code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Milestone2 (Post-Feedback)
#
# ## Names & ID
# * <NAME> (28222354)
# * <NAME> (18226282)
# ## Table of Contents
# * 1. Introduction
# * 2. Original Dataset
# * 3. Method Chaining
# * 4. Exploratory Data Analysis
# * 5. Analysis
# - 5.1 Research Questions
# - 5.2 Results
# * 6. Conclusion
# ## Introduction
# The "impacts.csv" is a dataset created by NASA, which includes a list of possible asteroid impacts, and characteristics of said asteroid such as probability, diameter, velocity, etc. NASA has gathered this information through their "Sentry" system, which is an automated collision monitoring system that scans through a catalog of asteroids to find possibilities of impact over the next 100 years. Our goal with this dataset is to do an Exploratory Data Analysis to help further our understanding. This will also help us answer any questions we may have on this dataset.
import pandas as pd
import seaborn as sns
import numpy as np
import matplotlib.pyplot as plt
# ## Original Dataset
# The DataFrame below includes information about possible asteriod impacts on earth and asteriod parameters. The dataset includes:
# - Columns named "Period Start and "Period End" show the initiation and ending of the risk factor.
# - Possible Impacts refers to a percentage of probability for impact
# - Asteriod Magnitude refers to the visual brightness relative to the earth , the higher the magnitude the less brightness and visibility.
# - Palermo Scale utilizes an in depth scale to quanitfy level of concern for future potential impacts.
# - Torino Scale indicates the impact's severity dedicated to notify the public.
#
df = pd.read_csv ('../../data/raw/impacts.csv')
df.head()
# ## Method Chaining
# - Through this method chaining I was able to get rid of all null values, assign units to parameters , and organize the data. I have renamed the colums for further description. I have also dropped the maximum torino scale because most of the values were 0 and would reproduce no plot.
# - I have used the load_and_process function. Calling upon this function will apply my method chaining to the dataframe.
#
df = (pd.read_csv('../../data/raw/impacts.csv')
.dropna()
.rename(columns={"Period Start": "Risk Period Start", "Period End": "Risk Period End", "Object Name": "Asteroid Name"})
.rename(columns={"Asteroid Velocity": "Asteroid Velocity (km/s)", "Asteroid Magnitude": "Asteroid Magnitude (m)"})
.drop(['Maximum Torino Scale'], axis=1)
.loc[0:450]
.sort_values(by='Risk Period Start', ascending=True)
.reset_index(drop=True)
)
df
from scripts import project_functions
df = project_functions.load_and_process('../../data/raw/impacts.csv')
df
# ## Exploratory Data Analysis
sns.set_theme(style="white",font_scale=1.4)
fig, x=plt.subplots(figsize=(35,8))
x = sns.countplot(x="Risk Period Start", data=df)
plt.title('Amount of Asteroids in each Risk Period Start')
x.set(ylabel='Number of Asteroids')
plt.tick_params(labelsize=13)
sns.despine()
plt.show()
# ### Observations
# In this part , I have used the count plot to demonstrate a branch of my data. I can see that the data is scattered throughout the diagram with no certain relationship between the two inputs. The year 2017 showed the highest number of asteriods in the initial risk period. The lowest number of asteriods (less than 5) is seen at the year 2048. The graph then fluctuates throughout the years.
fig, x=plt.subplots(figsize=(9,9))
sns.heatmap(df.corr(), annot = True, cmap = 'viridis')
plt.show()
# ### Observations
# - The heat map shows the correlation between the variables on each axis and the key on the right hand side of the image shows the correlation range.
# - Value closer to 0 such as risk period end and risk period start (0.0082) have no linear trend between the two variables
# - Value closer to 1 such as maxmium palermo scale and cumulative palermo scale (0.97) have positive correlation (as one increased so does the other)
# - value closer to -1 such as Cumulative Palermo Scale and Risk Period Start (-0.058) demonstrate that as one variable decreases another increases and vice versa
# - yellow squares are all 1 because those squares are correlating each variable to itself
sns.set_theme(style = 'white', font_scale = 1.7)
plt.subplots(figsize=(12,8))
plt.title('Lineplot Showing Relationship Between Probability for an Impact to Occur and Possible Impacts', fontsize = 16, y = 1.05)
graph1 = sns.lineplot(x = 'Possible Impacts', y = 'Cumulative Impact Probability', data = df)
graph1.set(xlabel = '# of Times Object can Potentially Impact Earth')
sns.despine()
# ### Observations
# This lineplot shows the relation between the number of possible impacts and the Cumulative Impact Probability. Other than an outlier just below 0.010, the rest of the data stays in between 0.000 and 0.004. Looking at this graph, I can conclude that even if the number of possible impacts increases, the cumulative impact probablity stays the same for the most part.
sns.set_theme(style = 'white', font_scale = 1.7)
plt.subplots(figsize=(12,8))
plt.title('How an Asteroids Velocity Affects # of Possible Impacts', fontsize = 16, y = 1.05)
graph4 = sns.histplot(x = 'Asteroid Velocity (km/s)', y = 'Possible Impacts', data = df)
graph4.set(ylabel = '# of Times Object can Potentially Impact Earth')
sns.despine()
# ### Observations
#
# This histplot shows the relationship between an asteroid's velocity and # possible impacts. From this plot, you can see around the 5-13 km/s range the # of possible impacts are the highest. As the velocity increases after 13 km/s, the # of possible impacts continues to get lower.
df2 = df[df["Cumulative Impact Probability"] < 1.500000e-07]
x = sns.relplot(data=df2, x="Possible Impacts", y="Cumulative Impact Probability")
plt.title('Relational Plot demonstrating the Probability of Asteroid Impacts', fontsize=12, y=1.05)
x.set(ylabel='Cumulative Probability of an Impact', xlabel = 'Possible Asteroid Impacts')
sns.despine()
plt.show()
# ### Comments
# For this section , I have used the relational plot. This plot shows the direct correlation between the number of impacts and the probabilities of the impacts. After feedback from the TA, I have added a column in the new dataframe called df2 to condense the data. The results show that asteriod impacts and their probabilties are skewed all over the plot. Although there is a less number in possible impacts for all ranges of probabilties , there are several outliers that show no concrete relationship between the two points of interest
sns.jointplot(x = 'Possible Impacts', y = 'Risk Period Start', kind = 'hex', data=df, color="#4CB391")
plt.show()
# ### Observations
# The jointplot demonstrates the amount of possible impacts and when their risk period initates. The plot clearly indicates that the year 2020 had the highest risk with more than 1000 possible impacts.The lowest risk period is revealed to be in 2040.
# ## Analysis
# ### Research Questions
# * Question 1 : What are the most common parameters that define an asteriod?
# * Question 2 : What factors of our dataset affect the probability of an impact occurring?
sns.set_theme(style="white",font_scale=1.4)
sns.displot(x = 'Asteroid Velocity (km/s)', data=df, bins = 25, color="Blue")
plt.title('Asteriod Count vs Asteriod Parameters')
sns.displot(x = 'Asteroid Magnitude (m)', data=df, bins = 25, color="Red")
plt.title('Asteriod Count vs Asteriod Parameters')
sns.displot(x = 'Asteroid Diameter (km)', data=df, bins = 25, color="Green")
plt.title('Asteriod Count vs Asteriod Parameters')
sns.despine()
plt.show()
# ### Observations
# These plots demonastrate relationships between asteriod counts and their relative parameters. We can see that the smaller the asteriod , the higher the count. Moreover , asteriod magnitudes of the range 25-30 show the highest counts.I have edited this section to produce a more colorful , visible plot demonstration
#
#
sns.set_theme(style="white",font_scale=1.4)
fig, x=plt.subplots(figsize=(15,10))
sns.lineplot(x="Asteroid Magnitude (m)", y="Asteroid Diameter (km)", data=df, palette="rocket")
plt.title('Relationship between Asteroid Mangnitude and Diameter')
sns.despine()
plt.show()
# ### Observations
# The graph decreases exponentially. In this plot , we can see the relationship between an asteriods diameter and magnitude. The higher the diamater the lower the magnitude of the asteriod. These parameters demonstrate and inverse relationship in our data. In other words , the wider the asteriod is in shape , the more visible and bright the asteriod is in relative to the earth.
sns.set_theme(style="white",font_scale=1.4)
sns.kdeplot(data=df, x = 'Asteroid Magnitude (m)', y = "Asteroid Velocity (km/s)", fill=True, thresh=0, levels=100, cmap="mako",)
plt.title('KDE Probability Distribution')
sns.despine()
plt.show()
# Observations:
# - This is a two-dimensional Kernel Distribution Estimation Plot which depicts the joint distribution both variables
# - Produce a plot that is less cluttered and more interpretable, especially when drawing multiple distributions
# - the lighter the area the more dense the distribution of variables is
#
sns.set_theme(style = 'white', font_scale = 1.5)
graph2 = sns.lmplot(x = 'Asteroid Diameter (km)', y = 'Cumulative Palermo Scale', data = df)
graph2.set(title = 'How an Asteroids Diameter Affects its Score on the Cumulative Palermo Scale', ylabel = 'Score on Cumulative Palermo Scale')
sns.despine()
# ### Observations
# This lmplot shows the relationship between the diameter of an asteroid and the Cumulative Palermo Scale. The Palermo Scale shows the seriousness of an impact, taking into account the probability of impact, and energy of impact. A number below -2 shows that there are no serious consequences, in between -2 and 0 shows that the object should be closely monitored, and above 0 means that there could be serious consequences. Looking at the plot, it shows that there is a positive relation between the diameter of the asteroid and the Cumulative Palermo Scale.
sns.set_theme(style = 'white', font_scale = 2)
plt.subplots(figsize=(12,8))
plt.title('How Cumulative Palermo Scale Score Affects the Probability for an Impact to Occur', fontsize = 16, y = 1.05)
graph3 = sns.scatterplot(x = 'Cumulative Palermo Scale', y = 'Cumulative Impact Probability', data = df)
graph3.set(xlabel = 'Score on Cumulative Palermo Scale')
sns.despine()
# ### Observations
# This scatterplot shows the relation between the Cumulative Palermo Scale and the Cumulative Impact Probability. Besides an outlier just below 0.010, most of the data ha a y-value in between 0.000 and 0.004. Towards the latter end of the plot, as the x-value increases, the y-value also slightly increases. Looking at this plot, I can conclude that there is a slight positive relation between the Cumulative Palermo Scale and the Cumulative Impact Probability
# ### Results
# * Question 1 : What are the most common parameters that define an asteriod?
# - Asteriods come in all shapes and sizes. Asteriod Classification has been conducted over the years in NASA's research. In this milestone , we come to discover the most common paramters that define an asteriod. First is Asteriod Velocity in space. Asteriods vary in velocity but seem to have a steady state of velocity between 6 and 15 km/s. Second, Asteriod sizes vary across the board but are most popular between 25 and 30 m in magnitude. Lastly , this parameter varies less across different asteriods as most demonstrate a diamater of around 0.2 Km.
#
# * Question 2 : What factors of our dataset affect the probability of an impact occurring?
# - There are a few factors of our dataset which affect the probability of an impact to occur. An important column in our dataset is the Cumulative Palermo Scale. This scale is important as it shows how serious an impact is, based on its energy at impact and the probability for one to occur. If you plot asteroids by diameter against the Cumulative Palermo Scale, it is shown that there is a positive relation between the two, meaning the larger the diameter of an asteroid results in a higher score on the scale. To further our understanding, if you plot asteroids by their score on the Cumulative Palermo Scale against their Cumulative Impact Probability, there is a slight positive relation between the two. This means that a higher score on the Cumulative Palermo Scale can lead to a slight increase in its Cumulative Impact Probability. Overall, factors like the diameter of an asteroid can affect the probability for an impact to occur.
# ## Conclusion
# Overall , the impacts.csv dataset shows a broad range of parameters and probabilties that might seem initally complex to dissect but has been analyzed according to our understanding.Annually , NASA conducts numerous projects to gather more information on asteriod parameters and their potential impacts.In this project , we scrutinize the dataset and gain valuable information from conducting an exploratory data analysis that reveals relationships between certain paramaters.Morever,we visualize the most common characteristics that define an asteriod.We then gather the factors of an asteriod and commence an analysis to see how they affect the probabilties of impacts.
| analysis/Task4/GroupSubmission.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
from random import sample
import networkx as nx
import numpy as np
from tqdm import tqdm, trange
sys.path.append('../')
from utils.aser_to_glucose import generate_aser_to_glucose_dict
from utils.glucose_utils import glucose_subject_list
# -
# Load the filtered ASER graph
aser = nx.read_gpickle('../../data/ASER_data//G_aser_core.pickle')
node2id_dict = np.load("../../dataset/ASER_core_node2id.npy", allow_pickle=True).item()
id2node_dict = dict([(node2id_dict[node], node) for node in node2id_dict])
# We test the coverage in the norm ASER
print_str = "\n\nStatistics in ASER_Norm:\n\n"
total_count, total_head, total_tail, total_both = 0, 0, 0, 0
for i in trange(1, 11):
list_count, list_head, list_tail, list_both = len(glucose_matching[i]), 0, 0, 0
for ind in range(len(glucose_matching[i])):
for h in glucose_matching[i][ind]['total_head']:
if h in node2id_dict.keys():
list_head += 1
break
for t in glucose_matching[i][ind]['total_tail']:
if t in node2id_dict.keys():
list_tail += 1
break
for h, t in glucose_matching[i][ind]['both']:
if h in node2id_dict.keys() and t in node2id_dict.keys():
list_both += 1
break
print_str += (
"In list {}, Total Head: {}\tMatched Head: {} ({}%)\tMatched Tail: {} ({}%)\tMatched Both: {} ({}%)\n"
.format(i, list_count, list_head, round(list_head / list_count, 3) * 100,
list_tail, round(list_tail / list_count, 3) * 100, list_both,
round(list_both / list_count, 3) * 100))
total_count += list_count
total_head += list_head
total_tail += list_tail
total_both += list_both
print_str += (
"\n\nIn total: Total Head: {}\tMatched Head: {} ({}%)\tMatched Tail: {} ({}%)\tMatched Both: {} ({}%)".format(
total_count, total_head, 100 * round(total_head / total_count, 3),
total_tail, 100 * round(total_tail / total_count, 3), total_both, 100 * round(total_both / total_count, 3)))
print(print_str)
# Do some ASER edge type statistics
all_edge_types = {}
for head, tail, feat_dict in aser.edges.data():
for r in feat_dict["edge_type"]:
if r in all_edge_types.keys():
all_edge_types[r] += 1
else:
all_edge_types[r] = 1
print("Edge types in ASER:")
print(all_edge_types)
def reverse_px_py(original: str):
return original.replace("PersonX", "[PX]").replace("PersonY", "[PY]").replace("[PX]", "PersonY").replace(
"[PY]", "PersonX")
def get_conceptualized_graph(G: nx.DiGraph):
G_conceptualized = nx.DiGraph()
for head, tail, feat_dict in tqdm(G.edges.data()):
head = id2node_dict[head]
tail = id2node_dict[tail]
head_split = head.split()
tail_split = tail.split()
head_subj = head_split[0]
tail_subj = tail_split[0]
relations = feat_dict["edge_type"]
for r in relations:
if head_subj == tail_subj and head_subj in glucose_subject_list:
new_rel = r + "_agent"
elif head_subj != tail_subj and head_subj in glucose_subject_list and tail_subj in glucose_subject_list:
new_rel = r + "_theme"
else:
new_rel = r + "_general"
_, re_head, re_tail, _ = generate_aser_to_glucose_dict(head, tail, True)
re_head_reverse, re_tail_reverse = reverse_px_py(re_head), reverse_px_py(re_tail)
if len(re_head) > 0 and len(re_tail) > 0:
if G_conceptualized.has_edge(re_head, re_tail):
G_conceptualized.add_edge(re_head, re_tail, relation=list(
set(G_conceptualized[re_head][re_tail]["relation"] + [new_rel])))
else:
G_conceptualized.add_edge(re_head, re_tail, relation=[new_rel])
if len(re_head_reverse) > 0 and len(re_tail_reverse) > 0:
if G_conceptualized.has_edge(re_head_reverse, re_tail_reverse):
G_conceptualized.add_edge(re_head_reverse, re_tail_reverse, relation=list(
set(G_conceptualized[re_head_reverse][re_tail_reverse]["relation"] + [new_rel])))
else:
G_conceptualized.add_edge(re_head_reverse, re_tail_reverse, relation=[new_rel])
return G_conceptualized
aser_conceptualized = get_conceptualized_graph(aser)
print("Before Conceptualization:\nNumber of Edges: {}\tNumber of Nodes: {}\n".format(len(aser.edges), len(aser.nodes)))
print("After Conceptualization:\nNumber of Edges: {}\tNumber of Nodes: {}\n".format(len(aser_conceptualized.edges),
len(aser_conceptualized.nodes)))
nx.write_gpickle(aser_conceptualized, '../../dataset/G_aser_concept.pickle')
# Let's sample some ASER conceptualization to check whether it's correct
for i in sample(list(aser_conceptualized.edges.data()), 30) + ['\n'] + sample(list(aser_conceptualized.nodes.data()), 10):
print(i)
# Now let's calculate the shortest path
def get_shortest_path(G , head, tail):
try:
p = nx.shortest_path_length(G, source=head, target=tail)
except nx.NodeNotFound:
return -1
except nx.NetworkXNoPath:
return -1
return p
full_path, norm_path = [], []
for i in range(1, 11):
for ind in trange(len(glucose_matching[i])):
norm_temp, full_temp = [], []
for h, t in glucose_matching[i][ind]['both']:
_, re_h, re_t, _ = generate_aser_to_glucose_dict(h, t, True)
if re_h in aser_conceptualized and re_t in aser_conceptualized:
norm_temp.append(get_shortest_path(aser_conceptualized, re_h, re_t))
if norm_temp:
try:
norm_path.append(min([i for i in norm_temp if i > 0]))
except ValueError:
norm_path.append(0)
else:
norm_path.append(0)
for h, t in glucose_matching[i][ind]['both']:
try:
hid = node2id_dict[h]
tid = node2id_dict[t]
except KeyError:
continue
if hid in aser and tid in aser:
full_temp.append(get_shortest_path(aser, hid, tid))
if full_temp:
try:
full_path.append(min([i for i in full_temp if i > 0]))
except ValueError:
full_path.append(0)
else:
full_path.append(0)
print("Average Shortest Path in Full ASER is: {}".format(np.mean([i for i in full_path if i > 0])))
print("Average Shortest Path in Norm ASER is: {}".format(np.mean([i for i in norm_path if i > 0])))
# +
# Calculate the average path in a simple graph:
G_simple = nx.Graph()
G_simple.add_nodes_from(aser_conceptualized)
G_simple.add_edges_from(aser_conceptualized.edges.data())
G_simple_full = nx.Graph()
G_simple_full.add_nodes_from(aser)
G_simple_full.add_edges_from(aser.edges.data())
full_path, norm_path = [], []
for i in range(1, 11):
for ind in trange(len(glucose_matching[i])):
norm_temp, full_temp = [], []
for h, t in glucose_matching[i][ind]['both']:
_, re_h, re_t, _ = generate_aser_to_glucose_dict(h, t, True)
if re_h in G_simple and re_t in G_simple:
norm_temp.append(get_shortest_path(G_simple, re_h, re_t))
if norm_temp:
try:
norm_path.append(min([i for i in norm_temp if i > 0]))
except ValueError:
norm_path.append(0)
else:
norm_path.append(0)
for h, t in glucose_matching[i][ind]['both']:
try:
hid = node2id_dict[h]
tid = node2id_dict[t]
except KeyError:
continue
if hid in G_simple_full and tid in G_simple_full:
full_temp.append(get_shortest_path(G_simple_full, hid, tid))
if full_temp:
try:
full_path.append(min([i for i in full_temp if i > 0]))
except ValueError:
full_path.append(0)
else:
full_path.append(0)
print("In No Direction Scenario:")
print("Average Shortest Path in Full ASER is: {}".format(np.mean([i for i in full_path if i > 0])))
print("Average Shortest Path in Norm ASER is: {}".format(np.mean([i for i in norm_path if i > 0])))
# -
# Now let's start merging with Glucose
G_Glucose = nx.read_gpickle('../../dataset/G_Glucose.pickle')
print("Node Coverage for Glucose Graph is: {}%\nEdge Coverage for Glucose Graph is: {}%".format(
100 * round(sum([node in aser_conceptualized for node in G_Glucose.nodes()]) / len(G_Glucose.nodes()), 4),
100 * round(sum([edge in aser_conceptualized.edges for edge in G_Glucose.edges()]) / len(G_Glucose.edges()), 4)))
print("Before Merging:\nEdges in ASER: {}\t\t\t\tNodes in ASER: {}\n".format(len(aser_conceptualized.edges()),
len(aser_conceptualized.nodes())))
aser_conceptualized.add_nodes_from(list(G_Glucose.nodes.data()))
aser_conceptualized.add_edges_from(list(G_Glucose.edges.data()))
print("\nAfter Merging:\nEdges in ASER+Glucose: {}\t\t\tNodes in ASER+Glucose: {}".format(len(aser_conceptualized.edges()),
len(aser_conceptualized.nodes())))
print("New Edges: {}\tNew Nodes: {}".format(len(aser_conceptualized.edges()) - 41336290,
len(aser_conceptualized.nodes()) - 11872745))
nx.write_gpickle(aser_conceptualized, '../../dataset/G_aser_glucose.pickle')
| preprocess/Glucose/build_graph/conceptualize_ASER.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Model Comparison
# ## Mixture of Unigram Model
#
# To reproduce the model results on the simulated data, please follow the following instruction:
# 1. Here we provide the code to preprocessing the simulated text file.
# 2. For this model to run, dowload the text file generated by the program below, turn this txt file into Unix Executable file in your local terminal. Re-upload and run with UMM model.
# 3. Go to Jupyter terminal, run "$ python pDMM.py --corpus simu --ntopics 10 --twords 10 --niters 500 --name unigram"
# 4. Check the "output" folder and the file named "unigram.topWords" produced.
import string
with open("stopword.txt", 'r') as s:
stopwords = s.readlines()
stpw = []
for word in stopwords:
stpw.append(word.strip())
with open("simulated.txt") as f:
corpus = f.readlines()
word_list = []
for line in corpus:
if line != "":
before = line.strip().split()
#Remove stopwords from the strings
for word in before:
if word.lstrip(string.punctuation).rstrip(string.punctuation).lower() not in stpw:
if word != "":
word_list.append(word.lstrip(string.punctuation).rstrip(string.punctuation).strip().lower())
new = []
for i in range(0,234,1):
line = " ".join(word_list[i*5:i*5+5])
new.append(line)
simu = "\n".join(new)
# +
output_file = open('simu.txt','w')
output_file.write(simu)
output_file.close()
#Then turn this txt file into Unix Executable file in local terminal. Reupload and run with UMM model.
# -
# #### Mixture of Unigrams Results
# Topic 0: luffy piece search treasure head law monkey fruit named pirate
#
# Topic 1: devil fruit user fruits animals race sea power powers haki
#
# Topic 2: sea grand red water mountain half rain runs seas wind
#
# Topic 3: series pirates roger humans gol merry manga video animation produced
#
# Topic 4: crew luffy robin ancient liberates sabaody archipelago ace navy government
#
# Topic 5: crew blue pirates navy straw joins named grand east pirate
#
# Topic 6: luffy nami sanji arlong chopper body properties crew usopp encounters
#
# Topic 7: pose calm belts log developed called thirteen animated feature films
#
# Topic 8: piece manga pirates king set island zou series eiichiro history
#
# Topic 9: pirates straw island luffy hat grand kingdom magnetic island's fishman
# ## Topic extraction with Non-negative Matrix Factorization (pLSI)
#
#
# This is an example of applying :class:`sklearn.decomposition.NMF` on a corpus
# of documents and extract additive models of the topic structure of the
# corpus. The output is a list of topics, each represented as a list of
# terms (weights are not shown).
#
# Non-negative Matrix Factorization has two different objective
# functions: the Frobenius norm, and the generalized Kullback-Leibler divergence.
# The latter is equivalent to Probabilistic Latent Semantic Indexing.
#
# The default parameters (n_samples / n_features / n_components) should make
# the example runnable in a couple of tens of seconds. You can try to
# increase the dimensions of the problem, but be aware that the time
# complexity is polynomial in NMF. In LDA, the time complexity is
# proportional to (n_samples * iterations).
# +
from sklearn.feature_extraction.text import TfidfVectorizer
import sklearn
from sklearn.decomposition import NMF
n_samples = 16
n_features = 1000
n_components = 10
n_top_words = 10
# -
def print_top_words(model, feature_names, n_top_words):
for topic_idx, topic in enumerate(model.components_):
message = "Topic #%d: " % topic_idx
message += " ".join([feature_names[i]
for i in topic.argsort()[:-n_top_words - 1:-1]])
print(message)
print()
import LDApackage
simulated_docs = LDApackage.read_documents_space('simulated.txt')
# Use tf-idf features for NMF.
tfidf_vectorizer = TfidfVectorizer(max_df=0.95, min_df=2,
max_features=n_features,
stop_words='english')
tfidf = tfidf_vectorizer.fit_transform(simulated_docs)
# +
# Fit the NMF model
nmf = sklearn.decomposition.NMF(n_components=n_components, random_state=1,
beta_loss='kullback-leibler', solver='mu', max_iter=1000, alpha=.1,
l1_ratio=.5).fit(tfidf)
#print("\nTopics in NMF model (generalized Kullback-Leibler divergence):")
tfidf_feature_names = tfidf_vectorizer.get_feature_names()
#print_top_words(nmf, tfidf_feature_names, n_top_words)
# -
# #### Results for NMF model (generalized Kullback-Leibler divergence):
# Topic 0: pirates line crew man search roger known king grand luffy
#
# Topic 1: history japan series date manga oda eiichiro body volumes world
#
# Topic 2: body devil animals used result users power presence time fruit
#
# Topic 3: grand time called sea currents line works island making specific
#
# Topic 4: law defeat mom sanji alliance caesar nami clown big straw
#
# Topic 5: group battles robin straw franky leading crew ancient save pluton
#
# Topic 6: used animals wind similar piece world various certain presence eiichiro
#
# Topic 7: ace huge grand luffy adoptive forced fish thousand led new
#
# Topic 8: soon cyborg island crew fishmen sabaody battle alias world archipelago
#
# Topic 9: blue humans usopp government going captures water sanji defeats creatures
# ## Discussion
# Some of the topics for mixture of unigram do not make sense. For example, topic 7 for mixture of unigram contains at least two topics one is related to 'belts', 'calm' and another is related to 'animated', 'film'. The inaccuracy may stem from the model itself which assumes that a document can only contain a single topic.
#
# For this simulated dataset, NMF model with KL divergence also exhibits unique topics across the results. It actually makes sense because NMF with KL divergence is equivalent to probabilistic latent semantic analysis (pLSA) when optimizing the same objective function. The two approaches only differ in how inference proceeds, but the underlying model is the same. Compared with LDA, NMF also model the total count of words in a document. For LDA, the total count of words in a document is assumed given.
#
# NMF with KL divergence (pLSI) runs faster than LDA, which can make a big difference if we want to apply topic modeling to large-scale datasets. Also, it is much easier to parallelize PLSI, at least compared to Gibbs sampling, not to mention the complicated implementation of the variational bayes approaches.
| Comparison.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import cv2
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import pickle
# Read in an image
image = mpimg.imread('sobel_direction.png')
# Define a function that applies Sobel x and y,
# then computes the direction of the gradient
# and applies a threshold.
def dir_threshold(img, sobel_kernel=3, thresh=(0, np.pi/2)):
# Apply the following steps to img
# 1) Convert to grayscale
gray=cv2.cvtColor(image,cv2.COLOR_RGB2GRAY)
# 2) Take the gradient in x and y separately
Sobelx=cv2.Sobel(gray,cv2.CV_64F,1,0,ksize=sobel_kernel)
Sobely=cv2.Sobel(gray,cv2.CV_64F,0,1,ksize=sobel_kernel)
# 3) Take the absolute value of the x and y gradients
abs_sobelx=np.absolute(Sobelx)
abs_sobely=np.absolute(Sobely)
# 4) Use np.arctan2(abs_sobely, abs_sobelx) to calculate the direction of the gradient
grad_direction=np.arctan2(abs_sobely,abs_sobelx)
# 5) Create a binary mask where direction thresholds are met
binary_output=np.zeros_like(grad_direction)
binary_output[(grad_direction > thresh[0]) & (grad_direction < thresh[1])] = 1
# 6) Return this mask as your binary_output image
return binary_output
# Run the function
dir_binary = dir_threshold(image, sobel_kernel=15, thresh=(0.7,1.3))
# Plot the result
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(24, 9))
f.tight_layout()
ax1.imshow(image)
ax1.set_title('Original Image', fontsize=50)
ax2.imshow(dir_binary, cmap='gray')
ax2.set_title('Thresholded Grad. Dir.', fontsize=50)
plt.subplots_adjust(left=0., right=1, top=0.9, bottom=0.)
| Sobel/sobel direction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# [← Back to Index](index.html)
# # Longest Common Subsequence
# To motivate dynamic time warping, let's look at a classic dynamic programming problem: find the **longest common subsequence (LCS)** of two strings ([Wikipedia](https://en.wikipedia.org/wiki/Longest_common_subsequence_problem)). A subsequence is not required to maintain consecutive positions in the original strings, but they must retain their order. Examples:
#
# lcs('cake', 'baker') -> 'ake'
# lcs('cake', 'cape') -> 'cae'
# We can solve this using recursion. We must find the optimal substructure, i.e. decompose the problem into simpler subproblems.
#
# Let `x` and `y` be two strings. Let `z` be the true LCS of `x` and `y`.
#
# If the first characters of `x` and `y` were the same, then that character must also be the first character of the LCS, `z`. In other words, if `x[0] == y[0]`, then `z[0]` must equal `x[0]` (which equals `y[0]`). Therefore, append `x[0]` to `lcs(x[1:], y[1:])`.
#
# If the first characters of `x` and `y` differ, then solve for both `lcs(x, y[1:])` and `lcs(x[1:], y)`, and keep the result which is longer.
# Here is the recursive solution:
def lcs(x, y):
if x == "" or y == "":
return ""
if x[0] == y[0]:
return x[0] + lcs(x[1:], y[1:])
else:
z1 = lcs(x[1:], y)
z2 = lcs(x, y[1:])
return z1 if len(z1) > len(z2) else z2
# Test:
pairs = [
('cake', 'baker'),
('cake', 'cape'),
('catcga', 'gtaccgtca'),
('zxzxzxmnxzmnxmznmzxnzm', 'nmnzxmxzmnzmx'),
('dfkjdjkfdjkjfdkfdkfjd', 'dkfjdjkfjdkjfkdjfkjdkfjdkfj'),
]
for x, y in pairs:
print lcs(x, y)
# The time complexity of the above recursive method is $O(2^{n_x+n_y})$. That is slow because we might compute the solution to the same subproblem multiple times.
# ### Memoization
# We can do better through memoization, i.e. storing solutions to previous subproblems in a table.
# Here, we create a table where cell `(i, j)` stores the length `lcs(x[:i], y[:j])`. When either `i` or `j` is equal to zero, i.e. an empty string, we already know that the LCS is the empty string. Therefore, we can initialize the table to be equal to zero in all cells. Then we populate the table from the top left to the bottom right.
def lcs_table(x, y):
nx = len(x)
ny = len(y)
# Initialize a table.
table = [[0 for _ in range(ny+1)] for _ in range(nx+1)]
# Fill the table.
for i in range(1, nx+1):
for j in range(1, ny+1):
if x[i-1] == y[j-1]:
table[i][j] = 1 + table[i-1][j-1]
else:
table[i][j] = max(table[i-1][j], table[i][j-1])
return table
# Let's visualize this table:
x = 'cake'
y = 'baker'
table = lcs_table(x, y)
table
xa = ' ' + x
ya = ' ' + y
print ' '.join(ya)
for i, row in enumerate(table):
print xa[i], ' '.join(str(z) for z in row)
# Finally, we backtrack, i.e. read the table from the bottom right to the top left:
def lcs(x, y, table, i=None, j=None):
if i is None:
i = len(x)
if j is None:
j = len(y)
if table[i][j] == 0:
return ""
elif x[i-1] == y[j-1]:
return lcs(x, y, table, i-1, j-1) + x[i-1]
elif table[i][j-1] > table[i-1][j]:
return lcs(x, y, table, i, j-1)
else:
return lcs(x, y, table, i-1, j)
for x, y in pairs:
table = lcs_table(x, y)
print lcs(x, y, table)
# Table construction has time complexity $O(mn)$, and backtracking is $O(m+n)$. Therefore, the overall running time is $O(mn)$.
# [← Back to Index](index.html)
| lcs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table class="ee-notebook-buttons" align="left">
# <td><a target="_parent" href="https://github.com/giswqs/geemap/tree/master/examples/notebooks/geemap_and_earthengine.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
# <td><a target="_parent" href="https://nbviewer.jupyter.org/github/giswqs/geemap/blob/master/examples/notebooks/geemap_and_earthengine.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
# <td><a target="_parent" href="https://colab.research.google.com/github/giswqs/geemap/blob/master/examples/notebooks/geemap_and_earthengine.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
# </table>
# ## Install Earth Engine API and geemap
# Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
# The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
# +
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# -
import ee
import geemap
# ## Create an interactive map
Map = geemap.Map(center=(40, -100), zoom=4)
Map
# ## Add Earth Engine Python script
# +
# Add Earth Engine dataset
image = ee.Image('USGS/SRTMGL1_003')
# Set visualization parameters.
vis_params = {
'min': 0,
'max': 4000,
'palette': ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']}
# Print the elevation of Mount Everest.
xy = ee.Geometry.Point([86.9250, 27.9881])
elev = image.sample(xy, 30).first().get('elevation').getInfo()
print('Mount Everest elevation (m):', elev)
# Add Earth Engine layers to Map
Map.addLayer(image, vis_params, 'SRTM DEM', True, 0.5)
Map.addLayer(xy, {'color': 'red'}, 'Mount Everest')
# -
# ## Change map positions
#
# For example, center the map on an Earth Engine object:
Map.centerObject(ee_object=xy, zoom=13)
# Set the map center using coordinates (longitude, latitude)
Map.setCenter(lon=-100, lat=40, zoom=4)
# ## Extract information from Earth Engine data based on user inputs
# +
import ee
import geemap
from ipyleaflet import *
from ipywidgets import Label
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
Map = geemap.Map(center=(40, -100), zoom=4)
Map.default_style = {'cursor': 'crosshair'}
# Add Earth Engine dataset
image = ee.Image('USGS/SRTMGL1_003')
# Set visualization parameters.
vis_params = {
'min': 0,
'max': 4000,
'palette': ['006633', 'E5FFCC', '662A00', 'D8D8D8', 'F5F5F5']}
# Add Earth Eninge layers to Map
Map.addLayer(image, vis_params, 'SRTM DEM', True, 0.5)
latlon_label = Label()
elev_label = Label()
display(latlon_label)
display(elev_label)
coordinates = []
markers = []
marker_cluster = MarkerCluster(name="Marker Cluster")
Map.add_layer(marker_cluster)
def handle_interaction(**kwargs):
latlon = kwargs.get('coordinates')
if kwargs.get('type') == 'mousemove':
latlon_label.value = "Coordinates: {}".format(str(latlon))
elif kwargs.get('type') == 'click':
coordinates.append(latlon)
# Map.add_layer(Marker(location=latlon))
markers.append(Marker(location=latlon))
marker_cluster.markers = markers
xy = ee.Geometry.Point(latlon[::-1])
elev = image.sample(xy, 30).first().get('elevation').getInfo()
elev_label.value = "Elevation of {}: {} m".format(latlon, elev)
Map.on_interaction(handle_interaction)
Map
# +
import ee
import geemap
from ipyleaflet import *
from bqplot import pyplot as plt
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
Map = geemap.Map(center=(40, -100), zoom=4)
Map.default_style = {'cursor': 'crosshair'}
# Compute the trend of nighttime lights from DMSP.
# Add a band containing image date as years since 1990.
def createTimeBand(img):
year = img.date().difference(ee.Date('1991-01-01'), 'year')
return ee.Image(year).float().addBands(img)
NTL = ee.ImageCollection('NOAA/DMSP-OLS/NIGHTTIME_LIGHTS') \
.select('stable_lights')
# Fit a linear trend to the nighttime lights collection.
collection = NTL.map(createTimeBand)
fit = collection.reduce(ee.Reducer.linearFit())
image = NTL.toBands()
figure = plt.figure(1, title='Nighttime Light Trend', layout={'max_height': '250px', 'max_width': '400px'})
count = collection.size().getInfo()
start_year = 1992
end_year = 2013
x = range(1, count+1)
coordinates = []
markers = []
marker_cluster = MarkerCluster(name="Marker Cluster")
Map.add_layer(marker_cluster)
def handle_interaction(**kwargs):
latlon = kwargs.get('coordinates')
if kwargs.get('type') == 'click':
coordinates.append(latlon)
markers.append(Marker(location=latlon))
marker_cluster.markers = markers
xy = ee.Geometry.Point(latlon[::-1])
y = image.sample(xy, 500).first().toDictionary().values().getInfo()
plt.clear()
plt.plot(x, y)
# plt.xticks(range(start_year, end_year, 5))
Map.on_interaction(handle_interaction)
# Display a single image
Map.addLayer(ee.Image(collection.select('stable_lights').first()), {'min': 0, 'max': 63}, 'First image')
# Display trend in red/blue, brightness in green.
Map.setCenter(30, 45, 4)
Map.addLayer(fit,
{'min': 0, 'max': [0.18, 20, -0.18], 'bands': ['scale', 'offset', 'scale']},
'stable lights trend')
fig_control = WidgetControl(widget=figure, position='bottomright')
Map.add_control(fig_control)
Map
| examples/notebooks/geemap_and_earthengine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# this file reads in the data from the GEO submission from Wen at al (JCI, 2019)
# data from each cell is provided in a seprarate file
import os
import glob
import pandas as pd
import numpy as np
files = glob.glob('WenData/Data/*gencode*gz')
len(files)
# iterate across all files and join to a single data frame
i = 0
for curr in files:
os.system('gzip -d ' + curr)
decompressed_file = curr[:-3]
curr_df = pd.read_csv(decompressed_file, sep = '\t', index_col= 0)
vec = curr_df.tpm
colName = '_'.join(decompressed_file.split('_')[1:4]) + str(i)
if curr == files[0]:
master_df = pd.DataFrame(vec)
master_df.columns = [colName]
else:
master_df[colName] = vec
i = i+1
os.system('gzip ' + decompressed_file)
master_df['gene'] = master_df.index
master_df = master_df.drop('gene', axis = 1)
# +
genes = [x.split('_')[0] for x in master_df.index]
genelist = list(set(genes))
for curr in genelist:
rows = master_df.iloc[np.where(np.array(genes) == curr)]
rows.sum(axis = 0)
vec = rows.sum(axis = 0)
if curr == genelist[0]:
df = pd.DataFrame(vec)
df.columns = [curr]
else:
df[curr] = vec
# -
df = pd.DataFrame(rows.sum(axis = 0))
df.columns =[ curr]
df
trans = df.transpose()
trans = trans.reset_index()
feather.write_dataframe(trans, 't.feather')
| Figures/Figure 4/wen_python_assembly.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pypermr
# language: python
# name: pypermr
# ---
# ### PySetPerm design
# The pysetperm.py module includes a number of classes that provide simple building blocks for testing set enrichments.
# Features can be anything: genes, regulatory elements etc. as long as they have chr, start (1-based!), end(1-based) and name columns:
# + language="bash"
# cd ..
# head -n3 data/genes.txt
# -
# Annotations are also simply specified:
# + language="bash"
# head -n3 data/kegg.txt
# -
# ### An example analysis
# We import features and annotaions via respective classes. Features can be altered with a distance (i.e. genes +- 2000 bp). Annotations can also be filtered to have a minimum set size (i.e. at least 5 genes)
import pysetperm as psp
features = psp.Features('data/genes.txt', 2000)
annotations = psp.AnnotationSet('data/kegg.txt', features.features_user_def, 5)
n_perms = 200000
cores = 10
# Initiate test groups using the Input class:
# +
e_input = psp.Input('data/eastern_candidates.txt',
'data/eastern_background.txt.gz',
features,
annotations)
c_input = psp.Input('data/central_candidates.txt',
'data/central_background.txt.gz',
features,
annotations)
i_input = psp.Input('data/internal_candidates.txt',
'data/internal_background.txt.gz',
features,
annotations)
# -
# A Permutation class holds the permuted datasets.
e_permutations = psp.Permutation(e_input, n_perms, cores)
c_permutations = psp.Permutation(c_input, n_perms, cores)
i_permutations = psp.Permutation(i_input, n_perms, cores)
# Once permutions are completed, we determine the distribution of the Pr. X of genes belonging to Set1...n, using the SetPerPerm class. This structure enables the easy generation of joint distributions.
e_per_set = psp.SetPerPerm(e_permutations,
annotations,
e_input,
cores)
c_per_set = psp.SetPerPerm(c_permutations,
annotations,
c_input,
cores)
i_per_set = psp.SetPerPerm(i_permutations,
annotations,
i_input,
cores)
# Here, we can use join_objects() methods for both Imput and SetPerPerm objects, to get the joint distribution of two or more indpendent tests.
# combine sims
ec_input = psp.Input.join_objects(e_input, c_input)
ec_per_set = psp.SetPerPerm.join_objects(e_per_set, c_per_set)
ei_input = psp.Input.join_objects(e_input, i_input)
ei_per_set = psp.SetPerPerm.join_objects(e_per_set, i_per_set)
ci_input = psp.Input.join_objects(c_input, i_input)
ci_per_set = psp.SetPerPerm.join_objects(c_per_set, i_per_set)
eci_input = psp.Input.join_objects(ec_input, i_input)
eci_per_set = psp.SetPerPerm.join_objects(ec_per_set, i_per_set)
# Call the make_results_table function to generate a pandas format results table.
# results
e_results = psp.make_results_table(e_input, annotations, e_per_set)
c_results = psp.make_results_table(c_input, annotations, c_per_set)
i_results = psp.make_results_table(i_input, annotations, i_per_set)
ec_results = psp.make_results_table(ec_input, annotations, ec_per_set)
ei_results = psp.make_results_table(ei_input, annotations, ei_per_set)
ci_results = psp.make_results_table(ci_input, annotations, ci_per_set)
eci_results = psp.make_results_table(eci_input, annotations, eci_per_set)
from itables import show
from IPython.display import display
from ipywidgets import HBox, VBox
import ipywidgets as widgets
display(e_results)
display(c_results)
display(i_results)
display(ec_results)
display(ei_results)
display(ci_results)
display(eci_results)
| notebooks/.ipynb_checkpoints/example_analysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## PHYS 105A: Introduction to Scientific Computing
#
# # Unix shells, Remote Login, Version Control, etc
#
# <NAME>
# In order to follow this hands-on exercise, you will need to have the following software installed on your system:
#
# * Terminal (built-in for windows, mac, and linux)
# * A Unix shell (Windows Subsystem for Linux; built-in for mac and linux)
# * Git
# * Jupyter Notebook or Jupyter Lab
# * Python 3.x
#
# The easiest way to install these software is to use [Anaconda](https://www.anaconda.com/).
# Nevertheless, you may use macports or homebrew to install these tools on Mac, and `apt` to install these tools on Linux. For windows, you may enable the Windows Subsystem for Linux (WSL) to get a Unix shell.
# ## Terminal and Unix Shell
#
# This step is trivial if you are on Linux or on a Mac. Simply open up your terminal and you will be given `bash` or `zsh`.
#
# If you are on Windows, you may start `cmd`, "Windows PowerShell", or the "Windows Subsystem for Linux".
# The former two don't give you a Unix shell, but you can still use it to navigate your file system, run `git`, etc.
# The last option is actually running a full Ubuntu envirnoment that will give you `bash`.
# ## Once you are on a bash/zsh, try the following commands
#
# echo "Hello World" # the infamous hello world program!
# ls
# echo "Hello World" > hw.txt
# ls
# cat hw.txt
# mv hw.txt hello.txt
# cp hello.txt world.txt
#
# Awesome! You are now a shell user!
# ## Git
#
# ### Pre-request
#
# * Have a GitHub account
# * Have Jupyter Lab set up on your machine
# * Have `git` (can be installed by conda)
#
# ### Set up development environment
#
# * `git config --global user.name "<NAME>"`
# * `git config --global user.email <EMAIL>`
#
# [Reference](https://git-scm.com/book/en/v2/Getting-Started-First-Time-Git-Setup)
# ## Clone Course Repository
#
# With a shell, you can now clone/copy the PHYS105A repository to your computer:
#
# git clone <EMAIL>:uarizona-2021spring-phys105a/phys105a.git
# cd phys105a
# ls
#
# You may now add a file, commit it to git, and push it back to GitHub:
#
# echo "Hello World" > hw.txt
# git add .
# git commit -m 'hello world'
# git log
# git push
# ## Jupyter Notebook
#
# A Jupyter Notebook is a JSON document containing multiple cells of text and codes. These cells maybe:
#
# * Markdown documentation
# * Codes in `python`, `R`, or even `bash`!
# * Output of the codes
# * RAW text
# ## Markdown Demo
#
# This is a markdown demo. Try edit this cell to see how things work.
#
# See https://guides.github.com/features/mastering-markdown/ for more details.
#
#
# # This is an `<h1>` tag
# ## This is an `<h2>` tag
# ###### This is an `<h6>` tag
#
# *This text will be italic*
# _This will also be italic_
#
# **This text will be bold**
# __This will also be bold__
#
# _You **can** combine them_
#
# * Item 1
# * Item 2
# * Item 2a
# * Item 2b
#
# 1. Item 1
# 1. Item 2
# 1. Item 3
# 1. Item 3a
# 1. Item 3b
# +
## Python and Jupyter
# We can finally try jupyter and python as your calculator.
# Type some math formulas, then press Shift+Enter, jupyter will interpret your equations by python and print the output.
# This is CK's equation:
1 + 2 + 3 + 4
# +
# EXERCISE: Now, type your own equation here and see the outcome.
# +
# EXERCISE: Try to create more cells and type out more equations.
# Note that Jupyter supports "shortcuts"/"hotkeys".
# When you are typing in a cell, there is a green box surrounding the cell.
# You may click outside the cell or press "ESC" to escape from the editing mode.
# The green box will turn blue.
# You can then press "A" or "B" to create additional cells above or below the current active cell.
# +
# One of the most powerful things programming langauges are able to do is to assign names to values.
# This is in contrast to spreadsheet software where each "cell" is prenamed (e.g., A1, B6).
# A value associated with a name is called a varaible.
# In python, we simply use the equal sign to assign a value to a variable.
a = 1
b = 1 + 1
# We can then reuse these variables.
a + b + 3
# +
# EXERCISE: create your own variables and check their values.
# +
# Sometime it is convenient to have mutiple output per cell.
# In such a case, you may use the python function `print()`
# Or the Jupyter function `display()`
print(1, 2)
print(a, b)
display(1, 2)
display(a, b)
# +
# EXERCISE: print and display results of equations
# +
# Speaking of print(), in python, you may use both single or double quotes to create "string" of characters.
'This is a string of characters.'
"This is also a string of characters."
# You can mix strings, numbers, etc in a single print statement.
print("Numbers:", 1, 2, 3)
# +
# EXERCISE: assign a string to a variable and then print it
# +
# In the lecture, we learned the different math operations in python: +, -, *, **, /, and //.
# Try to use them yourself.
# Pay special attention to the ** and // operators.
# * and ** are different
print(10*3)
print(10**3)
# / and // are different
print(10/3)
print(10//3)
# +
# EXERCISE: try out *, **, /, and // yourself.
# What if you use them for very big numbers? Very small numbers?
# Do you see any limitation?
# +
# COOL TRICK: In python, you use underscores to help writing very big numbers.
# E.g., python knows that 1_000_000 is 1000_000 is 1000000.
print(1_000_000)
print(1000_000)
print(1000000)
# +
# The integer division is logically equivalent to applying a floor function to the floating point division.
# However, the floor function is not a default (built-in) function.
# You need to import it from the math package:
from math import floor
print(10/3)
print(10//3)
print(floor(10/3))
# +
# EXERCISE: try to use the floor() function yourself
# +
# There are many useful functions and constants in the math package.
# See https://docs.python.org/3/library/math.html
from math import pi, sin, cos
print(pi)
print(sin(pi))
print(cos(pi))
# +
# EXERCISE: try to import additional functions from the math package and test them yourself.
# Python math package: https://docs.python.org/3/library/math.html
| 02/Handson.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # OpenEO Connection to EODC Backend
import logging
import openeo
from openeo.auth.auth_bearer import BearerAuth
logging.basicConfig(level=logging.INFO)
# +
# Define constants
# Connection
EODC_DRIVER_URL = "http://openeo.eodc.eu/api/v0.3"
OUTPUT_FILE = "/tmp/openeo_eodc_output.tiff"
EODC_USER = "user1"
EODC_PWD = "<PASSWORD>#"
# Data
PRODUCT_ID = "s2a_prd_msil1c"
DATE_START = "2017-01-01"
DATE_END = "2017-01-08"
IMAGE_LEFT = 652000
IMAGE_RIGHT = 672000
IMAGE_TOP = 5161000
IMAGE_BOTTOM = 5181000
IMAGE_SRS = "EPSG:32632"
# Processes
NDVI_RED = "B04"
NDVI_NIR = "B08"
# +
# Connect with EODC backend
connection = openeo.connect(EODC_DRIVER_URL,auth_type=BearerAuth, auth_options={"username": EODC_USER, "password": EODC_PWD})
# Login
#token = session.auth(EODC_USER, EODC_PWD, BearerAuth)
connection
# -
# Get available processes from the back end.
processes = connection.list_processes()
processes
# +
# Retrieve the list of available collections
collections = connection.list_collections()
list(collections)[:2]
# -
# Get detailed information about a collection
process = connection.describe_collection(PRODUCT_ID)
process
# +
# Select collection product
datacube = connection.imagecollection(PRODUCT_ID)
print(datacube.to_json())
# +
# Specifying the date range and the bounding box
datacube = datacube.filter_bbox(west=IMAGE_WEST, east=IMAGE_EAST, north=IMAGE_NORTH,
south=IMAGE_SOUTH, crs=IMAGE_SRS)
datacube = datacube.filter_daterange(extent=[DATE_START, DATE_END])
print(datacube.to_json())
# +
# Applying some operations on the data
datacube = datacube.ndvi(red=NDVI_RED, nir=NDVI_NIR)
datacube = datacube.min_time()
print(datacube.to_json())
# -
# Sending the job to the backend
job = datacube.send_job()
job.start_job()
job
# Describe Job
job.describe_job()
# +
# Download job result
#from openeo.rest.job import ClientJob
#job = ClientJob(107, session)
job.download(OUTPUT_FILE)
job
# -
# Showing the result
# !gdalinfo -hist "/tmp/openeo_eodc_output.tiff"
| examples/notebooks/EODC_Forum_2019/EODC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# binary search tree
# left_subtree (keys) ≤ node (key) ≤ right_subtree (keys)
class Node:
def __init__(self, data):
self.left = None
self.right = None
self.data = data
# insert method to create nodes
def insert(self, data):
if self.data: # if self.data is not None
if data < self.data: # check left subtree
if self.left is None: # if left subtree not exist
self.left = Node(data) # create a new node
else: # if left subtree is empty
self.left.insert(data) # insert data into it
elif data > self.data:
if self.right is None:
self.right = Node(data)
else:
self.right.insert(data)
else: # if self.data is None
self.data = data
# findval method to compare the value with nodes
# recursively traverse
def findval(self, lkpval):
if lkpval < self.data:
if self.left is None:
return str(lkpval)+" Not Found"
return self.left.findval(lkpval)
elif lkpval > self.data:
if self.right is None:
return str(lkpval)+" Not Found"
return self.right.findval(lkpval)
else:
print(str(self.data) + ' is found')
# Print the tree
# recursively traverse
def PrintTree(self):
if self.left:
self.left.PrintTree()
print(self.data)
if self.right:
self.right.PrintTree()
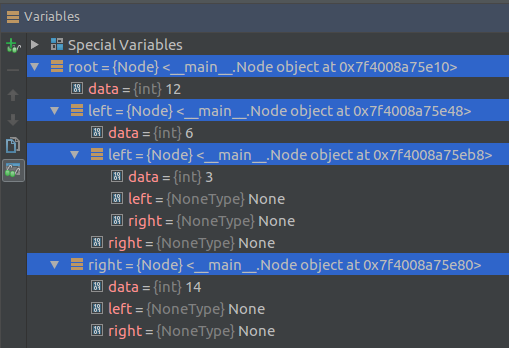
root = Node(12)
root.insert(6)
root.insert(14)
root.insert(3)
print(root.findval(7))
print(root.findval(14))
# -
# 
# [Python - Search Tree](https://www.tutorialspoint.com/python/python_binary_search_tree.htm)
# - [Searching a key](https://www.geeksforgeeks.org/binary-search-tree-set-1-search-and-insertion/)
#
# 1. Start from root.
# 2. Compare the inserting element with root, if less than root, then recurse for left, else recurse for right.
# 3. If element to search is found anywhere, return true, else return false.
#
# e.g.
# 
#
# ***
#
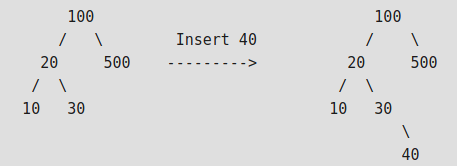
# - Insertion of a key
#
# 
#
# 1. Start from root.
# 2. Compare the inserting element with root, if less than root, then recurse for left, else recurse for right.
# 3. After reaching end,just insert that node at left(if less than current) else right.
#
# ***
#
# - Some Interesting Facts:
#
# 1. **Inorder traversal of BST always produces sorted output.**
# 2. We can construct a BST with only Preorder or Postorder or Level Order traversal. Note that we can always get inorder traversal by sorting the only given traversal.
# 3. Number of unique BSTs with n distinct keys is Catalan Number
#
# ***
#
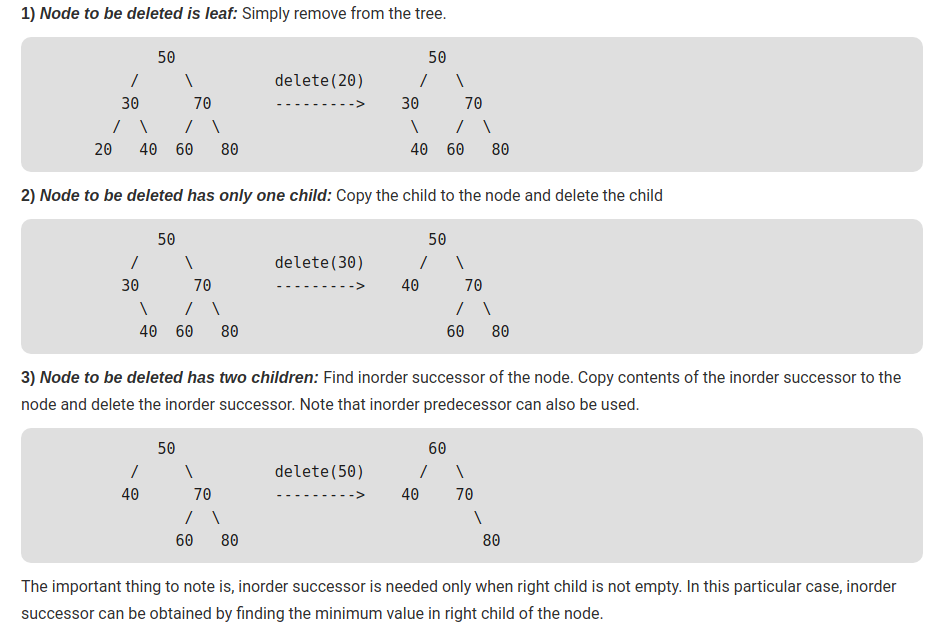
# - [Delete](https://www.geeksforgeeks.org/binary-search-tree-set-2-delete/)
#
# 
# +
# Python program to demonstrate delete operation
# in binary search tree
# A Binary Tree Node
class Node:
# Constructor to create a new node
def __init__(self, key):
self.key = key
self.left = None
self.right = None
# inorder traversal of BST
def inorder(root):
if root is not None:
inorder(root.left)
print(root.key)
inorder(root.right)
def insert(node, key):
# If the tree is empty, return a new node
if node is None:
return Node(key)
# Otherwise recur down the tree
if key < node.key:
node.left = insert(node.left, key)
else:
node.right = insert(node.right, key)
# return the (unchanged) node pointer
return node
# Given a non-empty binary search tree, return the node
# with minum key value found in that tree. Note that the
# entire tree does not need to be searched
def minValueNode(node): # it returns the address of node
current = node
# loop down to find the leftmost leaf
while (current.left is not None):
current = current.left
return current
# Given a binary search tree and a key, this function
# delete the key and returns the new root
def deleteNode(root, key):
# Base Case
if root is None:
return root
# If the key to be deleted is smaller than the root's
# key then it lies in left subtree
if key < root.key:
root.left = deleteNode(root.left, key)
# If the kye to be delete is greater than the root's key
# then it lies in right subtree
elif (key > root.key):
root.right = deleteNode(root.right, key)
# If key is same as root's key, then this is the node
# to be deleted
else:
# Node with only one child or no child
if root.left is None:
temp = root.right
root = None
return temp
elif root.right is None:
temp = root.left
root = None
return temp
# Node with two children: Get the inorder successor
# (smallest in the right subtree)
temp = minValueNode(root.right)
# Copy the inorder successor's content to this node
root.key = temp.key
# Delete the inorder successor
root.right = deleteNode(root.right, temp.key)
return root
# Driver program to test above functions
""" Let us create following BST
50
/ \
30 70
/ \ / \
20 40 60 80 """
root = None
root = insert(root, 50)
root = insert(root, 30)
root = insert(root, 20)
root = insert(root, 40)
root = insert(root, 70)
root = insert(root, 60)
root = insert(root, 80)
print("Inorder traversal of the given tree")
inorder(root)
print("\nMinVal is: ")
print(minValueNode(root))
print(minValueNode(root).key)
print("\nDelete 20")
root = deleteNode(root, 20)
print("Inorder traversal of the modified tree")
inorder(root)
print("\nDelete 30")
root = deleteNode(root, 30)
print("Inorder traversal of the modified tree")
inorder(root)
print("\nDelete 50")
root = deleteNode(root, 50)
print("Inorder traversal of the modified tree")
inorder(root)
| fundamentals/BST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 1. Setup
import sys
sys.path.append('../..')
# +
import config
import matplotlib.pyplot as plt
import numpy as np
import os
import warnings
from annotations import *
from density_maps import create_and_save_density_maps
from utils.data.data_ops import move_val_split_to_train
from utils.input_output.io import save_np_arrays, load_np_arrays, load_images
from utils.input_output.io import save_gt_counts, load_gt_counts
from utils.preprocessing.misc import gaussian_smoothing
# +
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
warnings.filterwarnings('ignore')
# -
# ## 2. Datasets
# ### 2.1 VGG Cells Dataset
# +
DATASET_PATH = '../../datasets/vgg_cells'
TRAIN_PATH = f'{DATASET_PATH}/train'
TRAIN_IMG_PATH = f'{TRAIN_PATH}/images'
TRAIN_GT_DOTS_PATH = f'{TRAIN_PATH}/gt_dots'
TRAIN_GT_COUNTS_PATH = f'{TRAIN_PATH}/gt_counts'
TRAIN_GT_DENSITY_MAPS_PATH = f'{TRAIN_PATH}/gt_density_maps'
VAL_PATH = f'{DATASET_PATH}/val'
TEST_PATH = f'{DATASET_PATH}/test'
TEST_IMG_PATH = f'{TEST_PATH}/images'
TEST_GT_DOTS_PATH = f'{TEST_PATH}/gt_dots'
TEST_GT_COUNTS_PATH = f'{TEST_PATH}/gt_counts'
TEST_GT_DENSITY_MAPS_PATH = f'{TEST_PATH}/gt_density_maps'
# -
move_val_split_to_train(VAL_PATH, TRAIN_PATH)
# +
# !rm -rf $TRAIN_GT_DENSITY_MAPS_PATH
# !rm -rf $TRAIN_GT_COUNTS_PATH
# !rm -rf $TEST_GT_DENSITY_MAPS_PATH
# !rm -rf $TEST_GT_COUNTS_PATH
# !mkdir $TRAIN_GT_DENSITY_MAPS_PATH
# !mkdir $TRAIN_GT_COUNTS_PATH
# !mkdir $TEST_GT_DENSITY_MAPS_PATH
# !mkdir $TEST_GT_COUNTS_PATH
# -
print(DATASET_PATH)
print(os.listdir(DATASET_PATH))
print(TRAIN_PATH)
print(os.listdir(TRAIN_PATH))
# +
train_img_names = sorted(os.listdir(TRAIN_IMG_PATH))
train_dots_names = sorted(os.listdir(TRAIN_GT_DOTS_PATH))
test_img_names = sorted(os.listdir(TEST_IMG_PATH))
test_dots_names = sorted(os.listdir(TEST_GT_DOTS_PATH))
print(f'train split: {len(train_img_names)} images')
print(train_img_names[:3])
print(train_dots_names[:3])
print(f'\ntest split: {len(test_img_names)} images')
print(test_img_names[:3])
print(test_dots_names[:3])
# +
train_dots_names = sorted(os.listdir(TRAIN_GT_DOTS_PATH))
test_dots_names = sorted(os.listdir(TEST_GT_DOTS_PATH))
print(TRAIN_GT_DOTS_PATH)
print(train_dots_names[:5])
print(TEST_GT_DOTS_PATH)
print(test_dots_names[:5])
# -
# #### Load dots images (.png)
# +
train_dots_images = load_dots_images(TRAIN_GT_DOTS_PATH, train_dots_names)
test_dots_images = load_dots_images(TEST_GT_DOTS_PATH, test_dots_names)
print(len(train_dots_images), train_dots_images[0].shape, train_dots_images[0].dtype,
train_dots_images[0].min(), train_dots_images[0].max(), train_dots_images[0].sum())
# -
# #### Save gt counts (from dots images)
train_counts = dots_images_to_counts(train_dots_images)
test_counts = dots_images_to_counts(test_dots_images)
save_gt_counts(train_counts, train_dots_names, TRAIN_GT_COUNTS_PATH)
save_gt_counts(test_counts, test_dots_names, TEST_GT_COUNTS_PATH)
# #### Create and save density maps (.npy)
create_and_save_density_maps(train_dots_images, config.VGG_CELLS_SIGMA,
train_img_names, TRAIN_GT_DENSITY_MAPS_PATH)
create_and_save_density_maps(test_dots_images, config.VGG_CELLS_SIGMA,
test_img_names, TEST_GT_DENSITY_MAPS_PATH)
# #### Load some train images and density maps
train_images = load_images(TRAIN_IMG_PATH, train_img_names, num_images=3)
print(len(train_images))
print(train_images[0].dtype)
train_gt_density_maps = load_np_arrays(TRAIN_GT_DENSITY_MAPS_PATH, num=3)
print(len(train_gt_density_maps))
print(train_gt_density_maps.dtype)
# +
NUM_PLOTS = 3
plt.figure(figsize=(15, 13))
for i in range(NUM_PLOTS):
count = train_dots_images[i].sum().astype(np.int)
plt.subplot(3, NUM_PLOTS, i + 1)
plt.title(f'Initial image (GT_dots count: {count})')
plt.imshow(train_images[i])
plt.subplot(3, NUM_PLOTS, NUM_PLOTS + i + 1)
plt.title(f'GT_dots count: {count}')
plt.imshow(train_dots_images[i], cmap='gray')
plt.subplot(3, NUM_PLOTS, 2 * NUM_PLOTS + i + 1)
plt.title(f'GT_dots: {count}, GT_density_map: {train_gt_density_maps[i].sum():.2f}')
plt.imshow(train_gt_density_maps[i], cmap='jet')
plt.colorbar()
| utils/data/density_map_generation_vgg_cells.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import torch
print(torch.__version__)
print(torch.cuda.is_available())
import torch
print(torch.cuda.current_device())
print(torch.cuda.device_count())
# + id="ULzsgA4bbtkr"
import os
import pandas as pd
import numpy as np #for transformation
import matplotlib.pyplot as plt #for plotting
from pathlib import Path
from sklearn.model_selection import train_test_split
import torch #pytorch package
import torch.nn as nn #basic building block for neural neteorks
import torch.nn.functional as F #import convolution functions like Relu
import torchvision
import torchvision.transforms as transforms #image transformation (cuz' the insufficiency of our data)
import torchvision.models as models #for finetune #commonly used model structures (including pre-trained models), such as AlexNet, VGG, ResNet, etc.
import torch.optim as optim #for optimize model(SGD, Adam, etc.)
#test
from skimage import io
from skimage import transform
import matplotlib.pyplot as plt
from torch.utils.data import Dataset, DataLoader
from torchvision.transforms import transforms
from torchvision.utils import make_grid
# -
# %pwd
# +
# 日常使用代码:将某个文件夹及其子目录下的所有图片格式改为.JPG格式
import os
import re
path='./imgs'
file_walk = os.walk(path)
fileNum = 0
filesPathList = []
for root, dirs, files in file_walk:
# print(root, end=',')
# print(dirs, end=',')
# print(files)
for file in files:
fileNum = fileNum + 1
filePath = root + '/' + file
#print(filePath)
filesPathList.append(filePath)
protion = os.path.splitext(filePath)
#print(protion[0],protion[1])
if protion[1].lower() != '.jpg':
#print("正在处理:" + filePath)
newFilePath = protion[0] + '.jpg'
os.rename(filePath, newFilePath)
print("這個dir已轉檔完成")
# + id="5ywYAy3_b0u2"
#training data
import glob
image= Path('imgs')
ext = ['png', 'jpg', 'gif','jfif','jpeg']# Add image formats here
file = []
[file.extend(image.glob(r'**/*.' + e)) for e in ext]
#file= list(image.glob('*.'+e) for e in ext)
labels = list(map(lambda x: os.path.split(os.path.split(x)[0])[1], file))
file= pd.Series(file, name='File').astype(str)
labels = pd.Series(labels, name='Label')
train_df= pd.concat([file, labels], axis=1)
train_df['Label'].value_counts()
# +
#確認資料寬高最大最小值
from os import listdir
from os.path import isfile, join
from PIL import Image
def print_data(data):
"""
Parameters
----------
data : dict
"""
print("Min width: %i" % data["min_width"])
print("Max width: %i" % data["max_width"])
print("Min height: %i" % data["min_height"])
print("Max height: %i" % data["max_height"])
def main(path):
"""
Parameters
----------
path : str
Path where to look for image files.
"""
onlyfiles = [f for f in listdir(path) if isfile(join(path, f))]
# Filter files by extension
onlyfiles = [f for f in onlyfiles if f.endswith(".jpg")]
data = {}
data["images_count"] = len(onlyfiles)
data["min_width"] = 10 ** 100 # No image will be bigger than that
data["max_width"] = 0
data["min_height"] = 10 ** 100 # No image will be bigger than that
data["max_height"] = 0
for filename in onlyfiles:
filename = str(path + '/'+filename)
im = Image.open(filename)
width, height = im.size
data["min_width"] = min(width, data["min_width"])
data["max_width"] = max(width, data["max_width"])
data["min_height"] = min(height, data["min_height"])
data["max_height"] = max(height, data["max_height"])
print_data(data)
if __name__ == "__main__":
main(path="marvel/train/black widow")
# +
#計算Normalization的mean跟sd
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import numpy as np
import torchvision
from torchvision import *
from torch.utils.data import Dataset, DataLoader
import matplotlib.pyplot as plt
import time
import copy
import os
from tqdm import tqdm
transform = transforms.Compose(
[ transforms.Resize([128, 128]),
transforms.ToTensor()
])
train_path = 'imgs/'
trainset = torchvision.datasets.ImageFolder(root=train_path, transform = transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=len(trainset))
####### COMPUTE MEAN / STD
# placeholders
psum = torch.tensor([0.0, 0.0, 0.0])
psum_sq = torch.tensor([0.0, 0.0, 0.0])
# loop through images
for inputs in tqdm(trainloader):
# print(inputs)
# inputs = np.asarray(inputs)
# inputs = np.squeeze(inputs)
# print(inputs.shape())
# psum += np.array(inputs[0]).sum(axis = [0, 2, 3])#B C H W 要求每C2的
inputs = inputs[0]
psum += inputs.sum(axis = (0, 2, 3))
psum_sq += (inputs ** 2).sum(axis = (0, 2, 3))
####### FINAL CALCULATIONS
# pixel count
image_size = 128
count = len(trainset) * image_size * image_size
# mean and std
total_mean = psum / count
total_var = (psum_sq/count) - (total_mean ** 2)
total_std = torch.sqrt(total_var)
# output
print('mean: ' + str(total_mean))
print('std: ' + str(total_std))
# +
# transform=transforms.Compose([
# # Random表示有可能做,所以也可能不做
# transforms.RandomHorizontalFlip(),# 水平翻转
# transforms.RandomVerticalFlip(), # 上下翻转
# transforms.RandomRotation(15), # 随机旋转-15°~15°
# transforms.RandomRotation([90, 180]), # 随机在90°、180°中选一个度数来旋转,如果想有一定概率不旋转,可以加一个0进去
# transforms.RandomCrop([28, 28]),
# ])
# +
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import numpy as np
import torchvision
from torchvision import *
from torch.utils.data import Dataset, DataLoader
import matplotlib.pyplot as plt
import time
import copy
import os
batch_size = 16
train_path = 'imgs/'
# mean: tensor([0.3850, 0.3505, 0.3395])
# std: tensor([0.3048, 0.2915, 0.2879])
transforms = transforms.Compose(
[
transforms.Resize([128,128]),
transforms.ToTensor(),
transforms.Normalize([0.3850, 0.3505, 0.3395],[0.3048, 0.2915, 0.2879])
])
allset = datasets.ImageFolder(root=train_path, transform=transforms)
print(allset.class_to_idx)
total_count = len(allset)
print(f"total count : {total_count}")
train_count = int(0.7 * total_count)
valid_count = int(0.2 * total_count)
test_count = total_count - train_count - valid_count
trainset ,validset ,testset = torch.utils.data.random_split(allset, (train_count, valid_count, test_count))
print('train num',train_count)
print('valid num',valid_count)
print('test num',test_count)
trainloader = DataLoader(trainset, batch_size=16, shuffle=True)
validloader = DataLoader(validset, batch_size=16, shuffle=True)
# test_dataloader = DataLoader(test_dataset, batch_size=batch_size, shuffle=True)
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
def imshow(inp, title=None):
inp = inp.cpu() if device else inp
inp = inp.numpy().transpose((1, 2, 0))
# mean = np.array([0.485, 0.456, 0.406])
# std = np.array([0.229, 0.224, 0.225])
# inp = std * inp + mean
# inp = np.clip(inp, 0, 1)
plt.figure(figsize=(128,512))
plt.subplot(3,3,1)
plt.imshow(inp)
plt.pause(0.001)
images, labels = next(iter(trainloader))
print("images-size:", images.shape)
out = torchvision.utils.make_grid(images)
print("out-size:", out.shape)
# imshow(out)
# + id="R7b_qYnKb7ut"
net = models.resnet34(pretrained=True)
net = nn.DataParallel(net)
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(net.parameters(), lr=0.0001, momentum=0.9,weight_decay=0.01)
#optimizer = optim.Adam(net.parameters(),lr=0.0001,weight_decay=0.1)
#exp_lr_scheduler = lr_scheduler.StepLR(optimizer, step_size=1, gamma=0.1) #lr 隨迭代改變
net.module.fc = torch.nn.Sequential(
torch.nn.Linear(in_features=512, out_features=9, bias=True),
#torch.nn.ReLU(inplace=True),
#torch.nn.Linear(in_features=128, out_features=8, bias=True)
)
net = net.cuda() if device else net
# + id="RfoYgJPCb-Ih"
# %%time
early_stop = 0
n_epochs = 50
valid_loss_min = np.Inf
val_loss = []
val_acc = []
train_loss = []
train_acc = []
total_step = len(trainloader)
valid_result = pd.DataFrame(columns=['pred', 'target'])
for epoch in range(1, n_epochs+1):
running_loss = 0.0
correct = 0
total=0
print(f'Epoch {epoch}\n')
for batch_idx, (data_, target_) in enumerate(trainloader):
data_, target_ = data_.to(device), target_.to(device)
outputs = net(data_)
optimizer.zero_grad()
loss = criterion(outputs, target_)
loss.backward()
optimizer.step()
running_loss += loss.item()
_,pred = torch.max(outputs, dim=1)
correct += torch.sum(pred==target_).item()
total += target_.size(0)
if (batch_idx) % 16 == 0:
print ('Epoch [{}/{}], Step [{}/{}], Loss: {:.4f}'
.format(epoch, n_epochs, batch_idx, total_step, loss.item()))
train_acc.append(100 * correct / total)
train_loss.append(running_loss/total_step)
print(f'\ntrain-loss: {np.mean(train_loss):.4f}, train-acc: {(100 * correct/total):.4f}')
batch_loss = 0
total_t=0
correct_t=0
with torch.no_grad():
net.eval()
for data_t, target_t in (validloader):
data_t, target_t = data_t.to(device), target_t.to(device)
outputs_t = net(data_t)
loss_t = criterion(outputs_t, target_t)
batch_loss += loss_t.item()
_,pred_t = torch.max(outputs_t, dim=1)
pred_t = pred_t.cpu()
target_t = target_t.cpu()
valid_result = valid_result.append({'pred':pred_t ,'target':target_t },ignore_index=True)
correct_t += torch.sum(pred_t==target_t).item()
total_t += target_t.size(0)
val_acc.append(100 * correct_t/total_t)
val_loss.append(batch_loss/len(testloader))
print(f'validation loss: {np.mean(val_loss):.4f}, validation acc: {(100 * correct_t/total_t):.4f}\n')
if batch_loss < valid_loss_min:
valid_loss_min = batch_loss
torch.save(net.state_dict(), 'resnet.pt')
print('Improvement-Detected, save-model')
early_stop = 0
else :
early_stop += 1
if early_stop == 5 :
print("Early Stop!")
break
#exp_lr_scheduler.step()#改變學習率
net.train()
print("Train End~")
valid_result.to_csv('分類結果.csv',index=False)
# + id="ytN5OtZVcAM1"
fig = plt.figure(figsize=(20,10))
plt.title("Train-Validation Accuracy")
plt.plot(train_acc, label='train')
plt.plot(val_acc, label='validation')
plt.xlabel('num_epochs', fontsize=12)
plt.ylabel('accuracy', fontsize=12)
plt.legend(loc='best')
# + id="68wVulTIcDOd"
PATH = './resnet.pt'
# -
# load back in our saved model
net.load_state_dict(torch.load(PATH))
#testloader = DataLoader(testset, batch_size=16, shuffle=True)
new_testset = torch.utils.data.ConcatDataset([testset, validset])
len(new_testset)
len(validset)
len(testset)
testloader = DataLoader(new_testset, batch_size=16, shuffle=True)
# + id="RWgMUj9hcTbS"
# %%time
correct = 0
total = 0
# since we're not training, we don't need to calculate the gradients for our outputs
with torch.no_grad():
for data in testloader:
images, labels = data
images, labels = images.to(device), labels.to(device)
# calculate outputs by running images through the network
outputs = net(images)
# the class with the highest energy is what we choose as prediction
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 874 test images: %d %%' % (
100 * correct / total))
# + id="GLyN6FBZcVse"
# prepare to count predictions for each class
correct_pred = {classname: 0 for classname in classes}
total_pred = {classname: 0 for classname in classes}
# again no gradients needed
with torch.no_grad():
for data in testloader:
images, labels = data
images, labels = images.to(device), labels.to(device)
outputs = net(images)
_, predictions = torch.max(outputs, 1)
# collect the correct predictions for each class
for label, prediction in zip(labels, predictions):
if label == prediction:
correct_pred[classes[label]] += 1
total_pred[classes[label]] += 1
# print accuracy for each class
for classname, correct_count in correct_pred.items():
accuracy = 100 * float(correct_count) / total_pred[classname]
print("The num of the class is {},Accuracy for class {:5s} is: {:.1f} %".format(total_pred[classname],classname,
accuracy))
# -
| AI_final_project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from osgeo import gdal, osr
gdal.PushErrorHandler('CPLQuietErrorHandler')
gdal.UseExceptions()
import sys, os, tqdm
from multiprocessing import Pool
# -
file_Input1 ='Mandiri/s1_konawe_selatan.tif'
file_Input2 ='Mandiri/s2_l8_konawe_selatan.tif'
file_location ='Mandiri/regular_points_fin.csv'
# +
print("Input Image 1 :", file_Input1)
print("Input Image 2 :", file_Input2)
##File Input 1##
ds2 = gdal.Open(file_Input1)
ulx, xres, xskew, uly, yskew, yres = ds2.GetGeoTransform()
lrx = ulx + (ds2.RasterXSize * xres)
lry = uly + (ds2.RasterYSize * yres)
gt = ds2.GetGeoTransform()
rows = ds2.RasterYSize
cols = ds2.RasterXSize
jbands = ds2.RasterCount
bands = jbands
proj_ref = ds2.GetProjectionRef()
training_features = np.zeros((rows*cols,bands),dtype=np.uint16)
for band in range(bands):
training_features[:,band]= (ds2.GetRasterBand(jbands-bands+band+1).ReadAsArray().astype(np.uint16)).flatten()
# -
data_locations = pd.read_csv(file_location)
#print (data_locations)
ndata = len(data_locations)
print ("NData:", ndata);
print ("ulx:", ulx, xres)
print ("uly:", uly, yres)
#data = data_locations.iloc[1,:]
print ("Rows:",rows)
print ("Cols:",cols)
ds2 = None
DNArray = np.zeros((ndata,bands+4),dtype=np.float64)
n=0
for i in range(ndata):
id = data_locations.iloc[i,0]
idcls = data_locations.iloc[i,2]
lon = data_locations.iloc[i,3]
lat = data_locations.iloc[i,4]
x= (lon-ulx)/xres
y= (lat-uly)/yres
xx = int(x)
yy = int(y)
#print (lon, lat, clsnm, xx,yy)
#DNArray[i,4] = clsnm
xy=0
if(xx>-1)and(xx<cols):
if(yy>-1)and(yy<rows):
xy = (yy*cols)+xx
DNArray[n,0] = id
DNArray[n,1] = idcls
DNArray[n,2] = lon
DNArray[n,3] = lat
DNArray[n,4:] = training_features[xy,:]
n=n+1
#print (DNArray[i,:])
#
DNArrayFinal = np.zeros((n,bands+4),dtype=np.float64)
DNArrayFinal[0:n,:] = DNArray[0:n,:]
#
data_1=pd.DataFrame(DNArrayFinal,columns=['id','Class_ID','xcoord','ycoord','S1_VVmin','S1_VHmin','S1_VVmed','S1_VHmed','S1_VVmax','S1_VHmax'])
##File Input 2#
ds2 = gdal.Open(file_Input2)
ulx, xres, xskew, uly, yskew, yres = ds2.GetGeoTransform()
lrx = ulx + (ds2.RasterXSize * xres)
lry = uly + (ds2.RasterYSize * yres)
gt = ds2.GetGeoTransform()
rows = ds2.RasterYSize
cols = ds2.RasterXSize
jbands = ds2.RasterCount
bands = jbands
proj_ref = ds2.GetProjectionRef()
training_features = np.zeros((rows*cols,bands),dtype=np.uint16)
for band in range(bands):
training_features[:,band]= (ds2.GetRasterBand(jbands-bands+band+1).ReadAsArray().astype(np.uint16)).flatten()
ds2 = None
DNArray = np.zeros((ndata,bands+4),dtype=np.float64)
n=0
for i in range(ndata):
id = data_locations.iloc[i,0]
idcls = data_locations.iloc[i,2]
lon = data_locations.iloc[i,3]
lat = data_locations.iloc[i,4]
x= (lon-ulx)/xres
y= (lat-uly)/yres
xx = int(x)
yy = int(y)
#print (lon, lat, clsnm, xx,yy)
#DNArray[i,4] = clsnm
xy=0
if(xx>-1)and(xx<cols):
if(yy>-1)and(yy<rows):
xy = (yy*cols)+xx
DNArray[n,0] = id
DNArray[n,1] = idcls
DNArray[n,2] = lon
DNArray[n,3] = lat
DNArray[n,4:] = training_features[xy,:]
n=n+1
#print (DNArray[i,:])
#
DNArrayFinal = np.zeros((n,bands+4),dtype=np.float64)
DNArrayFinal[0:n,:] = DNArray[0:n,:]
#
str_1='l8'
l8_=[]
for i in ['B2','B3','B4','B5','B6']:
for y in ['min','med','max','mean','stddev']:
l8_.append(str_1+'_'+i+y)
str_1='s2'
s2_=[]
for i in ['B2','B3','B4','B8','B11']:
for y in ['min','med','max','mean','stddev']:
s2_.append(str_1+'_'+i+y)
col_gab=['id','Class_ID','xcoord','ycoord']+l8_+s2_
data_2=pd.DataFrame(DNArrayFinal,columns=col_gab)[l8_+s2_]
gab_data_=data_1.merge(data_2,left_index=True,right_index=True)
gab_data_.to_csv('Mandiri/data_latihan_train.csv',sep=',',index=False)
| 03_WorkshopRS w Kustiyo Lapan/01_Preparing_data_and_bands.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pytorch Basics - Regressão Linear
# > Tutorial de como realizar um modelo de regressão linear no Pytorch.
#
# - toc: false
# - badges: true
# - comments: true
# - categories: [pytorch, regressaolinear]
# - image: images/pytorch.png
# O objetivo desse breve trabalho é apresentar como é realizado um modelo de regressão linear utilizando pytorch. Muitas das vezes utiliza-se regressão linear como uma primeira hipotese, devido a sua simplicidade, antes de partir para modelos mais complexos.
# ## Carregando as bibliotecas necessárias
# +
#Carregando o Pytorch
import torch
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import numpy as np
# -
# ## Carregando o conjunto de dados
# Para carregar o bando de dados que está em .csv, utilizamos o pandas, o qual consegue ler um arquivo localmente ou em um nuvem (url deve ser do raw do .csv)
df = pd.read_csv('https://raw.githubusercontent.com/lucastiagooliveira/lucas_repo/master/Kaggle/Revisiting%20a%20Concrete%20Strength%20regression/datasets_31874_41246_Concrete_Data_Yeh.csv')
# Mostrando as 5 primeiras linhas do dataframe carregado, isso é importante para verificarmos o se o dataframe está correto.
df.head()
# Apresentando um resumo estatístico dos dataframe por coluna, tais como: quantidade de dados, média, desvio padrão, mínimo, primeiro ao terceiro quartil e valor máximo.
df.describe()
# ## Plotando os gráficos de todas as váriaveis
# Para visualização da relação entre as váriaveis é interessante fazer a visualização gráfica da relação entre as variáveis. Para isso usamos a função PairGrid da biblioteca Seaborn aliado com um scatterplot da biblioteca MatplotLib.
sns.set(style="darkgrid")
g = sns.PairGrid(df)
g.map(plt.scatter)
# ## Correlação linear
# Para entendimento da correlação linear das variáveis entre si, temos a função "built-in" do Pandas que nos retorna o coeficiente de correlação que tem por padrão o método Pearson.
df.corr()
# Escolhendo as variáveis que serão utilizadas para criação do modelo.
var_used = ['cement', 'superplasticizer', 'age', 'water']
train = df[var_used]
target = df['csMPa']
# Tabela com somente as variáveis que serão utilizadas.
train.head()
# Para iniciarmos um modelo temos que fazer a transformação da base de dados que está com o tipo de DataFrame para tensor, que é utilizado pelo Pytorch. Todavia, uma das maneiras de fazer essa transformação é antes fazer a transformação da base de dados para um vetor do Numpy e depois transformar para um tensor do Pytorch.
#
# Obs.: Foi criado o vetor de uns para ser adicionado ao tensor dos parâmetros, pois essa coluna deverá multiplicar a constante da expressão (b), conforme o exemplo abaixo.
#
# Y = a*X + b
train = np.asarray(train)
a = np.ones((train.shape[0],1))
train = torch.tensor(np.concatenate((train, a), axis=1))
target = torch.tensor(np.asarray(target))
train.shape
# ## Criando o modelo
# Para iniciarmos precisamos criar uma função a qual definirá a equação da regressão linear a qual utilizará a função matmul para realizar a multiplicação entre os dois tensores dos parâmetros e variáveis dependentes.
def model(x,params):
return torch.matmul(x, params)
# Função que calcula o erro quadrático médio (MSE).
#
# Para saber mais sobre como é calculado acesso o link: https://pt.qwe.wiki/wiki/Mean_squared_error
def mse(pred, labels): return ((pred - labels)**2).mean()
# Para iniciar o treino do modelo primeiramente temos que criar um tensor o qual receberá os valores dos parâmetros que serão atualizados a cada iteração, quedo assim precisamos utilizar o método requires_grad_ assim será possível calcular o gradiente desse tensor quando necessário.
#
# Observe que o tipo do objeto criado é torch.float64.
params = torch.randn(5,1, dtype=torch.float64).requires_grad_()
params.dtype
# **Primeiro passo:** realizar as predições do modelo
pred = model(train, params)
# **Segundo passo:** calcular como o nosso modelo performou, ou seja, calcular MSE para averiguação da performace do modelo.
#
# Observe que o modelo vai apresentar um erro acentuado, pois os parâmetros ainda não foram *treinados*.
loss = mse(pred, target)
loss
# **Terceiro passo:** realizar o gradiente descente.
#
# Conceito do algoritmo de gradiente descendente: http://cursos.leg.ufpr.br/ML4all/apoio/Gradiente.html
loss.backward()
params.grad
# **Quarto passo:** Atualização dos parâmetros, para isso utiliza-se o valor do gradiente por meio do algoritmo descendente e é escalado (multiplicado) pelo taxa de aprendizado (*learning rate*).
#
# Após a realização da atulização dos parâmetros deve-se resetar o gradiente.
lr = 1e-5
params.data -= lr * params.grad.data
params.grad = None
# Primeira iteração realizada, pode-se observar o valor do erro do nosso modelo reduziu. A tendência é ocorrer uma diminuição até a cada iteração, até a estabilização do modelo.
pred = model(train, params)
loss = mse(pred, target)
loss
# Foi criada uma função que realiza todos os passos acima realizados.
def step(train, target, params, lr = 1e-6):
## realizando as predições
pred = model(train, params)
## caculando o erro
loss = mse(pred, target)
## realizando o gradiente descendente
loss.backward()
## atualizando os parâmtros
params.data -= lr * params.grad.data
## reset do gradiente
params.grad = None
## imprimindo na tela o erro
print('Loss:',loss.item())
## retornado as predições e os parâmetros atuzalizados na ultima iteração
return pred, params
# Criando um loop para realizar as itereções, é possível verificar a diminuição do erro a cada iteração, ou seja, se realizada mais iteração pode-se chegar a um resultado plausível (neste caso não cheramos a um, pois o modelo de regressão linear não é um modelo adequado para esses dados, somente como hipótese inicial).
for i in range(10): loss, params = step(train, target, params)
# Esté é o resultado dos parâmetros que serão utilizados para o modelo realizar futuras predições.
parameters = params
parameters #parametros do modelo
| _notebooks/2020-08-31-pytorch_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Q learning
# ## Brief
# Suppose we have 5 rooms in a building connected by doors as shown in the figure below. We'll number each room 0 through 4. The outside of the building can be thought of as one big room (5). Notice that doors 1 and 4 lead into the building from room 5 (outside). For this example, we'd like to put an agent in any room, and from that room, go outside the building (this will be our target room). In other words, the goal room is number 5.
# ### Map
# 
# ### Graph
# 
# ### Reference
# [Reference](http://mnemstudio.org/path-finding-q-learning-tutorial.htm)
# ### Key algorithm
# $$\Delta Q(s,a)=R(s,a)+\gamma\cdot\max_{\tilde{a}}{Q(\tilde{s},\tilde{a})}$$
#
# $$Q(s,a) = (1-\alpha)Q(s,a)+\alpha\Delta Q(s,a)$$
# ## Import
import numpy as np
import random
# ## Initialize
Q=np.zeros([6,6])
R=np.array([[-1,-1,-1,-1,0,-1],
[-1,-1,-1,0,-1,100],
[-1,-1,-1,0,-1,-1],
[-1,0,0,-1,0,-1],
[0,-1,-1,0,-1,100],
[-1,0,-1,-1,0,100]])
print("Q:\n{}\nR:\n{}".format(Q,R))
def train(R,targetState,Q=None,n_episode=20,learningRate=0.2,gamma=0.8,printInterval=None,verbose=False):
rShape=np.shape(R)
if rShape[0]!=rShape[1]:
raise ValueError("The number of columns and rows in R didn't match. ")
if Q==None:
Q=np.zeros([rShape[0],rShape[1]])
else:
qShape=np.shape(Q)
if qShape[0]!=rShape[0] or qShape[1]!=rShape[1]:
raise ValueError("Size of Q and R didn't match. ")
for episode in range(n_episode):
state=random.randint(0,rShape[0]-1)
step=0
while state!=targetState:
#Q(S,A) ← (1-α)*Q(S,A) + α*[R + γ*maxQ(S',a)]
step+=1
actionSet=np.argwhere(R[state]>=0)
action=actionSet[random.randint(0,np.shape(actionSet)[0]-1),0]
if verbose and printInterval!=None and (episode+1)%printInterval==0:
print("At episode {}\nState: {}\nAction: {}\nQ:\n{}".format(episode+1,state,action,Q))
Q[state,action]=(1-learningRate)*Q[state,action]+learningRate*(R[state,action]+
gamma*Q[action,np.argmax(Q[action])])
state=action
if printInterval!=None and (episode+1)%printInterval==0:
print("At episode {}\nQ:\n{}".format(episode+1,Q))
return Q
# ## Training #1
#Q(state, action) = R(state, action) + Gamma * Max[Q(next state, all actions)]
Q=train(R,targetState=5,n_episode=20,learningRate=1,gamma=0.8,printInterval=1,verbose=True)
print(Q)
# ## Online Inference
for state in range(5):
print("Initial state:{}".format(state))
while state!=5:
#Didn't bother to validate the action
actionSet=np.argwhere(Q[state]==np.max(Q[state]))
action=actionSet[random.randint(0,np.shape(actionSet)[0]-1),0]
print("Moving from room {} to room {}.".format(state,action))
state=action
# ## Training #2
#Q(S,A) ← (1-α)*Q(S,A) + α*[R + γ*maxQ(S',a)]
Q=train(R,n_episode=200,learningRate=0.3,gamma=0.85,targetState=5,printInterval=1,verbose=False)
# ## Online inference
for state in range(5):
print("Initial state:{}".format(state))
while state!=5:
#Didn't bother to validate the action
actionSet=np.argwhere(Q[state]==np.max(Q[state]))
action=actionSet[random.randint(0,np.shape(actionSet)[0]-1),0]
print("Moving from room {} to room {}.".format(state,action))
state=action
| Q-learning/Q-learning Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# language: python
# name: python3
# ---
# +
import pickle
import imageio
import sys
import pdb
import os
import cv2
import numpy as np
import trimesh
pickle_name = '15a3f5f564c538e2baa1282ba5a20d4c'
# person1: <PASSWORD>
# person2: <PASSWORD>
# person3: <PASSWORD>
OBJNAME = 'person1'
os.makedirs(f'output/{OBJNAME}/LASR/FlowFW/Full-Resolution/{OBJNAME}',exist_ok=True)
os.makedirs(f'output/{OBJNAME}/LASR/FlowBW/Full-Resolution/{OBJNAME}',exist_ok=True)
os.makedirs(f'output/{OBJNAME}/LASR/Annotations/Full-Resolution/{OBJNAME}',exist_ok=True)
os.makedirs(f'output/{OBJNAME}/LASR/JPEGImages/Full-Resolution/{OBJNAME}',exist_ok=True)
os.makedirs(f'output/{OBJNAME}/LASR/Meshes/Full-Resolution/{OBJNAME}',exist_ok=True)
# os.makedirs(f'output/{OBJNAME}/LASR/FlowFW/Full-Resolution/r{OBJNAME}',exist_ok=True)
# os.makedirs(f'output/{OBJNAME}/LASR/FlowBW/Full-Resolution/r{OBJNAME}',exist_ok=True)
with open(f'./MeshSeqs/{pickle_name}.pkl', 'rb') as file:
frames_dict = pickle.load(file)
for i in range(len(frames_dict)):
imageio.imsave(f'output/{OBJNAME}/LASR/JPEGImages/Full-Resolution/{OBJNAME}/{i:05d}.png',frames_dict[i]['rgb_frame'][...,:-1])
# miss mask
imageio.imsave(f'output/{OBJNAME}/LASR/Annotations/Full-Resolution/{OBJNAME}/{i:05d}.png',frames_dict[i]['rgb_frame'][...,-1] // 2)
# convert connectivity to faces
# faces = []
# for arr in map(lambda x:np.unique(np.concatenate(np.asarray(x))),frames_dict[i]['mesh_connectivity_graph'][0]):
# if len(arr) != 3:
# tri = trimesh.geometry.triangulate_quads([arr])
# faces.append(tri)
# # pdb.set_trace()
# else:
# faces.append([arr])
# faces = np.concatenate(faces)
connectivity_to_vertices = lambda x:np.unique(np.concatenate(np.asarray(x)))
quads = list(map(connectivity_to_vertices, frames_dict[i]['mesh_connectivity_graph'][0]))
faces = trimesh.geometry.triangulate_quads(quads)
mesh = trimesh.Trimesh(vertices=frames_dict[i]['mesh_coords_3d'][0,...], faces=faces)
trimesh.exchange.export.export_mesh(mesh,f'output/{OBJNAME}/LASR/Meshes/Full-Resolution/{OBJNAME}/{i:05d}.obj')
# save gif
imageio.mimsave(f'output/{OBJNAME}/{pickle_name}.gif',[frames_dict[i]['rgb_frame'][...,:-1] for i in range(len(frames_dict))])
fw = np.array([frames_dict[i]['forward_optical_flow'] for i in range(len(frames_dict))])[:-1,...]
bw = np.array([frames_dict[i]['backward_optical_flow'] for i in range(len(frames_dict))])[1:,...]
# write optical flow and occlusion map in LASR format
def write_pfm(path, image, scale=1):
"""Write pfm file.
Args:
path (str): pathto file
image (array): data
scale (int, optional): Scale. Defaults to 1.
"""
with open(path, "wb") as file:
color = None
if image.dtype.name != "float32":
raise Exception("Image dtype must be float32.")
image = np.flipud(image)
if len(image.shape) == 3 and image.shape[2] == 3: # color image
color = True
elif (
len(image.shape) == 2 or len(image.shape) == 3 and image.shape[2] == 1
): # greyscale
color = False
else:
raise Exception("Image must have H x W x 3, H x W x 1 or H x W dimensions.")
file.write("PF\n".encode() if color else "Pf\n".encode())
file.write("%d %d\n".encode() % (image.shape[1], image.shape[0]))
endian = image.dtype.byteorder
if endian == "<" or endian == "=" and sys.byteorder == "little":
scale = -scale
file.write("%f\n".encode() % scale)
image.tofile(file)
if 'no-flo' not in OBJNAME:
for i in range(len(fw)):
f = fw[i,...]
ones = np.ones_like(f[...,:1])
f = np.concatenate([f[...,1:], f[...,:1], ones],-1)
b = np.concatenate([-bw[i,...,1:],-bw[i,...,:1], ones],-1)
f = np.flip(f,0)
b = np.flip(b,0)
write_pfm(f'output/{OBJNAME}/LASR/FlowFW/Full-Resolution/{OBJNAME}/flo-{i:05d}.pfm',f)
write_pfm(f'output/{OBJNAME}/LASR/FlowBW/Full-Resolution/{OBJNAME}/flo-{i+1:05d}.pfm',b)
write_pfm(f'output/{OBJNAME}/LASR/FlowFW/Full-Resolution/{OBJNAME}/occ-{i:05d}.pfm',np.ones_like(f[...,0]))
write_pfm(f'output/{OBJNAME}/LASR/FlowBW/Full-Resolution/{OBJNAME}/occ-{i+1:05d}.pfm',np.ones_like(b[...,0]))
# -
for k in frames_dict[0]:
print(k)
# +
faces = []
for arr in map(lambda x:np.unique(np.concatenate(np.asarray(x))),frames_dict[0]['mesh_connectivity_graph'][0]):
if len(arr) != 3:
tri = trimesh.geometry.triangulate_quads([arr])
faces.append(tri)
# pdb.set_trace()
else:
faces.append([arr])
faces = np.concatenate(faces)
# -
trimesh.geometry.triangulate_quads(list(map(lambda x:np.unique(np.concatenate(np.asarray(x))),frames_dict[0]['mesh_connectivity_graph'][0])))
faces.shape
frames_dict[0]['num_vertices']
# +
import pickle
import imageio
import sys
import pdb
import os
import cv2
import numpy as np
import trimesh
import glob
os.makedirs(f'./MeshSeqs/preview/',exist_ok=True)
for filename in list(glob.glob('./MeshSeqs/*.pkl')):
with open(filename, 'rb') as f:
frames_dict = pickle.load(f)
pickle_name = filename.split('/')[-1]
imageio.mimsave(f'./MeshSeqs/preview/{pickle_name}.gif',[frames_dict[i]['rgb_frame'][...,:-1] for i in range(len(frames_dict))])
# -
filename.split('/')
mesh
cv2.imread('output/airplane/LASR/Annotations/Full-Resolution/airplane/00000.png').max()
# cv2.imread(f'output/{OBJNAME}/LASR/JPEGImages/Full-Resolution/{OBJNAME}/{i:05d}.png')
| mesh_sequence_checker.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%% Grabbing Excess Items (1)\n"}
a, b, *rest = range(5)
print(a)
print(b)
print(rest)
# + pycharm={"name": "#%% Grabbing Excess Items (2)\n"}
a, *middle, b = range(5)
print(a)
print(b)
print(middle)
# + pycharm={"name": "#%% Nested Unpacking\n"}
persons = [
('Sam', 42, 1.87, ('Hansbeat', 'Madeleine')),
('Shirley', 43, 1.61, ('John', 'Maya')),
]
fmt = '{:10} | {:2d} | {:1.3f} | {:10} | {:10}'
for person, age, height, (father, mother) in persons:
print(fmt.format(person, age, height, father, mother))
# ------------- Hint -----------------------
# For naming columns see namedtuple factory.
# ------------------------------------------
# + pycharm={"name": "#%% Splitting\n"}
my_list = [1, 2, 3, 4]
print(my_list[:2])
print(my_list[2:])
# + pycharm={"name": "#%% Replace parts of a list\n"}
my_list = list(range(0,10))
print(my_list)
my_list[8:10] = ['a', 'b', 'c']
print(my_list)
del my_list[0:2]
print(my_list)
my_list[1::2] = [42, 43, 44, 45]
print(my_list)
# + pycharm={"name": "#%% Tik, tak, toe right an wrong\n"}
# The right way:
tik_tak_toe = [['_'] * 3 for i in range(3)]
tik_tak_toe[0][0] = 'X'
print(tik_tak_toe)
"""
The wrong way: Three references to the same list.
"""
tik_tak_toe = [['_'] * 3] * 3
test = [['_'] * 3] * 3
tik_tak_toe[0][0] = 'X'
print(tik_tak_toe)
# + pycharm={"name": "#%% sort and sorted\n"}
"""
sort sorts the list returns 'None'. sorted returns a sorted list.
"""
names = ['Peter', 'Hans', 'Gustav', 'Adelheid', '<NAME>']
print(sorted(names))
print(sorted(names, reverse=True))
print(sorted(names, key=len))
print(names)
names.sort()
print(names)
# + pycharm={"name": "#%% Return position as search result of sorted sequence with bisect\n"}
"""
There is a bisect_right and bisect_left. bisect is an alias of
bisect_right.
"""
import bisect
to_be_searched = ['Adam', 'Carl', 'Daniel']
position = bisect.bisect(to_be_searched, 'Werner')
print(position)
# + pycharm={"name": "#%% Insert a element at the right place\n"}
my_list = ['Adam', 'Carl', 'Daniel']
bisect.insort(my_list, "Werner")
print(my_list)
bisect.insort(my_list, "Abraham")
print(my_list)
| notebooks/fp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
"""
基于滑动窗口的欧拉反卷积算法
通过基于滑动窗的欧拉反卷积方法估计简单异常模型的坐标。
该方法的使用条件是源要具有简单的几何形状,如球体, 垂直柱体,垂直板状体等。当异常源很复杂的时候,该方法效果不是很理想。
该程序通过常见的滑动窗口方案来测试欧拉反卷积
方案实现于:geoist.euler_moving_window.ipynb。
"""
from geoist.pfm import sphere, pftrans, euler, giutils
from geoist import gridder
from geoist.inversion import geometry
from geoist.vis import giplt
import matplotlib.pyplot as plt
# +
##合成磁数据测试欧拉反卷积
# 磁倾角,磁偏角
inc, dec = -45, 0
# 制作仅包含感应磁化的两个球体模型
model = [
geometry.Sphere(x=-1000, y=-1000, z=1500, radius=1000,
props={'magnetization': giutils.ang2vec(2, inc, dec)}),
geometry.Sphere(x=1000, y=1500, z=1000, radius=1000,
props={'magnetization': giutils.ang2vec(1, inc, dec)})]
print("Centers of the model spheres:")
print(model[0].center)
print(model[1].center)
# 从模型中生成磁数据
shape = (100, 100)
area = [-5000, 5000, -5000, 5000]
x, y, z = gridder.regular(area, shape, z=-150)
data = sphere.tf(x, y, z, model, inc, dec)
# 一阶导数
xderiv = pftrans.derivx(x, y, data, shape)
yderiv = pftrans.derivy(x, y, data, shape)
zderiv = pftrans.derivz(x, y, data, shape)
# +
#通过扩展窗方法实现欧拉反卷积
#给出2个解决方案,每一个扩展窗都靠近异常
#stutural_index=3表明异常源为球体
'''
===================================== ======== =========
源类型 SI (磁) SI (重力)
===================================== ======== =========
Point, sphere 3 2
Line, cylinder, thin bed fault 2 1
Thin sheet edge, thin sill, thin dyke 1 0
===================================== ======== =========
'''
#选择10 x 10个大小为1000 x 1000 m的窗口
solver = euler.EulerDeconvMW(x, y, z, data, xderiv, yderiv, zderiv,
structural_index=3, windows=(10, 10),
size=(1000, 1000))
#使用fit()函数来获取右下角异常的估计值
solver.fit()
#估计位置存储格式为一列[x, y, z] 坐标
print('Kept Euler solutions after the moving window scheme:')
print(solver.estimate_)
# 在磁数据上绘制异常估计值结果
# 异常源的中心的真正深度为1500 m 和1000 m。
plt.figure(figsize=(6, 5))
plt.title('Euler deconvolution with a moving window')
plt.contourf(y.reshape(shape), x.reshape(shape), data.reshape(shape), 30,
cmap="RdBu_r")
plt.scatter(solver.estimate_[:, 1], solver.estimate_[:, 0],
s=50, c=solver.estimate_[:, 2], cmap='cubehelix')
plt.colorbar(pad=0).set_label('Depth (m)')
plt.xlim(area[2:])
plt.ylim(area[:2])
plt.tight_layout()
plt.show()
| notebooks/euler_moving_window.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np, tensorflow as tf
from glob import glob
from analysis import plot_deep_features, plot, smooth_ranges_2d, text
from make_frames import make_frames
import os
import h5py
from pprint import pprint
from tqdm import tqdm
from IPython import display
import matplotlib.pyplot as plt
import matplotlib.patches as patches
# %matplotlib inline
# + language="sh"
# cd data/
# ls -l
# + language="sh"
# cd figures/animation/
# ls -l
# -
configs = [
{
'path': 'data/pre_train_lambda_0.01.h5',
'prefix': 'pre_train_lambda_0.01/frame'
},
{
'path': 'data/lambda_3.h5',
'prefix': 'lambda_3/frame'
},
{
'path': 'data/lambda_1.h5',
'prefix': 'lambda_1/frame'
},
{
'path': 'data/lambda_0.1.h5',
'prefix': 'lambda_0.1/frame'
},
{
'path': 'data/lambda_0.01.h5',
'prefix': 'lambda_0.01/frame'
},
{
'path': 'data/lambda_0.001.h5',
'prefix': 'lambda_0.001/frame'
}
]
def make_movie(config):
filepath = config['path']
frame_prefix = config['prefix']
make_frames(filepath, frame_prefix, title="MNIST LeNet++ Centroid Loss")
from multiprocessing import Pool
with Pool(4) as p:
p.map(make_movie, configs)
# + language="sh"
# # sudo shutdown -h now
# + language="sh"
# cd figures/animation/pre_train_lambda_0.01
# rm ../MNIST_LeNet_pre_train_lambda_0.01.mp4
# ffmpeg -framerate 10 -i frame_%03d.png -c:v libx264 -r 30 -pix_fmt yuv420p -vf "scale=trunc(iw/2)*2:trunc(ih/2)*2" ../MNIST_LeNet_pre_train_lambda_0.01.mp4
# + language="sh"
# cd figures/animation/lambda_3
# ffmpeg -framerate 10 -i frame_%03d.png -c:v libx264 -r 30 -pix_fmt yuv420p -vf "scale=trunc(iw/2)*2:trunc(ih/2)*2" ../MNIST_LeNet_lambda_3.mp4
# + language="sh"
# cd figures/animation/lambda_0.1
# ffmpeg -framerate 10 -i frame_%03d.png -c:v libx264 -r 30 -pix_fmt yuv420p -vf "scale=trunc(iw/2)*2:trunc(ih/2)*2" ../MNIST_LeNet_lambda_0.1.mp4
# + language="sh"
# cd figures/animation/lambda_0.01
# ffmpeg -framerate 10 -i frame_%03d.png -c:v libx264 -r 30 -pix_fmt yuv420p -vf "scale=trunc(iw/2)*2:trunc(ih/2)*2" ../MNIST_LeNet_lambda_0.01.mp4
# + language="sh"
# cd figures/animation/lambda_0.001
# ffmpeg -framerate 10 -i frame_%03d.png -c:v libx264 -r 30 -pix_fmt yuv420p -vf "scale=trunc(iw/2)*2:trunc(ih/2)*2" ../MNIST_LeNet_lambda_0.001.mp4
| misc/deep_learning_notes/Proj_Centroid_Loss_LeNet/LeNet_plus_centerloss/Animation of Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="KGo-kjeeHYz2"
# # PinSage graph model based recommender
# > Applying pinsage model on movielens-1m dataset
#
# - toc: false
# - badges: true
# - comments: true
# - categories: [graph, movie]
# - image:
# + id="3cJyL6GKJbfy"
# %reload_ext google.colab.data_table
# + id="D-GeM90JLOTw"
import warnings
warnings.filterwarnings('ignore')
# + id="_JkgOWMtcajy"
# !pip install dgl
# + id="gkinTrU2clkZ"
# # !wget https://s3.us-west-2.amazonaws.com/dgl-data/dataset/recsys/GATNE/example.zip && unzip example.zip
# # !wget https://s3.us-west-2.amazonaws.com/dgl-data/dataset/recsys/GATNE/amazon.zip && unzip amazon.zip
# # !wget https://s3.us-west-2.amazonaws.com/dgl-data/dataset/recsys/GATNE/youtube.zip && unzip youtube.zip
# # !wget https://s3.us-west-2.amazonaws.com/dgl-data/dataset/recsys/GATNE/twitter.zip && unzip twitter.zip
# + id="BhZPU2nIB_uR"
# !unzip example.zip
# + id="RLjy0-wdCB4Y"
# # !wget http://files.grouplens.org/datasets/movielens/ml-1m.zip && unzip ml-1m
# + colab={"base_uri": "https://localhost:8080/"} id="m88Ip1w9H7Qn" executionInfo={"status": "ok", "timestamp": 1621234814397, "user_tz": -330, "elapsed": 1470, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="1c1227ec-2959-4b8d-821c-589be230eb48"
# %%writefile builder.py
"""Graph builder from pandas dataframes"""
from collections import namedtuple
from pandas.api.types import is_numeric_dtype, is_categorical_dtype, is_categorical
import dgl
__all__ = ['PandasGraphBuilder']
def _series_to_tensor(series):
if is_categorical(series):
return torch.LongTensor(series.cat.codes.values.astype('int64'))
else: # numeric
return torch.FloatTensor(series.values)
class PandasGraphBuilder(object):
"""Creates a heterogeneous graph from multiple pandas dataframes.
Examples
--------
Let's say we have the following three pandas dataframes:
User table ``users``:
=========== =========== =======
``user_id`` ``country`` ``age``
=========== =========== =======
XYZZY U.S. 25
FOO China 24
BAR China 23
=========== =========== =======
Game table ``games``:
=========== ========= ============== ==================
``game_id`` ``title`` ``is_sandbox`` ``is_multiplayer``
=========== ========= ============== ==================
1 Minecraft True True
2 Tetris 99 False True
=========== ========= ============== ==================
Play relationship table ``plays``:
=========== =========== =========
``user_id`` ``game_id`` ``hours``
=========== =========== =========
XYZZY 1 24
FOO 1 20
FOO 2 16
BAR 2 28
=========== =========== =========
One could then create a bidirectional bipartite graph as follows:
>>> builder = PandasGraphBuilder()
>>> builder.add_entities(users, 'user_id', 'user')
>>> builder.add_entities(games, 'game_id', 'game')
>>> builder.add_binary_relations(plays, 'user_id', 'game_id', 'plays')
>>> builder.add_binary_relations(plays, 'game_id', 'user_id', 'played-by')
>>> g = builder.build()
>>> g.number_of_nodes('user')
3
>>> g.number_of_edges('plays')
4
"""
def __init__(self):
self.entity_tables = {}
self.relation_tables = {}
self.entity_pk_to_name = {} # mapping from primary key name to entity name
self.entity_pk = {} # mapping from entity name to primary key
self.entity_key_map = {} # mapping from entity names to primary key values
self.num_nodes_per_type = {}
self.edges_per_relation = {}
self.relation_name_to_etype = {}
self.relation_src_key = {} # mapping from relation name to source key
self.relation_dst_key = {} # mapping from relation name to destination key
def add_entities(self, entity_table, primary_key, name):
entities = entity_table[primary_key].astype('category')
if not (entities.value_counts() == 1).all():
raise ValueError('Different entity with the same primary key detected.')
# preserve the category order in the original entity table
entities = entities.cat.reorder_categories(entity_table[primary_key].values)
self.entity_pk_to_name[primary_key] = name
self.entity_pk[name] = primary_key
self.num_nodes_per_type[name] = entity_table.shape[0]
self.entity_key_map[name] = entities
self.entity_tables[name] = entity_table
def add_binary_relations(self, relation_table, source_key, destination_key, name):
src = relation_table[source_key].astype('category')
src = src.cat.set_categories(
self.entity_key_map[self.entity_pk_to_name[source_key]].cat.categories)
dst = relation_table[destination_key].astype('category')
dst = dst.cat.set_categories(
self.entity_key_map[self.entity_pk_to_name[destination_key]].cat.categories)
if src.isnull().any():
raise ValueError(
'Some source entities in relation %s do not exist in entity %s.' %
(name, source_key))
if dst.isnull().any():
raise ValueError(
'Some destination entities in relation %s do not exist in entity %s.' %
(name, destination_key))
srctype = self.entity_pk_to_name[source_key]
dsttype = self.entity_pk_to_name[destination_key]
etype = (srctype, name, dsttype)
self.relation_name_to_etype[name] = etype
self.edges_per_relation[etype] = (src.cat.codes.values.astype('int64'), dst.cat.codes.values.astype('int64'))
self.relation_tables[name] = relation_table
self.relation_src_key[name] = source_key
self.relation_dst_key[name] = destination_key
def build(self):
# Create heterograph
graph = dgl.heterograph(self.edges_per_relation, self.num_nodes_per_type)
return graph
# + colab={"base_uri": "https://localhost:8080/"} id="rL9meK9zIF6x" executionInfo={"status": "ok", "timestamp": 1621234848112, "user_tz": -330, "elapsed": 1070, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="e32ac217-2340-4528-e4a7-82207a4ec1d9"
# %%writefile data_utils.py
import torch
import dgl
import numpy as np
import scipy.sparse as ssp
import tqdm
import dask.dataframe as dd
# This is the train-test split method most of the recommender system papers running on MovieLens
# takes. It essentially follows the intuition of "training on the past and predict the future".
# One can also change the threshold to make validation and test set take larger proportions.
def train_test_split_by_time(df, timestamp, user):
df['train_mask'] = np.ones((len(df),), dtype=np.bool)
df['val_mask'] = np.zeros((len(df),), dtype=np.bool)
df['test_mask'] = np.zeros((len(df),), dtype=np.bool)
df = dd.from_pandas(df, npartitions=10)
def train_test_split(df):
df = df.sort_values([timestamp])
if df.shape[0] > 1:
df.iloc[-1, -3] = False
df.iloc[-1, -1] = True
if df.shape[0] > 2:
df.iloc[-2, -3] = False
df.iloc[-2, -2] = True
return df
df = df.groupby(user, group_keys=False).apply(train_test_split).compute(scheduler='processes').sort_index()
print(df[df[user] == df[user].unique()[0]].sort_values(timestamp))
return df['train_mask'].to_numpy().nonzero()[0], \
df['val_mask'].to_numpy().nonzero()[0], \
df['test_mask'].to_numpy().nonzero()[0]
def build_train_graph(g, train_indices, utype, itype, etype, etype_rev):
train_g = g.edge_subgraph(
{etype: train_indices, etype_rev: train_indices},
preserve_nodes=True)
# remove the induced node IDs - should be assigned by model instead
del train_g.nodes[utype].data[dgl.NID]
del train_g.nodes[itype].data[dgl.NID]
# copy features
for ntype in g.ntypes:
for col, data in g.nodes[ntype].data.items():
train_g.nodes[ntype].data[col] = data
for etype in g.etypes:
for col, data in g.edges[etype].data.items():
train_g.edges[etype].data[col] = data[train_g.edges[etype].data[dgl.EID]]
return train_g
def build_val_test_matrix(g, val_indices, test_indices, utype, itype, etype):
n_users = g.number_of_nodes(utype)
n_items = g.number_of_nodes(itype)
val_src, val_dst = g.find_edges(val_indices, etype=etype)
test_src, test_dst = g.find_edges(test_indices, etype=etype)
val_src = val_src.numpy()
val_dst = val_dst.numpy()
test_src = test_src.numpy()
test_dst = test_dst.numpy()
val_matrix = ssp.coo_matrix((np.ones_like(val_src), (val_src, val_dst)), (n_users, n_items))
test_matrix = ssp.coo_matrix((np.ones_like(test_src), (test_src, test_dst)), (n_users, n_items))
return val_matrix, test_matrix
def linear_normalize(values):
return (values - values.min(0, keepdims=True)) / \
(values.max(0, keepdims=True) - values.min(0, keepdims=True))
# + id="r9YUPFDbIWjM"
# !pip install dask[dataframe]
# + id="nK2226CYHmCC"
"""
Script that reads from raw MovieLens-1M data and dumps into a pickle
file the following:
* A heterogeneous graph with categorical features.
