code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
import torch
import random
from torch import nn
from torch.autograd import Variable
from torch.optim import Adam
import torch.nn.functional as F
import torchtext
from torchtext.datasets import Multi30k
from torchtext.data import Field, BucketIterator
from torch.nn.utils import clip_grad_norm
import spacy
# ## Convenience Functions
def sequence_to_text(sequence, field):
pad = field.vocab.stoi['<pad>']
return " ".join([field.vocab.itos[int(i)] for i in sequence])
# ## Load Multi30k English/German parallel corpus for NMT
# TorchText takes care of tokenization, padding, special character tokens and batching.
def load_dataset(batch_size, device=0):
spacy_de = spacy.load('de')
spacy_en = spacy.load('en')
def tokenize_de(text):
return [tok.text for tok in spacy_de.tokenizer(text)]
def tokenize_en(text):
return [tok.text for tok in spacy_en.tokenizer(text)]
DE = Field(tokenize=tokenize_de, init_token='<sos>', eos_token='<eos>')
EN = Field(tokenize=tokenize_en, init_token='<sos>', eos_token='<eos>')
train, val, test = Multi30k.splits(exts=('.de', '.en'), fields=(DE, EN))
DE.build_vocab(train.src)
EN.build_vocab(train.trg)
train_iter, val_iter, test_iter = BucketIterator.splits(
(train, val, test), batch_size=batch_size, device=device, repeat=False)
return train_iter, val_iter, test_iter, DE, EN
# ## Model Inputs
# Model inputs are (seq_len, batch_size) Tensors of word indices
train_iter, val_iter, test_iter, DE, EN = load_dataset(batch_size=5, device=-1)
example_batch = next(iter(train_iter))
example_batch.src, example_batch.trg
# We can recover the original text by looking up each index in the vocabularies we build with the `load_data` function.
print(sequence_to_text(example_batch.src[:, 0], DE))
print(sequence_to_text(example_batch.trg[:, 0], EN))
# ## Architecture
# NMT uses an encoder-decoder architecture to effectively translate source sequences and target sequences that are of different lengths
# 
# ## Encoder
# Encodes each word of the source sequence into a `hidden_dim` feature map. Sometimes called an `annotation`. Also returns the hidden state of the encoder bi-rnn.
class Encoder(nn.Module):
def __init__(self, source_vocab_size, embed_dim, hidden_dim,
n_layers, dropout):
super(Encoder, self).__init__()
self.hidden_dim = hidden_dim
self.embed = nn.Embedding(source_vocab_size, embed_dim, padding_idx=1)
self.gru = nn.GRU(embed_dim, hidden_dim, n_layers,
dropout=dropout, bidirectional=True)
def forward(self, source, hidden=None):
embedded = self.embed(source) # (batch_size, seq_len, embed_dim)
encoder_out, encoder_hidden = self.gru(
embedded, hidden) # (seq_len, batch, hidden_dim*2)
# sum bidirectional outputs, the other option is to retain concat features
encoder_out = (encoder_out[:, :, :self.hidden_dim] +
encoder_out[:, :, self.hidden_dim:])
return encoder_out, encoder_hidden
embed_dim = 256
hidden_dim = 512
n_layers = 2
dropout = 0.5
encoder = Encoder(source_vocab_size=len(DE.vocab), embed_dim=embed_dim,
hidden_dim=hidden_dim, n_layers=n_layers, dropout=dropout)
encoder_out, encoder_hidden = encoder(example_batch.src)
print('encoder output size: ', encoder_out.size()) # source, batch_size, hidden_dim
print('encoder hidden size: ', encoder_hidden.size()) # n_layers * num_directions, batch_size, hidden_dim
# ## Attention
# Currently the `encoder_output` is a length 14 sequence and the target is a length 13 sequence. We need to compress the information in the `encoder_output` into a `context_vector` which should have all the information the decoder needs to predict the next step of its output. We will use `Luong Attention` to create this context vector.
class LuongAttention(nn.Module):
"""
LuongAttention from Effective Approaches to Attention-based Neural Machine Translation
https://arxiv.org/pdf/1508.04025.pdf
"""
def __init__(self, dim):
super(LuongAttention, self).__init__()
self.W = nn.Linear(dim, dim, bias=False)
def score(self, decoder_hidden, encoder_out):
# linear transform encoder out (seq, batch, dim)
encoder_out = self.W(encoder_out)
# (batch, seq, dim) | (2, 15, 50)
encoder_out = encoder_out.permute(1, 0, 2)
# (2, 15, 50) @ (2, 50, 1)
return encoder_out @ decoder_hidden.permute(1, 2, 0)
def forward(self, decoder_hidden, encoder_out):
energies = self.score(decoder_hidden, encoder_out)
mask = F.softmax(energies, dim=1) # batch, seq, 1
context = encoder_out.permute(
1, 2, 0) @ mask # (2, 50, 15) @ (2, 15, 1)
context = context.permute(2, 0, 1) # (seq, batch, dim)
mask = mask.permute(2, 0, 1) # (seq2, batch, seq1)
return context, mask
# This will normally be part of the decoder as it takes the previous decoder hidden state as input, but just to show the inputs and outputs I will use it here.
# We will initialize the Decoder rnn's hidden state with the last hidden state from the encoder. Because the encoder is bi-directional we have to reshape it's hidden state in order to select the layer we want.
attention = LuongAttention(dim=hidden_dim)
context, mask = attention(encoder_hidden[-1:], encoder_out)
print(context.size()) # (1, batch, attention_dim) contect_vector
print(mask.size()) # the weights used to compute weighted sum over encoder out (1, batch, source_len)
# ## Decoder with attention
class Decoder(nn.Module):
def __init__(self, target_vocab_size, embed_dim, hidden_dim,
n_layers, dropout):
super(Decoder, self).__init__()
self.n_layers = n_layers
self.embed = nn.Embedding(target_vocab_size, embed_dim, padding_idx=1)
self.attention = LuongAttention(hidden_dim)
self.gru = nn.GRU(embed_dim + hidden_dim, hidden_dim, n_layers,
dropout=dropout)
self.out = nn.Linear(hidden_dim * 2, target_vocab_size)
def forward(self, output, encoder_out, decoder_hidden):
"""
decodes one output frame
"""
embedded = self.embed(output) # (1, batch, embed_dim)
context, mask = self.attention(decoder_hidden[:-1], encoder_out) # 1, 1, 50 (seq, batch, hidden_dim)
rnn_output, decoder_hidden = self.gru(torch.cat([embedded, context], dim=2),
decoder_hidden)
output = self.out(torch.cat([rnn_output, context], 2))
return output, decoder_hidden, mask
decoder = Decoder(target_vocab_size=len(EN.vocab), embed_dim=embed_dim,
hidden_dim=hidden_dim, n_layers=n_layers, dropout=dropout)
# To translate one word from German to English, the decoder needs:
# 1. `encoder_outputs`
# 2. `decoder_hidden` initially, the last n_layers of encoder_hidden then it's own returned hidden state.
# 3. `previous_output` feed a batch of start of string token (index 2) at the first step.
#
# The attention mask that the decoder returns is not used in training but can be used to visualize where the decoder is "looking" in the input sequence in order to generate its current output.
decoder_hidden = encoder_hidden[-decoder.n_layers:]
start_token = example_batch.trg[:1]
start_token
output, decoder_hidden, mask = decoder(start_token, encoder_out, decoder_hidden)
print('output size: ', output.size()) # (1, batch, target_vocab) # predicted probability distribution over all possible target words
print('decoder hidden size ', decoder_hidden.size())
print('attention mask size', mask.size())
# ## Decoding Helpers
# nmt models use teacher forcing during training and greedy decoding or beam search for inference. In order to accommodate these behaviors, I've made simple helper classes that get output from the decoder using each policy.
#
# The Teacher class sometimes feeds the previous target to the decoder rather than the model's previous prediction. this can help speed convergence but requires targets to be loaded to the helper at each step
class Teacher:
def __init__(self, teacher_forcing_ratio=0.5):
self.teacher_forcing_ratio = teacher_forcing_ratio
self.targets = None
self.maxlen = 0
def load_targets(self, targets):
self.targets = targets
self.maxlen = len(targets)
def generate(self, decoder, encoder_out, encoder_hidden):
outputs = []
masks = []
decoder_hidden = encoder_hidden[-decoder.n_layers:] # take what we need from encoder
output = self.targets[0].unsqueeze(0) # start token
for t in range(1, self.maxlen):
output, decoder_hidden, mask = decoder(output, encoder_out, decoder_hidden)
outputs.append(output)
masks.append(mask.data)
output = Variable(output.data.max(dim=2)[1])
# teacher forcing
is_teacher = random.random() < self.teacher_forcing_ratio
if is_teacher:
output = self.targets[t].unsqueeze(0)
return torch.cat(outputs), torch.cat(masks).permute(1, 2, 0) # batch, src, trg
decode_helper = Teacher()
decode_helper.load_targets(example_batch.trg)
outputs, masks = decode_helper.generate(decoder, encoder_out, encoder_hidden)
# ## Calc loss
# reshape outputs and targets, ignore sos token at start of target batch.
F.cross_entropy(outputs.view(-1, outputs.size(2)),
example_batch.trg[1:].view(-1), ignore_index=1)
# The greedy decoder simply chooses the highest scoring word as output.
# We cam use the `set_maxlen` method to generate sequences the same length as our targets to easily check perplexity and bleu score during evaluation steps.
class Greedy:
def __init__(self, maxlen=20, sos_index=2):
self.maxlen = maxlen
self.sos_index = sos_index
def set_maxlen(self, maxlen):
self.maxlen = maxlen
def generate(self, decoder, encoder_out, encoder_hidden):
seq, batch, _ = encoder_out.size()
outputs = []
masks = []
decoder_hidden = encoder_hidden[-decoder.n_layers:] # take what we need from encoder
output = Variable(torch.zeros(1, batch).long() + self.sos_index) # start token
for t in range(self.maxlen):
output, decoder_hidden, mask = decoder(output, encoder_out, decoder_hidden)
outputs.append(output)
masks.append(mask.data)
output = Variable(output.data.max(dim=2)[1])
return torch.cat(outputs), torch.cat(masks).permute(1, 2, 0) # batch, src, trg
decode_helper = Greedy()
decode_helper.set_maxlen(len(example_batch.trg[1:]))
outputs, masks = decode_helper.generate(decoder, encoder_out, encoder_hidden)
outputs.size()
F.cross_entropy(outputs.view(-1, outputs.size(2)),
example_batch.trg[1:].view(-1), ignore_index=1)
# ## seq2seq wrapper
class Seq2Seq(nn.Module):
def __init__(self, encoder, decoder):
super(Seq2Seq, self).__init__()
self.encoder = encoder
self.decoder = decoder
def forward(self, source, decoding_helper):
encoder_out, encoder_hidden = self.encoder(source)
outputs, masks = decoding_helper.generate(self.decoder, encoder_out, encoder_hidden)
return outputs, masks
seq2seq = Seq2Seq(encoder, decoder)
decoding_helper = Teacher(teacher_forcing_ratio=0.5)
# ## example iteration with wrapper
decoding_helper.load_targets(example_batch.trg)
outputs, masks = seq2seq(example_batch.src, decode_helper)
outputs.size(), masks.size()
F.cross_entropy(outputs.view(-1, outputs.size(2)),
example_batch.trg[1:].view(-1), ignore_index=1)
| nmt_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # operators
a=10
b=20
#addition
a+b
#subtraction
a-b
#division
a/b
# MODULUS- Divides left hand operand by right hand operand and returns remainder
#modulus
b%a
#multiplication
a*b
# +
#equals
c=a==b
print(c)
#notequals
print(a!=b)
# +
#greater than
print(a>b)
#less than
print(a<b)
# -
#exponent
a=10
print(a**2)
# # LOOPS
# ### For loops:-
# Executes a sequence of statements multiple times and abbreviates the code that manages the loop variable
#
# A for loop acts as an iterator in Python, it goes through items that are in a sequence or any other iterable item. Objects that we've learned about that we can iterate over include strings,lists,tuples, and even built in iterables for dictionaries, such as the keys or values.
# General syntax:-
#
# for item in object:
# code
#
# ## examples for for loop
# ### ex:1 Compute the square of each number in X and assign it to a list y print y and let be range 1-100.??
x=range(1,100)
y=[]
for i in x:
y.append(i**2)
print(y)
# ### ex:2 print all elements in the list
list=['praneeth','metuku','india','usa','texas','kingsville','starbucks','dominos','data science','colaberry']
for i in list:
print(i)
# ### ex3: for loops in dictionaries printing keys and values
# use .items() to iterate through the keys and values of a dictionary
dict={'key1':'value1','key2':'value2','key3':'values3'}
for k,v in dict.items():
print(k)
print(v)
# ## while loops:-
# Repeats a statement or group of statements while a given condition is TRUE. It tests the condition before executing the loop body
# + active=""
# The general syntax of a while loop is:
#
# while test:
# code statement
# else:
# final code statements
#
# -
# ## examples of while loops
# ### ex1: print x value 5 times and when its done print ALLDONE!
x=0
while x<=5:
print('x value is',x)
x=x+1
else:
print('ALLDONE!')
# # Decision making
#
# ### if-else conditions:-
#
# + active=""
# General syntax:-
#
# if case1:
# perform action1
# elif case2:
# perform action2
# else:
# perform action 3
# -
# ### examples of if-else conditions:-
# ### ex-1: Push even numbers in [0, 100] to the list variable even_numbers,print even_numbers?
# +
x=[]
for i in range(1,100):
if i%2==0:
x.append(i)
print(x)
# -
# # break, continue, pass in whileloops
# + active=""
# break: Breaks out of the current closest enclosing loop.
# continue: Goes to the top of the closest enclosing loop.
# pass: Does nothing at all
# -
#using break
x=0
while x<10:
print('x value is',x)
x=x+1
if x==7:
print('hey my lucky number')
break;
#using continue
x=0
while x<10:
print('x value is',x)
x=x+1
if x==7:
print('hey my lucky number')
else:
print('continue works')
# # Methods
# Methods are essentially functions built into objects
# General syntax:
#
# object.method(arg1,arg2,etc...)
#append is a method for list
list=[1,2,3,4]
list.append(6)
list
# # Functions(**)
# Function is a useful device that groups together a set of statements so they can be run more than once. They can also let us specify parameters that can serve as inputs to the functions
# Functions will be one of most basic levels of reusing code in Python, and it will also allow us to start thinking of program design
# + active=""
# General syntax:
#
# def name_of_function(arg1,arg2):
# '''
# This is where the function's Document String (doc-string) goes
# '''
# # Do stuff here
# #return desired result
# -
#addition of two numbers with functions
def addition(num1,num2):
return num1+num2
addition(3,4)
# +
#prime number detection with functions
def is_prime(n):
for i in range(2,n):
if n%i==0:
print('not prime')
break;
else:
print('prime')
break;
is_prime(31)
# + active=""
# EXAMPLE:
#
# Write a function, compute_sqrt(x), which takes a non-negative number as an input and returns a dictionary with key as number and square root of the number as its value.
#
# Invoke the function compute_sqrt(25), assign it to variable sqrt_25 and print it out.
#
# +
def compute_sqrt(x):
return{x:x**(1/2)}
compute_sqrt(25)
# -
# # lambda expressions(anonymus function)
# lambda expressions allow us to create "anonymous" functions. This basically means we can quickly make ad-hoc functions without needing to properly define a function using def
# lambda's body is a single expression, not a block of statements
#
#
#squre of a function with lambda expressions
square=lambda x:x**2
square(2)
# + active=""
# #insight of how lambda works
#
# def square(x): return x**2
#
# -
# # Scope of the variables
# L: Local — Names assigned in any way within a function (def or lambda)), and not declared global in that function.
#
# E: Enclosing function locals — Name in the local scope of any and all enclosing functions (def or lambda), from inner to outer.
#
# G: Global (module) — Names assigned at the top-level of a module file, or declared global in a def within the file.
#
# B: Built-in (Python) — Names preassigned in the built-in names module : open,range,SyntaxError,...
# # object oriented programming(python)
# In Python, everything is an object
# ### class
# The user defined objects are created using the class keyword. The class is a blueprint that defines a nature of a future object. From classes we can construct instances. An instance is a specific object created from a particular class.
# + active=""
# # Create a new object type called Sample
# class Sample(object):
# pass
#
# # Instance of Sample
# x = Sample()
#
# print type(x)
# -
# # Class Attributes
# An attribute is a characteristic of an object. A method is an operation we can perform with the object
# + active=""
# syntax for attribute:
#
# self.attribute = something
#
# There is a special method called:
#
# __init__()
# -
# ## example : creating a employee class with name and id
# +
class employee:
#class object
company='colaberry'
def __init__(self,name,id):
self.name=name
self.id=id
praneeth=employee('praneethmetuku',1)
# -
print(praneeth.name)
print(praneeth.id)
print(praneeth.company)
# + active=""
# Here employee is the class name
# name,id are the attributes of class
# __init__ is the special method
# praneeth is the instance of the class employee
#
# + active=""
# The special method
# __init__()
#
# is called automatically right after the object has been created:
# def __init__(self, name,id):
#
# Each attribute in a class definition begins with a reference to the instance object. It is by convention named self. The name,id is the argument. The value is passed during the class instantiation.
# self.name = name
# self.id=id
#
# -
# # Methods in a class
# Methods are functions defined inside the body of a class. They are used to perform operations with the attributes of our objects. Methods are essential in encapsulation concept of the OOP paradigm
#example: creating circle class with methods area()
class circle:
pi=3.13 #circle class object
def __init__(self,radius=1):#intializing radius=1
self.radius=radius
def area(self,radius):
return self.radius * self.radius * circle.pi
p=circle()
p.area(1)
# # Inheritance of a class
# Inheritance is a way to form new classes using classes that have already been defined. The newly formed classes are called derived classes, the classes that we derive from are called base classes. Important benefits of inheritance are code reuse and reduction of complexity of a program. The derived classes (descendants) override or extend the functionality of base classes (ancestors).
# +
#example of Inheritance animal and dog class
class Animal(object):
def __init__(self):
print ("Animal created")
def whoAmI(self):
print ("Animal")
def eat(self):
print ("Eating")
class Dog(Animal):
def __init__(self):
Animal.__init__(self)
print ("Dog created")
def whoAmI(self):
print ("Dog")
def bark(self):
print ("Woof!")
# + active=""
# In this example, we have two classes: Animal and Dog. The Animal is the base class, the Dog is the derived class.
#
# The derived class inherits the functionality of the base class.
#
# It is shown by the eat() method.
# The derived class modifies existing behavior of the base class.
#
# shown by the whoAmI() method.
# Finally, the derived class extends the functionality of the base class, by defining a new bark() method
# -
# # Errors and Excedptional handling
# https://docs.python.org/2/library/exceptions.html
# type of error and description is known as an Exception
#
# Even if a statement or expression is syntactically correct, it may cause an error when an attempt is made to execute it. Errors detected during execution are called exceptions and are not unconditionally fatal.
# ### Try and Expect
# The basic terminology and syntax used to handle errors in Python is the try and except statements. The code which can cause an exception to occue is put in the try block and the handling of the exception is the implemented in the except block of code
# + active=""
# General syntax of try and except:-
#
# try:
# You do your operations here...
# ...
# except ExceptionI:
# If there is ExceptionI, then execute this block.
# except ExceptionII:
# If there is ExceptionII, then execute this block.
# ...
# else:
# If there is no exception then execute this block
# -
try:
f = open('testfile','w')
f.write('Test write this')
except IOError:
# This will only check for an IOError exception and then execute this print statement
print ("Error: Could not find file or read data")
else:
print ("Content written successfully")
f.close()
# Now lets see what would happen if we did not have write permission (opening only with 'r'):
#
#
try:
f = open('testfile','r')
f.write('Test write this')
except IOError:
# This will only check for an IOError exception and then execute this print statement
print ("Error: Could not find file or read data")
else:
print ("Content written successfully")
f.close()
# Great! Notice how we only printed a statement! The code still ran and we were able to continue doing actions and running code blocks. This is extremely useful when you have to account for possible input errors in your code. You can be prepared for the error and keep running code, instead of your code just breaking as we saw above.
# # finally
# Great! Now we don't actually need to memorize that list of exception types! Now what if we kept wanting to run code after the exception occurred? This is where finally comes in.
# The finally: block of code will always be run regardless if there was an exception in the try code block. The syntax is:
#
#
#
# + active=""
# General syntax of finally:
#
# try:
# Code block here
# ...
# Due to any exception, this code may be skipped!
# finally:
# This code block would always be executed
# -
try:
f = open("testfile", "w")
f.write("Test write statement")
finally:
print ("Always execute finally code blocks")
# # Example for try and except
def askint():
try:
val = int(input("Please enter an integer: "))
except:
print ("Looks like you did not enter an integer!")
finally:
print ("Finally, I executed!")
print (val)
askint()
# Good! now try entering an string value and see the error
def askint():
try:
val = int(input("Please enter an integer: "))
except:
print ("Looks like you did not enter an integer!")
finally:
print ("Finally, I executed!")
print (val)
askint()
def askint():
try:
val = int(input("Please enter an integer: "))
except:
print ("Looks like you did not enter an integer!")
val = int(input("Try again-Please enter an integer: "))
finally:
print ("Finally, I executed!")
print (val)
askint()
# # Important Built-in Functions
# # map
# map() is a function that takes in two arguments: a function and a sequence iterable. In the form: map(function, sequence)
#
#
# The first argument is the name of a function and the second a sequence (e.g. a list). map() applies the function to all the elements of the sequence. It returns a new list with the elements changed by function
#map example
def calculateSquare(n):
return n*n
numbers = (1, 2, 3, 4)
result=map(calculateSquare, numbers)
set(result)
#map with lambda example
numbers = (1, 2, 3, 4)
result = map(lambda x: x*x, numbers)
set(result)
# # reduce
# The function reduce(function, sequence) continually applies the function to the sequence. It then returns a single value
from functools import reduce
lst =[47,11,42,13]
reduce(lambda x,y: x+y,lst)
from IPython.display import Image
Image('http://www.python-course.eu/images/reduce_diagram.png')
#reduce example
#Find the maximum of a sequence (This already exists as max())
max_find = lambda a,b: a if (a > b) else b
reduce(max_find,lst)
# # zip
# zip() makes an iterator that aggregates elements from each of the iterables.
#
#
#
numberList = [1, 2, 3]
strList = ['one', 'two', 'three']
# +
# Two iterables are passed
result = zip(numberList, strList)
# Converting itertor to set
resultSet = set(result)
print(resultSet)
# -
# # Enumerate
# Enumerate allows you to keep a count as you iterate through an object. It does this by returning a tuple in the form (count,element)
# +
lst = ['a','b','c']
for number,item in enumerate(lst):
print (number)
print (item)
# +
#enumerate example
grocery = ['bread', 'milk', 'butter']
for item in enumerate(grocery):
print(item)
print('\n')
for count, item in enumerate(grocery):
print(count, item)
print('\n')
# changing default start value
for count, item in enumerate(grocery, 100):
print(count, item)
# -
# # any() and all()
# all() and any() are built-in functions in Python that allow us to conveniently check for boolean matching in an iterable. all() will return True if all elements in an iterable are True. It is the same as this function code
list = [True,True,False,True]
any(list)
# Returns True because at least one of the elements in the list is True
#
#
all(list)
# Returns False because not all elements are True.
#
#
| Advanced Python/Advanced Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import cv2
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
def consecutive(data, stepsize=1):
return np.split(data, np.where(np.diff(data) != stepsize)[0]+1)
def segment_char(image_loc):
img = cv2.imread(image_loc,0)
img_copy = img.copy()
img_trans = np.transpose(img)
img_trans_copy = img_trans.copy()
width = np.size(img_trans,1)
line_ligature = []
for i , val in enumerate(img_trans):
if (sum(val) == 255 ):
line_ligature.append(i)
last = line_ligature[-1]
res = consecutive(line_ligature)
res = [i.tolist() for i in res]
print ("res is \n\n",res)
for val in res :
print (val)
times = 3
while(times > 0):
for i , val in enumerate(res):
try:
print (res[i+1][0] , val[-1] , res[i+1][0] - val[-1])
if (res[i+1][0] - val[-1] < 30):
val.append(res[i])
res.remove(res[i])
except:
pass
times = times - 1
print ("RES IS ",res,"\n\n\n\n\n\n")
print ("fianl RES",res)
rslt = []
for val in res:
n = (sum(val) // len(val) )
# if (len(val)>=7):
rslt.append(n)
print ("segm",rslt)
for val in rslt:
# if(last - val) > 50 :
img_trans_copy[val] = [255] * width
img_trans_copy = cv2.dilate(img_trans_copy , None , iterations = 1)
plt.imshow(np.transpose(img_trans_copy))
plt.show()
segment_char("lcts.png")
# segment_char("lrev.png")
# segment_char("lmat.png")
# segment_char("lcat.png")
# -
# find top and bottom layer
image_loc = "lcat.png"
img = cv2.imread(image_loc,0)
areas_of = []
for i , val in enumerate(img):
if (sum(val)>0):
areas_of.append(i)
else:
pass
print (areas_of[0],areas_of[-1])
plt.imshow(img)
38+380
418/2
img[198] = [255]*700
plt.imshow(img)
transpose = np.transpose(img)
plt.imshow(transpose)
transpose = cv2.dilate(transpose , None , iterations = 1)
plt.imshow(transpose)
255*4
line_ligature = []
for i ,val in enumerate(transpose):
if( (sum(val[198:]) == 255) and sum(val) == 255):
line_ligature.append(i)
print("Its There",i)
trans_copy = transpose.copy()
width = np.size(trans_copy,1)
for val in line_ligature:
trans_copy[val] = [255] * width
plt.imshow(np.transpose(trans_copy))
for val in line_ligature:
print(val,end=",")
def consecutive(data, stepsize=1):
return np.split(data, np.where(np.diff(data) != stepsize)[0]+1)
res = consecutive(line_ligature)
for val in res:
print (val)
arr = []
for i , val in enumerate(res):
if(len(val) > 7):
avg = sum(val) // len(val)
arr.append(avg)
arr
for val in arr:
# if()
trans_copy[val] = [255] * width
trans_copy = cv2.dilate(trans_copy,None,iterations=1)
plt.imshow(np.transpose(trans_copy))
cv2.imwrite("dear_t.png",np.transpose(trans_copy))
| Notebooks/myCharSeg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import re
from nltk.tokenize import sent_tokenize
class1_df = pd.read_csv('../data/eubank_class1_tagged.csv', sep=';', encoding='utf-8', index_col=0)
class1_df.head()
class2_df = pd.read_csv('../data/eubank_class2_tagged.csv', sep=';', encoding='utf-8', index_col=0)
class2_df.head()
def read_list_as_is(s):
if type(s) == list:
return s
## Strip brackets
s = s.strip('[').strip(']')
##
los = s.split(',')
los = list(set([s.strip() for s in los]))
return los
class1_df['entities'] = class1_df.entities.apply(lambda x: read_list_as_is(x))
class2_df['entities'] = class2_df.entities.apply(lambda x: read_list_as_is(x))
# +
class1_entity_list = class1_df.entities.tolist()
## Flattening list
class1_entity_list = list(set([e for sublist in class1_entity_list for e in sublist]))
class2_entity_list = class2_df.entities.tolist()
## Flattening list
class2_entity_list = list(set([e for sublist in class2_entity_list for e in sublist]))
full_entity_list = list(set(class1_entity_list + class2_entity_list))
# -
def entity_name(loe):
"""
entity_name consumes a list of entities, "loe", and
returns the dictionary of (entity name, tied entity name)
"""
#to_be_tied = [e for e in loe if " " in e]
tied = [e.replace(" ","_") for e in loe]
replace_dict = dict(zip(loe, tied))
replace_dict['.. Royal Bank of Scotland'] = 'Royal_Bank_of_Scotland'
return replace_dict
entity_dict = entity_name(full_entity_list)
# +
# https://stackoverflow.com/a/43966667
def replace_month_abbrev(input_str):
month_dict = {"Jan ": "January ",
"Feb ": "February ",
"Mar ": "March ",
"Apr ": "April ",
"May ": "May ",
"Jun ": "June ",
"Jul ": "July ",
"Aug ": "August ",
"Sep ": "September ",
"Sept ": "September ",
"Oct ": "October ",
"Nov ": "November ",
"Dec ": "December ",
"Jan. ": "January ",
"Feb. ": "February ",
"Mar. ": "March ",
"Apr. ": "April ",
"May. ": "May ",
"Jun. ": "June ",
"Jul. ": "July ",
"Aug. ": "August ",
"Sep. ": "September ",
"Sept. ": "September ",
"Oct. ": "October ",
"Nov. ": "November ",
"Dec. ": "December "}
# find all dates with abrev
abbrev_found = filter(lambda abbrev_month: abbrev_month in input_str, month_dict.keys())
# replace each date with its abbreviation
for abbrev in abbrev_found:
input_str = input_str.replace(abbrev, month_dict[abbrev])
# return the modified string (or original if no states were found)
return input_str
#combined['deadline'].map(replace_month_abbrev(h_abbr))
# -
def clean_paragraph(input_para, input_entity):
"""
The only 'brute force' involved in this part is when the period is succeeded by a non-numeric word and followed by a number.
We cannot simly tokenize the above case using str.split('.') since it will split the date apart.
i.e. Nov. 24
We will find all possible abbrevation of these months and write them out explicitly such that we do not have a period in between the date.
For the such fix, look at replace_month_abrev() function above.
"""
new_para = replace_month_abbrev(input_para)
for e in input_entity:
new_para = new_para.replace(e, entity_dict.get(e))
for old, new in entity_dict.items():
new_para = new_para.replace(old,new)
## List all possible cases where a space (or two) does not succeed the period.
## Alphanumeric followed by another alphanumeric (+special characters)
possible_exceptions = set(re.findall('([0-9A-Za-z]+[.][0-9A-Za-z]+)',input_para))
## $30.5m
allowed_list = set(re.findall('[0-9A-Za-z]*\d+[.]\d+[0-9A-Za-z]*',input_para))
exception_list = list(possible_exceptions.difference(allowed_list))
new_list = [e.replace('.','. ') for e in exception_list]
replace_dict = dict(zip(exception_list, new_list))
for old,new in replace_dict.items():
new_para = new_para.replace(old,new)
new_para = new_para.replace(';',',') ## Using ; as separator, so replacing any ; with ,
new_para = new_para.strip().strip('\t')
return new_para
def returnSentences(input_text):
## Don't have to run clean_paragraph twice.
#clean_para = clean_paragraph(input_text)
sent_list= sent_tokenize(input_text)
clean_sent_list = [sent.strip().strip('\t').strip('\b').strip('\n') for sent in sent_list]
return clean_sent_list
def returnRevelvantSentences(sent_list, entity_list):
if entity_list == []:
return []
## "set" to remove duplicate sentences. Sometimes a sentence can mention the same entity twice in the sentence.
relevant_sent_list = list(set([sent.strip().strip('\t').strip('\b') for sent in sent_list
if any(entity.lower() in sent.lower() for entity in entity_list)]))
return relevant_sent_list
class1_df.head()
#class1_df['entities'] = class1_df.entities.apply(lambda x: read_list_as_is(x))
class1_df['txt'] = class1_df.apply(lambda row: clean_paragraph(row['txt'], row['entities']), axis = 1)
class1_df['entities'] = class1_df.entities.apply(lambda row: list(map(entity_dict.get, row)))
class1_df['sentences'] = class1_df.txt.apply(lambda row: returnSentences(row))
class1_df['relevant_sent_list'] = class1_df.apply(lambda row: returnRevelvantSentences(row['sentences'], row['entities']), axis = 1)
class1_df = class1_df[class1_df['sentences'].apply(lambda x: len(x)) > 0]
class1_df = class1_df[class1_df['relevant_sent_list'].apply(lambda x: len(x)) > 0]
#class2_df['entities'] = class2_df.entities.apply(lambda x: read_list_as_is(x))
class2_df['txt'] = class2_df.apply(lambda row: clean_paragraph(row['txt'], row['entities']), axis = 1)
class2_df['entities'] = class2_df.entities.apply(lambda row: list(map(entity_dict.get, row)))
class2_df['sentences'] = class2_df.txt.apply(lambda row: returnSentences(row))
class2_df['relevant_sent_list'] = class2_df.apply(lambda row: returnRevelvantSentences(row['sentences'], row['entities']), axis = 1)
class2_df = class2_df[class2_df['sentences'].apply(lambda x: len(x)) > 0]
class2_df = class2_df[class2_df['relevant_sent_list'].apply(lambda x: len(x)) > 0]
#
#
## Labelling
class1_df['Y'] = 1
class2_df['Y'] = 0
print(class1_df.permid.nunique())
print(class2_df.permid.nunique())
# 12 banks only have articles that discuss the distress, not before and after the event. We exclude these banks from our analysis
excluded_permids = list(set(class1_df.permid.to_list()).difference(set(class2_df.permid.to_list())))
class2_df = class2_df[~(class2_df.permid.isin(excluded_permids))]
print(class2_df.shape)
# Explode list of relevant sentences to separte rows
# +
c1_df = class1_df.explode('relevant_sent_list')[['permid', 'relevant_sent_list','Y']].reset_index(drop = True)
c2_df = class2_df.explode('relevant_sent_list')[['permid', 'relevant_sent_list', "Y"]].reset_index(drop = True)
c1_df.head()
# -
print(c1_df.shape)
print(c2_df.shape)
c1_df.to_pickle('../data/class1.pkl')
c2_df.to_pickle('../data/class2.pkl')
| notebook/2. Extract Relevant Sentences.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# importing the necessary libraries
import os
import cv2
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import random
from tqdm import tqdm
import pandas as pd
from skimage.io import imread, imshow
from skimage.transform import resize
import scipy.ndimage
import IPython
from sklearn.preprocessing import StandardScaler
from tensorflow import keras
from tensorflow.keras import optimizers
import sklearn.model_selection
import tensorflow.keras.backend as K
# %matplotlib inline
base_dir = '/home/kiara/Desktop/CMR_Metric_Calculator/Dataset_Segmentation/'
train_img_dir = os.path.join(base_dir, 'PNG_images/')
train_label_dir = os.path.join(base_dir, 'PNG_labels/')
# obtaining the training image (and corresponding label (masks)) file names as a list
train_img_fname = os.listdir(train_img_dir)
train_label_fname = train_img_fname
# shuffling the image list randomply and saving it
train_img_fnames = random.sample(train_img_fname, len(train_img_fname))
train_label_fnames = train_img_fnames
print(len(train_label_fnames))
training_dataset, test_dataset = sklearn.model_selection.train_test_split(train_img_fnames, test_size=0.1)
train_img_fnames = training_dataset
train_label_fnames = train_img_fnames
test_img_fnames = test_dataset
test_label_fnames = test_img_fnames
IMG_WIDTH = 256
IMG_HEIGHT = 256
IMG_CHANNEL = 1
IMG_CHANNELS = 3
len(test_img_fnames)
# sanity check
print(len(train_img_fnames))
# creating an array of the same dimension as the input images
X_train = np.zeros((2*len(train_img_fnames), IMG_HEIGHT, IMG_WIDTH, IMG_CHANNEL), dtype = np.float32)
Y_train = np.zeros((2*len(train_img_fnames), IMG_HEIGHT, IMG_WIDTH, IMG_CHANNELS), dtype = np.float32)
#print("Resizing train images")
from numpy import asarray
from PIL import Image
for n, id_ in tqdm(enumerate(train_img_fnames), total=len(train_img_fnames)):
n=n*2
img = imread(train_img_dir + id_) # read the image
pixels=asarray(img).astype('float32')
pixels = resize(pixels, (IMG_HEIGHT, IMG_WIDTH, IMG_CHANNEL), mode = 'constant', preserve_range = True)
# convert from integers to floats
pixels = pixels.astype('float32')
# normalize to the range 0-1
pixels /= 255.0
# confirm the normalization
X_train[n] = pixels.astype('float32')
# rotate only
img = imread(train_img_dir + id_) # read the image
r_img = cv2.rotate(img, cv2.ROTATE_90_CLOCKWISE)
pixels1=asarray(r_img).astype('float32')
pixels1 = resize(pixels1, (IMG_HEIGHT, IMG_WIDTH, IMG_CHANNEL), mode = 'constant', preserve_range = True)
# convert from integers to floats
pixels1 = pixels1.astype('float32')
# normalize to the range 0-1
pixels1 /= 255.0
# confirm the normalization
X_train[n+1] = pixels1.astype('float32')
'''
img = imread(train_img_dir + id_) # read the image
r_img = cv2.rotate(img, cv2.ROTATE_90_COUNTERCLOCKWISE)
pixels1=asarray(r_img).astype('float32')
pixels1 = resize(pixels1, (IMG_HEIGHT, IMG_WIDTH, IMG_CHANNEL), mode = 'constant', preserve_range = True)
# convert from integers to floats
pixels1 = pixels1.astype('float32')
# normalize to the range 0-1
pixels1 /= 255.0
# confirm the normalization
X_train[n+2] = pixels1.astype('float32')
# rotate and CLAHE
img = cv2.imread((train_img_dir + id_), IMG_CHANNEL) # read the image
# rotate the image
r_img = cv2.rotate(img, cv2.ROTATE_90_CLOCKWISE)
#Converting image to LAB Color so CLAHE can be applied to the luminance channel
lab_img= cv2.cvtColor(r_img, cv2.COLOR_BGR2LAB)
#Splitting the LAB image to L, A and B channels, respectively
l, a, b = cv2.split(lab_img)
#Apply histogram equalization to the L channel
equ = cv2.equalizeHist(l)
#Combine the Hist. equalized L-channel back with A and B channels
updated_lab_img1 = cv2.merge((equ,a,b))
#Apply CLAHE to L channel
clahe = cv2.createCLAHE(clipLimit=3.0, tileGridSize=(8,8))
clahe_img = clahe.apply(l)
#Combine the CLAHE enhanced L-channel back with A and B channels
updated_lab_img2 = cv2.merge((clahe_img,a,b))
#Convert LAB image back to color (RGB)
CLAHE_img = cv2.cvtColor(updated_lab_img2, cv2.COLOR_LAB2BGR)
pixels=asarray(CLAHE_img).astype('float32')
pixels = resize(pixels, (IMG_HEIGHT, IMG_WIDTH, IMG_CHANNEL), mode = 'constant', preserve_range = True)
# convert from integers to floats
pixels = pixels.astype('float32')
# normalize to the range 0-1
pixels /= 255.0
X_train[n+3] = pixels.astype('float32')'''
'''# Remove comments to perform Augmentation
#print("-----------------------CLAHE and ROTATE------------------")
img = cv2.imread((train_img_dir + id_), IMG_CHANNEL) # read the image
# rotate the image
r_img = cv2.rotate(img, cv2.ROTATE_90_CLOCKWISE)
#Converting image to LAB Color so CLAHE can be applied to the luminance channel
lab_img= cv2.cvtColor(r_img, cv2.COLOR_BGR2LAB)
#Splitting the LAB image to L, A and B channels, respectively
l, a, b = cv2.split(lab_img)
#Apply histogram equalization to the L channel
equ = cv2.equalizeHist(l)
#Combine the Hist. equalized L-channel back with A and B channels
updated_lab_img1 = cv2.merge((equ,a,b))
#Apply CLAHE to L channel
clahe = cv2.createCLAHE(clipLimit=3.0, tileGridSize=(8,8))
clahe_img = clahe.apply(l)
#Combine the CLAHE enhanced L-channel back with A and B channels
updated_lab_img2 = cv2.merge((clahe_img,a,b))
#Convert LAB image back to color (RGB)
CLAHE_img = cv2.cvtColor(updated_lab_img2, cv2.COLOR_LAB2BGR)
pixels=asarray(CLAHE_img).astype('float32')
pixels = resize(pixels, (IMG_HEIGHT, IMG_WIDTH, IMG_CHANNEL), mode = 'constant', preserve_range = True)
# convert from integers to floats
pixels = pixels.astype('float32')
# normalize to the range 0-1
pixels /= 255.0
X_train[n+3] = pixels.astype('float32')'''
'''
#print("-----------------------CLAHE ONLY ------------------")
img = cv2.imread((train_img_dir + id_), IMG_CHANNEL) # read the image
# rotate the image
r_img = img
#Converting image to LAB Color so CLAHE can be applied to the luminance channel
lab_img= cv2.cvtColor(r_img, cv2.COLOR_BGR2LAB)
#Splitting the LAB image to L, A and B channels, respectively
l, a, b = cv2.split(lab_img)
#Apply histogram equalization to the L channel
equ = cv2.equalizeHist(l)
#Combine the Hist. equalized L-channel back with A and B channels
updated_lab_img1 = cv2.merge((equ,a,b))
#Apply CLAHE to L channel
clahe = cv2.createCLAHE(clipLimit=3.0, tileGridSize=(8,8))
clahe_img = clahe.apply(l)
#Combine the CLAHE enhanced L-channel back with A and B channels
updated_lab_img2 = cv2.merge((clahe_img,a,b))
#Convert LAB image back to color (RGB)
CLAHE_img = cv2.cvtColor(updated_lab_img2, cv2.COLOR_LAB2BGR)
pixels=asarray(CLAHE_img).astype('float32')
pixels = resize(pixels, (IMG_HEIGHT, IMG_WIDTH, IMG_CHANNEL), mode = 'constant', preserve_range = True)
# convert from integers to floats
pixels = pixels.astype('float32')
# normalize to the range 0-1
pixels /= 255.0
X_train[n+2] = pixels.astype('float32')
'''
'''#print("-----------------------CLAHE AND ROTATE COUNTER ONLY------------------")
img = cv2.imread((train_img_dir + id_), 1) # read the image
# rotate the image
r_img = cv2.rotate(img, cv2.ROTATE_90_COUNTERCLOCKWISE)
#Converting image to LAB Color so CLAHE can be applied to the luminance channel
lab_img= cv2.cvtColor(r_img, cv2.COLOR_BGR2LAB)
#Splitting the LAB image to L, A and B channels, respectively
l, a, b = cv2.split(lab_img)
#Apply histogram equalization to the L channel
equ = cv2.equalizeHist(l)
#Combine the Hist. equalized L-channel back with A and B channels
updated_lab_img1 = cv2.merge((equ,a,b))
#Apply CLAHE to L channel
clahe = cv2.createCLAHE(clipLimit=3.0, tileGridSize=(8,8))
clahe_img = clahe.apply(l)
#Combine the CLAHE enhanced L-channel back with A and B channels
updated_lab_img2 = cv2.merge((clahe_img,a,b))
#Convert LAB image back to color (RGB)
CLAHE_img = cv2.cvtColor(updated_lab_img2, cv2.COLOR_LAB2BGR)
pixels=asarray(CLAHE_img).astype('float32')
pixels = resize(pixels, (IMG_HEIGHT, IMG_WIDTH, IMG_CHANNEL), mode = 'constant', preserve_range = True)
# convert from integers to floats
pixels = pixels.astype('float32')
# normalize to the range 0-1
pixels /= 255.0
X_train[n+2] = pixels.astype('float32')'''
print("Resizing train images")
from numpy import asarray
from PIL import Image
for n, id_ in tqdm(enumerate(train_img_fnames), total=len(train_img_fnames)):
n=n*2
img = imread(train_label_dir + id_) # read the image
pixels=asarray(img).astype('float32')
pixels = resize(pixels, (IMG_HEIGHT, IMG_WIDTH, IMG_CHANNELS), mode = 'constant', preserve_range = True)
# convert from integers to floats
pixels = pixels.astype('float32')
# normalize to the range 0-1
pixels /= 255.0
Y_train[n] = pixels.astype('float32')
#clahe and rotate
img = imread(train_label_dir + id_) # read the image
r_img = cv2.rotate(img, cv2.ROTATE_90_CLOCKWISE)
pixels1=asarray(r_img).astype('float32')
pixels1 = resize(pixels1, (IMG_HEIGHT, IMG_WIDTH, IMG_CHANNELS), mode = 'constant', preserve_range = True)
# convert from integers to floats
pixels1 = pixels1.astype('float32')
# normalize to the range 0-1
pixels1 /= 255.0
# confirm the normalization
Y_train[n+1] = pixels1.astype('float32')
'''
# clahe and rotate counter
img = imread(train_label_dir + id_) # read the image
r_img = cv2.rotate(img, cv2.ROTATE_90_COUNTERCLOCKWISE)
pixels1=asarray(r_img).astype('float32')
pixels1 = resize(pixels1, (IMG_HEIGHT, IMG_WIDTH, IMG_CHANNELS), mode = 'constant', preserve_range = True)
# convert from integers to floats
pixels1 = pixels1.astype('float32')
# normalize to the range 0-1
pixels1 /= 255.0
# confirm the normalization
Y_train[n+2] = pixels1.astype('float32')
Y_train[n+3] = Y_train[n+1]'''
'''
# clahe only
#Y_train[n+2] = Y_train[n]'''
'''# clahe and rotate counter
img = imread(train_label_dir + id_) # read the image
r_img = cv2.rotate(img, cv2.ROTATE_90_COUNTERCLOCKWISE)
pixels1=asarray(r_img).astype('float32')
pixels1 = resize(pixels1, (IMG_HEIGHT, IMG_WIDTH, IMG_CHANNELS), mode = 'constant', preserve_range = True)
# convert from integers to floats
pixels1 = pixels1.astype('float32')
# normalize to the range 0-1
pixels1 /= 255.0
# confirm the normalization
Y_train[n+2] = pixels1.astype('float32')'''
#Y_train[n+3] = Y_train[n+1]
# plotting an image
seed = 17
np.random.seed = seed
image_x = random.randint(0, len(train_img_fnames)) # generate a random number between 0 and length of training ids
imshow(np.squeeze(X_train[image_x]))
#plt.savefig("image.pdf", format='pdf')
plt.show()
imshow(np.squeeze(Y_train[image_x]))
#plt.savefig("label.pdf", format='pdf')
plt.show()
X_test = np.zeros((len(test_img_fnames), IMG_HEIGHT, IMG_WIDTH, IMG_CHANNEL), dtype = np.float32)
sizes_test = []
print("Resizing test images")
for n, id_ in tqdm(enumerate(test_img_fnames), total=len(test_img_fnames)):
path = base_dir
img = imread(train_img_dir + id_) # read the image
# Uncomment to test on HELIX Dataset
#img = imread('/media/kiara/My Passport/HELIX/image/' + id_)
pixels=asarray(img).astype('float32')
pixels = resize(pixels, (IMG_HEIGHT, IMG_WIDTH, IMG_CHANNEL), mode = 'constant', preserve_range = True)
# convert from integers to floats
pixels = pixels.astype('float32')
# normalize to the range 0-1
pixels /= 255.0
X_test[n] = pixels.astype('float32')
Y_test = np.zeros((len(test_label_fnames), IMG_HEIGHT, IMG_WIDTH, IMG_CHANNELS), dtype = np.float32)
print("Resizing test images")
from numpy import asarray
from PIL import Image
for n, id_ in tqdm(enumerate(test_img_fnames), total=len(test_img_fnames)):
#path = base_dir
img = imread(train_label_dir + id_) # read the image
#img = imread('/media/kiara/My Passport/HELIX/label/ShortAxis/' + id_)
pixels=asarray(img).astype('float32')
pixels = resize(pixels, (IMG_HEIGHT, IMG_WIDTH, IMG_CHANNELS), mode = 'constant', preserve_range = True)
# convert from integers to floats
pixels = pixels.astype('float32')
# normalize to the range 0-1
pixels /= 255.0
Y_test[n] = pixels.astype('float32')
seed = 17
np.random.seed = seed
image_x = random.randint(0, len(test_img_fnames)) # generate a random number between 0 and length of training ids
imshow(np.squeeze(X_test[image_x]))
#plt.savefig("image.pdf", format='pdf')
plt.show()
imshow(np.squeeze(Y_test[image_x]))
#plt.savefig("label.pdf", format='pdf')
plt.show()
# ## Metrics and Function Definitions
# +
def iou_coef(y_true, y_pred, smooth=1):
intersection = K.sum(K.abs(y_true * y_pred), axis=[1,2,3])
union = K.sum(y_true,[1,2,3])+K.sum(y_pred,[1,2,3])-intersection
iou = K.mean((intersection + smooth) / (union + smooth), axis=0)
return iou
def precision(y_true, y_pred): #taken from old keras source code
true_positives = K.sum(K.round(K.clip(y_true * y_pred, 0, 1)))
predicted_positives = K.sum(K.round(K.clip(y_pred, 0, 1)))
precision = true_positives / (predicted_positives + K.epsilon())
return precision
def recall(y_true, y_pred): #taken from old keras source code
true_positives = K.sum(K.round(K.clip(y_true * y_pred, 0, 1)))
possible_positives = K.sum(K.round(K.clip(y_true, 0, 1)))
recall = true_positives / (possible_positives + K.epsilon())
return recall
# +
def DC(y_true, y_pred, smooth=1):
"""
Dice = (2*|X & Y|)/ (|X|+ |Y|)
= 2*sum(|A*B|)/(sum(A^2)+sum(B^2))
ref: https://arxiv.org/pdf/1606.04797v1.pdf
"""
intersection = K.sum(K.abs(y_true * y_pred), axis=-1)
return (2. * intersection + smooth) / (K.sum(K.square(y_true),-1) + K.sum(K.square(y_pred),-1) + smooth)
def dice_coef_loss(y_true, y_pred):
return 1-DC(y_true, y_pred)
# -
# importing packages necessary for model training
import os
import cv2
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import random
from tqdm import tqdm
import pandas as pd
from skimage.io import imread, imshow
from skimage.transform import resize
import scipy.ndimage
from sklearn.metrics import confusion_matrix
from sklearn.metrics import roc_curve
import IPython
from sklearn.preprocessing import StandardScaler
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras import optimizers
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Conv2D, MaxPooling2D
from tensorflow.keras.layers import Activation, Dropout, Flatten, Dense
from numpy import asarray
from PIL import Image
import sklearn.model_selection
# ## Model
IMG_WIDTH = 256
IMG_HEIGHT = 256
IMG_CHANNEL = 1
IMG_CHANNELS = 3
# defining input layer
inputs = tf.keras.layers.Input((IMG_WIDTH, IMG_HEIGHT, IMG_CHANNEL))
# pixels to floating point numbers
s = tf.keras.layers.Lambda(lambda x: (x/255))(inputs)
print(s)
# # MODEL 1 (Unet with Dropout Layers)
# +
c1 = tf.keras.layers.Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(s)
c1 = tf.keras.layers.Dropout(0.5)(c1)
c1 = tf.keras.layers.Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(c1)
p1 = tf.keras.layers.MaxPooling2D((2, 2))(c1)
c2 = tf.keras.layers.Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(p1)
c2 = tf.keras.layers.Dropout(0.5)(c2)
c2 = tf.keras.layers.Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(c2)
p2 = tf.keras.layers.MaxPooling2D((2, 2))(c2)
c3 = tf.keras.layers.Conv2D(128, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(p2)
c3 = tf.keras.layers.Dropout(0.5)(c3)
c3 = tf.keras.layers.Conv2D(128, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(c3)
p3 = tf.keras.layers.MaxPooling2D((2, 2))(c3)
c4 = tf.keras.layers.Conv2D(256, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(p3)
c4 = tf.keras.layers.Dropout(0.5)(c4)
c4 = tf.keras.layers.Conv2D(256, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(c4)
p4 = tf.keras.layers.MaxPooling2D(pool_size=(2, 2))(c4)
c5 = tf.keras.layers.Conv2D(512, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(p4)
c5 = tf.keras.layers.Dropout(0.5)(c5)
c5 = tf.keras.layers.Conv2D(512, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(c5)
c5 = tf.keras.layers.Dropout(0.5)(c5)
# Expansion Path
u6 = tf.keras.layers.Conv2DTranspose(256, (2, 2), strides=(2, 2), padding='same')(c5)
u6 = tf.keras.layers.concatenate([u6, c4])
c6 = tf.keras.layers.Conv2D(256, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(u6)
c6 = tf.keras.layers.Dropout(0.5)(c6)
c6 = tf.keras.layers.Conv2D(512, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(c6)
u7 = tf.keras.layers.Conv2DTranspose(128, (2, 2), strides=(2, 2), padding='same')(c6)
u7 = tf.keras.layers.concatenate([u7, c3])
c7 = tf.keras.layers.Conv2D(128, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(u7)
c7 = tf.keras.layers.Dropout(0.5)(c7)
c7 = tf.keras.layers.Conv2D(128, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(c7)
u8 = tf.keras.layers.Conv2DTranspose(64, (2, 2), strides=(2, 2), padding='same')(c7)
u8 = tf.keras.layers.concatenate([u8, c2])
c8 = tf.keras.layers.Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(u8)
c8 = tf.keras.layers.Dropout(0.5)(c8)
c8 = tf.keras.layers.Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(c8)
u9 = tf.keras.layers.Conv2DTranspose(32, (2, 2), strides=(2, 2), padding='same')(c8)
u9 = tf.keras.layers.concatenate([u9, c1], axis=3)
c9 = tf.keras.layers.Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(u9)
c9 = tf.keras.layers.Dropout(0.5)(c9)
c9 = tf.keras.layers.Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(c9)
outputs = tf.keras.layers.Conv2D(3, (1,1), activation='sigmoid')(c9)
# -
model = tf.keras.Model(inputs=[inputs], outputs=[outputs])
opt = keras.optimizers.Adam(learning_rate=0.1)
model.compile(optimizer=opt, loss=tf.keras.losses.BinaryCrossentropy(), metrics=[DC, iou_coef, 'acc', precision, recall])
model.summary()
# +
# model checkpoint
checkpointer = tf.keras.callbacks.ModelCheckpoint(filepath='/home/kiara/Desktop/CMR_Metric_Calculator/UNET_WITH_2000RERUN.h5', verbose = 2, save_weights_only = True)
callbacks = [
tf.keras.callbacks.EarlyStopping(patience = 50, monitor = 'val_loss'),
tf.keras.callbacks.TensorBoard(log_dir = 'Log_UNET_WITH_2000RERUN')
]
# change name to 2000RERUNAug
# FIT MODEL
results = model.fit(X_train, Y_train, validation_split = 0.1, batch_size = 4, epochs = 150, callbacks=callbacks)
model.save('model_UNET_WITH_2000RERUN')
model.save('model_UNET_WITH_2000RERUN.h5')
# -
model.evaluate(X_test, Y_test, verbose=1)
# # MODEL 2 (Unet without Dropout Layers)
# +
#Contraction path
c1 = tf.keras.layers.Conv2D(32, (3, 3), activation='tanh', kernel_initializer='he_normal', padding='same')(s)
#c1 = tf.keras.layers.Dropout(0.5)(c1)
c1 = tf.keras.layers.Conv2D(32, (3, 3), activation='tanh', kernel_initializer='he_normal', padding='same')(c1)
p1 = tf.keras.layers.MaxPooling2D((2, 2))(c1)
c2 = tf.keras.layers.Conv2D(64, (3, 3), activation='tanh', kernel_initializer='he_normal', padding='same')(p1)
#c2 = tf.keras.layers.Dropout(0.5)(c2)
c2 = tf.keras.layers.Conv2D(64, (3, 3), activation='tanh', kernel_initializer='he_normal', padding='same')(c2)
p2 = tf.keras.layers.MaxPooling2D((2, 2))(c2)
c3 = tf.keras.layers.Conv2D(128, (3, 3), activation='tanh', kernel_initializer='he_normal', padding='same')(p2)
#c3 = tf.keras.layers.Dropout(0.4)(c3)
c3 = tf.keras.layers.Conv2D(128, (3, 3), activation='tanh', kernel_initializer='he_normal', padding='same')(c3)
p3 = tf.keras.layers.MaxPooling2D((2, 2))(c3)
c4 = tf.keras.layers.Conv2D(256, (3, 3), activation='tanh', kernel_initializer='he_normal', padding='same')(p3)
#c4 = tf.keras.layers.Dropout(0.5)(c4)
c4 = tf.keras.layers.Conv2D(256, (3, 3), activation='tanh', kernel_initializer='he_normal', padding='same')(c4)
p4 = tf.keras.layers.MaxPooling2D(pool_size=(2, 2))(c4)
c5 = tf.keras.layers.Conv2D(512, (3, 3), activation='tanh', kernel_initializer='he_normal', padding='same')(p4)
#c5 = tf.keras.layers.Dropout(0.4)(c5)
c5 = tf.keras.layers.Conv2D(512, (3, 3), activation='tanh', kernel_initializer='he_normal', padding='same')(c5)
#c5 = tf.keras.layers.Dropout(0.5)(c5)
# Expansion Path
u6 = tf.keras.layers.Conv2DTranspose(256, (2, 2), strides=(2, 2), padding='same')(c5)
u6 = tf.keras.layers.concatenate([u6, c4])
c6 = tf.keras.layers.Conv2D(256, (3, 3), activation='tanh', kernel_initializer='he_normal', padding='same')(u6)
#c6 = tf.keras.layers.Dropout(0.4)(c6)
c6 = tf.keras.layers.Conv2D(512, (3, 3), activation='tanh', kernel_initializer='he_normal', padding='same')(c6)
u7 = tf.keras.layers.Conv2DTranspose(128, (2, 2), strides=(2, 2), padding='same')(c6)
u7 = tf.keras.layers.concatenate([u7, c3])
c7 = tf.keras.layers.Conv2D(128, (3, 3), activation='tanh', kernel_initializer='he_normal', padding='same')(u7)
#c7 = tf.keras.layers.Dropout(0.5)(c7)
c7 = tf.keras.layers.Conv2D(128, (3, 3), activation='tanh', kernel_initializer='he_normal', padding='same')(c7)
u8 = tf.keras.layers.Conv2DTranspose(64, (2, 2), strides=(2, 2), padding='same')(c7)
u8 = tf.keras.layers.concatenate([u8, c2])
c8 = tf.keras.layers.Conv2D(64, (3, 3), activation='tanh', kernel_initializer='he_normal', padding='same')(u8)
#c8 = tf.keras.layers.Dropout(0.5)(c8)
c8 = tf.keras.layers.Conv2D(64, (3, 3), activation='tanh', kernel_initializer='he_normal', padding='same')(c8)
u9 = tf.keras.layers.Conv2DTranspose(32, (2, 2), strides=(2, 2), padding='same')(c8)
u9 = tf.keras.layers.concatenate([u9, c1], axis=3)
c9 = tf.keras.layers.Conv2D(32, (3, 3), activation='tanh', kernel_initializer='he_normal', padding='same')(u9)
#c9 = tf.keras.layers.Dropout(0.4)(c9)
c9 = tf.keras.layers.Conv2D(32, (3, 3), activation='tanh', kernel_initializer='he_normal', padding='same')(c9)
outputs = tf.keras.layers.Conv2D(3, (1,1), activation='sigmoid')(c9)
# -
model = tf.keras.Model(inputs=[inputs], outputs=[outputs])
opt = keras.optimizers.Adam(learning_rate=0.1)
model.compile(optimizer=opt, loss=tf.keras.losses.BinaryCrossentropy(), metrics=[DC, iou_coef, 'acc', precision, recall])
model.summary()
# +
# model checkpoint
checkpointer = tf.keras.callbacks.ModelCheckpoint(filepath='/home/kiara/Desktop/CMR_Metric_Calculator/UNET_WITHOUT_10000AugRERUN.h5', verbose = 2, save_weights_only = True)
callbacks = [
tf.keras.callbacks.EarlyStopping(patience = 50, monitor = 'val_loss'),
tf.keras.callbacks.TensorBoard(log_dir = 'Log_UNET_WITHOUT_10000AugRERUN')
]
# FIT MODEL
results = model.fit(X_train, Y_train, validation_split = 0.1, batch_size = 4, epochs = 150, callbacks=callbacks)
model.save('model_UNET_WITHOUT_10000AugRERUN')
model.save('model_UNET_WITHOUT_10000AugRERUN.h5')
# -
model.evaluate(X_test, Y_test, verbose=1)
# # MODEL 3 (ResUnet)
# +
# taken directly from the original implementation https://arxiv.org/pdf/1711.10684.pdf
def bn_act(x, act=True):
x = tf.keras.layers.BatchNormalization()(x)
if act == True:
x = tf.keras.layers.Activation("tanh")(x)
#x = tf.keras.layers.Activation("sigmoid")(x)
return x
def conv_block(x, filters, kernel_size=(3, 3), padding="same", strides=1):
conv = bn_act(x)
conv = tf.keras.layers.Conv2D(filters, kernel_size, padding=padding, strides=strides)(conv)
return conv
def stem(x, filters, kernel_size=(3, 3), padding="same", strides=1):
conv = tf.keras.layers.Conv2D(filters, kernel_size, padding=padding, strides=strides)(x)
conv = conv_block(conv, filters, kernel_size=kernel_size, padding=padding, strides=strides)
shortcut = tf.keras.layers.Conv2D(filters, kernel_size=(1, 1), padding=padding, strides=strides)(x)
shortcut = bn_act(shortcut, act=False)
output = tf.keras.layers.Add()([conv, shortcut])
return output
def residual_block(x, filters, kernel_size=(3, 3), padding="same", strides=1):
res = conv_block(x, filters, kernel_size=kernel_size, padding=padding, strides=strides)
res = conv_block(res, filters, kernel_size=kernel_size, padding=padding, strides=1)
shortcut = tf.keras.layers.Conv2D(filters, kernel_size=(1, 1), padding=padding, strides=strides)(x)
shortcut = bn_act(shortcut, act=False)
output = tf.keras.layers.Add()([shortcut, res])
return output
def upsample_concat_block(x, xskip):
u = tf.keras.layers.UpSampling2D((2, 2))(x)
c = tf.keras.layers.concatenate([u, xskip])
return c
# -
# taken directly from the original implementation https://arxiv.org/pdf/1711.10684.pdf
def ResUNet():
#f = [24, 48, 96, 192, 384]
#f = [8, 16, 32, 64, 128]
#f = [64, 128, 256, 512, 1024]
#f = [8, 16, 32, 64, 128]#1st
# 16 2nd then 4 3rd
f = [4, 8, 16, 32, 64]
#f = [32, 64, 128, 256, 512]
inputs = keras.layers.Input((256, 256, 1))
## Encoder
e0 = inputs
e1 = stem(e0, f[0])
e2 = residual_block(e1, f[1], strides=2)
e3 = residual_block(e2, f[2], strides=2)
e4 = residual_block(e3, f[3], strides=2)
e5 = residual_block(e4, f[4], strides=2)
## Bridge
b0 = conv_block(e5, f[4], strides=1)
b1 = conv_block(b0, f[4], strides=1)
## Decoder
u1 = upsample_concat_block(b1, e4)
d1 = residual_block(u1, f[4])
u2 = upsample_concat_block(d1, e3)
d2 = residual_block(u2, f[3])
u3 = upsample_concat_block(d2, e2)
d3 = residual_block(u3, f[2])
u4 = upsample_concat_block(d3, e1)
d4 = residual_block(u4, f[1])
outputs = keras.layers.Conv2D(3, (1, 1), padding="same", activation="sigmoid")(d4)
model = keras.models.Model(inputs, outputs)
return model
model = ResUNet()
from keras.utils import to_categorical
opt = keras.optimizers.Adam(learning_rate=0.1)
model.compile(optimizer=opt, loss=tf.keras.losses.BinaryCrossentropy(), metrics=[DC, iou_coef, 'acc', precision, recall])
model.summary()
# +
# model checkpoint
checkpointer = tf.keras.callbacks.ModelCheckpoint(filepath='/home/kiara/Desktop/CMR_Metric_Calculator/RESUNET_10000.h5', verbose = 2, save_weights_only = True)
callbacks = [
tf.keras.callbacks.EarlyStopping(patience = 50, monitor = 'val_loss'),
tf.keras.callbacks.TensorBoard(log_dir = 'Log_RESUNET_10000')
]
# FIT MODEL
results = model.fit(X_train, Y_train, validation_split = 0.1, batch_size = 4, epochs = 150, callbacks=callbacks)
model.save('model_RESUNET_10000')
model.save('model_RESUNET_10000.h5')
# -
model.evaluate(X_test, Y_test, verbose=1)
# # MODEL 4 (FCN)
# +
c1 = tf.keras.layers.Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(s)
c1 = tf.keras.layers.Dropout(0.5)(c1)
c1 = tf.keras.layers.Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(c1)
p1 = tf.keras.layers.MaxPooling2D((2, 2))(c1)
c2 = tf.keras.layers.Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(p1)
c2 = tf.keras.layers.Dropout(0.5)(c2)
c2 = tf.keras.layers.Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(c2)
p2 = tf.keras.layers.MaxPooling2D((2, 2))(c2)
c3 = tf.keras.layers.Conv2D(128, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(p2)
c3 = tf.keras.layers.Dropout(0.5)(c3)
c3 = tf.keras.layers.Conv2D(128, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(c3)
p3 = tf.keras.layers.MaxPooling2D((2, 2))(c3)
c4 = tf.keras.layers.Conv2D(256, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(p3)
c4 = tf.keras.layers.Dropout(0.5)(c4)
c4 = tf.keras.layers.Conv2D(256, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(c4)
p4 = tf.keras.layers.MaxPooling2D(pool_size=(2, 2))(c4)
c5 = tf.keras.layers.Conv2D(512, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(p4)
c5 = tf.keras.layers.Dropout(0.5)(c5)
c5 = tf.keras.layers.Conv2D(512, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(c5)
c5 = tf.keras.layers.Dropout(0.5)(c5)
u6 = tf.keras.layers.Conv2DTranspose(256, (2, 2), strides=(2, 2), padding='same')(c5)
c6 = tf.keras.layers.Conv2D(256, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(u6)
c6 = tf.keras.layers.Dropout(0.5)(c6)
c6 = tf.keras.layers.Conv2D(512, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(c6)
u7 = tf.keras.layers.Conv2DTranspose(128, (2, 2), strides=(2, 2), padding='same')(c6)
c7 = tf.keras.layers.Conv2D(128, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(u7)
c7 = tf.keras.layers.Dropout(0.5)(c7)
c7 = tf.keras.layers.Conv2D(128, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(c7)
u8 = tf.keras.layers.Conv2DTranspose(64, (2, 2), strides=(2, 2), padding='same')(c7)
u8 = tf.keras.layers.concatenate([u8, c2])
c8 = tf.keras.layers.Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(u8)
c8 = tf.keras.layers.Dropout(0.5)(c8)
c8 = tf.keras.layers.Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(c8)
u9 = tf.keras.layers.Conv2DTranspose(32, (2, 2), strides=(2, 2), padding='same')(c8)
u9 = tf.keras.layers.concatenate([u9, c1], axis=3)
c9 = tf.keras.layers.Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(u9)
c9 = tf.keras.layers.Dropout(0.5)(c9)
c9 = tf.keras.layers.Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_normal', padding='same')(c9)
# In[13]:
outputs = tf.keras.layers.Conv2D(3, (1,1), activation='sigmoid')(c9)
# -
model = tf.keras.Model(inputs=[inputs], outputs=[outputs])
opt = keras.optimizers.Adam(learning_rate=0.1)
model.compile(optimizer=opt, loss=tf.keras.losses.BinaryCrossentropy(), metrics=[DC, iou_coef, 'acc', precision, recall])
model.summary()
# +
# model checkpoint
checkpointer = tf.keras.callbacks.ModelCheckpoint(filepath='/home/kiara/Desktop/CMR_Metric_Calculator/FCN_10000.h5', verbose = 2, save_weights_only = True)
callbacks = [
tf.keras.callbacks.EarlyStopping(patience = 50, monitor = 'val_loss'),
tf.keras.callbacks.TensorBoard(log_dir = 'Log_FCN_10000')
]
# FIT MODEL
results = model.fit(X_train, Y_train, validation_split = 0.1, batch_size = 4, epochs = 150, callbacks=callbacks)
model.save('model_FCN_10000')
model.save('model_FCN_10000.h5')
# -
model.evaluate(X_test, Y_test, verbose=1)
| RV_Segmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
from helper_func import *
# We start by using the ```get_data()``` function from the ```helper_func``` module, which uses the [pandas-datareader package](https://pandas-datareader.readthedocs.io/en/latest/) to load daily stock data and then plot the **closing prices**.
# +
ticker = 'race'
time_window = 1000
stock = get_data(ticker, time_window)
plt.figure(figsize=(15,5))
stock.Close.plot(lw=1)
plt.title(f'{ticker.upper()}')
plt.show()
# -
# Let's see what happens if we try to use **clustering** directly on the observed data *without* any smoothing applied. We use the ```plot_trend()``` function to plot the data.
#
# First, we evaluate the [logarithmic return](https://en.wikipedia.org/wiki/Rate_of_return#Comparing_ordinary_return_with_logarithmic_return) (```lr```), which has some useful properties; we can do it at once with [```np.diff```](https://numpy.org/doc/stable/reference/generated/numpy.diff.html).
#
# We then fit the k-means clustering algorithm, feeding ```lr``` as training instances. We also need to reshape the data, as this expects a 2D array.
# +
c = stock.Close.values
lr = np.diff(np.log(c))
km_obs = KMeans(n_clusters=3).fit(lr.reshape(-1,1))
# +
xr = np.arange(len(lr))
y = c[1:] # because len(c) = len(lr) + 1 (due to subtraction)
labels = km_obs.labels_
plot_trend(xr, y, labels, ticker)
# -
# Not a great help, without smoothing the time series: rough data lead to rough log-returns, which makes it very difficult for a clustering algorithm to separate the data.
#
# We now apply a smoothing technique. Many of these exist; we will consider a **forward-backward EMA** (Exponential Moving Average). What does this mean? Let's break it down:
# - *EMA* gives more weight to recent prices, as a result of which it can better capture the underlying trend in a faster way; with EMA old data points never leave the average, retaining a multiplier (albeit declining to zero) even if they are outside of the chosen time span. However, to *classify* trends well, the lag of regular moving averages is too significant.
# - *Forward-backward filtering* is a heavily used method in digital signal processing, which consists of applying a filter forward in time, then apply it once again backward. We apply a filter twice, hence the result is a two-passes filter, which results in an even smoother output -- more precisely, the filter [amplitude response is squared](https://ccrma.stanford.edu/~jos/fp/Forward_Backward_Filtering.html).
#
# It is important to understand that we are **not predicting anything** here: we just need indexes that tell us something about our data, so we can separate it -- and analyse it further.
# +
cs = stock.Close
# forward and forward-backward EMAs
f_ema = cs.ewm(span=20).mean()
fb_ema = f_ema[::-1].ewm(span=20).mean()[::-1]
plt.figure(figsize=(15,5))
plt.plot(stock.Close.values, c='tab:blue', alpha=0.8)
plt.plot(fb_ema.values, c='k', lw=1)
plt.show()
# -
# This looks pretty **smooth** and with **no lag**. We can use the ```get_trend()``` function, which simply puts all the pieces together.
c = stock.Close.values
x, EMA, labels = get_trend(c)
yy = c[1:]
plot_trend(x, EMA, labels, ticker)
plot_trend(x, yy, labels, ticker)
# It looks like it worked pretty well! From here, one could separate and analyse *by label*.
# +
up = len(labels[labels == 2])
down = len(labels[labels == 0])
still = len(labels[labels == 1])
days = up + down + still
up_perc = 100 * up / days
down_perc = 100 * down / days
still_perc = 100 - up_perc - down_perc
print(f'The {ticker.upper()} stock has been on an up-trend for {round(up_perc,1)}% of the time, on an down-trend for {round(down_perc,1)}% of the time')
# -
# We can finally put all the relevant images together, also relabelling the points to make them fall into three categories: *bullish*, *bearish* and *steady*.
# +
colors = ['tab:red','tab:orange','tab:green']
nclass = 3
freq_tic = int(len(stock.Close.values) / 15)
first_day, last_day = stock.Close.index.strftime('%Y-%m-%d').values[0], stock.Close.index.strftime('%Y-%m-%d').values[-1]
with plt.style.context('seaborn-white'):
# Figure
fig, (ax1, ax2) = plt.subplots(2, sharex=True, sharey=True,
gridspec_kw = {'hspace':0.035}, figsize=(15,12))
# Upper figure
f_ema = cs.ewm(span=20).mean()
fb_ema = f_ema[::-1].ewm(span=20).mean()[::-1]
ax1.plot(stock.Close.values, c='tab:blue', alpha=1, label='Closing price')
ax1.plot(fb_ema.values, c='k', alpha=0.9, lw=1.3, label='Forward-backward EMA')
ax1.legend(frameon=False)
ax1.grid(color='gray', linestyle='-', linewidth=0.2)
[s.set_visible(False) for s in ax1.spines.values()]
# Lower figure
for i in range(nclass):
xx = x[labels == i]
yy = y[labels == i]
ax2.scatter(xx, yy, c=colors[i], s=6, label=i)
handles, new_labels = ax2.get_legend_handles_labels()
new_labels = [f'Bearish ({int(round(down_perc,0))}% of the time)', 'Steady', f'Bullish ({int(round(up_perc,0))}% of the time)']
ax2.legend(handles, new_labels, frameon=False)
ax2.grid(color='gray', linestyle='-', linewidth=0.2)
[s.set_visible(False) for s in ax2.spines.values()]
ax1.set_title(f'{ticker.upper()} stock price (from {first_day} to {last_day})', fontsize='xx-large', weight='bold')
ax1.set_xlim(0, len(stock.Close.values))
ax2.set_xticks(range(0, len(stock.Close.values), freq_tic))
ax2.set_xticklabels(stock.Close.index.strftime('%Y-%m-%d').values[::freq_tic], rotation = 45)
plt.savefig(f'{ticker.upper()}_combined.png', dpi=300, bbox_inches='tight', facecolor='white')
plt.show()
| Historic_data_by_trend.ipynb |
# # Automatic generation of Notebook using PyCropML
# This notebook implements a crop model.
# +
import numpy as np
from copy import copy
from math import *
def diffusionlimitedevaporation(deficitOnTopLayers=5341.0,
soilDiffusionConstant=4.2):
"""
DiffusionLimitedEvaporation Model
Author: <NAME>
Reference: Modelling energy balance in the wheat crop model SiriusQuality2:
Evapotranspiration and canopy and soil temperature calculations
Institution: INRA Montpellier
Abstract: the evaporation from the diffusion limited soil
"""
if (deficitOnTopLayers / 1000.0 <= 0):
diffusionLimitedEvaporation = 8.3 * 1000
else:
if (deficitOnTopLayers / 1000 < 25):
diffusionLimitedEvaporation = (2 * soilDiffusionConstant * soilDiffusionConstant / (deficitOnTopLayers / 1000.0)) * 1000.0
else:
diffusionLimitedEvaporation = 0
return diffusionLimitedEvaporation
# -
# ## Run the model with a set of parameters.
# Each run will be defined in its own cell.
params= diffusionlimitedevaporation(
deficitOnTopLayers = 5341,
soilDiffusionConstant = 4.2,
)
print('diffusionLimitedEvaporation_estimated =')
print(round(params, 3))
# diffusionLimitedEvaporation_computed = 6605.505
| doc/ipynb/test3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pylab inline
from bokeh.plotting import *
from bokeh.layouts import *
from bokeh.models import *
from bokeh.models.widgets import *
from bokeh.application.handlers import FunctionHandler
from bokeh.application import Application
from bokeh.io import show
from bokeh.io import output_notebook
output_notebook()
@show
@Application
@FunctionHandler
def my_graph(doc):
x = np.array([x*0.005 for x in range(0, 200)])
y = x*1.0
source = ColumnDataSource(data=dict(x=x, y=y))
plot = Figure(plot_width=400, plot_height=400)
plot.line('x', 'y', source=source, line_width=3, line_alpha=0.6)
def callback(attr, old, new):
data = source.data
f = slider.value
x, y = data['x'], data['y']
y[:] = x**f
source.data = data #trigger('change', old=data, new=data)
slider = Slider(start=0.1, end=4, value=1, step=.1, title="power")
slider.on_change('value', callback)
doc.add_root(row(plot, slider))
| Interactive graphs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Plotting Velocity Fields on Horizontal Planes (Depth Slices)
#
# This notebook contains discussion, examples, and best practices for plotting velocity field results from NEMO. It extends the discussion of horizontal plane visualizations in the [Plotting Bathymetry Colour Meshes.ipynb](https://nbviewer.jupyter.org/github/SalishSeaCast/tools/blob/master/analysis_tools/Plotting%20Bathymetry%20Colour%20Meshes) and [Plotting Tracers on Horizontal Planes.ipynb](https://nbviewer.jupyter.org/github/SalishSeaCast/tools/blob/master/analysis_tools/Plotting%20Tracers%20on%20Horizontal%20Planes.ipynb) notebooks with plotting of quiver and streamline plots in addition to colour mesh plots.
# Topics include:
#
# * Reading velocity component values from NEMO `*grid_[UVW].nc` results files
# * Plotting colour meshes of velocity components
# * "Un-staggering" velocity component values for vector plots
# * Quiver plots of velocity vectors
# * Streamline plots of velocity fields
# We'll start with our imports, and activation of the Matplotlib inline backend:
# +
import matplotlib.pyplot as plt
import netCDF4 as nc
import numpy as np
from IPython.display import display, Math, Latex
from salishsea_tools import (
nc_tools,
viz_tools,
)
# -
# %matplotlib inline
# ## NEMO `grid_U`, `grid_V`, and `grid_W` Results Files
#
# In NEMO,
# velocity component values are stored at the u, v, and w grid point locations of an
# [Akawara C grid](http://clouds.eos.ubc.ca/~phil/numeric/labs/lab7/lab7.pdf#section.7)
# which NEMO calls the U-grid, V-grid, and W-grid,
# respectively.
#
# The results files that contain the velocity component values for a run have names like:
# ```bash
# SalishSea_1h_20021026_20021026_grid_U.nc
# SalishSea_1h_20021026_20021026_grid_V.nc
# SalishSea_1h_20021026_20021026_grid_W.nc
# ```
# The breakdown of the parts of those file names is the same as for the `grid_T` files
# described in the [NEMO grid_T Results Files](https://nbviewer.jupyter.org/github/SalishSeaCast/tools/blob/master/analysis_tools/Plotting%20Tracers%20on%20Horizontal%20Planes.ipynb#nemo-grid_t-results-files) section of the Plotting Tracers on Horizontal Planes notebook.
# We'll use results files from one of the spin-up runs that are stored in
# the `/results/SalishSea/spin-up/2002/` directory.
# Let's load the files into netCDF dataset objects and look at the dimensions of the U-grid dataset:
u_vel = nc.Dataset('/results/SalishSea/spin-up/2002/26oct/SalishSea_1h_20021026_20021026_grid_U.nc')
v_vel = nc.Dataset('/results/SalishSea/spin-up/2002/26oct/SalishSea_1h_20021026_20021026_grid_V.nc')
w_vel = nc.Dataset('/results/SalishSea/spin-up/2002/26oct/SalishSea_1h_20021026_20021026_grid_W.nc')
nc_tools.show_dimensions(u_vel)
# The dimensions of the velocity component datasets are the same as those of the tracers dataset
# (see the [Plotting Tracers on Horizontal Planes.ipynb](https://nbviewer.jupyter.org/github/SalishSeaCast/tools/blob/master/analysis_tools/Plotting%20Tracers%20on%20Horizontal%20Planes.ipynb#nemo-grid_t-results-files) notebook)
# with the exception of the depth dimension,
# which is called `depthu` rather than `deptht`.
# Likewise,
# the depths dimensions of the v and w velocity component datasets
# are `depthv` and `depthw`,
# respectively.
# These are simply nominal depths and so the u,v and t grid layer depths have the same values. But note that the w grid layer depth has different values.
#
# Oddly,
# the lateral dimensions,
# `x` and `y`, of the grid points of the T, U, V, and W which actually are different (not simply nominal),
# lack any such notation differentiation.
# Now let's look at the variables in the datasets:
nc_tools.show_variables(u_vel)
# The first 5 variables correspond to the dataset's dimensions:
#
# * `nav_lon` and `nav_lat` contain the longitudes and latitudes of lateral grid points
# * `depthu` contains the depths of the vertical grid layers
# * `time_counter` contains the model time values at the centre of each of the output intervals
# * `time_counter_bnds` contains the start and end times of each of the intervals in `time_counter`
#
# The other 2 are calculated values from NEMO:
#
# * `vozocrtx` is the u (zonal) velocity component
# * `u_wind_stress` is the u component of the surface wind stress
# In the V-grid dataset we have:
nc_tools.show_variables(v_vel)
# where:
#
# * `vomecrty` is the v (meridonal) velocity component
# * `v_wind_stress` is the v componenent of the surface wind stress
# and in the W-grid dataset we find:
nc_tools.show_variables(w_vel)
# where:
#
# * `vovecrtz` is the w (vertical) velocity component
# ##Plotting Velocity Component Colour Meshes
#
# Velocity component values are scalars,
# so plotting them as colour meshes is really no different than
# plotting temperature and salinity fields
# (see the [Plotting Tracers on Horizontal Planes.ipynb](https://nbviewer.jupyter.org/github/SalishSeaCast/tools/blob/master/analysis_tools/Plotting%20Tracers%20on%20Horizontal%20Planes.ipynb) notebook).
# ##"Un-staggering" u and v Components
#
# The u, v and w velocity components are all stored at different locations on the grid.
#
# 
#
# *Figure 4.1 from the NEMO 3.4 Book, <NAME>, and the NEMO Team, (2012), ISSN 1288-1619*
# Recall that our grid cells are nominally 500 m in the y direction by 440 m in the x direction.
# To obtain an accurate velocity vector value for a grid cell we interpolate the u and v
# component values to the T-grid point in the centre of the grid cell.
# The `salissea_tools.viz_tools.unstagger()` function is provided to do the necessary interploation.
help(viz_tools.unstagger)
# There is a significant scale difference between the lateral and vertical in the model.
# The vertical layer thicknesses vary from 1 m near the surface to 27 m near the bottom,
# and the w component values are typically an order of magnitude or more smaller than
# the u or v components at a given grid points.
# So,
# we don't worry about interpolating the w components to the T-grid point.
# ## Velocity Quiver Plots
#
# Let's alias the u and v velocity component variables,
# the depth layer midpoint variable,
# and the time counter variable,
# to save on keystrokes:
ugrid = u_vel.variables['vozocrtx']
vgrid = v_vel.variables['vomecrty']
zlevels = u_vel.variables['depthu']
timesteps = u_vel.variables['time_counter']
# Quiver plots are plots of velocity vectors
# (or other vector quantities)
# as arrows on a spatial grid.
# To avoid doing calculations on values that aren't going to be displayed
# we start by calculating the slices of the u and v arrays that we want to
# visualize and masking the zero values to apply an approximate land mask.
# Next,
# we unstagger those slices,
# then we plot the vectors,
# set the axes limits,
# and,
# add axes labels and a title.
#
# Note that the x and y slice arrays that we pass to `quiver()`
# have their first element dropped so that they match the shape
# of the un-staggered velocity component arrays.
# Define the slice to plot
t, zlevel = 4, 0
y_slice = np.arange(250, 370)
x_slice = np.arange(200, 320)
# Slice and mask the arrays
ugrid_tzyx = np.ma.masked_values(ugrid[t, zlevel, y_slice, x_slice], 0)
vgrid_tzyx = np.ma.masked_values(vgrid[t, zlevel, y_slice, x_slice], 0)
# "Unstagger" the velocity values by interpolating them to the T-grid points
u_tzyx, v_tzyx = viz_tools.unstagger(ugrid_tzyx, vgrid_tzyx)
# Plot the velocity vectors
fig, ax = plt.subplots(1, 1, figsize=(10, 8))
viz_tools.set_aspect(ax)
ax.quiver(x_slice[1:], y_slice[1:], u_tzyx, v_tzyx)
# Axes limits and grid
ax.set_xlim(x_slice[0], x_slice[-1])
ax.set_ylim(y_slice[0], y_slice[-1])
ax.grid()
# Axes label and title
ax.set_xlabel('x Index')
ax.set_ylabel('y Index')
ax.set_title(u't = {t:.1f}h, depth \u2248 {d:.2f}{z.units}'.format(t=timesteps[t] / 3600, d=zlevels[zlevel], z=zlevels))
# The `salishsea_tools.viz_tools.plot_land_mask()` function
# uses a bathymetry file or dataset to plot the land areas as
# filled polygons:
help(viz_tools.plot_land_mask)
# Let's "down-sample" the velocity field by plotting the vector at every 3rd grid point,
# and plot the land areas in black.
# +
t, zlevel = 4, 0
y_slice = np.arange(250, 370)
x_slice = np.arange(200, 320)
ugrid_tzyx = np.ma.masked_values(ugrid[t, zlevel, y_slice, x_slice], 0)
vgrid_tzyx = np.ma.masked_values(vgrid[t, zlevel, y_slice, x_slice], 0)
u_tzyx, v_tzyx = viz_tools.unstagger(ugrid_tzyx, vgrid_tzyx)
arrow_step = 3
y_slice_a = y_slice[::arrow_step]
x_slice_a = x_slice[::arrow_step]
fig, ax = plt.subplots(1, 1, figsize=(10, 8))
viz_tools.set_aspect(ax)
ax.quiver(x_slice_a[:], y_slice_a[:], u_tzyx[::3,::3], v_tzyx[::3,::3])
viz_tools.plot_land_mask(ax, '/data/dlatorne/MEOPAR/NEMO-forcing/grid/bathy_meter_SalishSea2.nc', xslice=x_slice, yslice=y_slice)
ax.set_xlim(x_slice[0], x_slice[-1])
ax.set_ylim(y_slice[0], y_slice[-1])
ax.grid()
ax.set_xlabel('x Index')
ax.set_ylabel('y Index')
ax.set_title(u't = {t:.1f}h, depth \u2248 {d:.2f}{z.units}'.format(t=timesteps[t] / 3600, d=zlevels[zlevel], z=zlevels))
# -
# Notice that changing the number of vector arrows plotted also
# changed the size of the arrows thanks to `quiver()`'s auto-scaling algorithm.
#
# Also,
# passing the x and y slices that define the section of the domain that we're working in
# to `viz_tools.plot_land_mask()` results in a significantly faster rendering of the plot.
# That is because calling `viz_tools.plot_land_mask()` without the slice arguments means
# that the land mask polygons for the entire domain are calculated and then those outside the
# plot limits are simply discarded.
# Similarly,
# if you are calling `viz_tools.plot_land_mask()` or `viz_tools.plot_coastline()` several times
# in a notebook,
# it is worth loading the bathymetry into a netCDF4 dataset once and passing the dataset object
# instead of its path and file name.
bathy = nc.Dataset('/data/dlatorne/MEOPAR/NEMO-forcing/grid/bathy_meter_SalishSea2.nc')
# Now,
# let's add some colour,
# move the arrows to that they are centred on the T-grid locations,
# and add a scale key for the arrow lengths:
# +
t, zlevel = 4, 0
y_slice = np.arange(250, 370)
x_slice = np.arange(200, 320)
arrow_step = 3
y_slice_a = y_slice[::arrow_step]
x_slice_a = x_slice[::arrow_step]
ugrid_tzyx = np.ma.masked_values(ugrid[t, zlevel, y_slice_a, x_slice_a], 0)
vgrid_tzyx = np.ma.masked_values(vgrid[t, zlevel, y_slice_a, x_slice_a], 0)
u_tzyx, v_tzyx = viz_tools.unstagger(ugrid_tzyx, vgrid_tzyx)
fig, ax = plt.subplots(1, 1, figsize=(10, 8))
viz_tools.set_aspect(ax)
quiver = ax.quiver(x_slice_a[1:], y_slice_a[1:], u_tzyx, v_tzyx, color='blue', pivot='mid')
viz_tools.plot_land_mask(ax, bathy, xslice=x_slice, yslice=y_slice)
ax.set_xlim(x_slice[0], x_slice[-1])
ax.set_ylim(y_slice[0], y_slice[-1])
ax.grid()
ax.set_xlabel('x Index')
ax.set_ylabel('y Index')
ax.set_title(u't = {t:.1f}h, depth \u2248 {d:.2f}{z.units}'.format(t=timesteps[t] / 3600, d=zlevels[zlevel], z=zlevels))
ax.quiverkey(quiver, 252, 302, 1, '1 m/s', coordinates='data', color='white', labelcolor='white')
# -
# `quiver()` has a number of optional arguments that allow you to
# tune the appearance of your plots
# (see the [docs](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.quiver) for details).
# Here we use the velocity magnitudes to apply a colour map to the arrows,
# and adjust the width of the arrow shafts:
# +
t, zlevel = 4, 0
y_slice = np.arange(250, 370)
x_slice = np.arange(200, 320)
arrow_step = 3
y_slice_a = y_slice[::arrow_step]
x_slice_a = x_slice[::arrow_step]
ugrid_tzyx = np.ma.masked_values(ugrid[t, zlevel, y_slice_a, x_slice_a], 0)
vgrid_tzyx = np.ma.masked_values(vgrid[t, zlevel, y_slice_a, x_slice_a], 0)
u_tzyx, v_tzyx = viz_tools.unstagger(ugrid_tzyx, vgrid_tzyx)
speeds = np.sqrt(np.square(u_tzyx) + np.square(v_tzyx))
fig, ax = plt.subplots(1, 1, figsize=(10, 8))
viz_tools.set_aspect(ax)
quiver = ax.quiver(
x_slice_a[1:], y_slice_a[1:], u_tzyx, v_tzyx, speeds,
pivot='mid', cmap='Reds', width=0.005)
viz_tools.plot_land_mask(ax, bathy, xslice=x_slice, yslice=y_slice)
ax.set_xlim(x_slice[0], x_slice[-1])
ax.set_ylim(y_slice[0], y_slice[-1])
ax.grid()
ax.set_xlabel('x Index')
ax.set_ylabel('y Index')
ax.set_title(u't = {t:.1f}h, depth \u2248 {d:.2f}{z.units}'.format(t=timesteps[t] / 3600, d=zlevels[zlevel], z=zlevels))
ax.quiverkey(quiver, 252, 302, 1, '1 m/s', coordinates='data', color='white', labelcolor='white')
# -
# Note the use of `np.square()` in the calculation of the velocity magnitudes:
#
# ```python
# speeds = np.sqrt(np.square(u_tzyx) + np.square(v_tzyx))
# ```
#
# in contrast to using the Python exponentiation operator:
#
# ```python
# speeds = np.sqrt(u_tzyx**2 + v_tzyx**2)
# ```
#
# The `u_tzyx` and `v_tzyx` arrays are NumPy masked arrays that contain `NaN`
# as mask values.
# While raising `NaN` to the power 2 doesn't seem to cause problems,
# calculating the square root of `NaN` results in a warning like:
#
# ```
# -c:3: RuntimeWarning: invalid value encountered in sqrt
# ```
#
# the first time the cell is executed.
# The warning appears only the first time the cell is run because,
# by default,
# Python only shows runtime errors once per interpreter session.
#
# Using `np.square()` and `np.sqrt()` ensures that the `NaN` values
# are handled properly through the entire calculation.
# ## Velocity Streamline Plots
#
# Streamline plots are plots of velocities
# (or other vector quantities)
# as line segments on a spatial grid.
#
# As for quiver plots,
# we want to avoid doing calculations on values that aren't going to be displayed,
# so we calculate the slices of the u and v arrays that we want to
# visualize and mask the zero values to apply an approximate land mask.
#
# We also need to un-stagger the velocity slices,
# and,
# as was the case for `quiver()`,
# the x and y slice arrays that we pass to `streamplot()`
# have their first element dropped so that they match the shape
# of the un-staggered velocity component arrays.
# +
t, zlevel = 4, 0
y_slice = np.arange(250, 370)
x_slice = np.arange(200, 320)
ugrid_tzyx = np.ma.masked_values(ugrid[t, zlevel, y_slice, x_slice], 0)
vgrid_tzyx = np.ma.masked_values(vgrid[t, zlevel, y_slice, x_slice], 0)
u_tzyx, v_tzyx = viz_tools.unstagger(ugrid_tzyx, vgrid_tzyx)
fig, ax = plt.subplots(1, 1, figsize=(10, 8))
viz_tools.set_aspect(ax)
ax.streamplot(x_slice[1:], y_slice[1:], u_tzyx, v_tzyx)
viz_tools.plot_land_mask(ax, bathy, xslice=x_slice, yslice=y_slice)
ax.set_xlim(x_slice[0], x_slice[-1])
ax.set_ylim(y_slice[0], y_slice[-1])
ax.grid()
ax.set_xlabel('x Index')
ax.set_ylabel('y Index')
ax.set_title(u't = {t:.1f}h, depth \u2248 {d:.2f}{z.units}'.format(t=timesteps[t] / 3600, d=zlevels[zlevel], z=zlevels))
# -
# The default operation of `streamplot()` is to divide the plot domain into a 25x25 grid.
# Each cell in that grid can have,
# at most,
# one traversing streamline.
# `streamplot()` takes a `density` argument with a default value of 1
# which linearly scales that grid and thereby controls
# the closeness of the streamlines.
# For different densities in each direction,
# use a `(density_x, density_y)` tuple value for `density`.
# +
t, zlevel = 4, 0
y_slice = np.arange(250, 370)
x_slice = np.arange(200, 320)
ugrid_tzyx = np.ma.masked_values(ugrid[t, zlevel, y_slice, x_slice], 0)
vgrid_tzyx = np.ma.masked_values(vgrid[t, zlevel, y_slice, x_slice], 0)
u_tzyx, v_tzyx = viz_tools.unstagger(ugrid_tzyx, vgrid_tzyx)
fig, axs = plt.subplots(1, 2, figsize=(16, 8))
densities = (0.5, 2)
for ax, density in zip(axs, densities):
viz_tools.set_aspect(ax)
ax.streamplot(
x_slice[1:], y_slice[1:], u_tzyx, v_tzyx,
density=density,
)
viz_tools.plot_land_mask(ax, bathy, xslice=x_slice, yslice=y_slice)
ax.set_xlim(x_slice[0], x_slice[-1])
ax.set_ylim(y_slice[0], y_slice[-1])
ax.grid()
ax.set_xlabel('x Index')
ax.set_ylabel('y Index')
ax.set_title(u't = {t:.1f}h, depth \u2248 {d:.2f}{z.units}'.format(t=timesteps[t] / 3600, d=zlevels[zlevel], z=zlevels))
# -
# `streamplot()` also accepts an optional `linewidth` argument
# to allow the line widths to be varied.
# We can use that to make the width of the streamlines indicate
# the velocity magnitude:
# +
t, zlevel = 4, 0
y_slice = np.arange(250, 370)
x_slice = np.arange(200, 320)
ugrid_tzyx = np.ma.masked_values(ugrid[t, zlevel, y_slice, x_slice], 0)
vgrid_tzyx = np.ma.masked_values(vgrid[t, zlevel, y_slice, x_slice], 0)
u_tzyx, v_tzyx = viz_tools.unstagger(ugrid_tzyx, vgrid_tzyx)
speeds = np.sqrt(np.square(u_tzyx) + np.square(v_tzyx))
max_speed = viz_tools.calc_abs_max(speeds)
fig, ax = plt.subplots(1, 1, figsize=(10, 8))
viz_tools.set_aspect(ax)
ax.streamplot(
x_slice[1:], y_slice[1:], u_tzyx, v_tzyx,
linewidth=7*speeds/max_speed,
)
viz_tools.plot_land_mask(ax, bathy, xslice=x_slice, yslice=y_slice)
ax.set_xlim(x_slice[0], x_slice[-1])
ax.set_ylim(y_slice[0], y_slice[-1])
ax.grid()
ax.set_xlabel('x Index')
ax.set_ylabel('y Index')
ax.set_title(u't = {t:.1f}h, depth \u2248 {d:.2f}{z.units}'.format(t=timesteps[t] / 3600, d=zlevels[zlevel], z=zlevels))
# -
# Another useful `streamplot()` argument is which allows a colour map to be applied
# along the streamline lenghts.
# We can use that to vary the colour intensity as well as the line width
# of the streamlines to indicate the velocity magnitude:
# +
t, zlevel = 4, 0
y_slice = np.arange(250, 370)
x_slice = np.arange(200, 320)
ugrid_tzyx = np.ma.masked_values(ugrid[t, zlevel, y_slice, x_slice], 0)
vgrid_tzyx = np.ma.masked_values(vgrid[t, zlevel, y_slice, x_slice], 0)
u_tzyx, v_tzyx = viz_tools.unstagger(ugrid_tzyx, vgrid_tzyx)
speeds = np.sqrt(np.square(u_tzyx) + np.square(v_tzyx))
max_speed = viz_tools.calc_abs_max(speeds)
fig, ax = plt.subplots(1, 1, figsize=(10, 8))
viz_tools.set_aspect(ax)
streams = ax.streamplot(
x_slice[1:], y_slice[1:], u_tzyx, v_tzyx,
linewidth=7*speeds/max_speed,
color=speeds, cmap='Blues',
)
viz_tools.plot_land_mask(ax, bathy, xslice=x_slice, yslice=y_slice)
ax.set_xlim(x_slice[0], x_slice[-1])
ax.set_ylim(y_slice[0], y_slice[-1])
ax.grid()
ax.set_xlabel('x Index')
ax.set_ylabel('y Index')
ax.set_title(u't = {t:.1f}h, depth \u2248 {d:.2f}{z.units}'.format(t=timesteps[t] / 3600, d=zlevels[zlevel], z=zlevels))
# -
# ## Changing the Coordinate System
# The NEMO grid is oriented at 29 degree N/W from the North in order to have it align with the major currents through the Georgia Strait. However, when performing comparisons with other sources we may need to rotate the grid to align it with the North, a more common reference point.
#
# First, we need to unstagger the velocities in order to be rotating them from a single centered point on the grid. Then we can rotate the velocities to have u velocities be E/W and v velocities be N/S. The w velocities will remain the same.
