code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Aggregating data
#
# Pandas has very convenient features to aggregate data. That is, to compute summary statistics about datasets either as a whole or after dividing them into subsets based on data values.
#
# 1. Loading a comma-separated-value (CSV) dataset
# 2. Computing descriptive statistics
# 3. Grouping data by value
# 4. Creating pivot tables
import pandas as pd
# +
#pandas is very good at reading data from many different types of file: JSON, text, excel, HDF
#In this case, we will load a data frame from the comma-separated-file tips.csv
open('tips.csv','r').readlines()[:10]
# +
#Pandas can read such a file with a read_csv function.
tips = pd.read_csv('tips.csv')
# -
tips.head()
# +
tips.mean()
#apply the aggregation function: mean, which will
#compute average values for all the columns for which it's meaningful to do so
# +
#describe the dataset for more information
tips.describe()
# -
# ## Grouping
# +
#Let's say you want to know how well men tip versus women.
#For that, we can tell pandas to group the data frame based
#on the value of the column: sex using the function groupby.
#And then, we can take the mean.
tips.groupby('sex').mean()
# +
tips.groupby(['sex','smoker']).mean()
#This creates a pandas multidimensional index
# -
# ## Pivot Tables
#
# we create groups and assign them to both index values and columns so that represent a multidimensional analysis of the data in tabular format. For instance, we'll create a pivot table for our tips data frame showing the total bill amount, grouped by sex and smoker status.
pd.pivot_table(tips,'total_bill','sex','smoker')
# +
#one more dimensional grouping
pd.pivot_table(tips,'total_bill',['sex','smoker'],['day','time'])
# -
| 11 aggregation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1A.algo - Optimisation sous contrainte (correction)
#
# Un peu plus de détails dans cet article : [Damped Arrow-Hurwicz algorithm for sphere packing](https://arxiv.org/abs/1605.05473).
from jyquickhelper import add_notebook_menu
add_notebook_menu()
# On rappelle le problème d'optimisation à résoudre :
#
# $\left \{ \begin{array}{l} \min_U J(U) = u_1^2 + u_2^2 - u_1 u_2 + u_2 \\ sous \; contrainte \; \theta(U) = u_1 + 2u_2 - 1 = 0 \; et \; u_1 \geqslant 0.5 \end{array}\right .$
#
# Les implémentations de l'algorithme Arrow-Hurwicz proposées ici ne sont pas génériques. Il n'est pas suggéré de les réutiliser à moins d'utiliser pleinement le calcul matriciel de [numpy](http://www.numpy.org/).
# ## Exercice 1 : optimisation avec cvxopt
#
# Le module [cvxopt](http://cvxopt.org/) utilise une fonction qui retourne la valeur de la fonction à optimiser, sa dérivée, sa dérivée seconde.
#
# $\begin{array}{rcl} f(x,y) &=& x^2 + y^2 - xy + y \\ \frac{\partial f(x,y)}{\partial x} &=& 2x - y \\ \frac{\partial f(x,y)}{\partial y} &=& 2y - x + 1 \\ \frac{\partial^2 f(x,y)}{\partial x^2} &=& 2 \\ \frac{\partial^2 f(x,y)}{\partial y^2} &=& 2 \\ \frac{\partial^2 f(x,y)}{\partial x\partial y} &=& -1 \end{array}$
# Le paramètre le plus complexe est la fonction ``F`` pour lequel il faut lire la documentation de la fonction [solvers.cp](http://cvxopt.org/userguide/solvers.html#problems-with-nonlinear-objectives) qui détaille les trois cas d'utilisation de la fonction ``F`` :
#
# * ``F()`` ou ``F(None,None)``, ce premier cas est sans doute le plus déroutant puisqu'il faut retourner le nombre de contraintes non linéaires et le premier $x_0$
# * ``F(x)`` ou ``F(x,None)``
# * ``F(x,z)``
#
# L'algorithme de résolution est itératif : on part d'une point $x_0$ qu'on déplace dans les directions opposés aux gradients de la fonction à minimiser et des contraintes jusqu'à ce que le point $x_t$ n'évolue plus. C'est pourquoi le premier d'utilisation de la focntion $F$ est en fait une initialisation. L'algorithme d'optimisation a besoin d'un premier point $x_0$ dans le domaine de défintion de la fonction $f$.
# +
from cvxopt import solvers, matrix
import random
def fonction(x=None,z=None) :
if x is None :
x0 = matrix ( [[ random.random(), random.random() ]])
return 0,x0
f = x[0]**2 + x[1]**2 - x[0]*x[1] + x[1]
d = matrix ( [ x[0]*2 - x[1], x[1]*2 - x[0] + 1 ] ).T
if z is None:
return f, d
else :
h = z[0] * matrix ( [ [ 2.0, -1.0], [-1.0, 2.0] ])
return f, d, h
A = matrix([ [ 1.0, 2.0 ] ]).trans()
b = matrix ( [[ 1.0] ] )
sol = solvers.cp ( fonction, A = A, b = b)
print (sol)
print ("solution:",sol['x'].T)
# -
# ## Exercice 2 : l'algorithme de Arrow-Hurwicz
# +
def fonction(X) :
x,y = X
f = x**2 + y**2 - x*y + y
d = [ x*2 - y, y*2 - x + 1 ]
return f, d
def contrainte(X) :
x,y = X
f = x+2*y-1
d = [ 1,2]
return f, d
X0 = [ random.random(),random.random() ]
p0 = random.random()
epsilon = 0.1
rho = 0.1
diff = 1
iter = 0
while diff > 1e-10 :
f,d = fonction( X0 )
th,dt = contrainte( X0 )
Xt = [ X0[i] - epsilon*(d[i] + dt[i] * p0) for i in range(len(X0)) ]
th,dt = contrainte( Xt )
pt = p0 + rho * th
iter += 1
diff = sum ( [ abs(Xt[i] - X0[i]) for i in range(len(X0)) ] )
X0 = Xt
p0 = pt
if iter % 100 == 0 :
print ("i {0} diff {1:0.000}".format(iter,diff),":", f,X0,p0,th)
print (diff,iter,p0)
print("solution:",X0)
# -
# La code proposé ici a été repris et modifié de façon à l'inclure dans une fonction qui s'adapte à n'importe quel type de fonction et contrainte dérivables : [Arrow_Hurwicz](http://www.xavierdupre.fr/app/ensae_teaching_cs/helpsphinx/td_1a/optimisation_contrainte.html?highlight=arrow#ensae_teaching_cs.td_1a.optimisation_contrainte.Arrow_Hurwicz). Il faut distinguer l'algorithme en lui-même et la preuve de sa convergence. Cet algorithme fonctionne sur une grande classe de fonctions mais sa convergence n'est assurée que lorsque les fonctions sont quadratiques.
# ## Exercice 3 : le lagrangien augmenté
# +
def fonction(X,c) :
x,y = X
f = x**2 + y**2 - x*y + y
d = [ x*2 - y, y*2 - x + 1 ]
v = x+2*y-1
v = c/2 * v**2
# la fonction retourne maintenant dv (ce qu'elle ne faisait pas avant)
dv = [ 2*(x+2*y-1), 4*(x+2*y-1) ]
dv = [ c/2 * dv[0], c/2 * dv[1] ]
return f + v, d, dv
def contrainte(X) :
x,y = X
f = x+2*y-1
d = [ 1,2]
return f, d
X0 = [ random.random(),random.random() ]
p0 = random.random()
epsilon = 0.1
rho = 0.1
c = 1
diff = 1
iter = 0
while diff > 1e-10 :
f,d,dv = fonction( X0,c )
th,dt = contrainte( X0 )
# le dv[i] est nouveau
Xt = [ X0[i] - epsilon*(d[i] + dt[i] * p0 + dv[i]) for i in range(len(X0)) ]
th,dt = contrainte( Xt )
pt = p0 + rho * th
iter += 1
diff = sum ( [ abs(Xt[i] - X0[i]) for i in range(len(X0)) ] )
X0 = Xt
p0 = pt
if iter % 100 == 0 :
print ("i {0} diff {1:0.000}".format(iter,diff),":", f,X0,p0,th)
print (diff,iter,p0)
print("solution:",X0)
# -
# ## Prolongement 1 : inégalité
#
# Le problème à résoudre est le suivant :
#
# $\left\{ \begin{array}{l} \min_U J(U) = u_1^2 + u_1^2 - u_1 u_2 + u_2 \\ \; sous \; contrainte \; \theta(U) = u_1 + 2u_2 - 1 = 0 \; et \; u_1 \geqslant 0.3 \end{array}\right.$
# +
from cvxopt import solvers, matrix
import random
def fonction(x=None,z=None) :
if x is None :
x0 = matrix ( [[ random.random(), random.random() ]])
return 0,x0
f = x[0]**2 + x[1]**2 - x[0]*x[1] + x[1]
d = matrix ( [ x[0]*2 - x[1], x[1]*2 - x[0] + 1 ] ).T
h = matrix ( [ [ 2.0, -1.0], [-1.0, 2.0] ])
if z is None: return f, d
else : return f, d, h
A = matrix([ [ 1.0, 2.0 ] ]).trans()
b = matrix ( [[ 1.0] ] )
G = matrix ( [[0.0, -1.0] ]).trans()
h = matrix ( [[ -0.3] ] )
sol = solvers.cp ( fonction, A = A, b = b, G=G, h=h)
print (sol)
print ("solution:",sol['x'].T)
# -
# ## Version avec l'algorithme de Arrow-Hurwicz
# +
import numpy,random
X0 = numpy.matrix ( [[ random.random(), random.random() ]]).transpose()
P0 = numpy.matrix ( [[ random.random(), random.random() ]]).transpose()
A = numpy.matrix([ [ 1.0, 2.0 ], [ 0.0, -1.0] ])
tA = A.transpose()
b = numpy.matrix ( [[ 1.0], [-0.30] ] )
epsilon = 0.1
rho = 0.1
c = 1
first = True
iter = 0
while first or abs(J - oldJ) > 1e-8 :
if first :
J = X0[0,0]**2 + X0[1,0]**2 - X0[0,0]*X0[1,0] + X0[1,0]
oldJ = J+1
first = False
else :
oldJ = J
J = X0[0,0]**2 + X0[1,0]**2 - X0[0,0]*X0[1,0] + X0[1,0]
dj = numpy.matrix ( [ X0[0,0]*2 - X0[1,0], X0[1,0]*2 - X0[0,0] + 1 ] ).transpose()
Xt = X0 - ( dj + tA * P0 ) * epsilon
Pt = P0 + ( A * Xt - b) * rho
if Pt [1,0] < 0 : Pt[1,0] = 0
X0,P0 = Xt,Pt
iter += 1
if iter % 100 == 0 :
print ("iteration",iter, J)
print (iter)
print ("solution:",Xt.T)
# -
# ## Prolongement 2 : optimisation d'une fonction linéaire
#
# Correction à venir.
| _doc/notebooks/td1a_algo/td1a_correction_session9.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/mrdbourke/tensorflow-deep-learning/blob/main/07_food_vision_milestone_project_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="z2PPYrQIztfX"
# # 07. Milestone Project 1: 🍔👁 Food Vision Big™
#
# In the previous notebook ([transfer learning part 3: scaling up](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/06_transfer_learning_in_tensorflow_part_3_scaling_up.ipynb)) we built Food Vision mini: a transfer learning model which beat the original results of the [Food101 paper](https://data.vision.ee.ethz.ch/cvl/datasets_extra/food-101/) with only 10% of the data.
#
# But you might be wondering, what would happen if we used all the data?
#
# Well, that's what we're going to find out in this notebook!
#
# We're going to be building Food Vision Big™, using all of the data from the Food101 dataset.
#
# Yep. All 75,750 training images and 25,250 testing images.
#
# And guess what...
#
# This time **we've got the goal of beating [DeepFood](https://www.researchgate.net/publication/304163308_DeepFood_Deep_Learning-Based_Food_Image_Recognition_for_Computer-Aided_Dietary_Assessment)**, a 2016 paper which used a Convolutional Neural Network trained for 2-3 days to achieve 77.4% top-1 accuracy.
#
# > 🔑 **Note:** **Top-1 accuracy** means "accuracy for the top softmax activation value output by the model" (because softmax ouputs a value for every class, but top-1 means only the highest one is evaluated). **Top-5 accuracy** means "accuracy for the top 5 softmax activation values output by the model", in other words, did the true label appear in the top 5 activation values? Top-5 accuracy scores are usually noticeably higher than top-1.
#
# | | 🍔👁 Food Vision Big™ | 🍔👁 Food Vision mini |
# |-----|-----|-----|
# | Dataset source | TensorFlow Datasets | Preprocessed download from Kaggle |
# | Train data | 75,750 images | 7,575 images |
# | Test data | 25,250 images | 25,250 images |
# | Mixed precision | Yes | No |
# | Data loading | Performanant tf.data API | TensorFlow pre-built function |
# | Target results | 77.4% top-1 accuracy (beat [DeepFood paper](https://arxiv.org/abs/1606.05675)) | 50.76% top-1 accuracy (beat [Food101 paper](https://data.vision.ee.ethz.ch/cvl/datasets_extra/food-101/static/bossard_eccv14_food-101.pdf)) |
#
# *Table comparing difference between Food Vision Big (this notebook) versus Food Vision mini (previous notebook).*
#
# Alongside attempting to beat the DeepFood paper, we're going to learn about two methods to significantly improve the speed of our model training:
# 1. Prefetching
# 2. Mixed precision training
#
# But more on these later.
#
# ## What we're going to cover
#
# * Using TensorFlow Datasets to download and explore data
# * Creating preprocessing function for our data
# * Batching & preparing datasets for modelling (**making our datasets run fast**)
# * Creating modelling callbacks
# * Setting up **mixed precision training**
# * Building a feature extraction model (see [transfer learning part 1: feature extraction](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/04_transfer_learning_in_tensorflow_part_1_feature_extraction.ipynb))
# * Fine-tuning the feature extraction model (see [transfer learning part 2: fine-tuning](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/05_transfer_learning_in_tensorflow_part_2_fine_tuning.ipynb))
# * Viewing training results on TensorBoard
#
# ## How you should approach this notebook
#
# You can read through the descriptions and the code (it should all run, except for the cells which error on purpose), but there's a better option.
#
# Write all of the code yourself.
#
# Yes. I'm serious. Create a new notebook, and rewrite each line by yourself. Investigate it, see if you can break it, why does it break?
#
# You don't have to write the text descriptions but writing the code yourself is a great way to get hands-on experience.
#
# Don't worry if you make mistakes, we all do. The way to get better and make less mistakes is to write more code.
#
# > 📖 **Resource:** See the full set of course materials on GitHub: https://github.com/mrdbourke/tensorflow-deep-learning
# + [markdown] id="rLaDq25mykWN"
# ## Check GPU
#
# For this notebook, we're going to be doing something different.
#
# We're going to be using mixed precision training.
#
# Mixed precision training was introduced in [TensorFlow 2.4.0](https://blog.tensorflow.org/2020/12/whats-new-in-tensorflow-24.html) (a very new feature at the time of writing).
#
# What does **mixed precision training** do?
#
# Mixed precision training uses a combination of single precision (float32) and half-preicison (float16) data types to speed up model training (up 3x on modern GPUs).
#
# We'll talk about this more later on but in the meantime you can read the [TensorFlow documentation on mixed precision](https://www.tensorflow.org/guide/mixed_precision) for more details.
#
# For now, before we can move forward if we want to use mixed precision training, we need to make sure the GPU powering our Google Colab instance (if you're using Google Colab) is compataible.
#
# For mixed precision training to work, you need access to a GPU with a compute compability score of 7.0+.
#
# Google Colab offers P100, K80 and T4 GPUs, however, **the P100 and K80 aren't compatible with mixed precision training**.
#
# Therefore before we proceed we need to make sure we have **access to a Tesla T4 GPU in our Google Colab instance**.
#
# If you're not using Google Colab, you can find a list of various [Nvidia GPU compute capabilities on Nvidia's developer website](https://developer.nvidia.com/cuda-gpus#compute).
#
# > 🔑 **Note:** If you run the cell below and see a P100 or K80, try going to to Runtime -> Factory Reset Runtime (note: this will remove any saved variables and data from your Colab instance) and then retry to get a T4.
# + colab={"base_uri": "https://localhost:8080/"} id="VAC_5rYJicZ4" outputId="7c6def0d-e61f-4361-e18a-ff3589d2b978"
# If using Google Colab, this should output "Tesla T4" otherwise,
# you won't be able to use mixed precision training
# !nvidia-smi -L
# + [markdown] id="oWgb38BYKhS_"
# Since mixed precision training was introduced in TensorFlow 2.4.0, make sure you've got at least TensorFlow 2.4.0+.
# + colab={"base_uri": "https://localhost:8080/"} id="8LpEDWLxKg46" outputId="aef28f24-4395-46f6-8129-6cbf84caa68a"
# Check TensorFlow version (should be 2.4.0+)
import tensorflow as tf
print(tf.__version__)
# + [markdown] id="pPwSfuFDzT5v"
# ## Get helper functions
#
# We've created a series of helper functions throughout the previous notebooks in the course. Instead of rewriting them (tedious), we'll import the [`helper_functions.py`](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/extras/helper_functions.py) file from the GitHub repo.
# + colab={"base_uri": "https://localhost:8080/"} id="iC2R6bOZzhQd" outputId="cfbe9a9f-fe5d-4497-c1de-6ac93013dfc0"
# Get helper functions file
# !wget https://raw.githubusercontent.com/mrdbourke/tensorflow-deep-learning/main/extras/helper_functions.py
# + id="ZqKKuFt7zYvf"
# Import series of helper functions for the notebook (we've created/used these in previous notebooks)
from helper_functions import create_tensorboard_callback, plot_loss_curves, compare_historys
# + [markdown] id="w5BE7WYl9b_8"
# ## Use TensorFlow Datasets to Download Data
#
# In previous notebooks, we've downloaded our food images (from the [Food101 dataset](https://www.kaggle.com/dansbecker/food-101/home)) from Google Storage.
#
# And this is a typical workflow you'd use if you're working on your own datasets.
#
# However, there's another way to get datasets ready to use with TensorFlow.
#
# For many of the most popular datasets in the machine learning world (often referred to and used as benchmarks), you can access them through [TensorFlow Datasets (TFDS)](https://www.tensorflow.org/datasets/overview).
#
# What is **TensorFlow Datasets**?
#
# A place for prepared and ready-to-use machine learning datasets.
#
# Why use TensorFlow Datasets?
#
# * Load data already in Tensors
# * Practice on well established datasets
# * Experiment with differet data loading techniques (like we're going to use in this notebook)
# * Experiment with new TensorFlow features quickly (such as mixed precision training)
#
# Why *not* use TensorFlow Datasets?
#
# * The datasets are static (they don't change, like your real-world datasets would)
# * Might not be suited for your particular problem (but great for experimenting)
#
# To begin using TensorFlow Datasets we can import it under the alias `tfds`.
#
# + id="YDMExkAG8ztE"
# Get TensorFlow Datasets
import tensorflow_datasets as tfds
# + [markdown] id="-TRPTGvpNuJm"
# To find all of the available datasets in TensorFlow Datasets, you can use the `list_builders()` method.
#
# After doing so, we can check to see if the one we're after (`"food101"`) is present.
# + colab={"base_uri": "https://localhost:8080/"} id="gXA8b2619s0X" outputId="8f0c8971-4a0a-4aec-f371-701022832a76"
# List available datasets
datasets_list = tfds.list_builders() # get all available datasets in TFDS
print("food101" in datasets_list) # is the dataset we're after available?
# + [markdown] id="bUK_zulYNfVY"
# Beautiful! It looks like the dataset we're after is available (note there are plenty more available but we're on Food101).
#
# To get access to the Food101 dataset from the TFDS, we can use the [`tfds.load()`](https://www.tensorflow.org/datasets/api_docs/python/tfds/load) method.
#
# In particular, we'll have to pass it a few parameters to let it know what we're after:
# * `name` (str) : the target dataset (e.g. `"food101"`)
# * `split` (list, optional) : what splits of the dataset we're after (e.g. `["train", "validation"]`)
# * the `split` parameter is quite tricky. See [the documentation for more](https://github.com/tensorflow/datasets/blob/master/docs/splits.md).
# * `shuffle_files` (bool) : whether or not to shuffle the files on download, defaults to `False`
# * `as_supervised` (bool) : `True` to download data samples in tuple format (`(data, label)`) or `False` for dictionary format
# * `with_info` (bool) : `True` to download dataset metadata (labels, number of samples, etc)
#
# > 🔑 **Note:** Calling the `tfds.load()` method will start to download a target dataset to disk if the `download=True` parameter is set (default). This dataset could be 100GB+, so make sure you have space.
# + colab={"base_uri": "https://localhost:8080/", "height": 347, "referenced_widgets": ["c464cf7f692747dd8a9e5c224b72e683", "e19cbd5c9eca4db6b023d501ae96447b", "465a84232f9344b7852bc0315aaf6147", "a15efd3f9c514ce1947bd75330ddc60f", "1630703c5da345df876cd4a38ac062d0", "48c761f4ebf541169f0b674e835081bd", "fc53cb8f72eb45d9987f22fa391e64ff", "5b48d0e1bd664416b3a8da6daedf29eb", "116932efaceb4e6889b9e9a7585a851f", "<KEY>", "<KEY>", "5ffe42dfcc8945058a1211325bb12276", "<KEY>", "02364bec4a004ed4a4b0970e1b572410", "<KEY>", "<KEY>", "16688ead8185417fbe6acf39649c222a", "5bdf272dff1049afac690fe78c530277", "<KEY>", "8f1b23e71fd14fcc97a32f0abdc96419", "daf7463ed2b14751bb7532a91429bf67", "<KEY>", "<KEY>", "7caebdb45e1f41ae88eddfabada5ff67", "da96cbabe8ff449388e052dd55213653", "<KEY>", "41908c7012174ecabfe801a2f5e3dee5", "103a4aded56c4acd81aac7fe81851956", "<KEY>", "6f9a55e4f5fd4f459eac237f02c37829", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "fe172d6537a642be985520c0e66d7801", "<KEY>", "e8ef9677adbf4f368a258d64732d152a", "acc1ea5a5531425c885afe2bdbce4273", "<KEY>", "<KEY>", "0624ceb3768e4c62a220b188c7ed1141", "<KEY>", "<KEY>", "7c0827a68eb141e5a1939597a171c593", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "8a925d410ebb413a808ce40f12e7d91e", "9a9338dbc9c04a078575071d48b25e51", "<KEY>", "<KEY>", "568e801778124de386e7badcf1e58456", "fe96151f8be04763bedf23083b6a6b7b", "142f3e1df4c1470c81be604db7f2aa95", "<KEY>"]} id="ClXZDWng-s8F" outputId="57febdf7-894d-4524-fb3b-9bc63d7c2d36"
# Load in the data (takes about 5-6 minutes in Google Colab)
(train_data, test_data), ds_info = tfds.load(name="food101", # target dataset to get from TFDS
split=["train", "validation"], # what splits of data should we get? note: not all datasets have train, valid, test
shuffle_files=True, # shuffle files on download?
as_supervised=True, # download data in tuple format (sample, label), e.g. (image, label)
with_info=True) # include dataset metadata? if so, tfds.load() returns tuple (data, ds_info)
# + [markdown] id="pSxo6soUwTQl"
# Wonderful! After a few minutes of downloading, we've now got access to entire Food101 dataset (in tensor format) ready for modelling.
#
# Now let's get a little information from our dataset, starting with the class names.
#
# Getting class names from a TensorFlow Datasets dataset requires downloading the "`dataset_info`" variable (by using the `as_supervised=True` parameter in the `tfds.load()` method, **note:** this will only work for supervised datasets in TFDS).
#
# We can access the class names of a particular dataset using the `dataset_info.features` attribute and accessing `names` attribute of the the `"label"` key.
# + colab={"base_uri": "https://localhost:8080/"} id="Zoy8Tu7VR2ji" outputId="e572e94c-d6d3-43bc-e049-ed4d73c84c96"
# Features of Food101 TFDS
ds_info.features
# + colab={"base_uri": "https://localhost:8080/"} id="g2UkCaLsDXaR" outputId="8ac531a8-f951-48d1-f9a1-dfdeabf6ac1b"
# Get class names
class_names = ds_info.features["label"].names
class_names[:10]
# + [markdown] id="TwsBAkGKwh08"
# ### Exploring the Food101 data from TensorFlow Datasets
#
# Now we've downloaded the Food101 dataset from TensorFlow Datasets, how about we do what any good data explorer should?
#
# In other words, "visualize, visualize, visualize".
#
# Let's find out a few details about our dataset:
# * The shape of our input data (image tensors)
# * The datatype of our input data
# * What the labels of our input data look like (e.g. one-hot encoded versus label-encoded)
# * Do the labels match up with the class names?
#
# To do, let's take one sample off the training data (using the [`.take()` method](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#take)) and explore it.
# + id="5eO2qVy3A-CC"
# Take one sample off the training data
train_one_sample = train_data.take(1) # samples are in format (image_tensor, label)
# + [markdown] id="hsZj4K3ETdvB"
# Because we used the `as_supervised=True` parameter in our `tfds.load()` method above, data samples come in the tuple format structure `(data, label)` or in our case `(image_tensor, label)`.
# + colab={"base_uri": "https://localhost:8080/"} id="m--0wDNDTU8S" outputId="27eaedbd-df55-4773-8a14-b35c40c8a4d0"
# What does one sample of our training data look like?
train_one_sample
# + [markdown] id="bP1MeznpTsbM"
# Let's loop through our single training sample and get some info from the `image_tensor` and `label`.
# + colab={"base_uri": "https://localhost:8080/"} id="Zjz4goiHBMO7" outputId="880b2873-15a9-4805-cfef-9e390c9f038f"
# Output info about our training sample
for image, label in train_one_sample:
print(f"""
Image shape: {image.shape}
Image dtype: {image.dtype}
Target class from Food101 (tensor form): {label}
Class name (str form): {class_names[label.numpy()]}
""")
# + [markdown] id="i4_od8dUUSHE"
# Because we set the `shuffle_files=True` parameter in our `tfds.load()` method above, running the cell above a few times will give a different result each time.
#
# Checking these you might notice some of the images have different shapes, for example `(512, 342, 3)` and `(512, 512, 3)` (height, width, color_channels).
#
# Let's see what one of the image tensors from TFDS's Food101 dataset looks like.
# + colab={"base_uri": "https://localhost:8080/"} id="FuZmVEH-WS4b" outputId="74ab49f2-d531-49a3-b0aa-13a0902846bf"
# What does an image tensor from TFDS's Food101 look like?
image
# + colab={"base_uri": "https://localhost:8080/"} id="3jJF7njRVKh6" outputId="5a10af78-6592-4499-f22b-eba95937ffb7"
# What are the min and max values?
tf.reduce_min(image), tf.reduce_max(image)
# + [markdown] id="P2GvO7HjVF5i"
# Alright looks like our image tensors have values of between 0 & 255 (standard red, green, blue colour values) and the values are of data type `unit8`.
#
# We might have to preprocess these before passing them to a neural network. But we'll handle this later.
#
# In the meantime, let's see if we can plot an image sample.
# + [markdown] id="llQyIBfJWc5x"
# ### Plot an image from TensorFlow Datasets
#
# We've seen our image tensors in tensor format, now let's really adhere to our motto.
#
# "Visualize, visualize, visualize!"
#
# Let's plot one of the image samples using [`matplotlib.pyplot.imshow()`](https://matplotlib.org/stable/api/_as_gen/matplotlib.pyplot.imshow.html) and set the title to target class name.
# + colab={"base_uri": "https://localhost:8080/", "height": 264} id="pK581hgPWyLm" outputId="f82b5267-3c9c-4d19-a3b8-6cf4998d4110"
# Plot an image tensor
import matplotlib.pyplot as plt
plt.imshow(image)
plt.title(class_names[label.numpy()]) # add title to image by indexing on class_names list
plt.axis(False);
# + [markdown] id="4mBAtGnPWQHy"
# Delicious!
#
# Okay, looks like the Food101 data we've got from TFDS is similar to the datasets we've been using in previous notebooks.
#
# Now let's preprocess it and get it ready for use with a neural network.
# + [markdown] id="UeRJnQMIYLcy"
# ## Create preprocessing functions for our data
#
# In previous notebooks, when our images were in folder format we used the method [`tf.keras.preprocessing.image_dataset_from_directory()`](https://www.tensorflow.org/api_docs/python/tf/keras/preprocessing/image_dataset_from_directory) to load them in.
#
# Doing this meant our data was loaded into a format ready to be used with our models.
#
# However, since we've downloaded the data from TensorFlow Datasets, there are a couple of preprocessing steps we have to take before it's ready to model.
#
# More specifically, our data is currently:
#
# * In `uint8` data type
# * Comprised of all differnet sized tensors (different sized images)
# * Not scaled (the pixel values are between 0 & 255)
#
# Whereas, models like data to be:
#
# * In `float32` data type
# * Have all of the same size tensors (batches require all tensors have the same shape, e.g. `(224, 224, 3)`)
# * Scaled (values between 0 & 1), also called normalized
#
# To take care of these, we'll create a `preprocess_img()` function which:
#
# * Resizes an input image tensor to a specified size using [`tf.image.resize()`](https://www.tensorflow.org/api_docs/python/tf/image/resize)
# * Converts an input image tensor's current datatype to `tf.float32` using [`tf.cast()`](https://www.tensorflow.org/api_docs/python/tf/cast)
#
# > 🔑 **Note:** Pretrained EfficientNetBX models in [`tf.keras.applications.efficientnet`](https://www.tensorflow.org/api_docs/python/tf/keras/applications/efficientnet) (what we're going to be using) have rescaling built-in. But for many other model architectures you'll want to rescale your data (e.g. get its values between 0 & 1). This could be incorporated inside your "`preprocess_img()`" function (like the one below) or within your model as a [`tf.keras.layers.experimental.preprocessing.Rescaling`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/Rescaling) layer.
# + id="NKuwdjm0CWc1"
# Make a function for preprocessing images
def preprocess_img(image, label, img_shape=224):
"""
Converts image datatype from 'uint8' -> 'float32' and reshapes image to
[img_shape, img_shape, color_channels]
"""
image = tf.image.resize(image, [img_shape, img_shape]) # reshape to img_shape
return tf.cast(image, tf.float32), label # return (float32_image, label) tuple
# + [markdown] id="m6kGGFa1Z3Nz"
# Our `preprocess_img()` function above takes image and label as input (even though it does nothing to the label) because our dataset is currently in the tuple structure `(image, label)`.
#
# Let's try our function out on a target image.
# + colab={"base_uri": "https://localhost:8080/"} id="BqPDUGCvHI4K" outputId="84cccb6f-5ace-4903-e33d-73dfa867c6ee"
# Preprocess a single sample image and check the outputs
preprocessed_img = preprocess_img(image, label)[0]
print(f"Image before preprocessing:\n {image[:2]}...,\nShape: {image.shape},\nDatatype: {image.dtype}\n")
print(f"Image after preprocessing:\n {preprocessed_img[:2]}...,\nShape: {preprocessed_img.shape},\nDatatype: {preprocessed_img.dtype}")
# + [markdown] id="uhIIvprqaHEZ"
# Excellent! Looks like our `preprocess_img()` function is working as expected.
#
# The input image gets converted from `uint8` to `float32` and gets reshaped from its current shape to `(224, 224, 3)`.
#
# How does it look?
# + colab={"base_uri": "https://localhost:8080/", "height": 264} id="wYtMxQzZY0F7" outputId="2dd69f3d-bfb0-4318-efb3-a40e98a558a2"
# We can still plot our preprocessed image as long as we
# divide by 255 (for matplotlib capatibility)
plt.imshow(preprocessed_img/255.)
plt.title(class_names[label])
plt.axis(False);
# + [markdown] id="gsIaJZEU7y_M"
# All this food visualization is making me hungry. How about we start preparing to model it?
# + [markdown] id="t2rd4_3CjdGE"
# ## Batch & prepare datasets
#
# Before we can model our data, we have to turn it into batches.
#
# # Why?
#
# Because computing on batches is memory efficient.
#
# We turn our data from 101,000 image tensors and labels (train and test combined) into batches of 32 image and label pairs, thus enabling it to fit into the memory of our GPU.
#
# To do this in effective way, we're going to be leveraging a number of methods from the [`tf.data` API](https://www.tensorflow.org/api_docs/python/tf/data).
#
# > 📖 **Resource:** For loading data in the most performant way possible, see the TensorFlow docuemntation on [Better performance with the tf.data API](https://www.tensorflow.org/guide/data_performance).
#
# Specifically, we're going to be using:
#
# * [`map()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#map) - maps a predefined function to a target dataset (e.g. `preprocess_img()` to our image tensors)
# * [`shuffle()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#shuffle) - randomly shuffles the elements of a target dataset up `buffer_size` (ideally, the `buffer_size` is equal to the size of the dataset, however, this may have implications on memory)
# * [`batch()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#batch) - turns elements of a target dataset into batches (size defined by parameter `batch_size`)
# * [`prefetch()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#prefetch) - prepares subsequent batches of data whilst other batches of data are being computed on (improves data loading speed but costs memory)
# * Extra: [`cache()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#cache) - caches (saves them for later) elements in a target dataset, saving loading time (will only work if your dataset is small enough to fit in memory, standard Colab instances only have 12GB of memory)
#
# Things to note:
# - Can't batch tensors of different shapes (e.g. different image sizes, need to reshape images first, hence our `preprocess_img()` function)
# - `shuffle()` keeps a buffer of the number you pass it images shuffled, ideally this number would be all of the samples in your training set, however, if your training set is large, this buffer might not fit in memory (a fairly large number like 1000 or 10000 is usually suffice for shuffling)
# - For methods with the `num_parallel_calls` parameter available (such as `map()`), setting it to`num_parallel_calls=tf.data.AUTOTUNE` will parallelize preprocessing and significantly improve speed
# - Can't use `cache()` unless your dataset can fit in memory
#
# Woah, the above is alot. But once we've coded below, it'll start to make sense.
#
# We're going to through things in the following order:
#
# ```
# Original dataset (e.g. train_data) -> map() -> shuffle() -> batch() -> prefetch() -> PrefetchDataset
# ```
#
# This is like saying,
#
# > "Hey, map this preprocessing function across our training dataset, then shuffle a number of elements before batching them together and make sure you prepare new batches (prefetch) whilst the model is looking through the current batch".
#
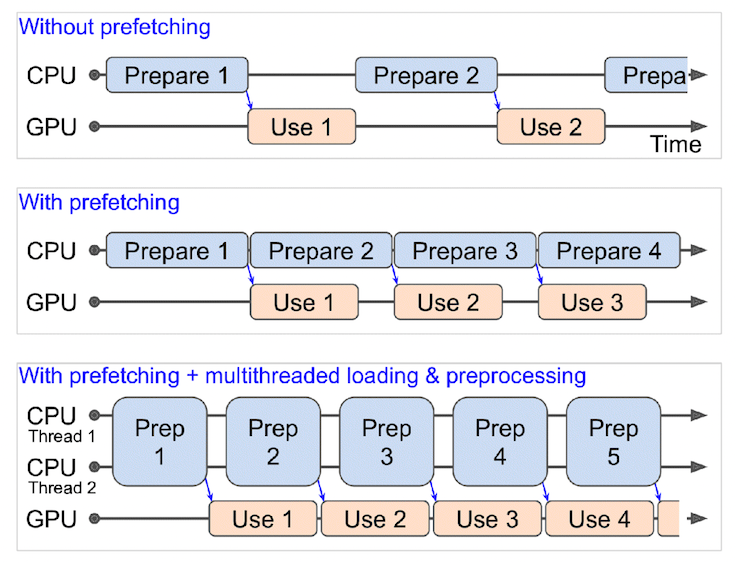
# 
#
# *What happens when you use prefetching (faster) versus what happens when you don't use prefetching (slower). **Source:** Page 422 of [Hands-On Machine Learning with Scikit-Learn, Keras & TensorFlow Book by <NAME>](https://www.oreilly.com/library/view/hands-on-machine-learning/9781492032632/).*
#
# + id="VhA4gq-pI2W3"
# Map preprocessing function to training data (and paralellize)
train_data = train_data.map(map_func=preprocess_img, num_parallel_calls=tf.data.AUTOTUNE)
# Shuffle train_data and turn it into batches and prefetch it (load it faster)
train_data = train_data.shuffle(buffer_size=1000).batch(batch_size=32).prefetch(buffer_size=tf.data.AUTOTUNE)
# Map prepreprocessing function to test data
test_data = test_data.map(preprocess_img, num_parallel_calls=tf.data.AUTOTUNE)
# Turn test data into batches (don't need to shuffle)
test_data = test_data.batch(32).prefetch(tf.data.AUTOTUNE)
# + [markdown] id="rnTPWyAhlKO3"
# And now let's check out what our prepared datasets look like.
# + colab={"base_uri": "https://localhost:8080/"} id="5_fBkGqfJFxT" outputId="176eab2e-d62a-433e-f7c1-32f111719e07"
train_data, test_data
# + [markdown] id="X1fxgyWnlQNU"
# Excellent! Looks like our data is now in tutples of `(image, label)` with datatypes of `(tf.float32, tf.int64)`, just what our model is after.
#
# > 🔑 **Note:** You can get away without calling the `prefetch()` method on the end of your datasets, however, you'd probably see significantly slower data loading speeds when building a model. So most of your dataset input pipelines should end with a call to [`prefecth()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#prefetch).
#
# Onward.
# + [markdown] id="Qj3umnpMvSw8"
# ## Create modelling callbacks
#
# Since we're going to be training on a large amount of data and training could take a long time, it's a good idea to set up some modelling callbacks so we be sure of things like our model's training logs being tracked and our model being checkpointed (saved) after various training milestones.
#
# To do each of these we'll use the following callbacks:
# * [`tf.keras.callbacks.TensorBoard()`](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/TensorBoard) - allows us to keep track of our model's training history so we can inspect it later (**note:** we've created this callback before have imported it from `helper_functions.py` as `create_tensorboard_callback()`)
# * [`tf.keras.callbacks.ModelCheckpoint()`](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/ModelCheckpoint) - saves our model's progress at various intervals so we can load it and resuse it later without having to retrain it
# * Checkpointing is also helpful so we can start fine-tuning our model at a particular epoch and revert back to a previous state if fine-tuning offers no benefits
# + id="wyYmxPnlXOwd"
# Create TensorBoard callback (already have "create_tensorboard_callback()" from a previous notebook)
from helper_functions import create_tensorboard_callback
# Create ModelCheckpoint callback to save model's progress
checkpoint_path = "model_checkpoints/cp.ckpt" # saving weights requires ".ckpt" extension
model_checkpoint = tf.keras.callbacks.ModelCheckpoint(checkpoint_path,
montior="val_acc", # save the model weights with best validation accuracy
save_best_only=True, # only save the best weights
save_weights_only=True, # only save model weights (not whole model)
verbose=0) # don't print out whether or not model is being saved
# + [markdown] id="DyXlCU50UElG"
# ## Setup mixed precision training
#
# We touched on mixed precision training above.
#
# However, we didn't quite explain it.
#
# Normally, tensors in TensorFlow default to the float32 datatype (unless otherwise specified).
#
# In computer science, float32 is also known as [single-precision floating-point format](https://en.wikipedia.org/wiki/Single-precision_floating-point_format). The 32 means it usually occupies 32 bits in computer memory.
#
# Your GPU has a limited memory, therefore it can only handle a number of float32 tensors at the same time.
#
# This is where mixed precision training comes in.
#
# Mixed precision training involves using a mix of float16 and float32 tensors to make better use of your GPU's memory.
#
# Can you guess what float16 means?
#
# Well, if you thought since float32 meant single-precision floating-point, you might've guessed float16 means [half-precision floating-point format](https://en.wikipedia.org/wiki/Half-precision_floating-point_format). And if you did, you're right! And if not, no trouble, now you know.
#
# For tensors in float16 format, each element occupies 16 bits in computer memory.
#
# So, where does this leave us?
#
# As mentioned before, when using mixed precision training, your model will make use of float32 and float16 data types to use less memory where possible and in turn run faster (using less memory per tensor means more tensors can be computed on simultaneously).
#
# As a result, using mixed precision training can improve your performance on modern GPUs (those with a compute capability score of 7.0+) by up to 3x.
#
# For a more detailed explanation, I encourage you to read through the [TensorFlow mixed precision guide](https://www.tensorflow.org/guide/mixed_precision) (I'd highly recommend at least checking out the summary).
#
# 
# *Because mixed precision training uses a combination of float32 and float16 data types, you may see up to a 3x speedup on modern GPUs.*
#
# > 🔑 **Note:** If your GPU doesn't have a score of over 7.0+ (e.g. P100 in Colab), mixed precision won't work (see: ["Supported Hardware"](https://www.tensorflow.org/guide/mixed_precision#supported_hardware) in the mixed precision guide for more).
#
# > 📖 **Resource:** If you'd like to learn more about precision in computer science (the detail to which a numerical quantity is expressed by a computer), see the [Wikipedia page](https://en.wikipedia.org/wiki/Precision_(computer_science)) (and accompanying resources).
#
# Okay, enough talk, let's see how we can turn on mixed precision training in TensorFlow.
#
# The beautiful thing is, the [`tensorflow.keras.mixed_precision`](https://www.tensorflow.org/api_docs/python/tf/keras/mixed_precision/) API has made it very easy for us to get started.
#
# First, we'll import the API and then use the [`set_global_policy()`](https://www.tensorflow.org/api_docs/python/tf/keras/mixed_precision/set_global_policy) method to set the *dtype policy* to `"mixed_float16"`.
#
# + colab={"base_uri": "https://localhost:8080/"} id="5BuEjmlybR7V" outputId="ebb171e8-4ae9-406d-c771-6cdf4f5b2a8d"
# Turn on mixed precision training
from tensorflow.keras import mixed_precision
mixed_precision.set_global_policy(policy="mixed_float16") # set global policy to mixed precision
# + [markdown] id="OLxlu7VyYoQm"
# Nice! As long as the GPU you're using has a compute capability of 7.0+ the cell above should run without error.
#
# Now we can check the global dtype policy (the policy which will be used by layers in our model) using the [`mixed_precision.global_policy()`](https://www.tensorflow.org/api_docs/python/tf/keras/mixed_precision/global_policy) method.
# + colab={"base_uri": "https://localhost:8080/"} id="qzSWJP8KkKae" outputId="90ae40e6-e28f-4c16-b565-0114cff046e2"
mixed_precision.global_policy() # should output "mixed_float16"
# + [markdown] id="gpnAW2ltXCpE"
# Great, since the global dtype policy is now `"mixed_float16"` our model will automatically take advantage of float16 variables where possible and in turn speed up training.
# + [markdown] id="rA8FBJwwvVoG"
# ## Build feature extraction model
#
# Callbacks: ready to roll.
#
# Mixed precision: turned on.
#
# Let's build a model.
#
# Because our dataset is quite large, we're going to move towards fine-tuning an existing pretrained model (EfficienetNetB0).
#
# But before we get into fine-tuning, let's set up a feature-extraction model.
#
# Recall, the typical order for using transfer learning is:
#
# 1. Build a feature extraction model (replace the top few layers of a pretrained model)
# 2. Train for a few epochs with lower layers frozen
# 3. Fine-tune if necessary with multiple layers unfrozen
#
# 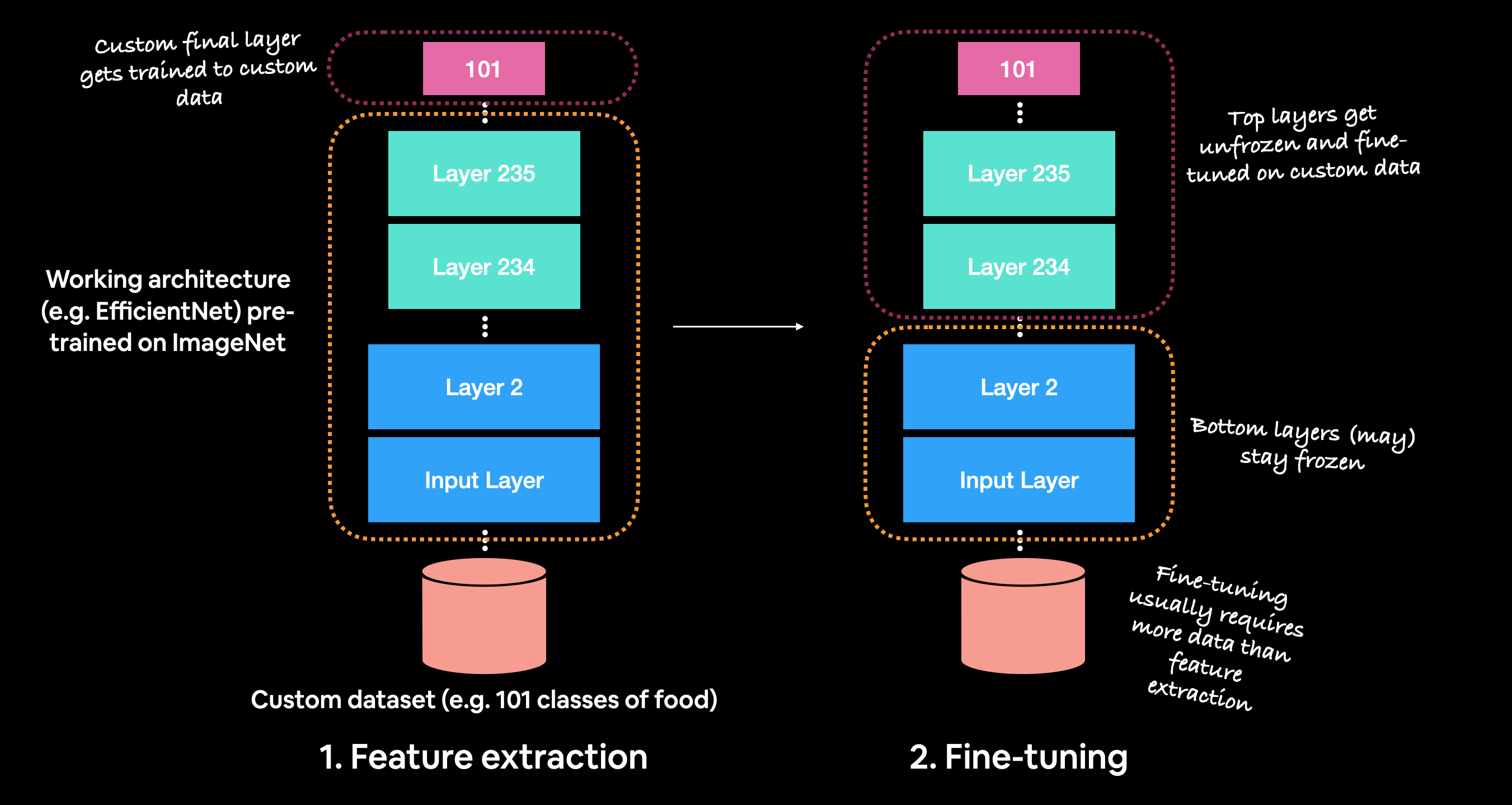
# *Before fine-tuning, it's best practice to train a feature extraction model with custom top layers.*
#
# To build the feature extraction model (covered in [Transfer Learning in TensorFlow Part 1: Feature extraction](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/04_transfer_learning_in_tensorflow_part_1_feature_extraction.ipynb)), we'll:
# * Use `EfficientNetB0` from [`tf.keras.applications`](https://www.tensorflow.org/api_docs/python/tf/keras/applications) pre-trained on ImageNet as our base model
# * We'll download this without the top layers using `include_top=False` parameter so we can create our own output layers
# * Freeze the base model layers so we can use the pre-learned patterns the base model has found on ImageNet
# * Put together the input, base model, pooling and output layers in a [Functional model](https://keras.io/guides/functional_api/)
# * Compile the Functional model using the Adam optimizer and [sparse categorical crossentropy](https://www.tensorflow.org/api_docs/python/tf/keras/losses/SparseCategoricalCrossentropy) as the loss function (since our labels **aren't** one-hot encoded)
# * Fit the model for 3 epochs using the TensorBoard and ModelCheckpoint callbacks
#
# > 🔑 **Note:** Since we're using mixed precision training, our model needs a separate output layer with a hard-coded `dtype=float32`, for example, `layers.Activation("softmax", dtype=tf.float32)`. This ensures the outputs of our model are returned back to the float32 data type which is more numerically stable than the float16 datatype (important for loss calculations). See the ["Building the model"](https://www.tensorflow.org/guide/mixed_precision#building_the_model) section in the TensorFlow mixed precision guide for more.
#
# 
# *Turning mixed precision on in TensorFlow with 3 lines of code.*
# + colab={"base_uri": "https://localhost:8080/"} id="GrkWpCzfXKE7" outputId="d3ae00b1-4c19-4d6e-c25f-5d0114a9168a"
from tensorflow.keras import layers
from tensorflow.keras.layers.experimental import preprocessing
# Create base model
input_shape = (224, 224, 3)
base_model = tf.keras.applications.EfficientNetB0(include_top=False)
base_model.trainable = False # freeze base model layers
# Create Functional model
inputs = layers.Input(shape=input_shape, name="input_layer")
# Note: EfficientNetBX models have rescaling built-in but if your model didn't you could have a layer like below
# x = preprocessing.Rescaling(1./255)(x)
x = base_model(inputs, training=False) # set base_model to inference mode only
x = layers.GlobalAveragePooling2D(name="pooling_layer")(x)
x = layers.Dense(len(class_names))(x) # want one output neuron per class
# Separate activation of output layer so we can output float32 activations
outputs = layers.Activation("softmax", dtype=tf.float32, name="softmax_float32")(x)
model = tf.keras.Model(inputs, outputs)
# Compile the model
model.compile(loss="sparse_categorical_crossentropy", # Use sparse_categorical_crossentropy when labels are *not* one-hot
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# + colab={"base_uri": "https://localhost:8080/"} id="wfEG8ud_jsNY" outputId="82b20036-498f-4efa-9ae1-d9e9e5c728e8"
# Check out our model
model.summary()
# + [markdown] id="lIXkEdnNGpKi"
# ## Checking layer dtype policies (are we using mixed precision?)
#
# Model ready to go!
#
# Before we said the mixed precision API will automatically change our layers' dtype policy's to whatever the global dtype policy is (in our case it's `"mixed_float16"`).
#
# We can check this by iterating through our model's layers and printing layer attributes such as `dtype` and `dtype_policy`.
# + colab={"base_uri": "https://localhost:8080/"} id="Zk__ebBLHC-Q" outputId="e94ff2a5-6adb-4745-fcd0-8cb2f2d5a083"
# Check the dtype_policy attributes of layers in our model
for layer in model.layers:
print(layer.name, layer.trainable, layer.dtype, layer.dtype_policy) # Check the dtype policy of layers
# + [markdown] id="7w6Gv6ySfpNY"
# Going through the above we see:
# * `layer.name` (str) : a layer's human-readable name, can be defined by the `name` parameter on construction
# * `layer.trainable` (bool) : whether or not a layer is trainable (all of our layers are trainable except the efficientnetb0 layer since we set it's `trainable` attribute to `False`
# * `layer.dtype` : the data type a layer stores its variables in
# * `layer.dtype_policy` : the data type a layer computes in
#
# > 🔑 **Note:** A layer can have a dtype of `float32` and a dtype policy of `"mixed_float16"` because it stores its variables (weights & biases) in `float32` (more numerically stable), however it computes in `float16` (faster).
#
# We can also check the same details for our model's base model.
#
# + colab={"base_uri": "https://localhost:8080/"} id="eL_THJCYGenQ" outputId="5339f5f1-0179-4b56-b65a-80ec350aee45"
# Check the layers in the base model and see what dtype policy they're using
for layer in model.layers[1].layers[:20]: # only check the first 20 layers to save output space
print(layer.name, layer.trainable, layer.dtype, layer.dtype_policy)
# + [markdown] id="GerkBr7GiDIj"
# > 🔑 **Note:** The mixed precision API automatically causes layers which can benefit from using the `"mixed_float16"` dtype policy to use it. It also prevents layers which shouldn't use it from using it (e.g. the normalization layer at the start of the base model).
# + [markdown] id="NJz5S66ojyUS"
# ## Fit the feature extraction model
#
# Now that's one good looking model. Let's fit it to our data shall we?
#
# Three epochs should be enough for our top layers to adjust their weights enough to our food image data.
#
# To save time per epoch, we'll also only validate on 15% of the test data.
# + colab={"base_uri": "https://localhost:8080/"} id="4v7rXZG-ZkNJ" outputId="e7f6f934-538b-42e1-f732-0bfd3b0ff13b"
# Fit the model with callbacks
history_101_food_classes_feature_extract = model.fit(train_data,
epochs=3,
steps_per_epoch=len(train_data),
validation_data=test_data,
validation_steps=int(0.15 * len(test_data)),
callbacks=[create_tensorboard_callback("training_logs",
"efficientnetb0_101_classes_all_data_feature_extract"),
model_checkpoint])
# + [markdown] id="xg01Gh3EnQSu"
# Nice, looks like our feature extraction model is performing pretty well. How about we evaluate it on the whole test dataset?
# + colab={"base_uri": "https://localhost:8080/"} id="jhV7fvTreV27" outputId="ca5d6920-1d60-46b4-abd3-38f0e8776718"
# Evaluate model (unsaved version) on whole test dataset
results_feature_extract_model = model.evaluate(test_data)
results_feature_extract_model
# + [markdown] id="TI0li4ZenctF"
# And since we used the `ModelCheckpoint` callback, we've got a saved version of our model in the `model_checkpoints` directory.
#
# Let's load it in and make sure it performs just as well.
# + [markdown] id="nNGoI1cS21um"
# ## Load and evaluate checkpoint weights
#
# We can load in and evaluate our model's checkpoints by:
#
# 1. Cloning our model using [`tf.keras.models.clone_model()`](https://www.tensorflow.org/api_docs/python/tf/keras/models/clone_model) to make a copy of our feature extraction model with reset weights.
# 2. Calling the `load_weights()` method on our cloned model passing it the path to where our checkpointed weights are stored.
# 3. Calling `evaluate()` on the cloned model with loaded weights.
#
# A reminder, checkpoints are helpful for when you perform an experiment such as fine-tuning your model. In the case you fine-tune your feature extraction model and find it doesn't offer any improvements, you can always revert back to the checkpointed version of your model.
# + colab={"base_uri": "https://localhost:8080/"} id="C5a_eh9RKBSY" outputId="71986b44-476b-4252-a8fc-2f4e1d77f073"
# Clone the model we created (this resets all weights)
cloned_model = tf.keras.models.clone_model(model)
cloned_model.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="3NBOvb9bkAHa" outputId="0ea80b8d-9570-4148-a784-f6fac093bc5e"
# Where are our checkpoints stored?
checkpoint_path
# + colab={"base_uri": "https://localhost:8080/"} id="mnagdIagKGZY" outputId="2787e959-6681-48fb-c37a-a324fad7f026"
# Load checkpointed weights into cloned_model
cloned_model.load_weights(checkpoint_path)
# + [markdown] id="Wh_-7URJlapr"
# Each time you make a change to your model (including loading weights), you have to recompile.
# + id="So0ybUwRKNSf"
# Compile cloned_model (with same parameters as original model)
cloned_model.compile(loss="sparse_categorical_crossentropy",
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# + colab={"base_uri": "https://localhost:8080/"} id="aZtbkOhHKKLs" outputId="c7e27c13-3a60-4a75-fdb3-ec32fdcb260d"
# Evalaute cloned model with loaded weights (should be same score as trained model)
results_cloned_model_with_loaded_weights = cloned_model.evaluate(test_data)
# + [markdown] id="_441Jd7PlkN4"
# Our cloned model with loaded weight's results should be very close to the feature extraction model's results (if the cell below errors, something went wrong).
# + id="zbcuQQgz5tNX"
# Loaded checkpoint weights should return very similar results to checkpoint weights prior to saving
import numpy as np
assert np.isclose(results_feature_extract_model, results_cloned_model_with_loaded_weights).all() # check if all elements in array are close
# + [markdown] id="n6j46R_L3VED"
# Cloning the model preserves `dtype_policy`'s of layers (but doesn't preserve weights) so if we wanted to continue fine-tuning with the cloned model, we could and it would still use the mixed precision dtype policy.
# + colab={"base_uri": "https://localhost:8080/"} id="YjCO0tm9Je3N" outputId="d894df48-2202-4f34-c079-3d619f4f4345"
# Check the layers in the base model and see what dtype policy they're using
for layer in cloned_model.layers[1].layers[:20]: # check only the first 20 layers to save space
print(layer.name, layer.trainable, layer.dtype, layer.dtype_policy)
# + [markdown] id="EvTGiIFv3eOe"
# ## Save the whole model to file
#
# We can also save the whole model using the [`save()`](https://www.tensorflow.org/api_docs/python/tf/keras/Model#save) method.
#
# Since our model is quite large, you might want to save it to Google Drive (if you're using Google Colab) so you can load it in for use later.
#
# > 🔑 **Note:** Saving to Google Drive requires mounting Google Drive (go to Files -> Mount Drive).
# + id="CH4jkVPBoPhe"
# ## Saving model to Google Drive (optional)
# # Create save path to drive
# save_dir = "drive/MyDrive/tensorflow_course/food_vision/07_efficientnetb0_feature_extract_model_mixed_precision/"
# # os.makedirs(save_dir) # Make directory if it doesn't exist
# # Save model
# model.save(save_dir)
# + [markdown] id="-P1L5fiwnApE"
# We can also save it directly to our Google Colab instance.
#
# > 🔑 **Note:** Google Colab storage is ephemeral and your model will delete itself (along with any other saved files) when the Colab session expires.
# + colab={"base_uri": "https://localhost:8080/"} id="RHKn4Ex57wzF" outputId="6f8706a9-d2f5-4774-b68f-8f59c40ab789"
# Save model locally (if you're using Google Colab, your saved model will Colab instance terminates)
save_dir = "07_efficientnetb0_feature_extract_model_mixed_precision"
model.save(save_dir)
# + [markdown] id="QKiEXBC6n83F"
# And again, we can check whether or not our model saved correctly by loading it in and evaluating it.
# + id="bKGDBKrU6rej"
# Load model previously saved above
loaded_saved_model = tf.keras.models.load_model(save_dir)
# + [markdown] id="CT50keE46i9y"
# Loading a `SavedModel` also retains all of the underlying layers `dtype_policy` (we want them to be `"mixed_float16"`).
# + colab={"base_uri": "https://localhost:8080/"} id="fUJXpxmMKnYV" outputId="8c0e1986-9282-48cd-c98a-e50c130afb46"
# Check the layers in the base model and see what dtype policy they're using
for layer in loaded_saved_model.layers[1].layers[:20]: # check only the first 20 layers to save output space
print(layer.name, layer.trainable, layer.dtype, layer.dtype_policy)
# + colab={"base_uri": "https://localhost:8080/"} id="5Qym9gSm6vL_" outputId="f1b77e21-a61e-47c8-b628-f532f36d4026"
# Check loaded model performance (this should be the same as results_feature_extract_model)
results_loaded_saved_model = loaded_saved_model.evaluate(test_data)
results_loaded_saved_model
# + id="4BUhHSXI-l0v"
# The loaded model's results should equal (or at least be very close) to the model's results prior to saving
# Note: this will only work if you've instatiated results variables
import numpy as np
assert np.isclose(results_feature_extract_model, results_loaded_saved_model).all()
# + [markdown] id="mXlDCU8zoUiK"
# That's what we want! Our loaded model performing as it should.
#
# > 🔑 **Note:** We spent a fair bit of time making sure our model saved correctly because training on a lot of data can be time-consuming, so we want to make sure we don't have to continaully train from scratch.
# + [markdown] id="bj21OVxBGlw9"
# ## Preparing our model's layers for fine-tuning
#
# Our feature-extraction model is showing some great promise after three epochs. But since we've got so much data, it's probably worthwhile that we see what results we can get with fine-tuning (fine-tuning usually works best when you've got quite a large amount of data).
#
# Remember our goal of beating the [DeepFood paper](https://arxiv.org/pdf/1606.05675.pdf)?
#
# They were able to achieve 77.4% top-1 accuracy on Food101 over 2-3 days of training.
#
# Do you think fine-tuning will get us there?
#
# Let's find out.
#
# To start, let's load in our saved model.
#
# > 🔑 **Note:** It's worth remembering a traditional workflow for fine-tuning is to freeze a pre-trained base model and then train only the output layers for a few iterations so their weights can be updated inline with your custom data (feature extraction). And then unfreeze a number or all of the layers in the base model and continue training until the model stops improving.
#
# Like all good cooking shows, I've saved a model I prepared earlier (the feature extraction model from above) to Google Storage.
#
# We can download it to make sure we're using the same model going forward.
# + colab={"base_uri": "https://localhost:8080/"} id="veoEmC6V-cZv" outputId="a52eff70-816d-4854-f092-050f4d905f63"
# Download the saved model from Google Storage
# !wget https://storage.googleapis.com/ztm_tf_course/food_vision/07_efficientnetb0_feature_extract_model_mixed_precision.zip
# + colab={"base_uri": "https://localhost:8080/"} id="F5uIf_0J-jRt" outputId="20f79229-9b94-4703-a6b8-a16662b59ff4"
# Unzip the SavedModel downloaded from Google Stroage
# !mkdir downloaded_gs_model # create new dir to store downloaded feature extraction model
# !unzip 07_efficientnetb0_feature_extract_model_mixed_precision.zip -d downloaded_gs_model
# + id="Xbs5ywA6_CWV"
# Load and evaluate downloaded GS model
loaded_gs_model = tf.keras.models.load_model("/content/downloaded_gs_model/07_efficientnetb0_feature_extract_model_mixed_precision")
# + colab={"base_uri": "https://localhost:8080/"} id="YEZSqPs6pQxy" outputId="4237c482-54c0-4534-99e6-4531113720b3"
# Get a summary of our downloaded model
loaded_gs_model.summary()
# + [markdown] id="AH6sS_DNzSe_"
# And now let's make sure our loaded model is performing as expected.
# + colab={"base_uri": "https://localhost:8080/"} id="IlGs5V3Tosx3" outputId="a325d32a-845b-415b-da8b-ae5edfd8c93e"
# How does the loaded model perform?
results_loaded_gs_model = loaded_gs_model.evaluate(test_data)
results_loaded_gs_model
# + [markdown] id="2ZokTvmxzXv8"
# Great, our loaded model is performing as expected.
#
# When we first created our model, we froze all of the layers in the base model by setting `base_model.trainable=False` but since we've loaded in our model from file, let's check whether or not the layers are trainable or not.
# + colab={"base_uri": "https://localhost:8080/"} id="S-RQ633CapPk" outputId="07a27634-668f-40bd-e1fc-8ce54b43f70c"
# Are any of the layers in our model frozen?
for layer in loaded_gs_model.layers:
layer.trainable = True # set all layers to trainable
print(layer.name, layer.trainable, layer.dtype, layer.dtype_policy) # make sure loaded model is using mixed precision dtype_policy ("mixed_float16")
# + [markdown] id="tTzYLrzGzs6W"
# Alright, it seems like each layer in our loaded model is trainable. But what if we got a little deeper and inspected each of the layers in our base model?
#
# > 🤔 **Question:** *Which layer in the loaded model is our base model?*
#
# Before saving the Functional model to file, we created it with five layers (layers below are 0-indexed):
# 0. The input layer
# 1. The pre-trained base model layer (`tf.keras.applications.EfficientNetB0`)
# 2. The pooling layer
# 3. The fully-connected (dense) layer
# 4. The output softmax activation (with float32 dtype)
#
# Therefore to inspect our base model layer, we can access the `layers` attribute of the layer at index 1 in our model.
# + colab={"base_uri": "https://localhost:8080/"} id="rob4ClPIa2hp" outputId="a4670a8a-4572-41b1-edbe-88fb084effad"
# Check the layers in the base model and see what dtype policy they're using
for layer in loaded_gs_model.layers[1].layers[:20]:
print(layer.name, layer.trainable, layer.dtype, layer.dtype_policy)
# + [markdown] id="BRmD0IvT1bR_"
# Wonderful, it looks like each layer in our base model is trainable (unfrozen) and every layer which should be using the dtype policy `"mixed_policy16"` is using it.
#
# Since we've got so much data (750 images x 101 training classes = 75750 training images), let's keep all of our base model's layers unfrozen.
#
# > 🔑 **Note:** If you've got a small amount of data (less than 100 images per class), you may want to only unfreeze and fine-tune a small number of layers in the base model at a time. Otherwise, you risk overfitting.
# + [markdown] id="m6_6m5jb2Nea"
# ## A couple more callbacks
#
# We're about to start fine-tuning a deep learning model with over 200 layers using over 100,000 (75k+ training, 25K+ testing) images, which means our model's training time is probably going to be much longer than before.
#
# > 🤔 **Question:** *How long does training take?*
#
# It could be a couple of hours or in the case of the [DeepFood paper](https://arxiv.org/pdf/1606.05675.pdf) (the baseline we're trying to beat), their best performing model took 2-3 days of training time.
#
# You will really only know how long it'll take once you start training.
#
# > 🤔 **Question:** *When do you stop training?*
#
# Ideally, when your model stops improving. But again, due to the nature of deep learning, it can be hard to know when exactly a model will stop improving.
#
# Luckily, there's a solution: the [`EarlyStopping` callback](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/EarlyStopping).
#
# The `EarlyStopping` callback monitors a specified model performance metric (e.g. `val_loss`) and when it stops improving for a specified number of epochs, automatically stops training.
#
# Using the `EarlyStopping` callback combined with the `ModelCheckpoint` callback saving the best performing model automatically, we could keep our model training for an unlimited number of epochs until it stops improving.
#
# Let's set both of these up to monitor our model's `val_loss`.
# + id="GcKFVlXVwjJy"
# Setup EarlyStopping callback to stop training if model's val_loss doesn't improve for 3 epochs
early_stopping = tf.keras.callbacks.EarlyStopping(monitor="val_loss", # watch the val loss metric
patience=3) # if val loss decreases for 3 epochs in a row, stop training
# Create ModelCheckpoint callback to save best model during fine-tuning
checkpoint_path = "fine_tune_checkpoints/"
model_checkpoint = tf.keras.callbacks.ModelCheckpoint(checkpoint_path,
save_best_only=True,
monitor="val_loss")
# + [markdown] id="14cdwkIi4WnG"
# Woohoo! Fine-tuning callbacks ready.
#
# If you're planning on training large models, the `ModelCheckpoint` and `EarlyStopping` are two callbacks you'll want to become very familiar with.
#
# We're almost ready to start fine-tuning our model but there's one more callback we're going to implement: [`ReduceLROnPlateau`](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/ReduceLROnPlateau).
#
# Remember how the learning rate is the most important model hyperparameter you can tune? (if not, treat this as a reminder).
#
# Well, the `ReduceLROnPlateau` callback helps to tune the learning rate for you.
#
# Like the `ModelCheckpoint` and `EarlyStopping` callbacks, the `ReduceLROnPlateau` callback montiors a specified metric and when that metric stops improving, it reduces the learning rate by a specified factor (e.g. divides the learning rate by 10).
#
# > 🤔 **Question:** *Why lower the learning rate?*
#
# Imagine having a coin at the back of the couch and you're trying to grab with your fingers.
#
# Now think of the learning rate as the size of the movements your hand makes towards the coin.
#
# The closer you get, the smaller you want your hand movements to be, otherwise the coin will be lost.
#
# Our model's ideal performance is the equivalent of grabbing the coin. So as training goes on and our model gets closer and closer to it's ideal performance (also called **convergence**), we want the amount it learns to be less and less.
#
# To do this we'll create an instance of the `ReduceLROnPlateau` callback to monitor the validation loss just like the `EarlyStopping` callback.
#
# Once the validation loss stops improving for two or more epochs, we'll reduce the learning rate by a factor of 5 (e.g. `0.001` to `0.0002`).
#
# And to make sure the learning rate doesn't get too low (and potentially result in our model learning nothing), we'll set the minimum learning rate to `1e-7`.
# + id="I794Kaiq4Ekk"
# Creating learning rate reduction callback
reduce_lr = tf.keras.callbacks.ReduceLROnPlateau(monitor="val_loss",
factor=0.2, # multiply the learning rate by 0.2 (reduce by 5x)
patience=2,
verbose=1, # print out when learning rate goes down
min_lr=1e-7)
# + [markdown] id="3v0BHURF6Lnz"
# Learning rate reduction ready to go!
#
# Now before we start training, we've got to recompile our model.
#
# We'll use sparse categorical crossentropy as the loss and since we're fine-tuning, we'll use a 10x lower learning rate than the Adam optimizers default (`1e-4` instead of `1e-3`).
# + id="GlpO9LflcVHW"
# Compile the model
loaded_gs_model.compile(loss="sparse_categorical_crossentropy", # sparse_categorical_crossentropy for labels that are *not* one-hot
optimizer=tf.keras.optimizers.Adam(0.0001), # 10x lower learning rate than the default
metrics=["accuracy"])
# + [markdown] id="yo1bio8SYxvc"
# Okay, model compiled.
#
# Now let's fit it on all of the data.
#
# We'll set it up to run for up to 100 epochs.
#
# Since we're going to be using the `EarlyStopping` callback, it might stop before reaching 100 epochs.
#
# > 🔑 **Note:** Running the cell below will set the model up to fine-tune all of the pre-trained weights in the base model on all of the Food101 data. Doing so with **unoptimized** data pipelines and **without** mixed precision training will take a fairly long time per epoch depending on what type of GPU you're using (about 15-20 minutes on Colab GPUs). But don't worry, **the code we've written above will ensure it runs much faster** (more like 4-5 minutes per epoch).
# + colab={"base_uri": "https://localhost:8080/"} id="LkUtOdVkbMPC" outputId="c316506e-0858-4b2e-81d7-c5921ef67368"
# Start to fine-tune (all layers)
history_101_food_classes_all_data_fine_tune = loaded_gs_model.fit(train_data,
epochs=100, # fine-tune for a maximum of 100 epochs
steps_per_epoch=len(train_data),
validation_data=test_data,
validation_steps=int(0.15 * len(test_data)), # validation during training on 15% of test data
callbacks=[create_tensorboard_callback("training_logs", "efficientb0_101_classes_all_data_fine_tuning"), # track the model training logs
model_checkpoint, # save only the best model during training
early_stopping, # stop model after X epochs of no improvements
reduce_lr]) # reduce the learning rate after X epochs of no improvements
# + [markdown] id="nC2HLXePh9Af"
# > 🔑 **Note:** If you didn't use mixed precision or use techniques such as [`prefetch()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#prefetch) in the *Batch & prepare datasets* section, your model fine-tuning probably takes up to 2.5-3x longer per epoch (see the output below for an example).
#
# | | Prefetch and mixed precision | No prefetch and no mixed precision |
# |-----|-----|-----|
# | Time per epoch | ~280-300s | ~1127-1397s |
#
# *Results from fine-tuning 🍔👁 Food Vision Big™ on Food101 dataset using an EfficienetNetB0 backbone using a Google Colab Tesla T4 GPU.*
#
# ```
# Saving TensorBoard log files to: training_logs/efficientB0_101_classes_all_data_fine_tuning/20200928-013008
# Epoch 1/100
# 2368/2368 [==============================] - 1397s 590ms/step - loss: 1.2068 - accuracy: 0.6820 - val_loss: 1.1623 - val_accuracy: 0.6894
# Epoch 2/100
# 2368/2368 [==============================] - 1193s 504ms/step - loss: 0.9459 - accuracy: 0.7444 - val_loss: 1.1549 - val_accuracy: 0.6872
# Epoch 3/100
# 2368/2368 [==============================] - 1143s 482ms/step - loss: 0.7848 - accuracy: 0.7838 - val_loss: 1.0402 - val_accuracy: 0.7142
# Epoch 4/100
# 2368/2368 [==============================] - 1127s 476ms/step - loss: 0.6599 - accuracy: 0.8149 - val_loss: 0.9599 - val_accuracy: 0.7373
# ```
# *Example fine-tuning time for non-prefetched data as well as non-mixed precision training (~2.5-3x longer per epoch).*
#
# Let's make sure we save our model before we start evaluating it.
# + id="mwjxnUsgI558"
# # Save model to Google Drive (optional)
# loaded_gs_model.save("/content/drive/MyDrive/tensorflow_course/food_vision/07_efficientnetb0_fine_tuned_101_classes_mixed_precision/")
# + colab={"base_uri": "https://localhost:8080/"} id="K1As0OhYHFX-" outputId="46408a84-943b-46f6-<PASSWORD>-2<PASSWORD>"
# Save model locally (note: if you're using Google Colab and you save your model locally, it will be deleted when your Google Colab session ends)
loaded_gs_model.save("07_efficientnetb0_fine_tuned_101_classes_mixed_precision")
# + [markdown] id="CcpNGcSAZ2UC"
# Looks like our model has gained a few performance points from fine-tuning, let's evaluate on the whole test dataset and see if managed to beat the [DeepFood paper's](https://arxiv.org/abs/1606.05675) result of 77.4% accuracy.
# + colab={"base_uri": "https://localhost:8080/"} id="2CR6q8MYM37K" outputId="d10180ce-1df4-4b77-a296-8f61ac7dd080"
# Evaluate mixed precision trained loaded model
results_loaded_gs_model_fine_tuned = loaded_gs_model.evaluate(test_data)
results_loaded_gs_model_fine_tuned
# + [markdown] id="rR-S5bKP0IxA"
# Woohoo!!!! It looks like our model beat the results mentioned in the DeepFood paper for Food101 (DeepFood's 77.4% top-1 accuracy versus our ~79% top-1 accuracy).
# + [markdown] id="a0H1rSG9RBBV"
# ## Download fine-tuned model from Google Storage
#
# As mentioned before, training models can take a significant amount of time.
#
# And again, like any good cooking show, here's something we prepared earlier...
#
# It's a fine-tuned model exactly like the one we trained above but it's saved to Google Storage so it can be accessed, imported and evaluated.
# + colab={"base_uri": "https://localhost:8080/"} id="qRuBSsPxI8Yg" outputId="86a5343d-d0eb-4fb4-9ed8-7acae14f7a9e"
# Download and evaluate fine-tuned model from Google Storage
# !wget https://storage.googleapis.com/ztm_tf_course/food_vision/07_efficientnetb0_fine_tuned_101_classes_mixed_precision.zip
# + [markdown] id="5_KhgOeA_hCG"
# The downloaded model comes in zip format (`.zip`) so we'll unzip it into the Google Colab instance.
# + colab={"base_uri": "https://localhost:8080/"} id="PNh0cPL7JBpv" outputId="c5ea36e4-1203-46eb-dd5a-285a8764d1e8"
# Unzip fine-tuned model
# !mkdir downloaded_fine_tuned_gs_model # create separate directory for fine-tuned model downloaded from Google Storage
# !unzip /content/07_efficientnetb0_fine_tuned_101_classes_mixed_precision -d downloaded_fine_tuned_gs_model
# + [markdown] id="blFdr0QJ_oQY"
# Now we can load it using the [`tf.keras.models.load_model()`](https://www.tensorflow.org/tutorials/keras/save_and_load) method and get a summary (it should be the exact same as the model we created above).
# + id="yBWWTb-QKuW1"
# Load in fine-tuned model from Google Storage and evaluate
loaded_fine_tuned_gs_model = tf.keras.models.load_model("/content/downloaded_fine_tuned_gs_model/07_efficientnetb0_fine_tuned_101_classes_mixed_precision")
# + colab={"base_uri": "https://localhost:8080/"} id="sysv2pDmJoe8" outputId="228088b7-0f87-4652-f894-3ef6accdb32a"
# Get a model summary (same model architecture as above)
loaded_fine_tuned_gs_model.summary()
# + [markdown] id="SxP8rbAj_4lY"
# Finally, we can evaluate our model on the test data (this requires the `test_data` variable to be loaded.
# + colab={"base_uri": "https://localhost:8080/"} id="Ms0R3LZ9Jrr5" outputId="957b540d-e05e-4f27-c379-39f7bce49da9"
# Note: Even if you're loading in the model from Google Storage, you will still need to load the test_data variable for this cell to work
results_downloaded_fine_tuned_gs_model = loaded_fine_tuned_gs_model.evaluate(test_data)
results_downloaded_fine_tuned_gs_model
# + [markdown] id="yMAXmidMc_hA"
# Excellent! Our saved model is performing as expected (better results than the DeepFood paper!).
#
# Congrautlations! You should be excited! You just trained a computer vision model with competitive performance to a research paper and in far less time (our model took ~20 minutes to train versus DeepFood's quoted 2-3 days).
#
# In other words, you brought Food Vision life!
#
# If you really wanted to step things up, you could try using the [`EfficientNetB4`](https://www.tensorflow.org/api_docs/python/tf/keras/applications/EfficientNetB4) model (a larger version of `EfficientNetB0`). At at the time of writing, the EfficientNet family has the [state of the art classification results](https://paperswithcode.com/sota/fine-grained-image-classification-on-food-101) on the Food101 dataset.
#
# > 📖 **Resource:** To see which models are currently performing the best on a given dataset or problem type as well as the latest trending machine learning research, be sure to check out [paperswithcode.com](http://paperswithcode.com/) and [sotabench.com](https://sotabench.com/).
# + [markdown] id="pFjooc5Gy0I2"
# ## View training results on TensorBoard
#
# Since we tracked our model's fine-tuning training logs using the `TensorBoard` callback, let's upload them and inspect them on TensorBoard.dev.
# + id="DW9x3o1kWlO3"
# !tensorboard dev upload --logdir ./training_logs \
# --name "Fine-tuning EfficientNetB0 on all Food101 Data" \
# --description "Training results for fine-tuning EfficientNetB0 on Food101 Data with learning rate 0.0001" \
# --one_shot
# + id="Cnqjq9aWM6pd"
View experiment: https://tensorboard.dev/experiment/2KINdYxgSgW2bUg7dIvevw/
# + [markdown] id="zvn5JfmuVSCf"
# Viewing at our [model's training curves on TensorBoard.dev](https://tensorboard.dev/experiment/2KINdYxgSgW2bUg7dIvevw/), it looks like our fine-tuning model gains boost in performance but starts to overfit as training goes on.
#
# See the training curves on TensorBoard.dev here: https://tensorboard.dev/experiment/2KINdYxgSgW2bUg7dIvevw/
#
# To fix this, in future experiments, we might try things like:
# * A different iteration of `EfficientNet` (e.g. `EfficientNetB4` instead of `EfficientNetB0`).
# * Unfreezing less layers of the base model and training them rather than unfreezing the whole base model in one go.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="fUpOtu_tNWcJ" outputId="1b9f45dd-95c6-4088-c939-761a491a3d3a"
# View past TensorBoard experiments
# !tensorboard dev list
# + colab={"base_uri": "https://localhost:8080/"} id="u-Rl-L0hNbEv" outputId="8b497600-ccd0-4f01-c4e5-defcb597310b"
# Delete past TensorBoard experiments
# # !tensorboard dev delete --experiment_id YOUR_EXPERIMENT_ID
# Example
# !tensorboard dev delete --experiment_id OAE6KXizQZKQxDiqI3cnUQ
# + [markdown] id="nDDrrqy5egTn"
# ## 🛠 Exercises
#
# 1. Use the same evaluation techniques on the large-scale Food Vision model as you did in the previous notebook ([Transfer Learning Part 3: Scaling up](https://github.com/mrdbourke/tensorflow-deep-learning/blob/main/06_transfer_learning_in_tensorflow_part_3_scaling_up.ipynb)). More specifically, it would be good to see:
# * A confusion matrix between all of the model's predictions and true labels.
# * A graph showing the f1-scores of each class.
# * A visualization of the model making predictions on various images and comparing the predictions to the ground truth.
# * For example, plot a sample image from the test dataset and have the title of the plot show the prediction, the prediction probability and the ground truth label.
# 2. Take 3 of your own photos of food and use the Food Vision model to make predictions on them. How does it go? Share your images/predictions with the other students.
# 3. Retrain the model (feature extraction and fine-tuning) we trained in this notebook, except this time use [`EfficientNetB4`](https://www.tensorflow.org/api_docs/python/tf/keras/applications/EfficientNetB4) as the base model instead of `EfficientNetB0`. Do you notice an improvement in performance? Does it take longer to train? Are there any tradeoffs to consider?
# 4. Name one important benefit of mixed precision training, how does this benefit take place?
# + [markdown] id="z8ncI458ZkpA"
# ## 📖 Extra-curriculum
#
# * Read up on learning rate scheduling and the [learning rate scheduler callback](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/LearningRateScheduler). What is it? And how might it be helpful to this project?
# * Read up on TensorFlow data loaders ([improving TensorFlow data loading performance](https://www.tensorflow.org/guide/data_performance)). Is there anything we've missed? What methods you keep in mind whenever loading data in TensorFlow? Hint: check the summary at the bottom of the page for a gret round up of ideas.
# * Read up on the documentation for [TensorFlow mixed precision training](https://www.tensorflow.org/guide/mixed_precision). What are the important things to keep in mind when using mixed precision training?
| 07_food_vision_milestone_project_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] cell_id="00000-7c9c2fd2-84b4-4d62-bcdc-a8da7f72feed" deepnote_app_coordinates={"x": 0, "y": null, "w": 12, "h": 5} deepnote_cell_type="markdown"
# # Lecture 1: Introduction to Data Science in Python
# ### [<NAME>](https://github.com/Qwerty71), [<NAME>](https://www.mci.sh), [<NAME>](https://www.vijayrs.ml)
# This notebook helps introduce some of the most basic tools that are commonly used for doing data science and statistics in Python.
#
# ## Note: you will need to run the following code cell every time you restart this notebook
# If this is your first time using Jupyter, click the block of code below and either press the Run button or press `Shift + Enter` on your keyboard.
# + cell_id="00001-d39b6262-0b35-4431-a304-1a15b4f84716" deepnote_to_be_reexecuted=false source_hash="2338102c" execution_start=1632835309692 execution_millis=13652 deepnote_app_coordinates={"x": 0, "y": 6, "w": 12, "h": 5} deepnote_cell_type="code"
# !pip install -r requirements.txt
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import statsmodels.api as sm
import seaborn as sns
from sklearn.linear_model import LinearRegression
from IPython.display import display
iris = sns.load_dataset('iris')
# + [markdown] tags=["jupyter-notebook"] cell_id="00002-28eb6661-b03e-451b-8727-5556ccecb17c" deepnote_app_coordinates={"x": 0, "y": 12, "w": 12, "h": 5} deepnote_cell_type="markdown"
# # Jupyter Notebook
# [Jupyter Notebook](https://jupyter.org/) is an interactive tool for running code and visualizing data. Each notebook consists of a series of _code cells_ and _Markdown cells_.
#
# * Code cells allow you to run code in a number of languages. Behind the scenes, Jupyter runs a "kernel" that processes the code whenever you execute a cell. Since this is a Python notebook, Jupyter is running the [IPython](https://ipython.org/) kernel. However, kernels also exist for Julia, R, and many other languages.
# * Markdown cells display text using the [Markdown language](https://jupyter-notebook.readthedocs.io/en/stable/examples/Notebook/Working%20With%20Markdown%20Cells.html). In addition to displaying text, you can write equations in these cells using $\LaTeX$.
#
# To run code, click a code cell (like the one below) and do one of the following:
# * Press `Shift + Enter` on your keyboard
# * On the toolbar next to the selected cell, press the Run button.
# + tags=["jupyter-notebook"] cell_id="00003-a0ab7299-d6b1-4716-b922-f0f35e4aac83" deepnote_to_be_reexecuted=false source_hash="a8a44a2d" execution_start=1632835323345 execution_millis=167 deepnote_app_coordinates={"x": 0, "y": 18, "w": 12, "h": 5} deepnote_cell_type="code"
print("Hello, world!")
# + [markdown] tags=["jupyter-notebook"] cell_id="00004-f4d71a30-33e8-4910-9c25-5bd726869771" deepnote_app_coordinates={"x": 0, "y": 24, "w": 12, "h": 5} deepnote_cell_type="markdown"
# You can render a markdown cell in the same way. Double click the text below, and try putting in some of the following items:
#
# # This is a large heading!
# ## This is a smaller heading!
# ### This is an even smaller heading!
# Here is some code: `x = y + z`
# And here is an equation: $x = y + z$
# + [markdown] tags=["jupyter-notebook"] cell_id="00005-26a7bfa9-c154-4089-bd26-014dcf001fe6" deepnote_app_coordinates={"x": 0, "y": 30, "w": 12, "h": 5} deepnote_cell_type="markdown"
#
# ## *Double-click this text!*
# + [markdown] cell_id="00006-f5ce4e43-6e02-4f8d-b6e4-c9b59f64c9be" deepnote_app_coordinates={"x": 0, "y": 36, "w": 12, "h": 5} deepnote_cell_type="markdown"
# ### Cell magic
# The IPython kernel provides some useful tools for programmers, including
#
# * [Magic commands](https://ipython.readthedocs.io/en/stable/interactive/magics.html), which allow you to do things like look up documentation and past commands that you've run, and
# * [Building graphical user interfaces (GUIs)](https://ipython.org/ipython-doc/stable/interactive/reference.html#gui-event-loop-support) to make it easier to interact with your code.
#
# Here's an example of a useful magic command. `?` will look up the documentation for a library, class, or function to help you figure out how to use it. For instance, if I want to learn about [pandas DataFrames](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.html), I can run the following:
# + cell_id="00007-7fab3cbf-546b-421a-97f3-d0d6a3b02ef8" deepnote_to_be_reexecuted=false source_hash="413901dd" execution_start=1632835323374 execution_millis=13 deepnote_app_coordinates={"x": 0, "y": 42, "w": 12, "h": 5} deepnote_cell_type="code"
# ?pd.DataFrame
# + [markdown] cell_id="00008-94b5dc0a-6abd-4151-a340-4022ea30ce93" deepnote_app_coordinates={"x": 0, "y": 48, "w": 12, "h": 5} deepnote_cell_type="markdown"
# If you want to see all the magic functions that IPython makes available to you, `%quickref` can give you a high-level overview.
# + cell_id="00009-7269edd3-eee6-4b55-9435-2b70a14cca02" deepnote_to_be_reexecuted=false source_hash="33232240" execution_start=1632835327290 execution_millis=1 deepnote_app_coordinates={"x": 0, "y": 54, "w": 12, "h": 5} deepnote_cell_type="code"
# %quickref
# + [markdown] tags=["jupyter-notebook"] cell_id="00010-a5ec21b5-6738-410e-a6bf-80fc6c2d222b" deepnote_app_coordinates={"x": 0, "y": 60, "w": 12, "h": 5} deepnote_cell_type="markdown"
# ### Widgets
# IPython and Jupyter Notebook also makes it easy to build [widgets](https://ipywidgets.readthedocs.io/en/latest/index.html), which give you a richer interface with which to interact with the notebook. Try running the code cell below. This code creates two plots, and displays them in adjacent tabs.
# + tags=["jupyter-notebook"] cell_id="00011-a3678e0e-57c0-43e0-ab80-2242a02aa276" deepnote_to_be_reexecuted=false source_hash="d3882c2c" execution_start=1632835344994 execution_millis=740 deepnote_app_coordinates={"x": 0, "y": 66, "w": 12, "h": 5} deepnote_cell_type="code"
# %matplotlib inline
import matplotlib.pyplot as plt
import ipywidgets as widgets
from scipy.stats import norm, linregress
out = [widgets.Output(), widgets.Output()]
tabs = widgets.Tab(children=[out[0], out[1]])
tabs.set_title(0, 'Linear regression')
tabs.set_title(1, 'Normal distribution')
with out[0]:
# Fit line to some random data
x = np.random.uniform(size=30)
y = x + np.random.normal(scale=0.1, size=30)
slope, intercept, _, _, _ = linregress(x,y)
u = np.linspace(0, 1)
# Plot
fig1, axes1 = plt.subplots()
axes1.scatter(x, y)
axes1.plot(u, slope * u + intercept, 'k')
plt.show(fig1)
with out[1]:
# Plot the probability distribution function (pdf) of the
# standard normal distribution.
x = np.linspace(-3.5, 3.5, num=100)
p = norm.pdf(x)
# Plot
fig2, axes2 = plt.subplots()
axes2.plot(x, p)
plt.show(fig2)
display(tabs)
# + [markdown] tags=["jupyter-notebook"] cell_id="00012-a477b6eb-7ab3-4ba4-bfcf-5bd0705b5deb" deepnote_app_coordinates={"x": 0, "y": 72, "w": 12, "h": 5} deepnote_cell_type="markdown"
# You can create much richer and more complex interfaces that include buttons, sliders, progress bars, and more with Jupyter's ipywidgets library ([docs](https://ipywidgets.readthedocs.io/en/latest/index.html)).
# + [markdown] tags=["pandas"] cell_id="00013-d3bc29ff-e3ce-46bb-a355-9057bddfc0bd" deepnote_app_coordinates={"x": 0, "y": 78, "w": 12, "h": 5} deepnote_cell_type="markdown"
# # Pandas
# [pandas](https://pandas.pydata.org/) is a Python library that provides useful data structures and tools for analyzing data.
#
# The fundamental type of the pandas library is the `DataFrame`. In the following code, we load the [iris flower dataset](https://en.wikipedia.org/wiki/Iris_flower_data_set) using the [seaborn library](https://seaborn.pydata.org/). By default, this dataset is stored in a pandas `DataFrame`.
# + tags=["pandas"] cell_id="00014-126a402e-911f-4ac3-93b5-825300819892" deepnote_to_be_reexecuted=false source_hash="ad6cb791" execution_start=1632835365980 execution_millis=88 deepnote_app_coordinates={"x": 0, "y": 84, "w": 12, "h": 5} deepnote_cell_type="code"
import pandas as pd
import seaborn as sns
iris = sns.load_dataset('iris')
# `iris` is stored as a pandas DataFrame
print('Type of "iris":', type(iris))
# Show the first few entries in this DataFrame
iris.head()
# + [markdown] tags=["pandas"] cell_id="00015-93accaa8-7129-40cc-87f5-38b0fe3539d2" deepnote_app_coordinates={"x": 0, "y": 90, "w": 12, "h": 5} deepnote_cell_type="markdown"
# Let's get some information about the iris dataset. Let's try to do the following:
#
# 1. Find out how many columns there are in the `DataFrame` object, and what kinds of data are in each column
# 2. Calculate the average petal length
# 3. Determine what species of flowers are in the dataset
# 4. Get an overall summary of the dataset
# + tags=["pandas"] cell_id="00016-1c3821b5-67f9-4712-a2cb-b660bde1a2b6" deepnote_to_be_reexecuted=false source_hash="67cf228" execution_start=1632835424142 execution_millis=18 deepnote_app_coordinates={"x": 0, "y": 96, "w": 12, "h": 5} deepnote_cell_type="code"
# 1. Column labels, and types of data in each column
print(iris.dtypes)
# + tags=["pandas"] cell_id="00017-232c7954-707e-4cdc-959f-d0a1629d94b1" deepnote_to_be_reexecuted=false source_hash="650ade12" execution_start=1632835435328 execution_millis=19 deepnote_app_coordinates={"x": 0, "y": 102, "w": 12, "h": 5} deepnote_cell_type="code"
# 2. Calculate the average petal length
print(iris['petal_length'].mean())
# + tags=["pandas"] cell_id="00018-21fc5a51-87a8-4870-b324-70139684a764" deepnote_to_be_reexecuted=false source_hash="56b279a5" execution_start=1632835438486 execution_millis=23 deepnote_app_coordinates={"x": 0, "y": 108, "w": 12, "h": 5} is_output_hidden=false deepnote_cell_type="code"
# 3. Determine which iris species are in the dataset
print(iris['species'].unique())
# + tags=["pandas"] cell_id="00019-9dbbd01a-da95-4d11-a626-76235a9e19db" deepnote_to_be_reexecuted=false source_hash="50ef09a6" execution_start=1632835441153 execution_millis=1048 deepnote_app_coordinates={"x": 0, "y": 114, "w": 12, "h": 5} deepnote_cell_type="code"
# 4. Summary of the data
iris.describe()
# + [markdown] cell_id="00020-6d5cffa0-7bca-482a-90e6-25b5f4577331" deepnote_app_coordinates={"x": 0, "y": 120, "w": 12, "h": 5} deepnote_cell_type="markdown"
# Sometimes we need to extract certain rows or columns of a DataFrame. For instance, in the following code we store each species of flower in its own variable:
# + tags=["pandas"] cell_id="00021-104446fe-73db-471a-b350-3b1b6f629a3c" deepnote_to_be_reexecuted=false source_hash="3b6d5c73" execution_start=1632829859729 execution_millis=88 deepnote_app_coordinates={"x": 0, "y": 126, "w": 12, "h": 5} deepnote_cell_type="code"
"""
IPython.display is a convenience function that works in Jupyter Notebook
(or, more generally, any IPython-based application) that will show
objects in a nicer way than using print(). We'll use it in this notebook
to show some pandas DataFrames.
"""
from IPython.display import display
"""
Create a DataFrame for each species of flower. I've provided two
methods for creating these DataFrames below; pick whichever you
prefer as they are equivalent.
"""
# Method 1: "query" function
setosa = iris.query('species == "setosa"')
versicolor = iris.query('species == "versicolor"')
# Method 2: index into the DataFrame
virginica = iris[iris['species'] == 'virginica']
"""
Show the first few entries of the DataFrame corresponding to each species
"""
print('Setosa data:')
display(setosa.head())
print('Versicolor data:')
display(versicolor.head())
print('Virginica data:')
display(virginica.head())
# + [markdown] cell_id="00022-92e41837-6b26-400d-8a20-25ab955034e8" deepnote_app_coordinates={"x": 0, "y": 132, "w": 12, "h": 5} deepnote_cell_type="markdown"
# To extract a column, we can either use `iris[column_name]` or `iris.iloc[:,column_index]`.
# + cell_id="00023-2e34c54f-5402-4371-804d-ee8f3f0c9f0b" deepnote_to_be_reexecuted=false source_hash="a15173b8" execution_start=1632835497714 execution_millis=120 deepnote_app_coordinates={"x": 0, "y": 138, "w": 12, "h": 5} deepnote_cell_type="code"
"""
Get the first column.
Note: whenever we extract a single column of a pandas DataFrame,
we get back a pandas Series object. To turn it back into a DataFrame,
we add the line `first_column = pd.DataFrame(first_column)`.
"""
first_column = iris.iloc[:,0]
first_column = pd.DataFrame(first_column)
print('First column:')
display(first_column.head())
"""
Get the first through third columns
"""
first_through_third_columns = iris.iloc[:,0:3]
print('First through third columns:')
display(first_through_third_columns.head())
"""
Get the 'species' column.
"""
species = iris['species']
species = pd.DataFrame(species)
print('Species column:')
display(species.head())
"""
Get all columns *except* the species column
"""
all_but_species = iris.iloc[:, iris.columns != 'species']
print("All columns *except* species:")
display(all_but_species.head())
# + [markdown] cell_id="00024-0f2cfc98-adf1-48cc-aa22-3db3889ea752" deepnote_app_coordinates={"x": 0, "y": 144, "w": 12, "h": 5} deepnote_cell_type="markdown"
# If you want to create your own pandas `DataFrame`, you have to specify the names of the columns and the items in the rows of the `DataFrame`.
# + cell_id="00025-56bca6ed-8a28-4b13-a39c-28a0bd74ab87" deepnote_to_be_reexecuted=false source_hash="f515e99d" execution_start=1632835634374 execution_millis=14 deepnote_app_coordinates={"x": 0, "y": 150, "w": 12, "h": 5} deepnote_cell_type="code"
column_labels = ['A', 'B']
column_entries = [
[1, 2],
[4, 5],
[7, 8]
]
pd.DataFrame(column_entries, columns=column_labels)
# + [markdown] cell_id="00026-f77e3587-d3bc-4da3-b095-ce64a0947f90" deepnote_app_coordinates={"x": 0, "y": 156, "w": 12, "h": 5} deepnote_cell_type="markdown"
# # NumPy
# [NumPy](https://www.numpy.org/) is another Python package providing useful data structures and mathematical functions. NumPy's fundamental data type is the array, `numpy.ndarray`, which is like a stripped-down version of a pandas `DataFrame`. However, the `numpy.ndarray` supports much faster operations, which makes it a lot more practical for scientific computing than, say, Python's list objects.
# + cell_id="00027-b1cb97d6-1190-4034-9ebe-a786b58e29a4" deepnote_to_be_reexecuted=false source_hash="9d29225b" execution_start=1632835740446 execution_millis=1 deepnote_app_coordinates={"x": 0, "y": 162, "w": 12, "h": 5} deepnote_cell_type="code"
import numpy as np
# 1. Create an array with the numbers [1, 2, 3]
x = np.array([1, 2, 3])
# 2. Create a 2 x 2 matrix with [1, 2] in the first row and [3, 4]
# in the second row.
x = np.array( [[1,2], [3,4]] )
# 3. Create an array with the numbers 0, 1, ... , 9. Equivalent to
# calling np.array(range(10))
x = np.arange(10)
# 4. Create a 2 x 2 matrix with zeros in all entries
x = np.zeros( (2,2) )
# 5. Get the total number of items in the matrix, and the shape of
# the matrix.
num_items = x.size
matrix_shape = x.shape
# + [markdown] cell_id="00028-6b5693fa-7cea-4f14-870e-73bb0074972b" deepnote_app_coordinates={"x": 0, "y": 168, "w": 12, "h": 5} deepnote_cell_type="markdown"
# Besides just providing data structures, though, NumPy provides many mathematical utilities as well.
# + cell_id="00029-08be4aa5-e868-47c0-bc7c-64eeecdf6b7a" deepnote_to_be_reexecuted=false source_hash="33ed6c7" execution_start=1632835746019 execution_millis=30 deepnote_app_coordinates={"x": 0, "y": 174, "w": 12, "h": 5} deepnote_cell_type="code"
### Constants: pi
print('π = %f' % np.pi)
print()
### Simple functions: sine, cosine, e^x, log, ...
print('sin(0) = %f' % np.sin(0))
print('cos(0) = %f' % np.cos(0))
print('e^1 = %f' % np.exp(1))
print('ln(1) = %f' % np.log(1))
print()
### Minimums, maximums, sums...
x = np.array([1,2,3])
print('Min of [1,2,3] = %d' % x.min())
print('Max of [1,2,3] = %d' % x.max())
print('Sum of [1,2,3] = %d' % x.sum())
print()
### Random numbers: uniform distribution, normal distribution, ...
print('Random numbers:')
print('Uniform([0,1]): %f' % np.random.uniform(0,1))
print('Normal(0,1): %f' % np.random.normal(loc=0, scale=1))
print('Poisson(1): %f' % np.random.poisson(1))
# + [markdown] cell_id="00030-f6bbe0e8-23f1-4cda-a491-7315fbfdbd5c" deepnote_app_coordinates={"x": 0, "y": 180, "w": 12, "h": 5} deepnote_cell_type="markdown"
# NumPy is primarily used to do large-scale operations on arrays of numbers. Because it has C code running behind the scenes, it can do these computations extremely quickly -- much faster than you could do with regular Python code. Among other things, with NumPy you can
#
# * add a number to every element of an array;
# * multiply every element of an array by a number;
# * add or multiply two arrays together; or
# * calculate a matrix-vector or matrix-matrix product between arrays.
# + cell_id="00031-1feba1ce-f7c7-487e-be7c-6365efb1ac4c" deepnote_to_be_reexecuted=false source_hash="e76a617a" execution_start=1632835752392 execution_millis=26 deepnote_app_coordinates={"x": 0, "y": 186, "w": 12, "h": 5} deepnote_cell_type="code"
x = np.array([1,2,3])
y = np.array([4,5,6])
print('1 + [1,2,3] =', 1 + x)
print('3 * [1,2,3] =', 3 * x)
print('[1,2,3] * [4,5,6] =', x * y)
print('[1,2,3] + [4,5,6] =', x + y)
print('Dot product of [1,2,3] and [4,5,6] =', x.dot(y))
# + [markdown] cell_id="00032-9dc9b5ea-ac56-4ea8-8a13-77996856ba2e" deepnote_app_coordinates={"x": 0, "y": 192, "w": 12, "h": 5} deepnote_cell_type="markdown"
# # Linear regression with scikit-learn and statsmodels
# In the last section of the notebook, we're going to use linear regression to try and predict the petal length of each iris from its sepal length, sepal width, and petal width.
#
# Before we even start using linear regression, let's look at how each of these variables are related to one another. Below we plot each pair of variables against the others, with the color of the points reflecting which species we're looking at. On the diagonal are box-and-whisker plots that tell us a little about how each of the variables is distributed.
# + cell_id="00033-6012eddb-48bc-4309-b79d-b81c6030c21b" deepnote_to_be_reexecuted=false source_hash="229e9196" execution_start=1632835776033 execution_millis=6163 deepnote_app_coordinates={"x": 0, "y": 198, "w": 12, "h": 5} deepnote_cell_type="code"
import seaborn as sns
sns.set()
sns.pairplot(iris, hue="species")
# + [markdown] cell_id="00034-e932e048-3cac-4c03-b422-d010457bc254" deepnote_app_coordinates={"x": 0, "y": 204, "w": 12, "h": 5} deepnote_cell_type="markdown"
# The most important conclusion we can draw from these plots is that each pair of variables is roughly linearly related. As a result, we can expect that we should be able to accurately predict petal length from the other three variables using linear regression.
#
# If you just want to do a basic regression and get the coefficients for each variable, you can use `LinearRegression` from the scikit-learn library:
# + cell_id="00035-adf77a24-4064-4ed3-bcf6-32fbfa4c120f" deepnote_to_be_reexecuted=false source_hash="528b4bca" execution_start=1632830090191 execution_millis=17 deepnote_app_coordinates={"x": 0, "y": 210, "w": 12, "h": 5} deepnote_cell_type="code"
from sklearn.linear_model import LinearRegression
"""
Get all of the irises of the species "setosa" and place them in
a pandas DataFrame called `data`.
Also try with 'setosa' replaced by 'versicolor' and 'virginica'
"""
data = iris[iris['species'] == 'setosa']
"""
Split the data into two pieces: the independent variables
(sepal_length, sepal_width, and petal_width), and the dependent
variable (petal_length).
"""
x = data[['sepal_length','sepal_width','petal_width']]
y = data['petal_length']
"""
Create a scikit-learn LinearRegression object which we will
fit to the data.