* A list with all the movie titles. The movie titles correspond to
the movie nodes in the heterogeneous graph.
This script exemplifies how to prepare tabular data with textual
features. Since DGL graphs do not store variable-length features, we
instead put variable-length features into a more suitable container
(e.g. torchtext to handle list of texts)
"""
# + id="zKlLCpOlCanU"
import os
import re
import argparse
import pickle
import pandas as pd
import numpy as np
import scipy.sparse as ssp
import dgl
import torch
import torchtext
from builder import PandasGraphBuilder
from data_utils import *
# + id="8ZUqdOZtI6o2"
# parser = argparse.ArgumentParser()
# parser.add_argument('directory', type=str)
# parser.add_argument('output_path', type=str)
# args = parser.parse_args()
directory = './ml-1m'
output_path = './ml-graph-data.pkl'
# + id="vJSbnupmI6lw"
## Build heterogeneous graph
# + id="JRf3Cn7KI6i5"
# Load data
users = []
with open(os.path.join(directory, 'users.dat'), encoding='latin1') as f:
for l in f:
id_, gender, age, occupation, zip_ = l.strip().split('::')
users.append({
'user_id': int(id_),
'gender': gender,
'age': age,
'occupation': occupation,
'zip': zip_,
})
users = pd.DataFrame(users).astype('category')
# + colab={"base_uri": "https://localhost:8080/", "height": 194} id="CpgWv-wLJAhV" executionInfo={"status": "ok", "timestamp": 1621235210404, "user_tz": -330, "elapsed": 1120, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="7a527419-c7fa-49ed-bcce-e42db9c19553"
users.head()
# + id="VsD6wICdJAch"
movies = []
with open(os.path.join(directory, 'movies.dat'), encoding='latin1') as f:
for l in f:
id_, title, genres = l.strip().split('::')
genres_set = set(genres.split('|'))
# extract year
assert re.match(r'.*\([0-9]{4}\)$', title)
year = title[-5:-1]
title = title[:-6].strip()
data = {'movie_id': int(id_), 'title': title, 'year': year}
for g in genres_set:
data[g] = True
movies.append(data)
movies = pd.DataFrame(movies).astype({'year': 'category'})
# + colab={"base_uri": "https://localhost:8080/", "height": 194} id="8eFQ-9yEJJNl" executionInfo={"status": "ok", "timestamp": 1621235248289, "user_tz": -330, "elapsed": 1027, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="9af3e74b-fc9c-451a-e963-852668d707ca"
movies.head().iloc[:,:10]
# + id="d4XufOxiJJLA"
ratings = []
with open(os.path.join(directory, 'ratings.dat'), encoding='latin1') as f:
for l in f:
user_id, movie_id, rating, timestamp = [int(_) for _ in l.split('::')]
ratings.append({
'user_id': user_id,
'movie_id': movie_id,
'rating': rating,
'timestamp': timestamp,
})
ratings = pd.DataFrame(ratings)
# + colab={"base_uri": "https://localhost:8080/", "height": 194} id="EHxuDrxsJXPU" executionInfo={"status": "ok", "timestamp": 1621235270306, "user_tz": -330, "elapsed": 1194, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="80e99e5e-09e7-4a6f-dd44-0c24aefc3476"
ratings.head()
# + id="I85IXVwAJXPW"
# Filter the users and items that never appear in the rating table.
distinct_users_in_ratings = ratings['user_id'].unique()
distinct_movies_in_ratings = ratings['movie_id'].unique()
users = users[users['user_id'].isin(distinct_users_in_ratings)]
movies = movies[movies['movie_id'].isin(distinct_movies_in_ratings)]
# + id="4hTUdj4JJXPX"
# Group the movie features into genres (a vector), year (a category), title (a string)
genre_columns = movies.columns.drop(['movie_id', 'title', 'year'])
movies[genre_columns] = movies[genre_columns].fillna(False).astype('bool')
movies_categorical = movies.drop('title', axis=1)
# + id="R5xWZCItJXPY"
# Build graph
graph_builder = PandasGraphBuilder()
graph_builder.add_entities(users, 'user_id', 'user')
graph_builder.add_entities(movies_categorical, 'movie_id', 'movie')
graph_builder.add_binary_relations(ratings, 'user_id', 'movie_id', 'watched')
graph_builder.add_binary_relations(ratings, 'movie_id', 'user_id', 'watched-by')
g = graph_builder.build()
# + id="EXnIIM2WJJGT"
# Assign features.
# Note that variable-sized features such as texts or images are handled elsewhere.
g.nodes['user'].data['gender'] = torch.LongTensor(users['gender'].cat.codes.values)
g.nodes['user'].data['age'] = torch.LongTensor(users['age'].cat.codes.values)
g.nodes['user'].data['occupation'] = torch.LongTensor(users['occupation'].cat.codes.values)
g.nodes['user'].data['zip'] = torch.LongTensor(users['zip'].cat.codes.values)
g.nodes['movie'].data['year'] = torch.LongTensor(movies['year'].cat.codes.values)
g.nodes['movie'].data['genre'] = torch.FloatTensor(movies[genre_columns].values)
g.edges['watched'].data['rating'] = torch.LongTensor(ratings['rating'].values)
g.edges['watched'].data['timestamp'] = torch.LongTensor(ratings['timestamp'].values)
g.edges['watched-by'].data['rating'] = torch.LongTensor(ratings['rating'].values)
g.edges['watched-by'].data['timestamp'] = torch.LongTensor(ratings['timestamp'].values)
# + colab={"base_uri": "https://localhost:8080/"} id="6-78nSGXLCKO" executionInfo={"status": "ok", "timestamp": 1621235692031, "user_tz": -330, "elapsed": 11215, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="e9ffbb28-7848-4211-a186-d533b63a61f4"
# Train-validation-test split
# This is a little bit tricky as we want to select the last interaction for test, and the
# second-to-last interaction for validation.
train_indices, val_indices, test_indices = train_test_split_by_time(ratings, 'timestamp', 'user_id')
# + id="7osDJmd0LeeY"
# Build the graph with training interactions only.
train_g = build_train_graph(g, train_indices, 'user', 'movie', 'watched', 'watched-by')
assert train_g.out_degrees(etype='watched').min() > 0
# + id="pxnbc5LbLh59"
# Build the user-item sparse matrix for validation and test set.
val_matrix, test_matrix = build_val_test_matrix(g, val_indices, test_indices, 'user', 'movie', 'watched')
# + id="OlbHm1BMLmQo"
## Build title set
movie_textual_dataset = {'title': movies['title'].values}
# + id="U32UKwxzLy11"
# The model should build their own vocabulary and process the texts. Here is one example
# of using torchtext to pad and numericalize a batch of strings.
# field = torchtext.data.Field(include_lengths=True, lower=True, batch_first=True)
# examples = [torchtext.data.Example.fromlist([t], [('title', title_field)]) for t in texts]
# titleset = torchtext.data.Dataset(examples, [('title', title_field)])
# field.build_vocab(titleset.title, vectors='fasttext.simple.300d')
# token_ids, lengths = field.process([examples[0].title, examples[1].title])
# + id="q79CfTh9IQdV"
## Dump the graph and the datasets
dataset = {
'train-graph': train_g,
'val-matrix': val_matrix,
'test-matrix': test_matrix,
'item-texts': movie_textual_dataset,
'item-images': None,
'user-type': 'user',
'item-type': 'movie',
'user-to-item-type': 'watched',
'item-to-user-type': 'watched-by',
'timestamp-edge-column': 'timestamp'}
with open(output_path, 'wb') as f:
pickle.dump(dataset, f)
# + colab={"base_uri": "https://localhost:8080/"} id="AmkRQU-rIQZ6" executionInfo={"status": "ok", "timestamp": 1621236152369, "user_tz": -330, "elapsed": 1393, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="45eebeaf-c40e-4c56-c260-6f1ce0cd5be6"
# %%writefile evaluation.py
import numpy as np
import torch
import pickle
import dgl
import argparse
def prec(recommendations, ground_truth):
n_users, n_items = ground_truth.shape
K = recommendations.shape[1]
user_idx = np.repeat(np.arange(n_users), K)
item_idx = recommendations.flatten()
relevance = ground_truth[user_idx, item_idx].reshape((n_users, K))
hit = relevance.any(axis=1).mean()
return hit
class LatestNNRecommender(object):
def __init__(self, user_ntype, item_ntype, user_to_item_etype, timestamp, batch_size):
self.user_ntype = user_ntype
self.item_ntype = item_ntype
self.user_to_item_etype = user_to_item_etype
self.batch_size = batch_size
self.timestamp = timestamp
def recommend(self, full_graph, K, h_user, h_item):
"""
Return a (n_user, K) matrix of recommended items for each user
"""
graph_slice = full_graph.edge_type_subgraph([self.user_to_item_etype])
n_users = full_graph.number_of_nodes(self.user_ntype)
latest_interactions = dgl.sampling.select_topk(graph_slice, 1, self.timestamp, edge_dir='out')
user, latest_items = latest_interactions.all_edges(form='uv', order='srcdst')
# each user should have at least one "latest" interaction
assert torch.equal(user, torch.arange(n_users))
recommended_batches = []
user_batches = torch.arange(n_users).split(self.batch_size)
for user_batch in user_batches:
latest_item_batch = latest_items[user_batch].to(device=h_item.device)
dist = h_item[latest_item_batch] @ h_item.t()
# exclude items that are already interacted
for i, u in enumerate(user_batch.tolist()):
interacted_items = full_graph.successors(u, etype=self.user_to_item_etype)
dist[i, interacted_items] = -np.inf
recommended_batches.append(dist.topk(K, 1)[1])
recommendations = torch.cat(recommended_batches, 0)
return recommendations
def evaluate_nn(dataset, h_item, k, batch_size):
g = dataset['train-graph']
val_matrix = dataset['val-matrix'].tocsr()
test_matrix = dataset['test-matrix'].tocsr()
item_texts = dataset['item-texts']
user_ntype = dataset['user-type']
item_ntype = dataset['item-type']
user_to_item_etype = dataset['user-to-item-type']
timestamp = dataset['timestamp-edge-column']
rec_engine = LatestNNRecommender(
user_ntype, item_ntype, user_to_item_etype, timestamp, batch_size)
recommendations = rec_engine.recommend(g, k, None, h_item).cpu().numpy()
return prec(recommendations, val_matrix)
if __name__ == '__main__':
parser = argparse.ArgumentParser()
parser.add_argument('dataset_path', type=str)
parser.add_argument('item_embedding_path', type=str)
parser.add_argument('-k', type=int, default=10)
parser.add_argument('--batch-size', type=int, default=32)
args = parser.parse_args()
with open(args.dataset_path, 'rb') as f:
dataset = pickle.load(f)
with open(args.item_embedding_path, 'rb') as f:
emb = torch.FloatTensor(pickle.load(f))
print(evaluate_nn(dataset, emb, args.k, args.batch_size))
# + colab={"base_uri": "https://localhost:8080/"} id="3xmC2B58IQWf" executionInfo={"status": "ok", "timestamp": 1621236173392, "user_tz": -330, "elapsed": 1163, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="e5281891-163f-4a06-c08d-42861e49ff9f"
# %%writefile layers.py
import torch
import torch.nn as nn
import torch.nn.functional as F
import dgl
import dgl.nn.pytorch as dglnn
import dgl.function as fn
def disable_grad(module):
for param in module.parameters():
param.requires_grad = False
def _init_input_modules(g, ntype, textset, hidden_dims):
# We initialize the linear projections of each input feature ``x`` as
# follows:
# * If ``x`` is a scalar integral feature, we assume that ``x`` is a categorical
# feature, and assume the range of ``x`` is 0..max(x).
# * If ``x`` is a float one-dimensional feature, we assume that ``x`` is a
# numeric vector.
# * If ``x`` is a field of a textset, we process it as bag of words.
module_dict = nn.ModuleDict()
for column, data in g.nodes[ntype].data.items():
if column == dgl.NID:
continue
if data.dtype == torch.float32:
assert data.ndim == 2
m = nn.Linear(data.shape[1], hidden_dims)
nn.init.xavier_uniform_(m.weight)
nn.init.constant_(m.bias, 0)
module_dict[column] = m
elif data.dtype == torch.int64:
assert data.ndim == 1
m = nn.Embedding(
data.max() + 2, hidden_dims, padding_idx=-1)
nn.init.xavier_uniform_(m.weight)
module_dict[column] = m
if textset is not None:
for column, field in textset.fields.items():
if field.vocab.vectors:
module_dict[column] = BagOfWordsPretrained(field, hidden_dims)
else:
module_dict[column] = BagOfWords(field, hidden_dims)
return module_dict
class BagOfWordsPretrained(nn.Module):
def __init__(self, field, hidden_dims):
super().__init__()
input_dims = field.vocab.vectors.shape[1]
self.emb = nn.Embedding(
len(field.vocab.itos), input_dims,
padding_idx=field.vocab.stoi[field.pad_token])
self.emb.weight[:] = field.vocab.vectors
self.proj = nn.Linear(input_dims, hidden_dims)
nn.init.xavier_uniform_(self.proj.weight)
nn.init.constant_(self.proj.bias, 0)
disable_grad(self.emb)
def forward(self, x, length):
"""
x: (batch_size, max_length) LongTensor
length: (batch_size,) LongTensor
"""
x = self.emb(x).sum(1) / length.unsqueeze(1).float()
return self.proj(x)
class BagOfWords(nn.Module):
def __init__(self, field, hidden_dims):
super().__init__()
self.emb = nn.Embedding(
len(field.vocab.itos), hidden_dims,
padding_idx=field.vocab.stoi[field.pad_token])
nn.init.xavier_uniform_(self.emb.weight)
def forward(self, x, length):
return self.emb(x).sum(1) / length.unsqueeze(1).float()
class LinearProjector(nn.Module):
"""
Projects each input feature of the graph linearly and sums them up
"""
def __init__(self, full_graph, ntype, textset, hidden_dims):
super().__init__()
self.ntype = ntype
self.inputs = _init_input_modules(full_graph, ntype, textset, hidden_dims)
def forward(self, ndata):
projections = []
for feature, data in ndata.items():
if feature == dgl.NID or feature.endswith('__len'):
# This is an additional feature indicating the length of the ``feature``
# column; we shouldn't process this.
continue
module = self.inputs[feature]
if isinstance(module, (BagOfWords, BagOfWordsPretrained)):
# Textual feature; find the length and pass it to the textual module.
length = ndata[feature + '__len']
result = module(data, length)
else:
result = module(data)
projections.append(result)
return torch.stack(projections, 1).sum(1)
class WeightedSAGEConv(nn.Module):
def __init__(self, input_dims, hidden_dims, output_dims, act=F.relu):
super().__init__()
self.act = act
self.Q = nn.Linear(input_dims, hidden_dims)
self.W = nn.Linear(input_dims + hidden_dims, output_dims)
self.reset_parameters()
self.dropout = nn.Dropout(0.5)
def reset_parameters(self):
gain = nn.init.calculate_gain('relu')
nn.init.xavier_uniform_(self.Q.weight, gain=gain)
nn.init.xavier_uniform_(self.W.weight, gain=gain)
nn.init.constant_(self.Q.bias, 0)
nn.init.constant_(self.W.bias, 0)
def forward(self, g, h, weights):
"""
g : graph
h : node features
weights : scalar edge weights
"""
h_src, h_dst = h
with g.local_scope():
g.srcdata['n'] = self.act(self.Q(self.dropout(h_src)))
g.edata['w'] = weights.float()
g.update_all(fn.u_mul_e('n', 'w', 'm'), fn.sum('m', 'n'))
g.update_all(fn.copy_e('w', 'm'), fn.sum('m', 'ws'))
n = g.dstdata['n']
ws = g.dstdata['ws'].unsqueeze(1).clamp(min=1)
z = self.act(self.W(self.dropout(torch.cat([n / ws, h_dst], 1))))
z_norm = z.norm(2, 1, keepdim=True)
z_norm = torch.where(z_norm == 0, torch.tensor(1.).to(z_norm), z_norm)
z = z / z_norm
return z
class SAGENet(nn.Module):
def __init__(self, hidden_dims, n_layers):
"""
g : DGLHeteroGraph
The user-item interaction graph.
This is only for finding the range of categorical variables.
item_textsets : torchtext.data.Dataset
The textual features of each item node.
"""
super().__init__()
self.convs = nn.ModuleList()
for _ in range(n_layers):
self.convs.append(WeightedSAGEConv(hidden_dims, hidden_dims, hidden_dims))
def forward(self, blocks, h):
for layer, block in zip(self.convs, blocks):
h_dst = h[:block.number_of_nodes('DST/' + block.ntypes[0])]
h = layer(block, (h, h_dst), block.edata['weights'])
return h
class ItemToItemScorer(nn.Module):
def __init__(self, full_graph, ntype):
super().__init__()
n_nodes = full_graph.number_of_nodes(ntype)
self.bias = nn.Parameter(torch.zeros(n_nodes))
def _add_bias(self, edges):
bias_src = self.bias[edges.src[dgl.NID]]
bias_dst = self.bias[edges.dst[dgl.NID]]
return {'s': edges.data['s'] + bias_src + bias_dst}
def forward(self, item_item_graph, h):
"""
item_item_graph : graph consists of edges connecting the pairs
h : hidden state of every node
"""
with item_item_graph.local_scope():
item_item_graph.ndata['h'] = h
item_item_graph.apply_edges(fn.u_dot_v('h', 'h', 's'))
item_item_graph.apply_edges(self._add_bias)
pair_score = item_item_graph.edata['s']
return pair_score
# + colab={"base_uri": "https://localhost:8080/"} id="zkv9CEXiNMSR" executionInfo={"status": "ok", "timestamp": 1621236200548, "user_tz": -330, "elapsed": 1282, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="40a83e2e-cc88-485b-d697-8b1c569eed29"
# %%writefile sampler.py
import numpy as np
import dgl
import torch
from torch.utils.data import IterableDataset, DataLoader
def compact_and_copy(frontier, seeds):
block = dgl.to_block(frontier, seeds)
for col, data in frontier.edata.items():
if col == dgl.EID:
continue
block.edata[col] = data[block.edata[dgl.EID]]
return block
class ItemToItemBatchSampler(IterableDataset):
def __init__(self, g, user_type, item_type, batch_size):
self.g = g
self.user_type = user_type
self.item_type = item_type
self.user_to_item_etype = list(g.metagraph()[user_type][item_type])[0]
self.item_to_user_etype = list(g.metagraph()[item_type][user_type])[0]
self.batch_size = batch_size
def __iter__(self):
while True:
heads = torch.randint(0, self.g.number_of_nodes(self.item_type), (self.batch_size,))
tails = dgl.sampling.random_walk(
self.g,
heads,
metapath=[self.item_to_user_etype, self.user_to_item_etype])[0][:, 2]
neg_tails = torch.randint(0, self.g.number_of_nodes(self.item_type), (self.batch_size,))
mask = (tails != -1)
yield heads[mask], tails[mask], neg_tails[mask]
class NeighborSampler(object):
def __init__(self, g, user_type, item_type, random_walk_length, random_walk_restart_prob,
num_random_walks, num_neighbors, num_layers):
self.g = g
self.user_type = user_type
self.item_type = item_type
self.user_to_item_etype = list(g.metagraph()[user_type][item_type])[0]
self.item_to_user_etype = list(g.metagraph()[item_type][user_type])[0]
self.samplers = [
dgl.sampling.PinSAGESampler(g, item_type, user_type, random_walk_length,
random_walk_restart_prob, num_random_walks, num_neighbors)
for _ in range(num_layers)]
def sample_blocks(self, seeds, heads=None, tails=None, neg_tails=None):
blocks = []
for sampler in self.samplers:
frontier = sampler(seeds)
if heads is not None:
eids = frontier.edge_ids(torch.cat([heads, heads]), torch.cat([tails, neg_tails]), return_uv=True)[2]
if len(eids) > 0:
old_frontier = frontier
frontier = dgl.remove_edges(old_frontier, eids)
#print(old_frontier)
#print(frontier)
#print(frontier.edata['weights'])
#frontier.edata['weights'] = old_frontier.edata['weights'][frontier.edata[dgl.EID]]
block = compact_and_copy(frontier, seeds)
seeds = block.srcdata[dgl.NID]
blocks.insert(0, block)
return blocks
def sample_from_item_pairs(self, heads, tails, neg_tails):
# Create a graph with positive connections only and another graph with negative
# connections only.
pos_graph = dgl.graph(
(heads, tails),
num_nodes=self.g.number_of_nodes(self.item_type))
neg_graph = dgl.graph(
(heads, neg_tails),
num_nodes=self.g.number_of_nodes(self.item_type))
pos_graph, neg_graph = dgl.compact_graphs([pos_graph, neg_graph])
seeds = pos_graph.ndata[dgl.NID]
blocks = self.sample_blocks(seeds, heads, tails, neg_tails)
return pos_graph, neg_graph, blocks
def assign_simple_node_features(ndata, g, ntype, assign_id=False):
"""
Copies data to the given block from the corresponding nodes in the original graph.
"""
for col in g.nodes[ntype].data.keys():
if not assign_id and col == dgl.NID:
continue
induced_nodes = ndata[dgl.NID]
ndata[col] = g.nodes[ntype].data[col][induced_nodes]
def assign_textual_node_features(ndata, textset, ntype):
"""
Assigns numericalized tokens from a torchtext dataset to given block.
The numericalized tokens would be stored in the block as node features
with the same name as ``field_name``.
The length would be stored as another node feature with name
``field_name + '__len'``.
block : DGLHeteroGraph
First element of the compacted blocks, with "dgl.NID" as the
corresponding node ID in the original graph, hence the index to the
text dataset.
The numericalized tokens (and lengths if available) would be stored
onto the blocks as new node features.
textset : torchtext.data.Dataset
A torchtext dataset whose number of examples is the same as that
of nodes in the original graph.
"""
node_ids = ndata[dgl.NID].numpy()
for field_name, field in textset.fields.items():
examples = [getattr(textset[i], field_name) for i in node_ids]
tokens, lengths = field.process(examples)
if not field.batch_first:
tokens = tokens.t()
ndata[field_name] = tokens
ndata[field_name + '__len'] = lengths
def assign_features_to_blocks(blocks, g, textset, ntype):
# For the first block (which is closest to the input), copy the features from
# the original graph as well as the texts.
assign_simple_node_features(blocks[0].srcdata, g, ntype)
assign_textual_node_features(blocks[0].srcdata, textset, ntype)
assign_simple_node_features(blocks[-1].dstdata, g, ntype)
assign_textual_node_features(blocks[-1].dstdata, textset, ntype)
class PinSAGECollator(object):
def __init__(self, sampler, g, ntype, textset):
self.sampler = sampler
self.ntype = ntype
self.g = g
self.textset = textset
def collate_train(self, batches):
heads, tails, neg_tails = batches[0]
# Construct multilayer neighborhood via PinSAGE...
pos_graph, neg_graph, blocks = self.sampler.sample_from_item_pairs(heads, tails, neg_tails)
assign_features_to_blocks(blocks, self.g, self.textset, self.ntype)
return pos_graph, neg_graph, blocks
def collate_test(self, samples):
batch = torch.LongTensor(samples)
blocks = self.sampler.sample_blocks(batch)
assign_features_to_blocks(blocks, self.g, self.textset, self.ntype)
return blocks
# + id="3n-wpT1VNWam"
import pickle
import argparse
import numpy as np
import torch
import torch.nn as nn
from torch.utils.data import DataLoader
import torchtext
import dgl
import tqdm
import layers
import sampler as sampler_module
import evaluation
# + id="dBZNZjVzNYtH"
class PinSAGEModel(nn.Module):
def __init__(self, full_graph, ntype, textsets, hidden_dims, n_layers):
super().__init__()
self.proj = layers.LinearProjector(full_graph, ntype, textsets, hidden_dims)
self.sage = layers.SAGENet(hidden_dims, n_layers)
self.scorer = layers.ItemToItemScorer(full_graph, ntype)
def forward(self, pos_graph, neg_graph, blocks):
h_item = self.get_repr(blocks)
pos_score = self.scorer(pos_graph, h_item)
neg_score = self.scorer(neg_graph, h_item)
return (neg_score - pos_score + 1).clamp(min=0)
def get_repr(self, blocks):
h_item = self.proj(blocks[0].srcdata)
h_item_dst = self.proj(blocks[-1].dstdata)
return h_item_dst + self.sage(blocks, h_item)
# + id="GX-PImUXNuZZ"
parser = argparse.ArgumentParser()
parser.add_argument('--dataset_path', type=str, default='./ml-graph-data.pkl')
parser.add_argument('--random-walk-length', type=int, default=2)
parser.add_argument('--random-walk-restart-prob', type=float, default=0.5)
parser.add_argument('--num-random-walks', type=int, default=10)
parser.add_argument('--num-neighbors', type=int, default=3)
parser.add_argument('--num-layers', type=int, default=2)
parser.add_argument('--hidden-dims', type=int, default=16)
parser.add_argument('--batch-size', type=int, default=32)
parser.add_argument('--device', type=str, default='cpu') # can also be "cuda:0"
parser.add_argument('--num-epochs', type=int, default=1)
parser.add_argument('--batches-per-epoch', type=int, default=20000)
parser.add_argument('--num-workers', type=int, default=0)
parser.add_argument('--lr', type=float, default=3e-5)
parser.add_argument('-k', type=int, default=10)
args, unknown = parser.parse_known_args()
# + id="Pt3zREy8Qtcl"
# Load dataset
with open(args.dataset_path, 'rb') as f:
dataset = pickle.load(f)
# + id="RO7FR1jnQ2ok"
g = dataset['train-graph']
val_matrix = dataset['val-matrix'].tocsr()
test_matrix = dataset['test-matrix'].tocsr()
item_texts = dataset['item-texts']
user_ntype = dataset['user-type']
item_ntype = dataset['item-type']
user_to_item_etype = dataset['user-to-item-type']
timestamp = dataset['timestamp-edge-column']
# + id="uPUJQ48eQ4rf"
device = torch.device(args.device)
# + id="OXlNFiRcQ8N2"
# Assign user and movie IDs and use them as features (to learn an individual trainable
# embedding for each entity)
g.nodes[user_ntype].data['id'] = torch.arange(g.number_of_nodes(user_ntype))
g.nodes[item_ntype].data['id'] = torch.arange(g.number_of_nodes(item_ntype))
# + id="Ps7F5tajQ8JN"
# Prepare torchtext dataset and vocabulary
fields = {}
examples = []
for key, texts in item_texts.items():
fields[key] = torchtext.legacy.data.Field(include_lengths=True, lower=True, batch_first=True)
for i in range(g.number_of_nodes(item_ntype)):
example = torchtext.legacy.data.Example.fromlist(
[item_texts[key][i] for key in item_texts.keys()],
[(key, fields[key]) for key in item_texts.keys()])
examples.append(example)
textset = torchtext.legacy.data.Dataset(examples, fields)
for key, field in fields.items():
field.build_vocab(getattr(textset, key))
#field.build_vocab(getattr(textset, key), vectors='fasttext.simple.300d')
# + id="2hSz7EwMNYoc"
# Sampler
batch_sampler = sampler_module.ItemToItemBatchSampler(g, user_ntype, item_ntype, args.batch_size)
neighbor_sampler = sampler_module.NeighborSampler(g, user_ntype, item_ntype, args.random_walk_length,
args.random_walk_restart_prob, args.num_random_walks,
args.num_neighbors, args.num_layers)
collator = sampler_module.PinSAGECollator(neighbor_sampler, g, item_ntype, textset)
dataloader = DataLoader(batch_sampler, collate_fn=collator.collate_train, num_workers=args.num_workers)
dataloader_test = DataLoader(torch.arange(g.number_of_nodes(item_ntype)), batch_size=args.batch_size,
collate_fn=collator.collate_test, num_workers=args.num_workers)
dataloader_it = iter(dataloader)
# + id="B4r309MtNYim"
# Model
model = PinSAGEModel(g, item_ntype, textset, args.hidden_dims, args.num_layers).to(device)
# + id="R7WgJFFsSI-R"
# Optimizer
opt = torch.optim.Adam(model.parameters(), lr=args.lr)
# + colab={"base_uri": "https://localhost:8080/"} id="_NsYPL1bSseZ" executionInfo={"status": "ok", "timestamp": 1621238437494, "user_tz": -330, "elapsed": 114798, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="dc66e775-67e8-4a01-b305-85e1ec5a6bf9"
# For each batch of head-tail-negative triplets...
for epoch_id in range(args.num_epochs):
model.train()
for batch_id in tqdm.trange(args.batches_per_epoch):
pos_graph, neg_graph, blocks = next(dataloader_it)
# Copy to GPU
for i in range(len(blocks)):
blocks[i] = blocks[i].to(device)
pos_graph = pos_graph.to(device)
neg_graph = neg_graph.to(device)
loss = model(pos_graph, neg_graph, blocks).mean()
opt.zero_grad()
loss.backward()
opt.step()
# + colab={"base_uri": "https://localhost:8080/"} id="KSe9dRhQS0t_" executionInfo={"status": "ok", "timestamp": 1621238440227, "user_tz": -330, "elapsed": 7378, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="37fa6835-78d9-4c96-8bc4-b76156e4a060"
# Evaluate HIT@10
model.eval()
with torch.no_grad():
item_batches = torch.arange(g.number_of_nodes(item_ntype)).split(args.batch_size)
h_item_batches = []
for blocks in dataloader_test:
for i in range(len(blocks)):
blocks[i] = blocks[i].to(device)
h_item_batches.append(model.get_repr(blocks))
h_item = torch.cat(h_item_batches, 0)
print(evaluation.evaluate_nn(dataset, h_item, args.k, args.batch_size))
# + [markdown] id="xE3cBBjIVev-"
# https://github.com/dmlc/dgl/tree/master/examples/pytorch/pinsage
| _docs/nbs/recograph-01-2021-06-23-pinsage-graph-movielens-1m.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 2020년 7월 10일 금요일
# ### HackerRank - Day14 : Scope (Python)
# ### 문제 : https://www.hackerrank.com/challenges/30-scope/problem
# ### 블로그 : https://somjang.tistory.com/entry/HackerRank-Day-14-Scope-Python
# ### 첫번째 시도
# +
class Difference:
def __init__(self, a):
self.__elements = a
self.maximumDifference = 0
def computeDifference(self):
max_num = max(self.__elements)
min_num = min(self.__elements)
max_differ = abs(max_num - min_num)
self.maximumDifference = max_differ
# Add your code here
# End of Difference class
_ = input()
a = [int(e) for e in input().split(' ')]
d = Difference(a)
d.computeDifference()
print(d.maximumDifference)
| DAY 101 ~ 200/DAY155_[HackerRank] Day14 Scope (Python).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="adTDe2CTh3MU"
# # Neural Machine Translation
#
# Welcome to your first programming assignment for this week!
#
# * You will build a Neural Machine Translation (NMT) model to translate human-readable dates ("25th of June, 2009") into machine-readable dates ("2009-06-25").
# * You will do this using an attention model, one of the most sophisticated sequence-to-sequence models.
#
# This notebook was produced together with NVIDIA's Deep Learning Institute.
# + [markdown] id="0LCkjDBFh3Md"
# ## Table of Contents
#
# - [Packages](#0)
# - [1 - Translating Human Readable Dates Into Machine Readable Dates](#1)
# - [1.1 - Dataset](#1-1)
# - [2 - Neural Machine Translation with Attention](#2)
# - [2.1 - Attention Mechanism](#2-1)
# - [Exercise 1 - one_step_attention](#ex-1)
# - [Exercise 2 - modelf](#ex-2)
# - [Exercise 3 - Compile the Model](#ex-3)
# - [3 - Visualizing Attention (Optional / Ungraded)](#3)
# - [3.1 - Getting the Attention Weights From the Network](#3-1)
# -
# <a name='0'></a>
# ## Packages
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 14508, "status": "ok", "timestamp": 1612468511651, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gip7OjOkdNkKxKDyWEQAq1o8ccGN_HrBTGdqjgQ=s64", "userId": "08094225471505108399"}, "user_tz": -300} id="RcBRMzPiiMmp" outputId="17e9a429-5bb6-4401-a23a-f8f756d6113d"
from tensorflow.keras.layers import Bidirectional, Concatenate, Permute, Dot, Input, LSTM, Multiply
from tensorflow.keras.layers import RepeatVector, Dense, Activation, Lambda
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.models import load_model, Model
import tensorflow.keras.backend as K
import tensorflow as tf
import numpy as np
from faker import Faker
import random
from tqdm import tqdm
from babel.dates import format_date
from nmt_utils import *
import matplotlib.pyplot as plt
# %matplotlib inline
# + [markdown] id="J0pkH-k0h3Mf"
# <a name='1'></a>
# ## 1 - Translating Human Readable Dates Into Machine Readable Dates
#
# * The model you will build here could be used to translate from one language to another, such as translating from English to Hindi.
# * However, language translation requires massive datasets and usually takes days of training on GPUs.
# * To give you a place to experiment with these models without using massive datasets, we will perform a simpler "date translation" task.
# * The network will input a date written in a variety of possible formats (*e.g. "the 29th of August 1958", "03/30/1968", "24 JUNE 1987"*)
# * The network will translate them into standardized, machine readable dates (*e.g. "1958-08-29", "1968-03-30", "1987-06-24"*).
# * We will have the network learn to output dates in the common machine-readable format YYYY-MM-DD.
#
# <!--
# Take a look at [nmt_utils.py](./nmt_utils.py) to see all the formatting. Count and figure out how the formats work, you will need this knowledge later. !-->
# + [markdown] id="8BhEaJvph3Mf"
# <a name='1-1'></a>
# ### 1.1 - Dataset
#
# We will train the model on a dataset of 10,000 human readable dates and their equivalent, standardized, machine readable dates. Let's run the following cells to load the dataset and print some examples.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 16981, "status": "ok", "timestamp": 1612468514155, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gip7OjOkdNkKxKDyWEQAq1o8ccGN_HrBTGdqjgQ=s64", "userId": "08094225471505108399"}, "user_tz": -300} id="gwIf5l17h3Mg" outputId="1fca5fb8-3a9b-4a78-f726-7aef8e14ee41"
m = 10000
dataset, human_vocab, machine_vocab, inv_machine_vocab = load_dataset(m)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 16972, "status": "ok", "timestamp": 1612468514156, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gip7OjOkdNkKxKDyWEQAq1o8ccGN_HrBTGdqjgQ=s64", "userId": "08094225471505108399"}, "user_tz": -300} id="zCTqMyPch3Mg" outputId="42c9d8aa-d07b-4618-ab8a-4db4e1b971e2"
dataset[:10]
# + [markdown] id="ao4Ffrkxh3Mg"
# You've loaded:
# - `dataset`: a list of tuples of (human readable date, machine readable date).
# - `human_vocab`: a python dictionary mapping all characters used in the human readable dates to an integer-valued index.
# - `machine_vocab`: a python dictionary mapping all characters used in machine readable dates to an integer-valued index.
# - **Note**: These indices are not necessarily consistent with `human_vocab`.
# - `inv_machine_vocab`: the inverse dictionary of `machine_vocab`, mapping from indices back to characters.
#
# Let's preprocess the data and map the raw text data into the index values.
# - We will set Tx=30
# - We assume Tx is the maximum length of the human readable date.
# - If we get a longer input, we would have to truncate it.
# - We will set Ty=10
# - "YYYY-MM-DD" is 10 characters long.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 16962, "status": "ok", "timestamp": 1612468514157, "user": {"displayName": "Mubsi K", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gip7OjOkdNkKxKDyWEQAq1o8ccGN_HrBTGdqjgQ=s64", "userId": "08094225471505108399"}, "user_tz": -300} id="Qdso90EBh3Mg" outputId="0a364ad8-8b25-4de3-f036-d5d8e40bdf8c"
Tx = 30
Ty = 10
X, Y, Xoh, Yoh = preprocess_data(dataset, human_vocab, machine_vocab, Tx, Ty)
print("X.shape:", X.shape)
print("Y.shape:", Y.shape)
print("Xoh.shape:", Xoh.shape)
print("Yoh.shape:", Yoh.shape)
# + [markdown] id="q9C0UY25h3Mh"
# You now have:
# - `X`: a processed version of the human readable dates in the training set.
# - Each character in X is replaced by an index (integer) mapped to the character using `human_vocab`.
# - Each date is padded to ensure a length of $T_x$ using a special character (< pad >).
# - `X.shape = (m, Tx)` where m is the number of training examples in a batch.
# - `Y`: a processed version of the machine readable dates in the training set.
# - Each character is replaced by the index (integer) it is mapped to in `machine_vocab`.
# - `Y.shape = (m, Ty)`.
# - `Xoh`: one-hot version of `X`
# - Each index in `X` is converted to the one-hot representation (if the index is 2, the one-hot version has the index position 2 set to 1, and the remaining positions are 0.
# - `Xoh.shape = (m, Tx, len(human_vocab))`
# - `Yoh`: one-hot version of `Y`
# - Each index in `Y` is converted to the one-hot representation.
# - `Yoh.shape = (m, Ty, len(machine_vocab))`.
# - `len(machine_vocab) = 11` since there are 10 numeric digits (0 to 9) and the `-` symbol.
# + [markdown] id="N7qKvWrTh3Mh"
# * Let's also look at some examples of preprocessed training examples.
# * Feel free to play with `index` in the cell below to navigate the dataset and see how source/target dates are preprocessed.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 16952, "status": "ok", "timestamp": 1612468514158, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gip7OjOkdNkKxKDyWEQAq1o8ccGN_HrBTGdqjgQ=s64", "userId": "08094225471505108399"}, "user_tz": -300} id="kUOayR4gh3Mh" outputId="d20994de-bbea-4cc7-ffaf-38a05974c9db"
index = 0
print("Source date:", dataset[index][0])
print("Target date:", dataset[index][1])
print()
print("Source after preprocessing (indices):", X[index])
print("Target after preprocessing (indices):", Y[index])
print()
print("Source after preprocessing (one-hot):", Xoh[index])
print("Target after preprocessing (one-hot):", Yoh[index])
# + [markdown] id="94o4RYbOh3Mi"
# <a name='2'></a>
# ## 2 - Neural Machine Translation with Attention
#
# * If you had to translate a book's paragraph from French to English, you would not read the whole paragraph, then close the book and translate.
# * Even during the translation process, you would read/re-read and focus on the parts of the French paragraph corresponding to the parts of the English you are writing down.
# * The attention mechanism tells a Neural Machine Translation model where it should pay attention to at any step.
#
# <a name='2-1'></a>
# ### 2.1 - Attention Mechanism
#
# In this part, you will implement the attention mechanism presented in the lecture videos.
# * Here is a figure to remind you how the model works.
# * The diagram on the left shows the attention model.
# * The diagram on the right shows what one "attention" step does to calculate the attention variables $\alpha^{\langle t, t' \rangle}$.
# * The attention variables $\alpha^{\langle t, t' \rangle}$ are used to compute the context variable $context^{\langle t \rangle}$ for each timestep in the output ($t=1, \ldots, T_y$).
#
# <table>
# <td>
# <img src="images/attn_model.png" style="width:500;height:500px;"> <br>
# </td>
# <td>
# <img src="images/attn_mechanism.png" style="width:500;height:500px;"> <br>
# </td>
# </table>
# <caption><center> **Figure 1**: Neural machine translation with attention</center></caption>
#
# + [markdown] id="b2TkQnykh3Mi"
# Here are some properties of the model that you may notice:
#
# #### Pre-attention and Post-attention LSTMs on both sides of the attention mechanism
# - There are two separate LSTMs in this model (see diagram on the left): pre-attention and post-attention LSTMs.
# - *Pre-attention* Bi-LSTM is the one at the bottom of the picture is a Bi-directional LSTM and comes *before* the attention mechanism.
# - The attention mechanism is shown in the middle of the left-hand diagram.
# - The pre-attention Bi-LSTM goes through $T_x$ time steps
# - *Post-attention* LSTM: at the top of the diagram comes *after* the attention mechanism.
# - The post-attention LSTM goes through $T_y$ time steps.
#
# - The post-attention LSTM passes the hidden state $s^{\langle t \rangle}$ and cell state $c^{\langle t \rangle}$ from one time step to the next.
# + [markdown] id="JpznWuWqh3Mi"
# #### An LSTM has both a hidden state and cell state
# * In the lecture videos, we were using only a basic RNN for the post-attention sequence model
# * This means that the state captured by the RNN was outputting only the hidden state $s^{\langle t\rangle}$.
# * In this assignment, we are using an LSTM instead of a basic RNN.
# * So the LSTM has both the hidden state $s^{\langle t\rangle}$ and the cell state $c^{\langle t\rangle}$.
# + [markdown] id="85btUzl4h3Mj"
# #### Each time step does not use predictions from the previous time step
# * Unlike previous text generation examples earlier in the course, in this model, the post-attention LSTM at time $t$ does not take the previous time step's prediction $y^{\langle t-1 \rangle}$ as input.
# * The post-attention LSTM at time 't' only takes the hidden state $s^{\langle t\rangle}$ and cell state $c^{\langle t\rangle}$ as input.
# * We have designed the model this way because unlike language generation (where adjacent characters are highly correlated) there isn't as strong a dependency between the previous character and the next character in a YYYY-MM-DD date.
# + [markdown] id="NYT3v7rUh3Mk"
# #### Concatenation of hidden states from the forward and backward pre-attention LSTMs
# - $\overrightarrow{a}^{\langle t \rangle}$: hidden state of the forward-direction, pre-attention LSTM.
# - $\overleftarrow{a}^{\langle t \rangle}$: hidden state of the backward-direction, pre-attention LSTM.
# - $a^{\langle t \rangle} = [\overrightarrow{a}^{\langle t \rangle}, \overleftarrow{a}^{\langle t \rangle}]$: the concatenation of the activations of both the forward-direction $\overrightarrow{a}^{\langle t \rangle}$ and backward-directions $\overleftarrow{a}^{\langle t \rangle}$ of the pre-attention Bi-LSTM.
# + [markdown] id="97GUKCqwh3Mk"
# #### Computing "energies" $e^{\langle t, t' \rangle}$ as a function of $s^{\langle t-1 \rangle}$ and $a^{\langle t' \rangle}$
# - Recall in the lesson videos "Attention Model", at time 6:45 to 8:16, the definition of "e" as a function of $s^{\langle t-1 \rangle}$ and $a^{\langle t \rangle}$.
# - "e" is called the "energies" variable.
# - $s^{\langle t-1 \rangle}$ is the hidden state of the post-attention LSTM
# - $a^{\langle t' \rangle}$ is the hidden state of the pre-attention LSTM.
# - $s^{\langle t-1 \rangle}$ and $a^{\langle t \rangle}$ are fed into a simple neural network, which learns the function to output $e^{\langle t, t' \rangle}$.
# - $e^{\langle t, t' \rangle}$ is then used when computing the attention $a^{\langle t, t' \rangle}$ that $y^{\langle t \rangle}$ should pay to $a^{\langle t' \rangle}$.
# + [markdown] id="scu_HnPNh3Mk"
# - The diagram on the right of figure 1 uses a `RepeatVector` node to copy $s^{\langle t-1 \rangle}$'s value $T_x$ times.
# - Then it uses `Concatenation` to concatenate $s^{\langle t-1 \rangle}$ and $a^{\langle t \rangle}$.
# - The concatenation of $s^{\langle t-1 \rangle}$ and $a^{\langle t \rangle}$ is fed into a "Dense" layer, which computes $e^{\langle t, t' \rangle}$.
# - $e^{\langle t, t' \rangle}$ is then passed through a softmax to compute $\alpha^{\langle t, t' \rangle}$.
# - Note that the diagram doesn't explicitly show variable $e^{\langle t, t' \rangle}$, but $e^{\langle t, t' \rangle}$ is above the Dense layer and below the Softmax layer in the diagram in the right half of figure 1.
# - We'll explain how to use `RepeatVector` and `Concatenation` in Keras below.
# + [markdown] id="_ukmqe_Yh3Ml"
# #### Implementation Details
#
# Let's implement this neural translator. You will start by implementing two functions: `one_step_attention()` and `model()`.
#
# #### one_step_attention
# * The inputs to the one_step_attention at time step $t$ are:
# - $[a^{<1>},a^{<2>}, ..., a^{<T_x>}]$: all hidden states of the pre-attention Bi-LSTM.
# - $s^{<t-1>}$: the previous hidden state of the post-attention LSTM
# * one_step_attention computes:
# - $[\alpha^{<t,1>},\alpha^{<t,2>}, ..., \alpha^{<t,T_x>}]$: the attention weights
# - $context^{ \langle t \rangle }$: the context vector:
#
# $$context^{<t>} = \sum_{t' = 1}^{T_x} \alpha^{<t,t'>}a^{<t'>}\tag{1}$$
#
# ##### Clarifying 'context' and 'c'
# - In the lecture videos, the context was denoted $c^{\langle t \rangle}$
# - In the assignment, we are calling the context $context^{\langle t \rangle}$.
# - This is to avoid confusion with the post-attention LSTM's internal memory cell variable, which is also denoted $c^{\langle t \rangle}$.
# + [markdown] id="LIfLKkwoh3Ml"
# <a name='ex-1'></a>
# ### Exercise 1 - one_step_attention
#
# Implement `one_step_attention()`.
#
# * The function `model()` will call the layers in `one_step_attention()` $T_y$ times using a for-loop.
# * It is important that all $T_y$ copies have the same weights.
# * It should not reinitialize the weights every time.
# * In other words, all $T_y$ steps should have shared weights.
# * Here's how you can implement layers with shareable weights in Keras:
# 1. Define the layer objects in a variable scope that is outside of the `one_step_attention` function. For example, defining the objects as global variables would work.
# - Note that defining these variables inside the scope of the function `model` would technically work, since `model` will then call the `one_step_attention` function. For the purposes of making grading and troubleshooting easier, we are defining these as global variables. Note that the automatic grader will expect these to be global variables as well.
# 2. Call these objects when propagating the input.
# * We have defined the layers you need as global variables.
# * Please run the following cells to create them.
# * Please note that the automatic grader expects these global variables with the given variable names. For grading purposes, please do not rename the global variables.
# * Please check the Keras documentation to learn more about these layers. The layers are functions. Below are examples of how to call these functions.
# * [RepeatVector()](https://www.tensorflow.org/api_docs/python/tf/keras/layers/RepeatVector)
# ```Python
# var_repeated = repeat_layer(var1)
# ```
# * [Concatenate()](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Concatenate)
# ```Python
# concatenated_vars = concatenate_layer([var1,var2,var3])
# ```
# * [Dense()](https://keras.io/layers/core/#dense)
# ```Python
# var_out = dense_layer(var_in)
# ```
# * [Activation()](https://keras.io/layers/core/#activation)
# ```Python
# activation = activation_layer(var_in)
# ```
# * [Dot()](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Dot)
# ```Python
# dot_product = dot_layer([var1,var2])
# ```
# + executionInfo={"elapsed": 16950, "status": "ok", "timestamp": 1612468514158, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gip7OjOkdNkKxKDyWEQAq1o8ccGN_HrBTGdqjgQ=s64", "userId": "08094225471505108399"}, "user_tz": -300} id="Cvop5Apyh3Mm"
# Defined shared layers as global variables
repeator = RepeatVector(Tx)
concatenator = Concatenate(axis=-1)
densor1 = Dense(10, activation="tanh")
densor2 = Dense(1, activation="relu")
activator = Activation(softmax, name='attention_weights') # We are using a custom softmax(axis = 1) loaded in this notebook
dotor = Dot(axes=1)
# + executionInfo={"elapsed": 16950, "status": "ok", "timestamp": 1612468514159, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gip7OjOkdNkKxKDyWEQAq1o8ccGN_HrBTGdqjgQ=s64", "userId": "08094225471505108399"}, "user_tz": -300} id="mZuMOnTDh3Mn"
# UNQ_C1 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: one_step_attention
def one_step_attention(a, s_prev):
"""
Performs one step of attention: Outputs a context vector computed as a dot product of the attention weights
"alphas" and the hidden states "a" of the Bi-LSTM.
Arguments:
a -- hidden state output of the Bi-LSTM, numpy-array of shape (m, Tx, 2*n_a)
s_prev -- previous hidden state of the (post-attention) LSTM, numpy-array of shape (m, n_s)
Returns:
context -- context vector, input of the next (post-attention) LSTM cell
"""
### START CODE HERE ###
# Use repeator to repeat s_prev to be of shape (m, Tx, n_s) so that you can concatenate it with all hidden states "a" (≈ 1 line)
s_prev = repeator(s_prev)
# Use concatenator to concatenate a and s_prev on the last axis (≈ 1 line)
# For grading purposes, please list 'a' first and 's_prev' second, in this order.
concat = concatenator([a, s_prev])
# Use densor1 to propagate concat through a small fully-connected neural network to compute the "intermediate energies" variable e. (≈1 lines)
e = densor1(concat)
# Use densor2 to propagate e through a small fully-connected neural network to compute the "energies" variable energies. (≈1 lines)
energies = densor2(e)
# Use "activator" on "energies" to compute the attention weights "alphas" (≈ 1 line)
alphas = activator(energies)
# Use dotor together with "alphas" and "a", in this order, to compute the context vector to be given to the next (post-attention) LSTM-cell (≈ 1 line)
context = dotor([alphas, a])
### END CODE HERE ###
return context
# +
# UNIT TEST
def one_step_attention_test(target):
m = 10
Tx = 30
n_a = 32
n_s = 64
#np.random.seed(10)
a = np.random.uniform(1, 0, (m, Tx, 2 * n_a)).astype(np.float32)
s_prev =np.random.uniform(1, 0, (m, n_s)).astype(np.float32) * 1
context = target(a, s_prev)
assert type(context) == tf.python.framework.ops.EagerTensor, "Unexpected type. It should be a Tensor"
assert tuple(context.shape) == (m, 1, n_s), "Unexpected output shape"
assert np.all(context.numpy() > 0), "All output values must be > 0 in this example"
assert np.all(context.numpy() < 1), "All output values must be < 1 in this example"
#assert np.allclose(context[0][0][0:5].numpy(), [0.50877404, 0.57160693, 0.45448175, 0.50074816, 0.53651875]), "Unexpected values in the result"
print("\033[92mAll tests passed!")
one_step_attention_test(one_step_attention)
# + [markdown] id="vcmC3WcQh3Mn"
# <a name='ex-2'></a>
# ### Exercise 2 - modelf
#
# Implement `modelf()` as explained in figure 1 and the instructions:
#
# * `modelf` first runs the input through a Bi-LSTM to get $[a^{<1>},a^{<2>}, ..., a^{<T_x>}]$.
# * Then, `modelf` calls `one_step_attention()` $T_y$ times using a `for` loop. At each iteration of this loop:
# - It gives the computed context vector $context^{<t>}$ to the post-attention LSTM.
# - It runs the output of the post-attention LSTM through a dense layer with softmax activation.
# - The softmax generates a prediction $\hat{y}^{<t>}$.
#
# Again, we have defined global layers that will share weights to be used in `modelf()`.
# + executionInfo={"elapsed": 16949, "status": "ok", "timestamp": 1612468514159, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gip7OjOkdNkKxKDyWEQAq1o8ccGN_HrBTGdqjgQ=s64", "userId": "08094225471505108399"}, "user_tz": -300} id="5RHgmZrVh3Mo"
n_a = 32 # number of units for the pre-attention, bi-directional LSTM's hidden state 'a'
n_s = 64 # number of units for the post-attention LSTM's hidden state "s"
# Please note, this is the post attention LSTM cell.
post_activation_LSTM_cell = LSTM(n_s, return_state=True) # Please do not modify this global variable.
output_layer = Dense(len(machine_vocab), activation=softmax)
# + [markdown] id="lGkKpb1Nh3Mo"
# Now you can use these layers $T_y$ times in a `for` loop to generate the outputs, and their parameters will not be reinitialized. You will have to carry out the following steps:
#
# 1. Propagate the input `X` into a bi-directional LSTM.
# * [Bidirectional](https://keras.io/layers/wrappers/#bidirectional)
# * [LSTM](https://keras.io/layers/recurrent/#lstm)
# * Remember that we want the LSTM to return a full sequence instead of just the last hidden state.
#
# Sample code:
#
# ```Python
# sequence_of_hidden_states = Bidirectional(LSTM(units=..., return_sequences=...))(the_input_X)
# ```
#
# 2. Iterate for $t = 0, \cdots, T_y-1$:
# 1. Call `one_step_attention()`, passing in the sequence of hidden states $[a^{\langle 1 \rangle},a^{\langle 2 \rangle}, ..., a^{ \langle T_x \rangle}]$ from the pre-attention bi-directional LSTM, and the previous hidden state $s^{<t-1>}$ from the post-attention LSTM to calculate the context vector $context^{<t>}$.
# 2. Give $context^{<t>}$ to the post-attention LSTM cell.
# - Remember to pass in the previous hidden-state $s^{\langle t-1\rangle}$ and cell-states $c^{\langle t-1\rangle}$ of this LSTM
# * This outputs the new hidden state $s^{<t>}$ and the new cell state $c^{<t>}$.
#
# Sample code:
# ```Python
# next_hidden_state, _ , next_cell_state =
# post_activation_LSTM_cell(inputs=..., initial_state=[prev_hidden_state, prev_cell_state])
# ```
# Please note that the layer is actually the "post attention LSTM cell". For the purposes of passing the automatic grader, please do not modify the naming of this global variable. This will be fixed when we deploy updates to the automatic grader.
# 3. Apply a dense, softmax layer to $s^{<t>}$, get the output.
# Sample code:
# ```Python
# output = output_layer(inputs=...)
# ```
# 4. Save the output by adding it to the list of outputs.
#
# 3. Create your Keras model instance.
# * It should have three inputs:
# * `X`, the one-hot encoded inputs to the model, of shape ($T_{x}, humanVocabSize)$
# * $s^{\langle 0 \rangle}$, the initial hidden state of the post-attention LSTM
# * $c^{\langle 0 \rangle}$, the initial cell state of the post-attention LSTM
# * The output is the list of outputs.
# Sample code
# ```Python
# model = Model(inputs=[...,...,...], outputs=...)
# ```
# + executionInfo={"elapsed": 16948, "status": "ok", "timestamp": 1612468514160, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gip7OjOkdNkKxKDyWEQAq1o8ccGN_HrBTGdqjgQ=s64", "userId": "08094225471505108399"}, "user_tz": -300} id="qeKbeDOvh3Mo"
# UNQ_C2 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: model
def modelf(Tx, Ty, n_a, n_s, human_vocab_size, machine_vocab_size):
"""
Arguments:
Tx -- length of the input sequence
Ty -- length of the output sequence
n_a -- hidden state size of the Bi-LSTM
n_s -- hidden state size of the post-attention LSTM
human_vocab_size -- size of the python dictionary "human_vocab"
machine_vocab_size -- size of the python dictionary "machine_vocab"
Returns:
model -- Keras model instance
"""
# Define the inputs of your model with a shape (Tx,)
# Define s0 (initial hidden state) and c0 (initial cell state)
# for the decoder LSTM with shape (n_s,)
X = Input(shape=(Tx, human_vocab_size))
s0 = Input(shape=(n_s,), name='s0')
c0 = Input(shape=(n_s,), name='c0')
s = s0
c = c0
# Initialize empty list of outputs
outputs = []
### START CODE HERE ###
# Step 1: Define your pre-attention Bi-LSTM. (≈ 1 line)
a = Bidirectional(LSTM(units=n_a, return_sequences=True))(X)
# Step 2: Iterate for Ty steps
for t in range(Ty):
# Step 2.A: Perform one step of the attention mechanism to get back the context vector at step t (≈ 1 line)
context = one_step_attention(a, s)
# Step 2.B: Apply the post-attention LSTM cell to the "context" vector.
# Don't forget to pass: initial_state = [hidden state, cell state] (≈ 1 line)
s, _, c = post_activation_LSTM_cell(inputs=context, initial_state=[s, c])
# Step 2.C: Apply Dense layer to the hidden state output of the post-attention LSTM (≈ 1 line)
out = output_layer(inputs=s)
# Step 2.D: Append "out" to the "outputs" list (≈ 1 line)
outputs.append(out)
# Step 3: Create model instance taking three inputs and returning the list of outputs. (≈ 1 line)
model = Model(inputs=[X, s0, c0], outputs=outputs)
### END CODE HERE ###
return model
# +
# UNIT TEST
from test_utils import *
def modelf_test(target):
m = 10
Tx = 30
n_a = 32
n_s = 64
len_human_vocab = 37
len_machine_vocab = 11
model = target(Tx, Ty, n_a, n_s, len_human_vocab, len_machine_vocab)
print(summary(model))
expected_summary = [['InputLayer', [(None, 30, 37)], 0],
['InputLayer', [(None, 64)], 0],
['Bidirectional', (None, 30, 64), 17920],
['RepeatVector', (None, 30, 64), 0, 30],
['Concatenate', (None, 30, 128), 0],
['Dense', (None, 30, 10), 1290, 'tanh'],
['Dense', (None, 30, 1), 11, 'relu'],
['Activation', (None, 30, 1), 0],
['Dot', (None, 1, 64), 0],
['InputLayer', [(None, 64)], 0],
['LSTM',[(None, 64), (None, 64), (None, 64)], 33024, [(None, 1, 64), (None, 64), (None, 64)],'tanh'],
['Dense', (None, 11), 715, 'softmax']]
assert len(model.outputs) == 10, f"Wrong output shape. Expected 10 != {len(model.outputs)}"
comparator(summary(model), expected_summary)
modelf_test(modelf)
# + [markdown] id="--RX7hSsh3Mo"
# Run the following cell to create your model.
# + executionInfo={"elapsed": 20837, "status": "ok", "timestamp": 1612468518050, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gip7OjOkdNkKxKDyWEQAq1o8ccGN_HrBTGdqjgQ=s64", "userId": "08094225471505108399"}, "user_tz": -300} id="psdd-Ac6h3Mp"
model = modelf(Tx, Ty, n_a, n_s, len(human_vocab), len(machine_vocab))
# + [markdown] id="nUJw7Xohh3Mp"
# #### Troubleshooting Note
# * If you are getting repeated errors after an initially incorrect implementation of "model", but believe that you have corrected the error, you may still see error messages when building your model.
# * A solution is to save and restart your kernel (or shutdown then restart your notebook), and re-run the cells.
# + [markdown] id="VgeU_I9_h3Mp"
# Let's get a summary of the model to check if it matches the expected output.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 20835, "status": "ok", "timestamp": 1612468518050, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gip7OjOkdNkKxKDyWEQAq1o8ccGN_HrBTGdqjgQ=s64", "userId": "08094225471505108399"}, "user_tz": -300} id="tX0vaYmPh3Mq" outputId="336b9248-70b0-4379-be95-95366874c02a"
model.summary()
# + [markdown] id="uiqCePt5h3Mr"
# **Expected Output**:
#
# Here is the summary you should see
# <table>
# <tr>
# <td>
# **Total params:**
# </td>
# <td>
# 52,960
# </td>
# </tr>
# <tr>
# <td>
# **Trainable params:**
# </td>
# <td>
# 52,960
# </td>
# </tr>
# <tr>
# <td>
# **Non-trainable params:**
# </td>
# <td>
# 0
# </td>
# </tr>
# <tr>
# <td>
# **bidirectional_1's output shape **
# </td>
# <td>
# (None, 30, 64)
# </td>
# </tr>
# <tr>
# <td>
# **repeat_vector_1's output shape **
# </td>
# <td>
# (None, 30, 64)
# </td>
# </tr>
# <tr>
# <td>
# **concatenate_1's output shape **
# </td>
# <td>
# (None, 30, 128)
# </td>
# </tr>
# <tr>
# <td>
# **attention_weights's output shape **
# </td>
# <td>
# (None, 30, 1)
# </td>
# </tr>
# <tr>
# <td>
# **dot_1's output shape **
# </td>
# <td>
# (None, 1, 64)
# </td>
# </tr>
# <tr>
# <td>
# **dense_3's output shape **
# </td>
# <td>
# (None, 11)
# </td>
# </tr>
# </table>
#
# + [markdown] id="8u3D9Odhh3Ms"
# <a name='ex-3'></a>
# ### Exercise 3 - Compile the Model
#
# * After creating your model in Keras, you need to compile it and define the loss function, optimizer and metrics you want to use.
# * Loss function: 'categorical_crossentropy'.
# * Optimizer: [Adam](https://keras.io/optimizers/#adam) [optimizer](https://keras.io/optimizers/#usage-of-optimizers)
# - learning rate = 0.005
# - $\beta_1 = 0.9$
# - $\beta_2 = 0.999$
# - decay = 0.01
# * metric: 'accuracy'
#
# Sample code
# ```Python
# optimizer = Adam(lr=..., beta_1=..., beta_2=..., decay=...)
# model.compile(optimizer=..., loss=..., metrics=[...])
# ```
# + executionInfo={"elapsed": 20835, "status": "ok", "timestamp": 1612468518051, "user": {"displayName": "Mubsi K", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gip7OjOkdNkKxKDyWEQAq1o8ccGN_HrBTGdqjgQ=s64", "userId": "08094225471505108399"}, "user_tz": -300} id="sBFRJ49rh3Ms"
### START CODE HERE ### (≈2 lines)
opt = Adam(lr=0.005, beta_1=0.9, beta_2=0.999, decay=0.01) # Adam(...)
model.compile(loss='categorical_crossentropy', optimizer=opt, metrics=['accuracy'])
### END CODE HERE ###
# +
# UNIT TESTS
assert opt.lr == 0.005, "Set the lr parameter to 0.005"
assert opt.beta_1 == 0.9, "Set the beta_1 parameter to 0.9"
assert opt.beta_2 == 0.999, "Set the beta_2 parameter to 0.999"
assert opt.decay == 0.01, "Set the decay parameter to 0.01"
assert model.loss == "categorical_crossentropy", "Wrong loss. Use 'categorical_crossentropy'"
assert model.optimizer == opt, "Use the optimizer that you have instantiated"
assert model.compiled_metrics._user_metrics[0] == 'accuracy', "set metrics to ['accuracy']"
print("\033[92mAll tests passed!")
# + [markdown] id="Qz71nM3oh3Ms"
# #### Define inputs and outputs, and fit the model
# The last step is to define all your inputs and outputs to fit the model:
# - You have input `Xoh` of shape $(m = 10000, T_x = 30, human\_vocab=37)$ containing the training examples.
# - You need to create `s0` and `c0` to initialize your `post_attention_LSTM_cell` with zeros.
# - Given the `model()` you coded, you need the "outputs" to be a list of 10 elements of shape (m, T_y).
# - The list `outputs[i][0], ..., outputs[i][Ty]` represents the true labels (characters) corresponding to the $i^{th}$ training example (`Xoh[i]`).
# - `outputs[i][j]` is the true label of the $j^{th}$ character in the $i^{th}$ training example.
# + executionInfo={"elapsed": 20833, "status": "ok", "timestamp": 1612468518051, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gip7OjOkdNkKxKDyWEQAq1o8ccGN_HrBTGdqjgQ=s64", "userId": "08094225471505108399"}, "user_tz": -300} id="USFiNKYhh3Mt"
s0 = np.zeros((m, n_s))
c0 = np.zeros((m, n_s))
outputs = list(Yoh.swapaxes(0,1))
# + [markdown] id="FVkITGi3h3Mt"
# Let's now fit the model and run it for one epoch.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 47944, "status": "ok", "timestamp": 1612468545172, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gip7OjOkdNkKxKDyWEQAq1o8ccGN_HrBTGdqjgQ=s64", "userId": "08094225471505108399"}, "user_tz": -300} id="tPuwY45bh3Mt" outputId="ec9dfc4c-1dcb-4577-d872-474f79c60d5f"
model.fit([Xoh, s0, c0], outputs, epochs=1, batch_size=100)
# + [markdown] id="SUikskCoh3Mt"
# While training you can see the loss as well as the accuracy on each of the 10 positions of the output. The table below gives you an example of what the accuracies could be if the batch had 2 examples:
#
# <img src="images/table.png" style="width:700;height:200px;"> <br>
# <caption><center>Thus, `dense_2_acc_8: 0.89` means that you are predicting the 7th character of the output correctly 89% of the time in the current batch of data. </center></caption>
#
#
# We have run this model for longer, and saved the weights. Run the next cell to load our weights. (By training a model for several minutes, you should be able to obtain a model of similar accuracy, but loading our model will save you time.)
# + executionInfo={"elapsed": 47942, "status": "ok", "timestamp": 1612468545173, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gip7OjOkdNkKxKDyWEQAq1o8ccGN_HrBTGdqjgQ=s64", "userId": "08094225471505108399"}, "user_tz": -300} id="ooiZCOx0h3Mu"
model.load_weights('models/model.h5')
# + [markdown] id="yUUD9yXxh3Mu"
# You can now see the results on new examples.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 53835, "status": "ok", "timestamp": 1612468551077, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gip7OjOkdNkKxKDyWEQAq1o8ccGN_HrBTGdqjgQ=s64", "userId": "08094225471505108399"}, "user_tz": -300} id="rQ8sd_cuh3Mv" outputId="c37e92ac-5c60-4caf-b843-6aaeaa37be25"
EXAMPLES = ['3 May 1979', '5 April 09', '21th of August 2016', 'Tue 10 Jul 2007', 'Saturday May 9 2018', 'March 3 2001', 'March 3rd 2001', '1 March 2001']
s00 = np.zeros((1, n_s))
c00 = np.zeros((1, n_s))
for example in EXAMPLES:
source = string_to_int(example, Tx, human_vocab)
#print(source)
source = np.array(list(map(lambda x: to_categorical(x, num_classes=len(human_vocab)), source))).swapaxes(0,1)
source = np.swapaxes(source, 0, 1)
source = np.expand_dims(source, axis=0)
prediction = model.predict([source, s00, c00])
prediction = np.argmax(prediction, axis = -1)
output = [inv_machine_vocab[int(i)] for i in prediction]
print("source:", example)
print("output:", ''.join(output),"\n")
# + [markdown] id="vjdEQiIDh3Mv"
# You can also change these examples to test with your own examples. The next part will give you a better sense of what the attention mechanism is doing--i.e., what part of the input the network is paying attention to when generating a particular output character.
# + [markdown] id="1XIxtN4xh3Mv"
# <a name='3'></a>
# ## 3 - Visualizing Attention (Optional / Ungraded)
#
# Since the problem has a fixed output length of 10, it is also possible to carry out this task using 10 different softmax units to generate the 10 characters of the output. But one advantage of the attention model is that each part of the output (such as the month) knows it needs to depend only on a small part of the input (the characters in the input giving the month). We can visualize what each part of the output is looking at which part of the input.
#
# Consider the task of translating "Saturday 9 May 2018" to "2018-05-09". If we visualize the computed $\alpha^{\langle t, t' \rangle}$ we get this:
#
# <img src="images/date_attention.png" style="width:600;height:300px;"> <br>
# <caption><center> **Figure 8**: Full Attention Map</center></caption>
#
# Notice how the output ignores the "Saturday" portion of the input. None of the output timesteps are paying much attention to that portion of the input. We also see that 9 has been translated as 09 and May has been correctly translated into 05, with the output paying attention to the parts of the input it needs to to make the translation. The year mostly requires it to pay attention to the input's "18" in order to generate "2018."
# + [markdown] id="FrP893IFh3Mv"
# <a name='3-1'></a>
# ### 3.1 - Getting the Attention Weights From the Network
#
# Lets now visualize the attention values in your network. We'll propagate an example through the network, then visualize the values of $\alpha^{\langle t, t' \rangle}$.
#
# To figure out where the attention values are located, let's start by printing a summary of the model .
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 53826, "status": "ok", "timestamp": 1612468551078, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gip7OjOkdNkKxKDyWEQAq1o8ccGN_HrBTGdqjgQ=s64", "userId": "08094225471505108399"}, "user_tz": -300} id="RfiLrfKIh3Mv" outputId="b6690603-209c-40d7-f352-235a689d1aea"
model.summary()
# + [markdown] id="zbcprBCPh3Mv"
# Navigate through the output of `model.summary()` above. You can see that the layer named `attention_weights` outputs the `alphas` of shape (m, 30, 1) before `dot_2` computes the context vector for every time step $t = 0, \ldots, T_y-1$. Let's get the attention weights from this layer.
#
# The function `attention_map()` pulls out the attention values from your model and plots them.
#
# **Note**: We are aware that you might run into an error running the cell below despite a valid implementation for Exercise 2 - `modelf` above. If you get the error kindly report it on this [Topic](https://discourse.deeplearning.ai/t/error-in-optional-ungraded-part-of-neural-machine-translation-w3a1/1096) on [Discourse](https://discourse.deeplearning.ai) as it'll help us improve our content.
#
# If you haven’t joined our Discourse community you can do so by clicking on the link: http://bit.ly/dls-discourse
#
# And don’t worry about the error, it will not affect the grading for this assignment.
# -
attention_map = plot_attention_map(model, human_vocab, inv_machine_vocab, "Tuesday 09 Oct 1993", num = 7, n_s = 64);
# + [markdown] id="pQ3qbIjqh3Mx"
# On the generated plot you can observe the values of the attention weights for each character of the predicted output. Examine this plot and check that the places where the network is paying attention makes sense to you.
#
# In the date translation application, you will observe that most of the time attention helps predict the year, and doesn't have much impact on predicting the day or month.
# + [markdown] id="IkpGu1Jkh3Mx"
# ### Congratulations!
#
#
# You have come to the end of this assignment
#
# #### Here's what you should remember
#
# - Machine translation models can be used to map from one sequence to another. They are useful not just for translating human languages (like French->English) but also for tasks like date format translation.
# - An attention mechanism allows a network to focus on the most relevant parts of the input when producing a specific part of the output.
# - A network using an attention mechanism can translate from inputs of length $T_x$ to outputs of length $T_y$, where $T_x$ and $T_y$ can be different.
# - You can visualize attention weights $\alpha^{\langle t,t' \rangle}$ to see what the network is paying attention to while generating each output.
# + [markdown] id="ZaKA2u4uh3My"
# Congratulations on finishing this assignment! You are now able to implement an attention model and use it to learn complex mappings from one sequence to another.
| Deep Learning Specialization/Sequence Models/Week 3/Neural_machine_translation_with_attention_v4a.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# +
import matplotlib.pyplot as plt
# Prepare the data
m = 51212.
f = 40742.
m_perc = m/(m+f)
f_perc = f/(m+f)
# Set the colors and labels
colors = ['navy','lightcoral']
labels = ["Male","Female"]
plt.figure(figsize=(12,12))
patches, texts, autotexts = plt.pie([m_perc,f_perc],labels=labels,autopct='%1.1f%%',\
explode=[0,0.05], startangle=0, colors=colors)
plt.title('Median Annual Earnings Ratio for Full-Time,\
Year-Round Workers in the US by Gender in 2015',fontsize=20)
# Format the text labels
for text in texts+autotexts:
text.set_fontsize(20)
text.set_fontweight('bold')
for text in autotexts:
text.set_color('white')
# Save the figure. Commented to save running time.
# plt.savefig('Section_2_annualearnings_pie.png',dpi=600)
plt.show()
# +
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
# Prepare the data from Figure 2b
saline_social = [123,96,111,85,74,115,74,91]
saline_opposite = [24,12,20,25,53,19,29,28]
ptz_social = [39,20,32,43,61,82,46,64,57,71]
ptz_opposite = [40,70,40,71,24,8,27,30,29,13]
n = 2
xloc = np.arange(2)
y1 = [np.mean(saline_social),np.mean(ptz_social)]
y2 = [np.mean(saline_opposite),np.mean(ptz_opposite)]
# Filling zeroes to draw positive error bars only
yerr1 = [(0,0),[np.std(saline_social),np.std(saline_opposite)]]
yerr2 = [(0,0),[np.std(ptz_social),np.std(ptz_opposite)]]
# Set the figure
fig = plt.figure(figsize=(3,5))
# Set the bar widths and locations
width = 0.3
# Draw the bar chart
plt.bar(xloc, y1, width=width, align='center', color='black', label='Social')
plt.bar(xloc+width, y2, width=width, edgecolor='black', facecolor='white',\
align='center', linewidth=3, label='Opposite')
# Format errorbars. More details will be covered in scientific plotting chapter.
plt.errorbar(xloc, y1, yerr=yerr1, ls='none', lw=3, color='black',\
capsize=8, capthick=3, xlolims=True)
plt.errorbar(xloc+width, y2, yerr=yerr2, ls='none', lw=3, color='black',\
capsize=8, capthick=3,xlolims=False)
# Get current axes and store it to ax
ax = plt.gca()
# Format the spines
ax.spines['left'].set_linewidth(3)
ax.spines['bottom'].set_linewidth(3)
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
# Format the axes
ax.xaxis.set_tick_params(width=3)
ax.yaxis.set_tick_params(width=3)
ax.yaxis.set_major_locator(ticker.MultipleLocator(50))
labels = ['Saline','PTZ']
plt.xticks(range(2), labels, fontsize=16, rotation=30)
plt.yticks(fontsize=16)
plt.ylabel('Sniffing time (s)', fontsize=18)
plt.ylim(0,150)
# Add a legend
plt.legend(loc='upper center',fontsize=18,frameon=False)
# Show the figure
plt.show()
# -
# +
#Adding Subplots
# +
import numpy as np
import matplotlib.pyplot as plt
# Prepare the data
y = np.arange(200)
# Draw the plots
# plt.subplot(nrows, ncols,plot_number)
ax1 = plt.subplot(121) # plt.figure(1) Create the first subplot (on left)
plt.plot(-y) # Plot on ax1
ax2 = plt.subplot(122) # Create the second subplot
plt.plot(y) # Plot on ax2
# Show the figure
plt.show()
# +
import matplotlib.pyplot as plt
import numpy as np
# Draw 2x2 subplots and assign them to axes variables
fig, axarr = plt.subplots(2,2) #fig = plt.figure(); ax =fig.add_subplot(111)
# Prepare data
x1 = np.linspace(5,5)
x2 = np.linspace(0,10)
y1 = x2
y2 = 10-x2
# Plot the data to respective subplots
axarr[0,0].plot(x2,y1)
axarr[0,1].plot(x2,y2)
axarr[1,0].plot(x1,y2)
axarr[1,1].plot(x1,y1)
# Show the figure
plt.show()
# +
import matplotlib.pyplot as plt
import numpy as np
# Draw 2x2 subplots and assign them to axes variables
fig, axarr = plt.subplots(2,2,sharex=True,sharey=True)
# Prepare data
x1 = np.linspace(5,5)
x2 = np.linspace(0,10)
y1 = x2
y2 = 10-x2
# Plot the data to respective subplots
axarr[0,0].plot(x2,y1)
axarr[0,1].plot(x2,y2)
axarr[1,0].plot(x1,y2)
axarr[1,1].plot(x1,y1)
# Show the figure
plt.show()
# -
# +
# Adjusting margins
# -
plt.figure()
ax = fig.add_axes([left, bottom, width, height])
fig, ax = plt.subplots()
plt.subplots_adjust(left=0.1, right=0.9, top=0.9, bottom=0.1)
# +
import matplotlib.pyplot as plt
import numpy as np
# Draw 2x2 subplots and assign them to axes variables
fig, axarr = plt.subplots(2,2,sharex=True,sharey=True)
# Prepare data
x1 = np.linspace(5,5)
x2 = np.linspace(0,10)
y1 = x2
y2 = 10-x2
# Plot the data to respective subplots
axarr[0,0].plot(x2,y1)
axarr[0,1].plot(x2,y2)
axarr[1,0].plot(x1,y2)
axarr[1,1].plot(x1,y1)
# Show the figure
plt.tight_layout(pad=0, w_pad=-1, h_pad=-1)
# -
ax1 = plt.subplot2grid((3, 3), (0, 0))
ax2 = plt.subplot2grid((3, 3), (1, 0))
ax3 = plt.subplot2grid((3, 3), (0, 2), rowspan=3)
ax4 = plt.subplot2grid((3, 3), (2, 0), colspan=2)
ax5 = plt.subplot2grid((3, 3), (0, 1), rowspan=2)
# +
#Drawing inset plots
# +
import matplotlib.pyplot as plt
import numpy as np
x = np.linspace(0,10,1000)
y2 = np.sin(x**2)
y1 = x**2
# Initiate a figure with subplot axes
fig, ax1 = plt.subplots()
# Set the inset plot dimensions
left, bottom, width, height = [0.22, 0.45, 0.3, 0.35]
ax2 = fig.add_axes([left, bottom, width, height])
# Draw the plots
ax1.plot(x,y1)
ax2.plot(x,y2)
# Show the figure
plt.show()
# -
from mpl_toolkits.axes_grid1.inset_locator import inset_axes
ax2 = inset_axes(ax1, width=6.5, height=45%, loc=5)
# +
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1.inset_locator import inset_axes
# Prepare the data
top10_arrivals_countries = ['CANADA','MEXICO','UNITED\nKINGDOM',\
'JAPAN','CHINA','GERMANY','SOUTH\nKOREA',\
'FRANCE','BRAZIL','AUSTRALIA']
top10_arrivals_values = [16.625687, 15.378026, 3.934508, 2.999718,\
2.618737, 1.769498, 1.628563, 1.419409,\
1.393710, 1.136974]
arrivals_countries = ['WESTERN\nEUROPE','ASIA','SOUTH\nAMERICA',\
'OCEANIA','CARIBBEAN','MIDDLE\nEAST',\
'CENTRAL\nAMERICA','EASTERN\nEUROPE','AFRICA']
arrivals_percent = [36.9,30.4,13.8,4.4,4.0,3.6,2.9,2.6,1.5]
# Set up the figure and the main subplot
fig, ax1 = plt.subplots(figsize=(20,12))
# Draw the bar plot
rects1 = ax1.bar(range(10),top10_arrivals_values, align='center',color='#3b5998')
# Set spines to be invisible
for spine in ax1.spines.values():
spine.set_visible(False)
# Format ticks and labels
plt.xticks(range(10),top10_arrivals_countries,fontsize=18)
for tic in ax1.xaxis.get_major_ticks():
tic.tick1On = tic.tick2On = False
plt.yticks(fontsize=18)
plt.xlabel('Top 10 tourist generating countries',fontsize=24,fontweight='semibold')
plt.ylabel('Arrivals (Million)',fontsize=24,fontweight='semibold')
# Prepare the inset axes
ax2 = inset_axes(ax1, width=6.5, height=6.5, loc=5)
# Store the pie chart sectors, sample labels and value labels
# Set the properties
explode = (0.08, 0.08, 0.05, 0.05,0.05,0.05,0.05,0.05,0.05)
patches, texts, autotexts = ax2.pie(arrivals_percent, \
labels=arrivals_countries,\
autopct='%1.1f%%', \
shadow=True, startangle=180,\
explode=explode, \
counterclock=False, \
pctdistance=0.72)
# Set properties of text in pie chart
for text in texts+autotexts:
text.set_fontsize(16)
text.set_fontweight('semibold')
# Add a super title to all the subplots
plt.suptitle('Non-Resident Arrivals to the US in 2016 (by Aug) by Regions',\
fontsize=36,color='navy',fontweight='bold')
# Show the figure
plt.show()
# -
# +
# Adding text annotations
# +
import matplotlib.pyplot as plt
import numpy as np
# create 1000 equally spaced points between -10 and 10
x = np.linspace(0, 10)
# Prepare the data
y1 = x
y2 = 10-x
# Plot the data
fig, ax = plt.subplots()
plt.plot(x,y1,label='Supply')
plt.plot(x,y2,label='Demand')
# Annotate the equilibrium point with arrow and text
ax.annotate("Equilibrium", xy=(5,5), xytext=(4,2), \
fontsize=12, fontweight='semibold',\
arrowprops=dict(linewidth=2, arrowstyle="->"))
# Label the axes
plt.xlabel('Quantity',fontsize=12,fontweight='semibold')
plt.ylabel('Price',fontsize=12,fontweight='semibold')
# Style the plot to a common demand-supply graph
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.xaxis.set_major_locator(plt.NullLocator())
ax.yaxis.set_major_locator(plt.NullLocator())
plt.legend()
plt.show()
# +
import matplotlib.pyplot as plt
import numpy as np
# create 1000 equally spaced points between -10 and 10
x = np.linspace(0, 10)
# Prepare the data
y1 = x
y2 = 10-x
# Plot the data
fig, ax = plt.subplots()
plt.plot(x,y1,label='Supply')
plt.plot(x,y2,label='Demand')
# Annotate the equilibrium point with arrow and text
ax.annotate("Equilibrium", xy=(5,5), xytext=(4,2), \
fontsize=12, fontweight='semibold',\
arrowprops=dict(linewidth=2, arrowstyle="->"))
# Label the axes
plt.xlabel('Quantity',fontsize=12,fontweight='semibold')
plt.ylabel('Price',fontsize=12,fontweight='semibold')
# Style the plot to a common demand-supply graph
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.xaxis.set_major_locator(plt.NullLocator())
ax.yaxis.set_major_locator(plt.NullLocator())
ax.text(9, 9.6, "Supply", ha="center", va="center", size=16, rotation=33,color='C0')
ax.text(9, 1.5, "Demand", ha="center", va="center", size=16, rotation=-33,color='C1')
plt.show()
# +
import matplotlib.pyplot as plt
import numpy as np
# create 1000 equally spaced points between -10 and 10
x = np.linspace(0, 10)
# Prepare the data
y1 = x
y2 = 10-x
# Plot the data
fig, ax = plt.subplots()
plt.plot(x,y1,label='Supply')
plt.plot(x,y2,label='Demand')
# Annotate the equilibrium point with arrow and text
bbox_props = dict(boxstyle="rarrow", fc=(0.8, 0.9, 0.9), ec="b", lw=2)
t = ax.text(2,5, "Equilibrium", ha="center", va="center", rotation=0,
size=12,bbox=bbox_props)
bb = t.get_bbox_patch()
bb.set_boxstyle("rarrow", pad=0.6)
# Label the axes
plt.xlabel('Quantity',fontsize=12,fontweight='semibold')
plt.ylabel('Price',fontsize=12,fontweight='semibold')
# Style the plot to a common demand-supply graph
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.xaxis.set_major_locator(plt.NullLocator())
ax.yaxis.set_major_locator(plt.NullLocator())
ax.text(9, 9.6, "Supply", ha="center", va="center", size=16, rotation=33,color='C0')
ax.text(9, 1.5, "Demand", ha="center", va="center", size=16, rotation=-33,color='C1')
plt.show()
# +
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1.inset_locator import inset_axes
# Prepare the bar value labelling function
def autolabel(rects):
#Attach a text label above each bar displaying its height
for rect in rects:
height = rect.get_height()
ax1.text(rect.get_x() + rect.get_width()/2., 1.02*height,
"{:,}".format(float(height)),
ha='center', va='bottom',fontsize=18)
# Prepare the data
top10_arrivals_countries = ['CANADA','MEXICO','UNITED\nKINGDOM',\
'JAPAN','CHINA','GERMANY','SOUTH\nKOREA',\
'FRANCE','BRAZIL','AUSTRALIA']
top10_arrivals_values = [16.625687, 15.378026, 3.934508, 2.999718,\
2.618737, 1.769498, 1.628563, 1.419409,\
1.393710, 1.136974]
arrivals_countries = ['WESTERN\nEUROPE','ASIA','SOUTH\nAMERICA',\
'OCEANIA','CARIBBEAN','MIDDLE\nEAST',\
'CENTRAL\nAMERICA','EASTERN\nEUROPE','AFRICA']
arrivals_percent = [36.9,30.4,13.8,4.4,4.0,3.6,2.9,2.6,1.5]
# Set up the figure and the main subplot
fig, ax1 = plt.subplots(figsize=(20,12))
# Draw the bar plot
rects1 = ax1.bar(range(10),top10_arrivals_values, align='center',color='#3b5998')
# Set spines to be invisible
for spine in ax1.spines.values():
spine.set_visible(False)
# Format ticks and labels
plt.xticks(range(10),top10_arrivals_countries,fontsize=18)
for tic in ax1.xaxis.get_major_ticks():
tic.tick1On = tic.tick2On = False
plt.yticks(fontsize=18)
plt.xlabel('Top 10 tourist generating countries',fontsize=24,fontweight='semibold')
plt.ylabel('Arrivals (Million)',fontsize=24,fontweight='semibold')
# Prepare the inset axes
ax2 = inset_axes(ax1, width=6.5, height=6.5, loc=5)
# Store the pie chart sectors, sample labels and value labels
# Set the properties
explode = (0.08, 0.08, 0.05, 0.05,0.05,0.05,0.05,0.05,0.05)
patches, texts, autotexts = ax2.pie(arrivals_percent, \
labels=arrivals_countries,\
autopct='%1.1f%%', \
shadow=True, startangle=180,\
explode=explode, \
counterclock=False, \
pctdistance=0.72)
# Set properties of text in pie chart
for text in texts+autotexts:
text.set_fontsize(16)
text.set_fontweight('semibold')
# Label bar values
autolabel(rects1)
# Add a super title to all the subplots
plt.suptitle('Non-Resident Arrivals to the US in 2016 (by Aug) by Regions',fontsize=36,\
color='navy',fontweight='bold')
# Show the figure
plt.show()
# -
# +
# Adding graphical annotations
# +
# https://www.nobelprize.org/nobel_prizes/lists/age.html
import re
prizes = {'Physics':[],'Chemistry':[],'Physiology or Medicine':[],'Economics':[],'Literature':[],'Peace':[]}
curr_age = 0
f = open('nobelbyage.txt')
for line in f.readlines():
if 'Age' in line:
curr_age = int(re.search('Age (\d+)',line).group(1))
elif len(line) > 0:
splitline = line.strip().rsplit(' ',1)
prize = splitline[0]
if prize in list(prizes):
year = int(splitline[1])
prizes[prize].append((year,curr_age))
f.close()
for prize in ['Physics','Chemistry','Physiology or Medicine','Economics','Literature','Peace']:
prizes[prize] = sorted(prizes[prize])
print(prize)
print(prizes[prize])
# +
import numpy as np
from matplotlib.patches import Circle, Wedge, Polygon, Ellipse
from matplotlib.collections import PatchCollection
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
patches = []
# Full and ring sectors drawn by Wedge((x,y),r,deg1,deg2)
leftstripe = Wedge((.46, .5), .15, 90,100) # Full sector by default
midstripe = Wedge((.5,.5), .15, 85,95)
rightstripe = Wedge((.54,.5), .15, 80,90)
lefteye = Wedge((.36, .46), .06, 0, 360, width=0.03) # Ring sector drawn when width <1
righteye = Wedge((.63, .46), .06, 0, 360, width=0.03)
nose = Wedge((.5, .32), .08, 75,105, width=0.03)
mouthleft = Wedge((.44, .4), .08, 240,320, width=0.01)
mouthright = Wedge((.56, .4), .08, 220,300, width=0.01)
patches += [leftstripe,midstripe,rightstripe,lefteye,righteye,nose,mouthleft,mouthright]
# Circles
leftiris = Circle((.36,.46),0.04)
rightiris = Circle((.63,.46),0.04)
patches += [leftiris,rightiris]
# Polygons drawn by passing coordinates of vertices
leftear = Polygon([[.2,.6],[.3,.8],[.4,.64]], True)
rightear = Polygon([[.6,.64],[.7,.8],[.8,.6]], True)
topleftwhisker = Polygon([[.01,.4],[.18,.38],[.17,.42]], True)
bottomleftwhisker = Polygon([[.01,.3],[.18,.32],[.2,.28]], True)
toprightwhisker = Polygon([[.99,.41],[.82,.39],[.82,.43]], True)
bottomrightwhisker = Polygon([[.99,.31],[.82,.33],[.81,.29]], True)
patches+=[leftear,rightear,topleftwhisker,bottomleftwhisker,toprightwhisker,bottomrightwhisker]
# Ellipse drawn by Ellipse((x,y),width,height)
body = Ellipse((0.5,-0.18),0.6,0.8)
patches.append(body)
# Draw the patches
colors = 100*np.random.rand(len(patches)) # set random colors
p = PatchCollection(patches, alpha=0.4)
p.set_array(np.array(colors))
ax.add_collection(p)
# Show the figure
plt.show()
# -
# +
# Adding image annotations
# +
import urllib
img_fetch = urllib.request.urlopen(url)
pyplot.imread(img_fetch, format)
imagebox = OffsetImage(img, zoom)
imagebox.image.axes = ax
ab = AnnotationBbox(imagebox, (x1,y1),
xybox=(x2,y2),
xycoords='data',
boxcoords='data',
pad=0.5,
arrowprops=dict(
arrowstyle="->",
connectionstyle="angle,angleA=90,angleB=0,rad=3")
)
ax.add_artist(ab)
# +
import re
prizes = {'Physics':[],'Chemistry':[],'Physiology or Medicine':[],\
'Economics':[],'Literature':[],'Peace':[]}
curr_age = 0
f = open('nobelbyage.txt')
for line in f.readlines():
if 'Age' in line:
curr_age = int(re.search('Age (\d+)',line).group(1))
elif len(line) > 0:
splitline = line.strip().rsplit(' ',1)
prize = splitline[0]
if prize in list(prizes):
year = int(splitline[1])
prizes[prize].append((year,curr_age))
f.close()
for prize in ['Physics','Chemistry','Physiology or Medicine',\
'Economics','Literature','Peace']:
prizes[prize] = sorted(prizes[prize])
print(prize)
print(prizes[prize])
# +
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
from matplotlib.offsetbox import (OffsetImage,AnnotationBbox)
import urllib
mpl.style.use('seaborn')
# A dictionary with the six fields as keys and tuples of years and ages of award received is prepared
# Data can be downloaded from our Github repository
# Prizes = {'Peace': [(2014, 17), (1976, 32), (2011, 32), ..., 'Chemistry':...}
# Prepare the subplots
fig, axarr = plt.subplots(2,3,sharex=True,sharey=True)
plt.subplots_adjust(left=0, right=1, top=1.5, bottom=0)
for i,prize in enumerate(['Physics','Chemistry','Physiology or Medicine','Economics',\
'Literature','Peace']):
years = [x[0] for x in prizes[prize]]
ages = [x[1] for x in prizes[prize]]
a = int(i/3) # subplot row index
b = i%3 # subplot column index
axarr[a,b].scatter(years,ages,s=2.5)
axarr[a,b].plot(years, np.poly1d(np.polyfit(years, ages, 3))(years),linewidth=2)
axarr[a,b].set_xlabel(prize,fontsize=16)
# Annotate with photo of the youngest Nobel laureate
img_fetch = urllib.request.urlopen("https://www.nobelprize.org/nobel_prizes/peace/\
laureates/2014/yousafzai_postcard.jpg") # fetch the online image
img = plt.imread(img_fetch, format='jpg')
imagebox = OffsetImage(img, zoom=0.2)
imagebox.image.axes = axarr[1,2]
ab1 = AnnotationBbox(imagebox, (2014,16),
xybox=(1920,28),
xycoords='data',
boxcoords='data',
pad=0.5,
arrowprops=dict(
arrowstyle="->",
connectionstyle="angle,angleA=90,angleB=0,rad=3")
)
axarr[1,2].add_artist(ab1)
# Annotate with photo of the oldest Nobel laureate
img_fetch = urllib.request.urlopen("http://www.nobelprize.org/nobel_prizes/\
economic-sciences/laureates/2007/hurwicz_postcard.jpg")
img = plt.imread(img_fetch, format='jpg')
imagebox = OffsetImage(img, zoom=0.2)
imagebox.image.axes = axarr[1,2]
ab2 = AnnotationBbox(imagebox, (2007,89),
xybox=(1930,60),
xycoords='data',
boxcoords='data',
pad=0.5,
arrowprops=dict(
arrowstyle="->",
connectionstyle="angle,angleA=90,angleB=0,rad=3")
)
axarr[1,0].add_artist(ab2)
# Show the figure
plt.show()
mpl.rcParams.update(mpl.rcParamsDefault)
# -
| Section 3/Section 3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# #!pip install graphviz --user
# #!echo $PYTHONPATH
# #!ls -ltr /eos/user/n/nmangane/.local/lib/python2.7/site-packages/
# #!export PATH=/eos/user/n/nmangane/.local/lib/python2.7/site-packages/:$PATH
# -
from __future__ import print_function
import ROOT
from IPython.display import Image, display, SVG
#import graphviz
ROOT.ROOT.EnableImplicitMT()
RS = ROOT.ROOT
RDF = RS.RDataFrame
with open("2017_booker.py", "r") as f:
for line in f:
#print(line, end="")
continue
# +
#FIXME: Need filter efficiency calculated for single lepton generator filtered sample. First approximation will be from MCCM (0.15) but as seen before, it's not ideal.
#May need to recalculate using genWeight/sumWeights instead of sign(genWeight)/(nPositiveEvents - nNegativeEvents), confirm if there's any difference.
lumi = {"2017": 41.53,
"2018": 1}
era = "2017"
minibooker = {
"tttt":{
"era": "2017",
"isData": False,
"nEvents": 2273928,
"nEventsPositive": 1561946,
"nEventsNegative": 711982,
"sumWeights": 18645.487772,
"sumWeights2": 1094.209551,
"isSignal": True,
"crossSection": 0.012,
"color": ROOT.kAzure-2,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/tttt_2017.root",
},
"tttt_orig":{
"era": "2017",
"isData": False,
"nEvents": 2273928,
"nEventsPositive": 1561946,
"nEventsNegative": 711982,
"sumWeights": 18645.487772,
"sumWeights2": 1094.209551,
"isSignal": False,
"crossSection": 0.012,
"color": ROOT.kRed,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/tttt-orig_2.root",
},
}
booker = {
"tttt":{
"era": "2017",
"isData": False,
"nEvents": 2273928,
"nEventsPositive": 1561946,
"nEventsNegative": 711982,
"sumWeights": 18645.487772,
"sumWeights2": 1094.209551,
"isSignal": True,
"crossSection": 0.012,
"color": ROOT.kAzure-2,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/tttt_2017.root",
},
"tt_SL-GF":{
"era": "2017",
"isData": False,
"nEvents": 8836856,
"nEventsPositive": 8794464,
"nEventsNegative": 42392,
"sumWeights": 2653328498.476976,
"sumWeights2": 812201885978.209229,
"isSignal": False,
"crossSection": 6,
"color": ROOT.kRed,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/tt_SL-GF_2017.root",
},
"tt_DL-GF":{
"era": "2017",
"isData": False,
"nEvents": 8510388,
"nEventsPositive": 8467543,
"nEventsNegative": 42845,
"sumWeights": 612101836.284397,
"sumWeights2": 44925503249.097206,
"isSignal": False,
"crossSection": 1.4705,
"color": ROOT.kRed-4,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/tt_DL-GF-*_2017.root",
},
"tt_SL":{
"era": "2017",
"isData": False,
"nEvents": 20122010,
"nEventsPositive": 20040607,
"nEventsNegative": 81403,
"sumWeights": 6052480345.748356,
"sumWeights2": 1850350248120.376221,
"isSignal": False,
"crossSection": 366.2073,
"color": ROOT.kRed,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/tt_SL_2017.root",
},
"tt_DL":{
"era": "2017",
"isData": False,
"nEvents": 69098644,
"nEventsPositive": 68818780,
"nEventsNegative": 279864,
"sumWeights": 4980769113.241218,
"sumWeights2": 364913493679.955078,
"isSignal": False,
"crossSection": 89.0482,
"color": ROOT.kRed-4,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/tt_DL_2017.root",
},
"ST_tW":{
"era": "2017",
"isData": False,
"nEvents": 7945242,
"nEventsPositive": 7914815,
"nEventsNegative": 30427,
"sumWeights": 277241050.840222,
"sumWeights2": 9823995766.508368,
"isSignal": False,
"crossSection": 35.8,
"color": ROOT.kYellow,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ST_tW_2017.root",
},
"ST_tbarW":{
"era": "2017",
"isData": False,
"nEvents": 7745276,
"nEventsPositive": 7715654,
"nEventsNegative": 30427,
"sumWeights": 270762750.172525,
"sumWeights2": 9611964941.797768,
"isSignal": False,
"crossSection": 35.8,
"color": ROOT.kYellow,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ST_tbarW_2017.root",
},
"ttH":{
"era": "2017",
"isData": False,
"nEvents": 8000000,
"nEventsPositive": 7916867,
"nEventsNegative": 83133,
"sumWeights": 4216319.315884,
"sumWeights2": 2317497.816608,
"isSignal": False,
"crossSection": 0.2934,
"color": ROOT.kMagenta,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ttH_2017.root",
},
"ttWJets":{
"era": "2017",
"isData": False,
"nEvents": 9425384,
"nEventsPositive": 9404856,
"nEventsNegative": 20528,
"sumWeights": 9384328.000000,
"sumWeights2": 9425384.000000,
"isSignal": False,
"crossSection": 0.611,
"color": ROOT.kViolet,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ttWJets_2017.root",
},
"ttZJets":{
"era": "2017",
"isData": False,
"nEvents": 8536618,
"nEventsPositive": 8527846,
"nEventsNegative": 8772,
"sumWeights": 8519074.000000,
"sumWeights2": 8536618.000000,
"isSignal": False,
"crossSection": 0.783,
"color": ROOT.kViolet,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ttZJets_2017.root",
},
"ttWH":{
"era": "2017",
"isData": False,
"nEvents": 200000,
"nEventsPositive": 199491,
"nEventsNegative": 509,
"sumWeights": 198839.680865,
"sumWeights2": 199704.039588,
"isSignal": False,
"crossSection": 0.001572,
"color": ROOT.kGreen,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ttWH_2017.root",
},
"ttWW":{
"era": "2017",
"isData": False,
"nEvents": 962000,
"nEventsPositive": 962000,
"nEventsNegative": 0,
"sumWeights": 962000.000000,
"sumWeights2": 962000.000000,
"isSignal": False,
"crossSection": 0.007882,
"color": ROOT.kGreen,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ttWW_2017.root",
},
"ttWZ":{
"era": "2017",
"isData": False,
"nEvents": 200000,
"nEventsPositive": 199379,
"nEventsNegative": 621,
"sumWeights": 198625.183551,
"sumWeights2": 199708.972601,
"isSignal": False,
"crossSection": 0.002974,
"color": ROOT.kGreen,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ttWZ_2017.root",
},
"ttZZ":{
"era": "2017",
"isData": False,
"nEvents": 200000,
"nEventsPositive": 199686,
"nEventsNegative": 314,
"sumWeights": 199286.628891,
"sumWeights2": 199816.306332,
"isSignal": False,
"crossSection": 0.001572,
"color": ROOT.kGreen,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ttZZ_2017.root",
},
"ttZH":{
"era": "2017",
"isData": False,
"nEvents": 200000,
"nEventsPositive": 199643,
"nEventsNegative": 357,
"sumWeights": 199192.234990,
"sumWeights2": 199794.753976,
"isSignal": False,
"crossSection": 0.01253,
"color": ROOT.kGreen,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ttZH_2017.root",
},
"ttHH":{
"era": "2017",
"isData": False,
"nEvents": 194817,
"nEventsPositive": 194516,
"nEventsNegative": 301,
"sumWeights": 194116.909912,
"sumWeights2": 194611.090542,
"isSignal": False,
"crossSection": 0.0007408,
"color": ROOT.kGreen,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ttHH_2017.root",
},
"tttW":{
"era": "2017",
"isData": False,
"nEvents": 200000,
"nEventsPositive": 199852,
"nEventsNegative": 148,
"sumWeights": 199552.187377,
"sumWeights2": 199697.648421,
"isSignal": False,
"crossSection": 0.007882,
"color": ROOT.kGreen,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/tttW_2017.root",
},
"tttJ":{
"era": "2017",
"isData": False,
"nEvents": 200000,
"nEventsPositive": 199273,
"nEventsNegative": 727,
"sumWeights": 198394.878491,
"sumWeights2": 199663.384954,
"isSignal": False,
"crossSection": 0.0004741,
"color": ROOT.kGreen,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/tttJ_2017.root",
},
"DYJets_DL":{
"era": "2017",
"isData": False,
"nEvents": 49125561,
"nEventsPositive": 49103859,
"nEventsNegative": 21702,
"sumWeights": 49082157.000000,
"sumWeights2": 49125561.000000,
"isSignal": False,
"crossSection": 5075.6,
"color": ROOT.kCyan,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/DYJets_DL_2017.root",
},
"MuMu_B":{
"era": "2017",
"subera": "B",
"channel": "MuMu",
"isData": True,
"nEvents": 14501767,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/MuMu_B_2017.root",
},
"MuMu_C":{
"era": "2017",
"subera": "C",
"channel": "MuMu",
"isData": True,
"nEvents": 49636525,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/MuMu_C_2017.root",
},
"MuMu_D":{
"era": "2017",
"subera": "D",
"channel": "MuMu",
"isData": True,
"nEvents": 23075733,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/MuMu_D_2017.root",
},
"MuMu_E":{
"era": "2017",
"subera": "E",
"channel": "MuMu",
"isData": True,
"nEvents": 51589091,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/MuMu_E_2017.root",
},
"MuMu_F":{
"era": "2017",
"subera": "F",
"channel": "MuMu",
"isData": True,
"nEvents": 79756560,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/MuMu_F_2017.root",
},
"ElEl_B":{
"era": "2017",
"subera": "B",
"channel": "ElEl",
"isData": True,
"nEvents": 58088760,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ElEl_B_2017.root",
},
"ElEl_C":{
"era": "2017",
"subera": "C",
"channel": "ElEl",
"isData": True,
"nEvents": 65181125,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ElEl_C_2017.root",
},
"ElEl_D":{
"era": "2017",
"subera": "D",
"channel": "ElEl",
"isData": True,
"nEvents": 25911432,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ElEl_D_2017.root",
},
"ElEl_E":{
"era": "2017",
"subera": "E",
"channel": "ElEl",
"isData": True,
"nEvents": 56233597,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ElEl_E_2017.root",
},
"ElEl_F":{
"era": "2017",
"subera": "F",
"channel": "ElEl",
"isData": True,
"nEvents": 74307066,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ElEl_F_2017.root",
},
"ElMu_B":{
"era": "2017",
"subera": "B",
"channel": "ElMu",
"isData": True,
"nEvents": 4453465,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ElMu_B_2017.root",
},
"ElMu_C":{
"era": "2017",
"subera": "C",
"channel": "ElMu",
"isData": True,
"nEvents": 15595214,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ElMu_C_2017.root",
},
"ElMu_D":{
"era": "2017",
"subera": "D",
"channel": "ElMu",
"isData": True,
"nEvents": 9164365,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ElMu_D_2017.root",
},
"ElMu_E":{
"era": "2017",
"subera": "E",
"channel": "ElMu",
"isData": True,
"nEvents": 19043421,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ElMu_E_2017.root",
},
"ElMu_F":{
"era": "2017",
"subera": "F",
"channel": "ElMu",
"isData": True,
"nEvents": 25776363,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/ElMu_F_2017.root",
},
"Mu_B":{
"era": "2017",
"subera": "B",
"channel": "Mu",
"isData": True,
"nEvents": 136300266,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/Mu_B_2017.root",
},
"Mu_C":{
"era": "2017",
"subera": "C",
"channel": "Mu",
"isData": True,
"nEvents": 165652756,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/Mu_C_2017.root",
},
"Mu_D":{
"era": "2017",
"subera": "D",
"channel": "Mu",
"isData": True,
"nEvents": 70361660,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/Mu_D_2017.root",
},
"Mu_E":{
"era": "2017",
"subera": "E",
"channel": "Mu",
"isData": True,
"nEvents": 154630534,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/Mu_E_2017.root",
},
"Mu_F":{
"era": "2017",
"subera": "F",
"channel": "Mu",
"isData": True,
"nEvents": 242135500,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/Mu_F_2017.root",
},
"El_B":{
"era": "2017",
"subera": "B",
"channel": "El",
"isData": True,
"nEvents": 60537490,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/El_B_2017.root",
},
"El_C":{
"era": "2017",
"subera": "C",
"channel": "El",
"isData": True,
"nEvents": 136637888,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/El_C_2017.root",
},
"El_D":{
"era": "2017",
"subera": "D",
"channel": "El",
"isData": True,
"nEvents": 51526710,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/El_D_2017.root",
},
"El_E":{
"era": "2017",
"subera": "E",
"channel": "El",
"isData": True,
"nEvents": 102121689,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/El_E_2017.root",
},
"El_F":{
"era": "2017",
"subera": "F",
"channel": "El",
"isData": True,
"nEvents": 128467223,
"color": ROOT.kBlack,
"source": "/eos/user/n/nmangane/SWAN_projects/LogicChainRDF/El_F_2017.root",
},
}
#Set up channel bits for selection and baseline. Separation not necessary in this stage, but convenient for loops
Chan = {}
Chan["ElMu_selection"] = 24576
Chan["MuMu_selection"] = 6144
Chan["ElEl_selection"] = 512
Chan["Mu_selection"] = 128
Chan["El_selection"] = 64
Chan["selection"] = Chan["ElMu_selection"] + Chan["MuMu_selection"] + Chan["ElEl_selection"] + Chan["Mu_selection"] + Chan["El_selection"]
Chan["ElMu_baseline"] = 24576
Chan["MuMu_baseline"] = 6144
Chan["ElEl_baseline"] = 512
Chan["Mu_baseline"] = 128
Chan["El_baseline"] = 64
Chan["baseline"] = Chan["ElMu_baseline"] + Chan["MuMu_baseline"] + Chan["ElEl_baseline"] + Chan["Mu_baseline"] + Chan["El_baseline"]
#samples["tt_DL-GF"] = {}
#samples["tt_DL-GF"]["path"] = base + "crab_tt_DL-GF_2017/results/tree*.root"
#booker["tttt"]['nEvents']
# -
def defineLeptons(input_df, input_lvl_filter=None, channel=None, isData=True, useBackupChannel=False):
"""Function to take in a dataframe and return one with new columns defined,
plus event filtering based on the criteria defined inside the function"""
if channel == None:
raise RuntimeError("channel must be selected, such as 'MuMu' or 'ElMu'")
elif channel == "MuMu":
nMuExp = 2
nElExp = 0
elif channel == "ElMu":
nMuExp = 1
nElExp = 1
elif channel == "MuMu":
nMuExp = 0
nElExp = 2
#Set up channel bits for selection and baseline. Separation not necessary in this stage, but convenient for loops
Chan = {}
Chan["ElMu_selection"] = 24576
Chan["MuMu_selection"] = 6144
Chan["ElEl_selection"] = 512
Chan["Mu_selection"] = 128
Chan["El_selection"] = 64
Chan["selection"] = Chan["ElMu_selection"] + Chan["MuMu_selection"] + Chan["ElEl_selection"] + Chan["Mu_selection"] + Chan["El_selection"]
Chan["ElMu_baseline"] = 24576
Chan["MuMu_baseline"] = 6144
Chan["ElEl_baseline"] = 512
Chan["Mu_baseline"] = 128
Chan["El_baseline"] = 64
Chan["baseline"] = Chan["ElMu_baseline"] + Chan["MuMu_baseline"] + Chan["ElEl_baseline"] + Chan["Mu_baseline"] + Chan["El_baseline"]
b = {}
b["baseline"] = "(ESV_TriggerAndLeptonLogic_baseline & {0}) > 0".format(Chan["ElMu_baseline"] + Chan["MuMu_baseline"] +
Chan["ElEl_baseline"] + Chan["Mu_baseline"] + Chan["El_baseline"])
b["ElMu_baseline"] = "(ESV_TriggerAndLeptonLogic_baseline & {0}) > 0".format(Chan["ElMu_baseline"])
b["MuMu_baseline"] = "(ESV_TriggerAndLeptonLogic_baseline & {0}) == 0 && (ESV_TriggerAndLeptonLogic_baseline & {1}) > 0".format(Chan["ElMu_baseline"],
Chan["MuMu_baseline"])
b["ElEl_baseline"] = "(ESV_TriggerAndLeptonLogic_baseline & {0}) == 0 && (ESV_TriggerAndLeptonLogic_baseline & {1}) > 0".format(Chan["ElMu_baseline"] + Chan["MuMu_baseline"],
Chan["ElEl_baseline"])
b["Mu_baseline"] = "(ESV_TriggerAndLeptonLogic_baseline & {0}) == 0 && (ESV_TriggerAndLeptonLogic_baseline & {1}) > 0".format(Chan["ElMu_baseline"] + Chan["MuMu_baseline"] + Chan["ElEl_baseline"], Chan["Mu_baseline"])
b["El_baseline"] = "(ESV_TriggerAndLeptonLogic_baseline & {0}) == 0 && (ESV_TriggerAndLeptonLogic_baseline & {1}) > 0".format(Chan["ElMu_baseline"] + Chan["MuMu_baseline"] +
Chan["ElEl_baseline"] + Chan["Mu_baseline"], Chan["El_baseline"])
b["selection"] = "ESV_TriggerAndLeptonLogic_selection > 0"
b["ElMu_selection"] = "(ESV_TriggerAndLeptonLogic_selection & {0}) > 0".format(Chan["ElMu_selection"])
b["MuMu_selection"] = "(ESV_TriggerAndLeptonLogic_selection & {0}) == 0 && (ESV_TriggerAndLeptonLogic_selection & {1}) > 0".format(Chan["ElMu_selection"], Chan["MuMu_selection"])
b["ElEl_selection"] = "(ESV_TriggerAndLeptonLogic_selection & {0}) == 0 && (ESV_TriggerAndLeptonLogic_selection & {1}) > 0".format(Chan["ElMu_selection"] + Chan["MuMu_selection"], Chan["ElEl_selection"])
b["Mu_selection"] = "(ESV_TriggerAndLeptonLogic_selection & {0}) == 0 && (ESV_TriggerAndLeptonLogic_selection & {1}) > 0".format(Chan["ElMu_selection"] + Chan["MuMu_selection"] + Chan["ElEl_selection"], Chan["Mu_selection"])
b["El_selection"] = "(ESV_TriggerAndLeptonLogic_selection & {0}) == 0 && (ESV_TriggerAndLeptonLogic_selection & {1}) > 0".format(Chan["ElMu_selection"] + Chan["MuMu_selection"] + Chan["ElEl_selection"]
+ Chan["Mu_selection"], Chan["El_selection"])
if input_lvl_filter == None:
rdf_input = input_df\
.Define("mu_mask", "Muon_pt > 0").Define("e_mask", "Electron_pt > 0")
else:
if "baseline" in input_lvl_filter:
lvl_type = "baseline"
elif "selection" in input_lvl_filter:
lvl_type = "selection"
else:
raise RuntimeError("No such level permissable: must contain 'selection' or 'baseline'")
rdf_input = input_df\
.Filter(b[input_lvl_filter], input_lvl_filter)\
.Define("mu_mask", "(Muon_OSV_{0} & {1}) > 0".format(lvl_type, Chan[input_lvl_filter]))\
.Define("e_mask", "(Electron_OSV_{0} & {1}) > 0".format(lvl_type, Chan[input_lvl_filter]))
indexDefineCode = '''ROOT::VecOps::RVec<int> i({0}.size());
std::iota(i.begin(), i.end(), 0);
return i;'''
rdf = rdf_input\
.Define("Muon_idx", indexDefineCode.format("mu_mask"))\
.Define("GMuon_idx", "Muon_idx[mu_mask]")\
.Define("GMuon_pfIsoId", "Muon_pfIsoId[mu_mask]")\
.Define("GMuon_looseId", "Muon_looseId[mu_mask]")\
.Define("GMuon_pt", "Muon_pt[mu_mask]")\
.Define("GMuon_eta", "Muon_eta[mu_mask]")\
.Define("GMuon_phi", "Muon_phi[mu_mask]")\
.Define("GMuon_mass", "Muon_mass[mu_mask]")\
.Define("GMuon_charge", "Muon_charge[mu_mask]")\
.Define("GMuon_dz", "Muon_dz[mu_mask]")\
.Define("GMuon_d0", "sqrt(Muon_dz*Muon_dz + Muon_dxy*Muon_dxy)[mu_mask]")\
.Define("GMuon_ip3d", "Muon_ip3d[mu_mask]")\
.Define("GMuon_jetIdx", "Muon_jetIdx[mu_mask]")\
.Define("nGMuon", "GMuon_pt.size()")\
.Define("nLooseGMuon", "Muon_looseId[mu_mask && Muon_looseId == true].size()")\
.Define("nMediumGMuon", "Muon_mediumId[mu_mask && Muon_mediumId == true].size()")\
.Define("Electron_idx", indexDefineCode.format("e_mask"))\
.Define("GElectron_idx", "Electron_idx[e_mask]")\
.Define("GElectron_cutBased", "Electron_cutBased[e_mask]")\
.Define("GElectron_pt", "Electron_pt[e_mask]")\
.Define("GElectron_eta", "Electron_eta[e_mask]")\
.Define("GElectron_phi", "Electron_phi[e_mask]")\
.Define("GElectron_mass", "Electron_mass[e_mask]")\
.Define("GElectron_charge", "Electron_charge[e_mask]")\
.Define("GElectron_dz", "Electron_dz[e_mask]")\
.Define("GElectron_d0", "sqrt(Electron_dz*Electron_dz + Electron_dxy*Electron_dxy)[e_mask]")\
.Define("GElectron_ip3d", "Electron_ip3d[e_mask]")\
.Define("GElectron_jetIdx", "Electron_jetIdx[e_mask]")\
.Define("nGElectron", "GElectron_pt.size()")\
.Define("nLooseGElectron", "Sum(GElectron_cutBased >= 2)")\
.Define("nMediumGElectron", "Sum(GElectron_cutBased >= 3)")\
.Define("GLepton_pt", "Concatenate(Muon_pt[mu_mask], Electron_pt[e_mask])")\
.Define("nGLepton", "GLepton_pt.size()")\
.Define("GLepton_jetIdx", "Concatenate(Muon_jetIdx[mu_mask], Electron_jetIdx[e_mask])")\
.Define("GLepton_pdgId", "Concatenate(Muon_pdgId[mu_mask], Electron_pdgId[e_mask])")\
.Define("GLepton_pt_lep0", "GLepton_pt.size() > 0 ? GLepton_pt.at(0) : -1")\
.Define("GLepton_pt_lep1", "GLepton_pt.size() > 1 ? GLepton_pt.at(1) : -1")\
.Define("GLepton_jetIdx_0", "GLepton_jetIdx.size() > 0 ? GLepton_jetIdx.at(0) : 999")\
.Define("GLepton_jetIdx_1", "GLepton_jetIdx.size() > 1 ? GLepton_jetIdx.at(1) : 999")
#.Define("GLepton_pt_lep0", "GMuon_pt.at(0)")\
#.Define("GLepton_pt_lep1", "GMuon_pt.at(1)")\
#.Define("GLepton_jetIdx_lep0_alt", "Muon_jetIdx[mu_mask].at(0)")\
#.Define("GLepton_jetIdx_lep0", "GMuon_jetIdx.at(0)")\
#.Define("GLepton_jetIdx_lep1", "GMuon_jetIdx.at(1)")
#elif channel == "ElMu":
# rdf = rdf\
# .Define("GLepton_pt", "Concatenate(Muon_pt[mu_mask], Electron_pt[e_mask])")\
# .Define("nGLepton", "GLepton_pt.size()")\
# .Define("GLepton_jetIdx", "Concatenate(Muon_jetIdx[mu_mask], Electron_jetIdx[e_mask])")
#.Define("GLepton_pt_lep0", "Max(GElectron_pt.at(0), GMuon_pt.at(0))")\
#.Define("GLepton_pt_lep1", "Min(GElectron_pt.at(0), GMuon_pt.at(0))")\
#.Define("GLepton_jetIdx_lep0", "GElectron_jetIdx.at(0)")\
#.Define("GLepton_jetIdx_lep1", "GMuon_jetIdx.at(0)")
#FIXME: Try Concatenate(GMuon_jetIdx, GElectron_jetIdx) to get unified RVec
#elif channel == "ElEl":
# rdf = rdf\
# .Define("GLepton_pt", "Concatenate(Muon_pt[mu_mask], Electron_pt[e_mask])")\
# .Define("nGLepton", "GLepton_pt.size()")\
# .Define("GLepton_jetIdx", "Concatenate(Muon_jetIdx[mu_mask], Electron_jetIdx[e_mask])")
#.Define("GLepton_pt_lep0", "GElectron_pt.at(0)")\
#.Define("GLepton_pt_lep1", "GElectron.at(1)")\
#.Define("GLepton_jetIdx_lep0", "GElectron_jetIdx.at(0)")\
#.Define("GLepton_jetIdx_lep1", "GElectron_jetIdx.at(1)")
#elif channel == "El":
# raise NotImplementedError("El channel not implemented yet")
#elif channel == "Mu":
# raise NotImplementedError("Mu channel not implemented yet")
#else:
# raise NotImplementedError("{0} channel not implemented yet".format(channel))
#Things that don't work...