# To do this we can perform our change of coordinate system by using the following equations, where theta is in radians:
display(Math(r'u = xcos(\theta)-ysin(\theta)'))
display(Math(r'v = xsin(\theta)+ycos(\theta)'))
t = nc.Dataset('/results/SalishSea/spin-up/2002/26oct/SalishSea_1h_20021026_20021026_grid_T.nc')
lats = t.variables['nav_lat']
lons = t.variables['nav_lon']
# +
t, zlevel = 4, 0
step = 3
y_slice = np.arange(250, 370)
x_slice = np.arange(200, 320)
lats_slice = lats[250:370, 200:320]
lons_slice = lons[250:370, 200:320]
theta=29
theta_rad=29 * np.pi / 180
ugrid_tzyx = np.ma.masked_values(ugrid[t, zlevel, y_slice, x_slice], 0)
vgrid_tzyx = np.ma.masked_values(vgrid[t, zlevel, y_slice, x_slice], 0)
u_tzyx, v_tzyx = viz_tools.unstagger(ugrid_tzyx, vgrid_tzyx)
u_E=u_tzyx * np.cos(theta_rad) - v_tzyx * np.sin(theta_rad)
v_N=u_tzyx * np.sin(theta_rad) + v_tzyx * np.cos(theta_rad)
fig, ax = plt.subplots(1, 1, figsize=(10, 8))
viz_tools.set_aspect(ax)
ax.quiver(lons_slice[1::3, 1::3], lats_slice[1::3, 1::3], u_E[::3,::3], v_N[::3,::3])
viz_tools.plot_land_mask(ax, bathy, coords='map', color='k')
ax.set_xlim([-123.5, -122.6])
ax.set_ylim([48.2, 49.0])
#ax.grid()
ax.set_xlabel('Longitude')
ax.set_ylabel('Latitude')
ax.set_title(u't = {t:.1f}h, depth \u2248 {d:.2f}{z.units}'.format(t=timesteps[t] / 3600, d=zlevels[zlevel], z=zlevels))
# -
| analysis_tools/Plotting Velocity Fields on Horizontal Planes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
d = {'k1': 1, 'k2': 2, 'k3': 3}
d.clear()
print(d)
# +
d = {'k1': 1, 'k2': 2, 'k3': 3}
removed_value = d.pop('k1')
print(d)
# -
print(removed_value)
# +
d = {'k1': 1, 'k2': 2, 'k3': 3}
# removed_value = d.pop('k4')
# print(d)
# KeyError: 'k4'
# +
d = {'k1': 1, 'k2': 2, 'k3': 3}
removed_value = d.pop('k4', None)
print(d)
# -
print(removed_value)
# +
d = {'k1': 1, 'k2': 2}
k, v = d.popitem()
print(k)
print(v)
print(d)
# -
k, v = d.popitem()
print(k)
print(v)
print(d)
# +
# k, v = d.popitem()
# KeyError: 'popitem(): dictionary is empty'
# +
d = {'k1': 1, 'k2': 2, 'k3': 3}
del d['k2']
print(d)
# +
d = {'k1': 1, 'k2': 2, 'k3': 3}
del d['k1'], d['k3']
print(d)
# +
d = {'k1': 1, 'k2': 2, 'k3': 3}
# del d['k4']
# print(d)
# KeyError: 'k4'
| notebook/dict_clear_pop_popitem_del.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from yahoofinancials import YahooFinancials
import numpy as np
import pandas as pd
import pickle
nifty_200 = pd.read_csv('database\ind_nifty200list.csv')
# +
start_date = '2014-01-01'
end_date = '2020-09-30'
price_df = pd.DataFrame()
for stock in nifty_200.Symbol:
print("fetching data for "+stock)
try:
current_price_data = pd.DataFrame(YahooFinancials(stock+'.NS').get_historical_price_data(start_date, end_date, 'monthly')[stock+'.NS']['prices'])
current_price_data['MarketCap'] = current_price_data['adjclose'] * YahooFinancials(stock+'.NS').get_num_shares_outstanding()
current_price_data['Symbol'] = stock
price_df = price_df.append(current_price_data)
except:
print("data not available for "+stock)
# -
price_df.to_pickle('database\\nifty_200_stocks_price_data_24Oct20.pickle')
price_df = pd.read_pickle('database\\nifty_200_stocks_price_data_24Oct20.pickle')
master_df = price_df.copy()
master_df['GrossProfit'] = 0
master_df['BookValue'] = 0
for stock in nifty_200.Symbol:
print("fetching data for "+stock)
try:
inc_stmt = YahooFinancials(stock+'.NS').get_financial_stmts('annual', 'income')['incomeStatementHistory'][stock+'.NS']
balance_sheet = YahooFinancials(stock+'.NS').get_financial_stmts('annual', 'balance')['balanceSheetHistory'][stock+'.NS']
for i in range(len(inc_stmt)):
current_date = list(inc_stmt[i].keys())[0]
gross_profit = inc_stmt[i][current_date]['grossProfit']
book_value = balance_sheet[i][current_date]['commonStock']
if i > 0:
previous_date = list(inc_stmt[i-1].keys())[0]
master_df.loc[(master_df.formatted_date>=current_date) & (master_df['Symbol'] == stock) & (master_df['formatted_date'] < previous_date),['GrossProfit']] = gross_profit
master_df.loc[(master_df.formatted_date>=current_date) & (master_df['Symbol'] == stock) & (master_df['formatted_date'] < previous_date) ,['BookValue']] = book_value
else:
master_df.loc[(master_df.formatted_date>=current_date) & (master_df['Symbol'] == stock),['GrossProfit']] = gross_profit
master_df.loc[(master_df.formatted_date>=current_date) & (master_df['Symbol'] == stock) ,['BookValue']] = book_value
except:
print("data not available for "+stock)
master_df.to_pickle('database\master_df_data_24Oct20.pickle')
| MarketCapDataHistory.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # LeNet on Cifar
#
# This is LeNet (6c-16c-120-84) on Cifar10. Adam algorithm (lr=0.001) with 100 epoches.
#
#
# #### LeNet
#
# Total params: 44,426
# Trainable params: 44,426
# Non-trainable params: 0
#
#
# #### LeNet with 10 intrinsic dim
#
# Total params: 682,076
# Trainable params: 10
# Non-trainable params: 682,066
#
# #### LeNet with 15000 intrinsic dim
# Total params: 930,167,006
# Trainable params: 15,000
# Non-trainable params: 930,152,006
import os, sys
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.pyplot import *
# %matplotlib inline
def extract_num(lines0):
valid_loss_str = lines0[-5]
valid_accuracy_str = lines0[-6]
train_loss_str = lines0[-8]
train_accuracy_str = lines0[-9]
run_time_str = lines0[-10]
valid_loss = float(valid_loss_str.split( )[-1])
valid_accuracy = float(valid_accuracy_str.split( )[-1])
train_loss = float(train_loss_str.split( )[-1])
train_accuracy = float(train_accuracy_str.split( )[-1])
run_time = float(run_time_str.split( )[-1])
return valid_loss, valid_accuracy, train_loss, train_accuracy, run_time
# +
results_dir = '../results/lenet_cifar_l2_0.001'
# dim = [0,250,500,750,1000,1250,1500,1750,1900,1950,2000,2050,2100,2250,2500,3000,4000,5000,5250,5500,5750,6000,6250,6500,6750,7000,7250,7500,7750,8000,8250,8500,8750,9000,9250,9500,9750,10000,15000,20000,25000,30000,35000,40000,45000,50000]
dim = [0,10,50,100,500,750,1000,1250,1500,1750,1900,1950,2000,2050,2100,2250,2500,5000,10000,15000]
i = 0
# filename list of diary
diary_names = []
for subdir, dirs, files in os.walk(results_dir):
for file in files:
if file == 'diary':
fname = os.path.join(subdir, file)
diary_names.append(fname)
dim_ = []
for f in diary_names:
dim_.append( int(f.split('_')[-2]) )
dim = sorted(dim_)
print dim
diary_names_ordered = []
for d in dim:
for f in diary_names:
if d == int(f.split('_')[-2]):
# print "%d is in" % d + f
diary_names_ordered.append(f)
# intrinsic update method
Rs, Acc = [], []
i = 0
for fname in diary_names_ordered:
with open(fname,'r') as ff:
lines0 = ff.readlines()
R = extract_num(lines0)
print "%d dim:\n"%dim[i] + str(R) + "\n"
i += 1
Rs.append(R)
Acc.append(R[1])
Rs = np.array(Rs)
Acc = np.array(Acc)
# 2.2 construct acc_solved_all and dim_solved_all
for id_d in range(len(dim)):
d = dim[id_d]
r = Rs[id_d,1]
if d==0:
test_acc_bl = r
# print "Acc goal is: " + str(test_acc_sl) + " for network with depth " + str(ll) + " width "+ str(w)
else:
test_acc = r
if test_acc>test_acc_bl*0.9:
acc_solved=test_acc
dim_solved=d
print "Intrinsic dim is: " + str(d) + " for LeNet"
# print "\n"
break
# -
print ','.join(['[%i, %s]' % (dim[n], Acc[n]) for n in xrange(len(Acc))])
# ## Performance comparison with Baseline
#
# "Baseline method" indicates optimization in the parameter space.
#
# The proposed method first embeds parameters into the intrinisic space (via orthogonal matrix), and optimization is the new space.
#
# The dimension of intrinsic space indicates the degree of freedom in the weights of neural nets.
# +
fig, ax = subplots(figsize=(6,5) )
font = {'size' : 12}
matplotlib.rc('font', **font)
plot(dim[1:], Rs[1:,1], 'o', mec='b', mfc=(.8,.8,1), ms=10)
plot(dim_solved, acc_solved, 'o', mec='b', mfc='b', ms=10)
axhline(Rs[0,1], ls='-', color='k',label='baseline')
axhline(Rs[0,1] * .9, ls=':', color='k',label='solved')
plt.legend()
ax.set_xlabel('Subspace dim $d$')
ax.set_ylabel('Validation accuracy')
# ax.set_title('width %d, depth %d' %(width[i], depth[j]))
plt.grid()
ax.set_ylim([0.3,0.7])
fig.savefig("figs/lenet_cifar_dim.pdf", bbox_inches='tight')
# -
# The above figure show that updating in the intrinsic space can prevent overfitting.
#
# +
nn = len(Rs)-1
fig, ax = subplots(figsize=(5,4) )
plt.scatter(dim[1:], Rs[1:,0], edgecolor="k", facecolor="w" )
ax.plot(dim[1:], Rs[0,0]*np.ones(nn)/0.9,'r-.', label="Testing: baseline")
ax.set_xlabel('Intrinsic Dim')
ax.set_ylabel('Accuracy')
plt.grid()
# ax.set_ylim([-0.1,1.1])
# -
# The above figure show that updating in the intrinsic space can prevent overfitting.
# +
results_dir = '../results/lrb_lenet_cifar'
i = 0
# filename list of diary
diary_names = []
for subdir, dirs, files in os.walk(results_dir):
for file in files:
if file == 'diary':
fname = os.path.join(subdir, file)
diary_names.append(fname)
print diary_names
dim_ = []
for f in diary_names:
dim_.append( int(f.split('/')[-2].split('_')[-1]) )
dim = sorted(dim_)
print dim
diary_names_ordered = []
for d in dim:
for f in diary_names:
if d == int(f.split('/')[-2].split('_')[-1]):
# print "%d is in" % d + f
diary_names_ordered.append(f)
# intrinsic update method
Rs, Acc = [], []
i = 0
for fname in diary_names_ordered:
with open(fname,'r') as ff:
lines0 = ff.readlines()
R = extract_num(lines0)
# print "%d dim:\n"%dim[i] + str(R) + "\n"
i += 1
Rs.append(R)
Acc.append(R[1])
Rs = np.array(Rs)
Acc = np.array(Acc)
print Rs.shape
# 2.2 construct acc_solved_all and dim_solved_all
for id_d in range(len(dim)):
d = dim[id_d]
r = Rs[id_d,1]
if d==0:
test_acc_bl = r
# print "Acc goal is: " + str(test_acc_sl) + " for network with depth " + str(ll) + " width "+ str(w)
else:
test_acc = r
if test_acc>test_acc_bl*0.9:
acc_solved=test_acc
dim_solved=d
print "Intrinsic dim is: " + str(d) + " for LeNet"
# print "\n"
break
# -
print ','.join(['[%i, %s]' % (dim[n], Acc[n]) for n in xrange(len(Acc))])
# +
""" Extract final stats from resman's diary file"""
def extract_num(lines0, is_reg=False):
if is_reg:
valid_loss_str = lines0[-5]
valid_accuracy_str = lines0[-6]
train_loss_str = lines0[-8]
train_accuracy_str = lines0[-9]
average_time_str = lines0[-10]
run_time_str = lines0[-11]
else:
valid_loss_str = lines0[-6]
valid_accuracy_str = lines0[-7]
train_loss_str = lines0[-10]
train_accuracy_str = lines0[-11]
average_time_str = lines0[-12]
run_time_str = lines0[-13]
valid_loss = float(valid_loss_str.split( )[-1])
valid_accuracy = float(valid_accuracy_str.split( )[-1])
train_loss = float(train_loss_str.split( )[-1])
train_accuracy = float(train_accuracy_str.split( )[-1])
run_time = float(run_time_str.split( )[-1])
return valid_loss, valid_accuracy, train_loss, train_accuracy, run_time
""" Extract number of total parameters for each net config from resman's diary file"""
def parse_num_params(lines0):
line_str = ''.join(lines0)
idx = line_str.find("Total params")
param_str = line_str[idx+14:idx+14+20] # 14 is the length of string "Total params: "
param_num = param_str.split("\n")[0]
return int(locale.atof(param_num))
def extract_perf_dim(results_dir):
# Dim: subspace dim
# Acc: Accuracy
# filename list of diary
diary_names = []
for subdir, dirs, files in os.walk(results_dir):
for f in files:
if f == 'diary':
fname = os.path.join(subdir, f)
diary_names.append(fname)
# print diary_names
dim = []
for f in diary_names:
# print f
tmp_str = f.split('/')[-2]
if tmp_str.split('_')[-3]=='LeNet':
d = int(tmp_str.split('_')[-2])
dim.append(d)
dim = list(set(dim))
dim = sorted(dim)
# print dim
diary_names_ordered = []
for d in dim:
for f in diary_names:
if '_LeNet_'+str(d)+'_' in f and f.split('_')[-3]=='LeNet':
# print "%d is in" % d + f
diary_names_ordered.append(f)
# print diary_names_ordered
# intrinsic update method
Dim= []
Acc = []
for fname in diary_names_ordered:
tmp_str = fname.split('/')[-2]
d = int(tmp_str.split('_')[-2])
with open(fname,'r') as ff:
lines0 = ff.readlines()
try:
r = extract_num(lines0,False)[1]
# print "%d dim:\n"% d + str(r) + "\n"
Dim.append(d)
Acc.append(r)
except ValueError:
print "%d dim:\n"%d + "Error \n"
pass
return Dim, Acc
# -
def plot_perf_dim(Dim, Acc, Cx=None):
fig, ax = subplots(figsize=(6,5) )
font = {'size' : 12}
matplotlib.rc('font', **font)
for i in range(len(Dim)):
if Cx == None:
if Acc[i]>Acc[0]*0.9 and i>10:
print "d_{int}="+str(Dim[i]) + ', acc:' + str(Acc[i])
break
else:
if Acc[i]>Cx and i>10:
print "d_{int}="+str(Dim[i]) + ', acc:' + str(Acc[i])
break
plot(Dim[4:], Acc[4:], 'o', mec='b', mfc=(.8,.8,1), ms=10)
plot(Dim[i], Acc[i], 'o', mec='b', mfc='b', ms=10)
axhline(Acc[0], ls='-', color='k',label='baseline')
axhline(Acc[0] * .9, ls=':', color='k',label='solved')
plt.legend()
ax.set_xlabel('Subspace dimension $d$')
ax.set_ylabel('Validation accuracy')
# ax.set_title('Cifar: Untied_LeNet' )
plt.grid()
ax.set_ylim([0.0,1.01])
# fig.savefig("figs/fnn_mnistPL_W"+str(width[i])+"_L"+str(depth[j])+".pdf", bbox_inches='tight')
# +
results_root = '/home/users/chunyuan.li/public_results/chun/'
exp_folder = 'results_cifar_LeNet'
results_dir = results_root + exp_folder
Dim, Acc= extract_perf_dim(results_dir)
# print Dim, Acc
plot_perf_dim(Dim, Acc, 0.5)
# -
print ','.join(['[%i, %s]' % (Dim[n], Acc[n]) for n in xrange(len(Dim))])
| intrinsic_dim/plots/more/lenet_cifar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import tensorflow as tf
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import sys
sys.path.insert(0, '/home/ubuntu/roger/Machine_Learning_Immunogenicity/src')
import onehot
from gzip import GzipFile
import copy
# %pwd
tcell_all=pd.read_table("/home/roger/other/Machine_Learning_Immunogenicity/data/tcell.txt.gz",compression='gzip')
tcell_all.shape
tcell_comp=pd.read_table('/home/roger/other/Machine_Learning_Immunogenicity/data/tcell_peptide_allele_nodups.txt.gz',compression='gzip')
print tcell_comp.shape
tcell_comp.head(10)
map(len,tcell_comp['Description'])
print np.histogram(map(len,tcell_comp['Description']),bins=30)
plt.hist(np.histogram(map(len,tcell_comp['Description']),bins=30))
plt.show()
print np.sum(np.asarray(map(len,tcell_comp['Description'])) <= 20)
# +
tcell_clean1=tcell_comp[tcell_comp['Allele Name'].notnull()]
tcell_clean2=tcell_clean1[tcell_clean1['Description'].notnull()]
tcell_clean=tcell_clean2[tcell_clean2['bin'].notnull()]
print tcell_clean.shape
tcell_clean
# -
print np.sum(tcell_clean['bin'])
tcell_clean['Allel']=pd.Categorical.from_array(tcell_clean['Allele Name']).codes
tcell_clean.head(10)
#plt.hist(np.histogram(tcell_clean['Allele Name'],bins=50))
#plt.show()
#print np.sum(np.asarray(tcell_all['MHC Allele ID']) < 50)
# scaling factors
print tcell_clean.columns
def to_ascii(letter):
if pd.isnull(letter):
return 0
else:
return ord(letter)
print tcell_clean.shape
tcell_clean.index
leng_seq=len(tcell_clean.Description.get(1))
print tcell_clean.Allel.get(0)
print tcell_clean.bin.get(0)
print leng_seq
map(to_ascii,tcell_clean.Description.get(1))[:15]
number_data=tcell_clean.shape[0]
number_feature=51
tcell_data=np.zeros([number_data,number_feature+1])
print number_data
j=0
for i in tcell_clean.index:
tcell_data[j,0]=tcell_clean.Allel.get(i)
tcell_data[j,number_feature]=tcell_clean.bin.get(i)
leng_seq=len(tcell_clean.Description.get(i))
if leng_seq <=number_feature-1:
tcell_data[j,1:leng_seq+1]=map(to_ascii,tcell_clean.Description.get(i))
else:
tcell_data[j,1:number_feature]=map(to_ascii,tcell_clean.Description.get(i))[:number_feature-1]
j=j+1
print tcell_data[3,:]
tcell_clean.head(10)
np.random.shuffle(tcell_data)
tcell_data[:2,:]
#tcell_shuf.dt
# +
tcell_array=np.asarray(tcell_data)
num_total=tcell_array.shape[0]
train_num=30000
valid_num=7000
test_num=num_total-train_num-valid_num
train_set=tcell_array[:train_num,1:number_feature]
#train_lable=tcell_array[:train_num,number_feature]
train_lable=(np.arange(2)==tcell_array[:train_num,number_feature][:,None]).astype(np.float32)
valid_set=tcell_array[train_num:train_num+valid_num,1:number_feature]
#valid_lable=tcell_array[]
valid_lable=(np.arange(2)==tcell_array[train_num:train_num+valid_num,number_feature][:,None]).astype(np.float32)
test_set=tcell_array[train_num+valid_num:,1:number_feature]
#test_lable=tcell_array[train_num+valid_num:,number_feature]
test_lable=(np.arange(2)==tcell_array[train_num+valid_num:,number_feature][:,None]).astype(np.float32)
print train_set.shape, train_lable.shape, valid_set.shape, valid_lable.shape, test_set.shape, test_lable.shape
# -
number_feature1=number_feature-1
batch_size=1000
hidden_size=20
num_lable=2
graph=tf.Graph()
with graph.as_default():
train_set_batch=tf.placeholder(tf.float32,shape=(batch_size,number_feature1))
train_lable_batch=tf.placeholder(tf.float32,shape=(batch_size,num_lable))
valid_set_tf=tf.constant(valid_set.astype(np.float32))
test_set_tf=tf.constant(test_set.astype(np.float32))
weights1=tf.Variable(tf.truncated_normal([number_feature1,hidden_size]))
bias1=tf.Variable(tf.zeros([hidden_size]))
weights2=tf.Variable(tf.truncated_normal([hidden_size,num_lable]))
bias2=tf.Variable(tf.zeros([num_lable]))
logits1=tf.matmul(train_set_batch,weights1)+bias1
act1=tf.nn.relu(logits1)
logits=tf.matmul(act1,weights2)+bias2
loss=tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits,train_lable_batch))
optimizer=tf.train.GradientDescentOptimizer(0.000005).minimize(loss)
train_pred=tf.nn.softmax(logits)
valid_pred=tf.nn.softmax(tf.matmul(tf.nn.relu(tf.matmul(valid_set_tf,weights1)+bias1),weights2)+bias2)
test_pred=tf.nn.softmax(tf.matmul(tf.nn.relu(tf.matmul(test_set_tf,weights1)+bias1),weights2)+bias2)
# +
num_step=10000
#def accuracy(predictions, labels):
# return (100.0 * np.sum(np.argmax(predictions, 1) == np.argmax(labels, 1))
# / predictions.shape[0])
def accuracy(predictions, labels):
pre=np.argmax(predictions, 1.)
lab=np.argmax(labels, 1.)
# print pre[:100]
# print lab[:100]
posi=np.sum(pre*lab).astype(np.float32)/np.sum(lab).astype(np.float32)
neg=np.sum((1.-pre)*(1.-lab)).astype(np.float32)/np.sum(1-lab).astype(np.float32)
return 100.*2.*posi*neg/(posi+neg)
with tf.Session(graph=graph) as session:
tf.initialize_all_variables().run()
print('Initialized')
for i in range(num_step):
offset=i*batch_size % train_num
train_set_feed=train_set[offset:offset+batch_size,:]
train_lable_feed=train_lable[offset:offset+batch_size]
feed_dict={train_set_batch:train_set_feed, train_lable_batch:train_lable_feed}
_,l,pred=session.run([optimizer,loss,train_pred],feed_dict=feed_dict)
if (i % 20)==0:
print("Minibatch loss at step %d: %f" % (i, l))
print("Minibatch accuracy: %.1f%%" % accuracy(pred, train_lable_feed))
print("Validation accuracy: %.1f%%" % accuracy(valid_pred.eval(), valid_lable))
print("Test accuracy: %.1f%%" % accuracy(test_pred.eval(), test_lable))
# -
train_set.shape
train_lable
| explore_without_Allel_tcell.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas
import numpy
import matplotlib.pyplot as plt
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.model_selection import KFold, cross_val_score
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import StandardScaler
import time
import datetime
pandas.set_option('display.max_columns', 200)
kf = KFold(n_splits=5, shuffle=True, random_state=42)
# -
# ## Предобработка данных
train = pandas.read_csv('features.csv', index_col='match_id')
train.describe().head(10)
# Целевой переменной является "radiant_win"
#Посчитаем в каких столбцах cколько пропусков
rows = train.shape[0]
counts = train.count()
counts_nan = counts[counts < rows]
counts_nan.sort_values().apply(lambda c: float((rows - c) * 100 / rows))
# #### Обоснование пропусков:
# * first_blood_player2 (второй игрок, причастный к событию) - первое убийство может быть совершено одним игроком, это достаточно частое явление и происходит в 45% случаев
# * first_blood_time (игровое время первой крови) - в 20% случаев первой крови нет в первые 5 минут игры
# * radiant_flying_courier_time (время приобретения предмета "flying_courier") - в 28% случаев команда не покупает "flying_courier" в первые 5 минут игры
#
# Остальные признаки аналогичны
#Замена пропусков
train = train.fillna(0).astype('int')
# удаление итогов матча
totals = ['duration',
'tower_status_radiant',
'tower_status_dire',
'barracks_status_radiant',
'barracks_status_dire',]
for total in totals:
del train[total]
# запись целевой переменной
y_train = train['radiant_win']
del train['radiant_win']
# удаление категориальных признаков
X_train = train.copy()
del X_train['lobby_type']
for i in range(1, 6):
del X_train[f'r{i}_hero']
del X_train[f'd{i}_hero']
# ## Градиентный бустинг
nums = (10, 20, 30, 80, 130, 230, 330)
scores = []
for n in nums:
print(f"# {n}", end='\t')
model = GradientBoostingClassifier(n_estimators=n, random_state=42)
start_time = datetime.datetime.now()
cvl = cross_val_score(model, X_train, y_train, scoring='roc_auc', cv=kf)
print(f"Time: {datetime.datetime.now() - start_time}")
scores.append(cvl.mean())
plt.plot(nums, scores)
plt.xlabel('n_estimators')
plt.ylabel('score')
plt.show()
# #### Результаты:
# Показатель метрики качества AUC-ROC равен 0.69 для градиентного бустинга с 30 деревьями, по времени это заняло 0:00:47. Увеличение количества деревьев до 200~ дает площадь порядка 0.71, дальше увеличение количества деревьев практически не улучшает результат, он стремится к 0.72
# ##### Для улучшения результатов модели / ускорения обучения можно:
# * Уменьшить размерность, избавившись от линейно-зависимых и ненужных признаков
# * Использовать не всю выборку, а некоторое ее подмножество, деревья малочувствительны к размеру.
# * Уменьшить глубину деревьев, в градиентном бустинге в отличие от случайного леса нам важна ширина.
# * Использовать стохастический градиентный бустинг, в нем случайные подвыборки.
# ## Логистическая регрессия
def best_C(X, Cs):
scores = []
text = []
for C in Cs:
start_time = datetime.datetime.now()
model = LogisticRegression(C=C, random_state=42)
cvs = cross_val_score(model, X, y_train, cv=kf, scoring='roc_auc')
start_time = datetime.datetime.now() - start_time
text.append(f'C={C:.5f}, AUC-ROC={cvs.mean():.4f}, time={start_time}')
scores.append(cvs.mean())
return scores, text
def pl(ans, range_):
plt.plot(range_, ans)
plt.xlabel('lg(C)')
plt.ylabel('score')
plt.show()
# X_train = train этой строкой можно проверить влияют ли на модель категориальные данные
X_train = StandardScaler().fit_transform(X_train)
range_ = range(-5, 6)
C_range = (10**i for i in range_)
'''
Для получения более точного значения С
range_ = range(1, 11)
C_range = (i/1000 for i in range_)
'''
ans = best_C(X_train, C_range)
print(*ans[1], sep='\n')
pl(ans[0], range_)
# ##### Вывод:
# Площадь при С=0.01 примерно 0.716. Наилучшее значение показателя AUC-ROC так же достигается при C = 0.007 и равно 0.717. Так же при проверке модели с категориальными признаками получается такой же результат, значит модель распознает их как шум.
#
# Надо заметить, что метрика AUC-ROC данной модели примерно равна 330 деревьям в градиентном бустинге, но при этом в разы быстрее.
# ##### Мешок слов
def bag_words(all_, X_):
X_hero = numpy.zeros((all_.shape[0], 112))
for i, match_id in enumerate(all_.index):
for p in range(1, 6):
X_hero[i, all_.ix[match_id, f'r{p}_hero']-1] = 1
X_hero[i, all_.ix[match_id, f'd{p}_hero']-1] = -1
X_ = pandas.DataFrame(X_)
X_hero = pandas.DataFrame(X_hero)
return pandas.concat([X_, X_hero], axis=1)
mx = 0
for i in range(1, 6):
mx = max(max(train[f'r{i}_hero']), max(train[f'd{i}_hero']), mx)
print(mx) # количество героев в выборке
# Формируем "мешок слов" по героям
X_train = bag_words(train, X_train)
range_ = range(-5, 6)
C_range = (10**i for i in range_)
'''
Для получения более точного значения С
range_ = range(5, 15)
C_range = (i/10 for i in range_)
'''
ans = best_C(X_train, C_range)
print(*ans[1], sep='\n')
pl(ans[0], range_)
# #### Результаты:
# После добавления "мешка слов" по героям площадь увеличилась. Наилучшее значение показателя AUC-ROC = 0.75 достигается при C = 0.1. Это произошло, потому что вместо отсутствия данных или шума, мы имеем осмысленную разреженную матрицу для построения предсказания.
# ## Предсказание для тестовой выборки:
test = pandas.read_csv('features_test.csv',index_col='match_id')
#Замена пропусков
test = test.fillna(0).astype('int')
# удаление категориальных признаков
X_test = test.copy()
del X_test['lobby_type']
for i in range(1, 6):
del X_test[f'r{i}_hero']
del X_test[f'd{i}_hero']
X_test = StandardScaler().fit_transform(X_test)
X_test = bag_words(test, X_test)
clf = LogisticRegression(C=0.1)
clf.fit(X_train, y_train)
pred = clf.predict_proba(X_test)[:, 1]
print(min(pred), max(pred))
| Dota2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## This demonstrates scikit-learn clustering for comparison with Tribuo clustering
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.cluster import KMeans
from sklearn.cluster import AgglomerativeClustering
from sklearn.cluster import MeanShift
from sklearn.cluster import DBSCAN
from sklearn.metrics import adjusted_mutual_info_score
from sklearn.metrics import normalized_mutual_info_score
# This dataset is generated in the notebook: scikit-learn Clustering - Data Setup
df = pd.read_csv('../../data/gaussianBlobs.csv')
# print(df)
# +
df_X = df.drop(['Cluster'], axis=1)
df_y = pd.DataFrame(df[['Cluster']])
X_train, X_test, y_train, y_test = train_test_split(df_X.values, df_y.values, test_size=0.2, random_state=1)
print('Training data size = %d, number of features = %d' % (len(X_train), len(df_X.columns)))
print('Testing data size = %d, number of features = %d' % (len(X_test), len(df_X.columns)))
# -
def evaluate(actual, predicted):
print('Clustering Evaluation')
print('Normalized MI = %.2f' % normalized_mutual_info_score(actual, predicted))
print('Adjusted MI = %.2f' % adjusted_mutual_info_score(actual, predicted))
km = KMeans(n_clusters=6, max_iter=100, n_jobs=4, random_state=1, init='random')
km_plus_plus = KMeans(n_clusters=6, max_iter=100, n_jobs=4, random_state=1, init='k-means++')
# This crashes the kernel everytime
# ag = AgglomerativeClustering(n_clusters=6)
# This doesn't finish after a reasonable amount of time
# ms = MeanShift(n_jobs=4)
# This also crashes the kernel everytime
# dbscan = DBSCAN(eps=3, min_samples=50, n_jobs=4)
print(km)
print(km_plus_plus)
# print(ag)
# print(ms)
# print(dbscan)
# +
# %time km.fit(X_train)
# run 1
# time: 17.4 s
# run 2
# time: 17.7 s
# run 3
# time: 16.0 s
# +
predicted = km.predict(X_test)
evaluate(y_test.ravel(), predicted)
# run 1
# Normalized MI = 1.00
# Adjusted MI = 1.00
# run 2
# Normalized MI = 1.00
# Adjusted MI = 1.00
# run 3
# Normalized MI = 1.00
# Adjusted MI = 1.00
# -
# +
# %time km_plus_plus.fit(X_train)
# run 1
# time: 19.2 s
# run 2
# time: 18.6 s
# run 3
# time: 17.9 s
# +
predicted = km_plus_plus.predict(X_test)
evaluate(y_test.ravel(), predicted)
# run 1
# Normalized MI = 1.00
# Adjusted MI = 1.00
# run 2
# Normalized MI = 1.00
# Adjusted MI = 1.00
# run 3
# Normalized MI = 1.00
# Adjusted MI = 1.00
# -
# +
# # %time ag.fit(X_train)
# +
# predicted = ag.predict(X_test)
# evaluate(y_test.ravel(), predicted)
# -
# +
# # %time ms.fit(X_train)
# +
# predicted = ms.predict(X_test)
# evaluate(y_test.ravel(), predicted)
# -
# +
# # %time dbscan.fit(X_train)
# +
# predicted = dbscan.predict(X_test)
# evaluate(y_test.ravel(), predicted)
| notebooks/clustering/scikit-learn Clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="iV4zt1ffHnd8"
# # Lab 03 : Vanilla neural networks -- demo
#
# # Creating a two-layer network
# + executionInfo={"elapsed": 5162, "status": "ok", "timestamp": 1633353050528, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14753845709256584186"}, "user_tz": -480} id="lv57ndKsHneB"
import torch
import torch.nn as nn
# + [markdown] id="XN_VbhDiHneC"
# ### In Pytorch, networks are defined as classes
# + executionInfo={"elapsed": 61, "status": "ok", "timestamp": 1633353050538, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14753845709256584186"}, "user_tz": -480} id="glmPTQvVHneD"
class two_layer_net(nn.Module):
def __init__(self, input_size, hidden_size, output_size):
super(two_layer_net , self).__init__()
self.layer1 = nn.Linear( input_size, hidden_size , bias=True)
self.layer2 = nn.Linear( hidden_size, output_size , bias=True)
def forward(self, x):
x = self.layer1(x)
x = torch.relu(x)
x = self.layer2(x)
p = torch.softmax(x, dim=0)
return p
# + [markdown] id="Yh09fuIeHneD"
# ### Create an instance that takes input of size 2, then transform it into something of size 5, then into something of size 3
# $$
# \begin{bmatrix}
# \times \\ \times
# \end{bmatrix}
# \longrightarrow
# \begin{bmatrix}
# \times \\ \times \\ \times \\ \times \\ \times
# \end{bmatrix}
# \longrightarrow
# \begin{bmatrix}
# \times \\ \times \\ \times
# \end{bmatrix}
# $$
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 60, "status": "ok", "timestamp": 1633353050538, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14753845709256584186"}, "user_tz": -480} id="APeqMsEiHneE" outputId="9afebe73-91ab-438b-e3e1-530b1067bca0"
net= two_layer_net(2,5,3)
print(net)
# + [markdown] id="OB_pI0sLHneF"
# ### Now we are going to make an input vector and feed it to the network:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 51, "status": "ok", "timestamp": 1633353050539, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14753845709256584186"}, "user_tz": -480} id="2UQRDBR8HneG" outputId="2d9bc723-dfe6-4903-b16e-25b36bdd58b0"
x=torch.Tensor([1,1])
print(x)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 48, "status": "ok", "timestamp": 1633353050540, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14753845709256584186"}, "user_tz": -480} id="NARI8kn7HneH" outputId="ab77cc29-e6e5-426f-e414-3b7cae6067fd"
p=net.forward(x)
print(p)
# + [markdown] id="Tx9OTxW8HneI"
# ### Syntactic easy for the forward method
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 45, "status": "ok", "timestamp": 1633353050541, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14753845709256584186"}, "user_tz": -480} id="p3Zn9pxYHneJ" outputId="3e2790a7-9974-4b21-d70e-7819b47e1ed8"
p=net(x)
print(p)
# + [markdown] id="9O66AvqUHneJ"
# ### Let's check that the probability vector indeed sum to 1:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 41, "status": "ok", "timestamp": 1633353050541, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14753845709256584186"}, "user_tz": -480} id="9OWPVuc1HneK" outputId="62e990d3-1f45-475f-dc76-aedcec9400cd"
print( p.sum() )
# + [markdown] id="hxEOJF_tHneK"
# ### This network is composed of two Linear modules that we have called layer1 and layer2. We can see this when we type:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 38, "status": "ok", "timestamp": 1633353050541, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14753845709256584186"}, "user_tz": -480} id="LUWDzf66HneL" outputId="4d7138b5-7e9e-452c-9007-1959267c4a20"
print(net)
# + [markdown] id="qRqBlEx2HneL"
# ### We can access the first module as follow:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 37, "status": "ok", "timestamp": 1633353050543, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14753845709256584186"}, "user_tz": -480} id="K5wg7D3HHneL" outputId="aa47e85d-b1f1-452a-cc23-f34b1a1cfa9a"
print(net.layer1)
# + [markdown] id="u3enZFg0HneM"
# ### To get the weights and bias of the first layer we do:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 35, "status": "ok", "timestamp": 1633353050544, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14753845709256584186"}, "user_tz": -480} id="iR3R8c8jHneM" outputId="742ff58b-ca68-4b0f-a41d-289ad1ef457c"
print(net.layer1.weight)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 34, "status": "ok", "timestamp": 1633353050546, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14753845709256584186"}, "user_tz": -480} id="fs15qz6yHneM" outputId="165d176b-4db1-4663-e051-ed0d1b707a94"
print(net.layer1.bias)
# + [markdown] id="PBNpOnslHneN"
# ### So to change the first row of the weights from layer 1 you would do:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 31, "status": "ok", "timestamp": 1633353050546, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14753845709256584186"}, "user_tz": -480} id="uP74F-69HneN" outputId="5cc1f6ce-f2e5-401f-ce20-2fe31c64150b"
with torch.no_grad():
net.layer1.weight[0,0]=10
net.layer1.weight[0,1]=20
print(net.layer1.weight)
# + [markdown] id="8bEuzw90HneN"
# ### Now we are going to feed $x=\begin{bmatrix}1\\1 \end{bmatrix}$ to this modified network:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 29, "status": "ok", "timestamp": 1633353050546, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14753845709256584186"}, "user_tz": -480} id="4hdc7arcHneO" outputId="184ffab9-a469-4c1d-aae3-04cb91d2d1e1"
p=net(x)
print(p)
# + [markdown] id="F8FEWMGcHneO"
# ### Alternatively, all the parameters of the network can be accessed by net.parameters().
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 29, "status": "ok", "timestamp": 1633353050549, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14753845709256584186"}, "user_tz": -480} id="btOOjpM-HneP" outputId="a484427e-6e60-4bcc-b17a-ed712be8d733"
list_of_param = list( net.parameters() )
print(list_of_param)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 474, "status": "ok", "timestamp": 1633353091968, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14753845709256584186"}, "user_tz": -480} id="ZDD9ICyBHneP" outputId="a16d70ad-d2ee-46d7-851c-6f25c2560f09"
print(net.parameters())
# + executionInfo={"elapsed": 26, "status": "ok", "timestamp": 1633353050552, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14753845709256584186"}, "user_tz": -480} id="YdD2JzunbR15"
| codes/labs_lecture03/lab03_vanilla_nn/vanilla_nn_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
'<NAME>'
"<NAME>"
'Este texto incluye unas " " '
"Este 'palabra' se encuentra escrita entre comillas simples"
"Esta \"palabra\" se encuentra escrita entre comillas dobles"
'Esta \'palabra\' se encuentra escrita entre comillas dobles'
"Una cadena"
'otra cadena'
'otra cadena más'
print("Una cadena")
print('otra cadena')
print('otra cadena más')
print("Un texto\tuna tabulación")
print("Un texto\nuna nueva línea")
print("C:\nombre\directorio")
print(r"C:\nombre\directorio")
print("""Una línea
otra línea
otra línea\tuna tabulación""")
c = "Esto es una cadena\ncon dos líneas"
c
print(c)
c + c
print(c + c)
s = c + " " + c
print(s)
s = "Una cadena " "compuesta de dos cadenas"
s
print(s)
c1 = "Una cadena"
c2 = "otra cadena"
print(c1 c2)
print(c1 + c2)
c1 = "Una cadena"
c2 = "otra cadena"
print(c1 + c2)
palabra = "Python"
print(palabra[0]) # Carácter en la posición 0
print(palabra[3]) # Carácter en la posición 3
palabra[-1]
palabra[-2]
palabra[-6]
palabra[0:2]
palabra[0:-1]
palabra[2:-1]
palabra[2:]
palabra[:2]
palabra[:]
palabra = "N" + palabra[1:]
palabra
| Fase 1 - Fundamentos de programacion/Tema 01 - Introduccion informal/Leccion 2 - Textos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] hideOutput=false hidePrompt=false slideshow={"slide_type": "-"}
# # Ecommerce Purchases Exercise - Week 1
# # <NAME> - ML01
#
# #### Import pandas and read in the Ecommerce Purchases csv file and set it to a DataFrame called ecom.
# + hidePrompt=false slideshow={"slide_type": "-"}
import pandas as pd
import seaborn as sns
# + hidePrompt=false
data=pd.read_csv('EcommercePurchases.csv')
# -
# ## The DataHead
# + slideshow={"slide_type": "-"}
#Data Head
data.head()
# -
# ## Detail information of the Data
# + slideshow={"slide_type": "-"}
data.info()
# -
# ### 1.How many Row and Column ?
#
# + hideCode=false
count_row=data.shape[0]
count_column=data.shape[1]
print('Row =',count_row,'and Column =',count_column)
# + [markdown] hideCode=false
# ### 2. The Average, Highest & Lowest 'Purchase Prices'
# + hidePrompt=false
#Average of Purchase Price
print('Average of Purchase Price:',data['Purchase Price'].mean())
# + hidePrompt=false
#Max of Purchase price
print('Max number of Purchase Price:',data['Purchase Price'].max())
# -
#Min of purchase price
print('Min number of Purchase Price:',data['Purchase Price'].min())
# ### 3. How many people have English 'en' as their Language of choice on the website?
data_en=data.loc[data['Language']=='en']
data_en_count=data_en['Language'].value_counts()
print('Total :',data_en_count)
# ### 4. How many people have the job title of "Lawyer" ?
#
data_lawyer=data.loc[data['Job']=='Lawyer']
data_lawyer['Job'].value_counts()
# ### 5.How many people made the purchase during the AM and how many people made the purchase during PM ?
data_time=data.loc[data['AM or PM'].isin(['AM','PM'])]
data_time['AM or PM'].value_counts()
# ### 6. What are the 5 most common Job Titles?
data['Job'].value_counts().nlargest(5)
# ### 7. Someone made a purchase that came from Lot: "90 WT" , what was the Purchase Price for this transaction?
data.loc[data['Lot']=="90 WT"]['Purchase Price']
# ### 8. What is the email of the person with the following Credit Card Number: 4926535242672853 ?
data.loc[data['Credit Card']==4926535242672853]['Email']
# ### 9. How many people have American Express as their Credit Card Provider *and* made a purchase above $95 ?
data_ccprovider=data.loc[(data['CC Provider']=='American Express') & (data['Purchase Price']>95)]
data_ccprovider['CC Provider'].value_counts()
# ### 10. (Hard): How many people have a credit card that expires in 2025?
# #### 10.1 List the credit card that expires in 2025
data['CC Exp Date']=pd.to_datetime(data['CC Exp Date'],format='%m/%y')
s2=data.loc[(data['CC Exp Date'] < "2026-01") & (data['CC Exp Date'] > "2024-12")]
s2['CC Exp Date'].value_counts()
#data[(data['CC Exp Date']>"2025-01-01" & (data['CC Exp Date']< "2025-12-01"))]
# #### 10.2 How many people (In total) have a Credit Card expires in 2025
#Total number of people have a credit card that expires in 2025
s2['CC Exp Date'].value_counts().sum()
# ### 11. (Hard): What are the top 5 most popular email providers/hosts (e.g. gmail.com, yahoo.com, etc...)
data_email=data['Email'].str.split('@').str[1]
data_email_count=data_email.value_counts()
data_email_plot=data_email.value_counts().nlargest(5)
data_email_plot
# ### 12. (Hard): What are the most popular Browser Info (or Browser version) ?
data['Browser Info'].unique()
data_browser=data['Browser Info'].str.split('(').str[0]
data_browser_plot=data_browser.value_counts().nlargest(10)
data_browser_plot
# # Data Visualization
#
# ### Implement a bar plot for top 5 most popular email providers/hosts
data_email_plot.plot(kind='barh',figsize=(8,5))
# ### Implement a Plot distribution of Purchase Price
# #### * Plot distribution of Purchase Price per Time (AM - PM)
sns.distplot(data[data['AM or PM']=="PM"]['Purchase Price'])
sns.distplot(data[data['AM or PM']=="AM"]['Purchase Price'])
# #### * Plot distribution of Purchase Price
sns.distplot(data['Purchase Price'])
# ### Implement countplot on Language
sns.catplot(x='Language',data=data,kind='count',aspect=2)
# ### Implement countplot on Browser
data_browser_plot.plot(kind='barh',figsize=(10,5))
# # Great Job!
| Ecommerce+Purchases+Exercise+.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework03: Topic Modeling with Latent Semantic Analysis
# Latent Semantic Analysis (LSA) is a method for finding latent similarities between documents treated as a bag of words by using a low rank approximation. It is used for document classification, clustering and retrieval. For example, LSA can be used to search for prior art given a new patent application. In this homework, we will implement a small library for simple latent semantic analysis as a practical example of the application of SVD. The ideas are very similar to PCA. SVD is also used in recommender systems in an similar fashion (for an SVD-based recommender system library, see [Surpise](http://surpriselib.com).
#
# We will implement a toy example of LSA to get familiar with the ideas. If you want to use LSA or similar methods for statistical language analysis, the most efficient Python libraries are probably [gensim](https://radimrehurek.com/gensim/) and [spaCy](https://spacy.io) - these also provide an online algorithm - i.e. the training information can be continuously updated. Other useful functions for processing natural language can be found in the [Natural Language Toolkit](http://www.nltk.org/).
# **Note**: The SVD from scipy.linalg performs a full decomposition, which is inefficient since we only need to decompose until we get the first k singluar values. If the SVD from `scipy.linalg` is too slow, please use the `sparsesvd` function from the [sparsesvd](https://pypi.python.org/pypi/sparsesvd/) package to perform SVD instead. You can install in the usual way with
# ```
# # !pip install sparsesvd
# ```
#
# Then import the following
# ```python
# from sparsesvd import sparsesvd
# from scipy.sparse import csc_matrix
# ```
#
# and use as follows
# ```python
# sparsesvd(csc_matrix(M), k=10)
# ```
# **Exercise 1 (20 points)**. Calculating pairwise distance matrices.
#
# Suppose we want to construct a distance matrix between the rows of a matrix. For example, given the matrix
#
# ```python
# M = np.array([[1,2,3],[4,5,6]])
# ```
#
# the distance matrix using Euclidean distance as the measure would be
# ```python
# [[ 0.000 1.414 2.828]
# [ 1.414 0.000 1.414]
# [ 2.828 1.414 0.000]]
# ```
# if $M$ was a collection of column vectors.
#
# Write a function to calculate the pairwise-distance matrix given the matrix $M$ and some arbitrary distance function. Your functions should have the following signature:
# ```
# def func_name(M, distance_func):
# pass
# ```
#
# 0. Write a distance function for the Euclidean, squared Euclidean and cosine measures.
# 1. Write the function using looping for M as a collection of row vectors.
# 2. Write the function using looping for M as a collection of column vectors.
# 3. Wrtie the function using broadcasting for M as a collection of row vectors.
# 4. Write the function using broadcasting for M as a collection of column vectors.
#
# For 3 and 4, try to avoid using transposition (but if you get stuck, there will be no penalty for using transposition). Check that all four functions give the same result when applied to the given matrix $M$.
# **Exercise 2 (20 points)**.
# **Exercise 2 (20 points)**. Write 3 functions to calculate the term frequency (tf), the inverse document frequency (idf) and the product (tf-idf). Each function should take a single argument `docs`, which is a dictionary of (key=identifier, value=document text) pairs, and return an appropriately sized array. Convert '-' to ' ' (space), remove punctuation, convert text to lowercase and split on whitespace to generate a collection of terms from the document text.
#
# - tf = the number of occurrences of term $i$ in document $j$
# - idf = $\log \frac{n}{1 + \text{df}_i}$ where $n$ is the total number of documents and $\text{df}_i$ is the number of documents in which term $i$ occurs.
#
# Print the table of tf-idf values for the following document collection
#
# ```
# s1 = "The quick brown fox"
# s2 = "Brown fox jumps over the jumps jumps jumps"
# s3 = "The the the lazy dog elephant."
# s4 = "The the the the the dog peacock lion tiger elephant"
#
# docs = {'s1': s1, 's2': s2, 's3': s3, 's4': s4}
# ```
# **Exercise 3 (20 points)**.
#
# 1. Write a function that takes a matrix $M$ and an integer $k$ as arguments, and reconstructs a reduced matrix using only the $k$ largest singular values. Use the `scipy.linagl.svd` function to perform the decomposition. This is the least squares approximation to the matrix $M$ in $k$ dimensions.
#
# 2. Apply the function you just wrote to the following term-frequency matrix for a set of $9$ documents using $k=2$ and print the reconstructed matrix $M'$.
# ```
# M = np.array([[1, 0, 0, 1, 0, 0, 0, 0, 0],
# [1, 0, 1, 0, 0, 0, 0, 0, 0],
# [1, 1, 0, 0, 0, 0, 0, 0, 0],
# [0, 1, 1, 0, 1, 0, 0, 0, 0],
# [0, 1, 1, 2, 0, 0, 0, 0, 0],
# [0, 1, 0, 0, 1, 0, 0, 0, 0],
# [0, 1, 0, 0, 1, 0, 0, 0, 0],
# [0, 0, 1, 1, 0, 0, 0, 0, 0],
# [0, 1, 0, 0, 0, 0, 0, 0, 1],
# [0, 0, 0, 0, 0, 1, 1, 1, 0],
# [0, 0, 0, 0, 0, 0, 1, 1, 1],
# [0, 0, 0, 0, 0, 0, 0, 1, 1]])
# ```
#
# 3. Calculate the pairwise correlation matrix for the original matrix M and the reconstructed matrix using $k=2$ singular values (you may use [scipy.stats.spearmanr](http://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.spearmanr.html) to do the calculations). Consider the fist 5 sets of documents as one group $G1$ and the last 4 as another group $G2$ (i.e. first 5 and last 4 columns). What is the average within group correlation for $G1$, $G2$ and the average cross-group correlation for G1-G2 using either $M$ or $M'$. (Do not include self-correlation in the within-group calculations.).
# **Exercise 4 (40 points)**. Clustering with LSA
#
# 1. Begin by loading a PubMed database of selected article titles using 'pickle'. With the following:
# ```import pickle
# docs = pickle.load(open('pubmed.pic', 'rb'))```
#
# Create a tf-idf matrix for every term that appears at least once in any of the documents. What is the shape of the tf-idf matrix?
#
# 2. Perform SVD on the tf-idf matrix to obtain $U \Sigma V^T$ (often written as $T \Sigma D^T$ in this context with $T$ representing the terms and $D$ representing the documents). If we set all but the top $k$ singular values to 0, the reconstructed matrix is essentially $U_k \Sigma_k V_k^T$, where $U_k$ is $m \times k$, $\Sigma_k$ is $k \times k$ and $V_k^T$ is $k \times n$. Terms in this reduced space are represented by $U_k \Sigma_k$ and documents by $\Sigma_k V^T_k$. Reconstruct the matrix using the first $k=10$ singular values.
#
# 3. Use agglomerative hierarchical clustering with complete linkage to plot a dendrogram and comment on the likely number of document clusters with $k = 100$. Use the dendrogram function from [SciPy ](https://docs.scipy.org/doc/scipy-0.15.1/reference/generated/scipy.cluster.hierarchy.dendrogram.html).
#
# 4. Determine how similar each of the original documents is to the new document `data/mystery.txt`. Since $A = U \Sigma V^T$, we also have $V = A^T U S^{-1}$ using orthogonality and the rule for transposing matrix products. This suggests that in order to map the new document to the same concept space, first find the tf-idf vector $v$ for the new document - this must contain all (and only) the terms present in the existing tf-idx matrix. Then the query vector $q$ is given by $v^T U_k \Sigma_k^{-1}$. Find the 10 documents most similar to the new document and the 10 most dissimilar.
# **Notes on the Pubmed articles**
#
# These were downloaded with the following script.
#
# ```python
# from Bio import Entrez, Medline
# Entrez.email = "YOUR EMAIL HERE"
# import cPickle
#
# try:
# docs = cPickle.load(open('pubmed.pic'))
# except Exception, e:
# print e
#
# docs = {}
# for term in ['plasmodium', 'diabetes', 'asthma', 'cytometry']:
# handle = Entrez.esearch(db="pubmed", term=term, retmax=50)
# result = Entrez.read(handle)
# handle.close()
# idlist = result["IdList"]
# handle2 = Entrez.efetch(db="pubmed", id=idlist, rettype="medline", retmode="text")
# result2 = Medline.parse(handle2)
# for record in result2:
# title = record.get("TI", None)
# abstract = record.get("AB", None)
# if title is None or abstract is None:
# continue
# docs[title] = '\n'.join([title, abstract])
# print title
# handle2.close()
# cPickle.dump(docs, open('pubmed.pic', 'w'))
# docs.values()
# ```
| homework/Homework07.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from bs4 import BeautifulSoup
import string
import requests
from selenium import webdriver
import spacy
import os
import pandas as pd
import re
from itertools import islice
import numpy as np
from nltk.tokenize import word_tokenize
import nltk
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
import csv
nltk.download('stopwords')
nltk.download('punkt')
import heapq
import csv
# ### 1.3 Parse downloaded pages
#
# This function **scrap_book** is the main core of our scraping part.
# Using the "BeautifulSoup" library, *scrap_book* takes as input the html page in soup_format and scrapes the different data that we need to build our dataset.
#
# As others input, there is **nlp** that is a tool for the Natural Language Processing and **n_link** that is useful to retrive the book's Url from the list previopus created
def scrap_book(page_soup , nlp , n_link):
n_link = int(n_link)
n_link = n_link-1
#Scrap the book's title
book_title = page_soup.find_all('h1', id ="bookTitle")[0].contents[0].replace('\n', '').strip()
print(book_title)
# Scrap the book's serie
series = page_soup.find_all('h2', id="bookSeries")[0].contents[0].replace('\n', '').strip()
#Scrap the book's author
author = page_soup.find_all('span', itemprop='name')[0].contents[0].replace('\n', '').strip()
#Scrap the rating value
rating_value = page_soup.find_all('span', itemprop='ratingValue')[0].contents[0].replace('\n', '').strip()
#Scrap the Rating_Count and Review_Count
ratings = page_soup.find_all('a', href="#other_reviews")
rating_count = -1
rating = -1
for raiting in ratings:
if raiting.find_all('meta', itemprop="ratingCount"):
rating_count = raiting.text.replace('\n', '').strip().split(' ')[0]
elif raiting.find_all('meta', itemprop="reviewCount"):
review_count = raiting.text.replace('\n', '').strip().split(' ')[0]
#Scrap the Plot
description = ' '.join([c for c in page_soup.find_all('span', id = re.compile(r'freeText\d'))[0].contents \
if isinstance(c, str)])
doc = nlp(description)
token_list_plot = [token for token in doc if not token.is_stop and not token.is_punct]
#Scrap number of pages
n_pages = page_soup.find_all('span', itemprop='numberOfPages')[0].contents[0].strip().split(' ')[0]
#Scrap Publishing date of the book
pub_date = page_soup.find_all('div', {"class": "row"})[1].contents[0].strip().split(' ')
pub_date = " ".join(pub_date)
pub_date = pub_date.replace('\n', '').strip()
pub_date = pub_date.replace(" ","")
pub_date = pub_date.split('d')[1]
#Scrap Characters
Characters = []
lenght_c = page_soup.find_all('a', {'href': re.compile(r'/characters/')})
l = len(lenght_c)
for i in range (0, l):
character = page_soup.find_all('a', {'href': re.compile(r'/characters/')})[i].contents[0]
Characters.append(character)
#Scrap Settings
Settings = []
lenght_s = page_soup.find_all('a', {'href': re.compile(r'/places/')})
l2 = len(lenght_s)
for i in range (0, l2):
setting = page_soup.find_all('a', {'href': re.compile(r'/places/')})[i].contents[0]
Settings.append(setting)
Settings = " ".join(Settings)
#scrap original book's link
f=open('lista_url.txt')
lines=f.readlines()
URL = lines[n_link]
return book_title,series,author,rating_value,rating_count, \
review_count,token_list_plot,n_pages,pub_date,Characters,Settings,URL
# ---------------------------------------------------------------------------------------
#
# This is the script that we launch in order to extract information from the html pages and store them in the file "articles.tsv".
#
# **filepath** : It is the forder in which there are the .html pages previous downloaded
#
# **articles.tsv** : It 's the file that we build. Using the function *scrap_book* we extract info about each page/book and put them in this file, in order to build our dataset.
#filepath = r"C:\Users\thoma\Desktop\HW3_ADM\html_folder3\articoli_7"
nlp = spacy.load('en_core_web_sm')
with open('articles.tsv', 'a', encoding="utf-8") as out_file:
tsv_writer = csv.writer(out_file, delimiter='\t')
#tsv_writer.writerow(['bookTitle', 'bookSeries','bookAuthors','ratingValue','ratingCount','reviewCount','Plot','NumberofPages','Published','characters','Settings','URL'])
for i in range (24001,30001):
try :
print(i)
string1 = "article_"
string2 = str(i)
string3 = ".html"
title = string1 + string2 + string3
soup = BeautifulSoup(open(os.path.join(filepath, title),encoding='utf-8' ), features="lxml")
bookTitle,bookSeries,bookAuthors,ratingValue,ratingCount, reviewCount, Plot, \
NumberofPages,Published, characters,Settings,URL = scrap_book(soup, nlp,string2)
riga_info = (bookTitle,bookSeries,bookAuthors,ratingValue,ratingCount, reviewCount, \
Plot, NumberofPages,Published,characters,Settings,URL)
tsv_writer.writerow([riga_info[0],riga_info[1],riga_info[2],riga_info[3],riga_info[4],riga_info[5], \
riga_info[6],riga_info[7],riga_info[8],riga_info[9],riga_info[10],riga_info[11]])
#For some reason related to the differents pages of the website could be an error in scrping process: we handle it.
except IndexError as e :
print(" lost a book because of scraping error")
| parser.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deadbeat controller for the double integrator
#
# Consider a mass of 1kg moving in one direction on a friction-free horizontal surface. We can apply a force to the mass (input signal $u$), and the mass is also subject to disturbance forces $v$. We are interested in controlling the position $z$ of the mass. In continuous time the dynamics are described by
# $$ \ddot{z} = u + v. $$
# Introducing the state variables $x_1=z$ and $x_2=\dot{z}$, the system can also be represented on state-space form with state vector $x = \begin{bmatrix} z & \dot{z}\end{bmatrix}^T$ as
# \begin{align}
# \dot{x} &= \underbrace{\begin{bmatrix} 0 & 1\\0 & 0\end{bmatrix}}_{A}x + \underbrace{\begin{bmatrix}0\\1\end{bmatrix}}_{B}u + \underbrace{\begin{bmatrix}0\\1\end{bmatrix}}_{B}v\\
# y &= \underbrace{\begin{bmatrix}1 & 0 \end{bmatrix}}_C x
# \end{align}
# ## Discrete-time state-space model
# The discrete-time state-space model using a sampling period $h$ is
# \begin{align}
# x(k+1) &= \Phi(h)x(k) + \Gamma(h)u + \Gamma(h)v\\
# y(k) &= Cx(k)
# \end{align}
# where
# $$ \Phi(h) = \mathrm{e}^{Ah} = \begin{bmatrix} 1 & h\\0 & 1 \end{bmatrix}$$
# and
# $$ \Gamma(h) = \int_0^h \mathrm{e}^{As}B ds = \begin{bmatrix} \frac{h^2}{2}\\h \end{bmatrix}.$$
# ### Verification by symbolic computation
# +
import numpy as np
import sympy as sy
sy.init_printing(use_latex='mathjax', order='lex')
h = sy.symbols('h', real=True, positive=True)
A = sy.Matrix([[0,1], [0,0]])
B = sy.Matrix([[0],[1]])
Phi = sy.simplify(sy.exp(A*h))
Phi
# -
s = sy.symbols('s')
Gamma = sy.integrate(sy.exp(A*s)*B, (s, 0, h))
Gamma
# ## Reachability
# The controllability matrix for this second order system becomes
# $$ W_c = \begin{bmatrix} \Gamma & \Phi\Gamma \end{bmatrix} = \begin{bmatrix} \frac{h^2}{2} & \frac{3h^2}{2}\\h & h\end{bmatrix}, $$
# with determinant
# $$\det W_c = \frac{h^3}{2}(1 - 3) = -h^3,$$
# which is different from zero since $h>0$.
# It is hence possible to reach any point in the state-space from any other point in just two steps (two sampling periods).
# ### Verification by symbolic computation
Wc = sy.BlockMatrix([[Gamma, Phi*Gamma]]).as_explicit()
Wc
sy.det(Wc)
# ## Designing an input sequence
# We now know that the system is reachable. This means that we can take the system from the origin in the state-space (position zero and velocity zero) to any other point in state-space. And it can be done in only two steps with the input sequence
# $$ u(0), \, u(1).$$
# Let's say we want to reach the point
# $$ x_d = \begin{bmatrix} a\\b \end{bmatrix},$$
# which in words is that we want the mass to be at $z=a$ and having the velocity $\dot{z}=b$. The general solution for an n-th order discrete-time state space system is
# \begin{align}
# x(n) &= \Phi^n x(0) + \Phi^{n-1}\Gamma u(0) + \Phi^{n-2}\Gamma u(1) + \cdots + \Gamma u(n-1)\\
# &= \Phi^n x(0) + W_cU,
# \end{align}
# where
# $$ U = \begin{bmatrix} u(n-1)\\u(n-2)\\\vdots\\u(0)\end{bmatrix}. $$
# In the case here we have $x(0)=0$ and this leads to the equation
# $$ W_cU = x_d, \qquad \text{with solution}$$
# \begin{align}
# U &= \begin{bmatrix}u(1)\\u(0)\end{bmatrix} = W_c^{-1}x_d = \begin{bmatrix} \frac{h^2}{2} & \frac{3h^2}{2}\\h & h\end{bmatrix}^{-1} \begin{bmatrix} a\\b \end{bmatrix}\\
# &= \frac{1}{-h^3} \begin{bmatrix} h & -\frac{3h^2}{2}\\-h & \frac{h^2}{2} \end{bmatrix} \begin{bmatrix} a\\b \end{bmatrix}\\
# &= \begin{bmatrix} -\frac{1}{h^2} & \frac{3}{2h}\\\frac{1}{h^2} & -\frac{1}{2h} \end{bmatrix} \begin{bmatrix} a\\b \end{bmatrix}\\
# &= \begin{bmatrix} -\frac{a}{h^2} + \frac{3b}{2h}\\ \frac{a}{h^2} - \frac{b}{2h} \end{bmatrix}.
# \end{align}
# Thus the input sequence becomes $u(0) = \frac{a}{h^2} - \frac{b}{2h}$, $u(1) = \frac{-a}{h^2} + \frac{3b}{2h}$.
# ### Verification with symbolic computation
# Verify
a,b = sy.symbols('a,b')
U = Wc.inv()*sy.Matrix([[a],[b]])
U
# Simulate
u0 = U[1,0]
u1 = U[0,0]
x0 = sy.Matrix([[0],[0]])
x1 = Phi*x0 + Gamma*u0
x2 = Phi*x1 + Gamma*u1
sy.simplify(x2)
# ## State feedback
# Introducing the state-feedback control law
# $$ u = -l_1x_1 - l_2 x_2 + l_0y_{ref} = -Lx + l_0y_{ref}$$
# gives the closed-loop state-space system
# \begin{align}
# x(k+1) &= \Phi x(k) +\Gamma\big(-Lx(k) + l_0y_{ref}(k)\big) + \Gamma v(k) = \left( \Phi - \Gamma L \right) x(k) + l_0\Gamma y_{ref}(k) + \Gamma v(k)\\
# y(k) &= C x(k)
# \end{align}
# with characteristic polynomial given by
# \begin{align}
# \det \left( zI - (\Phi-\Gamma L) \right) &= \det \left( \begin{bmatrix} z & 0\\0 & z \end{bmatrix} - \begin{bmatrix} 1 & h\\0 & 1 \end{bmatrix} + \begin{bmatrix} l_1\frac{h^2}{2} & l_2\frac{h^2}{2}\\ l_1h & l_2h \end{bmatrix} \right)\\
# &= \det \begin{bmatrix} z-1+l_1\frac{h^2}{2} & -h+l_2\frac{h^2}{2}\\l_1h & z-1+l_2h
# \end{bmatrix}\\
# &= (z-1+l_1\frac{h^2}{2})(z-1+l_2h) - l_1h(-h + l_2\frac{h^2}{2})\\
# &= z^2 + (-1+l_2h-1+l_1\frac{h^2}{2}) z + (1-l_2h - l_1\frac{h^2}{2} + l_1l_2\frac{h^3}{2} +l_1h^2 -l_1l_2\frac{h^3}{2})\\
# &= z^2 + (l_1\frac{h^2}{2}+l_2h-2) z + (1 +l_1\frac{h^2}{2} -l_2h)
# \end{align}
# ### Verification by symbolic computation
l1, l2 = sy.symbols('l1, l2', real=True)
z = sy.symbols('z')
L = sy.Matrix([[l1, l2]])
ch_poly = sy.Poly((z*sy.eye(2) - (Phi - Gamma*L)).det(), z)
ch_poly.as_expr()
# ### Desired closed-loop characteristic polynomial
# Here we are interested in designing a deadbeat controller, so the desired closed-loop poles are
# $$ p_1 = 0, \qquad p_2=0,$$
# and the desired characteristic polynomial is
# $$ A_c(z) = (z-p_1)(z-p_2) = z^2. $$
# In the same spirit as when designing an RST controller using the polynomial approach, we set the calculated characteristic polynomial - obtained when introducing the linear state feedback- equal to the desired characteristic polynomial.
# \begin{align}
# z^1: \qquad l_1\frac{h^2}{2} + l_2h -2 &= 0\\
# z^0: \qquad l_1\frac{h^2}{2} - l_2h+1 &= 0
# \end{align}
# which can be written as the system of equations
# $$ \underbrace{\begin{bmatrix} \frac{h^2}{2} & h\\\frac{h^2}{2} & -h \end{bmatrix}}_{M} \underbrace{\begin{bmatrix} l_1\\l_2\end{bmatrix}}_{L^T} = \underbrace{\begin{bmatrix}2\\-1\end{bmatrix}}_{b} $$
# with solution given by
#
# $$L^T = M^{-1}b = \frac{1}{-h^3} \begin{bmatrix} -h & -h\\-\frac{h^2}{2} & \frac{h^2}{2} \end{bmatrix} \begin{bmatrix} 2\\-1 \end{bmatrix}$$
# $$ = -\frac{1}{h^3} \begin{bmatrix} -2h+h\\-h^2-\frac{h^2}{2}\end{bmatrix} = \begin{bmatrix} \frac{1}{h^2}\\\frac{3}{2h} \end{bmatrix} $$
# ### Verification by symbolic calculation
des_ch_poly = sy.Poly(z*z, z)
dioph_eqn = ch_poly - des_ch_poly
sol = sy.solve(dioph_eqn.coeffs(), (l1,l2))
sol
# In the system of equations $ML^T=b$ above, note that the matrix $M$ can be written
# $$ M = \begin{bmatrix} \frac{h^2}{2} & h\\\frac{h^2}{2} & -h \end{bmatrix} = \begin{bmatrix}1 & 0\\-2 & 1\end{bmatrix}\underbrace{\begin{bmatrix} \frac{h^2}{2} & h \\ \frac{3h^2}{2} & h\end{bmatrix}}_{W_c^T}, $$
# so $M$ will be invertible if and only if $\det W_c^T = \det W_c \neq 0$.
# ## The resulting closed-loop system
# So, we have found the control law
# $$ u(k) = -Lx(k) + l_0y_{ref}(k) = -\begin{bmatrix} \frac{1}{h^2} & \frac{3}{2h} \end{bmatrix}x(k) + l_0 y_{ref}(k)$$
# which gives a closed-loop system with poles in the origin, i.e. deadbeat control. The closed-loop system becomes
# \begin{align*}
# x(k+1) &= \big( \Phi - \Gamma L \big) x(k) + \Gamma l_0 y_{ref}(k) + \Gamma v(k)\\
# &= \left( \begin{bmatrix} 1 & h\\0 & 1\end{bmatrix} - \begin{bmatrix} \frac{h^2}{2}\\h\end{bmatrix}\begin{bmatrix} \frac{1}{h^2} & \frac{3}{2h} \end{bmatrix} \right) x(k) + \Gamma l_0 y_{ref}(k) + \Gamma v(k)\\
# &= \left( \begin{bmatrix} 1 & h\\0 & 1\end{bmatrix} - \begin{bmatrix} \frac{1}{2} & \frac{3h}{4}\\ \frac{1}{h} & \frac{3}{2}\end{bmatrix}\right) x(k) + \Gamma l_0 y_{ref}(k) + \Gamma v(k)\\
# &= \underbrace{\begin{bmatrix} \frac{1}{2} & \frac{h}{4} \\-\frac{1}{h} & -\frac{1}{2}\end{bmatrix}}_{\Phi_c}x(k) + \begin{bmatrix}\frac{h^2}{2}\\h\end{bmatrix} l_0 y_{ref}(k) + \begin{bmatrix}\frac{h^2}{2}\\h\end{bmatrix} v(k)\\
# y(k) &= \begin{bmatrix} 1 & 0 \end{bmatrix} x(k)
# \end{align*}
# ### Verification using symbolic computations
L = sy.Matrix([[sol[l1], sol[l2]]])
Phic = Phi - Gamma*L
Phic
# ## Determining the reference signal gain $l_0$
# Consider the steady-state solution for a unit step in the reference signal. We set $y_{ref}=1$ and $v = 0$. This gives
# $$ x(k+1) = \Phi_c x(k) + \Gamma l_0. $$
# In steady-state there is no change in the state, so $x(k+1)=x(k)=x_{ss}$, which leads to
# $$ x_{ss} = \Phi_c x_{ss} + \Gamma l_0$$
# $$ (I - \Phi_c)x_{ss} = \Gamma l_0$$
# \begin{align}
# x_{ss} &= (I - \Phi_c)^{-1}\Gamma l_0\\
# &= \begin{bmatrix} \frac{1}{2} &-\frac{h}{4}\\ \frac{1}{h} & \frac{3}{2} \end{bmatrix}^{-1} \begin{bmatrix} \frac{h^2}{2}\\h \end{bmatrix} l_0\\
# &= \begin{bmatrix}\frac{3}{2} & \frac{h}{4}\\-\frac{1}{h} & \frac{1}{2} \end{bmatrix} \begin{bmatrix} \frac{h^2}{2}\\h\end{bmatrix} l_0\\
# &= \begin{bmatrix}\frac{3h^2}{4} + \frac{h^2}{4}\\-\frac{h}{2} + \frac{h}{2} \end{bmatrix}l_0= \begin{bmatrix}h^2\\ 0 \end{bmatrix}l_0\\
# \end{align}
# which means that the steady-state velocity $\dot{z}(\infty) = x_2(\infty) = 0$. This makes sense.
#
# We can now determine $l_0$. Since $y(k)=x_1(k)$ then $y_{ss} = h^2 l_0$ for a unit step in the reference signal. We would like the steady-state value $y_{ss}$ to be the same as the reference signal (which is equal to one, of course) so this gives
# $$ h^2l_0 = 1 \quad \Rightarrow \quad l_0 = \frac{1}{h^2}. $$
# ## Simulate step responses (symbolically)
# ### Step response from the reference
l0 = 1/(h*h)
C = sy.Matrix([[1,0]])
x = sy.Matrix([[0],[0]]) # Initial state
yref = sy.Matrix([[1]])
xs = [x] # List to hold state trajectory
us = [[0]] # and control signal
ys = [[0]] # and system output
for k in range(6): # No need to simulate too long. It is deadbeat control after all
us.append(-L*x + l0*yref)
x = Phic*x + Gamma*l0*yref
xs.append(x)
ys.append(C*x)
xs
us
# ### Step response from the disturbance
x = sy.Matrix([[0],[0]]) # Initial state
yref = sy.Matrix([[0]])
v = sy.Matrix([[1]])
xs = [x] # List to hold state trajectory
us = [[0]] # and control signal
ys = [[0]] # and system output
for k in range(6): # No need to simulate too long. It is deadbeat control after all
us.append(-L*x + l0*yref)
x = Phic*x + Gamma*l0*yref + Gamma*v
xs.append(x)
ys.append(C*x)
xs
# ## Simulate step-responses (numerically)
import control as ctrl
import matplotlib.pyplot as plt
# Convert to from sympy matrices to numpy
hval = .1
Phi_np = np.array(Phi.subs({h:hval})).astype(np.float64)
Gamma_np = np.array(Gamma.subs({h:hval})).astype(np.float64)
L_np = np.array(L.subs({h:hval})).astype(np.float64)
l0_np = np.array(l0.subs({h:hval})).astype(np.float64)
Phic_np = Phi_np - Gamma_np*L_np
C_np = np.array(C).astype(np.float64)
D_np = np.array([[0]])
sys_c = ctrl.ss(Phic_np, Gamma_np*l0_np, C_np, D_np, hval) # From ref signal
sys_cv = ctrl.ss(Phic_np, Gamma_np, C_np, D_np, hval) # From disturbance signal
tvec = np.asarray(np.arange(8))*hval
T, yout = ctrl.step_response(sys_c, tvec)
T, yout_v = ctrl.step_response(sys_cv, tvec)
plt.figure(figsize=(14,3))
plt.step(tvec, yout.flatten())
plt.figure(figsize=(14,3))
plt.step(tvec, yout_v.flatten())
# # Exercises
# ## Design a less agressive controller
# Consider to let the closed-loop poles be less fast. Choose something reasonable, for instance a double pole in $z=0.5$, or a pair of complex-conjugated poles in $z=0.6 \pm i0.3$. Redo the design, following the example above. Find the state feedback and simulate step-responses.
# ## Design a deadbeat controller for the DC-motor
# From the textbook (Åström & Wittenmark) Appendix:
# 
# 1. Use symbolic calculations to find the discrete-time state-space model for arbitrary sampling period $h$.
# 2. Design a deadbeat controller for arbitrary sampling period.
# 3. Assume a disturbance is acting on the input to the system, as an unknown torque on the motor shaft. This means that the disturbance enters into the system in the same way as the disturbance on the mass on frictionless surface analyzed above. Simulate step-responses for the closed-loop system.
| state-space/notebooks/Deadbeat controller for the double integrator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Logistic Regression for the clear area
import dask.dataframe as dd
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# Define the trainining data (X: input features, y : output feature)
# + tags=[]
df = dd.read_csv('test4_nosmoke*.csv')
df = df.compute()
print(df.shape)
X = df.iloc[:, :-1]
y = df.iloc[:, -1:]
X = X.to_numpy()
y = y.to_numpy().reshape(-1)
del df
# -
# Fit the LogisticRegression model with X and y
# Please note that through the GridSearchCV, we found the best performing C parameter: 100
# * _if you want to take a look at the code for the GridSearchCV, please take a look at the `python script` for the logistic regression_
from sklearn.linear_model import LogisticRegression
logreg = LogisticRegression(max_iter = 10000, n_jobs = -1, C = 100)
logreg.fit(X,y)
quant_names = pd.read_csv('clean_polyname.csv')
colors = ['blue' if abs(x) < 1.5 else 'red' for x in logreg.coef_[:,:78].reshape(-1)]
fig, ax = plt.subplots(figsize = (20,10))
ax.bar(x = quant_names.clean_polyname, height = logreg.coef_[:,:78].reshape(-1), color = colors)
ax.set_xticklabels(quant_names.clean_polyname, {'fontsize' : 13},rotation=90)
ax.set_yticklabels([-8,-6, -4,-2,0,2,4,6,8],{'fontsize' : 13},rotation=90)
[t.set_color(i) for (i,t) in
zip(colors,ax.xaxis.get_ticklabels())]
ax.set_title('Coefficients of the logistic model - Clear Area', {'fontsize': 30})
plt.savefig('logreg_coef.png')
plt.show()
colors = ['blue' if abs(x) < .5 else 'red' for x in logreg.coef_[:,78:].reshape(-1)]
fig, ax = plt.subplots(figsize = (20,10))
ax.bar(x = ['maskClose' + x for x in np.arange(25).astype(str)],height = logreg.coef_[:,78:].reshape(-1), color = colors)
ax.set_xticklabels( ['maskClose' + x for x in np.arange(25).astype(str)], {'fontsize' : 13},rotation=90)
ax.set_title('Coefficients of the logistic model (Categorical features) - Clear Area', {'fontsize': 30})
plt.savefig('logreg_coef_categories.png')
plt.show()
| Approach2/Step 4.1--------- Logistic Regrssion on the Clear Area.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problem1_TwoSum
# Given an array of integers, return indices of the two numbers such that they add up to a specific target.
#
# You may assume that each input would have exactly one solution.
# Example:
# Given nums = [2, 7, 11, 15], target = 9,
#
# Because nums[0] + nums[1] = 2 + 7 = 9,
# return [0, 1].
# The return format had been changed to zero-based indices. Please read the above updated description carefully.
# +
# O(n^2)
class Solution1(object):
def twoSum(self, nums, target):
"""
:type nums: List[int]
:type target: int
:rtype: List[int]
"""
for i in range(0,len(nums)):
for j in range(i+1,len(nums)):
if nums[i] + nums[j] == target:
return [i, j]
s1 = Solution1()
# -
s1.twoSum([3,2,4], 6)
import collections
# +
# O(n)
class Solution2(object):
def twoSum(self, nums, target):
"""
:type nums: List[int]
:type target: int
:rtype: List[int]
"""
ht = {}
for i, num in enumerate(nums):
ht[num] = i
for i, num in enumerate(nums):
if target - num in ht and i != ht[target - num]:
return [i, ht[target - num]]
s2 = Solution2()
# -
s2.twoSum([3,2,4], 6)
# %timeit s2.twoSum([2, 7, 11, 15], 9)
# %timeit s1.twoSum([2, 7, 11, 15], 9)
| Problem1_Two_Sum.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1.1 조건부 기댓값 Conditional expectation
# - 확률변수 Y의 기댓값 **by Conditional pdf** $f_{Y\vert X}(y|x)$
#
# $$\text{E}_Y[Y \vert X] = \int_{y=-\infty}^{y=\infty} y \, f_{Y \vert X}(y|x) dy$$
#
# $$\text{E}[Y \vert X] = \int y \, f(y \vert x) dy $$
#
# - 여기서 조건부 기댓값 $E[Y \vert X]$ 는 확률변수
# - eg. h(x): **조건 확률변수X** 받아, **결과값 확률변수 Y의 기댓값** 출력
#
# $$ \text{E}[Y \vert X=x] = h(x) $$
# # 1.2 전체 기댓값의 법칙 Law of iterated expectation
# - 두 번 기댓값 구한 것은 원래 확률별수의 보통 기댓값과 같다
#
# $$\text{E}_X[\text{E}_Y[Y \vert X]] = \text{E}_Y[Y]$$
#
# $$\text{E}[\text{E}[\text{Y} \vert \text{X}]] = \text{E}[\text{Y}]$$
# # 2.1 조건부 분산 Conditional variance
# - X=x일 때, Y는 어떤 분포를 따르는데, 그 폭을 나타냄
#
# $$
# \text{Var}_\text{Y} [Y \vert X]
# $$
#
# $$
# = E_Y[(Y - E_X[Y \vert X])^2 \vert X]
# $$
#
# $$
# = \int (Y - E_Y[Y \vert X])^2f_{Y \vert X}(y \vert x)dy
# $$
#
# # 2.2 전체 분산의 법칙 law of total variance
# - 확률변수의 분산 = 조건부 분산의 기댓값 + 조건부 기댓값의 분산
# - $\text{E}[\text{분산}] + \text{Var}[\text{평균}]$
#
# $$
# \text {Var}[Y] = E[Var[Y \vert X]] + Var[E[Y \vert X]]$$
#
| 06.Math/9.2.4 Conditional Expectation(important).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia
# language: julia
# name: julia-1.5
# ---
# # Cubic splines
#
# For illustration, here is a spline interpolant using just a few nodes.
using FundamentalsNumericalComputation
# +
f = x -> exp(sin(7*x))
plot(f,0,1,label="function")
t = [0, 0.075, 0.25, 0.55, 0.7, 1] # nodes
y = f.(t) # values at nodes
scatter!(t,y,label="nodes")
# +
S = FNC.spinterp(t,y)
plot!(S,0,1,label="spline")
# -
# Now we look at the convergence rate as the number of nodes increases.
# +
x = (0:10000)/1e4 # sample the difference at many points
n = @. round(Int,2^(3:0.5:8)) # numbers of nodes
err = zeros(0)
for n in n
t = (0:n)/n
S = FNC.spinterp(t,f.(t))
dif = @. f(x)-S(x)
push!(err,norm(dif,Inf))
end
pretty_table((n=n,error=err),backend=:html)
# -
# Since we expect convergence that is $O(h^4)=O(n^{-4})$, we use a log-log graph of error and expect a straight line of slope $-4$.
# +
order4 = @. (n/n[1])^(-4)
plot(n,[err order4],m=[:o :none],l=[:solid :dash],label=["error" "4th order"],
xaxis=(:log10,"n"), yaxis=(:log10,"\$\\| f-S \\|_\\infty\$"),
title="Convergence of spline interpolation")
| book/localapprox/demos/splines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Scattertext für mich
import numpy as np
import tensorflow as tf
import model_cnn
import preprocessing_classification as pre_c
np.set_printoptions(threshold=np.nan)
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import os, glob
# +
sess = tf.InteractiveSession()
text_length = 1000
num_authors = 5
input_cnn = tf.placeholder(tf.float32, [None, len(pre_c.alphabet), text_length, 1], name="input_x")
with tf.variable_scope("cnn"):
cnn_logits, cnn_variables, _ = model_cnn.inference(
input_x=input_cnn, keep_prob=1.0, num_authors=num_authors)
known_vars = []
known_vars = tf.global_variables()
saver = tf.train.Saver(var_list=known_vars)
saver.restore(sess, "../resources/1511967049/saves/cnn.ckpt-00009384")
print("cnn_classifier restored")
# -
# ## Vergleich von Filter 152 und 42
n = 7
ngram_act_dict = {}
activations_file_list = ["../resources/activations-five-authors/TrainSet-five_authors.txt-activations-"+str(i)+".npz" for i in range(35)]
text_file_list = ["../resources/activations-five-authors/TrainSet-five_authors.txt-texts-"+str(i)+".txt" for i in range(35)]
zipped = zip(activations_file_list, text_file_list)
ctr = 0
for acts_file, texts_file in zipped:
logits = np.load(acts_file)['logits']
filt = np.where(logits[:,0] == logits[:,1])
acts = np.load(acts_file)['act_7'].reshape(-1, 980, 256)[filt]
with open(texts_file, mode='r') as opened_texts:
texts = np.array(opened_texts.read().split('\n')[:-1])[filt]
for text_i, text in enumerate(texts):
for char_i in range(1, 20):
ngram = text[char_i:char_i+n]
if ngram not in ngram_act_dict:
ngram_act_dict[ngram] = [[0],[0]]
ngram_act_dict[ngram][0].append(np.mean(acts[text_i,0:char_i,42]))
ngram_act_dict[ngram][1].append(np.mean(acts[text_i,0:char_i,152]))
for char_i in range(20, len(text)-n):
ngram = text[char_i:char_i+n]
if ngram not in ngram_act_dict:
ngram_act_dict[ngram] = [[0],[0]]
ngram_act_dict[ngram][0].append(np.mean(acts[text_i,char_i-20:char_i,42]))
ngram_act_dict[ngram][1].append(np.mean(acts[text_i,char_i-20:char_i,152]))
ctr += 1
print("{} von {} Dateien ausgelesen.".format(ctr,len(activations_file_list)))
ngram_mean_act_dict = {}
for ngram in ngram_act_dict:
ngram_mean_act_dict[ngram] = [np.mean(np.array(a)) for a in ngram_act_dict[ngram]]
arr = np.array([ngram_mean_act_dict[ngram] for ngram in ngram_mean_act_dict.keys()])
df = pd.DataFrame(arr, index=ngram_mean_act_dict.keys(), columns=["filt_42", "filt_152"])
df.to_csv('ngram_mean_act_' + str(n) + '.csv')
# ## Visualization
df.plot.scatter(x='filt_42', y='filt_152', s=1, loglog=False, grid=True)
df[df.filt_42 > 0 ][df.filt_152 > 0].plot.hist(alpha=0.5, bins=100, logy=True)
# ## Wortlisten
# Top _42
df.nlargest(20, 'filt_42').reset_index()[['index','filt_42']].join(df.nlargest(20, 'filt_152').reset_index()[['index','filt_152']], lsuffix=' Top _42', rsuffix=' Top _152')
| src/new-scattertext.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
import numpy as np
# ### Torch and Numpy - rand
# +
xt = torch.rand(3,2)
print(xt)
xn = np.random.rand(3,2)
print(xn)
print(f'type of xt = {type(xt)}, type of xn = {type(xn)}')
# +
xt = torch.rand(3,2,3)
print(xt)
xn = np.random.rand(3,2,3)
print(xn)
print(f'type of xt = {type(xt)}, type of xn = {type(xn)}')
# -
print(xt*xn)
# ### Slicing
# +
x = torch.rand(3,4)
y = np.random.rand(3,4)
print(x)
# -
print(x[:, 1]) # only column #1
print(x[:, 1:3]) # only column #1 to until #3
print(x[1:, :]) # from #1th row until all row and all columns
# from row #1 row until all row
# from column #1 to until #3
print(x[1:, 1:3])
# + [markdown] tags=[]
# ### Resizing
# -
print(x.size())
y = x.view(12) # one dimentional
print(y)
y = x.view(2, 6) # 2x6
print(y)
# ### Simple Torch operations
# +
x = torch.rand(2,3)
y = torch.rand(2,3)
# ADDITIONS # SUB, MUL, DIV
z = x + y # - * /
z = torch.add(x, y) # sub mul div
y.add_(x) # inplace #
# -
# ### Torch <-> Numpy conversion
# +
# torch to numpy
print(x)
print(type(x))
n = x.numpy()
print(n)
print(type(n))
# -
# #### Important - deep copy
# +
x.add_(1) # inplace add 1
print(x)
print(n) # 1 is added to "n" too
# +
n = x.clone().numpy()
x.add_(1)
print(x)
print(n) # did not added
# -
| PyTorch learning 1 Declaration, slicing, resizing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import os
import sys
import numpy as np
MNM_nb_folder = os.path.join('..', 'network_builder')
sys.path.append(MNM_nb_folder)
python_lib_folder = os.path.join('..', '..', 'src', 'pylib')
sys.path.append(python_lib_folder)
from MNM_nb import *
data_folder = os.path.join('..', '..', 'data', 'input_files_33net')
nb = MNM_network_builder()
nb.load_from_folder(data_folder)
nb.graph.G
path_list = list()
path_ID_counter = 0
for O,D in nb.demand.demand_list:
O_node = nb.od.O_dict[O]
D_node = nb.od.D_dict[D]
print O, D, O_node, D_node
tmp_path_set = MNM_pathset()
tmp_path_set.origin_node = O_node
tmp_path_set.destination_node = D_node
tmp_path_node_list = list(nx.all_simple_paths(nb.graph.G, source=O_node, target=D_node))
for path_node_list in tmp_path_node_list:
tmp_path = MNM_path(path_node_list, path_ID_counter)
tmp_path.create_route_choice_portions(nb.config.config_dict['DTA']['max_interval'])
nb.path_table.ID2path[tmp_path.path_ID] = tmp_path
tmp_path_set.add_path(tmp_path)
path_ID_counter += 1
tmp_path_set.normalize_route_portions()
nb.route_choice_flag = True
nb.dump_to_folder(os.path.join('..', '..', 'data', 'input_files_33net'))
| side_project/MNM_fake_builder/generate_all_paths.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def a_n(a1, d, n):
'''Return the n-th term of the arithmetic sequence.
:a1: first term of the sequence. Integer or real.
:d: common difference of the sequence. Integer or real.
:n: the n-th term in sequence
returns: n-th term. Integer or real.'''
an = a1 + (n - 1)*d
return an
a_n(4, 3, 10)
def a_seq(a1, d, n):
'''Obtain the whole arithmetic sequence up to n.
:a1: first term of the sequence. Integer or real.
:d: common difference of the sequence. Integer or real.
:n: length of sequence
returns: sequence as a list.'''
sequence = []
for _ in range(n):
sequence.append(a1)
a1 = a1 + d
return sequence
a_seq(4, 3, 10)
# +
def infinite_a_sequence(a1, d):
while True:
yield a1
a1 = a1 + d
for i in infinite_a_sequence(4,3):
print(i, end=" ")
# -
sum(a_seq(4, 3, 10))
def a_series(a1, d, n):
result = n * (a1 + a_n(a1, d, n)) / 2
return result
a_series(4, 3, 10)
| Exercise01/Determining_the_nth_Term_of_an_Arithmetic_Sequence_and_Arithmetic_Series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
# Copyright 2021 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# -
# # PyTorch Image Classification Multi-Node Distributed Data Parallel Training on GPU using Vertex Training with Custom Container
# <table align="left">
# <td>
# <a href="https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/master/community-content/pytorch_image_classification_distributed_data_parallel_training_with_vertex_sdk/multi_node_ddp_nccl_vertex_training_with_custom_container.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
# View on GitHub
# </a>
# </td>
# </table>
# ## Setup
# + pycharm={"name": "#%%\n"}
PROJECT_ID = "YOUR PROJECT ID"
BUCKET_NAME = "gs://YOUR BUCKET NAME"
REGION = "YOUR REGION"
SERVICE_ACCOUNT = "YOUR SERVICE ACCOUNT"
# + pycharm={"name": "#%%\n"}
# ! gsutil ls -al $BUCKET_NAME
# + pycharm={"name": "#%%\n"}
content_name = "pt-img-cls-multi-node-ddp-cust-cont"
# -
# ## Vertex Training using Vertex SDK and Custom Container
# ### Built Custom Container
# + pycharm={"name": "#%%\n"}
hostname = "gcr.io"
image_name = content_name
tag = "latest"
custom_container_image_uri=f"{hostname}/{PROJECT_ID}/{image_name}:{tag}"
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Initialize Vertex SDK
# + pycharm={"name": "#%%\n"}
# ! pip install -r requirements.txt
# + pycharm={"name": "#%%\n"}
from google.cloud import aiplatform
aiplatform.init(
project=PROJECT_ID,
staging_bucket=BUCKET_NAME,
location=REGION,
)
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Create a Vertex Tensorboard Instance
# + pycharm={"name": "#%%\n"}
content_name = content_name + "-gpu"
tensorboard = aiplatform.Tensorboard.create(
display_name=content_name,
)
# -
# #### Option: Use a Previously Created Vertex Tensorboard Instance
#
# ```
# tensorboard_name = "Your Tensorboard Resource Name or Tensorboard ID"
# tensorboard = aiplatform.Tensorboard(tensorboard_name=tensorboard_name)
# ```
# ### Run a Vertex SDK CustomContainerTrainingJob
# + pycharm={"name": "#%%\n"}
display_name = content_name
gcs_output_uri_prefix = f"{BUCKET_NAME}/{display_name}"
replica_count = 4
machine_type = "n1-standard-4"
accelerator_count = 1
accelerator_type = "NVIDIA_TESLA_K80"
args = [
'--backend', 'nccl',
'--batch-size', '128',
'--epochs', '25',
]
# + pycharm={"name": "#%%\n"}
custom_container_training_job = aiplatform.CustomContainerTrainingJob(
display_name=display_name,
container_uri=custom_container_image_uri,
)
# + pycharm={"name": "#%%\n"}
custom_container_training_job.run(
args=args,
base_output_dir=gcs_output_uri_prefix,
machine_type=machine_type,
accelerator_count=accelerator_count,
accelerator_type=accelerator_type,
tensorboard=tensorboard.resource_name,
service_account=SERVICE_ACCOUNT,
)
# + pycharm={"name": "#%%\n"}
print(f'Custom Training Job Name: {custom_container_training_job.resource_name}')
print(f'GCS Output URI Prefix: {gcs_output_uri_prefix}')
# -
# ### Training Output Artifact
# + pycharm={"name": "#%%\n"}
# ! gsutil ls $gcs_output_uri_prefix
# -
# ## Clean Up Artifact
# + pycharm={"name": "#%%\n"}
# ! gsutil rm -rf $gcs_output_uri_prefix
| community-content/pytorch_image_classification_distributed_data_parallel_training_with_vertex_sdk/multi_node_ddp_nccl_vertex_training_with_custom_container.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Simpler Returns - Part I
# Download the data for Microsoft (‘MSFT’) from IEX for the period ‘2015-1-1’ until today.
# Apply the .**head()** and **.tail()** methods to check if the data is ok. Always pay attention to the dates. Try to get an idea about how the stock price changed during the period.
# ### Simple Rate of Return
# Calculate the simple returns of ‘MSFT’ for the given timeframe.
# $$
# \frac{P_1 - P_0}{P_0} = \frac{P_1}{P_0} - 1
# $$
| Python for Finance - Code Files/63 Simple Returns - Part I/Online Financial Data (APIs)/Python 3 APIs/Simple Returns - Part I - Exercise_IEX.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Challenge - Validating a Linear Regression
# ###### Import modules and ignore harmless seaborn warnings
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from sklearn.model_selection import cross_val_score
from sklearn import linear_model
from sklearn import preprocessing as preproc
import statsmodels.formula.api as smf
from statsmodels.sandbox.regression.predstd import wls_prediction_std
# %matplotlib inline
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
# ###### Build DataFrames from csv files
# +
ma_2013 = 'https://raw.githubusercontent.com/djrgit/coursework/master/thinkful/data_science/my_progress/unit_2_supervised_learning/table_8_offenses_known_to_law_enforcement_massachusetts_by_city_2013.csv'
nj_2013 = 'https://raw.githubusercontent.com/djrgit/coursework/master/thinkful/data_science/my_progress/unit_2_supervised_learning/table_8_offenses_known_to_law_enforcement_new_jersey_by_city_2013.csv'
ny_2013 = 'https://raw.githubusercontent.com/djrgit/coursework/master/thinkful/data_science/my_progress/unit_2_supervised_learning/table_8_offenses_known_to_law_enforcement_new_york_by_city_2013.csv'
ny_2014 = 'https://raw.githubusercontent.com/djrgit/coursework/master/thinkful/data_science/my_progress/unit_2_supervised_learning/Table_8_Offenses_Known_to_Law_Enforcement_by_New_York_by_City_2014.csv'
# 2013 - Massachusetts
df_ma_2013 = pd.read_csv(ma_2013, skiprows=59)
df_ma_2013 = df_ma_2013[:279]
# 2013 - New Jersey
df_nj_2013 = pd.read_csv(nj_2013, skiprows=59)
df_nj_2013 = df_nj_2013[:479]
# 2013 - New York
df_ny_2013 = pd.read_csv(ny_2013, skiprows=59)
df_ny_2013 = df_ny_2013[:348]
# 2014 - New York
df_ny_2014 = pd.read_csv(ny_2014, skiprows=4)
df_ny_2014 = df_ny_2014[:369]
# -
dfs = [df_ma_2013, df_nj_2013, df_ny_2013, df_ny_2014]
# ###### Preview the DataFrames before cleaning
df_ma_2013.head()
df_nj_2013.head()
df_ny_2013.head()
df_ny_2014.head()
df_ma_2013.info()
df_nj_2013.info()
df_ny_2013.info()
df_ny_2014.info()
# ###### Clean the data
def floatify(x):
try:
if ',' in str(x):
x = x.replace(',', '')
x = float(x)
except ValueError:
x = None
return x
def clean_df(df):
# Rename columns
df.columns = ['City', 'Population', 'Violent_crime', 'Murder', 'Rape_def1',
'Rape_def2', 'Robbery', 'Agg_assault', 'Prop_crime', 'Burglary',
'Larceny_theft', 'Mot_vehicle_theft', 'Arson']
# Ensure numerical data is treated as such
for col in df.columns:
if col != 'City':
df[col] = df[col].apply(floatify)
return df
for df in dfs:
clean_df(df)
# Drop columns for multiple states/years with many null values
df_ma_2013 = df_ma_2013.drop(columns=['Rape_def1', 'Rape_def2'])
df_nj_2013 = df_nj_2013.drop(columns=['Rape_def1', 'Rape_def2'])
df_ny_2013 = df_ny_2013.drop(columns=['Rape_def1', 'Rape_def2'])
df_ny_2014 = df_ny_2014.drop(columns=['Rape_def1', 'Rape_def2'])
# ###### Gather some basic statistical data for columns with numerical data
df_ma_2013.describe()
df_nj_2013.describe()
df_ny_2013.describe()
df_ny_2014.describe()
df_ma_2013 = df_ma_2013[(df_ma_2013['Population'] < 250000) & (df_ma_2013['Prop_crime'] > 0)]
df_nj_2013 = df_nj_2013[(df_nj_2013['Population'] < 250000) & (df_nj_2013['Prop_crime'] > 0)]
df_ny_2013 = df_ny_2013[(df_ny_2013['Population'] < 250000) & (df_ny_2013['Prop_crime'] > 0)]
df_ny_2014 = df_ny_2014[(df_ny_2014['Population'] < 250000) & (df_ny_2014['Prop_crime'] > 0)]
sns.set_style('darkgrid')
# ###### How does property crime vary with population (in cities with fewer than 250,000 people)?
# +
# Making a four-panel plot.
fig = plt.figure(figsize=(16,16))
fig.add_subplot(221)
sns.scatterplot(x='Population', y='Prop_crime', data=df_ma_2013)
plt.title('Property Crime Relative to Population\nMassachusetts (2013)')
plt.xlabel('Population')
plt.ylabel('Property Crime')
fig.add_subplot(222)
sns.scatterplot(x='Population', y='Prop_crime', data=df_nj_2013)
plt.title('Property Crime Relative to Population\nNew Jersey (2013)')
plt.xlabel('Population')
plt.ylabel('Property Crime')
fig.add_subplot(223)
sns.scatterplot(x='Population', y='Prop_crime', data=df_ny_2013)
plt.title('Property Crime Relative to Population\nNew York (2013)')
plt.xlabel('Population')
plt.ylabel('Property Crime')
fig.add_subplot(224)
sns.scatterplot(x='Population', y='Prop_crime', data=df_ny_2014)
plt.title('Property Crime Relative to Population\nNew York (2014)')
plt.xlabel('Population')
plt.ylabel('Property Crime')
plt.tight_layout()
plt.show()
# -
# ###### How does murder vary with population (in cities with fewer than 250,000 people)?
# +
# Making a four-panel plot.
fig = plt.figure(figsize=(16,16))
fig.add_subplot(221)
sns.scatterplot(x='Population', y='Murder', data=df_ma_2013)
plt.title('Murder Relative to Population\nMassachusetts (2013)')
plt.xlabel('Population')
plt.ylabel('Property Crime')
fig.add_subplot(222)
sns.scatterplot(x='Population', y='Murder', data=df_nj_2013)
plt.title('Murder Relative to Population\nNew Jersey (2013)')
plt.xlabel('Population')
plt.ylabel('Property Crime')
fig.add_subplot(223)
sns.scatterplot(x='Population', y='Murder', data=df_ny_2013)
plt.title('Murder Relative to Population\nNew York (2013)')
plt.xlabel('Population')
plt.ylabel('Property Crime')
fig.add_subplot(224)
sns.scatterplot(x='Population', y='Murder', data=df_ny_2014)
plt.title('Murder Relative to Population\nNew York (2014)')
plt.xlabel('Population')
plt.ylabel('Property Crime')
plt.tight_layout()
plt.show()
# -
# ###### How does robbery vary with population (in cities with fewer than 250,000 people)?
# +
# Making a four-panel plot.
fig = plt.figure(figsize=(16,16))
fig.add_subplot(221)
sns.scatterplot(x='Population', y='Robbery', data=df_ma_2013)
plt.title('Robbery Relative to Population\nMassachusetts (2013)')
plt.xlabel('Population')
plt.ylabel('Property Crime')
fig.add_subplot(222)
sns.scatterplot(x='Population', y='Robbery', data=df_nj_2013)
plt.title('Robbery Relative to Population\nNew Jersey (2013)')
plt.xlabel('Population')
plt.ylabel('Property Crime')
fig.add_subplot(223)
sns.scatterplot(x='Population', y='Robbery', data=df_ny_2013)
plt.title('Robbery Relative to Population\nNew York (2013)')
plt.xlabel('Population')
plt.ylabel('Property Crime')
fig.add_subplot(224)
sns.scatterplot(x='Population', y='Robbery', data=df_ny_2014)
plt.title('Robbery Relative to Population\nNew York (2014)')
plt.xlabel('Population')
plt.ylabel('Property Crime')
plt.tight_layout()
plt.show()
# -
# ###### Attempt to transform the property crime data for various states/years into a more normal distribution
# +
# Making a four-panel plot.
fig = plt.figure(figsize=(16,16))
fig.add_subplot(221)
plt.hist(np.log(df_ma_2013['Prop_crime']))
plt.title('Histogram\nLog of Property Crime in Massachusetts (2013)')
fig.add_subplot(222)
plt.hist(np.log(df_nj_2013['Prop_crime']))
plt.title('Histogram\nLog of Property Crime in New Jersey (2013)')
fig.add_subplot(223)
plt.hist(np.log(df_ny_2013['Prop_crime']))
plt.title('Histogram\nLog of Property Crime in New York (2013)')
fig.add_subplot(224)
plt.hist(np.log(df_ny_2014['Prop_crime']))
plt.title('Histogram\nLog of Property Crime in New York (2014)')
plt.tight_layout()
plt.show()
# +
# Making a four-panel plot.
fig = plt.figure(figsize=(16,16))
fig.add_subplot(221)
plt.hist(np.log(df_ma_2013['Population']))
plt.title('Histogram\nLog of Population in Massachusetts (2013)')
fig.add_subplot(222)
plt.hist(np.log(df_nj_2013['Population']))
plt.title('Histogram\nLog of Population in New Jersey (2013)')
fig.add_subplot(223)
plt.hist(np.log(df_ny_2013['Population']))
plt.title('Histogram\nLog of Population in New York (2013)')
fig.add_subplot(224)
plt.hist(np.log(df_ny_2014['Population']))
plt.title('Histogram\nLog of Population in New York (2014)')
plt.tight_layout()
plt.show()
# -
# ###### Construct model template
def build_model_df(df):
model = df[['Prop_crime', 'Population']].copy()
model['log_Prop_crime'] = np.log(model['Prop_crime'])
model['log_Pop'] = np.log(model['Population'])
model['Population^2'] = df['Population']**2
model['minmax_Burglary'] = preproc.minmax_scale(df[['Burglary']])
model['minmax_Murder'] = preproc.minmax_scale(df[['Murder']])
model['minmax_Robbery'] = preproc.minmax_scale(df[['Robbery']])
model['minmax_Larceny_theft'] = preproc.minmax_scale(df[['Larceny_theft']])
return model
# ###### Validate linear regression model
def validate_linear_reg(model, Xs, linear_formula):
# Instantiate and fit our model.
regr = linear_model.LinearRegression()
y = model['log_Prop_crime'].values.reshape(-1, 1)
X = model[Xs]
regr.fit(X, y)
# Inspect the results.
#print('\nCoefficients: \n', regr.coef_)
#print('\nIntercept: \n', regr.intercept_)
#print('\nR-squared:')
#print(regr.score(X, y))
# Extract predicted values.
predicted = regr.predict(X).ravel()
actual = model['log_Prop_crime']
# Calculate the error, also called the residual.
residual = actual - predicted
plt.hist(residual)
plt.title('Residual counts')
plt.xlabel('Residual')
plt.ylabel('Count')
plt.show()
# Scedasticity
plt.scatter(predicted, residual)
plt.xlabel('Predicted')
plt.ylabel('Residual')
plt.axhline(y=0)
plt.title('Residual vs. Predicted')
plt.show()
# Fit the model to our data using the formula.
lm = smf.ols(formula=linear_formula, data=model).fit()
print('Parameters (Intercept and Coefficients): ')
print(lm.params)
print('\n')
print('p-values: ')
print(lm.pvalues)
print('\n')
print('R-Squared: ')
print(lm.rsquared)
print('\n')
print('Confidence Intervals: ')
print(lm.conf_int())
print('\n')
print('Cross validation scores (5 folds): ')
print(cross_val_score(regr, X, y, cv=5))
X_params = ['log_Pop', 'minmax_Burglary', 'minmax_Murder', 'minmax_Robbery', 'minmax_Larceny_theft']
# Write out the model formula.
# Your dependent variable on the right, independent variables on the left
# Use a ~ to represent an '=' from the functional form
lin_form = 'log_Prop_crime ~ ' + '+'.join(X_params)
# ###### Test validation function with DataFrames
print('Massachusetts - 2013')
validate_linear_reg(build_model_df(df_ma_2013), X_params, lin_form)
print('New Jersey - 2013')
validate_linear_reg(build_model_df(df_nj_2013), X_params, lin_form)
print('New York - 2013')
validate_linear_reg(build_model_df(df_ny_2013), X_params, lin_form)
print('New York - 2014')
validate_linear_reg(build_model_df(df_ny_2014), X_params, lin_form)
# ### The p-values for 'minmax_Burglary' and 'minmax_Murder' are consistently higher than 0.05 for all datasets tested with this model, indicating that their inclusion in the model may not have much statistical significance with regards to predicting property crimes. Additionally, the confidence intervals for these parameters also cross 0.0 in _all_ tested datasets.