"""
lm = LinearRegression()
lm.fit(x,y)
for (coef,col) in zip(lm.coef_, x.columns):
print("%-30s %+.3f" % ("Coefficient for " + col + ":", coef))
# + [markdown] cell_id="00036-0b9f7011-d4df-49e5-a2d8-d4a924909431" deepnote_app_coordinates={"x": 0, "y": 216, "w": 12, "h": 5} deepnote_cell_type="markdown"
# However, when we run a linear regression we also want to find out other things about our linear model. For instance, we might want to get a confidence interval for each coefficient in the model. The [StatsModels library](https://www.statsmodels.org/stable/index.html) gives us this functionality with `statsmodel.api.sm`:
# + cell_id="00037-a2be049a-ab9a-48a3-90d0-3122be05c96a" deepnote_to_be_reexecuted=false source_hash="4460e923" execution_start=1632830101881 execution_millis=262 deepnote_app_coordinates={"x": 0, "y": 222, "w": 12, "h": 5} is_code_hidden=false is_output_hidden=false deepnote_cell_type="code"
import statsmodels.api as sm
# Also try with 'setosa' replaced by 'versicolor' and 'virginica'
data = iris[iris['species'] == 'setosa']
x = data[['sepal_length','sepal_width','petal_width']]
y = data['petal_length']
"""
Add y-intercept term to our linear model
"""
x = sm.add_constant(x)
"""
Perform linear regression
"""
lm = sm.OLS(y,x)
results = lm.fit()
"""
Print the results
"""
print(results.summary())
# + [markdown] cell_id="00038-7325b4de-90eb-4658-abce-109416185f74" deepnote_app_coordinates={"x": 0, "y": 228, "w": 12, "h": 5} deepnote_cell_type="markdown"
# You can also use R-style formulas to specify what variables you want to use to perform linear regression.
# + cell_id="00039-4a0a9d28-4593-4063-ba58-47fd27df6062" deepnote_to_be_reexecuted=false source_hash="8ce469a7" execution_start=1632800689229 execution_millis=34 deepnote_app_coordinates={"x": 0, "y": 234, "w": 12, "h": 5} is_output_hidden=false is_code_hidden=false deepnote_cell_type="code"
import statsmodels.formula.api as smf
df = iris[iris['species'] == 'setosa']
# Predict petal length from sepal_width, sepal_length, and petal_width
model = smf.ols(formula='petal_length ~ sepal_width + sepal_length + petal_width',
data=df)
results = model.fit()
print("Results (petal_length ~ sepal_width + sepal_length + petal_width)")
print(results.summary())
print('-' * 80)
# Predict petal length from just sepal_length and petal_width
model = smf.ols(formula='petal_length ~ sepal_length + petal_width',
data=df)
results = model.fit()
print("Results (petal_length ~ sepal_length + petal_width)")
print(results.summary())
# + [markdown] tags=[] is_collapsed=false cell_id="00040-c7b23cb8-6336-47a3-9648-c0762ab57535" deepnote_cell_type="text-cell-h1"
# # Try it yourself
# + [markdown] tags=[] cell_id="00041-f5b18891-96f7-4d67-bc5a-333da9646462" deepnote_cell_type="markdown"
# Use `sns.load_dataset('penguins')` to load the [palmerpenguins dataset](https://allisonhorst.github.io/palmerpenguins/) into a [pandas](https://pandas.pydata.org) `DataFrame` with the [seaborn library](https://seaborn.pydata.org/). Use the `.head()` method to display the first 5 rows of the `DataFrame`.
# + tags=[] cell_id="00041-abcec43c-21fd-4de5-84d8-d8d02b92ddcc" deepnote_to_be_reexecuted=false source_hash="7cd5ae39" execution_start=1632778645713 execution_millis=9 deepnote_cell_type="code"
import pandas as pd
import seaborn as sns
penguins = sns.load_dataset('penguins')
# Show the first few entries in this DataFrame
# YOUR CODE HERE
# + [markdown] tags=[] cell_id="00043-75599903-d8e5-46b1-a015-92b91c59ff8c" deepnote_cell_type="markdown"
# Looking at row 3, we can see that there are `nan` values in the dataset. These values will cause problems will calculations down the line so you should use the `.dropna()` method to remove them now.
# + tags=[] cell_id="00044-deeecef7-7eca-4c7a-991f-b2cf2f487c93" deepnote_to_be_reexecuted=false source_hash="8e7c4dfd" execution_start=1632778645722 execution_millis=1 deepnote_cell_type="code"
# Removing rows with 'nan' values
# YOUR CODE HERE
# + [markdown] tags=[] cell_id="00043-a7f81fa6-0e0a-4e8b-a763-ba9d95fd4982" deepnote_cell_type="markdown"
# Let's get some information about the penguins dataset. Let's try to do the following:
#
# 1. Find out how many columns there are in the `DataFrame` object, and what kinds of data are in each column
# 2. Calculate the average bill length
# 3. Determine what penguin species are in the dataset
# 4. Get an overall summary of the dataset
# + tags=[] cell_id="00044-f0f1d714-e3a5-4bf2-bd75-b34ef3fb21b6" deepnote_to_be_reexecuted=false source_hash="5bf2d5ad" execution_start=1632778645724 execution_millis=0 deepnote_cell_type="code"
# 1. Column labels, and types of data in each column
# YOUR CODE HERE
# + tags=[] cell_id="00045-dc194b98-693a-4f86-8940-1a2c1ffd2b14" deepnote_to_be_reexecuted=false source_hash="a663e912" execution_start=1632778645740 execution_millis=0 deepnote_cell_type="code"
# 2. Calculate the average bill length
# YOUR CODE HERE
# + tags=[] cell_id="00046-1aed981f-b682-494b-8a90-214c2ca9f5c2" deepnote_to_be_reexecuted=false source_hash="33099e6c" execution_start=1632778645741 execution_millis=0 deepnote_cell_type="code"
# 3. Determine which penguin species are in the dataset
# YOUR CODE HERE
# + tags=[] cell_id="00047-144302dd-cef6-4d57-b1e9-5b44d5904a5b" deepnote_to_be_reexecuted=false source_hash="848d87c3" execution_start=1632778645741 execution_millis=3 deepnote_cell_type="code"
# 4. Summary of the data
# YOUR CODE HERE
# + [markdown] tags=[] cell_id="00048-10b74706-30a0-4d60-92bc-dd8705fb378b" deepnote_cell_type="markdown"
# Store a `DataFrame` for each species in its own variable using either the `.query()` function or by indexing into the `DataFrame`. Use the `IPython.display` module to render the first five lines of each `DataFrame`.
# + tags=[] cell_id="00049-da75b50f-4060-4f66-a25a-5925cef6495e" deepnote_to_be_reexecuted=false source_hash="eca38295" execution_start=1632778645744 execution_millis=0 deepnote_cell_type="code"
from IPython.display import display
# Method 1: "query" function
# YOUR CODE HERE
# Method 2: index into the DataFrame
# YOUR CODE HERE
# Show the first few entries of the DataFrame corresponding to each species
# YOUR CODE HERE
# + [markdown] tags=[] cell_id="00050-db775712-1c86-4ad5-9cbd-a6032199c78e" deepnote_cell_type="markdown"
# Use either the `.iloc[:,column_index]` method or index into the dataframe using `column_name` to extract the following columns into their own `DataFrame`:
# 1. First Column
# 2. First Through Third Columns
# 3. `Sex` Column
# 4. Every Column Except The `Sex` Column
# + tags=[] cell_id="00051-85acc6de-a972-409c-84a4-c730a9937ae0" deepnote_to_be_reexecuted=false source_hash="4505bff9" execution_start=1632778645746 execution_millis=1 deepnote_cell_type="code"
# Get the first column
# YOUR CODE HERE
# Get the first through third columns
# YOUR CODE HERE
# Get the 'sex' column
# YOUR CODE HERE
# Get all columns *except* the 'sex' column
# YOUR CODE HERE
# + [markdown] tags=[] cell_id="00052-030c4e66-9050-4850-81c9-a65170650ec2" deepnote_cell_type="markdown"
# Use `sns.pairplot()` function to see how the variables in the `DataFrame` are related to each other. Pass `hue="species"` as an argument to function in order to distinguish between the penguin species.
# + tags=[] cell_id="00053-3e949723-3379-4412-803f-c44929c60b2c" deepnote_to_be_reexecuted=false source_hash="ac6aaae5" execution_start=1632778645753 execution_millis=2 deepnote_cell_type="code"
import seaborn as sns
sns.set()
# YOUR CODE HERE
# + [markdown] tags=[] cell_id="00056-d34161ab-0c2f-4eb8-b130-c51748c192af" deepnote_cell_type="markdown"
# Use the `LinearRegression` object from the `sklearn.linear_model` library to predict flipper length based on bill length, bill depth, and body mass for Adelie penguins. Print the coefficients for each of the dependent variables.
# + tags=[] cell_id="00054-d701f984-317c-496d-85e3-6180b5d98dec" deepnote_to_be_reexecuted=false source_hash="b6fa5d7" execution_start=1632835899139 execution_millis=1450 output_cleared=true deepnote_cell_type="code"
from sklearn.linear_model import LinearRegression
# Get all Adelie penguins and put them in a DataFrame called `data`.
# YOUR CODE HERE
"""
Split the data into two sets: the independent variables
(bill_length_mm, bill_depth_mm, body_mass_g), and
the dependent variable (flipper_length_mm)
"""
# YOUR CODE HERE
"""
Create a scikit-learn LinearRegression object which we will
fit to the data.
"""
# YOUR CODE HERE
# Print out the coefficients for each dependent variable
for (coef,col) in zip(lm.coef_, x.columns):
print("%-30s %+.3f" % ("Coefficient for " + col + ":", coef))
# + [markdown] tags=[] cell_id="00058-e888fcd2-9de6-4461-aed4-3e943556bc7a" deepnote_cell_type="markdown"
# Perform the same regression with `statsmodel.api.sm` from the [StatsModels library](https://www.statsmodels.org/stable/index.html) in order to get more information about the model.
# + tags=[] cell_id="00057-90cdb391-3d79-4886-a203-08afd55f6dec" deepnote_to_be_reexecuted=false source_hash="6d1b53eb" execution_start=1632778645791 execution_millis=23 is_code_hidden=false deepnote_cell_type="code"
import statsmodels.api as sm
# Get all Adelie penguins and put them in a DataFrame called `data`.
# YOUR CODE HERE
"""
Split the data into two sets: the independent variables
(bill_length_mm, bill_depth_mm, body_mass_g), and
the dependent variable (flipper_length_mm)
"""
# YOUR CODE HERE
"""
Add y-intercept term to our linear model
"""
# YOUR CODE HERE
"""
Perform linear regression
"""
# YOUR CODE HERE
"""
Print the results
"""
# YOUR CODE HERE
# + [markdown] tags=[] is_collapsed=false cell_id="00060-5aa977ff-4765-4ab4-91ed-2e51b093ae4e" deepnote_cell_type="text-cell-h1"
# # Solutions
# + [markdown] tags=[] cell_id="00061-a9239808-4a50-4046-a525-8b0619c3164b" deepnote_cell_type="markdown"
# Use `sns.load_dataset('penguins')` to load the [palmerpenguins dataset](https://allisonhorst.github.io/palmerpenguins/) into a [pandas](https://pandas.pydata.org) `DataFrame` with the [seaborn library](https://seaborn.pydata.org/). Use the `.head()` method to display the first 5 rows of the `DataFrame`.
# + tags=[] cell_id="00062-6deef299-85a6-4a2f-8a5b-27923e659ece" deepnote_to_be_reexecuted=false source_hash="a0c60d24" execution_start=1632778645792 execution_millis=15 is_code_hidden=true deepnote_cell_type="code"
import pandas as pd
import seaborn as sns
penguins = sns.load_dataset('penguins')
# `penguins` is stored as a pandas DataFrame
print('Type of "penguins":', type(penguins))
# Show the first few entries in this DataFrame
penguins.head()
# + [markdown] tags=[] cell_id="00063-15bf382a-a28e-4085-aa48-55fcb94d9210" deepnote_cell_type="markdown"
# Looking at row 3, we can see that there are `nan` values in the dataset. These values will cause problems will calculations down the line so you should use the `.dropna()` method to remove them now.
# + tags=[] cell_id="00064-1b65a1ca-c576-4494-92ae-df9e961d23b5" deepnote_to_be_reexecuted=false source_hash="8a26bc1f" execution_start=1632778645828 execution_millis=17 is_code_hidden=true deepnote_cell_type="code"
# Removing rows with 'nan' values
penguins = penguins.dropna()
display(penguins.head())
# + [markdown] tags=[] cell_id="00065-c365772b-d657-4ce4-8306-3f7682988e7e" deepnote_cell_type="markdown"
# Let's get some information about the penguins dataset. Let's try to do the following:
#
# 1. Find out how many columns there are in the `DataFrame` object, and what kinds of data are in each column
# 2. Calculate the average bill length
# 3. Determine what penguin species are in the dataset
# 4. Get an overall summary of the dataset
# + tags=[] cell_id="00066-cc291298-40e1-498b-b9b0-4c0ce1407df8" deepnote_to_be_reexecuted=false source_hash="d85aafac" execution_start=1632778645870 execution_millis=20 is_code_hidden=true deepnote_cell_type="code"
# 1. Column labels, and types of data in each column
penguins.dtypes
# + tags=[] cell_id="00067-eb335307-cc60-455e-9873-1751157e9dfb" deepnote_to_be_reexecuted=false source_hash="62ee4b79" execution_start=1632778645871 execution_millis=19 is_code_hidden=true deepnote_cell_type="code"
# 2. Calculate the average bill length
penguins['bill_length_mm'].mean()
# + tags=[] cell_id="00068-a8442092-e969-4e28-890b-a415649057f4" deepnote_to_be_reexecuted=false source_hash="ebe79347" execution_start=1632778645871 execution_millis=19 is_code_hidden=true deepnote_cell_type="code"
# 3. Determine which penguin species are in the dataset
penguins['species'].unique()
# + tags=[] cell_id="00069-59cf48db-e56b-4a5a-9b5c-6d925fad0606" deepnote_to_be_reexecuted=false source_hash="398fc003" execution_start=1632778645882 execution_millis=46 is_code_hidden=true deepnote_cell_type="code"
# 4. Summary of the data
penguins.describe()
# + [markdown] tags=[] cell_id="00070-510fefb5-3dd5-4b0a-b810-edfbaa9d037e" deepnote_cell_type="markdown"
# Store a `DataFrame` for each species in its own variable using either the `.query()` function or by indexing into the `DataFrame`. Use the `IPython.display` module to render the first five lines of each `DataFrame`.
# + tags=[] cell_id="00071-62a2d168-4f1e-455d-9bb8-1c2b38ee1997" deepnote_to_be_reexecuted=false source_hash="8f91bb57" execution_start=1632778645920 execution_millis=93 is_code_hidden=true deepnote_cell_type="code"
from IPython.display import display
# Method 1: "query" function
adelie = penguins.query('species == "Adelie"')
chinstrap = penguins.query('species == "Chinstrap"')
# Method 2: index into the DataFrame
gentoo = penguins[penguins['species'] == 'Gentoo']
# Show the first few entries of the DataFrame corresponding to each species
print('Adelie data:')
display(adelie.head())
print('Chinstrap data:')
display(chinstrap.head())
print('Gentoo data:')
display(gentoo.head())
# + [markdown] tags=[] cell_id="00072-e816a728-ab53-4e18-b711-daace6d40cf2" deepnote_cell_type="markdown"
# Use either the `.iloc[:,column_index]` method or index into the dataframe using `column_name` to extract the following columns into their own `DataFrame`:
# 1. First Column
# 2. First Through Third Columns
# 3. `Sex` Column
# 4. Every Column Except The `Sex` Column
# + tags=[] cell_id="00073-8e103dd4-9424-4304-956c-bf78bf20a454" deepnote_to_be_reexecuted=false source_hash="54c93fb4" execution_start=1632778646007 execution_millis=79 is_code_hidden=true deepnote_cell_type="code"
# Get the first column
first_column = penguins.iloc[:,0]
first_column = pd.DataFrame(first_column)
print('First column:')
display(first_column.head())
# Get the first through third columns
first_through_third_columns = penguins.iloc[:,0:3]
print('First through third columns:')
display(first_through_third_columns.head())
# Get the 'sex' column
sex = penguins['sex']
sex = pd.DataFrame(sex)
print('Sex column:')
display(sex.head())
# Get all columns *except* the 'sex' column
exclude_sex = penguins.iloc[:, penguins.columns != 'sex']
print('All columns *except* sex:')
display(exclude_sex.head())
# + [markdown] tags=[] cell_id="00074-ca18eeef-b70c-48ec-a0e4-031cb3155986" deepnote_cell_type="markdown"
# Use `sns.pairplot()` function to see how the variables in the `DataFrame` are related to each other. Pass `hue="species"` as an argument to function in order to distinguish between the penguin species.
# + tags=[] cell_id="00075-25f4df4b-cc54-4245-ac2f-58f3f8518ac9" deepnote_to_be_reexecuted=false source_hash="fe470c4a" execution_start=1632778646247 execution_millis=4549 is_code_hidden=true deepnote_cell_type="code"
import seaborn as sns
sns.set()
sns.pairplot(penguins, hue="species")
# + [markdown] tags=[] cell_id="00076-a33bf63e-9904-4477-beb1-af1338404ce4" deepnote_cell_type="markdown"
# Use the `LinearRegression` object from the `sklearn.linear_model` library to predict flipper length based on bill length, bill depth, and body mass for Adelie penguins. Print the coefficients for each of the dependent variables.
# + tags=[] cell_id="00077-cd7fee68-2dbd-4a75-8c8f-59c6c329ccb8" deepnote_to_be_reexecuted=false source_hash="380ae6a" execution_start=1632778650794 execution_millis=30 is_code_hidden=true deepnote_cell_type="code"
from sklearn.linear_model import LinearRegression
# Get all Adelie penguins and put them in a DataFrame called `data`.
data = penguins[penguins['species'] == 'Adelie']
"""
Split the data into two sets: the independent variables
(bill_length_mm, bill_depth_mm, body_mass_g), and
the dependent variable (flipper_length_mm)
"""
x = data[['bill_length_mm', 'bill_depth_mm', 'body_mass_g']]
y = data['flipper_length_mm']
"""
Create a scikit-learn LinearRegression object which we will
fit to the data.
"""
lm = LinearRegression()
lm.fit(x,y)
# Print out the coefficients for each dependent variable
for (coef,col) in zip(lm.coef_, x.columns):
print("%-30s %+.3f" % ("Coefficient for " + col + ":", coef))
# + [markdown] tags=[] cell_id="00078-a1ded61f-3d38-4b25-a052-d1c183af1907" deepnote_cell_type="markdown"
# Perform the same regression with `statsmodel.api.sm` from the [StatsModels library](https://www.statsmodels.org/stable/index.html) in order to get more information about the model.
# + tags=[] cell_id="00079-b7760413-4190-4cba-82b4-6783f2b3a409" deepnote_to_be_reexecuted=false source_hash="5f20fb9f" execution_start=1632778650806 execution_millis=24 is_code_hidden=true is_output_hidden=false deepnote_cell_type="code"
import statsmodels.api as sm
# Get all Adelie penguins and put them in a DataFrame called `data`.
data = penguins[penguins["species"] == "Adelie"]
"""
Split the data into two sets: the independent variables
(bill_length_mm, bill_depth_mm, body_mass_g), and
the dependent variable (flipper_length_mm)
"""
x = data[["bill_length_mm", "bill_depth_mm", "body_mass_g"]]
y = data["flipper_length_mm"]
"""
Add y-intercept term to our linear model
"""
x = sm.add_constant(x)
"""
Perform linear regression
"""
lm = sm.OLS(y, x)
results = lm.fit()
"""
Print the results
"""
print(results.summary())
# + [markdown] cell_id="00040-7eee5896-2528-4cea-8ca3-ed9ffcfd7fb3" deepnote_app_coordinates={"x": 0, "y": 240, "w": 12, "h": 5} deepnote_cell_type="markdown"
# ## Additional References
#
# * O'Reilly provides a couple of good books that go in-depth about these tools and more:
# * [Python Data Science Handbook](http://shop.oreilly.com/product/0636920034919.do)
# * [Python for Data Analysis](http://shop.oreilly.com/product/0636920023784.do) -- this book was published in 2012 and may be slightly dated. However, the author provides some Jupyter Notebooks for free in [this repository](https://github.com/wesm/pydata-book) that you may find helpful.
# * Check out the full documentation for Jupyter on the [Project Jupyter site](https://jupyter.org/documentation).
# * Plotting tools:
# * Matplotlib
# * [Documentation](https://matplotlib.org/contents.html)
# * [Tutorials](https://matplotlib.org/tutorials/index.html)
# * Seaborn
# * [Documentation](https://seaborn.pydata.org/api.html)
# * [Introduction](https://seaborn.pydata.org/introduction.html)
# * [Statsmodels documentation](https://www.statsmodels.org/stable/index.html)
# + [markdown] tags=[] created_in_deepnote_cell=true deepnote_cell_type="markdown"
# <a style='text-decoration:none;line-height:16px;display:flex;color:#5B5B62;padding:10px;justify-content:end;' href='https://deepnote.com?utm_source=created-in-deepnote-cell&projectId=575af5dc-65a8-40d2-9ecb-58d75726c7c0' target="_blank">
# <img alt='Created in deepnote.com' style='display:inline;max-height:16px;margin:0px;margin-right:7.5px;' src='data:image/svg+xml;base64,PD94bWwgdmVyc2lvbj0iMS4wIiBlbmNvZGluZz0iVVRGLTgiPz4KPHN2ZyB3aWR0aD0iODBweCIgaGVpZ2h0PSI4MHB4IiB2aWV3Qm94PSIwIDAgODAgODAiIHZlcnNpb249IjEuMSIgeG1sbnM9Imh0dHA6Ly93d3cudzMub3JnLzIwMDAvc3ZnIiB4bWxuczp4bGluaz0iaHR0cDovL3d3dy53My5vcmcvMTk5OS94bGluayI+CiAgICA8IS0tIEdlbmVyYXRvcjogU2tldGNoIDU0LjEgKDc2NDkwKSAtIGh0dHBzOi8vc2tldGNoYXBwLmNvbSAtLT4KICAgIDx0aXRsZT5Hcm91cCAzPC90aXRsZT4KICAgIDxkZXNjPkNyZWF0ZWQgd2l0aCBTa2V0Y2guPC9kZXNjPgogICAgPGcgaWQ9IkxhbmRpbmciIHN0cm9rZT0ibm9uZSIgc3Ryb2tlLXdpZHRoPSIxIiBmaWxsPSJub25lIiBmaWxsLXJ1bGU9ImV2ZW5vZGQiPgogICAgICAgIDxnIGlkPSJBcnRib2FyZCIgdHJhbnNmb3JtPS<KEY> > </img>
# Created in <span style='font-weight:600;margin-left:4px;'>Deepnote</span></a>
| DataSciIntro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
####################################
# SETUP Libraries and Import Dataset
####################################
# Standard modules
import os
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# Scikit Learn modules
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression, Lasso, Ridge
# Visualizations
from yellowbrick.regressor import ResidualsPlot, PredictionError
# Files
data = "../model/final.csv"
# load dataframe
df = pd.read_csv(filepath_or_buffer=data, sep=",", encoding="utf-8", header=0)
df = df.drop(columns=["Wind"])
pd.set_option('display.max_columns', None)
df.head(3)
# +
####################################
# Linear Regression, Lasso, Ridge, and ElasticNet
####################################
# set samples matrix [n_samples, n_features] (X) and target (y)
features = ['Population','sin_day','cos_day','Temperature','Dew','Sky','Visibility','ATM','Wind-Dir','Wind-Rate']
target = ['pm25']
# fit data to model
X = df[features]
print(X.shape)
y = df[target]
print(y.shape)
# Split into training and testing
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=0.8,test_size=0.2, random_state=42)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
# +
####################################
# Linear Regression
####################################
# instantiate the model
lr = LinearRegression()
visualizer = ResidualsPlot(model=lr, hist=False)
# fit data to model
visualizer.fit(X_train, y_train)
print(f'Coefficients: {lr.coef_}\n')
print(f'Intercept: {lr.intercept_}\n')
# predict
#pred = lr.predict(X_test)
#count = len(pred)
#print(f'Prediction #: {count}')
#print(pred[0:5])
# score model
visualizer.score(X_test, y_test)
print(f'R2 Score: {lr.score(X_test, y_test)}')
visualizer.show()
# +
####################################
# Lasso Regression
####################################
# instantiate the model
lasso = Lasso(alpha=0.2)
visualizer = PredictionError(model=lasso)
# fit data to model
#lasso.fit(X_train, y_train)
visualizer.fit(X_train, y_train)
print(f'Coefficients: {lasso.coef_} \n')
print(f'Intercept: {lasso.intercept_} \n')
# predict
#pred = lr.predict(X_test)
#count = len(pred)
#print(f'Prediction #: {count}')
#print(pred[0:5])
# score model
visualizer.score(X_test, y_test)
print(f'R2 Score: {lasso.score(X_test, y_test)}')
#visualizer.show()
# +
####################################
# Ridge Regression
####################################
# instantiate the model
ridge = Ridge(alpha=0.2)
visualizer = PredictionError(model=ridge)
# fit data to model
#lasso.fit(X_train, y_train)
visualizer.fit(X_train, y_train)
print(f'Coefficients: {ridge.coef_} \n')
print(f'Intercept: {ridge.intercept_} \n')
# predict
#pred = lr.predict(X_test)
#count = len(pred)
#print(f'Prediction #: {count}')
#print(pred[0:5])
# score model
print(f'R2 Score: {ridge.score(X_test, y_test)}')
visualizer.score(X_test, y_test)
#visualizer.show()
# -
| model/exploration/Regression_Jeremy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import math
class A():
def a1(self):
print("A-a1")
def a2(self, a):
print("A-a2", a)
class B(A):
def b1(self):
A.a1(self)
print("B-b1")
def a2(self, b):
print(math.pi)
print("Override method")
def main():
a = A()
a.a1()
a.a2(2)
b = B()
b.b1()
b.a2(1)
if __name__ == "__main__":
main()
# -
| 5-class.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pipeline A
#
# - Imputation
# - Feature engineering (Ratios, aggregation)
# - Standardization (min-max)
# - Normalization
# - Binning and Re-coding
# - Feature Selection
# - Modeling
# +
# for preprocessing/eda models
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.preprocessing import power_transform
from scipy import stats
from scipy.stats import boxcox
from scipy.stats import kurtosis, skew
import math
from scipy.stats import norm
# feature selection
from sklearn.feature_selection import RFE
# balancing
from imblearn.over_sampling import SMOTE
# accuracy metrics and data split models
from sklearn.model_selection import train_test_split
from sklearn import metrics, model_selection
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
from sklearn.metrics import roc_auc_score
from sklearn.metrics import accuracy_score
from sklearn.metrics import precision_recall_fscore_support
# machine learning
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import RandomForestClassifier
from sklearn import metrics
from sklearn.ensemble import ExtraTreesClassifier
from xgboost import XGBClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.neural_network import MLPClassifier
from sklearn.neighbors import KNeighborsClassifier
pd.set_option('display.max_columns', 500) # display max 500 rows
pd.set_option('display.max_rows', 140)
# -
# read in data to skip innitial steps from pipeline B
data = pd.read_csv('online_shoppers_intention-2.csv')
data1 = data.copy() # copy of original df
data1.head()
data1.Weekend.value_counts()
# ## Imputation
# - assumption that exit rates cannot be zero
# impute our assumption (beccomes a new feature)
data1['ExitRatesImpute'] = data1['ExitRates'].replace(0,np.NaN)
data1['ExitRatesImpute'] = data1['ExitRatesImpute'].fillna(data1['ExitRatesImpute'].median())
# ## Feature Engineering (Ratios, aggregation)
# - calculate ratios for count of page visits
# - combine bounce and exit rates (average and weighted average - new features)
# - they have a strong linear relationship
# - divide features that have strong non-linear relationship
# - solve zero-division by replacing it with 0 (e.g. 1/0 = n/a)
# +
# ratios for counts
data1['totalFracAdmin'] = data1['Administrative']/(data1['Administrative'] + data1['Informational'] + data1['ProductRelated'])
data1['totalFracInfo'] = data1['Informational']/(data1['Administrative'] + data1['Informational'] + data1['ProductRelated'])
data1['totalFracProd'] = data1['ProductRelated']/(data1['Administrative'] + data1['Informational'] + data1['ProductRelated'])
# average combining
data1['BounceExitAvg'] = (data1['BounceRates'] + data1['ExitRates'])/2
# weighted-average feature joining
data1['BounceExitW1'] = data1['BounceRates'] * 0.6 + data1['ExitRates'] * 0.4
data1['BounceExitW2'] = data1['BounceRates'] * 0.7 + data1['ExitRates'] * 0.3
data1['BounceExitW3'] = data1['BounceRates'] * 0.4 + data1['ExitRates'] * 0.6
data1['BounceExitW4'] = data1['BounceRates'] * 0.3 + data1['ExitRates'] * 0.7
# bounce and exit rates vs page values ratio
data1['BouncePageRatio'] = data1['BounceRates']/data1['PageValues']
data1['ExitPageRatio'] = data1['ExitRates']/data1['PageValues']
# durations vs page values, bounce and exit rates
data1['InfoPageRatio'] = data1['Informational_Duration']/data1['PageValues']
data1['ProdRelPageRatio'] = data1['ProductRelated_Duration']/data1['PageValues']
data1['InfoBounceRatio'] = data1['Informational_Duration']/data1['BounceRates']
data1['AdminBounceRatio'] = data1['Administrative_Duration']/data1['BounceRates']
data1['ProdRelBounceRatio'] = data1['ProductRelated_Duration']/data1['BounceRates']
data1['InfoExitRatio'] = data1['Informational_Duration']/data1['ExitRates']
data1['AdminBounceRatio'] = data1['Administrative_Duration']/data1['ExitRates']
data1['ProdRelExitRatio'] = data1['ProductRelated_Duration']/data1['ExitRates']
# page values, bounce and exit rates vs durations
#data1['PageInfoRatio'] = data1['PageValues']/data1['Informational_Duration']
#data1['PageProdRelRatio'] = data1['PageValues']/data1['ProductRelated_Duration']
#data1['BounceInfoRatio'] = data1['BounceRates']/data1['Informational_Duration']
#data1['BounceAdminRatio'] = data1['BounceRates']/data1['Administrative_Duration']
#data1['BounceProdRelRatio'] = data1['BounceRates']/data1['ProductRelated_Duration']
#data1['ExitInfoRatio'] = data1['ExitRates']/data1['Informational_Duration']
#data1['BounceAdminRatio'] = data1['ExitRates']/data1['Administrative_Duration']
#data1['ExitProdRelRatio'] = data1['ExitRates']/data1['ProductRelated_Duration']
# as there are many zero values --> e.g. x/0 (zero-division) = N/A or inf can occur
# if there is zero-division treat that as a zero
data1 = data1.fillna(0) # fill N/A with 0
data1 = data1.replace(np.inf, 0) # replace inf wit 0
data1 = data1.replace(-0, 0) # for some reason we also get -0 just fix it to 0
# -
print('Original number of features: ', len(data.columns))
print('Number of features added: ', len(data1.columns) - len(data.columns))
print('Total number of features after feature engineering: ', len(data1.columns) + len(data.columns))
# ## Standardization
# make a copy
standardize = data1.copy()
# select continous features (exclude Special Day)
standardize = standardize.select_dtypes(include='float64').drop('SpecialDay', axis = 1)
# +
# import MinMaxScaler module
from sklearn.preprocessing import MinMaxScaler
# use MinMaxScaler function
scaler = MinMaxScaler()
# min-max standerdize all continous columns
standardize[standardize.columns] = scaler.fit_transform(standardize[standardize.columns]) # fit the scaler to the model
standardize_done = standardize.add_suffix('_Scaled') # add suffix (new features)
# add new standerdized features to data1
data1 = pd.concat([data1, standardize_done], axis = 1 )
#use .describe() to prove standardization worked
standardize_done.describe()
# -
print('Number of features before standardization: ', len(data1.columns) - len(standardize_done.columns))
print('Number of features added: ', len(standardize_done.columns))
print('Total number of features after standardization: ', len(data1.columns))
# ## Normalization
# - Normalize continous values
# - Requirement: -0.5 < .skew() < 0.5
normalization_df = standardize_done.copy()
# ### Features that need to be Normalized
# - Note: original features that were not scaled were added as well
# find skewed features
normalization_needed = [] # empty list for cols that need normalizing
normalization_df_cols = normalization_df.columns
for col in normalization_df_cols:
if normalization_df[col].skew() > 0.5:
print('Right skew: ',col,normalization_df[col].skew())
normalization_needed.append(col)
elif normalization_df[col].skew() < -0.5:
print('Left skew: ',col,normalization_df[col].skew())
normalization_needed.append(col)
else:
""
print(len(normalization_needed),' features need normalizing')
# function to create a histogram
# source: https://stackoverflow.com/questions/20011122/fitting-a-normal-distribution-to-1d-data
def plot_hist(df, column, bin_amount=25):
# Fit a normal distribution to the data:
mu, std = norm.fit(df[column])
# Plot the histogram.
## Set the alpha value used for blending (how transparent the color is)
plt.hist(df[column], bins=bin_amount, density=True, alpha=0.8, color='g')
# Plot the PDF.
xmin, xmax = plt.xlim()
x = np.linspace(xmin, xmax, 100)
p = norm.pdf(x, mu, std)
plt.plot(x, p, 'k', linewidth=2)
title = "Fit results: mu = %.2f, std = %.2f" % (mu, std)
plt.title(title)
plt.xlabel(col)
plt.show()
# ### Histograms for all features before normalization
# - Some are highly skewed and might not be possible to normalize
for col in normalization_df_cols:
plot_hist(normalization_df, col)
# ### Applying Normalization
# +
# innitial yeo-johnson normalization
for col in normalization_needed:
if normalization_df[col].skew() > 0.5:
normalization_df[col] = stats.yeojohnson(normalization_df[col])[0]
# replace -0 with 0
normalization_df = normalization_df.replace(-0, 0)
# square-root transformation for positively/rightly skewed features that still have a skew > 0.5
for col in normalization_df_cols:
if normalization_df[col].skew() > 0.5:
normalization_df[col] = np.sqrt(normalization_df[col])
# left skew transformation
normalization_df['totalFracProd_Scaled'] = normalization_df['totalFracProd_Scaled']**2
# right manual transformation
normalization_df['AdminBounceRatio_Scaled'] = np.sqrt(normalization_df['AdminBounceRatio_Scaled'])
normalization_df['ProdRelBounceRatio_Scaled'] = np.sqrt(np.sqrt(normalization_df['ProdRelBounceRatio_Scaled']))
# -
# ### Skew after normalization
not_normalized = []
# if still not normal display
for col in normalization_df_cols:
if normalization_df[col].skew() > 0.5:
print('Right skew: ',col,normalization_df[col].skew())
not_normalized.append(col)
elif normalization_df[col].skew() < -0.5:
print('Left skew: ',col,normalization_df[col].skew())
not_normalized.append(col)
else:
print('Normalized', col,normalization_df[col].skew())
print('Number of features that were not normalized: ',len(not_normalized))
print('Number of features that were normalized: ', len(normalization_needed) - len(not_normalized))
# ### Checking distribution of features 10 features not fully normalized
# - they look almost binary (contain a lot of zero-values)
# - we will create new features by binning them on some threshold (e.g. zero or not)
for col in not_normalized:
plot_hist(normalization_df, col)
# ### Most frequent values for not fully normalized features
# - are moslty zeroes so we will bin them depending on whether the value is a zero or not
# - more zeroes occur in ratios as zero-division can be encountered
for col in not_normalized:
print(col, '\n',normalization_df[col].value_counts().head(1))
# ### Add suffix and merge with Data1
# - add suffix to columns that needed normalization even the ones that could not be fully normalized
for col in normalization_needed: # All needed normalization
normalization_done = normalization_df.add_suffix('_Norm') # add suffix to distinguish
# concat normalized features with scaled and original
data1 = pd.concat([data1,normalization_done], axis = 1)
# we now have 84 columns
data1.shape
# ## Binning and Encoding
# 1. Bin and encode original categorical and text data
# - Bin month to quarters and one-hot
# - Encode visitor type to numeric and one-hot
# - One-hot special day
# - Convert to correct data type
# 2. Bin not fully normalized features
# +
#1. Bin and encode original categorical and text data
#binning the Month column by quarter(as seen above)
#new column created-month_bin will have months binned by their respective quarters
def Month_bin(Month) :
if Month == 'Jan':
return 1
elif Month == 'Feb':
return 1
elif Month == 'Mar':
return 1
elif Month == 'Apr':
return 2
elif Month == 'May':
return 2
elif Month == 'June':
return 2
elif Month == 'Jul':
return 3
elif Month == 'Aug':
return 3
elif Month == 'Sep':
return 3
elif Month == 'Oct':
return 4
elif Month == 'Nov':
return 4
elif Month == 'Dec':
return 4
data1['Month_bin'] = data1['Month'].apply(Month_bin)
#binning VisitorType
#creating new column--VisitorType_bin
def VisitorType_bin(VisitorType) :
if VisitorType == 'Returning_Visitor':
return 1
elif VisitorType == 'New_Visitor':
return 2
elif VisitorType == 'Other':
return 3
# apply function
data1['VisitorType_bin'] = data1['VisitorType'].apply(VisitorType_bin)
# get dummies
data1 = pd.get_dummies(data1, columns=['VisitorType_bin','Month_bin','SpecialDay'])
# convert to bool
data1[['VisitorType_bin_1', 'VisitorType_bin_2', 'VisitorType_bin_3',
'Month_bin_1', 'Month_bin_2', 'Month_bin_3', 'Month_bin_4','SpecialDay_0.0', 'SpecialDay_0.2',
'SpecialDay_0.4', 'SpecialDay_0.6', 'SpecialDay_0.8', 'SpecialDay_1.0']] = data1[['VisitorType_bin_1',
'VisitorType_bin_2', 'VisitorType_bin_3','Month_bin_1', 'Month_bin_2', 'Month_bin_3', 'Month_bin_4','SpecialDay_0.0',
'SpecialDay_0.2','SpecialDay_0.4', 'SpecialDay_0.6', 'SpecialDay_0.8', 'SpecialDay_1.0']].astype(bool)
def browser_fun(series):
if series == 2 or series == 1:
return 1
elif series == 4 or series == 5 or series == 6 or series == 10 or series == 8 or series == 3:
return 2
else:
return 3
# apply function for browser
data1['Browser_Bin'] = data1['Browser'].apply(browser_fun)
def TrafficType_fun(series):
if series == 2 or series == 1 or series == 3 or series == 4:
return 1
elif series == 13 or series == 10 or series == 6 or series == 8 or series == 5 or series == 11 or series == 20:
return 2
else:
return 3
# apply function for TrafficType
data1['TrafficType_Bin'] = data1['TrafficType'].apply(TrafficType_fun)
def RegionFun(series):
if series == 1 or series == 3:
return 1
elif series == 4 or series == 2:
return 2
else:
return 3
# apply function for Region
data1['Region_Bin'] = data1['Region'].apply(RegionFun)
def OperatingSystemsFun(series):
if series == 2:
return 1
elif series == 1 or series == 3:
return 2
else:
return 3
# apply function for TrafficType
data1['OperatingSystems_Bin'] = data1['OperatingSystems'].apply(OperatingSystemsFun)
data1 = pd.get_dummies(data1, columns=['Browser_Bin','TrafficType_Bin','Region_Bin','OperatingSystems_Bin'])
# convert to bool
data1[['Browser_Bin_1','Browser_Bin_2','Browser_Bin_3','TrafficType_Bin_1','TrafficType_Bin_2','TrafficType_Bin_3',
'Region_Bin_1','Region_Bin_2','Region_Bin_3','OperatingSystems_Bin_1','OperatingSystems_Bin_2',
'OperatingSystems_Bin_3']] = data1[['Browser_Bin_1','Browser_Bin_2','Browser_Bin_3','TrafficType_Bin_1','TrafficType_Bin_2','TrafficType_Bin_3',
'Region_Bin_1','Region_Bin_2','Region_Bin_3','OperatingSystems_Bin_1','OperatingSystems_Bin_2',
'OperatingSystems_Bin_3']].astype(bool)
# +
# 2. Bin not fully normalized features
# if it has some value return 1 otherwise 0
def zero_or_not(series):
if series == 0:
return 0
else:
return 1
# apply function
not_normalized.pop(3) # remove left skew feature from the list
for col in not_normalized:
data1[col+'_Bin'] = data1[col].apply(zero_or_not)
# conver to bool (occupies less space than int)
data1[col+'_Bin'] = data1[col+'_Bin'].astype(bool)
# function for the left skewed feature
def left_skew(series):
if series >= 0.4:
return 1
else:
return 0
# convert to bool
data1['totalFracProd_Bin'] = data1['totalFracProd_Scaled'].apply(left_skew).astype(bool)
# -
print("Data shape: ", data1.shape)
print("Nbr dtype integer: ", len(data1.select_dtypes(include='int64').columns))
print("Nbr dtype float: ", len(data1.select_dtypes(include='float64').columns))
print("Nbr dtype bool: ", len(data1.select_dtypes(include='bool').columns))
print("Nbr dtype object: ", len(data1.select_dtypes(include='object').columns))
# # Merge Pipeline A and Pipeline B
# +
# drop original/unprocessed/object columns
pipeline_A = data1.drop(['Administrative','Administrative_Duration','Informational','Informational_Duration','ProductRelated','ProductRelated_Duration',
'BounceRates','ExitRates','PageValues','Month','VisitorType','ExitRatesImpute','totalFracAdmin','totalFracInfo',
'totalFracProd','BounceExitAvg','BounceExitW1','BounceExitW2','BounceExitW3','BounceExitW4','BouncePageRatio',
'ExitPageRatio','InfoPageRatio','ProdRelPageRatio','InfoBounceRatio','AdminBounceRatio',
'ProdRelBounceRatio','InfoExitRatio','ProdRelExitRatio', 'Browser','Region','TrafficType','Region','OperatingSystems'],
axis=1)
# Read in Data
pipeline_B = pd.read_csv('model_data_B.csv', index_col = 0)
pipeline_B = pipeline_B.select_dtypes(include = ['float64']).drop(['Administrative_Duration_Norm',
'Informational_Duration_Norm',
'ProductRelated_Duration_Norm', 'BounceRates_Norm', 'ExitRates_Norm',
'PageValues_Norm', 'ExitRatesImpute_Norm', 'totalFracAdmin_Norm',
'totalFracInfo_Norm', 'totalFracProd_Norm', 'BounceExitAvg_Norm',
'BounceExitW1_Norm', 'BounceExitW2_Norm', 'BounceExitW3_Norm',
'BounceExitW4_Norm', 'BouncePageRatio_Norm', 'ExitPageRatio_Norm',
'InfoPageRatio_Norm', 'ProdRelPageRatio_Norm', 'InfoBounceRatio_Norm',
'AdminBounceRatio_Norm', 'ProdRelBounceRatio_Norm',
'InfoExitRatio_Norm', 'ProdRelExitRatio_Norm'], axis = 1)
# MERGE
model_data = pd.concat([pipeline_A,pipeline_B], axis = 1)
# select X and y
X = model_data.drop('Revenue', axis =1) #features
x_col = model_data.drop('Revenue', axis =1)
y = model_data['Revenue'] #target
feature_name = X.columns.tolist()
# save preprocessed data
model_data.to_csv('all_model_data.csv')
# nbr of rows, nbr of columns
print(model_data.shape)
print(X.shape) # features
print(y.shape) # label/target
# -
model_data = pd.read_csv('all_model_data.csv', index_col = 0)
#model_data.head()
# ## 10 Way Feature Selection
# - select 50 features from 136
# - xxx_support: list to represent select this feature or not
# - xxx_feature: the name of selected features
#
# ### Methods:
# 1. Filter: Pearson, f_classif (Anova F value)
# 2. Wrapper: RFE with Logistic regression and XGBoost
# 3. Embeded: Logistic Regression, Random Forest, XGBoost, LassoCV, RidgeClassifierCV
#
# Source of Inspiration and modified: https://www.kaggle.com/sz8416/6-ways-for-feature-selection
# select X and y
X = model_data.drop('Revenue', axis = 1)
feature_name = X.columns.tolist()
y = model_data.Revenue
# ### 1 Filter
# #### 1.1 Pearson Correlation
def cor_selector(X, y):
cor_list = []
# calculate the correlation with y for each feature
for i in X.columns.tolist():
cor = np.corrcoef(X[i], y)[0, 1]
cor_list.append(cor)
# replace NaN with 0
cor_list = [0 if np.isnan(i) else i for i in cor_list]
# feature name
cor_feature = X.iloc[:,np.argsort(np.abs(cor_list))[-50:]].columns.tolist()
# feature selection? 0 for not select, 1 for select
cor_support = [True if i in cor_feature else False for i in feature_name]
return cor_support, cor_feature
cor_support, cor_feature = cor_selector(X, y)
print(str(len(cor_feature)), 'selected features')
# #### 1.2 f_classif
# - documentation for SelectKBest: http://scikit-learn.org/stable/modules/generated/sklearn.feature_selection.SelectKBest.html
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import f_classif
f_classif_selector = SelectKBest(f_classif, k=50)
f_classif_selector.fit(X, y)
f_classif_support = f_classif_selector.get_support()
f_classif_feature = X.loc[:,f_classif_support].columns.tolist()
print(str(len(f_classif_feature)), 'selected features')
# #### 2. Wrapper
# - documentation for RFE: http://scikit-learn.org/stable/modules/generated/sklearn.feature_selection.RFE.html
# - logistic regression and xgboost
#
# #### 2.1 RFE - Logistic Regression<br>
# RFE notes:
# - estimator = means what model to estimate it on
# - max_iter = if model doesn't converge, random state for reusability
# - step = how many observation to remove after each iteration
# - verbose = can get more visual output (doesn't change the model)
# packages you would need
#from sklearn.feature_selection import RFE
#from sklearn.linear_model import LogisticRegression
rfe_selector = RFE(estimator=LogisticRegression(max_iter = 1500,random_state=123), step = 10, n_features_to_select=50,
verbose=0)
rfe_selector.fit(X, y)
rfe_support = rfe_selector.get_support() # Get a mask, or integer index, of the features selected
rfe_feature = X.loc[:,rfe_support].columns.tolist() # get the column names of features selected and put them in a list
print(str(len(rfe_feature)), 'selected features')
# #### 2.2 RFE XGBOOST
rfe_selector_xgboost = RFE(estimator=XGBClassifier(random_state=123), n_features_to_select=50, step=10, verbose=0)
rfe_selector_xgboost.fit(X, y)
# transform
rfe_support_xgboost = rfe_selector_xgboost.get_support()
rfe_feature_xgboost = X.loc[:,rfe_support_xgboost].columns.tolist()
print(str(len(rfe_support_xgboost)), 'selected features')
# ### 3. Embeded
# - documentation for SelectFromModel: http://scikit-learn.org/stable/modules/generated/sklearn.feature_selection.SelectFromModel.html
#
# #### 3.1 Logistics Regression
from sklearn.feature_selection import SelectFromModel
#from sklearn.linear_model import LogisticRegression
# penalty l2 is default (regularization type for solver)
# threshold = minimum threshold applied (applied so it selects approx 50 features )
embeded_lr_selector = SelectFromModel(LogisticRegression(penalty="l2", random_state = 123, max_iter=1000), threshold = 0.2)
embeded_lr_selector.fit(X, y)
embeded_lr_support = embeded_lr_selector.get_support()
embeded_lr_feature = X.loc[:,embeded_lr_support].columns.tolist()
print(str(len(embeded_lr_feature)), 'selected features')
# #### 3.2 Random Forest
#from sklearn.feature_selection import SelectFromModel
#from sklearn.ensemble import RandomForestClassifier
# n_estimators = The number of trees in the forest (10-100)
embeded_rf_selector = SelectFromModel(RandomForestClassifier(n_estimators=50, random_state = 123), threshold=0.00775)
embeded_rf_selector.fit(X, y)
embeded_rf_support = embeded_rf_selector.get_support()
embeded_rf_feature = X.loc[:,embeded_rf_support].columns.tolist()
print(str(len(embeded_rf_feature)), 'selected features')
# #### 3.3 XGBoost
embeded_xgb_selector = SelectFromModel(XGBClassifier(n_estimators=50, random_state = 123))
embeded_xgb_selector.fit(X, y)
embeded_xgb_support = embeded_xgb_selector.get_support()
embeded_xgb_feature = X.loc[:,embeded_xgb_support].columns.tolist()
print(str(len(embeded_xgb_feature)), 'selected features')
# ### 3.4 LassoCV
# - Lasso linear model with iterative fitting along a regularization path (built-in cross validation)
# - https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LassoCV.html
from sklearn.linear_model import LassoCV
# cv = number of k for cross validation
embeded_lasso_selector = SelectFromModel(LassoCV(random_state = 123, cv = 10, max_iter = 2000),threshold = 0.0001)
embeded_lasso_selector.fit(X, y)
embeded_lasso_support = embeded_lasso_selector.get_support()
embeded_lasso_feature = X.loc[:,embeded_lasso_support].columns.tolist()
print(str(len(embeded_lasso_feature)), 'selected features')
# ### 3.5 Ridge Classifier CV
# - Ridge classifier with built-in cross-validation
# - https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.RidgeClassifierCV.html#sklearn.linear_model.RidgeClassifierCV
from sklearn.linear_model import RidgeClassifierCV
embeded_ridge_selector = SelectFromModel(RidgeClassifierCV(cv=10), threshold =0.059)
embeded_ridge_selector.fit(X, y)
embeded_ridge_support = embeded_ridge_selector.get_support()
embeded_ridge_feature = X.loc[:,embeded_ridge_support].columns.tolist()
print(str(len(embeded_ridge_feature)), 'selected features')
# ### 3.6 Linear SVC
# - https://scikit-learn.org/stable/modules/feature_selection.html
# +
from sklearn.svm import LinearSVC
embeded_svc_selector = SelectFromModel(LinearSVC(C=0.5, penalty='l1', dual=False, max_iter = 5000),threshold = 0.001)
embeded_svc_selector.fit(X, y)
embeded_svc_support = embeded_svc_selector.get_support()
embeded_svc_feature = X.loc[:,embeded_svc_support].columns.tolist()
print(str(len(embeded_svc_feature)), 'selected features')
# -
# ## Summary
# - Contains features that were derived from similar features
# - When modeling only one version (better one should be used)
pd.set_option('display.max_rows', 100)
# put all selection together
feature_selection_df = pd.DataFrame({'Feature':feature_name, 'Pearson':cor_support,'f_classif':f_classif_support,
'RFE-Log':rfe_support,'RFE-XGBoost': rfe_support_xgboost,'Logistics':embeded_lr_support,'LassoCV':embeded_lasso_support,
'RidgeClassifierCV':embeded_ridge_support,'Random Forest':embeded_rf_support,'XGBoost':embeded_xgb_support,
'LinearSVC':embeded_svc_support})
# count the selected times for each feature
feature_selection_df['Total'] = np.sum(feature_selection_df, axis=1)
# display the top 65
feature_selection_df = feature_selection_df.sort_values(['Total','Feature'] , ascending=False)
feature_selection_df.index = range(1, len(feature_selection_df)+1)
feature_selection_df.head(65)
feature_selection_df.to_csv('feature_selection.csv')
| project2-mem-master/3_Data_Preprocessing/pipelineA/PipelineA_full.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Module 5: Hierarchical Generators
# This module covers writing layout/schematic generators that instantiate other generators. We will write a two-stage amplifier generator, which instatiates the common-source amplifier followed by the source-follower amplifier.
# ## AmpChain Layout Example
# First, we will write a layout generator for the two-stage amplifier. The layout floorplan is drawn for you below:
# <img src="bootcamp_pics/5_hierarchical_generator/hierachical_generator_1.PNG" alt="Drawing" style="width: 400px;"/>
# This floorplan abuts the `AmpCS` instance next to `AmpSF` instance, the `VSS` ports are simply shorted together, and the top `VSS` port of `AmpSF` is ignored (they are connected together internally by dummy connections). The intermediate node of the two-stage amplifier is connected using a vertical routing track in the middle of the two amplifier blocks. `VDD` ports are connected to the top-most M6 horizontal track, and other ports are simply exported in-place.
#
# The layout generator is reproduced below, with some parts missing (which you will fill out later). We will walk through the important sections of the code.
# ```python
# class AmpChain(TemplateBase):
# def __init__(self, temp_db, lib_name, params, used_names, **kwargs):
# super(AmpChain, self).__init__(temp_db, lib_name, params, used_names, **kwargs)
# self._sch_params = None
#
# @property
# def sch_params(self):
# return self._sch_params
#
# @classmethod
# def get_params_info(cls):
# return dict(
# cs_params='common source amplifier parameters.',
# sf_params='source follower parameters.',
# show_pins='True to draw pin geometries.',
# )
#
# def draw_layout(self):
# """Draw the layout of a transistor for characterization.
# """
#
# # make copies of given dictionaries to avoid modifying external data.
# cs_params = self.params['cs_params'].copy()
# sf_params = self.params['sf_params'].copy()
# show_pins = self.params['show_pins']
#
# # disable pins in subcells
# cs_params['show_pins'] = False
# sf_params['show_pins'] = False
#
# # create layout masters for subcells we will add later
# cs_master = self.new_template(params=cs_params, temp_cls=AmpCS)
# # TODO: create sf_master. Use AmpSFSoln class
# sf_master = None
#
# if sf_master is None:
# return
#
# # add subcell instances
# cs_inst = self.add_instance(cs_master, 'XCS')
# # add source follower to the right of common source
# x0 = cs_inst.bound_box.right_unit
# sf_inst = self.add_instance(sf_master, 'XSF', loc=(x0, 0), unit_mode=True)
#
# # get VSS wires from AmpCS/AmpSF
# cs_vss_warr = cs_inst.get_all_port_pins('VSS')[0]
# sf_vss_warrs = sf_inst.get_all_port_pins('VSS')
# # only connect bottom VSS wire of source follower
# if sf_vss_warrs[0].track_id.base_index < sf_vss_warrs[1].track_id.base_index:
# sf_vss_warr = sf_vss_warrs[0]
# else:
# sf_vss_warr = sf_vss_warrs[1]
#
# # connect VSS of the two blocks together
# vss = self.connect_wires([cs_vss_warr, sf_vss_warr])[0]
#
# # get layer IDs from VSS wire
# hm_layer = vss.layer_id
# vm_layer = hm_layer + 1
# top_layer = vm_layer + 1
#
# # calculate template size
# tot_box = cs_inst.bound_box.merge(sf_inst.bound_box)
# self.set_size_from_bound_box(top_layer, tot_box, round_up=True)
#
# # get subcell ports as WireArrays so we can connect them
# vmid0 = cs_inst.get_all_port_pins('vout')[0]
# vmid1 = sf_inst.get_all_port_pins('vin')[0]
# vdd0 = cs_inst.get_all_port_pins('VDD')[0]
# vdd1 = sf_inst.get_all_port_pins('VDD')[0]
#
# # get vertical VDD TrackIDs
# vdd0_tid = TrackID(vm_layer, self.grid.coord_to_nearest_track(vm_layer, vdd0.middle))
# vdd1_tid = TrackID(vm_layer, self.grid.coord_to_nearest_track(vm_layer, vdd1.middle))
#
# # connect VDD of each block to vertical M5
# vdd0 = self.connect_to_tracks(vdd0, vdd0_tid)
# vdd1 = self.connect_to_tracks(vdd1, vdd1_tid)
# # connect M5 VDD to top M6 horizontal track
# vdd_tidx = self.grid.get_num_tracks(self.size, top_layer) - 1
# vdd_tid = TrackID(top_layer, vdd_tidx)
# vdd = self.connect_to_tracks([vdd0, vdd1], vdd_tid)
#
# # TODO: connect vmid0 and vmid1 to vertical track in the middle of two templates
# # hint: use x0
# vmid = None
#
# if vmid is None:
# return
#
# # add pins on wires
# self.add_pin('vmid', vmid, show=show_pins)
# self.add_pin('VDD', vdd, show=show_pins)
# self.add_pin('VSS', vss, show=show_pins)
# # re-export pins on subcells.
# self.reexport(cs_inst.get_port('vin'), show=show_pins)
# self.reexport(cs_inst.get_port('vbias'), net_name='vb1', show=show_pins)
# # TODO: reexport vout and vbias of source follower
# # TODO: vbias should be renamed to vb2
#
# # compute schematic parameters.
# self._sch_params = dict(
# cs_params=cs_master.sch_params,
# sf_params=sf_master.sch_params,
# )
# ```
# ## AmpChain Constructor
# ```python
# class AmpChain(TemplateBase):
# def __init__(self, temp_db, lib_name, params, used_names, **kwargs):
# super(AmpChain, self).__init__(temp_db, lib_name, params, used_names, **kwargs)
# self._sch_params = None
#
# @property
# def sch_params(self):
# return self._sch_params
#
# @classmethod
# def get_params_info(cls):
# return dict(
# cs_params='common source amplifier parameters.',
# sf_params='source follower parameters.',
# show_pins='True to draw pin geometries.',
# )
# ```
# First, notice that instead of subclassing `AnalogBase`, the `AmpChain` class subclasses `TemplateBase`. This is because we are not trying to draw transistor rows inside this layout generator; we just want to place and route multiple layout instances together. `TemplateBase` is the base class for all layout generators and it provides most placement and routing methods you need.
#
# Next, notice that the parameters for `AmpChain` are simply parameter dictionaries for the two sub-generators. The ability to use complex data structures as generator parameters solves the parameter explosion problem when writing generators with many levels of hierarchy.
# ## Creating Layout Master
# ```python
# # create layout masters for subcells we will add later
# cs_master = self.new_template(params=cs_params, temp_cls=AmpCS)
# # TODO: create sf_master. Use AmpSFSoln class
# sf_master = None
# ```
# Here, the `new_template()` function creates a new layout master, `cs_master`, which represents a generated layout cellview from the `AmpCS` layout generator. We can later instances of this master in the current layout, which are references to the generated `AmpCS` layout cellview, perhaps shifted and rotated. The main take away is that the `new_template()` function does not add any layout geometries to the current layout, but rather create a separate layout cellview which we may use later.
# ## Creating Layout Instance
# ```python
# # add subcell instances
# cs_inst = self.add_instance(cs_master, 'XCS')
# # add source follower to the right of common source
# x0 = cs_inst.bound_box.right_unit
# sf_inst = self.add_instance(sf_master, 'XSF', loc=(x0, 0), unit_mode=True)
# ```
#
# The `add_instance()` method adds an instance of the given layout master to the current cellview. By default, if no location or orientation is given, it puts the instance at the origin with no rotation. the `bound_box` attribute can then be used on the instance to get the bounding box of the instance. Here, the bounding box is used to determine the X coordinate of the source-follower.
# ## Get Instance Ports
# ```python
# # get subcell ports as WireArrays so we can connect them
# vmid0 = cs_inst.get_all_port_pins('vout')[0]
# vmid1 = sf_inst.get_all_port_pins('vin')[0]
# vdd0 = cs_inst.get_all_port_pins('VDD')[0]
# vdd1 = sf_inst.get_all_port_pins('VDD')[0]
# ```
# after adding an instance, the `get_all_port_pins()` function can be used to obtain a list of all pins as `WireArray` objects with the given name. In this case, we know that there's exactly one pin, so we use Python list indexing to obtain first element of the list.
# ## Routing Grid Object
# ```python
# # get vertical VDD TrackIDs
# vdd0_tid = TrackID(vm_layer, self.grid.coord_to_nearest_track(vm_layer, vdd0.middle))
# vdd1_tid = TrackID(vm_layer, self.grid.coord_to_nearest_track(vm_layer, vdd1.middle))
# ```
#
# the `self.grid` attribute of `TemplateBase` is a `RoutingGrid` objects, which provides many useful functions related to the routing grid. In this particular scenario, `coord_to_nearest_track()` is used to determine the vertical track index closest to the center of the `VDD` ports. These vertical tracks will be used later to connect the `VDD` ports together.
# ## Re-export Pins on Instances
# ```python
# # re-export pins on subcells.
# self.reexport(cs_inst.get_port('vin'), show=show_pins)
# self.reexport(cs_inst.get_port('vbias'), net_name='vb1', show=show_pins)
# # TODO: reexport vout and vbias of source follower
# # TODO: vbias should be renamed to vb2
# ```
# `TemplateBase` also provides a `reexport()` function, which is a convenience function to re-export an instance port in-place. The `net_name` optional parameter can be used to change the port name. In this example, the `vbias` port of common-source amplifier is renamed to `vb1`.
# ## Layout Exercises
# Now you should know everything you need to finish the two-stage amplifier layout generator. Fill in the missing pieces to do the following:
#
# 1. Create layout master for `AmpSF` using the `AmpSFSoln` class.
# 2. Using `RoutingGrid`, determine the vertical track index in the middle of the two amplifier blocks, and connect `vmid` wires together using this track.
# * Hint: variable `x0` is the X coordinate of the boundary between the two blocks.
# 3. Re-export `vout` and `vbias` of the source-follower. Rename `vbias` to `vb2`.
#
# Once you're done, evaluate the cell below, which will generate the layout and run LVS. If everything is done correctly, a layout should be generated inthe `DEMO_AMP_CHAIN` library, and LVS should pass.
# +
from bag.layout.routing import TrackID
from bag.layout.template import TemplateBase
from xbase_demo.demo_layout.core import AmpCS, AmpSFSoln
class AmpChain(TemplateBase):
def __init__(self, temp_db, lib_name, params, used_names, **kwargs):
super(AmpChain, self).__init__(temp_db, lib_name, params, used_names, **kwargs)
self._sch_params = None
@property
def sch_params(self):
return self._sch_params
@classmethod
def get_params_info(cls):
return dict(
cs_params='common source amplifier parameters.',
sf_params='source follower parameters.',
show_pins='True to draw pin geometries.',
)
def draw_layout(self):
"""Draw the layout of a transistor for characterization.
"""
# make copies of given dictionaries to avoid modifying external data.
cs_params = self.params['cs_params'].copy()
sf_params = self.params['sf_params'].copy()
show_pins = self.params['show_pins']
# disable pins in subcells
cs_params['show_pins'] = False
sf_params['show_pins'] = False
# create layout masters for subcells we will add later
cs_master = self.new_template(params=cs_params, temp_cls=AmpCS)
# TODO: create sf_master. Use AmpSFSoln class
sf_master = None
if sf_master is None:
return
# add subcell instances
cs_inst = self.add_instance(cs_master, 'XCS')
# add source follower to the right of common source
x0 = cs_inst.bound_box.right_unit
sf_inst = self.add_instance(sf_master, 'XSF', loc=(x0, 0), unit_mode=True)
# get VSS wires from AmpCS/AmpSF
cs_vss_warr = cs_inst.get_all_port_pins('VSS')[0]
sf_vss_warrs = sf_inst.get_all_port_pins('VSS')
# only connect bottom VSS wire of source follower
if sf_vss_warrs[0].track_id.base_index < sf_vss_warrs[1].track_id.base_index:
sf_vss_warr = sf_vss_warrs[0]
else:
sf_vss_warr = sf_vss_warrs[1]
# connect VSS of the two blocks together
vss = self.connect_wires([cs_vss_warr, sf_vss_warr])[0]
# get layer IDs from VSS wire
hm_layer = vss.layer_id
vm_layer = hm_layer + 1
top_layer = vm_layer + 1
# calculate template size
tot_box = cs_inst.bound_box.merge(sf_inst.bound_box)
self.set_size_from_bound_box(top_layer, tot_box, round_up=True)
# get subcell ports as WireArrays so we can connect them
vmid0 = cs_inst.get_all_port_pins('vout')[0]
vmid1 = sf_inst.get_all_port_pins('vin')[0]
vdd0 = cs_inst.get_all_port_pins('VDD')[0]
vdd1 = sf_inst.get_all_port_pins('VDD')[0]
# get vertical VDD TrackIDs
vdd0_tid = TrackID(vm_layer, self.grid.coord_to_nearest_track(vm_layer, vdd0.middle))
vdd1_tid = TrackID(vm_layer, self.grid.coord_to_nearest_track(vm_layer, vdd1.middle))
# connect VDD of each block to vertical M5
vdd0 = self.connect_to_tracks(vdd0, vdd0_tid)
vdd1 = self.connect_to_tracks(vdd1, vdd1_tid)
# connect M5 VDD to top M6 horizontal track
vdd_tidx = self.grid.get_num_tracks(self.size, top_layer) - 1
vdd_tid = TrackID(top_layer, vdd_tidx)
vdd = self.connect_to_tracks([vdd0, vdd1], vdd_tid)
# TODO: connect vmid0 and vmid1 to vertical track in the middle of two templates
# hint: use x0
vmid = None
if vmid is None:
return
# add pins on wires
self.add_pin('vmid', vmid, show=show_pins)
self.add_pin('VDD', vdd, show=show_pins)
self.add_pin('VSS', vss, show=show_pins)
# re-export pins on subcells.
self.reexport(cs_inst.get_port('vin'), show=show_pins)
self.reexport(cs_inst.get_port('vbias'), net_name='vb1', show=show_pins)
# TODO: reexport vout and vbias of source follower
# TODO: vbias should be renamed to vb2
# compute schematic parameters.
self._sch_params = dict(
cs_params=cs_master.sch_params,
sf_params=sf_master.sch_params,
)
import os
# import bag package
import bag
from bag.io import read_yaml
# import BAG demo Python modules
import xbase_demo.core as demo_core
# load circuit specifications from file
spec_fname = os.path.join(os.environ['BAG_WORK_DIR'], 'specs_demo/demo.yaml')
top_specs = read_yaml(spec_fname)
# obtain BagProject instance
local_dict = locals()
if 'bprj' in local_dict:
print('using existing BagProject')
bprj = local_dict['bprj']
else:
print('creating BagProject')
bprj = bag.BagProject()
demo_core.run_flow(bprj, top_specs, 'amp_chain_soln', AmpChain, run_lvs=True, lvs_only=True)
# -
# ## AmpChain Schematic Template
# Now let's move on to schematic generator. As before, we need to create the schematic template first. A half-complete schematic template is provided for you in library `demo_templates`, cell `amp_chain`, shown below:
# <img src="bootcamp_pics/5_hierarchical_generator/hierachical_generator_2.PNG" alt="Drawing" style="width: 400px;"/>
#
# The schematic template for a hierarchical generator is very simple; you simply need to instantiate the schematic templates of the sub-blocks (***Not the generated schematic!***). For the exercise, instantiate the `amp_sf` schematic template from the `demo_templates` library, named it `XSF`, connect it, then evaluate the following cell to import the `amp_chain` netlist to Python.
#
# +
import bag
# obtain BagProject instance
local_dict = locals()
if 'bprj' in local_dict:
print('using existing BagProject')
bprj = local_dict['bprj']
else:
print('creating BagProject')
bprj = bag.BagProject()
print('importing netlist from virtuoso')
bprj.import_design_library('demo_templates')
print('netlist import done')
# -
# ## AmpChain Schematic Generator
# With schematic template done, you are ready to write the schematic generator. It is also very simple, you just need to call the `design()` method, which you implemented previously, on each instance in the schematic. Complete the following schematic generator, then evaluate the cell to push it through the design flow.
# +
import os
from bag.design import Module
# noinspection PyPep8Naming
class demo_templates__amp_chain(Module):
"""Module for library demo_templates cell amp_chain.
Fill in high level description here.
"""
# hard coded netlist flie path to get jupyter notebook working.
yaml_file = os.path.join(os.environ['BAG_WORK_DIR'], 'BAG_XBase_demo',
'BagModules', 'demo_templates', 'netlist_info', 'amp_chain.yaml')
def __init__(self, bag_config, parent=None, prj=None, **kwargs):
Module.__init__(self, bag_config, self.yaml_file, parent=parent, prj=prj, **kwargs)
@classmethod
def get_params_info(cls):
# type: () -> Dict[str, str]
"""Returns a dictionary from parameter names to descriptions.
Returns
-------
param_info : Optional[Dict[str, str]]
dictionary from parameter names to descriptions.
"""
return dict(
cs_params='common-source amplifier parameters dictionary.',
sf_params='source-follwer amplifier parameters dictionary.',
)
def design(self, cs_params=None, sf_params=None):
self.instances['XCS'].design(**cs_params)
# TODO: design XSF
import os
# import bag package
import bag
from bag.io import read_yaml
# import BAG demo Python modules
import xbase_demo.core as demo_core
from xbase_demo.demo_layout.core import AmpChainSoln
# load circuit specifications from file
spec_fname = os.path.join(os.environ['BAG_WORK_DIR'], 'specs_demo/demo.yaml')
top_specs = read_yaml(spec_fname)
# obtain BagProject instance
local_dict = locals()
if 'bprj' in local_dict:
print('using existing BagProject')
bprj = local_dict['bprj']
else:
print('creating BagProject')
bprj = bag.BagProject()
demo_core.run_flow(bprj, top_specs, 'amp_chain', AmpChainSoln, sch_cls=demo_templates__amp_chain, run_lvs=True)
| workspace_setup/tutorial_files/5_hierarchical_generators.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from Maze import Maze
from sarsa_agent import SarsaAgent
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import animation
from IPython.display import HTML
# ## Designing the maze
# +
arr=np.array([[0,1,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0],
[0,1,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0],
[0,1,0,0,1,0,0,1,1,1,1,1,0,1,1,0,1,1,1,0],
[0,1,0,0,1,0,0,0,0,0,1,0,0,1,0,0,1,0,0,0],
[0,0,0,0,1,0,0,1,1,1,0,0,1,1,0,0,1,0,0,0],
[0,0,0,0,0,0,0,1,0,0,0,1,0,1,0,1,1,0,1,1],
[1,1,1,0,1,1,0,1,0,0,1,0,0,1,0,0,1,0,0,0],
[0,0,1,0,1,0,1,0,0,1,0,0,0,0,0,0,1,0,1,0],
[0,0,0,0,1,0,0,0,1,0,0,0,0,0,0,1,1,0,1,0],
[0,0,0,0,1,0,0,1,0,0,0,0,0,1,1,1,0,0,0,0],
[1,0,1,1,1,0,1,0,0,1,0,0,0,1,0,0,0,1,0,0],
[1,0,1,1,1,0,1,0,0,1,0,0,1,1,0,0,0,1,0,0],
[1,0,0,1,0,0,1,0,0,1,0,0,1,0,0,1,0,0,0,0],
[0,0,0,0,1,0,1,0,0,1,1,0,1,0,0,0,1,1,1,0],
[0,0,1,1,1,0,1,0,0,1,0,1,0,0,1,1,0,0,0,0],
[0,1,1,0,0,0,0,1,0,1,0,0,1,1,0,1,0,1,1,1],
[0,1,0,0,0,1,0,0,0,0,0,0,1,0,0,1,0,0,0,0],
[0,0,1,1,1,0,1,1,0,0,1,0,1,0,0,1,1,0,0,0],
[1,0,0,0,1,0,1,0,0,0,1,0,1,0,0,1,1,1,0,0],
[1,1,0,0,0,0,1,0,0,0,1,0,1,0,0,0,0,1,0,0]
],dtype=float)
#Position of the rat
rat=(0,0)
#If Cheese is None, cheese is placed in the bottom-right cell of the maze
cheese=None
#The maze object takes the maze
maze=Maze(arr,rat,cheese)
maze.show_maze()
# -
# ## Defining a Agent [Sarsa Agent because it uses Sarsa to solve the maze]
agent=SarsaAgent(maze)
# ## Making the agent play episodes and learn
agent.learn(episodes=1000)
# ## Plotting the maze
nrow=maze.nrow
ncol=maze.ncol
fig=plt.figure()
ax=fig.gca()
ax.set_xticks(np.arange(0.5,ncol,1))
ax.set_yticks(np.arange(0.5,nrow,1))
ax.set_xticklabels([])
ax.set_yticklabels([])
ax.grid('on')
img=ax.imshow(maze.maze,cmap="gray",)
a=5
# ## Making Animation of the maze solution
def gen_func():
maze=Maze(arr,rat,cheese)
done=False
while not done:
row,col,_=maze.state
cell=(row,col)
action=agent.get_policy(cell)
maze.step(action)
done=maze.get_status()
yield maze.get_canvas()
def update_plot(canvas):
img.set_data(canvas)
anim=animation.FuncAnimation(fig,update_plot,gen_func)
HTML(anim.to_html5_video())
anim.save("big_maze.gif",animation.PillowWriter())
| run.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from bs4 import BeautifulSoup
import requests
class stock:
def __init__(self,*stock_num):
# +
from bs4 import BeautifulSoup
import requests
import pymysql
import openpyxl
from openpyxl.styles import Font
import gspread
from oauth2client.service_account import ServiceAccountCredentials
from selenium import webdriver
from selenium.webdriver.support.ui import Select
from webdriver_manager.chrome import ChromeDriverManager
import time
class Stock:
def __init__(self, *stock_numbers):
self.stock_numbers = stock_numbers
def scrape(self):
result = list()
for stock_number in self.stock_numbers:
response = requests.get(
"https://tw.stock.yahoo.com/q/q?s=" + stock_number)
soup = BeautifulSoup(response.text.replace("加到投資組合", ""), "lxml")
stock_date = soup.find(
"font", {"class": "tt"}).getText().strip()[-9:] # 資料日期
tables = soup.find_all("table")[2] # 取得網頁中第三個表格
tds = tables.find_all("td")[0:11] # 取得表格中1到10格
result.append((stock_date,) +
tuple(td.getText().strip() for td in tds))
return result
def save(self, stocks):
db_settings = {
"host": "127.0.0.1",
"port": 3306,
"user": "root",
"password": "******",
"db": "stock",
"charset": "utf8"
}
try:
conn = pymysql.connect(**db_settings)
with conn.cursor() as cursor:
sql = """INSERT INTO market(
market_date,
stock_name,
market_time,
final_price,
buy_price,
sell_price,
ups_and_downs,
lot,
yesterday_price,
opening_price,
highest_price,
lowest_price)
VALUES(%s, %s, %s, %s, %s, %s, %s, %s, %s, %s, %s, %s)"""
for stock in stocks:
cursor.execute(sql, stock)
conn.commit()
except Exception as ex:
print("Exception:", ex)
def export(self, stocks):
wb = openpyxl.Workbook()
sheet = wb.create_sheet("Yahoo股市", 0)
response = requests.get(
"https://tw.stock.yahoo.com/q/q?s=2451")
soup = BeautifulSoup(response.text, "lxml")
tables = soup.find_all("table")[2]
ths = tables.find_all("th")[0:11]
titles = ("資料日期",) + tuple(th.getText() for th in ths)
sheet.append(titles)
for index, stock in enumerate(stocks):
sheet.append(stock)
if "△" in stock[6]:
sheet.cell(row=index+2, column=7).font = Font(color='FF0000')
elif "▽" in stock[6]:
sheet.cell(row=index+2, column=7).font = Font(color='00A600')
wb.save("yahoostock.xlsx")
def gsheet(self, stocks):
scopes = ["https://spreadsheets.google.com/feeds"]
credentials = ServiceAccountCredentials.from_json_keyfile_name(
"credentials.json", scopes)
client = gspread.authorize(credentials)
sheet = client.open_by_key(
"YOUR GOOGLE SHEET KEY").sheet1
response = requests.get(
"https://tw.stock.yahoo.com/q/q?s=2451")
soup = BeautifulSoup(response.text, "lxml")
tables = soup.find_all("table")[2]
ths = tables.find_all("th")[0:11]
titles = ("資料日期",) + tuple(th.getText() for th in ths)
sheet.append_row(titles, 1)
for stock in stocks:
sheet.append_row(stock)
def daily(self, year, month):
browser = webdriver.Chrome(ChromeDriverManager().install())
browser.get(
"https://www.twse.com.tw/zh/page/trading/exchange/STOCK_DAY_AVG.html")
select_year = Select(browser.find_element_by_name("yy"))
select_year.select_by_value(year) # 選擇傳入的年份
select_month = Select(browser.find_element_by_name("mm"))
select_month.select_by_value(month) # 選擇傳入的月份
stockno = browser.find_element_by_name("stockNo") # 定位股票代碼輸入框
result = []
for stock_number in self.stock_numbers:
stockno.clear() # 清空股票代碼輸入框
stockno.send_keys(stock_number)
stockno.submit()
time.sleep(2)
soup = BeautifulSoup(browser.page_source, "lxml")
table = soup.find("table", {"id": "report-table"})
elements = table.find_all(
"td", {"class": "dt-head-center dt-body-center"})
data = (stock_number,) + tuple(element.getText()
for element in elements)
result.append(data)
print(result)
stock = Stock('2451', '2454', '2369') # 建立Stock物件
stock.daily("2019", "7") # 動態爬取指定的年月份中,股票代碼的每日收盤價
# stock.gsheet(stock.scrape()) # 將爬取的股票當日行情資料寫入Google Sheet工作表
# stock.export(stock.scrape()) # 將爬取的股票當日行情資料匯出成Excel檔案
# stock.save(stock.scrape()) # 將爬取的股票當日行情資料存入MySQL資料庫中
# -
| sideproject/yahoo stock.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import tomocg as tc
import deformcg as dc
import dxchange
data = np.load('diff_int_projs_corr.npy')
theta = np.linspace(np.deg2rad(-85),np.deg2rad(85),data.shape[0]).astype('float32')
data = np.delete(data,-1,axis=0)[20:-20]
theta = np.delete(theta,316)[20:-20]
ntheta = data.shape[0]
nz = data.shape[1]
n = data.shape[2]
nz
import tomopy as tp
rot_center = tp.find_center(data, theta)
prj_algn = tp.prep.alignment.align_joint(data, theta, iters=64, upsample_factor=100, center=rot_center, algorithm='sirt')
plt.figure()
plt.plot(prj_algn[2])
# %matplotlib notebook
plt.figure(figsize=(9,3))
plt.subplot(131)
plt.imshow(data[0])
plt.title('raw')
plt.subplot(132)
plt.imshow(prj_algn[0][0])
plt.title('aligned')
plt.subplot(133)
plt.imshow(prj_algn[0][0]-data[0])
plt.title('difference')
plt.show()
plt.savefig('diff_int_prj.png',dpi=300)
from skimage.feature import register_translation
shift = np.zeros([2,ntheta])
shift_algn = np.zeros([2,ntheta])
for i in range(1,ntheta):
shift[:,i] = register_translation(data[i-1,:], data[i], 1000, return_error=False)
shift_algn[:,i] = register_translation(prj_algn[0][i-1,:], prj_algn[0][i], 1000, return_error=False)
plt.figure(figsize=(9,3))
plt.subplot(121)
plt.plot(shift[0,:-1],'.-')
plt.plot(shift_algn[0,:-1],'.-')
plt.title('x-axis')
plt.legend(['raw','aligned'])
plt.subplot(122)
plt.plot(shift[1,:-1],'.-')
plt.plot(shift_algn[1,:-1],'.-')
plt.title('y-axis')
plt.legend(['raw','aligned'])
plt.savefig("shifts.png",dpi=300)
rec = tp.recon(data, theta, center=rot_center, algorithm='gridrec')
rec_algn = tp.recon(prj_algn[0], theta, center=rot_center, algorithm='gridrec')
plt.figure(figsize = (9,8))
plt.subplot(221)
plt.imshow(rec[nz//2])
plt.title('before alignment')
plt.subplot(222)
plt.imshow(rec_algn[nz//2])
plt.title('after alignment')
plt.subplot(223)
plt.imshow(rec[:,n//2])
plt.title('before alignment')
plt.subplot(224)
plt.imshow(rec_algn[:,n//2])
plt.title('after alignment')
plt.show()
plt.savefig('diff_int_rec.png',dpi=300)
vmf = np.zeros((ntheta,nz))
vmf1 = np.zeros((ntheta,nz))
for i in range(ntheta):
vmf[i] = np.sum(data[i],axis=1)
vmf1[i] = np.sum(prj_algn[0][i],axis=1)
plt.figure(figsize=(9,2))
plt.subplot(121)
plt.imshow(vmf.T)
plt.title('raw')
# plt.plot(62.5+shift[1,:-1]*100,'r.-')
plt.subplot(122)
plt.imshow(vmf1.T)
plt.title('aligned')
# plt.plot(62.5+shift_algn[1,:-1]*100,'r.-')
plt.savefig('vmf.png',dpi=300)
dxchange.write_tiff_stack(rec.astype('float32'), fname=u'rec_raw/data.tiff', overwrite=True)
| align/diff_int_tomo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Looking Through Tree-Ring Data in the Southwestern USA Using Pandas
#
# **Pandas** provides a useful tool for the analysis of tabular data in Python, where previously we would have had to use lists of lists, or use R.
## Bringing in necessary pckages
# %config InlineBackend.figure_format = 'svg'
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import scipy.stats.mstats as stat
#
# ### The Data
#
# The dataset I included herein is an example of tree growth rates at many different sites in the southwestern United States (AZ, NM). Field crews used increment borers to core into live trees in these sites and extract core samples. These were then brought back to the lab and dated to determine tree age and growth rates. I obtained this during my master's work at Northern Arizona University.
#
# In this dataset, each individual row is a tree.
# The columns are as follows:
# * site: The code for either the plot name or study site at which the tree was surveyed
# * center.date: The innermost ring of the tree. The closes estimate for the establishment year of the tree
# * dbh: The diameter of the tree (cm) at 1.37m above ground level
# * dsh: The diameter of the tree (cm) at 0.4m above ground level
# * Age: Estimated age of the tree when the core was collected. $Age = inv.yr-center.date$
# * spp: Four letter species code for the tree. The first two letters of the genus and species
# * inv.yr: The year in which the core was collected
# * BA: The basal area of the tree. Basically the surface area of the top of a stump if the tree was cut at 1.37m. Given with the formula $BA = 0.00007854 * DBH^2$
# * BA/Age: Just what it sounds like
# * Annual BAI: An estimate of the square centimeters of basal area produced by the tree each year. A better measure of growth than annual growth on the core as it accounts for tree size in addition to ring thickness in the core.
#
# Similar datasets are available through the International Tree Ring Data Bank (ITRDB), and can be found on the [ITRDB Webpage](https://data.noaa.gov/dataset/international-tree-ring-data-bank-itrdb)
#
# The following codeblock reads in the data and displays the first few rows of the pandas data frame. The path should be changed to the location of the .csv file.
## Change the path below if being run on a different computer
data = pd.read_csv(r"/Users/kyle/Google Drive/UC Boulder PhD Stuff/Classes/Fall 2016/Spatiotemporal Methods/Combined_BaiData.csv")
data.head()
print "There are cores for "+str(len(data))+" trees"
filtered_data = data.dropna()
print ("After removing rows with missing values, there are cores for "+str(len(filtered_data))+
" trees. \nSo, there were "+str(len(data)-len(filtered_data))+" rows that had NaN's")
# #### A logical question may be:
#
# *What species is growing the fastest across the sites?*
# So, we can produce a simple boxplot to help visualize this.
filtered_data.boxplot(column = 'annual.bai', by = 'spp')
# It appears that *Abies lasiocarpa* - subalpine fir - may be the fastest growing species overall. We can also look at the median values for the different species to verify this
filtered_data.groupby('spp', as_index=False)['annual.bai'].median()
## Adapted from http://stackoverflow.com/questions/35816865/create-vectors-for-kruskal-wallis-h-test-python
groupednumbers = {}
for grp in filtered_data['spp'].unique():
groupednumbers[grp] = filtered_data['annual.bai'][filtered_data['spp']==grp].values
args = groupednumbers.values()
h,p = stat.kruskalwallis(*args)
print("The Kruskal-Wallis H-statistic is: "+str(round(h,2))+" and... \nThe p-value is: "+str(p))
# So the species have very different growth rates. We could take this a step further and perform pairwise comparisons between groups using Mann-Whitney tests with Bonferroni correction for multiple comparison, but a more robust analysis would likely use mixed-effects models or partial regression to account for the different growing conditions between sites, and perhaps to account for age and tree size as additional covariates.
| Kyle Rodman's Lab 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import dependencies
import pandas as pd
import numpy as np
import scipy as sp
import matplotlib.pyplot as plt
import matplotlib
# %matplotlib inline
from datetime import datetime
from sqlalchemy import create_engine
# +
def parser(x):
return datetime.strptime('190'+x, '%Y-%m')
from datetime import datetime
series = pd.read_csv('data/FinalDataset.csv', header=0, squeeze=True)
station = series[series['Station']=='SFIA']
origin = station[['DateTime','Origin']]
origin = origin.set_index('DateTime')
print(origin.head())
plt.rcParams["figure.figsize"] = (20,7)
plt.rcParams['lines.linewidth'] = 1
plt.rcParams['axes.grid'] = True
origin.plot()
plt.savefig('images/daily_volume_origin_SFO.png')
plt.show()
# -
# # Daily Volume Predictions
origin.index = pd.to_datetime(origin.index)
pd.plotting.register_matplotlib_converters()
daily = origin.resample("D").sum()
daily.rolling(30, center=True).sum().plot(style=[":", "--", "-"])
plt.savefig('images/daily_volume_historical_SFIA.png')
# Use Prophet to predict
df2 = daily
df2.reset_index(inplace=True)
# Prophet requires columnds ds (Date) and y (value)
df2 = df2.rename(columns={'DateTime': 'ds', 'Origin': 'y'})
df2.head()
# Import Prophet
import fbprophet
# Make the Prophet model and fit the data
df2_prophet = fbprophet.Prophet(interval_width = .95)
df2_prophet.fit(df2)
# Create future dataframe
df2_forecast = df2_prophet.make_future_dataframe(periods=30*12, freq='D')
# Make predictions
df2_forecast = df2_prophet.predict(df2_forecast)
pd.plotting.register_matplotlib_converters()
df2_prophet.plot(df2_forecast, xlabel = 'Datetime', ylabel = 'Origin')
plt.title('Predictions')
plt.savefig('images/prediction1_SFIA.png')
# Plot the trends and patterns
df2_prophet.plot_components(df2_forecast);
plt.savefig('images/prediction2_SFIA.png')
# # Hourly Volume Predictions
hourly = origin.resample("H").sum()
hourly.rolling(24, center=True).sum().plot(style=[":", "--", "-"])
plt.savefig('images/Historical_hourly_SFIA.png')
# Use Prophet to predict
df3 = hourly
df3.reset_index(inplace=True)
# Prophet requires columnds ds (Date) and y (value)
df3 = df3.rename(columns={'DateTime': 'ds', 'Origin': 'y'})
df3.head()
# Make the Prophet model and fit the data
df3_prophet = fbprophet.Prophet(interval_width = .95)
df3_prophet.fit(df3)
# Create future dataframe
df3_forecast = df3_prophet.make_future_dataframe(periods=300*12, freq='H')
# Make predictions
df3_forecast = df3_prophet.predict(df3_forecast)
pd.plotting.register_matplotlib_converters()
df3_prophet.plot(df3_forecast, xlabel = 'Datetime', ylabel = 'Origin')
plt.title('Predictions')
plt.savefig('images/Predictions_hourly_SFIA.png')
# Plot the trends and patterns
df3_prophet.plot_components(df3_forecast);
plt.savefig('images/Trends_hourly_SFIA.png')
| .ipynb_checkpoints/ml_prophet-SFO-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # set environment variables
# ```bash
# export PROFILE=
# export BUCKET=
# export FILE_S3=
# export FILE_LOCAL=
# export DYNAMODB_TABLE_NAME=
# ```
import os
import pkg
# +
PROFILE = os.environ["PROFILE"]
handler = pkg.aws.Handler(profile=PROFILE)
# -
buckets = handler.s3_list_bucket()
assert "Buckets" in buckets, "Buckets not found"
# +
BUCKET = os.environ["BUCKET"]
FILE_S3 = os.environ["FILE_S3"]
response = handler.rekognition_detect_labels(bucket=BUCKET, file=FILE_S3)
assert response is not None, "No response"
# +
FILE_LOCAL = os.environ["FILE_LOCAL"]
response = handler.rekognition_detect_labels_from_local(file=FILE_LOCAL)
assert response is not None, "No response"
# +
DYNAMODB_TABLE_NAME = os.environ["DYNAMODB_TABLE_NAME"]
response = handler.dynamodb_create_table(table_name=DYNAMODB_TABLE_NAME)
assert response is not None, "No response"
# -
response = handler.dynamodb_put_item(table_name=DYNAMODB_TABLE_NAME)
assert response is not None, "No response"
response = handler.dynamodb_get_item(table_name=DYNAMODB_TABLE_NAME)
assert "Item" in response
| notebooks/aws.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
tf.test.is_built_with_cuda()
# -
physical_devices = tf.config.list_physical_devices('GPU')
print("Num GPUs:", len(physical_devices))
# +
tf.debugging.set_log_device_placement(True)
# Create some tensors
a = tf.constant([[1.0, 2.0, 3.0], [4.0, 5.0, 6.0]])
b = tf.constant([[1.0, 2.0], [3.0, 4.0], [5.0, 6.0]])
c = tf.matmul(a, b)
print(c)
# -
| tensorflow_sandbox/check_install.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="3rqsGAHiVx-n"
# # React FFC 03 List & Spread Operator
#
#
# **References**
# - freeCodeCamp.org (2020) Full React Course 2020 - Learn Fundamentals, Hooks, Context API, React https://www.youtube.com/watch?v=4UZrsTqkcW4&ab_channel=freeCodeCamp.org
# - Tutorial by Mr. <NAME>
# + [markdown] id="tSBCdUsSlR8W"
# ## List
# If you have multiple instances to furnish a page with a list structure, make your instances into a list of variables and display them using a loop function.
# + id="Pe1evGVdR0FM"
from IPython.display import HTML, display
def set_css():
display(HTML('''<style> pre { white-space: pre-wrap; }</style>'''))
get_ipython().events.register('pre_run_cell', set_css)
# + id="J9NuQCHxC5rp"
Javascript("""
// Look at the booklist, we are repeating the recurrent components (book instances)
function Booklist() {
return <section className='booklist'>
<Book img={Book1.img} title={Book1.title} author={Book1.author}/>
<Book img={Book2.img} title={Book2.title} author={Book2.author}/>
<Book img={Book3.img} title={Book3.title} author={Book3.author}/>
<Book img={Book4.img} title={Book4.title} author={Book4.author}/>
<Book img={Book5.img} title={Book5.title} author={Book5.author}/>
<Book img={Book6.img} title={Book6.title} author={Book6.author}/>
<Book img={Book7.img} title={Book7.title} author={Book7.author}/>
<Book img={Book8.img} title={Book8.title} author={Book8.author}>
<p><NAME> is a top selling author of children’s books with over 65 million books sold worldwide...
</p>
</Book>
</section>
}
""")
# + id="ngTkJc9-C9C7" colab={"base_uri": "https://localhost:8080/", "height": 901} outputId="cdbd3129-9bc4-486b-e924-50eb2c256853"
# Using a list of instances
Javascript("""
console.log(newNames);
function Booklist() {
return <section className='booklist'>
{books.map((book) => {
const {img, title, author} = book;
return <div>
<h3>{title}</h3>
<h5>{author}</h5>
</div>;
})}</section>;
}
""")
# + [markdown] id="cbQE7cvkwLJB"
# ### Rendering a list
# + id="ba9Dvup3C93h" colab={"base_uri": "https://localhost:8080/", "height": 544} outputId="ecb1e2a2-e1e8-4287-e4d0-dc3fe3b3fc75"
Javascript("""
function Booklist() {
return (
<section className='booklist'>
{books.map((book) => {
const {img, title, author} = book; // pass book object as a prop.
return (
<Book book={book}></Book>
);
})}
</section>
);
}
const Book = (props) => {
const { img, title, author } = props.book;
return (
<article className='book'>
<img src={img} alt='' />
<h1>{title}</h1>
<h4>{author}</h4>
</article>
);
};
ReactDom.render(<Booklist/>, document.getElementById('root'));
""")
# + [markdown] id="Bvijli8B0S88"
# ## `...Spread Operator`
# + id="VwrVq5Th0WPW"
# before spreading
Javascript("""
function Booklist() {
return (
<section className='booklist'>
{books.map((book) => {
const {img, title, author, link} = book; // pass book object as a prop.
return (
<Book key={book.id} book={book}></Book>
);
})}
</section>
);
}
""")
# + id="fJfV-4PfC5hg"
# after spreading
Javascript("""
function Booklist() {
return (
<section className='booklist'>
{books.map((book) => {
return <Book key={book.id} {...book}></Book>;
})}
</section>
);
}
const Book = ({ img, title, author, link }) => {
return (
<article className='book'>
<a href={link} rel="noreferrer" target="_blank">
<img src={img} alt='' />
<h1>{title}</h1>
<h4>{author}</h4>
</a>
</article>
);
};
""")
| 0 Fundamentals/React FCC 03 List & Spread Operator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Logistic regression
#
# We want to minimize
# $$\min_{x\in {\mathbb R}^{d}} f(x)=-\frac{1}{n}\sum_{i=1}^n \left(y_i \log (s(a_i^\top x)) + (1 - y_i) \log (1 - s(a_i^\top x))\right) + \frac{\gamma}{2}\|x\|^2,$$
# where $a_i\in {\mathbb R}^{d}$, $y_i\in \{0, 1\}$, $s(z)=\frac{1}{1+\exp(-z)}$ is the sigmoid function.
# The gradient is given by $\nabla f(x) = \frac{1}{n}\sum_{i=1}^n a_i(s(a_i^\top x)-y_i) + \gamma x$.
#
# This is a smooth function with smoothness constant $L=\frac{1}{4}\lambda_{\max}(A^\top A) + \gamma$, where $\lambda_{\max}$ denotes the largest eigenvalue.
#
# For this experiment we used `w8a` dataset from the [LIBSVM](https://www.csie.ntu.edu.tw/~cjlin/libsvm/) library.
# +
import matplotlib
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import numpy.linalg as la
from sklearn.datasets import load_svmlight_file
from optimizers import Adgd, AdgdAccel, Armijo, NestLine
from loss_functions import logistic_loss, logistic_gradient
sns.set(style="whitegrid", font_scale=1.2, context="talk", palette=sns.color_palette("bright"), color_codes=False)
matplotlib.rcParams['mathtext.fontset'] = 'cm'
dataset = 'w8a'
data_path = './datasets/' + dataset
if dataset == 'covtype':
data_path += '.bz2'
if dataset == 'covtype':
it_max = 10000
elif dataset == 'w8a':
it_max = 8000
else:
it_max = 3000
def logistic_smoothness(X):
return 0.25 * np.max(la.eigvalsh(X.T @ X / X.shape[0]))
data = load_svmlight_file(data_path)
X, y = data[0].toarray(), data[1]
if (np.unique(y) == [1, 2]).all():
# Loss functions support only labels from {0, 1}
y -= 1
if (np.unique(y) == [-1, 1]).all():
y = (y+1) / 2
n, d = X.shape
L = logistic_smoothness(X)
l2 = L / n if dataset == 'covtype' else L / (10 * n)
w0 = np.zeros(d)
def loss_func(w):
return logistic_loss(w, X, y, l2)
def grad_func(w):
return logistic_gradient(w, X, y, l2)
# -
ar = Armijo(lr0=1 / L, loss_func=loss_func, grad_func=grad_func, it_max=it_max)
ar.run(w0=w0)
nest_ls = NestLine(lr0=1 / L, mu=0, loss_func=loss_func, grad_func=grad_func, it_max=it_max)
nest_ls.run(w0=w0)
adgd = Adgd(loss_func=loss_func, grad_func=grad_func, eps=0, it_max=it_max)
adgd.run(w0=w0)
ad_acc = AdgdAccel(loss_func=loss_func, grad_func=grad_func, it_max=it_max)
ad_acc.run(w0=w0)
# +
optimizers = [ar, nest_ls, adgd, ad_acc]
markers = [',', 'o', '*', '^', 'D', 's', '.', 'X']
for opt, marker in zip(optimizers, markers):
opt.compute_loss_on_iterates()
f_star = np.min([np.min(opt.losses) for opt in optimizers])
plt.figure(figsize=(8, 6))
labels = ['Armijo', 'Nesterov-LS', 'AdGD', 'AdGD-accel']
for opt, marker, label in zip(optimizers, markers + ['.', 'X'], labels):
opt.plot_losses(marker=marker, f_star=f_star, label=label)
plt.yscale('log')
plt.xlabel('# matrix-vector multiplications')
plt.ylabel(r'$f(x^k) - f_*$')
plt.legend()
plt.tight_layout()
plt.show()
# -
| linesearch_logistic_regression_w8a.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from datetime import datetime
import pandas as pd
from scipy import optimize
from scipy import integrate
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(style="darkgrid")
mpl.rcParams['figure.figsize'] = (16, 9)
pd.set_option('display.max_rows', 500)
# -
df_analyse=pd.read_csv('../data/processed/COVID_small_flat_table.csv',sep=';')
df_analyse.sort_values('date',ascending=True).head()
# +
# set some basic parameters
N0=1000000
beta=0.4
gamma=0.1
# condition = I0+S0+R0=N0
I0=df_analyse.Germany[35]
S0=N0-I0
R0=0
# -
df_analyse.Germany[35]
def SIR_model(SIR,beta,gamma):
''' Simple SIR model
S: susceptible population
I: infected people
R: recovered people
beta:
overall condition is that the sum of changes (differnces) sum up to 0
dS+dI+dR=0
S+I+R= N (constant size of population)
'''
S,I,R=SIR
dS_dt=-beta*S*I/N0
dI_dt=beta*S*I/N0-gamma*I
dR_dt=gamma*I
return([dS_dt,dI_dt,dR_dt])
# # Simulative approach to calculate SIR curves
# +
SIR=np.array([S0,I0,R0])
propagation_rates=pd.DataFrame(columns={'susceptible':S0,
'infected':I0,
'recoverd':R0})
for each_t in np.arange(100):
new_delta_vec=SIR_model(SIR,beta,gamma)
SIR=SIR+new_delta_vec
propagation_rates=propagation_rates.append({'susceptible':SIR[0],
'infected':SIR[1],
'recovered':SIR[2]}, ignore_index=True)
# +
fig, ax1 = plt.subplots(1, 1)
ax1.plot(propagation_rates.index,propagation_rates.infected,label='infected',color='k')
ax1.plot(propagation_rates.index,propagation_rates.recovered,label='recovered')
ax1.plot(propagation_rates.index,propagation_rates.susceptible,label='susceptible')
ax1.set_ylim(10, 1000000)
ax1.set_yscale('linear')
ax1.set_title('Szenario SIR simulations (demonstration purposes only)',size=16)
ax1.set_xlabel('time in days',size=16)
ax1.legend(loc='best',
prop={'size': 16});
# -
# # Fitting the parameters of SIR model
ydata = np.array(df_analyse.Germany[35:])
t=np.arange(len(ydata))
I0=ydata[0]
S0=N0-I0
R0=0
beta
def SIR_model_t(SIR,t,beta,gamma):
''' Simple SIR model
S: susceptible population
t: time step, mandatory for integral.odeint
I: infected people
R: recovered people
beta:
overall condition is that the sum of changes (differnces) sum up to 0
dS+dI+dR=0
S+I+R= N (constant size of population)
'''
S,I,R=SIR
dS_dt=-beta*S*I/N0
dI_dt=beta*S*I/N0-gamma*I
dR_dt=gamma*I
return dS_dt,dI_dt,dR_dt
def fit_odeint(x, beta, gamma):
'''
helper function for the integration
'''
return integrate.odeint(SIR_model_t, (S0, I0, R0), t, args=(beta, gamma))[:,1]
popt=[0.4,0.1]
fit_odeint(t, *popt)
# +
popt, pcov = optimize.curve_fit(fit_odeint, t, ydata)
perr = np.sqrt(np.diag(pcov))
print('standard deviation errors : ',str(perr), ' start infect:',ydata[0])
print("Optimal parameters: beta =", popt[0], " and gamma = ", popt[1])
# -
# Gettng the Final fitted curve
fitted=fit_odeint(t, *popt)
plt.semilogy(t, ydata, 'o')
plt.semilogy(t, fitted)
plt.title("Fit of SIR model for Germany cases")
plt.ylabel("Population infected")
plt.xlabel("Days")
plt.show()
print("Optimal parameters: beta =", popt[0], " and gamma = ", popt[1])
print("Basic Reproduction Number R0 " , popt[0]/ popt[1])
print("This ratio is derived as the expected number of new infections (these new infections are sometimes called secondary infections from a single infection in a population where all subjects are susceptible. @wiki")
# # Dynamic beta in SIR (infection rate)
t_initial=35
t_intro_measures=20
t_hold=50
t_relax=55
beta_max=0.3
beta_min=0.1
gamma=0.1
pd_beta=np.concatenate((np.array(t_initial*[beta_max]),
np.linspace(beta_max,beta_min,t_intro_measures),
np.array(t_hold*[beta_min]),
np.linspace(beta_min,beta_max,t_relax),
))
pd_beta
# +
SIR=np.array([S0,I0,R0])
propagation_rates=pd.DataFrame(columns={'susceptible':S0,
'infected':I0,
'recoverd':R0})
for each_beta in pd_beta:
new_delta_vec=SIR_model(SIR,each_beta,gamma)
SIR=SIR+new_delta_vec
propagation_rates=propagation_rates.append({'susceptible':SIR[0], 'infected':SIR[1], 'recovered':SIR[2]}, ignore_index=True)
# +
fig, ax1 = plt.subplots(1, 1)
ax1.plot(propagation_rates.index,propagation_rates.infected,label='infected',linewidth=3)
t_phases=np.array([t_initial,t_intro_measures,t_hold,t_relax]).cumsum()
ax1.bar(np.arange(len(ydata)),ydata, width=0.8,label=' current infected Germany',color='r')
ax1.axvspan(0,t_phases[0], facecolor='b', alpha=0.2,label='no measures')
ax1.axvspan(t_phases[0],t_phases[1], facecolor='b', alpha=0.3,label='hard measures introduced')
ax1.axvspan(t_phases[1],t_phases[2], facecolor='b', alpha=0.4,label='hold measures')
ax1.axvspan(t_phases[2],t_phases[3], facecolor='b', alpha=0.5,label='relax measures')
ax1.axvspan(t_phases[3],len(propagation_rates.infected), facecolor='b', alpha=0.6,label='repead hard measures')
ax1.set_ylim(10, 1.5*max(propagation_rates.infected))
ax1.set_yscale('log')
ax1.set_title('Szenario SIR simulations (demonstration purposes only)',size=16)
ax1.set_xlabel('time in days',size=16)
ax1.legend(loc='best',
prop={'size': 16});
# -
| notebooks/SIR_modeling_Joshua.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Day1
# ## Problem Statement
# On a particular date, If 70% number of confirmed case is zero, then Delete the column. i.e. whole February will be deleted and few more.
