code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="9k5d6qKkCjNq" colab_type="code" outputId="b0af5c99-d8b0-49c3-8fcc-8c8d8333322d" colab={"base_uri": "https://localhost:8080/", "height": 306}
# ! nvidia-smi
# + id="CMbVzSqAo3eJ" colab_type="code" colab={}
# ! mkdir /blazingsql
# + id="0Pgdxg4ppH3e" colab_type="code" outputId="23eafaf8-74e0-4f2c-be58-f946fd8fdef2" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# intall miniconda
# !wget -c https://repo.continuum.io/miniconda/Miniconda3-4.5.4-Linux-x86_64.sh
# !chmod +x Miniconda3-4.5.4-Linux-x86_64.sh
# !bash ./Miniconda3-4.5.4-Linux-x86_64.sh -b -f -p /usr/local
# install RAPIDS packages
# !conda install -q -y --prefix /usr/local -c nvidia -c rapidsai \
# -c numba -c conda-forge -c pytorch -c defaults \
# cudf=0.9 cuml=0.9 cugraph=0.9 python=3.6 cudatoolkit=10.0
# set environment vars
import sys, os, shutil
sys.path.append('/usr/local/lib/python3.6/site-packages/')
os.environ['NUMBAPRO_NVVM'] = '/usr/local/cuda/nvvm/lib64/libnvvm.so'
os.environ['NUMBAPRO_LIBDEVICE'] = '/usr/local/cuda/nvvm/libdevice/'
# # copy .so files to current working dir
for fn in ['libcudf.so', 'librmm.so']:
shutil.copy('/usr/local/lib/'+fn, os.getcwd())
# + id="2QhfPm6g9zJk" colab_type="code" outputId="b03cf0d8-5528-4f95-88c1-f49ccb8a320c" colab={"base_uri": "https://localhost:8080/", "height": 153}
import nvstrings, nvcategory, cudf
import io, requests
# download CSV file from GitHub
url="https://github.com/plotly/datasets/raw/master/tips.csv"
content = requests.get(url).content.decode('utf-8')
# read CSV from memory
tips_df = cudf.read_csv(io.StringIO(content))
tips_df['tip_percentage'] = tips_df['tip']/tips_df['total_bill']*100
# display average tip by dining party size
print(tips_df.groupby('size').tip_percentage.mean())
# + id="OuIkbAva9VVP" colab_type="code" outputId="3419c680-91e3-425e-9ae2-ff05c1b0fba9" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# Install BlazingSQL for CUDA 10.0
# ! conda install -q -y --prefix /usr/local -c conda-forge -c defaults -c nvidia -c rapidsai \
# -c blazingsql/label/cuda10.0 -c blazingsql \
# blazingsql-calcite blazingsql-orchestrator blazingsql-ral blazingsql-python
# + id="TrAD1GInIor3" colab_type="code" outputId="e0bea199-78c7-4f93-eef3-32ad8e1bccb2" colab={"base_uri": "https://localhost:8080/", "height": 85}
pip install flatbuffers
# + id="UiYEprQ0FoWM" colab_type="code" colab={}
sys.path.append('/usr/local/lib/python3.7/site-packages/')
# + colab_type="code" id="Y7705GbC9tkE" colab={}
from blazingsql import BlazingContextpython
# + id="lY98GEFP1g2_" colab_type="code" outputId="40ceac00-125e-4b6a-eba2-edec0e0d46b7" colab={"base_uri": "https://localhost:8080/", "height": 34}
bc = BlazingContext()
# + id="fd4aIbRUGKUO" colab_type="code" outputId="7e2e13c6-496f-4bef-e6b3-aacdcda1e316" colab={"base_uri": "https://localhost:8080/", "height": 289}
# ! wget https://github.com/plotly/datasets/raw/master/tips.csv
# + id="KU4WfapUGN6q" colab_type="code" outputId="ca2043d1-aed6-4877-e466-f6896a598abe" colab={"base_uri": "https://localhost:8080/", "height": 204}
# ! head sample_data/california_housing_train.csv
# ! pwd
# + id="PorSEjMaFzo-" colab_type="code" colab={}
bc.create_table('housing', '/content/sample_data/california_housing_train.csv')
# + id="SYApuO0zGk3u" colab_type="code" outputId="62272323-4d23-4219-91c9-14680fc66c58" colab={"base_uri": "https://localhost:8080/", "height": 918}
# Query
result = bc.sql('SELECT housing_median_age, AVG(median_house_value) FROM housing GROUP BY housing_median_age ORDER BY housing_median_age').get()
result_gdf = result.columns
#Print GDF
print(result_gdf)
# + id="XWSidqfbGpXG" colab_type="code" colab={}
| utils/blazing_conda_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import glob, os
from tqdm import tqdm
directory = "./2017/"
os.chdir(directory)
#print(glob.glob("*csv"))
all_files = glob.glob("*csv")
all_files[0]
len(all_files)
error_files = []
merged = pd.DataFrame()
for file in tqdm(all_files):
try:
data1 = pd.read_csv(file)
data1 = data1.loc[data1.index[5]:data1.index[5]]
merged = pd.concat([merged, data1])
except:
print('Error with file: ' + file)
error_files.append(file)
merged.head()
merged = merged.sort_values(by = 'Time')
merged.head()
merged.index = merged['Time']
len(error_files)
#len(error_files) / len(all_files)* 100
# +
#error_files[0]
# +
#error_files
# -
merged.head()
del merged['Time']
merged.head()
merged.to_csv('merged_17.csv')
| data/weather/v3/4_only_current_values.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Advanced Tricks
# ===
# So you've tried making normal waveforms and now you need to spice up your life by making some way more weird waveforms letting the detector be whatever you want it to be?
# You have come to the right place!
#
# By default fax uses some configuration file which is a huge pain to modify. So we made fax such that if you add a parameter in the instruction which corresponds to a parameter in the config it will overwrite what the value was in the config and let you deside what it should be!
#
# This example shows how to modify the electron lifetime and the anode voltage
# +
import numpy as np
import strax
import straxen
import wfsim
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm
from multihist import Histdd, Hist1d
from scipy import stats
# -
st = strax.Context(
config=dict(
detector='XENON1T',
fax_config='https://raw.githubusercontent.com/XENONnT/'
'strax_auxiliary_files/master/sim_files/fax_config_1t.json',
fax_config_override={'field_distortion_on':True, 's2_luminescence_model':'simple'},
**straxen.contexts.xnt_common_config),
**straxen.contexts.common_opts)
st.register(wfsim.RawRecordsFromFax1T)
# Just some id from post-SR1, so the corrections work
run_id = '000001'
strax.Mailbox.DEFAULT_TIMEOUT=10000
# +
dtype = wfsim.strax_interface.instruction_dtype
for new_dtype in [('electron_lifetime_liquid', np.int32),
('anode_voltage', np.int32)]:
if new_dtype not in dtype:
dtype.append(new_dtype)
def rand_instructions(c):
n = c['nevents'] = c['event_rate'] * c['chunk_size'] * c['nchunk']
c['total_time'] = c['chunk_size'] * c['nchunk']
instructions = np.zeros(2 * n, dtype=dtype)
uniform_times = c['total_time'] * (np.arange(n) + 0.5) / n
instructions['time'] = np.repeat(uniform_times, 2) * int(1e9)
instructions['event_number'] = np.digitize(instructions['time'],
1e9 * np.arange(c['nchunk']) * c['chunk_size']) - 1
instructions['type'] = np.tile([1, 2], n)
instructions['recoil'] = ['er' for i in range(n * 2)]
r = np.sqrt(np.random.uniform(0, 2500, n))
t = np.random.uniform(-np.pi, np.pi, n)
instructions['x'] = np.repeat(r * np.cos(t), 2)
instructions['y'] = np.repeat(r * np.sin(t), 2)
instructions['z'] = np.repeat(np.random.uniform(-100, 0, n), 2)
nphotons = np.random.uniform(2000, 2050, n)
nelectrons = 10 ** (np.random.uniform(1, 4, n))
instructions['amp'] = np.vstack([nphotons, nelectrons]).T.flatten().astype(int)
instructions['electron_lifetime_liquid'] = np.repeat(600e10,len(instructions))
instructions['anode_voltage'] = np.repeat(1e10,len(instructions))
return instructions
wfsim.strax_interface.rand_instructions = rand_instructions
wfsim.strax_interface.instruction_dtype = dtype
# -
st.set_config(dict(fax_file=None))
st.set_config(dict(nchunk=1, event_rate=1, chunk_size=100))
# +
# Remove any previously simulated data, if such exists
# # !rm -r strax_data
records = st.get_array(run_id,'raw_records', progress_bar=False)
peaks = st.get_array(run_id, ['peak_basics'], progress_bar=False)
data = st.get_df(run_id, 'event_info', progress_bar=False)
truth = st.get_df(run_id, 'truth', progress_bar=False)
# -
truth.head()
| docs/source/tutorials/Advanced_tricks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.6 64-bit (windows store)
# name: python3
# ---
# # Quartiles
# Quartile is a type of quantile which divides the number of data points into four parts, or quarters, of more-or-less equal size. The data must be ordered from smallest to largest to compute quartiles; as such, quartiles are a form of order statistics.
# - The first quartile $(Q_1)$ is defined as the middle number between the smallest number($minimum$) and the median of the data set. It is also known as the lower or $25^{th}\text{empirical quartile}$, as $25\%$ of the data is below this point.
# - The second quartile $(Q_2)$ is the median of the whole data set, thus $50\%$ of the data lies below this point.
# - The third quartile $(Q_3)$ is the middle value between the median and the highest value ($maximum$) of the data set. It is known as the $upper$ or $75^{th}\text{empirical quartile}$, as $75\%$ of the data lies below this point.
#
# $$minimum-----Q_1-----Q_2-----Q_3-----maximum$$
#
# Along with minimum and maximum of the data (which are also quartiles), the three quartiles described above provide a $\text{five-number summary}$ of the data. This summary is important in statistics because it provides information about both the center and the spread of the data. Knowing the lower and upper quartile provides information on how big the spread is and if the dataset is $skewed$ toward one side. Since quartiles divide the number of data points evenly, the range is not the same between quartiles (i.e., $Q_3-Q_2 \neq Q_2-Q_1$) and is instead known as the $\textbf{interquartile range (IQR)}$. While the maximum and minimum also show the spread of the data, the upper and lower quartiles can provide more detailed information on the location of specific data points, the presence of outliers in the data, and the difference in spread between the middle $50\%$ of the data and the outer data points.
#
# In desciptive statistics, the $\textbf{Interquartile range (IQR)}$ also called $midspread$, $middle\;50\%$, or $H-spread$, is a measure of $statistical\;dispersion$ being equal to the difference between $75^{th}$ and $25^{th}\;percentiles$. $IQR=Q_3-Q_1$
#
# <p align="center">
# <img src="https://upload.wikimedia.org/wikipedia/commons/thumb/1/1a/Boxplot_vs_PDF.svg/640px-Boxplot_vs_PDF.svg.png?1626778057933">
# </p>
#
#
# |Symbol|Names|Definition|
# |:---:|:---:|:---:|
# |$Q_1$|$25^{th}\;percentile$|splits off the lowest $25\%$ data from the highest $75\%$|
# |$Q_2$|$50^{th}\;percentile$|splits dataset in half|
# |$Q_3$|$75^{th}\;percentile$|splits off the highest $25\%$ data from the lowest $75\%$|
# +
import numpy as np
def quartiles(array):
# sort original array in ascending order
print(f"The original array is {array}") # Comment this out for large datasets
temp = 0
for i in range(0,len(array)):
for j in range(i+1,len(array)):
if (array[i]>array[j]):
temp = array[i]
array[i] = array[j]
array[j] = temp
# lower half of array
array1 = []
for i in range(0,len(array)//2):
array1.append(array[i])
# upper half of array
if len(array)%2==0:
array2 = []
for i in range(len(array)//2,len(array)):
array2.append(array[i])
elif len(array)%2==1:
array2 = []
for i in range((len(array)//2)+1,len(array)):
array2.append(array[i])
# Quartile values
Q1 = np.median(array1)
Q2 = np.median(array)
Q3 = np.median(array2)
# Either define a function to return the desired values or to print arrays and quartiles.
return array1,Q1,array,Q2,array2,Q3,Q3-Q1
'''
return values in the order -
Lower half, First quartile, whole array, second quartile(median of whole array), Upper half, third quartile, IQR = Q3-Q1
'''
# Alternatively if you don't want to use the values further you can print all the values by defining it in the function itself.
'''
print(f"The sorted array is {array}")
print(f"The lower half consists of {array1}, and it's Median: Q1 = {Q1}.")
print(f"The median of entire array {array} is Q2 = {Q2}.")
print(f"The upper half consists of {array2}, and its Median: Q3 = {Q3}.")
print(f"The interquartile range, IQR = {IQR}")
'''
# -
# Testing the function for odd and even number of elements in the array
# Odd number of elements in array
array = [5,7,1,4,2,9,10]
array1,Q1,array,Q2,array2,Q3,IQR = quartiles(array)
print(f"The sorted array is {array}")
print(f"The lower half consists of {array1}, and it's Median: Q1 = {Q1}.")
print(f"The median of entire array {array} is Q2 = {Q2}.")
print(f"The upper half consists of {array2}, and its Median: Q3 = {Q3}.")
print(f"The interquartile range, IQR = {IQR}")
# Even number of elements in array
a = [3,5,7,1,4,2,9,10]
array1,Q1,array,Q2,array2,Q3,IQR = quartiles(a)
print(f"The sorted array is {array}")
print(f"The lower half consists of {array1}, and it's Median: Q1 = {Q1}.")
print(f"The median of entire array {array} is Q2 = {Q2}.")
print(f"The upper half consists of {array2}, and its Median: Q3 = {Q3}.")
print(f"The interquartile range, IQR = {IQR}")
# Test with different array
b = [3,7,8,5,12,14,21,13,18]
array1,Q1,array,Q2,array2,Q3,IQR = quartiles(b)
print(f"The sorted array is {array}")
print(f"The lower half consists of {array1}, and it's Median: Q1 = {Q1}.")
print(f"The median of entire array {array} is Q2 = {Q2}.")
print(f"The upper half consists of {array2}, and its Median: Q3 = {Q3}.")
print(f"The interquartile range, IQR = {IQR}")
# # Using `statistics`
# +
from statistics import median
def quartiles(array):
# sort original array in ascending order
print(f"The original array is {array}") # Comment this out for large datasets
# Alternatively you can just use the .sort() function to arrange in order
# It changes the original array itself
array.sort()
# lower half of array
array1 = []
for i in range(0,len(array)//2):
array1.append(array[i])
# upper half of array
if len(array)%2==0:
array2 = []
for i in range(len(array)//2,len(array)):
array2.append(array[i])
elif len(array)%2==1:
array2 = []
for i in range((len(array)//2)+1,len(array)):
array2.append(array[i])
# Quartile values
Q1 = median(array1)
Q2 = median(array)
Q3 = median(array2)
# Either define a function to return the desired values or to print arrays and quartiles.
return array1,Q1,array,Q2,array2,Q3,Q3-Q1
'''
return values in the order -
Lower half, First quartile, whole array, second quartile(median of whole array), Upper half, third quartile, IQR = Q3-Q1
'''
# Alternatively if you don't want to use the values further you can print all the values by defining it in the function itself.
'''
print(f"The sorted array is {array}")
print(f"The lower half consists of {array1}, and it's Median: Q1 = {Q1}.")
print(f"The median of entire array {array} is Q2 = {Q2}.")
print(f"The upper half consists of {array2}, and its Median: Q3 = {Q3}.")
print(f"The interquartile range, IQR = {IQR}")
'''
# -
A = [56.0,32.7,90.4,54.2,50,49,51,52.9,51.3,53.1,55.1]
array1,Q1,array,Q2,array2,Q3,IQR = quartiles(A)
print(f"The sorted array is {array}")
print(f"The lower half consists of {array1}, and it's Median: Q1 = {Q1}.")
print(f"The median of entire array {array} is Q2 = {Q2}.")
print(f"The upper half consists of {array2}, and its Median: Q3 = {Q3}.")
print(f"The interquartile range, IQR = {IQR}")
| Math Programs/Quartiles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simple harmonic motion
# * $a = -kx$: acceleration $\propto$ displacement
# * $a = -\omega^2 x$ where angular frequency $\omega = 2\pi f = \dfrac{2\pi}{T}$ - (angle in radians) over (time for 1 revolution in seconds)
#
# Denote
# * $x$ the displacement from equilibrium position
# * $x_0$ the amplitude
#
# $\begin{aligned}
# v &= \pm \omega \sqrt{{x_0}^2 - x^2}\\
# E_T &= \frac{1}{2} m \omega^2 {x_0}^2\\
# E_k &= \frac{1}{2} m \omega^2 ({x_0}^2 - x^2)\\
# T_{\text{pendulum}} &= 2\pi \sqrt{\frac{l}{g}}\\
# T_{\text{mass on spring}} &= 2\pi \sqrt{\frac{m}{k}}\\
# \end{aligned}$
#
# $\begin{aligned}
# a &= -\omega^2 x(t)\\
# a &= -\omega^2 x_0 \sin(\omega t)\\
# v(t) &= \omega x_0 \cos(\omega t)\\
# x(t) &= x_0 \sin(\omega t)\\
# \end{aligned}$
# # Single slit diffraction
# $\theta = \dfrac{\lambda}{a}$
#
# ## Things to know
# * $\theta = \dfrac{n \lambda}{a}$ where $n$ is the number of minimum you are looking at
# * Intensity of 1st minimum ~5% of central maximum
#
#
# /|\
# / | \
# /|\/|\/ | \/|\/|\
# 2 1 𝜃 n=1
#
#
# # Multiple-slit interference
# The pattern of the multiple-slit interference is *modulated* by the single-slit interference (envelope).
# $$n \lambda = d \sin\theta$$
# ## Thin film interference
# A phenomenon that causes iridescence.
#
# 
# # Doppler effect
# When a source that is moving emits waves, the frequency observed by a static observer is "shifted".
#
# * Object moving towards observer: higher $f$ - blue shift for light;
# * object moving away from observer: lower $f$ - red shift
#
# For a moving source:
# $$f' = f\left(\dfrac{v}{v \pm u_s}\right)$$
# where
# * $v$: speed of wave
# * $u_s$: speed of source
#
# For a moving observer:
# $$f' = f\left(\dfrac{v \pm u_0}{v}\right)$$
# where
# * $u_o$: speed of observer
#
# For EM radiation:
# $$\dfrac{\Delta f}{f} = \dfrac{\Delta \lambda}{\lambda} = \dfrac{v}{c}$$
# ## Resolution
# The ability to distinguish 2 objects that are close to each other is the ability to **resolve** things.
#
# ```
# observer
# *---
# |/ angle between objects
# / The wavelength of light from objects
# * objects to observe
# <--> distance from objects to observer
# ```
# The Rayleigh criterion can be used to determine if two objects will be resolved.
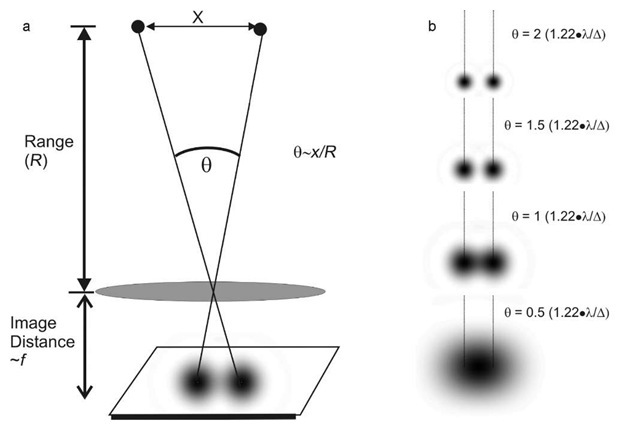
# 
# 
#
# The Rayleigh criterion states that for 2 images to be resolved, the principal maximum of the 1st diffraction pattern must be no closer than the first minimum of the secondary pattern.
#
# $$\theta = 1.22 \dfrac{\lambda}{b}$$
# * $\theta$: angle of resolution
# * $\lambda$: wavelength of light
# * $b$: width of aperture
# +
import matplotlib.pyplot as plt
import numpy as np
plt.style.use(['bmh', 'dark_background', 'seaborn-poster'])
# +
plt.plot(0.2, .59 * 9, 'ro', label=f"EG: 1.80mm")
plt.plot(0.2, .59 * 10, 'bo', label="JY: 1.62mm")
plt.xlim(0.19, 0.21)
xs = np.linspace(0.19, 0.21)
plt.plot(xs, 2e-3 / (1.22 * 500e-9) * 1.62e-2 * xs / 2, 'r')
plt.plot(xs, 2e-3 / (1.22 * 500e-9) * 1.80e-2 * xs / 2, 'b')
plt.xlabel("Aperture between two dots (cm)")
plt.ylabel("Distance at which we can't resolve (m)")
plt.legend()
plt.show()
# -
| notes/PHY/9-wave-phenomena.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: teachopencadd
# language: python
# name: teachopencadd
# ---
# + active=""
# # Run this if you get an ImportError below
# !pip install bravado
#
# -
#KLIFS
from bravado.client import SwaggerClient
KLIFS_API_DEFINITIONS = "http://klifs.vu-compmedchem.nl/swagger/swagger.json"
KLIFS_CLIENT = SwaggerClient.from_url(KLIFS_API_DEFINITIONS, config={'validate_responses': False})
def _all_kinase_families():
return KLIFS_CLIENT.Information.get_kinase_families().response().result
KLIFS_CLIENT.Information.get_kinase_families().response().result
def _kinases_from_family(family, species="HUMAN"):
return KLIFS_CLIENT.Information.get_kinase_names(kinase_family=family, species=species).response().result
KLIFS_CLIENT.Information.get_kinase_names(kinase_family=family, species=species).response().result
def _protein_and_ligand_structure(*kinase_ids):
structures = KLIFS_CLIENT.Structures.get_structures_list(kinase_ID=kinase_ids).response().result
molcomplex = KLIFS_CLIENT.Structures.get_structure_get_pdb_complex(structure_ID=structures[0].structure_ID).response().result
protein = KLIFS_CLIENT.Structures.get_structure_get_protein(structure_ID=structures[0].structure_ID).response().result
ligands = KLIFS_CLIENT.Ligands.get_ligands_list(kinase_ID=kinase_ids).response().result
return molcomplex, protein, [ligand.SMILES for ligand in ligands]
structures
pdb_code = '3w32'
def protein_binding_site(*kinase_ids):
molcomplex = KLIFS_CLIENT.Structures.get_structure_get_pdb_complex(structure_ID=structures[0].structure_ID).response().result
binding_site = KLIFS_CLIENT.Structures.get_structure_get_pocket(structure_ID=kinase_ID).response().result
return binding_site
# +
#import time
def KLIFS_binding_site():
try :
molcomplex, binding_site = protein_binding_site(kinase.kinase__ID)
except :
None
# -
Kinase_ID = KLIFS_CLIENT.Structures.get_structure_get_pdb_complex(structure_ID=structures[0].structure_ID).response().result
structures = KLIFS_CLIENT.Structures.get_structures_list(kinase_ID=[406]).response().result
structures
structures[0].structure_ID
molcomplex = KLIFS_CLIENT.Structures.get_structure_get_pdb_complex(structure_ID=structures[0].structure_ID).response().result
KLIFS_CLIENT.Structures.get_structure_get_pocket(structure_ID=782).response().result
binding_site = protein_binding_site(pdb_code)
binding_site = KLIFS_binding_site()
| examples/KLIFS_query.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
L = [1,2, 5, 7]
# +
import numpy as np
def softmax(L):
# convert to numpy array
L_np = np.array(L)
#calculate exponent of each element in l_np
expL = np.exp(L_np)
# calculate sum
sum_expL= sum(expL)
result = []
for i in expL:
result.append(i*1.0/sum_expL)
return result
# -
softmax(L)
| 2-softmax-function.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Convolutional Neural Network - Gap / Char Classification
# Using TensorFlow
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import tensorflow as tf
import cv2
# %matplotlib notebook
# Increase size of plots
plt.rcParams['figure.figsize'] = (9.0, 5.0)
# Creating CSV
import glob
import csv
# Helpers
from ocr.helpers import implt
from ocr.mlhelpers import TrainingPlot, DataSet
from ocr.datahelpers import loadGapData
print("OpenCV: " + cv2.__version__)
print("Numpy: " + np.__version__)
print("TensorFlow: " + tf.__version__)
# -
# ## Load Images and Lables in CSV
images, labels = loadGapData('data/gapdet/large/')
# +
print("Number of images: " + str(len(images)))
# Splitting on train and test data
div = int(0.90 * len(images))
trainData = images[0:div]
trainLabels = labels[0:div]
evalData = images[div:]
evalLabels = labels[div:]
print("Training images: %g" % div)
# -
# # Create classifier
# ### Dataset
# Prepare training dataset
trainSet = DataSet(trainData, trainLabels)
evalSet = DataSet(evalData, evalLabels)
# ## Convulation Neural Network
# ### Graph
# +
sess = tf.InteractiveSession()
# Help functions for standard layers
def conv2d(x, W, name=None):
return tf.nn.conv2d(x, W, strides=[1, 1, 1, 1], padding='SAME', name=name)
def max_pool_2x2(x, name=None):
return tf.nn.max_pool(x, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='SAME', name=name)
# Regularization scale - FOR TWEAKING
SCALE = 0.001
# Weighting cross entropy
POS_WEIGHT = (len(labels) - sum(labels)) / sum(labels)
# Place holders for data (x) and labels (y_)
x = tf.placeholder(tf.float32, [None, 7200], name='x')
targets = tf.placeholder(tf.int64, [None])
# Reshape input data
reshape_images = tf.reshape(x, [-1, 60, 120, 1])
# Image standardization
x_images = tf.map_fn(
lambda img: tf.image.per_image_standardization(img), reshape_images)
# 1. Layer - Convulation + Subsampling
W_conv1 = tf.get_variable('W_conv1', shape=[8, 8, 1, 10],
initializer=tf.contrib.layers.xavier_initializer())
b_conv1 = tf.Variable(tf.constant(0.1, shape=[10]), name='b_conv1')
h_conv1 = tf.nn.relu(conv2d(x_images, W_conv1) + b_conv1, name='h_conv1')
# 2. Layer - Max Pool
h_pool1 = max_pool_2x2(h_conv1, name='h_pool1')
# 3. Layer - Convulation + Subsampling
W_conv2 = tf.get_variable('W_conv2', shape=[5, 5, 10, 20],
initializer=tf.contrib.layers.xavier_initializer())
b_conv2 = tf.Variable(tf.constant(0.1, shape=[20]), name='b_conv2')
h_conv2 = tf.nn.relu(conv2d(h_pool1, W_conv2) + b_conv2, name='h_conv2')
# 4. Layer - Max Pool
h_pool2 = max_pool_2x2(h_conv2, name='h_pool2')
# 5. Fully Connected layer
W_fc1 = tf.get_variable('W_fc1', shape=[15*30*20, 1000],
initializer=tf.contrib.layers.xavier_initializer(),
regularizer=tf.contrib.layers.l2_regularizer(scale=SCALE))
b_fc1 = tf.Variable(tf.constant(0.1, shape=[1000]), name='b_fc1')
h_conv2_flat = tf.reshape(h_pool2, [-1, 15*30*20], name='h_conv2_flat')
h_fc1 = tf.nn.relu(tf.matmul(h_conv2_flat, W_fc1) + b_fc1, name='h_fc1')
# 6. Dropout
keep_prob = tf.placeholder(tf.float32, name='keep_prob')
h_fc1_drop = tf.nn.dropout(h_fc1, keep_prob, name='h_fc1_drop')
# 7. Output layer
W_fc2 = tf.get_variable('W_fc2', shape=[1000, 2],
initializer=tf.contrib.layers.xavier_initializer(),
regularizer=tf.contrib.layers.l2_regularizer(scale=SCALE))
b_fc2 = tf.Variable(tf.constant(0.1, shape=[2]), name='b_fc2')
y_conv = tf.matmul(h_fc1_drop, W_fc2) + b_fc2
# Activation function for real use in application
activation = tf.argmax(tf.matmul(h_fc1, W_fc2) + b_fc2, 1, name='activation')
# Cost: cross entropy + regularization
# Regularization with L2 Regularization with decaying learning rate
# cross_entropy = tf.nn.weighted_cross_entropy_with_logits(logits=y_conv, targets=y_)
weights = tf.multiply(targets, POS_WEIGHT) + 1
cross_entropy = tf.losses.sparse_softmax_cross_entropy(
logits=y_conv,
labels=targets,
weights=weights)
# Using cross entropy for sigmoid as loss
regularization = tf.get_collection(tf.GraphKeys.REGULARIZATION_LOSSES)
cost = tf.reduce_mean(cross_entropy) + sum(regularization)
# Optimizer
train_step = tf.train.AdamOptimizer(5e-5).minimize(cost, name='train_step')
# Evaluating
correct_prediction = tf.equal(tf.argmax(y_conv,1), targets)
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32), name='accuracy')
# -
# ### Training
# +
sess.run(tf.global_variables_initializer())
saver = tf.train.Saver()
### SETTINGS ###
TRAIN_STEPS = 500000
TEST_ITER = 150
COST_ITER = 50
SAVE_ITER = 2000
BATCH_SIZE = 64
# Graph for live ploting
trainPlot = TrainingPlot(TRAIN_STEPS, TEST_ITER, COST_ITER)
try:
for i in range(TRAIN_STEPS):
trainBatch, labelBatch = trainSet.next_batch(BATCH_SIZE)
if i%COST_ITER == 0:
# Plotting cost
tmpCost = cost.eval(feed_dict={x: trainBatch,
targets: labelBatch,
keep_prob: 1.0})
trainPlot.updateCost(tmpCost, i // COST_ITER)
if i%TEST_ITER == 0:
# Plotting accuracy
evalD, evalL = evalSet.next_batch(1000)
accEval = accuracy.eval(feed_dict={x: evalD,
targets: evalL,
keep_prob: 1.0})
accTrain = accuracy.eval(feed_dict={x: trainBatch,
targets: labelBatch,
keep_prob: 1.0})
trainPlot.updateAcc(accEval, accTrain, i // TEST_ITER)
if i%SAVE_ITER == 0:
# Saving model
saver.save(sess, 'models/gap-clas/large/CNN-CG')
train_step.run(feed_dict={x: trainBatch,
targets: labelBatch,
keep_prob: 0.4})
except KeyboardInterrupt:
pass
saver.save(sess, 'models/gap-clas/large/CNN-CG')
evalD, evalL = evalSet.next_batch(1000)
print("Accuracy %g" % accuracy.eval(feed_dict={x: evalD,
targets: evalL,
keep_prob: 1.0}))
sess.close()
| GapClassifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# %pylab inline
# # More Examples
# ## Additive model
#
# Example taken from JCGM 101:2008, Clause 9.2.
#
# This example considers the additive model
#
# $$ Y = X_1 + X_2 + X_3 + X_4 $$
#
# for three different sets of PDFs $g_{x_i}(\xi_i)$ assigned to the input quantities $X_i$, regarded as independent.
#
# #### Taks 1
# Assume that a Gaussian PDF is assigned to each $X_i$. The best estimates are $x_i=0$ with associated standard uncertainties $u(x_i)=1$. Report the Monte Carlo results for estimate, uncertainty and 95% coverage interval of $Y$ with three significant digits. Compare the Monte Carlo result to that of a standard GUM approach using the *law of propagation of uncertainty*.
# #### Task 2
# Assign a rectangular PDF to each $X_i$ so that $X_i$ has an expectation of zero and a standard deviation of unity. Report the results of $Y$ with three significant digits and compare to that obtained with the standard GUM approach.
# #### Taks 3
# Same as Taks 2, but with $X_4$ having standard uncertainty of 10 rathern than unity.
# ## Mass calibration
#
# Example taken from JCGM 101:2008, Clause 9.3
#
# Consider the calibration of a weight $W$ of mass density $\rho_W$ against a reference weight $R$ of mass density $\rho_R$ having nominally the same mass, using a balance operating in air of mass density $\rho_a$. Since $\rho_W$ and $\rho_R$ are generally diffeent, it is necessary to account for buoyancy effects.
#
# Applying Archimedes' principle, the model takes the form
#
# $$ m_W\left( 1 - \frac{\rho_a}{\rho_W} \right) = (m_R + \delta m_R)\left( 1-\frac{\rho_a}{\rho_R} \right) $$
#
# where $\delta m_R$ is the mass of a small weight of density $\rho_R$ added to $R$ to balance it with $W$. Working with so called "conventional masses", the model in this example ist
#
# $$ \delta m = (m_{R,c} + \delta m_{R,c})\left[ 1 + (\rho_a-\rho_{a_0})\left( \frac{1}{\rho_W}-\frac{1}{\rho_R} \right) \right] $$
# Knowledge about the input quantities is given as
#
# 
#
# Carry out the uncertainty evaluation using the Monte Carlo method to obtain an estimate, uncertainty and 99% coverage interval.
# ## Comparison loss
#
# Comparison loss in microwave power meter calibration example taken from JCGM 101:2008, Clause 9.4.
# During the calibration of a microwave power meter, the power meter and a standard power meter are connected in turn to a stable signal generator. The power absorbed by each meter will in general be different because their complex input voltage reflection coefficients are not identical. The ration $Y$ of the power $P_M$ absorbed by the meter being calibrated and that, $P_S$, by the standard meter is
#
# $$ Y = \frac{P_M}{P_S} = \frac{1-\vert \Gamma_M\vert^2}{1-\vert \Gamma_S\vert^2} \times \frac{\vert 1-\Gamma_S\Gamma_G\vert^2}{\vert 1-\Gamma_M\Gamma_G\vert^2} $$
#
# where $\Gamma_G$ is the voltage reflection coefficient of the signal generator, $\Gamma_M$ that of the meter being calibrated and $\Gamma_S$ the of the standard meter. This power ratio is an instance of "comparison loss".
# #### Task
#
# Considering the case that $\Gamma_S=\Gamma_G=0$, and measured values are obtained of the real and imaginary parts $X_1, X_2$ of $\Gamma_M$, the model used in this examples is $ Y = 1 - X_1^2 - X_2^2 $, better expressed as
#
# $$ \delta Y = 1 - Y = X_1^2 + X_2^2 $$
#
# $X_1$ and $X_2$ are usually not independent, but it may be difficult to gather information about the actual correlation coefficient. In such cases, the uncertainty evaluation can be repeated with different trial values for the correlation coefficient.
#
# Here we use $\rho=0$ and $\rho=0.9$. The estimate of $X_1$ is taken as $x_1 = 0.01$ with standard uncertainty $u(x_1) = 0.005$. The estimate of $X_2$ is taken as zero with an associated standard uncertainty of $u(x_2) = 0.005$.
#
# Carry out an uncertainty evaluation for $\delta Y$, calculating estimate and uncertainty using the *law of propagation of uncertainty* and the Monte Method for both correlation coefficients.
| .ipynb_checkpoints/10 More examples-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#
# COMMENTS TO DO
#
#Condensed code based on the code from: https://jmetzen.github.io/2015-11-27/vae.html
# %matplotlib inline
import tensorflow as tf
import tensorflow.contrib.layers as layers
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
import numpy as np
import os
import time
import glob
from tensorflow.examples.tutorials.mnist import input_data
def plot(samples, w, h, fw, fh, iw=28, ih=28):
fig = plt.figure(figsize=(fw, fh))
gs = gridspec.GridSpec(w, h)
gs.update(wspace=0.05, hspace=0.05)
for i, sample in enumerate(samples):
ax = plt.subplot(gs[i])
plt.axis('off')
ax.set_xticklabels([])
ax.set_yticklabels([])
ax.set_aspect('equal')
plt.imshow(sample.reshape(iw, ih), cmap='Greys_r')
return fig
def encoder(images, num_outputs_h0=8, num_outputs_h1=16, kernel_size=5, stride=2, num_hidden_fc=1024, z_dim=100):
print("Encoder")
h0 = layers.convolution2d(
inputs=images,
num_outputs=num_outputs_h0,
kernel_size=kernel_size,
stride=stride,
activation_fn=tf.nn.relu,
scope='e_cnn_%d' % (0,)
)
print("Convolution 1 -> {}".format(h0))
h1 = layers.convolution2d(
inputs=h0,
num_outputs=num_outputs_h1,
kernel_size=kernel_size,
stride=stride,
activation_fn=tf.nn.relu,
scope='e_cnn_%d' % (1,)
)
print("Convolution 2 -> {}".format(h1))
h1_dim = h1.get_shape().as_list()[1]
h2_flat = tf.reshape(h1, [-1, h1_dim * h1_dim * num_outputs_h1])
print("Reshape -> {}".format(h2_flat))
h2_flat =layers.fully_connected(
inputs=h2_flat,
num_outputs=num_hidden_fc,
activation_fn=tf.nn.relu,
scope='e_d_%d' % (0,)
)
print("FC 1 -> {}".format(h2_flat))
z_mean =layers.fully_connected(
inputs=h2_flat,
num_outputs=z_dim,
activation_fn=None,
scope='e_d_%d' % (1,)
)
print("Z mean -> {}".format(z_mean))
z_log_sigma_sq =layers.fully_connected(
inputs=h2_flat,
num_outputs=z_dim,
activation_fn=None,
scope='e_d_%d' % (2,)
)
return z_mean, z_log_sigma_sq
def decoder(z, num_hidden_fc=1024, h1_reshape_dim=7, kernel_size=5, h1_channels=16, h2_channels = 8, output_channels=1, strides=2, output_dims=784):
print("Decoder")
batch_size = tf.shape(z)[0]
h0 =layers.fully_connected(
inputs=z,
num_outputs=num_hidden_fc,
activation_fn=tf.nn.relu,
scope='d_d_%d' % (0,)
)
print("FC 1 -> {}".format(h0))
h1 =layers.fully_connected(
inputs=h0,
num_outputs=h1_reshape_dim*h1_reshape_dim*h1_channels,
activation_fn=tf.nn.relu,
scope='d_d_%d' % (1,)
)
print("FC 2 -> {}".format(h1))
h1_reshape = tf.reshape(h1, [-1, h1_reshape_dim, h1_reshape_dim, h1_channels])
print("Reshape -> {}".format(h1_reshape))
wdd2 = tf.get_variable('wd2', shape=(kernel_size, kernel_size, h2_channels, h1_channels), initializer=tf.contrib.layers.xavier_initializer())
bdd2 = tf.get_variable('bd2', shape=(h2_channels,), initializer=tf.constant_initializer(0))
h2 = tf.nn.conv2d_transpose(h1_reshape, wdd2, output_shape=(batch_size, h1_reshape_dim*2, h1_reshape_dim*2, h2_channels), strides=(1, strides, strides, 1), padding='SAME')
h2_out = tf.nn.relu(h2 + bdd2)
h2_out = tf.reshape(h2_out, (batch_size, h1_reshape_dim*2, h1_reshape_dim*2, h2_channels))
print("DeConv 1 -> {}".format(h2_out))
h2_dim = h2_out.get_shape().as_list()[1]
wdd3 = tf.get_variable('wd3', shape=(kernel_size, kernel_size, output_channels, h2_channels), initializer=tf.contrib.layers.xavier_initializer())
bdd3 = tf.get_variable('bd3', shape=(output_channels,), initializer=tf.constant_initializer(0))
h3 = tf.nn.conv2d_transpose(h2_out, wdd3, output_shape=(batch_size, h2_dim*2, h2_dim*2, output_channels), strides=(1, strides, strides, 1), padding='SAME')
h3_out = tf.nn.sigmoid(h3 + bdd3)
#Workaround to use dinamyc batch size...
h3_out = tf.reshape(h3_out, (batch_size, h2_dim*2, h2_dim*2, output_channels))
print("DeConv 2 -> {}".format(h3_out))
h3_reshape = tf.reshape(h3_out, [-1, output_dims])
print("Reshape -> {}".format(h3_reshape))
return h3_reshape
mnist = input_data.read_data_sets('DATASETS/MNIST_TF', one_hot=True)
#For reconstructing the same or a different image (denoising)
images = tf.placeholder(tf.float32, shape=(None, 784))
images_28x28x1 = tf.reshape(images, [-1, 28, 28, 1])
images_target = tf.placeholder(tf.float32, shape=(None, 784))
is_training_placeholder = tf.placeholder(tf.bool)
learning_rate_placeholder = tf.placeholder(tf.float32)
z_dim = 100
with tf.variable_scope("encoder") as scope:
z_mean, z_log_sigma_sq = encoder(images_28x28x1)
with tf.variable_scope("reparameterization") as scope:
eps = tf.random_normal(shape=tf.shape(z_mean), mean=0.0, stddev=1.0, dtype=tf.float32)
# z = mu + sigma*epsilon
z = tf.add(z_mean, tf.multiply(tf.sqrt(tf.exp(z_log_sigma_sq)), eps))
with tf.variable_scope("decoder") as scope:
x_reconstr_mean = decoder(z)
scope.reuse_variables()
##### SAMPLING #######
z_input = tf.placeholder(tf.float32, shape=[None, z_dim])
x_sample = decoder(z_input)
#reconstr_loss = tf.reduce_sum(tf.nn.sigmoid_cross_entropy_with_logits(logits=x_reconstr_mean, labels=images_target), reduction_indices=1)
offset=1e-7
obs_ = tf.clip_by_value(x_reconstr_mean, offset, 1 - offset)
reconstr_loss = -tf.reduce_sum(images_target * tf.log(obs_) + (1-images_target) * tf.log(1 - obs_), 1)
latent_loss = -.5 * tf.reduce_sum(1. + z_log_sigma_sq - tf.pow(z_mean, 2) - tf.exp(z_log_sigma_sq), reduction_indices=1)
cost = tf.reduce_mean(reconstr_loss + latent_loss)
optimizer=tf.train.AdamOptimizer(learning_rate=learning_rate_placeholder).minimize(cost)
init = tf.global_variables_initializer()
save_path = "MODELS_CVAE_MNIST/CONV_VAE_MNIST.ckpt"
CVAE_SAVER = tf.train.Saver()
# +
with tf.Session() as sess:
sess.run(init)
CVAE_SAVER.restore(sess, save_path)
print("Model restored in file: {}".format(save_path))
random_gen = sess.run(x_sample,feed_dict={z_input: np.random.randn(100, z_dim)})
fig=plot(random_gen, 10, 10, 10, 10)
plt.show()
# -
# # (READING) MNIST subset
# +
labels = 10
subset_size_per_label = 10
X_mini = np.load("DATASETS/MNIST_ALT/X_MINI_100")
Y_mini = np.load("DATASETS/MNIST_ALT/Y_MINI_100")
fig=plot(X_mini, 10, 10, 10, 10)
plt.show()
print(np.argmax(Y_mini, axis=1))
# -
# # Interpolating
# +
from numpy.linalg import norm
import progressbar
def slerp(p0, p1, t):
omega = np.arccos(np.dot(p0/norm(p0), p1/norm(p1)))
so = np.sin(omega)
return np.sin((1.0-t)*omega) / so * p0 + np.sin(t*omega)/so * p1
def linear(p0, p1, t):
return p0 * (1-t) + p1 * t
def interpolate(sample1, sample2, alphaValues, sess, method="linear"):
x_together = np.vstack((sample1, sample2))
z_samples = sess.run(z, feed_dict={images: x_together})
#fig=plot(z_samples, 1, 2, 10, 10, 10, 10)
#plt.show()
interpolation_steps = alphaValues.shape[0]
z_interpolations = np.zeros((interpolation_steps, z_dim))
for i, alpha in enumerate(alphaValues):
if method == "slerp":
z_interpolations[i] = slerp(z_samples[0], z_samples[1], alpha)
else:
z_interpolations[i] = linear(z_samples[0], z_samples[1], alpha)
x_interpolated = sess.run(x_sample, feed_dict={z_input: z_interpolations})
#fig=plot(x_interpolated, 1, INTERPOLATION_STEPS, 10, 10)
#plt.show()
return x_interpolated
labels = 10
INTERPOLATION_STEPS = 20
alphaValues = np.linspace(0, 1, INTERPOLATION_STEPS)
n_gen = labels * subset_size_per_label * (subset_size_per_label - 1) * INTERPOLATION_STEPS
print("Total gen: {}".format(n_gen))
x_pool = np.zeros((n_gen, X_mini.shape[1]))
y_pool = np.zeros((n_gen, Y_mini.shape[1]))
with tf.Session() as sess:
sess.run(init)
CVAE_SAVER.restore(sess, save_path)
print("Model restored in file: {}".format(save_path))
bar = progressbar.ProgressBar(max_value=n_gen)
bar.start()
counter = 0
for label in range(labels):
offset = label * subset_size_per_label
for i in range(subset_size_per_label):
samples_ind = list(range(subset_size_per_label))
samples_ind.remove(i)
x_sample_1 = X_mini[offset + i].copy()
for j in samples_ind:
x_sample_2 = X_mini[offset + j].copy()
x_output=interpolate(x_sample_1, x_sample_2, alphaValues, sess, method="linear")
x_pool[counter:counter+INTERPOLATION_STEPS] = x_output.copy()
y_pool[counter:counter+INTERPOLATION_STEPS, label] = 1
counter+=INTERPOLATION_STEPS
bar.update(counter)
bar.finish()
# +
fig=plot(x_pool[:100], 10, 10, 10, 10)
plt.show()
print(np.argmax(y_pool[:100], axis=1))
fig=plot(x_pool[-100:], 10, 10, 10, 10)
plt.show()
print(np.argmax(y_pool[-100:], axis=1))
perm = np.random.permutation(x_pool.shape[0])
x_pool = x_pool[perm]
y_pool = y_pool[perm]
perm = np.random.permutation(x_pool.shape[0])
x_pool = x_pool[perm]
y_pool = y_pool[perm]
fig=plot(x_pool[:100], 10, 10, 10, 10)
plt.show()
print(np.argmax(y_pool[:100], axis=1))
# -
# # Storing MINI-MNIST and GEN-MNIST
# +
fx = open("DATASETS/MNIST_ALT/X_GEN_18K_CVAE", "wb")
np.save(fx, x_pool)
fx.close()
fy = open("DATASETS/MNIST_ALT/Y_GEN_18K_CVAE", "wb")
np.save(fy, y_pool)
fy.close()
# -
| STEP2_2_GenerateInterpolations_x20.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Data Science in Medicine using Python
#
# ### Author: Dr <NAME>
# ### 1. Summary of the data processing we did so far and a little reminder
# +
# %%time
import datetime
import os
import pandas as pd
flist = [fle for fle in os.listdir('data') if 'slow_Measurement' in fle]
data_dict = {} # Creates an empty dictionary
for file in flist:
print(datetime.datetime.now(), file)
path = os.path.join('data', file,)
tag = file[11:-25]
data_dict[tag] = pd.read_csv(path, parse_dates = [['Date', 'Time']])
data_dict[tag] = data_dict[tag].set_index('Date_Time')
new_columns = [item[5:] for item in data_dict[tag].columns if item.startswith('5001')]
new_columns = ['Time [ms]', 'Rel.Time [s]'] + new_columns
data_dict[tag].columns = new_columns
data_dict[tag] = data_dict[tag].resample('1S').mean()
columns_to_drop = ['Tispon [s]', 'I:Espon (I-Part) [no unit]', 'I:Espon (E-Part) [no unit]']
data_dict[tag] = data_dict[tag].drop(columns_to_drop, axis = 1)
data_dict[tag].to_csv('%s' %tag)
# -
# ##### "Computer programs are for human to read and occasionally for computers to run"
#
# You want to be more verbose particularly when learning Python
# +
# %%time
# Import the required libraries
import datetime
import os
import pandas as pd
# From the files in 'Data' sub-directory only consider those ones which contain 'slow_Measurement'
flist = [fle for fle in os.listdir('data') if 'slow_Measurement' in fle]
data_dict = {} # Creates an empty dictionary
for file in flist: # Loop through all relevant data files
print(datetime.datetime.now(), file)
# The relative filepath to the files
path = os.path.join('data', file,)
# Use the specific part of the filename as a unique key for the dictionary
tag = file[11:-25]
# Import data, parse the 'Date' and 'Time' columns as datetime and combine them
data_dict[tag] = pd.read_csv(path, parse_dates = [['Date', 'Time']])
# Set the combined 'Date_Time' column as row index
data_dict[tag] = data_dict[tag].set_index('Date_Time')
# Remove the '5001' pre-tag from the column names
new_columns = [item[5:] for item in data_dict[tag].columns if item.startswith('5001')]
new_columns = ['Time [ms]', 'Rel.Time [s]'] + new_columns
data_dict[tag].columns = new_columns
# As data were retrieved in two batches every second, combine these data by using the mean() function
data_dict[tag] = data_dict[tag].resample('1S').mean()
# Drop columes which have barely any data
columns_to_drop = ['Tispon [s]', 'I:Espon (I-Part) [no unit]', 'I:Espon (E-Part) [no unit]']
data_dict[tag] = data_dict[tag].drop(columns_to_drop, axis = 1)
# Export processed data as .csv files with unique names
data_dict[tag].to_csv('%s' %tag)
# -
# ### 2. A quick look at allt the data
# Let us look at the data in more details
# This is a dictionary of DataFrames
data_dict;
data_dict.keys()
data_dict.values();
[len(value) for value in data_dict.values()]
[value.shape for value in data_dict.values()]
# ### 3. How to process the data further?
#
# Choose one of the 3 recordings initially and study further
data_dict['2019-01-14_124200.144']
data_dict['2019-01-14_124200.144'].info()
# Now all the data are in the right format, but...
#
# ##### Further issues:
# - Tidal volumes, minute volumes and compliance only make sense when normalised to body weight
# - We would like to know the distribution of the data to make sure that they make sense (are there non-sensical data or clear outliers)
# - We still have some missing data, what should we do with them?
# ### 4. Some parameters (`VTs, MVs, Cdyn`) only make sense if normalised to body weight
data_dict['2019-01-14_124200.144'].columns
to_normalise = ['MVe [L/min]', 'MVi [L/min]', 'Cdyn [L/bar]', 'MVespon [L/min]', 'MVemand [L/min]', 'VTmand [mL]',
'VTispon [mL]', 'VTmand [L]', 'VTspon [L]', 'VTemand [mL]', 'VTespon [mL]', 'VTimand [mL]', 'VT [mL]',
'MV [L/min]', 'VTspon [mL]', 'VTe [mL]', 'VTi [mL]', 'MVleak [L/min]',]
data_dict.keys()
# Here we are creating a dictionary with the same keys on the fly but it would be more usefuls (and less prone to error) to import weights from a csv or Excel file into a DataFrame
# Weights in kilogram
weights = {'2019-01-14_124200.144' : 0.575, '2019-01-16_090910.423' : 0.575 , '2020-11-02_134238.904' : 775}
for recording in data_dict: # I could have written datadict.keys(), it is the same
for par in to_normalise:
data_dict[recording][f'{par[:-1]}/kg{par[-1]}'] = data_dict[recording][par] / weights[recording]
data_dict['2019-01-14_124200.144']
data_dict['2019-01-14_124200.144'].columns
# ### 5. Analyse the distribution of the data
data_dict['2019-01-14_124200.144'].describe()
# You can customize the percentiles
percentiles_to_show = [0.001, 0.01, 0.05, 0.1, 0.25, 0.5, 0.75, 0.95, 0.99, 0.999]
data_dict['2019-01-14_124200.144'].describe(percentiles = percentiles_to_show )
# Too many zeros - you can round it
percentiles_to_show = [0.001, 0.01, 0.05, 0.1, 0.25, 0.5, 0.75, 0.95, 0.99, 0.999]
round(data_dict['2019-01-14_124200.144'].describe(percentiles = percentiles_to_show ), 2)
data_dict['2019-01-14_124200.144'].columns
# Too many columns - study them individually
percentiles_to_show = [0.001, 0.01, 0.05, 0.1, 0.25, 0.5, 0.75, 0.95, 0.99, 0.999]
round(data_dict['2019-01-14_124200.144']['VTemand [mL/kg]'].describe(percentiles = percentiles_to_show ), 2)
# ##### One picture speaks a thousand words
#
# The main purpose of generating graphs is not to present the data to others but for yourself to visualise and inspect them.
# You can use the default .plot() method of DataFrames
data_dict['2019-01-14_124200.144']['VTemand [mL]']
# This gives you a plot but not the one your want (shows time series data)
data_dict['2019-01-14_124200.144']['VTemand [mL]'].plot()
# Two useful way of displaying the distribution of continuous data are `boxplots` and `histograms`
# #### Boxplots
# +
# That is better but it does not look nice because it shows all the outliers
data_dict['2019-01-14_124200.144']['VTemand [mL]'].plot(kind = 'box')
# -
# You can customize it - to some extent
# Google DataFrame.plot()
#
# [this is the first hit](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.plot.html)
#
# +
# That is better but it does not look nice because it shows all the outliers
data_dict['2019-01-14_124200.144']['VTemand [mL]'].plot(kind = 'box', ylim = [0,5], ylabel = 'mL/kg')
# -
# Even better - use `matplotlib` for plotting
# +
import matplotlib.pyplot as plt # Imports the matplotlib plotting library
# First we create the figure and the subplot(s)
fig, ax = plt.subplots()
# Then we populate the subplot(s)
ax.boxplot(data_dict['2019-01-14_124200.144']['VTemand [mL]'])
# It is an empty plot (although it does not throw an error) - why ?
# -
# Check there is data !!
data_dict['2019-01-14_124200.144']['VTemand [mL]']
# Let us Google it: "matplotlib boxplot empty"
#
# [Look at Stackoverflow top hits](https://stackoverflow.com/questions/52960482/plt-boxplot-showing-up-blank-despite-thousands-of-varying-datapoints)
# +
import matplotlib.pyplot as plt # Imports the matplotlib plotting library
# Removing empty data points helps
fig, ax = plt.subplots()
ax.boxplot(data_dict['2019-01-14_124200.144']['VTemand [mL]'].dropna());
# -
# How can we improve this further ??
# +
# Now the outliers are removed and the mean is shown
fig, ax = plt.subplots()
ax.boxplot(data_dict['2019-01-14_124200.144']['VTemand [mL]'].dropna(), showfliers = False, showmeans = True,);
# -
# This looks better but I would like to modify the error bars and the layout
# +
# # Whiskers show by default, you can change it
fig, ax = plt.subplots()
ax.boxplot(data_dict['2019-01-14_124200.144']['VTemand [mL]'].dropna(),
whis = [10, 90], showfliers = False, showmeans = True,);
# +
# You can customize the plot further
# Define styling for each boxplot component
meanprops = {'marker':'s', 'markeredgecolor':'black', 'markerfacecolor':'firebrick'}
medianprops = {'color': 'black', 'linewidth': 2}
boxprops = {'color': 'blue', 'linestyle': '-'}
whiskerprops = { 'color': 'red', 'linestyle': '-'}
capprops = {'color': 'green', 'linestyle': '-'}
flierprops = {'color': 'black', 'marker': '.'}
fig, ax = plt.subplots()
ax.boxplot(data_dict['2019-01-14_124200.144']['VTemand [mL]'].dropna(),
whis = [5, 95], showfliers = False, showmeans = True, meanprops = meanprops,
medianprops=medianprops, boxprops=boxprops, whiskerprops=whiskerprops,
capprops=capprops,flierprops = flierprops);
# +
# Add more useful customisation
meanprops = {'marker':'s', 'markeredgecolor':'black', 'markerfacecolor':'black'}
medianprops = {'color': 'black', 'linewidth': 2}
boxprops = {'color': 'black', 'linestyle': '-'}
whiskerprops = { 'color': 'black', 'linestyle': '-'}
capprops = {'color': 'black', 'linestyle': '-'}
flierprops = {'color': 'black', 'marker': '.'}
fig, ax = plt.subplots()
ax.boxplot(data_dict['2019-01-14_124200.144']['VTemand [mL]'].dropna(),
whis = [5, 95], showfliers = False, showmeans = True, meanprops = meanprops,
medianprops=medianprops, boxprops=boxprops, whiskerprops=whiskerprops,
capprops=capprops,flierprops = flierprops);
ax.set_xlabel('VTemand')
ax.set_ylabel('mL/kg')
ax.set_ylim(1.5, 5.5)
ax.grid(True)
# Further customization is possible
# +
# How to save the graph
dpi = 600
filetype = 'pdf'
# Add more useful customisation
meanprops = {'marker':'s', 'markeredgecolor':'black', 'markerfacecolor':'black'}
medianprops = {'color': 'black', 'linewidth': 2}
boxprops = {'color': 'black', 'linestyle': '-'}
whiskerprops = { 'color': 'black', 'linestyle': '-'}
capprops = {'color': 'black', 'linestyle': '-'}
flierprops = {'color': 'black', 'marker': '.'}
fig, ax = plt.subplots()
ax.boxplot(data_dict['2019-01-14_124200.144']['VTemand [mL]'].dropna(),
whis = [5, 95], showfliers = False, showmeans = True, meanprops = meanprops,
medianprops=medianprops, boxprops=boxprops, whiskerprops=whiskerprops,
capprops=capprops,flierprops = flierprops);
ax.set_xlabel('VTemand')
ax.set_ylabel('mL/kg')
ax.set_ylim(1.5, 5.5)
fig.savefig(os.path.join('results', f'boxplot_1.{filetype}'), dpi = dpi, format = filetype,
bbox_inches='tight',);
# -
# #### Histograms
# +
# First attempt..
fig, ax = plt.subplots()
ax.hist(data_dict['2019-01-14_124200.144']['VTemand [mL]'], color = 'black', alpha = 0.7);
# You can customize the format further as for boxplots...
# +
# A logarithmic y axis is useful to appreciate the present of outliers
fig, ax = plt.subplots()
ax.hist(data_dict['2019-01-14_124200.144']['VTemand [mL]'], log= True);
# +
# Use more bins to reveal the actual distribution
fig, ax = plt.subplots()
ax.hist(data_dict['2019-01-14_124200.144']['VTemand [mL]'], bins = 50);
# +
# imports the numpy package
import numpy as np
bins = np.arange(0, 10, 0.2)
bins
# +
# You can define your own bins
# imports the numpy package
import numpy as np
bins = np.arange(0, 10, 0.2)
fig, ax = plt.subplots()
ax.hist(data_dict['2019-01-14_124200.144']['VTemand [mL]'], bins = bins);
# +
# You can define your own bins
# imports the numpy package
import numpy as np
bins = np.arange(0, 10, 0.2)
fig, ax = plt.subplots()
ax.hist(data_dict['2019-01-14_124200.144']['VTemand [mL]'], bins = bins);
# +
# I would like to show you a different way to produce histograms
# -
VTemand_binned = pd.cut(data_dict['2019-01-14_124200.144']['VTemand [mL]'], bins = 10)
VTemand_binned.head(10)
VTemand_binned.value_counts()
# Sort according the index, not the values
# Also, what one function returns can be passed on to the next function
VTemand_binned.value_counts().sort_index()
# Better but still not what you want
VTemand_binned.value_counts().sort_index().plot()
# Better but still not what you want
VTemand_binned.value_counts().sort_index().plot(kind = 'bar')
plot = VTemand_binned.value_counts().sort_index().plot(kind = 'bar')
# +
import matplotlib.pyplot as plt
plot = VTemand_binned.value_counts().sort_index().plot(kind = 'bar')
plt.savefig(fname = os.path.join('results', 'VTemand'))
# +
import matplotlib.pyplot as plt
plot = VTemand_binned.value_counts().sort_index().plot(kind = 'bar', color = 'black', alpha = 0.7,
xlabel = 'VTemand_kg', ylabel = 'number of inflations',)
#plt.grid(True)
plt.savefig(fname = os.path.join('results', 'VTemand'))
# +
import matplotlib.pyplot as plt
plot = VTemand_binned.value_counts().sort_index().plot(kind = 'bar', color = 'black', alpha = 0.7,
xlabel = 'VTemand_kg', ylabel = 'number of inflations', logy = True)
#plt.grid(True)
plt.savefig(fname = os.path.join('results', 'VTemand'))
# -
# ##### You need to do this for all relevant columns
data_dict['2019-01-14_124200.144'].columns
# +
columns_to_plot = [column for column in data_dict['2019-01-14_124200.144'].columns
if column not in ['Time [ms]', 'Rel.Time [s]']]
print(columns_to_plot)
# -
data_dict['2019-01-14_124200.144'].columns[2:]
'VTemand [mL/kg]'.split(' ')[0]
# +
dpi = 300
filetype = 'jpg'
for column in columns_to_plot:
print(datetime.datetime.now(), f'Working on {column}')
fig, ax = plt.subplots()
ax.hist(data_dict['2019-01-14_124200.144'][column], bins = 10)
fname = column.split(' ')[0]
plt.savefig(fname = os.path.join('results', f'{fname}.{filetype}'), dpi = dpi, format = filetype,
bbox_inches='tight',)
plt.close()
# -
for column in columns_to_plot:
print(datetime.datetime.now(), f'Working on {column}')
fig, ax = plt.subplots()
ax.hist(data_dict['2019-01-14_124200.144'][column], bins = 50,)
if column == 'C20/Cdyn [no unit]':
fname = 'C20_Cdyn'
else:
fname = column.split(' ')[0]
plt.savefig(fname = os.path.join('results', f'{fname}.{filetype}'), dpi = dpi, format = filetype,
bbox_inches='tight')
plt.close()
| Lecture_6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to XGBoost-Spark Cross Validation with GPU
#
# The goal of this notebook is to show you how to levarage GPU to accelerate XGBoost spark cross validatoin for hyperparameter tuning. The best model for the given hyperparameters will be returned.
#
# Note: CrossValidation can't be ran with the latest cudf v21.06.1 because of some API changes. We'll plan to release a new XGBoost jar with the fixing soon. We keep this notebook using cudf v0.19.2 & rapids-4-spark v0.5.0.
#
# Here takes the application 'Taxi' as an example.
#
# A few libraries are required for this notebook:
# 1. NumPy
# 2. cudf jar
# 2. xgboost4j jar
# 3. xgboost4j-spark jar
# #### Import the Required Libraries
from ml.dmlc.xgboost4j.scala.spark import XGBoostRegressionModel, XGBoostRegressor
from ml.dmlc.xgboost4j.scala.spark.rapids import CrossValidator
from pyspark.ml.evaluation import RegressionEvaluator
from pyspark.ml.tuning import ParamGridBuilder
from pyspark.sql import SparkSession
from pyspark.sql.types import FloatType, IntegerType, StructField, StructType
from time import time
# As shown above, here `CrossValidator` is imported from package `ml.dmlc.xgboost4j.scala.spark.rapids`, not the spark's `tuning.CrossValidator`.
# #### Create a Spark Session
spark = SparkSession.builder.appName("taxi-cv-gpu-python").getOrCreate()
# #### Specify the Data Schema and Load the Data
# +
label = 'fare_amount'
schema = StructType([
StructField('vendor_id', FloatType()),
StructField('passenger_count', FloatType()),
StructField('trip_distance', FloatType()),
StructField('pickup_longitude', FloatType()),
StructField('pickup_latitude', FloatType()),
StructField('rate_code', FloatType()),
StructField('store_and_fwd', FloatType()),
StructField('dropoff_longitude', FloatType()),
StructField('dropoff_latitude', FloatType()),
StructField(label, FloatType()),
StructField('hour', FloatType()),
StructField('year', IntegerType()),
StructField('month', IntegerType()),
StructField('day', FloatType()),
StructField('day_of_week', FloatType()),
StructField('is_weekend', FloatType()),
])
features = [ x.name for x in schema if x.name != label ]
train_data = spark.read.parquet('/data/taxi/parquet/train')
trans_data = spark.read.parquet('/data/taxi/parquet/eval')
# -
# #### Build a XGBoost-Spark CrossValidator
# First build a regressor of GPU version using *setFeaturesCols* to set feature columns
params = {
'eta': 0.05,
'maxDepth': 8,
'subsample': 0.8,
'gamma': 1.0,
'numRound': 100,
'numWorkers': 1,
'treeMethod': 'gpu_hist',
}
regressor = XGBoostRegressor(**params).setLabelCol(label).setFeaturesCols(features)
# Then build the evaluator and the hyperparameters
evaluator = (RegressionEvaluator()
.setLabelCol(label))
param_grid = (ParamGridBuilder()
.addGrid(regressor.maxDepth, [3, 6])
.addGrid(regressor.numRound, [100, 200])
.build())
# Finally the corss validator
cross_validator = (CrossValidator()
.setEstimator(regressor)
.setEvaluator(evaluator)
.setEstimatorParamMaps(param_grid)
.setNumFolds(3))
# #### Start Cross Validation by Fitting Data to CrossValidator
def with_benchmark(phrase, action):
start = time()
result = action()
end = time()
print('{} takes {} seconds'.format(phrase, round(end - start, 2)))
return result
model = with_benchmark('Cross-Validation', lambda: cross_validator.fit(train_data)).bestModel
# #### Transform On the Best Model
def transform():
result = model.transform(trans_data).cache()
result.foreachPartition(lambda _: None)
return result
result = with_benchmark('Transforming', transform)
result.select(label, 'prediction').show(5)
# #### Evaluation
accuracy = with_benchmark(
'Evaluation',
lambda: RegressionEvaluator().setLabelCol(label).evaluate(result))
print('RMSE is ' + str(accuracy))
spark.stop()
| examples/notebooks/python/cv-taxi-gpu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Zahlen in Python
#
# Zahlen funktionieren in Python ähnlich wie sonst überall auch - du kannst damit ganz normale Rechnungen berechnen.
print(5)
print(5)
print(6)
# ### Die Grundrechenarten
print(5 + 4)
print(5 - 4)
print(5 * 4)
print(5 / 4)
# Auch die Klammersetzung funktioniert natürlich wie gewohnt :-):
print((5 + 4) * 3)
print(1.25 + 2)
print(1 + 2)
# ## Übung
# * Was ist 3588 geteilt durch 11,3 ?
# * Was ist die Wurzel aus 345?
345**(1/2)
| 02 Python Teil 1/01 Zahlen.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tanzania] *
# language: python
# name: conda-env-tanzania-py
# ---
# # Research
# ## Models:
# These are the models that every data scientist should be familiar with.
# Favorite Classification Models:
# - LinearSVC
# - k-NN
# - Support Vector Machine Algorithm
# - XGBoost
# - Random Forest
# ### Hyperparameters to tune for each model type
# Below are the most common hyperparameters for the different models
# **k-NN**
# - **n_neighbors**: decreasing K decreases bias and increases variance, which leads to a more complex model
# - **leaf_size**: 'determines how many observations are captured in each leaf of either the BallTree of KDTree algorithms, which ultimately make the classification. The default equals 30. You can tune leaf_size by passing in a range of integers, like n_neighbors, to find the optimal leaf size. It is important to note that leaf_size can have a serious effect on run time and memory usage. Because of this, you tend not to run it on leaf_sizes smaller than 30 (smaller leafs equates to more leafs)'
# - **weights**: 'is the function that weights the data when making a prediction. “Uniform” is an equal weighted function, while “distance” weights the points by the inverse of their distance (i.e., location matters!). Utilizing the “distance” function will result in closer data points having a more significant influence on the classification'
# - **metric**: 'can be set to various distance metrics (see here) like Manhattan, Euclidean, Minkowski, or weighted Minkowski (default is “minkowski” with a p=2, which is the Euclidean distance). Which metric you choose is heavily dependent on what question you are trying to answer'
# **Random Forest, Decision Trees**
# - **n_estimators (random forest only)**: number of decision trees used in making the forest (default = 100). Generally speaking, the more uncorrelated trees in our forest, the closer their individual errors get to averaging out. However, more does not mean better since this can have an exponential effect on computation costs. After a certain point, there exists statistical evidence of diminishing returns. Bias-Variance Tradeoff: in theory, the more trees, the more overfit the model (low bias). However, when coupled with bagging, we need not worry'
# - **max_depth**: 'an integer that sets the maximum depth of the tree. The default is None, which means the nodes are expanded until all the leaves are pure (i.e., all the data belongs to a single class) or until all leaves contain less than the min_samples_split, which we will define next. Bias-Variance Tradeoff: increasing the max_depth leads to overfitting (low bias)'
# - **min_samples_split**: 'is the minimum number of samples required to split an internal node. Bias-Variance Tradeoff: the higher the minimum, the more “clustered” the decision will be, which could lead to underfitting (high bias)'
# - **min_samples_leaf**: 'defines the minimum number of samples needed at each leaf. The default input here is 1. Bias-Variance Tradeoff: similar to min_samples_split, if you do not allow the model to split (say because your min_samples_lear parameter is set too high) your model could be over generalizing the training data (high bias)'
# - **criterion**: 'measures the quality of the split and receives either “gini”, for Gini impurity (default), or “entropy”, for information gain. Gini impurity is the probability of incorrectly classifying a randomly chosen datapoint if it were labeled according to the class distribution of the dataset. Entropy is a measure of chaos in your data set. If a split in the dataset results in lower entropy, then you have gained information (i.e., your data has become more decision useful) and the split is worthy of the additional computational costs'
# **AdaBoost and Gradient Boosting**
# - **n_estimators**: is the maximum number of estimators at which boosting is terminated. If a perfect fit is reached, the algo is stopped. The default here is 50. Bias-Variance Tradeoff: the higher the number of estimators in your model the lower the bias.
# - **learning_rate**: is the rate at which we are adjusting the weights of our model with respect to the loss gradient. In layman’s terms: the lower the learning_rate, the slower we travel along the slope of the loss function. Important note: there is a trade-off between learning_rate and n_estimators as a tiny learning_rate and a large n_estimators will not necessarily improve results relative to the large computational costs.
# - **base_estimator (AdaBoost) / Loss (Gradient Boosting)**: is the base estimator from which the boosted ensemble is built. For AdaBoost the default value is None, which equates to a Decision Tree Classifier with max depth of 1 (a stump). For Gradient Boosting the default value is deviance, which equates to Logistic Regression. If “exponential” is passed, the AdaBoost algorithm is used.
#
# **Support Vector Machines (SVM)**
# - **C**: is the regularization parameter. As the documentation notes, the strength of regularization is inversely proportional to C. Basically, this parameter tells the model how much you want to avoid being wrong. You can think of the inverse of C as your total error budget (summed across all training points), with a lower C value allowing for more error than a higher value of C. Bias-Variance Tradeoff: as previously mentioned, a lower C value allows for more error, which translates to higher bias.
# - **gamma**: determines how far the scope of influence of a single training points reaches. A low gamma value allows for points far away from the hyperplane to be considered in its calculation, whereas a high gamma value prioritizes proximity. Bias-Variance Tradeoff: think of gamma as inversely related to K in KNN, the higher the gamma, the tighter the fit (low bias).
# - **kernel**: specifies which kernel should be used. Some of the acceptable strings are “linear”, “poly”, and “rbf”. Linear uses linear algebra to solve for the hyperplane, while poly uses a polynomial to solve for the hyperplane in a higher dimension (see Kernel Trick). RBF, or the radial basis function kernel, uses the distance between the input and some fixed point (either the origin or some of fixed point c) to make a classification assumption. More information on the Radial Basis Function can be found here.
# ## Evaluation Metrics
# ### Precision – What percent of your predictions were correct?
# Precision is the ability of a classifier not to label an instance positive that is actually negative. For each class it is defined as the ratio of true positives to the sum of true and false positives.
#
# TP – True Positives
# FP – False Positives
#
# Precision – Accuracy of positive predictions.
# Precision = TP/(TP + FP)
# ### Recall – What percent of the positive cases did you catch?
# Recall is the ability of a classifier to find all positive instances. For each class it is defined as the ratio of true positives to the sum of true positives and false negatives.
#
# FN – False Negatives
#
# Recall: Fraction of positives that were correctly identified.
# Recall = TP/(TP+FN)
# ### F1 score – What percent of positive predictions were correct?
# The F1 score is a weighted harmonic mean of precision and recall such that the best score is 1.0 and the worst is 0.0. Generally speaking, F1 scores are lower than accuracy measures as they embed precision and recall into their computation. **As a rule of thumb, the weighted average of F1 should be used to compare classifier models, not global accuracy**.
#
# F1 Score = 2*(Recall * Precision) / (Recall + Precision)
# ### Most important
# Recall will be the metric to focus on, because saying a well will fail when it is still fine, is no biggie.
#
# Saying a well is working when its actually broken is a biggie.
# ## Visualization Metrics
#
# ### Precision-Recall Curve
# Precision-Recall curves should be used when there is a moderate to large class imbalance.
#
# Our dataset has a very large class imbalance, so we chose to use the precision-recall curve.
#
# [https://machinelearningmastery.com/roc-curves-and-precision-recall-curves-for-imbalanced-classification](https://machinelearningmastery.com/roc-curves-and-precision-recall-curves-for-imbalanced-classification/)
# ### Confusion Matrix
# A confusion matrix is a table that is often used to describe the performance of a classification model (or “classifier”) on a set of test data for which the true values are known. It allows the visualization of the performance of an algorithm.
#
# [https://www.geeksforgeeks.org/confusion-matrix-machine-learning/](https://www.geeksforgeeks.org/confusion-matrix-machine-learning/)
# ## Methodology
# This project was built using th ROSEMED methodology.
# - **'R'**: Research the domain and relevant data science tools
# - **'O'**: Obtain the data
# - **'S'**: Scrub the data and remove any NaNs, missing values, duplicates, or outliers
# - **'E'**: Explore the data and look for correlations and insights
# - **'M'**: Model the data using the most relevant classifiers for the data
# - **'E'**: Evaluate the models and choose the model that is most suitable for the data
# - **'D'**: Deploy the models
| notebooks/research.ipynb |
# # Overfit-generalization-underfit
#
# In the previous notebook, we presented the general cross-validation framework
# and how it helps us quantify the training and testing errors as well
# as their fluctuations.
#
# In this notebook, we will put these two errors into perspective and show how
# they can help us know if our model generalizes, overfit, or underfit.
#
# Let's first load the data and create the same model as in the previous
# notebook.
# +
from sklearn.datasets import fetch_california_housing
housing = fetch_california_housing(as_frame=True)
data, target = housing.data, housing.target
target *= 100 # rescale the target in k$
# -
# <div class="admonition note alert alert-info">
# <p class="first admonition-title" style="font-weight: bold;">Note</p>
# <p class="last">If you want a deeper overview regarding this dataset, you can refer to the
# Appendix - Datasets description section at the end of this MOOC.</p>
# </div>
# +
from sklearn.tree import DecisionTreeRegressor
regressor = DecisionTreeRegressor()
# -
# ## Overfitting vs. underfitting
#
# To better understand the statistical performance of our model and maybe find
# insights on how to improve it, we will compare the testing error with the
# training error. Thus, we need to compute the error on the training set,
# which is possible using the `cross_validate` function.
# +
import pandas as pd
from sklearn.model_selection import cross_validate, ShuffleSplit
cv = ShuffleSplit(n_splits=30, test_size=0.2)
cv_results = cross_validate(regressor, data, target,
cv=cv, scoring="neg_mean_absolute_error",
return_train_score=True, n_jobs=2)
cv_results = pd.DataFrame(cv_results)
# -
# We will select the train and test score and take the error instead.
scores = pd.DataFrame()
scores[["train error", "test error"]] = -cv_results[
["train_score", "test_score"]]
# +
import matplotlib.pyplot as plt
scores.plot.hist(bins=50, edgecolor="black", density=True)
plt.xlabel("Mean absolute error (k$)")
_ = plt.title("Train and test errors distribution via cross-validation")
# -
# By plotting the distribution of the training and testing errors, we
# get information about whether our model is over-fitting, under-fitting (or
# both at the same time).
#
# Here, we observe a **small training error** (actually zero), meaning that
# the model is **not under-fitting**: it is flexible enough to capture any
# variations present in the training set.
#
# However the **significantly larger testing error** tells us that the
# model is **over-fitting**: the model has memorized many variations of the
# training set that could be considered "noisy" because they do not generalize
# to help us make good prediction on the test set.
#
# ## Validation curve
#
# Some model hyperparameters are usually the key to go from a model that
# underfits to a model that overfits, hopefully going through a region were we
# can get a good balance between the two. We can acquire knowledge by plotting
# a curve called the validation curve. This curve applies the above experiment
# and varies the value of a hyperparameter.
#
# For the decision tree, the `max_depth` parameter is used to control the
# tradeoff between under-fitting and over-fitting.
# +
# %%time
from sklearn.model_selection import validation_curve
max_depth = [1, 5, 10, 15, 20, 25]
train_scores, test_scores = validation_curve(
regressor, data, target, param_name="max_depth", param_range=max_depth,
cv=cv, scoring="neg_mean_absolute_error", n_jobs=2)
train_errors, test_errors = -train_scores, -test_scores
# -
# Now that we collected the results, we will show the validation curve by
# plotting the training and testing errors (as well as their deviations).
# +
plt.plot(max_depth, train_errors.mean(axis=1), label="Training error")
plt.plot(max_depth, test_errors.mean(axis=1), label="Testing error")
plt.legend()
plt.xlabel("Maximum depth of decision tree")
plt.ylabel("Mean absolute error (k$)")
_ = plt.title("Validation curve for decision tree")
# -
# The validation curve can be divided into three areas:
#
# - For `max_depth < 10`, the decision tree underfits. The training error and
# therefore the testing error are both high. The model is too
# constrained and cannot capture much of the variability of the target
# variable.
#
# - The region around `max_depth = 10` corresponds to the parameter for which
# the decision tree generalizes the best. It is flexible enough to capture a
# fraction of the variability of the target that generalizes, while not
# memorizing all of the noise in the target.
#
# - For `max_depth > 10`, the decision tree overfits. The training error
# becomes very small, while the testing error increases. In this
# region, the models create decision specifically for noisy samples harming
# its ability to generalize to test data.
#
# Note that for `max_depth = 10`, the model overfits a bit as there is a gap
# between the training error and the testing error. It can also
# potentially underfit also a bit at the same time, because the training error
# is still far from zero (more than 30 k\\$), meaning that the model might
# still be too constrained to model interesting parts of the data. However the
# testing error is minimal, and this is what really matters. This is the
# best compromise we could reach by just tuning this parameter.
#
# Be aware that looking at the mean errors is quite limiting. We should also
# look at the standard deviation to assess the dispersion of the score. We
# can repeat the same plot as before but this time, we will add some
# information to show the standard deviation of the errors as well.
# +
plt.errorbar(max_depth, train_errors.mean(axis=1),
yerr=train_errors.std(axis=1), label='Training error')
plt.errorbar(max_depth, test_errors.mean(axis=1),
yerr=test_errors.std(axis=1), label='Testing error')
plt.legend()
plt.xlabel("Maximum depth of decision tree")
plt.ylabel("Mean absolute error (k$)")
_ = plt.title("Validation curve for decision tree")
# -
# We were lucky that the variance of the errors was small compared to their
# respective values, and therefore the conclusions above are quite clear. This
# is not necessarily always the case.
# ## Summary:
#
# In this notebook, we saw:
#
# * how to identify whether a model is generalizing, overfitting, or
# underfitting;
# * how to check influence of an hyperparameter on the tradeoff
# underfit/overfit.
| notebooks/cross_validation_validation_curve.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Dependencies
import requests
from config import api_key
url = "http://www.omdbapi.com/?apikey=" + api_key + "&t="
movies = ["Aliens", "Sing", "Moana"]
responses = []
for movie in movies:
movie_data = requests.get(url + movie).json()
responses.append(movie_data)
print(f'The director of {movie} is {movie_data["Director"]}')
# -
responses
| 01-Lesson-Plans/06-Python-APIs/1/Activities/10-Stu_MovieLoop/Solved/Stu_MovieLoop.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: jax
# language: python
# name: jax
# ---
# ## 4.2 필기체를 분류하는 CNN 구현
# 2장, 3장에서 다룬 필기체 데이터를 CNN으로 분류합니다.
#
# ### 4.2.1 분류 CNN 모델링
# 1. 합성곱 계층들과 완전 연결 계층들이 결합하여 구성된 분류 CNN을 모델링하는 방법을 알아보겠습니다.
# - 모델링에 필요한 케라스 패키지들을 불러오는 것이 우선입니다.
# set to use CPU
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '-1'
# import library from keras
import keras
from keras import models, layers
from keras import backend
# - CNN 객체를 models.Sequential로 상속하여 연쇄 방식으로 모델을 구현합니다.
class CNN(models.Sequential):
def __init__(self, input_shape, num_classes):
super().__init__()
self.add(layers.Conv2D(32, kernel_size=(3, 3),
activation='relu',
input_shape=input_shape))
self.add(layers.Conv2D(64, (3, 3), activation='relu'))
self.add(layers.MaxPooling2D(pool_size=(2, 2)))
self.add(layers.Dropout(0.25))
self.add(layers.Flatten())
self.add(layers.Dense(128, activation='relu'))
self.add(layers.Dropout(0.5))
self.add(layers.Dense(num_classes, activation='softmax'))
self.compile(loss=keras.losses.categorical_crossentropy,
optimizer='rmsprop',
metrics=['accuracy'])
# ### 4.2.3 분류 CNN을 위한 데이터 준비
# 2. 분류 CNN에 사용할 데이터인 앞선 장들에 나왔던 MNIST이고 이에 대한 CNN을 위한 사전처리를 진행합니다.
# +
from keras import datasets
class DATA():
def __init__(self):
num_classes = 10
(x_train, y_train), (x_test, y_test) = datasets.mnist.load_data()
img_rows, img_cols = x_train.shape[1:]
if backend.image_data_format() == 'channels_first':
x_train = x_train.reshape(x_train.shape[0], 1, img_rows, img_cols)
x_test = x_test.reshape(x_test.shape[0], 1, img_rows, img_cols)
input_shape = (1, img_rows, img_cols)
else:
x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols, 1)
x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols, 1)
input_shape = (img_rows, img_cols, 1)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
self.input_shape = input_shape
self.num_classes = num_classes
self.x_train, self.y_train = x_train, y_train
self.x_test, self.y_test = x_test, y_test
# -
# ### 4.2.3 학습 효과 분석
# 3. 학습 효과를 분석하기 위해 그래프를 그리는 기능을 임포트합니다.
from keraspp.skeras import plot_loss, plot_acc
import matplotlib.pyplot as plt
# ### 4.2.4 분류 CNN 학습 및 성능 평가
# 4. 데이터와 모델이 준비되었으니 이제 이들을 이용해 학습과 성능 평가를 진행할 차례입니다.
# +
batch_size = 128
epochs = 10
data = DATA()
model = CNN(data.input_shape, data.num_classes)
history = model.fit(data.x_train, data.y_train,
batch_size=batch_size,
epochs=epochs,
validation_split=0.2)
score = model.evaluate(data.x_test, data.y_test)
print()
print('Test loss:', score[0])
print('Test accuracy:', score[1])
plot_loss(history)
plt.show()
plot_acc(history)
plt.show()
# -
# ---
# ### 4.2.4 전체 코드
# +
# File: ex4_1_cnn_mnist_cl.py
# 1. 분류 CNN 모델링
import keras
from keras import models, layers
from keras import backend
class CNN(models.Sequential):
def __init__(self, input_shape, num_classes):
super().__init__()
self.add(layers.Conv2D(32, kernel_size=(3, 3),
activation='relu',
input_shape=input_shape))
self.add(layers.Conv2D(64, (3, 3), activation='relu'))
self.add(layers.MaxPooling2D(pool_size=(2, 2)))
self.add(layers.Dropout(0.25))
self.add(layers.Flatten())
self.add(layers.Dense(128, activation='relu'))
self.add(layers.Dropout(0.5))
self.add(layers.Dense(num_classes, activation='softmax'))
self.compile(loss=keras.losses.categorical_crossentropy,
optimizer='rmsprop',
metrics=['accuracy'])
# 2. 분류 CNN을 위한 데이터 준비
from keras import datasets
class DATA():
def __init__(self):
num_classes = 10
(x_train, y_train), (x_test, y_test) = datasets.mnist.load_data()
img_rows, img_cols = x_train.shape[1:]
if backend.image_data_format() == 'channels_first':
x_train = x_train.reshape(x_train.shape[0], 1, img_rows, img_cols)
x_test = x_test.reshape(x_test.shape[0], 1, img_rows, img_cols)
input_shape = (1, img_rows, img_cols)
else:
x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols, 1)
x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols, 1)
input_shape = (img_rows, img_cols, 1)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
self.input_shape = input_shape
self.num_classes = num_classes
self.x_train, self.y_train = x_train, y_train
self.x_test, self.y_test = x_test, y_test
# 3. 학습 효과 분석
from keraspp.skeras import plot_loss, plot_acc
import matplotlib.pyplot as plt
# 4. 분류 CNN 학습 및 테스트
def main():
batch_size = 128
epochs = 10
data = DATA()
model = CNN(data.input_shape, data.num_classes)
history = model.fit(data.x_train, data.y_train,
batch_size=batch_size,
epochs=epochs,
validation_split=0.2)
score = model.evaluate(data.x_test, data.y_test)
print()
print('Test loss:', score[0])
print('Test accuracy:', score[1])
plot_loss(history)
plt.show()
plot_acc(history)
plt.show()
main()
| cpu_only/nb_ex4_1_cnn_mnist_cl-cpu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Revisiting Food-Safety Inspections from the Chicago Dataset - A Tutorial (Part 2)
# <NAME>, <NAME>, <NAME>
# * I switched name order here and put my bio second at the bottom
# ## 0. Foreward
# * probably touch this up
#
# Sustainabilist often works on data that is related to quality assurance and control (QA/QC) inspections of public or private infrastructure. Typically, this infrastructure takes the form of solar energy systems or energy efficiency upgrades for buildings. These data sets almost exclusively belong to private entities that have commissioned a study to evaluate how safe and/or well-installed the infrastructure that they financed is. For this reason, it has been very difficult to put anything up in the public sphere about how our work is conducted and any public documentation of what kind of analysis we do.
#
# Enter Epicodus, a coding bootcamp in Portland, OR. Several weeks ago, I met David and Russell - two eager coding students who were just learning how to code. They were attending the first meeting of CleanWeb Portland’s first meeting, which Sustainabilist organized. We were talking about the lack of public datasets in sustainability, and I mentioned how Chicago’s food science data set was very similar to many of the QA/QC data sets that I have looked at. Just like that, a project was born.
#
# The coding work demonstrated herein is 100% that of the student interns, under my guidance for how to structure, examine, and explore the data. The work was conducted using Google Collaboratory, iPython notebooks, and Anaconda’s scientific computing packages.
# ## 1. Review
# * foreward?
# * To prevent foodborne illness inspectors enforce stringent food codes, sometimes with the help of predictive violation models
# * We seek to expand the work of the CDPH, exploring highres predictions and neural nets
# * We want to to focus on helping restaurants prevent illness and avoid costly violations
# * We cleaned and pre-processed data from the following sources (databases)
# * ...(probably more stuff)
# ## 2. Feature engineering
# * something on how the model works, what we're building it for, the thing about blinding the model to outcome and then comparing it to actual outcome
# * how by training model to guess outcome for canvass inspections we're building a tool that we can feed same paramaters at any time to guess outcome of a simulated canvass inspection
# * Somthing on feature selection, why it makes sense to try out what we're trying out
# * should we explain features here or below? idk
# ## 3. Food Inspection Features
# * load inspections and select what we want from it to use as basis for model data
# * Something on what this data is, where it comes from, why we're using it?
# +
import numpy as np
import pandas as pd
import os.path
root_path = os.path.dirname(os.getcwd())
# Load food inspection data
inspections = pd.read_csv(os.path.join(root_path, "DATA/food_inspections.csv"))
# Create basis for model_data
data = inspections.loc[:, ["inspection_id", "license", "inspection_date", "facility_type"]]
# -
# ### 3.1. Pass / Fail Flags
# * pass fail flags denote inspection outcome, this is something that will be "covered" so model can guess it
# * converted to individual presence/absence flags to help with something or other (what and why specifically?)
# Create pass / fail flags
data["pass_flag"] = inspections.results.apply(lambda x: 1 if x == "Pass" else 0)
data["fail_flag"] = inspections.results.apply(lambda x: 1 if x == "Fail" else 0)
# ### 3.2. Facility Risk Flags
# * Facilities like restaurants pose greater risk than packaged food kiosks and are given higher risk levels
# * Higher risk levels mean greater inspection frequency also (unsure if this is relevant)
# * Again converted to numeric form to fit with (specs? what?)
# Create risk flags
data["risk_1"] = inspections.results.apply(lambda x: 1 if x == "Risk 1 (High)" else 0)
data["risk_2"] = inspections.results.apply(lambda x: 1 if x == "Risk 2 (Medium)" else 0)
data["risk_3"] = inspections.results.apply(lambda x: 1 if x == "Risk 3 (Low)" else 0)
# ### 3.3. Violation Data
# * Violation data is also something the model will be guessing, another part of the inspection outcome
# * The data consists of a bunch of rows (representing inspection outcomes) with binary values for whether a specific health code was violated in that inspection
# * Merged on inspection ID (each row of data is matched and merged with a violation data row with same ID. rows with no matches are excluded.)
#
# +
# Load violation data
values = pd.read_csv(os.path.join(root_path, "DATA/violation_values.csv"))
counts = pd.read_csv(os.path.join(root_path, "DATA/violation_counts.csv"))
# Merge with violation data, filtering missing data
data = pd.merge(data, values, on="inspection_id")
data = pd.merge(data, counts, on="inspection_id")
# -
# ### 3.4. Past Fails
# * Passed fails refers to the previous inspection outcome for that license (as a binary flag)
# * This is a strong predictor of inspection outcomes
# * Passed fails is something the model will have access to when predicting inspection outcomes, and will be used to guess the actual and current outcome.
# * We first create a dataframe of past data by arranging inspections chronologically, grouping by license and shifting each group of inspections by 1, so that the data for each inspection lines up with the row of the next inspection (the first row for each license will by empty and the last inspection is not used). The pre-grouping order is preserved upon shifting.
# * (this could use visualization)
# * We can then simply attach the fail_flag column to our data as past fails, setting the empty first value as 0 (no previous fail)
# +
# Sort inspections by date
grouped = data.sort_values(by="inspection_date", inplace=True)
# Find previous inspections by shifting each license group
past_data = data.groupby("license").shift(1)
# Add past fails, with 0 for first inspections
data["past_fail"] = past_data.fail_flag.fillna(0)
# -
# ### 3.5. Past Violation Data
# * individual past violation values might well be good for predicting individual violations (eg watch out mr. restaurant, you violated these codes last inspection so you're at risk for them)
# * We can use the same past_data to get past violation values
# * We'll modify the names to pv_1, etc
# * If we drop inspection_id we can just tack them on to the end of the data using join
# * first records are set to 0 (no past violation)
# * For past_critical, past_serious and past_minor we can similarly just grab each column and add it as a new column in data
# +
# Select past violation values, remove past inspection id
past_values = past_data[values.columns].drop("inspection_id", axis=1).add_prefix("p")
# Add past values to model data, with 0 for first records
data = data.join(past_values.fillna(0))
# -
# Add past violation counts, with 0 for first records
data["past_critical"] = past_data.critical_count.fillna(0)
data["past_serious"] = past_data.serious_count.fillna(0)
data["past_minor"] = past_data.minor_count.fillna(0)
# ### 3.6. Time Since Last
# * One potential risk factor is greater time since last inspection (do we say we got this from Chicago team or just give our own justification?)
# * To access this convert each inspection date to a python datetime, subtract the previous datetime from the later to create a series of delta objects and convert to days.
# * the default is set to two.
# +
# Calculate time since previous inspection
deltas = pd.to_datetime(data.inspection_date) - pd.to_datetime(past_data.inspection_date)
# Add years since previous inspection (default to 2)
data["time_since_last"] = deltas.apply(lambda x: x.days / 365.25).fillna(2)
# -
# ### 3.7. First Record
# * Actually not sure why this would matter in predicting outcomes? (check)
# * Maybe first records are more likely to fail?
# * To get it we simply put 1s for rows where data is absent in the shifted past_data.
# Check if first record
data["first_record"] = past_data.inspection_id.map(lambda x: 1 if pd.isnull(x) else 0)
# ## 4. Business License Features
# * These are the features derived from the busuiness license dataset
# * What is a business license? other background info?
# ### 4.1. Matching Inspections with Licenses
# * Load data, see publication 1
# Load business license data
licenses = pd.read_csv(os.path.join(root_path, "DATA/business_licenses.csv"))
# * In order to link food inspections to the business licenses of the facilities inspected we create a table of matches, each linking an inspection to a license
# * Many business licenses can be matched by license number to an inspection, but to account for licence discrepancies we also matched based on venue (street address and name)
# * Due to formatting differences it was necessary to use only the street number
# +
# Business licenses have numbers on end preventing simple match
# so using street number instead
def get_street_number(address):
return address.split()[0]
licenses["street_number"] = licenses.address.apply(get_street_number)
inspections["street_number"] = inspections.address.apply(get_street_number)
# Match based on DBA name and street number
venue_matches = pd.merge(inspections, licenses, left_on=["dba_name", "street_number"], right_on=["doing_business_as_name", "street_number"])
# Match based on license numbers
licence_matches = pd.merge(inspections, licenses, left_on="license", right_on="license_number")
# -
# * to create the working matches dataset we then appended venue and licence matches and dropped any duplicate inspection / business licence matches.
# +
# Join matches, reset index, drop duplicates
matches = venue_matches.append(license_matches, sort=False)
matches.reset_index(drop=True, inplace=True)
matches.drop_duplicates(["inspection_id", "id"], inplace=True)
# Restrict to matches where inspection falls within license period
matches = matches.loc[matches.inspection_date.between(matches.license_start_date, matches.expiration_date)]
# -
# ### 4.2. Filterering by Category
# * (This isn't a feature but is only convenient to do once we have the matches dataset. what to do?)
# * many non-retail establishments eg schools, hospitals follow different inspection schedules, so to ensure consistent data we filter matches to include only inspections of retail food establishments
# * to do this we select the inspection id's of all retail matches, drop any duplicates and merge these id's with the model data
# * by default merge includes only rows with keys present in each dataset (inner join)
# +
# Select retail food establishment inspection IDs
retail = matches.loc[matches.license_description == "Retail Food Establishment", ["inspection_id"]]
retail.drop_duplicates(inplace=True)
# FILTER: ONLY CONSIDER INSPECTIONS MATCHED WITH RETAIL LICENSES
data = pd.merge(data, retail, on="inspection_id")
# -
# ### 4.3. Calculating Age at Inspection
# * What might age at inspection tell?
# * One feature previously found significant in predicting inspection outcomes is the age of the facility
# * To calculate this we first convert all dates to datetime objects
# * We then group by licence and within each group find the earliest license start date
# * Finally we subtract this min date from the inspection date and merge the resulting age in with our model data
# +
# Convert dates to datetime format
matches.inspection_date = pd.to_datetime(matches.inspection_date)
matches.license_start_date = pd.to_datetime(matches.license_start_date)
def get_age_data(group):
min_date = group.license_start_date.min()
deltas = group.inspection_date - min_date
group["age_at_inspection"] = deltas.apply(lambda x: x.days / 365.25)
return group[["inspection_id", "age_at_inspection"]]
# Calculate (3 mins), drop duplicates
age_data = matches.groupby("license").apply(get_age_data).drop_duplicates()
# Merge in age_at_inspection
data = pd.merge(data, age_data, on="inspection_id", how="left")
# -
# ### 4.4. Calculating Category Data
# * The chicago team found the categories of licences attributed to an establishment to be significant in predicting violation outcomes
# * This data is derived from the licence_description column of the business licences dataset
# * We will be noting the presence or absence of these categories as a series of binary flags
# * To derive these features we first set up a dictionary linking the column entries to our desired snake case column titles
# * We then group matches by inspection id to gather all licence descriptions for each inspection
# * To generate the entries we apply our get_category_data method, using our dictionary to translate from licence_description entries to column titles
# * Finally we fill missing entries as 0 and merge the results in with our model data
# +
# Translate categories to snake-case titles
categories = {
"Consumption on Premises - Incidental Activity": "consumption_on_premises_incidental_activity",
"Tobacco": "tobacco",
"Package Goods": "package_goods",
"Limited Business License": "limited_business_license",
"Outdoor Patio": "outdoor_patio",
"Public Place of Amusement": "public_place_of_amusement",
"Children's Services Facility License": "childrens_services_facility_license",
"Tavern": "tavern",
"Regulated Business License": "regulated_business_license",
"Filling Station": "filling_station",
"Caterer's Liquor License": "caterers_liquor_license",
"Mobile Food License": "mobile_food_license"
}
# Create binary markers for license categories
def get_category_data(group):
df = group[["inspection_id"]].iloc[[0]]
for category in group.license_description:
if category in categories:
df[categories[category]] = 1
return df
# group by inspection, get categories (2 mins)
category_data = matches.groupby("inspection_id").apply(get_category_data)
# Reset index, set absent categories to 0
category_data.reset_index(drop=True, inplace=True)
category_data.fillna(0, inplace=True)
# Merge in category data, fill nan with 0
data = pd.merge(data, category_data, on="inspection_id", how="left").fillna(0)
# -
# ## 5. Crime Density
# Load observation datasets
burglaries = pd.read_csv(os.path.join(root_path, "DATA/burglaries.csv"))
carts = pd.read_csv(os.path.join(root_path, "DATA/garbage_carts.csv"))
complaints = pd.read_csv(os.path.join(root_path, "DATA/sanitation_complaints.csv"))
# Create datetime columns
inspections["datetime"] = pd.to_datetime(inspections.inspection_date)
burglaries["datetime"] = pd.to_datetime(burglaries.date)
carts["datetime"] = pd.to_datetime(carts.creation_date)
complaints["datetime"] = pd.to_datetime(complaints.creation_date)
# FILTER: consider only inspections since 2012
# Otherwise early inspections have few/no observations within window
inspections = inspections.loc[inspections.inspection_date >= "2012"]
# +
from datetime import datetime, timedelta
from scipy import stats
def get_kde(observations, column_name, window, bandwidth):
# Sort chronologically and index by datetime
observations.sort_values("datetime", inplace=True)
observations.index = observations.datetime.values
# Generate kernel from 90 days of observations
def get_kde_given_date(group):
stop = group.datetime.iloc[0]
start = stop - timedelta(days=window)
recent = observations.loc[start:stop]
x1 = recent.longitude
y1 = recent.latitude
values = np.vstack([x1, y1])
kernel = stats.gaussian_kde(values)
x2 = group.longitude
y2 = group.latitude
samples = np.vstack([x2, y2])
group[column_name] = kernel(samples)
return group[["inspection_id", column_name]]
# Group inspections by date, generate kernels, sample
return inspections.groupby("inspection_date").apply(get_kde_given_date)
# +
# Calculate burglary density estimates
burglary_kde = get_kde(burglaries, "burglary_kde", 90, 1)
# Calculate garbage cart density estimates
cart_kde = get_kde(carts, "cart_kde", 90, 1)
# Calculate sanitation complaint density estimates
complaint_kde = get_kde(complaints, "complaint_kde", 90, 1)
# -
# FILTER: only consider data since 2012 (with good kde data)
data = pd.merge(data, burglary_kde, on="inspection_id")
data = pd.merge(data, cart_kde, on="inspection_id")
data = pd.merge(data, complaint_kde, on="inspection_id")
# ## 6. Garbage Cart Density
# * Why we're including this feature
# * With our kernel density methods already defined...
# ## 7. Sanitation Complaint Density
# * Why we're including this feature
# * As with crime and garbage carts...
# ## 8. Weather Features
# +
# Load weather data
weather = pd.read_csv(os.path.join(root_path, "DATA/weather.csv"))
# Merge weather data with model data
data = pd.merge(data, weather, on="inspection_id")
# -
# ## 9. Next Steps
# * <NAME> is a web application developer with a great fondness for data driven decision making. Russell is excited to explore the applications of data science and machine learning in improving human judgement.
# * <NAME> is a seasoned corporate responsibility professional working to utilize technology to help improve the health and well being of human populations through environmental stewardship.
