write to many \d we use quantifiers
phone = re.search(r'\d{3}-\d{3}-\d{4}', text)
# -
phone
phone_pattern = re.compile(r'(\d{3})-(\d{3})-(\d{4})')
results = re.search(phone_pattern,text)
results.group()
results.group(1)
results.group(2)
results.group(3)
# ## Additional regex Syntax
re.search(r'cat|dog','The dog is here')
re.findall(r'.at','The cat in the hat sat there')
# to find a number at the begining of a string
re.findall(r'^\d', '1 is a number')
# to find a number at the endining of a string
re.findall(r'\d$', 'the number is 2')
phrase = "there are 3 numbers 34 inside 5 this sentence"
pattern = r'[^\d]+ '
re.findall(pattern,phrase)
test_phrase = 'This is a string! But it has punctuation. How canwe remove it?'
clean = re.findall(r'[^!.? ]+',test_phrase)
' '.join(clean)
text = 'Only find the hypen-words in this sentence. But you do not know how long-ish they are'
pattern = r'[\w]+-[\w]+'
re.findall(pattern,text)
| 12-Advanced Python Modules/RegularExpressionPractice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
sys.path.append(os.environ.get('NOTEBOOK_ROOT'))
import xarray as xr
import numpy as np
import pandas as pd
# Ensure string casts of NumPy arrays
# print as much as possible (no '...').
np.set_printoptions(threshold=sys.maxsize)
import matplotlib.pyplot as plt
# -
from odc_gee.earthengine import Datacube as GEE_Datacube
dc = GEE_Datacube()
dc.list_products()
ds = dc.load(product='ls8_l2_c1_t1_google',
# Lake Mead
# lat = (36.384826, 36.445486),
# lon = (-114.400928, -114.326445),
# time= ('2020-06-01', '2021-06-30'),
# Elizabeth River, VA (sml tst)
lat=(36.89, 36.895),
lon=(-76.40,-76.395),
time=('2014-01-01', '2014-06-30'),
# Elizabeth River, VA
# lat=(36.894872, 36.969353),
# lon=(-76.394949,-76.260025),
# time=('2014-01-01', '2014-06-30'),
# Prospect Lake, CO
# lat = (38.822297, 38.827771),
# lon = (-104.804378, -104.793997),
# Pueblo Reservoir, CO
# lat = (38.23319252391589, 38.28728420493292),
# lon = (-104.80201949531988, -104.72186163717585),
# time=('2013-01-01', '2019-12-31'),
measurements=['pixel_qa'],
group_by='solar_day')
# +
from utils.data_cube_utilities.clean_mask import landsat_qa_clean_mask
clean_da = \
landsat_qa_clean_mask(ds, 'LANDSAT_8', collection='c1', level='l2')
# -
from skimage.morphology import remove_small_objects
clean_da.values = remove_small_objects(clean_da.values, min_size=64)
# +
# clean_da_filtered = xr.DataArray(clean_da_filtered)
# +
# clean_da_filtered.mean('dim_0').plot()
# -
clean_da.mean('time').plot()
# + tags=[]
import importlib
from utils.data_cube_utilities.voxel_visualizer import voxel_visualizer
importlib.reload(voxel_visualizer)
from utils.data_cube_utilities.voxel_visualizer.voxel_visualizer import voxel_visualize
voxel_visualize(clean_da,
# Voxel distances
# x_scale=1, y_scale=1, z_scale=1, distance_scale=1,
# Voxel size
# voxel_size=4,
# voxel_opacity=0.75,
# show_stats=True,
# show_controls=False
)
# -
| notebooks/experimental/voxel_visualizer/Voxel_Visualizer_Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "skip"}
# **Important**: Click on "*Kernel*" > "*Restart Kernel and Clear All Outputs*" *before* reading this chapter in [JupyterLab <img height="12" style="display: inline-block" src="static/link_to_jp.png">](https://jupyterlab.readthedocs.io/en/stable/) (e.g., in the cloud on [MyBinder <img height="12" style="display: inline-block" src="static/link_to_mb.png">](https://mybinder.org/v2/gh/webartifex/intro-to-python/master?urlpath=lab/tree/07_sequences_00_content.ipynb))
# + [markdown] slideshow={"slide_type": "slide"}
# # Chapter 7: Sequential Data
# + [markdown] slideshow={"slide_type": "skip"}
# We studied numbers (cf., [Chapter 5 <img height="12" style="display: inline-block" src="static/link_to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/05_numbers_00_content.ipynb)) and textual data (cf., [Chapter 6 <img height="12" style="display: inline-block" src="static/link_to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/06_text_00_content.ipynb)) first mainly because objects of the presented data types are "simple." That is so for two reasons: First, they are *immutable*, and, as we saw in the "*Who am I? And how many?*" section in [Chapter 1 <img height="12" style="display: inline-block" src="static/link_to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/01_elements_00_content.ipynb#Who-am-I?-And-how-many?), mutable objects can quickly become hard to reason about. Second, they are "flat" in the sense that they are *not* composed of other objects.
#
# The `str` type is a bit of a corner case in this regard. While one could argue that a longer `str` object, for example, `"text"`, is composed of individual characters, this is *not* the case in memory as the literal `"text"` only creates *one* object (i.e., one "bag" of $0$s and $1$s modeling all characters).
#
# This chapter, [Chapter 8 <img height="12" style="display: inline-block" src="static/link_to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/08_mfr_00_content.ipynb), and [Chapter 9 <img height="12" style="display: inline-block" src="static/link_to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/09_mappings_00_content.ipynb) introduce various "complex" data types. While some are mutable and others are not, they all share that they are primarily used to "manage," or structure, the memory in a program (i.e., they provide references to other objects). Unsurprisingly, computer scientists refer to the ideas behind these data types as **[data structures <img height="12" style="display: inline-block" src="static/link_to_wiki.png">](https://en.wikipedia.org/wiki/Data_structure)**.
#
# In this chapter, we focus on data types that model all kinds of sequential data. Examples of such data are [spreadsheets <img height="12" style="display: inline-block" src="static/link_to_wiki.png">](https://en.wikipedia.org/wiki/Spreadsheet) or [matrices <img height="12" style="display: inline-block" src="static/link_to_wiki.png">](https://en.wikipedia.org/wiki/Matrix_%28mathematics%29) and [vectors <img height="12" style="display: inline-block" src="static/link_to_wiki.png">](https://en.wikipedia.org/wiki/Vector_%28mathematics_and_physics%29). These formats share the property that they are composed of smaller units that come in a sequence of, for example, rows/columns/cells or elements/entries.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Collections vs. Sequences
# + [markdown] slideshow={"slide_type": "skip"}
# [Chapter 6 <img height="12" style="display: inline-block" src="static/link_to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/06_text_00_content.ipynb#A-"String"-of-Characters) already describes the **sequence** properties of `str` objects. In this section, we take a step back and study these properties one by one.
#
# The [collections.abc <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/collections.abc.html) module in the [standard library <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/index.html) defines a variety of **abstract base classes** (ABCs). We saw ABCs already in [Chapter 5 <img height="12" style="display: inline-block" src="static/link_to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/05_numbers_00_content.ipynb#The-Numerical-Tower), where we use the ones from the [numbers <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/numbers.html) module in the [standard library <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/index.html) to classify Python's numeric data types according to mathematical ideas. Now, we take the ABCs from the [collections.abc <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/collections.abc.html) module to classify the data types in this chapter according to their behavior in various contexts.
#
# As an illustration, consider `numbers` and `text` below, two objects of *different* types.
# + slideshow={"slide_type": "slide"}
numbers = [7, 11, 8, 5, 3, 12, 2, 6, 9, 10, 1, 4]
text = "Lorem ipsum dolor sit amet."
# + [markdown] slideshow={"slide_type": "skip"}
# Among others, one commonality between the two is that we may loop over them with the `for` statement. So, in the context of iteration, both exhibit the *same* behavior.
# + slideshow={"slide_type": "fragment"}
for number in numbers:
print(number, end=" ")
# + slideshow={"slide_type": "fragment"}
for character in text:
print(character, end=" ")
# + [markdown] slideshow={"slide_type": "skip"}
# In [Chapter 4 <img height="12" style="display: inline-block" src="static/link_to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/04_iteration_00_content.ipynb#Containers-vs.-Iterables), we referred to such types as *iterables*. That is *not* a proper [English](https://dictionary.cambridge.org/spellcheck/english-german/?q=iterable) word, even if it may sound like one at first sight. Yet, it is an official term in the Python world formalized with the `Iterable` ABC in the [collections.abc <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/collections.abc.html) module.
#
# For the data science practitioner, it is worthwhile to know such terms as, for example, the documentation on the [built-ins <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html) uses them extensively: In simple words, any built-in that takes an argument called "*iterable*" may be called with *any* object that supports being looped over. Already familiar [built-ins <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html) include [enumerate() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#enumerate), [sum() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#sum), or [zip() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#zip). So, they do *not* require the argument to be of a certain data type (e.g., `list`); instead, any *iterable* type works.
# + slideshow={"slide_type": "slide"}
import collections.abc as abc
# + slideshow={"slide_type": "fragment"}
abc.Iterable
# + [markdown] slideshow={"slide_type": "skip"}
# As seen in [Chapter 5 <img height="12" style="display: inline-block" src="static/link_to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/05_numbers_00_content.ipynb#Goose-Typing), we can use ABCs with the built-in [isinstance() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#isinstance) function to check if an object supports a behavior.
#
# So, let's "ask" Python if it can loop over `numbers` or `text`.
# + slideshow={"slide_type": "slide"}
isinstance(numbers, abc.Iterable)
# + slideshow={"slide_type": "fragment"}
isinstance(text, abc.Iterable)
# + [markdown] slideshow={"slide_type": "skip"}
# Contrary to `list` or `str` objects, numeric objects are *not* iterable.
# + slideshow={"slide_type": "skip"}
isinstance(999, abc.Iterable)
# + [markdown] slideshow={"slide_type": "skip"}
# Instead of asking, we could try to loop over `999`, but this results in a `TypeError`.
# + slideshow={"slide_type": "skip"}
for digit in 999:
print(digit)
# + [markdown] slideshow={"slide_type": "skip"}
# Most of the data types in this and the next chapter exhibit three [orthogonal <img height="12" style="display: inline-block" src="static/link_to_wiki.png">](https://en.wikipedia.org/wiki/Orthogonality) (i.e., "independent") behaviors, formalized by ABCs in the [collections.abc <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/collections.abc.html) module as:
# - `Iterable`: An object may be looped over.
# - `Container`: An object "contains" references to other objects; a "whole" is composed of many "parts."
# - `Sized`: The number of references to other objects, the "parts," is *finite*.
#
# The characteristical operation supported by `Container` types is the `in` operator for membership testing.
# + slideshow={"slide_type": "slide"}
0 in numbers
# + slideshow={"slide_type": "fragment"}
"l" in text
# + [markdown] slideshow={"slide_type": "skip"}
# Alternatively, we could also check if `numbers` and `text` are `Container` types with [isinstance() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#isinstance).
# + slideshow={"slide_type": "fragment"}
isinstance(numbers, abc.Container)
# + slideshow={"slide_type": "fragment"}
isinstance(text, abc.Container)
# + [markdown] slideshow={"slide_type": "skip"}
# Numeric objects do *not* "contain" references to other objects, and that is why they are considered "flat" data types. The `in` operator raises a `TypeError`. Conceptually speaking, Python views numeric types as "wholes" without any "parts."
# + slideshow={"slide_type": "skip"}
isinstance(999, abc.Container)
# + slideshow={"slide_type": "skip"}
9 in 999
# + [markdown] slideshow={"slide_type": "skip"}
# Analogously, being `Sized` types, we can pass `numbers` and `text` as the argument to the built-in [len() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#len) function and obtain "meaningful" results. The exact meaning depends on the data type: For `numbers`, [len() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#len) tells us how many elements are in the `list` object; for `text`, it tells us how many [Unicode characters <img height="12" style="display: inline-block" src="static/link_to_wiki.png">](https://en.wikipedia.org/wiki/Unicode) make up the `str` object. *Abstractly* speaking, both data types exhibit the *same* behavior of *finiteness*.
# + slideshow={"slide_type": "slide"}
len(numbers)
# + slideshow={"slide_type": "fragment"}
len(text)
# + slideshow={"slide_type": "fragment"}
isinstance(numbers, abc.Sized)
# + slideshow={"slide_type": "fragment"}
isinstance(text, abc.Sized)
# + [markdown] slideshow={"slide_type": "skip"}
# On the contrary, even though `999` consists of three digits for humans, numeric objects in Python have no concept of a "size" or "length," and the [len() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#len) function raises a `TypeError`.
# + slideshow={"slide_type": "skip"}
isinstance(999, abc.Sized)
# + slideshow={"slide_type": "skip"}
len(999)
# + [markdown] slideshow={"slide_type": "skip"}
# These three behaviors are so essential that whenever they coincide for a data type, it is called a **collection**, formalized with the `Collection` ABC. That is where the [collections.abc <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/collections.abc.html) module got its name from: It summarizes all ABCs related to collections; in particular, it defines a hierarchy of specialized kinds of collections.
#
# Without going into too much detail, one way to read the summary table at the beginning of the [collections.abc <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/collections.abc.html) module's documention is as follows: The first column, titled "ABC", lists all collection-related ABCs in Python. The second column, titled "Inherits from," indicates if the idea behind the ABC is *original* (e.g., the first row with the `Container` ABC has an empty "Inherits from" column) or a *combination* (e.g., the row with the `Collection` ABC has `Sized`, `Iterable`, and `Container` in the "Inherits from" column). The third and fourth columns list the methods that come with a data type following an ABC. We keep ignoring the methods named in the dunder style for now.
#
# So, let's confirm that both `numbers` and `text` are collections.
# + slideshow={"slide_type": "slide"}
isinstance(numbers, abc.Collection)
# + slideshow={"slide_type": "fragment"}
isinstance(text, abc.Collection)
# + [markdown] slideshow={"slide_type": "skip"}
# They share one more common behavior: When looping over them, we can *predict* the *order* of the elements or characters. The ABC in the [collections.abc <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/collections.abc.html) module corresponding to this behavior is `Reversible`. While sounding unintuitive at first, it is evident that if something is reversible, it must have a forward order, to begin with.
#
# The [reversed() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#reversed) built-in allows us to loop over the elements or characters in reverse order.
# + slideshow={"slide_type": "slide"}
for number in reversed(numbers):
print(number, end=" ")
# + slideshow={"slide_type": "fragment"}
for character in reversed(text):
print(character, end=" ")
# + slideshow={"slide_type": "fragment"}
isinstance(numbers, abc.Reversible)
# + slideshow={"slide_type": "fragment"}
isinstance(text, abc.Reversible)
# + [markdown] slideshow={"slide_type": "skip"}
# Collections that exhibit this fourth behavior are referred to as **sequences**, formalized with the `Sequence` ABC in the [collections.abc <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/collections.abc.html) module.
# + slideshow={"slide_type": "slide"}
isinstance(numbers, abc.Sequence)
# + slideshow={"slide_type": "fragment"}
isinstance(text, abc.Sequence)
# + [markdown] slideshow={"slide_type": "skip"}
# Most of the data types introduced in the remainder of this chapter are sequences. Nevertheless, we also look at some data types that are neither collections nor sequences but still useful to model sequential data in practice.
#
# In Python-related documentations, the terms collection and sequence are heavily used, and the data science practitioner should always think of them in terms of the three or four behaviors they exhibit.
#
# Data types that are collections but not sequences are covered in [Chapter 9 <img height="12" style="display: inline-block" src="static/link_to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/09_mappings_00_content.ipynb).
# + [markdown] slideshow={"slide_type": "slide"}
# ## The `list` Type
# + [markdown] slideshow={"slide_type": "skip"}
# As already seen multiple times, to create a `list` object, we use the *literal notation* and list all elements within brackets `[` and `]`.
# + slideshow={"slide_type": "slide"}
empty = []
# + slideshow={"slide_type": "fragment"}
simple = [40, 50]
# + [markdown] slideshow={"slide_type": "skip"}
# The elements do *not* need to be of the *same* type, and `list` objects may also be **nested**.
# + slideshow={"slide_type": "fragment"}
nested = [empty, 10, 20.0, "Thirty", simple]
# + [markdown] slideshow={"slide_type": "skip"}
# [PythonTutor <img height="12" style="display: inline-block" src="static/link_to_py.png">](http://pythontutor.com/visualize.html#code=empty%20%3D%20%5B%5D%0Asimple%20%3D%20%5B40,%2050%5D%0Anested%20%3D%20%5Bempty,%2010,%2020.0,%20%22Thirty%22,%20simple%5D&cumulative=false&curInstr=0&heapPrimitives=nevernest&mode=display&origin=opt-frontend.js&py=3&rawInputLstJSON=%5B%5D&textReferences=false) shows how `nested` holds references to the `empty` and `simple` objects. Technically, it holds three more references to the `10`, `20.0`, and `"Thirty"` objects as well. However, to simplify the visualization, these three objects are shown right inside the `nested` object. That may be done because they are immutable and "flat" data types. In general, the $0$s and $1$s inside a `list` object in memory always constitute references to other objects only.
# + slideshow={"slide_type": "fragment"}
nested
# + [markdown] slideshow={"slide_type": "skip"}
# Let's not forget that `nested` is an object on its own with an *identity* and *data type*.
# + slideshow={"slide_type": "skip"}
id(nested)
# + slideshow={"slide_type": "skip"}
type(nested)
# + [markdown] slideshow={"slide_type": "skip"}
# Alternatively, we use the built-in [list() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#func-list) constructor to create a `list` object out of any (finite) *iterable* we pass to it as the argument.
#
# For example, we can wrap the [range() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#func-range) built-in with [list() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#func-list): As described in [Chapter 4 <img height="12" style="display: inline-block" src="static/link_to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/04_iteration_00_content.ipynb#Containers-vs.-Iterables), `range` objects, like `range(1, 13)` below, are iterable and generate `int` objects "on the fly" (i.e., one by one). The [list() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#func-list) around it acts like a `for`-loop and **materializes** twelve `int` objects in memory that then become the elements of the newly created `list` object. [PythonTutor <img height="12" style="display: inline-block" src="static/link_to_py.png">](http://pythontutor.com/visualize.html#code=r%20%3D%20range%281,%2013%29%0Al%20%3D%20list%28range%281,%2013%29%29&cumulative=false&curInstr=0&heapPrimitives=nevernest&mode=display&origin=opt-frontend.js&py=3&rawInputLstJSON=%5B%5D&textReferences=false) shows this difference visually.
# + slideshow={"slide_type": "slide"}
list(range(1, 13))
# + [markdown] slideshow={"slide_type": "skip"}
# Beware of passing a `range` object over a "big" horizon as the argument to [list() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#func-list) as that may lead to a `MemoryError` and the computer crashing.
# + slideshow={"slide_type": "fragment"}
list(range(999_999_999_999))
# + [markdown] slideshow={"slide_type": "skip"}
# As another example, we create a `list` object from a `str` object, which is iterable, as well. Then, the individual characters become the elements of the new `list` object!
# + slideshow={"slide_type": "fragment"}
list("iterable")
# + [markdown] slideshow={"slide_type": "slide"}
# ### Sequence Behaviors
# + [markdown] slideshow={"slide_type": "skip"}
# `list` objects are *sequences*. To reiterate that, we briefly summarize the *four* behaviors of a sequence and provide some more `list`-specific details below:
# + [markdown] slideshow={"slide_type": "slide"}
# - `Container`:
# - holds references to other objects in memory (with their own *identity* and *type*)
# - implements membership testing via the `in` operator
# - `Iterable`:
# - supports being looped over
# - works with the `for` or `while` statements
# - `Reversible`:
# - the elements come in a *predictable* order that we may loop over in a forward or backward fashion
# - works with the [reversed() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#reversed) built-in
# - `Sized`:
# - the number of elements is finite *and* known in advance
# - works with the built-in [len() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#len) function
# + [markdown] slideshow={"slide_type": "skip"}
# The "length" of `nested` is *five* because `empty` and `simple` count as *one* element each. In other words, `nested` holds five references to other objects, two of which are `list` objects.
# + slideshow={"slide_type": "slide"}
len(nested)
# + [markdown] slideshow={"slide_type": "skip"}
# With a `for`-loop, we can iterate over all elements in a *predictable* order, forward or backward. As `list` objects hold *references* to other *objects*, these have an *indentity* and may even be of *different* types; however, the latter observation is rarely, if ever, useful in practice.
# + slideshow={"slide_type": "slide"}
for element in nested:
print(str(element).ljust(10), str(id(element)).ljust(18), type(element))
# + slideshow={"slide_type": "slide"}
for element in reversed(nested):
print(element, end=" ")
# + [markdown] slideshow={"slide_type": "skip"}
# The `in` operator checks if a given object is "contained" in a `list` object. It uses the `==` operator behind the scenes (i.e., *not* the `is` operator) conducting a **[linear search <img height="12" style="display: inline-block" src="static/link_to_wiki.png">](https://en.wikipedia.org/wiki/Linear_search)**: So, Python implicitly loops over *all* elements and only stops prematurely if an element evaluates equal to the searched object. A linear search may, therefore, be relatively *slow* for big `list` objects.
# + slideshow={"slide_type": "slide"}
10 in nested
# + [markdown] slideshow={"slide_type": "skip"}
# `20` compares equal to the `20.0` in `nested`.
# + slideshow={"slide_type": "fragment"}
20 in nested
# + slideshow={"slide_type": "fragment"}
30 in nested
# + [markdown] slideshow={"slide_type": "slide"}
# ### Indexing
# + [markdown] slideshow={"slide_type": "skip"}
# Because of the *predictable* order and the *finiteness*, each element in a sequence can be labeled with a unique *index*, an `int` object in the range $0 \leq \text{index} < \lvert \text{sequence} \rvert$.
#
# Brackets, `[` and `]`, are the literal syntax for accessing individual elements of any sequence type. In this book, we also call them the *indexing operator* in this context.
# + slideshow={"slide_type": "slide"}
nested[0]
# + [markdown] slideshow={"slide_type": "skip"}
# The last index is one less than `len(nested)`, and Python raises an `IndexError` if we look up an index that is not in the range.
# + slideshow={"slide_type": "skip"}
nested[5]
# + [markdown] slideshow={"slide_type": "skip"}
# Negative indices are used to count in reverse order from the end of a sequence, and brackets may be chained to access nested objects. So, to access the `50` inside `simple` via the `nested` object, we write `nested[-1][1]`.
# + slideshow={"slide_type": "fragment"}
nested[-1]
# + slideshow={"slide_type": "fragment"}
nested[-1][1]
# + [markdown] slideshow={"slide_type": "slide"}
# ### Slicing
# + [markdown] slideshow={"slide_type": "skip"}
# Slicing `list` objects works analogously to slicing `str` objects: We use the literal syntax with either one or two colons `:` inside the brackets `[]` to separate the *start*, *stop*, and *step* values. Slicing creates a *new* `list` object with the elements chosen from the original one.
#
# For example, to obtain the three elements in the "middle" of `nested`, we slice from `1` (including) to `4` (excluding).
# + slideshow={"slide_type": "slide"}
nested[1:4]
# + [markdown] slideshow={"slide_type": "skip"}
# To obtain "every other" element, we slice from beginning to end, defaulting to `0` and `len(nested)` when omitted, in steps of `2`.
# + slideshow={"slide_type": "fragment"}
nested[::2]
# + [markdown] slideshow={"slide_type": "skip"}
# The literal notation with the colons `:` is *syntactic sugar*. It saves us from using the [slice() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#slice) built-in to create `slice` objects. [slice() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#slice) takes *start*, *stop*, and *step* arguments in the same way as the familiar [range() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#func-range), and the `slice` objects it creates are used just as *indexes* above.
#
# In most cases, the literal notation is more convenient to use; however, with `slice` objects, we can give names to slices and reuse them across several sequences.
# + slideshow={"slide_type": "skip"}
middle = slice(1, 4)
# + slideshow={"slide_type": "skip"}
type(middle)
# + slideshow={"slide_type": "skip"}
nested[middle]
# + slideshow={"slide_type": "skip"}
numbers[middle]
# + slideshow={"slide_type": "skip"}
text[middle]
# + [markdown] slideshow={"slide_type": "skip"}
# `slice` objects come with three read-only attributes `start`, `stop`, and `step` on them.
# + slideshow={"slide_type": "skip"}
middle.start
# + slideshow={"slide_type": "skip"}
middle.stop
# + [markdown] slideshow={"slide_type": "skip"}
# If not passed to [slice() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#slice), these attributes default to `None`. That is why the cell below has no output.
# + slideshow={"slide_type": "skip"}
middle.step
# + [markdown] slideshow={"slide_type": "skip"}
# A good trick to know is taking a "full" slice: This copies *all* elements of a `list` object into a *new* `list` object.
# + slideshow={"slide_type": "slide"}
nested_copy = nested[:]
# + slideshow={"slide_type": "fragment"}
nested_copy
# + [markdown] slideshow={"slide_type": "skip"}
# At first glance, `nested` and `nested_copy` seem to cause no pain. For `list` objects, the comparison operator `==` goes over the elements in both operands in a pairwise fashion and checks if they all evaluate equal (cf., the "*List Comparison*" section below for more details).
#
# We confirm that `nested` and `nested_copy` compare equal as could be expected but also that they are *different* objects.
# + slideshow={"slide_type": "fragment"}
nested == nested_copy
# + slideshow={"slide_type": "fragment"}
nested is nested_copy
# + [markdown] slideshow={"slide_type": "skip"}
# However, as [PythonTutor <img height="12" style="display: inline-block" src="static/link_to_py.png">](http://pythontutor.com/visualize.html#code=nested%20%3D%20%5B%5B%5D,%2010,%2020.0,%20%22Thirty%22,%20%5B40,%2050%5D%5D%0Anested_copy%20%3D%20nested%5B%3A%5D&cumulative=false&curInstr=0&heapPrimitives=nevernest&mode=display&origin=opt-frontend.js&py=3&rawInputLstJSON=%5B%5D&textReferences=false) reveals, only the *references* to the elements are copied, and not the objects in `nested` themselves! Because of that, `nested_copy` is a so-called **[shallow copy <img height="12" style="display: inline-block" src="static/link_to_wiki.png">](https://en.wikipedia.org/wiki/Object_copying#Shallow_copy)** of `nested`.
#
# We could also see this with the [id() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#id) function: The respective first elements in both `nested` and `nested_copy` are the *same* object, namely `empty`. So, we have three ways of accessing the *same* address in memory. Also, we say that `nested` and `nested_copy` partially share the *same* state.
# + slideshow={"slide_type": "skip"}
nested[0] is nested_copy[0]
# + slideshow={"slide_type": "skip"}
id(nested[0])
# + slideshow={"slide_type": "skip"}
id(nested_copy[0])
# + [markdown] slideshow={"slide_type": "skip"}
# Knowing this becomes critical if the elements in a `list` object are mutable objects (i.e., we can change them *in place*), and this is the case with `nested` and `nested_copy`, as we see in the next section on "*Mutability*".
#
# As both the original `nested` object and its copy reference the *same* `list` objects in memory, any changes made to them are visible to both! Because of that, working with shallow copies can easily become confusing.
#
# Instead of a shallow copy, we could also create a so-called **[deep copy <img height="12" style="display: inline-block" src="static/link_to_wiki.png">](https://en.wikipedia.org/wiki/Object_copying#Deep_copy)** of `nested`: Then, the copying process recursively follows every reference in a nested data structure and creates copies of *every* object found.
#
# To explicitly create shallow or deep copies, the [copy <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/copy.html) module in the [standard library <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/index.html) provides two functions, [copy() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/copy.html#copy.copy) and [deepcopy() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/copy.html#copy.deepcopy). We must always remember that slicing creates *shallow* copies only.
# + slideshow={"slide_type": "skip"}
import copy
# + slideshow={"slide_type": "skip"}
nested_deep_copy = copy.deepcopy(nested)
# + slideshow={"slide_type": "skip"}
nested == nested_deep_copy
# + [markdown] slideshow={"slide_type": "skip"}
# Now, the first elements of `nested` and `nested_deep_copy` are *different* objects, and [PythonTutor <img height="12" style="display: inline-block" src="static/link_to_py.png">](http://pythontutor.com/visualize.html#code=import%20copy%0Anested%20%3D%20%5B%5B%5D,%2010,%2020.0,%20%22Thirty%22,%20%5B40,%2050%5D%5D%0Anested_deep_copy%20%3D%20copy.deepcopy%28nested%29&cumulative=false&curInstr=0&heapPrimitives=nevernest&mode=display&origin=opt-frontend.js&py=3&rawInputLstJSON=%5B%5D&textReferences=false) shows that there are *six* `list` objects in memory.
# + slideshow={"slide_type": "skip"}
nested[0] is nested_deep_copy[0]
# + slideshow={"slide_type": "skip"}
id(nested[0])
# + slideshow={"slide_type": "skip"}
id(nested_deep_copy[0])
# + [markdown] slideshow={"slide_type": "skip"}
# As this [StackOverflow question <img height="12" style="display: inline-block" src="static/link_to_so.png">](https://stackoverflow.com/questions/184710/what-is-the-difference-between-a-deep-copy-and-a-shallow-copy) shows, understanding shallow and deep copies is a common source of confusion independent of the programming language.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Mutability
# + [markdown] slideshow={"slide_type": "skip"}
# In contrast to `str` objects, `list` objects are *mutable*: We may assign new elements to indices or slices and also remove elements. That changes the *references* in a `list` object. In general, if an object is *mutable*, we say that it may be changed *in place*.
# + slideshow={"slide_type": "slide"}
nested[0] = 0
# + slideshow={"slide_type": "fragment"}
nested
# + [markdown] slideshow={"slide_type": "skip"}
# When we re-assign a slice, we can even change the size of the `list` object.
# + slideshow={"slide_type": "fragment"}
nested[:4] = [100, 100, 100]
# + slideshow={"slide_type": "fragment"}
nested
# + slideshow={"slide_type": "fragment"}
len(nested)
# + [markdown] slideshow={"slide_type": "skip"}
# The `list` object's identity does *not* change. That is the main point behind mutable objects.
# + slideshow={"slide_type": "skip"}
id(nested) # same memory location as before
# + [markdown] slideshow={"slide_type": "skip"}
# `nested_copy` is unchanged!
# + slideshow={"slide_type": "slide"}
nested_copy
# + [markdown] slideshow={"slide_type": "skip"}
# Let's change the nested `[40, 50]` via `nested_copy` into `[1, 2, 3]` by replacing all its elements.
# + slideshow={"slide_type": "fragment"}
nested_copy[-1][:] = [1, 2, 3]
# + slideshow={"slide_type": "fragment"}
nested_copy
# + [markdown] slideshow={"slide_type": "skip"}
# That has a surprising side effect on `nested`.
# + slideshow={"slide_type": "fragment"}
nested
# + [markdown] slideshow={"slide_type": "skip"}
# That is precisely the confusion we talked about above when we said that `nested_copy` is a *shallow* copy of `nested`. [PythonTutor <img height="12" style="display: inline-block" src="static/link_to_py.png">](http://pythontutor.com/visualize.html#code=nested%20%3D%20%5B%5B%5D,%2010,%2020.0,%20%22Thirty%22,%20%5B40,%2050%5D%5D%0Anested_copy%20%3D%20nested%5B%3A%5D%0Anested%5B%3A4%5D%20%3D%20%5B100,%20100,%20100%5D%0Anested_copy%5B-1%5D%5B%3A%5D%20%3D%20%5B1,%202,%203%5D&cumulative=false&curInstr=0&heapPrimitives=nevernest&mode=display&origin=opt-frontend.js&py=3&rawInputLstJSON=%5B%5D&textReferences=false) shows how both reference the *same* nested `list` object that is changed *in place* from `[40, 50]` into `[1, 2, 3]`.
#
# Lastly, we use the `del` statement to remove an element.
# + slideshow={"slide_type": "slide"}
del nested[-1]
# + slideshow={"slide_type": "fragment"}
nested
# + [markdown] slideshow={"slide_type": "skip"}
# The `del` statement also works for slices. Here, we remove all references `nested` holds.
# + slideshow={"slide_type": "fragment"}
del nested[:]
# + slideshow={"slide_type": "fragment"}
nested
# + [markdown] slideshow={"slide_type": "skip"}
# Mutability for sequences is formalized by the `MutableSequence` ABC in the [collections.abc <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/collections.abc.html) module.
# + [markdown] slideshow={"slide_type": "slide"}
# ### List Methods
# + [markdown] slideshow={"slide_type": "skip"}
# The `list` type is an essential data structure in any real-world Python application, and many typical `list`-related algorithms from computer science theory are already built into it at the C level (cf., the [documentation <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/stdtypes.html#mutable-sequence-types) or the [tutorial <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/tutorial/datastructures.html#more-on-lists) for a full overview; unfortunately, not all methods have direct links). So, understanding and applying the built-in methods of the `list` type not only speeds up the development process but also makes programs significantly faster.
#
# In contrast to the `str` type's methods in [Chapter 6 <img height="12" style="display: inline-block" src="static/link_to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/06_text_00_content.ipynb#String-Methods) (e.g., [upper() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/stdtypes.html#str.upper) or [lower() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/stdtypes.html#str.lower)), the `list` type's methods that mutate an object do so *in place*. That means they *never* create *new* `list` objects and return `None` to indicate that. So, we must *never* assign the return value of `list` methods to the variable holding the list!
#
# Let's look at the following `names` example.
# + slideshow={"slide_type": "slide"}
names = ["Carl", "Peter"]
# + [markdown] slideshow={"slide_type": "skip"}
# To add an object to the end of `names`, we use the `append()` method. The code cell shows no output indicating that `None` must be the return value.
# + slideshow={"slide_type": "fragment"}
names.append("Eckardt")
# + slideshow={"slide_type": "fragment"}
names
# + [markdown] slideshow={"slide_type": "skip"}
# With the `extend()` method, we may also append multiple elements provided by an iterable. Here, the iterable is a `list` object itself holding two `str` objects.
# + slideshow={"slide_type": "slide"}
names.extend(["Karl", "Oliver"])
# + slideshow={"slide_type": "fragment"}
names
# + [markdown] slideshow={"slide_type": "skip"}
# Similar to `append()`, we may add a new element at an arbitrary position with the `insert()` method. `insert()` takes two arguments, an *index* and the element to be inserted.
# + slideshow={"slide_type": "slide"}
names.insert(1, "Berthold")
# + slideshow={"slide_type": "fragment"}
names
# + [markdown] slideshow={"slide_type": "skip"}
# `list` objects may be sorted *in place* with the [sort() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/stdtypes.html#list.sort) method. That is different from the built-in [sorted() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#sorted) function that takes any *finite* and *iterable* object and returns a *new* `list` object with the iterable's elements sorted!
# + slideshow={"slide_type": "slide"}
sorted(names)
# + [markdown] slideshow={"slide_type": "skip"}
# As the previous code cell created a *new* `list` object, `names` is still unsorted.
# + slideshow={"slide_type": "fragment"}
names
# + [markdown] slideshow={"slide_type": "skip"}
# Let's sort the elements in `names` instead.
# + slideshow={"slide_type": "fragment"}
names.sort()
# + slideshow={"slide_type": "fragment"}
names
# + [markdown] slideshow={"slide_type": "skip"}
# To sort in reverse order, we pass a keyword-only `reverse=True` argument to either the [sort() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/stdtypes.html#list.sort) method or the [sorted() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#sorted) function.
# + slideshow={"slide_type": "slide"}
names.sort(reverse=True)
# + slideshow={"slide_type": "fragment"}
names
# + [markdown] slideshow={"slide_type": "skip"}
# The [sort() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/stdtypes.html#list.sort) method and the [sorted() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#sorted) function sort the elements in `names` in alphabetical order, forward or backward. However, that does *not* hold in general.
#
# We mention above that `list` objects may contain objects of *any* type and even of *mixed* types. Because of that, the sorting is **[delegated <img height="12" style="display: inline-block" src="static/link_to_wiki.png">](https://en.wikipedia.org/wiki/Delegation_(object-oriented_programming))** to the elements in a `list` object. In a way, Python "asks" the elements in a `list` object to sort themselves. As `names` contains only `str` objects, they are sorted according the the comparison rules explained in [Chapter 6 <img height="12" style="display: inline-block" src="static/link_to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/06_text_00_content.ipynb#String-Comparison).
#
# To customize the sorting, we pass a keyword-only `key` argument to [sort() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/stdtypes.html#list.sort) or [sorted() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#sorted), which must be a `function` object accepting *one* positional argument. Then, the elements in the `list` object are passed to that one by one, and the return values are used as the **sort keys**. The `key` argument is also a popular use case for `lambda` expressions.
#
# For example, to sort `names` not by alphabet but by the names' lengths, we pass in a reference to the built-in [len() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#len) function as `key=len`. Note that there are *no* parentheses after `len`!
# + slideshow={"slide_type": "fragment"}
names.sort(key=len)
# + [markdown] slideshow={"slide_type": "skip"}
# If two names have the same length, their relative order is kept as is. That is why `"Karl"` comes before `"Carl" ` below. A [sorting algorithm <img height="12" style="display: inline-block" src="static/link_to_wiki.png">](https://en.wikipedia.org/wiki/Sorting_algorithm) with that property is called **[stable <img height="12" style="display: inline-block" src="static/link_to_wiki.png">](https://en.wikipedia.org/wiki/Sorting_algorithm#Stability)**.
#
# Sorting is an important topic in programming, and we refer to the official [HOWTO <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/howto/sorting.html) for a more comprehensive introduction.
# + slideshow={"slide_type": "fragment"}
names
# + [markdown] slideshow={"slide_type": "skip"}
# `sort(reverse=True)` is different from the `reverse()` method. Whereas the former applies some sorting rule in reverse order, the latter simply reverses the elements in a `list` object.
# + slideshow={"slide_type": "slide"}
names.reverse()
# + slideshow={"slide_type": "fragment"}
names
# + [markdown] slideshow={"slide_type": "skip"}
# The `pop()` method removes the *last* element from a `list` object *and* returns it. Below we **capture** the `removed` element to show that the return value is not `None` as with all the methods introduced so far.
# + slideshow={"slide_type": "slide"}
removed = names.pop()
# + slideshow={"slide_type": "fragment"}
removed
# + slideshow={"slide_type": "fragment"}
names
# + [markdown] slideshow={"slide_type": "skip"}
# `pop()` takes an optional *index* argument and removes that instead.
#
# So, to remove the second element, `"Eckhardt"`, from `names`, we write this.
# + slideshow={"slide_type": "fragment"}
removed = names.pop(1)
# + slideshow={"slide_type": "fragment"}
removed
# + slideshow={"slide_type": "fragment"}
names
# + [markdown] slideshow={"slide_type": "skip"}
# Instead of removing an element by its index, we can also remove it by its value with the `remove()` method. Behind the scenes, Python then compares the object to be removed, `"Peter"` in the example, sequentially to each element with the `==` operator and removes the *first* one that evaluates equal to it. `remove()` does *not* return the removed element.
# + slideshow={"slide_type": "slide"}
names.remove("Peter")
# + slideshow={"slide_type": "fragment"}
names
# + [markdown] slideshow={"slide_type": "skip"}
# Also, `remove()` raises a `ValueError` if the value is not found.
# + slideshow={"slide_type": "skip"}
names.remove("Peter")
# + [markdown] slideshow={"slide_type": "skip"}
# `list` objects implement an `index()` method that returns the position of the first element that compares equal to its argument. It fails *loudly* with a `ValueError` if no element compares equal.
# + slideshow={"slide_type": "slide"}
names
# + slideshow={"slide_type": "fragment"}
names.index("Oliver")
# + slideshow={"slide_type": "fragment"}
names.index("Karl")
# + [markdown] slideshow={"slide_type": "skip"}
# The `count()` method returns the number of elements that compare equal to its argument.
# + slideshow={"slide_type": "fragment"}
names.count("Carl")
# + slideshow={"slide_type": "skip"}
names.count("Karl")
# + [markdown] slideshow={"slide_type": "skip"}
# Two more methods, `copy()` and `clear()`, are *syntactic sugar* and replace working with slices.
#
# `copy()` creates a *shallow* copy. So, `names.copy()` below does the same as taking a full slice with `names[:]`, and the caveats from above apply, too.
# + slideshow={"slide_type": "skip"}
names_copy = names.copy()
# + slideshow={"slide_type": "skip"}
names_copy
# + [markdown] slideshow={"slide_type": "skip"}
# `clear()` removes all references from a `list` object. So, `names_copy.clear()` is the same as `del names_copy[:]`.
# + slideshow={"slide_type": "skip"}
names_copy.clear()
# + slideshow={"slide_type": "skip"}
names_copy
# + [markdown] slideshow={"slide_type": "skip"}
# Many methods introduced in this section are mentioned in the [collections.abc <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/collections.abc.html) module's documentation as well: While the `index()` and `count()` methods come with any data type that is a `Sequence`, the `append()`, `extend()`, `insert()`, `reverse()`, `pop()`, and `remove()` methods are part of any `MutableSequence` type. The `sort()`, `copy()`, and `clear()` methods are `list`-specific.