# Plot the graph
# - Country Wise
# - Date Wise
# - Continent Wise
# Dataset - https://data.humdata.org/hxlproxy/api/data-preview.csv?url=https%3A%2F%2Fraw.githubusercontent.com%2FCSSEGISandData%2FCOVID-19%2Fmaster%2Fcsse_covid_19_data%2Fcsse_covid_19_time_series%2Ftime_series_19-covid-Confirmed.csv&filename=time_series_2019-ncov-Confirmed.csv
# ## Importing Heavenly Bodies
#Calling the Gods
import pandas as pd
# %matplotlib notebook
import matplotlib.pyplot as plt
import matplotlib
import numpy as np
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
# Plot configurations
font = {'family' : 'normal',
'weight' : 'bold',
'size' : 7}
matplotlib.rc('font', **font)
# ## Loading the data and Extracting Desired columns
data = pd.read_csv("input/time_series_2019-ncov-Confirmed.csv")
data.head()
data = data.drop(columns = ['Province/State','Lat','Long'])
data2 = data.copy()
# +
#checking for 70% constraint
check = data.eq(0).sum().lt(341)
droplist = []
for col,val in check.iteritems():
if val==False:
droplist.append(col)
data = data.drop(columns = droplist)
# -
data.head()
# ## Grouping and adding dataframes based on Countries
# There are alot of Provinces/State of same countries, hence combining those to one.
df = data.groupby(['Country/Region']).sum().reset_index()
df.head()
# Full Dataset
data2 = data2.groupby(['Country/Region']).sum().reset_index()
data2.head()
# ## Countrywise plot
#Never work on original dataset, always create a copy and work
only_date_df = df.copy()
#Creating a dict for selection
column_list = list(only_date_df.columns)
mydate= {}
for i in column_list[1:]:
if i not in mydate.keys():
mydate[i] = i
#Function to plot a graph
def plot_country(date,n):
df = only_date_df.copy()
df = df.nlargest(n,date)
x = df['Country/Region'].tolist()
y = df[date].tolist()
plt.figure(figsize=(9,5))
for index, value in enumerate(y):
# print(index,value)
plt.text(value, index, str(value))
plt.barh(x,y)
n_dict = {'10':10,'20':20,'40':40}
# Here n represents top n countries infected the most, and date represents the date you want
# It is current set to 70% constraint
interact(plot_country,n = n_dict,date = mydate)
# ## Date wise plots
#
for_spec_coun_full = data2.copy()
for_spec_coun = df.copy()
mycountry= {}
country_list = for_spec_coun['Country/Region']
for i in country_list:
if i not in mycountry.keys():
mycountry[i] = i
# +
def plot_date(country,my_duration):
if my_duration == 0:
value = for_spec_coun.loc[for_spec_coun['Country/Region'] == country]
else:
value = for_spec_coun_full.loc[for_spec_coun_full['Country/Region'] == country]
x = list(value.values.flatten())[1:]
y = list(value.columns)[1:]
plt.figure(figsize=(9,5))
plt.xticks(rotation=90)
plt.plot(y,x)
# my_duration allows you to add or remove the 70% constaint and view the graph.
# you can select the country of your own choice..
my_duration={'less than 70':0,'full dataset':1}
interact(plot_date,country = country_list,my_duration = my_duration)
# -
# ## Continent wise
continent_data = pd.read_csv("input/continents.csv")
for_cont = data2.copy()
continent_data.head()
new_data = pd.merge(right=for_cont, left=continent_data, right_on='Country/Region', left_on='Country')
conti = new_data['Continent'].unique()
new_data.head()
column_list2 = list(new_data.columns)
mydate2= {}
for i in column_list2[3:]:
if i not in mydate2.keys():
mydate2[i] = i
# +
def plot_continent(date):
y = []
for c in conti:
y.append(new_data.loc[new_data['Continent'] == c][date].sum())
plt.figure(figsize=(9,5))
plt.barh(conti,y)
plt.xticks(rotation=90)
for index, value in enumerate(y):
plt.text(value, index, str(value))
interact(plot_continent,date = mydate2)
# -
# ## Acknowledgement
#
# 1. Continents Dataset - https://github.com/AbdulRashidReshamwala/10DayesOfML/blob/master/day_01/continents.csv
| Day1/.ipynb_checkpoints/Covid19_Dashboard-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Condensatore di Epino (II turno)
#
# Componenti del gruppo:
#
# * <NAME>
# * <NAME> 1141482
#
# **Premessa**
#
# Nel seguito verrà utilizzata la funzione `scipy.optimyze.curve_fit()` per stimare i parametri delle relazioni funzionale. Questo metodo implementa la regressione ai minimi quadrati insenso classico e restituisce, oltre ai parametri richiesti, la matrice delle covarianze in cui la diagonale contiene i valori della varianza dei parametri stimati.
#
# per la gestione dei dati verrà utilizzata la librearia `pandas`, per i calcoli e la rappresenzione verranno utilizzati `numpy` e `matplotlib.pyplot` rispettivamente.
#
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy.optimize import curve_fit
from scipy.constants import epsilon_0 as e0
# ## Risultati prima parte
# $$ tot = segnale - fondo $$
#
# calcolo del segnale netto
#
# possibili errori sistemici
df=pd.read_csv('data1parte.csv', header=0)
# carica totale
segnale, fondo = df.values[:,1], df.values[:,3]
q = segnale -fondo
volt = df.values[:,0]
df
# ## Interpretazione e commenti prima parte
#
# fit relazione :
# $$ Q = q_0 + C V $$
#
# in un condensatore piano indefinito:
# $$ C = \frac{ \Sigma \epsilon _0}{ d} $$
# +
#funzione fit
def Q(v,qo,c):
return qo + c*v
# ottimizzazione
param, covar = curve_fit(Q, volt, q)
qo, C = param
er_q, er_c = np.diag(covar)**.5
# creazione grafico
fig = plt.figure(dpi=200)
ax = fig.add_subplot(111)
plt.plot(volt, Q(volt, *param), 'r-', label='fit')
plt.plot(volt, q, marker = '+', linestyle= ':', label='data')
plt.title('scansione in ddp')
ax.set_ylabel('$carica (nC)$')
ax.set_xlabel('$ddp (V)$')
plt.legend(); plt.grid();plt.show()
# log
print('\nqo: {:.3f} +-{:.3f}nC'.format(qo, er_q) )
print('\nC: {:.3f} +-{:.3f}pF'.format(C, er_c))
#stima distanza iniziale, Area = 0.049
d0 = 0.049*e0/(C*10**(-12))
print('\ndistanza iniziale: {0:.3f}mm'.format(d0))
# -
# ## Risultati seconda parte
# * ddp in scansione: $ \Delta V = 60V $
# * passo della vite: $ passo = 1.5 \frac{mm}{giro} $
# * calcolo spostamenti relativi e carica totale
# $$ carica = segnale - fondo $$
df=pd.read_csv('data2parte.csv', header=0)
# voltaggio (V), passo (mm/giro)
volt, passo = 60, 1.5
# spostamento relativo d - d0 (mm)
spos_rel = df.values[:,0]*passo
# spostamento assoluto d (mm)
spos = spos_rel + d0
# carica q (nC)
q = df.values[:,1]-df.values[:,2]
new = {
'giri': df.values[:,0],
'd': spos,
'd-d0': spos_rel,
'segnale': df.values[:,1],
'fondo': df.values[:,2],
'q': q
}
pd.DataFrame(data=new)
# ## Interpretazione e commenti terza parte
#
# $$ Q(d) = \epsilon_0 \frac{ A}{ d+d_0} V + B $$
#
# dove:
#
# * $B$ è una costante di integrazione per compensare errori sistemici
# * $A = 0.049 m^2$ è l'area del piatto del condensatore
# +
# funzione da stimare
def Q(x,eo, do, B ):
# dati Area = 0.049, V= 60
return eo*60*0.049/(x+do) + B
# ottimizzazione
popt, pcov = curve_fit(Q, spos, q)
eo, do, B = popt
er_e, er_d, er_B = np.diag(pcov)**0.5
# creazione grafico
fig = plt.figure(dpi=200)
ax = fig.add_subplot(111)
plt.plot(spos, Q(spos, *popt), 'r-', label='fit')
plt.plot(spos, q, marker = '+', linestyle= ':', label='data')
# formattazione
plt.title('scansione in posizione')
ax.set_ylabel('$carica (nC)$')
ax.set_xlabel('$spostamento (mm)$')
plt.legend(); plt.grid();plt.show()
# parametri di ottimizzazione
print('\n eo:{:.3f} +-{:.3f}pF/m'.format(eo, er_e))
print('\n do:{:.3f} +-{:.3f}mm'.format(do, er_d))
print('\n B:{:.3f} +-{:.3f}nC'.format(B, er_B))
# -
# # Conclusioni
#
# * valore di e0
# * stima della distanza iniziale
# * validità dell'approssimazioni di condensatore indefinito
# # Bibliografia
# * Fisica in laborario Esculapio
# %load_ext version_information
# %version_information numpy, scipy, matplotlib, version_information
| 1esp/schio_demarchi_esperienza1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from __future__ import print_function
import os
import numpy as np
import pandas as pd
from matplotlib.patches import Rectangle as rect
import matplotlib.pyplot as plt
# ## Model background
# Here is an example based on the model of Freyberg, 1988. The synthetic model is a 2-dimensional MODFLOW model with 1 layer, 40 rows, and 20 columns. The model has 2 stress periods: an initial steady-state stress period used for calibration, and a 5-year transient stress period. The calibration period uses the recharge and well flux of Freyberg, 1988; the last stress period use 25% less recharge and 25% more pumping.
#
# The inverse problem has 761 parameters: hydraulic conductivity of each active model cell, calibration and forecast period recharge multipliers, storage and specific yield, calibration and forecast well flux for each of the six wells, and river bed conductance for each 40 cells with river-type boundary conditions. The inverse problem has 12 head obseravtions, measured at the end of the steady-state calibration period. The forecasts of interest include the sw-gw exchange flux during both stress periods (observations named ```sw_gw_0``` and ``sw_gw_1``), and the water level in well cell 6 located in at row 28 column 5 at the end of the stress periods (observations named ```or28c05_0``` and ```or28c05_1```). The forecasts are included in the Jacobian matrix as zero-weight observations. The model files, pest control file and previously-calculated jacobian matrix are in the `freyberg/` folder
#
#
# Freyberg, <NAME>. "AN EXERCISE IN GROUND‐WATER MODEL CALIBRATION AND PREDICTION." Groundwater 26.3 (1988): 350-360.
# +
import flopy
# load the model
model_ws = os.path.join("Freyberg","extra_crispy")
ml = flopy.modflow.Modflow.load("freyberg.nam",model_ws=model_ws)
# plot some model attributes
fig = plt.figure(figsize=(6,6))
ax = plt.subplot(111,aspect="equal")
ml.upw.hk.plot(axes=[ax],colorbar="K m/d",alpha=0.3)
ml.wel.stress_period_data.plot(axes=[ax])
ml.riv.stress_period_data.plot(axes=[ax])
# plot obs locations
obs = pd.read_csv(os.path.join("freyberg","misc","obs_rowcol.dat"),delim_whitespace=True)
obs_x = [ml.dis.sr.xcentergrid[r-1,c-1] for r,c in obs.loc[:,["row","col"]].values]
obs_y = [ml.dis.sr.ycentergrid[r-1,c-1] for r,c in obs.loc[:,["row","col"]].values]
ax.scatter(obs_x,obs_y,marker='.',label="obs")
#plot names on the pumping well locations
wel_data = ml.wel.stress_period_data[0]
wel_x = ml.dis.sr.xcentergrid[wel_data["i"],wel_data["j"]]
wel_y = ml.dis.sr.ycentergrid[wel_data["i"],wel_data["j"]]
for i,(x,y) in enumerate(zip(wel_x,wel_y)):
ax.text(x,y,"{0}".format(i+1),ha="center",va="center")
ax.set_ylabel("y")
ax.set_xlabel("x")
ax.add_patch(rect((0,0),0,0,label="well",ec="none",fc="r"))
ax.add_patch(rect((0,0),0,0,label="river",ec="none",fc="g"))
ax.legend(bbox_to_anchor=(1.5,1.0),frameon=False)
plt.savefig("domain.pdf")
# -
# The plot shows the Freyberg (1988) model domain. The colorflood is the hydraulic conductivity ($\frac{m}{d}$). Red and green cells coorespond to well-type and river-type boundary conditions. Blue dots indicate the locations of water levels used for calibration.
# ## Using `pyemu`
import pyemu
# First create a linear analysis object. We will use `MonteCarlo` derived type, which allows us to use some sampling based methods. We pass it the name of the jacobian matrix file. Since we don't pass an explicit argument for `parcov` or `obscov`, `pyemu` attempts to build them from the parameter bounds and observation weights in a pest control file (.pst) with the same base case name as the jacobian. Since we are interested in forecast uncertainty as well as parameter uncertainty, we also pass the names of the forecast sensitivity vectors we are interested in, which are stored in the jacobian as well. Note that the `forecasts` argument can be a mixed list of observation names, other jacobian files or PEST-compatible ASCII matrix files.
# get the list of forecast names from the pest++ argument in the pest control file
jco = os.path.join("freyberg","freyberg.jcb")
pst = pyemu.Pst(jco.replace("jcb","pst"))
mc = pyemu.MonteCarlo(jco=jco, forecasts=pst.pestpp_options["forecasts"].split(','),verbose=False)
print("observations,parameters in jacobian:",mc.jco.shape)
# ## Drawing from the prior
# Each ``MonteCarlo`` instance has a ``parensemble`` attribute which itself is an instance of ``Ensemble`` class, which is derived from ``pandas.DataFrame``. What all that means is that the parameter ensembles behave just like ```DataFrame```s
#
# ### ```draw```
# The ``draw`` method is the main entry point into getting realizations. It accepts several optional arguments. Without any args, it makes a single draw from the prior, which uses a $\boldsymbol{\mu}$ (mean) vector of the parameter values listed in the pest control file:
#
mc.draw()
print(mc.parensemble.shape)
# ``draw`` also accepts a ``num_reals`` argument to specify the number of draws to make:
mc.draw(num_reals=200)
print(mc.parensemble.shape)
print(mc.parensemble.mean().head())
# Notice that each call to ``draw`` overwrites the previous draws. ```draw``` also accepts a ``par_file`` argument in the case that you want to use a pest .par file as the $\boldsymbol{\mu}$ vector.
mc.draw(num_reals=200,par_file=jco.replace(".jcb",".par"))
print(mc.parensemble.mean().head())
# Notice how the mean value for ``rond00`` is different. ``draw`` also accepts an ``obs`` boolean flag to control include drawing a realization of observation noise. If ```obs``` is True, then a complimentary ```obsensemble``` attribute is also populated. The last optional flag for ```draw``` is ``enforce_bounds``, which controls whether parameter bounds are explicitly respected:
# ```.draw``` also accepts an optional ``how`` argument that controls the type of distribution to draw from. ``how`` can be either "gaussian" (default) or "uniform".
# ## plotting
#
# Since ```ParameterEnsemble``` is dervied from ```pandas.DataFrame```, it has all the cool methods and attributes we all love. Let's compare the results of drawing from a uniform vs a gaussian distribution. This may take some time.
mc.draw(num_reals=500,how="uniform")
ax = plt.subplot(111)
mc.parensemble.loc[:,"rch_1"].plot(kind="hist",bins=20,ax=ax,alpha=0.5)
mc.draw(num_reals=500,how="gaussian")
mc.parensemble.loc[:,"rch_1"].plot(kind="hist",bins=20,ax=ax,alpha=0.5)
# ## null-space projection
#
# This is too easy. Once you have drawn parameter realization, use the ```project_parensemble()``` method. This method accepts 3 optional arguemnts: ``nsing``: number of singular components to demarcate the solution-null space boundary, ``par_file``: a pest .par file to use as the final parameter values, and ``inplace``, which is a boolean flag to control whether a new ```Ensemble``` instance should be created and returned. The most important of these is ``nsing``. If it is not passed, then ``nsing`` is set based on the ratio between the largest and smallest singular values >= 1.0e-6
mc.draw(num_reals=1000,enforce_bounds=True)
ax = plt.subplot(111)
mc.parensemble.loc[:,"rch_1"].plot(kind="hist",bins=20,ax=ax,alpha=0.5)
mc.project_parensemble(nsing=50,par_file=jco.replace(".jcb",".par")) #use nsing=50 for demonstration purposes
mc.parensemble.loc[:,"rch_1"].plot(kind="hist",bins=20,ax=ax,alpha=0.5)
# We see that if we use a large number of singular components, then the null-space projection process greatly reduces the uncertainty in the ``rch_1`` parameter. Note that using 50 singular components greatly overeastimates the dimension of the range space of the normal matrix ($\mathbf{J}^T\mathbf{Q}\mathbf{J}$) and is likely not justifiable, since only 12 observations are being used for inversion. Let's redo the redo the null-space projection operation with 12 singular components:
mc.draw(num_reals=1000,enforce_bounds=True)
mc.parensemble.to_csv("prior.csv")
ax = plt.subplot(111)
mc.parensemble.loc[:,"rch_1"].plot(kind="hist",bins=20,ax=ax,alpha=0.5)
en = mc.project_parensemble(nsing=12,par_file=jco.replace(".jcb",".par"),inplace=False) #use nsing=50 for demonstration purposes
#mc.parensemble.loc[:,"rch_1"].plot(kind="hist",bins=20,ax=ax,alpha=0.5)
en.to_csv("proj.csv")
en.loc[:,"rch_1"].plot(kind="hist",bins=20,ax=ax,alpha=0.5)
# Now we see that the null-space projection operation only slightly increases the kurtosis of the distribution
#
# ## output
# Once the desired type of ensemble has been generated, ``pyemu`` offers several ways to output the ensemble. One option is to write a pest control file for each realization:
mc.draw(num_reals=10,enforce_bounds=True)
mc.project_parensemble()
mc.write_psts(jco.replace(".jcb","_real"),existing_jco="freyberg.jcb",noptmax=1)
# or, optionally, write pest .par files:
mc.parensemble.to_parfiles(jco.replace(".jcb","_par_real"))
# or, since the ``pyemu.Ensemble`` class is derived from the ``pandas.DataFrame``, we can also save the ensemble using any of the highly-optimized ``DataFrame`` output methods, such as ``.to_csv()``:
mc.parensemble.to_csv(jco+".csv")
# ## running the ensemble
#
# This is the hard part - typically, evaluating the realizations will require some form of high-throughput, distributed computing. This is why FOSM is so great!
| examples/NSMCexample_freyberg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={} tags=[]
# # Github - Get commits ranking from repository
# <a href="https://app.naas.ai/user-redirect/naas/downloader?url=https://raw.githubusercontent.com/jupyter-naas/awesome-notebooks/master/Github/Github_Get_commits_ranking_from_repository.ipynb" target="_parent"><img src="https://naasai-public.s3.eu-west-3.amazonaws.com/open_in_naas.svg"/></a>
# + [markdown] papermill={} tags=[]
# **Tags:** #github #commits #stats #naas_drivers #plotly #linechart
# + [markdown] papermill={} tags=[]
# ## Input
# + papermill={} tags=[]
import pandas as pd
import plotly.express as px
from naas_drivers import github
import naas
# + [markdown] papermill={} tags=[]
# ## Setup Github
# **How to find your personal access token on Github?**
#
# - First we need to create a personal access token to get the details of our organization from here: https://github.com/settings/tokens
# - You will be asked to select scopes for the token. Which scopes you choose will determine what information and actions you will be able to perform against the API.
# - You should be careful with the ones prefixed with write:, delete: and admin: as these might be quite destructive.
# - You can find description of each scope in docs here (https://docs.github.com/en/developers/apps/building-oauth-apps/scopes-for-oauth-apps).
# + papermill={} tags=[]
# Github repository url
REPO_URL = "https://github.com/jupyter-naas/awesome-notebooks"
# Github token
GITHUB_TOKEN = "ghp_fUYP0Z5i29AG4ggX8owctGnHUoVHi******"
# + [markdown] papermill={} tags=[]
# ## Model
# + [markdown] papermill={} tags=[]
# ### Get commits from repository url
# + papermill={} tags=[]
df_commits = github.connect(GITHUB_TOKEN).repositories.get_commits(REPO_URL)
df_commits
# + [markdown] papermill={} tags=[]
# ## Output
# + [markdown] papermill={} tags=[]
# ### Get commits ranking by user
# + papermill={} tags=[]
def get_commits(df):
# Exclude Github commits
df = df[(df.COMMITTER_EMAIL.str[-10:] != "github.com")]
# Groupby and count
df = df.groupby(["AUTHOR_NAME"], as_index=False).agg({"ID": "count"})
# Cleaning
df = df.rename(columns={"ID": "NB_COMMITS"})
return df.sort_values(by="NB_COMMITS", ascending=False).reset_index(drop=True)
df = get_commits(df_commits)
df
# + [markdown] papermill={} tags=[]
# ### Plot a bar chart of commit activity
# + papermill={} tags=[]
def create_barchart(df, repository):
# Get repository
repository = repository.split("/")[-1]
# Sort df
df = df.sort_values(by="NB_COMMITS")
# Calc commits
commits = df.NB_COMMITS.sum()
# Create fig
fig = px.bar(df,
y="AUTHOR_NAME",
x="NB_COMMITS",
orientation='h',
title=f"Github - {repository} : Commits by user <br><span style='font-size: 13px;'>Total commits: {commits}</span>",
text="NB_COMMITS",
labels={"AUTHOR_NAME": "Author",
"NB_COMMITS": "Nb commits"}
)
fig.update_traces(marker_color='black')
fig.update_layout(
plot_bgcolor="#ffffff",
width=1200,
height=800,
font=dict(family="Arial", size=14, color="black"),
paper_bgcolor="white",
xaxis_title=None,
xaxis_showticklabels=False,
yaxis_title=None,
margin_pad=10,
)
fig.show()
return fig
fig = create_barchart(df, REPO_URL)
# + [markdown] papermill={} tags=[]
# ### Save and export html
# + papermill={} tags=[]
output_path = f"{REPO_URL.split('/')[-1]}_commits_ranking.html"
fig.write_html(output_path)
naas.asset.add(output_path, params={"inline": True})
| Github/Github_Get_commits_ranking_from_repository.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import division, print_function, absolute_import
import tensorflow as tf
# -
from src.data import data
from datetime import datetime as dt
import os
import numpy as np
from IPython import display
from PIL import Image
from glob import glob
def get_filenames_and_labels(data_dir):
filenames = []
labels = []
for d in os.listdir(data_dir):
category_filenames = glob(f'{data_dir}/{d}/*')
filenames.extend(category_filenames)
labels.extend([d for x in range(len(category_filenames))])
mapping = {l: idx for idx, l in enumerate(np.unique(labels))}
category = [mapping[l] for l in labels]
category = tf.keras.utils.to_categorical(category, len(np.unique(labels)))
return filenames, category
import random
def create_tf_dataset(filenames, labels, shuffle_buffer_size=10000, n_repeat_times=100, batch_size=128, shuffle=True, return_dataset=False):
def _parse_function(filename, label):
image_string = tf.read_file(filename)
image_decoded = tf.image.decode_jpeg(image_string, channels=3)
image = tf.cast(image_decoded, tf.float32)
# resized_images = tf.image.resize_images(images, (img_size, img_size))
resized_image = tf.image.resize_images(image, (img_size, img_size))
# resized_image = tf.reshape(
# resized_image,
# (img_size * img_size * 3, ))
return resized_image/255, label
len_filenames = len(filenames)
tuples = [(f, l) for f, l in zip(filenames, labels)]
random.seed(14)
random.shuffle(tuples)
filenames = [t[0] for t in tuples]
labels = np.array([t[1] for t in tuples])
# step 1
filenames = tf.constant(filenames)
labels = tf.constant(labels)
# step 2: create a dataset returning slices of `filenames`
dataset = tf.data.Dataset.from_tensor_slices((filenames, labels))
# step 3: parse every image in the dataset using `map`
dataset = dataset.map(_parse_function)
if shuffle:
dataset = dataset.shuffle(shuffle_buffer_size,
reshuffle_each_iteration=True)
dataset = dataset.repeat(n_repeat_times).batch(batch_size if shuffle else 1024)
if return_dataset:
return dataset
# step 4: create iterator and final input tensor
iterator = tf.compat.v1.data.make_one_shot_iterator(dataset)
images, labels = iterator.get_next()
return images, labels
# +
""" Convolutional Neural Network.
Build and train a convolutional neural network with TensorFlow.
This example is using the MNIST database of handwritten digits
(http://yann.lecun.com/exdb/mnist/)
Author: <NAME>
Project: https://github.com/aymericdamien/TensorFlow-Examples/
"""
# Training Parameters
num_epochs = 100
batch_size = 64
num_classes = 6
img_size = 128
channels = 3
learning_rate = 0.001
display_step = 50
num_input = (img_size * img_size * channels)
dropout = 0.5 # Dropout, probability to keep units
# tf Graph input
X = tf.placeholder(tf.float32, [None, img_size, img_size, channels], name='X')
Y = tf.placeholder(tf.float32, [None, num_classes], name='y')
keep_prob = tf.placeholder(tf.float32) # dropout (keep probability)
# +
# Create some wrappers for simplicity
def conv2d(x, W, b, strides=1):
# Conv2D wrapper, with bias and relu activation
x = tf.nn.conv2d(x, W, strides=[1, strides, strides, 1], padding='SAME')
x = tf.nn.bias_add(x, b)
return tf.nn.relu(x)
def maxpool2d(x, k=2):
# MaxPool2D wrapper
return tf.nn.max_pool(x, ksize=[1, k, k, 1], strides=[1, k, k, 1],
padding='SAME')
# Create model
def conv_net(x, weights, biases, dropout):
# x = tf.reshape(x, shape=[-1, 224, 224, 3])
# Convolution Layer
conv1 = conv2d(x, weights['block_1_wc1'], biases['block_1_bc1'])
conv1 = tf.compat.v1.layers.batch_normalization(conv1,
training=True if dropout != 1 else False)
conv1 = conv2d(conv1, weights['block_1_wc2'], biases['block_1_bc2'])
conv1 = tf.compat.v1.layers.batch_normalization(conv1,
training=True if dropout != 1 else False)
# Max Pooling (down-sampling)
conv1 = maxpool2d(conv1, k=2)
# Apply Dropout
conv1 = tf.nn.dropout(conv1, 1 if dropout == 1 else 0.75)
# Convolution Layer
conv2 = conv2d(conv1, weights['block_2_wc1'], biases['block_2_bc1'])
conv2 = tf.compat.v1.layers.batch_normalization(conv2,
training=True if dropout != 1 else False)
conv2 = conv2d(conv2, weights['block_2_wc2'], biases['block_2_bc2'])
conv2 = tf.compat.v1.layers.batch_normalization(conv2,
training=True if dropout != 1 else False)
# Max Pooling (down-sampling)
conv2 = maxpool2d(conv2, k=2)
# Apply Dropout
conv2 = tf.nn.dropout(conv2, 1 if dropout == 1 else 0.75)
# Fully connected layer
# Reshape conv2 output to fit fully connected layer input
flatten = tf.contrib.layers.flatten(conv2)
fc1 = tf.contrib.layers.fully_connected(flatten, 512, )
# fc1 = tf.reshape(conv2, [-1, weights['wd1'].get_shape().as_list()[0]])
# fc1 = tf.add(tf.matmul(fc1, weights['wd1']), biases['bd1'])
# fc1 = tf.nn.relu(fc1)
# Apply Dropout
fc1 = tf.nn.dropout(fc1, dropout)
# Output, class prediction
out = tf.add(tf.matmul(fc1, weights['out']), biases['out'])
return out
# Store layers weight & bias
weights = {
# 5x5 conv, 1 input, 32 outputs
'block_1_wc1': tf.Variable(tf.random_normal([3, 3, 3, 32])),
'block_1_wc2': tf.Variable(tf.random_normal([3, 3, 32, 32])),
'block_2_wc1': tf.Variable(tf.random_normal([3, 3, 32, 64])),
'block_2_wc2': tf.Variable(tf.random_normal([3, 3, 64, 64])),
# fully connected, 7*7*64 inputs, 1024 outputs
'wd1': tf.Variable(tf.random_normal([7*7*64, 512])),
# 1024 inputs, 6 outputs (class prediction)
'out': tf.Variable(tf.random_normal([512, num_classes]))
}
biases = {
'block_1_bc1': tf.Variable(tf.random_normal([32])),
'block_1_bc2': tf.Variable(tf.random_normal([32])),
'block_2_bc1': tf.Variable(tf.random_normal([64])),
'block_2_bc2': tf.Variable(tf.random_normal([64])),
'bd1': tf.Variable(tf.random_normal([512])),
'out': tf.Variable(tf.random_normal([num_classes]))
}
# -
# ### Data
# +
path = data.PATH()
dataset = 'vision_based_dataset'
base_dir = f'{path.PROCESSED_DATA_PATH}/{dataset}'
train_dir = f'{base_dir}/train/'
validation_dir = f'{base_dir}/validation/'
test_dir = f'{base_dir}/test/'
filenames_train, labels_train = get_filenames_and_labels(train_dir)
filenames_valid, labels_valid = get_filenames_and_labels(validation_dir)
train_images, train_labels = create_tf_dataset(filenames_train, labels_train, batch_size=batch_size)
validation_images, validation_labels = create_tf_dataset(filenames_valid, labels_valid, batch_size=batch_size, shuffle=False)
# -
# ### Model
num_steps = int(num_epochs * len(filenames_train) / batch_size) - 1
from src.models import cnn_model
from tensorflow.keras import initializers, regularizers
image_input = tf.keras.Input(shape=(img_size, img_size, 3), name='input_layer')
SEED = 14
# +
conv1 = tf.keras.layers.Conv2D(64, kernel_size=(5, 5), strides=(1, 1),
activation='relu',
kernel_initializer=initializers.glorot_uniform(seed=SEED),
bias_initializer=initializers.Constant(0.1))(image_input)
conv1 = tf.keras.layers.BatchNormalization()(conv1)
conv1 = tf.keras.layers.Conv2D(64, (5, 5), activation='relu')(conv1)
conv1 = tf.keras.layers.BatchNormalization()(conv1)
conv1 = tf.keras.layers.MaxPooling2D(pool_size=(2, 2), strides=(2, 2))(conv1)
conv1 = tf.keras.layers.Dropout(0.25)(conv1)
conv2 = tf.keras.layers.Conv2D(128, (5, 5), activation='relu')(conv1)
conv2 = tf.keras.layers.BatchNormalization()(conv2)
conv2 = tf.keras.layers.Conv2D(128, (5, 5), activation='relu')(conv2)
conv2 = tf.keras.layers.BatchNormalization()(conv2)
conv2 = tf.keras.layers.MaxPooling2D(pool_size=(2, 2))(conv2)
conv2 = tf.keras.layers.Dropout(0.25)(conv2)
conv_flat = tf.keras.layers.Flatten()(conv2)
fc1 = tf.keras.layers.Dense(512, kernel_regularizer=regularizers.l2(0.01),
activation='relu',)(conv_flat)
fc1 = tf.keras.layers.Dropout(0.5)(fc1)
output = tf.keras.layers.Dense(num_classes, activation='softmax')(fc1)
# -
model = tf.keras.Model(inputs=image_input, outputs=[output])
model.summary()
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['categorical_accuracy'])
# +
train_dataset = create_tf_dataset(filenames_train, labels_train, batch_size=batch_size, return_dataset=True)
val_dataset = create_tf_dataset(filenames_valid, labels_valid, batch_size=batch_size, shuffle=False, return_dataset=True)
# -
history = model.fit(train_dataset,
steps_per_epoch=1000, # let's take just a couple of steps
epochs=1)
# +
# Construct model
logits = conv_net(image_input, weights, biases, keep_prob)
prediction = tf.nn.softmax(logits)
# -
# +
# Define loss and optimizer
loss_op = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(
logits=logits, labels=Y))
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate)
train_op = optimizer.minimize(loss_op)
# Evaluate model
correct_pred = tf.equal(tf.argmax(prediction, 1), tf.argmax(Y, 1))
accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
# Initialize the variables (i.e. assign their default value)
init = tf.global_variables_initializer()
saver = tf.train.Saver(max_to_keep=4, keep_checkpoint_every_n_hours=2)
# Start training
with tf.Session() as sess:
# Run the initializer
sess.run(init)
start = dt.now()
for step in range(1, num_steps+1):
#batch_x, batch_y = mnist.train.next_batch(batch_size)
# batch_x, batch_y = iterator.get_next()
batch_x, batch_y = sess.run([train_images, train_labels])
batch_start = dt.now()
# Run optimization op (backprop)
sess.run(train_op, feed_dict={X: batch_x, Y: batch_y, keep_prob: 0.5})
if step % display_step == 0 or step == 1:
# Calculate batch loss and accuracy
loss, acc = sess.run([loss_op, accuracy],
feed_dict={
X: batch_x,
Y: batch_y,
keep_prob: 1.0}
)
validation_batch_x, validation_batch_y = sess.run(
[validation_images, validation_labels])
val_loss, val_acc = sess.run([loss_op, accuracy],
feed_dict={
X: validation_batch_x,
Y: validation_batch_y,
keep_prob: 1.0
})
end_display = (dt.now() - batch_start).total_seconds() / 60
print(f"Step {step}/{num_steps} ({step*batch_size} imgs) >> minibatch loss: {np.round(loss, 2)}; acc {np.round(acc, 2)}; val_loss: {np.round(val_loss, 2)}; val_acc {np.round(val_acc, 2)} (Total mins: {np.round((dt.now() - start).total_seconds() / 60, 2)})")
saved_path = saver.save(sess, './tf_api_model', global_step=step)
print("Optimization Finished!")
# -
# +
# Calculate accuracy for 256 MNIST test images
print("Testing Accuracy:", \
sess.run(accuracy, feed_dict={X: mnist.test.images[:256],
Y: mnist.test.labels[:256],
keep_prob: 1.0}))
# -
| notebooks/tfApiVersion_v2.0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: graphs
# language: python
# name: graphs
# ---
# ### Introduction
# The source graph for this notebook was prepared using the map taken from: https://github.com/pszufe/OpenStreetMapX.jl/blob/master/test/data/reno_east3.osm.
#
# In order to follow the notebook you need to make sure you have the `folium` package installed. You can add it to your Python environment e.g. using the following command `conda install -c conda-forge folium` (or similar, depending on the Python configuration you use).
## path to datasets
datadir='../Datasets/'
import folium as flm
import igraph as ig
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# In this notebook we want to analyze the real data where the graph is representing the road network of Reno, NV, USA.
#
# We are interested in finding which intersections can be expected to be most busy. In order to perform this analysys we assume that citizens want to travel between any two randomly picked intersections via a shortest path linking them.
#
# In the notebook, in order to highlight how sensitive the results of the analysis are to the level of detail reflected by the graph, we present three scenarios:
# * assuming that each edge in the road graph has the same length and travel time;
# * assigning real road length but assuming that the driving speed on each edge is the same;
# * assuming real road lengths and differentiating driving speeds between roads (e.g. you can drive faster using a highway than using a gravel road).
# Of course our analysis will still be lacking many real-life details that are potentially important in practice like. Here are some major things we ignore in the analysis:
# * non-uniform distribution of source and destination locations of travel;
# * number of lanes on each road;
# * relationship between traffic on a road and effective average driving speed;
# * road usage restrictions for certain classes of vehicles;
# * effect of street lights;
# * restrictions on turning on intersections.
#
# We left these details from the analysis to keep the example simple enough. However, we encourage readers to try to experiment and extend the presented model with some of these details to check how they would influence the results.
# ### Ingesting the data
# We first read in the source data. It is stored in two files:
# * `nodeloc.csv` that for each node of the graph (intersection) contains information on its geographic location;
# * `weights.csv` that for each edge of the graph (road) contains information on its length (weight), and speed a car can drive on a given road.
# +
## build undirected weighted graph
g_edges = pd.read_csv(datadir+'Reno/weights.csv')
nv = 1 + max(max(g_edges["from"]), max(g_edges["to"]))
g = ig.Graph(directed=True)
g.add_vertices(nv)
for i in range(len(g_edges)):
g.add_edge(g_edges["from"][i], g_edges["to"][i])
g.es['weight'] = g_edges['w']
g.es['speed'] = g_edges['speed']
# +
## read lat/lon position of nodes (intersections)
meta = pd.read_csv(datadir+'Reno/nodeloc.csv')
g.vs['longitude'] = list(meta['lon'])
g.vs['latitude'] = list(meta['lat'])
g.vs['layout'] = [(v['longitude'],v['latitude']) for v in g.vs]
g.vs['color'] = "black"
# -
# Check that the graph is connected:
g.is_connected()
# Check the number of nodes and edges:
print(g.vcount(),'nodes and',g.ecount(),'edges')
# Verify the degree distribution of nodes:
pd.Series(g.indegree()).value_counts(normalize=True, sort=False)
pd.Series(g.outdegree()).value_counts(normalize=True, sort=False)
# Note that interestingly nodes having in- and out- degree 1, 2, and 3 have similar frequency, and in- and out- degree equal to 4 is less frequent.
# Finally lest us visualize our graph.
# First we do it using standard iGraph plotting:
ly = ig.Layout(g.vs['layout'])
ly.mirror(1)
ig.plot(g, layout=ly, vertex_size=3, edge_arrow_size=0.01, edge_arrow_width=0.01, edge_curved=0)
# Let us also learn how we can nicely overlay a graph on top of a map using the `folium` package:
# +
MAP_BOUNDS = ((39.5001-0.001, -119.802-0.001), (39.5435+0.001, -119.7065+0.001))
m_plot = flm.Map()
for v in g.vs:
flm.Circle(
(v['latitude'], v['longitude']),
radius=1, weight=1,
color=v['color'], fill=True, fill_color=v['color']).add_to(m_plot)
for e in g.es:
v1 = g.vs[e.source]
v2 = g.vs[e.target]
flm.PolyLine(
[(v1['latitude'], v1['longitude']), (v2['latitude'], v2['longitude'])],
color="black", weight=1).add_to(m_plot)
flm.Rectangle(MAP_BOUNDS, color="blue",weight=4).add_to(m_plot)
m_plot.fit_bounds(MAP_BOUNDS)
m_plot
# -
# Observe that the plot produced by `folium` is interactive: you can zoom it and move it around.
#
# This plot confirms that ineed we have nodes correctly aligned with intersections on a map of Reno, NV, USA. For instance we see that there are no roads crossing the airport.
# Let us now show how to plot nodes of different in- and out- degrees using different colors:
# in-degree:
# yellow - 1
# blue - 2
# red - 3
# green - 4
ig.plot(g, layout=ly, vertex_color=list(np.array(['yellow', 'blue', 'red', 'green'])[np.array(g.indegree())-1]),
vertex_size=5, edge_arrow_size=0.01, edge_arrow_width=0.01, edge_curved=0)
# out-degree:
# yellow - 1
# blue - 2
# red - 3
# green - 4
ig.plot(g, layout=ly, vertex_color=list(np.array(['yellow', 'blue', 'red', 'green'])[np.array(g.outdegree())-1]),
vertex_size=5, edge_arrow_size=0.01, edge_arrow_width=0.01, edge_curved=0)
# On the plots we see that we have large differences in road lengths in our graph. Let us inverstigate it.
#
# Notice in particular that most nodes lying on the highway are of in- and out- degree 1. This is due to the fact that the highway has only few entry/exit points but its representation in OpenStreetMaps consists of many road segments.
plt.hist(g_edges["w"], 50);
# Indeed most of the roads are short, but some of them are very long.
# Similarly we see that there are different road classes in our graph: there are highways, but there are also many local roads.
pd.Series(g_edges["speed"]).value_counts(normalize=True)
# Indeed we see that the vast majority of the roads allow for the lowest speed.
# Let us check if roads allowing speed `120` coincide with the highways on the map. This is easy to do visually using the following code:
# +
MAP_BOUNDS = ((39.5001-0.001, -119.802-0.001), (39.5435+0.001, -119.7065+0.001))
m_plot = flm.Map()
for v in g.vs:
flm.Circle(
(v['latitude'], v['longitude']),
radius=1, weight=1,
color=v['color'], fill=True, fill_color=v['color']).add_to(m_plot)
for i in range(g.ecount()):
e = g.es[i]
v1 = g.vs[e.source]
v2 = g.vs[e.target]
if g.es["speed"][i] == 120:
w = 3
c = 'green'
else:
w = 1
c = 'black'
flm.PolyLine(
[(v1['latitude'], v1['longitude']), (v2['latitude'], v2['longitude'])],
color=c, weight=w).add_to(m_plot)
flm.Rectangle(MAP_BOUNDS, color="blue",weight=4).add_to(m_plot)
m_plot.fit_bounds(MAP_BOUNDS)
m_plot
# -
# Indeed we see that the thick green edges are covering a highway.
# Having checked our input data we may turn to the analysis trying to answer which intersections can be expected to be most busy on the map.
# ## Basic analysis - each edge has weight 1
#
# We use betweenness centrality to identify how busy a given intersection is, as it measures number of shortest paths in the graph that go through a given node.
#
# In the plots we distinguish 3 types of nodes with respect to their betweenness centrality:
# * the very heavy ones (big circle), 99th percentile
# * heavy ones (small circle), 90th percentile
# * others (very small circle)
## compute betweenness and plot distribution
bet = g.betweenness()
plt.hist(bet, 50);
# +
## size w.r.t. 3 types of nodes
very_heavy_usage = np.quantile(bet, 0.99)
heavy_usage = np.quantile(bet, 0.9)
g.vs['size'] = [1 if b < heavy_usage else 7 if b < very_heavy_usage else 14 for b in bet]
# -
## plot highlighting intersections with high betweenness
ly = ig.Layout(g.vs['layout'])
ly.mirror(1)
ig.plot(g, layout=ly, vertex_size=g.vs['size'], vertex_color=g.vs['color'], edge_arrow_size=0.01, edge_arrow_width=0.01, edge_curved=0)
# +
MAP_BOUNDS = ((39.5001-0.001, -119.802-0.001), (39.5435+0.001, -119.7065+0.001))
m_plot = flm.Map()
for v in g.vs:
flm.Circle(
(v['latitude'], v['longitude']),
radius=1, color=v['color'], weight= v['size'],
fill=True, fill_color=v['color']).add_to(m_plot)
for e in g.es:
v1 = g.vs[e.source]
v2 = g.vs[e.target]
flm.PolyLine(
[(v1['latitude'], v1['longitude']), (v2['latitude'], v2['longitude'])],
color="black", weight=1).add_to(m_plot)
flm.Rectangle(MAP_BOUNDS, color="blue",weight=4).add_to(m_plot)
m_plot.fit_bounds(MAP_BOUNDS)
m_plot
# -
# In this simple analysis the most busy nodes are lying around the center of the map. We also have a set of relatively busy intersections in the most dense regions of the map.
#
# However, it seems that this analysis is too simple. We are ignoring the fact how distant are nodes in the calculation of betweenness. Let us include the road lengths in our model.
# ## Using betweenness with road length
#
## compute betweenness and plot distribution
bet = g.betweenness(weights=g.es['weight'])
plt.hist(bet, 50);
# +
## size w.r.t. 3 types of nodes
very_heavy_usage = np.quantile(bet, 0.99)
heavy_usage = np.quantile(bet, 0.9)
g.vs['size'] = [1 if b < heavy_usage else 7 if b < very_heavy_usage else 14 for b in bet]
# -
## plot highlighting intersections with high betweenness
ly = ig.Layout(g.vs['layout'])
ly.mirror(1)
ig.plot(g, layout=ly, vertex_size=g.vs['size'], vertex_color=g.vs['color'], edge_arrow_size=0.01, edge_arrow_width=0.01, edge_curved=0)
# +
MAP_BOUNDS = ((39.5001-0.001, -119.802-0.001), (39.5435+0.001, -119.7065+0.001))
m_plot = flm.Map()
for v in g.vs:
flm.Circle(
(v['latitude'], v['longitude']),
radius=10, color=v['color'], weight= v['size'],
fill=True, fill_color=v['color']).add_to(m_plot)
for e in g.es:
v1 = g.vs[e.source]
v2 = g.vs[e.target]
flm.PolyLine(
[(v1['latitude'], v1['longitude']), (v2['latitude'], v2['longitude'])],
color="black", weight=1).add_to(m_plot)
flm.Rectangle(MAP_BOUNDS, color="blue",weight=4).add_to(m_plot)
m_plot.fit_bounds(MAP_BOUNDS)
m_plot
# -
# This time we see that the most busy intersections lie on the main roads. However, surprisinly, the highways seem not bo be used much. This clearly is related to the fact that we ignore the speed that cars can drive with on different roads. Let us then add this dimension to our analysis.
# ## Using betweenness with travel time
## compute betweenness and plot distribution
bet = g.betweenness(weights=g.es['weight'] / g_edges["speed"])
plt.hist(bet, 50);
# +
## size w.r.t. 3 types of nodes
very_heavy_usage = np.quantile(bet, 0.99)
heavy_usage = np.quantile(bet, 0.9)
g.vs['size'] = [1 if b < heavy_usage else 7 if b < very_heavy_usage else 14 for b in bet]
# -
## plot highlighting intersections with high betweenness
ly = ig.Layout(g.vs['layout'])
ly.mirror(1)
ig.plot(g, layout=ly, vertex_size=g.vs['size'], vertex_color=g.vs['color'], edge_arrow_size=0.01, edge_arrow_width=0.01, edge_curved=0)
# +
MAP_BOUNDS = ((39.5001-0.001, -119.802-0.001), (39.5435+0.001, -119.7065+0.001))
m_plot = flm.Map()
for v in g.vs:
flm.Circle(
(v['latitude'], v['longitude']),
radius=10, color=v['color'], weight= v['size'],
fill=True, fill_color=v['color']).add_to(m_plot)
for e in g.es:
v1 = g.vs[e.source]
v2 = g.vs[e.target]
flm.PolyLine(
[(v1['latitude'], v1['longitude']), (v2['latitude'], v2['longitude'])],
color="black", weight=1).add_to(m_plot)
flm.Rectangle(MAP_BOUNDS, color="blue",weight=4).add_to(m_plot)
m_plot.fit_bounds(MAP_BOUNDS)
m_plot
# -
# We finally get what we would expect in practice - the most busy intersections go along the highway as it is the fastest way to travel.
# In this experiment we could observe that relatively small changes to the setting of the problem might lead to significantly different conclusions. Fortunately, in this case, the most realistic assumptions lead to the most realistic outcome!
| Python_Notebooks/Complementary_D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import scipy.io
import os
import matplotlib.pyplot as plt
import numpy as np
dirname = os.getcwd()
print(dirname)
mat = scipy.io.loadmat('pdataBoxplotV2-M-A-0-1.mat')
#globals()
pdatamasslist = list([mat['pdatamass1'][0],mat['pdatamass2'][0],mat['pdatamass3'][0],mat['pdatamass4'][0],mat['pdatamass5'][0],mat['pdatamass6'][0]])
pdatavollist = list([mat['pdatavol1'][0],mat['pdatavol2'][0],mat['pdatavol3'][0],mat['pdatavol4'][0],mat['pdatavol5'][0],mat['pdatavol6'][0]])
fig, ax = plt.subplots(figsize=(8, 6), dpi=80, facecolor='w', edgecolor='k')
plt.rc('font', size=30)
plt.rc('lines',linewidth=2)
# build a violin plot
parts=ax.violinplot(pdatavollist, points=100,showmeans=False, showmedians=False, showextrema=False, widths=0.9)
#quantiles=[[0.25,0.75], [0.25,0.75], [0.25,0.75], [0.25,0.75], [0.25,0.75],[0.25,0.75]])
# add x-tick labels
xticklabels = ['0', '1', '5', '10', '25', '50']
ax.set_xticks([1,2,3,4,5,6])
ax.set_xticklabels(xticklabels)
ax.set_xlabel('[Arp2/3] in nM')
ax.set_ylabel('Domain volume ($\mu m^3$)')
#Change color
for pc in parts['bodies']:
pc.set_facecolor('#D43F3A')
pc.set_edgecolor('#000000')
pc.set_alpha(1)
pc.set_linewidth(2)
#Draw quartiles
bp=ax.boxplot(pdatavollist, notch=True, widths=0.1)
# changing color and linewidth of whiskers
for whisker in bp['whiskers']:
whisker.set(color ='yellow',
linewidth = 2,
linestyle ="-")
# changing color and linewidth of caps
for cap in bp['caps']:
cap.set(color ='yellow',
linewidth = 2)
# changing color and linewidth of medians
for median in bp['medians']:
median.set(color ='white',
linewidth = 8)
# changing style of fliers
for flier in bp['fliers']:
flier.set(marker ='o',
markerfacecolor ='yellow',
markersize=8,
alpha = 1)
# add x-tick labels
xticklabels = ['0', '1', '5', '10', '25', '50']
ax.set_xticks([1,2,3,4,5,6])
ax.set_xticklabels(xticklabels)
plt.savefig('Domainvolume.png', dpi=300)
| src/DomainVolume_violinplot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
import matplotlib.pyplot as plt
import numpy
import pandas as pd
file_name = './class activities/final project/books_small.json'
with open(file_name) as f:
for line in f:
print(reviewline)
break
#During this Step I import all the libararies I will be using to complete project
# +
# The Class listed below will give you the name of the book, the score on a 5 star system. Plue it will telll you if the review is
#Negative, Neutral or Positive.
# -
Class Review:
def __init__(self,text,score)
SELF.TEXT = TEXT
SELF.SCORE = SCORE
SELF.SENTIMENTS
Class review
def __init__(self, text, score)
self.text = text
self.score = score
def get_sentiment(self) = self.get_sentiment
if self.score <= 2:
return "Negative"
elif self.score == 3:
return "Neutral"
else: #Score of 4 or 5
return "Positive"
# #### Data Prep
# sklearn.feature_extraction.text import
#
from sklearn.feature_extraction.text import
vectorizer = CountVectorizer()
>>> X = vectorizer.fit_transform(corpus)
#This section will fit my model and transform it.
training, test = train_test_split(reviews, test_size=0.33, random_state=42)
#We were taugh to have a train environment of 20 %, but in this project,
#I chose to extend it to 33% due to the quantity of positive reviews.
# +
train_x = [x.text for x in training]
train_y = [x.sentiment for x in training]
test_x = [x.text for x in test]
test_y = [x.sentiment for x in test]
# +
# I tested it in two different models
# +
#Naive Bayes
from sklearn.naive_bayes import GaussianNB
clf_gnb = DecisionTreeClassifier()
clf_gnb.fit(train_x_vector, train_y)
clf_gnb.predict(test_x_vectors[0])
# +
#Logistic Regression
from sklearn.linear_model import LogisticRegression
clf_log = LogisticRegression
clf_log.fit(train_x_vector, train_y)
clf_log.predict(test_x_vector[0])
# +
#Bag of Words
from sklearn.feature_extraction.text import CountVectorizer, TfidVectorizer
vectorizer = CounterVectorizer()
train_x_vector = vectorizer.fit_transform(train_x)
test_x_vector = vectorizer.transform(test_x)
print(train_x[0])
print(train_x_vector[0].toarray())
#The models were scoring 80 % on predicting positve outputs.
#Only 1% on neutral and negative
# +
#Bar Graph
#This was setup show how many books would show up in the 1-5 star category
labels = [positive, neurtral, negative]
values = [1,2,3,4,5]
plt.bar(labels, value)
plt.figure(figsize=(6,4))
plt.show
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from mlrun import get_run_db, mlconf
# specify the DB path (use 'http://mlrun-api:8080' for api service)
mlconf.dbpath = mlconf.dbpath or 'http://mlrun-api:8080'
db = get_run_db().connect()
# list all runs
db.list_runs('download').show()
# list all artifact for version "latest"
db.list_artifacts('', tag='latest', project='iris').show()
# check different artifact versions
db.list_artifacts('ch', tag='*').show()
db.del_runs(state='completed')
db.del_artifacts(tag='*')
| examples/mlrun_db.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (py2)
# language: python
# name: py2
# ---
""" load them libs """
import matplotlib.pyplot as plt
import networkx as nx
import random
import math
import pandas as pd
import statsmodels.api as sm
import glob
import os
import numpy as np
from PIL import Image
from helpers import *
import pickle
import time
#random.seed(100)
#tic = time.time()
# +
""" get all files """
files = glob.glob('./matrix_csvs/*')
all_files = sorted([f for f in files if (f.split('/')[-1].split('_')[-1]=='influence.csv')])
print all_files
# +
""" get a dict going with files and data """
data = {}
years = range(2007,2018)
years_m = [2007,2008,2009,2010,2012,2013,2014,2015,2016,2017]
for y in years_m:
fname_all = [f for f in all_files if ((str(y)+'.75') in f)][0]
data[y]={'year':y,
'fname_all':fname_all,
'data_all':0.0}
#2011 notes
data[2011]={'year': 2011,
'fname_all': './matrix_csvs/2011.5_por_green_influence.csv',
'data_all': 0.0}
for y in years:
print data[y]
# -
for y in years:
print y
data[y]['data_all']=pd.read_csv(data[y]['fname_all'], encoding='utf-8')
print data[y]['data_all'].shape
print list(data[y]['data_all'])
# +
""" fill out extra columns """
all_indices = []
for y in years:
all_indices += list(data[y]['data_all'])
print set(all_indices)
for y in years:
new_cols = [v for v in set(all_indices) if v not in data[y]['data_all']]
for v in new_cols:
data[y]['data_all'][v]=0
for y in years:
print data[y]['data_all'].shape
print list(data[2017]['data_all'])
# -
""" Let's see what data we have """
data_files = sorted(glob.glob('data_pickles/*'))
print data_files
# +
""" load pickles into a list of dataframes """
df_list = []
for f in data_files[2:]:
df_list.append(pickle.load( open(f, 'rb')))
print len(df_list)
print (df_list[0])
all_indices = []
for l in df_list:
#print l.shape
all_indices = all_indices + list(l.index)
all_indices = list(set(all_indices))
# +
""" for each dataframe add missing columns """
for df in df_list:
country_cols = [h for h in list(df) if len(h)<3]
new_countries = [val for val in all_indices if val not in country_cols]
#print new_countries
for co in new_countries:
df[co]=0
for l in df_list:
print l.shape
# +
""" add INFLUENCE_t-1 """
for i in range(1,len(df_list)):
#print i
#print df_list[i]
df_list[i] = df_list[i].merge(df_list[i-1][['INFLUENCE']], left_index=True, right_index=True, how='left', suffixes=('','_t-1') )
#print df_list[i][['INFLUENCE','INFLUENCE_t-1']]
df_list[i]['INFLUENCE_t-1'] = df_list[i]['INFLUENCE_t-1'].fillna(0.0)
#print df_list[i][['INFLUENCE','INFLUENCE_t-1']]
for l in df_list:
print l.shape
# +
""" concat whole list """
df = pd.concat(df_list[1:], axis=0)
df.DATE = df.DATE-2000
print df.shape
# +
""" Train an OLS model """
X = df.drop(['POR_GREEN','INFLUENCE'],axis=1)
#print list(X)
#print X.DATE
X.DATE = (X.DATE - X.DATE.min())/(X.DATE.max()-X.DATE.min())
X['INFLUENCE_t-1'] = (X['INFLUENCE_t-1'] - X['INFLUENCE_t-1'].min())/(X['INFLUENCE_t-1'].max()-X['INFLUENCE_t-1'].min())
#print X['INFLUENCE_t-1']
#X.to_csv('data.csv')
#print X.isnull().any().any()
print X.DATE
print X['INFLUENCE_t-1']
print Y
Y = df.POR_GREEN
#print list(X)
#print list(Y)
#X = sm.add_constant(X)
est = sm.OLS(Y,X)
est = est.fit()
print est.summary()
#with pd.option_context('display.max_rows', None, 'display.max_columns', 3):
# print est.params
# -
| data_ols.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Função horária do MCU
# ***
# Num MCU a velocidade angular ($W$) e a escalar ($V$) são sempre as mesmas em qualquer intervalo de tempo.
#
# Dividindo todos os termos da equação horária do movimento uniforme ($S = S_0 + V.t$) pelo raio $R$ da trajetória:
#
# \begin{align*}
# \frac{S}{R} & = \frac{S_0}{R} + \frac{V.t}{R} \\
# φ & = \frac{S}{R} \\
# φ_0 & = \frac{S_0}{R} \\
# W & = \frac{V}{R} \\
# φ & = φ_0 + W.t
# \end{align*}
#
# 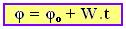
#
# A equação acima recebe o nome de equação (função) horária do MCU na forma angular, e aos termos $φ$ e $φ_0$ dá-se o nome de fases (ou ângulos) final e inicial, respectivamente.
# ***
# ### Movimento circular uniformemente variado (MCUV)
# ***
#
# Na figura abaixo o golfinho está efetuando movimento circular uniformemente retardado na subida e uniformemente acelerado na descida, ou seja, trata-se de um movimento circular uniformemente variado de equações: $S = S_0 + V_0.t + \frac{a.t^2}{2}$
#
# Dividindo por $R$:
#
# $$\frac{S}{R} = \frac{S_0}{R} + \frac{V_0.t}{R} + \frac{\frac{a}{R}.t^2}{2}$$
#
# 
#
# Para $V = V_0 + at$
#
# $$\frac{V}{R} = \frac{V_0}{R} + \frac{a}{R}.t$$
#
# 
#
# Para $V^2 = V_{0^2} + 2.a.ΔS$
#
# $$\left(\frac{V}{R}\right)^2 = \left(\frac{V_0}{R}\right)^2 + 2.\frac{a}{R}.Δφ$$
#
# 
#
# Logo:
#
# $$W = \frac{V}{R}$$
#
# $$λ = \frac{a}{R}$$
#
# $$Δφ = \frac{ΔS}{R}$$
#
# 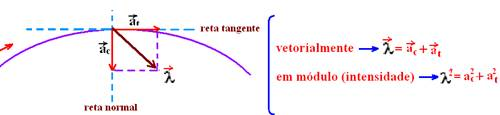
#
# $$a_t = a$$
#
# $$a_c = \frac{V^2}{R}$$
| Mecanica/cinematica/teoria/14_funcao_horaria_do_mcu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="4GKjLIFa8nDL"
# # BERT-based Recommendation System
# `Author: <NAME>`
#
# - 本notebook基于text_cnn baseline加以改进
# - 使用sentence-BERT预训练模型对非meta data做representation
# - 修改将电影类型用one-hot形式表示
# - 数据集使用[`MovieLens1M`](https://grouplens.org/datasets/movielens/)
# - 运行环境:google colab
#
# + [markdown] id="RkiSotsC8nDi"
# ## **0. 加载包**
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 10072, "status": "ok", "timestamp": 1605509693524, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="wC_uaNDAQkUw" outputId="4d1a4cfd-470c-470c-c333-5c208ba595ca"
# ! pip3 install sentence_transformers
# + executionInfo={"elapsed": 6567, "status": "ok", "timestamp": 1605509700908, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="u1MTfLvm8nEB"
import os
import pickle
import re
import time
import datetime
import random
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.decomposition import PCA
from collections import Counter
import tensorflow as tf
from tensorflow import keras
from tensorflow.python.ops import math_ops
from tensorflow.python.ops import summary_ops_v2
from sentence_transformers import SentenceTransformer, models
# + [markdown] id="mqSQJvW58nEo"
# ## **1. Data Preparing**
# + [markdown] id="n8sdu6nE8nEr"
# ### 1.1 查看数据
# 本项目使用的是MovieLens 1M 数据集,包含6000个用户在近4000部电影上的1亿条评论。
#
# 数据集分为三个文件:用户数据users.dat,电影数据movies.dat和评分数据ratings.dat。
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 16446, "status": "ok", "timestamp": 1605509721040, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="rphfmY4qPjgQ" outputId="c48d7e21-c58f-477d-83b8-6792f1cad662"
from google.colab import drive
drive.mount('/content/drive/')
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2630, "status": "ok", "timestamp": 1605509721041, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="Pge1Z62APkLX" outputId="c4c0425d-a2ed-4473-d11d-fd405fe18e43"
# %cd '/content/drive/My Drive/Movie_lens/'
# + [markdown] id="RhP5glF28nEt"
# #### 1.1.1 用户数据
# 分别有用户ID、性别、年龄、职业ID和邮编等字段。
#
# 数据中的格式:UserID::Gender::Age::Occupation::Zip-code
#
# - Gender is denoted by a "M" for male and "F" for female
# - Age is chosen from the following ranges:
#
# * 1: "Under 18"
# * 18: "18-24"
# * 25: "25-34"
# * 35: "35-44"
# * 45: "45-49"
# * 50: "50-55"
# * 56: "56+"
#
# - Occupation is chosen from the following choices:
#
# * 0: "other" or not specified
# * 1: "academic/educator"
# * 2: "artist"
# * 3: "clerical/admin"
# * 4: "college/grad student"
# * 5: "customer service"
# * 6: "doctor/health care"
# * 7: "executive/managerial"
# * 8: "farmer"
# * 9: "homemaker"
# * 10: "K-12 student"
# * 11: "lawyer"
# * 12: "programmer"
# * 13: "retired"
# * 14: "sales/marketing"
# * 15: "scientist"
# * 16: "self-employed"
# * 17: "technician/engineer"
# * 18: "tradesman/craftsman"
# * 19: "unemployed"
# * 20: "writer"
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 206} executionInfo={"elapsed": 969, "status": "ok", "timestamp": 1605509726989, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="qa2jymbs8nEt" outputId="ce74fca9-e44c-428a-c05c-0790d0a6d1e8"
users_title = ['UserID', 'Gender', 'Age', 'OccupationID', 'Zip-code']
users = pd.read_csv('./ml-1m/users.dat', sep='::', header=None, names=users_title, engine = 'python')
users.head()
# + [markdown] id="6fxSVkf48nE7"
# 可以看出UserID、Gender、Age和Occupation都是类别字段,其中邮编字段是我们不使用的。
# + [markdown] id="MGH3lKIi8nE7"
# #### 1.1.2 电影数据
# 分别有电影ID、电影名和电影风格等字段。
#
# 数据中的格式:MovieID::Title::Genres
#
# - Titles are identical to titles provided by the IMDB (including
# year of release)
# - Genres are pipe-separated and are selected from the following genres:
#
# * Action
# * Adventure
# * Animation
# * Children's
# * Comedy
# * Crime
# * Documentary
# * Drama
# * Fantasy
# * Film-Noir
# * Horror
# * Musical
# * Mystery
# * Romance
# * Sci-Fi
# * Thriller
# * War
# * Western
#
# + colab={"base_uri": "https://localhost:8080/", "height": 206} executionInfo={"elapsed": 972, "status": "ok", "timestamp": 1605509731083, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="CIkC6u2j8nE9" outputId="e4108b0c-8e72-4e22-ef24-cbe6d366401c"
movies_title = ['MovieID', 'Title', 'Genres']
movies = pd.read_csv('./ml-1m/movies.dat', sep='::', header=None, names=movies_title, engine = 'python')
movies.head()
# + [markdown] id="WA_PDU628nFI"
# MovieID是类别字段,Title是文本,Genres也是类别字段
# + [markdown] id="znWeBiEq8nFL"
# #### 1.1.3 评分数据
# 分别有用户ID、电影ID、评分和时间戳等字段。
#
# 数据中的格式:UserID::MovieID::Rating::Timestamp
#
# - UserIDs range between 1 and 6040
# - MovieIDs range between 1 and 3952
# - Ratings are made on a 5-star scale (whole-star ratings only)
# - Timestamp is represented in seconds since the epoch as returned by time(2)
# - Each user has at least 20 ratings
# + colab={"base_uri": "https://localhost:8080/", "height": 206} executionInfo={"elapsed": 6497, "status": "ok", "timestamp": 1605509739828, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="3KUyiMKA8nFL" outputId="2248226b-0d14-4de4-a76a-e0befb53e55c"
ratings_title = ['UserID','MovieID', 'Rating', 'timestamps']
ratings = pd.read_csv('./ml-1m/ratings.dat', sep='::', header=None, names=ratings_title, engine = 'python')
ratings.head()
# + [markdown] id="2GtONG2S8nFW"
# 评分字段Rating就是我们要学习的targets,时间戳字段我们不使用。
# + [markdown] id="4bWmdGHf8nFY"
# ### 1.2 数据预处理
# + [markdown] id="I8AgRCWN8nFb"
# - UserID、Occupation和MovieID不用变。
# - Gender字段:需要将‘F’和‘M’转换成0和1。
# - Age字段:要转成7个连续数字0~6。
# - Genres字段:是分类字段,使用One-hot embedding转成数字。
# - Title字段:使用Sentence BERT转成句向量。
# + [markdown] id="mP5tJTg7V-1m"
# #### 1.2.1 User dataset预处理
# + executionInfo={"elapsed": 1136, "status": "ok", "timestamp": 1605509743932, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="LSy4sBHqV92R"
#读取User数据
users_title = ['UserID', 'Gender', 'Age', 'JobID', 'Zip-code']
users = pd.read_csv('./ml-1m/users.dat', sep='::', header=None, names=users_title, engine = 'python')
users = users.filter(regex='UserID|Gender|Age|JobID')
users_orig = users.values
#改变User数据中性别和年龄
gender_map = {'F':0, 'M':1}
users['Gender'] = users['Gender'].map(gender_map)
age_map = {val:ii for ii,val in enumerate(set(users['Age']))}
users['Age'] = users['Age'].map(age_map)
# + [markdown] id="sGlHAspqW_FH"
# #### 1.2.2 Movie dataset预处理
# + executionInfo={"elapsed": 876, "status": "ok", "timestamp": 1605509747299, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="wg6Uuhu9XExJ"
#读取Movie数据
movies_title = ['MovieID', 'Title', 'Genres']
movies = pd.read_csv('./ml-1m/movies.dat', sep='::', header=None, names=movies_title, engine = 'python')
movies_orig = movies.values
# + [markdown] id="rThVfeq9Rjzd"
# ##### 1.2.2.1 Movie dataset 中Title字段处理
# + executionInfo={"elapsed": 935, "status": "ok", "timestamp": 1605509749434, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="5CAssjkObOam"
#将Title中的年份去掉
pattern = re.compile(r'^(.*)\((\d+)\)$')
title_map = {val:pattern.match(val).group(1) for ii,val in enumerate(set(movies['Title']))}
movies['Title'] = movies['Title'].map(title_map)
# + colab={"base_uri": "https://localhost:8080/", "height": 168, "referenced_widgets": ["80acd9370fbe4993b0f228c448904923", "0d0e5981669b413e80712704aac40df6", "070795ba80b34e2e8bfcc7cc84fdc6f1", "9f30ae648ad247ed89bf88c2fd5b1a1d", "67996e5c2f5e452882ba3b4f716b7ce0", "0fad55bfb5054db9beda3e9706740290", "75b37f74df754bbbad269dbe0e77dcc5", "408810d8df5b4c47aebf8bcd54a6a6e5", "8b81bd05377b423b92d210ee6026d215", "afe11a55aec842fa8691a8a13771a06f", "55f2949c3bbc45d380f96120fde9cb0c", "c340f86e52204715a4169a2a6c02c9a4", "b1d905aa87514c4195b4f55d5bd2e4d0", "<KEY>", "98ef0c080aad443ea696c4aea9ff02c6", "b323f97c4ae84d79804664d498bec7ff", "<KEY>", "f613a44183e14483b61dce20310541af", "<KEY>", "<KEY>", "4e0c8de536974c26afe729920729e108", "<KEY>", "316aa3b6500447a59d8e83449302db4e", "<KEY>"]} executionInfo={"elapsed": 159605, "status": "ok", "timestamp": 1605509910317, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="DOiDJHUjY7z-" outputId="549b3713-596f-4aa2-abe6-83e04ae2521b"
word_embedding_model = models.Transformer('bert-base-uncased', max_seq_length = 16) # 定义embedding模型
pooling_model = models.Pooling(word_embedding_model.get_word_embedding_dimension()) # 定义pooling层
model = SentenceTransformer(modules=[word_embedding_model, pooling_model]) # 总模型
title_encoded=model.encode(movies.Title[:]) # 对Title进行embedding
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 753, "status": "ok", "timestamp": 1605509915524, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="poQDSHdqSJaH" outputId="a0a2db99-c72e-4519-efb0-8fa7d1befa38"
"""
Sentence BERT模型的原理是:
将一个句子经过BERT网络,得到每个token的embedding(768维),对token embedding使用mean_over_time pooling得到整个句子的embedding(768维)
"""
title_encoded[0].shape
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 715, "status": "ok", "timestamp": 1605509917860, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="TQjlDQ2eWW14" outputId="6d4f31a7-e2c0-4877-c964-887e6f4bd02f"
#使用PCA将每个句子的embedding降到2维,可视化Sentence BERT embedding的效果
pca = PCA(n_components=2)
pca.fit(title_encoded)
print(pca.explained_variance_ratio_)
# + colab={"base_uri": "https://localhost:8080/", "height": 592} executionInfo={"elapsed": 1821, "status": "ok", "timestamp": 1605509921173, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="fam5iBueeKVZ" outputId="2bd68053-adb8-445e-9cd5-5ced1988fe9d"
#可视化50个Title embedding的效果
plot_only = 50
low_dim_embs = pca.fit_transform(title_encoded[:plot_only])
labels = [movies.Title[i] for i in range(plot_only)]
plt.figure(figsize=(10, 10))
for i, label in enumerate(labels):
x, y = low_dim_embs[i, :]
plt.scatter(x, y)
plt.annotate(label,xy=(x, y),xytext=(5, 2),textcoords='offset points',ha='right',va='bottom')
# + [markdown] id="hUTHS-XiWjgn"
# ##### 1.2.2.2 Movie dataset 中其他字段处理
# + id="bGV2Yyf4WmFq"
#电影类型one-hot编码
movies.Genres=movies.Genres.apply(lambda x: x.split('|'))
genres=[
'Action',
'Adventure',
'Animation',
"Children's",
'Comedy',
'Crime',
'Documentary',
'Drama',
'Fantasy',
'Film-Noir',
'Horror',
'Musical',
'Mystery',
'Romance',
'Sci-Fi',
'Thriller',
'War',
'Western'
]
for i in genres:
movies[i] = movies.Genres.apply(lambda x: i in x).astype(int)
classes = movies.iloc[:,3:].values
movies['Action']=movies['Action'].astype(object)
for i in range(len(movies)):
movies['Action'][i]=classes[i]
movies.drop(columns=['Genres','Adventure',
'Animation',
"Children's",
'Comedy',
'Crime',
'Documentary',
'Drama',
'Fantasy',
'Film-Noir',
'Horror',
'Musical',
'Mystery',
'Romance',
'Sci-Fi',
'Thriller',
'War',
'Western'],inplace=True)
movies = movies.rename(columns={'Action':'category'})
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1044, "status": "ok", "timestamp": 1605509943719, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="xTrHFyR1WzWr" outputId="20a97f8d-815c-4d91-f675-1b366ce8b972"
#把Title换成embedding
for i in range(len(movies)):
movies['Title'][i] = title_encoded[i]
# + [markdown] id="SQD77iLUVojm"
# #### 1.2.3 Dataset合并
# + executionInfo={"elapsed": 5785, "status": "ok", "timestamp": 1605510042864, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="O3SI8BXqd0z0"
#读取评分数据集
ratings_title = ['UserID','MovieID', 'ratings', 'timestamps']
ratings = pd.read_csv('./ml-1m/ratings.dat', sep='::', header=None, names=ratings_title, engine = 'python')
ratings = ratings.filter(regex='UserID|MovieID|ratings')
#合并三个表
data = pd.merge(pd.merge(ratings, users), movies)
#将数据分成feature dataframe和target dataframe
target_fields = ['ratings']
features_pd, targets_pd = data.drop(target_fields, axis=1), data[target_fields]
#转化成numpy数组
features = features_pd.values
targets_values = targets_pd.values
# + [markdown] id="w8ZCRBhq8nFx"
# #### 1.2.4 预处理后的数据一览
# + colab={"base_uri": "https://localhost:8080/", "height": 206} executionInfo={"elapsed": 657, "status": "ok", "timestamp": 1605510045753, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="VmcyIUWq8nF0" outputId="1387b2ea-0d7b-43dc-ddb6-5d239373754a"
users.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} executionInfo={"elapsed": 515, "status": "ok", "timestamp": 1605510046945, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="b-teBva59DXA" outputId="dec0a0b3-3a28-4055-a64f-1556396d0373"
movies.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} executionInfo={"elapsed": 681, "status": "ok", "timestamp": 1605510048649, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="Mlqge92y8nF5" outputId="b8b19692-1a1e-4d16-9ed9-a33482170965"
data.head()
# + [markdown] id="_zMAKaKA8nGU"
# ## **2. Model Design**
# + [markdown] id="egghvMDc8nGg"
# ### 2.1 预定义
# + [markdown] id="OTuBXqPxrE8U"
# #### 2.1.1 预定义变量
# + executionInfo={"elapsed": 975, "status": "ok", "timestamp": 1605510062788, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="_C0gSlO78nGg"
#嵌入矩阵的维度
embed_dim = 32
#用户ID个数
uid_max = max(features.take(0,1)) + 1 #共计6040
#性别个数
gender_max = max(features.take(2,1)) + 1 #共计2
#年龄类别个数
age_max = max(features.take(3,1)) + 1 #共计7
#职业个数
job_max = max(features.take(4,1)) + 1 #共计21
#电影ID个数
movie_id_max = max(features.take(1,1)) + 1 #共计3952
#电影Title embedding长度
sentences_size = title_count = 768
# + [markdown] id="UfA1ZLGm8nGm"
# #### 2.1.2 预定义超参
# + executionInfo={"elapsed": 660, "status": "ok", "timestamp": 1605510065214, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="JolsyUpT8nGn"
#Number of Epochs
num_epochs = 10
#Batch Size
batch_size = 256
#dropout
dropout_keep = 0.5
#Learning Rate
learning_rate = 0.0005
#Show stats for every n number of batches
show_every_n_batches = 20
#save path
save_dir = './save'
# + [markdown] id="QamhmNyc8nGr"
# #### 2.1.3 预定义占位符
# + executionInfo={"elapsed": 669, "status": "ok", "timestamp": 1605510067418, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="-g4kYV0A8nGu"
def get_inputs():
uid = tf.keras.layers.Input(shape=(1,), dtype='int32', name='uid')
user_gender = tf.keras.layers.Input(shape=(1,), dtype='int32', name='user_gender')
user_age = tf.keras.layers.Input(shape=(1,), dtype='int32', name='user_age')
user_job = tf.keras.layers.Input(shape=(1,), dtype='int32', name='user_job')
movie_id = tf.keras.layers.Input(shape=(1,), dtype='int32', name='movie_id')
movie_categories = tf.keras.layers.Input(shape=(18,), dtype='int32', name='movie_categories')
movie_titles = tf.keras.layers.Input(shape=(768,), dtype='int32', name='movie_titles')
return uid, user_gender, user_age, user_job, movie_id, movie_categories, movie_titles
# + [markdown] id="BZFUPF8x8nGz"
# ### 2.2 构建神经网络
# + [markdown] id="m7t8uLC98nG1"
# #### 2.2.1 定义User ID, age, job的嵌入矩阵
# + executionInfo={"elapsed": 700, "status": "ok", "timestamp": 1605510069472, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="1J1_0cXt8nG1"
def get_user_embedding(uid, user_gender, user_age, user_job):
uid_embed_layer = tf.keras.layers.Embedding(uid_max, embed_dim, input_length=1, name='uid_embed_layer')(uid)
gender_embed_layer = tf.keras.layers.Embedding(gender_max, embed_dim // 2, input_length=1, name='gender_embed_layer')(user_gender)
age_embed_layer = tf.keras.layers.Embedding(age_max, embed_dim // 2, input_length=1, name='age_embed_layer')(user_age)
job_embed_layer = tf.keras.layers.Embedding(job_max, embed_dim // 2, input_length=1, name='job_embed_layer')(user_job)
return uid_embed_layer, gender_embed_layer, age_embed_layer, job_embed_layer
# + [markdown] id="rUsn5hBB8nG7"
# #### 2.2.2将User的嵌入矩阵一起全连接生成User的特征
# + executionInfo={"elapsed": 672, "status": "ok", "timestamp": 1605510071570, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="GRChqrcm8nG8"
def get_user_feature_layer(uid_embed_layer, gender_embed_layer, age_embed_layer, job_embed_layer):
#第一层全连接
uid_fc_layer = tf.keras.layers.Dense(embed_dim, name="uid_fc_layer", activation='relu')(uid_embed_layer)
gender_fc_layer = tf.keras.layers.Dense(embed_dim, name="gender_fc_layer", activation='relu')(gender_embed_layer)
age_fc_layer = tf.keras.layers.Dense(embed_dim, name="age_fc_layer", activation='relu')(age_embed_layer)
job_fc_layer = tf.keras.layers.Dense(embed_dim, name="job_fc_layer", activation='relu')(job_embed_layer)
#第二层全连接
user_combine_layer = tf.keras.layers.concatenate([uid_fc_layer, gender_fc_layer, age_fc_layer, job_fc_layer], 2) #(?, 1, 128)
user_combine_layer = tf.keras.layers.Dense(200, activation='tanh')(user_combine_layer) #(?, 1, 200)
user_combine_layer_flat = tf.keras.layers.Reshape([200], name="user_combine_layer_flat")(user_combine_layer)
# dropout_layer = tf.keras.layers.Dropout(dropout_keep, name = "dropout_layer")(user_combine_layer_flat)
return user_combine_layer, user_combine_layer_flat
# + [markdown] id="2GSqTaU8tXh7"
# #### 2.2.3 定义Movie ID, genres, Title的嵌入矩阵
# + [markdown] id="V9WWJzG88nHF"
# ##### 2.2.3.1 定义Movie ID的嵌入矩阵
# + executionInfo={"elapsed": 673, "status": "ok", "timestamp": 1605510073493, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="4AUm__GV8nHF"
def get_movie_id_embed_layer(movie_id):
movie_id_embed_layer = tf.keras.layers.Embedding(movie_id_max, embed_dim, input_length=1, name='movie_id_embed_layer')(movie_id)
movie_id_embed_layer_flat = tf.keras.layers.Reshape([32], name="movie_id_embed_layer_flat")(movie_id_embed_layer)
return movie_id_embed_layer_flat
# + [markdown] id="0xj2Cr5B8nHK"
# ##### 2.2.3.2 定义Movie genres的嵌入矩阵
# + executionInfo={"elapsed": 702, "status": "ok", "timestamp": 1605510075412, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="Ge5Nx3NZHBE7"
def get_movie_categories_layers(movie_categories):
movie_categories_fc_layer = tf.keras.layers.Dense(embed_dim, name="movie_categories_fc_layer", activation='relu')(movie_categories)
return movie_categories_fc_layer
# + [markdown] id="EX-QjjY4tpFy"
# ##### 2.2.3.3 定义Movie Title的嵌入矩阵
# + executionInfo={"elapsed": 717, "status": "ok", "timestamp": 1605510077315, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="6FUeTyyk8nHL"
def get_movie_titles_layers(movie_titles):
movie_titles_fc_layer = tf.keras.layers.Dense(embed_dim, name="movie_titles_fc_layer", activation='relu')(movie_titles)
return movie_titles_fc_layer
# + [markdown] id="50i_PZhY8nHc"
# #### 2.2.4 将Movie的嵌入矩阵一起全连接生成movie的特征
# + executionInfo={"elapsed": 750, "status": "ok", "timestamp": 1605510084782, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="9djEwRQS8nHd"
def get_movie_feature_layer(movie_id_embed_layer_flat, movie_categories_fc_layer, movie_titles_fc_layer):
#第一层全连接
movie_id_fc_layer = tf.keras.layers.Dense(embed_dim, name="movie_id_fc_layer", activation='relu')(movie_id_embed_layer_flat)
#第二层全连接
movie_combine_layer = tf.keras.layers.concatenate([movie_id_fc_layer, movie_categories_fc_layer, movie_titles_fc_layer], 1)
movie_combine_layer = tf.keras.layers.Dense(200, activation='tanh')(movie_combine_layer)
movie_combine_layer_flat = tf.keras.layers.Reshape([200], name="movie_combine_layer_flat")(movie_combine_layer)
# dropout_layer1 = tf.keras.layers.Dropout(dropout_keep, name = "dropout_layer1")(movie_combine_layer_flat)
return movie_combine_layer, movie_combine_layer_flat
# + [markdown] id="OczD316N8nHt"
# #### 2.2.5 构建Tensorflow计算图
# + executionInfo={"elapsed": 1074, "status": "ok", "timestamp": 1605510086960, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="ulvsrkcO8nHu"
MODEL_DIR = "./models"
class mv_network(object):
def __init__(self, batch_size=256):
self.batch_size = batch_size
self.best_loss = 9999
self.losses = {'train': [], 'test': []}
# 获取输入占位符
uid, user_gender, user_age, user_job, movie_id, movie_categories, movie_titles = get_inputs()
# 获取User的4个嵌入向量
uid_embed_layer, gender_embed_layer, age_embed_layer, job_embed_layer = get_user_embedding(uid, user_gender, user_age, user_job)
# 得到用户特征
user_combine_layer, user_combine_layer_flat = get_user_feature_layer(uid_embed_layer, gender_embed_layer, age_embed_layer, job_embed_layer)
# 获取电影ID的嵌入向量
movie_id_embed_layer_flat = get_movie_id_embed_layer(movie_id)
# 获取电影种类的特征向量
movie_categories_fc_layer = get_movie_categories_layers(movie_categories)
# 获取电影名的特征向量
movie_titles_fc_layer = get_movie_titles_layers(movie_titles)
# 得到电影特征
movie_combine_layer, movie_combine_layer_flat = get_movie_feature_layer(movie_id_embed_layer_flat,movie_titles_fc_layer, movie_titles_fc_layer)
######## Plan 1 #########
# 将用户特征和电影特征做矩阵乘法得到一个预测评分的方案
inference = tf.keras.layers.Lambda(lambda layer:
tf.reduce_sum(layer[0] * layer[1], axis=1), name="inference")((user_combine_layer_flat, movie_combine_layer_flat))
inference = tf.keras.layers.Lambda(lambda layer: tf.expand_dims(layer, axis=1))(inference)
######## Plan 2 #########
# # 将用户特征和电影特征作为输入,经过全连接,输出一个值的方案
# inference_layer = tf.keras.layers.concatenate([user_combine_layer_flat, movie_combine_layer_flat],1) # (?, 400)
# # 你可以使用下面这个全连接层,试试效果
# inference_dense = tf.keras.layers.Dense(64, kernel_regularizer=tf.nn.l2_loss, activation='relu')(inference_layer)
# inference = tf.keras.layers.Dense(1, name="inference")(inference_layer) # inference_dense
self.model = tf.keras.Model(
inputs=[uid, user_gender, user_age, user_job, movie_id, movie_categories, movie_titles],
outputs=[inference])
self.model.summary()
self.optimizer = tf.keras.optimizers.Adam(learning_rate)
# MSE损失,将计算值回归到评分
self.ComputeLoss = tf.keras.losses.MeanSquaredError()
self.ComputeMetrics = tf.keras.metrics.MeanSquaredError()
if tf.io.gfile.exists(MODEL_DIR):
print('Removing existing model dir: {}'.format(MODEL_DIR))
tf.io.gfile.rmtree(MODEL_DIR)
pass
else:
tf.io.gfile.makedirs(MODEL_DIR)
train_dir = os.path.join(MODEL_DIR, 'summaries', 'train')
test_dir = os.path.join(MODEL_DIR, 'summaries', 'eval')
# self.train_summary_writer = summary_ops_v2.create_file_writer(train_dir, flush_millis=10000)
# self.test_summary_writer = summary_ops_v2.create_file_writer(test_dir, flush_millis=10000, name='test')
checkpoint_dir = os.path.join(MODEL_DIR, 'checkpoints')
self.checkpoint_prefix = os.path.join(checkpoint_dir, 'ckpt')
self.checkpoint = tf.train.Checkpoint(model=self.model, optimizer=self.optimizer)
# Restore variables on creation if a checkpoint exists.
self.checkpoint.restore(tf.train.latest_checkpoint(checkpoint_dir))
def compute_loss(self, labels, logits):
return tf.reduce_mean(tf.keras.losses.mse(labels, logits))
def compute_metrics(self, labels, logits):
return tf.keras.metrics.mse(labels, logits) #
@tf.function
def train_step(self, x, y):
# Record the operations used to compute the loss, so that the gradient
# of the loss with respect to the variables can be computed.
# metrics = 0
with tf.GradientTape() as tape:
logits = self.model([x[0],
x[1],
x[2],
x[3],
x[4],
x[5],
x[6]], training=True)
loss = self.ComputeLoss(y, logits)
# loss = self.compute_loss(labels, logits)
self.ComputeMetrics(y, logits)
# metrics = self.compute_metrics(labels, logits)
grads = tape.gradient(loss, self.model.trainable_variables)
self.optimizer.apply_gradients(zip(grads, self.model.trainable_variables))
return loss, logits
def training(self, features, targets_values, epochs=7, log_freq=50):
train_X, test_X, train_y, test_y = train_test_split(features, targets_values, test_size=0.2, random_state=0)
for epoch_i in range(epochs):
# 将训练集shuffle
state = np.random.get_state()
np.random.shuffle(train_X)
np.random.set_state(state)
np.random.shuffle(train_y)
train_batches = get_batches(train_X, train_y, self.batch_size)
batch_num = (len(train_X) // self.batch_size)
train_start = time.time()
# with self.train_summary_writer.as_default():
if True:
start = time.time()
# Metrics are stateful. They accumulate values and return a cumulative
# result when you call .result(). Clear accumulated values with .reset_states()
avg_loss = tf.keras.metrics.Mean('loss', dtype=tf.float32)
# avg_mae = tf.keras.metrics.Mean('mae', dtype=tf.float32)
# Datasets can be iterated over like any other Python iterable.
for batch_i in range(batch_num):
x, y = next(train_batches)
categories = np.zeros([self.batch_size, 18])
for i in range(self.batch_size):
categories[i] = x.take(6, 1)[i]
titles = np.zeros([self.batch_size, sentences_size])
for i in range(self.batch_size):
titles[i] = x.take(5, 1)[i]
loss, logits = self.train_step([np.reshape(x.take(0, 1), [self.batch_size, 1]).astype(np.float32),
np.reshape(x.take(2, 1), [self.batch_size, 1]).astype(np.float32),
np.reshape(x.take(3, 1), [self.batch_size, 1]).astype(np.float32),
np.reshape(x.take(4, 1), [self.batch_size, 1]).astype(np.float32),
np.reshape(x.take(1, 1), [self.batch_size, 1]).astype(np.float32),
categories.astype(np.float32),
titles.astype(np.float32)],
np.reshape(y, [self.batch_size, 1]).astype(np.float32))
avg_loss(loss)
self.losses['train'].append(loss)
if tf.equal(self.optimizer.iterations % log_freq, 0):
# summary_ops_v2.scalar('loss', avg_loss.result(), step=self.optimizer.iterations)
# summary_ops_v2.scalar('mae', self.ComputeMetrics.result(), step=self.optimizer.iterations)
# summary_ops_v2.scalar('mae', avg_mae.result(), step=self.optimizer.iterations)
rate = log_freq / (time.time() - start)
print('Step #{}\tEpoch {:>3} Batch {:>4}/{} Loss: {:0.6f} mse: {:0.6f} ({} steps/sec)'.format(
self.optimizer.iterations.numpy(),
epoch_i,
batch_i,
batch_num,
loss, (self.ComputeMetrics.result()), rate))
# print('Step #{}\tLoss: {:0.6f} mae: {:0.6f} ({} steps/sec)'.format(
# self.optimizer.iterations.numpy(), loss, (avg_mae.result()), rate))
avg_loss.reset_states()
self.ComputeMetrics.reset_states()
# avg_mae.reset_states()
start = time.time()
train_end = time.time()
print(
'\nTrain time for epoch #{} ({} total steps): {}'.format(epoch_i + 1, self.optimizer.iterations.numpy(),
train_end - train_start))
# with self.test_summary_writer.as_default():
self.testing((test_X, test_y), self.optimizer.iterations)
# self.checkpoint.save(self.checkpoint_prefix)
self.export_path = os.path.join(MODEL_DIR, 'export')
tf.saved_model.save(self.model, self.export_path)
def testing(self, test_dataset, step_num):
test_X, test_y = test_dataset
test_batches = get_batches(test_X, test_y, self.batch_size)
"""Perform an evaluation of `model` on the examples from `dataset`."""
avg_loss = tf.keras.metrics.Mean('loss', dtype=tf.float32)
# avg_mae = tf.keras.metrics.Mean('mae', dtype=tf.float32)
batch_num = (len(test_X) // self.batch_size)
for batch_i in range(batch_num):
x, y = next(test_batches)
categories = np.zeros([self.batch_size, 18])
for i in range(self.batch_size):
categories[i] = x.take(6, 1)[i]
titles = np.zeros([self.batch_size, sentences_size])
for i in range(self.batch_size):
titles[i] = x.take(5, 1)[i]
logits = self.model([np.reshape(x.take(0, 1), [self.batch_size, 1]).astype(np.float32),
np.reshape(x.take(2, 1), [self.batch_size, 1]).astype(np.float32),
np.reshape(x.take(3, 1), [self.batch_size, 1]).astype(np.float32),
np.reshape(x.take(4, 1), [self.batch_size, 1]).astype(np.float32),
np.reshape(x.take(1, 1), [self.batch_size, 1]).astype(np.float32),
categories.astype(np.float32),
titles.astype(np.float32)], training=False)
test_loss = self.ComputeLoss(np.reshape(y, [self.batch_size, 1]).astype(np.float32), logits)
avg_loss(test_loss)
# 保存测试损失
self.losses['test'].append(test_loss)
self.ComputeMetrics(np.reshape(y, [self.batch_size, 1]).astype(np.float32), logits)
# avg_loss(self.compute_loss(labels, logits))
# avg_mae(self.compute_metrics(labels, logits))
print('Model test set loss: {:0.6f} mse: {:0.6f}'.format(avg_loss.result(), self.ComputeMetrics.result()))
# print('Model test set loss: {:0.6f} mae: {:0.6f}'.format(avg_loss.result(), avg_mae.result()))
# summary_ops_v2.scalar('loss', avg_loss.result(), step=step_num)
# summary_ops_v2.scalar('mae', self.ComputeMetrics.result(), step=step_num)
# summary_ops_v2.scalar('mae', avg_mae.result(), step=step_num)
if avg_loss.result() < self.best_loss:
self.best_loss = avg_loss.result()
print("best loss = {}".format(self.best_loss))
self.checkpoint.save(self.checkpoint_prefix)
def forward(self, xs):
predictions = self.model(xs)
# logits = tf.nn.softmax(predictions)
return predictions
# + executionInfo={"elapsed": 743, "status": "ok", "timestamp": 1605510089430, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="Wa3Gui4I8nHz"
def get_batches(Xs, ys, batch_size):
for start in range(0, len(Xs), batch_size):
end = min(start + batch_size, len(Xs))
yield Xs[start:end], ys[start:end]
# + [markdown] id="LOCTMXlD8nH4"
# ### 2.3 训练神经网络
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 704441, "status": "ok", "timestamp": 1605510797037, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="aRESiEPr8nH6" outputId="0418c31f-84be-4077-e369-5361595c9375"
mv_net=mv_network()
mv_net.training(features, targets_values, epochs=7)
# + [markdown] id="_u8qVrowy8Zw"
# ### 2.4 可视化模型的训练结果
# + [markdown] id="Ih2l07ry8nII"
# #### 2.4.1 显示训练Loss
# + colab={"base_uri": "https://localhost:8080/", "height": 496} executionInfo={"elapsed": 1678, "status": "ok", "timestamp": 1605510813580, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="pmGw3DPl8nII" outputId="a171f91b-0d81-4b41-e457-77f35c892ddf"
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
plt.figure(figsize=(12,8))
plt.plot(mv_net.losses['train'], label='Training loss')
plt.xlabel('Step')
plt.ylabel('Training loss')
plt.legend()
_ = plt.ylim()
# + [markdown] id="2ng0hf308nIN"
# #### 2.4.2 显示测试Loss
# + colab={"base_uri": "https://localhost:8080/", "height": 496} executionInfo={"elapsed": 1909, "status": "ok", "timestamp": 1605510818152, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="-VmzxdBC8nIN" outputId="862719c9-0f72-44df-eed0-75685442d7be"
plt.figure(figsize=(12,8))
plt.plot(mv_net.losses['test'], label='Test loss')
plt.xlabel('Batch')
plt.ylabel('Test loss pre batch')
plt.legend()
_ = plt.ylim()
# + [markdown] id="cxVvFPEWzpX-"
# ## **3. 利用模型进行电影推荐**
# + [markdown] id="ezM8_KIc8nIZ"
# ### 3.1 生成Movie特征矩阵
# 将训练好的电影特征组合成电影特征矩阵并保存到本地
# + executionInfo={"elapsed": 63607, "status": "ok", "timestamp": 1605510909030, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="cFPEmeg58nIZ"
movie_layer_model = keras.models.Model(inputs=[mv_net.model.input[4], mv_net.model.input[5], mv_net.model.input[6]],
outputs=mv_net.model.get_layer("movie_combine_layer_flat").output)
movie_matrics = []
for item in movies.values:
categories = np.zeros([1, 18])
categories[0] = item.take(2)
titles = np.zeros([1, sentences_size])
titles[0] = item.take(1)
movie_combine_layer_flat_val = movie_layer_model([np.reshape(item.take(0), [1, 1]), categories, titles])
movie_matrics.append(movie_combine_layer_flat_val)
pickle.dump((np.array(movie_matrics).reshape(-1, 200)), open('movie_matrics.p', 'wb'))
movie_matrics = pickle.load(open('movie_matrics.p', mode='rb'))
# + [markdown] id="P65VdhuO8nIg"
# ### 3.2 生成User特征矩阵
# 将训练好的用户特征组合成用户特征矩阵并保存到本地
# + executionInfo={"elapsed": 120497, "status": "ok", "timestamp": 1605511032187, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="oD_HWGjW8nIh"
user_layer_model = keras.models.Model(inputs=[mv_net.model.input[0], mv_net.model.input[1], mv_net.model.input[2], mv_net.model.input[3]],
outputs=mv_net.model.get_layer("user_combine_layer_flat").output)
users_matrics = []
for item in users.values:
user_combine_layer_flat_val = user_layer_model([np.reshape(item.take(0), [1, 1]),
np.reshape(item.take(1), [1, 1]),
np.reshape(item.take(2), [1, 1]),
np.reshape(item.take(3), [1, 1])])
users_matrics.append(user_combine_layer_flat_val)
pickle.dump((np.array(users_matrics).reshape(-1, 200)), open('users_matrics.p', 'wb'))
users_matrics = pickle.load(open('users_matrics.p', mode='rb'))
# + [markdown] id="OGaswSaA8nIp"
# ### 3.3 开始推荐电影
# 使用生产的用户特征矩阵和电影特征矩阵做电影推荐
# + [markdown] id="mgkFlqfU8nIq"
# #### 3.3.1 推荐同类型的电影
# 思路是计算当前看的电影特征向量与整个电影特征矩阵的余弦相似度,取相似度最大的top_k个,这里加了些随机选择在里面,保证每次的推荐稍稍有些不同。
# + executionInfo={"elapsed": 741, "status": "ok", "timestamp": 1605511189417, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="32Sh_cgp7IVi"
#电影ID转下标的字典,数据集中电影ID跟下标不一致,比如第5行的数据电影ID不一定是5
movieid2idx = {val[0]:i for i, val in enumerate(movies.values)}
# + executionInfo={"elapsed": 886, "status": "ok", "timestamp": 1605511191418, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="EdHlwyDj8nIr"
def recommend_same_type_movie(movie_id_val, top_k = 20):
norm_movie_matrics = tf.sqrt(tf.reduce_sum(tf.square(movie_matrics), 1, keepdims=True))
normalized_movie_matrics = movie_matrics / norm_movie_matrics
#推荐同类型的电影
probs_embeddings = (movie_matrics[movieid2idx[movie_id_val]]).reshape([1, 200])
probs_similarity = tf.matmul(probs_embeddings, tf.transpose(normalized_movie_matrics))
sim = (probs_similarity.numpy())
# results = (-sim[0]).argsort()[0:top_k]
# print(results)
print("您看的电影是:{}".format(movies_orig[movieid2idx[movie_id_val]]))
print("以下是给您的推荐:")
p = np.squeeze(sim)
p[np.argsort(p)[:-top_k]] = 0
p = p / np.sum(p)
results = set()
while len(results) != 5:
c = np.random.choice(3883, 1, p=p)[0]
results.add(c)
for val in (results):
print(val)
print(movies_orig[val])
return results
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 699, "status": "ok", "timestamp": 1605511192720, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="xxElXSEY8nIw" outputId="e5207146-76ff-4cac-8876-cea58e2ea84e"
recommend_same_type_movie(1401, 20)
# + [markdown] id="15EJvv8r8nI0"
# #### 3.3.2 推荐用户喜欢的电影
# 思路是使用用户特征向量与电影特征矩阵计算所有电影的评分,取评分最高的top_k个,同样加了些随机选择部分。
# + executionInfo={"elapsed": 717, "status": "ok", "timestamp": 1605511196994, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="enDZgX2z8nI0"
def recommend_your_favorite_movie(user_id_val, top_k = 10):
#推荐您喜欢的电影
probs_embeddings = (users_matrics[user_id_val-1]).reshape([1, 200])
probs_similarity = tf.matmul(probs_embeddings, tf.transpose(movie_matrics))
sim = (probs_similarity.numpy())
# print(sim.shape)
# results = (-sim[0]).argsort()[0:top_k]
# print(results)
# sim_norm = probs_norm_similarity.eval()
# print((-sim_norm[0]).argsort()[0:top_k])
print("以下是给您的推荐:")
p = np.squeeze(sim)
p[np.argsort(p)[:-top_k]] = 0
p = p / np.sum(p)
results = set()
while len(results) != 5:
c = np.random.choice(3883, 1, p=p)[0]
results.add(c)
for val in (results):
print(val)
print(movies_orig[val])
return results
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 484, "status": "ok", "timestamp": 1605511197937, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="xrb4MsvZ8nI6" outputId="1c757669-0d62-4fc4-9a1a-dcd08a17c78f"
recommend_your_favorite_movie(234, 10)
# + [markdown] id="XNzraC8O8nI-"
# #### 3.3.3 看过这个电影的人还看了(喜欢)哪些电影
# - 首先选出喜欢某个电影的top_k个人,得到这几个人的用户特征向量。
# - 然后计算这几个人对所有电影的评分
# - 选择每个人评分最高的电影作为推荐
# - 同样加入了随机选择
# + executionInfo={"elapsed": 979, "status": "ok", "timestamp": 1605511201178, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="i7gr9us88nI_"
def recommend_other_favorite_movie(movie_id_val, top_k = 20):
probs_movie_embeddings = (movie_matrics[movieid2idx[movie_id_val]]).reshape([1, 200])
probs_user_favorite_similarity = tf.matmul(probs_movie_embeddings, tf.transpose(users_matrics))
favorite_user_id = np.argsort(probs_user_favorite_similarity.numpy())[0][-top_k:]
# print(normalized_users_matrics.numpy().shape)
# print(probs_user_favorite_similarity.numpy()[0][favorite_user_id])
# print(favorite_user_id.shape)
print("您看的电影是:{}".format(movies_orig[movieid2idx[movie_id_val]]))
print("喜欢看这个电影的人是:{}".format(users_orig[favorite_user_id-1]))
probs_users_embeddings = (users_matrics[favorite_user_id-1]).reshape([-1, 200])
probs_similarity = tf.matmul(probs_users_embeddings, tf.transpose(movie_matrics))
sim = (probs_similarity.numpy())
# results = (-sim[0]).argsort()[0:top_k]
# print(results)
# print(sim.shape)
# print(np.argmax(sim, 1))
p = np.argmax(sim, 1)
print("喜欢看这个电影的人还喜欢看:")
if len(set(p)) < 5:
results = set(p)
else:
results = set()
while len(results) != 5:
c = p[random.randrange(top_k)]
results.add(c)
for val in (results):
print(val)
print(movies_orig[val])
return results
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 557, "status": "ok", "timestamp": 1605511201179, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08237551511886881536"}, "user_tz": -480} id="BVb-X2uj8nJC" outputId="d7ffbf87-85ea-48cd-a4bd-3fd17918b1b6"
recommend_other_favorite_movie(1401, 20)
| bert_based.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Constructing an IMNN with on-the-fly additive noise
#
# The IMNN is fairly flexible and extendable. As an example, lets imagine that we have some signal that we want to do inference on, but there is an expensive additive noise model which is can be simulated, but simulations on the fly would be extremely expensive. In this case we could make a set of different noises and save them to file (or in memory, etc.) and add them randomly to the signal. If the signal model is not differentiable then we would also have to use numerical derivatives to obtain the derivative of the network outputs with respect to the model parameters. This requires specific ordering of which noises are grabbed for a single iteration.
#
# This example will show how we can add a limited set of Gaussian noise on the fly to sloped lines, i.e.
#
# $$y=mx+\epsilon$$
#
# where our parameter of interest is a noisy slope, $m = m' + \zeta$ where $\zeta\leftarrow N(0, 0.1)$.
from imnn import AggregatedNumericalGradientIMNN
from imnn.lfi import ApproximateBayesianComputation
import jax
import jax.numpy as np
import matplotlib.pyplot as plt
import tensorflow_probability
tfp = tensorflow_probability.substrates.jax
from functools import partial
from jax.experimental import stax
from jax.experimental import optimizers
rng = jax.random.PRNGKey(0)
# We will summarise the slope using `n_summaries=1` summaries and generate 2000 simulations and an extra 1000 simulations above and below the fiducial slope value to approximate the derivatives. For the fiducial value we'll choose $m'=1$ and $\delta m'=0.1$. We'll make $x$ a 100 length vector from 0 to 10. We'll make 500 noise realisations, $\epsilon$, drawn from a Gaussian with mean zero and variance 1.
# +
n_params = 1
n_summaries = 1
input_shape = (100,)
n_s = 2000
n_d = 1000
n_noise = 500
m_fid = np.array([1.])