#
# ### These parameters may not add much to the model's explanatory power and can probably be removed. Follow-up with this hypothesis by testing the new model (without the minmax_Burglary and minmax_Murder parameters) below.
# Removed 'minmax_Burglary' and 'minmax_Murder'
X_params2 = ['log_Pop', 'minmax_Robbery', 'minmax_Larceny_theft']
# Write out the model formula.
# Your dependent variable on the right, independent variables on the left
# Use a ~ to represent an '=' from the functional form
lin_form2 = 'log_Prop_crime ~ ' + '+'.join(X_params2)
print('Massachusetts - 2013')
validate_linear_reg(build_model_df(df_ma_2013), X_params2, lin_form2)
print('New Jersey - 2013')
validate_linear_reg(build_model_df(df_nj_2013), X_params2, lin_form2)
print('New York - 2013')
validate_linear_reg(build_model_df(df_ny_2013), X_params2, lin_form2)
print('New York - 2013')
validate_linear_reg(build_model_df(df_ny_2014), X_params2, lin_form2)
| thinkful/data_science/my_progress/unit_2_supervised_learning/Unit_2_-_Lesson_5_-_Challenge_-_Validating_a_Linear_Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="2qGrxQb-6gkn"
# %matplotlib inline
import torch
from torch import nn
from torch.nn import functional as F
import torchvision
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import random
import os
# -
# ## Model Architecture
# + colab={} colab_type="code" id="kgOEDAI46l5m"
layer1out = 512
layer2out = 256
layer3out = 128
layer4out = 64
z_size = 100
class Generator(nn.Module):
def __init__(self):
super(Generator, self).__init__()
self.conv1 = nn.ConvTranspose2d(z_size,layer1out,4,1,0,bias=False)
self.conv2 = nn.ConvTranspose2d(layer1out,layer2out,4,2,1,bias=False)
self.conv3 = nn.ConvTranspose2d(layer2out,layer3out,4,2,1,bias=False)
self.conv4 = nn.ConvTranspose2d(layer3out,layer4out,4,2,1,bias=False)
self.conv5 = nn.ConvTranspose2d(layer4out,1,4,2,1,bias=False)
self.leaky = nn.LeakyReLU()
self.bn1 = nn.BatchNorm2d(layer1out)
self.bn2 = nn.BatchNorm2d(layer2out)
self.bn3 = nn.BatchNorm2d(layer3out)
self.bn4 = nn.BatchNorm2d(layer4out)
self.tanh = nn.Tanh()
def forward(self, x):
x = self.leaky(self.bn1(self.conv1(x)))
x = self.leaky(self.bn2(self.conv2(x)))
x = self.leaky(self.bn3(self.conv3(x)))
x = self.leaky(self.bn4(self.conv4(x)))
x = self.tanh(self.conv5(x))
return x
# + colab={} colab_type="code" id="FtoaRDhAtGaU"
layer0out = 64
layer1out = 128
layer2out = 256
layer3out = 512
class Discriminator(nn.Module):
def __init__(self):
super(Discriminator, self).__init__()
self.conv0 = nn.Conv2d(1,layer0out,4,2,1,bias=False)
self.conv1 = nn.Conv2d(layer0out,layer1out,4,2,1,bias=False)
self.conv2 = nn.Conv2d(layer1out,layer2out,4,2,1,bias=False)
self.conv3 = nn.Conv2d(layer2out,layer3out,4,2,1,bias=False)
self.conv4 = nn.Conv2d(layer3out,1,4,1,0,bias=False)
self.bn0 = nn.BatchNorm2d(layer0out)
self.bn1 = nn.BatchNorm2d(layer1out)
self.bn2 = nn.BatchNorm2d(layer2out)
self.bn3 = nn.BatchNorm2d(layer3out)
def forward(self, x):
x = F.leaky_relu(self.conv0(x))
x = F.leaky_relu(self.bn1(self.conv1(x)))
x = F.leaky_relu(self.bn2(self.conv2(x)))
x = F.leaky_relu(self.bn3(self.conv3(x)))
x = torch.sigmoid(self.conv4(x))
return x
# + colab={} colab_type="code" id="3PwxH4o8N1l0"
def weights_init(m):
classname = m.__class__.__name__
if classname.find('Conv') != -1:
nn.init.normal_(m.weight.data,0.0,0.02)
elif classname.find('BatchNorm') != -1:
nn.init.normal_(m.weight.data, 1.0, 0.02)
nn.init.constant_(m.bias.data,0)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="yY9eqG0Rr5EH" outputId="9bc29e7b-53bd-4e11-e39b-2b37dcd640e5"
torch.backends.cudnn.enabled = False
d = Discriminator()
g = Generator()
d.apply(weights_init)
g.cuda()
d.cuda()
# + colab={"base_uri": "https://localhost:8080/", "height": 122} colab_type="code" id="ZbFCSUb6SwXT" outputId="aa204fb5-7bd2-45c5-d061-d7f12dfb62a1"
"""from google.colab import drive
drive.mount('/content/gdrive')
"""
FILE = './Images/'
size = 64
# + colab={} colab_type="code" id="L7ntnFT9NZ_Z"
batch_size = 32
dataset = torchvision.datasets.ImageFolder(root=FILE,
transform=torchvision.transforms.Compose([
torchvision.transforms.Grayscale(),
torchvision.transforms.Resize(size),
torchvision.transforms.CenterCrop(size),
torchvision.transforms.ToTensor(),
torchvision.transforms.Normalize((0.5,0.5,0.5),(0.5,0.5,0.5))]))
dataloader = torch.utils.data.DataLoader(dataset, batch_size=batch_size,shuffle=True)
device = torch.device('cuda:0')
# + colab={} colab_type="code" id="p_lOVaHHVtSP"
lr1 = 0.0002
lr2 = 0.0002
crit = nn.BCELoss()
optd = torch.optim.Adam(d.parameters(),lr=lr1,betas=(0.5,0.999))
optg = torch.optim.Adam(g.parameters(),lr=lr2,betas=(0.5,0.999))
# -
# ## Training
# + colab={"base_uri": "https://localhost:8080/", "height": 708} colab_type="code" id="fNI57pLzs14h" outputId="99e1c219-f2a3-4c20-90c8-3590d8ba43bb"
iters=0
epochs = 500
checkpoint = 246
log_list =[]
for epoch in range(checkpoint, epochs):
for i, data in enumerate(dataloader, 0):
# Discriminator
# real
d.zero_grad()
inps = data[0].to(device)
labels = torch.full((inps.shape[0],),random.uniform(0.9,1),device=device)
output = d(inps).view(-1)
loss_dis_real = crit(output,labels)
loss_dis_real.backward()
# fake
fake_inps = g(torch.randn(inps.shape[0],100,1,1,device=device))
labels.fill_(0)
output = d(fake_inps.detach()).view(-1)
loss_dis_fake = crit(output,labels)
loss_dis_fake.backward()
loss_dis = loss_dis_real + loss_dis_fake
optd.step()
# Generator
g.zero_grad()
labels.fill_(1)
output = d(fake_inps).view(-1)
loss_gen = crit(output,labels)
loss_gen.backward()
optg.step()
if i % 100 ==0:
log_text='Epoch: ' + str(epoch) + '\tloss_dis:' + str(float(loss_dis)) + '\tloss_gen: ' + str(float(loss_gen))
print(log_text)
log_list.append(log_text)
log_list.append('\n')
if epoch % 20 == 0:
f = open("log.txt",'w')
f.writelines(log_list)
log_list=[]
sample_pics = g(torch.randn(10,100,1,1,device=device))
for i,pic in enumerate(sample_pics):
plt.imsave('e'+str(epoch)+'_'+str(i),pic.cpu().detach().numpy().reshape(64,64),cmap='gray')
f.close()
# -
torch.save(d,'discrim')
torch.save(g,'generat')
# ## Generating Results
d = torch.load('discrim', map_location='cpu')
g = torch.load('generat', map_location='cpu')
#
d.zero_grad()
g.zero_grad()
generated = g(torch.randn(1000,100,1,1,device='cpu'))
out = d(generated).view(-1)
losses = []
for score in out:
losses.append(crit(score, torch.full((1,),1,device='cpu')))
sorted_loss_inds = np.argsort(losses)
best_images = generated[sorted_loss_inds][:100]
# From these results we see that there are a lot of well generated pictures such as the following.
plt.imshow(best_images[7].detach().numpy().reshape(64,64),cmap='gray')
plt.imshow(best_images[11].detach().numpy().reshape(64,64),cmap='gray')
plt.imshow(best_images[4].detach().numpy().reshape(64,64),cmap='gray')
plt.imshow(best_images[58].detach().numpy().reshape(64,64),cmap='gray')
# It seems the model has also undergone mode collapse as well with this recurring picture:
plt.imshow(best_images[12].detach().numpy().reshape(64,64),cmap='gray')
plt.imshow(best_images[20].detach().numpy().reshape(64,64),cmap='gray')
# Looking past the nightmare inducing features we're able to see that the recurring picture has the same facial structures however with bits and pieces of different dogs implanted. Note how the first dog's right eye is similar to that of a Maltese dog 
#
# while the second seems to be a different dog entirely.
# Another type of image generated include incomplete dogs:
plt.imshow(best_images[0].detach().numpy().reshape(64,64),cmap='gray')
plt.imshow(best_images[1].detach().numpy().reshape(64,64),cmap='gray')
plt.imshow(best_images[2].detach().numpy().reshape(64,64),cmap='gray')
# Further development of the model requires higher care and a more watchful eye over the outputs to prevent mode collapse. THe incompleteness of some of the dogs can be fixed with a longer run time as the losses converge further.
| dcgan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Given two strings S and T, return if they are equal when both are typed into empty text editors. # means a backspace character.
#
# Note that after backspacing an empty text, the text will continue empty.
#
# <b>Example 1:</b>
#
# Input: S = "ab#c", T = "ad#c"<br>
# Output: true<br>
# Explanation: Both S and T become "ac".<br>
# <b>Example 2:<br></b>
#
# Input: S = "ab##", T = "c#d#"<br>
# Output: true<br>
# Explanation: Both S and T become "".<br>
# <b>Example 3:<br></b>
#
# Input: S = "a##c", T = "#a#c"<br>
# Output: true<br>
# Explanation: Both S and T become "c".<br>
# <b>Example 4:<br></b>
#
# Input: S = "a#c", T = "b"<br>
# Output: false<br>
# Explanation: S becomes "c" while T becomes "b".<br>
# Note:
#
# 1 <= S.length <= 200<br>
# 1 <= T.length <= 200<br>
# S and T only contain lowercase letters and '#' characters.<br>
# Follow up:
#
# Can you solve it in O(N) time and O(1) space?
# ### Solution
# * For this solution stack is a good data structure.<br>
# * We need to simulate the result of every key stroke
# * If the character is not a # push to the stack else if stack is not empty pop from the stack
#
def backspace_compare(s,t):
def build(s):
stack=[]
for ch in s:
if ch!='#':
stack.append(ch)
elif stack !=[]:
stack.pop()
return "".join(stack)
return build(s)==build(t)
print(backspace_compare("ab#c","ad#c"))
print(backspace_compare("ab##","c#d#"))
print(backspace_compare("a##c","#a#c"))
print(backspace_compare("a#c","b"))
# ### Complexity
# #### Time complexity : O(M+N) , where m,n are the lengths of s and t
# #### Space complexity : O(M+N)
# 
def backspace_compare(s,t):
def build(s):
stack=[]
for ch in s:
print("\ncharac",ch)
if ch!='#':
stack.append(ch)
elif stack !=[]:
stack.pop()
print("stack",stack)
return "".join(stack)
return build(s)==build(t)
print(backspace_compare("ab#c","ad#c"))
| array_strings/ipynb/backspace_string_compare.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="JBMdxhsVohF2"
import pandas as pd
from sklearn.cluster import KMeans
import altair as alt
# + colab={} colab_type="code" id="iSwgbS3CooqD"
file_url = 'https://raw.githubusercontent.com/PacktWorkshops/The-Data-Science-Workshop/master/Chapter05/DataSet/taxstats2015.csv'
# + colab={} colab_type="code" id="91P_EE8NpNHg"
df = pd.read_csv(file_url, usecols=['Postcode', 'Average total business income', 'Average total business expenses'])
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="YTXNZwfYpQkg" outputId="6b1ab7f7-cc4c-434e-c482-db4063c6d68d"
df.head()
# + colab={} colab_type="code" id="86yRMdUtpgnI"
X = df[['Average total business income', 'Average total business expenses']]
# + colab={} colab_type="code" id="tU9JPKJJpzBg"
clusters = pd.DataFrame()
inertia = []
# + colab={} colab_type="code" id="Hu0Dv2tzqEB0"
clusters['cluster_range'] = range(1, 15)
# + colab={} colab_type="code" id="RlEkLMUkqsbq"
for k in clusters['cluster_range']:
kmeans = KMeans(n_clusters=k).fit(X)
inertia.append(kmeans.inertia_)
# + colab={"base_uri": "https://localhost:8080/", "height": 483} colab_type="code" id="slnfmG3Mqtyy" outputId="7ea3a4ec-e566-44d7-d48a-d2602d33feb9"
clusters['inertia'] = inertia
clusters
# + colab={"base_uri": "https://localhost:8080/", "height": 368} colab_type="code" id="WHUQBZ8uqvly" outputId="195438e3-b79a-41f8-afd5-e2430070aed9"
alt.Chart(clusters).mark_line().encode(alt.X('cluster_range'), alt.Y('inertia'))
# + colab={} colab_type="code" id="CUMT2XeIrIpf"
optim_cluster = 4
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="UTb91dx0rsqD" outputId="f84bdf2a-e092-46ff-cc4e-baddebcc2ccb"
kmeans = KMeans(random_state=42, n_clusters=optim_cluster)
kmeans.fit(X)
# + colab={} colab_type="code" id="4Zf9IGEsruV8"
df['cluster2'] = kmeans.predict(X)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="LfxHSNcUrzA0" outputId="c220d3eb-6145-4bcf-b53e-e4e0b35c7b11"
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 368} colab_type="code" id="JM-UjFvyr73R" outputId="c05e1733-37ea-4174-b279-ea028639d185"
alt.Chart(df).mark_circle().encode(x='Average total business income', y='Average total business expenses',color='cluster2:N', tooltip=['Postcode', 'cluster2', 'Average total business income', 'Average total business expenses']).interactive()
# + colab={} colab_type="code" id="aGMIPBYOscBQ"
| Chapter05/Exercise5.03/Exercise5_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python38-azureml
# kernelspec:
# display_name: Python 3.8 - AzureML
# language: python
# name: python38-azureml
# ---
# # How To: Adding Hunting Bookmarks from Notebooks
#
# __Notebook Version:__ 1.0<br>
# __Python Version:__ Python 3.8 - AzureML<br>
# __Platforms Supported:__<br>
# - Azure ML
# __Data Source Required:__<br>
# - no
#
# ### Description
# The sample notebook shows how to add hunting bookmarks to Azure Sentinel through Jupyter notebooks.
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1619122065407}
# Parameters for notebooks testing, can be ignored safely
test_run = False
# + gather={"logged": 1619122066467}
# Loading Python libraries
from azure.common.credentials import get_azure_cli_credentials
import requests
import json
import uuid
import pandas
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1619122067789}
# Functions will be used in this notebook
def read_config_values(file_path):
"This loads pre-generated parameters for Sentinel Workspace"
with open(file_path) as json_file:
if json_file:
json_config = json.load(json_file)
return (json_config["tenant_id"],
json_config["subscription_id"],
json_config["resource_group"],
json_config["workspace_id"],
json_config["workspace_name"])
return None
# Calling Sentinel API, the same template can be used for calling other Azure REST APIs with different parameters.
# For different environments, such as national clouds, you may need to use different root_url, please contact with your admins.
# It can be ---.azure.us, ---.azure.microsoft.scloud, ---.azure.eaglex.ic.gov, etc.
def call_azure_rest_api(token, resource_name, request_body, bookmark_id, api_version):
"Calling Sentinel REST API"
headers = {"Authorization": token, "content-type":"application/json" }
provider_name = "Microsoft.OperationalInsights"
provider2_name = "Microsoft.SecurityInsights"
target_resource_name = resource_name
api_version = api_version
root_url = "https://management.azure.com"
arm_rest_url_template = "{0}/subscriptions/{1}/resourceGroups/{2}/providers/{3}/workspaces/{4}/providers/{5}/{6}/{7}?api-version={8}"
arm_rest_url = arm_rest_url_template.format(root_url, subscription_id, resource_group, provider_name, workspace_name, provider2_name, target_resource_name, bookmark_id, api_version)
print(arm_rest_url)
response = requests.put(arm_rest_url, headers=headers, data=request_body)
return response
def display_result_name(response):
"Default to display column - name, you may change it to other columns"
column_name = "name"
if response != None:
entries = [item[column_name] for item in response.json()["value"]]
display(entries)
def display_result(response):
"Display the result set as pandas.DataFrame"
if response != None:
df = pandas.DataFrame(response.json()["value"])
display(df)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1619122070125}
# Calling the above function to populate Sentinel workspace parameters
# The file, config.json, was generated by the system, however, you may modify the values, or manually set the variables
tenant_id, subscription_id, resource_group, workspace_id, workspace_name = read_config_values('config.json');
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1619122091167}
# Azure CLI is used to get device code to login into Azure, you need to copy the code and open the DeviceLogin site.
# You may add [--tenant $tenant_id] to the command
if test_run == False:
# !az login --tenant $tenant_id --use-device-code
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1619130034528}
# Extract access token, which will be used to access Sentinel Watchlist API for your Watchlist data.
credentials, sub_id = get_azure_cli_credentials()
creds = credentials._get_cred(resource=None)
token = creds._token_retriever()[2]
access_token = token['accessToken']
header_token_value = "Bearer {}".format(access_token)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1619130035411}
name = "Bookmark test from notebook"
query = "AzureActivity | where TimeGenerated < ago(5d)"
entity_mappings = {}
entity_mappings.update({'550a6d02-d667-49d8-969a-e709cce03293': 'Account'})
entity_mappings.update({'192.168.3.11': 'Host'})
entities = r"{\"550a6d02-d667-49d8-969a-e709cce03293\": \"Account\", \"192.168.3.11\": \"Host\"}"
query_result = r"{\"Value\":0,\"Time\":\"2020-03-22T16:46:20.006499Z\",\"Legend\":\"F5Telemetry_LTM_CL\",\"__entityMapping\":" + entities + "}"
payload_data = "{\"properties\": { \"displayName\": \"" + name + "\", \"notes\": \"Testing from notebook\", \"labels\": [\"test\"], \"query\": \"" + query + "\", \"queryResult\": \"" + query_result + "\" }}"
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1619129877775}
# Calling Sentinel Watchlist API
response_bookmark = call_azure_rest_api(header_token_value, "bookmarks", payload_data, str(uuid.uuid4()), "2020-01-01")
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1619130037840}
response_bookmark.text
| HowTos/Adding Hunting Bookmarks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import math
import keras
import tensorflow as tf
print(pd.__version__)
import progressbar
import os
from os import listdir
# ## Print Dependencies
#
#
#
# Dependences are fundamental to record the computational environment.
# +
# %load_ext watermark
# python, ipython, packages, and machine characteristics
# %watermark -v -m -p pandas,keras,numpy,math,tensorflow,matplotlib,h5py
# date
print (" ")
# %watermark -u -n -t -z
# -
# ## Load of the data
#
# You can also load all of them! Writing "all_data"
os.listdir('../data/classifier/')
n = input('Which file do you want?')
if (n != 'all_data'):
print(2)
from process import loaddata
if n != 'all_data':
class_data = loaddata("../data/classifier/{}.csv".format(n))
if n == 'all_data':
files = os.listdir('../data/classifier/')
for filename in files:
class_data = loaddata("../data/classifier/{}".format(filename))
np.random.shuffle(class_data)
y = class_data[:,0]
x = class_data[:,1:]
x.shape
train_split = 0.75
train_limit = int(len(y)*train_split)
print("Training sample: {0} \nValuation sample: {1}".format(train_limit, len(y)-train_limit))
# +
x_train = x[:train_limit]
x_val = x[train_limit:]
y_train = y[:train_limit]
y_val = y[train_limit:]
# -
# ## Model Build
from keras.models import Sequential
from keras.layers.core import Dense
import keras.backend as K
from keras import optimizers
from keras import models
from keras import layers
def build_model() :
model = models.Sequential()
model.add (layers.Dense (16 , activation = "relu" , input_dim = 10))
model.add (layers.Dense (16, activation = "relu"))
model.add (layers.Dense (1 , activation = "sigmoid"))
model.compile(optimizer = "adam" , loss = "binary_crossentropy" , metrics =["accuracy"])
return model
model = build_model ()
history = model.fit ( x_train, y_train, epochs = 1000, batch_size = 10000 , validation_data = (x_val, y_val) )
model.save("../models/classifier/{}_nodropout.h5".format(n))
model.summary()
# ## Adding Dropout
#
# Dropout is one of the most effective and most commonly used regularization techniques for neural networks, developed by <NAME> and his students at the University of Toronto. Dropout, applied to a layer, consists of randomly dropping out (setting to zero) a number of output features of the layer during training. Let’s say a
# given layer would normally return a vector [0.2, 0.5, 1.3, 0.8, 1.1] for a given input sample during training. After applying dropout, this vector will have a few zero entries distributed at random: for example, [0, 0.5, 1.3, 0, 1.1]. The dropout rate is the fraction of the features that are zeroed out; it’s usually set between 0.2 and 0.5. At test time, no units are dropped out; instead, the layer’s output values are scaled down by a factor equal to the dropout rate, to balance for the fact that more units are active than at training time.
#
# ##### [From Deep Learning with Python, Chollet]
# +
import matplotlib.pyplot as plt
loss = history.history['loss']
val_loss = history.history['val_loss']
accuracy = history.history['accuracy']
val_accuracy = history.history['val_accuracy']
epochs = range(1, len(loss) + 1)
fig, ax1 = plt.subplots()
l1 = ax1.plot(epochs, loss, 'bo', label='Training loss')
vl1 = ax1.plot(epochs, val_loss, 'b', label='Validation loss')
ax1.set_title('Training and validation loss')
ax1.set_xlabel('Epochs')
ax1.set_ylabel('Loss')
ax2 = ax1.twinx()
ac2= ax2.plot(epochs, accuracy, 'o', c="red", label='Training acc')
vac2= ax2.plot(epochs, val_accuracy, 'r', label='Validation acc')
ax2.set_ylabel('Accuracy')
lns = l1 + vl1 + ac2 + vac2
labs = [l.get_label() for l in lns]
ax2.legend(lns, labs, loc="center right")
fig.tight_layout()
fig.savefig("acc+loss_drop.pdf")
fig.show()
# +
plt.clf() # clear figure
accuracy = history.history['accuracy']
val_accuracy = history.history['val_accuracy']
plt.plot(epochs, accuracy, 'bo', label='Training acc')
plt.plot(epochs, val_accuracy, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# -
def build_model() :
model = models.Sequential()
model.add (layers.Dense (16 , activation = "relu" , input_shape = input_shape = (None,10)))
model.add (layers.Dense (16, activation = "relu"))
model.add (layers.Dense (1 , activation = "sigmoid"))
model.compile(optimizer = "adam" , loss = "binary_crossentropy" , metrics =["accuracy"])
return model
model = build_model ()
history = model.fit ( x_train, y_train, epochs = 1000, batch_size = 10000 , validation_data = (x_val, y_val) )
model.save("../models/classifier/{}_nodropout.h5".format(n))
model.summary()
# +
import matplotlib.pyplot as plt
loss = history.history['loss']
val_loss = history.history['val_loss']
accuracy = history.history['accuracy']
val_accuracy = history.history['val_accuracy']
epochs = range(1, len(loss) + 1)
fig, ax1 = plt.subplots()
l1 = ax1.plot(epochs, loss, 'bo', label='Training loss')
vl1 = ax1.plot(epochs, val_loss, 'b', label='Validation loss')
ax1.set_title('Training and validation loss')
ax1.set_xlabel('Epochs')
ax1.set_ylabel('Loss')
ax2 = ax1.twinx()
ac2= ax2.plot(epochs, accuracy, 'o', c="red", label='Training acc')
vac2= ax2.plot(epochs, val_accuracy, 'r', label='Validation acc')
ax2.set_ylabel('Accuracy')
lns = l1 + vl1 + ac2 + vac2
labs = [l.get_label() for l in lns]
ax2.legend(lns, labs, loc="center right")
fig.tight_layout()
fig.savefig("acc+loss_drop.pdf")
fig.show()
# +
plt.clf() # clear figure
accuracy = history.history['accuracy']
val_accuracy = history.history['val_accuracy']
plt.plot(epochs, accuracy, 'bo', label='Training acc')
plt.plot(epochs, val_accuracy, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# -
# ## Performance summary and dropout vs no dropout
class_dropout = keras.models.load_model('../models/classifier/{}_dropout.h5'.format(n))
class_nodropout = keras.models.load_model('../models/classifier/{}_nodropout.h5'.format(n))
class_dropout.fit(x_val, y_val)
class_nodropout.fit(x_val, y_val)
class_dropout.fit(x_train, y_train)
class_nodropout.fit(x_train, y_train)
# ## Probability density distribution
| notebooks/Comp_Scatt_NeuralNetwork_Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="2V_OLlgtQ0DW"
# # Ungraded Lab: TFDV Exercise
#
#
# In this notebook, you will get to practice using [TensorFlow Data Validation (TFDV)](https://cloud.google.com/solutions/machine-learning/analyzing-and-validating-data-at-scale-for-ml-using-tfx), an open-source Python package from the [TensorFlow Extended (TFX)](https://www.tensorflow.org/tfx) ecosystem.
#
# TFDV helps to understand, validate, and monitor production machine learning data at scale. It provides insight into some key questions in the data analysis process such as:
#
# * What are the underlying statistics of my data?
#
# * What does my training dataset look like?
#
# * How does my evaluation and serving datasets compare to the training dataset?
#
# * How can I find and fix data anomalies?
#
# The figure below summarizes the usual TFDV workflow:
#
# <img src='img/tfdv.png' alt='picture of tfdv workflow'>
#
# As shown, you can use TFDV to compute descriptive statistics of the training data and generate a schema. You can then validate new datasets (e.g. the serving dataset from your customers) against this schema to detect and fix anomalies. This helps prevent the different types of skew. That way, you can be confident that your model is training on or predicting data that is consistent with the expected feature types and distribution.
#
# This ungraded exercise demonstrates useful functions of TFDV at an introductory level as preparation for this week's graded programming exercise. Specifically, you will:
#
# - **Generate and visualize statistics from a dataset**
# - **Detect and fix anomalies in an evaluation dataset**
#
# Let's begin!
# + [markdown] id="lyGr4CC8AUfu"
# ## Package Installation and Imports
# + id="GHi-tkOeBOis"
import tensorflow as tf
import tensorflow_data_validation as tfdv
import pandas as pd
from sklearn.model_selection import train_test_split
from util import add_extra_rows
from tensorflow_metadata.proto.v0 import schema_pb2
print('TFDV Version: {}'.format(tfdv.__version__))
print('Tensorflow Version: {}'.format(tf.__version__))
# + [markdown] id="Kzt9rO9eAGF-"
# ## Download the dataset
#
# You will be working with the [Census Income Dataset](http://archive.ics.uci.edu/ml/datasets/Census+Income), a dataset that can be used to predict if an individual earns more than or less than 50k US Dollars annually. The summary of attribute names with descriptions/expected values is shown below and you can read more about it [in this data description file.](https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.names)
#
#
# * **age**: continuous.
# * **workclass**: Private, Self-emp-not-inc, Self-emp-inc, Federal-gov, Local-gov, State-gov, Without-pay, Never-worked.
# * **fnlwgt**: continuous.
# * **education**: Bachelors, Some-college, 11th, HS-grad, Prof-school, Assoc-acdm, Assoc-voc, 9th, 7th-8th, 12th, Masters, 1st-4th, 10th, Doctorate, 5th-6th, Preschool.
# * **education-num**: continuous.
# * **marital-status**: Married-civ-spouse, Divorced, Never-married, Separated, Widowed, Married-spouse-absent, Married-AF-spouse.
# * **occupation**: Tech-support, Craft-repair, Other-service, Sales, Exec-managerial, Prof-specialty, Handlers-cleaners, Machine-op-inspct, Adm-clerical, Farming-fishing, Transport-moving, Priv-house-serv, Protective-serv, Armed-Forces.
# * **relationship**: Wife, Own-child, Husband, Not-in-family, Other-relative, Unmarried.
# * **race**: White, Asian-Pac-Islander, Amer-Indian-Eskimo, Other, Black.
# * **sex**: Female, Male.
# * **capital-gain**: continuous.
# * **capital-loss**: continuous.
# * **hours-per-week**: continuous.
# * **native-country**: United-States, Cambodia, England, Puerto-Rico, Canada, Germany, Outlying-US(Guam-USVI-etc), India, Japan, Greece, South, China, Cuba, Iran, Honduras, Philippines, Italy, Poland, Jamaica, Vietnam, Mexico, Portugal, Ireland, France, Dominican-Republic, Laos, Ecuador, Taiwan, Haiti, Columbia, Hungary, Guatemala, Nicaragua, Scotland, Thailand, Yugoslavia, El-Salvador, Trinadad&Tobago, Peru, Hong, Holand-Netherlands.
#
# Let's load the dataset and split it into training and evaluation sets. We will not shuffle them for consistent results in this demo notebook but you should otherwise in real projects.
# + id="WKTfuT2rga-_"
# Read in the training and evaluation datasets
df = pd.read_csv('data/adult.data', skipinitialspace=True)
# Split the dataset. Do not shuffle for this demo notebook.
train_df, eval_df = train_test_split(df, test_size=0.2, shuffle=False)
# + [markdown] id="0RQvgiMRq0pn"
# Let's see the first few columns of the train and eval sets.
# + id="uxDFD6dR0PYH"
# Preview the train set
train_df.head()
# + id="yyrmQLCm0a5V"
# Preview the eval set
eval_df.head()
# + [markdown] id="12_uIoIjr9lv"
# From these few columns, you can get a first impression of the data. You will notice that most are strings and integers. There are also columns that are mostly zeroes. In the next sections, you will see how to use TFDV to aggregate and process this information so you can inspect it more easily.
# -
# ### Adding extra rows
#
# To demonstrate how TFDV can detect anomalies later, you will add a few extra rows to the evaluation dataset. These are either malformed or have values that will trigger certain alarms later in this notebook. The code to add these can be seen in the `add_extra_rows()` function of `util.py` found in your Jupyter workspace. You can look at it later and even modify it after you've completed the entire exercise. For now, let's just execute the function and add the rows that we've defined by default.
# +
# add extra rows
eval_df = add_extra_rows(eval_df)
# preview the added rows
eval_df.tail(4)
# + [markdown] id="Duwwrsvf_9bK"
# ## Generate and visualize training dataset statistics
# + [markdown] id="5Nm5E1HAgPU0"
# You can now compute and visualize the statistics of your training dataset. TFDV accepts three input formats: TensorFlow’s TFRecord, Pandas Dataframe, and CSV file. In this exercise, you will feed in the Pandas Dataframes you generated from the train-test split.
#
# You can compute your dataset statistics by using the [`generate_statistics_from_dataframe()`](https://www.tensorflow.org/tfx/data_validation/api_docs/python/tfdv/generate_statistics_from_dataframe) method. Under the hood, it distributes the analysis via [Apache Beam](https://beam.apache.org/) which allows it to scale over large datasets.
#
# The results returned by this step for numerical and categorical data are summarized in this table:
#
# | Numerical Data | Categorical Data |
# |:-:|:-:|
# |Count of data records|Count of data records
# |% of missing data records|% of missing data records|
# |Mean, std, min, max|unique records|
# |% of zero values|Avg string length|
#
# + id="_vTx9Qkk4yGc"
# Generate training dataset statistics
train_stats = tfdv.generate_statistics_from_dataframe(train_df)
# + [markdown] id="86nhDglwuilJ"
# Once you've generated the statistics, you can easily visualize your results with the [`visualize_statistics()`](https://www.tensorflow.org/tfx/data_validation/api_docs/python/tfdv/visualize_statistics) method. This shows a [Facets interface](https://pair-code.github.io/facets/) and is very useful to spot if you have a high amount of missing data or high standard deviation. Run the cell below and explore the different settings in the output interface (e.g. Sort by, Reverse order, Feature search).
# + id="1D1wP3mm5ebW"
# Visualize training dataset statistics
tfdv.visualize_statistics(train_stats)
# + [markdown] id="DVQTOBpdgPU0"
# ## Infer data schema
# + [markdown] id="Ya6ecHE9gPU1"
# Next step is to create a data schema to describe your train set. Simply put, a schema describes standard characteristics of your data such as column data types and expected data value range. The schema is created on a dataset that you consider as reference, and can be reused to validate other incoming datasets.
#
# With the computed statistics, TFDV allows you to automatically generate an initial version of the schema using the [`infer_schema()`](https://www.tensorflow.org/tfx/data_validation/api_docs/python/tfdv/infer_schema) method. This returns a Schema [protocol buffer](https://developers.google.com/protocol-buffers) containing the result. As mentioned in the [TFX paper](http://stevenwhang.com/tfx_paper.pdf) (Section 3.3), the results of the schema inference can be summarized as follows:
#
# * The expected type of each feature.
# * The expected presence of each feature, in terms of a minimum count and fraction of examples that must contain
# the feature.
# * The expected valency of the feature in each example, i.e.,
# minimum and maximum number of values.
# * The expected domain of a feature, i.e., the small universe of
# values for a string feature, or range for an integer feature.
#
# Run the cell below to infer the training dataset schema.
# + id="W9skjM-M44Jz"
# Infer schema from the computed statistics.
schema = tfdv.infer_schema(statistics=train_stats)
# Display the inferred schema
tfdv.display_schema(schema)
# + [markdown] id="5oj_GIprgPU1"
# ## Generate and visualize evaluation dataset statistics
# + [markdown] id="rTYMPukogPU1"
# The next step after generating the schema is to now look at the evaluation dataset. You will begin by computing its statistics then compare it with the training statistics. It is important that the numerical and categorical features of the evaluation data belongs roughly to the same range as the training data. Otherwise, you might have distribution skew that will negatively affect the accuracy of your model.
#
# TFDV allows you to generate both the training and evaluation dataset statistics side-by-side. You can use the [`visualize_statistics()`](https://www.tensorflow.org/tfx/data_validation/api_docs/python/tfdv/visualize_statistics) function and pass additional parameters to overlay the statistics from both datasets (referenced as left-hand side and right-hand side statistics). Let's see what these parameters are:
#
# - `lhs_statistics`: Required parameter. Expects an instance of `DatasetFeatureStatisticsList `.
#
#
# - `rhs_statistics`: Expects an instance of `DatasetFeatureStatisticsList ` to compare with `lhs_statistics`.
#
#
# - `lhs_name`: Name of the `lhs_statistics` dataset.
#
#
# - `rhs_name`: Name of the `rhs_statistics` dataset.
# + id="bzZy1x3c6Mi0"
# Generate evaluation dataset statistics
eval_stats = tfdv.generate_statistics_from_dataframe(eval_df)
# Compare training with evaluation
tfdv.visualize_statistics(
lhs_statistics=eval_stats,
rhs_statistics=train_stats,
lhs_name='EVAL_DATASET',
rhs_name='TRAIN_DATASET'
)
# + [markdown] id="GODDgoHdgPU2"
# We encourage you to observe the results generated and toggle the menus to practice manipulating the visualization (e.g. sort by missing/zeroes). You'll notice that TFDV detects the malformed rows we introduced earlier. First, the `min` and `max` values of the `age` row shows `0` and `1000`, respectively. We know that those values do not make sense if we're talking about working adults. Secondly, the `workclass` row in the Categorical Features says that `0.02%` of the data is missing that particular attribute. Let's drop these rows to make the data more clean.
# +
# filter the age range
eval_df = eval_df[eval_df['age'] > 16]
eval_df = eval_df[eval_df['age'] < 91]
# drop missing values
eval_df.dropna(inplace=True)
# -
# You can then compute the statistics again and see the difference in the results.
# +
# Generate evaluation dataset statistics
eval_stats = tfdv.generate_statistics_from_dataframe(eval_df)
# Compare training with evaluation
tfdv.visualize_statistics(
lhs_statistics=eval_stats,
rhs_statistics=train_stats,
lhs_name='EVAL_DATASET',
rhs_name='TRAIN_DATASET'
)
# + [markdown] id="J3Lvnr-YgPU2"
# ## Calculate and display evaluation anomalies
# + [markdown] id="D1D9bNcigPU2"
# You can use your reference schema to check for anomalies such as new values for a specific feature in the evaluation data. Detected anomalies can either be considered a real error that needs to be cleaned, or depending on your domain knowledge and the specific case, they can be accepted.
#
# Let's detect and display evaluation anomalies and see if there are any problems that need to be addressed.
# + id="OR5dBqpW6ky2"
# Check evaluation data for errors by validating the evaluation dataset statistics using the reference schema
anomalies = tfdv.validate_statistics(statistics=eval_stats, schema=schema)
# Visualize anomalies
tfdv.display_anomalies(anomalies)
# + [markdown] id="HeEvhvgqzLu-"
# ## Revising the Schema
#
# As shown in the results above, TFDV is able to detect the remaining irregularities we introduced earlier. The short and long descriptions tell us what were detected. As expected, there are string values for `race`, `native-country` and `occupation` that are not found in the domain of the training set schema (you might see a different result if the shuffling of the datasets was applied). What you decide to do about the anomalies depend on your domain knowledge of the data. If an anomaly indicates a data error, then the underlying data should be fixed. Otherwise, you can update the schema to include the values in the evaluation dataset.
#
# TFDV provides a set of utility methods and parameters that you can use for revising the inferred schema. This [reference](https://www.tensorflow.org/tfx/data_validation/anomalies) lists down the type of anomalies and the parameters that you can edit but we'll focus only on a couple here.
#
# - You can relax the minimum fraction of values that must come from the domain of a particular feature (as described by `ENUM_TYPE_UNEXPECTED_STRING_VALUES` in the [reference](https://www.tensorflow.org/tfx/data_validation/anomalies)):
#
# ```python
# tfdv.get_feature(schema, 'feature_column_name').distribution_constraints.min_domain_mass = <float: 0.0 to 1.0>
# ```
#
# - You can add a new value to the domain of a particular feature:
#
# ```python
# tfdv.get_domain(schema, 'feature_column_name').value.append('string')
# ```
#
# Let's use these in the next section.
# + [markdown] id="HKECg6Lf6-ks"
# ## Fix anomalies in the schema
#
# Let's say that we want to accept the string anomalies reported as valid. If you want to tolerate a fraction of missing values from the evaluation dataset, you can do it like this:
# +
# Relax the minimum fraction of values that must come from the domain for the feature `native-country`
country_feature = tfdv.get_feature(schema, 'native-country')
country_feature.distribution_constraints.min_domain_mass = 0.9
# Relax the minimum fraction of values that must come from the domain for the feature `occupation`
occupation_feature = tfdv.get_feature(schema, 'occupation')
occupation_feature.distribution_constraints.min_domain_mass = 0.9
# -
# If you want to be rigid and instead add only valid values to the domain, you can do it like this:
# + id="hTCWS04p6lDh"
# Add new value to the domain of the feature `race`
race_domain = tfdv.get_domain(schema, 'race')
race_domain.value.append('Asian')
# -
# In addition, you can also restrict the range of a numerical feature. This will let you know of invalid values without having to inspect it visually (e.g. the invalid `age` values earlier).
# +
# Restrict the range of the `age` feature
tfdv.set_domain(schema, 'age', schema_pb2.IntDomain(name='age', min=17, max=90))
# Display the modified schema. Notice the `Domain` column of `age`.
tfdv.display_schema(schema)
# -
# With these revisions, running the validation should now show no anomalies.
# Validate eval stats after updating the schema
updated_anomalies = tfdv.validate_statistics(eval_stats, schema)
tfdv.display_anomalies(updated_anomalies)
# ## Examining dataset slices
#
# TFDV also allows you to analyze specific slices of your dataset. This is particularly useful if you want to inspect if a feature type is well-represented in your dataset. Let's walk through an example where we want to compare the statistics for male and female participants.
# First, you will use the [`get_feature_value_slicer`](https://github.com/tensorflow/data-validation/blob/master/tensorflow_data_validation/utils/slicing_util.py#L48) method from the `slicing_util` to get the features you want to examine. You can specify that by passing a dictionary to the `features` argument. If you want to get the entire domain of a feature, then you can map the feature name with `None` as shown below. This means that you will get slices for both `Male` and `Female` entries. This returns a function that can be used to extract the said feature slice.
# +
from tensorflow_data_validation.utils import slicing_util
slice_fn = slicing_util.get_feature_value_slicer(features={'sex': None})
# -
# With the slice function ready, you can now generate the statistics. You need to tell TFDV that you need statistics for the features you set and you can do that through the `slice_functions` argument of [`tfdv.StatsOptions`](https://www.tensorflow.org/tfx/data_validation/api_docs/python/tfdv/StatsOptions). Let's prepare that in the cell below. Notice that you also need to pass in the schema.
# Declare stats options
slice_stats_options = tfdv.StatsOptions(schema=schema,
slice_functions=[slice_fn],
infer_type_from_schema=True)
# You will then pass these options to the `generate_statistics_from_csv()` method. As of writing, generating sliced statistics only works for CSVs so you will need to convert the Pandas dataframe to a CSV. Passing the `slice_stats_options` to `generate_statistics_from_dataframe()` will not produce the expected results.
# +
# Convert dataframe to CSV since `slice_functions` works only with `tfdv.generate_statistics_from_csv`
CSV_PATH = 'slice_sample.csv'
train_df.to_csv(CSV_PATH)
# Calculate statistics for the sliced dataset
sliced_stats = tfdv.generate_statistics_from_csv(CSV_PATH, stats_options=slice_stats_options)
# -
# With that, you now have the statistics for the set slice. These are packed into a `DatasetFeatureStatisticsList` protocol buffer. You can see the dataset names below. The first element in the list (i.e. index=0) is named `All_Examples` which just contains the statistics for the entire dataset. The next two elements (i.e. named `sex_Male` and `sex_Female`) are the datasets that contain the stats for the slices. It is important to note that these datasets are of the type: `DatasetFeatureStatistics`. You will see why this is important after the cell below.
# +
print(f'Datasets generated: {[sliced.name for sliced in sliced_stats.datasets]}')
print(f'Type of sliced_stats elements: {type(sliced_stats.datasets[0])}')
# -
# You can then visualize the statistics as before to examine the slices. An important caveat is `visualize_statistics()` accepts a `DatasetFeatureStatisticsList` type instead of `DatasetFeatureStatistics`. Thus, at least for this version of TFDV, you will need to convert it to the correct type.
# +
from tensorflow_metadata.proto.v0.statistics_pb2 import DatasetFeatureStatisticsList
# Convert `Male` statistics (index=1) to the correct type and get the dataset name
male_stats_list = DatasetFeatureStatisticsList()
male_stats_list.datasets.extend([sliced_stats.datasets[1]])
male_stats_name = sliced_stats.datasets[1].name
# Convert `Female` statistics (index=2) to the correct type and get the dataset name
female_stats_list = DatasetFeatureStatisticsList()
female_stats_list.datasets.extend([sliced_stats.datasets[2]])
female_stats_name = sliced_stats.datasets[2].name
# Visualize the two slices side by side
tfdv.visualize_statistics(
lhs_statistics=male_stats_list,
rhs_statistics=female_stats_list,
lhs_name=male_stats_name,
rhs_name=female_stats_name
)
# -
# You should now see the visualization of the two slices and you can compare how they are represented in the dataset.
#
# We encourage you to go back to the beginning of this section and try different slices. Here are other ways you can explore:
#
# * If you want to be more specific, then you can map the specific value to the feature name. For example, if you want just `Male`, then you can declare it as `features={'sex': [b'Male']}`. Notice that the string literal needs to be passed in as bytes with the `b'` prefix.
#
# * You can also pass in several features if you want. For example, if you want to slice through both the `sex` and `race` features, then you can do `features={'sex': None, 'race': None}`.
#
# You might find it cumbersome or inefficient to redo the whole process for a particular slice. For that, you can make helper functions to streamline the type conversions and you will see one implementation in this week's assignment.
# + [markdown] id="YY812jDDgPU3"
# ## Wrap up
#
# This exercise demonstrated how you would use Tensorflow Data Validation in a machine learning project.
#
# * It allows you to scale the computation of statistics over datasets.
#
# * You can infer the schema of a given dataset and revise it based on your domain knowledge.
#
# * You can inspect discrepancies between the training and evaluation datasets by visualizing the statistics and detecting anomalies.
#
# * You can analyze specific slices of your dataset.
#
# You can consult this notebook in this week's programming assignment as well as these additional resources:
#
# * [TFDV Guide](https://www.tensorflow.org/tfx/data_validation/get_started)
# * [TFDV blog post](https://blog.tensorflow.org/2018/09/introducing-tensorflow-data-validation.html)
# * [Tensorflow Official Tutorial](https://colab.research.google.com/github/tensorflow/tfx/blob/master/docs/tutorials/data_validation/tfdv_basic.ipynb#scrollTo=mPt5BHTwy_0F)
# * [API Docs](https://www.tensorflow.org/tfx/data_validation/api_docs/python/tfdv)
# -
| course2/week1-ungraded-lab/C2_W1_Lab_1_TFDV_Exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings('ignore')
warnings.simplefilter('ignore')
pd.set_option('display.max_rows', 99)
pd.set_option('display.max_columns', 99)
df = pd.read_csv('wm_project.csv')
df.head()
df.shape
# <div class="alert alert-block alert-info">
# This dataset has 40886 observations and 21 variables.
# </div>
# # Data Cleaning
df.isna().sum()/df.shape[0]
# <div class="alert alert-block alert-info">
# I will use only the observations with not-null [hit] value to build my model. The test will be predicted later.
# </div>
# Take only the observations with hit as traning dataset, the rest as test dataset
train = df[df.hit.notnull()]
test = df[df.hit.isna()]
train.shape
# <div class="alert alert-block alert-info">
# Now the dataset is down to 33457 observations and 21 variables.
# </div>
# # EDA
train.info()
# track
round(train['track'].nunique()/df.shape[0],2)
# artist
round(train['artist'].nunique()/df.shape[0],2)
# uri
round(train['uri'].nunique()/df.shape[0],2)
# <div class="alert alert-block alert-info">
# Above 3 variables <b>track</b>, <b>artist</b> and <b>uri</b> have too many levels which won't help with building a effective model. I will drop those features.
# </div>
# +
# danceability
# -
sns.distplot(train.danceability, 20)
sns.boxplot(x="hit", y="danceability", data=train)
# <div class="alert alert-block alert-info">
# Hit songs on average have relatively higher danceability.
# </div>
# +
# energy
# -
sns.distplot(train.energy, 20)
sns.boxplot(x="hit", y="energy", data=train)
# <div class="alert alert-block alert-info">
# Hit songs on average have higher energy. Their distribution of energy is denser between around 0.5-0.8.
# </div>
# +
# key
# -
pd.crosstab(train['key'], train['hit']).plot(kind='bar', stacked=True)
# <div class="alert alert-block alert-info">
# Hit/Not-Hit distribution looks similar among different keys, which might lead to Key as a feature having no enough predictive power.
# </div>
# +
# loudness
# -
sns.distplot(train.loudness, 20)
sns.boxplot(x="hit", y="loudness", data=train)
# <div class="alert alert-block alert-info">
# Hit songs on average are louder
# </div>
# +
# mode
# -
pd.crosstab(train['mode'], train['hit']).plot(kind='bar', stacked=True)
# <div class="alert alert-block alert-info">
# There are higher percentage of hit songs in Major track (mode=1)
# </div>
# +
# speechiness
# -
fig, ax = plt.subplots(figsize=(10, 10))
sns.boxplot(x="hit", y="speechiness", data=train,ax=ax)
sns.distplot(train.speechiness, 20)
# <div class="alert alert-block alert-info">
# Overall songs have low speechiness level. Hit songs on average have even lower speechiness level compared with non-hit songs.
# </div>
# +
# acousticness
# -
sns.distplot(train.acousticness, 20)
sns.boxplot(x="hit", y="acousticness", data=train)
# <div class="alert alert-block alert-info">
# Hit songs have on average lower acousticness level.
# </div>
# +
# instrumentalness
# -
sns.distplot(train.instrumentalness, 20)
sns.boxplot(x="hit", y="instrumentalness", data=train)
train.groupby('hit')['instrumentalness'].mean()
# <div class="alert alert-block alert-info">
# Hit songs have more skewed distribution of instrumentalness while non-hit songs are mostly distributed around 0-0.5 instrumentalness.
# </div>
# +
# liveness
# -
sns.distplot(train.liveness, 20)
sns.boxplot(x="hit", y="liveness", data=train)
train.groupby('hit')['liveness'].mean()
# <div class="alert alert-block alert-info">
# Hit songs have lower level of liveness
# </div>
# +
# valence
# -
sns.distplot(train.valence, 20)
sns.boxplot(x="hit", y="valence", data=train)
# <div class="alert alert-block alert-info">
# Hit songs have higher level of valence on average
# </div>
# +
# tempo
# -
sns.distplot(train.tempo, 20)
fig, ax = plt.subplots(figsize=(10, 10))
sns.boxplot(x="hit", y="tempo", data=train,ax=ax)
train.groupby('hit')['tempo'].mean()
# <div class="alert alert-block alert-info">
# Hit songs have higher level of tempo on average
# </div>
# +
# duration_ms
# -
sns.distplot(train.duration_ms, 20)
fig, ax = plt.subplots(figsize=(10, 10))
sns.boxplot(x="hit", y="duration_ms", data=train, ax=ax)
train.groupby('hit')['duration_ms'].mean()
# <div class="alert alert-block alert-info">
# Hit songs on average is shorter
# </div>
# +
# time_signature
# -
train.time_signature.value_counts()
pd.crosstab(train['time_signature'], train['hit']).plot(kind='bar', stacked=True)
# <div class="alert alert-block alert-info">
# Most songs have 4 as time_signature, and time_signature=4 has a highest percentage of hit songs.
# </div>
# +
# chorus_hit
# -
sns.distplot(train.chorus_hit, 20)
fig, ax = plt.subplots(figsize=(10, 10))
sns.boxplot(x="hit", y="chorus_hit", data=train,ax=ax)
train.groupby('hit')['chorus_hit'].mean()
# <div class="alert alert-block alert-info">
# Hit songs on average have a smaller chorus_hit
# </div>
# +
# sections
# -
train.sections.nunique()
pd.crosstab(train['sections'], train['hit']).plot(kind='bar', stacked=True)
fig, ax = plt.subplots(figsize=(10, 10))
sns.boxplot(x="hit", y="sections", data=train,ax=ax)
train.groupby('hit')['sections'].mean()
# <div class="alert alert-block alert-info">
# It's better to take sections as a continuous numeric variable. Hit songs' sections have a more compact distribution where most of the value are between 0-75 sections, and hit songs have slightly lower number of sections.
# </div>
# +
# decade
# -
train.decade.value_counts()
pd.crosstab(train['decade'], train['hit']).plot(kind='bar', stacked=True)
# <div class="alert alert-block alert-info">
# 60s and 70s have higher percentage of hit songs.
# </div>
# # Feature Engineering
y = train.hit
X = train.loc[:, train.columns != 'hit']
# <div class="alert alert-block alert-info">
# As mentioned above, I will remove track, artist and uri.
# </div>
del X['id']
del X['track']
del X['artist']
del X['uri']
num_col = ['danceability','energy','loudness','speechiness',
'acousticness','instrumentalness','liveness','valence','tempo',
'duration_ms','chorus_hit','sections']
cat_col = ['key','mode','time_signature','decade']
for x in cat_col:
X[x] = X[x].astype('category')
# <div class="alert alert-block alert-info">
# One-hot-encoding for categorical variable
# </div>
pd.get_dummies(X[cat_col])
X_encoded = pd.concat([pd.get_dummies(X[cat_col]), X[num_col]], axis=1)
# # Feature Selection
train[train.hit == 1].shape[0]/df.shape[0] #40/60
# <div class="alert alert-block alert-info">
# In this sample, hit is around 40% of the entire population, therefore I will be using 40/60 as the class_weight going forward.
# </div>
# <div class="alert alert-block alert-info">
# Following I will use Boruta as my feature selection method.
# </div>
#
# Boruta method is based on two idea:
# - In Boruta, features do not compete among themselves. They compete with a randomized version of them.
# - The threshold is defined as the highest feature importance recorded among the shadow features. A feature is useful only if it’s capable of doing better than the best randomized feature.
# +
# Boruta
# -
from boruta import BorutaPy
from sklearn.ensemble import RandomForestClassifier
import numpy as np
# +
### initialize Boruta
forest = RandomForestClassifier(
n_jobs = -1,
max_depth = 5,
n_estimators=1000,
criterion='entropy',
class_weight={1:0.4,0:0.6},
random_state=42)
boruta = BorutaPy(
estimator = forest,
n_estimators = 'auto',
max_iter = 100 # number of trials to perform
)
### fit Boruta (it accepts np.array, not pd.DataFrame)
boruta.fit(np.array(X_encoded), np.array(y))
### print results
selected_features = X_encoded.columns[boruta.support_].to_list()
tbd_features = X_encoded.columns[boruta.support_weak_].to_list()
print('features selected:', selected_features)
print('features in consideration:', tbd_features)
# -
set(X_encoded.columns) - set(selected_features)
# <div class="alert alert-block alert-info">
# I will remove above features as they are not informative.
# </div>
X_encoded = X_encoded[selected_features]
# # XGBOOST
# conda install -c anaconda py-xgboost
import xgboost as xgb
from sklearn.model_selection import train_test_split
from sklearn.metrics import balanced_accuracy_score, roc_auc_score, make_scorer
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import confusion_matrix
from sklearn.metrics import plot_confusion_matrix
X_train, X_test, y_train, y_test = train_test_split(X_encoded, y, random_state=42, stratify=y)
# define a timer to keep track on training time
def timer(start_time=None):
if not start_time:
start_time = datetime.now()
return start_time
elif start_time:
thour, temp_sec = divmod((datetime.now()-start_time).total_seconds(), 3600)
tmin,tsec = divmod(temp_sec, 60)
print('\n Time taken: %i hours %i minutes and %s seconds.' % (thour, tmin, round(tsec,2)))
# defining a generic Function to give ROC_AUC Scores in table format for better readability
from sklearn.model_selection import cross_val_score
def crossvalscore(model):
scores = cross_val_score(model,X_encoded,y,cv=5,scoring='roc_auc',n_jobs=-1)
acc = cross_val_score(model,X_encoded,y,cv=5,scoring='accuracy',n_jobs=-1)
rand_scores = pd.DataFrame({
'cv':range(1,6),
'roc_auc score':scores,
'accuracy score':acc
})
print('AUC :',rand_scores['roc_auc score'].mean())
print('accuracy :',rand_scores['accuracy score'].mean())
return rand_scores.sort_values(by='roc_auc score',ascending=False)
# <div class="alert alert-block alert-info">
# Build preliminary XGB
# </div>
### initialize XGB
clf_xgb = xgb.XGBClassifier(objective='binary:logistic',
missing=None,
seed=42,
eval_metric='aucpr')
### fit XGB
clf_xgb.fit(X_train,
y_train,
early_stopping_rounds=10,
eval_metric='aucpr',
verbose=3,
eval_set=[(X_test, y_test)])
crossvalscore(clf_xgb)
# <div class="alert alert-block alert-info">
# Optimize XGB by tuning hyperparameter
# </div>
### Hyperparameter Optimization for Xgboost with RandomizedSearchCV
param_grid_3 = {
'max_depth':[25],
'learning_rate':[0.095],
'gamma':[5],
'reg_lambda':[72],
'scale_pos_weight':[1]}
from sklearn.model_selection import RandomizedSearchCV
random_search = RandomizedSearchCV(clf_xgb,
param_distributions=param_grid_3,
n_iter=5,
scoring='roc_auc',
n_jobs=-1,
cv=5,
verbose=3)
from datetime import datetime
start_time = timer(None)
random_search.fit(X_encoded,y)
timer(start_time)
### output optimized parameters
random_search.best_estimator_
### initialize and fit the model with optimized parameters
clf_xgb_optimized = xgb.XGBClassifier(seed=42,
objective='binary:logistic',
max_depth=25,
gamma=5,
learning_rate=0.095,
reg_lambda=72,
scale_pos_weight=1)
clf_xgb_optimized.fit(X_train,
y_train,
early_stopping_rounds=10,
eval_metric='aucpr',
eval_set=[(X_test, y_test)])
crossvalscore(clf_xgb_optimized)
# <div class="alert alert-block alert-info">
# Plot feature importance from optimized XGB
# </div>
# Cover. The number of times a feature is used to split the data across all
# trees weighted by the number of training data points that go through those
# splits.
xgb.plot_importance(clf_xgb_optimized, importance_type="cover")
# Gain. The average training loss reduction gained when using a feature for
# splitting.
xgb.plot_importance(clf_xgb_optimized, importance_type="gain")
# Weight. The number of times a feature is used to split the
# data across al trees
xgb.plot_importance(clf_xgb_optimized, importance_type="weight")
# +
# Calibrate Predict_proba with CalibratedClassifierCV
# -
from sklearn.calibration import CalibratedClassifierCV
calibrated = CalibratedClassifierCV(clf_xgb_optimized, method='isotonic', cv='prefit')
### fit the calibrated model with test dataset
calibrated = calibrated.fit(X_test, y_test)
### calibrated probability
probability = (calibrated.predict_proba(X_encoded))
# # Prediction
# +
# Handle test dataset
X_test = test.loc[:, test.columns != 'hit']
del X_test['id']
del X_test['track']
del X_test['artist']
del X_test['uri']
for x in cat_col:
X_test[x] = X_test[x].astype('category')
X_test_encoded = pd.concat([pd.get_dummies(X_test[cat_col]), X_test[num_col]], axis=1)
X_test_encoded = X_test_encoded[selected_features]
# -
pred = pd.DataFrame(calibrated.predict_proba(X_test_encoded))[1]
pred_v1=pd.concat([test.id.reset_index(drop=True).astype('str'),pred],axis=1, ignore_index=True)
pred_v1.columns = ['id', 'probability']
# <div class="alert alert-block alert-info">
# Predicted value for the observations with missing hit:
# </div>
pred_v1
| Hit_Song_Project_Fin.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Rovick1/CPEN21-BSCPE-1-2/blob/main/Midterm_Exam.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="FixTcusUA0fA"
# ##Midterm Exam
# + [markdown] id="kzTRE5HEClwo"
# ##Promblem 1
# + colab={"base_uri": "https://localhost:8080/"} id="WNHKlzaUC5M_" outputId="c0d31824-1596-49b9-8cc4-d68a92edb415"
fullname ="<NAME>"
studentnumber =" 202101632"
age =" 18"
birthday =" anuary 16, 2003"
address =" Maitim 1st, Amadeo"
course =" BSCpE 1-2"
print(fullname+studentnumber+age+birthday+address+course)
# + [markdown] id="-QxfroGAF6mn"
# ##Problem 2
# + colab={"base_uri": "https://localhost:8080/"} id="ExwuNxIFNtJY" outputId="736cacd0-dcd0-46c0-8742-7d215ceb53e4"
n=4
answ="Y"
print(bool(2<n) and (n<6)) #a
print(bool(2<n) or (n==6)) #b
print(bool(not (2<n) or (n==6))) #c
print(bool(not (n<6))) #d
print(bool(answ=="Y") or (answ=="y")) #e
print(bool(answ=="Y") and (answ=="y")) #f
print(bool(not(answ=="y"))) #g
print(bool((2<n) and (n==5+1)) or (answ=="No")) #h
print(bool((n==2) and (n==7)) or (answ=="Y")) #i
print(bool(n==2) and ((n==7) or (answ=="Y"))) #j
# + [markdown] id="HjmWQPXAGEwI"
# ##Problem 3
# + colab={"base_uri": "https://localhost:8080/"} id="x0v1m-J8Ggc2" outputId="a0f22ef2-78f3-4398-9896-f4e39e0a265f"
x=2
y=-3
w=7
z=-10
print(w/y)
print(w/y/x)
print(z/y%x)
print(x%-y*w)
print(x%y)
print(z%w-y/x*5+5)
print(9-x%(2+y))
print(z//w)
print((2+y)**2)
print(w/x*2)
| Midterm_Exam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Was ist Machine Learning?
#
# > Machine Learning ist die Wissenschaft, Computer so zu programmieren, dass sie anhand von Daten lernen.
#
# Gelernt wird mit Trainingsdaten(einem Trainingsdatensatz) wobei jedes Trainingsbeispiel auch Trainingsdatenpunkt-/Instanz genannt wird
# # Warum wird ML verwendet?
#
# ML wird verwendet, wenn:
# * die Aufgabe zu komplex für herkömmliche Aufgaben ist
# * eine Menge Handarbeit und lange Regeln geschrieben werden müssten
# * kein bekannter Algorithmus für diese Aufgabe existert
# * die Aufgabe mit sich stark verändernden Daten beschäftigt(ML ist Anpassungsfähig)
# * man riesige Datenmengen besitzt und daraus erkenntnisse gewinnen soll.(auch hier ist die Aufgabenstellung komplex)
# # Arten von ML
#
# Es wird unterschieden nach:
# * Überwachtes, unüberwachtes und halbüberwachtes Lernen sowie Reinforcement Learning
# * ständig dazu lernen(Online-learning) oder nicht(Batch-Learning)
# * Vergleicht der Algorithmus neue Datenpunkte mit bereits bekannten oder erkenner er Muster in den Daten
# ## Überwachtes Lernen
#
# Die Trainingsdaten sind **gelabelt**(sie enthalten die gewünschten Lösungen)
#
# ### Beispiele
#
# * Spamfilter -> **Klassifikation**
# * Vorhersagen eine Preises(nummerische Grösse) aufgrund von Merkmalen(**Prädikatoren**) -> **Regression**
#
#
# ### Algorithmen
#
# * **k-nearest-Neighbors**
# * **linear Regression**
# * **logistic Regression**
# * **Support Vector Machines**
# * **decision trees / random forests**
# * **neural networks**
# ## Unüberwachtes Lernen
#
# Es wird versucht ohne Anleitung zu lernen. Die Daten sind nicht gelabelt.
#
# ### Beispiele
#
# * Visualisierung -> Um grosse Datenmengen zu verstehen
# * Clustering -> Einteilung anhand von grossen Datenmengen vornehmen
# * Erkennen von Anomalien
# * Abfangen von Produktionsfehlern und entfernen von Ausreissern in Datensätzen
# * Lernen von Assoziationsregeln
#
# ### Algorithmen
#
# * **Clustering**
# * k-Means
# * hierarchical Cluster Analysis -> HCA
# * Expectaiton Maximation
# * Visualizing and Dimensionreduction
# * Principal component analysis -> PCA
# * Kernel PCA
# * locally-linear Embedding(LLE)
# * t-distributed stochastic neighbor embedding
# * learning with assoziation rules
# * apriori
# * eclat
#
# ### Dimensionsreduktion
#
# Die Daten vereinfachen, ohne Informationen zu verlieren.
# Auch **Extraktion von Merkmalen** genannt.
#
# ## Halbüberwachtes Lernen
#
# Der Grossteil der Trainingsdaten sind **nichtt** gelabelt, der kleine Teil dafür.
# Meist sind halbüberwachte Algorithmen Mischungen aus Überwachten- und Unüberwachten Algorithmen, diese werden einfach kobiniert für ein besseres Resultat.
#
# ### Beispiele
#
# * Google Photots erkennt mithilfe eines unüberwachten lernens, dass eine Person auf mehreren Fotos sichtbar ist. Aber um zu lernen, wie diese Person heisst, braucht es mindestens einen gelabelten Datensatz mit dem Namen.
#
# ### Algorithmen
#
# * Wie bereits erwähnt, sind dies meist Kombinationen von Überwachtem- und Unüberwachtem Lernen.
#
# ## Reinforcement Learning / Verstärkendes Lernen
#
# Der **Agent** hat verschiedene Möglichkeiten zu handeln.
# Diese führt er aus und wird danach **belohnt oder bestraft**(mit negativen Belohnungen).
# Durch diese Erfahrungen kann der Agent seine **Policy/Strategie** anpassen und weiss das nächste mal, welches die richtige Entscheidung ist.
#
# # Batch Learning
#
# Beim Batch-Lernen wird das System mit allen verfügbaren Daten trainiert und danach kann es verwendet werden.
# Ein solches System kann allerding nicht mit neuen Daten gefüttert/erweitert werden.
# Wenn das System trotzdem mit neuen Trainingsdaten erweitert werden soll, so muss es nochmals mit allen alten **und** den neuen Trainingsdaten trainiert werden.
# Dies benötigt sehr viel ressource, kann teuer werden und das trainierte System ist nicht anpassungsfähig.
#
# # Online Learning
#
# Die Trainingsdaten werden als erstes in kleine Stücke(batches) aufgeteilt, welche dann nacheinander trainiert werden und nach dem Training verworfen werden können, ausser man möchte einen alten Zustand wiederherstellen können.
# Das Aufteilen wird auch **out-of-core-lernen** genannt. Es werden weniger Ressourcen als beim Batch-Lernen benötigt.
#
# Ein Vorteil hier ist auch, das das System anpassungsfähig an neue Daten ist.
# Die Anpassungsfähigkeit kann eingestellt werden mithilfe einer **Lernrate**.
# Eine hohe Lernrate führt dazu, dass neues schnell angewendet und altes schnell vergessen wird.
# Eine tiefe sorgt im Gegenzug für eine gewisse Trägheit des System.
#
# Bei einer hohen Lernrate ist Vorsicht geboten, da Anomalien oder gefälschte Trainingsdaten die Qualität beeinflussen. Ein solches System sollte überwacht werden oder Anomalien zuvor entfernt werden.
# # Instanzbasiertes Lernen
#
# Das System wird mit den Trainingsdaten trainiert und lernt diese dadurch "Auswendig".
# Zusätzlich wird ein **Ähnlichkeitsmass** festgelegt. -> Bei **k-nearest-Neighbors** etwa die Anzahl Nachbarn **k**.
# Mit diesen zwei Dingen kann Verallgemeinert werden und neue Datensätze können vorhergesagt werden.(aufgrund der Ähnlichkeit mit bereits Trainierten Daten).
#
# # Modellbasiertes Lernen
#
# Das System versucht aus den Trainingsdaten ein Modell zu erstellen, welches auf möglichst alle Trainingsdaten möglichst gut zutrifft. Dieses Modell kann etwa eine lineare Gleichung sein. Neue Datensätze können in die Gleichung eingesetzt und somit vorhergesagt werden. Ein Beispiel ist die **Lineare Regression**.
# Die beste Gleichung wird durch den Algorithmus ermittelt.
#
# Der Algorithmus muss die Gleichung anhand der Trainingsdaten verbessern. Dazu muss er erst in der Lage sein die **Leistung der aktuellen Gleichung** zu messen.
# Dies wird mit einer
# * **Kostenfunktion** -> Wie schlecht ist das Modell?
# * Oder mit einer **Nutzenfunktion** -> Wie gut ist das Modell? /Güte des Modells
#
# geprüft.
#
# ## Lineare Gleichung
#
# Die Parameter **m** und **q** werden ermittelt um für **x** eine Vorhersage **y** zu treffen.
# $y=mx+q$
# $y=\Theta_0*x+\Theta_1$
# Im ML wird für die Parameter **m** und **q** der griechische Buchstabe **$\Theta$** verwendet.
# # Erstes Beispiel
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from sklearn import linear_model, neighbors
def prepare_country_stats(satisfaction, bipp):
satisfaction = satisfaction[satisfaction["INEQUALITY"]=="TOT"]
satisfaction = satisfaction.pivot(index="Country", columns="Indicator", values="Value")
bipp.rename(columns={"2015": "BIPP"}, inplace=True)
bipp.set_index("Country", inplace=True)
full_country_stats = pd.merge(left=satisfaction, right=bipp,
left_index=True, right_index=True)
full_country_stats.sort_values(by="BIPP", inplace=True)
remove_indices = [0, 1, 6, 8, 33, 34, 35]
keep_indices = list(set(range(36)) - set(remove_indices))
return full_country_stats[["BIPP", 'Life satisfaction']].iloc[keep_indices]
import os
datapath = os.path.join("datasets", "lifesat", "")
#Laden der Daten
satisfaction = pd.read_csv(datapath+'oecd_bli_2015.csv', thousands=',')
bipp = pd.read_csv(datapath +'gdp_per_capita.csv', thousands=',', delimiter='\t', encoding='latin1', na_values='n/a')
#Vorbereiten der Daten
country_stats = prepare_country_stats(satisfaction, bipp)
x = np.c_[country_stats['BIPP']]
y = np.c_[country_stats['Life satisfaction']]
#Visualisieren
country_stats.plot(kind='scatter', x='BIPP', y='Life satisfaction')
plt.show()
#Asuwahl des Algorithmus
#Modelbasiert
#model = linear_model.LinearRegression()
#Instanzbasiert
model = neighbors.KNeighborsRegressor()
model.fit(x, y)
#Vorhersage treffen
x_new = [[25000]]# Pro kopf BIP für Zypern
print(model.predict(x_new))
# # Schlechte Daten: Müll rein -> Müll raus
#
# * Zu wenig Trainingsdaten
# * Nicht repräsentative Trainingsdaten
# * Minderwertige Daten
# * Irrevelante Merkmale
#
# ## Zu wenig Trainingsdaten
# Hat man sehr viele Trainingsdaten zur verfügung, dann können primitive Algorithmen ähnlich gut Vorhersagen wie komplexe Algorithmen. Dies zeigt, wie wichtig die Menge an Trainingsdaten ist.
# Allerdings sollte man die Algorithmen trotzdem nicht ausser Acht lassen, denn auch sie spielen noch eine Rolle, gerade, wenn nicht viele Trainingsdaten zur verfügung stehen.
#
# ## Nicht repsäsentative Trainingsdaten
# Die Trainingsdaten sollten mit Vorsicht erhoben werden. Etwa bei einer Umfrage über das Kaufverhalten muss darauf geachtet werden, dass alle Demografischen Gruppen befragt werden, und von allen auch eine grosse Menge.
#
# ## Minderewertige Daten
# Die Daten können lückenhaft sein oder Ausreisser enthalten. Diese sollten korrigiert oder entfernt werden.
# Man könnte aber auch zwei Systeme trainieren, 1x mit den lückenhaften Daten und 1x ohne...
#
# ## Irrevelante Merkmale
# * Die Merkmale sollten sorgfältig ausgewählt werden, da das System aus wesentlichen/aussagekräftigen Merkmalen lernen soll.
# * Die Merkmale könnten etwa mit Dimensionsreduktion extrahiert werden
# * Neue Merkmale könnten erhoben werden.
# # Schlechter Algorithmus
# * Overfitting
# * Underfitting
#
# ## Overfitting
# Wie Menschen, so können auch Maschinen zu einer **übermässigen Verallgemeinerung** neigen, dies nennt man **Overfitting**.
# Bei ML tritt dies unter folgenden Bedingungen auf:
# 1. Zu komplexes Modell angesichts der Trainingsdaten:
# * Zu komplex bedeutet z.B, dass zu viele Parameter erwartet werden.
# * Zu wenig Trainingsdaten
# * Zu viel Rauschen und Ausreisser(Anomalien) in den Trainingsdaten
#
# Dem ersten kann man entgegewirken, indem man ein einfacheres Modell wählt oder man legt dem Modell Restriktionen auf, dies nennt man **Regularisieren**.
#
# $y=mx+q$
# $y=\Theta_0*x+\Theta_1$
#
# Man stelle sich nun vor, man würde $\Theta_1=0$ setzen, dann wäre das Modell vereinfacht, da sich nur noch $\Theta_0$ anpassen kann beim Trainieren.
# Man spricht hier auch von **Freiheitsgraden**. Statt zwei Freiheitsgraden besitzt das Modell jetzt nur noch einen.
# Würde man $\Theta_1$ nicht gleich $0$ setzen, aber trotzdem einen kleinen Wert erzwingen, so hätte das Modell zwischen 1 und 2 Freiheitsgraden.
#
# Um diese Regularisierung vorzunehmen, wendet man einen **Hyperparameter** auf die **Trainingsdaten**(nicht das Modell!) an. Dieser Hyperparameter bleibt konstant.
# Ein grosser Hyperparameter verhindert zwar Overfitting, reduziert aber auch die Leistung des Systems.
#
# ## Underfitting
#
# Der Gegenzug von Overfitting ist Underfitting. Ursachen:
# * Das Modell ist zu einfach für die Realität oder:
# >Die Realität ist komplexer als das Modell
#
# Man kann dies Beheben indem man ein komplexeres Modell wählt, welches auch fähig ist die in den Daten enthaltene Sruktur zu lernen, oder man verringert die Restriktionen des Modells(z.B. Hyperparameter verkleinern).
#
# # Testen und Validieren
#
# Um die Leistung(Wie gut verallgemeinert ein trainiertes Modell auf neue Datenpunkte) zu evaluieren testet man das Modell mit neuen Datenpunkten. Die Abweichung bei diesen neuen Datenpunkten nennt man **out-of-sample-error**.
#
# Die Daten aufteilen:
# 80% Trainingsdaten
# 20% Testdaten
#
#
# Man möchte den Besten Hyperparameter für ein Modell ermitteln.
# Vorgehen:
# 1. Testdaten und Trainingsdaten festlegen
# * Die Trainingsdaten werden in komplementäre Untermengen eingeteilt(Trainingsdaten und Validierungsdaten).
# * Jedes Modell wird mit einer anderen Kombination dieser Untermengen trainiert
# * Und mit den restlichen Validiert
# * Das am Besten abschneidende Modell wird nun mit den Testdaten getestet um den **Verallgemeinerungsfehler** festzulegen.
#
# ## No-Free-Lunch-Theorem
# Um zu bestimmen ob ein Modell "besser oder schlechter" ist als ein anderes, so muss zwingend eine Annahme über die Daten getroffen werden. Ohne eine Annahme über die Daten sind alle Modelle gleich gut/schlecht geeignet.
# >Kein Modell ist a priori(ohne weitere Beweise, hier eine Annahme über die Daten) besser als ein anderes.
# # Übungen
#
# **Wie würden sie ML definieren?**
# >ML bedeutet Maschinen aus Daten lernen zu lassen um Vorhersagen zu treffen.
#
# **Können sie vier Arten von Aufgaben nennen, für die Machine Learning geeignet ist?
# >1. Bilder klassifizieren
# * Preis eines Produktes anhand seiner Merkmale bestimmen
# * Einem Auto das Fahren beibringen
# * Die Kunden in Kategorien einordnen
#
# **Was ist ein gelabelter Trainingsdatensatz?**
# > Trainingsdaten mit den erwarteten Lösungen, also den erwünschten Ergebnissen.
# z.B. Bilder welches Klassifiziert werden soll, enthalten zusätzlich die Stichworte von dem was darauf abgebildet ist.
#
# **Was sind die zwei verbreitetesten Aufgaben beim überwachten Lernrn?**
# >1. Spamfilter für Emails
# * Preisvorhersagen aufgrund von Merkmalen
#
# **Können sie vier verbreitete Aufgaben für unüberwachtes Lernen nennen?**
# >1. Erkennen von Anomalien, etwa in Kreditkartentransaktionen
# * Erkennung von Assoziationen in riesigen Datenmengen, vielleicht sieht ein Modell mehr oder anderes als Menschen
# * Visualisierung von riesigen Datenmengen in 2d, oder 3d Diagrammen.
# * Clustering, Einteilungen anhand grosser Daten vornehmen
#
# **Was für einen ML-Algorithmus würden sie verewnden um einen Roboter über verschiedene Unbekannte Oberflächen laufen zu lassen?**
# >Einen Reinforcement-Learning Algorithmus eignet sich dazu, da er nicht direkt etwas vorhersagen muss, sondern eine Strategie lernen muss.
#
# **Welche Art Algorithmus würden sie verwenden, um Ihre Kunden in unterschiedliche Gruppen einzuteilen?**
# >Einen Clustering oder hierarchischen Clustering Algorithmus.
#
# **Würden sie die Aufgabe, Spam zu erkennen, als überwachte oder unüberwachte Lernaufgabe einstufen?**
# >Als überwachte, das Modell soll mit bereits gelabelten E-Mails trainiert werden.
#
# **Was ist ein Online-Lernsystem?**
# >Ein System, bei welchem ständig neue Datensätze dazu trainiert werden können. Es ist Anpassungsfähig. Die Lernrate bestimmt ob das System träge sein soll oder sich schnell an die neuen Daten anpassen soll.
#
# **Was ist Out-of-Core-Lernen?**
# >Dies kann bei einem Online-Lernsystem angewendet werden. Wenn die Trainingsdaten zu gross sind um sie alle im Hauptspeicher zu halten, so wird der Trainingsdatensatz in kleine Einheiten(Batches) eingeteilt, welche dann nacheinander in den Hauptspeicher geladen, trainiert und verworfen werden.
#
# **Welche Art Lernalgorithmus beruht auf einem Ähnlichkeitsmass, um Vorhersagen zu treffen?**
# >Der k-nearest-Neighbors Algorithmus. Er ist ein überwachter Lernalgorithmus.
#
# **Was ist der Unterschied zwischen einem Modellparameter und einem Hyperparameter eines Lernalgorithmus?**
# >1. Der Modellparameter wird während des Trainigs auf das Modell angewendet. Die Modellparameter passen sich während des Lernprozesses an. Dies ist das eigentliche Lernen.
# * Der Hyperparameter wird vor dem Training auf alle Trainingsdaten angewendet und kommt nie mit dem Modell selber in Berührung, er ist ausserdem konstant.
#
# **Wonach suchen modelbasierte Lernalgorithmen? Welches ist die häufigste Strategie, die zum Erfolg führt? Wie treffen sie Vorhersagen?**
# >Sie suchen nach einem Modell, welches Bestmöglich verallgemeinert und somit gute Vorhersagen auf neue unbekannte Daten treffen kann. Das Modell ist eine Gleichung mit verschiedenen trainierten Parametern, wird nun ein neuer Datensatz eingespeisst, so rechnet es aus $x$ das $y$ aus. Beim trainieren wird ständig die Leistung mithilfe einer Kosten- oder Nutzenfunktion(meist ersteres) gemessen und die Parameter des Modell werden angepasst, um die Leistung bestmöglich zu steigern.
#
# **Können sie vier der wichtigsten Herausforderungen beim ML bennenen?**
# >1. Overfitting/Underfitting
# * Zu wenig Trainingsdaten
# * Nicht repräsentative Trainingsdaten
# * Irrelevante Merkmale
#
# **Welches Problem liegt vor, wenn ihr Modell auf den Trainigsdaten ein sehr gute leistung erbringt, aber schlecht auf neue Daten verallgemeinert? Nennen sie drei Lösungsansätze**
# >Das Problem ist Overfitting.
# 1. Ein einfacheres Modell wählen
# * Regularisieren mithilfe von Hyperparametern
# * Mehr Trainingsdaten beschaffen.
#
# **Was ist ein Testdatensatz, und warum sollte man einen verwenden?**
# >Das Modell wurde mit Trainingsdaten trainiert und wird mit diesen eine sehr hohe Leistung erzielen, allerdings ist diese nicht Repräsentativ. Die Testdaten werden verwendet um die Leistung eines Modells zu bestimmen, da sie für das Modell noch unbekannt sind und wie neue reale Daten wirken.
#
# **Was ist der Zweck eines Validierungsdatensatzes?**
# >Er hilft, aus verschiedenen Varianten/Modellen das Beste auszusuchen. Er wird aber nicht zur Bestimmung der Leistung eines Modells verwendet.
#
# **Was kann schiefgehen, wenn sie Hyperparameter mithilfe der Testdaten einstellen?**
# >Der Hyperparameter wird dann so eingestelt, dass er zwar auf diese Testdaten eine Höchstleistung erzielt. Aber das Modell kann deswegen nicht gut verallgemeinern und wird auf neue Datensätze eine weniger gute Leistung erzielen.