# * <NAME>, Ph.D. is the managing partner at Sustainabilist and an expert in process improvement for distributed systems. Jason’s work portfolio includes the creation of novel data-driven methods for improving contractor performance, machine learning to optimize value in energy efficiency sales, and equipment maintenance optimization methodologies.
| PUBLICATIONS/publication_2/.ipynb_checkpoints/publication_2_draft_1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import pandas as pd
from IPython.display import display, HTML
import numpy as np
import matplotlib.pyplot as plt
import sklearn
from sklearn.model_selection import train_test_split
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
import statsmodels.api as sm
import seaborn as sb
from pylab import rcParams
DATABASE_PATH = '../data/simple_time_based_model.zip'
# Allow us to display all the columns in a dataframe
pd.options.display.max_columns = None
# -
df_time = pd.read_csv('simple_time_based_model.zip', compression='zip', header=0, sep=',', quotechar='"', low_memory=False, index_col=False)
# Calculate the proportion of positive examples
len(df_time[df_time['HEALTH_BASED_VIOLATION_THIS_YEAR'] == 1])/len(df_time)
df_time['EPA_REGION'].value_counts()
df_time.head(20)
df_time.set_index('PWSID', inplace=True)
df_time.head()
copy_of_initial_data = df_time.to_csv("Copy_of_Initial_Data.csv", encoding='utf-8')
df_time['PWSID'].value_counts
len(df_time)
df_time.info()
df_time.shape
df_codes = pd.read_csv('contaminant-codes.csv', header=0, sep=',', quotechar='"', low_memory=False, index_col=False)
df_group_codes = pd.read_csv('contaminant-group-codes.csv', header=0, sep=',', quotechar='"', low_memory=False, index_col=False)
df_violations = pd.read_csv('violations.csv', header=0, sep=',', quotechar='"', low_memory=False, index_col=False)
df_codes.head()
df_group_codes.head()
df_violations.head()
df_time.count()
df_time.shape
df_time.duplicated(subset=None, keep='first')
df_time.drop(['PRIMACY_TYPE', 'PRIMARY_SOURCE_CODE',
'PWS_ACTIVITY_CODE', 'PWSID', 'PWS_TYPE_CODE',
'SERVICE_CONNECTIONS_COUNT', 'ZIP_CODE'],
axis='columns', inplace=True)
df_time.count()
df_time = df_time.dropna(axis='index')
df_time['EPA_REGION'] = df_time['EPA_REGION'].astype(int)
df_time['STATE_CODE'] = df_time['STATE_CODE'].astype("category")
df_time['EPA_REGION'].value_counts()
df_time.reset_index(inplace=True)
df_time.drop('index', axis=1, inplace=True)
df_time.head()
one_hot_copy = df_time.copy()
one_hot_copy.head()
# ### Convert the Year column to datetime objects:
# +
# df = pd.read_csv('test.csv')
# df['date'] = pd.to_datetime(df['date'])
# df['date_delta'] = (df['date'] - df['date'].min()) / np.timedelta64(1,'D')
# city_data = df[df['city'] == 'London']
# result = sm.ols(formula = 'sales ~ date_delta', data = city_data).fit()
# -
type(one_hot_copy['YEAR'][0])
one_hot_copy['YEAR'] = one_hot_copy['YEAR'].astype(int)
one_hot_copy['YEAR'] = pd.to_datetime(one_hot_copy['YEAR'], format='%Y')
one_hot_copy['YEAR'].head(10)
one_hot_copy.head()
df_time['EPA_REGION'].value_counts()
# ### Here we're fixing the format and type of the year column so that we'll be able to run the logit regression later
# - Although the new year column is named the same as the old one, it's a separate column of ints rather than datetime objects (pandas is case sensitive so it can tell the difference)
# - I kept the name the same because it's easy to type 'Year'
def year_maker(date_value):
return date_value.year
one_hot_copy['Year'] = one_hot_copy['YEAR'].apply(year_maker)
one_hot_copy.head()
# ### Now just reorder the columns for visual ease:
cols_at_start = ['POPULATION_SERVED_COUNT', 'Year']
one_hot_copy = one_hot_copy[[c for c in cols_at_start if c in one_hot_copy] +
[c for c in one_hot_copy if c not in cols_at_start]]
one_hot_copy.head()
# ### We don't need the original "Year" column anymore so drop it:
one_hot_copy.drop('YEAR', axis=1, inplace=True)
# ### Check what different year values are in the new 'Year' column:
one_hot_copy.Year.unique()
one_hot_copy.head()
# ### We're just going to rename the columns here for visual ease:
one_hot_copy.rename(columns = {'POPULATION_SERVED_COUNT': 'Population',
'HEALTH_BASED_VIOLATION_THIS_YEAR': "Health_Violations_Year_0",
'NUM_HEALTH_BASED_VIOLATIONS_PREVIOUS_YEAR': 'Health_Violations_Year_1',
'NUM_NON_HEALTH_BASED_VIOLATIONS_PREVIOUS_YEAR': 'Other_Violations_Year_1',
'NUM_HEALTH_BASED_VIOLATIONS_2_YEARS_AGO': 'Health_Violations_Year_2',
'NUM_NON_HEALTH_BASED_VIOLATIONS_2_YEARS_AGO': 'Other_Violations_Year_2',
'NUM_HEALTH_BASED_VIOLATIONS_3_YEARS_AGO': 'Health_Violations_Year_3',
'NUM_NON_HEALTH_BASED_VIOLATIONS_3_YEARS_AGO': 'Other_Violations_Year_3',
'NUM_HEALTH_BASED_VIOLATIONS_4_YEARS_AGO': 'Health_Violations_Year_4',
'NUM_NON_HEALTH_BASED_VIOLATIONS_4_YEARS_AGO': 'Other_Violations_Year_4',
'NUM_HEALTH_BASED_VIOLATIONS_5_YEARS_AGO': 'Health_Violations_Year_5',
'NUM_NON_HEALTH_BASED_VIOLATIONS_5_YEARS_AGO': 'Other_Violations_Year_5',
'NUM_ENFORCEMENTS_PREVIOUS_YEAR': 'Enforcements_Year_1',
'NUM_ENFORCEMENTS_2_YEARS_AGO': 'Enforcements_Year_2',
'NUM_ENFORCEMENTS_3_YEARS_AGO': 'Enforcements_Year_3',
'NUM_ENFORCEMENTS_4_YEARS_AGO': 'Enforcements_Year_4',
'NUM_ENFORCEMENTS_5_YEARS_AGO': 'Enforcements_Year_5',
'EPA_REGION': 'Epa_Region',
'STATE_CODE': 'State'}, inplace=True)
one_hot_copy.head()
df_time['EPA_REGION'].value_counts()
# ## As of right now we're just working with the simple_time_based_model data
# ### Check to make sure the state codes are exclusive:
df_one_hot = pd.get_dummies(one_hot_copy, columns=['Epa_Region', 'State'], drop_first=True)
df_one_hot.count()
df_one_hot.head()
# - 0 if: none of the values on the given row were equal to
# This means that the reason of absence was not in this group
# 1 if: somewhere among these 14 columns we have observed the number 1
# - This means that the reason of absence was in this group
state_dummies = df_one_hot[df_one_hot.columns[df_one_hot.columns.get_loc('State_AL') :]]
# ### Below is the check the sum of the state dummies per row. Should equal 1 (if there was a violation) or 0 (if there wasn't)
df_one_hot['state_check'] = state_dummies.sum(axis=1)
df_one_hot.head()
df_one_hot.dtypes
# ### The below code is to check that the dummy variables where created succussfully. Should be an array of 1, 0:
df_one_hot.state_check.unique()
# ### Create a checkpoint of the time based data DF:
df_update1 = df_one_hot.copy()
df_update1.head()
# ### Convert violation data into integers rather than floats:
df_update1.dtypes
df_update1[df_update1.columns[
df_update1.columns.get_loc('Health_Violations_Year_0'): df_update1.columns.get_loc('Epa_Region_2')]] = df_update1[df_update1.columns[df_update1.columns.get_loc('Health_Violations_Year_0'): df_update1.columns.get_loc('Epa_Region_2')]].astype(int)
df_update1.dtypes
# ## Checking that the target variable is binary:
# Since we are building a model to predict whether or not there was a violation, our target is going to be "HEALTH_BASED_VIOLATION_THIS_YEAR" variable from the original simple_time_based_model dataframe. To make sure that it's a binary variable, let's use Seaborn's countplot() function.
# %matplotlib inline
rcParams['figure.figsize'] = 10, 8
sb.set_style('whitegrid')
sb.countplot(x='Health_Violations_Year_0',data=df_update1)
df_update1['Health_Violations_Year_0'].value_counts()
df_copy = df_update1.copy().drop(['state_check'], axis=1)
df_copy.dtypes
df_copy['Health_Violations_Year_0'].value_counts(normalize=True)[0]* 100,
df_copy['Health_Violations_Year_1'].value_counts(normalize=True)[0]* 100,
df_copy['Health_Violations_Year_2'].value_counts(normalize=True)[0]* 100,
df_copy['Health_Violations_Year_3'].value_counts(normalize=True)[0]* 100,
df_copy['Health_Violations_Year_4'].value_counts(normalize=True)[0]* 100,
df_copy['Health_Violations_Year_5'].value_counts(normalize=True)[0]* 100
# ## From the above, we can see that the the majority of the data is negative values (0) when it comes to negative cases when it comes to Health Violations across the years
# ### So we're going to perform a logistic regression before sampling the data sets (as done in the original 'Logit_Crappy_Model' but more accurately (I think)
# ### Then we're going to take the sampels and perform the logisitic regression again.
# ### So continuing on:
# ## Omitting the dummy variables from the Standardization:
# ### Big Issue: When we standardize the inputs with the above scaler object, we would also standardize the dummies (BAD PRACTICE)
# - When dummies are standardized they lose their interpretability
#
# - A unit change with dummy variables means:
# - Going from disregarding this dummy to taking ONLY this dummy into account
# - However, we previously standardized all the features (including the dummies) and now a unit change is uninterpretable.
#
# NOTE: We're could just perform the standardization before making the dummies, but I'm just trying to keep my coding skills... Disregard
# +
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.preprocessing import StandardScaler
class CustomScaler(BaseEstimator, TransformerMixin):
def __init__(self, columns, copy=True, with_mean=True, with_std=True):
self.scaler = StandardScaler(copy, with_mean, with_std)
self.columns = columns
self.mean_ = None
self.var_ = None
def fit(self, X, y=None):
self.scaler.fit(X[self.columns], y)
self.mean_ = np.mean(X[self.columns])
self.var_ = np.var(X[self.columns])
return self
def transform(self, X, y=None, copy=None):
init_col_order = X.columns
X_scaled = pd.DataFrame(self.scaler.transform(X[self.columns]),
columns = self.columns)
X_not_scaled = X.loc[:, ~X.columns.isin(self.columns)]
return pd.concat([X_not_scaled, X_scaled], axis=1)[init_col_order]
# -
df_copy.columns.values
columns_to_omit = ['Health_Violations_Year_0', 'Epa_Region_2', 'Epa_Region_3',
'Epa_Region_4', 'Epa_Region_5', 'Epa_Region_6', 'Epa_Region_7',
'Epa_Region_8', 'Epa_Region_9', 'Epa_Region_10', 'State_AL',
'State_AR', 'State_AS', 'State_AZ', 'State_BC', 'State_CA',
'State_CO', 'State_CT', 'State_DC', 'State_DE', 'State_FL',
'State_GA', 'State_GU', 'State_HI', 'State_IA', 'State_ID',
'State_IL', 'State_IN', 'State_KS', 'State_KY', 'State_LA',
'State_MA', 'State_MD', 'State_ME', 'State_MI', 'State_MN',
'State_MO', 'State_MP', 'State_MS', 'State_MT', 'State_NB',
'State_NC', 'State_ND', 'State_NE', 'State_NH', 'State_NJ',
'State_NM', 'State_NS', 'State_NV', 'State_NY', 'State_OH',
'State_OK', 'State_ON', 'State_OR', 'State_PA', 'State_PQ',
'State_PR', 'State_RI', 'State_SC', 'State_SD', 'State_TN',
'State_TX', 'State_UT', 'State_VA', 'State_VI', 'State_VT',
'State_WA', 'State_WI', 'State_WV', 'State_WY']
columns_to_scale = [x for x in df_copy.columns.values if x not in columns_to_omit]
columns_to_scale
# ### Still fixing the scaler:
# ### Now create the fixed scaler object:
data_scaler = CustomScaler(columns_to_scale)
# ### Now the scaler is fixed and we can continue on:
# - Below line will calculate and store the mean and standard deviation of each feature/variable from unscaled_inputs
# - Stored in data_scaler object
# - Whenever you get new data the standardization info is contained in data_scaler
data_scaler.fit(df_copy)
# ### We have just prepared the 'scaling mechanism'
# #### In order to apply it, we must call transform()
# - Transforms the unscaled inputs using the info contained in absentee_scaler
# - This subtracts the mean and divide by the standard deviation
scaled_inputs = data_scaler.transform(df_copy)
# ### Whenever you get new data, you will just apply below to reach same transformation as above (most common and useful way to transform new data when deploying a model):
#
# new_data_raw = pd.read_csv('new_data.csv')
#
# new_data_scaled = absentee_scaler.transform(new_data_raw)
# Below we can see that all the dummies remain untouched (0's and 1's):
scaled_inputs.head(20)
scaled_inputs.shape
# Split the data into independent and dependent variables. Also split into train and test sets.
Y = scaled_inputs['Health_Violations_Year_0'].copy()
Y is scaled_inputs['Health_Violations_Year_0']
X = scaled_inputs.drop(['Health_Violations_Year_0'], axis=1)
X.head()
# ### Checking for missing values:
scaled_inputs.isnull().sum().unique()
# ## Train/Test Split of Data:
# ### Overfitting:
# - Model learns the predict the train data so well that it fails on new data
# - So split the data into Train and Test to access the model's accuracy on predicting new data
# ### Data Shuffling:
# - In order to remove all types of dependencies that come from the order of the data set
#
x_train, x_test, y_train, y_test = train_test_split(X, Y,
test_size=0.2, random_state=42,
stratify=Y)
# ### Splitting the data and then training it:
display(x_train.shape)
display(x_test.shape)
display(len(x_test) / (len(x_test) + len(x_train)))
display(y_train.shape)
display(y_test.shape)
display(y_train)
display(len(y_train[y_train==1])/len(y_train))
display(len(y_test[y_test==1])/len(y_test))
# # Training the model and Assessing its Accuracy:
# ## Logistic Regression with sklearn:
from sklearn.linear_model import LogisticRegression
from sklearn import metrics
reg = LogisticRegression()
reg.fit(x_train, y_train)
reg.score(x_train, y_train)
# ## As we can see, even when run properly, we have a ridiclously high accuracy rate. The model we ran isn't THAT good. However this is probably because ~98% of the data set is all 0's, and this caused the model to just predict all those 0's rather than the 1's that we're looking for.
# ## We're going to MANUALLY check the accuracy to make sure that the model was correct:
# #### (Build score method above from scratch):
# - Accuracy:
# - Logistic Regression trained on train inputs
# - Finds outputs that are trying to be as close to the targets are possible
# - In this case 98.5% of the model ouputs match the targets
# - So to find accuracy of model manually, find the model outputs and compare them with the targets:
# - Find model ouputs with .predict()
# Model itself contained in variable 'reg'
model_outputs = reg.predict(x_train)
model_outputs[:100]
# Let's look at the targets to compare. Look similar but have differences
y_train.head(100)
# See which elements have been guessed correctly and whihc haven't
model_outputs == y_train
# #### How many matches are there?:
np.sum(model_outputs == y_train)
model_outputs.shape[0]
# #### Finally, check/verify the accuracy:
np.sum(model_outputs == y_train) / model_outputs.shape[0]
# # As of now, the logistic regression is working, but not accurately because we're training it on 98% negative cases. So we're going to improve it and work with the samples.
# ## We're going to try to run an "Informed Over Sample" which is a synthetic minority over-sampling technique.
# ### "This technique is followed to avoid overfitting which occurs when exact replicas of minority instances are added to the main dataset. A subset of data is taken from the minority class as an example and then new synthetic similar instances are created. These synthetic instances are then added to the original dataset. The new dataset is used as a sample to train the classification models."
# ### Let's see how many of the violation values are contributing to this massive 98% 0 value:
df_copy2 = df_update1.copy().drop(['state_check'], axis=1)
test1 = df_copy2[(df_copy2['Health_Violations_Year_0'] == 0) &
(df_copy2['Health_Violations_Year_1'] == 0) &
(df_copy2['Health_Violations_Year_2'] == 0) &
(df_copy2['Health_Violations_Year_3'] == 0) &
(df_copy2['Health_Violations_Year_4'] == 0) &
(df_copy2['Health_Violations_Year_5'] == 0) &
(df_copy2['Other_Violations_Year_1'] == 0) &
(df_copy2['Other_Violations_Year_2'] == 0) &
(df_copy2['Other_Violations_Year_3'] == 0) &
(df_copy2['Other_Violations_Year_4'] == 0) &
(df_copy2['Other_Violations_Year_5'] == 0)].copy()
test1 is df_copy2
# ### Let's confirm that test1 is giving us what we want. There should only be one value (0) for each violation column:
# +
# set
# -
# ### It worked properly, now let's see how many values there are:
len(test1)
# ### That's nearly half of the original DF! We don't need these immediately, at least not within our training data! Maybe we should include this within our clustering later on when we merge the other csv's from the SQL tables together. But not right now. So let's get rid of these for our analysis. But let's check how many target based 0's are left:
# len(df_copy[(df_copy['Health_Violations_Year_0'] == 0)]) - len(test1)
len(df_copy2) - len(df_copy2[df_copy2['Health_Violations_Year_0'] == 0])
# ### As we can see, we only have about 33,000 targets that have a value of 1. Yikes. So for our training data, let's not include our test1 data and just use about 33,000 random 0 values from the remaining target column of df_copy[df_copy['Health_Violations_Year_0'] == 0] without the test1 values:
# - This is the value of the size of the original dataset, which would split the target values of 0, 1 in half.
# - We'll choose at random to try to minimize any bias and reduce any patterns
len(df_copy2) - len(test1)
len(df_copy2[df_copy2['Health_Violations_Year_0'] == 0]) - len(test1)
# ### So there are about 400,000 values that are 0 remaining in the target column. Let's random sample about 33,000 of them for our training data:
df_less_test1 = df_copy2[~df_copy2.isin(test1)].dropna(how = 'all').copy()
# ### TODO: There may be excessive duplicates in this dataset. Search for duplicates later:
# +
# test7 = df_copy.drop_duplicates(subset=['Population', 'Year', 'Health_Violations_Year_0',
# 'Health_Violations_Year_1', 'Other_Violations_Year_1',
# 'Health_Violations_Year_2', 'Other_Violations_Year_2',
# 'Health_Violations_Year_3', 'Other_Violations_Year_3',
# 'Health_Violations_Year_4', 'Other_Violations_Year_4',
# 'Health_Violations_Year_5', 'Other_Violations_Year_5',
# 'Enforcements_Year_1', 'Enforcements_Year_2',
# 'Enforcements_Year_3', 'Enforcements_Year_4',
# 'Enforcements_Year_5', 'Epa_Region_2', 'Epa_Region_3',
# 'Epa_Region_4', 'Epa_Region_5', 'Epa_Region_6', 'Epa_Region_7',
# 'Epa_Region_8', 'Epa_Region_9', 'Epa_Region_10', 'State_AL',
# 'State_AR', 'State_AS', 'State_AZ', 'State_BC', 'State_CA',
# 'State_CO', 'State_CT', 'State_DC', 'State_DE', 'State_FL',
# 'State_GA', 'State_GU', 'State_HI', 'State_IA', 'State_ID',
# 'State_IL', 'State_IN', 'State_KS', 'State_KY', 'State_LA',
# 'State_MA', 'State_MD', 'State_ME', 'State_MI', 'State_MN',
# 'State_MO', 'State_MP', 'State_MS', 'State_MT', 'State_NB',
# 'State_NC', 'State_ND', 'State_NE', 'State_NH', 'State_NJ',
# 'State_NM', 'State_NS', 'State_NV', 'State_NY', 'State_OH',
# 'State_OK', 'State_ON', 'State_OR', 'State_PA', 'State_PQ',
# 'State_PR', 'State_RI', 'State_SC', 'State_SD', 'State_TN',
# 'State_TX', 'State_UT', 'State_VA', 'State_VI', 'State_VT',
# 'State_WA', 'State_WI', 'State_WV', 'State_WY'], keep=False)
# -
len(df_less_test1)
df_less_test1['Health_Violations_Year_0'].value_counts()
# ### Now we're getting the subsets that we want, however, the isin() method converted the int64's into something weird. They're still int's but they're added with a period (0 goes to 0.0). This format should output a float, but it's not, and it's going to messup the scaler later on. So let's try another method:
df_less_test1[df_less_test1.columns[
df_less_test1.columns.get_loc('Health_Violations_Year_0')]] = df_less_test1[df_less_test1.columns[df_less_test1.columns.get_loc('Health_Violations_Year_0')]].astype(int)
df_less_test1[df_less_test1.columns[
df_less_test1.columns.get_loc('Epa_Region_2'):]] = df_less_test1[df_less_test1.columns[df_less_test1.columns.get_loc('Epa_Region_2'):]].astype(int)
df_less_test1.head()
# ### Ok, that type conversion is complete. Let's carry on:
df_majority = df_less_test1[df_less_test1['Health_Violations_Year_0']==0]
df_minority = df_less_test1[df_less_test1['Health_Violations_Year_0']==1]
df_maj_reduced = df_majority.sample(n=34000, # To match about equal to minority class
replace=False)
df_downsampled = pd.concat([df_maj_reduced, df_minority])
len(df_downsampled)
df_downsampled['Health_Violations_Year_0'].unique()
df_downsampled.dtypes
df_downsampled.head()
df_downsampled['Health_Violations_Year_0'].value_counts(normalize=True)[0]* 100
df_downsampled['Health_Violations_Year_1'].value_counts(normalize=True)[0]* 100
df_downsampled['Health_Violations_Year_1'].value_counts(normalize=True)[0]* 100
df_downsampled['Health_Violations_Year_2'].value_counts(normalize=True)[0]* 100
df_downsampled['Health_Violations_Year_3'].value_counts(normalize=True)[0]* 100
df_downsampled['Health_Violations_Year_4'].value_counts(normalize=True)[0]* 100
df_downsampled['Health_Violations_Year_5'].value_counts(normalize=True)[0]* 100
df_downsampled.columns.values
unscaled_sample = df_downsampled.copy()
# +
from sklearn.preprocessing import RobustScaler
sample_scaler = StandardScaler()
# -
unscaled_sample[['Population', 'Year','Health_Violations_Year_1', 'Other_Violations_Year_1',
'Health_Violations_Year_2', 'Other_Violations_Year_2',
'Health_Violations_Year_3', 'Other_Violations_Year_3',
'Health_Violations_Year_4', 'Other_Violations_Year_4',
'Health_Violations_Year_5', 'Other_Violations_Year_5',
'Enforcements_Year_1', 'Enforcements_Year_2',
'Enforcements_Year_3', 'Enforcements_Year_4',
'Enforcements_Year_5']] = sample_scaler.fit_transform(unscaled_sample[['Population', 'Year','Health_Violations_Year_1', 'Other_Violations_Year_1',
'Health_Violations_Year_2', 'Other_Violations_Year_2',
'Health_Violations_Year_3', 'Other_Violations_Year_3',
'Health_Violations_Year_4', 'Other_Violations_Year_4',
'Health_Violations_Year_5', 'Other_Violations_Year_5',
'Enforcements_Year_1', 'Enforcements_Year_2',
'Enforcements_Year_3', 'Enforcements_Year_4',
'Enforcements_Year_5']])
unscaled_sample.head()
# ## We have scaled the sample correctly. Time to create another checkpoint from a copy:
scaled_sample = unscaled_sample.copy()
scaled_sample.shape
# Split the data into independent and dependent variables. Also split into train and test sets.
sample_Y = scaled_sample['Health_Violations_Year_0'].copy()
sample_Y is scaled_sample['Health_Violations_Year_0']
sample_X = scaled_sample.drop(['Health_Violations_Year_0'], axis=1)
sample_X.head()
# ### Checking for missing values:
scaled_sample.isnull().sum().unique()
# # Train/Test Split of Data:
# ### Overfitting:
# - Model learns the predict the train data so well that it fails on new data
# - So split the data into Train and Test to access the model's accuracy on predicting new data
# ### Data Shuffling:
# - In order to remove all types of dependencies that come from the order of the data set
#
# ### Splitting the data and then training it:
# +
# display(sample_x_train.shape)
# display(sample_x_test.shape)
# display(len(sample_x_test) / (len(sample_x_test) + len(sample_x_train)))
# display(sample_y_train.shape)
# display(sample_y_test.shape)
# display(sample_y_train)
# display(len(sample_y_train[sample_y_train==1])/len(sample_y_train))
# display(len(sample_y_test[sample_y_test==1])/len(sample_y_test))
# -
# # Training the model and Assessing its Accuracy:
# ## Logistic Regression with sklearn:
# ### StandardScaler:
from sklearn.linear_model import LogisticRegression
from sklearn import metrics
sample_reg = LogisticRegression()
sample_reg.fit(sample_x_train, sample_y_train)
sample_reg.score(sample_x_train, sample_y_train)
sample_reg.score(sample_x_test, sample_y_test)
# ## As we can see, even when run properly, we have a ridiclously high accuracy rate. The model we ran isn't THAT good. However this is probably because ~98% of the data set is all 0's, and this caused the model to just predict all those 0's rather than the 1's that we're looking for.
# ## We're going to MANUALLY check the accuracy to make sure that the model was correct:
# #### (Build score method above from scratch):
# - Accuracy:
# - Logistic Regression trained on train inputs
# - Finds outputs that are trying to be as close to the targets are possible
# - In this case 98.5% of the model ouputs match the targets
# - So to find accuracy of model manually, find the model outputs and compare them with the targets:
# - Find model ouputs with .predict()
# Model itself contained in variable 'reg'
model_outputs = reg.predict(x_train)
model_outputs[:100]
# Let's look at the targets to compare. Look similar but have differences
y_train.head(100)
# See which elements have been guessed correctly and whihc haven't
model_outputs == y_train
# #### How many matches are there?:
np.sum(model_outputs == y_train)
model_outputs.shape[0]
# #### Finally, check/verify the accuracy:
np.sum(model_outputs == y_train) / model_outputs.shape[0]
# # Try a Random Forest Classifier:
# ## These are much more accurate when it comes to imbalanced datasets, so we shouldn't have to give it the balanced DF, but let's just be safe at first:
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
from sklearn.metrics import roc_auc_score
# +
forest_Y = df_downsampled['Health_Violations_Year_0']
forest_X = df_downsampled.drop('Health_Violations_Year_0', axis=1)
class_forest = RandomForestClassifier()
class_forest.fit(forest_X, forest_Y)
# Predict on training set
pred_forest_Y = class_forest.predict(forest_X)
# Is our model predicting just one class? Should be an array of 0 and 1
print(np.unique(pred_forest_Y))
# How's our accuracy?
print(accuracy_score(forest_Y, pred_forest_Y) )
# What about AUROC?
prob_y_4 = class_forest.predict_proba(forest_X)
prob_y_4 = [p[1] for p in prob_y_4]
print( roc_auc_score(forest_Y, prob_y_4) )
# -
# ## Now let's try with the imbalanced dataset:
# +
forest_Y = df_copy2['Health_Violations_Year_0']
forest_X = df_copy2.drop('Health_Violations_Year_0', axis=1)
class_forest = RandomForestClassifier()
class_forest.fit(forest_X, forest_Y)
# Predict on training set
pred_forest_Y = class_forest.predict(forest_X)
# Is our model predicting just one class? Should be an array of 0 and 1
print(np.unique(pred_forest_Y))
# How's our accuracy?
print(accuracy_score(forest_Y, pred_forest_Y) )
# What about AUROC?
prob_y_4 = class_forest.predict_proba(forest_X)
prob_y_4 = [p[1] for p in prob_y_4]
print( roc_auc_score(forest_Y, prob_y_4) )
# -
X.describe()
# ## Extracting the Intercept and Coefficients from the Logistic Regression:
reg.intercept_
reg.coef_
# ### However we want to know which variable these coefficients refer to:
scaled_inputs.columns.values
# # Below we can do after logistic regression to check for sources of error or inaccuracy:
# ## Now, I have a feeling the number of people in a given region will have a factor in the output. So let's look at a comparison of how population is related to their health based violations in previous years:
df_update1['Population'].max()
# ### We're going to set a limit on the y-axis since the majority of the violations = 0, but we don't care about these in this case:
plot1 = sb.boxplot(x='Health_Violations_Year_1', y='Population', data=df_update1, palette='hls')
plot1.set(ylim=(0,80000))
plot1 = sb.boxplot(x='Health_Violations_Year_2', y='Population', data=df_update1, palette='hls')
plot1.set(ylim=(0,80000))
plot1 = sb.boxplot(x='Health_Violations_Year_3', y='Population', data=df_update1, palette='hls')
plot1.set(ylim=(0,80000))
# ### As expected, regions that serve smaller populations (10,000 people or less) experience the most number of health based violations as the lower 50% of violations occur the most with populations around even 5,000 people
df_update1.sort_values(['Year', 'Population', 'Health_Violations_Year_0'], ascending=False, inplace=True)
df_update1.head()
df_update2 = df_update1.copy()
df_update2.reset_index(drop=True, inplace=True)
df_update2.head()
df_update2['Health_Violations_Year_1'].value_counts()
df_update2['Health_Violations_Year_1'].value_counts()
sb.countplot(x='Health_Violations_Year_2', data=df_update2, palette='hls')
# ## Checking for independence between features:
# +
# sb.heatmap(one_hot_copy.corr(), robust=True)
# -
sb.heatmap(df_update2[0:.corr(), cmap='Set1')
# ### Convert Date's into dummies:
hot_copy_copy = one_hot_copy.copy()
import datetime as dt
hot_copy_copy['YEAR']= hot_copy_copy['YEAR'].map(dt.datetime.toordinal)
hot_copy_copy.YEAR.unique()
# +
unscaled_inputs = data_with_targets.iloc[:, :-1]
| projects/exploratory_phase/code/notebooks/Updated_Logit_Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Base enem 2016
# ## Predição se o aluno é treineiro.
#
# ## Primeiro teste:
#
# ### * Somente a limpeza dos dados
# ### * Sem balanceamento
# ### * Regressão Logística
#
# Score obtido: 87.921225
# +
import pandas as pd
import numpy as np
import warnings
from sklearn.preprocessing import OneHotEncoder
from sklearn.linear_model import LogisticRegression
warnings.filterwarnings('ignore')
pd.set_option('display.max_columns', 300)
pd.set_option('display.max_rows', 300)
# +
file_train = "train.csv"
file_test = "test.csv"
df_raw_train = pd.read_csv(file_train, index_col=False)
df_raw_test = pd.read_csv(file_test, index_col=False)
df_raw_train.shape, df_raw_test.shape
# +
columns_used=['NU_INSCRICAO','CO_UF_RESIDENCIA', 'SG_UF_RESIDENCIA', 'NU_IDADE', 'TP_SEXO', 'TP_COR_RACA',
'TP_NACIONALIDADE','TP_ST_CONCLUSAO','TP_ANO_CONCLUIU', 'TP_ESCOLA', 'TP_ENSINO',
'TP_PRESENCA_CN', 'TP_PRESENCA_CH', 'TP_PRESENCA_LC','NU_NOTA_CN', 'NU_NOTA_CH',
'NU_NOTA_LC','TP_LINGUA','TP_STATUS_REDACAO', 'NU_NOTA_COMP1', 'NU_NOTA_COMP2',
'NU_NOTA_COMP3','NU_NOTA_COMP4', 'NU_NOTA_COMP5', 'NU_NOTA_REDACAO','Q001', 'Q002',
'Q006', 'Q024', 'Q025', 'Q026', 'Q027', 'Q047', 'IN_TREINEIRO']
numerical_vars = ['NU_NOTA_CN', 'NU_NOTA_CH', 'NU_NOTA_LC','NU_NOTA_COMP1', 'NU_NOTA_COMP2', 'NU_NOTA_COMP3',
'NU_NOTA_COMP4','NU_NOTA_COMP5', 'NU_NOTA_REDACAO']
target = ['IN_TREINEIRO']
# +
df_train=df_raw_train[columns_used]
df_train.drop(['SG_UF_RESIDENCIA','TP_ENSINO'], inplace=True, axis=1)
df_test=df_raw_test[columns_used[:-1]]
df_test.drop(['SG_UF_RESIDENCIA','TP_ENSINO'], inplace=True, axis=1)
df_train[numerical_vars] = df_train[numerical_vars].fillna(0)
df_test[numerical_vars] = df_test[numerical_vars].fillna(0)
# -
df_train_clean = pd.DataFrame(index=df_train.index)
df_test_clean = pd.DataFrame(index=df_test.index)
df_train_clean['NU_INSCRICAO'] = df_raw_train['NU_INSCRICAO']
df_test_clean['NU_INSCRICAO'] = df_raw_test['NU_INSCRICAO']
def create_encoder(column, prefix):
#encoder = OneHotEncoder()
#train_column_df = pd.DataFrame(encoder.fit_transform(df_train[[column]]).toarray())
#test_column_df = pd.DataFrame(encoder.fit_transform(df_test[[column]]).toarray())
train_column_df = pd.get_dummies(df_train[column])
test_column_df = pd.get_dummies(df_test[column])
train_name_columns = df_train[column].sort_values().unique()
train_name_columns_co = [str(prefix) + str(train_name_column) for train_name_column in train_name_columns]
test_name_columns = df_test[column].sort_values().unique()
test_name_columns_co = [str(prefix) + str(test_name_column) for test_name_column in test_name_columns]
train_column_df.columns=train_name_columns_co
test_column_df.columns=test_name_columns_co
global df_train_clean
global df_test_clean
df_train_clean = pd.concat([df_train_clean, train_column_df ], axis=1)
df_test_clean = pd.concat([df_test_clean, test_column_df ], axis=1)
categorical_vars = {'CO_UF_RESIDENCIA' : 'co_uf_', 'TP_SEXO' : 'sexo_', 'TP_COR_RACA': 'raca_', 'TP_ST_CONCLUSAO': 'tp_st_con_',
'TP_ANO_CONCLUIU': 'tp_ano_con_', 'TP_ESCOLA': 'tp_esc_','TP_PRESENCA_CN': 'tp_pres_cn',
'TP_PRESENCA_CH': 'tp_pres_ch', 'TP_PRESENCA_LC': 'tp_pres_lc', 'TP_LINGUA': 'tp_ling_',
'Q001': 'q001_', 'Q002': 'q002_', 'Q006': 'q006_', 'Q024': 'q024_',
'Q025': 'q025_', 'Q026': 'q026_', 'Q047': 'q047_'}
for column, prefix in categorical_vars.items():
create_encoder(column, prefix)
# +
numerical_vars = ['NU_NOTA_CN', 'NU_NOTA_CH', 'NU_NOTA_LC','NU_NOTA_COMP1', 'NU_NOTA_COMP2', 'NU_NOTA_COMP3',
'NU_NOTA_COMP4','NU_NOTA_COMP5', 'NU_NOTA_REDACAO']
df_train_clean = pd.concat([df_train_clean, df_train[numerical_vars]], axis=1)
df_test_clean = pd.concat([df_test_clean, df_test[numerical_vars]], axis=1)
# -
X_train = df_train_clean.loc[:,'co_uf_11':]
y_train = df_train['IN_TREINEIRO']
X_test = df_test_clean.loc[:,'co_uf_11':]
X_train.shape, y_train.shape, X_test.shape
X_train_comp_X_test = X_train[X_test.columns]
X_train_comp_X_test.shape, y_train.shape, X_test.shape
regressor = LogisticRegression()
regressor.fit(X_train_comp_X_test, y_train)
y_pred = regressor.predict(X_test)
df_result_insc = pd.DataFrame(df_test_clean['NU_INSCRICAO'])
resultado = pd.concat([df_result_insc, pd.DataFrame(np.round(y_pred,3))], axis=1)
resultado.reset_index(inplace=True, drop=True)
resultado.columns=['NU_INSCRICAO', 'IN_TREINEIRO']
resultado.to_csv("answer.csv", index=False)
| enem-4/Week9-notasEnenChallenge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from fastai.text import *
path = Config().data_path()/'giga-fren'
path.ls()
# ## Load data
# We reuse the same functions as in the translation notebook to load our data.
def seq2seq_collate(samples:BatchSamples, pad_idx:int=1, pad_first:bool=True, backwards:bool=False) -> Tuple[LongTensor, LongTensor]:
"Function that collect samples and adds padding. Flips token order if needed"
samples = to_data(samples)
max_len_x,max_len_y = max([len(s[0]) for s in samples]),max([len(s[1]) for s in samples])
res_x = torch.zeros(len(samples), max_len_x).long() + pad_idx
res_y = torch.zeros(len(samples), max_len_y).long() + pad_idx
if backwards: pad_first = not pad_first
for i,s in enumerate(samples):
if pad_first:
res_x[i,-len(s[0]):],res_y[i,-len(s[1]):] = LongTensor(s[0]),LongTensor(s[1])
else:
res_x[i,:len(s[0]):],res_y[i,:len(s[1]):] = LongTensor(s[0]),LongTensor(s[1])
if backwards: res_x,res_y = res_x.flip(1),res_y.flip(1)
return res_x, res_y
class Seq2SeqDataBunch(TextDataBunch):
"Create a `TextDataBunch` suitable for training an RNN classifier."
@classmethod
def create(cls, train_ds, valid_ds, test_ds=None, path:PathOrStr='.', bs:int=32, val_bs:int=None, pad_idx=1,
pad_first=False, device:torch.device=None, no_check:bool=False, backwards:bool=False, **dl_kwargs) -> DataBunch:
"Function that transform the `datasets` in a `DataBunch` for classification. Passes `**dl_kwargs` on to `DataLoader()`"
datasets = cls._init_ds(train_ds, valid_ds, test_ds)
val_bs = ifnone(val_bs, bs)
collate_fn = partial(seq2seq_collate, pad_idx=pad_idx, pad_first=pad_first, backwards=backwards)
train_sampler = SortishSampler(datasets[0].x, key=lambda t: len(datasets[0][t][0].data), bs=bs//2)
train_dl = DataLoader(datasets[0], batch_size=bs, sampler=train_sampler, drop_last=True, **dl_kwargs)
dataloaders = [train_dl]
for ds in datasets[1:]:
lengths = [len(t) for t in ds.x.items]
sampler = SortSampler(ds.x, key=lengths.__getitem__)
dataloaders.append(DataLoader(ds, batch_size=val_bs, sampler=sampler, **dl_kwargs))
return cls(*dataloaders, path=path, device=device, collate_fn=collate_fn, no_check=no_check)
class Seq2SeqTextList(TextList):
_bunch = Seq2SeqDataBunch
_label_cls = TextList
# Refer to the translation notebook for creation of 'questions_easy.csv'.
df = pd.read_csv(path/'questions_easy.csv')
src = Seq2SeqTextList.from_df(df, path = path, cols='fr').split_by_rand_pct().label_from_df(cols='en', label_cls=TextList)
np.percentile([len(o) for o in src.train.x.items] + [len(o) for o in src.valid.x.items], 90)
np.percentile([len(o) for o in src.train.y.items] + [len(o) for o in src.valid.y.items], 90)
# As before, we remove questions with more than 30 tokens.
src = src.filter_by_func(lambda x,y: len(x) > 30 or len(y) > 30)
len(src.train) + len(src.valid)
data = src.databunch()
data.save()
# Can load from here when restarting.
data = load_data(path)
data.show_batch()
# ## Transformer model
# 
# ### Shifting
# We add a transform to the dataloader that shifts the targets right and adds a padding at the beginning.
def shift_tfm(b):
x,y = b
y = F.pad(y, (1, 0), value=1)
return [x,y[:,:-1]], y[:,1:]
data.add_tfm(shift_tfm)
# ### Embeddings
# The input and output embeddings are traditional PyTorch embeddings (and we can use pretrained vectors if we want to). The transformer model isn't a recurrent one, so it has no idea of the relative positions of the words. To help it with that, they had to the input embeddings a positional encoding which is cosine of a certain frequency:
class PositionalEncoding(nn.Module):
"Encode the position with a sinusoid."
def __init__(self, d:int):
super().__init__()
self.register_buffer('freq', 1 / (10000 ** (torch.arange(0., d, 2.)/d)))
def forward(self, pos:Tensor):
inp = torch.ger(pos, self.freq)
enc = torch.cat([inp.sin(), inp.cos()], dim=-1)
return enc
tst_encoding = PositionalEncoding(20)
res = tst_encoding(torch.arange(0,100).float())
_, ax = plt.subplots(1,1)
for i in range(1,5): ax.plot(res[:,i])
class TransformerEmbedding(nn.Module):
"Embedding + positional encoding + dropout"
def __init__(self, vocab_sz:int, emb_sz:int, inp_p:float=0.):
super().__init__()
self.emb_sz = emb_sz
self.embed = embedding(vocab_sz, emb_sz)
self.pos_enc = PositionalEncoding(emb_sz)
self.drop = nn.Dropout(inp_p)
def forward(self, inp):
pos = torch.arange(0, inp.size(1), device=inp.device).float()
return self.drop(self.embed(inp)* math.sqrt(self.emb_sz) + self.pos_enc(pos))
# ### Feed forward
# The feed forward cell is easy: it's just two linear layers with a skip connection and a LayerNorm.
def feed_forward(d_model:int, d_ff:int, ff_p:float=0., double_drop:bool=True):
layers = [nn.Linear(d_model, d_ff), nn.ReLU()]
if double_drop: layers.append(nn.Dropout(ff_p))
return SequentialEx(*layers, nn.Linear(d_ff, d_model), nn.Dropout(ff_p), MergeLayer(), nn.LayerNorm(d_model))
# ### Multi-head attention
# 
class MultiHeadAttention(nn.Module):
"MutiHeadAttention."
def __init__(self, n_heads:int, d_model:int, d_head:int=None, resid_p:float=0., attn_p:float=0., bias:bool=True,
scale:bool=True):
super().__init__()
d_head = ifnone(d_head, d_model//n_heads)
self.n_heads,self.d_head,self.scale = n_heads,d_head,scale
self.q_wgt = nn.Linear(d_model, n_heads * d_head, bias=bias)
self.k_wgt = nn.Linear(d_model, n_heads * d_head, bias=bias)
self.v_wgt = nn.Linear(d_model, n_heads * d_head, bias=bias)
self.out = nn.Linear(n_heads * d_head, d_model, bias=bias)
self.drop_att,self.drop_res = nn.Dropout(attn_p),nn.Dropout(resid_p)
self.ln = nn.LayerNorm(d_model)
def forward(self, q:Tensor, k:Tensor, v:Tensor, mask:Tensor=None):
return self.ln(q + self.drop_res(self.out(self._apply_attention(q, k, v, mask=mask))))
def _apply_attention(self, q:Tensor, k:Tensor, v:Tensor, mask:Tensor=None):
bs,seq_len = q.size(0),q.size(1)
wq,wk,wv = self.q_wgt(q),self.k_wgt(k),self.v_wgt(v)
wq,wk,wv = map(lambda x:x.view(bs, x.size(1), self.n_heads, self.d_head), (wq,wk,wv))
wq,wk,wv = wq.permute(0, 2, 1, 3),wk.permute(0, 2, 3, 1),wv.permute(0, 2, 1, 3)
attn_score = torch.matmul(wq, wk)
if self.scale: attn_score = attn_score.div_(self.d_head ** 0.5)
if mask is not None:
attn_score = attn_score.float().masked_fill(mask, -float('inf')).type_as(attn_score)
attn_prob = self.drop_att(F.softmax(attn_score, dim=-1))
attn_vec = torch.matmul(attn_prob, wv)
return attn_vec.permute(0, 2, 1, 3).contiguous().contiguous().view(bs, seq_len, -1)
def _attention_einsum(self, q:Tensor, k:Tensor, v:Tensor, mask:Tensor=None):
# Permute and matmul is a little bit faster but this implementation is more readable
bs,seq_len = q.size(0),q.size(1)
wq,wk,wv = self.q_wgt(q),self.k_wgt(k),self.v_wgt(v)
wq,wk,wv = map(lambda x:x.view(bs, x.size(1), self.n_heads, self.d_head), (wq,wk,wv))
attn_score = torch.einsum('bind,bjnd->bijn', (wq, wk))
if self.scale: attn_score = attn_score.mul_(1/(self.d_head ** 0.5))
if mask is not None:
attn_score = attn_score.float().masked_fill(mask, -float('inf')).type_as(attn_score)
attn_prob = self.drop_att(F.softmax(attn_score, dim=2))
attn_vec = torch.einsum('bijn,bjnd->bind', (attn_prob, wv))
return attn_vec.contiguous().view(bs, seq_len, -1)
# ### Masking
# The attention layer uses a mask to avoid paying attention to certain timesteps. The first thing is that we don't really want the network to pay attention to the padding, so we're going to mask it. The second thing is that since this model isn't recurrent, we need to mask (in the output) all the tokens we're not supposed to see yet (otherwise it would be cheating).
def get_padding_mask(inp, pad_idx:int=1):
return None
return (inp == pad_idx)[:,None,:,None]
def get_output_mask(inp, pad_idx:int=1):
return torch.triu(inp.new_ones(inp.size(1),inp.size(1)), diagonal=1)[None,None].byte()
return ((inp == pad_idx)[:,None,:,None].long() + torch.triu(inp.new_ones(inp.size(1),inp.size(1)), diagonal=1)[None,None] != 0)
# Example of mask for the future tokens:
torch.triu(torch.ones(10,10), diagonal=1).byte()
# ### Encoder and decoder blocks
# We are now ready to regroup these layers in the blocks we add in the model picture:
#
# 
class EncoderBlock(nn.Module):
"Encoder block of a Transformer model."
#Can't use Sequential directly cause more than one input...
def __init__(self, n_heads:int, d_model:int, d_head:int, d_inner:int, resid_p:float=0., attn_p:float=0., ff_p:float=0.,
bias:bool=True, scale:bool=True, double_drop:bool=True):
super().__init__()
self.mha = MultiHeadAttention(n_heads, d_model, d_head, resid_p=resid_p, attn_p=attn_p, bias=bias, scale=scale)
self.ff = feed_forward(d_model, d_inner, ff_p=ff_p, double_drop=double_drop)
def forward(self, x:Tensor, mask:Tensor=None): return self.ff(self.mha(x, x, x, mask=mask))
class DecoderBlock(nn.Module):
"Decoder block of a Transformer model."
#Can't use Sequential directly cause more than one input...
def __init__(self, n_heads:int, d_model:int, d_head:int, d_inner:int, resid_p:float=0., attn_p:float=0., ff_p:float=0.,
bias:bool=True, scale:bool=True, double_drop:bool=True):
super().__init__()
self.mha1 = MultiHeadAttention(n_heads, d_model, d_head, resid_p=resid_p, attn_p=attn_p, bias=bias, scale=scale)
self.mha2 = MultiHeadAttention(n_heads, d_model, d_head, resid_p=resid_p, attn_p=attn_p, bias=bias, scale=scale)
self.ff = feed_forward(d_model, d_inner, ff_p=ff_p, double_drop=double_drop)
def forward(self, x:Tensor, enc:Tensor, mask_in:Tensor=None, mask_out:Tensor=None):
y = self.mha1(x, x, x, mask_out)
return self.ff(self.mha2(y, enc, enc, mask=mask_in))
# ### The whole model
class Transformer(nn.Module):
"Transformer model"
def __init__(self, inp_vsz:int, out_vsz:int, n_layers:int=6, n_heads:int=8, d_model:int=256, d_head:int=32,
d_inner:int=1024, inp_p:float=0.1, resid_p:float=0.1, attn_p:float=0.1, ff_p:float=0.1, bias:bool=True,
scale:bool=True, double_drop:bool=True, pad_idx:int=1):
super().__init__()
self.enc_emb = TransformerEmbedding(inp_vsz, d_model, inp_p)
self.dec_emb = TransformerEmbedding(out_vsz, d_model, 0.)
self.encoder = nn.ModuleList([EncoderBlock(n_heads, d_model, d_head, d_inner, resid_p, attn_p,
ff_p, bias, scale, double_drop) for _ in range(n_layers)])
self.decoder = nn.ModuleList([DecoderBlock(n_heads, d_model, d_head, d_inner, resid_p, attn_p,
ff_p, bias, scale, double_drop) for _ in range(n_layers)])
self.out = nn.Linear(d_model, out_vsz)
self.out.weight = self.dec_emb.embed.weight
self.pad_idx = pad_idx
def forward(self, inp, out):
mask_in = get_padding_mask(inp, self.pad_idx)
mask_out = get_output_mask (out, self.pad_idx)
enc,out = self.enc_emb(inp),self.enc_emb(out)
for enc_block in self.encoder: enc = enc_block(enc, mask_in)
for dec_block in self.decoder: out = dec_block(out, enc, mask_in, mask_out)
return self.out(out)
model = Transformer(len(data.train_ds.x.vocab.itos), len(data.train_ds.y.vocab.itos), d_model=300)
# They use LabelSmoothing to get a slightly better accuracy and BLEU.
learn = Learner(data, model, metrics=accuracy, loss_func=FlattenedLoss(LabelSmoothingCrossEntropy, axis=-1))
learn.lr_find()
learn.recorder.plot()
learn.fit(8, 1e-3) #Without label smoothing
learn.fit(8, 1e-3) #With label smoothing
def get_predictions(learn, ds_type=DatasetType.Valid):
learn.model.eval()
inputs, targets, outputs = [],[],[]
with torch.no_grad():
for xb,yb in progress_bar(learn.dl(ds_type)):
out = learn.model(*xb)
for x,y,z in zip(xb[0],xb[1],out):
inputs.append(learn.data.train_ds.x.reconstruct(x))
targets.append(learn.data.train_ds.y.reconstruct(y))
outputs.append(learn.data.train_ds.y.reconstruct(z.argmax(1)))
return inputs, targets, outputs
inputs, targets, outputs = get_predictions(learn)
# Results with label smoothing
inputs[10],targets[10],outputs[10]
inputs[700],targets[700],outputs[700]
inputs[701],targets[701],outputs[701]
inputs[2501],targets[2501],outputs[2501]
inputs[4001],targets[4001],outputs[4001]
# Results without label smoothing
inputs[10],targets[10],outputs[10]
inputs[700],targets[700],outputs[700]
inputs[701],targets[701],outputs[701]
inputs[4001],targets[4001],outputs[4001]
# ### Test leakage
# If we change a token in the targets at position n, it shouldn't impact the predictions before that.
learn.model.eval();
xb,yb = data.one_batch(cpu=False)
inp1,out1 = xb[0][:1],xb[1][:1]
inp2,out2 = inp1.clone(),out1.clone()
out2[0,15] = 10
y1 = learn.model(inp1, out1)
y2 = learn.model(inp2, out2)
(y1[0,:15] - y2[0,:15]).abs().mean()
| dev_course/dl2/translation_transformer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# SUBPROCESSES
# UNIT TESTING
#
# +
import re
my_txt = "An investment in knowledge pays the best interest."
my_txt2 = "abc"
def LetterCompiler(txt):
# txt_combin = ['ab', 'bc', 'ac', 'ba', 'cb', 'ca']
result = re.findall(r'([a-c]).', txt)
# myresult = [combin[1] for combin in txt_combin if combin in txt]
# myresult = []
# for combin in txt_combin:
# if combin in txt and :
# myresult.append(combin[1])
return result
print(LetterCompiler(my_txt2))
# +
import unittest
class TestCompiler(unittest.TestCase):
def test_basic(self):
testcase = "The best preparation for tomorrow is doing your best today."
expected = ['b', 'a', 'a', 'b', 'a']
self.assertEqual(LetterCompiler(testcase), expected)
# -
unittest.main()
# %tb
# Yikes! SystemExit: **True** means an error occurred, as expected. The reason is that unittest.main( ) looks at sys.argv. In Jupyter, by default, the first parameter of sys.argv is what started the Jupyter kernel which is not the case when executing it from the command line. This default parameter is passed into unittest.main( ) as an attribute when you don't explicitly pass it attributes and is therefore what causes the error about the kernel connection file not being a valid attribute. Passing an explicit list to unittest.main( ) prevents it from looking at sys.argv.
#
# Let's pass it the list ['first-arg-is-ignored'] for example. In addition, we will pass it the parameter exit = False to prevent unittest.main( ) from shutting down the kernel process. Run the following cell with the argv and exit parameters passed into unittest.main( ) to rerun your automatic test.
# Use '*' for making text bold, italic and italic bold
unittest.main(argv = ['first-arg-is-ignored'], exit = False)
| JUPYTER_NOTEBOOK_NOTES/19. Managing data and processes.ipynb |
# ##### Copyright 2021 Google LLC.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# # assignment_sat
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/google/or-tools/blob/master/examples/notebook/sat/assignment_sat.ipynb"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/colab_32px.png"/>Run in Google Colab</a>
# </td>
# <td>
# <a href="https://github.com/google/or-tools/blob/master/ortools/sat/samples/assignment_sat.py"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/github_32px.png"/>View source on GitHub</a>
# </td>
# </table>
# First, you must install [ortools](https://pypi.org/project/ortools/) package in this colab.
# !pip install ortools
# +
# #!/usr/bin/env python3
# Copyright 2010-2021 Google LLC
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""Solve a simple assignment problem."""
# [START program]
# [START import]
from ortools.sat.python import cp_model
# [END import]
# Data
# [START data_model]
costs = [
[90, 80, 75, 70],
[35, 85, 55, 65],
[125, 95, 90, 95],
[45, 110, 95, 115],
[50, 100, 90, 100],
]
num_workers = len(costs)
num_tasks = len(costs[0])
# [END data_model]
# Model
# [START model]
model = cp_model.CpModel()
# [END model]
# Variables
# [START variables]
x = []
for i in range(num_workers):
t = []
for j in range(num_tasks):
t.append(model.NewBoolVar(f'x[{i},{j}]'))
x.append(t)
# [END variables]
# Constraints
# [START constraints]
# Each worker is assigned to at most one task.
for i in range(num_workers):
model.Add(sum(x[i][j] for j in range(num_tasks)) <= 1)
# Each task is assigned to exactly one worker.
for j in range(num_tasks):
model.Add(sum(x[i][j] for i in range(num_workers)) == 1)
# [END constraints]
# Objective
# [START objective]
objective_terms = []
for i in range(num_workers):
for j in range(num_tasks):
objective_terms.append(costs[i][j] * x[i][j])
model.Minimize(sum(objective_terms))
# [END objective]
# Solve
# [START solve]
solver = cp_model.CpSolver()
status = solver.Solve(model)
# [END solve]
# Print solution.
# [START print_solution]
if status == cp_model.OPTIMAL or status == cp_model.FEASIBLE:
print(f'Total cost = {solver.ObjectiveValue()}')
print()
for i in range(num_workers):
for j in range(num_tasks):
if solver.BooleanValue(x[i][j]):
print(
f'Worker {i} assigned to task {j} Cost = {costs[i][j]}')
else:
print('No solution found.')
# [END print_solution]
| examples/notebook/sat/assignment_sat.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
dataset_filename = "affinity_dataset.txt"
X = np.loadtxt(dataset_filename)
n_samples, n_features = X.shape
print("This dataset has {0} samples and {1} features".format(n_samples, n_features))
print(X[:5])
# The names of the features, for your reference.
features = ["bread", "milk", "cheese", "apples", "bananas"]
# In our first example, we will compute the Support and Confidence of the rule "If a person buys Apples, they also buy Bananas".
# First, how many rows contain our premise: that a person is buying apples
num_apple_purchases = 0
for sample in X:
if sample[3] == 1: # This person bought Apples
num_apple_purchases += 1
print("{0} people bought Apples".format(num_apple_purchases))
# How many of the cases that a person bought Apples involved the people purchasing Bananas too?
# Record both cases where the rule is valid and is invalid.
rule_valid = 0
rule_invalid = 0
for sample in X:
if sample[3] == 1: # This person bought Apples
if sample[4] == 1:
# This person bought both Apples and Bananas
rule_valid += 1
else:
# This person bought Apples, but not Bananas
rule_invalid += 1
print("{0} cases of the rule being valid were discovered".format(rule_valid))
print("{0} cases of the rule being invalid were discovered".format(rule_invalid))
# Now we have all the information needed to compute Support and Confidence
support = rule_valid # The Support is the number of times the rule is discovered.
confidence = rule_valid / num_apple_purchases
print("The support is {0} and the confidence is {1:.3f}.".format(support, confidence))
# Confidence can be thought of as a percentage using the following:
print("As a percentage, that is {0:.1f}%.".format(100 * confidence))
# +
from collections import defaultdict
# Now compute for all possible rules
valid_rules = defaultdict(int)
invalid_rules = defaultdict(int)
num_occurences = defaultdict(int)
for sample in X:
for premise in range(n_features):
if sample[premise] == 0: continue
# Record that the premise was bought in another transaction
num_occurences[premise] += 1
for conclusion in range(n_features):
if premise == conclusion: # It makes little sense to measure if X -> X.
continue
if sample[conclusion] == 1:
# This person also bought the conclusion item
valid_rules[(premise, conclusion)] += 1
else:
# This person bought the premise, but not the conclusion
invalid_rules[(premise, conclusion)] += 1
support = valid_rules
confidence = defaultdict(float)
for premise, conclusion in valid_rules.keys():
confidence[(premise, conclusion)] = valid_rules[(premise, conclusion)] / num_occurences[premise]
# -
for premise, conclusion in confidence:
premise_name = features[premise]
conclusion_name = features[conclusion]
print("Rule: If a person buys {0} they will also buy {1}".format(premise_name, conclusion_name))
print(" - Confidence: {0:.3f}".format(confidence[(premise, conclusion)]))
print(" - Support: {0}".format(support[(premise, conclusion)]))
print("")
def print_rule(premise, conclusion, support, confidence, features):
premise_name = features[premise]
conclusion_name = features[conclusion]
print("Rule: If a person buys {0} they will also buy {1}".format(premise_name, conclusion_name))
print(" - Confidence: {0:.3f}".format(confidence[(premise, conclusion)]))
print(" - Support: {0}".format(support[(premise, conclusion)]))
print("")
premise = 1
conclusion = 3
print_rule(premise, conclusion, support, confidence, features)
# Sort by support
from pprint import pprint
pprint(list(support.items()))
from operator import itemgetter
sorted_support = sorted(support.items(), key=itemgetter(1), reverse=True)
for index in range(5):
print("Rule #{0}".format(index + 1))
(premise, conclusion) = sorted_support[index][0]
print_rule(premise, conclusion, support, confidence, features)
sorted_confidence = sorted(confidence.items(), key=itemgetter(1), reverse=True)
for index in range(5):
print("Rule #{0}".format(index + 1))
(premise, conclusion) = sorted_confidence[index][0]
print_rule(premise, conclusion, support, confidence, features)
| Chapter 1/.ipynb_checkpoints/ch1_affinity-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# # Chapter 08 - Classification Example
#
# Shows how to use Azure Machine Learning AutoML for classification
# ## Installation of relevant packages
# skip this step if you already have seaborn installed.
# !pip install seaborn
# ## Getting Ready
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
import logging
# import AzureML SDK
import azureml.core
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
from azureml.train.automl import AutoMLConfig
# ## Preparing the Azure Machine Learning Workspace
#
# Ignore this section if the workspace has already been created in the Azure portal
# +
ws = Workspace.from_config()
workspace_info = {}
workspace_info['SDK version'] = azureml.core.VERSION
workspace_info['Subscription ID'] = ws.subscription_id
workspace_info['Workspace Name'] = ws.name
workspace_info['Resource Group'] = ws.resource_group
workspace_info['Location'] = ws.location
pd.set_option('display.max_colwidth', -1)
workspace_info = pd.DataFrame(data = workspace_info, index = [''])
workspace_info.T
# +
# Choose a name for the experiment and specify the project folder.
experiment_name = 'automl-classification'
project_folder = './book/automl-classification'
experiment = Experiment(ws, experiment_name)
# -
# ## Data Preparation
#define the column
columns = ['status_checking_acc', 'duration_months', 'credit_history', 'purpose', 'credit_amount',
'saving_acc_bonds', 'present_emp_since','installment_rate', 'personal_status','other_debtors',
'residing_since', 'property', 'age_years','inst_plans', 'housing', 'num_existing_credits',
'job', 'dependents', 'telephone', 'foreign_worker', 'status']
creditg_df = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/statlog/german/german.data', delim_whitespace = True, header = None )
creditg_df.columns = columns
creditg_df.head()
# +
# Get the unique values in the Status Column
creditg_df.status = creditg_df.status - 1
creditg_df['status'].unique()
# -
#creditg_df['status'] = creditg_df.status.astype(bool)
creditg_df.head()
# +
# Get the label column, and remove the label column from the dataframe
target = creditg_df["status"]
# when axis is 1, columns specified are dropped
creditg_df = creditg_df.drop(labels='status',axis=1)
# +
# Split into train and test data
X_train, X_test, y_train, y_test = train_test_split(creditg_df, target, test_size=0.3)
# convert y_train to a numpy array
y_train = y_train.values
y_test = y_test.values
# -
# Understand the variables
creditg_df.info()
import matplotlib.pyplot as plt
import seaborn as sn
sn.distplot(creditg_df.credit_amount)
plt.title("Distribution of Credit Amount")
plt.ylabel("Frequency")
sn.distplot(target, kde=False, rug=True )
plt.title("Credit Risk")
plt.ylabel("Frequency")
# ## Using AutoML to train the model
X_train.head()
len(y_train)
# Explore the metrics that are available for classification
azureml.train.automl.utilities.get_primary_metrics('classification')
# +
import time
automl_settings = {
"name": "AutoML_Book_CH08_Classification_{0}".format(time.time()),
"iteration_timeout_minutes": 10,
"iterations": 10,
"primary_metric": 'AUC_weighted',
"preprocess": True,
"max_concurrent_iterations": 10,
"verbosity": logging.INFO
}
# -
# ValueError: The training data contains datetime, categorical or text data. Please set preprocess flag as True
# Set Preprocess= True
automl_config = AutoMLConfig(task = 'classification',
debug_log = 'automl_errors.log',
X = X_train,
y = y_train,
**automl_settings,
n_cross_validations = 5,
path = project_folder
)
local_run = experiment.submit(automl_config, show_output = True)
# ValueError: Invalid primary metric specified for classificatio
# lease use on of: ['norm_macro_recall', 'precision_score_weighted', 'average_precision_score_weighted', 'accuracy', 'AUC_weighted']
local_run
# ## Understanding the results of the AutoML Run
# !pip install azureml-widgets
import azureml.widgets
from azureml.widgets import RunDetails
RunDetails(local_run).show()
# +
# Get all child runs
children = list(local_run.get_children())
metricslist = {}
for run in children:
properties = run.get_properties()
metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}
metricslist[int(properties['iteration'])] = metrics
rundata = pd.DataFrame(metricslist).sort_index(1)
rundata
# -
# ## Let's get the best model sofar
#
local_run
best_run, fitted_model = local_run.get_output(metric = "AUC_weighted")
print(best_run)
print(fitted_model)
# ## Testing the model
from sklearn.metrics import classification_report
from sklearn.metrics import roc_auc_score
y_pred = fitted_model.predict(X_test)
len(y_pred)
len(y_test)
target_names = ['0','1']
print (classification_report(y_test,y_pred, target_names=target_names))
print(roc_auc_score(y_test,y_pred))
# ## Explainability of Models
project_folder = './book/automl-classification-explain'
automl_explain_config = AutoMLConfig(task = 'classification',
model_explainability=True,
debug_log = 'automl_errors.log',
X = X_train,
y = y_train,
X_valid = X_test,
y_valid = y_test,
**automl_settings,
path = project_folder
)
# +
from azureml.core.experiment import Experiment
experiment_e=Experiment(ws, 'automl_explain')
local_explain_run = experiment_e.submit(automl_explain_config, show_output = True)
# -
best_run, fitted_model = local_explain_run.get_output()
# +
from azureml.widgets import RunDetails
RunDetails(local_explain_run).show()
# +
from azureml.train.automl.automlexplainer import retrieve_model_explanation
shap_values, expected_values, overall_summary, overall_imp, per_class_summary, per_class_imp = \
retrieve_model_explanation(best_run)
# +
print(overall_summary)
print(overall_imp)
# -
| Chapter 08 - Classification.ipynb |
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + gradient={"editing": false}
#hide
# !pip install -Uqq fastbook
import fastbook
fastbook.setup_book()
# + gradient={"editing": false}
#hide
from fastbook import *
# + gradient={"editing": false} active=""
# [[chapter_nlp_dive]]
# + [markdown] gradient={"editing": false}
# # A Language Model from Scratch
# + [markdown] gradient={"editing": false}
# We're now ready to go deep... deep into deep learning! You already learned how to train a basic neural network, but how do you go from there to creating state-of-the-art models? In this part of the book we're going to uncover all of the mysteries, starting with language models.
#
# You saw in <<chapter_nlp>> how to fine-tune a pretrained language model to build a text classifier. In this chapter, we will explain to you what exactly is inside that model, and what an RNN is. First, let's gather some data that will allow us to quickly prototype our various models.
# + [markdown] gradient={"editing": false}
# ## The Data
# + [markdown] gradient={"editing": false}
# Whenever we start working on a new problem, we always first try to think of the simplest dataset we can that will allow us to try out methods quickly and easily, and interpret the results. When we started working on language modeling a few years ago we didn't find any datasets that would allow for quick prototyping, so we made one. We call it *Human Numbers*, and it simply contains the first 10,000 numbers written out in English.
# + [markdown] gradient={"editing": false}
# > j: One of the most common practical mistakes I see even amongst highly experienced practitioners is failing to use appropriate datasets at appropriate times during the analysis process. In particular, most people tend to start with datasets that are too big and too complicated.
# -
# We can download, extract, and take a look at our dataset in the usual way:
# + gradient={}
from fastai.text.all import *
path = untar_data(URLs.HUMAN_NUMBERS)
# + gradient={}
#hide
Path.BASE_PATH = path
# + gradient={}
path.ls()
# -
# Let's open those two files and see what's inside. At first we'll join all of the texts together and ignore the train/valid split given by the dataset (we'll come back to that later):
# + gradient={}
lines = L()
with open(path/'train.txt') as f: lines += L(*f.readlines())
with open(path/'valid.txt') as f: lines += L(*f.readlines())
lines
# -
# We take all those lines and concatenate them in one big stream. To mark when we go from one number to the next, we use a `.` as a separator:
# + gradient={}
text = ' . '.join([l.strip() for l in lines])
text[:100]
# -
# We can tokenize this dataset by splitting on spaces:
# + gradient={}
tokens = text.split(' ')
tokens[:10]
# -
# To numericalize, we have to create a list of all the unique tokens (our *vocab*):
# + gradient={}
vocab = L(*tokens).unique()
vocab
# -
# Then we can convert our tokens into numbers by looking up the index of each in the vocab:
# + gradient={}
word2idx = {w:i for i,w in enumerate(vocab)}
nums = L(word2idx[i] for i in tokens)
nums
# -
# Now that we have a small dataset on which language modeling should be an easy task, we can build our first model.
# ## Our First Language Model from Scratch
# One simple way to turn this into a neural network would be to specify that we are going to predict each word based on the previous three words. We could create a list of every sequence of three words as our independent variables, and the next word after each sequence as the dependent variable.
#
# We can do that with plain Python. Let's do it first with tokens just to confirm what it looks like:
# + gradient={}
L((tokens[i:i+3], tokens[i+3]) for i in range(0,len(tokens)-4,3))
# -
# Now we will do it with tensors of the numericalized values, which is what the model will actually use:
# + gradient={}
seqs = L((tensor(nums[i:i+3]), nums[i+3]) for i in range(0,len(nums)-4,3))
seqs
# -
# We can batch those easily using the `DataLoader` class. For now we will split the sequences randomly:
# + gradient={}
bs = 64
cut = int(len(seqs) * 0.8)
dls = DataLoaders.from_dsets(seqs[:cut], seqs[cut:], bs=64, shuffle=False)
# -
# We can now create a neural network architecture that takes three words as input, and returns a prediction of the probability of each possible next word in the vocab. We will use three standard linear layers, but with two tweaks.
#
# The first tweak is that the first linear layer will use only the first word's embedding as activations, the second layer will use the second word's embedding plus the first layer's output activations, and the third layer will use the third word's embedding plus the second layer's output activations. The key effect of this is that every word is interpreted in the information context of any words preceding it.
#
# The second tweak is that each of these three layers will use the same weight matrix. The way that one word impacts the activations from previous words should not change depending on the position of a word. In other words, activation values will change as data moves through the layers, but the layer weights themselves will not change from layer to layer. So, a layer does not learn one sequence position; it must learn to handle all positions.
#
# Since layer weights do not change, you might think of the sequential layers as "the same layer" repeated. In fact, PyTorch makes this concrete; we can just create one layer, and use it multiple times.
# ### Our Language Model in PyTorch
# We can now create the language model module that we described earlier:
# + gradient={}
class LMModel1(Module):
def __init__(self, vocab_sz, n_hidden):
self.i_h = nn.Embedding(vocab_sz, n_hidden)
self.h_h = nn.Linear(n_hidden, n_hidden)
self.h_o = nn.Linear(n_hidden,vocab_sz)
def forward(self, x):
h = F.relu(self.h_h(self.i_h(x[:,0])))
h = h + self.i_h(x[:,1])
h = F.relu(self.h_h(h))
h = h + self.i_h(x[:,2])
h = F.relu(self.h_h(h))
return self.h_o(h)
# -
# As you see, we have created three layers:
#
# - The embedding layer (`i_h`, for *input* to *hidden*)
# - The linear layer to create the activations for the next word (`h_h`, for *hidden* to *hidden*)
# - A final linear layer to predict the fourth word (`h_o`, for *hidden* to *output*)
#
# This might be easier to represent in pictorial form, so let's define a simple pictorial representation of basic neural networks. <<img_simple_nn>> shows how we're going to represent a neural net with one hidden layer.
# <img alt="Pictorial representation of simple neural network" width="400" src="images/att_00020.png" caption="Pictorial representation of a simple neural network" id="img_simple_nn">
# Each shape represents activations: rectangle for input, circle for hidden (inner) layer activations, and triangle for output activations. We will use those shapes (summarized in <<img_shapes>>) in all the diagrams in this chapter.
# <img alt="Shapes used in our pictorial representations" width="200" src="images/att_00021.png" id="img_shapes" caption="Shapes used in our pictorial representations">
# An arrow represents the actual layer computation—i.e., the linear layer followed by the activation function. Using this notation, <<lm_rep>> shows what our simple language model looks like.
# <img alt="Representation of our basic language model" width="500" caption="Representation of our basic language model" id="lm_rep" src="images/att_00022.png">
# To simplify things, we've removed the details of the layer computation from each arrow. We've also color-coded the arrows, such that all arrows with the same color have the same weight matrix. For instance, all the input layers use the same embedding matrix, so they all have the same color (green).
#
# Let's try training this model and see how it goes:
# + gradient={}
learn = Learner(dls, LMModel1(len(vocab), 64), loss_func=F.cross_entropy,
metrics=accuracy)
learn.fit_one_cycle(4, 1e-3)
# -
# To see if this is any good, let's check what a very simple model would give us. In this case we could always predict the most common token, so let's find out which token is most often the target in our validation set:
# + gradient={}
n,counts = 0,torch.zeros(len(vocab))
for x,y in dls.valid:
n += y.shape[0]
for i in range_of(vocab): counts[i] += (y==i).long().sum()
idx = torch.argmax(counts)
idx, vocab[idx.item()], counts[idx].item()/n
# -
# The most common token has the index 29, which corresponds to the token `thousand`. Always predicting this token would give us an accuracy of roughly 15\%, so we are faring way better!
# > A: My first guess was that the separator would be the most common token, since there is one for every number. But looking at `tokens` reminded me that large numbers are written with many words, so on the way to 10,000 you write "thousand" a lot: five thousand, five thousand and one, five thousand and two, etc. Oops! Looking at your data is great for noticing subtle features and also embarrassingly obvious ones.
# This is a nice first baseline. Let's see how we can refactor it with a loop.
# ### Our First Recurrent Neural Network
# Looking at the code for our module, we could simplify it by replacing the duplicated code that calls the layers with a `for` loop. As well as making our code simpler, this will also have the benefit that we will be able to apply our module equally well to token sequences of different lengths—we won't be restricted to token lists of length three:
# + gradient={}
class LMModel2(Module):
def __init__(self, vocab_sz, n_hidden):
self.i_h = nn.Embedding(vocab_sz, n_hidden)
self.h_h = nn.Linear(n_hidden, n_hidden)
self.h_o = nn.Linear(n_hidden,vocab_sz)
def forward(self, x):
h = 0
for i in range(3):
h = h + self.i_h(x[:,i])
h = F.relu(self.h_h(h))
return self.h_o(h)
# -
# Let's check that we get the same results using this refactoring:
# + gradient={}
learn = Learner(dls, LMModel2(len(vocab), 64), loss_func=F.cross_entropy,
metrics=accuracy)
learn.fit_one_cycle(4, 1e-3)
# -
# We can also refactor our pictorial representation in exactly the same way, as shown in <<basic_rnn>> (we're also removing the details of activation sizes here, and using the same arrow colors as in <<lm_rep>>).
# <img alt="Basic recurrent neural network" width="400" caption="Basic recurrent neural network" id="basic_rnn" src="images/att_00070.png">
# You will see that there is a set of activations that are being updated each time through the loop, stored in the variable `h`—this is called the *hidden state*.
# > Jargon: hidden state: The activations that are updated at each step of a recurrent neural network.
# A neural network that is defined using a loop like this is called a *recurrent neural network* (RNN). It is important to realize that an RNN is not a complicated new architecture, but simply a refactoring of a multilayer neural network using a `for` loop.
#
# > A: My true opinion: if they were called "looping neural networks," or LNNs, they would seem 50% less daunting!
# Now that we know what an RNN is, let's try to make it a little bit better.
# ## Improving the RNN
# Looking at the code for our RNN, one thing that seems problematic is that we are initializing our hidden state to zero for every new input sequence. Why is that a problem? We made our sample sequences short so they would fit easily into batches. But if we order the samples correctly, those sample sequences will be read in order by the model, exposing the model to long stretches of the original sequence.
#
# Another thing we can look at is having more signal: why only predict the fourth word when we could use the intermediate predictions to also predict the second and third words?
#
# Let's see how we can implement those changes, starting with adding some state.
# ### Maintaining the State of an RNN
# Because we initialize the model's hidden state to zero for each new sample, we are throwing away all the information we have about the sentences we have seen so far, which means that our model doesn't actually know where we are up to in the overall counting sequence. This is easily fixed; we can simply move the initialization of the hidden state to `__init__`.
#
# But this fix will create its own subtle, but important, problem. It effectively makes our neural network as deep as the entire number of tokens in our document. For instance, if there were 10,000 tokens in our dataset, we would be creating a 10,000-layer neural network.
#
# To see why this is the case, consider the original pictorial representation of our recurrent neural network in <<lm_rep>>, before refactoring it with a `for` loop. You can see each layer corresponds with one token input. When we talk about the representation of a recurrent neural network before refactoring with the `for` loop, we call this the *unrolled representation*. It is often helpful to consider the unrolled representation when trying to understand an RNN.
#
# The problem with a 10,000-layer neural network is that if and when you get to the 10,000th word of the dataset, you will still need to calculate the derivatives all the way back to the first layer. This is going to be very slow indeed, and very memory-intensive. It is unlikely that you'll be able to store even one mini-batch on your GPU.
#
# The solution to this problem is to tell PyTorch that we do not want to back propagate the derivatives through the entire implicit neural network. Instead, we will just keep the last three layers of gradients. To remove all of the gradient history in PyTorch, we use the `detach` method.
#
# Here is the new version of our RNN. It is now stateful, because it remembers its activations between different calls to `forward`, which represent its use for different samples in the batch:
# + gradient={}
class LMModel3(Module):
def __init__(self, vocab_sz, n_hidden):
self.i_h = nn.Embedding(vocab_sz, n_hidden)
self.h_h = nn.Linear(n_hidden, n_hidden)
self.h_o = nn.Linear(n_hidden,vocab_sz)
self.h = 0
def forward(self, x):
for i in range(3):
self.h = self.h + self.i_h(x[:,i])
self.h = F.relu(self.h_h(self.h))
out = self.h_o(self.h)
self.h = self.h.detach()
return out
def reset(self): self.h = 0
# -
# This model will have the same activations whatever sequence length we pick, because the hidden state will remember the last activation from the previous batch. The only thing that will be different is the gradients computed at each step: they will only be calculated on sequence length tokens in the past, instead of the whole stream. This approach is called *backpropagation through time* (BPTT).
# > jargon: Back propagation through time (BPTT): Treating a neural net with effectively one layer per time step (usually refactored using a loop) as one big model, and calculating gradients on it in the usual way. To avoid running out of memory and time, we usually use _truncated_ BPTT, which "detaches" the history of computation steps in the hidden state every few time steps.
# To use `LMModel3`, we need to make sure the samples are going to be seen in a certain order. As we saw in <<chapter_nlp>>, if the first line of the first batch is our `dset[0]` then the second batch should have `dset[1]` as the first line, so that the model sees the text flowing.
#
# `LMDataLoader` was doing this for us in <<chapter_nlp>>. This time we're going to do it ourselves.
#
# To do this, we are going to rearrange our dataset. First we divide the samples into `m = len(dset) // bs` groups (this is the equivalent of splitting the whole concatenated dataset into, for example, 64 equally sized pieces, since we're using `bs=64` here). `m` is the length of each of these pieces. For instance, if we're using our whole dataset (although we'll actually split it into train versus valid in a moment), that will be:
# + gradient={}
m = len(seqs)//bs
m,bs,len(seqs)
# -
# The first batch will be composed of the samples:
#
# (0, m, 2*m, ..., (bs-1)*m)
#
# the second batch of the samples:
#
# (1, m+1, 2*m+1, ..., (bs-1)*m+1)
#
# and so forth. This way, at each epoch, the model will see a chunk of contiguous text of size `3*m` (since each text is of size 3) on each line of the batch.
#
# The following function does that reindexing:
# + gradient={}
def group_chunks(ds, bs):
m = len(ds) // bs
new_ds = L()
for i in range(m): new_ds += L(ds[i + m*j] for j in range(bs))
return new_ds
# -
# Then we just pass `drop_last=True` when building our `DataLoaders` to drop the last batch that does not have a shape of `bs`. We also pass `shuffle=False` to make sure the texts are read in order:
# + gradient={}
cut = int(len(seqs) * 0.8)
dls = DataLoaders.from_dsets(
group_chunks(seqs[:cut], bs),
group_chunks(seqs[cut:], bs),
bs=bs, drop_last=True, shuffle=False)
# -
# The last thing we add is a little tweak of the training loop via a `Callback`. We will talk more about callbacks in <<chapter_accel_sgd>>; this one will call the `reset` method of our model at the beginning of each epoch and before each validation phase. Since we implemented that method to zero the hidden state of the model, this will make sure we start with a clean state before reading those continuous chunks of text. We can also start training a bit longer:
# + gradient={}
learn = Learner(dls, LMModel3(len(vocab), 64), loss_func=F.cross_entropy,
metrics=accuracy, cbs=ModelResetter)
learn.fit_one_cycle(10, 3e-3)
# -
# This is already better! The next step is to use more targets and compare them to the intermediate predictions.
# ### Creating More Signal
# Another problem with our current approach is that we only predict one output word for each three input words. That means that the amount of signal that we are feeding back to update weights with is not as large as it could be. It would be better if we predicted the next word after every single word, rather than every three words, as shown in <<stateful_rep>>.
# <img alt="RNN predicting after every token" width="400" caption="RNN predicting after every token" id="stateful_rep" src="images/att_00024.png">
# This is easy enough to add. We need to first change our data so that the dependent variable has each of the three next words after each of our three input words. Instead of `3`, we use an attribute, `sl` (for sequence length), and make it a bit bigger:
# + gradient={}
sl = 16
seqs = L((tensor(nums[i:i+sl]), tensor(nums[i+1:i+sl+1]))
for i in range(0,len(nums)-sl-1,sl))
cut = int(len(seqs) * 0.8)
dls = DataLoaders.from_dsets(group_chunks(seqs[:cut], bs),
group_chunks(seqs[cut:], bs),
bs=bs, drop_last=True, shuffle=False)
# -
# Looking at the first element of `seqs`, we can see that it contains two lists of the same size. The second list is the same as the first, but offset by one element:
# + gradient={}
[L(vocab[o] for o in s) for s in seqs[0]]
# -
# Now we need to modify our model so that it outputs a prediction after every word, rather than just at the end of a three-word sequence:
# + gradient={}
class LMModel4(Module):
def __init__(self, vocab_sz, n_hidden):
self.i_h = nn.Embedding(vocab_sz, n_hidden)
self.h_h = nn.Linear(n_hidden, n_hidden)
self.h_o = nn.Linear(n_hidden,vocab_sz)
self.h = 0
def forward(self, x):
outs = []
for i in range(sl):
self.h = self.h + self.i_h(x[:,i])
self.h = F.relu(self.h_h(self.h))
outs.append(self.h_o(self.h))
self.h = self.h.detach()
return torch.stack(outs, dim=1)
def reset(self): self.h = 0
# -
# This model will return outputs of shape `bs x sl x vocab_sz` (since we stacked on `dim=1`). Our targets are of shape `bs x sl`, so we need to flatten those before using them in `F.cross_entropy`:
# + gradient={}
def loss_func(inp, targ):
return F.cross_entropy(inp.view(-1, len(vocab)), targ.view(-1))
# -
# We can now use this loss function to train the model:
# + gradient={}
learn = Learner(dls, LMModel4(len(vocab), 64), loss_func=loss_func,
metrics=accuracy, cbs=ModelResetter)
learn.fit_one_cycle(15, 3e-3)
# -
# We need to train for longer, since the task has changed a bit and is more complicated now. But we end up with a good result... At least, sometimes. If you run it a few times, you'll see that you can get quite different results on different runs. That's because effectively we have a very deep network here, which can result in very large or very small gradients. We'll see in the next part of this chapter how to deal with this.
#
# Now, the obvious way to get a better model is to go deeper: we only have one linear layer between the hidden state and the output activations in our basic RNN, so maybe we'll get better results with more.
# ## Multilayer RNNs
# In a multilayer RNN, we pass the activations from our recurrent neural network into a second recurrent neural network, like in <<stacked_rnn_rep>>.
# <img alt="2-layer RNN" width="550" caption="2-layer RNN" id="stacked_rnn_rep" src="images/att_00025.png">
# The unrolled representation is shown in <<unrolled_stack_rep>> (similar to <<lm_rep>>).
# <img alt="2-layer unrolled RNN" width="500" caption="Two-layer unrolled RNN" id="unrolled_stack_rep" src="images/att_00026.png">
# Let's see how to implement this in practice.
# ### The Model
# We can save some time by using PyTorch's `RNN` class, which implements exactly what we created earlier, but also gives us the option to stack multiple RNNs, as we have discussed:
# + gradient={}
class LMModel5(Module):
def __init__(self, vocab_sz, n_hidden, n_layers):
self.i_h = nn.Embedding(vocab_sz, n_hidden)
self.rnn = nn.RNN(n_hidden, n_hidden, n_layers, batch_first=True)
self.h_o = nn.Linear(n_hidden, vocab_sz)
self.h = torch.zeros(n_layers, bs, n_hidden)
def forward(self, x):
res,h = self.rnn(self.i_h(x), self.h)
self.h = h.detach()
return self.h_o(res)
def reset(self): self.h.zero_()
# + gradient={}
learn = Learner(dls, LMModel5(len(vocab), 64, 2),
loss_func=CrossEntropyLossFlat(),
metrics=accuracy, cbs=ModelResetter)
learn.fit_one_cycle(15, 3e-3)
# -
# Now that's disappointing... our previous single-layer RNN performed better. Why? The reason is that we have a deeper model, leading to exploding or vanishing activations.
# ### Exploding or Disappearing Activations
# In practice, creating accurate models from this kind of RNN is difficult. We will get better results if we call `detach` less often, and have more layers—this gives our RNN a longer time horizon to learn from, and richer features to create. But it also means we have a deeper model to train. The key challenge in the development of deep learning has been figuring out how to train these kinds of models.
#
# The reason this is challenging is because of what happens when you multiply by a matrix many times. Think about what happens when you multiply by a number many times. For example, if you multiply by 2, starting at 1, you get the sequence 1, 2, 4, 8,... after 32 steps you are already at 4,294,967,296. A similar issue happens if you multiply by 0.5: you get 0.5, 0.25, 0.125… and after 32 steps it's 0.00000000023. As you can see, multiplying by a number even slightly higher or lower than 1 results in an explosion or disappearance of our starting number, after just a few repeated multiplications.
#
# Because matrix multiplication is just multiplying numbers and adding them up, exactly the same thing happens with repeated matrix multiplications. And that's all a deep neural network is —each extra layer is another matrix multiplication. This means that it is very easy for a deep neural network to end up with extremely large or extremely small numbers.
#
# This is a problem, because the way computers store numbers (known as "floating point") means that they become less and less accurate the further away the numbers get from zero. The diagram in <<float_prec>>, from the excellent article ["What You Never Wanted to Know About Floating Point but Will Be Forced to Find Out"](http://www.volkerschatz.com/science/float.html), shows how the precision of floating-point numbers varies over the number line.
# <img alt="Precision of floating point numbers" width="1000" caption="Precision of floating-point numbers" id="float_prec" src="images/fltscale.svg">
# This inaccuracy means that often the gradients calculated for updating the weights end up as zero or infinity for deep networks. This is commonly referred to as the *vanishing gradients* or *exploding gradients* problem. It means that in SGD, the weights are either not updated at all or jump to infinity. Either way, they won't improve with training.
#
# Researchers have developed a number of ways to tackle this problem, which we will be discussing later in the book. One option is to change the definition of a layer in a way that makes it less likely to have exploding activations. We'll look at the details of how this is done in <<chapter_convolutions>>, when we discuss batch normalization, and <<chapter_resnet>>, when we discuss ResNets, although these details don't generally matter in practice (unless you are a researcher that is creating new approaches to solving this problem). Another strategy for dealing with this is by being careful about initialization, which is a topic we'll investigate in <<chapter_foundations>>.
#
# For RNNs, there are two types of layers that are frequently used to avoid exploding activations: *gated recurrent units* (GRUs) and *long short-term memory* (LSTM) layers. Both of these are available in PyTorch, and are drop-in replacements for the RNN layer. We will only cover LSTMs in this book; there are plenty of good tutorials online explaining GRUs, which are a minor variant on the LSTM design.
# ## LSTM
# LSTM is an architecture that was introduced back in 1997 by <NAME> and <NAME>. In this architecture, there are not one but two hidden states. In our base RNN, the hidden state is the output of the RNN at the previous time step. That hidden state is then responsible for two things:
#
# - Having the right information for the output layer to predict the correct next token
# - Retaining memory of everything that happened in the sentence
#
# Consider, for example, the sentences "Henry has a dog and he likes his dog very much" and "Sophie has a dog and she likes her dog very much." It's very clear that the RNN needs to remember the name at the beginning of the sentence to be able to predict *he/she* or *his/her*.
#
# In practice, RNNs are really bad at retaining memory of what happened much earlier in the sentence, which is the motivation to have another hidden state (called *cell state*) in the LSTM. The cell state will be responsible for keeping *long short-term memory*, while the hidden state will focus on the next token to predict. Let's take a closer look at how this is achieved and build an LSTM from scratch.
# ### Building an LSTM from Scratch
# In order to build an LSTM, we first have to understand its architecture. <<lstm>> shows its inner structure.
#
# <img src="images/LSTM.png" id="lstm" caption="Architecture of an LSTM" alt="A graph showing the inner architecture of an LSTM" width="700">
# In this picture, our input $x_{t}$ enters on the left with the previous hidden state ($h_{t-1}$) and cell state ($c_{t-1}$). The four orange boxes represent four layers (our neural nets) with the activation being either sigmoid ($\sigma$) or tanh. tanh is just a sigmoid function rescaled to the range -1 to 1. Its mathematical expression can be written like this:
#
# $$\tanh(x) = \frac{e^{x} - e^{-x}}{e^{x}+e^{-x}} = 2 \sigma(2x) - 1$$
#
# where $\sigma$ is the sigmoid function. The green circles are elementwise operations. What goes out on the right is the new hidden state ($h_{t}$) and new cell state ($c_{t}$), ready for our next input. The new hidden state is also used as output, which is why the arrow splits to go up.
#
# Let's go over the four neural nets (called *gates*) one by one and explain the diagram—but before this, notice how very little the cell state (at the top) is changed. It doesn't even go directly through a neural net! This is exactly why it will carry on a longer-term state.
#
# First, the arrows for input and old hidden state are joined together. In the RNN we wrote earlier in this chapter, we were adding them together. In the LSTM, we stack them in one big tensor. This means the dimension of our embeddings (which is the dimension of $x_{t}$) can be different than the dimension of our hidden state. If we call those `n_in` and `n_hid`, the arrow at the bottom is of size `n_in + n_hid`; thus all the neural nets (orange boxes) are linear layers with `n_in + n_hid` inputs and `n_hid` outputs.
#
# The first gate (looking from left to right) is called the *forget gate*. Since it’s a linear layer followed by a sigmoid, its output will consist of scalars between 0 and 1. We multiply this result by the cell state to determine which information to keep and which to throw away: values closer to 0 are discarded and values closer to 1 are kept. This gives the LSTM the ability to forget things about its long-term state. For instance, when crossing a period or an `xxbos` token, we would expect to it to (have learned to) reset its cell state.
#
# The second gate is called the *input gate*. It works with the third gate (which doesn't really have a name but is sometimes called the *cell gate*) to update the cell state. For instance, we may see a new gender pronoun, in which case we'll need to replace the information about gender that the forget gate removed. Similar to the forget gate, the input gate decides which elements of the cell state to update (values close to 1) or not (values close to 0). The third gate determines what those updated values are, in the range of –1 to 1 (thanks to the tanh function). The result is then added to the cell state.
#
# The last gate is the *output gate*. It determines which information from the cell state to use to generate the output. The cell state goes through a tanh before being combined with the sigmoid output from the output gate, and the result is the new hidden state.
#
# In terms of code, we can write the same steps like this:
# + gradient={}
class LSTMCell(Module):
def __init__(self, ni, nh):
self.forget_gate = nn.Linear(ni + nh, nh)
self.input_gate = nn.Linear(ni + nh, nh)
self.cell_gate = nn.Linear(ni + nh, nh)
self.output_gate = nn.Linear(ni + nh, nh)
def forward(self, input, state):
h,c = state
h = torch.cat([h, input], dim=1)
forget = torch.sigmoid(self.forget_gate(h))
c = c * forget
inp = torch.sigmoid(self.input_gate(h))
cell = torch.tanh(self.cell_gate(h))
c = c + inp * cell
out = torch.sigmoid(self.output_gate(h))
h = out * torch.tanh(c)
return h, (h,c)
# -
# In practice, we can then refactor the code. Also, in terms of performance, it's better to do one big matrix multiplication than four smaller ones (that's because we only launch the special fast kernel on the GPU once, and it gives the GPU more work to do in parallel). The stacking takes a bit of time (since we have to move one of the tensors around on the GPU to have it all in a contiguous array), so we use two separate layers for the input and the hidden state. The optimized and refactored code then looks like this:
# + gradient={}
class LSTMCell(Module):
def __init__(self, ni, nh):
self.ih = nn.Linear(ni,4*nh)
self.hh = nn.Linear(nh,4*nh)
def forward(self, input, state):
h,c = state
# One big multiplication for all the gates is better than 4 smaller ones
gates = (self.ih(input) + self.hh(h)).chunk(4, 1)
ingate,forgetgate,outgate = map(torch.sigmoid, gates[:3])
cellgate = gates[3].tanh()
c = (forgetgate*c) + (ingate*cellgate)
h = outgate * c.tanh()
return h, (h,c)
# -
# Here we use the PyTorch `chunk` method to split our tensor into four pieces. It works like this:
# + gradient={}
t = torch.arange(0,10); t
# + gradient={}
t.chunk(2)
# -
# Let's now use this architecture to train a language model!
# ### Training a Language Model Using LSTMs
# Here is the same network as `LMModel5`, using a two-layer LSTM. We can train it at a higher learning rate, for a shorter time, and get better accuracy:
# + gradient={}
class LMModel6(Module):
def __init__(self, vocab_sz, n_hidden, n_layers):
self.i_h = nn.Embedding(vocab_sz, n_hidden)
self.rnn = nn.LSTM(n_hidden, n_hidden, n_layers, batch_first=True)
self.h_o = nn.Linear(n_hidden, vocab_sz)
self.h = [torch.zeros(n_layers, bs, n_hidden) for _ in range(2)]
def forward(self, x):
res,h = self.rnn(self.i_h(x), self.h)
self.h = [h_.detach() for h_ in h]
return self.h_o(res)
def reset(self):
for h in self.h: h.zero_()
# + gradient={}
learn = Learner(dls, LMModel6(len(vocab), 64, 2),
loss_func=CrossEntropyLossFlat(),
metrics=accuracy, cbs=ModelResetter)
learn.fit_one_cycle(15, 1e-2)
# -
# Now that's better than a multilayer RNN! We can still see there is a bit of overfitting, however, which is a sign that a bit of regularization might help.
# ## Regularizing an LSTM
# Recurrent neural networks, in general, are hard to train, because of the problem of vanishing activations and gradients we saw before. Using LSTM (or GRU) cells makes training easier than with vanilla RNNs, but they are still very prone to overfitting. Data augmentation, while a possibility, is less often used for text data than for images because in most cases it requires another model to generate random augmentations (e.g., by translating the text into another language and then back into the original language). Overall, data augmentation for text data is currently not a well-explored space.
#
# However, there are other regularization techniques we can use instead to reduce overfitting, which were thoroughly studied for use with LSTMs in the paper ["Regularizing and Optimizing LSTM Language Models"](https://arxiv.org/abs/1708.02182) by <NAME>, <NAME>, and <NAME>. This paper showed how effective use of *dropout*, *activation regularization*, and *temporal activation regularization* could allow an LSTM to beat state-of-the-art results that previously required much more complicated models. The authors called an LSTM using these techniques an *AWD-LSTM*. We'll look at each of these techniques in turn.
# ### Dropout
# Dropout is a regularization technique that was introduced by <NAME> al. in [Improving neural networks by preventing co-adaptation of feature detectors](https://arxiv.org/abs/1207.0580). The basic idea is to randomly change some activations to zero at training time. This makes sure all neurons actively work toward the output, as seen in <<img_dropout>> (from "Dropout: A Simple Way to Prevent Neural Networks from Overfitting" by Nitish Srivastava et al.).
#
# <img src="images/Dropout1.png" alt="A figure from the article showing how neurons go off with dropout" width="800" id="img_dropout" caption="Applying dropout in a neural network (courtesy of Nitish Srivastava et al.)">
#
# Hinton used a nice metaphor when he explained, in an interview, the inspiration for dropout:
#
# > : I went to my bank. The tellers kept changing and I asked one of them why. He said he didn’t know but they got moved around a lot. I figured it must be because it would require cooperation between employees to successfully defraud the bank. This made me realize that randomly removing a different subset of neurons on each example would prevent conspiracies and thus reduce overfitting.
#
# In the same interview, he also explained that neuroscience provided additional inspiration:
#
# > : We don't really know why neurons spike. One theory is that they want to be noisy so as to regularize, because we have many more parameters than we have data points. The idea of dropout is that if you have noisy activations, you can afford to use a much bigger model.
# This explains the idea behind why dropout helps to generalize: first it helps the neurons to cooperate better together, then it makes the activations more noisy, thus making the model more robust.
# We can see, however, that if we were to just zero those activations without doing anything else, our model would have problems training: if we go from the sum of five activations (that are all positive numbers since we apply a ReLU) to just two, this won't have the same scale. Therefore, if we apply dropout with a probability `p`, we rescale all activations by dividing them by `1-p` (on average `p` will be zeroed, so it leaves `1-p`), as shown in <<img_dropout1>>.
#
# <img src="images/Dropout.png" alt="A figure from the article introducing dropout showing how a neuron is on/off" width="600" id="img_dropout1" caption="Why scale the activations when applying dropout (courtesy of Nitish Srivastava et al.)">
#
# This is a full implementation of the dropout layer in PyTorch (although PyTorch's native layer is actually written in C, not Python):
# + gradient={}
class Dropout(Module):
def __init__(self, p): self.p = p
def forward(self, x):
if not self.training: return x
mask = x.new(*x.shape).bernoulli_(1-p)
return x * mask.div_(1-p)
# -
# The `bernoulli_` method is creating a tensor of random zeros (with probability `p`) and ones (with probability `1-p`), which is then multiplied with our input before dividing by `1-p`. Note the use of the `training` attribute, which is available in any PyTorch `nn.Module`, and tells us if we are doing training or inference.
#
# > note: Do Your Own Experiments: In previous chapters of the book we'd be adding a code example for `bernoulli_` here, so you can see exactly how it works. But now that you know enough to do this yourself, we're going to be doing fewer and fewer examples for you, and instead expecting you to do your own experiments to see how things work. In this case, you'll see in the end-of-chapter questionnaire that we're asking you to experiment with `bernoulli_`—but don't wait for us to ask you to experiment to develop your understanding of the code we're studying; go ahead and do it anyway!
#
# Using dropout before passing the output of our LSTM to the final layer will help reduce overfitting. Dropout is also used in many other models, including the default CNN head used in `fastai.vision`, and is available in `fastai.tabular` by passing the `ps` parameter (where each "p" is passed to each added `Dropout` layer), as we'll see in <<chapter_arch_details>>.
# Dropout has different behavior in training and validation mode, which we specified using the `training` attribute in `Dropout`. Calling the `train` method on a `Module` sets `training` to `True` (both for the module you call the method on and for every module it recursively contains), and `eval` sets it to `False`. This is done automatically when calling the methods of `Learner`, but if you are not using that class, remember to switch from one to the other as needed.
# ### Activation Regularization and Temporal Activation Regularization
# *Activation regularization* (AR) and *temporal activation regularization* (TAR) are two regularization methods very similar to weight decay, discussed in <<chapter_collab>>. When applying weight decay, we add a small penalty to the loss that aims at making the weights as small as possible. For activation regularization, it's the final activations produced by the LSTM that we will try to make as small as possible, instead of the weights.
#
# To regularize the final activations, we have to store those somewhere, then add the means of the squares of them to the loss (along with a multiplier `alpha`, which is just like `wd` for weight decay):
#
# ``` python
# loss += alpha * activations.pow(2).mean()
# ```
# Temporal activation regularization is linked to the fact we are predicting tokens in a sentence. That means it's likely that the outputs of our LSTMs should somewhat make sense when we read them in order. TAR is there to encourage that behavior by adding a penalty to the loss to make the difference between two consecutive activations as small as possible: our activations tensor has a shape `bs x sl x n_hid`, and we read consecutive activations on the sequence length axis (the dimension in the middle). With this, TAR can be expressed as:
#
# ``` python
# loss += beta * (activations[:,1:] - activations[:,:-1]).pow(2).mean()
# ```
#
# `alpha` and `beta` are then two hyperparameters to tune. To make this work, we need our model with dropout to return three things: the proper output, the activations of the LSTM pre-dropout, and the activations of the LSTM post-dropout. AR is often applied on the dropped-out activations (to not penalize the activations we turned into zeros afterward) while TAR is applied on the non-dropped-out activations (because those zeros create big differences between two consecutive time steps). There is then a callback called `RNNRegularizer` that will apply this regularization for us.
# ### Training a Weight-Tied Regularized LSTM
# We can combine dropout (applied before we go into our output layer) with AR and TAR to train our previous LSTM. We just need to return three things instead of one: the normal output of our LSTM, the dropped-out activations, and the activations from our LSTMs. The last two will be picked up by the callback `RNNRegularization` for the contributions it has to make to the loss.
#
# Another useful trick we can add from [the AWD LSTM paper](https://arxiv.org/abs/1708.02182) is *weight tying*. In a language model, the input embeddings represent a mapping from English words to activations, and the output hidden layer represents a mapping from activations to English words. We might expect, intuitively, that these mappings could be the same. We can represent this in PyTorch by assigning the same weight matrix to each of these layers:
#
# self.h_o.weight = self.i_h.weight
#
# In `LMModel7`, we include these final tweaks:
# + gradient={}
class LMModel7(Module):
def __init__(self, vocab_sz, n_hidden, n_layers, p):
self.i_h = nn.Embedding(vocab_sz, n_hidden)
self.rnn = nn.LSTM(n_hidden, n_hidden, n_layers, batch_first=True)
self.drop = nn.Dropout(p)
self.h_o = nn.Linear(n_hidden, vocab_sz)
self.h_o.weight = self.i_h.weight
self.h = [torch.zeros(n_layers, bs, n_hidden) for _ in range(2)]
def forward(self, x):
raw,h = self.rnn(self.i_h(x), self.h)
out = self.drop(raw)
self.h = [h_.detach() for h_ in h]
return self.h_o(out),raw,out
def reset(self):
for h in self.h: h.zero_()
# -
# We can create a regularized `Learner` using the `RNNRegularizer` callback:
# + gradient={}
learn = Learner(dls, LMModel7(len(vocab), 64, 2, 0.5),
loss_func=CrossEntropyLossFlat(), metrics=accuracy,
cbs=[ModelResetter, RNNRegularizer(alpha=2, beta=1)])
# -
# A `TextLearner` automatically adds those two callbacks for us (with those values for `alpha` and `beta` as defaults), so we can simplify the preceding line to:
# + gradient={}
learn = TextLearner(dls, LMModel7(len(vocab), 64, 2, 0.4),
loss_func=CrossEntropyLossFlat(), metrics=accuracy)
# -
# We can then train the model, and add additional regularization by increasing the weight decay to `0.1`:
# + gradient={}
learn.fit_one_cycle(15, 1e-2, wd=0.1)
# -
# Now this is far better than our previous model!
# ## Conclusion
# You have now seen everything that is inside the AWD-LSTM architecture we used in text classification in <<chapter_nlp>>. It uses dropout in a lot more places:
#
# - Embedding dropout (inside the embedding layer, drops some random lines of embeddings)
# - Input dropout (applied after the embedding layer)
# - Weight dropout (applied to the weights of the LSTM at each training step)
# - Hidden dropout (applied to the hidden state between two layers)
#
# This makes it even more regularized. Since fine-tuning those five dropout values (including the dropout before the output layer) is complicated, we have determined good defaults and allow the magnitude of dropout to be tuned overall with the `drop_mult` parameter you saw in that chapter (which is multiplied by each dropout).