δm = np.array([0.1])
x = np.linspace(0, 10, input_shape[0])
# -
# The noises (which are not expensive in this case, but could be in general) are going to be drawn from a Gaussian, $ϵ\leftarrow N(0, 1)$
rng, key = jax.random.split(rng)
ϵ = jax.random.normal(key, shape=(n_noise,) + input_shape)
# We'll also make the variable element of the simulation, $\zeta$ for training and validation.
rng, key, validation_key = jax.random.split(rng, num=3)
ζ = np.sqrt(0.1) * jax.random.normal(key, shape=(n_s,))
validation_ζ = np.sqrt(0.1) * jax.random.normal(validation_key, shape=(n_s,))
# We can now make the datasets for the signal, i.e. $mx = (m^\textrm{fid} + \zeta)x$ and its derivatives $mx^{\pm}=(m^\textrm{fid}\pm\delta m/2 + \zeta)x$ for both the training and the validation
# +
mx = np.expand_dims(m_fid[0] + ζ, 1) * np.expand_dims(x, 0)
mx_mp = np.expand_dims(
np.stack(
[(np.expand_dims((m_fid[0] - δm[0] / 2) + ζ[:n_d], 1)
* np.expand_dims(x, 0)),
(np.expand_dims((m_fid[0] + δm[0] / 2) + ζ[:n_d], 1)
* np.expand_dims(x, 0))],
1),
2)
validation_mx = (np.expand_dims(m_fid[0] + validation_ζ, 1)
* np.expand_dims(x, 0))
validation_mx_mp = np.expand_dims(
np.stack(
[(np.expand_dims((m_fid[0] - δm[0] / 2) + validation_ζ[:n_d], 1)
* np.expand_dims(x, 0)),
(np.expand_dims((m_fid[0] + δm[0] / 2) + validation_ζ[:n_d], 1)
* np.expand_dims(x, 0))],
1),
2)
# -
# ### Constructing a new IMNN
#
# Now we can construct the new IMNN. For our example we'll imagine that we want to use the AggregatedNumericalGradientIMNN (normally used for large datasets where the derivative of the simulations with respect to the model parameters can not easily be calculated). In this case we can use AggregatedNumericalGradientIMNN as the parent class, and initialise all attributes and add just the noise as an input (from which we will get the number of noises available from its shape). Now we just need to edit two of the class functions to add random noises on the fly. First the generators provided in `_collect_input` needs to be editted to output a random key as well as the dataset. Aggregation is quite fiddly in the IMNN due to the need to properly slice the data - the fiducial datasets are transformed into a numpy iterator via a TensorFlow dataset, we need to therefore augment the normal dataset to return a key as well. For every XLA device used for aggregation then there is a separate dataset with `n_s // (n_devices * n_per_device)` yields with `n_per_device` elements. Our generator therefore must iterate through the dataset and return corresponding keys before resetting. To do this we will generate `n_s` keys and define a generator which takes in this list of keys and the dataset and yield the next iteration of data and the `i`th iteration of keys (which resets after `n_s // (n_devices * n_per_device)` iterations). This is true for the fiducial dataset, but the derivative dataset needs repeated keys for every parameter direction used for the numerical derivative. For this `n_d` of the keys are repeated `2 * n_params` times before being reshaped and returned by the same generator. With these generators made we now just need to change the `fn` function of `get_summary` such that it splits apart the signal data (called `d`) from the key, and we then use the key to grab a random integer between 0 and the amount of noise passed to the class and use that integer to get noise which is then added to the signal before the input to the neural network. Because of the way the generators are constructed for the derivative dataset the same noise will be added to each set of simulations for a single derivative.
class NoiseIMNN(AggregatedNumericalGradientIMNN):
def __init__(self, noise, **kwargs):
super().__init__(**kwargs)
self.noise = noise
self.n_noise = self.noise.shape[0]
def get_summary(self, inputs, w, θ, derivative=False, gradient=False):
def fn(inputs, w):
d, key = inputs
ϵ = self.noise[jax.random.randint(key, (), minval=0, maxval=self.n_noise)]
return self.model(w, d + ϵ)
if gradient:
dΛ_dx, d = inputs
dx_dw = jax.jacrev(fn, argnums=1)(d, w)
return self._construct_gradient(dx_dw, aux=dΛ_dx, func="einsum")
else:
return fn(inputs, w)
def _collect_input(self, key, validate=False):
def generator(dataset=None, key=None, total=None):
i = 0
while i < total:
yield next(dataset), key[i]
i += 1
i = 0
if validate:
fiducial = self.validation_fiducial
derivative = self.validation_derivative
else:
fiducial = self.fiducial
derivative = self.derivative
keys = np.array(jax.random.split(key, num=self.n_s))
return (
[partial(generator, dataset=fid, key=key, total=key.shape[0])()
for fid, key in zip(fiducial, keys.reshape(self.fiducial_batch_shape + (2,)))],
[partial(generator, dataset=der, key=key, total=key.shape[0])()
for der, key in zip(derivative, np.repeat(keys[:n_d], 2 * self.n_params, axis=0).reshape(
self.derivative_batch_shape + (2,)))])
# We're going to use jax's stax module to build a simple network with three hidden layers each with 128 neurons and which are activated by leaky relu before outputting the two summaries. The optimiser will be a jax Adam optimiser with a step size of 0.001.
model = stax.serial(
stax.Dense(128),
stax.LeakyRelu,
stax.Dense(128),
stax.LeakyRelu,
stax.Dense(128),
stax.LeakyRelu,
stax.Dense(n_summaries),
)
optimiser = optimizers.adam(step_size=1e-3)
# The NoiseIMNN can now be initialised setting up the network and the fitting routine (as well as the plotting function), where we will use the CPU as the host device and use the GPUs for calculating the summaries and say that we know that we can process 100 simulations at a time per device before running out of memory.
rng, key = jax.random.split(rng)
imnn = NoiseIMNN(
noise=ϵ,
n_s=n_s,
n_d=n_d,
n_params=n_params,
n_summaries=n_summaries,
input_shape=input_shape,
θ_fid=m_fid,
model=model,
optimiser=optimiser,
key_or_state=key,
fiducial=mx,
derivative=mx_mp,
δθ=δm,
host=jax.devices("cpu")[0],
devices=jax.devices(),
n_per_device=100,
validation_fiducial=validation_mx,
validation_derivative=validation_mx_mp)
# To set the scale of the regularisation we use a coupling strength 𝜆 whose value should mean that the determinant of the difference between the covariance of network outputs and the identity matrix is larger than the expected initial value of the determinant of the Fisher information matrix from the network. How close to the identity matrix the covariance should be is set by 𝜖 . These parameters should not be very important, but they will help with convergence time. Fitting can then be done simply by calling:
rng, key = jax.random.split(rng)
imnn.fit(λ=10., ϵ=0.1, rng=key, print_rate=1)
imnn.plot();
# ## Inference
#
# Now lets say we want to infer the slope of some data we can use this trained IMNN and the ABC module (with some premade simulations). I'm going to first generate some data to infer:
# +
rng, key = jax.random.split(rng)
target_m = 3.
y_target = (target_m * x + np.sqrt(0.1)
* jax.random.normal(key, shape=input_shape))
# -
# We'll make a bunch of new simulations for the ABC too. First we'll draw many (5000) different values of slopes and calculate $mx$ for each of these slopes. Then we'll sum all of the expensive noise realisations to every example to get 2500000 different realisations of noise and signal. We'll compress all of these using the IMNN and also repeat the parameter values used to make these simulations (because there are now 500 examples of each parameter due to the noise realisations).
# +
rng, key = jax.random.split(rng)
m_ABC = np.expand_dims(
jax.random.uniform(
key, minval=0., maxval=10., shape=(5000,)),
1)
y_ABC = np.einsum(
"ij,kj->ikj",
m_ABC * x,
ϵ).reshape((-1,) + input_shape)
parameters = np.repeat(m_ABC, n_noise, axis=0)
summaries = imnn.get_estimate(y_ABC)
# -
# We'll use TensorFlow Probability to define a prior distribution between zero and 10 for the slope
prior = tfp.distributions.Independent(
tfp.distributions.Uniform(low=[0.], high=[10.]),
reinterpreted_batch_ndims=1)
prior.low = np.array([0.])
prior.high = np.array([10.])
# We'll then initialise the ApproximateBayesianComputation module using the IMNN as the compression function for `y_target`
ABC = ApproximateBayesianComputation(
target_data=y_target,
prior=prior,
simulator=None,
compressor=imnn.get_estimate,
gridsize=100,
F=imnn.F)
# We can then run the ABC with an $\epsilon=0.1$ with the compressed examples with corresponding parameter values
ABC(ϵ=0.1, parameters=parameters, summaries=summaries);
# Plotting the accepted parameters gives us the posterior distribution of the value of possible slopes. We can also plot the value of the slope used to generate the target data for completeness.
plt.hist(ABC.parameters.accepted[0][:, 0],
range=[0, 10],
bins=25,
density=True)
plt.axvline(target_m, linestyle="dashed", color="black")
plt.xlabel(r"$m$");
| examples/additive_noise.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.2
# language: julia
# name: julia-1.7
# ---
# # Solving differential equations in Julia
#
# ## Define your model and find parameters
#
# The concentration of a decaying nuclear isotope could be described as an exponential decay:
#
# $$
# \frac{d}{dt}C(t) = - \lambda C(t)
# $$
#
# **State variable**
# - $C(t)$: The concentration of a decaying nuclear isotope.
#
# **Parameter**
# - $\lambda$: The rate constant of decay. The half-life $t_{\frac{1}{2}} = \frac{ln2}{\lambda}$
#
# Take a more complex model, the spreading of an contagious disease can be described by the [SIR model](https://www.maa.org/press/periodicals/loci/joma/the-sir-model-for-spread-of-disease-the-differential-equation-model):
#
# $$
# \begin{align}
# \frac{d}{dt}S(t) &= - \beta S(t)I(t) \\
# \frac{d}{dt}I(t) &= \beta S(t)I(t) - \gamma I(t) \\
# \frac{d}{dt}R(t) &= \gamma I(t)
# \end{align}
# $$
#
# **State variables**
#
# - $S(t)$ : the fraction of susceptible people
# - $I(t)$ : the fraction of infectious people
# - $R(t)$ : the fraction of recovered (or removed) people
#
# **Parameters**
#
# - $\beta$ : the rate of infection when susceptible and infectious people meet
# - $\gamma$ : the rate of recovery of infectious people
# ## Make a solver by yourself
#
# ### Forward Euler method
#
# The most straightforward approach to numerically solve differential equations is the forward Euler's (FE) method[^Euler].
#
# In each step, the next state variables ($\vec{u}_{n+1}$) is accumulated by the product of the size of time step (dt) and the derivative at the current state ($\vec{u}_{n}$):
#
# $$
# \vec{u}_{n+1} = \vec{u}_{n} + dt \cdot f(\vec{u}_{n}, t_{n})
# $$
# +
# The ODE model. Exponential decay in this example
# The input/output format is compatible to Julia DiffEq ecosystem
expdecay(u, p, t) = p * u
# Forward Euler stepper
step_euler(model, u, p, t, dt) = u .+ dt .* model(u, p, t)
# In house ODE solver
function mysolve(model, u0, tspan, p; dt=0.1, stepper=step_euler)
# Time points
ts = tspan[1]:dt:tspan[end]
# State variable at those time points
us = zeros(length(ts), length(u0))
# Initial conditions
us[1, :] .= u0
# Iterations
for i in 1:length(ts)-1
us[i+1, :] .= stepper(model, us[i, :], p, ts[i], dt)
end
# Results
return (t = ts, u = us)
end
tspan = (0.0, 2.0)
p = -1.0
u0 = 1.0
sol = mysolve(expdecay, u0, tspan, p, dt=0.1, stepper=step_euler)
# Visualization
using Plots
Plots.gr(lw=2)
# Numericalsolution
plot(sol.t, sol.u, label="FE method")
# True solution
plot!(x -> exp(-x), 0.0, 2.0, label="Analytical solution")
# +
# SIR model
function sir(u, p ,t)
s, i, r = u
β, γ = p
v1 = β * s * i
v2 = γ * i
return [-v1, v1-v2, v2]
end
p = (β = 1.0, γ = 0.3)
u0 = [0.99, 0.01, 0.00] # s, i, r
tspan = (0.0, 20.0)
sol = mysolve(sir, u0, tspan, p, dt=0.5, stepper=step_euler)
plot(sol.t, sol.u, label=["S" "I" "R"], legend=:right)
# -
# ### The fourth order Runge-Kutta (RK4) method
#
# One of the most popular ODE-solving methods is the fourth order Runge-Kutta ([RK4](https://en.wikipedia.org/wiki/Runge%E2%80%93Kutta_methods)) method.
#
# In each step, the next state is calculated in 5 steps, 4 of which are intermediate steps.
#
# $$
# \begin{align}
# k_1 &= dt \cdot f(\vec{u}_{n}, t_n) \\
# k_2 &= dt \cdot f(\vec{u}_{n} + 0.5k_1, t_n + 0.5dt) \\
# k_3 &= dt \cdot f(\vec{u}_{n} + 0.5k_2, t_n + 0.5dt) \\
# k_4 &= dt \cdot f(\vec{u}_{n} + k_3, t_n + dt) \\
# u_{n+1} &= \vec{u}_{n} + \frac{1}{6}(k_1 + 2k_2 + 2k_3 + k_4)
# \end{align}
# $$
#
# In homework 1, you are going to replace the Euler stepper with the RK4 one:
#
# ```julia
# step_rk4(f, u, p, t, dt) = """TODO"""
# ```
# ## Using DifferentialEquations.jl
#
# Documentation: <https://diffeq.sciml.ai/dev/index.html>
using Plots, DifferentialEquations
Plots.gr(linewidth=2)
# ### Exponential decay model
# +
# Parameter of exponential decay
p = -1.0
u0 = 1.0
tspan = (0.0, 2.0)
# Define a problem
prob = ODEProblem(expdecay, u0, tspan, p)
# Solve the problem
sol = solve(prob)
# Visualize the solution
plot(sol, legend=:right)
# -
# ### SIR model
# +
# Parameters of the SIR model
p = (β = 1.0, γ = 0.3)
u0 = [0.99, 0.01, 0.00] # s, i, r
tspan = (0.0, 20.0)
# Define a problem
prob = ODEProblem(sir, u0, tspan, p)
# Solve the problem
sol = solve(prob)
# Visualize the solution
plot(sol, label=["S" "I" "R"], legend=:right)
# -
plot(sol, vars=(0, 2), legend=:right)
plot(sol, vars=(1, 2), legend=:right)
# ## Using ModelingToolkit.jl
#
# [ModelingToolkit.jl](https://mtk.sciml.ai/dev/) is a high-level package for symbolic-numeric modeling and simulation ni the Julia DiffEq ecosystem.
using DifferentialEquations
using ModelingToolkit
using Plots
Plots.gr(linewidth=2)
# ### Exponential decay model
# +
@parameters λ # Decaying rate constant
@variables t C(t) # Time and concentration
D = Differential(t) # Differential operator
# Make an ODE system
@named expdecaySys = ODESystem([D(C) ~ -λ*C ])
# +
u0 = [C => 1.0]
p = [λ => 1.0]
tspan = (0.0, 2.0)
prob = ODEProblem(expdecaySys, u0, tspan, p)
sol = solve(prob)
plot(sol)
# -
# ### SIR model
# +
@parameters β γ
@variables t s(t) i(t) r(t)
D = Differential(t) # Differential operator
# Make an ODE system
@named sirSys = ODESystem(
[D(s) ~ -β * s * i,
D(i) ~ β * s * i - γ * i,
D(r) ~ γ * i])
# +
# Parameters of the SIR model
p = [β => 1.0, γ => 0.3]
u0 = [s => 0.99, i => 0.01, r => 0.00]
tspan = (0.0, 20.0)
prob = ODEProblem(sirSys, u0, tspan, p)
sol = solve(prob)
plot(sol)
# -
# ## Using Catalyst.jl
#
# [Catalyst.jl](https://github.com/SciML/Catalyst.jl) is a domain-specific language (DSL) package to solve "law of mass action" problems.
using Catalyst
using DifferentialEquations
using Plots
Plots.gr(linewidth=2)
# ### Exponential decay model
decayModel = @reaction_network begin
λ, C --> 0
end λ
# +
p = [1.0]
u0 = [1.0]
tspan = (0.0, 2.0)
prob = ODEProblem(decayModel, u0, tspan, p)
sol = solve(prob)
plot(sol)
# -
# ### SIR model
sirModel = @reaction_network begin
β, S + I --> 2I
γ, I --> R
end β γ
# +
# Parameters of the SIR model
p = (1.0, 0.3)
u0 = [0.99, 0.01, 0.00]
tspan = (0.0, 20.0)
prob = ODEProblem(sirModel, u0, tspan, p)
sol = solve(prob)
plot(sol, legend=:right)
| docs/intro/03-diffeq.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Kernel methods are widespread in machine learning and they were particularly common before deep learning became a dominant paradigm. The core idea is to introduce a new notion of distance between high-dimensional data points by replacing the inner product $(x_i, x_j)$ by a function that retains many properties of the inner product, yet which is nonlinear. This function $k(.,.)$ is called a kernel. Then, in many cases, wherever a learning algorithm would use an inner product, the kernel function is used instead.
#
# The intuition is that the kernel function acts as an inner product on a higher dimensional space and encompasses some $\phi(.)$ mapping from the original space of the data points to this space. So intuitively, the kernel function is $k(x_i, x_j)=(\phi(x_i), \phi(x_j))$. The hope is that points that were not linearly separable in the original space become linearly separable in the higher dimensional space. The $\phi(.)$ function may map to an infinite dimensional space and it does not actually have to be specified. As long as the kernel function is positive semidefinite, the idea works.
#
# Many kernel-based learning algorithms are instance-based, which means that the final model retains some or all of the training instances and they play a role in the actual prediction. Support vector machines belong here: support vectors are the training instances which are critically important in defining the boundary between two classes. Some important kernels are listed below.
#
# | Name | Kernel function|
# |------|-----------------|
# |Linear | $(x_i,x_j)$|
# |Polynomial| $((x_i,x_j)+c)^d$|
# |Radial basis function|$\exp(-\gamma\|x_i-x_j\|^2)$|
#
# The choice of kernel and the parameters of the kernel are often arbitrary and either some trial and error on the dataset or hyperparameter optimization helps choose the right combination. Quantum computers naturally give rise to certain kernels and it is worth looking at a specific variant of how it is constructed.
#
# # Thinking backward: learning methods based on what the hardware can do
#
# Instead of twisting a machine learning algorithm until it only contains subroutines that have quantum variants, we can reverse our thinking and ask: given a piece of quantum hardware and its constraints, can we come up with a new learning method? For instance, interference is a very natural thing to do: we showed an option in the first notebook on quantum states, and it can also be done with a Hadamard gate.
# For this to work we need to encode both training and testvectors as amplitudes in a statevector built up out of four registers:
#
# $|0\rangle_c|00..0\rangle_m|00..0\rangle_i|0\rangle_a$
#
# The amplitude of such state will be equal to the value of a feature in a training vector or test vector. To do that we use four registers. The first is a single bit, acting as the ancilla ancilla (a), which will will code for either a training (a=0) or a testvector (a=1). The second register, in the notebook example a single bit, will code for the m-th training vector. The third register, in the notebook example also reduced to a single bit, codes for the i-th feature. Lastly the class bit (c) codes for class -1 (c=0), or 1 (c=1).
# Hence, if after fully encoding all training data and test data into the state $|\psi>$ the state |1010> has coefficient 0.46 :
#
# $|\psi\rangle\ = ....+ 0.46|1010\rangle +....$ ,
#
# Then that implies that the second feature (i=1) of the first (m=0) training vector (a=0), which classifies as class 1 (c=1), has the value 0.46. Note, we assume both training vectors and test vector are normalized.
#
# In a more general expression we can write for a fully encoded state (NB we arrange the order of the registers to be consistent with the code below):
#
# $|\psi\rangle = \frac{1}{\sqrt{2M}}\sum_{m=0}^{M-1}|y_m\rangle|m\rangle|\psi_{x^m}\rangle|0\rangle + |y_m\rangle|m\rangle|\psi_{\tilde{x}}\rangle|1\rangle$
#
# with:
#
# $|\psi_{x^m}\rangle = \sum_{i=0}^{N-1}x_i^m|i\rangle, \; |\psi_{\tilde{x}}\rangle = \sum_{i=0}^{N-1}\tilde{x_i}|i\rangle. \quad$ N being equal to the number of features in the the training and test vectors
#
# As the last summation is independent on m, there will M copies of the test vector in the statevector, one for every training vector.
#
#
# We now only need to apply a Hadamard gate to the ancilla to interfere the test and training instances. Measuring and post-selecting on the ancilla gives rise to a kernel [[1](#1)].
#
# Let's start with initializations:
#
#
#
#
#
#
#
#
#
# +
from qiskit import ClassicalRegister, QuantumRegister, QuantumCircuit
from qiskit import execute
from qiskit import BasicAer as Aer
import numpy as np
# %matplotlib inline
np.set_printoptions(precision = 3)
q = QuantumRegister(4)
c = ClassicalRegister(4)
backend = Aer.get_backend('qasm_simulator')
# -
# We are constructing an instance-based classifier: we will calculate a kernel between all training instances and a test example. In this sense, this learning algorithm is lazy: no actual learning happens and each prediction includes the entire training set.
#
# As a consequence, state preparation is critical to this protocol. We have to encode the training instances in a superposition in a register, and the test instances in another register. Consider the following training instances of the [Iris dataset](https://archive.ics.uci.edu/ml/datasets/iris): $S = \{(\begin{bmatrix}0 \\ 1\end{bmatrix}, 0), (\begin{bmatrix}0.790 \\ 0.615\end{bmatrix}, 1)\}$, that is, one example from class 0 and one example from class 1. Furthermore, let's have two test instances, $\{\begin{bmatrix}-0.549\\ 0.836\end{bmatrix}, \begin{bmatrix}0.053 \\ 0.999\end{bmatrix}\}$. These examples were cherry-picked because they are relatively straightforward to prepare.
training_set = [[0, 1], [0.79, 0.615]]
labels = [0, 1]
test_set = [[-0.549, 0.836], [0.053 , 0.999]]
# To load the data vectors, we use amplitude encoding as explained above, which means that, for instance, the second training vector will be encoded as $0.78861006|0\rangle + 0.61489363|1\rangle$. Preparing these vectors only needs a rotation, and we only need to specify the corresponding angles. The first element of the training set does not even need that: it is just the $|1\rangle$ state, so we don't specify an angle for it.
# To get the angle we need to solve the equation $a|0\rangle + b|1\rangle=\cos\left(\frac{\theta}{2}\right)|0\rangle + i \sin \left(\frac{\theta}{2}\right) |1\rangle$. Therefore, we will use $\theta=2 \arccos(a)$
def get_angle(amplitude_0):
return 2*np.arccos(amplitude_0)
# In practice, the state preparation procedure we will consider requires the application of several rotations in order to prepare each data point in the good register. Don't hesitate to check it by yourself by running the circuit below with a pen and paper.
# The following function builds the circuit. We plot it and explain it in more details below.
def prepare_state(q, c, angles):
ancilla_qubit = q[0]
index_qubit = q[1]
data_qubit = q[2]
class_qubit = q[3]
circuit = QuantumCircuit(q, c)
# Put the ancilla and the index qubits into uniform superposition
circuit.h(ancilla_qubit)
circuit.h(index_qubit)
circuit.barrier()
# Prepare the test vector
circuit.cu3(angles[0], 0, 0, ancilla_qubit, data_qubit)
# Flip the ancilla qubit > this moves the input
# vector to the |0> state of the ancilla
circuit.x(ancilla_qubit)
circuit.barrier()
# Prepare the first training vector
# [0,1] -> class 0
# We can prepare this with a Toffoli
circuit.ccx(ancilla_qubit, index_qubit, data_qubit)
# Flip the index qubit > moves the first training vector to the
# |0> state of the index qubit
circuit.x(index_qubit)
circuit.barrier()
# Prepare the second training vector
# [0.790, 0.615] -> class 1
#
# Ideally we would do this with a double controlled, i.e a ccry, gate
# However in qiskit we cannot build such a gate, hence we resort to
# the following construction
circuit.ccx(ancilla_qubit, index_qubit, data_qubit)
circuit.ry(-angles[1], data_qubit)
circuit.ccx(ancilla_qubit, index_qubit, data_qubit)
circuit.ry(angles[1], data_qubit)
circuit.barrier()
# Flip the class label for training vector #2
circuit.cx(index_qubit, class_qubit)
return circuit
# Let us see the circuit for the distance-based classifier:
# +
from qiskit.tools.visualization import circuit_drawer
# Compute the angles for the testvectors
test_angles = [get_angle(test_set[0][0]), get_angle(test_set[1][0])]
# Compute the angle for the second training vector (as the first vector is trivial)
training_angle = get_angle(training_set[1][0])/2
angles = [test_angles[0], training_angle]
state_preparation_0 = prepare_state(q, c, angles)
circuit_drawer(state_preparation_0, output='mpl')
# -
# The vertical lines are barriers to make sure that all gates are finished by that point. They also make a natural segmentation of the state preparation.
#
# In the first section, the ancilla and index qubits are put into uniform superposition.
#
# The second section entangles the test vector with the ground state of the ancilla.
#
# In the third section, we prepare the state $|1\rangle$, which is the first training instance, and entangle it with the excited state of the ancilla and the ground state of the index qubit with a Toffoli gate and a Pauli-X gate. The Toffoli gate is also called the controlled-controlled-not gate, describing its action.
#
# The fourth section prepares the second training instance and entangles it with the excited state of the ancilla and the index qubit. Next, the class qubit is flipped conditioned on the index qubit being $|1\rangle$. This creates the connection between the encoded training instances and the corresponding class label.
#
# Let's dissect the last part where we prepare the second training state, which is $\begin{pmatrix}0.790 \\ 0.615\end{pmatrix}$ and we entangle it with the excited state of the ancilla and the excited state of the index qubit. We use `angles[1]`, which is ~`1.325/2`. Why? We have to rotate the basis state $|0\rangle$ to contain the vector we want. We could write this generic state as $\begin{pmatrix} \cos(\theta/2) \\ \sin(\theta/2)\end{pmatrix}$. Looking at the documentation of the gate implementing the rotation, you'll see that the function argument divides the angle by two, so we have to adjust for that -- this is why we divided $\theta$ by two. If you calculate the arccos or arcsin values, you will get the value in `angles[1]`.
#
#
#
# We need to apply the rotation to data qubit only if ancilla AND index qubits are 1, in other words, we have to implement a double controlled rotation. Qiskit does not have this type of gate. Hence, we'll build it in two stages of half the required angle, designed in such a way they either add or cancel. The quantum AND gate is the CCX (also known as Toffoli), which flips the target qubit if both controls are 1. Applying the CCX flips only the data qbit of the target state. The subsequent rotation over half the desired angle works on all states, but after applying the second CCX the targed state has actually rotated in the opposite direction. Applying the reverse rotation adds the second half of the rotation for the target state,ans cancels the rotation for all other states (See Bloch sphere diagram outlining these 4 steps)
#
#
# 
#
#
# <br>
# Let's now see what final state the circuit has produced. Note, the print is a non-normalized statevector:
val = ['Xtest x', 'Xtrn0 x','','','Xtest y','Xtrn0 y','','','','','Xtest x','Xtrn1 x','','','Xtest y','Xtrn1 y']
res = execute(state_preparation_0, Aer.get_backend('statevector_simulator')).result()
outp = 2* np.array(np.real(res.get_statevector(state_preparation_0)))
print('Statevector after insertion of data and testvectors\n\ncdia coefficient')
for z in range(outp.shape[0]):
print(format(z, '04b'),' % 5.4f ' %(round(outp[z],3)), val[z])
# From the table you can see how both the test vector (Xtst x, Xtsty), as well as the training vectors ((Xtrn0 x, Xtrn0 y) - class0) and ((Xtrn1 x,Xtrn1 y) - class1) are embedded in the state vector. The training vector class is indicated in the class bit (c). The test vector is coded by the 0-state of the ancilla (a), and the training vector is coded by the 1-state of the ancilla. Note also the data bit (d) coding for the value of the x or y feature of the training vectors, and the index bit (i) coding for training vector 1 or 2.
#
# We are now ready for the final step
# # A natural kernel on a shallow circuit
#
# Having done the state preparation, the actual prediction is nothing but a Hadamard gate applied on the ancilla, followed by measurements. Since the ancilla is in a uniform superposition at the end of the state preparation and it is entangled with the registers encoding the test and training instances, applying a second Hadamard on the ancilla interferes the entangled registers. The state before the measurement is $\frac{1}{2\sqrt{M}}\sum_{m=0}^{M-1}|y_m\rangle|m\rangle(|\psi_{x^m}\rangle+|\psi_{\tilde{x}}\rangle)|0\rangle+|y_m\rangle|m\rangle(|\psi_{x^m}\rangle-|\psi_{\tilde{x}}\rangle)|1\rangle$, where $|\psi_{\tilde{x}}\rangle$ is the encoded test instance and $\psi_{x^m}\rangle$ is the m-th training instance. For our example M, the number of training samples, equals 2.
#
#
def interfere_data_and_test_instances(circuit, q, c, angles):
circuit.h(q[0])
circuit.barrier()
circuit.measure(q, c)
return circuit
# If we measure the ancilla, the outcome probability of observing 0 will be $\frac{1}{4M}\sum_{i=0}^{M-1} |\tilde{x} + x_m|^2$. This creates a kernel of the following form:
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
x = np.linspace(-2, 2, 100)
plt.xlim(-2, 2)
plt.ylim(0, 1.1)
plt.plot(x, 1-x**2/4)
# This is the kernel that performs the classification. We perform the post-selection on observing 0 on the measurement on the ancilla and calculate the probabilities of the test instance belonging to either class:
def postselect(result_counts):
total_samples = sum(result_counts.values())
# define lambda function that retrieves only results where the ancilla is in the |0> state
post_select = lambda counts: [(state, occurences) for state, occurences in counts.items() if state[-1] == '0']
# perform the postselection
postselection = dict(post_select(result_counts))
postselected_samples = sum(postselection.values())
ancilla_post_selection = postselected_samples/total_samples
print('Ancilla post-selection probability was found to be ',round(ancilla_post_selection,3))
retrieve_class = lambda binary_class: [occurences for state, occurences in postselection.items() if state[0] == str(binary_class)]
prob_class0 = sum(retrieve_class(0))/postselected_samples
prob_class1 = sum(retrieve_class(1))/postselected_samples
print('Probability for class 0 is', round(prob_class0,3))
print('Probability for class 1 is', round(prob_class1,3))
# For the first instance we have:
circuit_0 = interfere_data_and_test_instances(state_preparation_0, q, c, angles)
job = execute(circuit_0, backend)
result = job.result()
count = result.get_counts(circuit_0)
print(count)
postselect(count)
# And for the second one:
angles = [test_angles[1], training_angle]
state_preparation_1 = prepare_state(q, c, angles)
circuit_1 = interfere_data_and_test_instances(state_preparation_1, q, c, angles)
job = execute(circuit_1, backend)
result = job.result()
count = result.get_counts(circuit_1)
print(count)
postselect(count)
# # References
#
# [1] <NAME>, <NAME>, <NAME>. (2017). [Implementing a distance-based classifier with a quantum interference circuit](https://doi.org/10.1209/0295-5075/119/60002). *Europhysics Letters*, 119(6), 60002. <a id='1'></a>
| qiskit_version/11_Kernel_Methods.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Vezérlési szerkezetek
#
# ---
#
# (2019. 11. 05. – 6. óra)
#
# <NAME>
# + [markdown] slideshow={"slide_type": "slide"}
# # Bevezető
#
# 1. szekvencia
# 1. elágazás
# 1. ciklus
# + [markdown] slideshow={"slide_type": "slide"}
# # 1. Szekvencia
#
# Utasítások egymásutánja:
#
# ```py
# utasitas1()
# utasitas2()
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# # 2. Elágazás
#
# Feltétel teljesülése esetén az egyik, nem teljesülése esetén egy másik utasítás fut le:
#
# ```py
# if ch in 'aeiou':
# print('mgh')
# else:
# print('msh')
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# # 2. Elágazás (folytatás)
#
# Az `else` ág opcionális (ha a feltétel nem teljesül, nem történik semmi):
#
# ```py
# if uj_fizetes > 147000:
# allast_valtok()
# ```
# + [markdown] slideshow={"slide_type": "fragment"}
# Lehet kettőnél több ágat is bevezetni:
#
# ```py
# if ch in 'aeiou':
# print('mgh')
# elif ch in 'bcdfghjklmnpqrstvwxyz':
# print('msh')
# else:
# print('nem betű')
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# # 3. Ciklus
#
# Ugyan azt az utasítás sokszor hajtjuk végre:
#
# - amíg egy feltétel teljesül: `while`
# - egy gyűjteményes adattípus minden elemére: `for`
# + [markdown] slideshow={"slide_type": "subslide"}
# # 3.1. *while*-ciklus
#
# Álláskeresés:
#
# ```py
# while fizetes < 200000:
# allast_keresek()
# ```
# + [markdown] slideshow={"slide_type": "fragment"}
# Számok kiírása:
#
# ```py
# x = 0
# while x < 10:
# x += 1
# print(x)
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# # 3.2. *for*-ciklus
#
# Lista / tuple / halmaz bejárása:
#
# ```py
# for i in container:
# print(i)
# ```
# + [markdown] slideshow={"slide_type": "fragment"}
# Szótár kulcsainak bejárása:
#
# ```py
# for key in mydict:
# print(key, mydict[key])
# ```
# + [markdown] slideshow={"slide_type": "fragment"}
# Szótár kulcs-érték párjainak bejárása:
#
# ```py
# for k, v in mydict.items():
# print(k, v)
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# # 3.3. *continue*, *break*
#
# `continue`: Ugrás a ciklus következő lépésére
#
# ```py
# for word in 'alma alma piros alma'.split():
# if word.startswith('a'):
# continue
# print(word)
# ```
# + [markdown] slideshow={"slide_type": "fragment"}
# `break`: kilépés a ciklusból
#
# ```py
# for word in 'alma alma piros alma'.split():
# if word == 'piros':
# break
# print(word)
# ```
# + [markdown] slideshow={"slide_type": "skip"}
# # programozási tetelek
#
# 1. összegzés
# - közvetlenül (számok egy listában)
# - közvetve (szavak hossza --> átlag)
# 1. számlálás: hány elemre teljesül egy tulajdonság?
# 1. maximum-keresés: leghosszabb szó
# 1. feltételes maximum-keresés: leghosszabb magánhangzóval kezdődő szó
# 1. lineáris keresés: keressük meg az első t-betűvel kezdődő szót egy szövegben
# 1. logaritmikus keresés: keressük meg az első t-betűvel kezdőső szót egy rendezett listában
# 1. rekurzió
# + [markdown] slideshow={"slide_type": "slide"}
# # Feladatok
#
# 1. Legyen egy számokat tartalmazó listánk. Adjuk össze az elemeit!
# 1. Legyen egy számokat tartalmazó listánk. Mennyi a számok átlaga?
# 1. Legyen egy szavakat tartalmazó listánk. Mekkora az átlagos szóhossz?
#
# ---
#
# 1. Legyen egy szavakat tartalmazó listánk. Hány szó kezdődik 'a'-val?
# 1. Legyen egy szavakat tartalmazó listánk. Hány szó kezdődik magánhangzóval?
#
# ---
#
# 1. Legyen egy szavakat tartalmazó listánk. Mely szavak kezdődnek 'a'-val?
# 1. Legyen egy szavakat tartalmazó listánk. Mely szavak kezdődnek magánhangzóval?
#
# ---
#
# 1. Legyen egy szövegünk. Melyik a leghosszabb szava?
# 1. Legyen egy szövegünk. Melyik a legtöbbféle betűt tartalmazó szava?
# 1. Legyen egy szövegünk. Melyik a leghosszabb, magánhangzóval kezdődő szó?
#
# + [markdown] slideshow={"slide_type": "subslide"}
# # További feladatok
#
# * Mi a közös a fenti feladatcsoportokban?
# * Legyen egy egész számokat tartalmazó listánk. Növeljük meg a lista minden elemét eggyel!
# * Legyen egy szövegünk. Számoljunk szógyakoriságot. Az eredményt tároljuk egy szótárban (`{szó: előfordulás}`).
# * Próbáljuk meg kiiratni a szótár elemeit gyakoriság szerint csökkenő sorrendben!
# * Írjuk ki a relatív gyakoriságot!
# + [markdown] slideshow={"slide_type": "slide"}
# # De jó lenne, ha ma eljutnánk idáig! (Vége.)
| 06.ctrl/ctrl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Scikit-Learn 소개
#
# - 다양한 머신 러닝 알고리즘을 구현하여 간단한 API로 제공하는 Python 라이브러리
# - 다양한 데이터집합 제공 :
# sklearn.datasets의 load_* ()함수
# - 데이터에 대한 모델을 평가하기 위해 학습 세트 와 테스트 세트 로 분리 :
# sklearn.model_selection의 train_test_split()함수
# - 다양한 머신러닝 알고리즘 제공 :
# - linear_model인 linear regression알고리즘 모듈
# from sklearn.linear_model import LinearRegressionsklearn.linear_model
# - naive_bayes의 GaussianNB 알고리즘 모듈
# from sklearn.naive_bayes import GaussianNB
# - neighbors의 KNeighborsClassifier 알고리즘 모듈
# from sklearn.neighbors import KNeighborsClassifier
# - 모델링
# - 모델 객체를 생성하여 학습데이터로 모델링
# model = LinearRegression(fit_intercept=True) #모델객체 생성
# model.fit(X, y) # 학습 데이터 X,y를 이용하여 모델링
# - 예측
# 새로운 데이터(nX)를 모델링 결과로 얻은 모델로 결과(ny) 예측
# ny = model.predict(nX)
# - 예측 성능확인
# sklearn.metrics의 accuracy_score()함수
#
# https://scikit-learn.org/stable/
#
# https://jakevdp.github.io/PythonDataScienceHandbook/05.02-introducing-scikit-learn.html
# ### (1)Supervised learning example: Simple linear regression
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
#(1)Scikit-Learn의 선형모델 linear regression 모듈 가져오기
from sklearn.linear_model import LinearRegression
#(2) 모델 객체 생성
model = LinearRegression(fit_intercept=True)
model
#(3) 데이터 준비
# 지도학습을 위한 학습 데이터 준비
# 데이터는 입력(x,feature)과 출력(y,target)으로 구성
# 50개의 랜덤 값 생성
rng = np.random.RandomState(10)
x = 10 * rng.rand(50)
y = 2 * x - 1 + rng.randn(50)
plt.scatter(x, y);
#(4) 데이터 구조 변경
# 데이터 구조
# 입력(x) : 2차원 특징(feature) 행렬
# 출력(y) : 1차원 목표(target) 벡터
X = x[:, np.newaxis]
X.shape
#(5)데이터를 이용하여 linear regression 모델링
#데이터로 얻은 선형 함수의 기울기(model.coef_)와 절편(model.intercept_)을 확인
model.fit(X, y)
print("model.coef_=", model.coef_)
print("model.intercept_ = ", model.intercept_)
#(6)새로운 입력(nx)에 대한 출력(ny) 예측
nx = np.linspace(-1, 11)
nX = nx[:, np.newaxis]
ny = model.predict(nX)
#(7)예측값에 대한 모델 (선형함수) 출력
plt.plot(nx, ny, color="orange" )
# -
# 모델 (선형 함수)
# $\hat{y}(w, x) = w_0 + w_1 x_1 + ... + w_p x_p$
#
# $w = (w_1,..., w_p)$ : model.coef_
# $w_0$ : model.intercept_.
#
# ### (2)Supervised learning example: Iris classification
#
# algorithm : Gaussian naive Bayes
# +
# (1)scikit learn 제공 iris 데이터 셋 가져오기
from sklearn.datasets import load_iris
iris = load_iris()
#iris는 key-value로 구성된 dictionary 와 유사한 bounch class
# dataset key값 확인
print("iris_dataset의 key: {}".format(iris.keys()))
#데이터 설명을 포함한 'DESCR' key의 value 확인
print(iris['DESCR'][:193] + "\n...")
#데이터 셋 확인
print("타깃의 이름: {}".format(iris['target_names']))
print("특성의 이름: {}".format(iris['feature_names']))
print("data의 타입: {}".format(type(iris['data']))) #x, features
print("data의 크기: {}".format(iris['data'].shape))
print("data의 처음 다섯 행:\n{}".format(iris['data'][:5]))
print("target의 타입: {}".format(type(iris['target'])))
print("타깃:\n{}".format(iris['target'])) #y, target, class
# 모델링을 위한 x, y 값 설정
irisX = iris['data']
irisy = iris['target']
print(irisX.shape)
# +
# (2)데이터에 대한 모델을 평가하기위해 학습 세트 와 테스트 세트 로 분리
from sklearn.model_selection import train_test_split
Xtrain, Xtest, ytrain, ytest = train_test_split(irisX, irisy,
random_state=1)
#(3)Modeling
# 학습데이터로 GaussianNB 알고리즘으로 모델링
from sklearn.naive_bayes import GaussianNB #GaussianNB 모듈 가져오기
model = GaussianNB() # 모델 객체 생성
model.fit(Xtrain, ytrain) #학습데이터를 이용하여 모델링
y_model = model.predict(Xtest) # 새로운 데이터 예측
#(4)예측결과에 대한 성능 확인
from sklearn.metrics import accuracy_score
accuracy_score(ytest, y_model)
# -
# ### (3)Unsupervised learning example: Iris dimensionality
#
# algorithm : PCA
# +
#(1) PCA 모델링
from sklearn.decomposition import PCA # PCA 모듈 가져오기
model = PCA(n_components=2) # PCA 모델객체 생성
model.fit(irisX) # iris 입력데이터로 차원축소 모델링
#(2)변환
#iris 입력데이터를 차원축소 모델을 이용하여 2차원 데이터로 변환
X_2D = model.transform(irisX)
#(3)iris 데이터프레임에 변환값 추가
irisdf = pd.DataFrame(iris.data, columns=iris.feature_names)
irisdf['Species'] = iris.target
irisdf['PCA1'] = X_2D[:, 0]
irisdf['PCA2'] = X_2D[:, 1]
#(3)변환값 확인
sns.lmplot("PCA1", "PCA2", hue='Species', data=irisdf, fit_reg=False);
plt.title('iris dimensionality')
# -
# ### (4)Unsupervised learning: clustering
# algorithm : k-Means
#
# $\sum_{i=0}^{n}\min_{\mu_j \in C}(||x_i - \mu_j||^2)$
# +
# (1)랜덤 데이터 생성 및 확인
from sklearn.datasets.samples_generator import make_blobs
X, y_true = make_blobs(n_samples=200, centers=3,
cluster_std=0.70, random_state=0)
plt.scatter(X[:, 0], X[:, 1], s=50);
plt.title('samples')
plt.show()
# (2)KMeans 모듈로 모델링
from sklearn.cluster import KMeans
kmeans = KMeans(n_clusters=3)
kmeans.fit(X)
#(3)결과 확인
y_kmeans = kmeans.predict(X)
plt.scatter(X[:, 0], X[:, 1], c=y_kmeans, s=50, cmap='viridis')
plt.title('clustering (center)')
centers = kmeans.cluster_centers_
plt.scatter(centers[:, 0], centers[:, 1], c='black', s=200, alpha=0.5);
plt.show()
# -
| datamining7_scikitlearn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
from plotly import tools
from plotly.graph_objs import *
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
init_notebook_mode(connected=True)
import plotly.graph_objs as go
import matplotlib as mpl
import matplotlib.cm as cm
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import os, json, itertools
from glob import glob
from tqdm import tqdm
from sklearn.grid_search import ParameterGrid
from sklearn import *
from copy import deepcopy
from scipy.stats import boxcox
from scipy.stats.stats import pearsonr, spearmanr
from shapely.geometry import Polygon
from collections import Counter
import statsmodels.api as sm
from scipy.stats.mstats import zscore
from modules import correlation, evaluation, query, residual, visualization
sns.set(color_codes=True)
loo = model_selection.LeaveOneOut()
scaler = preprocessing.StandardScaler()
data_dir = os.path.join('..','..','data')
# Dictionary File
df_dictionary = pd.read_csv(os.path.join(data_dir,'shapefiles','csv','addrcode-dictionary.csv'))
df_dictionary['addrcode'] = df_dictionary['addrcode'].astype('int')
df_dictionary = df_dictionary.fillna('')
df_dictionary.head(1)
# -
df_rainfall = pd.read_csv(os.path.join(data_dir,'weather','csv','rainfall.csv'))
df_rainfall['addrcode'] = df_rainfall['addrcode'].astype('int')
df_rainfall['date'] = pd.to_datetime(df_rainfall['date'], format='%Y-%m')
df_rainfall = df_rainfall.set_index('date')
df_rainfall = df_rainfall['2015':'2017']
df_rainfall = df_rainfall.set_index('addrcode')
df_rainfall = df_rainfall.sort_index()
df_rainfall = df_rainfall.drop('rainfall_sum', axis=1)
df_rainfall.columns = ['rainfall']
df_rainfall = df_rainfall.groupby('addrcode').sum()
df_rainfall.head()
df_temperature = pd.read_csv(os.path.join(data_dir,'weather','csv','temperature.csv'))
df_temperature['addrcode'] = df_temperature['addrcode'].astype('int')
df_temperature['date'] = pd.to_datetime(df_temperature['date'], format='%Y-%m')
df_temperature = df_temperature.set_index('date')
df_temperature = df_temperature['2015':'2017']
df_temperature = df_temperature.set_index('addrcode')
df_temperature = df_temperature.sort_index()
df_temperature.columns = ['temperature']
df_temperature = df_temperature.groupby('addrcode').mean()
df_temperature.head()
# +
def get_rainfall(df_rainfall, addrcode):
if addrcode in df_rainfall.index:
return df_rainfall.loc[addrcode]['rainfall']
return df_rainfall['rainfall'].mean()
def get_temperature(df_temperature, addrcode):
if addrcode in df_temperature.index:
return df_temperature.loc[addrcode]['temperature']
return df_temperature['temperature'].mean()
def get_category(df_residual_corr, addrcode, col_name, q):
q1, q2 = q
value = df_residual_corr.loc[addrcode][col_name]
if value > q2: return 'Bad'
elif value > q1: return 'Average'
else: return 'Good'
def get_residual_category(df_residual_corr):
q_entry = {
'error':[df_residual_corr.describe().loc['25%']['error'], df_residual_corr.describe().loc['75%']['error']],
'norm_error':[df_residual_corr.describe().loc['25%']['norm_error'], df_residual_corr.describe().loc['75%']['norm_error']],
}
addrcodes = df_residual_corr.index.values
error_categories, norm_error_categories, rainfalls, temperatures = [], [], [], []
for addrcode in addrcodes:
error_categories.append(get_category(df_residual_corr, addrcode, 'error', q_entry['error']))
norm_error_categories.append(get_category(df_residual_corr, addrcode, 'norm_error', q_entry['norm_error']))
rainfalls.append(get_rainfall(df_rainfall, addrcode))
temperatures.append(get_temperature(df_temperature, addrcode))
df_residual_corr['rainfall'] = rainfalls
df_residual_corr['temperature'] = temperatures
df_residual_corr['error_category'] = error_categories
df_residual_corr['norm_error_category'] = norm_error_categories
df_residual_corr['gsv_density'] = df_residual_corr['image_area']/df_residual_corr['land_area']
df_residual_corr['rainfall_density'] = df_residual_corr['rainfall']/df_residual_corr['land_area']
df_residual_corr = df_residual_corr.drop([
'jar','bin','bucket','tire','pottedplant','bowl','cup','vase',
'n_jar','n_bin','n_bucket','n_tire','n_pottedplant','n_bowl','n_cup','n_vase',
'a_jar','a_bin','a_bucket','a_tire','a_pottedplant','a_bowl','a_cup','a_vase',
'ngsv_jar','ngsv_bin','ngsv_bucket','ngsv_tire','ngsv_pottedplant','ngsv_bowl','ngsv_cup','ngsv_vase',
'image_area', 'land_area','area',
'n_total','total','population',
'rainfall','error','predicted',
'cases', 'normalized_cases'
], axis=1)
return df_residual_corr, q_entry
def plot_distribution(df_residual_corr, col_name):
_=plt.figure(figsize=((10,3)))
_=sns.set_context("poster", font_scale = 1)
fig = sns.boxplot(df_residual_corr[col_name])
_ = sns.swarmplot(df_residual_corr[col_name], color=".25", size=8)
_=plt.title('Distribution of Prediction Errors, '+col_name)
_=fig.set_xlabel('Residual Error')
def plot_residual_heatmap(df_residual_corr):
df_residual_corr['norm_error'] = 100*df_residual_corr['norm_error']
# plot_distribution(df_residual_corr, 'error')
plot_distribution(df_residual_corr, 'norm_error')
cols = df_residual_corr.columns.tolist()
cols = cols[-2:] + cols[:-2]
df_residual_corr = df_residual_corr[cols].copy()
df_residual_corr, q_entry = get_residual_category(df_residual_corr)
plt.figure(figsize=(10,1.5))
sns.set(font_scale=1.3)
sns.heatmap(df_residual_corr.corr().loc[['norm_error']], annot=True, fmt=".2f", cmap="RdYlBu", vmin=-0.7, vmax=0.7)
plt.show()
return df_residual_corr, q_entry
# +
def residual_corr_plot(df_residual_corr, title, x_col, y_col, x_axis_name, y_axis_name, out_filename, is_hue=True):
flatui = ["#2ecc71","#34495e", "#e74c3c"]
if is_hue:
if x_col == 'error':
hue = 'error_category'
elif x_col == 'norm_error':
hue = 'norm_error_category'
else:
hue = None
_=plt.figure(figsize=((10,10)))
_=sns.set_context("poster", font_scale = 1)
fig = sns.FacetGrid(data=df_residual_corr, hue=hue, size=8, hue_order=["Good", "Average", "Bad"], palette=sns.color_palette(flatui), hue_kws=dict(marker=["o", "D","s"]))\
.map(plt.scatter, x_col, y_col) \
.add_legend()
fig = sns.regplot(x=x_col, y=y_col, data=df_residual_corr, scatter=False)
X = df_residual_corr[x_col].values
y = df_residual_corr[y_col].values
pearson_val = pearsonr(X, y)
spearman_val = spearmanr(X, y)
_=fig.set_title(
title+ \
'\nPearson: '+str(round(pearson_val[0],4))+ ', p-value: '+str(round(pearson_val[1],4))+ \
'\nSpearman: '+str(round(spearman_val[0],4))+', p-value: '+str(round(spearman_val[1],4))
)
_=fig.set_xlabel(x_axis_name)
_=fig.set_ylabel(y_axis_name)
_=fig.figure.savefig(out_filename+'.svg', bbox_inches='tight')
_=fig.figure.savefig(out_filename+'.png', bbox_inches='tight')
def residual_corr_split_plot(df_residual_corr, title, x_col, y_col, x_axis_name, y_axis_name, out_filename):
flatui = ["#2ecc71","#34495e", "#e74c3c"]
if x_col == 'error':
hue = 'error_category'
elif x_col == 'norm_error':
hue = 'norm_error_category'
_=plt.figure(figsize=((10,10)))
_=sns.set_context("poster", font_scale = 1)
fig = sns.lmplot(x=x_col, y=y_col, data=df_residual_corr, size=8,
hue=hue, hue_order=["Good", "Average", "Bad"],
palette=sns.color_palette(flatui), markers=["o", "D","s"])
X = df_residual_corr[x_col].values
y = df_residual_corr[y_col].values
pearson_val = pearsonr(X, y)
spearman_val = spearmanr(X, y)
_=plt.title(
title+ \
'\nPearson: '+str(round(pearson_val[0],4))+ ', p-value: '+str(round(pearson_val[1],4))+ \
'\nSpearman: '+str(round(spearman_val[0],4))+', p-value: '+str(round(spearman_val[1],4))
)
_=plt.xlabel(x_axis_name)
_=plt.ylabel(y_axis_name)
# _=fig.figure.savefig(out_filename+'.svg', bbox_inches='tight')
# _=fig.figure.savefig(out_filename+'.png', bbox_inches='tight')
# -
# # 1. Bangkok
df_features.columns
# +
### Load Shapefile
with open(os.path.join(data_dir,'shapefiles','geojson','Bangkok-subdistricts.geojson')) as f:
data_polygon = json.load(f)
# data_polygon['features'][0]['properties']
df_features = pd.read_csv(os.path.join('combined-dengue-case-features', 'Bangkok.csv'))
df_features.set_index('addrcode', inplace=True)
# df_features = df_features.loc[df_features.error > df_features.error.mean()].copy()
# df_features.head(1)
# df_features.error = boxcox(df_features.error.values)[0]
# df_features.norm_error = boxcox(df_features.norm_error.values)[0]
df_residual_corr, q_entry = plot_residual_heatmap(df_features.copy())
# +
### Load Shapefile
with open(os.path.join(data_dir,'shapefiles','geojson','Bangkok-subdistricts.geojson')) as f:
data_polygon = json.load(f)
# data_polygon['features'][0]['properties']
df_features = pd.read_csv(os.path.join('combined-dengue-case-features', 'Bangkok.csv'))
df_features.set_index('addrcode', inplace=True)
df_features = df_features.loc[df_features.norm_error < df_features.norm_error.quantile()*3].copy()
# df_features.head(1)
# df_features.error = boxcox(df_features.error.values)[0]
# df_features.norm_error = boxcox(df_features.norm_error.values)[0]
df_residual_corr, q_entry = plot_residual_heatmap(df_features.copy())
# +
# category='error_category'
# x_col='error'
# x_axis_name='Absolute Error'
# y_col='gsv_density'
# y_axis_name='GSV Density'
# residual_corr_plot(df_residual_corr,
# title='Residual Analysis',
# x_col=x_col, x_axis_name=x_axis_name,
# y_col=y_col, y_axis_name=y_axis_name,
# out_filename='Bangkok-gsv_density-residual-corr-1',
# # is_hue=False
# )
# category='error_category'
# x_col='error'
# x_axis_name='Absolute Error'
# y_col='gsv_density'
# y_axis_name='GSV Density'
# residual_corr_plot(df_residual_corr,
# title='Residual Analysis',
# x_col=x_col, x_axis_name=x_axis_name,
# y_col=y_col, y_axis_name=y_axis_name,
# out_filename='Bangkok-gsv_density-residual-corr-2',
# is_hue=False
# )
# -
# +
# category='norm_error_category'
# x_col='norm_error'
# x_axis_name='Absolute Error'
# y_col='pop_density'
# y_axis_name='pop_density'
# residual_corr_plot(df_residual_corr,
# title='Residual Analysis',
# x_col=x_col, x_axis_name=x_axis_name,
# y_col=y_col, y_axis_name=y_axis_name,
# out_filename=''
# )
# residual_corr_split_plot(df_residual_corr,
# title='Residual Analysis',
# x_col=x_col, x_axis_name=x_axis_name,
# y_col=y_col, y_axis_name=y_axis_name,
# out_filename=''
# )
# residual_corr_plot(df_residual_corr.loc[df_residual_corr[category] == 'Good'].copy(),
# title='Residual Analysis',
# x_col=x_col, x_axis_name=x_axis_name,
# y_col=y_col, y_axis_name=y_axis_name,
# out_filename=''
# )
# residual_corr_plot(df_residual_corr.loc[df_residual_corr[category] == 'Average'].copy(),
# title='Residual Analysis',
# x_col=x_col, x_axis_name=x_axis_name,
# y_col=y_col, y_axis_name=y_axis_name,
# out_filename=''
# )
# residual_corr_plot(df_residual_corr.loc[df_residual_corr[category] == 'Bad'].copy(),
# title='Residual Analysis',
# x_col=x_col, x_axis_name=x_axis_name,
# y_col=y_col, y_axis_name=y_axis_name,
# out_filename=''
# )
# -
df_residual.describe()
# +
# df_residual = df_features[['error']].copy()
# df_residual.columns = ['residual']
# visualization.choropleth_plot(
# data_polygon,
# df_residual,
# df_dictionary,
# map_style='streets', # dark, satellite, streets, light, outdoors
# cmap_name='RdYlGn', # Blues, viridis, RdYlGn, autumn, summer
# none_data_rgba='rgba(255,255,255,0.5)',
# opacity=0.8,
# is_reverse=True,
# save_file=True,
# filename='Bangkok-dengue-case-residual.html'
# )
df_residual = df_features[['norm_error']].copy()
df_residual.columns = ['residual']
visualization.choropleth_plot(
data_polygon,
df_residual,
df_dictionary,
map_style='streets', # dark, satellite, streets, light, outdoors
cmap_name='RdYlGn', # Blues, viridis, RdYlGn, autumn, summer
none_data_rgba='rgba(255,255,255,0.5)',
opacity=0.8,
is_reverse=True,
save_file=True,
filename='Bangkok-dengue-case-smape.html'
)
# -
# # 2. Nakhon
# +
### Load Shapefile
with open(os.path.join(data_dir,'shapefiles','geojson','Nakhon-subdistricts.geojson')) as f:
data_polygon = json.load(f)
# data_polygon['features'][0]['properties']
df_features = pd.read_csv(os.path.join('combined-dengue-case-features', 'Nakhon.csv'))
df_features.set_index('addrcode', inplace=True)
# df_features = df_features.loc[df_features.norm_error < 0.5].copy()
# df_features.head(1)
# df_features.error = boxcox(df_features.error.values)[0]
# df_features.norm_error = boxcox(df_features.norm_error.values)[0]
df_residual_corr, q_entry = plot_residual_heatmap(df_features.copy())
# +
# ### Load Shapefile
# with open(os.path.join(data_dir,'shapefiles','geojson','Nakhon-subdistricts.geojson')) as f:
# data_polygon = json.load(f)
# # data_polygon['features'][0]['properties']
# df_features = pd.read_csv(os.path.join('combined-dengue-case-features', 'Nakhon.csv'))
# df_features.set_index('addrcode', inplace=True)
# df_features = df_features.loc[df_features.norm_error < df_features.norm_error.quantile()*3].copy()
# # df_features = df_features.loc[df_features.norm_error < 0.5].copy()
# # df_features.head(1)
# # df_features.error = boxcox(df_features.error.values)[0]
# # df_features.norm_error = boxcox(df_features.norm_error.values)[0]
# df_residual_corr, q_entry = plot_residual_heatmap(df_features.copy())
# +
# category='error_category'
# x_col='error'
# x_axis_name='Absolute Error'
# y_col='temperature'
# y_axis_name='Temperature'
# residual_corr_plot(df_residual_corr,
# title='Residual Analysis',
# x_col=x_col, x_axis_name=x_axis_name,
# y_col=y_col, y_axis_name=y_axis_name,
# out_filename='Nakhon-temperature-residual-corr',
# # is_hue=False
# )
# +
# df_residual = df_residual_corr[['error']].copy()
# df_residual.columns = ['residual']
# visualization.choropleth_plot(
# data_polygon,
# df_residual,
# df_dictionary,
# map_style='streets', # dark, satellite, streets, light, outdoors
# cmap_name='RdYlGn', # Blues, viridis, RdYlGn, autumn, summer
# none_data_rgba='rgba(255,255,255,0.5)',
# opacity=0.8,
# is_reverse=True,
# save_file=True,
# filename='Nakhon-dengue-case-residual.html'
# )
df_residual = df_residual_corr[['norm_error']].copy()
df_residual.columns = ['residual']
visualization.choropleth_plot(
data_polygon,
df_residual,
df_dictionary,
map_style='streets', # dark, satellite, streets, light, outdoors
cmap_name='RdYlGn', # Blues, viridis, RdYlGn, autumn, summer
none_data_rgba='rgba(255,255,255,0.5)',
opacity=0.8,
is_reverse=True,
save_file=True,
filename='Nakhon-dengue-case-smape.html'
)
# -
# # 3. Krabi
# +
### Load Shapefile
with open(os.path.join(data_dir,'shapefiles','geojson','Krabi-subdistricts.geojson')) as f:
data_polygon = json.load(f)
# data_polygon['features'][0]['properties']
df_features = pd.read_csv(os.path.join('combined-dengue-case-features', 'Krabi.csv'))
df_features.set_index('addrcode', inplace=True)
# df_features = df_features.loc[df_features.norm_error < 0.5].copy()
# df_features.head(1)
df_residual_corr, q_entry = plot_residual_heatmap(df_features.copy())
# +
# ### Load Shapefile
# with open(os.path.join(data_dir,'shapefiles','geojson','Krabi-subdistricts.geojson')) as f:
# data_polygon = json.load(f)
# # data_polygon['features'][0]['properties']
# df_features = pd.read_csv(os.path.join('combined-dengue-case-features', 'Krabi.csv'))
# df_features.set_index('addrcode', inplace=True)
# df_features = df_features.loc[df_features.norm_error < df_features.norm_error.quantile()*3].copy()
# # df_features.head(1)
# df_residual_corr, q_entry = plot_residual_heatmap(df_features.copy())
# +
df_residual = df_residual_corr[['norm_error']].copy()
df_residual.columns = ['residual']
visualization.choropleth_plot(
data_polygon,
df_residual,
df_dictionary,
map_style='streets', # dark, satellite, streets, light, outdoors
cmap_name='RdYlGn', # Blues, viridis, RdYlGn, autumn, summer
none_data_rgba='rgba(255,255,255,0.5)',
opacity=0.8,
is_reverse=True,
save_file=True,
filename='Krabi-dengue-case-smape.html'
)
# +
# residual_corr_plot(df_residual_corr,
# title='Residual Analysis',
# x_col='norm_error', x_axis_name='Symmetric Absolute Percentage Error',
# y_col='temperature',y_axis_name='Temperature',
# out_filename='Krabi-temerature-smape'
# )
# +
# residual_corr_plot(df_residual_corr,
# title='Residual Analysis',
# x_col='error', x_axis_name='Absolute Error',
# y_col='rainfall_density', y_axis_name='Rainfall Density',
# out_filename='Krabi-rainfall-residual-corr'
# )
# residual_corr_plot(df_residual_corr,
# title='Residual Analysis',
# x_col='error', x_axis_name='Absolute Error',
# y_col='a_total',y_axis_name='Total Breeding Site Density',
# out_filename='Krabi-bs_density-residual-corr'
# )
# residual_corr_plot(df_residual_corr,
# title='Residual Analysis',
# x_col='error', x_axis_name='Absolute Error',
# y_col='pop_density',y_axis_name='Population Density',
# out_filename='Krabi-pop_density-residual-corr'
# )
# +
# category='error_category'
# x_col='error'
# x_axis_name='Absolute Error'
# y_col='pop_density'
# y_axis_name='pop_density'
# residual_corr_plot(df_residual_corr,
# title='Residual Analysis',
# x_col=x_col, x_axis_name=x_axis_name,
# y_col=y_col, y_axis_name=y_axis_name,
# out_filename=''
# )
# residual_corr_split_plot(df_residual_corr,
# title='Residual Analysis',
# x_col=x_col, x_axis_name=x_axis_name,
# y_col=y_col, y_axis_name=y_axis_name,
# out_filename=''
# )
# residual_corr_plot(df_residual_corr.loc[df_residual_corr[category] == 'Good'].copy(),
# title='Residual Analysis',
# x_col=x_col, x_axis_name=x_axis_name,
# y_col=y_col, y_axis_name=y_axis_name,
# out_filename=''
# )
# residual_corr_plot(df_residual_corr.loc[df_residual_corr[category] == 'Average'].copy(),
# title='Residual Analysis',
# x_col=x_col, x_axis_name=x_axis_name,
# y_col=y_col, y_axis_name=y_axis_name,
# out_filename=''
# )
# residual_corr_plot(df_residual_corr.loc[df_residual_corr[category] == 'Bad'].copy(),
# title='Residual Analysis',
# x_col=x_col, x_axis_name=x_axis_name,
# y_col=y_col, y_axis_name=y_axis_name,
# out_filename=''
# )
# -
| src/correlation/correlation-dengue-cases-summary-v2.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia (24 threads) 1.5.3
# language: julia
# name: julia-(24-threads)-1.5
# ---
using Revise, BeamPropagation, Distributions, StatsBase, StaticArrays, PhysicalConstants.CODATA2018, Plots, BenchmarkTools
const h = PlanckConstant.val
const ħ = h / 2π
const λ = @with_unit 626 "nm"
const k = 2π / λ
const m = @with_unit 57 "u"
const Δv = ħ * k / m
;
vz_μ = @with_unit 125 "m/s"
vz_σ = @with_unit 25 "m/s"
vxy_μ = @with_unit 0 "m/s"
vxy_σ = @with_unit 25 "m/s"
exit_radius = @with_unit 4 "mm"
;
const r = (
Normal(0, exit_radius/2),
Normal(0, exit_radius/2),
Normal(0, 0)
)
const v = (
Normal(vxy_μ, vxy_σ),
Normal(vxy_μ, vxy_σ),
Normal(vz_μ, vz_σ)
)
const a = (
Normal(0, 0),
Normal(0, 0),
Normal(0, 0)
)
;
# +
VBRs = Weights([
0.9457, # to 000
0.0447, # 100
3.9e-3, # 0200
2.7e-3, # 200
9.9e-4, # 0220
7.5e-4, # 0110
3.8e-4, # 0110, N=2
3.9e-4, # 1200
1.5e-4, # 300
1.3e-4, # 1220
0.7e-4, # 110 (only N=1, assuming roughly 2/3 to 1/3 rotational branching)
0.4e-4, # 110, N=2
5.7e-5, # 220
4.3e-5 # other states
])
longitudinal = [
true, # to 000
true, # 100
true, # 0200
true, # 200
true, # 0220
true, # 0110
true, # 0110, N=2
true, # 1200
true, # 300
false, # 1220
false, # 110
false, # 110, N=2
false, # 220
false # other states
]
transverse = [
false, # to 000
false, # 100
false, # 0200
false, # 200
false, # 0220
false, # 0110
false, # 0110, N=2
false, # 1200
false, # 300
true, # 1220
true, # 110
false, # 110, N=2
false, # 220
false # other states
]
λs = 1e-9 .* [
626,
574,
629,
650,
630,
567,
623,
653,
595,
646,
566,
600,
600,
600
]
;
# +
struct Sideband
lower::Float64
upper::Float64
end
std_sideband = Sideband(0, 250)
sidebands = repeat([std_sideband], length(VBRs))
;
# +
@inline function transverse_on(z)
if 0.15 < z < 0.20
return true
elseif 0.45 < z < 0.50
return true
elseif 0.56 < z < 0.61
return true
end
return false
end
@inline function random_unit3Dvector()
θ = rand(Uniform(0, 2π))
z = rand(Uniform(-1, 1))
return @SVector [sqrt(1-z^2)*cos(θ), sqrt(1-z^2)*sin(θ), z]
end
;
# -
const detect_rad = @with_unit 0.5 "cm"
const detect_zloc = @with_unit 71 "cm"
const detect_zlen = @with_unit 0.5 "cm"
const dead_rad = @with_unit 1.0 "cm"
const dead_len = detect_zloc + detect_zlen
;
# +
@inline function simple_prop(r, v)
dist_detect = detect_zloc - r[3]
x_final = r[1] + v[1] * dist_detect / v[3]
y_final = r[2] + v[2] * dist_detect / v[3]
return sqrt(x_final^2 + y_final^2)
end
@inline discard(r, v) = (simple_prop(r, v) > dead_rad) || (r[3] > dead_len)
@inline is_detectable(r) = sqrt(r[1]^2 + r[2]^2) < detect_rad && (detect_zloc + detect_zlen > r[3] > detect_zloc)
;
# +
Γ = 2π * 6.4e6
Ng = 12
Ne = 4
s_ = 0.405
Γeff = Γ * 2Ne / (Ng + Ne)
seff = (s_/2) * (Ng + Ne) / 2Ng
Δv_to_Δf(Δv, λ) = (Δv / λ)
Δf_to_Δv(Δf, λ) = (λ * Δf)
sideband_spacing = @with_unit 4.5 "MHz"
@inline function Rsc(λ, Γ, s, v, sideband_spacing, sideband)
R = 0
sideband_spacing_v = Δf_to_Δv(sideband_spacing, λ)
spectrum = sideband.lower:sideband_spacing_v:sideband.upper # This is in terms of velocity
s /= length(spectrum)
for v_s in spectrum
Δ = Δv_to_Δf(v_s - v, λ)
R += (s/2) * Γ / (1 + 4(Δ/Γ)^2 + s)
end
return R
end
vs = 0:1:300
Rscs = zeros(Float64, length(vs), length(VBRs))
for (i,v) in enumerate(vs)
for state in 1:size(VBRs,1)
Rscs[i, state] = Rsc(λs[state], Γ, s_, v, sideband_spacing, sidebands[state])
end
end
@inline function interpolate_Rsc(Rscs, vz, state)
if vz < 0
return 0.0
end
v_idx = round(Int64, vz) + 1
Rsc = Rscs[v_idx, state]
return Rsc
end
;
# +
@inline function save(i, r, v, a, state, s)
s.vzs[i] = v[3]
s.detectable[i] = simple_prop(r, v) < detect_rad
return nothing
end
@inline function f(i, r, v, a, state, dt, p, s)
state′ = state
v′ = v
s.detectable[i] |= is_detectable(r)
if p.longitudinal[state] || (p.transverse[state] && transverse_on(r[3]))
p_scatter = min(1, interpolate_Rsc(p.Rscs, v[3], state) / p.scattering_rate)
if rand() < p_scatter
state′ = sample(1:14, p.VBRs)
v′ = @SVector [v[1], v[2], v[3] - Δv]
v′ += Δv .* random_unit3Dvector()
s.photons[i] += 1
end
end
s.vzs[i] = v′[3]
s.states[i] = state′
return (state′, v′, a)
end
@inline function f(i, r, v, a, state, dt, p, s)
return (state, v, a)
end
;
# -
n = Int64(1e6)
scattering_rate = @with_unit 1.2 "MHz"
delete_every = 15
dt = repeat([1 / scattering_rate], n)
max_steps = Int64(3.5e4)
;
# +
vzs = zeros(Float64, n)
photons = zeros(Int64, n)
detectable = zeros(Bool, n)
states = ones(Int64, n)
v_cutoff = @with_unit 50 "m/s"
p = @params (VBRs, transverse, longitudinal, v_cutoff, sidebands, Rscs, scattering_rate)
s = @params (vzs, photons, detectable, states)
# sf = deepcopy(s0)
# @time s0, sf = propagate!(n, dt, r, v, a, f, save, discard, delete_every, max_steps, p, s0, sf)
# bright = [longitudinal[final_state] || transverse[final_state] for final_state in sf.states]
# ;
# -
@time rs, vs, as, states, dead, idxs = initialize_dists(n, r, v, a)
;
using StructArrays
particles = StructArray{Particle}(undef, n)
dt_ = dt[1]
@btime propagate_particles!($dt_, $r, $v, $a, $particles, $f, $save, $discard, $delete_every, $max_steps, $p, $s)
;
sum(37.5 .< sf.vzs[sf.detectable .& bright] .< 42.5) * 2e7 / n
barhist(s0.vzs[s0.detectable], bins=0:10:225, alpha=0.5, xlim=[0,225], label="Unperturbed")
barhist!(sf.vzs[sf.detectable], bins=0:10:225, alpha=0.5, label="Slowed")
barhist!(sf.vzs[sf.detectable .& bright], bins=0:10:225, alpha=0.3, label="Slowed and detected")
xlabel!("Velocity (m/s)")
ylabel!("Molecule count")
v_class = 0 .< sₜ.vzs .< 100
histogram2d(sₜ.vzs[sₜ.detectable .& bright.& v_class], s₀.vzs[sₜ.detectable .& bright .& v_class], bins=30)
ylabel!("Initial velocity (m/s)")
xlabel!("Slowed velocity (m/s)")
delete_idxs = 1:100000;
@btime StructArrays.foreachfield(v -> deleteat!(v, $delete_idxs), particles_) setup=(particles_=deepcopy(particles_soa)) evals=1
;
# +
iters = 1
v_cutoffs = 55:2.0:75
data = zeros(iters, size(v_cutoffs, 1))
for i in 1:iters
for (j, v_cutoff) in enumerate(v_cutoffs)
vzs = zeros(Float64, n)
photons = zeros(Int64, n)
detectable = zeros(Bool, n)
states = ones(Int64, n)
s = @params (vzs, photons, detectable, states)
p = @params (VBRs, transverse, addressed, v_cutoff)
@time s₀, sₜ = propagate!(n, r, v, a, f, save, discard, delete_every, dt, max_steps, p, s)
bright = [addressed[final_state] for final_state in sₜ.states]
perc_under_10ms = sum(abs.(sₜ.vzs[sₜ.detectable .& bright]) .< 10) / sum(s₀.detectable)
data[i,j] = perc_under_10ms
end
end
# -
@time StructArrays.foreachfield(v -> deleteat!(v, delete_idxs), particles_soa)
scatter(v_cutoffs, mean(data, dims=1)')
# This assumes that the z-axis is discretized into 7100 elements across 71 cm (so each element covers 0.1 mm)
@inline function interpolate_forceprofile(forceprofile, r, v)
z = r[3]
force_idx = round(Int64, z * 1e4) + 1
force = forceprofile[force_idx]
return force
end
forceprofile = collect(0:0.0001:0.71)
r_ = @SVector [0.0, 0.0, 0.4]
v_ = @SVector [0.0, 0.0, 0.0]
@btime interpolate_forceprofile($forceprofile, $r_, $v_)
| examples/slowing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from scipy import stats
exp_sites = np.random.exponential(size=1000)
exp_sites
stats.kstest(exp_sites, 'expon')
norm_sites = np.random.normal(size=1000)
exp_sites
stats.kstest(norm_sites, 'norm')
| notebooks/Testing Other Input Dist KS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # !pip3 install kfp --upgrade --user
# # Presidential Election pipeline
EXPERIMENT_NAME = 'presidential-election'
# ## Imports
import kfp
from kfp import dsl
# ## Load components
# +
def preprocess_op():
return dsl.ContainerOp(
name='Preprocess Data',
image='twarik/presidential_pipeline_preprocessing:latest',
arguments=[],
file_outputs={
'x_train': '/app/x_train.npy',
'x_test': '/app/x_test.npy',
'y_train': '/app/y_train.npy',
'y_test': '/app/y_test.npy',
}
)
def train_op(x_train, y_train):
return dsl.ContainerOp(
name='Train Model',
image='twarik/presidential_pipeline_train:latest',
arguments=[
'--x_train', x_train,
'--y_train', y_train
],
file_outputs={
'model': '/app/model.pkl'
}
)
def test_op(x_test, y_test, model):
return dsl.ContainerOp(
name='Test Model',
image='twarik/presidential_pipeline_test:latest',
arguments=[
'--x_test', x_test,
'--y_test', y_test,
'--model', model
],
file_outputs={
'mean_squared_error': '/app/output.txt'
}
)
def deploy_model_op(model):
return dsl.ContainerOp(
name='Deploy Model',
image='twarik/presidential_pipeline_deploy_model:latest',
arguments=[
'--model', model
]
)
# -
# ## Build the Pipeline
@dsl.pipeline(
name='Presidential Elections Pipeline',
description='Presidential Elections pipeline that performs preprocessing, training, evaluation and model deployment.'
)
def presidential_pipeline():
_preprocess_op = preprocess_op()
_train_op = train_op(
dsl.InputArgumentPath(_preprocess_op.outputs['x_train']),
dsl.InputArgumentPath(_preprocess_op.outputs['y_train'])
).after(_preprocess_op)
_test_op = test_op(
dsl.InputArgumentPath(_preprocess_op.outputs['x_test']),
dsl.InputArgumentPath(_preprocess_op.outputs['y_test']),
dsl.InputArgumentPath(_train_op.outputs['model'])
).after(_train_op)
deploy_model_op(
dsl.InputArgumentPath(_train_op.outputs['model'])
).after(_test_op)
# ## Compile the Pipeline
# +
pipeline_func = presidential_pipeline
pipeline_filename = pipeline_func.__name__ + '.pipeline.zip'
import kfp.compiler as compiler
compiler.Compiler().compile(pipeline_func, pipeline_filename)
# -
# ## Create a Kubeflow Experiment
# +
client = kfp.Client(host='pipelines-api.kubeflow.svc.cluster.local:8888')
try:
experiment = client.get_experiment(experiment_name=EXPERIMENT_NAME)
except:
experiment = client.create_experiment(EXPERIMENT_NAME)
print(experiment)
# -
# ## Run the Pipeline
# +
arguments = {}
run_name = pipeline_func.__name__ + ' run'
run_result = client.run_pipeline(experiment.id,
run_name,
pipeline_filename,
arguments)
print(experiment.id)
print(run_name)
print(pipeline_filename)
print(arguments)
| pipelines/pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Motivación para usar métodos Monte Carlo
#
# En un problema que estamos resolviendo con estadística bayesiana cuando tenemos la posterior $P(\lambda|O)$, donde $\lambda$ representa un parámetro que queremos estimar y $O$ representa las observaciones disponibles, una de las primeras cantidades que queremos calcular es el valor esperado de $\lambda$:
#
# $$
# \langle \lambda \rangle = \int \lambda P(\lambda|O) d\lambda.
# $$
#
# En casos sencillos esta integral se puede resolver analíticamente pero en casos más complejos se debe hacer de manera numérica.
# Si $\lambda$ representa un parámetro unidimensional el cálculo numérico de la integral se puede hacer a través del métodos que discretizan la variable de integración en $N$ puntos para aproximar la integral como la suma de área de polígonos (i.e. el método del trapecio).
#
# Pero esta metodología no va a funcionar numéricamente para altas dimensiones. Si tenemos $m$ diferentes dimensiones y discretizamos cada dimensión en $N$ puntos. Esto implicaría evaluar la función a integrar en $N^d$ puntos; con $N=100$ y $d=10$ el problema se vuelve rápidamente incontrolable. La solución a este problema es hacer integración por métodos Monte Carlo.
# ## Ejemplo
# Pensemos en el ejemplo concreto de la función $f(x)=\exp(x)$ que quisiéramos integrar entre $0\leq x\leq 1$ con métodos Monte Carlo.
#
# En este caso la integral se va a aproximar de la siguiente manera
#
# $$
# \int_0^1 \exp(x) dx \approx \sum_{i=1}^{N} f(x_i)\Delta x \approx \frac{x_{\rm max}-x_{\rm min}}{N} \sum_{i=1}^{N} f(x_i) \approx \frac{1}{N} \sum_{i=1}^{N} f(x_i)
# $$
#
# Lo interesante del método Monte Carlo de integración es que los $x_i$ no deben estar equiespaciados. Es suficiente con que vengan
# de una distribución de probabilidad uniforme entre $x_{\rm min}$ y $x_{\rm max}$.
#
# Comparemos los resultados de la integración por Monte Carlo con el resultado analítico $\int_0^1 \exp(x) dx=e^{1}-1$
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# +
def f(x):
return np.exp(x)
def integral_analitica():
return np.exp(1) - 1
def integral_monte_carlo(N=100):
x = np.random.random(N)
return np.sum(f(x))/N
n_intentos = 10
puntos = np.int_(np.logspace(1,5,n_intentos))
diferencias = np.ones(n_intentos)
for i in range(n_intentos):
a = integral_analitica()
b = integral_monte_carlo(N=puntos[i])
diferencias[i] = (np.abs((a-b)/a))
# -
plt.plot(puntos, diferencias*100)
plt.loglog()
plt.xlabel("$N_{puntos}$")
plt.ylabel("Diferencia porcentual Monte Carlo vs. Analitica")
# En este ejemplo vemos que si queremos tener una integral con $1\%$ de precisión lo podemos lograr con cerca de $30000$ puntos.
# ### Ejercicio 1.1
#
# Calcule la integral $\int_0^1 \sin(x)dx$ con el método Monte Carlo y produzca una gráfica como la anterior
# donde se muestra la diferencia porcentual entre la solución analítica y numérica como función del número de puntos.
# # Integrando con distribuciones no uniformes
#
# En el ejemplo anterior utilizamos puntos que tienen una distribución uniforme para integrar la función exponencial.
# En general es posible utilizar puntos que siguen una distribución $g(x)$ para integrar una función $f(x)$.
#
# Escrito en forma de ecuaciones la siguiente identidad es válida
#
# $$
# \frac{\int g(x)f(x)dx}{\int g(x)dx} = \langle f(x)\rangle \approx \frac{1}{N}\sum_{i=1}^{N}f(x_i)
# $$
#
# donde $g(x)$ representa una distribución de densidad de probabilidad que puede no estar normalizada. Si estuviera normalizada la integral
# del denominador sería igual a uno.
#
#
#
# # Ejemplo
#
# Supongamos que queremos calcular la integral $\int_0 ^{\infty} e^{-x} \sin(x) dx$ con el método Monte Carlo.
# Para esto sería suficiente generar puntos que sigan la distribución de densidad de probabilidad $e^{-x}$ entre 0 e infinito
# y luego calcular el valor promedio de $\sin(x)$ sobre esos puntos.
#
# +
def f(x):
return np.sin(x)
def integral_analitica():
return 0.5
def integral_monte_carlo(N=100):
x = np.random.exponential(size=N) # esto ya no es una distribucion uniforme!
return np.sum(f(x))/N
n_intentos = 30
puntos = np.int_(np.logspace(1,5,n_intentos))
diferencias = np.ones(n_intentos) # aqui guardaremos la diferencia entre la sol. numerica y la analitica
for i in range(n_intentos):
a = integral_analitica()
b = integral_monte_carlo(N=puntos[i])
diferencias[i] = (np.abs((a-b)/a))
# -
plt.plot(puntos, diferencias*100)
plt.loglog()
plt.xlabel("$N_{puntos}$")
plt.ylabel("Diferencia porcentual Monte Carlo vs. Analitica")
# ### Ejercicio 1.2
#
# Calcule la integral $\int_0^{\pi/2} x \sin(x)dx$ con un método Monte Carlo y prepare una gráfica similar a la del Ejemplo 1.2.
#
| Metodos Computacionales Avanzados/secciones/05.MCMC/01_Motivacion_Integracion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
# # PyTorch basics - Linear Regression from scratch
#
# <!-- <iframe width="560" height="315" src="https://www.youtube.com/embed/ECHX1s0Kk-o?controls=0" frameborder="0" allow="accelerometer; autoplay; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe> -->
#
# Tutorial inspired from [FastAI development notebooks](https://github.com/fastai/fastai_v1/tree/master/dev_nb)
#
# ## Machine Learning
#
# <img src="https://i.imgur.com/oJEQe7k.png" width="500">
#
#
# ## Tensors & Gradients
# + [markdown] _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# ### Import Numpy & PyTorch
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
import numpy as np
import torch
# + [markdown] _uuid="34f006aa7eb4bbc683c39b7059021da900180908"
# A tensor is a number, vector, matrix or any n-dimensional array.
# + [markdown] _uuid="e22be3f71825128f990e78959fa00d1331d344e4"
# ### Create tensors. (Add "requires_grad=True" for variables that needs calculation of gradients)
# + _uuid="e22be3f71825128f990e78959fa00d1331d344e4"
x = torch.tensor(3.0)
w = torch.tensor(4.0, requires_grad=True)
b = torch.tensor(5.0, requires_grad=True)
# + [markdown] _uuid="3cb90767ff9bc2c12b72548b1a430984241d4910"
# ### Print tensors
# + _uuid="3cb90767ff9bc2c12b72548b1a430984241d4910"
print(x)
print(w)
print(b)
# + [markdown] _uuid="66a939ee0ec472705acd3f23654bc3ccea1cc8b4"
# We can combine tensors with the usual arithmetic operations.
# + [markdown] _uuid="0bd8fdeb252742e3449b7a2f08bcb188645dc9cf"
# ### Arithmetic operations
# + _uuid="0bd8fdeb252742e3449b7a2f08bcb188645dc9cf"
y = w * x + b
print(y)
# + [markdown] _uuid="64e0f175c65c3e875c671c40e4a9bf495e30b772"
# What makes PyTorch special, is that we can automatically compute the derivative of `y` w.r.t. the tensors that have `requires_grad` set to `True` i.e. `w` and `b`.
# + [markdown] _uuid="6c98996f00294f99eb11989b5a9ecdbda31864e1"
# ### Compute gradients
# + _uuid="6c98996f00294f99eb11989b5a9ecdbda31864e1"
y.backward()
# + [markdown] _uuid="47a62ffb26a76329e511f9f063c4c26cc6a7dc21"
# ### Display gradients
# + _uuid="47a62ffb26a76329e511f9f063c4c26cc6a7dc21"
print("dy/dw:", w.grad)
print("dy/db:", b.grad)
# + [markdown] _uuid="0b65b6bb4d15127b1d51f09abf616cfd29fa48b4"
# ## Problem Statement
# + [markdown] _uuid="c1beecda01bc332596edd193cade30006e3f6cbf"
# We'll create a model that predicts crop yeilds for apples and oranges (*target variables*) by looking at the average temperature, rainfall and humidity (*input variables or features*) in a region. Here's the training data:
#
# <img src="https://i.imgur.com/lBguUV9.png" width="500" />
#
# In a **linear regression** model, each target variable is estimated to be a weighted sum of the input variables, offset by some constant, known as a bias :
#
# ```
# yeild_apple = w11 * temp + w12 * rainfall + w13 * humidity + b1
# yeild_orange = w21 * temp + w22 * rainfall + w23 * humidity + b2
# ```
#
# Visually, it means that the yield of apples is a linear or planar function of the temperature, rainfall & humidity.
#
# <img src="https://i.imgur.com/mtkR2lB.png" width="540" >
#
#
# **Our objective**: Find a suitable set of *weights* and *biases* using the training data, to make accurate predictions.
# + [markdown] _uuid="c24b8195c0e9c6e8e13e169d264484f1f9b3b1ae"
# ## Training Data
# The training data can be represented using 2 matrices (inputs and targets), each with one row per observation and one column per variable.
# + [markdown] _uuid="dfda99005fc6daf3a49ae1cdd427ccac0aa446b1"
# ### Input (temp, rainfall, humidity)
# + _uuid="dfda99005fc6daf3a49ae1cdd427ccac0aa446b1"
inputs = np.array(
[[73, 67, 43], [91, 88, 64], [87, 134, 58], [102, 43, 37], [69, 96, 70]],
dtype="float32",
)
# + [markdown] _uuid="bf56faf74f7e29c9ed7523308718a9ab1acc0667"
# ### Targets (apples, oranges)
# + _uuid="bf56faf74f7e29c9ed7523308718a9ab1acc0667"
targets = np.array(
[[56, 70], [81, 101], [119, 133], [22, 37], [103, 119]], dtype="float32"
)
# + [markdown] _uuid="70d48f83ae4fce7aba7dd78fd58dddc77c598bfd"
# Before we build a model, we need to convert inputs and targets to PyTorch tensors.
# + [markdown] _uuid="931c1bad8788e607fa100d4338e1b1fe120e2339"
# ### Convert inputs and targets to tensors
# + _uuid="931c1bad8788e607fa100d4338e1b1fe120e2339"
inputs = torch.from_numpy(inputs)
targets = torch.from_numpy(targets)
print(inputs)
print(targets)
# + [markdown] _uuid="652647cd90bd0784ec4dc53472410f7358ee18c9"
# ## Linear Regression Model (from scratch)
#
# The *weights* and *biases* can also be represented as matrices, initialized with random values. The first row of `w` and the first element of `b` are use to predict the first target variable i.e. yield for apples, and similarly the second for oranges.
# + [markdown] _uuid="6f788ae559355b3f01667be1554a5d2bdcade8db"
# ### Weights and biases
# + _uuid="6f788ae559355b3f01667be1554a5d2bdcade8db"
w = torch.randn(2, 3, requires_grad=True)
b = torch.randn(2, requires_grad=True)
print(w)
print(b)
# + [markdown] _uuid="3579a065997cae41f7f504916b6bc07878ac768c"
# The *model* is simply a function that performs a matrix multiplication of the input `x` and the weights `w` (transposed) and adds the bias `b` (replicated for each observation).
#
# $$
# \hspace{2.5cm} X \hspace{1.1cm} \times \hspace{1.2cm} W^T \hspace{1.2cm} + \hspace{1cm} b \hspace{2cm}
# $$
#
# $$
# \left[ \begin{array}{cc}
# 73 & 67 & 43 \\
# 91 & 88 & 64 \\
# \vdots & \vdots & \vdots \\
# 69 & 96 & 70
# \end{array} \right]
# %
# \times
# %
# \left[ \begin{array}{cc}
# w_{11} & w_{21} \\
# w_{12} & w_{22} \\
# w_{13} & w_{23}
# \end{array} \right]
# %
# # +
# %
# \left[ \begin{array}{cc}
# b_{1} & b_{2} \\
# b_{1} & b_{2} \\
# \vdots & \vdots \\
# b_{1} & b_{2} \\
# \end{array} \right]
# $$
# + [markdown] _uuid="b1119f5ae9688a5f31dba438c7f78ca382deb7e3"
# ### Define the model
# + _uuid="b1119f5ae9688a5f31dba438c7f78ca382deb7e3"
def model(x):
return x @ w.T + b
# + [markdown] _uuid="8e0a4644cb1c4ed68a3bcf67a8a156341ac7c853"
# The matrix obtained by passing the input data to the model is a set of predictions for the target variables.
# + [markdown] _uuid="b042a3cf8f16f4c4380cccbac9d0892719c24190"
# ### Generate predictions
# + _uuid="b042a3cf8f16f4c4380cccbac9d0892719c24190"
preds = model(inputs)
print(preds)
# + [markdown] _uuid="5551ef933de7902c8b5a38ae3d8e4795cb244f38"
# ### Compare with targets
# + _uuid="5551ef933de7902c8b5a38ae3d8e4795cb244f38"
print(targets)
# + [markdown] _uuid="2c4a9cf2b3c9152f2f832176bce9a87381e2419c"
# Because we've started with random weights and biases, the model does not a very good job of predicting the target varaibles.
# + [markdown] _uuid="edaae7266f5d47c5e970e1438a812f10d8d35fb4"
# ## Loss Function
#
# We can compare the predictions with the actual targets, using the following method:
# * Calculate the difference between the two matrices (`preds` and `targets`).
# * Square all elements of the difference matrix to remove negative values.
# * Calculate the average of the elements in the resulting matrix.
#
# The result is a single number, known as the **mean squared error** (MSE).
# + [markdown] _uuid="dbf5bca8cbf2a3831089b454c70469e3748e9682"
# ### MSE loss
# -
# $$\operatorname{MSE}=\frac{1}{n}\sum_{i=1}^n(Y_i-\hat{Y_i})^2.$$
# + _uuid="dbf5bca8cbf2a3831089b454c70469e3748e9682"
def mse(t1, t2):
diff = t1 - t2
return torch.sum(diff * diff) / diff.numel()
# + [markdown] _uuid="90da6779aad81608c40cdca77c3c04b68a815c11"
# ### Compute loss
# + _uuid="90da6779aad81608c40cdca77c3c04b68a815c11"
loss = mse(preds, targets)
print(loss)
# + [markdown] _uuid="3ab3acadf389f30430b55c26c7979dcffaa974a5"
# The resulting number is called the **loss**, because it indicates how bad the model is at predicting the target variables. Lower the loss, better the model.
# + [markdown] _uuid="c61acf9c3cff205d769fc52ed3b1b76f5ae66233"
# ## Compute Gradients
#
# With PyTorch, we can automatically compute the gradient or derivative of the `loss` w.r.t. to the weights and biases, because they have `requires_grad` set to `True`.
# + [markdown] _uuid="ef66710c6ef1944567c4dc033e1ca316f35490ab"
# ### Compute gradients
# + _uuid="ef66710c6ef1944567c4dc033e1ca316f35490ab"
loss.backward()
# + [markdown] _uuid="6504cddcfb4bfb0817bf03ef460f08f3145a9091"
# The gradients are stored in the `.grad` property of the respective tensors.
# + [markdown] _uuid="5943d1cef604a178c95f5e8d255519d42d9f9982"
# ### Gradients for weights
# + _uuid="5943d1cef604a178c95f5e8d255519d42d9f9982"
print(w)
print(w.grad)
# + [markdown] _uuid="47278e318b156c6a5812e0842dbc4164c8362562"
# ### Gradients for bias
# + _uuid="47278e318b156c6a5812e0842dbc4164c8362562"
print(b)
print(b.grad)
# + [markdown] _uuid="466dc3a2cc2d4bd2c10ae4cf59cf4627b5cc9c75"
# A key insight from calculus is that the gradient indicates the rate of change of the loss, or the slope of the loss function w.r.t. the weights and biases.
#
# * If a gradient element is **postive**,
# * **increasing** the element's value slightly will **increase** the loss.
# * **decreasing** the element's value slightly will **decrease** the loss.
#
# <img src="https://i.imgur.com/2H4INoV.png" width="400" />
#
#
#
# * If a gradient element is **negative**,
# * **increasing** the element's value slightly will **decrease** the loss.
# * **decreasing** the element's value slightly will **increase** the loss.
#
# <img src="https://i.imgur.com/h7E2uAv.png" width="400" />
#
# The increase or decrease is proportional to the value of the gradient.
# + [markdown] _uuid="35ed968bfc135bd44eeb100ae401d0628fbc5c63"
# Finally, we'll reset the gradients to zero before moving forward, because PyTorch accumulates gradients.
# -
# ### Reset Gradients to Zero with "tensor.grad.zero_()" method
# + _uuid="5f02dc376c21857d4e545d98413952c5ac73039b"
w.grad.zero_()
b.grad.zero_()
print(w.grad)
print(b.grad)
# + [markdown] _uuid="5501c66c9729c4954e9b798a0634a9d84487e639"
# ## Adjust weights and biases using gradient descent
#
# We'll reduce the loss and improve our model using the gradient descent algorithm, which has the following steps:
#
# 1. Generate predictions
# 2. Calculate the loss
# 3. Compute gradients w.r.t the weights and biases
# 4. Adjust the weights by subtracting a small quantity proportional to the gradient
# 5. Reset the gradients to zero
# + [markdown] _uuid="ef0d2bd2d9c5acb60992e238439ee00c2223319f"
# ### Generate predictions
# + _uuid="ef0d2bd2d9c5acb60992e238439ee00c2223319f"
preds = model(inputs)
print(preds)
# + [markdown] _uuid="302ee8226da4ee5d0dad137c638573a79f8abded"
# ### Calculate the loss
# + _uuid="302ee8226da4ee5d0dad137c638573a79f8abded"
loss = mse(preds, targets)
print(loss)
# + [markdown] _uuid="01c596aecf87e4670033ddd4ed36e26b97e2f9ab"
# ### Compute gradients
# + _uuid="01c596aecf87e4670033ddd4ed36e26b97e2f9ab"
loss.backward()
# + [markdown] _uuid="ec1e2bdc8f91523e556fad55ee8c01eb5431ae24"
# ### Adjust weights & reset gradients
# -
# $learning\ rate=1e-5$
# + _uuid="ec1e2bdc8f91523e556fad55ee8c01eb5431ae24"
with torch.no_grad():
w -= w.grad * 1e-5
b -= b.grad * 1e-5
w.grad.zero_()
b.grad.zero_()
# + _uuid="1d61b6f61f49b19099d29d1be8ec5ae4967bbd51"
print(w)
# + [markdown] _uuid="6af10c29db7cb0d6e869b2c30966a34a48a011e2"
# With the new weights and biases, the model should have a lower loss.
# + [markdown] _uuid="c542b5fe75d82454f34cac13cdcff8b48dd1945c"
# ### Calculate loss
# + _uuid="c542b5fe75d82454f34cac13cdcff8b48dd1945c"
preds = model(inputs)
loss = mse(preds, targets)
print(loss)
# + [markdown] _uuid="5201901695f3ea13d7fdd5d985da7e0761c541d0"
# ## Train for multiple epochs
#
# To reduce the loss further, we repeat the process of adjusting the weights and biases using the gradients multiple times. Each iteration is called an epoch.
# + [markdown] _uuid="9f5f0ffeee666b30c5828636359f0be6addbef7c"
# ### Train for 100 epochs
# + _uuid="9f5f0ffeee666b30c5828636359f0be6addbef7c"
for i in range(100):
preds = model(inputs)
loss = mse(preds, targets)
loss.backward()
with torch.no_grad():
w -= w.grad * 1e-5
b -= b.grad * 1e-5
w.grad.zero_()
b.grad.zero_()
# + [markdown] _uuid="c4820ca48b78f4dc242d80a9ec3ec6aca1aef671"
# ### Calculate loss
# + _uuid="c4820ca48b78f4dc242d80a9ec3ec6aca1aef671"
preds = model(inputs)
loss = mse(preds, targets)
print(loss)
# + [markdown] _uuid="bbcd65fa7094cec187565e54c2107e683bea787b"
# ### Print predictions
# + _uuid="bbcd65fa7094cec187565e54c2107e683bea787b"
preds
# + [markdown] _uuid="addec2c4eca8edfcae5544ea2cc717182c21d90f"
# ### Print targets
# + _uuid="addec2c4eca8edfcae5544ea2cc717182c21d90f"
targets
# + [markdown] _uuid="ecc6e79cdfb6a8ca882895ccc895b61b960b0a04"
# ## Linear Regression Model using PyTorch built-ins
#
# Let's re-implement the same model using some built-in functions and classes from PyTorch.
# + [markdown] _uuid="ce66cf0d09a3f38bf2f00ea40418c56d98f1f814"
# ### Imports
# + _uuid="ce66cf0d09a3f38bf2f00ea40418c56d98f1f814"
import torch.nn as nn
# + [markdown] _uuid="74bb18bd01ac809079eeb8d05695206e8ba02069"
# ### Input (temp, rainfall, humidity)
# + _uuid="74bb18bd01ac809079eeb8d05695206e8ba02069"
inputs = np.array(
[
[73, 67, 43],
[91, 88, 64],
[87, 134, 58],
[102, 43, 37],
[69, 96, 70],
[73, 67, 43],
[91, 88, 64],
[87, 134, 58],
[102, 43, 37],
[69, 96, 70],
[73, 67, 43],
[91, 88, 64],
[87, 134, 58],
[102, 43, 37],
[69, 96, 70],
],
dtype="float32",
)
# + [markdown] _uuid="74bb18bd01ac809079eeb8d05695206e8ba02069"
# ### Targets (apples, oranges)
# + _uuid="74bb18bd01ac809079eeb8d05695206e8ba02069"
targets = np.array(
[
[56, 70],
[81, 101],
[119, 133],
[22, 37],
[103, 119],
[56, 70],
[81, 101],
[119, 133],
[22, 37],
[103, 119],
[56, 70],
[81, 101],
[119, 133],
[22, 37],
[103, 119],
],
dtype="float32",
)
# + _uuid="d94b355f55250e9c7dcff668920f02d7c5c04925"
inputs = torch.from_numpy(inputs)
targets = torch.from_numpy(targets)
# + [markdown] _uuid="a0665466eb5401f40a816b323a34450b2c052c41"
# ### Dataset and DataLoader
#
# We'll create a `TensorDataset`, which allows access to rows from `inputs` and `targets` as tuples. We'll also create a DataLoader, to split the data into batches while training. It also provides other utilities like shuffling and sampling.
# + [markdown] _uuid="206f5fd0473386476b23477bf38d2c327b6376c9"
# ### Import tensor dataset & data loader
# + _uuid="206f5fd0473386476b23477bf38d2c327b6376c9"
from torch.utils.data import TensorDataset, DataLoader
# + [markdown] _uuid="c47a4f2f86fda3918094e01cf7ab0698bbb5acc7"
# ### Define dataset
# + _uuid="c47a4f2f86fda3918094e01cf7ab0698bbb5acc7"
train_ds = TensorDataset(inputs, targets)
train_ds[0:3]
# + [markdown] _uuid="0a2f69126319d738b82ae67d5d404ecd6161bfac"
# ### Define data loader
# + _uuid="0a2f69126319d738b82ae67d5d404ecd6161bfac"
batch_size = 5
train_dl = DataLoader(train_ds, batch_size, shuffle=True)
next(iter(train_dl))
# + [markdown] _uuid="276a262e1b9e3a048bcd32989013f9c501c59037"
# ### nn.Linear
# Instead of initializing the weights & biases manually, we can define the model using `nn.Linear`.
# + [markdown] _uuid="59da3506559a0640d80d18f77b02726a1757be2f"
# ### Define model
# + _uuid="59da3506559a0640d80d18f77b02726a1757be2f"
model = nn.Linear(3, 2, bias=True)
print(model.weight)
print(model.bias)
# + [markdown] _uuid="b3a4a8c499a4680f2533329712de034671dd1cdd"
# ### Optimizer
# Instead of manually manipulating the weights & biases using gradients, we can use the optimizer `optim.SGD`.
# + [markdown] _uuid="1848398bd1ced8c25a7bb55612cf32a774500280"
# ### Define optimizer (SGD-Stochastic Gradient Descent)
# + _uuid="1848398bd1ced8c25a7bb55612cf32a774500280"
opt = torch.optim.SGD(model.parameters(), lr=1e-5)
# + [markdown] _uuid="28cbe62be55010bd11b31d819cff38da5a772b18"
# ### Loss Function
# Instead of defining a loss function manually, we can use the built-in loss function `mse_loss`.
# + [markdown] _uuid="69d7f4e8e27ccd077f711da27f8bede8aa711893"
# ### Import nn.functional
# + _uuid="69d7f4e8e27ccd077f711da27f8bede8aa711893"
import torch.nn.functional as F
# + [markdown] _uuid="a02ff888ed4be720fd9ca376022d8fdcf2559683"
# ### Define loss function
# + _uuid="a02ff888ed4be720fd9ca376022d8fdcf2559683"
loss_fn = F.mse_loss
# + _uuid="a540adf76725ea9968025f6c029fdd251bdada6c"
loss = loss_fn(model(inputs), targets)
print(loss)
# + [markdown] _uuid="e833614a69ff18c554a3d89f643ae2f11e0260f6"
# ### Train the model
#
# We are ready to train the model now. We can define a utility function `fit` which trains the model for a given number of epochs.
# + [markdown] _uuid="128bc7260221f5338edf8b503c75f0c7d1cce7e8"
# ### Define a utility function to train the model
# + _uuid="128bc7260221f5338edf8b503c75f0c7d1cce7e8"
def fit(num_epochs, model, loss_fn, opt):
for epoch in range(num_epochs):
for xb, yb in train_dl:
# take batch of items using dataloader
# Generate predictions
pred = model(xb)
loss = loss_fn(pred, yb)
# Perform gradient descent
loss.backward()
opt.step()
opt.zero_grad()
print("Training loss: ", loss_fn(model(inputs), targets))
# + [markdown] _uuid="ae8ca4686cf6a68f6c9ca93bf3d227abe96c2201"
# ### Train the model for 100 epochs
# + _uuid="ae8ca4686cf6a68f6c9ca93bf3d227abe96c2201"
fit(100, model, loss_fn, opt)
# + [markdown] _uuid="32588a47d0478772a1f08fa55874a322630bd0b6"
# ### Generate predictions
# + _uuid="32588a47d0478772a1f08fa55874a322630bd0b6"
preds = model(inputs)
preds
# + [markdown] _uuid="12d757c0f37c2e3af65cf9d4b59878cc10c65acf"
# ### Compare with targets
# + _uuid="12d757c0f37c2e3af65cf9d4b59878cc10c65acf"
targets
# + [markdown] _uuid="e182289ebf21d8296f11f13264c4732c100da14f"
# # Bonus: Feedfoward Neural Network
#
# 
#
# Conceptually, you think of feedforward neural networks as two or more linear regression models stacked on top of one another with a non-linear activation function applied between them.
#
# <img src="https://cdn-images-1.medium.com/max/1600/1*XxxiA0jJvPrHEJHD4z893g.png" width="640">
#
# To use a feedforward neural network instead of linear regression, we can extend the `nn.Module` class from PyTorch.
# + _uuid="c405e5075d6c4adb26ead75c17be90eaeb43f2d5"
class SimpleNet(nn.Module):
# Initialize the layers
def __init__(self):
super().__init__()
self.linear1 = nn.Linear(3, 3)
self.act1 = nn.ReLU() # Activation function
self.linear2 = nn.Linear(3, 2)
# Perform the computation
def forward(self, x):
x = self.linear1(x)
x = self.act1(x)
x = self.linear2(x)
return x
# + [markdown] _uuid="2448d9832722f4f2813f8bd80b91daefd901dc2e"
# Now we can define the model, optimizer and loss function exactly as before.
# + _uuid="a51ca222c2ea037c3caccaeab98ccdbcc30800cf"
model = SimpleNet()
opt = torch.optim.SGD(model.parameters(), 1e-5)
loss_fn = F.mse_loss
# + [markdown] _uuid="21000c9739ea39a173a256f87339bfc112c1a9b0"
# Finally, we can apply gradient descent to train the model using the same `fit` function defined earlier for linear regression.
#
# <img src="https://i.imgur.com/g7Rl0r8.png" width="500">
# + _uuid="e94de6868c76803a998c1c1934ed229c826f3b8c"
fit(100, model, loss_fn, opt)
| content/week-05/samples/calculus-02/Gradient Descent (PyTorch).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %run common.ipynb
t = ete3.Tree(WG_TREE)
print(t)
g = {
# subgenera - use basal divergence only
'Anopheles_subgenus':t.get_common_ancestor('Anopheles_sinensis','Anopheles_atroparvus'),
'Nyssorhynchus_subgenus':t.get_common_ancestor('Anopheles_albimanus','Anopheles_darlingi'),
'Celia_subgenus':t.get_common_ancestor('Anopheles_dirus','Anopheles_funestus'),
# series - use basal divergence only
'Myzomiya_series':t.get_common_ancestor('Anopheles_funestus','Anopheles_culicifacies'),
'Neocellia_series':t.get_common_ancestor('Anopheles_stephensi','Anopheles_maculatus'),
'Pyretophorus_series':t.get_common_ancestor('Anopheles_epiroticus','Anopheles_gambiae'),
'Neomyzomia_series':t.get_common_ancestor('Anopheles_farauti','Anopheles_dirus')
}
# add group names
for name, group in g.items():
for s in name.split('_'):
tf = ete3.TextFace(s, fsize=12)
tf.hz_align=1
tf.opacity=0.7
group.add_face(tf, column=0, position='branch-right')
# node appearance
ns = ete3.NodeStyle(size=0)
for n in t.traverse():
n.set_style(ns)
t.convert_to_ultrametric()
# tree appearance
ts = ete3.TreeStyle()
ts.orientation = 1
# ts.show_branch_support = True
# ts.show_leaf_name = True
# ts.show_branch_length = False
ts.show_scale = False
t.render(WG_TREE_FIG, tree_style=ts);
| work/7_species_id/7_wg_tree_vis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Django Shell-Plus
# language: python
# name: django_extensions
# ---
# # Decoding Filing Periods
# The raw data tables mix together filings from different reporting periods (e.g. quarterlys vs. semi-annual vs. pre-elections). But we need these filings to be sorted (or at least sortable) so that or users, for example, can compare the performance of two candidates in the same reporting period.