#NOPE doesn't work .Define("nLooseGMuon", "Sum(Muon_looseId[mu_mask])")\
return rdf
def defineWeights(input_df, crossSection=0, sumWeights=-1, lumi=0,
nEvents=-1, nEventsPositive=2, nEventsNegative=1,
channel=None, isData=True, verbose=False):
mc_def = {}
mc_def["wgt_NUMW"] = "({xs:s} * {lumi:s} * 1000 * genWeight) / (abs(genWeight) * ( {nevtp:s} - {nevtn:s} ) )"\
.format(xs=str(crossSection), lumi=str(lumi), nevt=str(nEvents),
nevtp=str(nEventsPositive), nevtn=str(nEventsNegative))
mc_def["wgt_SUMW"] = "({xs:s} * {lumi:s} * 1000 * genWeight) / {sumw:s}"\
.format(xs=str(crossSection), lumi=str(lumi), sumw=str(sumWeights))
mc_def["wgt_SUMW_PU"] = "wgt_SUMW * puWeight"
data_def = {}
data_def["wgt_NUMW"] = "1"
data_def["wgt_NUMW_V2"] = "1"
data_def["wgt_SUMW"] = "1"
data_def["wgt_SUMW_PU"] = "1"
#data_def["wgt_SUMW_LSF"] = "1"
if verbose == True:
print("===data and mc weight definitions===")
print(data_def)
print(mc_def)
if channel == "MuMu":
pass
#mc["wgt_SUMW_LSF"] = "wgt_SUMW*GMuon_"
if isData:
rdf = input_df\
.Define("wgt_NUMW", "1")\
.Define("wgt_SUMW", "1")\
.Define("wgt_SUMW_PU", "1")\
.Define("wgt_SUMW_LSF", "1")\
.Define("wgt_diff", "abs(wgt_NUMW - wgt_SUMW)/max(abs(wgt_SUMW), abs(wgt_NUMW))")
else:
rdf = input_df\
.Define("wgt_NUMW", mc_def["wgt_NUMW"])\
.Define("wgt_SUMW", mc_def["wgt_SUMW"])\
.Define("wgt_SUMW_PU", mc_def["wgt_SUMW"])\
.Define("wgt_diff", "abs(wgt_NUMW - wgt_SUMW)/max(abs(wgt_SUMW), abs(wgt_NUMW))")
return rdf
def defineJets(input_df, era="2017", doAK8Jets=False):
"""Function to take in a dataframe and return one with new columns defined,
plus event filtering based on the criteria defined inside the function"""
indexDefineCode = '''ROOT::VecOps::RVec<int> i({0}.size());
std::iota(i.begin(), i.end(), 0);
return i;'''
bTagWorkingPointDict = {
'2016':{
'DeepCSV':{'L': 0.2217, 'M': 0.6321, 'T': 0.8953, 'Var': 'btagDeepB'},
'DeepJet':{ 'L': 0.0614, 'M': 0.3093, 'T': 0.7221, 'Var': 'btagDeepFlavB'}
},
'2017':{
'CSVv2':{'L': 0.5803, 'M': 0.8838, 'T': 0.9693, 'Var': 'btagCSVV2'},
'DeepCSV':{'L': 0.1522, 'M': 0.4941, 'T': 0.8001, 'Var': 'btagDeepB'},
'DeepJet':{'L': 0.0521, 'M': 0.3033, 'T': 0.7489, 'Var': 'btagDeepFlavB'}
},
'2018':{
'DeepCSV':{'L': 0.1241, 'M': 0.4184, 'T': 0.7527, 'Var': 'btagDeepB'},
'DeepJet':{'L': 0.0494, 'M': 0.2770, 'T': 0.7264, 'Var': 'btagDeepFlavB'}
}
}
rdf = input_df\
.Define("jet_mask", "(Jet_OSV_baseline & {0}) > 0".format(23))\
.Define("jet_maskALT", "(Jet_pt > 20 && abs(Jet_eta) < 2.5 && Jet_jetId >= 2)")\
.Define("Jet_idx", indexDefineCode.format("jet_mask"))\
.Define("GJet_idx", "Jet_idx[jet_mask]")\
.Define("GJet_pt", "Jet_pt[jet_mask]")\
.Define("GJet_ptALT", "Jet_pt[jet_maskALT]")\
.Define("GJet_eta", "Jet_eta[jet_mask]")\
.Define("GJet_etaALT", "Jet_eta[jet_maskALT]")\
.Define("GJet_jetId", "Jet_jetId[jet_mask]")\
.Define("GJet_jetIdALT", "Jet_jetId[jet_maskALT]")\
.Define("GJet_btagDeepB_jet0", "Jet_btagDeepB[jet_mask].at(0)")\
.Define("GJet_btagDeepB_jet1", "Jet_btagDeepB[jet_mask].at(1)")\
.Define("nGJet", "GJet_pt.size()")\
.Define("GJet_MedDeepCSV", "Jet_{0}[Jet_{0} > {1}].size()"\
.format(bTagWorkingPointDict[era]["DeepCSV"]["Var"],
bTagWorkingPointDict[era]["DeepCSV"]["M"]))\
.Define("GJet_MediumDeepJet", "Jet_{}[Jet_{0} > {1}].size()"\
.format(bTagWorkingPointDict[era]["DeepJet"]["Var"],
bTagWorkingPointDict[era]["DeepJet"]["M"]))
return rdf
def fillHistos(input_df, input_name=None, wgtVar="wgt_SUMW", histos1D_dict=None, histos2D_dict=None, histosNS_dict=None,
doMuons=False, doElectrons=False, doLeptons=False, doJets=False, doWeights=False, doEventVars=False):
"""Method to fill histograms given an input RDataFrame, input sample/dataset name, input histogram dictionaries.
Has several options of which histograms to fill, such as Leptons, Jets, Weights, EventVars, etc.
Types of histograms (1D, 2D, those which will not be stacked(NS - histosNS)) are filled by passing non-None
value to that histosXX_dict variable. Internally stored with structure separating the categories of histos,
with 'Muons,' 'Electrons,' 'Leptons,' 'Jets,' 'EventVars,' 'Weights' subcategories."""
if doMuons == False and doElectrons == False and doLeptons == False\
and doJets==False and doWeights==False and doEventVars==False:
raise RuntimeError("Must select something to plot:"\
"Set do{Muons,Electrons,Leptons,Jets,Weights,EventVars,etc} = True in init method")
if doWeights == True:
if histosNS_dict != None:
if "EventVars" not in histosNS_dict:
histosNS_dict["EventVars"] = {}
histosNS[name][lvl]["event"]["wgt_NUMW"] = input_df.Histo1D("wgt_NUMW")
histosNS[name][lvl]["event"][wgtVar] = input_df.Histo1D(wgtVar)
if histos1D_dict != None:
if "EventVars" not in histos1D_dict:
histos1D_dict["EventVars"] = {}
histos1D_dict["event"]["wgt_diff"] = input_df.Histo1D(("wgt_diff",
"(wgt_NUMW - wgt_SUMW)/wgt_SUMW",
1000, -1, 1), "wgt_diff")
if doMuons == True:
if histos1D_dict != None:
if "Muons" not in histos1D_dict:
histos1D_dict["Muons"] = {}
histos1D_dict["Muons"]["idx"] = input_df.Histo1D(("idx", "", 5, 0, 5), "Muon_idx", wgtVar)
histos1D_dict["Muons"]["Gidx"] = input_df.Histo1D(("Gidx", "", 5, 0, 5), "GMuon_idx", wgtVar)
histos1D_dict["Muons"]["nMu"] = input_df.Histo1D(("nMuon", "", 5, 0, 5), "nGMuon", wgtVar)
histos1D_dict["Muons"]["nLooseMu"] = input_df.Histo1D(("nLooseMuon", "", 5, 0, 5), "nLooseGMuon", wgtVar)
histos1D_dict["Muons"]["nMediumMu"] = input_df.Histo1D(("nMediumMuon", "", 5, 0, 5), "nMediumGMuon", wgtVar)
histos1D_dict["Muons"]["pt"] = input_df.Histo1D(("Muon_pt", "", 100, 0, 500), "GMuon_pt", wgtVar)
histos1D_dict["Muons"]["eta"] = input_df.Histo1D(("Muon_eta", "", 104, -2.6, 2.6), "GMuon_eta", wgtVar)
histos1D_dict["Muons"]["phi"] = input_df.Histo1D(("Muon_phi", "", 64, -3.1416, 3.1416), "GMuon_phi", wgtVar)
histos1D_dict["Muons"]["mass"] = input_df.Histo1D(("Muon_mass", "", 50, 0, 1), "GMuon_mass", wgtVar)
histos1D_dict["Muons"]["iso"] = input_df.Histo1D(("Muon_iso", "", 8, 0, 8), "GMuon_pfIsoId", wgtVar)
histos1D_dict["Muons"]["dz"] = input_df.Histo1D(("Muon_dz", "", 100, -0.01, 0.01), "GMuon_dz", wgtVar)
histos1D_dict["Muons"]["d0"] = input_df.Histo1D(("Muon_d0", "", 100, -0.01, 0.01), "GMuon_d0", wgtVar)
histos1D_dict["Muons"]["ip3d"] = input_df.Histo1D(("Muon_ip3d", "", 100, -0.01, 0.01), "GMuon_ip3d", wgtVar)
if histos2D_dict != None:
if "Muons" not in histos2D_dict:
histos2D_dict["Muons"] = {}
histos2D_dict["Muons"]["eta_phi"] = input_df.Histo2D(("Muon_eta_phi", "",
104, -2.6, 2.6,
64, -3.1416, 3.1416),
"GMuon_eta", "GMuon_phi", wgtVar)
histos2D_dict["Muons"]["dz_ip3d"] = input_df.Histo2D(("Muon_dz_ip3d", "",
100, -0.01, 0.01,
100, 0, 0.01),
"GMuon_dz", "GMuon_ip3d", wgtVar)
if doElectrons == True:
if histos1D_dict != None:
if "Electrons" not in histos1D_dict:
histos1D_dict["Electrons"] = {}
histos1D_dict["Electrons"]["nEl"] = input_df.Histo1D(("nElectron", "", 5, 0, 5), "nGElectron", wgtVar)
histos1D_dict["Electrons"]["nLooseEl"] = input_df.Histo1D(("nLooseElectron", "", 5, 0, 5), "nLooseGElectron", wgtVar)
histos1D_dict["Electrons"]["nMediumEl"] = input_df.Histo1D(("nMediumElectron", "", 5, 0, 5), "nMediumGElectron", wgtVar)
histos1D_dict["Electrons"]["pt"] = input_df.Histo1D(("Electron_pt", "", 100, 0, 500), "GElectron_pt", wgtVar)
histos1D_dict["Electrons"]["eta"] = input_df.Histo1D(("Electron_eta", "", 104, -2.6, 2.6), "GElectron_eta", wgtVar)
histos1D_dict["Electrons"]["phi"] = input_df.Histo1D(("Electron_phi", "", 64, -3.1416, 3.1416), "GElectron_phi", wgtVar)
histos1D_dict["Electrons"]["mass"] = input_df.Histo1D(("Electron_mass", "", 50, 0, 1), "GElectron_mass", wgtVar)
histos1D_dict["Electrons"]["dz"] = input_df.Histo1D(("Electron_dz", "", 100, -0.01, 0.01), "GElectron_dz", wgtVar)
histos1D_dict["Electrons"]["d0"] = input_df.Histo1D(("Electron_d0", "", 100, 0, 0.01), "GElectron_d0", wgtVar)
histos1D_dict["Electrons"]["ip3d"] = input_df.Histo1D(("Electron_ip3d", "", 100, 0, 0.01), "GElectron_ip3d", wgtVar)
histos1D_dict["Electrons"]["cutBased"] = input_df.Histo1D(("Electron_cutBased", "", 5, 0, 5), "GElectron_cutBased", wgtVar)
if histos2D_dict != None:
if "Electrons" not in histos2D_dict:
histos2D_dict["Electrons"] = {}
histos2D_dict["Electrons"]["eta_phi"] = input_df.Histo2D(("Electron_eta_phi", "",
104, -2.6, 2.6,
64, -3.1416, 3.1416),
"GElectron_eta", "GElectron_phi", wgtVar)
histos2D_dict["Electrons"]["dz_ip3d"] = input_df.Histo2D(("Electron_dz_ip3d", "",
100, -0.01, 0.01,
100, 0, 0.01),
"GElectron_dz", "GElectron_ip3d", wgtVar)
if doLeptons == True:
if histos1D != None:
if "Leptons" not in histos1D_dict:
histos1D_dict["Leptons"] = {}
histos1D_dict["Leptons"]["pt_lep0"] = input_df\
.Histo1D(("GLepton_pt_lep0", "", 100, 0, 500),"GLepton_pt_lep0", wgtVar)
histos1D_dict["Leptons"]["pt_lep1"] = input_df\
.Histo1D(("GLepton_pt_lep1", "", 100, 0, 500),"GLepton_pt_lep1", wgtVar)
histos1D_dict["Leptons"]["nLepton"] = input_df\
.Histo1D(("nLepton", "", 5, 0, 5), "nGLepton", wgtVar)
histos1D_dict["Leptons"]["pdgId"] = input_df\
.Histo1D(("Lepton_pdgId", "", 32, -16, 16), "GLepton_pdgId", wgtVar)
histos1D_dict["Leptons"]["jetIdx"] = input_df\
.Histo1D(("Lepton_jetIdx", "", 20, 0, 20), "GLepton_jetIdx", wgtVar)
if doJets == True:
if histos1D != None:
if "Jets" not in histos1D_dict:
histos1D_dict["Jets"] = {}
histos1D_dict["Jets"]["pt"] = input_df.Histo1D(("Jet_pt", "", 100, 0, 500), "GJet_pt", wgtVar)
histos1D_dict["Jets"]["eta"] = input_df.Histo1D(("Jet_eta", "", 104, -2.6, 2.6), "GJet_eta", wgtVar)
histos1D_dict["Jets"]["phi"] = input_df.Histo1D(("Jet_phi", "", 64, -3.1416, 3.1416), "GJet_eta", wgtVar)
histos1D_dict["Jets"]["mass"] = input_df.Histo1D(("Jet_mass", "", 100, 0, 500), "GJet_eta", wgtVar)
histos1D_dict["Jets"]["jetId"] = input_df.Histo1D(("Jet_jetId", "", 8, 0, 8), "GJet_jetId", wgtVar)
histos1D_dict["Jets"]["ptALT"] = input_df.Histo1D(("Jet_ptALT", "", 100, 0, 500), "GJet_ptALT", wgtVar)
histos1D_dict["Jets"]["etaALT"] = input_df.Histo1D(("Jet_etaALT", "", 104, -2.6, 2.6), "GJet_etaALT", wgtVar)
histos1D_dict["Jets"]["jetIdALT"] = input_df.Histo1D(("Jet_jetIdALT", "", 8, 0, 8), "GJet_jetIdALT", wgtVar)
histos1D_dict["Jets"]["btagDeepB_jet0"] = input_df.Histo1D(("Jet_btagDeepB_jet0", "", 101, -0.01, 1), "GJet_btagDeepB_jet0", wgtVar)
histos1D_dict["Jets"]["btagDeepB_jet0"] = input_df.Histo1D(("Jet_btagDeepB_jet1", "", 101, -0.01, 1), "GJet_btagDeepB_jet1", wgtVar)
histos1D_dict["Jets"]["nMediumCSVv2"] = input_df.Histo1D(("nJet_MediumCSVv2", "", 10, 0, 10), "nGJet_MediumCSVv2", wgtVar)
histos1D_dict["Jets"]["nMediumDeepCSV"] = input_df.Histo1D(("nJet_MediumDeepCSV", "", 10, 0, 10), "nGJet_MediumDeepCSV", wgtVar)
histos1D_dict["Jets"]["nMediumDeepJet"] = input_df.Histo1D(("nJet_MediumDeepJet", "", 10, 0, 10), "nGJet_MediumDeepJet", wgtVar)
histos1D_dict["Jets"]["nJet"] = input_df.Histo1D(("nJet", "", 15, 0, 15), "nGJet", wgtVar)
histos1D_dict["Jets"]["nJetNUMW"] = input_df.Histo1D(("nJet_NUMW", "", 15, 0, 15), "nGJet", "wgt_NUMW_V2")
histos1D_dict["Jets"]["nJetSUMW_PU"] = input_df.Histo1D(("nJet_SUMW_PU", "", 15, 0, 15), "nGJet", "wgt_SUMW_PU")
histos1D_dict["Jets"]["nJetSUMW_LSF"] = input_df.Histo1D(("nJet_SUMW_LSF", "", 15, 0, 15), "nGJet", "wgt_SUMW_LSF")
if histos2D != None:
if "Jets" not in histos2d_dict:
histos2D_dict["Jets"] = {}
histos2D_dict["Jets"]["eta_phi"] = input_df.Histo2D(("Jet_eta_phi", "",
104, -2.6, 2.6,
64, -3.1416, 3.1416),
"GJet_eta", "GJet_phi", wgtVar)
# +
#Focus on limited set of events at a time
#levels_of_interest = set(["ElMu_baseline"])
#levels_of_interest = set(["selection", "ElMu_selection", "ElEl_selection", "MuMu_selection", "Mu_selection", "El_selection"])
levels_of_interest = set(["baseline", "MuMu_baseline", "ElEl_baseline", "selection", "MuMu_selection", "ElMu_selection"])
print("Creating selection and baseline bits")
b = {}
b["ElMu_baseline"] = "(ESV_TriggerAndLeptonLogic_baseline & {0}) > 0".format(Chan["ElMu_baseline"])
b["MuMu_baseline"] = "(ESV_TriggerAndLeptonLogic_baseline & {0}) == 0 && (ESV_TriggerAndLeptonLogic_baseline & {1}) > 0".format(Chan["ElMu_baseline"],
Chan["MuMu_baseline"])
b["ElEl_baseline"] = "(ESV_TriggerAndLeptonLogic_baseline & {0}) == 0 && (ESV_TriggerAndLeptonLogic_baseline & {1}) > 0".format(Chan["ElMu_baseline"] + Chan["MuMu_baseline"],
Chan["ElEl_baseline"])
b["Mu_baseline"] = "(ESV_TriggerAndLeptonLogic_baseline & {0}) == 0 && (ESV_TriggerAndLeptonLogic_baseline & {1}) > 0".format(Chan["ElMu_baseline"] + Chan["MuMu_baseline"] + Chan["ElEl_baseline"], Chan["Mu_baseline"])
b["El_baseline"] = "(ESV_TriggerAndLeptonLogic_baseline & {0}) == 0 && (ESV_TriggerAndLeptonLogic_baseline & {1}) > 0".format(Chan["ElMu_baseline"] + Chan["MuMu_baseline"] +
Chan["ElEl_baseline"] + Chan["Mu_baseline"], Chan["El_baseline"])
b["selection"] = "ESV_TriggerAndLeptonLogic_selection > 0"
b["ElMu_selection"] = "(ESV_TriggerAndLeptonLogic_selection & {0}) > 0".format(Chan["ElMu_selection"])
b["MuMu_selection"] = "(ESV_TriggerAndLeptonLogic_selection & {0}) == 0 && (ESV_TriggerAndLeptonLogic_selection & {1}) > 0".format(Chan["ElMu_selection"], Chan["MuMu_selection"])
b["ElEl_selection"] = "(ESV_TriggerAndLeptonLogic_selection & {0}) == 0 && (ESV_TriggerAndLeptonLogic_selection & {1}) > 0".format(Chan["ElMu_selection"] + Chan["MuMu_selection"], Chan["ElEl_selection"])
b["Mu_selection"] = "(ESV_TriggerAndLeptonLogic_selection & {0}) == 0 && (ESV_TriggerAndLeptonLogic_selection & {1}) > 0".format(Chan["ElMu_selection"] + Chan["MuMu_selection"] + Chan["ElEl_selection"], Chan["Mu_selection"])
b["El_selection"] = "(ESV_TriggerAndLeptonLogic_selection & {0}) == 0 && (ESV_TriggerAndLeptonLogic_selection & {1}) > 0".format(Chan["ElMu_selection"] + Chan["MuMu_selection"] + Chan["ElEl_selection"]
+ Chan["Mu_selection"], Chan["El_selection"])
filtered = {}
for name, vals in minibooker.items():
if name == "tttt_orig": continue
print("Caching - {}".format(name))
filtered[name] = {}
for lvl in levels_of_interest:
if lvl == "baseline":
filtered[name][lvl] = RDF("Events", vals["source"])#.Cache()
else:
filtered[name][lvl] = RDF("Events", vals["source"]).Filter(b[lvl], lvl)#.Cache()
# -
samples = {}
counts = {}
histos1D = {}
histos2D = {}
histosNS = {} #unstacked histograms
the_df = {}
print("Starting loop for booking")
for name, vals in minibooker.items():
if name == "tttt_orig": continue
#if name not in ["tttt", "ElMu_F"]: continue
print("Booking - {}".format(name))
counts[name] = {}
histos1D[name] = {}
histos2D[name] = {}
histosNS[name] = {}
the_df[name] = {}
#counts[name]["baseline"] = filtered[name].Count() #Unnecessary with baseline in levels of interest?
for lvl in levels_of_interest:
the_df[name][lvl] = defineLeptons(filtered[name][lvl],
input_lvl_filter=lvl,
channel="MuMu",
isData=vals["isData"],
useBackupChannel=False)
the_df[name][lvl] = defineWeights(the_df[name][lvl],
crossSection=vals["crossSection"],
sumWeights=vals["sumWeights"],
lumi=lumi[era],
nEvents=vals["nEvents"],
nEventsPositive=vals["nEventsPositive"],
nEventsNegative=vals["nEventsNegative"],
channel="MuMu",
isData=vals["isData"],
verbose=True)
counts[name][lvl] = the_df[name][lvl].Count()
histos1D[name][lvl] = {}
histosNS[name][lvl] = {}
histos2D[name][lvl] = {}
fillHistos(the_df[name][lvl], wgtVar="wgt_SUMW", histos1D_dict=histos1D[name][lvl],
histos2D_dict=histos2D[name][lvl], histosNS_dict=histosNS[name][lvl],
doMuons=True, doElectrons=True, doLeptons=True,
doJets=False, doWeights=False, doEventVars=False)
print("Warning: if filtered[name][lvl] RDFs are not reset, then calling define* on them will cause the error"\
"with 'program state reset'")
loopcounter = 0
for name, cnt in counts.items():
loopcounter += 1
print("==========={}/{}\n{}".format(loopcounter, len(counts), name))
if "baseline" in cnt:
print("Baseline = " + str(cnt["baseline"].GetValue()))
if "ElMu_baseline" in cnt:
print("\tElMu = {}".format(cnt["ElMu_baseline"].GetValue()),end='')
if "MuMu_baseline" in cnt:
print("\tMuMu = {}".format(cnt["MuMu_baseline"].GetValue()),end='')
if "ElEl_baseline" in cnt:
print("\tElEl = {}".format(cnt["ElEl_baseline"].GetValue()),end='')
if "Mu_baseline" in cnt:
print("\tMu = {}".format(cnt["Mu_baseline"].GetValue()),end='')
if "El_baseline" in cnt:
print("\tEl = {}".format(cnt["El_baseline"].GetValue()),end='')
print("")
if "ElMu_baseline" in cnt and "ElEl_baseline" in cnt and "MuMu_baseline" in cnt\
and "Mu_baseline" in cnt and "El_baseline" in cnt:
print("\nTotal = {}".format(cnt["ElMu_baseline"].GetValue() + cnt["MuMu_baseline"].GetValue() + cnt["ElEl_baseline"].GetValue() + cnt["Mu_baseline"].GetValue() + cnt["El_baseline"].GetValue()))
if "selection" in cnt:
print("Selection = " + str(cnt["selection"].GetValue()))
if "ElMu_selection" in cnt:
print("\tElMu = {}".format(cnt["ElMu_selection"].GetValue()),end='')
if "MuMu_selection" in cnt:
print("\tMuMu = {}".format(cnt["MuMu_selection"].GetValue()),end='')
if "ElEl_selection" in cnt:
print("\tElEl = {}".format(cnt["ElEl_selection"].GetValue()),end='')
if "Mu_selection" in cnt:
print("\tMu = {}".format(cnt["Mu_selection"].GetValue()),end='')
if "El_selection" in cnt:
print("\tEl = {}".format(cnt["El_selection"].GetValue()),end='')
print("")
if "ElMu_selection" in cnt and "ElEl_selection" in cnt and "MuMu_selection" in cnt\
and "Mu_selection" in cnt and "El_selection" in cnt:
print("\nTotal = {}".format(cnt["ElMu_selection"].GetValue() + cnt["MuMu_selection"].GetValue() + cnt["ElEl_selection"].GetValue() + cnt["Mu_selection"].GetValue() + cnt["El_selection"].GetValue()))
stacks = {}
stacksource = {} #Create sortable lists to fill stacks from
stacksource_data = {} #create separte list to append all the data to, so that they can be conbined into one hist file and added to the stacksoure at the end
for level, obj_dict in histos1D['tttt'].items():
if level not in levels_of_interest: continue
stacks[level] = {}
stacksource[level] = {}
stacksource_data[level] = {}
for obj_name, obj_val in obj_dict.items():
stacks[level][obj_name] = {}
stacksource[level][obj_name] = {}
stacksource_data[level][obj_name] = {}
for hname, hist in obj_val.items():
stacks[level][obj_name][hname] = []
stacks[level][obj_name][hname].append(ROOT.THStack("s_{}_{}_{}".format(level, obj_name, hname), "{}_{}_{}".format(level, obj_name, hname)))
stacksource[level][obj_name][hname] = []
stacksource_data[level][obj_name][hname] = []
for name, levels_dict in histos1D.items():
#if "DY" not in name and "t" not in name: continue
#if booker[name]["isData"] == True: continue
print(name, end='')
#print(booker[name].keys())
print(" - c=" + str(booker[name]["color"]))
for level, obj_dict in levels_dict.items():
if level not in levels_of_interest: continue
print("\t" + level)
for obj_name, obj_val in obj_dict.items():
print("\t\t" + obj_name)
for hname, hist in obj_val.items():
print("\t\t\t" + hname)
#help(hist)
hptr = hist.GetPtr().Clone()
hptr.SetFillColor(booker[name]["color"])
hptr.SetLineColor(booker[name]["color"])
#stacks[level][obj_name][hname].Add(hptr)
#stacksource[level][obj_name][hname].append((hptr, hptr.GetIntegral()))
#Integral fails sometimes, use sum of weights...
if booker[name]["isData"] == False:
stacksource[level][obj_name][hname].append((hptr, hptr.GetSumOfWeights(), booker[name]["isData"]))
else:
stacksource_data[level][obj_name][hname].append((hptr, hptr.GetSumOfWeights(), booker[name]["isData"]))
print()
#Now cycle through and sort each list, once it contains all hists from every source (outermost loop - name - above)
print(stacksource_data)
for level, obj_dict in histos1D['tttt'].items():
if level not in levels_of_interest: continue
for obj_name, obj_val in obj_dict.items():
for hname, hist in obj_val.items():
stacksource[level][obj_name][hname].sort(key=lambda b: b[1], reverse=False)
tmp = None
for hi, h_data in enumerate(stacksource_data[level][obj_name][hname]):
print("hi = {}".format(hi))
if hi == 0:
#take first histo
tmp = h_data[0]
else:
#hadd the other histos
tmp = tmp + h_data[0]
if tmp != None:
tmp.SetMarkerStyle(0)
tmp.SetLineColor(ROOT.kBlack)
tmp.SetFillColor(ROOT.kWhite)
stacks[level][obj_name][hname].append(tmp)
for hptrTup in stacksource[level][obj_name][hname]:
#add to the THStack in the first position of the tuple
stacks[level][obj_name][hname][0].Add(hptrTup[0])
# +
# %jsroot on
leg = ROOT.TLegend(0.75,0.85, 0.6, 0.75)
#leg.SetFillColor(0)
#leg.SetBorderSize(0)
leg.SetTextSize(0.03)
for obj_name, obj_dict in stacks["ElMu_selection"].items():
print(obj_name)
for sname, stack in obj_dict.items():
c = ROOT.TCanvas("cs_{}_{}".format(obj_name, sname), "", 800, 600)
c.cd()
#For data first
#stack[1].Draw("PE1")
#stack[0].Draw("HIST SAME")
#for MC first
stack[0].Draw("HIST S")
#FIXME - not drawing data stack[1].Draw("PE1 SAME")
# Add header
cms_label = ROOT.TLatex()
cms_label.SetTextSize(0.04)
cms_label.DrawLatexNDC(0.16, 0.92, "#bf{CMS Preliminary}")
header = ROOT.TLatex()
header.SetTextSize(0.03)
header.DrawLatexNDC(0.63, 0.92, "#sqrt{{s}} = 13 TeV, L_{{int}} = {0} fb^{{-1}}".format(lumi[era]))
leg.Draw()
c.Draw()
# +
#From example: https://root.cern.ch/doc/master/df103__NanoAODHiggsAnalysis_8py_source.html
#def plot(sig, bkg, data, x_label, filename):
# """
# Plot invariant mass for signal and background processes from simulated
# events overlay the measured data.
# """
# # Canvas and general style options
# ROOT.gStyle.SetOptStat(0)
# ROOT.gStyle.SetTextFont(42)
# d = ROOT.TCanvas("d", "", 800, 700)
# d.SetLeftMargin(0.15)
#
# # Get signal and background histograms and stack them to show Higgs signal
# # on top of the background process
# h_bkg = bkg
# h_cmb = sig.Clone()
#
# h_cmb.Add(h_bkg)
# h_cmb.SetTitle("")
# h_cmb.GetXaxis().SetTitle(x_label)
# h_cmb.GetXaxis().SetTitleSize(0.04)
# h_cmb.GetYaxis().SetTitle("N_{Events}")
# h_cmb.GetYaxis().SetTitleSize(0.04)
# h_cmb.SetLineColor(ROOT.kRed)
# h_cmb.SetLineWidth(2)
# h_cmb.SetMaximum(18)
# h_bkg.SetLineWidth(2)
# h_bkg.SetFillStyle(1001)
# h_bkg.SetLineColor(ROOT.kBlack)
# h_bkg.SetFillColor(ROOT.kAzure - 9)
#
# # Get histogram of data points
# h_data = data
# h_data.SetLineWidth(1)
# h_data.SetMarkerStyle(20)
# h_data.SetMarkerSize(1.0)
# h_data.SetMarkerColor(ROOT.kBlack)
# h_data.SetLineColor(ROOT.kBlack)
#
# # Draw histograms
# h_cmb.DrawCopy("HIST")
# h_bkg.DrawCopy("HIST SAME")
# h_data.DrawCopy("PE1 SAME")
#
# # Add legend
# legend = ROOT.TLegend(0.62, 0.70, 0.82, 0.88)
# legend.SetFillColor(0)
# legend.SetBorderSize(0)
# legend.SetTextSize(0.03)
# legend.AddEntry(h_data, "Data", "PE1")
# legend.AddEntry(h_bkg, "ZZ", "f")
# legend.AddEntry(h_cmb, "m_{H} = 125 GeV", "f")
# legend.Draw()
#
# # Add header
# cms_label = ROOT.TLatex()
# cms_label.SetTextSize(0.04)
# cms_label.DrawLatexNDC(0.16, 0.92, "#bf{CMS Open Data}")
# header = ROOT.TLatex()
# header.SetTextSize(0.03)
# header.DrawLatexNDC(0.63, 0.92, "#sqrt{s} = 8 TeV, L_{int} = 11.6 fb^{-1}")
#
# # Save plot
# d.SaveAs(filename)
# +
# #!top
| RDF/SelectionAnalyzer-20191029Backup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=false editable=false
# Copyright (c) 2020-2021 CertifAI Sdn. Bhd.
#
# This program is part of OSRFramework. You can redistribute it and/or modify
# it under the terms of the GNU Affero General Public License as published by
# the Free Software Foundation, either version 3 of the License, or
# (at your option) any later version.
#
# This program is distributed in the hope that it will be useful,
# but WITHOUT ANY WARRANTY; without even the implied warranty of
# MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the
# GNU Affero General Public License for more details.
#
# You should have received a copy of the GNU Affero General Public License
# along with this program. If not, see <http://www.gnu.org/licenses/>.
# -
# # Perform Classification by using K Nearest Neighbour (KNN)
# +
import numpy as np
from sklearn import datasets
from sklearn import model_selection
from sklearn import metrics
from sklearn import preprocessing
from imblearn.over_sampling import SMOTE
import pandas as pd
import requests as req
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ## Classifying Iris Dataset With KNN
# ### Load Data
# Here we will load the IRIS dataset from scikit-learn. We will be utilizing `iris.data` and `iris.target` as usual for our features and values.
iris = datasets.load_iris()
# As usual `dir(iris)` shows the attributes of the iris datasets.<br> `iris.data.shape` shows the shape of the data.<br>
# `iris.target_names` shows the classes that we want to classify.<br>
# `iris.feature_names` shows the name of features that we are training.
dir(iris)
iris.data.shape
iris.target_names
iris.feature_names
data = iris.data.astype(np.float32)
target = iris.target.astype(np.float32)
# Split data into train and test sets.
X_train, X_test, y_train, y_test = model_selection.train_test_split(
data, target, test_size=0.3, random_state=123
)
X_train.shape, y_train.shape
X_test.shape, y_test.shape
# ### Model Training
# We will use K Nearest Neighbours from scikit learn.
from sklearn.neighbors import KNeighborsClassifier
# Initialize the model.<br>
# Specify the number of neighbors to 3.
model = KNeighborsClassifier(n_neighbors=3)
# Train the model by using train dataset.
# TODO: Enter the code to call fit the training data into the model
model.fit(X_train,y_train)
# ### Evaluation
predictions = model.predict(X_test)
# The method `metrics.confusion_matrix` will visualize the performance of the model through a confusion matrix.
print(metrics.confusion_matrix(y_test,predictions))
metrics.accuracy_score(y_test, predictions)
# ## Classifying Glass Dataset from UCI Machine Learning Repository
# ### Load Data
#
# Here, we load the glass data from UCI ML Repository into a Dataframe using **pandas**.<br> `glass` will be storing the dataset, `description` will store the text with the description of the data.
glass = pd.read_csv(
"https://archive.ics.uci.edu/ml/machine-learning-databases/glass/glass.data",
names=['ID','Refractive Index','Na','Mg','Al','Si','K','Ca','Ba','Fe','Class']
)
description = req.get("https://archive.ics.uci.edu/ml/machine-learning-databases/glass/glass.names").text
print(description)
# The `glass` dataset is a combination of features and categories. From the description, we know that the features that we are interested are in columns **2 - 10**. <br>It is common practice that most of the data have their **expected value/ categories** in the last column, which is also the case in this dataset.<br><br> Using `iloc`, separate the data into `glass_data` which contains features, and `glass_target` which contains expected values/ categories.
glass_data = glass.iloc[:,1:-1]
glass_target = glass.iloc[:,-1]
# Notice that the amounts of data in each class varies too much. This is a showcase of what's called **imbalanced data**.<br><br>
# There are a few ways to tackle this problem. Here, we are choosing to use a method called **oversampling**.<br><br>
# **Oversampling** refers to increasing the number of data points in the minority classes.<br><br>
# There are a few techniques for oversampling:
# 1. Random sampling
# 2. SMOTE: Synthetic Minority Over-sampling Technique
# 3. ADASYN: Adaptive Synthetic Sampling
#
# For more details about oversampling do refer to https://machinelearningmastery.com/random-oversampling-and-undersampling-for-imbalanced-classification/.<br><br>
# In this case, we are going to utilize `SMOTE` as `SMOTE` can avoid overfitting.To oversample the data, we are going to utilize a external library called `imblearn`.<br><i>Note: To install this library, run this command: `pip install imblearn` in command line/ terminal.
# !pip install imblearn
oversample = SMOTE()
glass_data, glass_target = oversample.fit_resample(glass_data,glass_target)
# Split `glass_data` into **test and train data**.<br>Test size = 0.3
X_train2, X_test2, y_train2, y_test2 = model_selection.train_test_split(
glass_data, glass_target, test_size=0.3, random_state=123
)
# Perform **feature scaling** on the `X_train2`,`X_test2` into **`X_train2_scaled`** and **`X_test2_scaled`** respectively.<br>
# <I>Hint: fit_transform on the training data and transform only on the test data
scaler = preprocessing.StandardScaler()
X_train2_scaled = scaler.fit_transform(X_train2)
X_test2_scaled = scaler.transform(X_test2)
# ### Model Training
#
# Initialize KNN Model named `model_2` with `k=3`
model_2 = KNeighborsClassifier(n_neighbors=3)
model_2.fit(X_train2_scaled,y_train2)
# ### Evaluation
#
# Predict the values for the test data and do an **`accuracy test`** and a **`confusion matrix`**.
prediction = model_2.predict(X_test2_scaled)
metrics.accuracy_score(y_test2,prediction)
metrics.confusion_matrix(y_test2,prediction)
# Besides accuracy score and confusion matrix, **precision** and **recall** both provide some insights to any classification model that you're trying to train.<br>
# - **`Precision`** : the percentage of your results which are relevant.
# $$Precision = \frac{TP}{TP+FP}$$
# where: <br>
# $TP$ = True positive<br>
# $FP$ = False positive<br><br>
# - **`Recall`** :the percentage of total relevant results correctly classified by your algorithm.
# $$Recall = \frac{TP}{TP+FN}$$
# where: <br>
# $TP$ = True positive<br>
# $FN$ = False negative<br>
#
print(metrics.recall_score(y_test2,prediction,average=None))
print(metrics.precision_score(y_test2,prediction,average=None))
print(metrics.classification_report(y_test2,prediction))
# Occasionally we want to see if the model is overfit by the training data. In such cases we may try to measure the accuracy of the predictions by the training data itself.<br><br>
# Here we try to compare both the results.<br><br>
# If the accuracy is not that distinct from that of the test data, the model is well-fit.
print(metrics.accuracy_score(y_train2,model_2.predict(X_train2_scaled)))
# ## References
# <NAME> and <NAME> (1998). UCI repository of machine learning databases. University
# of California. [www http://www.ics.uci.edu/∼mlearn/MLRepository.html]
#
# <NAME>. (2019, November 18). Understanding a Classification Report For Your Machine Learning Model. Retrieved August 06, 2020, from https://medium.com/@kohlishivam5522/understanding-a-classification-report-for-your-machine-learning-model-88815e2ce397
| solution/08-0_K Nearest Neighbour(KNN).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.2 64-bit
# name: python3
# ---
# ## 1. Import Libraries
import pandas as pd
import numpy as np
import json
from nocasedict import NocaseDict
import re
import requests
import yaml
import math
import simplejson
# ## 2. Establish Key Variables
# +
# LGBF_DataFile_Url = input("Input Web Address of LGBF Data File")
# secrets_location = input("Enter the full file path for your secrets file"
LGBF_DataFile_Url = "https://www.improvementservice.org.uk/__data/assets/excel_doc/0021/23844/2019-20-lgbf-raw-data-may_2021_refresh.xlsx"
secrets_location = "P:\\My Documents\\Github\\Supporting Files\\secrets.yml"
secrets = open(secrets_location)
secrets = yaml.load(secrets,Loader=yaml.FullLoader)['SharePoint_LGBF']
SP_clientsecret = secrets['client_secret']
SP_tenantid = secrets['tenant_id']
SP_tenant = "stirlingcounciluk"
SP_clientid = secrets['client_id'] + '@' + SP_tenantid
# -
# ## 3. Establish Functions
# +
# Establish function to unpivot data as this will be repeated for each sheet. This function only works with the LGBF data file because the layout of the 4 sheets is the same.
def unpivotData (df) :
# Promote Headers
new_headers = df.iloc[0]
df = df[1:]
df.columns = new_headers
# Unpivot Data - !!!! This is creating additional blank rows for num/den dataframes - look into fix. Currently not causing any real issues (as far as I can tell from basic checks of the resulting csv) as additional blank rows can just be dropped!!!!
df = df.melt(id_vars = ['GSS Code', 'Local Authority'],var_name = 'Attribute',value_name = 'Value')
# For some reason only the indicators coded as corp contain a space between the code text and the number. Added a step here to correct it as it applies to all dataframes and causes an issue with splitting code and period from the single string format it is supplied in due to inconsistency.
df['Attribute'] = df['Attribute'].str.replace('CORP ','CORP')
return df
# Establish function that provides a signed difference between two values.
def distance(Current, Previous):
return (max(Previous, Current) - min(Previous, Current)) * (-1 if Previous > Current else 1)
# -
# ## 4. Establish Key Dataframes
# Get each sheet from the excel file and load them into their own distinct dataframes. Basic transformation steps will be applied here to ensure each dataframe has appropriate headers, data is unpivoted etc. More in depth transformation will be handled later.
# ### 4.1 Establish Dataframes from LGBF Raw Data XLSX File
# +
# Collect the excel file from the url/location provided
dataFile = pd.ExcelFile(LGBF_DataFile_Url)
# Write each sheet within the file to it's own data frame. These will eventually be combined into a single more usable dataframe. When rerunning this notebook be careful of the sheet names the Improvement Service have chosen (in particular trailing spaces).
cashIndicators = pd.read_excel(dataFile,'Cash Indicators')
cashIndicators_NumeratorDenominator = pd.read_excel(dataFile,' Cash Num-Den_Indi')
realIndicators = pd.read_excel(dataFile,'Real Indicators ')
realIndicators_NumeratorDenominator = pd.read_excel(dataFile,'Real Num_Den_Indi')
# Output realIndicators to give an example of the output from this step.
realIndicators
# +
# Unpivot all other dataframes so that they are in a more sensible starting point. Each dataframe will require it's own specific transformations so this is as far as the generic steps applied to all dataframes can go.
cashIndicators = unpivotData(cashIndicators)
realIndicators = unpivotData(realIndicators)
cashIndicators_NumeratorDenominator = unpivotData(cashIndicators_NumeratorDenominator)
realIndicators_NumeratorDenominator = unpivotData(realIndicators_NumeratorDenominator)
# Output realIndicators to give an example of the output from this step.
realIndicators
# -
# ### 4.2 Establish Dataframe from SharePoint Indicator Information List
# #### 4.2.1 Get Auth token using client credentials of established SP App for the LGBF site
# +
# Establish data to be passed as part of POST request to obtain session token from SharePoint
data = {
'grant_type':'client_credentials',
'resource': "00000003-0000-0ff1-ce00-000000000000/" + SP_tenant + ".sharepoint.com@" + SP_tenantid,
'client_id': SP_clientid,
'client_secret': SP_clientsecret,
}
# Establish headers to be passed as part of POST request to obtain session token from SharePoint
headers = {
'Content-Type':'application/x-www-form-urlencoded'
}
# Construct url variable for token request
url = "https://accounts.accesscontrol.windows.net/" + SP_tenantid + "/tokens/OAuth/2"
# Send POST request to obtain session token and then load the response into json_data
r = requests.post(url, data=data, headers=headers)
json_data = json.loads(r.text)
# -
# #### 4.2.2 Get SharePoint list data for Indicator Information list
# +
# Establish headers to be passed as part of GET request to obtain data from SharePoint list. Using the 'access_token' obtained in the previous step
headers = {
'Authorization': "Bearer " + json_data['access_token'],
'Accept':'application/json;odata=verbose',
"Accept-Charset": "utf-8",
'Content-Type': 'application/json;odata=verbose'
}
# Construct url variable for data request
url = "https://" + SP_tenant + ".sharepoint.com/sites/PPA/LGBF/_api/web/lists/GetByTitle('Indicator%20Information')/items"
# Send GET request to obtain list data. Extract and normalize this data into a dataframe.
indicatorInformation = requests.get(url, headers=headers)
indicatorInformation = indicatorInformation.json()
indicatorInformation = pd.json_normalize(indicatorInformation['d']['results'])
indicatorInformation
# -
# #### 4.2.3 Format Dataframe Appropriately
# +
# Drop system columns not needed in dataframe
indicatorInformation = indicatorInformation.drop(columns = ['FileSystemObjectType','Id','ServerRedirectedEmbedUri','ServerRedirectedEmbedUrl','GetDlpPolicyTip.__deferred.uri','FieldValuesAsHtml.__deferred.uri','FieldValuesAsText.__deferred.uri','FieldValuesForEdit.__deferred.uri','File.__deferred.uri','Folder.__deferred.uri','LikedByInformation.__deferred.uri','ParentList.__deferred.uri','Properties.__deferred.uri','Versions.__deferred.uri','Attachments','GUID','__metadata.id','__metadata.uri','__metadata.etag','__metadata.type','FirstUniqueAncestorSecurableObject.__deferred.uri','RoleAssignments.__deferred.uri','AttachmentFiles.__deferred.uri','AttachmentFiles.__deferred.uri','ContentType.__deferred.uri','ID','Modified','Created','AuthorId','EditorId','OData__UIVersionString','ContentTypeId','ComplianceAssetId'])
# Rename columns appropriately
indicatorInformation = indicatorInformation.rename(columns = {'Code_x0028_SortableA_x002d_Z_x00': 'Code_Sortable', 'Ranking_x0020_Type' : 'Ranking_Type', 'NumberFormat_x0028_notext_x0029_' : "NumberFormat_NoText", "GoldilocksRankingMidpoint_x0028_" : "Ranking_GoldilocksMidpoint"})
# Convert column types to ensure that they can be compared to other data later.
indicatorInformation['Code'] = indicatorInformation['Code'].astype(str)
indicatorInformation['Numerator_Match'] = indicatorInformation['Numerator_Match'].astype(str)
indicatorInformation['Denominator_Match'] = indicatorInformation['Denominator_Match'].astype(str)
indicatorInformation
# -
# ### 4.3 Establish Family Group Dataframe
# +
#Get Family Group Table information from the Improvement Service's website
html = requests.get('https://www.improvementservice.org.uk/benchmarking/how-do-we-compare-councils').content
htmlTables = pd.read_html(html)
FamilyGroups_CSWH = htmlTables[0]
FamilyGroups_ECLEDC = htmlTables[1]
#Unpivot the data in ECLEDC
FamilyGroups_ECLEDC = FamilyGroups_ECLEDC.assign(id= 1)
FamilyGroups_ECLEDC = pd.melt(FamilyGroups_ECLEDC, id_vars=['id'], value_vars=['Family Group 1', 'Family Group 2', 'Family Group 3', 'Family Group 4'],var_name='Family_Group', value_name='Local_Authority')
FamilyGroups_ECLEDC = FamilyGroups_ECLEDC.drop(columns = ['id'])
#The table on the Improvement Service's website does not separate the local authority values into separate table rows but instead puts them all in as one piece of text without a delimeter. In the HTML they are separated by </br> but this does not work with pandas read_html. The only delimeter that can be used is the transition from lower case to uppercase. This is because the only time words run together are between local authorities. Here we replace those transitions with ";" so they can be expanded to new rows.
FamilyGroups = []
Local_Authority = []
Type = []
for row in FamilyGroups_ECLEDC.itertuples() :
la_list = re.sub(r'(?<=[a-z])(?=[A-Z])', ';', row.Local_Authority)
Local_Authority.append(la_list)
FamilyGroups.append(row.Family_Group)
Type.append('Environmental, Culture & Leisure, Economic Development, Corporate and Property indicators')
#Create dataframe to contain the results and rename columns appropriately
FamilyGroups_ECLEDC = pd.DataFrame(Local_Authority)
FamilyGroups_ECLEDC = FamilyGroups_ECLEDC.assign(Type = Type)
FamilyGroups_ECLEDC = FamilyGroups_ECLEDC.assign(Family_Group = FamilyGroups)
FamilyGroups_ECLEDC = FamilyGroups_ECLEDC.rename(columns = {0 : 'Local_Authority'})
#Unpivot the data in CSWH
FamilyGroups_CSWH = FamilyGroups_CSWH.assign(id= 1)
FamilyGroups_CSWH = pd.melt(FamilyGroups_CSWH, id_vars=['id'], value_vars=['Family Group 1', 'Family Group 2', 'Family Group 3', 'Family Group 4'],var_name='Family_Group', value_name='Local_Authority')
FamilyGroups_CSWH = FamilyGroups_CSWH.drop(columns = ['id'])
#The table on the Improvement Service's website does not separate the local authority values into separate table rows but instead puts them all in as one piece of text without a delimeter. In the HTML they are separated by </br> but this does not work with pandas read_html. The only delimeter that can be used is the transition from lower case to uppercase. This is because the only time words run together are between local authorities. Here we replace those transitions with ";" so they can be expanded to new rows.
FamilyGroups = []
Local_Authority = []
Type = []
for row in FamilyGroups_CSWH.itertuples() :
la_list = re.sub(r'(?<=[a-z])(?=[A-Z])', ';', row.Local_Authority)
Local_Authority.append(la_list)
FamilyGroups.append(row.Family_Group)
Type.append('Children, Social Work and Housing indicators')
#Create dataframe to contain the results and rename columns appropriately
FamilyGroups_CSWH = pd.DataFrame(Local_Authority)
FamilyGroups_CSWH = FamilyGroups_CSWH.assign(Type = Type)
FamilyGroups_CSWH = FamilyGroups_CSWH.assign(Family_Group = FamilyGroups)
FamilyGroups_CSWH = FamilyGroups_CSWH.rename(columns = {0 : 'Local_Authority'})
#Concatenate dataframes together and expand the delimeted local authorities to new rows.
Family_Groups = pd.concat([FamilyGroups_ECLEDC, FamilyGroups_CSWH])
Family_Groups = Family_Groups.assign(Local_Authority=Family_Groups.Local_Authority.str.split(";")).explode('Local_Authority')
#There is a mismatch between the naming of Edinburgh on the Improvement Services Family Groupings web page and the naming in the raw data file. Replace the text here to allow merges with the raw data file later.
Family_Groups['Local_Authority'] = Family_Groups['Local_Authority'].str.replace('Edinburgh, City of','Edinburgh City')
Family_Groups
# -
# ## 5. Transform Each Dataframe Containing Indicator Data and Merge DataFrames Into One Master DataFrame
# Apply relevant transformation steps to each of the Dataframes below and join them into a single fact table containing Cash Values, Real Values, Cash Numerators, Cash Denominators, Real Numerators and Real Denominators.
# ### 5.1 Transform realIndicators Dataframe
#Import the realIndicators into what will become the final table
all_LGBFData = realIndicators
#Split the Attribute column into its parts using space as a delimeter
all_LGBFData[['Code','Period','Other']] = all_LGBFData['Attribute'].str.split(" ", n=2, expand=True)
#Remove the Attribute column and the redundant other column, rename columns to avoid spaces and to identify the value type as we intend to merge all dataframes into a single dataframe.
all_LGBFData = all_LGBFData.drop(columns = ['Attribute','Other'])
all_LGBFData = all_LGBFData.rename(columns = {'Value': 'Real_Value', 'Local Authority' : 'Local_Authority'})
# Save Scotland values only for merge with averages table later
ISScottishValues_Real = all_LGBFData[(all_LGBFData.Local_Authority == "Scotland")]
#Remove all of the "Scottish Average" Rows as we intend to provide both types of average later in order to provide a more "Complete" data set. Reset the index after this to ensure it is consistent.
all_LGBFData = all_LGBFData[(all_LGBFData.Local_Authority != "Scotland")]
all_LGBFData = all_LGBFData.reset_index(drop=True)
all_LGBFData
# ### 5.2 Transform cashIndicators Dataframe
#Split the Attribute column into its parts using space as a delimeter
cashIndicators[['Code','Period']] = cashIndicators['Attribute'].str.split(" ", n=2, expand=True)
#Remove the Attribute column and the redundant other column, rename columns to avoid spaces and to identify the value type as we intend merge into a single dataframe.
cashIndicators = cashIndicators.drop(columns = ['Attribute'])
cashIndicators = cashIndicators.rename(columns = {'Value': 'Cash_Value', 'Local Authority' : 'Local_Authority'})
#Merge with saved scotland real values to merge with Scottish Averages Dataframe later
ISScottishValues_Cash = cashIndicators[(cashIndicators.Local_Authority == "Scotland")]
ISScottishValues = ISScottishValues_Real.merge(ISScottishValues_Cash, how = 'left', on = ['Local_Authority','Code','Period','GSS Code'], suffixes = ('_Real','_Cash'))
ISScottishValues = ISScottishValues.drop(columns = ['GSS Code','Local_Authority'])
#Remove all of the "Scottish Average" Rows as we intend to provide both types of average later in order to provide a more "Complete" data set. Reset the index after this to ensure it is consistent.
cashIndicators = cashIndicators[(cashIndicators.Local_Authority != "Scotland")]
cashIndicators = cashIndicators.reset_index(drop=True)
cashIndicators
# ### 5.4 Transform NumeratorDenominator Dataframes
# Create a dictionary that can be searched independant of case from the indicatorInformation dataframe. This avoids having to loop the dataframe and exponentially speeds this step up.
indicatorInformation_dict = indicatorInformation.set_index('Code').to_dict('index')
indicatorInformation_dict = NocaseDict(indicatorInformation_dict)
# Display example dictionary item for reference
indicatorInformation_dict['env1a']
# +
# Define function to convert numerator/denominator sheets into column based dataframes that only contain the necessary columns
Types = []
Codes = []
Periods = []
Values = []
PreviousType = None
PreviousAttribute = None
def ConvertNumDenDF(df,prefix) :
#!!! This is a bit hacky - Requires proper fix in unpivot data table!!!
df = df[pd.notnull(df['Attribute'])]
df = df.rename(columns = {'Local Authority' : 'Local_Authority'})
# Filter the dataframe to remove Scotland values as they will be calculated later.
df = df[(df.Local_Authority != "Scotland")]
#Get first word from Attribute column (this will be the code where the value specified is not a numerator or denominator)
df['Code'] = df['Attribute'].str.split(' ').str[0]
df['Year'] = df['Attribute'].str.split(' ').str[1]
df['Value'] = df['Value'].apply(pd.to_numeric,errors='coerce')
#Loop dataframe and establish Value Type, Period and Code based on positioning and checking the code column in the indicatorInformation dataframe. This will have issues if new indicators are introduced and the codes have not yet been added to the Indicator Information SharePoint List. Manual checking required as not sure how to protect against this.
Types = []
Codes = []
Periods = []
Values = []
PreviousType = None
PreviousAttribute = None
for row in df.itertuples():
if row.Attribute == PreviousAttribute:
Types.append(PreviousType)
Codes.append(LastCode)
Periods.append(LastPeriod)
PreviousAttribute = row.Attribute
elif any(row.Code in string for string in indicatorInformation['Code']):
Types.append("Value")
LastCode = row.Code
Codes.append(LastCode)
LastPeriod = row.Year
Periods.append(LastPeriod)
PreviousType = "Value"
PreviousAttribute = row.Attribute
Numerator_Multiplier = indicatorInformation_dict.get(row.Code).get('Numerator_Multipier')
Denominator_Multiplier = indicatorInformation_dict.get(row.Code).get('Denominator_Multiplier')
elif PreviousType == "Value" :
Types.append("Numerator")
Codes.append(LastCode)
Periods.append(LastPeriod)
PreviousType = "Numerator"
PreviousAttribute = row.Attribute
elif PreviousType == "Numerator":
Types.append("Denominator")
Codes.append(LastCode)
Periods.append(LastPeriod)
PreviousType = "Denominator"
PreviousAttribute = row.Attribute
else:
Types.append(None)
Codes.append(None)
Periods.append(None)
PreviousAttribute = row.Attribute
if PreviousType == "Value":
Values.append(row.Value)
elif PreviousType == "Numerator":
Values.append(row.Value * Numerator_Multiplier)
elif PreviousType == "Denominator":
Values.append(row.Value * Denominator_Multiplier)
df['Code'] = Codes
df['Period'] = Periods
df['Type'] = Types
df['Value'] = Values
df = df[['GSS Code','Local_Authority','Code','Period','Type','Value']]
df = df[pd.notnull(df['Value'])]
# Establish new dataframe that contains only the values from realIndicators_NumeratorDenominator that are numerators. Rename the Value column appropriately
df_Numerators = df[(df.Type == "Numerator")]
df_Numerators = df_Numerators.rename(columns = {'Value' : prefix + '_Numerator'})
df_Numerators = df_Numerators.drop('Type', axis=1)
# Establish new dataframe that contains only the values from realIndicators_NumeratorDenominator that are denominators. Rename the Value column appropriately
df_Denominators = df[(df.Type == "Denominator")]
df_Denominators = df_Denominators.rename(columns={'Value': prefix + '_Denominator'})
df_Denominators = df_Denominators.drop('Type', axis=1)
# Merge realIndicators_Numerators and realIndicators_Denominators overwriting the previous dataframe realIndicators_NumeratorDenominator so we have a dataframe of the correct length with two value columns. One for numerator and one for denominator.
df = df_Numerators.merge(df_Denominators, how='left', on=['Local_Authority', 'Code', 'Period', 'GSS Code'], suffixes=('_Num', '_Den'))
return df
# -
# Convert sheets and output one as an example
realIndicators_NumeratorDenominator = ConvertNumDenDF(realIndicators_NumeratorDenominator,'Real')
cashIndicators_NumeratorDenominator = ConvertNumDenDF(cashIndicators_NumeratorDenominator,'Cash')
cashIndicators_NumeratorDenominator
# ### 5.5 Merge all Dataframes into all_LGBFData
# +
# Merge the dataframes together into all_LGBFData. Reorder columns after merge.
all_LGBFData = all_LGBFData.merge(cashIndicators, how = 'left', on = ['Local_Authority','Code','Period','GSS Code'], suffixes = ('_Real','_Cash'))
all_LGBFData = all_LGBFData.merge(realIndicators_NumeratorDenominator, how = 'left', on = ['Local_Authority','Code','Period','GSS Code'], suffixes = ('_all','_realNumDen'))
all_LGBFData = all_LGBFData.merge(cashIndicators_NumeratorDenominator, how = 'left', on = ['Local_Authority','Code','Period','GSS Code'], suffixes = ('_all','_cashNumDen'))
all_LGBFData = all_LGBFData[['GSS Code','Local_Authority','Code','Period','Real_Value','Real_Numerator','Real_Denominator','Cash_Value','Cash_Numerator','Cash_Denominator']]
# Convert value columns to numeric only using errors = 'coerce' to force any text values (dna, *, N/A etc.) to return NaN.
all_LGBFData[['Real_Value','Cash_Value']] = all_LGBFData[['Real_Value','Cash_Value']].apply(pd.to_numeric,errors='coerce')
# Remove rows which contain null or NaN in the Real_Value column as they are non-datapoints and could interfere with average, ranking and quartile calculations later.
all_LGBFData = all_LGBFData[pd.notnull(all_LGBFData['Real_Value'])]
# Convert Percentages to proper decimal percentages
all_LGBFData = all_LGBFData.merge(indicatorInformation[['Code','MeasureType']], how = 'left', on = ['Code'], suffixes = ('_all','_info'))
def ConvertPercent_Real(df) :
if df['MeasureType'] == "Percentage":
return df['Real_Value']/100
else :
return df['Real_Value']
def ConvertPercent_Cash(df) :
if df['MeasureType'] == "Percentage":
return df['Cash_Value']/100
else :
return df['Cash_Value']
all_LGBFData['Real_Value'] = all_LGBFData.apply(ConvertPercent_Real, axis = 1)
all_LGBFData['Cash_Value'] = all_LGBFData.apply(ConvertPercent_Cash, axis = 1)
all_LGBFData = all_LGBFData.drop(columns = ['MeasureType'])
all_LGBFData
# -
# ## 6. Calculate Averages
# There are two types of averages that will be applied to both real and cash values within both the Scottish 32 Councils whole group and the Family Groups for each indicator and period. All averages to be added are as below along with the column names to be assigned.
#
# 1. Scottish Averages
# 1. Average of Local Authority Real Values - ScotAv_LA_Real
# 2. Average of Local Authority Cash Values - ScotAv_LA_Cash
# 3. Sum of Real Numerators Divided by Sum of Real Denominators - ScotAv_NumDen_Real
# 4. Sum of Cash Numerators Divided by Sum of Cash Denominators - ScotAv_NumDen_Cash
# 5. Average of Local Authority Real Numerator Values - ScotAv_LA_Num_Real
# 6. Average of Local Authority Real Denominator Values - ScotAv_LA_Den_Real
# 5. Average of Local Authority Cash Numerator Values - ScotAv_LA_Num_Cash
# 6. Average of Local Authority Cash Denominator Values - ScotAv_LA_Den_Cash
# 2. Family Group Averages
# 1. Average of Local Authority Real Values - FamilyAv_LA_Real
# 2. Average of Local Authority Cash Values - FamilyAv_LA_Cash
# 3. Sum of Real Numerators Divided by Sum of Real Denominators - FamilyAv_NumDen_Real
# 4. Sum of Cash Numerators Divided by Sum of Cash Denominators - FamilyAv_NumDen_Cash
# 5. Average of Local Authority Real Numerator Values - FamilyAv_LA_Num_Real
# 6. Average of Local Authority Real Denominator Values - FamilyAv_LA_Den_Real
# 5. Average of Local Authority Cash Numerator Values - FamilyAv_LA_Num_Cash
# 6. Average of Local Authority Cash Denominator Values - FamilyAv_LA_Den_Cash
# ### 6.1 Scottish Averages
# +
#Sum then divide to get the average resulting from sum of numerator values divided by sum of denominator values for all Real Values.
real_NumeratorDenominatorAverages = realIndicators_NumeratorDenominator.groupby(['Code','Period'], as_index = False).sum()
real_NumeratorDenominatorAverages['ScotAv_NumDen_Real'] = real_NumeratorDenominatorAverages['Real_Numerator'] / real_NumeratorDenominatorAverages['Real_Denominator']
real_NumeratorDenominatorAverages = real_NumeratorDenominatorAverages[['Code','Period','ScotAv_NumDen_Real']]
real_NumeratorDenominatorAverages = real_NumeratorDenominatorAverages.replace([np.inf, -np.inf], np.nan)
#Sum then divide to get the average resulting from sum of numerator values divided by sum of denominator values for all Cash Values.
cash_NumeratorDenominatorAverages = cashIndicators_NumeratorDenominator.groupby(['Code','Period'], as_index = False).sum()
cash_NumeratorDenominatorAverages['ScotAv_NumDen_Cash'] = cash_NumeratorDenominatorAverages['Cash_Numerator'] / cash_NumeratorDenominatorAverages['Cash_Denominator']
cash_NumeratorDenominatorAverages = cash_NumeratorDenominatorAverages[['Code','Period','ScotAv_NumDen_Cash']]
cash_NumeratorDenominatorAverages = cash_NumeratorDenominatorAverages.replace([np.inf, -np.inf], np.nan)
#Calculate mean averages for all other columns and then merge dataframes together to finalise dataframe
ScottishAverages_LA = all_LGBFData.groupby(['Code','Period'], as_index = False).mean()
ScottishAverages = ScottishAverages_LA.merge(real_NumeratorDenominatorAverages, how = 'left', on = ['Code','Period'], suffixes = ('_all','_RealNumDen'))
ScottishAverages = ScottishAverages.merge(cash_NumeratorDenominatorAverages, how = 'left', on = ['Code','Period'], suffixes = ('_all','_CashNumDen'))
# Reorder and rename columns appropriately
ScottishAverages = ScottishAverages.rename(columns = {'Real_Value': 'ScotAv_LA_Real', 'Cash_Value' : 'ScotAv_LA_Cash', 'Real_Numerator' : 'ScotAv_LA_Num_Real', 'Real_Denominator' : 'ScotAv_LA_Den_Real','Cash_Numerator' : 'ScotAv_LA_Num_Cash', 'Cash_Denominator' : 'ScotAv_LA_Den_Cash'})
ScottishAverages = ScottishAverages.merge(indicatorInformation[['Code','Code_Sortable']], how = 'left', on = ['Code'], suffixes = ('_ScotAv','_info'))
ScottishAverages = ScottishAverages[['Code_Sortable','Period','ScotAv_LA_Real','ScotAv_LA_Num_Real','ScotAv_LA_Den_Real','ScotAv_NumDen_Real','ScotAv_LA_Cash','ScotAv_LA_Num_Cash','ScotAv_LA_Den_Cash','ScotAv_NumDen_Cash']]
ScottishAverages = ScottishAverages.rename(columns = {'Code_Sortable':'Code'})
# Add Relationship Key Column
ScottishAverages['Key_CodePeriod'] = ScottishAverages['Code'] + ScottishAverages['Period']
ScottishAverages
# -
# ### 6.2 Family Group Averages
# +
#Merge all_LGBFData with indicatorInformation and Family_Groups to add the additional grouping information that enables Family Group average calculations.
FamilyAverages_LA = all_LGBFData.merge(indicatorInformation[['Code','FamilyGrouping']], how = 'left', on = ['Code'], suffixes = ('_all','_info'))
FamilyAverages_LA = FamilyAverages_LA.rename(columns = {'FamilyGrouping' : 'Type'})
FamilyAverages_LA = FamilyAverages_LA.merge(Family_Groups,how ='left', on = ['Local_Authority','Type'], suffixes = ('_all','_group'))
#Sum then divide to get the average resulting from sum of numerator values divided by sum of denominator values for all Cash Values.
real_NumeratorDenominatorAverages = FamilyAverages_LA.groupby(['Code','Period','Family_Group'], as_index = False).sum()
real_NumeratorDenominatorAverages['FamilyAv_NumDen_Real'] = real_NumeratorDenominatorAverages['Real_Numerator'] / real_NumeratorDenominatorAverages['Real_Denominator']
real_NumeratorDenominatorAverages = real_NumeratorDenominatorAverages[['Code','Period','Family_Group','FamilyAv_NumDen_Real']]
real_NumeratorDenominatorAverages = real_NumeratorDenominatorAverages.replace([np.inf, -np.inf], np.nan)
#Sum then divide to get the average resulting from sum of numerator values divided by sum of denominator values for all Cash Values.
cash_NumeratorDenominatorAverages = FamilyAverages_LA.groupby(['Code','Period','Family_Group'], as_index = False).sum()
cash_NumeratorDenominatorAverages['FamilyAv_NumDen_Cash'] = cash_NumeratorDenominatorAverages['Cash_Numerator'] / cash_NumeratorDenominatorAverages['Cash_Denominator']
cash_NumeratorDenominatorAverages = cash_NumeratorDenominatorAverages[['Code','Period','Family_Group','FamilyAv_NumDen_Cash']]
cash_NumeratorDenominatorAverages = cash_NumeratorDenominatorAverages.replace([np.inf, -np.inf], np.nan)
#Calculate mean averages for all other columns and then merge dataframes together to finalise dataframe
FamilyAverages_LA = FamilyAverages_LA.groupby(['Code','Period','Family_Group'], as_index = False).mean()
FamilyAverages = FamilyAverages_LA.merge(real_NumeratorDenominatorAverages, how = 'left', on = ['Code','Period','Family_Group'], suffixes = ('_all','_RealNumDen'))
FamilyAverages = FamilyAverages.merge(cash_NumeratorDenominatorAverages, how = 'left', on = ['Code','Period','Family_Group'], suffixes = ('_all','_CashNumDen'))
# Reorder and rename columns appropriately
FamilyAverages = FamilyAverages.rename(columns = {'Real_Value': 'FamilyAv_LA_Real', 'Cash_Value' : 'FamilyAv_LA_Cash', 'Real_Numerator' : 'FamilyAv_LA_Num_Real', 'Real_Denominator' : 'FamilyAv_LA_Den_Real','Cash_Numerator' : 'FamilyAv_LA_Num_Cash', 'Cash_Denominator' : 'FamilyAv_LA_Den_Cash'})
FamilyAverages = FamilyAverages.merge(indicatorInformation[['Code','Code_Sortable']], how = 'left', on = ['Code'], suffixes = ('_ScotAv','_info'))
FamilyAverages = FamilyAverages[['Code_Sortable','Period','Family_Group','FamilyAv_LA_Real','FamilyAv_LA_Num_Real','FamilyAv_LA_Den_Real','FamilyAv_NumDen_Real','FamilyAv_LA_Cash','FamilyAv_LA_Num_Cash','FamilyAv_LA_Den_Cash','FamilyAv_NumDen_Cash']]
FamilyAverages = FamilyAverages.rename(columns = {'Code_Sortable':'Code'})
# Add Relationship Key Column
FamilyAverages['Key_CodePeriodFamily_Group'] = FamilyAverages['Code'] + FamilyAverages['Period'] + FamilyAverages['Family_Group']
FamilyAverages
# -
# ## 7 Calculate Ranks & Percentiles
# ### 7.1 Scottish Ranks & Percentiles
# +
#Copy all_LGBFData into ScottishRanks to avoid any steps affecting the all_LGBFData dataframe.
ScottishRanks = all_LGBFData.copy(deep=True)
#Add columns for both ascending and descending ranks and percentiles. The correct versions will be chosen later.
ScottishRanks['ScotRank_Desc'] = all_LGBFData.groupby(['Code','Period'])['Real_Value'].rank('min',ascending = False).astype(int)
ScottishRanks['ScotRank_Asc'] = all_LGBFData.groupby(['Code','Period'])['Real_Value'].rank('min',ascending = True).astype(int)
ScottishRanks['ScotRank_Desc_Pct'] = all_LGBFData.groupby(['Code','Period'])['Real_Value'].rank('min',ascending = False, pct = True).astype(float)
ScottishRanks['ScotRank_Asc_Pct'] = all_LGBFData.groupby(['Code', 'Period'])['Real_Value'].rank('min', ascending=True, pct=True).astype(float)
ScottishRanks = ScottishRanks.merge(indicatorInformation[['Code','Ranking_Type']], how = 'left', on = ['Code'], suffixes = ('_ScotAv','_Info'))
#Merge all_LGBFData with indicatorInformation to get the goldilocks mid points for the few indicators that use them.
GoldilocksScottishRanks = all_LGBFData.merge(indicatorInformation[['Code','Ranking_GoldilocksMidpoint']], how = 'left', on = ['Code'], suffixes = ('_Goldi','_Info'))
#Define function that returns unsigned difference between the real values and the goldilocks mid point. This uses the distance function established at the start of the notebook and then absolutes the values returned from it.
def DifferenceFromGoldilocksMidPoint(df) :
if df['Ranking_GoldilocksMidpoint'] == None :
return None
else :
return abs(distance(df['Real_Value'],df['Ranking_GoldilocksMidpoint']))
#Calculate ranking and percentile based on distance from goldilocks mid point.
GoldilocksScottishRanks['AbsoluteDifferenceFromGoldilocksMidPoint'] = GoldilocksScottishRanks.apply(DifferenceFromGoldilocksMidPoint, axis = 1)
GoldilocksScottishRanks = GoldilocksScottishRanks[pd.notnull(GoldilocksScottishRanks['AbsoluteDifferenceFromGoldilocksMidPoint'])]
GoldilocksScottishRanks['ScotRank_Goldi'] = GoldilocksScottishRanks.groupby(['Code','Period'])['AbsoluteDifferenceFromGoldilocksMidPoint'].rank('min',ascending = True).astype(int)
GoldilocksScottishRanks['ScotRank_Goldi_Pct'] = GoldilocksScottishRanks.groupby(['Code','Period'])['AbsoluteDifferenceFromGoldilocksMidPoint'].rank('min',ascending = True,pct = True).astype(float)
#Merge goldilocks ranking and percentile into ScottishRanks dataframe
ScottishRanks = ScottishRanks.merge(GoldilocksScottishRanks[['Code','Period','Local_Authority','ScotRank_Goldi','ScotRank_Goldi_Pct']], how = 'left', on = ['Code','Period','Local_Authority'], suffixes = ('_ScotRank','_Goldi'))
# Define functions needed to select correct ranking type and percentile type
def ScotRank_select(df) :
if df['Ranking_Type'] == "Ascending" :
return df['ScotRank_Asc']
elif df['Ranking_Type'] == "Descending" :
return df['ScotRank_Desc']
elif df['Ranking_Type'] == "Goldilocks" :
return df['ScotRank_Goldi']
else :
return None
def ScotRank_Pct_select(df) :
if df['Ranking_Type'] == "Ascending" :
return df['ScotRank_Asc_Pct']
elif df['Ranking_Type'] == "Descending" :
return df['ScotRank_Desc_Pct']
elif df['Ranking_Type'] == "Goldilocks" :
return df['ScotRank_Goldi_Pct']
else :
return None
# Apply functions above to create two new columns that contain the correct rank and percentile for each row
ScottishRanks['ScotRank'] = ScottishRanks.apply(ScotRank_select, axis = 1)
ScottishRanks['ScotPct'] = ScottishRanks.apply(ScotRank_Pct_select, axis = 1)
# Add Scottish Ranks to the main fact table
all_LGBFData = ScottishRanks[['GSS Code','Local_Authority','Code','Period','Real_Value','Real_Numerator','Real_Denominator','Cash_Value','Cash_Numerator','Cash_Denominator','ScotRank','ScotPct']]
all_LGBFData
# -
# ### 7.2 Family Group Ranks & Percentiles
# +
#Merge all_LGBFData with additional data from indicatorInformation and Family_Groups dataframes so that the dataframe contains all of the fields required to group for the purposes of ranking.
FamilyRanks_ForGroup = all_LGBFData.merge(indicatorInformation[['Code','FamilyGrouping']], how = 'left', on = ['Code'], suffixes = ('_all','_info'))
FamilyRanks_ForGroup = FamilyRanks_ForGroup.rename(columns = {'FamilyGrouping' : 'Type'})
FamilyRanks_ForGroup = FamilyRanks_ForGroup.merge(Family_Groups,how ='left', on = ['Local_Authority','Type'], suffixes = ('_all','_group'))
#Copy the dataframe to a new variable. THe FamilyRanks_ForGroup dataframe will be used again later to calculate goldilocks ranks and quartiles.
FamilyRanks = FamilyRanks_ForGroup.copy(deep = True)
#Add columns for both ascending and descending ranks and percentiles. The correct versions will be chosen later.
FamilyRanks['FamilyRank_Desc'] = FamilyRanks_ForGroup.groupby(['Code','Period','Family_Group'])['Real_Value'].rank('min',ascending = False).astype(int)
FamilyRanks['FamilyRank_Asc'] = FamilyRanks_ForGroup.groupby(['Code','Period','Family_Group'])['Real_Value'].rank('min',ascending = True).astype(int)
FamilyRanks['FamilyRank_Desc_Pct'] = FamilyRanks_ForGroup.groupby(['Code','Period','Family_Group'])['Real_Value'].rank('min',ascending = False, pct = True).astype(float)
FamilyRanks['FamilyRank_Asc_Pct'] = FamilyRanks_ForGroup.groupby(['Code','Period','Family_Group'])['Real_Value'].rank('min',ascending = True, pct = True).astype(float)
#Merge FamilyRanks_ForGroup with the midpoint for goldilocks ranks into a new dataframe
GoldilocksFamilyRanks = FamilyRanks_ForGroup.merge(indicatorInformation[['Code','Ranking_GoldilocksMidpoint']], how = 'left', on = ['Code'], suffixes = ('_Goldi','_Info'))
#Calculate ranking and percentile based on distance from goldilocks mid point.
GoldilocksFamilyRanks['AbsoluteDifferenceFromGoldilocksMidPoint'] = GoldilocksFamilyRanks.apply(DifferenceFromGoldilocksMidPoint, axis = 1)
GoldilocksFamilyRanks = GoldilocksFamilyRanks[pd.notnull(GoldilocksFamilyRanks['AbsoluteDifferenceFromGoldilocksMidPoint'])]
GoldilocksFamilyRanks['FamilyRank_Goldi'] = GoldilocksFamilyRanks.groupby(['Code','Period','Family_Group'])['AbsoluteDifferenceFromGoldilocksMidPoint'].rank('min',ascending = True).astype(int)
GoldilocksFamilyRanks['FamilyRank_Goldi_Pct'] = GoldilocksFamilyRanks.groupby(['Code','Period', 'Family_Group'])['AbsoluteDifferenceFromGoldilocksMidPoint'].rank('min',ascending = True,pct = True).astype(float)
#Merge relevant columns from GoldilocksFamilyRanks into the main FamilyRanks dataframe.
FamilyRanks = FamilyRanks.merge(GoldilocksFamilyRanks[['Code','Period','Local_Authority','FamilyRank_Goldi','FamilyRank_Goldi_Pct']], how = 'left', on = ['Code','Period','Local_Authority'], suffixes = ('_FamilyRank','_Goldi'))
#Merge the ranking type from the indicatorInformation dataframe. This will allow us to select the correct ranking and percentile in the next steps.
FamilyRanks = FamilyRanks.merge(indicatorInformation[['Code','Ranking_Type']], how = 'left', on = ['Code'], suffixes = ('_FamilyAv','_Info'))
# Define functions needed to select correct ranking type and percentile type
def FamilyRank_select(df) :
if df['Ranking_Type'] == "Ascending" :
return df['FamilyRank_Asc']
elif df['Ranking_Type'] == "Descending" :
return df['FamilyRank_Desc']
elif df['Ranking_Type'] == "Goldilocks" :
return df['FamilyRank_Goldi']
else :
return None
def FamilyRank_Pct_select(df) :
if df['Ranking_Type'] == "Ascending" :
return df['FamilyRank_Asc_Pct']
elif df['Ranking_Type'] == "Descending" :
return df['FamilyRank_Desc_Pct']
elif df['Ranking_Type'] == "Goldilocks" :
return df['FamilyRank_Goldi_Pct']
else :
return None
# Apply functions above to create two new columns that contain the correct rank and percentile for each row
FamilyRanks['FamilyRank'] = FamilyRanks.apply(FamilyRank_select, axis = 1)
FamilyRanks['FamilyPct'] = FamilyRanks.apply(FamilyRank_Pct_select, axis = 1)
# Merge the relevant columns from FamilyRanks into the main all_LGBFData dataframe
FamilyRanks = FamilyRanks[['GSS Code','Local_Authority','Code','Period','FamilyRank','FamilyPct']]
all_LGBFData = all_LGBFData.merge(FamilyRanks,how = 'left', on = ['GSS Code','Local_Authority','Code','Period'], suffixes = ('_ScotAv','_Info'))
all_LGBFData
# -
# ## 8 Adding Previous Period and Initial Row Data & Comparisons
# ### 8.1 Add previous and inital rows as dictionaries in their own columns
# +
#Sort rows of the all_LGBFData dataframe to ensure that we are getting the correct previous and first rows in the subsequent steps. This may not be strictly necessary as the data should already be in the correct sort order. It is here as a safeguard in case any sorting needs to be done in previous steps at a later date.
all_LGBFData.sort_values(by = ['Local_Authority','Code','Period'],inplace = True)
#Define all variables that will be used to record the changes from previous/first.
Previouss = []
Previous = None
Firsts = []
First = None
First_Save = None
Local_Authority = ""
Code = ""
Period = ""
Real_Value = ""
Real_Numerator = ""
Real_Denominator = ""
Cash_Value = ""
Cash_Numerator = ""
Cash_Denominator = ""
ScotRank = ""
ScotPct = ""
FamilyRank = ""
FamilyPct = ""
# Loop over the all_LGBFDataframe and record previous and first into a python dictionary object. Save the objects for each row into the two list variables (Previouss and Firsts)
for row in all_LGBFData.itertuples() :
# If the curently stored Local_Authority and Code are both equal to the current row then this is not the first row for this indicator and local authority combination. As such Previous is calculated using all of the currently stored values in the variables (these are written to at the end of each loop) and First is populated using the stored dictionary in First_Save
if Local_Authority == row.Local_Authority and Code == row.Code :
Previous = {
'Real_Value' : Real_Value,
'Real_Numerator' : Real_Numerator,
'Real_Denominator' : Real_Denominator,
'Cash_Value' : Cash_Value,
'Cash_Numerator' : Cash_Numerator,
'Cash_Denominator' : Cash_Denominator,
'ScotRank' : ScotRank,
'ScotPct' : ScotPct,
'FamilyRank' : FamilyRank,
'FamilyPct' : FamilyPct
}
First = First_Save
# If the curently stored Local_Authority and Code are both not equal to the current row then this is the first row for this indicator and local authority combination. as such the Previous object is set to None and the First object is populated using this rows values.
elif Local_Authority != row.Local_Authority or Code != row.Code :
First_Save = {
'Real_Value' : row.Real_Value,
'Real_Numerator' : row.Real_Numerator,
'Real_Denominator' : row.Real_Denominator,
'Cash_Value' : row.Cash_Value,
'Cash_Numerator' : row.Cash_Numerator,
'Cash_Denominator' : row.Cash_Denominator,
'ScotRank' : row.ScotRank,
'ScotPct' : row.ScotPct,
'FamilyRank' : row.FamilyRank,
'FamilyPct' : row.FamilyPct
}
First = None
Previous = None
# Append the First and Previous into their respective list variables.
Previouss.append(Previous)
Firsts.append(First)
# Set all other variables to their respective columns values in the current row. This is used to both evaluate the if criteria above and to populate the next previous object.
Local_Authority = row.Local_Authority
Code = row.Code
Period = row.Period
Real_Value = row.Real_Value
Real_Numerator = row.Real_Numerator
Real_Denominator = row.Real_Denominator
Cash_Value = row.Cash_Value
Cash_Numerator = row.Cash_Numerator
Cash_Denominator = row.Cash_Denominator
ScotRank = row.ScotRank
ScotPct = row.ScotPct
FamilyRank = row.FamilyRank
FamilyPct = row.FamilyPct
# Assign the two list variables into appropriately titled columns within our all_LGBFData dataframe
all_LGBFData['Previous_Row'] = Previouss
all_LGBFData['First_Row'] = Firsts
all_LGBFData
# -
# ### 8.2 Add comparisons to previous and first data
# +
# Define a function that will return an aim adjusted percentage change between two indicator values. There are two niche cases here. One where previous and current values are both 0 resulting in 0% in all cases. Another where only the previous value is 0 resulting in None being returned as it is not possible to calculate % change from 0. Having looked at the dataset this has only occured 3 times and only affects Orkney and Eilean Siar for CHN20b. Further to this changes in percentage indicators are calculated using 100 as a denominator rather than previous. This is to avoid situations where very small percentages return 1000% or more change (which for our purposes seemed unreasonable to report).
def PercentChange_AimAdjusted (Previous,Current,Code) :
Aim = None
SignedChange = None
PercentChange = None
GoldiMid = None
IsPercentage = False
if Previous == 0 and Current == 0 :
PercentChange = 0
indicatorInfo = indicatorInformation_dict[Code]
Aim = indicatorInfo['Ranking_Type']
GoldiMid = indicatorInfo['Ranking_GoldilocksMidpoint']
IsPercentage = indicatorInfo['MeasureType'] == "Percentage"
if IsPercentage == False and Previous != 0 :
if Aim == "Descending":
SignedChange = distance(Current,Previous)
PercentChange = SignedChange/Previous
if Aim == "Ascending":
SignedChange = -distance(Current,Previous)
PercentChange = SignedChange/Previous
if Aim == "Goldilocks":
Current_DistGoldi = abs(distance(Current,GoldiMid))
Previous_DistGoldi = abs(distance(Previous,GoldiMid))
SignedChange = distance(Previous_DistGoldi,Current_DistGoldi)
PercentChange = SignedChange/Previous_DistGoldi
elif IsPercentage == True and Previous != 0 :
if Aim == "Descending":
SignedChange = distance(Current,Previous)
PercentChange = SignedChange/100
if Aim == "Ascending":
SignedChange = -distance(Current,Previous)
PercentChange = SignedChange/100
if Aim == "Goldilocks":
Current_DistGoldi = abs(distance(Current,GoldiMid))
Previous_DistGoldi = abs(distance(Previous,GoldiMid))
SignedChange = distance(Previous_DistGoldi,Current_DistGoldi)
PercentChange = SignedChange/100
return PercentChange
# Define a function that calculates the changes in the current row by accessing values in the Previous_Row and First_Row python dictionaries.
def Changes(df) :
# Set the intial value of the return variable to none. This allows us to test to see if there were any changes present for a row and then return None instead of a dictionary of None values if not.
Changes = None
# Define all variables that will contain all of the relevant changes for a row.
ScotRank_ChangeSincePrevious = None
ScotPct_ChangeSincePrevious = None
FamilyRank_ChangeSincePrevious = None
FamilyPct_ChangeSincePrevious = None
Real_Value_ChangeSincePrevious = None
Real_Numerator_ChangeSincePrevious = None
Real_Denominator_ChangeSincePrevious = None
Cash_Value_ChangeSincePrevious = None
Cash_Numerator_ChangeSincePrevious = None
Cash_Denominator_ChangeSincePrevious = None
ScotRank_ChangeSinceFirst = None
ScotPct_ChangeSinceFirst = None
FamilyRank_ChangeSinceFirst = None
FamilyPct_ChangeSinceFirst = None
Real_Value_ChangeSinceFirst = None
Real_Numerator_ChangeSinceFirst = None
Real_Denominator_ChangeSinceFirst = None
Cash_Value_ChangeSinceFirst = None
Cash_Numerator_ChangeSinceFirst = None
Cash_Denominator_ChangeSinceFirst = None
PercentChange_AimAdjusted_SincePrevious = None
PercentChange_AimAdjusted_SinceFirst = None
# If the value currently in Previous_Row is not None then there exists a previous object to calculate the changes using.
if df['Previous_Row'] != None :
# Calculate all differences by comparing the current rows value to the same columns value in the Previous_Row dictionary. Ranks and Percentiles are always positive so the calculations are more simple. The other values use the distance function defined at the start of the notebook to determine the signed difference between the values (comparing a current value of -1 to a previous value of 2 will result in -3 difference.)
ScotRank_ChangeSincePrevious = - (df['ScotRank'] - df['Previous_Row'].get('ScotRank'))
ScotPct_ChangeSincePrevious = - (df['ScotPct'] - df['Previous_Row'].get('ScotPct'))
FamilyRank_ChangeSincePrevious = - (df['FamilyRank'] - df['Previous_Row'].get('FamilyRank'))
FamilyPct_ChangeSincePrevious = - (df['FamilyPct'] - df['Previous_Row'].get('FamilyPct'))
Real_Value_ChangeSincePrevious = distance(df['Real_Value'],df['Previous_Row'].get('Real_Value'))
Real_Numerator_ChangeSincePrevious = distance(df['Real_Numerator'],df['Previous_Row'].get('Real_Numerator'))
Real_Denominator_ChangeSincePrevious = distance(df['Real_Denominator'],df['Previous_Row'].get('Real_Denominator'))
Cash_Value_ChangeSincePrevious = distance(df['Cash_Value'],df['Previous_Row'].get('Cash_Value'))
Cash_Numerator_ChangeSincePrevious = distance(df['Cash_Numerator'],df['Previous_Row'].get('Cash_Numerator'))
Cash_Denominator_ChangeSincePrevious = distance(df['Cash_Denominator'],df['Previous_Row'].get('Cash_Denominator'))
PercentChange_AimAdjusted_SincePrevious = PercentChange_AimAdjusted(df['Previous_Row'].get('Real_Value'),df['Real_Value'],df['Code'])
#Set Changes to true to avoid creating a dictionary of None values
Changes = True
# If the value currently in First_Row is not None then there exists a previous object to calculate the changes using.
if df['First_Row'] != None :
# Calculate all differences by comparing the current rows value to the same columns value in the First_Row dictionary. Ranks and Percentiles are always positive so the calculations are more simple. The other values use the distance function defined at the start of the notebook to determine the signed difference between the values (comparing a current value of -1 to a previous value of 2 will result in -3 difference.)
ScotRank_ChangeSinceFirst = - (df['ScotRank'] - df['First_Row'].get('ScotRank'))
ScotPct_ChangeSinceFirst = - (df['ScotPct'] - df['First_Row'].get('ScotPct'))
FamilyRank_ChangeSinceFirst = - (df['FamilyRank'] - df['First_Row'].get('FamilyRank'))
FamilyPct_ChangeSinceFirst = - (df['FamilyPct'] - df['First_Row'].get('FamilyPct'))
Real_Value_ChangeSinceFirst = distance(df['Real_Value'],df['First_Row'].get('Real_Value'))
Real_Numerator_ChangeSinceFirst = distance(df['Real_Numerator'],df['First_Row'].get('Real_Numerator'))
Real_Denominator_ChangeSinceFirst = distance(df['Real_Denominator'],df['First_Row'].get('Real_Denominator'))
Cash_Value_ChangeSinceFirst = distance(df['Cash_Value'],df['First_Row'].get('Cash_Value'))
Cash_Numerator_ChangeSinceFirst = distance(df['Cash_Numerator'],df['First_Row'].get('Cash_Numerator'))
Cash_Denominator_ChangeSinceFirst = distance(df['Cash_Denominator'],df['First_Row'].get('Cash_Denominator'))
PercentChange_AimAdjusted_SinceFirst = PercentChange_AimAdjusted(df['First_Row'].get('Real_Value'),df['Real_Value'],df['Code'])
#Set Changes to true to avoid creating a dictionary of None values
Changes = True
# If there were changes recorded in the previous steps then write these changes into a python dictionary and assign this to Changes
if Changes != None :
Changes = {
"ScotRank_ChangeSincePrevious" : ScotRank_ChangeSincePrevious,
"ScotPct_ChangeSincePrevious" : ScotPct_ChangeSincePrevious,
"FamilyRank_ChangeSincePrevious" : FamilyRank_ChangeSincePrevious,
"FamilyPct_ChangeSincePrevious" : FamilyPct_ChangeSincePrevious,
"ScotRank_ChangeSinceFirst" : ScotRank_ChangeSinceFirst,
"ScotPct_ChangeSinceFirst" : ScotPct_ChangeSinceFirst,
"FamilyRank_ChangeSinceFirst" : FamilyRank_ChangeSinceFirst,
"FamilyPct_ChangeSinceFirst" : FamilyPct_ChangeSinceFirst,
"Real_Value_ChangeSincePrevious" : Real_Value_ChangeSincePrevious,
"Real_Numerator_ChangeSincePrevious" : Real_Numerator_ChangeSincePrevious,
"Real_Denominator_ChangeSincePrevious" : Real_Denominator_ChangeSincePrevious,
"Cash_Value_ChangeSincePrevious" : Cash_Value_ChangeSincePrevious,
"Cash_Numerator_ChangeSincePrevious" : Cash_Numerator_ChangeSincePrevious,
"Cash_Denominator_ChangeSincePrevious" : Cash_Denominator_ChangeSincePrevious,
"Real_Value_ChangeSinceFirst" : Real_Value_ChangeSinceFirst,
"Real_Numerator_ChangeSinceFirst" : Real_Numerator_ChangeSinceFirst,
"Real_Denominator_ChangeSinceFirst" : Real_Denominator_ChangeSinceFirst,
"Cash_Value_ChangeSinceFirst" : Cash_Value_ChangeSinceFirst,
"Cash_Numerator_ChangeSinceFirst" : Cash_Numerator_ChangeSinceFirst,
"Cash_Denominator_ChangeSinceFirst" : Cash_Denominator_ChangeSinceFirst,
"PercentChange_AimAdjusted_SincePrevious" : PercentChange_AimAdjusted_SincePrevious,
"PercentChange_AimAdjusted_SinceFirst" : PercentChange_AimAdjusted_SinceFirst
}
return Changes
#Apply the above function and output it to a new column named Changes.
all_LGBFData['Changes'] = all_LGBFData.apply(Changes, axis = 1)
all_LGBFData
# -
# ### 8.3 Convert Python Dictionaries to JSON
# This is to make the final file more usable in BI products such as PowerBI
# +
def PreviousConvertToJson (df) :
Previous_Row = simplejson.dumps(df['Previous_Row'],ignore_nan=True)
return Previous_Row
def FirstConvertToJson (df) :
First_Row = simplejson.dumps(df['First_Row'],ignore_nan=True)
return First_Row
def ChangesConvertToJson (df) :
Changes = simplejson.dumps(df['Changes'],ignore_nan=True)
return Changes
all_LGBFData['Previous_Row'] = all_LGBFData.apply(PreviousConvertToJson, axis = 1)
all_LGBFData['First_Row'] = all_LGBFData.apply(FirstConvertToJson, axis = 1)
all_LGBFData['Changes'] = all_LGBFData.apply(ChangesConvertToJson, axis = 1)
# Merge sortable codes into dataframe
all_LGBFData = all_LGBFData.merge(indicatorInformation[['Code','Code_Sortable']], how = 'left', on = ['Code'], suffixes = ('ISScot','_info'))
# Add relationship Key Columns
all_LGBFData['Key_CodePeriod'] = all_LGBFData['Code_Sortable'] + all_LGBFData['Period']
all_LGBFData['Key_CodePeriodLA'] = all_LGBFData['Code_Sortable'] + all_LGBFData['Period'] + all_LGBFData['Local_Authority']
all_LGBFData = all_LGBFData.merge(indicatorInformation[['Code_Sortable','FamilyGrouping']], how = 'left', on = ['Code_Sortable'], suffixes = ('_all','_info'))
all_LGBFData = all_LGBFData.rename(columns = {'FamilyGrouping' : 'Type'})
all_LGBFData = all_LGBFData.merge(Family_Groups,how ='left', on = ['Local_Authority','Type'], suffixes = ('_all','_group'))
all_LGBFData['Key_CodePeriodFamilyGroup'] = all_LGBFData['Code_Sortable'] + all_LGBFData['Period'] + all_LGBFData['Family_Group']
# Reorder and rename columns appropriately
all_LGBFData = all_LGBFData[['Key_CodePeriod','Key_CodePeriodFamilyGroup','Key_CodePeriodLA','GSS Code','Local_Authority','Code_Sortable','Period','Real_Value','Real_Numerator','Real_Denominator','Cash_Value','Cash_Numerator','Cash_Denominator','ScotRank','ScotPct','FamilyRank','FamilyPct','Previous_Row','First_Row','Changes']]
all_LGBFData = all_LGBFData.rename(columns = {'Code_Sortable' : 'Code'})
all_LGBFData
# -
# ## 9 Split Last Values Into Separate Dataframe
LastValues = all_LGBFData.copy(deep = True)
LastValues.sort_values(by = ['Code','Period','Local_Authority'],inplace = True)
LastValues = LastValues.groupby(['Code','Local_Authority']).tail(1)
LastValues['Key_CodePeriodLA'] = LastValues['Code'] + LastValues['Period'] + LastValues['Local_Authority']
LastValues = LastValues[['Key_CodePeriodLA','GSS Code','Local_Authority','Code','Period','Real_Value','Real_Numerator','Real_Denominator','Cash_Value','Cash_Numerator','Cash_Denominator','ScotRank','ScotPct','FamilyRank','FamilyPct','Previous_Row','First_Row','Changes']]
LastValues
# ## 10 Format Scottish Values
# +
# Convert value columns to numeric only using errors = 'coerce' to force any text values (dna, *, N/A etc.) to return NaN.
ISScottishValues[['Real_Value','Cash_Value']] = ISScottishValues[['Real_Value','Cash_Value']].apply(pd.to_numeric,errors='coerce')
# Remove rows which contain null or NaN in the Real_Value column as they are non-datapoints and could interfere with average, ranking and quartile calculations later.
ISScottishValues = ISScottishValues[pd.notnull(ISScottishValues['Real_Value'])]
# Convert Percentages to proper decimal percentages
ISScottishValues = ISScottishValues.merge(indicatorInformation[['Code','MeasureType']], how = 'left', on = ['Code'], suffixes = ('_all','_info'))
# These steps use the functions defined in step 5.5
ISScottishValues['Real_Value'] = ISScottishValues.apply(ConvertPercent_Real, axis = 1)
ISScottishValues['Cash_Value'] = ISScottishValues.apply(ConvertPercent_Cash, axis = 1)
ISScottishValues = ISScottishValues.drop(columns = ['MeasureType'])
# Merge sortable codes into dataframe
ISScottishValues = ISScottishValues.merge(indicatorInformation[['Code','Code_Sortable']], how = 'left', on = ['Code'], suffixes = ('ISScot','_info'))
# Add relationship Key Columns
ISScottishValues['Key_CodePeriod'] = ISScottishValues['Code_Sortable'] + ISScottishValues['Period']
# Reorder and rename columns appropriately
ISScottishValues = ISScottishValues[['Key_CodePeriod','Code_Sortable','Period','Real_Value','Cash_Value']]
ISScottishValues = ISScottishValues.rename(columns = {'Real_Value' : 'IS_Scot_Real_Value', 'Cash_Value' : 'IS_Scot_Cash_Value','Code_Sortable' : 'Code'})
ISScottishValues
# -
# ## 11 Basic Verification Checks
# +
# Default position is to assume checks are passed. If any of the checks (excluding numerator denominator checks) are failed below this will be changed and the final csv's will not be output. The numerator denominator errors should be checked at each refresh. The known errors (which exist within the LGBF raw data file) will be identified in the readme in the Error Outputs folder.
ChecksFailed = False
# ScotRank should be between 32 and 1 and should not contain any NaN values
if not all_LGBFData['ScotRank'].between(1,8).any() or all_LGBFData['ScotRank'].isnull().values.any() :
ChecksFailed = True
maxrank = str(max(all_LGBFData['ScotRank']))
minrank = str(min(all_LGBFData['ScotRank']))
countnull = str(all_LGBFData['ScotRank'].isna().sum())
print("ScotRank checks failed : Max - " + maxrank + ", Min - " + minrank + ", Count of Null - " + countnull)
# FamilyRank should be between 8 and 1 and should not contain any NaN values
if not all_LGBFData['FamilyRank'].between(1,8).any() or all_LGBFData['FamilyRank'].isnull().values.any() :
ChecksFailed = True
maxrank = str(max(all_LGBFData['FamilyRank']))
minrank = str(min(all_LGBFData['FamilyRank']))
countnull = str(all_LGBFData['FamilyRank'].isna().sum())
print("FamilyRank checks failed : Max - " + maxrank + ", Min - " + minrank + ", Count of Null - " + countnull)
# Code, Local_Authority, Period, Real_Value and Cash_Value should not contain any null values
if all_LGBFData[['Code','Local_Authority','Period','Real_Value','Cash_Value',]].isnull().values.any() :
ChecksFailed = True
countnullCode = str(all_LGBFData['Code'].isna().sum())
countnullLocal_Authority = str(all_LGBFData['Local_Authority'].isna().sum())
countnullPeriod = str(all_LGBFData['Period'].isna().sum())
countnullReal_Value = str(all_LGBFData['Real_Value'].isna().sum())
countnullCash_Value = str(all_LGBFData['Cash_Value'].isna().sum())
print("Null values found : Code - " + countnullCode + ", Local_Authority - " + countnullLocal_Authority + ", Period - " + countnullPeriod + ", Real_Value - " + countnullReal_Value + ", Cash_Value - " + countnullCash_Value)
# Value should equal numerator/denominator for both cash and real - These errors have been checked and exist in the original raw data file.
NumDenCheck = all_LGBFData.copy(deep = True)
NumDenCheck = NumDenCheck[pd.notnull(NumDenCheck['Real_Numerator'])]
Real_NumDenDivide_Checks = []
Real_NumDenDivide_Check = None
Cash_NumDenDivide_Checks = []
Cash_NumDenDivide_Check = None
FailReferences = []
for row in NumDenCheck.itertuples() :
if row.Real_Value == 0 or math.isnan(row.Real_Denominator) or math.isnan(row.Real_Numerator) :
Real_NumDenDivide_Check = None
else :
if math.isclose(row.Real_Numerator/row.Real_Denominator, row.Real_Value, rel_tol = 0.02):
Real_NumDenDivide_Check = True
else :
Real_NumDenDivide_Check = False
FailReferences.append("Real;" + row.Code + ";" + row.Period + ";" + row.Local_Authority + ";" + str(row.Real_Value) + ";" + str(row.Real_Numerator) + ";" + str(row.Real_Denominator))
if math.isclose(row.Cash_Numerator/row.Cash_Denominator, row.Cash_Value, rel_tol = 0.02):
Cash_NumDenDivide_Check = True
else :
Cash_NumDenDivide_Check = False
FailReferences.append("Cash;" + row.Code + ";" + row.Period + ";" + row.Local_Authority + ";" + str(row.Cash_Value) + ";" + str(row.Cash_Numerator) + ";" + str(row.Cash_Denominator))
Real_NumDenDivide_Checks.append(Real_NumDenDivide_Check)
Cash_NumDenDivide_Checks.append(Cash_NumDenDivide_Check)
if False in Real_NumDenDivide_Checks or False in Cash_NumDenDivide_Checks :
print("Numerator/Denominator values check failed : See Error Outputs for csv of failures")
FailReferences = sorted(list(set(FailReferences)))
FailReferences = pd.DataFrame([sub.split(";") for sub in FailReferences])
FailReferences = FailReferences.rename(columns = {0 : 'Type', 1 : 'Code', 2 : 'Period', 3 : 'Local Authority', 4 : 'Value', 5 : 'Numerator', 6 : 'Denominator'})
FailReferences.to_csv("Error Outputs//Numerator Denominator Fail References.csv", index = False, encoding='utf-8-sig')
# -
# ## 12 Output All Final Tables
if ChecksFailed == False :
all_LGBFData.to_csv("Data Files//Indicator Data.csv", index = False, encoding='utf-8-sig')
LastValues.to_csv("Data Files//Latest Values.csv", index = False, encoding='utf-8-sig')
FamilyAverages.to_csv("Data Files//Family Averages.csv", index = False, encoding='utf-8-sig')
ScottishAverages.to_csv("Data Files//Scottish Averages.csv", index = False, encoding='utf-8-sig')
ISScottishValues.to_csv("Data Files//Scottish Values.csv", index = False, encoding='utf-8-sig')
indicatorInformation.to_csv("Data Files//Indicator Information.csv", index = False, encoding='utf-8-sig')
Family_Groups.to_csv("Data Files//Family Groups.csv", index = False, encoding='utf-8-sig')
else :
print("Checks failed! Check output from section 11 for detail")
| Transform LGBF Data File.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Drich05/Linear-Algebra-58019/blob/main/Practical_Lab_Exam_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="KAChjpWPoLmd" outputId="cb01ac42-ca64-49bf-93f2-668fcb419515"
import numpy as np
A = np.array([[1,2,3],[4,5,6]])
B = np.array([[1,2,],[3,4,],[5,6,]])
C = np.array([[1,2,3],[4,5,6],[7,8,9]])
D = np.array([[1,2,],[3,4,]])
print(A)
print()
print(B)
print()
print(C)
print()
print(D)
print()
dot = np.dot(A,B)
print(dot)
print()
print(D+D)
print()
# + colab={"base_uri": "https://localhost:8080/"} id="wODFtjrhvQRd" outputId="7b31cf12-cee0-4d18-b468-a85e9ea7eb9f"
C = np.array([[1,2,3],[4,5,6],[7,8,9]])
S = 2*C
S
# + colab={"base_uri": "https://localhost:8080/"} id="Vx8Gu_Vuw0cN" outputId="08161e48-7796-4a51-b064-a2ce44e25a49"
import numpy as np
O = np.array([5,3,-1])
print(O)
print()
print('Type:', type(O))
print('Shape:', O.shape)
print('Dimension:', O.ndim)
| Practical_Lab_Exam_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Implement estimators of large-scale sparse Gaussian densities
# #### by <NAME> (email: <EMAIL>, <EMAIL>. Github: <a href="https://github.com/lambday">lambday</a>)<br/> Many many thanks to my mentor <NAME>, <NAME>, <NAME>, <NAME>
# This notebook illustrates large-scale sparse [Gaussian density](http://en.wikipedia.org/wiki/Normal_distribution) [likelihood](http://en.wikipedia.org/wiki/Likelihood_function) estimation. It first introduces the reader to the mathematical background and then shows how one can do the estimation with Shogun on a number of real-world data sets.
# <h2>Theoretical introduction</h2>
# <p><i>Multivariate Gaussian distributions</i>, i.e. some random vector $\mathbf{x}\in\mathbb{R}^n$ having probability density function
# $$p(\mathbf{x}|\boldsymbol\mu, \boldsymbol\Sigma)=(2\pi)^{-n/2}\text{det}(\boldsymbol\Sigma)^{-1/2} \exp\left(-\frac{1}{2}(\mathbf{x}-\boldsymbol\mu)^{T}\boldsymbol\Sigma^{-1}(\mathbf{x}-\boldsymbol\mu)\right)$$
# $\boldsymbol\mu$ being the mean vector and $\boldsymbol\Sigma$ being the covariance matrix, arise in numerous occassions involving large datasets. Computing <i>log-likelihood</i> in these requires computation of the log-determinant of the covariance matrix
# $$\mathcal{L}(\mathbf{x}|\boldsymbol\mu,\boldsymbol\Sigma)=-\frac{n}{2}\log(2\pi)-\frac{1}{2}\log(\text{det}(\boldsymbol\Sigma))-\frac{1}{2}(\mathbf{x}-\boldsymbol\mu)^{T}\boldsymbol\Sigma^{-1}(\mathbf{x}-\boldsymbol\mu)$$
# The covariance matrix and its inverse are symmetric positive definite (spd) and are often sparse, e.g. due to conditional independence properties of Gaussian Markov Random Fields (GMRF). Therefore they can be stored efficiently even for large dimension $n$.</p>
#
# <p>The usual technique for computing the log-determinant term in the likelihood expression relies on <i><a href="http://en.wikipedia.org/wiki/Cholesky_factorization">Cholesky factorization</a></i> of the matrix, i.e. $\boldsymbol\Sigma=\mathbf{LL}^{T}$, ($\mathbf{L}$ is the lower triangular Cholesky factor) and then using the diagonal entries of the factor to compute $\log(\text{det}(\boldsymbol\Sigma))=2\sum_{i=1}^{n}\log(\mathbf{L}_{ii})$. However, for sparse matrices, as covariance matrices usually are, the Cholesky factors often suffer from <i>fill-in</i> phenomena - they turn out to be not so sparse themselves. Therefore, for large dimensions this technique becomes infeasible because of a massive memory requirement for storing all these irrelevant non-diagonal co-efficients of the factor. While ordering techniques have been developed to permute the rows and columns beforehand in order to reduce fill-in, e.g. <i><a href="http://en.wikipedia.org/wiki/Minimum_degree_algorithm">approximate minimum degree</a></i> (AMD) reordering, these techniques depend largely on the sparsity pattern and therefore not guaranteed to give better result.</p>
#
# <p>Recent research shows that using a number of techniques from complex analysis, numerical linear algebra and greedy graph coloring, we can, however, approximate the log-determinant up to an arbitrary precision [<a href="http://link.springer.com/article/10.1007%2Fs11222-012-9368-y">Aune et. al., 2012</a>]. The main trick lies within the observation that we can write $\log(\text{det}(\boldsymbol\Sigma))$ as $\text{trace}(\log(\boldsymbol\Sigma))$, where $\log(\boldsymbol\Sigma)$ is the matrix-logarithm. Computing the log-determinant then requires extracting the trace of the matrix-logarithm as
# $$\text{trace}(\log(\boldsymbol\Sigma))=\sum_{j=1}^{n}\mathbf{e}^{T}_{j}\log(\boldsymbol\Sigma)\mathbf{e}_{j}$$
# where each $\mathbf{e}_{j}$ is a unit basis vector having a 1 in its $j^{\text{th}}$ position while rest are zeros and we assume that we can compute $\log(\boldsymbol\Sigma)\mathbf{e}_{j}$ (explained later). For large dimension $n$, this approach is still costly, so one needs to rely on sampling the trace. For example, using stochastic vectors we can obtain a <i><a href="http://en.wikipedia.org/wiki/Monte_Carlo_method">Monte Carlo estimator</a></i> for the trace -
# $$\text{trace}(\log(\boldsymbol\Sigma))=\mathbb{E}_{\mathbf{v}}(\mathbf{v}^{T}\log(\boldsymbol\Sigma)\mathbf{v})\approx \sum_{j=1}^{k}\mathbf{s}^{T}_{j}\log(\boldsymbol\Sigma)\mathbf{s}_{j}$$
# where the source vectors ($\mathbf{s}_{j}$) have zero mean and unit variance (e.g. $\mathbf{s}_{j}\sim\mathcal{N}(\mathbf{0}, \mathbf{I}), \forall j\in[1\cdots k]$). But since this is a Monte Carlo method, we need many many samples to get sufficiently accurate approximation. However, by a method suggested in Aune et. al., we can reduce the number of samples required drastically by using <i>probing-vectors</i> that are obtained from <a href="http://en.wikipedia.org/wiki/Graph_coloring">coloring of the adjacency graph</a> represented by the power of the sparse-matrix, $\boldsymbol\Sigma^{p}$, i.e. we can obtain -
# $$\mathbb{E}_{\mathbf{v}}(\mathbf{v}^{T}\log(\boldsymbol\Sigma)\mathbf{v})\approx \sum_{j=1}^{m}\mathbf{w}^{T}_{j}\log(\boldsymbol\Sigma)\mathbf{w}_{j}$$
# with $m\ll n$, where $m$ is the number of colors used in the graph coloring. For a particular color $j$, the probing vector $\mathbb{w}_{j}$ is obtained by filling with $+1$ or $-1$ uniformly randomly for entries corresponding to nodes of the graph colored with $j$, keeping the rest of the entries as zeros. Since the matrix is sparse, the number of colors used is usually very small compared to the dimension $n$, promising the advantage of this approach.</p>
#
# <p>There are two main issues in this technique. First, computing $\boldsymbol\Sigma^{p}$ is computationally costly, but experiments show that directly applying a <i>d-distance</i> coloring algorithm on the sparse matrix itself also results in a pretty good approximation. Second, computing the exact matrix-logarithm is often infeasible because its is not guaranteed to be sparse. Aune et. al. suggested that we can rely on rational approximation of the matrix-logarithm times vector using an approach described in <a href="http://eprints.ma.man.ac.uk/1136/01/covered/MIMS_ep2007_103.pdf">Hale et. al [2008]</a>, i.e. writing $\log(\boldsymbol\Sigma)\mathbf{w}_{j}$ in our desired expression using <i><a href="http://en.wikipedia.org/wiki/Cauchy's_integral_formula">Cauchy's integral formula</a></i> as -
# $$log(\boldsymbol\Sigma)\mathbf{w}_{j}=\frac{1}{2\pi i}\oint_{\Gamma}log(z)(z\mathbf{I}-\boldsymbol\Sigma)^{-1}\mathbf{w}_{j}dz\approx \frac{-8K(\lambda_{m}\lambda_{M})^{\frac{1}{4}}}{k\pi N} \boldsymbol\Sigma\Im\left(-\sum_{l=1}^{N}\alpha_{l}(\boldsymbol\Sigma-\sigma_{l}\mathbf{I})^{-1}\mathbf{w}_{j}\right)$$
# $K$, $k \in \mathbb{R}$, $\alpha_{l}$, $\sigma_{l} \in \mathbb{C}$ are coming from <i><a href="http://en.wikipedia.org/wiki/Jacobi_elliptic_functions">Jacobi elliptic functions</a></i>, $\lambda_{m}$ and $\lambda_{M}$ are the minimum/maximum eigenvalues of $\boldsymbol\Sigma$ (they have to be real-positive), respectively, $N$ is the number of contour points in the quadrature rule of the above integral and $\Im(\mathbf{x})$ represents the imaginary part of $\mathbf{x}\in\mathbb{C}^{n}$.</p>
#
# <p>The problem then finally boils down to solving the shifted family of linear systems $(\boldsymbol\Sigma-\sigma_{l}\mathbf{I})\mathbb{x}_{j}=\mathbb{w}_{j}$. Since $\boldsymbol\Sigma$ is sparse, matrix-vector product is not much costly and therefore these systems can be solved with a low memory-requirement using <i>Krylov subspace iterative solvers</i> like <i><a href="http://en.wikipedia.org/wiki/Conjugate_gradient_method">Conjugate Gradient</a></i> (CG). Since the shifted matrices have complex entries along their diagonal, the appropriate method to choose is <i>Conjugate Orthogonal Conjugate Gradient</i> (COCG) [<a href="http://ieeexplore.ieee.org/xpls/abs_all.jsp?arnumber=106415&tag=1"><NAME> et. al., 1990.</a>]. Alternatively, these systems can be solved at once using <i>CG-M</i> [<a href"http://arxiv.org/abs/hep-lat/9612014">Jegerlehner, 1996.</a>] solver which solves for $(\mathbf{A}+\sigma\mathbf{I})\mathbf{x}=\mathbf{b}$ for all values of $\sigma$ using as many matrix-vector products in the CG-iterations as required to solve for one single shifted system. This algorithm shows reliable convergance behavior for systems with reasonable condition number.</p>
#
# <p>One interesting property of this approach is that once the graph coloring information and shifts/weights are known, all the computation components - solving linear systems, computing final vector-vector product - are independently computable. Therefore, computation can be speeded up using parallel computation of these. To use this, a computation framework for Shogun is developed and the whole log-det computation works on top of it.</p>
#
# <h2>An example of using this approach in Shogun</h2>
# <p>We demonstrate the usage of this technique to estimate log-determinant of a real-valued spd sparse matrix with dimension $715,176\times 715,176$ with $4,817,870$ non-zero entries, <a href="http://www.cise.ufl.edu/research/sparse/matrices/GHS_psdef/apache2.html">apache2</a>, which is obtained from the <a href="http://www.cise.ufl.edu/research/sparse/matrices/">The University of Florida Sparse Matrix Collection</a>. Cholesky factorization with AMD for this sparse-matrix gives rise to factors with $353,843,716$ non-zero entries (from source). We use CG-M solver to solve the shifted systems. Since the original matrix is badly conditioned, here we added a ridge along its diagonal to reduce the condition number so that the CG-M solver converges within reasonable time. Please note that for high condition number, the number of iteration has to be set very high.
# +
# %matplotlib inline
from scipy.sparse import eye
from scipy.io import mmread
import numpy as np
from matplotlib import pyplot as plt
import os
import shogun as sg
SHOGUN_DATA_DIR=os.getenv('SHOGUN_DATA_DIR', '../../../data')
matFile=os.path.join(SHOGUN_DATA_DIR, 'logdet/apache2.mtx.gz')
M = mmread(matFile)
rows = M.shape[0]
cols = M.shape[1]
A = M + eye(rows, cols) * 10000.0
plt.title("A")
plt.spy(A, precision = 1e-2, marker = '.', markersize = 0.01)
plt.show()
# -
# First, to keep the notion of Krylov subspace, we view the matrix as a linear operator that applies on a vector, resulting a new vector. We use <a href="http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1SparseMatrixOperator.html">RealSparseMatrixOperator</a> that is suitable for this example. All the solvers work with <a href="http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1LinearOperator.html">LinearOperator</a> type objects. For computing the eigenvalues, we use <a href="http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1LanczosEigenSolver.html">LanczosEigenSolver</a> class. Although computation of the Eigenvalues is done internally within the log-determinant estimator itself (see below), here we explicitely precompute them.