#
# Another architecture that is very powerful, especially in "sequence-to-sequence" problems (that is, problems where the dependent variable is itself a variable-length sequence, such as language translation), is the Transformers architecture. You can find it in a bonus chapter on the [book's website](https://book.fast.ai/).
# ## Questionnaire
# 1. If the dataset for your project is so big and complicated that working with it takes a significant amount of time, what should you do?
# 1. Why do we concatenate the documents in our dataset before creating a language model?
# 1. To use a standard fully connected network to predict the fourth word given the previous three words, what two tweaks do we need to make to our model?
# 1. How can we share a weight matrix across multiple layers in PyTorch?
# 1. Write a module that predicts the third word given the previous two words of a sentence, without peeking.
# 1. What is a recurrent neural network?
# 1. What is "hidden state"?
# 1. What is the equivalent of hidden state in ` LMModel1`?
# 1. To maintain the state in an RNN, why is it important to pass the text to the model in order?
# 1. What is an "unrolled" representation of an RNN?
# 1. Why can maintaining the hidden state in an RNN lead to memory and performance problems? How do we fix this problem?
# 1. What is "BPTT"?
# 1. Write code to print out the first few batches of the validation set, including converting the token IDs back into English strings, as we showed for batches of IMDb data in <<chapter_nlp>>.
# 1. What does the `ModelResetter` callback do? Why do we need it?
# 1. What are the downsides of predicting just one output word for each three input words?
# 1. Why do we need a custom loss function for `LMModel4`?
# 1. Why is the training of `LMModel4` unstable?
# 1. In the unrolled representation, we can see that a recurrent neural network actually has many layers. So why do we need to stack RNNs to get better results?
# 1. Draw a representation of a stacked (multilayer) RNN.
# 1. Why should we get better results in an RNN if we call `detach` less often? Why might this not happen in practice with a simple RNN?
# 1. Why can a deep network result in very large or very small activations? Why does this matter?
# 1. In a computer's floating-point representation of numbers, which numbers are the most precise?
# 1. Why do vanishing gradients prevent training?
# 1. Why does it help to have two hidden states in the LSTM architecture? What is the purpose of each one?
# 1. What are these two states called in an LSTM?
# 1. What is tanh, and how is it related to sigmoid?
# 1. What is the purpose of this code in `LSTMCell`: `h = torch.cat([h, input], dim=1)`
# 1. What does `chunk` do in PyTorch?
# 1. Study the refactored version of `LSTMCell` carefully to ensure you understand how and why it does the same thing as the non-refactored version.
# 1. Why can we use a higher learning rate for `LMModel6`?
# 1. What are the three regularization techniques used in an AWD-LSTM model?
# 1. What is "dropout"?
# 1. Why do we scale the acitvations with dropout? Is this applied during training, inference, or both?
# 1. What is the purpose of this line from `Dropout`: `if not self.training: return x`
# 1. Experiment with `bernoulli_` to understand how it works.
# 1. How do you set your model in training mode in PyTorch? In evaluation mode?
# 1. Write the equation for activation regularization (in math or code, as you prefer). How is it different from weight decay?
# 1. Write the equation for temporal activation regularization (in math or code, as you prefer). Why wouldn't we use this for computer vision problems?
# 1. What is "weight tying" in a language model?
# ### Further Research
# 1. In ` LMModel2`, why can `forward` start with `h=0`? Why don't we need to say `h=torch.zeros(...)`?
# 1. Write the code for an LSTM from scratch (you may refer to <<lstm>>).
# 1. Search the internet for the GRU architecture and implement it from scratch, and try training a model. See if you can get results similar to those we saw in this chapter. Compare your results to the results of PyTorch's built in `GRU` module.
# 1. Take a look at the source code for AWD-LSTM in fastai, and try to map each of the lines of code to the concepts shown in this chapter.
# + gradient={}
| 12_nlp_dive.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://raw.githubusercontent.com/israeldi/quantlab/master/assets/images/Program-Logo.png" width="400px" align="right">
#
# # Binomial Pricing Model
# ### [(Go to Quant Lab)](https://israeldi.github.io/quantlab/)
#
# #### Source: Numerical Methods in Finance with C++
#
# © <NAME>, <NAME>
#
# <img src="https://raw.githubusercontent.com/israeldi/quantlab/master/assets/images/Numerical_Methods.jpg" width="200px" align="left">
# ## Table of Contents
#
# 1. [Random Numbers](#1.-Random-Numbers)
# 2. [Plotting Random Samples](#2.-Plotting-Random-Samples)
# 3. [Simulation](#3.-Simulation)
# - 3.1 [Random Variables](#3.1-Random-Variables)
# - 3.2 [Stochastic Processes](#3.2-Stochastic-Processes)
# - 3.2.1 [Geometric Brownian Motion](#3.2.1-Geometric-Brownian-Motion)
# - 3.2.2 [Square-Root Diffusion](#3.2.2-Square-Root-Diffusion)
# - 3.2.3 [Stochastic Processes](#3.2-Stochastic-Processes)
# - 3.2.4 [Stochastic Processes](#3.2-Stochastic-Processes)
# - 3.3 [Variance Reduction](#3.3-Variance-Reduction)
# 4. [Valuation](#4.-Valuation)
# - 4.1 [European Options](#4.1-European-Options)
# - 4.2 [American Options](#4.2-American-Options)
# 5. [Risk Measures](#5.-Risk-Measures)
# - 5.1 [Value-at-Risk](#5.1-Value-at-Risk)
# - 5.2 [Credit Value Adjustments](#5.2-Credit-Value-Adjustments)
#
# Initially import all the modules we will be using for our notebook
import math
import numpy as np
import numpy.random as npr
# from pylab import plt, mpl
import sys
import os
# ## 1.1 Creating Main Program
if __name__== "__main__":
print("Hi there")
input("Provide a character: ")
# Directory where we will save our plots
directory = "./images"
if not os.path.exists(directory):
os.makedirs(directory)
# ## 1.2 Entering Data
# #### ([Back to Top](#Table-of-Contents))
if __name__== "__main__":
S0 = float(input("Enter S_0: "))
U = float(input("Enter U: "))
D = float(input("Enter D: "))
R = float(input("Enter R: "))
# making sure that 0<S0, -1<D<U, -1<R
if (S0 <= 0.0 or U <= -1.0 or D <= -1.0 or U <= D or R <= -1.0):
print("Illegal data ranges")
print("Terminating program")
sys.exit()
# checking for arbitrage
if (R >= U or R <= D):
print("Arbitrage exists")
print("Terminating program")
sys.exit()
print("Input data checked")
print("There is no arbitrage\n")
# compute risk-neutral probability
print("q = ", (R - D) / (U - D))
# compute stock price at node n=3,i=2
n = 3; i = 2
print("n = ", n)
print("i = ", i)
print("S(n,i) = ", S0* math.pow(1 + U,i) * math.pow(1 + D, n - i))
# ## 1.3 Functions
# #### ([Back to Top](#Table-of-Contents))
# + uuid="4618b170-6bd3-4500-905a-0fe402f198c1"
# computing risk-neutral probability
def RiskNeutProb(U, D, R):
return (R - D) / (U - D)
# computing the stock price at node n,i
def S(S0, U, D, n, i):
return S0 * math.pow(1 + U,i) * math.pow(1 + D, n - i)
def isValidInput(S0, U, D, R):
# making sure that 0<S0, -1<D<U, -1<R
if (S0 <= 0.0 or U <= -1.0 or D <= -1.0 or U <= D or R <= -1.0):
print("Illegal data ranges")
print("Terminating program")
return 0
# checking for arbitrage
if (R >= U or R <= D):
print("Arbitrage exists")
print("Terminating program")
return 0
return 1
# inputting, displaying and checking model data
def GetInputData():
#entering data
params = ("S0", "U", "D", "R")
S0, U, D, R = [float(input("Enter %s: " % (var))) for var in params]
if not isValidInput(S0, U, D, R):
return 0
print("Input data checked")
print("There is no arbitrage\n")
d = locals().copy()
return d
if __name__== "__main__":
# compute risk-neutral probability
print("q = ", RiskNeutProb(U, D, R))
output = GetInputData()
if output == 0:
sys.exit()
# Update our parameters
locals().update(output)
# compute stock price at node n=3,i=2
n = 3; i = 2
print("n = ", n)
print("i = ", i)
print("S(n,i) = ", S(S0,U,D,n,i))
# -
# ## 2.0 Object Oriented European and American
# #### ([Back to Top](#Table-of-Contents))
# +
# Binomial Model (Parent class) ---------------------------------------------------
class BinModel:
def __init__(self, S0=100, U=0.1, D=-0.1, R=0):
self.S0 = S0
self.U = U
self.D = D
self.R = R
def RiskNeutProb(self):
return (self.R - self.D) / (self.U - self.D)
def S(self, n, i):
return self.S0 * math.pow(1 + self.U, i) * math.pow(1 + self.D, n - i)
# European Option class ------------------------------------------------------------------
class EurOption():
def PriceByCRR(self, Model, N=3):
q = Model.RiskNeutProb();
Price = np.zeros(N + 1)
for i in range(0, N + 1):
Price[i] = self.Payoff(Model.S(N, i))
for n in range(N - 1, -1, -1):
for i in range(0, n + 1):
Price[i] = (q * Price[i+1] + (1 - q) * Price[i]) / (1 + Model.R);
return Price[0]
# American Option class
class AmOption():
def PriceBySnell(self, Model, N=3):
q=Model.RiskNeutProb()
Price = np.zeros(N + 1)
for i in range(0, N + 1):
Price[i] = self.Payoff(Model.S(N, i))
for n in range(N - 1, -1, -1):
for i in range(0, n + 1):
ContVal = (q * Price[i+1] + (1 - q) * Price[i]) / (1 + Model.R)
Price[i] = self.Payoff(Model.S(n,i))
if (ContVal > Price[i]):
Price[i] = ContVal
return Price[0]
# Payoff Classes ----------------------------------------------------------------------------
class Call(EurOption, AmOption):
def __init__(self, K=1):
self.K = K
def Payoff(self, z):
if (z > self.K):
return z - self.K
return 0
class Put(EurOption, AmOption):
def __init__(self, K=1):
self.K = K
def Payoff(self, z):
if (z < self.K):
return self.K - z
return 0
model = BinModel(S0=10, U=0.2, D=-0.1, R=0.1)
Option1 = Call(K=22)
Option1.PriceByCRR(model, N=6)
# -
# ## Exercises:
#
# 1. Modify the `PriceByCRR()` function in `EurOption` Class to compute the time $0$ price of a European option using the **Cox–Ross–Rubinstein (CRR)** formula:
# $$H(0)=\frac{1}{(1+R)^{N}}\sum_{i=0}^{N}\frac{N!}{i!(N-i)!}q^{i}(1-q)^{N-i}h(S(N,I))$$
#
#
# 2. The payoff of a **digital call** with strike price $K$ is:
# $$h^{digit\thinspace call}(z)=\begin{cases}
# 1 & \textrm{{if} }K<z,\\
# 0 & \textrm{otherwise.}
# \end{cases}$$
# Include the ability to price digital calls in the program developed in the present section by adding the new payoff Class `DigitCall` just as was done for calls and puts.
#
#
# 3. Add the ability to price bull spreads and bear spreads by introducing new subclasses `BullSpread` and `BearSpread` of the `EurOption` and `AmOption` classes defined in our current program. The payoffs of a bull spread and a bear spread, which depend on two parameters $K1 < K2$, are given by:
# $$h^{bull}(z)=\begin{cases}
# 0 & \textrm{if }z\leq K_{1},\\
# z-K_{1} & \textrm{if }K_{1}<z<K_{2}\\
# K_{2}-K_{1} & \textrm{if }K_{2}\leq z,
# \end{cases}$$ and
# $$h^{bear}(z)=\begin{cases}
# K_{2}-K_{1} & \textrm{if }z\leq K_{1},\\
# K_{2}-z & \textrm{if }K_{1}<z<K_{2}\\
# 0 & \textrm{if }K_{2}<z,
# \end{cases}$$
#
#
# 4. Consider the **fixed-strike** Asian call option with process,
# $$A_{N}=\frac{1}{N}\sum_{k=0}^{N-1}S(k)$$ and payoff, $$C(N)=max\{A_{N},0\}$$
# Add a new Class `AsianOption` to price an Asian style Call Option.
| files/Bootcamp/.ipynb_checkpoints/discrete_finance-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Winpython Default checker
import warnings
#warnings.filterwarnings("ignore", category=DeprecationWarning)
#warnings.filterwarnings("ignore", category=UserWarning)
#warnings.filterwarnings("ignore", category=FutureWarning)
# warnings.filterwarnings("ignore") # would silence all warnings
# %matplotlib inline
# use %matplotlib widget for the adventurous
# ## Compilers: Numba and Cython
#
# ##### Requirement
# To get Cython working, Winpython 3.7+ users should install "Microsoft Visual C++ Build Tools 2017" (visualcppbuildtools_full.exe, a 4 Go installation) at https://beta.visualstudio.com/download-visual-studio-vs/
#
# To get Numba working, not-windows10 users may have to install "Microsoft Visual C++ Redistributable pour Visual Studio 2017" (vc_redist) at <https://beta.visualstudio.com/download-visual-studio-vs/>
#
# Thanks to recent progress, Visual Studio 2017/2018/2019 are cross-compatible now
#
# #### Compiler toolchains
# ##### Numba (a JIT Compiler)
# +
# checking Numba JIT toolchain
import numpy as np
image = np.zeros((1024, 1536), dtype = np.uint8)
#from pylab import imshow, show
import matplotlib.pyplot as plt
from timeit import default_timer as timer
from numba import jit
@jit
def create_fractal(min_x, max_x, min_y, max_y, image, iters , mandelx):
height = image.shape[0]
width = image.shape[1]
pixel_size_x = (max_x - min_x) / width
pixel_size_y = (max_y - min_y) / height
for x in range(width):
real = min_x + x * pixel_size_x
for y in range(height):
imag = min_y + y * pixel_size_y
color = mandelx(real, imag, iters)
image[y, x] = color
@jit
def mandel(x, y, max_iters):
c = complex(x, y)
z = 0.0j
for i in range(max_iters):
z = z*z + c
if (z.real*z.real + z.imag*z.imag) >= 4:
return i
return max_iters
# +
# Numba speed
start = timer()
create_fractal(-2.0, 1.0, -1.0, 1.0, image, 20 , mandel)
dt = timer() - start
fig = plt.figure()
print ("Mandelbrot created by numba in %f s" % dt)
plt.imshow(image)
plt.show()
# -
# ##### Cython (a compiler for writing C extensions for the Python language)
# WinPython 3.5 and 3.6 users may not have mingwpy available, and so need "VisualStudio C++ Community Edition 2015" https://www.visualstudio.com/downloads/download-visual-studio-vs#d-visual-c
# Cython + Mingwpy compiler toolchain test
# %load_ext Cython
# + magic_args="-a" language="cython"
# # with %%cython -a , full C-speed lines are shown in white, slowest python-speed lines are shown in dark yellow lines
# # ==> put your cython rewrite effort on dark yellow lines
# def create_fractal_cython(min_x, max_x, min_y, max_y, image, iters , mandelx):
# height = image.shape[0]
# width = image.shape[1]
# pixel_size_x = (max_x - min_x) / width
# pixel_size_y = (max_y - min_y) / height
#
# for x in range(width):
# real = min_x + x * pixel_size_x
# for y in range(height):
# imag = min_y + y * pixel_size_y
# color = mandelx(real, imag, iters)
# image[y, x] = color
#
# def mandel_cython(x, y, max_iters):
# cdef int i
# cdef double cx, cy , zx, zy
# cx , cy = x, y
# zx , zy =0 ,0
# for i in range(max_iters):
# zx , zy = zx*zx - zy*zy + cx , zx*zy*2 + cy
# if (zx*zx + zy*zy) >= 4:
# return i
# return max_iters
# +
#Cython speed
start = timer()
create_fractal_cython(-2.0, 1.0, -1.0, 1.0, image, 20 , mandel_cython)
dt = timer() - start
fig = plt.figure()
print ("Mandelbrot created by cython in %f s" % dt)
plt.imshow(image)
# -
# ## Graphics: Matplotlib, Pandas, Seaborn, Holoviews, Bokeh, bqplot, ipyleaflet, plotnine
# +
# Matplotlib 3.4.1
# for more examples, see: http://matplotlib.org/gallery.html
from mpl_toolkits.mplot3d import axes3d
import matplotlib.pyplot as plt
from matplotlib import cm
ax = plt.figure().add_subplot(projection='3d')
X, Y, Z = axes3d.get_test_data(0.05)
# Plot the 3D surface
ax.plot_surface(X, Y, Z, rstride=8, cstride=8, alpha=0.3)
# Plot projections of the contours for each dimension. By choosing offsets
# that match the appropriate axes limits, the projected contours will sit on
# the 'walls' of the graph
cset = ax.contourf(X, Y, Z, zdir='z', offset=-100, cmap=cm.coolwarm)
cset = ax.contourf(X, Y, Z, zdir='x', offset=-40, cmap=cm.coolwarm)
cset = ax.contourf(X, Y, Z, zdir='y', offset=40, cmap=cm.coolwarm)
ax.set_xlim(-40, 40)
ax.set_ylim(-40, 40)
ax.set_zlim(-100, 100)
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_zlabel('Z')
plt.show()
# -
# Seaborn
# for more examples, see http://stanford.edu/~mwaskom/software/seaborn/examples/index.html
import seaborn as sns
sns.set()
df = sns.load_dataset("iris")
sns.pairplot(df, hue="species", height=1.5)
# +
# altair-example
import altair as alt
alt.Chart(df).mark_bar().encode(
x=alt.X('sepal_length', bin=alt.Bin(maxbins=50)),
y='count(*):Q',
color='species:N',
#column='species',
).interactive()
# +
# temporary warning removal
import warnings
import matplotlib as mpl
warnings.filterwarnings("ignore", category=mpl.cbook.MatplotlibDeprecationWarning)
# Holoviews
# for more example, see http://holoviews.org/Tutorials/index.html
import numpy as np
import holoviews as hv
hv.extension('matplotlib')
dots = np.linspace(-0.45, 0.45, 11)
fractal = hv.Image(image)
layouts = {y: (fractal * hv.Points(fractal.sample([(i,y) for i in dots])) +
fractal.sample(y=y) )
for y in np.linspace(0, 0.45,11)}
hv.HoloMap(layouts, kdims=['Y']).collate().cols(2)
# +
# Bokeh 0.12.5
import numpy as np
from six.moves import zip
from bokeh.plotting import figure, show, output_notebook
N = 4000
x = np.random.random(size=N) * 100
y = np.random.random(size=N) * 100
radii = np.random.random(size=N) * 1.5
colors = ["#%02x%02x%02x" % (int(r), int(g), 150) for r, g in zip(50+2*x, 30+2*y)]
output_notebook()
TOOLS="hover,crosshair,pan,wheel_zoom,box_zoom,reset,tap,save,box_select,poly_select,lasso_select"
p = figure(tools=TOOLS)
p.scatter(x,y, radius=radii, fill_color=colors, fill_alpha=0.6, line_color=None)
show(p)
# +
# Datashader (holoviews+Bokeh)
import datashader as ds
import numpy as np
import holoviews as hv
from holoviews import opts
from holoviews.operation.datashader import datashade, shade, dynspread, spread, rasterize
from holoviews.operation import decimate
hv.extension('bokeh')
decimate.max_samples=1000
dynspread.max_px=20
dynspread.threshold=0.5
def random_walk(n, f=5000):
"""Random walk in a 2D space, smoothed with a filter of length f"""
xs = np.convolve(np.random.normal(0, 0.1, size=n), np.ones(f)/f).cumsum()
ys = np.convolve(np.random.normal(0, 0.1, size=n), np.ones(f)/f).cumsum()
xs += 0.1*np.sin(0.1*np.array(range(n-1+f))) # add wobble on x axis
xs += np.random.normal(0, 0.005, size=n-1+f) # add measurement noise
ys += np.random.normal(0, 0.005, size=n-1+f)
return np.column_stack([xs, ys])
def random_cov():
"""Random covariance for use in generating 2D Gaussian distributions"""
A = np.random.randn(2,2)
return np.dot(A, A.T)
# -
np.random.seed(1)
points = hv.Points(np.random.multivariate_normal((0,0), [[0.1, 0.1], [0.1, 1.0]], (50000,)),label="Points")
paths = hv.Path([0.15*random_walk(10000) for i in range(10)], kdims=["u","v"], label="Paths")
decimate(points) + rasterize(points) + rasterize(paths)
ropts = dict(colorbar=True, tools=["hover"], width=350)
rasterize( points).opts(cmap="kbc_r", cnorm="linear").relabel('rasterize()').opts(**ropts).hist() + \
dynspread(datashade( points, cmap="kbc_r", cnorm="linear").relabel("datashade()"))
#bqplot
from IPython.display import display
from bqplot import (Figure, Map, Mercator, Orthographic, ColorScale, ColorAxis,
AlbersUSA, topo_load, Tooltip)
def_tt = Tooltip(fields=['id', 'name'])
map_mark = Map(scales={'projection': Mercator()}, tooltip=def_tt)
map_mark.interactions = {'click': 'select', 'hover': 'tooltip'}
fig = Figure(marks=[map_mark], title='Interactions Example')
display(fig)
# +
# ipyleaflet (javascript library usage)
from ipyleaflet import (
Map, Marker, TileLayer, ImageOverlay, Polyline, Polygon,
Rectangle, Circle, CircleMarker, GeoJSON, DrawControl
)
from traitlets import link
center = [34.6252978589571, -77.34580993652344]
m = Map(center=[34.6252978589571, -77.34580993652344], zoom=10)
dc = DrawControl()
def handle_draw(self, action, geo_json):
print(action)
print(geo_json)
m
m
# -
dc.on_draw(handle_draw)
m.add_control(dc)
# +
# %matplotlib widget
# Testing matplotlib interactions with a simple plot
import matplotlib.pyplot as plt
import numpy as np
# warning ; you need to launch a second time %matplotlib widget, if after a %matplotlib inline
# %matplotlib widget
fig = plt.figure() #plt.figure(1)
plt.plot(np.sin(np.linspace(0, 20, 100)))
plt.show()
# -
# plotnine: giving a taste of ggplot of R langage (formerly we were using ggpy)
from plotnine import ggplot, aes, geom_blank, geom_point, stat_smooth, facet_wrap, theme_bw
from plotnine.data import mtcars
ggplot(mtcars, aes(x='hp', y='wt', color='mpg')) + geom_point() +\
facet_wrap("~cyl") + theme_bw()
# ## Ipython Notebook: Interactivity & other
import IPython;IPython.__version__
# Audio Example : https://github.com/ipython/ipywidgets/blob/master/examples/Beat%20Frequencies.ipynb
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from ipywidgets import interactive
from IPython.display import Audio, display
def beat_freq(f1=220.0, f2=224.0):
max_time = 3
rate = 8000
times = np.linspace(0,max_time,rate*max_time)
signal = np.sin(2*np.pi*f1*times) + np.sin(2*np.pi*f2*times)
print(f1, f2, abs(f1-f2))
display(Audio(data=signal, rate=rate))
try:
plt.plot(signal); #plt.plot(v.result);
except:
pass
return signal
v = interactive(beat_freq, f1=(200.0,300.0), f2=(200.0,300.0))
display(v)
# +
# Networks graph Example : https://github.com/ipython/ipywidgets/blob/master/examples/Exploring%20Graphs.ipynb
# %matplotlib inline
from ipywidgets import interact
import matplotlib.pyplot as plt
import networkx as nx
# wrap a few graph generation functions so they have the same signature
def random_lobster(n, m, k, p):
return nx.random_lobster(n, p, p / m)
def powerlaw_cluster(n, m, k, p):
return nx.powerlaw_cluster_graph(n, m, p)
def erdos_renyi(n, m, k, p):
return nx.erdos_renyi_graph(n, p)
def newman_watts_strogatz(n, m, k, p):
return nx.newman_watts_strogatz_graph(n, k, p)
@interact(n=(2,30), m=(1,10), k=(1,10), p=(0.0, 1.0, 0.001),
generator={'lobster': random_lobster,
'power law': powerlaw_cluster,
'Newman-Watts-Strogatz': newman_watts_strogatz,
u'Erdős-Rényi': erdos_renyi,
})
def plot_random_graph(n, m, k, p, generator):
g = generator(n, m, k, p)
nx.draw(g)
plt.title(generator.__name__)
plt.show()
# -
# ## Mathematical: statsmodels, lmfit,
# checking statsmodels
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('ggplot')
import statsmodels.api as sm
data = sm.datasets.anes96.load_pandas()
party_ID = np.arange(7)
labels = ["Strong Democrat", "Weak Democrat", "Independent-Democrat",
"Independent-Independent", "Independent-Republican",
"Weak Republican", "Strong Republican"]
plt.rcParams['figure.subplot.bottom'] = 0.23 # keep labels visible
plt.rcParams['figure.figsize'] = (6.0, 4.0) # make plot larger in notebook
age = [data.exog['age'][data.endog == id] for id in party_ID]
fig = plt.figure()
ax = fig.add_subplot(111)
plot_opts={'cutoff_val':5, 'cutoff_type':'abs',
'label_fontsize':'small',
'label_rotation':30}
sm.graphics.beanplot(age, ax=ax, labels=labels,
plot_opts=plot_opts)
ax.set_xlabel("Party identification of respondent")
ax.set_ylabel("Age")
plt.show()
# +
# lmfit test (from http://nbviewer.ipython.org/github/lmfit/lmfit-py/blob/master/examples/lmfit-model.ipynb)
import numpy as np
import matplotlib.pyplot as plt
def decay(t, N, tau):
return N*np.exp(-t/tau)
t = np.linspace(0, 5, num=1000)
data = decay(t, 7, 3) + np.random.randn(*t.shape)
from lmfit import Model
model = Model(decay, independent_vars=['t'])
result = model.fit(data, t=t, N=10, tau=1)
fig = plt.figure() # necessary to separate from previous ploot with %matplotlib widget
plt.plot(t, data) # data
plt.plot(t, decay(t=t, **result.values), color='orange', linewidth=5) # best-fit model
# -
# ## DataFrames: Pandas, Dask
# +
#Pandas
import pandas as pd
import numpy as np
idx = pd.date_range('2000', '2005', freq='d', closed='left')
datas = pd.DataFrame({'Color': [ 'green' if x> 1 else 'red' for x in np.random.randn(len(idx))],
'Measure': np.random.randn(len(idx)), 'Year': idx.year},
index=idx.date)
datas.head()
# -
# ### Split / Apply / Combine
# Split your data into multiple independent groups.
# Apply some function to each group.
# Combine your groups back into a single data object.
#
datas.query('Measure > 0').groupby(['Color','Year']).size().unstack()
# ## Web Scraping: Beautifulsoup
# +
# checking Web Scraping: beautifulsoup and requests
import requests
from bs4 import BeautifulSoup
URL = 'http://en.wikipedia.org/wiki/Franklin,_Tennessee'
req = requests.get(URL, headers={'User-Agent' : "Mining the Social Web"})
soup = BeautifulSoup(req.text, "lxml")
geoTag = soup.find(True, 'geo')
if geoTag and len(geoTag) > 1:
lat = geoTag.find(True, 'latitude').string
lon = geoTag.find(True, 'longitude').string
print ('Location is at', lat, lon)
elif geoTag and len(geoTag) == 1:
(lat, lon) = geoTag.string.split(';')
(lat, lon) = (lat.strip(), lon.strip())
print ('Location is at', lat, lon)
else:
print ('No location found')
# -
# ## Operations Research: Pulp
# +
# Pulp example : minimizing the weight to carry 99 pennies
# (from <NAME>)
# see https://www.youtube.com/watch?v=UmMn-N5w-lI#t=995
# Import PuLP modeler functions
from pulp import *
# The prob variable is created to contain the problem data
prob = LpProblem("99_pennies_Problem",LpMinimize)
# Variables represent how many of each coin we want to carry
pennies = LpVariable("Number_of_pennies",0,None,LpInteger)
nickels = LpVariable("Number_of_nickels",0,None,LpInteger)
dimes = LpVariable("Number_of_dimes",0,None,LpInteger)
quarters = LpVariable("Number_of_quarters",0,None,LpInteger)
# The objective function is added to 'prob' first
# we want to minimize (LpMinimize) this
prob += 2.5 * pennies + 5 * nickels + 2.268 * dimes + 5.670 * quarters, "Total_coins_Weight"
# We want exactly 99 cents
prob += 1 * pennies + 5 * nickels + 10 * dimes + 25 * quarters == 99, ""
# The problem data is written to an .lp file
prob.writeLP("99cents.lp")
prob.solve()
# print ("status",LpStatus[prob.status] )
print ("Minimal Weight to carry exactly 99 pennies is %s grams" % value(prob.objective))
# Each of the variables is printed with it's resolved optimum value
for v in prob.variables():
print (v.name, "=", v.varValue)
# -
# ## Deep Learning: see tutorial-first-neural-network-python-keras
# ## Symbolic Calculation: sympy
# checking sympy
import sympy
a, b =sympy.symbols('a b')
e=(a+b)**5
e.expand()
# ## SQL tools: sqlite, Ipython-sql, sqlite_bro, baresql, db.py
# checking Ipython-sql, sqlparse, SQLalchemy
# %load_ext sql
# + magic_args="sqlite:///.baresql.db" language="sql"
# DROP TABLE IF EXISTS writer;
# CREATE TABLE writer (first_name, last_name, year_of_death);
# INSERT INTO writer VALUES ('William', 'Shakespeare', 1616);
# INSERT INTO writer VALUES ('Bertold', 'Brecht', 1956);
# SELECT * , sqlite_version() as sqlite_version from Writer order by Year_of_death
# +
# checking baresql
from __future__ import print_function, unicode_literals, division # line needed only if Python2.7
from baresql import baresql
bsql = baresql.baresql(connection="sqlite:///.baresql.db")
bsqldf = lambda q: bsql.df(q, dict(globals(),**locals()))
users = ['Alexander', 'Billy', 'Charles', 'Danielle', 'Esmeralda', 'Franz', 'Greg']
# We use the python 'users' list like a SQL table
sql = "select 'Welcome ' || c0 || ' !' as say_hello, length(c0) as name_length from users$$ where c0 like '%a%' "
bsqldf(sql)
# -
# Transfering Datas to sqlite, doing transformation in sql, going back to Pandas and Matplotlib
bsqldf('''
select Color, Year, count(*) as size
from datas$$
where Measure > 0
group by Color, Year'''
).set_index(['Year', 'Color']).unstack().plot(kind='bar')
# checking db.py
from db import DB
db=DB(dbtype="sqlite", filename=".baresql.db")
db.query("select sqlite_version() as sqlite_version ;")
db.tables
# checking sqlite_bro: this should lanch a separate non-browser window with sqlite_bro's welcome
# !cmd start cmd /C sqlite_bro
# +
# pyodbc or pypyodbc or ceODBC
try:
import pyodbc
except ImportError:
import pypyodbc as pyodbc # on PyPy, there is no pyodbc currently
# look for pyodbc providers
sources = pyodbc.dataSources()
dsns = list(sources.keys())
sl = [' %s [%s]' % (dsn, sources[dsn]) for dsn in dsns]
print("pyodbc Providers: (beware 32/64 bit driver and python version must match)\n", '\n'.join(sl))
# +
# pythonnet
import clr
clr.AddReference("System.Data")
clr.AddReference('System.Data.Common')
import System.Data.OleDb as ADONET
import System.Data.Odbc as ODBCNET
import System.Data.Common as DATACOM
table = DATACOM.DbProviderFactories.GetFactoryClasses()
print("\n .NET Providers: (beware 32/64 bit driver and python version must match)")
for row in table.Rows:
print(" %s" % row[table.Columns[0]])
print(" ",[row[column] for column in table.Columns if column != table.Columns[0]])
# -
# ## Qt libraries Demo
#
#
# #### See [Dedicated Qt Libraries Demo](Qt_libraries_demo.ipynb)
# ## Wrap-up
# +
# optional scipy full test (takes up to 10 minutes)
# #!cmd /C start cmd /k python.exe -c "import scipy;scipy.test()"
# -
# %pip list
# !jupyter labextension list
# !pip check
# !pipdeptree
# !pipdeptree -p pip
| docs/Winpython_checker.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from datascience import *
# +
### COVID-19 PREDICTION TOOL ###
### Developer: <NAME> ###
### Sources: CSV file taken from https://github.com/nshomron/covidpred/blob/master/data/corona_tested_individuals_ver_006.english.csv.zip ###
### CSV file originally sourced from: https://data.gov.il/dataset/covid-19 ###
#--------------------------------------------------------------------------------------------------------------------------------------------#
# To classify a person, create a cell at the bottom of the notebook and run classify(<information list>) #
# The information list is a list containing 0s and 1s, corresponding to No/ Yes, in the following order: #
# cough || fever || sore_throat || shortness_of_breath || head_ache || age_60_and_above || sex(1: Male, 0: Female) #
# For instance, if a female under the age of 60 has cough, sore throat, and head ache, they would be classified using: #
# classify([1, 0, 1, 0, 1, 0, 0]) #
# +
# CSV file containing test-results is converted to a table. #
raw_data = Table.read_table("COVID_DataSet.csv")
raw_data.show(10)
# -
# Unnecessary columns are first dropped. #
filtered = raw_data.drop("test_date", "test_indication")
filtered.show(10)
# The data are now randomly shuffled precautionarily, to remove potential systematic ordering. #
shuffled = filtered.sample(with_replacement=False)
shuffled.show(10)
# 50,000 data points are initially chosen to train the classifier #
shortened = shuffled.take(range(50000))
# +
# Some data points have illegible values such as 'None' or 'Nan', these are filtered out. #
clean = shortened.where("age_60_and_above", are.contained_in(["No", "Yes"]))
clean = clean.where("corona_result", are.contained_in(["positive", "negative"]))
clean = clean.where("gender", are.contained_in(["female", "male"]))
clean = clean.where("cough", are.contained_in(["0", "1"]))
clean = clean.where("fever", are.contained_in(["0", "1"]))
clean = clean.where("sore_throat", are.contained_in(["0", "1"]))
clean = clean.where("shortness_of_breath", are.contained_in(["0", "1"]))
clean = clean.where("head_ache", are.contained_in(["0", "1"]))
clean.show(10)
# +
# To compute distances between various members of the population in the data sets, categorical values are converted to numerical. #
mapping_dict = {
"negative" : 0,
"positive" : 1,
"No" : 0,
"Yes" : 1,
"female" : 0,
"male" : 1
}
ages = np.array(clean.column("age_60_and_above"))
test_results = np.array(clean.column("corona_result"))
sexes = np.array(clean.column("gender"))
for i in range(clean.num_rows):
ages[i] = mapping_dict[ages[i]]
test_results[i] = mapping_dict[test_results[i]]
sexes[i] = mapping_dict[sexes[i]]
clean = clean.with_columns("corona_result", test_results, "age_60_and_above", ages, "sex", sexes).drop('gender')
# Note that the term gender has been changed to sex
clean.show(10)
# +
# Here, the data are split into a training and a test set. This ensures that the accuracy of the classifier can be deduced- #
# The classifier is trained only with the training set, and is later tested with the testing set. #
# The numbers are chosen to split the data, approximately, into two equal halves. #
test_set = clean.take(range(0, clean.num_rows // 2))
train_set = clean.take(range(clean.num_rows // 2, clean.num_rows))
# -
def table_to_int(table):
"""Returns a copy of table with all of its values converted to integers."""
cols = table.labels
new = table
for col in cols:
arr = np.array(table.column(col)).astype(int)
new = new.with_column(col, arr)
return new
# +
# This is necessary since in the original table, numerical values for symptoms (0/1) were entered as strings; #
# To compute distances between points, strings must be converted to numbers #
train_set = table_to_int(train_set)
test_set = table_to_int(test_set)
# -
def distance(new_data_point, training_data_table):
"""Returns a copy of the training data table, along with the Euclidian distances from the new data point to each point in the set."""
distances = make_array()
table_without_results = training_data_table.drop('corona_result')
for i in range(training_data_table.num_rows):
row_array = np.array(table_without_results.row(i))
distance = np.sqrt(sum((row_array - new_data_point) ** 2))
distances = np.append(distances, distance)
return training_data_table.with_column('distances', distances)
def k_nearest_neighbours(new_data_point, k):
""" Returns the 'k' nearest neighbours of a new data point in the training set """
neighbours = distance(new_data_point, train_set)
sorted_neighbours = neighbours.sort('distances',descending=False)
k_sorted_neighbours = sorted_neighbours.take(range(k))
return k_sorted_neighbours
def classify(new_data_point, k=5):
"""Returns the result (positive/negative) of classifying one new data point using 'k' nearest neighbours."""
nn = k_nearest_neighbours(new_data_point, k)
results = nn.select('corona_result')
counts = results.group('corona_result')
sorted_counts = counts.sort('count', descending=True)
result_key = sorted_counts.column(0).item(0)
if result_key == 0:
return 'NEGATIVE'
return 'POSITIVE'
def accuracy(k=5, n=10):
"""Returns the accuracy (%) of the classifier with 'k' nearest neighbours, using 'n' members of the test set."""
# Default values for k, n were chosen to accomodate lengthy processing times. #
test_set_without_results = test_set.drop('corona_result').take(range(n))
predictions = make_array()
for i in range(n):
prediction_string = classify(np.array(test_set_without_results.row(i)), k)
if prediction_string == 'NEGATIVE':
prediction_binary = 0
else:
prediction_binary = 1
predictions = np.append(predictions, prediction_binary)
comparison_table = test_set.take(range(n)).with_column('predicted_corona_result', predictions)
correct_guesses = sum(comparison_table.column('predicted_corona_result') == comparison_table.column('corona_result'))
return (correct_guesses / comparison_table.num_rows) * 100
# The classifier is 92.8% accurate when using 5 nearest neighbours with 1000 members of the testing set. #
print(accuracy(5, 1000))
| Covid19_Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # New Horizons launch and trajectory
#
# Main data source: Guo & Farquhar "New Horizons Mission Design" http://www.boulder.swri.edu/pkb/ssr/ssr-mission-design.pdf
# +
import matplotlib.pyplot as plt
plt.ion()
from astropy import time
from astropy import units as u
from poliastro.bodies import Sun, Earth, Jupiter
from poliastro.twobody import Orbit
from poliastro.plotting import plot, OrbitPlotter
from poliastro import iod
from poliastro.util import norm
# -
# ## Parking orbit
#
# Quoting from "New Horizons Mission Design":
#
# > It was first inserted into an elliptical Earth parking orbit
# of **perigee altitude 165 km** and **apogee altitude 215 km**. [Emphasis mine]
# +
r_p = Earth.R + 165 * u.km
r_a = Earth.R + 215 * u.km
a_parking = (r_p + r_a) / 2
ecc_parking = 1 - r_p / a_parking
parking = Orbit.from_classical(Earth, a_parking, ecc_parking,
0 * u.deg, 0 * u.deg, 0 * u.deg, 0 * u.deg, # We don't mind
time.Time("2006-01-19", scale='utc'))
plot(parking)
parking.v
# -
# ## Hyperbolic exit
#
# Hyperbolic excess velocity:
#
# $$ v_{\infty}^2 = \frac{\mu}{-a} = 2 \varepsilon = C_3 $$
#
# Relation between orbital velocity $v$, local escape velocity $v_e$ and hyperbolic excess velocity $v_{\infty}$:
#
# $$ v^2 = v_e^2 + v_{\infty}^2 $$
#
# ### Option a): Insert $C_3$ from report, check $v_e$ at parking perigee
# +
C_3_A = 157.6561 * u.km**2 / u.s**2 # Designed
a_exit = -(Earth.k / C_3_A).to(u.km)
ecc_exit = 1 - r_p / a_exit
exit = Orbit.from_classical(Earth, a_exit, ecc_exit,
0 * u.deg, 0 * u.deg, 0 * u.deg, 0 * u.deg, # We don't mind
time.Time("2006-01-19", scale='utc'))
norm(exit.v).to(u.km / u.s)
# -
# Quoting "New Horizons Mission Design":
#
# > After a short coast in the parking orbit, the spacecraft was then injected into
# the desired heliocentric orbit by the Centaur second stage and Star 48B third
# stage. At the Star 48B burnout, the New Horizons spacecraft reached the highest
# Earth departure speed, **estimated at 16.2 km/s**, becoming the fastest spacecraft
# ever launched from Earth. [Emphasis mine]
# +
v_estimated = 16.2 * u.km / u.s
print("Relative error of {:.2f} %".format((norm(exit.v) - v_estimated) / v_estimated * 100))
# -
# So it stays within the same order of magnitude. Which is reasonable, because real life burns are not instantaneous.
# +
op = OrbitPlotter()
op.plot(parking)
op.plot(exit)
plt.xlim(-8000, 8000)
plt.ylim(-20000, 20000)
plt.gcf().autofmt_xdate()
# -
# ### Option b): Compute $v_{\infty}$ using the Jupyter flyby
#
# According to Wikipedia, the closest approach occurred at 05:43:40 UTC. We can use this data to compute the solution of the Lambert problem between the Earth and Jupiter.
# +
nh_date = time.Time("2006-01-19 19:00", scale='utc')
nh_flyby_date = time.Time("2007-02-28 05:43:40", scale='utc')
nh_tof = nh_flyby_date - nh_date
nh_earth = Orbit.from_body_ephem(Earth, nh_date)
nh_r_0, v_earth = nh_earth.rv()
nh_jup = Orbit.from_body_ephem(Jupiter, nh_flyby_date)
nh_r_f, v_jup = nh_jup.rv()
(nh_v_0, nh_v_f), = iod.lambert(Sun.k, nh_r_0, nh_r_f, nh_tof)
# -
# The hyperbolic excess velocity is measured with respect to the Earth:
C_3_lambert = (norm(nh_v_0 - v_earth)).to(u.km / u.s)**2
C_3_lambert
print("Relative error of {:.2f} %".format((C_3_lambert - C_3_A) / C_3_A * 100))
# Which again, stays within the same order of magnitude of the figure given to the Guo & Farquhar report.
# ## From Earth to Jupiter
# +
from poliastro.plotting import BODY_COLORS
nh = Orbit.from_vectors(Sun, nh_r_0.to(u.km), nh_v_0.to(u.km / u.s), nh_date)
op = OrbitPlotter()
op.plot(nh_jup, label=Jupiter)
plt.gca().set_autoscale_on(False)
op.plot(nh_earth, label=Earth)
op.plot(nh, label="New Horizons");
| docs/source/examples/Exploring the New Horizons launch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SciPy
#
# SciPy é uma coleção de algoritmos matemáticos e funções de conveniência construídas sob Numpy
#
# Ele tem funções otimizadas pra trabalhar com matrizes (incluindo matrizes esparsas).
#
# * Determinante
# * Autovalor
# * Autovetor
# * Solução de sistemas de equações lineares
# * Etc...
#
# Além disso, inclui funções específicas para problemas clássicos de engenharia como:
#
# * Transformadas de Fourier
# * Funções de Bessel,
# * Solver para otimização de funções
# * Etc...
#
# Também tem um módulo estatístico, com diversar funções úteis:
#
# * T-student
# * Teste de normalidade
# * ANOVA
# * Correlação
# * Etc...
import numpy as np
# ## Algebra Linear
from scipy import linalg
A = np.array([[6,5,3,2],[4,8,4,3],[5,3,1,2],[10,9,8,7]])
A
# ### Calcule o determinante de uma matriz
linalg.det(A)
# ### Decomposição A = P L U
P, L, U = linalg.lu(A)
P
L
U
# ### Multiplicação matricial:
#
np.dot(P,L)
# Multiplicação matricial:
np.dot(np.dot(P,L),U)
# ### Autovalores e Autovetores
EW, EV = linalg.eig(A)
EV
EW
# ### Sistemas de equações lineares
#
v = np.array([[1],[1],[1],[1]])
s = linalg.solve(A,v)
s
# ## Transformadas de Fourier Usando SciPy
#
# Retirado de:
#
# (http://www.estruturas.ufpr.br/disciplinas/pos-graduacao/introducao-a-computacao-cientifica-com-python/introducao-python/4-4-transformada-rapida-de-fourier/)
#
# +
time_step = 0.02
period = 5.
time_vec = np.arange(0, 20, time_step)
sig = np.sin(2 * np.pi / period * time_vec) + \
0.5 * np.random.randn(time_vec.size)
from scipy import fftpack
sample_freq = fftpack.fftfreq(sig.size, d=time_step)
sig_fft = fftpack.fft(sig)
# -
sig_fft
pidxs = np.where(sample_freq > 0)
freqs = sample_freq[pidxs]
power = np.abs(sig_fft)[pidxs]
import matplotlib.pyplot as pl
pl.figure()
pl.plot(freqs, power)
pl.xlabel('Frequencia [Hz]')
pl.ylabel('Energia')
axes = pl.axes([0.3, 0.3, 0.5, 0.5])
pl.title('Pico de frequencia')
pl.plot(freqs[:8], power[:8])
pl.setp(axes, yticks=[])
pl.show()
freq = freqs[power.argmax()]
np.allclose(freq, 1./period) # checa se aquele frequência correta é encontrada
sig_fft[np.abs(sample_freq) > freq] = 0
ain_sig = fftpack.ifft(sig_fft)
plt.figure()
pl.plot(time_vec, sig)
pl.xlabel('Tempo [s]')
pl.ylabel('Amplitude')
# ## Otimização Linear Usando SciPy
# ##### Exemplo:
#
# Min: f = -1x + 4y
#
# Sujeito a:
#
# -3x + 1y <= 6
# 1x + 2y <= 4
# y >= -3
from scipy.optimize import linprog
# +
c = [-1, 4]
A = [[-3, 1], [1, 2]]
b = [6, 4]
method = 'Simplex' # Escolhe o solver, pode ser por ‘interior-point’ ou 'Simplex'
x_bounds = (None, None)
y_bounds = (-3, None)
res = linprog(c, A, b, bounds=(x_bounds, y_bounds), method=method, options={"disp": True})
res
# -
# ## ANOVA Usando SciPy
#
# Referência: (https://pythonfordatascience.org/anova-python/)
import scipy.stats as stats
import pandas as pd
df = pd.read_csv("https://raw.githubusercontent.com/Opensourcefordatascience/Data-sets/master/difficile.csv")
df.drop('person', axis= 1, inplace= True)
import statsmodels.api as sm
from statsmodels.formula.api import ols
df['dose'].replace({1: 'placebo', 2: 'low', 3: 'high'}, inplace= True)
df
stats.f_oneway(df['libido'][df['dose'] == 'high'],
df['libido'][df['dose'] == 'low'],
df['libido'][df['dose'] == 'placebo'])
results = ols('libido ~ C(dose)', data=df).fit()
results.summary()
aov_table = sm.stats.anova_lm(results, typ=2)
aov_table
# +
def anova_table(aov):
aov['mean_sq'] = aov[:]['sum_sq']/aov[:]['df']
aov['eta_sq'] = aov[:-1]['sum_sq']/sum(aov['sum_sq'])
aov['omega_sq'] = (aov[:-1]['sum_sq']-(aov[:-1]['df']*aov['mean_sq'][-1]))/(sum(aov['sum_sq'])+aov['mean_sq'][-1])
cols = ['sum_sq', 'df', 'mean_sq', 'F', 'PR(>F)', 'eta_sq', 'omega_sq']
aov = aov[cols]
return aov
anova_table(aov_table)
# -
#Teste Shapiro-Wilk de normalidade
stats.shapiro(results.resid)
# +
from statsmodels.stats.multicomp import pairwise_tukeyhsd
from statsmodels.stats.multicomp import MultiComparison
mc = MultiComparison(df['libido'], df['dose'])
mc_results = mc.tukeyhsd()
print(mc_results)
| curso/2 - Data Science Tool Kit/SciPy/DataScience Toolkit - SciPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc="true"
# # Table of Contents
# <div class="toc" style="margin-top: 1em;"><ul class="toc-item" id="toc-level0"></ul></div>
# +
import cmocean.cm as cm
import matplotlib.pyplot as plt
import numpy as np
import xarray as xr
# %matplotlib inline
# -
#orig = xr.open_dataset('/Users/sallen/Documents/MEOPAR/Results/test_benthos/original_ptrc.nc')
#benthos = xr.open_dataset('/Users/sallen/Documents/MEOPAR/Results/test_benthos/benthos_ptrc.nc')
#orig = xr.open_dataset('/Users/sallen/Documents/MEOPAR/Results/test_benthos/orig_aug31.nc')
orig = xr.open_dataset('/results/SalishSea/spinup.201905/31aug15/SalishSea_1h_20150831_20150831_ptrc_T.nc')
#benthos = xr.open_dataset('/Users/sallen/Documents/MEOPAR/Results/test_benthos/new_aug31.nc')
benthos = xr.open_dataset('/data/sallen/results/MEOPAR/test_benthos/SalishSea_1h_20130831_20130831_ptrc_T_20130831-20130831.nc')
it = 0
fig, axs = plt.subplots(1, 3, figsize=(15, 4))
orig.diatoms[it, :].sum(axis=0).plot(ax=axs[0], vmax=50, cmap=cm.algae)
orig.diatoms[it, :, 260:400, 75].plot(ax=axs[1], vmax=2)
orig.diatoms[it, :, 520, 170:].plot(ax=axs[2], vmax=1)
axs[1].invert_yaxis()
axs[1].set_ylim(30, 0)
axs[2].invert_yaxis()
axs[2].set_ylim(30, 0);
fig, axs = plt.subplots(1, 3, figsize=(15, 4))
benthos.diatoms[it, :].sum(axis=0).plot(ax=axs[0], vmax=80, cmap=cm.algae)
benthos.diatoms[it, :, 260:400, 75].plot(ax=axs[1], vmax=5)
benthos.diatoms[it, :, 520, 170:].plot(ax=axs[2], vmax=2)
axs[1].invert_yaxis()
axs[1].set_ylim(30, 0)
axs[2].invert_yaxis()
axs[2].set_ylim(30, 0);
fig, axs = plt.subplots(1, 3, figsize=(15, 4))
(benthos.diatoms[it, :] - orig.diatoms[it, :]).sum(axis=0).plot(ax=axs[0])
(benthos.diatoms[it, :, 260:400, 75] - orig.diatoms[it, :, 260:400, 75]).plot(ax=axs[1], vmax=2.5)
(benthos.diatoms[it, :, 520, 170:] - orig.diatoms[it, :, 520, 170:]).plot(ax=axs[2], vmax=2.5)
axs[1].invert_yaxis()
axs[1].set_ylim(30, 0)
axs[2].invert_yaxis()
axs[2].set_ylim(30, 0);
print ((benthos.diatoms[it, :] - orig.diatoms[it, :]).sum(axis=0).sum(axis=0).sum(axis=0))
((benthos.diatoms[it, :, 320:440, 50] - orig.diatoms[it, :, 320:440, 50]).sum(axis=0).sum(axis=0) /
orig.diatoms[it, :, 320:440, 50].sum(axis=0).sum(axis=0) )
((benthos.diatoms[it, :, 260:400, 75] - orig.diatoms[it, :, 260:400, 75]).sum(axis=0).sum(axis=0) /
orig.diatoms[it, :, 260:400, 75].sum(axis=0).sum(axis=0) )
((benthos.diatoms[it, :, 500, 200:] - orig.diatoms[it, :, 500, 200:]).sum(axis=0).sum(axis=0) /
orig.diatoms[it, :, 500, 200:].sum(axis=0).sum(axis=0))
((benthos.diatoms[it, :, 520, 170:] - orig.diatoms[it, :, 520, 170:]).sum(axis=0).sum(axis=0) /
orig.diatoms[it, :, 520, 170:].sum(axis=0).sum(axis=0))
| notebooks/SummerBenthicSink.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import collections as col
import importlib
import re
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import matplotlib.lines as lines
plot_aux_module = '/home/local/work/code/github/project-diploid-assembly/notebooks/aux_mods/plot_aux.py'
plot_aux_spec = importlib.util.spec_from_file_location("plot_aux", plot_aux_module)
plot_aux = importlib.util.module_from_spec(plot_aux_spec)
plot_aux_spec.loader.exec_module(plot_aux)
samples = plot_aux.load_sample_table()
hexcodes, rgbcodes, popmap = plot_aux.load_population_annotation()
prop = plot_aux.get_plot_property
save_plots = True
print_stats = False
pipeline_version = 'v12'
stats_path = '/home/local/work/data/hgsvc/figSX_panels/missing_regulatory'
out_path = os.path.join(stats_path, pipeline_version, 'figS13_PE_hap-assm-regbuild')
cache_file = os.path.join(stats_path, 'cache_{}.h5'.format(pipeline_version))
def load_regb_stats(file_path):
df = pd.read_csv(file_path, sep='\t', header=None, names=['value'], index_col=[0])
df = df.transpose()
for c in df.columns:
if c.endswith('_pct'):
df[c] = df[c].astype('float64')
else:
df[c] = df[c].astype('int64')
return df
if not os.path.isfile(cache_file):
stats_files = plot_aux.load_plot_data_files(
stats_path,
'.stats.tsv',
pipeline_version)
stats_data = [load_regb_stats(fp) for fp in stats_files]
sample_platform = [
plot_aux.extract_sample_platform(os.path.basename(f), multi_readset=True) for f in stats_files
]
row_index = []
for sample, platform in sample_platform:
super_pop = samples[sample]['super_population']
pop = samples[sample]['population']
row_index.append((sample, super_pop, pop, platform, 99))
row_index = pd.MultiIndex.from_tuples(
row_index,
names=['sample', 'super_pop', 'pop', 'platform', 'epsilon']
)
df = pd.concat(stats_data, axis=0)
df.index = row_index
df.to_hdf(cache_file, key='cache', mode='w', format='fixed')
df = pd.read_hdf(cache_file, 'cache')
df.sort_index(axis=0, inplace=True, level=['sample', 'super_pop', 'pop', 'platform'])
def plot_regb_statistic(sample_stats):
bar_width = prop('bar_width')
categories = [
'Heterozygous',
'Illumina missing',
'GRCh38 only'
]
selectors = {
'Heterozygous': ['total_Hap1_bp_pct', 'total_Hap2_bp_pct'],
'Illumina missing': ['total_NonIllumina_bp_pct'],
'GRCh38 only': ['total_RefOnly_bp_pct']
}
properties = {
'HiFi': {'color': 'red'}
}
x_pos_counter = 0
hom_stats = {
'CLR': [],
'HiFi': []
}
fig, axis = plt.subplots(figsize=(8,8))
for category in categories:
for platform in ['CLR', 'HiFi']:
dist_values = []
subset = sample_stats.xs(platform, level='platform')
if category == 'Heterozygous' and platform == 'CLR':
print(subset.loc[subset['total_Hap2_bp_pct'] > 5, 'total_Hap2_bp_pct'])
hom_stats[platform] = subset['total_HapBoth_bp_pct']
for selector in selectors[category]:
dist_values.extend(subset[selector].values.tolist())
axis.boxplot(
dist_values,
notch=False,
sym='x',
widths=bar_width,
positions=[x_pos_counter],
boxprops={
'color': plot_aux.get_platform_color(platform),
'linewidth': 2
},
meanprops={
'color': plot_aux.get_platform_color(platform),
'linewidth': 2
},
medianprops={
'color': plot_aux.get_platform_color(platform),
'linewidth': 2
},
whiskerprops={
'color': plot_aux.get_platform_color(platform),
'linewidth': 2
},
flierprops={
'color': plot_aux.get_platform_color(platform),
'linewidth': 2,
'markersize': 10
},
capprops={
'color': plot_aux.get_platform_color(platform),
'linewidth': 2
}
)
x_pos_counter += 1
# axis.set_xlabel(
# 'Samples',
# fontsize=prop('fontsize_axis_label')
# )
axis.set_ylabel(
'Regulatory Build regions (% bp)',
fontsize=prop('fontsize_axis_label')
)
axis.set_ylim(0, 6.5)
axis.spines['top'].set_visible(False)
axis.spines['right'].set_visible(False)
axis.set_xticks([0.5, 2.5, 4.5])
axis.set_xticklabels(categories)
axis.tick_params(
axis='both',
which='major',
labelsize=prop('fontsize_axis_ticks')
)
# build custom legend
custom_patches = [
patches.Patch(
facecolor=plot_aux.get_platform_color('CLR'),
edgecolor='white',
label='CLR'
),
patches.Patch(
facecolor=plot_aux.get_platform_color('HiFi'),
edgecolor='white',
label='HiFi',
),
]
axis.legend(
handles=custom_patches,
loc='upper right',
handlelength=3,
handleheight=1,
prop={'size': prop('fontsize_legend')}
)
hom_stats['HiFi'] = np.array(hom_stats['HiFi'], dtype=np.float64)
hom_stats['CLR'] = np.array(hom_stats['CLR'], dtype=np.float64)
hom_text = 'Homozygous:\nCLR {}%\nHiFi {}%'.format(
hom_stats['CLR'].mean().round(1),
hom_stats['HiFi'].mean().round(1)
)
axis.text(
x=0.05,
y=0.5,
s=hom_text,
fontdict={'size': prop('fontsize_legend')}
)
extra_artists = []
return fig, extra_artists
fig, exart = plot_regb_statistic(df)
if save_plots:
low_res_dpi = prop('dpi_low_res')
norm_res_dpi = prop('dpi_norm_res')
fig.savefig(
out_path + '.{}dpi.png'.format(low_res_dpi),
dpi=low_res_dpi,
bbox_inches='tight',
extra_artists=exart
)
fig.savefig(
out_path + '.{}dpi.png'.format(norm_res_dpi),
dpi=norm_res_dpi,
bbox_inches='tight',
extra_artists=exart
)
fig.savefig(
out_path + '.svg',
bbox_inches='tight',
extra_artists=exart
)
if print_stats:
for platform in ['CLR', 'HiFi']:
subset = df.xs(platform, level='platform')
data_values = []
for s in ['total_Hap1_bp_pct', 'total_Hap2_bp_pct']:
data_values.extend(subset[s].values.tolist())
print('=== {} summary'.format(platform))
data_values = np.array(data_values, dtype=np.float64)
print('- mean ', data_values.mean().round(2))
print('- stddev ', data_values.std().round(2))
print(subset['total_RefOnly_bp_pct'].describe())
print(subset['total_NonIllumina_bp_pct'].describe())
| notebooks/dev/hgsvc_figSX_panels/plot_regb_stats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Kd4ShwFw_Yy-"
# <NAME>
#
# **Practical 8**
# + [markdown] id="sNHpYxke_1ur"
# **Liner SVM**
# + id="l5eiQjfZ_H9T"
import sklearn
from sklearn import svm,datasets,metrics
# + id="UY9utA5R_sZg" outputId="f11696bb-6e3a-4ab4-c4d5-1c3c34ad9b29" colab={"base_uri": "https://localhost:8080/"}
X,Y = datasets.load_iris(return_X_y=True)
xtrain = X[range(0,150,2),:]
ytrain = Y[range(0,150,2)]
xtest = X[range(1,150,2),:]
ytest = Y[range(1,150,2)]
# linear svm
clf = svm.LinearSVC()
clf.fit(xtrain,ytrain)
ypred=clf.predict(xtest)
print("ACCURACY ",metrics.accuracy_score(ytest,ypred))
print("REPORT ",metrics.classification_report(ytest,ypred))
print("Confusion Metrix ",metrics.confusion_matrix(ytest,ypred))
# + [markdown] id="1ZCzmKOZAUVZ"
# **Grid** **Search**
# + id="unQTz_dTAP6o"
import numpy as np
import sklearn
from sklearn import datasets,metrics,neighbors
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import Ridge
from sklearn.linear_model import SGDClassifier
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
# + [markdown] id="R2kwvzIHAaKR"
# Grid search for knn
# + id="tiXvrvsVAOqR" outputId="5e41a831-d2fa-4eb2-95fd-f7d25cb42de4" colab={"base_uri": "https://localhost:8080/"}
X,Y =datasets.load_iris(return_X_y=True)
xtrain=X[range(0,150,2),:]
xtest=X[range(1,150,2),:]
ytrain=Y[range(0,150,2)]
ytest=Y[range(1,150,2)]
clf = neighbors.KNeighborsClassifier()
grid = {'n_neighbors':[1,5,7,11,17],'weights':['distance','uniform']}
gs = GridSearchCV(clf,param_grid=grid,cv=5)
clf=gs
clf.fit(xtrain,ytrain)
print("best params are\n"+str(clf.best_params_))
ypred=clf.predict(xtest)
print("Actual:",ytest)
print("Predicted:",ypred)
print(metrics.classification_report(ytest,ypred))
print(metrics.confusion_matrix(ytest,ypred))
print("\n accuracy",metrics.accuracy_score(ytest,ypred))
# + [markdown] id="lT-ZBW6NAnk5"
# Grid search for multiple regression
# + id="uT-YtsFrAmrT" outputId="157f99a7-44b6-4609-a2ee-0203ffe7ced1" colab={"base_uri": "https://localhost:8080/"}
X,Y=datasets.load_boston(return_X_y=True)
temp=np.ones(shape=(300,1))
xtrain=X[0:300,:]
xtrain=np.append(temp,xtrain,axis=1)
#print(xtrain[:10,:])
ytrain=Y[0:300]
ytrain=ytrain.astype('int')
xtest=np.append(np.ones(shape=(206,1)),X[300:,:],axis=1)
ytest=Y[300:]
ytest=ytest.astype('int')
model=StandardScaler()
model.fit(xtest)
xtest=model.transform(xtest)
model.fit(xtrain)
xtrain=model.transform(xtrain)
param_grid = {'alpha': [100, 1000, 10000], 'max_iter': [100,5000, 500]}
clf = SGDClassifier()
gs = GridSearchCV(clf, param_grid, cv=5)
clf=gs
clf.fit(xtrain, ytrain)
print(clf.best_params_)
ypred=clf.predict(xtest)
print(ypred)
print("MSE ",metrics.mean_squared_error(ytest,ypred));
print("MSA ",metrics.mean_absolute_error(ytest,ypred));
| SVM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to create Figure 4
#
#
# **Paper:** https://arxiv.org/pdf/2002.09301.pdf
#
#
# +
import numpy as np
import matplotlib.pyplot as plt
from difflikelihoods import odesolver
from difflikelihoods import linearised_odesolver as linsolver
from difflikelihoods import linearised_ode as linode
from difflikelihoods import statespace
from difflikelihoods import inverseproblem as ip
from difflikelihoods.optimisation import minimise_newton, minimise_gd, minimise_rs
from sampling_functions import *
# -
# At first we create the data, set the model parameters and the method parameters.
def create_data(solver, ivp, thetatrue, stepsize, ivpvar):
"""
Create artificial data for the inverse problem.
"""
ivp.params = thetatrue
tsteps, m, __, __, __ = solver.solve(ivp, stepsize)
means = odesolver.get_trajectory_ddim(m, 2, 0)
evalpts = np.array([1., 2., 3., 4., 5.])
evalpts = np.arange(.5, 5., 5/10)
assert(np.prod(np.in1d(evalpts, tsteps))==1), print(evalpts[np.in1d(evalpts, tsteps)==False])
noise = np.sqrt(ivpvar)*np.random.randn(len(evalpts)*2).reshape((len(evalpts), 2))
evalidcs = [list(tsteps).index(evalpt) for evalpt in evalpts]
data = means[evalidcs] + noise # this is 'wrong' noise
ipdata = ip.InvProblemData(evalpts, data, ivpvar)
return ipdata
# ## Optimisation Experiments
#
# We begin with the optimisation experiments. We begin by specifying the model and method parameters.
# +
np.random.seed(2)
# Set Model Parameters
initial_value = np.array([20, 20])
initial_time, end_time = 0., 5.
ivpvar = 1e-10
thetatrue = np.array([1.0, 0.1, 0.1, 1.0])
ivp = linode.LotkaVolterra(initial_time, end_time, params=thetatrue, initval=initial_value)
# Set Method Parameters
h_for_data = (end_time - initial_time)/10000
h = (end_time - initial_time)/100
solver = linsolver.LinearisedODESolver(statespace.IBM(q=1, dim=2))
ipdata = create_data(solver, ivp, thetatrue, h_for_data, ivpvar)
iplklhd = ip.InvProblemLklhd(ipdata, ivp, solver, h, with_jacob=True)
# -
# Next, we specify an initial value and compute the minisers for Newton, GD and RS as well as the respective root mean squared errors.
# +
niter = 100
init_theta = np.array([.8, .2, .05, 1.1])
traj_newton, obj_newton = minimise_newton(iplklhd.potenteval, iplklhd.gradeval, iplklhd.hesseval, niter, init_theta, lrate=0.5)
error_newton = np.sqrt(np.sum(np.abs(traj_newton - thetatrue)**2/(thetatrue**2) ,axis=-1))
traj_gd, obj_gd = minimise_gd(iplklhd.potenteval, iplklhd.gradeval, niter, init_theta, lrate=1e-9)
error_gd = np.sqrt(np.sum(np.abs(traj_gd - thetatrue)**2/(thetatrue**2) ,axis=-1))
traj_rs, obj_rs = minimise_rs(iplklhd.potenteval, niter, init_theta, lrate=1e-2)
error_rs = np.sqrt(np.sum(np.abs(traj_rs - thetatrue)**2/(thetatrue**2) ,axis=-1))
# -
print("Newton guess:", traj_newton[-1])
print("GD guess:", traj_gd[-1])
print("RS guess:", traj_rs[-1])
print("Truth:", thetatrue)
print("Init:", init_theta)
# Finally, we can visualise the results using both the log-likelihood values and the error in the parameter space as metrics.
# +
plt.style.use("./icmlstyle.mplstyle")
fig, (ax1, ax2) = plt.subplots(nrows=1, ncols=2)
prop_cycle = plt.rcParams['axes.prop_cycle']
colors = prop_cycle.by_key()['color']
ax1.set_xlabel("Iteration")
ax1.set_ylabel("Neg. log-likelihood")
mark_every = 5
ax1.semilogy((obj_gd), markevery=mark_every, color="gray", ls="-", marker="^", label="GD", alpha=0.8)
ax1.semilogy((obj_newton), markevery=mark_every, color="#999933", ls="-", marker="d", label="NWT", alpha=0.8)
ax1.semilogy((obj_rs), markevery=mark_every, color="#cc6677", ls="-", marker="s", label="RS", alpha=0.7)
ax2.set_xlabel("Iteration")
ax2.set_ylabel("Rel. Error")
ax2.semilogy(np.abs((traj_gd - thetatrue[np.newaxis, :])/thetatrue[np.newaxis, :]).mean(axis=1), markevery=mark_every, color="gray", ls="-", marker="^", label="GD", alpha=0.8)
ax2.semilogy(np.abs((traj_newton - thetatrue[np.newaxis, :])/thetatrue[np.newaxis, :]).mean(axis=1), markevery=mark_every, color="#999933", ls="-", marker="d", label="NWT", alpha=0.8)
ax2.semilogy(np.abs((traj_rs - thetatrue[np.newaxis, :])/thetatrue[np.newaxis, :]).mean(axis=1), markevery=mark_every, color="#cc6677", ls="-", marker="s", label="RS", alpha=0.7)
ax1.set_title("a", loc="left", fontweight='bold', ha='right')
ax2.set_title("b", loc="left", fontweight='bold', ha='right')
ax2.legend(loc='upper right', bbox_to_anchor=(1.0, 0.6))
ax1.minorticks_off()
ax2.minorticks_off()
plt.tight_layout()
plt.savefig("./figures/figure4_optim_left.pdf")
plt.show()
# -
# ## Sampling experiments
#
# Next, we repeat the experiments with the sampling algorithms. We start with specifying the model and method parameters.
# +
# Set Model Parameters
initial_value = np.array([20, 20])
initial_time, end_time = 0., 5.
ivpvar = 1e-2
thetatrue = np.array([1.0, 0.1, 0.1, 1.0])
ivp = linode.LotkaVolterra(initial_time, end_time, params=thetatrue, initval=initial_value)
# Set Method Parameters
h_for_data = (end_time - initial_time)/10000
h = (end_time - initial_time)/400
solver = linsolver.LinearisedODESolver(statespace.IBM(q=1, dim=2))
ipdata = create_data(solver, ivp, thetatrue, h_for_data, ivpvar)
iplklhd = ip.InvProblemLklhd(ipdata, ivp, solver, h, with_jacob=True)
# -
# Next, we sample from the posteriors with Langevin MCMC, Hamiltonian MC and random walk Metropolis-Hastings.
niter = 250
init_theta = np.array([1, .2, .01, 1.1])
np.random.seed(1)
_ninits = 50
samples_lang, probs_lang = langevin(niter, iplklhd, init_theta, stepsize=1.15, ninits=45)
np.random.seed(1)
samples_ham, probs_ham = hamiltonian(niter, iplklhd, init_theta, stepsize=0.35, nsteps=3, ninits=45)
np.random.seed(1)
samples_rw, probs_rw = randomwalk(niter, iplklhd, init_theta, stepsize=0.0005, ninits=45)
# Finally, we can visualise the results.
# +
# Plot results
plt.style.use("./icmlstyle.mplstyle")
fig, (ax1, ax2) = plt.subplots(nrows=1, ncols=2)
ax1.set_xlabel("Iteration")
ax1.set_ylabel("Neg. log-likelihood")
ax1.semilogy((probs_lang), ls='None', marker="^", label="PLMC", alpha=0.4, markevery=2)
ax1.semilogy((probs_ham), ls='None', marker="d", label="PHMC", alpha=0.4, markevery=2)
ax1.semilogy((probs_rw), ls='None', marker="s", label="RWM", alpha=0.4, markevery=2)
ax2.set_xlabel("Iteration")
ax2.set_ylabel("Rel. Error")
ax2.semilogy(np.abs((samples_lang - thetatrue[np.newaxis, :])/thetatrue[np.newaxis, :]).mean(axis=1), ls='None', marker="^", label="PLMC", alpha=0.4, markevery=2)
ax2.semilogy(np.abs((samples_ham - thetatrue[np.newaxis, :])/thetatrue[np.newaxis, :]).mean(axis=1), ls='None', marker="d", label="PHMC", alpha=0.4, markevery=2)
ax2.semilogy(np.abs((samples_rw - thetatrue[np.newaxis, :])/thetatrue[np.newaxis, :]).mean(axis=1), ls='None', marker="s", label="RWM", alpha=0.4, markevery=2)
ax1.set_title("c", loc="left", fontweight='bold', ha='right')
ax2.set_title("d", loc="left", fontweight='bold', ha='right')
ax2.legend()
ax1.minorticks_off()
ax2.minorticks_off()
plt.tight_layout()
plt.savefig("./figures/figure4_sampling_right.pdf")
plt.show()
# -
| experiments/Figure4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
s = pd.Series([1, 3, 5, np.nan, 6, 8])
s
dates = pd.date_range("20130228", periods=6)
dates
df = pd.DataFrame(np.random.randn(6,4), index = dates, columns = list("ABCD"))
#^randn: normally distributed random numbers
#(6,4) returns a 6 by 4 array (matrix) of random numbers
df
df2 = pd.DataFrame(
{
"Classification": "Housework",
"Date": pd.Timestamp("20220221"),
"Item": pd.Categorical(["cook","do dishes","laundry","vacuum"]),
}
)
df2
| notebooks/practice-pandas/10minPandas_practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from scipy.stats import entropy
a = np.array([0.1,0.5,0.3,0.1])
b = np.array([0.15,0.6,0.2,0.05])
c = np.array([0.15,0.4,0.4,0.05])
(a * b * c) / np.sum(a*b*c)
(((a * b) / np.sum(a*b)) * c) / np.sum((((a * b) / np.sum(a*b)) * c))
# +
bla = np.array([[0.4,0.5,0.05,0.05], [0.6,0.4,0.1,0.1], [0.3,0.65,0.025,0.025]])
# -
bla.shape
avg = np.mean(bla, axis=0)
print(avg)
entropy(avg)
bla2 = np.array([[0.4,0.5,0.05,0.05], [0.6,0.4,0.1,0.1], [0.3,0.65,0.025,0.025], [1.0, 0, 0, 0]])
avg2 = np.mean(bla2, axis=0)
print(avg2)
entropy(avg2)
| simple_accum_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from sklearn.ensemble import RandomForestRegressor
# +
def is_free_cancellation_policy(policy: str):
free_policy = "0D0N"
return free_policy == policy
def is_severe_policy(policy: str):
return policy == "365D100P_100P"
def get_least_precentage(data):
policy_string = data['cancellation_policy_code']
trip_days =data['Vacation_length']
if policy_string == "UNKNOWN":
return 90
least_problematic_policy = policy_string.split("_")[0]
if least_problematic_policy[-1] == "N":
nights_to_pay = (least_problematic_policy[:-1]).split("D")[-1]
return int(nights_to_pay) / trip_days * 100
else:
precentage_p = (least_problematic_policy[:-1]).split("D")[-1]
return int(precentage_p)
def is_unknown_policy(policy_string: str):
return policy_string == "UNKNOWN"
def get_group_size(data):
return data['no_of_adults'] + data['no_of_children']
# +
# parse_dates = ['booking_datetime', 'checkin_date', 'checkout_date', 'hotel_live_date', 'cancellation_datetime']
data = pd.read_csv(r"C:\Users\t8883217\Desktop\IML\Challenge\week1\agoda_cancellation_train.csv")
data['group_size'] = data.apply(get_group_size, axis =1)
data['cancellation_datetime']= pd.to_datetime(data['cancellation_datetime'])
data['checkin_date']= pd.to_datetime(data['checkin_date'],dayfirst = True)
data['checkout_date']= pd.to_datetime(data['checkout_date'],dayfirst = True)
data['booking_datetime']= pd.to_datetime(data['booking_datetime'],dayfirst = True)
display(data[["checkin_date", "booking_datetime"]])
data.loc[data['cancellation_datetime'].notnull(), 'cancellation'] = 1
data.loc[data['cancellation_datetime'].isnull(), 'cancellation'] = 0
data.loc[data['charge_option'] == "Pay Later", 'payment_option'] = 1
data.loc[data['charge_option'] == "Pay Now", 'payment_option'] = 0
data.loc[data['charge_option'] == "Pay at Check-in", 'payment_option'] = 0
data.loc[data['cancellation_policy_code'].str[-5:] == "_100P", 'no_show_100P'] = 1
data.loc[data['cancellation_policy_code'].str[-5:] != "_100P", 'no_show_100P'] = 0
data.loc[data['cancellation_policy_code'].str.contains("_"), 'policy_is_changing'] = 1
data.loc[~data['cancellation_policy_code'].str.contains("_"), 'policy_is_changing'] = 0
data['Vacation_length'] = ((data['checkout_date'] - data['checkin_date'])/np.timedelta64(1, 'D'))
data['Booking_to_checkin'] = ((data['checkin_date'] - data['booking_datetime'])/np.timedelta64(1, 'D'))
data['is_severe_policy'] = data['cancellation_policy_code'].apply(lambda x: is_severe_policy(x))
data['Days_from_checkin'] = ((data.loc[data['cancellation'] == 1]['checkin_date'] - data.loc[data['cancellation'] == 1]['cancellation_datetime'])/np.timedelta64(1, 'D'))
data['Days_after_booking'] = ((data.loc[data['cancellation'] == 1]['cancellation_datetime'] - data.loc[data['cancellation'] == 1]['booking_datetime'])/np.timedelta64(1, 'D'))
print(data['Days_from_checkin'])
data['min_precentage_to_pay'] = data.apply(get_least_precentage, axis =1)
display(data[["checkin_date", "booking_datetime", "Booking_to_checkin"]])
data = data.drop(['h_booking_id', 'booking_datetime', 'checkout_date',
'hotel_id', 'hotel_country_code', 'hotel_live_date',
'accommadation_type_name', 'h_customer_id', 'customer_nationality',
'guest_nationality_country_name', 'original_payment_type',"original_payment_currency",
'no_of_extra_bed', 'no_of_room', 'origin_country_code', 'language', "original_payment_method",
'request_nonesmoke', 'request_latecheckin', 'request_highfloor',
'request_largebed', 'request_twinbeds', 'request_airport', 'cancellation_policy_code',
'request_earlycheckin', 'hotel_area_code','no_of_children',
'hotel_brand_code', 'hotel_chain_code', 'hotel_city_code',"charge_option"],1) #TO ADD: , 'accommadation_type_name', original_payment_currency
data
# +
# Import train_test_split function
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from catboost import Pool, CatBoostClassifier
from sklearn import metrics
# Split dataset into features and labels
y=data["cancellation"]
X = data.drop(["cancellation", 'Days_from_checkin','Days_after_booking', "cancellation_datetime"],1)
# Split dataset into training set and test set
X_train, X_tv, y_train, y_tv = train_test_split(X, y, test_size=0.40, random_state=5) # 70% training and 30% test
X_validation, X_test, y_validation, y_test = train_test_split(X_tv, y_tv, test_size=0.50, random_state=5) # 70% training and 30% test
include_checkin_date = X_test
X_test = X_test.drop("checkin_date",1)
X_validation = X_validation.drop("checkin_date",1)
X_train = X_train.drop("checkin_date",1)
clf=RandomForestClassifier(n_estimators=300, min_samples_leaf=20, max_depth=19)
clf.fit(X_train,y_train)
# print("Accuracy:",metrics.accuracy_score(y_train, y_pred_train))
# print("Precision:",metrics.precision_score(y_train, y_pred_train))
# print("Re-call:",metrics.recall_score(y_train, y_pred_train))
feature_names = [featureName for (featureName, featureData) in X_test.iteritems()]
importances = clf.feature_importances_
forest_importances = pd.Series(importances, index=feature_names)
fig, ax = plt.subplots()
forest_importances.plot.bar(ax=ax)
ax.set_title("Feature importances using MDI")
ax.set_ylabel("Mean decrease in impurity")
fig.tight_layout()
# +
df_train = data.loc[data['cancellation_datetime'].notnull()]
X2 = df_train.drop(['Days_after_booking', "cancellation"], axis = 1)
Y2 = df_train['Days_from_checkin']
X_train_2, X_test_2, y_train_2, y_test_2 = train_test_split(
X2, Y2, test_size=0.33, random_state=42)
X_train_2 = X_train_2.drop(['cancellation_datetime',"Days_from_checkin", "checkin_date"], axis = 1)
X_test_2 = X_test_2.drop('cancellation_datetime', axis = 1)
regression = RandomForestRegressor(n_estimators = 300, max_depth = 17,random_state = 8)
regression.fit(X_train_2, y_train_2)
# +
parse_dates = ['booking_datetime', 'checkin_date', 'checkout_date', 'hotel_live_date']
data = pd.read_csv(r"C:\Users\t8883217\Desktop\IML\Challenge\week1\test_set_week_1.csv", parse_dates=parse_dates)
data['group_size'] = data.apply(get_group_size, axis =1)
data['checkin_date']= pd.to_datetime(data['checkin_date'])
data['checkout_date']= pd.to_datetime(data['checkout_date'])
data['booking_datetime']= pd.to_datetime(data['booking_datetime'])
data.loc[data['charge_option'] == "Pay Later", 'payment_option'] = 1
data.loc[data['charge_option'] == "Pay Now", 'payment_option'] = 0
data.loc[data['charge_option'] == "Pay at Check-in", 'payment_option'] = 0
data.loc[data['cancellation_policy_code'].str[-5:] == "_100P", 'no_show_100P'] = 1
data.loc[data['cancellation_policy_code'].str[-5:] != "_100P", 'no_show_100P'] = 0
data.loc[data['cancellation_policy_code'].str.contains("_"), 'policy_is_changing'] = 1
data.loc[~data['cancellation_policy_code'].str.contains("_"), 'policy_is_changing'] = 0
data['Vacation_length'] = ((data['checkout_date'] - data['checkin_date'])/np.timedelta64(1, 'D'))
data['Booking_to_checkin'] = ((data['checkin_date'] - data['booking_datetime'])/np.timedelta64(1, 'D'))
data['is_severe_policy'] = data['cancellation_policy_code'].apply(lambda x: is_severe_policy(x))
data['min_precentage_to_pay'] = data.apply(get_least_precentage, axis =1)
display(data[["checkin_date", "booking_datetime"]])
data = data.drop(['h_booking_id', 'booking_datetime', 'checkout_date',
'hotel_id', 'hotel_country_code', 'hotel_live_date',
'accommadation_type_name', 'h_customer_id', 'customer_nationality',
'guest_nationality_country_name', 'original_payment_type',"original_payment_currency",
'no_of_extra_bed', 'no_of_room', 'origin_country_code', 'language', "original_payment_method",
'request_nonesmoke', 'request_latecheckin', 'request_highfloor',
'request_largebed', 'request_twinbeds', 'request_airport', 'cancellation_policy_code',
'request_earlycheckin', 'hotel_area_code','no_of_children',
'hotel_brand_code', 'hotel_chain_code', 'hotel_city_code',"charge_option"],1) #TO ADD: , 'accommadation_type_name', original_payment_currency
data
# -
# +
include_checkin_date = data
test = include_checkin_date.drop("checkin_date",1)
y_pred=clf.predict(test)
cancel_date = include_checkin_date.loc[y_pred==1]['checkin_date']
# -
df_cancel = test.loc[y_pred == 1]
# df_cancel_true = df_cancel["Days_from_checkin"]
# df_cancel = df_cancel.drop(['Days_from_checkin', 'Days_after_booking'], axis = 1)
pred = regression.predict(df_cancel)
# train_pred = regression.predict(X_train_2)
# pd.DataFrame(df_cancel_true).hist(bins=10)
pd.DataFrame(pred).hist(bins=10)
# pd.DataFrame(train_pred).hist(bins=10)
# +
# display(pd.DataFrame(abs(regression.coef_)))
# display(X.columns)
# plt.bar(X_train.columns, abs(regression.coef_))
# plt.tight_layout()
display(pred.shape, cancel_date.shape)
pred_date = cancel_date - pd.to_timedelta(pred, unit = "D")
display(pred_date.to_frame())
start_date = pd.to_datetime('2018-12-07')
end_date = pd.to_datetime('2018-12-13')
prediction_binary = pred_date.between(start_date, end_date)
display(prediction_binary)
display(pred)
# -
# prediction_binary.count()
item_counts = prediction_binary.value_counts()
item_counts
# display(include_checkin_date[])
include_checkin_date.insert(1, "prediction_value", prediction_binary)
include_checkin_date
include_checkin_date["prediction_value"]=include_checkin_date["prediction_value"].fillna(False)
include_checkin_date
include_checkin_date.to_csv("213096175_.csv",index=False)
include_checkin_date["prediction_value"].to_csv("212422794_212618169_213096175.csv", index = False)
| data_challenge_week_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/islam-mirajul/Name-Based-Community-Prediction/blob/main/Word-Frequency-Analysis.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="dbTyAgsD7VaX" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="261f9b00-f8f1-4709-8960-9e5688e225a7"
#Full Name Analysis
import pandas as pd
df = pd.read_csv('/content/drive/MyDrive/name-religion/Religion-Prediction-large.csv')
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 173} id="tdEz0uQSAhYK" outputId="b6db5d9b-940c-491c-b10b-4562e11c84d0"
df.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="BscHBkUfAl4W" outputId="c52cd751-0ad4-4f0a-c672-281deceb8278"
msg = df.Name
msg_split = msg.str.split(expand=True)
msg_split.head()
# + colab={"base_uri": "https://localhost:8080/"} id="qOSqDDQtAwWl" outputId="856417c3-28d9-4650-f919-10c4225d3315"
msg_stack = msg_split.stack()
msg_stack.head()
# + colab={"base_uri": "https://localhost:8080/"} id="WbvocooDBBga" outputId="28530a39-0aa8-4679-e55e-71551450b139"
n_most_frequent_words = 40
msg_word_frequency = msg_stack.value_counts()
print(msg_word_frequency.head(n_most_frequent_words))
pd.set_option('display.max_rows', None)
# + colab={"base_uri": "https://localhost:8080/", "height": 313} id="F0pILffgC1hg" outputId="d091f3ad-53af-4c50-daaf-3c66d1822d87"
import matplotlib.pyplot as plt
# %matplotlib inline
import matplotlib.font_manager as fm
prop = fm.FontProperties(fname='/content/drive/MyDrive/gender project/kalpurush.ttf')
fontproperties=prop
plt.plot
plt.figure(figsize=(12,4))
plt.ylabel('Counts', fontsize=15)
plt.xlabel('Words', fontsize=15)
plt.xticks(rotation=90, fontsize=1000000000000, fontproperties=prop)
msg_word_frequency.head(n_most_frequent_words).plot.bar(msg_word_frequency,n_most_frequent_words);
# + [markdown] id="ANz3D6L8M7wo"
# **First Name Analysis**
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="v6qVWeq2F8kD" outputId="82f08eeb-6ca4-4fb2-b050-d76143ed245e"
df = pd.read_csv('/content/drive/MyDrive/name-religion/Religion-prediction_first-name_.csv')
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 173} id="d2-qVgAONh9o" outputId="a159c3b5-15d6-4b86-ef01-39b585043f90"
df.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="RvA02x7mNr1p" outputId="05c7a4be-f5d9-4d0d-d651-d1cf1d6947bb"
msg = df.FirstName
msg_split = msg.str.split(expand=True)
msg_split.head()
# + colab={"base_uri": "https://localhost:8080/"} id="Pr74at4xQ0Jg" outputId="c34250d8-2068-48db-f788-165df88ece2c"
msg_stack = msg_split.stack()
msg_stack.head()
# + colab={"base_uri": "https://localhost:8080/"} id="XpiSsL3QQ9Kj" outputId="6a966f0e-9ae0-41fb-ef6b-c8f1eed98446"
n_most_frequent_words = 40
msg_word_frequency = msg_stack.value_counts()
print(msg_word_frequency.head(n_most_frequent_words))
pd.set_option('display.max_rows', None)
# + colab={"base_uri": "https://localhost:8080/", "height": 309} id="d4rTWJmYRBNS" outputId="8ce19e70-39b6-47bc-8efb-7fe63ad699fa"
import matplotlib.pyplot as plt
# %matplotlib inline
import matplotlib.font_manager as fm
prop = fm.FontProperties(fname='/content/drive/MyDrive/gender project/kalpurush.ttf')
fontproperties=prop
plt.plot
plt.figure(figsize=(12,4))
plt.ylabel('Counts', fontsize=15)
plt.xlabel('Words', fontsize=15)
plt.xticks(rotation=90, fontsize=100, fontproperties=prop)
msg_word_frequency.head(n_most_frequent_words).plot.bar(msg_word_frequency,n_most_frequent_words);
# + [markdown] id="6smYA9H8SraV"
# **Last Name**
#
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="GI_vjImCRSLJ" outputId="a7171568-a302-43f4-e787-253e8a7ebb13"
df = pd.read_csv('/content/drive/MyDrive/name-religion/Religion-prediction_last-name_.csv')
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 173} id="yefSpTlDS2g0" outputId="c1cae062-0a05-437d-feff-80fd791e57ea"
df.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="UQITtiGNS8L_" outputId="92a67b36-a065-440e-f171-b706c7893d46"
msg = df.LastName
msg_split = msg.str.split(expand=True)
msg_split.head()
# + colab={"base_uri": "https://localhost:8080/"} id="81b1A8gqTB7G" outputId="0cabfe7a-80c2-4106-b549-c652a310c9d1"
msg_stack = msg_split.stack()
msg_stack.head()
# + colab={"base_uri": "https://localhost:8080/"} id="NV-5w-RMTGru" outputId="9aa9329e-bf60-4682-8eda-81adb6a3aa72"
n_most_frequent_words = 40
msg_word_frequency = msg_stack.value_counts()
print(msg_word_frequency.head(n_most_frequent_words))
pd.set_option('display.max_rows', None)
# + colab={"base_uri": "https://localhost:8080/", "height": 326} id="8auU81sQTJ09" outputId="4f60250d-c070-4fd4-9e08-2e66f10fc261"
import matplotlib.pyplot as plt
# %matplotlib inline
import matplotlib.font_manager as fm
prop = fm.FontProperties(fname='/content/drive/MyDrive/gender project/kalpurush.ttf')
fontproperties=prop
plt.plot
plt.figure(figsize=(12,4))
plt.ylabel('Counts', fontsize=15)
plt.xlabel('Words', fontsize=15)
plt.xticks(rotation=90, fontsize=100, fontproperties=prop)
msg_word_frequency.head(n_most_frequent_words).plot.bar(msg_word_frequency,n_most_frequent_words);
| Word-Frequency-Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy
import matplotlib.pyplot as plt
pd_data_train = pd.read_csv("./training_set.csv")
pd_data_test = pd.read_csv("./test_set.csv")
# data virtualization
pd_data_train.plot(x='Date', y='Close Price', title='Data virtualize')
plt.show()
print( numpy.isnan((pd_data_train).any()) )
print( numpy.isnan((pd_data_test).any()) )
pd_data_train.head(5)
# Use lambda to implement switch
switcher_month = {
'Jan': '01',
'Feb': '02',
'Mar': '03',
'Apr': '04',
'May': '05',
'Jun': '06',
'Jul': '07',
'Aug': '08',
'Sep': '09',
'Oct': '10',
'Nov': '11',
'Dec': '12',
}
print(pd_data_train['Date'][0][3:6])
print(switcher_month.get(pd_data_train['Date'][0][3:6]))
# +
# data preprocecing
pd_data_train_y = numpy.zeros(pd_data_train.shape[0], dtype = 'int')
for i in range(0, pd_data_train.shape[0]) :
tmp = pd_data_train['Date'][i]
pd_data_train.at[i, 'Date'] = tmp[7:11] + switcher_month.get(tmp[3:6]) + tmp[0:2]
if i != 0:
prev = pd_data_train.at[i-1, 'Close Price']
cur = pd_data_train.at[i, 'Close Price']
if prev > cur :
pd_data_train_y[i] = 0
else :
pd_data_train_y[i] = 1
pd_data_train.head(5)
# -
pd_data_train_y = pd.DataFrame(pd_data_train_y, columns=['result'])
pd_data_train_y.head(5)
pd_data_train_x = pd_data_train[['Date']]
# Because cann't get data of 2009/01/01, so the result of 2009/01/02 is unknowm
pd_data_train_x = pd_data_train_x.drop([0])
pd_data_train_y = pd_data_train_y.drop([0])
pd_data_train_y.head(5)
# +
pd_data_test_y = numpy.zeros(pd_data_test.shape[0], dtype = 'int')
for i in range(0, pd_data_test.shape[0]) :
tmp = pd_data_test['Date'][i]
pd_data_test.at[i, 'Date'] = tmp[7:11] + switcher_month.get(tmp[3:6]) + tmp[0:2]
if i != 0:
prev = pd_data_test.at[i-1, 'Close Price']
cur = pd_data_test.at[i, 'Close Price']
if prev > cur :
pd_data_test_y[i] = 0
else :
pd_data_test_y[i] = 1
pd_data_test.head(5)
# -
pd_data_test_y = pd.DataFrame(pd_data_test_y, columns=['result'])
pd_data_test_y.head(5)
pd_data_test_x = pd_data_test[['Date']]
# Because cann't get data of 2018/01/01, so the result of 2018/01/02 is unknowm
pd_data_test_x = pd_data_test_x.drop([0])
pd_data_test_y = pd_data_test_y.drop([0])
pd_data_test_y.head(5)
# +
from sklearn.datasets import load_iris
from sklearn.linear_model import LogisticRegression
model = LogisticRegression(solver='sag')
model = model.fit (pd_data_train_x,pd_data_train_y['result'])
model.score(pd_data_test_x,pd_data_test_y['result'])
# -
model.predict(pd_data_test_x)
# +
from sklearn.svm import SVC
model = SVC(gamma='auto', degree=1, class_weight='balanced')
model = model.fit (pd_data_train_x,pd_data_train_y['result'])
model.score(pd_data_test_x,pd_data_test_y['result'])
# -
model.predict(pd_data_test_x)
# +
from sklearn.neural_network import MLPClassifier
model = MLPClassifier(solver='lbfgs', alpha=1e-5,
hidden_layer_sizes=(15,2), random_state=1)
model = model.fit (pd_data_train_x,pd_data_train_y['result'])
model.score(pd_data_test_x,pd_data_test_y['result'])
# -
model.predict(pd_data_test_x)
| Stock-movement_prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Using Excel for parameter input and model runs
#
# Besides using a python script or a jupyter notebook it is also possible to read and run a model using excel files.
#
# The energySystemModelRunFromExcel() function reads a model from excel, optimizes it and stores it to an excel file.
#
# The readEnergySystemModelFromExcel() function reads a model from excel.
#
# The model run can also be started on double-klicking on the run.bat Windows batch script in the folder where this notebook is located (still requires that a Python version, the FINE package and an optmization solver are installed).
#
# Moreover, it is possible to run the model inside the Excel file with a VBA Macro (see scenarioInputWithMacro.xlsx)
# # Import FINE package
#
import FINE as fn
# %load_ext autoreload
# %autoreload 2
# # Call energySystemModelRunFromExcel function
# ### Read model from excel file, optimize and store to excel file
# Checkout the output excel file in the folder where this notebook is located
esM = fn.energySystemModelRunFromExcel()
# ### Read only
esM, esMData = fn.readEnergySystemModelFromExcel()
| examples/Model Run from Excel/Model Run from Excel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Tutorial Title
# A brief introduction to the tutorial that describes:
#
# - The problem that the tutorial addresses
# - Who the intended audience is
# - The expected experience level of that audience with a concept or tool
# - Which environment/language it runs in
#
# If there is another similar tutorial that's more appropriate for another audience, direct the reader there with a linked reference.
# ## How to Use This Tutorial
# A brief explanation of how the reader can use the tutorial. Can the reader copy each code snippet into a Python or other environment? Or can the reader run `<filename>` before or after reading through the explanations to understand how the code works?
# You can use this tutorial by *insert method(s) here*.
#
# A bulleted list of the tasks the reader will accomplish and skills he or she will learn. Begin each list item with a noun (Learn, Create, Use, etc.).
#
# You will accomplish the following:
#
# - First task or skill
# - Second task or skill
# - X task or skill
# ## Prerequisites
# Provide a *complete* list of the software, hardware, knowledge, and skills required to be successful using the tutorial. For each item, link the item to installation instructions, specs, or skill development tools, as appropriate. If good installation instructions aren't available for required software, start the tutorial with instructions for installing it.
# To complete this tutorial, you need:
#
# - [MXNet](https://mxnet.apache.org/install/#overview)
# - [Language](https://mxnet.apache.org/tutorials/)
# - [Tool](https://mxnet.apache.org/api/python/index.html)
# - [Familiarity with concept or tool](https://gluon.mxnet.io/)
#
# ## The Data
# Provide a link to where the data is hosted and explain how to download it. If it requires more than two steps, use a numbered list.
# You can download the data used in this tutorial from the [Site Name](http://) site. To download the data:
#
# 1. At the `<language>` prompt, type:
#
# `<command>`
# 2. Second task.
#
# 3. Last task.
# Briefly describe key aspects of the data. If there are two or more aspects of the data that require involved discussion, use subheads (### `<Concept or Sub-component Name>`). To include a graphic, introduce it with a brief description and use the image linking tool to include it. Store the graphic in GitHub and use the following format: <img width="517" alt="screen shot 2016-05-06 at 10 13 16 pm" src="https://cloud.githubusercontent.com/assets/5545640/15089697/d6f4fca0-13d7-11e6-9331-7f94fcc7b4c6.png">. You do not need to provide a title for your graphics.
# The data *add description here. (optional)*
# ## (Optional) Concept or Component Name
# If concepts or components need further introduction, include this section. If there are two or more aspects of the concept or component that require involved discussion, use subheads (### Concept or Sub-component Name).
# ## Prepare the Data
# If appropriate, summarize the tasks required to prepare the data, defining and explaining key concepts.
# To prepare the data, *provide explanation here.*
# Use a numbered procedure to explain how to prepare the data. Add code snippets or blocks that show the code that the user must type or that is used for this task in the Jupyter Notebook. To include code snippets, precede each line of code with four spaces and two tick marks. Always introduce input or output with a description or context or result, followed by a colon.
# To prepare the data:
#
# 1.
#
# 2.
#
# 3.
# If there are any aspects of data preparation that require elaboration, add it here.
# ## Create the Model
# If appropriate, summarize the tasks required to create the model, defining and explaining key concepts.
# To create the model, *provide explanation here.*
# Use a numbered procedure to explain how to create the data. Add code snippets or blocks that show the code that the user must type or that is used for this task in the Jupyter Notebook. To include code snippets, precede each line of code with four spaces and two tick marks. Always introduce input or output with a description or context or result, followed by a colon.
# To create the model:
#
# 1.
#
# 2.
#
# 3.
# If there are any aspects of model creation that require elaboration, add it here.
# ## Fit the Model
# If appropriate, summarize the tasks required to fit the model, defining and explaining key concepts.
# To fit the model, *provide explanation here.*
# Use a numbered procedure to explain how to fit the model. Add code snippets or blocks that show the code that the user must type or that is used for this task in the Jupyter Notebook. To include code snippets, precede each line of code with four spaces and two tick marks. Always introduce input or output with a description or context or result, followed by a colon.
# To fit the model:
#
# 1.
#
# 2.
#
# 3.
# If there are any aspects of model fitting that require elaboration, add it here.
# ## Evaluate the Model
# If appropriate, summarize the tasks required to evaluate the model, defining and explaining key concepts.
# To evaluate the model, *provide explanation here.*
# Use a numbered procedure to explain how to evaluate the model. Add code snippets or blocks that show the code that the user must type or that is used for this task in the Jupyter Notebook. To include code snippets, precede each line of code with four spaces and two tick marks. Always introduce input or output with a description or context or result, followed by a colon.
# To evaluate the model:
#
# 1.
#
# 2.
#
# 3.
# If there are any aspects of model evaluation that require elaboration, add it here.
# ## (Optional) Additional Tasks
# If appropriate, summarize the tasks required to perform the task, defining and explaining key concepts.
# To *perform the task*, *provide explanation here.*
# Use a numbered procedure to explain how to perform the task. Add code snippets or blocks that show the code that the user must type or that is used for this task in the Jupyter Notebook. To include code snippets, precede each line of code with four spaces and two tick marks. Always introduce input or output with a description or context or result, followed by a colon.
# To *perform the task*:
#
# 1.
#
# 2.
#
# 3.
# If there are any aspects of model evaluation that require elaboration, add it here.
# ## Summary
# Briefly describe the end result of the tutorial and how the user can use it or modify it to customize it.
# ## Next Steps
# Provide a bulleted list of other documents, tools, or tutorials that further explain the concepts discussed in this tutorial or build on this tutorial. Start each list item with a brief description of a user task followed by the title of the destination site or topic that is formatted as a link.