# There are two vectors at play here:
# 1. The "Statement Type", as described in CAL-ACCESS parlance, which indicates the length of time covered by the filing and how close it was filed to the election.
# 2. The actual time interval the filing covers, denoted by a start date and an end date.
# This notebook is pulling data from the downloads-website's dev database, which was last updated on...
from calaccess_processed.models.tracking import ProcessedDataVersion
ProcessedDataVersion.objects.latest()
# Will also need to execute some raw SQL, so I'll import a helper function in order to make the results more readable:
from project import sql_to_agate
# Let's start by examining the distinct values of the statement type on `CVR_CAMPAIGN_DISCLOSURE_CD`. And let's narrow the scope to only the Form 460 filings.
sql_to_agate(
"""
SELECT UPPER("STMT_TYPE"), COUNT(*)
FROM "CVR_CAMPAIGN_DISCLOSURE_CD"
WHERE "FORM_TYPE" = 'F460'
GROUP BY 1
ORDER BY COUNT(*) DESC;
"""
).print_table()
# Not all of these values are defined, as previously noted in our [docs](http://calaccess.californiacivicdata.org/documentation/calaccess-files/cvr-campaign-disclosure-cd/#fields):
# * `PR` might be pre-election
# * `QS` is pro probably quarterly statement
# * `YE` might be...I don't know "Year-end"?
# * `S` is probably semi-annual
#
# Maybe come back later and look at the actual filings. There aren't that many.
# There's another similar-named column on `FILER_FILINGS_CD`, but this seems to be a completely different thing:
sql_to_agate(
"""
SELECT FF."STMNT_TYPE", LU."CODE_DESC", COUNT(*)
FROM "FILER_FILINGS_CD" FF
JOIN "LOOKUP_CODES_CD" LU
ON FF."STMNT_TYPE" = LU."CODE_ID"
AND LU."CODE_TYPE" = 10000
GROUP BY 1, 2;
"""
).print_table()
# One of the tables that caught my eye is `FILING_PERIOD_CD`, which appears to have a row for each quarterly filing period:
sql_to_agate(
"""
SELECT *
FROM "FILING_PERIOD_CD"
"""
).print_table()
# Every period is described as a quarter, and the records are equally divided among them:
sql_to_agate(
"""
SELECT "PERIOD_DESC", COUNT(*)
FROM "FILING_PERIOD_CD"
GROUP BY 1;
"""
).print_table()
# The difference between every `START_DATE` and `END_DATE` is actually a three-month interval:
sql_to_agate(
"""
SELECT "END_DATE" - "START_DATE" AS duration, COUNT(*)
FROM "FILING_PERIOD_CD"
GROUP BY 1;
"""
).print_table()
# And they have covered every year between 1973 and 2334 (how optimistic!):
sql_to_agate(
"""
SELECT DATE_PART('year', "START_DATE")::int as year, COUNT(*)
FROM "FILING_PERIOD_CD"
GROUP BY 1
ORDER BY 1 DESC;
"""
).print_table()
# Filings are linked to filing periods via `FILER_FILINGS_CD.PERIOD_ID`. While that column is not always populated, it is if you limit your results to just the Form 460 filings:
sql_to_agate(
"""
SELECT ff."PERIOD_ID", fp."START_DATE", fp."END_DATE", fp."PERIOD_DESC", COUNT(*)
FROM "FILER_FILINGS_CD" ff
JOIN "CVR_CAMPAIGN_DISCLOSURE_CD" cvr
ON ff."FILING_ID" = cvr."FILING_ID"
AND ff."FILING_SEQUENCE" = cvr."AMEND_ID"
AND cvr."FORM_TYPE" = 'F460'
JOIN "FILING_PERIOD_CD" fp
ON ff."PERIOD_ID" = fp."PERIOD_ID"
GROUP BY 1, 2, 3, 4
ORDER BY fp."START_DATE" DESC;
"""
).print_table()
# Also, is Schwarzenegger running this cycle? Who else could be filing from so far into the future?
# AAANNNNYYYway...Also need to check to make sure the join between `FILER_FILINGS_CD` and `CVR_CAMPAIGN_DISCLOSURE_CD` isn't filtering out too many filings:
sql_to_agate(
"""
SELECT cvr."FILING_ID", cvr."FORM_TYPE", cvr."FILER_NAML"
FROM "CVR_CAMPAIGN_DISCLOSURE_CD" cvr
LEFT JOIN "FILER_FILINGS_CD" ff
ON cvr."FILING_ID" = ff."FILING_ID"
AND cvr."AMEND_ID" = ff."FILING_SEQUENCE"
WHERE cvr."FORM_TYPE" = 'F460'
AND (ff."FILING_ID" IS NULL OR ff."FILING_SEQUENCE" IS NULL)
ORDER BY cvr."FILING_ID";
"""
).print_table(max_column_width=60)
# So only a handful, mostly local campaigns or just nonsense test data.
# So another important thing to check is how well these the dates from the filing period look-up records line up with the dates on the Form 460 filing records. It would be bad if the `CVR_CAMPAIGN_DISCLOSURE_CD.FROM_DATE` were before `FILING_PERIOD_CD.START_DATE` or if `CVR_CAMPAIGN_DISCLOSURE_CD.THRU_DATE` were after `FILING_PERIOD_CD.END_DATE`.
sql_to_agate(
"""
SELECT
CASE
WHEN cvr."FROM_DATE" < fp."START_DATE" THEN 'filing from_date before period start_date'
WHEN cvr."THRU_DATE" > fp."END_DATE" THEN 'filing thru_date after period end_date'
ELSE 'okay'
END as test,
COUNT(*)
FROM "CVR_CAMPAIGN_DISCLOSURE_CD" cvr
JOIN "FILER_FILINGS_CD" ff
ON cvr."FILING_ID" = ff."FILING_ID"
AND cvr."AMEND_ID" = ff."FILING_SEQUENCE"
JOIN "FILING_PERIOD_CD" fp
ON ff."PERIOD_ID" = fp."PERIOD_ID"
WHERE cvr."FORM_TYPE" = 'F460'
GROUP BY 1;
"""
).print_table(max_column_width=60)
# So half of the time, the `THRU_DATE` on the filing is later than the `FROM_DATE` on the filing period. How big of a difference can exist between these two dates?
sql_to_agate(
"""
SELECT
cvr."THRU_DATE" - fp."END_DATE" as date_diff,
COUNT(*)
FROM "CVR_CAMPAIGN_DISCLOSURE_CD" cvr
JOIN "FILER_FILINGS_CD" ff
ON cvr."FILING_ID" = ff."FILING_ID"
AND cvr."AMEND_ID" = ff."FILING_SEQUENCE"
JOIN "FILING_PERIOD_CD" fp
ON ff."PERIOD_ID" = fp."PERIOD_ID"
WHERE cvr."FORM_TYPE" = 'F460'
AND cvr."THRU_DATE" > fp."END_DATE"
GROUP BY 1
ORDER BY COUNT(*) DESC;
"""
).print_table(max_column_width=60)
# Ugh. Looks like, in most of the problem cases, the from date can be a whole quarter later than the end date of the filing period. Let's take a closer look at these...
sql_to_agate(
"""
SELECT
cvr."FILING_ID",
cvr."AMEND_ID",
cvr."FROM_DATE",
cvr."THRU_DATE",
fp."START_DATE",
fp."END_DATE"
FROM "CVR_CAMPAIGN_DISCLOSURE_CD" cvr
JOIN "FILER_FILINGS_CD" ff
ON cvr."FILING_ID" = ff."FILING_ID"
AND cvr."AMEND_ID" = ff."FILING_SEQUENCE"
JOIN "FILING_PERIOD_CD" fp
ON ff."PERIOD_ID" = fp."PERIOD_ID"
WHERE cvr."FORM_TYPE" = 'F460'
AND 90 < cvr."THRU_DATE" - fp."END_DATE"
AND cvr."THRU_DATE" - fp."END_DATE" < 93
ORDER BY cvr."THRU_DATE" DESC;
"""
).print_table(max_column_width=60)
# So, actually, this sort of makes sense: Quarterly filings are for three month intervals, while the semi-annual filings are for six month intervals. And `FILING_PERIOD_CD` only has records for three month intervals. Let's test this theory by getting the distinct `CVR_CAMPAIGN_DISCLOSURE_CD.STMT_TYPE` values from these records:
sql_to_agate(
"""
SELECT UPPER(cvr."STMT_TYPE"), COUNT(*)
FROM "CVR_CAMPAIGN_DISCLOSURE_CD" cvr
JOIN "FILER_FILINGS_CD" ff
ON cvr."FILING_ID" = ff."FILING_ID"
AND cvr."AMEND_ID" = ff."FILING_SEQUENCE"
JOIN "FILING_PERIOD_CD" fp
ON ff."PERIOD_ID" = fp."PERIOD_ID"
WHERE cvr."FORM_TYPE" = 'F460'
AND 90 < cvr."THRU_DATE" - fp."END_DATE"
AND cvr."THRU_DATE" - fp."END_DATE" < 93
GROUP BY 1
ORDER BY COUNT(*) DESC;
"""
).print_table(max_column_width=60)
# At least this is mostly true.
| calaccess-exploration/decoding-filing-periods.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Example 1
name = '<NAME>'
name_split = name.split()
(name_split[1]+' '+name_split[0]).title()
name = '<NAME>'
name_split = name.split()
name_result = (name_split[1]+' '+name_split[0]). title()
print("Processed name: "+ name_result)
# Good introduction to python strings can be found at https://developers.google.com/edu/python/strings
# ## Exmaple 2
names = ['<NAME>', '<NAME>', '<NAME>']
for name in names:
name_split = name.split()
name_result = (name_split[1]+' '+name_split[0]). title()
print("Processed name: "+ name_result)
# Similar axample but with larger number of names and with some names having the same given or last name.
names = ['<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>',
'<NAME>']
for name in names:
name_split = name.split()
name_result = (name_split[1]+' '+name_split[0]). title()
print("Processed name: "+ name_result)
# ## Example 3
import pandas
names = pandas.read_excel('./names.xlsx')
names
names['Last name'] = 'DR '+ names['Last name'].str.upper()
names['First name'] = names['First name'].str.capitalize()
names
names = names[['First name', 'Last name']]
names
# Final code put together
# +
import pandas
names = pandas.read_excel('./names.xlsx')
names['Last name'] = 'DR '+ names['Last name'].str.upper()
names['First name'] = names['First name'].str.capitalize()
names = names[['First name', 'Last name']]
writer = pandas.ExcelWriter('names_processed.xlsx')
names.to_excel(writer, 'Sheet1')
writer.close()
# -
# ## Example 4
f = open('./names.txt')
firstnames = []
lastnames = []
for line in f.readlines():
if line.startswith('LN'):
lastnames.append(line.split()[1])
elif line.startswith('FN'):
firstnames.append(line.split()[1])
else:
pass
f.close()
firstnames
lastnames
# ## Example 5
import numpy
numpy.set_printoptions(precision=3 , suppress= True) # this is just to make the output look better
numbers = numpy.loadtxt('./numpers.txt')
numbers
reshaped = numbers.reshape(3,3)
reshaped
reshaped[:,2].mean()
# Final code put together
import numpy
numbers = numpy.loadtxt('./numpers.txt')
reshaped = numbers.reshape(3,3)
reshaped[:,2].mean()
| python_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/nahuelalmeira/graphMachineLearning/blob/main/Notebook5_Tutorial_PyTorch_Geometric.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# -
# # Clasificación de nodos utilizando redes neuronales de grafos
#
# En esta notebook utilizaremos redes neuronales de grafos (GNN) para la tarea de clasificación de nodos.
#
# Este tutorial está adaptado del tutorial [Node Classification with Graph Neural Networks](https://colab.research.google.com/drive/14OvFnAXggxB8vM4e8vSURUp1TaKnovzX?usp=sharing), de PyTorch-Geometric.
# ## Configuración general
# +
import torch
def format_pytorch_version(version):
return version.split('+')[0]
TORCH_version = torch.__version__
TORCH = format_pytorch_version(TORCH_version)
def format_cuda_version(version):
return 'cu' + version.replace('.', '')
CUDA_version = torch.version.cuda
CUDA = format_cuda_version(CUDA_version)
# !pip install torch-scatter -f https://pytorch-geometric.com/whl/torch-{TORCH}+{CUDA}.html
# !pip install torch-sparse -f https://pytorch-geometric.com/whl/torch-{TORCH}+{CUDA}.html
# !pip install torch-cluster -f https://pytorch-geometric.com/whl/torch-{TORCH}+{CUDA}.html
# !pip install torch-spline-conv -f https://pytorch-geometric.com/whl/torch-{TORCH}+{CUDA}.html
# !pip install torch-geometric
# + colab={"base_uri": "https://localhost:8080/"} id="F1op-CbyLuN4" outputId="4347455f-8568-43bc-a093-adeeb2a0293d"
# %matplotlib inline
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE
def visualize(h, color):
"""
Reduce la dimensionalidad del embedding a 2D utilizando t-SNE y
grafica.
"""
z = TSNE(n_components=2).fit_transform(h.detach().cpu().numpy())
plt.figure(figsize=(10,10))
plt.xticks([])
plt.yticks([])
plt.scatter(z[:, 0], z[:, 1], s=70, c=color, cmap="Set2")
plt.show()
# + [markdown] id="dszt2RUHE7lW"
# ## Dataset
#
# El dataset que utilizaremos se conoce como `Cora`, y contiene una **red de citaciones**, donde los nodos son artículos científicos y los enlaces entre ellos indican que uno cita al otro.
#
# Cada nodo está descripto además por un vector de 1433 features, los cuales se construyeron a partir de realizar una bolsa de palabras sobre el contenido del documento. Nuestra variable objetivo es la categoría a la cual pertenece cada documento. En total, tenemos 7 categorías posibles.
#
# Utilizaremos un conjunto de entrenamiento reducido, del orden del 5%, significativamente menor a lo que suele usarse en tareas habituales de aprendizaje supervisado.
#
# Este dataset fue introducido por [Yang et al. (2016)](https://arxiv.org/abs/1603.08861) como uno de los datasets del conjunto [`Planetoid`](https://pytorch-geometric.readthedocs.io/en/latest/modules/datasets.html#torch_geometric.datasets.Planetoid), el cual se utiliza como benchmark para modelos basados en grafos.
# + colab={"base_uri": "https://localhost:8080/"} id="imGrKO5YH11-" outputId="9ade2ede-d19e-4773-80bf-fdc66e2e5b54"
from torch_geometric.datasets import Planetoid
from torch_geometric.transforms import NormalizeFeatures
## Cargamos el dataset y normalizamos los features por fila
## (cada fila suma 1)
dataset = Planetoid(
root='data/Planetoid',
name='Cora',
transform=NormalizeFeatures()
)
print()
print(f'Dataset: {dataset}:')
print('======================')
print(f'Number of graphs: {len(dataset)}')
print(f'Number of features: {dataset.num_features}')
print(f'Number of classes: {dataset.num_classes}')
## Extraemos el primer (y único, en este caso) grafo del dataset
data = dataset[0]
print()
print(data)
print('===========================================================================================================')
# Imprimimos algunos datos estadísticos de nuestro dataset
print(f'Number of nodes: {data.num_nodes}')
print(f'Number of edges: {data.num_edges}')
print(f'Average node degree: {data.num_edges / data.num_nodes:.2f}')
print(f'Number of training nodes: {data.train_mask.sum()}')
print(f'Training node label rate: {int(data.train_mask.sum()) / data.num_nodes:.2f}')
print(f'Has isolated nodes: {data.has_isolated_nodes()}')
print(f'Has self-loops: {data.has_self_loops()}')
print(f'Is undirected: {data.is_undirected()}')
# + [markdown] id="5IRdAELVKOl6"
# ## Perceptrón Multicapa (MLP)
#
# Como modelo base, entrenamos un perceptrón multicapa. En teoría, deberíamos poder aprender de nuestros ejemplos etiquetados utilizando sólo los features de los nodos, sin tener en cuenta la información relacional (es decir, el grafo). Veamos qué tan bien se comporta este modelo.
# + colab={"base_uri": "https://localhost:8080/"} id="afXwPCA3KNoC" outputId="b578cca5-1b18-4161-82f6-a0ceb2444e74"
import torch
from torch.nn import Linear
import torch.nn.functional as F
class MLP(torch.nn.Module):
def __init__(self, hidden_channels):
super(MLP, self).__init__()
torch.manual_seed(12345)
self.lin1 = Linear(dataset.num_features, hidden_channels)
self.lin2 = Linear(hidden_channels, dataset.num_classes)
def forward(self, x):
x = self.lin1(x)
x = x.relu()
x = F.dropout(x, p=0.5, training=self.training)
x = self.lin2(x)
return x
model = MLP(hidden_channels=16)
print(model)
# + [markdown] id="L_PO9EEHL7J6"
# Nuestro MLP está definido por dos capas lineales, con función de activación [ReLU](https://pytorch.org/docs/stable/generated/torch.nn.ReLU.html?highlight=relu#torch.nn.ReLU) y regularización [dropout](https://pytorch.org/docs/stable/generated/torch.nn.Dropout.html?highlight=dropout#torch.nn.Dropout).
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="0YgHcLXMLk4o" outputId="c2a4ba8c-cec2-4609-951a-2caa34935ec5"
model = MLP(hidden_channels=16)
criterion = torch.nn.CrossEntropyLoss() # Define loss criterion.
optimizer = torch.optim.Adam(model.parameters(), lr=0.01, weight_decay=5e-4) # Define optimizer.
def train():
model.train()
optimizer.zero_grad() # Clear gradients.
out = model(data.x) # Perform a single forward pass.
loss = criterion(out[data.train_mask], data.y[data.train_mask]) # Compute the loss solely based on the training nodes.
loss.backward() # Derive gradients.
optimizer.step() # Update parameters based on gradients.
return loss
def test():
model.eval()
out = model(data.x)
pred = out.argmax(dim=1) # Use the class with highest probability.
test_correct = pred[data.test_mask] == data.y[data.test_mask] # Check against ground-truth labels.
test_acc = int(test_correct.sum()) / int(data.test_mask.sum()) # Derive ratio of correct predictions.
return test_acc
for epoch in range(1, 201):
loss = train()
if epoch % 5 == 0:
print(f'Epoch: {epoch:03d}, Loss: {loss:.4f}')
# + [markdown] id="kG4IKy9YOLGF"
# After training the model, we can call the `test` function to see how well our model performs on unseen labels.
# Here, we are interested in the accuracy of the model, *i.e.*, the ratio of correctly classified nodes:
# + colab={"base_uri": "https://localhost:8080/"} id="dBBCeLlAL0oL" outputId="59e278f5-996e-4c3d-d795-37089479ea0c"
test_acc = test()
print(f'Test Accuracy: {test_acc:.4f}')
# + [markdown] id="_jjJOB-VO-cw"
# As one can see, our MLP performs rather bad with only about 59% test accuracy.
# But why does the MLP do not perform better?
# The main reason for that is that this model suffers from heavy overfitting due to only a **small amount of training nodes**, and therefore generalizes poorly to unseen node representations.
#
# It also fails to incorporate an important bias into the model: **Cited papers are very likely related to the category of a document**.
# That is exactly where Graph Neural Networks come into play and can help to boost the performance of our model.
#
#
# + [markdown] id="_OWGw54wRd98"
# ## Entrenamiento de una red neuronal de grafos (GNN)
#
# Ahora cambiamos nuestras capas lineales `torch.nn.Linear` por GNN de PyTorch. Vamos a utilizar la clase [`GCNConv`](https://pytorch-geometric.readthedocs.io/en/latest/modules/nn.html#torch_geometric.nn.conv.GCNConv), introducida en [Kipf et al. (2017)](https://arxiv.org/abs/1609.02907). La misma está definida como
#
#
# $$
# \mathbf{h}_v^{(k + 1)} = \mathbf{W}^{(k + 1)} \sum_{w \in \mathcal{N}(v) \, \cup \, \{ v \}} \frac{1}{\sqrt{(k_v+1)(k_w+1)}} \cdot \mathbf{h}_w^{(k)},
# $$
#
# donde $\mathbf{W}^{(k + 1)}$ es una matriz de pesos entrenable de forma `[num_output_features, num_input_features]`. Para comparar, una capa lineal está definida como
#
#
# $$
# \mathbf{h}_v^{(k + 1)} = \mathbf{W}^{(k + 1)} \mathbf{h}_v^{(k)},
# $$
#
# la cual no utiliza información de los vecinos.
# + colab={"base_uri": "https://localhost:8080/"} id="fmXWs1dKIzD8" outputId="8ee4cc45-abfa-47a4-f790-2c582ac6671b"
from torch_geometric.nn import GCNConv
class GCN(torch.nn.Module):
def __init__(self, hidden_channels):
super(GCN, self).__init__()
torch.manual_seed(1234567)
self.conv1 = GCNConv(dataset.num_features, hidden_channels)
self.conv2 = GCNConv(hidden_channels, dataset.num_classes)
def forward(self, x, edge_index):
x = self.conv1(x, edge_index)
x = x.relu()
x = F.dropout(x, p=0.5, training=self.training)
x = self.conv2(x, edge_index)
return x
model = GCN(hidden_channels=16)
print(model)
# + [markdown] id="XhO8QDgYf_Q8"
# Visualizamos el embedding para la red GCN **sin entrenar**. Utilizamos para eso una reducción de la dimensión mediante el algoritmo [**t-SNE**](https://scikit-learn.org/stable/modules/generated/sklearn.manifold.TSNE.html) para pasar de las 7 dimensiones del embedding (una por clase) al plano.
# + colab={"base_uri": "https://localhost:8080/", "height": 578} id="ntt9qVFXlk6A" outputId="a5496675-1eb6-4518-ca0d-12768adf5ff6"
model = GCN(hidden_channels=16)
model.eval()
out = model(data.x, data.edge_index)
visualize(out, color=data.y)
# -
# Entrenamos el modelo
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="p3TAi69zI1bO" outputId="9a7a854e-e144-4fd9-bb47-8bc0aa587bdf"
model = GCN(hidden_channels=16)
optimizer = torch.optim.Adam(model.parameters(), lr=0.01, weight_decay=5e-4)
criterion = torch.nn.CrossEntropyLoss()
def train():
model.train()
optimizer.zero_grad() # Clear gradients.
out = model(data.x, data.edge_index) # Perform a single forward pass.
loss = criterion(out[data.train_mask], data.y[data.train_mask]) # Compute the loss solely based on the training nodes.
loss.backward() # Derive gradients.
optimizer.step() # Update parameters based on gradients.
return loss
def test():
model.eval()
out = model(data.x, data.edge_index)
pred = out.argmax(dim=1) # Use the class with highest probability.
test_correct = pred[data.test_mask] == data.y[data.test_mask] # Check against ground-truth labels.
test_acc = int(test_correct.sum()) / int(data.test_mask.sum()) # Derive ratio of correct predictions.
return test_acc
for epoch in range(1, 101):
loss = train()
print(f'Epoch: {epoch:03d}, Loss: {loss:.4f}')
# + [markdown] id="opBBGQHqg5ZO"
# Observamos las predicciones
# + colab={"base_uri": "https://localhost:8080/"} id="8zOh6IIeI3Op" outputId="4d493fec-4437-4d9a-fa13-2412ff1e7497"
test_acc = test()
print(f'Test Accuracy: {test_acc:.4f}')
# + [markdown] id="yhofzjaqhfY2"
# Proyectamos nuevamente el embedding en dos dimensiones para apreciar los clusters
# + colab={"base_uri": "https://localhost:8080/", "height": 578} id="9r_VmGMukf5R" outputId="b4f41cab-6203-42df-d5a8-86c6b8f13cdd"
model.eval()
out = model(data.x, data.edge_index)
visualize(out, color=data.y)
| notebooks/Notebook5_Tutorial_PyTorch_Geometric.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#hide
#default_exp training
from nbdev.showdoc import show_doc
# # Training with Letters Data
# > Train on letters data.
# +
from car_speech.fname_processing import load_fnames
from car_speech.pipeline import *
import string
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import tensorflow as tf
from tensorflow.keras.layers.experimental import preprocessing
from tensorflow.keras import layers
from tensorflow.keras import models
# -
# ## Configuration
# +
DATASET_TYPE = 'letters' # or 'letters'
label_strings = np.array(list(string.ascii_uppercase))
# load classified filenames
filenames = load_fnames('noise_levels/letter_noise_levels/IDL.data')
print('number of files:', len(filenames))
# -
# ## Pipeline (Audio to Spectrogram)
# +
# shuffle
filenames = shuffle_data(filenames)
# Train/Validation/Test Split
split_result = train_test_split(filenames)
train_files = split_result[0]
val_files = split_result[1]
test_files = split_result[2]
# Process data using the combined pipeline
spectrogram_ds = preprocess_dataset(train_files, DATASET_TYPE)
train_ds = spectrogram_ds
val_ds = preprocess_dataset(val_files, DATASET_TYPE)
test_ds = preprocess_dataset(test_files, DATASET_TYPE)
print("Pipeline Completed")
# -
# ## Train on training set
# Split data into batches
batch_size = 64
train_ds = train_ds.batch(batch_size)
val_ds = val_ds.batch(batch_size)
# Reduce read latency during training
# +
AUTOTUNE = tf.data.experimental.AUTOTUNE
train_ds = train_ds.cache().prefetch(AUTOTUNE)
val_ds = val_ds.cache().prefetch(AUTOTUNE)
# -
# Model
# +
for spectrogram, _ in spectrogram_ds.take(1):
input_shape = spectrogram.shape
print('Input shape:', input_shape)
num_labels = len(label_strings)
norm_layer = preprocessing.Normalization()
norm_layer.adapt(spectrogram_ds.map(lambda x, _: x))
model = models.Sequential([
layers.Input(shape=input_shape),
preprocessing.Resizing(32, 32),
norm_layer,
layers.Conv2D(32, 3, activation='relu'),
layers.Conv2D(64, 3, activation='relu'),
layers.MaxPooling2D(),
layers.Dropout(0.25),
layers.Flatten(),
layers.Dense(128, activation='relu'),
layers.Dropout(0.5),
layers.Dense(num_labels),
])
model.summary()
# -
model.compile(
optimizer=tf.keras.optimizers.Adam(),
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'],
)
# Train
EPOCHS = 25
history = model.fit(
train_ds,
validation_data=val_ds,
epochs=EPOCHS,
callbacks=tf.keras.callbacks.EarlyStopping(verbose=1, patience=2),
)
# Save model
model_dir = 'models'
# Create a folder and save the model
model_name = 'model_single_digit' #Make sure you change this name. DO NOT OVERWRITE TRAINED MODELS.
# model.save(os.path.join(model_dir, model_name))
# Plot loss
metrics = history.history
plt.plot(history.epoch, metrics['loss'], metrics['val_loss'])
plt.legend(['loss', 'val_loss'])
plt.show()
# ## Test
# +
test_audio = []
test_labels = []
for audio, label in test_ds:
test_audio.append(audio.numpy())
test_labels.append(label.numpy())
test_audio = np.array(test_audio)
test_labels = np.array(test_labels)
# +
y_pred = np.argmax(model.predict(test_audio), axis=1)
y_true = test_labels
test_acc = sum(y_pred == y_true) / len(y_true)
print(f'Test set accuracy: {test_acc:.0%}')
# -
# Plot confusion matrix
confusion_mtx = tf.math.confusion_matrix(y_true, y_pred)
plt.figure(figsize=(10, 8))
sns.heatmap(confusion_mtx, xticklabels=label_strings, yticklabels=label_strings,
annot=True, fmt='g')
plt.xlabel('Prediction')
plt.ylabel('Label')
plt.show()
| 06_Training_letters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="NuQ74qNmwU3H"
# !pip install transformers
# + id="Yy52ATY9wcco"
from transformers import pipeline
classifier = pipeline("zero-shot-classification",
model="facebook/bart-large-mnli")
# + id="X37h0fCsw731"
# !ls ~/.cache/huggingface/transformers
# + colab={"base_uri": "https://localhost:8080/"} id="nhaI5ppR0FY8" executionInfo={"status": "ok", "timestamp": 1618411044734, "user_tz": 240, "elapsed": 1454, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQ5abnImE35Fu-BNSrmYlpO4C_dqQpuG2gjJmy5Q=s64", "userId": "11004855458032977781"}} outputId="67e2d5fd-122d-4571-f07b-7220bdc5aed7"
sequence_to_classify = "one day I will see the world"
candidate_labels = ['travel', 'cooking', 'dancing']
output = classifier(sequence_to_classify, candidate_labels)
print(output)
# + [markdown] id="IvC6NvAk0bfT"
# [Source](https://huggingface.co/facebook/bart-large-mnli)
# + id="PXfOgcqi0kyW" executionInfo={"status": "ok", "timestamp": 1618411154399, "user_tz": 240, "elapsed": 507, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQ5abnImE35Fu-BNSrmYlpO4C_dqQpuG2gjJmy5Q=s64", "userId": "11004855458032977781"}} outputId="1108e06c-c04b-44c8-9d30-0f89344ec9cd" colab={"base_uri": "https://localhost:8080/"}
# !free -g
| classification/zero-shot-text-classification-bart-pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import math
import pickle
import matplotlib.pyplot as plt
# %matplotlib inline
def loading_data(filepath):
#loading data
ml = pd.read_csv(filepath, header=None)
ml.columns = ['User','Item','ItemRating']
return ml
def create_interaction_cov(ml):
# creating matrix from transactions
ml_user_item_matrix = ml.pivot(index='User', columns='Item', values='ItemRating')
ml_user_item_matrix = ml_user_item_matrix.fillna(0)
ml_user_item_matrix = ml_user_item_matrix.reindex(index=range(ml_user_item_matrix.index.max() + 1), columns= range(ml_user_item_matrix.columns.max() + 1), fill_value=0)
# create user covariance matrix
cov_ml = np.dot(ml_user_item_matrix.values,ml_user_item_matrix.T.values)
return ml_user_item_matrix, cov_ml
def neighbors(cov_ml, user):
# 'cov_ml' is the covariance matrix
nn = np.argsort(-cov_ml[user,:]) # all neighbors sorted descending
# I previously had a -1, double check
return nn
def prediction(ml_user_item_matrix, cov_ml, nn, user, item, number_of_n=10):
neighbors = []
# populating 'neighbors' with defined 'number_of_n'
for n in nn:
if len(neighbors) < number_of_n:
if n in ml_user_item_matrix.index:
# enforcing that neighbors have rated the item
if ml_user_item_matrix[item][n] != 0:
neighbors.append(n)
# total weight of N neighbors
total_distance = sum(cov_ml[user, neighbors])
# get the proportion of weight for each neighbor
weighted_input = cov_ml[user, neighbors] / total_distance
# creating prediction from weighted average of neighbors ratings
# getting the rating of the item to the predicted from each of the neighbors
neighbors_ratings = []
for e in neighbors:
neighbors_ratings.append(ml_user_item_matrix[item][e])
weighted_rate = neighbors_ratings * weighted_input
prediction_rate = weighted_rate.sum()
return weighted_input, neighbors_ratings, prediction_rate
# ### Load training set and Create interaction matrix
ml2 = loading_data("data/ml1m-train-clean")
ml_user_item_matrix, cov_ml = create_interaction_cov(ml2)
ml_user_item_matrix.shape, cov_ml.shape
# > There are 6041 users and 3953 items
# + When we create a User covariance similarity matrix we end up with a 6041x6041 U/U matrix
ml_user_item_matrix.head()
# find neighbors of user "1"
u = 1
nn = neighbors(cov_ml, u)
nn
i = 2018
# to make prediction for user ='1' and item '2018'
weighted_input, ratings_for_item, prediction_rate = prediction(ml_user_item_matrix, cov_ml, nn, u, i, number_of_n=10)
prediction_rate
# ## Load validation set
val = pd.read_csv("data/ml1m-validation-clean", header=None)
val.columns = ['User','Item','ItemRating']
val.head()
# ## Make Predictions
# To make predictions, we need to:
# + find the neighbors of user "u"
# + calculate a weighted average of the ratings of movie "i" for the neighbors of "u"
# Lets calculate the neighbors for each of the users in the test/val set and add them to the dataframe
val["Neighbors"] = val["User"].apply(lambda x: neighbors(cov_ml, x))
val.head()
users = val["User"].values
items = val["Item"].values
nn = val["Neighbors"].values
# Lets test the prediction process through the first **5 items** in the test set, using **10 nearest neighbors**
prediction_rates = []
for i in range(5):
prediction_rates.append(prediction(ml_user_item_matrix, cov_ml, nn[i], users[i], items[i], number_of_n=10)[2])
prediction_rates
# Adding predictions to dataframe
pred200 = pd.Series(prediction_rates200)
val200['Prediction'] = pred200.values
# ## Calculate RMSE
# The prediction loop was used for a range of neighbors. Each of the outputs added to a column named "Prediction" and the dataframe were pickled with the following names:
# + val200.pickle
# + val300.pickle
# + val400.pickle
# + val500.pickle
# + val600.pickle
# + val700.pickle
# + val800.pickle
# + val900.pickle
# + val1000.pickle
def calculate_RMSE(pickle_path):
# Load pickled file with "Prediction"
nn = pd.read_pickle(pickle_path)
# Calculate "Error" for each prediction
nn["Error"] = (nn["ItemRating"] - nn["Prediction"])**2
number_of_preds = nn.shape[0]
# Root mean square error
rmse = math.sqrt(nn["Error"].sum() / number_of_preds)
return rmse
rmse200 = calculate_RMSE("data/val200.pickle")
rmse200
rmse300 = calculate_RMSE("data/val300.pickle")
rmse400 = calculate_RMSE("data/val400.pickle")
rmse500 = calculate_RMSE("data/val500.pickle")
rmse600 = calculate_RMSE("data/val600.pickle")
rmse700 = calculate_RMSE("data/val700.pickle")
rmse800 = calculate_RMSE("data/val800.pickle")
rmse900 = calculate_RMSE("data/val900.pickle")
rmse1000 = calculate_RMSE("data/val1000.pickle")
x = np.linspace(200, 1000, 9)
x
rmse_values = [rmse200, rmse300, rmse400, rmse500, rmse600, rmse700, rmse800, rmse900, rmse1000]
data = { "neighbors": x, "COV-RMSE": rmse_values}
data
neighbors = pd.DataFrame(data = data, columns=["neighbors", "COV-RMSE"])
neighbors.head()
ax = neighbors.plot( x="neighbors",style=['rx'], figsize=(8,4));
ax.set_ylabel("RMSE");
ax.set_xlabel("Neighbors");
ax.set_title("Evaluation Scores");
# ## Comparing results
l2v_exp = pd.read_pickle("data/l2v-neighbors.pickle")
l2v_neighbors = l2v_exp[["pred_neighbors","eval_score"]]
l2v_neighbors.columns = ["pred_neighbors","L2V-RMSE"]
l2v_neighbors.head()
# +
# adding COV-RMSE to dataframe containing l2V-RMSE
# l2v_neighbors.loc[l2v_neighbors.pred_neighbors == 200, "COV-RMSE"] = rmse200
# -
ax = l2v_neighbors.plot( x="pred_neighbors",style=['o','rx'], figsize=(8,4));
ax.set_ylabel("RMSE");
ax.set_xlabel("Neighbors");
ax.set_title("Evaluation Scores");
# > The hyperparameters for L2v in the comparison are the experiments listed below (which can be found in "/CONFIG" folder in s3)
import sqlalchemy
import psycopg2
db_string = "postgresql://localhost:5433/jaimealmeida"
engine = sqlalchemy.create_engine(db_string)
the_frame = pd.read_sql_query("SELECT experimentid, pred_neighbors, eval_score FROM %s WHERE pred_neighbors > 100 and pred_neighbors < 1100;" % "experiments", engine)
the_frame.sort_values(by="pred_neighbors")
| notebooks/04-JJA-Covariance-Similarity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Resources
#
#
# ## Github source code Python
#
# 1. https://github.com/JWarmenhoven/ISLR-python
#
# 2. https://github.com/hardikkamboj/An-Introduction-to-Statistical-Learning
#
# 3. https://github.com/alexandrasouly/ISLR-but-python
#
#
# ## Course
#
# 1. https://www.dataschool.io/15-hours-of-expert-machine-learning-videos/
#
# 2. A summary of this book is available as a course by Stanford University on edX.
#
# https://learning.edx.org/course/course-v1:StanfordOnline+STATSX0001+1T2020/home
#
# You can enroll and watch the course for Free (If you have not used edX before, then during enrollment, you should select "Audit this Course" to watch it for free)
# Same is available on You tube, but some portions are cut off in Youtube
#
#
# ## Review
#
# 1. https://towardsdatascience.com/islr-review-an-ideal-textbook-to-start-your-ml-journey-7b548ba8977f
#
#
#
| Stat_Learning_Book/Resources.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Seaborn Refresher
#
# Let's review using Seaborn and Pandas to load up some data and then pair plot it.
#
# We'll be using the same tools that we used last week for this
# - [pandas](pandas.pydata.org) for data handling (our dataframe library)
# - [seaborn](seaborn.pydata.org) for _nice_ data visualization
#
# Shortly we'll also by trying out:
#
# - [scikit-learn](scikit-learn.org) an extensive machine learning library.
# - [numpy](numpy.org) - a fundamental maths library best used by people with a strong maths background. We won't explore it much today, but it does have some useful methods that we'll need. It underlies all other mathematical and plotting tools that we use in Python.
#
# We'll be using scikit-learn over the next few weeks, and it's well worth reading the documentation and high level descriptions.
#
# As before, the aim is to get familiar with code-sharing workflows - so we will be doing pair programming for the duration of the day! _You will probably want to take a moment to look at the documentation of the libraries above - especially pandas_
#
# The other useful resource is Stack Overflow - if you have a question that sounds like 'how do I do {x}' then someone will probably have answered it on SO. Questions are also tagged by library so if you have a particular pandas question you can do something like going to https://stackoverflow.com/questions/tagged/pandas (just replace the 'pandas' in the URL with whatever library you're trying to use.
#
# Generally answers on SO are probably a lot closer to getting you up and running than the documentation. Once you get used to the library then the documentation is generally a quicker reference. We will cover strategies for getting help in class.
#
# ## Git links
#
# We will be working through using GitHub and GitKraken to share code between pairs. We will go through all the workflow in detail in class but here are some useful links for reference:
#
# - GitKraken interface basics: https://support.gitkraken.com/start-here/interface
# - Staging and committing (save current state -> local history): https://support.gitkraken.com/working-with-commits/commits
# - Pushing and pulling (sync local history <-> GitHub history): https://support.gitkraken.com/working-with-repositories/pushing-and-pulling
# - Forking and pull requests (request to sync your GitHub history <-> someone else's history - requires a _review_):
# - https://help.github.com/articles/about-forks/
# - https://help.github.com/articles/creating-a-pull-request-from-a-fork/
#
# ## Step 1: Read in the dataset and pairplot
#
# For this exercise, we will be using the Tips dataset that you can find in the same directory as this notebook. This is a widely used dataset in machine learning, and while not related to minerals and energy, it is sufficient for our purpose. The dataset relates total bills at US restaurants to tip size, as well as the sex of the tipper, whether they smoke, the day of the week, the kind of meal, and the number of people.
#
# In pairs work out how to read this data into a pandas dataframe, then use Seaborn to pairplot the species in the dataset.
#
# Seaborn happens to have this dataset built in. Run the next cell to see the built in data. Then modify the code to open the dataset from a CSV file. The dataset can take a little while to load, so be patient - the dataset will appear.
import seaborn as sns
tips = sns.load_dataset('tips')
tips
import pandas as pd
tips = pd.read_csv('../data/tips.csv')
tips.head(10)
# ## Step 2 : Find a linear regression with Seaborn
#
# Now that you've seen some pairplots (tips vs tip size are the most meaningful comparisons), use Seaborn to find lines of best fit in this dataset.
#
# There are a few different ways to do this. Try using regplot.
#
# You may notice a "FutureWarning". Ignore this - Python is often in a state of flux and these types of warning are common. Often with major packages like Seaborn you'll find that a soon-to-be-released version of the library will not create these warnings.
sns.regplot("tip", "size", data=tips, scatter=True, color="blue")
sns.regplot("total_bill", "tip", data=tips, scatter=True, color="blue")
# ## Step 3: Linear regression with scikit-learn
#
# Scikit-learn provides machine learning tools in several categories. These include supervised learning and unsupervised learning. We'll start working with unsupervised learning next week. Supervised learning is about finding a model for features that can be measured and some labelling that we have for the available data. If, for example, we have lithium assays and we want to try to predict lithium based on sensor data from a portable spectrometer, then the lithium assays are the labels and the measured intensities at different wavelengths are the measured features. This kind of supervised learning is called regression.
#
# There's another kind of supervised learned which is called classification, this is what we're doing when we want to assign observed data to different discrete classes. Regression can sometimes be used, with minor additions, to classify data as well. For example, with our lithium spectral regression model we could classify samples as being high in lithium or low in lithium simply by using a threshold value that we set. There are more sophisticated ways to classify, which will be covered in later weeks.
#
# We use the estimator API of scikit-learn to do regression.
#
# ### The Estimator API of scikit-learn
#
# There are a few steps to follow when using the estimator API. These steps are the same for all methods that scikit-learn implements, not just for linear regression.
#
# 1. Choose a class of model by importing the appropriate estimator class. In our case we want to import Linear Regression. Here's how we can do it.
#
# First import LinearRegression from scikit-learn. Use this code:
#
# ```from sklearn.linear_model import LinearRegression```
from sklearn.linear_model import LinearRegression
# Now create an "instance" of the LinearRegression class. We can do it like this:
#
# ```model = LinearRegression(fit_intercept=True)```
#
# To check that this has worked look at the model object after it's created. It should tell you about some of its settings.
#
# ```model```
model = LinearRegression(fit_intercept=True)
model
# These settings are also called hyperparameters. We'll encounter hyperparameters again next week, and will talk about them in more detail then. They're often very important in working out whether our model is well fitted to the data.
#
# 2. Next we need to arrange a pandas dataframe (like "tips") into a features matrix and a target vector.
#
# Search on the Internet for this. I know that Stack Overflow will be helpful. You will need to look at the column names in the dataframe to find the names of the two columns that are important to us. Do this in the next cell.
#
# The notation is a bit strange! The two pairs of "[ ]" as "[[ ]]" that you will see is correct.
y = tips['tip']
y
x = tips[['total_bill']]
x
model.fit(x,y)
model.coef_
model.intercept_
# 3. Fit the model to your data by using the fit() method of the LinearRegression object.
#
# Again, look at the documentation for how to apply this. You'll need to provide your features matrix (X) and target vector (y) as parameters to the fit method.
#
# #### Congratulations you've trained your first machine learning model!
#
# As this is a two dimensional linear model, it has two parameters. The line's intercept and slope. The notation that scikit-learn uses is a little unfriendly. Its convention is to add underscores to the names of the parameters it finds. Also, it calls the slope "coef".
#
# After fitting the model, find the coefficients with ```model.coef_``` and ```model._intercept_```.
# #### Now that we've trained a model, we should make predictions!
#
# 6. Make predictions!
#
# This is also more complicated with scikit-learn than it is with Seaborn.
#
# For a given, single value for a feature (ie a meal cost) we can predict a label. For example, for a meal cost of $20, we could make a prediction with:
#
# ```predicted_tip = model.predict(20)```
#
# But to find the smooth line that seaborn finds we need to explicitly tell scikit-learn that we want to do a prediction for all of the meal costs that we're interested in. To do this we
# use a new library called "numpy" and a method called linspace (which is short for linear spacing).
#
# First we need to import numpy.
#
# ```import numpy as np```
#
# While I used predicted_tip above as an example of a predicted target array, and 20 is an example of x, I'll now switch to the usual y and x conventions used in tutorials with scikit-learn. You can of course use any variables names you, and in your own code it's best to use descriptive names that mean something in the domain of your industry, like 'predicted_tip", or "octane_rating".
#
# We need to use the linspace method in numpy. Use it like this:
#
# ```xfit = np.linspace(0, 60)```
#
# This will create a collection of meal costs, in order, starting from 0 dollars up to 60 dollars. This is what we need, but this collection isn't formatted correctly for scikit-learn. To make it work with scikit-learn we next have to adjust the format with this instruction:
#
# ```xfit_reshaped = xfit[:, np.newaxis]
# yfit = model.predict(xfit_reshaped).```
#
# yfit now contains our predicted tips. Type ```yfit``` to see them numerically.
#
# Try this all out in the next cell. Take it step by step. Don't try to run this all in one go, but build it up line by line, checking that you do not get errors after each line.
predicted_tip = model.predict(20)
predicted_tip
import numpy as np
xfit = np.linspace(0, 60)
xfit
xfit_reshaped = xfit[:, np.newaxis]
yfit = model.predict(xfit_reshaped)
yfit
# We can also plot these results, but lets stop here.
#
# We could use the default plotting functions that Pandas provides for this. But for report purposes you may, in future, want to find out how to use Seaborn for this.
#
# We can create dataframes from this data with code like:
#
# ```
# DataFrame({'meal_cost':xfit, 'tips':yfit})
# ```
df = pd.DataFrame({'meal_cost':xfit, 'tips':yfit})
sns.set()
ax = df.plot('meal_cost','tips')
tips.plot('total_bill','tip',marker='o',linewidth=0,ax=ax,color='red')
# ### Exercise: Create a linear function with scikit-learn. Then add noise.
#
# 1. Scikit-learn includes a built in function to quickly create datasets for experimenting with the estimator API.
#
# Find out about the make_regression method in sklearn.datasets.
#
# Use this to make a noisy line with 100 samples. Use n_features to set the number of features, and use noise to adjust gaussian noise that is added.
#
# Add outliers and see how the fitting is impacted. LinearRegression can report R^2 values. Use the "score" method. Google and Stack Overflow will help with usage.
# ### Exercise: Create a linear function with seaborn. Then add noise.
#
# While we'll be using scikit-learn over the next few weeks and it's helpful to keep working with it, it's really quite painful compared to Seaborn. Looking at residuals is an exploratory task that Seaborn is better suited to than scikit-learn.
#
# Try this code, which plots the residuals after gaussian noise is added to a simple y = x line. Try to get the gist of how it works.
#
# ```
# import numpy as np
# import seaborn as sns
# sns.set(style="whitegrid")
#
# rs = np.random.RandomState(7)
# x = rs.uniform(0, 100, 10000)
# y = x + rs.normal(0, 1, 10000)
#
# sns.residplot(x, y, lowess=True, color="g")
# ```
#
# The RandomState object is part of the Numpy numerical package which we won't explore in detail at this time. It is a collection of mathematics functions which underlies all other mathematical libraries that we've been using, such as Seaborn and scikit-learn. RandomState is used for generating random numbers from distributions.
#
# A uniform distribution means that all of the values that may be returned are equally likely. When we throw dice we are sampling from a uniform distribution. Here we tell Python that we want random numbers between 0 and 100, all equally likely, and we want 10000 of them.
#
# A normal (or gaussian) distribution returns values which are most likely to be near the mean, falling off symetrically to either side. It is the "bell" curve that you've seen many times. Here we say that we want the error that we add to our simple line to have a mean of zero, and a standard deviation of 1.
#
# Seaborn's residplot function plots the residuals after fitting a line to the data. With a normal distribution we expect to see these residuals evenly scattered around zero.
#
# An example of a heavy tailed distribution is the gamma distribution. This is often used to model failure likelihood for machines. Unlike the normal distribution it is not symmetric. In quality control applications it quickly peaks after a short lifetime, but then has a long tail that extends many years into the future. This makes sense as we expect most failures to be early in the life of a machine because of manufacturing faults, after that the failure time is less predictable, but we all know of machines or gadgets that seem to last forever. Google will quickly bring up examples of the shape.
#
# Try the code above again, but substitute ```rs.normal (0, 1, 10000)``` with ```rs.gamma(2, 2, 10000)```.
#
# How would you change this code to create a heteroscadistic error?
# +
import numpy as np
import seaborn as sns
sns.set(style="whitegrid")
rs = np.random.RandomState(7)
x = rs.uniform(0, 100, 10000) # random numbers between 0 and 100
y = x + rs.normal(0, 1, 10000) # add gaussian noise between 0 and 1
sns.residplot(x, y, lowess=True, color="g") # residuals plot
# +
y = x + rs.gamma(2, 2, 10000)
sns.residplot(x, y, lowess=True, color="g")
# -
# ### Exercise: Examine the linear dataset that you have brought
#
# Find a least-squares fit using scikit-learn, and plot the residuals. Are the residuals gaussian? Is there homoscedasticity? Do you have outliers?
# ### Exercise: Metal recovery vs %sulphur in feed
#
# In this exercise we're going to look at a typical minerals engineering problem. We have data collected in laboratory batch floatation tests on samples taken from different parts of a base metal orebody. It appears that there is a simple relationship between metal recovery and the percent of sulphur in the sample. We can see that recovery is increasing with sulphur.
#
# 1. Open and scatterplot the file metal_recovery_vs_sulphur.csv
df = pd.read_csv('../data/metal_recovery_vs_sulphur.csv')
df.head(10)
sns.regplot("Feed Sulphur", "Metal Recovery", data=df, scatter=True, color="blue")
# 2. Find the least-squares linear fit for this data, without using any data transformations. Plot the results. Using Pandas and Seaborn may be the easiest way to approach this.
#
# The results aren't terrible, but there are some problems. There is a definite curve in the data and the line is unable to fit through all points. It also poor at extrapolating. This curve will happily predict greater than 100% recovery at feed sulphur beyond around 2%. It's also happy to advise metal recoveries of around 40% with no feed sulphur. That may seem reasonable to a data scientist, but domain experts will regard that as ridiculous.
#
# 3. Try transforming the sulphur feed percentages before fitting. We'd like to know if the data can be made to look more linear through a simple algebraic relationship. Domain knowledge may help here. The general shape of the curve suggests that there may be a power relationship here. What happens if you regress again the square root of feed sulphur? What kind of transformation could lead a metal recovery that is limited below 100%? Maybe it's worth trying the reciprocal of feed sulphur?
df['Feed_Sulphur_Squared'] = df['Feed Sulphur']**2
sns.regplot("Feed_Sulphur_Squared", "Metal Recovery", data=df, scatter=True, color="blue")
df['Feed_Sulphur_Square_Root'] = df['Feed Sulphur']**0.5
sns.regplot("Feed_Sulphur_Square_Root", "Metal Recovery", data=df, scatter=True, color="blue")
df['Feed_Sulphur_Reciprocal'] = 1 / df['Feed Sulphur']
sns.regplot("Feed_Sulphur_Reciprocal", "Metal Recovery", data=df, scatter=True, color="blue")
| notebooks/session1and2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Data Structures
#
# We can think of data structures in Python as objects that hold data with special properties.
#
# Data can be numbers, strings, or even boolean (True or False) values.
#
# Two common data structures we use in Python are lists and dictionaries.
#
# We can use the special properties of these objects to manipulate them.
# ## Lists
#
# Lists are data structures, where the data is stored in between square brackets [ ] and separated by commas.
#
# For example, we can create a list of popular video games like this:
#
# my_game_list = ["Minecraft", "Grand Theft Auto", "Super Smash Bros", "Fortnite"]
#
# We can access elements in a list by "indexing" using square brackets [ ] and passing numbers 0,1,2,... to access its position in the list. Note, in Python, the left-most element has position 0.
#
# To access Super Smash Bros, for example, from `my_game_list` in Python we would say
#
# my_game_list[2]
#
# which will return
#
# “Super Smash Bros”
#
# Run the cell below to print the lists and see some of the properties.
# +
# List examples
my_number_list = [1,2,3,4]
my_string_list = ["a","b","c","d"]
my_mixed_list = [1,"a",2,"b"]
my_game_list = ["Minecraft", "Grand Theft Auto", "Super Sm<NAME>", "Fortnite"]
print(my_number_list)
print(my_string_list)
print(my_mixed_list)
print(my_game_list)
print("\n")
print("The first element in", my_number_list, " can be accessed as follows: my_number_list[0]")
print("The first element in", my_number_list, " is ", my_number_list[0])
# -
# ### 📗Challenge 1 (Level 1)
#
# Using the information above, find the **third** element in `my_number_list`
#
# Remember, indexing in Python starts with **0**.
#
# **[Hint]** The **second** element in a list can be accessed using the notation `my_number_list[1]`.
# ✏️your code here
# ## Dictionaries
#
# Dictionaries are another example of a data structure, but with different properties.
#
# In a paper dictionary, each word has its own definition (or multiple definitions). When we want to know the definition of a word, we find that word in the dictionary.
#
# Dictionaries in Python are similar, but instead of words being used to access definitions, **keys** are used to access other **data structures** (such as a string or a list).
#
# Here is what a dictionary in Python looks like:
#
# dictionary = {Key1: data_structure_1,
# Key2: data_structure_2,
# ...
# KeyN: data_structure_N}
#
# You can also think of Python dictionaries as magic spell books, where the name of the spell will access one specific spell, but no other.
#
# Spells = {"Slow-mo" : "Slow down an enemy for up to 10 seconds", "Fire-ball": "create a fireball from any element around you"}
#
# We can access elements in a dictionary by using square brackets **[]** that include the **key**. For example, to access the slow-mo spell we would say
#
# Spells["Slow-mo"]
#
# which will return
#
# "Slow down an enemy for up to 10 seconds"
#
# Run the code below.
# +
# Constructing a dictionary
spells = {"Slow-mo" : "Slow down an enemy for up to 10 seconds",
"Fire-ball": "Create a fireball from any element around you",
"Combo-spells": ["Venomous Plants","Storm"]}
print("Spell dictionary")
display(spells)
print("Accessing slow motion spell using Python dictionaries ")
print(spells["Slow-mo"])
# -
# ### 📗Challenge 2 (Level 1)
#
# Use the `spells` dictionary in the cell below to access the `Fire-ball` spell.
#
# **[Hint]** The code to access a value in a dictionary is `my_dictionary["key"]`.
# ✏️your response here
# ### 📗Challenge 3 (Level 2)
#
# Now that you have played with dictionaries, access the `Combo-spells` list from the spells dictionary.
#
# What is the first spell in the list?
# ✏️your response here
# ## Pandas Dataframes
#
# Pandas [dataframes](https://www.geeksforgeeks.org/python-pandas-dataframe/) are another type of data structures.
#
# Pandas dataframes are two-dimensional, which means they have rows and columns. They can have different data types such as [strings](https://realpython.com/python-data-types/#strings), [integers](https://realpython.com/python-data-types/#integers), [floating point (decimal) numbers](https://realpython.com/python-data-types/#floating-point-numbers), or [boolean](https://realpython.com/python-data-types/#boolean-type-boolean-context-and-truthiness).
#
# |Column 1| Column 2|
# | - | - |
# |Row 1: Value 1| Row 1: Value 2|
# |Row 2: Value 1| Row 2: Value 2|
#
# Pandas dataframes can be thought of as an extension of dictionaries, where the keys hold the name of the column and the keys are usually lists with variables or objects inside them. If you are a very experienced magician, you probably have a number of different spells and categories:
#
# |Freezing Spells|Attacking Spells| Healing Spells |
# | - | - | - |
# |Slow-mo spell| Fire-ball spell|Revive spell
# |Full-freeze spell| Teleportation spell| Help a friend spell|
#
# Freezing Spells
#
# Slow-mo spell: slows down enemy for up to 10 seconds
# Full-freeze spell: freeze enemy for 30 seconds
#
# Attacking Spells
#
# Fire-ball spell: create a fireball from any element around you
# Teleportation spell: use this spell to randomly relocate an enemy
#
# Healing Spells
#
# Revive spell: use this spell when your life points are near zero for a 100 boost
# Help a friend spell: use this spell to give 50 points to one of your team members
#
#
# We can access spells in Pandas by combining the properties of dictionaries and lists: **keys** and **indexing**. Use the notebook to explore data structures.
#
# Each column name is a key, and the values are a list of spells that we can access via indexing. Run the cell below to get started.
# Import the Pandas library. "pd" is an alias, i.e. a shorted name
import pandas as pd
# We then call the Dataframe function from the Pandas library using the "dot" notation:
#
# pd.Dataframe()
#
# This function takes as input a dictionary. Let's use our `spells` dictionary from the previous exercise and parse it as a Pandas dataframe.
# +
# Create a Pandas dataframe - we will create it by using the DataFrame function within the Pandas library
# we can do this by using the dot notation
SlowMo = {"Slow-mo":"Slow down an enemy for up to 10 seconds"}
FullFre = {"Full-freeze":"Freeze enemy for 30 seconds"}
FireB = {"Fire-ball": "Create a fireball from any element around you"}
TeleP = {"Teleportation": "Use this spell to randomly relocate an enemy"}
RevI = {"Revive": "Use this spell when your life points are near zero for a 100 boost"}
HelpF = {"Help a friend": "Use this spell to give 50 points to one of your team members"}
spells = {"Freezing Spells" : [SlowMo,FullFre],
"Attacking Spells": [FireB,TeleP],
"Healing Spelsl": [RevI,HelpF]}
# -
my_dataframe = pd.DataFrame(spells)
# Let's look at our dataframe.
my_dataframe
# ### 📗Challenge 4 (Level 3)
#
# Use your knowledge of dictionaries to access all "Freezing Spells" in `my_dataframe`.
#
# Within Freezing Spells, access the "Full-freeze" spell
#
# **[Hint 1]** Use the notation below to access, in order of appearance, the column name and the index
#
# my_dataframe["Spell_Category"][]
#
# The data structure will be a list.
#
# Access the first element in that list.
#
# **[Hint 2]** Use the notation below to access, in order of appearance, the column name and the index
#
# my_dataframe["Spell_Category"][index number]
#
# Now access the "Full-freeze" spell
#
# **[Hint 3]** `my_dataframe["Spell_Category"][index number]["Spell_name"]`
# ✏️your response here
# This concludes our data structures notebook. Next you will have an opportunity to apply your knowledge of data structures to analyse real data.
# [](https://github.com/callysto/curriculum-notebooks/blob/master/LICENSE.md)
| SouthwesternAlbertaOpenData/data-structures-in-python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="tvEh-krrYwgs"
# # !apt-get install openjdk-8-jdk-headless -qq > /dev/null
# # !wget -q https://www-us.apache.org/dist/spark/spark-2.4.5/spark-2.4.5-bin-hadoop2.7.tgz
# # !tar xf spark-2.4.5-bin-hadoop2.7.tgz
# # !unzip CORD-19-research-challenge.zip
# -
sc.stop()
# +
# # !pip3 install -q findspark
# + colab={} colab_type="code" id="O-gFAAClZLFf"
import os
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["SPARK_HOME"] = "spark-2.4.5-bin-hadoop2.7"
# + colab={} colab_type="code" id="q-4Kh_kDm1qP"
import findspark
findspark.init()
# + colab={} colab_type="code" id="K4FA5doToNoI"
from pyspark.sql import SparkSession
from pyspark import SparkContext
sc = SparkContext("local", "first app")
spark = SparkSession \
.builder.appName("Covid-19").config("spark.driver.memory", "9g").getOrCreate() #.config("spark.driver.memory", "10g")
# + colab={} colab_type="code" id="SarzpFaLZUfg"
import os
import json
import pandas as pd
from pyspark import SQLContext
from pyspark.ml.feature import IDF, Tokenizer
from pyspark.ml import Pipeline
from pyspark.ml.feature import CountVectorizer
from pyspark.ml.feature import PCA
from pyspark.ml.clustering import KMeans
from pyspark.sql.types import IntegerType
from pyspark.sql.types import DateType
from pyspark.sql.functions import *
# + colab={} colab_type="code" id="mGdl-1vKokJ1"
from pyspark.sql.types import (
ArrayType,
IntegerType,
MapType,
StringType,
StructField,
StructType,
)
# -
# # Data analysis
metadata = spark.read\
.format("csv")\
.option("header", "true")\
.load('metadata.csv')
journalstat = metadata.groupby("journal").count().toPandas()
journalstat.describe()
# +
licensestatdf = metadata.groupby("license").count()
licensestat= licensestatdf.collect()
from matplotlib import pyplot
indexes = list(range(len(licensestat)))
values = [r['count'] for r in licensestat]
labels = [r['license'] for r in licensestat]
bar_width = 0.35
pyplot.bar(indexes, values)
#add labels
labelidx = [i + bar_width for i in indexes]
pyplot.xticks(labelidx, labels)
pyplot.show()
# -
licensestatdf.toPandas().head()
source_xstatdf = metadata.groupby("source_x").count()
source_xstatdf.toPandas().head()
# ## Load MetaData
# + colab={} colab_type="code" id="FKjioqRmc2dB"
metadata = spark.read\
.format("csv")\
.option("header", "true")\
.load('metadata.csv').select("journal","abstract","title","license","source_x" , "publish_time","sha","pdf_json_files")
# -
# # Define The Json Schema
# + colab={"base_uri": "https://localhost:8080/", "height": 238} colab_type="code" executionInfo={"elapsed": 1079, "status": "ok", "timestamp": 1589752957469, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSN3WEwfveBmcF37wOzIy1hDQbfY21c0kGBG_I=s64", "userId": "02390443002865386025"}, "user_tz": -120} id="UUnh0bH_om6Y" outputId="409c9148-9ce7-458b-81c5-291bcda10c60"
def generate_schema():
author_fields = [
StructField("first", StringType()),
StructField("middle", ArrayType(StringType())),
StructField("last", StringType()),
StructField("suffix", StringType()),
]
authors_schema = ArrayType(
StructType(
author_fields
+ [
StructField(
"affiliation",
StructType(
[
StructField("laboratory", StringType()),
StructField("institution", StringType()),
StructField(
"location",
StructType(
[
StructField("settlement", StringType()),
StructField("country", StringType()),
]
),
),
]
),
),
StructField("email", StringType()),
]
)
)
spans_schema = ArrayType(
StructType(
[
StructField("start", IntegerType()),
StructField("end", IntegerType()),
StructField("text", StringType()),
StructField("ref_id", StringType()),
]
)
)
section_schema = ArrayType(
StructType(
[
StructField("text", StringType()),
StructField("cite_spans", spans_schema),
StructField("ref_spans", spans_schema),
StructField("eq_spans", spans_schema),
StructField("section", StringType()),
]
)
)
bib_schema = MapType(
StringType(),
StructType(
[
StructField("ref_id", StringType()),
StructField("title", StringType()),
StructField("authors", ArrayType(StructType(author_fields))),
StructField("year", IntegerType()),
StructField("venue", StringType()),
StructField("volume", StringType()),
StructField("issn", StringType()),
StructField("pages", StringType()),
StructField(
"other_ids",
StructType([StructField("DOI", ArrayType(StringType()))]),
),
]
),
True,
)
ref_schema = MapType(
StringType(),
StructType(
[
StructField("text", StringType()),
StructField("latex", StringType()),
StructField("type", StringType()),
]
),
)
return StructType(
[
StructField("paper_id", StringType()),
StructField(
"metadata",
StructType(
[
StructField("title", StringType()),
StructField("authors", authors_schema),
]
),
True,
),
StructField("abstract", section_schema),
StructField("body_text", section_schema),
StructField("bib_entries", bib_schema),
StructField("ref_entries", ref_schema),
StructField("back_matter", section_schema),
]
)
def extract_dataframe(spark,path):
return spark.read.json(path, schema=generate_schema(), multiLine=True)
# -
def ascii_ignore(x):
return x.encode('ascii', 'ignore').decode('ascii')
ascii_udf = udf(ascii_ignore)
# + colab={} colab_type="code" id="Cvpl_r7Lp18I"
def LoadSelectedFeatures(path):
# load trainging
df = extract_dataframe(spark,path)
sqlContext = SQLContext(sc)
selectedFeatures = df.select("paper_id","metadata.title","body_text.text")
selectedFeatures.createOrReplaceTempView("parquetFile")
view = spark.sql("SELECT paper_id,title as paper_title,text as paper_text FROM parquetFile")
return view;
def LoadDataAndSaveParquet(path):
selectedFeatures = LoadSelectedFeatures(path)
# !rm -R "datajson.parquet"
saveParq(selectedFeatures,"datajson.parquet")
return selectedFeatures;
def saveParq(datfrm , path):
# # ! rm -R path
datfrm.repartition(5).write.save(path);
def LoadParq(path):
return spark.read.parquet(path);
# + colab={} colab_type="code" id="HxvaKTc6p5Ft"
def LoadDataFromParquet():
parquetFile = LoadParq("datajson.parquet")
parquetFile.createOrReplaceTempView("parquetFile")
tmpview = spark.sql("SELECT paper_id,title as paper_title,text as paper_text FROM parquetFile")
return tmpview;
# + colab={} colab_type="code" id="2DXy3c1td83r"
def prepare_dataset(metadata,tmpview):
tmpview=tmpview.withColumn('fullText', array_join(tmpview.paper_text, " ")).drop(tmpview.paper_text)
tmpview = tmpview.withColumn("fullText", ascii_udf('fullText'))
metadata = metadata.withColumn('publish_time', metadata['publish_time'].cast(DateType()))
dataset= metadata.join(tmpview , metadata.sha == tmpview.paper_id)
dataset = dataset.dropDuplicates()
dataset = dataset.fillna('NA', subset=['journal'])
dataset = dataset.withColumn('publish_time',when(col('publish_time').isNull(),to_date(lit('01.01.1900'),'dd.MM.yyyy')).otherwise(col('publish_time')))
dataset.withColumn('publish_year',year(dataset.publish_time))
# dataset = dataset.withColumn('pub_week_of_year',weekofyear(dataset.publish_time))
return dataset;
# +
import requests
stop_words = requests.get('http://ir.dcs.gla.ac.uk/resources/linguistic_utils/stop_words').text.split()
x="!()-[]{};:'"+"\,<>./?@#$%^&*_~"+'"'
custom_stop_words = [ 'doi', 'preprint', 'copyright', 'peer', 'reviewed', 'org',
'https', 'et', 'al', 'author', 'figure','rights', 'reserved', 'permission', 'used', 'using',
'biorxiv', 'medrxiv', 'license', 'fig', 'fig.', 'al.', 'Elsevier', 'PMC', 'CZI', 'www']
Punctuation=[char for char in x]
all_stop_words=stop_words+custom_stop_words+Punctuation
# +
from pyspark.ml.feature import StopWordsRemover
from pyspark.ml.feature import IDF
from pyspark.sql.functions import *
from pyspark.ml.feature import OneHotEncoderEstimator, StringIndexer, VectorAssembler
def prepareStages():
tokenizer = Tokenizer().setInputCol('fullText').setOutputCol('words')
sw_filter = StopWordsRemover()\
.setStopWords(all_stop_words)\
.setCaseSensitive(False)\
.setInputCol("words")\
.setOutputCol("filtered")
cv = CountVectorizer(minTF=2., minDF=5., vocabSize=3000)\
.setInputCol("filtered")\
.setOutputCol("tf")
idf = IDF().\
setInputCol('tf').\
setOutputCol('tfidf')
# create the string indexers for the categoricalColumns in our selelect feature list
categoricalColumns = ["journal","license","source_x"]
stages = [tokenizer, sw_filter,cv,idf]
for categoricalCol in categoricalColumns:
stringIndexer = StringIndexer(inputCol = categoricalCol, outputCol = categoricalCol + 'Index',handleInvalid='keep')
#stringIndexer = StringIndexer(inputCol = categoricalCol, outputCol = categoricalCol + 'Index')
encoder = OneHotEncoderEstimator(inputCols=[stringIndexer.getOutputCol()], outputCols=[categoricalCol + "classVec"])
stages += [stringIndexer, encoder]
#create the VectorAssembler for the numColumns in our selelect feature list
numericCols =["publish_year"] #
assemblerInputs = [c + "classVec" for c in categoricalColumns] + numericCols +["tfidf"]
assembler = VectorAssembler(inputCols=assemblerInputs, outputCol="features")
stages += [assembler]
return stages;
# + colab={} colab_type="code" id="i7YGd3PXp9qR"
tmpview = LoadDataAndSaveParquet("document_parses/pdf_json/0*")
# tmpview = LoadDataFromParquet()
# + colab={} colab_type="code" id="u765iW3QqCfm"
tmpview.count()
# + colab={} colab_type="code" id="Rbkz7RmDYlgn"
dataset = prepare_dataset(metadata,tmpview)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 10526, "status": "ok", "timestamp": 1589750583786, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSN3WEwfveBmcF37wOzIy1hDQbfY21c0kGBG_I=s64", "userId": "02390443002865386025"}, "user_tz": -120} id="URa0KmP9fk8d" outputId="024a88fa-246e-4408-ac66-1b631510e362"
dataset.count()
# -
dataset.show(1)
# + colab={} colab_type="code" id="UkAoJz7eabvU"
# from pyspark.sql.functions import udf
# def ascii_ignore(x):
# return x.encode('ascii', 'ignore').decode('ascii')
# ascii_udf = udf(ascii_ignore)
# dataset = dataset.withColumn("paper_title", ascii_udf('paper_title')).withColumn("fullText", ascii_udf('fullText'))\
# .withColumn("abstract", ascii_udf('abstract')).withColumn("title", ascii_udf('title')).show()
# + colab={} colab_type="code" id="TCzNQUd6TZw4"
# training_df, validation_df, testing_df = dataset.randomSplit([0.6, 0.3, 0.1], seed=0)
# + colab={} colab_type="code" id="i3EU2G0mNNIY"
pipeline_estimator = Pipeline(stages=prepareStages())
# + colab={} colab_type="code" id="-nWmOmuR8Ola"
pipelineModel = pipeline_estimator.fit(dataset)
df = pipelineModel.transform(dataset)
# -
df.printSchema()
rf = df.select('filtered').take(1)
# + colab={} colab_type="code" id="-NpewYSq2Ydn"
df.count()
# -
# # Features reduction
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 6031, "status": "error", "timestamp": 1589752601985, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSN3WEwfveBmcF37wOzIy1hDQbfY21c0kGBG_I=s64", "userId": "02390443002865386025"}, "user_tz": -120} id="aMZ4ZDKLehXJ" outputId="7972f6f0-44b0-4e15-9abc-0adab8aee412"
pca = PCA(k=100, inputCol="features", outputCol="pcaFeatures")
pcaModel = pca.fit(df);
result=pcaModel.transform(df).select("pcaFeatures","title")
df = result.withColumnRenamed("pcaFeatures","features")
# result.show(truncate=False)
# -
# !rm -R "transformed_df.parquet"
saveParq(df,"transformed_df.parquet")
# + colab={} colab_type="code" id="hwp4-ZtC3fHt"
df = LoadParq("transformed_df.parquet")
# -
# # Kmeans K selections
# + colab={} colab_type="code" id="VFHnU6F208io"
import numpy as np
cost = {}
for k in range(2,400,5):
kmeans = KMeans().setK(k).setSeed(3).setFeaturesCol('features')
model = kmeans.fit(df)
cost[k] = model.computeCost(df)
# -
cost.keys()
# Plot the cost
df_cost = pd.DataFrame(cost.values())
df_cost.columns = ["cost"]
new_col = cost.keys()
df_cost.insert(0, 'cluster', new_col)
df_cost.to_csv("kselectionResult.csv")
kselectionResult= pd.read_csv("kselectionResult.csv")
# +
import pylab as pl
import matplotlib.pyplot as plt
plt.plot(kselectionResult.cluster, kselectionResult.cost)
plt.xlabel('Number of Clusters')
plt.ylabel('Score')
plt.title('Elbow Curve')
print(plt.show())
# -
kselectionResult= pd.read_csv("kselectionResult.csv")
# # using the selected k
kmeans = KMeans().setK(42).setSeed(3).setFeaturesCol('features')
model = kmeans.fit(df)
# + colab={} colab_type="code" id="_H16-bW9nGZK"
cluster = model.transform(df)
# + colab={"base_uri": "https://localhost:8080/", "height": 151} colab_type="code" executionInfo={"elapsed": 16061, "status": "ok", "timestamp": 1589752277966, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSN3WEwfveBmcF37wOzIy1hDQbfY21c0kGBG_I=s64", "userId": "02390443002865386025"}, "user_tz": -120} id="CRoxSzpJatVx" outputId="f9857145-08d6-4c44-a34d-16921057b24a"
cluster.groupby("prediction").count().show(42)
# -
cluster.printSchema()
# + colab={} colab_type="code" id="MPa7x5Nb7Mte"
from pyspark.sql.functions import broadcast
from pyspark.mllib.feature import Normalizer
from pyspark.sql.functions import udf
import random
def cosine_similarityfn(X,Y):
# return ;
# return #random.randint(0, 100);
denom = float(X.norm(2) * Y.norm(2))
if denom == 0.0:
return -1.0
else:
return float(X.dot(Y)) / float(denom)
cosine_similarity = udf(cosine_similarityfn)
def recommendPaper(paper,N):
targetdf = metadata.where(metadata.title == paper)
# load the json file for this paper
testdf = LoadSelectedFeatures(targetdf.select('pdf_json_files').take(1)[0].pdf_json_files)
testdataset= prepare_dataset(metadata,testdf)
pipresult = pipelineModel.transform(testdataset);
# pipresult.printSchema();
reducedfeature = pcaModel.transform(pipresult).select("title","pcaFeatures").withColumnRenamed("pcaFeatures","features")
reducedfeature.printSchema()
result = model.transform(reducedfeature)
dfclass = result.select('prediction').take(1)[0].prediction
PapersFromtheSameClass = cluster.where(cluster.prediction == dfclass)
result = result.withColumnRenamed('features','features2')
dfOfResult = result.select("features2")
PapersFromtheSameClass_withCosine = PapersFromtheSameClass.select('title','features').crossJoin(dfOfResult.hint("broadcast")) ;
# PapersFromtheSameClass_withCosine.count()
PapersFromtheSameClass_withCosine = PapersFromtheSameClass_withCosine.withColumn(
'COSINE_SIM', cosine_similarity("features",'features2')
).orderBy(col('COSINE_SIM').desc());
#
# PapersFromtheSameClass_withCosine.printSchema()
return PapersFromtheSameClass_withCosine.take(N)
# + colab_type="text" id="jyhMEwbkswnM"
r = recommendPaper('Sequence requirements for RNA strand transfer during nidovirus discontinuous subgenomic RNA synthesis',5)
# -
counter = 1
for i in r :
print(str(counter) + "\t" + str(i.COSINE_SIM)[:4]+ "\t"+i.title)
counter = counter+1
| COVID-19.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data description & Problem statement:
# This data set is collected from recordings of 30 human subjects captured via smartphones enabled with embedded inertial sensors.
# In this project, we calculate a model by which a smartphone can detect its owner’s activity precisely. For the dataset, 30 people were used to perform 6 different activities. Each of them was wearing a Samsung Galaxy SII on their waist. Using the smartphone’s embedded sensors (the accelerometer and the gyroscope), the user’s speed and acceleration were measured in 3-axial directions. We use the sensor’s data to predict user’s activity.
#
# * Dataset is imbalanced. The data has 10299 rows and 561 columns.
# * This is a Multiclass-classification problem.
#
# # Workflow:
# - Load the dataset, and define the required functions (e.g. for detecting the outliers)
# - Data Cleaning/Wrangling: Manipulate outliers, missing data or duplicate values, Encode categorical variables, etc.
# - Split data into training & test parts (utilize the training part for training & hyperparameter tuning of model, and test part for the final evaluation of model)
# # Model Training:
# - Build an Random Forest model, and evaluate it via C-V approach
#
# # Model Evaluation:
# - Evaluate the RF model with optimized hyperparameters on Test Dataset, by calculating:
# - f1_weighted
# - Confusion matrix
# +
import sklearn
import tensorflow as tf
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn import preprocessing
# %matplotlib inline
from scipy import stats
import warnings
warnings.filterwarnings("ignore")
# +
df_X=pd.read_csv('C:/Users/rhash/Documents/Datasets/Human activity recognition/UCI HAR Dataset/train/X_train.txt'
,delim_whitespace=True, header = None)
df_y=pd.read_csv('C:/Users/rhash/Documents/Datasets/Human activity recognition/UCI HAR Dataset/train/y_train.txt'
,delim_whitespace=True, header = None, names=['Label'])
df = pd.concat([df_y, df_X], axis=1)
# To Shuffle the data:
np.random.seed(42)
df=df.reindex(np.random.permutation(df.index))
df.reset_index(inplace=True, drop=True)
df.head(3)
# -
X=df.drop('Label', axis=1)
y=df['Label']
# We initially devide data into training & test folds:
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=42, stratify = y)
# +
# Build and fit the RF model:
from sklearn.ensemble import RandomForestClassifier
RF = RandomForestClassifier(max_features=560, n_estimators=100, max_depth=30, random_state=42, class_weight='balanced')
model = RF.fit(X_train, y_train)
# +
from sklearn.metrics import roc_curve, auc, confusion_matrix, classification_report
# Plot a confusion matrix.
# cm is the confusion matrix, names are the names of the classes.
def plot_confusion_matrix(cm, names, title='Confusion matrix', cmap=plt.cm.Blues):
fig, ax = plt.subplots(1,1,figsize=(8,8))
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(names))
plt.xticks(tick_marks, names, rotation=90)
plt.yticks(tick_marks, names)
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.show()
names = ["WALKING", "WALKING_UPSTAIRS", "WALKING_DOWNSTAIRS", "SITTING", "STANDING", "LAYING"]
# Compute confusion matrix
cm = confusion_matrix(y_test, model.predict(X_test))
np.set_printoptions(precision=2)
print('Confusion matrix, without normalization')
print(cm)
# Normalize the confusion matrix by row (i.e by the number of samples in each class)
cm_normalized = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print('Normalized confusion matrix')
print(cm_normalized)
plot_confusion_matrix(cm_normalized, names, title='Normalized confusion matrix')
# -
# Classification report:
report=classification_report(y_test, model.predict(X_test))
print(report)
| Projects in Python with Scikit-Learn- XGBoost- Pandas- Statsmodels- etc./Human Activity Recognition (Random Forest).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.1 64-bit
# metadata:
# interpreter:
# hash: dcacb0086e9a4f4eabd41c33bf4faac5ea0a3337ed3f5eff0680afa930572c04
# name: python3
# ---
# # Learning Objectives
#
# - [ ] PLACEHOLDER
# # 6 A Python REPL and Modules
#
# A **read–eval–print loop (REPL)**, also termed a language shell, is a simple interactive computer programming environment that
#
# 1. takes single user inputs,
# 2. executes them,
# 3. returns the result to the user,
# 4. loop back to 1.
#
# ## 6.1 IDLE, Python's REPL
#
# IDLE is Python’s **Integrated Development and Learning Environment**. It is installed with Python installation.
#
# #### Exercise
#
# 1. In IDLE, print `Hello World!`.
# 2. IDLE has autocompletion feature, accessible by using `tab` key. Try it out with some functions that you know, e.g. `print`. Also to defined a variable with a lengthy name and use the autocompletion feature, e.g. `this_is_a_long_variable_name=20`.
# 3. Try to use `help` function to find out information on the `print` function.
# 4. To copy a line, use `up` arrow key to move to the previous command and press enter. Try it out by assigning `50` to `this_is_a_long_variable_name`.
# 5. You can iterate through history commands on REPL. Following are shortcuts to move to previous/next history command.
# - Press `ALT + P` keys to get last history command.
# - Press `ALT + N` keys to see next history command.
# - To exit from history, press `CTRL + C` keys
# ## 6.2 Python Modules
#
# REPL is good for quick evaluation of single statement. But for multilines of code, it is common and much better off using a text editor to prepare the input and running it with that file as input. This is known as **scripting**. This input file is termed a **module**.
#
# Formally, a Python **module** is a file containing Python definitions and statements. The file name is the module name with the suffix `.py` appended.
#
# ### 6.2.1 Creating and Running Modules on IDLE
#
# On IDLE, go to menu `File > New Window` (or press `Ctrl + N` ) to start a new code window and type your code in the new window.
#
# #### Exercise
#
# In the module, enter the code `print('Hello World!')` and run the module in by either using `Run > Run Module` or pressing `F5`. Note that you will be prompted to save your file first before you can execute the code. Remember your file name for the next step.
#
# You can open the module by using `File > Open` or even `File > Open Module...` menu.
#
# #### Exercise
# On IDLE, create and save a module called `math_fun` that contains functions with the following names:
#
# 1. `cube` that takes in a float and return the cube of the float, e.g.
# >```
# >>>>cube(5)
# >125.0
# >```
# 2. `div_algo` that takes in 2 integers parameters that it returns a string that represents the larger integer in terms of the product of the lower integer with the quotient added with the remaindere, e.g.,
# >```
# >>>>div_algo(5,14)
# >14=2*5+4
# >>>>div_algo(13,6)
# >13=2*6+1
# >```
# 3. `lcm` that takes in 2 integer parameters and returns an integer that is the lowest common multiple of the two supplied integers. You might want to use the fact that for any integers $a$ and $b$, $$\gcd(a,b)\text{lcm}(a,b)=ab.$$
# >```
# >>>>lcm(15,10)
# >30
# >```
# ### 6.2.2 Importing Function from Other Modules
#
# Functions defined in another module can be accessed by using the `import` keyword with the following syntax:
# >```python
# >import MODULE_NAME_HERE
# >```
#
# The functions in the imported module can be accessed as a method of the module object, i.e. the syntax is
# >```python
# >MODULE_NAME_HERE.FUNCTION_NAME_IN_IMPORTED_MODULE(SOME_PARAMETERS)
# >```
#
# #### Example
# `math` is a standard Python module that comes withe some useful mathematical functions, e.g., `sqrt` is a function in the module `math` that takes in a float and return the positive square root of the inputted float. To find the value of $\sqrt{5}$, we use the following code in our main module. If we intend to use the function in the other module often, we can assign it to a local name.
# +
import math
print(math.sqrt(5))
sqrt=math.sqrt ##Assigning to a local name.
print(sqrt(5))
# -
# `from` statement is a variant of the `import` statement that imports names from a module directly into the importing module’s symbol table.
#
# #### Example
# +
from math import sqrt
print(sqrt(5))
# -
# #### Exercise
#
# On IDLE,
#
# 1. create a new module named `my_main`,
# 2. import the module `math_fun` you created earlier inside `my_main` by using `import` statement.
# 3. Try out the various functions that you have defined earlier.
# 4. Repeat 2 and 3, but now using the `from` statement.
#
# +
#YOUR CODE HERE
# -
# ## 6.3 Some Useful Modules
#
# - `csv`
# - `datetime`
# - `math`
# - `matplotlib`
# ### 6.3.4 Basic `matplotlib`
#
# If you're not installing through Anaconda, you might not have `matplotlib` package. Thus, to install, you can run `pip install matplotlib` through command prompt.
#
# After importing `matplotlib.pyplot`, we can use the `plot()` method to plot the list `y` against the list `x` on a cartesian plane. Syntax is
#
# >```python
# >matplotlib.pyplot.plot(x,y,point_options)
# >```
#
# The argument `point_options` has a default `b-` where `b` indicates the color blue and `-` means that the points will be connected with a straight line. You can also try with `rs,g--,b^` to see the different colors and styles. Please visit [https://matplotlib.org/stable/api/_as_gen/matplotlib.pyplot.plot.html#matplotlib.pyplot.plot] for the full documentation.
#
# The labels of the $x$ and $y$ axis can be adjusted via the self-explanatory `xlabel()` and `ylabel` methods. Syntax is
# >```python
# >matplotlib.pyplot.xlabel(your_xlabel_here)
# >matplotlib.pyplot.xlabel(your_ylabel_here)
# >```
#
# To display your plot, use the method `show()` on the object `matplotlib.pyplot.plot`
import matplotlib.pyplot
matplotlib.pyplot.plot([1, 2, 3, 4])
matplotlib.pyplot.ylabel('some numbers')
matplotlib.pyplot.show()
| Notes/Appendix_Python_Modules.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
df = pd.read_csv("./data/fashion/fashion-mnist_train.zip")
df.head()
X = df.iloc[:,1:]
pca=PCA(n_components=0.99)
pca.fit(X)
X_pca = pca.transform(X)
X_pca.shape
X.shape
pca.explained_variance_ratio_ # List of variance explained by each component
np.sum(pca.explained_variance_ratio_)
X_pca[0].shape
plt.imshow(pca.inverse_transform(X_pca[0]).reshape(28,28), cmap='gray')
plt.imshow(X.values[0].reshape(28,28), cmap='gray')
# ### Possible uses of PCA
#
# - Compression (reconstruction is quite good).
# - Use `X_pca` instead of `X` for training a model (either supervised / unsupervised learning task).
# - Visualiza the data better.
from matplotlib import colors
labels = [1,5]# sorted(df['label'].unique())
colors = ['black','red'] #list(colors.CSS4_COLORS)[0:len(labels)]
for i, color in enumerate(colors): #get index and value on the list (iterable)
x1 = X_pca[:,0][df['label'] == i]
x2 = X_pca[:,1][df['label'] == i]
plt.scatter(x1,x2,c=color)
plt.xlabel("First component")
plt.ylabel("Second component")
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(111, projection='3d')
for i, color in enumerate(colors): #get index and value on the list (iterable)
x1 = X_pca[:,0][df['label'] == i]
x2 = X_pca[:,1][df['label'] == i]
x3 = X_pca[:,2][df['label'] == i]
ax.scatter(x1,x2,c=color)
| 10 PCA - Dimensionality Reduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Model the Target as Binary Variable - Classification Problem
# #### Load Libraries
import pandas as pd
pd.options.display.max_columns = 200
pd.options.display.max_rows = 1000
from sklearn import preprocessing
import numpy as np
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline, make_union
from tpot.builtins import StackingEstimator
from tpot import TPOTClassifier
from sklearn.datasets import load_iris
from sklearn.metrics import roc_auc_score
from scipy import interp
from itertools import cycle
from sklearn import svm, datasets
from sklearn.metrics import roc_curve, auc
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import average_precision_score
from matplotlib import pyplot
from pandas import read_csv
from pandas import set_option
from pandas.tools.plotting import scatter_matrix
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.pipeline import Pipeline
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
from sklearn.ensemble import AdaBoostClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import ExtraTreesClassifier
import matplotlib.pyplot as plt
import seaborn as sns
from __future__ import print_function
import os
import subprocess
from sklearn.tree import DecisionTreeClassifier, export_graphviz
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
# #### Load the surrounding and sales data and join them to create a data set for learning
# +
surrounding_pdf = pd.read_csv('../UseCase_3_Datasets/store_surrounding_pdf_exp.csv')
surrounding_nf_pdf = pd.read_csv('../UseCase_3_Datasets/new_features.csv')
surrounding_pdf = pd.merge(surrounding_nf_pdf,surrounding_pdf,on='store_code',how='inner')
surrounding_pdf = surrounding_pdf.drop_duplicates()
y_column = 'total_sales'
# store_sales = pd.read_csv('../UseCase_3_Datasets/sales_daily_simple_sum.csv')
# sales_pdf_daily_sub = store_sales[['store_code','total_sales']]
#store_sales = pd.read_csv('../UseCase_3_Datasets/sales_per_hr.csv')
#sales_pdf_daily_sub = store_sales[['store_code','sales_per_hr']]
store_sales = pd.read_csv('../UseCase_3_Datasets/sales_per_day_period.csv')
sales_pdf_daily_sub = store_sales[['store_code','total_sales','Morning','Afternoon','Evening','sales_pattern']]
# sales_pdf_daily_sub = sales_pdf_daily_sub[(sales_pdf_daily_sub['sales_pattern'] == 'Evening') |
# (sales_pdf_daily_sub['sales_pattern'] == 'Afternoon')]
sales_pdf_daily_sub = sales_pdf_daily_sub[['store_code',y_column]]
# +
# # #Read other population realted information from wiki
# pop_features_pdf = pd.read_csv('../UseCase_3_Datasets/swiss_pop_features_2.csv',sep=';',header=0)
# pop_features_pdf.head()
# pop_features_pdf.columns
# pop_pdf = pd.read_csv('../UseCase_3_Datasets/swiss_pop.csv',sep=',')
# pop_pdf.head()
# # #Comnine with surrounding data using city code informartion
# surrounding_pdf = pd.merge(pop_features_pdf,surrounding_pdf,on='city',how='inner')
# surrounding_pdf = surrounding_pdf.drop_duplicates()
# surrounding_pdf = pd.merge(pop_pdf,surrounding_pdf,on='city',how='inner')
# surrounding_pdf = surrounding_pdf.drop_duplicates()
# surrounding_pdf['pop_above_20'] = 100-surrounding_pdf['pop_under_20']
# surrounding_pdf= surrounding_pdf.drop('cant',axis=1)
# surrounding_pdf.head()
# -
print("Surrounding Shape",surrounding_pdf.shape)
print("Sales Shape",sales_pdf_daily_sub.shape)
# ### Data Cleaning
# #### Remove variable that have ony one unique value
#Join Store and Surroounding datasets
sales_surrounding_pdf = pd.merge(sales_pdf_daily_sub,surrounding_pdf,on='store_code',how='inner')
sales_surrounding_pdf = sales_surrounding_pdf.drop_duplicates()
# Dropping columns that do not provide useful information for this analysis
for i in sales_surrounding_pdf.columns:
if sales_surrounding_pdf[i].nunique() == 1:
sales_surrounding_pdf.drop(i, axis = 1, inplace = True)
print("Sales Surrounding Shape",sales_surrounding_pdf.shape)
# +
# #Visualization of individual attributes
# #Histogram
# sales_surrounding_pdf.hist(sharex=False,sharey=False,xlabelsize=0.25,ylabelsize=0.25,figsize=(20,20))
# pyplot.show()
# +
# sales_surrounding_pdf.plot(kind='density', subplots=True, layout=(10,18), sharex=False, legend=False,fontsize=1,
# figsize=(20,20))
# pyplot.show()
# +
# fig = pyplot.figure(figsize=(15,15))
# ax = fig.add_subplot(111)
# cax = ax.matshow(sales_surrounding_pdf.corr(),vmin=-1, vmax=1)
# fig.colorbar(cax)
# pyplot.show()
# -
# #### One hot Encoding for categorical values
# +
import math
sales_surrounding_pdf_ohe = pd.get_dummies(sales_surrounding_pdf)
#Create Train,Test without imputation
#X = sales_surrounding_pdf.loc[:, sales_surrounding_pdf.columns != 'total_sales']
#X = sales_surrounding_pdf.loc[:, sales_surrounding_pdf.columns != 'store_code']
y = sales_surrounding_pdf_ohe[y_column].apply(lambda x : math.log(x+1))
X = sales_surrounding_pdf_ohe.drop([y_column,'store_code'],axis=1)
#X = sales_surrounding_pdf.drop('store_code',axis=1)
# print("Shape ",X.shape)
# print ("Shape ",y.shape )
# -
y.plot(kind='hist',subplots=True,fontsize=10,figsize=(10,10),grid=True,)
# y_test.hist(bins=20,figsize=(10,10))
pyplot.axvline(x=9.5,color='red')
pyplot.xlabel("Store Sales")
pyplot.show()
# #### Creation of Test and Train set
#Divide into train and test
validation_size = 0.20
seed = 33
X_train, X_test, y_train, y_test = train_test_split(X, y,test_size=validation_size,random_state=seed)
# #### Target Creation
# Since we plan to make a binary target, we try to find a split point, looking at the data
# Also, the split point is created in such a way, that we do dont create a unbalanced dataset to learn and predict.
# xtick_list = range(0,500000,50000)
# ytick_list = range(0,y_train.shape[0],30)
# y_train.plot(kind='hist',subplots=True,fontsize=10,figsize=(10,10),xticks=xtick_list,yticks=ytick_list)
y_train.hist(bins=30,figsize=(10,10))
# pyplot.xlabel("Store Sales")
# pyplot.show()
# +
# y_train[y_train.values < 8].hist(bins=30,figsize=(10,10),)
# pyplot.xlabel("Store Sales")
# pyplot.show()
# -
# xtick_list = range(0,500000,50000)
# ytick_list = range(0,y_test.shape[0],10)
#y_test.plot(kind='hist',subplots=True,fontsize=10,figsize=(10,10),grid=True,xticks=xtick_list,yticks=ytick_list)
y_test.hist(bins=20,figsize=(10,10))
pyplot.xlabel("Store Sales")
pyplot.show()
# +
# #Find the most balanced point of the data set :
# for i in np.arange(7,10,0.15):
# y_train_sp = y_train.apply(lambda x : 1 if x >=i else 0 )
# y_test_sp = y_test.apply(lambda x : 1 if x >=i else 0)
# print ("Split Point ", i)
# print ("% of 0-class in Test ",y_test_sp.value_counts()[0]/ (y_test_sp.value_counts()[1]+y_test_sp.value_counts()[0])*100)
# print ("% of 0-class in Train ",y_train_sp.value_counts()[0]/ (y_train_sp.value_counts()[1]+y_train_sp.value_counts()[0])*100)
# print("\n")
# +
#Based on the above histogram comparison threshold 50,000 is a good enough to make binary classification target
split_point = 8.95
y_train = y_train.apply(lambda x : 1 if x >=split_point else 0 )
y_test = y_test.apply(lambda x : 1 if x >=split_point else 0)
y = y.apply(lambda x : 1 if x >=split_point else 0)
y_test.value_counts()
#Binary Targets created.
# -
y_test.value_counts()[0] / (y_test.value_counts()[1]+y_test.value_counts()[0])*100
y_train.value_counts()
y_train.value_counts()[0] / (y_train.value_counts()[1]+y_train.value_counts()[0])*100
type(X_train)
# On the spot checks,try out some basic models and see, how they perform
# +
#Find Feature Importance via various methods
#1)Logistic Regression
#1.1) Logistic Regression with L1 Penalty (Lasso)
#2)Decision Tree
# Spot-Check Algorithms
models = []
num_folds = 10
seed = 42
scoring ='roc_auc'
models.append(('LR', LogisticRegression()))
models.append(('LR with L1 penalty', LogisticRegression(penalty='l1')))
models.append(('CART', DecisionTreeClassifier()))
models.append(('LDA', LinearDiscriminantAnalysis()))
models.append(('KNN', KNeighborsClassifier()))
results = []
names = []
for name, model in models:
kfold = KFold(n_splits=num_folds, random_state=seed)
cv_results = cross_val_score(model, X_train, y_train, cv=kfold, scoring=scoring)
results.append(cv_results)
names.append(name)
msg = "%s: %f (%f)" % (name,cv_results.mean(),cv_results.std())
print(msg)
# -
# Compare Algorithms
fig = pyplot.figure(figsize=(10,10))
fig.suptitle('Algorithm Comparison')
ax = fig.add_subplot(111)
pyplot.boxplot(results)
ax.set_xticklabels(names)
pyplot.show()
pipelines = []
pipelines.append(('Scaled LR', Pipeline([('Scaler', StandardScaler()),('LR',LogisticRegression())])))
pipelines.append(('Scaled LR with L1', Pipeline([('Scaler', StandardScaler()),('LR with L1',LogisticRegression(penalty='l1'))])))
pipelines.append(('Scaled CART', Pipeline([('Scaler', StandardScaler()),('CART',DecisionTreeClassifier(max_depth=5))])))
pipelines.append(('ScaledKNN', Pipeline([('Scaler', StandardScaler()),('KNN',KNeighborsClassifier())])))
results = []
names = []
for name, model in pipelines:
kfold = KFold(n_splits=num_folds, random_state=seed)
cv_results = cross_val_score(model, X_train, y_train, cv=kfold, scoring=scoring)
results.append(cv_results)
names.append(name)
msg = "%s: %f (%f)" % (name, cv_results.mean(), cv_results.std())
print(msg)
fig = pyplot.figure(figsize=(10,10))
fig.suptitle('Scaled Algorithm Comparison')
ax = fig.add_subplot(111)
pyplot.boxplot(results)
ax.set_xticklabels(names)
pyplot.show()
# Try, some ensemble model, and see their performance
# +
# ensembles
ensembles = []
ensembles.append(('RF', RandomForestClassifier()))
ensembles.append(('ET', ExtraTreesClassifier()))
results = []
names = []
for name, model in ensembles:
kfold = KFold(n_splits=num_folds, random_state=seed)
cv_results = cross_val_score(model, X_train, y_train, cv=kfold, scoring=scoring)
results.append(cv_results)
names.append(name)
msg = "%s: %f (%f)" % (name, cv_results.mean(), cv_results.std())
print(msg)
# -
# Compare Algorithms
fig = pyplot.figure(figsize=(10,10))
fig.suptitle('Ensemble Algorithm Comparison')
ax = fig.add_subplot(111)
pyplot.boxplot(results)
ax.set_xticklabels(names)
pyplot.show()
# +
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
# plt.xticks.set_color('black')
# plt.yticks.set_color('black')
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="red" if cm[i, j] > thresh else "black")
plt.tight_layout()
#ax.xaxis.label.set_color('red')
#plt.xlabel.set_color('black')
plt.ylabel('True label')
plt.xlabel('Predicted label')
def visualize_tree(tree,feature_names,file_name):
"""Create tree png using graphviz.
Args
----
tree -- scikit-learn DecsisionTree.
feature_names -- list of feature names.
"""
with open(file_name, 'w') as f:
export_graphviz(tree, out_file=f,feature_names=feature_names)
command = ["dot", "-Tpng", "dt.dot", "-o", "dt.png"]
try:
subprocess.check_call(command)
except:
exit("Could not run dot, ie graphviz, to produce visualization")
# -
# Since our goal is interpret-ability of our features, we will stick to models like Logistic regression, Decision Tree. Ensemble methods too can be used, since they provide a feature importance indication .
#
# In the below cells, we try out various algorithms. Use grid search to best tune the hyper parameters and using cross - validation, to ensure we do not over fit the training data.
# +
from sklearn.grid_search import GridSearchCV
from sklearn.datasets import make_classification
from sklearn.ensemble import RandomForestClassifier
seed = 13
dtcf = DecisionTreeClassifier(max_depth=1,random_state=seed)
# n_estimator = list(range(10,500,10))
#print(dtcf.get_params)
param_grid = {
'max_depth': list(range(1,10,3)),
'max_features': ['auto', 'sqrt', 'log2'],
'min_samples_split' : list(range(2,20,1)),
'criterion':['entropy','gini']
}
CV_dtcf = GridSearchCV(estimator=dtcf,param_grid=param_grid, cv= 5,scoring='roc_auc')
CV_dtcf.fit(X_train, y_train)
print (CV_dtcf.best_params_)
print (CV_dtcf.best_score_)
predictions_train = CV_dtcf.best_estimator_.predict_proba(X_train)[:,1]
print("Train ROC-AUC :",roc_auc_score(y_train, predictions_train))
predictions = CV_dtcf.best_estimator_.predict_proba(X_test)[:,1]
print("Test ROC_AUC :",roc_auc_score(y_test, predictions))
visualize_tree(CV_dtcf.best_estimator_, X_train.columns,"dt_cv.txt")
# -
seed=13
model = DecisionTreeClassifier(max_depth=3,criterion='entropy',random_state=seed)
model.fit(X_train, y_train)
#estimate on train data
predictions_train = model.predict_proba(X_train)[:,1]
print("Train ROC-AUC :",roc_auc_score(y_train, predictions_train))
predictions = model.predict_proba(X_test)[:,1]
print("Test ROC_AUC :",roc_auc_score(y_test, predictions))
visualize_tree(model, X_train.columns,"dt_entropy.txt")
# +
#Find Feature Importance with Random Forest Classifer and Extra Tress Classifier
# +
# from sklearn.grid_search import GridSearchCV
# from sklearn.datasets import make_classification
# from sklearn.ensemble import RandomForestClassifier
# rfc = RandomForestClassifier(n_jobs=-1,max_features= 'sqrt' ,n_estimators=50, oob_score = True)
# # n_estimator = list(range(10,500,10))
# # print(n_estimator)
# param_grid = {
# 'n_estimators': list(range(10,500,10)),
# 'max_features': ['auto', 'sqrt', 'log2'],
# 'max_depth' : list(range(3,9,1))
# }
# CV_rfc = GridSearchCV(estimator=rfc,param_grid=param_grid, cv= 5,scoring='roc_auc')
# CV_rfc.fit(X_train, y_train)
# print (CV_rfc.best_params_)
# -
# Build a forest and compute the feature importances
def visualizeFeatureImportance(model,feature_names,no_of_features):
forest = model
importances = forest.feature_importances_
std = np.std([tree.feature_importances_ for tree in forest.estimators_],axis=0)
indices = np.argsort(importances)[::-1]
features = no_of_features
# Print the feature ranking
print("Feature ranking:")
for f in range(features):
print("%d. feature %s (%f)" % (f + 1, feature_names[indices[f]], importances[indices[f]]))
# Plot the feature importances of the forest
plt.figure(figsize=(10,10))
plt.title("Feature importances")
plt.bar(range(features), importances[indices[:features]],color="r", yerr=std[indices[:features]], align="center")
plt.xticks(range(features), indices[:features])
plt.xlim([-1,features])
plt.show()
# +
seed = 19
rf_clf = RandomForestClassifier(n_estimators=5,max_depth=3,random_state=seed,max_features='sqrt')
rf_clf.fit(X_train,y_train)
predictions_train = rf_clf.predict(X_train)
print("Train ROC-AUC :",roc_auc_score(y_train, predictions_train))
# estimate accuracy on validation dataset
predictions = rf_clf.predict(X_test)
print("Test ROC-AUC :",roc_auc_score(y_test, predictions))
visualizeFeatureImportance(rf_clf,X_train.columns,10)
# +
# prepare the model
seed = 17
model_et = ExtraTreesClassifier(n_estimators=5,max_depth=3,bootstrap=True,max_features='sqrt',random_state=seed)
model_et.fit(X_train, y_train)
predictions_train = model_et.predict_proba(X_train)[:,1]
print("Train AUC :",roc_auc_score(y_train,predictions_train))
predictions = model_et.predict_proba(X_test)[:,1]
print("Test AUC :",roc_auc_score(y_test,predictions))
visualizeFeatureImportance(model_et,X_train.columns,10)
# +
from sklearn.grid_search import GridSearchCV,RandomizedSearchCV
from sklearn.datasets import make_classification
from sklearn.ensemble import RandomForestClassifier
from sklearn.preprocessing import Normalizer
validation_size = 0.30
seed = 13
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=validation_size,random_state=seed)
scaler = Normalizer().fit(X_train)
normalizedX = scaler.transform(X_train)
lr = LogisticRegression(C=1,n_jobs=10,penalty='l1')
param_grid = { 'C': np.arange(1,100,1)}
CV_lr = RandomizedSearchCV(estimator=lr,param_distributions=param_grid, cv=10,scoring='roc_auc',random_state=seed)
CV_lr.fit(normalizedX, y_train)
print (CV_lr.best_params_)
# +
lr = CV_lr.best_estimator_
lr.fit(normalizedX, y_train)
predictions_train = lr.predict_proba(normalizedX)[:,1]
print("Train AUC :",roc_auc_score(y_train,predictions_train))
normalizedX_test = scaler.transform(X_test)
predictions = lr.predict_proba(normalizedX_test)[:,1]
print("Test ROC_AUC :",roc_auc_score(y_test, predictions))
# +
# from tpot import TPOTClassifier
# # prepare the model
# model = TPOTClassifier(generations=5, population_size=50, verbosity=2,cv=10,scoring='roc_auc')
# model.fit(X_train, y_train)
# predictions = model.predict(X_test)
# print(roc_auc_score(y_test, predictions))
# print(confusion_matrix(y_test, predictions))
# -
# Sci-kit learn provide facilities from feature selection to improve estimators accuracy scores or to boost their performance on very high-dimensional datasets.
#
# We compute the ANOVA F-value for the data, using the SelectKBest methods and use these as inputs to various algorithms.