# +
op = sg.RealSparseMatrixOperator(A.tocsc())
# Lanczos iterative Eigensolver to compute the min/max Eigenvalues which is required to compute the shifts
eigen_solver = sg.LanczosEigenSolver(op)
# we set the iteration limit high to compute the eigenvalues more accurately, default iteration limit is 1000
eigen_solver.set_max_iteration_limit(2000)
# computing the eigenvalues
eigen_solver.compute()
print('Minimum Eigenvalue:', eigen_solver.get_min_eigenvalue())
print('Maximum Eigenvalue:', eigen_solver.get_max_eigenvalue())
# -
# Next, we use <a href="http://www.shogun-toolbox.org/doc/en/latest/ProbingSampler_8h_source.html">ProbingSampler</a> class which uses an external library <a href="http://www.cscapes.org/coloringpage/">ColPack</a>. Again, the number of colors used is precomputed for demonstration purpose, although computed internally inside the log-determinant estimator.
# We can specify the power of the sparse-matrix that is to be used for coloring, default values will apply a
# 2-distance greedy graph coloring algorithm on the sparse-matrix itself. Matrix-power, if specified, is computed in O(lg p)
trace_sampler = sg.ProbingSampler(op)
# apply the graph coloring algorithm and generate the number of colors, i.e. number of trace samples
trace_sampler.precompute()
print('Number of colors used:', trace_sampler.get_num_samples())
# <p>This corresponds to averaging over 13 source vectors rather than one (but has much lower variance as using 13 Gaussian source vectors). A comparison between the convergence behavior of using probing sampler and Gaussian sampler is presented later.</p>
#
# <p>Then we define <a href="http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CLogRationalApproximationCGM.html">LogRationalApproximationCGM</a> operator function class, which internally uses the Eigensolver to compute the Eigenvalues, uses <a href="http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CJacobiEllipticFunctions.html">JacobiEllipticFunctions</a> to compute the complex shifts, weights and the constant multiplier in the rational approximation expression, takes the probing vector generated by the trace sampler and then uses CG-M solver (<a href="http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CGMShiftedFamilySolver.html">CGMShiftedFamilySolver</a>) to solve the shifted systems. Precompute is not necessary here too.</p>
# +
cgm = sg.CGMShiftedFamilySolver()
# setting the iteration limit (set this to higher value for higher condition number)
cgm.set_iteration_limit(100)
# accuracy determines the number of contour points in the rational approximation (i.e. number of shifts in the systems)
accuracy = 1E-15
# we create a operator-log-function using the sparse matrix operator that uses CG-M to solve the shifted systems
op_func = sg.LogRationalApproximationCGM(op, eigen_solver, cgm, accuracy)
op_func.precompute()
print('Number of shifts:', op_func.get_num_shifts())
# -
# Finally, we use the <a href="http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1LogDetEstimator.html">LogDetEstimator</a> class to sample the log-determinant of the matrix.
# +
# number of log-det samples (use a higher number to get better estimates)
# (this is 5 times number of colors estimate in practice, so usually 1 probing estimate is enough)
num_samples = 5
log_det_estimator = sg.LogDetEstimator(trace_sampler, op_func)
estimates = log_det_estimator.sample(num_samples)
estimated_logdet = np.mean(estimates)
print('Estimated log(det(A)):', estimated_logdet)
# -
# To verify the accuracy of the estimate, we compute exact log-determinant of A using Cholesky factorization using <a href="http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1Statistics.html#a9931a4ea72310b239efdc05503442525">Statistics::log_det</a> method.
# +
# the following method requires massive amount of memory, for demonstration purpose
# the following code is commented out and direct value obtained from running it once is used
# from shogun import Statistics
# actual_logdet = Statistics.log_det(A)
actual_logdet = 7120357.73878
print('Actual log(det(A)):', actual_logdet)
plt.hist(estimates)
plt.plot([actual_logdet, actual_logdet], [0,len(estimates)], linewidth=3)
plt.show()
# -
# <h2>Statistics</h2>
# We use a smaller sparse-matrix, <a href="http://www.cise.ufl.edu/research/sparse/matrices/HB/west0479.html">'west0479'</a> in this section to demonstrate the benefits of using probing vectors over standard Gaussian vectors to sample the trace of matrix-logarithm. In the following we can easily observe the fill-in phenomena described earlier. Again, a ridge has been added to reduce the runtime for demonstration purpose.
# +
from scipy.sparse import csc_matrix
from scipy.sparse import identity
m = mmread(os.path.join(SHOGUN_DATA_DIR, 'logdet/west0479.mtx'))
# computing a spd with added ridge
B = csc_matrix(m.transpose() * m + identity(m.shape[0]) * 1000.0)
fig = plt.figure(figsize=(12, 4))
ax = fig.add_subplot(1,2,1)
ax.set_title('B')
ax.spy(B, precision = 1e-5, marker = '.', markersize = 2.0)
ax = fig.add_subplot(1,2,2)
ax.set_title('lower Cholesky factor')
dense_matrix = B.todense()
L = np.linalg.cholesky(dense_matrix)
ax.spy(csc_matrix(L), precision = 1e-5, marker = '.', markersize = 2.0)
plt.show()
# +
op = sg.RealSparseMatrixOperator(B)
eigen_solver = sg.LanczosEigenSolver(op)
# computing log-det estimates using probing sampler
probing_sampler = sg.ProbingSampler(op)
cgm.set_iteration_limit(500)
op_func = sg.LogRationalApproximationCGM(op, eigen_solver, cgm, 1E-5)
log_det_estimator = sg.LogDetEstimator(probing_sampler, op_func)
num_probing_estimates = 100
probing_estimates = log_det_estimator.sample(num_probing_estimates)
# computing log-det estimates using Gaussian sampler
from shogun import Statistics
num_colors = probing_sampler.get_num_samples()
normal_sampler = sg.NormalSampler(op.get_dimension())
log_det_estimator = sg.LogDetEstimator(normal_sampler, op_func)
num_normal_estimates = num_probing_estimates * num_colors
normal_estimates = log_det_estimator.sample(num_normal_estimates)
# average in groups of n_effective_samples
effective_estimates_normal = np.zeros(num_probing_estimates)
for i in range(num_probing_estimates):
idx = i * num_colors
effective_estimates_normal[i] = np.mean(normal_estimates[idx:(idx + num_colors)])
actual_logdet = Statistics.log_det(B)
print('Actual log(det(B)):', actual_logdet)
print('Estimated log(det(B)) using probing sampler:', np.mean(probing_estimates))
print('Estimated log(det(B)) using Gaussian sampler:', np.mean(effective_estimates_normal))
print('Variance using probing sampler:', np.var(probing_estimates))
print('Variance using Gaussian sampler:', np.var(effective_estimates_normal))
# +
fig = plt.figure(figsize=(15, 4))
ax = fig.add_subplot(1,3,1)
ax.set_title('Probing sampler')
ax.plot(np.cumsum(probing_estimates)/(np.arange(len(probing_estimates))+1))
ax.plot([0,len(probing_estimates)], [actual_logdet, actual_logdet])
ax.legend(["Probing", "True"])
ax = fig.add_subplot(1,3,2)
ax.set_title('Gaussian sampler')
ax.plot(np.cumsum(effective_estimates_normal)/(np.arange(len(effective_estimates_normal))+1))
ax.plot([0,len(probing_estimates)], [actual_logdet, actual_logdet])
ax.legend(["Gaussian", "True"])
ax = fig.add_subplot(1,3,3)
ax.hist(probing_estimates)
ax.hist(effective_estimates_normal)
ax.plot([actual_logdet, actual_logdet], [0,len(probing_estimates)], linewidth=3)
plt.show()
# -
# <h2>A motivational example - likelihood of the Ozone dataset</h2>
# <p>In <a href="http://arxiv.org/abs/1306.4032">Lyne et. al. (2013)</a>, an interesting scenario is discussed where the log-likelihood of a model involving large spatial dataset is considered. The data, collected by a satellite consists of $N=173,405$ ozone measurements around the globe. The data is modelled using three stage hierarchical way -
# $$y_{i}|\mathbf{x},\kappa,\tau\sim\mathcal{N}(\mathbf{Ax},\tau^{−1}\mathbf{I})$$
# $$\mathbf{x}|\kappa\sim\mathcal{N}(\mathbf{0}, \mathbf{Q}(\kappa))$$
# $$\kappa\sim\log_{2}\mathcal{N}(0, 100), \tau\sim\log_{2}\mathcal{N}(0, 100)$$
# Where the precision matrix, $\mathbf{Q}$, of a Matern SPDE model, defined on a fixed traingulation of the globe, is sparse and the parameter $\kappa$ controls for the range at which correlations in the field are effectively zero (see Girolami et. al. for details). The log-likelihood estiamate of the posterior using this model is
# $$2\mathcal{L}=2\log \pi(\mathbf{y}|\kappa,\tau)=C+\log(\text{det}(\mathbf{Q}(\kappa)))+N\log(\tau)−\log(\text{det}(\mathbf{Q}(\kappa)+\tau \mathbf{A}^{T}\mathbf{A}))− \tau\mathbf{y}^{T}\mathbf{y}+\tau^{2}\mathbf{y}^{T}\mathbf{A}(\mathbf{Q}(\kappa)+\tau\mathbf{A}^{T}\mathbf{A})^{−1}\mathbf{A}^{T}\mathbf{y}$$
# In the expression, we have two terms involving log-determinant of large sparse matrices. The rational approximation approach described in the previous section can readily be applicable to estimate the log-likelihood. The following computation shows the usage of Shogun's log-determinant estimator for estimating this likelihood (code has been adapted from an open source library, <a href="https://github.com/karlnapf/ozone-roulette.git">ozone-roulette</a>, written by <NAME>, one of the authors of the original paper).
#
# <b>Please note that we again added a ridge along the diagonal for faster execution of this example. Since the original matrix is badly conditioned, one needs to set the iteration limits very high for both the Eigen solver and the linear solver in absense of precondioning.</b>
# +
from scipy.io import loadmat
def get_Q_y_A(kappa):
# read the ozone data and create the matrix Q
ozone = loadmat(os.path.join(SHOGUN_DATA_DIR, 'logdet/ozone_data.mat'))
GiCG = ozone["GiCG"]
G = ozone["G"]
C0 = ozone["C0"]
kappa = 13.1
Q = GiCG + 2 * (kappa ** 2) * G + (kappa ** 4) * C0
# also, added a ridge here
Q = Q + eye(Q.shape[0], Q.shape[1]) * 10000.0
plt.spy(Q, precision = 1e-5, marker = '.', markersize = 1.0)
plt.show()
# read y and A
y = ozone["y_ozone"]
A = ozone["A"]
return Q, y, A
def log_det(A):
op = sg.RealSparseMatrixOperator(A)
eigen_solver = sg.LanczosEigenSolver(op)
probing_sampler = sg.ProbingSampler(op)
cgm = sg.CGMShiftedFamilySolver()
cgm.set_iteration_limit(100)
op_func = sg.LogRationalApproximationCGM(op, eigen_solver, cgm, 1E-5)
log_det_estimator = sg.LogDetEstimator(probing_sampler, op_func)
num_estimates = 1
return np.mean(log_det_estimator.sample(num_estimates))
def log_likelihood(tau, kappa):
Q, y, A = get_Q_y_A(kappa)
n = len(y);
AtA = A.T.dot(A)
M = Q + tau * AtA;
# Computing log-determinants")
logdet1 = log_det(Q)
logdet2 = log_det(M)
first = 0.5 * logdet1 + 0.5 * n * np.log(tau) - 0.5 * logdet2
# computing the rest of the likelihood
second_a = -0.5 * tau * (y.T.dot(y))
second_b = np.array(A.T.dot(y))
from scipy.sparse.linalg import spsolve
second_b = spsolve(M, second_b)
second_b = A.dot(second_b)
second_b = y.T.dot(second_b)
second_b = 0.5 * (tau ** 2) * second_b
log_det_part = first
quadratic_part = second_a + second_b
const_part = -0.5 * n * np.log(2 * np.pi)
log_marignal_lik = const_part + log_det_part + quadratic_part
return log_marignal_lik
L = log_likelihood(1.0, 15.0)
print('Log-likelihood estimate:', L)
# -
# <h2>Useful components</h2>
# <p>As a part of the implementation of log-determinant estimator, a number of classes have been developed, which may come useful for several other occassions as well.
# <h3>1. <a href="http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1LinearOperator.html">Linear Operators</a></h3>
# All the linear solvers and Eigen solvers work with linear operators. Both real valued and complex valued operators are supported for dense/sparse matrix linear operators.
# +
dim = 5
np.random.seed(10)
# create a random valued sparse matrix linear operator
A = csc_matrix(np.random.randn(dim, dim))
op = sg.RealSparseMatrixOperator(A)
# creating a random vector
np.random.seed(1)
b = np.array(np.random.randn(dim))
v = op.apply(b)
print('A.apply(b)=',v)
# create a dense matrix linear operator
B = np.array(np.random.randn(dim, dim)).astype(complex)
op = sg.ComplexDenseMatrixOperator(B)
print('Dimension:', op.get_dimension())
# -
# <h3>2. <a href="http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1LinearSolver.html">Linear Solvers</a></h3>
# <p> Conjugate Gradient based iterative solvers, that construct the Krylov subspace in their iteration by computing matrix-vector products are most useful for solving sparse linear systems. Here is an overview of CG based solvers that are currently available in Shogun.</p>
# <h4> <a href="http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CConjugateGradientSolver.html">Conjugate Gradient Solver</a></h4>
# This solver solves for system $\mathbf{Qx}=\mathbf{y}$, where $\mathbf{Q}$ is real-valued spd linear operator (e.g. dense/sparse matrix operator), and $\mathbf{y}$ is real vector.
# +
from scipy.sparse import csc_matrix
from scipy.sparse import identity
# creating a random spd matrix
dim = 5
np.random.seed(10)
m = csc_matrix(np.random.randn(dim, dim))
a = m.transpose() * m + csc_matrix(np.identity(dim))
Q = sg.RealSparseMatrixOperator(a)
# creating a random vector
y = np.array(np.random.randn(dim))
# solve the system Qx=y
# the argument is set as True to gather convergence statistics (default is False)
cg = sg.ConjugateGradientSolver(True)
cg.set_iteration_limit(20)
x = cg.solve(Q,y)
print('x:',x)
# verifying the result
print('y:', y)
print('Qx:', Q.apply(x))
residuals = cg.get_residuals()
plt.plot(residuals)
plt.show()
# -
# <h4><a href="http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1ConjugateOrthogonalCGSolver.html">Conjugate Orthogonal CG Solver</a></h4>
# Solves for systems $\mathbf{Qx}=\mathbf{z}$, where $\mathbf{Q}$ is symmetric but non-Hermitian (i.e. having complex entries in its diagonal) and $\mathbf{z}$ is real valued vector.
# +
# creating a random spd matrix
dim = 5
np.random.seed(10)
m = csc_matrix(np.random.randn(dim, dim))
a = m.transpose() * m + csc_matrix(np.identity(dim))
a = a.astype(complex)
# adding a complex entry along the diagonal
for i in range(0, dim):
a[i,i] += complex(np.random.randn(), np.random.randn())
Q = sg.ComplexSparseMatrixOperator(a)
z = np.array(np.random.randn(dim))
# solve for the system Qx=z
cocg = sg.ConjugateOrthogonalCGSolver(True)
cocg.set_iteration_limit(20)
x = cocg.solve(Q, z)
print('x:',x)
# verifying the result
print('z:',z)
print('Qx:',np.real(Q.apply(x)))
residuals = cocg.get_residuals()
plt.plot(residuals)
plt.show()
# -
# <h4><a href="http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CGMShiftedFamilySolver.html">CG-M Shifted Family Solver</a></h4>
# Solves for systems with real valued spd matrices with complex shifts. For using it with log-det, an option to specify the weight of each solution is also there. The solve_shifted_weighted method returns $\sum\alpha_{l}\mathbf{x}_{l}$ where $\mathbf{x}_{l}=(\mathbf{A}+\sigma_{l}\mathbf{I})^{-1}\mathbf{y}$, $\sigma,\alpha\in\mathbb{C}$, $\mathbf{y}\in\mathbb{R}$.
# +
cgm = sg.CGMShiftedFamilySolver()
# creating a random spd matrix
dim = 5
np.random.seed(10)
m = csc_matrix(np.random.randn(dim, dim))
a = m.transpose() * m + csc_matrix(np.identity(dim))
Q = sg.RealSparseMatrixOperator(a)
# creating a random vector
v = np.array(np.random.randn(dim))
# number of shifts (will be equal to the number of contour points)
num_shifts = 3;
# generating some random shifts
shifts = []
for i in range(0, num_shifts):
shifts.append(complex(np.random.randn(), np.random.randn()))
sigma = np.array(shifts)
print('Shifts:', sigma)
# generating some random weights
weights = []
for i in range(0, num_shifts):
weights.append(complex(np.random.randn(), np.random.randn()))
alpha = np.array(weights)
print('Weights:',alpha)
# solve for the systems
cgm = sg.CGMShiftedFamilySolver(True)
cgm.set_iteration_limit(20)
x = cgm.solve_shifted_weighted(Q, v, sigma, alpha)
print('x:',x)
residuals = cgm.get_residuals()
plt.plot(residuals)
plt.show()
# verifying the result with cocg
x_s = np.array([0+0j] * dim)
for i in range(0, num_shifts):
a_s = a.astype(complex)
for j in range(0, dim):
# moving the complex shift inside the operator
a_s[j,j] += sigma[i]
Q_s = sg.ComplexSparseMatrixOperator(a_s)
# multiplying the result with weight
x_s += alpha[i] * cocg.solve(Q_s, v)
print('x\':', x_s)
# -
# Apart from iterative solvers, a few more triangular solvers are added.
# <h4><a href="http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CDirectSparseLinearSolver.html">Direct Sparse Linear Solver</a></h4>
# This uses sparse Cholesky to solve for linear systems $\mathbf{Qx}=\mathbf{y}$, where $\mathbf{Q}$ is real-valued spd linear operator (e.g. dense/sparse matrix operator), and $\mathbf{y}$ is real vector.
# +
# creating a random spd matrix
dim = 5
np.random.seed(10)
m = csc_matrix(np.random.randn(dim, dim))
a = m.transpose() * m + csc_matrix(np.identity(dim))
Q = sg.RealSparseMatrixOperator(a)
# creating a random vector
y = np.array(np.random.randn(dim))
# solve the system Qx=y
chol = sg.DirectSparseLinearSolver()
x = chol.solve(Q,y)
print('x:',x)
# verifying the result
print('y:', y)
print('Qx:', Q.apply(x))
# -
# <h4><a href="http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CDirectLinearSolverComplex.html">Direct Linear Solver for Complex</a></h4>
# This solves linear systems $\mathbf{Qx}=\mathbf{y}$, where $\mathbf{Q}$ is complex-valued dense matrix linear operator, and $\mathbf{y}$ is real vector.
# +
# creating a random spd matrix
dim = 5
np.random.seed(10)
m = np.array(np.random.randn(dim, dim))
a = m.transpose() * m + csc_matrix(np.identity(dim))
a = a.astype(complex)
# adding a complex entry along the diagonal
for i in range(0, dim):
a[i,i] += complex(np.random.randn(), np.random.randn())
Q = sg.ComplexDenseMatrixOperator(a)
z = np.array(np.random.randn(dim))
# solve for the system Qx=z
solver = sg.DirectLinearSolverComplex()
x = solver.solve(Q, z)
print('x:',x)
# verifying the result
print('z:',z)
print('Qx:',np.real(Q.apply(x)))
# -
# <h3>References</h3>
# <ol>
# <li> <NAME>, <NAME>, <NAME>, <i>Parameter estimation in high dimensional Gaussian distributions</i>. Springer Statistics and Computing, December 2012.</li>
# <li> <NAME>, <NAME> and <NAME>, <i>Computing $A^{\alpha}$, $\log(A)$ and Related Matrix Functions by Contour Integrals</i>, MIMS EPrint: 2007.103</li>
# <li> <NAME>, <i>A Petrov-Galerkin Type Method for Solving $\mathbf{Ax}=\mathbf{b}$ Where $\mathbf{A}$ Is Symmetric Complex</i>, IEEE TRANSACTIONS ON MAGNETICS, VOL. 26, NO. 2, MARCH 1990</li>
# <li> <NAME>, <i>Krylov space solvers for shifted linear systems</i>, HEP-LAT heplat/9612014, 1996</li>
# <li> <NAME>, <NAME>, <NAME>, <NAME>, <NAME><i>Playing Russian Roulette with Intractable Likelihoods</i>,arXiv:1306.4032 June 2013</li>
#
| doc/ipython-notebooks/logdet/logdet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (ox)
# language: python
# name: ox
# ---
# # Calculate network routes between CHTS-derived origins and destinations
#
# ignoring freeways for plausibility and using igraph + multiprocessing for fast simulation
# +
import igraph as ig
import math
import matplotlib.pyplot as plt
import multiprocessing as mp
import networkx as nx
import numpy as np
import osmnx as ox
import pandas as pd
from scipy.stats import ttest_ind, ttest_rel
np.random.seed(0)
weight = 'length'
simulate_all_trips = True
# -
# ## Load network and ODs
# %%time
# load the networks
G_dir = ox.load_graphml('data/network/sf-directed-no-fwy.graphml')
G_undir = ox.load_graphml('data/network/sf-undirected-no-fwy.graphml')
print(nx.is_strongly_connected(G_dir))
print(nx.is_connected(G_undir))
# +
# load the origin-destination node pairs from CHTS
od_dir = pd.read_csv('data/od-directed-no-fwy.csv')
od_undir = pd.read_csv('data/od-undirected-no-fwy.csv')
# confirm the origins and destinations match between the two datasets
# (so we're simulating the same set of trips on each graph)
assert (od_dir['orig']==od_undir['orig']).sum() == len(od_dir) == len(od_undir)
assert (od_dir['dest']==od_undir['dest']).sum() == len(od_dir) == len(od_undir)
print(len(od_dir))
# -
# ## How many trips to simulate
# there are 1,133,333 daily trips that begin/end in SF
if simulate_all_trips:
total_daily_trips = 1133333
multiplier = total_daily_trips / len(od_dir)
multiplier = math.ceil(multiplier)
else:
multiplier = 1
multiplier
od_dir = pd.concat([od_dir]*multiplier, ignore_index=True)
len(od_dir)
od_undir = pd.concat([od_undir]*multiplier, ignore_index=True)
len(od_dir)
# ## Convert networkx graphs to igraph
# save osmid in attributes dict so we can retain it after integer-labeling nodes
for nodeid, data in G_dir.nodes().items():
data['osmid'] = nodeid
for nodeid, data in G_undir.nodes().items():
data['osmid'] = nodeid
# +
G_dir = nx.relabel.convert_node_labels_to_integers(G_dir)
G_dir_nodeid_to_osmid = {data['osmid']:nodeid for nodeid, data in G_dir.nodes().items()}
G_undir = nx.relabel.convert_node_labels_to_integers(G_undir)
G_undir_nodeid_to_osmid = {data['osmid']:nodeid for nodeid, data in G_undir.nodes().items()}
# +
# %%time
# convert directed networkx graph to igraph
G_dir_ig = ig.Graph(directed=True)
G_dir_ig.add_vertices(list(G_dir.nodes()))
G_dir_ig.add_edges(list(G_dir.edges()))
G_dir_ig.vs['osmid'] = list(nx.get_node_attributes(G_dir, 'osmid').values())
G_dir_ig.es[weight] = list(nx.get_edge_attributes(G_dir, weight).values())
assert len(G_dir.nodes()) == G_dir_ig.vcount()
assert len(G_dir.edges()) == G_dir_ig.ecount()
# +
# %%time
# convert undirected networkx graph to igraph
G_undir_ig = ig.Graph(directed=False)
G_undir_ig.add_vertices(list(G_undir.nodes()))
G_undir_ig.add_edges(list(G_undir.edges()))
G_undir_ig.vs['osmid'] = list(nx.get_node_attributes(G_undir, 'osmid').values())
G_undir_ig.es[weight] = list(nx.get_edge_attributes(G_undir, weight).values())
assert len(G_undir.nodes()) == G_undir_ig.vcount()
assert len(G_undir.edges()) == G_undir_ig.ecount()
# -
# # Simulate routes between origins and destinations
# +
def network_distance(G_ig, source, target, weight):
try:
return G_ig.shortest_paths(source=source, target=target, weights=weight)[0][0]
except:
return np.nan
def mp_paths(G_ig, orig, dest, weight, mapper):
sources = (mapper[o] for o in orig)
targets = (mapper[d] for d in dest)
args = ((G_ig, s, t, weight) for s, t in zip(sources, targets))
pool = mp.Pool(mp.cpu_count())
sma = pool.starmap_async(network_distance, args)
results = sma.get()
pool.close()
pool.join()
return results
# -
# %%time
# edges traversed along the real-world directed graph
G_ig = G_dir_ig
df = od_dir
mapper = G_dir_nodeid_to_osmid
w = None
col = 'edges_traversed'
df[col] = mp_paths(G_ig, df['orig'], df['dest'], w, mapper)
# %%time
# edges traversed along the bidirectional undirected graph
G_ig = G_undir_ig
df = od_undir
mapper = G_undir_nodeid_to_osmid
w = None
col = 'edges_traversed'
df[col] = mp_paths(G_ig, df['orig'], df['dest'], w, mapper)
# %%time
# meters traveled along the real-world directed graph
G_ig = G_dir_ig
df = od_dir
mapper = G_dir_nodeid_to_osmid
w = weight
col = 'meters_traveled'
df[col] = mp_paths(G_ig, df['orig'], df['dest'], w, mapper)
# %%time
# meters traveled along the bidirectional undirected graph
G_ig = G_undir_ig
df = od_undir
mapper = G_undir_nodeid_to_osmid
w = weight
col = 'meters_traveled'
df[col] = mp_paths(G_ig, df['orig'], df['dest'], w, mapper)
# # Analysis
#
# ### Topological distance (blocks traversed)
left = od_dir['edges_traversed'].describe().round(2)
left.name = 'edges_traversed_dir'
right = od_undir['edges_traversed'].describe().round(2)
right.name = 'edges_traversed_undir'
pd.concat([left, right], axis='columns')
# are the average blocks-traveled per trip significantly different?
a = od_dir['edges_traversed']
b = od_undir['edges_traversed']
diff = a.mean() - b.mean()
t, p = ttest_rel(a=a, b=b, alternative='greater', nan_policy='omit')
print('n={:}, δ={:0.2f}, t={:0.2f}, p={:0.4f}'.format(len(a), diff, t, p))
bw = 0.4
lw = 1.5
ax = a.plot.kde(label='Real-World Network', lw=lw, c='#666666', bw_method=bw)
ax = b.plot.kde(ax=ax, label='Two-Way Converted', lw=lw, c='k', ls='--', bw_method=bw)
ax.set_ylim(bottom=0)
ax.set_xlim(left=-14, right=100)
ax.set_xlabel('Blocks Traversed in Commute')
ax.set_ylabel('Probability Density')
plt.legend()
plt.show()
# ### Metric distance (meters traveled)
left = od_dir['meters_traveled'].describe().round(2)
left.name = 'meters_traveled_dir'
right = od_undir['meters_traveled'].describe().round(2)
right.name = 'meters_traveled_undir'
pd.concat([left, right], axis='columns')
# are the average meters-traveled per trip significantly different?
a = od_dir['meters_traveled']
b = od_undir['meters_traveled']
diff = a.mean() - b.mean()
t, p = ttest_rel(a=a, b=b, alternative='greater', nan_policy='omit')
print('n={:}, δ={:0.2f}, t={:0.2f}, p={:0.4f}'.format(len(a), diff, t, p))
a.mean() / b.mean()
difference = a - b
difference.describe().round(3)
# what % of trips had an increase between 1-way vs 2-way scenarios
(difference > 0).sum() / len(difference)
# of trips that did increase, what is the mean?
difference[difference > 0].describe().round(2)
ax = difference.plot.hist(bins=30, ec='w', color='#666666', alpha=0.8, zorder=2)
ax.set_xlim(left=0, right=500)
ax.set_ylim(bottom=0)
ax.grid(True)
ax.set_xlabel('Meters saved in trip on two-way network')
plt.show()
# # Surplus VKT and fuel consumption
# converters
miles_to_km = 1.60934 #factor to convert miles to km
mpg_to_kpl = 0.425144 #factor to convert fuel economy miles/gallon to km/liter
lb_per_gal_to_kg_per_liter = 0.119826 #factor to convert emissions from lb/gal to kg/liter
co2_usd_ton = 50 #factor to convert metric tons of co2 to US dollars
# #### Excess VKT per day and per year
#
# What is the daily VKT in San Francisco? There are competing figures out there.
#
# **SFCTA**'s TNCs Today report appears to estimate VMT for trips that both begin and end within the city: "Ride-hail vehicles drive approximately 570,000 vehicle miles within San Francisco on a typical weekday. This accounts for 20 percent of all local daily vehicle miles traveled" from https://www.sfcta.org/tncstoday
#
# **CalTrans**'s 2017 Public Roads Data report appears to estimate all the VMT that occurs on the city's (county's) streets, even if the trip is just passing through SF (9,648,730 miles/day). https://dot.ca.gov/-/media/dot-media/programs/research-innovation-system-information/documents/prd2017.pdf
#
# Finally, SFCTA estimates 1,133,333 occur each day entirely within SF. "On a typical weekday, ride-hail vehicles make more than 170,000 vehicle trips within San Francisco, approximately 12 times the number of taxi trips, representing 15 percent of all intra-San Francisco vehicle trips." from https://www.sfcta.org/tncstoday
#
# CA carbon pricing is contemporaneously \$14.67 per 1000 kg (metric ton): https://www.eia.gov/todayinenergy/detail.php?id=34792 In Europe in 2021, it's around $50.
#sf_daily_vmt = 9648730 #CalTrans estimate (all trips through SF)
sf_daily_vmt = 570000 * 5 #SFCTA estimate (only trips entirely within SF)
sf_daily_vkt = int(sf_daily_vmt * miles_to_km)
sf_daily_vkt
# estimate surplus VKT per day
surplus_vkt_daily = int(sf_daily_vkt - (sf_daily_vkt / (a.mean() / b.mean())))
surplus_vkt_daily
# estimate surplus VKT per year
surplus_vkt_annual = int(surplus_vkt_daily * 365)
surplus_vkt_annual
# #### Excess Fuel Consumption and GHG Emission
#
# Average US fuel economy is 24.7 miles/gallon: https://www.reuters.com/article/us-autos-emissions/u-s-vehicle-fuel-economy-rises-to-record-24-7-mpg-epa-idUSKBN1F02BX
#
# Burning 1 gallon of gasoline releases 20 lbs of CO2: https://www.fueleconomy.gov/feg/contentIncludes/co2_inc.htm
# what is the average US fuel economy in km/liter?
fuel_econ_mpg = 24.7 #per reuters
fuel_econ_kpl = fuel_econ_mpg * mpg_to_kpl
fuel_econ_kpl
# how many surplus liters of fuel does this waste per year?
surplus_fuel_liters = surplus_vkt_annual / fuel_econ_kpl
int(surplus_fuel_liters)
# how many kg of CO2 are released from combusting 1 liter of gasoline?
co2_lbs_per_gal = 20 #per fueleconomy.gov
co2_kg_per_liter = co2_lbs_per_gal * lb_per_gal_to_kg_per_liter
co2_kg_per_liter
# how many excess kg of CO2 are released each year?
co2_excess_kg = int(surplus_fuel_liters * co2_kg_per_liter)
co2_excess_kg
# price in USD for this excess CO2
co2_usd_ton * (co2_excess_kg / 1000)
# ## Save to disk
left = od_dir
right = od_undir[['edges_traversed', 'meters_traveled']]
df = pd.merge(left, right, left_index=True, right_index=True, suffixes=('_dir', '_undir'))
df.head()
# save to disk
df.to_csv('data/od_distances-no-fwy-igraph.csv', index=False, encoding='utf-8')
| analysis/03-test-network-routing-od.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.6.0
# language: julia
# name: julia-0.6
# ---
# # Introduction to DataFrames
# **[<NAME>](http://bogumilkaminski.pl/about/), May 23, 2018**
using DataFrames # load package
# ## Load and save DataFrames
# We do not cover all features of the packages. Please refer to their documentation to learn them.
#
# Here we'll load `CSV` to read and write CSV files and `JLD`, which allows us to work with a Julia native binary format.
using CSV
using JLD
# Let's create a simple `DataFrame` for testing purposes,
x = DataFrame(A=[true, false, true], B=[1, 2, missing],
C=[missing, "b", "c"], D=['a', missing, 'c'])
# and use `eltypes` to look at the columnwise types.
eltypes(x)
# Let's use `CSV` to save `x` to disk; make sure `x.csv` does not conflict with some file in your working directory.
CSV.write("x.csv", x)
# Now we can see how it was saved by reading `x.csv`.
print(read("x.csv", String))
# We can also load it back. `use_mmap=false` disables memory mapping so that on Windows the file can be deleted in the same session.
y = CSV.read("x.csv", use_mmap=false)
# When loading in a `DataFrame` from a `CSV`, all columns allow `Missing` by default. Note that the column types have changed!
eltypes(y)
# Now let's save `x` to a file in a binary format; make sure that `x.jld` does not exist in your working directory.
save("x.jld", "x", x)
# After loading in `x.jld` as `y`, `y` is identical to `x`.
y = load("x.jld", "x")
# Note that the column types of `y` are the same as those of `x`!
eltypes(y)
# Next, we'll create the files `bigdf.csv` and `bigdf.jld`, so be careful that you don't already have these files on disc!
#
# In particular, we'll time how long it takes us to write a `DataFrame` with 10^3 rows and 10^5 columns to `.csv` and `.jld` files. *You can expect JLD to be faster!* Use `compress=true` to reduce file sizes.
bigdf = DataFrame(Bool, 10^3, 10^2)
@time CSV.write("bigdf.csv", bigdf)
@time save("bigdf.jld", "bigdf", bigdf)
getfield.(stat.(["bigdf.csv", "bigdf.jld"]), :size)
# Finally, let's clean up. Do not run the next cell unless you are sure that it will not erase your important files.
foreach(rm, ["x.csv", "x.jld", "bigdf.csv", "bigdf.jld"])
| introductory-tutorials/broader-topics-and-ecosystem/intro-to-julia-DataFrames/04_loadsave.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import h5py
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import random
fname_long_term = 'E:\Dima\PhD\Papers\data\states\states_model_80_1_80_500_long_term.h5py'
fname_original = 'E:\Dima\PhD\Papers\data\states\states_model_80_1_80_500.h5py'
tokens_file = 'E:/Dima/PhD/Papers/data/states/tokens_index_model_80_1_80_500_long_term.csv'
m = h5py.File(fname_long_term, 'r')
f = h5py.File(fname_original, 'r')
df_tokens = pd.read_csv(tokens_file)
df_tokens.head()
p_long_term = max(m['y_' + str(20)][()])
p_original = max(f['y_' + str(20)][()])
# +
k = 0
prob_original_list = []
prob_long_term_list = []
p5_satisfies = []
for elem_id, row in df_tokens.iterrows():
elem = row[1] # long term string
p_long_term = max(m['y_' + str(elem_id)][()])
p_original = max(f['y_' + str(elem_id)][()])
if (p_original > 0.7) & (p_long_term > 0.65):
p5_satisfies.append(1)
else:
p5_satisfies.append(0)
prob_long_term_list.append(p_long_term)
prob_original_list.append(p_original)
k = k + 1
# if k % 1000 == 0:
# print("elem = {}. k = {}.".format(elem, k))
# -
len(f['h_' + str(elem_id)][()][0][0])
len(p5_satisfies)
2232/580741*100
df = pd.DataFrame({'p5_satisfies':p5_satisfies})
df.p5_satisfies.value_counts()
56371/df_tokens.shape[0]*100
# # SMC
def get_nu_rho(n_satisfies, n_sampled_so_far, alpha, beta):
ro = (n_satisfies + alpha) / (n_sampled_so_far + alpha + beta)
nu = np.sqrt(((alpha + n_satisfies) * (n_sampled_so_far - n_satisfies + beta)) / (
pow((alpha + n_sampled_so_far + beta), 2) * (alpha + n_sampled_so_far + beta + 1)))
return ro, nu
# +
# calculate time
elements_num = df_tokens.shape[0]
tmp = list(range(0, elements_num))
tokens_indx_rand = tmp.copy()
random.seed(42)
random.shuffle(tokens_indx_rand)
alpha = 1
beta = 1
n_satisfies_p5_list = []
n_sampled_so_far_list = []
smc_ro_estimates_p5 = []
smc_nu_estimates_p5 = []
n_satisfies_p5 = 0
n_sampled_so_far = 0
k = 0
for i in tokens_indx_rand:
# TODO: vectorize this function using apply()
# should be cumulative group by operation
if p5_satisfies[i] == 1:
n_satisfies_p5 = n_satisfies_p5 + 1
n_sampled_so_far = n_sampled_so_far + 1
rho_p5, nu_p5 = get_nu_rho(n_satisfies_p5, n_sampled_so_far, alpha, beta)
n_satisfies_p5_list.append(n_satisfies_p5)
n_sampled_so_far_list.append(n_sampled_so_far)
smc_ro_estimates_p5.append(rho_p5)
smc_nu_estimates_p5.append(nu_p5)
# if k % 1000 == 0:
# print(" k = {}.".format(k))
# k = k + 1
_df_p5_smc = pd.DataFrame({'token_id': tokens_indx_rand,
'satisfies_p5': n_satisfies_p5_list,
'sampled': n_sampled_so_far_list,
'rho_p5': smc_ro_estimates_p5,
'nu_p5': smc_nu_estimates_p5
})
# -
_df_p5_smc.shape
model_base_name = 'model_80_1_80_500'
verify_results_folder='E:/Dima/PhD/Papers/data/verify_results/'
_df_p5_smc.to_csv(verify_results_folder + "/" + model_base_name + "_p5_smc.csv", index=False)
| long_term_relationship.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
# !cat examples/ex5.csv
sentinels = {'message': ['world', 'NA'], 'something': ['three']}
pd.read_csv('examples/ex5.csv', na_values=sentinels)
frame2 = pd.DataFrame({'a': np.random.randn(100)})
frame2
store2 = pd.HDFStore('mydata2.h5')
store2
store2['obj2'] = frame2 #work like dict
store2
store2['obj2']
store2['obj2_col'] = frame2['a']
store2['obj2_col']
| ch06_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Customer satisfaction prediction
# ## Dataset structure
# | n | Variable | Description |
# | :--- | :---------------------------- | :----------------------------------------------------------- |
# | 1 | id | Client ID |
# | 2 | Gender | Client Gender |
# | 3 | Customer type | Client type: Premium or Standard |
# | 4 | Age | Client Age |
# | 5 | Price | Client Age |
# | 6 | New/Used | Client Age |
# | 7 | Category | Client Age |
# | 8 | Product description accuracy | Level of satisfaction on product description |
# | 9 | Manufacturer stainability | Level of satisfaction on the manufacturing sustainability process |
# | 10 | Packaging quality | Level of satisfaction on packaging |
# | 11 | Additional options | Level of satisfaction on extra options |
# | 12 | Reviews and ratings | Level of satisfaction on reviews and rating information |
# | 13 | Integrity of packaging | Level of satisfaction on packaging state |
# | 14 | Check-out procedure | Level of satisfaction on payment procedure |
# | 15 | Relevance of related products | Level of satisfaction on related product suggestion |
# | 16 | Costumer insurance | Level of satisfaction on insurance options |
# | 17 | Shipping delay in days | Delay of shipping in days |
# | 18 | Arrival delay in days | Arrival delay on days |
# | 19 | Satisfaction | Target: Satisfied, Not Satisfied |
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
df = pd.read_csv('../data/processed/final.csv', index_col=0)
df.head()
df_for_pca = df.iloc[:,:-1]
# ***
# ## Principal components analysis
import seaborn as sns
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(df_for_pca)
df_pca_scaled = scaler.transform(df_for_pca)
pca = PCA()
pca.fit(df_pca_scaled)
df_pca = pd.DataFrame(pca.transform(df_pca_scaled))
explained_var=pd.DataFrame(pca.explained_variance_ratio_).transpose()
explained_var.columns = ['PC1','PC2','PC3','PC4','PC5','PC6', 'PC7', 'PC8', 'PC9', 'PC10', 'PC11', 'PC12', 'PC13', 'PC14']
explained_var
cum_explained_var=np.cumsum(pca.explained_variance_ratio_)
cum_explained_var = pd.DataFrame(cum_explained_var)
cum_explained_var.columns = ['CUMSUM']
cum_explained_var
sns.set_style('white')
fig = plt.figure(1, figsize=(12,6))
ax = sns.barplot(data=explained_var, palette = 'viridis')
ax.set_ylabel('Explained Var')
ax2 = plt.twinx()
sns.lineplot(x = cum_explained_var.index, y = 'CUMSUM', data = cum_explained_var, ax = ax2, color = 'orange', marker = 'o', markersize = 9, linewidth = 2.5, label = 'CumSum');
plt.legend()
plt.savefig('../reports/figures/cumsum.png', bbox_inches = 'tight')
plt.show()
pd.DataFrame(pca.components_,index=['PC1','PC2','PC3','PC4','PC5','PC6', 'PC7', 'PC8', 'PC9', 'PC10', 'PC11', 'PC12', 'PC13', 'PC14'],columns=df.iloc[:,:-1].columns)
# >There is no semantic meaning for new components.
df_pca.columns=['PC1','PC2','PC3','PC4','PC5','PC6', 'PC7', 'PC8', 'PC9', 'PC10', 'PC11', 'PC12', 'PC13', 'PC14']
df_pca = pd.concat([df_pca, df['Satisfaction']], axis = 1)
df_pca
df_pca.to_csv('../data/processed/model_PCA.csv')
sns.scatterplot(x = 'PC1', y=[0]*(df_pca['Satisfaction'].size), data=df_pca, hue = 'Satisfaction', alpha = .8);
sns.scatterplot(x = 'PC1', y = 'PC2', data=df_pca, hue = 'Satisfaction', alpha = .8);
# +
from mpl_toolkits.mplot3d import axes3d
fig = plt.figure(figsize = (10,10))
ax = fig.add_subplot(111, projection='3d')
ax.scatter(df_pca['PC1'], df_pca['PC2'],df_pca['PC3'], c=df_pca['Satisfaction'], cmap = 'viridis', s=40, alpha = .8)
ax.set_xlabel('PC1')
ax.set_ylabel('PC2')
ax.set_zlabel('PC3')
ax.view_init(60, 60)
plt.show()
# -
| notebooks/2.1-fc-PCA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow2_p36
# language: python
# name: conda_tensorflow2_p36
# ---
# # Bring Your Own Model を SageMaker で hosting する
# * 先程学習したモデルを Notebook インスタンスで読み込み、改めて tensorflow で save し、自分で作成したモデルとする
# * 自作モデルを SageMaker で hosting & 推論する
#
# ## 処理概要
# * 先程学習したモデルを Notebook インスタンスにダウンロード
# * TensorFlow で読み込み、推論し、改めて保存し直す
# * 保存しなおしたモデルを S3 にアップロードする
# * モデルを hosting する
#
# 
# notebook のセルの横方向の表示範囲を広げる
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
import sagemaker, yaml, tarfile, boto3, os, json, boto3
import tensorflow as tf
from matplotlib import pyplot as plt
import numpy as np
from sagemaker.tensorflow import TensorFlowModel
print(f'Current tensorflow Version ={tf.__version__}')
# ## 設定読み込み
with open('./setting.yaml', 'r') as yml:
config = yaml.load(yml)
best_model_uri = config['best_model_uri']
name = config['name']
timestamp = config['timestamp']
print(best_model_uri)
# ## モデルを Notebook インスタンスにダウンロード
sagemaker.s3.S3Downloader.download(
s3_uri=best_model_uri,
local_path='./model/'
)
# tar.gz の解凍
with tarfile.open('./model/model.tar.gz') as tar:
tar.extractall('./model/')
# ## モデルの読み込み
# * SageMaker Training で学習したものは当然ながら TensorFlow で学習したものと同一
# * load してそのまま使えるかを確認する
model = tf.keras.models.load_model('./model/000000001/')
model.summary()
test_x = np.load('./test_x.npy')
plt.imshow(test_x[0,:,:,0],'gray')
np.argmax(model.predict(test_x[0:1,:,:,:]))
# ## ダウンロードしたモデルを削除し、改めて保存しなおす
# * tensorflow の save を利用して保存した後、tar.gz に圧縮する
# !rm -r ./model
model_dir = './000000002'
tar_name = os.path.join(model_dir, 'model.tar.gz')
model.save(model_dir)
with tarfile.open(tar_name, mode='w:gz') as tar:
tar.add(model_dir)
# ## tar.gz に圧縮したモデルを S3 にアップロードする
sess = sagemaker.session.Session()
bucket = sess.default_bucket()
model_s3_path = f's3://{bucket}/{name}-model-{timestamp}'
print(model_s3_path)
model_s3_uri = sagemaker.s3.S3Uploader.upload(
local_path = tar_name,
desired_s3_uri = model_s3_path
)
print(model_s3_uri)
# ## S3 にアップロードしたモデルを SageMaker 管理のモデルとして登録する
# * TensorFlow で作成したモデルは [TensorFlowModel](https://sagemaker.readthedocs.io/en/stable/frameworks/tensorflow/sagemaker.tensorflow.html?highlight=TensorFlowModel#sagemaker.tensorflow.model.TensorFlowModel) で読み込む
# * 推論に利用する TensorFlow のコンテナイメージを [sagemaker.image_uris.retrieve](https://sagemaker.readthedocs.io/en/stable/api/utility/image_uris.html?highlight=sagemaker.image_uris.retrieve#sagemaker.image_uris.retrieve) で事前に取得しておく(SageMaker にこのモデルは TensorFlow の 2.1 で作られたものであることを報せる必要がある)
container_image_uri = sagemaker.image_uris.retrieve(
"tensorflow",
boto3.Session().region_name,
version='2.1',
instance_type = 'ml.m5.large',
image_scope = 'inference'
)
print(container_image_uri)
tf_model = TensorFlowModel(
model_data=model_s3_uri,
role=sagemaker.get_execution_role(),
image_uri = container_image_uri,
)
# ## hosting して推論の確認を行う
# %%time
predictor = tf_model.deploy(
initial_instance_count=1,
instance_type='ml.m5.large',
)
np.argmax(predictor.predict(test_x[0:1,:,:,:])['predictions'])
# ## 推論のみを行う場合
# * 毎回モデルを読み込んで推論するわけではなく、ほとんどのケースでは モデルを hosting している endpoint で推論のみを行う
# * 今回は sagemaker sdk を利用して推論のみをする場合と、boto3 を用いて推論する場合の 2 種類を試す
# * 推論をするのに必要な情報は endpoint_name と推論するデータのみ
# endpoint_name を予め取得する (マネジメントコンソールでも確認可能)
endpoint_name = predictor.endpoint_name
print(endpoint_name)
# ### sagemaker SDK を利用する場合
predictor2 = sagemaker.predictor.Predictor(
endpoint_name,
serializer=sagemaker.serializers.JSONSerializer(),
deserializer=sagemaker.deserializers.JSONDeserializer(),
)
np.argmax(predictor2.predict(test_x[0:1,:,:,:])['predictions'])
# ### boto3 を利用する場合
client = boto3.client('sagemaker-runtime')
response = client.invoke_endpoint(
EndpointName=endpoint_name,
Body=json.dumps({"instances": test_x[0:1,:,:,:].tolist()}),
ContentType='application/json'
)
np.argmax(np.array(json.load(response['Body'])['predictions'][0]))
| 3_byom.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Introduction à Python
# -
# > présentée par <NAME>
# + [markdown] slideshow={"slide_type": "slide"}
# Ce n'est pas toujours très pratique d'enregistrer les données directement dans le code *(hard-coding)*. Il est souvent plus facile de lire les données depuis un fichier (comme un tableur au format csv).
# + [markdown] slideshow={"slide_type": "fragment"}
# Nous allons essayer d'écrire la liste des personnes avec lesquelles nous avons travaillé précédemment dans un fichier. Puis nous allons essayer de reconstruire l'information pour la manipuler avec python.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Manipulation de fichiers
# + [markdown] slideshow={"slide_type": "fragment"}
# Python fournit nativement une fonction `open` pour manipuler les fichiers. Elle retourne un `file object` qui dispose de certaines méthodes spécifiques, un peu comme nos objets *Personne* avec leur méthode `age()`. La fonction `open()` prend le nom du fichier comme premier argument arbitraire, et un tas d'options dont la plus utile est le mode de lecture du fichier (qui va définir la façon dont python va pouvoir manipuler le fichier : *read*, *write*, *append*)
# + [markdown] slideshow={"slide_type": "skip"}
# Note : [voir la liste de toutes les fonctions natives de python](https://docs.python.org/3/library/functions.html)
# + [markdown] slideshow={"slide_type": "skip"}
# Note : [voir la documentation de la fonction `open()`](https://docs.python.org/3/library/functions.html#open)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Ecriture dans un fichier
# + slideshow={"slide_type": "fragment"}
# solution naive
fichier = open("data/fichiers/nom_du_fichier.txt", mode="w") # mode write
fichier.write("La première ligne d'une longue histoire...") # là, on appelle la méthode write de l'object de type 'file object'
fichier.close() # on ferme le fichier
# + [markdown] slideshow={"slide_type": "fragment"}
# La fermeture du fichier est importante pour libérer immédiatement les ressources systèmes utilisées par celui-ci.
# + [markdown] slideshow={"slide_type": "fragment"}
# Note : Si vous ne fermez pas explicitement un fichier, le *garbage collector* de Python (une stratégie de python qui s'occupe de la gestion de la mémoire) finit par détruire l'objet et fermer le fichier ouvert, mais le fichier peut rester ouvert pendant un certain temps. Le risque associé est que différentes implémentations de Python effectuent ce nettoyage à différents moments, ce qui signifie que votre programme peut ne plus marcher d'une version à l'autre...
# + [markdown] slideshow={"slide_type": "slide"}
# ### Petite disgression et introduction à la notion de contexte
# + [markdown] slideshow={"slide_type": "fragment"}
# Comme c'est un peu pénible d'appeler des fonctions de manière répétitive (ou tout de moins celles auxquelles on ne pense pas toujours...), python a rajouté le mot clé natif `with` qui permet de gérer un contexte. `with` appelle la fonction `__enter__()` d'un objet quand on entre dans le contexte, et sa fonction `__exit__()` quand on en sort.
# + [markdown] slideshow={"slide_type": "subslide"}
# On pourrait voir la fonction open comme un objet implémenté commme ci:
# ```py
# class File(object):
# # Une fausse implémentation de la classe File, juste pour comprendre
#
# def __init__(self, filename, mode="r"):
# self.filename = filename
# self.mode = mode
#
# def __enter__(self):
# self.open_file = open(self.filename, self.mode)
# return self.open_file
#
# def __exit__(self):
# self.open_file.close()
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# Bref, voici comment utiliser un contexte avec les fichiers
# + slideshow={"slide_type": "fragment"}
# solution élégante
with open("data/fichiers/nom_du_fichier.txt", mode="w") as fichier:
fichier.write("La première ligne d'une longue histoire...")
# Une fois qu'on sort du contexte, le fichier est fermé. Note: la variable 'fichier' existe toujours
# + slideshow={"slide_type": "fragment"}
with open("data/fichiers/nom_du_fichier.txt", mode="w") as fichier:
fichier.write("La première ligne d'une longue histoire...")
print("Est-ce que le fichier est fermé dans le contexte ? {}".format(fichier.closed))
print("Est-ce que le fichier est fermé en dehors du contexte ? {}".format(fichier.closed))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Lecture d'un fichier
# + slideshow={"slide_type": "fragment"}
# solution naive
fichier = open("data/fichiers/nom_du_fichier.txt", "r")
contenu = fichier.read()
fichier.close()
contenu
# + slideshow={"slide_type": "fragment"}
# solution élégante
with open("data/fichiers/nom_du_fichier.txt", "r") as fichier:
contenu = fichier.read()
print(contenu)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Les exceptions
# + [markdown] slideshow={"slide_type": "fragment"}
# > Qu'arrive-t-il si le fichier n'existe pas ?
# + [markdown] slideshow={"slide_type": "fragment"}
# Python génère une erreur. On les appelle des exceptions. Si vous n'en n'avez pas encore rencontré :
# - soit vous suivez un tutoriel qui vous cache la véritée d'une vie de développeur.
# - soit vous n'avez pas encore essayer de programmer par vous même.
#
# Note : nous avions rencontrée une exception à la fin du premier chapitre. Cette introduction à python est clairement bien faite!
# + [markdown] slideshow={"slide_type": "subslide"}
# > Comment faire en sorte pour que, dans le pire des cas (le fichier n'existe pas), on soit capable d'afficher la valeur de la variable `contenu` ?
# + slideshow={"slide_type": "fragment"}
try: # Exécute les déclarations suivantes.
with open("un_nom_de_fichier_qui_n_existe_pas.txt", "r") as fichier:
contenu = fichier.read()
except FileNotFoundError as error: # Exécute les déclarations suivantes si une exception de type FileNotFoundError est soulevée.
contenu = None
print("Contenu : {}".format(contenu))
# + [markdown] slideshow={"slide_type": "slide"}
# [Prochain chapitre : Les snippets en pratique](08_Snippets_en_pratique.ipynb)
| 07_Snippets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bingo Tutorial 2: Zero Min Problem
#
# ## Goal: Find a list of numbers with zero magnitude through genetic optimization
# ### Chromosome
# The basic unit of bingo evolutionary analyses are Chromosomes. The chromosome used in this example is a `MultipleFloatChromosome`. The `MultipleFloatChromosome` contains a list of floating point values. It also has optional use of local optimization for some of those values.
from bingo.chromosomes.multiple_floats import MultipleFloatChromosome
chromosome = MultipleFloatChromosome([0., 1., 2., 3.])
print(type(chromosome))
print(chromosome)
# ### Chromosome Generator
# Chromosomes are created with a Generator. Generation of `MultipleValueChromosome` requires a function that returns floats to populate the list of values. In this example, that function is `get_random_float`.
#
# The Generator is initialized with the random value function, along with the desired size of the float list, and an optional list of indices on which to perform local optimization.
# The Generator is used to generate populations of Chromosomes on Islands.
# +
import numpy as np
from bingo.chromosomes.multiple_floats import MultipleFloatChromosomeGenerator
VALUE_LIST_SIZE = 8
np.random.seed(0)
def get_random_float():
return np.random.random_sample()
generator = MultipleFloatChromosomeGenerator(get_random_float, VALUE_LIST_SIZE, [1, 3, 4])
# -
# Example of Generator
chromosome = generator()
print(chromosome)
print(chromosome.get_number_local_optimization_params())
# ### Chromosome Variation
# Variation of `MultipleValueChromosome` individuals is performed with single-point crossover and/or single-point mutation.
# +
from bingo.chromosomes.multiple_values import SinglePointCrossover
from bingo.chromosomes.multiple_values import SinglePointMutation
crossover = SinglePointCrossover()
mutation = SinglePointMutation(get_random_float)
# -
# Example of Mutation
before_mutation = MultipleFloatChromosome([0., 0., 0., 0., 0., 0.])
after_mutation = mutation(before_mutation)
print("Mutation")
print("before: ", before_mutation)
print("after: ", after_mutation)
# Example of Crossover
parent_1 = MultipleFloatChromosome([0., 0., 0., 0., 0., 0.])
parent_2 = MultipleFloatChromosome([1., 1., 1., 1., 1., 1.])
child_1, child_2 = crossover(parent_1, parent_2)
print("Crossover")
print("parent 1: ", parent_1)
print("parent 1: ", parent_2)
print("child 1: ", child_1)
print("child 1: ", child_2)
# ### Fitness and Evaluation
# In order to Evaluate Chromosomes and assign them a fitness value, first we must define a `FitnessFunction`. For the Zero Min Problem, this Fitness Function calculates fitness by finding the norm of all the values in a Chromosome's list of values. Once a `FitnessFunction` has been defined, it can be passed to an Evaluation to be applied to a population. In this example, we also wrap the `FitnessFunction` with ContinuousLocalOptimization to perform local optimization on indices specified in the Generator class.
# +
from bingo.evaluation.fitness_function import FitnessFunction
from bingo.local_optimizers.continuous_local_opt import ContinuousLocalOptimization
from bingo.evaluation.evaluation import Evaluation
class ZeroMinFitnessFunction(FitnessFunction):
def __call__(self, individual):
return np.linalg.norm(individual.values)
fitness = ZeroMinFitnessFunction()
local_opt_fitness = ContinuousLocalOptimization(fitness)
evaluator = Evaluation(local_opt_fitness) # evaluates a population (list of chromosomes)
# -
# Example of fitness
chromosome = MultipleFloatChromosome([1., 1., 1., 1., 1., 1.],
needs_opt_list=[0, 3]) # perform local optimization on these indices
print(fitness(chromosome))
print(chromosome)
print(local_opt_fitness(chromosome))
print(chromosome)
# Notice that the values in the chromosome at indices 0 and 3 become very near zero. This occurs as part of the local optimization.
# ### Selection
# For this example, we use Tournament Selection to select `GOAL_POPULATION_SIZE` individuals to advance to the next generation.
# +
from bingo.selection.tournament import Tournament
GOAL_POPULATION_SIZE = 25
selection = Tournament(GOAL_POPULATION_SIZE)
# -
# ### Evolutionary Algorithm: Mu + Lambda
# The Evolutionary Algorithm used in this example is called `MuPlusLambda`. Mu represents the parent population and Lambda represents their offspring. MuPlusLambda means the parents and offspring are evaluated together and then the most fit individuals for the next generation are selected from both populations combined. We pass our previously defined Evaluation and Selection modules to MuPlusLambda, along with Crossover and Mutation which will be used to define the behaviors of Variation.
# +
from bingo.evolutionary_algorithms.mu_plus_lambda import MuPlusLambda
MUTATION_PROBABILITY = 0.4
CROSSOVER_PROBABILITY = 0.4
NUM_OFFSPRING = GOAL_POPULATION_SIZE
evo_alg = MuPlusLambda(evaluator,
selection,
crossover,
mutation,
CROSSOVER_PROBABILITY,
MUTATION_PROBABILITY,
NUM_OFFSPRING)
# -
# ### Hall of Fame
# A `HallOfFame` object can be used to keep track of the best individuals that occur during the evolution of a population. It is initialized with the maximum number of members to track, i.e., the 5 best individuals will be saved in the hall of fame in the example below. Optionally, a similarity function can be given as an argument, in order to identify similar individuals (and track only unique ones). It is passed to an `island` on initialization (see next subsection).
# +
from bingo.stats.hall_of_fame import HallOfFame
def similar_mfcs(mfc_1, mfc_2):
"""identifies if two MultpleFloatChromosomes have similar values"""
difference_in_values = 0
for i, j in zip(mfc_1.values, mfc_2.values):
difference_in_values += abs(i - j)
return difference_in_values < 1e-4
hof = HallOfFame(max_size=5, similarity_function=similar_mfcs)
# -
# ### Island
# An `Island` is where evolution takes place in bingo analyses. The `Island` class takes as arguments an Evolutionary Algorithm, a Generator with which to generate an initial population, and thesize of the population on the island. The `Island` will create a population and then execute generational steps of the Evolutionary Algorithm to evolve the population.
# +
from bingo.evolutionary_optimizers.island import Island
POPULATION_SIZE = 10
island = Island(evo_alg, generator, POPULATION_SIZE, hall_of_fame=hof)
# -
print("Island age:", island.generational_age,
" with best fitness:", island.get_best_fitness(), "\n")
for i, indv in enumerate(island.population):
print("indv", i, indv)
# ### Evolution
# There are two mechanisms for performing evolution in bingo.
#
# 1) Manually step through a set number of generations
# +
print("Island age:", island.generational_age,
" with best fitness:", island.get_best_fitness())
island.evolve(num_generations=10)
print("Island age:", island.generational_age,
" with best fitness:", island.get_best_fitness())
# -
# 2) Evolve automatically until convergence
# +
island.evolve_until_convergence(max_generations=1000,
fitness_threshold=0.05)
print("Island age:", island.generational_age,
" with best fitness:", island.get_best_fitness(), "\n")
print("Best indv: ", island.get_best_individual())
# -
# The hall of fame is automatically updated during evolution.
# Note that, for the most part, it can be treated like a list of individuals, in ascending order of fitness.
print("RANK FITNESS")
for i, member in enumerate(hof):
print(" ", i, " ", member.fitness)
# ### Animation of Evolution
# Reinitialize and rerun island while documenting best individual
island = Island(evo_alg, generator, POPULATION_SIZE)
best_indv_values = []
best_indv_values.append(island.get_best_individual().values)
for i in range(50):
island.evolve(1)
best_indv_values.append(island.get_best_individual().values)
# +
import matplotlib.pyplot as plt
import matplotlib.animation as animation
def animate_data(list_of_best_indv_values):
fig, ax = plt.subplots()
num_generations = len(list_of_best_indv_values)
x = np.arange(0, len(list_of_best_indv_values[0]))
y = list_of_best_indv_values
zero = [0]*len(x)
polygon = ax.fill_between(x, zero, y[0], color='b', alpha=0.3)
points, = ax.plot(x, y[0], 'bs')
points.set_label('Generation :' + str(0))
legend = ax.legend(loc='upper right', shadow=True)
def animate(i):
ax.collections.clear()
polygon = ax.fill_between(x, zero, y[i], color='b', alpha=0.3)
points.set_ydata(y[i]) # update the data
points.set_label('Generation :' + str(i))
legend = ax.legend(loc='upper right')
return points, polygon, legend
# Init only required for blitting to give a clean slate.
def init():
points.set_ydata(np.ma.array(x, mask=True))
return points, polygon, points
plt.xlabel('Chromosome Value Index', fontsize=15)
plt.ylabel('Value Magnitude', fontsize=15)
plt.title("Values of Best Individual in Island", fontsize=15)
plt.ylim(-0.01,0.5)
ax.tick_params(axis='y', labelsize=15)
ax.tick_params(axis='x', labelsize=15)
plt.close()
return animation.FuncAnimation(fig, animate, num_generations, init_func=init,
interval=250, blit=True)
# -
from IPython.display import HTML
HTML(animate_data(best_indv_values).to_jshtml())
| examples/Tutorial_2_Zero_Min.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# [](https://github.com/awslabs/aws-data-wrangler)
#
# # 1 - Introduction
# ## What is AWS Data Wrangler?
#
# An [open-source](https://github.com/awslabs/aws-data-wrangler>) Python package that extends the power of [Pandas](https://github.com/pandas-dev/pandas>) library to AWS connecting **DataFrames** and AWS data related services (**Amazon Redshift**, **AWS Glue**, **Amazon Athena**, **Amazon EMR**, etc).
#
# Built on top of other open-source projects like [Pandas](https://github.com/pandas-dev/pandas), [Apache Arrow](https://github.com/apache/arrow), [Boto3](https://github.com/boto/boto3), [SQLAlchemy](https://github.com/sqlalchemy/sqlalchemy), [Psycopg2](https://github.com/psycopg/psycopg2) and [PyMySQL](https://github.com/PyMySQL/PyMySQL), it offers abstracted functions to execute usual ETL tasks like load/unload data from **Data Lakes**, **Data Warehouses** and **Databases**.
#
# Check our [list of functionalities](https://aws-data-wrangler.readthedocs.io/en/latest/api.html).
# ## How to install?
#
# The Wrangler runs almost anywhere over Python 3.6, 3.7 and 3.8, so there are several different ways to install it in the desired enviroment.
#
# - [PyPi (pip)](https://aws-data-wrangler.readthedocs.io/en/latest/install.html#pypi-pip)
# - [Conda](https://aws-data-wrangler.readthedocs.io/en/latest/install.html#conda)
# - [AWS Lambda Layer](https://aws-data-wrangler.readthedocs.io/en/latest/install.html#aws-lambda-layer)
# - [AWS Glue Python Shell Jobs](https://aws-data-wrangler.readthedocs.io/en/latest/install.html#aws-glue-python-shell-jobs)
# - [AWS Glue PySpark Jobs](https://aws-data-wrangler.readthedocs.io/en/latest/install.html#aws-glue-pyspark-jobs)
# - [Amazon SageMaker Notebook](https://aws-data-wrangler.readthedocs.io/en/latest/install.html#amazon-sagemaker-notebook)
# - [Amazon SageMaker Notebook Lifecycle](https://aws-data-wrangler.readthedocs.io/en/latest/install.html#amazon-sagemaker-notebook-lifecycle)
# - [EMR Cluster](https://aws-data-wrangler.readthedocs.io/en/latest/install.html#emr-cluster)
# - [From source](https://aws-data-wrangler.readthedocs.io/en/latest/install.html#from-source)
#
# Some good practices for most of the above methods are:
# - Use new and individual Virtual Environments for each project ([venv](https://docs.python.org/3/library/venv.html))
# - On Notebooks, always restart your kernel after installations.
# ## Let's Install it!
# !pip install awswrangler
# > Restart your kernel after the installation!
# +
import awswrangler as wr
wr.__version__
# -
| tutorials/001 - Introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/niz11/Bachelor_Thesis_Colab_Files/blob/main/10_frame_seq_classifier_composite.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="EqMB5X65d0wf"
import numpy as np
from sklearn.model_selection import train_test_split
from keras.utils import to_categorical
# Load encoding
X = np.load('drive/My Drive/facea_seqs_one_frame_to_the_right/X.npy')
y = np.load('drive/My Drive/facea_seqs_one_frame_to_the_right/Y_label.npy')
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.3, random_state = 0)
y_train = to_categorical(y_train, num_classes=7)
y_test = to_categorical(y_test, num_classes=7)
# + id="mjG_Ek3ZuE9I" outputId="89713773-a426-4545-a440-4a5472153c0d" colab={"base_uri": "https://localhost:8080/"}
import keras
from keras.layers import Input, Flatten, Dense,LSTM,TimeDistributed,RepeatVector
from keras.models import Model,Sequential
from keras.utils import plot_model
def create_model(encoding_length=64,sequence_length=10):
# Define an input sequence and process it.
encoder_inputs = Input(shape=(sequence_length, encoding_length))
encoder = LSTM(encoding_length, return_sequences=True)(encoder_inputs)
encoder_outputs, state_h, state_c = LSTM(encoding_length, return_state=True, name='encoder')(encoder)
encoder_states = [state_h, state_c]
clssifier = Dense(7, activation='softmax')(encoder_outputs)
model = Model(inputs=encoder_inputs, outputs=[clssifier])
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
plot_model(model, show_shapes=True, to_file='lstm_autoencoder.png')
model.summary()
return model
model = create_model(128, 10)
model.load_weights("drive/My Drive/trained_models/composite_paper_3/encoderComposite_3.h5",by_name=True)
# + id="s46Hu_6xuMqP" outputId="8241a6c9-0583-4916-c05d-85462ad4a2c8" colab={"base_uri": "https://localhost:8080/"}
model.fit(X_train, y_train, epochs=10, verbose=1, validation_split=0.05)
# + id="cUclohW3uQZF"
def getFaceExpressionFromIndex(i):
if (i == 0):
return 'surprise'
elif (i == 1):
return 'smile'
elif (i == 2):
return 'sad'
elif (i == 3):
return 'anger'
elif (i == 4):
return 'fear'
elif (i == 5):
return 'disgust'
elif (i == 6):
return 'none'
else:
print(i)
# + id="qEDuKlpwuS-e" outputId="c72733c9-4db8-44c1-9e77-e1dda0cee551" colab={"base_uri": "https://localhost:8080/"}
test_predictions = model.predict(X_test)
correct = 0
for i in range(len(test_predictions)):
truth = np.argmax(y_test[i])
prediction = np.argmax(test_predictions[i])
if truth == prediction:
correct += 1
else:
print(f'Wrong classification, truth: {getFaceExpressionFromIndex(truth)}')
print(f'Wrong classification, prediction: {getFaceExpressionFromIndex(prediction)}')
print("---------------------------------------------------------------------------")
print(f'number of samples: {len(test_predictions)}')
print(f'correct: {correct}')
print(f'Accuracy of predicitons: {correct / len(test_predictions)}')
# + id="qDja8eYquVJY"
# Here and below is the network to classify the encodings
# + id="RoqTKsiCt-4h" outputId="acda1311-393c-4c3e-b2ec-9ae33bee6a51" colab={"base_uri": "https://localhost:8080/", "height": 34}
from keras.models import load_model
from keras.models import model_from_json
import json
# load json and create model
json_file = open('drive/My Drive/trained_models/composite_paper_one_frame_to_the_right/encoderCompositeFaceOneToRight.json', 'r')
loaded_model_json = json_file.read()
json_file.close()
encoder = model_from_json(loaded_model_json)
# load weights into new model
encoder.load_weights("drive/My Drive/trained_models/composite_paper_one_frame_to_the_right/encoderCompositeFaceOneToRight.h5")
print("Loaded model from disk")
# + id="uWKadq4vb4Gf"
def getFaceExpressionFromIndex(i):
if (i == 0):
return 'surprise'
elif (i == 1):
return 'smile'
elif (i == 2):
return 'sad'
elif (i == 3):
return 'anger'
elif (i == 4):
return 'fear'
elif (i == 5):
return 'disgust'
elif (i == 6):
return 'none'
else:
print(i)
# + id="XbX46PHcGRtp"
# Check here: better divinding to test/train set
import numpy as np
from sklearn.model_selection import train_test_split
from keras.utils import to_categorical
# Load encoding
X = np.load('drive/My Drive/facea_seqs_one_frame_to_the_right/X.npy')
y = np.load('drive/My Drive/facea_seqs_one_frame_to_the_right/Y_label.npy')
predictions = encoder.predict(X)
X_train, X_test, y_train, y_test = train_test_split(predictions, y, test_size = 0.3, random_state = 0)
y_train = to_categorical(y_train, num_classes=7)
y_test = to_categorical(y_test, num_classes=7)
# + id="eDOOfsmXXsMy"
from keras.models import Sequential
from keras.layers import Dense
from keras.optimizers import SGD
def classifier():
model = Sequential()
model.add(Dense(64, input_dim=128, activation='relu'))
model.add(Dense(32, activation='relu'))
model.add(Dense(7, activation='softmax'))
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
return model
# + id="kIbFL_mfYOrl" outputId="87f044cf-12e2-4db3-ac2e-7424b85997e5" colab={"base_uri": "https://localhost:8080/", "height": 1000}
model = classifier()
model.fit(X_train, y_train, epochs=100, verbose=1, validation_split=0.1)
# + id="vJxQwGS9YYA5" outputId="e54f8d88-0bcf-4690-bf9d-908471090ba6" colab={"base_uri": "https://localhost:8080/", "height": 1000}
test_predictions = model.predict(X_test)
correct = 0
for i in range(len(test_predictions)):
truth = np.argmax(y_test[i])
prediction = np.argmax(test_predictions[i])
if truth == prediction:
correct += 1
else:
print(f'Wrong classification, truth: {getFaceExpressionFromIndex(truth)}')
print(f'Wrong classification, prediction: {getFaceExpressionFromIndex(prediction)}')
print("---------------------------------------------------------------------------")
print(f'Accuracy of predicitons: {correct / len(test_predictions)}')
print(f'Got correct: {correct}')
print(f'Got wrong: {len(test_predictions) - correct}')
# + id="yyk0RtRMZoSq"
| 10_frame_seq_classifier_composite.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.2
# language: julia
# name: julia-1.6
# ---
# +
using GLM
using CSV
using Random
using StatsBase
using DataFrames
using Dagitty
using Turing
using StatsPlots
using StatisticalRethinking
using StatisticalRethinkingPlots
using Logging
default(labels=false)
Logging.disable_logging(Logging.Warn);
# -
# # 7.1 The problem with parameters
# Code 7.1
sppnames = ["afarensis", "africanus", "habilis", "boisei", "rudolfensis", "ergaster", "sapiens"]
brainvolcc = [438, 452, 612, 521, 752, 871, 1350]
masskg = [37.0, 35.5, 34.5, 41.5, 55.5, 61.0, 53.5]
d = DataFrame(:species => sppnames, :brain => brainvolcc, :mass => masskg);
# Code 7.2
d[!,:mass_std] = (d.mass .- mean(d.mass))./std(d.mass)
d[!,:brain_std] = d.brain ./ maximum(d.brain);
# Code 7.3
# +
Random.seed!(1)
@model function model_m7_1(mass_std, brain_std)
a ~ Normal(0.5, 1)
b ~ Normal(0, 10)
μ = @. a + b*mass_std
log_σ ~ Normal()
brain_std ~ MvNormal(μ, exp(log_σ))
end
m7_1_ch = sample(model_m7_1(d.mass_std, d.brain_std), NUTS(), 1000)
m7_1 = DataFrame(m7_1_ch)
precis(m7_1)
# -
# Code 7.4
X = hcat(ones(length(d.mass_std)), d.mass_std)
m = lm(X, d.brain_std)
# Code 7.5
# +
Random.seed!(12)
# do explicit simulation due to log_σ
s = [
rand(MvNormal((@. r.a + r.b * d.mass_std), exp(r.log_σ)))
for r ∈ eachrow(m7_1)
]
s = vcat(s'...);
r = mean.(eachcol(s)) .- d.brain_std;
resid_var = var(r, corrected=false)
outcome_var = var(d.brain_std, corrected=false)
1 - resid_var/outcome_var
# -
# Code 7.6
# function is implemented in a generic way to support any amount of b[x] coefficients
function R2_is_bad(df; sigma=missing)
degree = ncol(df[!,r"b"])
# build mass_std*degree matrix, with each col exponentiated to col's index
t = repeat(d.mass_std, 1, degree)
t = hcat(map(.^, eachcol(t), 1:degree)...)
s = [
begin
# calculate product on coefficient's vector
b = collect(r[r"b"])
μ = r.a .+ t * b
s = ismissing(sigma) ? exp(r.log_σ) : sigma
rand(MvNormal(μ, s))
end
for r ∈ eachrow(df)
]
s = vcat(s'...);
r = mean.(eachcol(s)) .- d.brain_std;
v1 = var(r, corrected=false)
v2 = var(d.brain_std, corrected=false)
1 - v1 / v2
end
# Code 7.7
# +
Random.seed!(1)
@model function model_m7_2(mass_std, brain_std)
a ~ Normal(0.5, 1)
b ~ MvNormal([0, 0], 10)
μ = @. a + b[1]*mass_std + b[2]*mass_std^2
log_σ ~ Normal()
brain_std ~ MvNormal(μ, exp(log_σ))
end
m7_2_ch = sample(model_m7_2(d.mass_std, d.brain_std), NUTS(), 10000)
m7_2 = DataFrame(m7_2_ch);
# -
# Code 7.8
#
# Implemented the sample in a general way
# +
Random.seed!(3)
@model function model_m7_n(mass_std, brain_std; degree::Int)
a ~ Normal(0.5, 1)
b ~ MvNormal(zeros(degree), 10)
# build matrix n*degree
t = repeat(mass_std, 1, degree)
# exponent its columns
t = hcat(map(.^, eachcol(t), 1:degree)...)
# calculate product on coefficient's vector
μ = a .+ t * b
log_σ ~ Normal()
brain_std ~ MvNormal(μ, exp(log_σ))
end
m7_3_ch = sample(model_m7_n(d.mass_std, d.brain_std, degree=3), NUTS(), 1000)
m7_3 = DataFrame(m7_3_ch);
m7_4_ch = sample(model_m7_n(d.mass_std, d.brain_std, degree=4), NUTS(), 1000)
m7_4 = DataFrame(m7_4_ch);
m7_5_ch = sample(model_m7_n(d.mass_std, d.brain_std, degree=5), NUTS(), 1000)
m7_5 = DataFrame(m7_5_ch);
# -
# Code 7.9
# +
Random.seed!(1)
@model function model_m7_6(mass_std, brain_std)
a ~ Normal(0.5, 1)
b ~ MvNormal(zeros(6), 10)
μ = @. a + b[1]*mass_std + b[2]*mass_std^2 + b[3]*mass_std^3 +
b[4]*mass_std^4 + b[5]*mass_std^5 + b[6]*mass_std^6
brain_std ~ MvNormal(μ, 0.001)
end
m7_6_ch = sample(model_m7_6(d.mass_std, d.brain_std), NUTS(), 1000)
m7_6 = DataFrame(m7_6_ch);
# -
# Code 7.10
# +
mass_seq = range(extrema(d.mass_std)...; length=100)
l = [
@. r.a + r.b * mass_seq
for r ∈ eachrow(m7_1)
]
l = vcat(l'...)
μ = mean.(eachcol(l))
ci = PI.(eachcol(l))
ci = vcat(ci'...)
scatter(d.mass_std, d.brain_std; title="1: R² = $(round(R2_is_bad(m7_1); digits=3))")
plot!(mass_seq, [μ μ]; fillrange=ci, c=:black, fillalpha=0.3)
# -
# reimplemented the brand_plot function to check my results
function brain_plot(df; sigma=missing)
degree = ncol(df[!,r"b"])
# build mass_seq*degree matrix, with each col exponentiated to col's index
t = repeat(mass_seq, 1, degree)
t = hcat(map(.^, eachcol(t), 1:degree)...)
l = [
r.a .+ t * collect(r[r"b"])
for r ∈ eachrow(df)
]
l = vcat(l'...)
μ = mean.(eachcol(l))
ci = PI.(eachcol(l))
ci = vcat(ci'...)
r2 = round(R2_is_bad(df, sigma=sigma); digits=3)
scatter(d.mass_std, d.brain_std; title="$degree: R² = $r2")
plot!(mass_seq, [μ μ]; fillrange=ci, c=:black, fillalpha=0.3)
end
plot(
brain_plot(m7_1),
brain_plot(m7_2),
brain_plot(m7_3),
brain_plot(m7_4),
brain_plot(m7_5),
brain_plot(m7_6, sigma=0.001);
size=(1000, 600)
)
# Code 7.11
i = 3
d_minus_i = d[setdiff(1:end,i),:];
function brain_loo_plot(model, data; title::String)
(a, b) = extrema(data.brain_std)
p = scatter(data.mass_std, data.brain_std; title=title, ylim=(a-0.1, b+0.1))
mass_seq = range(extrema(data.mass_std)...; length=100)
for i ∈ 1:nrow(data)
d_minus_i = data[setdiff(1:end,i),:]
df = DataFrame(sample(model(d_minus_i.mass_std, d_minus_i.brain_std), NUTS(), 1000))
degree = ncol(df[!,r"b"])
# build mass_seq*degree matrix, with each col exponentiated to col's index
t = repeat(mass_seq, 1, degree)
t = hcat(map(.^, eachcol(t), 1:degree)...)
l = [
r.a .+ t * collect(r[r"b"])
for r ∈ eachrow(df)
]
l = vcat(l'...)
μ = mean.(eachcol(l))
plot!(mass_seq, μ; c=:black)
end
p
end
# +
Random.seed!(1)
model_m7_4 = (mass, brain) -> model_m7_n(mass, brain, degree=4)
plot(
brain_loo_plot(model_m7_1, d, title="m7.1"),
brain_loo_plot(model_m7_4, d, title="m7.4");
size=(800, 400)
)
# -
# # 7.2 Entropy and accuracy
# Code 7.12
p = [0.3, 0.7]
-sum(p .* log.(p))
| 07-Chapter 7. Ulysses' Compass.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import gym
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
import json
import numpy as np
import scipy as sp
import scipy.stats as st
import scipy.integrate as integrate
from scipy.stats import multivariate_normal
from sklearn import linear_model
from sklearn.exceptions import ConvergenceWarning
import statsmodels.api as sm
from matplotlib.colors import LogNorm
sns.set_style("whitegrid")
sns.set_palette("colorblind")
palette = sns.color_palette()
figsize = (15,8)
legend_fontsize = 16
from matplotlib import rc
rc('font',**{'family':'sans-serif'})
rc('text.latex',preamble=r'\usepackage[utf8]{inputenc}')
rc('text.latex',preamble=r'\usepackage[russian]{babel}')
rc('figure', **{'dpi': 300})
# -
# ## OpenAI Gym
from gym import envs
print("\n".join(["%s" % x for x in envs.registry.all()]))
env = gym.make('FrozenLake-v1')
env.reset()
for _ in range(5):
env.render()
env.step(env.action_space.sample()) # take a random action
env.close()
env.env.P
# ## Policy iteration по уравнениям Беллмана
# +
nS, nA = env.env.nS, env.env.nA
final_states = np.where([ len(env.env.P[x][0]) == 1 and env.env.P[x][0][0][3] == True for x in env.env.P.keys() ])[0]
def get_random_V(env):
V = np.random.random(nS)
V[final_states] = 0.0
return V
def get_random_Q(env):
Q = np.random.random(size=(nS, nA))
Q[final_states, :] = 0.0
return Q
# +
def compute_V_by_policy(env, pi, gamma=1.0):
V = get_random_V(env)
while True:
new_V = np.array([ \
np.sum([ x[0] * ( x[2] + gamma * V[x[1]] ) for x in env.env.P[cur_state][pi[cur_state]] ]) \
for cur_state in range(nS) ])
if np.sum((V - new_V) ** 2) < 0.001:
break
V = new_V
return V
def compute_policy_by_V(env, V, gamma=1.0):
return np.argmax( np.array([[ \
np.sum([ x[0] * ( x[2] + gamma * V[x[1]] ) for x in env.env.P[s][a] ]) \
for a in range(nA) ] for s in range(nS)]), axis=1 )
# -
def compute_V_and_pi(env, gamma=1.0):
V = get_random_V(env)
pi = np.random.randint(nA, size=nS)
while True:
V = compute_V_by_policy(env, pi, gamma)
new_pi = compute_policy_by_V(env, V, gamma)
if np.array_equal(pi, new_pi):
break
pi = new_pi
return V, pi
# +
env = gym.make('FrozenLake-v1')
env._max_episode_steps = 10000
num_experiments = 200
num_steps, total_reward = [], []
V, pi = compute_V_and_pi(env)
for _ in range(num_experiments):
env.reset()
total_reward.append(0)
for step in range(1000):
observation, reward, done, info = env.step(pi[env.env.s])
total_reward[-1] += reward
if done:
num_steps.append(step+1)
print("Эпизод закончился за %d шагов в состоянии %s, общая награда %d" % (num_steps[-1], env.env.s, total_reward[-1]) )
break
env.close()
print("\nСредняя награда: %.6f\nСреднее число шагов: %.6f" % (np.mean(total_reward), np.mean(num_steps)))
# +
def conduct_experiments_pi(env, pi, num_experiments=1000):
num_steps, total_reward = [], []
for _ in range(num_experiments):
env.reset()
num_steps.append(0)
total_reward.append(0)
for _ in range(1000):
observation, reward, done, info = env.step(pi[env.env.s])
total_reward[-1] += reward
num_steps[-1] += 1
if done:
break
env.close()
return np.mean(total_reward), np.mean(num_steps)
def conduct_experiments(env, gamma=1.0, num_experiments=100, num_experiments_pi=10):
num_steps, total_reward = [], []
for _ in range(num_experiments):
V, pi = compute_V_and_pi(env, gamma=gamma)
cur_steps, cur_reward = conduct_experiments_pi(env, pi, num_experiments=num_experiments_pi)
num_steps.append(cur_steps)
total_reward.append(cur_reward)
return np.mean(num_steps), np.mean(total_reward)
# -
env = gym.make('FrozenLake-v1')
env._max_episode_steps = 10000
results = []
for gamma in np.linspace(0.5, 1.0, 10):
mean_reward, mean_steps = conduct_experiments(env, gamma, num_experiments=100, num_experiments_pi=10)
results.append([gamma, mean_reward, mean_steps])
print("gamma=%.4f, mean reward = %.4f, mean steps = %.4f" % (gamma, mean_reward, mean_steps) )
env.close()
# +
def plot_results(results):
gammas, rewards, numsteps = [x[0] for x in results], [x[1] for x in results], [x[2] for x in results]
fig, ax = plt.subplots(1, 1, figsize=(12, 6))
ax2 = ax.twinx()
ax2.grid(None)
line1 = ax.plot(gammas, rewards, label="Средние награды", color="C0")
line2 = ax2.plot(gammas, numsteps, label="Среднее число шагов", color="C1")
lines = line1 + line2
labels = [l.get_label() for l in lines]
ax.legend(lines, labels, loc=2)
ax.set_xlim((0.5, 1.0))
# ax.set_ylim((0.1, 0.8))
# ax2.set_ylim((10, 45))
return fig, ax
fig, ax = plot_results(results)
# -
# ## Value iteration по уравнениям Беллмана
# +
def compute_V_max(env, gamma=1.0):
V = get_random_V(env)
while True:
new_V = np.array([ [ \
np.sum([ x[0] * ( x[2] + gamma * V[x[1]] ) for x in env.env.P[cur_state][cur_action] ]) \
for cur_action in range(nA) ] for cur_state in range(nS) ])
new_V = np.max(new_V, axis=1)
if np.sum((V - new_V) ** 2) < 0.001:
break
V = new_V
return V
def compute_Q_max(env, gamma=1.0):
Q = get_random_Q(env)
while True:
new_Q = np.array([ [ \
np.sum([ x[0] * ( x[2] + gamma * np.max(Q[x[1], :]) ) for x in env.env.P[cur_state][cur_action] ]) \
for cur_action in range(nA) ] for cur_state in range(nS) ])
# new_V = np.max(new_V, axis=1)
if np.sum((Q - new_Q) ** 2) < 0.001:
break
Q = new_Q
return Q
def compute_policy_by_Q(env, Q, gamma=1.0):
return np.argmax( Q, axis=1 )
def conduct_experiments_max(env, gamma, use_Q=False, num_experiments=100, num_experiments_pi=200):
num_steps, total_reward = [], []
for _ in range(num_experiments):
if use_Q:
Q = compute_Q_max(env, gamma=gamma)
pi = compute_policy_by_Q(env, Q)
else:
V = compute_V_max(env, gamma=gamma)
pi = compute_policy_by_V(env, V)
result = conduct_experiments_pi(env, pi, num_experiments=num_experiments_pi)
num_steps.append(result[0])
total_reward.append(result[1])
return np.mean(num_steps), np.mean(total_reward)
# +
env = gym.make('FrozenLake-v1')
env._max_episode_steps = 10000
V = compute_V_max(env)
pi = compute_policy_by_V(env, V, gamma=0.2)
print(pi)
env.close()
# +
env = gym.make('FrozenLake-v1')
env._max_episode_steps = 10000
Q = compute_Q_max(env)
pi = compute_policy_by_Q(env, Q, gamma=0.2)
print(pi)
env.close()
# +
env = gym.make('FrozenLake-v1')
env._max_episode_steps = 10000
results_max = []
for gamma in np.linspace(0.5, 1.0, 20):
mean_reward, mean_steps = conduct_experiments_max(env, gamma, use_Q=True, num_experiments=20, num_experiments_pi=100)
results_max.append([gamma, mean_reward, mean_steps])
print("gamma=%.4f, mean reward = %.4f, mean steps = %.4f" % (gamma, mean_reward, mean_steps) )
env.close()
# -
fig, ax = plot_results(results_max)
| seminars/02-rlintro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="copyright"
# Copyright 2021 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="title:migration,new"
# # AI Platform (Unified) SDK: Train and deploy an XGBoost model with pre-built containers (formerly hosted runtimes)
#
#
# + [markdown] id="install_aip"
# ## Installation
#
# Install the latest (preview) version of AI Platform (Unified) SDK.
#
#
# + id="install_aip"
# ! pip3 install -U google-cloud-aiplatform --user
# + [markdown] id="install_storage"
# Install the Google *cloud-storage* library as well.
#
#
# + id="install_storage"
# ! pip3 install google-cloud-storage
# + [markdown] id="restart"
# ### Restart the Kernel
#
# Once you've installed the AI Platform (Unified) SDK and Google *cloud-storage*, you need to restart the notebook kernel so it can find the packages.
#
#
# + id="restart"
import os
if not os.getenv("AUTORUN"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
# + [markdown] id="before_you_begin"
# ## Before you begin
#
# ### GPU run-time
#
# *Make sure you're running this notebook in a GPU runtime if you have that option. In Colab, select* **Runtime > Change Runtime Type > GPU**
#
# ### Set up your GCP project
#
# **The following steps are required, regardless of your notebook environment.**
#
# 1. [Select or create a GCP project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
#
# 2. [Make sure that billing is enabled for your project.](https://cloud.google.com/billing/docs/how-to/modify-project)
#
# 3. [Enable the AI Platform APIs and Compute Engine APIs.](https://console.cloud.google.com/flows/enableapi?apiid=ml.googleapis.com,compute_component)
#
# 4. [Google Cloud SDK](https://cloud.google.com/sdk) is already installed in AI Platform Notebooks.
#
# 5. Enter your project ID in the cell below. Then run the cell to make sure the
# Cloud SDK uses the right project for all the commands in this notebook.
#
# **Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
#
#
# + id="set_project_id"
PROJECT_ID = "[your-project-id]" #@param {type:"string"}
# + id="autoset_project_id"
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
# shell_output = !gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
# + id="set_gcloud_project_id"
# ! gcloud config set project $PROJECT_ID
# + [markdown] id="region"
# #### Region
#
# You can also change the `REGION` variable, which is used for operations
# throughout the rest of this notebook. Below are regions supported for AI Platform (Unified). We recommend when possible, to choose the region closest to you.
#
# - Americas: `us-central1`
# - Europe: `europe-west4`
# - Asia Pacific: `asia-east1`
#
# You cannot use a Multi-Regional Storage bucket for training with AI Platform. Not all regions provide support for all AI Platform services. For the latest support per region, see [Region support for AI Platform (Unified) services](https://cloud.google.com/ai-platform-unified/docs/general/locations)
#
#
# + id="region"
REGION = 'us-central1' #@param {type: "string"}
# + [markdown] id="timestamp"
# #### Timestamp
#
# If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append onto the name of resources which will be created in this tutorial.
#
#
# + id="timestamp"
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
# + [markdown] id="gcp_authenticate"
# ### Authenticate your GCP account
#
# **If you are using AI Platform Notebooks**, your environment is already
# authenticated. Skip this step.
#
# *Note: If you are on an AI Platform notebook and run the cell, the cell knows to skip executing the authentication steps.*
#
#
# + id="gcp_authenticate"
import os
import sys
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your Google Cloud account. This provides access
# to your Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# If on AI Platform, then don't execute this code
if not os.path.exists('/opt/deeplearning/metadata/env_version'):
if 'google.colab' in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this tutorial in a notebook locally, replace the string
# below with the path to your service account key and run this cell to
# authenticate your Google Cloud account.
else:
# %env GOOGLE_APPLICATION_CREDENTIALS your_path_to_credentials.json
# Log in to your account on Google Cloud
# ! gcloud auth login
# + [markdown] id="bucket:batch_prediction"
# ### Create a Cloud Storage bucket
#
# **The following steps are required, regardless of your notebook environment.**
#
# This tutorial is designed to use training data that is in a public Cloud Storage bucket and a local Cloud Storage bucket for your batch predictions. You may alternatively use your own training data that you have stored in a local Cloud Storage bucket.
#
# Set the name of your Cloud Storage bucket below. It must be unique across all Cloud Storage buckets.
#
#
# + id="bucket"
BUCKET_NAME = "[your-bucket-name]" #@param {type:"string"}
# + id="autoset_bucket"
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "[your-bucket-name]":
BUCKET_NAME = PROJECT_ID + "aip-" + TIMESTAMP
# + [markdown] id="create_bucket"
# **Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
#
#
# + id="create_bucket"
# ! gsutil mb -l $REGION gs://$BUCKET_NAME
# + [markdown] id="validate_bucket"
# Finally, validate access to your Cloud Storage bucket by examining its contents:
#
#
# + id="validate_bucket"
# ! gsutil ls -al gs://$BUCKET_NAME
# + [markdown] id="setup_vars"
# ### Set up variables
#
# Next, set up some variables used throughout the tutorial.
# ### Import libraries and define constants
#
#
# + [markdown] id="import_aip"
# #### Import AI Platform (Unified) SDK
#
# Import the AI Platform (Unified) SDK into our Python environment.
#
#
# + id="import_aip"
import os
import sys
import time
from google.cloud.aiplatform import gapic as aip
from google.protobuf.struct_pb2 import Value
from google.protobuf.struct_pb2 import Struct
from google.protobuf.json_format import MessageToJson
from google.protobuf.json_format import ParseDict
# + [markdown] id="aip_constants"
# #### AI Platform (Unified) constants
#
# Setup up the following constants for AI Platform (Unified):
#
# - `API_ENDPOINT`: The AI Platform (Unified) API service endpoint for dataset, model, job, pipeline and endpoint services.
# - `PARENT`: The AI Platform (Unified) location root path for dataset, model and endpoint resources.
#
#
# + id="aip_constants"
# API Endpoint
API_ENDPOINT = "{0}-aiplatform.googleapis.com".format(REGION)
# AI Platform (Unified) location root path for your dataset, model and endpoint resources
PARENT = "projects/" + PROJECT_ID + "/locations/" + REGION
# + [markdown] id="clients"
# ## Clients
#
# The AI Platform (Unified) SDK works as a client/server model. On your side (the Python script) you will create a client that sends requests and receives responses from the server (AI Platform).
#
# You will use several clients in this tutorial, so set them all up upfront.
#
# - Model Service for managed models.
# - Endpoint Service for deployment.
# - Job Service for batch jobs and custom training.
# - Prediction Service for serving. *Note*: Prediction has a different service endpoint.
#
#
# + id="clients"
# client options same for all services
client_options = {"api_endpoint": API_ENDPOINT}
def create_model_client():
client = aip.ModelServiceClient(
client_options=client_options
)
return client
def create_endpoint_client():
client = aip.EndpointServiceClient(
client_options=client_options
)
return client
def create_prediction_client():
client = aip.PredictionServiceClient(
client_options=client_options
)
return client
def create_job_client():
client = aip.JobServiceClient(
client_options=client_options
)
return client
clients = {}
clients['model'] = create_model_client()
clients['endpoint'] = create_endpoint_client()
clients['prediction'] = create_prediction_client()
clients['job'] = create_job_client()
for client in clients.items():
print(client)
# -
# ## Prepare a trainer script
# ### Package assembly
# +
# Make folder for python training script
# ! rm -rf custom
# ! mkdir custom
# Add package information
# ! touch custom/README.md
setup_cfg = "[egg_info]\n\
tag_build =\n\
tag_date = 0"
# ! echo "$setup_cfg" > custom/setup.cfg
setup_py = "import setuptools\n\
setuptools.setup(\n\
install_requires=[\n\
],\n\
packages=setuptools.find_packages())"
# ! echo "$setup_py" > custom/setup.py
pkg_info = "Metadata-Version: 1.0\n\
Name: Custom XGBoost Iris\n\
Version: 0.0.0\n\
Summary: Demonstration training script\n\
Home-page: www.google.com\n\
Author: Google\n\
Author-email: <EMAIL>\n\
License: Public\n\
Description: Demo\n\
Platform: AI Platform (Unified)"
# ! echo "$pkg_info" > custom/PKG-INFO
# Make the training subfolder
# ! mkdir custom/trainer
# ! touch custom/trainer/__init__.py
# -
# ### Task.py contents
# +
# %%writefile custom/trainer/task.py
# Single Instance Training for Iris
import datetime
import os
import subprocess
import sys
import pandas as pd
import xgboost as xgb
import argparse
parser = argparse.ArgumentParser()
parser.add_argument('--model-dir', dest='model_dir',
default=os.getenv('AIP_MODEL_DIR'), type=str, help='Model dir.')
args = parser.parse_args()
# Download data
iris_data_filename = 'iris_data.csv'
iris_target_filename = 'iris_target.csv'
data_dir = 'gs://cloud-samples-data/ai-platform/iris'
# gsutil outputs everything to stderr so we need to divert it to stdout.
subprocess.check_call(['gsutil', 'cp', os.path.join(data_dir,
iris_data_filename),
iris_data_filename], stderr=sys.stdout)
subprocess.check_call(['gsutil', 'cp', os.path.join(data_dir,
iris_target_filename),
iris_target_filename], stderr=sys.stdout)
# Load data into pandas, then use `.values` to get NumPy arrays
iris_data = pd.read_csv(iris_data_filename).values
iris_target = pd.read_csv(iris_target_filename).values
# Convert one-column 2D array into 1D array for use with XGBoost
iris_target = iris_target.reshape((iris_target.size,))
# Load data into DMatrix object
dtrain = xgb.DMatrix(iris_data, label=iris_target)
# Train XGBoost model
bst = xgb.train({}, dtrain, 20)
# Export the classifier to a file
model_filename = 'model.bst'
bst.save_model(model_filename)
# Upload the saved model file to Cloud Storage
gcs_model_path = os.path.join(args.model_dir, model_filename)
subprocess.check_call(['gsutil', 'cp', model_filename, gcs_model_path],
stderr=sys.stdout)
# -
# ### Store training script on your Cloud Storage bucket
# ! rm -f custom.tar custom.tar.gz
# ! tar cvf custom.tar custom
# ! gzip custom.tar
# ! gsutil cp custom.tar.gz gs://$BUCKET_NAME/iris.tar.gz
# + [markdown] id="text_create_and_deploy_model:migration"
# ## Train a model
# + [markdown] id="0oqIBOSnJjkW"
# ### [projects.locations.customJobs.create](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.trainingPipelines/create)
# -
# #### Request
# +
TRAIN_IMAGE = 'gcr.io/cloud-aiplatform/training/xgboost-cpu.1-1:latest'
JOB_NAME = "custom_job_XGB" + TIMESTAMP
WORKER_POOL_SPEC = [
{
"replica_count": 1,
"machine_spec": {
"machine_type": 'n1-standard-4'
},
"python_package_spec": {
"executor_image_uri": TRAIN_IMAGE,
"package_uris": ["gs://" + BUCKET_NAME + "/iris.tar.gz"],
"python_module": "trainer.task",
"args": [
"--model-dir=" + 'gs://{}/{}'.format(BUCKET_NAME, JOB_NAME)
]
}
}
]
training_job = aip.CustomJob(
display_name = JOB_NAME,
job_spec = {
"worker_pool_specs": WORKER_POOL_SPEC
}
)
print(MessageToJson(
aip.CreateCustomJobRequest(
parent=PARENT,
custom_job=training_job
).__dict__["_pb"])
)
# + [markdown] id="datasets_import:migration,new,request"
# *Example output*:
# ```
# {
# "parent": "projects/migration-ucaip-training/locations/us-central1",
# "customJob": {
# "displayName": "custom_job_XGB20210323142337",
# "jobSpec": {
# "workerPoolSpecs": [
# {
# "machineSpec": {
# "machineType": "n1-standard-4"
# },
# "replicaCount": "1",
# "pythonPackageSpec": {
# "executorImageUri": "gcr.io/cloud-aiplatform/training/xgboost-cpu.1-1:latest",
# "packageUris": [
# "gs://migration-ucaip-trainingaip-20210323142337/iris.tar.gz"
# ],
# "pythonModule": "trainer.task",
# "args": [
# "--model-dir=gs://migration-ucaip-trainingaip-20210323142337/custom_job_XGB20210323142337"
# ]
# }
# }
# ]
# }
# }
# }
# ```
#
#
# -
# #### Call
request = clients["job"].create_custom_job(
parent=PARENT,
custom_job=training_job
)
# #### Response
print(MessageToJson(request.__dict__["_pb"]))
# + [markdown] id="datasets_import:migration,new,request"
# *Example output*:
# ```
# {
# "name": "projects/116273516712/locations/us-central1/customJobs/7371064379959148544",
# "displayName": "custom_job_XGB20210323142337",
# "jobSpec": {
# "workerPoolSpecs": [
# {
# "machineSpec": {
# "machineType": "n1-standard-4"
# },
# "replicaCount": "1",
# "diskSpec": {
# "bootDiskType": "pd-ssd",
# "bootDiskSizeGb": 100
# },
# "pythonPackageSpec": {
# "executorImageUri": "gcr.io/cloud-aiplatform/training/xgboost-cpu.1-1:latest",
# "packageUris": [
# "gs://migration-ucaip-trainingaip-20210323142337/iris.tar.gz"
# ],
# "pythonModule": "trainer.task",
# "args": [
# "--model-dir=gs://migration-ucaip-trainingaip-20210323142337/custom_job_XGB20210323142337"
# ]
# }
# }
# ]
# },
# "state": "JOB_STATE_PENDING",
# "createTime": "2021-03-23T14:23:45.067026Z",
# "updateTime": "2021-03-23T14:23:45.067026Z"
# }
# ```
#
#
# + id="training_pipeline_id:migration,new,response"
# The full unique ID for the custom training job
custom_training_id = request.name
# The short numeric ID for the custom training job
custom_training_short_id = custom_training_id.split('/')[-1]
print(custom_training_id)
# + [markdown] id="0oqIBOSnJjkW"
# ### [projects.locations.customJobs.get](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.trainingPipelines/get)
# -
# #### Call
request = clients['job'].get_custom_job(
name=custom_training_id
)
# #### Response
print(MessageToJson(request.__dict__["_pb"]))
# + [markdown] id="datasets_import:migration,new,request"
# *Example output*:
# ```
# {
# "name": "projects/116273516712/locations/us-central1/customJobs/7371064379959148544",
# "displayName": "custom_job_XGB20210323142337",
# "jobSpec": {
# "workerPoolSpecs": [
# {
# "machineSpec": {
# "machineType": "n1-standard-4"
# },
# "replicaCount": "1",
# "diskSpec": {
# "bootDiskType": "pd-ssd",
# "bootDiskSizeGb": 100
# },
# "pythonPackageSpec": {
# "executorImageUri": "gcr.io/cloud-aiplatform/training/xgboost-cpu.1-1:latest",
# "packageUris": [
# "gs://migration-ucaip-trainingaip-20210323142337/iris.tar.gz"
# ],
# "pythonModule": "trainer.task",
# "args": [
# "--model-dir=gs://migration-ucaip-trainingaip-20210323142337/custom_job_XGB20210323142337"
# ]
# }
# }
# ]
# },
# "state": "JOB_STATE_PENDING",
# "createTime": "2021-03-23T14:23:45.067026Z",
# "updateTime": "2021-03-23T14:23:45.067026Z"
# }
# ```
#
#
# + id="trainingpipelines_get:migration,new,wait"
while True:
response = clients["job"].get_custom_job(name=custom_training_id)
if response.state != aip.PipelineState.PIPELINE_STATE_SUCCEEDED:
print("Training job has not completed:", response.state)
if response.state == aip.PipelineState.PIPELINE_STATE_FAILED:
break
else:
print("Training Time:", response.end_time - response.start_time)
break
time.sleep(60)
# model artifact output directory on Google Cloud Storage
model_artifact_dir = response.job_spec.worker_pool_specs[0].python_package_spec.args[0].split("=")[-1]
print("artifact location " + model_artifact_dir)
# -
# ## Deploy the model
# + [markdown] id="COwVZtxhJjkW"
# ### [projects.locations.models.upload](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.models/upload)
# -
# #### Request
# +
DEPLOY_IMAGE = 'gcr.io/cloud-aiplatform/prediction/xgboost-cpu.1-1:latest'
model = {
"display_name": "custom_job_XGB" + TIMESTAMP,
"artifact_uri": model_artifact_dir,
"container_spec": {
"image_uri": DEPLOY_IMAGE,
"ports": [{"container_port": 8080}]
}
}
print(MessageToJson(
aip.UploadModelRequest(
parent=PARENT,
model=model
).__dict__["_pb"])
)
# + [markdown] id="datasets_import:migration,new,request"
# *Example output*:
# ```
# {
# "parent": "projects/migration-ucaip-training/locations/us-central1",
# "model": {
# "displayName": "custom_job_XGB20210323142337",
# "containerSpec": {
# "imageUri": "gcr.io/cloud-aiplatform/prediction/xgboost-cpu.1-1:latest",
# "ports": [
# {
# "containerPort": 8080
# }
# ]
# },
# "artifactUri": "gs://migration-ucaip-trainingaip-20210323142337/custom_job_XGB20210323142337"
# }
# }
# ```
#
#
# -
# #### Call
request = clients['model'].upload_model(
parent=PARENT,
model=model
)
# #### Response
# +
result = request.result()
print(MessageToJson(result.__dict__["_pb"]))
# + [markdown] id="datasets_import:migration,new,request"
# *Example output*:
# ```
# {
# "model": "projects/116273516712/locations/us-central1/models/2093698837704081408"
# }
# ```
#
#
# -
# The full unique ID for the model version
model_id = result.model
# + [markdown] id="make_batch_predictions:migration"
# ## Make batch predictions
#
#
# + [markdown] id="make_batch_prediction_file:migration,new"
# ### Make a batch prediction file
#
#
# + id="get_test_items:automl,icn,csv"
import json
import tensorflow as tf
INSTANCES = [
[1.4, 1.3, 5.1, 2.8],
[1.5, 1.2, 4.7, 2.4]
]
gcs_input_uri = "gs://" + BUCKET_NAME + "/" + "test.jsonl"
with tf.io.gfile.GFile(gcs_input_uri, 'w') as f:
for i in INSTANCES:
f.write(str(i) + '\n')
# ! gsutil cat $gcs_input_uri
# + [markdown] id="datasets_import:migration,new,request"
# *Example output*:
# ```
# [1.4, 1.3, 5.1, 2.8]
# [1.5, 1.2, 4.7, 2.4]
# ```
#
#
# + [markdown] id="batchpredictionjobs_create:migration,new"
# ### [projects.locations.batchPredictionJobs.create](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.batchPredictionJobs/create)
#
#
# + [markdown] id="request:migration"
# #### Request
#
#
# + id="batchpredictionjobs_create:migration,new,request,icn"
model_parameters = Value(struct_value=Struct(
fields={
"confidence_threshold": Value(number_value=0.5),
"max_predictions": Value(number_value=10000.0)
}
))
batch_prediction_job = {
"display_name": "custom_job_XGB" + TIMESTAMP,
"model": model_id,
"input_config": {
"instances_format": "jsonl",
"gcs_source": {
"uris": [gcs_input_uri]
}
},
"model_parameters": model_parameters,
"output_config": {
"predictions_format": "jsonl",
"gcs_destination": {
"output_uri_prefix": "gs://" + f"{BUCKET_NAME}/batch_output/"
}
},
"dedicated_resources": {
"machine_spec": {
"machine_type": "n1-standard-2"
},
"starting_replica_count": 1,
"max_replica_count": 1
}
}
print(MessageToJson(
aip.CreateBatchPredictionJobRequest(
parent=PARENT,
batch_prediction_job=batch_prediction_job
).__dict__["_pb"])
)
# + [markdown] id="batchpredictionjobs_create:migration,new,request,icn"
# *Example output*:
# ```
# {
# "parent": "projects/migration-ucaip-training/locations/us-central1",
# "batchPredictionJob": {
# "displayName": "custom_job_XGB20210323142337",
# "model": "projects/116273516712/locations/us-central1/models/2093698837704081408",
# "inputConfig": {
# "instancesFormat": "jsonl",
# "gcsSource": {
# "uris": [
# "gs://migration-ucaip-trainingaip-20210323142337/test.jsonl"
# ]
# }
# },
# "modelParameters": {
# "max_predictions": 10000.0,
# "confidence_threshold": 0.5
# },
# "outputConfig": {
# "predictionsFormat": "jsonl",
# "gcsDestination": {
# "outputUriPrefix": "gs://migration-ucaip-trainingaip-20210323142337/batch_output/"
# }
# },
# "dedicatedResources": {
# "machineSpec": {
# "machineType": "n1-standard-2"
# },
# "startingReplicaCount": 1,
# "maxReplicaCount": 1
# }
# }
# }
# ```
#
#
# + [markdown] id="call:migration"
# #### Call
#
#
# + id="batchpredictionjobs_create:migration,new,call"
request = clients["job"].create_batch_prediction_job(
parent=PARENT,
batch_prediction_job=batch_prediction_job
)
# + [markdown] id="response:migration"
# #### Response
#
#
# + id="print:migration,new,request"
print(MessageToJson(request.__dict__["_pb"]))
# + [markdown] id="batchpredictionjobs_create:migration,new,response,icn"
# *Example output*:
# ```
# {
# "name": "projects/116273516712/locations/us-central1/batchPredictionJobs/1415053872761667584",
# "displayName": "custom_job_XGB20210323142337",
# "model": "projects/116273516712/locations/us-central1/models/2093698837704081408",
# "inputConfig": {
# "instancesFormat": "jsonl",
# "gcsSource": {
# "uris": [
# "gs://migration-ucaip-trainingaip-20210323142337/test.jsonl"
# ]
# }
# },
# "modelParameters": {
# "confidence_threshold": 0.5,
# "max_predictions": 10000.0
# },
# "outputConfig": {
# "predictionsFormat": "jsonl",
# "gcsDestination": {
# "outputUriPrefix": "gs://migration-ucaip-trainingaip-20210323142337/batch_output/"
# }
# },
# "dedicatedResources": {
# "machineSpec": {
# "machineType": "n1-standard-2"
# },
# "startingReplicaCount": 1,
# "maxReplicaCount": 1
# },
# "manualBatchTuningParameters": {},
# "state": "JOB_STATE_PENDING",
# "createTime": "2021-03-23T14:25:10.582704Z",
# "updateTime": "2021-03-23T14:25:10.582704Z"
# }
# ```
#
#
# + id="batch_job_id:migration,new,response"
# The fully qualified ID for the batch job
batch_job_id = request.name
# The short numeric ID for the batch job
batch_job_short_id = batch_job_id.split('/')[-1]
print(batch_job_id)
# + [markdown] id="batchpredictionjobs_get:migration,new"
# ### [projects.locations.batchPredictionJobs.get](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.batchPredictionJobs/get)
#
#
# + [markdown] id="call:migration"
# #### Call
#
#
# + id="batchpredictionjobs_get:migration,new,call"
request = clients["job"].get_batch_prediction_job(
name=batch_job_id
)
# + [markdown] id="response:migration"
# #### Response
#
#
# + id="print:migration,new,request"
print(MessageToJson(request.__dict__["_pb"]))
# + [markdown] id="batchpredictionjobs_get:migration,new,response,icn"
# *Example output*:
# ```
# {
# "name": "projects/116273516712/locations/us-central1/batchPredictionJobs/1415053872761667584",
# "displayName": "custom_job_XGB20210323142337",
# "model": "projects/116273516712/locations/us-central1/models/2093698837704081408",
# "inputConfig": {
# "instancesFormat": "jsonl",
# "gcsSource": {
# "uris": [
# "gs://migration-ucaip-trainingaip-20210323142337/test.jsonl"
# ]
# }
# },
# "modelParameters": {
# "max_predictions": 10000.0,
# "confidence_threshold": 0.5
# },
# "outputConfig": {
# "predictionsFormat": "jsonl",
# "gcsDestination": {
# "outputUriPrefix": "gs://migration-ucaip-trainingaip-20210323142337/batch_output/"
# }
# },
# "dedicatedResources": {
# "machineSpec": {
# "machineType": "n1-standard-2"
# },
# "startingReplicaCount": 1,
# "maxReplicaCount": 1
# },
# "manualBatchTuningParameters": {},
# "state": "JOB_STATE_PENDING",
# "createTime": "2021-03-23T14:25:10.582704Z",
# "updateTime": "2021-03-23T14:25:10.582704Z"
# }
# ```
#
#
# + id="batchpredictionjobs_get:migration,new,wait"
def get_latest_predictions(gcs_out_dir):
''' Get the latest prediction subfolder using the timestamp in the subfolder name'''
# folders = !gsutil ls $gcs_out_dir
latest = ""
for folder in folders:
subfolder = folder.split('/')[-2]
if subfolder.startswith('prediction-'):
if subfolder > latest:
latest = folder[:-1]
return latest
while True:
response = clients["job"].get_batch_prediction_job(name=batch_job_id)
if response.state != aip.JobState.JOB_STATE_SUCCEEDED:
print("The job has not completed:", response.state)
if response.state == aip.JobState.JOB_STATE_FAILED:
break
else:
folder = get_latest_predictions(response.output_config.gcs_destination.output_uri_prefix)
# ! gsutil ls $folder/prediction*
# ! gsutil cat -h $folder/prediction*
break
time.sleep(60)
# + [markdown] id="batchpredictionjobs_get:migration,new,wait,icn"
# *Example output*:
# ```
# ==> gs://migration-ucaip-trainingaip-20210323142337/batch_output/prediction-custom_job_XGB20210323142337-2021_03_23T07_25_10_544Z/prediction.errors_stats-00000-of-00001 <==
#
# ==> gs://migration-ucaip-trainingaip-20210323142337/batch_output/prediction-custom_job_XGB20210323142337-2021_03_23T07_25_10_544Z/prediction.results-00000-of-00001 <==
# {"instance": [1.4, 1.3, 5.1, 2.8], "prediction": 2.0451931953430176}
# {"instance": [1.5, 1.2, 4.7, 2.4], "prediction": 1.9618644714355469}
# ```
#
#
# -
# ## Make online predictions
# + [markdown] id="endpoints_create:migration,new"
# ### [projects.locations.endpoints.create](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.endpoints/create)
#
#
# + [markdown] id="request:migration"
# #### Request
#
#
# + id="endpoints_create:migration,new,request"
endpoint = {
"display_name": "custom_job_XGB" + TIMESTAMP
}
print(MessageToJson(
aip.CreateEndpointRequest(
parent=PARENT,
endpoint=endpoint
).__dict__["_pb"])
)
# + [markdown] id="endpoints_create:migration,new,request"
# *Example output*:
# ```
# {
# "parent": "projects/migration-ucaip-training/locations/us-central1",
# "endpoint": {
# "displayName": "custom_job_XGB20210323142337"
# }
# }
# ```
#
#
# + [markdown] id="call:migration"
# #### Call
#
#
# + id="endpoints_create:migration,new,call"
request = clients["endpoint"].create_endpoint(
parent=PARENT,
endpoint=endpoint
)
# + [markdown] id="response:migration"
# #### Response
#
#
# + id="print:migration,new,response"
result = request.result()
print(MessageToJson(result.__dict__["_pb"]))
# + [markdown] id="endpoints_create:migration,new,response"
# *Example output*:
# ```
# {
# "name": "projects/116273516712/locations/us-central1/endpoints/1733903448723685376"
# }
# ```
#
#
# + id="endpoint_id:migration,new,response"
# The full unique ID for the endpoint
endpoint_id = result.name
# The short numeric ID for the endpoint
endpoint_short_id = endpoint_id.split('/')[-1]
print(endpoint_id)
# + [markdown] id="endpoints_deploymodel:migration,new"
# ### [projects.locations.endpoints.deployModel](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.endpoints/deployModel)
#
#
# + [markdown] id="request:migration"
# #### Request
#
#
# + id="endpoints_deploymodel:migration,new,request"
deployed_model = {
"model": model_id,
"display_name": "custom_job_XGB" + TIMESTAMP,
"dedicated_resources": {
"min_replica_count": 1,
"max_replica_count": 1,
"machine_spec": {
"machine_type": 'n1-standard-4',
"accelerator_count": 0
}
}
}
print(MessageToJson(
aip.DeployModelRequest(
endpoint=endpoint_id,
deployed_model=deployed_model,
traffic_split={"0": 100}
).__dict__["_pb"])
)
# + [markdown] id="endpoints_deploymodel:migration,new,request"
# *Example output*:
# ```
# {
# "endpoint": "projects/116273516712/locations/us-central1/endpoints/1733903448723685376",
# "deployedModel": {
# "model": "projects/116273516712/locations/us-central1/models/2093698837704081408",
# "displayName": "custom_job_XGB20210323142337",
# "dedicatedResources": {
# "machineSpec": {
# "machineType": "n1-standard-4"
# },
# "minReplicaCount": 1,
# "maxReplicaCount": 1
# }
# },
# "trafficSplit": {
# "0": 100
# }
# }
# ```
#
#
# + [markdown] id="call:migration"
# #### Call
#
#
# + id="endpoints_deploymodel:migration,new,call"
request = clients["endpoint"].deploy_model(
endpoint=endpoint_id,
deployed_model=deployed_model,
traffic_split={"0": 100}
)
# + [markdown] id="response:migration"
# #### Response
#
#
# + id="print:migration,new,response"
result = request.result()
print(MessageToJson(result.__dict__["_pb"]))
# + [markdown] id="endpoints_deploymodel:migration,new,response"
# *Example output*:
# ```
# {
# "deployedModel": {
# "id": "7407594554280378368"
# }
# }
# ```
#
#
# + id="deployed_model_id:migration,new,response"
# The unique ID for the deployed model
deployed_model_id = result.deployed_model.id
print(deployed_model_id)
# + [markdown] id="endpoints_predict:migration,new"
# ### [projects.locations.endpoints.predict](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.endpoints/predict)
#
#
# -
# ### Prepare file for online prediction
INSTANCES = [
[1.4, 1.3, 5.1, 2.8],
[1.5, 1.2, 4.7, 2.4]
]
# + [markdown] id="request:migration"
# #### Request
#
#
# + id="endpoints_predict:migration,new,request,icn"
prediction_request = {
"endpoint": endpoint_id,
"instances": INSTANCES
}
print(json.dumps(prediction_request, indent=2))
# + [markdown] id="endpoints_deploymodel:migration,new,request"
# *Example output*:
# ```
# {
# "endpoint": "projects/116273516712/locations/us-central1/endpoints/1733903448723685376",
# "instances": [
# [
# 1.4,
# 1.3,
# 5.1,
# 2.8
# ],
# [
# 1.5,
# 1.2,
# 4.7,
# 2.4
# ]
# ]
# }
# ```
#
#
# + [markdown] id="call:migration"
# #### Call
#
#
# + id="endpoints_predict:migration,new,call"
request = clients["prediction"].predict(
endpoint=endpoint_id,
instances=INSTANCES
)
# + [markdown] id="response:migration"
# #### Response
#
#
# + id="print:migration,new,request"
print(MessageToJson(request.__dict__["_pb"]))
# + [markdown] id="endpoints_predict:migration,new,response,icn"
# *Example output*:
# ```
# {
# "predictions": [
# 2.045193195343018,
# 1.961864471435547
# ],
# "deployedModelId": "7407594554280378368"
# }
# ```
#
#
# + [markdown] id="endpoints_undeploymodel:migration,new"
# ### [projects.locations.endpoints.undeployModel](https://cloud.google.com/ai-platform-unified/docs/reference/rest/v1beta1/projects.locations.endpoints/undeployModel)
#
#
# + [markdown] id="call:migration"
# #### Call
#
#
# + id="endpoints_undeploymodel:migration,new,call"
request = clients['endpoint'].undeploy_model(
endpoint=endpoint_id,
deployed_model_id=deployed_model_id,
traffic_split={}
)
# + [markdown] id="response:migration"
# #### Response
#
#
# + id="print:migration,new,response"
result = request.result()
print(MessageToJson(result.__dict__["_pb"]))
# + [markdown] id="endpoints_undeploymodel:migration,new,response"
# *Example output*:
# ```
# {}
# ```
#
#
# + [markdown] id="cleanup:migration,new"
# # Cleaning up
#
# To clean up all GCP resources used in this project, you can [delete the GCP
# project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
#
# Otherwise, you can delete the individual resources you created in this tutorial.
#
#
# + id="cleanup:migration,new"
delete_model = True
delete_endpoint = True
delete_pipeline = True
delete_batchjob = True
delete_bucket = True
# Delete the model using the AI Platform (Unified) fully qualified identifier for the model
try:
if delete_model:
clients['model'].delete_model(name=model_id)
except Exception as e:
print(e)
# Delete the endpoint using the AI Platform (Unified) fully qualified identifier for the endpoint
try:
if delete_endpoint:
clients['endpoint'].delete_endpoint(name=endpoint_id)
except Exception as e:
print(e)
# Delete the custom training using the AI Platform (Unified) fully qualified identifier for the custome training
try:
if custom_training_id:
clients['job'].delete_custom_job(name=custom_training_id)
except Exception as e:
print(e)
# Delete the batch job using the AI Platform (Unified) fully qualified identifier for the batch job
try:
if delete_batchjob:
clients['job'].delete_batch_prediction_job(name=batch_job_id)
except Exception as e:
print(e)
if delete_bucket and 'BUCKET_NAME' in globals():
# ! gsutil rm -r gs://$BUCKET_NAME
# -
| ai-platform-unified/notebooks/unofficial/migration/UJ9 unified Custom Training Prebuilt Container XGBoost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Altair Debugging Guide
#
# In this notebook we show you common debugging techniques that you can use if you run into issues with Altair.
#
# You can jump to the following sections:
#
# * [Installation and Setup](#Installation) when Altair is not installed correctly
# * [Display Issues](#Display-Troubleshooting) when you don't see a chart
# * [Invalid Specifications](#Invalid-Specifications) when you get an error
# * [Properties are Being Ignored](#Properties-are-Being-Ignored) when you don't see any errors or warnings
# * [Asking for Help](#Asking-for-Help) when you get stuck
# * [Reporting Issues](#Reporting-Issues) when you find a bug
#
# In addition to this notebook, you might find the [Frequently Asked Questions](https://altair-viz.github.io/user_guide/faq.html) and [Display Troubleshooting](https://altair-viz.github.io/user_guide/troubleshooting.html) guides helpful.