# - For more information on *topic*, see [Site Name](http://).
# - To learn more about using *tool or task*, see [Topic Title](http://).
# - To experiment with *service*, *tool*, or *object*, see [Site Name](http://).
# - For a more advanced tutorial on *subject*, see [Tutorial Title](http://).
| example/MXNetTutorialTemplate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.1
# language: julia
# name: julia-1.4
# ---
# # Tests
function test_eq(a, b)
@assert a==b
end
test_eq(10,10)
test_eq(10,30)
# # Matmul
using Flux, Flux.Data.MNIST, Images
using Statistics
labels = MNIST.labels();
images = MNIST.images();
length(labels)
images[1:5]
size(hcat(float.(vec.(images[1:50000]))...))
weights = randn(784, 10);
bias = zeros(10);
size(weights)
c = [1,2,3]
size(c)
reshape(collect(c),1,3)
# ## Matmul
function matmul(a,b)
return a*b
end
c = [1 2 3; 4 1 6; 7 8 1]
@time matmul(c,c)
# ## Normalize
size(images[1])
28*28
function Normalize(x, m,s)
@show return (x.-m)/s
end
function preprocess(img)
return hcat(float.(vec.(img))...)'
end
# +
# x_train=preprocess(normalize(images[1:50000]));
# y_train=labels[1:50000];
# x_valid=preprocess(normalize(images[50001:60000]));
# y_valid=labels[50001:60000];
# -
x_train=preprocess(images[1:50000]);
y_train=labels[1:50000];
x_valid=preprocess(images[50001:60000]);
y_valid=labels[50001:60000];
train_mean,train_std = Float64.(mean(x_train)), Float64.(std(x_train))
x_train = Normalize(x_train,train_mean,train_std);
x_valid = Normalize(x_valid,train_mean,train_std);
@show size(x_train)
@show size(y_train)
@show size(x_valid)
@show size(y_valid)
n,m = size(x_train)[1],size(x_train)[2]
c = maximum(y_train)+1
n,m,c
# # Basic arch
nh = 50
# +
w1= randn(m, nh)/sqrt(m);
b1 = zeros(nh);
w2 = randn(nh,1)/sqrt(n);
b2 = zeros(1);
# -
@info size(w1); @info size(b1); @info size(w2); @info size(b2)
mean(Float64.(x_valid)),std(Float64.(x_valid))
# ## Linear
function lin(x,w,b)
@show size(x)
@show size(w)
@show size(b)
@show size(x*w)
@show size((x*w).+b')
return (x*w).+b'
end
lin(x_valid,w1,b1);
# ## Relu
function relu(A)
A[A.<0.0] .= 0.0;
@show size(A)
return A.-0.5
end
@time relu([1.0,-1.0,-100000])
size(x_valid)
t = relu(lin(x_valid,w1,b1));
mean(t), std(t)
# ## Kaiming
nh = 50
m
# +
w1= randn(m, nh)*sqrt(2.0/m);
b1 = zeros(nh);
w2 = randn(nh,1)/sqrt(n);
b2 = zeros(1);
# -
@info size(w1); @info size(b1); @info size(w2); @info size(b2)
t = relu(lin(x_valid,w1,b1));
mean(t), std(t)
function model(xb)
l1 = lin(xb,w1,b1)
l2 = relu(l1)
l1 = lin(l2,w2,b2)
end
@time model(x_valid);
# ## MSE
size(model(x_valid))
size(reshape(randn(5000,1),(5000)))
function squeeze(x)
return reshape(x, size(x)[1])
end
size(squeeze(randn(500,1)))
function mse(output,targ)
return mean((squeeze(output).-targ).^2)
end
preds = model(x_train);
mse(preds, y_train)
# ## Gradients and backward pass
using AutoGrad
function mse(output,targ)
return mean((squeeze(output).-targ).^2)
end
function lin(x,w,b)
return (x*w).+b'
end
function model(xb)
l1 = lin(xb,w1,b1)
l2 = relu(l1)
l1 = lin(l2,w2,b2)
return l1
end
function weightFunc(h...; seed=nothing)
seed==nothing || srand(seed)
w = Any[]
x = 28*28
for y in [h..., 10]
push!(w, convert(Array{Float32}, 0.1*randn(y,x)))
push!(w, zeros(Float32, y))
x = y
end
return w
end
w=weightFunc()
# size(w)
# +
function forward_and_backward(inp,targ)
l1 = lin(inp, w1, b1)
l2 = relu(l1)
out = lin(l2, w2, b2)
loss = mse(out,targ)
return loss
end
# -
function model(xb)
l1 = lin(xb,w1,b1)
l2 = relu(l1)
l1 = lin(l2,w2,b2)
end
| streamlineAPI/fastaiFromScratch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: dviz
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Module 8: Histogram and CDF
#
# A deep dive into Histogram and boxplot.
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
import altair as alt
import pandas as pd
# %matplotlib inline
# + jupyter={"outputs_hidden": false}
import matplotlib
matplotlib.__version__
# -
# ## The tricky histogram with pre-counted data
# Let's revisit the table from the class
#
# | Hours | Frequency |
# |-------|-----------|
# | 0-1 | 4,300 |
# | 1-3 | 6,900 |
# | 3-5 | 4,900 |
# | 5-10 | 2,000 |
# | 10-24 | 2,100 |
# You can draw a histogram by just providing bins and counts instead of a list of numbers. So, let's try that.
bins = [0, 1, 3, 5, 10, 24]
data = {0.5: 4300, 2: 6900, 4: 4900, 7: 2000, 15: 2100}
# + jupyter={"outputs_hidden": false}
data.keys()
# -
# **Q: Draw histogram using this data.** Useful query: [Google search: matplotlib histogram pre-counted](https://www.google.com/search?client=safari&rls=en&q=matplotlib+histogram+already+counted&ie=UTF-8&oe=UTF-8#q=matplotlib+histogram+pre-counted)
# + jupyter={"outputs_hidden": false}
# TODO: draw a histogram with weighted data.
# -
# As you can see, the **default histogram does not normalize with binwidth and simply shows the counts**! This can be very misleading if you are working with variable bin width (e.g. logarithmic bins). So please be mindful about histograms when you work with variable bins.
#
# **Q: You can fix this by using the `density` option.**
# + jupyter={"outputs_hidden": false}
# TODO: fix it with density option.
# -
# ## Let's use an actual dataset
import vega_datasets
# + jupyter={"outputs_hidden": false}
movies = vega_datasets.data.movies()
movies.head()
# -
# Let's plot the histogram of IMDB ratings.
# + jupyter={"outputs_hidden": false}
plt.hist(movies['IMDB Rating'])
# -
# Did you get an error or a warning? What's going on?
#
# The problem is that the column contains `NaN` (Not a Number) values, which represent missing data points. The following command check whether each value is a `NaN` and returns the result.
# + jupyter={"outputs_hidden": false}
movies['IMDB Rating'].isna()
# -
# As you can see there are a bunch of missing rows. You can count them.
# + jupyter={"outputs_hidden": false}
sum(movies['IMDB Rating'].isna())
# -
# or drop them.
# + jupyter={"outputs_hidden": false}
IMDB_ratings_nan_dropped = movies['IMDB Rating'].dropna()
len(IMDB_ratings_nan_dropped)
# + jupyter={"outputs_hidden": false}
213 + 2988
# -
# The `dropna` can be applied to the dataframe too.
#
# **Q: drop rows from `movies` dataframe where either `IMDB_Rating` or `IMDB_Votes` is `NaN`.**
# TODO
# + jupyter={"outputs_hidden": false}
# Both should be zero.
print(sum(movies['IMDB Rating'].isna()), sum(movies['IMDB Votes'].isna()))
# -
# How does `matplotlib` decides the bins? Actually `matplotlib`'s `hist` function uses `numpy`'s `histogram` function under the hood.
# **Q: Plot the histogram of movie ratings (`IMDB_Rating`) using the `plt.hist()` function.**
# + jupyter={"outputs_hidden": false}
# TODO
# -
# Have you noticed that this function returns three objects? Take a look at the documentation [here](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.hist) to figure out what they are.
#
# To get the returned three objects:
# + jupyter={"outputs_hidden": false}
n_raw, bins_raw, patches = plt.hist(movies['IMDB Rating'])
print(n_raw)
print(bins_raw)
# -
# Here, `n_raw` contains the values of histograms, i.e., the number of movies in each of the 10 bins. Thus, the sum of the elements in `n_raw` should be equal to the total number of movies.
#
# **Q: Test whether the sum of values in `n_raw` is equal to the number of movies in the `movies` dataset**
# + jupyter={"outputs_hidden": false}
# TODO: test whether the sum of the numbers in n_raw is equal to the number of movies.
# -
# The second returned object (`bins_raw`) is a list containing the edges of the 10 bins: the first bin is \[1.4, 2.18\], the second \[2.18, 2.96\], and so on. What's the width of the bins?
# + jupyter={"outputs_hidden": false}
np.diff(bins_raw)
# -
# The width is same as the maximum value minus minimum value, divided by 10.
# + jupyter={"outputs_hidden": false}
min_rating = min(movies['IMDB Rating'])
max_rating = max(movies['IMDB Rating'])
print(min_rating, max_rating)
print( (max_rating-min_rating) / 10 )
# -
# Now, let's plot a normalized (density) histogram.
# + jupyter={"outputs_hidden": false}
n, bins, patches = plt.hist(movies['IMDB Rating'], density=True)
print(n)
print(bins)
# -
# The ten bins do not change. But now `n` represents the density of the data inside each bin. In other words, the sum of the area of each bar will equal to 1.
#
# **Q: Can you verify this?**
#
# Hint: the area of each bar is calculated as height * width. You may get something like 0.99999999999999978 instead of 1.
# + jupyter={"outputs_hidden": false}
# TODO
# -
# Anyway, these data generated from the `hist` function is calculated from `numpy`'s `histogram` function. https://docs.scipy.org/doc/numpy/reference/generated/numpy.histogram.html
#
# Note that the result of `np.histogram()` is same as that of `plt.hist()`.
# + jupyter={"outputs_hidden": false}
np.histogram(movies['IMDB Rating'])
# + jupyter={"outputs_hidden": false}
plt.hist(movies['IMDB Rating'])
# -
# If you look at the documentation, you can see that `numpy` uses simply 10 as the default number of bins. But you can set it manually or set it to be `auto`, which is the "Maximum of the `sturges` and `fd` estimators.". Let's try this `auto` option.
# + jupyter={"outputs_hidden": false}
_ = plt.hist(movies['IMDB Rating'], bins='auto')
# -
# ## Consequences of the binning parameter
#
# Let's explore the effect of bin size using small multiples. In `matplotlib`, you can use [subplot](https://www.google.com/search?client=safari&rls=en&q=matplotlib+subplot&ie=UTF-8&oe=UTF-8) to put multiple plots into a single figure.
#
# For instance, you can do something like:
# + jupyter={"outputs_hidden": false}
plt.figure(figsize=(10,5))
plt.subplot(1,2,1)
movies['IMDB Rating'].hist(bins=3)
plt.subplot(1,2,2)
movies['IMDB Rating'].hist(bins=20)
# -
# What does the argument in `plt.subplot(1,2,1)` mean? If you're not sure, check out: http://stackoverflow.com/questions/3584805/in-matplotlib-what-does-the-argument-mean-in-fig-add-subplot111
#
# **Q: create 8 subplots (2 rows and 4 columns) with the following `binsizes`.**
# + jupyter={"outputs_hidden": false}
nbins = [2, 3, 5, 10, 30, 40, 60, 100 ]
figsize = (18, 10)
# TODO
# -
# Do you see the issues with having too few bins or too many bins? In particular, do you notice weird patterns that emerge from `bins=30`?
#
# **Q: Can you guess why do you see such patterns? What are the peaks and what are the empty bars? What do they tell you about choosing the binsize in histograms?**
# + jupyter={"outputs_hidden": false}
# TODO: Provide your answer and evidence here
# -
# ## Formulae for choosing the number of bins.
#
# We can manually choose the number of bins based on those formulae.
# +
N = len(movies)
plt.figure(figsize=(12,4))
# Sqrt
nbins = int(np.sqrt(N))
plt.subplot(1,3,1)
plt.title("SQRT, {} bins".format(nbins))
movies['IMDB Rating'].hist(bins=nbins)
# Sturge's formula
nbins = int(np.ceil(np.log2(N) + 1))
plt.subplot(1,3,2)
plt.title("Sturge, {} bins".format(nbins))
movies['IMDB Rating'].hist(bins=nbins)
# Freedman-Diaconis
iqr = np.percentile(movies['IMDB Rating'], 75) - np.percentile(movies['IMDB Rating'], 25)
width = 2*iqr/np.power(N, 1/3)
nbins = int((max(movies['IMDB Rating']) - min(movies['IMDB Rating'])) / width)
plt.subplot(1,3,3)
plt.title("F-D, {} bins".format(nbins))
movies['IMDB Rating'].hist(bins=nbins)
# -
# But we can also use built-in formulae too. Let's try all of them.
# + jupyter={"outputs_hidden": false}
plt.figure(figsize=(20,4))
plt.subplot(161)
movies['IMDB Rating'].hist(bins='fd')
plt.subplot(162)
movies['IMDB Rating'].hist(bins='doane')
plt.subplot(163)
movies['IMDB Rating'].hist(bins='scott')
plt.subplot(164)
movies['IMDB Rating'].hist(bins='rice')
plt.subplot(165)
movies['IMDB Rating'].hist(bins='sturges')
plt.subplot(166)
movies['IMDB Rating'].hist(bins='sqrt')
# -
# Some are decent, but several of them tend to overestimate the good number of bins. As you have more data points, some of the formulae may overestimate the necessary number of bins. Particularly in our case, because of the precision issue, we shouldn't increase the number of bins too much.
# ### Then, how should we choose the number of bins?
# So what's the conclusion? use Scott's rule or Sturges' formula?
#
# No, I think the take-away is that you **should understand how the inappropriate number of bins can mislead you** and you should **try multiple number of bins** to obtain the most accurate picture of the data. Although the 'default' may work in most cases, don't blindly trust it! Don't judge the distribution of a dataset based on a single histogram. Try multiple parameters to get the full picture!
# ## CDF (Cumulative distribution function)
#
# Drawing a CDF is easy. Because it's very common data visualization, histogram has an option called `cumulative`.
# + jupyter={"outputs_hidden": false}
movies['IMDB Rating'].hist(cumulative=True)
# -
# You can also combine with options such as `histtype` and `density`.
# + jupyter={"outputs_hidden": false}
movies['IMDB Rating'].hist(histtype='step', cumulative=True, density=True)
# -
# And increase the number of bins.
# + jupyter={"outputs_hidden": false}
movies['IMDB Rating'].hist(cumulative=True, density=True, bins=1000)
# -
# This method works fine. By increasing the number of bins, you can get a CDF in the resolution that you want. But let's also try it manually to better understand what's going on. First, we should sort all the values.
# + jupyter={"outputs_hidden": false}
rating_sorted = movies['IMDB Rating'].sort_values()
rating_sorted.head()
# -
# We need to know the number of data points,
# + jupyter={"outputs_hidden": false}
N = len(rating_sorted)
N
# -
# And I think this may be useful for you.
# + jupyter={"outputs_hidden": false}
n = 50
np.linspace(1/n, 1.0, num=n)
# -
# **Q: now you're ready to draw a proper CDF. Draw the CDF plot of this data.**
# + jupyter={"outputs_hidden": false}
Y = np.linspace( 1/N, 1, num=N)
plt.xlabel("IMDB Rating")
plt.ylabel("CDF")
_ = plt.plot(rating_sorted,Y)
# -
# ## A bit more histogram with altair
#
# As you may remember, you can get a pandas dataframe from `vega_datasets` package and use it to create visualizations. But, if you use `altair`, you can simply pass the URL instead of the actual data.
# + jupyter={"outputs_hidden": false}
vega_datasets.data.movies.url
# + jupyter={"outputs_hidden": false}
# Choose based on your environment
#alt.renderers.enable('notebook')
#alt.renderers.enable('jupyterlab')
#alt.renderers.enable('default')
# -
# As mentioned before, in `altair` histogram is not special. It is just a plot that use bars (`mark_bar()`) where X axis is defined by `IMDB_Rating` with bins (`bin=True`), and Y axis is defined by `count()` aggregation function.
# + jupyter={"outputs_hidden": false}
alt.Chart(vega_datasets.data.movies.url).mark_bar().encode(
alt.X("IMDB Rating:Q", bin=True),
alt.Y('count()')
)
# -
# Have you noted that it is `IMDB_Rating:Q` not `IMDB_Rating`? This is a shorthand for
# + jupyter={"outputs_hidden": false}
alt.Chart(vega_datasets.data.movies.url).mark_bar().encode(
alt.X('IMDB Rating', type='quantitative', bin=True),
alt.Y(aggregate='count', type='quantitative')
)
# -
# In altair, you want to specify the data types using one of the four categories: quantitative, ordinal, nominal, and temporal. https://altair-viz.github.io/user_guide/encoding.html#data-types
# Although you can adjust the bins in `altair`, it does not encourage you to set the bins directly. For instance, although there is `step` parameter that directly sets the bin size, there are parameters such as `maxbins` (maximum number of bins) or `minstep` (minimum allowable step size), or `nice` (attemps to make the bin boundaries more human-friendly), that encourage you not to specify the bins directly.
# + jupyter={"outputs_hidden": false}
from altair import Bin
alt.Chart(vega_datasets.data.movies.url).mark_bar().encode(
alt.X("IMDB Rating:Q", bin=Bin(step=0.09)),
alt.Y('count()')
)
# + jupyter={"outputs_hidden": false}
alt.Chart(vega_datasets.data.movies.url).mark_bar().encode(
alt.X("IMDB Rating:Q", bin=Bin(nice=True, maxbins=20)),
alt.Y('count()')
)
# -
# ### Composing charts in altair
#
# `altair` has a very nice way to compose multiple plots. Two histograms side by side? just do the following.
chart1 = alt.Chart(vega_datasets.data.movies.url).mark_bar().encode(
alt.X("IMDB Rating:Q", bin=Bin(step=0.1)),
alt.Y('count()')
).properties(
width=300,
height=150
)
chart2 = alt.Chart(vega_datasets.data.movies.url).mark_bar().encode(
alt.X("IMDB Rating:Q", bin=Bin(nice=True, maxbins=20)),
alt.Y('count()')
).properties(
width=300,
height=150
)
# + jupyter={"outputs_hidden": false}
chart1 | chart2
# + jupyter={"outputs_hidden": false}
alt.hconcat(chart1, chart2)
# -
# Vertical commposition?
# + jupyter={"outputs_hidden": false}
alt.vconcat(chart1, chart2)
# + jupyter={"outputs_hidden": false}
chart1 & chart2
# -
# Shall we avoid some repetitions? You can define a *base* empty chart first and then assign encodings later when you put together multiple charts together. Here is an example: https://altair-viz.github.io/user_guide/compound_charts.html#repeated-charts
#
#
# Use base chart to produce the chart above:
# +
base = alt.Chart().mark_bar().encode(
alt.X("IMDB_Rating:Q", bin=Bin(nice=True, maxbins=20)),
alt.Y('count()')
).properties(
width=300,
height=150
)
chart = alt.vconcat(data=vega_datasets.data.movies.url)
for bin_param in [Bin(step=0.1), Bin(nice=True, maxbins=20)]:
row = alt.hconcat()
row |= base.encode(x=alt.X("IMDB Rating:Q", bin=bin_param), y='count()')
chart &= row
chart
# -
# **Q: Using the base chart approach to create a 2x2 chart where the top row shows the two histograms of `IMDB_Rating` with `maxbins`=10 and 50 respectively, and the bottom row shows another two histograms of `IMDB_Votes` with `maxbins`=10 and 50.**
# + jupyter={"outputs_hidden": false}
# TODO
| m08-histogram/m08-lab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table> <tr>
# <td style="background-color:#ffffff;">
# <a href="http://qworld.lu.lv" target="_blank"><img src="..\images\qworld.jpg" width="25%" align="left"> </a></td>
# <td style="background-color:#ffffff;vertical-align:bottom;text-align:right;">
# prepared by <NAME>, <NAME> and <NAME> (<a href="http://qworld.lu.lv/index.php/qturkey/" target="_blank">QTurkey</a>)
# </td>
# </tr></table>
# <table width="100%"><tr><td style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>
# $ \newcommand{\bra}[1]{\langle #1|} $
# $ \newcommand{\ket}[1]{|#1\rangle} $
# $ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
# $ \newcommand{\dot}[2]{ #1 \cdot #2} $
# $ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
# $ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
# $ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
# $ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
# $ \newcommand{\mypar}[1]{\left( #1 \right)} $
# $ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
# $ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
# $ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
# $ \newcommand{\onehalf}{\frac{1}{2}} $
# $ \newcommand{\donehalf}{\dfrac{1}{2}} $
# $ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
# $ \newcommand{\vzero}{\myvector{1\\0}} $
# $ \newcommand{\vone}{\myvector{0\\1}} $
# $ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
# $ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
# $ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
# $ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
# $ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
# $ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
# $ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
# $ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
# $ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
# <h1> Implementing Classical Gates Using Quantum Circuits </h1>
# In classical circuits, we use gates like $AND$, $OR$ and $NOT$. A gate set is universal if one can implement any boolean function $ f:\{0,1\}^n \rightarrow \{0,1\} $, using the gate set. An example universal set would be using $AND$, $OR$ and $NOT$ gates.
#
# - A $NOT$ gate takes one input $x_1 \in \{0,1\}$ and returns the negation of $x_1$.
#
# - An $AND$ gate takes two inputs $x_1,x_2 \in \{0,1\}$ and returns 1 if and only if both $x_1$ and $x_2$ are equal to 1, 0 otherwise.
#
# - An $OR$ gate takes two inputs $x_1, x_2 \in \{0, 1 \}$ and returns 1 if $x_1 = 1$ or $x_2 =1$.
#
#
# $
# NOT: \begin{array}{c|c} \mathbf{In} & \mathbf{Out} \\ \hline 0 & 1 \\ 1 & 0 \end{array}
# ~~~~~~~~
# AND: \begin{array}{cc|c} \mathbf{In} & \mathbf{In} & \mathbf{Out} \\ \hline 0 & 0 & 0 \\ 0 & 1 & 0 \\ 1 & 0 & 0 \\1 & 1 & 1 \end{array}
# ~~~~~~~~
# OR: \begin{array}{cc|c} \mathbf{In} & \mathbf{In} & \mathbf{Out} \\ \hline 0 & 0 & 0 \\ 0 & 1 & 1 \\ 1 & 0 & 1 \\1 & 1 & 1 \end{array}
# $
# <h3> Task 1 (Discuss) </h3>
#
# How can we implement $AND$ and $OR$ gates by a quantum circuit?
# ## Reversible Computing
# By looking at the output column of the tables of the $AND$ and $OR$ gates, we can not guess what the input is. We can say that the information or the entropy is lost by applying those gates and those operations are called **irreversible**. Irreversible copmutation dissipates heat to the environment.
#
# On the other hand, this is not the case for the $NOT$ gate as the input can be constructed by looking at the output. Such gates are called reversible and a computation which consists of only reversible operations is called a **reversible computation**.
#
# A set of gates is called **universal** if it is possible to implement any other gate using the gates in the set. Theoretically, it is possible to build a universal computer which only uses reversible gates. For instance, $AND$ and $NOT$ gates or the Toffoli gate ($CCNOT$) itself are universal sets of gates for classical computing. (Note that since $CCNOT$ is also a quantum gate, we conclude that any classical operation can be simulated by a quantum computer.)
# Since quantum computing is reversible according to the laws of physics, $AND$ and $OR$ gates should be implemented in a reversible way as well. The idea is to create a 3-qubit circuit, which does not modify the input bits and writes the output to the third bit. When the output bit is set to 0, then you exactly get the same output.
# <img src="../images/fcircuit.png" width="50%" align="center">
# <h3> Task 2 </h3>
# Complete the following table that corresponds to reversible $AND$ gate, where $\ket{x_1}$ and $\ket{x_2}$ are the inputs of the $AND$ gate and the $\ket{y} =0$ is the output. Which three-qubit quantum gate can we use to implement the $AND$ operator in a reversible manner?
# $
# AND: \begin{array}{ccc|ccc}
# \mathbf{In} & & &\mathbf{Out} & & \\
# \hline \mathbf{x_1} & \mathbf{x_2}& \mathbf{y} & \mathbf{x_1}& \mathbf{x_2}& \mathbf{y \oplus (x_1 \wedge x_2)} \\
# \hline 0 & 0 & 0 & & & \\
# \hline 0 & 1 & 0 & & & \\
# \hline 1 & 0 & 0 & & & \\
# \hline 1 & 1 & 0 & & & \\
# \end{array}
# $
# <a href="A01_Classical_Gates_Solutions.ipynb#task2">click for our solution</a>
# <h3> Task 3 </h3>
# Design a quantum (reversible) circuit for implementing $OR$ gate. Test your circuit on all possible two bit inputs. Measure only the output qubit.
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
for input in ['00','01','10','11']:
mycircuit1 = QuantumCircuit(3,1)
#
# Your code here
#
job = execute(mycircuit1,Aer.get_backend('qasm_simulator'),shots=1000)
counts = job.result().get_counts(mycircuit1)
print("Input:", input, "Output:", counts)
# -
# <a href="A01_Classical_Gates_Solutions.ipynb#task3">click for our solution</a>
# <h3>Task 4 </h3>
# Fredkin gate is a three qubit controlled swap gate which swaps the second and third qubits if the first qubit is in state $ \ket{1} $. It is represented with the following matrix.
# <img src="../images/fredkin.png" width="20%" align="center">
# Create a method named `fredkin` and implement Fredkin gate using $CNOT$ and $CCNOT$ operators. Use unitary simulator to check the unitary matrix corresponding to your circuit. (Follow the Qiskit order.)
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
def fredkin():
circuit = QuantumCircuit(3)
#
# Your code here
#
return circuit
# +
circuit = fredkin()
job = execute(circuit,Aer.get_backend('unitary_simulator'),shots=1)
u=job.result().get_unitary(circuit,decimals=3)
for i in range(len(u)):
s=""
for j in range(len(u)):
val = str(u[i][j].real)
while(len(val)<5): val = " "+val
s = s + val
print(s)
circuit.draw(output="mpl")
# -
# <a href="A01_Classical_Gates_Solutions.ipynb#task4">click for our solution</a>
# <h2> Implementing any Boolean Function </h2>
#
# Now having seen that we can implement a set of universal gates on a quantum computer, provided that we make them "reversible", we can say that it is possible to implement any boolean function $f:\{0,1\}^n \rightarrow \{0,1\}$. So given any boolean function $f(x)$, we propose that the following circuit will implement it in a quantum computer.
#
# <img src="../images/foperator.png" width="30%" align="center">
#
# Here $U_f$, the corresponding quantum operator, is defined as follows:
#
# $$U_f: \ket{x}\ket{y} \mapsto \ket{x}\ket{y \oplus f(x)} $$
#
# The symbol $\oplus$ denotes bitwise addition modulo 2 (XOR). This mapping is reversible although $f$ might not be invertible.
# <h3> Task 5</h3>
#
# Prove that $U_f$ is reversible, by showing that given $x$ and $y \oplus f(x)$, $y$ can be computed.
#
# _Hint: Use the fact that $x \oplus x = 0$ for any bit x and $\oplus$ operation is associative._
# <a href="A01_Classical_Gates_Solutions.ipynb#task5">click for our solution</a>
# <h3> Task 6</h3>
#
# After applying the operator $U_f$, what is the new state of the output qubit $\ket{y}$ in terms of $f(x)$ if initially:
#
# - $\ket{y} = \ket{0}$
#
# - $\ket{y} = \ket{1}$
#
# - $\ket{y} = \ket{-}$
# <a href="A01_Classical_Gates_Solutions.ipynb#task6">click for our solution</a>
| silver/.ipynb_checkpoints/A01_Classical_Gates-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Animation
from pythreejs import *
import ipywidgets
from IPython.display import display
# Reduce repo churn for examples with embedded state:
from pythreejs._example_helper import use_example_model_ids
use_example_model_ids()
view_width = 600
view_height = 400
# Let's first set up a basic scene with a cube and a sphere,
sphere = Mesh(
SphereBufferGeometry(1, 32, 16),
MeshStandardMaterial(color='red')
)
cube = Mesh(
BoxBufferGeometry(1, 1, 1),
MeshPhysicalMaterial(color='green'),
position=[2, 0, 4]
)
# as well as lighting and camera:
camera = PerspectiveCamera( position=[10, 6, 10], aspect=view_width/view_height)
key_light = DirectionalLight(position=[0, 10, 10])
ambient_light = AmbientLight()
# ## Keyframe animation
#
# The three.js animation system is built as a [keyframe](https://en.wikipedia.org/wiki/Key_frame) system. We'll demonstrate this by animating the position and rotation of our camera.
# First, we set up the keyframes for the position and the rotation separately:
positon_track = VectorKeyframeTrack(name='.position',
times=[0, 2, 5],
values=[10, 6, 10,
6.3, 3.78, 6.3,
-2.98, 0.84, 9.2,
])
rotation_track = QuaternionKeyframeTrack(name='.quaternion',
times=[0, 2, 5],
values=[-0.184, 0.375, 0.0762, 0.905,
-0.184, 0.375, 0.0762, 0.905,
-0.0430, -0.156, -0.00681, 0.987,
])
# Next, we create an animation clip combining the two tracks, and finally an animation action to control the animation. See the three.js docs for more details on the different responsibilities of the different classes.
camera_clip = AnimationClip(tracks=[positon_track, rotation_track])
camera_action = AnimationAction(AnimationMixer(camera), camera_clip, camera)
# Now, let's see it in action:
scene = Scene(children=[sphere, cube, camera, key_light, ambient_light])
controller = OrbitControls(controlling=camera)
renderer = Renderer(camera=camera, scene=scene, controls=[controller],
width=view_width, height=view_height)
# + tags=["nbval-ignore-output"]
renderer
# + tags=["nbval-ignore-output"]
camera_action
# -
# Let's add another animation clip, this time animating the color of the sphere's material:
# +
color_track = ColorKeyframeTrack(name='.material.color',
times=[0, 1], values=[1, 0, 0, 0, 0, 1]) # red to blue
color_clip = AnimationClip(tracks=[color_track], duration=1.5)
color_action = AnimationAction(AnimationMixer(sphere), color_clip, sphere)
# -
color_action
# Note how the two animation clips can freely be combined since they affect different properties. It's also worth noting that the color animation can be combined with manual camera control, while the camera animation cannot. When animating the camera, you might want to consider disabling the manual controls.
# ### Animating rotation
#
# When animating the camera rotation above, we used the camera's `quaternion`. This is the most robust method for animating free-form rotations. For example, the animation above was created by first moving the camera manually, and then reading out its `position` and `quaternion` properties at the wanted views. If you want more intuitive axes control, it is possible to animate the `rotation` sub-attributes instead, as shown below.
# +
f = """
function f(origu, origv, out) {
// scale u and v to the ranges I want: [0, 2*pi]
var u = 2*Math.PI*origu;
var v = 2*Math.PI*origv;
var x = Math.sin(u);
var y = Math.cos(v);
var z = Math.cos(u+v);
out.set(x,y,z)
}
"""
surf_g = ParametricGeometry(func=f, slices=16, stacks=16);
surf1 = Mesh(geometry=surf_g,
material=MeshLambertMaterial(color='green', side='FrontSide'))
surf2 = Mesh(geometry=surf_g,
material=MeshLambertMaterial(color='yellow', side='BackSide'))
surf = Group(children=[surf1, surf2])
camera2 = PerspectiveCamera( position=[10, 6, 10], aspect=view_width/view_height)
scene2 = Scene(children=[surf, camera2,
DirectionalLight(position=[3, 5, 1], intensity=0.6),
AmbientLight(intensity=0.5)])
renderer2 = Renderer(camera=camera2, scene=scene2,
controls=[OrbitControls(controlling=camera2)],
width=view_width, height=view_height)
display(renderer2)
# -
spin_track = NumberKeyframeTrack(name='.rotation[y]', times=[0, 2], values=[0, 6.28])
spin_clip = AnimationClip(tracks=[spin_track])
spin_action = AnimationAction(AnimationMixer(surf), spin_clip, surf)
spin_action
# Note that we are spinning the object itself, and that we are therefore free to manipulate the camera at will.
# ## Morph targets
#
# Set up a simple sphere geometry, and add a morph target that is an oblong pill shape:
# This lets three.js create the geometry, then syncs back vertex positions etc.
# For this reason, you should allow for the sync to complete before executing
# the next cell.
morph = BufferGeometry.from_geometry(SphereBufferGeometry(1, 32, 16))
# + tags=["nbval-skip"]
import numpy as np
# Set up morph targets:
vertices = np.array(morph.attributes['position'].array)
for i in range(len(vertices)):
if vertices[i, 0] > 0:
vertices[i, 0] += 1
morph.morphAttributes = {'position': [
BufferAttribute(vertices),
]}
morphMesh = Mesh(morph, MeshPhongMaterial(
color='#ff3333', shininess=150, morphTargets=True))
# -
# Set up animation for going back and forth between the sphere and pill shape:
# + tags=["nbval-skip"]
pill_track = NumberKeyframeTrack(
name='.morphTargetInfluences[0]', times=[0, 1.5, 3], values=[0, 2.5, 0])
pill_clip = AnimationClip(tracks=[pill_track])
pill_action = AnimationAction(AnimationMixer(morphMesh), pill_clip, morphMesh)
# + tags=["nbval-skip"]
camera3 = PerspectiveCamera( position=[5, 3, 5], aspect=view_width/view_height)
scene3 = Scene(children=[morphMesh, camera3,
DirectionalLight(position=[3, 5, 1], intensity=0.6),
AmbientLight(intensity=0.5)])
renderer3 = Renderer(camera=camera3, scene=scene3,
controls=[OrbitControls(controlling=camera3)],
width=view_width, height=view_height)
display(renderer3, pill_action)
# -
# ## Skeletal animation
#
# First, set up a skinned mesh with some bones:
# + tags=["nbval-skip"]
import numpy as np
N_BONES = 3
ref_cylinder = CylinderBufferGeometry(5, 5, 50, 5, N_BONES * 5, True)
cylinder = BufferGeometry.from_geometry(ref_cylinder)
# + tags=["nbval-skip"]
skinIndices = []
skinWeights = []
vertices = cylinder.attributes['position'].array
boneHeight = ref_cylinder.height / (N_BONES - 1)
for i in range(vertices.shape[0]):
y = vertices[i, 1] + 0.5 * ref_cylinder.height
skinIndex = y // boneHeight
skinWeight = ( y % boneHeight ) / boneHeight
# Ease between each bone
skinIndices.append([skinIndex, skinIndex + 1, 0, 0 ])
skinWeights.append([1 - skinWeight, skinWeight, 0, 0 ])
cylinder.attributes = dict(
cylinder.attributes,
skinIndex=BufferAttribute(skinIndices),
skinWeight=BufferAttribute(skinWeights),
)
shoulder = Bone(position=(0, -25, 0))
elbow = Bone(position=(0, 25, 0))
hand = Bone(position=(0, 25, 0))
shoulder.add(elbow)
elbow.add(hand)
bones = [shoulder, elbow, hand]
skeleton = Skeleton(bones)
mesh = SkinnedMesh(cylinder, MeshPhongMaterial(side='DoubleSide', skinning=True))
mesh.add(bones[0])
mesh.skeleton = skeleton
# + tags=["nbval-skip"]
helper = SkeletonHelper(mesh)
# -
# Next, set up some simple rotation animations for the bones:
# + tags=["nbval-skip"]
# Rotate on x and z axes:
bend_tracks = [
NumberKeyframeTrack(
name='.bones[1].rotation[x]',
times=[0, 0.5, 1.5, 2],
values=[0, 0.3, -0.3, 0]),
NumberKeyframeTrack(
name='.bones[1].rotation[z]',
times=[0, 0.5, 1.5, 2],
values=[0, 0.3, -0.3, 0]),
NumberKeyframeTrack(
name='.bones[2].rotation[x]',
times=[0, 0.5, 1.5, 2],
values=[0, -0.3, 0.3, 0]),
NumberKeyframeTrack(
name='.bones[2].rotation[z]',
times=[0, 0.5, 1.5, 2],
values=[0, -0.3, 0.3, 0]),
]
bend_clip = AnimationClip(tracks=bend_tracks)
bend_action = AnimationAction(AnimationMixer(mesh), bend_clip, mesh)
# Rotate on y axis:
wring_tracks = [
NumberKeyframeTrack(name='.bones[1].rotation[y]', times=[0, 0.5, 1.5, 2], values=[0, 0.7, -0.7, 0]),
NumberKeyframeTrack(name='.bones[2].rotation[y]', times=[0, 0.5, 1.5, 2], values=[0, 0.7, -0.7, 0]),
]
wring_clip = AnimationClip(tracks=wring_tracks)
wring_action = AnimationAction(AnimationMixer(mesh), wring_clip, mesh)
# + tags=["nbval-skip"]
camera4 = PerspectiveCamera( position=[40, 24, 40], aspect=view_width/view_height)
scene4 = Scene(children=[mesh, helper, camera4,
DirectionalLight(position=[3, 5, 1], intensity=0.6),
AmbientLight(intensity=0.5)])
renderer4 = Renderer(camera=camera4, scene=scene4,
controls=[OrbitControls(controlling=camera4)],
width=view_width, height=view_height)
display(renderer4)
# + tags=["nbval-skip"]
bend_action
# + tags=["nbval-skip"]
wring_action
# -
| examples/Animation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/AdityaKane2001/transformer2017/blob/main/transformer2017.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="YR7EwdYdzB6S"
import tensorflow as tf
from tensorflow.keras import layers
from tensorflow import keras
# + id="KDZCQPm-zJxV"
MAX_SEQ_LEN = 64
# BATCH_SIZE = 25000
DROPOUT_RATE = 0.1
EMBEDDING_DIMS = 512
VOCABULARY_SIZE = 4096
N_TRANSFORMERS = 6
FFNN_DIMS = 2048
NUM_HEADS = 8
KEY_DIMS = EMBEDDING_DIMS/ NUM_HEADS
VALUE_DIMS = EMBEDDING_DIMS/ NUM_HEADS
"""
Input pipeline:
1. We get batch_size number of pairs of sentences from the dataset:
batch_size x ("My name is <NAME>", "<start> Ich bin <NAME> <end>")
2. These sentences are then tokenized:
batch_size x ([2,3,4,5,6],[1,15,7,8,9,1000])
3. The sentences are then padded to the largest sentence:
batch_size x ([2,3,4,5,6,0,0,0,0], [1,15,7,8,9,1000,0,0,0])
4. They are then converted to embeddings:
batch_size x max_seq_len x embedding_dims
5. Add positional embeddings to this
batch_size x max_seq_len x embedding_dims
This is the input to our model.
"""
class PositionAwareEmbeddings(layers.Layer):
def __init__(self):
super().__init__()
self.dropout = layers.Dropout(DROPOUT_RATE)
self.embed_dims = EMBEDDING_DIMS
self.vocab_size = VOCABULARY_SIZE
self.embeddings = layers.Embedding(VOCABULARY_SIZE, EMBEDDING_DIMS,
input_length=MAX_SEQ_LEN)
self.max_seq_len = MAX_SEQ_LEN
def get_positional_embeddings(self, input_seq_len):
positions = tf.reshape(tf.range(input_seq_len, dtype=tf.double), (input_seq_len,1))
freqs = tf.math.pow(10000,
-tf.range(0, self.embed_dims, delta=2) / self.embed_dims)
sin_embs = tf.transpose(tf.cast(tf.math.sin(positions * freqs), tf.float32))
cos_embs = tf.transpose(tf.cast(tf.math.cos( positions* freqs), tf.float32))
expanded_sin_embs = tf.scatter_nd(
indices = [[i] for i in range(512) if i%2==1],
updates = sin_embs,
shape = ( self.embed_dims, input_seq_len)
)
expanded_cos_embs = tf.scatter_nd(
indices = [[i] for i in range(512) if i%2==0],
updates = cos_embs,
shape = ( self.embed_dims, input_seq_len)
)
pos_embs = tf.transpose(expanded_sin_embs + expanded_cos_embs)
return pos_embs #, expanded_sin_embs,expanded_cos_embs
def call(self, inputs):
input_seq_len = inputs.shape[-1]
pos_emb = self.get_positional_embeddings(input_seq_len)
outputs = self.embeddings(inputs)
outputs += pos_emb
return outputs
class MultiheadAttention(layers.Layer):
def __init__(self):
super().__init__()
self.heads = NUM_HEADS
def call(self, inputs):
pass
class ResidualAddNormMHA(layers.layer):
def __init__(self):
super().__init__()
pass
def call(self, inputs):
pass
# + id="5MgnAYQn0PoM"
pe = PositionAwareEmbeddings()
pos_embs, sin, cos = pe.get_positional_embeddings(100)
# print(pos_embs.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="BdV_bCoSh8Su" outputId="102ac24e-70c9-4442-92d3-0ac395e8fd2c"
tf.constant([[[5, 5, 5, 5], [6, 6, 6, 6],
[7, 7, 7, 7], [8, 8, 8, 8]],
[[5, 5, 5, 5], [6, 6, 6, 6],
[7, 7, 7, 7], [8, 8, 8, 8]]]).shape
# + id="RNHFd7plmjw2" colab={"base_uri": "https://localhost:8080/"} outputId="cc2d7bc4-e56f-4c6a-f72b-68024646855d"
import pathlib
text_file = tf.keras.utils.get_file(
fname="spa-eng.zip",
origin="http://storage.googleapis.com/download.tensorflow.org/data/spa-eng.zip",
extract=True,
)
text_file = pathlib.Path(text_file).parent / "spa-eng" / "spa.txt"
# + id="qoPL65kIKJ1D"
with open(text_file) as f:
lines = f.read().split("\n")[:-1]
text_pairs = []
for line in lines:
eng, spa = line.split("\t")
spa = "[start] " + spa + " [end]"
text_pairs.append((eng, spa))
# + colab={"base_uri": "https://localhost:8080/"} id="8y53hLaoKW5y" outputId="6fd50fda-3a08-42c9-ed5e-42962a4a1608"
import random
for _ in range(5):
print(random.choice(text_pairs))
# + id="Aq8IyRnDKaqr" outputId="9fc53b30-5fad-4c13-947f-87bf69c66a27" colab={"base_uri": "https://localhost:8080/"}
print(len(text_pairs
))
# + id="5kwu8V0RKf4a"
| transformer2017.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.chdir('..')
# +
import torch
import numpy as np
from util.misc import load
from model.videopose import TemporalModel
from model.pose_refinement import abs_to_hiprel, gmloss, capped_l2, capped_l2_euc_err, step_zero_velocity_loss
from databases.joint_sets import MuPoTSJoints
from databases.datasets import PersonStackedMuPoTsDataset
from training.preprocess import get_postprocessor, SaveableCompose, MeanNormalize3D
from training.callbacks import TemporalMupotsEvaluator
from training.loaders import UnchunkedGenerator
from training.torch_tools import get_optimizer
LOG_PATH = "../models"
# -
model_name = "29cbfa0fc1774b9cbb06a3573b7fb711"
def load_model(model_folder):
config = load(os.path.join(LOG_PATH, model_folder, "config.json"))
path = os.path.join(LOG_PATH, model_folder, "model_params.pkl")
# Input/output size calculation is hacky
weights = torch.load(path)
num_in_features = weights["expand_conv.weight"].shape[1]
m = TemporalModel(
num_in_features,
MuPoTSJoints.NUM_JOINTS,
config["model"]["filter_widths"],
dropout=config["model"]["dropout"],
channels=config["model"]["channels"],
layernorm=config["model"]["layernorm"],
)
m.cuda()
m.load_state_dict(weights)
m.eval()
return config, m
def get_dataset(config):
return PersonStackedMuPoTsDataset(
config["pose2d_type"],
config.get("pose3d_scaling", "normal"),
pose_validity="all",
)
config, model = load_model(model_name)
test_set = get_dataset(config)
def extract_post(model_name, test_set):
params_path = os.path.join(LOG_PATH, str(model_name), "preprocess_params.pkl")
transform = SaveableCompose.from_file(params_path, test_set, globals())
test_set.transform = transform
assert isinstance(transform.transforms[1].normalizer, MeanNormalize3D)
normalizer3d = transform.transforms[1].normalizer
return get_postprocessor(config, test_set, normalizer3d)
post_process_func = extract_post(model_name, test_set)
pad = (model.receptive_field() - 1) // 2
generator = UnchunkedGenerator(test_set, pad, True)
seqs = sorted(np.unique(test_set.index.seq))
preds = {}
# losses = {}
for i, (pose2d, valid) in enumerate(generator):
seq = seqs[i]
pred3d = model(torch.from_numpy(pose2d).cuda())
valid = valid[0]
# losses[seq] = self.loss(pred3d[0][valid], self.preprocessed3d[seq]) # .cpu().numpy()
pred_real_pose = post_process_func(pred3d[0], seq) # unnormalized output
pred_real_pose_aug = post_process_func(pred3d[1], seq)
pred_real_pose_aug[:, :, 0] *= -1
pred_real_pose_aug = test_set.pose3d_jointset.flip(pred_real_pose_aug)
pred_real_pose = (pred_real_pose + pred_real_pose_aug) / 2
preds[seq] = pred_real_pose[valid]
break
preds = TemporalMupotsEvaluator._group_by_seq(preds)
refine_config = load("../models/pose_refine_config.json")
pred = torch.cat(([preds[i] for i in range(1, 21)]))
# +
joint_set = MuPoTSJoints()
seqs = np.unique(test_set.index.seq)
losses = []
for seq in seqs:
inds = test_set.index.seq == seq # (20899,)
poses_pred = abs_to_hiprel(pred[inds], joint_set) / 1000 # (201, 17, 3)
# interpolate invisible poses, if required
poses_init = poses_pred.detach().clone()
kp_score = np.mean(test_set.poses2d[inds, :, 2], axis=-1) # (201,)
# if refine_config['smooth_visibility']:
# kp_score = ndimage.median_filter(kp_score, 9)
kp_score = torch.from_numpy(kp_score).cuda() # [201]
# poses_init = torch.from_numpy(poses_init).cuda() # [201, 17, 3]
# poses_pred = torch.from_numpy(poses_pred).cuda() # [201, 17, 3]
scale = torch.ones((len(kp_score), 1, 1)) # torch.Size([201, 1, 1])
poses_init.requires_grad = False
# poses_pred.requires_grad = True # TODO set to False
kp_score.requires_grad = False
scale.requires_grad = False
optimizer = get_optimizer(model.parameters(), refine_config)
for i in range(refine_config['num_iter']):
optimizer.zero_grad()
# smoothing formulation
if refine_config['pose_loss'] == 'gm':
pose_loss = torch.sum(kp_score.view(-1, 1, 1) * gmloss(poses_pred - poses_init, refine_config['gm_alpha']))
elif refine_config['pose_loss'] == 'capped_l2':
pose_loss = torch.sum(kp_score.view(-1, 1, 1) * capped_l2(poses_pred - poses_init,
torch.tensor(refine_config['l2_cap']).float().cuda()))
elif refine_config['pose_loss'] == 'capped_l2_euc_err':
pose_loss = torch.sum(kp_score.view(-1, 1) * capped_l2_euc_err(poses_pred, poses_init,
torch.tensor(refine_config['l2_cap']).float().cuda()))
else:
raise NotImplementedError('Unknown pose_loss' + refine_config['pose_loss'])
velocity_loss_hip = torch.sum(globals()[refine_config['smoothness_loss_hip']](poses_pred[:, [0], :], 1))
step = refine_config['smoothness_loss_hip_largestep']
vel_loss = globals()[refine_config['smoothness_loss_hip']](poses_pred[:, [0], :], step)
velocity_loss_hip_large = torch.sum((1 - kp_score[-len(vel_loss):]) * vel_loss)
velocity_loss_rel = torch.sum(globals()[refine_config['smoothness_loss_rel']](poses_pred[:, 1:, :], 1))
vel_loss = globals()[refine_config['smoothness_loss_rel']](poses_pred[:, 1:, :], step)
velocity_loss_rel_large = torch.sum((1 - kp_score[-len(vel_loss):]) * vel_loss)
total_loss = pose_loss + refine_config['smoothness_weight_hip'] * velocity_loss_hip \
+ refine_config['smoothness_weight_hip_large'] * velocity_loss_hip_large \
+ refine_config['smoothness_weight_rel'] * velocity_loss_rel \
+ refine_config['smoothness_weight_rel_large'] * velocity_loss_rel_large
# np.savez("pose_ref.npz",
# total_loss=total_loss.detach().cpu(),
# pose_loss=pose_loss.detach().cpu(),
# velocity_loss_hip=velocity_loss_hip.detach().cpu(),
# velocity_loss_hip_large=velocity_loss_hip_large.detach().cpu(),
# velocity_loss_rel=velocity_loss_rel.detach().cpu(),
# velocity_loss_rel_large=velocity_loss_rel_large.detach().cpu(),
# )
# exit()
total_loss.backward()
print(total_loss)
optimizer.step()
break
# poses_init = poses_init.detach().cpu().numpy() * 1000
# poses_pred = poses_pred.detach().cpu().numpy() * 1000
# poses_init = add_back_hip(poses_init, joint_set)
# poses_pred = add_back_hip(poses_pred, joint_set)
# smoothed_pred[inds] = poses_pred
# losses.append(total_loss.item())
break
# if _config.get('print_loss', False):
# print('Avg loss:', np.mean(losses))
# return smoothed_pred
# -
{'total_loss': total_loss.detach().cpu(),
'pose_loss': pose_loss.detach().cpu(),
'velocity_loss_hip': velocity_loss_hip.detach().cpu(),
'velocity_loss_hip_large': velocity_loss_hip_large.detach().cpu(),
'velocity_loss_rel': velocity_loss_rel.detach().cpu(),
'velocity_loss_rel_large': velocity_loss_rel_large.detach().cpu()}
| src/notebooks/04_00pure_nn_refine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Make-up assignment for XBUS 506: Visual Analytics
#
# Submit an end-to-end machine learning pipeline using Yellowbrick visualizations to support feature analysis (example here) and modeling (example here). Create a Jupyter notebook that:
# - Downloads a new data set from the UCI machine learning repository (Note: this should be a dataset you have not already explored in class as part of the certificate program).
# - Loads the data.
# - Performs a visual exploration of the data using Yellowbrick, annotated with text to explain your observations.
# - Fits and visually compares two or more Scikit-Learn models using Yellowbrick, and identifies the best performing model, providing textual descriptions to explain your reasoning.
# - Provides a brief conclusion with next steps that you would take (e.g. identifying one of the hyperparameters of your best performing model that you would experiment with to try to improve it's performance, explaining a cross-validation strategy, etc).
#
#
# ## default of credit card clients UCI Data Set
#
# Source:
#
# Name: <NAME>
# email addresses: (1) icyeh '@' chu.edu.tw (2) 140910 '@' mail.tku.edu.tw
# institutions: (1) Department of Information Management, Chung Hua University, Taiwan. (2) Department of Civil Engineering, Tamkang University, Taiwan.
# other contact information: 886-2-26215656 ext. 3181
#
#
# Data Set Information:
#
# This research aimed at the case of customers’ default payments in Taiwan and compares the predictive accuracy of probability of default among six data mining methods. From the perspective of risk management, the result of predictive accuracy of the estimated probability of default will be more valuable than the binary result of classification - credible or not credible clients. Because the real probability of default is unknown, this study presented the novel “Sorting Smoothing Method†to estimate the real probability of default. With the real probability of default as the response variable (Y), and the predictive probability of default as the independent variable (X), the simple linear regression result (Y = A + BX) shows that the forecasting model produced by artificial neural network has the highest coefficient of determination; its regression intercept (A) is close to zero, and regression coefficient (B) to one. Therefore, among the six data mining techniques, artificial neural network is the only one that can accurately estimate the real probability of default.
#
#
# Attribute Information:
#
# This research employed a binary variable, default payment (Yes = 1, No = 0), as the response variable. This study reviewed the literature and used the following 23 variables as explanatory variables:
# - LIMIT_BAL: Amount of the given credit (NT dollar): it includes both the individual consumer credit and his/her family (supplementary) credit.
# - SEX: Gender (1 = male; 2 = female).
# - Education: (1 = graduate school; 2 = university; 3 = high school; 4 = others).
# - MARRIAGE: Marital status (1 = married; 2 = single; 3 = others).
# - AGE: Age (year).
# - PAY: History of past payment. We tracked the past monthly payment records (from April to September, 2005) as follows: X6 = the repayment status in September, 2005; X7 = the repayment status in August, 2005; . . .;X11 = the repayment status in April, 2005. The measurement scale for the repayment status is: -1 = pay duly; 1 = payment delay for one month; 2 = payment delay for two months; . . .; 8 = payment delay for eight months; 9 = payment delay for nine months and above.
# - BILL_AMT: Amount of bill statement (NT dollar). X12 = amount of bill statement in September, 2005; X13 = amount of bill statement in August, 2005; . . .; X17 = amount of bill statement in April, 2005.
# - PAY_AMT: Amount of previous payment (NT dollar). X18 = amount paid in September, 2005; X19 = amount paid in August, 2005; . . .;X23 = amount paid in April, 2005.
# +
# %matplotlib inline
import os
import json
import time
import pickle
import requests
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings('ignore')
# -
# ## Data ingestion from the UCI repository
#
# Here we are retrieving data from the UCI repository. We do this by:
# - Write a function using os and requests
# - Define the URL
# - Define the file name
# - Define the location
# - Execute the function to fetch the data and save as CSV
# +
URL = "https://archive.ics.uci.edu/ml/machine-learning-databases/00350/default of credit card clients.xls"
def fetch_data(fname='cc_default.xls'):
"""
Helper method to retreive the ML Repository dataset.
"""
response = requests.get(URL)
outpath = os.path.abspath(fname)
with open(outpath, 'wb') as f:
f.write(response.content)
return outpath
# Fetch the data if required
DATA = fetch_data()
# -
print(DATA)
# +
FEATURES = [
"ID",
"LIMIT_BAL",
"SEX",
"EDUCATION",
"MARRIAGE",
"AGE",
"PAY_0",
"PAY_2",
"PAY_3",
"PAY_4",
"PAY_5",
"PAY_6",
"BILL_AMT1",
"BILL_AMT2",
"BILL_AMT3",
"BILL_AMT4",
"BILL_AMT5",
"BILL_AMT6",
"PAY_AMT1",
"PAY_AMT2",
"PAY_AMT3",
"PAY_AMT4",
"PAY_AMT5",
"PAY_AMT6",
"default_payment_next_month"
]
LABEL_MAP = {
1: "Yes",
0: "No",
}
# Read the data into a DataFrame
df = pd.read_excel(DATA, skiprows=2, header=None, names=FEATURES, index_col="ID")
# Convert class labels into text
df["default_payment_next_month"] = df["default_payment_next_month"].map(LABEL_MAP)
# Describe the dataset
print(df.describe())
df.columns
# -
df.head()
df.info()
# +
# Determine the shape of the data
print("{} instances with {} features\n".format(*df.shape))
# Determine the frequency of each class
print(df.groupby("default_payment_next_month")["default_payment_next_month"].count())
# -
# Replacing any null values with the median
df.fillna(df.median(),inplace=True)
print(df.describe())
# +
plt.figure(figsize=(4,3))
sns.countplot(data = df, x = "EDUCATION")
plt.figure(figsize=(4,3))
sns.countplot(data = df, x = "MARRIAGE")
plt.figure(figsize=(15,5))
sns.countplot(data = df, x = "AGE")
# -
# The first bar chart above shows us that the majority of people in this dataset have a college education.
#
# The next bar chart depicts that more people in this dataset are single.
#
# The last graph tells us that ages in this dataset ranges from 21 to ~75.
sns.countplot(y="EDUCATION", hue="default_payment_next_month", data=df,)
sns.countplot(y="MARRIAGE", hue="default_payment_next_month", data=df,)
plt.figure(figsize=(15,5))
sns.countplot(x="AGE", hue="default_payment_next_month", data=df,)
# The three bar chart above explore the same features as before, education, marriage, and age, only this time we've juxtaposed our target, defaulting on next month's payment.
# ### Data Wrangling
#
# Here we're using Scikit-Learn transformers to prepare data for ML. The sklearn.preprocessing package provides utility functions and transformer classes to help us transform input data so that it is better suited for ML. Here we're using LabelEncoder to encode the "default_payment" variable with a value between 0 and n_classes-1.
#
# +
from sklearn.preprocessing import LabelEncoder
# Extract our X and y data
X = df[FEATURES[1:-1]]
y = df["default_payment_next_month"]
# Encode our target variable
encoder = LabelEncoder().fit(y)
y = encoder.transform(y)
print(X.shape, y.shape)
# -
# ### Data Visualization
#
# Here we're using Pandas to create various visualizations of our data.
#
# First, I'm creating a matrix of scatter plots of the features in the dataset. This is useful for understanding how our features interact with eachother. For this section, I chose to only visualize 8 of the 23 features. This is useful for understanding how our features interact with eachother. I'm not sure I'm sensing any valuable insight from the scatter matrix below.
# Create a scatter matrix of the dataframe features
from pandas.plotting import scatter_matrix
scatter_matrix(df.ix[:,("LIMIT_BAL","SEX","EDUCATION","MARRIAGE","AGE","PAY_6","BILL_AMT6","PAY_AMT6",)]
, alpha=0.2, figsize=(12, 12), diagonal='kde')
plt.show()
# ### Rank Features
#
# Rank 2D is a two-dimensional ranking of features that utilizes a ranking algorithm that takes into account pairs of features at a time (e.g. joint plot analysis). The pairs of features are then ranked by score and visualized using the lower left triangle of a feature co-occurence matrix. The covariance and pearson plots below give us a better idea of which features we want to further explore.
# +
from yellowbrick.features import Rank2D
visualizer = Rank2D(algorithm="covariance", size=(1080, 720))
visualizer.fit_transform(X)
visualizer.poof()
# +
from yellowbrick.features import Rank2D
visualizer = Rank2D(algorithm="pearson", size=(1080, 720))
visualizer.fit_transform(X)
visualizer.poof()
# -
# ### RadViz
#
# RadViz is a multivariate data visualization algorithm that plots each feature dimension uniformly around the circumference of a circle then plots points on the interior of the circle such that the point normalizes its values on the axes from the center to each arc. This mechanism allows as many dimensions as will easily fit on a circle, greatly expanding the dimensionality of the visualization.
#
# Based on the Covariance and Pearson rankings, let's explore a few of the features with RadViz and Parallel Coordinates.
# +
# Extract our X and y data
X = df[FEATURES[1:12]]
y = df["default_payment_next_month"]
# Encode our target variable
encoder = LabelEncoder().fit(y)
y = encoder.transform(y)
# +
from yellowbrick.features import RadViz
# Specify the target classes
classes = [encoder.classes_]
# Instantiate the visualizer
visualizer = RadViz(size=(1080, 720))
visualizer.fit(X, y) # Fit the data to the visualizer
visualizer.transform(X) # Transform the data
visualizer.poof() # Draw/show/poof the data
# +
#Because the dataset is so large, I've decided to only take a 30% sample of the overall dataset.
from yellowbrick.features import ParallelCoordinates
_ = ParallelCoordinates(classes=encoder.classes_, normalize='standard', sample=0.3).fit_transform(X, y)
# -
#
# ### Data Extraction
# One way that we can structure our data for easy management is to save files on disk. The Scikit-Learn datasets are already structured this way, and when loaded into a Bunch (a class imported from the datasets module of Scikit-Learn) we can expose a data API that is very familiar to how we've trained on our toy datasets in the past. A Bunch object exposes some important properties:
#
# - data: array of shape n_samples * n_features
# - target: array of length n_samples
# - feature_names: names of the features
# - target_names: names of the targets
# - filenames: names of the files that were loaded
# - DESCR: contents of the readme
#
# Note: This does not preclude database storage of the data, in fact - a database can be easily extended to load the same Bunch API. Simply store the README and features in a dataset description table and load it from there. The filenames property will be redundant, but you could store a SQL statement that shows the data load.
#
# In order to manage our data set on disk, we'll structure our data as follows:
# +
from sklearn.datasets.base import Bunch
DATA_DIR = os.path.abspath(os.path.join( ".", "..", "Alternative Assignment"))
print(DATA_DIR)
# Show the contents of the data directory
for name in os.listdir(DATA_DIR):
if name.startswith("."): continue
print("- {}".format(name))
# +
def load_data(root=DATA_DIR):
# Construct the `Bunch` for the default dataset
filenames = {
'meta': os.path.join(root, 'meta.json'),
'rdme': os.path.join(root, 'README.md'),
'data': os.path.join(root, 'cc_default.xls'),
}
# Load the meta data from the meta json
with open(filenames['meta'], 'r') as f:
meta = json.load(f)
target_names = meta['target_names']
feature_names = meta['feature_names']
# Load the description from the README.
with open(filenames['rdme'], 'r') as f:
DESCR = f.read()
# Load the dataset from the excel file.
dataset = pd.read_excel('cc_default.xls', skiprows=2, header=None, names=FEATURES, index_col="ID")
# Extract the target from the data
data = dataset[[ "LIMIT_BAL", "SEX", "EDUCATION", "MARRIAGE","AGE", "PAY_0", "PAY_2", "PAY_3", "PAY_4", "PAY_5", "PAY_6", "BILL_AMT1", "BILL_AMT2", "BILL_AMT3", "BILL_AMT4", "BILL_AMT5", "BILL_AMT6", "PAY_AMT1", "PAY_AMT2", "PAY_AMT3","PAY_AMT4", "PAY_AMT5", "PAY_AMT6"]]
target = dataset["default_payment_next_month"]
# Create the bunch object
return Bunch(
data=data,
target=target,
filenames=filenames,
target_names=target_names,
feature_names=feature_names,
DESCR=DESCR
)
# Save the dataset as a variable we can use.
dataset = load_data()
print(dataset.data.shape)
print(dataset.target.shape)
# -
# ### Classification
#
# Now that we have a dataset Bunch loaded and ready, we can begin the classification process. Let's attempt to build a classifier with kNN, SVM, and Random Forest classifiers.
#
# - Load the Algorithms!
# - Metrics for evaluating performance
# - K-Folds cross-validator provides train/test indices to split data in train/test sets.
# - SVC algorithm
# - K Neighbors Classifier
# - Random Forest Classifier
# - Logistic Regression
# +
from sklearn import metrics
from sklearn.model_selection import KFold
from sklearn.svm import SVC
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.pipeline import Pipeline
from sklearn.metrics import f1_score
from sklearn.preprocessing import OneHotEncoder
from yellowbrick.classifier import ClassificationReport
# -
# Define a function to evaluate the performance of the models
# - Set our start time
# - Define an empty array for our scores variable
# - Define our training dataset and our test dataset
# - Define estimator and fit to data
# - Define predictor and set to data
# - Calculate metrics for evaluating models
# - Print evaluation report
# - Write estimator to disc for future predictions
# - Save model
def fit_and_evaluate(dataset, model, label, **kwargs):
"""
Because of the Scikit-Learn API, we can create a function to
do all of the fit and evaluate work on our behalf!
"""
start = time.time() # Start the clock!
scores = {'precision':[], 'recall':[], 'accuracy':[], 'f1':[]}
kf = KFold(n_splits = 12, shuffle=True)
for train, test in kf.split(dataset.data):
X_train, X_test = dataset.data.iloc[train], dataset.data.iloc[test]
y_train, y_test = dataset.target.iloc[train], dataset.target.iloc[test]
estimator = model(**kwargs)
estimator.fit(X_train, y_train)
expected = y_test
predicted = estimator.predict(X_test)
# Append our scores to the tracker
scores['precision'].append(metrics.precision_score(expected, predicted, average="weighted"))
scores['recall'].append(metrics.recall_score(expected, predicted, average="weighted"))
scores['accuracy'].append(metrics.accuracy_score(expected, predicted))
scores['f1'].append(metrics.f1_score(expected, predicted, average="weighted"))
# Report
print("Build and Validation of {} took {:0.3f} seconds".format(label, time.time()-start))
print("Validation scores are as follows:\n")
print(pd.DataFrame(scores).mean())
# Write official estimator to disk in order to use for future predictions on new data
estimator = model(**kwargs)
estimator.fit(dataset.data, dataset.target)
#saving model with the pickle model
outpath = label.lower().replace(" ", "-") + ".pickle"
with open(outpath, 'wb') as f:
pickle.dump(estimator, f)
print("\nFitted model written to:\n{}".format(os.path.abspath(outpath)))
# Perform SVC Classification
fit_and_evaluate(dataset, SVC, "Default SVM Classifier", gamma = 'auto')
# Perform kNN Classification
fit_and_evaluate(dataset, KNeighborsClassifier, "Default kNN Classifier", n_neighbors=12)
# Perform Random Forest Classification
fit_and_evaluate(dataset, RandomForestClassifier, "Default Random Forest Classifier")
fit_and_evaluate(dataset, LogisticRegression, "Default Logistic Regression")
# Creating a function to visualize estimators
def visual_fit_and_evaluate(X, y, estimator):
visualizer = ClassificationReport(estimator, classes=['No', 'Yes'], cmap='PRGn')
visualizer.fit(X, y)
visualizer.score(X, y)
visualizer.poof()
visual_fit_and_evaluate(X, y, SVC())
visual_fit_and_evaluate(X, y, KNeighborsClassifier())
visual_fit_and_evaluate(X, y, RandomForestClassifier(class_weight='balanced'))
visual_fit_and_evaluate(X, y, LogisticRegression(class_weight='balanced'))
# ## Conclusion
#
#
# While all estimators seem to be decent predictors, I believe the Random Forest classifier was the best in the preliminary fit and evaluation. This is because it has the highest F1 score, which is a measure of the test's accuracy taking into account both the precision and the recall. When it came to the visual fit and evaluation, the Random Forest classifier still seems to be the best model. However, all of the models yielded much higher F1 scores, especially the Random Forest classifier coming in at .96, which seems suspiciously high!
#
# The next thing to do is to look into is how generalizable our model is, as we seem to be in danger of overfitting our data. I would experiment with cross-validation for the Random Forest Classier to assess how generalizable the model is as well as grid searching to ensure that I'm using the best parameters for each model.
| Assignment for Visual Analytics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/dpk-a7/Deep-learning/blob/main/lstm_toxic_comments.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="Zu3kIccGKiPf" outputId="d1f60d93-9c84-4755-f79e-64da79abf1f1"
# !wget http://nlp.stanford.edu/data/glove.6B.zip
# !pip install -q kaggle
# !pip install -q kaggle-cli
# !mkdir -p ~/.kaggle
# !cp "kaggle.json" ~/.kaggle/
# !cat ~/.kaggle/kaggle.json
# !chmod 600 ~/.kaggle/kaggle.json# For competition datasets
# !kaggle competitions download -c jigsaw-toxic-comment-classification-challenge
# + colab={"base_uri": "https://localhost:8080/"} id="RITc5TNaLSE9" outputId="4051d597-95eb-49b1-9555-588c8f82d2b3"
# !unzip glove.6B.zip
# !unzip train.csv.zip
# !unzip test.csv.zip
# + id="rRf5HnUTLSPu"
import os
import sys
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from keras.models import Model
from keras.layers import Dense, Embedding, Input
from keras.layers import LSTM, Bidirectional, GlobalMaxPool1D, Dropout
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.optimizers import Adam
from sklearn.metrics import roc_auc_score
# + id="JLjoceoaMTXS"
MAX_SEQ_LENGTH = 100
MAX_VOCAB_SIZE = 20000
EMBEDDING_DIM = 100
VALIDATION_SPLIT = 0.2
BATCH_SIZE = 128
EPOCHS = 10
# + colab={"base_uri": "https://localhost:8080/"} id="g7XwGcQDM9y9" outputId="ed653cd8-e87a-499a-b994-edb29248ec72"
#loading word vectors..
word2vec = {} #key = word: value = vectors
with open(os.path.join("glove.6B.%sd.txt" % EMBEDDING_DIM)) as f:
for line in f:
values = line.split()
word = values[0]
vec = np.asarray(values[1:], dtype= 'float32')
word2vec[word] = vec
print("found %s word vectors" % len(word2vec))
# + colab={"base_uri": "https://localhost:8080/"} id="Yg2q7kfaM_5H" outputId="6ea519fe-16da-40f4-9e01-774abc826fdb"
word2vec['the']
# + colab={"base_uri": "https://localhost:8080/"} id="l5GfgU_kNDIt" outputId="0644747a-31dd-4007-c513-0c15ea78db17"
#Loading comments
train = pd.read_csv("train.csv")
train.columns
# + id="HMgL9OEPNGa4"
sentences = train["comment_text"].fillna("DUMMY_VALUE").values
possible_labels = ['toxic', 'severe_toxic', 'obscene',
'threat','insult', 'identity_hate']
targets = train[possible_labels].values
# + colab={"base_uri": "https://localhost:8080/"} id="cXiNPdD7NNw0" outputId="c6816718-5377-4f23-a236-ef5589f4cdd0"
print("max sequence length:",max(len(s)for s in sentences))
print("min sequence length:",min(len(s)for s in sentences))
s = sorted(len(s) for s in sentences)
print("median sequence length:", s[len(s)//2])
# + id="x3fwFl3QNIjk"
tokenizer = Tokenizer(num_words = MAX_VOCAB_SIZE)
tokenizer.fit_on_texts(sentences)
sequences = tokenizer.texts_to_sequences(sentences)
# + colab={"base_uri": "https://localhost:8080/"} id="xXWzG71LNObO" outputId="ec6fa307-be73-4175-8721-f9c7c7008e8c"
word2idx = tokenizer.word_index
print("found %s unique tokens" % len(word2idx))
# + colab={"base_uri": "https://localhost:8080/"} id="tQ1tSlGrNQx6" outputId="cadefdb5-9b89-4930-8774-91599906bb28"
data = pad_sequences(sequences, maxlen = MAX_SEQ_LENGTH)
print("Shape of data tensor:", data.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="Bo2UhHNMNTBb" outputId="01f731c8-cbef-4dae-b9a2-208c67152272"
print("Filling pre-trained embeddings..")
num_words = min(MAX_VOCAB_SIZE, len(word2idx)+1)
embedding_matrix = np.zeros((num_words, EMBEDDING_DIM))
for word, i in word2idx.items():
if i < MAX_VOCAB_SIZE:
embedding_vector = word2vec.get(word)
if embedding_vector is not None:
embedding_matrix[i] = embedding_vector
# + id="sJ4PCBG-NVHr"
embedding_layer = Embedding(
num_words,
EMBEDDING_DIM,
weights = [embedding_matrix],
input_length = MAX_SEQ_LENGTH,
trainable = False
)
# + id="s8vgKEWBNi_j"
# model build
input_ = Input(shape=(MAX_SEQ_LENGTH,))
x = embedding_layer(input_)
x = LSTM(15, return_sequences=True)(x)
# x = Bidirectional(LSTM(15, return_sequences=True))(x)
x = GlobalMaxPool1D()(x)
output = Dense(len(possible_labels), activation="sigmoid")(x)
model = Model(input_, output)
model.compile(
loss = "binary_crossentropy",
optimizer = Adam(learning_rate=0.01),
metrics=['accuracy']
)
# + colab={"base_uri": "https://localhost:8080/"} id="YYjlq8OVOo9d" outputId="e0f41d6d-7ca4-48fd-eb41-de2db94a2e40"
# Training model
r = model.fit(
data,
targets,
batch_size = BATCH_SIZE,
epochs= EPOCHS,
validation_split = VALIDATION_SPLIT
)
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="QumneMLRPNes" outputId="8711e339-4f09-49d7-cf58-fe2e02b7dcb9"
plt.plot(r.history['loss'], label='loss')
plt.plot(r.history['val_loss'], label='val_loss')
plt.legend()
plt.show();
# + colab={"base_uri": "https://localhost:8080/", "height": 268} id="kAfOM1hWPPnL" outputId="8438c6e8-86c9-46ca-f7b9-0c2d333f1349"
plt.plot(r.history['accuracy'], label='accuracy')
plt.plot(r.history['val_accuracy'], label='val_acc')
plt.legend()
plt.show();
# + colab={"base_uri": "https://localhost:8080/"} id="lMU2Ph-UPRdT" outputId="f5142585-8a8f-48a6-aa24-f9ca2c6f20d3"
p = model.predict(data)
aucs = []
del i
for i in range(5):
auc = roc_auc_score(targets[:,i], p[:,i])
aucs.append(auc)
print(np.mean(aucs)) # with cnn 0.9718710715825385
# + colab={"base_uri": "https://localhost:8080/"} id="YuGOkjGePTMh" outputId="24953d76-bfd9-4ec3-e858-06a47d2e698c"
def single_predict(sentence):
single_sequences = tokenizer.texts_to_sequences(sentence)
pad_input=[[j for i in single_sequences for j in i]]
single_data = pad_sequences(pad_input, maxlen=MAX_SEQ_LENGTH)
p = model.predict(single_data)
p = list(p)
# print(ans)
print(p)
for i in p[0]:
print(max([i]))
print('->',max(p[0]))
single_predict("very good")
# + colab={"base_uri": "https://localhost:8080/"} id="FEe4Nu9SPU-a" outputId="57e21922-c17d-4ba5-e129-c5b85baea815"
def single_predict(sentence):
single_sequences = tokenizer.texts_to_sequences(sentence)
single_data = pad_sequences(single_sequences, maxlen=MAX_SEQ_LENGTH)
p = model.predict(single_data)[-1]
lab = ["toxic", "severe_toxic", "obscene", "threat", "insult", "identity_hate"]
p = [i for i in p]
ans = lab[p.index(max(p))] if max(p) > 0.057 else "Neutral"
print(ans)
print(p)
single_predict("i am a bad boy")
# + colab={"base_uri": "https://localhost:8080/"} id="_mVK1_NsR8oM" outputId="9fd56725-714f-4593-f792-cc44016e5fa4"
single_sequences = tokenizer.texts_to_sequences("worst behaviour")
single_data = pad_sequences(single_sequences, maxlen=MAX_SEQ_LENGTH)
p = model.predict(single_data)
print(p)
# + colab={"base_uri": "https://localhost:8080/"} id="6VaQpem4S0Hf" outputId="4ada18f5-aee3-4923-93ed-5a325880a5a8"
single_sequences = tokenizer.texts_to_sequences("good boy")
single_data = pad_sequences(single_sequences, maxlen=MAX_SEQ_LENGTH)
p = model.predict(single_data)
print(p)
# + id="RwmqiGDlTuJt"
| lstm_toxic_comments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import and setting
# +
# ---------- import
import pickle
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# %matplotlib inline
# +
# ---------- figure size
plt.rcParams['figure.figsize'] =[8, 6]
# ---------- axes
plt.rcParams['axes.grid'] = True
plt.rcParams['axes.linewidth'] = 1.5
# ---------- ticks
plt.rcParams['xtick.direction'] = 'in'
plt.rcParams['ytick.direction'] = 'in'
plt.rcParams['xtick.major.width'] = 1.0
plt.rcParams['ytick.major.width'] = 1.0
plt.rcParams['xtick.major.size'] = 8.0
plt.rcParams['ytick.major.size'] = 8.0
# ---------- lines
plt.rcParams['lines.linewidth'] = 2.5
# ---------- grid
plt.rcParams['grid.linestyle'] = ':'
# ---------- font
plt.rcParams['font.size'] = 20
#plt.rcParams['pdf.fonttype'] = 42 # embed fonts in PDF using type42 (True type)
# -
# # Data
def load_pkl(filename):
with open(filename, 'rb') as f:
return pickle.load(f)
# +
rslt_data = load_pkl('./pkl_data/rslt_data.pkl')
# ---------- sort by Energy
rslt_data.sort_values(by=['E_eV_atom']).head(10)
# +
# ---------- Number of structures
ndata = len(rslt_data)
print('Number of data: {}'.format(ndata))
# ---------- check success and error
nsuccess = rslt_data['E_eV_atom'].count()
nerror = ndata - nsuccess
print('Success: {}'.format(nsuccess))
print('Error: {}'.format(nerror))
# ---------- minimum
Emin = rslt_data['E_eV_atom'].min()
print('Emin: {} eV/atom'.format(Emin))
# -
# # Energy vs. trial
# +
fig, ax = plt.subplots()
# ---------- axis
dx = 1
ax.set_xlim([0, ndata+dx])
ax.set_ylim([-0.5, 3])
# ---------- hline at zero
ax.hlines(0.0, -dx, ndata+dx, 'k', '--')
# ---------- plot
# x <-- ID + 1
ax.plot(rslt_data.index + 1, rslt_data['E_eV_atom'] - Emin, 'o', ms=15, mew=2.0, alpha=0.8)
# ---------- title and label
ax.set_title('Random search for Na$_8$Cl$_8$')
ax.set_xlabel('Number of trials')
ax.set_ylabel('Energy (eV/atom)')
# -
# ---------- save figure
fig.savefig('Na8Cl8_RS.png', bbox_inches='tight') # PNG
#fig.savefig('title_RS.png', bbox_inches='tight', dpi=300) # high dpi PNG
#fig.savefig('title_RS.pdf', bbox_inches='tight') # PDF
| example/VASP_qsub_RS_Na8Cl8/with_results/data/cryspy_analyzer_RS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/simecek/from0toheroin2h/blob/master/colab3_dataset_from_Google_Images.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] hide_input=false id="3PspAfmCvY0H" colab_type="text"
# # Creating your own dataset from Google Images
#
# *adapted from a notebook by [<NAME> and <NAME>](https://github.com/fastai/course-v3/blob/master/nbs/dl1/lesson2-download.ipynb). Inspired by [<NAME>](https://www.pyimagesearch.com/2017/12/04/how-to-create-a-deep-learning-dataset-using-google-images/)*
# + [markdown] hide_input=true id="ukDEw_2avY0K" colab_type="text"
# In this tutorial we will see how to easily create an image dataset through Google Images.
# + [markdown] id="l8fw9PKNvY0R" colab_type="text"
# ## Get a list of URLs
#
# **Note**: You will have to repeat these steps for any new category you want to Google (e.g once for dogs and once for cats).
# + [markdown] id="jAYHpdQmvY0S" colab_type="text"
# ### Search and scroll
# + [markdown] id="gWX36fXdvY0T" colab_type="text"
# Go to [Google Images](http://images.google.com) and search for the images you are interested in. The more specific you are in your Google Search, the better the results and the less manual pruning you will have to do.
#
# Scroll down until you've seen all the images you want to download, or until you see a button that says 'Show more results'. All the images you scrolled past are now available to download. To get more, click on the button, and continue scrolling. The maximum number of images Google Images shows is 700.
#
# It is a good idea to put things you want to exclude into the search query, for instance if you are searching for the Eurasian wolf, "canis lupus lupus", it might be a good idea to exclude other variants:
#
# "canis lupus lupus" -dog -arctos -familiaris -baileyi -occidentalis
#
# You can also limit your results to show only photos by clicking on Tools and selecting Photos from the Type dropdown.
# + [markdown] id="R_gmX7KAvY0U" colab_type="text"
# ### Download into file
# + [markdown] id="La5qHfUUvY0V" colab_type="text"
# Now you must run some Javascript code in your browser which will save the URLs of all the images you want for you dataset.
#
# Press `Ctrl`-`Shift`-`J` in Windows/Linux and `Cmd`-`Opt`-`J` in Mac, and a small window the javascript 'Console' will appear. That is where you will paste the JavaScript commands.
#
# You will need to get the urls of each of the images. You can do this by running the following commands (you might want to change `urls.txt` to `your_class_name.txt`):
#
# ```javascript
# javascript:document.body.innerHTML = `<a href="data:text/csv;charset=utf-8,${escape(Array.from(document.querySelectorAll('.rg_di .rg_meta')).map(el=>JSON.parse(el.textContent).ou).join('\n'))}" download="urls.txt">download urls</a>`;
# ```
# + [markdown] id="FuCTZrawvY0W" colab_type="text"
# ### Upload urls file into your server
# + [markdown] id="aQ9QcqHXvY0X" colab_type="text"
# In the left panel, select `Files`, click on `UPLOAD` and choose your file with urls. Make sure names of your url files are formatted as `your_class_name.txt`.
# + [markdown] id="G428HheWvY0y" colab_type="text"
# ## Download images
# + hide_input=false id="R5v8fJv6vY0M" colab_type="code" colab={}
from fastai.vision import *
import os, shutil
# + [markdown] id="TPckJ-ev2cb0" colab_type="text"
# First, let us create a list of your classes and check that everything is as it is supposed to be.
# + id="d68lp16xvY00" colab_type="code" colab={}
# you need to change classes to your classes
classes = ['konvalinka','kopretina','pampeliska']
for class_name in classes:
assert os.path.isfile(class_name + '.txt')
# + [markdown] id="-OGoykLY3lpd" colab_type="text"
# If everything is ok, create a folder for each class and copy url files into them.
# + id="MBQZzft530NQ" colab_type="code" colab={}
data_folder = Path('data/')
for class_name in classes:
class_folder = data_folder/class_name
class_folder.mkdir(parents=True, exist_ok=True)
shutil.copy(class_name + '.txt', class_folder)
# + [markdown] id="ghgz3Ms9vY0z" colab_type="text"
# Now you will need to download your images from their respective urls.
#
# fast.ai has a function that allows you to do just that. You just have to specify the urls filename as well as the destination folder and this function will download and save all images that can be opened. If they have some problem in being opened, they will not be saved.
#
# Let's download our images! Notice you can choose a maximum number of images to be downloaded. In this case we will not download all the urls.
#
# You will need to run this line once for every category.
# + id="hN43CasRvY03" colab_type="code" colab={}
for class_folder in classes:
print(class_folder)
file_name = class_folder + ".txt"
download_images(data_folder/class_folder/file_name, data_folder/class_folder, max_pics=195)
# + [markdown] id="uNsw8IJYvY0_" colab_type="text"
# Then we can remove any images that can't be opened:
# + id="vx1wzXcuvY1A" colab_type="code" colab={}
for class_folder in classes:
print(class_folder)
verify_images(data_folder/class_folder, delete=True, max_size=500)
# + [markdown] id="oGfkPOqSvY1D" colab_type="text"
# ## View data
# + id="WQWgtZ7GvY1E" colab_type="code" colab={}
np.random.seed(42)
data = ImageDataBunch.from_folder(data_folder, train=".", valid_pct=0.2,
ds_tfms=get_transforms(), size=224, num_workers=4).normalize(imagenet_stats)
# + id="dNZg1Zw2vY1H" colab_type="code" colab={}
# If you already cleaned your data, run this cell instead of the one before
# np.random.seed(42)
# data = ImageDataBunch.from_csv(path, folder=".", valid_pct=0.2, csv_labels='cleaned.csv',
# ds_tfms=get_transforms(), size=224, num_workers=4).normalize(imagenet_stats)
# + [markdown] id="qwv2kjeOvY1P" colab_type="text"
# Good! Let's take a look at some of our pictures then.
# + id="JKZ34KhsvY1Q" colab_type="code" colab={}
data.classes
# + id="cB00u7bOvY1X" colab_type="code" colab={}
data.show_batch(rows=3, figsize=(7,8))
# + id="lyyQ-WIevY1b" colab_type="code" colab={}
data.classes, data.c, len(data.train_ds), len(data.valid_ds)
# + [markdown] id="lWuetI62vY1f" colab_type="text"
# ## Train model
# + id="NEUaOxl0vY1g" colab_type="code" colab={}
learn = cnn_learner(data, models.resnet34, metrics=error_rate)
# + id="0pYu1-ZvvY1k" colab_type="code" colab={}
learn.fit_one_cycle(4)
# + id="46upSo-TvY1m" colab_type="code" colab={}
learn.save('stage-1')
# + id="ScR82vrEvY1o" colab_type="code" colab={}
learn.unfreeze()
# + id="FKaikiP3vY1s" colab_type="code" colab={}
learn.lr_find()
# + id="orPb4yoCvY1u" colab_type="code" colab={}
learn.recorder.plot()
# + id="m3D4bfUfvY1x" colab_type="code" colab={}
learn.fit_one_cycle(2, max_lr=slice(2e-5,2e-3))
# + id="JLOv2zyevY10" colab_type="code" colab={}
learn.save('stage-2')
# + [markdown] id="VaTtm024vY12" colab_type="text"
# ## Interpretation
# + id="uiEM9ueGvY14" colab_type="code" colab={}
learn.load('stage-2');
# + id="6kKIBm1hvY16" colab_type="code" colab={}
interp = ClassificationInterpretation.from_learner(learn)
# + id="02kmrYhavY19" colab_type="code" colab={}
interp.plot_confusion_matrix()
# + [markdown] id="TLslUx4DvY1_" colab_type="text"
# ## Cleaning Up - Not Working In Colab
#
# Some of our top losses aren't due to bad performance by our model. There are images in our data set that shouldn't be.
#
# Using the `ImageCleaner` widget from `fastai.widgets` we can prune our top losses, removing photos that don't belong.
# + id="OvDobr6xvY2A" colab_type="code" colab={}
from fastai.widgets import *
# + [markdown] id="9hlhzK9nvY2C" colab_type="text"
# First we need to get the file paths from our top_losses. We can do this with `.from_toplosses`. We then feed the top losses indexes and corresponding dataset to `ImageCleaner`.
#
# Notice that the widget will not delete images directly from disk but it will create a new csv file `cleaned.csv` from where you can create a new ImageDataBunch with the corrected labels to continue training your model.
# + [markdown] id="Dqt-NkcSvY2C" colab_type="text"
# In order to clean the entire set of images, we need to create a new dataset without the split. The video lecture demostrated the use of the `ds_type` param which no longer has any effect. See [the thread](https://forums.fast.ai/t/duplicate-widget/30975/10) for more details.
# + id="3EdNAuf1vY2D" colab_type="code" colab={}
db = (ImageList.from_folder(data_folder)
.no_split()
.label_from_folder()
.transform(get_transforms(), size=224)
.databunch()
)
# + id="cAZ1UoktvY2F" colab_type="code" colab={}
# If you already cleaned your data using indexes from `from_toplosses`,
# run this cell instead of the one before to proceed with removing duplicates.
# Otherwise all the results of the previous step would be overwritten by
# the new run of `ImageCleaner`.
# db = (ImageList.from_csv(path, 'cleaned.csv', folder='.')
# .no_split()
# .label_from_df()
# .transform(get_transforms(), size=224)
# .databunch()
# )
# + [markdown] id="iP8b84u7vY2G" colab_type="text"
# Then we create a new learner to use our new databunch with all the images.
# + id="Kt-KX7nLvY2H" colab_type="code" colab={}
learn_cln = cnn_learner(db, models.resnet34, metrics=error_rate)
learn_cln.load('stage-2');
# + id="5gkWiP7KvY2K" colab_type="code" colab={}
ds, idxs = DatasetFormatter().from_toplosses(learn_cln)
# + [markdown] id="1nVDvxcIvY2R" colab_type="text"
# Make sure you're running this notebook in Jupyter Notebook, not Jupyter Lab. That is accessible via [/tree](/tree), not [/lab](/lab). Running the `ImageCleaner` widget in Jupyter Lab or Colab is [not currently supported](https://github.com/fastai/fastai/issues/1539).
# + id="OYxel4OgvY2S" colab_type="code" colab={}
# do not run me in colab
ImageCleaner(ds, idxs, data_folder)
# + [markdown] id="SMDgeLcUvY2V" colab_type="text"
# Flag photos for deletion by clicking 'Delete'. Then click 'Next Batch' to delete flagged photos and keep the rest in that row. `ImageCleaner` will show you a new row of images until there are no more to show. In this case, the widget will show you images until there are none left from `top_losses.ImageCleaner(ds, idxs)`
# + [markdown] id="XcSAOtSjvY2W" colab_type="text"
# You can also find duplicates in your dataset and delete them! To do this, you need to run `.from_similars` to get the potential duplicates' ids and then run `ImageCleaner` with `duplicates=True`. The API works in a similar way as with misclassified images: just choose the ones you want to delete and click 'Next Batch' until there are no more images left.
# + [markdown] id="5Eb3L5l_vY2Y" colab_type="text"
# Make sure to recreate the databunch and `learn_cln` from the `cleaned.csv` file. Otherwise the file would be overwritten from scratch, loosing all the results from cleaning the data from toplosses.
# + id="KKslmXZ-vY2Z" colab_type="code" colab={}
ds, idxs = DatasetFormatter().from_similars(learn_cln)
# + id="UYNVrJaKvY2c" colab_type="code" colab={}
# do not run me in Colab
ImageCleaner(ds, idxs, path, duplicates=True)
# + [markdown] id="a1XnBEVlvY2g" colab_type="text"
# Remember to recreate your ImageDataBunch from your `cleaned.csv` to include the changes you made in your data!
# + [markdown] id="C-MN0Y9uKUTq" colab_type="text"
# ## Top losses
# + id="OfWcQIB2KsCu" colab_type="code" colab={}
interp.plot_top_losses(9, figsize=(15,11), heatmap=False)
# + [markdown] id="LPMtyN0AvY2y" colab_type="text"
# ## Things that can go wrong
# + [markdown] id="mkVcum6hvY2z" colab_type="text"
# - Most of the time things will train fine with the defaults
# - There's not much you really need to tune (despite what you've heard!)
# - Most likely are
# - Learning rate
# - Number of epochs
# + [markdown] id="mlOY2ZWLvY20" colab_type="text"
# ### Learning rate (LR) too high
# + id="lcOAZXNHvY20" colab_type="code" colab={}
learn = cnn_learner(data, models.resnet34, metrics=error_rate)
# + id="cOljVaxIvY22" colab_type="code" colab={}
learn.fit_one_cycle(1, max_lr=0.5)
# + [markdown] id="jT7PQho_vY26" colab_type="text"
# ### Learning rate (LR) too low
# + id="xHrvSyN6vY26" colab_type="code" colab={}
learn = cnn_learner(data, models.resnet34, metrics=error_rate)
# + [markdown] id="eHd7JWdEvY28" colab_type="text"
# Previously we had this result:
#
# ```
# Total time: 00:57
# epoch train_loss valid_loss error_rate
# 1 1.030236 0.179226 0.028369 (00:14)
# 2 0.561508 0.055464 0.014184 (00:13)
# 3 0.396103 0.053801 0.014184 (00:13)
# 4 0.316883 0.050197 0.021277 (00:15)
# ```
# + id="RflXvYssvY29" colab_type="code" colab={}
learn.fit_one_cycle(5, max_lr=1e-5)
# + id="TH7rUETAvY3B" colab_type="code" colab={}
learn.recorder.plot_losses()
# + [markdown] id="Cs8QMB3evY3D" colab_type="text"
# As well as taking a really long time, it's getting too many looks at each image, so may overfit.
# + [markdown] id="KeU75ZlXvY3E" colab_type="text"
# ### Too few epochs
# + id="RjodtYZ4vY3E" colab_type="code" colab={}
learn = cnn_learner(data, models.resnet34, metrics=error_rate, pretrained=False)
# + id="BadMQ4apvY3H" colab_type="code" colab={}
learn.fit_one_cycle(1)
# + [markdown] id="HnpE2KxwvY3J" colab_type="text"
# ### Too many epochs
# + id="80-SyRkXvY3J" colab_type="code" colab={}
np.random.seed(42)
data = ImageDataBunch.from_folder(path, train=".", valid_pct=0.9, bs=32,
ds_tfms=get_transforms(do_flip=False, max_rotate=0, max_zoom=1, max_lighting=0, max_warp=0
),size=224, num_workers=4).normalize(imagenet_stats)
# + id="xYWGNh4wvY3L" colab_type="code" colab={}
learn = cnn_learner(data, models.resnet50, metrics=error_rate, ps=0, wd=0)
learn.unfreeze()
# + id="AKoGYm8HvY3M" colab_type="code" colab={}
learn.fit_one_cycle(40, slice(1e-6,1e-4))
# + [markdown] id="QX05wqS4vY2h" colab_type="text"
# ## Putting your model in production
# + [markdown] id="1Mff_XdivY2i" colab_type="text"
# First thing first, let's export the content of our `Learner` object for production:
# + id="umcscfR9vY2i" colab_type="code" colab={}
learn.export()
# + [markdown] id="LHy9FgvEvY2k" colab_type="text"
# This will create a file named 'export.pkl' in the directory where we were working that contains everything we need to deploy our model (the model, the weights but also some metadata like the classes or the transforms/normalization used).
# + [markdown] id="pPfSmeikvY2l" colab_type="text"
# You probably want to use CPU for inference, except at massive scale (and you almost certainly don't need to train in real-time). If you don't have a GPU that happens automatically. You can test your model on CPU like so:
# + id="bzJ6EqHNvY2l" colab_type="code" colab={}
defaults.device = torch.device('cpu')
# + id="TK24p-eqvY2n" colab_type="code" colab={}
img = open_image(path/'black'/'00000021.jpg')
img
# + [markdown] id="AJkUk1ISvY2r" colab_type="text"
# We create our `Learner` in production enviromnent like this, just make sure that `path` contains the file 'export.pkl' from before.
# + id="a7b4d5qmvY2t" colab_type="code" colab={}
learn = load_learner(path)
# + id="K6fvc745vY2v" colab_type="code" colab={}
pred_class,pred_idx,outputs = learn.predict(img)
pred_class
# + [markdown] id="dbglLc0QvY2x" colab_type="text"
# For more information about putting your model into production, see [fast.ai webpage](https://course.fast.ai/deployment_render.html).
| colab3_dataset_from_Google_Images.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
os.environ['GOOGLE_APPLICATION_CREDENTIALS'] = '/home/husein/t5/prepare/mesolitica-tpu.json'
# +
from google.cloud import storage
client = storage.Client()
bucket = client.bucket('mesolitica-tpu-general')
directory = 't2t-paraphrase-base'
model = '50000'
# !mkdir {directory}
blob = bucket.blob(f'{directory}/model.ckpt-{model}.data-00000-of-00001')
blob.download_to_filename(f'{directory}/model.ckpt-{model}.data-00000-of-00001')
blob = bucket.blob(f'{directory}/model.ckpt-{model}.index')
blob.download_to_filename(f'{directory}/model.ckpt-{model}.index')
blob = bucket.blob(f'{directory}/model.ckpt-{model}.meta')
blob.download_to_filename(f'{directory}/model.ckpt-{model}.meta')
# +
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '3'
from tensor2tensor.data_generators import problem
from tensor2tensor.data_generators import text_problems
from tensor2tensor.data_generators import translate
from tensor2tensor.utils import registry
from tensor2tensor import problems
import tensorflow as tf
import os
import logging
logger = logging.getLogger()
tf.logging.set_verbosity(tf.logging.DEBUG)
# +
import sentencepiece as spm
vocab = 'sp10m.cased.t5.model'
sp = spm.SentencePieceProcessor()
sp.Load(vocab)
class Encoder:
def __init__(self, sp):
self.sp = sp
self.vocab_size = sp.GetPieceSize() + 100
def encode(self, s):
return self.sp.EncodeAsIds(s)
def decode(self, ids, strip_extraneous=False):
return self.sp.DecodeIds(list(ids))
encoder = Encoder(sp)
# +
from tqdm import tqdm
from glob import glob
@registry.register_problem
class Seq2Seq(text_problems.Text2TextProblem):
@property
def approx_vocab_size(self):
return 32100
@property
def is_generate_per_split(self):
return False
def feature_encoders(self, data_dir):
encoder = Encoder(sp)
return {
"inputs": encoder,
"targets": encoder
}
# -
DATA_DIR = os.path.expanduser('t2t-paraphrase/data')
TMP_DIR = os.path.expanduser('t2t-paraphrase/tmp')
PROBLEM = 'seq2_seq'
t2t_problem = problems.problem(PROBLEM)
# +
import tensorflow as tf
import os
ckpt_path = tf.train.latest_checkpoint('t2t-paraphrase-base')
ckpt_path
# -
from tensor2tensor import models
from tensor2tensor import problems
from tensor2tensor.layers import common_layers
from tensor2tensor.utils import trainer_lib
from tensor2tensor.utils import t2t_model
from tensor2tensor.utils import registry
from tensor2tensor.utils import metrics
from tensor2tensor.data_generators import problem
from tensor2tensor.data_generators import text_problems
from tensor2tensor.data_generators import translate
from tensor2tensor.utils import registry
# +
from tensor2tensor.layers import modalities
from tensor2tensor.layers import common_layers
def top_p_logits(logits, p):
with tf.variable_scope('top_p_logits'):
logits_sort = tf.sort(logits, direction = 'DESCENDING')
probs_sort = tf.nn.softmax(logits_sort)
probs_sums = tf.cumsum(probs_sort, axis = 1, exclusive = True)
logits_masked = tf.where(
probs_sums < p, logits_sort, tf.ones_like(logits_sort) * 1000
) # [batchsize, vocab]
min_logits = tf.reduce_min(
logits_masked, axis = 1, keepdims = True
) # [batchsize, 1]
return tf.where(
logits < min_logits,
tf.ones_like(logits, dtype = logits.dtype) * -1e10,
logits,
)
def sample(translate_model, features):
logits, losses = translate_model(features)
logits_shape = common_layers.shape_list(logits)
logits_p = logits[:,0,:,0,:] / translate_model.hparams.sampling_temp
logits_p = top_p_logits(logits_p, translate_model.hparams.top_p)
reshaped_logits = tf.reshape(logits_p, [-1, logits_shape[-1]])
choices = tf.multinomial(reshaped_logits, 1)
samples = tf.reshape(choices, logits_shape[:-1])
return samples, logits, losses
def nucleus_sampling(translate_model, features, decode_length):
"""A slow greedy inference method.
Quadratic time in decode_length.
Args:
features: an map of string to `Tensor`
decode_length: an integer. How many additional timesteps to decode.
Returns:
A dict of decoding results {
"outputs": integer `Tensor` of decoded ids of shape
[batch_size, <= decode_length] if beam_size == 1 or
[batch_size, top_beams, <= decode_length]
"scores": None
"logits": `Tensor` of shape [batch_size, time, 1, 1, vocab_size].