# +
from sklearn.datasets import load_iris
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import chi2, f_classif, mutual_info_classif
import pandas as pd
selector = SelectKBest(chi2, k=15)
selector.fit(X_train, y_train)
X_new = selector.transform(X)
X_new.shape
X.columns[selector.get_support(indices=True)]
chi2_list = X.columns[selector.get_support(indices=True)].tolist()
print(chi2_list,"\n")
selector = SelectKBest(f_classif, k=15)
selector.fit(X, y)
X_new = selector.transform(X)
X_new.shape
X.columns[selector.get_support(indices=True)]
f_classif_list = X.columns[selector.get_support(indices=True)].tolist()
print(f_classif_list,"\n")
selector = SelectKBest(mutual_info_classif, k=15)
selector.fit(X, y)
X_new = selector.transform(X)
X_new.shape
X.columns[selector.get_support(indices=True)]
mic_list = X.columns[selector.get_support(indices=True)].tolist()
print(mic_list,"\n")
# +
#First Level of segregation using f_classif_list
X_sub = X[X.columns.intersection(f_classif_list)]
y_sub = y
print("Shape ",X_sub.shape)
print ("Shape ",y.shape )
#Divide
validation_size = 0.30
seed = 19
X_train, X_test, y_train, y_test = train_test_split(X_sub,y_sub,test_size=validation_size,random_state=seed)
dtcf = DecisionTreeClassifier(max_depth=2,random_state=seed)
param_grid = {
'max_depth': list(range(2,10,1)),
'max_features': ['auto', 'sqrt', 'log2'],
'min_samples_split' : list(range(2,20,1)),
'criterion':['entropy','gini']
}
CV_dtcf = RandomizedSearchCV(estimator=dtcf,param_distributions=param_grid, cv= 5,scoring='roc_auc',
random_state=seed)
CV_dtcf.fit(X_train, y_train)
print (CV_dtcf.best_params_)
#print (CV_dtcf.best_score_)
predictions_train = CV_dtcf.best_estimator_.fit(X_train, y_train).predict_proba(X_train)[:,1]
print("Train ROC-AUC :",roc_auc_score(y_train, predictions_train))
predictions = CV_dtcf.best_estimator_.predict_proba(X_test)[:,1]
print("Test ROC-AUC :",roc_auc_score(y_test, predictions))
visualize_tree(CV_dtcf.best_estimator_,X_train.columns,"dt_f_classif_sub.txt")
# +
#Overall data set
validation_size = 0.30
seed = 17
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=validation_size,random_state=seed)
dtcf = DecisionTreeClassifier(max_depth=2,random_state=seed)
param_grid = {
'max_depth': list(range(2,10,1)),
'max_features': ['auto', 'sqrt', 'log2'],
'min_samples_split' : list(range(3,10,1)),
'criterion':['entropy','gini']
}
CV_dtcf = RandomizedSearchCV(estimator=dtcf,param_distributions=param_grid, cv= 5,scoring='roc_auc',random_state=seed)
CV_dtcf.fit(X_train, y_train)
print (CV_dtcf.best_params_)
#print (CV_dtcf.best_score_)
predictions_train = CV_dtcf.best_estimator_.fit(X_train, y_train).predict_proba(X_train)[:,1]
print("Train ROC-AUC :",roc_auc_score(y_train, predictions_train))
# estimate accuracy on validation dataset
predictions = CV_dtcf.best_estimator_.predict_proba(X_test)[:,1]
print("Test ROC-AUC :" , roc_auc_score(y_test, predictions))
visualize_tree(CV_dtcf.best_estimator_,X_train.columns,"dt_all_data.txt")
# +
# {criterion': 'gini', 'max_depth': 4, 'max_features': 'auto', 'min_samples_split': 15}
# {'criterion': 'gini', 'max_depth': 8, 'max_features': 'log2', 'min_samples_split': 4}
# {'criterion': 'gini', 'max_depth': 4, 'max_features': 'auto', 'min_samples_split': 11}
# +
#ensemble_input_col = list(set().union(chi2_list,f_classif_list,mic_list))
ensemble_input_col = list(set().union(f_classif_list))
X_sub = X[X.columns.intersection(ensemble_input_col)]
y_sub = y
print("Shape ",X_sub.shape)
print ("Shape ",y.shape )
#Divide
validation_size = 0.30
seed = 29
X_train, X_test, y_train, y_test = train_test_split(X_sub,y_sub,test_size=validation_size,random_state=seed)
from sklearn.grid_search import GridSearchCV, RandomizedSearchCV
from sklearn.datasets import make_classification
from sklearn.ensemble import RandomForestClassifier
from scipy.stats import randint as sp_randint
rfc = RandomForestClassifier(n_jobs=-1)
# n_estimator = list(range(10,500,10))
# print(n_estimator)
param_grid = {
'n_estimators': list(range(10,500,10)),
'max_features': ['auto', 'sqrt', 'log2'],
'max_depth' : list(range(3,9,1)),
"min_samples_split": sp_randint(2, 11),
"min_samples_leaf": sp_randint(2, 11),
"bootstrap": [True, False],
"criterion": ["gini", "entropy"]
}
CV_rfc = RandomizedSearchCV(estimator=rfc,param_distributions=param_grid,cv= 5,scoring='roc_auc')
CV_rfc.fit(X_train, y_train)
print (CV_rfc.best_params_)
seed = 29
rf_clf = CV_rfc.best_estimator_
rf_clf.fit(X_train,y_train)
predictions_train = rf_clf.predict(X_train)
print("Train ROC-AUC :",roc_auc_score(y_train, predictions_train))
# estimate accuracy on validation dataset
predictions = rf_clf.predict(X_test)
print("Test ROC-AUC :",roc_auc_score(y_test, predictions))
visualizeFeatureImportance(rf_clf,X_train.columns,10)
et_clf = ExtraTreesClassifier(n_jobs=-1)
# n_estimator = list(range(10,500,10))
# print(n_estimator)
param_grid = {
'n_estimators': list(range(10,500,10)),
'max_features': ['auto', 'sqrt', 'log2'],
'max_depth' : list(range(3,9,1)),
"min_samples_split": sp_randint(2, 11),
"min_samples_leaf": sp_randint(2, 11),
"bootstrap": [True, False],
"criterion": ["gini", "entropy"]
}
CV_et_clf = RandomizedSearchCV(estimator=et_clf,param_distributions=param_grid,cv= 5,scoring='roc_auc')
CV_et_clf.fit(X_train, y_train)
print (CV_et_clf.best_params_)
seed = 29
et_clf = CV_et_clf.best_estimator_
et_clf.fit(X_train,y_train)
predictions_train = et_clf.predict(X_train)
print("Train ROC-AUC :",roc_auc_score(y_train, predictions_train))
# estimate accuracy on validation dataset
predictions = et_clf.predict(X_test)
print("Test ROC-AUC :",roc_auc_score(y_test, predictions))
visualizeFeatureImportance(et_clf,X_train.columns,10)
# +
#Lasso Regression for variable selection and Random forest for feature imporatnce .
validation_size = 0.30
seed = 23
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=validation_size,random_state=seed)
from sklearn import linear_model
ls_clf = linear_model.Lasso(alpha=0.01,selection='random')
param_grid = { 'alpha': np.arange(0.1,1,0.01)}
CV_lr = GridSearchCV(estimator=ls_clf,param_grid=param_grid, cv=10,scoring='roc_auc')
CV_lr.fit(X_train, y_train)
print (CV_lr.best_params_)
CV_lr.best_estimator_.fit(X_train,y_train)
predictions_train = CV_lr.best_estimator_.predict(X_train)
print("Train AUC :",roc_auc_score(y_train,predictions_train))
predictions = CV_lr.best_estimator_.predict(X_test)
print("Test ROC_AUC :",roc_auc_score(y_test, predictions))
# print (len(CV_lr.best_estimator_.coef_))
# print (len(X.columns))
print (CV_lr.best_estimator_.alpha)
lasso_survival_list = []
for i in range(0,len(X.columns)):
if CV_lr.best_estimator_.coef_[i]!=0:
lasso_survival_list.append(X.columns[i])
print ("Features survied :",len(lasso_survival_list))
#Subselect the data using the variables survived from Lasso
X_sub = X[X.columns.intersection(lasso_survival_list)]
y_sub = y
#Divide
validation_size = 0.30
seed = 29
X_train, X_test, y_train, y_test = train_test_split(X_sub,y_sub,test_size=validation_size,random_state=seed)
from sklearn.grid_search import GridSearchCV, RandomizedSearchCV
from sklearn.datasets import make_classification
from sklearn.ensemble import RandomForestClassifier
from scipy.stats import randint as sp_randint
rfc = RandomForestClassifier(n_jobs=-1)
# n_estimator = list(range(10,500,10))
# print(n_estimator)
param_grid = {
'n_estimators': list(range(10,500,10)),
'max_features': ['auto', 'sqrt', 'log2'],
'max_depth' : list(range(3,9,1)),
"min_samples_split": sp_randint(2, 11),
"min_samples_leaf": sp_randint(2, 11),
"bootstrap": [True, False],
"criterion": ["gini", "entropy"]
}
CV_rfc = RandomizedSearchCV(estimator=rfc,param_distributions=param_grid,cv= 5,scoring='roc_auc')
CV_rfc.fit(X_train, y_train)
print (CV_rfc.best_params_)
seed = 29
rf_clf = CV_rfc.best_estimator_
rf_clf.fit(X_train,y_train)
predictions_train = rf_clf.predict(X_train)
print("Train ROC-AUC :",roc_auc_score(y_train, predictions_train))
# estimate accuracy on validation dataset
predictions = rf_clf.predict(X_test)
print("Test ROC-AUC :",roc_auc_score(y_test, predictions))
visualizeFeatureImportance(rf_clf,X_train.columns,10)
et_clf = ExtraTreesClassifier(n_jobs=-1)
# n_estimator = list(range(10,500,10))
# print(n_estimator)
param_grid = {
'n_estimators': list(range(10,500,10)),
'max_features': ['auto', 'sqrt', 'log2'],
'max_depth' : list(range(3,9,1)),
"min_samples_split": sp_randint(2, 11),
"min_samples_leaf": sp_randint(2, 11),
"bootstrap": [True, False],
"criterion": ["gini", "entropy"]
}
CV_et_clf = RandomizedSearchCV(estimator=et_clf,param_distributions=param_grid,cv= 5,scoring='roc_auc')
CV_et_clf.fit(X_train, y_train)
print (CV_et_clf.best_params_)
seed = 29
et_clf = CV_et_clf.best_estimator_
et_clf.fit(X_train,y_train)
predictions_train = et_clf.predict(X_train)
print("Train ROC-AUC :",roc_auc_score(y_train, predictions_train))
# estimate accuracy on validation dataset
predictions = et_clf.predict(X_test)
print("Test ROC-AUC :",roc_auc_score(y_test, predictions))
visualizeFeatureImportance(et_clf,X_train.columns,10)
# +
#Logistic Regression with L1 Penalty for variable selection and Random forest for feature imporatnce .
validation_size = 0.30
seed = 29
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=validation_size,random_state=seed)
from sklearn import linear_model
scaler = Normalizer().fit(X_train)
normalizedX = scaler.transform(X_train)
lr = LogisticRegression(C=1,n_jobs=10,penalty='l1')
param_grid = { 'C': np.arange(1,100,1)}
CV_lr = RandomizedSearchCV(estimator=lr,param_distributions=param_grid, cv=10,scoring='roc_auc')
CV_lr.fit(normalizedX, y_train)
print (CV_lr.best_params_)
lr = CV_lr.best_estimator_
lr.fit(normalizedX, y_train)
predictions_train = lr.predict_proba(normalizedX)[:,1]
print("Train AUC :",roc_auc_score(y_train,predictions_train))
normalizedX_test = scaler.transform(X_test)
predictions = lr.predict_proba(normalizedX_test)[:,1]
print("Test AUC :",roc_auc_score(y_test,predictions))
print (len(X.columns))
print (lr.C)
print (len(lr.coef_))
survival_list = []
for i in range(0,len(X.columns)):
if lr.coef_[:,i]!=0:
survival_list.append(X.columns[i])
print ("Features survived for RF :",len(survival_list))
#Subselect the data using the variables survived from Lasso
X_sub = X[X.columns.intersection(survival_list)]
y_sub = y
#Divide
validation_size = 0.30
seed = 29
X_train, X_test, y_train, y_test = train_test_split(X_sub,y_sub,test_size=validation_size,random_state=seed)
from sklearn.grid_search import GridSearchCV
from sklearn.datasets import make_classification
from sklearn.ensemble import RandomForestClassifier
rfc = RandomForestClassifier(n_jobs=-1)
# n_estimator = list(range(10,500,10))
# print(n_estimator)
param_grid = {
'n_estimators': list(range(10,100,10)),
'max_features': ['auto', 'sqrt', 'log2'],
'max_depth' : list(range(3,5,1)),
"min_samples_split": sp_randint(2, 11),
"min_samples_leaf": sp_randint(2, 11),
"bootstrap": [True, False],
"criterion": ["gini", "entropy"]
}
CV_rfc = RandomizedSearchCV(estimator=rfc,param_distributions=param_grid,cv= 5,scoring='roc_auc',random_state=seed)
CV_rfc.fit(X_train, y_train)
print (CV_rfc.best_params_)
rf_clf = CV_rfc.best_estimator_
rf_clf.fit(X_train,y_train)
predictions_train = rf_clf.predict(X_train)
print("Train ROC-AUC :",roc_auc_score(y_train, predictions_train))
# estimate accuracy on validation dataset
predictions = rf_clf.predict(X_test)
print("Test ROC-AUC :",roc_auc_score(y_test, predictions))
visualizeFeatureImportance(rf_clf,X_train.columns,10)
#Randomised search for ET Classifier
et_clf = ExtraTreesClassifier(n_jobs=-1)
# n_estimator = list(range(10,500,10))
# print(n_estimator)
param_grid = {
'n_estimators': list(range(10,500,10)),
'max_features': ['auto', 'sqrt', 'log2'],
'max_depth' : list(range(3,5,1)),
"min_samples_split": sp_randint(2, 11),
"min_samples_leaf": sp_randint(2, 11),
"bootstrap": [True, False],
"criterion": ["gini", "entropy"]
}
CV_et_clf = RandomizedSearchCV(estimator=et_clf,param_distributions=param_grid,cv= 5,scoring='roc_auc',
random_state=seed)
CV_et_clf.fit(X_train, y_train)
print (CV_et_clf.best_params_)
seed = 29
et_clf = CV_et_clf.best_estimator_
et_clf.fit(X_train,y_train)
predictions_train = et_clf.predict(X_train)
print("Train ROC-AUC :",roc_auc_score(y_train, predictions_train))
# estimate accuracy on validation dataset
predictions = et_clf.predict(X_test)
print("Test ROC-AUC :",roc_auc_score(y_test, predictions))
visualizeFeatureImportance(et_clf,X_train.columns,10)
# +
# effect_list = []
# for i in range(0,len(X.columns)):
# if lr.coef_[:,i]!=0:
# effect_list.append(X.columns[i]+str(lr.coef_[:,i]))
# print ("LR feature co-efficient :",effect_list)
# -
print("Train Accuracy :",accuracy_score(y_train, predictions_train))
# estimate accuracy on validation dataset
predictions = et_clf.predict(X_test)
print("Test Accuracy :",accuracy_score(y_test, predictions))
# +
# target_names = ['class 0', 'class 1']
# print(classification_report(y_test, predictions, target_names=target_names))
# +
# tn, fp, fn, tp = confusion_matrix(y_test, predictions).ravel()
# +
# tn, fp, fn, tp
# +
import numpy as np
import matplotlib.pyplot as plt
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
from sklearn.datasets import load_digits
from sklearn.model_selection import learning_curve
from sklearn.model_selection import ShuffleSplit
def plot_learning_curve(estimator, title, X, y, ylim=None, cv=None,
n_jobs=1, train_sizes=np.linspace(.1, 1.0, 5)):
"""
Generate a simple plot of the test and training learning curve.
Parameters
----------
estimator : object type that implements the "fit" and "predict" methods
An object of that type which is cloned for each validation.
title : string
Title for the chart.
X : array-like, shape (n_samples, n_features)
Training vector, where n_samples is the number of samples and
n_features is the number of features.
y : array-like, shape (n_samples) or (n_samples, n_features), optional
Target relative to X for classification or regression;
None for unsupervised learning.
ylim : tuple, shape (ymin, ymax), optional
Defines minimum and maximum yvalues plotted.
cv : int, cross-validation generator or an iterable, optional
Determines the cross-validation splitting strategy.
Possible inputs for cv are:
- None, to use the default 3-fold cross-validation,
- integer, to specify the number of folds.
- An object to be used as a cross-validation generator.
- An iterable yielding train/test splits.
For integer/None inputs, if ``y`` is binary or multiclass,
:class:`StratifiedKFold` used. If the estimator is not a classifier
or if ``y`` is neither binary nor multiclass, :class:`KFold` is used.
Refer :ref:`User Guide <cross_validation>` for the various
cross-validators that can be used here.
n_jobs : integer, optional
Number of jobs to run in parallel (default 1).
"""
plt.figure(figsize=(10,10))
plt.title(title)
if ylim is not None:
plt.ylim(*ylim)
plt.xlabel("Training examples")
plt.ylabel("Score")
train_sizes, train_scores, test_scores = learning_curve(
estimator, X, y, cv=cv, n_jobs=n_jobs, train_sizes=train_sizes)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
plt.grid()
plt.fill_between(train_sizes, train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std, alpha=0.1,
color="r")
plt.fill_between(train_sizes, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.1, color="g")
plt.plot(train_sizes, train_scores_mean, 'o-', color="r",
label="Training score")
plt.plot(train_sizes, test_scores_mean, 'o-', color="g",
label="Cross-validation score")
plt.legend(loc="best")
return plt
# -
title = "Learning Curves (Extra Tress Classifier)"
# Cross validation with 100 iterations to get smoother mean test and train
# score curves, each time with 20% data randomly selected as a validation set.
cv = ShuffleSplit(n_splits=50,test_size=0.3, random_state=0)
estimator = et_clf
plot_learning_curve(estimator, title, X, y, ylim=(0.5, 1.01), cv=cv, n_jobs=4)
plt.show()
title = "Learning Curves (Random Tress Classifier)"
# Cross validation with 100 iterations to get smoother mean test and train
# score curves, each time with 20% data randomly selected as a validation set.
cv = ShuffleSplit(n_splits=50,test_size=0.3, random_state=0)
estimator = rf_clf
plot_learning_curve(estimator, title, X, y, ylim=(0.5, 1.01), cv=cv)
plt.show()
title = "Learning Curves (Decision Tree Classifier)"
# Cross validation with 100 iterations to get smoother mean test and train
# score curves, each time with 20% data randomly selected as a validation set.
cv = ShuffleSplit(n_splits=50,test_size=0.3, random_state=0)
estimator = CV_dtcf.best_estimator_
plot_learning_curve(estimator, title, X, y, ylim=(0.5, 1.01), cv=cv)
plt.show()
title = "Learning Curves (Logistic Regression Classifier)"
# Cross validation with 100 iterations to get smoother mean test and train
# score curves, each time with 20% data randomly selected as a validation set.
cv = ShuffleSplit(n_splits=10,test_size=0.3, random_state=0)
estimator = CV_lr.best_estimator_
plot_learning_curve(estimator, title, X, y, ylim=(0.5, 1.01),cv=cv)
plt.show()
| Modelling Techniques - Total Sales Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2-Filtering
# This tutorial demonstrates how to filter PDB to create subsets of structures. For details see [filters](https://github.com/sbl-sdsc/mmtf-pyspark/tree/master/mmtfPyspark/filters) and [demos](https://github.com/sbl-sdsc/mmtf-pyspark/tree/master/demos/filters).
# ### Import pyspark and mmtfPyspark
from pyspark.sql import SparkSession
from mmtfPyspark.io import mmtfReader
from mmtfPyspark.filters import ContainsGroup, ContainsLProteinChain, PolymerComposition, Resolution
from mmtfPyspark.structureViewer import view_group_interaction
# ### Configure Spark
spark = SparkSession.builder.master("local[4]").appName("2-Filtering").getOrCreate()
# ### Read PDB structures
path = "../resources/mmtf_reduced_sample"
pdb = mmtfReader.read_sequence_file(path).cache()
pdb.count()
# ## Filter by Quality Metrics
# Structures can be filtered by [Resolution](https://pdb101.rcsb.org/learn/guide-to-understanding-pdb-data/resolution) and [R-free](https://pdb101.rcsb.org/learn/guide-to-understanding-pdb-data/r-value-and-r-free). Each filter takes a minimum and maximum values. The example below returns structures with a resolution in the inclusive range [0.0, 1.5]
pdb = pdb.filter(Resolution(0.0, 1.5))
pdb.count()
# ## Filter by Polymer Chain Types
# A number of filters are available to filter by the type of the polymer chain.
# ### Create a subset of structures that contain at least one L-protein chain
pdb = pdb.filter(ContainsLProteinChain())
pdb.count()
# ### Create a subset of structure that exclusively contain L-protein chains (e.g., exclude protein-nucleic acid complexes)
pdb = pdb.filter(ContainsLProteinChain(exclusive=True))
pdb.count()
# ### Keep protein structures that exclusively contain chains made out of the 20 standard amino acids
pdb = pdb.filter(PolymerComposition(PolymerComposition.AMINO_ACIDS_20, exclusive=True))
pdb.count()
# ## Find the subset of structures that contains ATP
pdb = pdb.filter(ContainsGroup("ATP"))
# ## Visualize the hits
view_group_interaction(pdb.keys().collect(),"ATP");
# ## Filter with a lambda expression
# Rather than using a pre-made filter, we can create simple filters using lambda expressions. The expression needs to evaluate to a boolean type.
#
# The variable t in the lambda expression below represents a tuple and t[1] is the second element in the tuple representing the mmtfStructure.
#
# Here, we filter by the number of atoms in an entry. You will learn more about extracting structural information from an mmtfStructure in future tutorials.
pdb = pdb.filter(lambda t: t[1].num_atoms < 500)
pdb.count()
# Or, we can filter by the key, represented by the first element in a tuple: t[0].
#
# **Keys are case sensitive. Always use upper case PDB IDs in mmtf-pyspark!**
pdb = pdb.filter(lambda t: t[0] in ["4AFF", "4CBU"])
pdb.count()
spark.stop()
| 3-mmtf-pyspark/2-Filtering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img style="float: right" src="https://upload.wikimedia.org/wikipedia/de/thumb/7/7b/DWD-Logo_2013.svg/500px-DWD-Logo_2013.svg.png" />
# # Unzip bz2 files
import bz2
import glob
# extract data that was downloaded from https://opendata.dwd.de/weather/nwp/icon-eu/
path = "/shared/eduard"
for filename in glob.glob("{}/*.bz2".format(path)):
print (filename)
with bz2.open(filename, "rb") as f:
# Decompress data from file
outfilename = filename.split(".bz2")[0]
with open(outfilename, 'wb') as out:
out.write(f.read())
print ("-> {}".format(outfilename))
# +
import xarray as xr
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# read dwd grib data from local directory
x = None
path = "/shared/eduard"
for filename in glob.glob("{}/icon-eu_*.grib2".format(path)):
print (filename)
tmp = xr.open_dataset(filename, engine='cfgrib')
tmp = tmp - 273.15 # Convert to celsius
if x is None:
x = tmp
else:
x = xr.concat([x, tmp], "valid_time")
# -
# display contents
x
# Plotting xarray https://xarray.pydata.org/en/v0.7.1/plotting.html
# plot 3 dimensional, optionally: wrap after 2 plots
x.t2m.plot(x="longitude", y="latitude", col="valid_time") #, col_wrap=2)
| dwd-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: v3
# language: python
# name: v-jpt-3
# ---
# # Code to explore structure of hdf5 data files
#
# June 9, 2020: Adding gaussian smoothing
# +
import numpy as np
import h5py
import os
import glob
import time
from scipy.ndimage import gaussian_filter ### For gausian filtering
import matplotlib.pyplot as plt
# -
# %matplotlib widget
# +
### Explore the hdf5 file
def f_explore_file(fname):
'''
Explore the structure of the hdf5 file
Attributes are : ['dataset_tag','seed9','universe_tag']
The Keys are : ['full', 'namePar', 'physPar', 'redshifts', 'unitPar']
'full' is an array of shape (512,512,512,4)
The last index 4 corresponds to red-shift. Eg. 0, 0.5, 1.5, 3.0
'''
dta=h5py.File(fname,'r')
### Attributes
attrs=dta.attrs
print('Attributes',[(i,attrs[i]) for i in attrs])
### Keys
keys=dta.keys()
print('\nKeys',keys)
print("\nThe key: 'full' ")
print('Shape of the array',dta['full'].shape)
print('\nOther keys')
for key in ['namePar', 'physPar', 'redshifts', 'unitPar']:
print(key,dta[key][:])
# -
# #### Sample exploration of files
fname='/global/cfs/cdirs/m3363/www/cosmoUniverse_2020_11_4parE_cGAN/Sg0.5/univ_ics_2019-03_a10582192.hdf5'
fname='/global/cfs/cdirs/m3363/www/cosmoUniverse_2020_08_4parEgrid/Om0.15_Sg0.5_H100.0/univ_ics_2019-03_a16305120.hdf5'
f_explore_file(fname)
512**3/(64**3)
# ### Read in list of file
### Location of hdf5 files
data_dir='/global/project/projectdirs/m3363/www/cosmoUniverse_2019_08_const/'
### Extract list of hdf5 files
f_list=glob.glob(data_dir+'*.hdf5')
len(f_list)
h5py.File(f_list[0],'r')['full'][:,:,:,0].shape
# +
# for i in f_list[:5]:
# f_explore_file(i)
# -
# ### Exploring Gaussian filtering
# **Gaussian blurring**: https://en.wikipedia.org/wiki/Gaussian_blur#:~:text=In%20image%20processing%2C%20a%20Gaussian,image%20noise%20and%20reduce%20detail \
# **Paper using it**: https://arxiv.org/abs/1801.09070
#
dta=h5py.File(fname,'r')
arr=np.array(dta['full'])
# %timeit filtered_arr=gaussian_filter(arr, sigma=0.5,mode='wrap')
def f_compare_pixel_intensity(img_lst,label_lst=['img1','img2'],bkgnd_arr=None,log_scale=True, normalize=True, mode='avg',bins=25, hist_range=None):
'''
Module to compute and plot histogram for pixel intensity of images
Has 2 modes : simple and avg
simple mode: No errors. Just flatten the input image array and compute histogram of full data
avg mode(Default) :
- Compute histogram for each image in the image array
- Compute errors across each histogram
bkgnd_arr : histogram of this array is plotting with +/- sigma band
'''
norm=normalize # Whether to normalize the histogram
def f_batch_histogram(img_arr,bins,norm,hist_range):
''' Compute histogram statistics for a batch of images'''
## Extracting the range. This is important to ensure that the different histograms are compared correctly
if hist_range==None : ulim,llim=np.max(img_arr),np.min(img_arr)
else: ulim,llim=hist_range[1],hist_range[0]
# print(ulim,llim)
### array of histogram of each image
hist_arr=np.array([np.histogram(arr.flatten(), bins=bins, range=(llim,ulim), density=norm) for arr in img_arr]) ## range is important
hist=np.stack(hist_arr[:,0]) # First element is histogram array
# print(hist.shape)
bin_list=np.stack(hist_arr[:,1]) # Second element is bin value
### Compute statistics over histograms of individual images
mean,err=np.mean(hist,axis=0),np.std(hist,axis=0)/np.sqrt(hist.shape[0])
bin_edges=bin_list[0]
centers = (bin_edges[:-1] + bin_edges[1:]) / 2
# print(bin_edges,centers)
return mean,err,centers
plt.figure()
## Plot background distribution
if bkgnd_arr is not None:
if mode=='simple':
hist, bin_edges = np.histogram(bkgnd_arr.flatten(), bins=bins, density=norm, range=hist_range)
centers = (bin_edges[:-1] + bin_edges[1:]) / 2
plt.errorbar(centers, hist, color='k',marker='*',linestyle=':', label='bkgnd')
elif mode=='avg':
### Compute histogram for each image.
mean,err,centers=f_batch_histogram(bkgnd_arr,bins,norm,hist_range)
plt.plot(centers,mean,linestyle=':',color='k',label='bkgnd')
plt.fill_between(centers, mean - err, mean + err, color='k', alpha=0.4)
### Plot the rest of the datasets
for img,label in zip(img_lst,label_lst):
if mode=='simple':
hist, bin_edges = np.histogram(img.flatten(), bins=bins, density=norm, range=hist_range)
centers = (bin_edges[:-1] + bin_edges[1:]) / 2
plt.errorbar(centers, hist, fmt='o-', label=label)
elif mode=='avg':
### Compute histogram for each image.
mean,err,centers=f_batch_histogram(img,bins,norm,hist_range)
# print('Centers',centers)
plt.errorbar(centers,mean,yerr=err,fmt='o-',label=label)
if log_scale:
plt.yscale('log')
plt.xscale('log')
plt.legend()
plt.xlabel('Pixel value')
plt.ylabel('Counts')
plt.title('Pixel Intensity Histogram')
f_compare_pixel_intensity([arr,filtered_arr],label_lst=['raw','filtered'],mode='simple',normalize=True)
fname='/global/cfs/cdirs/m3363/vayyar/cosmogan_data/raw_data/3d_data/dataset5_3dcgan_4univs_64cube_simple_splicing/Om0.3_Sg0.5_H70.0.npy'
a1=np.load(fname,mmap_mode='r')
print(a1.shape)
512**3/
| code/0a_data_preprocessing/0_explore_hdf5_files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Building a Basic CNN: The MNIST Dataset
#
# In this notebook, we will build a simple CNN-based architecture to classify the 10 digits (0-9) of the MNIST dataset. The objective of this notebook is to become familiar with the process of building CNNs in Keras.
#
# We will go through the following steps:
# 1. Importing libraries and the dataset
# 2. Data preparation: Train-test split, specifying the shape of the input data etc.
# 3. Building and understanding the CNN architecture
# 4. Fitting and evaluating the model
#
# Let's dive in.
# ## 1. Importing Libraries and the Dataset
#
# Let's load the required libraries. From Keras, we need to import two main components:
# 1. `Sequential` from `keras.models`: `Sequential` is the keras abstraction for creating models with a stack of layers (MLP has multiple hidden layers, CNNs have convolutional layers, etc.).
# 2. Various types of layers from `keras.layers`: These layers are added (one after the other) to the `Sequential` model
#
# The keras `backend` is needed for keras to know that you are using tensorflow (not Theano) at the backend (the backend is <a href="https://keras.io/backend/">specified in a JSON file</a>).
#
import numpy as np
import random
import keras
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
from keras import backend as K
# ! nvidia-smi
# Let's load the MNIST dataset from `keras.datasets`. The download may take a few minutes.
# load the dataset into train and test sets
(x_train, y_train), (x_test, y_test) = mnist.load_data()
print("train data")
print(x_train.shape)
print(y_train.shape)
print("\n test data")
print(x_test.shape)
print(y_test.shape)
# So we have 60,000 training and 10,000 test images each of size 28 x 28. Note that the images are grayscale and thus are stored as 2D arrays.<br>
#
# Also, let's sample only 20k images for training (just to speed up the training a bit).
# sample only 20k images for training
idx = np.random.randint(x_train.shape[0], size=20000) # sample 20k indices from 0-60,000
x_train = x_train[idx, :]
y_train = y_train[idx]
print(x_train.shape)
print(y_train.shape)
# ## 2. Data Preparation
#
# Let's prepare the dataset for feeding to the network. We will do the following three main steps:<br>
#
# #### 2.1 Reshape the Data
# First, let's understand the shape in which the network expects the training data.
# Since we have 20,000 training samples each of size (28, 28, 1), the training data (`x_train`) needs to be of the shape `(20000, 28, 28, 1)`. If the images were coloured, the shape would have been `(20000, 28, 28, 3)`.
#
# Further, each of the 20,000 images have a 0-9 label, so `y_train` needs to be of the shape `(20000, 10)` where each image's label is represented as a 10-d **one-hot encoded vector**.
#
# The shapes of `x_test` and `y_test` will be the same as that of `x_train` and `y_train` respectively.
#
# #### 2.2 Rescaling (Normalisation)
# The value of each pixel is between 0-255, so we will **rescale each pixel** by dividing by 255 so that the range becomes 0-1. Recollect <a href="https://stats.stackexchange.com/questions/185853/why-do-we-need-to-normalize-the-images-before-we-put-them-into-cnn">why normalisation is important for training NNs</a>.
#
# #### 2.3 Converting Input Data Type: Int to Float
# The pixels are originally stored as type `int`, but it is advisable to feed the data as `float`. This is not really compulsory, but advisable. You can read <a href="https://datascience.stackexchange.com/questions/13636/neural-network-data-type-conversion-float-from-int">why conversion from int to float is helpful here</a>.
#
# +
# specify input dimensions of each image
img_rows, img_cols = 28, 28
input_shape = (img_rows, img_cols, 1)
# batch size, number of classes, epochs
batch_size = 60
num_classes = 10
epochs = 12
# -
# Let's now reshape the array `x_train` from shape `(20000, 28, 28)`to `(20000, 28, 28, 1)`. Analogously for `x_test`.
# reshape x_train and x_test
x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols, 1)
x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols, 1)
print(x_train.shape)
print(x_test.shape)
# Now let's reshape `y_train` from `(20000,)` to `(20000, 10)`. This can be conveniently done using the keras' `utils` module.
# convert class labels (from digits) to one-hot encoded vectors
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
print(y_train.shape)
# Finally, let's convert the data type of `x_train` and `x_test` from int to float and normalise the images.
# originally, the pixels are stored as ints
x_train.dtype
# +
# convert int to float
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
# normalise
x_train /= 255
x_test /= 255
# -
# ## 3. Building the Model
# Let's now build the CNN architecture. For the MNIST dataset, we do not need to build a very sophisticated CNN - a simple shallow-ish CNN would suffice.
#
# We will build a network with:
# - two convolutional layers having 32 and 64 filters respectively,
# - followed by a max pooling layer,
# - and then `Flatten` the output of the pooling layer to give us a long vector,
# - then add a fully connected `Dense` layer with 128 neurons, and finally
# - add a `softmax` layer with 10 neurons
#
# The generic way to build a model in Keras is to instantiate a `Sequential` model and keep adding `keras.layers` to it. We will also use some dropouts.
# +
# model
model = Sequential()
# a keras convolutional layer is called Conv2D
# help(Conv2D)
# note that the first layer needs to be told the input shape explicitly
# first conv layer
model.add(Conv2D(32, kernel_size=(3, 3),
activation='relu',
input_shape=input_shape)) # input shape = (img_rows, img_cols, 1)
# second conv layer
model.add(Conv2D(64, kernel_size=(3, 3),
activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
# flatten and put a fully connected layer
model.add(Flatten())
model.add(Dense(128, activation='relu')) # fully connected
model.add(Dropout(0.5))
# softmax layer
model.add(Dense(num_classes, activation='softmax'))
# model summary
model.summary()
# -
# #### Understanding Model Summary
#
# It is a good practice to spend some time staring at the model summary above and verify the number of parameteres, output sizes etc. Let's do some calculations to verify that we understand the model deeply enough.
#
# - Layer-1 (Conv2D): We have used 32 kernels of size (3, 3), and each kernel has a single bias, so we have 32 x 3 x 3 (weights) + 32 (biases) = 320 parameters (all trainable). Note that the kernels have only one channel since the input images are 2D (grayscale). By default, a convolutional layer uses stride of 1 and no padding, so the output from this layer is of shape 26 x 26 x 32, as shown in the summary above (the first element `None` is for the batch size).
#
# - Layer-2 (Conv2D): We have used 64 kernels of size (3, 3), but this time, each kernel has to convolve a tensor of size (26, 26, 32) from the previous layer. Thus, the kernels will also have 32 channels, and so the shape of each kernel is (3, 3, 32) (and we have 64 of them). So we have 64 x 3 x 3 x 32 (weights) + 64 (biases) = 18496 parameters (all trainable). The output shape is (24, 24, 64) since each kernel produces a (24, 24) feature map.
#
# - Max pooling: The pooling layer gets the (24, 24, 64) input from the previous conv layer and produces a (12, 12, 64) output (the default pooling uses stride of 2). There are no trainable parameters in the pooling layer.
#
# - The `Dropout` layer does not alter the output shape and has no trainable parameters.
#
# - The `Flatten` layer simply takes in the (12, 12, 64) output from the previous layer and 'flattens' it into a vector of length 12 x 12 x 64 = 9216.
#
# - The `Dense` layer is a plain fully connected layer with 128 neurons. It takes the 9216-dimensional output vector from the previous layer (layer l-1) as the input and has 128 x 9216 (weights) + 128 (biases) = 1179776 trainable parameters. The output of this layer is a 128-dimensional vector.
#
# - The `Dropout` layer simply drops a few neurons.
#
# - Finally, we have a `Dense` softmax layer with 10 neurons which takes the 128-dimensional vector from the previous layer as input. It has 128 x 10 (weights) + 10 (biases) = 1290 trainable parameters.
#
# Thus, the total number of parameters are 1,199,882 all of which are trainable.
# ## 4. Fitting and Evaluating the Model
#
# Let's now compile and train the model.
# usual cross entropy loss
# choose any optimiser such as adam, rmsprop etc
# metric is accuracy
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
# fit the model
# this should take around 10-15 minutes when run locally on a windows/mac PC
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
# evaluate the model on test data
model.evaluate(x_test, y_test)
print(model.metrics_names)
# The final loss (on test data) is about 0.04 and the accuracy is 98.59%.
| CNN/Building+a+CNN+-+MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 介绍了LeNet-5网络的架构
import torch
import torch.nn.functional as F
class LeNet5(torch.nn.Module):
def __init__(self):
super(LeNet5, self).__init__()
# 卷积层
self.conv1 = torch.nn.Conv2d(1, 6, 3) # 1通道——>6通道,卷积核尺寸为3x3
self.conv2 = torch.nn.Conv2d(6, 16, 5) # 6通道——>16通道,卷积核尺寸为5x5
# 全连接层
self.fc1 = torch.nn.Linear(16*5*5, 120)
self.fc2 = torch.nn.Linear(120, 84)
self.fc3 = torch.nn.Linear(84, 10)
def forward(self, x):
# 输入图像尺寸为1*32*32
x = F.relu(self.conv1(x)) # 1*32*32 ——> 6*30*30
x = F.max_pool2d(x, (2, 2)) # 6*30*30 ——> 6*15*15
x = F.relu(self.conv2(x)) # 6*15*15 ——> 16*11*11
x = F.max_pool2d(x, (2, 2)) # 16*11*11 ——> 16*5*5
x = x.view(-1, self.num_flat_features(x)) # flatten
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
def num_flat_features(self, x):
# x和forward的原始输入并不一样
# forward定义的时候不需要显示声明dim=0的batch维度,但在真实前向计算时,是需要考虑这个维度的
size = x.size()[1:]
num_featurs = 1
for s in size:
num_featurs *= s
return num_featurs
net = LeNet5()
print(net)
| Pytorch_demos/basic_tutorail/2.3 LeNet-5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import csv
#Files to load
import_file_budget = os.path.join("budget_data.csv")
output_file_budget = os.path.join("budget_analysis.txt")
#Set up variables
total_months = 0
month_of_change = []
net_change_list = []
greatest_increase = ["", 0]
greatest_decrease = ["", 999999999999999999999]
total_net= 0
#Read csv
with open(import_file_budget) as financial_data:
reader= csv.reader(financial_data)
#Read header row
header = next(reader)
print(header)
#Look at first row to avoid Appending to net_change_list
first_row = next (reader)
print(first_row)
total_months = total_months + 1
print(first_row[1])
total_net = total_net + int(first_row[1])
prev_net = int(first_row[1])
print(total_net)
print(prev_net)
#For loop
for row in reader:
#Track total
total_months= total_months+1
total_net = total_net + int(row[1])
#print(total_net)
#Track new change
net_change = int(row[1]) - prev_net
prev_net = int(row[1])
net_change_list = net_change_list + [net_change]
month_of_change = month_of_change + [row[0]]
#print(net_change_list)
#print(month_of_change)
if net_change > greatest_increase[1]:
greatest_increase[0] = row [0]
greatest_increase[1] = net_change
#print("Greatest Increase")
#print(greatest_increase)
if net_change < greatest_decrease[1]:
greatest_decrease[0] = row[0]
greatest_decrease[1] = net_change
#Calculate Average Net Change
net_monthly_avg = sum(net_change_list)/len(net_change_list)
output = (
f"\nFinancial Analysis\n"
f"======================\n"
f"Total Months:{total_months}\n"
f"Total: ${total_net}\n"
f"Average Change: ${net_monthly_avg:.2f}\n"
f"Greatest Increase In Profits {greatest_increase[0]}, (${greatest_increase[1]})\n"
f"Greatest Decrease In Profits {greatest_decrease[0]}, (${greatest_decrease[1]})\n"
)
print (output)
#
with open(output_file_budget, "w") as txt_file:
txt_file.write(output)
# -
| PyBank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# +
# 2.1 Parsing a FASTA file
## <NAME>:
# with open() as f, open () as
# for i, line in enumerate(f):
# line=...rstrip()
# if i >0:
# f_out.write
with open('data/salmonella_spi1_region.fna','r') as f:
f_lines=f.readlines()
# -
np.shape(f_lines)
f_lines[:5]
f_nogap=''.join(f_lines[1:])
f_nogap = f_nogap.replace("\n", "")
# +
# 2.2 Pathogenicity islands
# a)
def segment_into_blocks(seq,block_size=1000):
"""segement the genome into blocks"""
blocks = [f_nogap[block_size*i:block_size*(i+1)] for i in range(len(seq)//block_size)]
return blocks
def gc_content_per_block(block):
"""compute gc content for one block"""
n_gc=block.count('G')+block.count('C')
gc_content=n_gc/len(block)
return gc_content
# -
5%3
5 //3
def gc_blocks(seq, block_size=1000):
blocks=segment_into_blocks(seq)
gc_list=[gc_content_per_block(i) for i in blocks]
return tuple(gc_list)
gc_tuple=gc_blocks(f_nogap)
gc_tuple
# +
# b)
def gc_map(seq, block_size=1000, gc_thresh=0.45):
block_tuple=tuple(segment_into_blocks(seq))
# gc tuple 直接用了上一问的结果
my_dict=dict(zip(gc_tuple,block_tuple))
gc_list=[]
for key,seq in my_dict.items():
if key<=gc_thresh:
seq=seq.lower()
gc_list.append(seq)
final_gc_list=''.join(gc_list)
return final_gc_list
# -
GC_map=gc_map(f_nogap)
# +
# d)
def segment_back_to_blocks(seq,block_size=1000):
"""segement the genome into blocks"""
#block_size=2000
blocks = [f_nogap[block_size*i:block_size*(i+1)]+'\n' for i in range(len(seq)//block_size)]
return blocks
f_back_to_segments=segment_back_to_blocks(GC_map,block_size=60)
# -
f_back_to_segments.insert(0,f_lines[0])
f_back_to_segments
''.join(f_back_to_segments)
with open('converted_FASTA', 'w') as f:
f.write(str(f_back_to_segments))
# +
# 2.3 ORF detection
# a)
def segmentation_to_three(seq,shift):
seg_list=[seq[x+shift:x+3+shift] for x in range(0,len(seq),3)] # some non-triple left-overs
print(seg_list)
start_pos=[i for i, x in enumerate(seg_list) if x == "ATG"]
stop_pos= [i for i, x in enumerate(seg_list) if x == ('TGA' or 'TAG' or 'TAA')]
return start_pos,stop_pos
# -
def find_long(start_pos,stop_pos):
max_length=0
max_start=0
max_end=0
i=0
pos_array=np.where(np.array(start_pos)<stop_pos[i])
print(np.where(np.array(start_pos)<stop_pos[i]))
print(len(pos_array[0]))
if len(pos_array[0]) >0:
start=np.min(np.where(np.array(start_pos)<stop_pos[i]))
print(start)
length=i-start+3
if length>max_length:
max_length=length
max_start=start
max_end=i
for i in range(1,len(stop_pos)):
pos_array=np.where(np.array(start_pos)<stop_pos[i] and np.array(start_pos)>stop_pos[i-1])
print(pos_array)
if len(pos_array[0]) >0:
start=np.min(np.where(np.array(start_pos)<stop_pos[i] and np.array(start_pos)>stop_pos[i-1] ))
print(start)
length=i-start+3
if length>max_length:
max_length=length
max_start=start
max_end=i
return max_length,max_start,max_end
def longest_orf(seq):
max0=find_long(*segmentation_to_three(seq,0))[0]
max1=find_long(*segmentation_to_three(seq,1))[0]
max3=find_long(*segmentation_to_three(seq,2))[0]
seq='GGATGATGATGTAAAACAGCTTCGATCGAT'
longest_orf(seq)
stop_pos
while True:
if seq[3*i:3*i+3]=='ATG':
find('TGA', 'TAG', 'TAA')
elif seq[3*i+1:3*i+4]=='ATG':
i+=1
def longest_orf(seq):
pos=seq.find('ATG')
y=[1,2,3,4,5,6,6]
y.index(6)
5==3 or 4 or 5
np.array([1,2,3])
len(np.array([ ]))
| day2_excercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Accessing data in a DataSet
#
# After a measurement is completed all the acquired data and metadata around it is accessible via a `DataSet` object. This notebook presents the useful methods and properties of the `DataSet` object which enable convenient access to the data, parameters information, and more. For general overview of the `DataSet` class, refer to [DataSet class walkthrough](DataSet-class-walkthrough.ipynb).
# ## Preparation: a DataSet from a dummy Measurement
#
# In order to obtain a `DataSet` object, we are going to run a `Measurement` storing some dummy data (see notebook on [Performing measurements using qcodes parameters and dataset](Performing-measurements-using-qcodes-parameters-and-dataset.ipynb) notebook for more details).
# +
import tempfile
import os
import numpy as np
import qcodes
from qcodes import initialise_or_create_database_at, \
load_or_create_experiment, Measurement, Parameter, \
Station
from qcodes.dataset.plotting import plot_dataset
# +
db_path = os.path.join(tempfile.gettempdir(), 'data_access_example.db')
initialise_or_create_database_at(db_path)
exp = load_or_create_experiment(experiment_name='greco', sample_name='draco')
# -
x = Parameter(name='x', label='Voltage', unit='V',
set_cmd=None, get_cmd=None)
t = Parameter(name='t', label='Time', unit='s',
set_cmd=None, get_cmd=None)
y = Parameter(name='y', label='Voltage', unit='V',
set_cmd=None, get_cmd=None)
y2 = Parameter(name='y2', label='Current', unit='A',
set_cmd=None, get_cmd=None)
q = Parameter(name='q', label='Qredibility', unit='$',
set_cmd=None, get_cmd=None)
# +
meas = Measurement(exp=exp, name='fresco')
meas.register_parameter(x)
meas.register_parameter(t)
meas.register_parameter(y, setpoints=(x, t))
meas.register_parameter(y2, setpoints=(x, t))
meas.register_parameter(q) # a standalone parameter
x_vals = np.linspace(-4, 5, 50)
t_vals = np.linspace(-500, 1500, 25)
with meas.run() as datasaver:
for xv in x_vals:
for tv in t_vals:
yv = np.sin(2*np.pi*xv)*np.cos(2*np.pi*0.001*tv) + 0.001*tv
y2v = np.sin(2*np.pi*xv)*np.cos(2*np.pi*0.001*tv + 0.5*np.pi) - 0.001*tv
datasaver.add_result((x, xv), (t, tv), (y, yv), (y2, y2v))
q_val = np.max(yv) - np.min(y2v) # a meaningless value
datasaver.add_result((q, q_val))
dataset = datasaver.dataset
# -
# For the sake of demonstrating what kind of data we've produced, let's use `plot_dataset` to make some default plots of the data.
plot_dataset(dataset)
# ## DataSet indentification
#
# Before we dive into what's in the `DataSet`, let's briefly note how a `DataSet` is identified.
dataset.captured_run_id
dataset.exp_name
dataset.sample_name
dataset.name
# ## Parameters in the DataSet
#
# In this section we are getting information about the parameters stored in the given `DataSet`.
#
# > Why is that important? Let's jump into *data*!
#
# As it turns out, just "arrays of numbers" are not enough to reason about a given `DataSet`. Even comping up with a reasonable deafult plot, which is what `plot_dataset` does, requires information on `DataSet`'s parameters. In this notebook, we first have a detailed look at what is stored about parameters and how to work with this information. After that, we will cover data access methods.
# ### Run description
#
# Every dataset comes with a "description" (aka "run description"):
dataset.description
# The description, an instance of `RunDescriber` object, is intended to describe the details of a dataset. In the future releases of QCoDeS it will likely be expanded. At the moment, it only contains an `InterDependencies_` object under its `interdeps` attribute - which stores all the information about the parameters of the `DataSet`.
#
# Let's look into this `InterDependencies_` object.
# ### Interdependencies
#
# `Interdependencies_` object inside the run description contains information about all the parameters that are stored in the `DataSet`. Subsections below explain how the individual information about the parameters as well as their relationships are captured in the `Interdependencies_` object.
interdeps = dataset.description.interdeps
interdeps
# #### Dependencies, inferences, standalones
# Information about every parameter is stored in the form of `ParamSpecBase` objects, and the releationship between parameters is captured via `dependencies`, `inferences`, and `standalones` attributes.
#
# For example, the dataset that we are inspecting contains no inferences, and one standalone parameter `q`, and two dependent parameters `y` and `y2`, which both depend on independent `x` and `t` parameters:
interdeps.inferences
interdeps.standalones
interdeps.dependencies
# `dependencies` is a dictionary of `ParamSpecBase` objects. The keys are dependent parameters (those which depend on other parameters), and the corresponding values in the dictionary are tuples of independent parameters that the dependent parameter in the key depends on. Coloquially, each key-value pair of the `dependencies` dictionary is sometimes referred to as "parameter tree".
#
# `inferences` follows the same structure as `dependencies`.
#
# `standalones` is a set - an unordered collection of `ParamSpecBase` objects representing "standalone" parameters, the ones which do not depend on other parameters, and no other parameter depends on them.
# #### ParamSpecBase objects
# `ParamSpecBase` object contains all the necessary information about a given parameter, for example, its `name` and `unit`:
ps = list(interdeps.dependencies.keys())[0]
print(f'Parameter {ps.name!r} is in {ps.unit!r}')
# `paramspecs` property returns a tuple of `ParamSpecBase`s for all the parameters contained in the `Interdependencies_` object:
interdeps.paramspecs
# Here's a trivial example of iterating through dependent parameters of the `Interdependencies_` object and extracting information about them from the `ParamSpecBase` objects:
for d in interdeps.dependencies.keys():
print(f'Parameter {d.name!r} ({d.label}, {d.unit}) depends on:')
for i in interdeps.dependencies[d]:
print(f'- {i.name!r} ({i.label}, {i.unit})')
# #### Other useful methods and properties
# `Interdependencies_` object has a few useful properties and methods which make it easy to work it and with other `Interdependencies_` and `ParamSpecBase` objects.
#
# For example, `non_dependencies` returns a tuple of all dependent parameters together with standalone parameters:
interdeps.non_dependencies
# `what_depends_on` method allows to find what parameters depend on a given parameter:
# +
t_ps = interdeps.paramspecs[2]
t_deps = interdeps.what_depends_on(t_ps)
print(f'Following parameters depend on {t_ps.name!r} ({t_ps.label}, {t_ps.unit}):')
for t_dep in t_deps:
print(f'- {t_dep.name!r} ({t_dep.label}, {t_dep.unit})')
# -
# ### Shortcuts to important parameters
#
# For the frequently needed groups of parameters, `DataSet` object itself provides convenient methods and properties.
#
# For example, use `dependent_parameters` property to get only dependent parameters of a given `DataSet`:
dataset.dependent_parameters
# This is equivalent to:
tuple(dataset.description.interdeps.dependencies.keys())
# ### Note on inferences
#
# Inferences between parameters is a feature that has not been used yet within QCoDeS. The initial concepts around `DataSet` included it in order to link parameters that are not directly dependent on each other as "dependencies" are. It is very likely that "inferences" will be eventually deprecated and removed.
# ### Note on ParamSpec's
#
# > `ParamSpec`s originate from QCoDeS versions prior to `0.2.0` and for now are kept for backwards compatibility. `ParamSpec`s are completely superseded by `InterDependencies_`/`ParamSpecBase` bundle and will likely be deprecated in future versions of QCoDeS together with the `DataSet` methods/properties that return `ParamSpec`s objects.
#
# In addition to the `Interdependencies_` object, `DataSet` also holds `ParamSpec` objects (not to be confused with `ParamSpecBase` objects from above). Similar to `Interdependencies_` object, the `ParamSpec` objects hold information about parameters and their interdependencies but in a different way: for a given parameter, `ParamSpec` object itself contains information on names of parameters that it depends on, while for the `InterDependencies_`/`ParamSpecBase`s this information is stored only in the `InterDependencies_` object.
# `DataSet` exposes `paramspecs` property and `get_parameters()` method, both of which return `ParamSpec` objects of all the parameters of the dataset, and are not recommended for use:
dataset.paramspecs
dataset.get_parameters()
dataset.parameters
# To give an example of what it takes to work with `ParamSpec` objects as opposed to `Interdependencies_` object, here's a function that one needs to write in order to find standalone `ParamSpec`s from a given list of `ParamSpec`s:
# +
def get_standalone_parameters(paramspecs):
all_independents = set(spec.name
for spec in paramspecs
if len(spec.depends_on_) == 0)
used_independents = set(d for spec in paramspecs for d in spec.depends_on_)
standalones = all_independents.difference(used_independents)
return tuple(ps for ps in paramspecs if ps.name in standalones)
all_parameters = dataset.get_parameters()
standalone_parameters = get_standalone_parameters(all_parameters)
standalone_parameters
# -
# ## Getting data from DataSet
#
# In this section methods for retrieving the actual data from the `DataSet` are discussed.
#
# ### `get_parameter_data` - the powerhorse
#
# `DataSet` provides one main method of accessing data - `get_parameter_data`. It returns data for groups of dependent-parameter-and-its-independent-parameters in a form of a nested dictionary of `numpy` arrays:
dataset.get_parameter_data()
# #### Avoid excessive calls to loading data
#
# Note that this call actually reads the data of the `DataSet` and in case of a `DataSet` with a lot of data can take noticable amount of time. Hence, it is recommended to limit the number of times the same data gets loaded in order to speed up the user's code.
# #### Loading data of selected parameters
#
# Sometimes data only for a particular parameter or parameters needs to be loaded. For example, let's assume that after inspecting the `InterDependencies_` object from `dataset.description.interdeps`, we concluded that we want to load data of the `q` parameter and the `y2` parameter. In order to do that, we just pass the names of these parameters, or their `ParamSpecBase`s to `get_parameter_data` call:
q_param_spec = list(interdeps.standalones)[0]
q_param_spec
y2_param_spec = interdeps.non_dependencies[-1]
y2_param_spec
dataset.get_parameter_data(q_param_spec, y2_param_spec)
# ### `to_pandas_dataframe_dict` and `to_pandas_dataframe` - for `pandas` fans
#
# `DataSet` provides two methods for accessing data with `pandas` - `to_pandas_dataframe` and `to_pandas_dataframe_dict`. The method `to_pandas_dataframe_dict` returns data for groups of dependent-parameter-and-its-independent-parameters in a form of a dictionary of `pandas.DataFrame` s, while `to_pandas_dataframe` returns a concatendated `pandas.DataFrame` for groups of dependent-parameter-and-its-independent-parameters:
# +
df_dict = dataset.to_pandas_dataframe_dict()
# For the sake of making this article more readable,
# we will print the contents of the `dfs` dictionary
# manually by calling `.head()` on each of the DataFrames
for parameter_name, df in df_dict.items():
print(f"DataFrame for parameter {parameter_name}")
print("-----------------------------")
print(f"{df.head()!r}")
print("")
# -
# Alternativly to concatinate the DataSet data into a single pandas Dataframe run the following:
df = dataset.to_pandas_dataframe()
print(f"{df.head()!r}")
# Similar to `get_parameter_data`, `to_pandas_dataframe_dict` and `to_pandas_dataframe_dict` also supports retrieving data for a given parameter(s), as well as `start`/`stop` arguments.
#
# Both `to_pandas_dataframe` and `to_pandas_dataframe_dict` is implemented based on `get_parameter_data`, hence the performance considerations mentioned above for `get_parameter_data` apply to these methods as well.
#
# For more details on `to_pandas_dataframe` refer to [Working with pandas and xarray article](Working-With-Pandas-and-XArray.ipynb).
# ### Exporting to other file formats
#
# The dataset support exporting to netcdf and csv via the `dataset.export` method. See [Exporting QCoDes Datasets](./Exporting-data-to-other-file-formats.ipynb) for more information.
#
# ### Data extraction into "other" formats
#
# If the user desires to export a QCoDeS `DataSet` into a format that is not readily supported by `DataSet` methods, we recommend to use `to_pandas_dataframe_dict` or `to_pandas_dataframe_dict` first, and then convert the resulting `DataFrame` s into a the desired format. This is becuase `pandas` package already implements converting `DataFrame` to various popular formats including comma-separated text file (`.csv`), HDF (`.hdf5`), xarray, Excel (`.xls`, `.xlsx`), and more; refer to [Working with pandas and xarray article](Working-With-Pandas-and-XArray.ipynb), and [`pandas` documentation](https://pandas.pydata.org/pandas-docs/stable/reference/frame.html#serialization-io-conversion) for more information.
#
# Refer to the docstrings of those methods for more information on how to use them.
| docs/examples/DataSet/Accessing-data-in-DataSet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# n a row of trees, the i-th tree produces fruit with type tree[i].
#
# You start at any tree of your choice, then repeatedly perform the following steps:
#
# Add one piece of fruit from this tree to your baskets. If you cannot, stop.
# Move to the next tree to the right of the current tree. If there is no tree to the right, stop.
# Note that you do not have any choice after the initial choice of starting tree: you must perform step 1, then step 2, then back to step 1, then step 2, and so on until you stop.
#
# You have two baskets, and each basket can carry any quantity of fruit, but you want each basket to only carry one type of fruit each.
#
# What is the total amount of fruit you can collect with this procedure?
#
#
#
# Example 1:
#
# <b>Input: [1,2,1]
#
# Output: 3</b>
#
# Explanation: We can collect [1,2,1].
#
# Example 2:
#
# <b>Input: [0,1,2,2]
#
# Output: 3</b>
# Explanation: We can collect [1,2,2].
# If we started at the first tree, we would only collect [0, 1].
#
# Example 3:
# <b>
# Input: [1,2,3,2,2]
#
# Output: 4
# </b>
#
# Explanation: We can collect [2,3,2,2].
# If we started at the first tree, we would only collect [1, 2].
# Example 4:
#
# <b>
# Input: [3,3,3,1,2,1,1,2,3,3,4]
#
# Output: 5
# </b>
#
# Explanation: We can collect [1,2,1,1,2].
# If we started at the first tree or the eighth tree, we would only collect 4 fruits.
#
#
# Note:
#
# 1 <= tree.length <= 40000
# 0 <= tree[i] < tree.length
class Solution(object):
def totalFruit(self, tree):
blocks = [(k, len(list(v)))
for k, v in itertools.groupby(tree)]
ans = i = 0
while i < len(blocks):
# We'll start our scan at block[i].
# types : the different values of tree[i] seen
# weight : the total number of trees represented
# by blocks under consideration
types, weight = set(), 0
# For each block from i and going forward,
for j in xrange(i, len(blocks)):
# Add each block to consideration
types.add(blocks[j][0])
weight += blocks[j][1]
# If we have 3 types, this is not a legal subarray
if len(types) >= 3:
i = j-1
break
ans = max(ans, weight)
# If we go to the last block, then stop
else:
break
return ans
import collections
class Solution(object):
def totalFruit(self, tree):
ans = i = 0
count = collections.Counter()
for j, x in enumerate(tree):
count[x] += 1
while len(count) >= 3:
count[tree[i]] -= 1
if count[tree[i]] == 0:
del count[tree[i]]
i += 1
ans = max(ans, j - i + 1)
return ans
test = Solution()
test.totalFruit([1,2,1])
# +
import collections
def fruit(tree):
ans = i = 0
count=collections.Counter()
for j,x in enumerate(tree):
count[x] += 1
print(count)
print(len(count))
print("j",j)
print("x",x)
while len(count) >= 3:
print("tree[i]",tree[i])
print("count[tree[i]]",count[tree[i]])
count[tree[i]] -= 1
if count[tree[i]] == 0:
del count[tree[i]]
i += 1
ans = max(ans, j - i + 1)
return ans
print(fruit([1,2,2,1,4]))
# -
| array_strings/ipynb/.ipynb_checkpoints/fruit_into_basket-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import model
import os
model_name = 'siamese-u-net'
DATASET_DIR = os.path.join(os.getcwd(), 'cluttered_omniglot/')
LOG_DIR = os.path.join(os.getcwd(), 'logs/' + model_name + '/')
# -
# ### Train
# +
# Train on all clutter levels sequentially
number_of_characters = [4, 8, 16, 32, 64, 128, 256]
for chars in range(len(number_of_characters)):
print('')
datadir = DATASET_DIR + '%.d_characters/'%(number_of_characters[chars])
logdir = LOG_DIR + '%.d_characters/'%(number_of_characters[chars])
model.training(datadir,
logdir,
epochs=20,
model=model_name,
feature_maps=24,
batch_size=250,
learning_rate=0.0005,
initial_training=True,
maximum_number_of_steps=0)
# -
# ### Evaluate
# +
# Evaluate all clutter levels sequentially
number_of_characters = [4, 8, 16, 32, 64, 128, 256]
for chars in range(len(number_of_characters)):
print('')
datadir = DATASET_DIR + '%.d_characters/'%(number_of_characters[chars])
logdir = LOG_DIR + '%.d_characters/'%(number_of_characters[chars])
model.evaluation(datadir,
logdir,
model=model_name,
feature_maps=24,
batch_size=250,
threshold=0.3,
max_steps=0)
| siamese-u-net.ipynb |
(defparameter grouping
(make-instance 'cljw::vbox :children
(vector
(make-instance 'cljw::hbox :children
(vector (make-instance 'cljw::button :description "a") (make-instance 'cljw::button :description "c")))
(make-instance 'cljw::hbox :children
(vector (make-instance 'cljw::button :description "b") (make-instance 'cljw::button :description "d")))
(make-instance 'cljw::text :value ""))))
grouping
(let ((textbox (aref (cljw::children grouping) 2)))
(defun add-to-textbox (button)
(setf (cljw::value textbox) (concatenate 'string (cljw::value textbox) (cljw::description button)))))
(let* ((children (cljw::children grouping))
(hbox1 (aref children 0))
(hbox1c (cljw::children hbox1))
(A (aref hbox1c 0))
(B (aref hbox1c 1))
(hbox2 (aref children 1))
(hbox2c (cljw::children hbox2))
(C (aref hbox2c 0))
(D (aref hbox2c 1)))
(cljw::on-click A #'add-to-textbox)
(cljw::on-click B #'add-to-textbox)
(cljw::on-click C #'add-to-textbox)
(cljw::on-click D #'add-to-textbox))
| src/demos/.ipynb_checkpoints/button-callbacks-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # KNN similarity search strategies
#
# This code generates Fig. 8
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# +
import numpy as np
import json
import h5py
import os
import sys
from time import time
import warnings
# Ignore warnings as they just pollute the output
warnings.filterwarnings('ignore')
# Enable importing modules from the parent directory
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
module_path = os.path.abspath(os.path.join('../experiments'))
if module_path not in sys.path:
sys.path.append(module_path)
# DNase-seq 2011, hg19
bw = 'data/ENCFF158GBQ.bigWig'
# -
# ## Download the data and the autoencoder
# +
from download import download_encode_file, download_file
from pathlib import Path
Path('data').mkdir(parents=True, exist_ok=True)
Path('models').mkdir(parents=True, exist_ok=True)
download_encode_file('ENCFF158GBQ.bigWig')
download_file(
"https://zenodo.org/record/2609763/files/dnase_w-12000_r-100.h5?download=1",
"dnase_w-12000_r-100.h5",
dir="models"
)
# -
# ## Helper methods
#
# #### Simple kNN search
# +
from scipy.spatial.distance import cdist
def knn(data, target_idx, k, metric='euclidean', sax = None, ignore: int = 0, sort_only: bool = False):
"""K nearest neighbors
Find the `k` nearest neighbors of a
"""
target = data[target_idx]
if sort_only:
dist = data
else:
if sax is None:
dist = cdist(data, target.reshape((1, target.size)), metric='euclidean').flatten()
else:
N = data.shape[0]
dist = np.zeros(N)
for i in range(N):
dist[i] = sax.distance_sax(target, data[i])
# Ensure that the target is always first
dist[target_idx] = -1
for i in range(1, ignore + 1):
dist[min(target_idx + i, data.shape[0] - 1)] = -1
dist[max(target_idx - i, 0)] = -1
return np.argsort(dist)[1 + (2 * ignore):k + 1 + (2 * ignore)]
# -
# #### DTW distance
# +
from scipy.spatial.distance import euclidean
from fastdtw import fastdtw
from multiprocessing import Pool
def dtw(data, target_idx: int, n: int, target = None, return_all = False, print_progress: bool = False):
N = data.shape[0]
dist = np.zeros(N)
if target is None:
target = data[target_idx]
p = ((np.arange(4) + 1) * (N // 4)).astype(int)
for i in np.arange(N):
if i in p and print_progress:
print('.', end='', flush=True)
d, _ = fastdtw(data[i], target, dist=euclidean)
dist[i] = d
if return_all:
return dist
return np.argsort(dist)[:n]
def pooled_dtw(data, target_idx: int, target = None, print_progress: bool = False, ignore: int = 0, num_threads: int = None):
if target is None:
target = data[target_idx]
with Pool(num_threads) as pool:
args = [[d, 0, -1, target, True, print_progress] for d in np.array_split(data, pool._processes)]
return np.concatenate(pool.starmap(dtw, args))
# -
# #### Normalized cross correlation search
# +
from scipy.signal import correlate
def norm(data, zero_norm: bool = False):
mean = np.mean(data) if zero_norm else 0
return (data - mean) / np.std(data)
def norm2d(data, zero_norm: bool = False):
mean = np.mean(data, axis=1).reshape(-1, 1) if zero_norm else np.zeros((data.shape[0], 1))
std = np.std(data, axis=1).reshape(-1, 1)
return (data - mean) / std
def xcorrelation(data, template_idx, n, normalize=False, zero_normalize=False, ignore: int = 0):
unknown = data
template = data[template_idx]
if norm:
unknown = norm2d(unknown, zero_norm=zero_normalize)
template = norm(template, zero_norm=zero_normalize)
xcorr = np.apply_along_axis(lambda m: correlate(m, template, mode='full'), axis=1, arr=unknown)
xcorr[np.where(np.isnan(xcorr))] = 0
max_xcorr = np.nanmax(xcorr, axis=1)
# Ensure that the target is always last
max_xcorr[template_idx] = -1
for i in range(1, ignore + 1):
max_xcorr[min(template_idx + i, data.shape[0] - 1)] = -1
max_xcorr[max(template_idx - i, 0)] = -1
return np.argsort(max_xcorr)[::-1][:n]
# -
# # 12 KB Search
# +
from server import bigwig
data_12kb = bigwig.chunk(bw, 12000, 100, 12000 / 6, ['chr1'], verbose=True)
# +
from ae.utils import plot_windows_from_data
k_12kb = 20 # Number of KNNs to be saved later on
targets_12kb = [80503, 43895, 33430, 42575, 6112, 91938, 82896, 1060, 11975]
targets_12kb_ex = 12933
with open('data/targets-12kb.json', 'w') as outfile:
json.dump(targets_12kb, outfile)
plot_windows_from_data(data_12kb, window_ids=targets_12kb)
# +
"""Compute the CAE latent space"""
from ae.utils import get_models, predict
encoder_12kb, decoder_12kb, autoencoder_12kb = get_models('models/dnase_w-12000_r-100.h5', loss_fn='bce')
t0 = time()
predicted_12kb, _, latent_12kb = predict(
encoder_12kb,
decoder_12kb,
data_12kb.reshape(data_12kb.shape[0], data_12kb.shape[1], 1)
)
print('Done! Took {:.2f} seconds ({:.1f} minutes).'.format(time() - t0, (time() - t0) / 60))
with h5py.File('data/cae_12kb.h5', 'w') as f:
f.create_dataset('latent_space', data=latent_12kb, dtype=np.float32)
# +
"""Compute UMAP embedding"""
import umap
t0 = time()
umap_embedding_12kb = umap.UMAP(
n_neighbors=10,
min_dist=0.01,
metric='l2',
n_components=10,
).fit_transform(data_12kb)
print('Done! Took {:.2f} seconds ({:.1f} minutes).'.format(time() - t0, (time() - t0) / 60))
with h5py.File('data/umap_12kb.h5', 'w') as f:
f.create_dataset('umap', data=umap_embedding_12kb, dtype=np.float32)
# +
"""Compute TSFRESH"""
import h5py
import pandas as pd
from tsfresh import extract_features
N = data_12kb.shape[0]
L = data_12kb.shape[1]
tsfresh_12kb_df = pd.DataFrame(
np.concatenate(
(
np.repeat(np.arange(data_12kb.shape[0]), data_12kb.shape[1]).reshape((-1, 1)),
data_12kb.reshape((-1, 1))
),
axis=1
),
columns=['id', 'value']
)
batch_size = 1000
t0 = time()
with h5py.File('data/tsfresh_12kb.h5', 'w') as f:
f.create_dataset('features', shape=(N, 794), dtype=np.float32)
for i in np.arange(0, N, batch_size):
batch = extract_features(
tsfresh_12kb_df[i * L:(i + batch_size) * L],
column_id='id',
n_jobs=4,
).values
f['features'][i:i + batch_size] = batch
print('Done! Took {:.2f} seconds ({:.1f} minutes).'.format(time() - t0, (time() - t0) / 60))
# +
"""Conpute DTW"""
import umap
dtw_12kb = np.zeros((data_12kb.shape[0], len(targets_12kb)))
print('Compute DTW:')
t0 = time()
for i, target in enumerate(targets_12kb):
t1 = time()
dtw_12kb[:,i] = pooled_dtw(data_12kb, target, num_threads=1)
print('Target #{} done! Took {:.2f} seconds ({:.1f} minutes).'.format(i, time() - t1, (time() - t1) / 60))
print('All done! Took {:.2f} seconds ({:.1f} minutes).'.format(time() - t0, (time() - t0) / 60))
with h5py.File('data/dtw_12kb.h5', 'w') as f:
f.create_dataset('dtw', data=dtw_12kb, dtype=np.float32)
# +
"""Compute SAX"""
from tslearn.piecewise import SymbolicAggregateApproximation
t0 = time()
sax_12kb = SymbolicAggregateApproximation(n_segments=120, alphabet_size_avg=10)
sax_data_12kb = sax_12kb.fit_transform(data_12kb)
print('Done! Took {:.2f} seconds ({:.1f} minutes).'.format(time() - t0, (time() - t0) / 60))
# +
from time import time
with h5py.File('data/cae_12kb.h5', 'r') as f:
cae_12kb = f['latent_space'][:]
with h5py.File('data/umap_12kb.h5', 'r') as f:
umap_12kb = f['umap'][:]
with h5py.File('data/dtw_12kb.h5', 'r') as f:
dtw_12kb = f['dtw'][:]
with h5py.File('data/tsfresh_12kb.h5', 'r') as f:
tsfresh_12kb = f['features'][:]
# Some features do not seem to computable. Lets set them to zero
tsfresh_12kb[np.isnan(tsfresh_12kb)] = 0.0
with h5py.File('data/12kb-similarity-search.h5', 'w') as f:
f.create_dataset('knn_ae', shape=(len(targets_12kb), k_12kb), dtype=np.int)
f.create_dataset('knn_eq', shape=(len(targets_12kb), k_12kb), dtype=np.int)
f.create_dataset('knn_sax', shape=(len(targets_12kb), k_12kb), dtype=np.int)
f.create_dataset('knn_dtw', shape=(len(targets_12kb), k_12kb), dtype=np.int)
f.create_dataset('knn_umap', shape=(len(targets_12kb), k_12kb), dtype=np.int)
f.create_dataset('knn_tsfresh', shape=(len(targets_12kb), k_12kb), dtype=np.int)
f.create_dataset('top_xcorr', shape=(len(targets_12kb), k_12kb), dtype=np.int)
for i, target in enumerate(targets_12kb):
t0 = time()
print('Search for window #{}'.format(target), end='', flush=True)
f['knn_ae'][i] = knn(latent_12kb, target, k_12kb, ignore=2)
print('.', end='', flush=True)
f['knn_eq'][i] = knn(data_12kb, target, k_12kb, ignore=2)
print('.', end='', flush=True)
f['knn_sax'][i] = knn(sax_data_12kb, target, k_12kb, sax=sax_12kb, ignore=2)
print('.', end='', flush=True)
f['knn_umap'][i] = knn(umap_embedding_12kb, target, k_12kb, ignore=2)
print('.', end='', flush=True)
f['top_xcorr'][i] = xcorrelation(data_12kb, target, k_12kb, normalize=True, zero_normalize=True, ignore=2)
print('.', end='', flush=True)
f['knn_tsfresh'][i] = knn(tsfresh_12kb, target, k_12kb)
print('.', end='', flush=True)
f['knn_dtw'][i] = knn(dtw_12kb[i], target, k_12kb, sort_only=True, ignore=2)
print('. done! Took {:.2f} seconds ({:.1f} minutes).'.format(time() - t0, (time() - t0) / 60))
# +
import h5py
import json
import matplotlib.pyplot as plt
import numpy as np
with h5py.File('data/12kb-similarity-search.h5', 'r') as f:
knn_ae_12kb = f['knn_ae'][:]
knn_eq_12kb = f['knn_eq'][:]
knn_sax_12kb = f['knn_sax'][:]
knn_dtw_12kb = f['knn_dtw'][:]
knn_umap_12kb = f['knn_umap'][:]
knn_tsfresh_12kb = f['knn_tsfresh'][:]
top_xcorr_12kb = f['top_xcorr'][:]
show = 5
N = (show + 1) * 7
T = len(targets_12kb)
sz = data_12kb[0].size
plt.figure(figsize=(6 * T, N))
ymax = 1.0
show_predictions = False
for i, target in enumerate(targets_12kb):
ax = plt.subplot(N, T, (i + 1))
ax.set_facecolor("#eeeeee")
plt.bar(np.arange(sz), data_12kb[target], color='#000000', width=1.0)
plt.ylim(0, ymax)
plt.xticks([], [])
plt.yticks([], [])
for j, hit in enumerate(knn_ae_12kb[i][:show]):
plt.subplot(N, T, ((j + 1) * T) + (i + 1))
plt.bar(np.arange(sz), data_12kb[hit], color='#d24f00', width=1.0) # orange = CAE
plt.ylim(0, ymax)
plt.xticks([], [])
plt.yticks([], [])
plt.subplots_adjust(top=0.9)
for j, hit in enumerate(knn_eq_12kb[i][:show]):
plt.subplot(N, T, ((j + 6) * T) + (i + 1))
plt.bar(np.arange(sz), data_12kb[hit], color='#008ca8', width=1.0) # blue = EQ
plt.ylim(0, ymax)
plt.xticks([], [])
plt.yticks([], [])
for j, hit in enumerate(knn_sax_12kb[i][:show]):
plt.subplot(N, T, ((j + 11) * T) + (i + 1))
plt.bar(np.arange(sz), data_12kb[hit], color='#a6227a', width=1.0) # purple = SAX
plt.ylim(0, ymax)
plt.xticks([], [])
plt.yticks([], [])
for j, hit in enumerate(knn_dtw_12kb[i][:show]):
plt.subplot(N, T, ((j + 16) * T) + (i + 1))
plt.bar(np.arange(sz), data_12kb[hit], color='#209e4e', width=1.0) # green = DTW
plt.ylim(0, ymax)
plt.xticks([], [])
plt.yticks([], [])
for j, hit in enumerate(top_xcorr_12kb[i][:show]):
plt.subplot(N, T, ((j + 21) * T) + (i + 1))
plt.bar(np.arange(sz), data_12kb[hit], color='#bf9f00', width=1.0) # yellow = Zero-nornalized X correlation
plt.ylim(0, ymax)
plt.xticks([], [])
plt.yticks([], [])
for j, hit in enumerate(knn_umap_12kb[i][:show]):
plt.subplot(N, T, ((j + 26) * T) + (i + 1))
plt.bar(np.arange(sz), data_12kb[hit], color='#bc2626', width=1.0) # red = UMAP
plt.ylim(0, ymax)
plt.xticks([], [])
plt.yticks([], [])
for j, hit in enumerate(knn_tsfresh_12kb[i][:show]):
plt.subplot(N, T, ((j + 31) * T) + (i + 1))
plt.bar(np.arange(sz), data_12kb[hit], color='#5943b2', width=1.0) # purple = tsfresh
plt.ylim(0, ymax)
plt.xticks([], [])
plt.yticks([], [])
| experiments/notebooks/Compare similarity search strategies (figure 8).ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .js
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: "JS (unsafe) \u2014 Jyve"
// language: javascript
// name: jyve-kyrnel-js-unsafe
// ---
// # Jyve JS
// Just like it says on the tin, this runs whatever JS your browser can currently support.
x = 1
++x
JSON.stringify(x)
// Modern browsers support:
// - the [`class` keyword](https://caniuse.com/#feat=es6-class) for inheritance without `prototype`
// -
// ```
// `${string} templates`
// ``` for concise string manipulation
// - [`async/await` keywords](https://caniuse.com/#feat=async-functions) for asynchronous code (inside functions)
// - and more!
class Adder {
add(a, b) {
document.write(
`🐍 says <i>${a} plussss ${b} issss ${a + b}</i><br/>`
)
return a + b;
}
}
this.Adder = Adder
let it = new this.Adder();
it.add(1, 1)
sir_not_appearing_in_this_notebook
| notebooks/JS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # **Amazon Lookout for Equipment** - Getting started
# *Part 3 - Model training*
# ## Initialization
# ---
# This repository is structured as follow:
#
# ```sh
# . lookout-equipment-demo
# |
# ├── data/
# | ├── interim # Temporary intermediate data are stored here
# | ├── processed # Finalized datasets are usually stored here
# | | # before they are sent to S3 to allow the
# | | # service to reach them
# | └── raw # Immutable original data are stored here
# |
# ├── getting_started/
# | ├── 1_data_preparation.ipynb
# | ├── 2_dataset_creation.ipynb
# | ├── 3_model_training.ipynb <<< THIS NOTEBOOK <<<
# | ├── 4_model_evaluation.ipynb
# | ├── 5_inference_scheduling.ipynb
# | └── 6_cleanup.ipynb
# |
# └── utils/
# └── lookout_equipment_utils.py
# ```
# ### Notebook configuration update
# Amazon Lookout for Equipment being a very recent service, we need to make sure that we have access to the latest version of the AWS Python packages. If you see a `pip` dependency error, check that the `boto3` version is ok: if it's greater than 1.17.48 (the first version that includes the `lookoutequipment` API), you can discard this error and move forward with the next cell:
# +
# !pip install --quiet --upgrade boto3 awscli aiobotocore botocore sagemaker tqdm
import boto3
print(f'boto3 version: {boto3.__version__} (should be >= 1.17.48 to include Lookout for Equipment API)')
# Restart the current notebook to ensure we take into account the previous updates:
from IPython.core.display import HTML
HTML("<script>Jupyter.notebook.kernel.restart()</script>")
# -
# ### Imports
# +
import config
import os
import pandas as pd
import sagemaker
import sys
# Helper functions for managing Lookout for Equipment API calls:
sys.path.append('../utils')
import lookout_equipment_utils as lookout
# -
ROLE_ARN = sagemaker.get_execution_role()
REGION_NAME = boto3.session.Session().region_name
BUCKET = config.BUCKET
PREFIX = config.PREFIX_LABEL
DATASET_NAME = config.DATASET_NAME
MODEL_NAME = config.MODEL_NAME
# Based on the label time ranges, we will use the following time ranges:
#
# * **Train set:** 1st January 2019 - 31st July 2019: Lookout for Equipment needs at least 180 days of training data and this period contains a few labelled ranges with some anomalies.
# * **Evaluation set:** 1st August 2019 - 27th October 2019 *(this test set includes both normal and abnormal data to evaluate our model on)*
# +
# Configuring time ranges:
training_start = pd.to_datetime('2019-01-01 00:00:00')
training_end = pd.to_datetime('2019-07-31 00:00:00')
evaluation_start = pd.to_datetime('2019-08-01 00:00:00')
evaluation_end = pd.to_datetime('2019-10-27 00:00:00')
print(f' Training period | from {training_start} to {training_end}')
print(f'Evaluation period | from {evaluation_start} to {evaluation_end}')
# -
# ## Model training
# ---
# +
# Prepare the model parameters:
lookout_model = lookout.LookoutEquipmentModel(model_name=MODEL_NAME,
dataset_name=DATASET_NAME,
region_name=REGION_NAME)
# Set the training / evaluation split date:
lookout_model.set_time_periods(evaluation_start,
evaluation_end,
training_start,
training_end)
# Set the label data location:
lookout_model.set_label_data(bucket=BUCKET,
prefix=PREFIX,
access_role_arn=ROLE_ARN)
# This sets up the rate the service will resample the data before
# training: we will keep the original sampling rate in this example
# (5 minutes), but feel free to use a larger sampling rate to accelerate
# the training time:
# lookout_model.set_target_sampling_rate(sampling_rate='PT15M')
# -
# The following method encapsulates a call to the [**CreateModel**](https://docs.aws.amazon.com/lookout-for-equipment/latest/ug/API_CreateModel.html) API:
# Actually create the model and train it:
lookout_model.train()
# A training is now in progress as captured by the console:
#
# 
#
# Use the following cell to capture the model training progress. **This model should take around 30-45 minutes to be trained.** Key drivers for training time usually are:
# * **Number of labels** in the label dataset (if provided)
# * Number of datapoints: this number depends on the **sampling rate**, the **number of time series** and the **time range**.
#
# The following method encapsulate a call to the [**DescribeModel**](https://docs.aws.amazon.com/lookout-for-equipment/latest/ug/API_DescribeModel.html) API and collect the model progress by looking at the `Status` field retrieved from this call:
lookout_model.poll_model_training(sleep_time=300)
# A model is now trained and we can visualize the results of the back testing on the evaluation window selected at the beginning on this notebook:
#
# 
# In the console, **you can click on each detected event**: Amazon Lookout for Equipment unpacks the ranking and display the top sensors contributing to the detected events.
#
# When you open this window, the first event is already selected and this is the detailed view you will get from the console:
#
# 
# This dataset contains 30 sensors:
# * If each sensor contributed the same way to this event, every sensors would **equally contribute** to this event (said otherwise, every sensor would have a similar feature importance of `100% / 30 = 3.33%`).
# * The top sensors (e.g. **Sensor19** with a **5.67% importance**) have a contribution that is significantly higher than this threshold, which is statistically relevant.
# * If the model continues outputing detected anomalies with a similar ranking, this might push a maintenance operator to go and have a look at the associated components.
# ## Conclusion
# ---
# In this notebook, we use the dataset created in part 2 of this notebook series and trained an Amazon Lookout for Equipment model.
#
# From here you can either head:
# * To the next notebook where we will **extract the evaluation data** for this model and use it to perform further analysis on the model results: this is optional and just gives you some pointers on how to post-process and visualize the data provided by Amazon Lookout for Equipment.
# * Or to the **inference scheduling notebook** where we will start the model, feed it some new data and catch the results.
| getting_started/3_model_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Xopt class, TNK test function
#
# This is the class method for running Xopt.
#
# TNK function
# $n=2$ variables:
# $x_i \in [0, \pi], i=1,2$
#
# Objectives:
# - $f_i(x) = x_i$
#
# Constraints:
# - $g_1(x) = -x_1^2 -x_2^2 + 1 + 0.1 \cos\left(16 \arctan \frac{x_1}{x_2}\right) \le 0$
# - $g_2(x) = (x_1 - 1/2)^2 + (x_2-1/2)^2 \le 0.5$
# +
# Import the class
from xopt import Xopt
# Notebook printing output
from xopt import output_notebook
output_notebook()
# -
# The `Xopt` object can be instantiated from a JSON or YAML file, or a dict, with the proper structure.
#
# Here we will make one
import yaml
# Make a proper input file.
YAML="""
xopt: {output_path: null}
algorithm:
name: cnsga
options:
max_generations: 50
population_size: 128
show_progress: True
simulation:
name: test_TNK
evaluate: xopt.tests.evaluators.TNK.evaluate_TNK
vocs:
variables:
x1: [0, 3.14159]
x2: [0, 3.14159]
objectives: {y1: MINIMIZE, y2: MINIMIZE}
constraints:
c1: [GREATER_THAN, 0]
c2: [LESS_THAN, 0.5]
linked_variables: {x9: x1}
constants: {a: dummy_constant}
"""
config = yaml.safe_load(YAML)
# +
# Optional: Connect the function directly
#from xopt.evaluators.test_TNK import evaluate_TNK
#config['simulation']['evaluate'] = evaluate_TNK
X = Xopt(config)
X
# -
# Note that the repr string contains all of the config information
X.random_evaluate()
# # Run CNSGA
#
# CNSGA is designed to run in parallel with an asynchronous executor as defined in PEP 3148
# +
#import logging, sys
#logging.basicConfig(format='%(message)s', level=logging.INFO, stream=sys.stdout)
# +
# Pick one of these
#from concurrent.futures import ThreadPoolExecutor as PoolExecutor
from concurrent.futures import ProcessPoolExecutor as PoolExecutor
executor = PoolExecutor()
# This will also work.
#executor=None
# -
# Change max generations
X.algorithm['options']['max_generations'] = 10
X.run(executor=executor)
# The last population is saved internally:
list(X.results)
X.results['outputs'][0:5]
# Increase the max_generations, and it will continue where it left off
X.algorithm['options']['max_generations'] = 20
X.run(executor=executor)
# # Run with MPI
# +
# X.save('test.json')
# # !mpirun -n 4 python -m mpi4py.futures -m xopt.mpi.run -vv --logfile xopt.log test.json
# -
# # Plot
# +
# Extract objectives from output
key1, key2 = list(X.vocs['objectives'])
x = [o[key1] for o in X.results['outputs']]
y = [o[key2] for o in X.results['outputs']]
import matplotlib.pyplot as plt
# %matplotlib inline
fig, ax = plt.subplots(figsize=(5,5))
ax.scatter(x, y, color='blue')
#ax.set_xlim(X_RANGE)
#ax.set_ylim(Y_RANGE)
ax.set_xlabel(key1)
ax.set_ylabel(key2)
ax.set_aspect('auto')
#ax.set_title(NAME)
| docs/examples/xopt_basic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Standard imports
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import sys
sys.path.insert(0, '/Users/tareen/Desktop/Research_Projects/2020_mavenn_github/mavenn_git_ssh_local')
# Load mavenn
import mavenn
print(mavenn.__path__)
# -
GB1_WT_seq = 'QYKLILNGKTLKGETTTEAVDAATAEKVFKQYANDNGVDGEWTYDDATKTFTVTE'
# raw single mutants data from the Gb1 paper.
GB1_data_single_mutants = pd.read_csv('GB1_single_mutant_data/oslon_data_single_mutants_ambler.csv')
GB1_data_single_mutants.head()
# +
def load_olson_data_GB1():
"""
Helper function to turn single mutant data provided by
Olson et al. into sequence-values arrays.
return
------
gb1_df: (pd dataframe)
dataframe containing sequences (single)
and their corresponding log2 enrichment values.
pseudo count of 1 is added to numerator and
denominator.
"""
# GB1 WT sequences
WT_seq = 'QYKLILNGKTLKGETTTEAVDAATAEKVFKQYANDNGVDGEWTYDDATKTFTVTE'
# WT sequence library and selection counts.
WT_input_count = 1759616
WT_selection_count = 3041819
# lists that will contain sequences and their values
sequences = []
enrichment = []
input_ct = []
selected_ct = []
# load single mutants data
oslon_single_mutant_positions_data = pd.read_csv('GB1_single_mutant_data/oslon_data_single_mutants_ambler.csv',
na_values="nan")
# add WT_sequence to top
sequences.append(GB1_WT_seq)
enrichment.append(1)
input_ct.append(1759616)
selected_ct.append(3041819)
for loop_index in range(len(oslon_single_mutant_positions_data)):
mut_index = int(oslon_single_mutant_positions_data['Position'][loop_index]) - 2
mut = oslon_single_mutant_positions_data['Mutation'][loop_index]
temp_seq = list(WT_seq)
temp_seq[mut_index] = mut
# calculate enrichment for sequence
input_count = oslon_single_mutant_positions_data['Input Count'][loop_index]
selection_count = oslon_single_mutant_positions_data['Selection Count'][loop_index]
input_ct.append(input_count)
selected_ct.append(selection_count)
# added pseudo count to ensure log doesn't throw up
temp_fitness = ((selection_count + 1) / (input_count+1)) / (WT_selection_count / WT_input_count)
sequences.append(''.join(temp_seq))
enrichment.append(temp_fitness)
enrichment = np.array(enrichment).copy()
gb1_df = pd.DataFrame({'x': sequences,'input_ct':input_ct,'selected_ct':selected_ct, 'y': np.log2(enrichment)}, columns=['x','input_ct', 'selected_ct','y'])
return gb1_df
gb1_single_mutants_df = load_olson_data_GB1()
# -
gb1_single_mutants_df['set']='training'
gb1_single_mutants_df
# +
# Load example data
data_df = mavenn.load_example_dataset('gb1')
# # Separate test from data_df
# ix_test = data_df['set']=='test'
# test_df = data_df[ix_test].reset_index(drop=True)
# print(f'test N: {len(test_df):,}')
# # Remove test data from data_df
# data_df = data_df[~ix_test].reset_index(drop=True)
# print(f'training + validation N: {len(data_df):,}')
data_df
# -
data_df = gb1_single_mutants_df.append(data_df, ignore_index=True).copy()
data_df.shape
gb1_single_mutants_df.to_csv('GB1_single_mutant_data/GB1_single_mutant_data_mavenn_format.csv.gz')
| mavenn/development/21.11.07_revision_development/21.12.19.get_GB1_single_mutant_data_from_raw_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# %reload_ext autoreload
# %autoreload 2
import multiprocessing
import os
from torch import autograd
from fastai.transforms import TfmType
from fasterai.transforms import *
from fastai.conv_learner import *
from fasterai.images import *
from fasterai.dataset import *
from fasterai.visualize import *
from fasterai.callbacks import *
from fasterai.loss import *
from fasterai.modules import *
from fasterai.training import *
from fasterai.generators import *
from fastai.torch_imports import *
from pathlib import Path
from itertools import repeat
import tensorboardX
torch.cuda.set_device(0)
plt.style.use('dark_background')
torch.backends.cudnn.benchmark=True
# +
IMAGENET = Path('data/imagenet/ILSVRC/Data/CLS-LOC/train')
IMAGENET_SMALL = IMAGENET/'n01440764'
colorizer_path = IMAGENET.parent/('colorize_gen_192.h5')
default_sz=400
torch.backends.cudnn.benchmark=True
# -
netG = Unet34(nf_factor=2).cuda()
load_model(netG, colorizer_path)
netG = netG.eval()
x_tfms = [BlackAndWhiteTransform()]
data_loader = ImageGenDataLoader(sz=256, bs=8, path=IMAGENET_SMALL, random_seed=42, keep_pct=1.0, x_tfms=x_tfms)
md = data_loader.get_model_data()
vis = ModelImageVisualizer(default_sz=default_sz)
vis.plot_transformed_image("test_images/overmiller.jpg", netG, md.val_ds, tfms=x_tfms, sz=400)
vis.plot_transformed_image("test_images/einstein_beach.jpg", netG, md.val_ds, tfms=x_tfms,sz=700)
vis.plot_transformed_image("test_images/abe.jpg", netG, md.val_ds, tfms=x_tfms, sz=380)
vis.plot_transformed_image("test_images/airmen1943.jpg", netG, md.val_ds, tfms=x_tfms, sz=650)
vis.plot_transformed_image("test_images/20sWoman.jpg", netG, md.val_ds, tfms=x_tfms, sz=480)
vis.plot_transformed_image("test_images/egypt-1.jpg", netG, md.val_ds,sz=460)
vis.plot_transformed_image("test_images/Rutherford_Hayes.jpg", netG, md.val_ds, tfms=x_tfms, sz=400)
vis.plot_transformed_image("test_images/einstein_portrait.jpg", netG, md.val_ds, tfms=x_tfms, sz=400)
vis.plot_transformed_image("test_images/pinkerton.jpg", netG, md.val_ds, tfms=x_tfms, sz=450)
vis.plot_transformed_image("test_images/marilyn_woods.jpg", netG, md.val_ds, tfms=x_tfms, sz=460)
vis.plot_transformed_image("test_images/WaltWhitman.jpg", netG, md.val_ds, tfms=x_tfms, sz=280)
vis.plot_transformed_image("test_images/dorothea-lange.jpg", netG, md.val_ds, tfms=x_tfms,sz=460)
vis.plot_transformed_image("test_images/Hemmingway2.jpg", netG, md.val_ds, tfms=x_tfms, sz=510)
vis.plot_transformed_image("test_images/Chief.jpg", netG, md.val_ds, tfms=x_tfms,sz=520)
vis.plot_transformed_image("test_images/hemmingway.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/smoking_kid.jpg", netG, md.val_ds, tfms=x_tfms,sz=550)
vis.plot_transformed_image("test_images/teddy_rubble.jpg", netG, md.val_ds, tfms=x_tfms, sz=360)
vis.plot_transformed_image("test_images/dustbowl_2.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/camera_man.jpg", netG, md.val_ds, tfms=x_tfms,sz=520)
vis.plot_transformed_image("test_images/migrant_mother.jpg", netG, md.val_ds, tfms=x_tfms,sz=590)
vis.plot_transformed_image("test_images/marktwain.jpg", netG, md.val_ds, tfms=x_tfms, sz=530)
vis.plot_transformed_image("test_images/HelenKeller.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/Evelyn_Nesbit.jpg", netG, md.val_ds, tfms=x_tfms, sz=580)
vis.plot_transformed_image("test_images/Eddie-Adams.jpg", netG, md.val_ds, tfms=x_tfms, sz=460)
vis.plot_transformed_image("test_images/soldier_kids.jpg", netG, md.val_ds, tfms=x_tfms, sz=400)
vis.plot_transformed_image("test_images/AnselAdamsYosemite.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/unnamed.jpg", netG, md.val_ds, tfms=x_tfms, sz=550)
vis.plot_transformed_image("test_images/workers_canyon.jpg", netG, md.val_ds, tfms=x_tfms,sz=570)
vis.plot_transformed_image("test_images/CottonMill.jpg", netG, md.val_ds, tfms=x_tfms,sz=500)
vis.plot_transformed_image("test_images/JudyGarland.jpeg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/kids_pit.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/last_samurai.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/AnselAdamsWhiteChurch.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/opium.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/dorothea_lange_2.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/rgs.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/wh-auden.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/w-b-yeats.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/marilyn_portrait.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/wilson-slaverevivalmeeting.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/ww1_trench.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/women-bikers.png", netG, md.val_ds, tfms=x_tfms, sz=450)
vis.plot_transformed_image("test_images/Unidentified1855.jpg", netG, md.val_ds, tfms=x_tfms, sz=400)
vis.plot_transformed_image("test_images/skycrapper_lunch.jpg", netG, md.val_ds, tfms=x_tfms, sz=550)
vis.plot_transformed_image("test_images/sioux.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/school_kids.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/royal_family.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/redwood_lumberjacks.jpg", netG, md.val_ds, tfms=x_tfms, sz=550)
vis.plot_transformed_image("test_images/poverty.jpg", netG, md.val_ds, tfms=x_tfms,sz=550)
vis.plot_transformed_image("test_images/paperboy.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/NativeAmericans.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/helmut_newton-.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/Greece1911.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/FatMenClub.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/EgyptColosus.jpg", netG, md.val_ds, tfms=x_tfms, sz=460)
vis.plot_transformed_image("test_images/egypt-2.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/dustbowl_sd.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/dustbowl_people.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/dustbowl_5.jpg", netG, md.val_ds, tfms=x_tfms, sz=450)
vis.plot_transformed_image("test_images/dustbowl_1.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/DriveThroughGiantTree.jpg", netG, md.val_ds, tfms=x_tfms, sz=520)
vis.plot_transformed_image("test_images/covered-wagons-traveling.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/civil-war_2.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/civil_war_4.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/civil_war_3.jpg", netG, md.val_ds, tfms=x_tfms, sz=550)
vis.plot_transformed_image("test_images/civil_war.jpg", netG, md.val_ds, tfms=x_tfms, sz=540)
vis.plot_transformed_image("test_images/BritishSlum.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/bicycles.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/brooklyn_girls_1940s.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/40sCouple.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/1946Wedding.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/Dolores1920s.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/TitanicGym.jpg", netG, md.val_ds, tfms=x_tfms, sz=550)
vis.plot_transformed_image("test_images/FrenchVillage1950s.jpg", netG, md.val_ds, tfms=x_tfms, sz=440)
vis.plot_transformed_image("test_images/ClassDivide1930sBrittain.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/1870sSphinx.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/1890Surfer.png", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/TV1930s.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/1864UnionSoldier.jpg", netG, md.val_ds, tfms=x_tfms, sz=510)
vis.plot_transformed_image("test_images/1890sMedStudents.png", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/BellyLaughWWI.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/PiggyBackRide.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/HealingTree.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/ManPile.jpg", netG, md.val_ds, tfms=x_tfms, sz=450)
vis.plot_transformed_image("test_images/1910Bike.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/FreeportIL.jpg", netG, md.val_ds, tfms=x_tfms, sz=402)
vis.plot_transformed_image("test_images/DutchBabyCoupleEllis.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/InuitWoman1903.png", netG, md.val_ds, tfms=x_tfms, sz=460)
vis.plot_transformed_image("test_images/1920sDancing.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/AirmanDad.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/1910Racket.png", netG, md.val_ds, tfms=x_tfms, sz=540)
vis.plot_transformed_image("test_images/1880Paris.jpg", netG, md.val_ds, tfms=x_tfms, sz=520)
vis.plot_transformed_image("test_images/Deadwood1860s.png", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/1860sSamauris.png", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/LondonUnderground1860.jpg", netG, md.val_ds, tfms=x_tfms, sz=460)
vis.plot_transformed_image("test_images/Mid1800sSisters.jpg", netG, md.val_ds, tfms=x_tfms, sz=480)
vis.plot_transformed_image("test_images/1860Girls.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/SanFran1851.jpg", netG, md.val_ds, tfms=x_tfms, sz=480)
vis.plot_transformed_image("test_images/Kabuki1870s.png", netG, md.val_ds, tfms=x_tfms,sz=450)
vis.plot_transformed_image("test_images/Mormons1870s.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/EgyptianWomenLate1800s.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/PicadillyLate1800s.jpg", netG, md.val_ds, tfms=x_tfms, sz=540)
vis.plot_transformed_image("test_images/SutroBaths1880s.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/1880sBrooklynBridge.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/ChinaOpiumc1880.jpg", netG, md.val_ds, tfms=x_tfms,sz=500)
vis.plot_transformed_image("test_images/Locomotive1880s.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/ViennaBoys1880s.png", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/VictorianDragQueen1880s.png", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/Sami1880s.jpg", netG, md.val_ds, tfms=x_tfms,sz=420)
vis.plot_transformed_image("test_images/ArkansasCowboys1880s.jpg", netG, md.val_ds, tfms=x_tfms, sz=450)
vis.plot_transformed_image("test_images/Ballet1890Russia.jpg", netG, md.val_ds, tfms=x_tfms, sz=480)
vis.plot_transformed_image("test_images/Rottindean1890s.png", netG, md.val_ds, tfms=x_tfms, sz=540)
vis.plot_transformed_image("test_images/1890sPingPong.jpg", netG, md.val_ds, tfms=x_tfms,sz=500)
vis.plot_transformed_image("test_images/London1937.png", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/Harlem1932.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/OregonTrail1870s.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/EasterNyc1911.jpg", netG, md.val_ds, tfms=x_tfms, sz=532)
vis.plot_transformed_image("test_images/1899NycBlizzard.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/1916Sweeden.jpg", netG, md.val_ds, tfms=x_tfms, sz=520)
vis.plot_transformed_image("test_images/Edinburgh1920s.jpg", netG, md.val_ds, tfms=x_tfms, sz=480)
vis.plot_transformed_image("test_images/1890sShoeShopOhio.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/1890sTouristsEgypt.png", netG, md.val_ds, tfms=x_tfms, sz=570)
vis.plot_transformed_image("test_images/1938Reading.jpg", netG, md.val_ds, tfms=x_tfms, sz=455)
vis.plot_transformed_image("test_images/1850Geography.jpg", netG, md.val_ds, tfms=x_tfms, sz=540)
vis.plot_transformed_image("test_images/1901Electrophone.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/Texas1938Woman.png", netG, md.val_ds, tfms=x_tfms, sz=520)
vis.plot_transformed_image("test_images/MaioreWoman1895NZ.jpg", netG, md.val_ds, tfms=x_tfms, sz=520)
vis.plot_transformed_image("test_images/WestVirginiaHouse.jpg", netG, md.val_ds, tfms=x_tfms, sz=520)
vis.plot_transformed_image("test_images/1920sGuadalope.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/1909Chicago.jpg", netG, md.val_ds, tfms=x_tfms, sz=540)
vis.plot_transformed_image("test_images/1920sFarmKid.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/ParisLate1800s.jpg", netG, md.val_ds, tfms=x_tfms,sz=410)
vis.plot_transformed_image("test_images/1900sDaytonaBeach.png", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/1930sGeorgia.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/NorwegianBride1920s.jpg", netG, md.val_ds, tfms=x_tfms, sz=550)
vis.plot_transformed_image("test_images/Depression.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/1888Slum.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/LivingRoom1920Sweeden.jpg", netG, md.val_ds, tfms=x_tfms, sz=540)
vis.plot_transformed_image("test_images/1896NewsBoyGirl.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/PetDucks1927.jpg", netG, md.val_ds, tfms=x_tfms, sz=400)
vis.plot_transformed_image("test_images/1899SodaFountain.jpg", netG, md.val_ds, tfms=x_tfms, sz=640)
vis.plot_transformed_image("test_images/TimesSquare1955.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/PuppyGify.jpg", netG, md.val_ds, tfms=x_tfms, sz=510)
vis.plot_transformed_image("test_images/1890CliffHouseSF.jpg", netG, md.val_ds, tfms=x_tfms, sz=590)
vis.plot_transformed_image("test_images/1908FamilyPhoto.jpg", netG, md.val_ds, tfms=x_tfms, sz=520)
vis.plot_transformed_image("test_images/1900sSaloon.jpg", netG, md.val_ds, tfms=x_tfms, sz=560)
vis.plot_transformed_image("test_images/1890BostonHospital.jpg", netG, md.val_ds, tfms=x_tfms,sz=500)
vis.plot_transformed_image("test_images/1870Girl.jpg", netG, md.val_ds, tfms=x_tfms, sz=450)
vis.plot_transformed_image("test_images/AustriaHungaryWomen1890s.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/Shack.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/Apsaroke1908.png", netG, md.val_ds, tfms=x_tfms, sz=520)
vis.plot_transformed_image("test_images/1948CarsGrandma.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/PlanesManhattan1931.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/WorriedKid1940sNyc.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/1920sFamilyPhoto.jpg", netG, md.val_ds, tfms=x_tfms, sz=550)
vis.plot_transformed_image("test_images/CatWash1931.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/1940sBeerRiver.jpg", netG, md.val_ds, tfms=x_tfms, sz=400)
vis.plot_transformed_image("test_images/VictorianLivingRoom.jpg", netG, md.val_ds, tfms=x_tfms, sz=560)
vis.plot_transformed_image("test_images/1897BlindmansBluff.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/1874Mexico.png", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/MadisonSquare1900.jpg", netG, md.val_ds, tfms=x_tfms, sz=450)
vis.plot_transformed_image("test_images/1867MusicianConstantinople.jpg", netG, md.val_ds, tfms=x_tfms, sz=550)
vis.plot_transformed_image("test_images/1925Girl.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/1907Cowboys.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/WWIIPeeps.jpg", netG, md.val_ds, tfms=x_tfms, sz=460)
vis.plot_transformed_image("test_images/BabyBigBoots.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/1895BikeMaidens.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/IrishLate1800s.jpg", netG, md.val_ds, tfms=x_tfms,sz=500)
vis.plot_transformed_image("test_images/LibraryOfCongress1910.jpg", netG, md.val_ds, tfms=x_tfms,sz=520)
vis.plot_transformed_image("test_images/1875Olds.jpg", netG, md.val_ds, tfms=x_tfms, sz=520)
vis.plot_transformed_image("test_images/SenecaNative1908.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/WWIHospital.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/GreekImmigrants1905.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/1892WaterLillies.jpg", netG, md.val_ds, tfms=x_tfms, sz=560)
vis.plot_transformed_image("test_images/GreekImmigrants1905.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/FatMensShop.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/KidCage1930s.png", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/FarmWomen1895.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/NewZealand1860s.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/JerseyShore1905.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/LondonKidsEarly1900s.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/NYStreetClean1906.jpg", netG, md.val_ds, tfms=x_tfms, sz=520)
vis.plot_transformed_image("test_images/Boston1937.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/Cork1905.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/BoxedBedEarly1900s.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/ZoologischerGarten1898.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/EmpireState1930.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/Agamemnon1919.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/AppalachianLoggers1901.jpg", netG, md.val_ds, tfms=x_tfms,sz=550)
vis.plot_transformed_image("test_images/WWISikhs.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/MementoMori1865.jpg", netG, md.val_ds, tfms=x_tfms, sz=500)
vis.plot_transformed_image("test_images/RepBrennanRadio1922.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/Late1800sNative.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/GasPrices1939.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/1933RockefellerCenter.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/Scotland1919.jpg", netG, md.val_ds, tfms=x_tfms)
vis.plot_transformed_image("test_images/SchoolDance1956.jpg", netG, md.val_ds, tfms=x_tfms)
| ColorizeVisualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Histogram Objects
import quasar
# ## Introduction
# A fundamental operation encountered in qubit hardware is the process of measurement. Generally, one prepares a quantum state by running a quantum circuit and then projectively measures each qubit to determine if that qubit ends up in the $|0\rangle$ or $|1\rangle$ state. The collective measurement result over all qubits is a binary string, e.g., $|0011\rangle$, which we refer to as a `Ket`. This procedure is repeated `nmeasurement` times, with different `Ket`s appearing probabilistically according to the square of the amplitude of each `Ket` in the quantum state. The fundamental output from this overall process is a histogram of `Ket` strings observed together with the the probability that each one was observed and the total number of measurements. We refer to this overall output as a `ProbabilityHistogram` object. An alternative output in terms of histogram of `Ket` strings observed together with the the integral number of times that each one was observed is provided as the `CountHistogram` object - utility functions exist to quickly convert back and forth between `ProbabilityHistogram` and `CountHistogram`.