#
# _This notebook is part of the [data visualization curriculum](https://github.com/uwdata/visualization-curriculum)._
# ## Installation
# These instructions follow [the Altair documentation](https://altair-viz.github.io/getting_started/installation.html) but focus on some specifics for this series of notebooks.
#
# In every notebook, we will import the [Altair](https://github.com/altair-viz/altair) and [Vega Datasets](https://github.com/altair-viz/vega_datasets) packages. If you are running this notebook on [Colab](https://colab.research.google.com), Altair and Vega Datasets should be preinstalled and ready to go. The notebooks in this series are designed for Colab but should also work in Jupyter Lab or the Jupyter Notebook (the notebook requires a bit more setup [described below](#Special-Setup-for-the-Jupyter-Notebook)) but additional packages are required.
#
# If you are running in Jupyter Lab or Jupyter Notebooks, you have to install the necessary packages by running the following command in your terminal.
#
# ```bash
# pip install altair vega_datasets
# ```
#
# Or if you use [Conda](https://conda.io)
#
# ```bash
# conda install -c conda-forge altair vega_datasets
# ```
#
# You can run command line commands from a code cell by prefixing it with `!`. For example, to install Altair and Vega Datasets with [Pip](https://pip.pypa.io/), you can run the following cell.
# !pip install altair vega_datasets
import altair as alt
from vega_datasets import data
# ### Make sure you are Using the Latest Version of Altair
# If you are running into issues with Altair, first make sure that you are running the latest version. To check the version of Altair that you have installed, run the cell below.
alt.__version__
# To check what the latest version of altair is, go to [this page](https://pypi.org/project/altair/) or run the cell below (requires Python 3).
import urllib.request, json
with urllib.request.urlopen("https://pypi.org/pypi/altair/json") as url:
print(json.loads(url.read().decode())['info']['version'])
# If you are not running the latest version, you can update it with `pip`. You can update Altair and Vega Datasets by running this command in your terminal.
#
# ```
# pip install -U altair vega_datasets
# ```
# ### Try Making a Chart
# Now you can create an Altair chart.
# +
iris = data.iris()
alt.Chart(iris).mark_point().encode(
x='petalLength',
y='petalWidth',
color='species'
)
# -
# ### Special Setup for the Jupyter Notebook
# If you are running in Colab or Jupyter Lab, you should be seeing a chart. If you are running in the Jupyter Notebook, you need to install an additional dependency and tell Altair to render charts for the Notebook.
#
# The additional dependency is the `vega` package, which you can install by running this command in your terminal
#
# ```bash
# pip install vega
# ```
#
# Then activate the Notebook renderer in a notebook cell
#
# ```python
# # for the notebook only (not for JupyterLab) run this command once per session
# alt.renderers.enable('notebook')
#
# ```
#
# These instruction follow [the instructions on the Altair website](https://altair-viz.github.io/getting_started/installation.html#installation-notebook).
# ## Display Troubleshooting
#
# If you are having issues with seeing a chart, make sure your setup is correct by following the [debugging instruction above](#installation). If you are still having issues, follow the [instruction about debugging display issues in the Altair documentation](https://altair-viz.github.io/user_guide/troubleshooting.html).
# ### Non Existent Fields
#
# A common error is [accidentally using a field that does not exit](https://altair-viz.github.io/user_guide/troubleshooting.html#plot-displays-but-the-content-is-empty).
# +
import pandas as pd
df = pd.DataFrame({'x': [1, 2, 3],
'y': [3, 1, 4]})
alt.Chart(df).mark_point().encode(
x='x:Q',
y='y:Q',
color='color:Q' # <-- this field does not exist in the data!
)
# -
# Check the spelling of your files and print the data source to confirm that the data and fields exist. For instance, here you see that `color` is not a vaid field.
df.head()
# ## Invalid Specifications
#
# Another common issue is creating an invalid specification and getting an error.
# ### Invalid Properties
#
# Altair might show an `SchemaValidationError` or `ValueError`. Read the error message carefully. Usually it will tell you what is going wrong.
# For example, if you forget the mark type, you will see this `SchemaValidationError`.
alt.Chart(data.cars()).encode(
y='Horsepower'
)
# Or if you use a non-existent channel, you get a `ValueError`.
alt.Chart(data.cars()).mark_point().encode(
z='Horsepower'
)
# ## Properties are Being Ignored
#
# Altair might ignore a property that you specified. In the chart below, we are using a `text` channel, which is only compatible with `mark_text`. You do not see an error or a warning about this in the notebook. However, the underlying Vega-Lite library will show a warning in the browser console. Press <kbd>Alt</kbd>+<kbd>Cmd</kbd>+<kbd>I</kbd> on Mac or <kbd>Alt</kbd>+<kbd>Ctrl</kbd>+<kbd>I</kbd> on Windows and Linux to open the developer tools and click on the `Console` tab. When you run the example in the cell below, you will see a the following warning.
#
# ```
# WARN text dropped as it is incompatible with "bar".
# ```
alt.Chart(data.cars()).mark_bar().encode(
y='mean(Horsepower)',
text='mean(Acceleration)'
)
# If you find yourself debugging issues related to Vega-Lite, you can open the chart in the [Vega Editor](https://vega.github.io/editor/) either by clicking on the "Open in Vega Editor" link at the bottom of the chart or in the action menu (click to open) at the top right of a chart. The Vega Editor provides additional debugging but you will be writing Vega-Lite JSON instead of Altair in Python.
#
# **Note**: The Vega Editor may be using a newer version of Vega-Lite and so the behavior may vary.
# ## Asking for Help
#
# If you find a problem with Altair and get stuck, you can ask a question on Stack Overflow. Ask your question with the `altair` and `vega-lite` tags. You can find a list of questions people have asked before [here](https://stackoverflow.com/questions/tagged/altair).
# ## Reporting Issues
#
# If you find a problem with Altair and believe it is a bug, please [create an issue in the Altair GitHub repo](https://github.com/altair-viz/altair/issues/new) with a description of your problem. If you believe the issue is related to the underlying Vega-Lite library, please [create an issue in the Vega-Lite GitHub repo](https://github.com/vega/vega-lite/issues/new).
| altair_debugging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Azure Cognitive Search sample
# ## Passing Images as Binary File References
#
# Skillsets that pass images to custom skills use a binary file reference to serialize the images before passing them to other skills. This sample demonstrates how skills can be configured to accept image inputs and return image outputs.
#
# While the other steps in this skillset, such as OCR and redaction, have relevance, the key takeaway is configuring and passing binary file references. The custom skill does the heavy lifting. Each input record contains an image that is serialized as a `Base64` encoded string. The input also contains the layout text of image, as returned from the OCR skill. Upon receiving the input, the custom skill segments the image into smaller images based on the coordinates of the layout text. It then returns a list of images, each `Base64` encoded, back to the skillset. While this is not a particularly realistic exercise, it demonstrates techniques that could be leverage in more interesting ways, such as in a [Custom Vision](https://github.com/Azure-Samples/azure-search-power-skills/tree/master/Vision/CustomVision) skill that performs useful inferences on your images.
#
# For more information about the skills used in this example, see [OCR skill](https://docs.microsoft.com/azure/search/cognitive-search-skill-ocr), [PII skill](https://docs.microsoft.com/azure/search/cognitive-search-skill-pii-detection), and [custom skills](https://docs.microsoft.com/azure/search/cognitive-search-custom-skill-web-api).
#
#
# ### Prerequisites
#
# + [Azure subscription](https://Azure.Microsoft.com/subscription/free)
# + [Azure Cognitive Search service](https://docs.microsoft.com/azure/search/search-create-service-portal) (get the full service endpoint and an admin API key)
# + [Azure Blob storage service](https://docs.microsoft.com/azure/storage/common/storage-account-create) (get the connection string)
# + [Python 3.6+](https://www.python.org/downloads/)
# + [Jupyter Notebook](https://jupyter.org/install)
# + [Visual Studio Code](https://code.visualstudio.com/download) with the [Azure Functions extension](https://marketplace.visualstudio.com/items?itemName=ms-azuretools.vscode-azurefunctions) and the [Python extension](https://marketplace.visualstudio.com/items?itemName=ms-python.python)
#
# If you adapt this exercise to include more image files, add [Azure Cognitive Services](https://docs.microsoft.com/azure/cognitive-services/cognitive-services-apis-create-account).
# ### Configure inputs
#
# Follow the instructions in the [readme](https://github.com/Azure-Samples/azure-search-python-samples/blob/master/Image-Processing/README.md) to set up the inputs used by the indexer, data source, and skillset.
#
# Besides connection information, you will need a blob container for the sample JPEG file, and a function app that provides the code used in the custom skill. All the necessary files are provided. The `SplitImage` folder contains an Azure function that will accept an input in the [custom skill format](https://docs.microsoft.com/azure/search/cognitive-search-custom-skill-web-api#skill-inputs).
# ### Create the enrichment pipeline
# In the next few steps, configure the Cognitive Search enrichment pipeline, creating these objects on your search service:
# 1. Create an indexer data source. The data source references a blob storage container with at least one image file.
# 2. Create a skillset that performs image analysis. The skillset references a Cognitive Services account, a custom function app, and a knowledge store.
# 3. Create a search index.
# 4. Create an indexer to move documents from the data source to the index while invoking the skillset.
#
# +
# !pip install azure-storage-blob
import os
import json
import requests
# Configure all required variables for this exerences. Replace each with the credentials from your accounts.
# Replace with a full search service endpoint the format "https://searchservicename.search.windows.net"
# Paste in an admin API key. Both values can be obtained from the Azure portal.
search_service = "https://<YOUR-SEARCH-SERVICE-NAME>.search.windows.net"
api_key = '<YOUR-SEARCH-ADMIN-API-KEY>'
# Leave the API version and content_type as they are listed here.
api_version = '2020-06-30'
content_type = 'application/json'
# Replace with a Cognitive Services account name and all-in-one key.
# Required only if processing more than 20 documents
cog_svcs_key = ''
cog_svcs_acct = ''
# Your Azure Storage account will be used for the datasource input and knowledge store output
# Replace with a connection string to your Azure Storage account.
STORAGECONNSTRING = "DefaultEndpointsProtocol=https;AccountName=<YOUR-STORAGE-ACCOUNT>;AccountKey=<YOUR-ACCOUNT-KEY>;EndpointSuffix=core.windows.net"
# Replace with the blob container containing your image file
datasource_container = 'bfr-sample'
# Container where the sliced images will be projected to. Use the value provided below.
know_store_container = "obfuscated"
# Replace with the Function HTTP URL of the app deployed to Azure Function
skill_uri = "<YOUR-FUNCTION-APP-URL"
# -
# Create a helper function to invoke the Cognitive Search REST APIs.
# +
def construct_Url(service, resource, resource_name, action, api_version):
if resource_name:
if action:
return service + '/'+ resource + '/' + resource_name + '/' + action + '?api-version=' + api_version
else:
return service + '/'+ resource + '/' + resource_name + '?api-version=' + api_version
else:
return service + '/'+ resource + '?api-version=' + api_version
headers = {'api-key': api_key, 'Content-Type': content_type}
# Test out the URLs to ensure that the configuration works
print(construct_Url(search_service, "indexes", "bfr-sample", "analyze", api_version))
print(construct_Url(search_service, "indexes", "bfr-sample", None, api_version))
print(construct_Url(search_service, "indexers", None, None, api_version))
# -
# #### Create the data source
# +
container = datasource_container
datsource_def = {
'name': f'{datasource_container}-ds',
'description': f'Datasource containing files with sample images',
'type': 'azureblob',
'subtype': None,
'credentials': {
'connectionString': f'{STORAGECONNSTRING}'
},
'container': {
'name': f'{datasource_container}'
},
}
r = requests.post(construct_Url(search_service, "datasources", None, None, api_version), data=json.dumps(datsource_def), headers=headers)
print(r)
res = r.json()
print(json.dumps(res, indent=2))
# -
# #### Create the skillset
#
# Binary image references are passed as inputs and outputs, starting with "/document/normalized_images/*" in the OCR skill. OCR output is text and layout. Only the text component is passed to PIIDectection for analysis and redactive formatting. In the custom skill, the image is sliced into component parts (text and layout from OCR, and PII entity created in the PIIDetection step).
#
# Besides skills, a skillset also specifies the knowledge store projections that shape the final output in Blob storage.
# +
skillset_name = f'{datasource_container}-ss'
skillset_def = {
'name': f'{skillset_name}',
'description': 'Skillset to demonstrate passing images to custom skills',
'skills': [
{
"@odata.type": "#Microsoft.Skills.Vision.OcrSkill",
"name": "OCRSkill",
"description": "OCR Skill",
"context": "/document/normalized_images/*",
"textExtractionAlgorithm": None,
"lineEnding": "Space",
"defaultLanguageCode": "en",
"detectOrientation": True,
"inputs": [
{
"name": "image",
"source": "/document/normalized_images/*"
}
],
"outputs": [
{
"name": "text",
"targetName": "text"
},
{
"name": "layoutText",
"targetName": "layoutText"
}
]
},
{
"@odata.type": "#Microsoft.Skills.Text.PIIDetectionSkill",
"name": "#1",
"description": "",
"context": "/document/merged_content",
"defaultLanguageCode": "en",
"minimumPrecision": 0.5,
"maskingMode": "replace",
"maskingCharacter": "*",
"inputs": [
{
"name": "text",
"source": "/document/merged_content"
}
],
"outputs": [
{
"name": "piiEntities",
"targetName": "pii_entities"
},
{
"name": "maskedText",
"targetName": "masked_text"
}
]
},
{
"@odata.type": "#Microsoft.Skills.Custom.WebApiSkill",
"name": "ImageSkill",
"description": "Segment Images",
"context": "/document/normalized_images/*",
"uri": f'{skill_uri}',
"httpMethod": "POST",
"timeout": "PT30S",
"batchSize": 1000,
"degreeOfParallelism": 1,
"inputs": [
{
"name": "image",
"source": "/document/normalized_images/*"
},
{
"name": "layoutText",
"source": "/document/normalized_images/*/layoutText"
},
{
"name": "pii_entities",
"source": "/document/merged_content/pii_entities"
}
],
"outputs": [
{
"name": "slices",
"targetName": "slices"
},
{
"name": "original",
"targetName": "original"
}
],
"httpHeaders": {}
},
{
"@odata.type": "#Microsoft.Skills.Text.MergeSkill",
"name": "MergeSkill",
"description": "Merge results from cracking with OCR text",
"context": "/document",
"insertPreTag": " ",
"insertPostTag": " ",
"inputs": [
{
"name": "text",
"source": "/document/content"
},
{
"name": "itemsToInsert",
"source": "/document/normalized_images/*/text"
},
{
"name": "offsets",
"source": "/document/normalized_images/*/contentOffset"
}
],
"outputs": [
{
"name": "mergedText",
"targetName": "merged_content"
}
]
}
],
'cognitiveServices':None,
'knowledgeStore': {
'storageConnectionString': f'{STORAGECONNSTRING}',
'projections': [
{
"tables": [],
"objects": [
{
"storageContainer": "layout",
"referenceKeyName": None,
"generatedKeyName": "layoutKey",
"source": "/document/normalized_images/*/layoutText",
"sourceContext": None,
"inputs": []
}
],
"files": [
{
"storageContainer": "slices",
"referenceKeyName": None,
"generatedKeyName": "slicesKey",
"source": "/document/normalized_images/*/slices/*",
"sourceContext": None,
"inputs": []
},
{
"storageContainer": "images",
"referenceKeyName": None,
"generatedKeyName": "imageKey",
"source": "/document/normalized_images/*",
"sourceContext": None,
"inputs": []
},
{
"storageContainer": f'{know_store_container}',
"referenceKeyName": None,
"generatedKeyName": "originalKey",
"source": "/document/normalized_images/*/original",
"sourceContext": None,
"inputs": []
}
]
}
]
}
}
r = requests.put(construct_Url(search_service, "skillsets", skillset_name, None, api_version), data=json.dumps(skillset_def), headers=headers)
print(r)
res = r.json()
print(json.dumps(res, indent=2))
# -
# #### Create the index
#
# A search index isn't used in this exercise, but because it's an indexer requirement, you'll create one anyway. You can use Search Explorer in the Azure portal to query the index on your own. It will contain text extracted from the image.
indexname = f'{datasource_container}-idx'
index_def = {
"name":f'{indexname}',
"defaultScoringProfile": "",
"fields": [
{
"name": "image_text",
"type": "Collection(Edm.String)",
"facetable": False,
"filterable": False,
"retrievable": True,
"searchable": True,
"analyzer": "standard.lucene",
"indexAnalyzer": None,
"searchAnalyzer": None,
"synonymMaps": [],
"fields": []
},
{
"name": "content",
"type": "Edm.String",
"facetable": False,
"filterable": False,
"key": False,
"retrievable": True,
"searchable": True,
"sortable": False,
"analyzer": "standard.lucene",
"indexAnalyzer": None,
"searchAnalyzer": None,
"synonymMaps": [],
"fields": []
},
{
"name": "metadata_storage_content_type",
"type": "Edm.String",
"searchable": False,
"filterable": False,
"retrievable": True,
"sortable": False,
"facetable": False,
"key": False,
"indexAnalyzer": None,
"searchAnalyzer": None,
"analyzer": None,
"synonymMaps": []
},
{
"name": "metadata_storage_size",
"type": "Edm.Int64",
"searchable": False,
"filterable": False,
"retrievable": True,
"sortable": False,
"facetable": False,
"key": False,
"indexAnalyzer": None,
"searchAnalyzer": None,
"analyzer": None,
"synonymMaps": []
},
{
"name": "metadata_storage_last_modified",
"type": "Edm.DateTimeOffset",
"searchable": False,
"filterable": False,
"retrievable": True,
"sortable": False,
"facetable": False,
"key": False,
"indexAnalyzer": None,
"searchAnalyzer": None,
"analyzer": None,
"synonymMaps": []
},
{
"name": "metadata_storage_content_md5",
"type": "Edm.String",
"searchable": False,
"filterable": False,
"retrievable": True,
"sortable": False,
"facetable": False,
"key": False,
"indexAnalyzer": None,
"searchAnalyzer": None,
"analyzer": None,
"synonymMaps": []
},
{
"name": "metadata_storage_name",
"type": "Edm.String",
"searchable": False,
"filterable": False,
"retrievable": True,
"sortable": False,
"facetable": False,
"key": False,
"indexAnalyzer": None,
"searchAnalyzer": None,
"analyzer": None,
"synonymMaps": []
},
{
"name": "metadata_storage_path",
"type": "Edm.String",
"searchable": False,
"filterable": False,
"retrievable": True,
"sortable": False,
"facetable": False,
"key": True,
"indexAnalyzer": None,
"searchAnalyzer": None,
"analyzer": None,
"synonymMaps": []
},
{
"name": "metadata_storage_file_extension",
"type": "Edm.String",
"searchable": False,
"filterable": False,
"retrievable": True,
"sortable": False,
"facetable": False,
"key": False,
"indexAnalyzer": None,
"searchAnalyzer": None,
"analyzer": None,
"synonymMaps": []
}
],
"scoringProfiles": [],
"corsOptions": None,
"suggesters": [
{
"name": "sg",
"searchMode": "analyzingInfixMatching",
"sourceFields": [
"metadata_storage_path"
]
}
],
"analyzers": [],
"tokenizers": [],
"tokenFilters": [],
"charFilters": [],
"encryptionKey": None,
"similarity": None
}
r = requests.post(construct_Url(search_service, "indexes", None, None, api_version), data=json.dumps(index_def), headers=headers)
print(r)
res = r.json()
print(json.dumps(res, indent=2))
# #### Create the indexer
#
# This step creates the index (you'll run it in a separate step). At run time, the indexer connects to the data source, invokes the skillset, and outputs results. This indexer is scheduled to run every two hours.
indexername = f'{datasource_container}-idxr'
indexer_def = {
"name": f'{indexername}',
"description": "Indexer to enrich hotel reviews",
"dataSourceName": f'{datasource_container}-ds',
"skillsetName": f'{datasource_container}-ss',
"targetIndexName": f'{datasource_container}-idx',
"disabled": None,
"schedule": {
"interval": "PT2H",
"startTime": "0001-01-01T00:00:00Z"
},
"parameters": {
"batchSize": None,
"maxFailedItems": 0,
"maxFailedItemsPerBatch": 0,
"base64EncodeKeys": None,
"configuration": {
"dataToExtract": "contentAndMetadata",
"parsingMode": "default",
"imageAction": "generateNormalizedImages"
}
},
"fieldMappings": [
{
"sourceFieldName": "metadata_storage_path",
"targetFieldName": "metadata_storage_path",
"mappingFunction": {
"name": "base64Encode"
}
}
],
"outputFieldMappings": [
{
"sourceFieldName": "/document/normalized_images/*/text",
"targetFieldName": "image_text"
}
]
}
r = requests.post(construct_Url(search_service, "indexers", None, None, api_version), data=json.dumps(indexer_def), headers=headers)
print(r)
res = r.json()
print(json.dumps(res, indent=2))
# #### Run the indexer
#
# This step executes the indexer you just created. It will take several minutes to process.
r = requests.post(construct_Url(search_service, "indexers", indexername, "run", api_version), data=None, headers=headers)
print(r)
#res = r.json()
#print(json.dumps(res, indent=2))
# #### Check status
#
# The final step in this exercise is to view results. Before doing so, make sure the lastResult status message indicates "success", which means that the indexer completed its work successfully, and the revised image now exists in blob storage.
r = requests.get(construct_Url(search_service, "indexers", indexername, "status", api_version), data=None, headers=headers)
print(r)
res = r.json()
print(res["lastResult"])
# ### View Results
# The following cell downloads the output image so that you can verify skillset success. If you get an error, check the indexer status to make sure the indexer is finished and that there were no errors.
# +
from IPython.display import Image
import base64
from azure.storage.blob import ContainerClient
count = 0
container = ContainerClient.from_connection_string(conn_str=STORAGECONNSTRING, container_name=know_store_container)
blob_list = container.list_blobs()
for blob in blob_list:
print(blob.name + '\n')
blob_client = container.get_blob_client(blob.name)
with open("image" + str(count) + ".jpg", "wb") as my_blob:
download_stream = blob_client.download_blob()
my_blob.write(download_stream.readall())
count = count + 1
if(count == 3):
break
Image(filename='image0.jpg')
# -
# ### Next Steps
# In this exercise, you learned how to pass images into skills and return the modified images to the skillset for further processing.
#
# As a next step, you can start from scratch and build a [custom AML Skill](https://docs.microsoft.com/azure/search/cognitive-search-aml-skill) to perform inferences on images, or use the Custom Vision service to build a skill. The power skills github repository has a [sample custom vision skill](https://github.com/Azure-Samples/azure-search-power-skills/tree/master/Vision/CustomVision) to help you get started.
| Image-Processing/BFR_Sample_Rest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stoichiometric Structure
# Part I of this book introduced the basics of dynamic simulation. The process for setting up dynamic equations, their simulation, and processing of the output was presented in Chapter 3. Several concepts of dynamic analysis of networks were illustrated through the use of simple examples of chemical reaction mechanisms in Chapters 4 through 6. Most of these examples were conceptual and had limited direct biological relevance. In Chapter 7 we began to estimate the numerical values and ranges for key quantities in dynamic models. With this background, we now begin the process of addressing issues that are important when one builds realistic dynamic models of biological functions. We start by exploring the consequences of reaction bilinearity and that of the stoichiometric structure of a network. In Part III we then extend the material in this chapter to well-known metabolic pathways.
#
# **MASSpy** will be used to demonstrate some of the topics in this chapter.
from mass import (
MassModel, MassMetabolite, MassReaction,
Simulation, MassSolution, strip_time)
from mass.util.matrix import nullspace, left_nullspace
from mass.visualization import plot_time_profile, plot_phase_portrait
# Other useful packages are also imported at this time.
import numpy as np
import pandas as pd
import sympy as sym
import matplotlib.pyplot as plt
XL_FONT = {"size": "x-large"}
# ## Bilinearity in Biochemical Reactions
# ### Bilinear reactions
# They are of the form:
#
# $$\begin{equation} x + y \rightarrow z \tag{8.1} \end{equation}$$
#
# Two molecules come together to form a new molecule through the breaking and forming of covalent bonds, or a complex through the formation of hydrogen bonds. As illustrated with the pool formations in the bilinear examples in Chapter 4, such reactions come with moiety exchanges.
#
# ### Enzyme classification
# Enzyme catalyzed reactions are classified in to seven categories by Enzyme Commission (EC) numbers, see Figure 8.1a. These categories are: oxidoreductases, transferases, hydrolases, lyases, isomerases, and ligases. All these chemical transformations are bilinear with the exception of isomerases that simply rearrange a molecule without the participation of other reactants. Thus, the vast majority of biochemical reactions are bilinear. An overall pseudo-elementary representation (i.e., without treating the enzyme itself as a reactant, and just representing the un-catalyzed reaction) is bilinear.
#
# 
#
# **Figure 8.1:** The bilinear nature of biochemical reactions. (a) The classification of enzyme catalyzed reactions into seven categories by the enzyme commission (EC) number system. (b) The detailed view of the role of coenzymes and prosthetic groups in enzyme catalyzed reactions. Coenzymes are often referred to as cofactors. Both images from Koolman, 2005 (reprinted with permission).
#
# ### Coenzymes and prosthetic groups
# There are coenzymes and prosthetic groups that are involved in many biochemical reactions. These molecules are involved in group transfer reactions as illustrated in Figure 8.1b. They can transfer various chemical moieties or redox equivalents, see Table 8.1. Coenzymes act like a reactant and product in a reaction. They can work with many enzymes performing reactions that need them. Prosthetic groups associate with a particular enzyme to give it chemical functionalities that the protein itself does not have, Figure 8.1b. The heme group on hemoglobin is perhaps the most familiar example (see Chapter 13) that allows the protein tetramer to acquire a ferrous ion thus enabling the binding of oxygen. This binding allows the red blood cell to perform its oxygen delivery functions. There are many such capabilities 'grafted' onto proteins in the form of prosthetic groups. Many of the vitamins confer functions on protein complexes.
# ## Bilinearity Leads to a Tangle of Cycles
# ### Moiety exchange:
# Biochemical reaction networks are primarily made up of bilinear reactions. A fundamental consequence of this characteristic is a deliberate exchange of chemical moieties and properties between molecules. This exchange is illustrated in Figure 8.2. Here, an incoming molecule, $XA$, puts the moiety, $A$, onto a carrier molecule, $C$. The carrier molecule, now in a 'charged' form $(CA)$, can donate the $A$ moiety to another molecule, $Y$, to form $YA$. The terms _coenzyme,_ _cofactor_ or _carrier_ are used to describe the $C$ molecule.
#
# 
#
# **Figure 8.2:** Carrier $(C)$ mediated transfer of chemical moiety $A$ from compound $X$ to compound $Y$.
#
# ### Formation of cycles:
# The ability of bilinear reactions to exchange moieties in this fashion leads to the formation of distribution networks of chemical moieties and other properties of interest through the formation of a deliberate 'supply-chain' network. The structure of such a network must be thermodynamically feasible and conform to environmental constraints.
#
# Bilinearization in biochemical reaction networks leads to a 'tangle of cycles,' where different moieties and properties are being moved around the network. While a property of all biochemical networks, this trafficking of chemical and other properties is best known in metabolism. The major chemical properties that are being exchanged in metabolism are summarized in Table 8.1. These properties include energy, redox potential, one-carbon units, two-carbon units, amide groups, amine groups, etc. We now consider some specific cases.
#
# **Table 8.1:** Some activated carriers or coenzymes in metabolism, modified from Kurganov, 1983.
#
# 
#
# #### Example: Redox and energy trafficking in the core _E. coli_ metabolic pathways
# Energy metabolism revolves around the generation of redox potential and chemical energy in the form of high-energy phosphate bonds. The degradation of substrates through a series of chemical reactions culminates in the storage of these properties on key carrier molecules; see Table 8.1.
#
# The core metabolic pathways in _E. coli_ illustrate this feature, Figure 8.3. The transmission of redox equivalents through this core set of pathways is shown in Figure 8.3a. Each pathway is coupled to a redox carrier in a particular way. This pathway map can be drawn to show the cofactors rather than the primary metabolites and the main pathways (Figure 8.3b). This figure clearly shows how the cofactors interact and how the bilinear property of the stoichiometry of the core set of pathways leads to a tangle of cycles among the redox carriers.
#
# 
#
# **Figure 8.3:** The tangle of cycles in trafficking of redox potential (R) in _E. coli_ core metabolic pathways. (a) A map organized around the core pathways. (b) The tangle of cycles seen by viewing the cofactors and how they are coupled. Prepared by <NAME>.
#
# #### Example: Protein trafficking in signaling pathways
# Although the considerations above are illustrated using well-known metabolic pathways, these same features are also observed in signaling pathways. Incoming molecules (ligands) trigger a well-defined series of charging and discharging of the protein that make up a signaling network, most often with a phosphate group.
# ## Trafficking of High-Energy Phosphate Bonds
# Given the bilinear nature of biochemical reaction networks and the key role that cofactors play, we begin the process of building biologically meaningful simulation models by studying the use and formation of high-energy phosphate bonds. Cellular energy is stored in high-energy phosphate bonds in ATP. The dynamic balance of the rates of use and formation of ATP is thus a common denominator in all cellular processes, and thus foundational to the living process. We study the dynamic properties of this system in a bottom-up fashion by starting with its simple elements and making the description progressively more complicated. Throughout the text we make explicit use of the basic methods in **MASSpy**.
#
# 
#
# **Figure 8.4:** Representation of the exchange of high energy phosphate bonds among the adenosine phosphates. (a) The chemical reactions. (b) The molecules with open circles showing the "vacant" places for high energy bonds. The capacity to carry high-energy phosphate bonds, the occupancy of high-energy bonds, and the energy charge are shown. (c) The reaction schema of (a) in pictorial form. The solid squares represent AMP and the solid circles the high energy phosphate bonds. (d) The same concepts as in (b) represented in pictorial form.
#
# ### Distribution of high-energy phosphate groups: adenylate kinase (EC 2.7.4.3)
# The Adenylate Kinase is an important part in intracellular energy homeostasis. Adenylate Kinase is a phosphotransferase enzyme and it is the enzyme responsible for the redistribution of the phosphate groups among the adenosine phosphates. The redistribution reaction the Adenylate Kinase catalyzes is seen in Figure 8.4a.
#
# #### The mass balance: adenylate kinase
# The redistribution of the phosphate groups among the adenosine phosphates by the adenylate kinase is given by the following kinetic equations:
#
# $$\begin{equation} \frac{d\text{ATP}}{dt} = v_{\mathrm{distr}}, \ \frac{d\text{ADP}}{dt} = -2\ v_{\mathrm{distr}}, \ \frac{d\text{AMP}}{dt} = v_{\mathrm{distr}} \tag{8.2} \end{equation}$$
#
# #### The reaction rates: adenylate kinase
# The mass action form of these basic reaction rates are
#
# $$\begin{equation} v_{\mathrm{distr}} = k_{\mathrm{distr}}^\rightarrow\text{ADP}^2 - k_{\mathrm{distr}}^\leftarrow\text{ATP}*\text{AMP} \tag{8.3} \end{equation}$$
#
# #### Numerical values: adenylate kinase
# The approximate numerical values of the parameters in this system can be estimated. In metabolically active tissues, the ATP concentration is about 1.6 mM, the ADP concentration is about 0.4 mM, and the AMP concentration is about 0.1 mM. Total adenosine phosphates are thus about 2.1 mM. Because this reaction is considerably faster compared to other metabolic processes, we set $k_{\mathrm{distr}}^\rightarrow$ to 1000/Min. $K_{\mathrm{distr}}$ for the distribution reaction is approximately unity. We then construct a model of the redistribution of phosphate groups among the adenosine phosphates by adenylate kinase using the above constraints. This is simple reversible reaction that equilibrates quickly.
#
# 
#
# **Figure 8.5:** The redistribution of phosphate groups among the adenosine phosphates by adenylate kinase.
# +
phos_traffic = MassModel("Phosphate_Trafficking", array_type="DataFrame",
dtype=np.int64)
# Define metabolites
atp = MassMetabolite("atp")
adp = MassMetabolite("adp")
amp = MassMetabolite("amp")
# Define reactions
v_distr = MassReaction("distr")
v_distr.add_metabolites({adp: -2, amp: 1, atp:1})
# Add reactions to model
phos_traffic.add_reactions([v_distr])
# Define initial conditions and parameters
atp.ic = 1.6
adp.ic = 0.4
amp.ic = 0.1
v_distr.kf = 1000
v_distr.Keq = 1
# -
# #### Null spaces: adenylate kinase
# The stoichiometric matrix is basically a column vector.
phos_traffic.S
# It has an empty null space; i.e. zero dimensional.
nullspace(phos_traffic.S, rtol=1e-1)
# However, the left null space has two dimensions and it thus has two conservation pools.
# +
# Obtain left nullspace
lns = left_nullspace(phos_traffic.S, rtol=1e-1)
# Iterate through left nullspace,
# dividing by the smallest value in each row.
for i, row in enumerate(lns):
minval = np.min(abs(row[np.nonzero(row)]))
new_row = np.array(row/minval)
# Round to ensure the left nullspace is composed of only integers
lns[i] = np.array([round(value) for value in new_row])
# Ensure positive stoichiometric coefficients if all are negative
for i, space in enumerate(lns):
lns[i] = np.negative(space) if all([num <= 0 for num in space]) else space
# Create a pandas.DataFrame to represent the left nullspace
pd.DataFrame(lns, index=["Vacancy", "Occupancy"],
columns=phos_traffic.metabolites, dtype=np.int64)
# -
# The interpretation of these pools is remarkably interesting: the first one counts the number of high energy phosphate bonds in the system, while the second counts the number of vacant spots where high energy phosphate bonds can be added. The left null space is spanned by these two vectors that we can think of as a conjugate pair. Furthermore, the summation of the two is the total amount of the 'A' nucleotide in the system times two; ie the total number of possible high-energy phosphate bonds that the system can carry.
# Sum the elements of each row to obtain the capacity pool
capacity = np.array([np.sum(lns, axis=0)])
pd.DataFrame(capacity, index=["Capacity"],
columns=phos_traffic.metabolites, dtype=np.int64)
# Note that any activity of this reaction does not change the sizes of these two pools as the left null space is orthogonal to the reaction vector (or the column vector of $(\textbf{S})$, that represents the direction of motion.
#
# ### Using and generating high-energy phosphate groups
# We now introduce the 'use' and 'formation' reactions for ATP into the above system. These represent aggregate processes in the cell using and forming high energy bonds.
#
# #### The mass balances: trafficking high-energy phosphate bonds
# $$\begin{align} \frac{d\text{ATP}}{dt} &= -v_{\mathrm{use}} + v_{\mathrm{form}} + v_{\mathrm{distr}} \tag{8.4} \\ \frac{d\text{ADP}}{dt} &= v_{\mathrm{use}} - v_{\mathrm{form}} - 2\ v_{\mathrm{distr}} \tag{8.5} \\ \frac{d\text{AMP}}{dt} &= v_{\mathrm{distr}} \tag{8.6} \end{align}$$
#
# where $v_{\mathrm{use}}$ is the rate of use of ATP, $v_{\mathrm{form}}$ is the rate of formation of ATP, and, as above, $v_{\mathrm{distr}}$ is the redistribution of the phosphate group among the adenosine phosphates by adenylate kinase.
#
# #### The reaction rates: trafficking high-energy phosphate bonds
# Elementary mass action form for the two additional rate equations are
#
# $$\begin{equation} v_{\mathrm{use}} = k_{\mathrm{use}}^\rightarrow \text{ATP},\ v_{\mathrm{form}} = k_{\mathrm{form}}^\rightarrow\text{ADP}\tag{8.7} \end{equation}$$
#
# #### Numerical values: trafficking high-energy phosphate bonds
# We use the equilibrium concentrations from the distribution model and estimate in the numerical values for the rate constants of ATP use and formation based on the fact that typical use and formation rates of ATP are about 10 mM/min. Using the steady state concentrations, we can calculate $k_{\mathrm{use}}^\rightarrow$ and $k_{\mathrm{form}}^\rightarrow$, resulting in $k_{\mathrm{use}}^\rightarrow=6.25\ min^{-1}$ and $k_{\mathrm{form}}^\rightarrow=25\ min^{-1}$. These constants are known as Pseudo-Elementary Rate Constants (PERCs). They are a ratio between the flux through a reaction and the concentrations of the involved species, and the simplify the network dynamic analysis. However they are condition dependent and result in a condition dependent kinetic model. What comprises the PERCs is explored further in the later chapters.
#
# We update the distribution model with the additional reactions and parameters.
#
# 
#
# **Figure 8.6:** The trafficking of high-energy phosphate bonds.
# +
# Create utilization reaction
v_use = MassReaction("use", reversible=False)
v_use.add_metabolites({atp: -1, adp: 1})
v_use.kf = 6.25
# Create formation reaction
v_form = MassReaction("form", reversible=False)
v_form.add_metabolites({adp: -1, atp: 1})
v_form.kf = 25
# Add reactions to model
phos_traffic.add_reactions([v_use, v_form])
# View rate of distribution reaction
print(v_distr.rate)
# -
# From the model we also see that the net rate for the redistribution of high-energy bonds is
#
# $$\begin{align} v_{\mathrm{distr}} &= k_{\mathrm{distr}}^\rightarrow\ \text{ADP}^2 - k_{\mathrm{distr}}^\leftarrow\text{ATP}*\text{AMP} \\ &= k_{\mathrm{distr}}^\rightarrow( \text{ADP}^2 - \text{ATP}*\text{AMP}/K_{\mathrm{distr}}) \end{align} \tag{8.8}$$
#
# #### Null spaces: trafficking high-energy phosphate bonds
# Now the stoichiometric matrix three columns.
phos_traffic.S
# It has a one-dimensional null space, that represents an internal loop as the use and formation reactions are the exact opposites of each other.
# +
# Obtain nullspace
ns = nullspace(phos_traffic.S, rtol=1e-1)
# Transpose and iterate through nullspace,
# dividing by the smallest value in each row.
ns = ns.T
for i, row in enumerate(ns):
minval = np.min(abs(row[np.nonzero(row)]))
new_row = np.array(row/minval)
# Round to ensure the nullspace is composed of only integers
ns[i] = np.array([round(value) for value in new_row])
# Ensure positive stoichiometric coefficients if all are negative
for i, space in enumerate(ns):
ns[i] = np.negative(space) if all([num <= 0 for num in space]) else space
# Revert transpose
ns = ns.T
# Create a pandas.DataFrame to represent the nullspace
pd.DataFrame(ns, index=[rxn.id for rxn in phos_traffic.reactions],
columns=["Path 1"], dtype=np.int64)
# -
# The left null space is now one-dimensional;
# +
# Obtain left nullspace
lns = left_nullspace(phos_traffic.S, rtol=1e-1)
# Iterate through left nullspace,
# dividing by the smallest value in each row.
for i, row in enumerate(lns):
minval = np.min(abs(row[np.nonzero(row)]))
new_row = np.array(row/minval)
# Round to ensure the left nullspace is composed of only integers
lns[i] = np.array([round(value) for value in new_row])
# Ensure positive stoichiometric coefficients if all are negative
for i, space in enumerate(lns):
lns[i] = np.negative(space) if all([num <= 0 for num in space]) else space
# Create a pandas.DataFrame to represent the left nullspace
pd.DataFrame(lns, index=["Total AxP"],
columns=phos_traffic.metabolites, dtype=np.int64)
# -
# #### Dynamic simulations: trafficking high-energy phosphate bonds
# The system is steady at the initial conditions given
# + tags=["remove_cell"]
t0, tf = (0, 1e3)
sim = Simulation(phos_traffic, verbose=True)
conc_sol, flux_sol = sim.simulate(
phos_traffic, time=(t0, tf, tf*10 + 1), interpolate=True,
verbose=True)
# + tags=["remove_cell"]
fig, axes = plt.subplots(nrows=2, ncols=1, figsize=(8, 4),
)
(ax1, ax2) = axes.flatten()
plot_time_profile(
conc_sol, ax=ax1,
legend="right outside",plot_function="semilogx",
xlabel="Time [min]", ylabel="Concentrations [mM]",
title=("Concentration Profile", XL_FONT));
plot_time_profile(
flux_sol, ax=ax2,
legend="right outside", plot_function="semilogx",
xlabel="Time [min]", ylabel="Fluxes [mM/min]",
title=("Flux Profile", XL_FONT));
# -
# We can induce motion in the system by taking 0.2 mM of ADP and splitting it into 0.1 mM addition to AMP and ATP, and set the initial conditions as ATP is 1.7 mM, ADP is 0.2 mM, and AMP is 0.2 mM and simulate the dynamic response. We graph the concentration profiles, as well as the two pools and the disequilibrium variable: $\text{ADP}^2 - \text{ATP}*\text{AMP}$ that is zero at the equilibrium
# +
# Define pools and perturbations
pools = {"Occupancy": "adp + 2*atp",
"Vacancy": "adp + 2*amp",
"Disequilibrium": "adp**2 - atp*amp"}
# Simulate with disturbance
conc_sol, flux_sol = sim.simulate(
phos_traffic, time=(t0, tf, tf*10 + 1),
perturbations={"atp": 1.7, "adp": 0.2, "amp": 0.2})
# Determine pools
for pool_id, equation_str in pools.items():
conc_sol.make_aggregate_solution(
pool_id, equation=equation_str, update=True)
# Visualize solutions
fig_8_7, axes = plt.subplots(nrows=3, ncols=1, figsize=(8, 6),)
(ax1, ax2, ax3) = axes.flatten()
plot_time_profile(
conc_sol, ax=ax1, observable=phos_traffic.metabolites,
legend="right outside", plot_function="semilogx", ylim=(0, 1.8),
xlabel="Time [min]", ylabel="Concentration [mM]",
title=("(a) Concentration Profile", XL_FONT));
plot_time_profile(
conc_sol, observable=["Occupancy", "Vacancy"], ax=ax2,
legend="right outside", plot_function="semilogx", ylim=(0., 4.),
xlabel="Time [min]", ylabel="Concentration [mM]",
title=("(b) Occupancy and Vacancy Pools", XL_FONT));
plot_time_profile(
conc_sol, observable=["Disequilibrium"], ax=ax3,
legend="right outside", plot_function="semilogx", ylim=(-.4, 0.1),
xlabel="Time [min]", ylabel="Concentration [mM]",
title=("(c) Disequilibrium Variable", XL_FONT));
# -
# **Figure 8.7:** The time response of the adenylate kinase reaction ("distr") and with the addition of ATP use and formation to a change in the initial conditions. (a) The concentrations. (b) The occupancy and capacity pools. (c) The disequilibrium variable.
#
# #### Towards a realistic simulation of a dynamic response
# Next, we simulate the response of this system to a more realistic perturbation: a 50% increase in the rate of ATP use. This would represent a sudden increase in energy use by a cell. At time zero, we have the network in a steady state and we change $k_{\mathrm{use}}^\rightarrow$ from $6.25/min$ to $1.5*6.25=9.375/min$, and the rate of ATP use instantly becomes 15 mM/min.
#
# The response of the system is perhaps best visualized by showing the phase portrait of the rate of ATP use versus ATP formation. Prior to the increased load, the system is on the 45 degree line, where the rate of ATP formation and use balances. Then at time zero it is instantly imbalanced by changing $k_{\mathrm{use}}^\rightarrow$ above or below its initial value. If $k_{\mathrm{use}}^\rightarrow$ is increased then the initial point moved into the region where more ATP is used than formed. From this initial perturbation the response of the system is to move directly towards the 45 degree line to regain balance between ATP use and formation.
# +
t0, tf = (0, 1e3)
# Simulate with disturbance
conc_sol, flux_sol = sim.simulate(
phos_traffic, time=(t0, tf, tf*10 + 1),
perturbations={"kf_use": "kf_use * 1.5"},
verbose=True)
# Determine pools
for pool_id, equation_str in pools.items():
conc_sol.make_aggregate_solution(
pool_id, equation=equation_str, update=True)
# + tags=["remove_cell"]
fig_8_8 = plt.figure(figsize=(15, 5))
gs = fig_8_8.add_gridspec(nrows=3, ncols=2, width_ratios=[1, 1.5])
ax1 = fig_8_8.add_subplot(gs[:, 0])
ax2 = fig_8_8.add_subplot(gs[0, 1])
ax3 = fig_8_8.add_subplot(gs[1, 1])
ax4 = fig_8_8.add_subplot(gs[2, 1])
label = "{0} [mM/min]"
plot_phase_portrait(
flux_sol, x=v_use, y=v_form, ax=ax1,
time_vector=np.linspace(t0, 1, int(1e4)),
xlabel=label.format(v_use.id), ylabel=label.format(v_form.id),
xlim=(4, 21), ylim=(4, 21),
title=("(a) Phase Portrait of ATP use vs. formation", XL_FONT),
annotate_time_points="endpoints",
annotate_time_points_labels=True);
line_data = [i for i in range(0, 22)]
ax1.plot(line_data, line_data, ls="--", color="black")
ax1.annotate("use < form", xy=(6, 15))
ax1.annotate("use > form", xy=(15, 6))
ax1.annotate("Steady-state line:\n use=form", xy=(15, 19))
ax1.annotate("initial perturbation", xy=(9.5, 9), xycoords="data")
ax1.annotate("", xy=(flux_sol[v_use.id][0], flux_sol[v_form.id][0]),
xytext=(10, 10), textcoords="data",
arrowprops=dict(arrowstyle="->",connectionstyle="arc3"))
plot_time_profile(
conc_sol, observable=phos_traffic.metabolites,
ax=ax2, legend="right outside",
time_vector=np.linspace(t0, 1, int(1e5)),
xlim=(t0, 1), ylim=(0, 2),
xlabel="Time [min]", ylabel="Concentration [mM]",
title=("(b) Concentration Profiles", XL_FONT));
plot_time_profile(
flux_sol, observable=[v_use],
ax=ax3, legend="right outside",
time_vector=np.linspace(t0, 1, int(1e5)),
xlim=(t0, 1), ylim=(12, 16),
xlabel="Time [min]", ylabel="Flux [mM/min]",
title=("(c) Net ATP use", XL_FONT));
plot_time_profile(
conc_sol, observable="Disequilibrium",
ax=ax4, legend="right outside",
time_vector=np.linspace(t0, 1, int(1e5)), plot_function="semilogx",
xlabel="Time [min]", ylabel="Concentration [mM]",
xlim=(1e-6, 1), ylim=(-.0001, 0.0015),
title=("(d) Disequilibrium", XL_FONT));
fig_8_8.tight_layout()
# -
# **Figure 8.8:** Dynamic responses for Eqs (8.4 - 8.8). (a) The phase portrait for the rates of use and formation of ATP. (b) The concentrations of ATP, ADP, and AMP. (c) Net ATP use (d) The disequilibrium variable for Adenylate kinase.
#
# #### Pooling and interpretation: trafficking high-energy phosphate bonds
# Since AMP is not being synthesized and degraded, the sum of $\text{ATP} + \text{ADP} +\text{AMP}$, or the capacity to carry high-energy phosphate bonds, is a constant. The Atkinson's energy charge
#
# $$\begin{equation} \text{E.C.} = \frac{2\ \text{ATP} + \text{ADP}}{2\ \text{ATP}+\text{ADP}+\text{AMP}} = \frac{\text{occupancy}}{\text{capacity}} \tag{8.9} \end{equation}$$
#
# shows a monotonic decay to a lower state in response to the increased load (see Figure 8.9).
pools.update({"EC": "(2*atp + adp) / (2*(atp + adp + amp))"})
# Determine pools
for pool_id, equation_str in pools.items():
conc_sol.make_aggregate_solution(
pool_id, equation=equation_str, update=True)
# + tags=["remove_cell"]
fig_8_9, axes = plt.subplots(nrows=1, ncols=2, figsize=(12, 4))
(ax1, ax2) = axes.flatten()
plot_time_profile(
conc_sol, observable=["EC"], ax=ax1, legend="best",
plot_function="semilogx", ylim= (.7, 1),
xlabel="Time [min]", ylabel="Concentration [mM]",
title=("(a) Energy Charge", XL_FONT));
plot_time_profile(
conc_sol, observable=["Occupancy", "Vacancy"], ax=ax2,
legend="right outside", plot_function="semilogx",
ylim=(0., 4.), xlabel="Time [min]", ylabel="Concentration [mM]",
title=("(b) Charge Pools", XL_FONT));
fig_8_9.tight_layout()
# -
# **Figure 8.9:** (a) The Atkinson's energy charge (Eq. 8.9). (b) The occupancy and vacancy pools move in the opposite directions. Increasing the load drops the occupancy pool and increases the vacancy pool as the system becomes discharged. Reduced loads have the opposite reaction.
#
# 
#
# **Figure 8.10:** Graphical representation of the energy charge (x-direction) versus the capacity (y-direction). The drop in the charge is indicated by the arrow. The capacity is a constant in this case.
#
# ### Buffering the energy charge
# #### Reaction mechanism: E.C. buffering
# In many situations, there is a buffering effect on the energy charge by a coupled carrier of high energy bonds. This exchange is:
#
# $$\begin{equation} \text{ATP}\ + \text{B} \leftrightharpoons \text{ADP}\ + \text{BP} \tag{8.10} \end{equation}$$
#
# where the buffering molecule, $\text{B}$, picks up the high-energy phosphate group through a fast equilibrating reaction.
#
# 
#
# **Figure 8.11:** The trafficking of high-energy phosphate bonds with the buffer molecule exchange reaction.
# +
# Copy the model to create a new, yet identical model instance
phos_buffered = phos_traffic.copy()
phos_buffered.id += "_Buffered"
# Create the buffer metabolites
b = MassMetabolite("b")
bp = MassMetabolite("bp")
# Create the buffer reaction and add the metaolites
v_buff = MassReaction("buff")
v_buff.add_metabolites({atp:-1, b:-1, adp:1, bp:1})
# Update model
phos_buffered.add_reactions(v_buff)
# -
# The rate equation of the buffering reaction is:
print(strip_time(phos_buffered.rates[v_buff]))
# #### Examples of buffer molecules
# In Eq. (8.10), $\text{B}$ represents a phosphagen, which is a compound containing a high-energy phosphate bond that is used as energy storage to buffer the ATP/ADP ratio. The most well-known phosphagen is creatine, which is found in the muscles of mammals. Marine organisms have other phosphagens (arginine, taurocyamine, glycocyamine), while earthworms use lombricine (Nguyen, 1960).
#
# #### Buffering:
# When the reaction in Eq. 8.10 is at equilibrium we have
#
# $$\begin{equation} k_{\mathrm{buff}}^\rightarrow\text{ATP}*\text{B} = k_{\mathrm{buff}}^\leftarrow \text{ADP}*\text{BP} \tag{8.11} \end{equation}$$
#
# This equation can be rearranged as
#
# $$\begin{equation} 4 K_{\mathrm{buff}} = \text{BP}/\text{B} \tag{8.12} \end{equation}$$
#
# where $\text{ATP}/\text{ADP}=1.6/0.4=4$ in the steady state, and $K_{\mathrm{buff}} = k_{\mathrm{buff}}/k_{-buff}$. If the buffering molecule is present in a constant amount, then
#
# $$\begin{equation} \text{B}_{\mathrm{tot}} = \text{B} + \text{BP} \tag{8.13} \end{equation}$$
#
# We can rearrange equations (8.12) and (8.13) as:
#
# $$\begin{equation} \frac{\text{BP}}{\text{B}_{\mathrm{tot}}} = \frac{4 K_{\mathrm{buff}}}{4 K_{\mathrm{buff}} + 1} \tag{8.14} \end{equation}$$
#
# In this equation, $\text{B}_{\mathrm{tot}}$ is the capacity of the buffer to carry the high energy phosphate bond whereas $\text{BP}/\text{B}_{\mathrm{tot}}$ is the energy charge of the buffer.
#
# We note that the value of $K_{\mathrm{buff}}$ is a key variable. If $K_{\mathrm{buff}} = 1/4$ then the buffer is half charged at equilibrium, whereas if $K_{\mathrm{buff}}=1$ then the buffer is 80% charged. Thus, this numerical value (a thermodynamic quantity) is key and will specify the relative charge on the buffer and the adenosine phosphates. The effect of $K_{\mathrm{buff}}$ can be determined through simulation.
#
# #### Updating the model with the buffering reaction
# It is assumed that the buffering reaction is at equilibrium and that the amount of buffering molecules is constant:
# +
# Use sympy to set up a symbolic equation for the buffer equilibrium
buff_equilibrium = sym.Eq(
sym.S.Zero, strip_time(phos_buffered.rates[v_buff]))
# Set amount of buffer molecules
btot = 10
# Use sympy to set up a symbolic equation for the buffer pool
b_sym = sym.Symbol(b.id)
bp_sym = sym.Symbol(bp.id)
buff_pool = sym.Eq(b_sym + bp_sym, btot)
# Pretty print the equations
sym.pprint(buff_equilibrium)
sym.pprint(buff_pool)
# -
# Solve the equilibrium system:
# +
# Obtain a dict of ic values for substitution into the sympy expressions
ic_dict = {sym.Symbol(met.id): ic
for met, ic in phos_buffered.initial_conditions.items()}
# Substitute known concentrations
buff_equilibrium = buff_equilibrium.subs(ic_dict)
# Obtain solutions for B and BP
buff_sol = sym.solve([buff_equilibrium, buff_pool], [b_sym, bp_sym])
# Pretty print the equation
print(buff_sol)
# -
# Set $K_{\mathrm{buff}}$ and $k_{\mathrm{buff}}^\rightarrow$:
# +
v_buff.kf = 1000
v_buff.Keq = 1
# Obtain a dict of parameter values for substitution into the sympy expressions
param_dict = {
sym.Symbol(parameter): value
for parameter, value in v_buff.parameters.items()}
buffer_ics = {
phos_buffered.metabolites.get_by_id(str(met)): float(expr.subs(param_dict))
for met, expr in buff_sol.items()}
# Update initial conditions with buffer molecule concentrations
phos_buffered.update_initial_conditions(buffer_ics)
for met, ic in phos_buffered.initial_conditions.items():
print("{0}: {1} mM".format(met, ic))
# -
# #### Null spaces: E.C. buffering
# With the addition of the buffer, stoichiometric matrix four columns.
# + tags=["remove_cell"]
phos_buffered.S
# -
# It has still has a one-dimensional null space, that represents and internal loop as the use and formation reactions are the exact opposites of each other.
# + tags=["remove_cell"]
# Obtain nullspace
ns = nullspace(phos_buffered.S, rtol=1e-1)
# Transpose and iterate through nullspace,
# dividing by the smallest value in each row.
ns = ns.T
for i, row in enumerate(ns):
minval = np.min(abs(row[np.nonzero(row)]))
new_row = np.array(row/minval)
# Round to ensure the nullspace is composed of only integers
ns[i] = np.array([round(value) for value in new_row])
# Ensure positive stoichiometric coefficients if all are negative
for i, space in enumerate(ns):
ns[i] = np.negative(space) if all([num <= 0 for num in space]) else space
# Revert transpose
ns = ns.T
# Create a pandas.DataFrame to represent the nullspace
pd.DataFrame(ns, index=[rxn.id for rxn in phos_buffered.reactions],
columns=["Path 1"], dtype=np.int64)
# -
# The left null space is two-dimensional. It represents conservation of the nucleotide and the buffer molecule. Neither AxP or B is produced or destroyed in the model;
# + tags=["remove_cell"]
# Obtain left nullspace
lns = left_nullspace(phos_buffered.S, rtol=1e-1)
# Iterate through left nullspace,
# dividing by the smallest value in each row.
for i, row in enumerate(lns):
minval = np.min(abs(row[np.nonzero(row)]))
new_row = np.array(row/minval)
# Round to ensure the left nullspace is composed of only integers
lns[i] = np.array([round(value) for value in new_row])
# Ensure positive stoichiometric coefficients if all are negative
for i, space in enumerate(lns):
lns[i] = np.negative(space) if all([num <= 0 for num in space]) else space
# Create a pandas.DataFrame to represent the left nullspace
pd.DataFrame(lns, index=["Total AxP", "Total B"],
columns=phos_buffered.metabolites, dtype=np.int64)
# -
# #### Dynamic simulation: E.C. buffering
# The model is initially in steady state.
t0, tf = (0, 1e3)
sim = Simulation(phos_buffered, verbose=True)
conc_sol, flux_sol = sim.simulate(phos_buffered, time=(t0, tf, tf*10 + 1),
verbose=True)
# + tags=["remove_cell"]
fig, axes = plt.subplots(nrows=2, ncols=1, figsize=(8, 4),
)
(ax1, ax2) = axes.flatten()
plot_time_profile(
conc_sol, ax=ax1, legend="right outside",
plot_function="semilogx",
xlabel="Time [min]", ylabel="Concentrations [mM]",
title=("Concentration Profile", XL_FONT));
plot_time_profile(
flux_sol, ax=ax2, legend="right outside",
plot_function="semilogx",
xlabel="Time [min]", ylabel="Fluxes [mM/min]",
title=("Flux Profile", XL_FONT));
# -
# We can compare the flux dynamics of the buffered vs. unbuffered system. The buffered system has a much longer response time. Once again, we consider a simulation where we increase the ATP use rate by a 'multiplier' in this figure:
# +
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(8, 4),
)
buff_strs = ["unbuffered", "buffered"]
linestyles = ["--", "-"]
t0, tf = (0, 1e3)
# Simulate both models with the disturbance
for i, model in enumerate([phos_traffic, phos_buffered]):
sim = Simulation(model)
conc_sol, flux_sol = sim.simulate(
model, time=(t0, tf, tf*10 + 1),
perturbations={"kf_use": "kf_use * 1.5"})
plot_time_profile(
flux_sol, observable=["use", "form"], ax=ax,
legend=(["use " + buff_strs[i], "form " + buff_strs[i]],
"right outside"),
plot_function="semilogx",
xlabel="Time [min]", ylabel="Fluxes [mM/min]",
color=["red", "blue"], linestyle=linestyles[i])
# -
# **Figure 8.12:** The fluxes of ATP use and formation respond more slowly when the ATP buffer is present.
#
# The response of the adenosine phosphate system can be simulated in the presence of a buffer. We choose the parameters as $\text{B}_{\mathrm{tot}}=10\ mM$, $K_{\mathrm{buff}}=1$, and $k_{\mathrm{buff}}=1000/min$ and all other conditions as in Figure 8.8. The results of the simulation are shown in Figure 8.13. The time response of the energy charge is shown, along with the buffer charge $\text{BP}/\text{B}_{\mathrm{tot}}$. We see that the fast response in the energy charge is now slower as the initial reaction is buffered by release of the high energy bonds that are bound to the buffer. The overall change in the energy charge is the same: it goes from 0.86 to 0.78. The charge of the buffer drops from 0.80 to 0.73 at the same time.
#
# 
#
# **Figure 8.13:** Pictorial representation of the phosphate exchange among the adenosine phosphates and a buffering molecule. (a) The reaction schema. (b) A pictorial representation of the molecules, their charged states, and the definition of pooled variables*
#
# #### Pooling and interpretation: E.C. buffering
#
# A pictorial representation of the phosphate buffering is given in Figure 8.13. Here, a generalized definition of the overall phosphate charge is:
#
# $$\begin{equation} \text{overall charge} = \frac{\text{overall occupancy}}{\text{overall capacity}} = \frac{2\ \text{ATP}+\text{ADP}+\text{BP}}{2\ (\text{ATP}+\text{ADP}+\text{AMP})+\text{BP} + \text{B}} \tag{8.15} \end{equation}$$
#
# This combined charge system can be represented similarly to the representation in Figure 8.10. Figure 8.14 shows a stacking of the buffer and adenosine phosphate capacity versus their charge. The total capacity to carry high-energy bonds is now 14.2 mM. The overall charge is 0.82 (or 11.64 mM concentration of high-energy bonds) in the system before the perturbation. The increased load brings the overall charge down to 0.74.
#
# 
#
# **Figure 8.14:** The representation of the energy and buffer charge versus the capacity (in mM on y-axis). The lumping of the two quantities into ‘overall’ quantities is illustrated. The case considered corresponds to the simulation in Figure 8.15.
#
# To understand this effect, we first define more pools:
pools.update({
"BC": "bp / (bp + b)",
"Overall_Charge": "(2*atp + adp + bp) / (2*(atp + adp + amp) + bp + b)"})
# and then plot the dynamic responses of the pools:
fig_8_15, axes = plt.subplots(nrows=2, ncols=1, figsize=(8, 6),)
(ax1, ax2) = axes.flatten()
legend_labels = ["E.C. Unbuffered", "E.C. Buffered"]
for i, model in enumerate([phos_traffic, phos_buffered]):
sim = Simulation(model)
conc_sol, flux_sol = sim.simulate(
model, time=(t0, tf, tf*10 + 1),
perturbations={"kf_use": "kf_use * 1.5"})
# Determine pools
for pool_id, equation_str in pools.items():
# Skip buffered charge for model with no buffer
if i == 0 and pool_id in ["BC", "Overall_Charge"]:
continue
conc_sol.make_aggregate_solution(
pool_id, equation=equation_str, update=True)
if i == 1:
# Plot the charge pools for the buffered solution
plot_time_profile(
conc_sol, observable=["EC", "BC", "Overall_Charge"], ax=ax1,
legend=(["E.C.", "B.C.", "Overall Charge"], "right outside"),
xlabel="Time [min]", ylabel="Charge",
xlim=(t0, 1), ylim=(.7, .9),
title=("(a) Charge Pools of Buffered Model", XL_FONT));
# Compare the buffered and unbuffered solutions
plot_time_profile(
conc_sol, observable=["EC"], ax=ax2,
legend=(legend_labels[i], "right outside"),
xlabel="Time [min]", ylabel="Charge",
xlim=(t0, 1), ylim=(.7, .9),
title=("(b) E.C. Unbuffered Vs Buffered", XL_FONT));
fig_8_15.tight_layout()
# **Figure 8.15:** Dynamic responses for Eqs.(8.4 - 8.8) with the buffering effect (Eq. (8.10)). (a) The Atkinson's energy charge (Eq. (8.9)) the buffer charge (Eq. (8.14)), and the overall charge (Eq. (8.15) are shown as a function of time. (b) Comparison of the buffered and unbuffered energy charge. $B_{\mathrm{tot}}=10 mM$, $K_{\mathrm{buff}}=1$ and $k_{\mathrm{buff}}=1000$. All other conditions are as in Figure 8.8; i.e., we simulate the response to a 'multiplier' increase in $k_{\mathrm{use}}$. Note the slower response of the E.C. in panel (b) when the system is buffered.
#
# ### Open system: long term adjustment of the capacity
# #### Inputs and outputs:
# Although the rates of formation and degradation of AMP are low, their effects can be significant. These fluxes will determine the total amount of the adenosine phosphates and thus their capacity to carry high energy bonds. The additional elementary rate laws needed to account for the rate of AMP formation and drain are:
#
# $$\begin{equation} v_{\mathrm{form,\ AMP}} = b_{1}, \ v_{\mathrm{drain}} = k_{\mathrm{drain}} * \text{AMP} \tag{8.16} \end{equation}$$
#
# where $b_1$ is the net synthesis rate of AMP. The numerical values used are $b_{1}=0.03\ mM/min$ and $k_{\mathrm{drain}} = (0.03\ mM/min)/(0.1\ mM) = 0.3\ mM/min$.
#
# #### Updating the model for long term capacity adjustment
# Define the AMP exchange reaction:
#
# 
#
# **Figure 8.16:** The trafficking of high-energy phosphate bonds with the buffer molecule and AMP exchange reactions.
# +
# Copy the model to create a new, yet identical model instance
phos_open = phos_buffered.copy()
phos_open.id += "_Open"
# Get MassMetabolite amp assoicated with the new copied model
amp = phos_open.metabolites.amp
# Define AMP formation
b1 = MassReaction("b1", reversible=False)
b1.add_metabolites({amp:1})
b1.kf = 0.03
# Define AMP drain
drain = MassReaction("drain", reversible=False)
drain.add_metabolites({amp:-1})
drain.kf = 0.3
# Add reactions to the model
phos_open.add_reactions([b1, drain])
# Set custom rate for formation of AMP
phos_open.add_custom_rate(b1, custom_rate=b1.kf_str)
# Display the net rate for AMP synthesis and draining
rate = strip_time(phos_open.rates[b1] - phos_open.rates[drain])
print(rate)
# Substitute values to check if steady state
print(rate.subs({
sym.Symbol('amp'): amp.ic, # AMP concentration at steady state
sym.Symbol('kf_drain'): drain.kf, # forward rate constant for drain reaction
sym.Symbol('kf_b1'): b1.kf})) # Synthesis rate
# -
# With the specified parameters and initial conditions, the system is in a steady state, i.e. no net exchange of AMP.
# #### Null spaces: long term capacity adjustment
# With the addition of the AMP exchanges, stoichiometric matrix six columns.
# + tags=["remove_cell"]
phos_open.S
# -
# It has still has a two-dimensional null space, that 1) represents and internal loop as the use and formation reactions are the exact opposites of each other, as before, and 2) an exchange pathways of AMP coming into the system and leaving the system.
# + tags=["remove_cell"]
# Obtain nullspace
ns = nullspace(phos_open.S, rtol=1e-1)
# Transpose and iterate through nullspace,
# dividing by the smallest value in each row.
ns = ns.T
for i, row in enumerate(ns):
minval = np.min(abs(row[np.nonzero(row)]))
new_row = np.array(row/minval)
# Round to ensure the nullspace is composed of only integers
ns[i] = np.array([round(value) for value in new_row])
# Ensure positive stoichiometric coefficients if all are negative
for i, space in enumerate(ns):
ns[i] = np.negative(space) if all([num <= 0 for num in space]) else space
# Revert transpose
ns = ns.T
# Create a pandas.DataFrame to represent the nullspace
pd.DataFrame(ns, index=[r.id for r in phos_open.reactions],
columns=["Path 1", "Path 2"], dtype=np.int64)
# -
# The left null space becomes one-dimensional. The total amount of A is no longer conserved as AMP can now enter or leave the system, i.e. pathway 2) can have a net flux. The buffer molecule, B, on the other hand is always contained within the system
# + tags=["remove_cell"]
# Obtain left nullspace
lns = left_nullspace(phos_open.S, rtol=1e-1)
# Iterate through left nullspace,
# dividing by the smallest value in each row.
for i, row in enumerate(lns):
minval = np.min(abs(row[np.nonzero(row)]))
new_row = np.array(row/minval)
# Round to ensure the left nullspace is composed of only integers
lns[i] = np.array([round(value) for value in new_row])
# Ensure positive stoichiometric coefficients if all are negative
for i, space in enumerate(lns):
lns[i] = np.negative(space) if all([num <= 0 for num in space]) else space
# Create a pandas.DataFrame to represent the left nullspace
pd.DataFrame(lns, index=["Total B"],
columns=phos_open.metabolites, dtype=np.int64)
# -
# #### Dynamic simulations: long term capacity adjustment
# Initially, the open system is in a steady-state. Once again, we consider a simulation where we increase the ATP use rate by a 'multiplier'. This system has a bi-phasic response for the values of the kinetic constants. We can start the system in a steady state at $t=0^-$ and simulate the response for increasing the ATP load by shifting the value of $k_{\mathrm{use}}^\rightarrow$ by a 'multiplier' at $t=0$, as before. The initial rapid response is similar to what is shown in Figure 8.8a, where the concentration of ATP drops in response to the load and the concentrations of ADP and AMP rise. This initial response is followed by a much slower response where all three concentrations drop.
# + tags=["remove_cell"]
t0, tf = (0, 1e3)
sim = Simulation(phos_open, verbose=True)
sim.find_steady_state(models=phos_open, strategy="simulate")
conc_sol, flux_sol = sim.simulate(
phos_open, time=(t0, tf, tf*10 + 1),
perturbations={"kf_use": "kf_use * 1.5"})
pools.update({"Capacity": "2*(atp + adp + amp)"})
# Determine pools
for pool_id, equation_str in pools.items():
# Skip buffered charge for model with no buffer
conc_sol.make_aggregate_solution(
pool_id, equation=equation_str, update=True)
# + tags=["remove_cell"]
fig, axes = plt.subplots(nrows=2, ncols=1, figsize=(8, 6),
)
(ax1, ax2) = axes.flatten()
plot_time_profile(
conc_sol, ax=ax1, observable=phos_open.metabolites,
legend="right outside",
plot_function="semilogx",
xlabel="Time [min]", ylabel="Concentrations [mM]",
title=("Concentration Profile", XL_FONT));
plot_time_profile(
flux_sol, ax=ax2, observable=phos_open.reactions,
legend="right outside",
plot_function="semilogx",
xlabel="Time [min]", ylabel="Fluxes [mM/min]",
title=("Flux Profile", XL_FONT));
fig.tight_layout()
# -
# #### Interpretation of the bi-phasic response
#
# This bi-phasic response can be examined further by looking at dynamic phase portraits of key fluxes (Figure 8.17) and key pools (Figure 8.18).
# + tags=["remove_cell"]
fig_8_17, axes = plt.subplots(nrows=1, ncols=2, figsize=(10, 5))
(ax1, ax2) = axes.flatten()
label = "{0} [mM/min]"
plot_phase_portrait(
flux_sol, x="use", y="form", ax=ax1,
xlim=(4, 21), ylim=(4, 21),
xlabel=label.format("use"), ylabel=label.format("form"),
title=("(a) Phase Portrait of ATP use vs. formation", XL_FONT),
annotate_time_points=[0, 1e-1, 1e0, 25, 150],
annotate_time_points_color=["red"],
annotate_time_points_labels=True);
# Annotate plot
line_data = [i for i in range(0, 22)]
ax1.plot(line_data, line_data, ls="--", color="black");
ax1.annotate("use < form", xy=(6, 15));
ax1.annotate("use > form", xy=(15, 6));
ax1.annotate("Steady-state line:\n use=form", xy=(15, 19));
ax1.annotate("initial perturbation", xy=(9.5, 9), xycoords="data");
ax1.annotate("", xy=(flux_sol["use"][0], flux_sol["form"][0]),
xytext=(10, 10), textcoords="data",
arrowprops=dict(arrowstyle="->",connectionstyle="arc3"));
plot_phase_portrait(
flux_sol, x="use", y="drain", ax=ax2,
xlim=(0, 21), ylim=(0, 0.1),
xlabel=label.format("use"), ylabel=label.format("drain"),
title=("(b) Phase Portrait of use vs. drain", XL_FONT),
annotate_time_points=[0, 1e-1, 1e0, 25, 150],
annotate_time_points_color=["red"],
annotate_time_points_labels=True);
# Annotate plot
ax2.plot(line_data, [0.03]*22, ls="--", color="black");
ax2.annotate("net AMP\ngain", xy=(1.5, 0.02));
ax2.annotate("net AMP\ndrain", xy=(1.5, 0.04));
fig_8_17.tight_layout()
# -
# **Figure 8.17:** Dynamic phase portraits of fluxes for the simulation of the adenosine phosphate system with formation and drain of AMP (Eq. (8.16)). (a) the ATP use $(v_{\mathrm{use}})$ versus the ATP formation rate $(v_{\mathrm{form}})$. (b) the ATP use $(v_{\mathrm{use}})$ versus the AMP drain $(v_{\mathrm{drain}})$.
# + tags=["remove_cell"]
fig_8_18 = plt.figure(figsize=(12, 4))
gs = fig_8_18.add_gridspec(nrows=1, ncols=2, width_ratios=[1, 1.5])
ax1 = fig_8_18.add_subplot(gs[0, 0])
ax2 = fig_8_18.add_subplot(gs[0, 1])
plot_phase_portrait(
conc_sol, x="Occupancy", y="Capacity", ax=ax1,
time_vector=np.linspace(t0, 10, int(1e6)),
xlim=(2.7, 4.3), ylim=(2.7, 4.3),
xlabel="Occupancy", ylabel="Capacity",
title=("(a) Occupancy vs. Capacity", XL_FONT),
annotate_time_points="endpoints",
annotate_time_points_labels=True);
plot_time_profile(
conc_sol, observable=["EC", "BC", "Overall_Charge"], ax=ax2,
legend=(["E.C.", "B.C.", "Overall Charge"], "right outside"),
time_vector=np.linspace(t0, 10, int(1e6)),
xlabel="Time [min]", ylabel="Charge",
xlim=(t0, 10), ylim=(0.65, 1),
title=("(b) Charge Responses", XL_FONT));
fig_8_18.tight_layout()
# -
# **Figure 8.18:** The Energy Charge response. (a) Dynamic phase portrait of 2ATP+ADP versus 2(ATP+ADP+AMP). (b) The response of E.C., B.C., and overall charge.
#
# * First, we examine how the system balances the use of ATP $(v_{\mathrm{use}})$ with its rate of formation $(v_{\mathrm{form}})$, see Figure 8.17. At $t=0$ the system is at rest at $v_{\mathrm{use}}=v_{\mathrm{form}}=10.0\ mM/min$. Then the system is perturbed by moving the ATP drain, $v_{\mathrm{use}}$, to 15.0 mM/min, as before. The initial response is to increase the formation rate of ATP to about 13 mM/min with the simultaneous drop in the use rate to about the same number, due to a net drop in the concentration of ATP during this period. The rate of ATP use and formation is approximately the same at this point in time. Then, during the slower response time, the use and formation rates of ATP are similar and the system moves along the 45 degree line to a new steady state point at 6.67 mM/min.
#
#
# * The slow dynamics are associated with the inventory of the adenosine phosphates (ATP + ADP + AMP). The AMP drain can be graphed versus the ATP use, see Figure 8.17b. Initially, the AMP drain increases rapidly as the increased ATP use leads to ADP buildup that gets converted into AMP by adenylate kinase $(v_{\mathrm{distr}})$. The AMP drain then drops and sets at the same rate to balance the formation rate, set at 0.03 mM/min.
#
#
# * We can graph the occupancy against the capacity (Figure 8.18a). During the initial response, the occupancy moves while the capacity is a constant. Then, during the slower phase, the two move at a constant ratio. This gives a bi-phasic response of the energy charge (Figure 8.18b). In about a minute, the energy charge changes from 0.86 to about 0.77 and then stays a constant. The energy charge is roughly a constant even though all the other concentrations are changing.
#
#
# This feature of keeping the energy charge a constant while the capacity is changing has a role in a variety of physiological responses, from blood storage to the ischemic response in the heart. Note that this property is a stoichiometric one; no regulation is required to produce this effect.
# ## Charging Substrates and Recovery of High-Energy Bonds
# ### Reaction mechanism:
# As discussed in Section 8.2, most catabolic pathways generate energy (and other metabolic resources) in the form of activated (or charged) carrier molecules. Before energy can be extracted from a compound, it is typically activated by the use of metabolic resources (a biological equivalent of "it takes money to make money"). This basic structure shown in Figure 2.5 is redrawn in Figure 8.19a where one ATP molecule is used to 'charge' a substrate $(x_1)$ with one high-energy bond to form an intermediate $(x_2)$. This intermediate is then degraded through a process wherein two ATP molecules are synthesized and an inorganic phosphate is incorporated. The net gain of ATP is 1 for every $(x_2)$ metabolized, and this ATP molecule can then be used to drive a process $v_{\mathrm{load}}$ that uses an ATP molecule. The trafficking of high-energy phosphate bonds is shown pictorially in Figure 8.19b.
#
# 
#
# **Figure 8.19:** Coupling of the adenosine phosphates with a skeleton metabolic pathway. (a) The reaction map. (b) A pictorial view of the molecules emphasizing the exchange of the high-energy phosphate group (solid circle). The blue square is AMP. The rate laws used are: $b_1 = 0.03\ mM/min.$; $b_2 = 5\ mM/min.$; $k_{\mathrm{drain}}=b_1/0.1$; $k_{\mathrm{load}}=5/1.6$; $k_1=5/0.4$. The flux of $b_2$ was set to 5 mM/min, as the ATP production rate is double that number, thus the steady state value for ATP production is 10 mM/min, to match what is discussed in section 8.3.
# +
# Create model
phos_recovery = MassModel("Phosphate_Recovery", array_type="dense",
dtype=np.int64)
# Define metabolites
atp = MassMetabolite("atp")
adp = MassMetabolite("adp")
amp = MassMetabolite("amp")
pi = MassMetabolite("pi")
x1 = MassMetabolite("x1")
x2 = MassMetabolite("x2")
x3 = MassMetabolite("x3")
# Define reactions
b1 = MassReaction("b1", reversible=False)
b1.add_metabolites({amp:1})
distr = MassReaction("distr")
distr.add_metabolites({adp: -2, amp: 1, atp:1})
load = MassReaction("load", reversible=False)
load.add_metabolites({atp: -1, adp: 1, pi: 1})
drain = MassReaction("drain", reversible=False)
drain.add_metabolites({amp:-1})
b2 = MassReaction("b2", reversible=False)
b2.add_metabolites({x1: 1})
v1 = MassReaction("v1", reversible=False)
v1.add_metabolites({atp: -1, x1: -1, adp: 1, x2: 1})
v2 = MassReaction("v2", reversible=False)
v2.add_metabolites({adp: -2, pi: -1, x2: -1, atp: 2, x3: 1})
DM_x3 = MassReaction("DM_x3", reversible=False)
DM_x3.add_metabolites({x3: -1})
# Add reactions to model
phos_recovery.add_reactions([b1, distr, load, drain, b2, v1, v2, DM_x3])
# Define initial conditions and parameters
atp.ic = 1.6
adp.ic = 0.4
amp.ic = 0.1
pi.ic = 2.5
x1.ic = 1
x2.ic = 1
x3.ic = 1
b1.kf = 0.03
distr.kf = 1000
distr.Keq = 1
load.kf = 5/1.6
drain.kf = 0.3
b2.kf = 5
v1.kf = 5/1.6
v2.kf = 5/0.4
DM_x3.kf = 5
# Set custom rate for source reactions
phos_recovery.add_custom_rate(b1, custom_rate=b1.kf_str)
phos_recovery.add_custom_rate(b2, custom_rate=b2.kf_str)
# -
# #### The dynamic mass balances:
# The dynamic mass balance equations that describe this process are:
#
# $$\begin{align} \frac{dx_1}{dt} &= b_2 - v_1 \\ \frac{dx_2}{dt} &= v_1 - v_2 \\ \frac{d\text{ATP}}{dt} &= -(v_1 + v_{\mathrm{load}}) + 2v_2 + v_{\mathrm{distr}} \\ \frac{d\text{ADP}}{dt} &= (v_1 + v_{\mathrm{load}}) - 2v_2 - 2v_{\mathrm{distr}} \\ \frac{d\text{AMP}}{dt} &= b_1 - v_{\mathrm{drain}} + v_{\mathrm{distr}} \\ \end{align} \tag{8.17}$$
#
# To integrate the reaction schema in Figure 8.13a with this skeleton pathway, we have replaced the use rate of ATP $(v_{\mathrm{use}})$ with $v_1 + v_{\mathrm{load}}$ and the formation rate of ATP $(v_{\mathrm{form}})$ with $2v_2$.
#
# #### Dynamic simulation:
# The flow of substrate into the cell, given by $b_2$, will be set to 5 mM/min in the simulation to follow to set the gross ATP production at 10 mM/min. The response of this system can be simulated to a change in the ATP load parameter, as in previous examples. The difference from the previous examples here is that the net ATP production rate is 5 mM/min.
#
# The time response of the concentrations and fluxes are shown in Figure 8.20, the flux phase portraits in Figure 8.21, and the pools and ratios in Figure 8.22.
# + tags=["remove_cell"]
t0, tf = (0, 100)
sim = Simulation(phos_recovery, verbose=True)
sim.find_steady_state(models=phos_recovery, strategy="simulate",
update_values=True)
conc_sol, flux_sol = sim.simulate(
phos_recovery, time=(t0, tf, tf*10 + 1),
perturbations={"kf_load": "kf_load * 1.5"},
interpolate=True)
pools = {"Occupancy": "adp + 2*atp",
"Capacity": "2*(atp + adp + amp)",
"EC": "(2*atp + adp) / (2*(atp + adp + amp))"}
for pool_id, equation_str in pools.items():
conc_sol.make_aggregate_solution(
pool_id, equation=equation_str, update=True)
netfluxes = {
"load_total": "v1 + load",
"generation": "2*v2",
"drain_total": "drain"}
for flux_id, equation_str in netfluxes.items():
# Skip buffered charge for model with no buffer
flux_sol.make_aggregate_solution(
flux_id, equation=equation_str, update=True)
# + tags=["remove_cell"]
fig_8_20, axes = plt.subplots(nrows=3, ncols=1, figsize=(8, 6))
(ax1, ax2, ax3) = axes.flatten()
plot_time_profile(
conc_sol, observable=phos_recovery.metabolites,
ax=ax1, legend="right outside",
xlim=(t0, 25), ylim=(0, 2.0),
xlabel="Time [min]", ylabel="Concentration [mM]",
title=("(a) Concentrations", XL_FONT));
plot_time_profile(
flux_sol, observable=["v1", "v2", "load"],
ax=ax2, legend="right outside",
xlim=(t0, 25), ylim=(4, 8),
xlabel="Time [min]", ylabel="Fluxes [mM/min]",
title=("(b) High-Flux Reactions", XL_FONT));
plot_time_profile(
flux_sol, observable=["distr", "drain"],
ax=ax3, legend="right outside",
xlim=(t0, 25), ylim=(0, .4),
xlabel="Time [min]", ylabel="Fluxes [mM/min]",
title=("(c) Low-Flux Reactions", XL_FONT));
fig_8_20.tight_layout()
# -
# **Figure 8.20:** The response of the systems shown in Figure 8.19 to a 50% increase in the ATP load rate constant. (a) Dynamic response of the concentrations on a fast and slow time scale. (b) Dynamic response of the main fluxes on a fast and slow time scale. (c) Dynamic response of the AMP determining fluxes on a fast and slow time scale. Parameter values are the same as in Figure 8.19.
# + tags=["remove_cell"]
fig_8_21, axes = plt.subplots(nrows=1, ncols=2, figsize=(10, 5))
(ax1, ax2) = axes.flatten()
plot_phase_portrait(
flux_sol, x="load_total", y="generation", ax=ax1,
xlabel="ATP load total", ylabel="ATP Synthesis",
xlim=(9, 13.5), ylim=(9, 13.5),
title=("(a) ATP Load vs. Synthesis", XL_FONT),
annotate_time_points="endpoints",
annotate_time_points_labels=True);
# Annotate plot
line_data = [i for i in range(8, 15)]
ax1.plot(line_data, line_data, ls="--", color="black");
ax1.annotate(
"", xy=(flux_sol["load_total"](0), flux_sol["generation"](0)),
xytext=(10, 10), textcoords="data",
arrowprops=dict(arrowstyle="->",connectionstyle="arc3"));
ax1.annotate("initial perturbation", xy=(
flux_sol["load_total"](0) - 1.7,
flux_sol["generation"](0) - 0.2));
plot_phase_portrait(
flux_sol, x="load_total", y="drain_total", ax=ax2,
xlabel="ATP load total", ylabel="AMP drain",
xlim=(8, 13.5), ylim=(0, 0.125),
title=("(a) ATP Load vs. Drain", XL_FONT),
annotate_time_points="endpoints",
annotate_time_points_labels=True);
ax2.plot(line_data, [0.03] * 7, ls="--", color="black");
fig_8_21.tight_layout()
# -
# **Figure 8.21:** The response of the system shown in Figure 8.19 to a change in the ATP load rate constant. (a) ATP load versus ATP synthesis rate. (b) ATP load versus AMP drainage rate. You can compare this response to Figure 8.17.
# + tags=["remove_cell"]
fig_8_22 = plt.figure(figsize=(10, 4))
gs = fig_8_22.add_gridspec(nrows=1, ncols=2, width_ratios=[1, 1.5])
ax1 = fig_8_22.add_subplot(gs[0, 0])
ax2 = fig_8_22.add_subplot(gs[0, 1])
plot_phase_portrait(
conc_sol, x="Occupancy", y="Capacity", ax=ax1,
xlim=(2.3, 4.4), ylim=(2.3, 4.4),
xlabel="Occupancy", ylabel="Capacity",
title=("(a) Occupancy vs. Capacity", XL_FONT),
annotate_time_points=[t0, 1e0, 50],
annotate_time_points_color=["red"],
annotate_time_points_labels=True);
# Annotate plot
ax1.annotate(" fast\nmotion\n", xy=(conc_sol["Occupancy"](0.3) - .25,
conc_sol["Capacity"](0.3) - .35))
plot_time_profile(
conc_sol, observable=["EC"], ax=ax2, legend="best",
xlim=(t0, 50), ylim=(0.65, 1),
xlabel="Time [min]", ylabel="Energy Charge",
title=("(b) Stoichiometric Disturbance Rejection Property", XL_FONT));
fig_8_22.tight_layout()
# -
# **Figure 8.22:** The response of the system shown in Figure 8.19 to a change in the ATP load rate constant. (a) Dynamic phase portrait of the pools 2ATP+ADP versus 2(ATP+ADP+AMP). (b) Energy charge ratio as a function of time. You can compare this response to Figure 8.18.
#
# ### Interpretation:
# We can make the following observations from this dynamic response:
#
# * The concentrations move on two principal time scales (Figure 8.20): a fast time scale that is about three to five minutes, and a slower time scale that is about 50 min. ATP and $x_1$ move primarily on the fast time scale, whereas ADP, AMP, and $x_2$ move on the slower time scale. You can see this clearly by changing time in Figure 8.20.
#
# * Initially $v_{\mathrm{load}}$ increases sharply, and $v_2$ increases and $v_1$ decreases to meet the increased load. The three high flux reactions $v_1$, $v_2$, and $v_{\mathrm{load}}$ restabilize at about 5 mM/min after about a three to five minute time frame, after which they are closely, but not fully, balanced (Figure 8.20).
#
# * The dynamic phase portrait, Figure 8.21a, shows that the overall ATP use $(v_1 + v_{\mathrm{load}})$ quickly moves to about 12.5 mM/min while the production rate $(2v_2)$ is about 10 mM/min. Following this initial response, the ATP use drops and the ATP synthesis rate increases to move towards the 45 degree line. The 45 degree line is not reached. After 0.1 min, $v_2$ starts to drop and the system moves somewhat parallel to the 45 degree line until 1.5 min have passed. At this time the ATP concentration has dropped to about 1.06 mM, which makes the ATP use and production rate approximately balanced. Following this point, both the use and production rate increase slowly and return the system back to the initial point where both have a value of 10 mM/min. Since the input rate of $x_1$ is a constant, the system has to return to the initial state.
#
# * AMP initially increases leading to a net drain of AMP from the system. This drain unfolds on a long time scale leading to a net flux through the adenylate kinase that decays on the slower time scale. The effects of AMP drainage can be seen in the flux phase portrait in Figure 8.21b. Initially the AMP drain increases as the ATP usage drops close to its eventual steady state. Then the vertical motion in the phase portrait shows that there is a slower motion in which the ATP usage does not change much but the AMP drainage rate drops to match its input rate at 0.03 mM/hr.
#
# * The dynamic response of the energy charge (Figure 8.22b) shows that it drops on the faster time scale from an initial value of 0.86 to reach a minimum of about 0.67 at about 1.5 min. This initial response results from the increase in the ATP load parameter of 50%. After this initial response, the energy charge increases on the slower time scale to an eventual value of about 0.82.
#
# * Notice that this secondary response is not a result of a regulatory mechanism, but is a property that is built into the stoichiometric structure and the values of the rate constants that lead to the time scale separation.
# ## Summary
#
# * Most biochemical reactions are bilinear. Six of the seven categories of enzymes catalyze bilinear reactions.
#
# * The bilinear properties of biochemical reactions lead to complex patterns of exchange of key chemical moieties and properties. Many such simultaneous exchange processes lead to a 'tangle of cycles' in biochemical reaction networks.
#
# * Skeleton (or scaffold) dynamic models of biochemical processes can be carried out using dynamic mass balances based on elementary reaction representations and mass action kinetics.
#
# * Complex kinetic models are built in a bottom-up fashion, adding more details in a step-wise fashion, making sure that every new feature is consistently integrated. This chapter demonstrated a four-step analysis of the ATP cofactor sub-network and then its integration to a skeleton ATP generating pathway.
#
# * Once dynamic network models are formulated, the perturbations to which we simulate their responses are in fluxes, typically the exchange and demand fluxes.
#
# * A recurring theme is the formation of pools and the state of those pools in terms of how their total concentration is distributed among its constituent members.
#
# * Some dynamic properties are a result of the stoichiometric structure and do not result from intricate regulatory mechanisms or complex kinetic expressions.
# $\tiny{\text{© <NAME> 2011;}\ \text{This publication is in copyright.}\\ \text{Subject to statutory exception and to the provisions of relevant collective licensing agreements,}\\ \text{no reproduction of any part may take place without the written permission of Cambridge University Press.}}$
| docs/education/sb2/chapters/sb2_chapter8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# basic functions and subject lists
# Read subject list, age list, and Jacobian Deter. of 3 study groups by sub_list
import nipype.interfaces.io as nio
import os
# read data: 3 study groups by sub_list
GROUPS=['PD','ET','NC']
OUT_DIR='/output/PD_ICA/'
SUB_LIST=[]; AGE_LIST=[]; JCOB_LIST=[];
print('Local data: ')
for group_name in GROUPS:
current_group=group_name
current_sub_list_file = '/codes/devel/PD_Marker/'+current_group+'_info_ICA.list'
# create dir for output
current_OUT_DIR=OUT_DIR+current_group+'/'
if not os.path.exists(current_OUT_DIR):
os.makedirs(current_OUT_DIR)
#read sub list
with open(current_sub_list_file, 'r') as f_sub:
sub_list_raw= f_sub.readlines()
sub_list = [x[0:-1].split('\t')[0] for x in sub_list_raw] # remove
age_list = [int(x[0:-1].split('\t')[1]) for x in sub_list_raw]
SUB_LIST.append(sub_list); AGE_LIST.append(age_list);
N_sub=len(sub_list)
print(group_name, ': ', N_sub)
# grab group Jacobians
ds_jacobian = nio.DataGrabber(infields=['sub_id'])
ds_jacobian.inputs.base_directory = current_OUT_DIR # database
ds_jacobian.inputs.template = '%s_desc-preproc_T1w_space-MNI2009c_Warp_Jacobian-masked.nii.gz' # from cwd
ds_jacobian.inputs.sort_filelist = True
ds_jacobian.inputs.sub_id = sub_list
res_jacobian = ds_jacobian.run()
jacobian_list=res_jacobian.outputs.outfiles
JCOB_LIST.append(jacobian_list)
pd_sub_list = SUB_LIST[0]; et_sub_list = SUB_LIST[1]; nc_sub_list = SUB_LIST[2];
pd_age_list = AGE_LIST[0]; et_age_list = AGE_LIST[1]; nc_age_list = AGE_LIST[2];
pd_jaco_list=JCOB_LIST[0]; et_jaco_list=JCOB_LIST[1]; nc_jaco_list=JCOB_LIST[2];
from nipype.interfaces.ants import ANTS
import os,time
OUT_DIR='/output/PD_ICA/'
if not os.path.exists(OUT_DIR):
os.makedirs(OUT_DIR)
atlas_09_masked='/templateflow/tpl-MNI152NLin2009cAsym/tpl-MNI152NLin2009cAsym_res-01_desc-brain_T1w.nii.gz'
# read group subject images given template
def dataGraber_sub(SUB_ID, DATA_DIR, TMPT_STR):
import nipype.interfaces.io as nio
import time
t0=time.time()
print('Grabbing files for: ', SUB_ID)
OUT_FILE=[]
out_len=len(TMPT_STR)
if out_len == 0:
print(SUB_ID+' has no files named: ', TMPT_STR)
return OUT_FILE
else:
for i in range(out_len):
TMP='%s/anat/%s_'+TMPT_STR[i]
ds = nio.DataGrabber(infields=['subject_id', 'subject_id'])
ds.inputs.base_directory = DATA_DIR # database
ds.inputs.template = TMP
ds.inputs.subject_id = [SUB_ID]
ds.inputs.sort_filelist = True
res = ds.run()
res_list = res.outputs.outfiles
OUT_FILE.append(res_list)
#print(SUB_ID+' files: ', OUT_FILE)
print('dataGraber takes: ', time.time()-t0 )
return OUT_FILE
# read subject affince and deformation fields from fMRIPrep results
def h5toWarp_nii(H5_FILE, OUT_DIR):
from nipype.interfaces.ants import CompositeTransformUtil
import os
import time
t0=time.time()
tran = CompositeTransformUtil()
tran.inputs.process = 'disassemble'
tran.inputs.in_file = H5_FILE
tran.inputs.out_file = OUT_DIR #bug
#print(tran.cmdline)
res=tran.run()
out_warp =OUT_DIR+'_Warp.nii.gz'
out_affine=OUT_DIR+'_Affine.txt'
os.system('mv '+res.outputs.displacement_field+' '+out_warp )
os.system('mv '+res.outputs.affine_transform+' '+ out_affine)
print('.h5 file disassemble takes: ', time.time()-t0 )
return [out_warp, out_affine]
# Get masked subject Jacobians
def GetJacobian_nii(IN_IMAGE, MASK, JACOB_IMG, JACOB_MASK_IMG):
# Jacobian of deformation field
from nipype.interfaces.ants import CreateJacobianDeterminantImage
from nipype.interfaces import fsl
import time
t0=time.time()
jacobian = CreateJacobianDeterminantImage()
jacobian.inputs.imageDimension = 3
jacobian.inputs.deformationField = IN_IMAGE
jacobian.inputs.outputImage = JACOB_IMG
jacobian.inputs.num_threads = 4
#print(jacobian.cmdline)
jacobian.run()
mask = fsl.ApplyMask(
in_file=JACOB_IMG,
out_file=JACOB_MASK_IMG,
mask_file=MASK)
mask.run()
print('Jacobian takes: ', time.time()-t0 )
return JACOB_MASK_IMG
# -
pd_jaco_list
# +
# MELODIC ICA (local pd+nc Jacobian masked)
from nipype.interfaces import fsl
from nipype.interfaces.ants import ANTS, ApplyTransforms,CreateJacobianDeterminantImage
t0=time.time()
ICA_LOCAL_PATH ='/output/PD_ICA/ICA_local_pd+nc'
if not os.path.exists(ICA_LOCAL_PATH):
os.makedirs(ICA_LOCAL_PATH)
dir_all_jacobian = pd_jaco_list+nc_jaco_list
merged_file = '/output/PD_ICA/local_pd+nc_4d.nii.gz'
#merger = fsl.Merge()
#merger.inputs.in_files = dir_all_jacobian
#merger.inputs.dimension = 'a'
#merger.inputs.merged_file = merged_file
#merger.cmdline
#merger.run()
t1=time.time()
print('Masked Jacobian:\n \tMegring 4D file takes: ', t1-t0)
melodic_setup = fsl.MELODIC()
melodic_setup.inputs.approach = 'tica'
melodic_setup.inputs.in_files = [merged_file]
melodic_setup.inputs.no_bet = True
melodic_setup.inputs.out_all = True
#melodic_setup.inputs.num_ICs = 30
melodic_setup.inputs.out_pca = True
melodic_setup.inputs.out_dir = ICA_LOCAL_PATH
melodic_setup.inputs.report = True
melodic_setup.cmdline
melodic_setup.run()
t2=time.time()
print('\tMELODIC ICA takes: ', t2-t1)
# +
# MELODIC ICA (local et+nc Jacobian masked)
from nipype.interfaces import fsl
from nipype.interfaces.ants import ANTS, ApplyTransforms,CreateJacobianDeterminantImage
t0=time.time()
ICA_LOCAL_PATH ='/output/PD_ICA/ICA_local_et+nc'
if not os.path.exists(ICA_LOCAL_PATH):
os.makedirs(ICA_LOCAL_PATH)
dir_all_jacobian = et_jaco_list+nc_jaco_list
merged_file = '/output/PD_ICA/local_et+nc_4d.nii.gz'
merger = fsl.Merge()
merger.inputs.in_files = dir_all_jacobian
merger.inputs.dimension = 'a'
merger.inputs.merged_file = merged_file
merger.cmdline
merger.run()
t1=time.time()
print('Masked Jacobian:\n \tMegring 4D file takes: ', t1-t0)
melodic_setup = fsl.MELODIC()
melodic_setup.inputs.approach = 'tica'
melodic_setup.inputs.in_files = [merged_file]
melodic_setup.inputs.no_bet = True
melodic_setup.inputs.out_all = True
melodic_setup.inputs.out_pca = True
melodic_setup.inputs.out_dir = ICA_LOCAL_PATH
melodic_setup.inputs.report = True
melodic_setup.cmdline
melodic_setup.run()
t2=time.time()
print('\tMELODIC ICA (ET+NC) takes: ', t2-t1)
# -
| codes/devel/PD_Marker/test_ica_p.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ceciestunepipe
# language: python
# name: ceciestunepipe
# ---
# ## Searching for bouts for a day of ephys recording
# - Microhpone already extracted and left in derived_data as a wav file in sglx_pipe-dev-sort-bouts-s_b1253_21-20210614
#
# +
# %matplotlib inline
import os
import glob
import socket
import logging
import numpy as np
import pandas as pd
from scipy.io import wavfile
from scipy import signal
from ceciestunepipe.util import sglxutil as sglu
from matplotlib import pyplot as plt
from importlib import reload
logger = logging.getLogger()
handler = logging.StreamHandler()
formatter = logging.Formatter(
'%(asctime)s %(name)-12s %(levelname)-8s %(message)s')
handler.setFormatter(formatter)
logger.addHandler(handler)
logger.setLevel(logging.INFO)
logger.info('Running on {}'.format(socket.gethostname()))
# -
from ceciestunepipe.file import bcistructure as et
# ### Get the file locations for a session (day) of recordings
# +
reload(et)
sess_par = {'bird': 'z_r12r13_21',
'sess': '2021-06-26',
'sort': 2}
exp_struct = et.get_exp_struct(sess_par['bird'], sess_par['sess'], ephys_software='sglx')
raw_folder = exp_struct['folders']['sglx']
# -
sess_epochs = et.list_sgl_epochs(sess_par)
sess_epochs
# +
### pick an epoch
reload(et)
reload(sglu)
epoch = sess_epochs[0] # g2 is the shortest
exp_struct = et.sgl_struct(sess_par, epoch)
sgl_folders, sgl_files = sglu.sgl_file_struct(exp_struct['folders']['sglx'])
# -
exp_struct['folders']
# #### search parameters
# Spectrograms are computed using librosa
#
# Additional parameters are for bout search criteria and functions to read the data
# +
# function for getting one channel out of a wave file
def read_wav_chan(wav_path: str, chan_id: int=0) -> tuple:
s_f, x = wavfile.read(wav_path, mmap=True)
if x.ndim==1:
if chan_id > 0:
raise ValueError('Wave file has only one channel, asking for channel {}'.format(chan_id))
x = x.reshape(-1, 1)
return s_f, x[:, chan_id]
def sess_file_id(f_path):
n = int(os.path.split(f_path)[1].split('-')[-1].split('.wav')[0])
return n
hparams = {
# spectrogram
'num_freq':1024, #1024# how many channels to use in a spectrogram #
'preemphasis':0.97,
'frame_shift_ms':5, # step size for fft
'frame_length_ms':10, #128 # frame length for fft FRAME SAMPLES < NUM_FREQ!!!
'min_level_db':-55, # minimum threshold db for computing spe
'ref_level_db':110, # reference db for computing spec
'sample_rate':None, # sample rate of your data
# spectrograms
'mel_filter': False, # should a mel filter be used?
'num_mels':1024, # how many channels to use in the mel-spectrogram
'fmin': 500, # low frequency cutoff for mel filter
'fmax': 12000, # high frequency cutoff for mel filter
# spectrogram inversion
'max_iters':200,
'griffin_lim_iters':20,
'power':1.5,
# Added for the searching
'read_wav_fun': read_wav_chan, # function for loading the wav_like_stream (has to returns fs, ndarray)
'file_order_fun': sess_file_id, # function for extracting the file id within the session
'min_segment': 5, # Minimum length of supra_threshold to consider a 'syllable' (ms)
'min_silence': 1500, # Minmum distance between groups of syllables to consider separate bouts (ms)
'min_bout': 200, # min bout duration (ms)
'peak_thresh_rms': 0.55, # threshold (rms) for peak acceptance,
'thresh_rms': 0.25, # threshold for detection of syllables
'mean_syl_rms_thresh': 0.3, #threshold for acceptance of mean rms across the syllable (relative to rms of the file)
'max_bout': 120000, #exclude bouts too long
'l_p_r_thresh': 100, # threshold for n of len_ms/peaks (typycally about 2-3 syllable spans
'waveform_edges': 1000, #get number of ms before and after the edges of the bout for the waveform sample
'bout_auto_file': 'bout_auto.pickle', # extension for saving the auto found files
'bout_curated_file': 'bout_checked.pickle', #extension for manually curated files (coming soon)
}
# -
# #### Get one wave file
exp_struct['folders']
# +
one_wav_path = os.path.join(exp_struct['folders']['derived'], 'wav_mic.wav')
s_f, x = read_wav_chan(one_wav_path)
hparams['sample_rate'] = s_f
# -
x.shape
plt.plot(x[:50000])
# ### try the function to search for bouts in the file
from ceciestunepipe.util.sound import boutsearch as bs
from joblib import Parallel, delayed
import pickle
# ### get all bouts of the day
reload(bs)
exp_struct['folders']['sglx']
# +
derived_folder = os.path.split(exp_struct['folders']['derived'])[0]
sess_files = et.get_sgl_files_epochs(derived_folder, file_filter='*mic.wav')
sess_files
# -
sess_files
# +
### Force run and save for a single epoch
one_wav_bout_pd = bs.get_bouts_in_long_file(sess_files[1], hparams)
i_folder = os.path.split(sess_files[1])[0]
epoch_bout_pd = one_wav_bout_pd[0]
epoch_bouts_path = os.path.join(i_folder, hparams['bout_auto_file'])
hparams_pickle_path = os.path.join(i_folder, 'bout_search_params.pickle')
with open(hparams_pickle_path, 'wb') as fh:
save_param = hparams.copy()
save_param['read_wav_fun'] = save_param['read_wav_fun'].__name__
save_param['file_order_fun'] = save_param['file_order_fun'].__name__
pickle.dump(save_param, fh)
logger.info('saving bouts pandas to ' + epoch_bouts_path)
epoch_bout_pd.to_pickle(epoch_bouts_path)
# -
plt.plot(one_wav_bout_pd[0]['start_sample'].values)
# +
def get_all_day_bouts(sess_par: dict, hparams:dict, n_jobs: int=28, ephys_software='sglx') -> pd.DataFrame:
logger.info('Will search for bouts through all session {}, {}'.format(sess_par['bird'], sess_par['sess']))
exp_struct = et.get_exp_struct(sess_par['bird'], sess_par['sess'], ephys_software=ephys_software)
# get all the paths to the wav files of the epochs of the day
source_folder = exp_struct['folders']['derived']
wav_path_list = get_files_epochs(source_folder, file_filter='*wav_mic.wav')
wav_path_list.sort()
logger.info('Found {} files'.format(len(wav_path_list)))
print(wav_path_list)
def get_file_bouts(i_path):
epoch_bout_pd = bs.get_bouts_in_long_file(i_path, hparams)[0]
i_folder = os.path.split(i_path)[0]
epoch_bouts_path = os.path.join(i_folder, hparams['bout_auto_file'])
hparams_pickle_path = os.path.join(i_folder, 'bout_search_params.pickle')
logger.info('saving bout detect parameters dict to ' + hparams_pickle_path)
with open(hparams_pickle_path, 'wb') as fh:
save_param = hparams.copy()
save_param['read_wav_fun'] = save_param['read_wav_fun'].__name__
save_param['file_order_fun'] = save_param['file_order_fun'].__name__
pickle.dump(save_param, fh)
logger.info('saving bouts pandas to ' + epoch_bouts_path)
epoch_bout_pd.to_pickle(epoch_bouts_path)
#epoch_bout_pd = pd.DataFrame()
return epoch_bout_pd
# Go parallel through all the paths in the day, get a list of all the pandas dataframes for each file
sess_pd_list = Parallel(n_jobs=n_jobs, verbose=100, backend=None)(delayed(get_file_bouts)(i) for i in wav_path_list)
#sess_pd_list = [get_file_bouts(i) for i in wav_path_list]
#concatenate the file and return it, eventually write to a pickle
sess_bout_pd = pd.concat(sess_pd_list)
return sess_bout_pd
# for large files set n_jobs to 1
sess_bout_pd = get_all_day_bouts(sess_par, hparams, n_jobs=6)
# -
hparams
sess_bout_pd.head()
import sys
sys.stdout.flush()
sess_bout_pd.shape
# + jupyter={"outputs_hidden": true}
sess_bout_pd
# -
# ### save the pandas for the day as a pickle
import pickle
import sys
# +
def save_auto_bouts(sess_bout_pd, sess_par, hparams):
exp_struct = et.get_exp_struct(sess_par['bird'], sess_par['sess'], ephys_software='sglx')
sess_bouts_dir = os.path.join(exp_struct['folders']['derived'], 'bouts_ceciestunepipe')
sess_bouts_path = os.path.join(sess_bouts_dir, hparams['bout_auto_file'])
hparams_pickle_path = os.path.join(sess_bouts_dir, 'bout_search_params.pickle')
os.makedirs(sess_bouts_dir, exist_ok=True)
logger.info('saving bouts pandas to ' + sess_bouts_path)
sess_bout_pd.to_pickle(sess_bouts_path)
logger.info('saving bout detect parameters dict to ' + hparams_pickle_path)
with open(hparams_pickle_path, 'wb') as fh:
pickle.dump(hparams, fh)
save_auto_bouts(sess_bout_pd, sess_par, hparams)
# -
sess_bout_pd.head(1)
# ### save the bouts of a pandas as wavs
bird_bouts_folder = os.path.abspath('/mnt/sphere/speech_bci/processed_data/s_b1253_21/bouts_wav')
bird_bouts_folder
os.makedirs(bird_bouts_folder, exist_ok=True)
a_bout = sess_bout_pd.iloc[0]
a_bout['file']
# +
def bout_to_wav(a_bout: pd.Series, sess_par, hparams, dest_dir):
file_name = '{}_{}_{}.wav'.format(sess_par['sess'],
os.path.split(a_bout['file'])[-1].split('.wav')[0],
a_bout['start_ms'])
file_path = os.path.join(dest_dir, file_name)
x = a_bout['waveform']
wavfile.write(file_path, hparams['sample_rate'], x)
return file_path
def bouts_to_wavs(sess_bout_pd, sess_par, hparams, dest_dir):
# make the dest_dir if does not exist
logger.info('Saving all session bouts to folder ' + dest_dir)
os.makedirs(dest_dir, exist_ok=True)
# write all the motifs to wavs
sess_bout_pd.apply(lambda x: bout_to_wav(x, sess_par, hparams, dest_dir), axis=1)
# write the hparams as pickle
hparams_pickle_path = os.path.join(dest_dir, 'bout_search_params_{}.pickle'.format(sess_par['sess']))
logger.info('saving bout detect parameters dict to ' + hparams_pickle_path)
with open(hparams_pickle_path, 'wb') as fh:
pickle.dump(hparams, fh)
# one example
sess_bouts_folder = os.path.join(bird_bouts_folder, sess_par['sess'])
os.makedirs(sess_bouts_folder, exist_ok=True)
#bout_to_wav(a_bout, sess_par, hparams, bout_folder)
# all the bouts in the sess
bouts_to_wavs(sess_bout_pd, sess_par, hparams, bird_bouts_folder)
# -
# ## Run this for all sessions of the bird with alsa recordings
# #### list all sessions
reload(et)
# + jupyter={"outputs_hidden": true}
all_sessions = et.list_sessions(sess_par['bird'], section='raw', ephys_software='alsa')
all_sessions.sort()
all_sessions
# -
all_sessions[-4:]
for sess in all_sessions[-3:]:
try:
sess_par['sess'] = sess
sess_bout_pd = get_all_day_bouts(sess_par, hparams, n_jobs=28)
save_auto_bouts(sess_bout_pd, sess_par, hparams)
sess_bouts_folder = os.path.join(bird_bouts_folder, sess)
bouts_to_wavs(sess_bout_pd, sess_par, hparams, sess_bouts_folder)
except:
logger.info('Something went wrong in session ' + sess)
logger.info('Error {}'.format(sys.exc_info()[0]))
| notebooks/searchbout_z_r12r13_21-ephys.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="q_Moh17hPano" colab_type="text"
# ### **Teste de performance dos diferentes algoritmos**
# + id="AeVHxt9EU0Z3" colab_type="code" colab={}
import pandas as pd
pd.set_option("display.precision", 2)
c1 = '../content/drive/My Drive/Temp/tp_v01.txt'
c2 = '../content/drive/My Drive/Temp/tp_v02.txt'
c3 = '../content/drive/My Drive/Temp/tp_v02_filtra_dic.txt'
tp1 = pd.read_csv(c1, names=['Palavra','Caracteres','Tempo','Erros'])
tp2 = pd.read_csv(c2, names=['Palavra','Caracteres','Tempo','Erros'])
tp3 = pd.read_csv(c3, names=['Palavra','Caracteres','Tempo','Erros'])
#Converte em milisegundos
tp1['Tempo'] *= 1000
tp2['Tempo'] *= 1000
tp3['Tempo'] *= 1000
# + [markdown] id="jCByLyoTQX7X" colab_type="text"
#
#
# ---
#
#
# + [markdown] id="p7NrVwWPP4NK" colab_type="text"
# ### **Formato do resultado de cada teste**
# + id="nBWxOsWR423A" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="de821ea1-f21f-44bc-ff8c-3187950f93c0"
tp1.head()
# + [markdown] colab_type="text" id="g5jmEIrKQP95"
#
#
# ---
#
#
# + [markdown] colab_type="text" id="3aXbLC3CQl_T"
# ### **Estatísticas de tentativas(letras) erradas**
# + id="9JyNSU3eYwbY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="d9ecbcc4-b502-43a3-ce63-573c40d83015"
erros = pd.concat([tp1['Erros'],tp2['Erros'],tp3['Erros']],axis=1)
erros.columns = ['v0.1','v0.2','v0.2f']
erros.describe()
# + [markdown] colab_type="text" id="nmdH2FCKQxiM"
#
#
# ---
#
#
# + [markdown] colab_type="text" id="T834S3RiQzH8"
# ### **Estatísticas de tempo de solução (ms)**
# + id="7mPVrHHJ26QJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="31f02047-7c41-48e5-9994-e49e7b28448c"
tempo = pd.concat([tp1['Tempo'],tp2['Tempo'],tp3['Tempo']],axis=1)
tempo.columns = ['v0.1','v0.2','v0.2f']
tempo.describe()
# + [markdown] colab_type="text" id="V_2edIEvRENF"
#
#
# ---
#
#
# + [markdown] colab_type="text" id="GOSn0K7kRGgQ"
# ### **Mediana de tentativas erradas por número de caracteres da palavra**
# + id="DoF0KnmpNUD9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 731} outputId="140b1fc3-23e7-4b1a-9807-68da57ad15ab"
e_tp1 = tp1.groupby('Caracteres')['Erros'].median()
e_tp2 = tp2.groupby('Caracteres')['Erros'].median()
e_tp3 = tp3.groupby('Caracteres')['Erros'].median()
e = pd.concat([e_tp1,e_tp2,e_tp3],axis=1)
e.columns = ['v0.1','v0.2','v0.2f']
e
# + [markdown] colab_type="text" id="Vf4MlL9mRmly"
#
#
# ---
#
#
# + [markdown] colab_type="text" id="ROxydGuYRrU0"
# ### **Mediana do tempo de solução por número de caracteres da palavra**
# + id="xCbRvhTPCvXQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 731} outputId="db61cb90-3572-4ece-dc74-f4988dd2e5a9"
t_tp1 = tp1.groupby('Caracteres')['Tempo'].median()
t_tp2 = tp2.groupby('Caracteres')['Tempo'].median()
t_tp3 = tp3.groupby('Caracteres')['Tempo'].median()
t = pd.concat([t_tp1,t_tp2,t_tp3],axis=1)
t.columns = ['v0.1','v0.2','v0.2f']
t
| Teste de Performance/Teste_Performance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Wine Classification using MLP
#
# ## Load the dataset and split it
# +
import numpy as np
import matplotlib.pyplot as plt
from sklearn import datasets
import pandas as pd
data = datasets.load_wine()
# convert sklearn dataset to pandas data frame
wine = pd.DataFrame(data= np.c_[data['data'], data['target']],
columns= data['feature_names'] + ['target'])
# +
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
X = wine.drop('target', axis=1)
y = wine['target']
sc = StandardScaler()
X = sc.fit_transform(X)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)
# -
# ## Create the classifier, train and use it to predict the test set
# +
from sklearn.neural_network import MLPClassifier
mlp = MLPClassifier(hidden_layer_sizes=(13,13,13),max_iter=500)
# -
mlp.fit(X_train,y_train)
predictions = mlp.predict(X_test)
# ## Display some metrics and graphs
from sklearn.metrics import classification_report,confusion_matrix
print(confusion_matrix(y_test,predictions))
print(classification_report(y_test,predictions))
len(mlp.coefs_)
len(mlp.coefs_[0])
# +
import matplotlib.pyplot as plt
plt.plot(range(len(mlp.loss_curve_)), mlp.loss_curve_)
plt.ylabel('Cost')
plt.xlabel('Epochs')
plt.tight_layout()
plt.show()
# +
from sklearn.model_selection import cross_val_score
model = MLPClassifier(hidden_layer_sizes=(100,),max_iter=500)
scores = cross_val_score(model, X, y, cv=10)
print scores
print("Accuracy: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
# +
from sklearn.model_selection import cross_val_score
model = MLPClassifier(hidden_layer_sizes=(50,100,50),max_iter=500)
scores = cross_val_score(model, X, y, cv=10)
print scores
print("Accuracy: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
# -
from sklearn.model_selection import cross_val_score
model = MLPClassifier(hidden_layer_sizes=(13, 13, 13),max_iter=500)
scores = cross_val_score(model, X, y, cv=10)
print scores
print("Accuracy: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
| notebooks/NeuralNetworks/Wine - MLP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from scipy.integrate import solve_ivp
import matplotlib.pyplot as plt
# +
# time = [0,5) in 1000 steps
t = np.linspace(0., 5., 1000)
G0 = 0
I0 = 0
y0 = [G0, I0]
# -
def gluc(t,y):
G, I = y
dG_dt = -2*G - 5*I + 1
dI_dt = + 0.2*G - 0.8*I
return dG_dt, dI_dt
# solve initial value system of ODEs via scipy.integrate.solve_ivp function thorugh Runge-Kutta 23 method
# .y at end extracts results as definition of solve_ivp
G, I = solve_ivp(gluc,(0,5),y0, method='RK23', t_eval=t).y
#simple plot
plt.plot(t, G, 'blue', t, I, 'red');
| Jupyter Notebooks/Code_Fig_25.10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import absolute_import
from __future__ import print_function
import numpy as np
import numpy
import PIL
from PIL import Image
np.random.seed(1337) # for reproducibility
import random
from keras.datasets import mnist
from keras.models import Sequential, Model
from keras.layers import Dense, Dropout, Input, Lambda
from keras.layers.convolutional import Conv2D
from keras.layers.convolutional import MaxPooling2D
from keras.layers import Flatten
from keras.optimizers import RMSprop
from keras import backend as K
def euclidean_distance(vects):
x, y = vects
return K.sqrt(K.sum(K.square(x - y), axis=1, keepdims=True))
def eucl_dist_output_shape(shapes):
shape1, shape2 = shapes
return (shape1[0], 1)
def contrastive_loss(y_true, y_pred):
'''Contrastive loss from Hadsell-et-al.'06
http://yann.lecun.com/exdb/publis/pdf/hadsell-chopra-lecun-06.pdf
'''
margin = 1
return K.mean(y_true * K.square(y_pred) + (1 - y_true) * K.square(K.maximum(margin - y_pred, 0)))
def kullback_leibler_divergence(y_true, y_pred):
y_true = K.clip(y_true, K.epsilon(), 1)
y_pred = K.clip(y_pred, K.epsilon(), 1)
return K.sum(y_true * K.log(y_true / y_pred), axis=-1)
def create_pairs(x, digit_indices):
'''Positive and negative pair creation.