#
# So, being a sequence does not only imply the four *behaviors* specified above, but also means that a data type comes with certain standardized methods.
# + [markdown] slideshow={"slide_type": "slide"}
# ### List Operations
# + [markdown] slideshow={"slide_type": "skip"}
# As with `str` objects, the `+` and `*` operators are overloaded for concatenation and always return a *new* `list` object. The references in this newly created `list` object reference the *same* objects as the two original `list` objects. So, the same caveat as with *shallow* copies from above applies!
# + slideshow={"slide_type": "slide"}
names
# + slideshow={"slide_type": "fragment"}
names + ["Diedrich", "Yves"]
# + slideshow={"slide_type": "fragment"}
2 * names
# + [markdown] slideshow={"slide_type": "skip"}
# Besides being an operator, the `*` symbol has a second syntactical use, as explained in [PEP 3132 <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://www.python.org/dev/peps/pep-3132/) and [PEP 448 <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://www.python.org/dev/peps/pep-0448/): It implements what is called **iterable unpacking**. It is *not* an operator syntactically but a notation that Python reads as a literal.
#
# In the example, Python interprets the expression as if the elements of the iterable `names` were placed between `"Achim"` and `"Xavier"` one by one. So, we do not obtain a nested but a *flat* list.
# + slideshow={"slide_type": "slide"}
["Achim", *names, "Xavier"]
# + [markdown] slideshow={"slide_type": "skip"}
# Effectively, Python reads that as if we wrote the following.
# + slideshow={"slide_type": "fragment"}
["Achim", names[0], names[1], names[2], "Xavier"]
# + [markdown] slideshow={"slide_type": "slide"}
# #### List Comparison
# + [markdown] slideshow={"slide_type": "skip"}
# The relational operators also work with `list` objects; yet another example of operator overloading.
#
# Comparison is made in a pairwise fashion until the first pair of elements does not evaluate equal or one of the `list` objects ends. The exact comparison rules depend on the elements and not the `list` objects. As with [sort() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/stdtypes.html#list.sort) or [sorted() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#sorted) above, comparison is *delegated* to the objects to be compared, and Python "asks" the elements in the two `list` objects to compare themselves. Usually, all elements are of the *same* type, and comparison is straightforward.
# + slideshow={"slide_type": "slide"}
names
# + slideshow={"slide_type": "fragment"}
names == ["Berthold", "Oliver", "Carl"]
# + slideshow={"slide_type": "fragment"}
names != ["Berthold", "Oliver", "Karl"]
# + slideshow={"slide_type": "fragment"}
names < ["Berthold", "Oliver", "Karl"]
# + [markdown] slideshow={"slide_type": "skip"}
# If two `list` objects have a different number of elements and all overlapping elements compare equal, the shorter `list` object is considered "smaller."
# + slideshow={"slide_type": "fragment"}
["Berthold", "Oliver"] < names < ["Berthold", "Oliver", "Carl", "Xavier"]
# + [markdown] slideshow={"slide_type": "slide"}
# ### Modifiers vs. Pure Functions
# + [markdown] slideshow={"slide_type": "skip"}
# As `list` objects are mutable, the caller of a function can see the changes made to a `list` object passed to the function as an argument. That is often a surprising *side effect* and should be avoided.
#
# As an example, consider the `add_xyz()` function.
# + slideshow={"slide_type": "slide"}
letters = ["a", "b", "c"]
# + slideshow={"slide_type": "fragment"}
def add_xyz(arg):
"""Append letters to a list."""
arg.extend(["x", "y", "z"])
return arg
# + [markdown] slideshow={"slide_type": "skip"}
# While this function is being executed, two variables, namely `letters` in the global scope and `arg` inside the function's local scope, reference the *same* `list` object in memory. Furthermore, the passed in `arg` is also the return value.
#
# So, after the function call, `letters_with_xyz` and `letters` are **aliases** as well, referencing the *same* object. We can also visualize that with [PythonTutor <img height="12" style="display: inline-block" src="static/link_to_py.png">](http://pythontutor.com/visualize.html#code=letters%20%3D%20%5B%22a%22,%20%22b%22,%20%22c%22%5D%0A%0Adef%20add_xyz%28arg%29%3A%0A%20%20%20%20arg.extend%28%5B%22x%22,%20%22y%22,%20%22z%22%5D%29%0A%20%20%20%20return%20arg%0A%0Aletters_with_xyz%20%3D%20add_xyz%28letters%29&cumulative=false&curInstr=0&heapPrimitives=nevernest&mode=display&origin=opt-frontend.js&py=3&rawInputLstJSON=%5B%5D&textReferences=false).
# + slideshow={"slide_type": "fragment"}
letters_with_xyz = add_xyz(letters)
# + slideshow={"slide_type": "fragment"}
letters_with_xyz
# + slideshow={"slide_type": "fragment"}
letters
# + [markdown] slideshow={"slide_type": "skip"}
# A better practice is to first create a copy of `arg` within the function that is then modified and returned. If we are sure that `arg` contains immutable elements only, we get away with a shallow copy. The downside of this approach is the higher amount of memory necessary.
#
# The revised `add_xyz()` function below is more natural to reason about as it does *not* modify the passed in `arg` internally. [PythonTutor <img height="12" style="display: inline-block" src="static/link_to_py.png">](http://pythontutor.com/visualize.html#code=letters%20%3D%20%5B%22a%22,%20%22b%22,%20%22c%22%5D%0A%0Adef%20add_xyz%28arg%29%3A%0A%20%20%20%20new_arg%20%3D%20arg%5B%3A%5D%0A%20%20%20%20new_arg.extend%28%5B%22x%22,%20%22y%22,%20%22z%22%5D%29%0A%20%20%20%20return%20new_arg%0A%0Aletters_with_xyz%20%3D%20add_xyz%28letters%29&cumulative=false&curInstr=0&heapPrimitives=nevernest&mode=display&origin=opt-frontend.js&py=3&rawInputLstJSON=%5B%5D&textReferences=false) shows that as well. This approach is following the **[functional programming <img height="12" style="display: inline-block" src="static/link_to_wiki.png">](https://en.wikipedia.org/wiki/Functional_programming)** paradigm that is going through a "renaissance" currently. Two essential characteristics of functional programming are that a function *never* changes its inputs and *always* returns the same output given the same inputs.
#
# For a beginner, it is probably better to stick to this idea and not change any arguments as the original `add_xyz()` above. However, functions that modify and return the argument passed in are an important aspect of object-oriented programming, as explained in [Chapter 10 <img height="12" style="display: inline-block" src="static/link_to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/10_classes_00_content.ipynb).
# + slideshow={"slide_type": "slide"}
letters = ["a", "b", "c"]
# + slideshow={"slide_type": "fragment"}
def add_xyz(arg):
"""Create a new list from an existing one."""
new_arg = arg[:]
new_arg.extend(["x", "y", "z"])
return new_arg
# + slideshow={"slide_type": "fragment"}
letters_with_xyz = add_xyz(letters)
# + slideshow={"slide_type": "fragment"}
letters_with_xyz
# + slideshow={"slide_type": "fragment"}
letters
# + [markdown] slideshow={"slide_type": "skip"}
# If we want to modify the argument passed in, it is best to return `None` and not `arg`, as does the final version of `add_xyz()` below. Then, the user of our function cannot accidentally create two aliases to the same object. That is also why the list methods above all return `None`. [PythonTutor <img height="12" style="display: inline-block" src="static/link_to_py.png">](http://pythontutor.com/visualize.html#code=letters%20%3D%20%5B%22a%22,%20%22b%22,%20%22c%22%5D%0A%0Adef%20add_xyz%28arg%29%3A%0A%20%20%20%20arg.extend%28%5B%22x%22,%20%22y%22,%20%22z%22%5D%29%0A%20%20%20%20return%0A%0Aadd_xyz%28letters%29&cumulative=false&curInstr=0&heapPrimitives=nevernest&mode=display&origin=opt-frontend.js&py=3&rawInputLstJSON=%5B%5D&textReferences=false) shows how there is only *one* reference to `letters` after the function call.
# + slideshow={"slide_type": "slide"}
letters = ["a", "b", "c"]
# + slideshow={"slide_type": "fragment"}
def add_xyz(arg):
"""Append letters to a list."""
arg.extend(["x", "y", "z"])
return # None
# + slideshow={"slide_type": "fragment"}
add_xyz(letters)
# + slideshow={"slide_type": "fragment"}
letters
# + [markdown] slideshow={"slide_type": "skip"}
# If we call `add_xyz()` with `letters` as the argument again, we end up with an even longer `list` object.
# + slideshow={"slide_type": "skip"}
add_xyz(letters)
# + slideshow={"slide_type": "skip"}
letters
# + [markdown] slideshow={"slide_type": "skip"}
# Functions that only work on the argument passed in are called **modifiers**. Their primary purpose is to change the **state** of the argument. On the contrary, functions that have *no* side effects on the arguments are said to be **pure**.
# + [markdown] slideshow={"slide_type": "slide"}
# ## The `tuple` Type
# + [markdown] slideshow={"slide_type": "skip"}
# To create a `tuple` object, we can use the same literal notation as for `list` objects *without* the brackets and list all elements.
# + slideshow={"slide_type": "slide"}
numbers = 7, 11, 8, 5, 3, 12, 2, 6, 9, 10, 1, 4
# + slideshow={"slide_type": "fragment"}
numbers
# + [markdown] slideshow={"slide_type": "skip"}
# However, to be clearer, many Pythonistas write out the optional parentheses `(` and `)`.
# + slideshow={"slide_type": "skip"}
numbers = (7, 11, 8, 5, 3, 12, 2, 6, 9, 10, 1, 4)
# + slideshow={"slide_type": "skip"}
numbers
# + [markdown] slideshow={"slide_type": "skip"}
# As before, `numbers` is an object on its own.
# + slideshow={"slide_type": "fragment"}
id(numbers)
# + slideshow={"slide_type": "fragment"}
type(numbers)
# + [markdown] slideshow={"slide_type": "skip"}
# While we could use empty parentheses `()` to create an empty `tuple` object ...
# + slideshow={"slide_type": "skip"}
empty_tuple = ()
# + slideshow={"slide_type": "skip"}
empty_tuple
# + slideshow={"slide_type": "skip"}
type(empty_tuple)
# + [markdown] slideshow={"slide_type": "skip"}
# ... we must use a *trailing comma* to create a `tuple` object holding one element. If we forget the comma, the parentheses are interpreted as the grouping operator and effectively useless!
# + slideshow={"slide_type": "skip"}
one_tuple = (1,) # we could ommit the parentheses but not the comma
# + slideshow={"slide_type": "skip"}
one_tuple
# + slideshow={"slide_type": "skip"}
type(one_tuple)
# + slideshow={"slide_type": "skip"}
no_tuple = (1)
# + slideshow={"slide_type": "skip"}
no_tuple
# + slideshow={"slide_type": "skip"}
type(no_tuple)
# + [markdown] slideshow={"slide_type": "skip"}
# Alternatively, we may use the [tuple() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#func-tuple) built-in that takes any iterable as its argument and creates a new `tuple` from its elements.
# + slideshow={"slide_type": "slide"}
tuple([1])
# + slideshow={"slide_type": "fragment"}
tuple("iterable")
# + [markdown] slideshow={"slide_type": "slide"}
# ### Tuples are like "Immutable Lists"
# + [markdown] slideshow={"slide_type": "skip"}
# Most operations involving `tuple` objects work in the same way as with `list` objects. The main difference is that `tuple` objects are *immutable*. So, if our program does not depend on mutability, we may and should use `tuple` and not `list` objects to model sequential data. That way, we avoid the pitfalls seen above.
#
# `tuple` objects are *sequences* exhibiting the familiar *four* behaviors. So, `numbers` holds a *finite* number of elements ...
# + slideshow={"slide_type": "slide"}
len(numbers)
# + [markdown] slideshow={"slide_type": "skip"}
# ... that we can obtain individually by looping over it in a predictable *forward* or *reverse* order.
# + slideshow={"slide_type": "fragment"}
for number in numbers:
print(number, end=" ")
# + slideshow={"slide_type": "fragment"}
for number in reversed(numbers):
print(number, end=" ")
# + [markdown] slideshow={"slide_type": "skip"}
# To check if a given object is *contained* in `numbers`, we use the `in` operator and conduct a linear search.
# + slideshow={"slide_type": "fragment"}
0 in numbers
# + slideshow={"slide_type": "skip"}
1 in numbers
# + slideshow={"slide_type": "skip"}
1.0 in numbers # in relies on == behind the scenes
# + [markdown] slideshow={"slide_type": "skip"}
# We may index and slice with the `[]` operator. The latter returns *new* `tuple` objects.
# + slideshow={"slide_type": "slide"}
numbers[0]
# + slideshow={"slide_type": "skip"}
numbers[-1]
# + slideshow={"slide_type": "fragment"}
numbers[6:]
# + [markdown] slideshow={"slide_type": "skip"}
# Index assignment does *not* work as tuples are *immutable* and results in a `TypeError`.
# + slideshow={"slide_type": "fragment"}
numbers[-1] = 99
# + [markdown] slideshow={"slide_type": "skip"}
# The `+` and `*` operators work with `tuple` objects as well: They always create *new* `tuple` objects.
# + slideshow={"slide_type": "skip"}
numbers + (99,)
# + slideshow={"slide_type": "skip"}
2 * numbers
# + [markdown] slideshow={"slide_type": "skip"}
# Being immutable, `tuple` objects only provide the `count()` and `index()` methods of `Sequence` types. The `append()`, `extend()`, `insert()`, `reverse()`, `pop()`, and `remove()` methods of `MutableSequence` types are *not* available. The same holds for the `list`-specific methods `sort()`, `copy()`, and `clear()`.
# + slideshow={"slide_type": "skip"}
numbers.count(0)
# + slideshow={"slide_type": "skip"}
numbers.index(1)
# + [markdown] slideshow={"slide_type": "skip"}
# The relational operators work in the *same* way as for `list` objects.
# + slideshow={"slide_type": "skip"}
numbers
# + slideshow={"slide_type": "skip"}
numbers == (7, 11, 8, 5, 3, 12, 2, 6, 9, 10, 1, 4)
# + slideshow={"slide_type": "skip"}
numbers != (99, 11, 8, 5, 3, 12, 2, 6, 9, 10, 1, 4)
# + slideshow={"slide_type": "skip"}
numbers < (99, 11, 8, 5, 3, 12, 2, 6, 9, 10, 1, 4)
# + [markdown] slideshow={"slide_type": "skip"}
# While `tuple` objects are immutable, this only relates to the references they hold. If a `tuple` object contains references to mutable objects, the entire nested structure is *not* immutable as a whole!
#
# Consider the following stylized example `not_immutable`: It contains *three* elements, `1`, `[2, ..., 11]`, and `12`, and the elements of the nested `list` object may be changed. While it is not practical to mix data types in a `tuple` object that is used as an "immutable list," we want to make the point that the mere usage of the `tuple` type does *not* guarantee a nested object to be immutable as a whole.
# + slideshow={"slide_type": "skip"}
not_immutable = (1, [2, 3, 4, 5, 6, 7, 8, 9, 10, 11], 12)
# + slideshow={"slide_type": "skip"}
not_immutable
# + slideshow={"slide_type": "skip"}
not_immutable[1][:] = [99, 99, 99]
# + slideshow={"slide_type": "skip"}
not_immutable
# + [markdown] slideshow={"slide_type": "slide"}
# ### Packing & Unpacking
# + [markdown] slideshow={"slide_type": "skip"}
# In the "*List Operations*" section above, the `*` symbol **unpacks** the elements of a `list` object into another one. This idea of *iterable unpacking* is built into Python at various places, even *without* the `*` symbol.
#
# For example, we may write variables on the left-hand side of a `=` statement in a `tuple` style. Then, any *finite* iterable on the right-hand side is unpacked. So, `numbers` is unpacked into *twelve* variables below.
# + slideshow={"slide_type": "slide"}
n1, n2, n3, n4, n5, n6, n7, n8, n9, n10, n11, n12 = numbers
# + slideshow={"slide_type": "fragment"}
n1
# + slideshow={"slide_type": "fragment"}
n2
# + slideshow={"slide_type": "fragment"}
n3
# + [markdown] slideshow={"slide_type": "skip"}
# Having to type twelve variables on the left is already tedious. Furthermore, if the iterable on the right yields a number of elements *different* from the number of variables, we get a `ValueError`.
# + slideshow={"slide_type": "skip"}
n1, n2, n3, n4, n5, n6, n7, n8, n9, n10, n11 = numbers
# + slideshow={"slide_type": "skip"}
n1, n2, n3, n4, n5, n6, n7, n8, n9, n10, n11, n12, n13 = numbers
# + [markdown] slideshow={"slide_type": "skip"}
# So, to make iterable unpacking useful, we prepend the `*` symbol to *one* of the variables on the left: That variable then becomes a `list` object holding the elements not captured by the other variables. We say that the excess elements from the iterable are **packed** into this variable.
#
# For example, let's get the `first` and `last` element of `numbers` and collect the rest in `middle`.
# + slideshow={"slide_type": "slide"}
first, *middle, last = numbers
# + slideshow={"slide_type": "fragment"}
first
# + slideshow={"slide_type": "fragment"}
middle # always a list!
# + slideshow={"slide_type": "fragment"}
last
# + [markdown] slideshow={"slide_type": "skip"}
# We already used unpacking before this section without knowing it. Whenever we write a `for`-loop over the [zip() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#zip) built-in, that generates a new `tuple` object in each iteration, which we unpack by listing several loop variables.
#
# So, the `name, position` acts like a left-hand side of an `=` statement and unpacks the `tuple` objects generated from "zipping" the `names` list and the `positions` tuple together.
# + slideshow={"slide_type": "skip"}
positions = ("goalkeeper", "defender", "midfielder", "striker", "coach")
# + slideshow={"slide_type": "skip"}
for name, position in zip(names, positions):
print(name, "is a", position)
# + [markdown] slideshow={"slide_type": "skip"}
# Without unpacking, [zip() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#zip) generates a series of `tuple` objects.
# + slideshow={"slide_type": "skip"}
for pair in zip(names, positions):
print(type(pair), pair, sep=" ")
# + [markdown] slideshow={"slide_type": "skip"}
# Unpacking also works for nested objects. Below, we wrap [zip() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#zip) with the [enumerate() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/functions.html#enumerate) built-in to have an index variable `number` inside the `for`-loop. In each iteration, a `tuple` object consisting of `number` and another `tuple` object is created. The inner one then holds the `name` and `position`.
# + slideshow={"slide_type": "skip"}
for number, (name, position) in enumerate(zip(names, positions), start=1):
print(f"{name} (jersey #{number}) is a {position}")
# + [markdown] slideshow={"slide_type": "slide"}
# #### Swapping Variables
# + [markdown] slideshow={"slide_type": "skip"}
# A popular use case of unpacking is **swapping** two variables.
#
# Consider `a` and `b` below.
# + slideshow={"slide_type": "slide"}
a = 0
b = 1
# + [markdown] slideshow={"slide_type": "skip"}
# Without unpacking, we must use a temporary variable `temp` to swap `a` and `b`.
# + slideshow={"slide_type": "fragment"}
temp = a
a = b
b = temp
del temp
# + slideshow={"slide_type": "fragment"}
a
# + slideshow={"slide_type": "fragment"}
b
# + [markdown] slideshow={"slide_type": "skip"}
# With unpacking, the solution is more elegant, and also a bit faster as well. *All* expressions on the right-hand side are evaluated *before* any assignment takes place.
# + slideshow={"slide_type": "slide"}
a, b = 0, 1
# + slideshow={"slide_type": "fragment"}
a, b = b, a
# + slideshow={"slide_type": "fragment"}
a, b
# + [markdown] slideshow={"slide_type": "slide"}
# ##### Example: [Fibonacci Numbers <img height="12" style="display: inline-block" src="static/link_to_wiki.png">](https://en.wikipedia.org/wiki/Fibonacci_number) (revisited)
# + [markdown] slideshow={"slide_type": "skip"}
# Unpacking allows us to rewrite the iterative `fibonacci()` function from [Chapter 4 <img height="12" style="display: inline-block" src="static/link_to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/04_iteration_00_content.ipynb#"Hard-at-first-Glance"-Example:-Fibonacci-Numbers-%28revisited%29) in a concise way.
# + slideshow={"slide_type": "slide"}
def fibonacci(i):
"""Calculate the ith Fibonacci number.
Args:
i (int): index of the Fibonacci number to calculate
Returns:
ith_fibonacci (int)
"""
a, b = 0, 1
for _ in range(i - 1):
a, b = b, a + b
return b
# + slideshow={"slide_type": "fragment"}
fibonacci(12)
# + [markdown] slideshow={"slide_type": "slide"}
# #### Function Definitions & Calls
# + [markdown] slideshow={"slide_type": "skip"}
# The concepts of packing and unpacking are also helpful when writing and using functions.
#
# For example, let's look at the `product()` function below. Its implementation suggests that `args` must be a sequence type. Otherwise, it would not make sense to index into it with `[0]` or take a slice with `[1:]`. In line with the function's name, the `for`-loop multiplies all elements of the `args` sequence. So, what does the `*` do in the header line, and what is the exact data type of `args`?
#
# The `*` is again *not* an operator in this context but a special syntax that makes Python *pack* all *positional* arguments passed to `product()` into a single `tuple` object called `args`.
# + slideshow={"slide_type": "slide"}
def product(*args):
"""Multiply all arguments."""
result = args[0]
for arg in args[1:]:
result *= arg
return result
# + [markdown] slideshow={"slide_type": "skip"}
# So, we can pass an *arbitrary* (i.e., also none) number of *positional* arguments to `product()`.
#
# The product of just one number is the number itself.
# + slideshow={"slide_type": "fragment"}
product(42)
# + [markdown] slideshow={"slide_type": "skip"}
# Passing in several numbers works as expected.
# + slideshow={"slide_type": "fragment"}
product(2, 5, 10)
# + [markdown] slideshow={"slide_type": "skip"}
# However, this implementation of `product()` needs *at least* one argument passed in due to the expression `args[0]` used internally. Otherwise, we see a *runtime* error, namely an `IndexError`. We emphasize that this error is *not* caused in the header line.
# + slideshow={"slide_type": "skip"}
product()
# + [markdown] slideshow={"slide_type": "skip"}
# Another downside of this implementation is that we can easily generate *semantic* errors: For example, if we pass in an iterable object like the `one_hundred` list, *no* exception is raised. However, the return value is also not a numeric object as we expect. The reason for this is that during the function call, `args` becomes a `tuple` object holding *one* element, which is `one_hundred`, a `list` object. So, we created a nested structure by accident.
# + slideshow={"slide_type": "slide"}
one_hundred = [2, 5, 10]
# + slideshow={"slide_type": "fragment"}
product(one_hundred) # a semantic error!
# + [markdown] slideshow={"slide_type": "skip"}
# This error does not occur if we unpack `one_hundred` upon passing it as the argument.
# + slideshow={"slide_type": "fragment"}
product(*one_hundred)
# + [markdown] slideshow={"slide_type": "skip"}
# That is the equivalent of writing out the following tedious expression. Yet, that does *not* scale for iterables with many elements in them.
# + slideshow={"slide_type": "fragment"}
product(one_hundred[0], one_hundred[1], one_hundred[2])
# + [markdown] slideshow={"slide_type": "skip"}
# In the "*Packing & Unpacking with Functions*" [exercise <img height="12" style="display: inline-block" src="static/link_to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/07_sequences_02_exercises.ipynb#Packing-&-Unpacking-with-Functions) at the end of this chapter, we look at `product()` in more detail.
#
# While we needed to unpack `one_hundred` above to avoid the semantic error, unpacking an argument in a function call may also be a convenience in general. For example, to print the elements of `one_hundred` in one line, we need to use a `for` statement, until now. With unpacking, we get away *without* a loop.
# + slideshow={"slide_type": "skip"}
print(one_hundred) # prints the tuple; we do not want that
# + slideshow={"slide_type": "skip"}
for number in one_hundred:
print(number, end=" ")
# + slideshow={"slide_type": "skip"}
print(*one_hundred) # replaces the for-loop
# + [markdown] slideshow={"slide_type": "slide"}
# ### The `namedtuple` Type
# + [markdown] slideshow={"slide_type": "skip"}
# Above, we proposed the idea that `tuple` objects are like "immutable lists." Often, however, we use `tuple` objects to represent a **record** of related **fields**. Then, each element has a *semantic* meaning (i.e., a descriptive name).
#
# As an example, think of a spreadsheet with information on students in a course. Each row represents a record and holds all the data associated with an individual student. The columns (e.g., matriculation number, first name, last name) are the fields that may come as *different* data types (e.g., `int` for the matriculation number, `str` for the names).
#
# A simple way of modeling a single student is as a `tuple` object, for example, `(123456, "John", "Doe")`. A disadvantage of this approach is that we must remember the order and meaning of the elements/fields in the `tuple` object.
#
# An example from a different domain is the representation of $(x, y)$-points in the $x$-$y$-plane. Again, we could use a `tuple` object like `current_position` below to model the point $(4, 2)$.
# + slideshow={"slide_type": "slide"}
current_position = (4, 2)
# + [markdown] slideshow={"slide_type": "skip"}
# We implicitly assume that the first element represents the $x$ and the second the $y$ coordinate. While that follows intuitively from convention in math, we should at least add comments somewhere in the code to document this assumption.
#
# A better way is to create a *custom* data type. While that is covered in depth in [Chapter 10 <img height="12" style="display: inline-block" src="static/link_to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/10_classes_00_content.ipynb), the [collections <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/collections.html) module in the [standard library <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/index.html) provides a [namedtuple() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/collections.html#collections.namedtuple) **factory function** that creates "simple" custom data types on top of the standard `tuple` type.
# + slideshow={"slide_type": "slide"}
from collections import namedtuple
# + [markdown] slideshow={"slide_type": "skip"}
# [namedtuple() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/collections.html#collections.namedtuple) takes two arguments. The first argument is the name of the data type. That could be different from the variable `Point` we use to refer to the new type, but in most cases it is best to keep them in sync. The second argument is a sequence with the field names as `str` objects. The names' order corresponds to the one assumed in `current_position`.
# + slideshow={"slide_type": "fragment"}
Point = namedtuple("Point", ["x", "y"])
# + [markdown] slideshow={"slide_type": "skip"}
# The `Point` object is a so-called **class**. That is what it means if an object is of type `type`. It can be used as a **factory** to create *new* `tuple`-like objects of type `Point`. In a way, [namedtuple() <img height="12" style="display: inline-block" src="static/link_to_py.png">](https://docs.python.org/3/library/collections.html#collections.namedtuple) gives us a way to create our own custom **constructors**.
# + slideshow={"slide_type": "fragment"}
id(Point)
# + slideshow={"slide_type": "fragment"}
type(Point)
# + [markdown] slideshow={"slide_type": "skip"}
# The value of `Point` is just itself in a *literal notation*.
# + slideshow={"slide_type": "fragment"}
Point
# + [markdown] slideshow={"slide_type": "skip"}
# We write `Point(4, 2)` to create a *new* object of type `Point`.
# + slideshow={"slide_type": "slide"}
current_position = Point(4, 2)
# + [markdown] slideshow={"slide_type": "skip"}
# Now, `current_position` has a somewhat nicer representation. In particular, the coordinates are named `x` and `y`.
# + slideshow={"slide_type": "fragment"}
current_position
# + [markdown] slideshow={"slide_type": "skip"}
# It is *not* a `tuple` any more but an object of type `Point`.
# + slideshow={"slide_type": "fragment"}
id(current_position)
# + slideshow={"slide_type": "fragment"}
type(current_position)
# + [markdown] slideshow={"slide_type": "skip"}
# We use the dot operator `.` to access the defined attributes.
# + slideshow={"slide_type": "slide"}
current_position.x
# + slideshow={"slide_type": "fragment"}
current_position.y
# + [markdown] slideshow={"slide_type": "skip"}
# As before, we get an `AttributeError` if we try to access an undefined attribute.
# + slideshow={"slide_type": "skip"}
current_position.z
# + [markdown] slideshow={"slide_type": "skip"}
# `current_position` continues to work like a `tuple` object! That is why we can use `namedtuple` as a replacement for `tuple`. The underlying implementations exhibit the *same* computational efficiencies and memory usages.
#
# For example, we can index into or loop over `current_position` as it is still a sequence with the familiar four properties.
# + slideshow={"slide_type": "slide"}
current_position[0]
# + slideshow={"slide_type": "skip"}
current_position[1]
# + slideshow={"slide_type": "fragment"}
for number in current_position:
print(number)
# + slideshow={"slide_type": "fragment"}
for number in reversed(current_position):
print(number)
# + [markdown] slideshow={"slide_type": "skip"}
# Because of that it has "length."
# + slideshow={"slide_type": "fragment"}
len(current_position)
# + [markdown] slideshow={"slide_type": "skip"}
# ## TL;DR
# + [markdown] slideshow={"slide_type": "skip"}
# **Sequences** are an *abstract* concept that summarizes *four* behaviors an object may or may not exhibit. Sequences are
# - **finite** and
# - **ordered**
# - **containers** that we may
# - **loop over**.
#
# Examples are the `list`, `tuple`, but also the `str` types.
#
# Objects that exhibit all behaviors *except* being ordered are referred to as **collections**.
#
# The objects inside a sequence are called its **elements** and may be labeled with a unique **index**, an `int` object in the range $0 \leq \text{index} < \lvert \text{sequence} \rvert$.
#
# `list` objects are **mutable**. That means we can change the references to the other objects it contains, and, in particular, re-assign them.
#
# On the contrary, `tuple` objects are like **immutable** lists: We can use them in place of any `list` object as long as we do *not* need to mutate it. Often, `tuple` objects are also used to model **records** of related **fields**.
# + [markdown] slideshow={"slide_type": "skip"}
# ## Further Resources
# + [markdown] slideshow={"slide_type": "skip"}
# A lecture-style **video presentation** of this chapter is integrated below (cf., the [video <img height="12" style="display: inline-block" src="static/link_to_yt.png">](https://www.youtube.com/watch?v=nx2sCDoeC3I&list=PL-2JV1G3J10lQ2xokyQowcRJI5jjNfW7f) or the entire [playlist <img height="12" style="display: inline-block" src="static/link_to_yt.png">](https://www.youtube.com/playlist?list=PL-2JV1G3J10lQ2xokyQowcRJI5jjNfW7f)).
# + slideshow={"slide_type": "skip"}
from IPython.display import YouTubeVideo
YouTubeVideo("nx2sCDoeC3I", width="60%")
| 07_sequences_00_content.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import json
import sys
import time
from sys import platform
from login import s, _print
dic = {}
file_index = 0
def upload_media(fpath, is_img=False):
global file_index
if not os.path.exists(fpath):
_print('File not exists')
return None
url_1 = 'https://file.wx2.qq.com/cgi-bin/mmwebwx-bin/webwxuploadmedia?f=json'
url_2 = 'https://file2.wx2.qq.com/cgi-bin/mmwebwx-bin/webwxuploadmedia?f=json'
flen = str(os.path.getsize(fpath))
ftype = mimetypes.guess_type(fpath)[0] or 'application/octet-stream'
files = {
'id': (None, 'WU_FILE_%s' % str(file_index)),
'name': (None, os.path.basename(fpath)),
'type': (None, ftype),
'lastModifiedDate': (None, time.strftime('%m/%d/%Y, %H:%M:%S GMT+0800 (CST)')),
'size': (None, flen),
'mediatype': (None, 'pic' if is_img else 'doc'),
'uploadmediarequest': (None, json.dumps({
'BaseRequest': dic['BaseRequest'],
'ClientMediaId': int(time.time()),
'TotalLen': flen,
'StartPos': 0,
'DataLen': flen,
'MediaType': 4,
})),
'webwx_data_ticket': (None, s.cookies['webwx_data_ticket']),
'pass_ticket': (None, dic['pass_ticket']),
'filename': (os.path.basename(fpath), open(fpath, 'rb'),ftype.split('/')[1]),
}
file_index += 1
try:
r = s.post(url_1, files=files)
if json.loads(r.text)['BaseResponse']['Ret'] != 0:
# 当file返回值不为0时则为上传失败,尝试第二服务器上传
r = s.post(url_2, files=files)
if json.loads(r.text)['BaseResponse']['Ret'] != 0:
_print('Upload media failure.')
return None
mid = json.loads(r.text)['MediaId']
return mid
except Exception,e:
return None
def send_img(fpath, friend):
mid = upload_media(fpath, is_img=True)
if mid is None:
return False
url = dic['base_uri'] + '/webwxsendmsgimg?fun=async&f=json'
data = {
'BaseRequest': self.base_request,
'Msg': {
'Type': 3,
'MediaId': mid,
'FromUserName': dic['My']['UserName'],
'ToUserName': friend["UserName"].encode('unicode_escape'),
'LocalID': str(time.time() * 1e7),
'ClientMsgId': str(time.time() * 1e7), }, }
if fpath[-4:] == '.gif':
url = dic['base_uri'] + '/webwxsendemoticon?fun=sys'
data['Msg']['Type'] = 47
data['Msg']['EmojiFlag'] = 2
try:
r = s.post(url, data=json.dumps(data))
res = json.loads(r.text)
if res['BaseResponse']['Ret'] == 0:
return True
else:
return False
except Exception,e:
return False
def init():
global dic
try:
with open("./logininfo.log", 'r') as f:
_print('login info time:\t' +
f.readline()[:-1])
dic = f.readline()
dic = eval(dic)
except:
if 'linux' in platform:
path = '/home/stevi/all2wechat/logininfo.log'
else:
path = 'E:/Github/all2wechat/logininfo.log'
with open(path, 'r') as f:
_print('login info time:\t' +
f.readline()[:-1])
dic = f.readline()
dic = eval(dic)
def webwxgetcontact():
global dic
try:
with open("./contactlist.log", 'r') as f:
ContactList = f.readline()
ContactList = eval(ContactList)
dic['ContactList'] = ContactList
except:
pass
#todo
if 'linux' in platform:
path = '/home/stevi/all2wechat/logininfo.log'
else:
path = 'E:/Github/all2wechat/logininfo.log'
with open(path, 'r') as f:
_print('loading login data from '+path)
_print('login info time:\t' +
f.readline()[:-1])
dic = f.readline()
dic = eval(dic)
# _print('Getting contactlist')
# url = dic['base_uri'] + "/webwxgetcontact?r=" + str(int(
# time.time()))
# r = s.post(url, json={})
# content = r.text.encode('unicode_escape').decode('string_escape')
# ContactList = json.loads(content)['MemberList']
# dic['ContactList'] = ContactList
# with open('contactlist.log', 'w') as f:
# f.write(str(ContactList))
# _print('Contactlist get')
def main():
global dic
init()
webwxgetcontact()
name = sys.argv[1].decode('utf8')
for f in dic['ContactList']:
if f['RemarkName'] == name or f['NickName'] == name:
send_img('./image/ScreenClip.png', f)
# webwxsendmsg(f, sys.argv[2])
print('Send')
break
def webwxsendmsg(friend, content):
clientMsgId = str(int(time.time()))
url = dic['base_uri'] + \
"/webwxsendmsg?lang=zh_CN&pass_ticket=" + dic['pass_ticket']
Msg = {
'Type': '1',
'Content': content,
'ClientMsgId': clientMsgId.encode('unicode_escape'),
'FromUserName': dic['My']['UserName'].encode('unicode_escape'),
'ToUserName': friend["UserName"].encode('unicode_escape'),
'LocalID': clientMsgId.encode('unicode_escape')
}
payload = {'BaseRequest': dic['BaseRequest'], 'Msg': Msg}
headers = {'ContentType': 'application/json; charset=UTF-8'}
data = json.dumps(payload, ensure_ascii=False)
r = s.post(url, data=data, headers=headers)
resp = json.loads(r.text)
if 'BaseResponse' in resp:
if 'Ret' in resp['BaseResponse']:
return True
return False
if __name__ == '__main__':
main()
| sendimg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# filtered the source and target sentence with length <=70 and the length ratio between 1/3 to 3
def data_filter(en_path,de_path,o_en_path,o_de_path):
f_en = open(en_path,'r')
f_de = open(de_path,'r')
fout_en = open(o_en_path,'w')
fout_de = open(o_de_path,'w')
for l_en,l_de in zip(f_en,f_de):
len_en = len(l_en.strip().split())
len_de = len(l_de.strip().split())
if len_en<=70 and len_de<=70 and len_en>0 and len_de>0:
if (len_en>=len_de and 1.0*len_en/len_de<=3) or \
(len_en<len_de and 1.0*len_de/len_en<=3):
fout_en.write(l_en)
fout_de.write(l_de)
f_en.close()
f_de.close()
fout_en.close()
fout_de.close()
# 合并数据集,输入是en-de
def merge_file(src_file,tgt_file):
fin_src = open(src_file,'r')
fin_tgt = open(tgt_file,'r')
fout_src = open('./parallel_data/2017/train.en','a')
fout_tgt = open('./parallel_data/2017/train.de','a')
for line1,line2 in zip(fin_src,fin_tgt):
fout_src.write(line1.strip()+'\n')
fout_tgt.write(line2.strip()+'\n')
fin_src.close()
fin_tgt.close()
fout_src.close()
fout_tgt.close()
| preprocess/data_filter_merge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Y1stijQKkKzM"
#
# # Time Series with Prodes Dataset
# [Prodes dataset](https://data.globalforestwatch.org/datasets/gfw::prodes-deforestation-in-amazonia/about)
#
# By <NAME>
#
# ## Cleaning the data
#
# The Prodes data set contains many data points. From observing the dataset, most of the satellite images are taken from August - September, which are the months of the year that the Amazon is not mostly covered by clouds. It is during this time, that the satellite images can be taken for evaluation.
#
# Because the clouds obstruct the image of the satellite data, there are no consistent patterns for date ranges that the data is collected.
#
# To clean the data, the following functions drop all columns except the columns that hold values for States, date the images were taken, and the areakm.
# After looking at the satellite images, I came to the discovery that the areakm data points detailed the measurment of deforestation from the previous year. Therefore, after the data is cleaned, the cummulative sum is the value the time series forecast model will use.
#
# In order to prepare the dataset for the time series forecast model, the data must be prepared with a datetime index, as well as the data points to fit the date time index. Because of the inconsistant data points (due to cloud cover), the redisturbution method works as follows:
#
# 1. The data points for each specific state of each specific year are redistributed to fit 365 days.
# 2. The left over rows that did not fit into 365 days are summed, and divided amoung the 356 rows evenly. This means- each row value is added with the divided remaining value. This was to keep the integrity of the total amount of areakm that was deforested, and to prepare the dataset for yearly forecasting.