"losses": a dictionary: {loss-name (string): floating point `Scalar`}
}
"""
if not features:
features = {}
inputs_old = None
if 'inputs' in features and len(features['inputs'].shape) < 4:
inputs_old = features['inputs']
features['inputs'] = tf.expand_dims(features['inputs'], 2)
# Save the targets in a var and reassign it after the tf.while loop to avoid
# having targets being in a 'while' frame. This ensures targets when used
# in metric functions stays in the same frame as other vars.
targets_old = features.get('targets', None)
target_modality = translate_model._problem_hparams.modality['targets']
def infer_step(recent_output, recent_logits, unused_loss):
"""Inference step."""
if not tf.executing_eagerly():
if translate_model._target_modality_is_real:
dim = translate_model._problem_hparams.vocab_size['targets']
if dim is not None and hasattr(
translate_model._hparams, 'vocab_divisor'
):
dim += (-dim) % translate_model._hparams.vocab_divisor
recent_output.set_shape([None, None, None, dim])
else:
recent_output.set_shape([None, None, None, 1])
padded = tf.pad(recent_output, [[0, 0], [0, 1], [0, 0], [0, 0]])
features['targets'] = padded
# This is inefficient in that it generates samples at all timesteps,
# not just the last one, except if target_modality is pointwise.
samples, logits, losses = sample(translate_model, features)
# Concatenate the already-generated recent_output with last timestep
# of the newly-generated samples.
top = translate_model._hparams.top.get(
'targets', modalities.get_top(target_modality)
)
if getattr(top, 'pointwise', False):
cur_sample = samples[:, -1, :, :]
else:
cur_sample = samples[
:, common_layers.shape_list(recent_output)[1], :, :
]
if translate_model._target_modality_is_real:
cur_sample = tf.expand_dims(cur_sample, axis = 1)
samples = tf.concat([recent_output, cur_sample], axis = 1)
else:
cur_sample = tf.to_int64(tf.expand_dims(cur_sample, axis = 1))
samples = tf.concat([recent_output, cur_sample], axis = 1)
if not tf.executing_eagerly():
samples.set_shape([None, None, None, 1])
# Assuming we have one shard for logits.
logits = tf.concat([recent_logits, logits[:, -1:]], 1)
loss = sum([l for l in losses.values() if l is not None])
return samples, logits, loss
# Create an initial output tensor. This will be passed
# to the infer_step, which adds one timestep at every iteration.
if 'partial_targets' in features:
initial_output = tf.to_int64(features['partial_targets'])
while len(initial_output.get_shape().as_list()) < 4:
initial_output = tf.expand_dims(initial_output, 2)
batch_size = common_layers.shape_list(initial_output)[0]
else:
batch_size = common_layers.shape_list(features['inputs'])[0]
if translate_model._target_modality_is_real:
dim = translate_model._problem_hparams.vocab_size['targets']
if dim is not None and hasattr(
translate_model._hparams, 'vocab_divisor'
):
dim += (-dim) % translate_model._hparams.vocab_divisor
initial_output = tf.zeros(
(batch_size, 0, 1, dim), dtype = tf.float32
)
else:
initial_output = tf.zeros((batch_size, 0, 1, 1), dtype = tf.int64)
# Hack: foldl complains when the output shape is less specified than the
# input shape, so we confuse it about the input shape.
initial_output = tf.slice(
initial_output, [0, 0, 0, 0], common_layers.shape_list(initial_output)
)
target_modality = translate_model._problem_hparams.modality['targets']
if (
target_modality == modalities.ModalityType.CLASS_LABEL
or translate_model._problem_hparams.get('regression_targets')
):
decode_length = 1
else:
if 'partial_targets' in features:
prefix_length = common_layers.shape_list(
features['partial_targets']
)[1]
else:
prefix_length = common_layers.shape_list(features['inputs'])[1]
decode_length = prefix_length + decode_length
# Initial values of result, logits and loss.
result = initial_output
vocab_size = translate_model._problem_hparams.vocab_size['targets']
if vocab_size is not None and hasattr(
translate_model._hparams, 'vocab_divisor'
):
vocab_size += (-vocab_size) % translate_model._hparams.vocab_divisor
if translate_model._target_modality_is_real:
logits = tf.zeros((batch_size, 0, 1, vocab_size))
logits_shape_inv = [None, None, None, None]
else:
# tensor of shape [batch_size, time, 1, 1, vocab_size]
logits = tf.zeros((batch_size, 0, 1, 1, vocab_size))
logits_shape_inv = [None, None, None, None, None]
if not tf.executing_eagerly():
logits.set_shape(logits_shape_inv)
loss = 0.0
def while_exit_cond(
result, logits, loss
): # pylint: disable=unused-argument
"""Exit the loop either if reach decode_length or EOS."""
length = common_layers.shape_list(result)[1]
not_overflow = length < decode_length
if translate_model._problem_hparams.stop_at_eos:
def fn_not_eos():
return tf.not_equal( # Check if the last predicted element is a EOS
tf.squeeze(result[:, -1, :, :]), 1
)
not_eos = tf.cond(
# We only check for early stopping if there is at least 1 element (
# otherwise not_eos will crash).
tf.not_equal(length, 0),
fn_not_eos,
lambda: True,
)
return tf.cond(
tf.equal(batch_size, 1),
# If batch_size == 1, we check EOS for early stopping.
lambda: tf.logical_and(not_overflow, not_eos),
# Else, just wait for max length
lambda: not_overflow,
)
return not_overflow
result, logits, loss = tf.while_loop(
while_exit_cond,
infer_step,
[result, logits, loss],
shape_invariants = [
tf.TensorShape([None, None, None, None]),
tf.TensorShape(logits_shape_inv),
tf.TensorShape([]),
],
back_prop = False,
parallel_iterations = 1,
)
if inputs_old is not None: # Restore to not confuse Estimator.
features['inputs'] = inputs_old
# Reassign targets back to the previous value.
if targets_old is not None:
features['targets'] = targets_old
losses = {'training': loss}
if 'partial_targets' in features:
partial_target_length = common_layers.shape_list(
features['partial_targets']
)[1]
result = tf.slice(
result, [0, partial_target_length, 0, 0], [-1, -1, -1, -1]
)
return {
'outputs': result,
'scores': None,
'logits': logits,
'losses': losses,
}
# +
class Model:
def __init__(self, HPARAMS = "transformer_base", DATA_DIR = 't2t/data'):
self.X = tf.placeholder(tf.int32, [None, None])
self.Y = tf.placeholder(tf.int32, [None, None])
self.top_p = tf.placeholder(tf.float32, None)
self.X_seq_len = tf.count_nonzero(self.X, 1, dtype=tf.int32)
self.maxlen_decode = tf.reduce_max(self.X_seq_len)
#self.maxlen_decode = tf.placeholder(tf.int32, None)
x = tf.expand_dims(tf.expand_dims(self.X, -1), -1)
y = tf.expand_dims(tf.expand_dims(self.Y, -1), -1)
features = {
"inputs": x,
"targets": y,
"target_space_id": tf.constant(1, dtype=tf.int32),
}
self.features = features
Modes = tf.estimator.ModeKeys
hparams = trainer_lib.create_hparams(HPARAMS, data_dir=DATA_DIR, problem_name=PROBLEM)
hparams.filter_size = 3072
hparams.hidden_size = 768
hparams.num_heads = 12
hparams.num_hidden_layers = 8
hparams.vocab_divisor = 128
hparams.dropout = 0.1
hparams.label_smoothing = 0.0
hparams.shared_embedding_and_softmax_weights = False
hparams.eval_drop_long_sequences = True
hparams.max_length = 1024
hparams.multiproblem_mixing_schedule = 'pretrain'
hparams.symbol_modality_num_shards = 1
hparams.attention_dropout_broadcast_dims = '0,1'
hparams.relu_dropout_broadcast_dims = '1'
hparams.layer_prepostprocess_dropout_broadcast_dims = '1'
translate_model = registry.model('transformer')(hparams, Modes.PREDICT)
self.translate_model = translate_model
logits, _ = translate_model(features)
self.logits = logits
translate_model.hparams.top_p = self.top_p
with tf.variable_scope(tf.get_variable_scope(), reuse=True):
self.fast_result = translate_model._greedy_infer(features, self.maxlen_decode)["outputs"]
self.beam_result = translate_model._beam_decode_slow(
features, self.maxlen_decode, beam_size=3,
top_beams=1, alpha=0.5)["outputs"]
self.nucleus_result = nucleus_sampling(translate_model, features, self.maxlen_decode)["outputs"]
self.nucleus_result = self.nucleus_result[:,:,0,0]
self.fast_result = tf.identity(self.fast_result, name = 'greedy')
self.beam_result = tf.identity(self.beam_result, name = 'beam')
self.nucleus_result = tf.identity(self.nucleus_result, name = 'nucleus')
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Model()
# -
var_lists = tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES)
saver = tf.train.Saver(var_list = var_lists)
saver.restore(sess, 't2t-paraphrase-base/model.ckpt-50000')
# +
import re
from unidecode import unidecode
def cleaning(string):
return re.sub(r'[ ]+', ' ', unidecode(string.replace('\n', ' '))).strip()
# +
import re
from unidecode import unidecode
from malaya.text.rules import normalized_chars
def filter_news(string):
string = string.lower()
return 'javascript is disabled' in string or 'requires javascript' in string or 'javascript' in string \
or 'président' in string
def make_cleaning(s, c_dict):
s = s.translate(c_dict)
return s
def transformer_textcleaning(string):
"""
use by any transformer model before tokenization
"""
string = unidecode(string)
string = ' '.join(
[make_cleaning(w, normalized_chars) for w in string.split()]
)
string = re.sub('\(dot\)', '.', string)
string = (
re.sub(re.findall(r'\<a(.*?)\>', string)[0], '', string)
if (len(re.findall(r'\<a (.*?)\>', string)) > 0)
and ('href' in re.findall(r'\<a (.*?)\>', string)[0])
else string
)
string = re.sub(
r'\w+:\/{2}[\d\w-]+(\.[\d\w-]+)*(?:(?:\/[^\s/]*))*', ' ', string
)
string = string.replace('\n', ' ')
string = re.sub(r'[ ]+', ' ', string).strip().split()
string = [w for w in string if w[0] != '@']
return ' '.join(string)
# +
import json
# # !wget https://f000.backblazeb2.com/file/malay-dataset/testset/paraphrase-set.json
with open('../summary/paraphrase-set.json') as fopen:
test = json.load(fopen)
test.keys()
# -
pad_sequences = tf.keras.preprocessing.sequence.pad_sequences
# +
from tqdm import tqdm
batch_size = 20
results = []
for i in tqdm(range(0, len(test['test_before']), batch_size)):
batch_x = test['test_before'][i: i + batch_size]
batches = []
for b in batch_x:
encoded = encoder.encode(f'parafrasa: {cleaning(b)}')
encoded = encoded[:1023] + [1]
batches.append(encoded)
batches = pad_sequences(batches, padding='post')
g = sess.run(model.fast_result, feed_dict = {model.X:batches})
for b in g:
results.append(encoder.decode(b.tolist()))
# +
from tensor2tensor.utils import bleu_hook
bleu_hook.compute_bleu(reference_corpus = test['test_after'],
translation_corpus = results)
# +
encoded = encoder.encode(f"parafrasa: {cleaning(test['test_before'][0])}") + [1]
f, b, n = sess.run([model.fast_result, model.beam_result, model.nucleus_result],
feed_dict = {model.X: [encoded], model.top_p: 0.7})
(encoder.decode(f[0].tolist()),
encoder.decode(b[0].tolist()),
encoder.decode(n[0].tolist()))
# -
test['test_after'][0]
# +
encoded = encoder.encode(f"parafrasa: {cleaning(test['test_before'][1])}") + [1]
f, b, n = sess.run([model.fast_result, model.beam_result, model.nucleus_result],
feed_dict = {model.X: [encoded], model.top_p: 0.7})
(encoder.decode(f[0].tolist()),
encoder.decode(b[0].tolist()),
encoder.decode(n[0].tolist()))
# -
test['test_after'][1]
# +
encoded = encoder.encode(f"parafrasa: {cleaning(test['test_before'][2])}") + [1]
f, b, n = sess.run([model.fast_result, model.beam_result, model.nucleus_result],
feed_dict = {model.X: [encoded], model.top_p: 0.7})
(encoder.decode(f[0].tolist()),
encoder.decode(b[0].tolist()),
encoder.decode(n[0].tolist()))
# -
test['test_after'][2]
saver = tf.train.Saver(tf.trainable_variables())
saver.save(sess, 'transformer-base/model.ckpt')
strings = ','.join(
[
n.name
for n in tf.get_default_graph().as_graph_def().node
if ('Variable' in n.op
or 'Placeholder' in n.name
or 'greedy' in n.name
or 'beam' in n.name
or 'nucleus' in n.name
or 'alphas' in n.name
or 'self/Softmax' in n.name)
and 'adam' not in n.name
and 'beta' not in n.name
and 'global_step' not in n.name
and 'modality' not in n.name
and 'Assign' not in n.name
]
)
strings.split(',')
def freeze_graph(model_dir, output_node_names):
if not tf.gfile.Exists(model_dir):
raise AssertionError(
"Export directory doesn't exists. Please specify an export "
'directory: %s' % model_dir
)
checkpoint = tf.train.get_checkpoint_state(model_dir)
input_checkpoint = checkpoint.model_checkpoint_path
absolute_model_dir = '/'.join(input_checkpoint.split('/')[:-1])
output_graph = absolute_model_dir + '/frozen_model.pb'
clear_devices = True
with tf.Session(graph = tf.Graph()) as sess:
saver = tf.train.import_meta_graph(
input_checkpoint + '.meta', clear_devices = clear_devices
)
saver.restore(sess, input_checkpoint)
output_graph_def = tf.graph_util.convert_variables_to_constants(
sess,
tf.get_default_graph().as_graph_def(),
output_node_names.split(','),
)
with tf.gfile.GFile(output_graph, 'wb') as f:
f.write(output_graph_def.SerializeToString())
print('%d ops in the final graph.' % len(output_graph_def.node))
freeze_graph('transformer-base', strings)
def load_graph(frozen_graph_filename):
with tf.gfile.GFile(frozen_graph_filename, 'rb') as f:
graph_def = tf.GraphDef()
graph_def.ParseFromString(f.read())
with tf.Graph().as_default() as graph:
tf.import_graph_def(graph_def)
return graph
g = load_graph('transformer-base/frozen_model.pb')
x = g.get_tensor_by_name('import/Placeholder:0')
greedy = g.get_tensor_by_name('import/greedy:0')
beam = g.get_tensor_by_name('import/beam:0')
nucleus = g.get_tensor_by_name('import/nucleus:0')
test_sess = tf.InteractiveSession(graph = g)
top_p = g.get_tensor_by_name('import/Placeholder_2:0')
g, b, n = test_sess.run([greedy, beam, nucleus], feed_dict = {x:[encoded],
top_p: 0.7})
(encoder.decode(f[0].tolist()),
encoder.decode(b[0].tolist()),
encoder.decode(n[0].tolist()))
| session/paraphrase/t2t/t2t-base-paraphrase.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analyzing, and Visualizing Data
# +
import matplotlib
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# -
matplotlib.style.use('ggplot')
df = pd.read_csv('twitter_archive_master.csv')
df.info()
df.head()
# +
# Convert columns to their appropriate types and set the timestamp as an index
df['tweet_id'] = df['tweet_id'].astype(object)
df['timestamp'] = pd.to_datetime(df.timestamp)
df['source'] = df['source'].astype('category')
df['dog_stage'] = df['dog_stage'].astype('category')
df['dog_gender'] = df['dog_gender'].astype('category')
df.set_index('timestamp', inplace=True)
df.info()
# -
df.describe()
# **Plot the correlation map to see the relationship between variables**
#
f,ax = plt.subplots(figsize=(18, 18))
sns.heatmap(df[['source', 'favorites','retweets',
'user_followers','rating_numerator']].corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax)
plt.title('Correlation Map')
# - The only strong correlation we see here is between favorites and retweet, this is normal (more favorites mean more retweets)
# - User followers and retweet have a weak negative correlation of -0.4 (this seems the opposite of normal prediction)
# - Rating don't get affected with any other variable from the ones we ploted
# #### Tweet Retweet Vs Favorites.
df.plot(kind='scatter',x='favorites',y='retweets', alpha = 0.5)
plt.xlabel('Favorites')
plt.ylabel('Retweets')
plt.title('Retweets and favorites Scatter plot')
# - As the correlation map shows that retweet is high correlated with favorites.
# ### Rating System
# Our range will be [0,16] taking of the two ouliers (1776 and 420)
df.plot(y ='rating_numerator', ylim=[0,16], style = '.', alpha = .2)
plt.title('Rating plot over Time')
plt.xlabel('Date')
plt.ylabel('Rating')
# - The page start with small rating than they adopt the system of rating numerator more than the denominator
#
df['retweets'].plot(color = 'red', label='Retweets')
df['favorites'].plot(color = 'blue', label='Favorites')
plt.style.use('seaborn-darkgrid')
plt.legend(loc='upper left')
plt.xlabel('Tweet timestamp')
plt.ylabel('Count')
plt.title('Retweets and favorites over time')
plt.savefig('retweets_favorites.png')
plt.show()
# - Brent has all the right to get mad (ratings getting higher with no specific reason)
#
sns.factorplot(kind='box',
y='rating_numerator',
x='dog_stage',
hue='dog_gender',
data=df[df['dog_stage'] != 'None'],
size=8,
aspect=1.5,
legend_out=False)
# - According to our treatment (getting the gender from the text of the tweet) we have male dogs more than female dogs in our dataset, whatever the female rating mean more than the male rating mean
# +
# Plot the data partitioned by dog stage
dog_stage_count = list(df[df['dog_stage'] != 'None']['dog_stage'].value_counts())[0:4]
dog_stages = df[df['dog_stage'] != 'None']['dog_stage'].value_counts().index.tolist()[0:4]
explode = (0.2, 0.1, 0.1, 0.1)
fig1, ax1 = plt.subplots()
ax1.pie(dog_stage_count, explode = explode, labels = dog_stages, shadow = True, startangle = 90)
ax1.axis('equal')
# -
df[df['dog_stage'] != 'None'].groupby('dog_stage')['rating_numerator'].mean()
# - Pupper is the most owned dog, but it has the lowest mean rating
# ### Conclusion
# The Twitter account `WeRateDogs` (@dog_rates) is devoted to humorously reviewing pictures of dogs doing adorable poses. Dogs are rated on a scale of one to ten, but are invariably given ratings in excess of the maximum.
#
# If you are thinking of adopting a dog, get a floof(er).
| act_report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### What is Machine Learning?
# Machine Learning is about building computational artifacts that learn over time based on experience. You have data and you try and infer information from that data using computationally applied statistical algorithms. It is a field of study that gives computers the ability to learn without being explicitly programmed.
#
# Here's a more formal definition:
#
# _A computer program is said to learn from experience E with respect to some task T and some performance measure P, if its performance on T, as measured by P, improves with experience E._
#
# Let's break this down in terms of an example. Let's say you want to create a spam mail classifier. How will that problem look like in terms of the above definition?
#
# - Task: Classifying mail as spam or not spam.
# - Experience: Watching us interact with the mailbox by throwing some mails in the junk folder and keeping the rest.
# - Performance: How many mails does our classifier correctly mark as spam/not spam.
#
# There isn't a simple algorithmic solution to this problem. You might think that most spam mails follow a certain kind of pattern or have certain tells which you can hardcode into an algorithm but that really isnt true anymore. Many spam mails are quite sophisticated now and dont really follow any set pattern. In fact, one person's spam could be another person's steak! So really the pattern we'll be looking for here is quite arbitary and is actually dependent on the person who is using the mailbox. This is wear statistical learning comes in.
# ### Supervised Learning
# Supervised learning can be thought of as **function approximation**. You have a bunch of data and labels corresponding to that data and you want to computationally approximate the most accurate relationship between the data and its labels anda then generalize the relationship so that you can accurately predict the labels of new datasets too.
#
# _Example:_
#
# <table>
# <tr>
# <td><b>Data (Input)</b></td>
# <td><b>Label (Output)</b></td>
# </tr>
# <tr>
# <td>1</td><td>1</td>
# </tr>
# <tr><td>2</td><td>4</td></tr>
# <tr><td>3</td><td>9</td></tr>
# <tr><td>4</td><td>16</td></tr>
# <tr><td>5</td><td>25</td></tr>
# <tr><td>6</td><td>36</td></tr>
# </table>
#
# Can we _approximate_ what the function behind this dataset is? Looks like $x^2$, doesnt it? Based on that can we predict what the output for the following input is:
#
# $$10 - ?$$
#
# Since, the function is $x^2$, the output will be:
#
# $$10 - 100$$
#
# However, this is still a leap of faith. We _assume_ the function is well behaved and is $x^2$ but what if the function is actually something like this:
#
# $$ f(x) = \begin{cases}
# if x < 10: x^2\\
# else: x
# \end{cases}
# $$
#
# In this case, our prediction for input $10$ is incorrect. Therefore, there are a few assumptions with which we approach supervised learning, one of them being that the function behind the data is a well behaved function. A well behaved function is continous.
# ### Unsupervised Learning
# Unlike supervised learning, in unsupervised learning we just have the inputs, without any outputs. We dont have the labels for our data and we have to derive some structure for the data using just the inputs.
#
# _Example_:
# You might be given a bunch of fruits without information about what each fruit is called (let's assume you dont know anything about fruits). And you can still start classifying them in groups based on shape or color or smell. Let's say you group them up by color, you put all orange color fruits in one group, all red colored fruits in one, and all yellow colored fruits in one. You have derived a structure between the fruits (based on color) by using just the data.
#
# So if supervised learning is about functional approximation, unsupervised learning is about **description**.
# ### Reinforcement Learning
# Learning from delayed rewards. Supervised learning is about playing a game _after_ learning about all the rules. So if you're playing tic tac toe, you know exactly where to move and what move is a good move and what move is a bad move. Reinforcement learning is playing a game _without_ knowing any of the rules, and discovering the best way to play the game after you have finished playing it. Again, if you're playing tic tac toe without knowing what it is and you keep playing multiple games of tic tac toe, you'll start discovering the various rules and moves of tic tac toe on your own and after a significant amount of play time, you'll become very good at tic tac toe.
# All these forms of learning can be thought of as some form of **optimizations**.
# - In supervised learning you're trying to optimize a funtion that labels data well.
# - In unsupervised learning you're trying to find the most optimal way to group together data.
# - In reinforcement learning you're trying to optimize a certain behavior.
#
# In machine learning, **data** is central. It differs slightly from a pure AI perspective where **algorithms** become central.
# ### How does a Machine Learning Project Look Like
# 1. **Define and Assess the Problem**: The first aim in any project, almost unanimously across industry, is to know what you're doing. The same applies to machine learning/data science/statistical analysis projects. Formulate the question/problem/value you are trying to answer/solve/add. Assess the problem. _Problems before requirements_, _requirements before solutions_, _solutions before design_, and _design before technology_.
#
#
# 2. **Gather the Data**: After defining the problem you're trying to solve and assessing its validity, we move on to actually looking at the data requirements. If you have correctly defined and assessed the problem, then you should know what kind of data you are looking for. For example, if you've defined your problem as being able to correctly predict the prices of houses in X locality; then maybe you've identified the data which can help you predict the prices as number of rooms in a house, types of rooms in a house, total area, location, nearby ammenities like schools and hospitals, crime in area and so on. These are also known as _features_ or _independent/predictor/input variables_ and there are different ways to select the best features. Of course, you'll also need the past prices of these houses to map the features onto. The past prices of the houses is called the _dependent/output variable_; or the _response variable_. So the next step will be to either find the data online or go and collect it.
#
#
# 3. **Prepare Data for Consumption**: This step is often referred to as _data wrangling_, a required process to turn “wild” data into “manageable” data. Data wrangling includes implementing data architectures for storage and processing, developing data governance standards for quality and control, data extraction (i.e. ETL and web scraping), and data cleaning to identify aberrant, missing, or outlier data points.
#
#
# 4. **Exploratory Analysis**: At this stage the idea is to understand our data better. This involves using statistics to describe our data, the distribution, the variance and standard deviations, it's spread, correlations etc and then infer knowledge from it. Here we also check for multicollinearity, if we can perform any kind of feature engineering—be it creating new features out of the set we have, reducing features; or selecting the best ones for modelling. Anybody who has ever worked with data knows, garbage-in, garbage-out (GIGO).
#
# 5. **Model Data: Like descriptive and inferential statistics, data modeling can either summarize the data or predict future outcomes. Your dataset and expected results, will determine the algorithms available for use. It's important to remember, algorithms are tools and not magical wands or silver bullets. You must still be the master craft (wo)man that knows how-to select the right tool for the job. An analogy would be asking someone to hand you a Philip screwdriver, and they hand you a flathead screwdriver or worst a hammer. At best, it shows a complete lack of understanding. At worst, it makes completing the project impossible. The same is true in data modelling. The wrong model can lead to poor performance at best and the wrong conclusion (that’s used as actionable intelligence) at worst.
#
# 6. Validate and Implement Data Model: After you've trained your model based on a subset of your data, it's time to test your model. This helps ensure you haven't overfit your model or made it so specific to the selected subset, that it does not accurately fit another subset from the same dataset. In this step we determine if our model overfit, generalize, or underfit our dataset.
#
# 7. Optimize and Strategize: This is the "bionic man" step, where you iterate back through the process to make it better...stronger...faster than it was before. As a data scientist, your strategy should be to outsource developer operations and application plumbing, so you have more time to focus on recommendations and design. Once you're able to package your ideas, this becomes your “currency exchange" rate.
# ### Regression Vs Classification
# - A person's age, height, or income, the value of a house, or the price of a stock are all examples of continuous variables. Problems which map continuous inputs to outputs are known as regression problems.
#
# - Discrete variables take on values in one of $K$ different classes, or categories. Like a person's gender, the brand of a product purchases, cancer diagnosis, or you can also convert a continuous variable to a discrete one. Problems involving discrete outputs are known as classification problems.
#
| .ipynb_checkpoints/Introduction-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # COVID and Toronto Licensed Child Care
#
# ## Objective
# Visualize the COVID cases in Licensed Child Care Centres, particularly in Toronto:
# - graph of cases over time since reporting began in September 2020
# - map LCCs with reported cases in Toronto since reporting begain in Septebmer 2020
#
# Data sourced from Ontario Data Catalogue for COVID cases in Ontario LCC + City of Toronto addresses/coordinates for Toronto LCCs.
#
# ## Remaining challenges:
# - multiple records of same LCC name in Toronto LCC data
# - mismatch of LCC name between Ontario COVID LCC data and Toronto LCC data; unfortunately there is no other unique identifier/key
# - finding limit of 1000+ locations can be mapped on ArcGIS - need to investigate this further
# - work on adding a "slider" that changes the date reported on the map
#
import pandas as pd
from io import StringIO
import requests
import urllib.request
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
url = 'https://data.ontario.ca/dataset/5bf54477-6147-413f-bab0-312f06fcb388/resource/eee282d3-01e6-43ac-9159-4ba694757aea/download/lccactivecovid.csv'
response = requests.get(url)
s = requests.get(url).text
covid_df = pd.read_csv(StringIO(s))
# +
# all cases in ontario
covid_all_df = covid_df
#covid_all_df.set_index('collected_date')
covid_sum_all = covid_all_df.groupby('collected_date')['total_confirmed_cases'].sum().to_frame(name='sum')
# cases in toronto
covid_tor_df = covid_df.loc[(covid_df['municipality'] == 'Toronto')]
#covid_tor_df.set_index('collected_date')
covid_sum_tor = covid_tor_df.groupby('collected_date')['total_confirmed_cases'].sum().to_frame(name='sum')
# +
# plot stacked bar chart
covid_sum_all.columns = ['Rest of Ontario']
covid_sum_tor.columns = ['Toronto']
plotdata = pd.concat([covid_sum_all,covid_sum_tor],axis=1)
plotdata['Rest of Ontario']=plotdata['Rest of Ontario']-plotdata['Toronto']
font = {'family' : 'normal',
'weight' : 'normal',
'size' : 22}
matplotlib.rc('font', **font)
plotdata.plot(figsize=(30,10),kind='bar',stacked=True)
plt.title("Covid in Ontario Licensed Childcare Centres")
plt.xlabel("Date")
plt.ylabel("Cases")
# -
covid_tor_df = covid_tor_df.reset_index(drop=True)
covid_tor_df.head()
covid_tor_df.count()
# Retrieve unique list of LCC names as will need to find corresponding mapping coordinates from separate datafram
unique_lcc = set(covid_tor_df['lcc_name'])
count_lcc = len(unique_lcc)
print(count_lcc)
# retrieve LCC mapping coordinates - https://open.toronto.ca/dataset/licensed-child-care-centres/
url = 'http://opendata.toronto.ca/childrens.services/licensed-child-care-centres/child-care.csv'
response = requests.get(url)
s = requests.get(url).text
lcc_df = pd.read_csv(StringIO(s))
lcc_df = lcc_df[['STR_NO', 'STREET','LOC_NAME','TOTSPACE','LONGITUDE','LATITUDE']]
lcc_df.head()
lcc_df['ADDRESS'] = lcc_df['STR_NO']+ " " +lcc_df['STREET']
lcc_df = lcc_df[['LOC_NAME','ADDRESS','TOTSPACE','LONGITUDE','LATITUDE']]
lcc_df.columns = ['lcc_name','Address','Total Capacity','Longitude','Latitude']
covid_tor_df.count()
# +
# due to the data quality issues:
# this left join returns more records than expected i.e. duplicates for some LCCs
# there are also many records where the address/longitude/latitude could not be returned
covid_lcc_df = covid_tor_df.merge(lcc_df, how='left',on=['lcc_name'])
# -
covid_lcc_df = covid_lcc_df[covid_lcc_df['Longitude'].notna()]
covid_lcc_df.head()
covid_lcc_df = covid_lcc_df.reset_index(drop=True)
from arcgis.gis import GIS
from arcgis.geocoding import geocode
from IPython.display import display
gis = GIS()
map1 = gis.map()
map1
# set the map's extent by geocoding the location
toronto = geocode("Toronto")[0]
map1.extent = toronto['extent']
covid_lcc_df.head()
covid_lcc_df = covid_lcc_df[['lcc_name','Address','Total Capacity','Longitude','Latitude']]
covid_lcc_df.columns = ['Name','Address', 'Total Capacity','Longitude','Latitude']
covid_lcc_df[covid_lcc_df['Longitude'].isnull()]
covid_lcc_df = covid_lcc_df.reset_index()
del covid_lcc_df['index']
covid_lcc_df = covid_lcc_df.drop_duplicates()
covid_lcc = gis.content.import_data(covid_lcc_df)
map1.add_layer(covid_lcc)
| .ipynb_checkpoints/COVID Toronto LCC-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="Se228pPjFl1W"
# # Understanding and implementing GRU
# GRU or Gated Recurrent Units are the also inspired form design of LSTM. Gated recurrent units were published in 2014 by Cho, et al. in a research paper named as Learning Phrase Representations using RNN Encoder–Decoder for Statistical Machine Translation.
#
# As we had gated in LSTM, GRU has 2 gates update, and reset gate. These two gates decide what information should be discarded and what information should be let pass through. Learnable parameters in these two gates can be trained to timely change the information content and make continuous updates. The flow diagram for GRU looks like as given below
#
# 
#
# Figure: Showing in detail structure of the GRU unit
# The four gates function are as follow
#
# **Update Gate:** This gate can be given by the following formula :
#
# $$z_t = \sigma (W^{(z)}X_t + U^{(z)}h_{t-1}) $$
#
# here the input $X_t$ is multiplied by its weight and previously hidden tensor which is carrying information of previous $t-1$ is multiplied by its weight. Then sigmoid squash them into a number between 1 and 0. Update gate determines how much past information to let go to the present time step. This gate helps in solving the problem related to the vanishing gradient. If the sigmoid gate value is 1 then all the information is preserved and solves vanishing gradient problem.
#
# **Reset gate: **This gate helps in how much information needs to be forgotten from the previous time steps
#
# $$r_t = \sigma(W^{(r)}x_t + U^{(r)}h_{h-1}) $$
#
# This equation seems to be very similar to the previous equation. Here the only difference is the weights are for reset gate. Next is using these gates to determine current memory content and final memory at the end to the output.
#
# **Current memory content:** This derived current memory content using reset gates value and current input value. as er discussed previously that the reset gate knows how much information to forget and it has a number between 0 and 1. if $ r_t $ is zero then the input information contained in the current time step will be ignored fully and if 1 then the entire information in the current input will be taken into cell state. The current memory content is calculated in the following way.
#
# 1. Taking Hadamard product of reset gate value $r_t$ and previous hidden state with its weight $Uh_{t-1}$.
# 2. Summing up above value with of $W_{x_t}$
# $$ h_t^{'} = tanh (Wx_t + r_t \odot Uh_{t-1}) $$
#
# **Final Memory at current time step:** final memory is constructed by taking help of the update gates result and the current memory content. Final Memory is formed by using the following steps.
#
# 1. Taking Hadamard product of update gate value and $h_{t-1}$ .
# 2. Taking Hadamard product of and current memory content $h^`_t$
#
#
#
# Summing up above two values
#
# $$h_t = z_t \odot h_{t-1} + (1-z_t) \odot h_t^{'} $$
#
# + [markdown] colab_type="text" id="DqQ1RzJ08OWU"
# # Importing Requirements
# + colab={"base_uri": "https://localhost:8080/", "height": 833} colab_type="code" id="6TEFf4WfvLRW" outputId="238b092a-10e2-4f5a-a956-798017af8437"
import json
import os
import random
import tarfile
import urllib
import zipfile
import chakin
import matplotlib.pyplot as plt
import nltk
import torch
from torch import nn, optim
from torchtext import data
from torchtext import vocab
from tqdm import tqdm
nltk.download('popular')
SEED = 1234
torch.manual_seed(SEED)
torch.cuda.manual_seed(SEED)
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# + [markdown] colab_type="text" id="2FdXgOS6nkHF"
# # Downloading required datasets
# To demonstrate how embeddings can help, we will be conducting an experiment on sentiment analysis task. I have used movie review dataset having 5331 positive and 5331 negative processed sentences. The entire experiment is divided into 5 sections.
#
# Downloading Dataset: Above discussed dataset is available at http://www.cs.cornell.edu/people/pabo/movie-review-data/rt-polaritydata.tar.gz.
#
#
# + colab={} colab_type="code" id="JtpoPziv8GJO"
data_exists = os.path.isfile('data/rt-polaritydata.tar.gz')
if not data_exists:
urllib.request.urlretrieve("http://www.cs.cornell.edu/people/pabo/movie-review-data/rt-polaritydata.tar.gz",
"data/rt-polaritydata.tar.gz")
tar = tarfile.open("data/rt-polaritydata.tar.gz")
tar.extractall(path='data/')
# + [markdown] colab_type="text" id="pG6zlPuBnkHI"
# # Downloading embedding
# The pre-trained embeddings are available and can be easily used in our model. we will be using the GloVe vector trained having 300 dimensions.
# + colab={"base_uri": "https://localhost:8080/", "height": 306} colab_type="code" id="4Oab-YjwI7DU" outputId="9b8efc41-3582-4622-87e0-171f9b168c2c"
embed_exists = os.path.isfile('../embeddings/glove.840B.300d.zip')
if not embed_exists:
print("Downloading Glove embeddings, if not downloaded properly, then delete the `../embeddings/glove.840B.300d.zip")
chakin.search(lang='English')
chakin.download(number=16, save_dir='../embeddings')
zip_ref = zipfile.ZipFile("../embeddings/glove.840B.300d.zip", 'r')
zip_ref.extractall("../embeddings/")
zip_ref.close()
# + [markdown] colab_type="text" id="13BKiVJhnkHM"
# # Preprocessing
# I am using TorchText to preprocess downloaded data. The preprocessing includes following steps:
#
# - Reading and parsing data
# - Defining sentiment and label fields
# - Dividing data into train, valid and test subset
# - forming the train, valid and test iterators
# + colab={} colab_type="code" id="k3sTlJzyvLRb"
SEED = 1
split = 0.80
# + colab={} colab_type="code" id="85Uzm_t7vLRe"
data_block = []
negative_data = open('data/rt-polaritydata/rt-polarity.neg',encoding='utf8',errors='ignore').read().splitlines()
for i in negative_data:
data_block.append({"sentiment":str(i.strip()),"label" : 0})
positve_data = open('data/rt-polaritydata/rt-polarity.pos',encoding='utf8',errors='ignore').read().splitlines()
for i in positve_data:
data_block.append({"sentiment":str(i.strip()),"label" : 1})
# + colab={} colab_type="code" id="SzntAD2CvLRi"
random.shuffle(data_block)
train_file = open('data/train.json', 'w')
test_file = open('data/test.json', 'w')
for i in range(0,int(len(data_block)*split)):
train_file.write(str(json.dumps(data_block[i]))+"\n")
for i in range(int(len(data_block)*split),len(data_block)):
test_file.write(str(json.dumps(data_block[i]))+"\n")
# + colab={} colab_type="code" id="m2xfhN6avLRl"
def tokenize(sentiments):
# print(sentiments)
return sentiments
def pad_to_equal(x):
if len(x) < 61:
return x + ['<pad>' for i in range(0, 61 - len(x))]
else:
return x[:61]
def to_categorical(x):
if x == 1:
return [0,1]
if x == 0:
return [1,0]
# + colab={} colab_type="code" id="a8JZt1mBvLRp"
SENTIMENT = data.Field(sequential=True , preprocessing =pad_to_equal , use_vocab = True, lower=True)
LABEL = data.Field(is_target=True,use_vocab = False, sequential=False, preprocessing =to_categorical)
fields = {'sentiment': ('sentiment', SENTIMENT), 'label': ('label', LABEL)}
# + [markdown] colab_type="text" id="sUYeX3eenkHi"
# **Splitting data in to test and train**
# + colab={} colab_type="code" id="fq8L7GjZvLRu"
train_data , test_data = data.TabularDataset.splits(
path = 'data',
train = 'train.json',
test = 'test.json',
format = 'json',
fields = fields
)
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" id="3WzzpHkXvLRy" outputId="17429aef-21f5-4d8f-8ebf-c45ce5b6c870"
print("Printing an example data : ",vars(train_data[1]))
# + colab={} colab_type="code" id="FEZRHuB7vLR3"
train_data, valid_data = train_data.split(random_state=random.seed(SEED))
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="3CFyFa5_vLR7" outputId="503e0e4b-96a4-4345-ee34-013d7f4dc2b3"
print('Number of training examples: ', len(train_data))
print('Number of validation examples: ', len(valid_data))
print('Number of testing examples: ',len(test_data))
# + [markdown] colab_type="text" id="iWod-X4lnkHp"
# **Loading Embedding to vocab**
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="LUF_n6AivLSY" outputId="95419a98-e271-467d-ac97-ebaa3659e93d"
vec = vocab.Vectors(name = "glove.840B.300d.txt",cache = "../embeddings/")
# + colab={} colab_type="code" id="_J_s0L0ovLSd"
SENTIMENT.build_vocab(train_data, valid_data, test_data, max_size=100000, vectors=vec)
# + [markdown] colab_type="text" id="mqWn6jGWnkHy"
# **Constructing Iterators**
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="qZKthaRvvLSj" outputId="a6a6202e-473a-47f4-cd45-27c23ca90873"
train_iter, val_iter, test_iter = data.Iterator.splits(
(train_data, valid_data, test_data), sort_key=lambda x: len(x.sentiment),
batch_sizes=(32,32,32), device=-1,)
# + colab={} colab_type="code" id="rLTLVlHQvLSr"
sentiment_vocab = SENTIMENT.vocab
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="X46F3yZE0Gx9" outputId="8598fd72-7005-4e27-b1a0-4bca3abf6a0e"
sentiment_vocab.vectors.shape
# + [markdown] colab_type="text" id="zlpB4gbKnkIB"
# # Training
# Training will be conducted for two models one with GRU pre-trained embedding and one with LSTM. I am using GloVe embeddings with a vector size of 300.
# One thing to note here is the GRU is using only one hidden state to deal with vanishing gradient problem whereas the LSTM uses two hidden states. Due to this GRU is a bit faster than the LSTM. Let's see their performance on the movie review dataset.
# + colab={} colab_type="code" id="p8sGtYZ1-Bkl"
def binary_accuracy(preds, y):
"""
Returns accuracy per batch, i.e. if you get 8/10 right, this returns 0.8, NOT 8
"""
rounded_preds = torch.argmax(preds, dim=1)
# print(rounded_preds)
correct = (rounded_preds == torch.argmax(y, dim=1)).float() #convert into float for division
acc = correct.sum()/len(correct)
return acc
# + [markdown] colab_type="text" id="QqPGWHX5B9Gj"
# ## Training using GRU
# + colab={} colab_type="code" id="cHhzpI4D41ZL"
class GRU_RNN(nn.Module):
def __init__(self, vocab_size, embedding_dim, hidden_dim, output_dim, n_layers, bidirectional, dropout, sentiment_vocab):
super(GRU_RNN, self).__init__()
self.embedding = nn.Embedding(vocab_size, embedding_dim)
self.rnn = nn.GRU(embedding_dim, hidden_dim, num_layers=n_layers, bidirectional=bidirectional, dropout=dropout)
self.fc = nn.Linear(hidden_dim * 2, output_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, x):
embedded = self.dropout(self.embedding(x))
output, hidden = self.rnn(embedded)
# concat the final forward (hidden[-2,:,:]) and backward (hidden[-1,:,:]) hidden layers
# and apply dropout
hidden = self.dropout(torch.cat((hidden[-2, :, :], hidden[-1, :, :]), dim=1))
return torch.softmax(self.fc(hidden.squeeze(0)),dim = 1)
# + colab={} colab_type="code" id="CRl3m_aN9mQN"
INPUT_DIM = len(SENTIMENT.vocab)
EMBEDDING_DIM = 300
HIDDEN_DIM = 256
OUTPUT_DIM = 2
BATCH_SIZE = 32
N_LAYERS = 2
BIDIRECTIONAL = True
DROPOUT = 0.5
gru_rnn = GRU_RNN(INPUT_DIM, EMBEDDING_DIM, HIDDEN_DIM, OUTPUT_DIM, N_LAYERS, BIDIRECTIONAL, DROPOUT, sentiment_vocab)
gru_rnn = gru_rnn.to(device)
# + colab={} colab_type="code" id="rcUokNSt9yGl"
optimizer = optim.SGD(gru_rnn.parameters(), lr=0.1)
criterion = nn.BCEWithLogitsLoss()
criterion = criterion.to(device)
# + colab={} colab_type="code" id="mSU<KEY>"
def train(gru_rnn, iterator, optimizer, criterion):
epoch_loss = 0
epoch_acc = 0
# vanila_rnn.train()
for batch in iterator:
optimizer.zero_grad()
predictions = gru_rnn(batch.sentiment.to(device)).squeeze(1)
loss = criterion(predictions.type(torch.FloatTensor), batch.label.type(torch.FloatTensor))
acc = binary_accuracy(predictions.type(torch.FloatTensor), batch.label.type(torch.FloatTensor))
loss.backward()
optimizer.step()
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
# + colab={"base_uri": "https://localhost:8080/", "height": 3451} colab_type="code" id="qkqjMC8e-Xzm" outputId="ad3eeffc-0044-4115-e5a2-f0466481b93d"
rnn_loss = []
rnn_accuracy = []
for i in tqdm(range(0,100)):
loss, accuracy = train(gru_rnn, train_iter, optimizer, criterion)
print("Loss : ",loss, "Accuracy : ", accuracy )
rnn_loss.append(loss)
rnn_accuracy.append(accuracy)
# + [markdown] colab_type="text" id="aPqBr_3LCFTp"
# ## Training using LSTM
# + colab={} colab_type="code" id="iAEv9frrjlMN"
class LSTM_RNN(nn.Module):
def __init__(self, vocab_size, embedding_dim, hidden_dim, output_dim, n_layers, bidirectional, dropout, sentiment_vocab):
super(LSTM_RNN, self).__init__()
self.embedding = nn.Embedding(vocab_size, embedding_dim)
self.rnn = nn.LSTM(embedding_dim, hidden_dim, num_layers=n_layers, bidirectional=bidirectional, dropout=dropout)
self.fc = nn.Linear(hidden_dim * 2, output_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, x):
embedded = self.dropout(self.embedding(x))
output, (hidden, cell)= self.rnn(embedded)
# concat the final forward (hidden[-2,:,:]) and backward (hidden[-1,:,:]) hidden layers
# and apply dropout
hidden = self.dropout(torch.cat((hidden[-2, :, :], hidden[-1, :, :]), dim=1))
return self.fc(hidden.squeeze(0))
# + colab={} colab_type="code" id="z2GsTDmPjlMS"
INPUT_DIM = len(SENTIMENT.vocab)
EMBEDDING_DIM = 300
HIDDEN_DIM = 256
OUTPUT_DIM = 2
BATCH_SIZE = 32
N_LAYERS = 2
BIDIRECTIONAL = True
DROPOUT = 0.5
lstm_rnn = LSTM_RNN(INPUT_DIM, EMBEDDING_DIM, HIDDEN_DIM, OUTPUT_DIM, N_LAYERS, BIDIRECTIONAL, DROPOUT, sentiment_vocab)
lstm_rnn = lstm_rnn.to(device)
# + colab={} colab_type="code" id="LD-nHs1Qz8AT"
optimizer = optim.SGD(lstm_rnn.parameters(), lr=0.1)
criterion = nn.BCEWithLogitsLoss()
criterion = criterion.to(device)
# + colab={"base_uri": "https://localhost:8080/", "height": 3451} colab_type="code" id="TLdlbaND-bvv" outputId="714478c8-b460-4c67-ec27-cc6911698d48"
lstm_loss = []
lstm_accuracy = []
for i in tqdm(range(0,100)):
loss, accuracy = train(lstm_rnn, train_iter, optimizer, criterion)
print("Loss : ",loss, "Accuracy : ", accuracy )
lstm_loss.append(loss)
lstm_accuracy.append(accuracy)
# + [markdown] colab_type="text" id="CFT-Uv9UCOcm"
# ## Comparision
# As shown in the above LSTM produce 95% accuracy and GRU produced 85% performance. However, for all the datasets this will not be the case some time GRU's performance was also found to be superior in some cases.
#
# 
#
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 283} colab_type="code" id="mkMZnGsRDgO0" outputId="b08d59e5-0061-4ba7-b533-e01909b2185c"
plt.plot(rnn_accuracy , label = "GRU Accuracy")
plt.plot(lstm_accuracy , label = "LSTM Accuracy")
plt.ylabel("Accuracy")
plt.xlabel("Epoch")
plt.legend(loc='upper left')
plt.show()
# + colab={} colab_type="code" id="sAqzXv9WLRfh"
| Chapter04/gru_and_lstm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="WBMC0GrjNRTM"
# # Genertors and Random Numbers
# This is our week 4 examples notebook and will be available on Github from the powderflask/cap-comp215 repository.
#
# As usual, the first code block just imports the modules we will use.
# + pycharm={"name": "#%%\n"}
import random
import matplotlib.pyplot as plt
from pprint import pprint
# -
# ## Examples: generator expressions
# + pycharm={"name": "#%%\n"}
neighbours = ((i-1, i, i+1) for i in range(1, 10))
print(neighbours)
# We can turn a generator into a list to look at all its elements (though this somewhat defeats the purpose!)
print(list(neighbours))
# Once the "stream" has flowed past, it is empty - there is no way to "replenish" it.
print(list(neighbours))
# -
# ## Pseudo-Random Numbers
# + pycharm={"name": "#%%\n"}
# A list of n random numbers - again we are turning a "generator" in to a list
n = 10
[random.random() for i in range(n)]
# + pycharm={"name": "#%%\n"}
# Scatter plot of random (x,y) coordinates
n = 20
fig, ax = plt.subplots()
ax.scatter([random.random() for i in range(n)], [random.random() for i in range(n)])
ax
# -
# ## Chaotic Feedback
#
# * some mathematical functions produce chaotic patterns when you feed their output back in as the next input.
# + pycharm={"name": "#%%\n"}
import time
class TwoDigitRand:
""" demonstrates 'chaotic feedback' algorithm - not crypographic quality randomness!! """
def __init__(self, seed=None):
seed = seed or int(time.time()*10**3) # clock time in milli-seconds
self.seed = seed % 100 # 2-digit only!
def rand(self):
""" chaotic feedback algorithm - the last output is used as input to compute the next one """
self.seed = self.seed//10 + 7*(self.seed%10)
return self.seed
r = TwoDigitRand()
[r.rand() for i in range(20)]
# + pycharm={"name": "#%%\n"}
# Scatter plot of (seed, rand(seed))
fig, ax = plt.subplots()
print("Sequence seed:", r.seed)
pairs = [(r.seed, r.rand()) for i in range(n)]
ax.plot([x for x,y in pairs], [y for x,y in pairs])
ax
# -
# ## Example: computational experiment with random numbers
#
# We'll use the Histogram class developed last week to count the outcomes of the experiment
# + pycharm={"name": "#%%\n"}
class Histogram:
""" A simple histogram with a nice API """
def __init__(self, title, xlabel=None):
fig, ax = plt.subplots()
ax.set_title(title)
if xlabel:
ax.set_xlabel(xlabel)
ax.set_ylabel('Count')
self.ax = ax
self.fig = fig
self.counts = {}
def count(self, category):
self.counts[category] = self.counts.get(category, 0) + 1
def plot(self):
self.ax.bar(self.counts.keys(), self.counts.values())
plt.show()
# + pycharm={"name": "#%%\n"}
# Experiment: Roll some dice
def roll_dice(sides=6):
return (random.randint(1,sides), random.randint(1,sides))
# number rolls to use in this experiment
n_rolls = 1000
rolls = Histogram("Distribution of fair dice rolls", xlabel='Sum of 2 dice')
for i in range(n_rolls):
rolls.count(sum(roll_dice()))
rolls.plot()
# + pycharm={"name": "#%%\n"}
| examples/week4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from sklearn.cluster import KMeans
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
from sklearn.decomposition import PCA
DATA_PATH = './data/'
# + pycharm={"name": "#%%\n"}
corpus = []
file_list = []
for i in range(1000):
file_list.append('News_' + str(i + 1) + '_C.txt')
for file_name in file_list:
file_path = DATA_PATH + file_name
file = open(file_path, encoding='utf-8')
corpus.append(file.read())
# + pycharm={"name": "#%%\n"}
# 分词向量化
vectorizer = CountVectorizer()
word_vec = vectorizer.fit_transform(corpus)
# + pycharm={"name": "#%%\n"}
#提取TF-IDF词向量
transformer = TfidfTransformer()
tfidf = transformer.fit_transform(word_vec)
tfidf_matrix = tfidf.toarray() #对应tfidf矩阵
# + pycharm={"name": "#%%\n"}
#压缩 Feature
pca = PCA(n_components=3)
tfidf_matrix = pca.fit_transform(tfidf_matrix)
# + pycharm={"name": "#%%\n"}
# K-Means
km_cluster = KMeans(n_clusters=20, max_iter=300, n_init=2,
init='random', verbose=False)
# 返回各自文本的所被分配到的类索引
labels = km_cluster.fit_predict(tfidf_matrix)
markers = ['^', '^', '^', '^', '^', '^', '^', 'o', 'o', 'o', 'o', 'o', 'o', '+', '+', '+', '+', '+', '+', 's', 's']
colors = ['r', 'g', 'b', 'm', 'k', 'y', 'g', 'r', 'b', 'm', 'k', 'y', 'b', 'g', 'r', 'm', 'k', 'y', 'r', 'g' ]
fig, ax = plt.subplots(1, 1, figsize=(10, 8))
for i in range(len(labels)):
plt.scatter(tfidf_matrix[i, 0], tfidf_matrix[i, 1], c=colors[labels[i]], marker=markers[labels[i]])
fig.savefig('./kMeans.png', transparent=False, dpi=600, bbox_inches="tight")
# +
# 三维可视化
label_pred = km_cluster.labels_ # 获取聚类标签
centroids = km_cluster.cluster_centers_ # 获取聚类中心
print(label_pred)
print(centroids)
fig = plt.figure()
ax = Axes3D(fig)
ax.scatter(tfidf_matrix[:, 0], tfidf_matrix[:, 1], tfidf_matrix[:, 2], c=labels, marker='*')
ax.scatter(centroids[:, 0], centroids[:, 1], centroids[:, 2], marker='>')
plt.axis([1, 0, 0, 1])
plt.show()
| Spider/K-Means_CN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Pandas is to work with the data
import pandas as pd
import numpy as np
#Matplot is to plot charts
import matplotlib.pyplot as plt
#Seaborn is to make the charts look prettier
import seaborn as sns; sns.set()
# %matplotlib inline
sns.set(style="darkgrid")
plt.style.use('classic')
pd.options.display.max_rows = 20
# -
#how do we use seaborn
sns.set(style="ticks")
#https://en.wikipedia.org/wiki/World_record_progression_50_metres_freestyle
df = pd.read_csv('Swimming_WRP.csv')
df = df.rename(index=str, columns = {"Age " : "Age"})
df_dropna = df.dropna()
df
# This data was collected from Wikipedia. Every stroke was initially seperated into its own data table so I used excel to combine it all into one big table.
# The columns "Gender" and "Stroke" and "Country were all added to the table table using excel. This allowed me to better represent the data by gender, event, and country.The column "Location" has the name of the of the city and country separated by a comma. For my purposes I only wanted the name of the country for each record set. I know that it is possible to loop through each row and only take the value after the comma but I could not get the code to work properly. So, instead I made a new column named "Country" and inserted the name of the country that each world record holder represented for. After cleaning the data I presented it as shown above.
g = sns.FacetGrid(df,col="Stroke", hue="Gender", aspect=1.5, col_wrap=2)
g.map(plt.scatter, "Date", "Age")
plt.subplots_adjust(top=0.9)
g.fig.suptitle('Figure 1: Age of World Record Holders')
g.axes[0].get_xaxis().get_major_formatter().set_useOffset(False)
g.add_legend();
# +
#g = sns.FacetGrid(df, col="Stroke", aspect=1.5, col_wrap=2)
#g.map(plt.scatter, "Date", "Time")
#g.axes[0].get_xaxis().get_major_formatter().set_useOffset(False)
#g.add_legend();
#g = sns.scatterplot(x="Date", y="Time", data=df)
g = sns.FacetGrid(df,col="Stroke", hue="Gender", aspect=1.5, col_wrap=2)
g.map(plt.plot, "Date", "Time")
plt.subplots_adjust(top=0.9)
g.fig.suptitle('Figure 2: World Record Progression')
g.axes[0].get_xaxis().get_major_formatter().set_useOffset(False)
g.add_legend();
#g = df.plot.scatter(x = "Date", y = "Time")
#g.xaxis.get_major_formatter().set_useOffset(False)
#plt.title("World Record Progression 50m")
#.set(xlim=(1960, 2020)
# +
g = sns.FacetGrid(df, col="Stroke", hue="Gender", aspect=1.5, col_wrap=2)
g.map(plt.scatter, "Age", "Time")
plt.subplots_adjust(top=0.9)
g.fig.suptitle('Figure 3:')
g.add_legend();
# -
| project1/Adams-Project1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Обнаружение статистически значимых отличий в уровнях экспрессии генов больных раком
# Это задание поможет вам лучше разобраться в методах множественной проверки гипотез и позволит применить ваши знания на данных из реального биологического исследования.
#
# #### В этом задании вы:
#
# вспомните, что такое t-критерий Стьюдента и для чего он применяется
# сможете применить технику множественной проверки гипотез и увидеть собственными глазами, как она работает на реальных данных
# почувствуете разницу в результатах применения различных методов поправки на множественную проверку
#
# ## Основные библиотеки и используемые методы:
#
# Библиотека scipy и основные статистические функции:http://docs.scipy.org/doc/scipy/reference/stats.html#statistical-functions
#
# Библиотека statmodels для методов коррекции при множественном сравнении:
#
# http://statsmodels.sourceforge.net/devel/stats.html
#
# Статья, в которой рассматриваются примеры использования statsmodels для множественной проверки гипотез:
#
# http://jpktd.blogspot.ru/2013/04/multiple-testing-p-value-corrections-in.html
#
# ## Описание используемых данных
#
# Данные для этой задачи взяты из исследования, проведенного в Stanford School of Medicine. В исследовании была предпринята попытка выявить набор генов, которые позволили бы более точно диагностировать возникновение рака груди на самых ранних стадиях.
#
# В эксперименте принимали участие 24 человек, у которых не было рака груди (normal), 25 человек, у которых это заболевание было диагностировано на ранней стадии (early neoplasia), и 23 человека с сильно выраженными симптомами (cancer).
#
#
# Ученые провели секвенирование биологического материала испытуемых, чтобы понять, какие из этих генов наиболее активны в клетках больных людей.
#
# Секвенирование — это определение степени активности генов в анализируемом образце с помощью подсчёта количества соответствующей каждому гену РНК.
#
# В данных для этого задания вы найдете именно эту количественную меру активности каждого из 15748 генов у каждого из 72 человек, принимавших участие в эксперименте.
#
# Вам нужно будет определить те гены, активность которых у людей в разных стадиях заболевания отличается статистически значимо.
#
# Кроме того, вам нужно будет оценить не только статистическую, но и практическую значимость этих результатов, которая часто используется в подобных исследованиях.
#
# Диагноз человека содержится в столбце под названием "Diagnosis".
#
# ## Практическая значимость изменения
#
# Цель исследований — найти гены, средняя экспрессия которых отличается не только статистически значимо, но и достаточно сильно. В экспрессионных исследованиях для этого часто используется метрика, которая называется fold change (кратность изменения). Определяется она следующим образом:
#
# $$F_{c}(C,T) = \begin{cases} \frac{T}{C}, T>C \\ -\frac{C}{T}, T<C \end{cases}$$
#
# где C,T — средние значения экспрессии гена в control и treatment группах соответственно. По сути, fold change показывает, во сколько раз отличаются средние двух выборок.
#
# ## Инструкции к решению задачи
#
# Задание состоит из трёх частей. Если не сказано обратное, то уровень значимости нужно принять равным 0.05.
#
# ### Часть 1: применение t-критерия Стьюдента
#
# В первой части вам нужно будет применить критерий Стьюдента для проверки гипотезы о равенстве средних в двух независимых выборках. Применить критерий для каждого гена нужно будет дважды:
#
# * для групп **normal (control)** и **early neoplasia (treatment)**
# * для групп **early neoplasia (control)** и **cancer (treatment)**
#
# В качестве ответа в этой части задания необходимо указать количество статистически значимых отличий, которые вы нашли с помощью t-критерия Стьюдента, то есть число генов, у которых p-value этого теста оказался меньше, чем уровень значимости.
import pandas as pd
import scipy.stats
df = pd.read_csv("gene_high_throughput_sequencing.csv")
control_df = df[df.Diagnosis == 'normal']
neoplasia_df = df[df.Diagnosis == 'early neoplasia']
cancer_df = df[df.Diagnosis == 'cancer']
# +
# scipy.stats.ttest_ind(data.Placebo, data.Methylphenidate, equal_var = False)
# -
genes = filter(lambda x: x not in ['Patient_id', 'Diagnosis'], df.columns.tolist())
control_vs_neoplasia = {}
neoplasia_vs_cancer = {}
for gene in genes:
control_vs_neoplasia[gene] = scipy.stats.ttest_ind(control_df[gene], neoplasia_df[gene], equal_var = False).pvalue
neoplasia_vs_cancer[gene] = scipy.stats.ttest_ind(cancer_df[gene], neoplasia_df[gene], equal_var = False).pvalue
print control_df['LOC643837'],neoplasia_df['LOC643837']
scipy.stats.ttest_ind(control_df['LOC643837'], neoplasia_df['LOC643837'], equal_var = False).pvalue
control_vs_neoplasia_df = pd.DataFrame.from_dict(control_vs_neoplasia, orient = 'index')
control_vs_neoplasia_df.columns = ['control_vs_neoplasia_pvalue']
neoplasia_vs_cancer_df = pd.DataFrame.from_dict(neoplasia_vs_cancer, orient = 'index')
neoplasia_vs_cancer_df.columns = ['neoplasia_vs_cancer_pvalue']
neoplasia_vs_cancer_df
pvalue_df = control_vs_neoplasia_df.join(neoplasia_vs_cancer_df)
pvalue_df.head()
pvalue_df[pvalue_df.control_vs_neoplasia_pvalue < 0.05].shape
pvalue_df[pvalue_df.neoplasia_vs_cancer_pvalue < 0.05].shape
# ### Часть 2: поправка методом Холма
#
# Для этой части задания вам понадобится модуль multitest из statsmodels.
#
# `import statsmodels.stats.multitest as smm`
#
# В этой части задания нужно будет применить поправку Холма для получившихся двух наборов достигаемых уровней значимости из предыдущей части. Обратите внимание, что поскольку вы будете делать поправку для каждого из двух наборов p-value отдельно, то проблема, связанная с множественной проверкой останется.
#
# Для того, чтобы ее устранить, достаточно воспользоваться поправкой Бонферрони, то есть использовать уровень значимости 0.05 / 2 вместо 0.05 для дальнейшего уточнения значений p-value c помощью метода Холма.
#
# В качестве ответа к этому заданию требуется ввести количество значимых отличий в каждой группе после того, как произведена коррекция Холма-Бонферрони. Причем это число нужно ввести с учетом практической значимости: посчитайте для каждого значимого изменения fold change и выпишите в ответ число таких значимых изменений, абсолютное значение fold change которых больше, чем 1.5.
#
# **Обратите внимание, что**
#
# применять поправку на множественную проверку нужно ко всем значениям достигаемых уровней значимости, а не только для тех, которые меньше значения уровня доверия.
# при использовании поправки на уровне значимости 0.025 меняются значения достигаемого уровня значимости, но не меняется значение уровня доверия (то есть для отбора значимых изменений скорректированные значения уровня значимости нужно сравнивать с порогом 0.025, а не 0.05)!
#
import statsmodels.stats.multitest as smm
pvalue_df['control_mean_expression'] = control_df.mean()
pvalue_df['neoplasia_mean_expression'] = neoplasia_df.mean()
pvalue_df['cancer_mean_expression'] = cancer_df.mean()
def abs_fold_change(c, t):
if t > c:
return t/c
else:
return c/t
pvalue_df['control_vs_neoplasia_fold_change'] = map(lambda x, y: abs_fold_change(x, y),
pvalue_df.control_mean_expression,
pvalue_df.neoplasia_mean_expression
)
pvalue_df['neoplasia_vs_cancer_fold_change'] = map(lambda x, y: abs_fold_change(x, y),
pvalue_df.neoplasia_mean_expression,
pvalue_df.cancer_mean_expression
)
pvalue_df['control_vs_neoplasia_rej_hb'] = smm.multipletests(pvalue_df.control_vs_neoplasia_pvalue, alpha=0.025, method='h')[0]
pvalue_df['neoplasia_vs_cancer_rej_hb'] = smm.multipletests(pvalue_df.neoplasia_vs_cancer_pvalue, alpha=0.025, method='h')[0]
pvalue_df[(pvalue_df.control_vs_neoplasia_rej_hb) & (pvalue_df.control_vs_neoplasia_fold_change > 1.5)].shape
pvalue_df[(pvalue_df.neoplasia_vs_cancer_rej_hb) & (pvalue_df.neoplasia_vs_cancer_fold_change > 1.5)].shape
# ### Часть 3: поправка методом Бенджамини-Хохберга
#
# Данная часть задания аналогична второй части за исключением того, что нужно будет использовать метод Бенджамини-Хохберга.
#
# Обратите внимание, что методы коррекции, которые контролируют FDR, допускает больше ошибок первого рода и имеют большую мощность, чем методы, контролирующие FWER. Большая мощность означает, что эти методы будут совершать меньше ошибок второго рода (то есть будут лучше улавливать отклонения от H0, когда они есть, и будут чаще отклонять H0, когда отличий нет).
#
# В качестве ответа к этому заданию требуется ввести количество значимых отличий в каждой группе после того, как произведена коррекция Бенджамини-Хохберга, причем так же, как и во второй части, считать только такие отличия, у которых abs(fold change) > 1.5.
#
pvalue_df['control_vs_neoplasia_rej_bh'] = smm.multipletests(pvalue_df.control_vs_neoplasia_pvalue, alpha=0.025, method='fdr_i')[0]
pvalue_df['neoplasia_vs_cancer_rej_bh'] = smm.multipletests(pvalue_df.neoplasia_vs_cancer_pvalue, alpha=0.025, method='fdr_i')[0]
pvalue_df.control_vs_neoplasia_rej_bh.value_counts()
pvalue_df[(pvalue_df.control_vs_neoplasia_rej_bh) & (pvalue_df.control_vs_neoplasia_fold_change > 1.5)].shape
pvalue_df[(pvalue_df.neoplasia_vs_cancer_rej_bh) & (pvalue_df.neoplasia_vs_cancer_fold_change > 1.5)].shape
| statistics/экспрессия генов.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SLU08 - Classification: Example notebook
# How to use the very useful sklearn's implementation of:
# - LogisticRegression
#
# to solve the last exercise of the Exercise Notebook of SLU08.
# +
import pandas as pd
import numpy as np
from sklearn.preprocessing import MinMaxScaler
from sklearn.linear_model import LogisticRegression
# -
# ### The Banknote Authentication Dataset
#
# There are 1372 items (images of banknotes — think Euro or dollar bill). There are 4 predictor variables (variance of image, skewness, kurtosis, entropy). The variable to predict is encoded as 0 (authentic) or 1 (forgery).
#
# Your quest, is to first analyze this dataset from the materials that you've learned in the previous SLUs and then create a logistic regression model that can correctly classify forged banknotes from authentic ones.
#
# The data is loaded for you below.
columns = ['variance','skewness','kurtosis','entropy', 'forgery']
data = pd.read_csv('data/data_banknote_authentication.txt',names=columns).sample(frac=1, random_state=1)
X_train = data.drop(columns='forgery').values
Y_train = data.forgery.values
# How does the dataset (features) and target look like?
X_train
Y_train
# # [MinMaxScaler](http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.MinMaxScaler.html)
# _Transforms features by scaling each feature to a given range._
#
# You can select the range for your final feature values with argument `feature_range=(0, 1)`
# +
# Init class
scaler = MinMaxScaler(feature_range=(0, 1))
# Fit your class
scaler.fit(X_train)
# -
# Transform your data
X_train = scaler.transform(X_train)
X_train
# So, now our features are scaled between 0 and 1.
# # [LogisticRegression](http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LogisticRegression.html)
# _Logistic Regression (aka logit, MaxEnt) classifier._ In this case let us use the L2 penalty (argument: `penalty='l2'`)
# +
# init with your arguments
logit_clf = LogisticRegression(penalty='l2', random_state=1)
# Fit it!
logit_clf.fit(X_train, Y_train)
# -
# What are the predicted probabilities on the training data (probability of being `1`) with our **Logit** classifier for the first 10 samples?
# First ten instances
logit_clf.predict_proba(X_train)[:, 1][:10]
# What about the predicted classes?
# First ten instances
logit_clf.predict(X_train)[:10]
# And the accuracy?
logit_clf.score(X_train, Y_train)
# How can we change the threshold from the default (0.5) to 0.9?
predictions = logit_clf.predict_proba(X_train)[:, 1]
predictions[predictions>=0.9] = 1
predictions[predictions<0.9] = 0
predictions[:10]
| S01 - Bootcamp and Binary Classification/SLU08 - Classification with Logistic Regression/Example notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf
# language: python
# name: tf
# ---
# <h1 style='font-size:20px;text-align: center'>
# A cloud effective radius (CER) and optical thickness (COT) retrieval framework using python; scikit-learn and TensorFlow:
# Application to the Moderate Resolution Imaging Spectroradiometer (MODIS) on board NASA's Terra & Aqua satellites.
# </h1>
#
# <p style='text-align: center'>Author: <NAME>
# <br><EMAIL>
# <br> <EMAIL>
# </p>
# +
import tensorflow as tf
import matplotlib.pyplot as plt
import numpy as np
import math
import matplotlib.patches as mpatches
import matplotlib.cm as cm
import pandas as pd
import matplotlib as mpl
import tensorflow_docs as tfdocs
import tensorflow_docs.plots
import tensorflow_docs.modeling
import os
import urllib.request
import tarfile
import warnings
from sklearn.gaussian_process import GaussianProcessRegressor
from sklearn.gaussian_process.kernels import RBF
from sklearn.gaussian_process.kernels import DotProduct, ConstantKernel as C
from sklearn import preprocessing
from pyhdf.SD import SD, SDC
from matplotlib.pyplot import figure
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import LinearRegression
from sklearn.pipeline import Pipeline
from sklearn.pipeline import make_pipeline
from sklearn.linear_model import Ridge
from pylab import figure, cm
from scipy import misc
from joblib import dump, load
from scipy.optimize import minimize, rosen, rosen_der
from tensorflow import keras
from tensorflow.keras import layers
warnings.filterwarnings('ignore')
# -
from IPython.core.display import HTML
HTML("""
<style>
.output_png {
display: table-cell;
text-align: center;
vertical-align: middle;
}
</style>
""")
# ### Download ice and liquid reflectance libraries
#
# [Look Up Tables (LUTs)](https://atmosphere-imager.gsfc.nasa.gov/products/cloud/luts)
# +
# %%time
if not os.path.exists('MODIS_C6_LUTS'):
# Download LUT libraries
url = 'https://atmosphere-imager.gsfc.nasa.gov/sites/default/files/ModAtmo/resources/modis_c6_luts.tar.gz'
downloaded_filename = 'modis_c6_luts.tar.gz'
urllib.request.urlretrieve(url, downloaded_filename)
# Unzip .tar.gz
# Ref: https://stackoverflow.com/questions/30887979/i-want-to-create-a-script-for-unzip-tar-gz-file-via-python
fname = 'modis_c6_luts.tar.gz'
if fname.endswith("tar.gz"):
tar = tarfile.open(fname, "r:gz")
tar.extractall()
tar.close()
# +
reflectance_ice_library = SD('./MODIS_C6_LUTS/modis_ice/reflectance_ice_library.hdf', SDC.READ)
print( reflectance_ice_library.info() )
# +
datasets_dic = reflectance_ice_library.datasets()
for idx,sds in enumerate(datasets_dic.keys()):
#print( idx,sds )
sds_obj = reflectance_ice_library.select(sds)
print( sds_obj.info() )
# +
MultiScatBDReflectance = reflectance_ice_library.select('MultiScatBDReflectance')
MultiScatBDReflectance = MultiScatBDReflectance.get()
print(MultiScatBDReflectance.shape)
fig = figure(num=None, figsize=(12, 10), dpi=80, facecolor='w', edgecolor='k')
ax = fig.add_subplot(111)
plt.imshow(MultiScatBDReflectance[8,3,13,:,4,:], cmap='hot', interpolation='nearest',origin='lower')
plt.colorbar()
ax.set_aspect('auto')
plt.show()
# +
OpticalThickness = reflectance_ice_library.select('OpticalThickness')
OpticalThickness = OpticalThickness.get()
print(OpticalThickness.shape)
print(OpticalThickness)
# +
ParticleRadius = reflectance_ice_library.select('ParticleRadius')
ParticleRadius = ParticleRadius.get()
print(ParticleRadius.shape)
print(ParticleRadius)
# +
y = MultiScatBDReflectance[8,3,13,:,4,:]
print(y.shape)
# -
print( y.max() )
print( y.min() )
# +
x1, x2 = np.meshgrid(np.arange(0,12, 1), np.arange(0,34, 1))
print(34*12)
x1 = x1.flatten()
x2 = x2.flatten()
x1 = ParticleRadius[x1]
x2 = OpticalThickness[x2]
y = y.flatten()
# -
X = np.stack((x1,x2),axis=-1)
X = pd.DataFrame(X,columns=['tau','cer'])
X_stats = X.describe()
X_stats = X_stats.transpose()
X_stats
def data_scaling_function(x):
return (x - X_stats['mean']) / X_stats['std']
# +
X = data_scaling_function(X)
test_stats = X.describe()
test_stats = test_stats.transpose()
test_stats
# -
y = pd.DataFrame(y,columns=['MultiScatBDReflectance'])
t = pd.concat((X,y),axis=1)
# +
t = t.sample(frac = 1)
t
# -
X_train = t[['tau','cer']]
y_train = t[['MultiScatBDReflectance']]
# +
def build_regression_model(intput_x):
model = keras.Sequential([
layers.Dense(10, activation='tanh', input_shape=[intput_x.shape[1]]),
layers.Dense(10, activation='tanh'),
layers.Dense(1)
])
opt = tf.keras.optimizers.RMSprop(0.01)
model.compile(loss='mse',
optimizer='adam',
metrics=['mse'])
return model
reg_model = build_regression_model(X_train)
# -
reg_model.fit(X_train, y_train, epochs=3000,
validation_split = 0.2, verbose=0, callbacks=[tfdocs.modeling.EpochDots()])
# +
X_test = np.stack((x1,x2),axis=-1)
X_test = pd.DataFrame(X_test,columns=['tau','cer'])
X_test = data_scaling_function(X_test)
# -
yhat = reg_model.predict(X_test)
print(yhat.min())
print(yhat.max())
# +
fig = figure(num=None, figsize=(12, 10), dpi=80, facecolor='w', edgecolor='k')
ax = fig.add_subplot(111)
#plt.imshow(yhat.reshape(34,12), cmap='hot', interpolation='nearest',origin='lower',vmin=0.0005173092,vmax=0.4885463)
plt.imshow(yhat.reshape(34,12), cmap='hot', interpolation='nearest',origin='lower',vmin=0.0005173092,vmax=0.4885463)
plt.colorbar()
ax.set_aspect('auto')
plt.show()
# +
y_test = y.to_numpy()
y_test = y_test.reshape(-1,1)
diff = np.sqrt( (yhat - y_test) * (yhat - y_test) )
for i in range(20):
print(yhat[i,0],y_test[i,0],diff[i,0])
# +
fig = figure(num=None, figsize=(12, 10), dpi=80, facecolor='w', edgecolor='k')
ax = fig.add_subplot(111)
plt.imshow(diff.reshape(34,12), cmap='hot', interpolation='nearest',origin='lower')
plt.colorbar()
ax.set_aspect('auto')
plt.show()
# +
x1, x2 = np.meshgrid(np.arange(0,60, 0.1), np.arange(0,160, 0.1))
x1 = x1.flatten()
x2 = x2.flatten()
X_test = np.stack((x1,x2),axis=-1)
X_test = pd.DataFrame(X_test,columns=['tau','cer'])
X_test = data_scaling_function(X_test)
yhat = reg_model.predict(X_test)
fig = figure(num=None, figsize=(12, 10), dpi=80, facecolor='w', edgecolor='k')
ax = fig.add_subplot(111)
plt.imshow(yhat.reshape(1600,600), cmap='hot', interpolation='nearest',origin='lower',vmin=0.0005173092,vmax=0.4885463)
plt.colorbar()
ax.set_aspect('auto')
plt.show()
# -
# +
y1 = MultiScatBDReflectance[8,3,13,:,0,:]
y2 = MultiScatBDReflectance[8,3,13,:,4,:]
y1 = y1.flatten()
y2 = y2.flatten()
Y = np.stack((y1,y2),axis=-1)
Y = pd.DataFrame(Y,columns=['band 1', 'band 6'])
Y
# +
t = pd.concat((X,Y),axis=1)
t = t.sample(frac = 1)
t
# -
X_train = t[['tau','cer']]
y_train = t[['band 1', 'band 6']]
# +
def build_multiple_regression_model(intput_x):
model = keras.Sequential([
layers.Dense(60, activation='tanh', input_shape=[intput_x.shape[1]]),
layers.Dense(60, activation='tanh'),
layers.Dense(60, activation='tanh'),
layers.Dense(2)
])
opt = tf.keras.optimizers.RMSprop(0.01)
model.compile(loss='mse',
optimizer='adam',
metrics=['mse'])
return model
mult_reg_model = build_multiple_regression_model(X_train)
# -
mult_reg_model.fit(X_train, y_train, epochs=20000,
validation_split = 0.2, verbose=0, callbacks=[tfdocs.modeling.EpochDots()])
# +
x1, x2 = np.meshgrid(np.arange(0,60, 1), np.arange(0,160, 1))
x1 = x1.flatten()
x2 = x2.flatten()
X_test = np.stack((x1,x2),axis=-1)
X_test = pd.DataFrame(X_test,columns=['tau','cer'])
X_test = data_scaling_function(X_test)
yhat = mult_reg_model.predict(X_test)
yhat
# +
fig = figure(num=None, figsize=(12, 10), dpi=80, facecolor='w', edgecolor='k')
ax = fig.add_subplot(111)
plt.imshow(yhat[:,0].reshape(160,60), cmap='hot', interpolation='nearest',origin='lower')
plt.colorbar()
ax.set_aspect('auto')
plt.show()
# +
fig = figure(num=None, figsize=(12, 10), dpi=80, facecolor='w', edgecolor='k')
ax = fig.add_subplot(111)
plt.imshow(yhat[:,1].reshape(160,60), cmap='hot', interpolation='nearest',origin='lower')
plt.colorbar()
ax.set_aspect('auto')
plt.show()
# -
# +
file = SD('./MODIS_C6_LUTS/modis_ice/reflectance_ice_library.hdf', SDC.READ)
x_band_idx = 0
y_band_idx = 4
ice_lut_shape = MultiScatBDReflectance.shape
figure(num=None, figsize=(12, 10), dpi=80, facecolor='w', edgecolor='k')
x = [MultiScatBDReflectance[8,3,13,:,x_band_idx,index_re] for index_re in np.arange(ice_lut_shape[5])]
y = [MultiScatBDReflectance[8,3,13,:,y_band_idx,index_re] for index_re in np.arange(ice_lut_shape[5])]
plt.plot(x,y, 'steelblue',linewidth=0.5)
x = [MultiScatBDReflectance[8,3,13,index_tau,x_band_idx,:] for index_tau in np.arange(ice_lut_shape[3])]
y = [MultiScatBDReflectance[8,3,13,index_tau,y_band_idx,:] for index_tau in np.arange(ice_lut_shape[3])]
plt.plot(x,y, 'coral',linewidth=0.5)
pop_a = mpatches.Patch(color='coral', label=r'Cloud Effective Radius ($\mu m$)')
pop_b = mpatches.Patch(color='steelblue', label='Cloud Optical Thickness')
plt.legend(handles=[pop_a,pop_b],fontsize=8)
plt.show()
plt.close()
# +
x1, x2 = np.meshgrid(np.arange(0,12, 1), np.arange(0,34, 1))
print(34*12)
x1 = x1.flatten()
x2 = x2.flatten()
x1 = ParticleRadius[x1]
x2 = OpticalThickness[x2]
X_test = np.stack((x1,x2),axis=-1)
X_test = pd.DataFrame(X_test,columns=['tau','cer'])
X_test = data_scaling_function(X_test)
yhat = mult_reg_model.predict(X_test)
yhat.shape
yhat = yhat.reshape(34,12,2)
# +
figure(num=None, figsize=(12, 10), dpi=80, facecolor='w', edgecolor='k')
x = [yhat[:,index_re,0] for index_re in np.arange(ice_lut_shape[5])]
y = [yhat[:,index_re,1] for index_re in np.arange(ice_lut_shape[5])]
plt.plot(x,y, 'steelblue',linewidth=0.5)
x = [yhat[index_tau,:,0] for index_tau in np.arange(ice_lut_shape[3])]
y = [yhat[index_tau,:,1] for index_tau in np.arange(ice_lut_shape[3])]
plt.plot(x,y, 'coral',linewidth=0.5)
pop_a = mpatches.Patch(color='steelblue', label=r'Cloud Effective Radius ($\mu m$)')
pop_b = mpatches.Patch(color='coral', label='Cloud Optical Thickness')
plt.legend(handles=[pop_a,pop_b],fontsize=8)
plt.show()
plt.close()
# +
figure(num=None, figsize=(12, 10), dpi=80, facecolor='w', edgecolor='k')
x = [MultiScatBDReflectance[8,3,13,:,x_band_idx,index_re] for index_re in np.arange(ice_lut_shape[5])]
y = [MultiScatBDReflectance[8,3,13,:,y_band_idx,index_re] for index_re in np.arange(ice_lut_shape[5])]
plt.plot(x,y, 'steelblue',linewidth=0.5)
x = [yhat[:,index_re,0] for index_re in np.arange(ice_lut_shape[5])]
y = [yhat[:,index_re,1] for index_re in np.arange(ice_lut_shape[5])]
plt.plot(x,y, 'steelblue',linewidth=0.5)
x = [MultiScatBDReflectance[8,3,13,index_tau,x_band_idx,:] for index_tau in np.arange(ice_lut_shape[3])]
y = [MultiScatBDReflectance[8,3,13,index_tau,y_band_idx,:] for index_tau in np.arange(ice_lut_shape[3])]
plt.plot(x,y, 'coral',linewidth=0.5)
x = [yhat[index_tau,:,0] for index_tau in np.arange(ice_lut_shape[3])]
y = [yhat[index_tau,:,1] for index_tau in np.arange(ice_lut_shape[3])]
plt.plot(x,y, 'coral',linewidth=0.5)
pop_a = mpatches.Patch(color='coral', label=r'Cloud Effective Radius ($\mu m$)')
pop_b = mpatches.Patch(color='steelblue', label='Cloud Optical Thickness')
plt.legend(handles=[pop_a,pop_b],fontsize=8)
plt.show()
plt.close()
# +
from graphviz import Digraph
from graphviz import Source
temp = '''
digraph G {
graph[ fontname = "Helvetica-Oblique",
fontsize = 12,
label = "",
size = "7.75,10.25" ];
rankdir = LR;
splines=false;
edge[style=invis];
ranksep= 1.4;
{
node [shape=circle, color=chartreuse, style=filled, fillcolor=chartreuse];
x1 [label=<CER>];
x2 [label=<TAU>];
}
{
node [shape=circle, color=dodgerblue, style=filled, fillcolor=dodgerblue];
a12 [label=<a<sub>1</sub><sup>(2)</sup>>];
a22 [label=<a<sub>2</sub><sup>(2)</sup>>];
a32 [label=<a<sub>3</sub><sup>(2)</sup>>];
a42 [label=<a<sub>4</sub><sup>(2)</sup>>];
a52 [label=<a<sub>5</sub><sup>(2)</sup>>];
a13 [label=<a<sub>1</sub><sup>(3)</sup>>];
a23 [label=<a<sub>2</sub><sup>(3)</sup>>];
a33 [label=<a<sub>3</sub><sup>(3)</sup>>];
a43 [label=<a<sub>4</sub><sup>(3)</sup>>];
a53 [label=<a<sub>5</sub><sup>(3)</sup>>];
}
{
node [shape=circle, color=coral1, style=filled, fillcolor=coral1];
O1 [label=<Band 1>];
O2 [label=<Band 6>];
O3 [label=<Band 7>];
}
{
rank=same;
x1->x2;
}
{
rank=same;
a12->a22->a32->a42->a52;
}
{
rank=same;
a13->a23->a33->a43->a53;
}
{
rank=same;
O1->O2->O3;
}
l0 [shape=plaintext, label="layer 1 (input layer)"];
l0->x1;
{rank=same; l0;x1};
l1 [shape=plaintext, label="layer 2 (hidden layer)"];
l1->a12;
{rank=same; l1;a12};
l2 [shape=plaintext, label="layer 3 (hidden layer)"];
l2->a13;
{rank=same; l2;a13};
l3 [shape=plaintext, label="layer 4 (output layer)"];
l3->O1;
{rank=same; l3;O1};
edge[style=solid, tailport=e, headport=w];
{x1; x2} -> {a12;a22;a32;a42;a52};
{a12;a22;a32;a42;a52} -> {a13;a23;a33;a43;a53};
{a13;a23;a33;a43;a53} -> {O1,O2,O3};
}'''
s = Source(temp)
s
# +
from graphviz import Digraph
from graphviz import Source
temp = '''
digraph G {
graph[ fontname = "Helvetica-Oblique",
fontsize = 12,
label = "",
size = "7.75,10.25" ];
rankdir = LR;
splines=false;
edge[style=invis];
ranksep= 1.4;
{
node [shape=circle, color=chartreuse, style=filled, fillcolor=chartreuse];
x1 [label=<x1>];
x2 [label=<x2>];
}
{
node [shape=circle, color=dodgerblue, style=filled, fillcolor=dodgerblue];
a12 [label=<a<sub>1</sub><sup>(2)</sup>>];
a22 [label=<a<sub>2</sub><sup>(2)</sup>>];
a32 [label=<a<sub>3</sub><sup>(2)</sup>>];
a42 [label=<a<sub>4</sub><sup>(2)</sup>>];
a52 [label=<a<sub>5</sub><sup>(2)</sup>>];
a13 [label=<a<sub>1</sub><sup>(3)</sup>>];
a23 [label=<a<sub>2</sub><sup>(3)</sup>>];
a33 [label=<a<sub>3</sub><sup>(3)</sup>>];
a43 [label=<a<sub>4</sub><sup>(3)</sup>>];
a53 [label=<a<sub>5</sub><sup>(3)</sup>>];
}
{
node [shape=circle, color=coral1, style=filled, fillcolor=coral1];
O1 [label=<y1>];
O2 [label=<y2>];
O3 [label=<y3>];
}
{
rank=same;
x1->x2;
}
{
rank=same;
a12->a22->a32->a42->a52;
}
{
rank=same;
a13->a23->a33->a43->a53;
}
{
rank=same;
O1->O2->O3;
}
l0 [shape=plaintext, label="layer 1 (input layer)"];
l0->x1;
{rank=same; l0;x1};
l1 [shape=plaintext, label="layer 2 (hidden layer)"];
l1->a12;
{rank=same; l1;a12};
l2 [shape=plaintext, label="layer 3 (hidden layer)"];
l2->a13;
{rank=same; l2;a13};
l3 [shape=plaintext, label="layer 4 (output layer)"];
l3->O1;
{rank=same; l3;O1};
edge[style=solid, tailport=e, headport=w];
{x1; x2} -> {a12;a22;a32;a42;a52};
{a12;a22;a32;a42;a52} -> {a13;a23;a33;a43;a53};
{a13;a23;a33;a43;a53} -> {O1,O2,O3};
}'''
s = Source(temp)
s
# -
| docs/train_deep_learning_model_from_modis_myd06_lut.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/PacktPublishing/Hands-On-Computer-Vision-with-PyTorch/blob/master/Chapter04/Image_augmentation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="8dqBSoIUfmoN"
import imgaug.augmenters as iaa
# + id="0Lrs6ufwfsRh"
from torchvision import datasets
import torch
data_folder = '/content/' # This can be any directory you want to download FMNIST to
fmnist = datasets.FashionMNIST(data_folder, download=True, train=True)
# + id="uNpLxbc6gGIl"
tr_images = fmnist.data
tr_targets = fmnist.targets
# + id="kuGaMbCGgHsd"
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
from torch.utils.data import Dataset, DataLoader
import torch
import torch.nn as nn
device = 'cuda' if torch.cuda.is_available() else 'cpu'
# + id="o2w1Ac2fgMEY" outputId="a8337b36-c6ce-4899-9f93-8c54bf4e7a6d" colab={"base_uri": "https://localhost:8080/", "height": 298}
plt.imshow(tr_images[0], cmap='gray')
plt.title('Original image')
# + id="x846czVPt6zf" outputId="39b48199-6882-4f3a-c9cf-a470d54a32f9" colab={"base_uri": "https://localhost:8080/", "height": 298}
aug = iaa.Affine(scale=2)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Scaled image')
# + id="px6aanrNt62B" outputId="34f60637-8e2e-4235-a682-c6fa7ce81ab2" colab={"base_uri": "https://localhost:8080/", "height": 298}
aug = iaa.Affine(translate_px=10)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Translated image by 10 pixels (right and bottom)')
# + id="svm9TAnJvbn6" outputId="c4a7c51b-7589-49db-b06a-684a03ae6cbd" colab={"base_uri": "https://localhost:8080/", "height": 314}
aug = iaa.Affine(translate_px={'x':10,'y':2})
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Translation of 10 pixels \nacross columns and 2 pixels over rows')
# + id="iCOcbS7sxpVg" outputId="a5ddf07a-125b-41a0-be7b-190d887d96d2" colab={"base_uri": "https://localhost:8080/", "height": 298}
aug = iaa.Affine(rotate=30)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Rotation of image by 30 degrees')
# + id="_JRiQsamxpXx" outputId="52ebfa9c-11c0-4544-83de-278631b7dacc" colab={"base_uri": "https://localhost:8080/", "height": 298}
aug = iaa.Affine(rotate=-30)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Rotation of image by -30 degrees')
# + id="WmxcpYbvx2ct" outputId="81edcd19-8d7f-4a99-de23-be16b07ee2ef" colab={"base_uri": "https://localhost:8080/", "height": 298}
aug = iaa.Affine(shear=30)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Shear of image by 30 degrees')
# + id="LQO41jTTx2fW" outputId="e93c8bfe-73e6-4171-e454-90448b57e9f4" colab={"base_uri": "https://localhost:8080/", "height": 298}
aug = iaa.Affine(shear=-30)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Shear of image by -30 degrees')
# + id="4RD-1tV0zuSm" outputId="504e918e-b18a-4868-8855-72137bbe1336" colab={"base_uri": "https://localhost:8080/", "height": 255}
plt.figure(figsize=(20,20))
plt.subplot(161)
plt.imshow(tr_images[0], cmap='gray')
plt.title('Original image')
plt.subplot(162)
aug = iaa.Affine(scale=2, fit_output=True)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Scaled image')
plt.subplot(163)
aug = iaa.Affine(translate_px={'x':10,'y':2}, fit_output=True)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Translation of 10 pixels across \ncolumns and 2 pixels over rows')
plt.subplot(164)
aug = iaa.Affine(rotate=30, fit_output=True)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Rotation of image \nby 30 degrees')
plt.subplot(165)
aug = iaa.Affine(shear=30, fit_output=True)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Shear of image \nby 30 degrees')
# + id="I4sSQxqPzuXa" outputId="71dc3ae0-b3c1-43c9-954b-679132d121df" colab={"base_uri": "https://localhost:8080/", "height": 255}
plt.figure(figsize=(20,20))
plt.subplot(161)
plt.imshow(tr_images[0], cmap='gray')
plt.title('Original image')
plt.subplot(162)
aug = iaa.Affine(scale=2, fit_output=True)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Scaled image')
plt.subplot(163)
aug = iaa.Affine(translate_px={'x':10,'y':2}, fit_output=True, cval = 255)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Translation of 10 pixels across \ncolumns and 2 pixels over rows')
plt.subplot(164)
aug = iaa.Affine(rotate=30, fit_output=True)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Rotation of image \nby 30 degrees')
plt.subplot(165)
aug = iaa.Affine(shear=30, fit_output=True)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Shear of image \nby 30 degrees')
# + id="qXCHS8M0zucP" outputId="f945c5f9-1630-45b3-dc29-299699ce8a2a" colab={"base_uri": "https://localhost:8080/", "height": 298}
aug = iaa.Affine(rotate=30, fit_output=True, cval=255)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Rotation of image by 30 degrees')
# + id="9JUsjQ8f4eZy" outputId="768ee835-3414-4dd6-bd36-561e837f3fbc" colab={"base_uri": "https://localhost:8080/", "height": 255}
plt.figure(figsize=(20,20))
plt.subplot(161)
aug = iaa.Affine(rotate=30, fit_output=True, cval=0, mode='constant')
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Rotation of image by \n30 degrees with constant mode')
plt.subplot(162)
aug = iaa.Affine(rotate=30, fit_output=True, cval=0, mode='edge')
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Rotation of image by 30 degrees \n with edge mode')
plt.subplot(163)
aug = iaa.Affine(rotate=30, fit_output=True, cval=0, mode='symmetric')
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Rotation of image by \n30 degrees with symmetric mode')
plt.subplot(164)
aug = iaa.Affine(rotate=30, fit_output=True, cval=0, mode='reflect')
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Rotation of image by 30 degrees \n with reflect mode')
plt.subplot(165)
aug = iaa.Affine(rotate=30, fit_output=True, cval=0, mode='wrap')
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.title('Rotation of image by \n30 degrees with wrap mode')
# + id="xguA31pNMfs4" outputId="ed34fbc8-629b-4d3e-b7e3-15f57367ba09" colab={"base_uri": "https://localhost:8080/", "height": 259}
plt.figure(figsize=(20,20))
plt.subplot(151)
aug = iaa.Affine(rotate=(-45,45), fit_output=True, cval=0, mode='constant')
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.subplot(152)
aug = iaa.Affine(rotate=(-45,45), fit_output=True, cval=0, mode='constant')
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.subplot(153)
aug = iaa.Affine(rotate=(-45,45), fit_output=True, cval=0, mode='constant')
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
plt.subplot(154)
aug = iaa.Affine(rotate=(-45,45), fit_output=True, cval=0, mode='constant')
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
# + id="e2fD-_dIPv7j"
# + id="9cFKjesAPv-y" outputId="3be2729d-a6a8-4e55-c1a3-517cf03e378c" colab={"base_uri": "https://localhost:8080/", "height": 282}
aug = iaa.Multiply(1)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray')
# + id="Jfyc8e13Pv5Q" outputId="3fc6cf7b-d793-4778-b108-2e2899661112" colab={"base_uri": "https://localhost:8080/", "height": 298}
aug = iaa.Multiply(0.5)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray',vmin = 0, vmax = 255)
plt.title('Pixels multiplied by 0.5')
# + id="wfBqBp6uMfqj" outputId="34502e00-0ab8-464b-87c2-0649f5a47c44" colab={"base_uri": "https://localhost:8080/", "height": 298}
aug = iaa.LinearContrast(0.5)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray',vmin = 0, vmax = 255)
plt.title('Pixel contrast by 0.5')
# + id="zqBbV3Zyk0Eb" outputId="d3491269-9924-4ba9-b377-083081226ab1" colab={"base_uri": "https://localhost:8080/", "height": 298}
aug = iaa.Dropout(p=0.2)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray',vmin = 0, vmax = 255)
plt.title('Random 20% pixel dropout')
# + id="E4y_n9Zjk0Cg" outputId="cc1bd40a-583e-4f00-ad39-0b6f8780f9bb" colab={"base_uri": "https://localhost:8080/", "height": 298}
aug = iaa.SaltAndPepper(0.2)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray',vmin = 0, vmax = 255)
plt.title('Random 20% salt and pepper noise')
# + id="m-y03oXDngcW" outputId="302719a8-cd41-4222-a959-54e529f42d9c" colab={"base_uri": "https://localhost:8080/", "height": 335}
plt.figure(figsize=(10,10))
plt.subplot(121)
aug = iaa.Dropout(p=0.2,)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray',vmin = 0, vmax = 255)
plt.title('Random 20% pixel dropout')
plt.subplot(122)
aug = iaa.SaltAndPepper(0.2,)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray',vmin = 0, vmax = 255)
plt.title('Random 20% salt and pepper noise')
# + id="khA0JNibph3U"
seq = iaa.Sequential([
iaa.Dropout(p=0.2,),
iaa.Affine(rotate=(-30,30))], random_order= True)
# + id="OzAZsENWph8X" outputId="f8f61e91-ae74-4356-c817-4071ba955290" colab={"base_uri": "https://localhost:8080/", "height": 314}
plt.imshow(seq.augment_image(tr_images[0]), cmap='gray',vmin = 0, vmax = 255)
plt.title('Image augmented using a \nrandom orderof the two augmentations')
# + id="2Jsk4U1clQFh" outputId="336afc19-91e4-466a-8e2d-7d0958da2de5" colab={"base_uri": "https://localhost:8080/", "height": 314}
aug = iaa.GaussianBlur(sigma=1)
plt.imshow(aug.augment_image(tr_images[0]), cmap='gray',vmin = 0, vmax = 255)
plt.title('Gaussian blurring of image\n with a sigma of 1')
| Chapter04/Image_augmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# This example shows how to interpret anomalies in *tabular* data.
#
# To verify the effectiveness of DeepAID, we use a synthetic dataset here.
# # Generate a synthetic dataset
# +
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
from sklearn.datasets import make_blobs
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
import random
# Generate Gaussian blobs (single cluster) consists 5050 100-dimension samples
X, _ = make_blobs(n_samples=5050, centers=1, n_features=100,
random_state=0, cluster_std=2.)
print('X.shape:',X.shape)
# first 5000 samples are used for training
X_train = X[:-50,:]
# generate anomalies by randomly perturbing 10 dimensions (1%) in the last 50 samples
random.seed(0)
noise_idx = random.choices(list(range(100)),k=10)
noise_idx.sort()
print('perturb index:',noise_idx)
noise_data = np.random.uniform(-20,-20,(50,10))
X_anomaly = X[-50:,:]
X_anomaly[:,noise_idx] += noise_data
# visualize anomaly and normal training data in 2D space
X_train_plot = X_train[:, noise_idx]
pca = PCA(n_components=2).fit(X_train_plot)
plt.scatter(X_train_plot[:, 0], X_train_plot[:, 1], alpha=0.5, s=3, label="training data (normal)")
X_anomaly_plot = X_anomaly[:,noise_idx]
pca = PCA(n_components=2).fit(X_anomaly_plot)
plt.scatter(X_anomaly_plot[:, 0], X_anomaly_plot[:, 1], alpha=1., s=4, c="r", label="anomaly")
plt.legend()
plt.show()
# -
# # Prepare a DL model for anomaly detection
# Train an autoencoder-based DL model
import numpy as np
import torch
from autoencoder import train, test, test_plot
import sys
sys.path.append('../../deepaid/')
from utils import validate_by_rmse, Normalizer
normer = Normalizer(X_train.shape[-1],online_minmax=False)
X_train = normer.fit_transform(X_train)
model, thres = train(X_train, X_train.shape[-1])
torch.save({'net':model,'thres':thres},'./save/autoencoder.pth.tar')
# Validate the performance of trained model
X_anomaly_norm = normer.transform(X_anomaly)
rmse_vec = test(model,thres,X_anomaly_norm)
test_plot(X_anomaly_norm, rmse_vec, thres) # ACC = 100%
# # Interpret the generated anomalies
# +
"""Load the model"""
from autoencoder import autoencoder
from utils import Normalizer
model_dict = torch.load('save/autoencoder.pth.tar')
model = model_dict['net']
thres = model_dict['thres']
""" Create a DeepAID Tabular Interpreter"""
import sys
sys.path.append("../../deepaid/interpreters/")
from tabular import TabularAID
my_interpreter = TabularAID(model,thres,input_size=100,k=10,steps=100,auto_params=False)
"""Interpret the anomalies"""
# for anomaly in X_anomaly:
anomaly = X_anomaly[5]
interpretation = my_interpreter(anomaly)
my_interpreter.show_table(anomaly,interpretation, normer)
print('perturb index:',noise_idx)
# -
# **As we can see that, the Interpreter successfully finds all perturbed dimensions**
| demos/tabular_synthesis/tabular_example_synthesis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import division
import os
from PIL import Image
import numpy as np
PREDICT_DIR = '../output/cihp_instance_part_maps'
INST_PART_GT_DIR = './Instance_part_val'
CLASSES = ['background', 'hat', 'hair', 'glove', 'sunglasses', 'upperclothes',
'dress', 'coat', 'socks', 'pants', 'tosor-skin', 'scarf', 'skirt',
'face', 'leftArm', 'rightArm', 'leftLeg', 'rightLeg', 'leftShoe', 'rightShoe']
IOU_THRE = [0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9]
#IOU_THRE = [0.1]
# compute mask overlap
def compute_mask_iou(mask_gt, masks_pre, mask_gt_area, masks_pre_area):
"""Calculates IoU of the given box with the array of the given boxes.
mask_gt: [H,W] # a mask of gt
masks_pre: [num_instances, height, width] predict Instance masks
mask_gt_area: the gt_mask_area , int
masks_pre_area: array of length masks_count. including all predicted mask, sum
Note: the areas are passed in rather than calculated here for
efficency. Calculate once in the caller to avoid duplicate work.
"""
intersection = np.logical_and(mask_gt, masks_pre)
intersection = np.where(intersection == True, 1, 0).astype(np.uint8)
intersection = NonZero(intersection)
# print('intersection:', intersection)
mask_gt_areas = np.full(len(masks_pre_area), mask_gt_area)
union = mask_gt_areas + masks_pre_area[:] - intersection[:]
iou = intersection / union
return iou
# compute the number of nonzero in mask
def NonZero(masks):
"""
:param masks: [N,h,w] a three-dimension array, includes N two-dimension mask arrays
:return: (N) return a tuple with length N. N is the number of non-zero elements in the two-dimension mask
"""
area = []
# print('NonZero masks',masks.shape)
for i in masks:
_, a = np.nonzero(i)
area.append(a.shape[0])
area = tuple(area)
return area
def compute_mask_overlaps(masks_pre, masks_gt):
"""Computes IoU overlaps between two sets of boxes.
masks_pre, masks_gt:
masks_pre [num_instances,height, width] Instance masks
masks_gt ground truth
For better performance, pass the largest set first and the smaller second.
"""
# Areas of masks_pre and masks_gt , get the nubmer of the non-zero element
area1 = NonZero(masks_pre)
area2 = NonZero(masks_gt)
# print(masks_pre.shape, masks_gt.shape)(1, 375, 1) (500, 375, 1)
# Compute overlaps to generate matrix [masks count, masks_gt count]
# Each cell contains the IoU value.
# print('area1',len(area1))
# print('area1',len(area2))
overlaps = np.zeros((masks_pre.shape[0], masks_gt.shape[0]))
# print('overlaps:',overlaps.shape)
for i in range(overlaps.shape[1]):
mask_gt = masks_gt[i]
# print('overlaps:',overlaps)
overlaps[:, i] = compute_mask_iou(mask_gt, masks_pre, area2[i], area1)
return overlaps
def voc_ap(rec, prec, use_07_metric=False):
"""
Compute VOC AP given precision and recall. If use_07_metric is true, uses the
VOC 07 11 point method (default:False).
Args:
rec: recall
prec: precision
use_07_metric:
Returns:
ap: average precision
"""
if use_07_metric:
# 11 point metric
ap = 0.
# arange([start, ]stop, [step, ]dtype=None)
for t in np.arange(0., 1.1, 0.1):
if np.sum(rec >= t) == 0:
p = 0
else:
p = np.max(prec[rec >= t])
ap += p / 11.
else:
# correct AP calculation
# first append sentinel values at the end
mrec = np.concatenate(([0.], rec, [1.]))
mpre = np.concatenate(([0.], prec, [0.]))
# compute the precision envelope
for i in range(mpre.size - 1, 0, -1):
mpre[i - 1] = np.maximum(mpre[i - 1], mpre[i])
# to calculate area under PR curve, look for points
# where X axis (recall) changes value
i = np.where(mrec[1:] != mrec[:-1])[0]
# and sum (\Delta recall) * prec
ap = np.sum((mrec[i + 1] - mrec[i]) * mpre[i + 1])
return ap
def convert2evalformat(inst_id_map, id_to_convert=None):
"""
param:
inst_id_map:[h, w]
id_to_convert: a set
return:
masks:[instances,h, w]
"""
masks = []
inst_ids = np.unique(inst_id_map)
# print("inst_ids:", inst_ids)
background_ind = np.where(inst_ids == 0)[0]
inst_ids = np.delete(inst_ids, background_ind)
if id_to_convert == None:
for i in inst_ids:
im_mask = (inst_id_map == i).astype(np.uint8)
masks.append(im_mask)
else:
for i in inst_ids:
if i not in id_to_convert:
continue
im_mask = (inst_id_map == i).astype(np.uint8)
masks.append(im_mask)
return masks, len(masks)
def compute_class_ap(image_id_list, class_id, iou_threshold):
"""Compute Average Precision at a set IoU threshold (default 0.5).
Input:
image_id_list : all pictures id list
gt_masks:all mask [N_pictures,num_inst,H,W]
pre_masks:all predict masks [N_pictures,num_inst,H,W]
pred_scores:scores for every predicted mask [N_pre_mask]
pred_class_ids: the indices of all predicted masks
Returns:
AP: Average Precision of specific class
"""
iou_thre_num = len(iou_threshold)
ap = np.zeros((iou_thre_num,))
gt_mask_num = 0
pre_mask_num = 0
tp = []
fp = []
scores = []
for i in range(iou_thre_num):
tp.append([])
fp.append([])
print("process class", CLASSES[class_id], class_id)
for image_id in image_id_list:
# print (image_id)
inst_part_gt = Image.open(os.path.join(INST_PART_GT_DIR, '%s.png' % image_id))
inst_part_gt = np.array(inst_part_gt)
rfp = open(os.path.join(INST_PART_GT_DIR, '%s.txt' % image_id), 'r')
gt_part_id = []
for line in rfp.readlines():
line = line.strip().split(' ')
gt_part_id.append([int(line[0]), int(line[1])])
rfp.close()
pre_img = Image.open(os.path.join(PREDICT_DIR, '%s.png' % image_id))
pre_img = np.array(pre_img)
rfp = open(os.path.join(PREDICT_DIR, '%s.txt' % image_id), 'r')
items = [x.strip().split(' ') for x in rfp.readlines()]
rfp.close()
pre_id = []
pre_scores = []
for i in range(len(items)):
if int(items[i][0]) == class_id:
pre_id.append(i+1)
pre_scores.append(float(items[i][1]))
gt_id = []
for i in range(len(gt_part_id)):
if gt_part_id[i][1] == class_id:
gt_id.append(gt_part_id[i][0])
gt_mask, n_gt_inst = convert2evalformat(inst_part_gt, set(gt_id))
pre_mask, n_pre_inst = convert2evalformat(pre_img, set(pre_id))
gt_mask_num += n_gt_inst
pre_mask_num += n_pre_inst
if n_pre_inst == 0:
continue
scores += pre_scores
if n_gt_inst == 0:
for i in range(n_pre_inst):
for k in range(iou_thre_num):
fp[k].append(1)
tp[k].append(0)
continue
gt_mask = np.stack(gt_mask)
pre_mask = np.stack(pre_mask)
# Compute IoU overlaps [pred_masks, gt_makss]
overlaps = compute_mask_overlaps(pre_mask, gt_mask)
# print('overlaps.shape',overlaps.shape)
max_overlap_ind = np.argmax(overlaps, axis=1)
# l = len(overlaps[:,max_overlap_ind])
for i in np.arange(len(max_overlap_ind)):
max_iou = overlaps[i][max_overlap_ind[i]]
# print('max_iou :', max_iou)
for k in range(iou_thre_num):
if max_iou > iou_threshold[k]:
tp[k].append(1)
fp[k].append(0)
else:
tp[k].append(0)
fp[k].append(1)
ind = np.argsort(scores)[::-1]
for k in range(iou_thre_num):
m_tp = tp[k]
m_fp = fp[k]
m_tp = np.array(m_tp)
m_fp = np.array(m_fp)
m_tp = m_tp[ind]
m_fp = m_fp[ind]
m_tp = np.cumsum(m_tp)
m_fp = np.cumsum(m_fp)
# print('m_tp : ',m_tp)
# print('m_fp : ', m_fp)
recall = m_tp / float(gt_mask_num)
precition = m_tp / np.maximum(m_fp+m_tp, np.finfo(np.float64).eps)
# Compute mean AP over recall range
ap[k] = voc_ap(recall, precition, False)
return ap
if __name__ == '__main__':
print("result of", PREDICT_DIR)
image_id_list = [x[:-4] for x in os.listdir(PREDICT_DIR) if x[-3:] == 'txt']
AP = np.zeros((len(CLASSES)-1, len(IOU_THRE)))
for ind in range(1, len(CLASSES)):
AP[ind - 1, :] = compute_class_ap(image_id_list, ind, IOU_THRE)
print("-----------------AP-----------------")
print(AP)
print("-------------------------------------")
mAP = np.mean(AP, axis=0)
print("-----------------mAP-----------------")
print(mAP)
print(np.mean(mAP))
print("-------------------------------------")
# -
| evaluation/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Accessing satellite data at AWS from within Python
# This example notebook shows how to obtain Sentinel-2 imagery and additional data from [Amazon Web Services (AWS) bucket](http://sentinel-s2-l1c.s3-website.eu-central-1.amazonaws.com/). The data at AWS is the same as original S-2 data provided by ESA. It is organized by [ESA products](http://sentinel-s2-l1c.s3-website.eu-central-1.amazonaws.com/#products/) and [ESA tiles](http://sentinel-s2-l1c.s3-website.eu-central-1.amazonaws.com/#tiles/).