Alternates between positive and negative pairs.
'''
pairs = []
labels = []
n = min([len(digit_indices[d]) for d in range(10)]) - 1
for d in range(10):
for i in range(n):
z1, z2 = digit_indices[d][i], digit_indices[d][i + 1]
pairs += [[x[z1], x[z2]]]
inc = random.randrange(1, 10)
dn = (d + inc) % 10
z1, z2 = digit_indices[d][i], digit_indices[dn][i]
pairs += [[x[z1], x[z2]]]
labels += [1, 0]
return np.array(pairs), np.array(labels)
def create_base_network():
'''Base network to be shared (eq. to feature extraction).
'''
seq = Sequential()
seq.add(Conv2D(30, (5, 5), input_shape=(28, 28,1), activation='relu'))
seq.add(MaxPooling2D(pool_size=(2, 2)))
seq.add(Dropout(0.2))
seq.add(Flatten())
seq.add(Dense(128, activation='relu'))
seq.add(Dropout(0.1))
seq.add(Dense(128, activation='relu'))
return seq
def compute_accuracy(predictions, labels):
'''Compute classification accuracy with a fixed threshold on distances.
'''
return labels[predictions.ravel() < 0.5].mean()
# -
from keras import backend as K
K.epsilon()
seed=7
numpy.random.seed(seed)
# +
(X_train, y_train), (X_test, y_test) = mnist.load_data()
X_train = X_train.reshape(60000, 784)
X_test = X_test.reshape(10000, 784)
X_test = X_test.astype('float32')
# +
X_train=list(X_train)
for i in range(0,10):
for j in range(201,216):
img = PIL.Image.open("/home/aniruddha/Documents/USPSdataset/%d/%d.jpg" %(i,j)).convert("L")
arr = np.array(img)
# ravel to convert 28x28 to 784 1D array
arr=arr.ravel()
X_train.append(arr)
X_train=np.array(X_train)
X_train=X_train.reshape(60150,784)
X_train=X_train.astype('float32')
print(X_train.shape)
# +
#X_train = X_train.reshape(X_train.shape[0], 1, 28, 28).astype('float32')
#X_test = X_test.reshape(X_test.shape[0], 1, 28, 28).astype('float32')
print(X_train.shape)
print(X_test.shape)
# -
X_train=X_train/255
X_test=X_test/255
# +
y_train=list(y_train)
for i in range(0,10):
for j in range(201,216):
y_train.append(i)
y_train=np.array(y_train)
num_classes = 10
print(y_train.shape)
print(y_test.shape)
# -
print(y_train)
input_dim = 784
nb_epoch = 20
# +
# create training+test positive and negative pairs
digit_indices = [np.where(y_train == i)[0] for i in range(10)]
tr_pairs, tr_y = create_pairs(X_train, digit_indices)
digit_indices = [np.where(y_test == i)[0] for i in range(10)]
te_pairs, te_y = create_pairs(X_test, digit_indices)
# -
base_network = create_base_network()
# +
X_temp=X_train.reshape(-1,28,28,1)
input_dim = X_temp.shape[1:]
print(input_dim)
# +
# network definition
base_network = create_base_network()
input_a = Input(shape=input_dim)
input_b = Input(shape=input_dim)
#input_a=K.reshape(input_a,(28,28,1))
#input_b=K.reshape(input_b,(28,28,1))
# because we re-use the same instance `base_network`,
# the weights of the network
# will be shared across the two branches
print(input_b.shape)
# +
processed_a = base_network(input_a)
processed_b = base_network(input_b)
distance = Lambda(euclidean_distance, output_shape=eucl_dist_output_shape)([processed_a, processed_b])
model = Model(input=[input_a, input_b], output=distance)
# -
test_model = Model(input = input_a, output = processed_a)
# +
tr_pairs1=tr_pairs.reshape(-1,2,28,28,1)
te_pairs1=te_pairs.reshape(-1,2,28,28,1)
print(tr_pairs1.shape)
print(te_pairs1.shape)
# -
print(tr_pairs1[:,0].shape)
print(tr_pairs[:, 1])
# +
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
SVG(model_to_dot(model).create(prog='dot', format='svg'))
# +
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
SVG(model_to_dot(base_network).create(prog='dot', format='svg'))
# -
nb_epoch=10
# +
# train
rms = RMSprop()
model.compile(loss=contrastive_loss, optimizer=rms)
history = model.fit([tr_pairs1[:,0], tr_pairs1[:, 1]], tr_y,
validation_data=([te_pairs1[:,0], te_pairs1[:, 1]], te_y),
batch_size=128,
nb_epoch=nb_epoch)
# -
model.save('Siamese+Conv_Train-mnist_150usps_Test-usps.hf')
model.save('Siamese+Conv_Train-mnist_150usps_Test-usps.h5')
print(history.history.keys())
# +
# compute final accuracy on training and test sets
pred = model.predict([tr_pairs1[:,0], tr_pairs1[:, 1]])
tr_acc = compute_accuracy(pred, tr_y)
pred = model.predict([te_pairs1[:,0], te_pairs1[:, 1]])
te_acc = compute_accuracy(pred, te_y)
print('* Accuracy on training set: %0.2f%%' % (100 * tr_acc))
print('* Accuracy on test set: %0.2f%%' % (100 * te_acc))
# -
y_test1=[]
for i in range(0,10):
for j in range(1,201):
y_test1.append(i)
y_test1=np.array(y_test1)
print(y_test1.size)
print(y_test1)
X_test1=[]
for i in range(0,10):
for j in range(1,201):
img = PIL.Image.open("/home/aniruddha/Documents/USPSdataset/%d/%d.jpg" %(i,j)).convert("L")
arr = np.array(img)
# ravel to convert 28x28 to 784 1D array
arr=arr.ravel()
X_test1.append(arr)
X_test1=np.array(X_test1)
print(X_test1.shape)
print(X_test.shape)
# +
X_test1 = X_test1.reshape(2000, 784)
X_test1 = X_test1.astype('float32')
X_test1 /= 255
print(X_test.shape)
digit_indices = [np.where(y_test1 == i)[0] for i in range(10)]
te_pairs, te_y = create_pairs(X_test1, digit_indices)
# +
te_pairs2=te_pairs.reshape(-1,2,28,28,1)
print(te_pairs2.shape)
# +
pred = model.predict([te_pairs2[:, 0], te_pairs2[:, 1]])
te_acc = compute_accuracy(pred, te_y)
# compute accuracy on new USPS dataset to check degree of transfer learning
# model only trained on MNIST dataset
print('* Accuracy on test set: %0.2f%%' % (100 * te_acc))
# +
# %matplotlib inline
from time import time
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import offsetbox
from sklearn import manifold, datasets, decomposition, ensemble, discriminant_analysis, random_projection
# +
def plot_embedding(mu, Y, title=None):
num_class = 1000 # data points per class
# x_min, x_max = np.min(mu, 0), np.max(mu, 0)
# mu = (mu - x_min) / (x_max - x_min)
# classes = [0, 1, 2, 3, 4, 6, 7, 8, 9, 10, 11, 12, 15, 16, 18, 19,
# 20, 21, 22, 25, 26, 27, 28, 29, 30, 31, 32, 34, 35, 36, 37, 39, 40, 42, 43, 44, 45, 46, 48, 49]
classes = [0,1,2,3,4,5,6,7,8,9]
data = [[] for i in classes]
for i, y in enumerate(Y):
data[classes.index(y)].append(np.array(mu[i]))
colors = ['b', 'g', 'r', 'c', 'm', 'y', 'k', 'olive', 'orange', 'mediumpurple']
l = [i for i in range(10)]
alphas = 0.3 * np.ones(10)
fig = plt.figure()
ax = fig.add_subplot(111)
ax.set_aspect(1)
font_size = 13
for i in range(10):
temp = np.array(data[i])
l[i] = plt.scatter(temp[:num_class, 0], temp[:num_class, 1], s = 5, c = colors[i], edgecolors = 'face', alpha=alphas[i])
leg = plt.legend((l[0],l[1],l[2],l[3],l[4],l[5],l[6],l[7],l[8],l[9]),
('0','1','2','3','4','5','6','7','8','9'), loc='center left', bbox_to_anchor=(1, 0.5), fontsize=font_size)
leg.get_frame().set_linewidth(0.0)
plt.xticks(fontsize=font_size)
plt.yticks(fontsize=font_size)
# -
X_test1=X_test1.reshape(-1,28,28,1)
test_model.compile(loss=contrastive_loss, optimizer=rms)
processed=test_model.predict(X_test1)
# +
print("Computing t-SNE embedding")
tsne_pred = manifold.TSNE(n_components=2, init='pca', random_state=0)
t0 = time()
X_tsne_pred = tsne_pred.fit_transform(processed)
plot_embedding(X_tsne_pred, y_test1,
"t-SNE embedding of the digits (time %.2fs)" %
(time() - t0))
# -
X_train11=X_train.reshape(-1,28,28,1)
X_test11=X_test.reshape(-1,28,28,1)
processed_train=test_model.predict(X_train11)
processed_test=test_model.predict(X_test11)
# +
num_pixels = 128
processed_train = processed_train.reshape(processed_train.shape[0], num_pixels).astype('float32')
processed_test = processed_test.reshape(processed_test.shape[0], num_pixels).astype('float32')
print(num_pixels)
print(processed_train.shape)
print(processed_test.shape)
# -
from keras.utils import np_utils
y_train1 = np_utils.to_categorical(y_train)
y_test1 = np_utils.to_categorical(y_test)
num_classes = 10
# define baseline model
def baseline_model1():
# create model
model = Sequential()
model.add(Dense(num_pixels, input_dim=num_pixels, kernel_initializer='normal', activation='relu'))
model.add(Dense(num_classes, kernel_initializer='normal', activation='softmax'))
# Compile model
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
return model
# +
# build the model
model1 = baseline_model1()
# Fit the model
model1.fit(processed_train, y_train1, validation_data=(processed_test, y_test1), epochs=10, batch_size=200, verbose=2)
# Final evaluation of the model
scores_test = model1.evaluate(processed_test, y_test1, verbose=1)
scores_train = model1.evaluate(processed_train, y_train1, verbose=1)
# -
#scores_train=scores_train/1.0
#scores_test=scores_test/1.0
print('* Accuracy on training set: %0.2f%%' % (100 * scores_train[1]))
print('* Accuracy on test set: %0.2f%%' % (100 * scores_test[1]))
y_test2=[]
for i in range(0,10):
for j in range(1,201):
y_test2.append(i)
y_test2=np.array(y_test2)
print(y_test2.size)
from keras.utils import np_utils
y_test2 = np_utils.to_categorical(y_test2)
num_classes = 10
scores_test_USPS=model1.evaluate(processed, y_test2, verbose=1)
print('* Accuracy on test USPS set: %0.2f%%' % (100 * scores_test_USPS[1]))
| Siamese+ConvLayer_Train-MNIST+USPS(150images)_Test-USPS .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="LAqYJKA3tHeA"
# # Assignment \#1
#
# **Due:** Friday, October 16 at 11:59 pm PT
#
# **Objective:**
# This assignment will give you experience using a Google Colab notebook to write text and code, including variables, mathematical operations, and string manipulations.
#
# **Instructions:**
# 1. This version of the assignment cannot be edited. To save an editable version, copy this Colab file to your individual class Google Drive folder ("OCEAN 215 - Autumn '20 - {your name}") by right clicking on this file and selecting "Move to".
# 2. Open the version you copied.
# 3. Complete the assignment by writing and executing text and code cells as specified. **For this assignment, do not use any features of Python that have not yet been discussed in the lessons or class sessions.**
# 4. When you're finished and are ready to submit the assignment, simply save the Colab file ("File" menu –> "Save") before the deadline, close the file, and keep it in your individual class Google Drive folder.
# 5. If you need more time, please see the section "Late work policy" in the syllabus for details.
#
# **Honor code:** In the space below, you can acknowledge and describe any assistance you've received on this assignment, whether that was from an instructor, classmate (either directly or on Piazza), and/or online resources other than official Python documentation websites like docs.python.org or numpy.org. Alternatively, if you prefer, you may acknowledge assistance at the relevant point(s) in your code using a Python comment (#).
# + [markdown] id="GdPsA50lJU_m"
# *Acknowledge assistance here:*
# + [markdown] id="vbwSV0ivx8t8"
# ## Question 1 (8 points)
#
# Would it make the most sense to use a Colab notebook ('A'), Jupyter notebook ('B'), Python script ('C'), or the Python command line ('D') for the following purposes?
#
# To provide your answers, create variables below and assign to them a string with each answer. For instance:
# + id="z_Chvuvzx-Vc"
question_1_part_6 = 'C' # this would indicate you've chosen a Python script for part (6) of the question
# + [markdown] id="zkze1GpFzAf2"
# Scenarios:
# 1. Exploring data you just collected from the ocean while conducting field work aboard a ship.
# 2. Quickly testing out a new Python function or package with a single line of code.
# 3. Writing sturdy code to regularly load and modify batches of data files that are generated in an identical format on a daily basis by a satellite.
# 4. Loading a fairly small data set, working up a preliminary analysis, and sharing the code and results with a teammate.
# + id="mEuH4FndzJA9"
# Provide your answers below:
# + [markdown] id="romb5WJwDINk"
# ## Question 2 (6 points)
#
# Although the goal of OCEAN 215 is not to teach "data science" per se, we hope this course will give you a window into the possibilities offered by data analysis. If you find that you enjoy working with data and writing computer code, you might want to consider a career in academic research or data science. But what is data science, precisely? To better understand this rapidly-growing field, we would like you to read the following article (which should take about 15 minutes), then answer the questions below. There are no wrong answers, but please respond thoughtfully with 1-3 sentences for each (2 points each).
#
# **Article:** "What is data science?" *Thinkful.* https://www.thinkful.com/blog/what-is-data-science/. Read up until the "Your Turn" section.
#
# 1. What data do you think you have generated today that might, at some point, be analyzed by a data scientist?
#
# 2. Can you think of an oceanography data set that could be considered "big data"? Why would it be classified as such?
#
# 3. The article describes how machine learning can be useful for making predictions and identifying relationships in data. Can you think of a research question or task in oceanography or your academic area of interest that might benefit from machine learning? Why might machine learning be particularly useful for that task?
# + [markdown] id="adanpZc9JSLB"
# *Provide your responses here:*
#
#
# + [markdown] id="Sd1TUGbrgwsb"
# ## Question 3 (10 points)
#
# Look at the following table of monthly weather averages in Honolulu, HI. (Source: [Google](https://www.google.com/search?rlz=1C5CHFA_enUS757US757&ei=oO90X-KsFdaU-gTg1paYCw&q=average+monthly+temperature+Honolulu&oq=average+monthly+temperature+Honolulu&gs_lcp=CgZwc3ktYWIQAzICCAAyBAgAEB4yBggAEAgQHjoECAAQRzoECAAQDVDCEVjIFWCoHGgAcAN4AIABPYgBdJIBATKYAQCgAQGqAQdnd3Mtd2l6yAEIwAEB&sclient=psy-ab&ved=0ahUKEwji_KHr4JHsAhVWip4KHWCrBbMQ4dUDCA0&uact=5))
#
# \\
#
# | Month | High T (˚F) | Low T (˚F) |
# |----------|:-------------:|:-------------:|
# | January | 81 | 65 |
# | February | 81 | 65 |
# | March | 82 | 67 |
# | April | 83 | 68 |
# | May | 85 | 69 |
# | June | 86 | 72 |
# | July | 87 | 73 |
# | August | 88 | 73 |
# | September| 88 | 73 |
# | October | 87 | 72 |
# | November | 84 | 70 |
# | December | 82 | 67 |
#
# \\
#
# Answer the following questions using Python code. Please have your code output (print) something for each part. This output should start with the question part – e.g. print('Part 1:',...) – and include the specified lists or calculations and the answers to questions.
#
# 1. Create and print a list containing the high temperatures for each month. What is the object type of the values inside of your list?
# 2. Create and print a new list with the original high temperatures converted into ˚C. If you do not know the conversion equation, feel free to look it up online, but acknowledge the source. What is the object type of the values inside of your new list?
# 3. What part of the conversion triggered this change in object type, and why? Print your answer (1 sentence).
# 4. In your new list from Part 2, replace the temperature in December with a temperature that is 1˚C colder. Print this list.
# 5. Create a list containing the low temperatures for each month.
# 6. Calculate the average low temperature (˚F).
# 7. Sort the low temperature list from low to high.
#
# + id="zohLDb7awjqW"
# Provide your answers below:
# + [markdown] id="L0B9RK4r6Yix"
# ## Question 4 (9 points)
# Challenger Deep, at a depth of 36,200 feet, is known as the deepest part of the ocean (source: [NOAA Ocean Facts](https://oceanservice.noaa.gov/facts/oceandepth.html)). Answer the following questions using Python code. Please have your code output (print) the solutions to each part, including the units! Start your output with the question part, e.g. print('Part 1:',...). If you do not know a conversion factor, feel free to look it up online, but acknowledge the source.
#
# 1. Convert the depth of Challenger Deep to miles.
# 2. Convert the depth of Challenger Deep to km.
# 3. Find the pressure in atm at the bottom of Challenger Deep. (Hint: 1 atm is approximately 10.33 m)
# 4. Convert the pressure to Pa. (Hint: 1 atm is approximately 101325 Pa)
# 5. Convert the pressure to dbar. (Interesting fact: depths in oceanographic data are often specified in dbar instead of meters. Despite pressure not being the same thing as depth, the pressure at 1000 m depth is nearly – but not exactly – equivalent to 1000 dbar. You can confirm this near-equivalence by comparing your answer here to Part 2.)
# 6. If a particle is sinking at a constant rate from the surface to the bottom of the ocean above Challenger Deep at 0.1 mm/s, how many days will it take to reach the bottom?
# + id="Dc9yJ5FX64DU"
# Provide your answers below:
# + [markdown] id="1xFc_ymI_gqB"
# ## Question 5 (10 points)
# Ribosomal RNA (rRNA) gene sequencing is often used in phylogenetic research. Each sequence is made up of combinations of 4 different nucleobases, adenine (A), cytosine (C), guanine (G), and uracil (U). Every species of bacteria and archaea has a slightly-unique version of its 16s rRNA gene. This means that 16s rRNA can be targeted to classify what marine bacteria are present in a given parcel of seawater.
#
# In the code box below, there are two strings containing different DNA gene sequences, which differ from RNA in that they contain thymine (T) nucleobases rather than uracil (U). Answer the following questions using Python. Please have your code output (print) the solutions to each, including any strings you are asked to create or slice. Start your output with the question part, e.g. print('Part 1:',...).
#
# 1. How many nucleobases are in each of the sequences?
# 2. How many times does adenine (A) occur in each of the sequences?
# 3. Do four thymine (T) nucleobases ever occur side-by-side in either of the sequences? Provide your answers as booleans.
# 4. Create and print two new rRNA sequences by replacing the thymine (T) bases with uracil (U) bases in both DNA sequences.
# 5. Slice and print the first 24 nucleobases from the new rRNA sequences. How many of each nucleobase (A,C,G,U) are in each of these shorter sequences?
# + id="NOVKnmep__Bq"
# Keep these starting lines of code – these may not be changed:
DNA1 = 'GGGGGGCAGCAGTGGGGAATATTGGGCAATGGACGAAAGTCTGACCCAGCCATGCCGCGTGTGTGAAGAAGGCTCTAGGGTTGTAATGCACTTTAAGTAGGGAGGAAAGGTTGTGTGTTAATAGCACATAGCTGTGACGTTACCTACAGAATAAGCACCGGCTAACTCCGTGCCAGCAGCCGCGGTAATACGGAGGGTGCAAGCGTTAATCGGAATTACTGGGCGTAAAGCGCGCGTAGGCGGTTATTTAAGCTAGATGTGAAAGCCCAGGGCTCAACCTTGGAATTGCATTTAGAACTGGGTAGCTAGAGTACAAGAGAGGGTGGTGGAATTTCCAGTGTAGCGGTGAAATGCGTAGAGATTGGAAGGAACATCAGTGGCGAAGGCGGCCACCTGGATTGATACTGACGCTGAGGTGCGAAAGCGTGGGGAGCAAACAGGATTAGATACCCCAGTAGTCCT'
DNA2 = 'GGGGCGCAGCAGTGGGGAATATTGCACAATGGGCGAAAGCCTGATGCAGCCATGCCGCGTGTGTGAAGAAGGCCTTCGGGTTGTAAAGCGCTTTCAGTTGTGAGGAAAGGGGTGTAGTTAATAGCTACATCCTGTGACGTTAGCAACAGAAGAAGCACCGGCTAACTTCGTGCCAGCAGCCTCGGTAATACGAGGGGTGCAAGCGTTAATCGGAATTACTGGGCGTAAAGCGTTCGTAGGCGGTTTGTTAAGCAAGATGTGAAAGCCCTGGGCTCAACCTGGGAACTGCATTTTGAACTGGCAAACTAGAGTACTGTAGAGGGTGGTGGAATTTCCAGTGTAGCGGTGAAATGCGTAGAGATTGGAAGGAACATCAGTGGCGAAGGCGGCCACCTGGACAGATACTGACGCTGAGGAACGAAAGCGTGGGGAGCAAACAGGATTAGATACCCTAGTAGTCTG'
# Provide your answers below:
# + [markdown] id="FQQ68HPc_uoo"
# ## Question 6 (7 points)
#
# In the code box below, there is a list of [rorqual whales](https://www.britannica.com/animal/rorqual) containing both their common name, their scientific name as a sub-list separated into genus and species, and their estimated populations (source: [The International Whaling Commission](https://iwc.int/status)). As before, begin your answers with the question part, e.g. print('Part 1:',...).
#
# 1. How many species are in this list?
# 2. Replace each of the inner scientific name sub-lists with a single string containing both the genus and the species separated by a space. Convert the population estimate strings to integers and replace them in the list. Make these changes "in-place," that is, without creating a new list. Print the updated list.
# 3. Remove the information about the Antarctic minke whale from the end of the list and insert it at the beginning of the list. Ensure that the overall format of the list stays the same.
# + id="30akt8mh_25u"
# Keep these starting lines of code – these may not be changed:
rorqual_whales = [['Blue whale',['Balaenoptera','musculus'],'8000'],
['Bryde\'s whale',['Balaenoptera','brydei'],'90000'],
['Common minke whale',['Balaenoptera','acutorostrata'],'200000'],
['Fin whale', ['Balaenoptera','physalus'],'90000'],
['Sei whale', ['Balaenoptera','borealis'],'50000'],
['Antarctic minke whale',['Balaenoptera', 'bonaerensis'],'510000']]
# Provide your answers below:
# + [markdown] id="WNXRVUUy45Xw"
# ## Question 7 (optional, for extra credit: 5 points)
#
# This is a coding puzzle. Write **4 or fewer lines of code** (to follow the 3 lines provided) such that, after executing the code:
# * the variable 'x' will be equal to 44,
# * the variable 'y' will be equal to 4,
# * and the variable 'z' will be equal to 40.
#
# Here's the catch: the code you write may contain mathematical operations (for instance, z = x + y), but **may not contain any numbers** and **may not add a variable to itself** (for instance, z = x + x).
# + id="TwKT6CXczLjf"
# Keep these starting lines of code – these may not be changed:
x = 4
y = 12
z = 'Hello'
# Write your own code below:
| materials/assignments/assignment_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="G0qZfMf4Yreh"
# # Few-shot OpenQA with ColBERT retrieval
# + id="NqUPAnWuYrej"
__author__ = "<NAME> and <NAME>"
__version__ = "CS224u, Stanford, Spring 2022"
# + [markdown] id="R6TRA3gNYrek"
# ## Contents
#
# 1. [Contents](#Contents)
# 1. [Overview](#Overview)
# 1. [Set-up](#Set-up)
# 1. [Google Colab set-up](#Google-Colab-set-up)
# 1. [General set-up](#General-set-up)
# 1. [Language model set-up](#Language-model-set-up)
# 1. [ColBERT set-up](#ColBERT-set-up)
# 1. [Language models](#Language-models)
# 1. [Answerhood](#Answerhood)
# 1. [Eleuther models from Hugging Face](#Eleuther-models-from-Hugging-Face)
# 1. [GPT-3](#GPT-3)
# 1. [SQuAD](#SQuAD)
# 1. [SQuAD dev](#SQuAD-dev)
# 1. [SQuAD dev sample](#SQuAD-dev-sample)
# 1. [SQuAD train](#SQuAD-train)
# 1. [Evaluation](#Evaluation)
# 1. [Open QA with no context](#Open-QA-with-no-context)
# 1. [Few-shot QA](#Few-shot-QA)
# 1. [ColBERT](#ColBERT)
# 1. [ColBERT parameters](#ColBERT-parameters)
# 1. [ColBERT index](#ColBERT-index)
# 1. [Search](#Search)
# 1. [Retrieval evaluation](#Retrieval-evaluation)
# 1. [Zero-shot OpenQA with ColBERT retrieval](#Zero-shot-OpenQA-with-ColBERT-retrieval)
# 1. [Homework questions](#Homework-questions)
# 1. [Few-shot OpenQA with no context [2 points]](#Few-shot-OpenQA-with-no-context-[2-points])
# 1. [Few-shot OpenQA [2 points]](#Few-shot-OpenQA-[2-points])
# 1. [Answer scoring [2 points]](#Answer-scoring-[2-points])
# 1. [Your original system [3 points]](#Your-original-system-[3-points])
# 1. [Bake-off [1 point]](#Bake-off-[1-point])
# + [markdown] id="MXqIujoUYrek"
# ## Overview
#
# The goal of this homework is to explore few-shot (or, prompt-based) learning in the context of open-domain question answering. This is an exciting area that brings together a number of recent task ideas and modeling innovations.
#
# Our core task is __open-domain question answering (OpenQA)__. In this task, all that is given by the dataset is a question text, and the task is to answer that question. By contrast, in modern QA tasks, the dataset provides a text and a gold passage, with a guarantee that the answer will be a substring of the passage.
#
# OpenQA is substantially harder than standard QA. The usual strategy is to use a _retriever_ to find passages in a large collection of texts and train a _reader_ to find answers in those passages. This means we have no guarantee that the retrieved passage will contain the answer we need. If we don't retrieve a passage containing the answer, our reader has no hope of succeeding. Although this is challenging, it is much more realistic and widely applicable than standard QA. After all, with the right retriever, an OpenQA system could be deployed over the entire Web.
#
# The task posed by this homework is harder even than OpenQA. We are calling this task __few-shot OpenQA__. The defining feature of this task is that the reader is simply a general purpose autoregressive language model. It accepts string inputs (prompts) and produces text in response. It is not trained to answer questions per se, and nothing about its structure ensures that it will respond with a substring of the prompt corresponding to anything like an answer.
#
# __Few-shot QA__ (but not OpenQA!) is explored in the famous GPT-3 paper ([Brown et al. 2020](https://arxiv.org/abs/2005.14165)). The authors are able to get traction on the problem using GPT-3, an incredible finding. Our task here – __few-shot OpenQA__ – pushes this even further by retrieving passages to use in the prompt rather than assuming that the gold passage can be used in the prompt. If we can make this work, then it should be a major step towards flexibly and easily deploying QA technologies in new domains.
#
# In summary:
#
# | Task | Passage given | Task-specific reader training |Task-specific retriever training |
# |-----------------:|:-------------:|:-----------------------------:|:--------------------------------:|
# | QA | yes | yes | n/a |
# | OpenQA | no | yes | maybe |
# | Few-shot QA | yes | no | n/a |
# | Few-shot OpenQA | no | no | maybe |
#
# Just to repeat: your mission (should you choose to accept it!) is to explore the final line in this table. The core notebook and assignment don't address the issue of training the retriever in a task-specific way, but we've given some pointers on this in the context of [the original system question at the bottom of this notebook](#Your-original-system-[3-points]).
#
# As usual, this notebook sets up the task and provides starter code. We proceed through a series of approaches:
#
# * _Open QA with no context_: the prompt consists of the question, and we just see what comes back. This is not particularly fair to the system since it doesn't unambiguously convey what we want it to do, but it's a start.
#
# * _Few-shot QA_: the prompt contains one or more examples formatted so as to indirectly convey what we want the system to do, and it uses the gold passage associated with the example. This is the approach of the GPT-3 paper. It works only for datasets with gold passages.
#
# * _Open QA with ColBERT retrieval_: This is roughly as in the previous case, but now we presume no access to a gold passage for our example. Rather, we retrieve a passage from a large corpus using the neural information retrieval model ColBERT.
#
# The above examples are followed by some assignment questions aimed at helping you to think creatively about the problem. These problems improve on the above approaches in various ways.
#
# All of this culminates in an original system question and some code and unlabeled data (here, just a list of questions) for the bake-off.
#
# It is a requirement of the bake-off that a pure autoregressive model be used. In particular, trained QA systems cannot be used at all. See the original system question at the bottom of this message for the full list of allowed models.
#
# Note: the models we are working with here are _big_. This poses a challenge that is increasingly common in NLP: you have to pay one way or another. You can pay to use the GPT-3 API, or you can pay to use an Eleuther model on a heavy-duty cluster computer, or you can pay with time by using an Eleuther model on a more modest computer. If none of these options is palatable, you might consider instead doing the [color reference assignment](hw_colors.ipynb)!
# + [markdown] id="dKB9zXRBYrel"
# ## Set-up
# + [markdown] id="1JwiISlVYrel"
# ### Google Colab set-up
# + [markdown] id="lbnxdvg7Yrem"
# We have sought to make this notebook self-contained so that it can easily be run as a Google Colab. If you are running it in Colab, make sure to select a GPU instance. The notebook will run on a CPU-only instance or CPU-only machine, but it should be much faster with GPU support.
#
# The following are all installed as part of course set-up, but you'll want to run this cell if you are working in a Colab:
# + colab={"base_uri": "https://localhost:8080/"} id="P2TXnN9HYrem" outputId="c87c8b28-c38f-4fbe-f9ae-6cd6dc0d4b44"
# !pip install torch==1.10.0
# !pip install ujson
# !pip install transformers
# !pip install datasets
# !pip install spacy
# !pip install gitpython
# + [markdown] id="hlk-FsfZYrem"
# If you are indeed on a GPU machine, then run the following to ensure complete CUDA support:
# + colab={"base_uri": "https://localhost:8080/"} id="Fl5wBNCxYrem" outputId="aacd2379-f6b7-428b-fec7-2dff54c2e9fa"
import torch
if torch.cuda.is_available():
# !pip install cupy-cuda111
# + [markdown] id="Ej8kZeh6Yren"
# If the above doesn't work, it might be because you don't have CUDA version 11.1. Run
# + id="PojRXsuPYren"
import torch
if torch.cuda.is_available():
# !nvcc --version
# + [markdown] id="VfZEOAL2Yren"
# and then install the corresponding `cupy-cuda`. See [this table](https://docs.cupy.dev/en/stable/install.html#installing-cupy-from-pypi) for details on which one to install for different scenarios.
# + [markdown] id="sFM54iO7Yren"
# ### General set-up
# + id="hL9AAtTzYren"
import collections
from contextlib import nullcontext
from collections import namedtuple
from datasets import load_dataset
import json
import numpy as np
import random
import re
import string
import torch
from typing import List
# + [markdown] id="vNXANb8fYreo"
# Try to set all the seeds for reproducibility (won't extend to GPT-3):
# + id="MIvsYoIpYreo"
seed = 1
np.random.seed(seed)
random.seed(seed)
torch.manual_seed(seed)
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
# + [markdown] id="oKp57bQZYrep"
# The following should install the version of [Faiss](https://github.com/facebookresearch/faiss) that will cooperate with your set-up:
# + colab={"base_uri": "https://localhost:8080/"} id="aGUj1O9wYrep" outputId="ffd634ba-c30c-4d77-c1cf-31533e05a876"
import torch
if torch.cuda.is_available():
# !pip install faiss-gpu==1.7.0
else:
# !pip install faiss-cpu==1.7.0
# + [markdown] id="-mp1C-oyYreq"
# ### Language model set-up
# + [markdown] id="pEUY5P8GYreq"
# To use the GPT-3 API, install the OpenAI library:
# + colab={"base_uri": "https://localhost:8080/", "height": 633} id="4FclKZiuYreq" outputId="53c0e8ca-e5fb-4bb1-b8be-c963d7517b90"
# !pip install openai
# + id="5cbE328hYrer"
import openai
import transformers
from transformers import AutoTokenizer, AutoModelForCausalLM
# + id="TKQEIYGDYrer"
transformers.logging.set_verbosity_error()
# + [markdown] id="axUPfnNmYrer"
# ### ColBERT set-up
#
# Our retriever will be a ColbERT-based model ([Khattab and Zaharia 2020](https://arxiv.org/abs/2004.12832)). ColBERT is a powerful neural information retrieval (Neural IR) model that has proven extremely successful in retrieval applications and as a component in a variety of different systems for OpenQA and other knowledge-intensive tasks (e.g., [Khattab et al. 2021a](https://aclanthology.org/2021.tacl-1.55/); [Khattab et al. 2021b](https://proceedings.neurips.cc/paper/2021/hash/e8b1cbd05f6e6a358a81dee52493dd06-Abstract.html); [Santhanam, Khattab, et al. 2021](https://arxiv.org/abs/2112.01488)).
#
# The following will clone the ColBERTv2 repository for use in this notebook:
# + colab={"base_uri": "https://localhost:8080/"} id="L5wP933JYrer" outputId="1bd1441c-7750-4d22-9519-9f68ff3cce74"
# !git clone -b cpu_inference https://github.com/stanford-futuredata/ColBERT.git
# + colab={"base_uri": "https://localhost:8080/"} id="6LWz4f-3Yres" outputId="11bb2cc3-140e-46d0-d04f-a35f74c97e12"
import os
import sys
sys.path.insert(0, 'ColBERT/')
from colbert.infra import Run, RunConfig, ColBERTConfig
from colbert.data import Collection
from colbert.searcher import Searcher
from utility.utils.dpr import has_answer, DPR_normalize
# + [markdown] id="U5ganiuNYres"
# ## Language models
#
# In few-shot OpenQA, the language model (LM) must read in a prompt and answer the question posed somewhere in the prompt. We propose two basic strategies:
#
# * [EleutherAI](https://www.eleuther.ai/) has released GPT-2-style models in a variety of sizes. These are free to use and easy to use via Hugging Face, and the larger ones very effective for our task, with GPT-J competitive with GPT-3 even though it has only 6B parameters (vs. 145B for GPT-3). The downside here is that the larger models in this family might be very slow and very difficult to work with unless you have access to really impressive GPU hardware. In testing with the free version of Google Colab, we were basically able to do everything we needed to do for the 1.3B parameter model, but the larger one caused too many problems to be viable.
#
# * OpenAI has outstanding API access to GPT-3. You can sign up for [a free account](https://beta.openai.com/signup), and, as of this writing, you get US$18 in credit when you sign up. This is more than enough for the current assignment provided you are careful about how much testing you do. The benefits here are that the API is blazingly fast and requires nothing of your computer in terms of GPU support, and you're getting responses from a 145B parameter model that is truly exceptional.
#
# Our suggestion is to do basic development with `"gpt-neo-125M"`, scale up to `"gpt-neo-1.3B"` once you have a sense for what your original system will be like, and then do your final bake-off entry with GPT-3. The functions `run_eleuther` and `run_gpt3` defined below are totally interchangeable, so this kind of development path should be easy to take.
# + [markdown] id="Fca8-RXjYres"
# ### Answerhood
# + id="U6KepnjTYret"
def _find_generated_answer(tokens, newline="\n" ):
"""Our LMs tend to insert initial newline characters before
they begin generating text. This function ensures that we
properly capture the true first line as the answer while
also ensuring that token probabilities are aligned."""
answer_token_indices = []
char_seen = False
for i, tok in enumerate(tokens):
# This is the main condition: a newline that isn't an initial
# string of newlines:
if tok == newline and char_seen:
break
# Keep the initial newlines for consistency:
elif tok == newline and not char_seen:
answer_token_indices.append(i)
# Proper tokens:
elif tok != newline:
char_seen = True
answer_token_indices.append(i)
return answer_token_indices
# + [markdown] id="klW12GkAYret"
# ### Eleuther models from Hugging Face
# + colab={"base_uri": "https://localhost:8080/", "height": 209, "referenced_widgets": ["19d0a74b2d134747880402c4e6f0fad0", "fcb0f73a2e314a8e935f0802f2f12f31", "9905ea5f04f34c5badc797bd6fa07d3a", "4a08f6612d9d4a3e98d00926e56415b0", "8a1a0c82837e45709424db63bc86ec86", "fb7cce9fecc441b1baf720d2dea6dbfc", "7086129269a04514aa49001d7e712216", "4742b73a39d2444d882aad5b47076911", "9dfca6ee005a4222aa369280c4381a7b", "7583f3ce599641d08a625337921e017f", "<KEY>", "<KEY>", "f208df7109cd475da4f6979e387af2fc", "<KEY>", "6ceb7c5f67e24c1490ff9d209b2e23dd", "<KEY>", "9c5dd297ecb9495aa82ca68bbec66e1c", "<KEY>", "c83a2859c1e941bca4849872a56baa53", "b3990b506419477b824530e11f8a42ef", "446124444abf4c739ad28e593f61562b", "d5a9ed85f6424d329727e3651094d394", "fb95bf53f6ab4ee19b1efeb17c2ea749", "<KEY>", "1a8214cc9451415483ce785b997032e6", "e131d608e26f44f89e1b155fa8042df9", "<KEY>", "83693e21aeae47b0a5b0ee42eb72a3b9", "<KEY>", "0e16502571d94ea286c1fd32d387e5ea", "<KEY>", "6b11d0d12a3c42f88defea028bc27545", "a5246564505c48ee98d7644deed5a255", "<KEY>", "<KEY>", "<KEY>", "c0ab47e3313549cd806f5c10d791d291", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "7efc61ef150b4287841c06cc291722c7", "38a2caefa5cd4722be6f553a1d9f7e8e", "06363edc1de54c15a0a00e104a08d07b", "<KEY>", "eb96d7ee6af840b2af48783ddad1ddb5", "<KEY>", "279a7470d04e4a34b1afdc7d4761c815", "<KEY>", "<KEY>", "40db9d36656c46238c5ed87adf8cb4ec", "<KEY>", "c99ffa80225b4a09ad4e7570b6c4e672", "6c7c0e3d3a844558b12dcdb952e76920", "<KEY>", "a8f1b2d780d74e4382e14974537b1e0b", "<KEY>", "577af7ffedd8430da0870d4a9f773056", "<KEY>", "cdb7e47723fe45e6845a02ad85871d44", "5d67fd950c8e4732a32250c3decb9aef", "e0d37a2cda934241b195138780fbe067", "<KEY>", "23bb67fe91d644469d5be8b442b2e3b6", "264be22c995f407ab22fb4605601f7c3"]} id="1-FkbTaUYret" outputId="f42b7a86-c3bb-4164-92bf-de57520f2bdf"
# "gpt-neo-125M" "gpt-neo-1.3B" "gpt-neo-2.7B" "gpt-j-6B"
eleuther_model_name = "gpt-neo-125M"
eleuther_tokenizer = AutoTokenizer.from_pretrained(
f"EleutherAI/{eleuther_model_name}",
padding_side="left",
padding='longest',
truncation='longest_first', max_length=2000)
eleuther_tokenizer.pad_token = eleuther_tokenizer.eos_token
eleuther_model = AutoModelForCausalLM.from_pretrained(
f"EleutherAI/{eleuther_model_name}")
# + id="rEo9YKwNYret"
def run_eleuther(prompts, temperature=0.1, top_p=0.95, **generate_kwargs):
"""
Parameters
----------
prompts : iterable of str
temperature : float
It seems best to set it low for this task!
top_p : float
For options for `generate_kwargs`, see:
https://huggingface.co/docs/transformers/master/en/main_classes/text_generation#transformers.generation_utils.GenerationMixin.generate
Options that are likely to be especially relevant include
`temperature`, `length_penalty`, and the parameters that
determine the decoding strategy. With `num_return_sequences > 1`,
the default parameters in this function do multinomial sampling.
Returns
-------
list of dicts
{"prompt": str,
"generated_text": str, "generated_tokens": list of str, "generated_probs": list of float,
"answer": str, "answer_tokens": list of str, "answer_probs": list of float
}
"""
prompt_ids = eleuther_tokenizer(
prompts, return_tensors="pt", padding=True).input_ids
with torch.inference_mode():
# Automatic mixed precision if possible.
with torch.cuda.amp.autocast() if torch.cuda.is_available() else nullcontext():
model_output = eleuther_model.generate(
prompt_ids,
temperature=temperature,
do_sample=True,
top_p=top_p,
max_new_tokens=16,
num_return_sequences=1,
pad_token_id=eleuther_tokenizer.eos_token_id,
return_dict_in_generate=True,
output_scores=True,
**generate_kwargs)
# Converting output scores using the helpful recipe here:
# https://discuss.huggingface.co/t/generation-probabilities-how-to-compute-probabilities-of-output-scores-for-gpt2/3175
gen_ids = model_output.sequences[:, prompt_ids.shape[-1] :]
gen_probs = torch.stack(model_output.scores, dim=1).softmax(-1)
gen_probs = torch.gather(gen_probs, 2, gen_ids[:, :, None]).squeeze(-1)
# Generated texts, including the prompts:
gen_texts = eleuther_tokenizer.batch_decode(
model_output.sequences, skip_special_tokens=True)
data = []
iterator = zip(prompts, gen_ids, gen_texts, gen_probs)
for prompt, gen_id, gen_text, gen_prob in iterator:
gen_tokens = eleuther_tokenizer.convert_ids_to_tokens(gen_id)
generated_text = gen_text[len(prompt): ]
gen_prob = [float(x) for x in gen_prob.numpy()] # float for JSON storage
ans_indices = _find_generated_answer(gen_tokens, newline="Ċ")
answer_tokens = [gen_tokens[i] for i in ans_indices]
answer_probs = [gen_prob[i] for i in ans_indices]
answer = "".join(answer_tokens).replace("Ġ", " ").replace("Ċ", "\n")
data.append({
"prompt": prompt,
"generated_text": generated_text,
"generated_tokens": gen_tokens,
"generated_probs": gen_prob,
"generated_answer": answer,
"generated_answer_probs": answer_probs,
"generated_answer_tokens": answer_tokens})
return data
# + id="7fIfuTEGYreu"
eleuther_ex = run_eleuther([
"What year was Stanford University founded?",
"In which year did Stanford first enroll students?"])
eleuther_ex
# + [markdown] id="oiN04KynYreu"
# ### GPT-3
# + id="5t5HTfRkYreu"
def run_gpt3(prompts, engine="text-curie-001", temperature=0.1, top_p=0.95, **gpt3_kwargs):
"""To use this function, sign up for an OpenAI account at
https://beta.openai.com/signup
That should give you $18 in free credits, which is more than enough
for this assignment assuming you are careful with testing.
Once your account is set up, you can get your API key from your
account dashboard and paste it in below as the value of
`openai.api_key`.
Parameters
----------
prompts : iterable of str
engine : str
This has to be one of the models whose name begins with "text".
The "instruct" class of models can't be used, since they seem
to depend on some kinds of QA-relevant supervision.
For options, costs, and other details:
https://beta.openai.com/docs/engines/gpt-3
temperature : float
It seems best to set it low for this task!
top_p : float
For information about values for `gpt3_kwargs`, see
https://beta.openai.com/docs/api-reference/completions
Returns
-------
list of dicts
"""
# Fill this in with the value from your OpenAI account. First
# verify that your account is set up with a spending limit that
# you are comfortable with. If you just opened your account,
# you should have $18 in credit and so won't need to supply any
# payment information.
openai.api_key = None
assert engine.startswith("text"), \
"Please use an engine whose name begins with 'text'."
response = openai.Completion.create(
engine=engine,
prompt=prompts,
temperature=temperature,
top_p=top_p,
echo=False, # This function will not work
logprobs=1, # properly if any of these
n=1, # are changed!
**gpt3_kwargs)
# From here, we parse each example to get the values
# we need:
data = []
for ex, prompt in zip(response["choices"], prompts):
tokens = ex["logprobs"]["tokens"]
logprobs = ex["logprobs"]["token_logprobs"]
probs = list(np.exp(logprobs))
if "<|endoftext|>" in tokens:
end_i = tokens.index("<|endoftext|>")
tokens = tokens[ : end_i] # This leaves off the "<|endoftext|>"
probs = probs[ : end_i] # token -- perhaps dubious.
ans_indices = _find_generated_answer(tokens)
answer_tokens = [tokens[i] for i in ans_indices]
answer_probs = [probs[i] for i in ans_indices]
answer = "".join(answer_tokens)
data.append({
"prompt": prompt,
"generated_text": ex["text"],
"generated_tokens": tokens,
"generated_probs": probs,
"generated_answer": answer,
"generated_answer_tokens": answer_tokens,
"generated_answer_probs": answer_probs})
return data
# + colab={"base_uri": "https://localhost:8080/"} id="y3jbKTAbYrev" outputId="6e560ddc-3762-48ce-8ba6-f8e23c0bda37"
gpt3_ex = run_gpt3([
"What year was Stanford University founded?",
"In which year did Stanford first enroll students?"])
gpt3_ex
# + [markdown] id="XGK3hCs9Yrev"
# ## SQuAD
#
# Our core development dataset is [SQuAD](https://rajpurkar.github.io/SQuAD-explorer/). We chose this dataset because it is well-known and widely used, and it is large enough to support lots of meaningful development work, without, though, being so large as to require lots of compute power. It is also useful that it has gold passages supporting the standard QA formulation, so we can see how well our LM performs with an "oracle" retriever that always retrieves the gold passage.
# + colab={"base_uri": "https://localhost:8080/", "height": 295, "referenced_widgets": ["fedae0a67e9a4f249d9d2ab527d973e2", "280ab82e1c5d4dd8abc4bd15e58034af", "8c59fe3638c344748d6eed8206e8975d", "5f72f99cd4094dc292146e35a4d67767", "35e94aa9d357405780025da79e3691af", "87991a91694f456f9318dc5afccec2c9", "32e27633a7054f17af4dca4ed5e43279", "1acc184682164c08aee48278ae399814", "368e5fc79b9e4ff9b4fe2e512947d677", "72570da75be34d37b32be2dae93222ec", "<KEY>", "bb922ea5e97a44d8b0e94d9f76773167", "32fbedf574754ccc85c4b59d83f743f4", "<KEY>", "b704aad9cc6e474f988e8966ca179da1", "<KEY>", "5c33f1097177468aa401020bac71a5be", "cef06e938024471cabfcfdad4411b961", "<KEY>", "<KEY>", "8639b41657e04b70ac4f3f3ca517a035", "5071e184457e42bf91a84cc5ea827c19", "<KEY>", "5c12675e10044c6db66daf30f4d4f948", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "73cdc7a035d544618e66ed586d7b9649", "<KEY>", "62196b4464bc48a59fa20f1f28f0fe7b", "c28e93ddbc77422484ef2de085794312", "b55b8143e71c4a2cbca7d399fd179797", "<KEY>", "d14af2ecb8664d26b0792eb47791741b", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "4b007c951e474e3db19aa16ca96c16a6", "<KEY>", "<KEY>", "73b8aa0fe175415a9864d017c5f88682", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "a7e04c08f551443481414a0dd619006e", "b40610e2ebb74029bddea0c818532604", "c05c539d546447a5882191f1f1e676a9", "<KEY>", "<KEY>", "<KEY>", "c3a64df8553f4deea779b13a44233546", "ad4e6f09e7b342a59235965668fc34a6", "<KEY>", "872202218d51454faa1211c35a96dc40", "<KEY>", "<KEY>", "<KEY>", "ce46384c19e84fd99447ede08bbaf243", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "f4047dc7f8fe40e3858efacfe3fa9db3", "<KEY>", "<KEY>", "90cb7850489648eab6264dd813719a9c", "2e25e5700521472e83e6888067336682", "ab24a9d6b7a84981ac9d335ff571f7e5", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "d833a94f93074a4085fb2de3583037f5", "<KEY>", "2badb7d424fa46f6b88b3e7acaf6ab48", "42c3232aaad049ff92e95957128e2f2d", "<KEY>", "ed33408a0cad4a46a8366263bdf81bfa", "5969f13e33594392ad1a9245313fd91f", "<KEY>", "d84e03f9117448ef970b930aa77fc968", "<KEY>", "cb5b0f438ff7492e88d9fe4cc9184e9d", "<KEY>", "<KEY>", "e7b24e1fa266446cb6bb61ec067fa412", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "304a8303bd0044e6aa53335b6ab30d5a", "d958f867d5eb40379753a14982302d08", "76b9355cce89436892b959f00b1cf702"]} id="YQ_do58EYrev" outputId="93464cd2-7658-40ac-f411-90cc3b7ba698"
squad = load_dataset("squad")
# + [markdown] id="VbCUz66GYrev"
# The following utility just reads a SQuAD split in as a list of `SquadExample` instances:
# + id="B9-0hkxgYrew"
SquadExample = namedtuple("SquadExample", "id title context question answers")
# + id="g2gt0dkeYrew"
def get_squad_split(squad, split="validation"):
"""
Use `split='train'` for the train split.
Returns
-------
list of SquadExample named tuples with attributes
id, title, context, question, answers
"""
fields = squad[split].features
data = zip(*[squad[split][field] for field in fields])
return [SquadExample(eid, title, context, question, answers["text"])
for eid, title, context, question, answers in data]
# -
# ### SQuAD dev
# + id="NatnOLsDYrew"
squad_dev = get_squad_split(squad)
# + colab={"base_uri": "https://localhost:8080/"} id="jb-ZrSzoYrew" outputId="c3d9b481-2620-434a-cd7b-43c90fc2d096"
squad_dev[0]
# -
# ### SQuAD dev sample
#
# We'll use this fixed but presumably quite random set of examples for exploration and system development:
dev_exs = sorted(squad_dev, key=lambda x: hash(x.id))[: 200]
# ### SQuAD train
#
# To build few-shot prompts, we will often sample SQuAD train examples, so we load that split here:
squad_train = get_squad_split(squad, "train")
# + [markdown] id="DZpXMk-0Yrew"
# ## Evaluation
#
# Our evaluation protocols are the standard ones for SQuAD and related tasks: exact match of the answer (EM) and token-level F1.
#
# We say further that the predicted answer is the first line of generated text after the prompt.
#
# The following evaluation code is taken from the [apple/ml-qrecc](https://github.com/apple/ml-qrecc/blob/main/utils/evaluate_qa.py) repository. It performs very basic string normalization before doing the core comparisons.
# + id="nHHntDSSYrew"
def normalize_answer(s: str) -> str:
"""Lower text and remove punctuation, articles and extra whitespace."""
def remove_articles(text):
regex = re.compile(r'\b(a|an|the)\b', re.UNICODE)
return re.sub(regex, ' ', text)
def white_space_fix(text):
return ' '.join(text.split())
def remove_punc(text):
exclude = set(string.punctuation)
return ''.join(ch for ch in text if ch not in exclude)
def lower(text):
return text.lower()
return white_space_fix(remove_articles(remove_punc(lower(s))))
def get_tokens(s: str) -> List[str]:
"""Normalize string and split string into tokens."""
if not s:
return []
return normalize_answer(s).split()
def compute_exact(a_gold: str, a_pred: str) -> int:
"""Compute the Exact Match score."""
return int(normalize_answer(a_gold) == normalize_answer(a_pred))
def compute_f1_from_tokens(gold_toks: List[str], pred_toks: List[str]) -> float:
"""Compute the F1 score from tokenized gold answer and prediction."""
common = collections.Counter(gold_toks) & collections.Counter(pred_toks)
num_same = sum(common.values())
if len(gold_toks) == 0 or len(pred_toks) == 0:
# If either is no-answer, then F1 is 1 if they agree, 0 otherwise
return int(gold_toks == pred_toks)
if num_same == 0:
return 0
precision = 1.0 * num_same / len(pred_toks)
recall = 1.0 * num_same / len(gold_toks)
f1 = (2 * precision * recall) / (precision + recall)
return f1
def compute_f1(a_gold: str, a_pred: str) -> float:
"""Compute the F1 score."""
gold_toks = get_tokens(a_gold)
pred_toks = get_tokens(a_pred)
return compute_f1_from_tokens(gold_toks, pred_toks)
# + [markdown] id="lJo6C7pgYrex"
# The following is our general evaluation function. We will make extensive use of it to evaluate different systems:
# + id="bJHSxnA6Yrex"
def evaluate(examples, prompts, gens):
"""Generic evalution function.
Parameters
----------
examples: iterable of `SquadExample` instances
prompts: list of str
preds: list of LM-generated texts to evaluate as answers
Returns
-------
dict with keys "em_per", "macro_f1", "examples", where
each "examples" value is a dict
"""
results = []
for ex, prompt, gen in zip(examples, prompts, gens):
answers = ex.answers
pred = gen['generated_answer']
# The result is the highest EM from the available answer strings:
em = max([compute_exact(ans, pred) for ans in answers])
f1 = max([compute_f1(ans, pred) for ans in answers])
gen.update({
"id": ex.id,
"question": ex.question,
"prediction": pred,
"answers": answers,
"em": em,
"f1": f1
})
results.append(gen)
data = {}
data["macro_f1"] = np.mean([d['f1'] for d in results])
data["em_per"] = sum([d['em'] for d in results]) / len(results)
data["examples"] = results
return data
# + [markdown] id="fj170C1PYrex"
# Here is a highly simplified example to help make the logic behind `evaluate` clearer:
# + colab={"base_uri": "https://localhost:8080/"} id="0bgFXLK3Yrex" outputId="e941a6f7-e326-4b26-e91e-8c0ccd11a0a0"
ex = namedtuple("SquadExample", "id title context question answers")
examples = [
ex("0", "CS224u",
"The course to take is NLU!",
"What is the course to take?",
["NLU", "CS224u"])]
prompts = ["Dear model, Please answer this question!\n\nQ: What is the course to take?\n\nA:"]
gens = [{"generated_answer": "NLU", "generated_text": "NLU\nWho am I?"}]
evaluate(examples, prompts, gens)
# + [markdown] id="MHZHve9NYrex"
# The bake-off uses `macro_f1` as the primary metric.
# + [markdown] id="_3LQv2lbYrex"
# ## Open QA with no context
#
# We now have all the pieces we need to begin building few-shot OpenQA systems. Our first system is the simplest and most naive: we simply feed the question text in as the prompt and hope that the model provides an answer as the first line of its generated text.
# + id="swF4V0ngYrex"
def evaluate_no_context(examples, gen_func=run_eleuther, batch_size=20):
prompts = []
gens = []
for i in range(0, len(examples), batch_size):
ps = [ex.question for ex in examples[i: i+batch_size]]
gs = gen_func(ps)
prompts += ps
gens += gs
return evaluate(examples, prompts, gens)
# + colab={"base_uri": "https://localhost:8080/"} id="aPuq5nzDYrex" outputId="cea1edaf-728b-4037-abbc-b58a2d82bfbd"
# %%time
nocontext_results = evaluate_no_context(dev_exs)
print(nocontext_results['macro_f1'])
# + colab={"base_uri": "https://localhost:8080/"} id="lWi1r1-oYrey" outputId="9ffe19d9-ddb1-4a91-9950-06ae521a2a77"
# %%time
nocontext_results_gpt3 = evaluate_no_context(dev_exs, gen_func=run_gpt3)
print(nocontext_results_gpt3['macro_f1'])
# + [markdown] id="csAimDMGYrey"
# ## Few-shot QA
#
# The above formulation is not especially fair to our model, since it doesn't convey anything about the intended structure of the prompt. We want the model to give us an answer to the input question, but we didn't specify that goal unambiguously. Perhaps we were looking for commentary on the question, or a count of the number of tokens it contains, or a passage containing the question string, or something else entirely.
#
# In few-shot QA, we construct a prompt that is intended to convey our intentions more clearly. The first part of the prompt gives some examples of what we want, and the final part provides the set-up for our actual question. In the current formulation, we assume access to the gold passage. For example, if our example of interest is
#
# ```
# Title: CS224u
#
# Background: The course to take is NLU!
#
# Q: What is the course to take?
# ```
#
# with gold answer ```NLU```, then we would create a prompt with, say, 2 additional examples preceding this, to yield a full prompt like this:
#
# ```
# Title: Pragmatics
#
# Background: Pragmatics is the study of language use.
#
# Q: What is pragmatics?
#
# A: The study of language use
#
# Title: Bert
#
# Background: Bert is a Muppet who is lives with Ernie.
#
# Q: Who is Bert?
#
# A: Bert is a Muppet
#
# Title: CS224u
#
# Background: The course to take is NLU!
#
# Q: What is the course to take?
#
# A:
# ```
# This is essentially the formulation used in the GPT-3 paper for SQuAD. The context examples are drawn randomly from the SQuAD train set. We will adopt this same protocol for now. (You might revisit this in the context of your original system.)
# + id="lhsl9yvHYrey"
def build_few_shot_qa_prompt(ex, squad_train, n_context=2, joiner="\n\n"):
segs = []
train_exs = random.sample(squad_train, k=n_context)
for t in train_exs:
segs += [
f"Title: {t.title}",
f"Background: {t.context}",
f"Q: {t.question}",
f"A: {t.answers[0]}"
]
segs += [
f"Title: {ex.title}",
f"Background: {ex.context}",
f"Q: {ex.question}",
f"A:"
]
return joiner.join(segs)
# + [markdown] id="XzLghzI5Yrez"
# Here's the sort of output we get with `n_context=1`:
# + colab={"base_uri": "https://localhost:8080/"} id="VEuVae4xYrez" outputId="1fc52cb8-1fa7-4d84-c911-4e3f2cafd682"
print(build_few_shot_qa_prompt(dev_exs[0], squad_train, n_context=1))
# + id="VUwgM625Yrez"
def evaluate_few_shot_qa(examples, squad_train, gen_func=run_eleuther, batch_size=20, n_context=2):
prompts = []
gens = []
for i in range(0, len(examples), batch_size):
batch = examples[i: i+batch_size]
ps = [build_few_shot_qa_prompt(ex, squad_train, n_context=n_context) for ex in batch]
gs = gen_func(ps)
prompts += ps
gens += gs
return evaluate(examples, prompts, gens)
# + colab={"base_uri": "https://localhost:8080/"} id="UelblRFCYrez" outputId="0587385d-207a-47bb-d363-5b33c3a20920"
# %%time
few_shot_qa_results = evaluate_few_shot_qa(dev_exs, squad_train, n_context=1)
print(few_shot_qa_results['macro_f1'])
# + id="-fdPsEbmYrez" outputId="be6e22d2-7655-4c98-89f9-a0fe81d204d8"
# %%time
few_shot_qa_results_gpt3 = evaluate_few_shot_qa(
dev_exs, squad_train, n_context=1, gen_func=run_gpt3)
print(few_shot_qa_results_gpt3['macro_f1'])
# + [markdown] id="WfvVdsGrYre0"
# ## ColBERT
#
# It's now just a short step to our core task, few-shot OpenQA. We just need to give up our beloved gold passage and instead try to retrieve the right passage or passages from a corpus.
#
# The first step is instantiating the ColBERT retriever and loading in an index. Our ColBERT retriever was initially trained on MS MARCO, and we have pre-indexed a collection of 100K documents that we know to be well-aligned with SQuAD and with the dataset used for the bake-off assessment. (See [the original system question](#Your-original-system-[3-points]) for tips on creating your own index.)
# -
index_home = os.path.join("experiments", "notebook", "indexes")
# + [markdown] id="NFYxJPpuYre0"
# ### ColBERT parameters
# + colab={"base_uri": "https://localhost:8080/"} id="5tnUU2UHYre0" outputId="9d7431a0-7f69-4549-960f-56f8575a8e97"
if not os.path.exists(os.path.join("data", "openqa", "colbertv2.0.tar.gz")):
# !mkdir -p data/openqa
# ColBERTv2 checkpoint trained on MS MARCO Passage Ranking (388MB compressed)
# !wget https://downloads.cs.stanford.edu/nlp/data/colbert/colbertv2/colbertv2.0.tar.gz -P data/openqa/
# !tar -xvzf data/openqa/colbertv2.0.tar.gz -C data/openqa/
# + [markdown] id="cjZ1hZJnYre0"
# If something went wrong with the above, you can just download the file https://downloads.cs.stanford.edu/nlp/data/colbert/colbertv2/colbertv2.0.tar.gz, unarchive it, and move the resulting `colbertv2.0` directory into the `data/openqa` directory.
# + [markdown] id="QzRjL61eYre0"
# ### ColBERT index
# + id="jD0U5Outa9HU"
if not os.path.exists(os.path.join(index_home, "cs224u.collection.2bits.tgz")):
# !wget https://web.stanford.edu/class/cs224u/data/cs224u.collection.2bits.tgz -P experiments/notebook/indexes
# !tar -xvzf experiments/notebook/indexes/cs224u.collection.2bits.tgz -C experiments/notebook/indexes
# + [markdown] id="61ergQcQYre0"
# If something went wrong with the above, download the file https://web.stanford.edu/class/cs224u/data/cs224u.collection.2bits.tgz, unarchive it, and move the resulting `cs224u.collection.2bits` directory into the `experiments/notebook/indexes` directory (which you will probably need to create).
# + colab={"base_uri": "https://localhost:8080/", "height": 71} id="XtEGC6MyYre0" outputId="93164bc8-b9da-4893-8b2f-d5eeca0403a3"
collection = os.path.join(index_home, "cs224u.collection.2bits", "cs224u.collection.tsv")
collection = Collection(path=collection)
f'Loaded {len(collection):,} passages'
# + id="BixmAYizYre0"
index_name = "cs224u.collection.2bits"
# -
# Now we create our `searcher`:
# + colab={"base_uri": "https://localhost:8080/"} id="N6wYpChzYre1" outputId="1eb08be0-a66c-4414-d69b-32733d74631b"
with Run().context(RunConfig(experiment='notebook')):
searcher = Searcher(index=index_name)
# + [markdown] id="dex5KPUTYre1"
# ### Search
#
# Now that the index is loaded, you can do searches over it. The index is limited, but retrieval is very solid!
# + colab={"base_uri": "https://localhost:8080/"} id="-v2jKaR8Yre1" outputId="6ae9e5ee-8a1d-4d00-9925-1dbf47d21412"
query = "linguistics"
print(f"#> {query}")
# Find the top-3 passages for this query
results = searcher.search(query, k=3)
# Print out the top-k retrieved passages
for passage_id, passage_rank, passage_score in zip(*results):
print(f"\t[{passage_rank}]\t{passage_score:.1f}\t {searcher.collection[passage_id]}")
# + [markdown] id="w9n0_xwdYre1"
# ### Retrieval evaluation
#
# For more rigorous evaluations of the retriever alone, we can use Sucess@`k` defined relative to the SQuAD passages and answers. We say that we have a "success" if a passage in the top `k` retrieved passages contains any of the answers substrings, and Sucess@`k` is the percentage of such success cases. This is very heuristic (perhaps the answer string happens to occur somewhere in a completely irrelevant passage), but it can still be good guidance.
# + id="fsNayHzdYre1"
def success_at_k(examples, k=20):
scores = []
for ex in examples:
scores.append(evaluate_retrieval_example(ex, k=5))
return sum(scores) / len(scores)
def evaluate_retrieval_example(ex, k=20):
results = searcher.search(ex.question, k=k)
for passage_id, passage_rank, passage_score in zip(*results):
passage = searcher.collection[passage_id]
score = has_answer([DPR_normalize(ans) for ans in ex.answers], passage)
if score:
return 1
return 0
# + [markdown] id="k2U4v58dYre1"
# Here is Sucess@20 for the SQuAD dev set:
# + colab={"base_uri": "https://localhost:8080/"} id="J2oEmspeYre1" outputId="bfff7ec2-26e4-47ef-a101-04fff7fd0621"
# %%time
if torch.cuda.is_available():
# This will take a few hours on a CPU:
print(success_at_k(squad_dev))
else:
# This should be reasonably fast and yields the
# same kind of result:
print(success_at_k(dev_exs))
# + [markdown] id="teKobQM8Yre1"
# ## Zero-shot OpenQA with ColBERT retrieval
#
# We're now in a position to define a system that does our full few-shot OpenQA task. To get this started, we define just a version that doesn't include any SQuaD-training examples in the prompt. So this is really zero-shot OpenQA. (The homework asks you to move to the true few-shot setting.)
# + id="jbn4BHr4Yre1"
def build_zero_shot_openqa_prompt(question, passage, joiner="\n\n"):
title, context = passage.split(" | ", 1)
segs = [
f"Title: {title}",
f"Background: {context}",
f"Q: {question}",
"A:"
]
return joiner.join(segs)
# + id="Xjz7l_yQYre2"
def evaluate_zero_shot_openqa(examples, joiner="\n\n", gen_func=run_eleuther, batch_size=20):
prompts = []
gens = []
for i in range(0, len(examples), batch_size):
exs = examples[i: i+batch_size]
results = [searcher.search(ex.question, k=1) for ex in exs]
passages = [searcher.collection[r[0][0]] for r in results]
ps = [build_zero_shot_openqa_prompt(ex.question, psg, joiner=joiner)
for ex, psg in zip(exs, passages)]
gs = gen_func(ps)
prompts += ps
gens += gs
return evaluate(examples, prompts, gens)
# + colab={"base_uri": "https://localhost:8080/"} id="21DfS0hHYre2" outputId="b653e3e8-252c-47f8-8481-c190aca6c286"
# %%time
zero_shot_openqa_results = evaluate_zero_shot_openqa(dev_exs)
print(zero_shot_openqa_results['macro_f1'])
# + colab={"base_uri": "https://localhost:8080/"} id="MNhMla71Yre2" outputId="d6a6d52b-4af5-4519-cd7f-2298e859cc79"
# %%time
zero_shot_openqa_results_gpt3 = evaluate_zero_shot_openqa(dev_exs, gen_func=run_gpt3)
zero_shot_openqa_results_gpt3['macro_f1']
# + [markdown] id="outblNTaYre2"
# ## Homework questions
#
# Please embed your homework responses in this notebook, and do not delete any cells from the notebook. (You are free to add as many cells as you like as part of your responses.)
# -
# ### Few-shot OpenQA with no context [2 points]
#
# In the section [Open QA with no context](#Open-QA-with-no-context) above, we simply prompted our LM with a question string and looked at what came back. This is arguably unfair to the LM, since we didn't convey anything about our intentions.
#
# For a fairer assessment of what the LM alone can do, we should move to the few-shot setting by giving the model a few examples of what we have in mind. The idea here is to create prompts that look like this:
#
# ```
# Q: What is pragmatics?
#
# A: The study of language use
#
# Q: Who is Bert?
#
# A: Bert is one of the Muppets.
#
# Q: What was Stanford University founded?
#
# A:
# ```
#
# This question asks you to write a function for creating such prompts, using SQuAD training examples, and a second function for evaluating this approach. The goal is to have a no context baseline for the other few-shot approaches we are considering.
#
# __Task 1___: Complete the function `build_few_shot_no_context_prompt` so that it builds prompts like the above. You can use `test_build_few_shot_no_context_prompt` to check that your function is returning prompts in the desired format.
#
# __Task 2__: Complete the function `evaluate_few_shot_no_context` so that you can evaluate this approach. You can use `test_evaluator` to check that your function is performing the desired kind of evaluation.
# +
def build_few_shot_no_context_prompt(question, train_exs, joiner="\n\n"):
"""No context few-shot OpenQA prompts.
Parameters
----------
question : str
train_exs : iterable of SQuAD train examples. These can be
obtained via a random sample
from `squad_train` as defined above.
joiner : str
The character to use to join pieces of the prompt into
a single str.
Returns
-------
str, the prompt
"""
##### YOUR CODE HERE
# -
def test_build_few_shot_no_context_prompt(func):
train_exs = [
SquadExample(0, "T1", "Q1", "C1", ["A1"]),
SquadExample(1, "T2", "Q2", "C2", ["A2"]),
SquadExample(2, "T3", "Q3", "C3", ["A3"])]
question = "My Q"
result = func(question, train_exs, joiner="\n")
expected = ""
tests = [
(1, "\n", 'Q: C1\nA: A1\nQ: My Q\nA:'),
(1, "\n\n", 'Q: C1\n\nA: A1\n\nQ: My Q\n\nA:'),
(2, "\n", 'Q: C1\nA: A1\nQ: C2\nA: A2\nQ: My Q\nA:')]
err_count = 0
for n_context, joiner, expected in tests:
result = func(question, train_exs[: n_context], joiner=joiner)
if result != expected:
err_count +=1
print(f"Error:\n\nExpected:\n\n{expected}\n\nGot:\n\n{result}")
if err_count == 0:
print("No errors detected in `build_few_shot_no_context_prompt`")
test_build_few_shot_no_context_prompt(build_few_shot_no_context_prompt)
def evaluate_few_shot_no_context(
examples,
squad_train,
batch_size=20,
n_context=2,
joiner="\n\n",
gen_func=run_eleuther):
"""Evaluate a few-shot OpenQA with no context approach
defined by `build_few_shot_no_context_prompt` and `gen_func`.
Parameters
----------
examples : iterable of SQuAD train examples
Presumably a subset of `squad_dev` as defined above.
squad_train : iterable of SQuAD train examples
batch_size : int
Number of examples to send to `gen_func` at once.
joiner : str
Used by `build_few_shot_open_qa_prompt` to join segments
of the prompt into a single str.
gen_func : either `run_eleuther` or `run_gpt3`
Returns
-------
dict as determined by `evaluate` above.
"""
# A list of strings that you build and feed into `gen_func`.
prompts = []
# A list of dicts that you get from `gen_func`.
gens = []
# Iterate through the examples in batches:
for i in range(0, len(examples), batch_size):
# Sample some SQuAD training examples to use with
# `build_few_shot_no_context_prompt` and `ex.question`,
# run the resulting prompt through `gen_func`, and
# add your prompts and results to `prompts` and `gens`.
##### YOUR CODE HERE
# Return value from a call to `evalaute`, with `examples`
# as provided by the user and the `prompts` and `gens`
# you built:
return evaluate(examples, prompts, gens)
def test_evaluator(func):
examples = [SquadExample(0, "T1", "Q1", "C1", ["A1"])]
squad_train = [SquadExample(0, "sT1", "sQ1", "sC1", ["sA1"])]
def gen_func(*prompts):
return [{
"generated_answer": "Constant output",
"generated_answer_tokens": ["Constant", "output"],
"generated_answer_probs": [0.1, 0.2]}]
batch_size = 1
n_context = 1
joiner = "\n"
result = func(
examples,
squad_train,
batch_size=1,
n_context=1,
joiner=joiner,
gen_func=gen_func)
expected_keys = {'em_per', 'examples', 'macro_f1'}
result_keys = set(result.keys())
if expected_keys != result_keys:
print(f"Unexpected keys in result. "
f"Expected: {expected_keys}; Got: {result_keys}")
return
expected_ex_keys = {
'f1', 'id', 'em', 'generated_answer_tokens', 'generated_answer_probs',
'prediction', 'generated_answer', 'question', 'answers'}
result_ex_keys = set(result["examples"][0].keys())
if expected_ex_keys != result_ex_keys:
print(f"Unexpected keys in result['examples']. "
f"Expected: {expected_ex_keys}; Got: {result_ex_keys}")
return
print("No errors detected in `evaluate_few_shot_open_qa`")
test_evaluator(evaluate_few_shot_no_context)
# + [markdown] id="w2mx3Z4HYre2"
# ### Few-shot OpenQA [2 points]
#
# In the section [Few-shot QA](Few-shot-QA) above, we used SQuAD training examples to build prompts that we hope will help the model infer our intended semantics for the prompts themselves. When we moved to the open formulation of the problem, in [Open QA with ColBERT retrieval](Open-QA-with-ColBERT-retrieval), we forced the model to deal with prompts that lack these context clues. This is a "zero-shot" formulation of the problem. The goal of this homework problem is to improve that system so that it truly supports few-shot OpenQA.
#
# __Task 1__: Complete the function `build_few_shot_open_qa_prompt` so that it builds prompts from a question, a passage, and a sample of SQuAD training examples. You can use `test_build_few_shot_open_qa_prompt` to check that your function is returning prompts in the desired format.
#
# __Task 2__: Complete the function `evaluate_few_shot_open_qa` so that you can evaluate this approach. You can use `test_evaluator` from above to check that your function is performing the desired kind of evaluation.
#
# We will be checking only that the tests pass. We will not be evaluating the quality of the results you obtain using this code.
# + id="HUuZ5l3gYre2"
def build_few_shot_open_qa_prompt(question, passage, train_exs, joiner="\n\n"):
"""Few-shot OpenQA prompts.
Parameters
----------
question : str
passage : str
Presumably something retrieved via search.
train_exs : iterable of SQuAD train examples
These can be obtained via a random sample from
`squad_train` as defined above.
joiner : str
The character to use to join pieces of the prompt
into a single str.
Returns
-------
str, the prompt
"""
##### YOUR CODE HERE
# + id="vgeNwTu4Yre2"
def test_build_few_shot_open_qa_prompt(func):
train_exs = [
SquadExample(0, "T1", "Q1", "C1", ["A1"]),
SquadExample(1, "T2", "Q2", "C2", ["A2"]),
SquadExample(2, "T3", "Q3", "C3", ["A3"])]
question = "My Q"
passage = "Title | target passage"
tests = [
(1, "\n", ('Title: T1\nBackground: Q1\nQ: C1\nA: A1\n'
'Title: Title\nBackground: target passage\nQ: My Q\nA:')),
(1, "\n\n", ('Title: T1\n\nBackground: Q1\n\nQ: C1\n\nA: A1\n\n'
'Title: Title\n\nBackground: target passage\n\nQ: My Q\n\nA:')),
(2, "\n", ('Title: T1\nBackground: Q1\nQ: C1\nA: A1\nTitle: T2\n'
'Background: Q2\nQ: C2\nA: A2\nTitle: Title\n'
'Background: target passage\nQ: My Q\nA:'))]
err_count = 0
for n_context, joiner, expected in tests:
result = func(question, passage, train_exs[: n_context], joiner=joiner)
if result != expected:
err_count +=1
print(f"Error:\n\nExpected:\n\n{expected}\n\nGot:\n\n{result}")
if err_count == 0:
print("No errors detected in `build_few_shot_open_qa_prompt`")
# + colab={"base_uri": "https://localhost:8080/"} id="lK991-2AYre3" outputId="b8134fd6-d477-41e4-9d70-cbed90f17a0c"
test_build_few_shot_open_qa_prompt(build_few_shot_open_qa_prompt)
# + id="DRhoMeEGYre3"
def evaluate_few_shot_open_qa(
examples,
squad_train,
batch_size=20,
n_context=2,
joiner="\n\n",
gen_func=run_eleuther):
"""Evaluate a few-shot OpenQA approach defined by
`build_few_shot_open_qa_prompt` and `gen_func`.
Parameters
----------
examples : iterable of SQuAD train examples
Presumably a subset of `squad_dev` as defined above.
squad_train : iterable of SQuAD train examples
batch_size : int
Number of examples to send to `gen_func` at once.
joiner : str
Used by `build_few_shot_open_qa_prompt` to join segments
of the prompt into a single str.
gen_func : either `run_eleuther` or `run_gpt3`
Returns
-------
dict as determined by `evaluate` above.
"""
# A list of strings that you build and feed into `gen_func`.
prompts = []
# A list of dicts that you get from `gen_func`.
gens = []
# Iterate through the examples in batches:
for i in range(0, len(examples), batch_size):
# Use the `searcher` defined above to get passages
# using `ex.question` as the query, and use your
# `build_few_shot_open_qa_prompt` to build prompts.
##### YOUR CODE HERE
# Return value from a call to `evalaute`, with `examples`
# as provided by the user and the `prompts` and `gens`
# you built:
return evaluate(examples, prompts, gens)
# + colab={"base_uri": "https://localhost:8080/"} id="beEyy_eOYre3" outputId="22b49855-289f-43e8-f214-5d18238a00b7"
test_evaluator(evaluate_few_shot_open_qa)
# + [markdown] id="oXeQzplkYre3"
# ### Answer scoring [2 points]
#
# We have so far been assuming that the top-ranked passage retrieved by ColBERT should be used in the prompt and that the single answer returned by the LM is our prediction. It may be possible to improve on this by scoring answers using the ColBERT scores and the probabilities returned by the LM. This question asks you to explore a basic approach to such scoring. The core scoring function:
#
# $$
# \textbf{score}_{\text{prompt-func}}(\textrm{answer}, \textrm{passage}, \textrm{question}) =
# P(\textrm{passage} \mid \textrm{question}) \cdot
# P(\textrm{answer} \mid \text{prompt-func}(\textrm{question}, \textrm{passage}) )
# $$
#
# where we estimate the two conditional probabilities as follows:
#
# * $P(\textrm{passage} \mid \textrm{question})$ is defined only for the top $k$ passages and defined by the softmax of the top $k$ scores returned by the retriever.
#
# * $P(\textrm{answer} \mid \text{prompt-func}(\textrm{question}, \textrm{passage}))$ is simply the product of the per-token probabilities of the generated answer given the prompt determined by $\text{prompt-func}(\textrm{question}, \textrm{passage})$. These values can be extracted from the return values of both `run_eleuther` and `run_gpt3` using the key `"generated_answer_probs"`. (Your prompt function might of course have other arguments not represented here.)
#
# __Your task__: Implement this scoring function for an individual example. The two required pieces are `get_passages_with_scores` and `answer_scoring`. Starter code for each is below, and each has a unit test you can run to check your work.
#
# (With this implemented, it is easy to create a new prediction function that uses the $\textrm{answer}$ from the highest-scoring $\textrm{answer}/\textrm{passage}$ pair as the prediction for input $\textrm{question}$. You are not required to implement such a prediction function, but you might do this as part of [your original system](#Your-original-system-[3-points]).)
# + id="m7PhfMNsYre3"
def get_passages_with_scores(question, k=5):
"""Pseudo-probabilities from the retriever.
Parameters
----------
question : str
k : int
Number of passages to retrieve.
Returns
-------
passages (list of str), passage_probs (np.array)
"""
# Use the `searcher` to get `k` passages for `questions`:
##### YOUR CODE HERE
# Softmax normalize the scores and convert the list to
# a NumPy array:
##### YOUR CODE HERE
# Get the passages as a list of texts:
##### YOUR CODE HERE
# + id="-vqCNOtMYre4"
def test_get_passages_with_scores(func):
question = "What is linguistics?"
passages, passage_probs = get_passages_with_scores(question, k=2)
if len(passages) != len(passage_probs):
print("`get_passages_with_scores` should return equal length "
"lists of passages and passage probabilities.")
return
if len(passages) != 2:
print(f"`get_passages_with_scores` should return `k` passages. Yours returns {len(passages)}")
return
if not all(isinstance(psg, str) for psg in passages):
print("The first return argument should be a list of passage strings.")
return
if not all(isinstance(p, (float, np.float32, np.float64)) for p in passage_probs):
print("The second return argument should be a list of floats.")
return
print("No errors detected in `get_passages_with_scores`")
# + colab={"base_uri": "https://localhost:8080/"} id="kfsb4pyHYre4" outputId="f13fc8e2-f21b-44ec-b16f-caa2855b8136"
test_get_passages_with_scores(get_passages_with_scores)
# + id="SwQVdCb6Yre4"
def answer_scoring(passages, passage_probs, prompts, gen_func=run_eleuther):
"""Implements our basic scoring strategy.
Parameters
----------
passages : list of str
passage_probs : list of float
prompts : list of str
gen_func : either `run_eleuther` or `run_gpt3`
Returns
-------
list of pairs (score, dict), sorted with the largest score first.
`dict` should be the return value of `gen_func` for an example.
"""
data = []
for passage, passage_prob, prompt in zip(passages, passage_probs, prompts):
# Run `gen_func` on [prompt] (crucially, the singleton list here),
# and get the dictionary `gen` from the singleton list `gen_func`
# returns, and then use the values to score `gen` according to our
# scoring method.
#
# Be sure to use "generated_answer_probs" for the scores.
##### YOUR CODE HERE
# Return `data`, sorted with the highest scoring `(score, gen)`
# pair given first.
##### YOUR CODE HERE
# + id="JD7d8ucgYre4"
def test_answer_scoring(func):
passages = [
"Pragmatics is the study of language use.",
"Phonology is the study of linguistic sound systems."]
passage_probs = [0.75, 0.25]
prompts = passages
def gen_func(*prompts):
return [{
"generated_answer": "Constant output",
"generated_answer_tokens": ["Constant", "output"],
"generated_answer_probs": [0.1, 0.2]}]
data = func(passages, passage_probs, prompts, gen_func=gen_func)
if not all(len(x) == 2 for x in data):
print("`answer_scoring` should return a list of pairs (score, gen)")
return
if not isinstance(data[0][0], (float, np.float32, np.float64)):
print("The first member of each pair in `data` should be a score (type `float`).")
return
if not isinstance(data[0][1], dict):
print("The second member of each pair in `data` should be a dict "
"created by running `gen_func` on a single example.")
return
if data[0][0] != max([x for x, y in data]):
print("`answer_scoring` should sort its data with the highest score first.")
return
print("No errors detected in `answer_scoring`")
# + colab={"base_uri": "https://localhost:8080/"} id="QWwwDzvBYre4" outputId="f591f713-3572-4f09-a550-08fd1595835f"
test_answer_scoring(answer_scoring)
# + id="OUcdfU9SYre4"
def answer_scoring_demo(question):
"""Example usage for answer_scoring. Here we extract the top-scoring
results, which can then be used in an evaluation."""
passages, passage_probs = get_passages_with_scores(question)
prompts = [build_zero_shot_openqa_prompt(question, psg) for psg in passages]
data = answer_scoring(passages, passage_probs, prompts)
# Top result:
return data[1]
# -
answer_scoring_demo("How long is Moby Dick?")
# + [markdown] id="PmURWvoJYre4"
# ### Your original system [3 points]
#
# This question asks you to design your own few-shot OpenQA system. All of the code above can be used and modified for this, and the requirement is just that you try something new that goes beyond what we've done so far.
#
# Terms for the bake-off:
#
# * You can make free use of SQuAD and other publicly available data.
# * The LM must be an autoregressive language model. No trained QA components can be used. Our list of preallowed models are those available via the OpenAI API whose names begin with "text" and the Eluether models "gpt-neo-125M", "gpt-neo-1.3B", "gpt-neo-2.7B", and "gpt-j-6B". If you would like to use a model outside of this set, please check with the teaching team first.
#
# Here are some ideas for the original system:
#
# * We have so far sampled randomly from the SQuaD train set to create few-shot prompts. One might instead sample passages that have some connection to the target question.
#
# * We have used actual SQuAD training examples to build contexts. These might be different in meaningful ways from the passages in our corpus. An alternative is to use the SQuAD question–answer pairs to retrieve passages that contain the answer and use the resulting question–answer–passage triple when building prompts.
#
# * There are a lot of parameters to our LMs that we have so far ignored. Exploring different values might lead to better results. The `temperature` parameter is highly impactful for our task.
#
# * We have distributed a fixed index of 100K passages. These cover SQuAD plus our bake-off data, but there might still be value in creating a different/expanded index. There is starter code for indexing data with ColBERT [here](https://github.com/stanford-futuredata/ColBERT/blob/new_api/docs/intro.ipynb).
#
# * [Khattab et al. (2021a)](https://aclanthology.org/2021.tacl-1.55/) fine-tune the retriever through a handful of successive rounds, using weak supervision from the QA dataset. This is an ambitious direction that could quickly build to an original project, as the role of retriever training is under-explored so far in the context of few-shot OpenQA.
#
# * In our "Answer scoring" question, we don't normalize scores by answer length. Such normalization might be fairer to long answers and so seems worth adding.
#
# * Our "Answer scoring" question is inspired by the Retrieval Augmented Generation (RAG) model of [Lewis et al. 2020](https://arxiv.org/abs/2005.11401). Their model fully marginalizes over $k$ retrieved passages to create a proper model of $P(\textrm{answer} \mid \textrm{question})$. Implementing this requires having the probabilities for the prompts. For GPT-3, these can be obtained with `echo=False`, which will lead you to have to make changes to the output processing of `run_gpt3`. For the Eleuther models, one needs to do another call to the model forward function. Here is some starter code that could be used to begin modifying `run_eleuther`:
#
# ```
# prompt_logits = eleuther_model(prompt_ids).logits
# prompt_probs = prompt_logits.softmax(-1)
# prompt_probs = torch.gather(prompt_probs, 2, prompt_ids[:, :, None]).squeeze(-1)
# prompt_probs = [list(prompt_prob.numpy()) for p in prompt_probs]
# ```
#
# __Original system instructions__:
#
# In the cell below, please provide a brief technical description of your original system, so that the teaching team can gain an understanding of what it does. This will help us to understand your code and analyze all the submissions to identify patterns and strategies.
#
# We also ask that you report the best macro F1 score your system got during development on `dev_exs` [as defined above](#SQuAD-dev-sample), just to help us understand how systems performed overall.
#
# Please review the descriptions in the following comment and follow the instructions.
# + id="D9pfDIrPYre4"
# PLEASE MAKE SURE TO INCLUDE THE FOLLOWING BETWEEN THE START AND STOP COMMENTS:
# 1) Textual description of your system.
# 2) The code for your original system.
# 3) The score achieved by your system in place of MY_NUMBER.
# With no other changes to that line.
# You should report your score as a decimal value <=1.0
# PLEASE MAKE SURE NOT TO DELETE OR EDIT THE START AND STOP COMMENTS
# NOTE: MODULES, CODE AND DATASETS REQUIRED FOR YOUR ORIGINAL SYSTEM
# SHOULD BE ADDED BELOW THE 'IS_GRADESCOPE_ENV' CHECK CONDITION. DOING
# SO ABOVE THE CHECK MAY CAUSE THE AUTOGRADER TO FAIL.
# START COMMENT: Enter your system description in this cell.
# My peak score was: MY_NUMBER
if 'IS_GRADESCOPE_ENV' not in os.environ:
pass
# STOP COMMENT: Please do not remove this comment.
# + [markdown] id="zS3G0Ss7Yre4"
# ## Bake-off [1 point]
#
# For the bake-off, you simply need to be able to run your system on the file
#
# ```data/openqa/cs224u-openqa-test-unlabeled.txt```
#
# The following code should download it for you if necessary:
# -
if not os.path.exists(os.path.join("data", "openqa", "cs224u-openqa-test-unlabeled.txt")):
# !mkdir -p data/openqa
# !wget https://web.stanford.edu/class/cs224u/data/cs224u-openqa-test-unlabeled.txt -P data/openqa/
# + [markdown] id="zS3G0Ss7Yre4"
# If the above fails, you can just download https://web.stanford.edu/class/cs224u/data/cs224u-openqa-test-unlabeled.txt and place it in `data/openqa`.
#
# This file contains only questions. The starter code below will help you structure this. It writes a file "cs224u-openqa-bakeoff-entry.json" to the current directory. That file should be uploaded as-is. Please do not change its name.
# + id="GyttYJxoYre4"
def create_bakeoff_submission():
filename = os.path.join("data", "openqa", "cs224u-openqa-test-unlabeled.txt")
# This should become a mapping from questions (str) to response
# dicts from your system.
gens = {}
with open(filename) as f:
questions = f.read().splitlines()
# `questions` is the list of questions you need to evaluate your system on.
# Put whatever code you need to in here to evaluate your system.
# All you need to be sure to do is create a list of dicts with at least
# the keys of the dicts returned by `run_gpt` and `run_eleuther`.
# Add those dicts to `gens`.
#
# Here is an example where we just do "Open QA with no context",
# for an "original system" that would not earn any credit (since
# it is not original!):
for question in questions:
gens[question] = run_eleuther([question])[0]
# Quick tests we advise you to run:
# 1. Make sure `gens` is a dict with the questions as the keys:
assert all(q in gens for q in questions)
# 2. Make sure the values are dicts and have the key we will use:
assert all(isinstance(d, dict) and "generated_answer" in d for d in gens.values())
# And finally the output file:
with open("cs224u-openqa-bakeoff-entry.json", "wt") as f:
json.dump(gens, f, indent=4)
# -
create_bakeoff_submission()
| hw_openqa.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Training a fully-connected neural network with 2 hidden layers and ReLU activations on MNIST dataset.
import torch
import torchvision
from torchvision import datasets, transforms
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torchviz import make_dot, make_dot_from_trace
from matplotlib import pyplot as plt
# ### Load Dataset
# +
# Converting the image into torch tensor and normalizing them to fall between 0 and 1
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,)),
])
# +
# Downloading the dataset
trainset = datasets.MNIST('.', download=True, train=True, transform=transform)
testset = datasets.MNIST('.', download=True, train=False, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64,shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=64, shuffle=True)
# -
# ### Explore data
training_data= enumerate(trainloader)
batch_idx, (images, labels) = next(training_data)
images = images.view(images.shape[0], -1)
print(images.shape)
labels.shape
# ### Defining Model :
# Train a fully-connected neural network with 2 hidden layers and ReLU activations.
# +
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
# define network layers
self.hidden1 = nn.Linear(in_features=784, out_features=100)
self.hidden2 = nn.Linear(in_features=100, out_features=50)
self.output = nn.Linear(in_features=50, out_features=10)
self.relu = nn.ReLU()
def forward(self, x):
x = self.relu(self.hidden1(x))
x = self.relu(self.hidden2(x))
x = self.output(x)
return x
# instantiate the model
model = Net()
print(model)
# -
# ### Define Loss and optimizer
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(model.parameters(), lr=0.001, momentum=0.9)
# ### Train the model
# +
epochs = 20
train_loss, val_loss = [], []
accuracy_total_train, accuracy_total_val = [], []
for epoch in range(epochs):
total_train_loss = 0
total_val_loss = 0
model.train()
total = 0
correct = 0
# training our model
for idx, (images, labels) in enumerate(trainloader):
images = images.view(images.shape[0], -1)
outputs = model(images)
optimizer.zero_grad()
loss = criterion(outputs, labels)
total_train_loss += loss.item()
loss.backward()
optimizer.step()
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
accuracy_train = correct / total
accuracy_total_train.append(accuracy_train)
total_train_loss = total_train_loss / (idx + 1)
train_loss.append(total_train_loss)
# validating our model
model.eval()
total = 0
correct = 0
for idx, (images, labels) in enumerate(testloader):
images = images.view(images.shape[0], -1)
outputs = model(images)
loss = criterion(outputs, labels)
total_val_loss += loss.item()
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
accuracy_val = correct/ total
accuracy_total_val.append(accuracy_val)
total_val_loss = total_val_loss / (idx + 1)
val_loss.append(total_val_loss)
if epoch % 2 == 0:
print("Epoch: {}/{} ".format(epoch, epochs),
"Training loss: {:.4f} ".format(total_train_loss),
"Testing loss: {:.4f} ".format(total_val_loss),
"Train accuracy: {:.4f} ".format(accuracy_train),
"Test accuracy: {:.4f} ".format(accuracy_val))
# -
# ### Plot Train Loss
plt.plot(train_loss, label='Training loss')
plt.plot(val_loss, label='Test loss')
plt.legend()
plt.grid()
# ### Plot Training Accuracy
plt.plot(accuracy_total_train, label='Training Accuracy')
plt.plot(accuracy_total_val, label='Test Accuracy')
plt.legend()
plt.grid()
| Deep Learning/MNIST_classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## The CSV File Format
#
# One simple way to store data in a text file is to write the data as a series of values separated by commas, called comma-separated values.
#
# The CSV files were downloaded from: https://github.com/ehmatthes/sitka_weather_hx
#
# `csv.reader()` creates a reader object associated with the file `f`
#
# `next()` return the next line in a csv file.
#
# BTW, Sitka is a city in Alaska and AKDT means Alaska Daylight Time.
#
# Max TemperatureF tells us that the second value in each line is the maximum temperature for that date in F degrees.
#
# ### Parsing the CSV File Headers
#
# +
import csv
filename = 'sitka_weather_07-2014.csv'
with open(filename) as f:
reader = csv.reader(f)
header_row = next(reader)
print (header_row)
print ("")
for index, column_header in enumerate(header_row):
print(index, column_header)
# -
# ### Extracting and Reading Data
#
# Initially, we want to plot the maximum temperature per day. So we need to extract the data from columns 0 and 1.
#
# Because we've already read the header ow, the loop will begin at the second line wheere the actual data begins.
# +
import csv
# Get high temperatures from file.
filename = 'sitka_weather_07-2014.csv'
with open(filename) as f:
reader = csv.reader(f)
header_row = next(reader)
highs = []
for row in reader:
high = int(row[1])
highs.append(high)
print (highs)
# -
# ### Plotting Data in a Temperature Chart
# +
from matplotlib import pyplot as plt
# Plot data.
fig = plt.figure(dpi=128, figsize=(10, 6))
plt.plot(highs, c='red')
# Format plot.
plt.title("Daily high temperatures, July 2014", fontsize=24)
plt.xlabel('', fontsize=16)
plt.ylabel("Temperature (F)", fontsize=16)
plt.tick_params(axis='both', which='major', labelsize=16)
plt.show()
# -
# ### The datetime Module
#
# The data will be read in as a string, so we need a way to convert the string '2014-7-1' to an object representing this date.
#
# `strptime()` takes the string containing the date as the first argument. The second argument tells Python how the date is formatted.
#
# In this example, '%Y-' tells Python to interpret the part of the string before the first dash as a four-digit year, and so on.
#
# `strptime()` can take a variety of arguments to determine how to interpret the date.
from datetime import datetime
first_date = datetime.strptime('2014-7-1', '%Y-%m-%d')
print(first_date)
# ### Plotting Dates
#
# We can now improve our plot of the temperature data by extracting date for the daily highs and passing the dates and the highs to `plot()`.
#
# `fig.autofmt_xdate()` draws the date labels diagonally to prevent them from overlapping.
# +
import csv
from datetime import datetime
from matplotlib import pyplot as plt
# Get dates, and high temperatures from file.
filename = 'sitka_weather_07-2014.csv'
with open(filename) as f:
reader = csv.reader(f)
header_row = next(reader)
dates, highs = [], []
for row in reader:
current_date = datetime.strptime(row[0], "%Y-%m-%d")
dates.append(current_date)
high = int(row[1])
highs.append(high)
# Plot data.
fig = plt.figure(dpi=128, figsize=(10, 6))
plt.plot(dates, highs, c='red')
# Format plot.
plt.title("Daily high temperatures, July 2014", fontsize=24)
plt.xlabel('', fontsize=16)
fig.autofmt_xdate()
plt.ylabel("Temperature (F)", fontsize=16)
plt.tick_params(axis='both', which='major', labelsize=16)
plt.show()
# -
# ### Plotting a longer Timeframe
#
# We modify the filename to use the new data file sitka_weather_2014.csv.
#
# We update the title of our plot to reflect the change in its content.
# +
import csv
from datetime import datetime
from matplotlib import pyplot as plt
# Get dates, and high temperatures from file.
filename = 'sitka_weather_2014.csv'
with open(filename) as f:
reader = csv.reader(f)
header_row = next(reader)
dates, highs = [], []
for row in reader:
current_date = datetime.strptime(row[0], "%Y-%m-%d")
dates.append(current_date)
high = int(row[1])
highs.append(high)
# Plot data.
fig = plt.figure(dpi=128, figsize=(10, 6))
plt.plot(dates, highs, c='red')
# Format plot.
plt.title("Daily high temperatures - 2014", fontsize=24)
plt.xlabel('', fontsize=16)
fig.autofmt_xdate()
plt.ylabel("Temperature (F)", fontsize=16)
plt.tick_params(axis='both', which='major', labelsize=16)
plt.show()
# -
# ### Plotting a Second Data Series
#
# We want to extract the low temperatures from the data file and then add them to our graph.
# +
import csv
from datetime import datetime
from matplotlib import pyplot as plt
# Get dates, high, and low temperatures from file.
filename = 'sitka_weather_2014.csv'
with open(filename) as f:
reader = csv.reader(f)
header_row = next(reader)
dates, highs, lows = [], [], []
for row in reader:
current_date = datetime.strptime(row[0], "%Y-%m-%d")
dates.append(current_date)
high = int(row[1])
highs.append(high)
low = int(row[3])
lows.append(low)
# Plot data.
fig = plt.figure(dpi=128, figsize=(10, 6))
plt.plot(dates, highs, c='red')
plt.plot(dates, lows, c='blue')
# Format plot.
plt.title("Daily high and low temperatures - 2014", fontsize=24)
plt.xlabel('', fontsize=16)
fig.autofmt_xdate()
plt.ylabel("Temperature (F)", fontsize=16)
plt.tick_params(axis='both', which='major', labelsize=16)
plt.show()
# -
# ### Shading an Area in the Chart
#
# `fill_between()` takes a series of x-values and 2 series of y-values, and fills the space between the 2 y-value series.
#
# `alpha` controls a color's transparency. Where 0 means completely transparent.
# +
import csv
from datetime import datetime
from matplotlib import pyplot as plt
# Get dates, high, and low temperatures from file.
filename = 'sitka_weather_2014.csv'
with open(filename) as f:
reader = csv.reader(f)
header_row = next(reader)
dates, highs, lows = [], [], []
for row in reader:
current_date = datetime.strptime(row[0], "%Y-%m-%d")
dates.append(current_date)
high = int(row[1])
highs.append(high)
low = int(row[3])
lows.append(low)
# Plot data.
fig = plt.figure(dpi=128, figsize=(10, 6))
plt.plot(dates, highs, c='red', alpha=0.5)
plt.plot(dates, lows, c='blue', alpha=0.5)
plt.fill_between(dates, highs, lows, facecolor='blue', alpha=0.1)
# Format plot.
plt.title("Daily high and low temperatures - 2014", fontsize=24)
plt.xlabel('', fontsize=16)
fig.autofmt_xdate()
plt.ylabel("Temperature (F)", fontsize=16)
plt.tick_params(axis='both', which='major', labelsize=16)
plt.show()
# -
# ### Error-Checking
#
# Missing data can result in exceptions that crash our programs if we don't handle them properly.
#
# On Feb 16, 2014 no data was recorded.
# +
import csv
from datetime import datetime
from matplotlib import pyplot as plt
# Get dates, high, and low temperatures from file.
filename = 'death_valley_2014.csv'
with open(filename) as f:
reader = csv.reader(f)
header_row = next(reader)
dates, highs, lows = [], [], []
for row in reader:
current_date = datetime.strptime(row[0], "%Y-%m-%d")
dates.append(current_date)
high = int(row[1])
highs.append(high)
low = int(row[3])
lows.append(low)
# Plot data.
fig = plt.figure(dpi=128, figsize=(10, 6))
plt.plot(dates, highs, c='red', alpha=0.5)
plt.plot(dates, lows, c='blue', alpha=0.5)
plt.fill_between(dates, highs, lows, facecolor='blue', alpha=0.1)
# Format plot.
plt.title("Daily high and low temperatures - 2014", fontsize=24)
plt.xlabel('', fontsize=16)
fig.autofmt_xdate()
plt.ylabel("Temperature (F)", fontsize=16)
plt.tick_params(axis='both', which='major', labelsize=16)
plt.show()
# -
# To address this issue, we'll run error checking code when the values are being read from the CSV file to handle exceptions that might arise when we parse our data sets.
# +
import csv
from datetime import datetime
from matplotlib import pyplot as plt
# Get dates, high, and low temperatures from file.
filename = 'death_valley_2014.csv'
with open(filename) as f:
reader = csv.reader(f)
header_row = next(reader)
dates, highs, lows = [], [], []
for row in reader:
try:
current_date = datetime.strptime(row[0], "%Y-%m-%d")
high = int(row[1])
low = int(row[3])
except ValueError:
print(current_date, 'missing data')
else:
dates.append(current_date)
highs.append(high)
lows.append(low)
# Plot data.
fig = plt.figure(dpi=128, figsize=(10, 6))
plt.plot(dates, highs, c='red', alpha=0.5)
plt.plot(dates, lows, c='blue', alpha=0.5)
plt.fill_between(dates, highs, lows, facecolor='blue', alpha=0.1)
# Format plot.
title = "Daily high and low temperatures - 2014\nDeath Valley, CA"
plt.title(title, fontsize=20)
plt.xlabel('', fontsize=16)
fig.autofmt_xdate()
plt.ylabel("Temperature (F)", fontsize=16)
plt.tick_params(axis='both', which='major', labelsize=16)
plt.show()
# -
# Here we used a `try-except-else` block to handle missing data. Sometimes you will use `continue` to skip over some data or `remove()` or `del` to eliminate some data after it's been extracted.
| 9_Project/2-Reading-data-from-CSV.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # [作業目標]
# - 利用範例的創建方式, 創建一組資料, 並練習如何取出最大值
# # [作業重點]
# - 練習創立 DataFrame (In[2])
# - 如何取出口數最多的國家 (In[3], Out[3])
# ## 練習時間
# 在小量的資料上,我們用眼睛就可以看得出來程式碼是否有跑出我們理想中的結果
#
# 請嘗試想像一個你需要的資料結構 (裡面的值可以是隨機的),然後用上述的方法把它變成 pandas DataFrame
#
# #### Ex: 想像一個 dataframe 有兩個欄位,一個是國家,一個是人口,求人口數最多的國家
#
# ### Hints: [隨機產生數值](https://blog.csdn.net/christianashannon/article/details/78867204)
import pandas as pd
import numpy as np
data ={}
data['國家'] = ['US','UK','Canada','Taiwan','New Zealand']
data['人口'] = np.random.randint(100000000, size=5)
df = pd.DataFrame(data)
print(df)
df[df.index==df['人口'].idxmax()]
| Day_005-1_HW.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Prediction
# This notebook is used to predict on the given values for the saved trained model.
import pandas as pd
import numpy as np
import pickle
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import f1_score
from math import sin, cos, sqrt, atan2, radians
# ## Methods
def loadModel(name):
'''
function to load model object with the given name.
'''
with open('store/'+name+'_model.pkl', 'rb') as f:
model = pickle.load(f)
return model
def discretizeTime(time):
'''
function to map 24-hours format time to one of the six 4 hours intervals.
'''
if '00:00' <= time < '04:00':
return 't1'
elif '04:00' <= time < '08:00':
return 't2'
elif '08:00' <= time < '12:00':
return 't3'
elif '12:00' <= time < '16:00':
return 't4'
elif '16:00' <= time < '20:00':
return 't5'
elif '20:00' <= time < '24:00':
return 't6'
def calculateDistance(src, dst):
'''
function to calculate the distance between two locations on earth
using src & dst tuples given in the format (latitude, longitude).
'''
# approximate radius of earth in km
R = 6373.0
# approximate 1 km to miles conversion
to_miles = 0.621371
lat1 = radians(abs(src[0]))
lon1 = radians(abs(src[1]))
lat2 = radians(abs(dst[0]))
lon2 = radians(abs(dst[1]))
dlon = lon2 - lon1
dlat = lat2 - lat1
a = sin(dlat / 2)**2 + cos(lat1) * cos(lat2) * sin(dlon / 2)**2
c = 2 * atan2(sqrt(a), sqrt(1 - a))
return R * c * to_miles
def isNear(location, data, radius):
'''
function to determine if the given location (latitude, longitude)
is near to any location in the given data (dataframe) based on the given radius.
'''
for index, row in data.iterrows():
if calculateDistance(location, (row['latitude'], row['longitude'])) <= radius:
return 1
return 0
def loadLabelEncoder(column):
'''
function to load label encoder object for the given column.