# 3. Due to the method that it was distubuted, no daily, or monthly trends should be observed. Only yearly.
#
# + [markdown] id="Qs7VJW_tlEwm"
# ## Prepare environment
# ## Cleaning data
#
# + id="jlkCzlUmkC2f"
import pandas as pd
import numpy as np
from datetime import datetime
import matplotlib.pyplot as plt
import seaborn as sns
import statsmodels.api as sm
from statsmodels.tsa.seasonal import seasonal_decompose
from statsmodels.tsa.filters.hp_filter import hpfilter
from statsmodels.tsa.holtwinters import SimpleExpSmoothing
from statsmodels.tsa.holtwinters import ExponentialSmoothing
from statsmodels.tsa.stattools import acovf, acf, pacf, pacf_yw, pacf_ols
from pandas.plotting import lag_plot
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
from sklearn.metrics import mean_squared_error, mean_absolute_error
from statsmodels.tsa.ar_model import AR, ARResults
from statsmodels.tsa.stattools import adfuller
from statsmodels.tools.eval_measures import mse, rmse, meanabs
# !pip install pmdarima
from pmdarima import auto_arima
from statsmodels.tsa.statespace.sarimax import SARIMAX
from statsmodels.tsa.arima_model import ARIMA, ARMA, ARMAResults, ARIMAResults
# + [markdown] id="cxOu2_wFlaKG"
# - sum all the km for every year.
# - First, organize by year- collect dfs of each year.
# - second, get sum for each year.
#
# + id="DaLulz50lVdI"
df = pd.read_csv('PRODES_Deforestation_in_Amazonia.csv')
# + id="F-lvuct8lljj"
def df_clean(df_):
df_.dropna(inplace=True)
df_.drop(columns=['FID', 'ORIGIN_ID', 'PATH_ROW', 'DEF_CLOUD', 'SCENE_ID', 'PUBLISH_YE', 'SOURCE', 'SATELLITE', 'SENSOR', 'MAIN_CLASS', 'CLASS_NAME', 'JULIAN_DAY', 'ID'], inplace=True)
df_.sort_values(by='IMAGE_DATE', inplace=True)
df_['IMAGE_DATE'] = pd.to_datetime(df_['IMAGE_DATE'], format='%Y-%m-%d', errors='coerce')
df_.set_index(df_['IMAGE_DATE'], inplace=True)
df_.dropna(inplace=True)
org_yearly(df_)
# + id="lP5FtZV5lpxg"
def org_yearly(df_):
df_name = {}
for year in df_.index.year.unique():
print('DF for {}'.format(year))
df_name[year] = df_['{}'.format(year)]
print(df_name[year])
# + id="kUF8nBRslrqr"
df_clean(df)
# + [markdown] id="XOXL9FVml7Fj"
# ### Preparing for time models
# + [markdown] id="GuJYBEXel1nR"
# - resample yearly
# - Add cumsum()
# + id="OxmSLnjkl9rg"
df_y = df.resample('A').sum()
# + id="pcn9gMdimDLO"
df_y['cumsum'] = df_y.AREA_KM.cumsum()
# + id="aSfkJ2K5mDwf"
df_y['cumsum'].plot()
# + [markdown] id="4d12IlyCmPwP"
# ### Time Series
# + id="QPiIm1LGmN6p"
df_y.index
# + id="A0Cq0dGMmN9W"
train_yearly = df_y.iloc[:9]
test_yearly = df_y.iloc[8:]
# + id="2AgmwrRqmOCL"
fitted_model = ExponentialSmoothing(train_yearly['cumsum'], trend='mul', seasonal_periods=1).fit()
test_predictions = fitted_model.forecast(10)
# + id="godx4OrymOEa"
train_yearly['cumsum'].plot(figsize=(12,5), legend=True, label='Train')
test_yearly['cumsum'].plot(legend=True, label='Test')
test_predictions.plot(legend=True, label='Prediction')
plt.title('Predictions from 2017 - 2026 \nAreakm of total deforestation \n All 9 State of Legal Amazonia', fontsize=20)
plt.xlabel('Years', fontsize=12)
plt.ylabel('Areakm total deforestation', fontsize=12)
# + id="GoTi5C6MmOG5"
test_predictions
# + id="HSCmjmL_mdgz"
df_y['cumsum']
# + [markdown] id="vZOE36r2mh7n"
# - put text preditions in a dataframe to download and send to flask server
# <br>
# - also add non predictions in a csv
# + id="EQhoPFVYmdi1"
from google.colab import drive
drive.mount('/content/gdrive')
# + id="19VtF-iSmdlm"
from google.colab import files
# + id="sMsUCY9Bmpih"
recorded_areakm = df_y['cumsum'].to_frame()
recorded_areakm.reset_index(level=0, inplace=True)
recorded_areakm.columns=['Recorded Years', 'Recorded Areakm Deforested']
# + id="qMTcyWbCmplR"
RecordedAreakm = recorded_areakm.to_csv()
with open('RecordedAreakm.csv', 'a') as f:
f.write(RecordedAreakm)
files.download('RecordedAreakm.csv')
# + id="MTpW5fe_mpno"
test_predictions_df=test_predictions.to_frame()
test_predictions_df.reset_index(level=0, inplace=True)
test_predictions_df.columns=['Prediction Year', 'Predicted Areakm Deforested']
# + id="F0nbMFUCmppu"
Brazil_ESM = test_predictions_df.to_csv()
with open('BrazilESM.csv', 'a') as f:
f.write(Brazil_ESM)
# + id="VwL2no0xpGf4"
# + id="USJQIueympsJ"
files.download('BrazilESM.csv')
# + [markdown] id="Bd-q1QJsntT6"
#
# #### Failed model:
# + [markdown] id="sHVx4orVnQL-"
#
# + id="1RTtOG-rnRln"
result_y = seasonal_decompose(df_y['cumsum'], model='multiplicative')
# + id="xR9iz_C-oLhV"
result_y.plot();
# + id="bZMs4x8SoPXQ"
auto_arima(df_y['cumsum']).summary() #m is for the seasonal periods
#SARIMAX(0, 1, 0)
# + id="RcJuzn0FoPZs"
train_sy = df_y.iloc[:10]
test_sy = df_y.iloc[10:]
# + id="RZTfW1K4oPb3"
model_sy = SARIMAX(train_sy['cumsum'], order=(0,1,0))
result_sy = model_sy.fit()
start_sy = len(train_sy)
end_sy = len(train_sy) + len(test_sy) -1
# + id="oh6YbaTAoPeP"
prediction_sy = result_sy.predict(start_sy, end_sy, type='levels').rename('SARIMAX Yearly Predictions')
# + id="iKafpIXBoPqE"
test_sy['cumsum'].plot(legend= True, figsize=(12, 8))
prediction_sy.plot(legend=True)
# + [markdown] id="YE5FGdhWopju"
# #### Visualizations
# + [markdown] id="Kw1GYcrgqo8-"
# #### Rate of deforestation over time from 2008 - 2019
# + id="2AD2pVUyox8U"
df.median()
df.describe()
df['STATE'].value_counts()
# + id="gUSsXdglo9tP"
state_group = df.groupby(['STATE'])
state_group['AREA_KM'].agg(['median', 'mean', 'sum'])
# + id="463AMbp_pA6S"
state_group.get_group('AM')
# + id="VOAWB6RlpCqc"
group_year = df.groupby(['IMAGE_DATE'])
group_year.get_group(2008)
# + id="IZydpLRSpI4j"
group_year['STATE'].value_counts()
# + id="8xYMhmF6pLR8"
group_year['AREA_KM'].sum() #.plot
# + id="o3uHjsBZpNJZ"
grp = df.groupby(by=['YEAR', 'STATE'])
# + id="JGtMtN_wpNn_"
grp['AREA_KM'].sum()
# + id="gIuwvbxmpNqb"
grp['AREA_KM'].sum().plot.bar()
# + [markdown] id="Pc-GMlaeqRPj"
#
# - The above visualizes the years 2008-2019. It is not clear by the marks, so for readability, each group of peaks represents a year.
# - From this visualization, 2008 is extremley high, while trends dip from 2009 - 2017.
# - Discovering from research, Brazil saw a decline in deforestation rates from 2008 to around 2018.
# According to an article published November 2020 by [BBC](https://www.bbc.com/news/world-latin-america-55130304), in collaboration with Prodes data:
# > "Amazon deforestation highest since 2008"
#
# Why? [BBC](https://www.bbc.com/news/world-latin-america-55130304) states:
#
# > "Scientists say it has suffered losses at an accelerated rate since <NAME> took office in January 2019.
# > The Brazilian president has encouraged agriculture and mining activities in the world's largest rainforest."
# + [markdown] id="qi5vlZBWqufb"
# #### Rate of deforestation by state in total from 2008-2019
# + id="IZA1E-qipNsb"
state_area = df.groupby(by='STATE')['AREA_KM'].sum().sort_values(ascending=False).reset_index()
state_area = state_area.sort_values(by='AREA_KM', ascending=True)
# + [markdown] id="Y3rqH33drMQk"
# Next step is to figure out size of each state and find the percentage that it has been deforested, then to look again at the rates of reforestation
# + id="GdnC3u0PrI1P"
plt.figure(figsize=(12,8))
ax = sns.barplot(x=state_area['STATE'], y=state_area['AREA_KM'], palette='Greens', alpha=0.85)
plt.title("Deforestation by State", fontsize = 25)
plt.xlabel("State", fontsize = 20)
plt.ylabel("Sum of km2 deforestation", fontsize = 20)
plt.xticks(fontsize = 15)
plt.yticks(fontsize = 15)
plt.legend(fontsize = 15)
# + [markdown] id="74DBteu7sKiU"
# **A similar process can be seen under the Failed Methods section: "Visualization of total cumsum deforested per state"**
#
#
# + [markdown] id="rJjdrhLfsUi_"
# # Failed Methods
# + [markdown] id="f1zmyIBYsFKf"
# ## Ceaning the data
# **A way of cleaning data, before fully understanding the capabilites of pandas dataframe methods offered**...
# + id="7vjTTqFKrSd7"
df = pd.read_csv('PRODES_Deforestation_in_Amazonia.csv')
def df_clean(df_):
df_.dropna(inplace=True)
df_.drop(columns=['FID', 'ORIGIN_ID', 'PATH_ROW', 'DEF_CLOUD', 'SCENE_ID', 'PUBLISH_YE', 'SOURCE', 'SATELLITE', 'SENSOR', 'MAIN_CLASS', 'CLASS_NAME', 'JULIAN_DAY', 'ID'], inplace=True)
df_.sort_values(by='IMAGE_DATE', inplace=True)
org_state(df_) #next function
#organize the df to clean for processing.
# + id="i3dPp2_8rSfy"
#organize the df into states
#the individual df of indv. states will pass through to the next function, bringing the state name with it
def org_state(df_):
df_.dropna(inplace=True) #put it here for further caution
df_name = {}
for state in df_.STATE.unique():
df_name[state] = df_.loc[df_['STATE'] == state].copy()
print('********************', '\n', 'Information for state of {} :'.format(state), '\n', '********************', '\n')
clean_date(df_name[state], state) #next function
# + id="ZgiM1OjxrSiD"
#change the Image_date to datetime
#individual df of the individual year will go through to the next function
#here a date range is created as well, this is carried through to the next function
def clean_date(df_, state):
df_.dropna(inplace=True)
df_year = {}
state = state
#turn image_date to a time series to get information about it and seperate it
df_['IMAGE_DATE'] = pd.to_datetime(df_['IMAGE_DATE'], format='%Y-%m-%d', errors='coerce').copy()
#time to organize the datetimes into seperate dfs. for each year
df_.set_index(df_['IMAGE_DATE'], inplace=True)
df_.dropna(inplace=True)
for year in df_.index.year.unique():
print('*************** CLEANING FOR {}'.format(year), '\n')
year = int(year)
df_year[year] = df_.loc[df_.YEAR == year].copy()
global date_rng
date_rng = pd.date_range(start='{}-01-01'.format(year), end='{}-12-31'.format(year), freq='D') #int year to sove the float problem
#print('date range length: ', len(date_rng))
apply_new_dates(df_year[year], date_rng, year, state)
# + id="EsK1iQl0sqFN"
#the individual state/year df will be processed to disperse across the date_range.
def apply_new_dates(df_, date_ranges, year, state):
df_.dropna(inplace=True)
unique_df_id = '{}'.format(state) + '{}'.format(year)
global cleaned_date_list
cleaned_date_list = []
global popped_numbers
popped_numbers = []
df_Htime = {}
subtract = {}
Y = {}
series_area = df_['AREA_KM']
if(len(series_area) > len(date_ranges)):
subtract_by = len(series_area) - len(date_ranges)
#print(subtract_by)
poped_rows = series_area[len(date_ranges):] #all the rows that cant fit to date range
rows_hourly = series_area[:(len(date_ranges))] #all the rows that fit in date range
#average = sum(poped_rows)/len(poped_rows)
average = sum(poped_rows) #add all the values together
dispurse_number = average / len(rows_hourly) #divide by rows in the timerange #len(poped_rows)
lst_date_rng = date_rng.tolist()
hourly_list = rows_hourly.tolist()
df_Htime[unique_df_id] = pd.DataFrame({'date':lst_date_rng, 'areakm':hourly_list, 'state':state})
disperse_areakms(df_Htime[unique_df_id], dispurse_number) #next function to disperse the left over numbers
create_big_csv(df_Htime[unique_df_id]) #next function to add it to a file
else:
area_sum = df_['AREA_KM'].sum()
dispurse_number = area_sum / len(date_ranges)
#print('Date range length: ', len(date_ranges))
#print('DF length: ', len(series_area))
#series_dates = date_ranges.to_series() #in order to merge into a df
lst_date_rng = date_ranges.to_list() #change it to list instead so index does not come with it
df_Htime[unique_df_id] = pd.DataFrame({'date':lst_date_rng, 'areakm':0, 'state':state})
df_Htime[unique_df_id].fillna(0, inplace=True)
#df_Htime[unique_df_id]['areakm'] = df_Htime[unique_df_id]['areakm'] + disperser
#send it to the function call instead
disperse_areakms(df_Htime[unique_df_id], dispurse_number) #next function to disperse the left over numbers
create_big_csv(df_Htime[unique_df_id]) #next function to add it to a file
# + id="8S0S-wKmstTN"
def disperse_areakms(df_, disperser):
df_['areakm'] = df_['areakm'] + disperser
print(df_.head())
return(df_)
# + id="T25cE0YQstYa"
df_clean(df)
# + id="W1D1kkGbstc1"
#Load the csv into a dataframe
#Take headers out
#shift first column out of header column
#rename the headers
#set index
df2 = pd.read_csv('df_dattime_byStateYears1.csv', header=None)
df2.rename(columns={0:'date', 1:'areakm', 2:'state'}, inplace=True)
df2['date'] = pd.to_datetime(df2['date'], format='%Y-%m-%d', errors='coerce')
df2.set_index(df2['date'], inplace=True)
# + id="yU81eJYhsthN"
df2.isnull().values.sum()
df2.isna().values.sum()
df2.state.unique()
# + id="26X6jhn_stk-"
#all states are complete and in order of time.
df_pa = df2.loc[df2['state'] == 'PA']
df_am = df2.loc[df2['state'] == 'AM']
df_rr = df2.loc[df2['state'] == 'RR']
df_ro = df2.loc[df2['state'] == 'RO']
df_mt = df2.loc[df2['state'] == 'MT']
df_ac = df2.loc[df2['state'] == 'AC']
df_ma = df2.loc[df2['state'] == 'MA']
df_to = df2.loc[df2['state'] == 'TO']
df_ap = df2.loc[df2['state'] == 'AP']
# + [markdown] id="u_Z8nj0HtGyp"
# ## Visualization of total cumsum deforested per state
# + id="UZII48o3stoI"
df_pa['areasum'] = df_pa.areakm.cumsum()
df_am['areasum'] = df_am.areakm.cumsum()
df_rr['areasum'] = df_rr.areakm.cumsum()
df_ro['areasum'] = df_ro.areakm.cumsum()
df_mt['areasum'] = df_mt.areakm.cumsum()
df_ac['areasum'] = df_ac.areakm.cumsum()
df_ma['areasum'] = df_ma.areakm.cumsum()
df_to['areasum'] = df_to.areakm.cumsum()
df_ap['areasum'] = df_ap.areakm.cumsum()
# + id="-S-CJ1WutOY0"
#make a bif df with all states once again
frames = [df_pa, df_am, df_rr, df_ro, df_mt, df_ac, df_ma, df_to, df_ap]
df_cs = pd.concat(frames)
# + id="KMkt51hCtOeO"
areasum_states = df_cs.groupby(by='state')['areasum'].sum().sort_values(ascending=False).reset_index()
areasum_states = areasum_states.sort_values(by='areasum', ascending=True)
# + id="KHGWdslttOiI"
plt.figure(figsize = (16, 9))
# plot
ax = sns.barplot(x = areasum_states['state'], y = areasum_states['areasum'], palette = "Reds", alpha = 0.85)
plt.title("States", fontsize = 25)
plt.xlabel("State", fontsize = 20)
plt.ylabel("areasum", fontsize = 20)
plt.xticks(fontsize = 15)
plt.yticks(fontsize = 15)
plt.legend(fontsize = 15)
# + id="MN3MwCjktOoq"
sns.lineplot(data=df_cs, x='date', y='areasum', hue='state')
# + id="sruhZtnTtOrb"
fig, axs = plt.subplots(3, 3, sharex='col', sharey='row')
axs[0, 0].plot(df_pa['date'], df_pa['areasum'])
axs[0, 0].set_title('PA')
axs[0, 1].plot(df_am['date'], df_am['areasum'])
axs[0,1].set_title('AM')
axs[0, 2].plot(df_rr['date'], df_rr['areasum'])
axs[0,2].set_title('RR')
axs[1, 0].plot(df_ro['date'], df_ro['areasum'])
axs[1,0].set_title('RO')
axs[1, 1].plot(df_mt['date'], df_mt['areasum'])
axs[1, 1].set_title('MT')
axs[1, 2].plot(df_ac['date'], df_ac['areasum'])
axs[1, 2].set_title('AC')
axs[2, 0].plot(df_ma['date'], df_ma['areasum'])
axs[2,0].set_title('MA')
axs[2, 1].plot(df_to['date'], df_to['areasum'])
axs[2,1].set_title('TO')
axs[2, 2].plot(df_ap['date'], df_ap['areasum'])
axs[2,2].set_title('AP')
# + [markdown] id="R_CwpM0jtxx8"
# ## Failed Models
# + [markdown] id="up1e8R1Yt6dF"
# Testing models on the state of Amazonia over the years of 2008-2019. df_am
#
# + [markdown] id="O8yWgHTcuOWY"
# ### Resampling, monthly
# + id="GZ44h75Dt80Z"
df_am_month = df_am.resample(rule='MS').mean()
# + id="EZVXi0G9t84s"
train_monthly = df_am_month.iloc[:121]
test_monthly = df_am_month.iloc[120:]
# + id="b-9zjK_mt88j"
fitted_model = ExponentialSmoothing(train_monthly['areasum'], trend='add', seasonal_periods=12).fit()
test_predictions = fitted_model.forecast(24) #2 years into future
# + id="FZBHIWFpt9As"
#plot them all
train_monthly['areasum'].plot(figsize=(12,5), legend=True, label='Train')
test_monthly['areasum'].plot(legend=True, label='Test')
test_predictions.plot(legend=True, label='Prediction')
# + [markdown] id="MaHkWTk8uhcj"
# ### Yearly
# + id="a1QA9IoNt9E3"
df_am_year = df_am.resample(rule='A').mean()
df_am_year.plot()
#areakm is white noise
# + id="BxZRzrlyt9Iy"
train_yearly = df_am_year.iloc[:9]
test_yearly = df_am_year.iloc[8:]
# + id="q7LJj8S5urPr"
#fitted_model = ExponentialSmoothing(train_monthly['areasum'], trend='add', seasonal_periods=12).fit()
fitted_model = ExponentialSmoothing(train_yearly['areasum'], trend='add', seasonal_periods=1).fit()
test_predictions = fitted_model.forecast(10) #2 years into future
# + id="LJPghDlMurTq"
train_yearly['areasum'].plot(figsize=(12,5), legend=True, label='Train')
test_yearly['areasum'].plot(legend=True, label='Test')
test_predictions.plot(legend=True, label='Prediction')
# + [markdown] id="a7b1UCBAuws1"
# ### Sarimax with Monthly data
# + id="PIky5agWurXp"
#look at the seasonal decompose:
result_m = seasonal_decompose(df_am_month['areasum'], model='add')
result_m.plot();
# + id="VA7vN2gnu3Mu"
auto_arima(df_am_month['areasum'], seasonal=True, m=12).summary() #monthly data, 12 rows per year
# + id="f5w54pYju3Ts"
#forecast into future- length of dataset is 144 rows
#train_sm = train Sarimax Monthly
train_sm = df_am_month.iloc[:132] #minus the last year
test_sm = df_am_month.iloc[132:]
# + id="16DSzrTku3Yz"
model = SARIMAX(train_sm['areasum'], order=(0,2,0))
# + id="HCo4GnSau9yE"
result_sm = model.fit()
result_sm.summary()
# + id="K9cMNOeiu91W"
start_sm = len(train_sm)
end_sm = len(train_sm) + len(test_sm) - 1
# + id="P2RyySSuu934"
#now create the predictions
prediction_sm = result_sm.predict(start_sm, end_sm, type='levels').rename('Sarima Monthly predications')
# + id="T8F_5n_ru96a"
test_sm['areasum'].plot(legend=True, figsize=(12, 8))
prediction_sm.plot(legend=True)
# + id="bzWY5rklu98y"
fcast = result_sm.predict(len(df_am_month), len(df_am_month) + 11, type='levels').rename('Sarima Forecast')
test_sm['areasum'].plot(legend=True, figsize=(12, 8))
prediction_sm.plot(legend=True)
fcast.plot(legend=True)
# + [markdown] id="F4H7421IvM78"
# ### Sarimax with Yearly data
# + id="c8-dcWfjvRe7"
result_y = seasonal_decompose(df_am_year['areasum'], model='add')
result_y.plot();
# + id="X4NzeSXavRiP"
auto_arima(df_am_year['areasum'], m=1).summary()
#SARIMAX(2, 1, 0)
# + id="uhRW9-TYvRn0"
train_sy = df_am_year.iloc[:10] #minus the last year
test_sy = df_am_year.iloc[10:]
# + id="CYqgFfMCvRrc"
model_y = SARIMAX(train_sy['areasum'], order=(2,1,0))
resuly_y = model_y.fit()
start_sy = len(train_sy)
end_sy = len(train_sy) + len(test_sy) - 1
# + id="Yw2fsDULvR0W"
#now create the predictions
prediction_sy = resuly_y.predict(start_sy, end_sy, type='levels').rename('Sarima Yearly predications')
# + id="6kX3KSvSvaNC"
test_sy['areasum'].plot(legend=True, figsize=(12, 8))
prediction_sy.plot(legend=True)
# + id="ej3xK4dpvaQo"
#future
fcast_y = resuly_y.predict(len(df_am_year), len(df_am_year) + 3, type='levels').rename('Sarima Forecast')
# + id="8PCK6li8vaTb"
test_sy['areasum'].plot(legend=True, figsize=(12, 8))
prediction_sy.plot(legend=True)
fcast_y.plot(legend=True)
# + [markdown] id="SrtCIBxJvvy4"
# ### More failed models
# + [markdown] id="j6GmIJF4v3Oi"
# Notes:
# - choosing arma/arima orders. chooseing the best p,q,a
# - finding out the orders of the ar and ma components.
# - finding out if the i component is needed
# - if the aurocorrelation plot shows positive autocorrelation at first lag (lag-1), suggested to use AR terms in relation to lag
# - if the autocorrelation plot shows negative autocorrelation, suggest using MA terms
# - p: number of lag observations included in ar component of the model
# - d: number of times the raw observations are differenced
# - q: size of moving average window.
# + [markdown] id="Td1EcB0SzUlz"
# ##Arima
# non seasonal arima (p, d, q) </br>
# p = corresponds to the AR poriont of model</br>
# d = Indegrated componend. Differencing- diff of observations. In order to make the time series stationary (statsmodels as the diff function) </br>
# q = corresponds to MA component. Plotting out moving average, and using the residual error </br>
#
# + id="xlIJRGupwpbb"
def adf_test(series, title=''):
print(f'Augmented Dickey-Fuller Test: {title}')
result=adfuller(series.dropna(), autolag='AIC')
labels=['ADF test stats', 'p-value', 'number of lags used', 'number of observations']
out = pd.Series(result[0:4], index=labels)
for key, val in result[4].items():
out[f'critical value ({key})'] = val
print(out.to_string())
if result[1] <= 0.05:
print('Strong evidence against null hypthesis.', '\n', 'Reject null hypothesis.', '\n', 'Data has no unit root and is stationary')
else:
print('Weake evidence against null hypthesis.', '\n', 'Fail to reject null hypothesis.', '\n', 'Data has a unit root and is non-stationary')
# + [markdown] id="dRev4kgrzZVY"
# model = AR(train_data['areasum'])
# + id="W3hogTaHzn73"
model = AR(train_data['areasum'])
AR1fit.params
# + id="6TVUuJN5zoEv"
start = len(train_data)
end = len(train_data) + len(test_data) -1
# + id="C1rs1r90zoKO"
AR1fit.predict(start=start, end=end)
# + id="tZHHcZ6yzvf-"
#compare predicted values to real known test values
prediction1 = AR1fit.predict(start=start, end=end)
#name it to keep track
prediction1 = prediction1.rename('AR(1) Predictions')
# + id="RDUxDsbWzvsX"
test_data['areasum'].plot(figsize=(12,8), legend=True, label='Origional')
prediction1.plot(legend=True)
#under predicting
# + id="uU1_aV9hzv0P"
#try to improve this by expanding order
#name it to keep track
AR2fit = model.fit(maxlag=2)
AR2fit.params
# + id="sVRScRRzzv7o"
prediction2 = AR2fit.predict(start=start, end=end)
prediction2 = prediction2.rename('AR2 predictions')
# + id="WLMhPT1Dz28o"
#now plot the three
test_data['areasum'].plot(figsize=(12,8), legend=True, label='Origional')
prediction1.plot(legend=True)
prediction2.plot(legend=True)
# + id="tEOyFpK4z3CM"
#finidng the correct order value, let statsmodels decide, do not specify maxlag
#look into the ic parameter
ARfit = model.fit(ic='t-stat')
ARfit.params
# + id="baFq_GvDz3G8"
prediction28 = ARfit.predict(start=start, end=end)
prediction28 = prediction28.rename('AR28 Predictions')
# + id="bMS_b0xqz3L7"
#evaluate it
labels = ['AR1', 'AR2', 'AR28']
preds = [prediction1, prediction2, prediction28]
# + id="WpaqazKpz3Pw"
for i in range(3):
#np.sqrt() #use if you want
error = mean_squared_error(test_data['areasum'], preds[i])
print(f'{labels[i]} error: {error}')
# + id="BIe2aKQq0Fym"
#plot all 4
test_data['areasum'].plot(figsize=(12,8), legend=True, label='Origional')
prediction1.plot(legend=True)
prediction2.plot(legend=True)
prediction28.plot(legend=True)
# + [markdown] id="SC2u0zqRwd3b"
# #### Arma with df_am_year1
# + id="yeSNMpJWvaWE"
#we know seasonal is false
#trace will show you the first few arima models that it is trying to fix
stepwise_fit = auto_arima(df_am['areasum'], start_P=0, start_q=0, max_p=6, max_q=3, seasonal=False, trace=True)
# + id="5oOV0KYqwLdY"
stepwise_fit.summary()
#why does it suggest a SARIMA model?
# + id="JvWBmE1lwLgW"
#look at first year
df_am_year1 = df_am[:366]
# + id="9VbZEhfcwLlv"
df_am_year1.index
# + id="d8be1joIwLoM"
df_am_year1['areakm'].plot(figsize=(12,8))
# + id="B10mdzeIwhl7"
adf_test(df_am_year1['areakm'])
# + id="FRIv3_etwhoZ"
auto_arima(df_am_year1['areakm'], seasonal=False).summary()
# + id="0dm0GwZ1whq7"
auto_arima(df_am['areakm'], seasonal=False).summary()
# + [markdown] id="gTlX43tgw1J7"
# Failed, moving on to **ARIMA**
# + [markdown] id="IT6DUT0Qw8jY"
# #### ARIMA with df_am
# + [markdown] id="k2vJK7kcw3lc"
# using df_am
# + id="gnuvL9Jrwhue"
test_am = df_am[:3650]
train_am = df_am[3650:]
# + id="rUjsl6GVwhxS"
model = ARIMA(train_am['areakm'], order=(2,1,3))
results = model.fit()
results = model.fit()
# + id="n2vUYDygwhzi"
#set start and end location (dates)
start_am = len(train_am)
end_am = len(train_am) + len(test_am) -1
# + id="vQzXWyNNwh2d"
predictions = results.predict(start_am,end_am).rename('ARIMA(2,1,3)')
# + id="b4Dfx39FxZIp"
test_am['areakm']
# + id="4DfAk8dyxZLK"
predictions
# + id="W6x16_vDxZN2"
test_am['areakm'].plot(figsize=(12,8), legend=True)
predictions.plot(legend=True)
# + [markdown] id="ytt47OdwxhxO"
# ### Resample yearly, SARIMAX
# + id="rHAYTlUdxZQS"
df_am_yearly = df_am.resample(rule='A').sum()
# + id="WhQddKuexZTO"
df_am_yearly.plot()
# + id="AX93xFIRxZVg"
results3 = auto_arima(df_am_yearly, start_P=0, start_q=0, max_p=6, max_q=3, seasonal=False, trace=True)
# + id="BNsH7VczxomX"
results3.summary()
# + [markdown] id="patKWZdwxvef"
# - the time series is white noise?
# - mean = 0
# - std is contant with time
# - correlatoin lag between lags is 0
# - is it stationary?
# - mean is constant
# - std is constant
# - no seasonality
#
# + id="gx0GOIs3xouJ"
smodel = SARIMAX(train_am['areakm'], order=(0,1,3), enforce_invertibility=False)
# + id="5ot7ul-Axoyf"
results = smodel.fit()
results.summary()
# + id="clRSdOL3xo1x"
start = len(train_am)
end = len(train_am) +len(test_am) -1
# + id="8b1nTdasxo4c"
predictions = results.predict(start,end).rename('Sarima')
predictions = results.predict(start,end).rename('Sarima')
test_am['areakm'].plot(figsize=(15,8))
predictions.plot()
# + [markdown] id="-wbDks8fybNC"
# # Evaluation Methods
# + [markdown] id="Zc7U0VPlyhJa"
# ###ACF and PACF
# -autocorrelation and partial autocorrelation tests </br>
# -correlation: closer to 0, weak relationship. +1 / -1 negativly or positively correlated
#
# - autocorrelation, compares a series to itself, lagged
# + id="xZhNOmKgyebP"
#check if frequency is set
#if none, then set it
df_am.index.freq= 'D'
print(df_am.index)
# + id="v5mERPhBykek"
#auto corelation
acf(df_am['areasum'])
#positivly correlated
# + id="nOr-BnROykl1"
lag_plot(df_am['areasum'])
#is this correct? this is purely strong autocorrelation
# + id="R_wPqRdTykpW"
plot_acf(df_am['areasum'], lags=40);
#confidence interval - correlation values outside of it, highly likely to be a correlation.
# + id="s1LjQ41YyksL"
#look at the same plot on the areakm column
plot_acf(df_am['areakm'], lags=40);
# + id="Z8UGMcpwykxa"
#what is this?
df_am['areakm'].plot()
# + id="Ytsfozljyk0u"
#PACF
#stationary
#if a sharp cutoff, indicator to add pr terms
plot_pacf(df_am['areakm'], lags=40);
# + id="IBEJtxCdyk3f"
#non stationary
plot_pacf(df_am['areasum'], lags=40);
# + id="EuYDxnIdyk6G"
# + id="VgPNgPmFyk8d"
# + id="U0NIceWtylA1"
# + id="EOVj8jkfylD6"
| data/notebooks/BirdsAI_ProdesDataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import k3d
vertices = [
-10, 0, -1,
10, 0, -1,
10, 0, 1,
-10, 0, 1,
]
indices = [
0, 1, 3,
1, 2, 3
]
plot = k3d.plot()
plot += k3d.mesh(vertices, indices)
plot.display()
# +
from ipywidgets import widgets,interact
vertex_attribute = [0, 1, 1, 0]
plot = k3d.plot()
mesh = k3d.mesh(vertices, indices, attribute=vertex_attribute, color_map=k3d.basic_color_maps.CoolWarm, color_range=[0.0, 1.0])
plot += mesh
basic_color_maps = [attr for attr in dir(k3d.basic_color_maps) if not attr.startswith('__')]
paraview_color_maps = [attr for attr in dir(k3d.paraview_color_maps) if not attr.startswith('__')]
matplotlib_color_maps = [attr for attr in dir(k3d.matplotlib_color_maps) if not attr.startswith('__')]
@interact(x=widgets.Dropdown(options=basic_color_maps, value=basic_color_maps[0], description='Basic ColorMap:'))
def g(x):
mesh.color_map = getattr(k3d.basic_color_maps, x)
@interact(x=widgets.Dropdown(options=paraview_color_maps, value=paraview_color_maps[0], description='ParaView ColorMap:'))
def g(x):
mesh.color_map = getattr(k3d.paraview_color_maps, x)
@interact(x=widgets.Dropdown(options=matplotlib_color_maps, value=matplotlib_color_maps[0], description='MatplolLib ColorMap:'))
def g(x):
mesh.color_map = getattr(k3d.matplotlib_color_maps, x)
plot.display()
# -
# show vertex numbers:
import numpy as np
v = np.array(vertices).reshape((len(vertices)//3, 3))
for i, pos in enumerate(v):
plot += k3d.text(text=str(i), position=pos, color=0, reference_point='cc')
| examples/mesh.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import theano.tensor as T
x = T.dmatrix("x")
y = T.dmatrix("y")
z = x + y
type(z)
type(z.owner)
z.owner.op.name
w = x * 2
w.owner.op.name
print(w.owner)
print(len(w.owner.inputs))
print(w.owner.inputs) # second input is not 2
print(z.owner)
print(len(z.owner.inputs))
print(z.owner.inputs)
w.owner.inputs[1].owner.inputs
# graph visualization
import theano
import pydot
v = theano.tensor.vector()
from IPython.display import SVG
SVG(theano.printing.pydotprint(v*2, return_image=True,
format='svg'))
# optimization visualization
import theano
import pydot
a = theano.tensor.vector("a")
b = a + a ** 10
f = theano.function([a], b)
print(f([0,1,2]))
theano.printing.pydotprint(b, outfile = './grap.png', var_with_name_simple=True)
theano.printing.pydotprint(f, outfile="./graph_opt.png", var_with_name_simple=True)
| theano tutorial 3 __graph structure__.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
from graphframes.examples import Graphs
graph = Graphs(sqlContext).friends()
page_rank = graph.pageRank(resetProbability=0.15, tol=0.1)
display(page_rank.vertices)
| ch11 - Graph Analysis with GraphFrames/PageRank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
from tensorflow.keras.models import Sequential
from sklearn.preprocessing import MinMaxScaler
from tensorflow.keras.callbacks import EarlyStopping
from tensorflow.keras.layers import *
from tensorflow.keras.optimizers import Adam
# Define the target polynomial
def makeY(x):
return 5*x**3 + 2*x**2 + 6*x +8
# this is our input vector, 30k evenly spaced points from -5 to 5
x = np.linspace(-5,5,30000)
# now generate our output values
y=makeY(x)
# and here's what it looks like
plt.plot(x,y)
plt.grid()
# +
# Neural network models generally train more easily when they deal with values
# between -1 and 1
# Use the MMScaler to squish our data into that range
xscaler = MinMaxScaler(feature_range=(-1,1))
yscaler = MinMaxScaler(feature_range=(-1,1))
x= np.array(x) # x was a python list and is now a vector
# the scaler doesn't want a vector it wants a 2d array
# [] is different from [[]]
x= np.reshape(x, (x.shape[0],1))
# define the transform (with fit)
# use the transform (transform) to return the data
x = xscaler.fit_transform(x)
y = np.array(y)
y = np.reshape(y, (y.shape[0],1))
y = yscaler.fit_transform(y)
# +
# implement the required model with one input neuron
# one hidden layer with 3 nodes
# and one output neuron
# tanh is used because we mapped our data from -1 to 1
# if we were between 0 and 1, sigmoid would have probably worked better
# mse loss is used because this is a regression problem
model= Sequential()
model.add(Dense(3, input_shape=(1,), activation='tanh'))
model.add(Dense(1, activation='tanh'))
model.compile(loss='mse', optimizer=Adam(lr=0.0001), metrics=['mse'])
# -
# +
# set a large epoch count and use EarlyStopping to bail out when
# we start to overfit
model.fit(x,y, epochs=100, validation_split=.1, callbacks=[EarlyStopping(patience=5)])
# +
# pick a new value of x within the training range
# pass the value through our polynomial
unscaled_x = np.pi
unscaled_y = makeY(unscaled_x)
# the model works with scaled inputs and produces scaled outputs
scaled_x = xscaler.transform([[unscaled_x]])
scaled_y = yscaler.transform([[unscaled_y]])
scaled_y_hat = model.predict(scaled_x)
print("actual scaled: {} scaled predicted: {}".format(scaled_y, scaled_y_hat))
print("actual unscaled: {} unscaled predicted: {}".format(unscaled_y, yscaler.inverse_transform(scaled_y_hat)))
# -
| polynomial regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# import numpy package for arrays and stuff
import numpy as np
# import matplotlib.pyplot for plotting our result
import matplotlib.pyplot as plt
# import pandas for importing csv files
import pandas as pd
# -
# Create artificial data set
dataset = np.array(
[[0, 'Model', 'Condition', 'Leslie', 'Price'],
[1, 'B3', 'excellent', 'yes', 4513],
[2, 'T202', 'fair', 'yes', 625],
[3, 'A100', 'good', 'yes', 1051],
[4, 'T202', 'good', 'no', 270],
[5, 'M102', 'good', 'yes', 870],
[6, 'A100', 'excellent', 'no', 1770],
[7, 'T202', 'fair', 'no', 99],
[8, 'A100', 'good', 'yes', 1900],
[9, 'E112', 'fair', 'no', 77]
])
df = pd.DataFrame(data=dataset[1:,1:], # values
index=dataset[1:,0], # 1st column as index
columns=dataset[0,1:]) # 1st row as the column names
df = df.astype(dtype={"Model":"category", "Condition":"category","Leslie":"category", "Price":np.int})
df
# +
# One hot encoding
X = pd.get_dummies(df[["Model","Condition","Leslie"]],drop_first=True)
y = df['Price']
#print(X)
# +
# import the regressor
from sklearn.tree import DecisionTreeRegressor
# create a regressor object
regressor = DecisionTreeRegressor(random_state = 0, max_depth = 2)
# fit the regressor with X and Y data
regressor.fit(X, y)
# -
from auxiliaryFunctions import rules
#generate JSON file
rules(regressor, X.columns.values, [y.name])
# +
import pydotplus
from sklearn.tree import export_graphviz
from IPython.display import Image
from IPython.display import display
dot_data = export_graphviz(
regressor,
out_file=None,
feature_names=X.columns.values,
filled=True,
rounded=True)
graph = pydotplus.graph_from_dot_data(dot_data)
display(Image(graph.create_png()))
| RegressionTree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
## Import libs
import findspark
findspark.init("/home/antonis/spark-2.3.0-bin-hadoop2.7")
import os.path
import pandas
import math
import time
from metrics.Correlation import Correlation
from IO.Output import Output
from IO.Input import Input
from metrics.RSquare import RSquare
from DataSet.Dataset import DataSet
import pyspark
from pyspark.sql import SparkSession
import sys
from pyspark import SparkContext
from pyspark.sql import SQLContext
from pyspark.sql.types import *
from pyspark.sql import Row
from pyspark.sql.functions import countDistinct,avg,stddev
from pyspark.ml.feature import (VectorAssembler,VectorIndexer,
OneHotEncoder , StringIndexer)
import DataSet.SnpsSelection as s
from pyspark.ml.tuning import CrossValidator, ParamGridBuilder
from pyspark.ml.evaluation import BinaryClassificationEvaluator
from pyspark.ml import Pipeline
from pyspark.ml.classification import LogisticRegression
from pyspark.ml.linalg import DenseVector
from pyspark.mllib.evaluation import MulticlassMetrics
from pyspark.ml.evaluation import RegressionEvaluator
from pyspark.ml.tuning import CrossValidator, ParamGridBuilder
from pyspark.ml.classification import RandomForestClassifier
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
from pyspark.ml.classification import LinearSVC
from pyspark.ml.classification import NaiveBayes
from pyspark.mllib.tree import DecisionTree, DecisionTreeModel
from pyspark.mllib.util import MLUtils
from pyspark.mllib.tree import GradientBoostedTrees, GradientBoostedTreesModel
from pyspark.ml.stat import Correlation
from pyspark.ml.evaluation import BinaryClassificationEvaluator
import math
import time
def writeCoef(path,snpsIds,sc,idToName,corr, name = None):
if not name:
print("give a name to file")
return
p = path + name + " ( " + time.strftime("%d-%m-%Y") + " ).txt "
i=1
while os.path.exists(p):
p = path + name + " ( " + time.strftime("%d-%m-%Y") + " ) " + '_' + str(i)+".txt"
i += 1
snps = []
for i in range(len(snpsIds)):
s = snpsIds[i]
#snps.append(idToName[s])
snps.append(s)
print("snpsIds = ",len(snpsIds))
print("idToName = ",len(idToName))
write = open(p,'w')
write.write("len = "+str(len(sc))+'\n')
write.write('corr = '+str(corr)+'\n')
for i in range(len(snps)):
write.write(str(snps[i])+'\t'+str(sc[i])+'\n')
write.close()
def calcCoeff(path,coefs, columns,corr):
ids = {}
ids['coef']={}
print("before = ",len(set(coefs)))
#for i in range(len(coefs)):
# coefs[i] = abs(coefs[i])
print("after = ",len(set(coefs)))
idToName = {}
nameToId = {}
for i in range(len(coefs)):
nameToId[coefs[i]] = []
for i in range(len(coefs)):
nameToId[coefs[i]].append(i)
idToName[i] = coefs[i]
ids['coef']['nameToId'] = nameToId
ids['coef']['idToName'] = idToName
sc = sorted(coefs,reverse=True)
top_30 = []
for i in range(30):
snp = ids['coef']['nameToId'][sc[i]][0]
ids['coef']['nameToId'][sc[i]].remove(snp)
top_30.append(columns[snp])
# top_30.append(snp)
#snpReduc['low'] = top_30
writeCoef(path,top_30,sc,columns,corr, name = 'mycutoffabs')
def showMetrics(c,value=1):
predictionAndLabels = c.map(lambda lp: (lp.prediction, float(lp.label)))
metrics = MulticlassMetrics(predictionAndLabels)
print("confusion matrix = ", metrics.confusionMatrix().toArray())
print("accuracy = ", metrics.accuracy)
print("recal = ", metrics.recall(value))
print("precision = ", metrics.precision(value))
print("f1 = ", metrics.fMeasure(1.0))
def showMyMetric(results,v1 = 1,v2 = 0):
if v1 == v2:
print("wrong values!!!v1 is the same with v2!!!!")