#
# **Naming:** The names of these two classes were selected after much debate. For `Ket`, we could have used `int` with a fixed endian condition, `str` which we would trust the user to always be composed of `0` and `1`, or a class which could alternatively be named `Ket`, `String`, or `Configuration`. We decided to use `Ket` to force the order, typing, and correctness of this concept, and to make the indexing of `ProbabilityHistogram` and `CountHistogram` also support direct indexing by `str` (cast to `Ket` under the hood) to make things easy. For `ProbabilityHistogram`, we could have used a `dict` or other map, or a class which could alternatively be named `Histogram`, `Counts`, `Shots`, `Probabilities` or any variation thereof. We decided to use `MeasurementResult` to keep things maximally explicit. Note that most users will encounter these objects as the output of `quasar` library methods, and will rarely need to explicitly construct `Ket`, `ProbabilityHistogram`, or `CountHistogram` objects. In this case, the manipulation of these data structures seems straightforward, and allows for easy casting to other data structures that the user might prefer.
# ## The ProbabilityHistogram and CountHistogram Objects
# To standardize the representation of the full histogram of results of a complete measurement process, `quasar` provides the `ProbabilityHistogram` object,
probabilities = quasar.ProbabilityHistogram(
nqubit=4,
histogram={
3 : 0.2, # 3 is 0011
12 : 0.8, # 12 is 1100
},
nmeasurement=1000,
)
print(probabilities)
# `ProbabilityHistogram` supports read-only indexing:
print(probabilities[3])
# For convenience, you can also use `str` objects to index a `ProbabilityHistogram`:
print(probabilities['0011'])
# The total number of measurements is provided in the `nmeasurement` attribute:
print(probabilities.nmeasurement)
# The number of qubits is provided in the `nqubit` attribute:
print(probabilities.nqubit)
# Many users are more familiar with the finite integer counts of each `Ket`, as opposed to the probabilities of each `Ket`. To deal with the former, we provide the utility to convert from a `ProbabilityHistogram` (based on floating point probabilities) to a `CountHistogram` (based on integral counts):
counts = probabilities.to_count_histogram()
print(counts)
print(counts['0011'])
# One can also convert the other direction from a `CountHistogram` to a `ProbabilityHistogram`:
probabilities2 = counts.to_probability_histogram()
print(probabilities2)
print(probabilities2['0011'])
# ## Infinite Sampling
# In working computations, the `ProbabilityHistogram` is preferred, as it allows for conceptually ideal infinite statistical sampling (indicated by `nmeasurement=None` to indicate infinite `nmeasurement`). No corresponding `CountHistogram` is valid (as all the counts would be infinite), so an error is thrown if one tries to convert an infinitely-sampled `ProbabilityHistogram` to `CountHistogram`:
probabilities = quasar.ProbabilityHistogram(
nqubit=4,
histogram={
3 : 0.2, # 3 is 0011
12 : 0.8, # 12 is 1100
},
nmeasurement=None,
)
print(probabilities)
# Throws due to infinite sampling
# counts = probabilities.to_count_histogram()
# ## Measurement in Action
# Here we show the main place a user would encounter these objects: calling `run_measurement` to sample the output of a given quantum circuit:
circuit = quasar.Circuit().H(0).CX(0,1).CX(1,2)
print(circuit)
# Here is an example with finite `nmeasurement`, which returns as a `ProbabilityHistogram` and is convertible to a `CountHistogram`:
backend = quasar.QuasarSimulatorBackend()
probabilities = backend.run_measurement(circuit, nmeasurement=1000)
print(probabilities)
counts = probabilities.to_count_histogram()
print(counts)
# Here is an example with infinite `nmeasurement`, which returns as a `ProbabilityHistogram` and is **not** convertible to a `CountHistogram`:
backend = quasar.QuasarSimulatorBackend()
probabilities = backend.run_measurement(circuit, nmeasurement=None)
print(probabilities)
# Throws due to infinite sampling
# counts = probabilities.to_count_histogram()
# print(counts)
import quasar
| notebooks/quasar-measurement.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Run the following:
#Run this from command line
# !python -c "import this"
#Inside a python console
import this
# ## From official website ( http://www.python.org/ ):
#
# Python is a programming language that lets you work more quickly and integrate
# your systems more effectively. You can learn to use Python and see almost
# immediate gains in productivity and lower maintenance costs.
#
# ## Executive summary from official website ( http://www.python.org/doc/essays/blurb.html )
#
# Python is an interpreted, object-oriented, high-level programming language
# with dynamic semantics. Its high-level built in data structures, combined
# with dynamic typing and dynamic binding, make it very attractive for Rapid
# Application Development, as well as for use as a scripting or glue language
# to connect existing components together. Python's simple, easy to learn syntax
# emphasizes readability and therefore reduces the cost of program maintenance.
# Python supports modules and packages, which encourages program modularity and
# code reuse. The Python interpreter and the extensive standard library are
# available in source or binary form without charge for all major platforms,
# and can be freely distributed.
#
# ### TO SUM UP
# - quick development
# - simple, readable, easy to learn syntax
# - general purpose
# - interpreted (not compiled)
# - object-oriented
# - high-level
# - dynamic semantics (aka execution semantics)
# - fully dynamic typing
# - dynamic binding
# - low programs manteinance cost
# - modularity and code reuse
# - no licensing costs
# - extensive standard library, "batteries included"
# - imperative and functional programming
# - automatic memory management
# execution semantics
for i in range(10):
print i**2
print "Outside for loop"
# dynamic binding
my_str = "hola"
type(my_str)
# dynamic binding
my_str = 90
type(my_str)
# fully dynamic typing
4 +5 +6
# fully dynamic typing
4 + "hola"
# ## HISTORY
# - http://docs.python.org/2/license.html
# - http://en.wikipedia.org/wiki/History_of_Python
# - http://en.wikipedia.org/wiki/Benevolent_Dictator_For_Life
# - http://www.python.org/dev/peps/pep-0001/
# - http://docs.python.org/3.0/whatsnew/3.0.html
# ## TOOLS
#
# **CPython** is the real name of default standard Python implementation
# - The interpreter is deployed together with standard library
# - It can take a file as argument to run. Otherwise it opens and interactive console
# - With '-m' parameter you can execute directly certain modules (debugging, profiling)
# - Implemented in C
#
# There are Python implementations in other languages:
# - Jython: Python 2.5 interpreter written in Java which runs bytecode in the JVM
# - IronPython: Similar approach for .NET Common Language Runtime
# - JS, C++, CIL...
# - Stackless Python: CPython fork with microthreads concurrency
# - PyPy: Python 2.7 interpreter implemented in Python. Really fast, multi-core...!!!
#
#
# **IPython**: create a comprehensive environment for interactive and exploratory computing
# An enhanced interactive Python shell
# An architecture for interactive parallel computing:
# - Other powerful interactive shells (terminal and Qt-based)
# - A browser-based notebook with support for code, text, mathematical expressions
# inline plots and other rich media
# - Support for interactive data visualization and use of GUI toolkits
# - Flexible, embeddable interpreters to load into your own projects
# - Easy to use, high performance tools for parallel computing
# - Recently funded with $1.15M from the Alfred P. Sloan Foundation
#
#
# **virtualenv**: a tool to create isolated Python environments
# It simply changes your PATH environment var to point to a different folder
#
#
# **PyPi**: The Python Package Index is a repository of software for the Python programming language.
# There are currently 89324 packages here.
# The packages are 'eggs'.
#
# **pip**: A tool for installing and managing Python packages
# It installs packages from PyPi, local folders or Git and other repositories
# It can read a list of packages from a file or generate the list of installed packages
# ### Installing pip and virtualenv for a python interpreter
#
# ```sh
# wget https://bootstrap.pypa.io/get-pip.py
# python get-pip.py
# pip install virtualenv
# virtualenv env
# source env/bin/activate
# # finish virtualenv running deactivate
# ```
#
# **IDE**?
# - PyCharm (we have licenses, ask EPG)
# - Eclipse + Pydev
# - NetBeans
# - Eric
# - NINJA IDE
# - Aptana Studio 3
# - SPE
# - Python's IDLE (not recommendable at all)
# - ...
# - Emacs
# - Vi
# http://wiki.python.org/moin/IntegratedDevelopmentEnvironments
# Lots of good IDEs, it's up to you!
| basic/0_What_is_python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Pandas
#
# Pandas is a library providing high-performance, easy-to-use data structures and data analysis tools. The core of pandas is its *dataframe* which is essentially a table of data. Pandas provides easy and powerful ways to import data from a variety of sources and export it to just as many. It is also explicitly designed to handle *missing data* elegantly which is a very common problem in data from the real world.
#
# The offical [pandas documentation](http://pandas.pydata.org/pandas-docs/stable/) is very comprehensive and you will be answer a lot of questions in there, however, it can sometimes be hard to find the right page. Don't be afraid to use Google to find help.
# Pandas has a standard convention for importing it which you will see used in a lot of documentation so we will follow that in this course:
import pandas as pd
from pandas import Series, DataFrame
# ## Series
#
# The simplest of pandas' data structures is the `Series`. It is a one-dimensional list-like structure.
# Let's create one from a `list`:
Series([14, 7, 3, -7, 8])
# There are three main components to this output.
# The first column (`0`, `2`, etc.) is the index, by default this is numbers each row starting from zero.
# The second column is our data, stored i the same order we entered it in our list.
# Finally at the bottom there is the `dtype` which stands for 'data type' which is telling us that all our data is being stored as a 64-bit integer.
# Usually you can ignore the `dtype` until you start doing more advanced things.
#
# In the first example above we allowed pandas to automatically create an index for our `Series` (this is the `0`, `1`, `2`, etc. in the left column) but often you will want to specify one yourself
s = Series([14, 7, 3, -7, 8], index=['a', 'b', 'c', 'd', 'e'])
print(s)
# We can use this index to retrieve individual rows
s['a']
# to replace values in the series
s['c'] = -1
# or to get a set of rows
s[['a', 'c', 'd']]
# ### Exercise 1
#
# - Create a Pandas `Series` with 10 or so elements where the indices are years and the values are numbers.
# - Experiment with retrieving elements from the `Series`.
# - Try making another `Series` with duplicate values in the index, what happens when you access those elements?
# - How does a Pandas `Series` differ from a Python `list` or `dict`?
# ## Series operations
#
# A `Series` is `list`-like in the sense that it is an ordered set of values. It is also `dict`-like since its entries can be accessed via key lookup. One very important way in which is differs is how it allows operations to be done over the whole `Series` in one go, a technique often referred to as 'broadcasting'.
#
# A simple example is wanting to double the value of every entry in a set of data. In standard Python, you might have a list like
my_list = [3, 6, 8, 4, 10]
# If you wanted to double every entry you might try simply multiplying the list by `2`:
my_list * 2
# but as you can see, that simply duplicated the elements. Instead you would have to use a `for` loop or a list comprehension:
[i * 2 for i in my_list]
# With a pandas `Series`, however, you can perform bulk mathematical operations to the whole series in one go:
my_series = Series(my_list)
print(my_series)
my_series * 2
# As well as bulk modifications, you can perform bulk selections by putting more complex statements in the square brackets:
s[s < 0] # All negative entries
s[(s * 2) > 4] # All entries which, when doubled are greater than 4
# These operations work because the `Series` index selection can be passed a series of `True` and `False` values which it then uses to filter the result:
(s * 2) > 4
# Here you can see that the rows `a`, `b` and `e` are `True` while the others are `False`. Passing this to `s[...]` will only show rows that are `True`.
# ### Multi-Series operations
#
# It is also possible to perform operations between two `Series` objects:
s2 = Series([23,5,34,7,5])
s3 = Series([7, 6, 5,4,3])
s2 - s3
# ### Exercise 2
#
# - Create two `Series` objects of equal length with no specified index and containing any values you like. Perform some mathematical operations on them and experiment to make sure it works how you think.
# - What happens then you perform an operation on two series which have different lengths? How does this change when you give the series some indices?
# - Using the `Series` from the first exercise with the years for the index, Select all entries with even-numbered years. Also, select all those with odd-numbered years.
# ## DataFrame
#
# While you can think of the `Series` as a one-dimensional list of data, pandas' `DataFrame` is a two (or possibly more) dimensional table of data. You can think of each column in the table as being a `Series`.
data = {'city': ['Paris', 'Paris', 'Paris', 'Paris',
'London', 'London', 'London', 'London',
'Rome', 'Rome', 'Rome', 'Rome'],
'year': [2001, 2008, 2009, 2010,
2001, 2006, 2011, 2015,
2001, 2006, 2009, 2012],
'pop': [2.148, 2.211, 2.234, 2.244,
7.322, 7.657, 8.174, 8.615,
2.547, 2.627, 2.734, 2.627]}
df = DataFrame(data)
# This has created a `DataFrame` from the dictionary `data`. The keys will become the column headers and the values will be the values in each column. As with the `Series`, an index will be created automatically.
df
# Or, if you just want a peek at the data, you can just grab the first few rows with:
df.head(3)
# Since we passed in a dictionary to the `DataFrame` constructor, the order of the columns will not necessarilly match the order in which you defined them. To enforce a certain order, you can pass a `columns` argument to the constructor giving a list of the columns in the order you want them:
DataFrame(data, columns=['year', 'city', 'pop'])
# When we accessed elements from a `Series` object, it would select an element by row. However, by default `DataFrame`s index primarily by column. You can access any column directly by using square brackets or by named attributes:
df['year']
df.city
# Accessing a column like this returns a `Series` which will act in the same way as those we were using earlier.
#
# Note that there is one additional part to this output, `Name: city`. Pandas has remembered that this `Series` was created from the `'city'` column in the `DataFrame`.
type(df.city)
df.city == 'Paris'
# This has created a new `Series` which has `True` set where the city is Paris and `False` elsewhere.
#
# We can use filtered `Series` like this to filter the `DataFrame` as a whole. `df.city == 'Paris'` has returned a `Series` containing booleans. Passing it back into `df` as an indexing operation will use it to filter based on the `'city'` column.
df[df.city == 'Paris']
# You can then carry on and grab another column after that filter:
df[df.city == 'Paris'].year
# If you want to select a **row** from a `DataFrame` then you can use the `.loc` attribute which allows you to pass index values like:
df.loc[2]
df.loc[2]['city']
# ## Adding new columns
#
# New columns can be added to a `DataFrame` simply by assigning them by index (as you would for a Python `dict`) and can be deleted with the `del` keyword in the same way:
df['continental'] = df.city != 'London'
df
del df['continental']
# ### Exercise 3
#
# - Create the `DataFrame` containing the census data for the three cities.
# - Select the data for the year 2001. Which city had the smallest population that year?
# - Find all the cities which had a population smaller than 2.6 million.
# ## Reading from file
#
# One of the most common situations is that you have some data file containing the data you want to read. Perhaps this is data you've produced yourself or maybe it's from a collegue. In an ideal world the file will be perfectly formatted and will be trivial to import into pandas but since this is so often not the case, it provides a number of features to make your life easier.
#
# Full information on reading and writing is available in the pandas manual on [IO tools](http://pandas.pydata.org/pandas-docs/stable/io.html) but first it's worth noting the common formats that pandas can work with:
# - Comma separated tables (or tab-separated or space-separated etc.)
# - Excel spreadsheets
# - HDF5 files
# - SQL databases
#
# For this course we will focus on plain-text CSV files as they are perhaps the most common format. Imagine we have a CSV file like (you can download this file from [city_pop.csv](https://raw.githubusercontent.com/milliams/data_analysis_python/master/city_pop.csv)):
# ! cat data/city_pop.csv # Uses the IPython 'magic' !cat to print the file
# We can use the pandas function `read_csv()` to read the file and convert it to a `DataFrame`. Full documentation for this function can be found in [the manual](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_csv.html) or, as with any Python object, directly in the notebook by putting a `?` after the name:
help(pd.read_csv)
pd.read_csv('data/city_pop.csv')
# We can see that by default it's done a fairly bad job of parsing the file (this is mostly because I've constructed the `city_pop.csv` file to be as obtuse as possible). It's making a lot of assumptions about the structure of the file but in general it's taking quite a naïve approach.
#
# The first this we notice is that it's treating the text at the top of the file as though it's data. Checking [the documentation](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_csv.html) we see that the simplest way to solve this is to use the `skiprows` argument to the function to which we give an integer giving the number of rows to skip:
pd.read_csv(
'data/city_pop.csv',
skiprows=5,
)
# The next most obvious problem is that it is not separating the columns at all. This is controlled by the `sep` argument which is set to `','` by default (hence *comma* separated values). We can simply set it to the appropriate semi-colon:
pd.read_csv(
'data/city_pop.csv',
skiprows=5,
sep=';'
)
# Reading the descriptive header of our data file we see that a value of `-1` signifies a missing reading so we should mark those too. This can be done after the fact but it is simplest to do it at import-time using the `na_values` argument:
pd.read_csv(
'data/city_pop.csv',
skiprows=5,
sep=';',
na_values='-1'
)
# The last this we want to do is use the `year` column as the index for the `DataFrame`. This can be done by passing the name of the column to the `index_col` argument:
df3 = pd.read_csv(
'data/city_pop.csv',
skiprows=5,
sep=';',
na_values='-1',
index_col='year'
)
df3
# ### Exercise 4
#
# - Alongside `data/city_pop.csv` there is another file called `data/cetml1659on.dat`. This contains some historical weather data for a location in the UK. Import that file as a Pandas `DataFrame` using `read_csv()`, making sure that you cover all the NaN values.
# - How many years had a negative average temperature in January?
# - What was the average temperature in June over the years in the data set? Tip: look in the [documentation](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.html) for which method to call.
#
# We will come back to this data set in a later stage.
| 12_pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.12 64-bit (''earth'': conda)'
# name: python3
# ---
# This notebook runs multiple linear regression model to predict the albedo from different combination of bands
import pandas as pd
import numpy as np
import os
import glob
import matplotlib.pyplot as plt
from sklearn import linear_model
from sklearn.model_selection import train_test_split
from scipy import stats
import seaborn as sns
import altair as alt
import plotly.express as px
# ## merge promice data to one dataframe
df = pd.read_csv('promice/promice.csv')
df['Longitude'] = df['Longitude'] * -1
# +
folderpath = "promice/multiSat90m"
searchCriteria = "*.csv"
globInput = os.path.join(folderpath, searchCriteria)
csvPath = glob.glob(globInput)
csvList = os.listdir(folderpath)
# -
# hourly
for i in range(len(csvList)):
# promice data
stationName = os.path.splitext(csvList[i])[0].replace("-", "*")
index = df.index[df.Station == stationName][0]
url = df.urlhourly[index]
dfs = pd.read_table(url, sep=r'\s{1,}', engine='python')
dfs = dfs[['Albedo_theta<70d', 'LatitudeGPS(degN)', 'LongitudeGPS(degW)', 'Year', 'MonthOfYear','DayOfMonth', 'HourOfDay(UTC)', 'CloudCover']]
dfs = dfs.replace(-999, np.nan)
dfs['lon'] = dfs['LongitudeGPS(degW)'].interpolate(method='linear',limit_direction='both') * -1
dfs['lat'] = dfs['LatitudeGPS(degN)'].interpolate(method='linear',limit_direction='both')
dfs['datetime'] = pd.to_datetime(dict(year=dfs.Year, month=dfs.MonthOfYear, day = dfs.DayOfMonth, hour = dfs['HourOfDay(UTC)']))
# cloud cover less than 50% and albedo must be valid value
dfs = dfs[(dfs['Albedo_theta<70d'] > 0) & (dfs['CloudCover'] < 0.5)]
dfs['Station'] = stationName
# satellite data
dfr = pd.read_csv(csvPath[i])
dfr = dfr.dropna(how='all', subset=['Blue', 'Green', 'Red', 'NIR', 'SWIR1', 'SWIR2'])
# dfr.datetime = pd.to_datetime(dfr.datetime).dt.date # keep only ymd
dfr.datetime = pd.to_datetime(dfr.datetime)
# join by datetime
dfmerge = pd.merge_asof(dfr.sort_values('datetime'), dfs, on='datetime',allow_exact_matches=False, tolerance=pd.Timedelta(hours=1),direction='nearest' )
# dfmerge = pd.merge_asof(dfr.sort_values('datetime'), dfs, on='datetime', tolerance=pd.Timedelta(hours=1) )
if i==0:
dfmerge.to_csv('promice vs satellite90m.csv', mode='w', index=False)
else:
dfmerge.to_csv('promice vs satellite90m.csv', mode='a', index=False, header=False)
# ## Multiple Lienar Regression: PROMICE VS Satellite data
dfmlr = pd.read_csv("promice vs satellite90m.csv")
# ProfileReport(df)
dfmlr = dfmlr[(dfmlr['MonthOfYear']>4) & (dfmlr['MonthOfYear']<10)] # (df['MonthOfYear']!=7
# dfmlr = dfmlr['Blue', 'Green', 'Red', 'NIR', 'SWIR1', 'SWIR2', 'Albedo_theta<70d']
# df = df[df['Albedo_theta<70d']<0.9]
fig, ax = plt.subplots(figsize=(6, 4))
sns.set_style("darkgrid")
sns.boxplot(data=dfmlr[['Blue', 'Green', 'Red', 'NIR', 'SWIR1', 'SWIR2', 'Albedo_theta<70d']], palette=['b', 'g', 'r', 'c', 'm', 'y', 'w'])
plt.xticks(rotation=45)
# boxplot = df.boxplot(column=['Blue', 'Green', 'Red', 'NIR', 'SWIR1', 'SWIR2', 'Albedo_theta<70d'], rot=45)
fig.savefig("print/boxplotSatPromice90.pdf", dpi=300, bbox_inches="tight")
sns.set_style("darkgrid")
dfmlr.Station.value_counts().plot(kind='bar')
plt.savefig("print/primiceHist90.pdf", dpi=300)
# +
# '''total band'''
# dfmlr = dfmlr.dropna(how='any', subset=['Blue', 'Green', 'Red', 'NIR', 'SWIR1', 'SWIR2', 'Albedo_theta<70d'])
# X = dfmlr[['Blue', 'Green', 'Red', 'NIR', 'SWIR1', 'SWIR2']]
# y = dfmlr['Albedo_theta<70d']
# X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
# ols = linear_model.LinearRegression()
# model = ols.fit(X_train, y_train)
# response = model.predict(X_test)
# r2 = model.score(X_test, y_test)
# '''viz band'''
# dfmlr = dfmlr.dropna(how='any', subset=['Blue', 'Green', 'Red', 'Albedo_theta<70d'])
# X = dfmlr[['Blue', 'Green', 'Red']]
# y = dfmlr['Albedo_theta<70d']
# X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
# ols = linear_model.LinearRegression()
# model = ols.fit(X_train, y_train)
# response = model.predict(X_test)
# r2 = model.score(X_test, y_test)
'''viz-nir band'''
dfmlr = dfmlr.dropna(how='any', subset=['Blue', 'Green', 'Red', 'NIR', 'Albedo_theta<70d'])
X = dfmlr[['Blue', 'Green', 'Red', 'NIR']]
y = dfmlr['Albedo_theta<70d']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
ols = linear_model.LinearRegression()
model = ols.fit(X_train, y_train)
response = model.predict(X_test)
r2 = model.score(X_test, y_test)
# -
print('R\N{SUPERSCRIPT TWO}: %.4f' % r2)
print(model.coef_)
# print("coefficients: Blue: %.4f, Green: %.4f, Red: %.4f, NIR: %.4f, SWIR1: %.4f, SWIR2: %.4f" %(model.coef_[0], model.coef_[1], model.coef_[2], model.coef_[3], model.coef_[4], model.coef_[5]))
# print("coefficients: Blue: %.4f, Red: %.4f, NIR: %.4f, SWIR1: %.4f, SWIR2: %.4f" %(model.coef_[0], model.coef_[1], model.coef_[2], model.coef_[3], model.coef_[4]))
print("intercept: %.4f" % model.intercept_)
len(dfmlr)
# +
colors = ['Positive' if c > 0 else 'Negative' for c in model.coef_]
figMLR = px.bar(
x=X.columns, y=model.coef_, color=colors,
color_discrete_sequence=['red', 'blue'],
labels=dict(x='band', y='Linear coefficient'),
title='Weight of each band for predicting albedo'
)
figMLR.show()
# figMLR.write_image("print/MLRcoefficient.jpg")
# +
fig, ax = plt.subplots(figsize=(8, 8))
plt.xlim(0, 1)
plt.ylim(0, 1)
sns.set_theme(style="darkgrid", font="Arial", font_scale=2)
# sns.set_theme(color_codes=True)
sns.scatterplot(x=response, y=y_test, s=20)
sns.regplot(x=response, y=y_test, scatter=False, color='red',)
ax.set_aspect('equal', 'box')
# sns.histplot(x=response, y=y, bins=50, pthresh=.1, cmap="viridis", cbar=True, cbar_kws={'label': 'frequency'})
# sns.kdeplot(x=response, y=y, levels=5, color="w", linewidths=1)
# ax.set(xlabel='Predicted Albedo (total bands)', ylabel='Albedo PROMICE')
# fig.savefig('print/totalMLR90.jpg', dpi=300, bbox_inches="tight")
# ax.set(xlabel='Predicted Albedo (vis bands)', ylabel='Albedo PROMICE')
# fig.savefig('print/visMLR90.jpg', dpi=300, bbox_inches="tight")
ax.set(xlabel='Predicted Albedo (vis-nir bands)', ylabel='Albedo PROMICE')
fig.savefig('print/visnirMLR90.jpg', dpi=300, bbox_inches="tight")
# +
dfmlr['response'] = response
alt.data_transformers.disable_max_rows() # this should be avoided but now let's disable the limit
alt.Chart(dfmlr).mark_circle().encode(
x='response',
y='Albedo_theta<70d',
color='Station',
tooltip=['datetime:T','Station','response','Albedo_theta<70d']
).interactive()
# chart + chart.transform_regression('x', 'y').mark_line()
# +
dfmlr['response'] = response
alt.data_transformers.disable_max_rows() # this should be avoided but now let's disable the limit
brush = alt.selection(type='interval')
points = alt.Chart(dfmlr).mark_circle().encode(
x='response',
y='Albedo_theta<70d',
color=alt.condition(brush, 'Station:O', alt.value('grey')),
tooltip=['datetime:T','Station','response','Albedo_theta<70d']
).add_selection(brush)
# Base chart for data tables
ranked_text = alt.Chart(dfmlr).mark_text().encode(
y=alt.Y('row_number:O',axis=None)
).transform_window(
row_number='row_number()'
).transform_filter(
brush
).transform_window(
rank='rank(row_number)'
).transform_filter(
alt.datum.rank<40
)
# Data Tables
stationalt = ranked_text.encode(text='Station').properties(title='station')
albedoalt = ranked_text.encode(text='Albedo_theta<70d:N').properties(title='Albedo')
predictedalt = ranked_text.encode(text='response:N').properties(title='predicted albedo')
timealt = ranked_text.encode(text='datetime:T').properties(title='time')
text = alt.hconcat(stationalt, albedoalt, predictedalt, timealt) # Combine data tables
# Build chart
alt.hconcat(
points,
text
).resolve_legend(
color="independent"
)
# chart + chart.transform_regression('x', 'y').mark_line()
# -
# # Liang et al. and Naegeli et al.
# +
albedo = 0.356 * dfmlr.Blue + 0.13 * dfmlr.Red + 0.373 * dfmlr.NIR + 0.085 * dfmlr.SWIR1 + 0.072 * dfmlr.SWIR2 - 0.018
slope, intercept, r_value, p_value, std_err = stats.linregress(y_test, albedo)
fig1, ax1 = plt.subplots(figsize=(8, 8))
# plt.sca(ax1)
sns.set_theme(style="darkgrid", font="Arial", font_scale=2)
sns.scatterplot(x=albedo, y=y_test, s=20)
sns.regplot(x=albedo, y=y_test, scatter=False, color='red',)
plt.xlim(0, 1)
plt.ylim(0, 1)
ax1.set(xlabel='Predicted Albedo', ylabel='Albedo PROMICE')
ax1.set_aspect('equal', 'box')
# sns.histplot(x=response, y=y, bins=50, pthresh=.1, cmap="viridis", cbar=True, cbar_kws={'label': 'frequency'})
# sns.kdeplot(x=response, y=y, levels=5, color="w", linewidths=1)
fig1.savefig('print/liang90.jpg', dpi=300, bbox_inches="tight")
| script/Promice vs satellite.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LA Restaurant Market and Health Data
# +
# import dependencies
import pandas as pd
import os
import numpy as np
os.getcwd()
from splinter import Browser
from bs4 import BeautifulSoup
from urllib.parse import urlencode
import pymongo
from pymongo import MongoClient
client = MongoClient()
# +
# pulling datasets
inspections_data = pd.read_csv('restaurant-and-market-health-inspections.csv')
violations_data = pd.read_csv('restaurant-and-market-health-violations.csv')
# +
# function to embed violations data into inspections data
def transform_inspection(inspection, violations_df, facilities_df):
VIOLATIONS_COLUMNS = ['row_id', 'violation_code', 'violation_description', 'violation_status', 'points']
serial_number = inspection['serial_number']
violations_dict = violations_df.loc[violations_df['serial_number'] == serial_number, VIOLATIONS_COLUMNS].to_dict(orient='records')
result = inspection.to_dict()
result['violations'] = violations_dict
result['violations_count'] = len(violations_dict)
rating = facilities_df.loc[(facilities_df['facility_name'] == inspections_data.iloc[0]['facility_name']) & (facilities_df['facility_address'] == inspections_data.iloc[0]['facility_address'])].loc[0]['rating']
if not np.isnan(rating):
result['rating'] = rating
return result
# -
# # Scrap Yelp For Ratings
# !which chromedriver
# +
executable_path = {'executable_path': '/usr/local/bin/chromedriver'}
browser = Browser('chrome', **executable_path, headless=False)
def get_url(restaurant_name):
url_prefix = 'https://www.yelp.com/search?'
params = { 'find_desc': restaurant_name, 'find_loc': 'Los Angeles,CA' }
param_string = urlencode(params)
return url_prefix + param_string
def is_matching_restaurant(name, address, scraped_name, scraped_address):
name = name.lower()
address = address.lower()
scraped_name = scraped_name.lower()
scraped_address = scraped_address.lower()
is_name_match = name.startswith(scraped_name) or scraped_name.startswith(name)
is_address_match = address.startswith(scraped_address) or scraped_address.startswith(address)
return is_name_match and is_address_match
def scrape_rating(html, name, address):
soup = BeautifulSoup(html, 'html.parser')
restaurants_li = soup.find_all('li')
for restaurant_li in restaurants_li:
try:
scraped_name = restaurant_li.h3.a['name']
scraped_address = restaurant_li.address.span.text
if (is_matching_restaurant(name, address, scraped_name, scraped_address)):
return float(restaurant_li.find('div', role="img")['aria-label'].replace(' star rating', ''))
except Exception as e:
{}
facilities_df = inspections_data[['facility_name', 'facility_address']].drop_duplicates()
facilities_df['rating'] = pd.Series()
for index, facility in facilities_df.iterrows():
if (index % 25 == 0):
print('processing record #' + str(index))
name = facility['facility_name']
address = facility['facility_address']
# visit site
browser.visit(get_url(name))
# send data to scrape rating and assign it to rating column
rating = scrape_rating(browser.html, name, address)
facilities_df.loc[index,'rating'] = rating
facilities_df.to_csv('yelp_rating.csv', sep=',')
# -
# # Load Into Mongo DB
conn = 'mongodb://localhost:27017'
client = pymongo.MongoClient(conn)
db = client.restaurant_market_health_db
collection = db.inspection_summary
for i, inspection_series in inspections_data.iterrows():
collection.insert_one(transform_inspection(inspection_series, violations_data, facilities_df))
executable_path = {'executable_path': '/usr/local/bin/chromedriver'}
browser = Browser('chrome', **executable_path, headless=False)
facilities_df.to_csv('yelp_rating.csv', sep=',')
| LA restaurant.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Cusp Project, What I have so far.
# +
# okay lets do this
import pandas as pd
import numpy as np
filesList = ['55deg.txt', '60deg.txt','65deg.txt', '70deg.txt', '75deg.txt','80deg.txt','85deg.txt']
anglesList = [55, 60, 65, 70, 75, 80, 85]
cma = []
npi = 0
for files in filesList:
df = pd.read_csv(files)
# print df.tail()
Xgse = df['DefaultSC.gse.X']
Zgse = df['DefaultSC.gse.Z']
angle = np.arctan2(df['DefaultSC.gse.Z'], df['DefaultSC.gse.X'])
# make it into an array for iteration (probably a faster way)
# print angle
#print len(angle)
count = 0
for x in angle:
if 0.2151<=x<=0.2849:
count+=1
# print count
cma.append([count])
npi+=1
print cma
from pylab import *
cdict = {'red': ((0.0, 0.0, 0.0),
(0.5, 1.0, 0.7),
(1.0, 1.0, 1.0)),
'green': ((0.0, 0.0, 0.0),
(0.5, 1.0, 0.0),
(1.0, 1.0, 1.0)),
'blue': ((0.0, 0.0, 0.0),
(0.5, 1.0, 0.0),
(1.0, 0.5, 1.0))}
my_cmap = matplotlib.colors.LinearSegmentedColormap('my_colormap',cdict,256)
pcolor(cma,cmap=my_cmap)
colorbar()
plt.show()
# -
| cuspStudy/Cusp Project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="zwRfuC6RDb1B" colab_type="text"
# # Seminar 2. PyTorch
# Hi! Today we are going to study PyTorch. We'll compare numpy and PyTorch commands, rewrite our previous neural network in two ways.
#
# !!! GPU ON !!!
# + id="ilG0uJwVcke4" colab_type="code" colab={}
# !pip install mnist
# + id="cMKPD9TZdokn" colab_type="code" colab={}
from IPython import display
import numpy as np
import random
import torch
# + id="7gsxpRimFHF-" colab_type="code" colab={}
np.random.seed(42)
random.seed(42)
torch.manual_seed(42)
torch.cuda.manual_seed(42)
# + [markdown] id="b7xxXYAeFofM" colab_type="text"
# ## Numpy vs Pytorch
# ### Initialization
# + id="kbIPHFs6FmkB" colab_type="code" colab={}
a = [1. , 1.4 , 2.5]
print(f"Simple way: {torch.tensor(a)}")
print(f"Zeros:\n {torch.zeros((2,3))}")
print(f"Range: {torch.arange(0, 10)}")
print(f"Complicated range: {torch.arange(4, 12, 2)}")
print(f"Space: {torch.linspace(1, 4, 6)}")
print(f"Identity matrix:\n {torch.eye(4)}")
# + [markdown] id="sWKCkXaSGK1U" colab_type="text"
# ### Random
# + id="OHFgnpzOF5yb" colab_type="code" colab={}
print(f"From 0 to 1: {torch.rand(1)}")
print(f"Vector from 0 to 1: {torch.rand(5)}")
print(f"Vector from 0 to 10: {torch.randint(10, size=(5,))}")
# + [markdown] id="7EmP3g0wGfzT" colab_type="text"
# ### Matrix Operation
# + id="KfUdENnEF8-l" colab_type="code" colab={}
a = torch.arange(10).type(torch.FloatTensor)
b = torch.linspace(-10, 10, 10)
print(f"a: {a}\nshape: {a.size()}")
print(f"b: {a}\nshape: {b.size()}")
print(f"a + b: {a + b},\n a * b: {a * b}")
print(f"Dot product: {a.dot(b)}")
print(f"Mean: {a.mean()}, STD: {a.std()}")
print(f"Sum: {a.sum()}, Min: {a.min()}, Max: {a.max()}")
print(f"Reshape:\n{a.reshape(-1, 1)}\nshape: {a.reshape(-1, 1).size()}")
c = a.reshape(-1, 1).repeat(1, 5)
print(f"Repeat:\n{c}\nshape: {c.size()}")
print(f"Transpose:\n{c.T}\nshape: {c.T.size()}")
print(f"Unique items: {torch.unique(c)}")
# + [markdown] id="pdhrhXj9H0m6" colab_type="text"
# ### Indexing
# + id="bHqXM0bQGrRv" colab_type="code" colab={}
a = torch.arange(100).reshape(10, 10)
print(f"Array:\n{a}\nshape: {a.size()}")
print(f"Get first column: {a[:, 0]}")
print(f"Get last row: {a[-1, :]}")
print(f"Add new awis:\n{a[:, np.newaxis]}\nshape: {a[:, np.newaxis].size()}")
print(f"Specific indexing:\n{a[4:6, 7:]}")
# + [markdown] id="rFQ0OwEkIHu8" colab_type="text"
# ### Numpy <-> Pytorch
# + id="oueWvyMzH6sZ" colab_type="code" colab={}
a = torch.normal(mean=torch.zeros(2,4))
a.numpy()
# + id="VRCr6c78QeH1" colab_type="code" colab={}
b = np.random.normal(size=(2, 4))
torch.from_numpy(b)
# + [markdown] id="fJyktewrROPL" colab_type="text"
# ### CUDA
# + id="JshOGbX0RAb1" colab_type="code" colab={}
a = torch.normal(mean=torch.zeros(2,4))
b = torch.normal(mean=torch.zeros(2, 4))
print(f"a:\n{a}\nb:\n{b}")
# + id="mKUxAfveR6iQ" colab_type="code" colab={}
a = a.cuda()
# + id="klvLnHmaSDbk" colab_type="code" colab={}
a + b
# + id="11l9FxZVSEDL" colab_type="code" colab={}
(a + b.cuda()).cpu()
# + [markdown] id="DFkGBACsSmL4" colab_type="text"
# ### Autograd
# + id="Yr46uuNNSR_U" colab_type="code" colab={}
a = torch.randn(2, requires_grad=True)
b = torch.normal(mean=torch.zeros(2))
c = torch.dot(a, b)
print(f'a:\n{a}\nb:\n{b}\n(a,b): {c}')
# + id="Jmiq2lLnSvO4" colab_type="code" colab={}
c.backward()
print(f'a:\n{a}\nb:\n{b}\n(a,b): {c}')
# + id="O8I2iLEyS3dU" colab_type="code" colab={}
print(f"Grad a: {a.grad}")
# + [markdown] id="YFD2nov7UDlf" colab_type="text"
# Add function!
# + id="-z2P8UECTnEO" colab_type="code" colab={}
a = torch.randn(2, requires_grad=True)
b = torch.normal(mean=torch.zeros(2))
c = torch.ones(1, requires_grad=True)
d = torch.sigmoid(torch.dot(a, b) + c)
print(f'a:\n{a}\nb:\n{b}\nSigmoid( (a,b) ): {d}')
# + id="Ur_QA6xbTzeG" colab_type="code" colab={}
print(f"Grad a: {a.grad}\nGrad c: {c.grad}")
# + id="PsheM7soT6sw" colab_type="code" colab={}
d.backward()
print(f"Grad a: {a.grad}\nGrad c: {c.grad}")
# + [markdown] id="Yf9pO1XJUFZI" colab_type="text"
# Okay, what about vectors?
# + id="9ginrHkBT_HC" colab_type="code" colab={}
a = torch.randn(2, requires_grad=True)
b = torch.normal(mean=torch.zeros(2))
c = a * b
c.backward()
# + [markdown] id="-sNYtrJ3YxV3" colab_type="text"
# ## Neural Network. Rewind
# + id="_MKsdG6GdorV" colab_type="code" colab={}
from copy import deepcopy
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
import seaborn as sns
import mnist
sns.set()
# + id="waQtAlplIh6N" colab_type="code" colab={}
images = mnist.train_images() / 255
labels = mnist.train_labels()
X_train, X_valid, y_train, y_valid = train_test_split(images, labels)
# + id="BOHbffeqHyeF" colab_type="code" colab={}
def get_batches(dataset, batch_size):
X = dataset[0].reshape(-1, 28 * 28)
Y = dataset[1]
n_samples = X.shape[0]
indices = np.arange(n_samples)
np.random.shuffle(indices)
for start in range(0, n_samples, batch_size):
end = min(start + batch_size, n_samples)
batch_idx = indices[start:end]
yield torch.FloatTensor(X[batch_idx]), torch.LongTensor(Y[batch_idx])
# + id="R402HnG8ULdl" colab_type="code" colab={}
class CustomLinear:
def __init__(self, in_size, out_size):
"""
Simple linear layer
"""
self.in_size = in_size
self.out_size = out_size
self.w = ...
self.b = ...
def __call__(self, x):
return ...
def zero_grad(self):
self.w.grad = None
self.b.grad = None
# + id="vzDzCXmtZivn" colab_type="code" colab={}
class CustomNeuralNetwork:
def __init__(self, dims, activation="sigmoid"):
"""
Simple deep networks, that joins several linear layers.
"""
self.dims = dims
self.linears = ...
if activation == "sigmoid":
self.activation = torch.sigmoid
elif activation == "relu":
self.activation = torch.relu
else:
raise NotImplementedError
def __call__(self, x):
return ...
def zero_grad(self):
for l in self.linears:
l.zero_grad()
# + id="LZtMZC_dbmFS" colab_type="code" colab={}
class CustomCrossEntropy:
def __call__(self, x, target):
return ...
# + id="t47gvIlUkFpT" colab_type="code" colab={}
def simple_sgd(model, config):
with torch.no_grad():
for l in model.linears:
l.w -= config['learning_rate'] * l.w.grad
l.b -= config['learning_rate'] * l.b.grad
# + id="4ER9Cs5jaoNi" colab_type="code" colab={}
net = CustomNeuralNetwork((28 * 28, 10, 10))
criterion = CustomCrossEntropy()
# + id="gOQeiTU0Jnkr" colab_type="code" colab={}
optimizer_config = {
"learning_rate": 1e-1
}
# + id="5nwAo4jfaxry" colab_type="code" colab={}
def train(model, optimizer_config, n_epoch=20, batch_size=256):
train_logs = {"Train Loss": [0,], "Steps": [0,]}
valid_logs = {"Valid Loss": [0,], "Valid Accuracy": [0,], "Steps": [0,]}
step = 0
best_valid_loss = np.inf
best_model = None
for i in range(n_epoch):
for x_batch, y_batch in get_batches((X_train, y_train), batch_size):
model.zero_grad()
predictions = model(x_batch)
loss = criterion(predictions, y_batch)
loss.backward()
simple_sgd(model, optimizer_config)
step += 1
train_logs["Train Loss"].append(loss.detach().item())
train_logs["Steps"].append(step)
sum_loss = 0
sum_acc = 0
count_valid_steps = 0
with torch.no_grad():
for x_batch, y_batch in get_batches((X_valid, y_valid), batch_size):
predictions = model(x_batch)
loss = criterion(predictions, y_batch)
sum_loss += loss.item()
sum_acc += accuracy_score(y_batch, np.argmax(predictions.numpy(), axis=1))
count_valid_steps += 1
valid_logs["Valid Loss"].append(sum_loss / count_valid_steps)
valid_logs["Valid Accuracy"].append(sum_acc / count_valid_steps)
valid_logs["Steps"].append(step)
if best_valid_loss > sum_loss / count_valid_steps:
best_valid_loss = sum_loss / count_valid_steps
best_model = deepcopy(model)
fig, ax = plt.subplots(1, 3, figsize=(20, 5))
sns.lineplot(x="Steps", y="Train Loss", data=train_logs, ax=ax[0])
sns.lineplot(x="Steps", y="Valid Loss", data=valid_logs, ax=ax[1])
sns.lineplot(x="Steps", y="Valid Accuracy", data=valid_logs, ax=ax[2])
plt.plot()
return best_model, train_logs, valid_logs
# + id="ufy_8neBa3hG" colab_type="code" colab={}
net, _, _ = train(net, optimizer_config)
# + [markdown] id="oWnI3eIEeLZT" colab_type="text"
# ## Neural Network. Rewind #2
# + id="sLgLKCUfcCG3" colab_type="code" colab={}
import torch.nn as nn
class Linear(nn.Module):
def __init__(self, in_size, out_size):
super().__init__()
self.in_size = in_size
self.out_size = out_size
self.w = ...
self.b = ...
def forward(self, x):
return ...
class BatchNorm(nn.Module):
def __init__(self, in_size, alpha=0.1):
super().__init__()
self.in_size = in_size
self.beta = ...
self.gamma = ...
self.epsilon = 1e-5
def forward(self, x):
return ...
class Dropout(nn.Module):
def __init__(self, p=0.5):
super().__init__()
self.p = p
def forward(self, x):
return ...
class Block(nn.Module):
def __init__(self, in_size, out_size, activation="sigmoid", p=0.5):
super().__init__()
self.linear = Linear(in_size, out_size)
self.dropout = Dropout(p)
self.batch_norm = BatchNorm(out_size)
if activation == "sigmoid":
self.activation = torch.sigmoid
elif activation == "relu":
self.activation = torch.relu
else:
raise NotImplementedError
def forward(self, x):
x = ...
x = ...
x = ...
return ...
class Net(nn.Module):
def __init__(self, dims, activation="relu", p=0.5):
super().__init__()
self.blocks = nn.ModuleList(
list(Block(d_0, d_1, activation=activation, p=p) for d_0, d_1 in zip(dims[:-2], dims[1:-1]))
)
self.cl = Linear(dims[-2], dims[-1])
def forward(self, x):
for m in self.blocks:
x = m(x)
return self.cl(x)
# + id="PcqQ6AwPczjI" colab_type="code" colab={}
net = Net((28 * 28, 100, 10), p=0.1).cuda()
criterion = nn.CrossEntropyLoss()
# + id="cZtluygsczgm" colab_type="code" colab={}
optimizer = torch.optim.SGD(net.parameters(), lr=1e-1)
# + id="o2iqWQojL3J7" colab_type="code" colab={}
def train(model, optimizer, n_epoch=20, batch_size=256, device="cpu"):
train_logs = {"Train Loss": [0,], "Steps": [0,]}
valid_logs = {"Valid Loss": [0,], "Valid Accuracy": [0,], "Steps": [0,]}
step = 0
best_valid_loss = np.inf
best_model = None
for i in range(n_epoch):
for x_batch, y_batch in get_batches((X_train, y_train), batch_size):
x_batch = x_batch.to(device)
y_batch = y_batch.to(device)
optimizer.zero_grad()
predictions = model(x_batch)
loss = criterion(predictions, y_batch)
loss.backward()
optimizer.step()
step += 1
train_logs["Train Loss"].append(loss.detach().item())
train_logs["Steps"].append(step)
sum_loss = 0
sum_acc = 0
count_valid_steps = 0
with torch.no_grad():
for x_batch, y_batch in get_batches((X_valid, y_valid), batch_size):
x_batch = x_batch.to(device)
y_batch = y_batch.to(device)
predictions = model(x_batch)
loss = criterion(predictions, y_batch)
sum_loss += loss.item()
sum_acc += accuracy_score(y_batch.cpu().numpy(), np.argmax(predictions.cpu().numpy(), axis=1))
count_valid_steps += 1
valid_logs["Valid Loss"].append(sum_loss / count_valid_steps)
valid_logs["Valid Accuracy"].append(sum_acc / count_valid_steps)
valid_logs["Steps"].append(step)
if best_valid_loss > sum_loss / count_valid_steps:
best_valid_loss = sum_loss / count_valid_steps
best_model = deepcopy(net)
fig, ax = plt.subplots(1, 3, figsize=(20, 5))
sns.lineplot(x="Steps", y="Train Loss", data=train_logs, ax=ax[0])
sns.lineplot(x="Steps", y="Valid Loss", data=valid_logs, ax=ax[1])
sns.lineplot(x="Steps", y="Valid Accuracy", data=valid_logs, ax=ax[2])
plt.plot()
return best_model, train_logs, valid_logs
# + id="QDkpB7TtL3IU" colab_type="code" colab={}
net, _, _ = train(net, optimizer, device="cuda:0")
# + [markdown] id="mcOEa7lcjiyO" colab_type="text"
# ### Neural Network. Rewind #3. Logging
#
# Logging systems:
# - [Tensorboard](https://pytorch.org/docs/stable/tensorboard.html)
# - [WandB](https://www.wandb.com/)
# + id="fjRHFJXSjn81" colab_type="code" colab={}
class Block(nn.Module):
def __init__(self, in_size, out_size, activation="relu", p=0.5):
super().__init__()
self.in_size = in_size
self.out_size = out_size
if activation == "sigmoid":
self.activation = nn.Sigmoid
elif activation == "relu":
self.activation = nn.ReLU
else:
raise NotImplementedError
self.fc = nn.Sequential(
...
)
def forward(self, x):
return self.fc(x)
net = nn.Sequential(Block(28 * 28, 100, p=0.2), nn.Linear(100, 10)).cuda()
optimizer = torch.optim.SGD(net.parameters(), lr=1e-1)
criterion = nn.CrossEntropyLoss()
# + id="QV1AxNFbfapY" colab_type="code" colab={}
# %load_ext tensorboard
# + id="Cgo_htQZfamn" colab_type="code" colab={}
from datetime import datetime
from pathlib import Path
from torch.utils.tensorboard import SummaryWriter
# + id="4qWa59wKf5Jd" colab_type="code" colab={}
def train(model, optimizer, n_epoch=20, batch_size=256, device="cpu"):
writer = SummaryWriter(Path("logs") / datetime.now().strftime("%Y%m%d-%H%M%S"))
step = 0
best_valid_loss = np.inf
best_model = None
for i in range(n_epoch):
for x_batch, y_batch in get_batches((X_train, y_train), batch_size):
x_batch = x_batch.to(device)
y_batch = y_batch.to(device)
optimizer.zero_grad()
predictions = model(x_batch)
loss = criterion(predictions, y_batch)
loss.backward()
optimizer.step()
step += 1
writer.add_scalar("Train Loss", loss.detach().item(), step)
sum_loss = 0
sum_acc = 0
count_valid_steps = 0
with torch.no_grad():
for x_batch, y_batch in get_batches((X_valid, y_valid), batch_size):
x_batch = x_batch.to(device)
y_batch = y_batch.to(device)
predictions = model(x_batch)
loss = criterion(predictions, y_batch)
sum_loss += loss.item()
sum_acc += accuracy_score(y_batch.cpu().numpy(), np.argmax(predictions.cpu().numpy(), axis=1))
count_valid_steps += 1
writer.add_scalar("Valid Loss", sum_loss / count_valid_steps, step)
writer.add_scalar("Valid Accuracy", sum_acc / count_valid_steps, step)
if best_valid_loss > sum_loss / count_valid_steps:
best_valid_loss = sum_loss / count_valid_steps
best_model = deepcopy(net)
return best_model
# + id="qptL6E_Mf5G5" colab_type="code" colab={}
net = train(net, optimizer, device="cuda:0")
# + id="SuzkIqc0f5FG" colab_type="code" colab={}
# %tensorboard --logdir logs
# + [markdown] id="7W1zyKnXFdxt" colab_type="text"
# ## Spoiler - train loop with [Catalyst](https://github.com/catalyst-team/catalyst)
#
# - [A comprehensive step-by-step guide to basic and advanced features](https://github.com/catalyst-team/catalyst#step-by-step-guide)
# - [Docs](https://catalyst-team.github.io/catalyst/)
# - [What is Runner?](https://catalyst-team.github.io/catalyst/api/core.html#runner)
# + id="8Y_cQnMeJJiU" colab_type="code" colab={}
# !pip install catalyst
# + id="Gm2SIio7f5Cd" colab_type="code" colab={}
import os
import torch
from torch.nn import functional as F
from torch.utils.data import DataLoader
from catalyst import dl
from catalyst.data.cv import ToTensor
from catalyst.contrib.datasets import MNIST
from catalyst.utils import metrics
net = nn.Sequential(Block(28 * 28, 100, p=0.2), nn.Linear(100, 10))
optimizer = torch.optim.Adam(net.parameters(), lr=0.02)
criterion = torch.nn.CrossEntropyLoss()
loaders = {
"train": DataLoader(MNIST(os.getcwd(), train=True, download=True, transform=ToTensor()), batch_size=32),
"valid": DataLoader(MNIST(os.getcwd(), train=False, download=True, transform=ToTensor()), batch_size=32),
}
class CustomRunner(dl.Runner):
def predict_batch(self, batch):
# model inference step
return self.model(batch[0].to(self.device).view(batch[0].size(0), -1))
def _handle_batch(self, batch):
# model train/valid step
x, y = batch
y_hat = self.model(x.view(x.size(0), -1))
loss = self.criterion(y_hat, y)
accuracy01, accuracy03 = metrics.accuracy(y_hat, y, topk=(1, 3))
self.batch_metrics.update(
{"loss": loss, "accuracy01": accuracy01, "accuracy03": accuracy03}
)
if self.is_train_loader:
loss.backward()
self.optimizer.step()
self.optimizer.zero_grad()
runner = CustomRunner()
runner.train(
model=net,
criterion=criterion,
optimizer=optimizer,
loaders=loaders,
logdir=Path("logs") / datetime.now().strftime("%Y%m%d-%H%M%S"),
num_epochs=5,
verbose=True,
load_best_on_end=True,
)
traced_model = runner.trace(loader=loaders["valid"])
# + id="ZUz3o_kJJIHe" colab_type="code" colab={}
| week-2/seminar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import sys
import numpy as np
import pandas as pd
import itertools
from collections import Counter
from scipy.stats import gaussian_kde
import pickle
saved_path = os.path.join(os.path.dirname(os.getcwd()), 'Saved Data\\')
import matplotlib.pyplot as plt
import matplotlib
import seaborn as sns
sns.set(color_codes=True)
# %matplotlib inline
plt.rcParams["figure.figsize"] = [16, 6]
import warnings
warnings.filterwarnings("ignore")
from matplotlib.axes._axes import _log as matplotlib_axes_logger
matplotlib_axes_logger.setLevel('ERROR')
'LOAD THE DATASET'
url = saved_path + 'requetes_hib_select.csv'
requetes = pd.read_csv(url, index_col=[0])
print('The number of queries is :',requetes.shape[0])
print('The number of queries with 0 lines returned : ',requetes[requetes['nbLignes'] == 0].shape[0])
"f(nombre de lignes) = temps d'execution"
fig, axes = plt.subplots(nrows = 1, ncols = 2)
requetes.plot.scatter(x = 'nbLignes', y = 'durationMS', label = 'hibernate queries', logy = False, ax = axes[0])
requetes.plot.scatter(x = 'nbLignes', y = 'durationMS', label = 'hibernate queries log scale',logx = False ,logy = True, ax = axes[1])
sns.FacetGrid(requetes, hue="DeclinaisonCOP", palette="husl", size=10).map(plt.scatter, "nbLignes", "durationMS").add_legend()
sns.catplot(x="DeclinaisonCOP", y="durationMS",jitter=False ,data=requetes,size =10)
sns.FacetGrid(requetes, hue="DeclinaisonCOP", palette="husl", size=10).map(plt.hist, "durationMS").set(yscale = 'log').add_legend()
sns.FacetGrid(requetes, hue="versionBDD", palette="husl", size=10).map(plt.scatter, "nbLignes", "durationMS").add_legend()
sns.FacetGrid(data = requetes, hue="DeclinaisonCOP", palette="husl", size=10).map(plt.scatter, "versionBDD", "durationMS").add_legend()
sns.FacetGrid(requetes, hue="versionBDD", palette="husl", size=10).map(plt.hist, "durationMS").set(yscale = 'log').add_legend()
sns.FacetGrid(requetes, hue="DeclinaisonCOP", palette="husl", size=10).map(plt.scatter, "long", "durationMS").add_legend().set(yscale = 'log')
sns.FacetGrid(requetes, hue="DeclinaisonCOP", palette="husl", size=10).map(plt.hist, "long").set(yscale = 'log').add_legend()
sns.catplot(x="scoreAnomalieRepartition", y="durationMS",jitter=False ,data=requetes,size=10)
sns.catplot(x="moyenneNbSessionsActives", y="durationMS",jitter=False ,data=requetes, size = 10)
sns.catplot(x="nbSessionBDBloquee", y="durationMS",jitter=False ,data = requetes, size = 10)
sns.catplot(x="zScoreNbPoolConnexionActif", y="durationMS",jitter=False ,data = requetes, size = 10)
requetes_uniques = pd.read_csv(saved_path + 'requetes_uniques.csv',index_col=[0])
requetes_uniques = requetes_uniques.set_index('requete')['frequence']
density = gaussian_kde(requetes_uniques.values[requetes_uniques.values > 100])
xs = np.linspace(0,10560,500)
density.covariance_factor = lambda : .5
density._compute_covariance()
plt.plot(xs,density(xs))
plt.show()
'TOP K queries'
top_requetes = requetes_uniques[:50]
dict_encod = pd.Series(np.arange(top_requetes.size),top_requetes.index).to_dict()
df = requetes[requetes['requete'].isin(top_requetes.index)]
df['requete'] = df['requete'].replace(dict_encod)
sns.catplot(x="requete", y="durationMS",jitter=False ,data=df,size = 10)
dict(map(reversed, dict_encod.items()))[10]
instances = df[df.requete == 10].instanceCode.value_counts().index[:10]
instances
g = sns.catplot(x="instanceCode", y="durationMS",jitter=False ,data=df[(df.requete == 10) & df.instanceCode.isin(instances)],size =10)
g.set_xticklabels(rotation=30)
| Code/sd-4sql/supplementary/query-analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="2MWcShQdrUu2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600205463313, "user_tz": 420, "elapsed": 17980, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gja41a6NZM-sXna8IDZYolHfTSms_knBAFrWLv8=s64", "userId": "08949961291514014620"}} outputId="93e292d7-cac9-4b89-a815-2033e0b473c2"
from google.colab import drive
drive.mount('/content/drive')
# + id="BOOeOyeerJj2" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1600206104350, "user_tz": 420, "elapsed": 350, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gja41a6NZM-sXna8IDZYolHfTSms_knBAFrWLv8=s64", "userId": "08949961291514014620"}}
from IPython.display import Image
import pandas as pd
# + [markdown] id="wKNB9ijIgRQy" colab_type="text"
# # Lab #-1 How to Use Python and Colab for Lab Notebooks
# + [markdown] id="QBLITMTMgeqy" colab_type="text"
# **Name: <NAME>**
#
# **Partner: <NAME>**
#
#
# + [markdown] id="D6JuDAPLIBRw" colab_type="text"
# # Table of contents
# *Note: The navigation bar on the left side of colab has a table of contents that you can use as a reference when creating your own.*
# * Objective
# * Introduction
# * Organization
# * Use of Sections
# * Images
# * Images Intro
# * Student Work
# * Mathematical Equations
# * LaTeX
# * Using Images
# * Student work
# * Data
# * Data taking
# * Loading data from Scope
# * Plotting Data
# * Student Work
# * Prelab for Lab #1
# * Conclusion
# + [markdown] id="rl2PsrfOg8ft" colab_type="text"
# # Objective
# In this section you describe in detail what you are going in the lab with goals...(which you know because you
# read the lab before, right?)
# + [markdown] id="--k5urp1hAqI" colab_type="text"
# # Introduction
# Some words about background and theory; include any new principles and equations you'll be using. If you don't know where to include the sketch of the circuit you're going to build, include it here.
# + [markdown] id="i6eTZg_7hMgA" colab_type="text"
# # Organization
# 1. Properly formated lab notebooks are much easier to read and follow (for your future self and for the
# TA marker)
# 2. So do use bullet lists and number lists
# 3. Each major section should use the '#' format
# 4. Each minor section should use the '##' format
# 5. Consider numbering your major sections; it makes it easier to refer back to them.
#
#
#
#
#
#
# + [markdown] id="-EWvhw0QifrK" colab_type="text"
# ## Use of Sections
# For each thought or concept, use a section break and add a new entry.
# + [markdown] id="C0UUfOWSimXJ" colab_type="text"
# # Images
#
# + [markdown] id="JI5nLVlfSUEy" colab_type="text"
# ## Images Intro
#
# Although there are many ways to add an image such that it shows up in the notebook view, I have only found **one** way to to make images show up in the exported pdf file.
#
# To insert an image:
# 1. Resize image to ~300px wide before uploading to your Google Drive (can be done in MS Paint for example)
# 2. Run `from Ipython.display import Image` somewhere in your notebook (probably the very top)
# 3. Then in a code cell run the line `Image(filename="<image path in google drive>")`
# *Note: Although you can specify width and height in the `Image` function, it will not work when it is exported to PDF*.
#
# + id="olE0MTSN1xP3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600141304254, "user_tz": 420, "elapsed": 469, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjtB83PLZbGcVLZH6fFJWZqfzyWhH1c0dgQgKV7lw=s64", "userId": "15145304080398626492"}} outputId="a36566ab-9db9-4d70-93a3-184c9a46446b"
from IPython.display import Image
from google.colab import drive
drive.mount('/content/drive')
# + id="SjWhjilVrpOp" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1600205676445, "user_tz": 420, "elapsed": 308, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gja41a6NZM-sXna8IDZYolHfTSms_knBAFrWLv8=s64", "userId": "08949961291514014620"}}
import os
ddir = 'drive/My Drive/ENPHYS_259/Intro'
# + id="ccpLlJFer59S" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} executionInfo={"status": "ok", "timestamp": 1600205704463, "user_tz": 420, "elapsed": 428, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gja41a6NZM-sXna8IDZYolHfTSms_knBAFrWLv8=s64", "userId": "08949961291514014620"}} outputId="74b6d756-05ca-443c-ad71-da9e89acf116"
os.listdir(ddir)
# + id="5jwMp0cN147U" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 317} executionInfo={"status": "ok", "timestamp": 1600205734376, "user_tz": 420, "elapsed": 924, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gja41a6NZM-sXna8IDZYolHfTSms_knBAFrWLv8=s64", "userId": "08949961291514014620"}} outputId="2b5b9f77-ea6d-4595-90ca-97ec1bd0745e"
Image(filename=os.path.join(ddir, 'cat_300.jpg')) # This path needs changing
# + [markdown] id="OCFdEVR02ryM" colab_type="text"
# *Figure 1: Safety Cat*
# + [markdown] id="cwpLj3y7ysJn" colab_type="text"
#
# <figure>
# <img src="https://drive.google.com/uc?id=1Bxb9qoPCrlfJ5cIZcqjGVE5D-7sJugxJ" width="400"/>
# <figcaption>Figure 2: Experimental setup for interferometer</figcaption>
# </figure>
# + [markdown] id="fQKJIcWKnrCI" colab_type="text"
# You should reference the figures in your entries. For example, Fig. 1 shows a good cat. DO NOT include a figure and then not discuss it...otherwise what is the point?
#
# Figure 2 shows an example of an
# actual experimental setup.
# + [markdown] id="WQQyuzhqky47" colab_type="text"
# ## Student Work
# Place an image below and properly caption it
# + id="TYis7UV2zC_b" colab_type="code" colab={}
# + [markdown] id="YdNHKbDKy-U0" colab_type="text"
# # Mathematical Equations
# + [markdown] id="2rFwFpa7zBSC" colab_type="text"
# There are two ways to put equations in:
#
# ## LaTeX
# The first is a widely used formatting scripting language called LaTex (or Tex) which is superior to render mathematical expressions and is used widely. You will use in later classes (ENPH 257, PHYS352 among others) so it will pay off to learn (and use) it now.
#
# To insert an expression wrap the LaTeX formatted equation in either `$` signs for inline equations, or `$$` for full line equations
#
# e.g. If you enter
#
# `$$\alpha^2+\beta^2=c^2$$`
#
# This is the result:
# $$\alpha^2+\beta^2=c^2$$
#
# Single `$` signs allow you to put equations in the $\alpha^2+\beta^2=c^2$ middle of sentences.
#
# Go to https://www.codecogs.com/latex/eqneditor.php or https://www.latex4technics.com/ for help with writing LaTeX equations.
#
# Here is an example of a more complicated formula,
#
# `$$P_\lambda = \frac{2 \pi h c^2}{\lambda^5 \left(e^{\left(\frac{h c}{\lambda k T}\right)} - 1\right)}$$`
# $$P_\lambda = \frac{2 \pi h c^2}{\lambda^5 \left(e^{\left(\frac{h c}{\lambda k T}\right)} - 1\right)}$$
# + [markdown] id="gS7OIMPq4YEZ" colab_type="text"
# ## Using Images
# If there is a long derivation if it often easier to write it out and take a picture as shown in Fig. 3.
#
# <figure>
# <img src="https://drive.google.com/uc?id=1-rIfkL7YHe-2OQT8mNryY-FqvdT9xkYH" width="400"/>
# <figcaption>Fig 3: Derivation of Uncertainty propagation</figcaption>
# </figure>
# <figure>
# <img src="https://drive.google.com/uc?id=1j8gmP3o-oKFqCB6v_EiHPeazBJwkUady" width="400"/>
# <figcaption>Fig 4: Bad picture of derivation of Uncertainty. There is no chance to read it</figcaption>
# </figure>
# + [markdown] id="z6X9jxBz5rC1" colab_type="text"
# ## Student Work
# Enter the Taylor expansion of sin(x)
# + id="URKq-egZ5x1H" colab_type="code" colab={}
# + [markdown] id="svqxGv2k5y4P" colab_type="text"
# # Data
# Plotting and analyzing data is one of the many things Python is well suited for.
#
# + [markdown] id="E4iFnAjFSae7" colab_type="text"
# ## Data Taking
# * Say you are going to record two sets of data.
# * First switch to a code section and type
#
# ```
# xdata = [1, 2, 3, 4, 5, 6, 7, 8] # this is a list with 8 data points
# ydata = [0.8, 3.8, 8.9, 17, 26, 35.8, 49.1, 63.3] # this is also a list with 8 data points
# ```
#
# * The `#` is a delimiting character for commenting
# * Lists are entered with `[]`
#
# Its often useful to work with arrays which can be multi-dimensional. For this we can use the `numpy` package in python. To use a numpy array, the `numpy` package must first be imported into the script using an `import` statement. It is common to import numpy like this: `import numpy as np`. The `as np` part just makes it quicker to type. Any import statements only need to be run once, but it doesn't hurt to run them multiple times. (Note: We usually import whatever packages we'll run at the top of our file.)
#
# e.g.
#
# ```
# import numpy as np
# xdata = np.array([1,2,3,4,5,6,7,8])
# ```
# To create a numpy array from scratch. Or
# ```
# ydata = np.array(ydata)
# ```
# To convert ydata from the `list` we entered above to a numpy `ndarray`
# + [markdown] id="ZnPekzbA6I_0" colab_type="text"
# ## Loading Data from Scope
#
# 1. Copy the files to the same folder as the `.ipynb` file you want to work with them in. This is your directory (e.g. `Python_Introduction` or `Lab1`).
# 2. Make sure your drive is mounted (i.e. give colab access to your google drive):
# * Open the help menu (`ctrl+M+H`).
# * Scroll down and find the option to "Mount Drive."
# * Create the following keyboard shortcut for it: `ctrl+D`
# * Press `ctrl+D` and follow the directions to mount your drive.
# 3. Set your data directory (see example below):
# + id="Vu7jRKh7TR3n" colab_type="code" colab={}
dir_path = 'ENPH259/Python_Introduction' # change this to your data directory path - e.g. ENPH259/Lab1
ddir = f'/content/drive/My Drive/{dir_path}'
# + [markdown] id="cv9kb4GUTT5h" colab_type="text"
# 4. List the contents of your directory to ensure your file is there:
# + id="NQAwzGkdTbvJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} executionInfo={"status": "ok", "timestamp": 1600206059963, "user_tz": 420, "elapsed": 280, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gja41a6NZM-sXna8IDZYolHfTSms_knBAFrWLv8=s64", "userId": "08949961291514014620"}} outputId="e70d1727-4687-46ae-d380-b7c52f745833"
print(os.listdir(ddir))
# + [markdown] id="RdvHDRkITcww" colab_type="text"
# 5. Load your data (example is a txt file, but you can do this with csv, xlsx...):
# + id="-duqzywGTs4g" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 419} executionInfo={"status": "ok", "timestamp": 1600206207307, "user_tz": 420, "elapsed": 332, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gja41a6NZM-sXna8IDZYolHfTSms_knBAFrWLv8=s64", "userId": "08949961291514014620"}} outputId="1ff8b616-d8f8-4066-91fa-506b81aff1fb"
data_path = os.path.join(ddir, 'square1khz.txt')
df_square = pd.read_csv(data_path, header=31, sep='\t') # 31 comment lines at the top, and the values are tab (\t) separated
df_square # df stands for dataframe
x = df_square['Time (s)']
y = df_square['Channel 2 (V)']
df_square
# + [markdown] id="4RwgfuEDLxaD" colab_type="text"
# ## Plotting Data
#
# To get started with plotting, we need to first `import` a useful library for plotting. There are many plotting packages available for Python, one of the most common is Matplotlib which as the name suggests, is very similar to Matlab plotting. Specifically we want the `matplotlib.pyplot` package. To make it easier to use we will import it `as plt` so that we only have to type `plt.` to use it. If you didn't see the import packages cell at the top of this tutorial, go back and take a look at it. Run the cell. You only need to import packages once per file, but we've included the code cell below as an example of the import syntax. You'll notice this is the same syntax we used at the beginning.
# + id="Ije9YISYL2C0" colab_type="code" colab={}
import matplotlib.pyplot as plt
import numpy as np # Importing numpy here so I can quickly make arrays of data
# + id="REqajvRdMAHM" colab_type="code" colab={}
x = np.linspace(0, 10, 100) # Using numpy to make a quick array of x coords from 0 to 10 with 100 points
y = np.sin(x)
# + id="-0GN9cCDMLZu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="e73affb9-c817-459a-e4b9-d71681bd5569"
fig, ax = plt.subplots(1,1) # Making a figure with 1 rows and 1 columns of axes to plot on (i.e. a figure with one plot)
ax.plot(x, y, label='sine wave'); # plotting the data onto the axes. (the label is for the legend we add next)
# + [markdown] id="2grQi6wrMm93" colab_type="text"
# All plots **MUST** have axis labels, titles, and legends. You can add these with some very simple commands
#
# + id="O2dW36ZyNli1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="b43aa756-8350-4355-8114-3c47bfe6732e"
# I can carry on using the figure and axes we made previously by using their handles (fig and ax)
ax.set_title('Title of Axes')
ax.set_xlabel('The X label')
ax.set_ylabel('The Y label')
ax.legend() # Turns the legend on and uses the labels that were given when the data was plotted
fig # This makes the figure show up again after this cell even though we didn't create it here
# + [markdown] id="7bGRzG9QNzfb" colab_type="text"
# ## Student Work
# * Make two arrays that are 10 elements long
# * Plot one against the other (add all the necessary extra information)
# + [markdown] id="nqTOmdpKOHmH" colab_type="text"
# # Prelab for Lab #1
#
# For second prelab question we are asked to plot the frequency response of an RC circuit for Lab #1
#
# * you can generate a linear or log space array using:
# `xlin = np.linspace(1, 100, 1000)` defines a vector xlin with a 1000 linearly spaced points from 1 to 100 `xlog = np.logspace(1, 100, 1000)` defines a vector xlog with a 1000 log spaced points from 1 to 100
#
# * Now the frequency response of the voltage across a capacitor normalized to the input voltage of an RC circuit is:
#
# + [markdown] id="B06q0Q2dOc7x" colab_type="text"
# # Conclusion
#
# Always include a conclusion!
# + id="yepCY5q5UJlk" colab_type="code" colab={}
# + id="n1pgM_egOeEN" colab_type="code" colab={}
| Tutorial 1/Lab_-1_Python_Lab_Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# Writing Tunable Templates and Using the Auto-tuner
# ==================================================
# **Author**: `<NAME> <https://github.com/merrymercy>`_
#
# This is an introduction tutorial to the auto-tuning module in TVM.
#
# There are two steps in auto-tuning.
# The first step is defining a search space.
# The second step is running a search algorithm to explore through this space.
# In this tutorial, you can learn how to perform these two steps in TVM.
# The whole workflow is illustrated by a matrix multiplication example.
#
# Note that this tutorial will not run on Windows or recent versions of macOS. To
# get it to run, you will need to wrap the body of this tutorial in a :code:`if
# __name__ == "__main__":` block.
#
# Install dependencies
# --------------------
# To use autotvm package in TVM, we need to install some extra dependencies.
# This step (installing xgboost) can be skipped as it doesn't need XGBoost
# (change "3" to "2" if you use python2):
#
# .. code-block:: bash
#
# pip3 install --user psutil xgboost cloudpickle
#
# To make TVM run faster in tuning, it is recommended to use cython
# as FFI of TVM. In the root directory of TVM, execute
# (change "3" to "2" if you use python2):
#
# .. code-block:: bash
#
# pip3 install --user cython
# sudo make cython3
#
# Now return to python code. Import packages.
#
#
# +
import logging
import sys
import numpy as np
import tvm
from tvm import te, testing
# the module is called `autotvm`
from tvm import autotvm
# -
# Step 1: Define the search space
# --------------------------------
# In this section, we will rewrite a deterministic TVM schedule code to a
# tunable schedule template. You can regard the process of search space definition
# as the parameterization of our existing schedule code.
#
# To begin with, here is how we implement a blocked matrix multiplication in TVM.
#
#
# Matmul V0: Constant tiling factor
def matmul_v0(N, L, M, dtype):
A = te.placeholder((N, L), name="A", dtype=dtype)
B = te.placeholder((L, M), name="B", dtype=dtype)
k = te.reduce_axis((0, L), name="k")
C = te.compute((N, M), lambda i, j: te.sum(A[i, k] * B[k, j], axis=k), name="C")
s = te.create_schedule(C.op)
# schedule
y, x = s[C].op.axis
k = s[C].op.reduce_axis[0]
yo, yi = s[C].split(y, 8)
xo, xi = s[C].split(x, 8)
s[C].reorder(yo, xo, k, yi, xi)
return s, [A, B, C]
# Parametrize the schedule
# ^^^^^^^^^^^^^^^^^^^^^^^^
# In the previous schedule code, we use a constant "8" as tiling factor.
# However, it might not be the best one because the best tiling factor depends
# on real hardware environment and input shape.
#
# If you want the schedule code to be portable across a wider range of input shapes
# and target hardware, it is better to define a set of candidate values and
# pick the best one according to the measurement results on target hardware.
#
# In autotvm, we can define a tunable parameter, or a "knob" for such kind of value.
#
#
# Matmul V1: List candidate values
@autotvm.template("tutorial/matmul_v1") # 1. use a decorator
def matmul_v1(N, L, M, dtype):
A = te.placeholder((N, L), name="A", dtype=dtype)
B = te.placeholder((L, M), name="B", dtype=dtype)
k = te.reduce_axis((0, L), name="k")
C = te.compute((N, M), lambda i, j: te.sum(A[i, k] * B[k, j], axis=k), name="C")
s = te.create_schedule(C.op)
# schedule
y, x = s[C].op.axis
k = s[C].op.reduce_axis[0]
# 2. get the config object
cfg = autotvm.get_config()
# 3. define search space
cfg.define_knob("tile_y", [1, 2, 4, 8, 16])
cfg.define_knob("tile_x", [1, 2, 4, 8, 16])
# 4. schedule according to config
yo, yi = s[C].split(y, cfg["tile_y"].val)
xo, xi = s[C].split(x, cfg["tile_x"].val)
s[C].reorder(yo, xo, k, yi, xi)
return s, [A, B, C]
# Here we make four modifications to the previous schedule code and get
# a tunable "template". We can explain the modifications one by one.
#
# 1. Use a decorator to mark this function as a simple template.
# 2. Get a config object:
# You can regard this :code:`cfg` as an argument of this function but
# we obtain it in a different way. With this argument, this function is no longer
# a deterministic schedule code. Instead, we can pass different configurations to
# this function and get different schedules, so this function is a "template".
#
# To make the template function more compact, we do two things in a single function.
# (1) define a search space and (2) schedule according to an entity in this space.
# To achieve this, we make :code:`cfg` be either
# a :any:`ConfigSpace` or a :any:`ConfigEntity` object.
#
# When it is a :any:`ConfigSpace`, it will collect all tunable knobs in this function and
# build the search space.
# When it is a :any:`ConfigEntity`, it will ignore all space definition API
# (namely, :code:`cfg.define_XXXXX(...)`). Instead, it stores deterministic values for
# all tunable knobs, and we schedule according to these values.
#
# During auto-tuning, we will first call this template with a :any:`ConfigSpace`
# object to build the search space. Then we call this template with different :any:`ConfigEntity`
# in the built space to get different schedules. Finally we will measure the code generated by
# different schedules and pick the best one.
#
# 3. Define two tunable knobs. The first one is :code:`tile_y` with
# 5 possible values. The second one is :code:`tile_x` with a same
# list of possible values. These two knobs are independent, so they
# span a search space with size = 5x5 = 25
# 4. Schedule according to the deterministic values in :code:`cfg`
#
#
#
# Use better space definition API
# ^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
# In the previous template, we manually list all possible values for a knob.
# This is the lowest level API to define the space.
# However, we also provide another set of API to make the space definition
# easier and smarter. It is recommended to use this set of high level API.
#
# In the following example, we use :any:`ConfigSpace.define_split` to define a split
# knob. It will enumerate all the possible ways to split an axis and construct
# the space.
#
# We also have :any:`ConfigSpace.define_reorder` for reorder knob and
# :any:`ConfigSpace.define_annotate` for annotation like unroll, vectorization,
# thread binding.
# When the high level API cannot meet your requirement, you can always fall
# back to use low level API.
#
#
@autotvm.template("tutorial/matmul")
def matmul(N, L, M, dtype):
A = te.placeholder((N, L), name="A", dtype=dtype)
B = te.placeholder((L, M), name="B", dtype=dtype)
k = te.reduce_axis((0, L), name="k")
C = te.compute((N, M), lambda i, j: te.sum(A[i, k] * B[k, j], axis=k), name="C")
s = te.create_schedule(C.op)
# schedule
y, x = s[C].op.axis
k = s[C].op.reduce_axis[0]
##### define space begin #####
cfg = autotvm.get_config()
cfg.define_split("tile_y", y, num_outputs=2)
cfg.define_split("tile_x", x, num_outputs=2)
##### define space end #####
# schedule according to config
yo, yi = cfg["tile_y"].apply(s, C, y)
xo, xi = cfg["tile_x"].apply(s, C, x)
s[C].reorder(yo, xo, k, yi, xi)
return s, [A, B, C]
# <div class="alert alert-info"><h4>Note</h4><p>More Explanation on :code:`cfg.defile_split`</p></div>
#
# In this template, :code:`cfg.define_split("tile_y", y, num_outputs=2)` will enumerate
# all possible combinations that can split axis y into two axes with factors of the length of y.
# For example, if the length of y is 32 and we want to split it into two axes
# using factors of 32, then there are 6 possible values for
# (length of outer axis, length of inner axis) pair, namely
# (32, 1), (16, 2), (8, 4), (4, 8), (2, 16) or (1, 32).
# They are just the 6 possible values of `tile_y`.
#
# During schedule, :code:`cfg["tile_y"]` is a :code:`SplitEntity` object.
# We stores the lengths of outer axes and inner axes in :code:`cfg['tile_y'].size`
# (a tuple with two elements).
# In this template, we apply it by using :code:`yo, yi = cfg['tile_y'].apply(s, C, y)`.
# Actually, this is equivalent to
# :code:`yo, yi = s[C].split(y, cfg["tile_y"].size[1])`
# or :code:`yo, yi = s[C].split(y, nparts=cfg['tile_y"].size[0])`
#
# The advantage of using cfg.apply API is that it makes multi-level split
# (when num_outputs >= 3) easier.
#
#
# Step 2: Search through the space
# ---------------------------------
# In step 1, we build the search space by extending our old schedule code
# into a template. The next step is to pick a tuner and explore in this space.
#
# Auto-tuners in TVM
# ^^^^^^^^^^^^^^^^^^
# The job for a tuner can be described by following pseudo code
#
# .. code-block:: c
#
# ct = 0
# while ct < max_number_of_trials:
# propose a batch of configs
# measure this batch of configs on real hardware and get results
# ct += batch_size
#
# When proposing the next batch of configs, the tuner can take different strategies. We
# provide four tuners with different strategies in autotvm.
#
# * :any:`RandomTuner`: Enumerate the space in a random order
# * :any:`GridSearchTuner`: Enumerate the space in a grid search order
# * :any:`GATuner`: Using genetic algorithm to search through the space
# * :any:`XGBTuner`: Uses a model based method. Train a XGBoost model to predict the speed of lowered IR and pick the next batch according to the prediction.
#
# You can choose the tuner according to the size of your space, your time budget and other factors.
# For example, if your space is very small (less than 1000), a gridsearch tuner or a
# random tuner is good enough. If your space is at the level of 10^9 (this is the space
# size of a conv2d operator on CUDA GPU), XGBoostTuner can explore more efficiently
# and find better configs.
#
#
# Begin tuning
# ^^^^^^^^^^^^
# Here we continue our matrix multiplication example.
# First we should create a tuning task.
# We can also inspect the initialized search space.
# In this case, for a 512x512 square matrix multiplication, the space size
# is 10x10=100
#
#
N, L, M = 512, 512, 512
task = autotvm.task.create("tutorial/matmul", args=(N, L, M, "float32"), target="llvm")
print(task.config_space)
# Then we need to define how to measure the generated code and pick a tuner.
# Since our space is small, a random tuner is just okay.
#
# We only make 10 trials in this tutorial for demonstration. In practice,
# you can do more trials according to your time budget.
# We will log the tuning results into a log file. This file can be
# used to get the best config later.
#
#
# +
# logging config (for printing tuning log to the screen)
logging.getLogger("autotvm").setLevel(logging.DEBUG)
logging.getLogger("autotvm").addHandler(logging.StreamHandler(sys.stdout))
# There are two steps for measuring a config: build and run.
# By default, we use all CPU cores to compile program. Then measure them sequentially.
# We measure 5 times and take average to reduce variance.
measure_option = autotvm.measure_option(builder="local", runner=autotvm.LocalRunner(number=5))
# Begin tuning with RandomTuner, log records to file `matmul.log`
# You can use alternatives like XGBTuner.
tuner = autotvm.tuner.RandomTuner(task)
tuner.tune(
n_trial=10,
measure_option=measure_option,
callbacks=[autotvm.callback.log_to_file("matmul.log")],
)
# -
# Finally we apply history best from the cache file and check its correctness.
# We can call the function :code:`matmul` directly under the
# :any:`autotvm.apply_history_best` context. When we call this function,
# it will query the dispatch context with its argument and get the best config
# with the same argument.
#
#
# +
# apply history best from log file
with autotvm.apply_history_best("matmul.log"):
with tvm.target.Target("llvm"):
s, arg_bufs = matmul(N, L, M, "float32")
func = tvm.build(s, arg_bufs)
# check correctness
a_np = np.random.uniform(size=(N, L)).astype(np.float32)
b_np = np.random.uniform(size=(L, M)).astype(np.float32)
c_np = a_np.dot(b_np)
c_tvm = tvm.nd.empty(c_np.shape)
func(tvm.nd.array(a_np), tvm.nd.array(b_np), c_tvm)
tvm.testing.assert_allclose(c_np, c_tvm.asnumpy(), rtol=1e-2)
| _downloads/979311f7aa6c4f64c571283c99b06367/tune_simple_template.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import numpy as np
np.warnings.filterwarnings('ignore', category=np.VisibleDeprecationWarning) # Debugging: 'ignore' -> 'error'
# # Hydrogen bonds
import molsysmt as msm
molecular_system = msm.demo['pentalanine']['traj.h5']
molecular_system = msm.convert(molecular_system, to_form='molsysmt.MolSys')
msm.info(molecular_system)
# ## Acceptor atoms
msm.hbonds.acceptor_inclusion_rules
msm.hbonds.acceptor_exclusion_rules
acceptors=msm.hbonds.get_acceptor_atoms(molecular_system)
acceptors
msm.info(molecular_system, target='atom', selection='@acceptors')
acceptors=msm.hbonds.get_acceptor_atoms(molecular_system, inclusion_rules=["atom_name=='CB'"],
exclusion_rules=["atom_type=='N' and group_name=='ALA'"])
msm.info(molecular_system, target='atom', selection='@acceptors')
msm.select(molecular_system, selection="all bonded to atom_type=='H'")
acceptors=msm.hbonds.get_acceptor_atoms(molecular_system,
default_inclusion_rules=False, default_exclusion_rules=False,
inclusion_rules=["atom_type=='C'"],
exclusion_rules=["all bonded to atom_type=='H'"])
msm.info(molecular_system, target='atom', selection='@acceptors')
# ## Donor atoms
msm.hbonds.donor_inclusion_rules
msm.hbonds.donor_exclusion_rules
donors = msm.hbonds.get_donor_atoms(molecular_system)
donors
msm.info(molecular_system, target='atom', selection='@donors[0]')
donors_heavy_atom=donors[:,0]
msm.info(molecular_system, target='atom', selection='@donors_heavy_atom')
# # Buch
hbonds = msm.hbonds.buch(molecular_system)
len(hbonds)
hbonds.keys()
| docs/contents/hbonds/hbonds.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import json
import csv
hum_ensembl_entrez_map = {}
with open('mart_export_hum.txt', 'r') as mar_export_hum_file:
mar_export_hum = csv.reader(mar_export_hum_file,delimiter='\t')
for row in mar_export_hum:
if len(row[1]) > 0:
hum_ensembl_entrez_map[row[0]] = row[1]
mus_refseq_entrez_map = {}
mus_refseq_ensembl_map = {}
with open('mart_export_mus.txt', 'r') as mar_export_mus_file:
mar_export_mus = csv.reader(mar_export_mus_file,delimiter='\t')
for row in mar_export_mus:
# if len(row[1]) > 0 and len(row[2]) > 0:
# print "NM and NR in same row", row[1],row[2]
if len(row[1]) > 0:
if len(row[0]) > 0:
mus_refseq_entrez_map[row[1]] = row[0]
if len(row[3]) > 0:
mus_refseq_ensembl_map[row[1]] = row[3]
if len(row[2]) > 0:
if len(row[0]) > 0:
mus_refseq_entrez_map[row[2]] = row[0]
if len(row[3]) > 0:
mus_refseq_ensembl_map[row[2]] = row[3]
# +
with open('genes_list.json') as hum_genes_file:
hum_data = json.load(hum_genes_file)
with open('genes_list_GRCm38.txt') as mus_genes_file:
mus_data = json.load(mus_genes_file)
print hum_data[0]['ensembl_id'].split('.')[0]
print mus_data[0]
# -
'hello'.upper()
'3333.333'.split('.')[0]
for idx,gene in enumerate(hum_data):
hum_data[idx]['name'] = hum_data[idx]['name'].upper()
if gene['ensembl_id'].split('.')[0] in hum_ensembl_entrez_map:
hum_data[idx]['entrez_id'] = hum_ensembl_entrez_map[gene['ensembl_id'].split('.')[0]]
else:
hum_data[idx]['entrez_id'] = ""
#hum_data[idx]['entrez_full'] = "Entrez" + hum_ensembl_entrez_map[gene['ensembl_id'][:-2]]
with open('genes_list.json2', 'w') as genes_list:
genes_list.write(json.dumps(hum_data))
for idx,gene in enumerate(mus_data):
if gene['ensembl_id'] in mus_refseq_entrez_map:
mus_data[idx]['entrez_id'] = mus_refseq_entrez_map[gene['ensembl_id']]
else:
mus_data[idx]['entrez_id'] = ""
if gene['ensembl_id'] in mus_refseq_ensembl_map:
mus_data[idx]['ensembl_id_real'] = mus_refseq_ensembl_map[gene['ensembl_id']]
else:
mus_data[idx]['ensembl_id_real'] = ""
with open('genes_list_GRCm38.txt2', 'w') as genes_list:
genes_list.write(json.dumps(mus_data))
print mus_data[0]
print hum_data[0]
| static/data/pre_processed/BioMart_Genes_Setup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('..')
# # VorDiff
#
# ## Introduction
#
# Automatic differentiation is a tool for calculating derivatives using machine accuracy. It has several advantages over traditional methods of derivative calculations such as symbolic and finite differentiation. Automatic differentiation is useful for calculating complex derivatives where errors are more likely with classical methods. For instance , with finite differentiation, h values that are too small will lead to accuracy errors though floating point roundoff error, while h values that are too large will start making vastly inaccurate approximations.
#
# Automatic differentiation is useful due to its practicality in real world applications that involve thousands of parameters in a complicated function, which would take a long runtime as well as strong possibility for error in calculating the derivatives individually.
#
# Our package allows users to calculate derivatives of complex functions, some with many parameters, allowing machine precision.
#
# ## Background
#
# Essentially automatic differentiation works by breaking down a complicated function and performing a sequence of elementary arithmetic such as addition, subtraction, multiplication, and division as well as elementary functions like exp, log, sin, etc. These operations are then repeated by the chain rule and the derivatives of these sequences are calculated. There are two ways that automatic differentiation can be implemented - forward mode and reverse mode.
#
#
# ### 2.1 The Chain Rule
#
# The chain rule makes up a fundamental component of auto differentiation. The basic idea is:
# For univariate function, $$ F(x) = f(g(x))$$
#
# $$F^{\prime} = (f(g))^{\prime} = f^{\prime}(g(x))g^{\prime}(x)$$
#
# For multivariate function, $$F(x) = f(g(x),h(x))$$
#
# $$ \frac{\partial F}{\partial x}=\frac{\partial f}{\partial g} \frac{\partial g}{\partial x}+\frac{\partial f}{\partial h} \frac{\partial h}{\partial x}$$
#
# For generalized cases, if F is a combination of more sub-functions, $$F(x) = f(g_{1}(x), g_{2}(x), …, g_{m}(x))$$
#
# $$\frac{\partial F}{\partial x}=\sum_{i=1}^{m}\frac{\partial F}{\partial g_{i}} \frac{\partial g_{i}}{\partial x}$$
#
# For F is a function f(g): f: $R^n$ -> $R^m$ and g: $R^m$ -> $R^k$,
#
# $$\mathbf{J}_{\mathrm{gof}}(\mathbf{x})=\mathbf{J}_{\mathrm{g}}(\mathbf{f}(\mathbf{x})) \mathbf{J}_{\mathrm{f}}(\mathbf{x})$$
#
# where $J(f) =\left[\begin{array}{ccc}{\frac{\partial \mathbf{f}}{\partial x_{1}}} & {\cdots} & {\frac{\partial \mathbf{f}}{\partial x_{n}}}\end{array}\right]=\left[\begin{array}{ccc}{\frac{\partial f_{1}}{\partial x_{1}}} & {\cdots} & {\frac{\partial f_{1}}{\partial x_{n}}} \\ {\vdots} & {\ddots} & {\vdots} \\ {\frac{\partial f_{m}}{\partial x_{1}}} & {\cdots} & {\frac{\partial f_{m}}{\partial x_{n}}}\end{array}\right]$ is the Jacobian Matrix.
#
#
#
# ### 2.2 Auto Differentiation: Forward Mode
#
# The forward mode automatic differentiation is accomplished by firstly splitting the function process into one-by-one steps, each including only one basic operation. It focuses on calculating two things in each step, the value of scalar or vector x in $R^n$, and the 'seed' vector for the derivatives or Jacobian Matrix. From the first node, the value and derivatives will be calculated based on the values and derivatives of forward nodes. AD exploits the fact that every computer program, no matter how complicated, executes a sequence of elementary arithmetic operations (addition, subtraction, multiplication, division, etc.) and elementary functions (exp, log, sin, cos, etc.). By applying the chain rule repeatedly to these operations, derivatives of arbitrary order can be computed automatically, accurately to working precision, and using at most a small constant factor more arithmetic operations than the original program.
#
# The automatic differentiation is superior to analytic or symbolic differentiation because it could be computed on modern machines. It is also superior to numerical differentiation because numeric method is not precise but AD deals with machine precision problems properly.
#
# An example of computational graph and table for forward mode AD is shown as follows:
#
# \begin{align}
# f\left(x,y\right) =\sin\left(xy\right)
# \end{align}
# We will be evaluating the function at $f(1, 0)$
#
# Evaluation trace:
#
# | Trace | Elementary Function | Current Value | Elementary Function Derivative | $\nabla_{x}$ Value | $\nabla_{y}$ Value |
# | :---: | :-----------------: | :-----------: | :----------------------------: | :-----------------: | :-----------------: |
# | $x_{1}$ | $x_{1}$ | $1$ | $\dot{x}_{1}$ | $1$ | $0$ |
# | $x_{2}$ | $x_{2}$ | $0$ | $\dot{x}_{2}$ | $0$ | $1$ |
# | $x_{3}$ | $x_{1}x_{2}$ | $0$ | $\dot{x}_{2}$ | $0$ | $1$ |
#
# 
#
#
#
# ### 2.3 Reverse Mode
#
# The reverse mode automatic differentiation has a process similar to the forward mode auto differentiation, but has another reverse process. It does not apply the chain rule and only partial derivatives to a node are stored. First, for the forward process, the partial derivatives are stored for each node. For the reverse process, it starts with the differentiation to the last node, and then activations in the forward process are deployed in the differentiation step by step.
#
#
# ### 2.4 Forward Mode v.s. Reverse Mode
#
# Two main aspects can be considered when choosing between Forward and Reverse mode auto differentiation.
# * Memory Storage & Time of Computation
#
# The forward mode needs memory storage for values and derivatives for each node, while the reverse mode only needs to store the activations of partial differentiation to each node. The forward mode do the computation at the same time as the variable evaluation, while the reverse mode do the calculation in the backward process.
# * Input & Output Dimensionality
#
# If the input dimension is much larger than output dimension, then reverse mode is more attractive. If the output dimension is much larger than the input dimension, the forward mode is much computational cheaper.
# ## Installation
#
# Our package can be installed from our GitHub Repository at: https://github.com/VoraciousFour/cs207-FinalProject.
#
# After the package is installed, it needs to be imported into their workspace. Doing so, will automatically download any dependencies that are required by our package such as math or numpy. Then, the user can create and activate a virtual envitronment to use the package in.
#
# The user can set up and use our package using their terminal as follows.
#
# 1. Clone the VorDiff package from our Github Repository into your directory
# git clone https://github.com/VoraciousFour/cs207-FinalProject.git
# 2. Create and activate a virtual environment
# '''Installing virtualenv'''
# sudo easy_install virtualenv
# '''Creating the Virtual Environment'''
# virtualenv env
# '''Activating the Virtual Environment'''
# source env/bin/activate
# 3. Install required dependencies
# pip install -r requirements.txt
# 4. Importing VorDiff package for use
# import VorDiff
# ## How to use VorDiff
#
# Our Automatic Differentiation package is called VorDiff. The two main objects you will interact with are `AutoDiff` and `Operator`. In short, the user will first instantiate a scalar variable as an `AutoDiff` object, and then feed those variables to operators specified in the `Operator` object. The `Operator` object allows users to build their own functions for auto-differentiation. Simple operations (e.g. addition, multiplication, power) may be used normally. More complex functions (e.g. log, sin, cos) must use the operations defined in the `Operator` class. Lastly, the user may retrieve the values and first derivatives from the objects defined above by using the `get()` method.
#
# A short example is provided below:
# +
from VorDiff.autodiff import AutoDiff as ad
from VorDiff.operator import Operator as op
# Define scalar variables
x = ad.scalar(3.14159)
y = ad.scalar(0)
# Build functions
fx = op.sin(x) + 3
fy = op.exp(y) + op.log(y+1)
# Get values and derivates
print(fx.get())
print(fy.get())
# Define vector variables
x = ad.vector([2, 1, 0, 3, 6])
y = ad.vector([-1, 2, 1])
# Build functions
def f1(a, b):
return 2*a + b
def f2(a):
return a**2
def f3(a, b):
return b/a
def f4(a, b, c):
return op.sin(a+2*b-c)
def f5(a):
return op.exp(a)
# vector functions
def F1(x):
x1, x2, x3, x4, x5 = x
return f5(f4(x3,f3(f2(f1(x1,x5)),x2),x4))
def F2(y):
y1, y2, y3 = y
return f1(f2(f4(y1, y2, y3)),y3)
# Get values and derivatives
vec_functions = [F1(x), F2(y)]
vals = []
derivatives = []
for function in vec_functions:
vals.append(function.get_val())
derivatives.append(function.get_derivatives())
print("Values of function F1:", vals[0])
print("Derivatives of function F1 with respect to vector of x:", derivatives[0])
print("Values of function F2:", vals[1])
print("Derivatives of function F2 with respect to vector of y:", derivatives[1])
# -
# For the reverse mode automatic differentiation, the two main objects you will interact with are `ReverseAutoDiff` and `ReverseOperator`. In short, the user will first instantiate a scalar or vector variable as an `ReverseAutoDiff` object, and then feed those variables to operators specified in the `ReverseOperator` object. The `ReverseOperator` object allows users to build their own functions for reveres mode auto-differentiation. The basic operations (e.g. addition, subtraction, multiplication, division, power, negation) may be used normally with the implemenation of `ReverseScalar` and `ReverseVector`. More complex functions (e.g. sin, cos, tan, arcsin, arccos, arctan, log, sqrt, etc.) must use the operations defined in the `ReverseOperator` class. Lastly, the user may retrieve the values from the objects by using the `get()` method and the derivatives from the objects by using the `._gradient` attribute.
# +
import sys
sys.path.append('../')
from VorDiff.reverse_operator import ReverseOperator as rop
from VorDiff.reverse_autodiff import ReverseAutoDiff as rad
def create_reverse_vector(array):
x, y = rad.reverse_vector(array)
return x, y
# for scalar
x,y = create_reverse_vector([[1, 2, 3], [1,3,6]])
f = 1 / (x[1]) + rop.sin(1/x[1])
print("Partial derivative of f with respect to x:", rad.partial_scalar(f))
# for vector
x,y = create_reverse_vector([[1, 2, 3], [1,3,6]])
g = rop.cos(y)**2
print("Partial derivative of f with respect to vector of y:", rad.partial_vector(g,y))
# -
# ## Software Organization
#
# ### Directory Structure
# The package's directory will be structured as follows:
# ```
# VorDiff/
# __init__.py
# nodes/
# __init__.py
# scalar.py
# reverse_scalar.py
# reverse_vector.py
# vector.py
# tests/
# __init__.py
# test_autodiff.py
# test_node.py
# test_operator.py
# test_reverse_autodiff.py
# test_reverse_operator.py
# test_reverse_scalar.py
# test_reverse_vector.py
# test_scaler.py
# test_vector.py
# autodiff.py
# operator.py
# reverse_autodiff.py
# reverse_operator.py
# README.md
# ...
# demo/
# demo_reverse.py
# demo_scalar.py
# demo_vector.py
# docs/
# ...
# ```
# ### Modules
# - VorDiff: The VorDiff module contains the operator class to be directly used by users to evaluate functions and calculate their derivatives, and an autodiff class that acts as the central interface for automatic differentiation.
#
# - Nodes: The Nodes module contains the the scalar and vector classes, which define the basic operations that can be performed on scalar and vector variables for the autodiff class.
#
# - Test_Vordiff: The Test_Vordiff module contains the test suite for this project. TravisCI and CodeCov are used to test our operator classes, node classes, and auto-differentiator.
#
# - Demo: The Demo module contains python files demonstrating how to perform automatic differentiation with the implemented functions.
#
# ### Testing
# In this project we use TravisCI to perform continuous integration testing and CodeCov to check the code coverage of our test suite. The status us TravisCI and CodeCov can be found in README.md, in the top level of our package. Since the test suite is included in the project distribution, users can also install the project package and use pytest and pytest-cov to check the test results locally.
#
# ### Distribution:
# Our open-source VorDiff package will be uploaded to PyPI by using twine because it uses a verified connection for secure authentication to PyPI over HTTPS. Users will be able to install our project package by using the convential `pip install VorDiff`.
#
#
#
# ## Implementation
#
# ### Scalar
# The `Scalar` class represents a single scalar node in the computational graph of a function. It implements the interface for user defined scalar variables. The object contains two hidden attributes, `._val` and `._der`, which can be retrieved with the `get()` method.
'''Docstrings hidden'''
class Scalar():
def __init__(self, value, *kwargs):
self._val = value
if len(kwargs) == 0:
self._der = 1
else:
self._der = kwargs[0]
def get(self):
return self._val, self._der
def __add__(self, other):
try:
return Scalar(self._val+other._val, self._der+other._der)
except AttributeError:
return self.__radd__(other)
def __radd__(self, other):
return Scalar(self._val+other, self._der)
'''etc'''
# ### Vector
# The `Vector` class represents a single vector variable. Vectors are comprised of `Element` objects, which implement much of the computation necessary for vector automatic differentiation. Vectors contain two hidden attributes: a list `_elements`, and a numpy array `_jacob`.