'''
with open('store/'+column+'_label_encoder.pkl', 'rb') as f:
le = pickle.load(f)
return le
def createDataPoint(day, district, longitude, latitude, month, time):
columns = [
loadLabelEncoder('day').transform(np.array([day]))[0],
loadLabelEncoder('district').transform(np.array([district]))[0],
longitude,
latitude,
loadLabelEncoder('month').transform(np.array([month]))[0],
loadLabelEncoder('time_interval').transform(np.array([discretizeTime(time)]))[0],
0,
isNear((latitude, longitude), pd.read_pickle('store/facilities.pkl', compression='gzip'), 1),
isNear((latitude, longitude), pd.read_pickle('store/private_spaces.pkl', compression='gzip'), 1),
isNear((latitude, longitude), pd.read_pickle('store/colleges.pkl', compression='gzip'), 1),
isNear((latitude, longitude), pd.read_pickle('store/public_open_spaces.pkl', compression='gzip'), 1),
isNear((latitude, longitude), pd.read_pickle('store/commuter_stops.pkl', compression='gzip'), 1),
isNear((latitude, longitude), pd.read_pickle('store/public_park.pkl', compression='gzip'), 1),
isNear((latitude, longitude), pd.read_pickle('store/landmarks.pkl', compression='gzip'), 1),
isNear((latitude, longitude), pd.read_pickle('store/schools.pkl', compression='gzip'), 1)
]
return np.array(columns).reshape(1, -1)
def getLabel(pred):
return loadLabelEncoder('label').inverse_transform(pred)[0]
# ## Model
model = loadModel('randomForest')
getLabel(model.predict(createDataPoint('sunday', 'tenderloin',-122.414406, 37.784191, 'february', '02:00')))
| data_prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
# +
# MASTER ONLY
# %load_ext prog_edu_assistant_tools.magics
from prog_edu_assistant_tools.magics import report, autotest
# -
# ```
# # ASSIGNMENT METADATA
# assignment_id: "1-1"
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# ## 練習
# + [markdown] slideshow={"slide_type": "-"}
# 黄金比を求めてください。黄金比とは、5 の平方根に 1 を加えて 2 で割ったものです。約 1.618 になるはずです。
#
# ```
# # EXERCISE METADATA
# exercise_id: "GoldenRatio"
# ```
#
# -
# %%solution
import math
""" # BEGIN PROMPT
golden_ratio = ....
print(golden_ratio)
""" # END PROMPT
# BEGIN SOLUTION
golden_ratio = (math.sqrt(5.) + 1)/2
print(golden_ratio)
# END SOLUTION
# %%studenttest StudentTest_GoldenRatio
assert abs(golden_ratio - 1.618) < 0.001
# %%inlinetest AutograderTest_GoldenRatio
assert 'golden_ratio' in globals(), "Did you define the variable golden_ratio?"
assert str(golden_ratio.__class__) == "<class 'float'>", f"Did you compute a number? Found {golden_ratio.__class__}"
assert abs(golden_ratio - 1.618) < 0.001, f"Did you compute golden ratio correctly? Expected a number close to 1.618, but got {golden_ratio}"
assert golden_ratio != 1.618, f"Did you use math.sqrt() to compute the golden ratio? Expected a number with many digits after the point, but got exactly 1.618"
# %%submission
# Incorrect submission
golden_ratio = 1.618
# Test the incorrect submission against an autograder test
result, log = %autotest AutograderTest_GoldenRatio
assert not result.results['passed']
assert 'got exactly 1.618' in str(result.results['error'])
report(AutograderTest_GoldenRatio, results=result.results)
| 1/1-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Model Inference
# Inference using `pyhf` Python API
import pyhf
import json
# +
with open("data/2-bin_1-channel.json") as serialized:
spec = json.load(serialized)
workspace = pyhf.Workspace(spec)
model = workspace.model(poi_name="mu")
# -
pars = model.config.suggested_init()
data = workspace.data(model)
# Creating the log-likelihood conditioned on the data
# $$
# L(\vec{\theta}|\vec{x}) = k \cdot p(\vec{x} | \vec{\theta})
# $$
model.logpdf(pars, data)
# moar inference
bestfit_pars, twice_nll = pyhf.infer.mle.fit(data, model, return_fitted_val=True)
print(bestfit_pars)
# objective function is twice the negative log-likelihood
-2 * model.logpdf(bestfit_pars, data) == twice_nll
# ## Hypothesis Testing
# Very often in physics analyses we want to compute the $\mathrm{CL}_{s}$ — a modified pseudo-frequentist $p$-value. We can use the API that `pyhf` provides for hypothesis testing to compute the expected and observed $\mathrm{CL}_{s}$.
test_mu = 1.0
CLs_obs, CLs_exp = pyhf.infer.hypotest(test_mu, data, model, return_expected=True)
print('Observed: {}, Expected: {}'.format(CLs_obs, CLs_exp))
# As a quick aside, remembering that
# $$
# \mathrm{CL}_{s} = \frac{\mathrm{CL}_{s+b}}{\mathrm{CL}_{b}} = \frac{p_{s+b}}{1-p_{b}}
# $$
CLs_obs, p_values = pyhf.infer.hypotest(test_mu, data, model, return_tail_probs=True)
print('Observed CL_s: {}, CL_sb: {}, CL_b: {}'.format(CLs_obs, p_values[0], p_values[1]))
# we can explicitly check `pyhf` with
assert CLs_obs == p_values[0]/p_values[1]
# We often are intersted in computing a band of expected $\mathrm{CL}_{s}$ values (the "Brazil band")
# +
import numpy as np
CLs_obs, CLs_exp_band = pyhf.infer.hypotest(test_mu, data, model, return_expected_set=True)
print('Observed CL_s: {}\n'.format(CLs_obs))
for p_value, n_sigma in enumerate(np.arange(-2,3)):
print('Expected CL_s{}: {}'.format(' ' if n_sigma==0 else '({} σ)'.format(n_sigma),CLs_exp_band[p_value]))
# -
# We can scan across test values of our POI ($\mu$) to perform hypothesis testing for our model
results = []
poi_vals = np.linspace(0,5,41)
for test_poi in poi_vals:
results.append(
pyhf.infer.hypotest(test_poi, data, model, return_expected_set=True)
)
# and then invert those hypothesis tests to set upper limits on the model parameters
# +
obs = [cls_vals[0][0] for cls_vals in results]
exp = [cls_vals[1][2][0] for cls_vals in results]
alpha = 0.05
cls_limit = (1.-alpha)*100
print(f"{cls_limit}% CLs Upper Limit (observed): µ = {np.interp(0.05, obs[::-1], poi_vals[::-1]):0.3}")
print(f"{cls_limit}% CLs Upper Limit (expected): µ = {np.interp(0.05, exp[::-1], poi_vals[::-1]):0.3}")
# -
# ## Visualization
# We can visualize this by plotting the observed and expected results (the "Brazil band")
import matplotlib.pyplot as plt
import pyhf.contrib.viz.brazil
# +
fig, ax = plt.subplots(1,1)
fig.set_size_inches(7, 5)
ax.set_xlabel(r"$\mu$ (POI)")
ax.set_ylabel(r"$\mathrm{CL}_{s}$")
pyhf.contrib.viz.brazil.plot_results(ax, poi_vals, results)
| book/inference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Operations in sets
#In this tutorial, I want to explain the operations in sets with a couple of examples
#First of all you should know, there is no order in set elements
#Second of all, repetitive items will be removed in sets
#With this introduction let's learning set's operations in Python
# add method
#Assume there is bmj1 as a set which has been filled as follows:
bmj1={12, 13, "tit", "reza", 15}
print(bmj1)
# "add method" add a new element to a set
bmj1.add(14)
print (bmj1)
bmj1.add("ali")
print (bmj1)
# +
# You are nto allowed to add two or more items with add method simultaneously
#If you add a element twice the set takes no notice of the repetitive new
bmj1={12, 13, "tit", "reza", 15}
bmj1.add("tit")
print(bmj1)
bmj1={12, 13, "tit", "reza", 15}
bmj1.add(13)
print(bmj1)
# +
#We can remove the items with "remove method"
var1={42.3, 43.67, "Dev", "Python", "ANN"}
var1.remove("Dev")
print(var1)
bmj1={12, 13, "tit", "reza", 15}
bmj1.remove("tit")
print(bmj1)
var1={42.3, 43.67, "Dev", "Python", "ANN"}
var1.remove("Dev")
var1.remove("Python")
var1.remove("ANN")
print(var1)
# +
#We can verify an item is in the set or not using "in method" as following form
mo={"xc", "gh","es", "fr", "wa"}
r= "ds" in mo
print(bool(r))
r="gh" in mo
print(bool(r))
r= "xs" in mo
print(bool(r))
r="xc" in mo
print(bool(r))
# +
# Mathematical operation in sets
#intersection of two sets make a set containing the elements of both sets
Set1={1,"bm", 3, 4, 23, 54, "as"}
Set2={12, "rty","ali", "bm", 23, "er"}
Set3=Set1.intersection(Set2)
print(Set1)
print(Set2)
print(Set3)
#Overlapping in sets can be computed using "&"
#In Python, we just place "ampersand"= sign "&" between two sets to calculate their overlaps
#Therefore, both ampersand & and "intersection method" are same
Set1={1,"bm", 3, 4, 23, 54, "as"}
Set2={12, "rty","ali", "bm", 23, "er"}
Set3=Set1 & Set2
print(Set1)
print(Set2)
print(Set3)
#Union of set
#Global sets
#To calculate union of set, we can use "union method" as follows
Set1={1,"bm", 3, 4, 23, 54, "as"}
Set2={12, "rty","ali", "bm", 23, "er"}
Set3={"bm", "er", 12}
U1=Set1.union(Set2)
U2=Set2.union(Set3)
U3=Set1.union(Set3)
print("Union Set1 and Set2: ", U1)
print("Union Set1 and Set2: ", U2)
print("Union Set1 and Set2: ", U3)
#Subset of set
#Python allows you to check whether a set is a subset for another set or not
#To check subset you just use "issubset method" and get results as "True" or "False"
Set1={1,"bm", 3, 4, 23, 54, "as"}
Set2={12, "rty","ali", "bm", 23, "er"}
Set3={"bm", "er", 12}
r1=Set3.issubset(Set2)
r2=Set2.issubset(Set3)
r3=Set1.issubset(Set2)
print("Set3 is a subset of Set2: ",bool(r1))
print("Set3 is a subset of Set2: ",bool(r2))
print("Set3 is a subset of Set2: ",bool(r3))
# Find the difference in set1 but not set2
Set1={1,"bm", 3, 4, 23, 54, "as"}
Set2={12, "rty","ali", "bm", 23, "er"}
Set3={"bm", "er", 12}
Setd1=Set1.difference(Set2)
Setd2=Set2.difference(Set1)
Setd3=Set1.difference(Set3)
print("Difference Set1 than Set2: ", Setd1)
print("Difference Set2 than Set1: ", Setd2)
print("Difference Set1 than Set3: ", Setd3)
# -
| SetOperations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Tariffs in General equilibrium
# ## Background
#
# This notebook builds the diagrams for the general equilibrium analysis of a production subsidy or a tariff (production subsidy plus consumption tax).
#
# **THIS IS A DRAFT:** The notebook thus far illustrates just the supply side distortions of a production subsidy. In the next iteration I will also analyze the effects of a tariff on production and consumption/utility, and plots will properly display consumption indifference curves.
from sfm import *
# + tags=["hide-input"]
plt.rcParams["figure.figsize"] = [7,7]
import warnings
warnings.filterwarnings('ignore', 'The iteration is not making good progress')
# -
# ### The production possibility frontier
#
# We will use a bowed-out production possibility frontier. The details of how this is built don't really matter to the analysis at hand but it will be easy to illustrate by leveraging our earlier analysis of the specific factors model.
#
# Agriculture requires specific capital $T$ and mobile labor:
#
# $$Q_a = F(\bar T, L_a)$$
#
# Manufacturing production requires specific capital $K$ and mobile labor:
#
# $$Q_m = G(\bar K, L_m)$$
#
# The quantity of land in the agricultural sector and the quantity of capital in the manufacturing sector are both in fixed inelastic supply during the period of analysis.
#
# The market for mobile labor is competitive and the market clears at a wage where the sum of labor demands from each sector equals total labor supply.
#
# $$
# L_a + L_m = \bar{L}
# $$
# From this it is easy to build a production possibility frontier (to wit: at every level of $L_A$ calculate and plot $Q_a=F(K,L_a)$ and $Q_m=G(K, \bar L - L_a)$.
#
# See the SFM notebook for details on the parameters used to create these visualizations.
ppf(Tbar=100, Kbar=100, Lbar=400)
# ### The closed economy
#
# Let $p$ be the relative price of manufactures
#
# $$
# p=\frac{P_m}{P_a}
# $$
#
# With the assumed technologies and parameters, the economy in autarky will have an autarky relative equilibrium price of $p=1$ and would look like this:
open_trade(p=1, t = 0)
# ### Production subsidy and tariff
# An ad-valorem subsidy on manufactures of $t$ will raises the effective price received by the domestic supplier from $P_M$ to $P_m(1+t)$. If we assume no tax or subsidy on agriculture then this also raises the relative price of manufactures from $p$ to $p(1+t)$.
#
# A tariff can be thought of as a production subsidy to suppliers plus a consumption tax to consumers.
#
# Suppose the small but closed economy with market equilibrium relative price $p^a$ opens to trade. Since it is a small economy its own domestic prices will now be set by the world relative price, let's call it $p^w$.
#
# The following diagram shows the open economy of a country which had a comparative advantage in the production of agricultural goods. As depicted as it opened to trade the relative price of manufactured goods falls from $p=p^a=1$ to $p=p^w=1/2$. We depict the market equilibrium (with no subsidy or tariff). As the price rises producers cut back on manufacturing production and expand agricultural production to take advantage of the price change. Meanwhile consumers substitute away from the now relatively more expensive agricultural good.
#
# In this new equilibrium the country exports agricultural goods in exchange for manufactures.
open_trade(p=1/2, t = 0)
# ### A production subsidy to manufacturers
#
# The graph below depicts the effects of a production subsidy on equlibrium.
# Producers faced the distorted prices (steeper price line in graph below). The subsidy induced rise in the relative price of manufactures leads the economy to a new production point along the PPF (where the PPF is tangent to the new distorted price line). This production bundle is less valuable measured at world prices than the early bundle produced when the country followed it's comparative advantage. That can be seen by comparing the world price line (national income at world prices) passing through this new bundle compared to a similar world price line passing through the free trade bundle that was chosen before the country distorted its domestic prices (not drawn below, but see the following figure) which can be seen to have higher intercepts (meaning measured in real terms in terms of either good national income is higher in the undistorted open economy equilibrium).
#
# We can also see (see this and following diagram) how consumer welfare is reduced.
open_trade(p=1/2, t = 1, dt=True)
interact(open_trade, p=(0.25,2,0.25), t = (0,1,0.05) );
# It gets a bit busy but if we draw the situation before and after on the same graph we can see clearly how the production subsidy, while it does raise the production of manufactures, also lowers national income measured at world prices as well as welfare.
open_trade(p=1/2, t = 0, dt=False)
open_trade(p=1/2, t = 1, dt=True)
def twoplot(t):
open_trade(p=1/2, t = 0, dt=False)
open_trade(p=1/2, t = t, dt=True)
interact(twoplot,t=(0,2,0.1));
# ### A tariff
#
# A tariff can be analyzed just as the production subsidy above, only that now consumers also face higher prices $p(1+t)$.
#
# The country still trades with the world at world prices $p=p^w$ but consumers face the price distortion introduced by the tariff.
#
# We don't draw this situatino but look at the world price line (or national income line) at world prices running through the subsidy distorted production bundle. The country must trade along this line. But if consumers face distorted prices consumption must be at a point where the community indifference is tangent to these distorted price lines. Technically we can think of moving the distorted price line out parallel from the PPF along this world price line until we touch an indifference curve tangent to this distorted line. It's easy to verify that this will take us to an indifference curve below the lowest indifference curve above. Hence we see that in addition to the deadweight loss (lost national income) from producing the wrong bundle of goods, when analysing a tariff, we also see the consumer loss of welfare from the consumer tax.
| notebooks/trade/Tariff_in_general_equilibrium.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="K80Nz7J48L4a"
# ## **Preliminary Implementation of the LFattNet Approach Over Different LF Images**
#
# ##### $ Training: ${'additional/museum', 'additional/kitchen'}
# ##### $ Validation: ${stratified/backgammon', 'stratified/dots'}
# ##### $ Testing: ${'LFattNet_output/backgammon', 'LFattNet_output/dots', 'LFattNet_output/boxes', 'LFattNet_output/cotton'}
# + [markdown] id="82apOZ4F8L1U"
# **Preliminary Implementation Properties:**
# * Modified to work with newer Tensorflow, Keras, and Python versions
# * Defualt Dataset
# * Removed unwanted warnings
# + [markdown] id="VnEAWiK68LyO"
# After uploading the HCL scene images to Google Drive, and all the necessary modified version of the code. The following steps show the preliminary implementation and results of the code.
# + [markdown] id="MM7z52vC8LvB"
# **First:** Connect w/ Google Drive
# + id="DpiEz28oezR_" colab={"base_uri": "https://localhost:8080/"} outputId="8e439247-80ad-4a28-b8bf-3992ebbc6ac6"
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="v1hB0TY0-OCu"
# **Second:** Navigate through drive
# + colab={"base_uri": "https://localhost:8080/"} id="jir9grNeCBWB" outputId="de8716b1-9dcd-40bf-be5f-0a5a8c79004b"
# cd gdrive/MyDrive/LFattNet-master/
# + [markdown] id="Ci8gEIGj-Ov8"
# **Third:** Check current working packages' version for code testing
# + id="eMbDaEM4FY4-" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="25dc30d0-50fa-4f1b-9c5a-1b65f7ff16ca"
import tensorflow as tf
tf.__version__
# + colab={"base_uri": "https://localhost:8080/"} id="2YkMjPd_UiSs" outputId="1aa2c1b2-fad1-4329-b053-643b7d4fcc24"
import keras
print(keras.__version__)
# + [markdown] id="K8x1OJTf-P1P"
# **Forth:** Download needed packages
# + id="Up404HJb-imQ" colab={"base_uri": "https://localhost:8080/"} outputId="a61fb747-6864-4e1a-df74-e2edf4ed47f5"
pip install tensorflow-addons==0.15
# + [markdown] id="wuIxdbW1-Qaz"
# ### **Fifth:** Train & Validate the model...
#
# *Note: uncomment LFattNet_train.py (Lines 171,172, 183, 198), LFattNet_func/func_model_81.py (Line 250)*
# + colab={"base_uri": "https://localhost:8080/"} id="NrxgSJiZ4TYd" outputId="a82df9e9-944f-44ca-dea0-ffdc5b94940c"
# !python LFattNet_train.py
# + [markdown] id="vqGU99dM-RtC"
# ### **Sixth:** Test the model...
#
# *Note: uncomment LFattNet_evalution.py (Line 71,72), LFattNet_func/func_model_81.py (Line 253)*
# + id="WOp3UN6C-ijt" colab={"base_uri": "https://localhost:8080/"} outputId="c5b08b38-875a-4ea7-90c2-ce6485f903e4"
# !python LFattNet_evalution.py
# + [markdown] id="PiUAYoLT_5eD"
# ### ** Test validation set...
# + id="OgS8HIPA-iaI" colab={"base_uri": "https://localhost:8080/"} outputId="fd0739c8-5750-4964-9661-07c8af5741b6"
# !python LFattNet_evalution.py
# + [markdown] id="MjKrNCZOAErP"
# Metric performances are shown above. For further inspection regarding the output images. Please review the paper's figures and the result part.
# + id="hrL4_qK7AM5H"
| LFNatt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Implementation of adaptive sampling protocols guided by SAXS and hybrid information in kinetic Monte Carlo simulations on the MSM of MoaD-MoaE asssociation.
from msmbuilder.utils import io
import numpy as np
# Load the previously constructed Markov state model (MSM) on the MoaD-MoaE association.
# Number of states: 500, Lag time: 40 ns
msm = io.load("MSM.pkl")
print(msm.n_states_, msm.lag_time)
# Load the calculated SAXS discrepancy scores between the SAXS profiles of each state and the target.
SAXS = np.loadtxt("MoaD-MoaE_discrepancy.txt")
def loadSAXSdata(SAXS):
N_states = np.shape(SAXS)[0]
SAXS_dict = {}
for i in range(N_states):
state = int(SAXS[i][0])
intensity = SAXS[i][1]
SAXS_dict[state] = intensity
return SAXS_dict
saxs_dict = loadSAXSdata(SAXS)
# Load the average distances between the top 5 evolutionarily coupled residue pairs at the MoaD-MoaE interface.
EC = np.loadtxt("MoaD-MoaE_avg_dists.txt")
def loadECdata(EC):
N_states = np.shape(EC)[0]
EC_dict = {}
for i in range(N_states):
state = int(i)
dist = EC[i]
EC_dict[state] = dist
return EC_dict
ec_dict = loadECdata(EC)
# Define the initial and the target states where kinetic Monte Carlo simulations start from and end at.
init = 359
end = 241
# Visualize the 500 MSM states, projected onto x, y axis on SAXS discrepancy score and average residue pair distance.
import matplotlib.pyplot as plt
# %matplotlib inline
import math
from matplotlib import rc
import matplotlib.colors as plc
rc('font',**{'family':'sans-serif','sans-serif':['Helvetica'], 'size': '24'})
params = {'mathtext.default': 'regular'}
plt.rcParams.update(params)
plt.rcParams['figure.figsize'] = [8, 8]
import numpy as np
from matplotlib.font_manager import FontProperties
fontP = FontProperties()
fontP.set_size('24')
fig, ax = plt.subplots()
ax.scatter(np.transpose(SAXS)[1], np.transpose(EC), lw=0.3, color="#f46855", \
s=msm.populations_/np.max(msm.populations_)*200, zorder=2, label='all')
ax.scatter(np.transpose(SAXS)[1][init], np.transpose(EC)[init], lw=0.3, color="blue", \
s=msm.populations_[init]/np.max(msm.populations_)*200, zorder=2, label='initial')
ax.scatter(np.transpose(SAXS)[1][end], np.transpose(EC)[end], lw=0.3, color="green", \
s=msm.populations_[end]/np.max(msm.populations_)*2500, marker='*', zorder=3, label='target')
ax.set_xlim(1,10000)
ax.set_xscale("log")
ax.set_ylim(0,200)
ax.set_yticks((0,50,100,150,200))
ax.set(xlabel=r'SAXS discrepancy $\chi$$^{2}$', ylabel='Average residue pair distance (${\AA}$)')
ax.legend(loc=2, ncol=1, frameon=False, prop=fontP, numpoints=1, scatterpoints=1)
ax.tick_params(axis='x', which='minor', top='off')
# ### Perform SAXS-guided adaptive sampling in kinetic Monte Carlo simulations.
# Define the kernel to run kinetic Monte Carlo simulation on MSM.
def run(init,msm,step):
try:
traj = msm.sample_discrete(state=init, n_steps=step, random_state=None)
except KeyError:
print("ERROR, using random state for starting!")
traj = msm.sample_discrete(state=None, n_steps=step)
return traj
# #### Step 1: Running 10 short parallel simulations starting from the initial states
Num=10 # Number of parallel simulations to run
step=5 # Individual trajectory length, 5 * lag time = 200 ns
trajs = []
initRoundFlag = True
inits = np.zeros(Num, dtype=int)
trajs_rnd = []
if initRoundFlag:
for i in range(Num): # Run 10 parallel simulations
traj = run(init, msm, step)
trajs_rnd.append(traj)
traj = [ traj ]
trajs.append(traj)
unique = np.unique(trajs_rnd)
SAXS_diff = [ saxs_dict.get(state) for state in unique ]
print(trajs) # 10 trajectories sampled from kinetic Monte Carlo simulations
print(unique) # sampled states from the first round of samplings
print(len(unique)) # total number of cluster states sampled from kinetic Monte Carlo simulations
# #### Step 2: Clustering the sampled trajectories based on some structural features; (skipped as clustering is already performed)
# #### Step 3: Picking the 10 cluster states with the lowest SAXS-discrepancy scores.
# +
if np.shape(unique)[0] < 10:
for i in range(np.shape(unique)[0]):
inits[i] = int(unique[i])
for i in range(10-np.shape(unique)[0]):
inits[np.shape(unique)[0] + i] = int(np.random.choice(unique,1))
if np.shape(unique)[0] == 10:
for i in range(np.shape(unique)[0]):
inits[i] = int(unique[i])
if np.shape(unique)[0] > 10:
sorted_SAXS_diff = np.unique(SAXS_diff)
for i in range(Num):
index = 0
for saxs in SAXS_diff:
if saxs == sorted_SAXS_diff[i]:
inits[i] = int(unique[index])
index = index + 1
initRoundFlag = False
print(unique, inits)
# -
# Visualize the sampled states and the 10 adaptive seeds selected for the next round sampling.
sampled_saxs = [ np.transpose(SAXS)[1][unique[i]] for i in range(len(unique)) ]
sampled_dist = [ np.transpose(EC)[unique[i]] for i in range(len(unique)) ]
sampled_pop = [ msm.populations_[unique[i]]/np.max(msm.populations_)*200 for i in range(len(unique)) ]
init_saxs = [ np.transpose(SAXS)[1][inits[i]] for i in range(len(inits)) ]
init_dist = [ np.transpose(EC)[inits[i]] for i in range(len(inits)) ]
init_pop = [ msm.populations_[inits[i]]/np.max(msm.populations_)*200 for i in range(len(inits)) ]
fig, ax = plt.subplots()
ax.scatter(np.transpose(SAXS)[1], np.transpose(EC), lw=0.3, color="#f46855", \
s=msm.populations_/np.max(msm.populations_)*200, zorder=1, label='all')
ax.scatter(np.transpose(SAXS)[1][init], np.transpose(EC)[init], lw=0.3, color="blue", \
s=msm.populations_[init]/np.max(msm.populations_)*200, zorder=1, label='initial')
ax.scatter(np.transpose(SAXS)[1][end], np.transpose(EC)[end], lw=0.3, color="green", \
s=msm.populations_[end]/np.max(msm.populations_)*2500, marker='*', zorder=1, label='target')
ax.scatter(sampled_saxs, sampled_dist, lw=0.3, color="grey", \
s=sampled_pop, zorder=2, label="sampled states")
ax.scatter(init_saxs, init_dist, lw=0.3, color="purple", \
s=init_pop, zorder=2, label="adaptive seeds")
ax.set_xlim(1,10000)
ax.set_xscale("log")
ax.set_ylim(0,200)
ax.set_yticks((0,50,100,150,200))
ax.set(xlabel=r'SAXS discrepancy $\chi$$^{2}$', ylabel='Average residue pair distance (${\AA}$)')
ax.legend(loc=2, ncol=1, frameon=False, prop=fontP, numpoints=1, scatterpoints=1)
ax.tick_params(axis='x', which='minor', top='off')
# #### Step 4: Starting 10 parallel simulations in the second round from the 10 picked cluster states.
# +
if not initRoundFlag:
inits_iter = inits
inits = np.zeros(Num, dtype=int)
trajs_rnd = []
for i in range(Num):
traj = run(inits_iter[i], msm, step)
trajs[i].append(traj)
trajs_rnd.append(traj)
unique = np.unique(trajs)
# -
# Visualize the sampled states after the second round of sampling.
sampled_saxs = [ np.transpose(SAXS)[1][unique[i]] for i in range(len(unique)) ]
sampled_dist = [ np.transpose(EC)[unique[i]] for i in range(len(unique)) ]
sampled_pop = [ msm.populations_[unique[i]]/np.max(msm.populations_)*200 for i in range(len(unique)) ]
fig, ax = plt.subplots()
ax.scatter(np.transpose(SAXS)[1], np.transpose(EC), lw=0.3, color="#f46855", \
s=msm.populations_/np.max(msm.populations_)*200, zorder=1, label='all')
ax.scatter(np.transpose(SAXS)[1][init], np.transpose(EC)[init], lw=0.3, color="blue", \
s=msm.populations_[init]/np.max(msm.populations_)*200, zorder=1, label='initial')
ax.scatter(np.transpose(SAXS)[1][end], np.transpose(EC)[end], lw=0.3, color="green", \
s=msm.populations_[end]/np.max(msm.populations_)*2500, marker='*', zorder=1, label='target')
ax.scatter(sampled_saxs, sampled_dist, lw=0.3, color="grey", \
s=sampled_pop, zorder=2, label="sampled states")
ax.set_xlim(1,10000)
ax.set_xscale("log")
ax.set_ylim(0,200)
ax.set_yticks((0,50,100,150,200))
ax.set(xlabel=r'SAXS discrepancy $\chi$$^{2}$', ylabel='Average residue pair distance (${\AA}$)')
ax.legend(loc=2, ncol=1, frameon=False, prop=fontP, numpoints=1, scatterpoints=1)
ax.tick_params(axis='x', which='minor', top='off')
# ### Perform adaptive sampling in kinetic Monte Carlo simulations guided by hybrid information
# In the step 3 as noted above, additional information such as the interfacial residue pair distance can be combined with SAXS to choose the seeding clusters for adaptive sampling. Below is the example of choosing 5 states with the minimal SAXS discrepancy scores (adaptive seeds: set A) and 5 states with the minimal residue pair distances (adaptive seeds: set B), respectively.
# +
unique = np.unique(trajs)
SAXS_diff = [ saxs_dict.get(state) for state in unique ]
EC_diff = [ ec_dict.get(state) for state in unique ]
if np.shape(unique)[0] < 10:
for i in range(np.shape(unique)[0]):
inits[i] = int(unique[i])
for i in range(10-np.shape(unique)[0]):
inits[np.shape(unique)[0] + i] = int(np.random.choice(unique,1))
if np.shape(unique)[0] == 10:
for i in range(np.shape(unique)[0]):
inits[i] = int(unique[i])
if np.shape(unique)[0] > 10:
sorted_SAXS_diff = np.unique(SAXS_diff)
for i in range(int(Num/2)):
index = 0
for saxs in SAXS_diff:
if saxs == sorted_SAXS_diff[i]:
inits[i] = int(unique[index])
index = index + 1
sorted_EC_diff = np.unique(EC_diff)
for i in range(int(Num/2),Num):
index = 0
for ec in EC_diff:
if ec == sorted_EC_diff[i-int(Num/2)]:
inits[i] = int(unique[index])
index = index + 1
# -
# Visualize the sampled states and the adaptive seeds (set A and B) selected for the next round sampling.
sampled_saxs = [ np.transpose(SAXS)[1][unique[i]] for i in range(len(unique)) ]
sampled_dist = [ np.transpose(EC)[unique[i]] for i in range(len(unique)) ]
sampled_pop = [ msm.populations_[unique[i]]/np.max(msm.populations_)*200 for i in range(len(unique)) ]
init_saxs = [ np.transpose(SAXS)[1][inits[i]] for i in range(len(inits)) ]
init_dist = [ np.transpose(EC)[inits[i]] for i in range(len(inits)) ]
init_pop = [ msm.populations_[inits[i]]/np.max(msm.populations_)*200 for i in range(len(inits)) ]
fig, ax = plt.subplots()
ax.scatter(np.transpose(SAXS)[1], np.transpose(EC), lw=0.3, color="#f46855", \
s=msm.populations_/np.max(msm.populations_)*200, zorder=1, label='all')
ax.scatter(np.transpose(SAXS)[1][init], np.transpose(EC)[init], lw=0.3, color="blue", \
s=msm.populations_[init]/np.max(msm.populations_)*200, zorder=1, label='initial')
ax.scatter(np.transpose(SAXS)[1][end], np.transpose(EC)[end], lw=0.3, color="green", \
s=msm.populations_[end]/np.max(msm.populations_)*2500, marker='*', zorder=1, label='target')
ax.scatter(sampled_saxs, sampled_dist, lw=0.3, color="grey", \
s=sampled_pop, zorder=2, label="sampled states")
ax.scatter(init_saxs[0:int(Num/2)], init_dist[0:int(Num/2)], lw=0.3, color="purple", \
s=init_pop, zorder=2, label="adaptive seeds: set A")
ax.scatter(init_saxs[int(Num/2):Num], init_dist[int(Num/2):Num], lw=0.3, color="yellow", \
s=init_pop, zorder=2, label="adaptive seeds: set B")
ax.set_xlim(1,10000)
ax.set_xscale("log")
ax.set_ylim(0,200)
ax.set_yticks((0,50,100,150,200))
ax.set(xlabel=r'SAXS discrepancy $\chi$$^{2}$', ylabel='Average residue pair distance (${\AA}$)')
ax.legend(loc=2, ncol=1, frameon=False, prop=fontP, numpoints=1, scatterpoints=1)
ax.tick_params(axis='x', which='minor', top='off')
print(inits)
| example/MoaD-MoaE-Association.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Two sum
#
# Given an array of integers, return **indices** of the two numbers such that they add up to a specific target.
# You may assume that each input would have *exactly* one solution, and you may not use the same element twice.
def two_sum(nums, target):
# number to index mapping table
mapping = {num : ix for ix, num in enumerate(nums)}
for ix, n in enumerate(nums):
diff = target - n
if diff in nums:
if mapping[diff] != ix:
return ix, mapping[diff]
nums = [3, 2, 4, 5]
target = 6
two_sum(nums, target)
# ## Reverse Integer
#
# Not working !!!
def rev_int(x):
rev = 0
INT_MAX = 2**31 - 1
INT_MIN = -2**31
while x !=0:
pop = x % 10
x = x//10
if (rev > INT_MAX//10) or (rev == INT_MAX//10 and pop > 6):
return 0
if (rev < INT_MIN//10) or (rev == INT_MIN//10 and pop < -8):
return 0
rev = rev*10 + pop
return rev
rev_int(-123)
# ## Palindrome Number
#
# Discuss all cases:
# - Negative integers are not palindromes
# - If the last digit is 0, palindrome should also start with 0. Which is possible only in the case of 0
# - We should care about INT_MAX overflow
#
# Algorithm:
# - In order to NOT interfere with the
def palindrome(num):
if num == 0:
return True
if num < 0 or num%10 == 0:
return False
rev = 0
length = len(str(num))
half = length // 2
for _ in range(half):
pop = num%10
num = num//10
rev = rev*10 + pop
if length%2 == 0:
if rev == num:
return True
else:
return False
else:
if rev == num//10:
return True
else:
return False
palindrome(763182)
# ### Remove Duplicates from Sorted Array (Lists)
#
# Discuss:
# - Arrays contains nums = integers only
# - Don't use extra memory. Allowance O(1) only
#
# Algorithm:
# - In-place algorithm means: manipulate/modify object without copyting it to another variable
# - **i** stays and waits until **j** goes and find the element which is different from nums[i]
# - Once it finds it, we copy that elements to the next slot: nums[i+1] (so i++)
# - **j** continues search, now it tries to find the element which is different from **i**'s new spot
# - Continue until j reaches the end of array and return the **i**! >> Number of different integers!
# +
def removeDuplicates(nums):
if len(nums) == 0:
return 0
i = 0
length = len(nums)
for j in range(1, length):
if nums[i] != nums[j]:
i += 1
nums[i] = nums[j]
return i+1 #since i is index
#---------------------------------------------------------------------------------------------------------------#
def removeDuplicates_long(arr):
from collections import Counter
unique_nums = Counter(arr)
for item, value in unique_nums.items():
if value > 1:
temp = value
while temp != 1:
arr.remove(item)
temp -= 1
print(arr)
print(len(arr))
# -
removeDuplicates([1,2,3,4,4])
# ### Merge Two Sorted Lists
#
# Discuss:
# - Done with ListNodes, where given Listnode is a head node of the linked list!
#
# Algorithm:
# - If one of the linked lists is empty, return the other one!
# - Do a recursive solution: call the function everytime for the new search!
# - Try alterbative approach NOT using recursive method!
# +
# write list nodes here!!!
def mergeTwoLists(self, l1: ListNode, l2: ListNode) -> ListNode:
if l1 is None: #pointing to None
return l2
if l2 is None:
return l1
if l1.val <= l2.val:
l1.next = self.mergeTwoLists(l1.next, l2)
return l1
else:
l2.next = self.mergeTwoLists(l2.next, l1)
return l2
# -
# ### Merge Sorted Array
#
# Discuss:
# - Without using stacks
# - in-place modification, nothing to return
#
# Algorithm:
# - Compare the max length with its content length
# -sort the first m+n elements together, and replace nums1 by it
# +
## too long -- later
def merge(nums1, m, nums2, n):
if m == 0:
nums1 = nums2
elif n != 0:
i = m
j = n
nums1_copy = nums1[:m]
for k in reversed(range(m+n)):
if nums1_copy[i-1] >= nums2[j-1]:
temp = nums1_copy[i-1]
i -= 1
else:
temp = nums2[j-1]
j -= 1
print(temp, i, j, k)
if (j == -1):
break
elif (i == -1):
nums1[:k] = nums2[:j]
else:
nums1[k] = temp
print(nums1)
return nums1
def merge_sorted(nums1, m, nums2, n):
nums1[:] = sorted(nums1[1:m] + nums2)
return(nums1)
merge([2,0], 1, [1], 1)
# -
# ### Square root of a number without using any math:
#
# Algorithm:
# - Use Newton's approximation: Taylor one series expansion
# +
def mySqrt(x: int) -> int:
if x < 1:
return 0
elif x < 4:
return 1
elif x == 4:
return 2
else:
res_prev = x
while True:
prev_temp = res_prev
res_next = 1/2*(prev_temp + x/prev_temp)
print('prev is %d', res_prev)
print('next is %d', res_next)
if res_prev - res_next < 0.01:
break
else:
temp_next = res_next
res_prev = temp_next
return int(res_next)
mySqrt(8)
# -
# ### Implmenet strStr()
#
# Discussion:
# - qjeq
#
# Algorithm:
# - Start from the edge cases
# +
def strStr(self, haystack: str, needle: str) -> int:
if not haystack
# -
'banana'.count('na')
print(not 'a')
| exercises_easy_all_categories.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# # Azure Notebook Climate Change Analysis
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from sklearn.linear_model import LinearRegression
import seaborn as sns; sns.set()
yearsBase, meanBase = np.loadtxt('5-year-mean-1951-1980.csv', delimiter=',', usecols=(0, 1), unpack=True)
years, mean = np.loadtxt('5-year-mean-1882-2014.csv', delimiter=',', usecols=(0, 1), unpack=True)
# ## Create Scatter Plot
plt.scatter(yearsBase, meanBase)
plt.title('scatter plot of mean temp difference vs year')
plt.xlabel('years', fontsize=12)
plt.ylabel('mean temp difference', fontsize=12)
plt.show()
# ## Perform linear regression
# +
# Creates a linear regression from the data points
m,b = np.polyfit(yearsBase, meanBase, 1)
# This is a simple y = mx + b line function
def f(x):
return m*x + b
# This generates the same scatter plot as before, but adds a line plot using the function above
plt.scatter(yearsBase, meanBase)
plt.plot(yearsBase, f(yearsBase))
plt.title('scatter plot of mean temp difference vs year')
plt.xlabel('years', fontsize=12)
plt.ylabel('mean temp difference', fontsize=12)
plt.show()
# Prints text to the screen showing the computed values of m and b
print(' y = {0} * x + {1}'.format(m, b))
plt.show()
# -
# ## Perform linear regression with scikit-learn
yearsBase.shape
yearsBase[:, np.newaxis].shape
# +
# Pick the Linear Regression model and instantiate it
model = LinearRegression(fit_intercept=True)
# Fit/build the model
model.fit(yearsBase[:, np.newaxis], meanBase)
mean_predicted = model.predict(yearsBase[:, np.newaxis])
# Generate a plot like the one in the previous exercise
plt.scatter(yearsBase, meanBase)
plt.plot(yearsBase, mean_predicted)
plt.title('scatter plot of mean temp difference vs year')
plt.xlabel('years', fontsize=12)
plt.ylabel('mean temp difference', fontsize=12)
plt.show()
print(' y = {0} * x + {1}'.format(model.coef_[0], model.intercept_))
# -
# ## Perform linear regression with Seaborn
plt.scatter(years, mean)
plt.title('scatter plot of mean temp difference vs year')
plt.xlabel('years', fontsize=12)
plt.ylabel('mean temp difference', fontsize=12)
sns.regplot(yearsBase, meanBase)
plt.show()
| Azure/notebooks/climatechange.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dendrograms
# Visualization of dendrograms as SVG images.
# + pycharm={"is_executing": false}
from IPython.display import SVG
# + pycharm={"is_executing": false}
import numpy as np
# + pycharm={"is_executing": false}
from sknetwork.data import karate_club, painters, movie_actor
from sknetwork.hierarchy import Paris
from sknetwork.visualization import svg_graph, svg_digraph, svg_bigraph
from sknetwork.visualization import svg_dendrogram
# -
# ## Graphs
# + pycharm={"is_executing": false}
graph = karate_club(metadata=True)
adjacency = graph.adjacency
position = graph.position
labels = graph.labels
# + pycharm={"is_executing": false}
# graph
image = svg_graph(adjacency, position, labels=labels)
SVG(image)
# -
# hierarchical clustering
paris = Paris()
dendrogram = paris.fit_transform(adjacency)
# visualization
image = svg_dendrogram(dendrogram)
SVG(image)
# add names, set colors
n = adjacency.shape[0]
image = svg_dendrogram(dendrogram, names=np.arange(n), n_clusters=5, color='gray')
SVG(image)
# export
svg_dendrogram(dendrogram, filename='dendrogram_karate_club')
# ## Directed graphs
# + pycharm={"is_executing": false}
graph = painters(metadata=True)
adjacency = graph.adjacency
names = graph.names
position = graph.position
# + pycharm={"is_executing": false}
# graph
image = svg_digraph(adjacency, position, names)
SVG(image)
# -
# hierarchical clustering
paris = Paris()
dendrogram = paris.fit_transform(adjacency)
# visualization
image = svg_dendrogram(dendrogram, names, n_clusters=3, rotate=True)
SVG(image)
# ## Bipartite graphs
# + pycharm={"is_executing": false}
graph = movie_actor(metadata=True)
biadjacency = graph.biadjacency
names_row = graph.names_row
names_col = graph.names_col
# + pycharm={"is_executing": false}
# graph
image = svg_bigraph(biadjacency, names_row, names_col)
SVG(image)
# -
# hierarchical clustering
paris = Paris()
paris.fit(biadjacency)
dendrogram_row = paris.dendrogram_row_
dendrogram_col = paris.dendrogram_col_
dendrogram_full = paris.dendrogram_full_
# visualization
image = svg_dendrogram(dendrogram_row, names_row, n_clusters=3, rotate=True)
SVG(image)
image = svg_dendrogram(dendrogram_col, names_col, n_clusters=3, rotate=True)
SVG(image)
| docs/tutorials/visualization/dendrograms.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-1:081325390199:image/datascience-1.0
# ---
# # Training and Deploying a BlazingText model
# <img align="left" width="130" src="https://raw.githubusercontent.com/PacktPublishing/Amazon-SageMaker-Cookbook/master/Extra/cover-small-padded.png"/>
#
# This notebook contains the code to help readers work through one of the recipes of the book [Machine Learning with Amazon SageMaker Cookbook: 80 proven recipes for data scientists and developers to perform ML experiments and deployments](https://www.amazon.com/Machine-Learning-Amazon-SageMaker-Cookbook/dp/1800567030)
# ### How to do it...
import sagemaker
from sagemaker import get_execution_role
import json
import boto3
session = sagemaker.Session()
role = get_execution_role()
region_name = boto3.Session().region_name
# %store -r s3_bucket
# %store -r prefix
s3_train_data = 's3://{}/{}/input/{}'.format(
s3_bucket,
prefix,
"synthetic.train.txt"
)
s3_validation_data = 's3://{}/{}/input/{}'.format(
s3_bucket,
prefix,
"synthetic.validation.txt"
)
s3_output_location = 's3://{}/{}/output'.format(
s3_bucket,
prefix
)
# +
from sagemaker.image_uris import retrieve
container = retrieve(
"blazingtext",
region_name,
"1"
)
# -
estimator = sagemaker.estimator.Estimator(
container,
role,
instance_count=1,
instance_type='ml.c4.xlarge',
input_mode= 'File',
output_path=s3_output_location,
sagemaker_session=session
)
estimator.set_hyperparameters(
mode="supervised",
min_count=2
)
# +
from sagemaker.inputs import TrainingInput
train_data = TrainingInput(
s3_train_data,
distribution='FullyReplicated',
content_type='text/plain',
s3_data_type='S3Prefix'
)
validation_data = TrainingInput(
s3_validation_data,
distribution='FullyReplicated',
content_type='text/plain',
s3_data_type='S3Prefix'
)
# -
data_channels = {
'train': train_data,
'validation': validation_data
}
# %%time
estimator.fit(
inputs=data_channels,
logs=True
)
endpoint = estimator.deploy(
initial_instance_count = 1,
instance_type = 'ml.r5.large'
)
# +
sentences = [
"that is bad",
"the apple tastes good",
"i would recommend it to my friends"
]
payload = {"instances" : sentences}
# +
from sagemaker.serializers import JSONSerializer
endpoint.serializer = JSONSerializer()
response = endpoint.predict(payload)
predictions = json.loads(response)
print(json.dumps(predictions, indent=2))
# -
tn = estimator.latest_training_job.name
training_job_name = tn
# %store training_job_name
# +
# endpoint.delete_endpoint()
| Chapter08/03 - Training and Deploying a BlazingText model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.12 ('tensorflow26')
# language: python
# name: python3
# ---
import os
import json
from utils import load_datasets, load_target
import models
from models.tuning import beyesian_optimization
from models.evaluation import cross_validation_score
import json
import numpy as np
config = json.load(open('./config/default.json'))
# X_train, X_test = load_datasets(["Age", "AgeSplit", "EducationNum"])
X_train, X_test = load_datasets(config['features'])
y_train = load_target('Y')
lgbm1 = models.Lgbm(json.load(open('./config/Lgbm-depth-5.json')))
lgbm2 = models.Lgbm(json.load(open('./config/Lgbm-depth-15.json')))
lgbm3 = models.Lgbm(json.load(open('./config/Lgbm-depth-inf.json')))
nn1 = models.NN(json.load(open('./config/NN-shallow.json')))
nn2 = models.NN(json.load(open('./config/NN-deep.json')))
rf = models.RandomForest(json.load(open('./config/RandomForest.json')))
ert = models.RandomForest(json.load(open('./config/ERT.json')))
model_list = [lgbm1, lgbm2, lgbm3, nn1, nn2, rf, ert]
# +
from models.evaluation import stacking
oof, y_preds = stacking(model_list, X_train, y_train, X_test, 1)
# -
y_pred = [np.argmax(y_oofs, axis=1) for i in y_preds]
y_pred = sum(y_pred) / len(y_pred)
y_pred = (y_pred > 0.5).astype(int)
import pandas as pd
sample_submit = pd.read_csv(
'./data/input/sample_submit.csv', names=['id', 'Y'])
sample_submit['Y'] = y_pred
sample_submit.to_csv('./data/output/2022-2-19-submit.csv', header=False, index=False)
| tabular-playground-series-feb-2022/stacking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 2. Topic Modeling with NMF and SVD
# Topic modeling is a great way to get started with matrix factorizations. We start with a **term-document matrix**:
#
# <img src="images/document_term.png" alt="term-document matrix" style="width: 80%"/>
# (source: [Introduction to Information Retrieval](http://player.slideplayer.com/15/4528582/#))
#
# We can decompose this into one tall thin matrix times one wide short matrix (possibly with a diagonal matrix in between).
#
# Notice that this representation does not take into account word order or sentence structure. It's an example of a **bag of words** approach.
# ### Motivation
# Consider the most extreme case - reconstructing the matrix using an outer product of two vectors. Clearly, in most cases we won't be able to reconstruct the matrix exactly. But if we had one vector with the relative frequency of each vocabulary word out of the total word count, and one with the average number of words per document, then that outer product would be as close as we can get.
#
# Now consider increasing that matrices to two columns and two rows. The optimal decomposition would now be to cluster the documents into two groups, each of which has as different a distribution of words as possible to each other, but as similar as possible amongst the documents in the cluster. We will call those two groups "topics". And we would cluster the words into two groups, based on those which most frequently appear in each of the topics.
# ### Today
# We'll take a dataset of documents in several different categories, and find topics (consisting of groups of words) for them. Knowing the actual categories helps us evaluate if the topics we find make sense.
#
# We will try this with two different matrix factorizations: **Singular Value Decomposition (SVD)** and **Non-negative Matrix Factorization (NMF)**
import numpy as np
from sklearn.datasets import fetch_20newsgroups
from sklearn import decomposition
from scipy import linalg
import matplotlib.pyplot as plt
# %matplotlib inline
np.set_printoptions(suppress=True)
# ## Additional Resources
# - [Data source](http://scikit-learn.org/stable/datasets/twenty_newsgroups.html): Newsgroups are discussion groups on Usenet, which was popular in the 80s and 90s before the web really took off. This dataset includes 18,000 newsgroups posts with 20 topics.
# - [Chris Manning's book chapter](https://nlp.stanford.edu/IR-book/pdf/18lsi.pdf) on matrix factorization and LSI
# - Scikit learn [truncated SVD LSI details](http://scikit-learn.org/stable/modules/decomposition.html#lsa)
#
# ### Other Tutorials
# - [Scikit-Learn: Out-of-core classification of text documents](http://scikit-learn.org/stable/auto_examples/applications/plot_out_of_core_classification.html): uses [Reuters-21578](https://archive.ics.uci.edu/ml/datasets/reuters-21578+text+categorization+collection) dataset (Reuters articles labeled with ~100 categories), HashingVectorizer
# - [Text Analysis with Topic Models for the Humanities and Social Sciences](https://de.dariah.eu/tatom/index.html): uses [British and French Literature dataset](https://de.dariah.eu/tatom/datasets.html) of <NAME>en, Charlotte Bronte, <NAME>, and more
# ## Set up data
# Scikit Learn comes with a number of built-in datasets, as well as loading utilities to load several standard external datasets. This is a [great resource](http://scikit-learn.org/stable/datasets/), and the datasets include Boston housing prices, face images, patches of forest, diabetes, breast cancer, and more. We will be using the newsgroups dataset.
#
# Newsgroups are discussion groups on Usenet, which was popular in the 80s and 90s before the web really took off. This dataset includes 18,000 newsgroups posts with 20 topics.
categories = ['alt.atheism', 'talk.religion.misc', 'comp.graphics', 'sci.space']
remove = ('headers', 'footers', 'quotes')
newsgroups_train = fetch_20newsgroups(subset='train', categories=categories, remove=remove)
newsgroups_test = fetch_20newsgroups(subset='test', categories=categories, remove=remove)
newsgroups_train.filenames.shape, newsgroups_train.target.shape
# Let's look at some of the data. Can you guess which category these messages are in?
print("\n".join(newsgroups_train.data[:3]))
# hint: definition of *perijove* is the point in the orbit of a satellite of Jupiter nearest the planet's center
np.array(newsgroups_train.target_names)[newsgroups_train.target[:3]]
# The target attribute is the integer index of the category.
newsgroups_train.target[:10]
num_topics, num_top_words = 6, 8
# Next, scikit learn has a method that will extract all the word counts for us.
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
vectorizer = CountVectorizer(stop_words='english')
vectors = vectorizer.fit_transform(newsgroups_train.data).todense() # (documents, vocab)
vectors.shape #, vectors.nnz / vectors.shape[0], row_means.shape
print(len(newsgroups_train.data), vectors.shape)
vocab = np.array(vectorizer.get_feature_names())
vocab.shape
vocab[7000:7020]
# ## Singular Value Decomposition (SVD)
# "SVD is not nearly as famous as it should be." - <NAME>
# We would clearly expect that the words that appear most frequently in one topic would appear less frequently in the other - otherwise that word wouldn't make a good choice to separate out the two topics. Therefore, we expect the topics to be **orthogonal**.
#
# The SVD algorithm factorizes a matrix into one matrix with **orthogonal columns** and one with **orthogonal rows** (along with a diagonal matrix, which contains the **relative importance** of each factor).
#
# <img src="images/svd_fb.png" alt="" style="width: 80%"/>
# (source: [Facebook Research: Fast Randomized SVD](https://research.fb.com/fast-randomized-svd/))
#
# SVD is an **exact decomposition**, since the matrices it creates are big enough to fully cover the original matrix. SVD is extremely widely used in linear algebra, and specifically in data science, including:
#
# - semantic analysis
# - collaborative filtering/recommendations ([winning entry for Netflix Prize](https://datajobs.com/data-science-repo/Recommender-Systems-%5BNetflix%5D.pdf))
# - calculate Moore-Penrose pseudoinverse
# - data compression
# - principal component analysis (will be covered later in course)
# %time U, s, Vh = linalg.svd(vectors, full_matrices=False)
print(U.shape, s.shape, Vh.shape)
# Confirm this is a decomposition of the input.
# #### Answer
#Exercise: confrim that U, s, Vh is a decomposition of the var Vectors
# Confirm that U, V are orthonormal
# #### Answer
#Exercise: Confirm that U, Vh are orthonormal
# #### Topics
# What can we say about the singular values s?
plt.plot(s);
plt.plot(s[:10])
# +
num_top_words=8
def show_topics(a):
top_words = lambda t: [vocab[i] for i in np.argsort(t)[:-num_top_words-1:-1]]
topic_words = ([top_words(t) for t in a])
return [' '.join(t) for t in topic_words]
# -
show_topics(Vh[:10])
# We get topics that match the kinds of clusters we would expect! This is despite the fact that this is an **unsupervised algorithm** - which is to say, we never actually told the algorithm how our documents are grouped.
# We will return to SVD in **much more detail** later. For now, the important takeaway is that we have a tool that allows us to exactly factor a matrix into orthogonal columns and orthogonal rows.
# ## Non-negative Matrix Factorization (NMF)
# + [markdown] heading_collapsed=true
# #### Motivation
# + [markdown] hidden=true
# <img src="images/face_pca.png" alt="PCA on faces" style="width: 80%"/>
#
# (source: [NMF Tutorial](http://perso.telecom-paristech.fr/~essid/teach/NMF_tutorial_ICME-2014.pdf))
#
# A more interpretable approach:
#
# <img src="images/face_outputs.png" alt="NMF on Faces" style="width: 80%"/>
#
# (source: [NMF Tutorial](http://perso.telecom-paristech.fr/~essid/teach/NMF_tutorial_ICME-2014.pdf))
# + [markdown] heading_collapsed=true
# #### Idea
# + [markdown] hidden=true
# Rather than constraining our factors to be *orthogonal*, another idea would to constrain them to be *non-negative*. NMF is a factorization of a non-negative data set $V$: $$ V = W H$$ into non-negative matrices $W,\; H$. Often positive factors will be **more easily interpretable** (and this is the reason behind NMF's popularity).
#
# <img src="images/face_nmf.png" alt="NMF on faces" style="width: 80%"/>
#
# (source: [NMF Tutorial](http://perso.telecom-paristech.fr/~essid/teach/NMF_tutorial_ICME-2014.pdf))
#
# Nonnegative matrix factorization (NMF) is a non-exact factorization that factors into one skinny positive matrix and one short positive matrix. NMF is NP-hard and non-unique. There are a number of variations on it, created by adding different constraints.
# + [markdown] heading_collapsed=true
# #### Applications of NMF
# + [markdown] hidden=true
# - [Face Decompositions](http://scikit-learn.org/stable/auto_examples/decomposition/plot_faces_decomposition.html#sphx-glr-auto-examples-decomposition-plot-faces-decomposition-py)
# - [Collaborative Filtering, eg movie recommendations](http://www.quuxlabs.com/blog/2010/09/matrix-factorization-a-simple-tutorial-and-implementation-in-python/)
# - [Audio source separation](https://pdfs.semanticscholar.org/cc88/0b24791349df39c5d9b8c352911a0417df34.pdf)
# - [Chemistry](http://ieeexplore.ieee.org/document/1532909/)
# - [Bioinformatics](https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-015-0485-4) and [Gene Expression](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2623306/)
# - Topic Modeling (our problem!)
#
# <img src="images/nmf_doc.png" alt="NMF on documents" style="width: 80%"/>
#
# (source: [NMF Tutorial](http://perso.telecom-paristech.fr/~essid/teach/NMF_tutorial_ICME-2014.pdf))
# + [markdown] hidden=true
# **More Reading**:
#
# - [The Why and How of Nonnegative Matrix Factorization](https://arxiv.org/pdf/1401.5226.pdf)
# + [markdown] heading_collapsed=true
# ### NMF from sklearn
# + [markdown] hidden=true
# First, we will use [scikit-learn's implementation of NMF](http://scikit-learn.org/stable/modules/generated/sklearn.decomposition.NMF.html):
# + hidden=true
m,n=vectors.shape
d=5 # num topics
# + hidden=true
clf = decomposition.NMF(n_components=d, random_state=1)
W1 = clf.fit_transform(vectors)
H1 = clf.components_
# + hidden=true
show_topics(H1)
# + [markdown] heading_collapsed=true
# ### TF-IDF
# + [markdown] hidden=true
# [Topic Frequency-Inverse Document Frequency](http://www.tfidf.com/) (TF-IDF) is a way to normalize term counts by taking into account how often they appear in a document, how long the document is, and how commmon/rare the term is.
#
# TF = (# occurrences of term t in document) / (# of words in documents)
#
# IDF = log(# of documents / # documents with term t in it)
# + hidden=true
vectorizer_tfidf = TfidfVectorizer(stop_words='english')
vectors_tfidf = vectorizer_tfidf.fit_transform(newsgroups_train.data) # (documents, vocab)
# + hidden=true
W1 = clf.fit_transform(vectors_tfidf)
H1 = clf.components_
# + hidden=true
show_topics(H1)
# + hidden=true
plt.plot(clf.components_[0])
# + hidden=true
clf.reconstruction_err_
# + [markdown] heading_collapsed=true
# ### NMF in summary
# + [markdown] hidden=true
# Benefits: Fast and easy to use!
#
# Downsides: took years of research and expertise to create
# + [markdown] hidden=true
# Notes:
# - For NMF, matrix needs to be at least as tall as it is wide, or we get an error with fit_transform
# - Can use df_min in CountVectorizer to only look at words that were in at least k of the split texts
# + [markdown] heading_collapsed=true
# ### NMF from scratch in numpy, using SGD
# + [markdown] hidden=true
# #### Gradient Descent
# + [markdown] hidden=true
# The key idea of standard **gradient descent**:
# 1. Randomly choose some weights to start
# 2. Loop:
# - Use weights to calculate a prediction
# - Calculate the derivative of the loss
# - Update the weights
# 3. Repeat step 2 lots of times. Eventually we end up with some decent weights.
#
# **Key**: We want to decrease our loss and the derivative tells us the direction of **steepest descent**.
#
# Note that *loss*, *error*, and *cost* are all terms used to describe the same thing.
#
# Let's take a look at the [Gradient Descent Intro notebook](gradient-descent-intro.ipynb) (originally from the [fast.ai deep learning course](https://github.com/fastai/courses)).
# + [markdown] heading_collapsed=true hidden=true
# #### Stochastic Gradient Descent (SGD)
# + [markdown] hidden=true
# **Stochastic gradient descent** is an incredibly useful optimization method (it is also the heart of deep learning, where it is used for backpropagation).
#
# For *standard* gradient descent, we evaluate the loss using **all** of our data which can be really slow. In *stochastic* gradient descent, we evaluate our loss function on just a sample of our data (sometimes called a *mini-batch*). We would get different loss values on different samples of the data, so this is *why it is stochastic*. It turns out that this is still an effective way to optimize, and it's much more efficient!
#
# We can see how this works in this [excel spreadsheet](graddesc.xlsm) (originally from the [fast.ai deep learning course](https://github.com/fastai/courses)).
#
# **Resources**:
# - [SGD Lecture from <NAME>'s Coursera ML course](https://www.coursera.org/learn/machine-learning/lecture/DoRHJ/stochastic-gradient-descent)
# - <a href="http://wiki.fast.ai/index.php/Stochastic_Gradient_Descent_(SGD)">fast.ai wiki page on SGD</a>
# - [Gradient Descent For Machine Learning](http://machinelearningmastery.com/gradient-descent-for-machine-learning/) (<NAME>- Machine Learning Mastery)
# - [An overview of gradient descent optimization algorithms](http://sebastianruder.com/optimizing-gradient-descent/)
# + [markdown] heading_collapsed=true hidden=true
# #### Applying SGD to NMF
# + [markdown] hidden=true
# **Goal**: Decompose $V\;(m \times n)$ into $$V \approx WH$$ where $W\;(m \times d)$ and $H\;(d \times n)$, $W,\;H\;>=\;0$, and we've minimized the Frobenius norm of $V-WH$.
#
# **Approach**: We will pick random positive $W$ & $H$, and then use SGD to optimize.
# + [markdown] hidden=true
# **To use SGD, we need to know the gradient of the loss function.**
#
# **Sources**:
# - Optimality and gradients of NMF: http://users.wfu.edu/plemmons/papers/chu_ple.pdf
# - Projected gradients: https://www.csie.ntu.edu.tw/~cjlin/papers/pgradnmf.pdf
# + hidden=true
lam=1e3
lr=1e-2
m, n = vectors_tfidf.shape
# + hidden=true
W1 = clf.fit_transform(vectors)
H1 = clf.components_
# + hidden=true
show_topics(H1)
# + hidden=true
mu = 1e-6
def grads(M, W, H):
R = W@H-M
return R@H.T + penalty(W, mu)*lam, W.T@R + penalty(H, mu)*lam # dW, dH
# + hidden=true
def penalty(M, mu):
return np.where(M>=mu,0, np.min(M - mu, 0))
# + hidden=true
def upd(M, W, H, lr):
dW,dH = grads(M,W,H)
W -= lr*dW; H -= lr*dH
# + hidden=true
def report(M,W,H):
print(np.linalg.norm(M-W@H), W.min(), H.min(), (W<0).sum(), (H<0).sum())
# + hidden=true
W = np.abs(np.random.normal(scale=0.01, size=(m,d)))
H = np.abs(np.random.normal(scale=0.01, size=(d,n)))
# + hidden=true
report(vectors_tfidf, W, H)
# + hidden=true
upd(vectors_tfidf,W,H,lr)
# + hidden=true
report(vectors_tfidf, W, H)
# + hidden=true
for i in range(50):
upd(vectors_tfidf,W,H,lr)
if i % 10 == 0: report(vectors_tfidf,W,H)
# + hidden=true
show_topics(H)
# + [markdown] hidden=true
# This is painfully slow to train! Lots of parameter fiddling and still slow to train (or explodes).
# + [markdown] heading_collapsed=true
# ### PyTorch
# + [markdown] hidden=true
# [PyTorch](http://pytorch.org/) is a Python framework for tensors and dynamic neural networks with GPU acceleration. Many of the core contributors work on Facebook's AI team. In many ways, it is similar to Numpy, only with the increased parallelization of using a GPU.
#
# From the [PyTorch documentation](http://pytorch.org/tutorials/beginner/blitz/tensor_tutorial.html):
#
# <img src="images/what_is_pytorch.png" alt="pytorch" style="width: 80%"/>
#
# **Further learning**: If you are curious to learn what *dynamic* neural networks are, you may want to watch [this talk](https://www.youtube.com/watch?v=Z15cBAuY7Sc) by <NAME>, Facebook AI researcher and core PyTorch contributor.
#
# If you want to learn more PyTorch, you can try this [tutorial](http://pytorch.org/tutorials/beginner/deep_learning_60min_blitz.html) or this [learning by examples](http://pytorch.org/tutorials/beginner/pytorch_with_examples.html).
# + [markdown] hidden=true
# **Note about GPUs**: If you are not using a GPU, you will need to remove the `.cuda()` from the methods below. GPU usage is not required for this course, but I thought it would be of interest to some of you. To learn how to create an AWS instance with a GPU, you can watch the [fast.ai setup lesson](http://course.fast.ai/lessons/aws.html).
# + hidden=true
import torch
import torch.cuda as tc
from torch.autograd import Variable
# + hidden=true
def V(M): return Variable(M, requires_grad=True)
# + hidden=true
v=vectors_tfidf.todense()
# + hidden=true
t_vectors = torch.Tensor(v.astype(np.float32)).cuda()
# + hidden=true
mu = 1e-5
# + hidden=true
def grads_t(M, W, H):
R = W.mm(H)-M
return (R.mm(H.t()) + penalty_t(W, mu)*lam,
W.t().mm(R) + penalty_t(H, mu)*lam) # dW, dH
def penalty_t(M, mu):
return (M<mu).type(tc.FloatTensor)*torch.clamp(M - mu, max=0.)
def upd_t(M, W, H, lr):
dW,dH = grads_t(M,W,H)
W.sub_(lr*dW); H.sub_(lr*dH)
def report_t(M,W,H):
print((M-W.mm(H)).norm(2), W.min(), H.min(), (W<0).sum(), (H<0).sum())
# + hidden=true
t_W = tc.FloatTensor(m,d)
t_H = tc.FloatTensor(d,n)
t_W.normal_(std=0.01).abs_();
t_H.normal_(std=0.01).abs_();
# + hidden=true
d=6; lam=100; lr=0.05
# + hidden=true
for i in range(1000):
upd_t(t_vectors,t_W,t_H,lr)
if i % 100 == 0:
report_t(t_vectors,t_W,t_H)
lr *= 0.9
# + hidden=true
show_topics(t_H.cpu().numpy())
# + hidden=true
plt.plot(t_H.cpu().numpy()[0])
# + hidden=true
t_W.mm(t_H).max()
# + hidden=true
t_vectors.max()
# + [markdown] heading_collapsed=true
# ### PyTorch: autograd
# + [markdown] hidden=true
# Above, we used our knowledge of what the gradient of the loss function was to do SGD from scratch in PyTorch. However, PyTorch has an automatic differentiation package, [autograd](http://pytorch.org/docs/autograd.html) which we could use instead. This is really useful, in that we can use autograd on problems where we don't know what the derivative is.
#
# The approach we use below is very general, and would work for almost any optimization problem.
#
# In PyTorch, Variables have the same API as tensors, but Variables remember the operations used on to create them. This lets us take derivatives.
# + [markdown] heading_collapsed=true hidden=true
# #### PyTorch Autograd Introduction
# + [markdown] hidden=true
# Example taken from [this tutorial](http://pytorch.org/tutorials/beginner/former_torchies/autograd_tutorial.html) in the official documentation.
# + hidden=true
x = Variable(torch.ones(2, 2), requires_grad=True)
print(x)
# + hidden=true
print(x.data)
# + hidden=true
print(x.grad)
# + hidden=true
y = x + 2
print(y)
# + hidden=true
z = y * y * 3
out = z.sum()
print(z, out)
# + hidden=true
out.backward()
print(x.grad)
# + hidden=true
# + [markdown] hidden=true
# #### Using Autograd for NMF
# + hidden=true
lam=1e6
# + hidden=true
pW = Variable(tc.FloatTensor(m,d), requires_grad=True)
pH = Variable(tc.FloatTensor(d,n), requires_grad=True)
pW.data.normal_(std=0.01).abs_()
pH.data.normal_(std=0.01).abs_();
# + hidden=true
def report():
W,H = pW.data, pH.data
print((M-pW.mm(pH)).norm(2).data[0], W.min(), H.min(), (W<0).sum(), (H<0).sum())
def penalty(A):
return torch.pow((A<0).type(tc.FloatTensor)*torch.clamp(A, max=0.), 2)
def penalize(): return penalty(pW).mean() + penalty(pH).mean()
def loss(): return (M-pW.mm(pH)).norm(2) + penalize()*lam
# + hidden=true
M = Variable(t_vectors).cuda()
# + hidden=true
opt = torch.optim.Adam([pW,pH], lr=1e-3, betas=(0.9,0.9))
lr = 0.05
report()
# + [markdown] hidden=true
# How to apply SGD, using autograd:
# + hidden=true
for i in range(1000):
opt.zero_grad()
l = loss()
l.backward()
opt.step()
if i % 100 == 99:
report()
lr *= 0.9 # learning rate annealling
# + hidden=true
h = pH.data.cpu().numpy()
show_topics(h)
# + hidden=true
plt.plot(h[0]);
# -
# ### Comparing Approaches
# + [markdown] heading_collapsed=true
# #### Scikit-Learn's NMF
# - Fast
# - No parameter tuning
# - Relies on decades of academic research, took experts a long time to implement
#
# <img src="images/nimfa.png" alt="research on NMF" style="width: 80%"/>
# source: [Python Nimfa Documentation](http://nimfa.biolab.si/)
#
# #### Using PyTorch and SGD
# - Took us an hour to implement, didn't have to be NMF experts
# - Parameters were fiddly
# - Not as fast (tried in numpy and was so slow we had to switch to PyTorch)
# -
# ## Truncated SVD
# We saved a lot of time when we calculated NMF by only calculating the subset of columns we were interested in. Is there a way to get this benefit with SVD? Yes there is! It's called truncated SVD. We are just interested in the vectors corresponding to the **largest** singular values.
# <img src="images/svd_fb.png" alt="" style="width: 80%"/>
# (source: [Facebook Research: Fast Randomized SVD](https://research.fb.com/fast-randomized-svd/))
# #### Shortcomings of classical algorithms for decomposition:
# - Matrices are "stupendously big"
# - Data are often **missing or inaccurate**. Why spend extra computational resources when imprecision of input limits precision of the output?
# - **Data transfer** now plays a major role in time of algorithms. Techniques the require fewer passes over the data may be substantially faster, even if they require more flops (flops = floating point operations).
# - Important to take advantage of **GPUs**.
#
# (source: [Halko](https://arxiv.org/abs/0909.4061))
# #### Advantages of randomized algorithms:
# - inherently stable
# - performance guarantees do not depend on subtle spectral properties
# - needed matrix-vector products can be done in parallel
#
# (source: [Halko](https://arxiv.org/abs/0909.4061))
# ### Randomized SVD
# Reminder: full SVD is **slow**. This is the calculation we did above using Scipy's Linalg SVD:
vectors.shape
# %time U, s, Vh = linalg.svd(vectors, full_matrices=False)
print(U.shape, s.shape, Vh.shape)
# Fortunately, there is a faster way:
# %time u, s, v = decomposition.randomized_svd(vectors, 5)
# The runtime complexity for SVD is $\mathcal{O}(\text{min}(m^2 n,\; m n^2))$
# **Question**: How can we speed things up? (without new breakthroughs in SVD research)
# **Idea**: Let's use a smaller matrix (with smaller $n$)!
#
# Instead of calculating the SVD on our full matrix $A$ which is $m \times n$, let's use $B = A Q$, which is just $m \times r$ and $r << n$
#
# We haven't found a better general SVD method, we are just using the method we have on a smaller matrix.
# %time u, s, v = decomposition.randomized_svd(vectors, 5)
u.shape, s.shape, v.shape
show_topics(v)
# Here are some results from [Facebook Research](https://research.fb.com/fast-randomized-svd/):
#
# <img src="images/randomizedSVDbenchmarks.png" alt="" style="width: 80%"/>
# **Johnson-Lindenstrauss Lemma**: ([from wikipedia](https://en.wikipedia.org/wiki/Johnson%E2%80%93Lindenstrauss_lemma)) a small set of points in a high-dimensional space can be embedded into a space of much lower dimension in such a way that distances between the points are nearly preserved.
#
# It is desirable to be able to reduce dimensionality of data in a way that preserves relevant structure. The Johnson–Lindenstrauss lemma is a classic result of this type.
# ### Implementing our own Randomized SVD
from scipy import linalg
# The method `randomized_range_finder` finds an orthonormal matrix whose range approximates the range of A (step 1 in our algorithm above). To do so, we use the LU and QR factorizations, both of which we will be covering in depth later.
#
# I am using the [scikit-learn.extmath.randomized_svd source code](https://github.com/scikit-learn/scikit-learn/blob/14031f65d144e3966113d3daec836e443c6d7a5b/sklearn/utils/extmath.py) as a guide.
# computes an orthonormal matrix whose range approximates the range of A
# power_iteration_normalizer can be safe_sparse_dot (fast but unstable), LU (imbetween), or QR (slow but most accurate)
def randomized_range_finder(A, size, n_iter=5):
Q = np.random.normal(size=(A.shape[1], size))
for i in range(n_iter):
Q, _ = linalg.lu(A @ Q, permute_l=True)
Q, _ = linalg.lu(A.T @ Q, permute_l=True)
Q, _ = linalg.qr(A @ Q, mode='economic')
return Q
# And here's our randomized SVD method:
def randomized_svd(M, n_components, n_oversamples=10, n_iter=4):
n_random = n_components + n_oversamples
Q = randomized_range_finder(M, n_random, n_iter)
# project M to the (k + p) dimensional space using the basis vectors
B = Q.T @ M
# compute the SVD on the thin matrix: (k + p) wide
Uhat, s, V = linalg.svd(B, full_matrices=False)
del B
U = Q @ Uhat
return U[:, :n_components], s[:n_components], V[:n_components, :]
u, s, v = randomized_svd(vectors, 5)
# %time u, s, v = randomized_svd(vectors, 5)
u.shape, s.shape, v.shape
show_topics(v)
# Write a loop to calculate the error of your decomposition as you vary the # of topics. Plot the result
# #### Answer
#Exercise: Write a loop to calculate the error of your decomposition as you vary the # of topics
plt.plot(range(0,n*step,step), error)
# **Further Resources**:
# - [a whole course on randomized algorithms](http://www.cs.ubc.ca/~nickhar/W12/)
# + [markdown] heading_collapsed=true
# ### More Details
# + [markdown] hidden=true
# Here is a process to calculate a truncated SVD, described in [Finding Structure with Randomness: Probabilistic Algorithms for Constructing Approximate Matrix Decompositions](https://arxiv.org/pdf/0909.4061.pdf) and [summarized in this blog post](https://research.fb.com/fast-randomized-svd/):
#
# 1\. Compute an approximation to the range of $A$. That is, we want $Q$ with $r$ orthonormal columns such that $$A \approx QQ^TA$$
#
#
# 2\. Construct $B = Q^T A$, which is small ($r\times n$)
#
#
# 3\. Compute the SVD of $B$ by standard methods (fast since $B$ is smaller than $A$), $B = S\,\Sigma V^T$
#
# 4\. Since $$ A \approx Q Q^T A = Q (S\,\Sigma V^T)$$ if we set $U = QS$, then we have a low rank approximation $A \approx U \Sigma V^T$.
# + [markdown] hidden=true
# #### So how do we find $Q$ (in step 1)?
# + [markdown] hidden=true
# To estimate the range of $A$, we can just take a bunch of random vectors $w_i$, evaluate the subspace formed by $Aw_i$. We can form a matrix $W$ with the $w_i$ as it's columns. Now, we take the QR decomposition of $AW = QR$, then the columns of $Q$ form an orthonormal basis for $AW$, which is the range of $A$.
#
# Since the matrix $AW$ of the product has far more rows than columns and therefore, approximately, orthonormal columns. This is simple probability - with lots of rows, and few columns, it's unlikely that the columns are linearly dependent.
# + [markdown] hidden=true
# #### The QR Decomposition
# + [markdown] hidden=true
# We will be learning about the QR decomposition **in depth** later on. For now, you just need to know that $A = QR$, where $Q$ consists of orthonormal columns, and $R$ is upper triangular. Trefethen says that the QR decomposition is the most important idea in numerical linear algebra! We will definitely be returning to it.
# + [markdown] hidden=true
# #### How should we choose $r$?
# + [markdown] hidden=true
# Suppose our matrix has 100 columns, and we want 5 columns in U and V. To be safe, we should project our matrix onto an orthogonal basis with a few more rows and columns than 5 (let's use 15). At the end, we will just grab the first 5 columns of U and V
#
# So even although our projection was only approximate, by making it a bit bigger than we need, we can make up for the loss of accuracy (since we're only taking a subset later).
# + hidden=true
# %time u, s, v = decomposition.randomized_svd(vectors, 5)
# + hidden=true
# %time u, s, v = decomposition.randomized_svd(vectors.todense(), 5)
# -
# ## End
| Mathematics/Numerical Methods/numerical-linear-algebra-grad-course/2. Topic Modeling with NMF and SVD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 (Develer Science)
# language: python
# name: develer-science
# ---
# # SciPy Challenge
#
# ## SciPy at a Glance
#
# The SciPy framework builds on top of the low-level NumPy framework for multidimensional arrays, and provides a large number of higher-level scientific algorithms. Some of the topics that SciPy covers are:
#
# * Special functions ([scipy.special](http://docs.scipy.org/doc/scipy/reference/special.html))
# * Integration ([scipy.integrate](http://docs.scipy.org/doc/scipy/reference/integrate.html))
# * Optimization ([scipy.optimize](http://docs.scipy.org/doc/scipy/reference/optimize.html))
# * Interpolation ([scipy.interpolate](http://docs.scipy.org/doc/scipy/reference/interpolate.html))
# * Fourier Transforms ([scipy.fftpack](http://docs.scipy.org/doc/scipy/reference/fftpack.html))
# * Signal Processing ([scipy.signal](http://docs.scipy.org/doc/scipy/reference/signal.html))
# * Linear Algebra ([scipy.linalg](http://docs.scipy.org/doc/scipy/reference/linalg.html))
# * Sparse Eigenvalue Problems ([scipy.sparse](http://docs.scipy.org/doc/scipy/reference/sparse.html))
# * Statistics ([scipy.stats](http://docs.scipy.org/doc/scipy/reference/stats.html))
# * Multi-dimensional image processing ([scipy.ndimage](http://docs.scipy.org/doc/scipy/reference/ndimage.html))
# * File IO ([scipy.io](http://docs.scipy.org/doc/scipy/reference/io.html))
# ## Sparse Matrices
#
# **Sparse Matrices** are very nice in some situations.
#
# For example, in some machine learning tasks, especially those associated
# with textual analysis, the data may be mostly zeros.
#
# Storing all these zeros is very inefficient.
#
# We can create and manipulate sparse matrices using the `scipy.sparse` module.
# There exists different implementations of sparse matrices, which are supposed to be efficient in different scenarios:
#
# - CSR: Compressed Sparse Rows
# - CSC: Compressec Sparse Colums
# - DOK: Dictionary of Keys
# - LIL: List of Lists
# - BSR: Block Sparse Row
# ## Ex 1.1
#
# Create a big numpy **dense** matrix filled with random numbers in
# `[0, 1)`.
# Generate a random number within this range and subsitute all the elements in the matrix **less than** this number with a zero.
#
# Save resulting matrix as a `DOK` sparse matrix
# ## Ex 1.2
#
# Repeat the previous exercise, but this time use a `CSR` sparse matrix.
# ## Ex 1.3
#
# Transform the previously generated sparse matrix back to a full dense `numpy.array`.
# ## Ex 1.4
#
# Generate two sparse Matrix and sum them together, choosing the most appropriate internal representation (i.e. `LIL`, `CSR`, `DOK`...).
#
# #### Hint: Oh c'mon.. :)
| 1_apprentice/2.3. Scipy Challenge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pandas Data Grammar Illustration
#
# This notebook illustrates the concepts of grammar of data in pandas using the bank dataset.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# +
# Read the data file into a DataFrame
import os
PROJECT_NAME = "bank"
home_dir = os.path.expanduser("~")
course_id = 'cpsc6300'
project_dir = os.path.join(home_dir, course_id, PROJECT_NAME)
project_data_dir = os.path.join(project_dir, "data", "bank")
data_file_path = os.path.join(project_data_dir, "bank-additional-bank-additional-full.csv")
df = pd.read_csv(data_file_path, sep=";")
# -
# ## Select a data slice
# +
### head() and tail()
# -
df.head(5)
df.tail(5)
# ### bracket selection
df[['age', 'job']]
# ### iloc
### Read the iloc documentation
help(df.iloc)
df.iloc[[10, 12, 15], [0, 1]]
# ### loc
# df.loc
help(df.loc)
df.loc[[0, 13]]
df.loc[[0, 13], ["age", "job"]]
df.loc[df.education == 'basic.4y']
# ## Sort
### sort_values
help(df.sort_values)
df.sort_values(by=["education", "age"])
### sort_values
help(df.sort_index)
df.sort_index(ascending=False)
df.loc[df.education == 'basic.4y']
# ### Unqiue values
### unqiue
help(pd.Series.unique)
df["job"].unique()
# ### drop_duplicates
help(pd.DataFrame.drop_duplicates)
df[["education", "job"]].drop_duplicates()
# ## Transform
education_map = {
'illiterate': 0,
'basic.4y': 4,
'basic.6y': 6,
'basic.9y': 9,
'high.school': 12,
'professional.course': 12,
'university.degree': 16,
'unknown': np.NaN,
}
education_map
df['education_numeric'] = df["education"].map(education_map)
help(pd.Series.map)
df.head()
# ### Assignment
df.loc[df.poutcome == 'nonexistent', "poutcome"] = np.nan
# ### Sample
df.sample(n=5, random_state=1)
# ## Aggregated statistics
df.age.max(), df.age.min()
df[["age", "nr.employed"]].describe()
# ## Delete Columns
df.drop(columns=["contact"], inplace=True)
# ## Group By
df.groupby("education").sum()
help(df.groupby)
| numpy_pandas_matplotlib/data_grammar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="mw2VBrBcgvGa"
# # Week 1 Assignment: Housing Prices
#
# In this exercise you'll try to build a neural network that predicts the price of a house according to a simple formula.
#
# Imagine that house pricing is as easy as:
#
# A house has a base cost of 50k, and every additional bedroom adds a cost of 50k. This will make a 1 bedroom house cost 100k, a 2 bedroom house cost 150k etc.
#
# How would you create a neural network that learns this relationship so that it would predict a 7 bedroom house as costing close to 400k etc.
#
# Hint: Your network might work better if you scale the house price down. You don't have to give the answer 400...it might be better to create something that predicts the number 4, and then your answer is in the 'hundreds of thousands' etc.
# + id="PUNO2E6SeURH"
import tensorflow as tf
import numpy as np
# + id="B-74xrKrBqGJ"
# GRADED FUNCTION: house_model
def house_model():
### START CODE HERE
# Define input and output tensors with the values for houses with 1 up to 6 bedrooms
# Hint: Remember to explictly set the dtype as float
xs = None
ys = None
# Define your model (should be a model with 1 dense layer and 1 unit)
model = None
# Compile your model
# Set the optimizer to Stochastic Gradient Descent
# and use Mean Squared Error as the loss function
model.compile(optimizer=None, loss=None)
# Train your model for 1000 epochs by feeding the i/o tensors
model.fit(None, None, epochs=None)
### END CODE HERE
return model
# -
# Now that you have a function that returns a compiled and trained model when invoked, use it to get the model to predict the price of houses:
# Get your trained model
model = house_model()
# Now that your model has finished training it is time to test it out! You can do so by running the next cell.
# + id="kMlInDdSBqGK"
new_y = 7.0
prediction = model.predict([new_y])[0]
print(prediction)
# -
# If everything went as expected you should see a prediction value very close to 4. **If not, try adjusting your code before submitting the assignment.** Notice that you can play around with the value of `new_y` to get different predictions. In general you should see that the network was able to learn the linear relationship between `x` and `y`, so if you use a value of 8.0 you should get a prediction close to 4.5 and so on.
# **Congratulations on finishing this week's assignment!**
#
# You have successfully coded a neural network that learned the linear relationship between two variables. Nice job!
#
# **Keep it up!**
| C1/W1/assignment/C1W1_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # End to End MLPerf Submission example
#
# This is following the [General MLPerf Submission Rules](https://github.com/mlcommons/policies/blob/master/submission_rules.adoc).
#
# ### Get the MLPerf Inference Benchmark Suite source code
#
# You run this notebook from the root of the 'mlcommons/inference' repo that you cloned with
# ```
# git clone --recurse-submodules https://github.com/mlcommons/inference.git --depth 1
# ```
# ### Build loadgen
# build loadgen
# !pip install pybind11
# !cd loadgen; CFLAGS="-std=c++14 -O3" python setup.py develop
# !cd vision/classification_and_detection; python setup.py develop
# ### Set Working Directory
# %cd vision/classification_and_detection
# # Download data
#
# For this example, the ImageNet and/or COCO validation data should already be on the host system. See the [MLPerf Image Classification task](https://github.com/mlcommons/inference/tree/master/vision/classification_and_detection#datasets) for more details on obtaining this. For the following step each validation dataset is stored in /workspace/data/. You should change this to the location in your setup.
# + language="bash"
#
# mkdir data
# ln -s /workspace/data/imagenet2012 data/
# ln -s /workspace/data/coco data/
# -
# ### Download models
# + language="bash"
#
# mkdir models
#
# # resnet50
# wget -q https://zenodo.org/record/2535873/files/resnet50_v1.pb -O models/resnet50_v1.pb
# wget -q https://zenodo.org/record/2592612/files/resnet50_v1.onnx -O models/resnet50_v1.onnx
#
# # ssd-mobilenet
# wget -q http://download.tensorflow.org/models/object_detection/ssd_mobilenet_v1_coco_2018_01_28.tar.gz -O models/ssd_mobilenet_v1_coco_2018_01_28.tar.gz
# tar zxvf ./models/ssd_mobilenet_v1_coco_2018_01_28.tar.gz -C ./models; mv models/ssd_mobilenet_v1_coco_2018_01_28/frozen_inference_graph.pb ./models/ssd_mobilenet_v1_coco_2018_01_28.pb
# wget -q https://zenodo.org/record/3163026/files/ssd_mobilenet_v1_coco_2018_01_28.onnx -O models/ssd_mobilenet_v1_coco_2018_01_28.onnx
#
# # ssd-resnet34
# wget -q https://zenodo.org/record/3345892/files/tf_ssd_resnet34_22.1.zip -O models/tf_ssd_resnet34_22.1.zip
# unzip ./models/tf_ssd_resnet34_22.1.zip -d ./models; mv models/tf_ssd_resnet34_22.1/resnet34_tf.22.1.pb ./models
# wget -q https://zenodo.org/record/3228411/files/resnet34-ssd1200.onnx -O models/resnet34-ssd1200.onnx
# -
# ### Run benchmarks using the reference implementation
#
# Lets prepare a submission for ResNet-50 on a cloud datacenter server with a NVIDIA T4 GPU using TensorFlow.
#
# The following script will run those combinations and prepare a submission directory, following the general submission rules documented [here](https://github.com/mlcommons/policies/blob/master/submission_rules.adoc).
# +
import logging
import os
logger = logging.getLogger()
logger.setLevel(logging.CRITICAL)
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
os.environ['CUDA_VISIBLE_DEVICES'] = "0"
# final results go here
ORG = "mlperf-org"
DIVISION = "closed"
SUBMISSION_ROOT = "/tmp/mlperf-submission"
SUBMISSION_DIR = os.path.join(SUBMISSION_ROOT, DIVISION, ORG)
os.environ['SUBMISSION_ROOT'] = SUBMISSION_ROOT
os.environ['SUBMISSION_DIR'] = SUBMISSION_DIR
os.makedirs(SUBMISSION_DIR, exist_ok=True)
os.makedirs(os.path.join(SUBMISSION_DIR, "measurements"), exist_ok=True)
os.makedirs(os.path.join(SUBMISSION_DIR, "code"), exist_ok=True)
# + language="bash"
#
# # where to find stuff
# export DATA_ROOT=`pwd`/data
# export MODEL_DIR=`pwd`/models
#
# # options for official runs
# gopt="--max-batchsize 8 --samples-per-query 40 --threads 2 --qps 145"
#
#
# function one_run {
# # args: mode count framework device model ...
# scenario=$1; shift
# count=$1; shift
# framework=$1
# device=$2
# model=$3
# system_id=$framework-$device
# echo "====== $model/$scenario ====="
#
# case $model in
# resnet50)
# cmd="tools/accuracy-imagenet.py --imagenet-val-file $DATA_ROOT/imagenet2012/val_map.txt"
# offical_name="resnet";;
# ssd-mobilenet)
# cmd="tools/accuracy-coco.py --coco-dir $DATA_ROOT/coco"
# offical_name="ssd-small";;
# ssd-resnet34)
# cmd="tools/accuracy-coco.py --coco-dir $DATA_ROOT/coco"
# offical_name="ssd-large";;
# esac
# output_dir=$SUBMISSION_DIR/results/$system_id/$offical_name
#
# # accuracy run
# ./run_local.sh $@ --scenario $scenario --accuracy --output $output_dir/$scenario/accuracy
# python $cmd --mlperf-accuracy-file $output_dir/$scenario/accuracy/mlperf_log_accuracy.json \
# > $output_dir/$scenario/accuracy/accuracy.txt
# cat $output_dir/$scenario/accuracy/accuracy.txt
#
# # performance run
# cnt=0
# while [ $cnt -lt $count ]; do
# let cnt=cnt+1
# ./run_local.sh $@ --scenario $scenario --output $output_dir/$scenario/performance/run_$cnt
# done
#
# # setup the measurements directory
# mdir=$SUBMISSION_DIR/measurements/$system_id/$offical_name/$scenario
# mkdir -p $mdir
# cp ../../mlperf.conf $mdir
#
# # reference app uses command line instead of user.conf
# echo "# empty" > $mdir/user.conf
# touch $mdir/README.md
# impid="reference"
# cat > $mdir/$system_id"_"$impid"_"$scenario".json" <<EOF
# {
# "input_data_types": "fp32",
# "retraining": "none",
# "starting_weights_filename": "https://zenodo.org/record/2535873/files/resnet50_v1.pb",
# "weight_data_types": "fp32",
# "weight_transformations": "none"
# }
# EOF
# }
#
# function one_model {
# # args: framework device model ...
# one_run SingleStream 1 $@ --max-latency 0.0005
# one_run Server 1 $@
# one_run Offline 1 $@ --qps 1000
# one_run MultiStream 1 $@
# }
#
#
# # run image classifier benchmarks
# export DATA_DIR=$DATA_ROOT/imagenet2012
# one_model tf gpu resnet50 $gopt
# -
# There might be large trace files in the submission directory - we can delete them.
# !find {SUBMISSION_DIR}/ -name mlperf_log_trace.json -delete
# ### Complete submission directory
#
# Add the required meta data to the submission.
# + language="bash"
#
# #
# # setup systems directory
# #
# if [ ! -d ${SUBMISSION_DIR}/systems ]; then
# mkdir ${SUBMISSION_DIR}/systems
# fi
#
# cat > ${SUBMISSION_DIR}/systems/tf-gpu.json <<EOF
# {
# "division": "closed",
# "status": "available",
# "submitter": "mlperf-org",
# "system_name": "tf-gpu",
# "system_type": "datacenter",
#
# "number_of_nodes": 1,
# "host_memory_capacity": "32GB",
# "host_processor_core_count": 1,
# "host_processor_frequency": "3.50GHz",
# "host_processor_model_name": "Intel(R) Xeon(R) CPU E5-1620 v3 @ 3.50GHz",
# "host_processors_per_node": 1,
# "host_storage_capacity": "512GB",
# "host_storage_type": "SSD",
#
# "accelerator_frequency": "-",
# "accelerator_host_interconnect": "-",
# "accelerator_interconnect": "-",
# "accelerator_interconnect_topology": "-",
# "accelerator_memory_capacity": "16GB",
# "accelerator_memory_configuration": "none",
# "accelerator_model_name": "T4",
# "accelerator_on-chip_memories": "-",
# "accelerators_per_node": 1,
#
# "framework": "v1.14.0-rc1-22-gaf24dc9",
# "operating_system": "ubuntu-18.04",
# "other_software_stack": "cuda-11.2",
# "sw_notes": ""
# }
# EOF
# + language="bash"
#
# #
# # setup code directory
# #
# dir=${SUBMISSION_DIR}/code/resnet/reference
# mkdir -p $dir
# echo "git clone https://github.com/mlcommons/inference.git" > $dir/VERSION.txt
# git rev-parse HEAD >> $dir/VERSION.txt
# -
# ### What's in the submission directory now ?
#
# !find {SUBMISSION_ROOT}/ -type f
# If we look at some files:
# !echo "-- SingleStream Accuracy"; head {SUBMISSION_DIR}/results/tf-gpu/resnet/SingleStream/accuracy/accuracy.txt
# !echo "\n-- SingleStream Summary"; head {SUBMISSION_DIR}/results/tf-gpu/resnet/SingleStream/performance/run_1/mlperf_log_summary.txt
# !echo "\n-- Server Summary"; head {SUBMISSION_DIR}/results/tf-gpu/resnet/Server/performance/run_1/mlperf_log_summary.txt
# ## Run the submission checker
#
# Finally, run the submission checker tool that does some sanity checking on your submission.
# We run it at the end and attach the output to the submission.
# !python ../../tools/submission/submission-checker.py --input {SUBMISSION_ROOT} > {SUBMISSION_DIR}/submission-checker.log 2>&1
# !cat {SUBMISSION_DIR}/submission-checker.log
| SubmissionExample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.5
# language: julia
# name: julia-1.0
# ---
using JuliaAcademyData; activate("Foundations of machine learning")
# # Representing data in a computer
#
# The core of data science and machine learning is **data**: we are interested in extracting knowledge from data.
#
# But how exactly do computers represent data? Let's find out exactly what an "artificial intelligence" has at its disposal to learn from.
# ## Data is represented as arrays
#
# Let's take a look at some fruit. Using the `Images.jl` library, we can load in some images:
# +
using Images
apple = load(datapath("data/10_100.jpg"))
# -
banana = load(datapath("data/104_100.jpg"))
# Here we have images of apples and bananas. We would eventually like to build a program that can automatically distinguish between the two. However, the computer doesn't "see" an apple or a banana; instead, it just sees numbers.
#
# An image is encoded in something called an **array**, which is like a container that has boxes or slots for individual pieces of data:
#
# An array is a bunch of numbers in connected boxes; the figure above shows a 1-dimensional array. Our images are instead 2-dimensional arrays, or matrices, of numbers, arranged something like this:
# <img src="https://raw.githubusercontent.com/JuliaComputing/JuliaAcademyData.jl/master/courses/Foundations%20of%20machine%20learning/data/array2d.png" alt="attachment:array2d.png" width="500"/>
# For example, `apple` is an image, consisting of a 100x100 array of numbers:
typeof(apple)
size(apple)
a = [ 1 2 3;4 5 6]
typeof(a)
size(a)
# We can grab the datum stored in the box at row `i` and column `j` by *indexing* using square brackets: `[i, j]`. For example, let's get the pixel (piece of the image) in box $(40, 60)$, i.e. in the 40th row and 60th column of the image:
apple
dump(apple[40, 60])
apple[18:20,29:31]
# We see that Julia displays a coloured box! Julia, via the `Colors.jl` package, is clever enough to display colours in a way that is useful to us humans!
#
# So, in fact, an image is a 2D array, in which each element of the array is an object (a collection of numbers) describing a coloured pixel.
# ## Colors as numbers
#
# How, then, are these colors actually stored? Computers store colors in RGB format, that is they store a value between 0 and 1 for each of three "channels": red, green, and blue. Here, 0 means none of that color and 1 means the brightest form of that color. The overall color is a combination of those three colors.
#
# For example, we can pull out the `red` value using the function `red` applied to the color. Since internally the actual value is stored in a special format, we choose to convert it to a standard floating-point number using the `Float64` function:
Float64(red(apple[40, 60]))
using Statistics
[ mean(float.(c.(img))) for c = [red,green,blue], img = [apple,banana] ]
using Plots
histogram(float.(green.(apple[:])),color="red",label="apple", normalize=true, nbins=25)
histogram!(float.(green.(banana[:])),color="yellow",label="banana",normalize=true, nbins=25)
apple
float(red(banana[50,20]))
banana[50,20]
# +
pixel = apple[40, 60]
red_value = Float64( red(pixel) )
green_value = Float64( green(pixel) )
blue_value = Float64( blue(pixel) )
print("The RGB values are ($red_value, $green_value, $blue_value)")
# -
# Since the red value is high while the others are low, this means that at pixel `(40, 60)`, the picture of the apple is very red. If we do the same at one of the corners of the picture, we get the following:
# +
pixel = apple[1, 1]
red_value = Float64( red(pixel) )
green_value = Float64( green(pixel) )
blue_value = Float64( blue(pixel) )
print("The RGB values are ($red_value, $green_value, $blue_value)")
# -
apple
# We see that every color is bright, which corresponds to white.
# ## Working on an image as a whole
#
# In Julia, to apply a function to the whole of an array, we place a `.` between the function name and the left parenthesis (`(`), so the following gives us the `red` value of every pixel in the image:
redpartofapple = Float64.(red.(apple))
mean(redpartofapple)
using Plots
gr()
histogram(redpartofapple[:],color=:red,label="redness in the apple")
# Note that we get a 2D array (matrix) back.
# Julia's [mathematical standard library](https://docs.julialang.org/en/stable/stdlib/math/#Mathematics-1) has many mathematical functions built in. One of them is the `mean` function, which computes the average value. If we apply this to our apple:
mean(Float64.(red.(apple)))
# we see that the value indicates that the average amount of red in the image is a value between the amount of red in the apple and the amount of red in the white background.
#
# *Somehow we need to teach a computer to use this information about a picture to recognize that there's an apple there!*
# ## A quick riddle
#
# Here's a quick riddle. Let's check the average value of red in the image of the banana.
mean(Float64.(red.(banana)))
# Oh no, that's more red than our apple? This isn't a mistake and is actually true! Before you move onto the next exercise, examine the images of the apple and the banana very carefully and see if you can explain why this is expected.
# #### Exercise 1
#
# What is the average value of blue in the banana?
#
# (To open a new box use <ESC>+b (b is for "below", what do you think a does?))
mean(Float64.(blue.(banana)))
# #### Solution
#
# We can calculate the average value of blue in the banana via
mean(Float64.(blue.(banana)))
# The result is approximately `0.8`.
# #### Exercise 2
#
# Does the banana have more blue or more green?
banana_blue = mean(Float64.(blue.(banana)))
banana_green = mean(Float64.(green.(banana)))
if banana_blue > banana_green
print("Banana has more blue $banana_blue")
else
print("Banana has more green $banana_green")
end
# #### Solution
#
# The average value of green in the banana is
mean(Float64.(green.(banana)))
# which gives approximately `0.88`. The banana has more green on average.
# # Modeling data 1
#
# Machine learning and data science is about modeling data. **Modeling** is the representation of an idea with some parameters and a mathematical representation which we will encode in software. All machine learning methods are about training a computer to fit a model to some data. Even the fanciest neural networks are simply choices for models. In this notebook, we will begin to start building our first computational model of data.
# ## Modeling data is hard!
#
# Let's pick up where we left off in notebook 1 with fruit. We were left with a riddle: when we load images of apples and bananas,
# +
# using Pkg; Pkg.add("Images")
using Images, Statistics
apple = load(datapath("data/10_100.jpg"))
# -
banana = load(datapath("data/104_100.jpg"))
# and then compare their average value for the color red, we end up with something that is perhaps surprising:
# +
apple_red_amount = mean(Float64.(red.(apple)))
banana_red_amount = mean(Float64.(red.(banana)));
"The average value of red in the apple is $apple_red_amount, " *
"while the average value of red in the banana is $banana_red_amount."
# -
# We see that the banana's mean red value is higher than the apple's, even though the apple looks much redder. Can you guess why?
#
# There are actually two reasons. One of the reasons is the background: the image of the banana has a lot more white background than the apple, and that white background has a red value of 1! In our minds we ignore the background and say "the banana is bright yellow, the apple is dark red", but a computer just has a bundle of numbers and does not know where it should be looking.
#
# The other issue is that "bright yellow" isn't a color that exists in a computer. The computer has three colors: red, green, and blue. "Bright yellow" in a computer is a mixture of red and green, and it just so happens that to get this color yellow, it needs more red than the apple!
"The amount of red in the apple at (60, 60) is $(Float64(red(apple[60, 60]))), " *
"while the amount of red in the banana at (60, 60) is $(Float64(red(banana[60, 60])))."
apple[60,60]
banana[60,60]
# This is a clear example that modeling data is hard!
# ### A note on string interpolation
#
# In the last two input cells, we *interpolated a string*. This means that when we write the string using quotation marks (`" "`), we insert a placeholder for some **value** we want the string to include. When the string is evaluated, the value we want the string to include replaces the placeholder. For example, in the following string,
#
# ```julia
# mystring = "The average value of red in the apple is $apple_red_amount"
# ```
#
# `$apple_red_amount` is a placeholder for the value stored in the variable `apple_red_amount`. Julia knows that we want to use the value bound to the variable `apple_red_amount` and *not* the word "apple_red_amount" because of the dollar sign, `$`, that comes before `apple_red_amount`.
# #### Exercise 1
#
# Execute the following code to see what the dollar sign does:
#
# ```julia
# mypi = 3.14159
# println("I have a variable called mypi that has a value of $mypi.")
# ```
# #### Solution
mypi = 3.14159
println("I have a variable called mypi that has a value of $mypi.")
# #### Exercise 2
#
# Alter and execute the code that creates `mystring` below
#
# ```julia
# apple_blue_amount = mean(Float64.(blue.(apple)))
# mystring = "The average amount of blue in the apple is apple_blue_amount"
# ```
#
# so that `println(mystring)` prints a string that reports the mean value of blue coloration in our image of an apple.
# #### Solution
# Add a `$` in front of `apple_blue_amount`:
apple_blue_amount = mean(Float64.(blue.(apple)))
mystring = "The average amount of blue in the apple is $apple_blue_amount."
# ## Take some time to think about the data
#
# Apples and bananas are very different, but how could we use the array of RGB values (which is how the images are represented in the computer, as we saw in notebook 1) to tell the difference between the two? Here are some quick ideas:
#
# - We could use the shape of the object in the image. But how can we encode ideas about shape from an array?
# - We could use the size of the object in the image. But how do we calculate that size?
# - We could use another color, or combinations of colors, from the image. Which colors?
#
# Let's go with the last route. The banana is yellow, which is a combination of red and green, while the apple is red. This means that the color that clearly differentiates between the two is not red, but green!
# +
apple_green_amount = mean(Float64.(green.(apple)))
banana_green_amount = mean(Float64.(green.(banana)));
"The average value of green in the apple is $apple_green_amount, " *
"while the average value of green in the banana is $banana_green_amount."
# -
# The processes that we just went through are assigned fancy names: feature selection and data munging.
#
# **Feature selection** is the process of subsetting the data to a more relevant and informative set. We took the full image data and decided to select out the green channel.
#
# **Data munging** is transforming the data into a format more suitable for modeling. Here, instead of keeping the full green channel, we transformed it down to a single data point: the average amount of green.
# ## Building a model
#
# We want to model the connection between "the average amount of green" and "is an apple or banana".
#
# <img src="https://raw.githubusercontent.com/JuliaComputing/JuliaAcademyData.jl/master/courses/Foundations%20of%20machine%20learning/data/data_flow.png" alt="Drawing" style="width: 800px;"/>
#
# This model is a mathematical function which takes in our data and spits out a number that we will interpret as "is an apple" or "is a banana".
#
# <img src="https://raw.githubusercontent.com/JuliaComputing/JuliaAcademyData.jl/master/courses/Foundations%20of%20machine%20learning/data/what_is_model.png" alt="Drawing" style="width: 500px;"/>
#
#
# We will interpret the output of the function as "is an apple" if the output is close to 0, and "is a banana" if it's close to 1. Anything in the middle is something we are unsure about. Here we're using a mathematical function to perform a **classification**.
#
# Knowing how to declare and work with functions will allow us to model our data in the coming sections, so this is the subject of the next notebook!
| 0300.Representing-data-with-models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### 1. Import pandas library
# #### 2. Import pymysql and sqlalchemy as you have learnt in the lesson of importing/exporting data
#
# #### 3. Create a mysql engine to set the connection to the server. Check the connection details in [this link](https://relational.fit.cvut.cz/search?tableCount%5B%5D=0-10&tableCount%5B%5D=10-30&dataType%5B%5D=Numeric&databaseSize%5B%5D=KB&databaseSize%5B%5D=MB)
# #### 4. Import the users table
# #### 5. Rename Id column to userId
# #### 6. Import the posts table.
# #### 7. Rename Id column to postId and OwnerUserId to userId
# #### 8. Define new dataframes for users and posts with the following selected columns:
# **users columns**: userId, Reputation,Views,UpVotes,DownVotes
# **posts columns**: postId, Score,userID,ViewCount,CommentCount
# #### 8. Merge both dataframes, users and posts.
# You will need to make a [merge](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.merge.html) of posts and users dataframes.
# #### 9. How many missing values do you have in your merged dataframe? On which columns?
# #### 10. You will need to make something with missing values. Will you clean or filling them? Explain.
# **Remember** to check the results of your code before passing to the next step
# #### 11. Adjust the data types in order to avoid future issues. Which ones should be changed?
# #### Bonus: Identify extreme values in your merged dataframe as you have learned in class, create a dataframe called outliers with the same columns as our data set and calculate the bounds. The values of the outliers dataframe will be the values of the merged_df that fall outside that bounds. You will need to save your outliers dataframe to a csv file on your-code folder.
| module-1/lab-data_cleaning/your-code/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/teampurpler/Mechine_Learning_Projects/blob/main/Supervised_Learning_Project_Bank_Customer_Churn_Prediction.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="R88Ms0MTi0Ma"
# ## Bank Customer Churn Prediction
#
# > Indented block
#
#
# + [markdown] id="WA6lL1fni0Mb"
# In this project, we use supervised learning models to identify customers who are likely to churn in the future. Furthermore, we will analyze top factors that influence user retention. [Dataset information](https://www.kaggle.com/adammaus/predicting-churn-for-bank-customers).
# + [markdown] id="bO94-bXZi0Md"
# ## Contents
# + [markdown] id="SIvRSRqAi0Md"
#
# * [Part 1: Data Exploration](#Part-1:-Data-Exploration)
# * [Part 2: Feature Preprocessing](#Part-2:-Feature-Preprocessing)
# * [Part 3: Model Training and Results Evaluation](#Part-3:-Model-Training-and-Result-Evaluation)
# + [markdown] id="TUoI2S7Bi6iR"
# # Part 0: Setup Google Drive Environment / Data Collection
# check this [link](https://colab.research.google.com/notebooks/io.ipynb) for more info
# + id="neechzbWi7rV"
# install pydrive to load data
# !pip install -U -q PyDrive
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
# + id="UScKyL2TjARW"
# the same way we get id from last class
#https://drive.google.com/file/d/1szdCZ98EK59cfJ4jG03g1HOv_OhC1oyN/view?usp=sharing
id = "1szdCZ98EK59cfJ4jG03g1HOv_OhC1oyN"
file = drive.CreateFile({'id':id})
file.GetContentFile('bank_churn.csv')
# + id="nK7A1qhYSDxM" colab={"base_uri": "https://localhost:8080/", "height": 250} outputId="99d34d85-6b53-46aa-b389-c07f164f3652"
import pandas as pd
df = pd.read_csv('bank_churn.csv')
df.head()
# + [markdown] id="a6bG_gAPi0Me"
# # Part 1: Data Exploration
# + [markdown] id="bspx2K6fi0Me"
# ### Part 1.1: Understand the Raw Dataset
# + id="kuTHKjk-i0Mf"
import pandas as pd
import numpy as np
churn_df = pd.read_csv('bank_churn.csv')
# + id="hHNZRs2Ti0Mi" colab={"base_uri": "https://localhost:8080/", "height": 250} outputId="49ad1a6d-96c6-4159-ce78-39dd34f6079e"
churn_df.head()
# + id="ht5YOBdx8NLV" colab={"base_uri": "https://localhost:8080/"} outputId="4673cfc7-2d33-4767-ac53-f69ac3659efd"
# check data info
churn_df.info()
# + id="ZASeB8_089yA" colab={"base_uri": "https://localhost:8080/"} outputId="03cd3ca5-ab90-431d-ea90-a4edc6de20fa"
# check the unique values for each column
churn_df.nunique()
# + id="4ec5r_Qdi0NL"
# Get target variable
y = churn_df['Exited']
# + [markdown] id="SsAbAjhvi0Mx"
# ### Part 1.2: Understand the features
# + id="_t1xsBp--_0K" colab={"base_uri": "https://localhost:8080/"} outputId="70d00dbc-26a2-4232-c241-bf7d2abcba1f"
# check missing values
churn_df.isnull().sum()
# + id="BIqBIpOt_COM" colab={"base_uri": "https://localhost:8080/", "height": 300} outputId="d4ebe775-0f3e-4d93-985d-e7787ab09a46"
# understand Numerical feature
# discrete/continuous
# 'CreditScore', 'Age', 'Tenure', 'NumberOfProducts'
# 'Balance', 'EstimatedSalary'
churn_df[['CreditScore', 'Age', 'Tenure', 'NumOfProducts','Balance', 'EstimatedSalary']].describe()
# + id="uSWC_9arxlfk"
# check the feature distribution
# pandas.DataFrame.describe()
# boxplot, distplot, countplot
import matplotlib.pyplot as plt
import seaborn as sns
# + id="E6o4PlZbuSYy" colab={"base_uri": "https://localhost:8080/", "height": 623} outputId="e37bd6bb-3d76-4e98-b034-06a2f611c8cc"
# boxplot for numerical feature
_,axss = plt.subplots(2,3, figsize=[20,10])
sns.boxplot(x='Exited', y ='CreditScore', data=churn_df, ax=axss[0][0])
sns.boxplot(x='Exited', y ='Age', data=churn_df, ax=axss[0][1])
sns.boxplot(x='Exited', y ='Tenure', data=churn_df, ax=axss[0][2])
sns.boxplot(x='Exited', y ='NumOfProducts', data=churn_df, ax=axss[1][0])
sns.boxplot(x='Exited', y ='Balance', data=churn_df, ax=axss[1][1])
sns.boxplot(x='Exited', y ='EstimatedSalary', data=churn_df, ax=axss[1][2])
# + colab={"base_uri": "https://localhost:8080/", "height": 623} id="U0xZ-y3cw8JJ" outputId="8020e101-9650-4d7a-d108-c8b0f3e7c717"
# understand categorical feature
# 'Geography', 'Gender'
# 'HasCrCard', 'IsActiveMember'
_,axss = plt.subplots(2,2, figsize=[20,10])
sns.countplot(x='Exited', hue='Geography', data=churn_df, ax=axss[0][0])
sns.countplot(x='Exited', hue='Gender', data=churn_df, ax=axss[0][1])
sns.countplot(x='Exited', hue='HasCrCard', data=churn_df, ax=axss[1][0])
sns.countplot(x='Exited', hue='IsActiveMember', data=churn_df, ax=axss[1][1])
# + [markdown] id="aFa4d6t3i0NH"
# # Part 2: Feature Preprocessing
# + id="3sfa2fQx2xXa"
# Get feature space by dropping useless feature
to_drop = ['RowNumber','CustomerId','Surname','Exited']
X = churn_df.drop(to_drop, axis=1)
# + id="lZFFVHUgftso" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="4c20b302-c13c-4558-ddc2-47433adfd69b"
X.head()
# + colab={"base_uri": "https://localhost:8080/"} id="i6WyuvNyxNeE" outputId="626e731c-2b1b-4370-ce8e-4da5dfdc0f95"
X.dtypes
# + id="QSna9_kTcDx8"
cat_cols = X.columns[X.dtypes == 'object']
num_cols = X.columns[(X.dtypes == 'float64') | (X.dtypes == 'int64')]
# + id="XVAaLjGsgCPq" colab={"base_uri": "https://localhost:8080/"} outputId="8b8ca360-7427-4b07-8aca-de506667bf4b"
num_cols
# + id="V784I6eGgAH4" colab={"base_uri": "https://localhost:8080/"} outputId="2f3b6358-b3fe-414f-d64b-89ff93f76789"
cat_cols
# + [markdown] id="77OjmSl9i0Nf"
# Split dataset
# + id="Uay8Md5li0Nh" colab={"base_uri": "https://localhost:8080/"} outputId="b953666c-8679-414c-90e1-fb45fce01c48"
# Splite data into training and testing
# 100 -> 75:y=1, 25:y=0
# training(80): 60 y=1; 20 y=0
# testing(20): 15 y=1; 5 y=0
from sklearn import model_selection
# Reserve 25% for testing
# stratify example:
# 100 -> y: 80 '0', 20 '1' -> 4:1
# 80% training 64: '0', 16:'1' -> 4:1
# 20% testing 16:'0', 4: '1' -> 4:1
X_train, X_test, y_train, y_test = model_selection.train_test_split(X, y, test_size=0.25, stratify = y, random_state=1) #stratified sampling
print('training data has ' + str(X_train.shape[0]) + ' observation with ' + str(X_train.shape[1]) + ' features')
print('test data has ' + str(X_test.shape[0]) + ' observation with ' + str(X_test.shape[1]) + ' features')
# + [markdown] id="ODAexq7nyppM"
# * 10000 -> 8000 '0' + 2000 '1'
#
# * 25% test 75% training
# ---
# without stratified sampling:
# • extreme case:
# ---
# 1. testing: 2000 '1' + 500 '0'
# 2. training: 7500 '0'
# ---
# with stratified sampling:
# 1. testing: 2000 '0' + 500 '1'
# 2. training: 6000 '0' + 1500 '1'
#
# + [markdown] id="JMTIEpY7IfPp"
# Read more for handling [categorical feature](https://github.com/scikit-learn-contrib/categorical-encoding), and there is an awesome package for [encoding](http://contrib.scikit-learn.org/category_encoders/).
# + id="apcEXk0Eh978" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="0fcf1ce1-afd1-45e3-80cc-e93af48837cc"
X_train.head()
# + id="iWEjSK9leWyH" colab={"base_uri": "https://localhost:8080/"} outputId="559251db-dd8c-4dc3-9e8a-005c7bd58ed2"
# One hot encoding
# another way: get_dummies
from sklearn.preprocessing import OneHotEncoder
def OneHotEncoding(df, enc, categories):
transformed = pd.DataFrame(enc.transform(df[categories]).toarray(), columns=enc.get_feature_names(categories))
return pd.concat([df.reset_index(drop=True), transformed], axis=1).drop(categories, axis=1)
categories = ['Geography']
enc_ohe = OneHotEncoder()
enc_ohe.fit(X_train[categories])
X_train = OneHotEncoding(X_train, enc_ohe, categories)
X_test = OneHotEncoding(X_test, enc_ohe, categories)
# + colab={"base_uri": "https://localhost:8080/", "height": 268} id="txvvIi_Y1G1_" outputId="f4f035bd-71a2-4a0e-fd7f-e55783876085"
X_train.head()
# + id="6f3JCVj3ouWj"
# Ordinal encoding
from sklearn.preprocessing import OrdinalEncoder
categories = ['Gender']
enc_oe = OrdinalEncoder()
enc_oe.fit(X_train[categories])
X_train[categories] = enc_oe.transform(X_train[categories])
X_test[categories] = enc_oe.transform(X_test[categories])
# + id="qFaSPAc7sP8W" colab={"base_uri": "https://localhost:8080/", "height": 250} outputId="513b2736-ebf3-41c5-b474-c5416e1d8444"
X_train.head()
# + [markdown] id="vecyDzf8eXgg"
# Standardize/Normalize Data
# + id="JuPhtUkJi0NW"
# Scale the data, using standardization
# standardization (x-mean)/std
# normalization (x-x_min)/(x_max-x_min) ->[0,1]
# 1. speed up gradient descent
# 2. same scale
# 3. algorithm requirments
# for example, use training data to train the standardscaler to get mean and std
# apply mean and std to both training and testing data.
# fit_transform does the training and applying, transform only does applying.
# Because we can't use any info from test, and we need to do the same modification
# to testing data as well as training data
# https://scikit-learn.org/stable/auto_examples/preprocessing/plot_all_scaling.html#sphx-glr-auto-examples-preprocessing-plot-all-scaling-py
# https://scikit-learn.org/stable/modules/preprocessing.html
# min-max example: (x-x_min)/(x_max-x_min)
# [1,2,3,4,5,6,100] -> fit(min:1, max:6) (scalar.min = 1, scalar.max = 6) -> transform [(1-1)/(6-1),(2-1)/(6-1)..]
# scalar.fit(train) -> min:1, max:100
# scalar.transform(apply to x) -> apply min:1, max:100 to X_train
# scalar.transform -> apply min:1, max:100 to X_test
# scalar.fit -> mean:1, std:100
# scalar.transform -> apply mean:1, std:100 to X_train
# scalar.transform -> apply mean:1, std:100 to X_test
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(X_train[num_cols])
X_train[num_cols] = scaler.transform(X_train[num_cols])
X_test[num_cols] = scaler.transform(X_test[num_cols])
# + colab={"base_uri": "https://localhost:8080/", "height": 270} id="UydCNgAZ9ajK" outputId="48eebf24-3cca-40c3-e3a2-a4adb202e383"
X_train.head()
# + [markdown] id="q3x9ySX_i0Nd"
# # Part 3: Model Training and Result Evaluation
# + [markdown] id="c4UTtCQTi0Nl"
# ### Part 3.1: Model Training
# + id="EAhSxINLi0Nl"
#@title build models
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.linear_model import LogisticRegression
# Logistic Regression
classifier_logistic = LogisticRegression()
# K Nearest Neighbors
classifier_KNN = KNeighborsClassifier()
# Random Forest
classifier_RF = RandomForestClassifier()
# + id="Av0IRSoBQ3pe" colab={"base_uri": "https://localhost:8080/"} outputId="21cf9504-6d3d-442c-8e82-752dae55b0a1"
# Train the model
classifier_logistic.fit(X_train, y_train)
# + id="EiLuzUDJRBNi" colab={"base_uri": "https://localhost:8080/"} outputId="74a5faba-936f-4255-c50d-ff5e10bda714"
# Prediction of test data
classifier_logistic.predict(X_test)
# + id="XjMV04mKRJ30" colab={"base_uri": "https://localhost:8080/"} outputId="ee1722ca-d69e-4710-9aaa-5076d1f59290"
# Accuracy of test data
classifier_logistic.score(X_test, y_test)
# + [markdown] id="7J-23z78i0Ns"
# ### Part 3.2: Use Grid Search to Find Optimal Hyperparameters
# alternative: random search
# + id="lqTL8zwnANHr"
#Loss/cost function --> (wx + b - y) ^2 + ƛ * |w| --> ƛ is a hyperparameter
# + id="Hpe9PEAAi0Nt"
from sklearn.model_selection import GridSearchCV
# helper function for printing out grid search results
def print_grid_search_metrics(gs):
print ("Best score: " + str(gs.best_score_))
print ("Best parameters set:")
best_parameters = gs.best_params_
for param_name in sorted(best_parameters.keys()):
print(param_name + ':' + str(best_parameters[param_name]))
# + [markdown] id="qvYo9I5Ti0Nv"
# #### Part 3.2.1: Find Optimal Hyperparameters - LogisticRegression
# + id="wOc48syxi0Nx" colab={"base_uri": "https://localhost:8080/"} outputId="10277260-a8f0-4199-bdfe-630dd437ad03"
# Possible hyperparamter options for Logistic Regression Regularization
# Penalty is choosed from L1 or L2
# C is the 1/lambda value(weight) for L1 and L2
# solver: algorithm to find the weights that minimize the cost function
# ('l1', 0.01)('l1', 0.05) ('l1', 0.1) ('l1', 0.2)('l1', 1)
# ('12', 0.01)('l2', 0.05) ('l2', 0.1) ('l2', 0.2)('l2', 1)
parameters = {
'penalty':('l2','l1'),
'C':(0.01, 0.05, 0.1, 0.2, 1)
}
Grid_LR = GridSearchCV(LogisticRegression(solver='liblinear'),parameters, cv=5)
Grid_LR.fit(X_train, y_train)
# + id="nN5rU0e-i0N1" colab={"base_uri": "https://localhost:8080/"} outputId="4cefe5e3-df6f-451b-92eb-34f7ad21a48b"
# the best hyperparameter combination
# C = 1/lambda
print_grid_search_metrics(Grid_LR)
# + id="TtkDsXgui0N3"
# best model
best_LR_model = Grid_LR.best_estimator_
# + colab={"base_uri": "https://localhost:8080/"} id="epIrhEO3DimN" outputId="6c8bb931-449c-478b-9a18-e73a29e8828a"
best_LR_model.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="zF6EfOVaDZvH" outputId="64340637-d907-42dc-d174-905815f2c2b2"
best_LR_model.score(X_test, y_test)
# + id="Jy5hU5KymPT7" colab={"base_uri": "https://localhost:8080/", "height": 280} outputId="55d69111-7d0d-40b6-fa2f-175625734da7"
LR_models = pd.DataFrame(Grid_LR.cv_results_)
res = (LR_models.pivot(index='param_penalty', columns='param_C', values='mean_test_score')
)
_ = sns.heatmap(res, cmap='viridis')
# + [markdown] id="9u9YFedOi0N6"
# #### Part 3.2.2: Find Optimal Hyperparameters: KNN
# + id="o78422XVi0N6" colab={"base_uri": "https://localhost:8080/"} outputId="bca9caca-8550-436d-de1d-a6fc2b8bab1a"
# Possible hyperparamter options for KNN
# Choose k
parameters = {
'n_neighbors':[1,3,5,7,9]
}
Grid_KNN = GridSearchCV(KNeighborsClassifier(),parameters, cv=5)
Grid_KNN.fit(X_train, y_train)
# + id="ydaRZVAIi0N_" colab={"base_uri": "https://localhost:8080/"} outputId="4a505d67-7761-4192-b053-03e25f6062f6"
# best k
print_grid_search_metrics(Grid_KNN)
# + id="Nq_qfVpXUJcx"
best_KNN_model = Grid_KNN.best_estimator_
# + [markdown] id="nKn_oKLSi0OB"
# #### Part 3.2.3: Find Optimal Hyperparameters: Random Forest
# + id="NniAZIPfi0OC" colab={"base_uri": "https://localhost:8080/"} outputId="e23e6356-54f7-48af-9c33-092c8fe7fd30"
# Possible hyperparamter options for Random Forest
# Choose the number of trees
parameters = {
'n_estimators' : [60,80,100],
'max_depth': [1,5,10]
}
Grid_RF = GridSearchCV(RandomForestClassifier(),parameters, cv=5)
Grid_RF.fit(X_train, y_train)
# + id="ScPiI-Bfi0OE" colab={"base_uri": "https://localhost:8080/"} outputId="96765a7f-a210-4384-d6b4-8ae90d1a2fd6"
# best number of tress
print_grid_search_metrics(Grid_RF)
# + id="xJgfri_Mi0OG"
# best random forest
best_RF_model = Grid_RF.best_estimator_
# + id="LjmwORLjqCwZ" colab={"base_uri": "https://localhost:8080/"} outputId="335faa80-2acc-4386-ecba-7209bc51d0a3"
best_RF_model
# + [markdown] id="xxDAOrGIi0OI"
# ####Part 3.3: Model Evaluation - Confusion Matrix (Precision, Recall, Accuracy)
#
# class of interest as positive
#
# TP: correctly labeled real churn
#
# Precision(PPV, positive predictive value): tp / (tp + fp);
# Total number of true predictive churn divided by the total number of predictive churn;
# High Precision means low fp, not many return users were predicted as churn users.
#
#
# Recall(sensitivity, hit rate, true positive rate): tp / (tp + fn)
# Predict most postive or churn user correctly. High recall means low fn, not many churn users were predicted as return users.