r = results.rdd
r3 = r.collect()
same0 = 0
same1 = 0
sum0 = 0
sum1 = 0
sumall = 0
for i in r3:
if i[v1] == 0:
sum0 += 1
if i[v2] == 0:
same0 += 1
elif i[v1] == 1:
sum1 += 1
if i[v2] == 1:
same1 += 1
sumall += 1
print('sum0 = ', sum0)
print('sum1 = ', sum1)
print('same0 = ', same0)
print('same1 = ', same1)
print('all = ', sumall)
print('all2 = ', sum0+sum1)
def calculateAvgMetrics(results,classLabel=1):
metircs = {}
predictionAndLabels = results.map(lambda lp: (lp.prediction, float(lp.label)))
metrics = MulticlassMetrics(predictionAndLabels)
#metrics["confusion_matrix"] = metrics.confusionMatrix().toArray()
metrics["accuracy"] = metrics.accuracy
metrics["recal"]= metrics.recall(classLabel)
metrics["precision"] = metrics.precision(classLabel)
metrics["f1"] = metrics.fMeasure(float(classLabel))
return metrics
def split(numFold = 0, df = None):
samples = {}
if numFold == 0:
print("wrong!!!!! num fold is zero (0)")
return
dfCount = df.count()
k = int (dfCount / numFold)
dfsplit = df
# print("k = ", k)
for i in range(1, numFold):
# print("count = ",dfCount)
x = (k*100) / dfCount
x = x / 100
split1, split2 = dfsplit.randomSplit([x,1-x],seed=2018)
dfsplit= split2
dfCount = dfCount - k
samples[i] = split1
samples[numFold] = split2
return samples
def balanedData(df):
label1 = df.filter("label == 1").count()
label0 = df.filter("label == 0").count()
print("label0 = ", label0)
print("label1 = ", label1)
l0 = label1 * 2
x = (l0 * 100) / label0
x = x / 100
print("label1 = ", label1)
print("l2 = ",l0)
print("label0 = ", label0)
print("x = ", x)
print()
t0 = df.filter('label == 0')
t1 = df.filter('label == 1')
train0,t2 = t0.randomSplit([x,1-x],seed = 11)
train_data = train0.union(t1)
return train_data
def reduceDismension(train_data, test_data):
li = len(df.columns)
input_data = train_data.rdd.map(lambda x: (x[li-1], DenseVector(x[:li-1])))
train0 = spark.createDataFrame(input_data, ["label", "features"])
final_data1 = train0.select('features')
corr = Correlation.corr(final_data1, "features")
corr = corr.head()[0].toArray()
snpsRed = []
snpsRed = s.lowCorrelation(corr, threshold=0.7, up=100, down=99)#oso megalytero threshold toso perissotero omoia einai
###############################################################################################################################
features = []
for i in range(0,len(train_data.columns)):
if 'rs' in train_data.columns[i] and train_data.columns[i] !='label':
features.append(train_data.columns[i])
snpsRed1 = []
for i in snpsRed:
snpsRed1.append(features[i])
snpsRed1.append('label')
features = []
for i in range(0,len(train_data.columns)):
if train_data.columns[i] not in snpsRed1:
features.append(train_data.columns[i])
dok_train = train_data.drop(*features)
dok_test = test_data.drop(*features)
li = len(dok_train.columns)
############################################################################################################################
input_data = dok_test.rdd.map(lambda x: (x[li-1], DenseVector(x[:li-1])))
test = spark.createDataFrame(input_data, ["label", "features"])
input_data = dok_train.rdd.map(lambda x: (x[li-1], DenseVector(x[:li-1])))
train = spark.createDataFrame(input_data, ["label", "features"])
print("train = ",train.head()[1].toArray().shape)
print("test = ",test.head()[1].toArray().shape)
return train, test
def crossVal(numFold = 0,data = None,classifier = None):
avgMetrics = {}
if data = None:
print("data not given")
return
if classifier = None:
print("classifier not given")
return
samples = split(numFold = numFold, df = data)
if samples = None:
return
for i in range(1,numFold + 1):
test_data = samples[i]
if i <= 2:
train_data = samples[3].union(samples[4])
s1 = 3
s2 = 4
else:
train_data = samples[1].union(samples[2])
s1 = 1
s2 = 2
for j in range(1,numFold + 1):
if j != i and j!= s2 and j != s1:
train_data = train_data.join(samples[j])
train,test = reduceDismension(train_data,test_data)
train = balanedData(train)
model = classifier.fit(train)
results = lr_model.transform(test)
evaluate = BinaryClassificationEvaluator(rawPredictionCol='prediction',labelCol='label')
AUC = evaluate.evaluate(results)
results = cross_results.select('prediction','label')
showMetrics(cross_results.rdd,value=1)
showMyMetric(results,v1 = 1,v2 = 0)
# -
# +
from pyspark import SparkConf, SparkContext
conf = (SparkConf()
.set("spark.driver.maxResultSize", "20g")
.set('spark_executor_cores',"3")
.set('spark.graphx.pregel.checkpointInterval','-1')
.set('spark.network.timeout','100000000')
.set('spark.executor.heartbeatInterval','10000000'))
#spark = SparkSession.builder.appName('melanoma').getOrCreate()
sc = SparkContext(conf=conf)
spark = SQLContext(sc)
# +
#spark.stop()
# -
# # Tests
# +
path = '/media/antonis/red/newdata/maf = 0.05/pvalue = 0.001/'
#pathSnp = '/media/antonis/red/newSet/maf/maf = 0.05/assoc/pvalue = 0.001/snpCodeTest.csv'
#pathSnp = '/media/antonis/red/newSet/maf/maf = 0.05/assoc/pvalue = 0.01/snp2.txt'
#pathSnp = '/media/antonis/red/newdata/maf = 0.05/pvalue = 0.001/snp2.txt'
pathSnp = '/media/antonis/red/newdata/maf = 0.05/pvalue = 0.001/snpCodeTest1.csv'
#pathSnp = '/media/antonis/Antonis_Moulopoulos/newSet/pvalue = 0.001/snp1.txt'
#pathSnp = '/media/antonis/Antonis_Moulopoulos/newdata/maf = 0.05/pvalue = 0.001/snp2.txt'
data = spark.read.option("maxColumns", 80000).csv(pathSnp,inferSchema=True,header=True)
data=data.withColumnRenamed('TARGET','label')
# +
print("data columns = ",len(data.columns))
features = []
for i in data.columns:
if 'rs' not in i and i !='label':
features.append(i)
print(len(features))
#print((features))
# +
#data.select('patients','label').show(10)
# +
from pyspark.ml.linalg import DenseVector
d = data.drop('patients')
for i in features:
d = d.drop(i)
print("columns = ",len(d.columns))
train_data,test_data = d.randomSplit([0.8,0.2],seed=18)
# +
samples = split(numFold = 10, df = d)
s = 0
for i in samples.keys():
s = s + samples[i].count()
print("sum = ",s)
# -
len(samples.keys())
'''t0 = train_data.count()
t1 = test_data.count()
print("test = ",t1)
print("train = ",t0)'''
# # Balanced Label1 to train
# +
label1 = train_data.filter("label == 1").count()
label0 = train_data.filter("label == 0").count()
print("label0 = ", label0)
print("label1 = ", label1)
l0 = label1 * 2
x = (l0 * 100) / label0
x = x / 100
print("label1 = ", label1)
print("l2 = ",l0)
print("label0 = ", label0)
print("x = ", x)
print()
t0 = train_data.filter('label == 0')
t1 = train_data.filter('label == 1')
train0,t2 = t0.randomSplit([x,1-x],seed = 11)
train_data = train0.union(t1)
#train1,t3 = t1.randomSplit([0.5,0.5],seed = 11)
#train_data = train0.union(train1)
#t2 = t2.union(t3)
#test_data = test_data.union(t2)
t0 = train_data.count()
t1 = test_data.count()
print("label0 = ",label0)
print('label1 = ', label1)
print("l1 = ",l0)
print('len test label1 = ',train_data.filter('label==1').count())
print('len test label0 = ',train_data.filter('label==0').count())
print('train = ', t0)
print('test = ', t1)
print('all = ', t0 + t1)
print('d = ',d.count())
# -
# # UnBalanced Label1 to train
# +
label1 = train_data.filter("label == 1").count()
label0 = train_data.filter("label == 0").count()
l1 = (label1/2) + ((label1/2) * 0.5)
x = (l1 * 100) / label0
x = x / 100
print("label1 = ", label1)
print("l1 = ",l1)
print("label0 = ", label0)
print("x = ", x)
t0 = train_data.filter('label == 0')
train0,t2 = t0.randomSplit([x,1-x],seed = 11)
train1 = train_data.filter("label == 1")
train_data = train0.union(train1)
test_data = test_data.union(t2)
t0 = train_data.count()
t1 = test_data.count()
print("label0 = ",label0)
print('label1 = ', label1)
print("l1 = ",l1)
print('train = ', t0)
print('test = ', t1)
print('all = ', t0 + t1)
print('d = ',d.count())
# -
# # dokimh
# +
li = len(d.columns)
input_data = train_data.rdd.map(lambda x: (x[li-1], DenseVector(x[:li-1])))
train0 = spark.createDataFrame(input_data, ["label", "features"])
final_data1 = train0.select('features')
corr = Correlation.corr(final_data1, "features")
corr = corr.head()[0].toArray()
# -
snpsRed = []
snpsRed = s.lowCorrelation(corr, threshold=0.7, up=100, down=99)#oso megalytero threshold toso perissotero omoia einai
# +
features = []
for i in range(0,len(d.columns)):
if 'rs' in d.columns[i] and d.columns[i] !='label':
features.append(d.columns[i])
print("len features1 = ",len(features))
snpsRed1 = []
for i in snpsRed:
snpsRed1.append(features[i])
snpsRed1.append('label')
print('len snpsRed1 = ', len(snpsRed1))
features = []
for i in range(0,len(d.columns)):
if d.columns[i] not in snpsRed1:
features.append(d.columns[i])
print("len features2 = ",len(features))
dok_train = train_data.drop(*features)
dok_test = test_data.drop(*features)
print("columns dok train = ",len(dok_train.columns))
print("columns dok test = ",len(dok_test.columns))
li = len(dok_train.columns)
print('first column = ', dok_train.columns[0])
print('last column = ', dok_train.columns[li - 1])
print('test first column = ', dok_test.columns[0])
print('test last column = ', dok_test.columns[li - 1])
# +
li = len(dok_train.columns)
#li = len(test_data.columns)
input_data = dok_test.rdd.map(lambda x: (x[li-1], DenseVector(x[:li-1])))
test = spark.createDataFrame(input_data, ["label", "features"])
input_data = dok_train.rdd.map(lambda x: (x[li-1], DenseVector(x[:li-1])))
train = spark.createDataFrame(input_data, ["label", "features"])
'''input_data = test_data.rdd.map(lambda x: (x[li-1], DenseVector(x[:li-1])))
test = spark.createDataFrame(input_data, ["label", "features"])
input_data = train_data.rdd.map(lambda x: (x[li-1], DenseVector(x[:li-1])))
train = spark.createDataFrame(input_data, ["label", "features"])'''
# -
# # telos dokimhs
print("train = ",train.head()[1].toArray().shape)
print("test = ",test.head()[1].toArray().shape)
# # Linear Regression
log_reg = LogisticRegression(featuresCol='features',labelCol='label',maxIter=10)
lr_model = log_reg.fit(train)
# +
results = lr_model.transform(test)
from pyspark.ml.evaluation import BinaryClassificationEvaluator
evaluate = BinaryClassificationEvaluator(rawPredictionCol='prediction',labelCol='label')
AUC = evaluate.evaluate(results)
print("auc = ",AUC)
'''0.7909133853963526'''
# +
'''Param for metric name in evaluation. Supports: - "rmse" (default): root mean squared error -
"mse": mean squared error - "r2": R^2^ metric - "mae": mean absolute error metricName='mae' '''
'''pipeline = Pipeline(stages=[log_reg])
modelEvaluator=RegressionEvaluator(predictionCol='prediction', labelCol='label',metricName='mse')
paramGrid = ParamGridBuilder().build()
crossval = CrossValidator(estimator=log_reg,
estimatorParamMaps=paramGrid,
evaluator=modelEvaluator,
numFolds=4)
cvModel = crossval.fit(train)
cross_results = cvModel.transform(test)
results = cross_results.select('prediction','label')
#results.show(10)'''
# +
'''Param for metric name in evaluation. Supports: - "rmse" (default): root mean squared error -
"mse": mean squared error - "r2": R^2^ metric - "mae": mean absolute error metricName='mae' '''
pipeline = Pipeline(stages=[log_reg])
modelEvaluator=MulticlassClassificationEvaluator(predictionCol='prediction', labelCol='label',metricName='accuracy')
paramGrid = ParamGridBuilder().build()
crossval = CrossValidator(estimator=log_reg,
estimatorParamMaps=paramGrid,
evaluator=modelEvaluator,
numFolds=4)
cvModel = crossval.fit(train)
cross_results = cvModel.transform(test)
results = cross_results.select('prediction','label')
#results.show(10)
# -
avgResults = cvModel.avgMetrics
#print(len(d.columns))
#print(d.columns[len(d.columns)-1])
calcCoeff(path,lr_model.coefficientMatrix.toArray()[0],d.columns,0.2)
'''trainingSummary = cvModel.bestModel.stages[-1].summary
trainingSummary.roc.show()
print("areaUnderROC: " + str(trainingSummary.areaUnderROC))'''
showMetrics(cross_results.rdd,value=1)
showMyMetric(results,v1 = 1,v2 = 0)
# # RFR
rf = RandomForestClassifier(featuresCol='features',labelCol='label', numClasses=2, categoricalFeaturesInfo={},
numTrees=50, featureSubsetStrategy="auto",
impurity='gini', maxDepth=4, maxBins=32)
rf_model = rf.fit(train)
results = rf_model.transform(test)
from pyspark.ml.evaluation import BinaryClassificationEvaluator
evaluate = BinaryClassificationEvaluator(rawPredictionCol='prediction',labelCol='label')
AUC = evaluate.evaluate(results)
AUC
# +
modelEvaluator=RegressionEvaluator(predictionCol='prediction', labelCol='label')
paramGrid = ParamGridBuilder().build()
crossval = CrossValidator(estimator=rf,
estimatorParamMaps=paramGrid,
evaluator=modelEvaluator,
numFolds=4)
cvModel = crossval.fit(train)
cross_results = cvModel.transform(test)
results = cross_results.select('prediction','label')
#results.show(10)
# -
showMetrics(cross_results.rdd,value=1)
showMyMetric(results,v1 = 1,v2 = 0)
# # SVM
#lsvc = LinearSVC(maxIter=10, regParam=0.1)
lsvc = LinearSVC(featuresCol='features',labelCol='label',maxIter=10)
lsvcModel = lsvc.fit(train)
lsvc.setThreshold(0.5)
results = lsvcModel.transform(test)
from pyspark.ml.evaluation import BinaryClassificationEvaluator
evaluate = BinaryClassificationEvaluator(rawPredictionCol='prediction',labelCol='label')
AUC = evaluate.evaluate(results)
AUC
# +
modelEvaluator=MulticlassClassificationEvaluator(predictionCol='prediction', labelCol='label',metricName='accuracy')
paramGrid = ParamGridBuilder().build()
crossval = CrossValidator(estimator=lsvc,
estimatorParamMaps=paramGrid,
evaluator=modelEvaluator,
numFolds=4)
cvModel = crossval.fit(train)
cross_results = cvModel.transform(test)
results = cross_results.select('prediction','label')
#results.show(10)
# -
showMetrics(cross_results.rdd,value=1)
showMyMetric(results,v1 = 1,v2 = 0)
lsvc.clearThreshold()
# +
#calcCoeff(path,lr_model.coefficientMatrix.toArray()[0],d.columns)
# -
'''print('len test label1 = ',train_data.filter('label==1').count())
print('len test label0 = ',train_data.filter('label==0').count())'''
# # Bernoulli
nb = NaiveBayes(predictionCol='prediction', labelCol='label',smoothing=0.2)
nbg = nb.fit(train)
results = nbg.transform(test)
from pyspark.ml.evaluation import BinaryClassificationEvaluator
evaluate = BinaryClassificationEvaluator(rawPredictionCol='prediction',labelCol='label')
AUC = evaluate.evaluate(results)
AUC
# +
modelEvaluator=MulticlassClassificationEvaluator(predictionCol='prediction', labelCol='label',metricName='accuracy')
paramGrid = ParamGridBuilder().build()
crossval = CrossValidator(estimator=nb,
estimatorParamMaps=paramGrid,
evaluator=modelEvaluator,
numFolds=4)
cvModel = crossval.fit(train)
cross_results = cvModel.transform(test)
results = cross_results.select('prediction','label')
#results.show(10)
# -
showMetrics(cross_results.rdd,value=1)
showMyMetric(results,v1 = 1,v2 = 0)
# # Tree
# +
from pyspark.mllib.regression import LabeledPoint
from pyspark.sql.functions import col
from pyspark.ml.regression import DecisionTreeRegressor
from pyspark.ml.classification import DecisionTreeClassifier
'''features = []
for i in dok_train.columns:
features.append(i)
assempler = VectorAssembler(inputCols=features,outputCol='features')
output = assempler.transform(data)
tr = output.select('label','features').rdd.map(lambda row: LabeledPoint(row.label, row.features))
treeD = DecisionTree.trainClassifier(tr, numClasses=2, categoricalFeaturesInfo={},
impurity='gini', maxDepth=4, maxBins=32)'''
treeD = DecisionTreeClassifier(predictionCol='prediction', labelCol='label')
tree = treeD.fit(train)
#tree = treeD.fit(train0)
print("ok")
# -
results = tree.transform(test)
from pyspark.ml.evaluation import BinaryClassificationEvaluator
evaluate = BinaryClassificationEvaluator(rawPredictionCol='prediction',labelCol='label')
AUC = evaluate.evaluate(results)
AUC
# +
modelEvaluator=MulticlassClassificationEvaluator(predictionCol='prediction', labelCol='label',metricName='accuracy')
paramGrid = ParamGridBuilder().build()
crossval = CrossValidator(estimator=treeD,
estimatorParamMaps=paramGrid,
evaluator=modelEvaluator,
numFolds=4)
cvModel = crossval.fit(train)
cross_results = cvModel.transform(test)
results = cross_results.select('prediction','label')
#results.show(10)
# -
showMetrics(cross_results.rdd,value=1)
showMyMetric(results,v1 = 1,v2 = 0)
modelFile = path+'treeRules.txt'
f = open(modelFile,"w")
f.write(tree.toDebugString)
f.close()
# # Gradient-Boosted Trees (GBTs)
# +
modelGBT = GradientBoostedTrees.trainClassifier(train,categoricalFeaturesInfo={}, numIterations=10)
#modelGBT = GradientBoostedTrees(predictionCol='prediction', labelCol='label')
gbt = treeD.fit(train)
# -
results = gbt.transform(test)
from pyspark.ml.evaluation import BinaryClassificationEvaluator
evaluate = BinaryClassificationEvaluator(rawPredictionCol='prediction',labelCol='label')
AUC = evaluate.evaluate(results)
AUC
# +
modelEvaluator=RegressionEvaluator(predictionCol='prediction', labelCol='label')
paramGrid = ParamGridBuilder().build()
crossval = CrossValidator(estimator=gbt,
estimatorParamMaps=paramGrid,
evaluator=modelEvaluator,
numFolds=4)
cvModel = crossval.fit(train)
cross_results = cvModel.transform(test)
results = cross_results.select('prediction','label')
#results.show(10)
# -
showMetrics(cross_results.rdd,value=1)
showMyMetric(results,v1 = 1,v2 = 0)
| code/spark_final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import math
""" Confidence Interval of 95% Function
mean = the population mean
std = standard deviation of the population
sample_size = size of the sample
"""
#Standard error method that comes into computation
def standard_error(std, sample_size):
return (std/(math.sqrt(sample_size)))
def ci_95(mean, se):
z = 1.96
ci_minus = mean - (z * se)
ci_plus = mean + (z * se)
ci = (ci_minus,ci_plus)
#returns a tuple
return ci
# -
# # 4.7 Chronic illness
# **Given Data :**
# <br/>
# point estimate = 45%<br/>
# Standard Error = 1.2%
# Creating a 95 % confidence interval for the proportion of US adults with one pr more chronic conditions.
ci_95(45,1.2)
# **<u>Inference : </u> : ** We are 95% confident that the proporion of U.S. citizen living with one or more chronic conditions is between 42.648 and 47.352
| Chapter 4/Exercises/4.7_Chronic_Illness_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow import keras
import seaborn as sns
from os.path import join
plt.style.use(["seaborn", "thesis"])
# -
plt.rc("figure", figsize= (8, 4))
# # Fetch Dataset
# +
from SCFInitialGuess.utilities.dataset import AbstractDataset
data_path = "../../dataset/EthenT/"
postfix = "EthenT"
dim = 72
basis = "6-311++g**"
n_electrons = 16
#data_path = "../butadien/data/"
#postfix = ""
#dim = 26
S = np.load(join(data_path, "S" + postfix + ".npy"))
P = np.load(join(data_path, "P" + postfix + ".npy"))
index = np.load(join(data_path, "index" + postfix + ".npy"))
molecules = np.load(join(data_path, "molecules" + postfix + ".npy"))
ind = int(0.8 * len(index))
ind_val = int(0.8 * ind)
molecules = (
molecules[:ind_val],
molecules[ind_val:ind],
molecules[ind:]
)
s_triu_norm, mu, std = AbstractDataset.normalize(S)
def split(x, y, ind):
return x[:ind], y[:ind], x[ind:], y[ind:]
s_train, p_train, s_test, p_test = split(S, P, ind)
s_train, p_train, s_val, p_val = split(s_train, p_train, ind_val)
# -
# # Define Candidates
# +
from SCFInitialGuess.descriptors.cutoffs import behler_cutoff_1
def plot_cutoff_model(model, R_c, t=None):
mus, sigmas = model
if t is None:
t = np.linspace(0, 1.2 * R_c, 200)
for r_s, eta in zip(mus, sigmas):
plt.plot(
t,
np.exp(-1 * eta*(t - r_s)**2) * \
behler_cutoff_1(t, R_c)
)
def plot_damped_model(model, tau, t=None):
mus, sigmas = model
if t is None:
t = np.linspace(0, 1.2 * tau, 200)
for r_s, eta in zip(mus, sigmas):
plt.plot(
t,
np.exp(-1 * eta*(t - r_s)**2) * \
np.exp(- t / tau)
)
def plot_normal_model(model, t):
for r_s, eta in zip(model[0], model[1]):
plt.plot(t, np.exp(-1 * eta*(t - r_s)**2))
def plot_periodic_model(model, t):
period = model[2]
for r_s, eta in zip(model[0], model[1]):
plt.plot(t,
np.exp(-1 * eta * ((t % period) - r_s)**2) + \
np.exp(-1 * eta * ((t % period) - period - r_s)**2)
)
# -
# ## Radial
#
x = np.linspace(0, 6, 200)
plt.plot(x, np.exp(-x / 5))
plt.plot(x, behler_cutoff_1(x, 5))
plt.show()
# ### Origin Centered
r_model_origin = (
[0.0] * 50,
np.linspace(10, 0.1, 50)
)
plot_cutoff_model(r_model_origin, 5)
plot_damped_model(r_model_origin, 2, np.linspace(0, 5*1.2, 200))
# ### Evenly distributed
r_model_evenly = (
np.linspace(0.1,4, 50),
np.linspace(30, 10, 50),
)
plot_cutoff_model(r_model_evenly, 5)
plot_damped_model(r_model_evenly, 2, np.linspace(0, 5*1.2, 200))
# ### Highest density around average bonding length
r_model_concentrated = (
list(np.linspace(0.2, 0.7, 11))[:-1] + \
list(np.linspace(0.7, 2.5, 30)) + \
list(np.linspace(2.5, 4, 11))[1:],
[500]*10 + \
[1000]*30 + \
[200]*10,
)
plot_cutoff_model(r_model_concentrated, 5)
plot_damped_model(r_model_concentrated, 2, np.linspace(0, 5*1.2, 200))
# ## Aizimutahl
phi_model_evenly = (
np.linspace(0, 2*np.pi, 50),
[300] * 50,
2 * np.pi
)
plot_periodic_model(phi_model_evenly, np.linspace(-0.5, 2.5*np.pi, 200))
plot_normal_model(phi_model_evenly, np.linspace(-0.5*np.pi, 2.5*np.pi, 200))
# ### Polar, das gleiche wie bei Azimuthal
theta_model_evenly = (
np.linspace(0, np.pi, 50),
[700] * 50,
np.pi
)
plot_periodic_model(theta_model_evenly, np.linspace(-0.5, 2.5*np.pi, 200))
plot_normal_model(theta_model_evenly, np.linspace(-0.5*np.pi, 2.5*np.pi, 200))
#
# # Train Benchmarksystem
# ## Utilities
# +
from SCFInitialGuess.utilities.constants import number_of_basis_functions as N_BASIS
from SCFInitialGuess.utilities.dataset import extract_triu_batch, extract_triu
from SCFInitialGuess.utilities.dataset import StaticDataset
def extract_dataset(molecules, p_batch, descriptor):
from SCFInitialGuess.utilities.dataset import AbstractDataset
G = []
for mol in molecules:
G.append(
descriptor.calculate_all_descriptors(mol).flatten()
)
#G = np.asarray(G)
p_triu = extract_triu_batch(p_batch, dim)
return G, p_triu
def make_dataset(descriptor, dim):
inputs_test, outputs_test = extract_dataset(
molecules[2],
p_test.reshape(-1, dim, dim),
descriptor
)
inputs_validation, outputs_validation = extract_dataset(
molecules[1],
p_val.reshape(-1, dim, dim),
descriptor
)
inputs_train, outputs_train = extract_dataset(
molecules[0],
p_train.reshape(-1, dim, dim),
descriptor
)
_, mu, std = StaticDataset.normalize(inputs_train + inputs_validation + inputs_test)
dataset = StaticDataset(
train=(
StaticDataset.normalize(inputs_train, mean=mu, std=std)[0],
np.asarray(outputs_train)
),
validation=(
StaticDataset.normalize(inputs_validation, mean=mu, std=std)[0],
np.asarray(outputs_validation)
),
test=(
StaticDataset.normalize(inputs_test, mean=mu, std=std)[0],
np.asarray(outputs_test)
),
mu=mu,
std=std
)
return dataset
# -
# ### NN
# +
#intializer = keras.initializers.TruncatedNormal(mean=0.0, stddev=0.01)
def make_model(
input_dim,
output_dim,
activation="elu",
learning_rate=1e-3
):
model = keras.Sequential()
# linear model => 1 layer
model.add(keras.layers.Dense(
output_dim,
#activation=activation,
input_dim=input_dim,
#kernel_initializer=intializer,
#bias_initializer='zeros',
kernel_regularizer=keras.regularizers.l2(1e-7)
))
model.compile(
optimizer=keras.optimizers.Adam(learning_rate),
loss='MSE',
metrics=['mse']
)
return model
# +
early_stopping = keras.callbacks.EarlyStopping(
monitor="val_mean_squared_error",
min_delta=1e-7,
patience=100,
verbose=1
)
reduce_lr = keras.callbacks.ReduceLROnPlateau(
monitor='val_mean_squared_error',
factor=0.5,
patience=20,
verbose=1,
mode='auto',
min_delta=1e-6,
cooldown=10,
min_lr=1e-8
)
EPOCHS = 5000
def train_model(model, dataset, learning_rate=1e-3, epochs=EPOCHS):
history = model.fit(
x = dataset.training[0],
y = dataset.training[1],
epochs=epochs,
shuffle=True,
validation_data=dataset.validation,
verbose=0,
callbacks=[
early_stopping,
reduce_lr
]
)
ind = np.argmin(history.history["val_mean_squared_error"])
return history.history["mean_squared_error"][ind], history.history["val_mean_squared_error"][ind]
# -
# ### Benchmarking
dim_triu = dim * (dim + 1) // 2
# +
DIM = {
"H": int(7*8 / 2),
"C": int(22 * 23 / 2)
}
number_of_atoms = 6
def run_test(descriptor, n_tests=5):
dataset = make_dataset(descriptor, dim)
errors_train, errors_val = [], []
for i in range(n_tests):
print("\n\n--> Training: " + str(i+1))
keras.backend.clear_session()
model = make_model(
descriptor.number_of_descriptors * number_of_atoms,
dim_triu
)
errors = train_model(model, dataset)
errors_train.append(errors[0])
errors_val.append(errors[1])
print("\n--> Result: Train: {:0.4e} Vaidation: {:0.4e}".format(errors[0], errors[1]))
return errors_train, errors_val
# -
# # Fetch old results
results = np.load("data/DescriptorBenchmarkResults.npy")
# # Do Benchmark
from SCFInitialGuess.descriptors.models import make_uniform
#plot_normal_model(make_uniform(5, 5, eta_max=10, eta_min=1), np.linspace(0, 5, 200))
plot_normal_model(r_model_concentrated, np.linspace(0, 5, 200))
# +
from SCFInitialGuess.descriptors.high_level import NonWeighted, AtomicNumberWeighted, ElectronegativityWeighted
from SCFInitialGuess.descriptors.coordinate_descriptors import Gaussians, PeriodicGaussians
from SCFInitialGuess.descriptors.coordinate_descriptors import IndependentAngularDescriptor, SPHAngularDescriptor
from SCFInitialGuess.descriptors.cutoffs import BehlerCutoff1, Damping
from SCFInitialGuess.descriptors.models import \
make_uniform, make_periodic_uniform
N_TESTS = 5
angular_descriptor_list = [
IndependentAngularDescriptor(
PeriodicGaussians(*make_periodic_uniform(20, 2*np.pi)),
PeriodicGaussians(*make_periodic_uniform(10, np.pi))
),
SPHAngularDescriptor(3),
SPHAngularDescriptor(6)
]
radial_descriptor_list = [
Gaussians(*r_model_concentrated),
Gaussians(*make_uniform(5, 5, eta_max=10, eta_min=1)),
Gaussians(*make_uniform(25, 5, eta_max=60, eta_min=20)),
Gaussians(*make_uniform(50, 5, eta_max=120, eta_min=40))
]
counter = 0
results = []
for HighLevelDescriptor in [NonWeighted, AtomicNumberWeighted, ElectronegativityWeighted]:
for CutOff in [BehlerCutoff1(2.5), BehlerCutoff1(5), Damping(2)]:
for radial_descriptor in radial_descriptor_list:
for angular_descriptor in angular_descriptor_list:
counter += 1
print("\n\n\n\nTesting model: {0}\n".format(counter))
descriptor = HighLevelDescriptor(
radial_descriptor,
angular_descriptor,
CutOff
)
print("dim_in: " + str(descriptor.number_of_descriptors * 6))
errors_train, errors_val = run_test(descriptor, n_tests=N_TESTS)
results.append((
[counter] * N_TESTS,
errors_train,
errors_val
))
# -
np.save("data/DescriptorBenchmarkResults.npy", results)
results = np.load("data/DescriptorBenchmarkResults.npy")
# # Plot
labels = []
x_data = []
y_train = []
y_val = []
for x, train, val in results:
labels.append("Model {0}".format(int(x[0])))
x_data += list(x)
y_train += list(train)
y_val += list(val)
np.isnan(np.array([1.0, 1 +2j, np.nan]))
# +
fig = plt.figure()
ax = plt.gca()
ax.scatter(x_data, y_train, s=30, marker="x", label="Training")
ax.scatter(x_data, y_val, s=30, marker="x", label="Validation")
ax.set_yscale('log')
plt.ylim(1e-6, 1e-1)
#xticks = np.arange(9) * 12 + 1
xticks = np.arange(0, 112, 5)
plt.xticks(xticks, xticks, rotation=-70)
#plt.xticks(np.arange(1, len(labels)+1), labels, rotation=-70, fontsize=4)
plt.xlabel("Model Nr. / 1")
plt.ylabel("MSE / 1")
#plt.legend()
plt.legend(bbox_to_anchor=(0.92, 1))
plt.tight_layout()
fig.subplots_adjust(right=0.9)
#plt.tight_layout(rect=[0, 0, 0.88, 1])
plt.savefig("/home/jo/Repos/MastersThesis/GenericDescriptors/figures/DescriptorBenchmark.pdf")
plt.show()
# -
3*3*4*3
| thesis/notebooks/Descriptors/DescriptorBenchmark.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
from keras.layers.core import Dense, Dropout, Flatten
from keras.layers.convolutional import Conv2D, MaxPooling2D
from keras.models import Sequential
from keras.models import load_model
from keras.utils import np_utils
from keras.datasets import mnist
from keras.callbacks import EarlyStopping
import numpy as np
(X_train, y_train), (X_test, y_test) = mnist.load_data()
X_train = np.array(X_train, dtype = np.float32)
X_test = np.array(X_test, dtype = np.float32)
X_train = X_train.reshape(60000, 28,28,1)
X_test = X_test.reshape(10000, 28,28,1)
y_train = np_utils.to_categorical(y_train, 10)
y_test = np_utils.to_categorical(y_test, 10)
#Using ConvNets
convmodel = Sequential()
convmodel.add(Conv2D(filters=32, padding = "same", kernel_size = (5,5), activation = "relu", input_shape = (28,28,1)))
convmodel.add(Conv2D(filters=32, padding = "same", kernel_size = (5,5), activation = "relu"))
convmodel.add(MaxPooling2D(pool_size=(2,2)))
convmodel.add(Flatten())
convmodel.add(Dense(10, activation = "softmax"))
convmodel.compile(optimizer = "adam", loss = "categorical_crossentropy", metrics=["accuracy"])
convhist = convmodel.fit(X_train, y_train, batch_size=128, epochs=10, verbose = 1, validation_data=(X_test, y_test))
score = convmodel.evaluate(X_test, y_test,verbose=0)
score
plt.plot(range(1, len(convhist.history["val_loss"])+1), convhist.history['val_loss'], 'b')
plt.xlabel('Epochs')
plt.ylabel('Validation score')
plt.show()
plt.plot(range(1, len(convhist.history["val_acc"])+1), convhist.history['val_acc'], 'b')
plt.xlabel('Epochs')
plt.ylabel('Validation score')
plt.show()
convmodel.save("SimpleConvModel.h5")
| Models/Notebooks/SimpleConvModel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
## Bibliotheken importieren
import pandas as pd
# +
## Top 1000-Liste von switch.ch importieren
directory = 'data/import/'
all_data = pd.DataFrame()
for filename in os.listdir(directory):
if filename.endswith(".csv"):
date = filename[8:14]
urls_import = pd.read_csv(directory + filename, header=0, names=["domainname", date])
all_data = pd.concat([all_data, urls_import.set_index('domainname')], axis=1, sort=False)
# -
## Datensatz transponieren
data_tp = all_data.T
## Datetimeindex erstellen
data_tp["date"] = pd.to_datetime(data_tp.index, format="%y%m")
data_tp = data_tp.set_index("date")
## zusammengefasste und formatierte Daten zwischenspeichern ##
data_tp.to_pickle('data/all_data.pkl')
## Und gleich wieder laden:
data_tp = pd.read_pickle('data/all_data.pkl')
# Zur Visualisierung in R bereitstellen:
data_tp.reset_index().to_feather('data/all_data.feather')
# Anzahl eindeutiger Domains ermitteln
len(data_tp.columns)
# ### Meta-Daten zu URLs zur späteren Klassifizierung mittels Beautifulsoup ermitteln:
## Bibliotheken importieren
from bs4 import BeautifulSoup
import requests
import pprint
import sys
# urls_metadata = []
for website in data_tp.columns[747:]:
url_metadata = []
url_metadata.append(website)
url = "http://www.{}".format(website)
try:
response = requests.get(url)
soup = BeautifulSoup(response.text)
metas = soup.find_all('meta')
title = soup.find('title')
if (title != None):
title = title.text
url_metadata.append(title)
for meta in metas:
if ('name' in meta.attrs and meta.attrs['name'] == 'description' and 'content' in meta.attrs):
description = meta.attrs['content']
url_metadata.append(description)
elif ('name' in meta.attrs and meta.attrs['name'] == 'keywords' and 'content' in meta.attrs):
keywords = meta.attrs['content']
url_metadata.append(keywords)
except requests.exceptions.RequestException as e:
print(e)
urls_metadata.append(url_metadata)
# adidas.ch bei index 746 führt zu einem Hänger in der Schleife - deshalb wurde es übersprungen.
## Metadaten zwischenspeichern ##
pd.DataFrame.from_records(urls_metadata, columns=["url", "title", "description", "keywords", "NaN", "NaN"]).to_pickle('data/metadata.pkl')
## Und gleich wieder laden:
urls_metadata_df = pd.read_pickle('data/metadata.pkl')
# ### Offene Ports der einzelnen Domains ermitteln (um allfällige andere Aufgaben hinter den Domains zu finden):
## Bibliotheken importieren
import nmap
import numpy as np
nm = nmap.PortScanner()
## Portscanner Test
scanresult = nm.scan('tagesanzeiger.ch', arguments='-Pn')
# + active=""
# #### Script zum Scannen der Ports aller Top-1000-Domains ####
# ##### Wird auf Droplet ausgeführt #####
# url_ports = pd.DataFrame(columns=['url', "ip"])
# url_list = ["tagesanzeiger.ch", "google.ch", "chiflado.ch"]
# for url in data_tp.columns:
# scanresult = nm.scan(url, arguments='-Pn')
# for ip in scanresult["scan"]:
# if "tcp" in scanresult["scan"][ip]:
# for ports, meta in scanresult["scan"][ip]["tcp"].items():
# if str(ports) + "-" + meta["name"] not in url_ports.columns.values:
# url_ports[str(ports) + "-" + meta["name"]] = ""
# x = pd.DataFrame([[""] * len(url_ports.columns.values)], columns = url_ports.columns.values)
# x["url"] = url
# x["ip"] = ip
# if "tcp" in scanresult["scan"][ip]:
# for ports, meta in scanresult["scan"][ip]["tcp"].items():
# x[str(ports) + "-" + meta["name"]] = meta["state"]
# url_ports = url_ports.append(x, ignore_index=True)
# -
## URLs und ihre offenen Ports (die via Droplet ermittelt wurden) einlesen:
url_ports = pd.read_pickle('data/url_ports.pkl')
## Domains ohne Webserver ermitteln
for website in url_ports[(url_ports["80-http"] != "open") & (url_ports["443-https"] != "open")].url:
line = url_ports[url_ports["url"] == website].T
print("{} - {}".format(website, line.index[line.iloc[:,0] == "open"].values))
url_ports
# ### Domains mit Hilfe von externer Webseite klassifizieren:
## Bibliotheken importieren
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.support.ui import Select
from selenium.webdriver.support.ui import WebDriverWait
import json
import requests
import time
import re
from pathlib import Path
## Webseiten Mithilfe von Bluecoat Sitereview klassifizieren
driver = webdriver.Chrome("./tools/chromedriver")
url = "https://sitereview.bluecoat.com/"
url_categories = []
driver.get(url)
driver.implicitly_wait(100)
for url in data_tp.columns[1226:]:
url_entry = [url]
time.sleep(1)
driver.find_element_by_id("txtSearch").send_keys(url)
time.sleep(1)
driver.find_element_by_id("btnLookupSubmit").click()
time.sleep(1)
category_entry = []
categories = driver.find_elements_by_class_name("clickable-category")
if categories:
for category in categories:
category_entry.append(category.text)
url_entry.append(category_entry)
url_categories.append(url_entry)
try:
driver.find_element_by_xpath("/html[1]/body[1]/app-root[1]/div[1]/section[1]/div[1]/ng-component[1]/form[1]/p[1]/a[1]").click()
except:
driver.find_element_by_id("btnOk").click()
driver.find_element_by_xpath("/html[1]/body[1]/app-root[1]/div[1]/section[1]/div[1]/ng-component[1]/form[1]/p[1]/a[1]").click()
driver.quit()
## DF erstellen
url_categories_df = pd.DataFrame(columns=['url', 'n_categories'])
for url in url_categories:
n_categories = 0
if (len(url[1]) > 0):
for idx, category in enumerate(url[1]):
if category not in url_categories_df.columns.values:
url_categories_df[category] = ""
n_categories = idx + 1
x = pd.DataFrame([[False] * len(url_categories_df.columns.values)], columns = url_categories_df.columns.values)
x['url'] = url[0]
x['n_categories'] = n_categories
if len(url[1]) > 0:
for idx, category in enumerate(url[1]):
x[category] = True
url_categories_df = url_categories_df.append(x)
## URL-Kategorien zwischenspeichern ##
url_categories_df.to_pickle('data/url_categories.pkl')
# und URL-Kategorien gleich wieder einlesen:
url_categories_df = pd.read_pickle('data/url_categories.pkl')
## Blick in die Daten
url_categories_df.url[url_categories_df['Bösartige ausgehende Daten/Botnets (Malicious Outbound Data/Botnets)'] == True]
## Anzahl Kategorien ermitteln
len(url_categories_df.columns) - 2
# +
## Durchschnittswerte pro Kategorie ermitteln (Beliebtheit pro Kategorie)
from datetime import datetime
import numpy as np
categories_data = pd.DataFrame(columns=["Name", "n_urls", "avg_1901", "avg_1812", "avg_1811", "avg_1810", "avg_1809", "avg_1808", "avg_1807", "avg_1806", "avg_1805", "avg_1804", "avg_1803", "avg_1802", "avg_1801", "avg_1712", "avg_1711", "avg_1710", "avg_1709", "avg_1708", "avg_1707"])
for cat in url_categories_df.columns.values[2:]:
visits = {
"avg_1901": 0,
"avg_1812": 0,
"avg_1811": 0,
"avg_1810": 0,
"avg_1809": 0,
"avg_1808": 0,
"avg_1807": 0,
"avg_1806": 0,
"avg_1805": 0,
"avg_1804": 0,
"avg_1803": 0,
"avg_1802": 0,
"avg_1801": 0,
"avg_1712": 0,
"avg_1711": 0,
"avg_1710": 0,
"avg_1709": 0,
"avg_1708": 0,
"avg_1707": 0
}
for website in (url_categories_df.url[url_categories_df[cat] == True]):
for average in visits:
visits[average] += 0 if np.isnan(data_tp[website][datetime.strftime(datetime.strptime(average[4:], '%y%m'), "%Y-%m-%d")][0]) else data_tp[website][datetime.strftime(datetime.strptime(average[4:], '%y%m'), "%Y-%m-%d")][0]
for average in visits:
visits[average] = visits[average]/url_categories_df.url[url_categories_df[cat] == True].count()
visits["Name"] = cat
visits["n_urls"] = url_categories_df.url[url_categories_df[cat] == True].count()
categories_data = categories_data.append(visits, ignore_index=True)
# -
## Metadaten zu Kategorien zwischenspeichern und für Visualisierung in R exportieren ##
categories_data.to_pickle('data/categories_data.pkl')
# und URL-Kategorien gleich wieder einlesen:
categories_data = pd.read_pickle('data/categories_data.pkl')
# ---
## duchschnittliche Besuchswerte pro Kategorie ermitteln
categories_data[categories_data["Name"] == "Phishing (Phishing)"]
### Domains mit grossen Veränderungen in den DNS-Anfrage von einem Monat zum nächsten ermitteln
big_changes = []
for website in data_tp.columns:
comparison = [0]
for key, value in data_tp[website].iteritems():
comparison.append(value)
if (comparison[-1] != 0) and (comparison[-2] != 0):
change = (100/comparison[-2]*comparison[-1]) - 100
if abs(change) > 60:
big_changes.append([key.strftime('%Y-%m'), website, change])
### Blick in die Daten
big_changes
### Bösartige URLs ermitteln
url_categories_df.url[(url_categories_df['Bösartige ausgehende Daten/Botnets (Malicious Outbound Data/Botnets)'] == True) |
(url_categories_df['Phishing (Phishing)'] == True) |
(url_categories_df['Spam (Spam)'] == True) |
(url_categories_df['Bösartige Quellen/Malnets (Malicious Sources/Malnets)'] == True) |
(url_categories_df['Verdächtig (Suspicious)'] == True) |
(url_categories_df['Raubkopien/Urheberrechtsverletzung (Piracy/Copyright Concerns)'] == True) |
(url_categories_df['Möglicherweise unerwünschte Software (Potentially Unwanted Software)'] == True)]
| 1_switch_domains.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="R9Dcw7yC_FTM" colab_type="text"
# # Feedforward Neural Networks
#
# This notebook accompanies the Intro to Deep Learning workshop run by Hackers at Cambridge
# + [markdown] id="0FSW41K5_FTN" colab_type="text"
# ## Importing Data and Dependencies
#
#
# First, we will import the dependencies - **numpy**, the python linear algebra library, **pandas** to load and preprocess the input data and **matplotlib** for visualisation purposes.