#
# The ```sentinelhub``` package therefore supports obtaining data by products and by tiles. It can download data either to the same file structure as it is at AWS or it can download data into original ```.SAFE``` file structure described by ESA.
# ## Imports
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from sentinelhub import AwsProductRequest, AwsTileRequest, AwsTile
# ## Aws Tile
# S-2 tile can be uniquely defined either with ESA tile ID (e.g. `L1C_T01WCV_A012011_20171010T003615`) or with tile name (e.g. `T38TML` or `38TML`), sensing time/date and AWS index. The AWS index is the last number in tile AWS url (e.g. http://sentinel-s2-l1c.s3-website.eu-central-1.amazonaws.com/#tiles/10/U/EV/2017/11/17/0/ → `0`).
#
# The package works with the second tile definition. To transform tile ID to `(tile_name, time, aws_index)` do the following:
tile_id = 'S2A_OPER_MSI_L1C_TL_MTI__20151219T100121_A002563_T38TML_N02.01'
tile_name, time, aws_index = AwsTile.tile_id_to_tile(tile_id)
tile_name, time, aws_index
# Next we can download the tile data. Let's download only bands `B8A` and `B10`. Let's also download meta data files `tileInfo.json`, `preview.jp2` and pre-calculated cloud mask `qi/MSK_CLOUDS_B00`. We will save everything into folder `./AwsData`
# +
bands = ['B8A', 'B10']
metafiles = ['tileInfo', 'preview', 'qi/MSK_CLOUDS_B00']
data_folder = './AwsData'
request = AwsTileRequest(tile=tile_name, time=time, aws_index=aws_index,
bands=bands, metafiles=metafiles, data_folder=data_folder)
request.save_data()
# -
# Note that upon calling this method again the data won't be re-downloaded unless we set the parameter `redownload=True`.
#
# To obtain downloaded data we can simply do:
data_list = request.get_data()
b8a, b10, tile_info, preview, cloud_mask = data_list
# Download and reading could also be done in a single call `request.get_data(save_data=True)`.
plt.imshow(preview)
plt.imshow(b8a)
# ## Aws Product
# S-2 product is uniquely defined by ESA product ID. We can obtain data for the whole product
# +
product_id = 'S2A_MSIL1C_20171010T003621_N0205_R002_T01WCV_20171010T003615'
request = AwsProductRequest(product_id=product_id, data_folder=data_folder)
data_list = request.get_data(save_data=True)
# -
# If `bands` parameter is not defined all bands will be downloaded. If `metafiles` parameter is not defined no additional metadata files will be downloaded.
# ## Data into .SAFE structure
# The data can also be downloaded into .SAFE structure by specifying `safe_format=True` in upper examples.
tile_request = AwsTileRequest(tile=tile_name, time=time, aws_index=aws_index,
bands=bands, metafiles=metafiles, data_folder=data_folder, safe_format=True)
tile_request.save_data()
# +
product_id = 'S2A_OPER_PRD_MSIL1C_PDMC_20160121T043931_R069_V20160103T171947_20160103T171947'
product_request = AwsProductRequest(product_id=product_id, bands=['B01'], data_folder=data_folder, safe_format=True)
product_request.save_data()
# -
# Older products contain multiple tiles. In case would like to download only some tiles it is also possible to specify a list of tiles to download.
product_request = AwsProductRequest(product_id=product_id, tile_list = ['T14PNA', 'T13PHT'], data_folder=data_folder, safe_format=True)
product_request.save_data()
# ## Searching for data available at AWS
#
# The archive can be manually searched using either [Sentinel L1C webpage](http://sentinel-s2-l1c.s3-website.eu-central-1.amazonaws.com/) in case of L1C data or with [aws_cli](https://docs.aws.amazon.com/cli/latest/reference/s3/ls.html) in case of L2A data. E.g.:
#
# ```bash
# aws s3 ls s3://sentinel-s2-l2a/tiles/33/U/WR/ --request-payer
# ```
#
# The archive can also be searched automatically and according to specified area and time interval using Sentinel Hub Web Feature Service (WFS):
# +
from sentinelhub import WebFeatureService, BBox, CRS, DataSource
INSTANCE_ID = '' # In case you put instance ID into cofniguration file you can leave this unchanged
search_bbox = BBox(bbox=[46.16, -16.15, 46.51, -15.58], crs=CRS.WGS84)
search_time_interval = ('2017-12-01', '2017-12-15')
wfs_iterator = WebFeatureService(search_bbox, search_time_interval,
data_source=DataSource.SENTINEL2_L1C,
maxcc=1.0, instance_id=INSTANCE_ID)
for tile_info in wfs_iterator:
print(tile_info)
# -
# From obtained WFS iterator we can extract info which uniquely defines each tile.
wfs_iterator.get_tiles()
| examples/aws_request.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# We will apply the modelling script to draft model protein CCNI (accession code Q14094)
# +
from Bio.Blast import NCBIWWW
from Bio import SeqIO
from io import StringIO
import requests
idname='Q14094'
# retrieve info
baseUrl="http://www.uniprot.org/uniprot/"
currentUrl=baseUrl+idname+".fasta"
response = requests.post(currentUrl)
cData=''.join(response.text)
Seq=StringIO(cData)
# -
# print in FASTA format
sequery=""
for record in SeqIO.parse(Seq, "fasta"):
print(">"+record.description)
print(record.seq)
print("Found match for "+idname)
filename="files/"+idname+"_blast.xml"
print("Calling BLAST against the PDB, and saving the XML file "+filename)
result_handle=NCBIWWW.qblast("blastp", "pdb", record.seq, hitlist_size=100)
a=str(record.seq[0:4])
blastXMLfh = open(filename, "w")
blastXMLfh.write(result_handle.read())
blastXMLfh.close()
result_handle.close()
print("Done!")
sequery = str(record.seq)
# +
from Bio.Blast import NCBIXML
import urllib
result_handle = open("files/"+idname+"_blast.xml")
blast_records=list(NCBIXML.parse(result_handle)) #putting the results into a list is convenient
#to do some extra work with them
fileh = open ("files/seq.fasta","w")
fileh.write(">"+idname+"\n")
fileh.write(sequery+"\n")
E_VALUE_THRESH = 1e-9
listofknowns = []
for blast_record in blast_records:
for alignment in blast_record.alignments:
for hsp in alignment.hsps:
if hsp.expect < E_VALUE_THRESH:
print("\nALIGNMENT\n=========\n")
print(alignment.title)
fileh.write(">"+alignment.title+"\n")
print("E value:",hsp.expect)
print(hsp.query[0:75]+"...")
print(hsp.match[0:75]+"...")
print(hsp.sbjct[0:75]+"...")
fileh.write(hsp.sbjct+"\n")
print("found good match: ",alignment.hit_id)
PDBcode = alignment.hit_id.split('|')[1]
PDBfile = 'data/'+PDBcode+'.pdb'
print("retrieving PDB file: ",PDBcode)
urllib.request.urlretrieve('http://files.rcsb.org/download/'+PDBcode+'.pdb', PDBfile)
listofknowns.append(PDBfile)
fileh.close()
print(listofknowns)
# +
from Bio import AlignIO
from Bio.Align.Applications import ClustalOmegaCommandline
# use these lines to ensure clustalo can be found. This works for my installation using
# the above conda instructions
import os
os.environ['PATH'] += ':~/miniconda3/bin/'
# this is an example using the complete collection of sequences. You should try it with your collection
# obtained from the BALST calculation
file='files/seq.fasta'
outfile='files/seq_aligned.fasta'
cline = ClustalOmegaCommandline(infile= file, outfile= outfile, verbose=True, auto=True, force=True)
stdout, stderr = cline()
# -
print(stdout)
# you can check what you created:
# !cat files/seq_aligned.fasta
# Once you have the list of PDB files and the alignment you are ready to use MODELLER.
# Check the [SaliLab web site](https://salilab.org) for registration to the program.
# ```
# conda config --add channels salilab
# conda install modeller
# ```
# In the same site you can find examples of use.
#
# QUESTION: so, did you get how to obtain your models with MODELLER?
#
from modeller import *
from modeller.automodel import *
#from modeller import d
env = environ()
a = automodel(env, alnfile='files/seq_aligned.fasta',
knowns=listofknowns, sequence='qseq1',
assess_methods=(assess.DOPE,
#soap_protein_od.Scorer(),
assess.GA341))
a.starting_model = 1
a.ending_model = 5
a.make()
# Now you are ready to check the quality of your models with [SAVES](https://servicesn.mbi.ucla.edu/SAVES/)
| .ipynb_checkpoints/HomologyModellingCCNI-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # T81-558: Applications of Deep Neural Networks
# **Class 5: Backpropagation.**
# * Instructor: [<NAME>](https://sites.wustl.edu/jeffheaton/), School of Engineering and Applied Science, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
# * For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# # Common Functions from Before
# +
from sklearn import preprocessing
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from sklearn.cross_validation import train_test_split
# Encode text values to dummy variables(i.e. [1,0,0],[0,1,0],[0,0,1] for red,green,blue)
def encode_text_dummy(df,name):
dummies = pd.get_dummies(df[name])
for x in dummies.columns:
dummy_name = "{}-{}".format(name,x)
df[dummy_name] = dummies[x]
df.drop(name, axis=1, inplace=True)
# Encode text values to indexes(i.e. [1],[2],[3] for red,green,blue).
def encode_text_index(df,name):
le = preprocessing.LabelEncoder()
df[name] = le.fit_transform(df[name])
return le.classes_
# Encode a numeric column as zscores
def encode_numeric_zscore(df,name,mean=None,sd=None):
if mean is None:
mean = df[name].mean()
if sd is None:
sd = df[name].std()
df[name] = (df[name]-mean)/sd
# Convert all missing values in the specified column to the median
def missing_median(df, name):
med = df[name].median()
df[name] = df[name].fillna(med)
# Convert a Pandas dataframe to the x,y inputs that TensorFlow needs
def to_xy(df,target):
result = []
for x in df.columns:
if x != target:
result.append(x)
# find out the type of the target column. Is it really this hard? :(
target_type = df[target].dtypes
target_type = target_type[0] if hasattr(target_type, '__iter__') else target_type
# Encode to int for classification, float otherwise. TensorFlow likes 32 bits.
if target_type in (np.int64, np.int32):
# Classification
return df.as_matrix(result).astype(np.float32),df.as_matrix([target]).astype(np.int32)
else:
# Regression
return df.as_matrix(result).astype(np.float32),df.as_matrix([target]).astype(np.float32)
# Nicely formatted time string
def hms_string(sec_elapsed):
h = int(sec_elapsed / (60 * 60))
m = int((sec_elapsed % (60 * 60)) / 60)
s = sec_elapsed % 60
return "{}:{:>02}:{:>05.2f}".format(h, m, s)
# +
# %matplotlib inline
import matplotlib.pyplot as plt
from sklearn.metrics import roc_curve, auc
# Plot a confusion matrix.
# cm is the confusion matrix, names are the names of the classes.
def plot_confusion_matrix(cm, names, title='Confusion matrix', cmap=plt.cm.Blues):
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(names))
plt.xticks(tick_marks, names, rotation=45)
plt.yticks(tick_marks, names)
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# Plot an ROC. pred - the predictions, y - the expected output.
def plot_roc(pred,y):
fpr, tpr, _ = roc_curve(y_test, pred)
roc_auc = auc(fpr, tpr)
plt.figure()
plt.plot(fpr, tpr, label='ROC curve (area = %0.2f)' % roc_auc)
plt.plot([0, 1], [0, 1], 'k--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver Operating Characteristic (ROC)')
plt.legend(loc="lower right")
plt.show()
# Plot a lift curve. pred - the predictions, y - the expected output.
def chart_regression(pred,y):
t = pd.DataFrame({'pred' : pred.flatten(), 'y' : y_test.flatten()})
t.sort_values(by=['y'],inplace=True)
plt.plot(t['y'].tolist(),label='expected')
plt.plot(t['pred'].tolist(),label='prediction')
plt.ylabel('output')
plt.legend()
plt.show()
# -
# # Classic Backpropagation
# Backpropagation is the primary means by which a neural network's weights are determined during training. Backpropagation works by calculating a weight change amount ($v_t$) for every weight($\theta$) in the neural network. This value is subtracted from every weight by the following equation:
#
# $ \theta_t = \theta_{t-1} - v_t $
#
# This process is repeated for every iteration($t$). How the weight change is calculated depends on the training algorithm. Classic backpropagation simply calculates a gradient ($\nabla$) for every weight in the neural network with respect to the error function ($J$) of the neural network. The gradient is scaled by a learning rate ($\eta$).
#
# $ v_t = \eta \nabla_{\theta_{t-1}} J(\theta_{t-1}) $
#
# The learning rate is an important concept for backpropagation training. Setting the learning rate can be complex:
#
# * Too low of a learning rate will usually converge to a good solution; however, the process will be very slow.
# * Too high of a learning rate will either fail outright, or converge to a higher error than a better learning rate.
#
# Common values for learning rate are: 0.1, 0.01, 0.001, etc.
#
# Gradients:
#
# 
#
# The following link, from the book, shows how a simple [neural network is trained with backpropagation](http://www.heatonresearch.com/aifh/vol3/).
# # Momentum Backpropagation
#
# Momentum adds another term to the calculation of $v_t$:
#
# $ v_t = \eta \nabla_{\theta_{t-1}} J(\theta_{t-1}) + \lambda v_{t-1} $
#
# Like the learning rate, momentum adds another training parameter that scales the effect of momentum. Momentum backpropagation has two training parameters: learning rate ($\lambda$) and momentum ($\eta$). Momentum simply adds the scaled value of the previous weight change amount ($v_{t-1}$) to the current weight change amount($v_t$).
#
# This has the effect of adding additional force behind a direction a weight was moving. This might allow the weight to escape a local minima:
#
# 
#
# A very common value for momentum is 0.9.
#
# # Batch and Online Backpropagation
#
# How often should the weights of a neural network be updated? Gradients can be calculated for a training set element. These gradients can also be summed together into batches and the weights updated once per batch.
#
# * **Online Training** - Update the weights based on gradients calculated from a single training set element.
# * **Batch Training** - Update the weights based on the sum of the gradients over all training set elements.
# * **Batch Size** - Update the weights based on the sum of some batch size of training set elements.
# * **Mini-Batch Training** - The same as batch size, but with a very small batch size. Mini-batches are very popular and they are often in the 32-64 element range.
#
# Because the batch size is smaller than the complete training set size, it may take several batches to make it completely through the training set. You may have noticed TensorFlow reporting both steps and epochs when a neural network is trained:
#
# ```
# Step #100, epoch #7, avg. train loss: 23.02969
# Step #200, epoch #15, avg. train loss: 2.67576
# Step #300, epoch #23, avg. train loss: 1.33839
# Step #400, epoch #30, avg. train loss: 0.86830
# Step #500, epoch #38, avg. train loss: 0.67166
# Step #600, epoch #46, avg. train loss: 0.54569
# Step #700, epoch #53, avg. train loss: 0.47544
# Step #800, epoch #61, avg. train loss: 0.39358
# Step #900, epoch #69, avg. train loss: 0.36052
# ```
#
# * **Step/Iteration** - The number of batches that were processed.
# * **Epoch** - The number of times the complete training set was processed.
#
# # Stochastic Gradient Descent
#
# Stochastic gradient descent (SGD) is currently one of the most popular neural network training algorithms. It works very similarly to Batch/Mini-Batch training, except that the batches are made up of a random set of training elements.
#
# This leads to a very irregular convergence in error during training:
#
# 
# [Image from Wikipedia](https://en.wikipedia.org/wiki/Stochastic_gradient_descent)
#
# Because the neural network is trained on a random sample of the complete training set each time, the error does not make a smooth transition downward. However, the error usually does go down.
#
# Advantages to SGD include:
#
# * Computationally efficient. Even with a very large training set, each training step can be relatively fast.
# * Decreases overfitting by focusing on only a portion of the training set each step.
#
# # Other Techniques
#
# One problem with simple backpropagation training algorithms is that they are highly sensative to learning rate and momentum. This is difficult because:
#
# * Learning rate must be adjusted to a small enough level to train an accurate neural network.
# * Momentum must be large enough to overcome local minima, yet small enough to not destabilize the training.
# * A single learning rate/momentum is often not good enough for the entire training process. It is often useful to automatically decrease learning rate as the training progresses.
# * All weights share a single learning rate/momentum.
#
# Other training techniques:
#
# * **Resilient Propagation** - Use only the magnitude of the gradient and allow each neuron to learn at its own rate. No need for learning rate/momentum; however, only works in full batch mode.
# * **Nesterov accelerated gradient** - Helps midigate the risk of choosing a bad mini-batch.
# * **Adagrad** - Allows an automatically decaying per-weight learning rate and momentum concept.
# * **Adadelta** - Extension of Adagrad that seeks to reduce its aggressive, monotonically decreasing learning rate.
# * **Non-Gradient Methods** - Non-gradient methods can *sometimes* be useful, though rarely outperform gradient-based backpropagation methods. These include: [simulated annealing](https://en.wikipedia.org/wiki/Simulated_annealing), [genetic algorithms](https://en.wikipedia.org/wiki/Genetic_algorithm), [particle swarm optimization](https://en.wikipedia.org/wiki/Particle_swarm_optimization), [Nelder Mead](https://en.wikipedia.org/wiki/Nelder%E2%80%93Mead_method), and [many more](https://en.wikipedia.org/wiki/Category:Optimization_algorithms_and_methods).
# # ADAM Update
#
# ADAM is the first training algorithm you should try. It is very effective. Kingma and Ba (2014) introduced the Adam update rule that derives its name from the adaptive moment estimates that it uses. Adam estimates the first (mean) and second (variance) moments to determine the weight corrections. Adam begins with an exponentially decaying average of past gradients (m):
#
# $ m_t = \beta_1 m_{t-1} + (1-\beta_1) g_t $
#
# This average accomplishes a similar goal as classic momentum update; however, its value is calculated automatically based on the current gradient ($g_t$). The update rule then calculates the second moment ($v_t$):
#
# $ v_t = \beta_2 v_{t-1} + (1-\beta_2) g_t^2 $
#
# The values $m_t$ and $v_t$ are estimates of the first moment (the mean) and the second moment (the uncentered variance) of the gradients respectively. However, they will have a strong bias towards zero in the initial training cycles. The first moment’s bias is corrected as follows.
#
# $ \hat{m}_t = \frac{m_t}{1-\beta^t_1} $
#
# Similarly, the second moment is also corrected:
#
# $ \hat{v}_t = \frac{v_t}{1-\beta_2^t} $
#
# These bias-corrected first and second moment estimates are applied to the ultimate Adam update rule, as follows:
#
# $ \theta_t = \theta_{t-1} - \frac{\eta}{\sqrt{\hat{v}_t}+\eta} \hat{m}_t $
#
# Adam is very tolerant to initial learning rate (η) and other training parameters. Kingma and Ba (2014) propose default values of 0.9 for $\beta_1$, 0.999 for $\beta_2$, and 10-8 for $\eta$.
# # Methods Compared
#
# The following image shows how each of these algorithms train (image credits: [author](<NAME>), [where I found it](http://sebastianruder.com/optimizing-gradient-descent/index.html#visualizationofalgorithms) ):
#
# 
#
#
# # Specifying the Update Rule in Tensorflow
#
# TensorFlow allows the update rule to be set to one of:
#
# * Adagrad
# * **Adam**
# * Ftrl
# * Momentum
# * RMSProp
# * **SGD**
#
#
# +
# %matplotlib inline
from matplotlib.pyplot import figure, show
from numpy import arange
import tensorflow.contrib.learn as skflow
import pandas as pd
import os
import numpy as np
import tensorflow as tf
from sklearn import metrics
from scipy.stats import zscore
import matplotlib.pyplot as plt
path = "./data/"
filename_read = os.path.join(path,"auto-mpg.csv")
df = pd.read_csv(filename_read,na_values=['NA','?'])
# create feature vector
missing_median(df, 'horsepower')
df.drop('name',1,inplace=True)
encode_numeric_zscore(df, 'horsepower')
encode_numeric_zscore(df, 'weight')
encode_numeric_zscore(df, 'cylinders')
encode_numeric_zscore(df, 'displacement')
encode_numeric_zscore(df, 'acceleration')
encode_text_dummy(df, 'origin')
# Encode to a 2D matrix for training
x,y = to_xy(df,['mpg'])
# Split into train/test
x_train, x_test, y_train, y_test = train_test_split(
x, y, test_size=0.25, random_state=42)
# Create a deep neural network with 3 hidden layers of 50, 25, 10
regressor = skflow.TensorFlowDNNRegressor(
hidden_units=[50, 25, 10],
batch_size = 32,
#momentum=0.9,
optimizer='SGD',
learning_rate=0.01,
steps=5000)
# Early stopping
early_stop = skflow.monitors.ValidationMonitor(x_test, y_test,
early_stopping_rounds=200, print_steps=50)
# Fit/train neural network
regressor.fit(x_train, y_train, monitor=early_stop)
# Measure RMSE error. RMSE is common for regression.
pred = regressor.predict(x_test)
score = np.sqrt(metrics.mean_squared_error(pred,y_test))
print("Final score (RMSE): {}".format(score))
# Plot the chart
chart_regression(pred,y_test)
# -
| t81_558_class5_backpropagation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyDeepLearning-1.0
# language: python
# name: pydeeplearning
# ---
# <img src=attachment:88c90614-07ea-4ccb-aab3-7b63dc27a603.png width=400, height=41>
#
# Author: [<NAME>](mailto:<EMAIL>)
# # Python Face Blurer
#
# This Script will scan any images in the *Images*-folder or any videos in the *Video*-folder for faces/persons using the [Faster RCNN Open Images V4 Neural Network](https://tfhub.dev/google/faster_rcnn/openimages_v4/inception_resnet_v2/1) and apply a blur to all findings.
#
#
#
# #### Table of Contents:
# 1. Initializing the Project
# - Creating the Folder Structure
# - Imports
# - Configuration
# 2. Initializing the Blur
# - Functions for bluring objects
# - Functions for drawing boxes
# 3. Loading the Classifier
# - Pulling the Classifier
# - Input and Output Parsing
# 4. Fetching all Images and Video
# 5. Run Classifier and Apply Blur
# - Run Classifier over every Image and Video
# - Save to Output folder
# - Compact Frames into a Video
# 6. Cleanup
#
# -----------------------------------------------------
# ## Initializing the Project
# ### Creating the Folder Structure
#
# For Python to find the Images and Videos it needs the path to the folders where the Images are stored. By default the notebook expects the following folder structure, but you can change the paths below if you want:
# <pre>
# ├── face_blurer.ipynb
# ├── Images
# │ ├── Image.png
# │ └── ...
# ├── Images_Out
# ├── Videos
# │ ├── Video.mp4
# │ └── ...
# └── Videos_Out
# </pre>
#
# Usage:
# - **Images**: Put all Images here that you want to have blured. OpenCV can load almost all file formats, but for a complete list look [here](https://docs.opencv.org/3.4/d4/da8/group__imgcodecs.html#ga288b8b3da0892bd651fce07b3bbd3a56).
# - **Images_Out**: After processing the new Images will be placed here. The name of the processed file can be specified in the config with *FILENAME_AFTER_BLURRING*.
# - **Videos**: Same deal as with the Images. Put all Videos here that you want to have processed.
# - **Videos_Out**: All processed Videos will be placed here after applying the algorithm. The new name will be the same as for the Images.
#
# > Note: If you want to load the files from a different place in storage you can do so by editing the variable in the configuration. More on that below!
# ### Imports
#
# #### *Please use the PyDeepLearning*-kernel or the kernel from **kernel.ipynb** to run this
#
# Import everything we need for the whole operation.
# - The main package for the model will be tensorflow, which will internally use [TensorflowHub](https://tfhub.dev/) to download the network.
# - We will apply the blur using PIL, the Python Image Library that nativly gets delivered with python.
# - matplotlib will be used to display images in the notebook. Usefull if you want to see the results of the blurring directly.
# +
#######################################
# Disable GPU Usage for TensorFlow
#######################################
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '-1'
# Main package for the ANN
import tensorflow as tf
import tensorflow_hub as hub
# Opening Files and Bluring
import cv2 as cv
import numpy as np
# Displaying the image
import matplotlib.pyplot as plt
from time import time # For measuring the inference time.
from os import listdir # Getting available files
import re # Config
import tempfile
import shutil
print("tensorflow version: " + tf.__version__)
print("OpenCV version: " + cv.__version__)
# Check available GPU devices.
print("\nNum GPUs Available: ", len(tf.config.experimental.list_physical_devices("GPU")))
# -
# ### Configuration
#
# Here you can configure every aspect of the algorithm
#
# The most interesting option is probably *FILTER_OBJETCS*, where you can specify what object should be detected/blurred. You can see all available object classes in [this image](https://storage.googleapis.com/openimages/2018_04/bbox_labels_600_hierarchy_visualizer/circle.html) or take a direct look at the [Open Image Dataset](https://storage.googleapis.com/openimages/web/visualizer/index.html?). The filter is interpreted as a Regex, so you can also filter for multiple classes, eg. by chaining them together with `|` or using wildcard symbols like `.`. In most cases you'll probably want to use `Person` or `Human face`
# +
FILTER_OBJECTS = "Person" # Regex. What Object you want to detect. Everything else will be ignored.
FILENAME_AFTER_BLURRING = "{}_blured" # {} replaces the original filename. Don't add the extension (.jpg, .mp3) here
DRAW_BOXES = False
BLUR_OBJECTS = True
BLUR_INTENSITY = 20
EXTEND_BOXES = 20 # Increases the size of the boxes by x pixels in each direction
FORCE_IMAGE_SIZE = (None, None) # Leave at None for no changes. You are free to only resize one Dimension and leave the other
FORCE_VIDEO_SIZE = (None, None) # Same as above
CROP_VIDEO_LENGTH = None # Time in seconds after which videos will be cropped. Recommended for testing
VIDEO_CACHE_FOLDER = None # Before we output the final video all frames will be saved as a .png-file in a folder. If you specify a folder here they won't be deleted afterwards. Leave None to use a tempdir.
FORCE_VIDEO_FORMAT = None # Leave None for Original. You must not include the preceding "."
VIDEO_BITRATE = None # String or Integer. Leave None for default
ENCODER = "h264" # E.g. "h264", "hevc"(h265), "asv1" (AV1)
# When paths are relative the origin will be the folder where this notebook is. You MUST put an / behind the folder paths
IMAGE_INPUT_FOLDER = "./Images/"
IMAGE_OUTPUT_FOLDER = "./Images_Out/"
VIDEO_INPUT_FOLDER = "./Videos/"
VIDEO_OUTPUT_FOLDER = "./Videos_Out/"
# -
# ----------------------
# ## Initializing the Blur
# ### Functions for bluring objects
# +
def blur_objects(image, boxes, names, confidence, min_score=0.1):
"""
Apply the results from the network to the image and blur the objects specified above.
From the network return values we need to pass where the objects are, what each object is and how confident the network if about the detection.
Since those properties are given in seperate lists, each identified object needs to be in the same index for every parameter.
:param image: (numpy.ndarray) The image to be drawn on.
:param boxes: (numpy.ndarray) An Array of shape (n, 4) with the 4 coordinates for each box. Cordinate must be [ymin, xmin, ymax, xmax]
:param names: (numpy.ndarray) An Array of shape (n) with the name of each Object.
:param confidence: (numpy.ndarray) An Array of shape (n) with the confidence for each Object.
:param min_score: (optional, int) The required confidence for the box to be applied
"""
for i in range(boxes.shape[0]):
is_confident = confidence[i] >= min_score
is_in_filter = re.search(FILTER_OBJECTS, names[i]) is not None
if is_confident and is_in_filter:
blur_object(image, boxes[i])
def blur_object(image, coordinates, intensity=BLUR_INTENSITY):
height, width = image.shape[:2]
(ymin, xmin, ymax, xmax) = coordinates
(x, w, y, h) = (int(xmin * width), int(xmax * width), int(ymin * height), int(ymax * height))
# Blur the area of the face
sub_image = image[y:h, x:w]
sub_image = cv.blur(sub_image, (intensity, intensity))
# Apply blurred face back to image
image[y:h, x:w] = sub_image
# -
# ### Functions for drawing boxes
# +
def draw_boxes(image, boxes, names, confidence, min_score=0.1):
"""
Apply the results from the network to the image and draw a box around each identified object.
From the network return values we need to pass where the boxes are, what object each box represents and confident the network if about the detection.
Since those properties are given in seperate lists, each identified object needs to be in the same index for every parameter.
:param image: (numpy.ndarray) The image to be drawn on.
:param boxes: (numpy.ndarray) An Array of shape (n, 4) with the 4 coordinates for each box. Cordinate must be [ymin, xmin, ymax, xmax]
:param names: (numpy.ndarray) An Array of shape (n) with the name of each Object.
:param confidence: (numpy.ndarray) An Array of shape (n) with the confidence for each Object.
:param min_score: (optional, int) The required confidence for the box to be applied
"""
for i in range(boxes.shape[0]):
is_confident = confidence[i] >= min_score
is_in_filter = re.search(FILTER_OBJECTS, names[i]) is not None
if is_confident and is_in_filter:
display_str = f"{names[i]}: {int(100 * confidence[i])}%"
draw_bounding_box_on_image(image, boxes[i], display_str)
def draw_bounding_box_on_image(image, coordinates, label="", color=(0, 0, 255), thickness=4):
"""
Adds a bounding box to an image.
:param image: (np.ndarray) The Image to be drawn on.
:param coordinates: (tuple) Coordinates of the Box: (ymin, xmin, ymax, xmax). Each coordinate must be between 0 and 1.
:param color: (optional, str) 7-digit String representing the Color in the format of '#rrggbb'.
:param thickness: (optional, int) How thick the box should be.
"""
# Draw the Box itself
height, width = image.shape[:2]
(left, top) = (int(coordinates[1] * width) - EXTEND_BOXES, int(coordinates[0] * height) - EXTEND_BOXES)
(width, height) = (int(coordinates[3] * width) - left + 2 * EXTEND_BOXES, int(coordinates[2] * height) - top + 2 * EXTEND_BOXES)
image = cv.rectangle(image, (left, top, width, height), color, thickness)
# Calculate text specs
(width, height), _ = cv.getTextSize(label, cv.FONT_HERSHEY_SIMPLEX, 0.6, 1)
height += 10
width += 10
position = top if top < height else top - height # Move label to the inside if not enough space above the box
# Draw Text
rect = (left, position, width, height)
image = cv.rectangle(image, rect, color, -1)
image = cv.putText(image, label, (left + 5, position + height - 5), cv.FONT_HERSHEY_SIMPLEX, 0.6, (255, 255, 255))
# -
# --------------
# ## Loading the Classifier
# ### Pulling the Classifier
#
# Here we use tensorflow_hub to Pull the ANN from the [Tensorflow Hub](https://tfhub.dev/google/faster_rcnn/openimages_v4/inception_resnet_v2/1). Be aware that depending on your setup the loading can take a few minutes.
module_handle = "./model"
classifier = hub.load(module_handle).signatures['default']
# # Input and Output Parsing
#
# These Functions will load an Image as an numpy.ndarray, which will then be passed to *run_detector*, were they will be converted to the right format and passed to the classifier
# +
def display_image(image):
"""Displays the image to the screen."""
plt.figure(figsize=(20, 15))
plt.grid(False)
plt.axis("off")
plt.imshow(image)
def open_image(path, display=False, resolution=FORCE_IMAGE_SIZE):
"""
Open and format an image from disk.
Open a picture from file in the path variable. After if gets resized and converted to RGB if neccesary. If chosen if will also be displayed.
:param path: (str) The path to the Picture.
:param display: (boolean, optional) Whether you want the image displayed
:param width: (tuple, optional) The desired resolution after rescaling.
"""
image = cv.imread(path, cv.IMREAD_COLOR)
if (resolution != (None, None)):
width = resolution[0] or image.shape[1]
height = resolution[1] or image.shape[0]
image = cv.resize(image, dsize=(width, height), interpolation=cv.INTER_CUBIC)
if display:
display_image(image)
return image
def run_detector(detector, image):
"""
Runs the given classifier on the given image. Only works with detectors that accept the same input format as the original one.
:param detector: (A trackable Object) The Classifier.
:param image: (np.ndarray) The Image.
:return: (Dict) A Dictionary with the objects found by the Classifier.
"""
img = image.astype(np.float32)
img = np.expand_dims(img, axis=0) # add required additional dimension. Same as img[np.newaxis, :], but better readable
img /= 255 # Normalize
result = detector(tf.convert_to_tensor(img))
# Extract the numeric values
result = {key: value.numpy() for key, value in result.items()}
result["detection_class_entities"] = [name.decode('ascii') for name in result["detection_class_entities"]]
return result
# -
# --------------
# ## Fetching all Images and Videos
# +
images = listdir(IMAGE_INPUT_FOLDER)
videos = listdir(VIDEO_INPUT_FOLDER)
# Remove hidden files (Starting with .) and convert to an tuple with (name, format)
images = [[file.split(".")[0], file.split(".")[-1]] for file in images if not file.startswith(".")]
videos = [[file.split(".")[0], file.split(".")[-1]] for file in videos if not file.startswith(".")]
print(f"Found {len(images)} Images")
print(f"Found {len(videos)} Videos")
# -
# ---------------------
#
# ## Run Classifier and Blur
# ### Run Classifier on every Image
for file in images:
image = open_image(IMAGE_INPUT_FOLDER + file[0] + "." + file[1])
start_time = time()
#######################################
# Code that failes
#######################################
result = run_detector(classifier, image)
if BLUR_OBJECTS:
blur_objects(image, result["detection_boxes"], result["detection_class_entities"], result["detection_scores"])
if DRAW_BOXES:
draw_boxes(image, result["detection_boxes"], result["detection_class_entities"], result["detection_scores"])
cv.imwrite(IMAGE_OUTPUT_FOLDER + FILENAME_AFTER_BLURRING.format(file[0]) + "." + file[1], image)
print(f"Name: {file[0]}\t Format: {file[1]}\t Size: {image.shape[1]}x{image.shape[0]}\t Time: {time() - start_time:.2f}s")
# ### Run Classifier on every Video
# +
cache_folder = VIDEO_CACHE_FOLDER if VIDEO_CACHE_FOLDER else tempfile.mkdtemp() + "/"
print(f"Using folder {cache_folder}")
for file in videos:
video = cv.VideoCapture(VIDEO_INPUT_FOLDER + file[0] + "." + file[1])
start_time = time()
width = FORCE_VIDEO_SIZE[0] or int(video.get(cv.CAP_PROP_FRAME_WIDTH))
height = FORCE_VIDEO_SIZE[1] or int(video.get(cv.CAP_PROP_FRAME_HEIGHT))
fps = video.get(cv.CAP_PROP_FPS)
resolution = (width, height)
frame_count = video.get(cv.CAP_PROP_FRAME_COUNT)
# Will be needed for ffmpeg
file.append(fps)
# Create output folder
folderpath = f"{cache_folder}{file[0]}-{file[1]}/"
# !mkdir $folderpath
file.append(folderpath)
counter = 1
length = frame_count if CROP_VIDEO_LENGTH is None else int(CROP_VIDEO_LENGTH * fps)
while counter <= length:
success, frame = video.read()
# If no new Frame could be loaded (aka the video ended)
if not success:
break
# Print progress
if counter % int(length / 100) == 0:
percentage = int(counter * 100 / length) + 1
string = 'X' * percentage
print(f"{percentage}% [{string.ljust(100)}] {counter}/{int(length)}", end="\r")
frame = cv.cvtColor(frame, cv.COLOR_BGR2RGB)
result = run_detector(classifier, frame)
if BLUR_OBJECTS:
blur_objects(frame, result["detection_boxes"], result["detection_class_entities"], result["detection_scores"])
if DRAW_BOXES:
draw_boxes(frame, result["detection_boxes"], result["detection_class_entities"], result["detection_scores"])
# Resize Frames
if (FORCE_VIDEO_SIZE != (None, None)):
frame = cv.resize(frame, dsize=resolution, interpolation=cv.INTER_CUBIC)
Save as png
frame = cv.cvtColor(frame, cv.COLOR_RGB2BGR)
name = f"{folderpath}{str(counter).zfill(6)}.png"
cv.imwrite(name, frame, [cv.IMWRITE_PNG_COMPRESSION, 4])
counter += 1
video.release()
print(f"\nName: {file[0]}\t Format: {file[1]}\t Size: {resolution}\t FPS: {fps:.2f}\t Duration: {(frame_count / fps):.1f}s\t Time: {(time() - start_time):.2f}s\n")
# -
# ### Compact Frames into a Video
#
# As a final step the single Frames will be compressed into a Video-File
#
# TODO: Add Audio
for file in videos:
print(file)
input_files = f"{file[3]}%06d.png"
output_filename = f"{VIDEO_OUTPUT_FOLDER}{FILENAME_AFTER_BLURRING.format(file[0])}." + (file[1] if FORCE_VIDEO_FORMAT is None else FORCE_VIDEO_FORMAT)
fps = file[2]
options = ""
if VIDEO_BITRATE is not None:
options += f" -b:v {VIDEO_BITRATE}"
# !ffmpeg -i $input_files -framerate $fps -y $options -loglevel +info $output_filename
# ## Cleanup
#
# Delete all tempdirs we created
#
if not VIDEO_CACHE_FOLDER:
shutil.rmtree(cache_folder)
| 002-Methods/001-Computing/PyDeepLearningVersion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=false editable=false dc={"key": "4"} run_control={"frozen": true} tags=["context"]
# ## 1. American Sign Language (ASL)
# <p>American Sign Language (ASL) is the primary language used by many deaf individuals in North America, and it is also used by hard-of-hearing and hearing individuals. The language is as rich as spoken languages and employs signs made with the hand, along with facial gestures and bodily postures.</p>
# <p><img src="https://assets.datacamp.com/production/project_509/img/asl.png" alt="american sign language"></p>
# <p>A lot of recent progress has been made towards developing computer vision systems that translate sign language to spoken language. This technology often relies on complex neural network architectures that can detect subtle patterns in streaming video. However, as a first step, towards understanding how to build a translation system, we can reduce the size of the problem by translating individual letters, instead of sentences.</p>
# <p><strong>In this notebook</strong>, we will train a convolutional neural network to classify images of American Sign Language (ASL) letters. After loading, examining, and preprocessing the data, we will train the network and test its performance.</p>
# <p>In the code cell below, we load the training and test data. </p>
# <ul>
# <li><code>x_train</code> and <code>x_test</code> are arrays of image data with shape <code>(num_samples, 3, 50, 50)</code>, corresponding to the training and test datasets, respectively.</li>
# <li><code>y_train</code> and <code>y_test</code> are arrays of category labels with shape <code>(num_samples,)</code>, corresponding to the training and test datasets, respectively.</li>
# </ul>
# + dc={"key": "4"} tags=["sample_code"]
# Import packages and set numpy random seed
import numpy as np
np.random.seed(5)
import tensorflow as tf
tf.set_random_seed(2)
from datasets import sign_language
import matplotlib.pyplot as plt
# %matplotlib inline
# Load pre-shuffled training and test datasets
(x_train, y_train), (x_test, y_test) = sign_language.load_data()
# + deletable=false editable=false dc={"key": "11"} run_control={"frozen": true} tags=["context"]
# ## 2. Visualize the training data
# <p>Now we'll begin by creating a list of string-valued labels containing the letters that appear in the dataset. Then, we visualize the first several images in the training data, along with their corresponding labels.</p>
# + dc={"key": "11"} tags=["sample_code"]
# Store labels of dataset
labels = ...
# Print the first several training images, along with the labels
fig = plt.figure(figsize=(20,5))
for i in range(36):
ax = fig.add_subplot(3, 12, i + 1, xticks=[], yticks=[])
ax.imshow(np.squeeze(x_train[i]))
ax.set_title("{}".format(labels[y_train[i]]))
plt.show()
# + deletable=false editable=false dc={"key": "18"} run_control={"frozen": true} tags=["context"]
# ## 3. Examine the dataset
# <p>Let's examine how many images of each letter can be found in the dataset.</p>
# <p>Remember that dataset has already been split into training and test sets for you, where <code>x_train</code> and <code>x_test</code> contain the images, and <code>y_train</code> and <code>y_test</code> contain their corresponding labels.</p>
# <p>Each entry in <code>y_train</code> and <code>y_test</code> is one of <code>0</code>, <code>1</code>, or <code>2</code>, corresponding to the letters <code>'A'</code>, <code>'B'</code>, and <code>'C'</code>, respectively.</p>
# <p>We will use the arrays <code>y_train</code> and <code>y_test</code> to verify that both the training and test sets each have roughly equal proportions of each letter.</p>
# + dc={"key": "18"} tags=["sample_code"]
# Number of A's in the training dataset
num_A_train = sum(y_train==0)
# Number of B's in the training dataset
num_B_train = ...
# Number of C's in the training dataset
num_C_train = ...
# Number of A's in the test dataset
num_A_test = sum(y_test==0)
# Number of B's in the test dataset
num_B_test = ...
# Number of C's in the test dataset
num_C_test = ...
# Print statistics about the dataset
print("Training set:")
print("\tA: {}, B: {}, C: {}".format(num_A_train, num_B_train, num_C_train))
print("Test set:")
print("\tA: {}, B: {}, C: {}".format(num_A_test, num_B_test, num_C_test))
# + deletable=false editable=false dc={"key": "25"} run_control={"frozen": true} tags=["context"]
# ## 4. One-hot encode the data
# <p>Currently, our labels for each of the letters are encoded as categorical integers, where <code>'A'</code>, <code>'B'</code> and <code>'C'</code> are encoded as <code>0</code>, <code>1</code>, and <code>2</code>, respectively. However, recall that Keras models do not accept labels in this format, and we must first one-hot encode the labels before supplying them to a Keras model.</p>
# <p>This conversion will turn the one-dimensional array of labels into a two-dimensional array.</p>
# <p><img src="https://assets.datacamp.com/production/project_509/img/onehot.png" alt="one-hot encoding"></p>
# <p>Each row in the two-dimensional array of one-hot encoded labels corresponds to a different image. The row has a <code>1</code> in the column that corresponds to the correct label, and <code>0</code> elsewhere. </p>
# <p>For instance, </p>
# <ul>
# <li><code>0</code> is encoded as <code>[1, 0, 0]</code>, </li>
# <li><code>1</code> is encoded as <code>[0, 1, 0]</code>, and </li>
# <li><code>2</code> is encoded as <code>[0, 0, 1]</code>.</li>
# </ul>
# + dc={"key": "25"} tags=["sample_code"]
from keras.utils import np_utils
# One-hot encode the training labels
y_train_OH = ...
# One-hot encode the test labels
y_test_OH = ...
# + deletable=false editable=false dc={"key": "32"} run_control={"frozen": true} tags=["context"]
# ## 5. Define the model
# <p>Now it's time to define a convolutional neural network to classify the data.</p>
# <p>This network accepts an image of an American Sign Language letter as input. The output layer returns the network's predicted probabilities that the image belongs in each category.</p>
# + dc={"key": "32"} tags=["sample_code"]
from keras.layers import Conv2D, MaxPooling2D
from keras.layers import Flatten, Dense
from keras.models import Sequential
model = Sequential()
# First convolutional layer accepts image input
model.add(Conv2D(filters=5, kernel_size=5, padding='same', activation='relu',
input_shape=(50, 50, 3)))
# Add a max pooling layer
model.add(...)
# Add a convolutional layer
model.add(...)
# Add another max pooling layer
model.add(...)
# Flatten and feed to output layer
model.add(Flatten())
model.add(Dense(3, activation='softmax'))
# Summarize the model
model.summary()
# + deletable=false editable=false dc={"key": "39"} run_control={"frozen": true} tags=["context"]
# ## 6. Compile the model
# <p>After we have defined a neural network in Keras, the next step is to compile it! </p>
# + dc={"key": "39"} tags=["sample_code"]
# Compile the model
model.compile(optimizer=...,
loss=...,
metrics=...)
# + deletable=false editable=false dc={"key": "46"} run_control={"frozen": true} tags=["context"]
# ## 7. Train the model
# <p>Once we have compiled the model, we're ready to fit it to the training data.</p>
# + dc={"key": "46"} tags=["sample_code"]
# Train the model
hist = model.fit(...)
# + deletable=false editable=false dc={"key": "53"} run_control={"frozen": true} tags=["context"]
# ## 8. Test the model
# <p>To evaluate the model, we'll use the test dataset. This will tell us how the network performs when classifying images it has never seen before!</p>
# <p>If the classification accuracy on the test dataset is similar to the training dataset, this is a good sign that the model did not overfit to the training data. </p>
# + dc={"key": "53"} tags=["sample_code"]
# Obtain accuracy on test set
score = model.evaluate(x=...,
y=...,
verbose=0)
print('Test accuracy:', score[1])
# + deletable=false editable=false dc={"key": "61"} run_control={"frozen": true} tags=["context"]
# ## 9. Visualize mistakes
# <p>Hooray! Our network gets very high accuracy on the test set! </p>
# <p>The final step is to take a look at the images that were incorrectly classified by the model. Do any of the mislabeled images look relatively difficult to classify, even to the human eye? </p>
# <p>Sometimes, it's possible to review the images to discover special characteristics that are confusing to the model. However, it is also often the case that it's hard to interpret what the model had in mind!</p>
# + dc={"key": "61"} tags=["sample_code"]
# Get predicted probabilities for test dataset
y_probs = ...
# Get predicted labels for test dataset
y_preds = ...
# Indices corresponding to test images which were mislabeled
bad_test_idxs = ...
# Print mislabeled examples
fig = plt.figure(figsize=(25,4))
for i, idx in enumerate(bad_test_idxs):
ax = fig.add_subplot(2, np.ceil(len(bad_test_idxs)/2), i + 1, xticks=[], yticks=[])
ax.imshow(np.squeeze(x_test[idx]))
ax.set_title("{} (pred: {})".format(labels[y_test[idx]], labels[y_preds[idx]]))
| ASL Recognition with Deep Learning/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
#
# Head model and forward computation
# ==================================
#
# The aim of this tutorial is to be a getting started for forward
# computation.
#
# For more extensive details and presentation of the general
# concepts for forward modeling. See `ch_forward`.
#
# +
import os.path as op
import mne
from mne.datasets import sample
data_path = sample.data_path()
# the raw file containing the channel location + types
raw_fname = data_path + '/MEG/sample/sample_audvis_raw.fif'
# The paths to Freesurfer reconstructions
subjects_dir = data_path + '/subjects'
subject = 'sample'
# -
# Computing the forward operator
# ------------------------------
#
# To compute a forward operator we need:
#
# - a ``-trans.fif`` file that contains the coregistration info.
# - a source space
# - the :term:`BEM` surfaces
#
#
# Compute and visualize BEM surfaces
# ----------------------------------
#
# The :term:`BEM` surfaces are the triangulations of the interfaces between
# different tissues needed for forward computation. These surfaces are for
# example the inner skull surface, the outer skull surface and the outer skin
# surface, a.k.a. scalp surface.
#
# Computing the BEM surfaces requires FreeSurfer and makes use of either of
# the two following command line tools:
#
# - `gen_mne_watershed_bem`
# - `gen_mne_flash_bem`
#
# Or by calling in a Python script one of the functions
# :func:`mne.bem.make_watershed_bem` or :func:`mne.bem.make_flash_bem`.
#
# Here we'll assume it's already computed. It takes a few minutes per subject.
#
# For EEG we use 3 layers (inner skull, outer skull, and skin) while for
# MEG 1 layer (inner skull) is enough.
#
# Let's look at these surfaces. The function :func:`mne.viz.plot_bem`
# assumes that you have the the *bem* folder of your subject FreeSurfer
# reconstruction the necessary files.
#
#
mne.viz.plot_bem(subject=subject, subjects_dir=subjects_dir,
brain_surfaces='white', orientation='coronal')
# Visualization the coregistration
# --------------------------------
#
# The coregistration is operation that allows to position the head and the
# sensors in a common coordinate system. In the MNE software the transformation
# to align the head and the sensors in stored in a so-called **trans file**.
# It is a FIF file that ends with ``-trans.fif``. It can be obtained with
# :func:`mne.gui.coregistration` (or its convenient command line
# equivalent `gen_mne_coreg`), or mrilab if you're using a Neuromag
# system.
#
# For the Python version see :func:`mne.gui.coregistration`
#
# Here we assume the coregistration is done, so we just visually check the
# alignment with the following code.
#
#
# +
# The transformation file obtained by coregistration
trans = data_path + '/MEG/sample/sample_audvis_raw-trans.fif'
info = mne.io.read_info(raw_fname)
# Here we look at the dense head, which isn't used for BEM computations but
# is useful for coregistration.
mne.viz.plot_alignment(info, trans, subject=subject, dig=True,
meg=['helmet', 'sensors'], subjects_dir=subjects_dir,
surfaces='head-dense')
# -
#
# Compute Source Space
# --------------------
#
# The source space defines the position and orientation of the candidate source
# locations. There are two types of source spaces:
#
# - **surface-based** source space when the candidates are confined to a
# surface.
#
# - **volumetric or discrete** source space when the candidates are discrete,
# arbitrarily located source points bounded by the surface.
#
# **Surface-based** source space is computed using
# :func:`mne.setup_source_space`, while **volumetric** source space is computed
# using :func:`mne.setup_volume_source_space`.
#
# We will now compute a surface-based source space with an OCT-6 resolution.
# See `setting_up_source_space` for details on source space definition
# and spacing parameter.
#
#
src = mne.setup_source_space(subject, spacing='oct6',
subjects_dir=subjects_dir, add_dist=False)
print(src)
# The surface based source space ``src`` contains two parts, one for the left
# hemisphere (4098 locations) and one for the right hemisphere
# (4098 locations). Sources can be visualized on top of the BEM surfaces
# in purple.
#
#
mne.viz.plot_bem(subject=subject, subjects_dir=subjects_dir,
brain_surfaces='white', src=src, orientation='coronal')
# To compute a volume based source space defined with a grid of candidate
# dipoles inside a sphere of radius 90mm centered at (0.0, 0.0, 40.0)
# you can use the following code.
# Obviously here, the sphere is not perfect. It is not restricted to the
# brain and it can miss some parts of the cortex.
#
#
# +
sphere = (0.0, 0.0, 40.0, 90.0)
vol_src = mne.setup_volume_source_space(subject, subjects_dir=subjects_dir,
sphere=sphere)
print(vol_src)
mne.viz.plot_bem(subject=subject, subjects_dir=subjects_dir,
brain_surfaces='white', src=vol_src, orientation='coronal')
# -
# To compute a volume based source space defined with a grid of candidate
# dipoles inside the brain (requires the :term:`BEM` surfaces) you can use the
# following.
#
#
# +
surface = op.join(subjects_dir, subject, 'bem', 'inner_skull.surf')
vol_src = mne.setup_volume_source_space(subject, subjects_dir=subjects_dir,
surface=surface)
print(vol_src)
mne.viz.plot_bem(subject=subject, subjects_dir=subjects_dir,
brain_surfaces='white', src=vol_src, orientation='coronal')
# -
# With the surface-based source space only sources that lie in the plotted MRI
# slices are shown. Let's see how to view all sources in 3D.
#
#
fig = mne.viz.plot_alignment(subject=subject, subjects_dir=subjects_dir,
surfaces='white', coord_frame='head',
src=src)
mne.viz.set_3d_view(fig, azimuth=173.78, elevation=101.75,
distance=0.30, focalpoint=(-0.03, -0.01, 0.03))
#
# Compute forward solution
# ------------------------
#
# We can now compute the forward solution.
# To reduce computation we'll just compute a single layer BEM (just inner
# skull) that can then be used for MEG (not EEG).
#
# We specify if we want a one-layer or a three-layer BEM using the
# conductivity parameter.
#
# The BEM solution requires a BEM model which describes the geometry
# of the head the conductivities of the different tissues.
#
#
conductivity = (0.3,) # for single layer
# conductivity = (0.3, 0.006, 0.3) # for three layers
model = mne.make_bem_model(subject='sample', ico=4,
conductivity=conductivity,
subjects_dir=subjects_dir)
bem = mne.make_bem_solution(model)
# Note that the :term:`BEM` does not involve any use of the trans file. The BEM
# only depends on the head geometry and conductivities.
# It is therefore independent from the MEG data and the head position.
#
# Let's now compute the forward operator, commonly referred to as the
# gain or leadfield matrix.
#
# See :func:`mne.make_forward_solution` for details on parameters meaning.
#
#
fwd = mne.make_forward_solution(raw_fname, trans=trans, src=src, bem=bem,
meg=True, eeg=False, mindist=5.0, n_jobs=2)
print(fwd)
# We can explore the content of fwd to access the numpy array that contains
# the gain matrix.
#
#
leadfield = fwd['sol']['data']
print("Leadfield size : %d sensors x %d dipoles" % leadfield.shape)
# To extract the numpy array containing the forward operator corresponding to
# the source space `fwd['src']` with cortical orientation constraint
# we can use the following:
#
#
fwd_fixed = mne.convert_forward_solution(fwd, surf_ori=True, force_fixed=True,
use_cps=True)
leadfield = fwd_fixed['sol']['data']
print("Leadfield size : %d sensors x %d dipoles" % leadfield.shape)
# This is equivalent to the following code that explicitly applies the
# forward operator to a source estimate composed of the identity operator:
#
#
# +
import numpy as np # noqa
n_dipoles = leadfield.shape[1]
vertices = [src_hemi['vertno'] for src_hemi in fwd_fixed['src']]
stc = mne.SourceEstimate(1e-9 * np.eye(n_dipoles), vertices, tmin=0., tstep=1)
leadfield = mne.apply_forward(fwd_fixed, stc, info).data / 1e-9
# -
# To save to disk a forward solution you can use
# :func:`mne.write_forward_solution` and to read it back from disk
# :func:`mne.read_forward_solution`. Don't forget that FIF files containing
# forward solution should end with *-fwd.fif*.
#
# To get a fixed-orientation forward solution, use
# :func:`mne.convert_forward_solution` to convert the free-orientation
# solution to (surface-oriented) fixed orientation.
#
#
# Exercise
# --------
#
# By looking at
# `sphx_glr_auto_examples_forward_plot_forward_sensitivity_maps.py`
# plot the sensitivity maps for EEG and compare it with the MEG, can you
# justify the claims that:
#
# - MEG is not sensitive to radial sources
# - EEG is more sensitive to deep sources
#
# How will the MEG sensitivity maps and histograms change if you use a free
# instead if a fixed/surface oriented orientation?
#
# Try this changing the mode parameter in :func:`mne.sensitivity_map`
# accordingly. Why don't we see any dipoles on the gyri?
#
#
| dev/_downloads/e6e21719275dd1fe5b070e9c49c33bc5/plot_forward.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Using a separate (non-Wikidata) Wikibase instance
# ### In this example, we have downloaded and run the wikibase docker images from here: https://github.com/wmde/wikibase-docker
# #### *Before running this example, I created an item on my local wikibase with the label "Greg"*
from wikidataintegrator import wdi_core, wdi_login, wdi_helpers
from wikidataintegrator.wdi_core import WDItemEngine
# set urls
mediawiki_api_url = 'http://localhost:8181/w/api.php'
sparql_endpoint_url = 'http://localhost:8282/proxy/wdqs/bigdata/namespace/wdq/sparql'
# create a WDItemEngine class with the urls preset
LocalItemEngine = WDItemEngine.wikibase_item_engine_factory(mediawiki_api_url, sparql_endpoint_url)
# we can now retrieve this item from our local wikibase
litem = LocalItemEngine(wd_item_id="Q2")
litem.get_label()
# and can also still access regular wikidata
item = WDItemEngine(wd_item_id="Q2")
item.get_label()
# to login:
local_login = wdi_login.WDLogin("test", "password", mediawiki_api_url)
| doc/non_wikidata_wikibase.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import torch
import os
# +
tokenizer = None
import numpy as np
import torch
import os
config_switch=os.getenv('DOCKER', 'local')
if config_switch=='local':
startup_nodes = [{"host": "127.0.0.1", "port": "30001"}, {"host": "127.0.0.1", "port":"30002"}, {"host":"127.0.0.1", "port":"30003"}]
else:
startup_nodes = [{"host": "rgcluster", "port": "30001"}, {"host": "rgcluster", "port":"30002"}, {"host":"rgcluster", "port":"30003"}]
try:
from redisai import ClusterClient
redisai_cluster_client = ClusterClient(startup_nodes=startup_nodes)
except:
print("Redis Cluster is not available")
def loadTokeniser():
global tokenizer
from transformers import BertTokenizerFast
tokenizer = BertTokenizerFast.from_pretrained("bert-large-uncased-whole-word-masking-finetuned-squad")
return tokenizer
def qa_redisai(question, sentence_key,hash_tag):
### question is encoded
### use pre-computed context/answer text tensor
global tokenizer
if not tokenizer:
tokenizer=loadTokeniser()
token_key = f"tokenized:bert:qa:{sentence_key}"
input_ids_question = tokenizer.encode(question, add_special_tokens=True, truncation=True, return_tensors="np")
input_ids_context=redisai_cluster_client.tensorget(token_key)
input_ids = np.append(input_ids_question,input_ids_context)
print(input_ids.shape)
print(input_ids)
attention_mask = np.array([[1]*len(input_ids)])
input_idss=np.array([input_ids])
print(input_idss.shape)
print("Attention mask shape ",attention_mask.shape)
num_seg_a=input_ids_question.shape[1]
print(num_seg_a)
num_seg_b=input_ids_context.shape[0]
print(num_seg_b)
token_type_ids = np.array([0]*num_seg_a + [1]*num_seg_b)
print("Segments id",token_type_ids.shape)
redisai_cluster_client.tensorset(f'input_ids{hash_tag}', input_idss)
redisai_cluster_client.tensorset(f'attention_mask{hash_tag}', attention_mask)
redisai_cluster_client.tensorset(f'token_type_ids{hash_tag}', token_type_ids)
redisai_cluster_client.modelrun(f'bert-qa{hash_tag}', [f'input_ids{hash_tag}', f'attention_mask{hash_tag}', f'token_type_ids{hash_tag}'],
[f'answer_start_scores{hash_tag}', f'answer_end_scores{hash_tag}'])
print(f"Model run on {hash_tag}")
answer_start_scores = redisai_cluster_client.tensorget(f'answer_start_scores{hash_tag}')
answer_end_scores = redisai_cluster_client.tensorget(f'answer_end_scores{hash_tag}')
answer_start = np.argmax(answer_start_scores)
answer_end = np.argmax(answer_end_scores) + 1
answer = tokenizer.convert_tokens_to_string(tokenizer.convert_ids_to_tokens(input_ids[answer_start:answer_end], skip_special_tokens = True))
print(answer)
return answer
# -
question="What about frequencies of occurenence RNA?"
qa_redisai(question,"PMC222961.xml:{06S}:26",'{06S}')
question="Effectiveness of community contact reduction"
sentence_key="PMC261870.xml:{06S}:26"
token_key = f"tokenized:bert:qa:{sentence_key}"
redisai_cluster_client.connection_pool
# %%time
slot = redisai_cluster_client.connection_pool.nodes.keyslot(sentence_key)
node = redisai_cluster_client.connection_pool.get_master_node_by_slot(slot)
connection = redisai_cluster_client.connection_pool.get_connection_by_node(node)
connection.send_command('RG.TRIGGER',"RunQABERT",sentence_key,question)
print(connection.__dict__)
print(redisai_cluster_client.parse_response(connection,"RG.TRIGGER"))
# %%time
slot = redisai_cluster_client.connection_pool.nodes.keyslot(sentence_key)
node = redisai_cluster_client.connection_pool.get_master_node_by_slot(slot)
connection = redisai_cluster_client.connection_pool.get_connection_by_node(node)
connection.send_command('RG.TRIGGER',"RunQABERT",sentence_key,question)
print(connection.__dict__)
print(redisai_cluster_client.parse_response(connection,"RG.TRIGGER"))
question
from rediscluster import RedisCluster
startup_nodes = [{"host": "127.0.0.1", "port": "30001"}, {"host": "127.0.0.1", "port":"30002"}, {"host":"127.0.0.1", "port":"30003"}]
rc = RedisCluster(startup_nodes=startup_nodes, decode_responses=True)
object_methods = [method_name for method_name in dir(rc)
if callable(getattr(rc, method_name))]
sentence_key="PMC261870.xml:{06S}:26"
question="Effectiveness of community contact reduction"
rc.execute_command('RG.TRIGGER',"RunQABERT",sentence_key,question)
command='RG.TRIGGER'
rc.determine_node('RG.TRIGGER',"RunQABERT",sentence_key,question)
print(rc.nodes_flags.get(command))
args=[1,2]
len(args)>=1
rc.execute_command('RG.TRIGGER',"RunQABERT",sentence_key,question)
from rediscluster import RedisCluster
# +
import logging
from rediscluster import RedisCluster
logging.basicConfig()
logger = logging.getLogger('rediscluster')
logger.setLevel(logging.DEBUG)
logger.propagate = True
# -
rc = RedisCluster(startup_nodes=startup_nodes, decode_responses=True)
rc.execute_command('RG.TRIGGER',"RunQABERT",sentence_key,question)
rc.connection_pool.nodes.random_node()
list(rc.connection_pool.nodes.all_masters())
rc.get(sentence_key)
print(rc.parse_response(connection,"RG.TRIGGER"))
result=rc.get("cache{06S}_PMC261870.xml:{06S}:26_Effectiveness of community contact reduction")
print(result)
# +
tokenizer = None
model = None
import torch
def loadTokeniser():
global tokenizer
from transformers import AutoTokenizer
tokenizer = AutoTokenizer.from_pretrained("bert-large-uncased-whole-word-masking-finetuned-squad", torchscript=True)
return tokenizer
def loadModel():
global model
from transformers import AutoModelForQuestionAnswering
model = AutoModelForQuestionAnswering.from_pretrained("bert-large-uncased-whole-word-masking-finetuned-squad", torchscript=True)
return model
def qa(question, content_text):
global tokenizer, model
if not tokenizer:
tokenizer=loadTokeniser()
if not model:
model=loadModel()
inputs = tokenizer.encode_plus(question, content_text, add_special_tokens=True, return_tensors="pt")
input_ids = inputs["input_ids"].tolist()[0]
print(input_ids)
answer_start_scores, answer_end_scores = model(**inputs,return_dict=False)
answer_start = torch.argmax(
answer_start_scores
) # Get the most likely beginning of answer with the argmax of the score
answer_end = torch.argmax(answer_end_scores) + 1 # Get the most likely end of answer with the argmax of the score
answer = tokenizer.convert_tokens_to_string(tokenizer.convert_ids_to_tokens(input_ids[answer_start:answer_end]))
return answer
# -
content_text="The frequencies of occurrence for i nucleotides were compared to the random RNA counterparts having the same base proportion in order to compute the a value that reflected their i nucleotide bias Table 2"
question="What about frequencies of occurenence RNA?"
# +
tokenizer = None
model = None
import torch
def loadTokeniser():
global tokenizer
from transformers import AutoTokenizer
tokenizer = AutoTokenizer.from_pretrained("bert-large-uncased-whole-word-masking-finetuned-squad")
return tokenizer
def loadModel():
global model
from transformers import AutoModelForQuestionAnswering
model = AutoModelForQuestionAnswering.from_pretrained("bert-large-uncased-whole-word-masking-finetuned-squad")
return model
if not tokenizer:
tokenizer=loadTokeniser()
if not model:
model=loadModel()
inputs = tokenizer.encode_plus(question, content_text, add_special_tokens=True, return_tensors="pt")
input_ids = inputs["input_ids"].tolist()[0]
print(input_ids)
answer_start_scores, answer_end_scores = model(**inputs,return_dict=False)
answer_start = torch.argmax(
answer_start_scores
) # Get the most likely beginning of answer with the argmax of the score
answer_end = torch.argmax(answer_end_scores) + 1 # Get the most likely end of answer with the argmax of the score
answer = tokenizer.convert_tokens_to_string(tokenizer.convert_ids_to_tokens(input_ids[answer_start:answer_end]))
# -
print(answer)
# %%time
print(qa(question,content_text))
"cache{06S}_PMC222961.xml:{06S}:26_%s" % question
question="When air samples collected?"
"cache{5M5}_PMC140314.xml:{5M5}:44_%s" % question
rc.get("cache{5M5}_PMC261870.xml:{5M5}:26_%s" % question)
def print_tokens(input_ids):
# BERT only needs the token IDs, but for the purpose of inspecting the
# tokenizer's behavior, let's also get the token strings and display them.
tokens = tokenizer.convert_ids_to_tokens(input_ids)
# For each token and its id...
for token, id in zip(tokens, input_ids):
# If this is the [SEP] token, add some space around it to make it stand out.
if id == tokenizer.sep_token_id:
print('')
# Print the token string and its ID in two columns.
print('{:<12} {:>6,}'.format(token, id))
if id == tokenizer.sep_token_id:
print('')
def answer_question(question, answer_text):
'''
Takes a `question` string and an `answer_text` string (which contains the
answer), and identifies the words within the `answer_text` that are the
answer. Prints them out.
'''
# ======== Tokenize ========
# Apply the tokenizer to the input text, treating them as a text-pair.
input_ids = tokenizer.encode(question, answer_text)
# Report how long the input sequence is.
print('Query has {:,} tokens.\n'.format(len(input_ids)))
# ======== Set Segment IDs ========
# Search the input_ids for the first instance of the `[SEP]` token.
sep_index = input_ids.index(tokenizer.sep_token_id)
# The number of segment A tokens includes the [SEP] token istelf.
num_seg_a = sep_index + 1
# The remainder are segment B.
num_seg_b = len(input_ids) - num_seg_a
# Construct the list of 0s and 1s.
segment_ids = [0]*num_seg_a + [1]*num_seg_b
# There should be a segment_id for every input token.
assert len(segment_ids) == len(input_ids)
# ======== Evaluate ========
# Run our example question through the model.
start_scores, end_scores = model(torch.tensor([input_ids]), # The tokens representing our input text.
token_type_ids=torch.tensor([segment_ids]),return_dict=False) # The segment IDs to differentiate question from answer_text
# ======== Reconstruct Answer ========
# Find the tokens with the highest `start` and `end` scores.
answer_start = torch.argmax(start_scores)
answer_end = torch.argmax(end_scores)
# Get the string versions of the input tokens.
tokens = tokenizer.convert_ids_to_tokens(input_ids)
# Start with the first token.
answer = tokens[answer_start]
# Select the remaining answer tokens and join them with whitespace.
for i in range(answer_start + 1, answer_end + 1):
# If it's a subword token, then recombine it with the previous token.
if tokens[i][0:2] == '##':
answer += tokens[i][2:]
# Otherwise, add a space then the token.
else:
answer += ' ' + tokens[i]
print('Answer: "' + answer + '"')
answer_question(question, content_text)
"""
hget sentence:PMC222961.xml:{06S} 26
"The frequencies of occurrence for i nucleotides were compared to the random RNA counterparts having the same base proportion in order to compute the a value that reflected their i nucleotide bias Table 2
""""
context="Finally there are many other possible scenarios and as stated in the conclusion of the wHO panel held on 15 17 May 2003 in Geneva Participants from the main outbreak sites noted the striking similarity of the pattern of outbreaks in different countries and the consistent effectiveness of specific control measures including early identification and isolation of patients vigorous contact tracing management of close contacts by home confinement or quarantine and public information and education to encourage prompt reporting of symptoms"
question="What about community contact reduction?"
answer_question(question, context)
| the-pattern-api/qasearch/experiments/!TestBERTQA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
import pandas as pd
import numpy as np
from Bio import SeqIO
from Bio.SeqRecord import SeqRecord
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import stats
def calc_frequencies(cov, gene, window):
#Find percent polymorphism at each site
#Also determine whether polymorphism is silent or replacement
input_file_outgroup = '../../config/'+str(cov)+'_'+str(gene)+'_reference.gb'
input_file_alignment = '../../results/aligned_'+str(cov)+'_'+str(gene)+'.fasta'
metafile = '../../results/metadata_'+str(cov)+'_'+str(gene)+'.tsv'
#Subset data based on time windows
meta = pd.read_csv(metafile, sep = '\t')
meta.drop(meta[meta['date']=='?'].index, inplace=True)
meta['year'] = meta['date'].str[:4].astype('int')
date_range = meta['year'].max() - meta['year'].min()
#Group viruses by time windows
virus_time_subset = {}
if window == 'all':
years = str(meta['year'].min()) + '-' + str(meta['year'].max())
virus_time_subset[years] = meta['strain'].tolist()
else:
date_window_start = meta['year'].min()
date_window_end = meta['year'].min() + window
while date_window_end <= meta['year'].max():
years = str(date_window_start) + '-' + str(date_window_end)
strains = meta[(meta['year']>=date_window_start) & (meta['year']<date_window_end)]['strain'].tolist()
virus_time_subset[years] = strains
date_window_end += window
date_window_start += window
#initiate lists to record all time windows
year_windows = []
seqs_in_window = []
polymorphic_frequencies = []
replacement_frequencies = []
silent_frequencies = []
for years, subset_viruses in virus_time_subset.items():
if len(subset_viruses) != 0:
year_windows.append(years)
seqs_in_window.append(len(subset_viruses))
outgroup_seq = ''
outgroup_aa_seq = ''
with open(input_file_outgroup, "r") as outgroup_handle:
for outgroup in SeqIO.parse(outgroup_handle, "genbank"):
outgroup_seq = outgroup.seq
outgroup_aa_seq = outgroup.seq.translate()
count_polymorphic = np.zeros(len(outgroup.seq))
count_total_unambiguous = np.zeros(len(outgroup.seq))
count_replacement_mutations = np.zeros(len(outgroup.seq))
count_silent_mutations = np.zeros(len(outgroup.seq))
with open(input_file_alignment, "r") as aligned_handle:
for virus in SeqIO.parse(aligned_handle, "fasta"):
#Only viruses in time window
if virus.id in subset_viruses:
#check
if len(virus.seq) != len(outgroup_seq):
print(virus)
elif len(virus.seq) == len(outgroup_seq):
for pos in range(len(outgroup_seq)):
outgroup_nt = str(outgroup_seq[pos])
virus_nt = str(virus.seq[pos])
#skip ambiguous sites
if virus_nt != 'N':
count_total_unambiguous[pos]+=1
if virus_nt != outgroup_nt:
count_polymorphic[pos]+=1
#determine silent or replacement
codon = math.floor(pos/3)
codon_pos = pos-(codon*3)
if codon_pos == 0:
codon_nt = virus.seq[pos:(pos+3)]
elif codon_pos == 1:
codon_nt = virus.seq[(pos-1):(pos+2)]
elif codon_pos == 2:
codon_nt = virus.seq[(pos-2):(pos+1)]
codon_aa = codon_nt.translate()
outgroup_aa = outgroup_aa_seq[codon]
if codon_aa != outgroup_aa:
count_replacement_mutations[pos]+=1
elif codon_aa == outgroup_aa:
count_silent_mutations[pos]+=1
polymorphic_frequencies_window = count_polymorphic/count_total_unambiguous
replacement_frequencies_window = count_replacement_mutations/count_polymorphic
silent_frequencies_window = count_silent_mutations/count_polymorphic
polymorphic_frequencies.append(polymorphic_frequencies_window)
replacement_frequencies.append(replacement_frequencies_window)
silent_frequencies.append(silent_frequencies_window)
return year_windows, seqs_in_window, polymorphic_frequencies, replacement_frequencies, silent_frequencies
# +
def bhatt_variables(cov, gene, window):
(year_windows,seqs_in_window, polymorphic_frequencies, replacement_frequencies, silent_frequencies) = calc_frequencies(cov, gene, window)
#Initiate lists to store a values
window_midpoint = []
adaptive_substitutions = []
#Categorize sites into fixation, low freq, medium freq and high freq polymorphisms (and silent or replacement)
for years_window in range(len(polymorphic_frequencies)):
#don't use windows with fewer than 3 sequences
if seqs_in_window[years_window] >= 3:
window_start = int(year_windows[years_window][0:4])
window_end = int(year_windows[years_window][-4:])
window_midpoint.append((window_start + window_end)/2)
sf = 0
rf = 0
sh = 0
rh = 0
sm = 0
rm = 0
sl = 0
rl = 0
window_polymorphic_freqs = polymorphic_frequencies[years_window]
for site in range(len(window_polymorphic_freqs)):
pfreq = window_polymorphic_freqs[site]
#ignore sites with no polymorphisms?
if pfreq!= 0:
if pfreq == 1:
sf+= (pfreq*silent_frequencies[years_window][site])
rf+= (pfreq*replacement_frequencies[years_window][site])
elif pfreq > 0.75:
sh+= (pfreq*silent_frequencies[years_window][site])
rh+= (pfreq*replacement_frequencies[years_window][site])
elif pfreq > 0.15 and pfreq < 0.75:
sm+= (pfreq*silent_frequencies[years_window][site])
rm+= (pfreq*replacement_frequencies[years_window][site])
elif pfreq < 0.15:
sl+= (pfreq*silent_frequencies[years_window][site])
rl+= (pfreq*replacement_frequencies[years_window][site])
# print(year_windows[years_window])
# print(sf, rf, sh, rh, sm, rm, sl, rl)
#Calculate equation 1: number of nonneutral sites
al = rl - sl*(rm/sm)
ah = rh - sh*(rm/sm)
af = rf - sf*(rm/sm)
#set negative a values to zero
if al < 0:
al = 0
if ah < 0:
ah = 0
if af < 0:
af = 0
# print(al, ah, af)
#Calculate the number and proportion of all fixed or high-freq sites that have undergone adaptive change
number_adaptive_substitutions = af + ah
adaptive_substitutions.append(number_adaptive_substitutions)
proportion_adaptive_sites = (af + ah)/(rf +rh)
# get coeffs of linear fit
slope, intercept, r_value, p_value, std_err = stats.linregress(window_midpoint, adaptive_substitutions)
ax = sns.regplot(x= window_midpoint, y=adaptive_substitutions,
line_kws={'label':"y={0:.1f}x+{1:.1f}".format(slope,intercept)})
plt.ylabel('number of adaptive substitutions')
plt.xlabel('year')
ax.legend()
plt.show()
# -
bhatt_variables('oc43', 'spike', 5)
# +
#Bhatt method to calculate rate of adaptation
def calc_frequencies_old(cov, gene):
#Find percent polymorphism at each site
#Also determine whether polymorphism is silent or replacement
input_file_outgroup = '../../config/'+str(cov)+'_'+str(gene)+'_reference.gb'
input_file_alignment = '../../results/aligned_'+str(cov)+'_'+str(gene)+'.fasta'
outgroup_seq = ''
outgroup_aa_seq = ''
with open(input_file_outgroup, "r") as outgroup_handle:
for outgroup in SeqIO.parse(outgroup_handle, "genbank"):
outgroup_seq = outgroup.seq
outgroup_aa_seq = outgroup.seq.translate()
count_polymorphic = np.zeros(len(outgroup.seq))
count_total_unambiguous = np.zeros(len(outgroup.seq))
count_replacement_mutations = np.zeros(len(outgroup.seq))
count_silent_mutations = np.zeros(len(outgroup.seq))
with open(input_file_alignment, "r") as aligned_handle:
for virus in SeqIO.parse(aligned_handle, "fasta"):
#check
if len(virus.seq) != len(outgroup_seq):
print(virus)
elif len(virus.seq) == len(outgroup_seq):
for pos in range(len(outgroup_seq)):
outgroup_nt = str(outgroup_seq[pos])
virus_nt = str(virus.seq[pos])
#skip ambiguous sites
if virus_nt != 'N':
count_total_unambiguous[pos]+=1
if virus_nt != outgroup_nt:
count_polymorphic[pos]+=1
#determine silent or replacement
codon = math.floor(pos/3)
codon_pos = pos-(codon*3)
if codon_pos == 0:
codon_nt = virus.seq[pos:(pos+3)]
elif codon_pos == 1:
codon_nt = virus.seq[(pos-1):(pos+2)]
elif codon_pos == 2:
codon_nt = virus.seq[(pos-2):(pos+1)]
codon_aa = codon_nt.translate()
outgroup_aa = outgroup_aa_seq[codon]
if codon_aa != outgroup_aa:
count_replacement_mutations[pos]+=1
elif codon_aa == outgroup_aa:
count_silent_mutations[pos]+=1
polymorphic_frequencies = count_polymorphic/count_total_unambiguous
replacement_frequencies = count_replacement_mutations/count_polymorphic
silent_frequencies = count_silent_mutations/count_polymorphic
return polymorphic_frequencies, replacement_frequencies, silent_frequencies
# +
def calc_bhatt_variables_old(cov, gene):
(polymorphic_frequencies, replacement_frequencies, silent_frequencies) = calc_frequencies(cov, gene)
#Categorize sites into fixation, low freq, medium freq and high freq polymorphisms (and silent or replacement)
sf = 0
rf = 0
sh = 0
rh = 0
sm = 0
rm = 0
sl = 0
rl = 0
for site in range(len(polymorphic_frequencies)):
pfreq = polymorphic_frequencies[site]
#ignore sites with no polymorphisms?
if pfreq!= 0:
if pfreq == 1:
sf+= (pfreq*silent_frequencies[site])
rf+= (pfreq*replacement_frequencies[site])
elif pfreq > 0.75:
sh+= (pfreq*silent_frequencies[site])
rh+= (pfreq*replacement_frequencies[site])
elif pfreq > 0.15 and pfreq < 0.75:
sm+= (pfreq*silent_frequencies[site])
rm+= (pfreq*replacement_frequencies[site])
elif pfreq < 0.15:
sl+= (pfreq*silent_frequencies[site])
rl+= (pfreq*replacement_frequencies[site])
# print(sf, rf, sh, rh, sm, rm, sl, rl)
#Calculate equation 1: number of nonneutral sites
al = rl - sl*(rm/sm)
ah = rh - sh*(rm/sm)
af = rf - sf*(rm/sm)
print(al, ah, af)
#Calculate the proportion of all fixed or high-freq sites that have undergone adaptive change
proportion_adaptive_sites = (af + ah)/(rf +rh)
# print(adaptive_sites)
# -
calc_bhatt_variables_old('hku1', 'spike')
| data-wrangling/.ipynb_checkpoints/bhatt_2011-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Learn to calculate with seq2seq model
#
# In this assignment, you will learn how to use neural networks to solve sequence-to-sequence prediction tasks. Seq2Seq models are very popular these days because they achieve great results in Machine Translation, Text Summarization, Conversational Modeling and more.
#
# Using sequence-to-sequence modeling you are going to build a calculator for evaluating arithmetic expressions, by taking an equation as an input to the neural network and producing an answer as it's output.
#
# The resulting solution for this problem will be based on state-of-the-art approaches for sequence-to-sequence learning and you should be able to easily adapt it to solve other tasks. However, if you want to train your own machine translation system or intellectual chat bot, it would be useful to have access to compute resources like GPU, and be patient, because training of such systems is usually time consuming.
#
# ### Libraries
#
# For this task you will need the following libraries:
# - [TensorFlow](https://www.tensorflow.org) — an open-source software library for Machine Intelligence.
# - [scikit-learn](http://scikit-learn.org/stable/index.html) — a tool for data mining and data analysis.
#
# If you have never worked with TensorFlow, you will probably want to read some tutorials during your work on this assignment, e.g. [Neural Machine Translation](https://www.tensorflow.org/tutorials/seq2seq) tutorial deals with very similar task and can explain some concepts to you.
# ### Data
#
# One benefit of this task is that you don't need to download any data — you will generate it on your own! We will use two operators (addition and subtraction) and work with positive integer numbers in some range. Here are examples of correct inputs and outputs:
#
# Input: '1+2'
# Output: '3'
#
# Input: '0-99'
# Output: '-99'
#
# *Note, that there are no spaces between operators and operands.*
#
#
# Now you need to implement the function *generate_equations*, which will be used to generate the data.
import random
def generate_equations(allowed_operators, dataset_size, min_value, max_value):
"""Generates pairs of equations and solutions to them.
Each equation has a form of two integers with an operator in between.
Each solution is an integer with the result of the operaion.
allowed_operators: list of strings, allowed operators.
dataset_size: an integer, number of equations to be generated.
min_value: an integer, min value of each operand.
max_value: an integer, max value of each operand.
result: a list of tuples of strings (equation, solution).
"""
sample = []
for _ in range(dataset_size):
######################################
######### YOUR CODE HERE #############
######################################
a = random.randint(min_value, max_value)
b = random.randint(min_value, max_value)
op = random.choice(allowed_operators)
equation = str(a) + op + str(b)
solution = str(eval(equation))
sample.append((equation, solution))
return sample
# To check the correctness of your implementation, use *test_generate_equations* function:
def test_generate_equations():
allowed_operators = ['+', '-']
dataset_size = 10
for (input_, output_) in generate_equations(allowed_operators, dataset_size, 0, 100):
if not (type(input_) is str and type(output_) is str):
return "Both parts should be strings."
if eval(input_) != int(output_):
return "The (equation: {!r}, solution: {!r}) pair is incorrect.".format(input_, output_)
return "Tests passed."
print(test_generate_equations())
# Finally, we are ready to generate the train and test data for the neural network:
from sklearn.model_selection import train_test_split
# +
allowed_operators = ['+', '-']
dataset_size = 100000
data = generate_equations(allowed_operators, dataset_size, min_value=0, max_value=9999)
train_set, test_set = train_test_split(data, test_size=0.2, random_state=42)
# -
# ## Prepare data for the neural network
#
# The next stage of data preparation is creating mappings of the characters to their indices in some vocabulary. Since in our task we already know which symbols will appear in the inputs and outputs, generating the vocabulary is a simple step.
#
# #### How to create dictionaries for other task
#
# First of all, you need to understand what is the basic unit of the sequence in your task. In our case, we operate on symbols and the basic unit is a symbol. The number of symbols is small, so we don't need to think about filtering/normalization steps. However, in other tasks, the basic unit is often a word, and in this case the mapping would be *word $\to$ integer*. The number of words might be huge, so it would be reasonable to filter them, for example, by frequency and leave only the frequent ones. Other strategies that your should consider are: data normalization (lowercasing, tokenization, how to consider punctuation marks), separate vocabulary for input and for output (e.g. for machine translation), some specifics of the task.
word2id = {symbol:i for i, symbol in enumerate('#^$+-1234567890')}
id2word = {i:symbol for symbol, i in word2id.items()}
# #### Special symbols
start_symbol = '^'
end_symbol = '$'
padding_symbol = '#'
# You could notice that we have added 3 special symbols: '^', '\$' and '#':
# - '^' symbol will be passed to the network to indicate the beginning of the decoding procedure. We will discuss this one later in more details.
# - '\$' symbol will be used to indicate the *end of a string*, both for input and output sequences.
# - '#' symbol will be used as a *padding* character to make lengths of all strings equal within one training batch.
#
# People have a bit different habits when it comes to special symbols in encoder-decoder networks, so don't get too much confused if you come across other variants in tutorials you read.
# #### Padding
# When vocabularies are ready, we need to be able to convert a sentence to a list of vocabulary word indices and back. At the same time, let's care about padding. We are going to preprocess each sequence from the input (and output ground truth) in such a way that:
# - it has a predefined length *padded_len*
# - it is probably cut off or padded with the *padding symbol* '#'
# - it *always* ends with the *end symbol* '$'
#
# We will treat the original characters of the sequence **and the end symbol** as the valid part of the input. We will store *the actual length* of the sequence, which includes the end symbol, but does not include the padding symbols.
# Now you need to implement the function *sentence_to_ids* that does the described job.
def sentence_to_ids(sentence, word2id, padded_len):
""" Converts a sequence of symbols to a padded sequence of their ids.
sentence: a string, input/output sequence of symbols.
word2id: a dict, a mapping from original symbols to ids.
padded_len: an integer, a desirable length of the sequence.
result: a tuple of (a list of ids, an actual length of sentence).
"""
sent_len = min(len(sentence) + 1, padded_len)
plen = max(0, padded_len - sent_len)
sent_ids = [word2id[word] for word in sentence[:sent_len - 1]] + [word2id[end_symbol]] + [word2id[padding_symbol]]*plen
return sent_ids, sent_len
# Check that your implementation is correct:
def test_sentence_to_ids():
sentences = [("123+123", 7), ("123+123", 8), ("123+123", 10)]
expected_output = [([5, 6, 7, 3, 5, 6, 2], 7),
([5, 6, 7, 3, 5, 6, 7, 2], 8),
([5, 6, 7, 3, 5, 6, 7, 2, 0, 0], 8)]
for (sentence, padded_len), (sentence_ids, expected_length) in zip(sentences, expected_output):
output, length = sentence_to_ids(sentence, word2id, padded_len)
if output != sentence_ids:
return("Convertion of '{}' for padded_len={} to {} is incorrect.".format(
sentence, padded_len, output))
if length != expected_length:
return("Convertion of '{}' for padded_len={} has incorrect actual length {}.".format(
sentence, padded_len, length))
return("Tests passed.")
print(test_sentence_to_ids())
# We also need to be able to get back from indices to symbols:
def ids_to_sentence(ids, id2word):
""" Converts a sequence of ids to a sequence of symbols.
ids: a list, indices for the padded sequence.
id2word: a dict, a mapping from ids to original symbols.
result: a list of symbols.
"""
return [id2word[i] for i in ids]
# #### Generating batches
# The final step of data preparation is a function that transforms a batch of sentences to a list of lists of indices.
def batch_to_ids(sentences, word2id, max_len):
"""Prepares batches of indices.
Sequences are padded to match the longest sequence in the batch,
if it's longer than max_len, then max_len is used instead.
sentences: a list of strings, original sequences.
word2id: a dict, a mapping from original symbols to ids.
max_len: an integer, max len of sequences allowed.
result: a list of lists of ids, a list of actual lengths.
"""
max_len_in_batch = min(max(len(s) for s in sentences) + 1, max_len)
batch_ids, batch_ids_len = [], []
for sentence in sentences:
ids, ids_len = sentence_to_ids(sentence, word2id, max_len_in_batch)
batch_ids.append(ids)
batch_ids_len.append(ids_len)
return batch_ids, batch_ids_len
# The function *generate_batches* will help to generate batches with defined size from given samples.
def generate_batches(samples, batch_size=64):
X, Y = [], []
for i, (x, y) in enumerate(samples, 1):
X.append(x)
Y.append(y)
if i % batch_size == 0:
yield X, Y
X, Y = [], []
if X and Y:
yield X, Y
# To illustrate the result of the implemented functions, run the following cell:
sentences = train_set[0]
ids, sent_lens = batch_to_ids(sentences, word2id, max_len=10)
print('Input:', sentences)
print('Ids: {}\nSentences lengths: {}'.format(ids, sent_lens))
# ## Encoder-Decoder architecture
#
# Encoder-Decoder is a successful architecture for Seq2Seq tasks with different lengths of input and output sequences. The main idea is to use two recurrent neural networks, where the first neural network *encodes* the input sequence into a real-valued vector and then the second neural network *decodes* this vector into the output sequence. While building the neural network, we will specify some particular characteristics of this architecture.
import tensorflow as tf
# Let us use TensorFlow building blocks to specify the network architecture.
class Seq2SeqModel(object):
pass
# First, we need to create [placeholders](https://www.tensorflow.org/api_guides/python/io_ops#Placeholders) to specify what data we are going to feed into the network during the execution time. For this task we will need:
# - *input_batch* — sequences of sentences (the shape will equal to [batch_size, max_sequence_len_in_batch]);
# - *input_batch_lengths* — lengths of not padded sequences (the shape equals to [batch_size]);
# - *ground_truth* — sequences of groundtruth (the shape will equal to [batch_size, max_sequence_len_in_batch]);
# - *ground_truth_lengths* — lengths of not padded groundtruth sequences (the shape equals to [batch_size]);
# - *dropout_ph* — dropout keep probability; this placeholder has a predifined value 1;
# - *learning_rate_ph* — learning rate.
def declare_placeholders(self):
"""Specifies placeholders for the model."""
# Placeholders for input and its actual lengths.
self.input_batch = tf.placeholder(shape=(None, None), dtype=tf.int32, name='input_batch')
self.input_batch_lengths = tf.placeholder(shape=(None, ), dtype=tf.int32, name='input_batch_lengths')
# Placeholders for groundtruth and its actual lengths.
self.ground_truth = tf.placeholder(shape=(None, None), dtype=tf.int32, name='ground_truth')
self.ground_truth_lengths = tf.placeholder(shape=(None, ), dtype=tf.int32, name='ground_truth_lengths')
self.dropout_ph = tf.placeholder_with_default(tf.cast(1.0, tf.float32), shape=[])
self.learning_rate_ph = tf.placeholder(dtype=tf.float32, shape=[])
Seq2SeqModel.__declare_placeholders = classmethod(declare_placeholders)
# Now, let us specify the layers of the neural network. First, we need to prepare an embedding matrix. Since we use the same vocabulary for input and output, we need only one such matrix. For tasks with different vocabularies there would be multiple embedding layers.
# - Create embeddings matrix with [tf.Variable](https://www.tensorflow.org/api_docs/python/tf/Variable). Specify its name, type (tf.float32), and initialize with random values.
# - Perform [embeddings lookup](https://www.tensorflow.org/api_docs/python/tf/nn/embedding_lookup) for a given input batch.
def create_embeddings(self, vocab_size, embeddings_size):
"""Specifies embeddings layer and embeds an input batch."""
random_initializer = tf.random_uniform((vocab_size, embeddings_size), -1.0, 1.0)
self.embeddings = tf.Variable(random_initializer, dtype=tf.float32)
# Perform embeddings lookup for self.input_batch.
self.input_batch_embedded = tf.nn.embedding_lookup(self.embeddings, self.input_batch)
Seq2SeqModel.__create_embeddings = classmethod(create_embeddings)
# #### Encoder
#
# The first RNN of the current architecture is called an *encoder* and serves for encoding an input sequence to a real-valued vector. Input of this RNN is an embedded input batch. Since sentences in the same batch could have different actual lengths, we also provide input lengths to avoid unnecessary computations. The final encoder state will be passed to the second RNN (decoder), which we will create soon.
#
# - TensorFlow provides a number of [RNN cells](https://www.tensorflow.org/api_guides/python/contrib.rnn#Core_RNN_Cells_for_use_with_TensorFlow_s_core_RNN_methods) ready for use. We suggest that you use [GRU cell](https://www.tensorflow.org/api_docs/python/tf/contrib/rnn/GRUCell), but you can also experiment with other types.
# - Wrap your cells with [DropoutWrapper](https://www.tensorflow.org/api_docs/python/tf/contrib/rnn/DropoutWrapper). Dropout is an important regularization technique for neural networks. Specify input keep probability using the dropout placeholder that we created before.
# - Combine the defined encoder cells with [Dynamic RNN](https://www.tensorflow.org/api_docs/python/tf/nn/dynamic_rnn). Use the embedded input batches and their lengths here.
# - Use *dtype=tf.float32* everywhere.
def build_encoder(self, hidden_size):
"""Specifies encoder architecture and computes its output."""
# Create GRUCell with dropout.
encoder_cell = tf.nn.rnn_cell.DropoutWrapper(tf.nn.rnn_cell.GRUCell(hidden_size), input_keep_prob=self.dropout_ph)
# Create RNN with the predefined cell.
_, self.final_encoder_state = tf.nn.dynamic_rnn(encoder_cell,
self.input_batch_embedded,
sequence_length=self.input_batch_lengths,
dtype=tf.float32)
Seq2SeqModel.__build_encoder = classmethod(build_encoder)
# #### Decoder
#
# The second RNN is called a *decoder* and serves for generating the output sequence. In the simple seq2seq arcitecture, the input sequence is provided to the decoder only as the final state of the encoder. Obviously, it is a bottleneck and [Attention techniques](https://www.tensorflow.org/tutorials/seq2seq#background_on_the_attention_mechanism) can help to overcome it. So far, we do not need them to make our calculator work, but this would be a necessary ingredient for more advanced tasks.
#
# During training, decoder also uses information about the true output. It is feeded in as input symbol by symbol. However, during the prediction stage (which is called *inference* in this architecture), the decoder can only use its own generated output from the previous step to feed it in at the next step. Because of this difference (*training* vs *inference*), we will create two distinct instances, which will serve for the described scenarios.
#
# The picture below illustrates the point. It also shows our work with the special characters, e.g. look how the start symbol `^` is used. The transparent parts are ignored. In decoder, it is masked out in the loss computation. In encoder, the green state is considered as final and passed to the decoder.
# <img src="encoder-decoder-pic.png" style="width: 500px;">
# Now, it's time to implement the decoder:
# - First, we should create two [helpers](https://www.tensorflow.org/api_guides/python/contrib.seq2seq#Dynamic_Decoding). These classes help to determine the behaviour of the decoder. During the training time, we will use [TrainingHelper](https://www.tensorflow.org/api_docs/python/tf/contrib/seq2seq/TrainingHelper). For the inference we recommend to use [GreedyEmbeddingHelper](https://www.tensorflow.org/api_docs/python/tf/contrib/seq2seq/GreedyEmbeddingHelper).
# - To share all parameters during training and inference, we use one scope and set the flag 'reuse' to True at inference time. You might be interested to know more about how [variable scopes](https://www.tensorflow.org/programmers_guide/variables) work in TF.
# - To create the decoder itself, we will use [BasicDecoder](https://www.tensorflow.org/api_docs/python/tf/contrib/seq2seq/BasicDecoder) class. As previously, you should choose some RNN cell, e.g. GRU cell. To turn hidden states into logits, we will need a projection layer. One of the simple solutions is using [OutputProjectionWrapper](https://www.tensorflow.org/api_docs/python/tf/contrib/rnn/OutputProjectionWrapper).
# - For getting the predictions, it will be convinient to use [dynamic_decode](https://www.tensorflow.org/api_docs/python/tf/contrib/seq2seq/dynamic_decode). This function uses the provided decoder to perform decoding.
def build_decoder(self, hidden_size, vocab_size, max_iter, start_symbol_id, end_symbol_id):
"""Specifies decoder architecture and computes the output.
Uses different helpers:
- for train: feeding ground truth
- for inference: feeding generated output
As a result, self.train_outputs and self.infer_outputs are created.
Each of them contains two fields:
rnn_output (predicted logits)
sample_id (predictions).
"""
# Use start symbols as the decoder inputs at the first time step.
batch_size = tf.shape(self.input_batch)[0]
start_tokens = tf.fill([batch_size], start_symbol_id)
ground_truth_as_input = tf.concat([tf.expand_dims(start_tokens, 1), self.ground_truth], 1)
# Use the embedding layer defined before to lookup embedings for ground_truth_as_input.
self.ground_truth_embedded = tf.nn.embedding_lookup(self.embeddings, ground_truth_as_input)
# Create TrainingHelper for the train stage.
train_helper = tf.contrib.seq2seq.TrainingHelper(self.ground_truth_embedded,
self.ground_truth_lengths)
# Create GreedyEmbeddingHelper for the inference stage.
# You should provide the embedding layer, start_tokens and index of the end symbol.
infer_helper = tf.contrib.seq2seq.GreedyEmbeddingHelper(self.embeddings, start_tokens, end_symbol_id)
def decode(helper, scope, reuse=None):
"""Creates decoder and return the results of the decoding with a given helper."""
with tf.variable_scope(scope, reuse=reuse):
# Create GRUCell with dropout. Do not forget to set the reuse flag properly.
decoder_cell = tf.nn.rnn_cell.DropoutWrapper(tf.nn.rnn_cell.GRUCell(hidden_size, reuse=reuse),
input_keep_prob=self.dropout_ph)
# Create a projection wrapper.
decoder_cell = tf.contrib.rnn.OutputProjectionWrapper(decoder_cell, vocab_size, reuse=reuse)
# Create BasicDecoder, pass the defined cell, a helper, and initial state.
# The initial state should be equal to the final state of the encoder!
decoder = tf.contrib.seq2seq.BasicDecoder(decoder_cell, helper, self.final_encoder_state)
# The first returning argument of dynamic_decode contains two fields:
# rnn_output (predicted logits)
# sample_id (predictions)
outputs, _, _ = tf.contrib.seq2seq.dynamic_decode(decoder=decoder, maximum_iterations=max_iter,
output_time_major=False, impute_finished=True)
return outputs
self.train_outputs = decode(train_helper, 'decode')
self.infer_outputs = decode(infer_helper, 'decode', reuse=True)
Seq2SeqModel.__build_decoder = classmethod(build_decoder)
# In this task we will use [sequence_loss](https://www.tensorflow.org/api_docs/python/tf/contrib/seq2seq/sequence_loss), which is a weighted cross-entropy loss for a sequence of logits. Take a moment to understand, what is your train logits and targets. Also note, that we do not want to take into account loss terms coming from padding symbols, so we will mask them out using weights.
def compute_loss(self):
"""Computes sequence loss (masked cross-entopy loss with logits)."""
weights = tf.cast(tf.sequence_mask(self.ground_truth_lengths), dtype=tf.float32)
self.loss = tf.contrib.seq2seq.sequence_loss(self.train_outputs.rnn_output, self.ground_truth, weights)
Seq2SeqModel.__compute_loss = classmethod(compute_loss)
# The last thing to specify is the optimization of the defined loss.
# We suggest that you use [optimize_loss](https://www.tensorflow.org/api_docs/python/tf/contrib/layers/optimize_loss) with Adam optimizer and a learning rate from the corresponding placeholder. You might also need to pass global step (e.g. as tf.train.get_global_step()) and clip gradients by 1.0.
def perform_optimization(self):
"""Specifies train_op that optimizes self.loss."""
self.train_op = tf.contrib.layers.optimize_loss(loss=self.loss,
global_step=tf.train.get_global_step(),
learning_rate=self.learning_rate_ph,
optimizer='Adam',
clip_gradients=1.0)
Seq2SeqModel.__perform_optimization = classmethod(perform_optimization)
# Congratulations! You have specified all the parts of your network. You may have noticed, that we didn't deal with any real data yet, so what you have written is just recipies on how the network should function.
# Now we will put them to the constructor of our Seq2SeqModel class to use it in the next section.
def init_model(self, vocab_size, embeddings_size, hidden_size,
max_iter, start_symbol_id, end_symbol_id, padding_symbol_id):
self.__declare_placeholders()
self.__create_embeddings(vocab_size, embeddings_size)
self.__build_encoder(hidden_size)
self.__build_decoder(hidden_size, vocab_size, max_iter, start_symbol_id, end_symbol_id)
# Compute loss and back-propagate.
self.__compute_loss()
self.__perform_optimization()
# Get predictions for evaluation.
self.train_predictions = self.train_outputs.sample_id
self.infer_predictions = self.infer_outputs.sample_id
Seq2SeqModel.__init__ = classmethod(init_model)
# ## Train the network and predict output
#
# [Session.run](https://www.tensorflow.org/api_docs/python/tf/Session#run) is a point which initiates computations in the graph that we have defined. To train the network, we need to compute *self.train_op*. To predict output, we just need to compute *self.infer_predictions*. In any case, we need to feed actual data through the placeholders that we defined above.
def train_on_batch(self, session, X, X_seq_len, Y, Y_seq_len, learning_rate, dropout_keep_probability):
feed_dict = {
self.input_batch: X,
self.input_batch_lengths: X_seq_len,
self.ground_truth: Y,
self.ground_truth_lengths: Y_seq_len,
self.learning_rate_ph: learning_rate,
self.dropout_ph: dropout_keep_probability
}
pred, loss, _ = session.run([
self.train_predictions,
self.loss,
self.train_op], feed_dict=feed_dict)
return pred, loss
Seq2SeqModel.train_on_batch = classmethod(train_on_batch)
# We implemented two prediction functions: *predict_for_batch* and *predict_for_batch_with_loss*. The first one allows only to predict output for some input sequence, while the second one could compute loss because we provide also ground truth values. Both these functions might be useful since the first one could be used for predicting only, and the second one is helpful for validating results on not-training data during the training.