#
# **Was ist Kreuzvalidierung, und warum sollten Sie diese einem Validierungsdatensatz vorziehen?**
# >Dies ist ein Vorgehen um Modelle miteinander zu vergleichen, ohne das weitere Daten benötigt werden.
#
#
| 01_the_machine_learning_landscape_summary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/wanderloop/WanderlustAI/blob/master/db_search_predict.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="tk_F1tseZxcc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 202} outputId="60e76e2e-28d2-44bd-99e5-816ed69bec61"
from datetime import datetime
from math import radians, sin, cos, atan2, sqrt
def Union(lst1,
lst2,):
final_list = list(set(lst1) | set(lst2))
return final_list
def distance(lat1,
lon1,
lat2,
lon2,
u,):
radius = 6371 # km
dlat, dlon = radians(lat2-lat1), radians(lon2-lon1)
a = sin(dlat/2) * sin(dlat/2) + cos(radians(lat1)) \
* cos(radians(lat2)) * sin(dlon/2) * sin(dlon/2)
c = 2 * atan2(sqrt(a),
sqrt(1-a))
d = radius * c
return True if d < u else False
def date_time_match_check(time1,
date1,
time2,
date2,
goku,):
def date_time_match(time1, date1,
time2, date2,):
def date_time(time,
date,):
def time_split(time):
x=time.split(':')
return(int(x[0]),
int(x[1]),)
def date_split(date):
k=date.split('/')
return(int(k[1]),
int(k[0]),)
m, n=time_split(time)
g, h=date_split(date)
day1 = datetime(2020, h, g,
m, n, 0,)
return(day1)
day1=date_time(time1,
date1,)
day2 = date_time(time2,
date2,)
diff = day1-day2
e=str(diff).split('days')
e=e[0].split(':')
#print(e)
hrs, mins, secs=int(e[0]), int(e[1]), int(e[0])
if hrs<1:
if mins<goku:
return('True')
else:
return('False')
else:
return('False')
try:
x=date_time_match(time1, date1,
time2, date2,)
return(x)
except:
return('False')
def string_ParseTo_Float(mystring):
alist=[]
k=mystring.split("X:")
j=k[0]
x=j.split("ID:")
j=x[1]
ID=j
m=k[1].split("Y:")
long=m[0]
n=m[1].split("TIME:")
lat=n[0]
c=n[1].split("DATE: ")
time=c[0]
date=c[1]
return(ID,
long,
lat,
time,
date,)
with open('text_database') as f:
content = f.readlines()
#print('Is the file closed?', f.closed)
content = [x.strip() for x in content]
j=0
rows, cols = (len(content), 5)
arr = [[0]*cols]*rows
IDLis=[]
longLis=[]
latLis=[]
timeLis=[]
dateLis=[]
for i in range(1,
len(content),):
try:
ID, long, lat, time, date=string_ParseTo_Float(content[i])
IDLis.append(ID)
longLis.append(long)
latLis.append(lat)
timeLis.append(time)
dateLis.append(date)
except:
continue
item=input('Enter Patient ID : ')
def find(item, where):
j=0
note=[]
item=' '+item+' ' # Only applicable for Notepad data
for i in where:
if i==item:
note.append(j)
j=j+1
else:
j=j+1
return(note)
y=find(item,
IDLis)
'''
print(y)
Generate list1 backward
'''
def backward(y):
note=[]
for i in y:
note.append(int(i)-1)
return(note)
note=backward(y)
#print(note)
def forward(y):
note=[]
for i in y:
note.append(int(i)+1)
return(note)
note=forward(y)
'''
print(note)
----------------------------------------------------------------------------
all values of dates of infected person (General)
'''
inf_Gen_dates=[]
for i in y:
inf_Gen_dates.append(dateLis[int(i)])
#print(inf_Gen_dates)
inf_Gen_time=[]
for i in y:
inf_Gen_time.append(timeLis[int(i)])
'''
print(inf_Gen_time)
------------------------------------------------------------------------------
all values of dates of close_match person (Backward)
'''
note=backward(y)
inf_back_dates=[]
for i in note:
inf_back_dates.append(dateLis[int(i)])
#print(inf_back_dates)
inf_back_time=[]
for i in note:
inf_back_time.append(timeLis[int(i)])
'''
print(inf_back_time)
------------------------------------------------------------------------------
Back dates and time matchup with general
'''
h=0
date_time_result=[]
goku=input('Enter Time Difference in minutes for Backward Check : ')
for i in range(0,
len(y),):
s=date_time_match_check(inf_Gen_time[h],
inf_Gen_dates[h],
inf_back_time[h],
inf_back_dates[h],
int(goku),)
date_time_result.append(s)
h=h+1
'''
print(date_time_result)
-------------------------------------------------------------------------------
extracting True Positives for further proceeding
'''
posLis=[]
q=0
for i in date_time_result:
if i=='True':
posLis.append(q)
q=q+1
else:
q=q+1
#print(posLis)
new_lis=[]
for i in posLis:
new_lis.append(y[i])
#print(new_lis)
y=new_lis
'''
print(y)
----------------------------------------------------------------------------
Now we test distance--------------------------------General Distance--------
'''
inf_Gen_longitude=[]
for i in y:
inf_Gen_longitude.append(longLis[int(i)])
'''
print(inf_Gen_longitude)
-----------------------------------------------------
latitude
'''
inf_Gen_latitude=[]
for i in y:
inf_Gen_latitude.append(latLis[int(i)])
'''
print(inf_Gen_latitude)
---------------------------------------------------Backward longitude
'''
inf_Gen_longitude_back=[]
note=backward(y)
for i in note:
inf_Gen_longitude_back.append(longLis[int(i)])
'''
print(inf_Gen_longitude_back)
------------------------------------------------------------Backward latitude
'''
inf_Gen_latitude_back=[]
note=backward(y)
for i in note:
inf_Gen_latitude_back.append(latLis[int(i)])
'''
print(inf_Gen_latitude_back)
------------------------------------------------------------Distance calculation (General vs. Backward)
x=distance(lat1,
lon1,
lat2,
lon2,)
print(x)
'''
h=0
lat_long_result_backward=[]
u=input('Enter Distance Range in meters For Backward Check : ')
for i in range(0,
len(y),):
s=distance(float(inf_Gen_longitude[h]),
float(inf_Gen_latitude[h]),
float(inf_Gen_longitude_back[h]),
float(inf_Gen_latitude_back[h]),
float(u),)
lat_long_result_backward.append(s)
h=h+1
'''
print(lat_long_result_backward)
-----------------------------------from sucessful TRUE extracting the ID Values
'''
f=0
note=backward(y)
imp_ret=[]
for i in lat_long_result_backward:
if i==True:
imp_ret.append(IDLis[note[f]])
f=f+1
else:
f=f+1
'''
print('BACKWARD RESULT')
print(imp_ret)
'''
BACKWARD_RESULT=imp_ret
'''
---------------------------------------------------------------------------
STEPS FOR FORWARD CHECK
-----------------------------------------------------------------------
-----------------------------------------------------------------------
-----------------------------------------------------------------------
-----------------------------------------------------------------------
-----------------------------------------------------------------------
'''
def find(item,
where,):
j=0
note=[]
item=' '+item+' ' # Only applicable for Notepad data
for i in where:
if i==item:
note.append(j)
j=j+1
else:
j=j+1
return(note)
y=find(item,
IDLis,)
#print(y)
def forward(y):
note=[]
for i in y:
note.append(int(i)+1)
return(note)
note=forward(y)
'''
print(note)
------------------------
all values of dates of infected person (General)
'''
inf_Gen_dates=[]
for i in y:
inf_Gen_dates.append(dateLis[int(i)])
#print(inf_Gen_dates)
inf_Gen_time=[]
for i in y:
inf_Gen_time.append(timeLis[int(i)])
'''
print(inf_Gen_time)
--------------------------------
all values of dates of close_match person(Forward)
'''
note=forward(y)
inf_forward_dates=[]
for i in note:
inf_forward_dates.append(dateLis[int(i)])
#print(inf_back_dates)
inf_forward_time=[]
for i in note:
inf_forward_time.append(timeLis[int(i)])
'''
print(inf_back_time)
-------------------------------------------------
Back dates and time matchup with general
'''
h=0
date_time_result=[]
goku=input('Enter Time Difference in minutes for Forward Check : ')
for i in range(0,
len(y),):
s=date_time_match_check(inf_Gen_time[h],
inf_Gen_dates[h],
inf_forward_time[h],
inf_forward_dates[h],
int(goku),)
date_time_result.append(s)
h=h+1
'''
print(date_time_result)
--------------------------------------------------------extracting True Positives for further proceeding
'''
posLis=[]
q=0
for i in date_time_result:
if i=='True':
posLis.append(q)
q=q+1
else:
q=q+1
#print(posLis)
new_lis=[]
for i in posLis:
new_lis.append(y[i])
#print(new_lis)
y=new_lis
#print(y)
'''
--------------------------------------------------
Now we test distance--------------------------------General Distance
'''
inf_Gen_longitude=[]
for i in y:
inf_Gen_longitude.append(longLis[int(i)])
'''
print(inf_Gen_longitude)
-----------------------------------------------------latitude
'''
inf_Gen_latitude=[]
for i in y:
inf_Gen_latitude.append(latLis[int(i)])
'''
print(inf_Gen_latitude)
---------------------------------------------------forward longitude
'''
inf_Gen_longitude_forward=[]
note=forward(y)
for i in note:
inf_Gen_longitude_forward.append(longLis[int(i)])
'''
print(inf_Gen_longitude_back)
--------------------------------------------------forward latitude
'''
inf_Gen_latitude_forward=[]
note=forward(y)
for i in note:
inf_Gen_latitude_forward.append(latLis[int(i)])
'''
print(inf_Gen_latitude_back)
------------------------------------------------------------Distance calculation (General vs Backward)
x=distance(lat1,
lon1,
lat2,
lon2,)
print(x)
'''
h=0
lat_long_result_forward=[]
u=input('Enter Distance Range in meters for Forward Check : ')
for i in range(0,
len(y),):
s=distance(float(inf_Gen_longitude[h]),
float(inf_Gen_latitude[h]),
float(inf_Gen_longitude_forward[h]),
float(inf_Gen_latitude_forward[h]),
float(u),)
lat_long_result_forward.append(s)
h=h+1
'''
print(lat_long_result_backward)
----------------------------------from sucessful TRUE extracting the ID Values
'''
f=0
note=forward(y)
imp_ret=[]
for i in lat_long_result_forward:
if i==True:
imp_ret.append(IDLis[note[f]])
f=f+1
else:
f=f+1
#print('FORWARD RESULT')
#print(imp_ret)
FORWARD_RESULT=imp_ret
result=Union(FORWARD_RESULT, BACKWARD_RESULT)
print('\n')
print(' UNION RESULT of 1st ITERATION Check: ')
print('\n')
print(result)
| db_search_predict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + deletable=true editable=true
# Import libraries and modules
import tensorflow as tf
import numpy as np
import pandas as pd
import shutil
print tf.__version__
print np.__version__
print pd.__version__
np.set_printoptions(threshold=np.inf)
# + deletable=true editable=true
# change these to try this notebook out
BUCKET = 'youtube8m-4-train'
PROJECT = 'qwiklabs-gcp-8d3d0cd07cef9252'
REGION = 'us-central1'
# + deletable=true editable=true
# Import os environment variables
import os
os.environ['BUCKET'] = BUCKET
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
# + [markdown] deletable=true editable=true
# # Local Development
# + deletable=true editable=true
# Set logging verbosity to INFO for richer output
tf.logging.set_verbosity(tf.logging.INFO)
# + deletable=true editable=true
# The number of video classes
NUM_CLASSES = 4716
# + deletable=true editable=true
arguments = {}
arguments["train_file_pattern"] = "gs://youtube-8m-team/1/video_level/train/train*.tfrecord"
arguments["eval_file_pattern"] = "gs://youtube-8m-team/1/video_level/validate/validate-0.tfrecord"
arguments["output_dir"] = "trained_model"
arguments["batch_size"] = 10
arguments["train_steps"] = 100
arguments["hidden_units"] = [1024, 256, 64]
arguments["top_k"] = 5
arguments["start_delay_secs"] = 60
arguments["throttle_secs"] = 30
# + deletable=true editable=true
# Create an input function to read our training and validation data
# Then provide the results to the Estimator API
def read_dataset_video(file_pattern, mode, batch_size):
def _input_fn():
print("\nread_dataset_video: _input_fn: file_pattern = {}".format(file_pattern))
print("read_dataset_video: _input_fn: mode = {}".format(mode))
print("read_dataset_video: _input_fn: batch_size = {}".format(batch_size))
# This function will decode frame examples from the frame level TF Records
def decode_example(serialized_examples):
# Create feature map
feature_map = {
'video_id': tf.FixedLenFeature(shape = [], dtype = tf.string),
'labels': tf.VarLenFeature(dtype = tf.int64),
'mean_rgb': tf.FixedLenFeature(shape = [1024], dtype = tf.float32),
'mean_audio': tf.FixedLenFeature(shape = [128], dtype = tf.float32)
}
# Parse TF Records into our features
features = tf.parse_single_example(serialized = serialized_examples, features = feature_map)
print("\nread_dataset_video: _input_fn: decode_example: features = {}".format(features)) # shape = video_id = (), mean_rgb = (1024,), mean_audio = (128,), labels = SparseTensor object
# Extract and format labels
sparse_labels = features.pop("labels") # SparseTensor object
print("read_dataset_video: _input_fn: decode_example: sparse_labels = {}\n".format(sparse_labels))
labels = tf.cast(x = tf.sparse_to_dense(sparse_indices = sparse_labels.values, output_shape = (NUM_CLASSES,), sparse_values = 1, validate_indices = False), dtype = tf.float32)
print("read_dataset_video: _input_fn: decode_example: labels = {}\n".format(labels)) # shape = (NUM_CLASSES,)
return features, labels
# Create list of files from file pattern
file_list = tf.gfile.Glob(filename = file_pattern)
#print("read_dataset_video: _input_fn: file_list = {}".format(file_list))
# Create dataset from file list
dataset = tf.data.TFRecordDataset(filenames = file_list)
print("read_dataset_video: _input_fn: dataset.TFRecordDataset = {}".format(dataset))
# Decode TF Record dataset examples
dataset = dataset.map(map_func = lambda x: decode_example(serialized_examples = x))
print("read_dataset_video: _input_fn: dataset.map = {}".format(dataset))
# Determine amount of times to repeat file and if we should shuffle based on if we are training or evaluating
if mode == tf.estimator.ModeKeys.TRAIN:
num_epochs = None # read files forever
# Shuffle the dataset within a buffer
dataset = dataset.shuffle(buffer_size = batch_size * 10, seed = None)
print("read_dataset_video: _input_fn: dataset.shuffle = {}".format(dataset))
else:
num_epochs = 1 # read files only once
# Repeat files num_epoch times
dataset = dataset.repeat(count = num_epochs)
print("read_dataset_video: _input_fn: dataset.repeat = {}".format(dataset))
# Group the data into batches
dataset = dataset.batch(batch_size = batch_size)
print("read_dataset_video: _input_fn: dataset.batch = {}".format(dataset))
# Create a iterator and then pull the next batch of features and labels from the example queue
batch_features, batch_labels = dataset.make_one_shot_iterator().get_next()
print("read_dataset_video: _input_fn: batch_features = {}".format(batch_features))
print("read_dataset_video: _input_fn: batch_labels = {}\n".format(batch_labels))
return batch_features, batch_labels
return _input_fn
# + deletable=true editable=true
def try_input_function():
with tf.Session() as sess:
fn = read_dataset_video(file_pattern = "gs://youtube-8m-team/1/video_level/train/train*.tfrecord", mode = tf.estimator.ModeKeys.TRAIN, batch_size = 5)
batch_features, batch_labels = fn()
features, labels = sess.run([batch_features, batch_labels])
print("\ntry_input_function: features = {}".format(features))
print("try_input_function: labels = {}\n".format(labels))
# + deletable=true editable=true
try_input_function()
# + deletable=true editable=true
# Create our model function to be used in our custom estimator
def video_level_model(features, labels, mode, params):
print("\nvideo_level_model: features = {}".format(features))
print("video_level_model: labels = {}".format(labels))
print("video_level_model: mode = {}".format(mode))
# 0. Configure network
# Get dynamic batch size
current_batch_size = tf.shape(features['mean_rgb'])[0]
print("video_level_model: current_batch_size = {}".format(current_batch_size))
# Stack all of the features into a 3-D tensor
combined_features = tf.concat(values = [features['mean_rgb'], features['mean_audio']], axis = 1) # shape = (current_batch_size, 1024 + 128)
print("video_level_model: combined_features = {}".format(combined_features))
# 1. Create the DNN structure now
# Create the input layer to our frame DNN
network = combined_features # shape = (current_batch_size, 1024 + 128)
print("video_level_model: network = combined_features = {}".format(network))
# Add hidden layers with the given number of units/neurons per layer
for units in params['hidden_units']:
network = tf.layers.dense(inputs = network, units = units, activation = tf.nn.relu) # shape = (current_batch_size, units)
print("video_level_model: network = {}, units = {}".format(network, units))
# Connect the final hidden layer to a dense layer with no activation to get the logits
logits = tf.layers.dense(inputs = network, units = NUM_CLASSES, activation = None) # shape = (current_batch_size, NUM_CLASSES)
print("video_level_model: logits = {}".format(logits))
# Select the top k logits in descending order
top_k_logits = tf.nn.top_k(input = logits, k = params['top_k'], sorted = True) # shape = (current_batch_size, top_k)
print("video_level_model: top_k_logits = {}".format(top_k_logits))
# Since this is a multi-class, multi-label problem we will apply a sigmoid, not a softmax, to each logit to get its own probability
probabilities = tf.sigmoid(logits) # shape = (current_batch_size, NUM_CLASSES)
print("video_level_model: probabilities = {}".format(probabilities))
# Select the top k probabilities in descending order
top_k_probabilities = tf.sigmoid(top_k_logits.values) # shape = (current_batch_size, top_k)
print("video_level_model: top_k_probabilities = {}".format(top_k_probabilities))
# Select the top k classes in descending order of likelihood
top_k_classes = top_k_logits.indices # shape = (current_batch_size, top_k)
print("video_level_model: top_k_classes = {}".format(top_k_classes))
# The 0/1 predictions based on a threshold, in this case the threshold is if the probability it greater than random chance
predictions = tf.where(
condition = probabilities > 1.0 / NUM_CLASSES, # shape = (current_batch_size, NUM_CLASSES)
x = tf.ones_like(tensor = probabilities),
y = tf.zeros_like(tensor = probabilities))
print("video_level_model: predictions = {}".format(predictions))
top_k_predictions = tf.where(
condition = top_k_probabilities > 1.0 / NUM_CLASSES, # shape = (current_batch_size, top_k)
x = tf.ones_like(tensor = top_k_probabilities),
y = tf.zeros_like(tensor = top_k_probabilities))
print("video_level_model: top_k_predictions = {}\n".format(top_k_predictions))
# 2. Loss function, training/eval ops
if mode == tf.estimator.ModeKeys.TRAIN or mode == tf.estimator.ModeKeys.EVAL:
# Since this is a multi-class, multi-label problem, we will use sigmoid activation and cross entropy loss
loss = tf.losses.sigmoid_cross_entropy(multi_class_labels = labels, logits = logits)
train_op = tf.contrib.layers.optimize_loss(
loss = loss,
global_step = tf.train.get_global_step(),
learning_rate = 0.01,
optimizer = "Adam")
eval_metric_ops = {
"accuracy": tf.metrics.mean_per_class_accuracy(labels = labels, predictions = predictions, num_classes = NUM_CLASSES)
}
else:
loss = None
train_op = None
eval_metric_ops = None
# 3. Create predictions
predictions_dict = {"logits": top_k_logits.values,
"probabilities": top_k_probabilities,
"predictions": top_k_predictions,
"classes": top_k_classes}
# 4. Create export outputs
export_outputs = {"predict_export_outputs": tf.estimator.export.PredictOutput(outputs = predictions_dict)}
# 5. Return EstimatorSpec
return tf.estimator.EstimatorSpec(
mode = mode,
predictions = predictions_dict,
loss = loss,
train_op = train_op,
eval_metric_ops = eval_metric_ops,
export_outputs = export_outputs)
# + deletable=true editable=true
# Create our serving input function to accept the data at serving and send it in the right format to our custom estimator
def serving_input_fn():
# This function fixes the shape and type of our input strings
def fix_shape_and_type_for_serving(placeholder):
# String split each string in the batch and output the values from the resulting SparseTensors
split_string = tf.map_fn(
fn = lambda x: tf.string_split(source = [placeholder[x]], delimiter=',').values,
elems = tf.range(start = 0, limit = tf.shape(input = placeholder)[0]),
dtype = tf.string) # shape = (batch_size, input_sequence_length)
print("serving_input_fn: fix_shape_and_type_for_serving: split_string = {}".format(split_string))
# Convert each string in the split tensor to float
feature_tensor = tf.string_to_number(string_tensor = split_string, out_type = tf.float32) # shape = (batch_size, input_sequence_length)
print("serving_input_fn: fix_shape_and_type_for_serving: feature_tensor = {}".format(feature_tensor))
return feature_tensor
# This function fixes dynamic shape ambiguity of last dimension so that we will be able to use it in our DNN (since tf.layers.dense require the last dimension to be known)
def get_shape_and_set_modified_shape_2D(tensor, additional_dimension_sizes):
# Get static shape for tensor and convert it to list
shape = tensor.get_shape().as_list()
# Set outer shape to additional_dimension_sizes[0] since we know that this is the correct size
shape[1] = additional_dimension_sizes[0]
# Set the shape of tensor to our modified shape
tensor.set_shape(shape = shape) # shape = (batch_size, additional_dimension_sizes[0])
print("serving_input_fn: get_shape_and_set_modified_shape_2D: tensor = {}, additional_dimension_sizes = {}".format(tensor, additional_dimension_sizes))
return tensor
# Create placeholders to accept the data sent to the model at serving time
feature_placeholders = { # all features come in as a batch of strings, shape = (batch_size,), this was so because of passing the arrays to online ml-engine prediction
'video_id': tf.placeholder(dtype = tf.string, shape = [None]),
'mean_rgb': tf.placeholder(dtype = tf.string, shape = [None]),
'mean_audio': tf.placeholder(dtype = tf.string, shape = [None])
}
print("\nserving_input_fn: feature_placeholders = {}".format(feature_placeholders))
# Create feature tensors
features = {
"video_id": feature_placeholders["video_id"],
"mean_rgb": fix_shape_and_type_for_serving(placeholder = feature_placeholders["mean_rgb"]),
"mean_audio": fix_shape_and_type_for_serving(placeholder = feature_placeholders["mean_audio"])
}
print("serving_input_fn: features = {}".format(features))
# Fix dynamic shape ambiguity of feature tensors for our DNN
features["mean_rgb"] = get_shape_and_set_modified_shape_2D(tensor = features["mean_rgb"], additional_dimension_sizes = [1024])
features["mean_audio"] = get_shape_and_set_modified_shape_2D(tensor = features["mean_audio"], additional_dimension_sizes = [128])
print("serving_input_fn: features = {}\n".format(features))
return tf.estimator.export.ServingInputReceiver(features = features, receiver_tensors = feature_placeholders)
# + deletable=true editable=true
# Create custom estimator's train and evaluate function
def train_and_evaluate(args):
# Create our custome estimator using our model function
estimator = tf.estimator.Estimator(
model_fn = video_level_model,
model_dir = args['output_dir'],
params = {'hidden_units': args['hidden_units'], 'top_k': args['top_k']})
# Create train spec to read in our training data
train_spec = tf.estimator.TrainSpec(
input_fn = read_dataset_video(
file_pattern = args['train_file_pattern'],
mode = tf.estimator.ModeKeys.TRAIN,
batch_size = args['batch_size']),
max_steps = args['train_steps'])
# Create exporter to save out the complete model to disk
exporter = tf.estimator.LatestExporter(name = 'exporter', serving_input_receiver_fn = serving_input_fn)
# Create eval spec to read in our validation data and export our model
eval_spec = tf.estimator.EvalSpec(
input_fn = read_dataset_video(
file_pattern = args['eval_file_pattern'],
mode = tf.estimator.ModeKeys.EVAL,
batch_size = args['batch_size']),
steps = None,
exporters = exporter,
start_delay_secs = args['start_delay_secs'],
throttle_secs = args['throttle_secs'])
# Create train and evaluate loop to train and evaluate our estimator
tf.estimator.train_and_evaluate(estimator = estimator, train_spec = train_spec, eval_spec = eval_spec)
# + deletable=true editable=true
# Run the training job
shutil.rmtree(arguments['output_dir'], ignore_errors = True) # start fresh each time
train_and_evaluate(args = arguments)
# + [markdown] deletable=true editable=true
# # Training
# + [markdown] deletable=true editable=true
# ### Locally
# + deletable=true editable=true
# %bash
OUTDIR=trained_model
# rm -rf $OUTDIR
export PYTHONPATH=$PYTHONPATH:$PWD/trainer
python -m trainer.task \
--train_file_pattern="gs://youtube-8m-team/1/video_level/train/train*.tfrecord" \
--eval_file_pattern="gs://youtube-8m-team/1/video_level/validate/validate-0.tfrecord" \
--output_dir=$OUTDIR \
--batch_size=10 \
--train_steps=100 \
--hidden_units="1024 512 256" \
--top_k=5 \
--job-dir=./tmp
# + [markdown] deletable=true editable=true
# ### GCloud
# + deletable=true editable=true
# %bash
OUTDIR=gs://$BUCKET/youtube_8m_video_level_datasets/trained_model
JOBNAME=job_youtube_8m_video_level_datasets$(date -u +%y%m%d_%H%M%S)
# echo $OUTDIR $REGION $JOBNAME
gsutil -m rm -rf $OUTDIR
gcloud ml-engine jobs submit training $JOBNAME \
--region=$REGION \
--module-name=trainer.task \
--package-path=$PWD/trainer \
--job-dir=$OUTDIR \
--staging-bucket=gs://$BUCKET \
--scale-tier=STANDARD_1 \
--runtime-version=1.5 \
-- \
--train_file_pattern="gs://youtube-8m-team/1/video_level/train/train*.tfrecord" \
--eval_file_pattern="gs://youtube-8m-team/1/video_level/validate/validate-0.tfrecord" \
--output_dir=$OUTDIR \
--batch_size=50 \
--train_steps=10000 \
--hidden_units="1024 512 256" \
--top_k=5 \
--job-dir=$OUTDIR
# -
# ### Hyperparameter tuning
# %writefile hyperparam.yaml
trainingInput:
scaleTier: STANDARD_1
hyperparameters:
hyperparameterMetricTag: accuracy
goal: MAXIMIZE
maxTrials: 30
maxParallelTrials: 1
params:
- parameterName: batch_size
type: INTEGER
minValue: 8
maxValue: 512
scaleType: UNIT_LOG_SCALE
- parameterName: hidden_units
type: CATEGORICAL
categoricalValues: ["64 32", "256 128 16", "64 64 64 8"]
# %bash
OUTDIR=gs://$BUCKET/youtube_8m_video_level_datasets/hyperparam
JOBNAME=job_youtube_8m_video_level_datasets$(date -u +%y%m%d_%H%M%S)
# echo $OUTDIR $REGION $JOBNAME
gsutil -m rm -rf $OUTDIR
gcloud ml-engine jobs submit training $JOBNAME \
--region=$REGION \
--module-name=trainer.task \
--package-path=$PWD/trainer \
--job-dir=$OUTDIR \
--staging-bucket=gs://$BUCKET \
--scale-tier=STANDARD_1 \
--config=hyperparam.yaml \
--runtime-version=1.5 \
-- \
--train_file_pattern="gs://youtube-8m-team/1/video_level/train/train*.tfrecord" \
--eval_file_pattern="gs://youtube-8m-team/1/video_level/validate/validate-0.tfrecord" \
--output_dir=$OUTDIR \
--train_steps=10000 \
--top_k=5 \
--job-dir=$OUTDIR
# + [markdown] deletable=true editable=true
# # Deploy
# + deletable=true editable=true
# %bash
MODEL_NAME="youtube_8m_video_level_datasets"
MODEL_VERSION="v1"
MODEL_LOCATION=$(gsutil ls gs://$BUCKET/youtube_8m_video_level_datasets/trained_model/export/exporter/ | tail -1)
# echo "Deleting and deploying $MODEL_NAME $MODEL_VERSION from $MODEL_LOCATION ... this will take a few minutes"
#gcloud ml-engine versions delete ${MODEL_VERSION} --model ${MODEL_NAME}
#gcloud ml-engine models delete ${MODEL_NAME}
gcloud ml-engine models create $MODEL_NAME --regions $REGION
gcloud ml-engine versions create $MODEL_VERSION --model $MODEL_NAME --origin $MODEL_LOCATION --runtime-version 1.5
# + [markdown] deletable=true editable=true
# # Prediction
# + [markdown] deletable=true editable=true
# ### Prep
# + deletable=true editable=true
# Let's call our input function to decode our data to put into BigQuery for testing predictions
video_list = []
with tf.Session() as sess:
fn = read_dataset_video(
file_pattern = "gs://youtube-8m-team/1/video_level/validate/validate-0.tfrecord",
mode = tf.estimator.ModeKeys.EVAL,
batch_size = 1)
batch_features, batch_labels = fn()
for key,value in batch_features.items():
batch_features[key] = tf.squeeze(batch_features[key])
fixed_batch_features = batch_features
fixed_batch_labels = tf.squeeze(batch_labels)
while True:
features, labels = sess.run([fixed_batch_features, fixed_batch_labels])
features["labels"] = labels
video_list.append(features)
# + deletable=true editable=true
# This is the number of videos from the video level file we just processed
len(video_list)
# + deletable=true editable=true
# Convert the nd-arrays to lists and cast to strings (video_id is already a single string)
for items in video_list:
items["labels"] = str(items["labels"].tolist())
items["mean_rgb"] = str(items["mean_rgb"].tolist())
items["mean_audio"] = str(items["mean_audio"].tolist())
# + deletable=true editable=true
# Create a dataframe from the list
video_df = pd.DataFrame(video_list)
video_df = video_df[["video_id", "mean_rgb", "mean_audio", "labels"]]
video_df.head()
# + deletable=true editable=true
# Export dataframe to BigQuery
import datalab.bigquery as bq
bigquery_dataset_name = 'ryan_youtube'
bigquery_table_name = 'tbl_video_level'
# Define BigQuery dataset and table\n",
dataset = bq.Dataset(bigquery_dataset_name)
table = bq.Table(bigquery_dataset_name + '.' + bigquery_table_name)
# Create BigQuery dataset
if not dataset.exists():
dataset.create()
# Create or overwrite the existing table if it exists\n",
table_schema = bq.Schema.from_dataframe(video_df)
table.create(schema = table_schema, overwrite = True)
video_df.to_gbq(destination_table = bigquery_dataset_name + '.' + bigquery_table_name, project_id = "qwiklabs-gcp-8d3d0cd07cef9252", if_exists = "replace")
# + deletable=true editable=true
# Create SQL query
query="""
SELECT
video_id,
mean_rgb,
mean_audio
FROM
`qwiklabs-gcp-8d3d0cd07cef9252.ryan_youtube.tbl_video_level`
LIMIT
3
"""
# + deletable=true editable=true
# Export BigQuery results to dataframe
import google.datalab.bigquery as bq2
df_predict = bq2.Query(query).execute().result().to_dataframe()
df_predict.head()
# + [markdown] deletable=true editable=true
# ### Local prediction from local model
# + deletable=true editable=true
# Format dataframe to new line delimited json strings and write out to json file
with open('video_level.json', 'w') as outfile:
for idx, row in df_predict.iterrows():
json_string = "{\"video_id\": \"" + row["video_id"] + "\", \"mean_rgb\": \"" + row["mean_rgb"].replace(" ","").replace("[","").replace("]","") + "\", \"mean_audio\": \"" + row["mean_audio"].replace(" ","").replace("[","").replace("]","") + "\"}"
outfile.write("%s\n" % json_string)
# + deletable=true editable=true
# %bash
model_dir=$(ls $PWD/trained_model/export/exporter | tail -1)
gcloud ml-engine local predict \
--model-dir=$PWD/trained_model/export/exporter/$model_dir \
--json-instances=./video_level.json
# + [markdown] deletable=true editable=true
# ### GCloud ML-Engine prediction from deployed model
# + deletable=true editable=true
# Format dataframe to instances list to get sent to ML-Engine
instances = [{"video_id": row["video_id"], "mean_rgb": row["mean_rgb"].replace(" ","").replace("[","").replace("]",""), "mean_audio": row["mean_audio"].replace(" ","").replace("[","").replace("]","")} for idx, row in df_predict.iterrows()]
# + deletable=true editable=true
# Send instance dictionary to receive response from ML-Engine for online prediction
from googleapiclient import discovery
from oauth2client.client import GoogleCredentials
import json
credentials = GoogleCredentials.get_application_default()
api = discovery.build('ml', 'v1', credentials=credentials)
request_data = {"instances": instances}
parent = 'projects/%s/models/%s/versions/%s' % (PROJECT, 'youtube_8m_video_level_datasets', 'v1')
response = api.projects().predict(body = request_data, name = parent).execute()
print("response = {}".format(response))
# + deletable=true editable=true
| courses/machine_learning/asl/open_project/ASL_youtube8m_models/video_using_datasets/youtube_8m_video_level_datasets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="Ene-8n_Nv4gS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 459} outputId="56ee3177-6951-45b0-9294-883d9f98b37f"
# !pip install split-folders tqdm
# !pip install -U -q kaggle
# !apt install pv
# !pip install -U -q PyDrive
# + id="jYepCM3QwBSs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} outputId="ea19a5e5-8289-448a-f80c-2a98beae4295"
import pandas as pd
import numpy as np
import os
import keras
import matplotlib.pyplot as plt
import splitfolders
from keras.layers import Dense,GlobalAveragePooling2D
from keras.applications import MobileNet
from keras.preprocessing import image
from keras.applications.mobilenet import preprocess_input
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Model
from keras.optimizers import Adam
from google.colab import files
import seaborn as sns
import matplotlib.pyplot as plt
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
# + [markdown] id="h_3jBou7oTLO" colab_type="text"
# ## Get the dataset and split it into test train and eval sets
# + id="LjDtV8iLwUtK" colab_type="code" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 89} outputId="e98660b5-0304-40e1-82ad-85d845269f5e"
files.upload()
# + id="QbvoE_vUo51g" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="f58b16d7-4dc8-4056-abc4-0f159fc1ce5d"
# !mkdir -p ~/.kaggle
# !cp kaggle.json ~/.kaggle/
# !kaggle datasets download -d iarunava/cell-images-for-detecting-malaria
# + id="xPJzUVt1wl6N" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="789e8a14-f1c5-4fa0-839c-d5941f57acd0"
# !unzip -o /content/cell-images-for-detecting-malaria.zip| pv -l >/dev/null
os.remove('cell-images-for-detecting-malaria.zip')
# + id="Sz8SYP4cw1K2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="27458ec7-36dd-4539-9986-27f47fb94472"
splitfolders.ratio("/content/cell_images/cell_images", output="/content/cell_images/cell_images_split", seed=1337, ratio=(.8, .1, .1), group_prefix=None)
# + [markdown] id="LisKjzU_yjk8" colab_type="text"
# ## Generate the model using TensorFlow & Keras
# + id="sa-d07JPyvMX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="21accb68-d57a-4334-f2ff-32efeea39387"
base_model=MobileNet(weights='imagenet',include_top=False) #imports the mobilenet model and discards the last 1000 neuron layer.
x=base_model.output
x=GlobalAveragePooling2D()(x)
x=Dense(1024,activation='relu')(x) #we add dense layers so that the model can learn more complex functions and classify for better results.
x=Dense(1024,activation='relu')(x) #dense layer 2
x=Dense(512,activation='relu')(x) #dense layer 3
preds=Dense(2,activation='softmax')(x) #final layer with softmax activation
# + id="XQKKxRZbyzEF" colab_type="code" colab={}
model=Model(inputs=base_model.input,outputs=preds)
for layer in model.layers[:20]:
layer.trainable=False
for layer in model.layers[20:]:
layer.trainable=True
# + id="GOS64-_Vy6_U" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 714} outputId="9329982c-478e-4512-de35-7129586c3f5c"
train_datagen= ImageDataGenerator(preprocessing_function=preprocess_input) #included in our dependencies
train_generator=train_datagen.flow_from_directory('/content/cell_images/cell_images_split/train',
target_size=(224,224),
color_mode='rgb',
batch_size=32,
class_mode='categorical', shuffle=True)
model.compile(optimizer='Adam',loss='categorical_crossentropy',metrics=['accuracy'])
# Adam optimizer
# loss function will be categorical cross entropy
# evaluation metric will be accuracy
step_size_train=train_generator.n//train_generator.batch_size
hist = model.fit_generator(generator=train_generator,
steps_per_epoch=step_size_train,
epochs=20)
# + [markdown] id="BzvyYLtAwt3D" colab_type="text"
# ## Saving the Model
# + id="_LyZwa_owtlf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="499a350a-7e8e-4130-b9ec-a333866c5dcc"
# 1. Authenticate and create the PyDrive client.
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
# 2. Save Keras Model or weights on google drive
# create on Colab directory
model.save('model.h5')
model_file = drive.CreateFile({'title' : 'model.h5'})
model_file.SetContentFile('model.h5')
model_file.Upload()
# download to google drive
drive.CreateFile({'id': model_file.get('id')})
# + [markdown] id="oNFYU2qY1SlK" colab_type="text"
# ## Evaluating the model
# + id="eG69IAOnCb4R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 573} outputId="13be1172-f18d-4d96-c1f6-609fdd5917f1"
def plotLearningCurve(hist,epochs):
epochRange = range(1,epochs+1)
plt.plot(epochRange,hist.history['accuracy'])
plt.title('Model Accuracy')
plt.xlabel('Epoch')
plt.ylabel('Accuracy')
plt.legend(['Train'],loc='upper left')
plt.show()
plt.plot(epochRange,hist.history['loss'])
plt.title('Model Loss')
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.legend(['Train'],loc='upper left')
plt.show()
plotLearningCurve(hist,20)
| Malaria_Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/milayacharlieCvSU/OOP-1-1/blob/main/Python_Classes_and_Objects.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="l2tD3expzX6q"
# Class
# + id="2Da_NsBuzOiJ"
# Class without Attributes and methods
class MyClass:
pass
# + colab={"base_uri": "https://localhost:8080/"} id="HS0PC1Rnztoh" outputId="4b04407d-7dc5-4d03-cf42-f4cd606811ec"
# Creat a class with Attributes and Methods
class MyClass:
def __init__(self, name, age):
# Attributes
self.name = name
self.age = age
# Method
def display(self):
print(self.name, self.age, "years old")
# Object Name Creation
person = MyClass("<NAME>", 18)
person.display()
# + colab={"base_uri": "https://localhost:8080/"} id="G3w_A8zi0AAq" outputId="5648f1d0-f96c-4f71-f2e6-c8119e428b13"
# Application 1 - Write a Python program involving classes that computes for
# the area of a rectangle. Formula: A = l*w
class Rectangle:
def __init__(self, length, width):
# Attibutes
self.length = length
self.width = width
# Method
def area(self):
print(self.length * self.width)
rect1 = Rectangle(7, 3)
rect1.area()
| Python_Classes_and_Objects.ipynb |
# # Test Package Command
# %load_ext literary.module
# +
import logging
from concurrent.futures import ProcessPoolExecutor
from pathlib import Path
from traitlets import Int, List, Unicode, default
from ..core.test import ProjectTester
from .app import LiteraryApp
# -
class LiteraryTestApp(LiteraryApp):
description = "Test literary notebooks in parallel"
aliases = {
**LiteraryApp.aliases,
"ignore": "ProjectTester.ignore_patterns",
"jobs": "ProjectTester.jobs",
"packages": "ProjectTester.packages_dir",
"extras": "ProjectTester.extra_sources"
}
source = List(trait=Unicode(help="source directory or notebooks to run")).tag(
config=True
)
jobs = Int(
allow_none=True, default_value=None, help="number of parallel jobs to run"
).tag(config=True)
ignore = List(help="glob pattern to ignore during recursion", trait=Unicode()).tag(
config=True
)
def start(self):
tester = ProjectTester(parent=self)
tester.run()
| src/literary/commands/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Frequently Used Commands
# ### Shift+Enter to execute code in a cell
print('Hello World!')
# 
# ### Other commands
# - Esc-a: adds an extra cell _above_ the current one
# - Pressing a after this sequence adds more cells
# - Esc-b: adds an extra cell _below_ the current one
# - Pressing b after this sequence adds more cells
# - Esc-x: Deletes current cell
import matplotlib.pyplot as plt
# %matplotlib inline
# ### Show Tab Completion and Shift-Tab Parameter Context Help
# +
x = [0,82.0442626658045,164.140532825633,249.101916493353,332.182520454561,416.254670963008,495.330844802651,575.152803567278,653.205144040749,738.002602107511,818.027625347191,898.118246791891,980.079475188722,1061.14169245002,1139.27372549306,1222.46124841828,1300.35875796241,1375.56916894815,1450.58381247379,1528.61926864683,1605.58397275636,1690.73526419816,1765.8607155748,1844.76374052086,1927.68455634244,2007.86892722346,2086.83606915306,2166.71913598734,2251.80135023691,2335.70816516882,2420.79263834332,2499.86354911873,2577.05890165442,2654.19208140119,2730.09283946298,2809.07577275313,2894.08624312766,2969.05670691318,3049.93958564477,3129.75861740531,3206.68277248806,3283.60346903165,3364.58552562287,3449.40362446521,3525.4406380069,3608.13685182861,3690.05929028829,3773.14656467608,3850.22786804579,3930.30737684189,4015.46434790825,4093.50879464788,4172.64841837878,4252.66450124259,4335.57956650011,4420.52164467734,4495.52991250898,4575.39742270308,4650.49642074232,4730.52574677055,4805.68851119975,4880.60019974235,4960.70865715944,5043.67168991259,5118.73724644963,5203.54753389807,5283.38430140739,5359.51383129058,5438.3504224767,5519.19106748314,5597.38879699402]
y = [69,70.5331166725557,71.5580608629997,71.7925736363559,71.3468902075964,70.8280365103217,70.6414982437025,70.4451053764375,70.3795952412711,69.7645648576213,69.5426154537094,69.3004779067882,68.9026747371149,68.599154906399,68.519069857217,68.3772433121902,69.1486631584102,68.7548264957532,68.3609585242701,68.5575021969504,68.5786288310157,70.1801398605569,69.7914710297875,70.1491959166127,71.3308639026079,71.9361568322099,71.7841530239482,71.5632100706621,70.9532872882161,70.4120650812618,69.7956890967353,69.6193830861858,69.6154149236862,69.6130427165012,69.7102463368457,69.5264209638887,68.9050613087613,69.1051712486311,68.8193037564598,68.6066819000086,68.6092749335593,68.6204785117934,68.3360360051156,68.5812312774717,68.3713770000006,69.5440095449469,70.5260768620176,72.0285425812187,72.0349254824144,71.8158332177273,71.1892654017154,71.1169346409977,70.8948191898231,70.6895661675202,70.2298913978404,69.6255849641529,69.7340391269025,69.5070018796742,69.6617353391113,69.4627544694931,69.630774569752,69.8026157834872,69.5269217792645,69.0549816241028,69.1984712324886,68.3562401007611,68.1422906748129,68.2062726166005,68.0498033914326,68.6018257745021,68.7606159741864]
plt.plot()
#plt.plot(x,y)
plt.show()
# -
# # Jupyter Shutdown
# 
# 
# 
# # Commenting
# This is a single-line comment
print('Hello World!') # Rest of the line will be ignored
""" This is a multiple-line comment just in
case the comment is very long """
print('Hello World!')
# # Markdown
# This notebook gives some examples of Jupyter markdown language.
#
# Don't worry that you might not understand all the code in this notebook, yet. It's main prupose is to show you what markdown is possible and to provide a template.
# # Preliminary Jupyter Environment Set-up
# The code below loads the required Python packages into the notebook.
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# # Data Input
# The code below reads data in from a text file and loads it into a DataFrame object as defined by the pandas package. In order for this cell to be executed successfully, the CSV file must be located in the same file folder as the Jupyter notebook.
df_oz = pd.read_csv('ozone.csv')
# Here is a quick look at the top five rows of the data:
df_oz.head()
# # Analysis
# Analysis can take the form of graphing as demonstrated below. This first graph shows the relationship of Wind Speed and Ozone level.
""" Format of the scatterplot method is as follows: ax.scatter(x-series, y-series) """
fig, ax = plt.subplots()
ax.scatter(df_oz['wind'], df_oz['ozone'], alpha=0.5) # the 'alpha' parameter controls dot opacity
# alpha is a parameter that controls the transparency of the dots: 1 = solid, <1 = various transparency levels, 0 = no mark
fig.suptitle('Ozone vs. Wind Speed')
ax.xaxis.set_label_text('Wind Speed')
ax.yaxis.set_label_text('Ozone')
fig.set_size_inches(7,5)
plt.show()
# This next graph uses a Python package called Seaborn to investigate the relationships of multiple variables simulataneously.
sns.set(style="ticks", color_codes=True)
g = sns.pairplot(df_oz)
g.savefig('ozone.jpg')
# This cell really has nothing to do with the preceding analysis, but it is simply an example of including *LaTex* typesetting language into a jupyter notebook.
#
# $\sum_{i=0}^{n}{i} = \frac{n(n+1)}{2}$
# # Conclusions
# This cell, mostly, is just a demonstration of bullet points of various formats:
#
# - Ozone and Wind Speed are negatively correlated
#
# * Ozone and Temperature are positively correlated
#
# - Wind Speed and Radiation appear not to be correlated
# - The scatter plot is not tightly grouped around any postiviely or negatively-sloped line
# # You Can Include Images
# We will use some data about these three species of Iris.
# 
# # References for Markdown
# The following links provide cheatsheets on Jupyter markdown language.
# [Link 1](https://beegit.com/markdown-cheat-sheet)
#
# [Link 2](https://medium.com/ibm-data-science-experience/markdown-for-jupyter-notebooks-cheatsheet-386c05aeebed)
#
# [Link 3](https://datascience.ibm.com/docs/content/analyze-data/markd-jupyter.html)
#
# [Link 4](http://jupyter-notebook.readthedocs.io/en/stable/examples/Notebook/Working%20With%20Markdown%20Cells.html)
#
# [Link 5](https://github.com/adam-p/markdown-here/wiki/Markdown-Cheatsheet)
#
# [Link 6](http://nestacms.com/docs/creating-content/markdown-cheat-sheet)
| HomeworkAssignment/JupyterIntroduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import division, print_function, absolute_import
import collections
import os
import random
import urllib
import zipfile
import numpy as np
import tensorflow as tf
# +
# Training Parameters.
learning_rate = 0.1
batch_size = 128
num_steps = 3000000
display_step = 10000
eval_step = 200000
# Evaluation Parameters.
eval_words = ['five', 'of', 'going', 'hardware', 'american', 'britain']
# Word2Vec Parameters.
embedding_size = 200 # Dimension of the embedding vector.
max_vocabulary_size = 50000 # Total number of different words in the vocabulary.
min_occurrence = 10 # Remove all words that does not appears at least n times.
skip_window = 3 # How many words to consider left and right.
num_skips = 2 # How many times to reuse an input to generate a label.
num_sampled = 64 # Number of negative examples to sample.
# -
# Download a small chunk of Wikipedia articles collection.
url = 'http://mattmahoney.net/dc/text8.zip'
data_path = 'text8.zip'
if not os.path.exists(data_path):
print("Downloading the dataset... (It may take some time)")
filename, _ = urllib.urlretrieve(url, data_path)
print("Done!")
# Unzip the dataset file. Text has already been processed.
with zipfile.ZipFile(data_path) as f:
text_words = f.read(f.namelist()[0]).lower().split()
# +
# Build the dictionary and replace rare words with UNK token.
count = [('UNK', -1)]
# Retrieve the most common words.
count.extend(collections.Counter(text_words).most_common(max_vocabulary_size - 1))
# Remove samples with less than 'min_occurrence' occurrences.
for i in range(len(count) - 1, -1, -1):
if count[i][1] < min_occurrence:
count.pop(i)
else:
# The collection is ordered, so stop when 'min_occurrence' is reached.
break
# Compute the vocabulary size.
vocabulary_size = len(count)
# Assign an id to each word.
word2id = dict()
for i, (word, _)in enumerate(count):
word2id[word] = i
data = list()
unk_count = 0
for word in text_words:
# Retrieve a word id, or assign it index 0 ('UNK') if not in dictionary.
index = word2id.get(word, 0)
if index == 0:
unk_count += 1
data.append(index)
count[0] = ('UNK', unk_count)
id2word = dict(zip(word2id.values(), word2id.keys()))
print("Words count:", len(text_words))
print("Unique words:", len(set(text_words)))
print("Vocabulary size:", vocabulary_size)
print("Most common words:", count[:10])
# -
data_index = 0
# Generate training batch for the skip-gram model.
def next_batch(batch_size, num_skips, skip_window):
global data_index
assert batch_size % num_skips == 0
assert num_skips <= 2 * skip_window
batch = np.ndarray(shape=(batch_size), dtype=np.int32)
labels = np.ndarray(shape=(batch_size, 1), dtype=np.int32)
# get window size (words left and right + current one).
span = 2 * skip_window + 1
buffer = collections.deque(maxlen=span)
if data_index + span > len(data):
data_index = 0
buffer.extend(data[data_index:data_index + span])
data_index += span
for i in range(batch_size // num_skips):
context_words = [w for w in range(span) if w != skip_window]
words_to_use = random.sample(context_words, num_skips)
for j, context_word in enumerate(words_to_use):
batch[i * num_skips + j] = buffer[skip_window]
labels[i * num_skips + j, 0] = buffer[context_word]
if data_index == len(data):
buffer.extend(data[0:span])
data_index = span
else:
buffer.append(data[data_index])
data_index += 1
# Backtrack a little bit to avoid skipping words in the end of a batch.
data_index = (data_index + len(data) - span) % len(data)
return batch, labels
# +
# Ensure the following ops & var are assigned on CPU
# (some ops are not compatible on GPU).
with tf.device('/cpu:0'):
# Create the embedding variable (each row represent a word embedding vector).
embedding = tf.Variable(tf.random.normal([vocabulary_size, embedding_size]))
# Construct the variables for the NCE loss.
nce_weights = tf.Variable(tf.random.normal([vocabulary_size, embedding_size]))
nce_biases = tf.Variable(tf.zeros([vocabulary_size]))
def get_embedding(x):
with tf.device('/cpu:0'):
# Lookup the corresponding embedding vectors for each sample in X.
x_embed = tf.nn.embedding_lookup(embedding, x)
return x_embed
def nce_loss(x_embed, y):
with tf.device('/cpu:0'):
# Compute the average NCE loss for the batch.
y = tf.cast(y, tf.int64)
loss = tf.reduce_mean(
tf.nn.nce_loss(weights=nce_weights,
biases=nce_biases,
labels=y,
inputs=x_embed,
num_sampled=num_sampled,
num_classes=vocabulary_size))
return loss
# Evaluation.
def evaluate(x_embed):
with tf.device('/cpu:0'):
# Compute the cosine similarity between input data embedding and every embedding vectors
x_embed = tf.cast(x_embed, tf.float32)
x_embed_norm = x_embed / tf.sqrt(tf.reduce_sum(tf.square(x_embed)))
embedding_norm = embedding / tf.sqrt(tf.reduce_sum(tf.square(embedding), 1, keepdims=True), tf.float32)
cosine_sim_op = tf.matmul(x_embed_norm, embedding_norm, transpose_b=True)
return cosine_sim_op
# Define the optimizer.
optimizer = tf.optimizers.SGD(learning_rate)
# -
# Optimization process.
def run_optimization(x, y):
with tf.device('/cpu:0'):
# Wrap computation inside a GradientTape for automatic differentiation.
with tf.GradientTape() as g:
emb = get_embedding(x)
loss = nce_loss(emb, y)
# Compute gradients.
gradients = g.gradient(loss, [embedding, nce_weights, nce_biases])
# Update W and b following gradients.
optimizer.apply_gradients(zip(gradients, [embedding, nce_weights, nce_biases]))
# +
# Words for testing.
x_test = np.array([word2id[w] for w in eval_words])
# Run training for the given number of steps.
for step in xrange(1, num_steps + 1):
batch_x, batch_y = next_batch(batch_size, num_skips, skip_window)
run_optimization(batch_x, batch_y)
if step % display_step == 0 or step == 1:
loss = nce_loss(get_embedding(batch_x), batch_y)
print("step: %i, loss: %f" % (step, loss))
# Evaluation.
if step % eval_step == 0 or step == 1:
print("Evaluation...")
sim = evaluate(get_embedding(x_test)).numpy()
for i in xrange(len(eval_words)):
top_k = 8 # number of nearest neighbors.
nearest = (-sim[i, :]).argsort()[1:top_k + 1]
log_str = '"%s" nearest neighbors:' % eval_words[i]
for k in xrange(top_k):
log_str = '%s %s,' % (log_str, id2word[nearest[k]])
print(log_str)
| Tutorials/TensorFlow_V2/notebooks/2_BasicModels/word2vec.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #
#
#
#
#
#
#
#
# # Group Project March 2021
# ReGeneration Academy on Big Data & Artificial Intelligence (powered by Microsoft) | A case study for predicting the price of an Airbnb listing in Athens using Microsoft Azure
# ## 5 Detailed Objectives
# ### 5.4 Modelling
#
# + pycharm={"name": "#%%\n"}
import pandas as pd
import numpy as np
import shap as shap
import xgboost
from sklearn.linear_model import LogisticRegression, Ridge,LinearRegression
from sklearn.metrics import accuracy_score, mean_absolute_error, r2_score, mean_squared_error
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.neighbors import KNeighborsClassifier, KNeighborsRegressor
from sklearn.preprocessing import MinMaxScaler, StandardScaler, OneHotEncoder
import matplotlib.pyplot as plt
from xgboost import XGBRegressor
from helper_functions.clearAmenity import *
from helper_functions.cleanRows import *
from helper_functions.DataFrames import *
from helper_functions.cleanBinary import *
from helper_functions.fill_beds import *
from helper_functions.fill_bedrooms import *
from helper_functions.LabelEncoding import *
from helper_functions.functionKeyword import *
from helper_functions.FillNaNValues import *
import warnings
warnings.filterwarnings('ignore')
from scipy import stats
import xgboost as xgb
# + pycharm={"name": "#%%\n"}
clean_data =pd.read_csv('csv_creation/final_clean.csv',index_col=0)
#filtering outlier prices
clean_data=clean_data[clean_data['price']<200]
clean_data
# + pycharm={"name": "#%%\n"}
#initialize X,y
y=clean_data["price"]
X=clean_data.drop(['price'],axis=1)
#Splitting dataset
X_train, X_test, y_train, y_test = train_test_split(X,y, random_state=1)
#Scaling X_train,X_test
sc = StandardScaler()
sc.fit(X_train)
X_train_scaled = sc.transform(X_train)
X_test_scaled = sc.transform(X_test)
# + pycharm={"name": "#%%\n"}
print(X_train.shape)
print(X_test.shape)
#print(selCols)
# + pycharm={"name": "#%%\n"}
#-------------------------------KNeighborsRegressor-------------------------------#
KNeighborsRegr = KNeighborsRegressor(n_neighbors=2)
# fit model
KNeighborsRegr.fit(X_train_scaled, y_train)
# make predictions
preds = KNeighborsRegr.predict(X_test_scaled)
print("Train Accuracy:",KNeighborsRegr.score(X_train_scaled, y_train))
print("Test Accuracy:",KNeighborsRegr.score(X_test_scaled, y_test))
# evaluate predictions
mae = mean_absolute_error(y_test, preds)
r2 = r2_score(y_test, preds)
mse = mean_squared_error(y_test, preds)
print('MAE: %.3f' % mae)
print('r2: %.3f' % r2)
print('MSE: %.3f' % mse)
# + pycharm={"name": "#%%\n"}
#-------------------------------KNeighborsRegressor CV-------------------------------
# scores = cross_val_score(KNeighborsRegr, X_train_scaled, y_train, cv=5)
# #printing the averaged score over the 5 Kfolds
# print("Accuracy: %.3f%% (%.3f%%)" % (scores.mean()*100.0, scores.std()*100.0))
# + pycharm={"name": "#%%\n"}
#-------------------------------LinearRegression-------------------------------#
regression = LinearRegression(fit_intercept = True)
# fit model
regression.fit(X_train_scaled, y_train)
# make predictions
preds = regression.predict(X_test_scaled)
print("Train Accuracy:",regression.score(X_train_scaled, y_train))
print("Test Accuracy:",regression.score(X_test_scaled, y_test))
# evaluate predictions
mae = mean_absolute_error(y_test, preds)
r2 = r2_score(y_test, preds)
mse = mean_squared_error(y_test, preds)
print('MAE: %.3f' % mae)
print('r2: %.3f' % r2)
print('MSE: %.3f' % mse)
# + pycharm={"name": "#%%\n"}
#-------------------------------Ridge-------------------------------#
from sklearn.linear_model import Ridge
ridge = Ridge(fit_intercept = True)
# fit model
ridge.fit(X_train_scaled, y_train)
# make predictions
preds = ridge.predict(X_test_scaled)
print("Train Accuracy:",ridge.score(X_train_scaled, y_train))
print("Test Accuracy:",ridge.score(X_test_scaled, y_test))
# evaluate predictions
mae = mean_absolute_error(y_test, preds)
r2 = r2_score(y_test, preds)
mse = mean_squared_error(y_test, preds)
print('MAE: %.3f' % mae)
print('r2: %.3f' % r2)
print('MSE: %.3f' % mse)
# + pycharm={"name": "#%%\n"}
#-------------------------------XGBRegressor-------------------------------#
# Create instace
xgb_reg = xgb.XGBRegressor()
# Fit the model on training data
xgb_reg.fit(X_train_scaled, y_train)
# Predict
training_preds_xgb_reg = xgb_reg.predict(X_train_scaled)
# Validate
val_preds_xgb_reg = xgb_reg.predict(X_test_scaled)
# evaluate predictions
mae = mean_absolute_error(y_test, val_preds_xgb_reg)
r2 = r2_score(y_test, val_preds_xgb_reg)
mse = mean_squared_error(y_test, val_preds_xgb_reg)
print('MAE: %.3f' % mae)
print('r2: %.3f' % r2)
print('MSE: %.3f' % mse)
# + pycharm={"name": "#%%\n"}
clf = XGBRegressor(n_estimators=100)
clf.fit(X_train_scaled,
y_train,
eval_metric='mae',
verbose=True,
eval_set=[(X_test_scaled, y_test)],
early_stopping_rounds=50
)
clf.get_params()
fig,ax=plt.subplots(figsize=(12,50))
xgboost.plot_importance(clf,ax=ax)
# + pycharm={"name": "#%%\n"}
#some plots of feature importance
shap.initjs()
explainer = shap.TreeExplainer(clf)
explainer.expected_value
p_df = pd.DataFrame()
p_df['pred'] = clf.predict(X_test_scaled)
p_df['price'] = y_test.reset_index(drop=True)
p_df['mae'] = p_df[['pred','price']].apply(lambda x:abs(x[1]-x[0]),axis=1)
p_df[p_df['mae']<5].head()
shap_values = explainer.shap_values(X_test_scaled)
fig =shap.summary_plot(shap_values, X_test_scaled,show=False)
plt.savefig('sometest.png')
| Deliverable_04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
n=int(input())
arr=[int(x) for x in input().split()]
def binarySearch(arr,element):
start = 0
end = len(arr)-1
while(start<=end):
mid = (start+end)//2
if(arr[mid] == element):
return mid
elif(arr[mid] < element):
start = mid + 1
else:
end = mid - 1
return -1
index = binarySearch(arr,n)
print(index)
# +
def binarySearch(arr,element):
start = 0
end = len(arr)-1
while(start<=end):
mid = (start+end)//2
if(arr[mid] == element):
return mid
elif(arr[mid] < element):
start = mid + 1
else:
end = mid - 1
return -1
arr = [1,3,8,9,11,13,70,89,98]
index = binarySearch(arr,3)
print(index)
# +
def binarySearch(arr,element):
start = 0
end = len(arr)-1
while(start<=end):
mid = (start+end)//2
if(arr[mid] == element):
return mid
elif(arr[mid] < element):
start = mid + 1
else:
end = mid - 1
return -1
arr = [1,3,8,9,11,13,70,89,98]
index = binarySearch(arr,90)
print(index)
# -
| random/binarySearch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from groupdocstranslationcloud.configuration import Configuration
from groupdocstranslationcloud.api.translation_api import TranslationApi
from groupdocstranslationcloud.api.storage_api import StorageApi
from groupdocstranslationcloud.models.translate_text import TranslateText
from groupdocstranslationcloud.models.translate_document import TranslateDocument
# ## Please enter valid client_secret and client_id to run this demo.
#
# ## See documentation for details: https://docs.groupdocs.cloud/total/creating-and-managing-application/
configuration = Configuration(client_secret="", client_id="")
translation_api = TranslationApi(configuration)
storage_api = StorageApi(configuration)
# ## Plain text translation: setting parameters
pair = "en-fr"
text = "Welcome to Paris"
# ## Plain text translation: translating text
translator = TranslateText(pair, text)
request = translator.to_string()
res = translation_api.post_translate_text(request)
print(res.translation)
# ## Document translation: setting parameters
# +
upload_path = "test.docx"
download_path = "translated.docx"
pair = "en-fr"
_format = "docx"
outformat = "docx"
storage = "First Storage"
name = "test.docx"
folder = ""
savepath = ""
savefile = "translated.docx"
masters = False
elements = []
# -
# ## Document translation: uploading document
upload_res = storage_api.upload_file(name, upload_path)
print("File uploaded")
# ## Document translation: translating document
translator = TranslateDocument(pair, _format, outformat, storage, name, folder, savepath, savefile, masters, elements)
request = translator.to_string()
res = translation_api.post_translate_document(request)
print(res.message)
| demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
def metadata(titulo, ejex, ejey):
plt.title(titulo, fontsize = 60, fontweight='bold')
plt.xlabel(ejex, fontsize = 35)
plt.ylabel(ejey, fontsize = 35)
plt.xticks(fontsize = 25)
plt.yticks(fontsize = 25)
return
def cifras_signif(i, cifras):
return str('{:g}'.format(float('{:.{p}g}'.format(i, p = cifras))))
def umbral(a, sigma, ro, d):
umbral = a * ((sigma-ro)*9.8*d/ro)**(1/2)
return umbral
def diametro(a, sigma, ro, umbral):
# Umbral en m/s
diam = (umbral/a)**2 * ro/(9.8*(sigma-ro))
return diam
datos = pd.read_csv(
'C:\\Users\\nahue\\Desktop\\Agro 2\\Tp_5\\Ej_4.txt',
delimiter = ' ',
decimal = '.',
)
datos.head()
datos['24km'] = datos['Ul']/100 * 24
datos['54km'] = datos['Ul']/100 * 54
datos.head(10)
plt.figure(figsize = (18, 9))
metadata('Viento luego de la barrera', 'Distancia (H)', 'Velocidad (Km/h)')
plt.plot(datos['Distancia[H]'], datos['24km'])
plt.plot(datos['Distancia[H]'], datos['54km'])
plt.legend(['24 km/h', '54 km/h'], fontsize = 15,
title = 'Viento antes de la barrera', title_fontsize = 20
)
plt.axhline(18, color = 'red', ls = '--')
plt.savefig('C:\\Users\\nahue\\Desktop\\Agro 2\\Tp_5\\erosion.png')
| Tp 5/Ej_4_y_5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Add logs to invalid jobs
# Add id to executed job
# Add logs to executed job
# -
from mason.util.notebook_environment import NotebookEnvironment
env = NotebookEnvironment()
# !aws s3 ls spg-mason-demo/merged_csv/
job = env.run("table", "query", parameters="database_name:test-database,query_string:SELECT * FROM test-table LIMIT 5", config_id="1")
job_id = "9c59c226-0396-4e25-ba92-48149fd135bc"
job = env.run("job", "get", parameters=f"job_id:{job_id}", config_id="1")
job.to_response(Response())
table.schema.to_dict()
result = env.run(
"table",
"infer",
parameters="storage_path:spg-mason-demo/merged_csv/,database_name:crawler_poc",
config_id="3",
log_level="trace",
return_response=True
)
response.response.responses
| Notebooks/Mason.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Python library and module imports
import os
import sys
import numpy as np
import matplotlib.pyplot as plt
import math
import h5py
import torch
# Add the path to the parent directory to augment search for module
par_dir = os.path.abspath(os.path.join(os.getcwd(), os.pardir))
if par_dir not in sys.path:
sys.path.append(par_dir)
# Import the custom plotting module
from plot_utils import plot_utils
# WatChMaL imports
from io_utils.data_handling_2 import WCH5Dataset
# PyTorch imports
from torch.utils.data import DataLoader
from torch.utils.data.sampler import SubsetRandomSampler
from collections import Counter
# -
# ### Use the watchmal WCH5Dataset handler to load the dataset
train_batch_size = 1024
dset=WCH5Dataset("/fast_scratch/NuPRISM_4pi_full_tank_grid/out.h5", 0.1, 0.1, reduced_dataset_size=1000000)
train_loader = DataLoader(dset, batch_size=train_batch_size, shuffle=False,
pin_memory=True, sampler=SubsetRandomSampler(dset.train_indices))
# ### Iterate over the dataset and collect the positions per example
# +
labels = []
positions = []
for data in iter(train_loader):
labels.append(data[1].numpy())
positions.append(data[5].numpy())
# -
print(len(labels), labels[0].shape,
len(positions), positions[0].shape)
labels = np.concatenate(labels, axis=0)
positions = np.concatenate(positions, axis=0)
print(labels.shape, positions.shape)
# ### Check the distribution of labels
labels = labels.reshape(-1)
print(Counter(labels))
# ### Plot the per label position $r$ distribution
label_dict = {0:["gamma","red"], 1:["e","blue"], 2:["mu","green"]}
positions = positions.reshape(-1, positions.shape[2])
rs = np.array([math.sqrt(position[0]**2 + position[1]**2 + position[2]**2) for position in positions])
print(rs.shape)
# +
rs_dict = {}
for label in label_dict.keys():
rs_dict[label] = rs[labels == label]
fig, axes = plt.subplots(3, 1, figsize=(16,9), sharex=True)
for label in rs_dict.keys():
axes[label].hist(rs_dict[label], bins=200, density=False, label=label_dict[label][0], alpha=0.8,
color=label_dict[label][1])
axes[label].legend()
axes[label].set_ylabel("Frequency")
plt.xlabel(r"Position, $r = \sqrt{x^2 + y^2 + z^2}$")
# -
# ### Plot the per label position component distributions
print(len(positions[labels == 0]))
# +
pos_x_dict = {}
pos_y_dict = {}
pos_z_dict = {}
for label in label_dict.keys():
pos_x_dict[label] = positions[labels == label][:,0]
pos_y_dict[label] = positions[labels == label][:,1]
pos_z_dict[label] = positions[labels == label][:,2]
for pos_dict, axis in zip([pos_x_dict, pos_y_dict, pos_z_dict], ["x", "y", "z"]):
fig, axes = plt.subplots(3, 1, figsize=(16,9), sharex=True)
for label in label_dict.keys():
axes[label].hist(pos_dict[label], bins=200, density=False, label=label_dict[label][0], alpha=0.8,
color=label_dict[label][1])
axes[label].legend()
axes[label].set_ylabel("Frequency")
plt.xlabel(r" ${0}$ position".format(axis))
plt.show()
# -
| notebooks/2810 - New data exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Copyright (c) Facebook, Inc. and its affiliates.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# -
# # KubeFlow Pipelines : Pytorch Cifar10 Image classification
#
# This notebook shows PyTorch CIFAR10 end-to-end classification example using Kubeflow Pipelines.
#
# An example notebook that demonstrates how to:
#
# * Get different tasks needed for the pipeline
# * Create a Kubeflow pipeline
# * Include Pytorch KFP components to preprocess, train, visualize and deploy the model in the pipeline
# * Submit a job for execution
# * Query(prediction and explain) the final deployed model
#
# ! pip uninstall -y kfp
# ! pip install --no-cache-dir kfp
# ## import the necessary packages
# +
import kfp
import json
import os
from kfp.onprem import use_k8s_secret
from kfp import components
from kfp.components import load_component_from_file, load_component_from_url
from kfp import dsl
from kfp import compiler
import numpy as np
import logging
kfp.__version__
# -
# ## Enter your gateway and the auth token
# [Use this extension on chrome to get token]( https://chrome.google.com/webstore/detail/editthiscookie/fngmhnnpilhplaeedifhccceomclgfbg?hl=en)
#
# 
# ## Update values for the ingress gateway and auth session
INGRESS_GATEWAY='http://istio-ingressgateway.istio-system.svc.cluster.local'
AUTH="<enter your auth token>"
NAMESPACE="kubeflow-user-example-com"
COOKIE="authservice_session="+AUTH
EXPERIMENT="Default"
# ## Set the Log bucket and Tensorboard Image
MINIO_ENDPOINT="http://minio-service.kubeflow:9000"
LOG_BUCKET="mlpipeline"
TENSORBOARD_IMAGE="public.ecr.aws/pytorch-samples/tboard:latest"
# ## Set the client and create the experiment
client = kfp.Client(host=INGRESS_GATEWAY+"/pipeline", cookies=COOKIE)
client.create_experiment(EXPERIMENT)
experiments = client.list_experiments(namespace=NAMESPACE)
my_experiment = experiments.experiments[0]
my_experiment
# ## Set the Inference parameters
DEPLOY_NAME="torchserve"
MODEL_NAME="cifar10"
ISVC_NAME=DEPLOY_NAME+"."+NAMESPACE+"."+"example.com"
INPUT_REQUEST="https://raw.githubusercontent.com/kubeflow/pipelines/master/samples/contrib/pytorch-samples/cifar10/input.json"
# ## Load the the components yaml files for setting up the components
# ! python utils/generate_templates.py cifar10/template_mapping.json
# +
prepare_tensorboard_op = load_component_from_file("yaml/tensorboard_component.yaml")
prep_op = components.load_component_from_file(
"yaml/preprocess_component.yaml"
)
train_op = components.load_component_from_file(
"yaml/train_component.yaml"
)
deploy_op = load_component_from_file("yaml/deploy_component.yaml")
pred_op = load_component_from_file("yaml/prediction_component.yaml")
minio_op = components.load_component_from_file(
"yaml/minio_component.yaml"
)
# -
# ## Define the pipeline
# +
@dsl.pipeline(
name="Training Cifar10 pipeline", description="Cifar 10 dataset pipeline"
)
def pytorch_cifar10( # pylint: disable=too-many-arguments
minio_endpoint=MINIO_ENDPOINT,
log_bucket=LOG_BUCKET,
log_dir=f"tensorboard/logs/{dsl.RUN_ID_PLACEHOLDER}",
mar_path=f"mar/{dsl.RUN_ID_PLACEHOLDER}/model-store",
config_prop_path=f"mar/{dsl.RUN_ID_PLACEHOLDER}/config",
model_uri=f"s3://mlpipeline/mar/{dsl.RUN_ID_PLACEHOLDER}",
tf_image=TENSORBOARD_IMAGE,
deploy=DEPLOY_NAME,
isvc_name=ISVC_NAME,
model=MODEL_NAME,
namespace=NAMESPACE,
confusion_matrix_log_dir=f"confusion_matrix/{dsl.RUN_ID_PLACEHOLDER}/",
checkpoint_dir="checkpoint_dir/cifar10",
input_req=INPUT_REQUEST,
cookie=COOKIE,
ingress_gateway=INGRESS_GATEWAY,
):
def sleep_op(seconds):
"""Sleep for a while."""
return dsl.ContainerOp(
name="Sleep " + str(seconds) + " seconds",
image="python:alpine3.6",
command=["sh", "-c"],
arguments=[
'python -c "import time; time.sleep($0)"',
str(seconds)
],
)
"""This method defines the pipeline tasks and operations"""
pod_template_spec = json.dumps({
"spec": {
"containers": [{
"env": [
{
"name": "AWS_ACCESS_KEY_ID",
"valueFrom": {
"secretKeyRef": {
"name": "mlpipeline-minio-artifact",
"key": "accesskey",
}
},
},
{
"name": "AWS_SECRET_ACCESS_KEY",
"valueFrom": {
"secretKeyRef": {
"name": "mlpipeline-minio-artifact",
"key": "secretkey",
}
},
},
{
"name": "AWS_REGION",
"value": "minio"
},
{
"name": "S3_ENDPOINT",
"value": f"{minio_endpoint}",
},
{
"name": "S3_USE_HTTPS",
"value": "0"
},
{
"name": "S3_VERIFY_SSL",
"value": "0"
},
]
}]
}
})
prepare_tb_task = prepare_tensorboard_op(
log_dir_uri=f"s3://{log_bucket}/{log_dir}",
image=tf_image,
pod_template_spec=pod_template_spec,
).set_display_name("Visualization")
prep_task = (
prep_op().after(prepare_tb_task
).set_display_name("Preprocess & Transform")
)
confusion_matrix_url = f"minio://{log_bucket}/{confusion_matrix_log_dir}"
script_args = f"model_name=resnet.pth," \
f"confusion_matrix_url={confusion_matrix_url}"
# For GPU, set number of gpus and accelerator type
ptl_args = f"max_epochs=1, gpus=0, accelerator=None, profiler=pytorch"
train_task = (
train_op(
input_data=prep_task.outputs["output_data"],
script_args=script_args,
ptl_arguments=ptl_args
).after(prep_task).set_display_name("Training")
)
# For GPU uncomment below line and set GPU limit and node selector
# ).set_gpu_limit(1).add_node_selector_constraint
# ('cloud.google.com/gke-accelerator','nvidia-tesla-p4')
(
minio_op(
bucket_name="mlpipeline",
folder_name=log_dir,
input_path=train_task.outputs["tensorboard_root"],
filename="",
).after(train_task).set_display_name("Tensorboard Events Pusher")
)
(
minio_op(
bucket_name="mlpipeline",
folder_name=checkpoint_dir,
input_path=train_task.outputs["checkpoint_dir"],
filename="",
).after(train_task).set_display_name("checkpoint_dir Pusher")
)
minio_mar_upload = (
minio_op(
bucket_name="mlpipeline",
folder_name=mar_path,
input_path=train_task.outputs["checkpoint_dir"],
filename="cifar10_test.mar",
).after(train_task).set_display_name("Mar Pusher")
)
(
minio_op(
bucket_name="mlpipeline",
folder_name=config_prop_path,
input_path=train_task.outputs["checkpoint_dir"],
filename="config.properties",
).after(train_task).set_display_name("Conifg Pusher")
)
model_uri = str(model_uri)
# pylint: disable=unused-variable
isvc_yaml = """
apiVersion: "serving.kubeflow.org/v1beta1"
kind: "InferenceService"
metadata:
name: {}
namespace: {}
spec:
predictor:
serviceAccountName: sa
pytorch:
storageUri: {}
resources:
requests:
cpu: 4
memory: 8Gi
limits:
cpu: 4
memory: 8Gi
""".format(deploy, namespace, model_uri)
# For GPU inference use below yaml with gpu count and accelerator
gpu_count = "1"
accelerator = "nvidia-tesla-p4"
isvc_gpu_yaml = """# pylint: disable=unused-variable
apiVersion: "serving.kubeflow.org/v1beta1"
kind: "InferenceService"
metadata:
name: {}
namespace: {}
spec:
predictor:
serviceAccountName: sa
pytorch:
storageUri: {}
resources:
requests:
cpu: 4
memory: 8Gi
limits:
cpu: 4
memory: 8Gi
nvidia.com/gpu: {}
nodeSelector:
cloud.google.com/gke-accelerator: {}
""".format(deploy, namespace, model_uri, gpu_count, accelerator)
# Update inferenceservice_yaml for GPU inference
deploy_task = (
deploy_op(action="apply", inferenceservice_yaml=isvc_yaml
).after(minio_mar_upload).set_display_name("Deployer")
)
# Wait here for model to be loaded in torchserve for inference
sleep_task = sleep_op(5).after(deploy_task).set_display_name("Sleep")
# Make Inference request
pred_task = (
pred_op(
host_name=isvc_name,
input_request=input_req,
cookie=cookie,
url=ingress_gateway,
model=model,
inference_type="predict",
).after(sleep_task).set_display_name("Prediction")
)
(
pred_op(
host_name=isvc_name,
input_request=input_req,
cookie=cookie,
url=ingress_gateway,
model=model,
inference_type="explain",
).after(pred_task).set_display_name("Explanation")
)
dsl.get_pipeline_conf().add_op_transformer(
use_k8s_secret(
secret_name="mlpipeline-minio-artifact",
k8s_secret_key_to_env={
"secretkey": "MINIO_SECRET_KEY",
"accesskey": "MINIO_ACCESS_KEY",
},
)
)
# -
# ## Compile the pipeline
compiler.Compiler().compile(pytorch_cifar10, 'pytorch.tar.gz', type_check=True)
# ## Execute the pipeline
run = client.run_pipeline(my_experiment.id, 'pytorch-cifar10', 'pytorch.tar.gz')
# ## Wait for inference service below to go to READY True state
# !kubectl get isvc $DEPLOY
# ## Get the Inference service name
INFERENCE_SERVICE_LIST = ! kubectl get isvc {DEPLOY_NAME} -n {NAMESPACE} -o json | python3 -c "import sys, json; print(json.load(sys.stdin)['status']['url'])"| tr -d '"' | cut -d "/" -f 3
INFERENCE_SERVICE_NAME = INFERENCE_SERVICE_LIST[0]
INFERENCE_SERVICE_NAME
# ## Use the deployed model for prediction request and save the output into a json
# !curl -v -H "Host: $INFERENCE_SERVICE_NAME" -H "Cookie: $COOKIE" "$INGRESS_GATEWAY/v1/models/$MODEL_NAME:predict" -d @./cifar10/input.json > cifar10_prediction_output.json
# ! cat cifar10_prediction_output.json
# ## Use the deployed model for explain request and save the output into a json
# !curl -v -H "Host: $INFERENCE_SERVICE_NAME" -H "Cookie: $COOKIE" "$INGRESS_GATEWAY/v1/models/$MODEL_NAME:explain" -d @./cifar10/input.json > cifar10_explanation_output.json
# ## Clean up
# #### Delete Viewers, Inference Services and Completed pods
# ! kubectl delete --all isvc -n $NAMESPACE
# ! kubectl delete pod --field-selector=status.phase==Succeeded -n $NAMESPACE
| samples/contrib/pytorch-samples/Pipeline-Cifar10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="59BRf43IJiQ1"
from gym import spaces
import numpy as np
import random
from itertools import groupby
from itertools import product
# -
curr_state = [np.nan for _ in range(9)]
curr_state[0:4] = [1, 2, 3, 4]
#curr_state = [1,3,nan,nan,nan,nan,nan,nan,nan]
curr_state
np.nan
# +
# Row wise evaluation:
sum_row1 = curr_state[0] + curr_state[1] + curr_state[2]
sum_row2 = curr_state[3] + curr_state[4] + curr_state[5]
sum_row3 = curr_state[6] + curr_state[7] + curr_state[8]
print(sum_row1, sum_row2, sum_row3)
if(sum_row1 == 6
or sum_row2 == 15
or sum_row3 == 15):
print('winner')
# -
state = [np.nan for _ in range(9)]
[i for i in range(1, len(state) + 1)]
# +
max_epsilon = 1.0
min_epsilon = 0.001
time = np.arange(0,5000000)
epsilon = []
for i in range(0,5000000):
epsilon.append(min_epsilon + (max_epsilon - min_epsilon) * np.exp(-0.000001*i))
epsilon
# -
np.random.random()
# +
# !python TCGame_Env1.py
from TCGame_Env1 import TicTacToe
env3 = TicTacToe()
curr_state = env2.state #curr_state is state_start
valid_Actions = [i for i in env3.action_space(curr_state)[0]] ###### -------please call your environment as env
#curr_state = [np.nan for _ in range(9)]
#reward is not calculated for the immediately
#curr_action = np.random.choice(np.arange(0,m+1)) #randomly choosing and action
curr_action = random.choice(valid_Actions)
state = [np.nan for _ in range(9)]
curr_state = [i for i in range(1, len(state) + 1)]
print(valid_Actions)
print(env3.testFunctions())
#next_state, reward, terminal = env2.step(curr_state, curr_action)
# -
| ai-playing-tic-tac-toe/TicTacToe_Testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <font color='blue'>Data Science Academy - Python Fundamentos - Capítulo 10</font>
#
# ## Download: http://github.com/dsacademybr
# ## Mini-Projeto 3 - Guia de Modelagem Preditiva com Linguagem Python e TensorFlow
# 
# Neste Mini-Projeto vamos apresentar um guia básico de modelagem preditiva usando Linguagem Python e TensorFlow, o principal framework para construção de modelos de Machine Learning e Deep Learning e para construir aplicações comerciais de Inteligência Artificial.
#
# Este é um guia básico pois o TensorFlow é um framework extenso. O TensorFlow é abordado em detalhes nos cursos da <a href="https://www.datascienceacademy.com.br/bundle/formacao-inteligencia-artificial">Formação Inteligência Artificial</a> (especialmente no curso Deep Learning Frameworks) e na <a href="https://www.datascienceacademy.com.br/bundle/formacao-inteligencia-artificial-aplicada-a-medicina">Formação IA Aplicada à Medicina</a>.
#
# Na <a href="https://www.datascienceacademy.com.br/bundle/formacao-cientista-de-dados">Formação Cientista de Dados</a>, no curso de Machine Learning também há um módulo sobre TensorFlow. Alguns projetos com TensorFlow podem ser encontrados no curso de IA Aplicada a Finanças da <a href="https://www.datascienceacademy.com.br/bundle/formacao-engenheiro-blockchain">Formação Engenheiro Blockchain</a>.
# ## Instalando e Carregando Pacotes
# Versão da Linguagem Python
from platform import python_version
print('Versão da Linguagem Python Usada Neste Jupyter Notebook:', python_version())
# Instala o TensorFlow
# !pip install -q tensorflow==2.5
# Instala o Pydot
# !pip install -q pydot
# + colab={"base_uri": "https://localhost:8080/", "height": 70} colab_type="code" id="Yh0AtXXOXVuD" outputId="ec529b08-63f5-42a6-c058-56b3d81421b7"
# Imports
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="zP5cGu1RWLxw" outputId="d7957dbd-6a86-4c68-9f92-b97cf7e2826c"
# Imports
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
print(tf.__version__)
# -
# ## Carregando os Dados
# + [markdown] colab_type="text" id="CTetrCMSca0w"
# ### Boston House Prices Dataset
#
# https://www.cs.toronto.edu/~delve/data/boston/bostonDetail.html
#
# #### Características:
#
# * Número de Observções: 506
# * Os primeiros 13 recursos são recursos preditivos numéricos / categóricos.
# * O último (atributo 14): o valor mediano é a variável de destino.
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="3bAedq0NWThz" outputId="cbfdce8f-2dbc-494c-b2ab-32fa66159aaf"
# Download dos dados
import ssl
ssl._create_default_https_context = ssl._create_unverified_context
dataset_path = keras.utils.get_file("housing.data", "https://archive.ics.uci.edu/ml/machine-learning-databases/housing/housing.data")
# + colab={"base_uri": "https://localhost:8080/", "height": 343} colab_type="code" id="U3Pv_Om-WwHN" outputId="24075064-33e9-471c-8199-8f3691643587"
# Nomes das colunas
nomes_colunas = ['CRIM',
'ZN',
'INDUS',
'CHAS',
'NOX',
'RM',
'AGE',
'DIS',
'RAD',
'TAX',
'PTRATION',
'B',
'LSTAT',
'MEDV']
# -
# Carrega os dados
dataset = pd.read_csv(dataset_path,
names = nomes_colunas,
na_values = "?",
comment = '\t',
sep = " ",
skipinitialspace = True)
# Shape
dataset.shape
# Visualiza os dados
dataset.head()
# + [markdown] colab_type="text" id="nf8QJRMeh910"
# Vamos dividir os dados em treino e teste com proporção 80/20.
# + colab={} colab_type="code" id="Vd0uyKsFgyQ0"
# Split dos dados
dados_treino = dataset.sample(frac = 0.8, random_state = 0)
dados_teste = dataset.drop(dados_treino.index)
# + [markdown] colab_type="text" id="4vC8cz0chjL1"
# ## Modelagem Preditiva - Regressão Linear Simples
#
# Na regressão linear simples desejamos modelar a relação entre a variável dependente (y) e uma variável independente (x).
#
# * Variável independente: 'RM'
# * Variável dependente: 'MEDV'
#
# Queremos prever o valor da mediana das casas ocupadas por proprietários com base no número de quartos.
# + [markdown] colab_type="text" id="KY0vmL_PhIPn"
# Vamos criar um plot mostrando a relação atual entre as variáveis.
#
# Basicamente temos $MEDV=f(RM)$ e queremos estimar a função $f()$ usando regressão linear.
# + colab={"base_uri": "https://localhost:8080/", "height": 279} colab_type="code" id="E_R3ADmzgmo0" outputId="5bffce03-ca2a-4d56-9710-4c280cdb6482"
# Representação visual dos dados de treino
fig, ax = plt.subplots()
x = dados_treino['RM']
y = dados_treino['MEDV']
ax.scatter(x, y, edgecolors = (0, 0, 0))
ax.set_xlabel('RM')
ax.set_ylabel('MEDV')
plt.show()
# + [markdown] colab_type="text" id="M-EpBay_M3_X"
# Vamos separar x e y.
# + colab={} colab_type="code" id="dgOBhSaniDHx"
# Divisão
x_treino = dados_treino['RM']
y_treino = dados_treino['MEDV']
x_teste = dados_teste['RM']
y_teste = dados_teste['MEDV']
# + [markdown] colab_type="text" id="VXA8J28QWYMH"
# ## Criação do Modelo
#
# Queremos encontrar os parâmetros (**W**) que permitem prever a saída y a partir da entrada x:
#
# $y = w_1 x + w_0$
#
# A fórmula acima pode ser definida com a seguinte camada densa em um modelo de rede neural artificial:
#
# *layers.Dense(1, use_bias=True, input_shape=(1,))*
# -
# Função para construir o modelo
def modelo_linear():
# Cria o modelo
model = keras.Sequential([layers.Dense(1, use_bias = True, input_shape = (1,), name = 'layer')])
# Otimizador
optimizer = tf.keras.optimizers.Adam(learning_rate = 0.01,
beta_1 = 0.9,
beta_2 = 0.99,
epsilon = 1e-05,
amsgrad = False,
name = 'Adam')
# Compila o modelo
model.compile(loss = 'mse',
optimizer = optimizer,
metrics = ['mae','mse'])
return model
# MAE = Mean Absolute Error
#
# MSE = Mean Squared Error
# Cria o modelo
modelo = modelo_linear()
# + colab={"base_uri": "https://localhost:8080/", "height": 185} colab_type="code" id="okLiiirkWaot" outputId="a60d4638-bedc-47cf-8236-ab23a81dc2cc"
# Plot do modelo
tf.keras.utils.plot_model(modelo,
to_file = 'imagens/modelo.png',
show_shapes = True,
show_layer_names = True,
rankdir = 'TB',
expand_nested = False,
dpi = 100)
# + [markdown] colab_type="text" id="6ny4wr5l3M_k"
# ### Treinamento do Modelo
# -
# Hiperparâmetros
n_epochs = 4000
batch_size = 256
n_idle_epochs = 100
n_epochs_log = 200
n_samples_save = n_epochs_log * x_treino.shape[0]
print('Checkpoint salvo a cada {} amostras'.format(n_samples_save))
# Callback
earlyStopping = tf.keras.callbacks.EarlyStopping(monitor = 'val_loss',
patience = n_idle_epochs,
min_delta = 0.001)
# Lista para as previsões
predictions_list = []
# Caminho ppara salvar o checkpoint
checkpoint_path = "dados/"
# Create a callback that saves the model's weights every n_samples_save
checkpointCallback = tf.keras.callbacks.ModelCheckpoint(filepath = "dados/",
verbose = 1,
save_weights_only = True,
save_freq = n_samples_save)
# Salva a primeira versão do modelo
modelo.save_weights(checkpoint_path.format(epoch = 0))
# + colab={"base_uri": "https://localhost:8080/", "height": 235} colab_type="code" id="YKMSaOXnXmOT" outputId="df8ca3f0-5632-4e40-9707-8e00b6726795"
# Treinamento
history = modelo.fit(x_treino,
y_treino,
batch_size = batch_size,
epochs = n_epochs,
validation_split = 0.1,
verbose = 1,
callbacks = [earlyStopping, checkpointCallback])
# -
# Métricas do histórico de treinamento
print('keys:', history.history.keys())
# MSE = Mean Squared Error
# Retornando os valores desejados para o plot
mse = np.asarray(history.history['mse'])
val_mse = np.asarray(history.history['val_mse'])
# Prepara os valores para o dataframe
num_values = (len(mse))
values = np.zeros((num_values, 2), dtype = float)
values[:,0] = mse
values[:,1] = val_mse
# Cria o dataframe
steps = pd.RangeIndex(start = 0, stop = num_values)
df = pd.DataFrame(values, steps, columns = ["MSE em Treino", "MSE em Validação"])
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 302} colab_type="code" id="uVvegIVGXvbv" outputId="be0dac35-2301-434d-97ab-07900016e970"
# Plot
sns.set(style = "whitegrid")
sns.lineplot(data = df, palette = "tab10", linewidth = 2.5)
# -
# Previsões com o modelo treinado
previsoes = modelo.predict(x_teste).flatten()
# + colab={} colab_type="code" id="KniAsElx9cqB"
# Imprime as previsões
previsoes
# -
# # Fim
# ### Obrigado
#
# ### Visite o Blog da Data Science Academy - <a href="http://blog.dsacademy.com.br">Blog DSA</a>
| Data Science Academy/PythonFundamentos/Cap10/Notebooks/DSA-Python-Cap10-Mini-Projeto3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:GEM_py36]
# language: python
# name: conda-env-GEM_py36-py
# ---
# # Load dependencies
# +
import cobra
import libsbml
import lxml
models_directory = '/media/sf_Shared/Systems_biology/Metabolic_models/'
OB3b_directory = '/media/sf_Shared/GEM_OB3b/'
memote_directory = '/home/ensakz/Desktop/memote_m_trichosporium/'
fastas_directory = '/media/sf_Shared/Systems_biology/Fastas_and_annotations/'
inparanoid_directory = '/media/sf_Shared/Systems_biology/InParanoid_runs/'
draft_gems_dir_8 = 'draft_gems/8.OB3b_draft_gem_during_pathway_manual_curation/'
# -
iMsOB3b = cobra.io.load_json_model(memote_directory + draft_gems_dir_8 + "bigg_OB3b_16_01_2020.json")
model_Bath = cobra.io.load_json_model(models_directory + "iMcBath.json")
model_universal = cobra.io.load_json_model(models_directory + 'universal_model.json')
# # Curate electron transport chain and nitrogen metabolism
# Look at cofactors present in electron transport chain
# ## Draw all reactions belonging to ETC on Escher map
for reaction in iMsOB3b.metabolites.ficytc_c.reactions:
print(reaction.name, reaction)
for reaction in iMsOB3b.metabolites.qh2_c.reactions:
if "iomass" not in reaction.name:
print(reaction.name, reaction)
for reaction in iMsOB3b.metabolites.nadh_c.reactions:
if "iomass" not in reaction.name:
print(reaction.name, reaction)
iMsOB3b.metabolites.q8_c.reactions
iMsOB3b.metabolites.q8h2_c.reactions
import re
for reaction in iMsOB3b.reactions:
if re.search("transhydrogenase", reaction.name, re.IGNORECASE):
print(reaction.id, reaction.name)
import re
for reaction in iMsOB3b.reactions:
if re.search("FMN reducta", reaction.name, re.IGNORECASE):
print(reaction.id, reaction.name)
import re
for reaction in iMsOB3b.reactions:
if re.search("Inorganic", reaction.name, re.IGNORECASE):
print(reaction.id, reaction.name)
import re
for reaction in iMsOB3b.reactions:
if re.search("oxidoreductase", reaction.name, re.IGNORECASE):
print(reaction.id, reaction.name)
import re
for reaction in iMsOB3b.reactions:
if re.search("ubiquinone", reaction.name, re.IGNORECASE):
print(reaction.id, reaction.name)
import re
for reaction in iMsOB3b.reactions:
if re.search("translocatin", reaction.name, re.IGNORECASE):
print(reaction.id, reaction.name)
import re
for reaction in iMsOB3b.reactions:
if re.search("nitrate r", reaction.name, re.IGNORECASE):
print(reaction.id, reaction.name)
import re
for reaction in iMsOB3b.reactions:
if re.search("rubre", reaction.name, re.IGNORECASE):
print(reaction.id, reaction.name)
iMsOB3b.reactions.EX_DM_rbrdxOX
iMsOB3b.metabolites.rbrdxOX_c.reactions
import re
for reaction in iMsOB3b.reactions:
if re.search("pyridi", reaction.name, re.IGNORECASE):
print(reaction.id, reaction.name)
iMsOB3b.reactions.SUCCD1
# ## Draw all reactions belonging to Nitrogen metabolism on Escher map
# ### Molecular nitrogen:
iMsOB3b.metabolites.n2_c.reactions
iMsOB3b.metabolites.n2_e.reactions
iMsOB3b.reactions.EX_n2_e
# ### Ammonia
iMsOB3b.metabolites.nh4_e
iMsOB3b.metabolites.nh4_p
iMsOB3b.metabolites.nh4_c
iMsOB3b.reactions.GLUD
# ### Nitrate
iMsOB3b.metabolites.no3_c
iMsOB3b.reactions.NODOy
iMsOB3b.metabolites.no3_e
iMsOB3b.metabolites.no3_p
iMsOB3b.metabolites.no3_c
# ### Nitrite
iMsOB3b.metabolites.no2_e
iMsOB3b.metabolites.no2_p
iMsOB3b.metabolites.no2_c
iMsOB3b.reactions.NTRIR4
iMsOB3b.reactions.NTPPD11
# ### Search for ammonia metabolism by keywords:
# Within OB3b GEM:
# Nitrite:
import re
for reaction in iMsOB3b.reactions:
if re.search("ferredoxin", reaction.name, re.IGNORECASE):
print(reaction.id, reaction.name)
# Nitrate:
import re
for reaction in iMsOB3b.reactions:
if re.search("nitrate red", reaction.name, re.IGNORECASE):
print(reaction.id, reaction.name)
import re
for reaction in iMsOB3b.reactions:
if re.search("ABC t", reaction.name, re.IGNORECASE):
print(reaction.id, reaction.name)
# Ammonia
import re
for reaction in iMsOB3b.reactions:
if re.search("ammon", reaction.name, re.IGNORECASE):
print(reaction.id, reaction.name)
iMsOB3b.reactions.GMPS
iMsOB3b.reactions.SAMMOi
# Molecular nitrogen:
import re
for reaction in iMsOB3b.reactions:
if re.search("nitr", reaction.name, re.IGNORECASE):
print(reaction.id, reaction.name)
# ## Check each reaction with KEGG database
# Check compliance with KEGG annotation first, then see whether some KEGG pathway reactions are missing from OB3b GEM.
# <br> oxidative phosphorylation --> https://www.genome.jp/kegg-bin/show_pathway?mtw00190
# <br> nitrogen metabolism --> https://www.genome.jp/kegg-bin/show_pathway?mtw00910
# Import orangecontrib.bioinformatics.kegg:
# +
import orangecontrib.bioinformatics.kegg
organism = orangecontrib.bioinformatics.kegg.Organism("Methylosinus trichosporium")
genes = organism.genes
# -
# ### ETC
# <br> For KEGG-based curation, please refer to: https://www.genome.jp/kegg-bin/show_pathway?mtw00190
iMsOB3b.reactions.THD2pp.gene_reaction_rule = '(CQW49_13075 or CQW49_13080) and CQW49_13070'
# NADH-quinone oxidoreductase
for gene in iMsOB3b.reactions.NADH11.genes:
gene_entry = genes["mtw:"+gene.id]
print(gene, gene_entry.orthology)
gene_entry = genes["mtw:"+'CQW49_18615']
print(locus_tag, gene_entry)
# Succinate dehydrogenase
for gene in iMsOB3b.reactions.SUCCD1.genes:
gene_entry = genes["mtw:"+gene.id]
print(gene, gene_entry.orthology)
# ubiquinol_cytochrome_c_reductase:
# <br> CQW49_00050 and CQW49_00055 are added based on ortholog analysis
for gene in iMsOB3b.reactions.UQCYOR_2p.genes:
gene_entry = genes["mtw:"+gene.id]
print(gene, gene_entry.orthology)
# Complex IV reactions
# <br>cytochrome d ubiquinol oxidase subunit II
iMsOB3b.reactions.CYOD.name = 'cytochrome bd ubiquinol oxidase'
for gene in iMsOB3b.reactions.CYOD.genes:
gene_entry = genes["mtw:"+gene.id]
print(gene, gene_entry.orthology)
# CYO_2p
iMsOB3b.reactions.CYO_2p
for gene in iMsOB3b.reactions.CYO_2p.genes:
gene_entry = genes["mtw:"+gene.id]
print(gene, gene_entry.orthology)
# CYOO_4p
iMsOB3b.reactions.CYOO_4p
for gene in iMsOB3b.reactions.CYOO_4p.genes:
gene_entry = genes["mtw:"+gene.id]
print(gene, gene_entry.orthology)
# ATPS4rpp
iMsOB3b.reactions.ATPS4rpp
for gene in iMsOB3b.reactions.ATPS4rpp.genes:
gene_entry = genes["mtw:"+gene.id]
print(gene, gene_entry.orthology)
# FMNRx2
# <br> gapfilling reaction
iMsOB3b.reactions.FMNRx2#.flux
with iMsOB3b:
iMsOB3b.reactions.FMNRx2.knock_out()
iMsOB3b.optimize()
print(iMsOB3b.reactions.Biomass_Mextorquens_AM1_core.flux)
for gene in iMsOB3b.reactions.FMNRx2.genes:
gene_entry = genes["mtw:"+gene.id]
print(gene, gene_entry.orthology)
with iMsOB3b:
iMsOB3b.reactions.FMNRx2.reaction = 'fmnh2_c + nad_c <=> fmn_c + h_c + nadh_c'
iMsOB3b.optimize()
print(iMsOB3b.reactions.Biomass_Mextorquens_AM1_core.flux)
iMsOB3b.metabolites.fmnh2_c.summary()
# ETFDH
iMsOB3b.reactions.ETFDH
for gene in iMsOB3b.reactions.ETFDH.genes:
gene_entry = genes["mtw:"+gene.id]
print(gene, gene_entry.orthology)
# PPA
iMsOB3b.reactions.PPA
for gene in iMsOB3b.reactions.PPA.genes:
gene_entry = genes["mtw:"+gene.id]
print(gene, gene_entry.orthology)
# FRDOr
# <br> Gapfilling reaction
iMsOB3b.reactions.FRDOr
iMsOB3b.metabolites.fdxrd_c.summary()
iMsOB3b.metabolites.nadp_c.summary()
with iMsOB3b:
iMsOB3b.reactions.FRDOr.knock_out()
iMsOB3b.optimize()
print(iMsOB3b.reactions.Biomass_Mextorquens_AM1_core.flux)
with iMsOB3b:
iMsOB3b.reactions.FRDOr.reaction = '2.0 fdxox_c + nadph_c <=> 2.0 fdxrd_c + h_c + nadp_c'
iMsOB3b.optimize()
print(iMsOB3b.reactions.Biomass_Mextorquens_AM1_core.flux)
for gene in iMsOB3b.reactions.FRDOr.genes:
gene_entry = genes["mtw:"+gene.id]
print(gene, gene_entry.orthology)
iMsOB3b.metabolites.indpyr_c
import re
for metabolite in iMsOB3b.metabolites:
if re.search("indol", metabolite.name, re.IGNORECASE):
print(metabolite.id, metabolite.name)
# PPAppt
iMsOB3b.reactions.PPAppt
for gene in iMsOB3b.reactions.PPAppt.genes:
gene_entry = genes["mtw:"+gene.id]
print(gene, gene_entry.orthology)
with iMsOB3b:
iMsOB3b.reactions.PPAppt.knock_out()
iMsOB3b.optimize()
print(iMsOB3b.reactions.Biomass_Mextorquens_AM1_core.flux)
iMsOB3b.reactions.PPAppt.remove_from_model()
# NO3Ras and NO3Rpp and NO3R1
iMsOB3b.reactions.NO3Ras.gene_reaction_rule = 'CQW49_19300 or CQW49_17890'
iMsOB3b.reactions.NO3R1.gene_reaction_rule = 'CQW49_19300 or CQW49_17890'
iMsOB3b.reactions.NO3Rpp
for gene in iMsOB3b.reactions.NO3R1.genes:
gene_entry = genes["mtw:"+gene.id]
print(gene, gene_entry.orthology)
iMsOB3b.reactions.NO3Rpp.remove_from_model()
# EAOXRED
# <br> Included based on KEGG information --> https://www.genome.jp/dbget-bin/www_bget?mtw:CQW49_14855
iMsOB3b.reactions.EAOXRED.gene_reaction_rule = 'CQW49_14855'
import re
for metabolite in iMsOB3b.metabolites:
if re.search("polyp", metabolite.name, re.IGNORECASE):
print(metabolite.id, metabolite.name)
iMsOB3b.metabolites.polypi_c
iMsOB3b.reactions.POLYPIP
iMsOB3b.add_reaction(model_universal.reactions.PPK2)
iMsOB3b.reactions.PPK2.gene_reaction_rule = 'CQW49_02850'
# SUCCD1
# <br> gene reaction rule fixing
for gene in iMsOB3b.reactions.SUCCD1.genes:
gene_entry = genes["mtw:"+gene.id]
print(gene, gene_entry.orthology)
iMsOB3b.reactions.SUCCD1.gene_reaction_rule = '(CQW49_10805 or CQW49_22885) and CQW49_10810 and (CQW49_10825 or CQW49_22875) and CQW49_10800'
# heme o synthase
import re
for reaction in iMsOB3b.reactions:
if re.search("heme", reaction.name, re.IGNORECASE):
print(reaction.id, reaction.name)
iMsOB3b.reactions.HEMEOS
# Save GEM for now:
cobra.io.save_json_model(iMsOB3b, memote_directory + draft_gems_dir_8 + "bigg_OB3b_18_01_2020.json")
# ### Nitrogen metabolism
# For nitrogen metabolism, please refer to Escher_map_17_01_2020_nitrogen_metabolism.json map
# <br> KEGG nitrogen metabolism pathway --> https://www.genome.jp/kegg-bin/show_pathway?mtw00910
# Molecular nitrogen
iMsOB3b.reactions.EX_n2_e
iMsOB3b.reactions.N2tex
iMsOB3b.reactions.N2trpp
iMsOB3b.reactions.NIT_mc
# Nitrate
iMsOB3b.reactions.EX_no3_e
iMsOB3b.reactions.NO3tex
# Nitrite
iMsOB3b.reactions.EX_no2_e
iMsOB3b.reactions.NO2tex
# 2 nitropropane dioxygenase:
# <br>KEGG and Genbank does not comply with each other
iMsOB3b.reactions.NTPPD11.remove_from_model()
# Ammonia
iMsOB3b.reactions.EX_nh4_e
iMsOB3b.reactions.NH4tex
iMsOB3b.reactions.NH4tpp
iMsOB3b.reactions.PAMMOipp
iMsOB3b.reactions.SAMMOi.gene_reaction_rule = 'CQW49_14985'
iMsOB3b.reactions.CYP460ipp.remove_from_model()
iMsOB3b.reactions.HAORipp.remove_from_model()
iMsOB3b.reactions.NO2tpp
iMsOB3b.reactions.NO3t7pp.gene_reaction_rule = 'CQW49_19295'
iMsOB3b.reactions.NITR_AMpp.remove_from_model()
iMsOB3b.reactions.NTRIR2x.gene_reaction_rule = 'CQW49_19305 or CQW49_19310'
iMsOB3b.reactions.NTRIR4.remove_from_model()
import re
for reaction in iMsOB3b.reactions:
if re.search("sulfite", reaction.name, re.IGNORECASE):
print(reaction.id, reaction.name)
iMsOB3b.reactions.SULR.gene_reaction_rule = 'CQW49_00335'
iMsOB3b.reactions.NODOy.remove_from_model()
iMsOB3b.reactions.NTRIR2y.remove_from_model()
iMsOB3b.reactions.NITR_NOpp.remove_from_model()
iMsOB3b.reactions.NORZ2pp
# Check ammonia assimilation:
iMsOB3b.reactions.GLNS
iMsOB3b.reactions.GLUSy
# Remove GLUD from GEM based on KEGG annotations:
# <br> https://www.genome.jp/dbget-bin/www_bget?mtw:CQW49_02775
iMsOB3b.reactions.GLUD.remove_from_model()
iMsOB3b.reactions.GLUDxi
iMsOB3b.reactions.GLUDy
# KEGG-based manual curation:
iMsOB3b.metabolites.n2o_p
iMsOB3b.metabolites.n2o_c
iMsOB3b.metabolites.no_c
iMsOB3b.metabolites.no_p
iMsOB3b.reactions.NORZ2pp.reaction = '2.0 focytc_c + 2.0 h_c + 2.0 no_c --> 2.0 ficytc_c + h2o_c + n2o_c'
iMsOB3b.reactions.NORZ2pp.id = "NORZ2"
iMsOB3b.reactions.PMMOipp.gene_reaction_rule
iMsOB3b.reactions.SMMOi.gene_reaction_rule
iMsOB3b.reactions.PAMMOipp.gene_reaction_rule = "CQW49_10250 and (CQW49_10255 or CQW49_01220) and CQW49_10260"
iMsOB3b.reactions.SAMMOi.gene_reaction_rule = "CQW49_12480 and CQW49_12475 and CQW49_12465 and CQW49_12470 and CQW49_12455 and CQW49_12485 and CQW49_12495"
# Adapt HAMR from http://bigg.ucsd.edu/universal/reactions/HAMR
iMsOB3b.add_reaction(model_universal.reactions.HAMR)
iMsOB3b.reactions.HAMR.gene_reaction_rule = 'CQW49_14985'
iMsOB3b.reactions.HAMR
# Include hydroxylamine utilizing reactions based on experimental evidence of NO2 production:
#
# https://www.ncbi.nlm.nih.gov/pubmed/3921227
iMsOB3b.add_reaction(model_Bath.reactions.CYP460ipp)
iMsOB3b.add_reaction(model_Bath.reactions.HAORipp)
# Save GEM:
cobra.io.save_json_model(iMsOB3b, memote_directory + draft_gems_dir_8 + "bigg_OB3b_18_01_2020.json")
# # Some stuff for progress report
nitrogen_reactions = ['NO2tpp', 'NO3t7pp', 'PAMMOipp', 'SAMMOi', 'HAMR', 'CYP460ipp', 'HAORipp', 'NO3Ras', 'NO3R1',
'NTRIR2x', 'NORZ2']
ammonia_assimilation = ['GLNS', 'GLUSy', 'GLUDxi', 'GLUDy']
for reaction_id in nitrogen_reactions:
print(reaction_id, iMsOB3b.reactions.get_by_id(reaction_id).name)
for reaction_id in ammonia_assimilation:
print(reaction_id, iMsOB3b.reactions.get_by_id(reaction_id).name)
ETC = ['THD2pp', 'NADH11', 'SUCCD1', 'UQCYOR_2p', 'CYOD', 'CYO_2p', 'CYOO_4p', 'ATPS4rpp', 'FMNRx2',
'ETFDH', 'PPA', 'FRDOr', 'EAOXRED']
for reaction_id in ETC:
print(reaction_id, iMsOB3b.reactions.get_by_id(reaction_id).name)
| supplementary_scripts/5. Curate electron transport chain and nitrogen metabolism.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="MNd7jnnSkuUD" colab_type="text"
# #Setup
# Specify your desired blender version and the path to your blend file within google drive.
#
# ###Info
# If you do need more information on parameters etc. look here: [Blender CLI Wiki](https://docs.blender.org/manual/en/latest/advanced/command_line/arguments.html)
# + id="8dFNjYGTgNjR" colab_type="code" colab={}
#@title Setup
#@markdown Please configure your setup
blender_version = 'blender2.90.1' #@param ["blender2.80", "blender2.81", "blender2.82", "blender2.83", "blender2.90.1"] {allow-input: false}
path_to_blend = 'path/to/blend.blend' #@param {type: "string"}
output_path = 'output/dir/in/drive/output####.png' #@param {type: "string"}
gpu_enabled = True #@param {type:"boolean"}
cpu_enabled = False #@param {type:"boolean"}
#@markdown ---
# + id="GPJ9fmoB6PWM" colab_type="code" colab={}
if blender_version == "blender2.80":
download_path="https://ftp.halifax.rwth-aachen.de/blender/release/Blender2.80/blender-2.80-linux-glibc217-x86_64.tar.bz2"
elif blender_version == "blender2.81":
download_path="https://ftp.halifax.rwth-aachen.de/blender/release/Blender2.81/blender-2.81-linux-glibc217-x86_64.tar.bz2"
elif blender_version == "blender2.82":
download_path="https://ftp.halifax.rwth-aachen.de/blender/release/Blender2.82/blender-2.82-linux64.tar.xz"
elif blender_version == "blender2.83":
download_path="https://ftp.halifax.rwth-aachen.de/blender/release/Blender2.83/blender-2.83.0-linux64.tar.xz"elif blender_version == "blender2.90.1":
download_path="https://ftp.halifax.rwth-aachen.de/blender/release/Blender2.90/blender-2.90.1-linux64.tar.xz"
# + id="OQ54OjLVjb26" colab_type="code" colab={}
# !mkdir $blender_version
if blender_version == "blender2.81" or "blender2.80":
# !wget -O '{blender_version}.tar.bz2' -nc $download_path
# !tar -xf '{blender_version}.tar.bz2' -C ./$blender_version --strip-components=1else:
# !wget -O '{blender_version}.tar.xz' -nc $download_path
# !tar xf '{blender_version}.tar.xz' -C ./$blender_version --strip-components=1
# + [markdown] id="s3uIDMZLbYOE" colab_type="text"
# This Block is required as some weird behaviors with libtcmalloc appeared in the colab VM
# + id="h6vohA2q2BDF" colab_type="code" colab={}
import os
os.environ["LD_PRELOAD"] = ""
# !apt update
# !apt remove libtcmalloc-minimal4
# !apt install libtcmalloc-minimal4
os.environ["LD_PRELOAD"] = "/usr/lib/x86_64-linux-gnu/libtcmalloc_minimal.so.4.3.0"
# !echo $LD_PRELOAD
# + [markdown] id="UUwwvaq5BxzN" colab_type="text"
# Unpack and move blender
# + id="RU3p_G4hDk97" colab_type="code" colab={}
# !apt install libboost-all-dev
# !apt install libgl1-mesa-dev
# !apt install libglu1-mesa libsm-dev
# + [markdown] id="Wh4i8msvAanq" colab_type="text"
# Required for Blender to use the GPU as expected
# + id="30Tv3lIeVaC6" colab_type="code" colab={}
data = "import re\n"+\
"import bpy\n"+\
"scene = bpy.context.scene\n"+\
"scene.cycles.device = 'GPU'\n"+\
"prefs = bpy.context.preferences\n"+\
"prefs.addons['cycles'].preferences.get_devices()\n"+\
"cprefs = prefs.addons['cycles'].preferences\n"+\
"print(cprefs)\n"+\
"# Attempt to set GPU device types if available\n"+\
"for compute_device_type in ('CUDA', 'OPENCL', 'NONE'):\n"+\
" try:\n"+\
" cprefs.compute_device_type = compute_device_type\n"+\
" print('Device found',compute_device_type)\n"+\
" break\n"+\
" except TypeError:\n"+\
" pass\n"+\
"#for scene in bpy.data.scenes:\n"+\
"# scene.render.tile_x = 64\n"+\
"# scene.render.tile_y = 64\n"+\
"# Enable all CPU and GPU devices\n"+\
"for device in cprefs.devices:\n"+\
" if not re.match('intel', device.name, re.I):\n"+\
" print('Activating',device)\n"+\
" device.use = "+str(gpu_enabled)+"\n"+\
" else:\n"+\
" device.use = "+str(cpu_enabled)+"\n"
with open('setgpu.py', 'w') as f:
f.write(data)
# + id="QRzNmiHN8Xr0" colab_type="code" colab={}
from google.colab import drive
drive.mount('/gdrive')
# + [markdown] id="RPiQkHVbCDu2" colab_type="text"
# Use this if you want to render all Frames
#
# Use -s to speficy the start frame.
# eg: -s 10
#
# Use -e to speficy the end frame.
# eg: -e 20
#
# **THE ORDER IS IMPORTANT. BOTH -s AND -e MUST BE SPEFICIED BEFORE -a**
# + id="Fpk2w3yM8XqQ" colab_type="code" colab={}
# !sudo ./$blender_version/blender -P './setgpu.py' -b '/gdrive/My Drive/{path_to_blend}' -E CYCLES -o '/gdrive/My Drive/{output_path}' -a
# + [markdown] id="qmZg_8Eylgxe" colab_type="text"
# Use to render a single frame.
# Specify the frame with -f *frame_number*
# + id="VFVx8omJCGBN" colab_type="code" colab={}
# !sudo ./$blender_version/blender -P 'setgpu.py' -b '/gdrive/My Drive/{path_to_blend}' -P 'setgpu.py' -o '/gdrive/My Drive/{output_path}' -f 1
| runblender.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Scala (2.13)
// language: scala
// name: scala213
// ---
// <p style="float: left;"><a href="upper-type-bounds.ipynb" target="_blank">Previous</a></p>
// <p style="float: right;"><a href="inner-classes.ipynb" target="_blank">Next</a></p>
// <p style="text-align:center;">Tour of Scala</p>
// <div style="clear: both;"></div>
//
// # Lower Type Bounds
//
// While [upper type bounds](upper-type-bounds.ipynb) limit a type to a subtype of another type, *lower type bounds* declare a type to be a supertype of another type. The term `B >: A` expresses that the type parameter `B` or the abstract type `B` refer to a supertype of type `A`. In most cases, `A` will be the type parameter of the class and `B` will be the type parameter of a method.
//
// Here is an example where this is useful:
// + attributes={"classes": ["tut"], "id": ""}
trait Node[+B] {
def prepend(elem: B): Node[B]
}
case class ListNode[+B](h: B, t: Node[B]) extends Node[B] {
def prepend(elem: B): ListNode[B] = ListNode(elem, this)
def head: B = h
def tail: Node[B] = t
}
case class Nil[+B]() extends Node[B] {
def prepend(elem: B): ListNode[B] = ListNode(elem, this)
}
// -
// This program implements a singly-linked list. `Nil` represents an empty element (i.e. an empty list). `class ListNode` is a node which contains an element of type `B` (`head`) and a reference to the rest of the list (`tail`). The `class Node` and its subtypes are covariant because we have `+B`.
//
// However, this program does _not_ compile because the parameter `elem` in `prepend` is of type `B`, which we declared *co*variant. This doesn't work because functions are *contra*variant in their parameter types and *co*variant in their result types.
//
// To fix this, we need to flip the variance of the type of the parameter `elem` in `prepend`. We do this by introducing a new type parameter `U` that has `B` as a lower type bound.
// + attributes={"classes": ["tut"], "id": ""}
trait Node[+B] {
def prepend[U >: B](elem: U): Node[U]
}
case class ListNode[+B](h: B, t: Node[B]) extends Node[B] {
def prepend[U >: B](elem: U): ListNode[U] = ListNode(elem, this)
def head: B = h
def tail: Node[B] = t
}
case class Nil[+B]() extends Node[B] {
def prepend[U >: B](elem: U): ListNode[U] = ListNode(elem, this)
}
// -
// Now we can do the following:
// + attributes={"classes": ["tut"], "id": ""}
trait Bird
case class AfricanSwallow() extends Bird
case class EuropeanSwallow() extends Bird
val africanSwallowList= ListNode[AfricanSwallow](AfricanSwallow(), Nil())
val birdList: Node[Bird] = africanSwallowList
birdList.prepend(new EuropeanSwallow)
// -
// The `Node[Bird]` can be assigned the `africanSwallowList` but then accept `EuropeanSwallow`s.