#
# We'll import the datasets using nice loader functions from **sklearn**.
# + id="w2tsb3eK_FTP" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
from sklearn.datasets import load_boston,load_breast_cancer
# + [markdown] id="ozNhjbuS_FTT" colab_type="text"
# We'll load the house dataset, using the load_boston(return_X_y=True) function. This returns our inputs $X$ and our labels $y$ as a tuple.
# If you read them in, you'll notice they're not the right dimensions, so you'll need to reshape them.
#
# It is also good practice to normalise the data (subtract mean and then divide by standard deviation) as this speeds up learning.
#
# Finally, split the data into training and test data sets (80:20 split) - we'll keep the test data to the side to evaluate our model at the end.
# + id="XUe4J9h8BRwd" colab_type="code" colab={}
X,y = load_boston(return_X_y=True)
#X,y = load_breast_cancer(return_X_y=True)
# make sure dimensions are right
X = X.T
y = np.reshape(y, (1,y.shape[0]))
#normalise the data
mean = np.mean(X, axis=1, keepdims=True)
std = np.std(X, axis=1,keepdims = True)
X -=mean
X /= std
#split data into train and test set
X_train = X[:,:4*X.shape[0]//5]
Y_train = y[:,:4*X.shape[0]//5]
X_test = X[:,4*X.shape[0]//5:]
Y_test = y[:,4*X.shape[0]//5:]
# + [markdown] id="Pe4zmPEu_FTc" colab_type="text"
# ## Creating the neural network:
#
# Having preprocessed our data into matrices, it is now time to create the feedforward neural network.
# + [markdown] id="sUfdUeJQ_FTd" colab_type="text"
# First we need to initialise parameters: the weights and biases for each layer.
#
# The weights for layer *$l$* are stored in *$ W^{[l]}$*, a *$n_l$ x $n_{(l-1)}$* matrix, where *$n_l$* is the number of units in layer *$l$*.
# We initialise the weights randomly to break symmetry, and multiply by 0.001 to ensure weights aren't too large.
#
# The biases for layer *$l$* are stored in *$ b^{[l]}$*, which is a *$n_l$ x 1* matrix.
# + id="fQI5mNln_FTf" colab_type="code" colab={}
def initialise_parameters(layers_units):
parameters = {} # create a dictionary containing the parameters
for l in range(1, len(layers_units)):
parameters['W' + str(l)] = 0.001* np.random.randn(layers_units[l],layers_units[l-1])
parameters['b' + str(l)] = np.zeros((layers_units[l],1))
return parameters
# + [markdown] id="QBvP-WeS_FTi" colab_type="text"
# The activation function $g(z)$ we will be using is the ReLU function $g(z) = max(0,z)$ in the hidden layers.
#
#
# NB: Although the ReLU function is technically non-differentiable when $z=0$, in practice we can set the derivative=0 at $z=0$.
# + id="-zS22wKC_FTj" colab_type="code" colab={}
def sigmoid(z):
return 1/(1+np.exp(-z))
def relu(z, deriv = False):
if(deriv):
return z>0
else:
return np.multiply(z, z>0)
# + [markdown] id="Xdw2iwNX_FTl" colab_type="text"
# We can now write the code for the forward propagation step.
#
# In each layer $l$ , we matrix multiply the output of the previous layer $A^{(l-1)}$ by a weight matrix $W^{(l)}$ and then add a bias term $b^{(l)}$. We then take the result $Z^{(l)}$ and apply the activation function $g(z)$ to it to get the output $A^{(l)}$. $L$ = number of layers.
#
# The equations are thus:
# $$Z^{(l)}=W^{[l]}A^{([l-1]} + b^{[l]}$$
# $$A^{[l]}=g(Z^{[l]})$$
#
# + id="G1mGl8Ol_FTl" colab_type="code" colab={}
def forward_propagation(X,parameters):
cache = {}
L = len(parameters)//2 #final layer
cache["A0"] = X #ease of notation since input = layer 0
for l in range(1, L):
cache['Z' + str(l)] = np.dot(parameters['W' + str(l)],cache['A' + str(l-1)]) + parameters['b' + str(l)]
cache['A' + str(l)] = relu(cache['Z' + str(l)])
#final layer
cache['Z' + str(L)] = np.dot(parameters['W' + str(L)],cache['A' + str(L-1)]) + parameters['b' + str(L)]
cache['A' + str(L)] =cache['Z' + str(L)]
return cache
# + [markdown] id="jwuBdFF9_FTo" colab_type="text"
# ## Implementing the Learning
#
#
# $m$ = number of training examples, $(x^{(i)},y^{(i)})$ is the $i^{th}$ training example, $a^{[L](i)}$ is the output of the final layer $L$ for that $i^{th}$ training example.
#
#
# **Mean Squared Error:**
#
# $$ J(W^{(1)}, b^{(1)},...) = \frac{1}{2m} \sum_{i=1}^{m} (a^{[L](i)} - y^{(i)})^2 $$
#
# + id="h614AF-__FTp" colab_type="code" colab={}
def cost_function(AL,Y):
m = Y.shape[1]
cost = (1/(2*m))*(np.sum(np.square(AL-Y)))
return cost
# + [markdown] id="GTMg3uoB_FTs" colab_type="text"
# ### Backpropagation:
#
# Calculating the gradients:
#
# For the final layer:
#
# $$\frac{\partial \mathcal{J} }{\partial Z^{(L)}} = A^{(L)} - Y$$
#
#
# For a general layer $l$,
#
# $$ \frac{\partial \mathcal{J} }{\partial Z^{[l]}} = \frac{\partial \mathcal{J} }{\partial A^{[l]}}*g^{'}(Z^{[l]})$$
#
# $$ \frac{\partial \mathcal{J} }{\partial W^{[l]}} = \frac{1}{m}\frac{\partial \mathcal{J} }{\partial Z^{[l]}} A^{[l-1] T} $$
#
# $$ \frac{\partial \mathcal{J} }{\partial b^{(l)}} = \frac{1}{m} \sum_{i = 1}^{m} \frac{\partial \mathcal{J} }{\partial Z^{(l)(i)}}$$
#
# $$ \frac{\partial \mathcal{J} }{\partial A^{[l-1]}} = W^{[l] T} \frac{\partial \mathcal{J} }{\partial Z^{[l]}} $$
#
# <br>
#
# If you are keen, it's a good exercise to derive them yourself or alternatively check this [post](https://mukul-rathi.github.io/2018/08/31/Backpropagation.html) for a deeper dive into the intuition behind it.
#
# + id="JMoNeqUU_FTt" colab_type="code" colab={}
def backpropagation(cache,Y,parameters):
L = len(parameters)//2
m = Y.shape[1]
grads = {}
#code up the last layer explicitly
grads["dZ" + str(L)]= cache["A" + str(L)] - Y
grads["dW" + str(L)]= (1/m)*np.dot(grads["dZ" + str(L)],cache["A" + str(L-1)].T)
grads["db" + str(L)]= (1/m)*np.sum(grads["dZ" + str(L)],axis=1,keepdims=True)
for l in range(L-1,0,-1):
grads["dA" + str(l)]= np.dot(parameters["W" + str(l+1)].T,grads["dZ" + str(l+1)])
grads["dZ" + str(l)]= np.multiply(grads["dA" + str(l)], relu(cache["Z" + str(l)], deriv = True))
grads["dW" + str(l)]= (1/m)*np.dot(grads["dZ" + str(l)],cache["A" + str(l-1)].T)
grads["db" + str(l)]= (1/m)*np.sum(grads["dZ" + str(l)],axis=1,keepdims=True)
return grads
# + [markdown] id="FtDypLdm_FTz" colab_type="text"
# ### Gradient Descent
#
# Now let's combine the functions created so far to create a model and train it using gradient descent.
#
#
# The update equations for the parameters are as follows:
# $$ W^{[l]} = W^{[l]} - \alpha \frac{\partial \mathcal{J} }{\partial W^{[l]}} $$
#
# $$ b^{[l]} = b^{[l]} - \alpha \frac{\partial \mathcal{J} }{\partial b^{[l]}} $$
#
# where $\alpha$ is the learning rate parameter.
# + id="DPXbsDfq_FTz" colab_type="code" colab={}
def train_model(X_train, Y_train,num_epochs,layers_units,learning_rate): #epoch = one cycle through the dataset
train_costs = []
parameters = initialise_parameters(layers_units)
L = len(layers_units)-1
for epoch in range (num_epochs):
#perform one cycle of forward and backward propagation to get the partial derivatives w.r.t. the weights
#and biases. Calculate the cost - used to monitor training
cache = forward_propagation(X_train,parameters)
cost = cost_function(cache["A" + str(L)],Y_train)
grads = backpropagation(cache,Y_train,parameters)
#iterate through and update the parameters using gradient descent
for l in range(1,L+1):
parameters["W" + str(l)] = parameters["W" + str(l)] - learning_rate*grads["dW" + str(l)]
parameters["W" + str(l)] = parameters["W" + str(l)] - learning_rate*grads["db" + str(l)]
#periodically output an update on the current cost and performance on the dev set for visualisation
train_costs.append(cost)
if(epoch%(num_epochs//10)==0):
print("Training the model, epoch: " + str(epoch+1))
print("Cost after epoch " + str((epoch)) + ": " + str(cost))
print("Training complete!")
#return the trained parameters and the visualisation metrics
return parameters, train_costs
# + [markdown] id="lhvigzxs_FT1" colab_type="text"
# To evaluate the model, we'll visualise the training set error over the number of iterations. We then output the final value of the evaluation metric for training and test sets. (I've used *matplotlib* to plot the graph).
# + id="z6Pfr4gB_FT1" colab_type="code" colab={}
def evaluate_model(train_costs,parameters,X_train, Y_train, X_test, Y_test):
#plot the graphs of training set error
plt.plot(np.squeeze(train_costs))
plt.ylabel('Cost')
plt.xlabel('Iterations')
plt.title("Training Set Error")
plt.show()
L = len(parameters)//2
#For train and test sets, perform a step of forward propagation to obtain the trained model's
#predictions and evaluate this
train_cache = forward_propagation(X_train,parameters)
train_AL = train_cache["A"+ str(L)]
print("The train set MSE is: "+str(cost_function(train_AL,Y_train)))
test_cache = forward_propagation(X_test,parameters)
test_AL = test_cache["A"+ str(L)]
print("The test set MSE is: "+str(cost_function(test_AL,Y_test)))
# + [markdown] id="7qHYA2ql8kX-" colab_type="text"
# ## Training the model
#
# Now it's time to train the model using our helper functions.
#
# Let's define our hyperparameters - I encourage you to play around with these - e.g. add more layers, change number of iterations.
#
# You might find the model does much worse on the test set - this is called **overfitting** - again you can read up more about it [here](https://mukul-rathi.github.io/2018/09/02/DebuggingLearningCurve.html)
#
#
# + id="vQwKmiqV_FT6" colab_type="code" colab={}
#define the hyperparameters for the model
num_epochs = 1500 #number of passes through the training set
layers_units = [X.shape[0], 1] #layer 0 is the input layer - each value in list = number of nodes in that layer
learning_rate = 1e-4
# + id="few8mbeH_FT8" colab_type="code" outputId="d1f75e8b-f9e2-4100-b5c1-80e1b8ae3266" colab={"base_uri": "https://localhost:8080/", "height": 374}
parameters, train_costs = train_model(X_train, Y_train ,num_epochs,layers_units,learning_rate)
# + id="HxRFpsk0Kd9D" colab_type="code" outputId="2b059dca-42ca-47bd-9a18-d80eb5d5bfe9" colab={"base_uri": "https://localhost:8080/", "height": 410}
evaluate_model(train_costs,parameters,X_train, Y_train, X_test, Y_test)
# + [markdown] id="2U7b7ebN9cRP" colab_type="text"
# ## Summary and Extensions:
#
# ## Summary and Extensions:
#
# You've just trained your first deep learning model! As an extension, try running the code again, but this time, use the **load_cancer()** function instead of **load boston()**. This is a dataset that classifies breast cancer as malignant/benign.
#
# Remember that sigmoid function in the lectures? We can use it to predict probabilities for classification, so all you need to do is apply it to the output of the final layer.
#
# A couple of other minor tweaks - for classification, the network uses the **cross-entropy loss** as a cost function instead of mean-square error, and you'll want to print out accuracy not MSE in the evaluation function.
#
# But the cool thing is that the network structure is the **same**! The same network, just with a sigmoid function applied to the output, can be trained on a *completely different task* and still work.
#
# That's the power of deep learning! Stay tuned for future workshops on specialised deep learning models for computer vision and natural language processing. If you want to dive deeper, head over to the [blog](http://mukul-rathi.github.io/blog.html).
# + id="SI2k1ZTt_PRV" colab_type="code" colab={}
| Reference Notebooks/FeedforwardNeuralNet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook is intended to demonstrate how vessel segmentation methods of ITKTubeTK can be applied to binary data.
# +
import itk
from itk import TubeTK as ttk
from itkwidgets import view
import numpy as np
# +
ImageType = itk.Image[itk.F, 3]
im1 = itk.imread("../Data/Binary/DSA-Binary.mha")
resample = ttk.ResampleImage.New(Input=im1, MakeIsotropic=True)
resample.Update()
im1Iso = resample.GetOutput()
# +
imMath = ttk.ImageMath.New(im1Iso)
imMath.Threshold(1,1,0,1)
im1Inv = imMath.GetOutputFloat()
distFilter = itk.DanielssonDistanceMapImageFilter.New(im1Inv)
distFilter.SetUseImageSpacing(True)
distFilter.SetInputIsBinary(True)
distFilter.SetSquaredDistance(False)
distFilter.Update()
im1Dist = distFilter.GetOutput()
view(im1Dist)
# +
numSeeds = 200
vSeg = ttk.SegmentTubes.New(Input=im1Dist)
#vSeg.SetVerbose(True)
vSeg.SetMinCurvature(0.0)
vSeg.SetMinRoundness(0.0)
vSeg.SetMinRidgeness(0.9)
vSeg.SetMinLevelness(0.0)
vSeg.SetRadiusInObjectSpace( 0.4 )
vSeg.SetSeedMask( im1Dist )
vSeg.SetSeedRadiusMask( im1Dist )
vSeg.SetOptimizeRadius(False)
vSeg.SetUseSeedMaskAsProbabilities(True)
vSeg.SetSeedExtractionMinimumProbability(0.4)
#vSeg.SetSeedMaskMaximumNumberOfPoints( numSeeds )
vSeg.ProcessSeeds()
# -
tubeMaskImage = vSeg.GetTubeMaskImage()
view(tubeMaskImage)
SOWriter = itk.SpatialObjectWriter[3].New(vSeg.GetTubeGroup())
SOWriter.SetFileName( "BinaryImageVessels.tre" )
SOWriter.Update()
| examples/BinaryImage/Demo-SegmentVesselsFromBinaryImage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Approximation Methods
# Interpolation
# -------------
#
# Given a set of *N* points $(x_i, y_i)$ with $i = 1, 2, …N$, we sometimes need a function $\hat{f}(x)$ which returns $y_i = f(x_i)$ where $x == x_i$, and which in addition provides some interpolation of the data $(x_i, y_i)$ for all $x$.
#
# The function `y0 = scipy.interpolate.interp1d(x,y,kind=’..’)` does this interpolation based on splines of varying order. Note that the function `interp1d` returns *a function* `y0` which will then interpolate the x-y data for any given $x$ when called as $y0(x)$.
#
# The code below demonstrates this, and shows the different interpolation kinds.
# +
import numpy as np
import scipy.interpolate
import pylab
# %matplotlib inline
def create_data(n):
"""Given an integer n, returns n data points
x and values y as a numpy.array."""
xmax = 5.
x = np.linspace(0, xmax, n)
y = - x**2
#make y-data somewhat irregular
y += 1.5 * np.random.normal(size=len(x))
return x, y
#main program
n = 10
x, y = create_data(n)
#use finer and regular mesh for plot
xfine = np.linspace(0.1, 4.9, n * 100)
#interpolate with piecewise constant function (p=0)
y0 = scipy.interpolate.interp1d(x, y, kind='nearest')
#interpolate with piecewise linear func (p=1)
y1 = scipy.interpolate.interp1d(x, y, kind='linear')
#interpolate with piecewise constant func (p=3)
y3 = scipy.interpolate.interp1d(x, y, kind='cubic')
print(y3(4.5))
#interpolate with cubic Hermite
yfineH = scipy.interpolate.pchip_interpolate(x, y, xfine)
pylab.figure()
pylab.plot(x, y, 'o', label='data point')
pylab.plot(xfine, y0(xfine), label='nearest')
pylab.plot(xfine, y1(xfine), label='linear')
pylab.plot(xfine, y3(xfine), label='cubic')
pylab.plot(xfine, yfineH, label='cubic Hermite')
pylab.legend(loc='upper left',bbox_to_anchor=(1.02,0.5,0.5,0.5))
pylab.xlabel('x')
pylab.ylabel('y')
# plot differences
pylab.figure()
pylab.plot(x, y+x**2, 'o', label='data point')
pylab.plot(xfine, y0(xfine)+xfine**2, label='nearest')
pylab.plot(xfine, y1(xfine)+xfine**2, label='linear')
pylab.plot(xfine, y3(xfine)+xfine**2, label='cubic')
pylab.plot(xfine, yfineH+xfine**2, label='cubic Hermite')
pylab.legend(loc='upper left',bbox_to_anchor=(1.02,0.5,0.5,0.5))
pylab.xlabel('x')
pylab.ylabel('y-y_function = y+x**2')
# -
# ## Root finding
# ### Root finding using iterative relaxation method
# +
import numpy as np
import scipy as sc
import matplotlib.pyplot as plt
# %matplotlib inline
def g(x):
return np.cos(x) # np.sin(x) # 2-np.exp(-x)#
nIter = 50
na = range(nIter)
xa = np.zeros(nIter)
x = 0.6 # initial guess
for k in na:
xa[k] = x
x = g(x)
plt.plot(na,xa)
plt.xlabel('n')
plt.ylabel('x')
print('Solution = ',xa[-1],', precision = ',np.abs(xa[-1]-xa[-2]))
# -
# ### Root finding using the bisection method
#
# First we introduce the `bisect` algorithm which is (i) robust and (ii) slow but conceptually very simple.
#
# Suppose we need to compute the roots of *f*(*x*)=*x*<sup>3</sup> − 2*x*<sup>2</sup>. This function has a (double) root at *x* = 0 (this is trivial to see) and another root which is located between *x* = 1.5 (where *f*(1.5)= − 1.125) and *x* = 3 (where *f*(3)=9). It is pretty straightforward to see that this other root is located at *x* = 2. Here is a program that determines this root numerically:
# +
from scipy.optimize import bisect
def f(x):
"""returns f(x)=x^3-2x^2. Has roots at
x=0 (double root) and x=2"""
return x ** 3 - 2 * x ** 2
# main program starts here
x = bisect(f, 1.5, 3, xtol=1e-6)
print("The root x is approximately x=%14.12g,\n"
"the error is less than 1e-6." % (x))
print("The exact error is %g." % (2 - x))
# -
# The `bisect()` method takes three compulsory arguments: (i) the function *f*(*x*), (ii) a lower limit *a* (for which we have chosen 1.5 in our example) and (ii) an upper limit *b* (for which we have chosen 3). The optional parameter `xtol` determines the maximum error of the method.
# ### Root finding using Brent method
#
# This is a classic method to find a zero of the function f on the sign changing interval [a , b]. It is a safe version of the secant method that uses inverse quadratic extrapolation. Brent’s method combines root bracketing, interval bisection, and inverse quadratic interpolation.
# +
from scipy.optimize import brentq
def f(x):
return x ** 3 - 2 * x ** 2
x = brentq(f, 1.5, 3, xtol=1e-6)
print("The root x is approximately x=%14.12g,\n"
"the error is less than 1e-6." % (x))
print("The exact error is %g." % (2 - x))
# -
# ### Root finding using the `fsolve` function
#
# A (often) better (in the sense of “more efficient”) algorithm than the bisection algorithm is implemented in the general purpose `fsolve()` function for root finding of (multidimensional) functions. This algorithm needs only one starting point close to the suspected location of the root (but is not guaranteed to converge).
#
# Here is an example:
# +
from scipy.optimize import fsolve
def f(x):
return x ** 3 - 2 * x ** 2
x0 = [0,2.5]
x = fsolve(f, x0 ) # specify starting points
print("Number of roots is",len(x))
print("The root(s) are ",x)
print("Error of the initial guess = ",x0[0]-0,x0[1]-2)
print("error : ",x[0]-0,x[1]-2)
# -
# The input to `fsolve` is the function and the array of initial locations for the roots. The return value of `fsolve` is a numpy array of length *n* for a root finding problem with *n* variables. In the example above, we have *n* = 2.
# +
import matplotlib
import matplotlib.pyplot as plt
x1=np.linspace(-1,3,100)
fig, ax = plt.subplots()
ax.plot(x1, f(x1))
ax.set(xlabel='x', ylabel='f(x)',
title='Visual check')
ax.grid()
plt.show()
# -
| Week12/Lecture08.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python tf>=2.0
# language: python
# name: tf2gpu
# ---
# ## detect_particles.py
# +
import sys
import matplotlib.pyplot as plt
import numpy as np
from tomo_encoders.misc_utils.feature_maps_vis import view_midplanes
import cupy as cp
import time
import h5py
#from recon_subvol import fbp_filter, recon_patch
from tqdm import tqdm
from tomo_encoders import DataFile, Patches
import os
from tomo_encoders.misc_utils.phantom_generators import get_sub_vols_elliptical_voids
from tomo_encoders.misc_utils.viewer import view_midplanes
from tomo_encoders.tasks.sparse_segmenter.detect_voids import wrapper_label, to_regular_grid, upsample_sub_vols
# -
# ### to-do:
# 1. make everything with numpy
# 2. use cupy, what is performance again assuming array is on cpu first?
# ### ('b' stands for binned = detector binning): e.g., vol_seg_b, sub_vols_voids_b, p3d_voids_b
BINNED_VOL_SHAPE = (1000,1000,1000)
DEFAULT_VOL_SHAPE = (3000, 3000, 3000)
DET_BINNING = 3
PATCH_SIZE_REC = (64,64,64)
N_MAX_DETECT = 200
# ## Create voids in sub-volumes and assign to output volume
vol_seg_b = np.ones(BINNED_VOL_SHAPE, dtype = np.uint8)
n_voids_gt = 3000 #len(p)
patch_size = (32,32,32)
p = Patches(BINNED_VOL_SHAPE, \
initialize_by = 'regular-grid', \
patch_size = patch_size, \
n_points = n_voids_gt)
y = get_sub_vols_elliptical_voids(patch_size, len(p), ellipse_range = (1.5, 1.8), rad_range = (5,8))
p.fill_patches_in_volume(y, vol_seg_b, TIMEIT = True)
view_midplanes(vol_seg_b)
void_size_gt = np.cbrt(np.prod(patch_size) - np.sum(y, axis = (1,2,3)))
plt.scatter(np.arange(len(p)), void_size_gt, marker ='*', color = 'black', s = 10)
plt.xlabel("void id")
plt.ylabel("void size")
# **Comments** The void size is defined as ${{volume}^{1/3}}$ where the volume of each void is counted as the sum of voxels inside the void. In above figure, the distribution of void size in the randomly generated elliptical voids is shown. Note that the patches are NOT sorted by size. So when these are assigned on a grid inside the big volume *vol_seg_b*, they are assigned to random locations.
fig, ax = plt.subplots(1,3, figsize = (8,4))
view_midplanes(vol_seg_b, ax = ax)
# ## Test wrapper for label() - connected components
sub_vols_voids_b, p3d_voids_b = wrapper_label(vol_seg_b, N_MAX_DETECT, TIMEIT = False)
plt.plot(p3d_voids_b.features[:,1])
assert len(p3d_voids_b) <= N_MAX_DETECT, "test failed: number of detected particles is greater than the maximum requested."
# assert n_detected == n_voids_gt, "test failed: number of detected particles is not equal to the ground-truth."
# ### Assert if labels are sorted by size automatically
plt.scatter(p3d_voids_b.features[:,0], p3d_voids_b.features[:,1], marker ='*', color = 'black', s = 10)
plt.xlabel("void id")
plt.ylabel("void size")
fig = plt.figure(figsize = (8,8))
ax = fig.add_subplot(projection='3d')
p3d_voids_b.plot_3D_feature(1, ax)
# check that p3d_voids and sub_vols are sorted, by checking the size
for ii in range(len(sub_vols_voids_b)):
assert np.cbrt(np.sum(sub_vols_voids_b[ii])) == p3d_voids_b.features[ii,1], "voids sub_vol and p3d_voids do not match at %i"%ii
# ## get_patches_on_grid()
# fill patches and resize
vol_lab_b = np.empty(vol_seg_b.shape, dtype = np.uint8)
p3d_voids_b.fill_patches_in_volume(sub_vols_voids_b, vol_lab_b)
fig, ax = plt.subplots(1,3, figsize = (8,4))
view_midplanes(vol_lab_b > 0, ax = ax)
p3d_grid_1_voids = to_regular_grid(sub_vols_voids_b, \
p3d_voids_b, \
PATCH_SIZE_REC,\
DET_BINNING, \
DEFAULT_VOL_SHAPE)
y_2 = upsample_sub_vols(y, 4, TIMEIT = True)
view_midplanes(y_2[2800])
view_midplanes(y[2800])
# +
# ip = 10
# sub_vol = vol_lab_b[s_voids[ip]]
# np.cbrt(np.sum(sub_vol == ip + 1)) == p3d_voids.features[ip,1], "size calculation error"
# print("void size %i"%p3d_voids.features[ip,1])
# print("void id %i"%p3d_voids.features[ip,0])
# view_midplanes(sub_vol == ip + 1)
# -
exit()
| tests/labeling/detect_voids.ipynb |
# # 9.4. Finding the equilibrium state of a physical system by minimizing its potential energy
import numpy as np
import scipy.optimize as opt
import matplotlib.pyplot as plt
# %matplotlib inline
g = 9.81 # gravity of Earth
m = .1 # mass, in kg
n = 20 # number of masses
e = .1 # initial distance between the masses
l = e # relaxed length of the springs
k = 10000 # spring stiffness
P0 = np.zeros((n, 2))
P0[:, 0] = np.repeat(e * np.arange(n // 2), 2)
P0[:, 1] = np.tile((0, -e), n // 2)
# + podoc={"output_text": "<matplotlib.figure.Figure at 0x7f93deeab668>"}
A = np.eye(n, n, 1) + np.eye(n, n, 2)
# We display a graphic representation of
# the matrix.
f, ax = plt.subplots(1, 1)
ax.imshow(A)
ax.set_axis_off()
# -
L = l * (np.eye(n, n, 1) + np.eye(n, n, 2))
for i in range(n // 2 - 1):
L[2 * i + 1, 2 * i + 2] *= np.sqrt(2)
I, J = np.nonzero(A)
def dist(P):
return np.sqrt((P[:,0]-P[:,0][:,np.newaxis])**2 +
(P[:,1]-P[:,1][:,np.newaxis])**2)
def show_bar(P):
fig, ax = plt.subplots(1, 1, figsize=(5, 4))
# Wall.
ax.axvline(0, color='k', lw=3)
# Distance matrix.
D = dist(P)
# Get normalized elongation in [-1, 1].
elong = np.array([D[i, j] - L[i, j]
for i, j in zip(I, J)])
elong_max = np.abs(elong).max()
# The color depends on the spring tension, which
# is proportional to the spring elongation.
colors = np.zeros((len(elong), 4))
colors[:, -1] = 1 # alpha channel is 1
# Use two different sequentials colormaps for
# positive and negative elongations, to show
# compression and extension in different colors.
if elong_max > 1e-10:
# We don't use colors if all elongations are
# zero.
elong /= elong_max
pos, neg = elong > 0, elong < 0
colors[pos] = plt.cm.copper(elong[pos])
colors[neg] = plt.cm.bone(-elong[neg])
# We plot the springs.
for i, j, c in zip(I, J, colors):
ax.plot(P[[i, j], 0],
P[[i, j], 1],
lw=2,
color=c,
)
# We plot the masses.
ax.plot(P[[I, J], 0], P[[I, J], 1], 'ok',)
# We configure the axes.
ax.axis('equal')
ax.set_xlim(P[:, 0].min() - e / 2,
P[:, 0].max() + e / 2)
ax.set_ylim(P[:, 1].min() - e / 2,
P[:, 1].max() + e / 2)
ax.set_axis_off()
return ax
# + podoc={"output_text": "<matplotlib.figure.Figure at 0x7157668>"}
ax = show_bar(P0)
ax.set_title("Initial configuration")
# -
def energy(P):
# The argument P is a vector (flattened matrix).
# We convert it to a matrix here.
P = P.reshape((-1, 2))
# We compute the distance matrix.
D = dist(P)
# The potential energy is the sum of the
# gravitational and elastic potential energies.
return (g * m * P[:, 1].sum() +
.5 * (k * A * (D - L)**2).sum())
energy(P0.ravel())
bounds = np.c_[P0[:2, :].ravel(),
P0[:2, :].ravel()].tolist() + \
[[None, None]] * (2 * (n - 2))
P1 = opt.minimize(energy, P0.ravel(),
method='L-BFGS-B',
bounds=bounds).x.reshape((-1, 2))
# + podoc={"output_text": "<matplotlib.figure.Figure at 0x74b6ef0>"}
ax = show_bar(P1)
ax.set_title("Equilibrium configuration")
| chapter09_numoptim/04_energy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.7 64-bit (''django'': conda)'
# language: python
# name: python_defaultSpec_1616329314126
# ---
import numpy as np
import pandas as pd
from sklearn.feature_extraction.text import CountVectorizer
sentences = [
'I come to China to travel',
'This is a car polupar in China',
# 'I love tea and Apple',
# 'The work is to write some papers in science',
# 'You forgot Jack',
# 'Jack forgot you'
]
vec = CountVectorizer(ngram_range=(1,2), stop_words=None)
X = vec.fit_transform(sentences)
# 特征向量每个维度所代表的词或元组,似乎认为单个字符不是单词,例如 'I' 没有出现
vec.get_feature_names()
len(vec.get_feature_names())
df = pd.DataFrame(X.toarray(), columns=vec.get_feature_names())
df.head()
a = (1,2)
for each in range(a[0],a[1]+1):
print(each)
# ## 2. DataLoader测试
from sklearn.model_selection import train_test_split
train_path = '../sentiment-analysis-on-movie-reviews/train.tsv'
# 该数据集是用 tab 作为分隔符
train_df = pd.read_csv(train_path, sep='\t')
train_df.head()
# 短语
train_df['Phrase']
# 类别标签
train_df['Sentiment']
X_data = train_df['Phrase'].values[:100]
y_data = train_df['Sentiment'].values[:100]
y_data = np.array(y_data).reshape(len(y_data), 1)
# 提取特征
from Vectorization import *
bow = BagOfWord(True)
X_features = bow.fit_transform(X_data)
print(X_features.shape)
print(y_data.shape)
# 数据集划分
X_train_Bow, X_test_Bow, y_train_Bow, y_test_Bow = train_test_split(X_features, y_data, test_size=0.2, random_state=1, stratify=y_data)
print(X_train_Bow.shape)
print(y_train_Bow.shape)
print(X_test_Bow.shape)
print(y_test_Bow.shape)
# ## numpy乘法测试
a = np.ones((5,1))
b = np.ones((4,1))
a.reshape(1, -1).T.shape
b.reshape(1, 4).shape
a.reshape(1, -1).T.dot(b.reshape(1, 4)).shape
np.zeros_like(a.flatten())
a.flatten().shape[0]
# ## dataloader Test
class DataLoader:
"""
对数据集进行划分batch,根据batch_size调整是SGD、BGD还是MGBD
"""
def __init__(self, dataset, labels, batch_size=1):
"""
dataset - (bs, feature_size)
labels - (bs,)
"""
self.batch_size = batch_size
self.dataset_len, self.feature_size = dataset.shape
self.loader_len = self.dataset_len // self.batch_size
# 打乱数据
rand_index = np.arange(dataset_len)
np.random.shuffle(rand_index)
# 构建loader
self.data_loader = np.zeros((self.loader_len, self.batch_size, self.feature_size))
self.label_loader = np.zeros((self.loader_len, self.batch_size))
for i, di in enumerate(range(0, dataset_len-batch_size+1, batch_size)):
self.data_loader[i] = dataset[di:di+batch_size]
self.label_loader[i] = labels[di:di+batch_size]
def __len__(self):
"""
重载 len() 函数
"""
return self.loader_len
def __getitem__(self, index):
"""
重载[],返回第 index batch的data和label
"""
return self.data_loader[index], self.label_loader[index]
y_train_Bow = y_train_Bow.reshape(-1)
dataloader = DataLoader(X_train_Bow, y_train_Bow, batch_size=15)
len(dataloader)
for x, y in dataloader:
print(x.shape)
print(y.shape)
a = np.array([1,2,3])
b = np.array([2,3,4])
print(a.dot(b))
a * b
np.sum(np.power(b-a, 2))
t = (b-a).dot(a)
print(t)
b-a-t
| p0/Some_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TREC-6
# # DistilBERT finetuning
# ## Librairy
# +
# # !pip install transformers==4.8.2
# # !pip install datasets==1.7.0
# +
import os
import time
import pickle
import numpy as np
import torch
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, recall_score, precision_score, f1_score
from transformers import DistilBertTokenizer, DistilBertTokenizerFast
from transformers import DistilBertForSequenceClassification, AdamW
from transformers import Trainer, TrainingArguments
from transformers import EarlyStoppingCallback
from transformers.data.data_collator import DataCollatorWithPadding
from datasets import load_dataset, Dataset, concatenate_datasets
# -
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
device
# ## Global variables
BATCH_SIZE = 24 # cf. paper Sun et al.
NB_EPOCHS = 4 # cf. paper Sun et al.
CURRENT_PATH = '~/Results/BERT_finetune' # put your path here
RESULTS_FILE = os.path.join(CURRENT_PATH, 'trec-6_results.pkl')
RESULTS_DIR = os.path.join(CURRENT_PATH,'trec-6/')
CACHE_DIR = '~/Data/huggignface/' # put your path here
# ## Dataset
# +
# download dataset
raw_datasets = load_dataset('trec', cache_dir=CACHE_DIR)
# -
raw_datasets
# +
# tokenize
tokenizer = DistilBertTokenizer.from_pretrained("distilbert-base-uncased")
def tokenize_function(examples):
return tokenizer(examples["text"], padding=True, truncation=True)
tokenized_datasets = raw_datasets.map(tokenize_function, batched=True)
tokenized_datasets = tokenized_datasets.rename_column('label-coarse', 'labels') # 'label-fine'
tokenized_datasets.set_format(type='torch', columns=['input_ids', 'attention_mask', 'labels'])
train_dataset = tokenized_datasets["train"].shuffle(seed=42)
train_val_datasets = train_dataset.train_test_split(train_size=0.8)
train_dataset = train_val_datasets['train']
val_dataset = train_val_datasets['test']
test_dataset = tokenized_datasets["test"].shuffle(seed=42)
# +
# get number of labels
num_labels = len(set(train_dataset['labels'].tolist()))
num_labels
# -
# ## Model
# #### Model
model = DistilBertForSequenceClassification.from_pretrained("distilbert-base-uncased", num_labels=num_labels)
model.to(device)
# #### Training
# +
training_args = TrainingArguments(
# output
output_dir=RESULTS_DIR,
# params
num_train_epochs=NB_EPOCHS, # nb of epochs
per_device_train_batch_size=BATCH_SIZE, # batch size per device during training
per_device_eval_batch_size=BATCH_SIZE, # cf. paper Sun et al.
learning_rate=2e-5, # cf. paper Sun et al.
# warmup_steps=500, # number of warmup steps for learning rate scheduler
warmup_ratio=0.1, # cf. paper Sun et al.
weight_decay=0.01, # strength of weight decay
# # eval
evaluation_strategy="steps", # cf. paper Sun et al.
eval_steps=50, # 20 # cf. paper Sun et al.