# +
import numpy as np
'''Docstrings hidden'''
class Vector():
def __init__(self, vec, *kwargs):
self._vec = np.array(vec)
if len(kwargs) == 0:
self._jacob = np.eye(len(vec))
else:
self._jacob = np.array(kwargs[0])
elements = []
for i in range(len(vec)):
elements.append(Element(self._vec[i], self._jacob[i]))
self._elements = elements
def __getitem__(self, idx):
return self._elements[idx]
class Element():
"""
The Element object has an evaluated value (it can be the value of function compositions with
the input of user defined values) and a list of current derivatives with respect to each variable.
"""
def __init__(self, val, jacob):
self._val = val
self._jacob = np.array(jacob)
def get_val(self):
return self._val
def get_derivatives(self):
return self._jacob
def __add__(self, other):
try:
val = self._val+other._val
jacob = self._jacob+other._jacob
return Element(val, jacob)
except AttributeError:
return self.__radd__(other)
'''etc'''
# -
# ### Operator
# The operator class contains all mathematical operations that users can call to build their functions. Each function returns a `Scalar` object, a `Vector` object, or a numeric constant, depending on the input type. Each function raises an error if its input falls outside its domain. All functions in the class are static.
# +
from VorDiff.nodes.scalar import Scalar
from VorDiff.nodes.vector import Vector,Element
'''Docstrings hidden'''
class Operator():
@staticmethod
def sin(x):
try:
return Element(np.sin(x._val), np.cos(x._val)*x._jacob)
except AttributeError: # If constant
try: # If scalar variable
return Scalar(np.sin(x._val), x._der*np.cos(x._val))
except AttributeError: # If constant
return np.sin(x)
@staticmethod
def cos(x):
try:
return Element(np.cos(x._val), -np.sin(x._val)*x._jacob)
except AttributeError: # If constant
try: # If scalar variable
return Scalar(np.cos(x._val), -np.sin(x._val)*x._der)
except AttributeError: # If constant
return np.cos(x)
'''etc'''
# -
# ### AutoDiff
# The `AutoDiff` class will allow the user to easily create variables and build auto-differentiable functions, without having to interface with the any of the node classes. It will make use of the auto-differentiator much more intuitive for the user.
# +
from VorDiff.nodes.scalar import Scalar
from VorDiff.nodes.vector import Vector, Element
import numpy as np
class AutoDiff():
'''
The AutoDiff class allows users to define Scalar variables and
interface with the auto-differentiator.
'''
@staticmethod
def scalar(val):
return Scalar(val, 1)
def element(val,jacob):
return Element(val,jacob)
@staticmethod
def vector(vec):
return Vector(vec, np.eye(len(vec)))
# -
# # Advanced Features (Reverse AD)
# ### ReverseScalar
# The ReverseScalar class allows a single user defined variable capable of reverse mode of automatic differentiation. The ReverseScalar objects contain two hidden attributes, ._val and ._gradient. The attribute ._val and the function compute_gradient can be retrieved with the get() method. The dunder methods are also implemented in this class so that user can do the basic reverse mode computation and operations with the objects.
class ReverseScalar():
def __init__(self, val: float):
self._val = val
self._gradient = 1
self._children = {}
def get(self):
return self._val, self.compute_gradient()
def compute_gradient(self):
pass
def __add__(self, other):
pass
'etc'
# ### ReverseVector
# The ReverseVector class allows a vector of multiple user defined variables capable of reverse mode of automatic differentiation. The ReverseVector objects contain two hidden attributes, ._val and ._gradient. The attribute ._val and can be retrieved with the get() method. The partial derivatives with respect to the user defined variables can be retrieved by just calling the attribute ._gradient of the objects. The dunder methods are also implemented in this class so that user can do the basic reverse mode computation and operations with the objects.
class ReverseVector():
def __init__(self, vals: list):
self._val = np.array(vals)
self._children = {}
self._gradient = np.zeros(len(vals))
self._reverse_scalars = [ReverseScalar(val) for val in vals]
def __getitem__(self, idx):
return self._reverse_scalars[idx]
def get(self):
return self._val
def _init_children(self):
pass
def compute_gradient(self, var):
pass
def __add__(self, other):
pass
'''etc'''
# ### Reverse AutoDiff
# The `ReverseAutoDiff` class allows users to create variables in the reverse mode and build auto-differentiable functions, without having to interface with the `Node` class. It will make use of the reverse auto-differentiator much more intuitive for the user. The `reverse_scalar` and `reverse_vector` function specifically allows users to create variables and functions. The `partial_scalar` and `partial_vector` allows users to calculate the derivatives in a single-vriable function and the partial derivatives in a multi-variables function.
#
# The sturcture of this class is:
class ReverseAutoDiff():
@staticmethod
def reverse_scalar(val):
pass
@staticmethod
def reverse_vector(vals):
pass
def partial_vector(f, x):
pass
def partial_scalar(f):
pass
# ### Reverse Operator
#
# The `ReverseOperator` class contains all mathematical operations that users can call to build their functions. Each function returns a `Vector` object, `Scalar` object, or a numeric constant, depending on the input type. Each function raises an erro if its input falls outside its domain.
#
# In this implementation, we include the following elementary functions. Derivatives are calculated with the with the reverse mode.
class ReverseOperator:
@staticmethod
def sin(x):
pass
@staticmethod
def cos(x):
pass
@staticmethod
def tan(x):
pass
@staticmethod
def sqrt(x):
pass
'''etc'''
# ## Future Features
#
# ### 1. Option for higher-order derivatives
#
# There are plenty of ways we could improve our package. The first is to grant users the option to compute higher-order derivatives like Hessians. We can recursively apply AD first to the target function, i.e., producing the first-order derivative, then moving the operations of the first-order derivatives into a new computational graph then applying AD again. In short, higher order derivatives would be accomplished by repeatedly applying automatic differentiation to function and its derivatives.
#
# ### 2. Application using AD library to find the roots of functions
#
# A second way we could extend our work is by writing a separate library to find the roots of given functions. For example, this could include an implementation of Newton’s Method that calculates the exact Hessian matrix of a function using AD to get second-order partial derivatives. We would use Newton's Method to search for the approximations by calculating the exact Hessian matrix of the function using AD to get the second-order partial derivatives.
#
# ### 3. Backpropagation in neural networks
#
# We can also extend our implementation of automatic differentiation to the neural networks. Neural networks are able to gradually increase accuracy with every training session through the process of gradient descent. In gradient descent, we aim to minimize the loss (i.e. how inaccurate the model is) through tweaking the weights and biases.
#
# By finding the partial derivative of the loss function, we know how much (and in what direction) we must adjust our weights and biases to decrease loss. In that series, we calculate the derivative mean squared error loss function of a single-neuron neural network.
#
# For computers to calculate the partial derivatives of an expression in neural networks, we can implement the automatic differentiation for both forward pass and back propagation. Then we can calculate the partial derivatives in both scalar and vector modes.
#
| docs/documentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torchvision import datasets, transforms
torch.manual_seed(4242)
train_loader = torch.utils.data.DataLoader(
datasets.MNIST('../data/p1ch2/mnist', train=True, download=True,
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])),
batch_size=64, shuffle=True)
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 10, kernel_size=5)
self.conv2 = nn.Conv2d(10, 20, kernel_size=5)
self.conv2_drop = nn.Dropout2d()
self.fc1 = nn.Linear(320, 50)
self.fc2 = nn.Linear(50, 10)
def forward(self, x):
x = F.relu(F.max_pool2d(self.conv1(x), 2))
x = F.relu(F.max_pool2d(self.conv2_drop(self.conv2(x)), 2))
x = x.view(-1, 320)
x = F.relu(self.fc1(x))
x = F.dropout(x, training=self.training)
x = self.fc2(x)
return F.log_softmax(x, dim=1)
model = Net()
optimizer = optim.SGD(model.parameters(), lr=0.01, momentum=0.5)
for epoch in range(10):
for batch_idx, (data, target) in enumerate(train_loader):
optimizer.zero_grad()
output = model(data)
loss = F.nll_loss(output, target)
loss.backward()
optimizer.step()
print('Current loss', float(loss))
torch.save(model.state_dict(), '../data/p1ch2/mnist/mnist.pth')
pretrained_model = Net()
pretrained_model.load_state_dict(torch.load('../data/p1ch2/mnist/mnist.pth'))
| p1ch2/4_mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
train_df = pd.read_parquet('../../input/train.parquet')
test_df = pd.read_parquet('../../input/test.parquet')
# +
# [x for x in train_df.columns]
| notebooks/EDA/EDA-003-Features_For_Baseline_Models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from textblob import TextBlob
import spacy
data = pd.read_csv('data/joe_biden_tweets_2020.csv')
data.head()
# - Using TextBlob evaluate the sentiment and polarization of each tweet
# - show the five most positive tweets (based on polarity)
# - show the five most negative tweets (based on polarity)
# - show the five most subjective tweets
# - Using spacy perform named entity extraction
# - What are the five most frequent `PERSON` entities
# - What are the five most frequent `EVENT` entities
sentiment = data.tweet.apply(lambda tweet: TextBlob(tweet).sentiment)
data['sentiment'] = sentiment.apply(lambda x: x.polarity)
data['subjectivity'] = sentiment.apply(lambda x: x.subjectivity)
data.head()
data.sort_values('sentiment', ascending=False).tweet.head()
list(_)
data.sort_values('sentiment', ascending=False).tweet.iloc[-5:]
list(_)
data.sort_values('subjectivity', ascending=False).tweet.head()
list(_)
nlp = spacy.load('en_core_web_sm')
def extract_entities(text):
doc = nlp(text)
return [(ent.text, ent.label_) for ent in doc.ents]
# test
extract_entities(data.tweet.iloc[0])
data['entities'] = data.tweet.apply(extract_entities)
entities = pd.DataFrame(data.entities.sum(), columns=['text', 'entity'])
entities['count'] = 1
entities.head()
counts = entities.groupby(['entity', 'text'])['count'].count()
counts.loc['PERSON'].sort_values(ascending=False).head()
counts.loc['EVENT'].sort_values(ascending=False).head()
| 34-problem-solution_semantics_sentiment_and_named_entities.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #Real Estate Predictor
import pandas as pd
housing = pd.read_csv("data.csv")
housing.head()
# ## We should check info of the dataset to check weather there is any null set
housing.info()
housing.describe()
# Now let me check the value count of CHAS variable
housing['CHAS'].value_counts()
import matplotlib.pyplot as plt
# %matplotlib inline
housing.hist(bins =50, figsize = (20,15))
# ## Train-Test Splitting
# for learning purpose
import numpy as np
def split_test_train(data, test_ratio):
np.random.seed(42)
shuffled = np.random.permutation(len(data))
test_set_size = int(len(data) * test_ratio)
test_indices = shuffled[:test_set_size]
train_indices = shuffled[test_set_size:]
return data.iloc[train_indices], data.iloc[test_indices]
# +
# train_set, test_set = split_test_train(housing, 0.2)
# -
# The function that we have generated above is already available in scikit-learn package so we are going to comment it and use the same same functyion from scikit-learn module
from sklearn.model_selection import train_test_split
train_set, test_set = train_test_split(housing, test_size = 0.2, random_state = 42)
print(f"Rows in train set:{len(train_set)}\nRows in test set:{len(test_set)}")
| Real Estate Project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Tuples
#
# In Python tuples are very similar to lists, however, unlike lists they are *immutable* meaning they can not be changed. You would use tuples to present things that shouldn't be changed, such as days of the week, or dates on a calendar.
#
# In this section, we will get a brief overview of the following:
#
# 1.) Constructing Tuples
# 2.) Basic Tuple Methods
# 3.) Immutability
# 4.) When to Use Tuples.
#
# You'll have an intuition of how to use tuples based on what you've learned about lists. We can treat them very similarly with the major distinction being that tuples are immutable.
#
# ## Constructing Tuples
#
# The construction of a tuples use () with elements separated by commas. For example:
# Can create a tuple with mixed types
t = (1,2,3)
# Check len just like a list
len(t)
# +
# Can also mix object types
t = ('one',2)
# Show
t
# -
# Use indexing just like we did in lists
t[0]
# Slicing just like a list
t[-1]
# ## Basic Tuple Methods
#
# Tuples have built-in methods, but not as many as lists do. Lets look at two of them:
# Use .index to enter a value and return the index
t.index('one')
# Use .count to count the number of times a value appears
t.count('one')
# ## Immutability
#
# It can't be stressed enough that tuples are immutable. To drive that point home:
t[0]= 'change'
# Because of this immutability, tuples can't grow. Once a tuple is made we can not add to it.
t.append('nope')
# ## When to use Tuples
#
# You may be wondering, "Why bother using tuples when they have fewer available methods?" To be honest, tuples are not used as often as lists in programming, but are used when immutability is necessary. If in your program you are passing around an object and need to make sure it does not get changed, then tuple become your solution. It provides a convenient source of data integrity.
#
# You should now be able to create and use tuples in your programming as well as have an understanding of their immutability.
#
# Up next Files!
| Complete-Python-Bootcamp-master/Tuples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import os
import tensorflow as tf
from google.protobuf import json_format
from tensorflow_serving.apis import prediction_log_pb2
from bulk_inferrer import executor
from tfx.proto import bulk_inferrer_pb2
from tfx.types import artifact_utils
from tfx.types import standard_artifacts
# -
executor.__file__
# !pwd
_source_data_dir = '/home/jupyter/tfx_0.23.0/tfx/tfx/components/testdata'
_output_data_dir = '/home/jupyter/dataflow_example'
# +
_examples = standard_artifacts.Examples()
_examples.uri = os.path.join(_source_data_dir, 'csv_example_gen')
_examples.split_names = artifact_utils.encode_split_names(['unlabelled'])
_model = standard_artifacts.Model()
_model.uri = os.path.join(_source_data_dir, 'trainer/current')
_model_blessing = standard_artifacts.ModelBlessing()
_model_blessing.uri = os.path.join(_source_data_dir, 'model_validator/blessed')
_model_blessing.set_int_custom_property('blessed', 1)
_inference_result = standard_artifacts.InferenceResult()
_prediction_log_dir = os.path.join(_output_data_dir, 'prediction_logs')
_inference_result.uri = _prediction_log_dir
# Create context
_tmp_dir = os.path.join(_output_data_dir, '.temp')
_context = executor.Executor.Context(
tmp_dir=_tmp_dir, unique_id='2')
# -
prediction_log_pb2.PredictionLog()
def _get_results(prediction_log_path):
results = []
filepattern = os.path.join(
prediction_log_path,
executor._PREDICTION_LOGS_DIR_NAME) + '-?????-of-?????.gz'
for f in tf.io.gfile.glob(filepattern):
record_iterator = tf.compat.v1.python_io.tf_record_iterator(
path=f,
options=tf.compat.v1.python_io.TFRecordOptions(
tf.compat.v1.python_io.TFRecordCompressionType.GZIP))
for record_string in record_iterator:
prediction_log = prediction_log_pb2.PredictionLog()
prediction_log.MergeFromString(record_string)
results.append(prediction_log)
return results
# +
input_dict = {
'examples': [_examples],
'model': [_model],
'model_blessing': [_model_blessing],
}
output_dict = {
'inference_result': [_inference_result],
}
# Create exe properties.
exec_properties = {
'data_spec':
json_format.MessageToJson(
bulk_inferrer_pb2.DataSpec(), preserving_proto_field_name=True),
'model_spec':
json_format.MessageToJson(
bulk_inferrer_pb2.ModelSpec(),
preserving_proto_field_name=True),
'component_id':
'test_component' #self.component_id,
}
# Run executor.
bulk_inferrer = executor.Executor(_context)
bulk_inferrer.Do(input_dict, output_dict, exec_properties)
# -
result[0]
result = _get_results(_prediction_log_dir)
json_format.MessageToJson(result[0])
json_format.MessageToDict(result[0])
{
"namespace": "example.avro",
"type": "record",
"name": "User",
"fields": [
{"name": "name",
"type": "string"},
{"name": "favorite_number",
"type": ["int", "null"]},
{"name": "favorite_color",
"type": ["string", "null"]}
]
}
type(json_format.MessageToDict(result[0]))
result[0].classify_log.response
table_spec = 'res-nbcupea-dev-ds-sandbox-001:oneapp.test_bigquery_beam'
table_schema = 'source:STRING, quote:STRING'
# +
import apache_beam as beam
with beam.Pipeline() as p:
quotes = p | beam.Create([
{
'source': '<NAME>', 'quote': 'My life is my message.'
},
{
'source': 'Yoda', 'quote': "Do, or do not. There is no 'try'."
},
])
quotes | beam.io.WriteToBigQuery(
table_spec,
schema=table_schema,
write_disposition=beam.io.BigQueryDisposition.WRITE_TRUNCATE,
create_disposition=beam.io.BigQueryDisposition.CREATE_IF_NEEDED,
custom_gcs_temp_location='gs://benk-dev-sandbox/temp'
)
# -
# + [markdown] id="R_vFbXN2T7oo"
# ##### Copyright © 2020 The TensorFlow Authors.
# + id="NhL09apzT-W_"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://urldefense.com/v3/__https://www.apache.org/licenses/LICENSE-2.0*5Cn__;JQ!!PIZeeW5wscynRQ!55X7LkP-oqeRJ6UUmtPVv4vn3HLBOqlFNpBPTKq9j6A_FsUVIeCO-4Rxhel3RfE$ #
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="23R0Z9RojXYW"
# # TFX – Developing With Apache Beam
#
# [Apache Beam](https://urldefense.com/v3/__https://www.tensorflow.org/tfx/guide/beam__;!!PIZeeW5wscynRQ!55X7LkP-oqeRJ6UUmtPVv4vn3HLBOqlFNpBPTKq9j6A_FsUVIeCO-4RxlMaOx6c$ ) is an open source, unified model for defining both batch and streaming data-parallel processing pipelines. TFX uses Apache Beam to implement data-parallel pipelines. The pipeline is then executed by one of Beam's supported distributed processing back-ends or "runners", which include [Dataflow](https://urldefense.com/v3/__https://cloud.google.com/dataflow__;!!PIZeeW5wscynRQ!55X7LkP-oqeRJ6UUmtPVv4vn3HLBOqlFNpBPTKq9j6A_FsUVIeCO-4Rx_xDiz6Q$ ), [Apache Flink](https://urldefense.com/v3/__https://flink.apache.org/__;!!PIZeeW5wscynRQ!55X7LkP-oqeRJ6UUmtPVv4vn3HLBOqlFNpBPTKq9j6A_FsUVIeCO-4RxUIpCch4$ ), [Apache Spark](https://urldefense.com/v3/__https://spark.apache.org/__;!!PIZeeW5wscynRQ!55X7LkP-oqeRJ6UUmtPVv4vn3HLBOqlFNpBPTKq9j6A_FsUVIeCO-4RxVA2PkcU$ ), [Apache Samza](https://urldefense.com/v3/__http://samza.apache.org/__;!!PIZeeW5wscynRQ!55X7LkP-oqeRJ6UUmtPVv4vn3HLBOqlFNpBPTKq9j6A_FsUVIeCO-4Rx5goppac$ ), and others.
#
# TFX is designed to be scalable to very large datasets which require substantial resources. Distributed pipeline frameworks such as Apache Beam offer the ability to distribute processing across compute clusters and apply the resources required. Many of the standard TFX components use Apache Beam, and custom components that you may write may also benefit from using Apache Beam for distibuted processing.
#
# This notebook introduces the concepts and code patterns for developing with the Apache Beam Python API. This is especially important for developers who will be creating custom TFX components which may require substantial computing resources, and will benefit from distributed processing using beam.
# + [markdown] id="EnOzbFhBKqwf"
# ### [DirectRunner](https://beam.apache.org/documentation/runners/direct/)
#
# In this notebook, to avoid the requirement for a distributed processing cluster we will instead use Beam's [`DirectRunner`](https://urldefense.com/v3/__https://beam.apache.org/documentation/runners/direct/__;!!PIZeeW5wscynRQ!55X7LkP-oqeRJ6UUmtPVv4vn3HLBOqlFNpBPTKq9j6A_FsUVIeCO-4RxE4HG0-g$ ). The `DirectRunner` is also very convenient when developing Beam pipelines. The `DirectRunner` executes pipelines on your machine and is designed to validate that pipelines adhere to the Beam programming model as closely as possible. Instead of focusing on efficient pipeline execution, the `DirectRunner` performs additional checks to ensure that users do not rely on semantics that are not guaranteed by the model.
# + [markdown] id="2GivNBNYjb3b"
# ## Setup
# First, we install the necessary packages, download data, import modules and set up paths.
#
# ### Install Apache Beam
# + id="7zhcJBLoMXQE"
# !pip install -q -U \
# apache-beam \
# apache-beam[interactive]
# + [markdown] id="N-ePgV0Lj68Q"
# ### Import packages
# We import necessary packages, including Beam.
# -
# !pip install -q -U graphviz
# + id="YIqpWK9efviJ"
import apache_beam as beam
from apache_beam import pvalue
from apache_beam.runners.interactive.display import pipeline_graph
import graphviz
# + [markdown] id="p4XfH7dyRZtd"
# Check the version.
# + id="0uuFfIiHviyW"
print('Beam version: {}'.format(beam.__version__))
# + [markdown] id="nCb-bS_eMkMf"
# ## Beam Python Syntax
#
# Beam uses a [special Python syntax](https://urldefense.com/v3/__https://beam.apache.org/documentation/programming-guide/*applying-transforms__;Iw!!PIZeeW5wscynRQ!55X7LkP-oqeRJ6UUmtPVv4vn3HLBOqlFNpBPTKq9j6A_FsUVIeCO-4Rxw19zxq8$ ) to define and invoke transforms. For example, in this line:
#
# >`result = pass_this | 'name this step' >> to_this_call`
#
# The method `to_this_call` is being invoked and passed the object called `pass_this`, and this operation will be referred to as [name this step](https://urldefense.com/v3/__https://stackoverflow.com/questions/50519662/what-does-the-redirection-mean-in-apache-beam-python__;!!PIZeeW5wscynRQ!55X7LkP-oqeRJ6UUmtPVv4vn3HLBOqlFNpBPTKq9j6A_FsUVIeCO-4RxMFC5UkE$ ) in a stack trace or pipeline diagram. The result of the call to `to_this_call` is returned in `result`. You will often see stages of a pipeline chained together like this:
#
# >`result = apache_beam.Pipeline() | 'first step' >> do_this_first() | 'second step' >> do_this_last()`
#
# and since that started with a new pipeline, you can continue like this:
#
# >`next_result = result | 'doing more stuff' >> another_function()`
# + [markdown] id="qAika7-6gLvI"
# ## Create a Beam Pipeline
#
# Create a pipeline, including a simple [`PCollection`](https://urldefense.com/v3/__https://beam.apache.org/releases/javadoc/2.1.0/org/apache/beam/sdk/values/PCollection.html__;!!PIZeeW5wscynRQ!55X7LkP-oqeRJ6UUmtPVv4vn3HLBOqlFNpBPTKq9j6A_FsUVIeCO-4Rxgmi9VMc$ ) and a [`ParDo()`](https://urldefense.com/v3/__https://beam.apache.org/releases/javadoc/2.0.0/org/apache/beam/sdk/transforms/ParDo.html__;!!PIZeeW5wscynRQ!55X7LkP-oqeRJ6UUmtPVv4vn3HLBOqlFNpBPTKq9j6A_FsUVIeCO-4RxfbsGH5U$ ) transform.
#
# * A `PCollection<T>` is an **immutable collection** of values of type `T`. A `PCollection` can contain either a bounded or unbounded number of elements. Bounded and unbounded `PCollections` are produced as the output of `PTransforms` (including root `PTransforms` like `Read` and `Create`), and can be passed as the inputs of other `PTransforms`.
# * `ParDo` is the core **element-wise** transform in Apache Beam, invoking a user-specified function on each of the elements of the input `PCollection` to produce zero or more output elements, all of which are collected into the output `PCollection`.
#
# This can be done in different ways, depending on coding style preferences. First, use the `.run()` method.
# + id="X_XV_cm2_E1x"
first_pipeline = beam.Pipeline()
lines = (first_pipeline
| "Create" >> beam.Create(["Hello", "World", "!!!"]) # PCollection
| "Print" >> beam.ParDo(print)) # ParDo transform
result = first_pipeline.run()
result.state
# + [markdown] id="qx1xFZ-BYccz"
# Display the structure of this pipeline.
# + id="bdnEk2p3Jnfs"
def display_pipeline(pipeline):
graph = pipeline_graph.PipelineGraph(pipeline)
return graphviz.Source(graph.get_dot())
display_pipeline(first_pipeline)
# + [markdown] id="JSaFbVyIYNIA"
# Using a different coding style, invoke run inside a `with` block.
# + id="P4O8EAYNULbF"
with beam.Pipeline() as with_pipeline:
lines = (with_pipeline
| "Create" >> beam.Create(["Hello", "World", "!!!"])
| "Print" >> beam.ParDo(print))
display_pipeline(with_pipeline)
# + [markdown] id="yLJf3Zr5STWR"
# Notice that both pipelines and their outputs are the same.
# + [markdown] id="bMYiWcBtIBF7"
# ### Now You Try
#
# This is a good point to stop reading and try writing code yourself, as a way to help you learn.
#
# **Exercise 1 — Creating and Running Your Beam Pipeline**
#
# 1. Build a Beam pipeline that creates a PCollection containing integers 0 to 10 and prints them.
# 2. Add a step in the pipeline to square each item.
# 3. Display the pipeline.
#
# *Warning*: the `ParDo()` method must either return `None` or a list.
# + id="0aPlwJjQKmo6"
# + id="8f_Nf9XkH-vn"
# + id="W46JR_uDH-lz"
# + id="4f8NJsj-K1hG"
# + id="Ai6SmbkuK1SC"
# + [markdown] id="J-b7ZdFYLoH2"
# 
# + [markdown] id="Vbw50WdgMsLU"
# **Solution**:
# + id="PSUBzUGqfX4K"
def passthrough(label, x): # Utility for printing and returning the element
print(label, x)
return x
# This includes printing stages for illustration.
with beam.Pipeline() as with_pipeline:
lines = (with_pipeline
| "Create" >> beam.Create(range(10 + 1))
| 'Print0' >> beam.Map(lambda x: passthrough('x =', x))
| "Square" >> beam.ParDo(lambda x: [x ** 2])
| "Print1" >> beam.Map(lambda x: passthrough('x^2 =', x)))
display_pipeline(with_pipeline)
# + [markdown] id="C6lzLoIQz1Oe"
# # Core Transforms
# ---
# Beam has a set of core transforms on data that is contained in `PCollections`. In the cells that follow, explore several core transforms and observe the results in order to develop some understanding and intuition for what each transform does.
#
# ## [Map](https://beam.apache.org/documentation/transforms/python/elementwise/map/)
# The `Map` transform applies a simple 1-to-1 mapping function over each element in the collection. `Map` accepts a function that returns a single element for every input element in the `PCollection`. You can pass functions with multiple arguments to `Map`. They are passed as additional positional arguments or keyword arguments to the function.
#
# First, compare the results of a `ParDo` transform and a `Map` transform for a multiply operation.
# + id="bkzO7gANza49"
with beam.Pipeline() as pipeline:
lines = (pipeline
| "Create" >> beam.Create([1, 2, 3])
| 'Print0' >> beam.Map(lambda x: passthrough('x =', x))
| "Multiply" >> beam.ParDo(lambda number: [number * 2]) # ParDo with integers
| 'Print1' >> beam.Map(lambda x: passthrough('x * 2 =', x)))
# + id="dZWhKgqn02Ow"
with beam.Pipeline() as pipeline:
lines = (pipeline
| "Create" >> beam.Create([1, 2, 3])
| 'Print0' >> beam.Map(lambda x: passthrough('x =', x))
| "Multiply" >> beam.Map(lambda number: number * 2) # Map with integers
| 'Print1' >> beam.Map(lambda x: passthrough('x * 2 =', x)))
# + [markdown] id="gh1R4stdUglj"
# Notice that the results are the same. Now try it with a split operation.
# + id="7bsEdlKj0Wce"
with beam.Pipeline() as pipeline:
lines = (pipeline
| "Create" >> beam.Create(["Hello Beam", "This is cool"])
| 'Print0' >> beam.Map(lambda x: passthrough('Sentence:', x))
| "Split" >> beam.ParDo(lambda sentence: sentence.split()) # ParDo with strings
| 'Print1' >> beam.Map(lambda x: passthrough('Word:', x)))
# + id="GY490-duaQFh"
with beam.Pipeline() as pipeline:
lines = (pipeline
| "Create" >> beam.Create(["Hello Beam", "This is cool"])
| 'Print0' >> beam.Map(lambda x: passthrough('Sentence:', x))
| "Split" >> beam.Map(lambda sentence: sentence.split()) # Map with strings
| 'Print1' >> beam.Map(lambda x: passthrough('Words:', x)))
# + [markdown] id="USH3bn-VcHsl"
# Notice that `ParDo` returned individual elements, while `Map` returned lists.
# + [markdown] id="wtcELhYDUsY0"
# ## DoFn and [FlatMap](https://beam.apache.org/documentation/transforms/python/elementwise/flatmap/)
# Now try working with a `DoFn` in a `ParDo`, and compare that to using `FlatMap`.
# -
import apache_beam as beam
# + id="t0-gNKm70oJD"
class BreakIntoWordsDoFn(beam.DoFn):
def process(self, element):
return element.split()
with beam.Pipeline() as pipeline:
lines = (pipeline
| "Create" >> beam.Create(["Hello Beam", "This is cool"])
| 'Print0' >> beam.Map(lambda x: passthrough('Sentence:', x))
| "Split" >> beam.ParDo(BreakIntoWordsDoFn()) # Apply a DoFn with a process method
| 'Print1' >> beam.Map(lambda x: passthrough('Word:', x)))
# + id="99n6mPVF1C1_"
with beam.Pipeline() as pipeline:
lines = (pipeline
| "Create" >> beam.Create(["Hello Beam", "This is cool"])
| 'Print0' >> beam.Map(lambda x: passthrough('Sentence:', x))
| "Split" >> beam.FlatMap(lambda sentence: sentence.split()) # Compare to a FlatMap
| 'Print1' >> beam.Map(lambda x: passthrough('Word:', x)))
# + [markdown] id="kD0CwIu5220t"
# Again, notice that the results are the same. There is often more than one way to generate the same result!
# + [markdown] id="Gbthg5uU1X4a"
# ## [GroupByKey](https://beam.apache.org/documentation/transforms/python/aggregation/groupbykey/)
# `GroupByKey` takes a keyed collection of elements and produces a collection where each element consists of a key and all values associated with that key.
#
# `GroupByKey` is a transform for processing collections of key/value pairs. It’s a parallel reduction operation, analogous to the Shuffle phase of a Map/Shuffle/Reduce-style algorithm. The input to `GroupByKey` is a collection of key/value pairs that represents a multimap, where the collection contains multiple pairs that have the same key, but different values. Given such a collection, you use `GroupByKey` to collect all of the values associated with each unique key.
#
# `GroupByKey` is a good way to aggregate data that has something in common. For example, if you have a collection that stores records of customer orders, you might want to group together all the orders from the same postal code (wherein the “key” of the key/value pair is the postal code field, and the “value” is the remainder of the record).
# + id="PTX7JoS11LQd"
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.Create(['apple', 'ball', 'car', 'bear', 'cheetah', 'ant'])
| 'Words' >> beam.Map(lambda x: passthrough('Word:', x))
| beam.Map(lambda word: (word[0], word))
| 'Keyed' >> beam.Map(lambda x: passthrough('Keyed Tuple:', x))
| beam.GroupByKey()
| 'Groups' >> beam.Map(lambda x: passthrough('Grouped:', x)))
# + [markdown] id="DKNvm2H7hrsw"
# ### Now You Try
#
# This is a good point to stop reading and try writing code yourself, as a way to help you learn.
#
# **Exercise 2 — Group Items by Key**
#
# 1. Build a Beam pipeline that creates a PCollection containing integers 0 to 10 and prints them.
# 2. Add a step in the pipeline to add a key to each item that will indicate whether it is even or odd.
# 3. Use `GroupByKey` to group even items together and odd items together.
# + id="KwinUX5-hrs0"
def passthrough(label, value):
print(label, value)
return value
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.Create(range(11))
| beam.Map(lambda x: passthrough('num', x))
| beam.Map(lambda x: (x%2, x))
| beam.GroupByKey()
| beam.Map(lambda x: passthrough('grouped', x))
)
# + id="9rm6a2W7hrs3"
# + id="3ituEBJzhrs5"
# + id="IqjIVuc1hrs8"
# + id="BFW9cwPMhrtA"
# + [markdown] id="OkrvfvXohrtB"
# 
# + [markdown] id="euNLIh_ThrtC"
# **Solution**:
# + id="K3_dXXXIiM1w"
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.Create(range(10 + 1))
| 'Number' >> beam.Map(lambda x: passthrough('Number:', x))
| beam.Map(lambda x: ("odd" if x % 2 else "even", x))
| 'Even/Odd' >> beam.Map(lambda x: passthrough('Even/Odd:', x))
| beam.GroupByKey()
| 'Groups' >> beam.Map(lambda x: passthrough('Group:', x)))
# + [markdown] id="bsFACrDYgZ9-"
# `CoGroupByKey` can combine multiple `PCollections`, assuming every element is a tuple whose first item is the key to join on.
# + id="G3Mefr2d1fJa"
pipeline = beam.Pipeline()
# Create two PCollections
fruits = pipeline | 'Fruits' >> beam.Create(['apple',
'banana',
'cherry'])
print('fruits is a {}'.format(type(fruits)))
countries = pipeline | 'Countries' >> beam.Create(['australia',
'brazil',
'belgium',
'canada'])
print('countries is a {}'.format(type(countries)))
def add_key(word):
return (word[0], word)
# Create PCollections of keyed tuples
fruits_with_keys = (fruits | "fruits_with_keys" >> beam.Map(add_key)
| 'Fruit' >> beam.Map(lambda x: passthrough('Fruit:', x)))
countries_with_keys = (countries | "countries_with_keys" >> beam.Map(add_key)
| 'Country' >> beam.Map(lambda x: passthrough('Country:', x)))
fruitcountries = {"fruits": fruits_with_keys, "countries": countries_with_keys}
# Print the PCollections
print('fruits_with_keys is a {}'.format(type(fruits_with_keys)))
_ = (fruits_with_keys | 'Print0' >> beam.combiners.ToList()
| beam.Map(lambda x: print('fruits_with_keys:', x)))
print('countries_with_keys is a {}'.format(type(countries_with_keys)))
_ = (countries_with_keys | 'Print1' >> beam.combiners.ToList()
| beam.Map(lambda x: print('countries_with_keys:', x)))
print('fruitcountries: {}'.format(fruitcountries))
(fruitcountries | beam.CoGroupByKey() | beam.Map(lambda x: print('CoGrouped:', x)))
print('\nRun the pipeline')
pipeline.run().state # Try commenting out this line
# + [markdown] id="pLfHkghk3JGd"
# Try commenting out the last line, and notice that while a `PCollection` is created for the `CoGroupByKey` operation, the pipeline is not actually executed.
#
# Also try running the cell above a few times, and note how the order of the print statements changes.
# + [markdown] id="LPi2bXuh5a9l"
# ## [Combine](https://beam.apache.org/documentation/programming-guide/#combine)
# `Combine` is a transform for combining collections of elements or values. `Combine` has variants that work on entire `PCollections`, and some that combine the values for each key in `PCollections` of key/value pairs.
#
# To apply a `Combine` transform, you must provide the function that contains the logic for combining the elements or values. The combining function should be **commutative and associative**, as the function is not necessarily invoked exactly once on all values with a given key. Because the input data (including the value collection) **may be distributed across multiple workers**, the combining function might be called multiple times to perform partial combining on subsets of the value collection. The Beam SDK also provides some pre-built combine functions for common numeric combination operations such as `sum`, `min`, and `max`.
#
# Simple combine operations, such as sums, can usually be implemented as a simple function. More complex combination operations might require you to create a subclass of `CombineFn` that has an accumulation type distinct from the input/output type.
#
# Try it first with a sum.
# + id="mWC8ywHC5KQ9"
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.Create([1, 2, 3, 4, 5])
| 'Numbers' >> beam.Map(lambda x: passthrough('Number:', x))
| beam.CombineGlobally(sum)
| 'Sums' >> beam.Map(lambda x: passthrough('Sum:', x)))
# + [markdown] id="B9Jsahga-YSE"
# Now try calculating the mean using the pre-built combine function.
# + id="DS58V5cd5pQH"
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.Create([1, 2, 3, 4, 5])
| 'Numbers' >> beam.Map(lambda x: passthrough('Number:', x))
| beam.combiners.Mean.Globally()
| 'Means' >> beam.Map(lambda x: passthrough('Mean:', x)))
# + [markdown] id="m0yQwqUB-r8N"
# Now try creating a custom `CombineFn` to calculate the mean. Note the required concrete implementations.
# + id="1R9zt-xX5h-Y"
class AverageFn(beam.CombineFn):
def create_accumulator(self):
return (0.0, 0)
def add_input(self, accumulator, input_):
total, count = accumulator
total += input_
count += 1
return (total, count)
def merge_accumulators(self, accumulators):
totals, counts = zip(*accumulators)
return sum(totals), sum(counts)
def extract_output(self, accumulator):
total, count = accumulator
return total / count if count else float("NaN")
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.Create([1, 2, 3, 4, 5])
| 'Numbers' >> beam.Map(lambda x: passthrough('Number:', x))
| beam.CombineGlobally(AverageFn())
| 'Means' >> beam.Map(lambda x: passthrough('Mean:', x)))
# + [markdown] id="a4VBhAcc_pt_"
# Now try a simple `Count` using the pre-built combine function. This one does a `PerElement` count.
# + id="9N0So44g6_8M"
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.Create(['bob', 'alice', 'alice', 'bob', 'charlie', 'alice'])
| 'Persons' >> beam.Map(lambda x: passthrough('Person:', x))
| beam.combiners.Count.PerElement()
| 'Counts' >> beam.Map(lambda x: passthrough('Count:', x)))
# + [markdown] id="LI3BEiBIANHl"
# We can also create a count by using `CombinePerKey` with the Python `sum` function, using the people as keys.
# + id="Vafplz7d56CF"
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.Create(['bob', 'alice', 'alice', 'bob', 'charlie', 'alice'])
| 'Persons' >> beam.Map(lambda x: passthrough('Person:', x))
| beam.Map(lambda word: (word, 1))
| 'Keyed' >> beam.Map(lambda x: passthrough('Key:', x))
| beam.CombinePerKey(sum)
| 'Counts' >> beam.Map(lambda x: passthrough('Count:', x)))
# + [markdown] id="B6KyLL8wA4EO"
# Now try a simple `Count` using the pre-built combine function. This one does a count `Globally`.
# + id="lswZZCAe6Th5"
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.Create(['bob', 'alice', 'alice', 'bob', 'charlie', 'alice'])
| 'Persons' >> beam.Map(lambda x: passthrough('Person:', x))
| beam.combiners.Count.Globally()
| 'Counts' >> beam.Map(lambda x: passthrough('Count:', x)))
# + [markdown] id="tgoLTv8gi0WW"
# ### Now You Try
#
# This is a good point to stop reading and try writing code yourself, as a way to help you learn.
#
# **Exercise 3 — Combine Items**
#
# 1. Start with Beam pipeline you built in the previous exercise: it creates a `PCollection` containing integers 0 to 10, and labels them as odd or even.
# 2. Add another step to make the pipeline compute the square of each number.
# 3. Add a step that groups all of the odd or even numbers and computes the mean of each group (i.e., the mean of all odd numbers between 0 and 10, and the mean of all even numbers between 0 and 10). You can use the `AverageFn` that we created above.
# + id="Cjw3Sqjci0Wd"
with beam.Pipeline() as pipeline:
result = (
pipeline
| 'Create Array' >> beam.Create(range(11))
| 'Square Element' >> beam.Map(lambda x: x**2)
| 'Generate Key' >> beam.Map(lambda x: (x%2, x))
| 'Average by Key' >> beam.CombinePerKey(AverageFn())
| 'Print' >> beam.Map(lambda x: passthrough('Avg', x))
)
# 0 2 4 6 8 10
# 1 3 5 7 9
display_pipeline(pipeline)
# + id="AalDelXSi0Wh"
# + id="EPb9-f9Ri0Wj"
# + id="LQ3EL2QFi0Wl"
# + id="pgbf6O42i0Wo"
# + [markdown] id="6SzSYQ2ei0Wp"
# 
# + [markdown] id="qMYzcG46i0Wq"
# **Solution**:
# + id="KSOWJXA_jt1r"
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.Create(range(10 + 1))
| 'Numbers' >> beam.Map(lambda x: passthrough('Number:', x))
| 'Label OddEven' >> beam.Map(lambda x: ("odd" if x % 2 else "even", x))
| 'Print OddEven' >> beam.Map(lambda x: passthrough('OddEven:', x))
| 'Square' >> beam.Map(lambda x: (x[0], x[1] ** 2))
| 'Print Squares' >> beam.Map(lambda x: passthrough('Squared:', x))
| 'Calculate Means' >> beam.CombinePerKey(AverageFn())
| 'Print Means' >> beam.Map(lambda x: passthrough('Mean:', x)))
display_pipeline(pipeline)
# + [markdown] id="n-SEoKnd8cea"
# ## [Flatten](https://beam.apache.org/documentation/programming-guide/#flatten)
# `Flatten` is a transform for `PCollection` objects that store **the same data type**. `Flatten` merges multiple `PCollection` objects into a single logical `PCollection`.
#
# #### Data encoding in merged collections
# By default, the coder for the output `PCollection` is the same as the coder for the first `PCollection` in the input `PCollectionList`. However, the input `PCollection` objects can each use different coders, **as long as they all contain the same data type** in your chosen language.
#
# #### Merging windowed collections
# When using `Flatten` to merge `PCollection` objects that have a windowing strategy applied, all of the `PCollection` objects you want to merge must use a compatible windowing strategy and window sizing. For example, all the collections you're merging must all use (hypothetically) identical 5-minute fixed windows or 4-minute sliding windows starting every 30 seconds.
#
# If your pipeline attempts to use `Flatten` to merge `PCollection` objects with incompatible windows, Beam generates an `IllegalStateException` error when your pipeline is constructed.
# + id="0BVhknBM7Frz"
pipeline = beam.Pipeline()
wordsStartingWithA = pipeline | 'Words starting with A' >> beam.Create(['apple', 'ant', 'arrow'])
# Note the required parens for multi-line
wordsStartingWithB = (pipeline
| 'Words starting with B' >> beam.Create(['ball', 'book', 'bow']))
result = (wordsStartingWithA, wordsStartingWithB) | beam.Flatten()
# Print the PCollections
_ = (wordsStartingWithA | 'PrintListA' >> beam.combiners.ToList()
| 'Print A' >> beam.Map(lambda x: passthrough('wordsStartingWithA:', x)))
_ = (wordsStartingWithB | 'PrintListB' >> beam.combiners.ToList()
| 'Print B' >> beam.Map(lambda x: passthrough('wordsStartingWithB:', x)))
_ = (result | 'Result' >> beam.combiners.ToList()
| 'Print Result' >> beam.Map(lambda x: passthrough('result:', x)))
pipeline.run().state
display_pipeline(pipeline)
# + [markdown] id="etCS1bQlQNJx"
# Try running the cell above a few times, and note how the order of the print statements changes.
# + [markdown] id="rvaC0Zvj9Cse"
# ## [Partition](https://beam.apache.org/documentation/programming-guide/#partition)
# `Partition` is a transform for `PCollection` objects that store **the same data type**. `Partition` splits a single `PCollection` into a fixed number of smaller collections.
#
# `Partition` divides the elements of a `PCollection` according to a **partitioning function** that you provide. The partitioning function contains the logic that determines how to split up the elements of the input `PCollection` into each resulting partition `PCollection`. The number of partitions must be determined at graph construction time. You can, for example, pass the number of partitions as a command-line option at runtime (which will then be used to build your pipeline graph), but you cannot determine the number of partitions in mid-pipeline (based on data calculated after your pipeline graph is constructed, for instance).
#
# Let's partition a group of numbers in ranges of 100.
# + id="P6wjB7Y48olW"
def partition_fn(number, num_partitions):
partition = number // 100
return min(partition, num_partitions - 1)
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.Create([1, 110, 2, 350, 4, 5, 100, 150, 3])
| beam.Partition(partition_fn, 3))
# Print the PCollections
_ = (lines[0] | 'PrintSmallList' >> beam.combiners.ToList()
| 'Print Small' >> beam.Map(lambda x: passthrough('Small:', x)))
_ = (lines[1] | 'PrintMediumList' >> beam.combiners.ToList()
| 'Print Medium' >> beam.Map(lambda x: passthrough('Medium:', x)))
_ = (lines[2] | 'PrintLargeList' >> beam.combiners.ToList()
| 'Print Large' >> beam.Map(lambda x: passthrough('Large:', x)))
display_pipeline(pipeline)
# + [markdown] id="kBiNHmlcFBb1"
# ## [Side Inputs](https://beam.apache.org/documentation/programming-guide/#side-inputs)
# In addition to the main input `PCollection`, you can provide additional inputs to a `ParDo` transform in the form of side inputs. A side input is an additional input that your `DoFn` can access each time it processes an element in the input `PCollection`. When you specify a side input, you create a view of some other data that can be read from within the `ParDo` transform’s `DoFn` while processing each element.
#
# Side inputs are useful if your `ParDo` needs to inject additional data when processing each element in the input PCollection, but the additional data needs to be determined at runtime (and not hard-coded). Such values might be determined by the input data, or depend on a different branch of your pipeline.
#
# Let's use a side input to increment numbers by a configurable amount. First, we'll use the default for our `ParDo` transform.
# + id="YTZWcmdB9eB9"
def increment(number, inc=1):
return number + inc
with beam.Pipeline() as pipeline:
lines = (pipeline
| "Create" >> beam.Create([1, 2, 3, 4, 5])
| "Increment" >> beam.Map(increment)
| "Print" >> beam.ParDo(print))
# + [markdown] id="MKg0SV4ja4A1"
# Next, we'll pass a number to increment by.
# + id="Men6SXjGFkKH"
with beam.Pipeline() as pipeline:
lines = (pipeline
| "Create" >> beam.Create([1, 2, 3, 4, 5])
| "Increment" >> beam.Map(increment, 10) # Pass a side input of 10
| "Print" >> beam.ParDo(print))
# + [markdown] id="j2LAoh1fupfp"
# ## [Additional Outputs](https://beam.apache.org/documentation/programming-guide/#additional-outputs)
# While `ParDo` always produces a main output `PCollection` (as the return value from `apply`), you can also have your `ParDo` produce any number of additional output `PCollections`. If you choose to have multiple outputs, your `ParDo` returns all of the output `PCollections` (including the main output) bundled together.
#
# To emit elements to multiple output `PCollections`, invoke `with_outputs()` on the `ParDo`, and specify the expected tags for the outputs. `with_outputs()` returns a `DoOutputsTuple` object. Tags specified in `with_outputs` are attributes on the returned `DoOutputsTuple` object. The tags give access to the corresponding output `PCollections`.
#
# Let's try creating two `PCollections`, one each for odd and even numbers.
# + id="hlv1qnPeFt54"
def compute(number):
if number % 2 == 0:
yield number
else:
yield pvalue.TaggedOutput("odd", number + 10)
with beam.Pipeline() as pipeline:
even, odd = (pipeline
| "Create" >> beam.Create([1, 2, 3, 4, 5, 6, 7])
| "Increment" >> beam.ParDo(compute).with_outputs("odd", main="even"))
# Print the PCollections
_ = (even | 'PrintEvenList' >> beam.combiners.ToList()
| 'Print Even' >> beam.Map(lambda x: passthrough('Evens:', x)))
_ = (odd | 'PrintOddList' >> beam.combiners.ToList()
| 'Print Odd' >> beam.Map(lambda x: passthrough('Odds:', x)))
display_pipeline(pipeline)
# + [markdown] id="jTubowaGcu4J"
# Note how the `Increment` stage returns two different `PCollections` in the graph above. **Untagged elements** will always be placed in the "main" `PCollection`.
#
# Try tagging even elements and see what happens!
# + [markdown] id="ZYFrovl8IGsB"
# ## [Branching](https://beam.apache.org/documentation/programming-guide/#applying-transforms)
# A transform does not consume or otherwise alter the input collection – remember that a `PCollection` is immutable by definition. This means that you can apply multiple transforms to the same input `PCollection` to create a branching pipeline.
#
# Let's create a pipeline with 3 branches, which produces 3 different `PCollections`.
# + id="cc9hrASXGmug"
with beam.Pipeline() as branching_pipeline:
numbers = (branching_pipeline | beam.Create([1, 2, 3, 4, 5]))
mult5_results = numbers | 'Multiply by 5' >> beam.Map(lambda num: num * 5)
mult10_results = numbers | 'Multiply by 10' >> beam.Map(lambda num: num * 10)
# Print the PCollections
_ = (numbers | 'PrintNumbersList' >> beam.combiners.ToList()
| 'Print Numbers' >> beam.Map(lambda x: passthrough('Numbers:', x)))
_ = (mult5_results | 'Print5List' >> beam.combiners.ToList()
| 'Print Mult 5' >> beam.Map(lambda x: passthrough('Mult 5:', x)))
_ = (mult10_results | 'Print10List' >> beam.combiners.ToList()
| 'Print Mult 10' >> beam.Map(lambda x: passthrough('Mult 10:', x)))
display_pipeline(branching_pipeline)
# + [markdown] id="1-_FQ3S9IotY"
# ## [Composite Transforms](https://beam.apache.org/documentation/programming-guide/#composite-transforms)
# Transforms can have a nested structure, where a complex transform performs multiple simpler transforms (such as more than one `ParDo`, `Combine`, `GroupByKey`, or even other composite transforms). These transforms are called ***composite transforms***. Nesting multiple transforms inside a single composite transform can make your code more modular and easier to understand.
#
# Your composite transform's parameters and return value must match the initial input type and final return type for the entire transform, even if the transform's intermediate data changes type multiple times.
#
# To create a composite transform, create a subclass of the `PTransform` class and override the `expand` method to specify the actual processing logic. Then use this transform just as you would a built-in transform.
# + id="I8GplhSxIY8I"
# Subclass beam.PTransform to create a composite transform
class ExtractAndMultiplyNumbers(beam.PTransform):
def expand(self, pcollection):
return (pcollection
| beam.FlatMap(lambda line: line.split(","))
| beam.Map(lambda num: int(num) * 10))
with beam.Pipeline() as composite_pipeline:
number_PCollection = (composite_pipeline
| beam.Create(['1,2,3,4,5', '6,7,8,9,10'])
| ExtractAndMultiplyNumbers())
# Print the PCollection
_ = (number_PCollection | 'PrintNumbersList' >> beam.combiners.ToList()
| 'Print Numbers' >> beam.Map(lambda x: passthrough('Numbers:', x)))
display_pipeline(composite_pipeline)
# + [markdown] id="b7NFpWUMJIJy"
# ## [Filter](https://beam.apache.org/documentation/transforms/python/elementwise/filter/)
# `Filter`, given a predicate, filters out all elements that don't satisfy that predicate. `Filter` may also be used to filter based on an inequality with a given value based on the comparison ordering of the element. You can pass functions with multiple arguments to `Filter`. They are passed as additional positional arguments or keyword arguments to the function. If the `PCollection` has a single value, such as the average from another computation, passing the `PCollection` as a *singleton* accesses that value. If the `PCollection` has multiple values, pass the `PCollection` as an *iterator*. This accesses elements lazily as they are needed, so it is possible to iterate over large `PCollections` that won't fit into memory.
#
# > Note: You can pass the `PCollection` as a list with `beam.pvalue.AsList(PCollection)`, but this requires that all the elements fit into memory.
#
# > Note: You can pass the `PCollection` as a dictionary with `beam.pvalue.AsDict(PCollection)`. Each element must be a (key, value) pair. Note that all the elements of the `PCollection` must fit into memory.
#
# If the `PCollection` won't fit into memory, use `beam.pvalue.AsIter(PCollection)` instead.
#
# First, let's try filtering out even numbers using a `ParDo` transform.
# + id="BuFN36jVI4gH"
class FilterOddNumbers(beam.DoFn):
def process(self, element, *args, **kwargs):
if element % 2 == 1:
yield element
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.Create(range(1, 11))
| beam.ParDo(FilterOddNumbers())
| beam.Map(lambda x: passthrough('Odd number:', x)))
# + [markdown] id="hqlPQSCwTsFS"
# Next, let's use `Filter` to do the same thing.
# + id="8zQlSBJWJYrY"
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.Create(range(1, 11))
| beam.Filter(lambda num: num % 2 == 1)
| beam.Map(lambda x: passthrough('Odd number:', x)))
# + [markdown] id="Ic7kHP5sJphJ"
# ## [Aggregation](https://beam.apache.org/documentation/programming-guide/)
# Beam uses [windowing](https://urldefense.com/v3/__https://beam.apache.org/documentation/programming-guide/*windowing__;Iw!!PIZeeW5wscynRQ!55X7LkP-oqeRJ6UUmtPVv4vn3HLBOqlFNpBPTKq9j6A_FsUVIeCO-4RxDyJyXgQ$ ) to divide a continuously updating unbounded `PCollection` into logical windows of finite size. These logical windows are determined by some characteristic associated with a data element, such as a timestamp. [Aggregation transforms](https://urldefense.com/v3/__https://beam.apache.org/documentation/transforms/python/overview/*aggregation__;Iw!!PIZeeW5wscynRQ!55X7LkP-oqeRJ6UUmtPVv4vn3HLBOqlFNpBPTKq9j6A_FsUVIeCO-4RxGg6bt2E$ ) (such as GroupByKey and Combine) work on a per-window basis — as the data set is generated, they process each `PCollection` as a succession of these finite windows.
#
# A related concept, called [triggers](https://urldefense.com/v3/__https://beam.apache.org/documentation/programming-guide/*triggers__;Iw!!PIZeeW5wscynRQ!55X7LkP-oqeRJ6UUmtPVv4vn3HLBOqlFNpBPTKq9j6A_FsUVIeCO-4RxSPrnvUA$ ), determines when to emit the results of aggregation as unbounded data arrives. You can use triggers to refine the windowing strategy for your `PCollection`. Triggers allow you to deal with late-arriving data, or to provide early results.
#
# First, let's count the elements in a `PCollection`.
# + id="O3W-h0AzJg3f"
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.Create(range(1, 11))
| beam.combiners.Count.Globally() # Count
| beam.Map(lambda x: passthrough('Count =', x)))
# + [markdown] id="TMHW9Je3VCUP"
# Next, let's sum the elements with a `CombineGlobally` aggregator and Python's sum.
# + id="GZVWg2AvKxUz"
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.Create(range(1, 11))
| beam.CombineGlobally(sum) # CombineGlobally sum
| beam.Map(lambda x: passthrough('Sum =', x)))
# + [markdown] id="tRXBNpUTVkRw"
# Next, let's calculate the mean.
# + id="PpkdeF6YK2gc"
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.Create(range(1, 11))
| beam.combiners.Mean.Globally() # Mean
| beam.Map(lambda x: passthrough('Mean =', x)))
# + [markdown] id="Zkw5rjkEVv7M"
# Next, let's return the smallest element in the `PCollection`.
# + id="EYJCGAgdK227"
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.Create(range(1, 11))
| beam.combiners.Top.Smallest(1) # Top Smallest
| beam.Map(lambda x: passthrough('Smallest =', x)))
# + [markdown] id="XnQpGZTYV5t-"
# Next, let's return the largest element in the `PCollection`.
# + id="aXySuL7QK4sb"
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.Create(range(1, 11))
| beam.combiners.Top.Largest(1) # Top Largest
| beam.Map(lambda x: passthrough('Largest =', x)))
# + [markdown] id="GMKY3LCWLGXu"
# # [Pipeline I/O](https://beam.apache.org/documentation/programming-guide/#pipeline-io)
# When you create a pipeline, you often need to read data from some external source, such as a file or a database. Likewise, you may want your pipeline to output its result data to an external storage system. Beam provides read and write transforms for a [number of common data storage types](https://beam.apache.org/documentation/io/built-in/). If you want your pipeline to read from or write to a data storage format that isn’t supported by the built-in transforms, you can [implement your own read and write transforms](https://beam.apache.org/documentation/io/developing-io-overview/)
# ### Download example data
# Download the sample dataset for use with the cells below.
# + id="BywX6OUEhAqn"
import os, tempfile, urllib
DATA_PATH = 'https://raw.githubusercontent.com/tensorflow/tfx/master/tfx/examples/chicago_taxi_pipeline/data/simple/data.csv'
_data_root = tempfile.mkdtemp(prefix='tfx-data')
_data_filepath = os.path.join(_data_root, "data.csv")
urllib.request.urlretrieve(DATA_PATH, _data_filepath)
# + [markdown] id="4pwOXlPQYiJn"
# Let's take a quick look at the data.
# + id="Hqn4wST2Bex5"
# !head {_data_filepath}
# + [markdown] id="R9BQpD-JYlte"
# Let's filter out only the lines that we're interested in.
# + id="hz4oEI90mB-m"
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.io.ReadFromText(_data_filepath)
| beam.Filter(lambda line: line.startswith("13,16.45,11,12,3"))
| beam.Map(lambda x: passthrough('Match:', x)))
# + [markdown] id="8siWql-89oq1"
# ### Putting Everything Together
#
# Use several of the concepts, classes, and methods discussed above in a concrete example.
#
# **Exercise 4 — Reading, Filtering, Parsing, Grouping and Averaging**
#
# Write a Beam pipeline that reads the dataset, computes the mean fare for each company, and takes the top 5 companies with the largest mean fares.
#
# *Hints*:
# * Use the code above to read the dataset.
# * Add a `Map` step to split each row on the commas.
# * Filter out the header row and any rows without values for company.
# * Calculate the mean fare for each company
# * (Optional) Print the mean fares for each company
# * Take the top 5 companies by mean fare
# * Print the list of the top 5 companies
# + id="I_jeg125l6sR"
with beam.Pipeline() as pipeline:
average_per_company = (
pipeline
| beam.io.ReadFromText(_data_filepath)
| beam.Map(lambda x: x.split(','))
| beam.Filter(lambda x: x[0] != 'pickup_community_area' and x[-4])
| beam.Map(lambda x: (x[-4], float(x[1])))
| beam.CombinePerKey(AverageFn())
)
print_average_fare = (
average_per_company
| beam.Map(lambda x: passthrough('avg({}) ='.format(x[0]), x[1]))
)
print_average_top5 = (
average_per_company
| beam.Map(lambda x: (x[1], x[0]))
| beam.combiners.Top.Largest(5)
| beam.Map(lambda x: passthrough('Top 5 Companies =', x))
)
display_pipeline(pipeline)
# + id="3q6OTO5Xl6sU"
# + id="U5sVJaojl6sW"
# + id="Y7LfH_rHl6sZ"
# + id="lNSVxnmnl6sa"
# + [markdown] id="u8pjA5uFl6sc"
# 
# + [markdown] id="I2mfIjMbl6sc"
# **Solution**:
# + id="QGLY9QRJLB8y"
with beam.Pipeline() as pipeline:
lines = (pipeline
| beam.io.ReadFromText(_data_filepath)
| beam.Map(lambda line: line.split(",")) # CSV
| beam.Filter(lambda cols: len(cols[-4]) > 0 and cols[0] != 'pickup_community_area')
| beam.Map(lambda cols: (cols[-4], float(cols[1]))) # (Company, Mean Fare) for PerKey
| beam.combiners.Mean.PerKey() # Mean fare by company
| beam.Map(lambda cols: (cols[1], cols[0])) # Switch to (Mean Fare, Company) for Top
| beam.Map(lambda x: passthrough('(Mean Fare, Company):', x))
| beam.combiners.Top.Largest(5) # Top 5 mean fares
| beam.Map(lambda x: passthrough('Top 5:', x)))
# + [markdown] id="SZlIAvT4o0tW"
# ## Beam I/O and TFX
#
# Since TensorFlow and TFX often use the [TFRecord](https://urldefense.com/v3/__https://www.tensorflow.org/tutorials/load_data/tfrecord__;!!PIZeeW5wscynRQ!55X7LkP-oqeRJ6UUmtPVv4vn3HLBOqlFNpBPTKq9j6A_FsUVIeCO-4RxBE9fjf0$ ) format, let's use Beam I/O with TFRecord files.
#
# First, let's create a test file containing 10 records of "Record X" strings in UTF-8.
# + id="IGEzjUGaRHCW"
# #!pip install -q -U "tensorflow>=2.1,<=2.2"
import tensorflow as tf
with tf.io.TFRecordWriter("test.tfrecord") as tfrecord_file:
for index in range(10):
tfrecord_file.write("Record {}".format(index).encode("utf-8"))
dataset = tf.data.TFRecordDataset('test.tfrecord')
for record in dataset:
print(record.numpy().decode('utf-8'))
# + [markdown] id="FzAt26WBqgKE"
# Now let's read the test file and process each of the records. We'll write the results to a second `TFRecord` file, one record for each record processed.
# + id="hgRAt4nRR-qf"
with beam.Pipeline() as rw_pipeline:
lines = (rw_pipeline
| 'Read test file' >> beam.io.ReadFromTFRecord("test.tfrecord")
| 'Process each record' >> beam.Map(lambda line: line + b' processed')
| 'Write the results to a new file' >> beam.io.WriteToTFRecord("test_processed.tfrecord")
| 'Print the filename' >> beam.ParDo(print))
display_pipeline(rw_pipeline)
# + [markdown] id="X_j06O0ktIdy"
# Now let's read the results file and print each record.
# + id="sT_bo1VxTbTb"
with beam.Pipeline() as utf_pipeline:
lines = (utf_pipeline
| "Read the results file" >> beam.io.ReadFromTFRecord("test_processed.tfrecord-00000-of-00001")
| "Decode UTF-8" >> beam.Map(lambda line: line.decode('utf-8'))
| "Print each record" >> beam.ParDo(print))
display_pipeline(utf_pipeline)
# + [markdown] id="KCAysNHLQK80"
# Note that there are many [other built-in I/O transforms](https://urldefense.com/v3/__https://beam.apache.org/documentation/io/built-in/__;!!PIZeeW5wscynRQ!55X7LkP-oqeRJ6UUmtPVv4vn3HLBOqlFNpBPTKq9j6A_FsUVIeCO-4RxE3x8M-c$ ).
# + [markdown] id="KZDYehqpPpXY"
# # [Windowing](https://beam.apache.org/documentation/programming-guide/#windowing)
# As discussed above, windowing subdivides a `PCollection` according to the timestamps, or some other ordering or grouping, of its individual elements.
#
# Some Beam transforms, such as `GroupByKey` and `Combine`, group multiple elements by a common key. Ordinarily, that grouping operation groups all of the elements that have the same key within the entire data set. With an unbounded data set, it is impossible to collect all of the elements, since new elements are constantly being added and may be infinitely many (e.g. streaming data). If you are working with unbounded `PCollections`, windowing is especially useful.
#
# In the Beam model, any `PCollection` (including unbounded `PCollections`) can be subdivided into logical windows. Each element in a `PCollection` is assigned to one or more windows according to the `PCollection`'s windowing function, and each individual window contains a finite number of elements. Grouping transforms then consider each `PCollection`'s elements on a per-window basis. `GroupByKey`, for example, implicitly groups the elements of a `PCollection` by key and window.
#
# Additional information on Beam Windowing is available in the [Beam Programming Guide](https://urldefense.com/v3/__https://beam.apache.org/documentation/programming-guide/*windowing__;Iw!!PIZeeW5wscynRQ!55X7LkP-oqeRJ6UUmtPVv4vn3HLBOqlFNpBPTKq9j6A_FsUVIeCO-4RxDyJyXgQ$ ).
# + [markdown] id="tLMh3chYC1sE"
# Let's begin by creating a pipeline to:
#
# * Read our CSV file above
# * Create fixed windows for each week
# * Calculate the mean fare by company for each week
# * Group the companies by week
# * Print out 2016 only
# + id="Qd3a4hwlVqfF"
import datetime
DAYS = 24 * 60 * 60
class BuildRecordFn(beam.DoFn):
def process(self, element, window=beam.DoFn.WindowParam):
window_end = str(window.end.to_utc_datetime())
return [(window_end, element)]
class AssignTimestamps(beam.DoFn):
def process(self, element):
ts = int(element[5])
yield beam.window.TimestampedValue(element, ts)
with beam.Pipeline() as window_pipeline:
lines = (window_pipeline
| 'Read text file' >> beam.io.ReadFromText(_data_filepath)
| 'Parse CSV' >> beam.Map(lambda line: line.split(","))
| 'Filter out' >> beam.Filter(lambda cols: len(cols[-4]) > 0 and cols[0] != 'pickup_community_area')
| 'Assign timestamps' >> beam.ParDo(AssignTimestamps())
| 'Create week windows' >> beam.WindowInto(beam.window.FixedWindows(7*DAYS))
| '(Company, Mean Fare) for PerKey' >> beam.Map(lambda cols: (cols[-4], float(cols[1])))
| 'Mean fare by company' >> beam.combiners.Mean.PerKey()
| 'Add window end' >> beam.ParDo(BuildRecordFn())
| 'Group windows' >> beam.GroupByKey()
| 'Only include 2016' >> beam.Filter(lambda win: win[0] > '2016' and win[0] < '2017')
# | 'Sort by Key' >> beam.SortByKey()
| 'Print' >> beam.Map(lambda x: passthrough('(Window end, [Mean fares by company]):', x)))
display_pipeline(window_pipeline)
# -
| tfx_apache_beam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Loading Data Set
import pandas as pd
# # Formatting Gene Expressions
# +
# loading gene counts
gene_fpkm = pd.read_csv('data/gene_count.txt', sep='\t', index_col='GENE_ID')
# removing data not collected at the first trail
for col in gene_fpkm.columns:
if '_1_' not in col:
del gene_fpkm[col]
# transpose matrix, delete patients and gene with all nan, and replace remainder missing by zero
gene_fpkm = gene_fpkm.T.dropna(how='all', axis=0).dropna(how='any', axis=1)
# replace id column name
gene_fpkm.index.name = 'ID'
# normalize index value transforming mmrf ids to integers
gene_fpkm.index = [''.join(col.split('_')[:2]) for col in gene_fpkm.index]
# selected class
# gene_details = pd.read_csv('data/gene_details.tsv', sep='\t')
# gene_selected_class = pd.read_csv('data/gene_selected_class.tsv', sep='\t')
# gene_selected_class = gene_details.merge(gene_selected_class, on='gene_biotype').set_index('ensembl_gene_id')
# gene_selected_class = [gen for gen in gene_selected_class.index if gen in gene_fpkm.columns]
# gene_fpkm = gene_fpkm[gene_selected_class]
# removing genes with zero sum
gene_fpkm = gene_fpkm[list(gene_fpkm.sum()[gene_fpkm.sum() > 0].index)]
# removing duplicated index
gene_fpkm = gene_fpkm.loc[~gene_fpkm.index.duplicated(keep='first')]
# removing duplicated columns
gene_fpkm = gene_fpkm.loc[:,~gene_fpkm.columns.duplicated()]
gene_fpkm.index.name = 'ID'
gene_fpkm.to_csv('data/gene_count.tsv', sep='\t')
print(gene_fpkm.shape)
gene_fpkm.iloc[:8,:8]
| 00_02_clean_gen_exp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "NGA Tennis Financial Calculations"
# > "Quick calculation shows that closing the tennis courts will not help in the financial deficit, if there is one."
#
# - toc: true
# - branch: master
# - badges: true
# - comments: true
# - categories: [fastpages, jupyter, nga, tennis, financial]
# - hide: false
# - search_exclude: true
# - metadata_key1: metadata_value1
# - metadata_key2: metadata_value2
# ## Overview
# NGA released a follow-up circular regarding the NGA Tennis Court Conversion topic with financial information. However, NGA has neglected to show the expected "financial deficit" that "is clearly forthcoming." Therefore, this is our attempt to calculate that financial deficit.
#
# According to the limited information given, it does not show that the conversion of the tennis court will help in the financial status significantly. By cutting off the expenses of the tennis court, there will be an increase of around 0.2% in cash balance per annum. Furthermore, current estimates show that NGA cash balance will still stay positive.
# ## Methodology
# 1. Calculate the ratio of expenses of same period pre quarantine and during quarantine.
# 2. Calculate the estimated yearly expense of during quarantine using the ratio calculated in step 1
# 3. Calculate the estimated yearly total collection of during quarantine period
# 4. Calculate the estimated yearly loss by maintaining tennis court
# 5. Calculate the estimated yearly cash balance during quarantine by finding the difference between step 2 and step 3
# 6. Calculate the percentage component of step 4 to step 5
# ## Data Prep: NGA Financial Data
# In the circular dated 18 July 2020, NGA shared unaudited revenue summary. We will be using these information:
# 1. Membership Dues (same for 2019 and 2020)
# 2. Tennis Collection (past 5 years)
# 3. Tennis Expenses (past 5 years)
# 4. Same period expenses (Jan-June 2019 and 2020)
# 5. Same period collection (Jan-June 2019 and 2020)
#
# Also in the same circular, NGA has provided additional insights that are useful:
# 1. The present Homeowners Association Dues covers ONLY about 25% on an average of our total yearly expenses.
# 2. NGA has already collected an estimate of 90% of this Year 2020 revenues as of this writing.
#
# This section will prepare the data to be used in the calculation based on the statements above.
# +
# Data Prep: Financial Information
mem_dues = {
'2019': 8152483.9
}
mem_dues['2020'] = mem_dues['2019']
tennis = {
'expense': 2297120,
'collection': 1930940.25
}
same_period_expense = {
'2019': 19037930.42,
'2020': 12928366.14
}
same_period_collection = {
'2019': 33782575.40,
'2020': 22844454.50
}
# Other constants
mem_dues_perc = 0.25
collection_rate = 0.9
tennis_years = 5
# -
# ## Calculations
# This section is to calculate the values listed in 1.2 Methodology
# ### Estimated Yearly Expenses: Pre Quarantine vs During Quarantine
# +
# Calculate estimated yearly expenses by using mem_dues_perc for 2019
est_year_expense = {}
est_year_expense['2019'] = mem_dues['2019']/mem_dues_perc
# Calculate estimated yearly expenses for quarantine
ratio_same_period_expense = same_period_expense['2020']/same_period_expense['2019']
est_year_expense['2020'] = est_year_expense['2019']*ratio_same_period_expense
# -
# ### Estimated Collection: Quarantine
# Use collection_rate to project total collection of 2020
est_total_collection = {}
est_total_collection['2020'] = same_period_collection['2020'] / collection_rate
# ### Tennis Calculations
# Estimate yearly loss by averaging the delta of collection and expense over 5 years
tennis_ave_yearly_delta = (tennis['collection'] - tennis['expense'])/tennis_years
# ### Estimated Yearly Cash Balance: During Quarantine
# Project yearly cash balance for quarantine period
est_yearly_cash_balance = est_total_collection['2020'] - est_year_expense['2020']
# ### Estimate Percentage of Tennis to Total Cash
est_tennis_percentage = tennis_ave_yearly_delta / est_yearly_cash_balance
# ## Results
# String formatting
result_values = [
'{:,.2f}'.format(abs(tennis_ave_yearly_delta)),
'{:,.2f}'.format(abs(est_year_expense['2020'])),
'{:,.2f}'.format(est_total_collection['2020']),
'{:,.2f}'.format(abs(est_tennis_percentage)*100)
]
#collapse
print(
'''
By removing tennis courts, NGA will save around {} php per year.
The estimated yearly expenses amount to {} php per year during quarantine.
The estimated yearly collection amount to {} php.
Therefore, estimated yearly collection is projected to be higher than estimated yearly expenses.
If we remove the tennis court, then the cash balance would increase by about {}% per annum
'''.format(
result_values[0],
result_values[1],
result_values[2],
result_values[3]
)
)
| _notebooks/2020-07-18-NGA-Tennis-Calculation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import time
import sys
from selenium import webdriver
from selenium.webdriver.common.by import By
from selenium.webdriver.common.keys import Keys
from selenium.webdriver import ActionChains
from selenium.common.exceptions import NoSuchElementException, TimeoutException
from selenium.webdriver.support import expected_conditions as EC
from selenium.webdriver.support.wait import WebDriverWait
import pandas as pd
import csv
import random
import pandas as pd
# +
driver = webdriver.Chrome('/Users/leijunjie/JUNJIE/myScaper/Seleuim/chromedriver')
driver.get("http://www.baidu.com")
input = driver.find_element_by_id('kw')
input.send_keys("失信")
time.sleep(1)
input.send_keys("被执行人")
time.sleep(1)
input.send_keys(Keys.ENTER)
WebDriverWait(driver, 80).until(EC.visibility_of_element_located((By.CSS_SELECTOR, 'div.op_trust_mainBox')))
def fetch_page_data():
Name_list = []
Ids = []
val1 = []
val2 = []
val3 = []
val4 = []
val5 = []
val6 = []
val7 = []
val8 = []
items = driver.find_elements_by_css_selector('li.op_trust_item')
for item in items:
try:
Name = item.find_element_by_css_selector(
'span.op_trust_name').text
Id_number = item.find_element_by_css_selector(
'span.op_trust_fl').text
# Click & Open baidu Bulletin Board
ActionChains(driver).click(item).perform()
values = item.find_elements_by_css_selector(
'tbody td.op_trust_tdRight')
except StaleElementReferenceException:
# if element attachment error then we rest for 1 sec and repeat this process;
try:
print("出现异常,等待重试...")
time.sleep(1)
ActionChains(self.driver).click(item).perform()
values = item.find_elements_by_css_selector(
'tbody td.op_trust_tdRight')
except StaleElementReferenceException:
print("重试失败,跳过该公告...")
continue
if len(values) == 7:
Name_list.append(Name)
Ids.append(Id_number)
val1.append(values[0].text)
val2.append(values[1].text)
val3.append(values[2].text)
val4.append(values[3].text)
val5.append(values[4].text)
val6.append(values[5].text)
val7.append(values[6].text)
val8.append('individual')
else:
Name_list.append(Name)
Ids.append(Id_number)
val1.append(values[1].text)
val2.append(values[2].text)
val3.append(values[3].text)
val4.append(values[4].text)
val5.append(values[5].text)
val6.append(values[6].text)
val7.append(values[7].text)
val8.append(values[0].text)
df_temp = pd.DataFrame(
{
"Name": Name_list,
"ID_Num": Id_number,
"执行法院": val1,
"省份":val2,
"案号":val3 ,
"生效法律文书":val4,
"被执行人情况":val5,
"具体行为":val6,
"发布时间":val7,
"if_company":val8
})
return df_temp
df = fetch_page_data()
pagenum = 1
while (pagenum < 3):
print("Scraping Page %d..." % pagenum)
next_btn = driver.find_element_by_css_selector(
'div.op_trust_page span.op_trust_page_next')
ActionChains(driver).click(next_btn).perform()
time.sleep(random.randint(10,12))
df_2 = fetch_page_data()
df = pd.merge(df, df_2, how='outer')
time.sleep(random.randint(5,9))
pagenum +=1
print('finished!')
# -
df.info()
df
| dataCollection/SX_Spider_SE/SE_SpiderDemo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Spotify Data Collection
# ## <NAME>
# ---
# imports
import numpy as np
import pandas as pd
from dotenv import load_dotenv
from os import getenv
from spotipy import Spotify, SpotifyException
from spotipy.oauth2 import SpotifyClientCredentials
from pycountry import countries
# +
# Instantiate authorized Spotify API object
load_dotenv()
client_id = getenv('CLIENT_ID')
client_secret = getenv('CLIENT_SECRET')
auth_manager = SpotifyClientCredentials(
client_id=client_id,
client_secret=client_secret
)
spotify = Spotify(auth_manager=auth_manager)
# -
# Function for gathering Spotify category playlists by country
def get_playlists():
"""
Creates a list of Spotify markets,
and queries the Spotify API for "decades" playlists
returns:
-------
playlists: dict
Dictionary of decades playlists from each supported country
"""
# Collect category data for all Spotify markets
markets = spotify.available_markets()['markets']
print(f"Number of Spotify markets: {len(markets)}")
# Get decades playlists for each country
categories = [spotify.categories(market) for market in markets]
playlists = {}
for market, category in zip(markets, categories):
items = category['categories']['items']
playlist_ids = [item['id'] for item in items]
if 'decades' in playlist_ids:
playlists[market] = spotify.category_playlists('decades', country=market)
print(f"Number of countries with decades playlists: {len(playlists)}")
return playlists
playlists = get_playlists()
# +
# Create country index
country_index = {}
for market in playlists.keys():
country = countries.get(alpha_2=market)
if country is not None:
country_index[market] = country.name
country_index
# -
playlists['AE']
| notebooks/data_collection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Exercise for Homework
#
# Since we had limited time during the lab, here are some exercises that can help you with homework 1. The answers are hidden in the cell just below the empty code cells for you.
#
# ## Part A: Basic Operations
#
# Calculate the average (mean) of `12, 13, 14, 15, 16, 199` and save it in a variable `x`.
# +
# your code here. Click the three dots below to see answer
# + jupyter={"source_hidden": true}
#You can store the numbers in a vector using the c() command
num <- c(12, 13, 14, 15, 16, 199)
#Then, you can compute the mean by taking the sum of x/6
x <- sum(num)/6
#Or use directly the function mean()
x <- mean(num)
# -
# Load the data from internal demo dataset `iris`.
# + jupyter={"source_hidden": true}
data("iris")
# -
# Show first 10 rows of the data. Use the head() function
# + jupyter={"outputs_hidden": true, "source_hidden": true}
head(iris, n=10)
# -
# Accessing values in a data frame: Find the value on row 15, column 4
# + jupyter={"outputs_hidden": true, "source_hidden": true}
iris[15,4]
# -
# The `[ ]` (e.g. `a[]`) is somehow similar to a function call `()`, except that it is an internal function specified in the object `a`.
#
# If `a` is a dataframe, `a[<rows>,<columns>]` will get you the specified part of the dataframe.
#
# The first number `<rows>` you type in, represents a row (or rows) in the data frame.
#
# The second number `<columns>` you type in, after the comma, represents a column (or columns) in the data frame.
#
# However, you do not need to specify both number. If you leave it blank, it will by default get you all the columns or rows:
# + jupyter={"source_hidden": true}
iris[1,] # gets you all the columns but only gives you the first row
# -
# If you know the name of the desired variable/column, you can directly use the name of the column, rather than the number.
# + jupyter={"outputs_hidden": true, "source_hidden": true}
iris[,"Sepal.Length"] # note that this will give you a list, not a data.frame
# + jupyter={"outputs_hidden": true, "source_hidden": true}
iris$Sepal.Length # alternatively
# -
# ## Part 2: Loading & Processing Data
#
# Import `"coffee.csv`, and store it as `CoffeeData`.
#
# Note that the delimiter of this file is not comma, it is semicolon (`;`). We need to specify the delimiter in read.csv() function with the `sep` parameter.
#
# Fill out the following code, get help from the contexual help in JupyterLab.
# +
# CoffeeData <- read.csv( )
# + jupyter={"source_hidden": true}
CoffeeData <- read.csv("coffee.csv",sep = ";")
# -
# Summarize the variables in coffee data, check for potential missing values
# + jupyter={"outputs_hidden": true, "source_hidden": true}
summary(CoffeeData)
# -
# Or try using `is.na`. It works for both lists and dataframe.
# + jupyter={"outputs_hidden": true, "source_hidden": true}
# You can also check rows with missing values explicitly using is.na(). You can pass any R object to it.
is.na(CoffeeData)
# -
# To find out the index of missing data identified by `is.na`, use `which` function. However, `which` works best with lists, it does not work well with dataframe.
# + jupyter={"source_hidden": true}
which(is.na(CoffeeData$COGS))
# -
# ### Arranging Data
#
# What is the `ProductID` of the most profitable coffee?
#
# First, we sort the data based on Profit, in decreasing order:
# + jupyter={"source_hidden": true}
CoffeeData_sorted <- CoffeeData[order(CoffeeData$Profit,decreasing = TRUE),]
# + jupyter={"outputs_hidden": true, "source_hidden": true}
#Option 1: We can look at the head of the sorted dataset, and specify n =1
head(CoffeeData_sorted,n=1)
#Option 2: look at the dataframe, row 1
CoffeeData_sorted[1,]
#Option 3: from the original dataset,
# take the Profit column CoffeeData$Profit and ask for the row
# that is exactly equal (==) to the maximum of the values in the Profit column
CoffeeData[CoffeeData$Profit==max(CoffeeData$Profit),]
# -
# ### Subsetting Data
#
# Find the subset of data with `Sales` larger than `200`
# + jupyter={"source_hidden": true}
CoffeeData_subset <- subset(CoffeeData, Sales > 200)
# -
# ## Part 3 Visualization
#
# Plot the distribution of `COGS`, using the original coffee dataset:
# + jupyter={"outputs_hidden": true, "source_hidden": true}
# built-in histogram
hist(CoffeeData$COGS,main = "COGS", col = "Blue", xlab = "Cost of Goods", xlim = c(0, 400))
# + jupyter={"outputs_hidden": true, "source_hidden": true}
#ggplot, example 1
# install.packages("ggplot2") # Un-comment this line to install ggplot if you havn't already
library("ggplot2")
ggplot(data = CoffeeData, aes(x = COGS)) +
geom_histogram(color = "darkblue" , fill = "blue", alpha = 0.5) +
xlim (0, 400)
# + jupyter={"outputs_hidden": true, "source_hidden": true}
#ggplot, example 2: get a different gradient of color depending on the count
ggplot(data = CoffeeData, aes(x = COGS, fill = ..count..)) +
geom_histogram(alpha = 0.5) +
xlim (0, 400) +
scale_fill_gradient(low="blue", high="red")
# -
# Now let's try boxplot. Plot the distribution of profit across different products.
# + jupyter={"outputs_hidden": true, "source_hidden": true}
#basic boxplot
boxplot(Profit~ProductId,CoffeeData, ylim = c(-700, 700))
# + jupyter={"outputs_hidden": true, "source_hidden": true}
# example using ggplot
ggplot(data = CoffeeData, aes(y=Profit, group =ProductId, color = ProductId)) +
geom_boxplot(notch = TRUE, outlier.colour="red", outlier.shape=8, outlier.size=2)
# Note the outlier arguments in `geom_boxplot`. You can know more from the contexual help.
# -
# Try scatterplot. Plot the relationship between marketing and sales:
# + jupyter={"outputs_hidden": true, "source_hidden": true}
#example using plot
plot(CoffeeData$Marketing, CoffeeData$Sales, xlim=c(0,200), ylim = c(0, 1000), xlab = "Marketing",
ylab = "Sales")
# + jupyter={"outputs_hidden": true, "source_hidden": true}
#example using qplot
qplot(data = CoffeeData, x = CoffeeData$Marketing, y = CoffeeData$Sales, col = CoffeeData$ProductId)
# -
| Lab 1 - Intro to R/Part 3 Exercise For Homework.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + raw_mimetype="text/restructuredtext" active=""
# .. _networks::
#
# |
# |
#
# Download This Notebook: :download:`Networks.ipynb`
#
# -
# # Networks
# ## Introduction
# This tutorial gives an overview of the microwave network analysis
# features of **skrf**. For this tutorial, and the rest of the scikit-rf documentation, it is assumed that **skrf** has been imported as `rf`. Whether or not you follow this convention in your own code is up to you.
import skrf as rf
from pylab import *
# If this produces an import error, please see [Installation ](Installation.ipynb).
# ## Creating Networks
#
# **skrf** provides an object for a N-port microwave [Network](../api/network.rst). A [Network](../api/network.rst) can be created in a number of ways:
# - from a Touchstone file
# - from S-parameters
# - from Z-parameters
# - from other RF parameters (Y, ABCD, T, etc.)
#
# Some examples for each situation is given below.
# ### Creating Network from Touchstone file
# [Touchstone file](https://en.wikipedia.org/wiki/Touchstone_file) (`.sNp` files, with `N` being the number of ports) is a _de facto_ standard to export N-port network parameter data and noise data of linear active devices, passive filters, passive devices, or interconnect networks. Creating a Network from a Touchstone file is simple:
# +
from skrf import Network, Frequency
ring_slot = Network('data/ring slot.s2p')
# -
# Note that some softwares, such as ANSYS HFSS, add additional information to the Touchstone standard, such as comments, simulation parameters, Port Impedance or Gamma (wavenumber). These data are also imported if detected.
#
# A short description of the network will be printed out if entered onto the command line
#
ring_slot
# ### Creating Network from s-parameters
# Networks can also be created by directly passing values for the `frequency`, `s`-parameters (and optionally the port impedance `z0`).
#
# The scattering matrix of a N-port Network is expected to be a Numpy array of shape `(nb_f, N, N)`, where `nb_f` is the number of frequency points and `N` the number of ports of the network.
# dummy 2-port network from Frequency and s-parameters
freq = Frequency(1, 10, 101, 'ghz')
s = rand(101, 2, 2) + 1j*rand(101, 2, 2) # random complex numbers
# if not passed, will assume z0=50. name is optional but it's a good practice.
ntwk = Network(frequency=freq, s=s, name='random values 2-port')
ntwk
ntwk.plot_s_db()
# Often, s-parameters are stored in separate arrays. In such case, one needs to forge the s-matrix:
# +
# let's assume we have separate arrays for the frequency and s-parameters
f = np.array([1, 2, 3, 4]) # in GHz
S11 = np.random.rand(4)
S12 = np.random.rand(4)
S21 = np.random.rand(4)
S22 = np.random.rand(4)
# Before creating the scikit-rf Network object, one must forge the Frequency and S-matrix:
freq2 = rf.Frequency.from_f(f, unit='GHz')
# forging S-matrix as shape (nb_f, 2, 2)
# there is probably smarter way, but less explicit for the purpose of this example:
s = np.zeros((len(f), 2, 2))
s[:,0,0] = S11
s[:,0,1] = S12
s[:,1,0] = S21
s[:,1,1] = S22
# constructing Network object
ntw = rf.Network(frequency=freq2, s=s)
print(ntw)
# -
# If necessary, the characteristic impedance can be passed as a scalar (same for all frequencies), as a list or an array:
ntw2 = rf.Network(frequency=freq, s=s, z0=25, name='same z0 for all ports')
print(ntw2)
ntw3 = rf.Network(frequency=freq, s=s, z0=[20, 30], name='different z0 for each port')
print(ntw3)
ntw4 = rf.Network(frequency=freq, s=s, z0=rand(101,2), name='different z0 for each frequencies and ports')
print(ntw4)
# ### from z-parameters
# As networks are also defined from their Z-parameters, there is `from_z()` method of the Network:
# +
# 1-port network example
z = 10j
Z = np.full((len(freq), 1, 1), z) # replicate z for all frequencies
ntw = rf.Network()
ntw = ntw.from_z(Z)
ntw.frequency = freq
print(ntw)
# -
# ### from other network parameters (Z, Y, ABCD, T)
# It is also possible to generate Networks from other kind of RF parameters, using the conversion functions: `z2s`, `y2s`, `a2s`, `t2s`, `h2s`. For example, the [ABCD parameters](https://en.wikipedia.org/wiki/Two-port_network#ABCD-parameters) of a serie-impedance is:
# $$
# \left[
# \begin{array}{cc}
# 1 & Z \\
# 0 & 1
# \end{array}
# \right]
# $$
# +
z = 20
abcd = array([[1, z],
[0, 1]])
s = rf.a2s(tile(abcd, (len(freq),1,1)))
ntw = Network(frequency=freq, s=s)
print(ntw)
# -
# ## Basic Properties
#
#
# The basic attributes of a microwave [Network](../api/network.rst) are provided by the
# following properties :
#
# * `Network.s` : Scattering Parameter matrix.
# * `Network.z0` : Port Characteristic Impedance matrix.
# * `Network.frequency` : Frequency Object.
# The [Network](../api/network.rst) object has numerous other properties and methods. If you are using IPython, then these properties and methods can be 'tabbed' out on the command line.
#
# In [1]: ring_slot.s<TAB>
# ring_slot.line.s ring_slot.s_arcl ring_slot.s_im
# ring_slot.line.s11 ring_slot.s_arcl_unwrap ring_slot.s_mag
# ...
#
# All of the network parameters are represented internally as complex `numpy.ndarray`. The s-parameters are of shape (nfreq, nport, nport)
shape(ring_slot.s)
# ## Slicing
# You can slice the `Network.s` attribute any way you want.
ring_slot.s[:11,1,0] # get first 10 values of S21
# Slicing by frequency can also be done directly on Network objects like so
ring_slot[0:10] # Network for the first 10 frequency points
# or with a human friendly string,
ring_slot['80-90ghz']
# Notice that slicing directly on a Network **returns a Network**. So, a nice way to express slicing in both dimensions is
ring_slot.s11['80-90ghz']
# ## Plotting
# Amongst other things, the methods of the [Network](../api/network.rst) class provide convenient ways to plot components of the network parameters,
#
# * `Network.plot_s_db` : plot magnitude of s-parameters in log scale
# * `Network.plot_s_deg` : plot phase of s-parameters in degrees
# * `Network.plot_s_smith` : plot complex s-parameters on Smith Chart
# * ...
#
# If you would like to use skrf's plot styling,
# %matplotlib inline
rf.stylely()
#
# To plot all four s-parameters of the `ring_slot` on the Smith Chart.
ring_slot.plot_s_smith()
# Combining this with the slicing features,
# +
from matplotlib import pyplot as plt
plt.title('Ring Slot $S_{21}$')
ring_slot.s11.plot_s_db(label='Full Band Response')
ring_slot.s11['82-90ghz'].plot_s_db(lw=3,label='Band of Interest')
# -
# For more detailed information about plotting see [Plotting](Plotting.ipynb).
# ## Arithmetic Operations
#
# Element-wise mathematical operations on the scattering parameter matrices are accessible through overloaded operators. To illustrate their usage, load a couple Networks stored in the `data` module.
# +
from skrf.data import wr2p2_short as short
from skrf.data import wr2p2_delayshort as delayshort
short - delayshort
short + delayshort
short * delayshort
short / delayshort
# -
# All of these operations return [Network](../api/network.rst) types. For example, to plot the complex difference between `short` and `delay_short`,
difference = (short - delayshort)
difference.plot_s_mag(label='Mag of difference')
# Another common application is calculating the phase difference using the division operator,
(delayshort/short).plot_s_deg(label='Detrended Phase')
# Linear operators can also be used with scalars or an `numpy.ndarray` that ais the same length as the [Network](../api/network.rst).
hopen = (short*-1)
hopen.s[:3,...]
rando = hopen *rand(len(hopen))
rando.s[:3,...]
# + raw_mimetype="text/restructuredtext" active=""
# .. notice ::
#
# Note that if you multiply a Network by an `numpy.ndarray` be sure to place the array on right side.
# -
# ## Comparison of Network
# Comparison operators also work with networks:
short == delayshort
short != delayshort
# ## Cascading and De-embedding
#
# Cascading and de-embeding 2-port Networks can also be done through operators. The `cascade` function can be called through the power operator, `**`. To calculate a new network which is the cascaded connection of the two individual Networks `line` and `short`,
short = rf.data.wr2p2_short
line = rf.data.wr2p2_line
delayshort = line ** short
# De-embedding can be accomplished by cascading the *inverse* of a network. The inverse of a network is accessed through the property `Network.inv`. To de-embed the `short` from `delay_short`,
# +
short_2 = line.inv ** delayshort
short_2 == short
# -
# The cascading operator also works for to 2N-port Networks. This is illustrated in [this example on balanced Networks](../examples/networktheory/Balanced%20Network%20De-embedding.ipynb). For example, assuming you want to cascade three 4-port Network `ntw1`, `ntw2` and `ntw3`, you can use:
# `resulting_ntw = ntw1 ** ntw2 ** ntw3`. Note that the port scheme assumed by the `**` cascading operator with 4-port networks is:
# + [markdown] raw_mimetype="text/markdown"
# ```
# ntw1 ntw2 ntw3
# +----+ +----+ +----+
# 0-|0 2|--|0 2|--|0 2|-2
# 1-|1 3|--|1 3|--|1 3|-3
# +----+ +----+ +----+
# ```
# -
# More documentation on Connecting Network is available here: [Connecting Networks](./Connecting_Networks.ipynb).
# ## Connecting Multi-ports
#
# **skrf** supports the connection of arbitrary ports of N-port networks. It accomplishes this using an algorithm called sub-network growth[[1]](#References), available through the function `connect()`.
#
# As an example, terminating one port of an ideal 3-way splitter can be done like so,
tee = rf.data.tee
tee
# To connect port `1` of the tee, to port `0` of the delay short,
terminated_tee = rf.connect(tee, 1, delayshort, 0)
terminated_tee
# Note that this function takes into account port impedances. If two connected ports have different port impedances, an appropriate impedance mismatch is inserted.
#
# More information on connecting Networks is available here: [connecting Networks](./Connecting_Networks.ipynb).
#
# For advanced connections between many arbitrary N-port Networks, the `Circuit` object is more relevant since it allows defining explicitly the connections between ports and Networks. For more information, please refer to the [Circuit documentation](Circuit.ipynb).
#
# ## Interpolation and Concatenation
#
# A common need is to change the number of frequency points of a [Network](../api/network.rst). To use the operators and cascading functions the networks involved must have matching frequencies, for instance. If two networks have different frequency information, then an error will be raised,
# +
from skrf.data import wr2p2_line1 as line1
line1
# -
# line1+line
#
# ---------------------------------------------------------------------------
# IndexError Traceback (most recent call last)
# <ipython-input-49-82040f7eab08> in <module>()
# ----> 1 line1+line
#
# /home/alex/code/scikit-rf/skrf/network.py in __add__(self, other)
# 500
# 501 if isinstance(other, Network):
# --> 502 self.__compatible_for_scalar_operation_test(other)
# 503 result.s = self.s + other.s
# 504 else:
#
# /home/alex/code/scikit-rf/skrf/network.py in __compatible_for_scalar_operation_test(self, other)
# 701 '''
# 702 if other.frequency != self.frequency:
# --> 703 raise IndexError('Networks must have same frequency. See `Network.interpolate`')
# 704
# 705 if other.s.shape != self.s.shape:
#
# IndexError: Networks must have same frequency. See `Network.interpolate`
#
#
# This problem can be solved by interpolating one of Networks along the frequency axis using `Network.resample`.
line1.resample(201)
line1
# And now we can do things
line1 + line
# You can also interpolate from a `Frequency` object. For example,
line.interpolate_from_f(line1.frequency)
# A related application is the need to combine Networks which cover different frequency ranges. Two Networks can be concatenated (aka stitched) together using `stitch`, which concatenates networks along their frequency axis. To combine a WR-2.2 Network with a WR-1.5 Network,
#
# +
from skrf.data import wr2p2_line, wr1p5_line
big_line = rf.stitch(wr2p2_line, wr1p5_line)
big_line
# -
# ## Reading and Writing
#
#
# For long term data storage, **skrf** has support for reading and partial support for writing [touchstone file format](http://en.wikipedia.org/wiki/Touchstone_file). Reading is accomplished with the Network initializer as shown above, and writing with the method `Network.write_touchstone()`.
#
# For **temporary** data storage, **skrf** object can be [pickled](http://docs.python.org/2/library/pickle.html) with the functions `skrf.io.general.read` and `skrf.io.general.write`. The reason to use temporary pickles over touchstones is that they store all attributes of a network, while touchstone files only store partial information.
rf.write('data/myline.ntwk',line) # write out Network using pickle
ntwk = Network('data/myline.ntwk') # read Network using pickle
# + raw_mimetype="text/restructuredtext" active=""
# .. warning::
#
# Pickling methods cant support long term data storage because they require the structure of the object being written to remain unchanged. something that cannot be guaranteed in future versions of skrf. (see http://docs.python.org/2/library/pickle.html)
#
# -
# Frequently there is an entire directory of files that need to be analyzed. `rf.read_all` creates Networks from all files in a directory quickly. To load all **skrf** files in the `data/` directory which contain the string `'wr2p2'`.
dict_o_ntwks = rf.read_all(rf.data.pwd, contains = 'wr2p2')
dict_o_ntwks
# Other times you know the list of files that need to be analyzed. `rf.read_all` also accepts a files parameter. This example file list contains only files within the same directory, but you can store files however your application would benefit from.
import os
dict_o_ntwks_files = rf.read_all(files=[os.path.join(rf.data.pwd, test_file) for test_file in ['ntwk1.s2p', 'ntwk2.s2p']])
dict_o_ntwks_files
# ## Other Parameters
#
# This tutorial focuses on s-parameters, but other network representations are available as well. Impedance and Admittance Parameters can be accessed through the parameters `Network.z` and `Network.y`, respectively. Scalar components of complex parameters, such as `Network.z_re`, `Network.z_im` and plotting methods are available as well.
#
# Other parameters are only available for 2-port networks, such as wave cascading parameters (`Network.t`), and ABCD-parameters (`Network.a`)
ring_slot.z[:3,...]
ring_slot.plot_z_im(m=1,n=0)
# ## References
#
#
# [1] <NAME>.; , "Perspectives in microwave circuit analysis," Circuits and Systems, 1989., Proceedings of the 32nd Midwest Symposium on , vol., no., pp.716-718 vol.2, 14-16 Aug 1989. URL: http://ieeexplore.ieee.org/stamp/stamp.jsp?tp=&arnumber=101955&isnumber=3167
#
| doc/source/tutorials/Networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext Cython
# + magic_args="--annotate" language="cython"
# import numpy as np
# import array
# from libc.time cimport clock
# #import time <-you can use it of course
# from cython cimport boundscheck, wraparound
#
# def main():
# cdef:
# # 点数
# int N = 100000000
# # 刻み
# double h = 0.00000004
# # 初期化
# #double[:] y=array.array('d',np.zeros(N))
# double[:] y=np.zeros(N)
# # 数値計算
# y[0]=1.0
# start = clock()
# cdef int n
# with boundscheck(False), wraparound(False):
# for n in range(N-1):
# y[n+1]=y[n]-h*y[n]
#
# elapsed_time = (clock() - start)/1000
# print("Elapsed Time(Python): %.2f [s]" % elapsed_time)
# -
main()
| cythonTest/eulerMethod/eulerCython.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Funzione print
a = 5 #operatore di assegnazione
b = 2
print('a =',a)
print('b =',b)
print('a+b =',a+b)
# ci sono 33 parole riservate in python
import keyword
parole_riservate = keyword.kwlist
print("Parole riservate= ",parole_riservate)
print("numero di parole riservate= ",len(parole_riservate))
print(a,b,12)
#print ha anche il parametro 'sep' (separator) e 'end'.
print(a,b,12,sep='?', end='#\n')
print(11,25, sep='.', end='\n')
print('Ciao','Belli', sep='-', end='\n')
# ### Input da tastiera
#esiste la funzione input(..) per passare delle stringhe(!!!) al programma in esecuzione,
#ed assegnarle a delle variabili.
#Esempio:
base_del_triangolo = input('base del Triangolo:')
#nella variabile base_del_triangolo verrà memorizzata una stringa.
# +
#Esercizio1
#domanda "come ti chiami?" Mario
#domanda "dove vivi?" Milano
#stampa : Mario vive a Milano.
nome = input("Come ti chiami?") # Mario
città = input('dove vivi?') # Milano
print(nome, 'vive a', città)
# +
#Esercizio2
#input "Base del rettangolo:" 4
#input "Altezza del rettangolo:" 3
#stampa : L'area del rettangolo è 12.
base = int(input('Base del rettangolo:'))
altezza = int(input('Altezza del rettangolo'))
print("L'area del rettangolo è", base*altezza)
# -
#con la funzione int(..) abbiamo convertito la stringa corrispondente alla base (e all'altezza)
#del triangolo da tipo stringa a tipo intero! Questo è uno step necessario se vogliamo
#effettuare operazioni algebriche tra le variabili.
type(base) #la funzione type(..) ritorna il tipo della variabile.
#Esercizio 3
#codice che prende in input un orario "europeo"(??) 15, 17, 19 e lo
#trasforma in orario americano 3,5,7 (le 12 e le 24 verranno convertite entrambi a 0)
orario_eu = int(input('orario europeo:'))
orario_us = orario_eu%12
print('orario americano:', orario_us)
#abbiamo usato l'operatore % (modulo).
# x%y ritorna il resto della divisione tra l'intero x e y.
# x//y tirona il quoziente delle divisione tra interi.
#Esercizio 4
#codice che prende in input i minuti 65 e lo trasforma in 1h:5min
minuti = int(input('minuti:'))
ore = minuti//60
minuti_resto = minuti - ore*60
print(ore,'h:',minuti_resto,'min',sep='')
# ### String.format
# format è un metodo della classe stringa che permette di inserire delle stringhe al posto di
# alcuni "placeholder" indentificati tramite parentesi graffe {} (da tastiera altGr+Shift+\[\] )
s = '{} - {}'.format('ciao', 'belli')
print(s)
#place order
s = '{1} {0}'.format('ciao', 'belli')
print(s)
s = '{1} {0}'.format('ciao', 'belli')
print(s)
s = '{1}-{1}-{0}'.format('ciao', 'belli')
print(s)
#assegnare delle chiavi
nome = 'Mario'
anni = 65
s = '{name} ha {age} anni'.format(name=nome,age=anni)
print(s)
#Esercizio4 (bis)
#prende in input i minuti 65 e lo trasforma in 1h:5min
minuti = int(input('minuti:'))
ore = minuti//60
minuti_resto = minuti - ore*60
orario_nuovo_formato='{}h:{}min'.format(ore,minuti)
print(orario_nuovo_formato)
# ### Funzioni
# + active=""
# def nome_funzione(a,b):
# return a+b
# -
def perimetro_quadrato(lato): #lato è un parametro di input
perimetro = 4*lato # scope della funzione
return perimetro
a = perimetro_quadrato(10)
print(a)
def raddoppia(n): # input
d=n*2
return d
def prodotto(n,m):
return n*m
prodotto(2,3)
prodotto(n=2,m=3)
def diviso(n,m):
return n/m
diviso(n=10,m=5)
diviso(m=10,n=5)
# le funzioni possono avere dei valori di default per alcuni parametri
def potenza(n,m):
p=n**m
return p
potenza(100)
# ### A cosa servono le funzioni?
#
#scrivere codice in maniera comprensibile e pulita
#esempio
partenza = '12:52'
arrivo = '17:23'
orario_partenza = partenza.split(':')
orario_arrivo = arrivo.split(':')
tot = int(orario_arrivo[1])+int(orario_arrivo[0])*60 - (int(orario_partenza[1])+int(orario_partenza[0])*60)
print('{}:{}'.format(tot//60,tot%60))
#inoltre evitiamo la duplicazione di codice.
def minuti_totali(orario_string):
orario = orario_string.split(':')
return int(orario[1])+int(orario[0])*60
def to_string(minuti):
return '{}:{}'.format(minuti//60,minuti%60)
partenza = '12:52'
arrivo = '17:23'
tot=minuti_totali(arrivo) - minuti_totali(partenza)
print(to_string(tot))
minuti_totali(arrivo)
# # scope di una funzione
# #le variabili vivono all'interno di uno scope (letteralmente: portata, campo, scopo) ben delimitato.
# #per esempio
# def potenza2(a):
# c=a*a
# return c
# #la variabile c non esiste al di fuori dello scope della funzione potenza2
# potenza2(2)
# print(c)
| Lez02/Lez2.ipynb |