// <p style="float: left;"><a href="upper-type-bounds.ipynb" target="_blank">Previous</a></p>
// <p style="float: right;"><a href="inner-classes.ipynb" target="_blank">Next</a></p>
// <p style="text-align:center;">Tour of Scala</p>
// <div style="clear: both;"></div>
| notebooks/scala-tour/lower-type-bounds.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Imports
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
# tensorflow
import tensorflow as tf
# Estimators
from tensorflow.contrib import learn
# Model builder
from tensorflow.contrib.learn.python.learn.estimators import model_fn as model_fn_lib
print (tf.__version__) # tested with v1.1
# Input function
from tensorflow.python.estimator.inputs import numpy_io
# numpy
import numpy as np
# Enable TensorFlow logs
tf.logging.set_verbosity(tf.logging.INFO)
# keras
from tensorflow.contrib.keras.python.keras.preprocessing import sequence
from tensorflow.contrib.keras.python.keras.layers import Embedding, GRU, Dense, SimpleRNN
from tensorflow.contrib.keras.python.keras.layers import Reshape, Activation
# data
from tensorflow.contrib.keras.python.keras.datasets import imdb
# Run an experiment
from tensorflow.contrib.learn.python.learn import learn_runner
# -
# ## Helpers
# +
# map word to index
word_to_index = imdb.get_word_index()
# map index to word
index_to_word = {}
num_words = 0
for k in word_to_index:
index_to_word[word_to_index[k]] = k
num_words += 1
# turn a sequence into a sentence
def get_sentence(seq):
sentence = ''
for v in seq:
if v != 0: # 0 means it was just added to the sentence so it could have maxlen words
sentence += index_to_word[int(v)] + ' '
return sentence
# turn a sentence into a sequence
def gen_sequence(sentence):
seq = []
for word in sentence:
seq.append(word_to_index[word])
return np.asarray(seq, dtype=np.float32)
print('there are', num_words, 'words in the files')
# -
# ## Visualizing data
# +
# ------------------- negative
print('-' * 30)
print('Example of a negative review')
print('-' * 30)
x = open('data/train/neg/0_3.txt')
r = x.readline()
print(r)
# ------------------ positive
print()
print('-' * 30)
print('Example of a positive review')
print('-' * 30)
x = open('data/train/pos/0_9.txt')
r = x.readline()
print(r)
# +
print('Loading data')
(x_train, y_train), (x_test, y_test) = imdb.load_data(num_words=num_words)
# lets make things faster
limit = 3200
maxlen = 200
x_train = sequence.pad_sequences(x_train, maxlen=maxlen)
x_test = sequence.pad_sequences(x_test, maxlen=maxlen)
x_train = x_train[:limit].astype('float32')
y_train = y_train[:limit].astype('int32')
x_test = x_test[:limit].astype('float32')
y_test = y_test[:limit].astype('int32')
# y to onehot
y_train_one_hot = np.zeros((limit, 2), dtype=np.float32)
for i in range(limit):
y_train_one_hot[i][y_train[i]] = 1
y_test_one_hot = np.zeros((limit, 2), dtype=np.float32)
for i in range(limit):
y_test_one_hot[i][y_test[i]] = 1
#print(y_train)
#print(y_train_one_hot)
# +
# parameters
LEARNING_RATE = 0.01
BATCH_SIZE = 32
STEPS = 1000
# Define the model, using Keras
def model_fn(features, targets, mode, params):
embed = Embedding(num_words, 128)(features['x'])
gru = GRU(128)(embed)
logits = Dense(2)(gru)
logits_softmax = Activation('softmax')(logits)
# make logits shape the same as the targets: (BATCH_SIZE, 2)
if mode != learn.ModeKeys.PREDICT:
logits = tf.reshape(logits, shape=[BATCH_SIZE, 2])
logits_softmax = tf.reshape(logits, shape=[BATCH_SIZE, 2])
targets = tf.reshape(targets, shape=[BATCH_SIZE, 2])
loss = tf.losses.softmax_cross_entropy(
onehot_labels=targets, logits=logits)
train_op = tf.contrib.layers.optimize_loss(
loss=loss,
global_step=tf.contrib.framework.get_global_step(),
learning_rate=params["learning_rate"],
optimizer="Adam")
predictions = {
"probabilities": tf.nn.softmax(logits)
}
eval_metric_ops = {
"accuracy": tf.metrics.accuracy(
tf.argmax(input=logits_softmax, axis=1),
tf.argmax(input=targets, axis=1))
}
return model_fn_lib.ModelFnOps(
mode=mode,
predictions=predictions,
loss=loss,
train_op=train_op,
eval_metric_ops=eval_metric_ops)
# +
# In[ ]:
# Input functions
# this couldn't possibly be right...
x_train_dict = {'x': x_train }
train_input_fn = numpy_io.numpy_input_fn(
x_train_dict, y_train_one_hot, batch_size=BATCH_SIZE,
shuffle=False, num_epochs=2,
queue_capacity=1000, num_threads=1)
x_test_dict = {'x': x_test }
test_input_fn = numpy_io.numpy_input_fn(
x_test_dict, y_test_one_hot, batch_size=BATCH_SIZE, shuffle=False, num_epochs=1)
# In[ ]:
model_params = {"learning_rate": LEARNING_RATE}
# create estimator
estimator = tf.contrib.learn.Estimator(model_fn=model_fn, params=model_params)
# create experiment
def generate_experiment_fn():
"""
Create an experiment function given hyperparameters.
Returns:
A function (output_dir) -> Experiment where output_dir is a string
representing the location of summaries, checkpoints, and exports.
this function is used by learn_runner to create an Experiment which
executes model code provided in the form of an Estimator and
input functions.
All listed arguments in the outer function are used to create an
Estimator, and input functions (training, evaluation, serving).
Unlisted args are passed through to Experiment.
"""
def _experiment_fn(output_dir):
train_input = train_input_fn
test_input = test_input_fn
return tf.contrib.learn.Experiment(
estimator,
train_input_fn=train_input,
eval_input_fn=test_input
)
return _experiment_fn
# run experiment
learn_runner.run(generate_experiment_fn(), '/tmp/outputdir')
# -
# generate predictions
preds = list(estimator.predict(input_fn=test_input_fn))
# +
# number of outputs we want to see the prediction
NUM_EVAL = 10
def check_prediction(x, y, p, index):
print('prediction:', np.argmax(p[index]))
print('target:', np.argmax(y[index]))
print('sentence:', get_sentence(x[index]))
for i in range(NUM_EVAL):
index = np.random.randint(limit)
print('test:', index)
print('-' * 30)
print(np.asarray(x_test[index], dtype=np.int32))
check_prediction(x_test, y_test_one_hot, preds, index)
print()
# -
| code_samples/RNN/sentiment_analysis/.ipynb_checkpoints/keras-Copy3-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (sso-lc)
# language: python
# name: sso-lc
# ---
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
from lsst.sims.movingObjects import Orbits
modelfile = '/epyc/users/lynnej/sso/l7/L7SyntheticModel-v09.txt'
columns = ['a', 'e', 'inc', 'Omega', 'argPeri', 'meanAnomaly', 'H', 'j', 'type', 'inner', 'outer']
model = pd.read_csv(modelfile, delim_whitespace=True, comment='#', names=columns, index_col=False)
model = model.assign(epoch=2453157.50000 - 2400000.5)
print(len(model))
model[0:5]
subsetLen = 5000
# Pick a random subset of the model:
sub = model.sample(subsetLen)
plt.figure(figsize=(16, 7))
plt.subplot(1,2,1)
plt.plot(model.a, model.e, 'k.')
plt.plot(sub.a, sub.e, 'r.')
plt.xlabel("Semimajor axis (AU)")
plt.ylabel("Eccentricity")
plt.subplot(1,2,2)
plt.plot(model.a, model.inc, 'k.')
plt.plot(sub.a, sub.inc, 'r.')
plt.xlabel("Semimajor axis (AU)")
plt.ylabel("inclination (deg)")
# +
subsetLen = 5000
subpieces = 500
root = 'l7_5k'
subset = Orbits()
subset.setOrbits(model.sample(subsetLen))
print(len(model), len(subset), len(subset.orbits.objId.unique()))
subset.orbits.to_csv('%s.txt' % root, index=False, sep=' ')
x = np.arange(0, subsetLen+1, subpieces)
try:
os.mkdir('split')
except FileExistsError:
pass
for i, (xi, xo) in enumerate(zip(x[:-1], x[1:])):
subset.orbits[xi:xo].to_csv('split/%s_%d.txt' % (root, i), index=False, sep=' ')
# +
# snapshot
from lsst.sims.movingObjects import PyOrbEphemerides
pyephs = PyOrbEphemerides()
pyephs.setOrbits(subset)
ephs = pyephs.generateEphemerides(times = [subset.orbits.epoch.iloc[0]])
# -
plt.figure(figsize=(12, 7))
plt.plot(ephs['ra'], ephs['dec'], 'k.')
plt.xlabel('RA')
plt.ylabel('Dec')
| MAF_TEST/l7/L7 model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="47_7sBsU8JYt"
# ## UmojaHack Africa 2021 #3: Financial Resilience Challenge (BEGINNER) by UmojaHack Africa
#
# Can you predict if an individual will be able to make a payment in an emergency situation?
#
# The objective of this challenge is to build a machine learning model to predict which individuals across Africa and around the world are most likely to be financially resilient.
#
#
# 
#
# This is a simple Python starter notebook to get you started with the Financial Resilience Challenge.
#
# This notebook covers:
# - Loading the data
# - Simple EDA and an example of feature enginnering
# - Data preprocessing and data wrangling
# - Creating a simple model
# - Making a submission
# - Some tips for improving your score
# + id="GsxsCJyNFxBZ"
# Import libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.metrics import auc, classification_report, roc_auc_score
from sklearn.model_selection import train_test_split
from lightgbm import LGBMClassifier
from sklearn.ensemble import RandomForestClassifier
pd.set_option('max_colwidth', 500)
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
# + id="0_7BVS92U0Ti"
# Load files
train = pd.read_csv('Train.csv')
test = pd.read_csv('Test.csv')
samplesubmission = pd.read_csv('SampleSubmission.csv')
variable_definations = pd.read_csv('VariableDefinitions.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 635} id="f_LhTb6nU0Se" outputId="b9882c53-9626-41b1-da7e-0299c7d2a3f6"
# Preview the first five rows of the train set
train.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 602} id="c0KotvwrU0PP" outputId="9eef902c-3d8e-47a9-a340-6bc9c7aae1a0"
# Preview the first five rows of the test set
test.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="wU1bcDZSU0Nj" outputId="370f83aa-3fe6-475f-d2ed-d966a3431647"
# Preview the first five rows of the sample submission file
samplesubmission.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="UFh6oAKDFyws" outputId="b7801036-dfb5-4172-b045-b1155e723706"
# Preview variable definations
variable_definations
# + colab={"base_uri": "https://localhost:8080/"} id="PdVvmpG_GRxA" outputId="2efee873-8ff7-44de-8570-a7575c8cf0f4"
# Check the shape of the train and test sets
print(f'The shape of the train set is: {train.shape}\nThe shape of the test set is: {test.shape}')
# -
train.isnull().sum()
# + [markdown] id="9CO1R3hi8FJQ"
# ### Check for missing values
# + colab={"base_uri": "https://localhost:8080/", "height": 609} id="iWgHvxqYGRux" outputId="979fb804-723c-466c-81ff-7216d3e760cd"
# Check if there any missing values in train set
ax = train.isna().sum().sort_values().plot(kind = 'barh', figsize = (9, 10))
plt.title('Percentage of Missing Values Per Column in Train Set', fontdict={'size':15})
for p in ax.patches:
percentage ='{:,.0f}%'.format((p.get_width()/train.shape[0])*100)
width, height =p.get_width(),p.get_height()
x=p.get_x()+width+0.02
y=p.get_y()+height/2
ax.annotate(percentage,(x,y))
# + colab={"base_uri": "https://localhost:8080/", "height": 609} id="6A9jdEtoGRsi" outputId="ca04afd2-6372-4b8a-b8af-8ff177185145"
# Check if there missing values in test set
ax = test.isna().sum().sort_values().plot(kind = 'barh', figsize = (9, 10))
plt.title('Percentage of Missing Values Per Column in Test Set', fontdict={'size':15})
for p in ax.patches:
percentage ='{:,.1f}%'.format((p.get_width()/test.shape[0])*100)
width, height =p.get_width(),p.get_height()
x=p.get_x()+width+0.02
y=p.get_y()+height/2
ax.annotate(percentage,(x,y))
# + colab={"base_uri": "https://localhost:8080/"} id="PnGVLoE2GRqL" outputId="6f96e2e1-54f7-4d22-ee10-8ad69ec8bb8f"
# Check for duplicates
train.duplicated().any(), test.duplicated().any()
# -
# since the total count of 'Q7' is so small
# so filling data will decrease the reliablity
train = train.drop('Q7', axis=1)
test = test.drop('Q7', axis=1)
# Checking each Q column's count of 1.0, 2.0, 3.0, 4.0
for j in range(1,29):
if j == 7:
print(f'this is {j}')
continue
o, t, th, f = 0, 0, 0, 0
try:
for i in train[f'Q{j}']:
if i == 1.0: o+=1
elif i == 2.0: t+=1
elif i == 3.0: th+=1
elif i == 4.0: f+=1
print(f"this is Q{j}",len(train[f'Q{j}']))
print(o, t, th, f)
except:
try:
for i in train[f'Q{j}a']:
if i == 1.0: o+=1
elif i == 2.0: t+=1
elif i == 3.0: th+=1
elif i == 4.0: f+=1
print(f"this is Q{j}a",len(train[f'Q{j}a']))
print(o, t, th, f)
except:
try:
for i in train[f'Q{j}a']:
if i == 1.0: o+=1
elif i == 2.0: t+=1
elif i == 3.0: th+=1
elif i == 4.0: f+=1
print(f"this is Q{j}a",len(train[f'Q{j}a']))
print(o, t, th, f)
except:
pass
# since the next large null column is Q23
# fill the data using maximum count
train['Q23'] = train['Q23'].fillna(2.0)
test['Q23'] = test['Q23'].fillna(2.0)
# Check for duplicates
train.duplicated().any(), test.duplicated().any()
# + [markdown] id="SiZR9Zu6I5xo"
# ## Distribution of the target variable
# + colab={"base_uri": "https://localhost:8080/", "height": 404} id="J5xyqTXqHCYo" outputId="a5e94eb4-1475-4144-c4ea-0281f8784acd"
plt.figure(figsize=(7, 6))
sns.countplot(train.target)
plt.title('Target Variable Distribution');
# + [markdown] id="2r3BS3IcKP7f"
# This shows us that the target (Can you make a payment if you were in an emergency) is fairly balanced. The majority class in this dataset are people who can make a payment incase of an emergency
# + colab={"base_uri": "https://localhost:8080/", "height": 523} id="izjup23WHCVr" outputId="d4fc229a-9618-40f2-99c0-7aed0dbfe7fb"
# Q1 - Has ATM/debit card
plt.figure(figsize=(8, 7))
ax =sns.countplot(train.Q1)
ax.set_xticklabels(['Yes', 'No', 'Don"t Know', 'Refused to answer'], rotation=45 )
plt.title('Distribution of Q1 - Has ATM/debit card', fontdict = {'size': 15});
# + [markdown] id="4WpL_l95Nlxq"
# This shows that most people do not own a debit or ATM card
# + [markdown] id="g9oZ14qYN1FY"
# ## Combine train and test set for easy preprocessing
# + colab={"base_uri": "https://localhost:8080/"} id="8PH2LhntHCTj" outputId="b0c166b3-28bf-4dc4-f072-4219e99d4c0c"
# Combine train and test set
ntrain = train.shape[0] # to be used to split train and test set from the combined dataframe
all_data = pd.concat((train, test)).reset_index(drop=True)
print(f'The shape of the combined dataframe is: {all_data.shape}')
# + colab={"base_uri": "https://localhost:8080/"} id="4chstsvnHCRS" outputId="528302e0-53f9-4f9c-9289-9952fd05bb3a"
# Check the column names and datatypes
all_data.info()
# + colab={"base_uri": "https://localhost:8080/"} id="3AADdmJ6HCOr" outputId="c4b4a051-37e9-403d-f042-567cb49351e4"
# Category columns
cat_cols = ['country', 'region', 'owns_mobile'] + [x for x in all_data.columns if x.startswith('Q')]
num_cols = ['age', 'population']
# Change columns to their respective datatypes
all_data[cat_cols] = all_data[cat_cols].astype('category')
# Confirm whether the changes have been successful
all_data.info()
# + [markdown] id="_QFo9TxSkY6w"
# ### Number of unique values per categorical column
# + colab={"base_uri": "https://localhost:8080/"} id="2gc8aFu_kcU6" outputId="8eff12c2-e4c7-4ff8-cbd6-a31b66d5f592"
# Check unique values for each categorical column
for col in cat_cols:
print(col, all_data[col].nunique())
# +
# o, t, th, f = 0, 0, 0, 0
# for i in train['Q27']:
# if i == 1.0: o+=1
# elif i == 2.0: t+=1
# elif i == 3.0: th+=1
# elif i == 4.0: f+=1
# print(len(train['Q27']))
# print(o, t, th, f)
# + [markdown] id="wUA3IqW42Oeq"
# ## Fill in missing values
# Missing values can be filled using different strategies
# - Mean
# - Max
# - Min
# - for categorical variables - mode
# - [sklearn SimpleImputer](https://scikit-learn.org/stable/modules/generated/sklearn.impute.SimpleImputer.html)
# - Others... do more reasearch
# + colab={"base_uri": "https://localhost:8080/"} id="VQ4t3UbC2Nsy" outputId="176e804c-12a8-4f76-8a12-af4de9a2665f"
# Fill in missing values
# For cat cols and date cols fill in with mode and for num cols fill in with 9999
from sklearn.impute import SimpleImputer
imp = SimpleImputer(strategy='mean')
for col in all_data.columns:
if col in cat_cols:
all_data[col] = all_data[col].fillna(all_data[col].mode()[0])
elif col in num_cols:
all_data[col] = imp.fit_transform(np.array(all_data[col]).reshape(-1, 1))
# all_data[col] = all_data[col].fillna(all_data[col].fillna(9999))
# Confirm that there aren't any missing values
all_data[all_data.columns.difference(['target'])].isna().sum().any()
# -
# fill age column
imp = SimpleImputer(strategy='mean')
train['age'] = imp.fit_transform(np.array(train['age']).reshape(-1, 1))
train.isnull().sum()
# + [markdown] id="3Hd9viOb2_c3"
# ### Feature Engineering
# #### Try different strategies of dealing with categorical variables
# - One hot encoding
# - Label encoding
# - Target encoding
# - Reduce the number of unique values...
# + colab={"base_uri": "https://localhost:8080/"} id="X2DUKqTb3UY-" outputId="428bb9c0-2d5c-4d68-b206-4ac1069a613d"
# Shape of data before encoding
all_data.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 278} id="vQtjAQw92Np4" outputId="782fe9cb-276c-45f1-de3a-861f918fac54"
# Use one hot encoding to turn categorical features to numerical features
# Encode categorical features
all_data = pd.get_dummies(data = all_data, columns = cat_cols)
all_data.head()
# + colab={"base_uri": "https://localhost:8080/"} id="xijxGVyB3Z3K" outputId="aaeda529-cd48-4d0d-e3d3-941e7e518c26"
# Shape of data after encoding
all_data.shape
# + colab={"base_uri": "https://localhost:8080/"} id="xXRt7bHS3fhH" outputId="c21bc672-e690-4c0e-a106-304978846a4f"
# Separate train and test data from the combined dataframe
train_df = all_data[:ntrain]
test_df = all_data[ntrain:]
# Check the shapes of the split dataset
train_df.shape, test_df.shape
# + [markdown] id="eDBSVM-S4E-h"
# ### Training and making predictions
#
# - Is lgbm the best model for this challenge?
# - Parameter tuning
# - Grid search, random search, perhaps bayesian search works better...
# + colab={"base_uri": "https://localhost:8080/"} id="MzMVccR34ARy" outputId="b74bc80f-ec2a-41cb-de11-7e1793bb92c3"
# Select main columns to be used in training
main_cols = all_data.columns.difference(['ID', 'target'])
X = train_df[main_cols]
y = train_df.target.astype(int)
# Split data into train and test sets
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.3, random_state=42)
# Train model
model = RandomForestClassifier()
model.fit(X_train, y_train)
# Make predictions
y_pred = model.predict_proba(X_test)[:, 1]
# Check the auc score of the model
print(f'RandomForest AUC score on the X_test is: {roc_auc_score(y_test, y_pred)}\n')
# print classification report
print(classification_report(y_test, [1 if x >= 0.5 else 0 for x in y_pred]))
# + [markdown] id="tFULqvbM659e"
# ### Train different model and compare results
# +
# Select main columns to be used in training
main_cols = all_data.columns.difference(['ID', 'target'])
X = train_df[main_cols]
y = train_df.target.astype(int)
# Split data into train and test sets
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.13, random_state=41)
# +
# Train model
model2 = LGBMClassifier()
model2.fit(X_train.values, y_train.values)
# Make predictions
y_pred = model2.predict_proba(X_test)[:, 1]
# Check the auc score of the model
print(f'LGBM AUC score on the X_test is: {roc_auc_score(y_test, y_pred)}\n')
# print classification report
print(classification_report(y_test, [1 if x >= 0.5 else 0 for x in y_pred]))
# +
# Select main columns to be used in training
main_cols = all_data.columns.difference(['ID', 'target'])
X = train_df[main_cols]
y = train_df.target.astype(int)
# Split data into train and test sets
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.3, random_state=41)
# +
from xgboost import XGBClassifier
# Train model
model3 = XGBClassifier()
model3.fit(X_train.values, y_train.values)
# Make predictions
y_pred = model3.predict_proba(X_test)[:, 1]
# Check the auc score of the model
print(f'XGB AUC score on the X_test is: {roc_auc_score(y_test, y_pred)}\n')
# print classification report
print(classification_report(y_test, [1 if x >= 0.5 else 0 for x in y_pred]))
# +
# Select main columns to be used in training
main_cols = all_data.columns.difference(['ID', 'target'])
X = train_df[main_cols]
y = train_df.target.astype(int)
# Split data into train and test sets
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.3, random_state=41)
# +
from catboost import CatBoostClassifier
# Train model
seed = 42
model = CatBoostClassifier(iterations=1000, verbose=0)
model.fit(X_train.values, y_train.values)
# Make predictions
y_pred = model.predict_proba(X_test)[:, 1]
# Check the auc score of the model
print(f'CatBoostClassifier AUC score on the X_test is: {roc_auc_score(y_test, y_pred)}\n')
# print classification report
print(classification_report(y_test, [1 if x >= 0.5 else 0 for x in y_pred]))
# +
# Make prediction on the test set
test_df = test_df[main_cols]
# model.fit(X,y)
model2.fit(X.values,y)
# model3.fit(X,y)
predictions1 = model.predict_proba(test_df)[:, 1]
predictions2 = model2.predict_proba(test_df)[:, 1]
# predictions3 = model3.predict_proba(test_df)[:, 1]
predictions = (predictions1*0.6 + predictions2*0.3) #+ predictions3*0.1)
# # Create a submission file
sub_file = samplesubmission.copy()
sub_file.target = predictions
# # Check the distribution of your predictions
sns.countplot([1 if x >= 0.5 else 0 for x in sub_file.target])
plt.title('Predicted Variable Distribution');
# + [markdown] id="bXKU5avW6uaT"
# - There is a light improvement when using LGBMClassifier, XGBoostClassifier, CatBoostClassifier
#
# [More on AUC score](https://developers.google.com/machine-learning/crash-course/classification/roc-and-auc#:~:text=AUC%20represents%20the%20probability%20that,has%20an%20AUC%20of%201.0.)
# + [markdown] id="2L7-CX1c7l-u"
# ### Making predictions of the test set and creating a submission file
# + colab={"base_uri": "https://localhost:8080/", "height": 281} id="2CY70-KV5lYO" outputId="a928aaf6-c5c9-4e76-eda1-377e999617a1"
# Make prediction on the test set
test_df = test_df[main_cols]
predictions = model.predict_proba(test_df)[:, 1]
# # Create a submission file
sub_file = samplesubmission.copy()
sub_file.target = predictions
# # Check the distribution of your predictions
sns.countplot([1 if x >= 0.5 else 0 for x in sub_file.target])
plt.title('Predicted Variable Distribution');
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="oroCrCK27v-4" outputId="6de7be09-47f7-41d7-e59e-1c30726c8c3f"
# Create a csv file and upload to zindi
sub_file.to_csv('Baseline.csv', index = False)
sub_file.head()
# + [markdown] id="co8AT8lL7y5s"
# ###More Tips
# - Thorough EDA and domain knowledge sourcing
# - Re-group Categorical features
# - More Feature Engineering
# - Dataset balancing - oversampling, undersampling, SMOTE...
# - Ensembling of models
# - Cross-validation: Group folds, Stratified...
# + [markdown] id="ih0ejC9w7617"
# # ******************* GOOD LUCK!!! ***************************
| Initial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Import Library
import pandas
import configparser
import psycopg2
# +
config = configparser.ConfigParser()
config.read('config.ini')
host = config['myaws']['host']
db = config['myaws']['db']
user = config['myaws']['user']
pwd = config['myaws']['pwd']
# -
conn = psycopg2.connect(host = host,
user = user,
password = <PASSWORD>,
dbname = db
)
cur = conn.cursor()
# # Create House Table
table_sql = """
CREATE TABLE IF NOT EXISTS gp23.house
(
price integer,
bed integer,
bath integer,
area integer,
address VARCHAR(200),
PRIMARY KEY(address)
);
"""
conn.rollback()
# +
cur.execute(table_sql)
conn.commit()
# -
# # Define the Table
# +
url = 'https://www.trulia.com/VA/Ashburn/20147/'
# -
import urllib.request
response = urllib.request.urlopen(url)
html_data= response.read()
print(html_data.decode('utf-8'))
# +
from bs4 import BeautifulSoup
soup = BeautifulSoup(html_data,'html.parser')
print (soup)
# -
for li_class in soup.find_all('li', class_ = 'Grid__CellBox-sc-144isrp-0 SearchResultsList__WideCell-b7y9ki-2 jiZmPM'):
try:
for price_div in li_class.find_all('div',{'data-testid':'property-price'}):
price =int(price_div.text.replace('$','').replace(",",""))
for bed_div in li_class.find_all('div', {'data-testid':'property-beds'}):
bed= int(bed_div.text.replace('bd','').replace(",",""))
for bath_div in li_class.find_all('div',{'data-testid':'property-baths'}):
bath =int(bath_div.text.replace('ba','').replace(",",""))
for area_div in li_class.find_all('div',{'data-testid':'property-floorSpace'}):
area=int(area_div.text.split('sqft')[0].replace(",",""))
for address_div in li_class.find_all('div',{'data-testid':'property-address'}):
address =address_div.text
try:
sql_insert = """
insert into gp23.house(price,bed,bath,area,address)
values('{}','{}','{}','{}','{}')
""".format(price,bed,bath,area,address)
cur.execute(sql_insert)
conn.commit()
except:
conn.rollback()
except:
pass
# # Query the Table
df = pandas.read_sql_query('select * from gp23.house ', conn)
df[:]
# # Basic Stat
df.describe()
# # Price Distribution
df['price'].hist()
# # Bed vs Bath
df.plot.scatter(x='bed',y='bath')
| Lab06.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Global Fishing Effort Purse Seiners
import time
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.basemap import Basemap
from matplotlib import colors,colorbar
import matplotlib
# %matplotlib inline
import csv
import math
# from scipy import stats
import bq
client = bq.Client.Get()
def Query(q):
t0 = time.time()
answer = client.ReadTableRows(client.Query(q)['configuration']['query']['destinationTable'])
print 'Query time: ' + str(time.time() - t0) + ' seconds.'
return answer
# +
q = '''
SELECT
INTEGER(latitude*2) lat_bin,
INTEGER(longitude*2) lon_bin,
SUM(hours_since_last_timestamp) fishing_hours,
avg(distance_to_shore) distance_to_shore
FROM (
SELECT
latitude,
longitude,
CASE WHEN hours_since_last_timestamp>12 THEN 12
WHEN hours_since_last_timestamp IS NULL THEN 0
ELSE hours_since_last_timestamp END hours_since_last_timestamp,
distance_to_shore
FROM
[scratch_global_fishing_raster.Jan_July_2015_FishingVessels_Time]
WHERE
mmsi IN (
SELECT
mmsi
FROM
[scratch_global_fishing_raster.classification_20160324]
WHERE
label = 'Purse seine' and score >.5)
AND sog < 3
// AND sog < 5.5
and distance_to_shore>5
)
GROUP BY
lat_bin,
lon_bin
'''
fishing_grid = Query(q)
# +
cellsize = .5
one_over_cellsize = 2
max_lat = 90
min_lat = -90
min_lon = -180
max_lon = 180
num_lats = (max_lat-min_lat)*one_over_cellsize
num_lons = (max_lon-min_lon)*one_over_cellsize
grid = np.zeros(shape=(num_lats,num_lons))
for row in fishing_grid:
lat = int(row[0])
lon = int(row[1])
lat_index = lat-min_lat*one_over_cellsize
lon_index = lon-min_lon*one_over_cellsize
grid[lat_index][lon_index] = float(row[2])
# +
plt.rcParams["figure.figsize"] = [12,7]
cutoff = 0 # 4 degress away from the pole
firstlat = 90-cutoff
lastlat = -90+cutoff
firstlon = -180
lastlon = 180
scale = cellsize
one_over_cellsize = 2
fishing_days_truncated = grid[one_over_cellsize*cutoff:(180*one_over_cellsize)-cutoff*one_over_cellsize][:]
numlats = int((firstlat-lastlat)*one_over_cellsize+.5)
numlons = int((lastlon-firstlon)*one_over_cellsize+.5)
lat_boxes = np.linspace(lastlat,firstlat,num=numlats,endpoint=False)
lon_boxes = np.linspace(firstlon,lastlon,num=numlons,endpoint=False)
fig = plt.figure()
m = Basemap(llcrnrlat=lastlat, urcrnrlat=firstlat,
llcrnrlon=lastlon, urcrnrlon=firstlon, lat_ts=0, projection='robin',resolution="h", lon_0=0)
m.drawmapboundary(fill_color='#111111')
# m.drawcoastlines(linewidth=.2)
m.fillcontinents('#111111',lake_color='#111111')#, lake_color, ax, zorder, alpha)
x = np.linspace(-180, 180, 360*one_over_cellsize)
y = np.linspace(lastlat, firstlat, (firstlat-lastlat)*one_over_cellsize)
x, y = np.meshgrid(x, y)
converted_x, converted_y = m(x, y)
from matplotlib import colors,colorbar
maximum = grid.max()
minimum = 1
norm = colors.LogNorm(vmin=minimum, vmax=maximum)
# norm = colors.Normalize(vmin=0, vmax=1000)
m.pcolormesh(converted_x, converted_y, fishing_days_truncated, norm=norm, vmin=minimum, vmax=maximum, cmap = plt.get_cmap('viridis'))
t = "Fishing Hours for Purse Seiners, January 2015 to June 2015\nOnly Likely Fishing Vessesl - 4,731 Vessels "
plt.title(t, color = "#ffffff", fontsize=18)
ax = fig.add_axes([0.2, 0.1, 0.4, 0.02]) #x coordinate ,
norm = colors.LogNorm(vmin=minimum, vmax=maximum)
# norm = colors.Normalize(vmin=0, vmax=1000)
lvls = np.logspace(np.log10(minimum),np.log10(maximum),num=8)
cb = colorbar.ColorbarBase(ax,norm = norm, orientation='horizontal', ticks=lvls, cmap = plt.get_cmap('viridis'))
the_labels = []
for l in lvls:
if l>=1:
l = int(l)
the_labels.append(l)
#cb.ax.set_xticklabels(["0" ,round(m3**.5,1), m3, round(m3**1.5,1), m3*m3,round(m3**2.5,1), str(round(m3**3,1))+"+"], fontsize=10)
cb.ax.set_xticklabels(the_labels, fontsize=10, color = "#ffffff")
cb.set_label('Fishing Hours by Two Degree Grid',labelpad=-40, y=0.45, color = "#ffffff")
ax.text(1.7, -0.5, 'Data Source: Orbcomm\nMap by Global Fishing Watch',
verticalalignment='bottom', horizontalalignment='right',
transform=ax.transAxes,
color='#ffffff', fontsize=6)
plt.savefig("fishing_hours_purseseiners_2015Jan-Jun.png",bbox_inches='tight',dpi=300,transparent=True,pad_inches=.1, facecolor="#000000")
plt.show()
# +
q = '''
SELECT
INTEGER(latitude*2) lat_bin,
INTEGER(longitude*2) lon_bin,
SUM(hours_since_last_timestamp) fishing_hours,
avg(distance_from_shore) distance_from_shore
FROM (
SELECT
lat latitude,
lon longitude,
CASE WHEN hours_since_last_timestamp>12 THEN 12
WHEN hours_since_last_timestamp IS NULL THEN 0
ELSE hours_since_last_timestamp END hours_since_last_timestamp,
distance_from_shore
FROM
[scratch_global_fishing_raster.Jan_July_2015_AllVessels_Time]
WHERE
mmsi IN (
SELECT
mmsi
FROM
[scratch_global_fishing_raster.classification_20160324]
WHERE
label = 'Purse seine')
AND sog < 3
//AND sog < 5.5
and distance_from_shore>5
)
GROUP BY
lat_bin,
lon_bin
'''
fishing_grid_allvessels = Query(q)
# +
cellsize = .5
one_over_cellsize = 2
max_lat = 90
min_lat = -90
min_lon = -180
max_lon = 180
num_lats = (max_lat-min_lat)*one_over_cellsize
num_lons = (max_lon-min_lon)*one_over_cellsize
grid = np.zeros(shape=(num_lats,num_lons))
for row in fishing_grid_allvessels:
lat = int(row[0])
lon = int(row[1])
lat_index = lat-min_lat*one_over_cellsize
lon_index = lon-min_lon*one_over_cellsize
grid[lat_index][lon_index] = float(row[2])
# +
plt.rcParams["figure.figsize"] = [12,7]
cutoff = 0 # 4 degress away from the pole
firstlat = 90-cutoff
lastlat = -90+cutoff
firstlon = -180
lastlon = 180
scale = cellsize
one_over_cellsize = 2
fishing_days_truncated = grid[one_over_cellsize*cutoff:(180*one_over_cellsize)-cutoff*one_over_cellsize][:]
numlats = int((firstlat-lastlat)*one_over_cellsize+.5)
numlons = int((lastlon-firstlon)*one_over_cellsize+.5)
lat_boxes = np.linspace(lastlat,firstlat,num=numlats,endpoint=False)
lon_boxes = np.linspace(firstlon,lastlon,num=numlons,endpoint=False)
fig = plt.figure()
m = Basemap(llcrnrlat=lastlat, urcrnrlat=firstlat,
llcrnrlon=lastlon, urcrnrlon=firstlon, lat_ts=0, projection='robin',resolution="h", lon_0=0)
m.drawmapboundary(fill_color='#111111')
# m.drawcoastlines(linewidth=.2)
m.fillcontinents('#111111',lake_color='#111111')#, lake_color, ax, zorder, alpha)
x = np.linspace(-180, 180, 360*one_over_cellsize)
y = np.linspace(lastlat, firstlat, (firstlat-lastlat)*one_over_cellsize)
x, y = np.meshgrid(x, y)
converted_x, converted_y = m(x, y)
from matplotlib import colors,colorbar
maximum = grid.max()
minimum = 1
norm = colors.LogNorm(vmin=minimum, vmax=maximum)
# norm = colors.Normalize(vmin=0, vmax=1000)
m.pcolormesh(converted_x, converted_y, fishing_days_truncated, norm=norm, vmin=minimum, vmax=maximum, cmap = plt.get_cmap('viridis'))
t = "Fishing Hours for Purse Seiners, January 2015 to June 2015\nAll Classified Vessels - 13,945 Vessels"
plt.title(t, color = "#ffffff", fontsize=18)
ax = fig.add_axes([0.2, 0.1, 0.4, 0.02]) #x coordinate ,
norm = colors.LogNorm(vmin=minimum, vmax=maximum)
# norm = colors.Normalize(vmin=0, vmax=1000)
lvls = np.logspace(np.log10(minimum),np.log10(maximum),num=8)
cb = colorbar.ColorbarBase(ax,norm = norm, orientation='horizontal', ticks=lvls, cmap = plt.get_cmap('viridis'))
the_labels = []
for l in lvls:
if l>=1:
l = int(l)
the_labels.append(l)
#cb.ax.set_xticklabels(["0" ,round(m3**.5,1), m3, round(m3**1.5,1), m3*m3,round(m3**2.5,1), str(round(m3**3,1))+"+"], fontsize=10)
cb.ax.set_xticklabels(the_labels, fontsize=10, color = "#ffffff")
cb.set_label('Fishing Hours by Two Degree Grid',labelpad=-40, y=0.45, color = "#ffffff")
ax.text(1.7, -0.5, 'Data Source: Orbcomm\nMap by Global Fishing Watch',
verticalalignment='bottom', horizontalalignment='right',
transform=ax.transAxes,
color='#ffffff', fontsize=6)
plt.savefig("fishing_hours_purseseiner_2015Jan-Jun_allvessels.png",bbox_inches='tight',dpi=300,transparent=True,pad_inches=.1, facecolor="#000000")
plt.show()
# +
# how many vessels?
q = '''
SELECT
count(distinct mmsi)
FROM
[scratch_global_fishing_raster.Jan_July_2015_FishingVessels_Time]
WHERE
mmsi IN (
SELECT
mmsi
FROM
[scratch_global_fishing_raster.classification_20160324]
WHERE
label = 'Purse seine' and score >.5)
// AND sog > 2.5
AND sog < 3
and distance_to_shore>5
'''
likely_fishing = Query(q)
# +
# how many vessels?
q = '''
SELECT
count(distinct mmsi)
FROM
[scratch_global_fishing_raster.Jan_July_2015_AllVessels_Time]
WHERE
mmsi IN (
SELECT
mmsi
FROM
[scratch_global_fishing_raster.classification_20160324]
WHERE
label = 'Purse seine' and score >.5)
// AND sog > 2.5
AND sog < 3
and distance_from_shore>5
'''
all_vessels = Query(q)
# -
number_of_classified = Query('select count(*) from [scratch_global_fishing_raster.classification_20160324]')
print "Classified Purse Seiners:", all_vessels[0][0]
print "Classified Purse Seiners that are Likely Fishing Vessels:", likely_fishing[0][0]
print "All Classified Vessels of All Types:", number_of_classified[0][0]
| initial_maps/Global_Fishing_Effort_PurseSeiners_V0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The GP-SFH module
# ## Creating different shapes using SFH-tuples
#
# The `dense_basis` code contains a module for creating smooth star formation history from a tuple consisting of (M$_*$, SFR, {$t_X$}) - the stellar mass, star formation rate, and a set of lookback times at which the galaxy forms N equally spaced quantiles of its stellar mass.
#
# This parametrization comes with a lot of flexibility, and allows us to create a large range of SFH shapes even with a small number of parameters. Here we show a few examples, showing how we create a variety of different SFH shapes with just 2 free parameters - the SFR and the t$_{50}$.
import dense_basis as db
import numpy as np
import matplotlib.pyplot as plt
Nparam = 1
redshift = 1.0
logMstar = 10.0
# Let's start with an `SFH that is rising` throughout a galaxy's lifetime, such as may be expected for high-redshift star forming galaxies. Since we are considering a galaxy with $M_* = 10^{10}M_\odot$ at z=1, we choose a reasonably high SFR of 10 $M_\odot/yr$. Since the SFR is rising, we also choose a short $t_{50}$, since it is rapidly building forming its stars. Running this through the model, we get:
# +
logSFR = 1.0
t50 = 0.6 # t50, lookback time, in Gyr
sfh_tuple = np.hstack([logMstar, logSFR, Nparam, db.scale_t50(t50,redshift)])
sfh, timeax = db.tuple_to_sfh(sfh_tuple, redshift)
fig = db.plot_sfh(timeax, sfh, lookback=True)
plt.title('Rising SFH')
plt.show()
# -
# We next consider the case of reasonably `steady star formation`. This is different from constant star formation, because SFR goes to 0 smoothly as we approach the big bang. In this case, we choose an SFR closer to the expected lifetime average for a massive galaxy at z=1, and a $t_{50}$ close to half the age of the universe at the redshift of observation. Doing this gives us:
# +
logSFR = 0.335
t50 = 2.3 # t50, lookback time, in Gyr
sfh_tuple = np.hstack([logMstar, logSFR, Nparam, db.scale_t50(t50,redshift)])
sfh, timeax = db.tuple_to_sfh(sfh_tuple, redshift)
fig = db.plot_sfh(timeax, sfh, lookback=True)
plt.title('Steady SFH')
plt.show()
# -
# We now look at the class of quenched and quenching galaxies.
#
# For the `post-starburst SFH`, we create a similar setup to the rising SFH, but with a low SFR at the time of observation. Since the galaxy still formed a lot of stars in the recent past but is not doing so now, this creates the distinctive post-starburst shape.
# +
logSFR = 0.5
t50 = 0.6 # t50, lookback time, in Gyr
sfh_tuple = np.hstack([logMstar, logSFR, Nparam, db.scale_t50(t50,redshift)])
sfh, timeax = db.tuple_to_sfh(sfh_tuple, redshift)
fig = db.plot_sfh(timeax, sfh, lookback=True)
plt.title('Post-starburst SFH')
plt.show()
# -
# We also consider two simple types of `quenched galaxies`, obtained easily by setting the recent SFR to a very low value. To consider the different possible shapes for a quenched SFH, we use a recent and an older value for the $t_{50}$, to obtain SFHs that quenched either gradually or aburptly.
# +
logSFR = -3.0
t50 = 4.6 # t50, lookback time, in Gyr
sfh_tuple = np.hstack([logMstar, logSFR, Nparam, db.scale_t50(t50,redshift)])
sfh, timeax = db.tuple_to_sfh(sfh_tuple, redshift)
fig = db.plot_sfh(timeax, sfh, lookback=True)
plt.title('Old Quiescent SFH')
plt.show()
# +
logSFR = -3.0
t50 = 1.7 # t50, lookback time, in Gyr
sfh_tuple = np.hstack([logMstar, logSFR, Nparam, db.scale_t50(t50,redshift)])
sfh, timeax = db.tuple_to_sfh(sfh_tuple, redshift)
fig = db.plot_sfh(timeax, sfh, lookback=True)
plt.title('Young Quiescent SFH')
plt.show()
# -
# Finally, we also consider the case of a `rejuvenated SFH`, which had a significant lull between two periods of active star formation. To create an example of this kind of SFH, we use a reasonably large $t_{50}$, which tells the GP-SFH module that the galaxy formed 50% of its stars early on. Coupled with an SFR that indicates active star formation at the time of observation, this means that there had to be a period between these two when the galaxy did not form a lot of mass, leading to this distinctive shape.
# +
logSFR = 0.5
t50 = 4.0 # t50, lookback time, in Gyr
sfh_tuple = np.hstack([logMstar, logSFR, Nparam, db.scale_t50(t50,redshift)])
sfh, timeax = db.tuple_to_sfh(sfh_tuple, redshift)
fig = db.plot_sfh(timeax, sfh, lookback=True)
plt.title('Rejuvenated SFH')
plt.show()
# -
| docs/tutorials/the_gp_sfh_module.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cnosolar as cno
# ## 1. Configuración Inicial
# +
# Información Geográfica
latitude = 9.789103 #Latitud
longitude = -73.722451 #Longitud
tz = 'America/Bogota' #Huso Horario
altitude = 50 #Altitud
surface_type = 'soil' #irradiance.SURFACE_ALBEDOS
surface_albedo = 0.17
# Base de Datos
file_name = './data/tmy_elpaso.csv'
# Configuración del Sistema
## Inversor
inverters_database = 'CECInverter' # ['CECInverter', 'SandiaInverter', 'ADRInverter']
inverter_name = 'Power_Electronics__FS1275CU15__690V_'
## Módulo
modules_database = 'CECMod' # ['CECMod', 'SandiaMod']
module_name = 'Jinko_Solar_Co___Ltd_JKM345M_72'
module_type = 'open_rack_glass_polymer' #['open_rack_glass_glass', 'close_mount_glass_glass', 'open_rack_glass_polymer', 'insulated_back_glass_polymer']
surface_azimuth=None
surface_tilt=None
## Tracker
with_tracker = True
axis_tilt = 0
axis_azimuth = 180 #Heading south
max_angle = 60
racking_model = 'open_rack'
module_height = None
## Arreglo
num_arrays = 2
per_mppt = 1
modules_per_string = [30, 30] #Modules Per String
strings_per_inverter = [168, 6] #Strings Per Inverter
# Modelado de Producción con PVlib
ac_model='sandia'
loss = 26.9
num_inverter = 4*12
resolution = 60
energy_units = 'Wh'
# -
# ## 2. Base de Datos
df = cno.data.load_csv(file_name=file_name, tz=tz)
df.head()
# ## 3. Datos Meteorológicos
# +
location, solpos, airmass, etr_nrel = cno.location_data.get_parameters(latitude,
longitude,
tz,
altitude,
datetime=df.index)
solpos.head()
# -
# ## 4. Descomposición + Transposición
# ### 4.1. Descomposición: DISC
# +
disc = cno.irradiance_models.decomposition(ghi=df.GHI,
solpos=solpos,
datetime=df.index)
disc.head()
# -
# ### 4.2. Montaje y Seguidor
# +
mount, tracker = cno.pvstructure.get_mount_tracker(with_tracker=with_tracker,
surface_tilt=surface_tilt,
surface_azimuth=surface_azimuth,
solpos=solpos,
axis_tilt=axis_tilt,
axis_azimuth=axis_azimuth,
max_angle=max_angle,
racking_model=racking_model,
module_height=module_height)
print('MOUNT\n', mount, '\n')
print('TRACKER\n', tracker)
# -
# ### 4.3. Transposición: Perez-Ineichen 1990
# +
poa = cno.irradiance_models.transposition(with_tracker=with_tracker,
tracker=tracker,
surface_tilt=surface_tilt,
surface_azimuth=surface_azimuth,
solpos=solpos,
disc=disc,
ghi=df.GHI,
etr_nrel=etr_nrel,
airmass=airmass,
surface_albedo=surface_albedo,
surface_type=surface_type)
poa.head()
# -
# ## 5. Configuración del Sistema
# ### 5.1. Inversor y Módulo
# +
inverter = cno.components.get_inverter(inverters_database=inverters_database,
inverter_name=inverter_name,
inv=None)
print('INVERTER\n', inverter, '\n')
module = cno.components.get_module(modules_database=modules_database,
module_name=module_name,
mod=None)
print('MODULE\n', module)
# -
# ### 5.2. Arreglos Fotovoltaicos
# +
string_arrays = cno.def_pvsystem.get_arrays(mount=mount,
surface_albedo=surface_albedo,
surface_type=surface_type,
module_type=module_type,
module=module,
mps=modules_per_string,
spi=strings_per_inverter)
string_arrays
# -
# ### 5.3. Sistema Fotovoltaico
# +
system = cno.def_pvsystem.get_pvsystem(with_tracker=with_tracker,
tracker=tracker,
string_array=string_arrays,
surface_tilt=surface_tilt,
surface_azimuth=surface_azimuth,
surface_albedo=surface_albedo,
surface_type=surface_type,
module_type=module_type,
module=module,
inverter=inverter,
racking_model=racking_model)
system
# -
# ## 6. Modelado de Producción con PVlib
# ### 6.1. Temperatura de las Celdas
# +
temp_cell = cno.cell_temperature.from_tnoct(poa=poa.poa_global,
temp_air=df['Temperature'],
tnoct=module['T_NOCT'],
mount_temp=0)
temp_cell.head()
# -
# ### 6.2. Producción DC, Potencia AC y Energía
dc, ac, energy = cno.production.production_pipeline(poa=poa.poa_global,
cell_temperature=temp_cell,
module=module,
inverter=inverter,
system=system,
ac_model=ac_model,
loss=loss,
resolution=resolution,
num_inverter=num_inverter,
per_mppt=per_mppt,
energy_units=energy_units)
dc
ac
energy
# ## 7. Capacidad Efectiva Neta (CEN)
# +
route()
import cno_cen
perc = 99 # Percentil CEN
cen_per, cen_pmax = cno_cen.get_cen(ac=ac,
perc=perc,
decimals=2,
curve=True)
# -
# ## 8. ENFICC
# +
route()
import cno_energia_firme
# Prepare Data
df_hora = df[['GHI', 'Temperature']]
df_hora['GHI'] = df_hora['GHI'] / 1000 # W to kW
df_hora = df_hora.loc[df_hora.GHI != 0]
ghi = df_hora.resample('M').apply(lambda x: x.quantile(0.95)).GHI # https://stackoverflow.com/questions/39246664/calculate-percentiles-quantiles-for-a-timeseries-with-resample-or-groupby-pand
insolation = df_hora['GHI'].resample('M').sum() # kWh/m2 / month
temp = df_hora['Temperature'].resample('M').mean()
df_mes = pd.DataFrame({'GHI': ghi, 'Insolation': insolation,'Temperature': temp})
# ENFICC CREG 201 de 2017
efirme, enficc_t = cno_energia_firme.enficc_creg(df=df_mes,
Kinc=1.1981,
IHF=0.1,
CEN=cen_per,
a=1.10e-05,
b=-0.0007,
c=0.0185,
d=-0.1157,
Kmedt=0.8540)
# Energía Firme PVlib + CREG
__, enficc_v2 = cno_energia_firme.efirme_pvlib_creg(energy=energy)
# Energía Firme: PVlib + Min
enficc_v3 = cno_energia_firme.efirme_pvlib_min(energy=energy)
# Energía Firme: PVlib + Percentil
enficc_v4 = cno_energia_firme.efirme_pvlib_percentile(energy=energy,
percentile=95)
# +
#Energy Error Comparison Plot
months = ['Jan\n2019', 'Feb', 'Mar', 'Abr', 'May', 'Jun', 'Jul', 'Ago', 'Sep', 'Oct', 'Nov', 'Dec',
'Jan\n2020', 'Feb', 'Mar', 'Abr', 'May', 'Jun', 'Jul', 'Ago', 'Sep', 'Oct', 'Nov', 'Dec']
x = np.arange(len(months))
#Bar Plots
route()
import cno_plots_metrics
hor, ver = 13, 5
plt.figure(figsize=(hor,ver))
plt.bar(x, energy['month']['energy'].tail(12*2), color='#1580E4',
label=f'ENFICC CREG 201 = {enficc_t} kWh/día\
\n\nEF PVlib-CREG = {enficc_v2} kWh/día\
\n\nEF PVlib-Min = {enficc_v3} kWh/día\
\n\nEF PVlib-Perc ({perc} %) = {enficc_v4} kWh/día')
plt.xticks(x, months);
cno_plots_metrics.plot_specs(title='Energía Mensual',
ylabel=f'Energía, ${energy_units}$',
xlabel='Tiempo',
rot=0,
ylim_min=0, ylim_max=None,
xlim_min=None, xlim_max=None,
loc='best')
plt.legend(loc='best', bbox_to_anchor=(1,1), fontsize=9.5);
# -
# ## 9. Relación Recurso-Potencia
# +
route()
import cno_recurso_potencia
cno_recurso_potencia.get_curve(poa=poa.poa_global,
ac=ac,
ac_units='MW')
# -
# ---
| .ipynb_checkpoints/CuadernoInteractivo_ElPaso-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://www.kaggle.com/code/stiwar1/tps-may22-data-preprocessing?scriptVersionId=95242244" target="_blank"><img align="left" alt="Kaggle" title="Open in Kaggle" src="https://kaggle.com/static/images/open-in-kaggle.svg"></a>
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 1.16602, "end_time": "2022-05-10T12:34:06.725884", "exception": false, "start_time": "2022-05-10T12:34:05.559864", "status": "completed"} tags=[]
import numpy as np
import pandas as pd
pd.set_option('precision', 4)
pd.set_option('display.max_columns', None)
from sklearn.model_selection import StratifiedKFold
SEED = 2311
# + papermill={"duration": 14.99395, "end_time": "2022-05-10T12:34:21.737475", "exception": false, "start_time": "2022-05-10T12:34:06.743525", "status": "completed"} tags=[]
train = pd.read_csv('../input/tabular-playground-series-may-2022/train.csv')
test = pd.read_csv('../input/tabular-playground-series-may-2022/test.csv')
# + papermill={"duration": 0.05662, "end_time": "2022-05-10T12:34:21.811426", "exception": false, "start_time": "2022-05-10T12:34:21.754806", "status": "completed"} tags=[]
train.head()
# + papermill={"duration": 0.286221, "end_time": "2022-05-10T12:34:22.115792", "exception": false, "start_time": "2022-05-10T12:34:21.829571", "status": "completed"} tags=[]
#Missing values
train.isna().sum().sum(), test.isna().sum().sum()
# + papermill={"duration": 0.039719, "end_time": "2022-05-10T12:34:22.173558", "exception": false, "start_time": "2022-05-10T12:34:22.133839", "status": "completed"} tags=[]
#Target class balance
train.target.value_counts(normalize=True)
# + papermill={"duration": 0.197584, "end_time": "2022-05-10T12:34:22.389622", "exception": false, "start_time": "2022-05-10T12:34:22.192038", "status": "completed"} tags=[]
train.info()
# + papermill={"duration": 0.071201, "end_time": "2022-05-10T12:34:22.480908", "exception": false, "start_time": "2022-05-10T12:34:22.409707", "status": "completed"} tags=[]
features = [f for f in train.columns if f not in ('id', 'target')]
num_features = train.select_dtypes('float64').columns.to_list()
# + papermill={"duration": 0.030023, "end_time": "2022-05-10T12:34:22.530297", "exception": false, "start_time": "2022-05-10T12:34:22.500274", "status": "completed"} tags=[]
def split_and_encode(df, feature='f_27'):
'''
Handles feature 'f_27' by splitting the string,
creating new columns for each character and then
encoding them with integers. Also creates a column
for count of unique characters in feature 'f_27'.
Each string in 'f_27' has exactly 10 uppercase
characters for both train and test sets.
'''
expanded_df = pd.DataFrame(
df[feature].apply(list).to_list(),
columns=[(feature + '_' + str(i)) for i in range(10)])
for col in expanded_df.columns:
expanded_df[col] = expanded_df[col].apply(lambda x: ord(x) - ord('A'))
expanded_df[feature + '_unique'] = df[feature].apply(set).apply(len)
return expanded_df
# + papermill={"duration": 19.820167, "end_time": "2022-05-10T12:34:42.36929", "exception": false, "start_time": "2022-05-10T12:34:22.549123", "status": "completed"} tags=[]
train = pd.concat([train, split_and_encode(train, 'f_27')], axis=1)
test = pd.concat([test, split_and_encode(test, 'f_27')], axis=1)
# + papermill={"duration": 0.606649, "end_time": "2022-05-10T12:34:42.994822", "exception": false, "start_time": "2022-05-10T12:34:42.388173", "status": "completed"} tags=[]
train.drop(['f_27'], axis=1, inplace=True)
test.drop(['f_27'], axis=1, inplace=True)
# + papermill={"duration": 0.026598, "end_time": "2022-05-10T12:34:43.041362", "exception": false, "start_time": "2022-05-10T12:34:43.014764", "status": "completed"} tags=[]
#updating features list with created columns
features = [f for f in train.columns if f not in ('id', 'target')]
num_features.append('f_27_unique')
cat_features = [f for f in features if f not in num_features]
# + papermill={"duration": 1.927654, "end_time": "2022-05-10T12:34:44.988565", "exception": false, "start_time": "2022-05-10T12:34:43.060911", "status": "completed"} tags=[]
train[cat_features] = train[cat_features].astype('category')
test[cat_features] = test[cat_features].astype('category')
# + papermill={"duration": 0.305566, "end_time": "2022-05-10T12:34:45.313832", "exception": false, "start_time": "2022-05-10T12:34:45.008266", "status": "completed"} tags=[]
N_SPLITS = 5
skf = StratifiedKFold(n_splits=N_SPLITS, shuffle=True, random_state=SEED)
train['fold'] = -1
for fold, (_, idx) in enumerate(skf.split(X=train, y=train['target'])):
train.loc[idx, 'fold'] = fold
# + papermill={"duration": 0.056973, "end_time": "2022-05-10T12:34:45.389795", "exception": false, "start_time": "2022-05-10T12:34:45.332822", "status": "completed"} tags=[]
train.head()
# + papermill={"duration": 0.057274, "end_time": "2022-05-10T12:34:45.466716", "exception": false, "start_time": "2022-05-10T12:34:45.409442", "status": "completed"} tags=[]
test.head()
# + papermill={"duration": 61.716209, "end_time": "2022-05-10T12:35:47.203357", "exception": false, "start_time": "2022-05-10T12:34:45.487148", "status": "completed"} tags=[]
train.to_csv('train_processed.csv', index=False)
test.to_csv('test_processed.csv', index=False)
| tps-may22/01-data-preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-1:081325390199:image/datascience-1.0
# ---
# ## Regression with Amazon SageMaker Linear Learner algorithm for Taxi ride fare prediction
# _**Single machine training for regression with Amazon SageMaker Linear Learner algorithm**_
# ## Introduction
#
# This notebook demonstrates the use of Amazon SageMaker’s implementation of the Linear Learner algorithm to train and host a regression model to predict taxi fare. This notebook uses the [New York City Taxi and Limousine Commission (TLC) Trip Record Data] (https://registry.opendata.aws/nyc-tlc-trip-records-pds/#) to train the model. We are not using the whole dataset from above but a small subset of the dataset to train our model here. You will download this subset of data in below steps.
#
#
# ---
# ## Setup
#
#
# This notebook was tested in Amazon SageMaker Studio on a ml.t3.medium instance with Python 3 (Data Science) kernel.
#
# Let's start by specifying:
# 1. The S3 buckets and prefixes that you want to use for training data and model data. This should be within the same region as the Notebook Instance, training, and hosting ( us-east-1).
# 1. The IAM role arn used to give training and hosting access to your data. See the documentation for how to create these. Note, if more than one role is required for notebook instances, training, and/or hosting, please replace the boto regexp with a the appropriate full IAM role arn string(s).
# cell 01
# !pip install numpy==1.19.5
# !pip install pandas==0.25.3
# cell 02
import os
import boto3
import re
import sagemaker
import numpy as np
# +
# cell 03
role = sagemaker.get_execution_role()
sess = sagemaker.Session()
region = boto3.Session().region_name
# S3 bucket for training data.
# this will create bucket like 'Sagemaker-<region>-<Your AccountId>'
data_bucket=sess.default_bucket()
data_prefix = "1p-notebooks-datasets/taxi/text-csv"
# S3 bucket for saving code and model artifacts.
output_bucket = data_bucket
output_prefix = "sagemaker/DEMO-linear-learner-taxifare-regression"
# -
# ### Before running the below cell make sure that you uploaded the nyc-taxi.csv file in Sagemaker Studio, provided to you, in the same folder where this Studio notebook is residing.
#
#
# +
# cell 04
import boto3
FILE_TRAIN = "nyc-taxi.csv"
# s3 = boto3.client("s3")
# s3.download_file(data_bucket, f"{FILE_TRAIN}", FILE_TRAIN)
import pandas as pd # Read in csv and store in a pandas dataframe
# df = pd.read_csv(FILE_TRAIN, sep=",", encoding="latin1")
df = pd.read_csv(FILE_TRAIN, sep=",", encoding="latin1", names=["fare_amount","vendor_id","pickup_datetime","dropoff_datetime","passenger_count","trip_distance","pickup_longitude","pickup_latitude","rate_code","store_and_fwd_flag","dropoff_longitude","dropoff_latitude","payment_type","surcharge","mta_tax","tip_amount","tolls_amount","total_amount"])
print(df.head(5))
# -
# cell 05
df.info()
# #### We have 18 features "fare_amount", "vendor_id", "pickup_datetime", "dropoff_datetime", "passenger_count", "trip_distance", "pickup_longitude", "pickup_latitude", "rate_code", "store_and_fwd_flag", "dropoff_longitude", "dropoff_latitude", "payment_type", "surcharge", "mta_tax", "tip_amount", "tolls_amount", "total_amount" in the dataset
#
# Lets explore the dataset
# +
# cell 06
# Frequency tables for each categorical feature
for column in df.select_dtypes(include=['object']).columns:
display(pd.crosstab(index=df[column], columns='% observations', normalize='columns'))
# Histograms for each numeric features
display(df.describe())
# %matplotlib inline
hist = df.hist(bins=30, sharey=True, figsize=(10, 10))
# -
# #### As we can see that store_and_fwd_flg column doesn't have much variance in it ( as 98% of the column values are N and 2% are Y) hence this column won't have much impact on target variable ( fare_amount ). Also from our domain knowledge we can see that payment_type column value doesn't impact on trip fare hence we can drop both of these features from dataset
# cell 07
df = df.drop(['payment_type', 'store_and_fwd_flag'], axis=1)
df.info()
# #### we can see that in the dataset there are 2 features 'pickup_datetime' and 'dropoff_datetime' which depict when ride started and when did it end. As we know that taxi fare is highly dependent on duration of the drive hence as part of feature engineering we will create a feature which will calculate ride duration using these features
# cell 08
df['dropoff_datetime']= pd.to_datetime(df['dropoff_datetime'])
df['pickup_datetime']= pd.to_datetime(df['pickup_datetime'])
df['journey_time'] = (df['dropoff_datetime'] - df['pickup_datetime'])
df['journey_time'] = df['journey_time'].dt.total_seconds()
df['journey_time']
# #### after creation of 'journey_time feature' we can drop 'pickup_datetime' and 'dropoff_datetime' features
# cell 09
df = df.drop(['dropoff_datetime', 'pickup_datetime'], axis=1)
df.info()
# #### As you can see that vedor_id is still a categorical feature and we need to chage it to float ( using dummuies 0) so that dataset can be passed to Liner learner algorithm
# cell 10
df = pd.get_dummies(df, dtype=float)
df.info()
# #### Split the dataframe in train, test and validation
# +
# cell 11
import numpy as np
train_data, validation_data, test_data = np.split(df.sample(frac=1, random_state=1729), [int(0.7 * len(df)), int(0.9 * len(df))])
train_data.to_csv('train.csv', header=False, index=False)
validation_data.to_csv('validation.csv', header=False, index=False)
test_data.to_csv('test.csv', header=False, index=False)
# -
# cell 12
boto3.Session().resource('s3').Bucket(data_bucket).Object(os.path.join(data_prefix, 'train/train.csv')).upload_file('train.csv')
boto3.Session().resource('s3').Bucket(data_bucket).Object(os.path.join(data_prefix, 'validation/validation.csv')).upload_file('validation.csv')
boto3.Session().resource('s3').Bucket(data_bucket).Object(os.path.join(data_prefix, 'test/test.csv')).upload_file('test.csv')
#
# ---
# Let us prepare the handshake between our data channels and the algorithm. To do this, we need to create the `sagemaker.session.s3_input` objects from our [data channels](https://sagemaker.readthedocs.io/en/v1.2.4/session.html#). These objects are then put in a simple dictionary, which the algorithm consumes. Notice that here we use a `content_type` as `text/csv` for the pre-processed file in the data_bucket. We use two channels here one for training and the second one for validation. The testing samples from above will be used on the prediction step.
# +
# cell 13
# creating the inputs for the fit() function with the training and validation location
s3_train_data = f"s3://{data_bucket}/{data_prefix}/train"
print(f"training files will be taken from: {s3_train_data}")
s3_validation_data = f"s3://{data_bucket}/{data_prefix}/validation"
print(f"validtion files will be taken from: {s3_validation_data}")
s3_test_data = f"s3://{data_bucket}/{data_prefix}/test"
print(f"test files will be taken from: {s3_test_data}")
output_location = f"s3://{output_bucket}/{output_prefix}/output"
print(f"training artifacts output location: {output_location}")
# generating the session.s3_input() format for fit() accepted by the sdk
train_data = sagemaker.inputs.TrainingInput(
s3_train_data,
distribution="FullyReplicated",
content_type="text/csv",
s3_data_type="S3Prefix",
record_wrapping=None,
compression=None,
)
validation_data = sagemaker.inputs.TrainingInput(
s3_validation_data,
distribution="FullyReplicated",
content_type="text/csv",
s3_data_type="S3Prefix",
record_wrapping=None,
compression=None,
)
# -
# ## Training the Linear Learner model
#
# First, we retrieve the image for the Linear Learner Algorithm according to the region.
#
# Then we create an [estimator from the SageMaker Python SDK](https://sagemaker.readthedocs.io/en/stable/api/training/estimators.html) using the Linear Learner container image and we setup the training parameters and hyperparameters configuration.
#
# +
# cell 14
# getting the linear learner image according to the region
from sagemaker.image_uris import retrieve
container = retrieve("linear-learner", boto3.Session().region_name, version="1")
print(container)
# +
# cell 15
# # %%time
import boto3
import sagemaker
from time import gmtime, strftime
sess = sagemaker.Session()
job_name = "DEMO-linear-learner-taxifare-regression-" + strftime("%H-%M-%S", gmtime())
print("Training job", job_name)
linear = sagemaker.estimator.Estimator(
container,
role,
input_mode="File",
instance_count=1,
instance_type="ml.m4.xlarge",
output_path=output_location,
sagemaker_session=sess,
)
linear.set_hyperparameters(
epochs=16,
wd=0.01,
loss="absolute_loss",
predictor_type="regressor",
normalize_data=True,
optimizer="adam",
mini_batch_size=1000,
lr_scheduler_step=100,
lr_scheduler_factor=0.99,
lr_scheduler_minimum_lr=0.0001,
learning_rate=0.1,
)
# -
# ---
# After configuring the Estimator object and setting the hyperparameters for this object. The only remaining thing to do is to train the algorithm. The following cell will train the algorithm. Training the algorithm involves a few steps. Firstly, the instances that we requested while creating the Estimator classes are provisioned and are setup with the appropriate libraries. Then, the data from our channels are downloaded into the instance. Once this is done, the training job begins. The provisioning and data downloading will take time, depending on the size of the data. Therefore it might be a few minutes before we start getting data logs for our training jobs. The data logs will also print out Mean Average Precision (mAP) on the validation data, among other losses, for every run of the dataset once or one epoch. This metric is a proxy for the quality of the algorithm.
#
# Once the job has finished a "Job complete" message will be printed. The trained model can be found in the S3 bucket that was setup as output_path in the estimator.
# cell 16
# # %%time
linear.fit(inputs={"train": train_data, "validation": validation_data}, job_name=job_name)
# ## Set up hosting for the model
#
# Once the training is done, we can deploy the trained model as an Amazon SageMaker real-time hosted endpoint. This will allow us to make predictions (or inference) from the model. Note that we don't have to host on the same instance (or type of instance) that we used to train. Training is a prolonged and compute heavy job that require a different of compute and memory requirements that hosting typically do not. We can choose any type of instance we want to host the model. In our case we chose the ml.m4.xlarge instance to train, but we choose to host the model on the less expensive cpu instance, ml.c4.xlarge. The endpoint deployment can be accomplished as follows:
# cell 17
# # %%time
# creating the endpoint out of the trained model
linear_predictor = linear.deploy(initial_instance_count=1, instance_type="ml.c4.xlarge")
print(f"\ncreated endpoint: {linear_predictor.endpoint_name}")
# #### Copy the endpoint name of the deployed model from above and save it for later
# ## Inference
#
# Now that the trained model is deployed at an endpoint that is up-and-running, we can use this endpoint for inference. To do this, we are going to configure the [predictor object](https://sagemaker.readthedocs.io/en/v1.2.4/predictors.html) to parse contents of type text/csv and deserialize the reply received from the endpoint to json format.
#
# +
# cell 18
# configure the predictor to accept to serialize csv input and parse the reposne as json
from sagemaker.serializers import CSVSerializer
from sagemaker.deserializers import JSONDeserializer
linear_predictor.serializer = CSVSerializer()
linear_predictor.deserializer = JSONDeserializer()
# -
# ---
# We then use the test file containing the records of the data that we kept to test the model prediction. By running below cell multiple times we are selecting random sample from the testing samples to perform inference with.
# +
# cell 19
# # %%time
import json
from itertools import islice
import math
import struct
import boto3
import random
# downloading the test file from data_bucket
FILE_TEST = "test.csv"
s3 = boto3.client("s3")
s3.download_file(data_bucket, f"{data_prefix}/test/{FILE_TEST}", FILE_TEST)
# getting testing sample from our test file
test_data = [l for l in open(FILE_TEST, "r")]
sample = random.choice(test_data).split(",")
actual_fare = sample[0]
payload = sample[1:] # removing target variable from the sample
payload = ",".join(map(str, payload))
print('payload: ', payload, type(payload))
# Invoke the predicor and analyise the result
result = linear_predictor.predict(payload)
print('Result:', result)
# extracting the prediction value
result = round(float(result["predictions"][0]["score"]), 2)
accuracy = str(round(100 - ((abs(float(result) - float(actual_fare)) / float(actual_fare)) * 100), 2))
print(f"Actual fare: {actual_fare}\nPrediction: {result}\nAccuracy: {accuracy}")
# -
# cell 20
| lab7/TaxiFare_Predict_Liner_Learner.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/EOKELLO/IP-WEEK1-CORE/blob/master/2Moringa_Data_Science_Core_W1_Independent_Project_2019_07_Elizabeth_Okello_Python_Notebook.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="BRU2vCZajm3c" colab_type="code" colab={}
#import important libraries
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import sklearn
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from scipy import stats
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier, export_graphviz
from sklearn import metrics
# + [markdown] id="Q1-zb98pCw2w" colab_type="text"
# **Define** **the** **question**
# + id="q0Z_nyLHC5bP" colab_type="code" colab={}
#here we define our research question which is
#To figure out how we can predict which individuals are most likely to have or use a bank account
# + [markdown] id="tHQv4qTzDc4A" colab_type="text"
# **The metric for success**
# + id="hJVvI6KjDkl8" colab_type="code" colab={}
#Perform univariate analysis using visualization
#perform bivariate analysis using visualization
#mutivariate analysis using multiple regression and reduction techniques in this case PCA
#Make reccommendation and comments on the findings
# + [markdown] id="CYh4FAVkEqXX" colab_type="text"
# **The context**
# + id="nT5S6qFtEykP" colab_type="code" colab={}
#The countries in this study include uganda,kenya,rwanda,tanzania
#the refences are listed below
#FinAccess Kenya 2018.
#Finscope Rwanda 2016
#Finscope Tanzania 2017
#Finscope Uganda 2018
#Dataset: http://bit.ly/FinancialDataset
#Variable Definitions: http://bit.ly/VariableDefinitions
# + [markdown] id="_pirBF1KFczl" colab_type="text"
# **Experimental design**
# + id="SM2bffcFFi7x" colab_type="code" colab={}
#load the dataset
#clean the dataset by dealing with outliers, anomalies, and missing data within the dataset if any
#Plot univariate and bivariate summaries recording your observations
#Implement the solution by performing the respective analysis i.e. reduction, modeling
#Challenge your solution by providing insights on how you can make improvements
# + [markdown] id="ePYkwOWrGy7-" colab_type="text"
# **The appropriateness of the available data to answer the given question**
# + id="ju7UhHRJHVaf" colab_type="code" colab={}
#How is the dataset in regard to our research question
#Is the Data sufficient
#is it relevant
#is the dataset reliable
#what about the external sources are they helpfull?
# + id="66ydDVSMj9Xj" colab_type="code" outputId="8d6ac760-54d0-46ce-c400-02d0ae02d510" colab={"base_uri": "https://localhost:8080/", "height": 377}
#we now load our dataset
url='http://bit.ly/FinancialDataset'
df = pd.read_csv(url)
df.head()
# + id="1HdFcOlhkI3Y" colab_type="code" colab={}
#we now remane out dataset columns to make the column name uniform with the informaton captured in the data description
df = df.rename(columns={'The relathip with head':'relationship_with_head',
'Type of Location':'location_type',
'Cell Phone Access':'cellphone_access',
'Respondent Age':'age_of_respondent',
'Level of Educuation':'education_level',
'Type of Job':'job_type'})
# + id="Yo3Bgf8vrbkD" colab_type="code" outputId="363f7f5c-abdb-455b-c43d-6bea708f484d" colab={"base_uri": "https://localhost:8080/", "height": 360}
df.head()
# + id="D-0VAKa0rpHK" colab_type="code" outputId="f0389d52-ba08-4236-e656-cb067e54e92e" colab={"base_uri": "https://localhost:8080/", "height": 34}
#we want to understand our dataset by checking the number of records,missing values and data types
df.shape#here we are checking the size of our dataset is the volume sufficient or its very little data to enable us gain useful insights?
# + id="SCgxdVZBr2si" colab_type="code" outputId="7e08d585-e4cc-4216-ef9c-858e8512a909" colab={"base_uri": "https://localhost:8080/", "height": 255}
#the correct data format is key and we check if our dataset mets this requirement by
df.dtypes
# + id="ZueNp1B9r64y" colab_type="code" outputId="9f4cc104-5c0a-4ce7-d1ec-423e9f662c1c" colab={"base_uri": "https://localhost:8080/", "height": 255}
#we establish whether there are missing values using the code below
df.isnull().sum()
# + id="YqX4a_DisASf" colab_type="code" outputId="392a1f10-36a7-4952-b48a-2a4ab1b72919" colab={"base_uri": "https://localhost:8080/", "height": 1000}
#we the proceed to check our missing values by column and what quantity is missing for each variables
missing_data=df.isnull()# assign all mising values to a name missing_data
for column in missing_data.columns.values.tolist():#we use the for loops to avoid repitition of code for each variable
print(column)
print (missing_data[column].value_counts())
print("")
# + id="q3JMDGTuwHUd" colab_type="code" colab={}
#we now fill in missing values with the respective modes of each attribute in the columns enlisted
categorical_columns = ['country','gender_of_respondent', 'relationship_with_head', 'marital_status','location_type','cellphone_access','education_level','job_type']
for column in categorical_columns:# we do this to avoid repiting the same procedure for each column.
df[column] = df[column].fillna(df[column].mode().iloc[0])
# + id="cHOM7EvgzUvj" colab_type="code" colab={}
#we replace NaN value with the most common house_hold size
df['household_size'] = df['household_size'].fillna(df['household_size'].mode().iloc[0])
# + id="vCQ7Q_x-0lqS" colab_type="code" colab={}
#find the mode in the column age_of_respondent and use it to replace the nan values.
df['age_of_respondent'] = df['age_of_respondent'].fillna(df['age_of_respondent'].mode().iloc[0])
# + id="b9Hnxf18zpKU" colab_type="code" colab={}
#there is an anomaly in the education level
df['education_level'].unique()
df['education_level']=df['education_level']
# + id="y7OY3GmSyHRm" colab_type="code" colab={}
#we replace NaN value with the most frequent value for Has a Bank account
df['Has a Bank account'] = df['Has a Bank account'].fillna(df['Has a Bank account'].mode().iloc[0])
# + id="vVjwywbF2xTV" colab_type="code" outputId="de46f356-ffbf-4e6a-d60c-50228f3f76a4" colab={"base_uri": "https://localhost:8080/", "height": 255}
#check whether we successfully treated the missing values
df.isnull().sum()
# + id="6roCLcsmTtuN" colab_type="code" outputId="a974e939-a2bc-4a6d-baa4-f87548b22716" colab={"base_uri": "https://localhost:8080/", "height": 255}
#we check whether our variable data types are as expected
df.dtypes
# + id="d6VaqoPlUGXl" colab_type="code" colab={}
#all our ariable data types are fine except for house hold size so to correct this we do the conversion
df['household_size']=df['household_size'].astype("float")
# + id="OHBIdv-MUtdT" colab_type="code" outputId="eb1f437f-02d5-41dd-ab98-f8b2748a091d" colab={"base_uri": "https://localhost:8080/", "height": 255}
#we check if this is converted
df.dtypes
# + id="MHJoBWn7U1Pz" colab_type="code" colab={}
#we now have all missing values treated, variable data types in the correct format
# + id="NcNN8RDBcnQj" colab_type="code" colab={}
#we delete the columns we may not need like the uniqueid,year
df=df.drop(['uniqueid','year'],axis=1)
# + id="ipoNKgTgcnMy" colab_type="code" outputId="44d27294-564c-4b9a-fb60-a1931345a6c1" colab={"base_uri": "https://localhost:8080/", "height": 199}
df.head(2)
# + [markdown] id="eTcI8fbesErA" colab_type="text"
# **UNIVARIATE** **ANALYSIS**
# + id="fRK0ZR--yRTR" colab_type="code" colab={}
#we perform a univariate analysis which majorly highlghts the distribution of data variables.
# + id="ShqDQI1BoheM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 286} outputId="a6b70959-dff6-4d3b-d884-4aa89b1093bd"
df['age_of_respondent'].hist()
#in the below histogram we observe some positive skewness.
#we can say that those aged between 20 and 50 account for most of the respondents by age.
# + id="mHNRPLoiopfN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="5d9ab33f-bc45-4425-bc6c-7c04c71a188c"
#getting the descriptive statistics
df['age_of_respondent'].describe()
# + id="F4wdHHv3q3KS" colab_type="code" colab={}
#from the above statistics we can infer that since the mean and the median(50%quartile) are slightly far from each other there could be
#a presence of relatively many outliers
# + id="xhgZux25taXT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 301} outputId="0b5a0554-e7cc-404f-e871-8f20d876288c"
#we visualize to check if truely there are outliers using the box plot and yes there are outliers!as in the box plot below.
sns.boxplot(df['age_of_respondent'],showmeans=True)
# + id="Bcl_YrC1f6HX" colab_type="code" outputId="52af503d-ed17-4904-d844-ee9b0323ed1d" colab={"base_uri": "https://localhost:8080/", "height": 170}
#we proceed to perform a univariate analysis of the household_size
df['household_size'].describe()
# + id="dQnXK8epo5Ow" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 286} outputId="5f27bb76-b809-4b6c-bfbc-1272bed42ec1"
df['household_size'].hist()
#in the histogram below we observe positive skewness
#respondents of household_size 5 and below account for most of the respondents going by household size
# + id="w0ip3azoS3C7" colab_type="code" outputId="49cc8c89-63fc-47e1-9e0c-28318a42ac59" colab={"base_uri": "https://localhost:8080/", "height": 301}
#we see that the diference between the mean and the median is small indicating the presence of outliers thouhg not too many
#this comes out clearly given the visualization below
sns.boxplot(df['household_size'],showmeans=True)
# + id="bLZDQ-wMWSCd" colab_type="code" outputId="0c6859f6-6465-4ce6-bfcd-748f42d9e723" colab={"base_uri": "https://localhost:8080/", "height": 51}
#we check for distinct values in the column
df['household_size'].unique()
# + id="bcB2ogv4WR3n" colab_type="code" outputId="6f09d6e5-4653-4fda-9c15-2edd25266cc2" colab={"base_uri": "https://localhost:8080/", "height": 296}
#we still see that the trend holds ie household_size of less than 5 account for most of the respondents in the survey.
household_size = df['household_size']
plt.hist(household_size, bins=5, histtype='bar', rwidth=0.9, color='maroon')
plt.xlabel('household_size')
plt.ylabel('frequency')
plt.title('Histogram of household_size')
plt.show()
# + id="tbFNnASv8Ksy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 486} outputId="a644ed73-21a9-4992-a7f9-7fa7d94dc8cd"
#we now use a bargraph since the attributes are categorical in nature.
#from the below plot we see that in regard to gender majority of the respondents were women at a percentage of 59.06%.
#we could attribute this to the fact that since it was a household survey more females were found at home compared to males
#not necessarily that it could point to more females holding accounts than men.but it could be an interesting variable to investigate
#but this helps us to understand that of the respondents by gender women accounted for 59.06%
ax = df['gender_of_respondent'].value_counts().plot(kind='bar', figsize=(10,7),
color="indigo", fontsize=13);
ax.set_alpha(0.8)
ax.set_title("gender of respondents", fontsize=18)
ax.set_ylabel("frequency", fontsize=18);
ax.set_yticks([0, 5, 10, 15, 20,25, 30, 35, 40])
# create a list to collect the plt.patches data
totals = []
# find the values and append to list
for i in ax.patches:
totals.append(i.get_height())
# set individual bar lables using above list
total = sum(totals)
# set individual bar lables using above list
for i in ax.patches:
# get_x pulls left or right; get_height pushes up or down
ax.text(i.get_x()-.03, i.get_height()+.5, \
str(round((i.get_height()/total)*100, 2))+'%', fontsize=15,
color='dimgrey')
# + id="m6Bg8tLe9oxj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 478} outputId="b48dbae8-4424-4e0b-bc25-dc9ba1ee6a6e"
#from the below grapg we see that most o the respondents were drawn from rural areas and account for 61% of the respodents by
#location.typically banks are widespread across the countries and location of an individual maynot necessarily be a strong pointer to
#an individual having an account or not. may thereore not necessarily include it in my model but it paints a picture of the distribution
#of respondents by location type
ax = df['location_type'].value_counts().plot(kind='bar', figsize=(10,7),
color="green", fontsize=13);
ax.set_alpha(0.8)
ax.set_title("location_type", fontsize=18)
ax.set_ylabel("frequency", fontsize=18);
ax.set_yticks([0, 5, 10, 15, 20])
# create a list to collect the plt.patches data
totals = []
# find the values and append to list
for i in ax.patches:
totals.append(i.get_height())
# set individual bar lables using above list
total = sum(totals)
# set individual bar lables using above list
for i in ax.patches:
# get_x pulls left or right; get_height pushes up or down
ax.text(i.get_x()-.03, i.get_height()+.5, \
str(round((i.get_height()/total)*100, 2))+'%', fontsize=15,
color='dimgrey')
# + id="D8N_RkCB-Abp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 461} outputId="c9a40159-1078-437e-faf8-9c4abff544b6"
#fom the plots below 74.22% of the respondents have access to cell phones.
#with the development of ict this looks an interesting variable to include in my model
#growth of mobile money across the countries included in the study add strength to this variable as an interesting one to
#consider in the model.
ax = df['cellphone_access'].value_counts().plot(kind='bar', figsize=(10,7),
color="green", fontsize=13);
ax.set_alpha(0.8)
ax.set_title("cellphone_access", fontsize=18)
ax.set_ylabel("frequency", fontsize=18);
ax.set_yticks([0, 5, 10, 15, 20])
# create a list to collect the plt.patches data
totals = []
# find the values and append to list
for i in ax.patches:
totals.append(i.get_height())
# set individual bar lables using above list
total = sum(totals)
# set individual bar lables using above list
for i in ax.patches:
# get_x pulls left or right; get_height pushes up or down
ax.text(i.get_x()-.03, i.get_height()+.5, \
str(round((i.get_height()/total)*100, 2))+'%', fontsize=15,
color='dimgrey')
# + id="l97BOKt6-AZd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 592} outputId="3d8740a3-f02a-480f-bd9b-848b9e6c3add"
#from the plots those married accounted for 45.79% of the respondents by marital status.
#other groups in this category have also sufficiently contributed to this survey
#its an interesting factoe and on the groung it clearly is a contributing factor to having or not having an account
#
ax = df['marital_status'].value_counts().plot(kind='bar', figsize=(10,7),
color="blue", fontsize=13);
ax.set_alpha(0.8)
ax.set_title("marital status", fontsize=18)
ax.set_ylabel("frequency", fontsize=18);
ax.set_yticks([0, 5, 10, 15, 20,25, 30, 35, 40])
# create a list to collect the plt.patches data
totals = []
# find the values and append to list
for i in ax.patches:
totals.append(i.get_height())
# set individual bar lables using above list
total = sum(totals)
# set individual bar lables using above list
for i in ax.patches:
# get_x pulls left or right; get_height pushes up or down
ax.text(i.get_x()-.03, i.get_height()+.5, \
str(round((i.get_height()/total)*100, 2))+'%', fontsize=15,
color='dimgrey')
# + id="qACivHLt-1P5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 650} outputId="ad8def92-248a-4b27-e8d4-3272b03fb38b"
#from the plotted graphs we see that the respondents according to job_type self employed,informally employed and fish farming
#accounted for the greatest percentage. in this survey it looks an interesting variable to investigate
ax = df['job_type'].value_counts().plot(kind='bar', figsize=(10,7),
color="gold", fontsize=13);
ax.set_alpha(0.8)
ax.set_title("job_type", fontsize=18)
ax.set_ylabel("frequency", fontsize=18);
ax.set_yticks([0, 5, 10, 15, 20,25, 30, 35, 40])
# create a list to collect the plt.patches data
totals = []
# find the values and append to list
for i in ax.patches:
totals.append(i.get_height())
# set individual bar lables using above list
total = sum(totals)
# set individual bar lables using above list
for i in ax.patches:
# get_x pulls left or right; get_height pushes up or down
ax.text(i.get_x()-.03, i.get_height()+.5, \
str(round((i.get_height()/total)*100, 2))+'%', fontsize=15,
color='dimgrey')
# + id="N8zYcXXu_IfV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 566} outputId="42afa0b9-8cbf-4668-9dcf-1d9a5cde6a7a"
#heads of households accounted for the biggest percentage of respondents in this category.
#from te survey of the three countries this variable does not appear key to pointing whether one can have an account or not.
#rom the plots however we are able to understand the distribution of the respondents in this category with non relatives forming the smallest
#percentage of respondents.
ax = df['relationship_with_head'].value_counts().plot(kind='bar', figsize=(10,7),
color="brown", fontsize=13);
ax.set_alpha(0.8)
ax.set_title("relationship with head", fontsize=18)
ax.set_ylabel("frequency", fontsize=18);
ax.set_yticks([0, 5, 10, 15, 20,25, 30, 35, 40])
# create a list to collect the plt.patches data
totals = []
# find the values and append to list
for i in ax.patches:
totals.append(i.get_height())
# set individual bar lables using above list
total = sum(totals)
# set individual bar lables using above list
for i in ax.patches:
# get_x pulls left or right; get_height pushes up or down
ax.text(i.get_x()-.03, i.get_height()+.5, \
str(round((i.get_height()/total)*100, 2))+'%', fontsize=15,
color='dimgrey')
# + id="mYvPLxlL_b2h" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 641} outputId="26aff970-9a86-433e-8a19-b3a344041743"
#this variable is highly correlated to an individual having an account or not and would obviously be factored in while creatint the model
#indiviiduals who have attained education upto primary level are the biggest portion of the respondents at 54.55%
ax = df['education_level'].value_counts().plot(kind='bar', figsize=(10,7),
color="maroon", fontsize=13);
ax.set_alpha(0.8)
ax.set_title("education_level", fontsize=18)#for the title of the graph
ax.set_ylabel("frequency", fontsize=18);#
ax.set_yticks([0, 5, 10, 15, 20,25, 30, 35, 40])
# create a list to collect the plt.patches data
totals = []
# find the values and append to list
for i in ax.patches:
totals.append(i.get_height())
# set individual bar lables using above list
total = sum(totals)
# set individual bar lables using above list
for i in ax.patches:
# get_x pulls left or right; get_height pushes up or down
ax.text(i.get_x()-.03, i.get_height()+.5, \
str(round((i.get_height()/total)*100, 2))+'%', fontsize=15,
color='dimgrey')
# + [markdown] id="30CjGGPLryRd" colab_type="text"
# BIVARIATE ANALYSIS
# + id="51bRHb8l5ehE" colab_type="code" colab={}
#we want to check for the distribution od variables and how they relate to one another . we shall also be establishing
#the strength of relationship if there exists
# + id="6qVWzkKljnD7" colab_type="code" outputId="814d3de4-3462-4187-fceb-b7e9837b9b43" colab={"base_uri": "https://localhost:8080/", "height": 403}
plt.figure(figsize=(8, 6))
sns.pairplot(df, diag_kind='kde')
# + id="p89y9JkF7mMo" colab_type="code" colab={}
#in the above plots we see no relationship between the two variables at all
# + id="OE_TctRFjnAp" colab_type="code" outputId="dd10d2a4-72a3-4e1d-9121-6055489973bf" colab={"base_uri": "https://localhost:8080/", "height": 306}
sns.heatmap(df.corr())
#the black shades confirm that there is completely no relationship between the two variables
# + [markdown] id="bRYO1H0DjDJh" colab_type="text"
# Distributions of observations within categories
# + id="QnOpJcyncNYY" colab_type="code" outputId="5a37c185-5665-42d1-e11c-17c5dae6b6b0" colab={"base_uri": "https://localhost:8080/", "height": 420}
#we can use this to visualize how our observations are distributed within the various categories.
#we can say that the ages of the respondents in the category of educ level those with no formal education were more and older compared to the rest
sns.boxplot(x='education_level',y='age_of_respondent',data=df,palette='rainbow')
plt.xticks(rotation = 45)
plt.title('education level vs age of respondent')
# + id="VIDfB-JSjm4_" colab_type="code" outputId="e28aa7b3-e2bb-475b-c63d-3da001c85ba9" colab={"base_uri": "https://localhost:8080/", "height": 424}
sns.boxplot(x='job_type',y='age_of_respondent',data=df,palette='coolwarm')
plt.xticks(rotation = 45)
plt.title('job_type vs age of respondent')
# + id="68xOPAJkf8ts" colab_type="code" outputId="9011cd1d-def2-4a89-adf3-0a8f7db8b989" colab={"base_uri": "https://localhost:8080/", "height": 336}
#the plot below suggests that the ages of the respondents whether male or female are around the same bracket
sns.boxplot(x='gender_of_respondent',y='age_of_respondent',data=df,palette='rainbow')
plt.xticks(rotation = 45)
plt.title('gender_of_respondent vs age of respondent')
# + id="cqoN5UCRgd2R" colab_type="code" outputId="e1b8a417-7d10-4a34-9d2d-c145193d33a2" colab={"base_uri": "https://localhost:8080/", "height": 336}
#the plot suggests that there was no big variance in the number of males and females with regard to the household size
sns.boxplot(x='gender_of_respondent',y='household_size',data=df,palette='rainbow')
plt.xticks(rotation = 45)
plt.title('gender_of_respondent vs household_size')
# + id="fFqpkQltg07A" colab_type="code" outputId="0fdbb528-880d-47d9-86da-86f9195c99ee" colab={"base_uri": "https://localhost:8080/", "height": 393}
#
sns.boxplot(x='marital_status',y='age_of_respondent',data=df,palette='rainbow')
plt.xticks(rotation = 45)
plt.title('marital_status vs age of respondent')
# + id="VIDSWYsNZopK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 550} outputId="2246698c-0e50-4392-a5cb-fd738a5b1453"
#this plot suggests that most of the respondents were self employed in this jobtype category
sns.catplot(x="job_type", y="household_size", data=df)
plt.xticks(rotation = 90)
plt.title('job_type vs household size')
# + id="tNRUGKJiiDoQ" colab_type="code" colab={}
# + [markdown] id="ERSYQzM30kAz" colab_type="text"
# MULTIVARIATE ANALYSIS
# + id="xrmkU06M-Xyx" colab_type="code" colab={}
#multiple regression
from sklearn.linear_model import LinearRegression
from sklearn import model_selection
from sklearn.metrics import r2_score
from sklearn.model_selection import train_test_split
import statsmodels.api as sm
from sklearn.metrics import accuracy_score
# + id="DTO39M8_0KXy" colab_type="code" colab={}
#From our dataset we to divide dataset
# into features and corresponding labels and then divide the resultant dataset into training and test sets.
X = df.iloc[:, 2:11]
y = df.iloc[:, 1]
# + id="F5GftEtt8q7-" colab_type="code" colab={}
#because the data is categorical we have to make them in aa form that we can run a regression model on it
X=pd.get_dummies(X)
# + id="5QpbB2wC9MAP" colab_type="code" colab={}
from sklearn.preprocessing import LabelEncoder
label_encoder = LabelEncoder()
y = label_encoder.fit_transform(y)
# + id="EBkn2Idq-pWb" colab_type="code" outputId="cd818959-08d3-4b4a-9f5e-066e52158f9f" colab={"base_uri": "https://localhost:8080/", "height": 258}
X.head()
# + id="x_AzkPYwiqkX" colab_type="code" outputId="19c7ea31-732b-4cf5-c58f-c2908b19cfe8" colab={"base_uri": "https://localhost:8080/", "height": 34}
X.shape
# + id="7eUGHpKu1B5I" colab_type="code" colab={}
#the following code divides data into training and test sets
#
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
# + id="cWxhf2NSbnXQ" colab_type="code" colab={}
# We now need to perform feature scaling. We execute the following code to do so:
# this is components are sensitive to scale of measurements so we need to standardize our dataset
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# + id="CU5_3XE-n_im" colab_type="code" outputId="26577205-f453-43be-9ea4-79188605dd91" colab={"base_uri": "https://localhost:8080/", "height": 1000}
#we want to check if there is a relationship between the variables and also the strength of the relationship
X.corr()
# + id="AHxvHUoloC8i" colab_type="code" outputId="69af210b-5ccc-4789-bff2-4314d474ea5c" colab={"base_uri": "https://localhost:8080/", "height": 493}
#the heat map below a majority of the variables display very weak relationship with each other.
#but we can try to use multiple regression and see its performance
sns.heatmap(X.corr())
# + id="C0RMADsZoJtm" colab_type="code" colab={}
#We then create and train the model by fitting the linear regression model on the training data.
lm = LinearRegression()
results = lm.fit(X_train,y_train)
# + id="I4UQ20ypofS2" colab_type="code" colab={}
#Grabbing predictions off the test set and see how well it did
predictions = lm.predict(X_test)
# + id="yOP9mQnFo29g" colab_type="code" outputId="b84660e6-8161-4f12-a4e7-6d62799da0c3" colab={"base_uri": "https://localhost:8080/", "height": 1000}
X2 = sm.add_constant(X.values)
model = sm.OLS(y, X2).fit()
print(model.summary())
# + id="xaA0QHxI182w" colab_type="code" colab={}
#this model is not appropriate for modelling this kind of data. having assumed that there exists a linear relationship the relationship is too weak and absent in most variables
#therefore yields a very poor model looking at the rsquared and adjusted -squared
# + [markdown] id="BxE1lynYnKps" colab_type="text"
# **Principal Component Analysis**
# + id="g8-C_Se3dC5U" colab_type="code" colab={}
#we can also try pca to do some reduction on out dataset
from sklearn.decomposition import PCA
pca = PCA(0.95)
X_train = pca.fit_transform(X_train)
X_test = pca.transform(X_test)
# + id="hbP4a9AWdPFb" colab_type="code" colab={}
# Using Principal Component we set the threshhold of 95%
# Let's first try to use principal component to reduce our dataset by picking the items with the largest variance, execute the following code:
#
from sklearn.decomposition import PCA
pca = PCA(0.95)
X_train = pca.fit_transform(X_train)
X_test = pca.transform(X_test)
# + id="8HSrb_9Sdfxz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} outputId="5c8ad1fa-56d8-4c55-f543-e78c42ee363a"
#Training and Making Predictions
# In this case we'll use random forest classification for making the predictions.
#
from sklearn.ensemble import RandomForestClassifier
classifier = RandomForestClassifier(max_depth=2, random_state=0)
classifier.fit(X_train, y_train)
# Predicting the Test set results
y_pred = classifier.predict(X_test)
# + id="alPiVph2drZo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="399db76e-1dea-4b04-90cc-856e8aff68a3"
#Performance Evaluation
#
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
cm = confusion_matrix(y_test, y_pred)
print(cm)
print(accuracy_score(y_test, y_pred))
# + id="NR0SsvSBdfts" colab_type="code" colab={}
#from the above methods we attained an accuracy of 86%
#much as our data was sufficient we cant give a conclusion because looking at the confusion matrix we cannot make any reasonable conclusion.
#this then leads us to try and use a different method in this case Logistic Regression as the results we expect are binary in nature
# + id="7Tke7ezP-oDu" colab_type="code" colab={}
# + [markdown] id="5zVvLU9L-pnQ" colab_type="text"
# **Challenging our model using Logistic regression**
# + id="EnNw4YMO1M_j" colab_type="code" colab={}
#we want to use logistic regression to check whether we can get better results.
#following the assumptions of logistic regression that are known to us namely
#Binary logistic regression requires the dependent variable to be binary.(Our dataset meets this condition)
#For a binary regression, the factor level 1 of the dependent variable should represent the desired outcome.(we already did the conversions to dummy variable)
#Only the meaningful variables should be included.this is done by th use of PCA
#The independent variables should be independent of each other. That is, the model should have little or no multicollinearity.