# evaluation_strategy='no', # no more evaluation, takes time
# log
logging_dir=RESULTS_DIR+'logs',
logging_strategy='steps',
logging_steps=50, # 20
# save
save_strategy='steps',
save_total_limit=1,
# save_steps=20, # default 500
load_best_model_at_end=True, # cf. paper Sun et al.
metric_for_best_model='eval_loss'
)
# -
def compute_metrics(p):
pred, labels = p
pred = np.argmax(pred, axis=1)
accuracy = accuracy_score(y_true=labels, y_pred=pred)
return {"val_accuracy": accuracy}
trainer = Trainer(
model=model,
args=training_args,
tokenizer=tokenizer,
train_dataset=train_dataset,
eval_dataset=val_dataset,
# compute_metrics=compute_metrics,
# callbacks=[EarlyStoppingCallback(early_stopping_patience=5)]
)
results = trainer.train()
training_time = results.metrics["train_runtime"]
training_time_per_epoch = training_time / training_args.num_train_epochs
training_time_per_epoch
trainer.save_model(os.path.join(RESULTS_DIR, 'checkpoint_best_model'))
# ## Results
# +
# load model
model_file = os.path.join(RESULTS_DIR, 'checkpoint_best_model')
finetuned_model = DistilBertForSequenceClassification.from_pretrained(model_file, num_labels=num_labels)
finetuned_model.to(device)
finetuned_model.eval()
# compute test acc
test_trainer = Trainer(finetuned_model, data_collator=DataCollatorWithPadding(tokenizer))
raw_preds, labels, _ = test_trainer.predict(test_dataset)
preds = np.argmax(raw_preds, axis=1)
test_acc = accuracy_score(y_true=labels, y_pred=preds)
# save results
results_d = {}
results_d['accuracy'] = test_acc
results_d['training_time'] = training_time
# -
results_d
# +
# save results
with open(RESULTS_FILE, 'wb') as fh:
pickle.dump(results_d, fh)
# +
# # load results
# with open(RESULTS_FILE, 'rb') as fh:
# results_d = pickle.load(fh)
# results_d
# -
| notebooks_paper_2022/DistilBERT_finetuning_v2/DistilBERT_finetune_TREC-6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="7a4n8t6oUIW1"
# # Importación
# + id="GWfiikYqYU1P" colab={"base_uri": "https://localhost:8080/"} outputId="44f2adaa-f1e4-4f62-c913-11720833496d"
# Librería Yahoo Finance
# !pip install yfinance
# + id="bdyRFV0eP2FV"
import pandas as pd
import yfinance as yf
from datetime import datetime
from dateutil.relativedelta import relativedelta
from google.colab import auth
auth.authenticate_user()
import gspread
from oauth2client.client import GoogleCredentials
# + colab={"base_uri": "https://localhost:8080/"} id="_s20BRMsUEHh" outputId="e56a334a-014f-4714-c8fe-0d7ddceb6c26"
# Importar el fichero import_data_stock_prediction.py, el cual se conecta a Yahoo Finance e importa los datos
from google.colab import drive
drive.mount('/content/drive')
execfile('/content/drive/MyDrive/StockPrediction/import_data_stock_prediction.py')
# + colab={"base_uri": "https://localhost:8080/"} id="KWhveGOYZ13K" outputId="a55c35d8-501e-42c0-95b5-f1a1be94c9b7"
# Importar el ficher lstm_stock_prediction.py, el cual contiene el modelo predictivo LSTM
drive.mount('/content/drive', force_remount=True)
execfile('/content/drive/MyDrive/StockPrediction/lstm_stock_prediction.py')
# + colab={"base_uri": "https://localhost:8080/"} id="a9x5osNe2fyf" outputId="d366e65b-e917-41d9-ee7f-7394d31b36b3"
# Se comprueba que se tiene activado la aceleración con GPU
print("GPU available: ", tf.config.list_physical_devices('GPU'))
# + [markdown] id="Wp0OYv_6UJlO"
# # Leer de Google Sheets
# + id="A2L68NpiQGBG"
def OpenSheet(sheetName):
gc = gspread.authorize(GoogleCredentials.get_application_default())
worksheet = gc.open('Cartera StockPrediction').worksheet(sheetName)
# get_all_values gives a list of rows.
rows = worksheet.get_all_values()
# Convert to a DataFrame and render.
stock_wallet = pd.DataFrame.from_records(rows)
stock_wallet.columns = stock_wallet.iloc[0]
stock_wallet = stock_wallet.iloc[1: , :]
return worksheet, stock_wallet
# + colab={"base_uri": "https://localhost:8080/"} id="KV1WTFfvJk3W" outputId="37cf4a45-7d55-4283-8842-9d82c2257394"
worksheet, stock_wallet = OpenSheet('Prediccion 27122021')
stock_wallet.iloc[0:]['Ticker'].head(5)
# + [markdown] id="5N87YTs8K4oC"
# # Analizar una compañía
# + id="i0EATJd5K6zS"
tickerSymbol = 'AAPL'
# Obtenemos los datos desde el día actual hasta hace 5 años
one_year_ago = datetime.now() - relativedelta(years=5)
start_date = one_year_ago.strftime('%Y-%m-%d')
#one_year_ago = datetime.now() - relativedelta(years=1)
end_date = datetime.now().strftime('%Y-%m-%d')
# Se obtiene el datafram de la empresa seleccionada y en el rango de fechas
tickerDf = GetStockDataByTicker(tickerSymbol, start_date, end_date)
# + colab={"base_uri": "https://localhost:8080/", "height": 231} id="vIfW64wxLNAR" outputId="ad941405-75cc-439d-aba9-b7a10583d2cc"
GetStockInformationByTicker(tickerSymbol, start_date, end_date)
# + colab={"base_uri": "https://localhost:8080/", "height": 499} id="aI9k9EtPLmKs" outputId="dbb7cc83-9932-4959-c75c-b6e3f90a3c6d"
Graph_StockEvolutionByTime(tickerSymbol, tickerDf, 180)
# + colab={"base_uri": "https://localhost:8080/", "height": 539} id="3DL08LMFLNgP" outputId="33685e67-f06c-45dd-b8b4-94b70194594c"
x_train, y_train, dataset, training_data_len, scaler, scaled_data = TransformData(tickerDf, split_data)
LSTM_model = LSTM_Model(x_train, model_dropout, model_neuron, model_learning_rate)
LSTM_history = LSTM_fit(LSTM_model, x_train, y_train, model_epochs, model_batch_size)
predictions, y_test = LSTM_Predictions(LSTM_model, training_data_len, scaled_data, scaler, dataset)
LSTM_PlotPredictions(tickerDf.filter(['Adj Close']), training_data_len, predictions)
print("RSME = ", LSTM_RMSE(predictions, y_test))
# + [markdown] id="-VycV2LSWxz1"
# # Predicción en Bucle
# + [markdown] id="UyidXA-ML8kV"
# Parámetros de predicción
# + id="TQaMs3MRL7ZT"
train_last_days = 60
split_data = 0.8 #80% train, 20% test
model_neuron = 50
model_epochs = 25
model_batch_size = 32
model_learning_rate = 1e-4
model_dropout = 0.2
# + id="s3DigXmnW9hH"
from random import randint
from time import sleep
# last_company = Ult numeración del excel - 2 casillas
def PredictWeek_SP500(sheet_name,
last_company,
train_last_days = 60,
split_data = 0.8,
model_neuron = 50,
model_epochs = 25,
model_batch_size = 32,
model_learning_rate = 1e-4,
model_dropout = 0.2):
# Se abre la hoja de cálculo
worksheet, stock_wallet = OpenSheet(sheet_name)
# Obtenemos los datos desde el día actual hasta hace 5 años
five_year_ago = datetime.now() - relativedelta(years=5)
start_date = five_year_ago.strftime('%Y-%m-%d')
end_date = datetime.now().strftime('%Y-%m-%d')
row = last_company + 2
for tickerSymbol in stock_wallet.iloc[last_company:]['Ticker']:
tickerSymbol = tickerSymbol.replace(".", "-")
# Temporizador aleatorio de parada para evitar que Google Colab aborte la sesión
#sleep(randint(10,100))
# Se obtiene el datafram de la empresa seleccionada y en el rango de fechas
tickerDf = GetStockDataByTicker(tickerSymbol, start_date, end_date)
# Tratamiento de datos
x_train, y_train, dataset, training_data_len, scaler, scaled_data = TransformData(tickerDf, split_data)
# Entrenamiento
LSTM_model = LSTM_Model(x_train, model_dropout, model_neuron, model_learning_rate)
LSTM_history = LSTM_fit(LSTM_model, x_train, y_train, model_epochs, model_batch_size)
# Testeo
predictions, y_test = LSTM_Predictions(LSTM_model, training_data_len, scaled_data, scaler, dataset)
rsme = LSTM_RMSE(predictions, y_test)
# Actualizar lista de valores del Excel
worksheet.update_cell(row, 5, str(rsme).replace('.', ',')) # RMSE
# Predicción de la última semana
column = 0
for predicted_day in reversed(range(1,6)):
day, real, predicted = LSTM_PredictNextDay(tickerDf, scaler, LSTM_model, train_last_days, predicted_day)
if predicted_day != 0:
worksheet.update_cell(row, 7+column, str(real).replace('.', ',')) # Real
worksheet.update_cell(row, 8+column, str(predicted).replace('.', ',')) # Predecido
else:
worksheet.update_cell(row, 8+column, str(predicted).replace('.', ',')) # Predecido
column += 3
row += 1
# + id="7FO_aXRXC7Vz"
sheet_name = 'Prediccion 27122021'
last_company = 476 - 2
PredictWeek_SP500(sheet_name, last_company) #train_last_days, split_data, model_neuron, model_epochs, model_batch_size, model_learning_rate, model_dropout
# + [markdown] id="jRrkQwaYODX-"
# # Actualizar Estudio Cartera o SP_500
#
# + id="mzqN37wHttVp"
def UpdateExcelByCompany(sheetName, company, row):
# Fechas de Inicio y Fin para obtener los datos.
five_years_ago = datetime.now() - relativedelta(years=5)
start_date = five_years_ago.strftime('%Y-%m-%d')
end_date = datetime.now().strftime('%Y-%m-%d')
# Obtenemos los datos por companía y fechas
newCompany = GetStockInformationByTicker(company, start_date, end_date)
# Se accede al Excel
gc = gspread.authorize(GoogleCredentials.get_application_default())
worksheet = gc.open('Cartera StockPrediction').worksheet(sheetName)
# Se actualizan los valores del Excel
worksheet.update_cell(row, 2, str(newCompany['Short Name'][0])) # Short Name
worksheet.update_cell(row, 3, str(newCompany['Sector'][0])) # Sector
worksheet.update_cell(row, 4, str(newCompany['Industry'][0])) # Industry
worksheet.update_cell(row, 5, str(newCompany['Recommendation'][0])) # Recommendation
worksheet.update_cell(row, 6, str(newCompany['Recommendation Mean'][0]).replace('.', ',')) # Recommendation Mean
worksheet.update_cell(row, 8, str(newCompany['Target Low Price'][0]).replace('.', ',')) # Target Low Price
worksheet.update_cell(row, 9, str(newCompany['Target High Price'][0]).replace('.', ',')) # Target High Price
worksheet.update_cell(row, 10, str(newCompany['Target Mean Price'][0]).replace('.', ',')) # Target Mean Price
worksheet.update_cell(row, 11, str(newCompany['52 Week Change'][0]).replace('.', ',')) # 52 Week Change
worksheet.update_cell(row, 14, str(newCompany['EMA10'][0]).replace('.', ',')) # EMA10
worksheet.update_cell(row, 15, str(newCompany['SMA20'][0]).replace('.', ',')) # SMA20
worksheet.update_cell(row, 16, str(newCompany['SMA50'][0]).replace('.', ',')) # SMA50
worksheet.update_cell(row, 17, str(newCompany['SMA100'][0]).replace('.', ',')) # SMA100
worksheet.update_cell(row, 18, str(newCompany['SMA200'][0]).replace('.', ',')) # SMA200
worksheet.update_cell(row, 25, str(newCompany['EBITDA'][0]).replace('.', ',')) # EBITDA
worksheet.update_cell(row, 26, str(newCompany['EBITDA Margins'][0]).replace('.', ',')) # EBITDA Margins
worksheet.update_cell(row, 27, str(newCompany['Fecha Actualización'][0])) # Fecha Actualización
# + id="VB33NGAfOMUG"
from random import randint
from time import sleep
# Nombre de la hoja donde escribir los datos
sheetName = '<NAME>'
# sheetName = 'SP_500'
worksheet, stock_wallet = OpenSheet(sheetName)
last_company = 0
row = last_company + 2
for company in stock_wallet.iloc[last_company:]['Ticker']:
wait_time = randint(7,30)
sleep(wait_time)
company = company.replace(".", "-")
UpdateExcelByCompany(sheetName, company, row)
row += 1
| ipynb y Resultados/5_Sheets_Stock_Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Scalable Multitask GP Regression (w/ KISS-GP)
#
# This notebook demonstrates how to perform a scalable multitask regression.
#
# It does everything that the [standard multitask GP example](./Multitask_GP_Regression.ipynb) does, but using the SKI scalable GP approximation. This can be used on much larger datasets (up to 100,000+ data points).
#
# For more information on SKI, check out the [scalable regression example](../04_Scalable_GP_Regression_1D/KISSGP_Regression_1D.ipynb).
# +
import math
import torch
import gpytorch
from matplotlib import pyplot as plt
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# -
# ### Set up training data
#
# In the next cell, we set up the training data for this example. We'll be using 1000 regularly spaced points on [0,1] which we evaluate the function on and add Gaussian noise to get the training labels.
#
# We'll have two functions - a sine function (y1) and a cosine function (y2).
#
# For MTGPs, our `train_targets` will actually have two dimensions: with the second dimension corresponding to the different tasks.
# +
train_x = torch.linspace(0, 1, 1000)
train_y = torch.stack([
torch.sin(train_x * (2 * math.pi)) + torch.randn(train_x.size()) * 0.2,
torch.cos(train_x * (2 * math.pi)) + torch.randn(train_x.size()) * 0.2,
], -1)
# -
# ## Set up the model
#
# The model should be somewhat similar to the `ExactGP` model in the [simple regression example](../01_Simple_GP_Regression/Simple_GP_Regression.ipynb).
#
# The differences:
#
# 1. We're going to wrap ConstantMean with a `MultitaskMean`. This makes sure we have a mean function for each task.
# 2. Rather than just using a RBFKernel, we're using that in conjunction with a `MultitaskKernel`. This gives us the covariance function described in the introduction.
# 3. We're using a `MultitaskMultivariateNormal` and `MultitaskGaussianLikelihood`. This allows us to deal with the predictions/outputs in a nice way. For example, when we call MultitaskMultivariateNormal.mean, we get a `n x num_tasks` matrix back.
#
# In addition, we're going to wrap the RBFKernel in a `GridInterpolationKernel`.
# This approximates the kernel using SKI, which makes GP regression very scalable.
#
# You may also notice that we don't use a ScaleKernel, since the IndexKernel will do some scaling for us. (This way we're not overparameterizing the kernel.)
# +
class MultitaskGPModel(gpytorch.models.ExactGP):
def __init__(self, train_x, train_y, likelihood):
super(MultitaskGPModel, self).__init__(train_x, train_y, likelihood)
# SKI requires a grid size hyperparameter. This util can help with that
grid_size = gpytorch.utils.grid.choose_grid_size(train_x)
self.mean_module = gpytorch.means.MultitaskMean(
gpytorch.means.ConstantMean(), num_tasks=2
)
self.covar_module = gpytorch.kernels.MultitaskKernel(
gpytorch.kernels.GridInterpolationKernel(
gpytorch.kernels.RBFKernel(), grid_size=grid_size, num_dims=1,
), num_tasks=2, rank=1
)
def forward(self, x):
mean_x = self.mean_module(x)
covar_x = self.covar_module(x)
return gpytorch.distributions.MultitaskMultivariateNormal(mean_x, covar_x)
likelihood = gpytorch.likelihoods.MultitaskGaussianLikelihood(num_tasks=2)
model = MultitaskGPModel(train_x, train_y, likelihood)
# -
# ## Train the model hyperparameters
# +
# Find optimal model hyperparameters
model.train()
likelihood.train()
# Use the adam optimizer
optimizer = torch.optim.Adam([
{'params': model.parameters()}, # Includes GaussianLikelihood parameters
], lr=0.1)
# "Loss" for GPs - the marginal log likelihood
mll = gpytorch.mlls.ExactMarginalLogLikelihood(likelihood, model)
n_iter = 50
for i in range(n_iter):
optimizer.zero_grad()
output = model(train_x)
loss = -mll(output, train_y)
loss.backward()
print('Iter %d/%d - Loss: %.3f' % (i + 1, n_iter, loss.item()))
optimizer.step()
# -
# ## Make predictions with the model
# +
# Set into eval mode
model.eval()
likelihood.eval()
# Initialize plots
f, (y1_ax, y2_ax) = plt.subplots(1, 2, figsize=(8, 3))
# Test points every 0.02 in [0,1]
# Make predictions
with torch.no_grad(), gpytorch.fast_pred_var():
test_x = torch.linspace(0, 1, 51)
observed_pred = likelihood(model(test_x))
# Get mean
mean = observed_pred.mean
# Get lower and upper confidence bounds
lower, upper = observed_pred.confidence_region()
# This contains predictions for both tasks, flattened out
# The first half of the predictions is for the first task
# The second half is for the second task
# Plot training data as black stars
y1_ax.plot(train_x.detach().numpy(), train_y[:, 0].detach().numpy(), 'k*')
# Predictive mean as blue line
y1_ax.plot(test_x.numpy(), mean[:, 0].numpy(), 'b')
# Shade in confidence
y1_ax.fill_between(test_x.numpy(), lower[:, 0].numpy(), upper[:, 0].numpy(), alpha=0.5)
y1_ax.set_ylim([-3, 3])
y1_ax.legend(['Observed Data', 'Mean', 'Confidence'])
y1_ax.set_title('Observed Values (Likelihood)')
# Plot training data as black stars
y2_ax.plot(train_x.detach().numpy(), train_y[:, 1].detach().numpy(), 'k*')
# Predictive mean as blue line
y2_ax.plot(test_x.numpy(), mean[:, 1].numpy(), 'b')
# Shade in confidence
y2_ax.fill_between(test_x.numpy(), lower[:, 1].numpy(), upper[:, 1].numpy(), alpha=0.5)
y2_ax.set_ylim([-3, 3])
y2_ax.legend(['Observed Data', 'Mean', 'Confidence'])
y2_ax.set_title('Observed Values (Likelihood)')
None
# -
| examples/03_Multitask_GP_Regression/Multitask_GP_Regression_Scalable_With_KISSGP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import time
from volatility import GARCH_MIDAS
from weights import beta_
# +
df = pd.read_csv('C:/Users/peter/Desktop/volatility-forecasting/data/Stocks/AMD.csv')
df['Unnamed: 0'] = df['Unnamed: 0'].astype("datetime64[ms]")
df = df.rename(columns = {"Unnamed: 0": 'Date', 'open': 'Open', 'high': 'High', 'low': 'Low', 'close': 'Close', 'volume': 'Volume'})
df['LogReturn'] = np.log(df.Close).diff().fillna(0)*10
df['Volatility'] = df['LogReturn'] ** 2.0
df = df.iloc[1023:3019, :]
df.set_index(df.Date, inplace = True)
vix = pd.read_csv('C:/Users/peter/Desktop/volatility-forecasting/data/Macroeconomic/VIXCLS.csv')
vix = vix[vix.VIXCLS != '.'].reset_index(drop=True)
vix['DATE'] = vix['DATE'].astype("datetime64[ms]")
vix = vix.rename(columns = {'DATE': 'Date', 'VIXCLS': 'VIX'})
vix = vix[(vix.Date >= df.Date.min()) & (vix.Date <= df.Date.max())]
vix['VIX'] = vix['VIX'].astype('float64')
vix.set_index(vix.Date, inplace = True)
cfnai = pd.read_csv('C:/Users/peter/Desktop/volatility-forecasting/data/Macroeconomic/CFNAI.csv')
cfnai = cfnai[cfnai.CFNAI != '.'].reset_index(drop = True)
cfnai['DATE'] = cfnai['DATE'].astype('datetime64[ms]')
cfnai = cfnai.rename(columns = {'DATE': 'Date', 'CFNAI': 'NAI'})
cfnai = cfnai[(cfnai.Date >= df.Date.min()) & (cfnai.Date <= df.Date.max())]
cfnai['NAI'] = cfnai['NAI'].astype('float64')
cfnai.set_index(cfnai.Date, inplace = True)
indpro = pd.read_csv('C:/Users/peter/Desktop/volatility-forecasting/data/Macroeconomic/INDPRO.csv')
indpro = indpro[indpro.Value != '.'].reset_index(drop = True)
indpro['Date'] = indpro['Date'].astype('datetime64[ms]')
indpro = indpro.rename(columns = {'Date': 'Date', 'Value': 'IND'})
indpro = indpro[(indpro.Date >= df.Date.min()) & (indpro.Date <= df.Date.max())]
indpro['IND'] = indpro['IND'].astype('float64')
indpro['IND'] = np.log(indpro.IND).diff().fillna(0.0)
indpro.set_index(indpro.Date, inplace = True)
ppi = pd.read_csv('C:/Users/peter/Desktop/volatility-forecasting/data/Macroeconomic/PPIACO.csv')
ppi = ppi[ppi.Value != '.'].reset_index(drop = True)
ppi['Date'] = ppi['Date'].astype('datetime64[ms]')
ppi = ppi.rename(columns = {'Date': 'Date', 'Value': 'PPI'})
ppi = ppi[(ppi.Date >= df.Date.min()) & (ppi.Date <= df.Date.max())]
ppi['PPI'] = ppi['PPI'].astype('float64')
ppi['PPI'] = np.log(ppi['PPI']).diff().fillna(0.0)
ppi.set_index(ppi.Date, inplace = True)
houst = pd.read_csv('C:/Users/peter/Desktop/volatility-forecasting/data/Macroeconomic/HOUST.csv')
houst = houst[houst.HOUST != '.'].reset_index(drop = True)
houst['DATE'] = houst['DATE'].astype('datetime64[ms]')
houst = houst.rename(columns = {'DATE': 'Date', 'HOUST': 'HST'})
houst = houst[(houst.Date >= df.Date.min()) & (houst.Date <= df.Date.max())]
houst['HST'] = houst['HST'].astype('float64')
houst['HST'] = np.log(houst['HST']).diff().fillna(0.0)
houst.set_index(houst.Date, inplace = True)
# -
data = pd.concat([df.LogReturn, df.Volatility], axis = 1)
data = pd.concat([data, vix.VIX], axis = 1)
data = pd.concat([data, cfnai.NAI], axis = 1)
data = pd.concat([data, indpro.IND], axis = 1)
data = pd.concat([data, ppi.PPI], axis = 1)
data = pd.concat([data, houst.HST], axis = 1)
data = data.fillna(method = 'ffill')
data = data.fillna(0.0)
ret = data.iloc[1:, 0] * 10
data = data.iloc[1:, 2:]
# +
res = ['', 'pos', 'pos', 'pos', '', 'pos', 'pos', 'pos', 'pos', 'pos', 'pos', 'pos']
model = GARCH_MIDAS()
start = time.time()
model.fit(res, data, ret)
print(time.time() - start, 'ms to run')
# -
fit = model.predict(data, ret)
plt.figure(figsize=(15, 5))
plt.plot(data.index, fit / 100)
plt.title('Volatility')
plt.tight_layout()
plt.show()
plt.figure(figsize=(15, 5))
plt.plot(model.tau)
plt.title('Long-term Component (Tau)')
plt.tight_layout()
plt.show()
| Examples/M-GARCH.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Causal Impact (tfCausalImpact) sample
# ## Ref.
# https://github.com/google/CausalImpact
# https://github.com/WillianFuks/tfcausalimpact
#
# https://towardsdatascience.com/implementing-causal-impact-on-top-of-tensorflow-probability-c837ea18b126
# ## Requirements:
# ```
# python{3.6, 3.7, 3.8}
# matplotlib
# jinja2
# tensorflow>=2.3.0
# tensorflow_probability>=0.11.0
# ```
pip install tfcausalimpact
# ## Examples 1
# Here's a simple example (which can also be found in the original Google's R implementation) running in Python:
# 施策効果を、施策リリース前後の数値をもとに推定・可視化するサンプル
# +
import pandas as pd
from causalimpact import CausalImpact
data = pd.read_csv('https://raw.githubusercontent.com/WillianFuks/tfcausalimpact/master/tests/fixtures/arma_data.csv')[['y', 'X']]
# -
data.info()
data.describe()
# +
# treatment effect
data['y'][70:] += 5
# treatment release period
pre_period = [0, 69]
post_period = [70, 99]
# -
ci = CausalImpact(data, pre_period, post_period)
print(ci.summary())
print(ci.summary(output='report'))
ci.plot()
data.plot()
# ## Examples 2
# 「Paypal のプラットフォームにおいて暗号資産を受け入れる」 というニュースが、ビットコイン価格へと与えた影響の推定・可視化
# (参考として、大手テクノロジー企業の株価も合わせてplot)
#
# Ref.
# https://towardsdatascience.com/implementing-causal-impact-on-top-of-tensorflow-probability-c837ea18b126
pip install pandas-datareader
# +
import datetime
import pandas_datareader as pdr
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
btc_data = pdr.get_data_yahoo(['BTC-USD'],
start=datetime.datetime(2018, 1, 1),
end=datetime.datetime(2020, 12, 3))['Close']
btc_data = btc_data.reset_index().drop_duplicates(subset='Date', keep='last').set_index('Date').sort_index()
btc_data = btc_data.resample('D').fillna('nearest')
X_data = pdr.get_data_yahoo(['TWTR', 'GOOGL', 'AAPL', 'MSFT', 'AMZN', 'FB', 'GOLD'],
start=datetime.datetime(2018, 1, 1),
end=datetime.datetime(2020, 12, 2))['Close']
X_data = X_data.reset_index().drop_duplicates(subset='Date', keep='last').set_index('Date').sort_index()
X_data = X_data.resample('D').fillna('nearest')
data = pd.concat([btc_data, X_data], axis=1)
data.dropna(inplace=True)
data = data.resample('W-Wed').last() # Weekly is easier to process. We select Wednesday to 2020-10-21 is available.
data = data.astype(np.float32)
np.log(data).plot(figsize=(15, 12))
plt.axvline('2020-10-14', 0, np.max(data['BTC-USD']), lw=2, ls='--', c='red', label='PayPal Impact')
plt.legend(loc='upper left')
# +
from causalimpact import CausalImpact
pre_period=['2018-01-03', '2020-10-14']
post_period=['2020-10-21', '2020-11-25']
ci = CausalImpact(data, pre_period, post_period)
print(ci.summary())
print(ci.summary(output='report'))
ci.plot()
# -
| examples/causal_inference/.ipynb_checkpoints/causal_impact-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py3]
# language: python
# name: conda-env-py3-py
# ---
# # TreeCorr Tutorial
# This tutorial is intended to give you a rough overview of TreeCorr functionality. It doesn't cover every feature by a long shot, but it should hopefully give you enough of a flavor of how TreeCorr works to be able to find more information in the full [TreeCorr documentation](http://rmjarvis.github.io/TreeCorr/html/index.html)
#
# ---
#
# This script lives in the [TreeCorr/tests](https://github.com/rmjarvis/TreeCorr/tree/master/tests) directory, and should run successfully if executed from there. The [Aardvark.fit](https://github.com/rmjarvis/TreeCorr/wiki/Aardvark.fit) file we use here will be automatically downloaded by the test suite, so if you run nosetests in that directory, it should be downloaded for you.
# First some imports that we'll use below
from __future__ import print_function
import treecorr
import fitsio
import numpy
import time
import pprint
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
# For this demo, we'll use input data from a cosmological simulation for DES, called Aardvark, of which the [Aardvark.fit](https://github.com/rmjarvis/TreeCorr/wiki/Aardvark.fit) file we are using is just a subset.
# First, let's take a look at what's in that file.
file_name = 'data/Aardvark.fit'
data = fitsio.read(file_name)
print(data.dtype) # Includes ra, dec, redshift, and lensing observables
print(data.shape) # 390K objects
# ### Using a configuration file
#
# There are two modes for running treecorr. You can either write a config file that specifies
# the nature of the data and what you want TreeCorr to calculate. Or you can do each step
# yourself within python. We'll start with the first mode by running a config file that will
# compute a shear-shear correlation function on these data. This [config file](https://github.com/rmjarvis/TreeCorr/blob/master/tests/Aardvark.yaml) is located in
# the [treecorr/tests](https://github.com/rmjarvis/TreeCorr/tree/master/tests) directory.
config_file = 'Aardvark.yaml'
with open(config_file) as fin:
print(''.join(fin.readlines()))
# The corr2 executable installed when you install treecorr will use this config file to process the data and output the correlation to an ASCII file (output/Aardvark.out). From the shell prompt, you would run:
# ```
# $ corr2 Aardvark.yaml
# ```
# The same thing can be done from within python, so we'll do it that way here. It is a two step process:
#
# First we need to read the config file in a python dict. There is a treecorr utility function to do this called `treecorr.read_config`. In this case, since the config file is a yaml file, it is equivalent to doing `yaml.load(open(config_file))`.
config = treecorr.read_config(config_file)
print(pprint.pformat(config))
# And then the treecorr function `corr2` takes this dict as a parameter
treecorr.corr2(config)
# The output file has a number of columns:
#
# - R_nom is the nominal center of each bin (exp(logr) where logr is spaced uniformly)
# - meanR is the actual measured mean of the pairs that went into that bin
# - meanlogR is the measured mean log(R) of the pairs
# - xip is the computed xi+(R)
# - xim is the computed xi-(R)
# - xip_im, xim_im are the imaginary components of these, since they are technically complex valued
# - sigma_xi is an estimated error bar for both xip and xim from the shape noise
# - weight is the total weight of the pairs: sum(w1 w2) if the catalog entries were given weights.
# - npairs is the total number of pairs in each bin.
#
# **IMPORTANT**:
# The sigma_xi estimate only includes shot noise. It does not (and cannot) include sample variance,
# which is almost always an important component of the total uncertainty of these
# values. Furthermore, the measured xi values are correlated, so the covariance
# matrix is not diagonal.
# Therefore, in general, you would want to use something like jackknife to get a better
# estimate of the covariance matrix of these points. sigma_xi is just a very rough
# (under-)estimate to help guide the eye in plots.
with open('output/Aardvark.out') as f:
print(''.join(f.readlines()[:10])) # Just the first 10 lines for brevity
# ---
# ### Reading in an input Catalog
# Now let's repeat that process within Python.
# The first step is to read in the input data into a treecorr.Catalog object.
#
# For reference, this corresponds to these lines in the config file:
# ```
# file_name: data/Aardvark.fit
#
# ra_col: RA
# dec_col: DEC
# ra_units: degrees
# dec_units: degrees
#
# g1_col: GAMMA1
# g2_col: GAMMA2
# k_col: KAPPA
# ```
cat = treecorr.Catalog(file_name, ra_col='RA', dec_col='DEC', ra_units='deg', dec_units='deg', g1_col='GAMMA1',
g2_col='GAMMA2', k_col='KAPPA')
# ### Computing a shear-shear correlation function
# Next we need to define what kind of correlation function we want to build including how we want the binning to work, which in the config file corresponds to these lines:
# ```
# min_sep: 1
# max_sep: 400
# nbins: 100
# sep_units: arcmin
# ```
gg = treecorr.GGCorrelation(min_sep=1, max_sep=400, nbins=100, sep_units='arcmin')
# I recommend usually using `bin size <= 0.1`. If you use fatter bins than this, then internally
# TreeCorr will set [bin_slop](http://rmjarvis.github.io/TreeCorr/html/correlation2.html#treecorr.BinnedCorr2)) such that `bin_slop * bin_size = 0.1` to avoid various numerical problems that show up with fat bins.
#
# In this case, `bin_size` is inferred from `min_sep`, `max_sep`, `nbins`, and comes out to ~0.06.
print('bin_size = %.6f'%gg.bin_size)
# Now the actual calculation is done with the GGCorrelation.process method.
#
# As a rule of thumb, this takes around 1 minute per million objects (for typical kinds of binning choices).
#
# Using smaller bins or bin_slop < 1 will be slower. Fatter bins will be faster. Also, count-count correlations (which we'll get to later) are faster, since there is less work to do for each pair.
t1 = time.time()
gg.process(cat) # Takes approx 1 minute / million objects
t2 = time.time()
print('Time for calculating gg correlation = ',t2-t1)
# Rather than write to disk, we can now access the correlation function directly. In most
# cases, the correlation function is called xi. For gg, there are two: xi+ and xi-, which
# are named xip and xim in TreeCorr.
# These are numpy arrays that you could plot or send on to some likelihood calculation, etc.
#
# Note: The error bars here are significantly underestimated, since they only include
# shape noise, not sample variance, which is a significant contributor to the total
# variance. The sample variance (and indeed the full covariance matrix) is typically
# estimated either from the data via jackknife or similar technique, or fromtheory/simulations.
#
# +
r = numpy.exp(gg.meanlogr)
xip = gg.xip
xim = gg.xim
sig = numpy.sqrt(gg.varxip)
plt.plot(r, xip, color='blue')
plt.plot(r, -xip, color='blue', ls=':')
plt.errorbar(r[xip>0], xip[xip>0], yerr=sig[xip>0], color='blue', lw=0.1, ls='')
plt.errorbar(r[xip<0], -xip[xip<0], yerr=sig[xip<0], color='blue', lw=0.1, ls='')
lp = plt.errorbar(-r, xip, yerr=sig, color='blue')
plt.plot(r, xim, color='green')
plt.plot(r, -xim, color='green', ls=':')
plt.errorbar(r[xim>0], xim[xim>0], yerr=sig[xim>0], color='green', lw=0.1, ls='')
plt.errorbar(r[xim<0], -xim[xim<0], yerr=sig[xim<0], color='green', lw=0.1, ls='')
lm = plt.errorbar(-r, xim, yerr=sig, color='green')
plt.xscale('log')
plt.yscale('log', nonposy='clip')
plt.xlabel(r'$\theta$ (arcmin)')
plt.legend([lp, lm], [r'$\xi_+(\theta)$', r'$\xi_-(\theta)$'])
plt.xlim( [1,200] )
plt.ylabel(r'$\xi_{+,-}$')
plt.show()
# -
# ### Computing a count-count correlation function
# Next let's calculate the regular count-count correlation function.
#
# The letter TreeCorr uses for galaxy counts is N, so the count-count correlation function
# is called NNCorrelation in TreeCorr.
# In this case, we will also need a random catalog, which we'll get to below, but for now we
# can calculate the data auto-correlation, which is traditionally called dd for data-data.
dd = treecorr.NNCorrelation(min_sep=0.01, max_sep=10, bin_size=0.2, sep_units='degrees')
dd.process(cat)
# Now we have to make a random catalog. For real data (not a simulation), this would typically involve a sophisticated treatment of the mask, the variation in the survey depth, etc. It's normally a rather complicated calculation. But for simulated data, as we have here, we can just randomly place points within the ra, dec range in the catalog.
#
# Note: for points to be uniformly distributed on the sky (which is what the random catalog should be), the positions should have uniform distributions in RA and sin(Dec). Not uniform in Dec.
# +
ra_min = numpy.min(cat.ra)
ra_max = numpy.max(cat.ra)
dec_min = numpy.min(cat.dec)
dec_max = numpy.max(cat.dec)
print('ra range = %f .. %f' % (ra_min, ra_max))
print('dec range = %f .. %f' % (dec_min, dec_max))
rand_ra = numpy.random.uniform(ra_min, ra_max, 10**6)
rand_sindec = numpy.random.uniform(numpy.sin(dec_min), numpy.sin(dec_max), 10**6)
rand_dec = numpy.arcsin(rand_sindec)
# -
# In this case, our catalog has a weird shape, which is moderately well approximated by the following mask. Close enough for the purpose of this demo, but for real science work, you should be more careful than this, since it doesn't get the curved edge exactly right.
#
# +
rand_ra_pi = rand_ra / numpy.pi
rand_cosdec = numpy.cos(rand_dec)
mask = ( (rand_cosdec < 0.1*(1 + 2*rand_ra_pi + 8*(rand_ra_pi)**2)) &
(rand_cosdec < 0.1*(1 + 2*(0.5-rand_ra_pi) + 8*(0.5-rand_ra_pi)**2)) )
rand_ra = rand_ra[mask]
rand_dec = rand_dec[mask]
# Check that the randoms cover the same space as the data
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(15,5))
ax1.scatter(cat.ra * 180/numpy.pi, cat.dec * 180/numpy.pi, color='blue', s=0.1)
ax1.scatter(rand_ra * 180/numpy.pi, rand_dec * 180/numpy.pi, color='green', s=0.1)
ax1.set_xlabel('RA (degrees)')
ax1.set_ylabel('Dec (degrees)')
ax1.set_title('Randoms on top of data')
# Repeat in the opposite order
ax2.scatter(rand_ra * 180/numpy.pi, rand_dec * 180/numpy.pi, color='green', s=0.1)
ax2.scatter(cat.ra * 180/numpy.pi, cat.dec * 180/numpy.pi, color='blue', s=0.1)
ax2.set_xlabel('RA (degrees)')
ax2.set_ylabel('Dec (degrees)')
ax2.set_title('Data on top of randoms')
plt.show()
# -
# It's also fun to blow up a portion of this to see the clustering in the data.
# Note: the cos(dec) factor makes the ra values real angles on the sky (locally), so the points are not stretched out along the RA direction due to Mercator projection.
# +
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(15,5))
ax1.scatter(rand_ra * 180/numpy.pi * numpy.cos(rand_dec), rand_dec * 180/numpy.pi, color='green', s=0.1)
ax1.set_xlabel('RA * cos(Dec)')
ax1.set_ylabel('Dec')
ax1.set_xlim(3,5)
ax1.set_ylim(83,85)
ax1.set_title('Randoms')
ax2.scatter(cat.ra * 180/numpy.pi * numpy.cos(cat.dec), cat.dec * 180/numpy.pi, color='blue', s=0.1)
ax2.set_xlabel('RA * cos(Dec)')
ax2.set_ylabel('Dec')
ax2.set_xlim(3,5)
ax2.set_ylim(83,85)
ax2.set_title('Data')
plt.show()
# -
# Now we can make the Catalog for the randoms. Rather than use a file on disk, here we will
# build the Catalog from the existing numpy arrays that we just made. Also, the NNCorrelation
# doesn't need g1, g2, or kappa values, so we don't bother to give anything for them.
#
# We also need to make a second NNCorrelation object to do the same processing on the randoms that we did on the data.
rand = treecorr.Catalog(ra=rand_ra, dec=rand_dec, ra_units='radians', dec_units='radians')
rr = treecorr.NNCorrelation(min_sep=0.01, max_sep=10, bin_size=0.2, sep_units='degrees')
rr.process(rand)
# Then the simple estimate of the correlation function xi = (dd-rr)/rr is calculated via the function dd.calculateXi(rr). (We'll get to the more accurate Landy-Szalay formula below.)
#
# +
xi, varxi = dd.calculateXi(rr)
r = numpy.exp(dd.meanlogr)
sig = numpy.sqrt(varxi)
plt.plot(r, xi, color='blue')
plt.plot(r, -xi, color='blue', ls=':')
plt.errorbar(r[xi>0], xi[xi>0], yerr=sig[xi>0], color='blue', lw=0.1, ls='')
plt.errorbar(r[xi<0], -xi[xi<0], yerr=sig[xi<0], color='blue', lw=0.1, ls='')
leg = plt.errorbar(-r, xi, yerr=sig, color='blue')
plt.xscale('log')
plt.yscale('log', nonposy='clip')
plt.xlabel(r'$\theta$ (degrees)')
plt.legend([leg], [r'$w(\theta)$'], loc='lower left')
plt.xlim([0.01,10])
plt.show()
# -
# A better estimator for the correlation function is the Landy-Szalay formula (dd-2dr+rr)/rr.
# For this we need a cross-correlation between the data and the randoms.
# This is also done with the process function, but now with two arguments.
dr = treecorr.NNCorrelation(min_sep=0.01, max_sep=10, bin_size=0.2, sep_units='degrees')
dr.process(cat, rand)
# If you pass both rr and dr to calculateXi, then it will use the Landy-Szalay formula, which is more accurate than the simple formula used above. Note that both curves roughly follow a power law as expected at small scales, but the Landy-Szalay formula recovers the power law out to somewhat higher separations than the simple formula.
#
# Also, similar to what we saw with the GG correlation function, the error bars are clearly underestimated, since they only include shot noise, not sample variance.
# +
xi, varxi = dd.calculateXi(rr, dr)
sig = numpy.sqrt(varxi)
plt.plot(r, xi, color='blue')
plt.plot(r, -xi, color='blue', ls=':')
plt.errorbar(r[xi>0], xi[xi>0], yerr=sig[xi>0], color='blue', lw=0.1, ls='')
plt.errorbar(r[xi<0], -xi[xi<0], yerr=sig[xi<0], color='blue', lw=0.1, ls='')
leg = plt.errorbar(-r, xi, yerr=sig, color='blue')
plt.xscale('log')
plt.yscale('log', nonposy='clip')
plt.xlabel(r'$\theta$ (degrees)')
plt.legend([leg], [r'$w(\theta)$'], loc='lower left')
plt.xlim([0.01,10])
plt.show()
# -
# ---
# ### Links for further reading
#
# - The [2004 paper](http://adsabs.harvard.edu/abs/2004MNRAS.352..338J) that describes the basic algorithm implemented in TreeCorr (section 4.1).