# +
def predict_for_batch(self, session, X, X_seq_len):
feed_dict = {
self.input_batch: X,
self.input_batch_lengths: X_seq_len
}
pred = session.run([
self.infer_predictions
], feed_dict=feed_dict)[0]
return pred
def predict_for_batch_with_loss(self, session, X, X_seq_len, Y, Y_seq_len):
feed_dict = {
self.input_batch: X,
self.input_batch_lengths: X_seq_len,
self.ground_truth: Y,
self.ground_truth_lengths: Y_seq_len
}
pred, loss = session.run([
self.infer_predictions,
self.loss,
], feed_dict=feed_dict)
return pred, loss
# -
Seq2SeqModel.predict_for_batch = classmethod(predict_for_batch)
Seq2SeqModel.predict_for_batch_with_loss = classmethod(predict_for_batch_with_loss)
# ## Run your experiment
#
# Create *Seq2SeqModel* model with the following parameters:
# - *vocab_size* — number of tokens;
# - *embeddings_size* — dimension of embeddings, recommended value: 20;
# - *max_iter* — maximum number of steps in decoder, recommended value: 7;
# - *hidden_size* — size of hidden layers for RNN, recommended value: 512;
# - *start_symbol_id* — an index of the start token (`^`).
# - *end_symbol_id* — an index of the end token (`$`).
# - *padding_symbol_id* — an index of the padding token (`#`).
#
# Set hyperparameters. You might want to start with the following values and see how it works:
# - *batch_size*: 128;
# - at least 10 epochs;
# - value of *learning_rate*: 0.001
# - *dropout_keep_probability* equals to 0.5 for training (typical values for dropout probability are ranging from 0.1 to 1.0); larger values correspond smaler number of dropout units;
# - *max_len*: 20.
# +
tf.reset_default_graph()
model = Seq2SeqModel(vocab_size=len(word2id),
embeddings_size=20,
hidden_size=512,
max_iter=7,
start_symbol_id=word2id['^'],
end_symbol_id=word2id['$'],
padding_symbol_id=word2id['#'])
batch_size = 128
n_epochs = 10
learning_rate = 0.001
dropout_keep_probability = 0.5
max_len = 20
n_step = int(len(train_set) / batch_size)
# -
# Finally, we are ready to run the training! A good indicator that everything works fine is decreasing loss during the training. You should account on the loss value equal to approximately 2.7 at the beginning of the training and near 1 after the 10th epoch.
# +
session = tf.Session()
session.run(tf.global_variables_initializer())
invalid_number_prediction_counts = []
all_model_predictions = []
all_ground_truth = []
print('Start training... \n')
for epoch in range(n_epochs):
random.shuffle(train_set)
random.shuffle(test_set)
print('Train: epoch', epoch + 1)
for n_iter, (X_batch, Y_batch) in enumerate(generate_batches(train_set, batch_size=batch_size)):
######################################
######### YOUR CODE HERE #############
######################################
# prepare the data (X_batch and Y_batch) for training
# using function batch_to_ids
X, X_seq_len = batch_to_ids(X_batch, word2id, max_len)
Y, Y_seq_len = batch_to_ids(Y_batch, word2id, max_len)
predictions, loss = model.train_on_batch(session, X, X_seq_len, Y, Y_seq_len, learning_rate, dropout_keep_probability)
if n_iter % 200 == 0:
print("Epoch: [%d/%d], step: [%d/%d], loss: %f" % (epoch + 1, n_epochs, n_iter + 1, n_step, loss))
X_sent, Y_sent = next(generate_batches(test_set, batch_size=batch_size))
######################################
######### YOUR CODE HERE #############
######################################
# prepare test data (X_sent and Y_sent) for predicting
# quality and computing value of the loss function
# using function batch_to_ids
X, X_seq_len = batch_to_ids(X_sent, word2id, max_len)
Y, Y_seq_len = batch_to_ids(Y_sent, word2id, max_len)
predictions, loss = model.predict_for_batch_with_loss(session, X, X_seq_len, Y, Y_seq_len)
print('Test: epoch', epoch + 1, 'loss:', loss,)
for x, y, p in list(zip(X, Y, predictions))[:3]:
print('X:',''.join(ids_to_sentence(x, id2word)))
print('Y:',''.join(ids_to_sentence(y, id2word)))
print('O:',''.join(ids_to_sentence(p, id2word)))
print('')
model_predictions = []
ground_truth = []
invalid_number_prediction_count = 0
# For the whole test set calculate ground-truth values (as integer numbers)
# and prediction values (also as integers) to calculate metrics.
# If generated by model number is not correct (e.g. '1-1'),
# increase invalid_number_prediction_count and don't append this and corresponding
# ground-truth value to the arrays.
for X_batch, Y_batch in generate_batches(test_set, batch_size=batch_size):
######################################
######### YOUR CODE HERE #############
######################################
X, X_seq_len = batch_to_ids(X_batch, word2id, max_len)
Y, Y_seq_len = batch_to_ids(Y_batch, word2id, max_len)
predictions = model.predict_for_batch(session, X, X_seq_len)
for y, p in zip(Y, predictions):
y_sent = ''.join(ids_to_sentence(y, id2word))
y_sent = y_sent[:y_sent.find('$')]
p_sent = ''.join(ids_to_sentence(p, id2word))
index = p_sent.find('$')
if index != -1:
p_sent = p_sent[:index]
if p_sent.isdigit() or (p_sent.startswith('-') and p_sent[1:].isdigit()):
model_predictions.append(int(p_sent))
ground_truth.append(int(y_sent))
else:
invalid_number_prediction_count += 1
all_model_predictions.append(model_predictions)
all_ground_truth.append(ground_truth)
invalid_number_prediction_counts.append(invalid_number_prediction_count)
print('\n...training finished.')
# -
# ## Evaluate results
#
# Because our task is simple and the output is straight-forward, we will use [MAE](https://en.wikipedia.org/wiki/Mean_absolute_error) metric to evaluate the trained model during the epochs. Compute the value of the metric for the output from each epoch.
from sklearn.metrics import mean_absolute_error
for i, (gts, predictions, invalid_number_prediction_count) in enumerate(zip(all_ground_truth,
all_model_predictions,
invalid_number_prediction_counts), 1):
mae = mean_absolute_error(gts, predictions)
print("Epoch: %i, MAE: %f, Invalid numbers: %i" % (i, mae, invalid_number_prediction_count))
| Natural Language Processing/Week4/week4-seq2seq.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# This plots the epochs at which RR Lyrae spectra were taken
# created 2017 Dec 19 by E.S.
# -
import astropy
from astropy.io import fits
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from os import listdir
from os.path import isfile, join
from datetime import datetime
from astropy import time, coordinates as coord, units as u
from astropy.time import Time
from dateutil.parser import parse
stem = "/Users/bandari/Documents/git.repos/rrlyrae_metallicity/" + \
"rrlyrae_metallicity/src/mcdonald_spectra/original_fits_files/"
import os
arr_files = os.listdir(stem) # list files
# +
# read in each FITS file in the directory and obtain epoch from header
fileList_2012 = []
dateList_2012 = []
fileList_2013 = []
dateList_2013 = []
epochList_2012 = []
epochList_2013 = []
for f in range(0,len(arr_files)): # loop over filenames
# retrieve header
image, header = fits.getdata(stem+arr_files[f],
0,
header=True)
# observation epoch
epoch = header['DATE-OBS']+' '+header['UT']
# parse
epoch_dateTime = datetime.strptime(epoch, '%Y-%m-%d %H:%M:%S.%f')
if (epoch_dateTime.year == 2012):
fileList_2012.append(arr_files[f])
dateList_2012.append(epoch_dateTime)
epochList_2012.append(epoch)
else:
fileList_2013.append(arr_files[f])
dateList_2013.append(epoch_dateTime)
epochList_2013.append(epoch)
# +
ut_2012 = {'file':fileList_2012,'ut_epoch':epochList_2012}
df_ut_2012 = pd.DataFrame(ut_2012)
# print dataframe.
df_ut_2012
ut_2013 = {'file':fileList_2013,'ut_epoch':epochList_2013}
df_ut_2013 = pd.DataFrame(ut_2013)
# print dataframe.
df_ut_2013
df_ut_2012.to_csv("mcd_2012_run.csv")
df_ut_2013.to_csv("mcd_2013_run.csv")
# +
# set observatory coordinates
loc_mcdonald = coord.EarthLocation.from_geodetic(lon=-104.0215753,lat=30.6715396,height=2076,ellipsoid='WGS84')
loc_macadam = coord.EarthLocation.from_geodetic(lon=-84.503712,lat=38.033891,height=298,ellipsoid='WGS84')
# +
# convert UTC times to isot format, then compile into list of astropy Time object
t_spectra_2012_iso = [Time(dateList_2012[i].isoformat(), format='isot', scale='utc') for i in range(len(dateList_2012))]
t_spectra_2013_iso = [Time(dateList_2013[i].isoformat(), format='isot', scale='utc') for i in range(len(dateList_2013))]
# +
# convert isot-format times to MJD or JD (Astropy seems to make more accurate conversion from MJD->BJD)
t_spectra_2012_mjd = [t_spectra_2012_iso[i].mjd for i in range(len(t_spectra_2012_iso))]
t_spectra_2013_mjd = [t_spectra_2013_iso[i].mjd for i in range(len(t_spectra_2013_iso))]
t_spectra_2012_jd = [t_spectra_2012_iso[i].jd for i in range(len(t_spectra_2012_iso))]
t_spectra_2013_jd = [t_spectra_2013_iso[i].jd for i in range(len(t_spectra_2013_iso))]
# +
# fcn to convert MJD to BJD times
def convert_mjd_to_bjd(mjdTimes,observatoryLoc,skyCoordObj):
timesObj = time.Time(mjdTimes, format='mjd', scale='utc', location=observatoryLoc)
ltt_bary = timesObj.light_travel_time(skyCoordObj)
time_barycentre = timesObj.tdb + ltt_bary
# note the returned type is still mendaciously called 'mjd'
return np.add(time_barycentre.mjd,0.5) # I think 0.5 day is missing from the code
# +
# fcn to convert MJD to HJD times
def convert_mjd_to_hjd(mjdTimes,observatoryLoc,skyCoordObj):
timesObj = time.Time(mjdTimes, format='mjd', scale='utc', location=observatoryLoc)
ltt_helio = timesObj.light_travel_time(skyCoordObj, 'heliocentric')
times_heliocentre = timesObj.utc + ltt_helio
# note the returned type is still mendaciously called 'mjd'
return np.add(times_heliocentre.mjd,0.5) # I think 0.5 day is missing from the code
# +
# combine all data for across-the-board comparison
allFileList = np.hstack((fileList_2012,fileList_2013))
allSpecEpochList_utc = np.hstack((dateList_2012,dateList_2013))
allSpecEpochList_mjd = np.hstack((t_spectra_2012_mjd,t_spectra_2013_mjd))
allSpecEpochList_jd = np.hstack((t_spectra_2012_jd,t_spectra_2013_jd))
#allSpecEpochList_bjd = np.hstack((t_spectra_2012_bjd,t_spectra_2013_bjd))
# -
allSpecEpochList_utc
# +
# read in star name, return file names and BJDs of spectra observations
def return_star_bjds(fileNames,mjdTimes,starNameFile,starNameGeneric,observatoryLoc):
if len(fileNames) != len(mjdTimes): # something must be wrong!
return
# initialize a pandas dataframe
df = pd.DataFrame()
# make a star coordinate object
coord_star = coord.SkyCoord.from_name(starNameGeneric)
fileNamesThisStar = []
mjdsThisStar = []
hjdsThisStar = []
bjdsThisStar = []
for t in range(0,len(fileNames)):
if starNameFile in fileNames[t]:
print(fileNames[t])
fileNamesThisStar.append(fileNames[t])
mjdsThisStar.append(mjdTimes[t])
hjdsThisStar.append(convert_mjd_to_hjd(mjdTimes[t],observatoryLoc,coord_star))
bjdsThisStar.append(convert_mjd_to_bjd(mjdTimes[t],observatoryLoc,coord_star))
#bjdsThisStar = np.add(bjdsThisStar,0.5) # I think 0.5 day is missing from the code
elapsed_bjd = np.subtract(bjdsThisStar,np.min(bjdsThisStar))
df['filenames'] = fileNamesThisStar
df['mjd'] = mjdsThisStar
df['hjd'] = hjdsThisStar
df['bjd'] = bjdsThisStar
df['elapsed_bjd_since_spec_01'] = elapsed_bjd
return df
# +
# star names as they appear in the filenames
star_names_files = ['RW_Ari','X_Ari','UY_Cam','RR_Cet','SV_Eri',
'VX_Her','RR_Leo','TT_Lyn','TV_Lyn','TW_Lyn',
'RR_Lyr','V_535','V445','AV_Peg','BH_Peg',
'AR_Per','RU_Psc','T_Sex','TU_UMa']
# star names for SIMBAD lookup
star_names_simbad = ['RW Ari','X Ari','UY Cam','RR Cet','SV Eri',
'VX Her','RR Leo','TT Lyn','TV Lyn','TW Lyn',
'RR Lyr','V535 Mon','V445 Oph','AV Peg','BH Peg',
'AR Per','RU Psc','T Sex','TU UMa']
# -
allSpecEpochList_mjd
# +
# find BJDs and concatenate everything into one dataframe
dfAll = pd.DataFrame()
for star in range(0,len(star_names_files)):
df_thisStar = return_star_bjds(allFileList,allSpecEpochList_mjd,star_names_files[star],star_names_simbad[star],loc_mcdonald)
dfAll = pd.concat([dfAll,df_thisStar])
# +
# write filenames, spectra epochs out csv
dfAll.to_csv('junk.csv')
# +
# print filenames, spectra epochs
dfAll
# +
########################################
## MAKE PLOTS
########################################
# +
# fcn for generating a plot to visualize epochs
def spec_epoch_plot(fileArray,epochArray,plotName):
fig, ax = plt.subplots()
fig.set_size_inches(200, 10)
ax.scatter(epochArray, np.ones(len(epochArray)))
#textPos = np.max(spectData.flux)+0.01
[ax.text(epochArray[i], 2.5, fileArray[i], rotation='vertical') for i in range(len(epochArray))]
ax.set_ylim([0,3.6])
ax.set_xlim([np.min(epochArray),np.max(epochArray)])
plt.savefig(plotName)
# +
# write out plots of when spectra were observed
spec_epoch_plot(fileList_2012,dateList_2012,'test_2012.pdf')
spec_epoch_plot(fileList_2013,dateList_2013,'test_2013.pdf')
| notebooks_for_development/find_spec_epochs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
from IPython.display import display
import optoanalysis
ConvFactor = 1.8e5
dat = optoanalysis.sim_data.SimData([0, 0.1], 5e6, [75e3, 160e3, 180e3], 4000.0, 3.1e-19, ConvFactor, 1e-30, dt=1e-9)
# +
# tArray = dat.sde_solvers[0].tArray.get_array()
# fig,ax = plt.subplots()
# ax.plot(tArray, dat.sde_solvers[0].q)
# ax.plot(tArray, dat.sde_solvers[1].q)
# ax.plot(tArray, dat.sde_solvers[2].q)
# -
dat.plot_PSD([0, 300])
# +
# dat.extract_ZXY_motion?
# -
z, x, y, t, fig, ax = dat.extract_ZXY_motion([75e3, 160e3, 180e3], 3e3, [10e3, 5e3, 5e3], subSampleFraction=2)
t, z, vz = dat.calc_phase_space(75e3, ConvFactor, FractionOfSampleFreq=2)
t, x, vx = dat.calc_phase_space(160e3, ConvFactor, FractionOfSampleFreq=2)
t, y, vy = dat.calc_phase_space(180e3, ConvFactor, FractionOfSampleFreq=2)
TimeTuple = [500e-6, 1000e-6]
alpha=0.8
plt.plot(t, z, alpha=alpha)
plt.plot(dat.simtime.get_array(), dat.TrueSignals[0, 0], alpha=alpha)
plt.xlim(TimeTuple)
plt.plot(t[1:], vz, alpha=alpha)
plt.plot(dat.simtime.get_array(), dat.TrueSignals[0, 1], alpha=alpha)
plt.xlim(TimeTuple)
plt.plot(t, x, alpha=alpha)
plt.plot(dat.simtime.get_array(), dat.TrueSignals[1, 0], alpha=alpha)
plt.xlim(TimeTuple)
plt.plot(t[1:], vx, alpha=alpha)
plt.plot(dat.simtime.get_array(), dat.TrueSignals[1, 1], alpha=alpha)
plt.xlim(TimeTuple)
plt.plot(t, y, alpha=alpha)
plt.plot(dat.simtime.get_array(), dat.TrueSignals[2, 0], alpha=alpha)
plt.xlim(TimeTuple)
plt.plot(t[1:], vy, alpha=alpha)
plt.plot(dat.simtime.get_array(), dat.TrueSignals[2, 1], alpha=alpha)
plt.xlim(TimeTuple)
dat.get_fit_auto(75e3)
dir(dat)
| sim_data_usage_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from scipy import signal
from bokeh.plotting import figure, output_file, show
from bokeh.io import output_notebook
from bokeh.layouts import row
import numpy as np
import math
import sympy as sy
output_notebook()
# + code_folding=[0, 7, 42]
def lti_to_sympy(lsys, symplify=True):
""" Convert Scipy's LTI instance to Sympy expression """
s = sy.Symbol('s')
G = sy.Poly(lsys.num, s) / sy.Poly(lsys.den, s)
return sy.simplify(G) if symplify else G
def sympy_to_lti(xpr, s=sy.Symbol('s')):
""" Convert Sympy transfer function polynomial to Scipy LTI """
num, den = sy.simplify(xpr).as_numer_denom() # expressions
p_num_den = sy.poly(num, s), sy.poly(den, s) # polynomials
c_num_den = [sy.expand(p).all_coeffs() for p in p_num_den] # coefficients
l_num, l_den = [sy.lambdify((), c)() for c in c_num_den] # convert to floats
return signal.lti(l_num, l_den)
def polyphase_iir_to_ba_coeffs(directCoeffs,delayedCoeffs):
#For z^-1:
z_1 = sy.Symbol('z_1')
directPath = 1
delayedPath = 1
for c in directCoeffs:
directPath = directPath * (sy.Poly([1,0,c], z_1) / sy.Poly([c,0,1], z_1))
for c in delayedCoeffs:
delayedPath = delayedPath * (sy.Poly([1,0,c], z_1) / sy.Poly([c,0,1], z_1))
tranferFunction = 0.5 *( directPath + sy.Poly([1,0], z_1) * delayedPath)
n,d = sy.fraction(sy.simplify(tranferFunction))
b = [float(c) for c in sy.Poly(n).all_coeffs()]
a = [float(c) for c in sy.Poly(d).all_coeffs()]
return b,a
""" Adapted from:
Digital signal processing schemes for efficient interpolation and decimation
<NAME>, <NAME>
IEEE 1983
transitionBandwitdth between 0 and 0.5 ( 0.5 = half nyquist)
stopbandAttenuation in dB
"""
def design_polyphase_halfband_iir(transitionBandwidth, stopbandAttenuation):
k = np.tan((np.pi-2*np.pi*transitionBandwidth)/4)**2
kp = np.sqrt(1-k**2)
e = 0.5 * (1 -np.sqrt(kp))/(1+np.sqrt(kp))
q = e + 2*e**5 + 15*e**9 + 150*e**13
ds = 10**(-stopbandAttenuation/20)
k1 = ds**2 / (1 - ds**2)
n = int(math.ceil(math.log(k1**2/16)/math.log(q)))
if n % 2 ==0:
n += 1
if n ==1 :
n = 3
print("Order: %d" % n)
q1 = q**n
k1 = 4 * math.sqrt(q1)
ds = math.sqrt(k1 / (1+k1))
dp = 1 - math.sqrt(1-ds**2)
def ellipticSum1(q,n,i):
s = 0
for m in range(5):
s += (-1)**m *q**(m*(m+1)) * math.sin((2*m+1)*(math.pi*i)/n)
return s
def ellipticSum2(q,n,i):
s = 0
for m in range(1,5):
s += (-1)**m *q**(m*m) * math.cos(2*m*(math.pi*i)/n)
return s
wi = [ 2*math.pow(q,0.25) * ellipticSum1(q,n,i)/(1+2*ellipticSum2(q,n,i)) for i in range(1,int((n-1)/2)+1) ]
ai = [ math.sqrt((1-(w**2)*k)*(1-(w**2)/k))/(1+w**2) for w in wi ]
ai = [ (1-a)/(1+a) for a in ai ]
#print(ai)
return ai[0::2], ai[1::2]
# + code_folding=[0]
def plot_filter(b,a,name):
p1 = figure(plot_width = 500, plot_height=300, title = "%s Gain" % name, y_range = (-80,6))
p1.xaxis.axis_label = 'Frequency [rad/sample]'
p1.yaxis.axis_label = 'Amplitude [dB]'
w, h = signal.freqz(b,a)
p1.line(w, 20 * np.log10(abs(h)))
w, gd = signal.group_delay((b, a))
p2 = figure(plot_width = 500, plot_height=300, title = "%s Group delay" % name)
p2.xaxis.axis_label = 'Frequency [rad/sample]'
p2.yaxis.axis_label = 'Group delay [samples]'
p2.line(w,gd)
show(row(p1,p2))
# -
#b, a = signal.iirdesign(0., 0.3, 5, 50, ftype='cheby1')
b,a = signal.cheby1(5,1,0.5)
np.set_printoptions(precision=16)
#print(a)
#print(b)
plot_filter(b,a,'Cheby1')
b,a = signal.butter(7,0.5)
plot_filter(b,a,'Butterworth')
b,a = signal.cheby2(5,40,0.5)
plot_filter(b,a,'Cheby2')
b,a = polyphase_iir_to_ba_coeffs([1.0 / (5 + 2 * np.sqrt(5))],[5-2*np.sqrt(5)])
plot_filter(b,a,'ButterPolyphase')
int(math.ceil(0.9))
directCoeffs,delayedCoeffs = design_polyphase_halfband_iir(0.1, 30)
print(directCoeffs)
print(delayedCoeffs)
b,a = polyphase_iir_to_ba_coeffs(directCoeffs,delayedCoeffs)
plot_filter(b,a,'Cheby-ish polyphase')
[i for i in range(1,int((7-1)/2)+1)]
[1.0 / (5 + 2 * np.sqrt(5))],[5-2*np.sqrt(5)]
| tests/FiltersTests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Run SQL Workloads
#
# This Jupyter Notebook contains code to run SQL workloads across databases.
#
# The Db2 Data Management Console is more than a graphical user interface. It is a set of microservices that you can use to build custom applications to automate your use of Db2.
#
# This Jupyter Notebook contains examples of how to use the Open APIs and the composable interface that are available in the Db2 Data Management Console. Everything in the User Interface is also available through an open and fully documented RESTful Services API. The full set of APIs are documented as part of the Db2 Data Management Console user interface. In this hands on lab you can connect to the documentation directly through this link: [Db2 Data Management Console RESTful APIs](http://localhost:11080/dbapi/api/index_enterprise.html).
#
# You can also embed elements of the full user interface into an IFrame by constructing the appropriate URL.
#
# This hands on lab will be calling the Db2 Data Management Console as a service. However you can explore it through the user interface as well. Just click on the following link to try out the console that is already and setup in this lab: http://localhost:11080/console. If you have not already logged in you can use the following:
# * Userid: db2inst1
# * Password: <PASSWORD>
# ### Import Helper Classes
# For more information on these classes, see the lab: [Automate Db2 with Open Console Services](http://localhost:8888/notebooks/Db2_Data_Management_Console_Overview.ipynb)
# %run ./dmc_setup.ipynb
# ### Db2 Data Management Console Connection
# To connect to the Db2 Data Management Console service you need to provide the URL, the service name (v4) and profile the console user name and password as well as the name of the connection profile used in the console to connect to the database you want to work with. For this lab we are assuming that the following values are used for the connection:
# * Userid: db2inst1
# * Password: <PASSWORD>
# * Connection: sample
#
# **Note:** If the Db2 Data Management Console has not completed initialization, the connection below will fail. Wait for a few moments and then try it again.
# +
# Connect to the Db2 Data Management Console service
Console = 'http://localhost:11080'
profile = 'SAMPLE'
user = 'DB2INST1'
password = '<PASSWORD>'
# Set up the required connection
profileURL = "?profile="+profile
databaseAPI = Db2(Console+'/dbapi/v4')
if databaseAPI.authenticate(user, password, profile) :
print("Token Created")
else :
print("Token Creation Failed")
database = Console
# -
# ### Confirm the connection
# To confirm that your connection is working you can check the status of the moitoring service. You can also check your console connection to get the details of the specific database connection you are working with. Since your console user id and password may be limited as to which databases they can access you need to provide the connection profile name to drill down on any detailed information for the database.
# List Monitoring Profile
r = databaseAPI.getProfile(profile)
json = databaseAPI.getJSON(r)
print(json)
# ### SQL Scripts Used to Generate Work
# We are going to define a few scripts that we will use during this lab.
sqlScriptWorkload1 = \
'''
WITH SALARYBY (DEPTNO, TOTAL) AS
(SELECT DEPT.DEPTNO DNO, SUM(BIGINT(EMP.SALARY)) TOTAL_SALARY
FROM EMPLOYEES EMP, DEPARTMENTS DEPT
WHERE DEPT.DEPTNO = EMP.DEPTNO AND EMP.SALARY > 190000
GROUP BY DEPT.DEPTNO
ORDER BY DNO)
SELECT DEPT.DEPTNAME NAME, SALARYBY.TOTAL COST, DEPT.REVENUE, DEPT.REVENUE-SALARYBY.TOTAL PROFIT
FROM SALARYBY, DEPARTMENTS DEPT
WHERE DEPT.DEPTNO = SALARYBY.DEPTNO
AND REVENUE > TOTAL
ORDER BY PROFIT
'''
print("Defined Workload 1 Script")
sqlScriptWorkload2 = \
'''
SELECT DEPT.DEPTNO DNO, SUM(FLOAT(EMP.SALARY)) TOTAL_SALARY
FROM EMPLOYEES EMP, DEPARTMENTS DEPT
WHERE DEPT.DEPTNO = EMP.DEPTNO
AND EMP.SALARY < 50000
AND YEAR(EMP.HIREDATA) > 2010
GROUP BY DEPT.DEPTNO
ORDER BY DNO;
SELECT DEPT.DEPTNO DNO, SUM(FLOAT(EMP.SALARY)) TOTAL_SALARY
FROM EMPLOYEES EMP, DEPARTMENTS DEPT
WHERE DEPT.DEPTNO = EMP.DEPTNO
AND EMP.SALARY < 190000
AND YEAR(EMP.HIREDATA) > 2010
GROUP BY DEPT.DEPTNO
ORDER BY DNO;
SELECT DEPT.DEPTNO, DEPT.REVENUE
FROM DEPARTMENTS DEPT WHERE DEPT.REVENUE > 450000000;
'''
print("Defined Workload 2 Script")
# ### Creating a Routine to Run an SQL Script
# To make things easier we can create reusable routines that will included everything we have developed so far. By running the next two steps, you create two routines that you can call by passing parameters to them.
#
# While we could create a single routine to run SQL and then display the results, we are creating two different routines so that we can display the results differently later in the lab.
def runSQL(profile,user, password, sqlText):
if databaseAPI.authenticate(user, password, profile) :
# Run the SQL Script and return the runID for later reference
runID = databaseAPI.getJSON(databaseAPI.runSQL(sqlText))['id']
# See if there are any results yet for this job
json = databaseAPI.getJSON(databaseAPI.getSQLJobResult(runID))
# If the REST call returns an error return the json with the error to the calling routine
if 'errors' in json :
return json
# Append the results from each statement in the script to the overall combined JSON result set
fulljson = json
while json['results'] != [] or (json['status'] != "completed" and json['status'] != "failed") :
json = databaseAPI.getJSON(databaseAPI.getSQLJobResult(runID))
# Get the results from each statement as they return and append the results to the full JSON
for results in json['results'] :
fulljson['results'].append(results)
# Wait 250 ms for more results
time.sleep(0.25)
return fulljson
else :
print('Could not authenticate')
print('runSQL routine defined')
def displayResults(json):
for results in json['results']:
print('Statement: '+str(results['index'])+': '+results['command'])
print('Runtime ms: '+str(results['runtime_seconds']*1000))
if 'error' in results :
print(results['error'])
elif 'rows' in results :
df = pd.DataFrame(results['rows'],columns=results['columns'])
print(df)
else :
print('No errors. Row Affected: '+str(results['rows_affected']))
print()
print('displayResults routine defined')
# ### Running multiple scripts across multiple databases - Summarized Results
# Now that we have our tables created on both databases, we can run workloads and measure their performance. By repeatedly running the scripts across multiple databases in a single Db2 instance we can simulate a real database environemt.
#
# Instead of using the displayResults routine we are going to capture runtime metrics for each run of the SQL Query workloads so that we can analyze performance. The appendResults routine builds this dataframe with each pass.
#
# runScripts lets use define the database connection profiles we want to run against, the scripts to run, and now many times to repeat the runs for each profile and for each script.
# This routine builds up a Data Frame containing the run results as we run workloads across databases
def appendResults(df, profile, json) :
error = ''
rows = 0
if 'error' in json :
print('SQL Service Failed')
else :
for results in json['results']:
if 'error' in results :
error = results['error']
if 'rows_affected' in results :
rows = results['rows_affected']
df = df.append({'profile':profile,'index':results['index'], 'statement':results['command'], 'error':error, 'rows_affected': rows, 'runtime_ms':(results['runtime_seconds']*1000)}, ignore_index=True)
return df
print('appendResults routine defined')
# This routine runs multistatment scripts across multiple databases.
# The scripts are run repeatedly for each profile (database)
def runScripts(profileList, scriptList, user, password, profileReps, scriptReps) :
df = pd.DataFrame(columns=['profile', 'index', 'statement', 'error', 'rows_affected', 'runtime_ms'])
for x in range(0, profileReps):
print("Running repetition: "+str(x))
for profile in profileList :
print(" Running scripts against: "+profile)
for y in range(0, scriptReps) :
print(" Running script repetition: "+str(y))
for script in scriptList :
json = runSQL(profile, user, password, script)
while 'errors' in json:
print(' * Trying again *')
json = runSQL(profile, user, password, script)
df = appendResults(df, profile, json)
return df
print('runScripts routine defined')
# The next cell loops through a list of databases as well as a list of scripts and run then repeatedly. You an set the number of times the scripts are run against each database and the number of times the runs against both databases is repeated.
# +
profileList = ['SAMPLE','HISTORY']
scriptList = [sqlScriptWorkload1, sqlScriptWorkload2]
user = 'DB2INST1'
password = '<PASSWORD>'
profileReps = 20
scriptReps = 5
df = runScripts(profileList, scriptList, user, password, profileReps, scriptReps)
display(df)
# -
# ### Analyze Results
# Now we can use the results in the dataframe to look at the results statistically. First we can see the average runtime for each statement across the databases.
print('Mean runtime in ms')
pd.set_option('display.max_colwidth', 100)
stmtMean = df.groupby(['statement']).mean()
print(stmtMean)
# We can also display the total runtime for each statement across databases.
print('Total runtime in ms')
pd.set_option('display.max_colwidth', 100)
stmtSum = df.groupby(['statement']).sum()
print(stmtSum)
# We can even graph the total run time for all the statements can compare database performance. Since there are more rows in the employees table in the SAMPLE database it takes longer for the queries to run.
print('Mean runtime in ms')
pd.set_option('display.max_colwidth', 100)
profileSum = df.groupby(['profile']).sum()
profileSum.plot(kind='bar')
plt.show()
# #### Credits: IBM 2019, <NAME> [<EMAIL>]
| Db2_Data_Management_Console_Run_Workload_One.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pylab inline
# %config InlineBackend.figure_format = 'retina'
from ipywidgets import interact
# # Question 1
# # A
# Consider the function $f(x)=x^3 -9x^2 +11x-11$, which has a root in the interval $(-10,10)$. Calculate by hand the first 3 iterations of Newton's method to estimate the root of $f(x)$, using the initial guess $x_0=0$. What happens?
# ---------------------------------------
# ## Solution
# Starting with the initial guess $x_0=0$, the first three iterations using Newton's method are
# \begin{align}
# x_1 &= 0 - \frac{-11}{11} = 1 \\
# x_2 &= 1 - \frac{1^3-9\cdot1^2+11\cdot1-11}{3\cdot1^2-18\cdot1+11} = 1-\frac{-8}{-4} = -1 \\
# x_3 &= -1 - \frac{(-1)^3-9(-1)^2+11(-1)-11}{3(-1)^2-18(-1)+11} = -1 - \frac{-32}{32} = 0 = x_0
# \end{align}
# Computing the first three iterates we find that Newton's method with $x_0=0$ results in a cycle with period 3.
# # B
# Write a python code that uses bisection to determine a better choice of $x_0$, then feed this new initial guess to a Newton's method solver. Iterate your Newton's method solver until you have found the root of $f(x)$ to within a tolerance of $\epsilon=10^{-6}$, and report the value of the root.
# +
# ====== bisection method followed by Newton's method ======
def f(x):
return x**3 - 9*x**2 + 11*x - 11
def df(x):
return 3*x**2 - 18*x + 11
# ------ endpoints, max iterations, and tolerance ------
a=-10 #left endpoint
b=10 #right endpoint
N=25 #max iterations for bisection & newton's
tol=(b-1)/2 #very broad tolerance for bisection. NOTE: this was chosen aftercomputing bisection by hand
#until the subinterval no longer contained -1, 0, or 1. See text below
# ------ bisection (Burden & Faires Numerical Analysis 9th ed) ------
i=1
FA=f(a)
while i<=N:
p = a + (b-a)/2
FP=f(p)
if FP==0 or (b-a)/2<tol: break #return p=x0 for newton's
i+=1
if FA*FP>0:
a=p
FA=FP
else:
b=p
print('Bisection complete. Using x0 =',p,'as initial guess for Newton\'s method')
# ------ newton's method adapted from Lab 2 ------
tol=10**(-6) #update tolerance
x=p #switch from p to x so notation is familiar
# ------ compute x_1 directly so we can do 1st comparison ------
xn = x - f(x)/df(x)
i=1 #initialize an iteration counter
while abs(xn-x)>tol and i<=N:
x=xn
xn = x - f(x)/df(x)
i+=1
print('Convergence for Newton\'s method was achieved using',i,'iterations')
print('The root of f(x) is',x)
# -
# # Question 2
# # A
# Derive a third order method for solving $f(x) = 0$ in a way similar to the derivation of Newton’s method, using evaluations of $f(x_n)$, $f’(x_n)$, and $f’’(x_n)$. Show that in the course of derivation, a quadratic equation arises, and therefore two distinct schemes can be derived. **Hint: Expand $f(x)$ around $x_n$.**
# ---------------------------------------
# ## Solution
# Expand $f(x)$ around $x_n$ to get
# $$ f(x) = f(x_n) + (x - x_n)f'(x_n) + \frac{(x - x_n)^2}{2}f''(x_n) + O((x - x_n)^3). $$
# Set $x = \hat{x}$ and use $f(\hat{x})=0$ to get
# $$ 0 = f(x_n) + (\hat{x} - x_n)f'(x_n) + \frac{(\hat{x} - x_n)^2}{2}f''(x_n) + O((\hat{x} - x_n)^3). $$
# Solving for $\hat{x}$, we have
# $$\frac{1}{2}f''(x_n) \hat{x}^2 + (f'(x_n) - x_n f''(x_n))\hat{x} + f(x_n) - x_n f'(x_n) + \frac{x_n^2}{2}f''(x_n) = O((\hat{x} - x_n)^3).$$
# Setting the right hand side to zero, we find two solutions for $\hat{x}$,
# \begin{equation}
# \hat{x} = x_n - \frac{f'(x_n)}{f''(x_n)}\left[1 \pm \sqrt{1 - \frac{2 f''(x_n) f(x_n)}{f'(x_n)^2}}\right].
# \end{equation}
# **There is one valid solution if we require $x_n \to \hat{x}$ as $f(x_n)\to 0$.** To see this, set $f(x_n)=0$ in the above equation and solve for $y=\hat{x}-x_n$. Only one of the two solutions is zero (assuming that $f'(\hat{x}) \neq 0$). This might seem counter intuitive, but it is not once you look at it in simpler terms. There are two (possibly complex) roots of the polynomial $ax^2 + bx + c = 0$ (assuming $a \neq 0$), but only one of them will vanish as $c\to 0$ (assuming $b \neq 0$).
#
# Hence,
# \begin{equation}
# \hat{x} = x_n - \frac{f'(x_n)}{f''(x_n)}\left[1 - \sqrt{1 - \frac{2 f''(x_n) f(x_n)}{f'(x_n)^2}}\right] + O((\hat{x} - x_n)^3).
# \end{equation}
# Using the above, we propose the fixed point iteration
# $$ x_{n+1} = g(x_n) = x_n - \frac{f'(x_n)}{f''(x_n)}\left[1 - \sqrt{1 - \frac{2 f''(x_n) f(x_n)}{f'(x_n)^2}}\right]$$
#
# ------------------------------------------------------------------------------------------
# **Comment:** The above is sufficient for full credit. However, one can futher simplify the method by expanding the square root term around $y = 0$, where $y = f(x_n)$. Assuming that $\hat{x}$ is a simple root, we have $f(x_n) = O(x_n - \hat{x})$, $f'(x_n) = O(1)$, and $f''(x_n) = O(1)$. To maintain the same accuracy, we need to expand the square root to explicitly include $O(y^2)$ terms. The $O(y^3)$ terms can be ignored and the same accuracy will be maintained. This leads to the iteration formula
# $$ x_{n+1} = x_n - \frac{f(x_n)}{f'(x_n)} - \frac{f''(x_n)f(x_n)^2}{f'(x_n)^3}.$$
# One can show that the above iteration method also has cubic convergence.
# # B
# Show that the order of convergence (under appropriate conditions) is cubic.
# ---------------------------------------
# ## Solution
# From the solution of part A we have
# \begin{equation}
# \hat{x} = x_n - \frac{f'(x_n)}{f''(x_n)}\left[1 - \sqrt{1 - \frac{2 f''(x_n) f(x_n)}{f'(x_n)^2}}\right] + O((\hat{x} - x_n)^3),
# \end{equation}
# and
# $$ x_{n+1} = g(x_n) = x_n - \frac{f'(x_n)}{f''(x_n)}\left[1 - \sqrt{1 - \frac{2 f''(x_n) f(x_n)}{f'(x_n)^2}}\right].$$
# Taking the difference of the above two equation yields
# $$ \vert x_{n+1} - \hat{x} \vert = O((\hat{x} - x_n)^3). $$
# # C
# Implement the root-finding method in Python to compute the root of $f(x) = x^3 - 2$. Add a stopping criterion that requires $\vert f(x_n) \vert \leq 10^{-8}$. Save the value of $x_n$ at each iteration and create a plot showing the convergence rate.
# +
## parameters
max_steps = 50 ## max number of iterations to use
tol = 1e-8 ## 10^(-8)
x0 = 2.
xhat = 2.**(1/3.)
def f(x):
return x**3 - 2.
def fp(x):
return 3*x**2
def fpp(x):
return 6*x
x = [x0]
for j in arange(max_steps):
fj = f(x[j])
fpj = fp(x[j])
fppj = fpp(x[j])
x.append(x[j] - fpj/fppj*(1 - sqrt(1 - 2*fppj*fj/fpj**2)))
if absolute(fj) < tol:
break ## this will stop the for loop
x = array(x) ## convert the list into an array
figure(1, [7, 4])
xplot = linspace(-1, 2, 200)
plot(xplot, 0*xplot, 'k') ## plot the line y=0
plot(xplot, f(xplot)) ## plot f(x)
plot(x, f(x), '*') ## plot the iterates of bisection
xlabel(r'$x$', fontsize=24) ## x axis label
ylabel(r'$f(x)$', fontsize=24); ## y axis label
## Convergence plot
figure(2, [7, 4])
err = absolute(x - xhat)
loglog(err[:-1], err[1:], '*') ## plot the iterates of bisection
err_plot = array([1e-3, 1e-2, 1e-1, 1.])
conv = err_plot**3 # quadratic. the theoretecal convergence curve
loglog(err_plot, conv, 'k')
xlim(1e-3, 1)
xlabel(r'$\vert x_n - \hat{x}\vert$', fontsize=24) ## x axis label
ylabel(r'$\vert x_{n+1} - \hat{x}\vert$', fontsize=24) ## y axis label
title('cubic convergence');
# -
# # D
# Using your code and the function $f$ defined in part C, numerically estimate the number of iterations needed to reduce the initial error, $\mathcal{E}_0 = \vert \hat{x} - x_0\vert$, by factor of $10^m$ for $m=1, \ldots 4$. Do this for each of the initial guesses $x_0 = 0.25, 1.25$.
# +
x_exact = 2.**(1./3.)
def number_of_iterations(x0, reduce_factor):
initial_error = absolute(x0 - x_exact)
x = [x0]
for j in arange(max_steps):
fj = f(x[j])
fpj = fp(x[j])
fppj = fpp(x[j])
x.append(x[j] - fpj/fppj*(1 - sqrt(1 - 2*fppj*fj/fpj**2)))
if absolute(x[-1] - x_exact) < initial_error/reduce_factor:
break ## this will stop the for loop
return j + 1
print('for x0 = 0.25')
print('number of iterations needed:')
print([number_of_iterations(0.25, 10.**m) for m in arange(1, 5)])
print('')
print('for x0 = 1.25')
print('number of iterations needed:')
print([number_of_iterations(1.25, 10.**m) for m in arange(1, 5)])
# -
# ## JUST FOR FUN!!!
# +
x_exact = 2.**(1./3.)
def number_of_iterations_V2(x0, reduce_factor):
initial_error = absolute(x0 - x_exact)
x = [x0]
for j in arange(max_steps):
fj = f(x[j])
fpj = fp(x[j])
fppj = fpp(x[j])
x.append(x[j] - fj/fpj - fppj*fj**2/fpj**3) ## See the comment in my answer to part A
if absolute(x[-1] - x_exact) < initial_error/reduce_factor:
break ## this will stop the for loop
return j + 1
print('for x0 = -1')
print('number of iterations needed:')
print([number_of_iterations_V2(-1., 10.**m) for m in arange(1, 5)])
print('')
print('for x0 = 0.1')
print('number of iterations needed:')
print([number_of_iterations_V2(0.1, 10.**m) for m in arange(1, 5)])
print('')
print('for x0 = 1.25')
print('number of iterations needed:')
print([number_of_iterations_V2(1.25, 10.**m) for m in arange(1, 5)])
print('')
print('for x0 = 2.25')
print('number of iterations needed:')
print([number_of_iterations_V2(2.25, 10.**m) for m in arange(1, 5)])
print('')
print('for x0 = 3.25')
print('number of iterations needed:')
print([number_of_iterations_V2(3.25, 10.**m) for m in arange(1, 5)])
# -
| Homework 4 Solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
import numpy as np
tf.set_random_seed(777) # for reproducibility
xy = np.loadtxt('data-5-diabetes.csv', delimiter=',',dtype=np.float32)
x_data = xy[:,0:-1]
y_data = xy[:,[-1]]
print(x_data.shape, y_data.shape)
X = tf.placeholder(tf.float32,shape=[None,8])
Y = tf.placeholder(tf.float32,shape=[None,1])
W = tf.Variable(tf.random_normal([8,1]),name='weight')
b = tf.Variable(tf.random_normal([1]), name = 'bias')
# +
hypothesis = tf.sigmoid(tf.matmul(X,W)+b)
cost = -tf.reduce_mean(Y*tf.log(hypothesis) + (1-Y)*tf.log(1-hypothesis))
# -
train = tf.train.GradientDescentOptimizer(learning_rate=0.001).minimize(cost)
predicted = tf.cast(hypothesis > 0.5, dtype=tf.float32)
accuracy = tf.reduce_mean(tf.cast(tf.equal(predicted, Y), dtype=tf.float32))
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for step in range(100001):
cost_val, _ = sess.run([cost,train],feed_dict={X:x_data,Y:y_data})
if step % 1000 == 0:
print(cost_val)
h, c, a = sess.run([hypothesis, predicted, accuracy],
feed_dict={X: x_data, Y: y_data})
print("\nHypothesis: ", h, "\nCorrect (Y): ", c, "\nAccuracy: ", a)
| ML_project/5-2.Classifying diabetes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/soumilbaldota/TensorFlow_Notebooks_101/blob/main/HiddenMarkowModel_Basic_TensorFlow.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="jh0b6U6cC43-"
# %tensorflow_version 2.x
# + id="peHrZiqiITTA"
import tensorflow_probability as tfp
import tensorflow as tf
import numpy as np
# + id="hCvmcKU4Jg2G"
tfd=tfp.distributions
initial_distribution=tfd.Categorical(probs=[0.8,0.2])
transition_distribution=tfd.Categorical(probs=[[0.7,0.3],[0.2,0.8]])
observation_distribution=tfd.Normal(loc=[0.,15.],scale=[5.,10.])
# + id="KWDh4uaMKPzw"
model=tfd.HiddenMarkovModel(initial_distribution=initial_distribution,
transition_distribution=transition_distribution,
observation_distribution=observation_distribution,
num_steps=7)
# + id="tQUHuiKrMpJo" outputId="0268ff73-efd3-438d-cdcc-d020df20c5e7" colab={"base_uri": "https://localhost:8080/", "height": 34}
mean=model.mean()
with tf.compat.v1.Session() as sess:
print(mean.numpy())
| HiddenMarkowModel_Basic_TensorFlow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Building and deploying machine learning solutions with Vertex AI: Challenge Lab
# This Challenge Lab is recommended for students who have enrolled in the [**Building and deploying machine learning solutions with Vertex AI**](). You will be given a scenario and a set of tasks. Instead of following step-by-step instructions, you will use the skills learned from the labs in the quest to figure out how to complete the tasks on your own! An automated scoring system (shown on the Qwiklabs lab instructions page) will provide feedback on whether you have completed your tasks correctly.
#
# When you take a Challenge Lab, you will not be taught Google Cloud concepts. To build the solution to the challenge presented, use skills learned from the labs in the Quest this challenge lab is part of. You are expected to extend your learned skills and complete all the **`TODO:`** comments in this notebook.
#
# Are you ready for the challenge?
# ## Scenario
# You were recently hired as a Machine Learning Engineer at a startup movie review website. Your manager has tasked you with building a machine learning model to classify the sentiment of user movie reviews as positive or negative. These predictions will be used as an input in downstream movie rating systems and to surface top supportive and critical reviews on the movie website application. The challenge: your business requirements are that you have just 6 weeks to productionize a model that achieves great than 75% accuracy to improve upon an existing bootstrapped solution. Furthermore, after doing some exploratory analysis in your startup's data warehouse, you found that you only have a small dataset of 50k text reviews to build a higher performing solution.
#
# To build and deploy a high performance machine learning model with limited data quickly, you will walk through training and deploying a custom TensorFlow BERT sentiment classifier for online predictions on Google Cloud's [Vertex AI](https://cloud.google.com/vertex-ai) platform. Vertex AI is Google Cloud's next generation machine learning development platform where you can leverage the latest ML pre-built components and AutoML to significantly enhance your development productivity, scale your workflow and decision making with your data, and accelerate time to value.
#
# 
#
# First, you will progress through a typical experimentation workflow where you will build your model from pre-trained BERT components from TF-Hub and `tf.keras` classification layers to train and evaluate your model in a Vertex Notebook. You will then package your model code into a Docker container to train on Google Cloud's Vertex AI. Lastly, you will define and run a Kubeflow Pipeline on Vertex Pipelines that trains and deploys your model to a Vertex Endpoint that you will query for online predictions.
# ## Learning objectives
# * Train a TensorFlow model locally in a hosted [**Vertex Notebook**](https://cloud.google.com/vertex-ai/docs/general/notebooks?hl=sv).
# * Containerize your training code with [**Cloud Build**](https://cloud.google.com/build) and push it to [**Google Cloud Artifact Registry**](https://cloud.google.com/artifact-registry).
# * Define a pipeline using the [**Kubeflow Pipelines (KFP) V2 SDK**](https://www.kubeflow.org/docs/components/pipelines/sdk/v2/v2-compatibility) to train and deploy your model on [**Vertex Pipelines**](https://cloud.google.com/vertex-ai/docs/pipelines).
# * Query your model on a [**Vertex Endpoint**](https://cloud.google.com/vertex-ai/docs/predictions/getting-predictions) using online predictions.
# ## Setup
# ### Define constants
# Add installed library dependencies to Python PATH variable.
# PATH=%env PATH
# %env PATH={PATH}:/home/jupyter/.local/bin
# Retrieve and set PROJECT_ID and REGION environment variables.
# TODO: fill in PROJECT_ID.
PROJECT_ID = ""
REGION = "us-central1"
# TODO: Create a globally unique Google Cloud Storage bucket for artifact storage.
GCS_BUCKET = ""
# !gsutil mb -l $REGION $GCS_BUCKET
# ### Import libraries
# +
import os
import shutil
import logging
# TensorFlow model building libraries.
import tensorflow as tf
import tensorflow_text as text
import tensorflow_hub as hub
# Re-create the AdamW optimizer used in the original BERT paper.
from official.nlp import optimization
# Libraries for data and plot model training metrics.
import pandas as pd
import matplotlib.pyplot as plt
# Import the Vertex AI Python SDK.
from google.cloud import aiplatform as vertexai
# -
# ### Initialize Vertex AI Python SDK
# Initialize the Vertex AI Python SDK with your GCP Project, Region, and Google Cloud Storage Bucket.
vertexai.init(project=PROJECT_ID, location=REGION, staging_bucket=GCS_BUCKET)
# ## Build and train your model locally in a Vertex Notebook
# Note: this lab adapts and extends the official [TensorFlow BERT text classification tutorial](https://www.tensorflow.org/text/tutorials/classify_text_with_bert) to utilize Vertex AI services. See the tutorial for additional coverage on fine-tuning BERT models using TensorFlow.
# ### Lab dataset
# In this lab, you will use the [Large Movie Review Dataset](https://ai.stanford.edu/~amaas/data/sentiment) that contains the text of 50,000 movie reviews from the Internet Movie Database. These are split into 25,000 reviews for training and 25,000 reviews for testing. The training and testing sets are balanced, meaning they contain an equal number of positive and negative reviews. Data ingestion and processing code has been provided for you below:
# ### Import dataset
DATA_URL = "https://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz"
LOCAL_DATA_DIR = "."
def download_data(data_url, local_data_dir):
"""Download dataset.
Args:
data_url(str): Source data URL path.
local_data_dir(str): Local data download directory path.
Returns:
dataset_dir(str): Local unpacked data directory path.
"""
if not os.path.exists(local_data_dir):
os.makedirs(local_data_dir)
dataset = tf.keras.utils.get_file(
fname="aclImdb_v1.tar.gz",
origin=data_url,
untar=True,
cache_dir=local_data_dir,
cache_subdir="")
dataset_dir = os.path.join(os.path.dirname(dataset), "aclImdb")
train_dir = os.path.join(dataset_dir, "train")
# Remove unused folders to make it easier to load the data.
remove_dir = os.path.join(train_dir, "unsup")
shutil.rmtree(remove_dir)
return dataset_dir
DATASET_DIR = download_data(data_url=DATA_URL, local_data_dir=LOCAL_DATA_DIR)
# Create a dictionary to iteratively add data pipeline and model training hyperparameters.
HPARAMS = {
# Set a random sampling seed to prevent data leakage in data splits from files.
"seed": 42,
# Number of training and inference examples.
"batch-size": 32
}
def load_datasets(dataset_dir, hparams):
"""Load pre-split tf.datasets.
Args:
hparams(dict): A dictionary containing model training arguments.
Returns:
raw_train_ds(tf.dataset): Train split dataset (20k examples).
raw_val_ds(tf.dataset): Validation split dataset (5k examples).
raw_test_ds(tf.dataset): Test split dataset (25k examples).
"""
raw_train_ds = tf.keras.preprocessing.text_dataset_from_directory(
os.path.join(dataset_dir, 'train'),
batch_size=hparams['batch-size'],
validation_split=0.2,
subset='training',
seed=hparams['seed'])
raw_val_ds = tf.keras.preprocessing.text_dataset_from_directory(
os.path.join(dataset_dir, 'train'),
batch_size=hparams['batch-size'],
validation_split=0.2,
subset='validation',
seed=hparams['seed'])
raw_test_ds = tf.keras.preprocessing.text_dataset_from_directory(
os.path.join(dataset_dir, 'test'),
batch_size=hparams['batch-size'])
return raw_train_ds, raw_val_ds, raw_test_ds
raw_train_ds, raw_val_ds, raw_test_ds = load_datasets(DATASET_DIR, HPARAMS)
# +
AUTOTUNE = tf.data.AUTOTUNE
CLASS_NAMES = raw_train_ds.class_names
train_ds = raw_train_ds.prefetch(buffer_size=AUTOTUNE)
val_ds = raw_val_ds.prefetch(buffer_size=AUTOTUNE)
test_ds = raw_test_ds.prefetch(buffer_size=AUTOTUNE)
# -
# Let's print a few example reviews:
for text_batch, label_batch in train_ds.take(1):
for i in range(3):
print(f'Review {i}: {text_batch.numpy()[i]}')
label = label_batch.numpy()[i]
print(f'Label : {label} ({CLASS_NAMES[label]})')
# ### Choose a pre-trained BERT model to fine-tune for higher accuracy
# [**Bidirectional Encoder Representations from Transformers (BERT)**](https://arxiv.org/abs/1810.04805v2) is a transformer-based text representation model pre-trained on massive datasets (3+ billion words) that can be fine-tuned for state-of-the art results on many natural language processing (NLP) tasks. Since release in 2018 by Google researchers, its has transformed the field of NLP research and come to form a core part of significant improvements to [Google Search](https://www.blog.google/products/search/search-language-understanding-bert).
#
# To meet your business requirements of achieving higher accuracy on a small dataset (20k training examples), you will use a technique called transfer learning to combine a pre-trained BERT encoder and classification layers to fine tune a new higher performing model for binary sentiment classification.
# For this lab, you will use a smaller BERT model that trades some accuracy for faster training times.
#
# The Small BERT models are instances of the original BERT architecture with a smaller number L of layers (i.e., residual blocks) combined with a smaller hidden size H and a matching smaller number A of attention heads, as published by
#
# <NAME>, <NAME>, <NAME>, <NAME>: ["Well-Read Students Learn Better: On the Importance of Pre-training Compact Models"](https://arxiv.org/abs/1908.08962), 2019.
#
# They have the same general architecture but fewer and/or smaller Transformer blocks, which lets you explore tradeoffs between speed, size and quality.
#
# The following preprocessing and encoder models in the TensorFlow 2 SavedModel format use the implementation of BERT from the [TensorFlow Models Github repository](https://github.com/tensorflow/models/tree/master/official/nlp/bert) with the trained weights released by the authors of Small BERT.
HPARAMS.update({
# TF Hub BERT modules.
"tfhub-bert-preprocessor": "https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3",
"tfhub-bert-encoder": "https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-2_H-128_A-2/2",
})
# Text inputs need to be transformed to numeric token ids and arranged in several Tensors before being input to BERT. TensorFlow Hub provides a matching preprocessing model for each of the BERT models discussed above, which implements this transformation using TF ops from the TF.text library. Since this text preprocessor is a TensorFlow model, It can be included in your model directly.
# For fine-tuning, you will use the same optimizer that BERT was originally trained with: the "Adaptive Moments" (Adam). This optimizer minimizes the prediction loss and does regularization by weight decay (not using moments), which is also known as [AdamW](https://arxiv.org/abs/1711.05101).
# For the learning rate `initial-learning-rate`, you will use the same schedule as BERT pre-training: linear decay of a notional initial learning rate, prefixed with a linear warm-up phase over the first 10% of training steps `n_warmup_steps`. In line with the BERT paper, the initial learning rate is smaller for fine-tuning.
HPARAMS.update({
# Model training hyperparameters for fine tuning and regularization.
"epochs": 3,
"initial-learning-rate": 3e-5,
"dropout": 0.1
})
# +
epochs = HPARAMS['epochs']
steps_per_epoch = tf.data.experimental.cardinality(train_ds).numpy()
n_train_steps = steps_per_epoch * epochs
n_warmup_steps = int(0.1 * n_train_steps)
OPTIMIZER = optimization.create_optimizer(init_lr=HPARAMS['initial-learning-rate'],
num_train_steps=n_train_steps,
num_warmup_steps=n_warmup_steps,
optimizer_type='adamw')
# -
# ### Build and compile a TensorFlow BERT sentiment classifier
# Next, you will define and compile your model by assembling pre-built TF-Hub components and tf.keras layers.
def build_text_classifier(hparams, optimizer):
"""Define and compile a TensorFlow BERT sentiment classifier.
Args:
hparams(dict): A dictionary containing model training arguments.
Returns:
model(tf.keras.Model): A compiled TensorFlow model.
"""
text_input = tf.keras.layers.Input(shape=(), dtype=tf.string, name='text')
# TODO: Add a hub.KerasLayer for BERT text preprocessing using the hparams dict.
# Name the layer 'preprocessing' and store in the variable preprocessor.
encoder_inputs = preprocessor(text_input)
# TODO: Add a trainable hub.KerasLayer for BERT text encoding using the hparams dict.
# Name the layer 'BERT_encoder' and store in the variable encoder.
outputs = encoder(encoder_inputs)
# For the fine-tuning you are going to use the `pooled_output` array which represents
# each input sequence as a whole. The shape is [batch_size, H].
# You can think of this as an embedding for the entire movie review.
classifier = outputs['pooled_output']
# Add dropout to prevent overfitting during model fine-tuning.
classifier = tf.keras.layers.Dropout(hparams['dropout'], name='dropout')(classifier)
classifier = tf.keras.layers.Dense(1, activation=None, name='classifier')(classifier)
model = tf.keras.Model(text_input, classifier, name='bert-sentiment-classifier')
loss = tf.keras.losses.BinaryCrossentropy(from_logits=True)
metrics = tf.metrics.BinaryAccuracy()
model.compile(optimizer=optimizer,
loss=loss,
metrics=metrics)
return model
model = build_text_classifier(HPARAMS, OPTIMIZER)
# Visualize your fine-tuned BERT sentiment classifier.
tf.keras.utils.plot_model(model)
TEST_REVIEW = ['this is such an amazing movie!']
BERT_RAW_RESULT = model(tf.constant(TEST_REVIEW))
print(BERT_RAW_RESULT)
# ### Train and evaluate your BERT sentiment classifier
# +
HPARAMS.update({
# TODO: Save your BERT sentiment classifier locally.
# Hint: Save it to './bert-sentiment-classifier-local'. Note the key name in model.save().
})
# -
# **Note:** training your model locally will take about 8-10 minutes.
def train_evaluate(hparams):
"""Train and evaluate TensorFlow BERT sentiment classifier.
Args:
hparams(dict): A dictionary containing model training arguments.
Returns:
history(tf.keras.callbacks.History): Keras callback that records training event history.
"""
# dataset_dir = download_data(data_url, local_data_dir)
raw_train_ds, raw_val_ds, raw_test_ds = load_datasets(DATASET_DIR, hparams)
train_ds = raw_train_ds.cache().prefetch(buffer_size=AUTOTUNE)
val_ds = raw_val_ds.cache().prefetch(buffer_size=AUTOTUNE)
test_ds = raw_test_ds.cache().prefetch(buffer_size=AUTOTUNE)
epochs = hparams['epochs']
steps_per_epoch = tf.data.experimental.cardinality(train_ds).numpy()
n_train_steps = steps_per_epoch * epochs
n_warmup_steps = int(0.1 * n_train_steps)
optimizer = optimization.create_optimizer(init_lr=hparams['initial-learning-rate'],
num_train_steps=n_train_steps,
num_warmup_steps=n_warmup_steps,
optimizer_type='adamw')
mirrored_strategy = tf.distribute.MirroredStrategy()
with mirrored_strategy.scope():
model = build_text_classifier(hparams=hparams, optimizer=optimizer)
logging.info(model.summary())
history = model.fit(x=train_ds,
validation_data=val_ds,
epochs=epochs)
logging.info("Test accuracy: %s", model.evaluate(test_ds))
# Export Keras model in TensorFlow SavedModel format.
model.save(hparams['model-dir'])
return history
# Based on the `History` object returned by `model.fit()`. You can plot the training and validation loss for comparison, as well as the training and validation accuracy:
history = train_evaluate(HPARAMS)
# +
history_dict = history.history
print(history_dict.keys())
acc = history_dict['binary_accuracy']
val_acc = history_dict['val_binary_accuracy']
loss = history_dict['loss']
val_loss = history_dict['val_loss']
epochs = range(1, len(acc) + 1)
fig = plt.figure(figsize=(10, 6))
fig.tight_layout()
plt.subplot(2, 1, 1)
# "bo" is for "blue dot"
plt.plot(epochs, loss, 'r', label='Training loss')
# b is for "solid blue line"
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
# plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.subplot(2, 1, 2)
plt.plot(epochs, acc, 'r', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend(loc='lower right');
# -
# In this plot, the red lines represent the training loss and accuracy, and the blue lines are the validation loss and accuracy. Based on the plots above, you should see model accuracy of around 78-80% which exceeds your business requirements target of greater than 75% accuracy.
# ## Containerize your model code
# Now that you trained and evaluated your model locally in a Vertex Notebook as part of an experimentation workflow, your next step is to train and deploy your model on Google Cloud's Vertex AI platform.
# To train your BERT classifier on Google Cloud, you will you will package your Python training scripts and write a Dockerfile that contains instructions on your ML model code, dependencies, and execution instructions. You will build your custom container with Cloud Build, whose instructions are specified in `cloudbuild.yaml` and publish your container to your Artifact Registry. This workflow gives you the opportunity to use the same container to run as part of a portable and scalable [Vertex Pipelines](https://cloud.google.com/vertex-ai/docs/pipelines/introduction) workflow.
#
#
# You will walk through creating the following project structure for your ML mode code:
# ```
# |--/bert-sentiment-classifier
# |--/trainer
# |--__init__.py
# |--model.py
# |--task.py
# |--Dockerfile
# |--cloudbuild.yaml
# |--requirements.txt
# ```
# ### 1. Write a `model.py` training script
#
# First, you will tidy up your local TensorFlow model training code from above into a training script.
MODEL_DIR = "bert-sentiment-classifier"
# +
# %%writefile {MODEL_DIR}/trainer/model.py
import os
import shutil
import logging
import tensorflow as tf
import tensorflow_text as text
import tensorflow_hub as hub
from official.nlp import optimization
DATA_URL = 'https://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz'
LOCAL_DATA_DIR = './tmp/data'
AUTOTUNE = tf.data.AUTOTUNE
def download_data(data_url, local_data_dir):
"""Download dataset.
Args:
data_url(str): Source data URL path.
local_data_dir(str): Local data download directory path.
Returns:
dataset_dir(str): Local unpacked data directory path.
"""
if not os.path.exists(local_data_dir):
os.makedirs(local_data_dir)
dataset = tf.keras.utils.get_file(
fname='aclImdb_v1.tar.gz',
origin=data_url,
untar=True,
cache_dir=local_data_dir,
cache_subdir="")
dataset_dir = os.path.join(os.path.dirname(dataset), 'aclImdb')
train_dir = os.path.join(dataset_dir, 'train')
# Remove unused folders to make it easier to load the data.
remove_dir = os.path.join(train_dir, 'unsup')
shutil.rmtree(remove_dir)
return dataset_dir
def load_datasets(dataset_dir, hparams):
"""Load pre-split tf.datasets.
Args:
hparams(dict): A dictionary containing model training arguments.
Returns:
raw_train_ds(tf.dataset): Train split dataset (20k examples).
raw_val_ds(tf.dataset): Validation split dataset (5k examples).
raw_test_ds(tf.dataset): Test split dataset (25k examples).
"""
raw_train_ds = tf.keras.preprocessing.text_dataset_from_directory(
os.path.join(dataset_dir, 'train'),
batch_size=hparams['batch-size'],
validation_split=0.2,
subset='training',
seed=hparams['seed'])
raw_val_ds = tf.keras.preprocessing.text_dataset_from_directory(
os.path.join(dataset_dir, 'train'),
batch_size=hparams['batch-size'],
validation_split=0.2,
subset='validation',
seed=hparams['seed'])
raw_test_ds = tf.keras.preprocessing.text_dataset_from_directory(
os.path.join(dataset_dir, 'test'),
batch_size=hparams['batch-size'])
return raw_train_ds, raw_val_ds, raw_test_ds
def build_text_classifier(hparams, optimizer):
"""Define and compile a TensorFlow BERT sentiment classifier.
Args:
hparams(dict): A dictionary containing model training arguments.
Returns:
model(tf.keras.Model): A compiled TensorFlow model.
"""
text_input = tf.keras.layers.Input(shape=(), dtype=tf.string, name='text')
# TODO: Add a hub.KerasLayer for BERT text preprocessing using the hparams dict.
# Name the layer 'preprocessing' and store in the variable preprocessor.
preprocessor = hub.KerasLayer(hparams['tfhub-bert-preprocessor'], name='preprocessing')
encoder_inputs = preprocessor(text_input)
# TODO: Add a trainable hub.KerasLayer for BERT text encoding using the hparams dict.
# Name the layer 'BERT_encoder' and store in the variable encoder.
encoder = hub.KerasLayer(hparams['tfhub-bert-encoder'], trainable=True, name='BERT_encoder')
outputs = encoder(encoder_inputs)
# For the fine-tuning you are going to use the `pooled_output` array which represents
# each input sequence as a whole. The shape is [batch_size, H].
# You can think of this as an embedding for the entire movie review.
classifier = outputs['pooled_output']
# Add dropout to prevent overfitting during model fine-tuning.
classifier = tf.keras.layers.Dropout(hparams['dropout'], name='dropout')(classifier)
classifier = tf.keras.layers.Dense(1, activation=None, name='classifier')(classifier)
model = tf.keras.Model(text_input, classifier, name='bert-sentiment-classifier')
loss = tf.keras.losses.BinaryCrossentropy(from_logits=True)
metrics = tf.metrics.BinaryAccuracy()
model.compile(optimizer=optimizer,
loss=loss,
metrics=metrics)
return model
def train_evaluate(hparams):
"""Train and evaluate TensorFlow BERT sentiment classifier.
Args:
hparams(dict): A dictionary containing model training arguments.
Returns:
history(tf.keras.callbacks.History): Keras callback that records training event history.
"""
dataset_dir = download_data(data_url=DATA_URL,
local_data_dir=LOCAL_DATA_DIR)
raw_train_ds, raw_val_ds, raw_test_ds = load_datasets(dataset_dir=dataset_dir,
hparams=hparams)
train_ds = raw_train_ds.cache().prefetch(buffer_size=AUTOTUNE)
val_ds = raw_val_ds.cache().prefetch(buffer_size=AUTOTUNE)
test_ds = raw_test_ds.cache().prefetch(buffer_size=AUTOTUNE)
epochs = hparams['epochs']
steps_per_epoch = tf.data.experimental.cardinality(train_ds).numpy()
n_train_steps = steps_per_epoch * epochs
n_warmup_steps = int(0.1 * n_train_steps)
optimizer = optimization.create_optimizer(init_lr=hparams['initial-learning-rate'],
num_train_steps=n_train_steps,
num_warmup_steps=n_warmup_steps,
optimizer_type='adamw')
mirrored_strategy = tf.distribute.MirroredStrategy()
with mirrored_strategy.scope():
model = build_text_classifier(hparams=hparams, optimizer=optimizer)
logging.info(model.summary())
history = model.fit(x=train_ds,
validation_data=val_ds,
epochs=epochs)
logging.info("Test accuracy: %s", model.evaluate(test_ds))
# Export Keras model in TensorFlow SavedModel format.
model.save(hparams['model-dir'])
return history
# -
# ### 2. Write a `task.py` file as an entrypoint to your custom model container
# +
# %%writefile {MODEL_DIR}/trainer/task.py
import os
import argparse
from trainer import model
if __name__ == '__main__':
parser = argparse.ArgumentParser()
# Vertex custom container training args. These are set by Vertex AI during training but can also be overwritten.
parser.add_argument('--model-dir', dest='model-dir',
default=os.environ['AIP_MODEL_DIR'], type=str, help='GCS URI for saving model artifacts.')
# Model training args.
parser.add_argument('--tfhub-bert-preprocessor', dest='tfhub-bert-preprocessor',
default='https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3', type=str, help='TF-Hub URL.')
parser.add_argument('--tfhub-bert-encoder', dest='tfhub-bert-encoder',
default='https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-2_H-128_A-2/2', type=str, help='TF-Hub URL.')
parser.add_argument('--initial-learning-rate', dest='initial-learning-rate', default=3e-5, type=float, help='Learning rate for optimizer.')
parser.add_argument('--epochs', dest='epochs', default=3, type=int, help='Training iterations.')
parser.add_argument('--batch-size', dest='batch-size', default=32, type=int, help='Number of examples during each training iteration.')
parser.add_argument('--dropout', dest='dropout', default=0.1, type=float, help='Float percentage of DNN nodes [0,1] to drop for regularization.')
parser.add_argument('--seed', dest='seed', default=42, type=int, help='Random number generator seed to prevent overlap between train and val sets.')
args = parser.parse_args()
hparams = args.__dict__
model.train_evaluate(hparams)
# -
# ### 3. Write a `Dockerfile` for your custom model container
# Third, you will write a `Dockerfile` that contains instructions to package your model code in `bert-sentiment-classifier` as well as specifies your model code's dependencies needed for execution together in a Docker container.
# +
# %%writefile {MODEL_DIR}/Dockerfile
# Specifies base image and tag.
# https://cloud.google.com/vertex-ai/docs/training/pre-built-containers
FROM us-docker.pkg.dev/vertex-ai/training/tf-cpu.2-6:latest
# Sets the container working directory.
WORKDIR /root
# Copies the requirements.txt into the container to reduce network calls.
COPY requirements.txt .
# Installs additional packages.
RUN pip3 install -U -r requirements.txt
# b/203105209 Removes unneeded file from TF2.5 CPU image for python_module CustomJob training.
# Will be removed on subsequent public Vertex images.
RUN rm -rf /var/sitecustomize/sitecustomize.py
# Copies the trainer code to the docker image.
COPY . /trainer
# Sets the container working directory.
WORKDIR /trainer
# Sets up the entry point to invoke the trainer.
ENTRYPOINT ["python", "-m", "trainer.task"]
# -
# ### 4. Write a `requirements.txt` file to specify additional ML code dependencies
# These are additional dependencies for your model code not included in the pre-built Vertex TensorFlow images such as TF-Hub, TensorFlow AdamW optimizer, and TensorFlow Text needed for importing and working with pre-trained TensorFlow BERT models.
# %%writefile {MODEL_DIR}/requirements.txt
tf-models-official==2.6.0
tensorflow-text==2.6.0
tensorflow-hub==0.12.0
# ## Use Cloud Build to build and submit your model container to Google Cloud Artifact Registry
# Next, you will use [Cloud Build](https://cloud.google.com/build) to build and upload your custom TensorFlow model container to [Google Cloud Artifact Registry](https://cloud.google.com/artifact-registry).
#
# Cloud Build brings reusability and automation to your ML experimentation by enabling you to reliably build, test, and deploy your ML model code as part of a CI/CD workflow. Artifact Registry provides a centralized repository for you to store, manage, and secure your ML container images. This will allow you to securely share your ML work with others and reproduce experiment results.
#
# **Note**: the initial build and submit step will take about 16 minutes but Cloud Build is able to take advantage of caching for faster subsequent builds.
# ### 1. Create Artifact Registry for custom container images
ARTIFACT_REGISTRY="bert-sentiment-classifier"
# TODO: create a Docker Artifact Registry using the gcloud CLI. Note the required respository-format and location flags.
# Documentation link: https://cloud.google.com/sdk/gcloud/reference/artifacts/repositories/create
# ### 2. Create `cloudbuild.yaml` instructions
IMAGE_NAME="bert-sentiment-classifier"
IMAGE_TAG="latest"
IMAGE_URI=f"{REGION}-docker.pkg.dev/{PROJECT_ID}/{ARTIFACT_REGISTRY}/{IMAGE_NAME}:{IMAGE_TAG}"
# +
cloudbuild_yaml = f"""steps:
- name: 'gcr.io/cloud-builders/docker'
args: [ 'build', '-t', '{IMAGE_URI}', '.' ]
images:
- '{IMAGE_URI}'"""
with open(f"{MODEL_DIR}/cloudbuild.yaml", "w") as fp:
fp.write(cloudbuild_yaml)
# -
# ### 3. Build and submit your container image to Artifact Registry using Cloud Build
# **Note:** your custom model container will take about 16 minutes initially to build and submit to your Artifact Registry. Artifact Registry is able to take advantage of caching so subsequent builds take about 4 minutes.
# +
# TODO: use Cloud Build to build and submit your custom model container to your Artifact Registry.
# Documentation link: https://cloud.google.com/sdk/gcloud/reference/builds/submit
# Hint: make sure the config flag is pointed at {MODEL_DIR}/cloudbuild.yaml defined above and you include your model directory.
# -
# ## Define a pipeline using the KFP V2 SDK
# To address your business requirements and get your higher performing model into production to deliver value faster, you will define a pipeline using the [**Kubeflow Pipelines (KFP) V2 SDK**](https://www.kubeflow.org/docs/components/pipelines/sdk/v2/v2-compatibility) to orchestrate the training and deployment of your model on [**Vertex Pipelines**](https://cloud.google.com/vertex-ai/docs/pipelines) below.
import datetime
# google_cloud_pipeline_components includes pre-built KFP components for interfacing with Vertex AI services.
from google_cloud_pipeline_components import aiplatform as gcc_aip
from kfp.v2 import dsl
# +
TIMESTAMP=datetime.datetime.now().strftime('%Y%m%d%H%M%S')
DISPLAY_NAME = "bert-sentiment-{}".format(TIMESTAMP)
GCS_BASE_OUTPUT_DIR= f"{GCS_BUCKET}/{MODEL_DIR}-{TIMESTAMP}"
USER = "" # TODO: change this to your name.
PIPELINE_ROOT = "{}/pipeline_root/{}".format(GCS_BUCKET, USER)
print(f"Model display name: {DISPLAY_NAME}")
print(f"GCS dir for model training artifacts: {GCS_BASE_OUTPUT_DIR}")
print(f"GCS dir for pipeline artifacts: {PIPELINE_ROOT}")
# -
# Pre-built Vertex model serving container for deployment.
# https://cloud.google.com/vertex-ai/docs/predictions/pre-built-containers
SERVING_IMAGE_URI = "us-docker.pkg.dev/vertex-ai/prediction/tf2-cpu.2-6:latest"
# The pipeline consists of three components:
#
# * `CustomContainerTrainingJobRunOp` [(documentation)](https://google-cloud-pipeline-components.readthedocs.io/en/google-cloud-pipeline-components-0.2.0/google_cloud_pipeline_components.aiplatform.html#google_cloud_pipeline_components.aiplatform.CustomContainerTrainingJobRunOp): trains your custom model container using Vertex Training. This is the same as configuring a Vertex Custom Container Training Job using the Vertex Python SDK you covered in the Vertex AI: Qwik Start lab.
#
# * `EndpointCreateOp` [(documentation)](https://google-cloud-pipeline-components.readthedocs.io/en/google-cloud-pipeline-components-0.2.0/google_cloud_pipeline_components.aiplatform.html#google_cloud_pipeline_components.aiplatform.EndpointCreateOp): Creates a Google Cloud Vertex Endpoint resource that maps physical machine resources with your model to enable it to serve online predictions. Online predictions have low latency requirements; providing resources to the model in advance reduces latency.
#
# * `ModelDeployOp`[(documentation)](https://google-cloud-pipeline-components.readthedocs.io/en/google-cloud-pipeline-components-0.2.0/google_cloud_pipeline_components.aiplatform.html#google_cloud_pipeline_components.aiplatform.ModelDeployOp): deploys your model to a Vertex Prediction Endpoint for online predictions.
@dsl.pipeline(name="bert-sentiment-classification", pipeline_root=PIPELINE_ROOT)
def pipeline(
project: str = PROJECT_ID,
location: str = REGION,
staging_bucket: str = GCS_BUCKET,
display_name: str = DISPLAY_NAME,
container_uri: str = IMAGE_URI,
model_serving_container_image_uri: str = SERVING_IMAGE_URI,
base_output_dir: str = GCS_BASE_OUTPUT_DIR,
):
#TODO: add and configure the pre-built KFP CustomContainerTrainingJobRunOp component using
# the remaining arguments in the pipeline constructor.
# Hint: Refer to the component documentation link above if needed as well.
model_train_evaluate_op = gcc_aip.CustomContainerTrainingJobRunOp(
# Vertex AI Python SDK authentication parameters.
project=project,
location=location,
staging_bucket=staging_bucket,
# WorkerPool arguments.
replica_count=1,
machine_type="c2-standard-4",
# TODO: fill in the remaining arguments from the pipeline constructor.
)
# Create a Vertex Endpoint resource in parallel with model training.
endpoint_create_op = gcc_aip.EndpointCreateOp(
# Vertex AI Python SDK authentication parameters.
project=project,
location=location,
display_name=display_name
)
# Deploy your model to the created Endpoint resource for online predictions.
model_deploy_op = gcc_aip.ModelDeployOp(
# Link to model training component through output model artifact.
model=model_train_evaluate_op.outputs["model"],
# Link to the created Endpoint.
endpoint=endpoint_create_op.outputs["endpoint"],
# Define prediction request routing. {"0": 100} indicates 100% of traffic
# to the ID of the current model being deployed.
traffic_split={"0": 100},
# WorkerPool arguments.
dedicated_resources_machine_type="n1-standard-4",
dedicated_resources_min_replica_count=1,
dedicated_resources_max_replica_count=2
)
# ## Compile the pipeline
from kfp.v2 import compiler
compiler.Compiler().compile(
pipeline_func=pipeline, package_path="bert-sentiment-classification.json"
)
# ## Run the pipeline on Vertex Pipelines
# The `PipelineJob` is configured below and triggered through the `run()` method.
#
# **Note:** This pipeline run will take around 30-40 minutes to train and deploy your model. Follow along with the execution using the URL from the job output below.
vertex_pipelines_job = vertexai.pipeline_jobs.PipelineJob(
display_name="bert-sentiment-classification",
template_path="bert-sentiment-classification.json",
parameter_values={
"project": PROJECT_ID,
"location": REGION,
"staging_bucket": GCS_BUCKET,
"display_name": DISPLAY_NAME,
"container_uri": IMAGE_URI,
"model_serving_container_image_uri": SERVING_IMAGE_URI,
"base_output_dir": GCS_BASE_OUTPUT_DIR},
enable_caching=True,
)
vertex_pipelines_job.run()
# ## Query deployed model on Vertex Endpoint for online predictions
# Finally, you will retrieve the `Endpoint` deployed by the pipeline and use it to query your model for online predictions.
#
# Configure the `Endpoint()` function below with the following parameters:
#
# * `endpoint_name`: A fully-qualified endpoint resource name or endpoint ID. Example: "projects/123/locations/us-central1/endpoints/456" or "456" when project and location are initialized or passed.
# * `project_id`: GCP project.
# * `location`: GCP region.
#
# Call `predict()` to return a prediction for a test review.
# Retrieve your deployed Endpoint name from your pipeline.
ENDPOINT_NAME = vertexai.Endpoint.list()[0].name
# +
#TODO: Generate online predictions using your Vertex Endpoint.
endpoint = vertexai.Endpoint(
)
# -
#TODO: write a movie review to test your model e.g. "The Dark Knight is the best Batman movie!"
test_review = ""
# TODO: use your Endpoint to return prediction for your test_review.
prediction =
print(prediction)
# Use a sigmoid function to compress your model output between 0 and 1. For binary classification, a threshold of 0.5 is typically applied
# so if the output is >= 0.5 then the predicted sentiment is "Positive" and < 0.5 is a "Negative" prediction.
print(tf.sigmoid(prediction.predictions[0]))
# ## Next steps
# Congratulations! You walked through a full experimentation, containerization, and MLOps workflow on Vertex AI. First, you built, trained, and evaluated a BERT sentiment classifier model in a Vertex Notebook. You then packaged your model code into a Docker container to train on Google Cloud's Vertex AI. Lastly, you defined and ran a Kubeflow Pipeline on Vertex Pipelines that trained and deployed your model container to a Vertex Endpoint that you queried for online predictions.
# ## License
# +
# Copyright 2021 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
| quests/vertex-ai/vertex-challenge-lab/vertex-challenge-lab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Practice functions
#
# Review [`Intro_to_functions`](Intro_to_functions.ipynb) before coming in here.
#
# Our goal for this notebook is to get some practice writing functions.
#
# In doing so, we will implement a function to compute reflection coefficients from sequences of Vp and density values.
# +
import numpy as np
% matplotlib inline
import matplotlib.pyplot as plt
# -
# Make some dummy data:
vp = [2300, 2400, 2500, 2300, 2600]
rho = [2.45, 2.35, 2.45, 2.55, 2.80]
# Sometimes Vp data is in km/s and density is in g/cm<sup>3</sup>. Let's make a simple function to convert them to SI units.
def convert_si(n):
"""
Convert vp or rhob from cgs to SI system.
"""
if n < 10:
n = n * 1000
return n
convert_si(2400), convert_si(2.4)
convert_si(vp)
# + [markdown] tags=["exercise"]
# <div class="alert alert-success">
# <b>Exercise</b>:
# <ul>
# <li>- Make a looping version of `convert_si()` called `convert_all()` that will treat the whole list at once. Use the `convert_si()` function inside it. If you get stuck, write the loop on its own first, then put it in a function.</li>
# <li>- Can you write a function containing a `for` loop to implement this equation?</li>
# <li>$$ Z = \rho V_\mathrm{P} $$</li>
# <li>You will find the function `zip()` useful. Try the code below to see what it does.</li>
# </ul>
# </div>
# -
for pair in zip([1,2,3], [10,11,12]):
print(pair)
def impedance(vp, rho):
# Your code here!
return z
impedance(vp, rho)
# This should give you:
#
# [5635.0, 5640.0, 6125.0, 5865.0, 7279.999999999999]
# + tags=["hide"]
def convert_all(data):
"""
Convert vp or rhob from cgs to SI system.
"""
result = []
for d in data:
result.append(convert_si(d))
return result
# + tags=["hide"]
def convert_all(data):
"""
Convert vp or rhob from cgs to SI system.
"""
return [convert_si(d) for d in data]
# + tags=["hide"]
convert_all(vp)
# + tags=["hide"]
convert_all(rho)
# + tags=["hide"]
def impedance(vp, rho):
"""
Compute impedance given sequences of vp and rho.
"""
z = []
for v, r in zip(vp, rho):
z.append(v * r)
return z
# + tags=["hide"]
impedance(vp, rho)
# -
# ## Docstrings and doctests
#
# Let's add a docstring and doctests to our original function.
def convert_si(n):
"""
Convert vp or rhob from cgs to SI system.
>>> convert_si(2400)
2400
>>> convert_si(2.4)
2400.0
"""
if n < 10:
n = n * 1000
return n
import doctest
doctest.testmod()
# + [markdown] tags=["exercise"]
# <div class="alert alert-success">
# <b>Exercise</b>:
# <ul>
# <li>- Add docstrings and doctests to the functions we already wrote.</li>
# <li>- Can you rewrite your loop as a list comprehension? Make sure it still passes the tests.</li>
# <li>- Use the `convert_si` function inside your function to make sure we have the right units.</li>
# <li>- Make sure your tests still pass.</li>
# </ul>
# </div>
# + tags=["hide"]
def impedance2(vp, rho):
"""
Compute impedance given sequences of vp and rho.
>>> impedance([2300, 2400], [2450, 2350])
[5635000, 5640000]
"""
return [v*r for v, r in zip(vp, rho)]
# + tags=["hide"]
impedance(vp, rho)
# + tags=["hide"]
import doctest
doctest.testmod()
# -
# ## Compute reflection coefficients
# + [markdown] tags=["exercise"]
# <div class="alert alert-success">
# <b>Exercise</b>:
# <ul>
# <li>Can you implement the following equation?</li>
# <li>$$ \mathrm{rc} = \frac{Z_\mathrm{lower} - Z_\mathrm{upper}}{Z_\mathrm{lower} + Z_\mathrm{upper}} $$</li>
# <li>You will need to use slicing to implement the concept of upper and lower layers.</li>
# </ul>
# </div>
# -
z = impedance(vp, rho)
rc_series(z)
# You should get:
#
# [0.0004434589800443459,
# 0.04122396940076498,
# -0.021684737281067557,
# 0.10764549258273108]
# + tags=["hide"]
def compute_rc(upper, lower):
return (lower - upper) / (lower + upper)
def rc_series(z):
"""
Computes RC series.
"""
upper = z[:-1]
lower = z[1:]
rc = []
for u, l in zip(upper, lower):
rc.append(compute_rc(u, l))
return rc
# + tags=["hide"]
def rc_series2(z):
upper = z[:-1]
lower = z[1:]
return [(l-u)/(l+u) for l, u in zip(lower, upper)]
# + tags=["hide"]
rc_series2(z)
# + tags=["hide"]
# %timeit rc_series(z)
# %timeit rc_series2(z)
# + [markdown] tags=["exercise"]
# <div class="alert alert-success">
# <b>Exercise</b>:
# <ul>
# <li>Write a function to convert a slowness DT log, in microseconds per metre, into a velocity log, in m/s. </li>
# </ul>
# </div>
# -
dt = [400, 410, 420, 400, 430, 450, 440]
# +
def vp_from_dt(dt):
# Your code here!
return vp
vp = vp_from_dt(dt)
vp
# -
# You should get
#
# [2500.0,
# 2439.0243902439024,
# 2380.9523809523807,
# 2500.0,
# 2325.5813953488373,
# 2222.222222222222,
# 2272.7272727272725]
# + tags=["hide"]
def vp_from_dt(dt):
"""
Compute Vp from DT log.
Args:
dt (list): A sequence of slowness measurements.
Returns:
list. The data transformed to velocity.
TODO:
Deal with microseconds/ft.
Example:
>>> vp = vp_from_dt([400, 410])
[2500.0, 2439.0243902439024]
"""
return [1e6 / s for s in dt]
# + tags=["hide"]
vp = vp_from_dt(dt)
vp
# + tags=["hide"]
import doctest
doctest.testmod()
# + tags=["hide"]
vp_from_dt(450)
# + tags=["hide"]
def vp_from_dt(dt):
try:
v = [1e6 / s for s in dt]
except TypeError:
# Treat as scalar.
v = 1e6 / dt
return v
# + tags=["hide"]
vp_from_dt(dt)
# + tags=["hide"]
vp_from_dt(450)
# + tags=["hide"]
vp_from_dt('450')
# + [markdown] tags=["exercise"]
# <div class="alert alert-success">
# <b>Exercise</b>:
# <ul>
# <li>- Put the functions `impedance`, `rc_series`, and `vp_from_dt()` into a file called `utils.py`.</li>
# </ul>
# </div>
# -
# ## Reading data from files
#
# Go to the Reading_data_from_files notebook and do the first exercise.
# + [markdown] tags=["exercise"]
# <div class="alert alert-success">
# <b>Exercise</b>:
# <ul>
# <li>Remind yourself how you solved the problem of reading the 'tops' files in the notebook [`Ex_Reading_data_from_files.ipynb`](Ex_Reading_data_from_files.ipynb). </li>
# <li>Your challenge is to turn this into a function, complete with docstring and any options you want to try to implement. For example:</li>
# <li>- Try putting everything, including the file reading into the function. Better yet, write functions for each main 'chunk' of the workflow.</li>
# <li>- Perhaps the user can pass the number of lines to skip as a parameter.</li>
# <li>- You could also let the user choose different 'comment' characters.</li>
# <li>- Let the user select different delimiters, other than a comma.</li>
# <li>- Transforming the case of the names should probably be optional.</li>
# <li>- Print some 'progress reports' as you go, so the user knows what's going on.</li>
# <li>- As a final challenge: can you add a passing doctest? Make sure it passes on `B-41_tops.txt`.</li>
# <li>When you're done, add the function to `utils.py`.</li>
# </ul>
# </div>
# -
# + tags=["hide"]
def get_tops_from_file(fname, skip=0, comment='#', delimiter=',', null=-999.25, fix_case=True):
"""
Docstring.
>>> len(get_tops_from_file("../data/B-41_tops.txt"))
Changed depth: Upper Missisauga Fm
6
"""
with open(fname, 'r') as f:
data = f.readlines()[skip:]
tops = {}
for line in data:
# Skip comment rows.
if line.startswith(comment):
continue
# Assign names to elements.
name, dstr = line.split(delimiter)
if fix_case:
name = name.title()
dstr = dstr.strip()
if not dstr.isnumeric():
dstr = dstr.lower().rstrip('mft')
# Skip NULL entries.
if (not dstr) or (dstr == str(null)):
continue
# Correct for other negative values.
depth = float(dstr)
if depth < 0:
depth *= -1
print('Changed depth: {}'.format(name))
tops[name] = depth
return tops
# + tags=["hide"]
import doctest
doctest.testmod()
# + tags=["hide"]
tops = get_tops_from_file("../data/B-41_tops.txt")
# + tags=["hide"]
tops
| instructor/Practice_functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Spam Filters, Naive Bayes, and Wrangling
# This tutorial (ch.4) discusses how Naive Bayes could be used for spam filtering.
# \begin{equation}
# p(y|x)p(x)=p(x,y)=p(x|y)p(y)
# \end{equation}
# Using this equation:
# \begin{equation}
# p(y|x) = \frac{p(x|y)p(y)}{p(x)}
# \end{equation}
#
# - Naive Bayes is a spam filter that combines words.
# - Using a binary vector $x$ to represent an email and $c$ to denote "is spam", Naive Bayes evaluates:
# \begin{equation}
# p(x|c) = \prod_j\theta_{jc}^{x_j}(1-\theta_{jc})^{1-x_j}
# \end{equation}
# where $\theta$ is the probability that an individual word is present in a spam email.
#
# We model the words *independently* (aka *independent trials*), which is why we take the product on the righthand side of the preceding formula and don't count how many times they are present. That's why this called *"naive"*, because we know that there are actually certain words that tend to appear together, and we're ignoring this.
#
# We apply log to both sides to transform the preceding equation to summation:
# \begin{equation}
# log(p(x|c)) = \sum_jx_jlog\left(\frac{\theta_j}{1-\theta_j}\right)+\sum_jlog(1-\theta_j)
# \end{equation}
#
# The term $log(\theta_j/(1-\theta_j))$ does not depend on a given email, just the word, so let's rename it $w_j$ and assume we've computed it once and stored it. Same with quantity $\sum_j\log(1-\theta_j)=w_0$. Now we have:
# \begin{equation}
# log(p(x|c)) = \sum_jx_jw_j+w_0
# \end{equation}
#
# The accompanying code tutorial is from [here](https://dzone.com/articles/naive-bayes-tutorial-naive-bayes-classifier-in-pyt)
#
# Data
# ## step by step implementation
# Stages:
# 1. Handle data
# 2. Summarize data
# 3. Make predictions
# 4. Evaluate accuracy
# **Step 1: Handle data**
import csv
import math
import random
lines = csv.reader(open(r'diabetes.csv'))
dataset = list(lines)
len(dataset)
for i in range(1, len(dataset)):
dataset[i] = [float(x) for x in dataset[i]]
def splitDataset(dataset, splitRatio):
trainSize = int(len(dataset) * splitRatio)
trainSet = []
copy = list(dataset)
while len(trainSet) < trainSize:
index = random.randrange(len(copy))
trainSet.append(copy.pop(index))
return [trainSet, copy]
# **Step 2: Summarize the data**
def separateByClass(dataset):
separated = {}
for i in range(len(dataset)):
vector = dataset[i]
if (vector[-1] not in separated):
separated[vector[-1]] = []
separated[vector[-1]].append(vector)
return separated
dataset.pop(0)
print("classes:{}".format(list(separateByClass(dataset).keys())))
def mean(numbers):
return sum(numbers)/float(len(numbers))
# calculate standard deviation
def stdev(numbers):
avg = mean(numbers)
variance = sum([pow(x-avg, 2) for x in numbers])/float(len(numbers)-1)
return math.sqrt(variance)
def summarize(dataset):
summaries = [(mean(attribute), stdev(attribute)) for attribute in zip(*dataset)]
del summaries[-1]
return summaries
summarize(dataset)
def summarizeByClass(dataset):
separated = separateByClass(dataset)
summaries = {}
for classValue, instances in separated.items():
summaries[classValue] = summarize(instances)
return summaries
summarizeByClass(dataset)
# **Step 3: Making predictions**
# calculate the Gaussian probability density function
def calculateProbability(x, mean, stdev):
exponent = math.exp(-(math.pow(x-mean, 2)/(2 * math.pow(stdev, 2))))
return (1/(math.sqrt(2*math.pi)*stdev))*exponent
# calculate class probabilities
def calculateClassProbabilities(summaries, inputVector):
probabilities = {}
for classValue, classSummaries in summaries.items():
probabilities[classValue] = 1
for i in range(len(classSummaries)):
mean, stdev = classSummaries[i]
x = inputVector[i]
probabilities[classValue] *= calculateProbability(x, mean, stdev)
return probabilities
def predict(summaries, inputVector):
probabilities = calculateClassProbabilities(summaries, inputVector)
bestLabel, bestProb = None, -1
for classValue, probability in probabilities.items():
print(classValue, probability)
if bestLabel is None or probability > bestProb:
bestProb = probability
bestLabel = classValue
return bestLabel
def getPredictions(summaries, testSet):
predictions = []
for i in range(len(testSet)):
result = predict(summaries, testSet[i])
predictions.append(result)
return predictions
def getAccuracy(testSet, predictions):
correct = 0
for i in range(len(testSet)):
if testSet[i][-1] == predictions[i]:
correct += 1
return (correct/float(len(testSet))) * 100.0
# **Putting it all together**
trainingSet, testSet = splitDataset(dataset, 0.67)
len(trainingSet)
len(testSet)
print('Split {0} rows into train = {1} and test = {2} rows'.format(len(dataset),len(trainingSet),len(testSet)))
summaries = summarizeByClass(trainingSet)
summaries
testSet[0]
predictions = getPredictions(summaries, [testSet[0]])
len(predictions)
accuracy = getAccuracy(testSet, predictions)
print('Accuracy: {}%'.format(accuracy))
# ## Naive Bayes using Scikit-Learn
from sklearn import datasets
from sklearn import metrics
from sklearn.naive_bayes import GaussianNB, BernoulliNB
dataset = datasets.load_iris()
model = GaussianNB()
model.fit(dataset.data, dataset.target)
expected = dataset.target
len(expected)
predicted = model.predict(dataset.data)
len(predicted)
print(metrics.classification_report(expected, predicted))
print(metrics.confusion_matrix(expected, predicted))
bernModel = BernoulliNB()
bernModel.fit(dataset.data, dataset.target)
predicted = bernModel.predict(dataset.data)
print(metrics.classification_report(expected, predicted))
print(metrics.confusion_matrix(expected, predicted))
| NaiveBayes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: CUDAtorch
# language: python
# name: cudatorch
# ---
import pandas as pd
import torch
import pytorch_lightning as pl
from tqdm import tqdm
import torchmetrics
import math
from urllib.request import urlretrieve
from zipfile import ZipFile
import os
import torch.nn as nn
import numpy as np
# ## Settings
WINDOW_SIZE = 20
# ## Data
urlretrieve("http://files.grouplens.org/datasets/movielens/ml-1m.zip", "movielens.zip")
ZipFile("movielens.zip", "r").extractall()
# +
users = pd.read_csv(
"ml-1m/users.dat",
sep="::",
names=["user_id", "sex", "age_group", "occupation", "zip_code"],
)
ratings = pd.read_csv(
"ml-1m/ratings.dat",
sep="::",
names=["user_id", "movie_id", "rating", "unix_timestamp"],
)
movies = pd.read_csv(
"ml-1m/movies.dat", sep="::", names=["movie_id", "title", "genres"]
)
# +
## Movies
movies["year"] = movies["title"].apply(lambda x: x[-5:-1])
movies.year = pd.Categorical(movies.year)
movies["year"] = movies.year.cat.codes
## Users
users.sex = pd.Categorical(users.sex)
users["sex"] = users.sex.cat.codes
users.age_group = pd.Categorical(users.age_group)
users["age_group"] = users.age_group.cat.codes
users.occupation = pd.Categorical(users.occupation)
users["occupation"] = users.occupation.cat.codes
users.zip_code = pd.Categorical(users.zip_code)
users["zip_code"] = users.zip_code.cat.codes
#Ratings
ratings['unix_timestamp'] = pd.to_datetime(ratings['unix_timestamp'],unit='s')
# +
# Save primary csv's
if not os.path.exists('data'):
os.makedirs('data')
users.to_csv("data/users.csv",index=False)
movies.to_csv("data/movies.csv",index=False)
ratings.to_csv("data/ratings.csv",index=False)
# +
## Movies
movies["movie_id"] = movies["movie_id"].astype(str)
## Users
users["user_id"] = users["user_id"].astype(str)
##Ratings
ratings["movie_id"] = ratings["movie_id"].astype(str)
ratings["user_id"] = ratings["user_id"].astype(str)
# + id="6_CC3yYCLxVN"
genres = [
"Action",
"Adventure",
"Animation",
"Children's",
"Comedy",
"Crime",
"Documentary",
"Drama",
"Fantasy",
"Film-Noir",
"Horror",
"Musical",
"Mystery",
"Romance",
"Sci-Fi",
"Thriller",
"War",
"Western",
]
for genre in genres:
movies[genre] = movies["genres"].apply(
lambda values: int(genre in values.split("|"))
)
# + [markdown] id="0KsqW_4rLxVN"
# ### Transform the movie ratings data into sequences
#
# First, let's sort the the ratings data using the `unix_timestamp`, and then group the
# `movie_id` values and the `rating` values by `user_id`.
#
# The output DataFrame will have a record for each `user_id`, with two ordered lists
# (sorted by rating datetime): the movies they have rated, and their ratings of these movies.
# + id="D5v700zTLxVN"
ratings_group = ratings.sort_values(by=["unix_timestamp"]).groupby("user_id")
ratings_data = pd.DataFrame(
data={
"user_id": list(ratings_group.groups.keys()),
"movie_ids": list(ratings_group.movie_id.apply(list)),
"ratings": list(ratings_group.rating.apply(list)),
"timestamps": list(ratings_group.unix_timestamp.apply(list)),
}
)
# + [markdown] id="USa6rk0eLxVN"
# Now, let's split the `movie_ids` list into a set of sequences of a fixed length.
# We do the same for the `ratings`. Set the `sequence_length` variable to change the length
# of the input sequence to the model. You can also change the `step_size` to control the
# number of sequences to generate for each user.
# + id="XdhRJlxULxVN"
sequence_length = 8
step_size = 1
def create_sequences(values, window_size, step_size):
sequences = []
start_index = 0
while True:
end_index = start_index + window_size
seq = values[start_index:end_index]
if len(seq) < window_size:
seq = values[-window_size:]
if len(seq) == window_size:
sequences.append(seq)
break
sequences.append(seq)
start_index += step_size
return sequences
ratings_data.movie_ids = ratings_data.movie_ids.apply(
lambda ids: create_sequences(ids, sequence_length, step_size)
)
ratings_data.ratings = ratings_data.ratings.apply(
lambda ids: create_sequences(ids, sequence_length, step_size)
)
del ratings_data["timestamps"]
# + [markdown] id="5dYEduaqLxVN"
# After that, we process the output to have each sequence in a separate records in
# the DataFrame. In addition, we join the user features with the ratings data.
# + id="gM5_RBACLxVO"
ratings_data_movies = ratings_data[["user_id", "movie_ids"]].explode(
"movie_ids", ignore_index=True
)
ratings_data_rating = ratings_data[["ratings"]].explode("ratings", ignore_index=True)
ratings_data_transformed = pd.concat([ratings_data_movies, ratings_data_rating], axis=1)
ratings_data_transformed = ratings_data_transformed.join(
users.set_index("user_id"), on="user_id"
)
ratings_data_transformed.movie_ids = ratings_data_transformed.movie_ids.apply(
lambda x: ",".join(x)
)
ratings_data_transformed.ratings = ratings_data_transformed.ratings.apply(
lambda x: ",".join([str(v) for v in x])
)
del ratings_data_transformed["zip_code"]
ratings_data_transformed.rename(
columns={"movie_ids": "sequence_movie_ids", "ratings": "sequence_ratings"},
inplace=True,
)
# + [markdown] id="YnxbQ-wQLxVO"
# With `sequence_length` of 4 and `step_size` of 2, we end up with 498,623 sequences.
#
# Finally, we split the data into training and testing splits, with 85% and 15% of
# the instances, respectively, and store them to CSV files.
# + id="0lPMjBoRLxVO"
random_selection = np.random.rand(len(ratings_data_transformed.index)) <= 0.85
train_data = ratings_data_transformed[random_selection]
test_data = ratings_data_transformed[~random_selection]
train_data.to_csv("data/train_data.csv", index=False, sep=",")
test_data.to_csv("data/test_data.csv", index=False, sep=",")
# -
test_data
| prepare_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Python NRT
# ## (Not a Real Tutorial)
#
# <br>A brief brief tour around</br>
# Python 2.7 & Pandas library
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Python language
#
# + [markdown] slideshow={"slide_type": "subslide"}
# * Good language to start programming with
# * Simple, powerful, mature
# * Easy to read, intuitive
# + [markdown] slideshow={"slide_type": "fragment"}
# ```
# >>> print "how "+"are you?"
# how are you?
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# ## Running Python
# + [markdown] slideshow={"slide_type": "subslide"}
# * From Python console
# + [markdown] slideshow={"slide_type": "fragment"}
# ```
# $ python
# Python 2.7.12 (default, Jun 29 2016, 14:05:02)
# [GCC 4.2.1 Compatible Apple LLVM 7.3.0 (clang-703.0.31)] on darwin
# Type "help", "copyright", "credits" or "license" for more information.
# >>>
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# * Run a python script
# + [markdown] slideshow={"slide_type": "fragment"}
# ```
# $ python myprogram.py
# ```
# + [markdown] slideshow={"slide_type": "fragment"}
# * Use an interactive web console like Jupyter
# + [markdown] slideshow={"slide_type": "fragment"}
# ```
# $ jupyter notebook
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# ## Syntax
# + [markdown] slideshow={"slide_type": "subslide"}
# * No termination character
# + slideshow={"slide_type": "fragment"}
name = "Pepe"
# + [markdown] slideshow={"slide_type": "subslide"}
# * Blocks specified by indentation (not braces nor brackets)
# * First statement of a block ends in colon (:)
# + slideshow={"slide_type": "fragment"}
def myfunction(x):
pass
if x > 10:
pass
pass
return "bigger"
else:
pass
pass
return "smaller"
print "This number is: " + myfunction(5)
# + [markdown] slideshow={"slide_type": "subslide"}
# * Comments use number/hash symbol (#) and triple quotes (""")
# + slideshow={"slide_type": "fragment"}
"""This is a comment that spands for
more than one line"""
# This is a one line comment
print "This line is executed"
# + [markdown] slideshow={"slide_type": "slide"}
# ## Modules
# + slideshow={"slide_type": "fragment"}
import pandas as pd
from time import clock
# + [markdown] slideshow={"slide_type": "slide"}
# ## Lists and selections
# + slideshow={"slide_type": "subslide"}
months = ["Jan", "Feb", 3, 4, "May", "Jun"]
print months[0]
# + slideshow={"slide_type": "fragment"}
print months[1:3] # slice operator :
# + slideshow={"slide_type": "fragment"}
print months[-2:]
# + [markdown] slideshow={"slide_type": "slide"}
# ## Tuples
# + [markdown] slideshow={"slide_type": "subslide"}
# Similar to lists, sequence of elements that conforms an immutable object.
# + slideshow={"slide_type": "fragment"}
tup = ('physics', 'chemistry', 1997, 2000)
print tup[0]
# + slideshow={"slide_type": "fragment"}
print tup[1:3]
# + [markdown] slideshow={"slide_type": "slide"}
# ## Functions & Methods
# + slideshow={"slide_type": "subslide"}
"""functions are pieces of code that you can
call/execute, they are defined with the def keyword"""
def hola_mundo():
print "Hola Mundo!"