#this is evident in our correlation table above
#Logistic regression requires quite large sample sizes.(our data is sufficient as it has volume)
# + id="O_M1xiMI2KGw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 156} outputId="70f8892f-8162-4428-e7ad-3e6ad5ab96ce"
#we therefore proceed to import our logistic regression
#we then fit our logistic regression model
from sklearn.linear_model import LogisticRegression
from sklearn import metrics
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0)
logreg = LogisticRegression()
logreg.fit(X_train, y_train)
# + id="ohiVlNPD30Lz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="41fc782a-ce35-4de5-db11-f79492c4fd40"
#Predicting the test set results and calculating the accuracy
y_pred = logreg.predict(X_test)
print('Accuracy of logistic regression classifier on test set: {:.2f}'.format(logreg.score(X_test, y_test)))
# + id="cSHadFcc4WHU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="d85b765b-6c8f-4fce-bca1-257d9bb82f1f"
#we can import confusion matrix
from sklearn.metrics import confusion_matrix
confusion_matrix = confusion_matrix(y_test, y_pred)
print(confusion_matrix)
# + id="EbYNfyzJ4idz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="0cdf893a-1c55-4ba1-b261-38756439f2ed"
#we evaluate our model
from sklearn.metrics import classification_report
print(classification_report(y_test, y_pred))
# + id="puA7I8HY495p" colab_type="code" colab={}
#Interpretation: Of the entire test set, 86% of the respondents were likely to have accounts.
| 2Moringa_Data_Science_Core_W1_Independent_Project_2019_07_Elizabeth_Okello_Python_Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
def single_replication(wks, lmbda):
data = pd.DataFrame(columns= ['C', 'U', 'I', 'B', 'R'], data=np.zeros((wks, 5)))
data.loc[0, 'I'] = lmbda
#Use Poisson to reverse solve for random vars
data.loc[:, 'C'] = np.random.poisson(lmbda, size=(wks,1))
for i in range(1,wks):
#Note: U_n has been shifted to accommodate the for loop
data.loc[i, 'I'] = (data.loc[i-1, 'I'] + data.loc[i-1, 'B'] - data.loc[i, 'C'] - data.loc[i-1,'U']) * (data.loc[i-1, 'I'] + data.loc[i-1, 'B'] - data.loc[i, 'C'] - data.loc[i-1,'U'] >= 0)
data.loc[i, 'U'] = (data.loc[i, 'C'] - data.loc[i, 'I']) * (data.loc[i, 'C'] >= data.loc[i, 'I'])
data.loc[i, 'R'] = lmbda + data.loc[i,'U']
data.loc[i, 'B'] = data.loc[i-1, 'R']
return data
# + pycharm={"name": "#%%\n"}
#Aggregate data over 100 simulations
reps = 100
weeks = 52
lambda_value = 1046
avg_data = single_replication(weeks, lambda_value)
stderr_data = avg_data.pow(2)
for i in range(reps-1):
avg_data = avg_data.add(single_replication(weeks, lambda_value))
stderr_data = stderr_data.add(single_replication(weeks, lambda_value).pow(2))
#Get standard error of data by getting variance and square rooting it (i.e. std dev)
avg_data = avg_data/reps
stderr_data = ((1/(reps-1))*(stderr_data - avg_data.pow(2))).pow(0.5)
avg_data.head()
# + pycharm={"name": "#%%\n"}
sns.set()
#Fill lines with standard error levels
colors = ['r', 'b', 'g', 'k', 'm']
for idj, j in enumerate(['C', 'U', 'I', 'B', 'R']):
plt.fill_between(avg_data.index, avg_data.loc[:,j] - stderr_data.loc[:,j]/np.sqrt(reps), avg_data.loc[:,j] + stderr_data.loc[:,j]/np.sqrt(reps), color=colors[idj], alpha=0.2)
plt.plot(avg_data)
#Match the colours with the ones above
for index, line in enumerate(plt.gca().get_lines()):
line.set_color(colors[index])
#Format plots
plt.gca().legend(('C','U','I','B','R'))
plt.gca().set_title("Supply Chain of Beer Store")
plt.gca().set_xlabel("Week")
plt.gca().set_ylabel("Number of Beer Cases")
plt.savefig('bullwhipeffect_beerstore.png', dpi=800)
plt.show()
| mat231-bullwhipmodel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import requests
import json
import pandas as pd
service = "http://172.16.31.10:80"
# service = "http://localhost:80"
sites = ['AGLT2', 'MWT2']
dataset = 'AUG'
GB = 1024 * 1024 * 1024
TB = 1024 * GB
PB = 1024 * TB
# -
# ### Import data
# +
all_accesses = []
for si, site in enumerate(sites):
print('Loading:', site)
all_accesses.append(pd.read_hdf(site + '_' + dataset + '.h5', key=site, mode='r'))
all_accesses[si]['site'] = 'xc_' + site
# print(all_accesses[si].head())
print(all_accesses[si].filesize.count(), "files")
print(all_accesses[si].index.unique().shape[0], " unique files")
print(all_accesses[si].filesize.sum() / PB, "PB")
print(all_accesses[si].filesize.mean() / GB, "GB avg. file size")
print('----------------------------')
all_data = pd.concat(all_accesses).sort_values('transfer_start')
print('---------- merged data -----------')
print(all_data.shape[0], 'files\t\t', all_data.index.unique().shape[0], 'unique files' )
print(all_data.filesize.sum() / PB, "PB")
print(all_data.filesize.mean() / GB, "GB avg. file size")
# print(all_data.head(100))
# -
# ### running requests
# +
print('---------- start requests ----------')
acs = []
dac = []
accesses = [0, 0, 0, 0]
dataaccc = [0, 0, 0, 0]
count = 0
payload = []
with requests.Session() as session:
for index, row in all_data.iterrows():
if count > 300000:
break
fs = row['filesize']
payload.append({'filename': index, 'site': row['site'], 'filesize': fs, 'time': row['transfer_start']})
# print(payload)
try:
if count % 100 and count > 0:
r = session.post(service + '/simulate', json=payload)
if r.status_code != 200:
print(r)
accs = r.json()
for i, j in enumerate(accs['counts']):
accesses[i] += int(j)
dataaccc[i] += accs['sizes'][i]
payload = []
except requests.exceptions.RequestException as e:
print(e)
if not count % 5000 and count > 0:
# print(count, accesses, dataaccc)
acs.append(accesses.copy())
dac.append(dataaccc.copy())
pacce = []
pdata = []
for i in range(len(accesses)):
pacce.append(accesses[i] / sum(accesses))
pdata.append(dataaccc[i] / sum(dataaccc))
print(count, pacce, pdata)
count += 1
print('final: ', accesses, dataaccc)
accdf = pd.DataFrame(acs)
dacdf = pd.DataFrame(dac)
dacdf=dacdf/(1024*1024*1024*1024)
# -
# ### ploting results
# +
accdf.columns = ['level 1', 'level 2', 'level 3', 'origin']
dacdf.columns = ['level 1', 'level 2', 'level 3', 'origin']
fig, axs = plt.subplots(nrows=2, ncols=1, constrained_layout=True,figsize=(8,10))
# plt.subplot(211)
accdf.plot(ax=axs[0])
axs[0].set_ylabel('hits')
axs[0].set_xlabel('reqeusts [x1000]')
axs[0].legend()
dacdf.plot(ax=axs[1])
axs[1].set_ylabel('data delivered [TB]')
axs[1].set_xlabel('reqeusts [x1000]')
axs[1].legend()
plt.show()
fig.savefig('filling_up.png')
# -
# ### Network states
# +
res = requests.get(service + '/status')
status = json.loads(res.json())
# print(status)
tp=[]
for site in status:
# print(site[0])
# print(site[1])
tp.append([site[0],site[1]['requests_received'],site[1]['files_delivered'],site[1]['data_delivered']/(1024*1024*1024*1024)])
sites=pd.DataFrame(tp)
sites.columns=['xcache','requests','hits','data delivered']
sites = sites[sites.requests!=0]
sites.head(20)
# +
fig, ax = plt.subplots(constrained_layout=True,figsize=(8,8))
sites.plot(x="xcache", y=["requests", "hits", "data delivered"], kind="bar", ax=ax,secondary_y= 'data delivered')
fig.savefig('xcache_sites.png')
# -
| analytics/Stresser.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="EFyh_Kfx5jkw"
# **Chapter 15 – Processing Sequences Using RNNs and CNNs**
#
# **Chapter 16 – Natural Language Processing with RNNs and Attention**
# + [markdown] id="TQs1xzgo5jkz"
# _This notebook contains the sample from https://github.com/ageron/handson-ml2/ and https://github.com/fchollet/deep-learning-with-python-notebooks_
# + [markdown] id="qIV46eeo5jkz"
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/phonchi/nsysu-math604/blob/master/static_files/presentations/04_Recurrent Neural Networks.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# </td>
# <td>
# <a target="_blank" href="https://kaggle.com/kernels/welcome?src=https://github.com/phonchi/nsysu-math604/blob/master/static_files/presentations/04_Recurrent Neural Networks.ipynb"><img src="https://kaggle.com/static/images/open-in-kaggle.svg" /></a>
# </td>
# </table>
# + [markdown] id="PdY3OIJ15jk0"
# # Setup
# + [markdown] id="ykB4PPYw5jk0"
# First, let's import a few common modules, ensure MatplotLib plots figures inline and prepare a function to save the figures. We also check that Python 3.5 or later is installed (although Python 2.x may work, it is deprecated so we strongly recommend you use Python 3 instead), as well as Scikit-Learn ≥0.20 and TensorFlow ≥2.0.
# + id="0cmPr4J_5jk1"
# Python ≥3.5 is required
import sys
assert sys.version_info >= (3, 5)
# Is this notebook running on Colab or Kaggle?
IS_COLAB = "google.colab" in sys.modules
IS_KAGGLE = "kaggle_secrets" in sys.modules
# Scikit-Learn ≥0.20 is required
import sklearn
assert sklearn.__version__ >= "0.20"
# TensorFlow ≥2.0 is required
import tensorflow as tf
from tensorflow import keras
assert tf.__version__ >= "2.0"
if not tf.config.list_physical_devices('GPU'):
print("No GPU was detected. LSTMs and CNNs can be very slow without a GPU.")
if IS_COLAB:
print("Go to Runtime > Change runtime and select a GPU hardware accelerator.")
if IS_KAGGLE:
print("Go to Settings > Accelerator and select GPU.")
# Common imports
import numpy as np
import os
from pathlib import Path
# to make this notebook's output stable across runs
np.random.seed(42)
tf.random.set_seed(42)
# To plot pretty figures
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "rnn"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
# + [markdown] id="eAE6jniY5jk2"
# # Basic RNNs for forecasting times series
# + [markdown] id="PhK1Eyvw5jk3"
# ## Generate the Dataset
# + [markdown] id="ZNbWWTJm9_WN"
# Suppose you are studying the number of active users per hour on your website, or the daily temperature in your city, or your company’s financial health, measured quarterly using multiple metrics. In all these cases, the data will be a sequence of one or more values per time step. This is called a time series. In the first two examples there is a single value per time step, so these are **univariate time series**, while in the financial example there are multiple values per time step (e.g., the company’s revenue, debt, and so on), so it is a **multivariate time series**. A typical task is to predict future values, which is called forecasting. Another common task is to fill in the blanks: to predict (or rather “postdict”) missing values from the past. This is called imputation.
#
# For simplicity, we are using a time series generated by the generate_time_series() function, shown here:
# + id="4XG7SRs35jk3"
def generate_time_series(batch_size, n_steps):
freq1, freq2, offsets1, offsets2 = np.random.rand(4, batch_size, 1)
time = np.linspace(0, 1, n_steps)
series = 0.5 * np.sin((time - offsets1) * (freq1 * 10 + 10)) # wave 1
series += 0.2 * np.sin((time - offsets2) * (freq2 * 20 + 20)) # + wave 2
series += 0.1 * (np.random.rand(batch_size, n_steps) - 0.5) # + noise
return series[..., np.newaxis].astype(np.float32)
# + [markdown] id="3mT-BlwO-QMs"
# The function returns a NumPy array of shape `[batch size, time steps, 1]`, where each series is the sum of two sine waves of fixed amplitudes but random frequencies and phases, plus a bit of noise.
#
# When dealing with time series (and other types of sequences such as sentences), the input features are generally represented as 3D arrays of shape `[batch size, time steps, dimensionality]`,where dimensionality is 1 for univariate time series and more for multivariate
# time series.
#
# + id="jGFqAitJ5jk4"
np.random.seed(42)
n_steps = 50
series = generate_time_series(10000, n_steps + 1)
X_train, y_train = series[:7000, :n_steps], series[:7000, -1]
X_valid, y_valid = series[7000:9000, :n_steps], series[7000:9000, -1]
X_test, y_test = series[9000:, :n_steps], series[9000:, -1]
# + id="UXAjbgOt5jk4" outputId="5de60e8b-2b38-47ac-e541-a42fcd6ba328" colab={"base_uri": "https://localhost:8080/"}
X_train.shape, y_train.shape
# + [markdown] id="LXt8l_8F-2Ed"
# `X_train` contains 7,000 time series (i.e., its shape is `[7000, 50, 1]`), while `X_valid` contains 2,000 (from the 7,000th time series to the 8,999th) and X_test contains 1,000 (from the 9,000 to the 9,999 ). **Since we want to
# forecast a single value for each series**, the targets are column vectors (e.g., `y_train` has a shape of `[7000, 1]`).
# + id="_LQXcP-r5jk4" outputId="27d1aa45-0690-437f-b77a-ae48446f7b66" colab={"base_uri": "https://localhost:8080/", "height": 314}
def plot_series(series, y=None, y_pred=None, x_label="$t$", y_label="$x(t)$", legend=True):
plt.plot(series, ".-")
if y is not None:
plt.plot(n_steps, y, "bo", label="Target")
if y_pred is not None:
plt.plot(n_steps, y_pred, "rx", markersize=10, label="Prediction")
plt.grid(True)
if x_label:
plt.xlabel(x_label, fontsize=16)
if y_label:
plt.ylabel(y_label, fontsize=16, rotation=0)
plt.hlines(0, 0, 100, linewidth=1)
plt.axis([0, n_steps + 1, -1, 1])
if legend and (y or y_pred):
plt.legend(fontsize=14, loc="upper left")
fig, axes = plt.subplots(nrows=1, ncols=3, sharey=True, figsize=(12, 4))
for col in range(3):
plt.sca(axes[col])
plot_series(X_valid[col, :, 0], y_valid[col, 0],
y_label=("$x(t)$" if col==0 else None),
legend=(col == 0))
save_fig("time_series_plot")
plt.show()
# + [markdown] id="mVW_UPLp5jk5"
# ## Computing Some Baselines
# + [markdown] id="opK47Oz95jk5"
# Before we start using RNNs, it is often a good idea to have a few baseline metrics, or else we may end up thinking our model works great when in fact it is doing worse than basic models. The simplest approach is to predict the last value in each series. This is called *naive forecasting*
# + id="zQIzWhQQ5jk5" outputId="be199c4b-cff9-45d6-91bd-ba16b2191cd9" colab={"base_uri": "https://localhost:8080/"}
y_pred = X_valid[:, -1]
np.mean(keras.losses.mean_squared_error(y_valid, y_pred))
# + id="4y1kxSFw5jk6" outputId="0f0b9c0d-9df8-46ae-917e-4bb3610496cf" colab={"base_uri": "https://localhost:8080/", "height": 293}
plot_series(X_valid[0, :, 0], y_valid[0, 0], y_pred[0, 0])
plt.show()
# + [markdown] id="9Zi14gly5jk6"
# Another simple approach is to use a fully connected network. Since it expects a flat list of features for each input, we need to add a `Flatten layer`. Let’s just use a simple Linear Regression model so that each prediction will be a linear combination of the values in the time series:
# + id="Tr1yTxkj5jk6" outputId="7368b75f-6b0c-4019-9113-96586a7c23f2" colab={"base_uri": "https://localhost:8080/"}
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[50, 1]),
keras.layers.Dense(1)
])
model.compile(loss="mse", optimizer="adam")
history = model.fit(X_train, y_train, epochs=20,
validation_data=(X_valid, y_valid))
# + id="swohpJKX5jk6" outputId="df2883b3-43eb-432e-a2cf-5188ae484c6a" colab={"base_uri": "https://localhost:8080/"}
model.evaluate(X_valid, y_valid)
# + [markdown] id="nDRQiwp7_7Zd"
# If we compile this model using the MSE loss and the default Adam optimizer, then fit it on the training set for 20 epochs and evaluate it on the validation set, we get an MSE of about 0.004. That’s much better than the naive approach!
# + id="kckEzMwk5jk7" outputId="cbed5fa3-36cc-4f07-dc94-eb94a923594e" colab={"base_uri": "https://localhost:8080/", "height": 291}
def plot_learning_curves(loss, val_loss):
plt.plot(np.arange(len(loss)) + 0.5, loss, "b.-", label="Training loss")
plt.plot(np.arange(len(val_loss)) + 1, val_loss, "r.-", label="Validation loss")
plt.gca().xaxis.set_major_locator(mpl.ticker.MaxNLocator(integer=True))
plt.axis([1, 20, 0, 0.05])
plt.legend(fontsize=14)
plt.xlabel("Epochs")
plt.ylabel("Loss")
plt.grid(True)
plot_learning_curves(history.history["loss"], history.history["val_loss"])
plt.show()
# + id="HhHs0TEs5jk7" outputId="8ec7db41-b8ce-4486-8629-5cf8ff537d14" colab={"base_uri": "https://localhost:8080/", "height": 293}
y_pred = model.predict(X_valid)
plot_series(X_valid[0, :, 0], y_valid[0, 0], y_pred[0, 0])
plt.show()
# + [markdown] id="BaS5Yqbx5jk7"
# ## Using a Simple RNN
# + [markdown] id="4H7cLxnfAVEW"
# Let’s see if we can beat that with a simple RNN. It just contains a single layer, with a single neuron. We do not need to specify the length of the input sequences (unlike in the previous model), **since a recurrent neural network can process any number of time steps** (this is why we set the first input dimension to None). By default, the SimpleRNN layer **uses the hyperbolic tangent activation function**. It works exactly as we saw earlier: the initial state $h_{init}$ is set to 0, and it is passed to a single recurrent neuron, along with the value of the first time step, $x_0$. The neuron computes a weighted sum of these values and applies the hyperbolic tangent activation function to the result, and this gives the first output, $y_0$. In a simple RNN, this output is also the new state $h_0$. This new state is passed to the same recurrent neuron along with the next input value, $x_1$, and the process is repeated until the last time step. Then the layer just outputs the last value, $y_{49}$. All of this is performed simultaneously for every time series.
# + id="LlXfar6n5jk7"
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.SimpleRNN(1, input_shape=[None, 1])
])
# + [markdown] id="h11FMMnGCCeg"
# Note that for each neuron, a linear model has one parameter per input and per time step, plus a bias term (in the simple linear model we used, that’s a total of 51 parameters). In contrast, for each recurrent neuron in a simple
# RNN, **there is just one parameter per input and per hidden state dimension** (in a simple RNN, that’s just the number of recurrent neurons in the layer), plus a bias term.
# + colab={"base_uri": "https://localhost:8080/"} id="E082xqnTBttm" outputId="47389102-2276-41ab-c5fd-173c8a079662"
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="DFkksB7EBsjd" outputId="4e89dcd8-f38c-4e73-86eb-5bb18bf7ca58"
optimizer = keras.optimizers.Adam(learning_rate=0.005)
model.compile(loss="mse", optimizer=optimizer)
history = model.fit(X_train, y_train, epochs=20,
validation_data=(X_valid, y_valid))
# + id="xp6oMcGi5jk8" outputId="345df898-d649-4d5b-88f4-465dcea37f37" colab={"base_uri": "https://localhost:8080/"}
model.evaluate(X_valid, y_valid)
# + id="ArbK1I4Z5jk8" outputId="54db10c9-856c-4d19-f5d0-dcc062d70344" colab={"base_uri": "https://localhost:8080/", "height": 291}
plot_learning_curves(history.history["loss"], history.history["val_loss"])
plt.show()
# + id="-r6pZD_05jk8" outputId="4be1cf45-3e15-4119-e0dc-b52eb6601d20" colab={"base_uri": "https://localhost:8080/", "height": 293}
y_pred = model.predict(X_valid)
plot_series(X_valid[0, :, 0], y_valid[0, 0], y_pred[0, 0])
plt.show()
# + [markdown] id="AqHSj9gsDGCv"
# There are many other models to forecast time series, such as weighted moving average models or autoregressive integrated moving average (ARIMA) models. Some of them require you to first remove the trend and seasonality. Once the model is
# trained and starts making predictions, you would have to add them back. When using RNNs, it is generally not necessary to do all this, but it may improve performance in some cases, since the model will not have to learn the trend or the seasonality.
# + [markdown] id="xWFI7MMA5jk8"
# ## Deep RNNs
# + [markdown] id="5Za9vyD9DRtu"
# Implementing a deep RNN with `tf.keras` is quite simple: just stack recurrent layers. In this example, we use three SimpleRNN layers. Make sure to set `return_sequences=True` for all recurrent layers (except the last one, if you only care about the last output). **If you don’t, they will output a 2D array (containing only the output of the last time step)** instead of a 3D array (containing outputs for all time steps), and the next recurrent layer will complain that you are not feeding it sequences in the expected 3D format
# + id="S63hdy8B5jk8" outputId="7b254938-a536-4839-e0da-25876725fb38" colab={"base_uri": "https://localhost:8080/"}
np.random.seed(42)
tf.random.set_seed(42)
# By default, recurrent layers in Keras only return the final output.
# To make them return one output per time step, you must set return_sequences=True
# number of parameters https://d2l.ai/chapter_recurrent-neural-networks/rnn.html#recurrent-neural-networks-with-hidden-states
model = keras.models.Sequential([
keras.layers.SimpleRNN(20, return_sequences=True, input_shape=[None, 1]), #1*20+20+20*20
keras.layers.SimpleRNN(20, return_sequences=True), #20*20++20+20*20
keras.layers.SimpleRNN(1) #20*1+1+1*1
])
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="SfAv-77vDlcm" outputId="7138b70e-7342-4ca4-8549-28f58f537b8d"
model.compile(loss="mse", optimizer="adam")
history = model.fit(X_train, y_train, epochs=20,
validation_data=(X_valid, y_valid))
# + id="42_p59jH5jk8" outputId="4bd43337-53b7-451b-87a7-4acf6cc28e3a" colab={"base_uri": "https://localhost:8080/"}
model.evaluate(X_valid, y_valid)
# + id="fsQp1KDq5jk9" outputId="4d353299-c25f-4a96-da4b-f6d63ae212d6" colab={"base_uri": "https://localhost:8080/", "height": 291}
plot_learning_curves(history.history["loss"], history.history["val_loss"])
plt.show()
# + id="HAS8HZnX5jk9" outputId="f53af0f5-72cc-43d4-d48c-d6214f166d76" colab={"base_uri": "https://localhost:8080/", "height": 293}
y_pred = model.predict(X_valid)
plot_series(X_valid[0, :, 0], y_valid[0, 0], y_pred[0, 0])
plt.show()
# + [markdown] id="7Mk8hrHT5jk9"
# Since a SimpleRNN layer uses the tanh activation function by default, the predicted values must lie within the range –1 to 1. It might be preferable to
# replace the output layer with a Dense layer: it would run slightly faster, the accuracy would be roughly the same, and it would allow us to choose any output activation function we want. If you make this change, also make sure to remove `return_sequences=True` from the second (now last) recurrent layer
# + id="nT3gQCrq5jk9" outputId="2aa5a397-0474-4438-9733-360a6ecc9278" colab={"base_uri": "https://localhost:8080/"}
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.SimpleRNN(20, return_sequences=True, input_shape=[None, 1]),
keras.layers.SimpleRNN(20),
keras.layers.Dense(1)
])
model.summary()
# + [markdown] id="IXZoG7QA5jk-"
# ## Forecasting Several Steps Ahead
# + [markdown] id="cg0V6AMCJ8Zg"
# So far we have only predicted the value at the next time step, but we could just as easily have predicted the value several steps ahead by changing the targets appropriately (e.g., to predict 10 steps ahead, just change the targets
# to be the value 10 steps ahead instead of 1 step ahead). But what if we want to predict the next 10 values?
#
# The first option is to use the model we already trained, make it predict the next value, then add that value to the inputs (acting as if this predicted value had actually occurred), and use the model again to predict the following
# value, and so on, as in the following code:
# + id="aEXPkSXD5jk-"
np.random.seed(43) # not 42, as it would give the first series in the train set
series = generate_time_series(1, n_steps + 10)
X_new, Y_new = series[:, :n_steps], series[:, n_steps:]
X = X_new
for step_ahead in range(10):
y_pred_one = model.predict(X[:, step_ahead:])[:, np.newaxis, :]
X = np.concatenate([X, y_pred_one], axis=1)
Y_pred = X[:, n_steps:]
# + id="2QT45Nd85jk-" outputId="4b717995-5a87-43c0-8814-4c06a28b59a3" colab={"base_uri": "https://localhost:8080/"}
Y_pred.shape
# + id="sJ_ocxoY5jk-" outputId="c614333d-4fdd-41fe-97ea-a9d5fe9577a4" colab={"base_uri": "https://localhost:8080/", "height": 314}
def plot_multiple_forecasts(X, Y, Y_pred):
n_steps = X.shape[1]
ahead = Y.shape[1]
plot_series(X[0, :, 0])
plt.plot(np.arange(n_steps, n_steps + ahead), Y[0, :, 0], "bo-", label="Actual")
plt.plot(np.arange(n_steps, n_steps + ahead), Y_pred[0, :, 0], "rx-", label="Forecast", markersize=10)
plt.axis([0, n_steps + ahead, -1, 1])
plt.legend(fontsize=14)
plot_multiple_forecasts(X_new, Y_new, Y_pred)
save_fig("forecast_ahead_plot")
plt.show()
# + [markdown] id="SkRQp2eMLsDS"
# As you might expect, the prediction for the next step will usually be more accurate than the predictions for later time steps, since the errors might accumulate. If you only want to forecast a few time steps ahead, on more
# complex tasks, this approach may work well.
# + id="r3mvSba_5jk-"
np.random.seed(42)
n_steps = 50
series = generate_time_series(10000, n_steps + 10)
X_train, Y_train = series[:7000, :n_steps], series[:7000, -10:, 0]
X_valid, Y_valid = series[7000:9000, :n_steps], series[7000:9000, -10:, 0]
X_test, Y_test = series[9000:, :n_steps], series[9000:, -10:, 0]
# + [markdown] id="HDbQm3hU5jk-"
# Now let's predict the next 10 values one by one:
# + id="Nb9XFiww5jk_"
X = X_valid
for step_ahead in range(10):
y_pred_one = model.predict(X)[:, np.newaxis, :]
X = np.concatenate([X, y_pred_one], axis=1)
Y_pred = X[:, n_steps:, 0]
# + id="ioAnpOO75jk_" outputId="e2c25792-d471-4410-e9c7-639ea3b9bc71" colab={"base_uri": "https://localhost:8080/"}
Y_pred.shape
# + id="JSeX3I_y5jk_" outputId="9c0fe992-efbc-4dd4-cf9b-c6e34d711664" colab={"base_uri": "https://localhost:8080/"}
np.mean(keras.metrics.mean_squared_error(Y_valid, Y_pred))
# + [markdown] id="mLeMjfsH5jk_"
# Let's compare this performance with some baselines: naive predictions:
# + id="oYYPT19r5jk_" outputId="3c3443e1-ff3a-4da8-da44-54a5c20273c1" colab={"base_uri": "https://localhost:8080/"}
Y_naive_pred = np.tile(X_valid[:, -1], 10) # take the last time step value, and repeat it 10 times
np.mean(keras.metrics.mean_squared_error(Y_valid, Y_naive_pred))
# + [markdown] id="Hb7kFoGB5jk_"
# The second option is to train an RNN to predict all 10 next values at once. We can still use a sequence-to-vector model, but it will output 10 values instead of 1. Now we just need the output layer to have 10 units instead of 1:
# + colab={"base_uri": "https://localhost:8080/"} id="3JAX6ywcMpTZ" outputId="094bf7d4-fef8-4c70-faec-59eadcab1aba"
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.SimpleRNN(20, return_sequences=True, input_shape=[None, 1]),
keras.layers.SimpleRNN(20),
keras.layers.Dense(10)
])
model.summary()
# + id="v6AHQGbF5jk_" outputId="8bacd10d-4c87-41c4-a0b1-42ac4eb4c5fa" colab={"base_uri": "https://localhost:8080/"}
model.compile(loss="mse", optimizer="adam")
history = model.fit(X_train, Y_train, epochs=20,
validation_data=(X_valid, Y_valid))
# + id="CTOKPakr5jlA"
np.random.seed(43)
series = generate_time_series(1, 50 + 10)
X_new, Y_new = series[:, :50, :], series[:, -10:, :]
Y_pred = model.predict(X_new)[..., np.newaxis]
# + id="J2sKchZu5jlA" outputId="8ac7905d-f1cd-4f9b-e2e4-742405e7668a" colab={"base_uri": "https://localhost:8080/", "height": 293}
plot_multiple_forecasts(X_new, Y_new, Y_pred)
plt.show()
# + [markdown] id="2lxUhMIvNId7"
# This model works nicely: the MSE for the next 10 time steps is about 0.008. That’s much better than predicted one by one. But we can still do better: indeed, instead of training the model to forecast the next 10 values only at the
# very last time step, **we can train it to forecast the next 10 values at each and every time step.** In other words, we can turn this sequence-to-vector RNN into a sequence-to-sequence RNN. The advantage of this technique is that the loss will contain a term for the output of the RNN at each and every time step, not just the output at the last time step. This means there will be many more error gradients flowing through the model, and they won’t have to flow only through time; they will also flow from the output of each time step. This will both stabilize and speed up training.
#
# Now let's create an RNN that predicts the next 10 steps at each time step. That is, instead of just forecasting time steps 50 to 59 based on time steps 0 to 49, it will forecast time steps 1 to 10 at time step 0, then time steps 2 to 11 at time step 1, and so on, and finally it will forecast time steps 50 to 59 at the last time step. So each target must be a sequence of the same
# length as the input sequence, containing a 10-dimensional vector at each step. Let’s prepare these target sequences:
# + id="sRRkwZGd5jlA"
np.random.seed(42)
# Notice that the model is still causal: when it makes predictions at
# any time step, it can only see past time steps.
n_steps = 50
series = generate_time_series(10000, n_steps + 10)
X_train = series[:7000, :n_steps]
X_valid = series[7000:9000, :n_steps]
X_test = series[9000:, :n_steps]
Y = np.empty((10000, n_steps, 10)) # each target is a sequence of 10D vectors
for step_ahead in range(1, 10 + 1):
Y[:,:, step_ahead - 1] = series[:, step_ahead:step_ahead + n_steps, 0]
Y_train = Y[:7000]
Y_valid = Y[7000:9000]
Y_test = Y[9000:]
# + id="G0w2fo8M5jlA" outputId="95d507e9-1197-4ccb-d732-ae536a6f508d" colab={"base_uri": "https://localhost:8080/"}
X_train.shape, Y_train.shape, series.shape
# + [markdown] id="QSdLOYW3PFgj"
# To turn the model into a sequence-to-sequence model, we must set `return_sequences=True` in all recurrent layers (even the last one), and we must **apply the output Dense layer at every time step**. Keras offers a
# `TimeDistributed` layer for this very purpose: it wraps any layer (e.g., a Dense layer) and applies it at every time step of its input sequence. It does this efficiently, by reshaping the inputs so that each time step is treated as a
# separate instance (i.e., it reshapes the inputs from `[batch size, time steps, input dimensions]` to `[batch size×time steps, input dimensions]`. In this example, the number of input dimensions is 20 because the previous SimpleRNN
# layer has 20 units), then it runs the Dense layer, and finally it reshapes the outputs back to sequences (i.e., it reshapes the outputs from `[batch size × time steps, output dimensions]` to `[batch size, time steps, output dimensions]`; in this example the number of output dimensions is 10, since the Dense layer has 10 units). Here is the updated model:
# + id="lkFg1r3-5jlA" outputId="425807e6-1b3c-45b2-e8a9-09f18f961983" colab={"base_uri": "https://localhost:8080/"}
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.SimpleRNN(20, return_sequences=True, input_shape=[None, 1]),
keras.layers.SimpleRNN(20, return_sequences=True),
keras.layers.TimeDistributed(keras.layers.Dense(10))
])
model.summary()
# + [markdown] id="835XgNffQAVq"
# It makes it clear that the Dense layer is applied independently at each time step and that the model will output a sequence, not just a single vector.
#
# All outputs are needed during training, but only the output at the last time step is useful for predictions and for evaluation. So although we will rely on the MSE over all the outputs for training, we will use a custom metric for
# evaluation, to only compute the MSE over the output at the last time step:
# + colab={"base_uri": "https://localhost:8080/"} id="ZfljBHwIPzVS" outputId="1b640b93-6771-4644-d48d-c7e1f76f2ce5"
def last_time_step_mse(Y_true, Y_pred):
return keras.metrics.mean_squared_error(Y_true[:, -1], Y_pred[:, -1])
model.compile(loss="mse", optimizer=keras.optimizers.Adam(learning_rate=0.01), metrics=[last_time_step_mse])
history = model.fit(X_train, Y_train, epochs=20,
validation_data=(X_valid, Y_valid))
# + id="LT0dimq-5jlA"
np.random.seed(43)
series = generate_time_series(1, 50 + 10)
X_new, Y_new = series[:, :50, :], series[:, 50:, :]
Y_pred = model.predict(X_new)[:, -1][..., np.newaxis]
# + id="7-GHQBIcZR31"
model.evaluate(X_valid, Y_valid)
# + id="DICzAf4-5jlB" outputId="7b867ee3-336f-4d78-9ac7-068ace0a8fc2" colab={"base_uri": "https://localhost:8080/", "height": 293}
plot_multiple_forecasts(X_new, Y_new, Y_pred)
plt.show()
# + [markdown] id="C2xRfWneMqEu"
# You might find https://www.tensorflow.org/api_docs/python/tf/keras/utils/timeseries_dataset_from_array or https://www.tensorflow.org/api_docs/python/tf/keras/preprocessing/sequence/TimeseriesGenerator useful
# + [markdown] id="rAKE21VC5jlB"
# ## Deep RNNs with Layer Norm
# + [markdown] id="QnIADsRAU6lL"
# Let’s use tf.keras to implement Layer Normalization within a simple memory cell. We need to define a custom memory cell. It is just like a regular layer, except its `call()` method takes two arguments: the inputs at the current time step and the hidden states from the previous time step. Note that the states argument is a list containing one or more tensors. In the case of a simple RNN cell it contains a single tensor equal to the outputs of the previous time step, but other cells may have multiple state tensors (e.g., an LSTMCell has a long-term state and a short-term state). A cell must also have a `state_size` attribute and an `output_size` attribute. In a simple RNN, both are simply equal to the number of units. The following code implements a custom memory cell which will behave like a SimpleRNNCell, except it will also apply Layer Normalization at each time step:
# + id="BQ5QfPbp5jlB"
from tensorflow.keras.layers import LayerNormalization
# + id="tMtn5tDX5jlB"
class LNSimpleRNNCell(keras.layers.Layer):
def __init__(self, units, activation="tanh", **kwargs):
super().__init__(**kwargs)
self.state_size = units
self.output_size = units
self.simple_rnn_cell = keras.layers.SimpleRNNCell(units,
activation=None)
self.layer_norm = LayerNormalization()
self.activation = keras.activations.get(activation)
def get_initial_state(self, inputs=None, batch_size=None, dtype=None):
if inputs is not None:
batch_size = tf.shape(inputs)[0]
dtype = inputs.dtype
return [tf.zeros([batch_size, self.state_size], dtype=dtype)]
def call(self, inputs, states):
outputs, new_states = self.simple_rnn_cell(inputs, states)
# in a SimpleRNNCell, the outputs are just equal to the hidden states: new_states[0] is equal to outputs,
# so we can safely ignore new_states in the rest of the call() method.
norm_outputs = self.activation(self.layer_norm(outputs))
return norm_outputs, [norm_outputs]
# + [markdown] id="K7qg9UpqVdDd"
# Similarly, you could create a custom cell to apply dropout between each time step. But there’s a simpler way: all recurrent layers and all cells provided by Keras have a dropout hyperparameter and a recurrent_dropout hyperparameter: the **former defines the dropout rate to apply to the inputs** (at each time step), and the latter defines the **dropout rate for the hidden states** (also at each time step).
# + id="jZa2bF8R5jlB" outputId="0f7d7af8-1726-4807-9f24-91b9c22e7e2e" colab={"base_uri": "https://localhost:8080/"}
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.RNN(LNSimpleRNNCell(20), return_sequences=True,
input_shape=[None, 1]),
keras.layers.RNN(LNSimpleRNNCell(20), return_sequences=True),
keras.layers.TimeDistributed(keras.layers.Dense(10))
])
model.compile(loss="mse", optimizer="adam", metrics=[last_time_step_mse])
history = model.fit(X_train, Y_train, epochs=20,
validation_data=(X_valid, Y_valid))
# + colab={"base_uri": "https://localhost:8080/"} id="4EfwMTdzZOMf" outputId="86db371a-a6da-4814-b7ee-d4e7a30671ec"
model.evaluate(X_valid, Y_valid)
# + [markdown] id="3QR4Q74s5jlC"
# ## LSTMs
# + [markdown] id="6VSPGUazWzI1"
# In Keras, you can simply use the LSTM layer instead of the SimpleRNN layer:
# + id="hFClcQIw5jlC" outputId="9286bdac-5599-4c79-e1c7-7db3b147024e" colab={"base_uri": "https://localhost:8080/"}
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.LSTM(20, return_sequences=True, input_shape=[None, 1]),
keras.layers.LSTM(20, return_sequences=True),
keras.layers.TimeDistributed(keras.layers.Dense(10))
])
model.summary()
# + id="-YL5WRMMXxvr"
# keras.layers.RNN(keras.layers.LSTMCell(20), return_sequences=True, input_shape=[None, 1]) also works
# However, the LSTM layer uses an optimized implementation when running on a GPU
# RNN layer is mostly useful when you define custom cells, as we did earl
# + colab={"base_uri": "https://localhost:8080/"} id="LdL-BteUXio0" outputId="e9748267-a240-43d3-c18b-a0c67924364d"
model.compile(loss="mse", optimizer="adam", metrics=[last_time_step_mse])
history = model.fit(X_train, Y_train, epochs=20,
validation_data=(X_valid, Y_valid))
# + id="H-BCnT5d5jlC" outputId="b2a3e1da-15b6-4093-dec0-5d20c4f99f66" colab={"base_uri": "https://localhost:8080/"}
model.evaluate(X_valid, Y_valid)
# + id="dHvxuwIw5jlC" outputId="8e240048-c556-4fed-84a9-e7a1eff224af" colab={"base_uri": "https://localhost:8080/", "height": 291}
plot_learning_curves(history.history["loss"], history.history["val_loss"])
plt.show()
# + id="yJIAA4wP5jlC"
np.random.seed(43)
series = generate_time_series(1, 50 + 10)
X_new, Y_new = series[:, :50, :], series[:, 50:, :]
Y_pred = model.predict(X_new)[:, -1][..., np.newaxis]
# + id="9Q83vZbA5jlC" outputId="fca5256c-633c-415a-8d97-38d8a15b4f6d" colab={"base_uri": "https://localhost:8080/", "height": 293}
plot_multiple_forecasts(X_new, Y_new, Y_pred)
plt.show()
# + [markdown] id="V5kAzvP15jlD"
# ## GRUs
# + colab={"base_uri": "https://localhost:8080/"} id="U0pp1tbVcFEP" outputId="3dedff16-df49-459d-aa13-4f607d010e1a"
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.GRU(20, return_sequences=True, input_shape=[None, 1]),
keras.layers.GRU(20, return_sequences=True),
keras.layers.TimeDistributed(keras.layers.Dense(10))
])
model.summary()
# + id="cV-aoeD95jlD" outputId="aa2fe19b-4891-4f77-e507-0751f2053dde" colab={"base_uri": "https://localhost:8080/"}
model.compile(loss="mse", optimizer="adam", metrics=[last_time_step_mse])
history = model.fit(X_train, Y_train, epochs=20,
validation_data=(X_valid, Y_valid))
# + id="TNOeSZW_5jlD" outputId="7e827299-42da-4e52-e341-3ed81d0333e9" colab={"base_uri": "https://localhost:8080/"}
model.evaluate(X_valid, Y_valid)
# + [markdown] id="9gDnHqvg5jlD"
# ## Using One-Dimensional Convolutional Layers to Process Sequences
# + [markdown] id="yTdnZoTLYjGN"
# The following model is the same as earlier, except it starts with a 1D convolutional layer that downsamples the input sequence by a factor of 2, using
# a stride of 2. By shortening the sequences, the convolutional layer may help the GRU layers detect longer patterns. Note that we must also crop off the first three time steps in the targets (since the kernel’s size is 4, the first output of the convolutional layer will be based on the input time steps 0 to 3), and downsample the targets by a factor of 2:
# + id="9PKOidly5jlE" outputId="d77c8947-1acc-497e-b7b0-df3dcbfb0fe9" colab={"base_uri": "https://localhost:8080/"}
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.Conv1D(filters=20, kernel_size=4, strides=2, padding="valid",
input_shape=[None, 1]),
keras.layers.GRU(20, return_sequences=True),
keras.layers.GRU(20, return_sequences=True),
keras.layers.TimeDistributed(keras.layers.Dense(10))
])
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="kMugaQpAcKFB" outputId="2b4564bd-988a-4735-b9a2-c6e029300045"
model.compile(loss="mse", optimizer="adam", metrics=[last_time_step_mse])
history = model.fit(X_train, Y_train[:, 3::2], epochs=20,
validation_data=(X_valid, Y_valid[:, 3::2]))
# + colab={"base_uri": "https://localhost:8080/"} id="DlBLXViLcOxn" outputId="cf835442-e81f-4be9-8b09-22942cca6d01"
model.evaluate(X_valid, Y_valid[:, 3::2])
# + [markdown] id="e16QVbDxcPb2"
# # Natural-language processing
# + [markdown] id="Vq_YGZBzcZsn"
# ## Preparing text data
# + [markdown] id="ACRmbWfKdbvv"
# Vectorizing process using Python may be done as follows
# + id="JPAOQHVAdZTR"
import string
class Vectorizer:
def standardize(self, text):
text = text.lower()
return "".join(char for char in text if char not in string.punctuation)
def tokenize(self, text):
text = self.standardize(text)
return text.split()
def make_vocabulary(self, dataset):
self.vocabulary = {"": 0, "[UNK]": 1}
for text in dataset:
text = self.standardize(text)
tokens = self.tokenize(text)
for token in tokens:
if token not in self.vocabulary:
self.vocabulary[token] = len(self.vocabulary)
self.inverse_vocabulary = dict(
(v, k) for k, v in self.vocabulary.items())
def encode(self, text):
text = self.standardize(text)
tokens = self.tokenize(text)
return [self.vocabulary.get(token, 1) for token in tokens]
def decode(self, int_sequence):
return " ".join(
self.inverse_vocabulary.get(i, "[UNK]") for i in int_sequence)
vectorizer = Vectorizer()
dataset = [
"I write, erase, rewrite",
"Erase again, and then",
"A poppy blooms.",
]
vectorizer.make_vocabulary(dataset)
# + colab={"base_uri": "https://localhost:8080/"} id="VGgm3yP3drce" outputId="6b368c1a-3ec3-48e9-e7f1-ad80eae4dc9c"
test_sentence = "I write, rewrite, and still rewrite again"
encoded_sentence = vectorizer.encode(test_sentence)
print(encoded_sentence)
# + colab={"base_uri": "https://localhost:8080/"} id="yeVLfOrAdxRf" outputId="7f564f9f-8911-46dd-c9bd-04f4ffde58a4"
decoded_sentence = vectorizer.decode(encoded_sentence)
print(decoded_sentence)
# + [markdown] id="cF6Xq7YMcip3"
# However, using something like this wouldn’t be very performant. In practice, you’ll work with the Keras `TextVectorization` layer, which is fast and efficient and can be dropped directly into a `tf.data` pipeline or a Keras model.
# + id="7oWACab-cTGJ"
from tensorflow.keras.layers import TextVectorization
# Configures the layer to return sequences of words encoded
# as integer indices.
text_vectorization = TextVectorization(
output_mode="int",
)
# + [markdown] id="t8PWeaj0eUSp"
# By default, the TextVectorization layer will use the setting “convert to lowercase and remove punctuation” for text standardization, and “split on whitespace” for tokenization.
#
# But importantly, you can provide custom functions for standardization and tokenization, which means the layer is flexible enough to handle any use case.To index the vocabulary of a text corpus, just call the `adapt()` method of the layer with a `Dataset` object that yields strings, or just with a list of Python strings:
# + id="9IqhI0sceL0Q"
dataset = [
"I write, erase, rewrite",
"Erase again, and then",
"A poppy blooms.",
]
text_vectorization.adapt(dataset)
# + [markdown] id="tMljNvynex1J"
# Note that you can retrieve the computed vocabulary via `get_vocabulary()`—this can be useful if you need to convert text encoded as integer sequences back into words. The first two entries in the vocabulary are the mask token (index 0) and the OOV token (index 1). Entries in the vocabulary list are sorted by frequency, so with a realworld dataset, very common words like “the” or “a” would come first.
# + colab={"base_uri": "https://localhost:8080/"} id="y3XeB2HYegB6" outputId="3398c130-6067-4b33-9979-679a4d2eb66d"
text_vectorization.get_vocabulary()
# + colab={"base_uri": "https://localhost:8080/"} id="3OldsH4Qe4Qb" outputId="da56367e-1b43-4285-ebf7-36d48b1451a6"
vocabulary = text_vectorization.get_vocabulary()
test_sentence = "I write, rewrite, and still rewrite again"
encoded_sentence = text_vectorization(test_sentence)
print(encoded_sentence)
# + colab={"base_uri": "https://localhost:8080/"} id="v94pHFXUe-MO" outputId="b3a5e18d-79df-4594-8d6f-79f0933a4fa4"
inverse_vocab = dict(enumerate(vocabulary))
decoded_sentence = " ".join(inverse_vocab[int(i)] for i in encoded_sentence)
print(decoded_sentence)
# + [markdown] id="DVxg9nzSfPuY"
# Before dive into the modeling part. We’ll demonstrate each approach on a well-known text classification benchmark: the IMDB movie review sentiment-classification dataset. Let’s process the raw
# IMDB text data, just like you would do when approaching a new text-classification problem in the real world.
#
# https://ai.stanford.edu/~amaas/data/sentiment/
#
# Let’s start by downloading the dataset from the Stanford page of <NAME> and uncompressing it
# + colab={"base_uri": "https://localhost:8080/"} id="WWTJRDZXfOtX" outputId="b342fa45-bee0-423a-936a-d0176d393283"
# !curl -O https://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz
# !tar -xf aclImdb_v1.tar.gz
# + [markdown] id="9Firy_UuhRf4"
# There’s also a `train/unsup` subdirectory in there, which we don’t need. Let’s
# delete it:
# + id="y5jDnHImg4lT"
# !rm -r aclImdb/train/unsup
# + [markdown] id="jdRkfu9uhXbx"
# Take a look at the content of a few of these text files. Whether you’re working with text data or image data, remember to always inspect what your data looks like before you dive into modeling it.
# + colab={"base_uri": "https://localhost:8080/"} id="DzJMWYO0hZOO" outputId="250da9ff-4194-402a-a1a4-d6d791c92d1e"
# !cat aclImdb/train/pos/4077_10.txt
# + [markdown] id="oEplxGV0hcyS"
# Next, let’s prepare a validation set by setting apart 20% of the training text files in a new directory, aclImdb/val:
# + id="uI1RHMXfheLP"
import os, pathlib, shutil, random
base_dir = pathlib.Path("aclImdb")
val_dir = base_dir / "val"
train_dir = base_dir / "train"
for category in ("neg", "pos"):
os.makedirs(val_dir / category)
files = os.listdir(train_dir / category)
random.Random(1337).shuffle(files)
num_val_samples = int(0.2 * len(files))
val_files = files[-num_val_samples:]
for fname in val_files:
shutil.move(train_dir / category / fname,
val_dir / category / fname)
# + [markdown] id="9rwl4rFphkdx"
# Let’s create three Dataset objects for training, validation, and testing just like previous lab:
# + colab={"base_uri": "https://localhost:8080/"} id="672YOyLXhnyI" outputId="63913c02-61eb-44ab-bff5-7e830afa81e2"
from tensorflow import keras
batch_size = 32
train_ds = keras.utils.text_dataset_from_directory(
"aclImdb/train", batch_size=batch_size
)
val_ds = keras.utils.text_dataset_from_directory(
"aclImdb/val", batch_size=batch_size
)
test_ds = keras.utils.text_dataset_from_directory(
"aclImdb/test", batch_size=batch_size
)
# + [markdown] id="tILAl18fh9FA"
# These datasets yield inputs that are TensorFlow `tf.string` tensors and targets that are `int32` tensors encoding the value “0” or “1.”
# + colab={"base_uri": "https://localhost:8080/"} id="gD_XsKfih3XR" outputId="9a9ceccd-17c1-4f8b-d128-6449ef5f28c0"
for inputs, targets in train_ds:
print("inputs.shape:", inputs.shape)
print("inputs.dtype:", inputs.dtype)
print("targets.shape:", targets.shape)
print("targets.dtype:", targets.dtype)
print("inputs[0]:", inputs[0])
print("targets[0]:", targets[0])
break
# + [markdown] id="nXU16HSgiGAK"
# ## Processing words as a set: The bag-of-words approach
# + [markdown] id="6akoHLcIkVXr"
# ### Single words (unigrams) with binary encoding
# + [markdown] id="E0-NGwoeiNvx"
# First, let’s process our raw text datasets with a `TextVectorization` layer so that they yield multi-hot encoded binary word vectors. Our layer will only look at single words (that is to say, unigrams). We will limit the vocabulary to the 20,000 most frequent words. Otherwise we’d be indexing every word in the training data— potentially tens of thousands of terms that only occur once or
# twice and thus aren’t informative. In general, 20,000 is the right vocabulary size for text classification.
# + id="SV_WnlbKh2xK"
# Encode the output tokens as multi-hot binary vectors.
text_vectorization = TextVectorization(
max_tokens=20000,
output_mode="multi_hot",
)
# Prepare a dataset that only yields raw text inputs (no labels).
text_only_train_ds = train_ds.map(lambda x, y: x)
text_vectorization.adapt(text_only_train_ds)
binary_1gram_train_ds = train_ds.map(
lambda x, y: (text_vectorization(x), y),
num_parallel_calls=4)
binary_1gram_val_ds = val_ds.map(
lambda x, y: (text_vectorization(x), y),
num_parallel_calls=4)
binary_1gram_test_ds = test_ds.map(
lambda x, y: (text_vectorization(x), y),
num_parallel_calls=4)
# + colab={"base_uri": "https://localhost:8080/"} id="rgJwHXctiuIH" outputId="b167570c-c6d3-4b6e-c0d3-d4ac00780454"
for inputs, targets in binary_1gram_train_ds:
print("inputs.shape:", inputs.shape)
print("inputs.dtype:", inputs.dtype)
print("targets.shape:", targets.shape)
print("targets.dtype:", targets.dtype)
print("inputs[0]:", inputs[0])
print("targets[0]:", targets[0])
break
# + id="DqwEhyqfiyma"
from tensorflow import keras
from tensorflow.keras import layers
# A densely connected NN
def get_model(max_tokens=20000, hidden_dim=16):
inputs = keras.Input(shape=(max_tokens,))
x = layers.Dense(hidden_dim, activation="relu")(inputs)
x = layers.Dropout(0.5)(x)
outputs = layers.Dense(1, activation="sigmoid")(x)
model = keras.Model(inputs, outputs)
model.compile(optimizer="nadam",
loss="binary_crossentropy",
metrics=["accuracy"])
return model
# + [markdown] id="yMTMyNB5j0xq"
# Finally, let’s train and test our model.
# + colab={"base_uri": "https://localhost:8080/"} id="h1ws10OIjt_X" outputId="eba30619-0f49-4880-d7e9-219acae9bc26"
model = get_model()
model.summary()
callbacks = [
keras.callbacks.ModelCheckpoint("binary_1gram.keras",
save_best_only=True)
]
model.fit(binary_1gram_train_ds.cache(),
validation_data=binary_1gram_val_ds.cache(),
epochs=10,
callbacks=callbacks)
model = keras.models.load_model("binary_1gram.keras")
print(f"Test acc: {model.evaluate(binary_1gram_test_ds)[1]:.3f}")
# + [markdown] id="YbuIiJwYkGiR"
# This gets us to a test accuracy of 88.4%: not bad!
# + [markdown] id="2SoPSzSrkOXL"
# ### Bigrams with binary encoding
# + [markdown] id="YOwpIaJjkeka"
# The `TextVectorization` layer can be configured to return arbitrary N-grams: bigrams, trigrams, etc. Just pass an `ngrams=N` argument as in the following listing.
# + id="GC3ZrIQmj3VB"
text_vectorization = TextVectorization(
ngrams=2,
max_tokens=20000,
output_mode="multi_hot",
)
# + colab={"base_uri": "https://localhost:8080/"} id="SdEQkw3mkhk9" outputId="5cfc6f2d-3521-43e2-8c05-d995b9824d7c"
text_vectorization.adapt(text_only_train_ds)
binary_2gram_train_ds = train_ds.map(
lambda x, y: (text_vectorization(x), y),
num_parallel_calls=4)
binary_2gram_val_ds = val_ds.map(
lambda x, y: (text_vectorization(x), y),
num_parallel_calls=4)
binary_2gram_test_ds = test_ds.map(
lambda x, y: (text_vectorization(x), y),
num_parallel_calls=4)
model = get_model()
model.summary()
callbacks = [
keras.callbacks.ModelCheckpoint("binary_2gram.keras",
save_best_only=True)
]
model.fit(binary_2gram_train_ds.cache(),
validation_data=binary_2gram_val_ds.cache(),
epochs=10,
callbacks=callbacks)
model = keras.models.load_model("binary_2gram.keras")
print(f"Test acc: {model.evaluate(binary_2gram_test_ds)[1]:.3f}")
# + [markdown] id="0aGRj_DRkqtD"
# We’re now getting 89.5% test accuracy, a marked improvement! Turns out local order is pretty important.
# + [markdown] id="CB3pEfxck15o"
# ### Bigrams with TF-IDF encoding
# + [markdown] id="tPG88cwWk5zS"
# TF-IDF is so common that it’s built into the TextVectorization layer. All you need to do to start using it is to switch the output_mode argument to `tf_idf`.
# + id="uRaK89lhnenM"
from keras.layers import TextVectorization as TexVec
# + colab={"base_uri": "https://localhost:8080/", "height": 503} id="m2V7i76UklOO" outputId="fd33c64c-57cd-44ba-829c-99ddcb61c660"
text_vectorization = TexVec(
ngrams=2,
max_tokens=20000,
output_mode="tf_idf",
)
text_vectorization.adapt(text_only_train_ds)
# + id="MDp4V3VjlGVh"
text_vectorization.adapt(text_only_train_ds)
tfidf_2gram_train_ds = train_ds.map(
lambda x, y: (text_vectorization(x), y),
num_parallel_calls=4)
tfidf_2gram_val_ds = val_ds.map(
lambda x, y: (text_vectorization(x), y),
num_parallel_calls=4)
tfidf_2gram_test_ds = test_ds.map(
lambda x, y: (text_vectorization(x), y),
num_parallel_calls=4)
model = get_model()
model.summary()
callbacks = [
keras.callbacks.ModelCheckpoint("tfidf_2gram.keras",
save_best_only=True)
]
model.fit(tfidf_2gram_train_ds.cache(),
validation_data=tfidf_2gram_val_ds.cache(),
epochs=10,
callbacks=callbacks)
model = keras.models.load_model("tfidf_2gram.keras")
print(f"Test acc: {model.evaluate(tfidf_2gram_test_ds)[1]:.3f}")
# + [markdown] id="qoz77TP4oewU"
# This gets us an 89.8% test accuracy on the IMDB classification task: it doesn’t seem to be particularly helpful in this case. However, for many text-classification datasets, it would be typical to see a one-percentage-point increase when using TF-IDF compared to plain binary encoding.
# + [markdown] id="ZC7yKzENvV5N"
# ## Processing words as a sequence: The sequence model approach
# + [markdown] id="IC3yN0qevZM2"
# Let’s try out a first sequence model in practice. First, let’s prepare datasets that return integer sequences. In order to keep a manageable input size, **we’ll truncate the inputs after the first 600 words.**
#
# This is a reasonable choice, since the average review length is 233 words, and only 5% of reviews are longer than 600 words.
# + id="8__1VvdClKay"
max_length = 600
max_tokens = 20000
text_vectorization = layers.TextVectorization(
max_tokens=max_tokens,
output_mode="int",
output_sequence_length=max_length,
)
text_vectorization.adapt(text_only_train_ds)
int_train_ds = train_ds.map(
lambda x, y: (text_vectorization(x), y),
num_parallel_calls=4)
int_val_ds = val_ds.map(
lambda x, y: (text_vectorization(x), y),
num_parallel_calls=4)
int_test_ds = test_ds.map(
lambda x, y: (text_vectorization(x), y),
num_parallel_calls=4)
# + [markdown] id="ezd52NzTvmwl"
# Next, let’s make a model. The simplest way to convert our integer sequences to vector sequences is to one-hot encode the integers (each dimension would represent one possible term in the vocabulary). On top of these one-hot vectors, we’ll add a simple bidirectional LSTM.
# + colab={"base_uri": "https://localhost:8080/"} id="jEGBs6wjvs5s" outputId="29b2e209-9bff-41ad-8f6d-cd443ffe2480"
inputs = keras.Input(shape=(None,), dtype="int64") # One input is a sequence of integers
embedded = tf.one_hot(inputs, depth=max_tokens) # A 3D tensor of shape [batch size, time steps, embedding size]
x = layers.Bidirectional(layers.LSTM(32))(embedded)
x = layers.Dropout(0.5)(x)
outputs = layers.Dense(1, activation="sigmoid")(x) # Classification layer
model = keras.Model(inputs, outputs)
model.compile(optimizer="nadam",
loss="binary_crossentropy",
metrics=["accuracy"])
model.summary()
# + [markdown] id="fUDH3SWRz9u3"
# A first observation: this model will train very slowly, especially compared to the lightweight model of the previous section. This is because our inputs are quite large: each input sample is encoded as a matrix of size `(600, 20000)` (600 words per sample, 20,000 possible words). That’s 12,000,000 floats for a single movie review. Our bidirectional LSTM has a lot of work to do.
#
# Let's try word embedding. What makes a good word-embedding space depends heavily on your task: the perfect word-embedding space for an English-language movie-review sentiment-analysis model may look different from the perfect embedding space for an English-language legal-document classification model, because the importance of certain semantic relationships varies from task to task. It’s thus reasonable to learn a new embedding space with every new task. Fortunately, backpropagation makes this easy, and Keras makes it even easier. It’s about learning the weights of a layer: the Embedding layer.
# + id="YYAaa3CNzbWY"
embedding_layer = layers.Embedding(input_dim=max_tokens, output_dim=256)
# + [markdown] id="aspfIuYU1b83"
# The Embedding layer is best understood as a dictionary that maps integer indices (which stand for specific words) to dense vectors. The Embedding layer takes as input a rank-2 tensor of integers, of shape `(batch_size,sequence_length)`, where each entry is a sequence of integers. The layer then returns a 3D floating-point tensor of shape` (batch_size, sequence_length, embedding_dimensionality)`.
#
# When you instantiate an Embedding layer, its weights (its internal dictionary of
# token vectors) are initially random, just as with any other layer. During training, these word vectors are gradually adjusted via backpropagation, structuring the space into something the downstream model can exploit. Once fully trained, the embedding space will show a lot of structure—a kind of structure specialized for the specific problem for which you’re training your model. We will talk more about embedding in Lecture 7.
# + colab={"base_uri": "https://localhost:8080/"} id="d1F2Z7fj1UJ-" outputId="0d76ca08-5e7c-4951-8d2f-508e0cc0f415"
inputs = keras.Input(shape=(None,), dtype="int64")
embedded = layers.Embedding(input_dim=max_tokens, output_dim=256)(inputs)
x = layers.Bidirectional(layers.LSTM(32))(embedded)
x = layers.Dropout(0.5)(x)
outputs = layers.Dense(1, activation="sigmoid")(x)
model = keras.Model(inputs, outputs)
model.compile(optimizer="rmsprop",
loss="binary_crossentropy",
metrics=["accuracy"])
model.summary()
callbacks = [
keras.callbacks.ModelCheckpoint("embeddings_bidir_gru.keras",
save_best_only=True)
]
model.fit(int_train_ds, validation_data=int_val_ds, epochs=10, callbacks=callbacks)
model = keras.models.load_model("embeddings_bidir_gru.keras")
print(f"Test acc: {model.evaluate(int_test_ds)[1]:.3f}")
# + [markdown] id="_3dQg0bM2Ks3"
# We’re still some way off from the results of our basic bigram model. Part of the reason why is simply that the model is looking at slightly less data:
# the bigram model processed full reviews, while our sequence model truncates sequences after 600 words. One thing that’s slightly hurting model performance here is that our input sequences are full of zeros. This comes from our use of the `output_sequence_length=max_length` option in `TextVectorization` (with max_length equal to 600): **sentences longer than 600 tokens are truncated to a length of 600 tokens, and sentences shorter than 600 tokens are padded with zeros** at the end so that they can be concatenated together with other sequences to form contiguous batches.
#
# We’re using a bidirectional RNN: two RNN layers running in parallel, with one processing the tokens in their natural order, and the other processing the same
# tokens in reverse. **The RNN that looks at the tokens in their natural order will spend its last iterations seeing only vectors that encode padding—possibly for several hundreds of iterations if the original sentence was short**. The information stored in the internal state of the RNN will gradually fade out as it gets exposed to these meaningless inputs.
#
# We need some way to tell the RNN that it should skip these iterations. There’s an API for that: *masking*. The Embedding layer is capable of generating a“mask” that corresponds to its input data. This mask is a tensor of ones and zeros (or `True/False` booleans), of shape `(batch_size, sequence_length)`,where the entry `mask[i, t]` indicates where timestep t of sample i should be skipped or not (the timestep will be skipped if `mask[i, t]` is 0 or False, and processed otherwise).
# + colab={"base_uri": "https://localhost:8080/"} id="h--MrGck12LD" outputId="48d231b8-a68d-4644-c2a3-d9add6b8cd68"
inputs = keras.Input(shape=(None,), dtype="int64")
embedded = layers.Embedding(
input_dim=max_tokens, output_dim=256, mask_zero=True)(inputs) # You can turn it on by passing mask_zero=True
x = layers.Bidirectional(layers.LSTM(32))(embedded)
x = layers.Dropout(0.5)(x)
outputs = layers.Dense(1, activation="sigmoid")(x)
model = keras.Model(inputs, outputs)
model.compile(optimizer="rmsprop",
loss="binary_crossentropy",
metrics=["accuracy"])
model.summary()
callbacks = [
keras.callbacks.ModelCheckpoint("embeddings_bidir_gru_with_masking.keras",
save_best_only=True)
]
model.fit(int_train_ds, validation_data=int_val_ds, epochs=10, callbacks=callbacks)
model = keras.models.load_model("embeddings_bidir_gru_with_masking.keras")
print(f"Test acc: {model.evaluate(int_test_ds)[1]:.3f}")
# + [markdown] id="hycMo2qy3uO_"
# ## The Transformer encoder
# + [markdown] id="ozI7Y4wx6ScJ"
# The encoder part of transformer can be used for text classification—it’s a very generic module that ingests a sequence and learns to turn it into a more useful representation. Let’s implement a Transformer encoder using Kera subclassing API.
# + id="3mjNWl3L5SFK"
class TransformerEncoder(layers.Layer):
def __init__(self, embed_dim, dense_dim, num_heads, **kwargs):
super().__init__(**kwargs)
self.embed_dim = embed_dim # Size of the input token vectors
self.dense_dim = dense_dim # Size of the inner dense layer
self.num_heads = num_heads # Number of attention heads
self.attention = layers.MultiHeadAttention(
num_heads=num_heads, key_dim=embed_dim)
self.dense_proj = keras.Sequential(
[layers.Dense(dense_dim, activation="relu"),
layers.Dense(embed_dim),]
)
self.layernorm_1 = layers.LayerNormalization()
self.layernorm_2 = layers.LayerNormalization()
# Computation goes in call().
def call(self, inputs, mask=None):
# The mask that will be generated by the Embedding layer will be 2D, but
# the attention layer expects to be 3D or 4D, so we expand its rank.
if mask is not None:
mask = mask[:, tf.newaxis, :]
attention_output = self.attention(
inputs, inputs, attention_mask=mask)
proj_input = self.layernorm_1(inputs + attention_output)
proj_output = self.dense_proj(proj_input)
return self.layernorm_2(proj_input + proj_output)
# Implement serialization so we can save the model.
def get_config(self):
config = super().get_config()
config.update({
"embed_dim": self.embed_dim,
"num_heads": self.num_heads,
"dense_dim": self.dense_dim,
})
return config
# + [markdown] id="-NEaOFJs7aTW"
# When you write custom layers, make sure to implement the `get_config` method: this enables the layer to be reinstantiated from its config dict, which is useful during model saving and loading.
#
# To add positional encoding, we’ll do something simpler and more effective: we’ll learn positionembedding vectors the same way we learn to embed word indices. We’ll then proceed to add our position embeddings to the corresponding word embeddings, to obtain a position-aware word embedding. This technique is called **“positional embedding.”** Let’s implement it. **It is noted that vector.
# neural networks don’t like very large input values, or discrete input distributions** therefore simply adding a position information as interger is not a good idea.
# + id="kaYlKHpS7hgI"
class PositionalEmbedding(layers.Layer):
# A downside of position embeddings is that the sequence length needs to be known in advance.
def __init__(self, sequence_length, input_dim, output_dim, **kwargs):
super().__init__(**kwargs)
# Prepare an Embedding layer for the token indices.
self.token_embeddings = layers.Embedding(
input_dim=input_dim, output_dim=output_dim)
# And another one for the token positions
self.position_embeddings = layers.Embedding(
input_dim=sequence_length, output_dim=output_dim)
self.sequence_length = sequence_length
self.input_dim = input_dim
self.output_dim = output_dim
def call(self, inputs):
length = tf.shape(inputs)[-1]
positions = tf.range(start=0, limit=length, delta=1)
embedded_tokens = self.token_embeddings(inputs)
embedded_positions = self.position_embeddings(positions)
# Add both embedding vectors together
return embedded_tokens + embedded_positions
# Like the Embedding layer, this layer should be able to generate a
# mask so we can ignore padding 0s in the inputs. The compute_mask
# method will called automatically by the framework, and the
# mask will get propagated to the next layer.
def compute_mask(self, inputs, mask=None):
return tf.math.not_equal(inputs, 0)
def get_config(self):
config = super().get_config()
config.update({
"output_dim": self.output_dim,
"sequence_length": self.sequence_length,
"input_dim": self.input_dim,
})
return config
# + [markdown] id="ac1jhOCK-W0q"
# All you have to do to start taking word order into account is swap the old Embedding layer with our position-aware version.
# + colab={"base_uri": "https://localhost:8080/"} id="hA5Bdn6B-d-i" outputId="3c3ad580-9b4b-49cc-acb3-acee89a813dd"
vocab_size = 20000
sequence_length = 600
embed_dim = 256
num_heads = 2
dense_dim = 32
inputs = keras.Input(shape=(None,), dtype="int64")
x = PositionalEmbedding(sequence_length, vocab_size, embed_dim)(inputs)
x = TransformerEncoder(embed_dim, dense_dim, num_heads)(x)
x = layers.GlobalMaxPooling1D()(x)
x = layers.Dropout(0.5)(x)
outputs = layers.Dense(1, activation="sigmoid")(x)
model = keras.Model(inputs, outputs)
model.compile(optimizer="rmsprop",
loss="binary_crossentropy",
metrics=["accuracy"])
model.summary()
callbacks = [
keras.callbacks.ModelCheckpoint("full_transformer_encoder.keras",
save_best_only=True)
]
model.fit(int_train_ds, validation_data=int_val_ds, epochs=20, callbacks=callbacks)
model = keras.models.load_model(
"full_transformer_encoder.keras",
custom_objects={"TransformerEncoder": TransformerEncoder,
"PositionalEmbedding": PositionalEmbedding})
print(f"Test acc: {model.evaluate(int_test_ds)[1]:.3f}")
# + id="9cSp01jOCD_R"
| static_files/presentations/04_Recurrent_Neural_Networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_amazonei_mxnet_p36
# language: python
# name: conda_amazonei_mxnet_p36
# ---
# # Plagiarism Text Data
#
# In this project, you will be tasked with building a plagiarism detector that examines a text file and performs binary classification; labeling that file as either plagiarized or not, depending on how similar the text file is when compared to a provided source text.
#
# The first step in working with any dataset is loading the data in and noting what information is included in the dataset. This is an important step in eventually working with this data, and knowing what kinds of features you have to work with as you transform and group the data!
#
# So, this notebook is all about exploring the data and noting patterns about the features you are given and the distribution of data.
#
# > There are not any exercises or questions in this notebook, it is only meant for exploration. This notebook will note be required in your final project submission.
#
# ---
# ## Read in the Data
#
# The cell below will download the necessary data and extract the files into the folder `data/`.
#
# This data is a slightly modified version of a dataset created by <NAME> (Information Studies) and <NAME> (Computer Science), at the University of Sheffield. You can read all about the data collection and corpus, at [their university webpage](https://ir.shef.ac.uk/cloughie/resources/plagiarism_corpus.html).
#
# > **Citation for data**: <NAME>. and <NAME>. Developing A Corpus of Plagiarised Short Answers, Language Resources and Evaluation: Special Issue on Plagiarism and Authorship Analysis, In Press. [Download]
# +
# # !wget https://s3.amazonaws.com/video.udacity-data.com/topher/2019/January/5c4147f9_data/data.zip
# # !unzip data
# -
# import libraries
import pandas as pd
import numpy as np
import os
# This plagiarism dataset is made of multiple text files; each of these files has characteristics that are is summarized in a `.csv` file named `file_information.csv`, which we can read in using `pandas`.
# +
csv_file = 'data/file_information.csv'
plagiarism_df = pd.read_csv(csv_file)
# print out the first few rows of data info
plagiarism_df.head(10)
# -
# ## Types of Plagiarism
#
# Each text file is associated with one **Task** (task A-E) and one **Category** of plagiarism, which you can see in the above DataFrame.
#
# ### Five task types, A-E
#
# Each text file contains an answer to one short question; these questions are labeled as tasks A-E.
# * Each task, A-E, is about a topic that might be included in the Computer Science curriculum that was created by the authors of this dataset.
# * For example, Task A asks the question: "What is inheritance in object oriented programming?"
#
# ### Four categories of plagiarism
#
# Each text file has an associated plagiarism label/category:
#
# 1. `cut`: An answer is plagiarized; it is copy-pasted directly from the relevant Wikipedia source text.
# 2. `light`: An answer is plagiarized; it is based on the Wikipedia source text and includes some copying and paraphrasing.
# 3. `heavy`: An answer is plagiarized; it is based on the Wikipedia source text but expressed using different words and structure. Since this doesn't copy directly from a source text, this will likely be the most challenging kind of plagiarism to detect.
# 4. `non`: An answer is not plagiarized; the Wikipedia source text is not used to create this answer.
# 5. `orig`: This is a specific category for the original, Wikipedia source text. We will use these files only for comparison purposes.
#
# > So, out of the submitted files, the only category that does not contain any plagiarism is `non`.
#
# In the next cell, print out some statistics about the data.
# print out some stats about the data
print('Number of files: ', plagiarism_df.shape[0]) # .shape[0] gives the rows
# .unique() gives unique items in a specified column
print('Number of unique tasks/question types (A-E): ', (len(plagiarism_df['Task'].unique())))
print('Unique plagiarism categories: ', (plagiarism_df['Category'].unique()))
# You should see the number of text files in the dataset as well as some characteristics about the `Task` and `Category` columns. **Note that the file count of 100 *includes* the 5 _original_ wikipedia files for tasks A-E.** If you take a look at the files in the `data` directory, you'll notice that the original, source texts start with the filename `orig_` as opposed to `g` for "group."
#
# > So, in total there are 100 files, 95 of which are answers (submitted by people) and 5 of which are the original, Wikipedia source texts.
#
# Your end goal will be to use this information to classify any given answer text into one of two categories, plagiarized or not-plagiarized.
# ### Distribution of Data
#
# Next, let's look at the distribution of data. In this course, we've talked about traits like class imbalance that can inform how you develop an algorithm. So, here, we'll ask: **How evenly is our data distributed among different tasks and plagiarism levels?**
#
# Below, you should notice two things:
# * Our dataset is quite small, especially with respect to examples of varying plagiarism levels.
# * The data is distributed fairly evenly across task and plagiarism types.
# +
# Show counts by different tasks and amounts of plagiarism
# group and count by task
counts_per_task=plagiarism_df.groupby(['Task']).size().reset_index(name="Counts")
print("\nTask:")
display(counts_per_task)
# group by plagiarism level
counts_per_category=plagiarism_df.groupby(['Category']).size().reset_index(name="Counts")
print("\nPlagiarism Levels:")
display(counts_per_category)
# group by task AND plagiarism level
counts_task_and_plagiarism=plagiarism_df.groupby(['Task', 'Category']).size().reset_index(name="Counts")
print("\nTask & Plagiarism Level Combos :")
display(counts_task_and_plagiarism)
# -
# It may also be helpful to look at this last DataFrame, graphically.
#
# Below, you can see that the counts follow a pattern broken down by task. Each task has one source text (original) and the highest number on `non` plagiarized cases.
# +
import matplotlib.pyplot as plt
# %matplotlib inline
# counts
group = ['Task', 'Category']
counts = plagiarism_df.groupby(group).size().reset_index(name="Counts")
plt.figure(figsize=(8,5))
plt.bar(range(len(counts)), counts['Counts'], color = 'blue')
# -
# ## Category
df_ex = plagiarism_df.copy()
dic_cat_class = {'cut':0, 'heavy':1, 'light':2, 'non':3, 'orig':-1}
for i,value in enumerate(df_ex['Category']):
df_ex['Category'][i] = dic_cat_class[value]
df_ex
counts_per_category['Category']
# ## Up Next
#
# This notebook is just about data loading and exploration, and you do not need to include it in your final project submission.
#
# In the next few notebooks, you'll use this data to train a complete plagiarism classifier. You'll be tasked with extracting meaningful features from the text data, reading in answers to different tasks and comparing them to the original Wikipedia source text. You'll engineer similarity features that will help identify cases of plagiarism. Then, you'll use these features to train and deploy a classification model in a SageMaker notebook instance.
# Read in a csv file and return a transformed dataframe
def numerical_dataframe(csv_file='data/file_information.csv'):
'''Reads in a csv file which is assumed to have `File`, `Category` and `Task` columns.
This function does two things:
1) converts `Category` column values to numerical values
2) Adds a new, numerical `Class` label column.
The `Class` column will label plagiarized answers as 1 and non-plagiarized as 0.
Source texts have a special label, -1.
:param csv_file: The directory for the file_information.csv file
:return: A dataframe with numerical categories and a new `Class` label column'''
# your code here
dic_cat_class = {'non':0, 'heavy':1, 'light':2, 'cut':3, 'orig':-1}
plagiarism_df = pd.read_csv(csv_file)
classes = []
for i,value in enumerate(plagiarism_df['Category']):
plagiarism_df['Category'][i] = dic_cat_class[value]
if value == 'non':
classes.append(0)
else:
classes.append(1)
plagiarism_df['Class'] = classes
return plagiarism_df
# +
# informal testing, print out the results of a called function
# create new `transformed_df`
transformed_df = numerical_dataframe(csv_file ='data/file_information.csv')
# check work
# check that all categories of plagiarism have a class label = 1
transformed_df.head(10)
# +
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
import helpers
# create a text column
text_df = helpers.create_text_column(transformed_df)
text_df.head()
# +
# after running the cell above
# check out the processed text for a single file, by row index
row_idx = 0 # feel free to change this index
sample_text = text_df.iloc[0]['Text']
print('Sample processed text:\n\n', sample_text)
# +
random_seed = 1 # can change; set for reproducibility
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
import helpers
# create new df with Datatype (train, test, orig) column
# pass in `text_df` from above to create a complete dataframe, with all the information you need
complete_df = helpers.train_test_dataframe(text_df, random_seed=random_seed)
# check results
complete_df
# -
from sklearn.feature_extraction.text import CountVectorizer
text = {}
for line in range(0,complete_df.shape[0]):
if(complete_df.iloc[line]['File'] == 'g0pA_taska.txt'):
text['answer'] = complete_df.iloc[line]['Text']
if(complete_df.iloc[line]['File'] == 'orig_taska.txt'):
text['orig'] = complete_df.iloc[line]['Text']
n = 1
to_vec = []
to_vec.append(text['answer'])
to_vec.append(text['orig'])
vectorizer2 = CountVectorizer(analyzer='word', ngram_range=(n, n))
X2 = vectorizer2.fit_transform(to_vec)
intersec = 0
count = 0
for i in range(0, len(X2.toarray()[0])):
if(X2.toarray()[0][i] > 0 and X2.toarray()[1][i] > 0 ):
intersec = intersec + 1
if(X2.toarray()[0][i] > 0):
count = count + 1
print(count)
print(intersec)
intersec/count
A = "i think pagerank is a link analysis algorithm used by google that uses a system of weights attached to each element of a hyperlinked set of documents"
S = "pagerank is a link analysis algorithm used by the google internet search engine that assigns a numerical weighting to each element of a hyperlinked set of documents"
array_a = A.split(" ")
array_s = S.split(" ")
list_word = []
for i in range(0,len(array_a)):
for j in range(0, len(array_s)):
if(array_a[i] == array_s[j]):
list_word.append(array_a[i])
del array_s[j]
break
list_word
# Compute the normalized LCS given an answer text and a source text
def lcs_norm_word(answer_text, source_text):
'''Computes the longest common subsequence of words in two texts; returns a normalized value.
:param answer_text: The pre-processed text for an answer text
:param source_text: The pre-processed text for an answer's associated source text
:return: A normalized LCS value'''
array_a = answer_text.split(" ")
array_s = source_text.split(" ")
list_word = []
for i in range(0,len(array_a)):
for j in range(0, len(array_s)):
if(array_a[i] == array_s[j]):
list_word.append(array_a[i])
del array_s[j]
break
return len(list_word)/len(array_a)
# importing tests
import problem_unittests as tests
# +
df = complete_df.copy()
test_index = 10 # file 10
# get answer file text
answer_text = df.loc[test_index, 'Text']
# get text for orig file
# find the associated task type (one character, a-e)
task = df.loc[test_index, 'Task']
# we know that source texts have Class = -1
orig_rows = df[(df['Category'] == -1)]
orig_row = orig_rows[(orig_rows['Task'] == task)]
source_text = orig_row['Text'].values[0]
# +
# calculate LCS
test_val = lcs_norm_word(answer_text, source_text)
test_val
# tests.test_lcs(complete_df, lcs_norm_word)
# -
features_df = pd.read_csv('features.csv')
ind_train = complete_df[(complete_df['Datatype'] == 'train')].index
ind_test = complete_df[(complete_df['Datatype'] == 'test')].index
test_selection = list(features_df)[1:3] # first couple columns as a test
test_selection
len(complete_df.iloc[ind_train,3].values)
f_df = features_df.copy()
ind_column = [i for i, el in enumerate(features_df.columns) if el in test_selection]
f_df = features_df.iloc[ind_train,ind_column]
train_x = f_df.values
len(train_x)
task = features_df.loc[test_index, 'Task']
# Takes in dataframes and a list of selected features (column names)
# and returns (train_x, train_y), (test_x, test_y)
def train_test_data(complete_df, features_df, selected_features):
'''Gets selected training and test features from given dataframes, and
returns tuples for training and test features and their corresponding class labels.
:param complete_df: A dataframe with all of our processed text data, datatypes, and labels
:param features_df: A dataframe of all computed, similarity features
:param selected_features: An array of selected features that correspond to certain columns in `features_df`
:return: training and test features and labels: (train_x, train_y), (test_x, test_y)'''
ind_train = complete_df[(complete_df['Datatype'] == 'train')].index
ind_test = complete_df[(complete_df['Datatype'] == 'test')].index
# get the training features
f_df = features_df.copy()
ind_column = [i for i, el in enumerate(features_df.columns) if el in test_selection]
f_df = features_df.iloc[ind_train,ind_column]
train_x = f_df.values
# And training class labels (0 or 1)
train_y = complete_df.iloc[ind_train,3].values
# get the test features and labels
f_df = features_df.copy()
ind_column = [i for i, el in enumerate(features_df.columns) if el in test_selection]
f_df = features_df.iloc[ind_test,ind_column]
test_x = f_df.values
test_y = complete_df.iloc[ind_test,3].values
return (train_x, train_y), (test_x, test_y)
# +
(train_x, train_y), (test_x, test_y) = train_test_data(complete_df, features_df, test_selection)
# params: generated train/test data
tests.test_data_split(train_x, train_y, test_x, test_y)
# -
| .ipynb_checkpoints/1_Data_Exploration-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# coding: utf-8
# (c) 2016-02-12 <NAME>
import netCDF4
import shutil
import numpy as np
std_tmp = '/home/okada/Data/ob500_std_i_param_v1_NL1_{0:04d}.nc'
std_main = '/home/okada/Data/ob500_std_i_param_v1_NL1.nc'
std_zeros = '/home/okada/Data/ob500_std_i_zeros.nc'
grdfile = '/home/okada/Data/ob500_grd-11_3.nc'
vnames = ['temp', 'salt', 'NO3', 'NH4', 'chlorophyll', 'phytoplankton', 'zooplankton',
'LdetritusN', 'SdetritusN', 'oxygen', 'PO4', 'LdetritusP', 'SdetritusP']
shutil.copyfile(std_zeros, std_main)
main = netCDF4.Dataset(std_main, 'a')
for i in range(12):
stdfile = std_tmp.format(i+1)
nc = netCDF4.Dataset(stdfile, 'r')
for vname in vnames:
main[vname][i] = nc[vname][0]
nc.close()
main.close()
# -
| reduce_std.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %%
# code by <NAME> @graykode
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
def make_batch():
input_batch, target_batch = [], []
for seq in seq_data:
input = [word_dict[n] for n in seq[:-1]] # 'm', 'a' , 'k' is input
target = word_dict[seq[-1]] # 'e' is target
input_batch.append(np.eye(n_class)[input])
target_batch.append(target)
return input_batch, target_batch
class TextLSTM(nn.Module):
def __init__(self):
super(TextLSTM, self).__init__()
self.lstm = nn.LSTM(input_size=n_class, hidden_size=n_hidden)
self.W = nn.Linear(n_hidden, n_class, bias=False)
self.b = nn.Parameter(torch.ones([n_class]))
def forward(self, X):
input = X.transpose(0, 1) # X : [n_step, batch_size, n_class]
hidden_state = torch.zeros(1, len(X), n_hidden) # [num_layers(=1) * num_directions(=1), batch_size, n_hidden]
cell_state = torch.zeros(1, len(X), n_hidden) # [num_layers(=1) * num_directions(=1), batch_size, n_hidden]
outputs, (_, _) = self.lstm(input, (hidden_state, cell_state))
outputs = outputs[-1] # [batch_size, n_hidden]
model = self.W(outputs) + self.b # model : [batch_size, n_class]
return model
if __name__ == '__main__':
n_step = 3 # number of cells(= number of Step)
n_hidden = 128 # number of hidden units in one cell
char_arr = [c for c in 'abcdefghijklmnopqrstuvwxyz']
word_dict = {n: i for i, n in enumerate(char_arr)}
number_dict = {i: w for i, w in enumerate(char_arr)}
n_class = len(word_dict) # number of class(=number of vocab)
seq_data = ['make', 'need', 'coal', 'word', 'love', 'hate', 'live', 'home', 'hash', 'star']
model = TextLSTM()
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=0.001)
input_batch, target_batch = make_batch()
input_batch = torch.FloatTensor(input_batch)
target_batch = torch.LongTensor(target_batch)
# Training
for epoch in range(1000):
optimizer.zero_grad()
output = model(input_batch)
loss = criterion(output, target_batch)
if (epoch + 1) % 100 == 0:
print('Epoch:', '%04d' % (epoch + 1), 'cost =', '{:.6f}'.format(loss))
loss.backward()
optimizer.step()
inputs = [sen[:3] for sen in seq_data]
predict = model(input_batch).data.max(1, keepdim=True)[1]
print(inputs, '->', [number_dict[n.item()] for n in predict.squeeze()])
# -
| 3-2.TextLSTM/TextLSTM_pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Guide to Linear Regression in Python
#
# This notebook is designed to guide you in learning how to do a basic linear regression using Python. The notebook will be divided into two parts. In the first, we will create data with some known function. Then, we will add some random fluctuations to the data and see if we can recover the function. In the second part, we will use a public dataset to test this schema and perform the linear regression.
#
# This notebook will use a variety of packages.
#
# * [Numpy](https://numpy.org/doc/stable/reference/index.html) for array handling and basic numerical functions.
# * [SciKit-Learn](https://scikit-learn.org/stable/modules/classes.html) to perform the [linear regression](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html#sklearn.linear_model.LinearRegression) and provide [some data](https://scikit-learn.org/stable/modules/classes.html#module-sklearn.datasets)
# * [Matplotlib](https://matplotlib.org/stable/contents.html) for plotting and its simplified plotting module [Pyplot](https://matplotlib.org/stable/api/_as_gen/matplotlib.pyplot.html?highlight=pyplot#module-matplotlib.pyplot)
#
# ## Imports
#
# We will begin by *importing* the necessary packages below.
# +
# Import Numpy
import numpy as np # For arrays and basic numerical computation
# Import Packages from SciKit-Learn (sklearn)
# sklearn is such a large project that objects must be imported directly
from sklearn import datasets
from sklearn.linear_model import LinearRegression
# Import Matplotlib
from matplotlib import pyplot as plt
# -
# ## Part 1 - Known Functions
#
# ### Example - Velocity as a Function of Position
#
# Suppose we have a particle traveling along the $x$ axis and get the following data.
#
# | x (m) | v (m/s) |
# | ----- | ------- |
# | 0.00 | 3.04 |
# | 1.00 | 4.96 |
# | 2.00 | 7.19 |
# | 3.00 | 9.03 |
# | 4.00 | 10.84 |
# | 5.00 | 13.11 |
# | 6.00 | 15.39 |
# | 7.00 | 17.28 |
# | 8.00 | 18.79 |
# | 9.00 | 20.62 |
# | 10.00 | 22.81 |
#
# As it turns out, this data was simply generated by the equation
#
# \begin{equation}
# v(x) = 2 x + 3
# \end{equation}
#
# with some additional random, gaussian noise. Therefore, when we do linear regression on this data, we *should* recover a slope of 2 and an offset of 3.
#
# Normally, your data might be entered on a spreadsheet which could be ported with either [pandas](https://pandas.pydata.org/) or Google's gsheet plugin, but we will enter the data manually below.
#
# SciKit-Learn requires that data be entered as 2D arrays where the "observations" appear on each row and the "features" on each column. Therefore, we will cast our data into just such an array.
# +
# Store the Positions
# We will store this a 2D row vector first the transpose it to a column vector
# A 1D array is created with one set of brackets whereas a 2D row vector is
# stored with two brackets
# Note the `T` operator at the end of the parentheses. This is what is performing
# the transposition
# Although this is not incredibly relevant now, python/numpy assumes numerical data
# without decimals is integer data. Therefore, my dtype statement is telling
# numpy that the data should be treated as a float (number with decimals)
X = np.array([[0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10]], dtype='float64').T
# Store the Velocities
# This procedure matches the statement above
# Note that statements in parentheses can be broken over multiple lines
# This is called implicit line breaking
Y = np.array([[
3.04, 4.96, 7.19, 9.03, 10.84, 13.11, 15.39, 17.28, 18.79, 20.62, 22.81
]]).T
# -
# Now that we have stored our data in the variables `X` and `Y`, we can plot the scatter data before the regression to get a sense for what the data look like. Since this is a simple plot, we will use the pyplot [scatter](scatter) function. I'll also plot the function from which the data were generated with a simple [line plot](https://matplotlib.org/stable/api/_as_gen/matplotlib.pyplot.plot.html#matplotlib.pyplot.plot).
# +
# Plot the Sample Data as a scatter plot
# The scatter function takes the independent variable first and the dependent second
# The label keyword is for displaying the legend later
_ = plt.scatter(X, Y, label='Sample Data')
# Plot the Function
# I'll generate another set of variables to make this plot
xLine = np.arange(-2., 12) # Generates data from -2 to 11 with stepsize 1
yLine = 2*xLine + 3 # Create y from the x data and known function
_ = plt.plot(xLine, yLine, ':k', label='v(x)')
# Set the Limits for the axis
plt.xlim(-2, 11)
plt.ylim( 0, 25)
# Label Everything!
plt.xlabel('Position (m)')
plt.ylabel('Velocity (m/s)')
plt.title("Measuring a Particle's Velocity")
# Add the grid if wanted
# Uncomment the next line if you want the plot to be gridded
# plt.grid()
# Create the Legend
_ = plt.legend()
# -
# Now that we are comfortable with plotting, let's move on the the linear regression.
# +
# Create/Fit the Model
# sklearn is built around a framework of fitting data to models. Here, we are trying
# to fit our data to a linear model with linear regression.
# Below, we are creating a linear regression model, fitting the model to our data
# then we are storing that information in the variable named model
model = LinearRegression().fit(X, Y)
# Print out the fit
# Note that the slope parameter is stored as the model.coef_ and the
# intercept is stored as model.intercept_
# We store the equation in a string for future use
eqStr = 'v(x) = {:.2f} x + {:.2f}'.format(model.coef_.item(), model.intercept_.item())
print('Model Fit')
print('v(x) = m x + b')
print(eqStr)
# -
# As can be seen above, we were able to approximate the generating function based on the given data. Although the "random" fluctuations created a slight error in our coeficient and intercept, we got close to the predicted value.
#
# Now, let's recreate the plot above with our regression too.
# +
# Plot the Sample Data as a scatter plot
_ = plt.scatter(X, Y, label='Sample Data')
# Plot the Regression Model
# For this, we need an xLine that is a 2D column vector like above
# We will calculate the y values by predicting with our model
xLine = np.arange(-2., 12)[:, np.newaxis] # Generates data from -2 to 11 with stepsize 1
yLine = model.predict(xLine) # Create y from the x data and known function
_ = plt.plot(xLine, yLine, 'C1', label='Trendline')
# Add the equation string we printed above on the graph
_ = plt.text(-1, 15, eqStr)
# Plot the Function
yLine = 2*xLine + 3 # Create y from the x data and known function
_ = plt.plot(xLine, yLine, ':k', label='v(x)')
# Set the Limits for the axis
plt.xlim(-2, 11)
plt.ylim( 0, 25)
# Label Everything!
plt.xlabel('Position (m)')
plt.ylabel('Velocity (m/s)')
plt.title("Measuring a Particle's Velocity")
# Add the grid if wanted
# Uncomment the next line if you want the plot to be gridded
# plt.grid()
# Create the Legend
_ = plt.legend()
# -
# As you can see, the model fits the data well and almost perfectly recovers the generating function.
#
# ### Your Turn
#
# Now it's your turn, input the following data into the variables below and let my code from above take care of the rest.
#
# | x (m) | v (m/s) |
# | ----- | ------- |
# | 0.00 | 2.09 |
# | 2.00 | 7.91 |
# | 4.00 | 14.45 |
# | 6.00 | 20.07 |
# | 8.00 | 25.63 |
# | 10.00 | 32.25 |
# | 12.00 | 38.91 |
# | 14.00 | 44.66 |
# | 16.00 | 49.51 |
# | 18.00 | 55.11 |
# | 20.00 | 61.56 |
#
# +
# Store the Positions
X = np.array([[]], dtype='float64').T
# Store the Velocities
Y = np.array([[
]]).T
# Create/Fit the Model
model = LinearRegression().fit(X, Y)
# Print out the fit
eqStr = 'v(x) = {:.2f} x + {:.2f}'.format(model.coef_.item(), model.intercept_.item())
print('Model Fit')
print('v(x) = m x + b')
print(eqStr)
# Plot the Sample Data as a scatter plot
_ = plt.scatter(X, Y, label='Sample Data')
# Plot the Regression Model
xLine = np.arange(-2., 22)[:, np.newaxis] # Generates data from -2 to 11 with stepsize 1
yLine = model.predict(xLine) # Create y from the x data and known function
_ = plt.plot(xLine, yLine, 'C1', label='Trendline')
# Add the equation string we printed above on the graph
_ = plt.text(-1, 40, eqStr)
# Plot the Function
yLine = 3*xLine + 2 # Create y from the x data and known function
_ = plt.plot(xLine, yLine, ':k', label='v(x)')
# Set the Limits for the axis
plt.xlim(-2, 21)
plt.ylim( 0, 65)
# Label Everything!
plt.xlabel('Position (m)')
plt.ylabel('Velocity (m/s)')
plt.title("Measuring a Particle's Velocity")
# Add the grid if wanted
# Uncomment the next line if you want the plot to be gridded
# plt.grid()
# Create the Legend
_ = plt.legend()
| 04-ElementaryStatistics/LinearRegression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.3
# language: julia
# name: julia-1.0
# ---
# # Radiative recombination of initially helium-like ions
using JAC
#
# Here, we calculate the radiative recombination cross sections for initially helium-like and multiply-charged ions. For these ions, we consider the
#
#
# Therefore, let us first have a look how these settings are defined internally:
#
? PhotoRecombination.Settings
# # **The following part of this nootebook is still under construction.**
defaultSettings = PhotoRecombination.Settings()
#
# As seen from these settings, the default only includes the E1 photoionization amplitudes, the use of Coulomb gauge, and *no* photon energies are yet defined.
#
# We ....
#
setDefaults("unit: energy", "eV")
recSettings = PhotoRecombination.Settings()
#
# With these settings, we can proceed along standard lines and compute the cross section and $\beta_2$ parameters by:
#
| tutorials/56-compute-radiative-recombination-rates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas
from sklearn.metrics import f1_score
from sklearn.metrics import accuracy_score
import matplotlib.pyplot as plt; plt.rcdefaults()
import numpy as np
from sklearn import preprocessing
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn.preprocessing import LabelEncoder
from yellowbrick.text import TSNEVisualizer
# ## Load in Data
# +
DF_FL0 = pandas.read_excel('/Users/alice.naghshineh/Desktop/michael_data/Florida.xlsx')
DF_Dependent = DF_FL0[['Q12PresidentialVote']].copy()
DF_Independent = DF_FL0[[
'Party',
'Issue',
'Hispanic',
'Race',
'Marital Status',
'Education',
'Age',
'Gender',
'Raceparty',
'Genderparty']]
DF_FL0 = DF_FL0.drop(['Q8Clinton', 'Q9Trump'], axis=1)
# -
DF_FL0.head(10)
DF_FL0.describe()
# ## Visualize Feature Counts (Raw & Normalized)
# +
candidates = ('Clinton', 'Trump', 'Johnson', 'Stein', 'Undecided')
y_pos = np.arange(len(candidates))
counts = DF_FL0['Q12PresidentialVote'].value_counts().tolist()
plt.bar(y_pos, counts, align='center', alpha=0.5)
plt.xticks(y_pos, candidates)
plt.ylabel('Counts')
plt.title('Votes per Candidate')
plt.show()
# -
def get_counts_percent(column):
c = DF_FL0[column].value_counts()
p = DF_FL0[column].value_counts(normalize=True)
t = pandas.concat([c,p], axis=1, keys=['counts', '%'])
print('\n{}:\n{}'.format(column,t))
for column in DF_Independent:
get_counts_percent(column)
for column in ['Marital Status','Age','Gender']:
categories = DF_FL0[column].unique().tolist()
y_pos = np.arange(len(categories))
counts = DF_FL0[column].value_counts().tolist()
plt.bar(y_pos, counts, align='center', alpha=0.5)
plt.xticks(y_pos, categories)
plt.ylabel('Counts')
plt.title('{}'.format(column))
plt.show()
ax = sns.countplot(y='Party', hue='Race', data=DF_FL0)
ax = sns.countplot(y='Q12PresidentialVote', hue='Race', data=DF_FL0)
ax = sns.countplot(y='Race', hue='Issue', data=DF_FL0)
plt.legend(loc='lower right');
# ## Prepare Categorical Variables for Modeling
# +
DF_Independent_dummies = pandas.get_dummies(DF_Independent)
le = preprocessing.LabelEncoder()
le.fit(DF_Dependent['Q12PresidentialVote'])
target = le.transform(DF_Dependent['Q12PresidentialVote'])
# -
[x_train, x_test, y_train, y_test] = train_test_split(DF_Independent_dummies, target, test_size=0.2)
# ## Try KNeighbors
# +
knn = KNeighborsClassifier(n_neighbors = 1).fit(DF_Independent_dummies, target)
# accuracy on X_test
accuracyknn = knn.score(DF_Independent_dummies, target)
print(accuracyknn)
# -
knn = KNeighborsClassifier(n_neighbors = 1).fit(x_train, y_train)
predicted = knn.predict(x_test)
print("{}: {}".format(knn.__class__.__name__, f1_score(y_test, predicted, average='micro')))
# ## ... and a Clustering Visualization
# +
tsne = TSNEVisualizer(size=(1080, 720))
tsne.fit(DF_Independent_dummies, target)
print(tsne.poof())
| ADS-Spring2019/voting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Machine Learning Project
# This notebook presents a machine learning algorithm for the powerproduction dataset provided as part of the final project for Module 52954 "Machine Learning and Statistics", GMIT Higher Diploma in Computer Science. It applies a tensorflow learning algorithm to the dataset to allow prediction of power production based on wind speed.
# ***
# ## Import Modules
# +
#Import Modules
# Neural networks.
import tensorflow.keras as kr
# Numerical arrays
import numpy as np
# Data frames.
import pandas as pd
# Plotting
import matplotlib.pyplot as plt
# -
# ***
# ## Import dataset
#
# Read in data into pandas dataframe
df = pd.read_csv("powerproduction.csv")
# Sanity check on import
df.head()
df.tail()
# Basic descriptive stats on dataset
df.max()
df.min()
df.mean()
df.median()
# Quick plot of dataset to visualise relationship
plt.plot(df['speed'], df['power'])
plt.xlabel('Speed')
plt.ylabel('Power')
# ***
# ## Clean dataset
# On inspection, it appears that there's a sigmoid relationship between speed and power. It would also appear that there are datapoints with winds above 10, but no power production. This may be due to data entry error, recording error, or wind turbines facing perpendicular to the wind. For the sake of training a model, we will remove these points to clean up the dataset for training.
# +
#Remove power of zero where windspeed is above 10
df = df.drop(df[(df.speed > 10) & (df.power == 0)].index)
# Confirm removal (one was present in tail)
df.tail()
# -
# Quick visual confirmation that these datapoints have been removed
plt.plot(df['speed'], df['power'])
plt.xlabel('Speed')
plt.ylabel('Power')
# ***
# ## Prepare dataset for training and testing
# We'll split the set in preparation for training and testing, using an 80/20 split.
# +
# Extract 80% of the sample randomly for training
#Adapted from: https://www.geeksforgeeks.org/how-to-randomly-select-rows-from-pandas-dataframe/
train = df.sample(frac = 0.8)
# Sort into index order again
train.sort_index()
# +
# Use the remaining 20% of the dataset for testing
#https://stackoverflow.com/a/18360223
train_list = (train.index.values)
#train_list
# Sort into index order again
test = df.drop(train_list)
test.sort_index()
# -
# ***
# ## Train and fit model
#Train model
model = kr.models.Sequential()
model.add(kr.layers.Dense(50, input_shape=(1,), activation='sigmoid', kernel_initializer="glorot_uniform", bias_initializer="glorot_uniform"))
model.add(kr.layers.Dense(1, activation='linear', kernel_initializer="glorot_uniform", bias_initializer="glorot_uniform"))
model.compile(kr.optimizers.Adam(lr=0.001), loss='mean_squared_error')
# Fit the training data
model.fit(train['speed'], train['power'], epochs=500, batch_size=10)
# It would appear that the model 'bottoms-out' at a loss of around 15. This might be adequate for the purposes of this dataset. Let's see how it looks:
# Visualise how well the model fits the training data
plt.plot(train['speed'], train['power'], 'o', label='actual')
plt.plot(train['speed'], model.predict(train['speed']), 'x', label='prediction')
plt.xlabel('Speed')
plt.ylabel('Power')
plt.legend()
# The model fits the training data quite well. Next we will visualise if the test data set aligns with the overall dataset:
# Visualise the test dataset predictions against the actual dataset
plt.plot(df['speed'], df['power'], 'o', label='actual')
plt.plot(test['speed'], model.predict(test['speed']), 'x', label='prediction')
plt.xlabel('Speed')
plt.ylabel('Power')
plt.legend()
# The test data set matches the actual dataset closely. We can also quantify the 'best' loss for this model:
# Evaluate the neural network on the test data.
model.evaluate(test['speed'], test['power'])
# Finally, as a sanity check, we will run a few predictions and see if the predicted value makes 'sense':
print(f'The predicted power production with a wind-speed of 3.2 is {model.predict([3.2])}')
print(f'The predicted power production with a wind-speed of 14.7 is {model.predict([14.7])}')
print(f'The predicted power production with a wind-speed of 25.5 is {model.predict([25.5])}')
# These values appear sensical. The model is ready to deploy.
# ***
# # END
| .ipynb_checkpoints/Machine Learning & Statistics Final Project-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
msg=pd.read_csv('prog6_dataset.csv',names=['message','label']) #Tabular form data
print('Total instances in the dataset:',msg.shape[0])
msg['labelnum']=msg.label.map({'pos':1,'neg':0})
X=msg.message
Y=msg.labelnum
print('\nThe message and its label of first 5 instances are listed below')
X5, Y5 = X[0:5], msg.label[0:5]
for x, y in zip(X5,Y5):
print(x,',',y)
# Splitting the dataset into train and test data
from sklearn.model_selection import train_test_split
xtrain,xtest,ytrain,ytest=train_test_split(X,Y)
print('\nDataset is split into Training and Testing samples')
print('Total training instances :', xtrain.shape[0])
print('Total testing instances :', xtest.shape[0])
# Output of count vectoriser is a sparse matrix
# CountVectorizer - stands for 'feature extraction'
from sklearn.feature_extraction.text import CountVectorizer
count_vect = CountVectorizer()
xtrain_dtm = count_vect.fit_transform(xtrain) #Sparse matrix
xtest_dtm = count_vect.transform(xtest)
print('\nTotal features extracted using CountVectorizer:',xtrain_dtm.shape[1])
print('\nFeatures for first 5 training instances are listed below')
df=pd.DataFrame(xtrain_dtm.toarray(),columns=count_vect.get_feature_names())
print(df[0:5])#tabular representation
#print(xtrain_dtm) #Same as above but sparse matrix representation
# Training Naive Bayes (NB) classifier on training data.
from sklearn.naive_bayes import MultinomialNB
clf = MultinomialNB().fit(xtrain_dtm,ytrain)
predicted = clf.predict(xtest_dtm)
print('\nClassstification results of testing samples are given below')
for doc, p in zip(xtest, predicted):
pred = 'pos' if p==1 else 'neg'
print('%s -> %s ' % (doc, pred))
#printing accuracy metrics
from sklearn import metrics
print('\nAccuracy metrics')
print('Accuracy of the classifer is',metrics.accuracy_score(ytest,predicted))
print('Recall :',metrics.recall_score(ytest,predicted),
'\nPrecison :',metrics.precision_score(ytest,predicted))
print('Confusion matrix')
print(metrics.confusion_matrix(ytest,predicted))
# -
| .ipynb_checkpoints/Prog6-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Random Forest Classification with Normalizer
# This Code template is for the Classification tasks using a simple RandomForestClassifier based on the Ensemble Learning technique and feature rescaling technique Normalize. It is a meta estimator that fits multiple decision trees and uses averaging to improve the predictive accuracy and control over-fitting.