#
# - An annotated [sample configuration file](https://github.com/rmjarvis/TreeCorr/blob/master/sample_config.yaml), which shows, among many other things, how to specify multiple input files, including randoms, in a configuration file.
#
# - Full documentation of all [allowed configuration parameters](https://github.com/rmjarvis/TreeCorr/wiki/Configuration-Parameters).
#
# - All the possible parameters for a [Catalog object](http://rmjarvis.github.io/TreeCorr/html/catalog.html), including things like flip_g1/flip_g2 to
# adjust shear definition conventions, how to specify weights, how to skip objects with
# particular (e.g. non-zero) flags, and more.
#
# - Documentation about the different [Metric](http://rmjarvis.github.io/TreeCorr/html/metric.html) options that are possible in TreeCorr including Rlens, which uses the perpendicular distance at the location of the first object (typically a lens) and Rperp which is approximately the perpendicular separation a the geometric mean distance to the two objects in a pair.
| tests/Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import statsmodels.formula.api as smf
import statsmodels.api as sm
import seaborn as sns
import scipy.stats as stats
import matplotlib.pyplot as plt
import datetime
from datetime import datetime, timedelta
pd.set_option('display.float_format', lambda x: '%.4f' % x)
path='proshares_analysis_data.xlsx'
# Reference: The code in this notebook is modified from the HW2_2020_solution.ipynb notebook on GitHub.
# ## 1. Summary Statistics
df_hedge_fund=pd.read_excel(path,sheet_name='hedge_fund_series').set_index('date')
df_merrill_factors=pd.read_excel(path,sheet_name='merrill_factors').set_index('date')
df_hedge_fund['SPY US Equity']=df_merrill_factors['SPY US Equity']
df_hedge_fund.head()
mean_hf=df_hedge_fund.mean()*12
std_hf=df_hedge_fund.std()*np.sqrt(12)
sharpe_ratio_hf=mean_hf/std_hf
df_stats=pd.DataFrame({'Mean':mean_hf, 'Volatility':std_hf, 'Sharpe Ratio':sharpe_ratio_hf})
df_stats
# ## 2. Summary Tail-Risk Statistics
# Reference: The calculate_MDD_with_Dates function is modified from https://www.quantstart.com/articles/Event-Driven-Backtesting-with-Python-Part-VII/
# +
portfolio_names=df_hedge_fund.columns
df_tail_risk_hf = pd.DataFrame(columns=portfolio_names)
def calculate_MDD_with_Dates(data):
# Calculate the cumulative returns curve
# and set up the High Water Mark
# Then create the drawdown and duration series
hwm = [0]
idx = data.index
drawdown = pd.Series(index = idx)
duration = pd.Series(index = idx)
# Loop over the index range
for t in range(1, len(idx)):
cur_hwm = max(hwm[t-1], data[t])
hwm.append(cur_hwm)
drawdown[t]= hwm[t] - data[t]
duration[t]= 0 if drawdown[t] == 0 else duration[t-1] + 1
max_date=drawdown[drawdown==drawdown.max()].index[0]
min_date=max_date+timedelta(days=duration.max())
return drawdown.max(),max_date,min_date,duration.max()
def calculate_VaR(data, percentile):
VaR=np.percentile(data,percentile)
return VaR
def calculate_CVaR(data,VaR):
CVaR=data[data<=VaR].mean()
return CVaR
for portfolio in portfolio_names:
df_tail_risk_hf.loc['Skewness', portfolio] = df_hedge_fund[portfolio].skew()
df_tail_risk_hf.loc['Excess Kurtosis', portfolio] = df_hedge_fund[portfolio].kurtosis()-3
VaR_5= calculate_VaR(df_hedge_fund[portfolio],5)
df_tail_risk_hf.loc['VaR(.05)', portfolio] = VaR_5
df_tail_risk_hf.loc['CVaR(.05)', portfolio] = calculate_CVaR(df_hedge_fund[portfolio],VaR_5)
df_tail_risk_hf.loc['Maximum Drawdown', portfolio] =calculate_MDD_with_Dates(df_hedge_fund[portfolio])[0]
df_tail_risk_hf.loc['MDD Max Date', portfolio]=calculate_MDD_with_Dates(df_hedge_fund[portfolio])[1]
df_tail_risk_hf.loc['MDD Min Date', portfolio]=calculate_MDD_with_Dates(df_hedge_fund[portfolio])[2]
df_tail_risk_hf.loc['MDD Duration', portfolio]=calculate_MDD_with_Dates(df_hedge_fund[portfolio])[3]
# -
df_tail_risk_hf
# +
factor='SPY US Equity'
df_tail_risk_hf.loc['Skewness', factor] = df_merrill_factors[factor].skew()
df_tail_risk_hf.loc['Excess Kurtosis', factor] = df_merrill_factors[factor].kurtosis()-3
VaR_5= calculate_VaR(df_merrill_factors[factor],5)
df_tail_risk_hf.loc['VaR(.05)', factor] = VaR_5
df_tail_risk_hf.loc['CVaR(.05)', factor] = calculate_CVaR(df_merrill_factors[factor],VaR_5)
df_tail_risk_hf.loc['Maximum Drawdown', factor] =calculate_MDD_with_Dates(df_merrill_factors[factor])[0]
df_tail_risk_hf.loc['MDD Max Date', factor]=calculate_MDD_with_Dates(df_merrill_factors[factor])[1]
df_tail_risk_hf.loc['MDD Min Date', factor]=calculate_MDD_with_Dates(df_merrill_factors[factor])[2]
df_tail_risk_hf.loc['MDD Duration', factor]=calculate_MDD_with_Dates(df_merrill_factors[factor])[3]
# -
# ## 3. Summary Regression Statistics
# +
#df_merrill_factors=pd.read_excel(path,sheet_name='merrill_factors').set_index('date')
df_hedge_fund_excess=df_hedge_fund.subtract(df_merrill_factors['USGG3M Index'],axis=0)
rhs=sm.add_constant(df_hedge_fund_excess['SPY US Equity'])
df_params=pd.DataFrame(columns=portfolio_names)
for portfolio in portfolio_names:
lhs=df_hedge_fund_excess[portfolio]
res=sm.OLS(lhs,rhs,missing='drop').fit()
df_params.loc['const', portfolio]=12*res.params['const']
df_params.loc['Market Beta', portfolio]=res.params['SPY US Equity']
df_params.loc['Treynor Ratio', portfolio]=12*df_hedge_fund_excess[portfolio].mean()/res.params['SPY US Equity']
df_params.loc['Information Ratio', portfolio]=12*res.params['const']/(res.resid.std()*np.sqrt(12))
# -
df_params.T
# ## 4. Relative Performance
df_stats
# ### (a) SPY vs. Hedge-Fund Series
# SPY has the largest mean return, volatility, and Sharpe Ratio, and also the highest Treynor Ratio and Information Ratio among all the securities in the hedge fund series.
# ### (b) HDG vs. QAI
# HDG has higher mean, higher volatility, and lower Sharpe Ratio than QAI, so it only outperforms QAI in mean return, but QAI outperforms HDG in Sharpe Ratio.
# ### (c) HDG, ML, and HFRI
# HFRI has a much higher mean return and Sharpe Ratio than the ML indexes and HDG, and HFRI also has a slightly higher volatility than the ML series and HDG. Overall HDG and the ML seriies capture the high mean, high volatility features of HFRI, and HDG and the ML series also have similar beta that are slightly lower than HFRI.
# ## 5. Correlation Matrix
# ### (a) Heatmap
corrmat=df_hedge_fund_excess.corr()
# ignore self-corr
corrmat[corrmat==1] = None
sns.heatmap(corrmat,annot=True)
# ### (b) Highest and Lowest Correlations
# +
corr_rank = corrmat.unstack().sort_values().dropna()
pair_max = corr_rank.index[-1]
pair_min = corr_rank.index[0]
print(f'MIN Correlation pair is {pair_min}')
print(f'MAX Correlation pair is {pair_max}')
# -
# ## 6. Replicate HFRI
merrill_factors=df_merrill_factors.columns
df_merrill_factors['HFRIFWI Index']=df_hedge_fund['HFRIFWI Index']
portfolio_names=['HFRIFWI Index']
rhs=sm.add_constant(df_merrill_factors[merrill_factors])
df_params=pd.DataFrame(columns=portfolio_names)
df_other=pd.DataFrame(columns=portfolio_names)
df_fitted=pd.DataFrame(columns=portfolio_names)
df_residuals=pd.DataFrame(columns=portfolio_names)
for portfolio in portfolio_names:
lhs=df_merrill_factors[portfolio]
res=sm.OLS(lhs,rhs,missing='drop').fit()
df_params[portfolio]=res.params
df_params.loc['const', portfolio]=12*res.params['const']
df_params.loc['Mean Fitted Values', portfolio]=np.mean(res.fittedvalues)
df_fitted[portfolio]=res.fittedvalues
df_other.loc['R-Squared',portfolio]=res.rsquared
df_residuals[portfolio]=res.resid
df_fitted_with_intercept=df_fitted
# ### (a) Intercept and Betas
df_params_with_intercept=df_params.T
df_params_with_intercept
# ### (b) Betas
# The betas require a relatively huge short position in USGG3M index, but the sizes are realistic to use in replicating portfolios.
# ### (c) R-squared
print('\nR-squared = {:.5f}'.format(np.array(df_other.T['R-Squared'])[0]))
# ### (d) Tracing Error
print('Tracking error = {:.5f}'.format(np.array(df_residuals.std() * np.sqrt(12))[0]))
# ## 7. Replication Out-of-sample
def oos_reg(df,factors,t):
date_range =df.index[t:]
oos_fitted = pd.Series(index=date_range, name='OOS_fit')
for date in date_range:
date_month_prior = pd.DatetimeIndex([date]).shift(periods=-1, freq='M')[0]
df_subset = df[:date_month_prior]
rhs = sm.add_constant(df_subset[factors])
lhs = df_subset['HFRIFWI Index']
res = sm.OLS(lhs, rhs, drop="missing").fit()
alpha = res.params['const']
beta = res.params.drop(index='const')
x_t = df.loc[date, factors]
predicted_next_value = alpha + x_t @ beta
oos_fitted[date] = predicted_next_value
output=(pd.DataFrame([oos_fitted, df_merrill_factors.loc[df_merrill_factors.index[61]:,'HFRIFWI Index']])).T.corr()
'''
oos_fitted[df.index[t]:].plot(figsize=(14,3))
df.loc[df.index[t]:,'HFRIFWI Index'].plot()
plt.legend()
plt.show()
None
(oos_fitted[df.index[t]:] + 1).cumprod().plot(figsize=(14,3))
(df.loc[df.index[t]:,'HFRIFWI Index'] + 1).cumprod().plot()
plt.legend()
plt.show()
None
'''
return output
oos_reg(df_merrill_factors,merrill_factors,61)
oos_reg(df_merrill_factors,merrill_factors,62)
# +
months=len(df_merrill_factors)
pd.set_option('display.max_rows', None)
df_list=[]
for m in range(61,months):
df=oos_reg(df_merrill_factors,merrill_factors,m)
df_list.append(df)
pd.concat(df_list)
# -
# While we may worry that the strong replication results were dependent on being in-sample, for t=61 and t=62, the OOS results show almost the same level of replicability - roughly 93% correlation between the replicating portfolio and the HFRI. The rolling 60-month regression result also shows high correlation for most of the time values, but the correlation decreases to 70%-80% for the last 10 time values.
# ## 8. Replication without an Intercept
df_merrill_factors=pd.read_excel(path,sheet_name='merrill_factors').set_index('date')
merrill_factors=df_merrill_factors.columns
df_merrill_factors['HFRIFWI Index']=df_hedge_fund['HFRIFWI Index']
portfolio_names=['HFRIFWI Index']
rhs=df_merrill_factors[merrill_factors]
df_params=pd.DataFrame(columns=portfolio_names)
df_other=pd.DataFrame(columns=portfolio_names)
df_fitted=pd.DataFrame(columns=portfolio_names)
df_residuals=pd.DataFrame(columns=portfolio_names)
for portfolio in portfolio_names:
lhs=df_merrill_factors[portfolio]
res=sm.OLS(lhs,rhs,missing='drop').fit()
df_params[portfolio]=res.params
df_params.loc['Mean Fitted Values', portfolio]=np.mean(res.fittedvalues)
df_fitted[portfolio]=res.fittedvalues
df_other.loc['R-Squared',portfolio]=res.rsquared
df_residuals[portfolio]=res.resid
# ### (a) (b) Regression Beta and Mean Fitted Values
df_params_no_intercept=df_params.T
df_params_no_intercept
df_compare_reg=pd.concat([df_params_with_intercept,df_params_no_intercept],sort=True)
df_compare_reg
# The replication without intercept has a lower mean fitted values and smaller beta for all US Equity series except SPY, and USGG2M has a large positive beta in the no-intercept replication, while the beta for USGG2M with intercept is a large negative number. Also, both of the regressions have mean of fitted values for HFRIFWI smaller than the mean of HFRIFWI (0.0508), and the replication with intercept has a closer value.
# ### (c) Correlations of Fitted value to the HFRI
# +
df_fitted_no_intercept=df_fitted
df_fitted_no_intercept['HFRIFWI Index'].corr(df_hedge_fund['HFRIFWI Index'])
# -
df_fitted_with_intercept['HFRIFWI Index'].corr(df_hedge_fund['HFRIFWI Index'])
# The correlation for the fitted values of the no-intercept replication is a little bit smaller than the correlation for the fitted values of the replication with intercept.
# Since replication with intercept has higher mean fitted value, which indicates higher return, Merrill and ProShares may fit their replicators with an intercept to make the hedge funds more attractive.
| solutions/hw2/HW2-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:asp]
# language: python
# name: conda-env-asp-py
# ---
# # Data analysis
# +
import glob, os
import pandas as pd
import matplotlib.pyplot as plt
from scipy.stats import sigmaclip as clip
import numpy as np
from astropy.stats import sigma_clip
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
# -
# !ls
# +
draco = pd.concat(map(pd.read_json, glob.glob('draco_runtimes*.json')))
cql = pd.concat(map(pd.read_json, glob.glob('cql_runtimes*.json')))
df = pd.concat([cql, draco])
df = df.reindex()
# -
df
# +
gb = df.groupby(['system', 'encodings', 'fields'])
results = pd.DataFrame(columns=['system', 'encodings', 'fields', 'runtime'])
for i, (name, group) in enumerate(gb):
# print(group)
# group.runtime.hist()
before = group.runtime
after = pd.Series(sigma_clip(before)).dropna()
print(name, len(before), len(after))
results = results.append(pd.DataFrame.from_dict([{
'system': name[0],
'encodings': name[1],
'fields': name[2],
'mean': np.mean(after),
'stdev': np.std(after),
'min': np.min(before),
'max': np.max(before),
'runtime': np.median(before)
}]))
# plt.show()
# -
results
results.to_json('results.json',orient='records')
| benchmark/Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install psycopg2-binary
# !pwd
import psycopg2
dir(psycopg2)
help(psycopg2.connect)
dbname = 'sjnmahbj'
user = 'sjnmahbj'
password = '<PASSWORD>'
host = 'rajje.db.elephantsql.com (rajje-01)' # port should be included by degfult
pgconn = psycopg2.connect(dbname=dbname, user = user, password=password, host=host)
# #ETL - RPG data from SQLite to Postgresql
# We';d like to get the RG dst ojt g SWQ<nte snd j setryt jt k t
#
#
# !mv 'rpg_db.sqlinte)?raw=True rpg_db.sqlite)
'
# !ls
characters[:5]
# +
# we need a new table
# -
len (characters)
sl_curs = sl.conn_cursor()
# +
# we need a new table witht eh apprpropriaate schemaa
# -
sl_curs.executre("PRAGMA_table_(charactercreator_character)"").fetchall()
# +
create_character_table = """
CREATE TABLE charactercreator_character(
character_id SETISL PRIMSRY KEY,
name VRCHAR(30)
level INT,
dexterity INT,
wisdom INT,
strength INT,
intelligence INT,
)
"""
pg_curse.execute(thing)
pg_conn.conn()
## UUID
# -
characters[0]
# +
example_instert """
INSERT INTO charactercreator_character
(name, level, hp, experience, dexterity, wisdom)
VALUES = """ + str(characters[0][1:]) + ";"
print example_insert
# +
real_instert """
INSERT INTO charactercreator_character
(name, level, hp, experience, dexterity, wisdom)
VALUES = """ + str(characters[0][1:]) + ";"
print example_insert
# +
# loops
for c in characters:
insert_character = """
INSERT INTO charactercreator_character
(name, level, hp, experience, dexterity, wisdom)
VALUES = """ + str(c[0][1:])
pg_currs.execute(insert_character)
pg_conn.commit()
# print example_insert
# +
# stretch insert all at one go
pg_curs.execute("SELECT * FROM chaaractercreator_character")
pg_curse.fetchall()
# +
# YESTERDY DIRECTY QUERY SQLITE DBV TO ANSWER QUESTIONS
# CONTINUE WIORKING ON LIVE LECTURE TASK
# TITANIC!
# SCHEMA
# PUT IT INTO A PG TABLEW
# office hours
# Joins
# -
| module2-sql-for-analysis/jud-Unit3-Playground.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
import pandas as pd
import matplotlib.pyplot as plt
import commons
import textwrap
import math
# +
# load data and ratings about open-source projects
features = set()
scores = set()
data = []
score_weights = {}
def process_used_score_value(used_value, row, score_weights):
score_type = used_value['score']['type']
if used_value['isNotApplicable']:
row[score_type] = 'N/A'
else:
row[score_type] = used_value['value']
if score_type not in score_weights:
score_weights[score_type] = used_value['weight']
elif score_weights[score_type] != used_value['weight']:
raise Exception('Oops! Weights do not match!')
def process_vulnerability_value(used_value, row):
row['# of vulnerabilities'] = len(used_value['vulnerabilities']['entries'])
row['# of unpatched vulnerabilities'] = 0
row['# of patched vulnerabilities'] = 0
row['# of vulnerabilities without introduced date'] = 0
row['# of vulnerabilities without fixed date'] = 0
for entry in used_value['vulnerabilities']['entries']:
if entry['resolution'] == 'UNPATCHED':
row['# of unpatched vulnerabilities'] = row['# of unpatched vulnerabilities'] + 1
elif entry['resolution'] == 'PATCHED':
row['# of patched vulnerabilities'] = row['# of patched vulnerabilities'] + 1
if entry['introduced'] is None:
row['# of vulnerabilities without introduced date'] = row['# of vulnerabilities without introduced date'] + 1
if entry['fixed'] is None:
row['# of vulnerabilities without fixed date'] = row['# of vulnerabilities without fixed date'] + 1
def process_sub_scores(score_value, row, score_weights):
for used_value in score_value['usedValues']:
if used_value['type'] == 'ScoreValue':
scores.add(used_value['score']['type'])
process_used_score_value(used_value, row, score_weights)
process_sub_scores(used_value, row, score_weights)
continue
feature = used_value['feature']['name']
features.add(feature)
if used_value['type'] == 'UnknownValue':
row[feature] = 'unknown'
elif used_value['type'] == 'VulnerabilitiesValue':
process_vulnerability_value(used_value, row)
elif used_value['type'] == 'IntegerValue':
row[feature] = used_value['number']
elif used_value['type'] == 'BooleanValue':
row[feature] = used_value['flag']
elif used_value['type'] == 'DateValue':
row[feature] = used_value['date']
elif used_value['type'] == 'LanguagesValue':
row[feature] = used_value['languages']['elements']
elif used_value['type'] == 'PackageManagersValue':
row[feature] = used_value['packageManagers']['packageManagers']
elif used_value['type'] == 'LgtmGradeValue':
row[feature] = used_value['value']
else:
raise Exception('Unknown value type: ' + used_value['type'])
with open('../../../../../docs/oss/security/github_projects.json') as json_file:
json_data = json.load(json_file)
for json_project_data in json_data:
row = {}
row['URL'] = json_project_data['url']
rating_value = json_project_data['ratingValue']
if rating_value is None:
continue
row['Label'] = rating_value['label'][1]
score_value = json_project_data['ratingValue']['scoreValue']
row['Score'] = score_value['value']
row['Confidence'] = score_value['confidence']
process_sub_scores(score_value, row, score_weights)
data.append(row)
project_data = pd.DataFrame(data)
project_data.fillna('unknown', inplace = True)
project_data
# -
df = project_data['Label'].value_counts()
plt.bar(range(len(df)), df.values, align='center')
plt.xticks(range(len(df)), df.index.values, size='small')
plt.title('Distribution of labels')
plt.show()
df = project_data
plt.figure(figsize=[10, 10])
f, ax = plt.subplots(1, 1)
f.set_figwidth(10)
f.set_figheight(5)
column = 'Score'
d = df[df[column] != 'N/A']
d = d.loc[:, column]
ax.hist(d, histtype='bar', bins=100)
ax.set_title(column)
plt.tight_layout()
df = project_data
plt.figure(figsize=[10, 10])
f, ax = plt.subplots(1, 1)
f.set_figwidth(10)
f.set_figheight(5)
column = 'Confidence'
d = df[df[column] != 'N/A']
d = d.loc[:, column]
ax.hist(d, histtype='bar', bins=10)
ax.set_title(column)
plt.tight_layout()
df = project_data[['ProjectActivityScore', 'ProjectPopularityScore',
'CommunityCommitmentScore', 'ProjectSecurityAwarenessScore']]
plt.figure(figsize=[10, 10])
f, a = plt.subplots(2, 2)
f.set_figwidth(10)
f.set_figheight(10)
a = a.ravel()
for idx, ax in enumerate(a):
column = df.columns[idx]
d = df[df[column] != 'N/A']
d = d.iloc[:,idx]
ax.hist(d, histtype='bar', bins='auto')
ax.set_title('{} ({:f})'.format(column, score_weights[column]))
plt.tight_layout()
df = project_data[['DependencyScanScore', 'FuzzingScore',
'MemorySafetyTestingScore', 'NoHttpToolScore']]
plt.figure(figsize=[10, 10])
f, a = plt.subplots(2, 2)
f.set_figwidth(15)
f.set_figheight(10)
a = a.ravel()
for idx, ax in enumerate(a):
column = df.columns[idx]
d = df.replace('N/A', -10)
d = d.iloc[:,idx]
ax.hist(d, histtype='bar', bins='auto')
ax.set_title('{} ({:f})'.format(column, score_weights[column]))
plt.tight_layout()
df = project_data[['FindSecBugsScore', 'LgtmScore', 'StaticAnalysisScore',
'UnpatchedVulnerabilitiesScore']]
plt.figure(figsize=[10, 10])
f, a = plt.subplots(2, 2)
f.set_figwidth(15)
f.set_figheight(10)
a = a.ravel()
for idx, ax in enumerate(a):
if idx == len(df.columns): break
column = df.columns[idx]
d = df.replace('N/A', -10)
d = d.iloc[:,idx]
ax.hist(d, histtype='bar', bins='auto')
ax.set_title('{} ({:f})'.format(column, score_weights[column]))
plt.tight_layout()
df = project_data[['ProjectSecurityTestingScore', 'VulnerabilityDiscoveryAndSecurityTestingScore']]
plt.figure(figsize=[10, 10])
f, a = plt.subplots(1, 2)
f.set_figwidth(15)
f.set_figheight(10)
a = a.ravel()
for idx, ax in enumerate(a):
if idx == len(df.columns): break
column = df.columns[idx]
d = df.replace('N/A', -10)
d = d.iloc[:,idx]
ax.hist(d, histtype='bar', bins='auto')
ax.set_title('{} ({:f})'.format(column, score_weights[column]))
plt.tight_layout()
pd.DataFrame.from_dict(score_weights, orient='index')
project_data[
['URL',
'# of vulnerabilities',
'# of unpatched vulnerabilities',
'# of patched vulnerabilities']
][project_data['# of vulnerabilities'] > 3].sort_values(by = '# of vulnerabilities', ascending=False)
project_data[
['URL',
'# of vulnerabilities',
'# of vulnerabilities without introduced date',
'# of vulnerabilities without fixed date']
][project_data['# of vulnerabilities'] > 3].sort_values(by = '# of vulnerabilities', ascending=False)
# +
def autolabel(rects):
"""Attach a text label above each bar in *rects*, displaying its height."""
for rect in rects:
height = rect.get_height()
ax.annotate('{}'.format(height),
xy=(rect.get_x() + rect.get_width() / 2, height),
xytext=(0, 3), # 3 points vertical offset
textcoords="offset points",
ha='center', va='bottom')
def draw_boolean_hists(df, columns, a):
a = a.ravel()
for idx, ax in enumerate(a):
if idx == len(df.columns): break
column = df.columns[idx]
if column not in columns: continue
vc = df[column].replace(True, 'Yes').replace(False, 'No').value_counts().to_dict()
if 'Yes' not in vc: vc['Yes'] = 0
if 'No' not in vc: vc['No'] = 0
if 'unknown' not in vc: vc['unknown'] = 0
rects = ax.bar(vc.keys(), vc.values(), width=0.5)
autolabel(rects)
ax.set_title(column)
# +
columns = [
'If an open-source project uses AddressSanitizer',
'If an open-source project uses MemorySanitizer',
'If an open-source project uses UndefinedBehaviorSanitizer',
'If an open-source project is included to OSS-Fuzz project',
]
df = project_data[columns]
plt.figure(figsize=[10, 10])
f, a = plt.subplots(2, 2)
f.set_figwidth(10)
f.set_figheight(10)
draw_boolean_hists(df, columns, a)
f.tight_layout()
# +
columns = [
'If an open-source project uses FindSecBugs',
'If a project uses LGTM checks for commits',
'If a project uses nohttp tool',
'If an open-source project is regularly scanned for vulnerable dependencies',
'If a project uses Dependabot',
'If a project uses GitHub as the main development platform',
'If a project uses OWASP Enterprise Security API (ESAPI)',
'If a project uses OWASP Java HTML Sanitizer',
'If a project uses OWASP Java Encoder'
]
df = project_data[columns]
plt.figure(figsize=[10, 10])
f, a = plt.subplots(3, 3)
f.set_figwidth(15)
f.set_figheight(10)
draw_boolean_hists(df, columns, a)
f.tight_layout()
# +
columns = [
'If an open-source project is supported by a company',
'If an open-source project belongs to Apache Foundation',
'If an open-source project belongs to Eclipse Foundation',
'If an open-source project has a security policy',
'If an open-source project has a security team',
'If a project uses signed commits',
'If a project has a bug bounty program',
]
df = project_data[columns]
plt.figure(figsize=[10, 10])
f, a = plt.subplots(3, 3)
f.set_figwidth(15)
f.set_figheight(10)
draw_boolean_hists(df, columns, a)
f.tight_layout()
# +
unclear_threshold = 8.0
bad_ratings_fraction = 0.30
moderate_ratings_fraction = 0.50
good_ratings_fraction = 0.20
if bad_ratings_fraction + moderate_ratings_fraction + good_ratings_fraction != 1:
raise Exception('Oops!')
n = len(project_data.index)
target_bad_ratings_number = bad_ratings_fraction * n
target_moderate_ratings_number = moderate_ratings_fraction * n
moderate_threshold = 0
good_threshold = 0
i = 0
for index, value in project_data['Score'].sort_values().items():
i = i + 1
if moderate_threshold == 0 and i > target_bad_ratings_number:
moderate_threshold = value
i = 0
if good_threshold == 0 and i > target_moderate_ratings_number:
good_threshold = value
break
print('New threshold for MODERATE = {:f}'.format(moderate_threshold))
print('New threshold for GOOD = {:f}'.format(good_threshold))
# +
def get_new_label(row):
s = row['Score']
c = row['Confidence']
if c < unclear_threshold: return 'UNCLEAR'
if s >= good_threshold: return 'GOOD'
if s >= moderate_threshold: return 'MODERATE'
return 'BAD'
project_data['Label'] = project_data.apply(lambda row: get_new_label(row), axis=1)
df = project_data['Label'].value_counts()
plt.bar(range(len(df)), df.values, align='center')
plt.xticks(range(len(df)), df.index.values, size='small')
plt.title('New distribution of labels')
plt.show()
# -
# save the new thresholds
data = {}
data['moderate'] = moderate_threshold
data['good'] = good_threshold
data['unclear'] = unclear_threshold
filename = '../../../../../src/main/resources/com/sap/sgs/phosphor/fosstars/model/rating/oss/OssSecurityRatingThresholds.json'
with open(filename, 'w') as file:
json.dump(data, file, sort_keys=True, indent=2)
| src/main/jupyter/oss/security/SecurityRatingAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### This notebook shows how to use Permutation Importnace using eli5 library and LGBM
#
# Only 1% of data is used in this notebook while computing Permutation Importnace
# +
import os
import sys
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
pd.options.display.max_rows = 1000
# -
sys.path.insert(0, "/opt/vssexclude/personal/kaggle/kaggle_tab_mar/src")
# %load_ext autoreload
# %autoreload 2
import munging.process_data_util as process_data
import common.com_util as common
import config.constants as constants
import modeling.train_util as model
# +
SEED = 42
EXP_DETAILS = "Permutation Importance with raw features"
TARGET = 'target'
MODEL_TYPE = "lgb"
OBJECTIVE = "binary"
BOOSTING_TYPE = "gbdt"
METRIC = "auc"
VERBOSE = 100
N_THREADS = -1
NUM_LEAVES = 31
MAX_DEPTH = -1
N_ESTIMATORS = 10000
LEARNING_RATE = 0.1
EARLY_STOPPING_ROUNDS = 100
lgb_params = {
'objective': OBJECTIVE,
'boosting_type': BOOSTING_TYPE,
'learning_rate': LEARNING_RATE,
'num_leaves': NUM_LEAVES,
'tree_learner': 'serial',
'n_jobs': N_THREADS,
'seed': SEED,
'max_depth': MAX_DEPTH,
'max_bin': 255,
'metric': METRIC,
'verbose': -1,
'n_estimators': N_ESTIMATORS
}
LOGGER_NAME = 'main'
logger = common.get_logger(LOGGER_NAME)
common.set_seed(SEED)
train_df, test_df, sample_submission_df = process_data.read_processed_data(
logger, constants.PROCESSED_DATA_DIR, train=True, test=True, sample_submission=True, frac=0.1)
combined_df = pd.concat([train_df.drop('target', axis=1), test_df])
target = train_df[TARGET]
cat_fetaures = [name for name in train_df.columns if "cat" in name]
logger.info("Label Encoding the categorcal features")
for name in cat_fetaures:
lb = LabelEncoder()
combined_df[name] = lb.fit_transform(combined_df[name])
train_df = combined_df.loc[train_df.index]
train_df[TARGET] = target
test_df = combined_df.loc[test_df.index]
train_X = train_df.drop([TARGET], axis=1)
train_Y = train_df[TARGET]
test_X = test_df
logger.info(f"Shape of train_X: {train_X.shape}, train_Y: {train_Y.shape}, test_X: {test_X.shape}")
predictors = list(train_X.columns)
logger.info(f"List of predictors {predictors}")
sk = StratifiedKFold(n_splits=5, shuffle=False)
permu_imp_df, top_imp_df = model.lgb_train_perm_importance_on_cv(logger,
run_id=123,
train_X=train_X,
train_Y=train_Y,
kf=sk,
features=predictors,
params=lgb_params,
early_stopping_rounds=EARLY_STOPPING_ROUNDS,
cat_features=cat_fetaures,
display_imp=True,
verbose_eval=100,
)
# -
permu_imp_df.head()
top_imp_df.head()
plt.figure(figsize=(16, 12))
sns.barplot(
x="weight", y="feature", data=top_imp_df.sort_values(
by="weight", ascending=False))
plt.show()
| notebooks/feature_selection/permutation_importnace.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from pandas import Series,DataFrame
df1 = DataFrame(np.arange(8).reshape(2,4),
index=pd.Index(['LA','SF'],name='city'),
columns=pd.Index(['A','B','C','D'],name='letters'))
df1
# +
df_st = df1.stack()
df_st
# -
df_st.unstack()
df_st.unstack('letters')
df_st.unstack('city')
# how to stack/unstack handle null values
ser1 = Series([0,1,2],index=['Q','X','Y'])
ser2 = Series([4,5,6],index=['X','Y','Z'])
ser1
ser2
# +
df = pd.concat([ser1,ser2], keys=['Alpha','Bravo'])
df
# -
df.unstack()
# remove null values while stacking
df.unstack().stack()
# keep null values while stacking
df.unstack().stack(dropna=False)
| python/udemy-data-analysis-and-visualization/lecture33_data_reshaping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import tensorflow as tf
X1 = tf.constant(np.random.uniform(size=(1000, 3)))
X2 = tf.constant(np.random.uniform(size=(1000, 4)))
X3 = tf.concat([X1, X2], axis=1)
print(X3.shape)
zero_vals = tf.constant(np.zeros((1000, 7)))
one_vals = tf.constant(np.ones((1000, 7)))
X5 = tf.where(X3 > 0.2, one_vals, zero_vals)
X6 = tf.where(tf.logical_or(X3 < 0.2, X3 > 0.8), one_vals, zero_vals)
v1 = tf.reduce_mean(X5)
v2 = tf.reduce_mean(X6)
with tf.Session() as sess:
out1, out2 = sess.run([v1, v2])
print(out1, out2)
| old/t03_controls_logic_operators.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # 预测波士顿房价
#
# ## 在 SageMaker 中使用 XGBoost(批转换)
#
# _机器学习工程师纳米学位课程 | 开发_
#
# ---
#
# 为了介绍 SageMaker 的高阶 Python API,我们将查看一个相对简单的问题。我们将使用[波士顿房价数据集](https://www.cs.toronto.edu/~delve/data/boston/bostonDetail.html)预测波士顿地区的房价中位数。
#
# 高阶 API 的文档位于 [ReadTheDocs 页面](http://sagemaker.readthedocs.io/en/latest/)
#
# ## 一般步骤
#
# 通常,在 notebook 实例中使用 SageMaker 时,你需要完成以下步骤。当然,并非每个项目都要完成每一步。此外,有很多步骤有很大的变化余地,你将在这些课程中发现这一点。
#
# 1. 下载或检索数据。
# 2. 处理/准备数据。
# 3. 将处理的数据上传到 S3。
# 4. 训练所选的模型。
# 5. 测试训练的模型(通常使用批转换作业)。
# 6. 部署训练的模型。
# 7. 使用部署的模型。
#
# 在此 notebook 中,我们将仅介绍第 1-5 步,因为只是大致了解如何使用 SageMaker。在后面的 notebook 中,我们将详细介绍如何部署训练的模型。
# ## 第 0 步:设置 notebook
#
# 先进行必要的设置以运行 notebook。首先,加载所需的所有 Python 模块。
# +
# %matplotlib inline
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.datasets import load_boston
import sklearn.model_selection
# -
# 除了上面的模块之外,我们还需要导入将使用的各种 SageMaker 模块。
# +
import sagemaker
from sagemaker import get_execution_role
from sagemaker.amazon.amazon_estimator import get_image_uri
from sagemaker.predictor import csv_serializer
# This is an object that represents the SageMaker session that we are currently operating in. This
# object contains some useful information that we will need to access later such as our region.
session = sagemaker.Session()
# This is an object that represents the IAM role that we are currently assigned. When we construct
# and launch the training job later we will need to tell it what IAM role it should have. Since our
# use case is relatively simple we will simply assign the training job the role we currently have.
role = get_execution_role()
# -
# ## 第 1 步:下载数据
#
# 幸运的是,我们可以使用 sklearn 检索数据集,所以这一步相对比较简单。
boston = load_boston()
# ## 第 2 步:准备和拆分数据
#
# 因为使用的是整洁的表格数据,所以不需要进行任何处理。但是,我们需要将数据集中的各行拆分成训练集、测试集和验证集。
# +
# First we package up the input data and the target variable (the median value) as pandas dataframes. This
# will make saving the data to a file a little easier later on.
X_bos_pd = pd.DataFrame(boston.data, columns=boston.feature_names)
Y_bos_pd = pd.DataFrame(boston.target)
# We split the dataset into 2/3 training and 1/3 testing sets.
X_train, X_test, Y_train, Y_test = sklearn.model_selection.train_test_split(X_bos_pd, Y_bos_pd, test_size=0.33)
# Then we split the training set further into 2/3 training and 1/3 validation sets.
X_train, X_val, Y_train, Y_val = sklearn.model_selection.train_test_split(X_train, Y_train, test_size=0.33)
# -
# ## 第 3 步:将数据文件上传到 S3
#
# 使用 SageMaker 创建训练作业后,进行训练操作的容器会执行。此容器可以访问存储在 S3 上的数据。所以我们需要将用来训练的数据上传到 S3。此外,在执行批转换作业时,SageMaker 要求输入数据存储在 S3 上。我们可以使用 SageMaker API 完成这一步,它会在后台自动处理一些步骤。
#
# ### 将数据保存到本地
#
# 首先,我们需要创建测试、训练和验证 csv 文件,并将这些文件上传到 S3。
# This is our local data directory. We need to make sure that it exists.
data_dir = '../data/boston'
if not os.path.exists(data_dir):
os.makedirs(data_dir)
# +
# We use pandas to save our test, train and validation data to csv files. Note that we make sure not to include header
# information or an index as this is required by the built in algorithms provided by Amazon. Also, for the train and
# validation data, it is assumed that the first entry in each row is the target variable.
X_test.to_csv(os.path.join(data_dir, 'test.csv'), header=False, index=False)
pd.concat([Y_val, X_val], axis=1).to_csv(os.path.join(data_dir, 'validation.csv'), header=False, index=False)
pd.concat([Y_train, X_train], axis=1).to_csv(os.path.join(data_dir, 'train.csv'), header=False, index=False)
# -
# ### 上传到 S3
#
# 因为目前正在 SageMaker 会话中运行,所以可以使用代表此会话的对象将数据上传到默认的 S3 存储桶中。注意,建议提供自定义 prefix(即 S3 文件夹),以防意外地破坏了其他 notebook 或项目上传的数据。
# +
prefix = 'boston-xgboost-HL'
test_location = session.upload_data(os.path.join(data_dir, 'test.csv'), key_prefix=prefix)
val_location = session.upload_data(os.path.join(data_dir, 'validation.csv'), key_prefix=prefix)
train_location = session.upload_data(os.path.join(data_dir, 'train.csv'), key_prefix=prefix)
# -
# ## 第 4 步:训练 XGBoost 模型
#
# 将训练和验证数据上传到 S3 后,我们可以构建 XGBoost 模型并训练它。我们将使用高阶 SageMaker API 完成这一步,这样的话代码更容易读懂,但是灵活性较差。
#
# 为了构建一个评估器(即我们要训练的对象),我们需要提供训练代码所在的容器的位置。因为我们使用的是内置算法,所以这个容器由 Amazon 提供。但是,容器的完整名称比较长,取决于我们运行所在的区域。幸运的是,SageMaker 提供了一个实用方法,叫做 `get_image_uri`,它可以为我们构建镜像名称。
#
# 为了使用 `get_image_uri` 方法,我们需要向其提供当前所在区域(可以从 session 对象中获得),以及要使用的算法的名称。在此 notebook 中,我们将使用 XGBoost,但是你也可以尝试其他算法。[常见参数](https://docs.aws.amazon.com/sagemaker/latest/dg/sagemaker-algo-docker-registry-paths.html)中列出了 Amazon 的内置算法。
# +
# As stated above, we use this utility method to construct the image name for the training container.
container = get_image_uri(session.boto_region_name, 'xgboost')
# Now that we know which container to use, we can construct the estimator object.
xgb = sagemaker.estimator.Estimator(container, # The image name of the training container
role, # The IAM role to use (our current role in this case)
train_instance_count=1, # The number of instances to use for training
train_instance_type='ml.m4.xlarge', # The type of instance to use for training
output_path='s3://{}/{}/output'.format(session.default_bucket(), prefix),
# Where to save the output (the model artifacts)
sagemaker_session=session) # The current SageMaker session
# -
container
# 在要求 SageMaker 开始训练作业之前,我们需要设置模型超参数。如果使用 XGBoost 算法,可以设置的超参数有很多,以下只是其中几个。如果你想修改下面的超参数或修改其他超参数,请参阅 [XGBoost 超参数页面](https://docs.aws.amazon.com/sagemaker/latest/dg/xgboost_hyperparameters.html)
xgb.set_hyperparameters(max_depth=5,
eta=0.2,
gamma=4,
min_child_weight=6,
subsample=0.8,
objective='reg:linear',
early_stopping_rounds=10,
num_round=200)
# 完全设置好 estimator 对象后,可以训练它了。我们需要告诉 SageMaker 输入数据是 csv 格式,然后调用 `fit` 方法。
# +
# This is a wrapper around the location of our train and validation data, to make sure that SageMaker
# knows our data is in csv format.
s3_input_train = sagemaker.s3_input(s3_data=train_location, content_type='csv')
s3_input_validation = sagemaker.s3_input(s3_data=val_location, content_type='csv')
xgb.fit({'train': s3_input_train, 'validation': s3_input_validation})
# -
# ## 第 5 步:测试模型
#
# 将模型拟合训练数据并使用验证数据避免过拟合后,我们可以测试模型了。我们将使用 SageMaker 的批转换功能。首先,我们需要根据拟合的模型构建一个 transformer 对象。
xgb_transformer = xgb.transformer(instance_count = 1, instance_type = 'ml.m4.xlarge')
# 接着,要求 SageMaker 使用训练过的模型开始批转换作业,并将其应用到之前存储到 S3 上的测试数据上。我们需要指定输入数据的类型,即 `text/csv`,使 SageMaker 知道如何将数据拆分成一份份,以防整个数据集太大了,无法一次性发送给模型。
#
# 注意,SageMaker 将在后台执行批转换作业。因为我们需要等待此作业的结果,然后才能继续,所以将使用 `wait()` 方法。使用该方法还有一个好处,即可以从批转换作业中获得一些输出结果,看看是否出现任何问题。
xgb_transformer.transform(test_location, content_type='text/csv', split_type='Line')
xgb_transformer.wait()
# 现在批转换作业已经运行完毕,输出结果存储到了 S3 上。因为我们想要在 notebook 中分析输出结果,所以将使用一个 notebook 功能将输出文件从 S3 复制到本地。
# !aws s3 cp --recursive $xgb_transformer.output_path $data_dir
# 为了查看模型的运行效果,我们可以绘制一个简单的预测值与真实值散点图。如果模型的预测完全准确的话,那么散点图将是一条直线 $x=y$。可以看出,我们的模型表现不错,但是还有改进的余地。
Y_pred = pd.read_csv(os.path.join(data_dir, 'test.csv.out'), header=None)
plt.scatter(Y_test, Y_pred)
plt.xlabel("Median Price")
plt.ylabel("Predicted Price")
plt.title("Median Price vs Predicted Price")
# ## 可选步骤:清理数据
#
# SageMaker 上的默认 notebook 实例没有太多的可用磁盘空间。当你继续完成和执行 notebook 时,最终会耗尽磁盘空间,导致难以诊断的错误。完全使用完 notebook 后,建议删除创建的文件。你可以从终端或 notebook hub 删除文件。以下单元格中包含了从 notebook 内清理文件的命令。
# +
# First we will remove all of the files contained in the data_dir directory
# !rm $data_dir/*
# And then we delete the directory itself
# !rmdir $data_dir
# -
| Tutorials/Boston Housing - XGBoost (Batch Transform) - High Level.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
# Loading Datasets
# We'll be using the Kagle Disease UCI dataset as an example.You can find it here:
# https://www.kaggle.com/ronitf/heart-disease-uci
#
# +
import numpy as np
import pandas as pd
import pickle
filename ="heart.csv"
# -
# # pandas's read_csv
# Handles the most edge cases,datetime and file issues best.
df=pd.read_csv(filename)
df
# # nmpy's loadtxt and genfromtxt
# Designed for loading in data saved using np.savetext, not meant to be a robust loader.
data= np.loadtxt(filename,delimiter=",",skiprows=1)
print (data)
data=np.genfromtxt(filename, delimiter=",", dtype=None, names=True, encoding="utf-8-sig")
print (data)
print (data.dtype)
# # Manual Loading
#
# For completly weird file structures
def load_file(filename):
with open (filename,encoding="utf-8-sig") as f:
data,cols=[],[]
for i,line in enumerate(f.read().splitlines()):
if i ==0:
cols+=line.split(",")
else:
data.append([float(x) for x in line.split(",")])
df=pd.DataFrame(data,columns=cols)
return df
load_file(filename).head()
# # Pickles !
# Some danger using pickles as encoding changes.use an industry standard like hd5 instead if you can.Note IF you're working with dataframes , dont use python's pickle, pandas has their own implementation -df.to_pickle and df.read_pickle.