# + slideshow={"slide_type": "subslide"}
""" methods are attributes of an object that
you can call over the object with and "." """
s = "How are you"
print s.split(" ")
# + [markdown] slideshow={"slide_type": "slide"}
# ## Control flow
# + [markdown] slideshow={"slide_type": "subslide"}
# * Loops (while, for)
# + slideshow={"slide_type": "fragment"}
for numbers in range(1,5):
print numbers
# + [markdown] slideshow={"slide_type": "subslide"}
# * Conditionals (if, elif, else)
# + slideshow={"slide_type": "fragment"}
united_kingdom = ["England", "Scotland", "Wales", "N Ireland"]
one = "France"
if one in united_kingdom:
print "UK"
elif one == "France":
print "Not UK. Bon jour!"
else:
print "Not UK"
# + [markdown] slideshow={"slide_type": "slide"}
# ## Help!
# + [markdown] slideshow={"slide_type": "subslide"}
# # "house".len()?
#
# # len(house)?
# + slideshow={"slide_type": "fragment"}
help(len)
# + slideshow={"slide_type": "fragment"}
len("house")
# + slideshow={"slide_type": "subslide"}
help(list)
# + [markdown] slideshow={"slide_type": "slide"}
# # PANDAS LIBRARY
# + [markdown] slideshow={"slide_type": "subslide"}
# Open source library providing high-performance **structures** and **data analysis tools** for the Python programming language.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Import
# + slideshow={"slide_type": "fragment"}
import pandas as pd
# + [markdown] slideshow={"slide_type": "slide"}
# ## Structures
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Pandas Series
# + slideshow={"slide_type": "fragment"}
ss = pd.Series([1,2,3],
index = ['a','b','c'])
ss
# + [markdown] slideshow={"slide_type": "subslide"}
# Selection
# + slideshow={"slide_type": "fragment"}
ss = pd.Series([1,2,3],
index = ['a','b','c'])
print ss[0] # as a list
print ss.iloc[0] # by position, integer
print ss.loc['a'] # by label of the index
print ss.ix['a'] # label (priority)
print ss.ix[0] # position if no label
# + slideshow={"slide_type": "subslide"}
"""Be careful with the slice operator
using positions or labels"""
print ss.iloc[0:2] # positions 0,1
print ss.loc['a':'c'] # labels 'a','b','c'
# + [markdown] slideshow={"slide_type": "subslide"}
# Built-in methods
# + slideshow={"slide_type": "fragment"}
pd.Series([1, 2, 3]).mean()
# + slideshow={"slide_type": "fragment"}
pd.Series([1, 2, 3]).sum()
# + slideshow={"slide_type": "fragment"}
pd.Series([1, 2, 3]).std()
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Pandas Dataframe
# + slideshow={"slide_type": "fragment"}
df = pd.DataFrame(
data =[[1, 2, 3], [4, 5, 6], [7, 8, 9]],
index=['row1', 'row2', 'row3'],
columns=['col1', 'col2', 'col3'])
df
# + [markdown] slideshow={"slide_type": "subslide"}
# Selection
# + [markdown] slideshow={"slide_type": "subslide"}
# Select columns
# + slideshow={"slide_type": "fragment"}
df['col1'] # one col => Series
# + slideshow={"slide_type": "fragment"}
df[['col1']] # list of cols => DataFrame
# + [markdown] slideshow={"slide_type": "subslide"}
# Select rows
# + slideshow={"slide_type": "subslide"}
df.loc['row1'] # by row using label
# + slideshow={"slide_type": "fragment"}
df.iloc[0] # by row using position
# + slideshow={"slide_type": "fragment"}
df.ix['row1'] # by row, using label
print df.ix[0] # by row, using position
# + [markdown] slideshow={"slide_type": "subslide"}
# Combined selection
# + slideshow={"slide_type": "fragment"}
print df.loc['row1',['col1', 'col3']] # labels
print df.loc[['row1','row3'],'col1' : 'col3']
# + slideshow={"slide_type": "fragment"}
df.iloc[0:2,[0,2]] # row position 0,1
# + slideshow={"slide_type": "subslide"}
print df.ix[0,['col2','col3']] # position & label
print df.ix['row1':'row3', :]
# + [markdown] slideshow={"slide_type": "fragment"}
# Should I use always .ix()?
# + [markdown] slideshow={"slide_type": "subslide"}
# .ix() selector gotcha!
# + slideshow={"slide_type": "fragment"}
df2 = pd.DataFrame(
data =[[1, 2, 3], [4, 5, 6], [7, 8, 9]],
index=[1, 2, 3],
columns=['col1', 'col2', 'col3'])
print df2.ix[1] # priority is label
# df2.ix[0] ERROR!!
# + slideshow={"slide_type": "subslide"}
df2 = pd.DataFrame(
data =[[1, 2, 3], [4, 5, 6], [7, 8, 9]],
index=[1, 2, 3],
columns=['col1', 'col2', 'col3'])
print df2.ix[1:3] # LABELS!! (1,2,3)
# + slideshow={"slide_type": "subslide"}
# these two dataframes are the same!!
df2 = pd.DataFrame(
data =[[1, 2, 3], [4, 5, 6], [7, 8, 9]],
index=[1, 2, 3],
columns=[1, 2, 3])
df3 = pd.DataFrame(
data =[[1, 2, 3], [4, 5, 6], [7, 8, 9]])
df3
# + [markdown] slideshow={"slide_type": "subslide"}
# DataFrame Selection Summary
# + slideshow={"slide_type": "fragment"}
df['col1'] # by columns
df.loc['row1'] # by row, using label
df.iloc[0] # by row, using position
df.ix['row2'] # by row, using label
df.ix[1] # by row, using position
# + [markdown] slideshow={"slide_type": "subslide"}
# Built-in method
# + slideshow={"slide_type": "fragment"}
df.mean() # operates by columns (axis=0)
# + [markdown] slideshow={"slide_type": "subslide"}
# Pandas Axis
# + [markdown] slideshow={"slide_type": "fragment"}
# | axis | axis | along | each |
# |--------|:--------------:|------------------:|---------------:|
# | axis=1 | axis="columns" | along the columns |for each row |
# | axis=0 | axis="index" | along the rows |for each column |
#
#
# + slideshow={"slide_type": "subslide"}
df2 = pd.DataFrame(
data =[[1, 2], [4, 5], [7, 8]],
columns=["A", "B"])
df2
# + slideshow={"slide_type": "fragment"}
df2.mean(axis=1) # mean for each row
# + slideshow={"slide_type": "subslide"}
df2 = pd.DataFrame(
data =[[1, 2], [4, 5], [7, 8]],
columns=["A", "B"])
df2
# + slideshow={"slide_type": "fragment"}
df2.drop("A", axis=1) # drop columns for each row
# + [markdown] slideshow={"slide_type": "slide"}
# ## Let's do it!!
| Python_NRT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
num=5
for i in range(0,num):
for j in range(0,i+1):
print("*",end=" ")
print()
# -
import random
random.randrange(3,10)
random.randrange(20,30)
random.random()
random.uniform(2,6)
a=random.uniform(2,6)
a
def fun(a,b):
return random.uniform(a,b)
fun(4,6)
import numpy as np
np.array()
L=[10,20,30,40,50]
T=(100,200,300,400,500)
S="Today is a very beautiful day"
random.choice(S)
random.sample(L,3)
random.sample(T,4)
random.shuffle(L)
L
random.shuffle(T)
random.shuffle(S)
# +
val1=int(input("Enter the first value:"))
val2=int(input("Enter the second value:"))
if val1*val2<100:
print(val1*val2)
else:
print(val1+val2)
# +
val1=int(input("Enter the first value:"))
val2=int(input("Enter the second value:"))
if val1*val2<100:
print(val1*val2)
else:
print(val1+val2)
# -
L=[2,3,4,5]
L1=[2,6,12,20]
def fun(L):
newL=[L[0] if i==0 else L[i]*L[i-1] for i in range(len(L))]
return newL
def fun1(L):
newL=[L[0] if i==0 else L[i]*L[i-1] for i,j in enumerate(L)]
return newL
fun([10,20,30])
fun1([10,20,30])
def fun2(L):
newL=[]
for i in range(len(L)):
if i==0:
newL.append(L[0])
else:
newL.append(L[i]*L[i-1])
fun([10,20,30])
fun1([10,20,30])
fun2([10,20,30])
def fun(L):
newL=[L[i] for i in range(len(L)) if i%2==0]
return sum(newL)
L=[1,3,4,1,5,6,7,3,8]
fun(L)
def fun1(L):
if L[0]==L[-1]:
return True
else:
return False
def fun2(L):
return L[0]==L[-1]
fun1([2,3,3,8])
fun2([2,3,3,8])
L=[2,4,5,1,-9,8/9]
L[::-1]
list(reversed(L))
def fun(L):
return L==list(reversed(L))
fun([1,2,3,2,1])
fun([-1,3,4,6,7/8])
def fun(L1,L2):
return[i for i in L1 if i%3==0]+[j for j in L2 if j%5==0]
fun([2,3,6,12,-9,8/9],[5,3,4,-1,5/6])
(lambda L1,L2:[i for i in L1 if i%3==0]+[j for j in L2 if j%5==0])([2,3,6,12,-9,8/9],[5,3,4,-1,5/6])
def fun(L):
return len(L)>=1 and (L[0]==6 or L[-1]==6)
fun([])
fun([-1])
fun([0])
| Online Python workshop webinar session Day 08.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas as pd
from textblob.classifiers import NaiveBayesClassifier
from sklearn.cross_validation import train_test_split
# pos = pd.read_csv('positive.csv')
# neg = pd.read_csv('negative.csv')
# p = pos
# n = neg
# res = pd.concat([p,n])
# res.to_csv("headlines.csv", index=False, header=False)
headlines = pd.read_csv('headlines.csv')
# +
# train, test = train_test_split(headlines,test_size=0.2,random_state=0)
# train.to_csv("train.csv", index=False, header=False)
# test.to_csv("test.csv", index=False, header=False)
# -
with open("train.csv", "r") as f:
cl = NaiveBayesClassifier(f, format='csv')
with open("test.csv", "r") as f:
print cl.accuracy(f, format='csv')
h = pd.read_csv('headlines.csv')
train, test = train_test_split(h, test_size=0.2, random_state=0)
| Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + slideshow={"slide_type": "skip"}
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# + slideshow={"slide_type": "skip"}
def clean_ax(ax, x_range, y_range, spines=False, legend=False):
if legend:
ax.legend(loc="upper left")
if not spines:
ax.spines["top"].set_visible(False)
ax.spines["bottom"].set_visible(False)
ax.spines["left"].set_visible(False)
ax.spines["right"].set_visible(False)
plt.setp(ax, xticks=(), yticks=(), xlim=x_range, ylim=y_range)
# + [markdown] slideshow={"slide_type": "slide"}
# # Dynamical Movement Primitive (DMPs)
# + [markdown] slideshow={"slide_type": "subslide"}
# DMPs are trajectory representations that
#
# * are goal-directed
# * are robust to perturbations and noise (like potential fields)
# * are arbitrarily shapeable
# * can be scaled and translated arbitrarily, even online
# * are parametrized policy representations that can be used for reinforcement learning
# * can be used for imitation learning
# + [markdown] slideshow={"slide_type": "subslide"}
# They have drawbacks:
#
# * an inverse (velocity) kinematics is required to execute task space trajectories
# * they are usually not able to learn from multiple demonstrations
# + [markdown] slideshow={"slide_type": "subslide"}
# There are different variants of DMPs in the literature [1, 2, 3].
#
# [1] Ijspeert, <NAME>; <NAME>; <NAME>; <NAME>; <NAME>: Dynamical movement primitives: learning attractor models for motor behaviors, Neural computation 25 (2), pp. 328-373, 2013.
#
# [2] <NAME>; <NAME>; <NAME>; <NAME>: Learning and generalization of motor skills by learning from demonstration, Proceedings of the IEEE International Conference on Robotics and Automation, pp. 763-768, 2009.
#
# [3] <NAME>; <NAME>; <NAME>; <NAME>: Learning to Select and Generalize Striking Movements in Robot Table Tennis, International Journal of Robotics Research 32 (3), 2013.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Dynamical System
# + [markdown] slideshow={"slide_type": "subslide"}
# A fixed rule describes how a point in geometrical space depends on time.
#
# * At any time $t$, a dynamical system has a state $\boldsymbol{x} \in S \subseteq \mathbb{R}^n$
# * Small changes in the systems state create small changes in $\boldsymbol{x}$
# * The evolution rule $f(\boldsymbol{x}_t) = \boldsymbol{x}_{t+1}$ is a fixed, deterministic rule that describes what future states follow from the current state
# + [markdown] slideshow={"slide_type": "slide"}
# ## Mass-spring-damper
# + [markdown] slideshow={"slide_type": "subslide"}
# 
# + [markdown] slideshow={"slide_type": "slide"}
# * An oscillatory force $F_s = -k x$
# * and a damping force $F_d = -c \dot{x}$
# * result in a total force $F_{tot} = m \ddot{x}$
#
# $$F_{tot} = F_s + F_d \Leftrightarrow \ddot{x} + \frac{c}{m} \dot{x} + \frac{k}{m} x = 0$$
#
# The system is **critically damped**, i.e. it converges to zero as fast as possible without oscillating (overshoot can occur), iff
# $$c = 2 \sqrt{mk} \Leftrightarrow \zeta = \frac{c}{2 \sqrt{mk}} = 1$$
# It is **over-damped** (too slow) iff
# $$\zeta > 1$$
# and **under-damped** (oscillating) iff
# $$0 \leq \zeta < 1$$
#
# [Wikipedia: Damping](http://en.wikipedia.org/wiki/Damping)
# + [markdown] slideshow={"slide_type": "slide"}
# The dynamical system that we use in a Dynamical Movement Primitive is based on a spring-damper system with $m=1$. The original formulation by Ijspeert et al. computes the acceleration by
# $$\ddot{x} = \frac{\alpha}{\tau^2} \left(\beta (g - x) - \tau \dot{x}\right),$$
# which can be reformulated as
# $$x + \frac{\tau}{\beta} \dot{x} + \frac{\tau^2}{\alpha \beta} \ddot{x} = g,$$
# so that critical damping is achieved with
# $$\frac{\tau}{\beta} = 2 \sqrt{\frac{\tau^2}{\alpha \beta}}
# \Leftrightarrow \frac{\alpha}{4} = \beta.$$
# + slideshow={"slide_type": "slide"}
def spring_damper(x0, g, tau, dt, alpha, beta, observe_for):
X = [x0]
xd = 0.0
xdd = 0.0
t = 0.0
while t < observe_for:
X.append(X[-1] + xd * dt)
x = X[-1]
xd += xdd * dt
xdd = alpha / (tau ** 2) * (beta * (g - x) - tau * xd)
t += dt
return X
x0, g = 0.0, 1.0
tau = 1.0
observe_for = 2.0 * tau
dt = 0.01
plt.figure(figsize=(10, 5))
plt.xlabel("Time")
plt.ylabel("Position")
plt.xlim((0.0, observe_for))
diff = g - x0
plt.ylim((x0 - 0.1 * diff, g + 0.5 * diff))
for alpha, beta in [(25.0, 6.25), (25.0, 1.5), (25.0, 25.0)]:
X = spring_damper(x0, g, tau, dt, alpha, beta, observe_for)
plt.plot(np.arange(0.0, observe_for + dt, dt), X, lw=3,
label="$\\alpha = %g,\\quad \\beta = %g$" % (alpha, beta))
plt.scatter([0.0, tau], [x0, g], marker="*", s=500, label="$x_0, g$")
plt.legend(loc="lower right")
# + [markdown] slideshow={"slide_type": "slide"}
# In addition, the DMP has a nonlinear forcing term that defines the shape of the movement over time.
# $$\tau^2 \ddot{x} = \alpha \left(\beta (g - x) - \tau \dot{x}\right) + f(s),$$
# where $s$ is the so called phase.
# + slideshow={"slide_type": "slide"}
from dmp import DMP, trajectory, potential_field
x0 = np.array([0, 0], dtype=np.float64)
g = np.array([1, 1], dtype=np.float64)
tau = 1.0
w = np.array([[-50.0, 100.0, 300.0],
[-200.0, -200.0, -200.0]])
o = np.array([1.0, 0.5])
dt = 0.01
dmp = DMP()
x_range = (-0.2, 1.2)
y_range = (-0.2, 1.2)
n_tics = 10
G, _, _ = trajectory(dmp, w, x0, g, tau, dt, o, shape=False, avoidance=False)
T, _, _ = trajectory(dmp, w, x0, g, tau, dt, o, shape=True, avoidance=False)
O, _, _ = trajectory(dmp, w, x0, g, tau, dt, o, shape=True, avoidance=True)
fig = plt.figure(figsize=(5, 5))
plt.plot(G[:, 0], G[:, 1], lw=3, color="g", label="Goal-directed")
plt.plot(T[:, 0], T[:, 1], lw=3, color="r", label="Shaped")
plt.plot(O[:, 0], O[:, 1], lw=3, color="black", label="Obstacle avoidance")
plt.plot(x0[0], x0[1], "o", color="b", markersize=10)
plt.plot(g[0], g[1], "o", color="g", markersize=10)
plt.plot(o[0], o[1], "o", color="y", markersize=10)
clean_ax(plt.gca(), x_range=x_range, y_range=y_range, legend=True)
# + slideshow={"slide_type": "skip"}
def potential_trajectory(dmp, t_max, dt, shape, avoidance):
T, Td, _ = trajectory(dmp, w, x0, g, tau, dt, o, shape, avoidance)
X, Y, sd, f, C, acc = potential_field(
dmp, t_max, Td[t_max - 1],
w, x0, g, tau, dt, o, x_range, y_range, n_tics)
if not avoidance:
acc -= C
return T[:t_max], X, Y, sd, f, C, acc
# + slideshow={"slide_type": "slide"}
fig = plt.figure(figsize=(10, 10))
plt.subplots_adjust(left=0.0, bottom=0.0, right=1.0, top=1.0, wspace=0.01, hspace=0.01)
for i, t in enumerate([1, 5, 10, 15, 20, 25, 30, 50, 99]):
ax = plt.subplot(3, 3, 1 + i, aspect="equal")
T, X, Y, sd, f, C, acc = potential_trajectory(
dmp, t, dt, shape=True, avoidance=True)
plt.plot(T[:, 0], T[:, 1], lw=5, color="black")
quiver_scale = np.abs(acc).max() * n_tics
plt.quiver(X, Y, sd[:, :, 0], sd[:, :, 1], scale=quiver_scale, color="g")
plt.quiver(X, Y, f[:, :, 0], f[:, :, 1], scale=quiver_scale, color="r")
plt.quiver(X, Y, C[:, :, 0], C[:, :, 1], scale=quiver_scale, color="y")
plt.quiver(X, Y, acc[:, :, 0], acc[:, :, 1], scale=quiver_scale, color="black")
plt.plot(x0[0], x0[1], "o", color="b", markersize=10)
plt.plot(g[0], g[1], "o", color="g", markersize=10)
plt.plot(o[0], o[1], "o", color="y", markersize=10)
clean_ax(ax, x_range=x_range, y_range=y_range, spines=True, legend=False)
# + [markdown] slideshow={"slide_type": "skip"}
# Dependencies
#
# pip install ipywidgets
# + slideshow={"slide_type": "slide"}
import ipywidgets
def plot(step):
t = int(step)
fig = plt.figure(figsize=(7, 7))
ax = fig.add_subplot(111)
T, X, Y, sd, f, C, acc = potential_trajectory(
dmp, t, dt, shape=True, avoidance=True)
plt.plot(T[:, 0], T[:, 1], lw=5, color="black")
quiver_scale = np.abs(acc).max() * n_tics
plt.quiver(X, Y, sd[:, :, 0], sd[:, :, 1], scale=quiver_scale, color="g")
plt.quiver(X, Y, f[:, :, 0], f[:, :, 1], scale=quiver_scale, color="r")
plt.quiver(X, Y, C[:, :, 0], C[:, :, 1], scale=quiver_scale, color="y")
plt.quiver(X, Y, acc[:, :, 0], acc[:, :, 1], scale=quiver_scale, color="black")
plt.plot(x0[0], x0[1], "o", color="b", markersize=10)
plt.plot(g[0], g[1], "o", color="g", markersize=10)
plt.plot(o[0], o[1], "o", color="y", markersize=10)
clean_ax(ax, x_range=x_range, y_range=y_range, spines=True, legend=False)
return fig
ipywidgets.interact(plot, step=ipywidgets.IntSlider(min=5, max=95, step=5, value=5));
# + [markdown] slideshow={"slide_type": "slide"}
# ## Imitation Learning
#
# $$\tau^2 \ddot{x} = \alpha \left(\beta (g - x) - \tau \dot{x}\right) + f(s)$$
# can be rearranged to
# $$f(s) = \tau^2 \ddot{x} - \alpha \left(\beta (g - x) - \tau \dot{x}\right)$$
# + [markdown] slideshow={"slide_type": "slide"}
# TODO
# ====
#
# * imitation learning
# * Muelling's DMP
# * goal adaption
# * parametric DMPs
# * episodic REPS
| DMP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import requests
from scipy.stats import norm
from random import uniform
from mpl_toolkits.mplot3d import Axes3D
# +
def f(x,y):
return np.cos(x**2 + y**2)
num_rows, num_cols = 0,0
fig, axes = plt.subplots(num_rows, num_cols, figsize=(16,12))
for i in range(num_rows):
for j in range(num_cols):
x = np.linspace(-3,3,100)
y = x
axes[i, j].scatter(x, f(x,y), alpha=0.6)
t = f'$i = {i}, \quad j = {j}$'
axes[i, j].set(title=t, xticks=[-3,0,3], yticks=[-3,0,3])
# +
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.cm as cm
grid = np.linspace(-3,3,100)
x, y = np.meshgrid(grid, grid)
z = np.cos((x**2)+(y**2))
m = np.asarray(z)
plt.imshow(m, cmap=cm.hot)
plt.colorbar()
plt.show()
# +
# %matplotlib notebook
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.cm as cm
from mpl_toolkits.mplot3d import Axes3D
res=200
grid = np.linspace(-3,3,res)
x, y = np.meshgrid(grid,grid)
color = np.empty([res,res,res])
for i in range(res):
z = np.cos((x**2)+(y**2)+((((i/res)*6)-3)**2))
color[i] = z
color.shape=(res**3)
fig = plt.figure(figsize=(8,8))
ax = fig.add_subplot(111, projection='3d')
X, Y, Z = np.meshgrid(grid, grid, grid)
ax.scatter(X, Y, Z,
c=color, cmap=cm.hot,
alpha=0.5, s=0.01)
# -
| Exercises/QuantEcon/Packages/heatmap examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Spatio-temporal Transcriptomics
#
# Toy dataset from López-Lopera et al. (2019)
#
# - Data download: https://github.com/anfelopera/PhysicallyGPDrosophila
# + pycharm={"name": "#%%\n"}
import numpy as np
import torch
from torch.nn import Parameter
from matplotlib import pyplot as plt
from mpl_toolkits.axes_grid1 import ImageGrid
from pathlib import Path
from scipy.interpolate import interp1d
from torch.optim import Adam
from gpytorch.optim import NGD
from alfi.models import MultiOutputGP, PartialLFM, generate_multioutput_rbf_gp
from alfi.models.pdes import ReactionDiffusion
from alfi.utilities.data import dros_ground_truth
from alfi.utilities.fenics import interval_mesh
from alfi.datasets import DrosophilaSpatialTranscriptomics, HomogeneousReactionDiffusion
from alfi.trainers import PartialPreEstimator, PDETrainer
from alfi.plot import plot_spatiotemporal_data, Plotter1d
from alfi.utilities.torch import softplus, inv_softplus
from alfi.configuration import VariationalConfiguration
# + pycharm={"name": "#%%\n"}
drosophila = False
if drosophila:
gene = 'kr'
dataset = DrosophilaSpatialTranscriptomics(
gene=gene, data_dir='../../../data', scale=True, disc=20)
params = dict(lengthscale=10,
**dros_ground_truth(gene),
parameter_grad=False,
warm_epochs=-1,
natural=True,
zero_mean=True,
clamp=True)
disc = dataset.disc
else:
data = 'toy-spatial'
dataset = HomogeneousReactionDiffusion(data_dir='../../../data')
params = dict(lengthscale=0.2,
sensitivity=1,
decay=0.1,
diffusion=0.01,
warm_epochs=-1,
dp=0.025,
natural=True,
clamp=False)
disc = 1
data = next(iter(dataset))
tx, y_target = data
lengthscale = params['lengthscale']
zero_mean = params['zero_mean'] if 'zero_mean' in params else False
# + [markdown] pycharm={"name": "#%% md\n"}
# We can either create a simple unit interval mesh
# + pycharm={"name": "#%%\n"}
from dolfin import *
mesh = UnitIntervalMesh(40)
plot(mesh)
# -
# Alternatively, if our spatial data is not uniformly spaced, we can define a custom mesh as follows.
# + pycharm={"name": "#%%\n"}
# We calculate a mesh that contains all possible spatial locations in the dataset
spatial = np.unique(tx[1, :])
mesh = interval_mesh(spatial)
plot(mesh)
# The mesh coordinates should match up to the data:
print('Matching:', (spatial == mesh.coordinates().reshape(-1)).all())
# + [markdown] pycharm={"name": "#%% md\n"}
# Set up GP model
# + pycharm={"name": "#%%\n"}
# Define GP
if tx.shape[1] > 1000:
num_inducing = int(tx.shape[1] * 3/6)
else:
num_inducing = int(tx.shape[1] * 5/6)
use_lhs = True
if use_lhs:
print('tx', tx.shape)
from smt.sampling_methods import LHS
ts = tx[0, :].unique().sort()[0].numpy()
xs = tx[1, :].unique().sort()[0].numpy()
xlimits = np.array([[ts[0], ts[-1]],[xs[0], xs[-1]]])
sampling = LHS(xlimits=xlimits)
inducing_points = torch.tensor(sampling(num_inducing)).unsqueeze(0)
else:
inducing_points = torch.stack([
tx[0, torch.randperm(tx.shape[1])[:num_inducing]],
tx[1, torch.randperm(tx.shape[1])[:num_inducing]]
], dim=1).unsqueeze(0)
gp_kwargs = dict(learn_inducing_locations=False,
natural=params['natural'],
use_tril=True)
gp_model = generate_multioutput_rbf_gp(
1, inducing_points,
initial_lengthscale=lengthscale,
ard_dims=2,
zero_mean=zero_mean,
gp_kwargs=gp_kwargs)
gp_model.covar_module.lengthscale = lengthscale
# lengthscale_constraint=Interval(0.1, 0.3),
gp_model.double()
print(inducing_points.shape)
plt.figure(figsize=(2, 2))
plt.scatter(inducing_points[0,:,0], inducing_points[0, :, 1], s=1)
# -
# Set up PDE (fenics module)
# + pycharm={"name": "#%%\n"}
# Define fenics model
ts = tx[0, :].unique().sort()[0].numpy()
t_range = (ts[0], ts[-1])
print(t_range)
time_steps = dataset.num_discretised
fenics_model = ReactionDiffusion(t_range, time_steps, mesh)
config = VariationalConfiguration(
initial_conditions=False,
num_samples=5
)
# + pycharm={"name": "#%%\n"}
# Define LFM
parameter_grad = params['parameter_grad'] if 'parameter_grad' in params else True
sensitivity = Parameter(
inv_softplus(torch.tensor(params['sensitivity'])) * torch.ones((1, 1), dtype=torch.float64),
requires_grad=False)
decay = Parameter(
inv_softplus(torch.tensor(params['decay'])) * torch.ones((1, 1), dtype=torch.float64),
requires_grad=parameter_grad)
diffusion = Parameter(
inv_softplus(torch.tensor(params['diffusion'])) * torch.ones((1, 1), dtype=torch.float64),
requires_grad=parameter_grad)
fenics_params = [sensitivity, decay, diffusion]
train_ratio = 0.3
num_training = int(train_ratio * tx.shape[1])
lfm = PartialLFM(1, gp_model, fenics_model, fenics_params, config, num_training_points=num_training)
# + pycharm={"name": "#%%\n"}
if params['natural']:
variational_optimizer = NGD(lfm.variational_parameters(), num_data=num_training, lr=0.01)
parameter_optimizer = Adam(lfm.nonvariational_parameters(), lr=0.01)
optimizers = [variational_optimizer, parameter_optimizer]
else:
optimizers = [Adam(lfm.parameters(), lr=0.005)]
track_parameters = list(lfm.fenics_named_parameters.keys()) +\
list(map(lambda s: f'gp_model.{s}', dict(lfm.gp_model.named_hyperparameters()).keys()))
# As in Lopez-Lopera et al., we take 30% of data for training
train_mask = torch.zeros_like(tx[0, :])
train_mask[torch.randperm(tx.shape[1])[:int(train_ratio * tx.shape[1])]] = 1
orig_data = dataset.orig_data.squeeze().t()
trainer = PDETrainer(lfm, optimizers, dataset,
clamp=params['clamp'],
track_parameters=track_parameters,
train_mask=train_mask.bool(),
warm_variational=-1,
lf_target=orig_data)
tx = trainer.tx
num_t_orig = orig_data[:, 0].unique().shape[0]
num_x_orig = orig_data[:, 1].unique().shape[0]
num_t = tx[0, :].unique().shape[0]
num_x = tx[1, :].unique().shape[0]
ts = tx[0, :].unique().sort()[0].numpy()
xs = tx[1, :].unique().sort()[0].numpy()
extent = [ts[0], ts[-1], xs[0], xs[-1]]
# -
# Now let's see some samples from the GP and corresponding LFM output
#
# + pycharm={"name": "#%%\n"}
time = orig_data[:, 0].unique()
latent = torch.tensor(orig_data[trainer.t_sorted, 2]).unsqueeze(0)
latent = latent.repeat(lfm.config.num_samples, 1, 1)
latent = latent.view(lfm.config.num_samples, 1, num_t_orig, num_x_orig)
time_interp = tx[0].unique()
time_interp[-1] -= 1e-5
latent = torch.from_numpy(interp1d(time, latent, axis=2)(time_interp))
# gp_model.covar_module.lengthscale = 0.3*0.3 * 2
out = gp_model(trainer.tx.transpose(0, 1))
sample = out.sample(torch.Size([lfm.config.num_samples])).permute(0, 2, 1)
plot_spatiotemporal_data(
[
sample.mean(0)[0].detach().view(num_t, num_x).t(),
latent[0].squeeze().view(num_t, num_x).t(),
],
extent,
titles=['Prediction', 'Ground truth']
)
sample = sample.view(lfm.config.num_samples, 1, num_t, num_x)
output_pred = lfm.solve_pde(sample).mean(0)
output = lfm.solve_pde(latent).mean(0)
print(output.shape)
plot_spatiotemporal_data(
[
output_pred.squeeze().detach().t(),
output.squeeze().detach().t(),
trainer.y_target.view(num_t_orig, num_x_orig).t()
],
extent,
titles=['Prediction', 'Prediction with real LF', 'Ground truth']
)
# + pycharm={"name": "#%%\n"}
print(sensitivity.shape)
num_t = tx[0, :].unique().shape[0]
num_x = tx[1, :].unique().shape[0]
y_target = trainer.y_target[0]
y_matrix = y_target.view(num_t_orig, num_x_orig)
pde_func, pde_target = lfm.fenics_model.interpolated_gradient(tx, y_matrix, disc=disc, plot=True)
u = orig_data[trainer.t_sorted, 2].view(num_t_orig, num_x_orig)
u = u.view(1, -1)
print(u.shape)
plot_spatiotemporal_data([pde_target.view(num_t, num_x).t()],
extent=extent, figsize=(3,3))
# + pycharm={"name": "#%%\n"}
train_ratio = 0.3
num_training = int(train_ratio * tx.shape[1])
print(num_training)
if params['natural']:
variational_optimizer = NGD(lfm.variational_parameters(), num_data=num_training, lr=0.04)
parameter_optimizer = Adam(lfm.nonvariational_parameters(), lr=0.03)
optimizers = [variational_optimizer, parameter_optimizer]
else:
optimizers = [Adam(lfm.parameters(), lr=0.09)]
pre_estimator = PartialPreEstimator(
lfm, optimizers, dataset, pde_func,
input_pair=(trainer.tx, trainer.y_target), target=pde_target,
train_mask=trainer.train_mask
)
# + pycharm={"name": "#%%\n"}
import time
t0 = time.time()
lfm.pretrain(True)
lfm.config.num_samples = 50
times = pre_estimator.train(60, report_interval=5)
lfm.config.num_samples = 5
# + pycharm={"name": "#%%\n"}
from alfi.utilities.torch import q2, cia
lfm.eval()
f = lfm(tx)
print(f.mean.shape)
f_mean = f.mean.detach()
f_var = f.variance.detach()
y_target = trainer.y_target[0]
print(f_mean.shape, y_target.shape, f_var.shape)
print('prot Q2', q2(y_target.squeeze(), f_mean.squeeze()))
print('prot CA', cia(y_target.squeeze(), f_mean.squeeze(), f_var.squeeze()))
gp = lfm.gp_model(tx.t())
lf_target = orig_data[trainer.t_sorted, 2]
f_mean = gp.mean.detach().view(num_t, num_x)[::disc].reshape(-1)
f_var = gp.variance.detach().view(num_t, num_x)[::disc].reshape(-1)
print('mrna Q2', q2(lf_target.squeeze(), f_mean.squeeze()))
print('mrna CA', cia(lf_target.squeeze(), f_mean.squeeze(), f_var.squeeze()))
print(np.stack(times).shape)
plt.plot(np.stack(times)[:, 1])
# + pycharm={"name": "#%%\n"}
#print(hihi)
trainer.train(5)
# + pycharm={"name": "#%%\n"}
with torch.no_grad():
lfm.config.num_samples = 5
lfm.eval()
u = lfm.gp_model(trainer.tx.t()).sample(torch.Size([5])).permute(0, 2, 1)
u = u.view(*u.shape[:2], num_t, num_x)
dy_t_ = pde_func(
trainer.y_target,
u[:, :, ::disc].contiguous(),
sensitivity,
decay,
diffusion)[0]
out_predicted = lfm.solve_pde(u.view(5, 1, num_t, num_x)).mean(0)
ts = tx[0, :].unique().numpy()
xs = tx[1, :].unique().numpy()
extent = [ts[0], ts[-1], xs[0], xs[-1]]
axes = plot_spatiotemporal_data(
[
trainer.y_target.view(num_t_orig, num_x_orig).t(),
pde_target.reshape(num_t_orig, num_x_orig).t(),
dy_t_.view(num_t_orig, num_x_orig).t(),
latent[0].view(num_t, num_x).t(),
u.mean(0).view(num_t, num_x).t(),
],
extent, titles=['y', 'target dy_t', 'pred dy_t_', 'target latent', 'pred latent']
)
plot_spatiotemporal_data(
[
# real.t().detach(),
trainer.y_target.view(num_t_orig, num_x_orig).t(),
out_predicted.t().detach()
],
extent, titles=['Target', 'Predicted']
)
print([softplus(param).item() for param in lfm.fenics_parameters])
# + pycharm={"name": "#%%\n"}
lfm = PartialLFM.load(filepath,
gp_cls=MultiOutputGP,
gp_args=[inducing_points, 1],
gp_kwargs=gp_kwargs,
lfm_args=[1, fenics_model, fenics_params, config])
# lfm = PartialLFM(gp_model, fenics_model, fenics_params, config)
gp_model = lfm.gp_model
optimizer = torch.optim.Adam(lfm.parameters(), lr=0.07)
trainer = PDETrainer(lfm, optimizer, dataset, track_parameters=list(lfm.fenics_named_parameters.keys()))
# + pycharm={"name": "#%%\n"}
from alfi.utilities.torch import smse, cia, q2
num_t = tx[0, :].unique().shape[0]
num_x = tx[1, :].unique().shape[0]
# f_mean = lfm(tx).mean.detach()
# f_var = lfm(tx).variance.detach()
y_target = trainer.y_target[0]
ts = tx[0, :].unique().sort()[0].numpy()
xs = tx[1, :].unique().sort()[0].numpy()
t_diff = ts[-1] - ts[0]
x_diff = xs[-1] - xs[0]
extent = [ts[0], ts[-1], xs[0], xs[-1]]
print(y_target.shape, f_mean.squeeze().shape)
f_mean_test = f_mean.squeeze()
f_var_test = f_var.squeeze()
print(q2(y_target, f_mean.squeeze()))
print(cia(y_target, f_mean_test, f_var_test).item())
print(smse(y_target, f_mean_test).mean().item())
# + pycharm={"name": "#%%\n"}
plotter = Plotter1d(lfm, np.arange(1))
titles = ['Sensitivity', 'Decay', 'Diffusion']
kinetics = list()
for key in lfm.fenics_named_parameters.keys():
kinetics.append(softplus(trainer.parameter_trace[key][-1]).squeeze().numpy())
kinetics = np.array(kinetics).reshape(3, 1)
plotter.plot_double_bar(kinetics, titles=titles)
# plotter.plot_latents()
# + pycharm={"name": "#%%\n"}
| notebooks/pde/pde.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # `Probability Distributions`
# %matplotlib inline
# for inline plots in jupyter
import matplotlib.pyplot as plt# import matplotlib
import seaborn as sns
import warnings
warnings.simplefilter("ignore")
from ipywidgets import interact
styles = ['seaborn-notebook', 'seaborn', 'seaborn-darkgrid', 'classic',
'_classic_test', 'seaborn-poster', 'tableau-colorblind10', 'grayscale',
'fivethirtyeight', 'seaborn-ticks', 'seaborn-dark',
'dark_background', 'seaborn-pastel',
'fast', 'Solarize_Light2', 'seaborn-colorblind', 'seaborn-white',
'seaborn-dark-palette',
'bmh', 'seaborn-talk', 'seaborn-paper', 'seaborn-deep', 'seaborn-bright',
'seaborn-muted',
'seaborn-whitegrid', 'ggplot']
# ## `3. Bernoulli Distribution`
# A random variable X is said to be a Bernoulli random variable with parameter p, shown as X∼Bernoulli(p), if its PMF is given by
#
# \begin{equation}
# \nonumber P_X(x) = \left\{
# \begin{array}{l l}
# p& \quad \text{for } x=1\\
# 1-p & \quad \text{ for } x=0\\
# 0 & \quad \text{ otherwise}
# \end{array} \right.
# \end{equation}
#
# where 0<p<1.
# %20color.png)
# import bernoulli
from scipy.stats import bernoulli
# * Bernoulli random variable can take either 0 or 1 using certain probability as a parameter. To generate 10000, bernoulli random numbers with success probability p =0.3, we will use bernoulli.rvs with two arguments.
# generate bernoulli
data_bern = bernoulli.rvs(size=10000,p=0.3)
ax= sns.distplot(data_bern,
kde=False,
hist_kws={"linewidth": 15,'alpha':1})
ax.set(xlabel='Bernoulli', ylabel='Frequency')
# We can see from the plot below out of 10000 trials with success probability 0.3,
# we get about 3000 successes
def BernoulliDistribution(palette="dark",kde = False,style = "ggplot"):
plt.figure(figsize=(13,10))
plt.style.use(style)
sns.set_palette(palette)
ax= sns.distplot(data_bern,
kde=kde,
hist_kws={"linewidth": 15,'alpha':1})
ax.set(xlabel='Bernoulli', ylabel='Frequency')
plt.show()
interact(BernoulliDistribution,palette = ["deep", "muted", "pastel", "bright",
"dark", "colorblind","Set3","Set2"],kde = [True,False],style = styles);
# ## `4. Binomial Distribution`
#
# 1. The binomial distribution is a discrete probability distribution that gives the probability that n identical independently distributed Bernoulli trials sum to k, where n is any natural number.
# 
from scipy.stats import binom
# * Binomial distribution is a discrete probability distributionlike Bernoulli. It can be used to obtain the number of successes from N Bernoulli trials. For example, to find the number of successes in 10 Bernoulli trials with p =0.5, we will use
binom.rvs(n=10,p=0.5)
# * We can also use binom.rvs to repeat the trials with size argument. If we want to repeat 5 times, we will use
data_binom = binom.rvs(n=10,p=0.5,size=10000)
ax = sns.distplot(data_binom,
kde=False,
hist_kws={"linewidth": 15,'alpha':1})
ax.set(xlabel='Binomial', ylabel='Frequency')
def BinomialDistribution(palette="dark",kde = False,style = "ggplot"):
plt.figure(figsize=(13,10))
plt.style.use(style)
sns.set_palette(palette)
ax = sns.distplot(data_binom,
kde=kde,
hist_kws={"linewidth": 15,'alpha':1})
ax.set(xlabel='Binomial', ylabel='Frequency')
plt.show()
interact(BinomialDistribution,palette = ["deep", "muted", "pastel", "bright",
"dark", "colorblind","Set3","Set2"],kde = [True,False],style = styles);
| DataScience365/Day-13/Probability Distributions part-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Classification
# In this notebook you will go through a classification problem.
#
# Unlike the previous lecture, where we dealt with a regression problem,
# this time you are going to predict a categorical value, i.e., a class.
#
# # Setup
#
# + jupyter={"outputs_hidden": false}
# To support both python 2 and python 3
from __future__ import division, print_function, unicode_literals
# Common imports
import numpy as np
import os
import pickle
# to make this notebook's output stable across runs
np.random.seed(42)
# To plot pretty figures
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# Where to save the figures
PROJECT_ROOT_DIR = "."
DATA_DIR = os.path.join(PROJECT_ROOT_DIR,'data')
IMG_DIR = os.path.join(PROJECT_ROOT_DIR, 'img')
CHAPTER_ID = "classification"
def save_fig(fig_id, tight_layout=True):
path = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID, fig_id + ".png")
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format='png', dpi=300)
# -
# ## MNIST
# MNIST is likely the most famous dataset in ML.
#
# It is a collection of 70.000 handwritten digits.
#
# Each image is associated with its actual digit.
#
# `sklearn` provides some helper function to fetch this dataset.
#
# **Warning**: `fetch_mldata()` is deprecated since Scikit-Learn 0.20. You should use `fetch_openml()` instead. However, it returns the unsorted MNIST dataset, whereas `fetch_mldata()` returned the dataset sorted by target (the training set and the test test were sorted separately). In general, this is fine, but if you want to get the exact same results as before, you need to sort the dataset using the following function:
# + jupyter={"outputs_hidden": false}
def sort_by_target(mnist):
reorder_train = np.array(sorted([(target, i) for i, target in enumerate(mnist.target[:60000])]))[:, 1]
reorder_test = np.array(sorted([(target, i) for i, target in enumerate(mnist.target[60000:])]))[:, 1]
mnist.data[:60000] = mnist.data.iloc[reorder_train]
mnist.target[:60000] = mnist.target.iloc[reorder_train]
mnist.data[60000:] = mnist.data.iloc[reorder_test + 60000]
mnist.target[60000:] = mnist.target.iloc[reorder_test + 60000]
# -
def maybe_download_mnist():
if not os.path.isdir(DATA_DIR):
os.mkdir(DATA_DIR)
download = False
if not os.listdir(DATA_DIR):
download = True
# empty directory download the file
try:
from sklearn.datasets import fetch_openml
mnist = fetch_openml('mnist_784', version=1, cache=True)
mnist.target = mnist.target.astype(np.int8) # fetch_openml() returns targets as strings
sort_by_target(mnist) # fetch_openml() returns an unsorted dataset
except ImportError:
from sklearn.datasets import fetch_mldata
mnist = fetch_mldata('MNIST original')
mode = 'wb' if download else 'rb'
with open(os.path.join(DATA_DIR,'mnist.pickle'), mode) as f:
if download:
pickle.dump(mnist, f)
else:
mnist = pickle.load(f)
return mnist
mnist = maybe_download_mnist()
# The object you just downloaded has a dict-like structure.
#
# The actual data are stored in the ``.data`` field of the object.
#
# + jupyter={"outputs_hidden": false}
print([k for k in mnist])
# -
# Take a look at the shape of your dataset.
#
# + jupyter={"outputs_hidden": false}
mnist.data.shape
# + jupyter={"outputs_hidden": false}
X, y = mnist["data"].values, mnist["target"].values
X.shape
# + jupyter={"outputs_hidden": false}
y.shape
# + jupyter={"outputs_hidden": false}
28*28
# -
# There are 70.000 images with 784 features.
# Each feature represents the density of a pixel, ranging from 0 to 255.
#
# Each row is the rollout vector of a 28x28 image.
#
# You can plot some data by reshaping the vector into the original matrix and then you
# can use ``matplotlib.imshow``.
# + jupyter={"outputs_hidden": false}
some_digit = X[36000]
def draw(sample, save_path = None):
some_digit_image = some_digit.reshape(28, 28)
plt.imshow(some_digit_image, cmap = mpl.cm.binary,
interpolation="nearest")
plt.axis("off")
if save_path is not None:
save_fig(os.path.join(IMG_DIR, save_path))
draw(some_digit)
# + jupyter={"outputs_hidden": false}
def plot_digits(instances, images_per_row=10, **options):
size = 28
images_per_row = min(len(instances), images_per_row)
images = [instance.reshape(size,size) for instance in instances]
n_rows = (len(instances) - 1) // images_per_row + 1
row_images = []
n_empty = n_rows * images_per_row - len(instances)
images.append(np.zeros((size, size * n_empty)))
for row in range(n_rows):
rimages = images[row * images_per_row : (row + 1) * images_per_row]
row_images.append(np.concatenate(rimages, axis=1))
image = np.concatenate(row_images, axis=0)
plt.imshow(image, cmap = mpl.cm.binary, **options)
plt.axis("off")
# + jupyter={"outputs_hidden": false}
plt.figure(figsize=(9,9))
example_images = np.r_[X[:12000:600], X[13000:30600:600], X[30600:60000:590]]
plot_digits(example_images, images_per_row=10)
plt.show()
# -
# Now let's split the data
# + jupyter={"outputs_hidden": false}
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size=1/7, random_state=42 )
X_train.shape
# -
# # Training a Binary Classifier
#
# Before diving into the details of the problem,
# let's focus on slightly simpler sub-problem.
#
# You have to train a _binary classifier_.
#
# The label _5__ is regarded as the positive class.
#
# Let's create the new target vector for this problem.
# + jupyter={"outputs_hidden": false}
y_train_5 = (y_train == 5)
y_test_5 = (y_test == 5)
some_digit = X[36000]
# -
# Let's pick the **Stochastic Gradient Descent** classifier and train it upon the
# new dataset.
#
# The main advantage of this classifier is its ability to handle large datasets in a very efficient way.
#
# + jupyter={"outputs_hidden": false}
from sklearn.linear_model import SGDClassifier
# train it yourself
sgd_clf = SGDClassifier(max_iter=5, tol=-np.infty, random_state=42)
sgd_clf.fit(X_train, y_train_5)
# + jupyter={"outputs_hidden": false}
draw(some_digit)
sgd_clf.predict([some_digit])
# -
# ## Performance Measures for Classification Problem.
#
# When we introduced the regression problem we discussed about two commonly
# used performance measures: RMSE, MAE.
#
# **Question**: Are they useful in this context?
#
# *Answer*:
#
# ----
#
# ## Measuring Accuracy with Cross-Validation
# Accuracy is defined as the ratio of
# the number of correct predictions on the total number of predictions.
#
# More formally:
#
# $$
# ACCURACY = \frac{\#\: right predictions}{\#\: predictions}
# $$
#
# ### Implementing Cross-Validation
# Occasionally you will need more control over the cross-validation process than what
# ``cross_val_score`` and similar functions provide.
#
# In these cases, you can implement
# cross-validation yourself; it is actually fairly straightforward.
#
# The ``StratifiedKFold`` class performs stratified sampling
# to produce folds that contain a representative ratio of each class.
#
# Every iteration of the following code involves the following steps:
#
# 1. Create a clone of the classifier
# 2. Train the classifier against the training folds
# 3. Compute predicitons over the validation fold
#
# + jupyter={"outputs_hidden": false}
from sklearn.model_selection import StratifiedKFold
from sklearn.base import clone
# stratified splitting of the training samples
skfolds = StratifiedKFold(n_splits=3)
for train_index, test_index in skfolds.split(X_train, y_train_5):
clone_clf = clone(sgd_clf)
X_train_folds = X_train[train_index]
y_train_folds = (y_train_5[train_index])
X_test_fold = X_train[test_index]
y_test_fold = (y_train_5[test_index])
clone_clf.fit(X_train_folds, y_train_folds)
y_pred = clone_clf.predict(X_test_fold)
n_correct = sum(y_pred == y_test_fold)
print(n_correct / len(y_pred))
# -
# If your are lazy, just go with sklearn:
# + jupyter={"outputs_hidden": false}
from sklearn.model_selection import cross_val_score
cross_val_scores = cross_val_score(sgd_clf, X_train, y_train_5, cv=3, scoring="accuracy")
cross_val_scores.max()
# -
# More than 90%! It is great, isn't it?
#
# **Question**: Should we be satisfied with this result?
#
# Let's implement another classifier. A dummy classifier that always predict
# ``False``.
# + jupyter={"outputs_hidden": false}
from sklearn.base import BaseEstimator
class Never5Classifier(BaseEstimator):
def fit(self, X, y=None):
pass
def predict(self, X):
return np.zeros((len(X), 1), dtype=bool)
# + jupyter={"outputs_hidden": false}
never_5_clf = Never5Classifier()
cross_val_score(never_5_clf, X_train, y_train_5, cv=3, scoring="accuracy")
# -
# **Question**: What's just happened?
#
# Accuracy is often a misleading metric,
# especially when data are skewed.
#
# ## Confusion Matrix
#
# A much better way to evaluate the performance of a classifier is to look at the confusion matrix.
#
# The general idea is to count the number of times instances of class A are
# classified as class B.
#
# The confusion matrix can be computed with function ``confusion_matrix`` of
# ``sklearn.metrics``.
#
# First you need to compute some predictions.
# + jupyter={"outputs_hidden": false}
from sklearn.model_selection import cross_val_predict
y_train_pred = cross_val_predict(sgd_clf, X_train, y_train_5, cv=3)
# + jupyter={"outputs_hidden": false}
def display_cofusion_matrix(matrix, classes):
s = "\t"+"\t".join(classes)
for i, row in enumerate(matrix):
s += "\n{}\t{}".format(classes[i], "\t".join([str(x) for x in row]))
return s
from sklearn.metrics import confusion_matrix
print(f"Ground Truth:{y_train_5.sum()}, Predicted: {y_train_pred.sum()}")
print(display_cofusion_matrix(confusion_matrix(y_train_5, y_train_pred), ['not-5', '5']))
# -
# Each row represents the actual class while the column represents the predicted class.
#
# Since we are dealing with binary classification, we can devise 4 categories:
# * True Positive - samples predicted as positive that are actually positive (right classification)
# * True Negative - samples predicted as negative that are actually negative (right classification)
# * False Positive - samples predicted as positive that are actually negative (wrong classification)
# * False Negative - samples predicted as negative that are actually positibe( wrong classification)
#
#
# + jupyter={"outputs_hidden": false}
from sklearn.metrics import plot_confusion_matrix
plot_confusion_matrix(sgd_clf, X_train, y_train_5)
# -
# A perfect binary classifier would be the following
# + jupyter={"outputs_hidden": false}
y_train_perfect_predictions = y_train_5
# + jupyter={"outputs_hidden": false}
confusion_matrix(y_train_5, y_train_perfect_predictions)
# -
# ## Precision and Recall
# The confusion matrix is the building block for two other measures:
#
# * Precision - it is given by:
# $$PRECISION = \frac{TP}{TP+FP}$$
#
# It measures how many times
# prediction of a positive sample were actually positive.
#
# * Recall - it is given by: $$ RECALL = \frac{TP}{TP+FN}$$
#
# It measures how many positve samples are captured from the set of all the positive samples.
#
# If we apply these measures over the trained classifier we get:
# + jupyter={"outputs_hidden": false}
from sklearn.metrics import precision_score, recall_score
precision_score(y_train_5, y_train_pred)
# + jupyter={"outputs_hidden": false}
recall_score(y_train_5, y_train_pred)
# -
# See? The model is far from being perfect as the >90% precision suggested before.
#
# Precision and recall, combined together, define the so called $F_1$ score denoted as:
#
# $$
# F_1 = \frac{2}{\frac{1}{precision} + \frac{1}{recall}} = 2 \times \frac{precision \times recall}{precision + recall}
# $$
# The $F_1$ score favors classifiers that have similar precision and recall.
#
# **Question**: Are precision and recall equally important?
#
# *Answer*....
#
# **Question**: Is there any correlation between precision and recall?
#
# *Answer*....
#
# + jupyter={"outputs_hidden": false}
from sklearn.metrics import f1_score
f1_score(y_train_5, y_train_pred)
# -
#
# ## Precision/Recall Trade-off
#
# ``SGDClassifier`` associates to each point a score,
# based on which it takes its final decision about the class of a sample.
#
# In order to take this decision, sgd uses a threshold value.
# If the score of a sample exceeds this threshold then it is considered
# a positive sample.
#
#
# **Question** How, and to what degree, does the threshold impact on both precision and recall?
#
# *Answer*
#
# ---
# Let's see what happen when you increase or decrease the threshold.
#
# With the method ``decision_function`` you can access the score assigned to
# the sample given as input.
# + jupyter={"outputs_hidden": false}
y_scores = sgd_clf.decision_function([some_digit])
y_scores
# + jupyter={"outputs_hidden": false}
threshold = 0
y_some_digit_pred = (y_scores > threshold)
# + jupyter={"outputs_hidden": false}
y_some_digit_pred
# -
# You can try different values of threshold
# + jupyter={"outputs_hidden": false}
threshold = -200000
y_some_digit_pred = (y_scores > threshold)
y_some_digit_pred
# -
# If you want to obtain scores instead of classes you can use set ``method="decision_function"``.
#
# + jupyter={"outputs_hidden": false}
y_scores = cross_val_predict(sgd_clf, X_train, y_train_5, cv=3,
method="decision_function")
y_scores.shape
# -
# Setting manually a threshold is not a common approach.
#
# A better way is to plot the precision recall curve.
#
# + jupyter={"outputs_hidden": false}
from sklearn.metrics import precision_recall_curve
precisions, recalls, thresholds = precision_recall_curve(y_train_5, y_scores)
print(precisions.shape, recalls.shape, thresholds.shape)
# + jupyter={"outputs_hidden": false}
def plot_precision_recall_vs_threshold(precisions, recalls, thresholds):
plt.plot(thresholds, precisions[:-1], "b--", label="Precision", linewidth=2)
plt.plot(thresholds, recalls[:-1], "g-", label="Recall", linewidth=2)
plt.xlabel("Threshold", fontsize=16)
plt.legend(loc="upper left", fontsize=16)
plt.ylim([0, 1])
plt.figure(figsize=(8, 4))
plot_precision_recall_vs_threshold(precisions, recalls, thresholds)
plt.xlim([thresholds.min(), thresholds.max()])
plt.show()
# -
thresholds.max()
# Based on this curve you can select the threshold that gives you the best
# precision/recall tradeoff.
#
# Another way to select a good precision/recall tradeoff is to plot
# precision directly against recall.
# + jupyter={"outputs_hidden": false}
y_train_pred_90 = (y_scores > 70000)
# + jupyter={"outputs_hidden": false}
precision_score(y_train_5, y_train_pred_90)
# + jupyter={"outputs_hidden": false}
recall_score(y_train_5, y_train_pred_90)
# +
def plot_precision_vs_recall(precisions, recalls):
plt.plot(recalls, precisions, "b-", linewidth=2)
plt.xlabel("Recall", fontsize=16)
plt.ylabel("Precision", fontsize=16)
plt.axis([0, 1, 0, 1])
plt.figure(figsize=(8, 6))
plot_precision_vs_recall(precisions, recalls)
plt.show()
# -
# __Take Home Lesson__
#
# At this point you should be aware of the fact that accuracy is not the only metrics.
#
# Clearly, having 90% precision is not useful if either recall or precision are too low.
#
# ## ROC curves
# The receiver operating characteristic curve is another common tool used
# with binary classifier.
#
# The ROC curve shows the true-positive-rate (another name for Recall) against the false-positive-rate.
#
# This second quantity, the FPR, is the ratio of negative samples misclassified as positive.
#
# Another quantity is the true-negative-rate, i.e., the ratio of negative samples correctly classified as negative (TNR)
#
# The false positive rate can be computed as $FPR = 1-FNR$
#
# The ROC curve combines all these information.
#
# In short:
#
# * True positive rate (recall/sensitivity)
# $$TPR = \frac{TP}{TP+FN}$$
#
# * False positive rate
# $$
# FPR = \frac{FP}{FP+TN}
# $$
# * True negative rate (specificity)
# $$
# TNR = \frac{TN}{TN+FP}
# $$
#
# To plot the ROC curve, you first need to compute the TPR and FPR for various threshold values, using the ``roc_curve`` function.
#
# The ROC curve plots *sensitivity* versus 1-*specificity*.
# + jupyter={"outputs_hidden": false}
from sklearn.metrics import roc_curve
fpr, tpr, thresholds = roc_curve(y_train_5, y_scores)
# + jupyter={"outputs_hidden": false}
def plot_roc_curve(fpr, tpr, label=None):
plt.plot(fpr, tpr, linewidth=2, label=label)
plt.plot([0, 1], [0, 1], 'k--')
plt.axis([0, 1, 0, 1])
plt.xlabel('False Positive Rate', fontsize=16)
plt.ylabel('True Positive Rate', fontsize=16)
plt.figure(figsize=(8, 6))
plot_roc_curve(fpr, tpr)
plt.show()
# -
# Again, the ROC curve highlights the negative correlation between precision and recall.
#
# In fact, increasing recall usually leads to decreasing accuracy.
#
# The dotted line represents the ROC curve associated to a purely random classifier.
#
# The performance of a classifier are proportional to the distance between its
# ROC curve and the one related to the random classifiers.
#
#
# One can directly measure the area under the ROC curve, that is called AUC.
#
# It must be as closer as possible to one.
#
# sklearn provides a function for computing this area.
#
# + jupyter={"outputs_hidden": false}
from sklearn.metrics import roc_auc_score
roc_auc_score(y_train_5, y_scores)
# -
# ### Review
#
# You have seen a number of different ways for measuring the performance of a classifier.
#
# > How to choose the correct one?
#
# There are general rule of thumbs.
#
# Precision Recall comes in handy when the dataset is unbalanced, with few positive samples
# as opposed to negative ones. Also, if you are more interested in keeping false positive low
# rather than false negative.
#
# In other situations, ROC/AUC is the way to go.
#
# This result suggests that your model works pretty good.
#
# However, there is still room for improvement.
# In fact, performance are good because the dataset is strongly unbalanced towards the negative class.
#
# **Exercise**
#
# Train a ``RandomForestClassifier`` (Hint: pass ``predict_proba`` as the ``method`` argument).
#
# Display the ROC curve, precision and recall.
# + jupyter={"outputs_hidden": false}
#your code here
# -
# # Multiclass classification
# From this point on, we will address the original multiclass classification problem.
#
#
# Instead of having to choose between only a pair of labels, you need to assign a label amongst a number
# labels.
#
# Many algorithms like the Random Forest Classifier
# or the Naive Bayes Classifier are able to genuinely handle multiclass
# classification problems.
#
# However, other algorithm needs to be properly extended in order to
# embrace a multiclass classification problem.
#
# The main strategy is to adapt their binary version.
#
# Two approaches are:
#
# 1. One vs All (OVA) - if you are asked to predict a class amongst N different ones, you can train N different binary classifiers. Each one of them is trained in order to detect a single class against all the others.
#
# 2. One vs One (OVO) - given N different class, you can N x (N-1)/2 different binary classifier. You will need to train a classifier for every possible pair of classes.
#
# For most binary classification algorithms the OVA approach is the one adopted by default.
#
# sklearn automatically detects when you need to solve a multiclass classification problem.
#
# + jupyter={"outputs_hidden": false}
sgd_clf.fit(X_train, y_train)
sgd_clf.predict([some_digit])
# -
# Actually, under the hood,
# scikit-Learn actually trained 10 binary classifiers, it obtained their decision scores for the
# image, and then it selected the class with the highest score.
#
# To convince yourself about the actual strategy adopted by sklearn you can
# compute the scores associated with the ``some_digit``.
# + jupyter={"outputs_hidden": false}
some_digit_scores = sgd_clf.decision_function([some_digit])
some_digit_scores
# -
# Each score **reflects** a probability, i.e., the likelihood the a certain sample belongs to
# the corresponding class.
#
# The class is assigned computing the argmax on this vector of scores
# + jupyter={"outputs_hidden": false}
np.argmax(some_digit_scores)
draw(some_digit)
# -
# **Note**: Be careful, this 5 actually is the index of the maximum element, not the class!
#
# The classes upon which the classifier is trained can be accessed via the ``classes_`` field of the estimator
# + jupyter={"outputs_hidden": false}
sgd_clf.classes_
# + jupyter={"outputs_hidden": false}
sgd_clf.classes_[5]
# -
# You can force sklearn to adopt the OVO by wrapping a classifier with the
# ``OneVsOneClassifier`` .
# You need to pass a regular binary classifier and then call the usual fit method.
# + jupyter={"outputs_hidden": false}
from sklearn.multiclass import OneVsOneClassifier
ovo_clf = OneVsOneClassifier(SGDClassifier(max_iter=5, tol=-np.infty, random_state=42))
ovo_clf.fit(X_train, y_train)
ovo_clf.predict([some_digit])
# + jupyter={"outputs_hidden": false}
len(ovo_clf.estimators_)
# -
# Now, try with a **RandomForestClassifier**
# + jupyter={"outputs_hidden": false}
forest_clf.fit(X_train, y_train)
forest_clf.predict([some_digit])
# + jupyter={"outputs_hidden": false}
forest_clf.predict_proba([some_digit])
# + jupyter={"outputs_hidden": false}
cross_val_score(sgd_clf, X_train, y_train, cv=3, scoring="accuracy")
# -
# It gets over 84% on all test folds.
#
# A random classifier would get 10% accuracy, not a bad score.
# But you can improve.
#
# We did not use any preprocessing techniques yet.
#
# + jupyter={"outputs_hidden": false}
from sklearn.preprocessing import StandardScaler, MinMaxScaler
from sklearn.pipeline import Pipeline
#your code here
p = Pipeline([
('scaler', MinMaxScaler()),
('classifier', sgd_clf)
])
cross_val_score(p, X_train, y_train, cv=3, scoring="accuracy")
# -
# **Exercise**: Modify the above cell to train the same classifier but after applying the `StandardScaler`
#
#
# ## Error Analysis
# Let assume we found a promising model.
#
# One thing to do is to evaluate its errors.
#
# You need to make predictions using the ``cross_val_predict`` fnction and then
# call the ``confusion_matrix()`` function, just like
# you did earlier:
# + jupyter={"outputs_hidden": false}
X_train_scaled = StandardScaler().fit_transform(X_train)
y_train_pred = cross_val_predict(sgd_clf, X_train_scaled, y_train, cv=3)
conf_mx = confusion_matrix(y_train, y_train_pred)
conf_mx
# + jupyter={"outputs_hidden": false}
import seaborn as sns
def plot_confusion_matrix(matrix):
fig = plt.figure(figsize=(8,8))
ax = fig.add_subplot(111)
sns.heatmap(matrix,
annot=True,
cmap="YlGnBu")
plot_confusion_matrix(conf_mx)
# -
# Let’s focus the plot on the errors.
#
# First, you need to divide each value in the confusion
# matrix by the number of images in the corresponding class, so you can compare error
# rates instead of absolute number of errors.
#
# + jupyter={"outputs_hidden": false}
row_sums = conf_mx.sum(axis=1, keepdims=True) # num of samples for each class
col_sum = conf_mx.sum(axis=0, keepdims=True) # num of predictions for each class
norm_conf_mx = conf_mx / row_sums
# -
# Now let’s fill the diagonal with zeros to keep only the errors, and plot the result:
# + jupyter={"outputs_hidden": false}
np.fill_diagonal(norm_conf_mx, 0)
plot_confusion_matrix(norm_conf_mx)
# -
# The columns corresponding to 8 and 9 are quite dark,
# which tells you that many images get misclassified as 8s or 9s.
#
# Similarly, the rows for classes 8 and 9 are also quite dark, telling you that 8s
# and 9s are often confused with other digits.
#
# Conversely, some rows are pretty bright,
# such as ow 1: this means that most 1s are classified correctly.
# # Multilabel classification
#
# Until now each instance has always been assigned to just one class. In some cases you
# may want your classifier to output multiple classes for each instance. For example,
# consider a face-recognition classifier: what should it do if it recognizes several people
# on the same picture?
#
# Such a classification system that outputs multiple
# binary labels is called a multilabel classification system.
# + jupyter={"outputs_hidden": false}
from sklearn.neighbors import KNeighborsClassifier
y_train_large = (y_train >= 7)
y_train_odd = (y_train % 2 == 1)
y_multilabel = np.c_[y_train_large, y_train_odd]
knn_clf = KNeighborsClassifier()
knn_clf.fit(X_train, y_multilabel)
# -
# This code creates a y_multilabel array containing two target labels for each digit
# image: the first indicates whether or not
# the digit is large (7, 8, or 9) and the second
# indicates whether or not it is odd. The next lines create a KNeighborsClassifier,
# Now you can make a prediction, and notice
# that it outputs two labels:
# + jupyter={"outputs_hidden": false}
knn_clf.predict([some_digit])
# -
# And it gets it right! The digit 5 is indeed not large ( False ) and odd ( True ).
#
# There are many ways to evaluate a multilabel classifier, and selecting the right metric
# really depends on your project. For example, one approach is to measure the F 1 score
# for each individual label (or any other binary classifier metric discussed earlier), then
# simply compute the average score. This code computes the average F 1 score across all
# labels:
# + jupyter={"outputs_hidden": false}
y_train_knn_pred = cross_val_predict(knn_clf, X_train, y_multilabel, cv=3, n_jobs=-1)
f1_score(y_multilabel, y_train_knn_pred, average="macro")
# -
# This assumes that all labels are equally important, which may not be the case.
#
# One simple option is to give each label a weight equal to its support (i.e., the number of instances with that
# target label). To do this, simply set average="weighted" in the preceding code.
# # Exercises
# Train different classifiers and test their performance on the test set
# ## KNN classifier
# ## SVM
# # Decision Tree
# + jupyter={"outputs_hidden": false}
# -
# ## Logistic Regression
# ## 1. An MNIST Classifier With Over 97% Accuracy
# `KNeighborsClassifiers` seems to perform pretty well.
#
# Let's try to push its performance with
# automated parameter tuning.
#
# The algorithm has just one parameter: k, which represents the n
# ## 2. Testing and Selecting the best Model
# Now, you are required to build an entire pipeline.
# You need to select a number of models, tuning upon each of them and then you need to report in a table the following
# information:
#
# 1. Name of The estimator
# 2. Best Configuration
# 3. accuracy
# 3. precision
# 4. recall
# 5. AUC score
#
# Of course, these results must be obtained wrt the **VALIDATION SET**.
| 03-classification/classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/hila-chefer/NLP_Final_Project/blob/main/Explainability_Based_Attention_Head_Analysis_for_Transformers.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="HiplefhuIUDd"
# # **Explainability-Based Attention Head Analysis for Transformers**
# + colab={"base_uri": "https://localhost:8080/"} id="3ogYpvQAAH4s" outputId="679a9655-fdc6-43a1-ca45-08dc7e64927c"
# !git clone https://github.com/hila-chefer/NLP_Final_Project
import os
os.chdir(f'./NLP_Final_Project')
# !pip install -r requirements.txt
# + [markdown] id="P8sl0DTeHuKx"
# # **LXMERT**
# + [markdown] id="lMKASCZ2HzQh"
# **Examples from paper**
# + id="or8UETbZAYY3"
from lxmert.lxmert.src.modeling_frcnn import GeneralizedRCNN
import lxmert.lxmert.src.vqa_utils as utils
from lxmert.lxmert.src.processing_image import Preprocess
from transformers import LxmertTokenizer
from lxmert.lxmert.src.huggingface_lxmert import LxmertForQuestionAnswering
from lxmert.lxmert.src.lxmert_lrp import LxmertForQuestionAnswering as LxmertForQuestionAnsweringLRP
from lxmert.lxmert.src.VisualizationsGenerator import GeneratorOurs as VisGen
from lxmert.lxmert.src.ExplanationGenerator import HeadPrune, LayerPrune
import random
import torch
import cv2
# + id="fWKGyu2YAeSV"
OBJ_URL = "https://raw.githubusercontent.com/airsplay/py-bottom-up-attention/master/demo/data/genome/1600-400-20/objects_vocab.txt"
ATTR_URL = "https://raw.githubusercontent.com/airsplay/py-bottom-up-attention/master/demo/data/genome/1600-400-20/attributes_vocab.txt"
VQA_URL = "https://raw.githubusercontent.com/airsplay/lxmert/master/data/vqa/trainval_label2ans.json"
# + id="pns9sG9eAhho"
class ModelUsage:
def __init__(self):
self.vqa_answers = utils.get_data(VQA_URL)
# load models and model components
self.frcnn_cfg = utils.Config.from_pretrained("unc-nlp/frcnn-vg-finetuned")
self.frcnn_cfg.MODEL.DEVICE = "cuda"
self.frcnn = GeneralizedRCNN.from_pretrained("unc-nlp/frcnn-vg-finetuned", config=self.frcnn_cfg)
self.image_preprocess = Preprocess(self.frcnn_cfg)
self.lxmert_tokenizer = LxmertTokenizer.from_pretrained("unc-nlp/lxmert-base-uncased")
self.lxmert_vqa = LxmertForQuestionAnsweringLRP.from_pretrained("unc-nlp/lxmert-vqa-uncased").to("cuda")
self.lxmert_vqa_no_lrp = LxmertForQuestionAnswering.from_pretrained("unc-nlp/lxmert-vqa-uncased").to("cuda")
self.lxmert_vqa.eval()
self.lxmert_vqa_no_lrp.eval()
self.model = self.lxmert_vqa
def forward(self, item):
URL, question = item
self.image_file_path = URL
# run frcnn
images, sizes, scales_yx = self.image_preprocess(URL)
output_dict = self.frcnn(
images,
sizes,
scales_yx=scales_yx,
padding="max_detections",
max_detections= self.frcnn_cfg.max_detections,
return_tensors="pt"
)
inputs = self.lxmert_tokenizer(
question,
truncation=True,
return_token_type_ids=True,
return_attention_mask=True,
add_special_tokens=True,
return_tensors="pt"
)
self.question_tokens = self.lxmert_tokenizer.convert_ids_to_tokens(inputs.input_ids.flatten())
self.text_len = len(self.question_tokens)
# Very important that the boxes are normalized
normalized_boxes = output_dict.get("normalized_boxes")
features = output_dict.get("roi_features")
self.image_boxes_len = features.shape[1]
self.bboxes = output_dict.get("boxes")
self.output = self.lxmert_vqa(
input_ids=inputs.input_ids.to("cuda"),
attention_mask=inputs.attention_mask.to("cuda"),
visual_feats=features.to("cuda"),
visual_pos=normalized_boxes.to("cuda"),
token_type_ids=inputs.token_type_ids.to("cuda"),
return_dict=True,
output_attentions=False,
)
return self.output
def forward_prune(self, item, text_prune, image_prune):
URL, question = item
self.image_file_path = URL
# run frcnn
images, sizes, scales_yx = self.image_preprocess(URL)
output_dict = self.frcnn(
images,
sizes,
scales_yx=scales_yx,
padding="max_detections",
max_detections= self.frcnn_cfg.max_detections,
return_tensors="pt"
)
inputs = self.lxmert_tokenizer(
question,
truncation=True,
return_token_type_ids=True,
return_attention_mask=True,
add_special_tokens=True,
return_tensors="pt"
)
# Very important that the boxes are normalized
normalized_boxes = output_dict.get("normalized_boxes")
features = output_dict.get("roi_features")
output = self.lxmert_vqa_no_lrp(
input_ids=inputs.input_ids.to("cuda"),
attention_mask=inputs.attention_mask.to("cuda"),
visual_feats=features.to("cuda"),
visual_pos=normalized_boxes.to("cuda"),
token_type_ids=inputs.token_type_ids.to("cuda"),
return_dict=True,
output_attentions=False,
text_head_prune=text_prune,
image_head_prune=image_prune,
)
answer = self.vqa_answers[output.question_answering_score.argmax()]
return answer
# + id="4FqWPLMCApCo"
def save_image_vis(image_file_path, bbox_scores, pert_step=0):
img = cv2.imread(image_file_path)
mask = torch.zeros(img.shape[0], img.shape[1])
for index in range(len(bbox_scores)):
[x, y, w, h] = model.bboxes[0][index]
curr_score_tensor = mask[int(y):int(h), int(x):int(w)]
new_score_tensor = torch.ones_like(curr_score_tensor)*bbox_scores[index].item()
mask[int(y):int(h), int(x):int(w)] = torch.max(new_score_tensor,mask[int(y):int(h), int(x):int(w)])
if mask.max() > 0:
mask = (mask - mask.min()) / (mask.max() - mask.min())
mask = mask.unsqueeze_(-1)
mask = mask.expand(img.shape)
img = img * mask.cpu().data.numpy()
cv2.imwrite(
'experiments/lxmert/result_pert_{0}.jpg'.format(pert_step), img)
# + id="fHn4tvHKAsHt" colab={"base_uri": "https://localhost:8080/"} outputId="fb2204cc-386f-45c3-e3ab-ca6f56e86697"
model = ModelUsage()
vis_gen = VisGen(model)
head_prune = HeadPrune(model)
image_ids = [
# giraffe
'COCO_val2014_000000054123'
]
test_questions_for_images = [
################## paper samples
# zebra
"how many zebras are facing away from the camera?"
################## paper samples
]
# + id="CaxJXarlE5rm"
pert_steps = [0, 0.4, 0.6, 0.9]
# + id="mS3mophB_TeL"
def generate_perturbation_visualizations(URL, text):
scores_text, scores_image, _, _ = head_prune.generate_ours((URL, text))
num_text_layers = len(scores_text)
num_image_layers = len(scores_image)
num_text_heads = scores_text[0].shape[0]
num_image_heads = scores_image[0].shape[0]
tot_num = num_text_layers * num_text_heads + num_image_layers * num_image_heads
scores_text = torch.stack(scores_text)
scores_image = torch.stack(scores_image)
joint_scores = torch.cat([scores_text, scores_image]).to("cuda")
all_text_scores = []
for step_idx, step in enumerate(pert_steps):
# find top step heads
curr_num = int((1 - step) * tot_num)
joint_scores = joint_scores.flatten()
_, top_heads = joint_scores.topk(k=curr_num, dim=-1)
heads_indicator = torch.zeros_like(joint_scores)
heads_indicator[top_heads] = 1
heads_indicator = heads_indicator.reshape(num_text_layers + num_image_layers, num_image_heads)
heads_indicator_text = heads_indicator[:num_text_layers, :]
heads_indicator_image = heads_indicator[num_text_layers:, :]
R_t_t, R_t_i = vis_gen.generate_ours((URL, text), head_prune_text=heads_indicator_text, head_prune_image=heads_indicator_image)
model.forward_prune((URL, text), heads_indicator_text, heads_indicator_image)
image_scores = R_t_i[0]
text_scores = R_t_t[0, 1:-1]
save_image_vis(URL, image_scores, pert_step=step)
all_text_scores.append(text_scores)
return model.question_tokens, all_text_scores
# + id="C3nIj4sVEEEa"
import matplotlib.pyplot as plt
from PIL import Image
from captum.attr import visualization
import requests
# + id="QXoI89YgAyem" colab={"base_uri": "https://localhost:8080/", "height": 569} outputId="bd99482d-4f4f-47a7-c471-7d3100110b02"
URL = 'experiments/lxmert/{0}_orig.jpg'.format(image_ids[0])
question, all_text_scores = generate_perturbation_visualizations(URL, test_questions_for_images[0])
orig_image = Image.open(model.image_file_path)
fig, axs = plt.subplots(ncols=5, figsize=(20, 5))
axs[0].imshow(orig_image);
axs[0].axis('off');
axs[0].set_title('original');
for step_idx, step in enumerate(pert_steps):
masked_image = Image.open('experiments/lxmert/result_pert_{0}.jpg'.format(step))
axs[step_idx+1].imshow(masked_image);
axs[step_idx+1].axis('off');
axs[step_idx+1].set_title('{0} heads masked'.format(step));
text_scores = all_text_scores[step_idx]
text_scores = (text_scores - text_scores.min()) / (text_scores.max() - text_scores.min())
print("Text explainability after pruning {0}% of the heads:".format(step*100))
vis_data_records = [visualization.VisualizationDataRecord(text_scores,0,0,0,0,0,model.question_tokens[1:-1],1)]
visualization.visualize_text(vis_data_records)
# + [markdown] id="4YR9oY6oH5bS"
# # **Online Examples**
#
# To upload your own example, simply modify the URL to your image url, and the question to your question.
# + colab={"base_uri": "https://localhost:8080/", "height": 568} id="lgEMI4ZvHoJn" outputId="9cae8c09-21d4-4d4e-ac6d-64a9f8e0efbb"
URL = "https://vqa.cloudcv.org/media/val2014/COCO_val2014_000000549112.jpg"
question = 'where is the knife?'
# save image to experiments folder
im = Image.open(requests.get(URL, stream=True).raw)
im.save('experiments/lxmert/online_image.jpg', 'JPEG')
URL = 'experiments/lxmert/online_image.jpg'
question, all_text_scores = generate_perturbation_visualizations(URL, question)
orig_image = Image.open(model.image_file_path)
fig, axs = plt.subplots(ncols=5, figsize=(20, 5))
axs[0].imshow(orig_image);
axs[0].axis('off');
axs[0].set_title('original');
for step_idx, step in enumerate(pert_steps):
masked_image = Image.open('experiments/lxmert/result_pert_{0}.jpg'.format(step))
axs[step_idx+1].imshow(masked_image);
axs[step_idx+1].axis('off');
axs[step_idx+1].set_title('{0} heads masked'.format(step));
text_scores = all_text_scores[step_idx]
text_scores = (text_scores - text_scores.min()) / (text_scores.max() - text_scores.min())
print("Text explainability after pruning {0}% of the heads:".format(step*100))
vis_data_records = [visualization.VisualizationDataRecord(text_scores,0,0,0,0,0,model.question_tokens[1:-1],1)]
visualization.visualize_text(vis_data_records)
# + [markdown] id="3vVAdf7pJ7Uw"
# # **VisualBERT**
#
# Since MMF does not support single image loading, we present the result images for each pruning step as saved by running the VisualBERT visualizations script. To run this script and get the results locally, see our README: https://github.com/hila-chefer/NLP_Final_Project#34generating-visualizations
# + colab={"base_uri": "https://localhost:8080/", "height": 209} id="WTiwAL5YQqIw" outputId="2fa0e14b-9c83-48a4-a260-b79748466147"
URL = 'experiments/visualBERT/COCO_val2014_000000097230_orig.jpg'
question = 'how tall is the grass?'
# save image to experiments folder
orig_image = Image.open(URL)
fig, axs = plt.subplots(ncols=5, figsize=(20, 5))
axs[0].imshow(orig_image);
axs[0].axis('off');
axs[0].set_title('original');
print("QUESTION: ", question)
for step_idx, step in enumerate(pert_steps):
masked_image = Image.open('experiments/visualBERT/COCO_val2014_000000097230_{0}.jpg'.format(step*100))
axs[step_idx+1].imshow(masked_image);
axs[step_idx+1].axis('off');
axs[step_idx+1].set_title('{0} heads masked'.format(step));
| Explainability_Based_Attention_Head_Analysis_for_Transformers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Excercises Electric Machinery Fundamentals
# ## Chapter 6
# ## Problem 6-19
# + slideshow={"slide_type": "skip"}
# %pylab notebook
# -
# ### Description
# A dc test is performed on a 460-V $\Delta$-connected 100-hp induction motor. If $V_{DC} = 21\,V$ and $I_{DC} = 72\,A$,
#
# * What is the stator resistance $R_1$? Why is this so?
Vdc = 21.0 # [V]
Idc = 72.0 # [A]
# ### SOLUTION
# If this motor’s armature is connected in delta, then there will be two phases in parallel with one phase between the lines tested.
# <img src="figs/Problem_6-19.jpg" width="30%">
# Therefore, the stator resistance $R_1$ will be:
# $$\frac{V_{DC}}{I_{DC}} = \frac{R_1(R_1 + R_2)}{R_1 + (R_1 + R_2)} = \frac{2}{3}R_1$$
# $$R_1 = \frac{3}{2}\frac{V_{DC}}{I_{DC}}$$
R1 = 3/2 * Vdc/Idc
print('''
R1 = {:.3f} Ω
============'''.format(R1))
| Chapman/Ch6-Problem_6-19.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Tce3stUlHN0L"
# ##### Copyright 2020 The TensorFlow Authors.
# + cellView="form" id="tuOe1ymfHZPu"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="qFdPvlXBOdUN"
# # Keras 예제의 양자화 인식 훈련
# + [markdown] id="MfBg1C5NB3X0"
# <table class="tfo-notebook-buttons" align="left">
# <td><a target="_blank" href="https://www.tensorflow.org/model_optimization/guide/quantization/training_example"><img src="https://www.tensorflow.org/images/tf_logo_32px.png">TensorFlow.org에서 보기</a></td>
# <td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ko/model_optimization/guide/quantization/training_example.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png">Run in Google Colab</a></td>
# <td><a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ko/model_optimization/guide/quantization/training_example.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">GitHub에서 소스 보기</a></td>
# <td><a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ko/model_optimization/guide/quantization/training_example.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png">노트북 다운로드하기</a></td>
# </table>
# + [markdown] id="Bjmi3qZeu_xk"
# ## 개요
#
# *양자화 인식 훈련*의 엔드 투 엔드 예제를 시작합니다.
#
# ### 기타 페이지
#
# 양자화 인식 훈련이 무엇인지에 대한 소개와 이를 사용해야 하는지에 대한 결정(지원되는 내용 포함)은 [개요 페이지](https://www.tensorflow.org/model_optimization/guide/quantization/training.md)를 참조하세요.
#
# 사용 사례에 필요한 API를 빠르게 찾으려면(8bit로 모델을 완전히 양자화하는 것 이상), [종합 가이드](https://www.tensorflow.org/model_optimization/guide/quantization/training_comprehensive_guide.md)를 참조하세요.
#
# ### 요약
#
# 이 튜토리얼에서는 다음을 수행합니다.
#
# 1. MNIST용 `tf.keras` 모델을 처음부터 훈련합니다.
# 2. 양자화 인식 학습 API를 적용하여 모델을 미세 조정하고, 정확성을 확인하고, 양자화 인식 모델을 내보냅니다.
# 3. 모델을 사용하여 TFLite 백엔드에 대해 실제로 양자화된 모델을 만듭니다.
# 4. TFLite와 4배 더 작아진 모델에서 정확성의 지속성을 확인합니다. 모바일에서의 지연 시간 이점을 확인하려면, [TFLite 앱 리포지토리](https://www.tensorflow.org/lite/models)에서 TFLite 예제를 사용해 보세요.
# + [markdown] id="yEAZYXvZU_XG"
# ## 설정
# + id="zN4yVFK5-0Bf"
# ! pip uninstall -y tensorflow
# ! pip install -q tf-nightly
# ! pip install -q tensorflow-model-optimization
# + id="yJwIonXEVJo6"
import tempfile
import os
import tensorflow as tf
from tensorflow import keras
# + [markdown] id="psViY5PRDurp"
# ## 양자화 인식 훈련 없이 MNIST 모델 훈련하기
# + id="pbY-KGMPvbW9"
# Load MNIST dataset
mnist = keras.datasets.mnist
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
# Normalize the input image so that each pixel value is between 0 to 1.
train_images = train_images / 255.0
test_images = test_images / 255.0
# Define the model architecture.
model = keras.Sequential([
keras.layers.InputLayer(input_shape=(28, 28)),
keras.layers.Reshape(target_shape=(28, 28, 1)),
keras.layers.Conv2D(filters=12, kernel_size=(3, 3), activation='relu'),
keras.layers.MaxPooling2D(pool_size=(2, 2)),
keras.layers.Flatten(),
keras.layers.Dense(10)
])
# Train the digit classification model
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
model.fit(
train_images,
train_labels,
epochs=1,
validation_split=0.1,
)
# + [markdown] id="K8747K9OE72P"
# ## 양자화 인식 훈련으로 사전 훈련된 모델 복제 및 미세 조정
#
# + [markdown] id="F19k7ExXF_h2"
# ### 모델 정의하기
# + [markdown] id="JsZROpNYMWQ0"
# 전체 모델에 양자화 인식 훈련을 적용하고 모델 요약에서 이를 확인합니다. 이제 모든 레이어 앞에 "quant"가 붙습니다.
#
# 결과 모델은 양자화를 인식하지만, 양자화되지는 않습니다(예: 가중치가 int8 대신 float32임). 다음 섹션에서는 양자화 인식 모델에서 양자화된 모델을 만드는 방법을 보여줍니다.
#
# [종합 가이드](https://www.tensorflow.org/model_optimization/guide/quantization/training_comprehensive_guide.md)에서 모델 정확성의 향상을 위해 일부 레이어를 양자화하는 방법을 볼 수 있습니다.
# + id="oq6blGjgFDCW"
import tensorflow_model_optimization as tfmot
quantize_model = tfmot.quantization.keras.quantize_model
# q_aware stands for for quantization aware.
q_aware_model = quantize_model(model)
# `quantize_model` requires a recompile.
q_aware_model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
q_aware_model.summary()
# + [markdown] id="uDr2ijwpGCI-"
# ### 기준선과 비교하여 모델 훈련 및 평가하기
# + [markdown] id="XUBEn94hXYB1"
# 하나의 epoch 동안 모델을 훈련한 후 미세 조정을 시연하려면 훈련 데이터의 하위 집합에 대한 양자화 인식 훈련으로 미세 조정합니다.
# + id="_PHDGJryE31X"
train_images_subset = train_images[0:1000] # out of 60000
train_labels_subset = train_labels[0:1000]
q_aware_model.fit(train_images_subset, train_labels_subset,
batch_size=500, epochs=1, validation_split=0.1)
# + [markdown] id="-byC2lYlMkfN"
# 이 예제의 경우, 기준선과 비교하여 양자화 인식 훈련 후 테스트 정확성의 손실이 거의 없습니다.
# + id="6bMFTKSSHyyZ"
_, baseline_model_accuracy = model.evaluate(
test_images, test_labels, verbose=0)
_, q_aware_model_accuracy = q_aware_model.evaluate(
test_images, test_labels, verbose=0)
print('Baseline test accuracy:', baseline_model_accuracy)
print('Quant test accuracy:', q_aware_model_accuracy)
# + [markdown] id="2IepmUPSITn6"
# ## TFLite 백엔드를 위한 양자화 모델 생성하기
# + [markdown] id="1FgNP4rbOLH8"
# 다음을 통해 int8 가중치 및 uint8 활성화를 사용하여 실제로 양자화된 모델을 얻게 됩니다.
# + id="w7fztWsAOHTz"
converter = tf.lite.TFLiteConverter.from_keras_model(q_aware_model)
converter.optimizations = [tf.lite.Optimize.DEFAULT]
quantized_tflite_model = converter.convert()
# + [markdown] id="BEYsyYVqNgeY"
# ## TF에서 TFLite까지 정확성의 지속성 확인하기
# + [markdown] id="saadXD4JQsBK"
# 테스트 데이터세트에 대해 TF Lite 모델을 평가하는 도우미 함수를 정의합니다.
# + id="b8yBouuGNqls"
import numpy as np
def evaluate_model(interpreter):
input_index = interpreter.get_input_details()[0]["index"]
output_index = interpreter.get_output_details()[0]["index"]
# Run predictions on every image in the "test" dataset.
prediction_digits = []
for i, test_image in enumerate(test_images):
if i % 1000 == 0:
print('Evaluated on {n} results so far.'.format(n=i))
# Pre-processing: add batch dimension and convert to float32 to match with
# the model's input data format.
test_image = np.expand_dims(test_image, axis=0).astype(np.float32)
interpreter.set_tensor(input_index, test_image)
# Run inference.
interpreter.invoke()
# Post-processing: remove batch dimension and find the digit with highest
# probability.
output = interpreter.tensor(output_index)
digit = np.argmax(output()[0])
prediction_digits.append(digit)
print('\n')
# Compare prediction results with ground truth labels to calculate accuracy.
prediction_digits = np.array(prediction_digits)
accuracy = (prediction_digits == test_labels).mean()
return accuracy
# + [markdown] id="TuEFS4CIQvUw"
# 양자화 모델을 평가하고 TensorFlow의 정확성이 TFLite 백엔드까지 유지되는지 확인합니다.
# + id="VqQTyqz4NsWd"
interpreter = tf.lite.Interpreter(model_content=quantized_tflite_model)
interpreter.allocate_tensors()
test_accuracy = evaluate_model(interpreter)
print('Quant TFLite test_accuracy:', test_accuracy)
print('Quant TF test accuracy:', q_aware_model_accuracy)
# + [markdown] id="z8D7WnFF5DZR"
# ## 양자화로 4배 더 작아진 모델 확인하기
# + [markdown] id="I1c2IecBRCdQ"
# float TFLite 모델을 생성한 다음 TFLite 양자화 모델이 4배 더 작아진 것을 확인합니다.
# + id="jy_Lgfh8VkyX"
# Create float TFLite model.
float_converter = tf.lite.TFLiteConverter.from_keras_model(model)
float_tflite_model = float_converter.convert()
# Measure sizes of models.
_, float_file = tempfile.mkstemp('.tflite')
_, quant_file = tempfile.mkstemp('.tflite')
with open(quant_file, 'wb') as f:
f.write(quantized_tflite_model)
with open(float_file, 'wb') as f:
f.write(float_tflite_model)
print("Float model in Mb:", os.path.getsize(float_file) / float(2**20))
print("Quantized model in Mb:", os.path.getsize(quant_file) / float(2**20))
# + [markdown] id="0O5xuci-SonI"
# ## 결론
# + [markdown] id="O2I7xmyMW5QY"
# 이 튜토리얼에서는 TensorFlow Model Optimization Toolkit API를 사용하여 양자화 인식 모델을 만든 다음 TFLite 백엔드용 양자화 모델을 만드는 방법을 살펴보았습니다.
#
# 정확성 차이를 최소화하면서 MNIST 모델의 크기를 4배 압축하는 이점을 확인했습니다. 모바일에서의 지연 시간 이점을 확인하려면, [TFLite 앱 리포지토리](https://www.tensorflow.org/lite/models)에서 TFLite 예제를 사용해 보세요.
#
# 이 새로운 기능은 리소스가 제한된 환경에서 배포할 때 특히 중요하므로 사용해 볼 것을 권장합니다.
#
| site/ko/model_optimization/guide/quantization/training_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# In this notebook, I added the random walk process to a randomly selected sources among the ICRF3 defing source list and computed the global spin of the CRF.
# +
import random
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
from astropy.table import Table, join
# My progs
from myprogs.vlbi.ts_func import get_ts
from tool_func import vsh_fit_for_pm, random_walk
from linear_fit import linfit2d
np.random.seed(3)
# +
apm_all = Table.read("../data/ts_nju_pm_fit_10sigma-10step.dat", format="ascii.csv")
# convert mas/yr into muas/yr
apm_all["pmra"] = apm_all["pmra"] * 1e3
apm_all["pmra_err"] = apm_all["pmra_err"] * 1e3
apm_all["pmdec"] = apm_all["pmdec"] * 1e3
apm_all["pmdec_err"] = apm_all["pmdec_err"] * 1e3
# -
icrf3_def = Table.read("../data/icrf3sx-def-sou.txt", format="ascii")
# +
mask = apm_all["num_cln"] >= 5
apm_all = apm_all[mask]
apm_def = join(icrf3_def, apm_all, keys="iers_name")
# +
sample_size = 100
sou_idx = random.choices(np.arange(len(apm_def)), k=sample_size)
# -
resample_wx = np.zeros(sample_size)
resample_wy = np.zeros(sample_size)
resample_wz = np.zeros(sample_size)
resample_w = np.zeros(sample_size)
resample_ra = np.zeros(sample_size)
resample_dec = np.zeros(sample_size)
for i, idx in enumerate(sou_idx):
# Get the name for the selected source
sou_name = apm_def["iers_name"][idx]
# Coordinate time series for the source
coordts = get_ts(
sou_name,
data_dir="/Users/Neo/Astronomy/data/vlbi/nju/series-10step",
calc_oft=True)
# Add random walk process to the coordinate time series for this source
dra_rw, ddec_rw = random_walk(coordts["jyear"], t_scale=5, sigma_var=3)
# Form the new coordinate offset series
# coordts["dra"] = coordts["dra"] + dra_rw
# coordts["ddec"] = coordts["ddec"] + ddec_rw
coordts["dra"] = dra_rw
coordts["ddec"] = ddec_rw
# Re-fit the apparent proper motion
res1 = linfit2d(coordts["jyear"] - np.median(coordts["jyear"]),
coordts["dra"],
coordts["ddec"],
x_err=coordts["ra_err"],
y_err=coordts["dec_err"],
xy_cor=coordts["ra_dec_corr"],
fit_type="sep")
# Update the APM data
apm_def["pmra"][idx] = res1["x1"] * 1e3
apm_def["pmra_err"][idx] = res1["x1_err"] * 1e3
apm_def["pmdec"][idx] = res1["y1"] * 1e3
apm_def["pmdec_err"][idx] = res1["y1_err"] * 1e3
apm_def["pmra_pmdec_cor"][idx] = res1["x1y1_cor"]
# VSH fitting
pmt, sig, output = vsh_fit_for_pm(apm_def)
# Record the results
resample_wx[i] = pmt[0]
resample_wy[i] = pmt[1]
resample_wz[i] = pmt[2]
resample_w[i] = pmt[3]
resample_ra[i] = output["R_ra"]
resample_dec[i] = output["R_dec"]
# +
bin_size = 0.05
bin_array = np.arange(-1.5, 0, bin_size)
fig, ax = plt.subplots()
ax.hist(resample_wx,
bins=bin_array,
color="grey",
label="All")
ax.set_xlabel("$\\omega_{\\rm x}$ ($\\mu$as$\,$yr$^{-1}$)", fontsize=15)
ax.set_ylabel("Nb sources in bins", fontsize=15)
plt.tight_layout()
# plt.savefig("../plots/spin-x-from-resampled-apm.eps")
# +
bin_size = 0.05
bin_array = np.arange(-.5, 1.0, bin_size)
fig, ax = plt.subplots()
ax.hist(resample_wy,
bins=bin_array,
color="grey",
label="All")
ax.set_xlabel("$\\omega_{\\rm y}$ ($\\mu$as$\,$yr$^{-1}$)", fontsize=15)
ax.set_ylabel("Nb sources in bins", fontsize=15)
plt.tight_layout()
# plt.savefig("../plots/spin-y-from-resampled-apm.eps")
# +
bin_size = 0.05
bin_array = np.arange(-.1, 1.6, bin_size)
fig, ax = plt.subplots()
ax.hist(resample_wz,
bins=bin_array,
color="grey",
label="All")
# ax.plot(bin_array, rvs_wz.pdf(bin_array)*sample_num*bin_size, "r--")
# ax.text(-2., 55, "$\mu={:+.2f}$".format(mu_wz), fontsize=15)
# ax.text(-2., 45, "$\sigma={:.2f}$".format(std_wz), fontsize=15)
ax.set_xlabel("$\\omega_{\\rm z}$ ($\\mu$as$\,$yr$^{-1}$)", fontsize=15)
ax.set_ylabel("Nb sources in bins", fontsize=15)
plt.tight_layout()
# plt.savefig("../plots/spin-z-from-resampled-apm.eps")
# +
bin_size = 5
bin_array = np.arange(0, 150, bin_size)
fig, ax = plt.subplots()
ax.hist(resample_w,
bins=bin_array,
color="grey",
label="All")
ax.set_xlabel("$\\omega$ ($\\mu$as$\,$yr$^{-1}$)", fontsize=15)
ax.set_ylabel("Nb sources in bins", fontsize=15)
plt.tight_layout()
# +
bin_size = 10
bin_array = np.arange(0, 361, bin_size)
fig, ax = plt.subplots()
ax.hist(resample_ra,
bins=bin_array,
color="grey",
label="All")
# ax.plot(bin_array, rvs_ra.pdf(bin_array)*sample_num*bin_size, "r--")
# ax.text(0, 75, "$\mu={:.0f}$".format(mu_ra), fontsize=15)
# ax.text(0, 65, "$\sigma={:.0f}$".format(std_ra), fontsize=15)
ax.set_xlabel("$\\alpha_{\\rm apex}$ (degree))", fontsize=15)
ax.set_ylabel("Nb sources in bins", fontsize=15)
plt.tight_layout()
# +
bin_size = 5
bin_array = np.arange(-90, 91, bin_size)
fig, ax = plt.subplots()
ax.hist(resample_dec,
bins=bin_array,
color="grey",
label="All")
# ax.plot(bin_array, rvs_dec.pdf(bin_array)*sample_num*bin_size, "r--")
# ax.text(-80, 85, "$\mu={:-.0f}$".format(mu_dec), fontsize=15)
# ax.text(-80, 75, "$\sigma={:.0f}$".format(std_dec), fontsize=15)
ax.set_xlabel("$\\delta_{\\rm apex}$ (degree))", fontsize=15)
ax.set_ylabel("Nb sources in bins", fontsize=15)
plt.tight_layout()
| progs/influence-of-random-walk-on-spin-estimate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Install latest version from GitHub
# !pip install -q -U git+https://github.com/jdvelasq/techminer
# # Citations by terms by year
# +
import matplotlib.pyplot as plt
import pandas as pd
from techminer import DataFrame, Plot, heatmap
#
# Data loading
#
df = DataFrame(
pd.read_json(
"https://raw.githubusercontent.com/jdvelasq/techminer/master/data/tutorial/"
+ "cleaned-data.json",
orient="records",
lines=True,
)
)
#
# Columns of the dataframe
#
df.columns
# -
# ## Document Type example
df.citations_by_term_per_year("Document Type").head(40)
df.citations_by_term_per_year("Document Type", as_matrix=True)
plt.figure(figsize=(7, 4))
Plot(df.citations_by_term_per_year("Document Type", as_matrix=True)).heatmap(cmap='Blues')
| sphinx/tutorial/22-citations-by-terms-by-year.ipynb |