# ### Required Packages
import warnings
import numpy as np
import pandas as pd
import seaborn as se
import matplotlib.pyplot as plt
from sklearn.preprocessing import LabelEncoder,Normalizer
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import classification_report,plot_confusion_matrix
warnings.filterwarnings('ignore')
# ### Initialization
#
# Filepath of CSV file
#filepath
file_path= ""
# List of features which are required for model training .
#x_values
features=[]
# Target feature for prediction.
#y_value
target=''
# ### Data Fetching
#
# Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
#
# We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
df=pd.read_csv(file_path)
df.head()
# ### Feature Selections
#
# It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
#
# We will assign all the required input features to X and target/outcome to Y.
X = df[features]
Y = df[target]
# ### Data Preprocessing
#
# Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
#
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
def EncodeY(df):
if len(df.unique())<=2:
return df
else:
un_EncodedT=np.sort(pd.unique(df), axis=-1, kind='mergesort')
df=LabelEncoder().fit_transform(df)
EncodedT=[xi for xi in range(len(un_EncodedT))]
print("Encoded Target: {} to {}".format(un_EncodedT,EncodedT))
return df
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=EncodeY(NullClearner(Y))
X.head()
# #### Correlation Map
#
# In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
# #### Distribution Of Target Variable
plt.figure(figsize = (10,6))
se.countplot(Y)
# ### Data Splitting
#
# The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size = 0.2, random_state = 123)#performing datasplitting
# ### Feature Rescaling:
#
# Scale input vectors individually to unit norm (vector length).
# [For more information click here](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.normalize.html)
normalize = Normalizer()
X_train = normalize.fit_transform(X_train)
X_test = normalize.transform(X_test)
# ### Model
#
# A random forest is a meta estimator that fits a number of decision tree classifiers on various sub-samples of the dataset and uses averaging to improve the predictive accuracy and control over-fitting. The sub-sample size is controlled with the <code>max_samples</code> parameter if <code>bootstrap=True</code> (default), otherwise the whole dataset is used to build each tree.
#
# #### Model Tuning Parameters
#
# 1. n_estimators : int, default=100
# > The number of trees in the forest.
#
# 2. criterion : {“gini”, “entropy”}, default=”gini”
# > The function to measure the quality of a split. Supported criteria are “gini” for the Gini impurity and “entropy” for the information gain.
#
# 3. max_depth : int, default=None
# > The maximum depth of the tree.
#
# 4. max_features : {“auto”, “sqrt”, “log2”}, int or float, default=”auto”
# > The number of features to consider when looking for the best split:
#
# 5. bootstrap : bool, default=True
# > Whether bootstrap samples are used when building trees. If False, the whole dataset is used to build each tree.
#
# 6. oob_score : bool, default=False
# > Whether to use out-of-bag samples to estimate the generalization accuracy.
#
# 7. n_jobs : int, default=None
# > The number of jobs to run in parallel. fit, predict, decision_path and apply are all parallelized over the trees. <code>None</code> means 1 unless in a joblib.parallel_backend context. <code>-1</code> means using all processors. See Glossary for more details.
#
# 8. random_state : int, RandomState instance or None, default=None
# > Controls both the randomness of the bootstrapping of the samples used when building trees (if <code>bootstrap=True</code>) and the sampling of the features to consider when looking for the best split at each node (if <code>max_features < n_features</code>).
#
# 9. verbose : int, default=0
# > Controls the verbosity when fitting and predicting.
# Build Model here
model = RandomForestClassifier(n_jobs = -1,random_state = 123)
model.fit(X_train, y_train)
# #### Model Accuracy
#
# score() method return the mean accuracy on the given test data and labels.
#
# In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
print("Accuracy score {:.2f} %\n".format(model.score(X_test,y_test)*100))
# #### Confusion Matrix
#
# A confusion matrix is utilized to understand the performance of the classification model or algorithm in machine learning for a given test set where results are known.
plot_confusion_matrix(model,X_test,y_test,cmap=plt.cm.Blues)
# #### Classification Report
# A Classification report is used to measure the quality of predictions from a classification algorithm. How many predictions are True, how many are False.
#
# * **where**:
# - Precision:- Accuracy of positive predictions.
# - Recall:- Fraction of positives that were correctly identified.
# - f1-score:- percent of positive predictions were correct
# - support:- Support is the number of actual occurrences of the class in the specified dataset.
print(classification_report(y_test,model.predict(X_test)))
# #### Feature Importances.
#
# The Feature importance refers to techniques that assign a score to features based on how useful they are for making the prediction.
plt.figure(figsize=(8,6))
n_features = len(X.columns)
plt.barh(range(n_features), model.feature_importances_, align='center')
plt.yticks(np.arange(n_features), X.columns)
plt.xlabel("Feature importance")
plt.ylabel("Feature")
plt.ylim(-1, n_features)
# #### Creator: <NAME>, Github: [Profile]( https://github.com/neel-ntp)
| Classification/Random Forest/RandomForestClassifier_Normalize.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import toyplot
# ## Before
# +
canvas = toyplot.Canvas(width=300)
axes = canvas.cartesian(yshow=False)
x = [-4, -3, -2, -1, 0]
y = [4, 2, 3, 1, 0]
axes.plot(x, y, color=toyplot.color.black)
x = [1, 1, 1, 1, 1]
y = [4, 3, 2, 1, 0]
text = ["a", "b", "c", "d", "e"]
axes.text(x, y, text, color=toyplot.color.black);
# -
# ## After
# +
canvas = toyplot.Canvas(width=300)
axes = canvas.cartesian(yshow=False)
x = [-4, -3, -2, -1, 0]
y = [4, 2, 3, 1, 0]
axes.plot(x, y, color=toyplot.color.black)
x = [1, 1, 1, 1, 1]
y = [4, 3, 2, 1, 0]
text = ["a", "b", "c", "d", "e"]
axes.text(x, y, text, color=toyplot.color.black)
axes.x.ticks.locator = toyplot.locator.Explicit([-4, -3, -2, -1, 0])
| notebooks/tick-repellent.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Основы Pandas
# ### Загрузка данных в датафрейм
import pandas as pd
pd.read_html('http://www.cbr.ru')[3]
data = pd.read_csv('/Users/aleksandr/Dropbox/Python для работы с данными/2. Python для анализа данных/1. Python для анализа данных numpy и scipy/Python_5_pandas/power.csv')
type(data)
data.head()
# +
# если надо указать свои заголовки и разделитель
# data = pd.read_csv('power.csv', names = ['страна', 'год', 'количество', 'категория'], sep = '\t', header=0)
# data.head()
# +
# количество строк в датафрейме
len(data)
# +
# или так
data.shape
# -
# ### Упражнение
# Вам дана статистика продаж в файле transactions.csv. Вам необходимо загрузить этот файл в датафрейм и посчитать его размеры.
# ### Основные сведения о датафрейме
data.info()
# +
# немного статистики
data.describe()
# -
data.tail()
data['year'].head()
# +
# уникальные значения в столбце
data['category'].unique()
# +
# количество уникальных значений в столбце
len(data['category'].unique())
# +
# распределение количества строк по значениям столбца
data['category'].value_counts()
# +
# если надо в процентах
data['category'].value_counts(normalize=True)
# -
# ### Упражнение
# Используем файл transactions.csv. Определите какой товар (столбец Product) упоминается в файле чаще всего?
# # Фильтры
data = pd.read_csv('/Users/aleksandr/Dropbox/Python для работы с данными/2. Python для анализа данных/1. Python для анализа данных numpy и scipy/Python_5_pandas/power.csv')
data.head()
# +
# выбрать несколько столбцов
country_stats = data.filter(items = ['country', 'quantity'])
country_stats.head()
# -
average_level = data['quantity'].mean()
average_level
'quantity > {}'.format(average_level)
# +
# строки с потреблением больше среднего
average_level = data['quantity'].mean()
country_stats.query('quantity > {}'.format(average_level)).head()
# +
# самый популярный способ
data[ data.quantity > average_level ].head()
# +
# фильтр на подстроку
# найдем как называется Россия в этом датафрейме
data[ data['country'].str.contains('us', case=False) ]['country'].unique()
# +
# фильтр на несколько условий сразу
# | - условие ИЛИ
# & AND
# () | (() | () & ())
filtered_countries = data[ (data['country']=='Russian Federation') | (data['country']=='Belarus') ]
filtered_countries.head()
# -
filtered_countries['country'].unique()
data[ ['country', 'quantity'] ].head()
# +
# фильтры на строки
data.loc[1000:1005]
# -
# # Сортировка
# +
# Сортировка по столбцу
data.sort_values(by='quantity').head()
# +
# сортировка по убыванию
data.sort_values('quantity', ascending=False).head()
# +
# сортировка по нескольким столбцам
data.sort_values(by=['country', 'year', 'quantity'], ascending=[False, True, False]).head(50)
# -
data.sort_values('country', ascending=True, inplace=True)
# +
# параметр inplace
data = data.sort_values(by=['country', 'year', 'quantity'], ascending=[True, True, False])
# чтобы сократить это выражение используем inplace:
data.sort_values(by=['country', 'year', 'quantity'], ascending=[True, True, False], inplace=True)
# -
# ### Упражнение
# Используем transactions.csv.
#
# Для какой транзакции (столбец ID) были наибольшие расходы (столбец Cost) в категории "_8" (столбец Product)?
| Lectures notebooks/(Lectures notebooks) netology Mathematics and Python/5. Python_5_numpy_scipy/Python_5_pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# n=b
# # Print Pattern
# - Problem Code: [PPATTERN](https://www.codechef.com/problems/PPATTERN)
for _ in range(int(input())):
n=int(input())
a=[[None]*n for i in range(n)]
k=1
for i in range(n):
for j in range(i+1):
a[j][i-j]=k
k+=1
for i in range(1,n):
for j in range(n-i):
a[i+j][n-1-j]=k
k+=1
#display
for i in range(n):
for j in range(n):
print(a[i][j],end=' ')
print()
a
a[0][0]=1
a
a[1][0]=2
a
| Codechef/Practise/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Analyzer and mappings to enable shingles (bigrams and trigrams) for clinical trials, instead of allowing them to be auto-created. Then index as usual.
# **PUT** *{{elasticsearch}}*/trials
# ```json
# {
# "settings": {
# "number_of_shards": 5,
# "analysis": {
# "filter": {
# "my_shingle_filter": {
# "type": "shingle",
# "min_shingle_size": 2,
# "max_shingle_size": 3,
# "output_unigrams": false
# }
# },
# "analyzer": {
# "my_shingle_analyzer": {
# "type": "custom",
# "tokenizer": "standard",
# "filter": [
# "lowercase",
# "my_shingle_filter"
# ]
# }
# }
# }
# }
# }
# ```
# **PUT** *{{elasticsearch}}*/trials/_mapping/trials
#
# ```json
# {
# "trials": {
# "properties": {
# "exclusion": {
# "type": "string",
# "fields": {
# "shingles": {
# "type": "string",
# "analyzer": "my_shingle_analyzer"
# }
# }
# },
# "id": {
# "type": "text",
# "fields": {
# "keyword": {
# "type": "keyword",
# "ignore_above": 256
# }
# }
# },
# "inclusion": {
# "type": "string",
# "fields": {
# "shingles": {
# "type": "string",
# "analyzer": "my_shingle_analyzer"
# }
# }
# },
# "maximum_age": {
# "type": "long"
# },
# "minimum_age": {
# "type": "long"
# },
# "sex": {
# "type": "text",
# "fields": {
# "keyword": {
# "type": "keyword",
# "ignore_above": 256
# }
# }
# },
# "summary": {
# "type": "string",
# "fields": {
# "shingles": {
# "type": "string",
# "analyzer": "my_shingle_analyzer"
# }
# }
# },
# "title":{
# "type": "string",
# "fields": {
# "shingles": {
# "type": "string",
# "analyzer": "my_shingle_analyzer"
# }
# }
# }
# }
# }
# }
# ```
| python/trials/trials_elastic_config.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
import osmnx as ox, networkx as nx, pandas as pd, geopandas as gpd, time, matplotlib.pyplot as plt, math, ast, re
import matplotlib.cm as cm
from matplotlib.collections import PatchCollection
from descartes import PolygonPatch
from shapely.geometry import Point, Polygon, MultiPolygon
import statsmodels.api as sm, numpy as np
from geopy.distance import great_circle
from shapely.geometry import Polygon
# %matplotlib inline
ox.config(use_cache=True, log_file=True, log_console=True, log_filename='analyze_stats_every_us_city',
data_folder='G:/Geoff/osmnx/cities-usa', cache_folder='G:/Geoff/osmnx/cache/cities-usa')
df = pd.read_csv('stats_every_city.csv', encoding='utf-8')
df.head()
# ## Inspect the stats
len(df)
df.describe()
df[['state', 'city', 'time', 'streets_per_node_avg', 'n', 'm', 'circuity_avg']].sort_values(by='m', ascending=False).head()
df.groupby('state').median().describe()
df['streets_per_node_avg'].hist(bins=50)
df_plot = df.dropna(subset=['n'])
# +
fig, ax = plt.subplots(figsize=(6, 6))
x = df_plot['n'] / 1000
y = df_plot['street_length_total'] / 1000 / 1000
xlim = [0, math.ceil(x.max()/10)*10] #round to nearest greater 100
ylim = [0, math.ceil(y.max()/2)*2] #round to nearest greater 20
# regress y on x
results = sm.OLS(y, sm.add_constant(x)).fit()
# calculate estimated y values for regression line
x_line = pd.Series(xlim)
y_est = x_line * results.params[1] + results.params[0]
# draw regression line and scatter plot the points
ax.plot(x_line, y_est, c='k', alpha=0.7, linewidth=1, zorder=1)
ax.scatter(x=x, y=y, c='#336699', linewidths=0, alpha=0.7, s=10)
ax.set_xlim(xlim)
ax.set_ylim(ylim)
#ax.set_xlim((0,15000))
#ax.set_ylim((0,3000))
ax.set_title('Total street length vs number of nodes')
ax.set_xlabel('Node count, thousands')
ax.set_ylabel('Total street length (km, thousands)')
fig.savefig(filename='images/street-length-vs-nodes', dpi=300, bbox_inches='tight')
plt.show()
# -
print('r-squared:', round(results.rsquared, 2))
results.params
cols = ['area_km', 'avg_neighbor_degree_avg',
'avg_weighted_neighbor_degree_avg', 'circuity_avg',
'clustering_coefficient_avg', 'clustering_coefficient_weighted_avg',
'count_intersections', 'degree_centrality_avg', 'edge_density_km',
'edge_length_avg', 'edge_length_total', 'geoid',
'int_1_streets_prop', 'int_3_streets_prop', 'int_4_streets_prop', 'intersection_density_km',
'k_avg', 'm', 'n', 'city', 'node_density_km', 'pagerank_max',
'pagerank_min',
'self_loop_proportion', 'street_density_km', 'street_length_avg',
'street_length_total', 'street_segments_count', 'streets_per_node_avg', 'city']
summary = df[cols]
summary = summary.describe().T.drop(['count', '25%', '75%'], axis=1).applymap(lambda x: round(x, 3))
summary.to_csv('summary.csv', encoding='utf-8')
df.groupby('state')['streets_per_node_avg'].median().sort_values()
df.columns
# node and edge density per km are hard to interpret when using municipal boundaries
# as municipal boundaries vary greatly in their extents around the built-up area
by_state = df.groupby('state')[['intersection_density_km',
'streets_per_node_avg',
'circuity_avg',
'street_length_avg']].median().sort_values(by='intersection_density_km').applymap(lambda x: round(x, 2))
by_state.sort_index().to_csv('by_state.csv', encoding='utf-8')
by_state.sort_index()
variable = 'streets_per_node_avg'
df[df['state']=='NE'][[variable, 'city']].sort_values(by=variable)
# ## Distribution of street segment lengths
#
# Tends to follow approx lognormal distribution in most cities with a variety of streets. Show one example here.
query = 'Oakland, CA, USA'
G = ox.graph_from_place(query, network_type='drive')
lengths = [data['length'] for u, v, key, data in G.edges(keys=True, data=True)]
ax = pd.Series(lengths).hist(bins=100, fc='k', alpha=0.6)
ax.set_xlim((0, 1000))
ax.set_xlabel('Street segment length (m)')
ax.set_ylabel('Count')
ax.set_title('Distribution of street segment lengths\n{}'.format(query))
plt.show()
| cities-usa/analyze_stats_every_us_city.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to add new gauge-optimizations to GST results
# This example demonstrates how to take a previously computed `Results` object and add new gauge-optimized version of to one of the estimates. First, let's "pre-compute" a `ModelEstimateResults` object using `StandardGST`, which contains a single `Estimate` called "TP":
import pygsti
from pygsti.modelpacks import smq1Q_XYI
# +
#Generate some fake data and run GST on it.
exp_design = smq1Q_XYI.get_gst_experiment_design(max_max_length=4)
mdl_datagen = smq1Q_XYI.target_model().depolarize(op_noise=0.1, spam_noise=0.001)
ds = pygsti.construction.generate_fake_data(mdl_datagen, exp_design.all_circuits_needing_data, nSamples=1000, seed=1234)
data = pygsti.protocols.ProtocolData(exp_design, ds)
gst = pygsti.protocols.StandardGST("TP", gaugeopt_suite={'go0': {'itemWeights': {'gates': 1, 'spam': 1}}})
results = gst.run(data)
results.write("example_files/regaugeopt_example")
# -
# Next, let's load in the pre-computed results and use the `add_gauge_optimization` method of the `pygsti.objects.Estimate` object to add a new gauge-optimized version of the (gauge un-fixed) model estimate stored in `my_results.estimates['default']`. The first argument of `add_gauge_optimization` is just a dictionary of arguments to `pygsti.gaugeopt_to_target` **except** that you don't need to specify the `Model` to gauge optimize or the target `Model` (just like the `gaugeOptParams` argument of `do_long_sequence_gst`). The optional "`label`" argument defines the key name for the gauge-optimized `Model` and the corresponding parameter dictionary within the `Estimate`'s `.models` and `.goparameters` dictionaries, respectively.
my_results = pygsti.io.load_results_from_dir("example_files/regaugeopt_example", name="StandardGST")
# +
estimate = my_results.estimates['TP']
estimate.add_gaugeoptimized( {'itemWeights': {'gates': 1, 'spam': 0.001}}, label="Spam 1e-3" )
mdl_gaugeopt = estimate.models['Spam 1e-3']
print(list(estimate.goparameters.keys())) # 'go0' is the default gauge-optimization label
print(mdl_gaugeopt.frobeniusdist(estimate.models['target']))
# -
# One can also perform the gauge optimization separately and specify it using the `model` argument (this is useful when you want or need to compute the gauge optimization elsewhere):
mdl_unfixed = estimate.models['final iteration estimate']
mdl_gaugefixed = pygsti.gaugeopt_to_target(mdl_unfixed, estimate.models['target'], {'gates': 1, 'spam': 0.001})
estimate.add_gaugeoptimized( {'any': "dictionary",
"doesn't really": "matter",
"but could be useful it you put gaugeopt params": 'here'},
model=mdl_gaugefixed, label="Spam 1e-3 custom" )
print(list(estimate.goparameters.keys()))
print(estimate.models['Spam 1e-3 custom'].frobeniusdist(estimate.models['Spam 1e-3']))
# You can look at the gauge optimization parameters using `.goparameters`:
import pprint
pp = pprint.PrettyPrinter()
pp.pprint(dict(estimate.goparameters['Spam 1e-3']))
# Finally, note that if, in the original creation of `StandardGST`, you set **`gaugeopt_suite=None`** then no gauge optimizations are performed (there would be no "`go0`" elements) and you start with a blank slate to perform whatever gauge optimizations you want on your own.
| jupyter_notebooks/Examples/GOpt-AddingNewOptimizations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: enu
# kernelspec:
# display_name: ENU
# language: python
# name: enu
# ---
# # Generates tables for the manuscript
#
# **Authored by:** <NAME>, <NAME> and <NAME>
#
# This notebook produces the tables for the main manuscript and the supplementary material. It assumes the directory containing the manuscript latex is named `mutation_classifier_manuscript` and is a sister directory to the one containing this file.
#
# The notebook further assumes installation of numerous python libraries. Those are defined in the top level imports below.
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
import os
import re
import textwrap
import numpy
from tqdm import tqdm
from cogent3.util.misc import open_
from cogent3 import LoadTable
from ms_scripts.makefig import MakeMatch
from ms_scripts.getrefs import get_ms_supp_labels
from ms_scripts.maketab import (classifier_summary_stats, format_latex_table,
format_positions, format_pvalue, format_direction,
format_group)
os.makedirs('figs_n_tabs', exist_ok=True)
kwargs = dict(category="name", order=["M", "M+I", "M+I+2D", "M+I+2Dp", "FS"])
def clean_latex(latex):
latex = latex.splitlines()
result = []
for line in latex:
if line.rstrip().endswith(r'\\'):
result.append(line)
continue
line = line.strip()
result.extend(textwrap.wrap(line.strip(), break_long_words=False, break_on_hyphens=False))
latex = '\n'.join(result)
return latex
def get_relative_dir(path):
relative_dir = os.path.basename(os.path.dirname(path))
return relative_dir
def get_summary_stats(table, stat, k):
matcher = MakeMatch({0: lambda x: x == "lr",
1: lambda x: x == k,
2: lambda x: x in kwargs["order"]})
selected = collated.filtered(matcher, columns=["algorithm", "k", "name"])
t = classifier_summary_stats(selected, 'auc', ['name', 'size'])
return t
def is_tab(val):
bits = val.split(':')
result = bits[0].endswith('tab')
return result
# +
outdir_ms = "../mutation_classifier_manuscript"
all_ms_tables = {}
all_supp_tables = {}
all_floats = {} # for storing all latex float text (figs, tables)
unused_labels = []
ms_labels, supp_labels = get_ms_supp_labels(is_tab, texdir=outdir_ms)
# -
# ## Sample sizes
label = "suptab:sample-sizes"
if label in supp_labels:
chroms = [1, 2, 3, 4, 5, 6, 7, 8, 9,10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 'XY']
enu_size_dict = {1: 16977, 2: 21100, 3: 11228, 4: 13973, 5: 14509, 6: 13039, 7: 20864, 8: 11232,
9: 14010, 10: 11315, 11: 17101, 12: 8022, 13: 9085, 14: 8395, 15: 9342, 16: 7266,
17: 11981, 18: 6356, 19: 7529, 'XY': 853}
sp_sizes_dict = {1: 17848, 2: 20051, 3: 11713, 4: 16936, 5: 16028, 6: 12097, 7: 19161, 8: 13465,
9: 15662, 10: 12641, 11: 19626, 12: 8817, 13: 8939, 14: 8868, 15: 11079, 16: 8117,
17: 12168, 18: 7732, 19: 8635, 'XY': 5097}
rows = []
for chrom in chroms:
num_enu = enu_size_dict[chrom]
num_sp = sp_sizes_dict[chrom]
rows.append([chrom, num_enu, num_sp])
header = ['Chromosome', 'ENU-induced', 'Spontaneous']
all_size_table = LoadTable(header=header, rows=rows, column_templates={"ENU-induced": "{:,}".format, "Spontaneous": "{:,}".format})
all_size_table.title = r"By-chromosome sample sizes of genetic variants from the ENU induced and spontaneous "\
+r"germline mutations."
all_supp_tables[label] = format_latex_table(all_size_table, justify="rrrl", label=label)
all_size_table
else:
unused_labels.append(label)
# # Log-linear
# ### For manuscript
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
label = "tab:enu_v_germline:a-g"
if label in ms_labels:
# fns = !ls loglin/results/ENU_vs_germline/autosomes/directions/AtoG/summary.txt
fns
tab_enu_v_sp = LoadTable(fns[0], sep="\t")
tab_enu_v_sp = tab_enu_v_sp.with_new_column("Position(s)", format_positions, columns=["Position"])
tab_enu_v_sp = tab_enu_v_sp.get_columns(["Position(s)", "Deviance", "df", "prob"])
tab_enu_v_sp = tab_enu_v_sp.with_new_header("prob", "p-value")
tab_enu_v_sp.format_column("p-value", format_pvalue)
tab_enu_v_sp.format_column("Deviance", "%.1f")
tab_enu_v_sp = tab_enu_v_sp.sorted(columns=["df", "Deviance"])
tab_enu_v_sp.title = r"Log-linear analysis of mutation motif comparison between mouse germline and ENU-induced "\
+r"A$\rightarrow$G mutations. Deviance is from the log-linear model, with df degrees-of-freedom "\
+r"and corresponding $p$-value obtained from the $\chi^2$ distribution."
all_ms_tables[label] = format_latex_table(tab_enu_v_sp, justify="rrrl", label=label)
print(tab_enu_v_sp)
else:
unused_labels.append(labels)
# -
# # For supplementary
# ## Log-linear
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
label = "suptab:spectra:enu_spontaneous"
if label in supp_labels:
# fns = !ls loglin/results/ENU_vs_germline/autosomes/combined/spectra_summary.txt
tab_spectra = LoadTable(fns[0], sep="\t")
tab_spectra = tab_spectra.get_columns(["direction", "group", "ret"])
tab_spectra = tab_spectra.with_new_header("direction", "Direction")
tab_spectra = tab_spectra.with_new_header("group", "Class")
tab_spectra = tab_spectra.with_new_header("ret", "RET")
tab_spectra.format_column("Direction", format_direction)
tab_spectra.format_column("Class", format_group)
tab_spectra.format_column("RET", "%.3f")
tab_spectra = tab_spectra.sorted(columns=["RET"])
tab_spectra.title = r"Comparison of mutation spectra between Spontaneous and ENU-induced "\
+r"germline point mutations. RET values are proportional to deviance generated from the log-linear model \citep{zhu2017statistical}, and "\
+r"$p$-value are obtained from the $\chi^2$ distribution. All $p$-values were below the limit of detection."
all_supp_tables[label] = format_latex_table(tab_spectra, justify="rrrl", label=label)
tab_spectra
else:
unused_labels.append(labels)
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
label = "suptab:a-g:enu"
if label in supp_labels:
# fns = !ls loglin/results/ENU_variants/autosomes/directions/AtoG/*.txt
tab_enu = LoadTable(fns[0], sep="\t")
tab_enu = tab_enu.with_new_column("Position(s)", format_positions, columns=["Position"])
tab_enu = tab_enu.get_columns(["Position(s)", "Deviance", "df", "prob"])
tab_enu = tab_enu.with_new_header("prob", "p-value")
tab_enu.format_column("p-value", format_pvalue)
tab_enu.format_column("Deviance", "%.1f")
tab_enu = tab_enu.sorted(columns=["df", "Deviance"])
tab_enu.title = r"Log-linear analysis of ENU-induced A$\rightarrow$G mutation. "\
+r"Position(s) are relative to the index position. Deviance is from the log-linear model, "\
+r"with df degrees-of-freedom and corresponding $p$-value obtained from the $\chi^2$ "\
+r"distribution. $p$-values listed as 0.0 are below the limit of detection. "\
+r"See \citet{zhu2017statistical} for a more detailed description of the log-linear models."
all_supp_tables[label] = format_latex_table(tab_enu, justify="rrrl", label=label)
tab_enu
else:
unused_labels.append(label)
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
label = "suptab:p_sum_tab"
if label in supp_labels:
# fns = !ls loglin/results/ENU_vs_germline/autosomes/directions/*/summary.txt
p_sum_rows = []
for fn in fns:
start, end = fn.split('/')[-2].split('to')
mut_dir = start + r'$\rightarrow$' + end
summary_tab = LoadTable(fn, sep="\t")
p_vals = summary_tab.get_columns(["Position", "prob"]).tolist()
first_p_vals = []
second_p_vals = []
third_p_vals = []
forth_p_vals = []
for record in p_vals:
poses = record[0]
order = poses.count(':') + 1
if order == 1:
first_p_vals.append(record[1])
if order == 2:
second_p_vals.append(record[1])
if order == 3:
third_p_vals.append(record[1])
if order == 4:
forth_p_vals.append(record[1])
first_p_num = sum(p < 0.05 for p in first_p_vals)
second_p_num = sum(p < 0.05 for p in second_p_vals)
third_p_num = sum(p < 0.05 for p in third_p_vals)
forth_p_num = sum(p < 0.05 for p in forth_p_vals)
p_sum_rows.append([mut_dir, first_p_num, second_p_num, third_p_num, forth_p_num])
p_sum_header = ["Mutation direction", "1st-order", "2nd-order", "3rd-order", "4th-order"]
p_sum_tab = LoadTable(header=p_sum_header, rows=p_sum_rows, sep='\t')
p_sum_tab.title = r"Number of positions showing significant differences between ENU-induced and "\
+r"spontaneous germline point mutations from analysis of 5-mers. A $p$-value $\le 0.05$ was classified as significant. "\
+r"$p$-values were from the log-linear analysis."
all_supp_tables[label] = format_latex_table(p_sum_tab, justify="ccccc", label=label)
p_sum_tab
else:
unused_labels.append(label)
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
def convert_to_subtable(table):
table = table.splitlines()
table[0] = r"\begin{subtable}[t]{1.0\textwidth}"
table[-1] = r"\end{subtable}"
table = "\n".join(table)
return table
label = "suptab:long-flank"
if label in supp_labels:
header = ['Direction', 'RE$_{max}(1)$', 'RE Dist.', 'p-val Dist.']
rows =[['A$\\rightarrow$C', '0.0374', '6', '10'],
['A$\\rightarrow$G', '0.0402', '4', '10'],
['A$\\rightarrow$T', '0.0638', '2', '10'],
['C$\\rightarrow$A', '0.0632', '2', '10'],
['C$\\rightarrow$T', '0.0703', '2', '10'],
['G$\\rightarrow$A', '0.0710', '2', '10'],
['G$\\rightarrow$T', '0.0624', '2', '10'],
['T$\\rightarrow$A', '0.0606', '2', '10'],
['T$\\rightarrow$C', '0.0395', '4', '10'],
['T$\\rightarrow$G', '0.0373', '6', '10']]
lflank_enu = LoadTable(header=header, rows=rows, title="ENU-induced")
lflank_enu = format_latex_table(lflank_enu, "rrrc", label="suptab:long-flank-nbrsize:enu")
lflank_enu = convert_to_subtable(lflank_enu)
d = r"""Direction,RE$_{max}(1)$,RE Dist.,p-val Dist.
A$\rightarrow$C,0.0047,8,10
A$\rightarrow$G,0.0118,3,10
A$\rightarrow$T,0.0194,3,10
C$\rightarrow$A,0.0332,4,10
C$\rightarrow$T,0.0505,1,10
G$\rightarrow$A,0.0508,1,10
G$\rightarrow$T,0.0351,3,10
T$\rightarrow$A,0.0117,2,10
T$\rightarrow$C,0.0152,2,10
T$\rightarrow$G,0.0148,2,10""".splitlines()
header = d.pop(0).split(",")
rows = [r.split(",") for r in d]
lflank_spontab = LoadTable(header=header, rows=rows, title="Spontaneous")
lflank_spon = format_latex_table(lflank_spontab, "rrrc")
lflank_spon = convert_to_subtable(lflank_spon)
lflank_tmp = '\n'.join([r"\begin{table}",
r"\centering", "", "%s", "", "%s",
r"\caption{Longer range neighbourhood effect log-linear analyses results of (a) ENU-induced "
"mutations and (b) germline spontaneous mutations. For both subtables, the most distant "
"positions from the mutation with RE$(1)\ge10\%%$ of RE$_{max}(1)$. RE$(1)$ is the"
" first order RE for the position, and RE$_{max}(1)$ the largest RE from a first "
"order effect for the surveyed positions. RE Dist. is the furthest position with "
"an RE value $\ge 0.1\times\mathrm{RE}_{max}$. p-val Dist. is the corresponding"
" distance based on the $p$-value$\le 0.05$. As the analysis was limited to "
"a flank size of 10bp either side of the mutating base, the maximum possible distance is 10.}",
r"\label{%s}",
r"\end{table}"])
all_supp_tables[label] = lflank_tmp % (lflank_enu, lflank_spon, label)
lflank_spontab
else:
unused_labels.append(label)
# -
# ## Data properties
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
auto_xy = lambda x: re.search(r'chrom([0-9]{1,2}|XY)\.', x)
get_chrom = lambda x: re.findall(r'(?<=chrom).{1,2}(?=\.)', x)[0]
def get_chrom_paths(paths):
result = {}
for path in paths:
if not auto_xy(path):
continue
chrom = get_chrom(path)
try:
chrom = int(chrom)
except ValueError:
pass
result[chrom] = path
return result
def get_num_records(path):
data = LoadTable(path)
return data.shape[0]
# enu_fns = !ls ../variant_data/ENU/*.tsv.gz
enu_fns = get_chrom_paths(enu_fns)
# spn_fns = !ls ../variant_data/Germline/*.tsv.gz
spn_fns = get_chrom_paths(spn_fns)
rows = []
for chrom in enu_fns:
enu_count = get_num_records(enu_fns[chrom])
spn_count = get_num_records(spn_fns[chrom])
rows.append([chrom, enu_count, spn_count])
rows = sorted(rows, key=lambda x: ({True: 100}.get(type(x[0]) == str, x[0]), x))
data_sizes = LoadTable(header=['Chromosome', 'ENU-induced', 'Spontaneous'], rows=rows)
data_sizes.format_column('ENU-induced', '{:,}'.format)
data_sizes.format_column('Spontaneous', '{:,}'.format)
data_sizes.title = 'By-chromosome sample sizes of genetic variants from the ENU induced and spon-taneous germline mutations.'
# -
# # Classifier
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
collated = LoadTable("classifier/chrom1_train/collated/collated.tsv.gz",
static_column_types=True)
collated = collated.with_new_column('k', lambda x: 2 * x + 1, columns='flank_size')
stat = 'auc'
columns = ['algorithm', stat, 'k', 'name', 'size', 'flank_size', 'feature_dim', 'usegc', 'proximal']
collated = collated.get_columns(columns)
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
three = get_summary_stats(collated, 'auc', 3)
three.title = "Summary of AUC scores from LR classifiers using 3-mers."
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
five = get_summary_stats(collated, 'auc', 5)
five.title = "Summary of AUC scores from LR classifiers using 5-mers."
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
seven = get_summary_stats(collated, 'auc', 7)
seven.title = "Summary of AUC scores from LR classifiers using 7-mers."
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
fifty_nine = get_summary_stats(collated, 'auc', 59)
fifty_nine.title = "Summary of AUC scores from LR classifiers using 59-mers."
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
tables = []
for t, l in [(three, 'suptab:LR_aucs_3mer'), (five, 'suptab:LR_aucs_5mer'),
(seven, 'suptab:LR_aucs_7mer'),
(fifty_nine, 'suptab:LR_aucs_59mer')]:
all_supp_tables[l] = format_latex_table(t, 'rrcccc', label=l)
# + [markdown] inputHidden=false outputHidden=false
# # Writing ms tables
# -
for label in ms_labels:
if label not in all_ms_tables:
print('ms label missing', label)
continue
opath = os.path.join(outdir_ms, label.replace(':', '-') + '.tex')
print(opath)
with open(opath, 'w') as outfile:
table = all_ms_tables[label]
outfile.write(clean_latex(table) + '\n\n\n')
# # Writing supp tables
with open(os.path.join(outdir_ms, 'sup_tables.tex'), 'w') as outfile:
for label in supp_labels:
if label not in all_supp_tables:
print('ms label missing', label)
continue
table = all_supp_tables[label]
outfile.write(clean_latex(table) + '\n\n\n')
| make_manuscript_tables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Kód k vylepšení
# +
import ai
import utils
from random import randrange
def vyhodnot(pole):
# Funkce vezme hrací pole a vrátí výsledek
# na základě aktuálního stavu hry
if "xxx" in pole: #Vyhrál hráč s křížky
return "x"
elif "ooo" in pole: #Vyhrál hráč s kolečky.
return "o"
elif "-" not in pole: #Nikdo nevyhrál
return "!"
else: #Hra ještě neskončila.
return "-"
def tah_pocitace(pole):
"Počítač vybere pozici, na kterou hrát, a vrátí herní pole se zaznamenaným tahem počítače"
delka=len(pole)
while True:
pozice=randrange(1,delka-1)
if "-" in pole[pozice]:
if "o" in pole[pozice+1] or "o" in pole[pozice-1] or "x" in pole[pozice+1] or "x" in pole[pozice-1]: #počítač hraje strategicky
return pozice
# -
# ## Vylepšená verze
# +
from random import randrange
def vyhodnot(pole):
"""
Funkce vezme hrací pole a vrátí výsledek
na základě aktuálního stavu hry
"""
if "xxx" in pole: #Vyhrál hráč s křížky
return "x"
elif "ooo" in pole: #Vyhrál hráč s kolečky.
return "o"
elif "-" not in pole: #Nikdo nevyhrál
return "!"
else: #Hra ještě neskončila.
return "-"
def tah_pocitace(pole):
"""
Počítač vybere pozici, na kterou hrát,
a vrátí ideální pozici k tahu
"""
delka = len(pole)
while True:
pozice = randrange(1, delka - 1)
if "-" in pole[pozice]:
if "o" in pole[pozice + 1] or "o" in pole[pozice - 1] or \
"x" in pole[pozice + 1] or "x" in pole[pozice - 1]: #počítač hraje strategicky
return pozice
# -
# ## Co je tady navíc?
pozice = int(randrange(len(pole)))
# ## Obecné rady a doporučení
# - Importované moduly je třeba použít nebo jejich import smazat. V kódu je pak větší pořádek.
# - Mezi importy a zbytek kódu je pro přehlednost dobré vložit alespoň jeden prázdný řádek, ale správně by každé definici funkce měly předcházet dva prázdné řádky.
# - Pro dokumentační řetězec se používá """text""" místo #
# - Je nezbytné ošetřovat vstupy od uživatele a naopak zcela zbytečné ošetřovat "vstupy" od počítače
| original/v1/s005-modules/ostrava/Feedback k domácím projektům - moduly.ipynb |
# ---
# title: "Show errors in bar, plot, and boxplot"
# date: 2020-04-12T14:41:32+02:00
# author: "<NAME>"
# type: technical_note
# draft: false
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import pandas as pd
summer2016 = pd.read_csv('summer2016.csv', index_col=0)
mens_rowing = summer2016[summer2016['Sport'] == 'Rowing']
mens_gymnastics = summer2016[summer2016['Sport'] == 'Gymnastics']
seattle_weather = pd.read_csv('seattle_weather.csv').loc[:11]
austin_weather = pd.read_csv('austin_weather.csv')
seattle_weather['MONTH'] = pd.to_datetime(seattle_weather['DATE'], format='%m').dt.month_name().str.slice(stop=3)
austin_weather['MONTH'] = pd.to_datetime(austin_weather['DATE'], format='%m').dt.month_name().str.slice(stop=3)
# -
# ### Adding error-bars to a bar chart
# +
fig, ax = plt.subplots()
# Add a bar for the rowing "Height" column mean/std
ax.bar("Rowing", mens_rowing['Height'].mean(), yerr=mens_rowing['Height'].std())
# Add a bar for the gymnastics "Height" column mean/std
ax.bar("Gymnastics", mens_gymnastics['Height'].mean(), yerr=mens_gymnastics['Height'].std())
# Label the y-axis
ax.set_ylabel("Height (cm)")
plt.show()
# -
# ### Adding error-bars to a plot
# +
fig, ax = plt.subplots()
# Add Seattle temperature data in each month with error bars
ax.errorbar(seattle_weather['MONTH'], seattle_weather['MLY-TAVG-NORMAL'], yerr=seattle_weather['MLY-TAVG-STDDEV'])
# Add Austin temperature data in each month with error bars
ax.errorbar(austin_weather['MONTH'], austin_weather['MLY-TAVG-NORMAL'], yerr=austin_weather['MLY-TAVG-STDDEV'])
# Set the y-axis label
ax.set_ylabel('Temperature (Fahrenheit)')
plt.show()
# -
# ### Creating boxplots
# Tell us what the median of the distribution is, what the inter-quartile range is and also what the expected range of approximately 99% of the data should be. Outliers beyond this range are particularly highlighted.
# +
fig, ax = plt.subplots()
# Add a boxplot for the "Height" column in the DataFrames
ax.boxplot([mens_rowing['Height'], mens_gymnastics['Height']])
# Add x-axis tick labels:
ax.set_xticklabels(['Rowing', 'Gymnastics'])
# Add a y-axis label
ax.set_ylabel('Height (cm)')
plt.show()
# -
fig, ax = plt.subplots()
ax.hist(mens_rowing['Height'], histtype='step', label='Rowing')
ax.hist(mens_gymnastics['Height'], histtype='step', label='Gymnastics')
ax.legend()
plt.show()
| courses/datacamp/notes/python/matplotlibTMP/errorsplot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import wradlib
import numpy as np
import os
import datetime as dt
# %matplotlib
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
from matplotlib.collections import PatchCollection
import datetime
import warnings
warnings.simplefilter('once', DeprecationWarning)
from scipy import ndimage as ndi
from skimage import feature
from skimage.feature import match_template
import h5py
import pandas as pd
import matplotlib
from matplotlib.patches import Circle, Wedge, Polygon, Rectangle
from matplotlib.collections import PatchCollection
from skimage import measure
from skimage import filters
from scipy import ndimage
from skimage.measure import label, regionprops
import math
from matplotlib.patches import Ellipse
for_year = "2016"
rootdir = r"e:\data\radolan\ry"
tmpdir = r"e:\data\radolan\tmp"
h5file = 'ry_%s.hdf5' % for_year
hourlyfile = 'hdf/ry_hourly_%s.hdf5' % for_year
hourlyobjfile = 'hdf/ry_hourly_objects_%s.pickle' % for_year
tstart = "%s-01-01" % for_year
tend = "%s-12-31" % for_year
dffile = "exc_%s.csv" % for_year
nx = 900
ny = 900
thresh = 20. # mm/h
minarea = 10
maxarea = 1500
# ## Extract hourly features (initial detection)
days = wradlib.util.from_to(dt.datetime.strptime(tstart, "%Y-%m-%d"),
dt.datetime.strptime(tend, "%Y-%m-%d"), tdelta=3600*24)
dtimes = wradlib.util.from_to(days[0].strftime("%Y-%m-%d 00:00:00"),
(days[-1]+dt.timedelta(days=1)).strftime("%Y-%m-%d 00:00:00"), tdelta=60*60)
hrs = np.arange(24).astype("i4")
dummy = regionprops(np.ones((4,4)).astype("i4"), intensity_image=np.ones((4,4)) )
keys = list(dummy[0])
props = [ dummy[0].__getitem__(key) for key in keys ]
keys.insert(0, "dtime")
props.insert(0, "1900-01-01 00:00:00")
df = pd.DataFrame( dict([(key, [props[i]]) for i,key in enumerate(keys)] ) )
with h5py.File(hourlyfile, 'r') as f:
for day in days:
print(day.strftime("%Y/%m/%d"), end="")
try:
dset = f[day.strftime("%Y/%m/%d")][:]
except KeyError:
print(" does not exist.")
continue
found = 0
for i, hr in enumerate(hrs):
hset = dset[i]
label_im = measure.label(hset > thresh, background=0)
nb_labels = len(np.unique(label_im))
regions = regionprops(label_im, intensity_image=hset)
for region in regions:
if (region.area < minarea) or (region.area > maxarea):
continue
found += 1
thetime = day.strftime("%Y-%m-%d") + " %02d:00:00" % hr
theprops = [region.__getitem__(prop) for prop in region]
theprops.insert(0, thetime)
df = df.append(dict([(key, theprops[i]) for i,key in enumerate(keys)] ), ignore_index=True)
print(" Found %d regions." % found)
#df.to_pickle(hourlyobjfile)
df = pd.read_pickle(hourlyobjfile)
df = df.set_index(pd.DatetimeIndex(df['dtime']))
df.keys()
# ## Analyse spatial extent of cells
expandby = np.arange(50)
toleft = np.zeros((len(df), len(expandby))) * np.nan
toright = toleft.copy()
tobottom = toleft.copy()
totop = toleft.copy()
with h5py.File(hourlyfile, 'r') as f:
for i in range(len(df)):
dtime = dt.datetime.strptime(df.dtime.iloc[i], "%Y-%m-%d %H:%M:%S")
print(dtime)
try:
hset = f[dtime.strftime("%Y/%m/%d")][dtime.hour]
except KeyError:
continue
left, bottom, right, top = df.bbox.iloc[i][0], df.bbox.iloc[i][1], df.bbox.iloc[i][2], df.bbox.iloc[i][3]
for j, step in enumerate(expandby):
try:
toleft[i,j] = np.nanmean(hset[(left-step):right, bottom:top])
except IndexError:
continue
try:
toright[i,j] = np.nanmean(hset[left:(right+step), bottom:top])
except IndexError:
continue
try:
tobottom[i,j] = np.nanmean(hset[left:right, (bottom-step):top])
except IndexError:
continue
try:
totop[i,j] = np.nanmean(hset[left:right, bottom:(top+step)])
except IndexError:
continue
# +
leftnorm = toleft / toleft[:,0].reshape((-1,1))
leftnorm[leftnorm>1] = np.nan
rightnorm = toright / toright[:,0].reshape((-1,1))
rightnorm[rightnorm>1] = np.nan
bottomnorm = tobottom / tobottom[:,0].reshape((-1,1))
bottomnorm[bottomnorm>1] = np.nan
topnorm = totop / totop[:,0].reshape((-1,1))
topnorm[topnorm>1] = np.nan
# -
print("left")
for i, item in enumerate(leftnorm):
plt.plot(expandby, np.ma.masked_invalid(item), "b-", alpha=0.005)
print("right")
for i, item in enumerate(rightnorm):
plt.plot(expandby, np.ma.masked_invalid(item), "r-", alpha=0.005)
print("bottom")
for i, item in enumerate(bottomnorm):
plt.plot(expandby, np.ma.masked_invalid(item), "g-", alpha=0.005)
print("top")
for i, item in enumerate(topnorm):
plt.plot(expandby, np.ma.masked_invalid(item), "k-", alpha=0.005)
# ## Analyse impact of threshold on mean intensity
# Thresholds of hourly rainfall depths for detection of contiguous regions
threshs = np.arange(19,0,-1)
def get_regions(im, thresh):
"""Extract regions from im which exceed thresh.
"""
label_im = measure.label(im > thresh, background=0)
nb_labels = len(np.unique(label_im))
regions = regionprops(label_im, intensity_image=im)
return(regions)
# +
#means = np.load("hdf/means_2016.numpy.npy")
#areas = np.load("hdf/areas_2016.numpy.npy")
# -
means = np.zeros( (len(df), len(threshs)+1) )
areas = np.zeros( (len(df), len(threshs)+1) )
_dtime = None
with h5py.File(hourlyfile, 'r') as f:
for i in range(len(df)):
dtime = dt.datetime.strptime(df.dtime.iloc[i], "%Y-%m-%d %H:%M:%S")
bottom, left, top, right = df.bbox.iloc[i][0], df.bbox.iloc[i][1], df.bbox.iloc[i][2], df.bbox.iloc[i][3]
means[i, 0] = df.mean_intensity.iloc[i]
areas[i, 0] = df.area.iloc[i]
if dtime != _dtime:
print("")
print(dtime, end="")
# Only process new hourly set for a new datetime
try:
hset = f[dtime.strftime("%Y/%m/%d")][dtime.hour]
except KeyError:
continue
threshregions = [get_regions(hset, thresh) for thresh in threshs]
else:
print(".", end="")
for trix, tr in enumerate(threshregions):
for r in tr:
# Looking for region that contains core region
if (left >= r.bbox[1]) and \
(right <= r.bbox[3]) and \
(bottom >= r.bbox[0]) and \
(top <= r.bbox[2]):
# Found
means[i,trix+1] = r.mean_intensity
areas[i,trix+1] = r.area
_dtime = dtime
np.save("hdf/means_2016", means)
np.save("hdf/areas_2016", areas)
# +
meansnorm = means / means[:,0].reshape((-1,1))
#meansnorm[meansnorm>1] = np.nan
areasnorm = areas / areas[:,0].reshape((-1,1))
#areasnorm[areasnorm>1] = np.nan
vols = areas * means
volsnorm = (vols - vols[:,0].reshape((-1,1)) ) / vols[:,0].reshape((-1,1))
# -
for i, item in enumerate(areasnorm):
plt.plot(np.arange(20,0,-1), item, "b-", alpha=0.005)
matplotlib.rcParams.update({'font.size': 7})
plt.figure(figsize=(14,10))
for i, item in enumerate(range(1300,1400)):
ax1 = plt.subplot(10,10,i+1)
plt.plot(np.arange(20,0,-1), means[item], "b-")
plt.ylim(0,30)
plt.grid()
plt.title(df.dtime.iloc[item] + ", " + str(df.label.iloc[item]), fontsize=7)
ax2 = ax1.twinx()
plt.semilogy(np.arange(20,0,-1), areas[item], "r-")
plt.ylim(10,10000)
plt.tight_layout()
from scipy.signal import argrelextrema
np.gradient(areas[item])
argrelextrema(np.gradient(areas[item]), np.greater)[0]
blacklist_hours = ["2016-06-29 02:00:00"
"2016-06-29 13:00:00",
"2016-06-29 14:00:00",
"2016-07-05 05:00:00",
"2016-07-05 16:00:00",
"2016-07-05 17:00:00"]
blacklist_days = ["2016-06-16", "2016-06-29", "2016-07-04", "2016-07-05"]
for day in blacklist_days:
df.mean_intensity.loc[day] = -9999
bigx = np.argsort(df.mean_intensity)[::-1]
plt.figure()
plt.plot( np.array(df.mean_intensity)[bigx])
plt.ylim(0,60)
matplotlib.rcParams.update({'font.size': 7})
plt.figure(figsize=(14,10))
# Look at the 100 most intensive objects
for i, item in enumerate(bigx[0:100]):
ax1 = plt.subplot(10,10,i+1)
plt.plot(np.arange(20,0,-1), np.gradient(areas[item]), "b-")
# for local maxima
extr = argrelextrema(np.gradient(areas[item]), np.greater)[0]
plt.plot(np.arange(20,0,-1)[extr], np.gradient(areas[item])[extr], "bo")
plt.grid()
plt.ylim(0,20)
plt.title(df.dtime.iloc[item] + ", " + str(df.label.iloc[item]), fontsize=7)
ax2 = ax1.twinx()
plt.plot(np.arange(20,0,-1), areasnorm[item], "g-")
plt.grid()
#plt.plot(np.arange(20,0,-1), areas[item], "r-")
plt.ylim(1,10)
plt.tight_layout()
plt.figure(figsize=(6,6))
ax1 = plt.subplot(2,1,1)
ax2 = plt.subplot(2,1,2)
for item in range(len(means[1:])):
ax1.plot(np.arange(20,0,-1), meansnorm[item], "b-", alpha=0.005)
ax1.grid()
ax2.semilogy(np.arange(20,0,-1), areasnorm[item], "r-", alpha=0.005)
plt.figure(figsize=(6,6))
for i, item in enumerate(volsnorm):
plt.plot(np.arange(20,0,-1), item, "b-", alpha=0.005)
# ## Analyze impact of temporal duration
time_window = np.arange(1, 7)
tdeltas = [dt.timedelta(seconds=i*3600.) for i in time_window]
tmeans = np.zeros( (len(df), len(time_window)+1) )
tareas = np.zeros( (len(df), len(time_window)+1) )
_dtime = None
with h5py.File(hourlyfile, 'r') as f:
for i in range(len(df)):
dtime = dt.datetime.strptime(df.dtime.iloc[i], "%Y-%m-%d %H:%M:%S")
bottom, left, top, right = df.bbox.iloc[i][0], df.bbox.iloc[i][1], df.bbox.iloc[i][2], df.bbox.iloc[i][3]
tmeans[i, 0] = df.mean_intensity.iloc[i]
tareas[i, 0] = df.area.iloc[i]
if dtime != _dtime:
print("")
print(dtime, end="")
# Only process new hourly set for a new datetime
thewindow = [dtime + item for item in tdeltas]
daystrings = [item.strftime("%Y/%m/%d") for item in thewindow]
hours = [item.hour for item in thewindow]
hsets = np.zeros((len(time_window), 900, 900)) * np.nan
for i in range(len(time_window)):
try:
hsets[i] = f[dtime.strftime("%Y/%m/%d")][dtime.hour]
except KeyError:
continue
hsets = np.cumsum(hsets, axis=0)
threshregions = [get_regions(hset, 20.) for hset in hsets]
else:
print(".", end="")
for trix, tr in enumerate(threshregions):
for r in tr:
# Looking for region that contains core region
if (left >= r.bbox[1]) and \
(right <= r.bbox[3]) and \
(bottom >= r.bbox[0]) and \
(top <= r.bbox[2]):
# Found
tmeans[i,trix+1] = r.mean_intensity
tareas[i,trix+1] = r.area
_dtime = dtime
np.save("hdf/tmeans_2016", tmeans)
np.save("hdf/tareas_2016", tareas)
# +
tmeansnorm = tmeans / tmeans[:,0].reshape((-1,1))
#meansnorm[meansnorm>1] = np.nan
tareasnorm = tareas / tareas[:,0].reshape((-1,1))
# -
matplotlib.rcParams.update({'font.size': 7})
plt.figure(figsize=(14,10))
for i, item in enumerate(range(1300,1400)):
ax1 = plt.subplot(10,10,i+1)
plt.plot(np.arange(0,7), tmeans[item], "b-")
plt.grid()
plt.title(df.dtime.iloc[item] + ", " + str(df.label.iloc[item]), fontsize=7)
ax2 = ax1.twinx()
plt.plot(np.arange(0,7), tareas[item], "r-")
plt.tight_layout()
# ## View specific situations
radolan_grid_xy = wradlib.georef.get_radolan_grid(900,900)
x = radolan_grid_xy[:,:,0]
y = radolan_grid_xy[:,:,1]
# +
dtime = "2016-06-07 21:00:00"
dtime_ = dt.datetime.strptime(dtime, "%Y-%m-%d %H:%M:%S")
with h5py.File(hourlyfile, 'r') as f:
hset = f[dtime_.strftime("%Y/%m/%d")][dtime_.hour]
sub = df.loc[dtime]
cmap=plt.cm.nipy_spectral
norm = matplotlib.colors.BoundaryNorm(np.arange(0,21), cmap.N)
plt.figure(figsize=(8,8))
pm = plt.pcolormesh(np.ma.masked_array(hset, ~np.isfinite(hset)), cmap=cmap, norm=norm)
plt.xlabel("RADOLAN easting (km)")
plt.ylabel("RADOLAN northing (km)")
plt.colorbar(pm)
ax = plt.gca()
patches = []
for i in range(0,len(sub)):
polygon = Rectangle(
(sub.iloc[i]["bbox"][1], sub.iloc[i]["bbox"][0]), # (x,y)
sub.iloc[i]["bbox"][3]-sub.iloc[i]["bbox"][1], # height
sub.iloc[i]["bbox"][2]-sub.iloc[i]["bbox"][0] # width
)
patches.append(polygon)
p = PatchCollection(patches, facecolor="None", edgecolor="white", linewidth=2)
ax.add_collection(p)
for i in range(0,len(sub)):
plt.text(sub.iloc[i].centroid[1], sub.iloc[i].centroid[0], str(sub.iloc[i].label), color="red", fontsize=18)
# -
sub.centroid
# +
fig = plt.figure(figsize=(8, 8))
ax = fig.add_subplot(111, aspect="equal")
patches = []
for i in range(1,len(df)):
polygon = Rectangle(
df.iloc[i]["bbox"][0:2], # (x,y)
df.iloc[i]["bbox"][2]-df.iloc[i]["bbox"][0], # width
df.iloc[i]["bbox"][3]-df.iloc[i]["bbox"][1], # height
)
patches.append(polygon)
colors = 100*np.random.rand(len(patches))
p = PatchCollection(patches, alpha=0.4)
p.set_array(np.array(colors))
ax.add_collection(p)
plt.xlim(0,900)
plt.ylim(0,900)
#plt.draw()
# -
# ## Junkyard
proj = wradlib.georef.create_osr("dwd-radolan")
watersheds_shp = r"E:\src\git\heisterm_bitbucket\tsms_data\tsms-data-misc\shapefiles\watersheds_kocher.shp"
dataset, inLayer = wradlib.io.open_vector(watersheds_shp)
cats, ids = wradlib.georef.get_vector_coordinates(inLayer, dest_srs=proj,
key="value")
ids = np.array(ids)
left, right, bottom, top = inLayer.GetExtent()
# +
fig = plt.figure(figsize=(8, 8))
ax = fig.add_subplot(111, aspect="equal")
patches = []
for i in range(1,len(df)):
polygon = Rectangle(
df.iloc[i]["bbox"][0:2], # (x,y)
df.iloc[i]["bbox"][2]-df.iloc[i]["bbox"][0], # width
df.iloc[i]["bbox"][3]-df.iloc[i]["bbox"][1], # height
)
patches.append(polygon)
colors = 100*np.random.rand(len(patches))
p = PatchCollection(patches, alpha=0.4)
p.set_array(np.array(colors))
ax.add_collection(p)
#for i in range(1,len(df)):
# plt.plot(df.ix[i]["centroid"][0], df.ix[i]["centroid"][1], "bo")
plt.xlim(0,900)
plt.ylim(0,900)
#plt.draw()
#wradlib.vis.add_lines(ax, cats, color='red', lw=0.5, zorder=4, alpha=0.3)
#plt.xlim(-40,20)
#plt.ylim(-4440,-4390)
# -
toobigx = np.where(df["area"]>1500)[0]
print(len(toobigx))
for i in toobigx:
plt.figure()
plt.pcolormesh(df.iloc[i]["image"])
i
plt.hist(df["area"], bins=100, range=(0,200), log=True)
# +
plt.figure(figsize=(8,8))
#plt.imshow(im.mean(axis=0), cmap=plt.cm.gray, origin="lower")
plt.imshow(frames[start:end].sum(axis=0), cmap=plt.cm.gray, origin="lower", vmin=0, vmax=30)
plt.xlabel("RADOLAN easting (km)")
plt.ylabel("RADOLAN northing (km)")
plt.title("Rainfall accumulation and cell tracks\nMay 29, 2016, 15:00-18:00 UTC")
ax = plt.gca()
for label in labels[1:]:
#for i in range(len(im)):
tmp = (label_im == label).astype("int")
#tmp = label_im[i]
regions = regionprops(tmp, intensity_image=im)
centx, centy = [], []
for region in regions:
y0, x0 = region.centroid
centx.append(x0)
centy.append(y0)
orientation = region.orientation
angle=-np.rad2deg( orientation)
e = Ellipse([x0,y0], region.major_axis_length, region.minor_axis_length,
angle=angle, facecolor="none", edgecolor="blue", linewidth=1.3, alpha=0.5)
ax.add_artist(e)
#plt.plot(x0, y0, "o", markerfacecolor=plt.cm.rainbow(i/len(im)), markeredgecolor="none", alpha=0.5)
plt.contour(tmp, [0.5], linewidths=1., colors="red", alpha=0.5)
#pm=plt.scatter([], [], c=[], cmap=plt.cm.rainbow, vmin=0, vmax=len(im)*5)
#cb=plt.colorbar(pm, label="Minutes from 2016-05-29 16:00", shrink=0.75)
# -
fig = plt.figure(figsize=(12,12))
ax = fig.add_subplot(111)
for i, label in enumerate(labels):
tmp = (label_im == label)
areal_avg = np.array([np.mean(frames[i][tmp]) for i in range(len(frames))])
ax.plot(np.cumsum(areal_avg))
from matplotlib import animation
# +
# Animate features
# Prepare canvas
fig = plt.figure(figsize=(5,5))
ax = plt.subplot(111,aspect="equal")
im1 = ax.imshow(frames[0], origin="lower", cmap="gray", interpolation="none", vmin=10, vmax=20)
plt.xlabel("Easting (km)")
plt.ylabel("Northing (km)")
plt.grid(color="white")
plt.xlim(150,450)
plt.ylim(550,900)
#ax1.plot(x[0,goodtrack], y[0,goodtrack], linestyle="None", marker="o", mfc="None", mec="limegreen")
#ax1.plot(x[0,~goodtrack], y[0,~goodtrack], linestyle="None", marker="o", mfc="None", mec="red")
ax.grid(color="white")
tstamp1 = ax.text(160, 560, dtimes[0].isoformat(), color="white", fontsize=12)
def animate(j):
im1.set_array(frames[0+j])
tstamp1.set_text(dtimes[0+j].isoformat())
for label in labels[1:]:
#break
tmp = (label_im[j] == label).astype("int")
#tmp = label_im[i]
regions = regionprops(tmp, intensity_image=im[j])
centx, centy = [], []
for region in regions:
y0, x0 = region.centroid
centx.append(x0)
centy.append(y0)
orientation = region.orientation
angle=-np.rad2deg( orientation)
e = Ellipse([x0,y0], region.major_axis_length, region.minor_axis_length,
angle=angle, facecolor="none", edgecolor=plt.cm.rainbow(j/len(im)), linewidth=1.3, alpha=0.3)
ax.add_artist(e)
#ax.plot(x0, y0, "o", markerfacecolor=plt.cm.rainbow(j/len(im)), markeredgecolor="none", alpha=0.5)
tstamp1.set_text(dtimes[0+j].isoformat())
return im1
# ATTENTION: THIS IS SLOW - Rendering each frame of the animation might take more time than the interval between the frames
# This can cause the temporal sequence to be confused in the matplotlib interactive mode.
# The animation thus looks better if saved as movie, or you have to increase the interval argument
# Animation not shown in notebook if you use %pylab inline
maxi = len(frames)-1
ani = animation.FuncAnimation(fig, animate, frames=np.arange(0, maxi), interval=400, blit=False)
ani.save("features.gif", writer="imagemagick", dpi=150)
# -
len(region)
# +
#fig, ax = plt.subplots()
plt.imshow(im, cmap=plt.cm.gray, origin="lower")
plt.contour(label_im, [0.5], linewidths=1.2, colors='y')
plt.xlabel("RADOLAN easting (km)")
plt.ylabel("RADOLAN northing (km)")
plt.title("Snaphot at 2016-05-29 16:00 UTC")
ax = plt.gca()
for i, props in enumerate(regions):
y0, x0 = props.centroid
orientation = props.orientation
x1 = x0 + math.cos(orientation) * 0.5 * props.major_axis_length
y1 = y0 - math.sin(orientation) * 0.5 * props.major_axis_length
x2 = x0 - math.sin(orientation) * 0.5 * props.minor_axis_length
y2 = y0 - math.cos(orientation) * 0.5 * props.minor_axis_length
#plt.plot((x0, x1), (y0, y1), '--r', linewidth=2)
#plt.plot((x0, x2), (y0, y2), '--r', linewidth=2)
#plt.plot(x0, y0, '.r', markersize=15)
angle=-np.rad2deg( props.orientation)
e = Ellipse([x0,y0], props.major_axis_length, props.minor_axis_length,
angle=angle, facecolor="none", edgecolor="red", linewidth=2)
ax.add_artist(e)
minr, minc, maxr, maxc = props.bbox
bx = (minc, maxc, maxc, minc, minc)
by = (minr, minr, maxr, maxr, minr)
#plt.plot(bx, by, '-b', linewidth=2.5)
try:
label = "ID=%s\navg=%d mm/h\nmax=%d mm/h" % (props.label, props.mean_intensity, props.max_intensity)
except:
label = "ID=%s, avg=%s mm/h, max=%s mm/h" % (props.label, "nan", "nan")
plt.text((minc+maxc)/2, maxr+2, label, color="red", fontsize=10, horizontalalignment='center')
#plt.axis((0, 900, 900, 0))
plt.xlim(200,900)
plt.ylim(0,470)
# -
minr, minc, maxr, maxc = props.bbox
plt.imshow(im[minr:maxr, minc:maxc])
# +
im2 = frames[1]
fig = plt.figure(figsize=(8, 8))
ax2 = plt.subplot(1, 1, 1)
for i, props in enumerate(regions):
minr, minc, maxr, maxc = props.bbox
roi = im[minr:maxr, minc:maxc]
result = match_template(im2, roi)
ij = np.unravel_index(np.argmax(result), result.shape)
x, y = ij[::-1]
print(ij)
#ax1.imshow(roi, cmap=plt.cm.gray)
#ax1.set_axis_off()
#ax1.set_title('Feature #1 at t+0')
ax2.imshow(im2, cmap=plt.cm.gray, origin="lower")
ax2.set_axis_off()
ax2.set_title('Feature #1 at t+2')
# highlight matched region
hcoin, wcoin = roi.shape
rect = plt.Rectangle((x, y), wcoin, hcoin, edgecolor='r', facecolor='none')
ax2.add_patch(rect)
plt.plot(x,y,".r")
plt.plot(ij[0],ij[1],".b")
# highlight matched region
bx = (minc, maxc, maxc, minc, minc)
by = (minr, minr, maxr, maxr, minr)
plt.plot(bx, by, '-b', linewidth=1.)
# -
ij
ndimage.find_objects(label_im==15)
# +
image = frames[2]
coin = roi
result = match_template(image, coin)
ij = np.unravel_index(np.argmax(result), result.shape)
x, y = ij[::-1]
fig = plt.figure(figsize=(8, 3))
ax1 = plt.subplot(1, 2, 1)
ax2 = plt.subplot(1, 2, 2, adjustable='box-forced')
ax1.imshow(coin, cmap=plt.cm.gray)
ax1.set_axis_off()
ax1.set_title('Feature #1 at t+0')
ax2.imshow(image, cmap=plt.cm.gray)
ax2.set_axis_off()
ax2.set_title('Feature #1 at t+2')
# highlight matched region
hcoin, wcoin = coin.shape
rect = plt.Rectangle((x, y), wcoin, hcoin, edgecolor='r', facecolor='none')
ax2.add_patch(rect)
# -
| extract_depth_features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PythonData
# language: python
# name: pythondata
# ---
# # Pyber Challenge
# ### 4.3 Loading and Reading CSV files
# +
# Add Matplotlib inline magic command
# %matplotlib inline
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
# File to Load (Remember to change these)
city_data_to_load = "./Resources/city_data.csv"
ride_data_to_load = "./Resources/ride_data.csv"
# Read the City and Ride Data
city_data_df = pd.read_csv(city_data_to_load)
ride_data_df = pd.read_csv(ride_data_to_load)
# -
# ### Merge the DataFrames
# +
# Combine the data into a single dataset
pyber_data_df = pd.merge(ride_data_df, city_data_df, how="left", on=["city", "city"])
# Display the data table for preview
pyber_data_df.head()
# -
# ## Deliverable 1: Get a Summary DataFrame
# 1. Get the total rides for each city type
total_rides_by_type = pyber_data_df.groupby(["type"]).count()["ride_id"]
total_rides_by_type
# 2. Get the total drivers for each city type
total_drivers_by_type = city_data_df.groupby(["type"]).sum()["driver_count"]
total_drivers_by_type
# 3. Get the total amount of fares for each city type
total_fare_by_type = pyber_data_df.groupby(["type"]).sum()["fare"]
total_fare_by_type
# 4. Get the average fare per ride for each city type.
avg_fare_per_ride = total_fare_by_type / total_rides_by_type
avg_fare_per_ride
# 5. Get the average fare per driver for each city type.
avg_fare_per_driver = total_fare_by_type / total_drivers_by_type
avg_fare_per_driver
# 6. Create a PyBer summary DataFrame.
summary = {
"Total Rides": total_rides_by_type,
"Total Drivers": total_drivers_by_type,
"Total Fares":total_fare_by_type,
"Average Fare per Ride": avg_fare_per_ride,
"Average Fare per Driver": avg_fare_per_driver
}
pyber_ride_summary_df = pd.DataFrame(summary)
pyber_ride_summary_df
# 7. Cleaning up the DataFrame. Delete the index name
pyber_ride_summary_df.index.name = None
pyber_ride_summary_df
# 8. Format the columns.
pyber_ride_summary_df["Total Rides"] = pyber_ride_summary_df["Total Rides"].map("{:,}".format)
pyber_ride_summary_df["Total Drivers"] = pyber_ride_summary_df["Total Drivers"].map("{:,}".format)
pyber_ride_summary_df["Total Fares"] = pyber_ride_summary_df["Total Fares"].map("${:,.2f}".format)
pyber_ride_summary_df["Average Fare per Ride"] = pyber_ride_summary_df["Average Fare per Ride"].map("${:,.2f}".format)
pyber_ride_summary_df["Average Fare per Driver"] = pyber_ride_summary_df["Average Fare per Driver"].map("${:,.2f}".format)
pyber_ride_summary_df
# ## Deliverable 2. Create a multiple line plot that shows the total weekly of the fares for each type of city.
# 1. Read the merged DataFrame
pyber_data_df
# 2. Using groupby() to create a new DataFrame showing the sum of the fares
# for each date where the indices are the city type and date.
sum_of_fares_df = pyber_data_df.groupby(["type", "date"]).sum()["fare"]
sum_of_fares_df
# 3. Reset the index on the DataFrame you created in #1. This is needed to use the 'pivot()' function.
# df = df.reset_index()
sum_of_fares_df = sum_of_fares_df.reset_index()
# 4. Create a pivot table with the 'date' as the index, the columns ='type', and values='fare'
# to get the total fares for each type of city by the date.
fares_by_type_df = sum_of_fares_df.pivot(index="date", columns="type", values="fare")
fares_by_type_df
# 5. Create a new DataFrame from the pivot table DataFrame using loc on the given dates, '2019-01-01':'2019-04-29'.
fares_by_type_clean_df=fares_by_type_df.loc['1/1/2019':'4/28/2019']
fares_by_type_clean_df.tail()
# 6. Set the "date" index to datetime datatype. This is necessary to use the resample() method in Step 8.
# df.index = pd.to_datetime(df.index)
fares_by_type_clean_df.index = pd.to_datetime(fares_by_type_clean_df.index)
# 7. Check that the datatype for the index is datetime using df.info()
fares_by_type_clean_df.info()
# 8. Create a new DataFrame using the "resample()" function by week 'W' and get the sum of the fares for each week.
weekly_fares_df = fares_by_type_clean_df.resample('W').sum()
weekly_fares_df
# +
# 8. Using the object-oriented interface method, plot the resample DataFrame using the df.plot() function.
# Import the style from Matplotlib.
from matplotlib import style
# Use the graph style fivethirtyeight.
style.use('fivethirtyeight')
ax = weekly_fares_df.plot(figsize=(20,6))
# Title
ax.set_title("Total Fare by City Type")
# x and y labels
ax.set_xlabel("Month")
ax.set_ylabel("Fare ($USD)")
plt.savefig("analysis/Challenge_fare_summary.png")
plt.show()
# -
| PyBer_Challenge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: codeforecon
# language: python
# name: codeforecon
# ---
# (working-with-data)=
# # Working with Data
#
# ## Introduction
#
# The previous chapter was just a quick tour of what can be done with a single tabular dataset (a 'dataframe'). In this chapter, we'll go deeper into working with data.
#
# The ability to extract, clean, and analyse data is one of the core skills any economist needs. Fortunately, the (open source) tools that are available for data analysis have improved enormously in recent years, and working with them can be a delight--even the most badly formatted data can be beaten into shape.
#
# In this chapter, you'll get really good introduction to the [**pandas**](https://pandas.pydata.org/) package, the core data manipulation library in Python. The name is derived from 'panel data' but it's suited to any tabular data, and can be used to work with more complex data structures too. We *won't* cover reading in or writing data here; see the next chapter for that.
#
# This chapter is hugely indebted to the fantastic [Python Data Science Handbook](https://jakevdp.github.io/PythonDataScienceHandbook/) by <NAME>. Remember, if you get stuck with pandas, there is brilliant [documentation](https://pandas.pydata.org/docs/user_guide/index.html) and a fantastic set of [introductory tutorials](https://pandas.pydata.org/pandas-docs/stable/getting_started/intro_tutorials/index.html) on their website. These notes are heavily indebted to those introductory tutorials.
#
# This chapter uses the **pandas**, **seaborn**, and **numpy** packages. If you're running this code, you may need to install these packages, which you can do using either `conda install packagename` or `pip install packagename` on your computer's command line. (If you're not sure what a command line or terminal is, take a quick look at the basics of coding chapter.)
#
#
# ### Using tidy data
#
# As an aside, if you're working with tabular data, it's good to try and use a so-called 'tidy data' format. This is data with one observation per row, and one variable per column, like so:
#
# 
#
# Tidy data aren't going to be appropriate *every* time and in every case, but they're a really, really good default for tabular data. Once you use it as your default, it's easier to think about how to perform subsequent operations. Some plotting libraries, such as **seaborn**, take that your data are in tidy format as a given. And many operations that you can perform on dataframes (the objects that hold tabular data within many programming languages) are easier when you have tidy data. If you're writing out data to file to share, putting it in tidy format is a really good idea.
#
# Of course, *getting* your messy dataset into a tidy format may take a bit of work... but we're about to enter the exciting world of coding for data analysis and the tools you'll see in the rest of this chapter will help you to 'wrangle' even the most messy of datasets.
#
# Having said that tidy data are great, and they are, one of standard data library **pandas**' advantages relative to other data analysis libraries is that it isn't *too* tied to tidy data and can navigate awkward non-tidy data manipulation tasks happily too.
# ## Dataframes and series
#
# Let's start with the absolute basics. The most basic **pandas** object is a dataframe. A DataFrame is a 2-dimensional data structure that can store data of different types (including characters, integers, floating point values, categorical data, even lists) in columns.
#
# 
#
# We'll now import some useful packages, set a random number seed (some examples use randomly generated data), set up some nice plot settings, and load a dataset.
#
# We'll look at a dataframe of the *penguins* dataset. To show just the first 5 rows, I'll be using the `head()` method (there's also a `tail()` method).
#
# +
import seaborn as sns
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import os
# Set seed for reproducibility
np.random.seed(10)
# Plot settings
plt.style.use(
"https://github.com/aeturrell/coding-for-economists/raw/main/plot_style.txt"
)
df = sns.load_dataset("penguins")
df.head()
# -
# What just happened? We loaded a pandas dataframe called `df` and showed its contents. You can see the column names in bold, and the index on the left hand side. Just to double check it *is* a pandas dataframe, we can call type on this.
type(df)
# And if we want a bit more information about what we imported (including the datatypes of the columns):
df.info()
# Remember that everything in Python is an object, and our dataframe is no exception. Each dataframe is made up of a set of series that, in a dataframe, become columns: but you can turn a single series into a dataframe too.
#
# 
#
# Let's see a couple of ways of creating some series from raw data:
# + tags=["hide-output"]
# From a list:
s1 = pd.Series([1.0, 6.0, 19.0, 2.0])
print(s1)
print("\n")
# From a dictionary
population_dict = {
"California": 38332521,
"Texas": 26448193,
"New York": 19651127,
"Florida": 19552860,
"Illinois": 12882135,
}
s2 = pd.Series(population_dict)
print(s2)
# -
# Note that in each case there is no column name (because this is a series, not a dataframe), and there *is* an index. The index is automatically created if we don't specify it; in the third example, by passing a dictionary we implicitly asked for the index to be the locations we supplied.
#
# If you ever need to get the data 'out' of a series or dataframe, you can just call the `values` method on the object:
s2.values
# If you ever want to turn a series into a dataframe, just called `pd.DataFrame(series)` on it. Note that while series have an index and an object name (eg `s2` above), they don't have any column labels because they only have one column.
#
# Now let's try creating our own dataframe with more than one column of data using a *dictionary*:
df = pd.DataFrame(
{
"A": 1.0,
"B": pd.Series(1, index=list(range(4)), dtype="float32"),
"C": [3] * 4,
"D": pd.Categorical(["test", "train", "test", "train"]),
"E": "foo",
}
)
df
# Remember, curly brackets in the format `{key: value}` denote a dictionary. In the above example, the `pd.DataFrame()` function understands that any single value entries in the dictionary that is passed, such as `{'A': 1.}`, should be repeated as many times as are needed to match the longest series in the dictionary (4 in the above example).
#
# Another way to create dataframes is to pass a bunch of series (note that `index`, `columns`, and `dtype` are optional--you can just specify the data):
df = pd.DataFrame(
data=np.reshape(range(36), (6, 6)),
index=["a", "b", "c", "d", "e", "f"],
columns=["col" + str(i) for i in range(6)],
dtype=float,
)
df
# Note that `reshape` takes an input and puts it into a given shape (a 6 by 6 matrix in the above example).
# ### Values, columns, and index
#
# You'll have seen that there are three different things that make up a dataframe: the values that are in the cells, the column names, and the index. The column and index can take on values that are the same as the values in a dataframe do; string, int, float, datetime, and more. It's pretty obvious what role the columns play: they keep track of the name of different sets of values. But for people who may have seen other dataframe-like libraries, the role played by the index may be less familiar. The easiest way to think about a **pandas** index is that it does for row values what the columns do for columnar values: it's a way of keeping track of what individual roles are and it *doesn't* get used for calculations (just as summing a column ignores the name of the row).
#
# Here's an example to show this. Let's first create a simple dataframe:
df = pd.DataFrame(
data={
"col0": [0, 0, 0, 0],
"col1": [0, 0, 0, 0],
"col2": [0, 0, 0, 0],
"col3": ["a", "b", "b", "a"],
"col4": ["alpha", "gamma", "gamma", "gamma"],
},
index=["row" + str(i) for i in range(4)],
)
df.head()
# If we add one to the integer columns in the dataframe, this is what we get (note we're not saving the result):
df[["col0", "col1", "col2"]] + 1
# Now let's use `col0` as our index instead of the original labels we created and add one to the remaining numeric columns:
df = df.set_index("col0")
df[["col1", "col2"]] = df[["col1", "col2"]] + 1
df.head()
# What was a column name has become an index name (which you can change with `df.index.name='newname'`) and, when we do add one, it isn't applied to the index. Even though their datatype is `int`, for integer, the index entries are now acting as a label for each row--not as values in the dataframe.
#
# An index can be useful for keeping track of what's going on, and it's particularly convenient for some datetime operations.
#
# Whenever you use `groupby` (and some other operations), the columns you use to perform the operation are set as the index of the returned dataframe (you can have multiple index columns). To get back those back to being columns, use the `reset_index()` method like so:
df.groupby(["col3", "col4"]).sum()
df.groupby(["col3", "col4"]).sum().reset_index()
# ## Datatypes
# Pandas has some built-in datatypes (some are the basic Python datatypes) that will make your life a *lot* easier if you work with them. Why bother specifying datatypes? Languages like Python let you get away with having pretty much anything in your columns. But this can be a problem: sometimes you'll end up mixing integers, strings, the generic 'object' datatype, and more by mistake. By ensuring that columns conform to a datatype, you can save yourself from some of the trials that come with these mixed datatypes. Some of the most important datatypes for dataframe are string, float, categorical, datetime, int, and boolean.
#
# Typically, you'll read in a dataset where the dataypes of the columns are a mess. One of the first things you'll want to do is sort these out. Here's an example dataset showing how to set the datatypes:
# +
data = [
["string1", "string2"],
[1.2, 3.4],
["type_a", "type_b"],
["01-01-1999", "01-01-2000"],
[1, 2],
[0, 1],
]
columns = [
"string_col",
"double_col",
"category_col",
"datetime_col",
"integer_col",
"bool_col",
]
df = pd.DataFrame(data=np.array(data).T, columns=columns)
df.info()
# -
# Note that the data type for all of these columns is the generic 'Object' (you can see this from the `Dtype` column that is printed when you use `df.info()`). Let's fix that:
#
df = df.assign(
string_col=df["string_col"].astype("string"),
double_col=df["double_col"].astype("double"),
category_col=df["category_col"].astype("category"),
datetime_col=df["datetime_col"].astype("datetime64[ns]"),
integer_col=df["integer_col"].astype("int"),
bool_col=df["bool_col"].astype("bool"),
)
df.info()
# If you're creating a series or dataframe from scratch, here's how to start off with these datatypes:
#
# +
str_s = pd.Series(["string1", "string2"], dtype="string")
float_s = pd.Series([1.2, 3.4], dtype=float)
cat_s = pd.Series(["type_a", "type_b"], dtype="category")
date_s = pd.Series(["01-01-1999", "01-01-2000"], dtype="datetime64[ns]")
int_s = pd.Series([1, 2], dtype=int)
bool_s = pd.Series([True, False], dtype=bool)
df = pd.concat([str_s, float_s, cat_s, date_s, int_s, bool_s], axis=1)
df.info()
# -
# ### Categorical variables
#
# Categorical variables can be especially useful and there are a couple of convenience functions that allow you to create them from other types of columns. Cut splits input data into a given number of (evenly spaced) bins that you can optionally give names to via the `labels=` keyword. The default behaviour is for the order of the labels to matter so that, in the below example, the bins will be evenly spaced from the smallest value to the largest, with the smallest receiving the label `bad`.
#
#
pd.cut([1, 7, 5, 4, 6, 3], 3, labels=["bad", "medium", "good"])
# We can also pass the bins directly:
pd.cut([1, 7, 5, 4, 6, 3], bins=[-5, 0, 5, 10])
# Another useful function is qcut, which provides a categorical breakdown according to a given number of quantiles (eg 4 produces quartiles):
pd.qcut(range(1, 10), 4)
# ## Accessing and slicing
#
# Now you know how to put data in a dataframe, how do you access the bits of it you need? There are various ways. If you want to access an entire column, the syntax is very simple; `df['columname']` (you can also use `df.columname`).
#
# 
#
#
#
df = pd.DataFrame(
data=np.reshape(range(36), (6, 6)),
index=["a", "b", "c", "d", "e", "f"],
columns=["col" + str(i) for i in range(6)],
dtype=float,
)
df["col1"]
# To access a particular row, it's `df.loc['rowname']` or `df.loc[['rowname1', 'rowname1']]`.
#
# 
#
df.loc[["a", "b"]]
# As well as the `.loc` method, there is the `.iloc` method that accesses rows or columns based on their position, for example `df.iloc[i, :]` for the ith row and `df.iloc[:, j]` for the jth column (but remember the numbers start from zero).
#
# To access an individual value from within the dataframe, we have two options: pass an index value and a column name to `.loc[rowname, columnname]` or retrieve the value by using its position using `.iloc[row, column]`:
# Using .loc
print(df.loc["b", "col1"])
# Using .iloc
print(df.iloc[1, 0])
# With all of these different ways to access values in dataframes, it can get confusing. These are the different ways to get the first column of a dataframe (when that first column is called `column` and the dataframe is `df`):
#
# - `df.column`
# - `df["column"]`
# - `df.loc[:, "column"]`
# - `df.iloc[:, 0]`
#
# The ways to access rows are similar (here assuming the first row is called `row`):
#
# - `df.loc["row", :]`
# - `df.iloc[0, :]`
#
# And to access the first value (ie the value in first row, first column):
#
# - `df.column[0]`
# - `df["column"][0]`
# - `df.iloc[0, 0]`
# - `df.loc["row", "column"]`
#
# In the above examples, square brackets are instructions about *where* to grab bits from the dataframe. They are a bit like an address system for values within a dataframe.
#
# Square brackets *also* denote lists though. So if you want to select *multiple* columns or rows, you might see syntax like this:
#
# `df.loc[["row0", "row1"], ["column0", "column2"]]`
#
# which picks out two rows and two columns via the lists `["row0", "row1"]` and `["column0", "column2"]`. Because there are lists alongside the usual system of selecting values, there are two sets of square brackets.
# So often what we really want is a subset of values (as opposed to *all* values or just *one* value). This is where *slicing* comes in. If you've looked at the Basics of Coding chapter, you'll know a bit about slicing and indexing already, but we'll cover the basics here too.
#
# The syntax for slicing is similar to what we've seen already: there are two methods `.loc` to access items by name, and `.iloc` to access them by position. The syntax for the former is `df.loc[start:stop:step, start:stop:step]`, where the first position is index name and the second is column name (and the same applies for numbers and `df.iloc`). Let's see an example using the storms dataset, and do some cuts.
df.loc["a":"f":2, "col1":"col3"]
# As you can see, slicing even works on names! By asking for rows `'a':'f':2`, we get every other row from 'a' to 'f' (inclusive). Likewise, for columns, we asked for every column between `col1` and `col3` (inclusive). `iloc` works in a very similar way.
df.iloc[1:, :-1]
# In this case, we asked for everything from row 1 onwards, and everything up to (but excluding) the last column.
#
# It's not just strings and positions that can be sliced though, here's an example using *dates* (pandas support for dates is truly excellent):
index = pd.date_range("1/1/2000", periods=12, freq="Q")
df = pd.DataFrame(np.random.randint(0, 10, (12, 5)), index=index, columns=list("ABCDE"))
df
# Now let's do some slicing!
df.loc["2000-01-01":"2002-01-01", :]
# Two important points to note here: first, pandas doesn't mind that we supplied a date that didn't actually exist in the index. It worked out that by '2000-01-01' we meant a datetime and compared the values of the index to that datetime in order to decide what rows to return from the dataframe. The second thing to notice is the use of `:` for the column names; this explicitly says 'give me all the columns'.
# ## Operations on dataframes
#
# Columns in dataframes can undergo all the usual arithmetic operations you'd expect of addition, multiplication, division, and so on. If the underlying datatypes of two columns have a group operation, then the dataframe columns will use that.
#
# 
#
# The results of these manipulations can just be saved as a new series, eg, `new_series = df['A'] + df['B']` or created as a new column of the dataframe:
df["new_col"] = df["A"] * (df["B"] ** 2) + 1
df
# Boolean variables and strings have group operations (eg concatenation is via `+` with strings), and so work well with column operations too:
df = pd.DataFrame(
{"a": [1, 0, 1], "b": [0, 1, 1], "c": [0, 1, 1], "d": [1, 1, 0]}, dtype=bool
)
print(df)
print("\n a and c:\n")
print(df["a"] & df["c"])
print("\n b or d:\n")
print(df["b"] | df["d"])
# More complex operations on whole dataframes are supported, but if you're doing very heavy lifting you might want to just switch to using numpy arrays (**numpy** is basically Matlab in Python). As examples though, you can transpose and exponentiate easily:
df = pd.DataFrame(np.random.randint(0, 5, (3, 5)), columns=list("ABCDE"))
print("\n Dataframe:")
print(df)
print("\n Exponentiation:")
print(np.exp(df))
print("\n Transpose:")
print(df.T)
# ## Aggregation
#
# **pandas** has built-in aggregation functions such as
#
# | Aggregation | Description |
# | ----------- | ----------- |
# | `count()` | Number of items |
# | `first()`, `last()` | First and last item |
# | `mean()`, `median()` | Mean and median |
# | `min()`, `max()` | Minimum and maximum |
# | `std()`, `var()` | Standard deviation and variance |
# | `mad()` | Mean absolute deviation |
# | `prod()` | Product of all items |
# | `sum()` | Sum of all items |
# | `value_counts()` | Counts of unique values |
#
# these can applied to all entries in a dataframe, or optionally to rows or columns using `axis=0` or `axis=1` respectively.
#
df.sum(axis=0)
# ## Split, apply, and combine
#
# Splitting a dataset, applying a function, and combining the results are three key operations that we'll want to use again and again. Splitting means differentiating between rows or columns of data based on some conditions, for instance different categories or different values. Applying means applying a function, for example finding the mean or sum. Combine means putting the results of these operations back into the dataframe, or into a variable. The figure gives an example
#
# 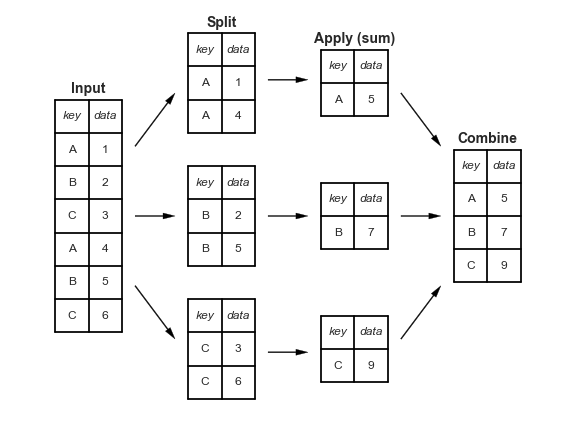
#
# Note that the 'combine' part doesn't always have to result in a new dataframe; it could create new columns in an existing dataframe.
#
# Let's first see a really simple example of splitting a dataset into groups and finding the mean across those groups using the *penguins* dataset. We'll group the data by island and look at the means.
df = sns.load_dataset("penguins")
df.groupby("island").mean()
# The aggregations from the previous part all work on grouped data. An example is `df['body_mass_g'].groupby('island').std()` for the standard deviation of body mass by island.
#
# You can also pass other functions via the `agg` method (short for aggregation). Here we pass two numpy functions:
#
df.groupby("species").agg([np.mean, np.std])
# Multiple aggregations can also be performed at once on the entire dataframe by using a dictionary to map columns into functions. You can also group by as many variables as you like by passing the groupby method a list of variables. Here's an example that combines both of these features:
#
df.groupby(["species", "island"]).agg({"body_mass_g": "sum", "bill_length_mm": "mean"})
# Sometimes, inheriting the column names becomes problematic. There's a slightly fussy syntax to help with that:
#
df.groupby(["species", "island"]).agg(
count_bill=("bill_length_mm", "count"),
mean_bill=("bill_length_mm", "mean"),
std_flipper=("flipper_length_mm", np.std),
)
# Finally, you should know about the `apply` method, which takes a function and applies it to a given axis (`axis=0` for index, `axis=1` for columns) or column. The simple example below shows how it works, though in practice you'd just use `df['body_mass_kg'] = df['body_mass_g]/1e3` to do this.
# +
def g_to_kg(mass_in_g):
return mass_in_g / 1e3
df["mass_in_kg"] = df["body_mass_g"].apply(g_to_kg)
df.head()
# -
# ## Filter, transform, apply, and assign
#
# ### Filter
#
# Filtering does exactly what it sounds like, but it can make use of group-by commands. In the example below, all but one species is filtered out.
#
# In the example below, `filter` passes a grouped version of the dataframe into the `filter_func` we've defined (imagine that a dataframe is passed for each group). Because the passed variable is a dataframe, and variable `x` is defined in the function, the `x` within `filter_func` body behaves like our dataframe--including having the same columns.
# +
def filter_func(x):
return x["bill_length_mm"].mean() > 48
df.groupby("species").filter(filter_func).head()
# -
# ### Transform
#
# Transforms return a transformed version of the data that has the same shape as the input. This is useful when creating new columns that depend on some grouped data, for instance creating group-wise means. Here's an example using the datetime group to subtract a yearly mean. First let's create some synthetic data with some data, a datetime index, and some groups:
index = pd.date_range("1/1/2000", periods=10, freq="Q")
data = np.random.randint(0, 10, (10, 2))
df = pd.DataFrame(data, index=index, columns=["values1", "values2"])
df["type"] = np.random.choice(["group" + str(i) for i in range(3)], 10)
df
# Now we take the yearly means by type. `pd.Grouper(freq='A')` is an instruction to take the `A`nnual mean using the given datetime index. You can group on as many coloumns and/or index properties as you like: this example groups by a property of the datetime index and on the `type` column, but performs the computation on the `values1` column.
df["v1_demean_yr_type"] = df.groupby([pd.Grouper(freq="A"), "type"])[
"values1"
].transform(lambda x: x - x.mean())
df
# You'll have seen there's a `lambda` keyword here. Lambda (or anonymous) functions have a rich history in mathematics, and were used by scientists such as Church and Turing to create proofs about what is computable *before electronic computers existed*. They can be used to define compact functions:
multiply_plus_one = lambda x, y: x * y + 1
multiply_plus_one(3, 4)
# ### Apply
#
#
# Both regular functions and lambda functions can be used with the more general apply method, which takes a function and applies it to a given axis (`axis=0` for index, `axis=1` for columns):
df["val1_times_val2"] = df.apply(lambda row: row["values1"] * row["values2"], axis=1)
df
# Of course, the much easier way to do this very common operation is `df['val1_times_val2'] = df['values1']*df['values2']`, but there are times when you need to run more complex functions element-wise and, for those, `apply` is really useful.
# ### Assign
#
# Assign is a method that allows you to return a new object with all the original columns in addition to new ones. Existing columns that are re-assigned will be overwritten. This is *really* useful when you want to perform a bunch of operations together in a concise way and keep the original columns. For instance, to demean the 'values1' column by year-type and to recompute the 'val1_times_val2' column using the newly demeaned 'values1' column:
df.assign(
values1=(
df.groupby([pd.Grouper(freq="A"), "type"])["values1"].transform(
lambda x: x - x.mean()
)
),
val1_times_val2=lambda x: x["values1"] * x["values2"],
)
# ## Time series, resampling, and rolling windows
#
# The support for time series and the datetime type is excellent in pandas. It is very easy to manipulate datetimes. The [relevant part](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html) of the documentation has more info; here we'll just see a couple of the most important bits. First, let's create some synthetic data to work with:
# +
def recursive_ts(n, x=0.05, beta=0.6, alpha=0.2):
shock = np.random.normal(loc=0, scale=0.6)
if n == 0:
return beta * x + alpha + shock
else:
return beta * recursive_ts(n - 1, x=x) + alpha + shock
t_series = np.cumsum([recursive_ts(n) for n in range(12)])
index = pd.date_range("1/1/2000", periods=12, freq="M")
df = pd.DataFrame(t_series, index=index, columns=["values"])
df.loc["2000-08-31", "values"] = np.nan
df
# -
# Now let's imagine that there are a number of issues with this time series. First, it's been recorded wrong: it actually refers to the start of the next month, not the end of the previous as recorded; second, there's a missing number we want to interpolate; third, we want to take the difference of it to get to something stationary; fourth, we'd like to add a lagged column. We can do all of those things!
#
# +
# Change freq to next month start
df.index += pd.tseries.offsets.DateOffset(days=1)
df["values"] = df["values"].interpolate(method="time")
df["diff_values"] = df["values"].diff(1)
df["lag_diff_values"] = df["diff_values"].shift(1)
df
# -
# Two other useful time series functions to be aware of are `resample` and `rolling`. `resample` can upsample or downsample time series. Downsampling is by aggregation, eg `df['values].resample('Q').mean()` to downsample to quarterly ('Q') frequency by taking the mean within each quarter. Upsampling involves a choice about how to fill in the missing values; examples of options are `bfill` (backfill) and `ffill` (forwards fill).
#
# Rolling is for taking rolling aggregations, as you'd expect; for example, the 3-month rolling mean of our first difference time series:
df["diff_values"].rolling(3).mean()
#
# ## Method chaining
#
# Sometimes, rather than splitting operations out into multiple lines, it can be more concise and clear to chain methods together. A typical time you might do this is when reading in a dataset and perfoming all of the initial cleaning. Tom Augsperger has a [great tutorial](https://tomaugspurger.github.io/method-chaining) on this, which I've reproduced parts of here. For more info on the `pipe` function used below, check out these short [video tutorials](https://calmcode.io/pandas-pipe/introduction.html).
#
# To chain methods together, both the input and output must be a pandas dataframe. Many functions already do input and output these, for example the `df.rename(columns={'old_col': 'new_col'})` takes in `df` and outputs a dataframe with one column name changed.
#
# But occassionally, we'll want to use a function that we've defined (rather than an already existing one). For that, we need the `pipe` method; it 'pipes' the result of one operation to the next operation. When objects are being passed through multiple functions, this can be much clearer. Compare, for example,
#
# ```python
# f(g(h(df), g_arg=a), f_arg=b)
# ```
#
# that is, dataframe `df` is being passed to function `h`, and the results of that are being passed to a function `g` that needs a key word argument `g_arg`, and the results of *that* are being passed to a function `f` that needs keyword argument `f_arg`. The nested structure is barely readable. Compare this with
#
# ```python
# (df.pipe(h)
# .pipe(g, g_arg=a)
# .pipe(f, f_arg=b)
# )
# ```
#
# Let's see a method chain in action on a real dataset so you get a feel for it. We'll use 1,000 rows of flight data from BTS (a popular online dataset for demos of data cleaning!). TODO use github path. (For further info on method chaining in Python, [see these videos](https://calmcode.io/method-chains/introduction.html)--but be aware they assume advanced knowledge of the language.)
df = pd.read_csv(
"https://github.com/aeturrell/coding-for-economists/raw/main/data/flights1kBTS.csv",
index_col=0,
)
df.head()
# We'll try and do a number of operations in one go: putting column titles in lower case, discarding useless columns, creating precise depature and arrival times, turning some of the variables into categoricals, creating a demeaned delay time, and creating a new categorical column for distances according to quantiles that will be called 'near', 'less near', 'far', and 'furthest'. Some of these operations require a separate function, so we first define those. When we do the cleaning, we'll pipe our dataframe to those functions (optionally passing any arguments).
# +
def extract_city_name(df):
"""
Chicago, IL -> Chicago for origin_city_name and dest_city_name
"""
cols = ["origin_city_name", "dest_city_name"]
city = df[cols].apply(lambda x: x.str.extract("(.*), \w{2}", expand=False))
df = df.copy()
df[["origin_city_name", "dest_city_name"]] = city
return df
def time_to_datetime(df, columns):
"""
Combine all time items into datetimes.
2014-01-01,0914 -> 2014-01-01 09:14:00
"""
df = df.copy()
def converter(col):
timepart = (
col.astype(str)
.str.replace("\.0$", "") # NaNs force float dtype
.str.pad(4, fillchar="0")
)
return pd.to_datetime(
df["fl_date"]
+ " "
+ timepart.str.slice(0, 2)
+ ":"
+ timepart.str.slice(2, 4),
errors="coerce",
)
df[columns] = df[columns].apply(converter)
return df
df = (
df.drop([x for x in df.columns if "Unnamed" in x], axis=1)
.rename(columns=str.lower)
.pipe(extract_city_name)
.pipe(time_to_datetime, ["dep_time", "arr_time"])
.assign(
fl_date=lambda x: pd.to_datetime(x["fl_date"]),
dest=lambda x: pd.Categorical(x["dest"]),
origin=lambda x: pd.Categorical(x["origin"]),
tail_num=lambda x: pd.Categorical(x["tail_num"]),
arr_delay=lambda x: pd.to_numeric(x["arr_delay"]),
op_unique_carrier=lambda x: pd.Categorical(x["op_unique_carrier"]),
arr_delay_demean=lambda x: x["arr_delay"] - x["arr_delay"].mean(),
distance_group=lambda x: (
pd.qcut(x["distance"], 4, labels=["near", "less near", "far", "furthest"])
),
)
)
df.head()
# -
# ### Pyjanitor and more extensive method chaining
#
# Although there's enough support for functional style method chaining to get by, you might find that doing this or that operation isn't always as efficient or as concise as you'd like. Let's look at two examples where this bites and what a wonderful little extension to **pandas** called [**Pyjanitor**](https://pyjanitor.readthedocs.io) does about it.
#
# First, you'll have seen that the syntax for assigning a new column as part of a method chain is relatively fussy, using as it does the `new_column=lambda x: func(x['old_column])` syntax. If you already have a dataframe named `df`, **pyjanitor** gives you the option to create new columns like so (using an example from the documentation):
#
# ```python
#
# df = (
# df.add_columns(
# prop_late_departures=df.num_departing_late / df.total_num_trips,
# prop_late_arrivals=df.num_arriving_late / df.total_num_trips
# )
# )
# ```
# What's great about this is not only that it's cleaner to read, but that all you need do to get the functionality is to import the pyjanitor library and the extra functions will appear magically as options to use on the usual pandas dataframes.