# Underlying algorhythm is the same, but less code for you to type and supports compression.
# +
# df= pd.read_pickle("heart.pkl")
# df.head()
| Intro loading/Loading.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
import scipy.stats as ss
import numpy as np
import matplotlib.pyplot as plt
from src.classes import *
import time
from IPython.display import display, clear_output
import seaborn as sns
sns.set(style="whitegrid", palette="deep", color_codes=True)
# Colors
Scol = "#07D5E6"
Ecol = "#FFD700"
Icol = "#FB9214"
Rcol = "#3B7548"
Dcol = "#EA1313"
Ccol = "#123456"
# -
# ### Parameter investigation
# +
fig = plt.subplots(nrows=2, ncols=2,figsize = (14,8))
plt.sca(fig[0].axes[0])
# Incubation
rv = ss.beta(a=5,b=15,loc=2,scale=19)
x = np.linspace(0,21,10000)
plt.plot(x,rv.pdf(x))
plt.title("Incubation process", fontsize = 16)
print(rv.cdf(4))
print(rv.cdf(10))
# Death process
plt.sca(fig[0].axes[1])
rv = ss.beta(a=5,b=10,loc=6,scale=10)
x = np.linspace(0,16,10000)
plt.title("Death process", fontsize = 16)
plt.plot(x,rv.pdf(x))
# Recovery
plt.sca(fig[0].axes[2])
rv = ss.beta(a=4,b=4,loc=7,scale=21)
x = np.linspace(0,28,10000)
plt.title("Recovery process", fontsize = 16)
plt.plot(x,rv.pdf(x))
# Mutation
plt.sca(fig[0].axes[3])
rv = ss.erlang(3500)
x = np.linspace(3300,3700,10000)
plt.title("Mutation process", fontsize = 16)
plt.plot(x,rv.pdf(x))
plt.tight_layout()
plt.savefig('../reports/figures/Ebola_processes.pdf', format='pdf')
plt.show()
# -
# ### Actual model
# +
# Current time and set seed
t = time.time()
np.random.seed(1)
# Parameters
beta = 0.14
new_beta = 0.055
beta_change = 150
population = 24_500_000
init_exposed = 4
prob_dead = 11314/28637 # =0.395
# Processeces
incubation_process = ss.beta(a=5,b=15,loc=2,scale=19)
recovery_process = ss.beta(a=4,b=4,loc=7,scale=21)
death_process = ss.beta(a=5,b=10,loc=6,scale=10)
mutation_process = ss.erlang(3500)
es = Ebola_SEIRSD(
beta,
incubation_process,
recovery_process,
death_process,
mutation_process,
population=population,
init_exposed=init_exposed,
prob_dead=prob_dead,
beta_change = beta_change,
new_beta=new_beta
)
history = [(day, S, E, I, R, D, C) for __, day, (S, E, I, R, D, C) in es.run_until(1500)]
day, S, E, I, R, D, C = zip(*history)
print("Model done! Took {} seconds".format(time.time()-t))
# Plot
fig = plt.figure(figsize = (14, 8))
plt.plot(day, S, label = "Susceptible", color = Scol)
plt.plot(day, E, label = "Exposed", color = Ecol)
plt.plot(day, I, label = "Infected", color = Icol)
plt.plot(day, R, label = "Recovered", color = Rcol)
plt.plot(day, D, label = "Deceased", color = Dcol)
plt.axvline(x=beta_change, c = "k", ls = "--", label = "Actions taken")
plt.legend(fontsize = 14)
plt.title("Model of Ebola using SR_SEIRSD", fontsize = 16)
plt.xlabel("Days", fontsize = 14)
plt.ylabel("Individuals", fontsize = 14)
plt.show()
# Log plot
fig = plt.figure(figsize = (14, 8))
plt.plot(day, S, label = "Susceptible", color = Scol)
plt.plot(day, E, label = "Exposed", color = Ecol)
plt.plot(day, I, label = "Infected", color = Icol)
plt.plot(day, R, label = "Recovered", color = Rcol)
plt.plot(day, D, label = "Deceased", color = Dcol)
plt.axvline(x=beta_change, c = "k", ls = "--", label = "Actions taken")
plt.yscale('log')
plt.legend(fontsize = 14)
plt.title("Model of Ebola using SR_SEIRSD (log-scale)", fontsize = 16)
plt.xlabel("Days", fontsize = 14)
plt.ylabel("Individuals", fontsize = 14)
plt.savefig('../reports/figures/Ebola_log.pdf', format='pdf')
plt.show()
fig = plt.figure(figsize = (14, 8))
plt.plot(day, D, label = "Deceased", color = Dcol)
plt.plot(day, C, label = "Total Cases", color = Ccol)
plt.axvline(x=beta_change, c = "k", ls = "--", label = "Actions taken")
plt.legend(fontsize = 14)
plt.title("Model of Ebola using SR_SEIRSD", fontsize = 16)
plt.xlabel("Days", fontsize = 14)
plt.ylabel("Individuals", fontsize = 14)
plt.savefig('../reports/figures/Ebola_DC.pdf', format='pdf')
plt.show()
# Print time
print("ALl done! Toke {} seconds".format(time.time()-t))
print(es.state)
# -
# Notes:
#
# - Den initiale beta-værdi er sat ret lavt. Ift. vores model kan det begrundes med, hvor rural Afrika er.
#
# - Key number: https://en.wikipedia.org/wiki/West_African_Ebola_virus_epidemic_timeline_of_reported_cases_and_deaths "Thus Ebola virus disease spread for several months before it was recognized as such." Derfor cutoff ved 150 dage. Se grafer nederst for sammenligning
#
# - Processer: https://en.wikipedia.org/wiki/Ebola
#
# +
# Current time and set seed
t = time.time()
np.random.seed(42)
# Parameters
beta = 0.14
new_beta = 0.055
beta_change = 150
population = 24_500_000
init_exposed = 4
prob_dead = 11314/28637 # =0.395
# Processeces
incubation_process = ss.beta(a=5,b=15,loc=2,scale=19)
recovery_process = ss.beta(a=4,b=4,loc=7,scale=21)
death_process = ss.beta(a=5,b=10,loc=6,scale=10)
mutation_process = ss.erlang(3500)
num_sim = 1000
day = [None]*num_sim
S = [None]*num_sim
E = [None]*num_sim
I = [None]*num_sim
R = [None]*num_sim
D = [None]*num_sim
C = [None]*num_sim
for i in range(num_sim):
es = Ebola_SEIRSD(
beta,
incubation_process,
recovery_process,
death_process,
mutation_process,
population=population,
init_exposed=init_exposed,
prob_dead=prob_dead,
beta_change = beta_change,
new_beta=new_beta
)
history = [(day, S, E, I, R, D, C) for __, day, (S, E, I, R, D, C) in es.run_until(3000)]
day[i], S[i], E[i], I[i], R[i], D[i], C[i] = zip(*history)
if (i % 10) == 0:
clear_output(wait=True)
display(str(i)+' iterations complete')
print("ALl done! Toke {} seconds".format(time.time()-t))
# -
D_estimates = [d[-1] for d in D]
C_estimates = [c[-1] for c in C]
day_estimates = [d[-1] for d in day]
print(np.mean(D_estimates), np.quantile(D_estimates,[0.025,0.975]))
print(np.mean(C_estimates), np.quantile(C_estimates,[0.025,0.975]))
# +
fig = plt.subplots(nrows=1, ncols=2,figsize = (16,6), sharey=True)
plt.sca(fig[0].axes[0])
plt.hist(D_estimates, bins = 20)
plt.title("Total number of deaths based on 1.000 simulations", fontsize = 16)
plt.xlabel("Number of deaths", fontsize = 14)
plt.axvline(x=np.quantile(D_estimates,0.025), c = "k", ls = "--")
plt.axvline(x=np.quantile(D_estimates,0.975), c = "k", ls = "--")
plt.sca(fig[0].axes[1])
plt.hist(C_estimates, bins = 20)
plt.title("Total number of cases based on 1.000 simulations", fontsize = 16)
plt.xlabel("Number of cases", fontsize = 14)
plt.axvline(x=np.quantile(C_estimates,0.025), c = "k", ls = "--")
plt.axvline(x=np.quantile(C_estimates,0.975), c = "k", ls = "--")
plt.savefig('../reports/figures/Ebola_hist.pdf', format='pdf')
plt.show()
# -
fig = plt.figure(figsize = (14, 8))
plt.plot(day[766], S[766], label = "Susceptible", color = Scol)
plt.plot(day[766], E[766], label = "Exposed", color = Ecol)
plt.plot(day[766], I[766], label = "Infected", color = Icol)
plt.plot(day[766], R[766], label = "Recovered", color = Rcol)
plt.plot(day[766], D[766], label = "Deceased", color = Dcol)
plt.axvline(x=beta_change, c = "k", ls = "--", label = "Actions taken")
plt.yscale('log')
plt.legend(fontsize = 14)
plt.title("Model of Ebola using SR_SEIRSD (log-scale)", fontsize = 16)
plt.xlabel("Days", fontsize = 14)
plt.ylabel("Individuals", fontsize = 14)
plt.show()
np.argmax(day_estimates)
| notebooks/Ebola.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from bs4 import BeautifulSoup
import pandas as pd
import numpy as np
import requests as req
import scrapy
html_text = req.get('https://www.reed.co.uk/jobs/data-scientist-jobs-in-london').text
soup = BeautifulSoup(html_text, 'lxml')
jobs_count = soup.find('span', class_ = 'count').text.replace(' ','')
listings = soup.find('article', class_ = 'job-result')
company = listings.find('a', class_ = 'gtmJobListingPostedBy').text
salary = listings.find('li', class_ = "salary").text
location = listings.find('li', class_ = "location").text.replace(' ','')
print(listings)
| script/python/scrapy_project/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="H7OLbevlbd_Z"
# # Lambda School Data Science Module 143
#
# ## Introduction to Bayesian Inference
#
# !['Detector! What would the Bayesian statistician say if I asked him whether the--' [roll] 'I AM A NEUTRINO DETECTOR, NOT A LABYRINTH GUARD. SERIOUSLY, DID YOUR BRAIN FALL OUT?' [roll] '... yes.'](https://imgs.xkcd.com/comics/frequentists_vs_bayesians_2x.png)
#
# *[XKCD 1132](https://www.xkcd.com/1132/)*
#
# + [markdown] colab_type="text" id="3mz8p08BsN6p"
# ## Prepare - Bayes' Theorem and the Bayesian mindset
# + [markdown] colab_type="text" id="GhycNr-Sbeie"
# Bayes' theorem possesses a near-mythical quality - a bit of math that somehow magically evaluates a situation. But this mythicalness has more to do with its reputation and advanced applications than the actual core of it - deriving it is actually remarkably straightforward.
#
# ### The Law of Total Probability
#
# By definition, the total probability of all outcomes (events) if some variable (event space) $A$ is 1. That is:
#
# $$P(A) = \sum_n P(A_n) = 1$$
#
# The law of total probability takes this further, considering two variables ($A$ and $B$) and relating their marginal probabilities (their likelihoods considered independently, without reference to one another) and their conditional probabilities (their likelihoods considered jointly). A marginal probability is simply notated as e.g. $P(A)$, while a conditional probability is notated $P(A|B)$, which reads "probability of $A$ *given* $B$".
#
# The law of total probability states:
#
# $$P(A) = \sum_n P(A | B_n) P(B_n)$$
#
# In words - the total probability of $A$ is equal to the sum of the conditional probability of $A$ on any given event $B_n$ times the probability of that event $B_n$, and summed over all possible events in $B$.
#
# ### The Law of Conditional Probability
#
# What's the probability of something conditioned on something else? To determine this we have to go back to set theory and think about the intersection of sets:
#
# The formula for actual calculation:
#
# $$P(A|B) = \frac{P(A \cap B)}{P(B)}$$
#
# We can see how this relates back to the law of total probability - multiply both sides by $P(B)$ and you get $P(A|B)P(B) = P(A \cap B)$ - replaced back into the law of total probability we get $P(A) = \sum_n P(A \cap B_n)$.
#
# This may not seem like an improvement at first, but try to relate it back to the above picture - if you think of sets as physical objects, we're saying that the total probability of $A$ given $B$ is all the little pieces of it intersected with $B$, added together. The conditional probability is then just that again, but divided by the probability of $B$ itself happening in the first place.
#
# ### Bayes Theorem
#
# Here is is, the seemingly magic tool:
#
# $$P(A|B) = \frac{P(B|A)P(A)}{P(B)}$$
#
# In words - the probability of $A$ conditioned on $B$ is the probability of $B$ conditioned on $A$, times the probability of $A$ and divided by the probability of $B$. These unconditioned probabilities are referred to as "prior beliefs", and the conditioned probabilities as "updated."
#
# Why is this important? Scroll back up to the XKCD example - the Bayesian statistician draws a less absurd conclusion because their prior belief in the likelihood that the sun will go nova is extremely low. So, even when updated based on evidence from a detector that is $35/36 = 0.972$ accurate, the prior belief doesn't shift enough to change their overall opinion.
#
# There's many examples of Bayes' theorem - one less absurd example is to apply to [breathalyzer tests](https://www.bayestheorem.net/breathalyzer-example/). You may think that a breathalyzer test that is 100% accurate for true positives (detecting somebody who is drunk) is pretty good, but what if it also has 8% false positives (indicating somebody is drunk when they're not)? And furthermore, the rate of drunk driving (and thus our prior belief) is 1/1000.
#
# What is the likelihood somebody really is drunk if they test positive? Some may guess it's 92% - the difference between the true positives and the false positives. But we have a prior belief of the background/true rate of drunk driving. Sounds like a job for Bayes' theorem!
#
# $$
# \begin{aligned}
# P(Drunk | Positive) &= \frac{P(Positive | Drunk)P(Drunk)}{P(Positive)} \\
# &= \frac{1 \times 0.001}{0.08} \\
# &= 0.0125
# \end{aligned}
# $$
#
# In other words, the likelihood that somebody is drunk given they tested positive with a breathalyzer in this situation is only 1.25% - probably much lower than you'd guess. This is why, in practice, it's important to have a repeated test to confirm (the probability of two false positives in a row is $0.08 * 0.08 = 0.0064$, much lower), and Bayes' theorem has been relevant in court cases where proper consideration of evidence was important.
# + [markdown] colab_type="text" id="htI3DGvDsRJF"
# ## Live Lecture - Deriving Bayes' Theorem, Calculating Bayesian Confidence
# + [markdown] colab_type="text" id="moIJNQ-nbfe_"
# Notice that $P(A|B)$ appears in the above laws - in Bayesian terms, this is the belief in $A$ updated for the evidence $B$. So all we need to do is solve for this term to derive Bayes' theorem. Let's do it together!
# + colab={} colab_type="code" id="ke-5EqJI0Tsn"
# Activity 2 - Use SciPy to calculate Bayesian confidence intervals
# https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.bayes_mvs.html#scipy.stats.bayes_mvs
# -
import scipy.stats as stats
import numpy as np
# +
# Frequentist approach (from yesterday)
def confidence_interval(data, confidence=0.95):
"""
Calculate a confidence interval around a sample mean for given data.
Using t-distribution and two-tailed test, default 95% confidence.
Arguments:
data - iterable (list or numpy array) of sample observations
confidence - level of confidence for the interval
Returns:
tuple of (mean, lower bound, upper bound)
"""
data = np.array(data)
mean = np.mean(data)
n = len(data)
stderr = stats.sem(data)
interval = stderr * stats.t.ppf((1 + confidence) / 2., n - 1)
return (mean, mean - interval, mean + interval)
# -
size =coinflips = np.random.binomial(n=1, p=0.5, size =100)
confidence_interval(coinflips), stats.bayes_mvs(coinflips)
# +
treatment_group = np.random.binomial(n=1, p=0.65, size = 100)
nontreated_group = np.random.binomial(n=1, p=0.4, size =100)
import pandas as pd
df = pd.DataFrame((treatment_group, nontreated_group), index = None)
# -
df
# + [markdown] colab_type="text" id="P-DzzRk5bf0z"
# ## Assignment - Code it up!
#
# Most of the above was pure math - write Python code to reproduce the results. This is purposefully open ended - you'll have to think about how you should represent probabilities and events. You can and should look things up, and as a stretch goal - refactor your code into helpful reusable functions!
#
# If you're unsure where to start, check out [this blog post of Bayes theorem with Python](https://dataconomy.com/2015/02/introduction-to-bayes-theorem-with-python/) - you could and should create something similar!
#
# Stretch goal - apply a Bayesian technique to a problem you previously worked (in an assignment or project work) on from a frequentist (standard) perspective.
# + colab={} colab_type="code" id="xpVhZyUnbf7o"
# TODO - code!
# -
def bayes(BcA,A,B):
return (A*BcA)/B
bayes(1,.001,0.08)
#drunk driving example
#double check the breathelyze
A = 0.08*.08
A
bayes(1,0.001,0.0064)
# ### How many breathylyzer tests do you need to be more than 50% sure
def fifty_percent_sure(BcA,A,B):
count = 1
while bayes(BcA,A,B) < .5:
B = B**2
count +=1
print("you need to test "+str(count)+" times to be more than 50% sure of the results")
fifty_percent_sure(1,.001,.07992)
# (Sending this to channel as it's a good question - regarding the breathalyzer calculation from the prewatch video).
#
# Technically the 0.08 is rounded - but it's very close to the true value. Overall we can consider P(Positive) = P(Positive|Drunk)P(Drunk) + P(Positive|Not Drunk)P(Not Drunk) (this is referred to as the "total probability of P(Positive)").
#
# - P(Positive|Drunk)P(Drunk) = 1 * 0.001 = 0.001
# - P(Positive|Not Drunk)P(Not Drunk) = 0.08 * 0.999 = 0.07992
# - 0.07992 + 0.001 = 0.08092, which is approximately 0.08
def bayes_for_test():
tp = float(input("what is the true positive rate of your test? \n please use decimals i.e. 0.01\n"))
gp = float(input("what is the probability of the general public meeting this condition\nplease use decimals i.e. 0.01\n"))
fp = float(input("What is the false positive rate of the test?\n"))
s = bayes(tp,gp,tp-1+fp)
print('the probability of the condition being true, given the positive test is '+str(round(s,2))+'%.')
fifty_percent_sure(tp,gp,tp-1+fp)
bayes_for_test()
# ## bayes theorem sampling
#
import numpy as np
data_coin_flips = np.random.randint(2, size=1000)
np.mean(data_coin_flips)
bernoulli_flips = np.random.binomial(n=1, p=.5, size =1000)
np.mean(bernoulli_flips)
# +
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
sns.set(style='ticks', palette='Set2')
params = np.linspace(0, 1, 100)
p_x = [np.product(stats.bernoulli.pmf(data_coin_flips, p)) for p in params]
p_x =p_x/np.sum(p_x)
plt.plot(params, p_x)
sns.despine()
max(p_x)
# -
from IPython.display import Latex
# ### breast cancer & BRCA
# - (1/8) women will be diagonosed with breast cancer
# - 0.25% of people carry a mutuated BRCA gene
# - ~60% of BRCA1 carriers develop cancer by 70
# - 45% of BRCA2 carriers develop cancer by 70
# - for convenience, i will use 52.5% of BRCA carriers will develop cancer
#
#
# What Percent of women diagnosed with cancer have the mutation?
#
#
# \begin{align}
# P(BRCA | cancer) &= \frac {P(cancer | BRCA) \times P(BRCA)}{P(cancer)} \\
# P(BRCA) &= 0.0025 \\
# P(Cancer) &= (1/8) = .125 \\
# P(Cancer | BRCA) &= .525 \\
# P(BRCA | cancer) &= \frac {.525 \times 0.0025} {0.125} \\
# P(BRCA | cancer) &= 0.0105
# \\
# \\
# \end{align}
#
#
# https://www.nationalbreastcancer.org/what-is-brca
#
# https://www.nationalbreastcancer.org/genetic-testing-for-breast-cancer
# \\
# P(NoCancer) &= (7/8) = .875 \\
# P(No BRCA) &= .9975 \\
# P(No Cancer | No BRCA) &= P(No Cancer * No BRCA)/P(NoBRCA) \\
# P(No Cancer | No BRCA) &= \frac{P(No BRCA | No Cancer) \times P(NoCancer)}{P(NoBRCA)}\\
# P(Cancer | BRCA) &= .525 \\
# P(NoCancer | BRCA) &= .475 \\
# P(Cancer | BRCA) &= P(Cancer * BRCA)/P(BRCA)\\
# \end{align}
| module3-introduction-to-bayesian-inference/LS_DS_143_Introduction_to_Bayesian_Inference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ana_py37tf]
# language: python
# name: conda-env-ana_py37tf-py
# ---
# # 07.13 - Modeling - SARIMAX IV + XGBoost
# ## Imports & setup
# +
import pathlib
import warnings
from datetime import datetime
import sys
import pickle
import joblib
import gc
import pandas as pd
import numpy as np
# Plotting
import matplotlib as mpl
import matplotlib.pyplot as plt
from pandas.plotting import register_matplotlib_converters
register_matplotlib_converters()
from matplotlib.dates import DateFormatter
import matplotlib.dates as mdates
# Imports
sys.path.append("..")
from src.utils.utils import (AnnualTimeSeriesSplit,
RollingAnnualTimeSeriesSplit,
bound_precision,
run_cross_val,
run_data_split_cross_val,
save_run_results)
from src.features.features import CyclicalToCycle
from src.models.models import SK_SARIMAX, SK_Prophet, SetTempAsPower, SK_Prophet_1
from src.visualization.visualize import (plot_prediction,
plot_joint_plot,
residual_plots,
print_residual_stats,
resids_vs_preds_plot)
#b # Packages
from sklearn.pipeline import Pipeline
from skoot.feature_selection import FeatureFilter
from skoot.preprocessing import SelectiveRobustScaler
from sklearn.metrics import mean_absolute_error
from scipy.stats import norm
from statsmodels.graphics.gofplots import qqplot
from pandas.plotting import autocorrelation_plot
from statsmodels.graphics.tsaplots import plot_acf
import statsmodels.api as sm
from fbprophet import Prophet
from xgboost.sklearn import XGBRegressor
# Display
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
figsize=(15,7)
warnings.filterwarnings(action='ignore')
# %matplotlib inline
# Data
PROJECT_DIR = pathlib.Path.cwd().parent.resolve()
CLEAN_DATA_DIR = PROJECT_DIR / 'data' / '05-clean'
MODELS_DIR = PROJECT_DIR / 'data' / 'models'
RESULTS_PATH = PROJECT_DIR / 'data' /'results' / 'results.csv'
# -
# ## Load Daily Data & Inspect
df = pd.read_csv(CLEAN_DATA_DIR / 'clean-features.csv', parse_dates=True, index_col=0)
X = df.copy(deep=True)
X = X.loc['1994': '2008']
y = X.pop('daily_peak')
X.head()
y.tail()
# ## SARIMAX Model + XGBoost Residuals Model
# +
N_SPLITS = 10
def run_model(X,y, param=None):
scores_dicts = {'train':
{'mae': [],
'bound': []},
'test':
{'mae': [],
'bound': []}}
# Pre-processing
# Robust Scaling
robust_scaler_cols = ['hmdxx_min', 'hmdxx_max', 'hmdxx_median-1', 'temp_min','temp_max',
'dew_point_temp_max', 'visibility_mean']
# Cyclical Transform
rscaler = SelectiveRobustScaler(cols=robust_scaler_cols, trans_col_name=robust_scaler_cols)
cyclical0 = CyclicalToCycle(cycle_name='hmdxx_max_hour', periods_per_cycle=24)
cyclical1 = CyclicalToCycle(cycle_name='sun_rise', periods_per_cycle=24)
cyclical2 = CyclicalToCycle(cycle_name='sun_set', periods_per_cycle=24)
cyclical3 = CyclicalToCycle(cycle_name='day_of_week', periods_per_cycle=24)
cyclical4 = CyclicalToCycle(cycle_name='week_of_year', periods_per_cycle=24)
ts_model = SK_SARIMAX(order=(1,0,1), seasonal_order=(1,0,0,96), trend='c')
# Residuals model
resid_model = XGBRegressor(max_depth=8,
objective='reg:squarederror',
n_estimators=200, #200
learning_rate=0.01,
subsample=0.8,
colsample_bytree=1.0,
gamma=0.3,
importance_type='gain')
ratscv = RollingAnnualTimeSeriesSplit(n_splits=N_SPLITS, goback_years=5)
for train_indces, test_indces in ratscv.split(X, y):
X_train = X.iloc[train_indces]; y_train = y.iloc[train_indces]
X_test = X.iloc[test_indces] ; y_test = y.iloc[test_indces]
# Pre-processing Cyclic features
cyclical0.fit(X_train, y_train)
X_t_train = cyclical0.transform(X_train)
X_t_test = cyclical0.transform(X_test)
cyclical1.fit(X_t_train, y_train)
X_t_train = cyclical1.transform(X_t_train)
X_t_test = cyclical1.transform(X_t_test)
cyclical2.fit(X_t_train, y_train)
X_t_train = cyclical2.transform(X_t_train)
X_t_test = cyclical2.transform(X_t_test)
cyclical3.fit(X_t_train, y_train)
X_t_train = cyclical3.transform(X_t_train)
X_t_test = cyclical3.transform(X_t_test)
cyclical4.fit(X_t_train, y_train)
X_t_train = cyclical4.transform(X_t_train)
X_t_test = cyclical4.transform(X_t_test)
# Pre-processing Robust Scaler
rscaler.fit(X_t_train, y_train)
X_t_train = rscaler.transform(X_t_train)
X_t_test = rscaler.transform(X_t_test)
# LH Initial y Prediction
ts_model.fit(X_t_train, y_train)
y_hat_train = ts_model.predict(X_t_train)
y_hat_test = ts_model.predict(X_t_test)
# LH Get Residuals
r_train = y_train - y_hat_train
r_test = y_test - y_hat_test
# LH Fit & Predict Residuals
resid_model.fit(X_t_train, r_train)
r_hat_train = resid_model.predict(X_t_train)
r_hat_test = resid_model.predict(X_t_test)
# LH Adder - Add Residual Prediction to Initial y Prediction
y_hat_plus_train = y_hat_train + r_hat_train
y_hat_plus_test = y_hat_test + r_hat_test
mae_train = mean_absolute_error(y_train, y_hat_plus_train)
mae_test = mean_absolute_error(y_test, y_hat_plus_test)
bound_prec_train = bound_precision(y_train, y_hat_plus_train)
bound_prec_test = bound_precision(y_test, y_hat_plus_test)
scores_dicts['train']['mae'].append(mae_train)
scores_dicts['train']['bound'].append(bound_prec_train)
scores_dicts['test']['mae'].append(mae_test)
scores_dicts['test']['bound'].append(bound_prec_test)
# Use the time series pred vals DataFrame as a template
pred_vals = ts_model.get_pred_values()
for col in pred_vals.columns:
# Leave the is_forecast in place to tell the plot functions the train/test split
if col != 'is_forecast':
pred_vals[col] = np.NaN
# Fill in the values for y
pred_vals.loc[y_train.index, 'y'] = y_train.values
pred_vals.loc[y_test.index, 'y'] = y_test.values
# Replace the Time Series Models predictions with the
# combined prediction values
pred_vals.loc[y_hat_plus_train.index, 'yhat'] = y_hat_plus_train.values
pred_vals.loc[y_hat_plus_test.index, 'yhat'] = y_hat_plus_test.values
# Calculate the residuals
pred_vals['resid'] = pred_vals['y'].subtract(pred_vals['yhat'])
# Eliminate the obsolete columns such as confidence intervals
pred_vals = pred_vals[['y', 'yhat', 'resid', 'is_forecast']]
return pred_vals, scores_dicts
# -
pred_vals, d = run_model(X, y)
print(d)
print('Train')
print(np.mean(d['train']['mae']))
print(np.mean(d['train']['bound']))
print('Test')
print(np.mean(d['test']['mae']))
print(np.mean(d['test']['bound']))
pred_vals
# ### Review the last model run in the Cross Validation
fig, ax = plot_prediction(pred_vals, goback_years=20)
plt.show();
fig, ax = plot_prediction(pred_vals, goback_years=2)
plt.show();
fig, ax = plot_joint_plot(pred_vals, goback_years=1)
plt.show();
fig, ax = resids_vs_preds_plot(pred_vals)
plt.show()
fig, ax = residual_plots(pred_vals)
plt.show()
print_residual_stats(pred_vals)
df = save_run_results(X, N_SPLITS, 'SARIMAX IV + XGBoost', d, RESULTS_PATH)
df
| notebooks/07.13 - Modeling - SARIMAX IV + XGBoost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deep Deterministic Policy Gradient
#
# In this notebook you will teach a __pytorch__ neural network to do Deterministic Policy Gradient.
# +
# # !pip install -r ../requirements.txt
# +
import math
import random
import gym
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torch.distributions import Normal
# -
# from IPython.display import clear_output
import matplotlib.pyplot as plt
# %matplotlib inline
use_cuda = torch.cuda.is_available()
device = torch.device("cuda" if use_cuda else "cpu")
# ## Environment
# ### Normalize action space
class NormalizedActions(gym.ActionWrapper):
def action(self, action):
low_bound = self.action_space.low
upper_bound = self.action_space.high
action = low_bound + (action + 1.0) * 0.5 * (upper_bound - low_bound)
action = np.clip(action, low_bound, upper_bound)
return action
def reverse_action(self, action):
low_bound = self.action_space.low
upper_bound = self.action_space.high
action = 2 * (action - low_bound) / (upper_bound - low_bound) - 1
action = np.clip(action, low_bound, upper_bound)
return actions
# ### Exploration - GaussNoise
# Adding Normal noise to the actions taken by the deterministic policy<br>
class GaussNoise:
"""
For continuous environments only.
Adds spherical Gaussian noise to the action produced by actor.
"""
def __init__(self, sigma):
super().__init__()
self.sigma = sigma
def get_action(self, action):
noisy_action = np.random.normal(action, self.sigma)
return noisy_action
# <h1> Continuous control with deep reinforcement learning</h1>
# <h2><a href="https://arxiv.org/abs/1509.02971">Arxiv</a></h2>
# +
class ValueNetwork(nn.Module):
def __init__(
self,
num_inputs,
num_actions,
hidden_size,
init_w=3e-3
):
super().__init__()
self.net = nn.Sequential(
nn.Linear(num_inputs + num_actions, hidden_size),
nn.ReLU(),
nn.Linear(hidden_size, hidden_size),
nn.ReLU(),
)
self.head = nn.Linear(hidden_size, 1)
self.head.weight.data.uniform_(-init_w, init_w)
self.head.bias.data.uniform_(-init_w, init_w)
def forward(self, state, action):
x = torch.cat([state, action], 1)
x = self.net(x)
x = self.head(x)
return x
class PolicyNetwork(nn.Module):
def __init__(
self,
num_inputs,
num_actions,
hidden_size,
init_w=3e-3
):
super().__init__()
self.net = nn.Sequential(
nn.Linear(num_inputs, hidden_size),
nn.ReLU(),
nn.Linear(hidden_size, hidden_size),
nn.ReLU(),
)
self.head = nn.Linear(hidden_size, num_actions)
self.head.weight.data.uniform_(-init_w, init_w)
self.head.bias.data.uniform_(-init_w, init_w)
def forward(self, state):
x = state
x = self.net(x)
x = self.head(x)
return x
def get_action(self, state):
state = torch.tensor(state, dtype=torch.float32)\
.unsqueeze(0).to(device)
action = self.forward(state)
action = action.detach().cpu().numpy()[0]
return action
# -
# <h2>DDPG Update</h2>
def ddpg_update(
state,
action,
reward,
next_state,
done,
gamma = 0.99,
min_value=-np.inf,
max_value=np.inf,
soft_tau=1e-2,
):
state = torch.tensor(state, dtype=torch.float32).to(device)
next_state = torch.tensor(next_state, dtype=torch.float32).to(device)
action = torch.tensor(action, dtype=torch.float32).to(device)
reward = torch.tensor(reward, dtype=torch.float32).unsqueeze(1).to(device)
done = torch.tensor(np.float32(done)).unsqueeze(1).to(device)
policy_loss = value_net(state, policy_net(state))
policy_loss = -policy_loss.mean()
next_action = target_policy_net(next_state)
target_value = target_value_net(next_state, next_action.detach())
expected_value = reward + (1.0 - done) * gamma * target_value
expected_value = torch.clamp(expected_value, min_value, max_value)
value = value_net(state, action)
value_loss = value_criterion(value, expected_value.detach())
policy_optimizer.zero_grad()
policy_loss.backward()
policy_optimizer.step()
value_optimizer.zero_grad()
value_loss.backward()
value_optimizer.step()
for target_param, param in zip(target_value_net.parameters(), value_net.parameters()):
target_param.data.copy_(
target_param.data * (1.0 - soft_tau) + param.data * soft_tau
)
for target_param, param in zip(target_policy_net.parameters(), policy_net.parameters()):
target_param.data.copy_(
target_param.data * (1.0 - soft_tau) + param.data * soft_tau
)
# ### Experience replay buffer
#
# 
class ReplayBuffer:
def __init__(self, capacity):
self.capacity = capacity
self.buffer = []
self.position = 0
def push(self, state, action, reward, next_state, done):
if len(self.buffer) < self.capacity:
self.buffer.append(None)
self.buffer[self.position] = (state, action, reward, next_state, done)
self.position = (self.position + 1) % self.capacity
def sample(self, batch_size):
batch = random.sample(self.buffer, batch_size)
state, action, reward, next_state, done = map(np.stack, zip(*batch))
return state, action, reward, next_state, done
def __len__(self):
return len(self.buffer)
# ---
# +
batch_size = 128
def generate_session(t_max=1000, train=False):
"""play env with ddpg agent and train it at the same time"""
total_reward = 0
state = env.reset()
for t in range(t_max):
action = policy_net.get_action(state)
if train:
action = noise.get_action(action)
next_state, reward, done, _ = env.step(action)
if train:
replay_buffer.push(state, action, reward, next_state, done)
if len(replay_buffer) > batch_size:
states, actions, rewards, next_states, dones = \
replay_buffer.sample(batch_size)
ddpg_update(states, actions, rewards, next_states, dones)
total_reward += reward
state = next_state
if done:
break
return total_reward
# +
env = NormalizedActions(gym.make("Pendulum-v0"))
noise = GaussNoise(sigma=0.3)
state_dim = env.observation_space.shape[0]
action_dim = env.action_space.shape[0]
hidden_dim = 256
value_net = ValueNetwork(state_dim, action_dim, hidden_dim).to(device)
policy_net = PolicyNetwork(state_dim, action_dim, hidden_dim).to(device)
target_value_net = ValueNetwork(state_dim, action_dim, hidden_dim).to(device)
target_policy_net = PolicyNetwork(state_dim, action_dim, hidden_dim).to(device)
for target_param, param in zip(target_value_net.parameters(), value_net.parameters()):
target_param.data.copy_(param.data)
for target_param, param in zip(target_policy_net.parameters(), policy_net.parameters()):
target_param.data.copy_(param.data)
value_lr = 1e-3
policy_lr = 1e-4
value_optimizer = optim.Adam(value_net.parameters(), lr=value_lr)
policy_optimizer = optim.Adam(policy_net.parameters(), lr=policy_lr)
value_criterion = nn.MSELoss()
replay_buffer_size = 10000
replay_buffer = ReplayBuffer(replay_buffer_size)
# +
max_steps = 500
valid_mean_rewards = []
for i in range(100):
session_rewards_train = [
generate_session(t_max=max_steps, train=True)
for _ in range(10)
]
session_rewards_valid = [
generate_session(t_max=max_steps, train=False)
for _ in range(10)
]
print(
"epoch #{:02d}\tmean reward (train) = {:.3f}\tmean reward (valid) = {:.3f}".format(
i, np.mean(session_rewards_train), np.mean(session_rewards_valid))
)
valid_mean_rewards.append(np.mean(session_rewards_valid))
if len(valid_mean_rewards) > 5 and np.mean(valid_mean_rewards[-5:]) > -200:
print("You Win!")
break
# -
# ---
# record sessions
import gym.wrappers
env = gym.wrappers.Monitor(
NormalizedActions(gym.make("Pendulum-v0")),
directory="videos_ddpg",
force=True)
sessions = [generate_session(t_max=max_steps, train=False) for _ in range(10)]
env.close()
# +
# show video
from IPython.display import HTML
import os
video_names = list(
filter(lambda s: s.endswith(".mp4"), os.listdir("./videos_ddpg/")))
HTML("""
<video width="640" height="480" controls>
<source src="{}" type="video/mp4">
</video>
""".format("./videos/"+video_names[-1])) # this may or may not be _last_ video. Try other indices
| 2020/code/DDPG.ipynb |