#
# Pyjanitor isn't just about method chaining--it does a lot of other things too, like introducing a bunch of convenience commands with easily understandable verbs behind them. (Though do note it has a hefty download size.)
# ## Reshaping data
#
# The main options for reshaping data are `pivot`, `melt`, `stack`, `unstack`, `pivot_table`, `get_dummies`, `cross_tab`, and `explode`. We’ll look at some of these here.
#
#
# ### Pivoting data from tidy to, err, untidy
#
# At the start of this chapter, I said you should use tidy data--one row per observation, one column per variable--whenever you can. But there are times when you will want to take your lovingly prepared tidy data and pivot it into a wider format. `pivot` and `pivot_table` help you to do that.
#
# 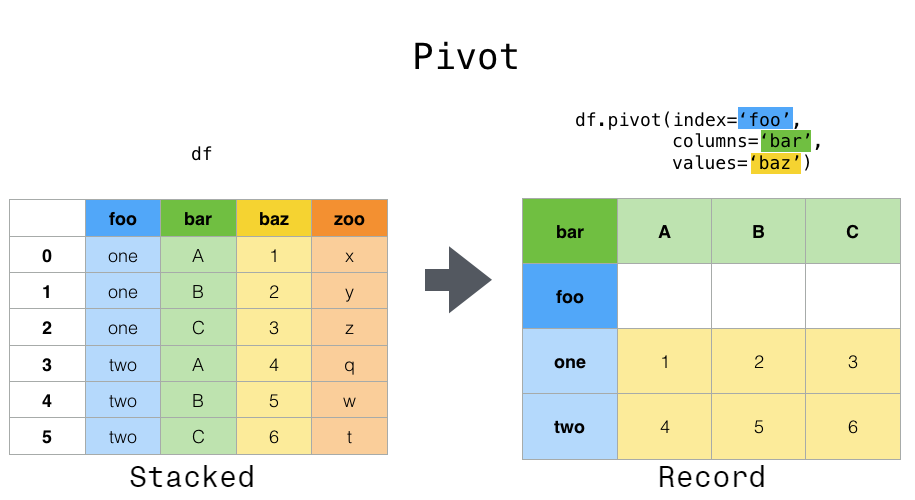
#
# This can be especially useful for time series data, where operations like `shift` or `diff` are typically applied assuming that an entry in one row follows (in time) from the one above. Here's an example:
data = {
"value": np.random.randn(20),
"variable": ["A"] * 10 + ["B"] * 10,
"date": (
list(pd.date_range("1/1/2000", periods=10, freq="M"))
+ list(pd.date_range("1/1/2000", periods=10, freq="M"))
),
}
df = pd.DataFrame(data, columns=["date", "variable", "value"])
df.sample(5)
# If we just run `shift` on this, it's going to shift variable B's and A's together. So we pivot to a wider format (and then we can shift safely).
df.pivot(index="date", columns="variable", values="value").shift(1)
#
# ### Melt
#
# `melt` can help you go from untidy to tidy data (from wide data to long data), and is a *really* good one to remember. Of course, I have to look at the documentation every single time myself, but I'm sure you'll do better.
#
# 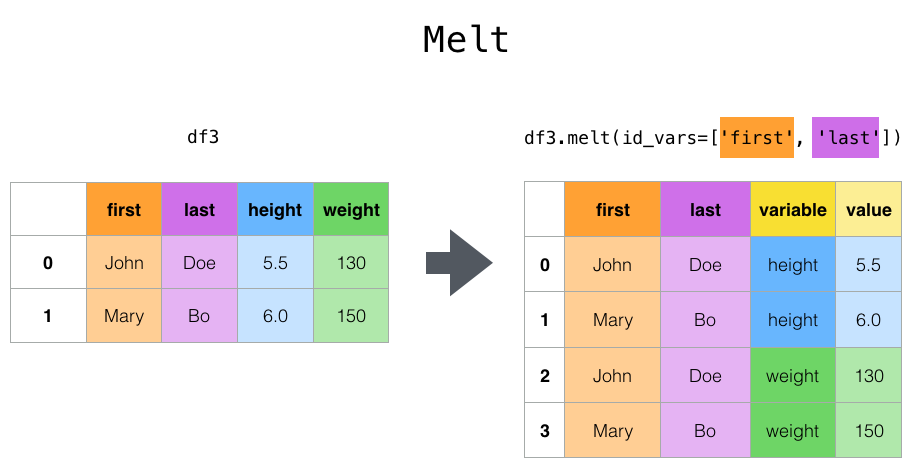
#
# Here's an example of it in action:
df = pd.DataFrame(
{
"first": ["John", "Mary"],
"last": ["Doe", "Bo"],
"height": [5.5, 6.0],
"weight": [130, 150],
}
)
print("\n Unmelted: ")
print(df)
print("\n Melted: ")
df.melt(id_vars=["first", "last"], var_name="quantity")
# If you don't wan the headscratching of melt, there's also `wide_to_long`, which is really useful for typical data cleaning cases where you have data like this:
df = pd.DataFrame(
{
"A1970": {0: "a", 1: "b", 2: "c"},
"A1980": {0: "d", 1: "e", 2: "f"},
"B1970": {0: 2.5, 1: 1.2, 2: 0.7},
"B1980": {0: 3.2, 1: 1.3, 2: 0.1},
"X": dict(zip(range(3), np.random.randn(3))),
"id": dict(zip(range(3), range(3))),
}
)
df
# i.e. data where there are different variables and time periods across the columns. Wide to long is going to let us give info on what the stubnames are ('A', 'B'), the name of the variable that's always across columns (here, a year), any values (X here), and an id column.
pd.wide_to_long(df, ["A", "B"], i="id", j="year")
# ### Stack and unstack
#
# Stack, `stack()` is a shortcut for taking a single type of wide data variable from columns and turning it into a long form dataset, but with an extra index.
#
# 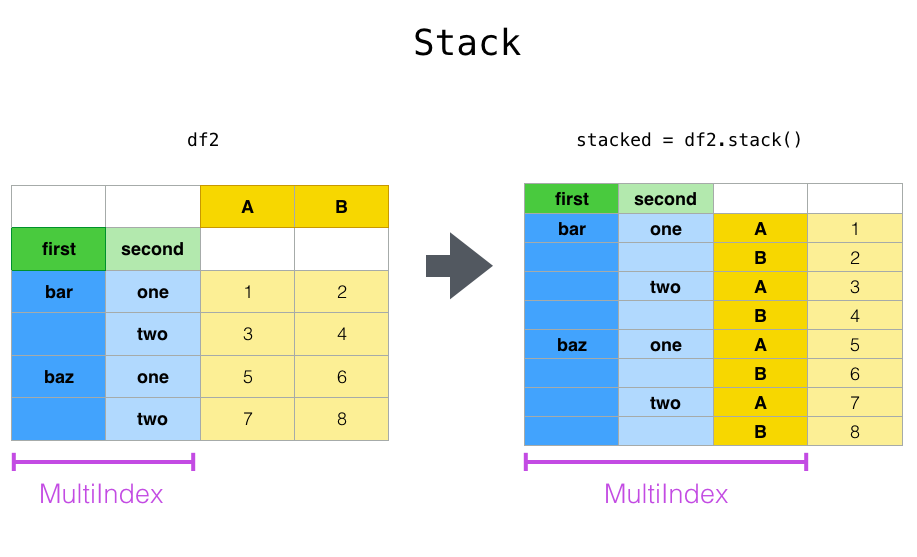
#
# Unstack, `unstack()` unsurprisingly does the same operation, but in reverse.
#
# 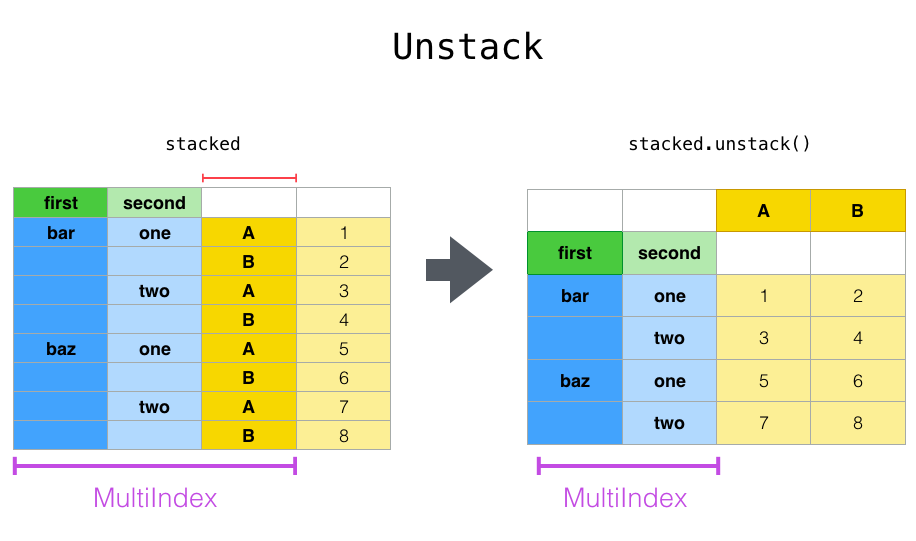
#
# Let's define a multi-index dataframe to demonstrate this:
tuples = list(
zip(
*[
["bar", "bar", "baz", "baz", "foo", "foo", "qux", "qux"],
["one", "two", "one", "two", "one", "two", "one", "two"],
]
)
)
index = pd.MultiIndex.from_tuples(tuples, names=["first", "second"])
df = pd.DataFrame(np.random.randn(8, 2), index=index, columns=["A", "B"])
df
# Let's stack this to create a tidy dataset:
df = df.stack()
df
# Now let's see unstack but, instead of unstacking the 'A', 'B' variables we began with, let's unstack the 'first' column by passing `level=0` (the default is to unstack the innermost index). This diagram shows what's going on:
#
# 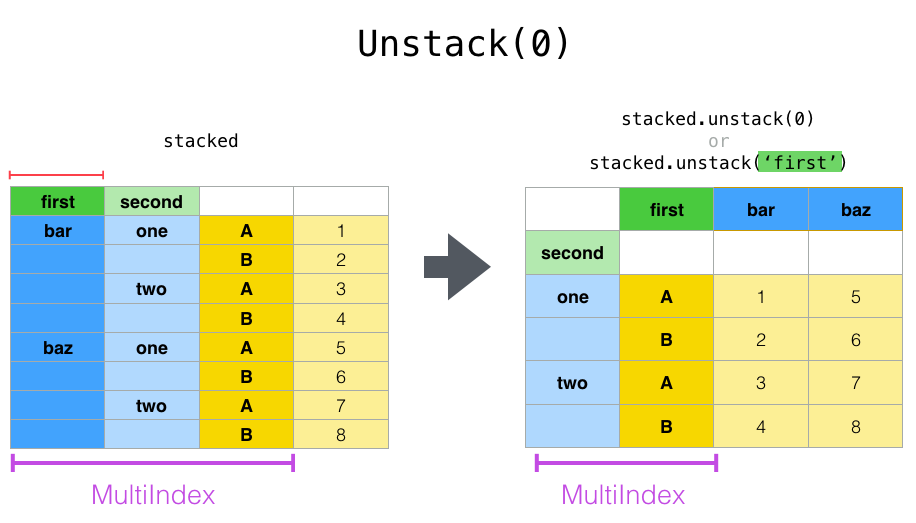
#
# And here's the code:
df.unstack(level=0)
# ### Get dummies
#
# This is a really useful reshape command for when you want (explicit) dummies in your dataframe. When running simple regressions, you can achieve the same effect by declaring the column only be included as a fixed effect, but there are some machine learning packages where converting to dummies may be easier.
#
# Here's an example:
# +
df = pd.DataFrame(
{"group_var": ["group1", "group2", "group3"], "B": ["c", "c", "b"], "C": [1, 2, 3]}
)
print(df)
pd.get_dummies(df, columns=["group_var"])
# -
#
# ## Combining data
#
# **pandas** has a really rich set of options for combining one or more dataframes. The two most important are concatenate and merge.
#
# ### Concatenate
#
# If you have two or more dataframes with the same index or the same columns, you can glue them together into a single dataframe using `pd.concat`.
#
# 
#
# For the same columns, pass `axis=0` to glue the index together; for the same index, pass `axis=1` to glue the columns together. The concatenate function will typically be used on a list of dataframes.
#
# If you want to track where the original data came from in the final dataframe, use the `keys` keyword.
#
# Here's an example using data on two different states' populations that also makes uses of the `keys` option:
# +
base_url = "http://www.stata-press.com/data/r14/"
state_codes = ["ca", "il"]
end_url = "pop.dta"
# This grabs the two dataframes, one for each state
list_of_state_dfs = [pd.read_stata(base_url + state + end_url) for state in state_codes]
# Concatenate the list of dataframes
df = pd.concat(list_of_state_dfs, keys=state_codes, axis=0)
df
# -
# ### Merge
#
# There are so many options for merging dataframes using `pd.merge(left, right, on=..., how=...` that we won't be able to cover them all here. The most important features are: the two dataframes to be merged, what variables (aka keys) to merge on (and these can be indexes) via `on=`, and *how* to do the merge (eg left, right, outer, inner) via `how=`. This diagram shows an example of a merge using keys from the left-hand dataframe:
#
# 
#
# The `how=` keyword works in the following ways:
# - `how='left'` uses keys from the left dataframe only to merge.
# - `how='right'` uses keys from the right dataframe only to merge.
# - `how='inner'` uses keys that appear in both dataframes to merge.
# - `how='outer'` uses the cartesian product of keys in both dataframes to merge on.
#
# Let's see examples of some of these:
left = pd.DataFrame(
{
"key1": ["K0", "K0", "K1", "K2"],
"key2": ["K0", "K1", "K0", "K1"],
"A": ["A0", "A1", "A2", "A3"],
"B": ["B0", "B1", "B2", "B3"],
}
)
right = pd.DataFrame(
{
"key1": ["K0", "K1", "K1", "K2"],
"key2": ["K0", "K0", "K0", "K0"],
"C": ["C0", "C1", "C2", "C3"],
"D": ["D0", "D1", "D2", "D3"],
}
)
# Right merge
pd.merge(left, right, on=["key1", "key2"], how="right")
# Note that the key combination of K2 and K0 did not exist in the left-hand dataframe, and so its entries in the final dataframe are NaNs. But it *does* have entries because we chose the keys from the right-hand dataframe.
#
# What about an inner merge?
pd.merge(left, right, on=["key1", "key2"], how="inner")
# Now we see that the combination K2 and K0 are excluded because they didn't exist in the overlap of keys in both dataframes.
#
# Finally, let's take a look at an outer merge that comes with some extra info via the `indicator` keyword:
pd.merge(left, right, on=["key1", "key2"], how="outer", indicator=True)
# Now we can see that the products of all key combinations are here. The `indicator=True` option has caused an extra column to be added, called '_merge', that tells us which dataframe the keys on that row came from.
#
# For more on the options, see the **pandas** [merging documentation](https://pandas.pydata.org/docs/user_guide/merging.html#database-style-dataframe-or-named-series-joining-merging).
# ## (Advanced) alternatives to **pandas**
#
# Feel free to skip this section if you're just interested in getting going as quickly as possible.
#
# **pandas** isn't the only game in town, not by a long way--though it's by far the best supported and the most fully featured. But it's always good to have options--or, put another way, options have value! Other dataframe libraries may have a syntax that you prefer or provide a speed-up (perhaps in certain situations, for example when working with very large datasets).
#
# If you're specifically interested in how different dataframe options perform on increasingly large datasets, take a look at the benchmarks [here](https://h2oai.github.io/db-benchmark/).
#
# Here's a quick run-through of some alternatives to **pandas**:
# ### Datatable
#
# [**datatable**](https://datatable.readthedocs.io/en/latest/) is another dataframe based way to do analysis, and it has quite a different syntax to **pandas** for data manipulation. **datatable** is very fast, not the fastest dataframe option out there, but it holds its own. Its other major advantage is that it is comfortable running on extremely large (think 50GB) datasets. Most other dataframe packages (including in other languages) cannot cope with this.
#
# I am indebted to the very well-written datatable documentation for the rest of this demonstration.
#
# In datatable, almost all operations are achieved via so-called 'square-bracket notation'. Operations with a (data) Frame are almost all expressed in the form
#
# ```python
# DT[i, j, ...]
# ```
#
# where `DT` is a datatable dataframe, `i` is the row selector, `j` is the column selector, and `...` is a space that could be filled by other commands (potentially several). However, `i` and `j` aren't just positional, as they might be in a **numpy** array or a list, they do a lot more than that.
#
# Datatable allows `i` to be anything that can conceivably be interpreted as a row selector: an integer to select a row, a slice, a range, a list of integers, a list of slices, an expression, a boolean-valued Frame, an integer-valued Frame, an integer **numpy** array, a generator, and more.
#
# Likewise, `j` is also really versatile. It can be used to select a column by name or position, but it will also accept a list of columns, a slice, a list of booleans indicating which columns to pick, an expression, a list of expressions, and a dictionary of expressions. The j expression can even be a python type (such as int or dt.float32), selecting all columns matching that type!
#
# To change an entry (entries), you can use `DT[i, j] = new_value`, and to remove it (or them), it's `del DT[i, j]`.
#
# It's probably going to be easiest to see some examples, so let's do a quick run through of some functions with the storms dataset.
# +
from datatable import dt, f, by, g, join, sort, update, ifelse
DT = dt.fread("https://vincentarelbundock.github.io/Rdatasets/csv/dplyr/storms.csv")
DT.head(5)
# -
# Yes, that's right, you're seeing columns colour coded by data type! Otherwise, what you can see here is that the syntax so far looks quite similar but don't worry because it's about to get weird.
#
# The square brackets accept expressions, i.e. functions. This is achieved through a special import, simply denoted `f`, that says do this thing to the current datatable. For example, to filter by a certain value in the 'status' column (to avoid overloading with data, I'll just use head to show the first few rows):
DT[f.status == "tropical storm", :].head(4)
# Let's get rid of some columns we're not using right now
del DT[:, "year":"long"]
DT.head(4)
# Now, to select only those columns that are strings:
#
DT[:, str].head(4)
# Or to select only those columns that contain 'diameter'
DT[:, [x for x in DT.names if "diameter" in x]].tail(4)
# To create a new column based on an existing one and just look at the last few columns:
DT[:, update(atmospheres=f.pressure / 1013.0)]
DT[:, -4:].tail(4)
# To sort all values by multiple columns:
DT[:, :, sort("wind", "pressure", reverse=[True, True])].head(4)
# And to perform group-by operations:
DT[:, dt.mean(f.wind), by("status")]
# And, finally, one that I always find of great practical use--the within-group transform, with the new column going back into the original data(frame/table). In **pandas**, this is achieved by `transform`, here we use `extend`. Let's demonstrate by showing the maximum wind speed by status group:
DT[:, f[:].extend({"max_wind": dt.min(f.wind)}), by("status")].head(4)
# ### cuDF
#
# [**cuDF**](https://github.com/rapidsai/cudf) is still under development and so doesn't yet have all the features of **pandas**. It's a Python GPU DataFrame library built on the blisteringly fast Apache Arrow columnar memory format. The parts of the library that have been implemented follow the same commands and structure as **pandas**, so it should be easy to use it should you need to.
#
# **cuDF** is *the* fastest dataframe library out there in any of Python, Julia, Rust, or R with the caveats that:
# - this only applies to in-memory datasets, ie datasets smaller than a few GB
# - it doesn't yet do everything
# - it's only currently available on the Linux operating system
# - you need a GPU (and associated software) to use it!
# ### Polars
#
# [**Polars**]() is almost as fast as **cuDF**, but is further down the development path. It also uses Apache Arrow as backend. It currently consists of an 'eager' (for datasets smaller than approximately a few GB) interface that's very similar to **pandas** and a 'lazy' interface (don't worry if you don't know what that means, it's a big data thing) that is somewhat similar to spark (a big data tool). **Polars** is built on the Rust language. It's particularly effective at merging datasets.
# ### Plydata
#
# [Plydata](https://plydata.readthedocs.io/en/stable/index.html) uses the syntax of the **dplyr** package in the R statistical language (the package is a really important part of the data ecosystem in R). It's built around method chaining and everything being in a tidy format, which has pros and cons. Largely, the benefits are in readability and a smaller number of commands to cover the ground that you need to. The downsides are that it plays best with tidy data and when chained methods go wrong it can be more difficult to find out what the issue is.
#
#
# ### dfplyr
#
# [dfplyr](https://github.com/kieferk/dfply) also follows the syntax of the **dplyr** package in R. *Note* that it does not appear to be under active development.
# ## Review
#
# If you know:
#
# - ✅ what tidy data are;
# - ✅ how to create series and dataframes with different datatypes;
# - ✅ how to access values in series and dataframes;
# - ✅ how to perform operations on columns;
# - ✅ how to chain methods;
# - ✅ how to reshape data;
# - ✅ how to combine different dataframes; and
# - ✅ what some alternatives to **pandas** are
#
# then you are well on your way to becoming a data analysis master!
#
| data-intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/shangeth/Google-ML-Academy/blob/master/2-Deep-Neural-Networks/2_5_Bias_Variance.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="AnrLRReKtZ_4"
# <hr>
# <h1 align="center"><a href='https://shangeth.com/courses/'>Deep Learning - Beginners Track</a></h1>
# <h3 align="center">Instructor: <a href='https://shangeth.com/'><NAME></a></h3>
# <hr>
# + [markdown] colab_type="text" id="ihTTDTLYuAu8"
# # Bias & Variance
#
# Let us train a DNN model for a simple regression problem.
# + colab={"base_uri": "https://localhost:8080/", "height": 541} colab_type="code" id="hVSCpKf9tafe" outputId="e15ca2bb-acbd-471c-c644-da00075f2f8f"
import numpy as np
import matplotlib.pyplot as plt
def dataset(show=True):
X = np.arange(-5, 5, 0.01)
y = 8 * np.sin(X) + np.random.randn(1000)
if show:
yy = 8 * np.sin(X)
plt.figure(figsize=(15,9))
plt.scatter(X, y)
plt.plot(X, yy, color='red', linewidth=7)
plt.show()
return X, y
X, y = dataset(show=True)
# + [markdown] colab_type="text" id="zsnVkai4wqDP"
# Lets train 2 models for this dataset
#
# - a very simple linear model
# - a very complex DNN model
# + [markdown] colab_type="text" id="xPTsSruMxD9E"
# ## Simple Linear Model
#
# We are going to split the dataset into 5 groups(random shuffle) and use each of that 5 groups to train 5 different linear models. We will use sklearn's StratifiedKFold to split the dataset into 5. Check the [docs](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.StratifiedKFold.html).
# + colab={"base_uri": "https://localhost:8080/", "height": 541} colab_type="code" id="CTOdFdj87Ck6" outputId="7f986eae-203a-4fed-a784-6d4615833010"
import tensorflow as tf
from tensorflow import keras
import numpy as np
import matplotlib.pyplot as plt
tf.keras.backend.clear_session()
import random
predictions = []
for i in range(5):
idx = random.choices(np.arange(1000), k=700)
X_train, y_train = X[idx], y[idx]
model = tf.keras.Sequential([keras.layers.Dense(units=1, input_shape=[1]) ])
optimizer = tf.keras.optimizers.Adam(lr=0.001)
model.compile(optimizer=optimizer, loss='mean_squared_error')
tf_history = model.fit(X_train, y_train, batch_size=100, epochs=200, verbose=False)
prediction = model.predict(X)
predictions.append(prediction)
plt.figure(figsize=(12,9))
plt.plot(X, predictions[0])
plt.plot(X, predictions[1])
plt.plot(X, predictions[2])
plt.plot(X, predictions[3])
plt.plot(X, predictions[4])
plt.plot(X, 8 * np.sin(X), linewidth=5, label='True curve y')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="seJGd25PB67H"
# ## Deep Neural Network model
# + colab={"base_uri": "https://localhost:8080/", "height": 541} colab_type="code" id="eUqaLDuJCBvX" outputId="bd29400c-1fb1-4cc4-8627-f71f4e6b1028"
import tensorflow as tf
from tensorflow import keras
import numpy as np
import matplotlib.pyplot as plt
tf.keras.backend.clear_session()
import random
predictions = []
for i in range(5):
idx = random.choices(np.arange(1000), k=100)
X_train, y_train = X[idx], y[idx]
model = tf.keras.Sequential([
keras.layers.Dense(units=50, input_shape=[1]),
keras.layers.Activation('relu'),
keras.layers.Dense(units=50),
keras.layers.Activation('relu'),
keras.layers.Dense(units=1),
])
optimizer = tf.keras.optimizers.Adam(lr=0.001)
model.compile(optimizer=optimizer, loss='mean_squared_error')
tf_history = model.fit(X_train, y_train, batch_size=100, epochs=200, verbose=False)
prediction = model.predict(X)
predictions.append(prediction)
plt.figure(figsize=(12,9))
plt.plot(X, predictions[0])
plt.plot(X, predictions[1])
plt.plot(X, predictions[2])
plt.plot(X, predictions[3])
plt.plot(X, predictions[4])
plt.plot(X, 8 * np.sin(X), linewidth=5, label='True curve y')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="UQ_bQW1J1iTy"
# ## Bias
#
# Bias is defined as $ Bias = E[\hat{y}] - y$
#
# It is the difference between the expected value of prediction and the true curve. The expected value will be calculated by splitting the data into n parts and training n model on those n data parts and average of that n model prediction will be expected value.
#
# You can see the bias for first model will be very high as the model predicts a straight line, but the true curve is sinusoidal. But the bias for 2nd model will be lower than 1st model.
#
#
# ## Variance
#
# Variance as you should know defines how much a data is varying.
# $Variance(\hat{y}) = E[(\hat{y} - E[\hat{y}])^2]$
# Although the predictions are not good, but the variance of 2nd model will be higher than 1st model, as the 2nd comple model will try to fit the data more.
#
#
# | Model | Bias | Variance |
# |-------------------- |------ |---------- |
# | Simple Model | High | Low |
# | Very Complex model | Low | High |
# + [markdown] colab_type="text" id="L64aVialQUdP"
# # Bias-Variance Tradeoff
#
# Let's do some math first and discuss about it.
#
# ## Bias-Variance Decomposition
# $MSE = E[(y - \hat{y})^2] = E[y^2 - 2.y.\hat{y} + \hat{y}^2]$
#
# here the random variable is $\hat{y}$ as it is dependent on $X$.
#
# $ MSE = y^2 - 2.y.E[\hat{y}] + E[\hat{y}^2]$
#
# $Bias = E[\hat{y}] - y$
#
# $Bias^2 = (E[\hat{y}] - y)^2 = E[\hat{y}]^2 + y^2 - 2yE[\hat{y}]$
#
# $Variance = E[(\hat{y} - E[\hat{y}])^2] = = E[\hat{y}^2] + E[\hat{y}]^2 - 2E[\hat{y} E[\hat{y}]] = E[\hat{y}^2] + E[\hat{y}]^2 - 2E[\hat{y}]^2 = E[\hat{y}^2] - E[\hat{y}]^2$
#
# $Bias^2 + Variance = y^2 - 2.y.E[\hat{y}] + E[\hat{y}^2] = MSE$
#
# $Bias^2 + Variance = MSE$
#
# - when the bias is high(Simple Model), MSE is high, We don't want high Loss, so **we don't want high bias**
# - when the variance is high(complex model), again MSE is high, so **we don't want high variance**
#
# Conclusion is we need to choose a model which doesn't have high bias or high variance, somthing optimal bias-variance in between will do good.
#
# 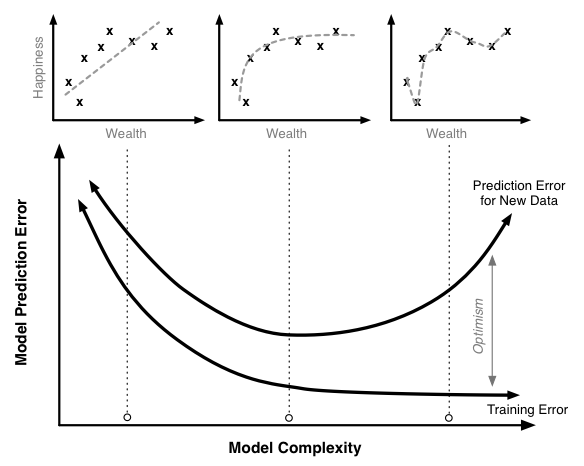
#
# [Image Source](http://scott.fortmann-roe.com)
# + [markdown] colab_type="text" id="C5z6dh27PMjr"
# # Underfitting
#
# When a model have high bias, then the model is **"Underfitting"**.
# Let's see an example first
# + colab={"base_uri": "https://localhost:8080/", "height": 541} colab_type="code" id="qYsrKiG_SZpk" outputId="f1f79dc7-8911-46e7-96fe-65f2c0cf2b8e"
import numpy as np
import matplotlib.pyplot as plt
def dataset(show=True):
X = np.arange(-5, 5, 0.1)
y = 8 * np.sin(X) + np.random.randn(100)
if show:
yy = 8 * np.sin(X)
plt.figure(figsize=(15,9))
plt.scatter(X, y)
plt.plot(X, yy, color='red', linewidth=7)
plt.show()
return X, y
X, y = dataset(show=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="TSHYBB48xbJe" outputId="d09d2335-465e-4b9c-f9cb-3b624bdc7594"
import tensorflow as tf
from tensorflow import keras
import numpy as np
import matplotlib.pyplot as plt
tf.keras.backend.clear_session()
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, shuffle=True)
model = tf.keras.Sequential([keras.layers.Dense(units=1, input_shape=[1]) ])
optimizer = tf.keras.optimizers.Adam(lr=0.001)
model.compile(optimizer=optimizer, loss='mean_squared_error')
tf_history = model.fit(X_train, y_train, batch_size=100, epochs=200, verbose=True, validation_data=(X_test, y_test))
prediction = model.predict(X)
plt.figure(figsize=(12,9))
plt.plot(X, prediction)
plt.plot(X, 8 * np.sin(X), linewidth=5, label='True curve y')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="CrTo-T-KRI_d"
# You can see the Training data loss and Validation data loss both are bad, the model performance is not good. This is called Underfitting.
#
# Underfitting may happen because the model is not complex enough, or need more training. So, using a deeper network or training for more time may help.
# + [markdown] colab_type="text" id="RSmnmkCmRheA"
# # Overfitting
#
# Let's train a more complex model with less training data.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="P_xa0uUIPsW2" outputId="a5dc8e8a-8e6c-48c3-d05d-b7454d0c98df"
import tensorflow as tf
from tensorflow import keras
import numpy as np
import matplotlib.pyplot as plt
tf.keras.backend.clear_session()
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.9, shuffle=True)
model = tf.keras.Sequential([
keras.layers.Dense(units=50, input_shape=[1]),
keras.layers.Activation('relu'),
keras.layers.Dense(units=50),
keras.layers.Activation('relu'),
keras.layers.Dense(units=1),
])
optimizer = tf.keras.optimizers.Adam(lr=0.001)
model.compile(optimizer=optimizer, loss='mean_squared_error')
tf_history = model.fit(X_train, y_train, batch_size=100, epochs=1000, verbose=True, validation_data=(X_test, y_test))
prediction = model.predict(X_train)
plt.figure(figsize=(12,9))
plt.scatter(X_train, prediction,label='Training Data Prediction')
plt.scatter(X_test, model.predict(X_test), color='r', marker='x', label='Test Data Prediction')
plt.plot(X, 8 * np.sin(X), linewidth=1, label='True curve y')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="B3It44w0T-GE"
# Here you can see, although the model is comple and can learn more complex featreus of the data, the Validation loss is way higher than training loss. This is called Overfitting. This means the model fits the training data so much that it does not generalize and perform very poor in new unseen data. Adding more data can help to prevent overfitting.
| 2-Deep-Neural-Networks/2_5_Bias_Variance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Task: Quantum Circuit Simulator
#
# The goal here is to implement simple quantum circuit simulator.
#
# ## Introduction
#
# Before we start coding:
#
#
# ### Qubit
#
# Qubit is the basic unit of quantum information. It is a two-state (or two-level) quantum-mechanical system, and can be represented by a linear superposition of its two orthonormal basis states (or basis vectors). The vector representation of a single qubit is: ${\vert a\rangle =v_{0} \vert 0\rangle +v_{1} \vert 1\rangle \rightarrow {\begin{bmatrix}v_{0}\\v_{1}\end{bmatrix}}}$,
# Here, ${\displaystyle v_{0}}v_{0}$ and ${\displaystyle v_{1}}v_{1}$ are the complex probability amplitudes of the qubit. These values determine the probability of measuring a 0 or a 1, when measuring the state of the qubit.
#
# Code:
# +
# Qubit in |0> state (100% probability of measuring 0)
q0 = [1, 0]
# Qubit in |1> state (100% probability of measuring 1)
q1 = [0, 1]
# Qubit |+> state (superposition: 50% probability of measuring 0 and 50% probability of measuring 1)
q2 = [0.7071067811865475, 0.7071067811865475]
# Qubit |-> state (superposition: 50% probability of measuring 0 and 50% probability of measuring 1) with phase pi
q3 = [0.7071067811865475, -0.7071067811865475]
# Qubit |i> state (superposition: 50% probability of measuring 0 and 50% probability of measuring 1) with phase pi/2
q3 = [0.7071067811865475, 0+0.7071067811865475j]
# Qubit |-i> state (superposition: 50% probability of measuring 0 and 50% probability of measuring 1) with phase -pi/2
q4 = [0.7071067811865475, 0-0.7071067811865475j]
# -
# Note that vector contains probability amplitudes - not probabilities. Probability amplitude is complex number and can be negative. Probability is calculated as absolute value squared:
# +
import numpy as np
q4 = np.array([0.7071067811865475+0j, 0-0.7071067811865475j])
p4 = np.abs(q4)**2
print(p4)
# -
# ### State vector
#
# The combined state of multiple qubits is the tensor product of their states (vectors). The tensor product is denoted by the symbol ${\displaystyle \otimes }$.
#
# The vector representation of two qubits is:
#
# ${\displaystyle \vert ab\rangle =\vert a\rangle \otimes \vert b\rangle =v_{00}\vert 00\rangle +v_{01}\vert 01\rangle +v_{10}\vert 10\rangle +v_{11}\vert 11\rangle \rightarrow {\begin{bmatrix}v_{00}\\v_{01}\\v_{10}\\v_{11}\end{bmatrix}}}$
#
# Example:
# +
# Qubit in |0> state (100% probability of measuring 0)
q0 = [1, 0]
# Qubit in |1> state (100% probability of measuring 1)
q1 = [0, 1]
combined_state = np.kron(q0, q1)
print(combined_state)
# -
# Now, what this vector tells us?
#
# It will be more clear if we write vector elements in a column with index expressed in binary format:
#
# ```
# Index (dec) Index (bin) Amplitude Probability
# ================================================
# 0 00 0 0 ( 0%)
# 1 01 1 1 (100%)
# 2 10 0 0 ( 0%)
# 3 11 0 0 ( 0%)
# ```
#
# - First element (binary: 00) is probability of measuring 0 on both qubits.
# - Second element (binary: 01) is probability of measuring 0 on first qubit and 1 on second qubit.
# - Third element (binary: 10) is probability of measuring 1 on first qubit and 0 on second qubit.
# - Fourth element (binary: 11) is probability of measuring 1 on both qubits.
#
# #### Endianness
#
# It is important to say that different quantum programming frameworks use different orientation of bitstrings (endianness). In previous example, left bit belongs to first qubit and righ bit belongs to second qubit. This enconding is called "big endian".
#
# But, in some frameworks (like Qiskit), encoding is opposite: rightmost bit belongs to first qubit and leftmost bit belongs to last qubit. This is called "little endian".
#
# So, vector from our example in Qiskit's "little endian" encoding will look like this:
#
# ```
# Index (dec) Index (bin) Amplitude Probability
# ================================================
# 0 00 0 0 ( 0%)
# 1 01 0 0 ( 0%)
# 2 10 1 1 (100%)
# 3 11 0 0 ( 0%)
# ```
#
# "Little endian" encoding:
#
# - First element (binary: 00) is probability of measuring 0 on both qubits.
# - Second element (binary: 01) is probability of measuring 0 on second qubit and 1 on first qubit.
# - Third element (binary: 10) is probability of measuring 1 on second qubit and 0 on first qubit.
# - Fourth element (binary: 11) is probability of measuring 1 on both qubits.
#
# # Quantum gates
#
# Quantum gates are basic units of quantum processing. Gates are represented as unitary matrices. The action of the gate on a specific quantum state is found by multiplying the vector ${\displaystyle \vert \psi _{1}\rangle }$ which represents the state, by the matrix ${\displaystyle U}$ representing the gate. The result is a new quantum state ${\displaystyle \vert \psi _{2}\rangle }$
#
# ${\displaystyle U\vert \psi _{1}\rangle =\vert \psi _{2}\rangle }$
#
# Quantum gates (usually) act on small number of qubits. We have single-qubit and multi-qubit gates. n-qubit gate is represented as $2^n\times2^n$ unitary matrix.
#
# Examples:
#
# #### Single qubit gates
#
# X (aka NOT) gate:
#
# $X = \begin{bmatrix} 0 & 1 \\ 1 & 0 \end{bmatrix}$
#
# Hadamard gate:
#
# $H = \tfrac{1}{\sqrt{2}}\begin{bmatrix} 1 & 1 \\ 1 & -1 \end{bmatrix}$
#
# General single qubit rotation gate:
#
# $U_3(\theta, \phi, \lambda) = \begin{bmatrix} \cos(\theta/2) & -e^{i\lambda}\sin(\theta/2) \\
# e^{i\phi}\sin(\theta/2) & e^{i\lambda+i\phi}\cos(\theta/2)
# \end{bmatrix}$
#
# #### Two-qubit gates:
#
# Controlled-X (aka CNOT) gate:
#
# ${CNOT} = \begin{bmatrix} 1 & 0 & 0 & 0 \\
# 0 & 1 & 0 & 0 \\
# 0 & 0 & 0 & 1 \\
# 0 & 0 & 1 & 0 \\
# \end{bmatrix}$
#
# SWAP gate:
#
# ${SWAP} = \begin{bmatrix} 1 & 0 & 0 & 0 \\
# 0 & 0 & 1 & 0 \\
# 0 & 1 & 0 & 0 \\
# 0 & 0 & 0 & 1 \\
# \end{bmatrix}$
#
# #### Examples
#
# Let's see how single-qubit gate modifies state of the qubit:
# +
import numpy as np
# Let's start with qubit in state |0> (100% probability of measuring 0)
q0 = np.array([1, 0])
print("Initial state:\t", q0)
# Define X (NOT) gate:
X = np.array([
[0, 1],
[1, 0]
])
# Now apply X gate to a qubit (matrix-vector dot product):
q0 = np.dot(X, q0)
print("Final state:\t", q0)
# -
# After applying X gate, qubit flips from state $|0\rangle$ to state $|1\rangle$.
#
# Now, let's see how Hadamard gate works:
# +
import numpy as np
# Let's start with qubit in state |0> (100% probability of measuring 0)
q0 = np.array([1, 0])
print("Initial state:\t", q0)
# Define H (Hadamard) gate:
H = np.array([
[1/np.sqrt(2), 1/np.sqrt(2)],
[1/np.sqrt(2), -1/np.sqrt(2)]
])
# Now apply H gate to a qubit (matrix-vector dot product):
q0 = np.dot(H, q0)
print("Final state:\t", q0)
# -
# After applying Hadamard gate on qubit in state $|0\rangle$ it evolves to state $|+\rangle$ which is equal superposition.
# ### Matrix operator
#
# Quantum program eveloves quantum state by multiplying state vector with each gate's unitary matrix (dot product). Note that dimension of the state vector and dimension of the unitary matrix describing a gate usually don't match. For example: 3-qubit quantum circuit's state vector has $2^n=2^3=8$ elements, but single-qubit gate has $2^n\times2^n=2^1\times2^1=2\times2$ elements. In order to perform matrix-vector multiplication, we need to "resize" gate's matrix to the dimension of the state vector. Let's call that matrix a **matrix operator**.
#
# Note that size of the matrix operator is $2^n\times2^n$ where $n$ is total number of qubits in the circuit, so storing it into memory and calculating it requires a lot of memory and cpu power for bigger circuits. Optimizing this code is most interesting and challenging part, but for our purpose it is enough if you make it work smoothly with 8 qubits (the more - the better).
#
# #### Matrix operator for single-qubit gates
#
# Matrix operator for single-qubit gate can be calculated by performing tensor product of gate's unitary matrix and $2\times2$ identity matrices in correct order.
#
# Example for single-qubit gate $U$ in 3-qubit circuit:
#
# - gate on qubit 0: ${O=U \otimes I \otimes I}$
# - gate on qubit 1: ${O=I \otimes U \otimes I}$
# - gate on qubit 2: ${O=I \otimes I \otimes U}$
#
# Example matrix operator for X gate acting on third qubit in 3-qubit circuit can be calculated like this:
# +
import numpy as np
# Let's define state vector of the 3-qubit circuit in "ground state" (all qubits in state |0>)
psi = [1, 0, 0, 0, 0, 0, 0, 0]
print("Initial state:", psi)
# Define X (NOT) gate:
X = np.array([
[0, 1],
[1, 0]
])
# Define 2x2 identity
I = np.identity(2)
# Calculate operator for X gate acting on third qubit in 3-qubit circuit
O = np.kron(np.kron(I, I), X)
print("\nOperator:\n\n", O, "\n")
# And finally, apply operator
psi = np.dot(psi, O)
print("Final state:", psi)
# -
# We are dealing with "big endian" encoding, so this result is correct: third qubit is flipped to state $|1\rangle$ and other qubits are not changed.
#
# **Note**: if we want vector in "little endian" encoding (like Qiskit), then order in which we perform tensor product to calculate operator is opposite. Instead ${O=I \otimes I \otimes U}$ we would do ${O=U \otimes I \otimes I}$.
# #### Matrix operator for multi-qubit gates
#
# If we want to apply two qubit gate on subsequent qubits ( 0-1, 1-2, 2-3 etc.) then we can use the same technique like we do with single qubit gates:
#
# For 3-qubit circuit, CNOT gate:
#
# - acting on first and second qubit, operator is ${O=CNOT \otimes I}$
#
# - acting on second and third qubit, operator is ${O=I \otimes CNOT}$
#
# But, multi-qubit gates can be applied to qubits which are not consequent, so this is not that trivial.
#
# The main feature of a controlled-$U$ operation, for any unitary $U$, is that it (coherently) performs an operation on some qubits depending on the value of some single qubit. The way that we can write this explicitly algebraically (with the control on the first qubit) is:
#
# $\mathit{CU} \;=\; \vert{0}\rangle\!\langle{0}\vert \!\otimes\! \mathbf 1 \,+\, \vert{1}\rangle\!\langle{1}\vert \!\otimes\! U$
#
# where ${\mathbf 1}$ is an identity matrix of the same dimension as $U$. Here, ${\ket{0}\!\bra{0}}$ and ${\ket{1}\!\bra{1}}$ are projectors onto the states ${\ket{0}}$ and ${\ket{1}}$ of the control qubit — but we are not using them here as elements of a measurement, but to describe the effect on the other qubits depending on one or the other subspace of the state-space of the first qubit.
#
# We can use this to derive the matrix for the gate ${\mathit{CX}_{1,3}}$ which performs an $X$ operation on qubit 3, coherently conditioned on the state of qubit 1, by thinking of this as a controlled-${(\mathbf 1_2 \!\otimes\! X)}$ operation on qubits 2 and 3:
#
# $\begin{aligned}
# \mathit{CX}_{1,3} \;&=\;
# \vert{0}\rangle\!\langle{0}\vert \otimes \mathbf 1_4 \,+\, \vert{1}\rangle\!\langle{1}\vert \otimes (\mathbf 1_2 \otimes X)
# \\[1ex]&=\;
# \begin{bmatrix}
# \mathbf 1_4 & \mathbf 0_4 \\
# \mathbf 0_4 & (\mathbf 1_2 \!\otimes\! X)
# \end{bmatrix}
# \;=\;
# \begin{bmatrix}
# \mathbf 1_2 & \mathbf 0_2 & \mathbf 0_2 & \mathbf 0_2 \\
# \mathbf 0_2 & \mathbf 1_2 & \mathbf 0_2 & \mathbf 0_2 \\
# \mathbf 0_2 & \mathbf 0_2 & X & \mathbf 0_2 \\
# \mathbf 0_2 & \mathbf 0_2 & \mathbf 0_2 & X
# \end{bmatrix},
# \end{aligned}$
#
# where the latter two are block matrix representations to save on space (and sanity).
#
# Better still: we can recognise that — on some mathematical level where we allow ourselves to realise that the order of the tensor factors doesn't have to be in some fixed order — the control and the target of the operation can be on any two tensor factors, and that we can fill in the description of the operator on all of the other qubits with $\mathbf 1_2$. This would allow us to jump straight to the representation
#
# $\begin{aligned}
# \mathit{CX}_{1,3} \;&=&\;
# \underbrace{\vert{0}\rangle\!\langle{0}\vert}_{\text{control}} \otimes \underbrace{\;\mathbf 1_2\;}_{\!\!\!\!\text{uninvolved}\!\!\!\!} \otimes \underbrace{\;\mathbf 1_2\;}_{\!\!\!\!\text{target}\!\!\!\!}
# &+\,
# \underbrace{\vert{1}\rangle\!\langle{1}\vert}_{\text{control}} \otimes \underbrace{\;\mathbf 1_2\;}_{\!\!\!\!\text{uninvolved}\!\!\!\!} \otimes \underbrace{\; X\;}_{\!\!\!\!\text{target}\!\!\!\!}
# \\[1ex]&=&\;
# \begin{bmatrix}
# \mathbf 1_2 & \mathbf 0_2 & \phantom{\mathbf 0_2} & \phantom{\mathbf 0_2} \\
# \mathbf 0_2 & \mathbf 1_2 & \phantom{\mathbf 0_2} & \phantom{\mathbf 0_2} \\
# \phantom{\mathbf 0_2} & \phantom{\mathbf 0_2} & \phantom{\mathbf 0_2} & \phantom{\mathbf 0_2} \\
# \phantom{\mathbf 0_2} & \phantom{\mathbf 0_2} & \phantom{\mathbf 0_2} & \phantom{\mathbf 0_2}
# \end{bmatrix}
# \,&+\,
# \begin{bmatrix}
# \phantom{\mathbf 0_2} & \phantom{\mathbf 0_2} & \phantom{\mathbf 0_2} & \phantom{\mathbf 0_2} \\
# \phantom{\mathbf 0_2} & \phantom{\mathbf 0_2} & \phantom{\mathbf 0_2} & \phantom{\mathbf 0_2} \\
# \phantom{\mathbf 0_2} & \phantom{\mathbf 0_2} & X & \mathbf 0_2 \\
# \phantom{\mathbf 0_2} & \phantom{\mathbf 0_2} & {\mathbf 0_2} & X
# \end{bmatrix}
# \end{aligned}$
#
# and also allows us to immediately see what to do if the roles of control and target are reversed:
#
# $\begin{aligned}
# \mathit{CX}_{3,1} \;&=&\;
# \underbrace{\;\mathbf 1_2\;}_{\!\!\!\!\text{target}\!\!\!\!} \otimes \underbrace{\;\mathbf 1_2\;}_{\!\!\!\!\text{uninvolved}\!\!\!\!} \otimes \underbrace{\vert{0}\rangle\!\langle{0}\vert}_{\text{control}}
# \,&+\,
# \underbrace{\;X\;}_{\!\!\!\!\text{target}\!\!\!\!} \otimes \underbrace{\;\mathbf 1_2\;}_{\!\!\!\!\text{uninvolved}\!\!\!\!} \otimes \underbrace{\vert{1}\rangle\!\langle{1}\vert}_{\text{control}}
# \\[1ex]&=&\;
# {\scriptstyle\begin{bmatrix}
# \!\vert{0}\rangle\!\langle{0}\vert\!\! & & & \\
# & \!\!\vert{0}\rangle\!\langle{0}\vert\!\! & & \\
# & & \!\!\vert{0}\rangle\!\langle{0}\vert\!\! & \\
# & & & \!\!\vert{0}\rangle\!\langle{0}\vert
# \end{bmatrix}}
# \,&+\,
# {\scriptstyle\begin{bmatrix}
# & & \!\!\vert{1}\rangle\!\langle{1}\vert\!\! & \\
# & & & \!\!\vert{1}\rangle\!\langle{1}\vert \\
# \!\vert{1}\rangle\!\langle{1}\vert\!\! & & & \\
# & \!\!\vert{1}\rangle\!\langle{1}\vert & &
# \end{bmatrix}}
# \\[1ex]&=&\;
# \left[{\scriptstyle\begin{matrix}
# 1 & 0 & 0 & 0 \\
# 0 & 0 & 0 & 0 \\
# 0 & 0 & 1 & 0 \\
# 0 & 0 & 0 & 0 \\
# 0 & 0 & 0 & 0 \\
# 0 & 1 & 0 & 0 \\
# 0 & 0 & 0 & 0 \\
# 0 & 0 & 0 & 1
# \end{matrix}}\right.\,\,&\,\,\left.{\scriptstyle\begin{matrix}
# 0 & 0 & 0 & 0 \\
# 0 & 1 & 0 & 0 \\
# 0 & 0 & 0 & 0 \\
# 0 & 0 & 0 & 1 \\
# 1 & 0 & 0 & 0 \\
# 0 & 0 & 0 & 0 \\
# 0 & 0 & 1 & 0 \\
# 0 & 0 & 0 & 0
# \end{matrix}}\right].
# \end{aligned}$
#
# But best of all: if you can write down these operators algebraically, you can take the first steps towards dispensing with the giant matrices entirely, instead reasoning about these operators algebraically using expressions such as $\mathit{CX}_{1,3} =
# \vert{0}\rangle\!\langle{0}\vert \! \otimes\!\mathbf 1_2\! \otimes\! \mathbf 1_2 +
# \vert{1}\rangle\!\langle{1}\vert \! \otimes\! \mathbf 1_2 \! \otimes\! X$
# and
# $\mathit{CX}_{3,1} =
# \mathbf 1_2 \! \otimes\! \mathbf 1_2 \! \otimes \! \vert{0}\rangle\!\langle{0}\vert +
# X \! \otimes\! \mathbf 1_2 \! \otimes \! \vert{1}\rangle\!\langle{1}\vert$.
#
#
# For Example, let's calculate operator for Controlled-X (CNOT) on first qubit as control and third qubit as target in 3 qubit quantum circuit:
# +
import numpy as np
# Define X gate (CNOT is controlled-X):
X = np.array([
[0, 1],
[1, 0]
])
# Define 2x2 Identity
I = np.identity(2)
# Define projection operator |0><0|
P0x0 = np.array([
[1, 0],
[0, 0]
])
# Define projection operator |1><1|
P1x1 = np.array([
[0, 0],
[0, 1]
])
# And now calculate our operator:
O = np.kron(np.kron(P0x0, I), I) + np.kron(np.kron(P1x1, I), X)
print("CNOT(0, 2) for 3-qubit circuit, operator is:\n")
print(O)
O = (np.kron(P0x0, I)) + np.kron(P1x1, X)
# -
# In order to implement simulator, it is best if you have function which returns operator for any unitary targeting any qubit(s) for any circuit size, something like:
#
# ```
# get_operator(total_qubits, gate_unitary, target_qubits)
# ```
#
# But this is not trivial so **it is enough if you can implement it for any 1-qubit gates and CNOT only.**
#
# If you are still enthusiastic and you wish to implement universal operator function then please refer to:
#
# - [qosf-simulator-task-additional-info.pdf](https://github.com/quantastica/qosf-mentorship/blob/master/qosf-simulator-task-additional-info.pdf)
#
#
# - Book *<NAME>.; <NAME> (2000). Quantum Computation and Quantum Information, 10th Anniversary Edition, Section 8.2.3, Operator-Sum Representation*
# ### Measurement
#
# State vector of the real quantum computer cannot be directly observed. All we can read out of qubit is a single classical bit. So best we can get as output from quantum computer is bitstring of size $n$ where $n$ is number of qubits. Reading the state from a qubit is called "measurement". When qubit is in superposition, measurment puts qubit in one of two classical states. If we read 1 from qubit, it will "collapse" to state |1> and will stay there - superposition is "destroyed", and any subsequent measurement will return the same result.
#
# Measurement is non-unitary operation on the state vector. But for simplicity, and because we can access state vector of our simulator, it is easier if we do it with a trick:
#
# We can simulate measurement by choosing element from the state vector with weighted random function. Elements with larger probability amplitude will be returned more often, and elements with smaller probability amplitude will be returned less often. Elements with zero probability will never be returned.
#
# For example, this state vector:
#
# ```
# Index (dec) Index (bin) Amplitude Probability
# =================================================
# 0 00 0.70710678 0.5 (50%)
# 1 01 0 0 ( 0%)
# 2 10 0 0 ( 0%)
# 3 11 0.70710678 0.5 (50%)
# ```
#
# Our random function should return elements 00 and 11 equaly often and never return 01 and 10. If we execute it 1000 times (1000 shots) we should get something like this:
#
# ```
# {
# "00": 494,
# "11": 506
# }
# ```
# *(this is random, so it usually is not exactly 500/500 and that is completelly fine)*
#
# ## Requirements
#
# It is expected that simulator can perform following:
#
# - initialize state
#
# - read program, and for each gate:
# - calculate matrix operator
# - apply operator (modify state)
#
# - perform multi-shot measurement of all qubits using weighted random technique
#
# It is up to you how you will organize code, but this is our suggestion:
#
# ### Input format (quantum program)
#
# It is enough if simulator takes program in following format:
#
# ```
# [
# { "unitary": [[0.70710678, 0.70710678], [0.70710678, -0.70710678]], "target": [0] },
# { "unitary": [ [1, 0, 0, 0], [0, 1, 0, 0], [0, 0, 0, 1], [0, 0, 1, 0] ], "target": [0, 1] }
# ...
# ]
# ```
#
# Or, you can define unitaries in the code, and accept program with gate names instead:
#
# ```
# [
# { "gate": "h", "target": [0] },
# { "gate": "cx", "target": [0, 1] }
# ...
# ]
# ```
#
# ### Simulator program
#
# Not engraved into stone, but you can do something like this:
# ## Extra Files for the simulator task
#
# For my simulator proposal I need some extra documents which I added in the notebook to handle in a general way the different processes and classes involved in the circuits, these are:
#
# <ul>
# <li>gates_basic.py</li>
# <li>find_gate.py</li>
# <li>gates_x.py</li>
# <li>gates_h.py</li>
# <li>gates_swap.py</li>
# <li>operator_size.py</li>
# </ul>
#
# ##### Note: those files were added in the notebook to make it more accessible to understand the project.
#
# +
########### gates_basic.py ######################
#### This static class is intended ######
#### to define in methods the base ######
#### matrices of size 2x2: ######
#### -projection of the operator |0><0|, ######
#### -projection of the operator |1><1|, ######
#### -the Identity, ######
#### -and U3(theta,phi,landa). ######
##################################################
#### define the four method for the basic matrices of size 2x2
#### in the static class gates_basic
#### using import numpy as np library for the matrices
class gates_basic:
### Define of the operator |0><0|
def p0x0():
return np.array([[1, 0], ## return the matrix
[0, 0]]) ## equals to |0><0|
### Define of the operator |0><0|
def p1x1():
return np.array([[0, 0], ## return the matrix
[0, 1]]) ## equals to |1><1|
## Define the Identity matriz of size 2x2
def i():
return np.array([[1, 0], ## return the identity
[0, 1]]) ## matrix of 2x2
## Define the U3 matriz of size 2x2
## This matrix needs three angles:
## -theta
## -lambda
## -ph
## and works with complex numbers (0+1j = a+bi mathematic expresion)
## in order to modify the qubits scalars values (alpha and beta)
def u3(theta,lambda_,phi):
return np.array([[np.cos(theta/2), -np.exp(0+1j * lambda_) * np.sin(theta / 2)],
[np.exp(0+1j * phi) * np.sin(theta / 2),
np.exp(0+1j * lambda_ + 0+1j * phi) * np.cos(theta / 2)]])
### this matrix it was define in the tutorial part of this notebook.
# +
########### find_gate.py ######################
#### It consists of a static class that ######
#### has different methods to indicate ######
#### in the case of using multiple ######
#### qubits how to perform the tensor ######
#### product for 2x2 and 4x4 gates ######
#### with its different possibilities. ######
################################################
#### define the method fwhen needs
#### the product tensfor for matrices of size 2x2;
#### when are using matrices of size 4x4 with tensor product
#### in the original and inverse matrix;
#### finally all the posibilities when use a matrix of size 4x4
#### using qubits moe than 2.
class find_gate:
#### Define the method when is neccesary apply the tensor product
#### in a matrix of size 2x2
### needs the number of qubits, the qubit to modify
### and the matrix of size 2x2.
def gate_1xn(total_qubits,target,gate):
operators = [] ## using a list for every gate by qubit
for j in range(total_qubits): ## iteration on all the qubits
if j == target: ## when find the value for the qubit
operators.append(gate) ## to apply the gate we append in the list
else: ## in other case we append the identity gate
operators.append(gates_basic.i())
O_state = operators[0] ## when finish the iterative for
for j in range (1,len(operators)): ## whe apply the tensor product
O_state = np.kron(operators[j],O_state) ## for all the gates o size 2x2
return O_state ## return the matriz result of size 2**nẍ2**n
## where n = total_qubits
#### The method of constructing a CU gate
#### by considering the U matriz of size 2x2,
#### the target qubit and the control qubit.
#### when control qubit is less than target qubit
def gate_i_i2xn(target_0,target_1,gate): ## is neccesary two parts to build the gate
left_part = gates_basic.i() ## left part is initialized with an identity gate
right_part = gates_basic.i() ## right part is initialized with an identity gate
for j in range(target_0,target_1): ## iterative function to find the gates
## between target and controls qubits positions
if j == target_0: ## when find the control qubit value
left_part = np.kron(gates_basic.p0x0(),left_part) ## apply the tensor product
right_part = np.kron(gates_basic.p1x1(),right_part) ## between left part and |0><0!
## and right_part with |1><1| and assign
## in its respectively variable left_part or right part
elif j == target_1-1: ## When find the target qubit position we apply
left_part = np.kron(left_part,gates_basic.i()) ## product tensor between left and
right_part = np.kron(right_part,gate) ## right part with Identity and
## U gate rrespectively
else: ## in other case apply on left_part and right_part
left_part = np.kron(left_part,gates_basic.i()) ## the tensor product with
right_part = np.kron(right_part,gates_basic.i()) ## identities gate of size 2x2
O_state = left_part + right_part ## finally, adder the left_part matrix with right_part matrix
return O_state ## return the final matrix
#### The method of constructing a CU gate
#### by considering the U matriz of size 2x2,
#### the target qubit and the control qubit.
#### when control qubit is more than target qubit
def gate_i_i2xn_inv(target_0,target_1,gate):
left_part = gates_basic.i() ## left part is initialized with an identity gate
right_part = gates_basic.i() ## right part is initialized with an identity gate
for j in range(target_0,target_1): ## iterative function to find the gates
## between target and controls qubits positions
if j == target_1-1: ## When find the Control qubit position we apply
left_part = np.kron(left_part,gates_basic.p0x0()) ## apply the tensor product
right_part = np.kron(right_part,gates_basic.p1x1()) ## between left part and |0><0!
## and right_part with |1><1| and assign
## in its respectively variable left_part or right part
elif j == target_0: ## When find the target qubit position we apply
left_part = np.kron(gates_basic.i(),left_part) ## product tensor between left and
right_part = np.kron(gate,right_part) ## right part with Identity and
## U gate rrespectively
else: ## in other case apply on left_part and right_part
left_part = np.kron(left_part,gates_basic.i()) ## the tensor product with
right_part = np.kron(right_part,gates_basic.i()) ## identities gate of size 2x2
O_state = left_part + right_part ## finally, adder the left_part matrix with right_part matrix
return O_state ## return the final matrix
#######################################################################
########## For the version 2: #######
########## The gate_i_i2xn() and gate_i_i2xn_inv() methods #######
#########$ can work in a single method under certain conditions #######
#######################################################################
#### In case of having more than 3 qubits and the CU matrix is obtained,
#### within the simulator, the tensor product must be completed
#### in order to work correctly using the total_qubits,
#### the CU matrix and the qubit's position of the target and control qubits
def gate_moving(total_qubits,target_0,target_1,O_state):
if target_1 == 0: ## When control qubit is in the position 0
O_state = find_gate.gate_1xn(total_qubits-target_0,target_1,O_state)
## consider
## apply the same idea of tensor product
## with a gate of sie 2x2
elif target_0 == total_qubits-1: ## When target qubit is in the final qubit position
O_state = find_gate.gate_1xn(target_1+1,target_1,O_state)
## apply the same idea of tensor product
## with a gate of sie 2x2
elif target_1 != 0 and target_0 != total_qubits-1: ## when the gate is between
## the first and last qubit
O_state = find_gate.gate_1xn(total_qubits-target_0 + target_1,target_1,O_state)
## for identify the remaining qubits
return O_state ## return the final matrix after the tensor product
# +
########### gates_x.py ##############################
#### The operators X and CX are defined ######
#### and considering all the possibilities ######
#### of being carried out from one qubit or ######
#### two depending on the matrix up to N qubits ######
#### and may correspond to the expected result. ######
######################################################
#### define the methods for the posibilities for X and CX
#### when have more qubits than 2.
#### using import numpy as np library for the matrices
class gates_x:
# using u3 to define x gate where
# theta = pi
# lamda = 0
# phi = pi
one_size = gates_basic.u3(np.pi,0,np.pi) # assign a variable x gate matrix
cu_size = np.array([[1, 0, 0, 0], # generate cx matrix
[0, 1, 0, 0], # and assign a variable
[0, 0, 0, 1],
[0, 0, 1, 0]])
cu_inv_size = np.array([[1, 0, 0, 0], # matrices of 4x4 and higher
[0, 0, 0, 1], # need an inverse part that
[0, 0, 1, 0], # changes the order of their values
[0, 1, 0, 0]])
def _1(self): # method when is X for a qubit in the simulation
return gates_x.one_size # return the X gate matrix
def _n(self,total_qubits,target): # method when there is more than one qubit
#and the X gate must be applied
return find_gate.gate_1xn(total_qubits,target,gates_x.one_size)
# return the tensor product
def cu(self): # method when is CX when control value is more bigger than target target
return gates_x.cu_size # return cx matrix
def cu_inv(self): # method when is CH when control value is more less than target target
return gates_x.cu_inv_size # return the inverse of cx
def cu_1xn(self,total_qubits,target): # method when is cx when control value is more bigger
# than target by 1 and have 3 or more qubits
# in the quantum circuit
return find_gate.gate_1xn(total_qubits,target,gates_x.cu_size) #return the tensor product
def cu_1xn_inv(self,total_qubits,target): # method when is cx when control value is more bless
# than target by 1 and have 3 or more qubits
# in the quantum circuit
return find_gate.gate_1xn(total_qubits,target,gates_x.cu_inv_size) #return the tensor product
def cu_i_i2xn(self,target_0,target_1):# method when is cx when control value is more bless
# than target by more than 1 and have 3 or more qubits
# in the quantum circuit
return find_gate.gate_i_i2xn(target_0,target_1,gates_x.one_size)#return the tensor product
def cu_i_i2xn_inv(self,target_0,target_1):# method when is cx when control value is more bless
# than target by more than 1 and have 3 or more qubits
# in the quantum circuit
return find_gate.gate_i_i2xn_inv(target_0,target_1,gates_x.one_size)#return the tensor product
# +
########### gates_h.py ##############################
#### The operators H and CH are defined ######
#### and considering all the possibilities ######
#### of being carried out from one qubit or ######
#### two depending on the matrix up to N qubits ######
#### and may correspond to the expected result. ######
#### This class was added in the simulator to ######
#### validate that the structure of the ######
#### find_gate class can be used for any ######
#### matrix u and cu. ######
######################################################
#### define the methods for the posibilities for H and CH
#### when have more qubits than 2.
#### using import numpy as np library for the matrices
class gates_h:
# using u3 to define H gate where
# theta = pi/2
# lamda = 0
# phi = pi
one_size = gates_basic.u3(np.pi/2,0,np.pi) # assign a variable H gate matrix
cu_size = np.array([[1, 0, 0, 0],# generate CH matrix
[0, 1, 0, 0], #assign a variable
[0, 0, 1/np.sqrt(2), 1/np.sqrt(2)],
[0, 0, 1/np.sqrt(2), -1/np.sqrt(2)]])
cu_inv_size = np.array([[1/np.sqrt(2), 1/np.sqrt(2), 0, 0], # matrices of 4x4 and higher
[1/np.sqrt(2), -1/np.sqrt(2), 0, 0], # need an inverse part that
[0, 0, 1, 0],
[0, 0, 0, 1]])
def _1(self): # method when is H for a qubit in the simulation
return gates_h.one_size # return the H gate matrix
def _n(self,total_qubits,target): # method when there is more than one qubit
#and the H gate must be applied
return find_gate.gate_1xn(total_qubits,target,gates_h.one_size)
# return the tensor product
def cu(self): # method when is CH when control value is more bigger than target target
return gates_h.cu_size # return CH matrix
def cu_inv(self): # method when is CH when control value is more less than target target
return gates_h.cu_inv_size # return the inverse of CH
def cu_1xn(self,total_qubits,target): # method when is CH when control value is more bigger
# than target by 1 and have 3 or more qubits
# in the quantum circuit
return find_gate.gate_1xn(total_qubits,target,gates_h.cu_size)#return the tensor product
def cu_1xn_inv(self,total_qubits,target):# method when is CH when control value is more bless
# than target by 1 and have 3 or more qubits
# in the quantum circuit
return find_gate.gate_1xn(total_qubits,target,gates_h.cu_inv_size)#return the tensor product
def cu_i_i2xn(self,target_0,target_1):# method when is CH when control value is more bless
# than target by more than 1 and have 3 or more qubits
# in the quantum circuit
return find_gate.gate_i_i2xn(target_0,target_1,gates_h.one_size)#return the tensor product
def cu_i_i2xn_inv(self,target_0,target_1):# method when is CH when control value is more bless
# than target by more than 1 and have 3 or more qubits
# in the quantum circuit
return find_gate.gate_i_i2xn_inv(target_0,target_1,gates_h.one_size)#return the tensor product
# +
########### operator_size.py ########################
#### The static class operator_size has two ######
#### methods to identify the order of the ######
#### output unitary matrix depending on the ######
#### input qubits and targets. ######
######################################################
#### The methods aim to obtain the tensor product
#### for gates of size 2x2 and 4x4
#### when there are more than 1 or 2 qubits respectively.
class operator_size:
def tam_1(total_qubits,target_0,gate): ## method for the two cases of a gate of size 2x2
if total_qubits == 1: ## when has a qubit
O_state = gate._1() ## return the original gate
else: ## in other case call gate._n to find the product tensor
O_state = gate._n(total_qubits, target_0) ## assign in the output
return O_state ## and return the output matrix
def tam_2(total_qubits,target_qubits,gate): ## method for all the cases of a gate of size 4x4
size_cu = total_qubits-1 ## how is gate 4x4 is necessary subtract 1 from the number of qubits
if target_qubits[0] > target_qubits[1]: ## when control's qubit is more than target's qubit
if size_cu == 1: ## case when have 2 qubis
O_state = gate.cu() ## apply direct the original gate
elif target_qubits[0] - target_qubits[1] == 1: ## when the control's index is next to
## with the target's index
O_state = gate.cu_1xn(size_cu,target_qubits[1]) # apply the tensor product cu_1xn
else: ## others case we apply the method cu_i_i2xn
O_state = gate.cu_i_i2xn(target_qubits[1],target_qubits[0])
if O_state.shape[0] < 2**total_qubits: ## in case we need a bigger matrix
O_state = find_gate.gate_moving(total_qubits,target_qubits[0],## apply the method
target_qubits[1],O_state) ## gate_moving
elif target_qubits[0] < target_qubits[1]: ## when control's qubit is less than target's qubit
if size_cu == 1: ## case when have 2 qubis
O_state = gate.cu_inv() ## apply direct the inverse original gate
elif target_qubits[1] - target_qubits[0] == 1:## when the control's index is next to
## with the target's index
O_state = gate.cu_1xn_inv(size_cu,target_qubits[0]) # apply the tensor product cu_1xn
else: ## others case we apply the method cu_i_i2xn
O_state = gate.cu_i_i2xn_inv(target_qubits[0],target_qubits[1])
if O_state.shape[0] < 2**total_qubits: ## in case we need a bigger matrix
O_state = find_gate.gate_moving(total_qubits,target_qubits[1],## apply the method
target_qubits[0],O_state) ## gate_moving
return O_state ## return the final matrix with the size 2**nx2**n with n = total_qubits
# -
# ## Main functions for the simulation
# +
########### task functios ############################
#### The functions required to generate the ######
#### simulator, which consists of generating the ######
#### initial state zero for N qubits, applying ######
#### the gates to the input qubits, reading the ######
#### quantum circuit and measuring ######
#### the output qubits. ######
######################################################
#It is necessary to use the methods of the classes:
# -gates_basic
# -find_gate
#and objects of type:
# -gates_x
# -gates_h
#Furthermore, is neccesary a exra module random
#for apply the weighted random.
import random #
# return vector of size 2**num_qubits with all zeroes except first element which is 1
def get_ground_state(num_qubits):
vector = [0]* (2**num_qubits) # generar a vector of zeros with a sizeof 2^num_qubits
vector[0] = 1 # modify the first element to 1
return vector # return the vector result
# return unitary operator of size 2**n x 2**n for given gate and target qubits
def get_operator(total_qubits, gate_unitary, target_qubits, params):
O_state = gates_basic.i() #the O_state variable of the output gate is
#initialized with the 2x2 identity matrix
gate_list = ['h','x','u3','cx','ch'] # list of the unitary operators or quantum gatess
if gate_unitary == gate_list[0]: # case h
gate_i = gates_h() # an object of class gates_h is instantiated.
O_state = operator_size.tam_1(total_qubits,target_qubits[0],gate_i)
#assign the matrix of size 2**nx2**n with the H gate in the target qubit position
elif gate_unitary == gate_list[1]:# case x
gate_i = gates_x() # an object of class gates_x is instantiated.
O_state = operator_size.tam_1(total_qubits,target_qubits[0],gate_i)
#assign the matrix of size 2**nx2**n with the X gate in the target qubit position
elif gate_unitary == gate_list[2]: # case U3
## works similar with the method tam_! of class operator_size but don't exist
## the class U3 so work with the static method of fates_basic
if total_qubits == 1: ## when has a qubit, then return the original gate
O_state = gates_basic.u3(params['theta'],params['lambda'],params['phi'])
else: ## other case apply the tensor product with n-1 Identities gate
## where n = total_qubits
O_state = find_gate.gate_1xn(total_qubits,target_qubits[0], ## assing U3 matrix to
gates_basic.u3(params['theta'], ## variable O_state
params['lambda'],params['phi']))
elif gate_unitary == gate_list[3]: ## case cx
gate_i = gates_x() ## an object of class gates_x is instantiated.
O_state = operator_size.tam_2(total_qubits,target_qubits,gate_i)
#assign the matrix of size 2**nx2**n with the CX gate in the target qubit position
elif gate_unitary == gate_list[4]: ## case ch
gate_i = gates_h() ## an object of class gates_h is instantiated.
O_state = operator_size.tam_2(total_qubits,target_qubits,gate_i)
#assign the matrix of size 2**nx2**n with the CH gate in the target qubit position
return O_state ## return the final matrix
def run_program(initial_state, program,global_params=None):
# read program, and for each gate:
# - calculate matrix operator
# - multiply state with operator
# return final state
# code
total_qubits = int(np.log2(len(initial_state))) #obtain the numbers of qubits
if global_params: ## global parameters when work whit quantum variational circuits
if len(global_params) == 2:
global_params["lambda"] = -3.1415 ## deault value or the example
global_params["theta"] = global_params.pop("global_1") ## global_1 params to theta
global_params["phi"] = global_params.pop("global_2") ## global:! params to phi
for i in program: ## run the program
matrix_unitary = get_operator(total_qubits, i['gate'], i['target'],global_params)
initial_state = np.dot(matrix_unitary, initial_state) ## apply the dot product
else:
for i in program: ## run the program
if 'params' in i: ## if exist params in the input's data
## get the input with params values
matrix_unitary = get_operator(total_qubits, i['gate'], i['target'],i['params'])
else: ## in other case get the input's data without params values
matrix_unitary = get_operator(total_qubits, i['gate'], i['target'],None)
initial_state = np.dot(matrix_unitary, initial_state) ## apply the dot product
## between the operator and the urrent vector state
return initial_state # return the output_state after apply the unitary matrix with vector's input
## choose element from state_vector using weighted random and return it's index
def measure_all(state_vector):
state_vector_output = [] ## list of the index of each output state
state_vector_prob = [] ## list of the scalar number of each output state
lenght = len(state_vector) ## ientify the len of the state vector
for i in range(lenght): ## and apply a iterative function depend of the lenght
index = bin(i)[2:] ## pass the integer value index to binary number
while len(index) < np.log2(lenght): ## in case to don't have the same lenght we adder '0'
index = '0' + index ## in the left part
state_vector_output.append(str(index)) ## append the index binary value in a list
state_vector_prob.append((abs(state_vector[i])**2)) ## append the probability value in a list
return random.choices( ## apply choice which generate a weighted random
population=state_vector_output, ## list to the states from the output
weights=state_vector_prob, ## weights/probabilities of the every state
k=1) ## apply random 1 time to find the output
# simply execute measure_all in a loop num_shots times and
# return object with statistics in following form:
# {
# element_index: number_of_ocurrences,
# element_index: number_of_ocurrences,
# element_index: number_of_ocurrences,
# ...
# }
# (only for elements which occoured - returned from measure_all)
def get_counts(state_vector, num_shots):
measurment_dict = {} ## the dict output
for i in range(num_shots): ## apply a iterative function num_shots times
index_dict = measure_all(state_vector)[0] ## obtain the measure_all result
if not index_dict in measurment_dict: ## in case not exist the value in the dict
measurment_dict[index_dict] = 1 ## we assign with 1 value
else: ## in other case
measurment_dict[index_dict] += 1 ## increment the index value with 1
return measurment_dict ## finally return the dict output
## for all measurment
# -
# ### Example usage
#
# If your code is organized as we suggested, then usage will look like this:
# +
# Define program:
my_circuit = [
{ "gate": "h", "target": [0] },
{ "gate": "cx", "target": [0, 1] }
]
# Create "quantum computer" with 2 qubits (this is actually just a vector :) )
my_qpu = get_ground_state(2)
# Run circuit
final_state = run_program(my_qpu, my_circuit)
# Read results
counts = get_counts(final_state, 1000)
print(counts)
# Should print something like:
# {
# "00": 502,
# "11": 498
# }
# Voila!
# -
# ## Bonus requirements
#
# If you have implemented simulator as described above: congratulations!
#
# Now, if you wish you can continue improving it, first and useful thing to do would be to allow parametric gates:
#
#
# ### Parametric gates
#
# For example, following gate:
#
# ```
# [
# ["cos(theta/2)", "-exp(i * lambda) * sin(theta / 2)"],
# ["exp(i * phi) * sin(theta / 2)", "exp(i * lambda + i * phi) * cos(theta / 2)"]
# ]
# ```
#
# Contains strings with expressions, and expressions can contain variables (usually angles in radians).
#
# When your program gets gate like this, it should parse and evaluate expressions (with variables) and make a "normal" unitary matrix with values, which is then applied to the state vector.
#
# Example program with parametric gates:
#
# ```
# [
# { "unitary": [["cos(theta/2)", "-exp(i * lambda) * sin(theta / 2)"], ["exp(i * phi) * sin(theta / 2)", "exp(i * lambda + i * phi) * cos(theta / 2)"]], "params": { "theta": 3.1415, "phi": 1.15708, "lambda": -3.1415 }, "target": [0] }
# ...
# ]
# ```
#
# Or, if you have defined unitaries somewhere in the program, then:
#
# ```
# [
# { "gate": "u3", "params": { "theta": 3.1415, "phi": 1.5708, "lambda": -3.1415 }, "target": [0] }
# ...
# ]
# ```
#
# Which your program translates to:
#
# ```
# [
# [ 0+0j, 1+0j],
# [ 0+1j, 0+0j]
# ]
# ```
#
#
# ### Allow running variational quantum algorithms
#
# With support for parametric gates, all you need to do is to allow global params - and your simulator will be able to run variational quantum algorithms!
#
# In that case, parametrized gates in your program will contain strings instead parameter values:
#
# ```
# [
# { "gate": "u3", "params": { "theta": "global_1", "phi": "global_2", "lambda": -3.1415 }, "target": [0] }
# ...
# ]
# ```
#
# Notice `global_1` and `global_2` instead angle values, which you pass to `run_program` method:
#
# ```
# final_state = run_program(my_qpu, my_circuit, { "global_1": 3.1415, "global_2": 1.5708 })
# ```
#
# And that way you can use it in variational algorithms:
#
# ```
# mu_qpu = [...]
# my_circuit = [...]
#
# def objective_function(params):
# final_state = run_program(my_qpu, my_circuit, { "global_1": params[0], "global_2": params[1] })
#
# counts = get_counts(final_state, 1000)
#
# # ...calculate cost here...
#
# return cost
#
# # initial values
# params = np.array([3.1415, 1.5708])
#
# # minimize
# minimum = minimize(objective_function, params, method="Powell", tol=1e-6)
# ```
#
# ### Parametric gates
# example for the parametric gate
# +
# Define program:
my_circuit = [
{ "gate": "u3", "params": { "theta": 3.1415, "phi": 1.5708, "lambda": -3.1415 }, "target": [0] }
]
# using y3 gate with the index params wiht theta, phi, lambda values
# Create "quantum computer" with 1 qubits (this is actually just a vector :) )
my_qpu = get_ground_state(1)
print('Show the U3 gate is equal to X gate') ## indicate that is an example of U3
print()
# Run circuit
final_state = run_program(my_qpu, my_circuit)
# Read the final_state
print(final_state)
# Read results
counts = get_counts(final_state, 1000)
print(counts)
# Expect output
#[
# [ 0+0j, 1+0j],
# [ 0+1j, 0+0j]
#]
# -
# ### Allow running variational quantum algorithms
#
# +
from scipy.optimize import minimize ## need the mthod minimize of scipy.optimize module
my_qpu = get_ground_state(1)
my_circuit = [
{ "gate": "u3", "target": [0] }
]
real_value = np.random.randn(2,) ## real_value
print("real number: ", real_value)
def calc_cost(real_value, circuit_value):
diff = real_value - circuit_value ## calculates the sum of squares
mod_abs = diff.real**2 + diff.imag**2 ## of mod of difference between corresponding elements
cost = np.sum(mod_abs) ## of real_value and circuit_value
return cost ## returns the cost
def objective_function(params):
final_state = run_program(my_qpu, my_circuit,
{ "global_1": params[0], "global_2": params[1] })
#counts = get_counts(final_state, 1000) ## return the values of the measure
# ...calculate cost here...
return calc_cost(real_value, final_state)
# initial values
params = np.array([3.1415, 1.5708])
# minimize
minimum = minimize(objective_function, params, method="Powell", tol=1e-6)
print(minimum)
# -
# ## Extra examples
#
# Show different inputs and prove the result of the simulator
# +
# Define program:
my_circuit = [
{ "gate": "h", "target": [0] },
{ "gate": "cx", "target": [0, 5] }
]
# Create "quantum computer" with 2 qubits (this is actually just a vector :) )
my_qpu = get_ground_state(6)
# Run circuit
final_state = run_program(my_qpu, my_circuit)
# Read results
counts = get_counts(final_state, 1000)
print(counts)
# Should print something like:
# {
# "00": 502,
# "11": 498
# }
# Voila!
# +
# Define program:
my_circuit = [
{ "gate": "x", "target": [5] },
{ "gate": "cx", "target": [5, 0] }
]
# Create "quantum computer" with 2 qubits (this is actually just a vector :) )
my_qpu = get_ground_state(6)
# Run circuit
final_state = run_program(my_qpu, my_circuit)
# Read results
counts = get_counts(final_state, 1000)
print(counts)
# Should print something like:
# {
# "00": 502,
# "11": 498
# }
# Voila!
# +
# Define program:
my_circuit = [
{ "gate": "h", "target": [0] },
{ "gate": "ch", "target": [0, 4] }
]
# Create "quantum computer" with 2 qubits (this is actually just a vector :) )
my_qpu = get_ground_state(6)
# Run circuit
final_state = run_program(my_qpu, my_circuit)
# Read results
counts = get_counts(final_state, 1000)
print(counts)
# Should print something like:
# {
# "00": 502,
# "11": 498
# }
# Voila!
# +
# Define program:
my_circuit = [
{ "gate": "x", "target": [2] },
{ "gate": "ch", "target": [2, 4] }
]
# Create "quantum computer" with 2 qubits (this is actually just a vector :) )
my_qpu = get_ground_state(6)
# Run circuit
final_state = run_program(my_qpu, my_circuit)
# Read results
counts = get_counts(final_state, 1000)
print(counts)
# Should print something like:
# {
# "00": 502,
# "11": 498
# }
# Voila!
# +
# Define program:
my_circuit = [
{ "gate": "h", "target": [2] },
{ "gate": "cx", "target": [2, 3] } ,
{ "gate": "u3", "params": { "theta": 2.1415, "phi": 1.5708, "lambda": -3.1415 }, "target": [0] }
]
# Create "quantum computer" with 2 qubits (this is actually just a vector :) )
my_qpu = get_ground_state(6)
# Run circuit
final_state = run_program(my_qpu, my_circuit)
# Read results
counts = get_counts(final_state, 1000)
print(counts)
# Should print something like:
# {
# "00": 502,
# "11": 498
# }
# Voila!
# -
# ## Additional info/help
#
# Any questions? Ping us on Slack!
#
# To be blamed: <NAME>,
# Quantastica
#
# May the force be with you!
# ## Next steps:
#
# Implement ccx, swap, fredkin gate, works with swap test i nthe quantum variational circuits.
# Also, I can do a better implementation of the classes and methods
# ## Final comments:
#
# Very good task and with good activities, I hope to continue improving this project and that it can be a great simulator regardless of the result, I had a lot of fun making it and I think it can be improved, thank you very much for your resources.
| challenge-2021.02-feb/maldoalberto/qosf-simulator-task.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %reset -f
# %matplotlib inline
import matplotlib as mpl
import numpy as np
from numpy import array as a
import matplotlib.pyplot as plt
import numpy.random as rng
from scipy.special import expit as sigmoid
np.set_printoptions(precision = 2, suppress = True)
import time
rng.seed(int(time.time())) # seed the random number generator
# specify a weights matrix
N = 2
hiWgt, loWgt = 8.0, -6.0
W = loWgt * np.ones((N,N), dtype=float)
for i in range(N): W[i,i] = hiWgt
print(W)
# make up an array with each row being one of the binary patterns. Do 'em all.
hidpats = np.array([[0 if (i & (1 << bit) == 0) else 1 for bit in range(N)] for i in range(2**N)])
vispats = np.array([[0 if (i & (1 << bit) == 0) else 1 for bit in range(N)] for i in range(2**N)])
# calculate the true probability distribution over hidden pats for each RBM, under the generative model.
pHid = {}
total = 0.0
for pat in hidpats:
phiVis = np.dot(W.T, pat)
logP_star = np.sum(np.log(1+np.exp(phiVis)))
pHid[tuple(pat)] = np.exp(logP_star)
total += pHid[tuple(pat)]
for pat in pHid.keys():
pHid[pat] = pHid[pat] / total
for pat in hidpats:
print (pat, pHid[tuple(pat)])
# form the joint distribution over hiddens AND visibles
pHV = {}
for vis in vispats:
for hA in hidpats:
for hB in hidpats:
phi = np.dot(W.T, hA) + np.dot(W.T, hB)
pVis = np.prod(vis * sigmoid(phi) + (1-vis) * (1 - sigmoid(phi)))
pHV[(tuple(hA),tuple(hB),tuple(vis))] = pHid[tuple(hA)] * pHid[tuple(hB)] * pVis
print('visible probabilities under generative model:')
for vis in vispats:
total = 0.0
for hA in hidpats:
for hB in hidpats:
total += pHV[(tuple(hA),tuple(hB),tuple(vis))]
print(vis, ' prob: ',total)
print('hidden probabilities, given each visible in turn:')
for vis in vispats:
print('vis: ',vis)
normalisation = 0.0
for hA in hidpats:
for hB in hidpats:
normalisation += pHV[(tuple(hA),tuple(hB),tuple(vis))]
for hA in hidpats:
for hB in hidpats:
if pHV[(tuple(hA),tuple(hB),tuple(vis))]/normalisation > 0.01:
print ('\t hA,hB: ', hA, hB, ' ',pHV[(tuple(hA),tuple(hB),tuple(vis))]/normalisation)
| ExploratoryCode/compute_exact_posterior.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import altair as alt
# We are using default data transformer for html chart to work offline
# alt.data_transformers.enable('default')
alt.data_transformers.enable('json')
# alt.data_transformers.enable('csv')
# alt.data_transformers.enable('default', max_rows=1000000)
# alt.renderers.enable('notebook')
# alt.data_transformers.enable('data_server')
alt.renderers.enable('notebook', embed_options={'renderer': 'svg'})
from vega_datasets import data
import pandas as pd
from altair import Scale,Color
file = 'annual_county_level.csv'
# Reading the original dataframe
dforg = pd.read_csv(file,header=0)
# Looking at the data in the dataframe
dforg.head()
# Looking at the statistics of the data
dforg.describe()
# Looking at the data-types of the columns
dforg.dtypes
# Changing the data-types of non-numeric to str
dforg['Pill_per_pop'] = dforg["Pill_per_pop"].astype(int)
dforg['BUYER_COUNTY'] = dforg["BUYER_COUNTY"].astype(str)
dforg['state'] = dforg["state"].astype(str)
# Year will be used as column names, hence, we have to convert to str
dforg['year'] = dforg["year"].astype(str)
# Subset the data selecting the required columns
fdf = dforg[['fips','year','Pill_per_pop','BUYER_COUNTY','state']]
# Rows having pills per person >150 are =150 to cap off.
fdf.loc[fdf['Pill_per_pop']>150,['Pill_per_pop']] = 150
# +
# We now take another subset from the above subsetted data
fdfn = fdf[['fips','year','Pill_per_pop']]
fdfn = fdfn.pivot(index='fips',columns='year',values='Pill_per_pop').reset_index('fips')
# -
# Storing a list of column names for plotting
columns = [str(i) for i in range(2006,2013)]+['BUYER_COUNTY','state']
# Joining dataframes to add BUYER_COUNTY and state columns
fdfn = fdfn.set_index('fips').join(fdf[['fips','BUYER_COUNTY','state']].set_index('fips'))
# Resetting index to index(fips) column
fdfn = fdfn.reset_index()
# Setting plotting properties
dummy = 'nothing' #This holds the default fill-in color
scheme = 'yellowgreenblue'
scale_type = 'linear'
# Using based on the color scheme chosen, we check the values in
# MatplotLib cmaps and take the appropriate hex-code value for dummy
# Since yellowgreenblue is not present in matplotlib, we go to:
# https://htmlcolorcodes.com/color-picker/
# and select the dummy fill-in color of choice.
# Otherwise, if color-scheme is in matplotlib, we use the following code:
# cmap = cm.get_cmap('viridis_r', 15)
# cutoff = 0
# for i in range(cmap.N):
# rgb = cmap(i)[:3] # will return rgba, we take only first 3 so we get rgb
# print(matplotlib.colors.rgb2hex(rgb))
# if i == cutoff:
# dummy = matplotlib.colors.rgb2hex(rgb)
dummy = '#EEFBB3'
# Getting the json to plot states and counties
# Make sure there is a defference in the urls
states = alt.topo_feature(data.us_10m.url, 'states')
counties = alt.topo_feature(data.us_10m.url+'#', 'counties')
cols = [str(i) for i in range(2006,2013)]
cols
columns
# +
us_counties = alt.topo_feature(data.us_10m.url, 'counties')
slider = alt.binding_range(min=2006, max=2012, step=1)
select_year = alt.selection_single(name="year", fields=['year'],
bind=slider, init={'year': 2006})
a = alt.Chart(counties).mark_geoshape(
stroke='black',
strokeWidth=0.05
).project(
type='albersUsa'
).transform_lookup(
lookup='id',
from_=alt.LookupData(fdfn, 'fips', columns)
).transform_fold(
cols, as_=['year', 'Pill_per_pop']
).transform_calculate(
year='parseInt(datum.year)',
Pill_per_pop='isValid(datum.Pill_per_pop) ? datum.Pill_per_pop : -1'
).encode(
color = alt.condition(
'datum.Pill_per_pop > 0',
alt.Color('Pill_per_pop:Q', scale=alt.Scale(type='linear',scheme='yellowgreenblue')),
alt.value(dummy),
tooltip=['BUYER_COUNTY:N', 'state:N','Pill_per_pop:Q','year:Q']
)).add_selection(
select_year
).properties(
width=700,
height=400
).transform_filter(
select_year
)
b = alt.Chart(states).mark_geoshape(stroke='black',strokeWidth=0.15).project(
type='albersUsa'
)
c = alt.Chart(counties).mark_geoshape(fillOpacity=0
).project(
type='albersUsa'
).transform_lookup(
lookup='id',
from_=alt.LookupData(fdfn, 'fips', columns)
).transform_fold(
cols, as_=['year', 'Pill_per_pop']
).transform_calculate(
year='parseInt(datum.year)',
Pill_per_pop='isValid(datum.Pill_per_pop) ? datum.Pill_per_pop : -1'
).encode(
tooltip=['BUYER_COUNTY:N', 'state:N','Pill_per_pop:Q','year:Q']
).properties(
title='Pills per person',
width=700,
height=400
)
chart = a+b+c
# -
chart.save('chart.html', embed_options={'renderer':'svg'})
chart.display(renderer='svg')
# +
# # Plotting function
# def choropleth(data,level,color=True,scheme='yellowgreenblue',dummy=dummy,stroke='black',strokeWidth=0.05,type='linear',fips='fips',columns=columns):
# foldcols = columns.copy()
# foldcols.remove('BUYER_COUNTY')
# foldcols.remove('state')
# slider = alt.binding_range(min=2006, max=2012, step=1)
# select_year = alt.selection_single(name="key", fields=['key'],
# bind=slider, init={'key': 2006})
# if color:
# chart = alt.Chart(level).mark_geoshape(
# stroke=stroke,
# strokeWidth=strokeWidth
# ).project(
# type='albersUsa'
# ).transform_lookup(
# lookup='id',
# from_=alt.LookupData(data, fips, columns)
# ).transform_fold(
# foldcols,as_=['year','Pill_per_pop']
# ).transform_calculate(
# year='parseInt(datum.year)',
# Pill_per_pop='isValid(datum.Pill_per_pop) ? datum.Pill_per_pop : -1'
# ).encode(
# color = alt.condition(
# 'datum.Pill_per_pop > 0',
# alt.Color('Pill_per_pop:Q', scale=Scale(scheme=scheme,type=type)),
# alt.value(dummy)
# ),
# tooltip=['BUYER_COUNTY:N', 'state:N','Pill_per_pop:Q','year:Q']
# ).add_selection(
# select_year
# ).properties(
# width=700,
# height=400,
# title='Pills per person'
# ).transform_filter(
# select_year
# )
# else:
# chart = alt.Chart(level).mark_geoshape(
# stroke=stroke,
# strokeWidth=strokeWidth
# ).project(
# type='albersUsa'
# ).transform_lookup(
# lookup='id',
# from_=alt.LookupData(data, fips, columns)
# ).transform_fold(
# foldcols,as_=['year','Pill_per_pop']
# ).transform_calculate(
# year='parseInt(datum.year)',
# Pill_per_pop='isValid(datum.Pill_per_pop) ? datum.Pill_per_pop : -1'
# ).encode(
# tooltip=['BUYER_COUNTY:N', 'state:N','Pill_per_pop:Q','year:Q']
# ).add_selection(
# select_year
# ).properties(
# width=700,
# height=400,
# title='Pills per person'
# ).transform_filter(
# select_year
# )
# return chart
# +
# def plot_year(year,fdf=fdf,counties=counties,states=states,state_stroke='black',state_strokew=0.15):
# chart = choropleth(fdf1,counties)
# outline = alt.Chart(states).mark_geoshape(stroke=state_stroke,strokeWidth=state_strokew).project(
# type='albersUsa'
# )
# chart1 = choropleth(fdf1,counties,color=False,year=year)
# return chart+outline+chart1
# +
# chart = choropleth(fdfn,counties)
# +
# Useful links:
# https://github.com/altair-viz/altair/issues/611
# ellisonbg
| .ipynb_checkpoints/main-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # AGENT #
#
# An agent, as defined in 2.1 is anything that can perceive its <b>environment</b> through sensors, and act upon that environment through actuators based on its <b>agent program</b>. This can be a dog, robot, or even you. As long as you can perceive the environment and act on it, you are an agent. This notebook will explain how to implement a simple agent, create an environment, and create a program that helps the agent act on the environment based on its percepts.
#
# Before moving on, review the </b>Agent</b> and </b>Environment</b> classes in <b>[agents.py](https://github.com/aimacode/aima-python/blob/master/agents.py)</b>.
#
# Let's begin by importing all the functions from the agents.py module and creating our first agent - a blind dog.
# +
from agents import *
class BlindDog(Agent):
def eat(self, thing):
print("Dog: Ate food at {}.".format(self.location))
def drink(self, thing):
print("Dog: Drank water at {}.".format( self.location))
dog = BlindDog()
# -
# What we have just done is create a dog who can only feel what's in his location (since he's blind), and can eat or drink. Let's see if he's alive...
print(dog.alive)
# 
# This is our dog. How cool is he? Well, he's hungry and needs to go search for food. For him to do this, we need to give him a program. But before that, let's create a park for our dog to play in.
# # ENVIRONMENT #
#
# A park is an example of an environment because our dog can perceive and act upon it. The <b>Environment</b> class in agents.py is an abstract class, so we will have to create our own subclass from it before we can use it. The abstract class must contain the following methods:
#
# <li><b>percept(self, agent)</b> - returns what the agent perceives</li>
# <li><b>execute_action(self, agent, action)</b> - changes the state of the environment based on what the agent does.</li>
# +
class Food(Thing):
pass
class Water(Thing):
pass
class Park(Environment):
def percept(self, agent):
'''prints & return a list of things that are in our agent's location'''
things = self.list_things_at(agent.location)
print(things)
return things
def execute_action(self, agent, action):
'''changes the state of the environment based on what the agent does.'''
if action == "move down":
agent.movedown()
elif action == "eat":
items = self.list_things_at(agent.location, tclass=Food)
if len(items) != 0:
if agent.eat(items[0]): #Have the dog pick eat the first item
self.delete_thing(items[0]) #Delete it from the Park after.
elif action == "drink":
items = self.list_things_at(agent.location, tclass=Water)
if len(items) != 0:
if agent.drink(items[0]): #Have the dog drink the first item
self.delete_thing(items[0]) #Delete it from the Park after.
def is_done(self):
'''By default, we're done when we can't find a live agent,
but to prevent killing our cute dog, we will or it with when there is no more food or water'''
no_edibles = not any(isinstance(thing, Food) or isinstance(thing, Water) for thing in self.things)
dead_agents = not any(agent.is_alive() for agent in self.agents)
return dead_agents or no_edibles
# -
# ## Wumpus Environment
# +
from ipythonblocks import BlockGrid
from agents import *
color = {"Breeze": (225, 225, 225),
"Pit": (0,0,0),
"Gold": (253, 208, 23),
"Glitter": (253, 208, 23),
"Wumpus": (43, 27, 23),
"Stench": (128, 128, 128),
"Explorer": (0, 0, 255),
"Wall": (44, 53, 57)
}
def program(percepts):
'''Returns an action based on it's percepts'''
print(percepts)
return input()
w = WumpusEnvironment(program, 7, 7)
grid = BlockGrid(w.width, w.height, fill=(123, 234, 123))
def draw_grid(world):
global grid
grid[:] = (123, 234, 123)
for x in range(0, len(world)):
for y in range(0, len(world[x])):
if len(world[x][y]):
grid[y, x] = color[world[x][y][-1].__class__.__name__]
def step():
global grid, w
draw_grid(w.get_world())
grid.show()
w.step()
# -
step()
# # PROGRAM #
# Now that we have a <b>Park</b> Class, we need to implement a <b>program</b> module for our dog. A program controls how the dog acts upon it's environment. Our program will be very simple, and is shown in the table below.
# <table>
# <tr>
# <td><b>Percept:</b> </td>
# <td>Feel Food </td>
# <td>Feel Water</td>
# <td>Feel Nothing</td>
# </tr>
# <tr>
# <td><b>Action:</b> </td>
# <td>eat</td>
# <td>drink</td>
# <td>move up</td>
# </tr>
#
# </table>
#
# +
class BlindDog(Agent):
location = 1
def movedown(self):
self.location += 1
def eat(self, thing):
'''returns True upon success or False otherwise'''
if isinstance(thing, Food):
print("Dog: Ate food at {}.".format(self.location))
return True
return False
def drink(self, thing):
''' returns True upon success or False otherwise'''
if isinstance(thing, Water):
print("Dog: Drank water at {}.".format(self.location))
return True
return False
def program(percepts):
'''Returns an action based on it's percepts'''
for p in percepts:
if isinstance(p, Food):
return 'eat'
elif isinstance(p, Water):
return 'drink'
return 'move down'
# +
park = Park()
dog = BlindDog(program)
dogfood = Food()
water = Water()
park.add_thing(dog, 0)
park.add_thing(dogfood, 5)
park.add_thing(water, 7)
park.run(10)
# -
# That's how easy it is to implement an agent, its program, and environment. But that was a very simple case. What if our environment was 2-Dimentional instead of 1? And what if we had multiple agents?
#
# To make our Park 2D, we will need to make it a subclass of <b>XYEnvironment</b> instead of Environment. Also, let's add a person to play fetch with the dog.
class Park(XYEnvironment):
def percept(self, agent):
'''prints & return a list of things that are in our agent's location'''
things = self.list_things_at(agent.location)
print(things)
return things
def execute_action(self, agent, action):
'''changes the state of the environment based on what the agent does.'''
if action == "move down":
agent.movedown()
elif action == "eat":
items = self.list_things_at(agent.location, tclass=Food)
if len(items) != 0:
if agent.eat(items[0]): #Have the dog pick eat the first item
self.delete_thing(items[0]) #Delete it from the Park after.
elif action == "drink":
items = self.list_things_at(agent.location, tclass=Water)
if len(items) != 0:
if agent.drink(items[0]): #Have the dog drink the first item
self.delete_thing(items[0]) #Delete it from the Park after.
def is_done(self):
'''By default, we're done when we can't find a live agent,
but to prevent killing our cute dog, we will or it with when there is no more food or water'''
no_edibles = not any(isinstance(thing, Food) or isinstance(thing, Water) for thing in self.things)
dead_agents = not any(agent.is_alive() for agent in self.agents)
return dead_agents or no_edibles
| agents.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 64-bit (conda)
# language: python
# name: python3
# ---
import pandas as pd
data = [
[1, 2, 3],
[4, 5, 6],
[7, 8, 9]
]
data
list('321')
df = pd.DataFrame(data, list('321'), list('ZYX'))
df
df.sort_index(inplace = True) # ordena pelo indice
df
df.sort_index(inplace = True, axis = 1) # ordena pela coluna
df
df.sort_values(by = 'X', inplace = True) # ordena pela coluna X
df
df.sort_values(by = '3', axis = 1, inplace = True) # ordena
df
df.sort_values(by = ['X', 'Y'], inplace = True)
df
# ## Exercicio - funcionamento do sort_index
data = [[9, 6, 3],
[8, 5, 2],
[7, 4, 1]
]
df = pd.DataFrame(data, list('ZYX'), list('CBA'))
df
df.sort_index()
df.sort_index(axis = 1)
df
| data/extras/dados/organizando-dataframes-sort.ipynb |