# + id="o-tP94iFi0OI"
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
# calculate accuracy, precision and recall, [[tn, fp],[]]
def cal_evaluation(classifier, cm):
tn = cm[0][0]
fp = cm[0][1]
fn = cm[1][0]
tp = cm[1][1]
accuracy = (tp + tn) / (tp + fp + fn + tn + 0.0)
precision = tp / (tp + fp + 0.0)
recall = tp / (tp + fn + 0.0)
print (classifier)
print ("Accuracy is: " + str(accuracy))
print ("precision is: " + str(precision))
print ("recall is: " + str(recall))
print ()
# print out confusion matrices
def draw_confusion_matrices(confusion_matricies):
class_names = ['Not','Churn']
for cm in confusion_matrices:
classifier, cm = cm[0], cm[1]
cal_evaluation(classifier, cm)
# + id="OpSGaN49i0OL" colab={"base_uri": "https://localhost:8080/"} outputId="d89c5aac-bc3a-4c0e-c86a-b499161144cd"
# Confusion matrix, accuracy, precison and recall for random forest and logistic regression
confusion_matrices = [
("Random Forest", confusion_matrix(y_test,best_RF_model.predict(X_test))),
("Logistic Regression", confusion_matrix(y_test,best_LR_model.predict(X_test))),
("K nearest neighbor", confusion_matrix(y_test, best_KNN_model.predict(X_test)))
]
draw_confusion_matrices(confusion_matrices)
# + [markdown] id="OvHlyhPBi0OT"
# ### Part 3.4: Model Evaluation - ROC & AUC
# + [markdown] id="jx_3XkgKi0OW"
# RandomForestClassifier, KNeighborsClassifier and LogisticRegression have predict_prob() function
# + [markdown] id="-Os_ZLTvi0OX"
# #### Part 3.4.1: ROC of RF Model
# + id="UypvQMVBi0OY"
from sklearn.metrics import roc_curve
from sklearn import metrics
# Use predict_proba to get the probability results of Random Forest
y_pred_rf = best_RF_model.predict_proba(X_test)[:, 1]
fpr_rf, tpr_rf, _ = roc_curve(y_test, y_pred_rf)
# + id="RyADmHiPTtln" colab={"base_uri": "https://localhost:8080/"} outputId="b7b757b0-90a5-449d-cf18-439f03b58b06"
best_RF_model.predict_proba(X_test)
# + id="s3PR-PdPi0Ob" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="a7dd517a-760e-4583-9273-aea2816e0ac6"
# ROC curve of Random Forest result
import matplotlib.pyplot as plt
plt.figure(1)
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr_rf, tpr_rf, label='RF')
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
plt.title('ROC curve - RF model')
plt.legend(loc='best')
plt.show()
# + id="R89IUMYDi0Oe" colab={"base_uri": "https://localhost:8080/"} outputId="6713cdc6-b405-4dbe-855d-942daedc7ac5"
from sklearn import metrics
# AUC score
metrics.auc(fpr_rf,tpr_rf)
# + [markdown] id="-1DVqnJVi0Oh"
# #### Part 3.4.1: ROC of LR Model
# + id="t-q5XJPoi0Oi"
# Use predict_proba to get the probability results of Logistic Regression
y_pred_lr = best_LR_model.predict_proba(X_test)[:, 1]
fpr_lr, tpr_lr, thresh = roc_curve(y_test, y_pred_lr)
# + id="zc4k8gUYcpNE" colab={"base_uri": "https://localhost:8080/"} outputId="6814a897-86a8-40b9-9a9d-9a2ed3ea33e1"
best_LR_model.predict_proba(X_test)
# + id="KZSrN-1Mi0Ok" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="c110ec4a-31e5-448f-82d8-f3ce81a7eef8"
# ROC Curve
plt.figure(1)
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr_lr, tpr_lr, label='LR')
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
plt.title('ROC curve - LR Model')
plt.legend(loc='best')
plt.show()
# + id="LHAyxishi0On" colab={"base_uri": "https://localhost:8080/"} outputId="7c63cb46-7430-4194-c215-54041abec938"
# AUC score
metrics.auc(fpr_lr,tpr_lr)
# + [markdown] id="gHHurD8Ii0Oq"
# # Part 4: Model Extra Functionality
# + [markdown] id="dSx4TPO-i0Or"
# ### Part 4.1: Logistic Regression Model
# + [markdown] id="BtLHUixoi0Ot"
# The corelated features that we are interested in
# + id="mbNTNeb7saCy" colab={"base_uri": "https://localhost:8080/", "height": 305} outputId="73d8027e-a19e-42c1-8005-9025859fc06c"
X_with_corr = X.copy()
X_with_corr = OneHotEncoding(X_with_corr, enc_ohe, ['Geography'])
X_with_corr['Gender'] = enc_oe.transform(X_with_corr[['Gender']])
X_with_corr['SalaryInRMB'] = X_with_corr['EstimatedSalary'] * 6.4
X_with_corr.head()
# + id="cQaXOIsUi0Ou" colab={"base_uri": "https://localhost:8080/"} outputId="a1d4f952-14e5-4ba0-e676-bcccd6cab637"
# add L1 regularization to logistic regression
# check the coef for feature selection
scaler = StandardScaler()
X_l1 = scaler.fit_transform(X_with_corr)
LRmodel_l1 = LogisticRegression(penalty="l1", C = 0.04, solver='liblinear')
LRmodel_l1.fit(X_l1, y)
indices = np.argsort(abs(LRmodel_l1.coef_[0]))[::-1]
print ("Logistic Regression (L1) Coefficients")
for ind in range(X_with_corr.shape[1]):
print ("{0} : {1}".format(X_with_corr.columns[indices[ind]],round(LRmodel_l1.coef_[0][indices[ind]], 4)))
# + id="majifZZqi0O9" colab={"base_uri": "https://localhost:8080/"} outputId="9144702e-dc89-461d-fd6d-1d99cdd7cdd8"
# add L2 regularization to logistic regression
# check the coef for feature selection
np.random.seed()
scaler = StandardScaler()
X_l2 = scaler.fit_transform(X_with_corr)
LRmodel_l2 = LogisticRegression(penalty="l2", C = 0.1, solver='liblinear', random_state=42)
LRmodel_l2.fit(X_l2, y)
LRmodel_l2.coef_[0]
indices = np.argsort(abs(LRmodel_l2.coef_[0]))[::-1]
print ("Logistic Regression (L2) Coefficients")
for ind in range(X_with_corr.shape[1]):
print ("{0} : {1}".format(X_with_corr.columns[indices[ind]],round(LRmodel_l2.coef_[0][indices[ind]], 4)))
# + [markdown] id="uqs41ydLi0O_"
# ### Part 4.2: Random Forest Model - Feature Importance Discussion
# + id="z6HNnaBRvE1T" colab={"base_uri": "https://localhost:8080/", "height": 305} outputId="7ff4f758-a7b0-4ffa-f2df-e45995c7b089"
X_RF = X.copy()
X_RF = OneHotEncoding(X_RF, enc_ohe, ['Geography'])
X_RF['Gender'] = enc_oe.transform(X_RF[['Gender']])
X_RF.head()
# + id="MPxUM2lei0PA" colab={"base_uri": "https://localhost:8080/"} outputId="3f2614df-4980-45af-e43d-a34460390fd7"
# check feature importance of random forest for feature selection
forest = RandomForestClassifier()
forest.fit(X_RF, y)
importances = forest.feature_importances_
indices = np.argsort(importances)[::-1]
# Print the feature ranking
print("Feature importance ranking by Random Forest Model:")
for ind in range(X.shape[1]):
print ("{0} : {1}".format(X_RF.columns[indices[ind]],round(importances[indices[ind]], 4)))
| Supervised_Learning_Project_Bank_Customer_Churn_Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import plotly.express as px
from jupyter_dash import JupyterDash
import dash_daq as daq
import dash_core_components as dcc
import dash_html_components as html
from dash.dependencies import Input, Output
import plotly.express as px
import pandas as pd
from plotly.subplots import make_subplots
df = pd.read_csv("../data/input-magnitude.csv", decimal=".")
dfe = pd.read_csv("../data/input-extra.csv", decimal=".")
disaster = df["Disaster_Type"].unique()
YEARS = df["Decade"].unique()
external_stylesheets = ['https://codepen.io/anon/pen/mardKv.css']
app = JupyterDash(__name__, external_stylesheets=external_stylesheets)
# +
app.layout = html.Div([
dcc.Dropdown(
id="dropdown",
options=[{"label": x, "value": x} for x in disaster],
value=disaster[0],
clearable=False,
style={'backgroundColor': 'black'}
),
html.Div(
id="slider-container",
children=[
html.P(
id="slider-text",
children="Drag the slider to change the year:",
),
dcc.RangeSlider(
id="years-slider",
min=1900,
max=2090,
step=10,
value=[1900, 1950],
marks={
str(year): {
"label": str(year),
"style": {"color": "#7fafdf"},
}
for year in YEARS
},
),
],
),
html.Div(
children=[
daq.ToggleSwitch(
id='Impact-Selector',
label=['Human', 'Financial'],
value=False,
style={
'width': '100%',
'marginTop': '30px',
'marginBottom': '30px'},
theme={
'dark': False,
'detail': '#4525F2',
'primary': '#4525F2',
'secondary': '#F2F4F8',
}
),
],
),
html.Br(),
html.Div(
dcc.Graph(id="histogram"),
),
html.Div(
dcc.Graph(id="bc_DOHI"),
),
html.Div(
dcc.Graph(id="bc_ImpactDisasterType"),
),
])
@app.callback(
Output("histogram", "figure"),
Output("bc_DOHI", "figure"),
Output("bc_ImpactDisasterType", "figure"),
Input("dropdown", "value"),
Input("years-slider", "value"),
Input("Impact-Selector", "value")
)
def update_charts(disaster, year, ImpactType):
c = dfe.groupby(['Decade'])['°C'].mean().reset_index()
df['Temperature'] = df.Decade.map(c.set_index('Decade')['°C'])
is_disaster = df["Disaster_Type"] == disaster
df_disaster = df[is_disaster]
if not ImpactType:
# color = "reds"
impact_type = 'Human_Impact'
else:
# color = "purples"
impact_type = 'Financial_Impact'
# Si on veut définir toutes les couleurs une par une nous même:
# color_codes= ['#CCFFFF','#CCCCFF','#CC99FF','#009999','#0033FF','#003333',
# '#9900CC','#FFFF33','#339966','#CC6666','#996633','#009900','#6666FF','#330033',
# '#FF3333','#FFCCFF','#33FF99','#33FF99','#9999FF','#CC3300','#3300CC','#9999FF']
# color={}
# for i in range(21):
# color[i] = color_codes[i]
df_fig = df_disaster.query('Decade >=@year[0] and Decade <@year[1]')
bins = int((int(year[1]) - int(year[0])) / 10)
fig = px.histogram(df_fig,
x="Decade",
y=impact_type,
color="UN_Geosheme_Subregion",
# color_discrete_sequence = color,
template='plotly',
nbins=bins,
# barmode="stack",
title='Evolution of {0} caused by {1} per Regions'.format(impact_type, disaster),
# animation_frame="Decade",
labels={'x': 'Decade', 'y': 'Total Financial Impact', 'color': 'Region'},
)
# fig.update_xaxes(type='category')
fig.update_layout(xaxis={'categoryorder': 'total descending'})
subfig = make_subplots(specs=[[{"secondary_y": True}]])
fig2 = px.histogram(df_fig,
x="Decade",
y="DO",
template='plotly',
# barmode="stack",
color_discrete_sequence={0: '#CC0000'},
nbins=bins,
title='Evolution of disaster occurence and human impact worldwide',
# animation_frame="Decade", #ne autoscale pas malheureusement
labels={'Decade': 'Decade', 'DO': 'Disaster Occurence'},
)
fig2.update_xaxes(type='category')
fig3 = px.line(df_fig, x="Decade", y="Temperature", labels={'°C': 'Average Temperature'})
fig3.update_traces(yaxis="y2", showlegend=True, name='Temperatures', line_color='black')
subfig.add_traces(fig2.data + fig3.data)
subfig.update_xaxes(type='category')
subfig.layout.xaxis.title = "Decades"
subfig.layout.yaxis.title = "Occurences"
subfig.layout.yaxis2.title = "Temperatures"
subfig.layout.title = "{0} Occurence vs Temperature".format(disaster)
fig4 = px.histogram(df.query('Decade >=@year[0] and Decade <@year[1]'),
x="Decade",
y=impact_type,
color="Disaster_Type",
color_discrete_sequence={0: '#FFAE5D', 1: '#C5EBFD', 2: '#B561F4'},
template='plotly',
nbins=bins,
title='Total {0} per Disaster Type'.format(impact_type),
# animation_frame="Decade",
labels={'x': 'Decade', 'y': 'Total Financial Impact', 'color': 'Disaster Type'},
)
fig4.update_xaxes(type='category')
return fig, subfig, fig4
if __name__ == '__main__':
app.run_server(mode='inline')
# -
| Dashboard/charts/charts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # unconstrained
pwd
# ### Unconstrained growth of a population
# +
#This code performs the necessary steps to prepare to plot.
from pylab import * #Import plotting module matplotlib as well as other modules
#into the global namespace.
#Set the default plot to be inline rather than a new window.
# %matplotlib inline
import inspect #This allows us to print the source code in this notebook.
from unconstrained import * #Import the custom functions from this project.
goldenratio=1/2*(1+sqrt(5)) #The next few lines are used for the size of plots
fsx=7 #Width (in inches) for the figures.
fsy=fsx/goldenratio #Height (in inches) for the figures.
# + jupyter={"outputs_hidden": true}
#Simulate
timeLst, populationLst = unconstrained()
# -
#Graph
figure(figsize=(fsx, fsy))
plot(timeLst, populationLst, 'b-')
xlabel('Time')
ylabel('population')
title('Population vs Time')
show()
| resources/CalculusReview/unconstrained.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch, torchvision
from torchvision import transforms
import matplotlib.pyplot as plt
from torch import nn, optim
# Download MNIST manually using 'wget' then uncompress the file
# !wget www.di.ens.fr/~lelarge/MNIST.tar.gz
# !tar -zxvf MNIST.tar.gz
transform = transforms.Compose([transforms.ToTensor()])
trainset = torchvision.datasets.MNIST(root='./', train=True, transform=transform, download=True)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
class Net(nn.Module):
def __init__(self):
super().__init__()
self.hidden = nn.Linear(28*28, 512)
self.output = nn.Linear(512, 10)
self.sigmoid = nn.Sigmoid()
self.softmax = nn.LogSoftmax(dim=1)
def forward(self, x):
x = self.hidden(x)
x = self.sigmoid(x)
x = self.output(x)
x = self.softmax(x)
return x
model = Net()
model
criterion = nn.NLLLoss()
images, labels = next(iter(trainloader))
images = images.view(images.shape[0], -1)
logits = model(images)
loss = criterion(logits, labels)
loss
loss.backward()
optimizer = optim.SGD(model.parameters(), lr=0.01)
optimizer.step()
| federated_learning/Introduction_to_PyTorch/7_Single_Epoch_Training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os, sys
sys.path.insert(1, os.path.join(sys.path[0], '..'))
import database
from snowballing.graph import Graph
from snowballing.operations import reload
reload()
Graph("graph", delayed=False)
| snowballing/example/notebooks/CitationGraph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Ambulance Routing Ithaca Dataset
#
# One potential application of reinforcement learning involves positioning a server or servers (in this case an ambulance) in an optimal way geographically to respond to incoming calls while minimizing the distance traveled by the servers. This is closely related to the [k-server problem](https://en.wikipedia.org/wiki/K-server_problem), where there are $k$ servers stationed in a space that must respond to requests arriving in that space in such a way as to minimize the total distance traveled.
#
# The ambulance routing problem addresses the problem by modeling an environment where there are ambulances stationed at locations, and calls come in that one of the ambulances must be sent to respond to. The goal of the agent is to minimize both the distance traveled by the ambulances between calls and the distance traveled to respond to a call by optimally choosing the locations to station the ambulances. The ambulance environment has been implemented in two different ways; as a 1-dimensional number line $[0,1]$ along which ambulances will be stationed and calls will arrive, and a graph with nodes where ambulances can be stationed and calls can arrive, and edges between the nodes that ambulances travel along.
# In this notebook, we walk through the Ambulance Routing problem with on a graph. There is a structured as a graph of nodes $V$ with edges between the nodes $E$. Each node represents a location where an ambulance could be stationed or a call could come in. The edges between nodes are undirected and have a weight representing the distance between those two nodes.
#
# The nearest ambulance to a call is determined by computing the shortest path from each ambulance to the call, and choosing the ambulance with the minimum length path. The calls arrive using a prespecified iid probability distribution. The default is for the probability of call arrivals to be evenly distributed over all the nodes; however, the user can also choose different probabilities for each of the nodes that a call will arrive at that node. For example, in the following graph the default setting would be for each call to have a 0.25 probability of arriving at each node, but the user could instead specify that there is a 0.1 probability of a call at node 0, and a 0.3 probability of a call arriving at each of the other three nodes.
#
#
# We do a single experiment using the underlying graph taken as the Ithaca city, with arrivals from a historical police call dataset.
# This is an example with 2 ambulances (k=2) at nodes 2 and 3.
# 
#
#
# ### Package Installation
#
# +
import or_suite
import numpy as np
import copy
import os
from stable_baselines3.common.monitor import Monitor
from stable_baselines3 import PPO
from stable_baselines3.ppo import MlpPolicy
from stable_baselines3.common.env_util import make_vec_env
from stable_baselines3.common.evaluation import evaluate_policy
import pandas as pd
import gym
# -
# ### Experimental Parameters
#
# The ambulance routing problem has several experiment parameters
# * The parameter `epLen`, an int, represents the length of each episode
# * `nEps` is an int representing the number of episodes. The default is set to 2.
# * `numIters`, an int, is the number of iterations. Here it is set to 50.
# * `seed` allows random numbers to be generated.
# * `dirPath`, a string, is the location where the data files are stored.
# * `deBug`, a bool, prints information to the command line when set true.
# * `save_trajectory`, a bool, saves the trajectory information of the ambulance when set to true.
# * `render` renders the algorithm when set to true.
# * `pickle` is a bool that saves the information to a pickle file when set to true.
#
# Along with experiment parameters, the ambulance problem has several environmental parameters.
# * `alpha`, a float controlling the proportional difference between the cost to move ambulances in between calls and the cost to move the ambulance to respond to a call. If `alpha` is 0, there is no cost to move between calls. If `alpha` is one, there is no cost to respond to calls.
# * `num_ambulance`, an int which represents the number of ambulances in the system.
#
# +
# Experiment Parameters
# Getting out configuration parameter for the environment
CONFIG = or_suite.envs.env_configs.ambulance_graph_ithaca_config
# Specifying training iteration, epLen, number of episodes, and number of iterations
epLen = CONFIG['epLen']
nEps = 2
numIters = 50
# Configuration parameters for running the experiment
DEFAULT_SETTINGS = {'seed': 1,
'recFreq': 1,
'dirPath': '../data/ambulance/',
'deBug': False,
'nEps': nEps,
'numIters': numIters,
'saveTrajectory': True, # save trajectory for calculating additional metrics
'epLen' : 5,
'render': False,
'pickle': False # indicator for pickling final information
}
alpha = CONFIG['alpha']
num_ambulance = CONFIG['num_ambulance']
ambulance_env = gym.make('Ambulance-v1', config=CONFIG)
mon_env = Monitor(ambulance_env)
# -
# ### Specifying Agent
# We have several heuristics implemented for each of the environments defined, in addition to a `Random` policy, and some `RL discretization based` algorithms.
#
# The `Stable` agent only moves ambulances when responding to an incoming call and not in between calls. This means the policy $\pi$ chosen by the agent for any given state $X$ will be $\pi_h(X) = X$
#
# The `Median` agent takes a list of all past call arrivals sorted by arrival location, and partitions it into $k$ quantiles where $k$ is the number of ambulances. The algorithm then selects the middle data point in each quantile as the locations to station the ambulances.
# The `Mode` agent chooses to stattion the ambulances at the nodes where the most calls have come in the past. This uses the policy $\pi$ chosen by a state $X$ will be $\pi_h(X) = $`mode`
#
agents = { 'SB PPO': PPO(MlpPolicy, mon_env, gamma=1, verbose=0, n_steps=epLen),
'Random': or_suite.agents.rl.random.randomAgent(),
'Stable': or_suite.agents.ambulance.stable.stableAgent(CONFIG['epLen']),
'Mode': or_suite.agents.ambulance.mode_graph.modeAgent(CONFIG['epLen']),
'Median': or_suite.agents.ambulance.median_graph.medianAgent(CONFIG['epLen'], CONFIG['edges'], CONFIG['num_ambulance']),
}
# Run the different heuristics in the environment
# ### Running Algorithm
#
# +
path_list_line = []
algo_list_line = []
path_list_radar = []
algo_list_radar= []
for agent in agents:
print(agent)
DEFAULT_SETTINGS['dirPath'] = '../data/ambulance_metric_'+str(agent)+'_'+str(num_ambulance)+'_'+str(alpha)+'/'
if agent == 'SB PPO':
or_suite.utils.run_single_sb_algo(mon_env, agents[agent], DEFAULT_SETTINGS)
elif agent == 'AdaQL' or agent == 'Unif QL' or agent == 'AdaMB' or agent == 'Unif MB':
or_suite.utils.run_single_algo_tune(ambulance_env, agents[agent], scaling_list, DEFAULT_SETTINGS)
else:
or_suite.utils.run_single_algo(ambulance_env, agents[agent], DEFAULT_SETTINGS)
path_list_line.append('../data/ambulance_metric_'+str(agent)+'_'+str(num_ambulance)+'_'+str(alpha))
algo_list_line.append(str(agent))
if agent != 'SB PPO':
path_list_radar.append('../data/ambulance_metric_'+str(agent)+'_'+str(num_ambulance)+'_'+str(alpha))
algo_list_radar.append(str(agent))
fig_path = '../figures/'
fig_name = 'ambulance_metric'+'_'+str(num_ambulance)+'_'+str(alpha)+'_line_plot'+'.pdf'
or_suite.plots.plot_line_plots(path_list_line, algo_list_line, fig_path, fig_name, int(nEps / 40)+1)
additional_metric = {}
fig_name = 'ambulance_metric'+'_'+str(num_ambulance)+'_'+str(alpha)+'_radar_plot'+'.pdf'
or_suite.plots.plot_radar_plots(path_list_radar, algo_list_radar,
fig_path, fig_name,
additional_metric
)
# -
# Here we see with a quick set-up that the best-performing algorithm in this limited data regime is the Mode algorithm, essentially putting the ambulances at the estimated mode from the observed data thus far.
| examples/ambulance_ithaca_environment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## SQL ALCHEMY
#
from sqlalchemy import create_engine,or_
import pandas as pd
from sqlalchemy.orm import sessionmaker
from sqlalchemy.ext.declarative import declarative_base
SERVER = 'DESKTOP-V20Q7QI'
DATABASE = 'sample'
DRIVER = 'SQL Server Native Client 11.0'
DATABASE_CONNECTION = f'mssql://@{SERVER}/{DATABASE}?driver={DRIVER}'
engine = create_engine(DATABASE_CONNECTION)
connection = engine.connect()
# ### running a query to check the connection
data = pd.read_sql_query("select * from [sample].[dbo].[Market_Basket_Optimisation$]", connection)
data
data.dtypes
# ### session
# +
Session = sessionmaker(bind=engine)
session = Session()
Base = declarative_base()
# -
# #### basic queries
# +
from sqlalchemy import Column, String, Integer
# create a table
class Student(Base):
__tablename__ = 'student'
id = Column(Integer, primary_key=True)
name = Column(String(50))
age = Column(Integer)
grade = Column(String(50))
Base.metadata.create_all(engine)
# table is created
# -
# Insert data
student1 = Student(name="vijaya",age=24, grade='Masters') # creating instance
session.add(student1) # adding data to session ( single record)
session.commit() # commit changes to DB
# +
# Insert multiple data
student2 = Student(name="vijaya1",age=23, grade='Bachelors') # creating instance
student3 = Student(name="vijaya2",age=22, grade='Highschool') # creating instance
session.add_all([student2, student3]) # adding data to session ( multiple record)
session.commit() # commit changes to DB
# +
# Query the table - read
# get all the data
students = session.query(Student)
for student in students:
print(student.name, student.age, student.grade)
# +
# get data in order
students = session.query(Student).order_by(Student.name)
for student in students:
print(student.name)
# +
# Get data by filtering
student = session.query(Student).filter(Student.name=="vijaya").first()
for student in students:
print(student.name)
print(student.name, student.age)
# +
student = session.query(Student).filter(or_(Student.name=="vijaya", Student.name=='vijaya1'))
for student in students:
print(student.name, student.age)
# +
# count the results
student_count = session.query(Student).filter(or_(Student.name=="vijaya", Student.name=='vijaya1')).count()
print(student_count)
# +
# update data
student = session.query(Student).filter(Student.name=="vijaya").first()
student.name = "laxmi"
session.commit()
# +
# get all the data
students = session.query(Student)
for student in students:
print(student.name, student.age, student.grade)
# -
# Delete the data
student = session.query(Student).filter(Student.name=="vijaya1").first()
session.delete(student)
session.commit()
# +
# get all the data
students = session.query(Student)
for student in students:
print(student.name, student.age, student.grade)
# -
| SQL ALCHEMY.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# Generate spatial index for catchments based on their topology
#
# Save parent catchment info for every child catchment so that it can be quickly queried, O(n), eventually it is much faster with column value indices.
#
# 
#
# So, for every parent of a given catchment, a new feature is generated, where an id of its child catchment is saved. This way, all parent catchment ids can be found using a single query.
# Note, that rivers index is generated using QGIS: __Join attributes by location__ tool.
#
# Before join, HydroBASINS layer was converted to contain only a single field HYBAS_ID - changed to text to avoid int overflow. Used __Refactor field__ tool.
#
# hydro-engine\data\HydroBASINS\l05\hybas_lev05_v1c_id.dbf
# +
# %matplotlib inline
import glob
import os
import logging
import sys
import json
import math
import shapefile
import shapely.geometry, shapely.wkt
import shapely as sl
import numpy as np
import networkx as nx
import matplotlib.pyplot as plt
import pylab
# -
pylab.rcParams['figure.figsize'] = (17.0, 15.0)
logging.basicConfig(stream=sys.stderr, level=logging.INFO)
def generate_index(src, dst):
shp = shapefile.Reader(src)
# 1. fill directed graph
graph = nx.DiGraph()
list_hybas_id = [r[0] for r in shp.records()]
list_next_down = [r[1] for r in shp.records()]
list_shape_box = [s.bbox for s in shp.shapes()]
edges = zip(list_hybas_id, list_next_down)
edges = [e for e in edges if e[1] != 0]
edges = [e for e in edges if e[0] != 0]
graph.add_nodes_from([(hybas_id) for (hybas_id) in list_hybas_id])
graph.add_edges_from([(node_from, node_to) for (node_from, node_to) in edges])
# 2. traverse parent nodes
index = []
for r in zip(list_hybas_id, list_shape_box):
hybas_id = r[0]
bbox = r[1]
x0 = bbox[0]
x1 = bbox[2]
y0 = bbox[1]
y1 = bbox[3]
poly = [[[x0, y0], [x1, y0], [x1, y1], [x0, y1], [x0, y0]]]
parents = nx.bfs_predecessors(graph.reverse(), hybas_id)
for parent in zip(parents.keys(), parents.values()):
index.append([hybas_id, parent[0], parent[1], poly])
# endorheic
if len(parents) == 0:
index.append([hybas_id, 0, hybas_id, poly])
return index
# 3. write
print('{0} -> {1}'.format(src, dst))
print(len(shp.shapes()))
pass
# +
files = glob.glob('../data/HydroBASINS/*lev09*.shp')
import geojson
l = None
for src in files:
file_dir = os.path.dirname(src)
file_name = os.path.basename(src)
dst = '../data/HydroBASINS_indexed/' + file_name
print('Processing {0} ...'.format(src))
index = generate_index(src, dst)
w = shapefile.Writer(shapeType=shapefile.POLYGON)
w.field('HYBAS_ID', 'N', 16)
w.field('PARENT_FROM', 'N', 16)
w.field('PARENT_TO', 'N', 16)
features = []
for i in index:
w.poly(i[3])
w.record(i[0], i[1], i[2])
w.save(dst)
# -
w = shapefile.Writer(shapefile.POINT)
w.point(1,1)
w.point(3,1)
w.point(4,3)
w.point(2,2)
w.field('FIRST_FLD', 'N',16)
w.field('SECOND_FLD','C','40')
w.record(1,'Point')
w.record(1,'Point')
w.record(2,'Point')
w.record(3,'Point')
w.save('s')
# DONE
# +
shp = shapefile.Reader('../data/HydroBASINS\hybas_af_lev05_v1c.shp')
# 1. fill directed graph
graph = nx.DiGraph()
# 2. traverse all parent nodes
list_hybas_id = [r[0] for r in shp.records()]
list_next_down = [r[1] for r in shp.records()]
edges = zip(list_hybas_id, list_next_down)
edges = [e for e in edges if e[1] != 0]
edges = [e for e in edges if e[0] != 0]
graph.add_nodes_from([(hybas_id) for (hybas_id) in list_hybas_id])
graph.add_edges_from([(node_from, node_to) for (node_from, node_to) in edges])
# -
len(graph.nodes())
len(graph.edges())
# +
pos = nx. (nx.Graph(graph), iterations=50, scale=1.0)
nx.draw_networkx_nodes(graph, pos, node_color='b', alpha=0.85, node_size=10)
nx.draw_networkx_edges(graph, pos, alpha=0.2)
# Niger: https://code.earthengine.google.com/0a846953681587649ed9bc6deee974cf
start_node = 1050022420
# traverse up, DFS
parent_nodes = graph.subgraph(nx.dfs_predecessors(graph.reverse(), start_node))
nx.draw_networkx_edges(parent_nodes, pos, edge_color='r', alpha=0.2)
nx.draw_networkx_nodes(parent_nodes, pos, node_color='r', alpha=0.85, node_size=30)
nx.draw_networkx_nodes(graph.subgraph([start_node]), pos, node_color='r', alpha=0.85, node_size=30)
nx.draw_networkx_nodes(graph.subgraph(graph.predecessors(start_node)), pos, node_color='r', alpha=0.85, node_size=30)
# -
graph.predecessors(start_node)
predecesors = nx.bfs_predecessors(graph.reverse(), start_node)
list(predecesors.keys())
| notebooks/generate_index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# Disable warnings
import warnings
warnings.filterwarnings('ignore')
# ### Re-Run Set Up Code from Intro Notebook
# +
import datetime as dt
import json
import os
import urllib.request
import numpy as np
import pandas as pd
import tensorflow as tf
from sklearn.preprocessing import MinMaxScaler
import matplotlib.pyplot as plt
data_source = 'kaggle' # alphavantage or kaggle
if data_source == 'alphavantage':
api_key = '<KEY>'
# American Airlines stock market prices
ticker = 'AAL'
# JSON file with all the stock market data for AAL from the last 20 years
url_string = f'https://www.alphavantage.co/query?function=TIME_SERIES_DAILY&symbol={ticker}&outputsize=full&apikey={api_key}'
# Save data to this file
file_to_save = f'../data/raw/stock_market_data-{ticker}.csv'
# If you haven't already saved data,
# Go ahead and grab the data from the url
# And store date, low, high, volume, close, open values to a Pandas DataFrame
if not os.path.exists(file_to_save):
with urllib.request.urlopen(url_string) as url:
data = json.loads(url.read().decode())
# extract stock market data
data = data['Time Series (Daily)']
df = pd.DataFrame(columns=['Date', 'Low', 'High', 'Close', 'Open'])
for k, v in data.items():
date = dt.datetime.strptime(k, '%Y-%m-%d')
data_row = [date.date(), float(v['3. low']), float(v['2. high']), float(v['4. close']),
float(v['1. open'])]
df.loc[-1, :] = data_row
df.index = df.index + 1
print(f'Data saved to : {file_to_save}')
df.to_csv(file_to_save)
# If the data is already there, just load it from the CSV
else:
print('File already exists. Loading data from CSV')
df = pd.read_csv(file_to_save)
else:
# You will be using HP's data. Feel free to experiment with other data.
# But while doing so, be careful to have a large enough dataset and also pay attention to the data normalization
df = pd.read_csv(os.path.join('../data/external/Stocks', 'hpq.us.txt'), delimiter=',',
usecols=['Date', 'Open', 'High', 'Low', 'Close'])
print('Loaded data from the Kaggle repository')
# Sort DataFrame by date
df = df.sort_values('Date')
df.head()
# First calculate the mid prices from the highest and lowest
high_prices = df.loc[:, 'High'].to_numpy()
low_prices = df.loc[:, 'Low'].to_numpy()
mid_prices = (high_prices + low_prices) / 2.0
# Split data into training and test sets
train_data = mid_prices[:11000]
test_data = mid_prices[11000:]
# Scale the data to be between 0 and 1
# When scaling remember! You normalize both test and train data with respect to training data
# Because you are not supposed to have access to test data
scaler = MinMaxScaler()
train_data = train_data.reshape(-1, 1)
test_data = test_data.reshape(-1, 1)
# Train the Scaler with training data and smooth data
smoothing_window_size = 2500
for di in range(0, 10000, smoothing_window_size):
scaler.fit(train_data[di:di + smoothing_window_size, :])
train_data[di:di + smoothing_window_size, :] = scaler.transform(train_data[di:di + smoothing_window_size, :])
# You normalize the last bit of remaining data
scaler.fit(train_data[di + smoothing_window_size:, :])
train_data[di + smoothing_window_size:, :] = scaler.transform(train_data[di + smoothing_window_size:, :])
# Reshape both train and test data
train_data = train_data.reshape(-1)
# Normalize test data
test_data = scaler.transform(test_data).reshape(-1)
# Now perform exponential moving average smoothing
# So the data will have a smoother curve than the original ragged data
EMA = 0.0
gamma = 0.1
for ti in range(11000):
EMA = gamma * train_data[ti] + (1 - gamma) * EMA
train_data[ti] = EMA
# Used for visualization and test purposes
all_mid_data = np.concatenate([train_data, test_data], axis=0)
# -
# ## Intro to Long Short-Term Memory Models
#
# Long Short-Term Memory models are powerful time-series models which can make predictions an arbitrary number of steps (i.e., periods of time into the future). An LSTM module, or cell, is comprised of the following components:
# - Cell state ($c_t$) - This represents the internal memory of the cell which stores both short term memory and long-term memories.
# - Hidden state ($h_t$) - This is output state information calculated with respect to current input, previous hidden state and current cell input which you eventually use to predict the future stock market prices. Additionally, the hidden state can decide to only retrive the short or long-term or both types of memory stored in the cell state to make the next prediction.
# - Input gate ($i_t$) - Decides how much information from current input flows to the cell state.
# - Forget gate ($f_t$) - Decides how much information from the current input and the previous cell state flows into the current cell state.
# - Output gate ($o_t$) - Decides how much information from the current cell state flows into the hidden state, so that if needed LSTM can only pick the long-term memories or short-term memories and long-term memories.
#
# The image below illustrates the composition of an LSTM cell.
from IPython.display import HTML, display
display(HTML("<img src='img/lstm.png'>"))
# The equations for calculating each of the components are as follows:
# - $i_t=\sigma(W_{ix}x_t+W_{ih}h_{t-1}+b_i)$
# - $\tilde{c}_t=\sigma(W_{cx}x_t+W_{ch}h_{t-1}+b_c)$
# - $f_t=\sigma(W_{fx}x_t+W_{fh}h_{t-1}+b_f)$
# - $c_t=f_tc_{t-1}+i_t\tilde{c}_t$
# - $o_t=\sigma(W_{ox}x_t+W_{oh}h_{t-1}+b_o)$
# - $h_t=o_t\tanh{(c_t)}$
# ### Data Generation and Augmentation
#
# First, we need to create a data generator to train our LSTM model. The `.unroll_batches(...)` method will output a set of a specified number of batches of input data, ordered sequentially. Each batch of data will be of the specified size and will have a corresponding batch of output data.
#
# To make our model more robust, we will not make the output for $x_t$ always be $x_{t+1}$. Instead, we will randomly sample an output from the set $x_{t+1},\ x_{t+2},\ ...,\ x_{t+N}$ where $N$ is a small window size. In essence, we will randomly select an output for $x_t$ which can be any observation in the time series that comes after $x_t$ and that falls within the specified window of the series, which is of size $N$. Note that, we are assuming that $x_{t+1},\ x_{t+2},\ ...,\ x_{t+N}$ will be relatively close to each other in the series. The following image illustrates this data augmentation process.
display(HTML("<img src='img/batch.png'>"))
# +
class DataGeneratorSeq(object):
def __init__(self, prices, batch_size, num_unroll):
self._prices = prices
self._prices_length = len(self._prices) - num_unroll
self._batch_size = batch_size
self._num_unroll = num_unroll
self._segments = self._prices_length // self._batch_size
self._cursor = [offset * self._segments for offset in range(self._batch_size)]
def next_batch(self):
batch_data = np.zeros(self._batch_size, dtype=np.float32)
batch_labels = np.zeros(self._batch_size, dtype=np.float32)
for b in range(self._batch_size):
if self._cursor[b] + 1 >= self._prices_length:
# self._cursor[b] = b * self._segments
self._cursor[b] = np.random.randint(0, (b + 1) * self._segments)
batch_data[b] = self._prices[self._cursor[b]]
batch_labels[b] = self._prices[self._cursor[b] + np.random.randint(0, 5)]
self._cursor[b] = (self._cursor[b] + 1) % self._prices_length
return batch_data, batch_labels
def unroll_batches(self):
unroll_data, unroll_labels = [], []
init_data, init_label = None, None
for ui in range(self._num_unroll):
data, labels = self.next_batch()
unroll_data.append(data)
unroll_labels.append(labels)
return unroll_data, unroll_labels
def reset_indices(self):
for b in range(self._batch_size):
self._cursor[b] = np.random.randint(0, min((b + 1) * self._segments, self._prices_length - 1))
dg = DataGeneratorSeq(train_data, 5, 5)
u_data, u_labels = dg.unroll_batches()
for ui, (dat, lbl) in enumerate(zip(u_data, u_labels)):
print('\n\nUnrolled index %d' % ui)
dat_ind = dat
lbl_ind = lbl
print('\tInputs: ', dat)
print('\n\tOutput:', lbl)
# -
# ### Defining Hyperparameters
#
# Here we will define several hyperparamters which can be tweaked to optimize our model.
# +
D = 1 # Dimensionality of the data. Since your data is 1-D this would be 1
num_unrollings = 50 # Number of time steps you look into the future.
batch_size = 500 # Number of samples in a batch
num_nodes = [200, 200, 150] # Number of hidden nodes (neurons) in each layer of the deep LSTM stack we're using
n_layers = len(num_nodes) # number of layers
dropout = 0.2 # dropout amount
tf.compat.v1.reset_default_graph() # This is important in case you run this multiple times
# -
# ### Defining Inputs and Outputs
#
# Now we will define placeholders for training inputs and labels. We create a list of input placeholders, where each placeholder contains a single batch of data. This list has `num_unrollings` placeholders specified which will all be used at once for a single optimization step.
# +
# Input data.
train_inputs, train_outputs = [], []
# You unroll the input over time defining placeholders for each time step
tf.compat.v1.disable_eager_execution()
for ui in range(num_unrollings):
train_inputs.append(tf.compat.v1.placeholder(tf.float32, shape=[batch_size, D], name='train_inputs_%d' % ui))
train_outputs.append(tf.compat.v1.placeholder(tf.float32, shape=[batch_size, 1], name='train_outputs_%d' % ui))
# -
# ### Defining Parameters of the LSTM and Regression Layer
#
# Our model has three layers of LSTMs and a linear regression layer, denoted by `w` and `b`, which uses the output of the final LSTM cell to produce a prediction for the stock price at the next time step. Dropout-implemented LSTM cells are used to improve performance and reduce overfitting.
# +
lstm_cells = [
tf.compat.v1.nn.rnn_cell.LSTMCell(
num_units=num_nodes[li],
state_is_tuple=True,
initializer=tf.compat.v1.keras.initializers.VarianceScaling(
scale=1.0,
mode="fan_avg",
distribution="uniform"
)
)
for li in range(n_layers)
]
drop_lstm_cells = [
tf.compat.v1.nn.rnn_cell.DropoutWrapper(
lstm, input_keep_prob=1.0, output_keep_prob=1.0 - dropout, state_keep_prob=1.0 - dropout
)
for lstm in lstm_cells
]
drop_multi_cell = tf.keras.layers.StackedRNNCells(drop_lstm_cells)
multi_cell = tf.keras.layers.StackedRNNCells(lstm_cells)
w = tf.compat.v1.get_variable('w', shape=[num_nodes[-1], 1],
initializer=tf.compat.v1.keras.initializers.VarianceScaling(scale=1.0, mode="fan_avg",
distribution="uniform"))
b = tf.compat.v1.get_variable('b', initializer=tf.random.uniform([1], -0.1, 0.1))
# -
# ### Calculating LSTM Output and Feeding it to the Regression Layer to Get Final Prediction
#
# Now, we create TensorFlow variables `c` and `h` which hold the cell state and hidden state of the LSTM cell. We then transform the training inputs and feed them to the `dynamic_rnn(...)` function which calculates LSTM outputs, and split that output back into `num_unrollings` tensors.
# +
# Create cell state and hidden state variables to maintain the state of the LSTM
c, h = [],[]
initial_state = []
for li in range(n_layers):
c.append(tf.Variable(tf.zeros([batch_size, num_nodes[li]]), trainable=False))
h.append(tf.Variable(tf.zeros([batch_size, num_nodes[li]]), trainable=False))
initial_state.append(tf.compat.v1.nn.rnn_cell.LSTMStateTuple(c[li], h[li]))
# Do several tensor transofmations, because the function dynamic_rnn requires the output to be of
# a specific format. Read more at: https://www.tensorflow.org/api_docs/python/tf/nn/dynamic_rnn
all_inputs = tf.concat([tf.expand_dims(t,0) for t in train_inputs],axis=0)
# all_outputs is [seq_length, batch_size, num_nodes]
all_lstm_outputs, state = tf.compat.v1.nn.dynamic_rnn(
drop_multi_cell, all_inputs, initial_state=tuple(initial_state),
time_major = True, dtype=tf.float32)
all_lstm_outputs = tf.reshape(all_lstm_outputs, [batch_size*num_unrollings,num_nodes[-1]])
all_outputs = tf.compat.v1.nn.xw_plus_b(all_lstm_outputs,w,b)
split_outputs = tf.split(all_outputs,num_unrollings,axis=0)
# -
# ### Loss Calculation and Optimizer
#
# Now, we calculate the loss of our predictions by summing together the Mean Squared Error (MSE) of each prediction within each batch of data. Then, we define an optimizer which will be used to optimize the nueral network.
# +
# When calculating the loss you need to be careful about the exact form, because you calculate
# loss of all the unrolled steps at the same time
# Therefore, take the mean error or each batch and get the sum of that over all the unrolled steps
print('Defining training Loss')
loss = 0.0
with tf.control_dependencies([tf.compat.v1.assign(c[li], state[li][0]) for li in range(n_layers)]+
[tf.compat.v1.assign(h[li], state[li][1]) for li in range(n_layers)]):
for ui in range(num_unrollings):
loss += tf.reduce_mean(0.5*(split_outputs[ui]-train_outputs[ui])**2)
print('Learning rate decay operations')
global_step = tf.Variable(0, trainable=False)
inc_gstep = tf.compat.v1.assign(global_step,global_step + 1)
tf_learning_rate = tf.compat.v1.placeholder(shape=None,dtype=tf.float32)
tf_min_learning_rate = tf.compat.v1.placeholder(shape=None,dtype=tf.float32)
learning_rate = tf.maximum(
tf.compat.v1.train.exponential_decay(tf_learning_rate, global_step, decay_steps=1, decay_rate=0.5, staircase=True),
tf_min_learning_rate)
# Optimizer.
print('TF Optimization operations')
optimizer = tf.compat.v1.train.AdamOptimizer(learning_rate)
gradients, v = zip(*optimizer.compute_gradients(loss))
gradients, _ = tf.clip_by_global_norm(gradients, 5.0)
optimizer = optimizer.apply_gradients(
zip(gradients, v))
print('\tAll done')
# -
# ### Prediction Related Calculations
#
# The last step before running the LSTM is to define prediction-related TensorFlow operations. We first define a placeholder for feeding in our inputs (`sample_inputs`), and define state variables for prediction (`sample_c` and `sample_h`). Finally, we calculate predictions with the `dynamic_rnn(...)` function and send the output through the regression layer (`w` and `b`). We also define the `reset_sample_state` operation which resets the cell state and hidden state of the LSTM cells, and should be executed just before making each batch of predictions.
# +
print('Defining prediction related TF functions')
sample_inputs = tf.compat.v1.placeholder(tf.float32, shape=[1,D])
# Maintaining LSTM state for prediction stage
sample_c, sample_h, initial_sample_state = [],[],[]
for li in range(n_layers):
sample_c.append(tf.Variable(tf.zeros([1, num_nodes[li]]), trainable=False))
sample_h.append(tf.Variable(tf.zeros([1, num_nodes[li]]), trainable=False))
initial_sample_state.append(tf.compat.v1.nn.rnn_cell.LSTMStateTuple(sample_c[li],sample_h[li]))
reset_sample_states = tf.group(*[tf.compat.v1.assign(sample_c[li],tf.zeros([1, num_nodes[li]])) for li in range(n_layers)],
*[tf.compat.v1.assign(sample_h[li],tf.zeros([1, num_nodes[li]])) for li in range(n_layers)])
sample_outputs, sample_state = tf.compat.v1.nn.dynamic_rnn(multi_cell, tf.expand_dims(sample_inputs,0),
initial_state=tuple(initial_sample_state),
time_major = True,
dtype=tf.float32)
with tf.control_dependencies([tf.compat.v1.assign(sample_c[li],sample_state[li][0]) for li in range(n_layers)]+
[tf.compat.v1.assign(sample_h[li],sample_state[li][1]) for li in range(n_layers)]):
sample_prediction = tf.compat.v1.nn.xw_plus_b(tf.reshape(sample_outputs,[1,-1]), w, b)
print('\tAll done')
# -
# ### Running the LSTM
#
# Here we train and predict stock price movements for several (30) epochs and see how the network performs over time. The following procedure is used:
# 1. Define a test set of starting points (`test_points_seq`) on the time series to evaluate the model on
# 2. For each epoch
# 1. For full sequence length of training data
# 1. Unroll a set of `num_unrollings` batches
# 2. Train the neural network with the unrolled batches
# 2. Calculate the average training loss
# 3. For each starting point in the test set
# 1. Update the LSTM state by iterating through the previous `num_unrollings` data points found before the test point
# 2. Make predictions for `n_predict_once` steps continuously, using the previous prediction as the current input
# 3. Calculate the MSE loss between the `n_predict_once` points predicted and the true stock prices at those time stamps
#
# +
epochs = 30
valid_summary = 1 # Interval you make test predictions
n_predict_once = 50 # Number of steps you continously predict for
train_seq_length = train_data.size # Full length of the training data
train_mse_ot = [] # Accumulate Train losses
test_mse_ot = [] # Accumulate Test loss
predictions_over_time = [] # Accumulate predictions
session = tf.compat.v1.InteractiveSession()
tf.compat.v1.global_variables_initializer().run()
# Used for decaying learning rate
loss_nondecrease_count = 0
loss_nondecrease_threshold = 2 # If the test error hasn't increased in this many steps, decrease learning rate
print('Initialized')
average_loss = 0
# Define data generator
data_gen = DataGeneratorSeq(train_data,batch_size,num_unrollings)
x_axis_seq = []
# Points you start your test predictions from
test_points_seq = np.arange(11000,12000,50).tolist()
for ep in range(epochs):
# ========================= Training =====================================
for step in range(train_seq_length//batch_size):
u_data, u_labels = data_gen.unroll_batches()
feed_dict = {}
for ui,(dat,lbl) in enumerate(zip(u_data,u_labels)):
feed_dict[train_inputs[ui]] = dat.reshape(-1,1)
feed_dict[train_outputs[ui]] = lbl.reshape(-1,1)
feed_dict.update({tf_learning_rate: 0.0001, tf_min_learning_rate:0.000001})
_, l = session.run([optimizer, loss], feed_dict=feed_dict)
average_loss += l
# ============================ Validation ==============================
if (ep+1) % valid_summary == 0:
average_loss = average_loss/(valid_summary*(train_seq_length//batch_size))
# The average loss
if (ep+1)%valid_summary==0:
print('Average loss at step %d: %f' % (ep+1, average_loss))
train_mse_ot.append(average_loss)
average_loss = 0 # reset loss
predictions_seq = []
mse_test_loss_seq = []
# ===================== Updating State and Making Predicitons ========================
for w_i in test_points_seq:
mse_test_loss = 0.0
our_predictions = []
if (ep+1)-valid_summary==0:
# Only calculate x_axis values in the first validation epoch
x_axis=[]
# Feed in the recent past behavior of stock prices
# to make predictions from that point onwards
for tr_i in range(w_i-num_unrollings+1,w_i-1):
current_price = all_mid_data[tr_i]
feed_dict[sample_inputs] = np.array(current_price).reshape(1,1)
_ = session.run(sample_prediction,feed_dict=feed_dict)
feed_dict = {}
current_price = all_mid_data[w_i-1]
feed_dict[sample_inputs] = np.array(current_price).reshape(1,1)
# Make predictions for this many steps
# Each prediction uses previous prediciton as it's current input
for pred_i in range(n_predict_once):
pred = session.run(sample_prediction,feed_dict=feed_dict)
our_predictions.append(np.asscalar(pred))
feed_dict[sample_inputs] = np.asarray(pred).reshape(-1,1)
if (ep+1)-valid_summary==0:
# Only calculate x_axis values in the first validation epoch
x_axis.append(w_i+pred_i)
mse_test_loss += 0.5*(pred-all_mid_data[w_i+pred_i])**2
session.run(reset_sample_states)
predictions_seq.append(np.array(our_predictions))
mse_test_loss /= n_predict_once
mse_test_loss_seq.append(mse_test_loss)
if (ep+1)-valid_summary==0:
x_axis_seq.append(x_axis)
current_test_mse = np.mean(mse_test_loss_seq)
# Learning rate decay logic
if len(test_mse_ot)>0 and current_test_mse > min(test_mse_ot):
loss_nondecrease_count += 1
else:
loss_nondecrease_count = 0
if loss_nondecrease_count > loss_nondecrease_threshold :
session.run(inc_gstep)
loss_nondecrease_count = 0
print('\tDecreasing learning rate by 0.5')
test_mse_ot.append(current_test_mse)
print('\tTest MSE: %.5f'%np.mean(mse_test_loss_seq))
predictions_over_time.append(predictions_seq)
print('\tFinished Predictions')
# -
# ### Visualizing the Predictions
#
# Generally speaking, the MSE of our predictions goes down as the model continues to be trained. We can compare the best MSE of our model above back to that of our standard moving average model, at which point we see that our LSTM neural network performed significantly better. Make sure to replace the value of `best_prediction_epoch` with the step number of the epoch in which the test MSE is lowest.
# +
best_prediction_epoch = 18 # replace this with the epoch that you got the best results when running the plotting code
plt.figure(figsize = (18,18))
plt.subplot(2,1,1)
plt.plot(range(df.shape[0]),all_mid_data,color='b')
# Plotting how the predictions change over time
# Plot older predictions with low alpha and newer predictions with high alpha
start_alpha = 0.25
alpha = np.arange(start_alpha,1.1,(1.0-start_alpha)/len(predictions_over_time[::3]))
for p_i,p in enumerate(predictions_over_time[::3]):
for xval,yval in zip(x_axis_seq,p):
plt.plot(xval,yval,color='r',alpha=alpha[p_i])
plt.title('Evolution of Test Predictions Over Time',fontsize=18)
plt.xlabel('Date',fontsize=18)
plt.ylabel('Mid Price',fontsize=18)
plt.xlim(11000,12075)
plt.subplot(2,1,2)
# Predicting the best test prediction you got
plt.plot(range(df.shape[0]),all_mid_data,color='b')
for xval,yval in zip(x_axis_seq,predictions_over_time[best_prediction_epoch - 1]):
plt.plot(xval,yval,color='r')
plt.title('Best Test Predictions Over Time',fontsize=18)
plt.xlabel('Date',fontsize=18)
plt.ylabel('Mid Price',fontsize=18)
plt.xlim(11000,12075)
plt.show()
# -
# Note that the model is making predictions that fall roughly in the range of 0 to 1.0 (i.e., not the actual stock prices). This is because we're trying to predict the movement of the stock prices rather than the prices themselves.
| notebooks/lstm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.9 64-bit
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import mpl_toolkits.mplot3d.axes3d as p3
import numpy as np
from matplotlib import animation
from Quadrotor import Quadrotor_linear
from MPC_controller import mpc_control,mpc_control_stable,OTS,get_observer_gain,luenberger_observer
from visualization import data_for_cylinder_along_z
from convexification import get_intermediate_goal, convexify
from test import get_terminal_set_corners
np.random.seed(seed=0)
drone = [0,0,0.05] #pos_x,pos_y,radius
obs1=np.array([-3,1,1]) #pos_x,pos_y,radius
obs2=np.array([-2,-3,1]) #pos_x,pos_y,radius
obs3=np.array([0,2,1]) #pos_x,pos_y,radius
obs4=np.array([-5,-1.9,1]) #pos_x,pos_y,radius
obs5=np.array([0.5,-2,1]) #pos_x,pos_y,radius
obstacle_list=[obs1,obs2,obs3,obs4,obs5]#,obs6]#,obs1*2,obs2*2,obs3*2,obs4*2,obs5*2]
goal = np.array([-5,-5,2]) #pos_x,pos_y,pos_z
sensor_noise_sigma=np.array([0.05,0.05,0.05,0.001,0.001,0.001,0.001,0.001,0.001,0.001])
real_disturbance=np.random.normal(loc=0,scale=0.01,size=3)
print("real _dist", real_disturbance)
Cd= np.zeros((10,3))
Bd= np.array([[0, 0, 0],
[0, 0, 0],
[0, 0, 0],
[1, 0, 0],
[0, 1, 0],
[0, 0, 1],
[0, 0, 0],
[0, 0, 0],
[0, 0, 0],
[0, 0, 0]])
obs_eigen_values= np.array([-0.1, -0.1, -0.1, -0.03, -0.03, -0.03, 0.3, 0.3, 0.6, 0.6, -0.05, -0.05, -0.05])
def animate(i):
line.set_xdata(real_trajectory['x'][:i + 1])
line.set_ydata(real_trajectory['y'][:i + 1])
line.set_3d_properties(real_trajectory['z'][:i + 1])
line_est.set_xdata(real_trajectory['x'][:i + 1])
line_est.set_ydata(real_trajectory['y'][:i + 1])
line_est.set_3d_properties(real_trajectory['z'][:i + 1])
point.set_xdata(real_trajectory['x'][i])
point.set_ydata(real_trajectory['y'][i])
point.set_3d_properties(real_trajectory['z'][i])
if __name__ == "__main__":
N = 10
quadrotor_linear = Quadrotor_linear()
# Some initialisation
x_init = np.zeros(10)
x_init[0]=drone[0]
x_init[1]=drone[1]
x_target = np.zeros(10)
x_target[0] = goal[0]
x_target[1] = goal[1]
x_target[2] = goal[2]
x_hat = x_init
x_real = x_init
d_hat=np.zeros((3,1))
output = x_init
L=get_observer_gain(quadrotor_linear, Bd,Cd,obs_eigen_values)
# Get convex workspace with constraints for obstacles
A,b = convexify(x_init[:2].flatten(),drone[2],obstacle_list)
#Get intermediate goal in this convex space
inter_goal=get_intermediate_goal(x_init[:2].flatten(), 0, x_target[:2].flatten(), A,b).flatten()
x_intergoal=np.zeros(10)
x_intergoal[:2]=inter_goal
x_intergoal[2] = x_target[2]
real_trajectory = {'x': [], 'y': [], 'z': []}
est_trajectory = {'x': [], 'y': [], 'z': []}
output_trajectory = {'x': [], 'y': [], 'z': []}
d_hat_list=[]
# Optimal target selector
x_ref,u_ref = OTS(quadrotor_linear,x_intergoal,d_hat, A, b, Bd, Cd)
i = 0
while np.linalg.norm(x_intergoal[:3].flatten()-x_target[:3]) > 0.1 and i<500: #until intergoal and final goal are equal
i += 1 #limit number of iterations
# Get obstacle matrix
A_obs,b_obs=convexify(x_hat[:2].flatten(),drone[2],obstacle_list)
output = quadrotor_linear.disturbed_output(x_real,real_disturbance, Cd, sensor_noise_sigma)
#Compute the input with mpc
u = mpc_control(quadrotor_linear, N, x_hat.flatten(), x_ref.flatten(),u_ref.flatten(),A_obs,b_obs,1)
if u is None:
# if failure from MPC, u=0, which means hover input
print("no solution")
u=np.zeros((4,1))
else:
u = u.reshape(-1,1)
est_trajectory['x'].append(x_hat[0])
est_trajectory['y'].append(x_hat[1])
est_trajectory['z'].append(x_hat[2])
real_trajectory['x'].append(x_real[0])
real_trajectory['y'].append(x_real[1])
real_trajectory['z'].append(x_real[2])
output_trajectory['x'].append(output[0])
output_trajectory['y'].append(output[1])
output_trajectory['z'].append(output[2])
# Plant
x_real = quadrotor_linear.disturbed_next_x(x_real,u,real_disturbance,Bd)
#Observer
x_hat, d_hat = luenberger_observer(quadrotor_linear, x_hat, d_hat, output, u, Bd, Cd, L)
d_hat_list.append(d_hat)
#Recompute obstacles and intermediate goal
A_obs,b_obs = convexify(x_hat[:2].flatten(),drone[2],obstacle_list)
x_intergoal[:2] = get_intermediate_goal(output[:2].flatten(), 0,x_target[:2].flatten(), A_obs,b_obs).flatten()
x_ref,u_ref = OTS(quadrotor_linear, x_intergoal, d_hat, A_obs, b_obs, Bd, Cd)
if x_ref is None : #if failure from target selector
x_ref = x_intergoal
u_ref = np.zeros((4,1))
print(i)
print("x_real :",x_real)
#intermediate goal = final goal, so we pass on stable MPC with time invariant constraints
A,b = convexify(x_hat[:2].flatten(),drone[2],obstacle_list)
print("***")
while np.linalg.norm(x_real[:3].flatten() - x_target[:3]) >= 0.3 and i<=1000: #until reach final goal
i+=1
output = quadrotor_linear.disturbed_output(x_real,real_disturbance, Cd, sensor_noise_sigma)
# Use the mpc with terminal cost and set
u = mpc_control_stable(quadrotor_linear, 3*N, x_hat.flatten(), x_ref.flatten(),u_ref.flatten(),A,b,0.01,1)
if u is None:
print("no solution")
u=np.zeros((4,1))
else:
u = u.reshape(-1,1)
est_trajectory['x'].append(x_hat[0])
est_trajectory['y'].append(x_hat[1])
est_trajectory['z'].append(x_hat[2])
real_trajectory['x'].append(x_real[0])
real_trajectory['y'].append(x_real[1])
real_trajectory['z'].append(x_real[2])
output_trajectory['x'].append(output[0])
output_trajectory['y'].append(output[1])
output_trajectory['z'].append(output[2])
#Plant
x_real = quadrotor_linear.disturbed_next_x(x_real,u,real_disturbance,Bd)
#Observer
x_hat, d_hat = luenberger_observer(quadrotor_linear, x_hat, d_hat, output, u, Bd, Cd, L)
d_hat_list.append(d_hat)
#Optimal target selector
x_ref,u_ref = OTS(quadrotor_linear,x_intergoal,d_hat, A,b,Bd,Cd)
print(i)
print("x_real :",x_real)
if x_ref is None :#if failure from target selector
x_ref = x_intergoal
u_ref = np.zeros((4,1))
# +
""" Visualisation """
fig = plt.figure(1)
ax1 = p3.Axes3D(fig) # 3D place for drawing
real_trajectory['x'] = np.array(real_trajectory['x'], dtype=float)
real_trajectory['y'] = np.array(real_trajectory['y'], dtype=float)
real_trajectory['z'] = np.array(real_trajectory['z'], dtype=float)
est_trajectory['x'] = np.array(est_trajectory['x'], dtype=float)
est_trajectory['y'] = np.array(est_trajectory['y'], dtype=float)
est_trajectory['z'] = np.array(est_trajectory['z'], dtype=float)
output_trajectory['x'] = np.array(output_trajectory['x'], dtype=float)
output_trajectory['y'] = np.array(output_trajectory['y'], dtype=float)
output_trajectory['z'] = np.array(output_trajectory['z'], dtype=float)
point, = ax1.plot([real_trajectory['x'][0]], [real_trajectory['y'][0]], [real_trajectory['z'][0]], 'ro', ms=2.5, label='Quadrotor')
line, = ax1.plot([real_trajectory['x'][0]], [real_trajectory['y'][0]], [real_trajectory['z'][0]], label='Real_Trajectory')
line_est, = ax1.plot([est_trajectory['x'][0]], [est_trajectory['y'][0]], [est_trajectory['z'][0]], label='est_Trajectory')
ax1.set_xlabel('x')
ax1.set_ylabel('y')
ax1.set_zlabel('z')
ax1.set_xlim3d((-7, 3))
ax1.set_ylim3d((-6, 4))
ax1.set_zlim3d((0, 3))
ax1.set_title('3D animate')
ax1.view_init(30, 35)
ax1.legend(loc='lower right')
for obstacle in obstacle_list:
Xc,Yc,Zc = data_for_cylinder_along_z(obstacle[0],obstacle[1],obstacle[2],2)
ax1.plot_surface(Xc, Yc, Zc, alpha=0.5)
points = get_terminal_set_corners(quadrotor_linear, x_target[:3], 0.01)
for point in points:
ax1.scatter3D(point[0], point[1], point[2], s=5)
ani = animation.FuncAnimation(fig=fig,
func=animate,
frames=len(real_trajectory['x']),
interval=5,
repeat=False,
blit=False)
plt.show()
| test/terminal_set.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W3D3_OptimalControl/W3D3_Tutorial1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# -
# # Tutorial 1: Optimal Control for Discrete States
# **Week 3, Day 3: Optimal Control**
#
# **By Neuromatch Academy**
#
# __Content creators:__ <NAME>, <NAME>, <NAME>
#
# __Content reviewers:__ <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>
# **Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
#
# <p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
# ---
# # Tutorial Objectives
#
# In this tutorial, we will implement a binary control task: a Partially Observable Markov Decision Process (POMDP) that describes fishing. The agent (you) seeks reward from two fishing sites without directly observing where the school of fish is (a group of fish is called a school!). This makes the world a Hidden Markov Model. Based on when and where you catch fish, you keep updating your belief about the fish location, _i.e._ the posterior of the fish given past observations. You should control your position to get the most fish while minimizing the cost of switching sides.
#
# You've already learned about stochastic dynamics, latent states, and measurements. Now we introduce you to the new concepts of **control, utility, and policy**.
# + cellView="form"
# @title Tutorial slides
# @markdown These are the slides for all videos in this tutorial.
from IPython.display import IFrame
IFrame(src=f"https://mfr.ca-1.osf.io/render?url=https://osf.io/8j5rs/?direct%26mode=render%26action=download%26mode=render", width=854, height=480)
# -
# ---
# ## Setup
#
#
# +
# Imports
import numpy as np
import scipy
from scipy.linalg import inv
from math import isclose
import matplotlib.pyplot as plt
# + cellView="form"
#@title Figure Settings
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import ipywidgets as widgets
from ipywidgets import interact, fixed, HBox, Layout, VBox, interactive, Label
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
# + cellView="form"
# @title Plotting Functions
def plot_fish(fish_state, ax=None):
"""
Plot the fish dynamics
"""
T = len(fish_state)
showlen = min(T, 200)
startT = 0
endT = startT + showlen
showT = range(startT, endT)
time_range = np.linspace(0, showlen - 1)
if not ax:
fig, ax = plt.subplots(1, 1, figsize=(12, 2.5))
ax.plot(- fish_state[showT], color='dodgerblue', markersize=10, linewidth=3.0)
ax.set_xlabel('time', fontsize=18)
ax.set_ylabel('Fish state', rotation=360, fontsize=18)
ax.yaxis.set_label_coords(-0.1, 0.25)
ax.set_xticks([0, showlen, showlen])
ax.tick_params(axis='both', which='major', labelsize=18)
ax.set_xlim([0, showlen])
ax.set_ylim([-1.1, 1.1])
ax.set_yticks([-1, 1])
ax.tick_params(axis='both', which='major', labelsize=18)
labels = [item.get_text() for item in ax.get_yticklabels()]
labels[0] = 'Right'
labels[1] = 'Left'
ax.set_yticklabels(labels)
def plot_measurement(measurement, ax=None):
"""
Plot the measurements
"""
T = len(measurement)
showlen = min(T, 200)
startT = 0
endT = startT + showlen
showT = range(startT, endT)
time_range = np.linspace(0, showlen - 1)
if not ax:
fig, ax = plt.subplots(1, 1, figsize=(12, 2.5))
ax.plot(measurement[showT], 'r*', markersize=5)
ax.set_xlabel('time', fontsize=18)
ax.set_ylabel('Measurement', rotation=360, fontsize=18)
ax.yaxis.set_label_coords(-0.2, 0.4)
ax.set_xticks([0, showlen, showlen])
ax.tick_params(axis='both', which='major', labelsize=18)
ax.set_xlim([0, showlen])
ax.set_ylim([-.1, 1.1])
ax.set_yticks([0, 1])
ax.set_yticklabels(['no fish', 'caught fish'])
def plot_act_loc(loc, act, ax_loc=None):
"""
Plot the action and location of 200 time points
"""
T = len(act)
showlen = min(T, 200)
startT = 0
endT = startT + showlen
showT = range(startT, endT)
time_range = np.linspace(0, showlen - 1)
if not ax_loc:
fig, ax_loc = plt.subplots(1, 1, figsize=(12, 2.5))
act_int = (act == "switch").astype(int)
ax_loc.plot(-loc[showT], 'g.-', markersize=8, linewidth=5)
ax_loc.plot((act_int[showT] * 4 - 3) * .5, 'rv', markersize=12,
label='switch')
ax_loc.set_xlabel('time', fontsize=18)
ax_loc.set_ylabel('Your state', rotation=360, fontsize=18)
ax_loc.legend(loc="upper right", fontsize=12)
ax_loc.set_xlim([0, showlen])
ax_loc.set_ylim([-1.1, 1.1])
ax_loc.set_yticks([-1, 1])
ax_loc.set_xticks([0, showlen, showlen])
ax_loc.tick_params(axis='both', which='major', labelsize=18)
labels = [item.get_text() for item in ax_loc.get_yticklabels()]
labels[1] = 'Left'
labels[0] = 'Right'
ax_loc.set_yticklabels(labels)
def plot_belief(belief, ax1=None, choose_policy=None):
"""
Plot the belief dynamics of 200 time points
"""
T = belief.shape[1]
showlen = min(T, 200)
startT = 0
endT = startT + showlen
showT = range(startT, endT)
time_range = np.linspace(0, showlen - 1)
if not ax1:
fig, ax1 = plt.subplots(1, 1, figsize=(12, 2.5))
ax1.plot(belief[0, showT], color='dodgerblue', markersize=10, linewidth=3.0)
ax1.yaxis.set_label_coords(-0.1, 0.25)
ax1.set_xlabel('time', rotation=360, fontsize=18)
ax1.set_ylabel('Belief on \n left', rotation=360, fontsize=18)
ax1.tick_params(axis='both', which='major', labelsize=18)
ax1.set_xlim([0, showlen])
ax1.set_yticks([0, 1])
ax1.set_ylim([0, 1.1])
ax1.set_xticks([0, showlen, showlen])
if choose_policy == "threshold":
ax2 = ax1.twinx()
ax2.plot(time_range, threshold * np.ones(time_range.shape), 'r--')
ax2.plot(time_range, (1 - threshold) * np.ones(time_range.shape), 'c--')
ax2.set_yticks([threshold, 1 - threshold])
ax2.set_ylim([0, 1.1])
ax2.tick_params(axis='both', which='major', labelsize=18)
labels = [item.get_text() for item in ax2.get_yticklabels()]
labels[0] = 'threshold to switch \n from left to right'
labels[-1] = 'threshold to switch \n from right to left'
ax2.set_yticklabels(labels)
def plot_dynamics(belief, loc, act, meas, fish_state, choose_policy):
"""
Plot the dynamics of 200 time points
"""
if choose_policy == 'threshold':
fig, [ax0, ax_loc, ax1, ax_bel] = plt.subplots(4, 1, figsize=(12, 9))
plot_fish(fish_state, ax=ax0)
plot_belief(belief, ax1=ax_bel)
plot_measurement(meas, ax=ax1)
plot_act_loc(loc, act, ax_loc=ax_loc)
else:
fig, [ax0, ax1, ax_bel] = plt.subplots(3, 1, figsize=(12, 7))
plot_fish(fish_state, ax=ax0)
plot_belief(belief, ax1=ax_bel)
plot_measurement(meas, ax=ax1)
plt.tight_layout()
plt.show()
def belief_histogram(belief, bins=100):
"""
Plot the histogram of belief states
"""
fig, ax = plt.subplots(1, 1, figsize=(8, 6))
ax.hist(belief, bins)
ax.set_xlabel('belief', fontsize=18)
ax.set_ylabel('count', fontsize=18)
plt.show()
def plot_value_threshold(cost_sw=0.5, T=10000, p_stay=.95,
high_rew_p=.4, low_rew_p=.1, step=.05):
"""
Helper function to plot the value function and threshold
"""
params = [T, p_stay, high_rew_p, low_rew_p, _]
threshold_array, value_array = value_threshold(params, cost_sw, step)
yrange = np.max(value_array) - np.min(value_array)
fig_, ax = plt.subplots(1, 1, figsize=(8, 6))
ax.plot(threshold_array, value_array, 'b')
ax.set_ylim([np.min(value_array) - yrange * .1, np.max(value_array) + yrange * .1])
ax.set_title(f'threshold vs value with switching cost c = {cost_sw:.2f}',
fontsize=20)
ax.set_xlabel('threshold', fontsize=16)
ax.set_ylabel('value', fontsize=16)
plt.show()
# + cellView="form"
# @title Helper Functions
binomial = np.random.binomial
class ExcerciseError(AssertionError):
pass
def test_policy_threshold():
well_done = True
for loc in [-1, 1]:
threshold = 0.4
belief = np.array([.2, .3])
if policy_threshold(threshold, belief, loc) != "switch":
raise ExcerciseError("'policy_threshold' function is not correctly implemented!")
for loc in [1, -1]:
threshold = 0.6
belief = np.array([.7, .8])
if policy_threshold(threshold, belief, loc) != "stay":
raise ExcerciseError("'policy_threshold' function is not correctly implemented!")
print("Well Done!")
def test_value_function():
measurement = np.array([0, 0, 0, 1, 0, 0, 0, 0, 1, 1])
act = np.array(["switch", "stay", "switch", "stay", "stay",
"stay", "switch", "switch", "stay", "stay"])
cost_sw = .5
if not isclose(value_function(measurement, act, cost_sw), .1):
raise ExcerciseError("'value_function' function is not correctly implemented!")
print("Well Done!")
# -
# ---
# # Section 1: Dynamics of Fishing
# + cellView="form"
# @title Video 1: Gone fishing
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV1FL411p7o5", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="3oIwUFpolVA", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# There are two locations for the fish and you (Left and Right). If you're on the same side as the fish, you'll catch more, with probabilty $q_{\rm high}$ per discrete time step. Otherwise you may still catch fish with probability $q_{\rm low}$. One fish is worth 1 "point".
#
# The fish location $s^{\rm fish}$ is latent. The only information you get about the fish location is when you catch one. Secretly at each time step, the fish may switch sides with a certain probability $p_{\rm sw} = 1 - p_{\rm stay}$.
#
#
# You are in control of your own location. You may stay on your current side with no cost, or switch to the other side and incur an action cost $C$ (again, in units of fish).
#
# You select controls or actions by following a **policy**. This defines what to do in any situation. Here the situation is specified by your location and your belief $b_t$ about the fish location. For optimal control we assume that this belief is the posterior probability over the current fish location, given all the past measurements. We only need one number for this, since the fish are either on the left or the right. So we write
#
# $$b_t = p(s^{\rm fish}_t = {\rm Right}\ |\ m_{0:t}, a_{0:t-1})$$
#
# where $m$ are the measurements, and $a$ are the controls or actions (stay or switch).
#
# Ultimately we will parameterize the policy by a simple threshold on beliefs. (This happens to be optimal if you pick the right threshold!) When your belief that fish are on your current side falls below a threshold $\theta$, you switch to the other side.
#
# Your **overall goals** in this tutorial are:
# 1. Measure when fish are caught, first if the school of fish doesn't move.
# 2. For moving fish, plot their dynamics and your belief about it based on your measurements.
# 3. Compute the value for a given control policy.
# 4. Find the optimal policy for controlling your position.
# ## Interactive Demo 1: Examining fish dynamics
#
# In this demo, we will look at the dynamics of the fish moving from side to side. We will use the helper class `binaryHMM`, implemented in the next cell. Please take some time to study this class and its methods. You will then see a demo where you can change the probability of switching states and examing the resulting dynamics of the fish.
#
# + cellView="both"
class binaryHMM():
def __init__(self, params, fish_initial=-1, loc_initial=-1):
self.params = params
self.fish_initial = fish_initial
self.loc_initial = loc_initial
def fish_state_telegraph(self, fish_past, p_stay):
"""
fish state update according to telegraph process
Args:
fish_past (int): the fish location (-1 for left side, 1 for right side)
p_stay : the probability that the state of a certain site stays the same
Returns:
fish_new (int): updated fish location
"""
# we use logical operation XOR (denoted by ^ in python)
fish_new = (1 - binomial(1, p_stay)) ^ ((fish_past + 1) // 2)
fish_new = fish_new * 2 - 1
return fish_new
def fish_dynamics(self):
"""
fish state dynamics according to telegraph process
Returns:
fish_state (numpy array of int)
"""
T, p_stay, _, _, _ = self.params
fish_state = np.zeros(T, int) # -1: left side ; 1: right side
# initialization
fish_state[0] = self.fish_initial
for t in range(1, T):
fish_state[t] = self.fish_state_telegraph(fish_state[t - 1], p_stay)
return fish_state
def generate_process_lazy(self):
"""
fish dynamics and measurements if you always stay in the intial location
without changing sides
Returns:
fish_state (numpy array of int): locations of the fish
loc (numpy array of int): left or right site, -1 for left, and 1 for right
measurement (numpy array of binary): whether a reward is obtained
"""
T, _, high_rew_p, low_rew_p, _ = self.params
rew_p_vector = np.array([low_rew_p, high_rew_p])
fish_state = self.fish_dynamics()
loc = np.zeros(T, int) # -1: left side, 1: right side
measurement = np.zeros(T, int) # 0: no food, 1: get food
for t in range(0, T):
loc[t] = self.loc_initial
# new measurement
measurement[t] = binomial(1, rew_p_vector[(fish_state[t] == loc[t]) * 1])
return fish_state, loc, measurement
# -
def update_ex_1(p_stay=.95, high_rew_p=.4, low_rew_p=.1, T=200):
"""
p_stay: probability fish stay
high_rew_p: p(catch fish) when you're on their side
low_rew_p : p(catch fish) when you're on other side
"""
params = [T, p_stay, high_rew_p, low_rew_p, _]
#### initial condition for fish [fish_initial] and you [loc_initial] ####
binaryHMM_test = binaryHMM(params, fish_initial=-1, loc_initial=-1)
fish_state = binaryHMM_test.fish_dynamics()
plot_fish(fish_state)
# + cellView="form"
# @markdown Make sure you execute this cell to enable the widget!
widget=interactive(update_ex_1, {'manual': True},
high_rew_p=fixed(.4),
low_rew_p=fixed(.1),
p_stay=(.5, 1., .001),
T=fixed(200))
widget.children[-2].description='Run Simulation'
widget.children[-2].style.button_color='lightgreen'
controls = HBox(widget.children[:-1], layout=Layout(flex_flow='row wrap'))
output = widget.children[-1]
display(VBox([controls, output]))
# -
# ---
# # Section 2: Catching fish
# + cellView="form"
# @title Video 2: Catch some fish
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV1kD4y1m7Lo", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="1-Wionllt9U", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# ## Interactive Demo 2: Catching fish
#
# Now set $p_{\rm stay} = 1$ so that the state of the two sites are fixed, and we can directly see the chances of catching fish on each side. The variable `fish_initial` indicates the initial side of the fish, and `loc_initial` indicates your initial location. They each take value $-1$ for left and $1$ for right.
#
# **Instructions:**
# 1. set the two locations (`fish_initial` and `loc_initial`) to be the _same_, and measure when you catch fish.
# 2. set the two locations (`fish_initial` and `loc_initial`) to be the _different_, and measure when you catch fish.
# 3. visually compare the measurements from 1 and 2.
# 4. Finally, you can also play around with `high_rew_p` (high reward probability) and `low_rew_p` (low reward probability) sliders.
def update_ex_2(p_stay=1., high_rew_p=.6, low_rew_p=.05, T=100):
"""
p_stay: probability fish stay
high_rew_p: p(catch fish) when you're on their side
low_rew_p : p(catch fish) when you're on other side
"""
params = [T, p_stay, high_rew_p, low_rew_p, _]
#### initial condition for fish [fish_initial] and you [loc_initial] ####
binaryHMM_test = binaryHMM(params, fish_initial=-1, loc_initial=-1)
fish_state, loc, measurement = binaryHMM_test.generate_process_lazy()
plot_measurement(measurement)
# + cellView="form"
#@markdown Make sure you execute this cell to enable the widget!
widget=interactive(update_ex_2, {'manual': True},
high_rew_p=(.0, 1., .001),
low_rew_p=(.0, 1., .001),
p_stay=fixed(1.),
T=fixed(100))
widget.children[-2].description='Run Simulation'
widget.children[-2].style.button_color='lightgreen'
controls = HBox(widget.children[:-1], layout=Layout(flex_flow='row wrap'))
output = widget.children[-1]
display(VBox([controls, output]))
# -
# ---
# # Section 3: Belief dynamics and belief distributions
# + cellView="form"
# @title Video 3: Where are the fish?
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV19t4y1Q7VH", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="wCzVnnd4bmg", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# We have provided a class for the binary task, with the agent always staying at one side following a lazy policy function `def policy_lazy(belief, loc)` that we provided. Now in this exercise, you will extend the module to generate the real dynamics, including beliefs and a moving agent. With the generated data, we will see how beliefs change over time, and how often different beliefs happen.
#
# For convenience, your belief at time *t* is actually a 2-dimensional vector. The first element is the belief that the fish are on the left, and the second element is the belief the fish are on the right. At every time, these elements sum to $1$.
#
# We will first check the dynamics with lazy policy, and then explore the case with a threshold-based policy.
#
#
# Evaluate the cells below to setup the lazy policy.
#
# + cellView="both"
class binaryHMM_belief(binaryHMM):
def __init__(self, params, fish_initial = -1, loc_initial = -1,
choose_policy = 'threshold'):
binaryHMM.__init__(self, params, fish_initial, loc_initial)
self.choose_policy = choose_policy
def generate_process(self):
"""
fish dynamics and measurements based on the choosen policy
Returns:
belief (numpy array of float): belief on the states of the two sites
act (numpy array of string): actions over time
loc (numpy array of int): left or right site
measurement (numpy array of binary): whether a reward is obtained
fish_state (numpy array of int): fish locations
"""
T, p_stay, high_rew_p, low_rew_p, threshold = self.params
fish_state = self.fish_dynamics() # -1: left side; 1: right side
loc = np.zeros(T, int) # -1: left side, 1: right side
measurement = np.zeros(T, int) # 0: no food, 1: get food
act = np.empty(T, dtype='object') # "stay", or "switch"
belief = np.zeros((2, T), float) # the probability that the fish is on the left (1st element)
# or on the right (2nd element),
# the beliefs on the two boxes sum up to be 1
rew_prob = np.array([low_rew_p, high_rew_p])
# initialization
loc[0] = -1
measurement[0] = 0
belief_0 = np.random.random(1)[0]
belief[:, 0] = np.array([belief_0, 1 - belief_0])
act[0] = self.policy(threshold, belief[:, 0], loc[0])
for t in range(1, T):
if act[t - 1] == "stay":
loc[t] = loc[t - 1]
else:
loc[t] = - loc[t - 1]
# new measurement
measurement[t] = binomial(1, rew_prob[(fish_state[t] == loc[t]) * 1])
belief[0, t] = self.belief_update(belief[0, t - 1] , loc[t],
measurement[t], p_stay,
high_rew_p, low_rew_p)
belief[1, t] = 1 - belief[0, t]
act[t] = self.policy(threshold, belief[:, t], loc[t])
return belief, loc, act, measurement, fish_state
def policy(self, threshold, belief, loc):
"""
chooses policy based on whether it is lazy policy
or a threshold-based policy
Args:
threshold (float): the threshold of belief on the current site,
when the belief is lower than the threshold, switch side
belief (numpy array of float): the belief on the two sites
loc (int) : the location of the agent
Returns:
act (string): "stay" or "switch"
"""
if self.choose_policy == "threshold":
act = policy_threshold(threshold, belief, loc)
if self.choose_policy == "lazy":
act = policy_lazy(belief, loc)
return act
def belief_update(self, belief_past, loc, measurement, p_stay,
high_rew_p, low_rew_p):
"""
using PAST belief on the LEFT box, CURRENT location and
and measurement to update belief
"""
rew_prob_matrix = np.array([[1 - high_rew_p, high_rew_p],
[1 - low_rew_p, low_rew_p]])
# update belief posterior, p(s[t] | measurement(0-t), act(0-t-1))
belief_0 = (belief_past * p_stay + (1 - belief_past) * (1 - p_stay)) *\
rew_prob_matrix[(loc + 1) // 2, measurement]
belief_1 = ((1 - belief_past) * p_stay + belief_past * (1 - p_stay)) *\
rew_prob_matrix[1-(loc + 1) // 2, measurement]
belief_0 = belief_0 / (belief_0 + belief_1)
return belief_0
# -
def policy_lazy(belief, loc):
"""
This function is a lazy policy where stay is also taken
"""
act = "stay"
return act
# ## Interactive Demo 3: Task dynamics following a **lazy** policy
#
# The parameter for policy `choose_policy` can be either "*lazy*" or "*threshold*". In the following example, use the lazy policy.
#
# **Instructions:**
# * With the class defined above, we have created an object of `binaryHMM_belief` given parameters of the dynamics, *params*, and a parameter for policy.
# * Run the dynamics and explain the time series of the beliefs you see.
#
def update_ex_3(p_stay=.98, threshold=.2, high_rew_p=.4, low_rew_p=.1, T=200):
"""
p_stay: probability fish stay
high_rew_p: p(catch fish) when you're on their side
low_rew_p : p(catch fish) when you're on other side
threshold: threshold of belief below which switching is taken
"""
params = [T, p_stay, high_rew_p, low_rew_p, threshold]
#### initial condition for fish [fish_initial] and you [loc_initial] ####
binaryHMM_test = binaryHMM_belief(params, choose_policy="lazy",
fish_initial=-1, loc_initial=-1)
belief, loc, act, measurement, fish_state = binaryHMM_test.generate_process()
plot_dynamics(belief, loc, act, measurement, fish_state,
binaryHMM_test.choose_policy)
# + cellView="form"
#@markdown Make sure you execute this cell to enable the widget!
widget=interactive(update_ex_3, {'manual': True},
high_rew_p=(.0, 1., .001),
low_rew_p=(.0, 1., .001),
p_stay=(.5, 1., .001),
T=fixed(200),
threshold=fixed(.2))
widget.children[-2].description='Run Simulation'
widget.children[-2].style.button_color='lightgreen'
controls = HBox(widget.children[:-1], layout=Layout(flex_flow='row wrap'))
output = widget.children[-1]
display(VBox([controls, output]))
# -
# ---
# # Section 4: Implementing threshold policy
# + cellView="form"
# @title Video 4: How should you act?
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV1ri4y137cj", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="G3fNz23IDUg", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# ## Coding Exercise 4: dynamics following a **threshold-based** policy.
#
# **Instructions:**
#
# * You need to code a new policy `def policy_threshold(threshold, belief, loc)`. The policy takes three inputs: your belief about the fish state, your location ("Left" or "Right"), and a belief _threshold_: when your belief that you are on the same side as the fish drops below this threshold, you choose to switch; otherwise you stay.
#
# * You should return an action for each time *t*, which takes the value of "stay" or "switch".
#
# * After you complete the code for the policy based on threshold, create an object of `binaryHMM_belief` and set the policy parameter to be `choose_policy = threshold`.
# * We have provided an example of the parameters. You should play with the parameters to see the various dynamics.
# + cellView="both"
def policy_threshold(threshold, belief, loc):
"""
chooses whether to switch side based on whether the belief
on the current site drops below the threshold
Args:
threshold (float): the threshold of belief on the current site,
when the belief is lower than the threshold, switch side
belief (numpy array of float, 2-dimensional): the belief on the
two sites at a certain time
loc (int) : the location of the agent at a certain time
-1 for left side, 1 for right side
Returns:
act (string): "stay" or "switch"
"""
############################################################################
## Insert code to:
## generate actions (Stay or Switch) for current belief and location
##
## Belief is a 2d vector: first element = Prob(fish on Left | measurements)
## second element = Prob(fish on Right | measurements)
## Returns "switch" if Belief that fish are in your current location < threshold
## "stay" otherwise
##
## Hint: use loc value to determine which row of belief you need to use
## see the docstring for more information about loc
##
## complete the function and remove
raise NotImplementedError("Student exercise: Please complete <act>")
############################################################################
# Write the if statement
if ...:
# action below threshold
act = ...
else:
# action above threshold
act = ...
return act
# Test your function
test_policy_threshold()
# +
# to_remove solution
def policy_threshold(threshold, belief, loc):
"""
chooses whether to switch side based on whether the belief
on the current site drops below the threshold
Args:
threshold (float): the threshold of belief on the current site,
when the belief is lower than the threshold, switch side
belief (numpy array of float, 2-dimensional): the belief on the
two sites at a certain time
loc (int) : the location of the agent at a certain time
-1 for left side, 1 for right side
Returns:
act (string): "stay" or "switch"
"""
# Write the if statement
if belief[(loc + 1) // 2] <= threshold:
# action below threshold
act = "switch"
else:
# action above threshold
act = "stay"
return act
# Test your function
test_policy_threshold()
# -
# ## Interactive Demo 4: Dynamics with different thresholds
def update_ex_4(p_stay=.98, threshold=.2, high_rew_p=.4, low_rew_p=.1, T=200):
"""
p_stay: probability fish stay
high_rew_p: p(catch fish) when you're on their side
low_rew_p : p(catch fish) when you're on other side
threshold: threshold of belief below which switching is taken
"""
params = [T, p_stay, high_rew_p, low_rew_p, threshold]
#### initial condition for fish [fish_initial] and you [loc_initial] ####
binaryHMM_test = binaryHMM_belief(params, fish_initial=-1, loc_initial=-1,
choose_policy="threshold")
belief, loc, act, measurement, fish_state = binaryHMM_test.generate_process()
plot_dynamics(belief, loc, act, measurement,
fish_state, binaryHMM_test.choose_policy)
# + cellView="form"
#@markdown Make sure you execute this cell to enable the widget!
widget=interactive(update_ex_4, {'manual': True},
high_rew_p=fixed(.4),
low_rew_p=fixed(.1),
p_stay=fixed(.95),
T=fixed(200),
threshold=(.0, 1., .001))
widget.children[-2].description='Run Simulation'
widget.children[-2].style.button_color='lightgreen'
controls = HBox(widget.children[:-1], layout=Layout(flex_flow='row wrap'))
output = widget.children[-1]
display(VBox([controls, output]))
# -
# ---
# # Section 5: Implementing a value function
# + cellView="form"
# @title Video 5: Evaluate policy
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV1TD4y1D7K3", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="aJhffROC74w", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# ## Coding Exercise 5: Implementing a value function
#
# Now we have generated behavior for a policy parameterized by a threshold. While it seems clear that this is at least better than being lazy, we want to know how good it is. For that, we will calculate a _value function_. We will use this value to compare different policies, and maximize the amount of fish we catch while minimizing our effort.
#
# Specifically, here the value is total expected utility per unit time.
#
# $$V(\theta) = \frac{1}{T}\left(\sum_t U_s(s_t) + U_a(a_t)\right)$$
#
# where $U_s(s_t)$ is the instantaneous utility (reward) from the site, and $U_a(a_t)$ is the utility (negative cost) for the chosen action. Here, the action cost is 0 if you stay, and `cost_sw` if you switch.
#
# We could take this average mathematically over the probabilities of rewards and actions. More simply, we get the same answer by simply averaging the _actual_ rewards and costs over a long time, so that's what you should do.
#
#
# **Instructions**
# * Fill in the function `value_function(measurement, act, cost_sw)` given a sequence of measurements, actions, and the cost of switching.
# * Visually find the threshold that yields the highest total value. We have provided code for plotting value versus threshold. The threshold $\theta^*$ with the highest value gives the optimal policy for controlling where you should fish.
# + cellView="both"
def value_function(measurement, act, cost_sw):
"""
value function
Args:
act (numpy array of string): length T with each element
taking value "stay" or "switch"
cost_sw (float): the cost of switching side
measurement (numpy array of binary): whether a reward is obtained
Returns:
value (float): expected utility per unit time
"""
act_int = (act == "switch").astype(int)
T = len(measurement)
############################################################################
## Insert your code here to:
## compute the value function = rate of catching fish - costs
##
## complete the function and remove
raise NotImplementedError("Student exercise: Please complete <value>")
############################################################################
# Calculate the value function
value = ...
return value
# Test your function
test_value_function()
# +
# to_remove solution
def value_function(measurement, act, cost_sw):
"""
value function
Args:
act (numpy array of string): length T with each element
taking value "stay" or "switch"
cost_sw (float): the cost of switching side
measurement (numpy array of binary): whether a reward is obtained
Returns:
value (float): expected utility per unit time
"""
act_int = (act == "switch").astype(int)
T = len(measurement)
# Calculate the value function
value = np.sum(measurement - act_int * cost_sw) / T
return value
# Test your function
test_value_function()
# +
# Brute force search for optimal policy: loop over thresholds and compute value for each.
# This function is needed for the second exercise.
def value_threshold(params, cost_sw, step):
threshold_array = np.arange(0, .5 + step, step)
value_array = np.zeros(threshold_array.shape)
T, p_stay, high_rew_p, low_rew_p, _ = params
for i in range(len(threshold_array)):
threshold = threshold_array[i]
params = [T, p_stay, high_rew_p, low_rew_p, threshold]
binaryHMM_test = binaryHMM_belief(params, choose_policy="threshold")
belief, loc, act, measurement, fish_state = binaryHMM_test.generate_process()
value_array[i] = value_function(measurement, act, cost_sw)
return threshold_array, value_array
plot_value_threshold(cost_sw=0.5, p_stay=0.95, high_rew_p=0.4, low_rew_p=0.1, T=10000)
# -
# ---
# # Summary
#
# In this tutorial, you have used the Hidden Markov Models you learned about yesterday to figure out where the fish are, and then you have acted (moved location) to catch the most fish. In particular, you have found an optimal policy to guide your actions.
# ---
# # Bonus
# ---
# ## Bonus Section 1: Different task, different optimal policy
# + cellView="form"
# @title Video 6: Sensitivity of optimal policy
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV1QK4y1e7N9", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="wd8IVsKoEfA", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# ### Bonus Interactive Demo 1
#
# **Instructions:**
# After plotting value versus threshold, adjust various task parameters using the sliders below, and observe how the optimal threshold moves with
# * switching cost (`cost_sw`)
# * fish dynamics (`p_switch`)
# * probability of catching fish on each side, `low_rew_p` and `high_rew_p`
#
# Can you explain why the optimal threshold changes with these parameters?
#
# EXPLAIN why the optimal threshold changes for:
# * lower switching cost?
# * faster fish dynamics?
# * rarer fish?
#
# Note that it may require long simulations to see subtle changes in values of different policies, so look for coarse trends first.
#
#
#
# + cellView="form"
#@title
#@markdown Make sure you execute this cell to enable the widget!
widget=interactive(plot_value_threshold, {'manual': True},
T=fixed(10000),
p_stay=(0.5, 1., 0.001),
high_rew_p=(0., 1., 0.001),
low_rew_p=(0., 1., 0.001),
cost_sw=(0., 2., .1),
step=fixed(0.1))
widget.children[-2].description='Run Simulation'
widget.children[-2].style.button_color='lightgreen'
controls = HBox(widget.children[:-1], layout=Layout(flex_flow='row wrap'))
output = widget.children[-1]
display(VBox([controls, output]))
# +
# to_remove explanation
"""
* High switching cost means that you should be more certain that the other side
is better before committing to change sides. This means that beliefs must fall
below a threshold before acting. Conversely, a lower switching cost allows you
more flexibility to switch at less stringent thresholds. In the limit of _zero_
switching cost, you should always switch whenever you think the other side is
better, even if it's just 51%, and even if you switch every time step.
* Faster fish dynamics (lower `p_stay`) also promotes faster switching, because
you cannot plan as far into the future. In that case you must base your decisions
on more immediate evidence, but since you still pay the same switching cost that
cost is a higher fraction of your predictable rewards. And thus you should be more
conservative, and switch only when you are more confident.
* When `high_rew_p` and/or `low_rew_p` decreases, your predictions become less reliable,
again encouraging you to require more confidence before committing to a switch.
"""
# + cellView="form"
# @title Video 7: From discrete to continuous control
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV1JA411v7jy", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="ndCMgdjv9Gg", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
| tutorials/W3D3_OptimalControl/W3D3_Tutorial1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <NAME> 817518 Assignment 2
# ____
# ### Roadmap:
# - Data Loading
# - Data Preprocessing
# - Model Definition
# - Training
# - Regularization
# - Test with Feature Selection
# - Test Prediction
# +
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
import keras.backend as K
from keras.utils.generic_utils import get_custom_objects
from keras.layers import Activation, LeakyReLU
from tensorflow.keras.activations import sigmoid
from tensorflow.keras import regularizers
import numpy as np
import tensorflow as tf
from tensorflow import keras
from keras.layers import Dense
from keras.models import Sequential
from tensorflow.keras import layers
# -
# ### Data Loading
x_train = pd.read_csv("X_train.csv", index_col=False)
x_train_originale=x_train
x_train
y_train = pd.read_csv("Y_train.csv", index_col=False)
y_train
x_train= pd.merge(x_train, y_train, on='ID')
x_train.head()
# ### Data Preprocessing
# Ho eliminato le istanze che presentavano valori scorretti del campo EDUCATION
# elimino righe con valore 0
x_train["EDUCATION"].value_counts()
x_train = x_train.loc[(x_train["EDUCATION"]!=0)]
# Ho eliminato le istanze che presentavano un valore scorretto del campo MARRIAGE
# elimino valori scorretti
x_train["MARRIAGE"].value_counts()
x_train = x_train.loc[(x_train["MARRIAGE"]!=0)]
# la target è sbilanciata
x_train["default.payment.next.month"].value_counts()
def undersampling(df):
_, uno= df["default.payment.next.month"].value_counts()
df_0 = df[df["default.payment.next.month"] == 0]
df_0 = df_0.sample(uno, random_state=1)
df_1 = df[df["default.payment.next.month"] == 1]
df = pd.concat([df_0, df_1])
return df
# Undersampling per bilanciare la distribuzione della target
x_train = undersampling(x_train)
x_train["default.payment.next.month"].value_counts()
x_train.isna().sum()
# Analizzo le statistiche descrittive e trasformo le features ridimensionandole in un intervallo [0, 1].
x_train.describe()
def preprocess_data(X, scaler=None):
if not scaler:
scaler = MinMaxScaler()
scaler.fit(X)
X = scaler.transform(X)
return X, scaler
y_train = x_train["default.payment.next.month"]
x_train = x_train.drop(['default.payment.next.month'], axis=1)
x_train = x_train.drop(['ID'], axis=1)
x_train_pulito=x_train
y_train_finale=y_train
x_train
x_train, x_scaler = preprocess_data(x_train)
x_train, x_validation, y_train, y_validation = train_test_split(x_train, y_train, test_size=0.1, random_state=0)
def f1(y_true, y_pred): #taken from old keras source code
true_positives = K.sum(K.round(K.clip(y_true * y_pred, 0, 1)))
possible_positives = K.sum(K.round(K.clip(y_true, 0, 1)))
predicted_positives = K.sum(K.round(K.clip(y_pred, 0, 1)))
precision = true_positives / (predicted_positives + K.epsilon())
recall = true_positives / (possible_positives + K.epsilon())
f1_val = 2*(precision*recall)/(precision+recall+K.epsilon())
return f1_val
# +
def gelu(x):
return 0.5 * x * (1 + tf.tanh(tf.sqrt(2 / np.pi) * (x + 0.044715 * tf.pow(x, 3))))
get_custom_objects().update({'gelu': Activation(gelu)})
# Add the alpha parameter of the Leaky-Relu function
get_custom_objects().update({'leaky-relu': Activation(LeakyReLU(alpha=0.2))})
# Add the Swish function
def swish(x, beta = 1):
return (x * sigmoid(beta * x))
get_custom_objects().update({'swish': Activation(swish)})
act_func = ['relu', 'elu', 'leaky-relu', 'selu', 'gelu', 'swish']
# +
initializer = tf.keras.initializers.GlorotUniform(seed=1)
def NeuralNetwork(X, activation):
inputs = keras.Input(shape = (x_train.shape[1]))
x = layers.Dense(13, activation = activation, kernel_initializer=initializer) (inputs)
x = layers.Dense(6, activation = activation, kernel_initializer=initializer) (x)
x = layers.Dense(1, activation= "sigmoid", kernel_initializer=initializer) (x)
model = keras.Model(inputs, x)
model.compile(loss = 'binary_crossentropy', optimizer='adam', metrics=["accuracy", f1])
return model
# -
x_train.shape[1]
# Scelgo l'activation function migliore
result = []
for activation in act_func:
print('\nTraining with -->{0}<-- activation function\n'.format(activation))
model = NeuralNetwork(x_train, activation)
history = model.fit(x_train, y_train,
batch_size=16,
epochs=20,
verbose=1,
validation_data=(x_validation, y_validation))
result.append(history)
K.clear_session()
del model
# +
f1_finale = []
def risultati(activation_function, rmse):
for i in range(len(activation_function)):
print('RMSE of the activation function {} is {}'.format(activation_function[i], round(rmse[i], 5)))
def plot_act_func_results(results, activation_functions = []):
plt.figure(figsize=(10,7))
plt.style.use('dark_background')
plt.figure(figsize=(10,10))
for act_func in results:
plt.plot(act_func.history['val_f1'])
f1_finale.append(act_func.history['val_f1'][-1])
plt.title('Model F1')
plt.ylabel('Validation F1')
plt.xlabel('Epoch')
plt.legend(activation_functions)
plt.show()
plot_act_func_results(result, act_func)
# -
activation= "relu"
# ### Model Definition
inputs = keras.Input(shape = (x_train.shape[1]))
x = layers.Dense(32, activation = activation, kernel_initializer=initializer) (inputs)
x = layers.Dense(16, activation = activation, kernel_initializer=initializer) (x)
x = layers.Dense(1, activation= "sigmoid", kernel_initializer=initializer) (x)
model = keras.Model(inputs, x)
model.compile(loss = 'binary_crossentropy', optimizer='adam', metrics=["accuracy", f1])
model.summary()
# ### Training
history = model.fit(x_train, y_train, batch_size=16, epochs=100, verbose=1, validation_data=(x_validation, y_validation))
history.history['accuracy'][-1]
# +
x_plot = list(range(1,101))
def plot_history(history):
plt.figure()
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.plot(x_plot, history.history['loss'])
plt.plot(x_plot, history.history['val_loss'])
plt.legend(['Training', 'Validation'])
plt.figure()
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.plot(x_plot, history.history['accuracy'])
plt.plot(x_plot, history.history['val_accuracy'])
plt.legend(['Training', 'Validation'], loc='lower right')
plt.show()
plot_history(history)
# -
# ## Regularization
# ### L1
# +
inputs = keras.Input(shape = (x_train.shape[1]))
x = layers.Dense(32, activation = activation, kernel_initializer=initializer, kernel_regularizer=regularizers.l1(0.01)) (inputs)
x = layers.Dense(16, activation = activation, kernel_initializer=initializer, kernel_regularizer=regularizers.l1(0.01)) (x)
x = layers.Dense(1, activation= "sigmoid", kernel_initializer=initializer) (x)
model_l1 = keras.Model(inputs, x)
model_l1.compile(loss = 'binary_crossentropy', optimizer='adam', metrics=["accuracy", f1])
history = model_l1.fit(x_train, y_train, batch_size=16, epochs=100, verbose=0, validation_data=(x_validation, y_validation))
# -
history.history['accuracy'][-1]
# ### L2
# +
inputs = keras.Input(shape = (x_train.shape[1]))
x = layers.Dense(32, activation = activation, kernel_initializer=initializer, kernel_regularizer=regularizers.l2(0.01)) (inputs)
x = layers.Dense(16, activation = activation, kernel_initializer=initializer, kernel_regularizer=regularizers.l2(0.01)) (x)
x = layers.Dense(1, activation= "sigmoid", kernel_initializer=initializer) (x)
model_l2 = keras.Model(inputs, x)
model_l2.compile(loss = 'binary_crossentropy', optimizer='adam', metrics=["accuracy", f1])
history = model_l2.fit(x_train, y_train, batch_size=16, epochs=100, verbose=1, validation_data=(x_validation, y_validation))
# -
history.history['accuracy'][-1]
# ### Dropout
inputs = keras.Input(shape = (x_train.shape[1]))
x = layers.Dense(32, activation = activation, kernel_initializer=initializer) (inputs)
x = layers.Dropout(0.2)(x)
x = layers.Dense(16, activation = activation, kernel_initializer=initializer) (x)
x = layers.Dropout(0.2)(x)
x = layers.Dense(1, activation= "sigmoid", kernel_initializer=initializer) (x)
model_dp = keras.Model(inputs, x)
model_dp.compile(loss = 'binary_crossentropy', optimizer='adam', metrics=["accuracy", f1])
history = model_dp.fit(x_train, y_train, batch_size=16, epochs=100, verbose=1, validation_data=(x_validation, y_validation))
history.history['f1'][-1]
model_dp.summary()
# ### Early Stopping
# +
callback = tf.keras.callbacks.EarlyStopping(monitor='loss', patience=3)
inputs = keras.Input(shape = (x_train.shape[1]))
x = layers.Dense(32, activation = activation, kernel_initializer=initializer) (inputs)
x = layers.Dense(16, activation = activation, kernel_initializer=initializer) (x)
x = layers.Dense(1, activation= "sigmoid", kernel_initializer=initializer) (x)
model_es = keras.Model(inputs, x)
model_es.compile(loss = 'binary_crossentropy', optimizer='adam', metrics=["accuracy", f1])
history = model_es.fit(x_train, y_train, batch_size=16, epochs=1000, verbose=1, callbacks=[callback], validation_data=(x_validation, y_validation))
# -
history.history['f1'][-1]
# ### Weights
# Modello base
print('Layers name:', model.weights[2].name)
print('Layers kernel shape:', model.weights[2].shape)
model.weights[2][0]
# L1
print('Layers name:', model_l1.weights[2].name)
print('Layers kernel shape:', model_l1.weights[2].shape)
model_l1.weights[2][0]
# L2
print('Layers name:', model_l2.weights[2].name)
print('Layers kernel shape:', model_l2.weights[2].shape)
model_l2.weights[2][0]
# Dropout
print('Layers name:', model_dp.weights[2].name)
print('Layers kernel shape:', model_dp.weights[2].shape)
model_dp.weights[2][0]
# Early Stopping
print('Layers name:', model_es.weights[2].name)
print('Layers kernel shape:', model_es.weights[2].shape)
model_es.weights[2][0]
print('Sum of the values of the weights without regularization:', sum(abs(model.weights[2][0])).numpy())
print('Sum of the values of the weights with regularization l1:', sum(abs(model_l1.weights[2][0])).numpy())
print('Sum of the values of the weights with regularization l2:', sum(abs(model_l2.weights[2][0])).numpy())
print('Sum of the values of the weights with dropout:', sum(abs(model_dp.weights[2][0])).numpy())
print('Sum of the values of the weights with early stopping:',sum(abs(model_es.weights[2][0])).numpy())
# ### Test with Feature Selection
x_train_pulito["target"]=y_train_finale
x_train_pulito
x_train_pulito.corr()
prova = x_train_pulito.corr()
prova=prova.loc[(prova["target"]>=0)]
print(prova.index)
x_train=x_train_pulito[prova.index]
y=x_train["target"]
del x_train["target"]
x_train, _ = preprocess_data(x_train)
x_train, x_validation, y_train, y_validation = train_test_split(x_train, y, test_size=0.1, random_state=0)
# +
callback = tf.keras.callbacks.EarlyStopping(monitor='loss', patience=3)
inputs = keras.Input(shape = (x_train.shape[1]))
x = layers.Dense(32, activation = activation, kernel_initializer=initializer) (inputs)
x = layers.Dense(16, activation = activation, kernel_initializer=initializer) (x)
x = layers.Dense(1, activation= "sigmoid", kernel_initializer=initializer) (x)
model_test = keras.Model(inputs, x)
model_test.compile(loss = 'binary_crossentropy', optimizer='adam', metrics=["accuracy", f1])
history = model_test.fit(x_train, y_train, batch_size=16, epochs=1000, verbose=1, callbacks=[callback], validation_data=(x_validation, y_validation))
# -
history.history['f1'][-1]
# ### Test Prediction
x_test = pd.read_csv("X_test.csv", index_col=False)
x_test
x_test["EDUCATION"].value_counts()
x_test = x_test.loc[(x_test["EDUCATION"]!=0)]
x_test["MARRIAGE"].value_counts()
x_test = x_test.loc[(x_test["MARRIAGE"]!=0)]
x_test = x_test.drop(['ID'], axis=1)
x_test, _ = preprocess_data(x_test, x_scaler)
y_test = model_es.predict(x_test)
y_test.round()
# +
file = open("Gabriele_Ferrario_817518_score2.txt", "w")
for row in y_test.round():
np.savetxt(file, row)
file.close()
| assignment2/Gabriele_Ferrario_817518.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="images/usm.jpg" width="480" height="240" align="left"/>
# # MAT281 - Laboratorio N°04
#
# ## Objetivos del laboratorio
#
# * Reforzar conceptos básicos de reducción de dimensionalidad.
# ### <NAME>.
# ### 201510509-K
# ## Contenidos
#
# * [Problema 01](#p1)
#
# <a id='p1'></a>
# ## I.- Problema 01
#
#
# <img src="https://www.goodnewsnetwork.org/wp-content/uploads/2019/07/immunotherapy-vaccine-attacks-cancer-cells-immune-blood-Fotolia_purchased.jpg" width="360" height="360" align="center"/>
#
#
# El **cáncer de mama** es una proliferación maligna de las células epiteliales que revisten los conductos o lobulillos mamarios. Es una enfermedad clonal; donde una célula individual producto de una serie de mutaciones somáticas o de línea germinal adquiere la capacidad de dividirse sin control ni orden, haciendo que se reproduzca hasta formar un tumor. El tumor resultante, que comienza como anomalía leve, pasa a ser grave, invade tejidos vecinos y, finalmente, se propaga a otras partes del cuerpo.
#
# El conjunto de datos se denomina `BC.csv`, el cual contine la información de distintos pacientes con tumosres (benignos o malignos) y algunas características del mismo.
#
#
# Las características se calculan a partir de una imagen digitalizada de un aspirado con aguja fina (FNA) de una masa mamaria. Describen las características de los núcleos celulares presentes en la imagen.
# Los detalles se puede encontrar en [<NAME> and <NAME>: "Robust Linear Programming Discrimination of Two Linearly Inseparable Sets", Optimization Methods and Software 1, 1992, 23-34].
#
#
# Lo primero será cargar el conjunto de datos:
# +
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
# %matplotlib inline
sns.set_palette("deep", desat=.6)
sns.set(rc={'figure.figsize':(11.7,8.27)})
# -
# cargar datos
df = pd.read_csv(os.path.join("data","BC.csv"), sep=",")
df['diagnosis'] = df['diagnosis'] .replace({'M':1,'B':0})
df.head()
# Basado en la información presentada responda las siguientes preguntas:
#
#
# 1. Normalizar para las columnas numéricas con procesamiento **StandardScaler**.
# 2. Realice un gráfico de correlación. Identifique la existencia de colinealidad.
# 3. Realizar un ajuste PCA con **n_components = 10**. Realice un gráfico de la varianza y varianza acumulada. Interprete.
# 4. Devuelva un dataframe con las componentes principales.
# 5. Aplique al menos tres modelos de clasificación. Para cada modelo, calule el valor de sus métricas.
# __Pregunta 1__
# +
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
col_num = []
col_no_num = ['id', 'diagnosis']
for col in df.columns:
if col not in col_no_num:
col_num.append(col)
df[col_num] = scaler.fit_transform(df[col_num])
#Ahora sacaremos las dos primeras columnas, ya que no aportan información relevante para el análisis de los tumores
df = df.drop(['id', 'diagnosis'], axis = 1)
df.head()
# -
# __Pregunta 2__
# +
#Calculamos la correlación de df
corr = df.corr()
#No contamos valores que no sirven
mask = np.zeros_like(corr, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
f, ax = plt.subplots(figsize=(11, 9))
cmap = sns.diverging_palette(10, 250, as_cmap=True)
sns.heatmap(corr, mask=mask, cmap=cmap,
square=True,
linewidths=.5, cbar_kws={"shrink": .5}, ax=ax)
# -
# __Es directo ver que hay cierta linealidad entre algunas variables (los cuadrados con tonos azulados). Pero también hay un gran número de variables que no poseen una linealidad óptima (los cuadrados con tonos rojizos).__
# __Pregunta 3__
# +
#Gráfico de Varianza
# ajustar modelo
from sklearn.decomposition import PCA
pca = PCA(n_components=10)
principalComponents = pca.fit_transform(df.values)
# Graficar varianza por componente
percent_variance = np.round(pca.explained_variance_ratio_* 100, decimals =2)
columns = ['PC1', 'PC2', 'PC3', 'PC4', 'PC5', 'PC6', 'PC7', 'PC8', 'PC9', 'PC10']
plt.figure(figsize=(12,4))
plt.bar(x= range(1,11), height=percent_variance, tick_label=columns)
plt.ylabel('Percentate of Variance Explained')
plt.xlabel('Principal Component')
plt.title('PCA Scree Plot')
plt.show()
# +
#Gráfico de varianzas acumuladas
percent_variance_cum = np.cumsum(percent_variance)
columns = ['PC1', 'PC1 a PC2', 'PC1 a PC3', 'PC1 a PC4', 'PC1 a PC5', 'PC1 a PC6', 'PC1 a PC7', 'PC1 a PC8', 'PC1 a PC9', 'PC1 a PC10']
plt.figure(figsize=(12,4))
plt.bar(x = range(1,11), #Eje x
height = percent_variance_cum, #Valores
tick_label = columns) #Etiquetas eje x
#Etiquetas
plt.ylabel('Percentate of Variance Explained')
plt.xlabel('Principal Component Cumsum')
plt.title('PCA Scree Plot')
plt.show()
# -
# __Vemos del primer gráfico que la varianza de PC1 es la mayor por lejos, y dicha varianza va bajando considerablemente hasta el PC10 que posee una varianza mínima. Por otro lado, el segundo gráfico se ve que las últimas varianzas acumuladas son similares, puesto que, de PC7 a PC10 sus varianzas son mínimas, por lo que no debería haber mucha diferencia.__
# __Pregunta 4__
pca = PCA(n_components=10)
principalComponents = pca.fit_transform(df)
principalDataframe = pd.DataFrame(data = principalComponents, columns = ['PC1', 'PC2', 'PC3', 'PC4', 'PC5',
'PC6', 'PC7', 'PC8', 'PC9', 'PC10'])
principalDataframe
# __Pregunta 5__
| labs/01_python/laboratorio_10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tree_problem_0 as tree
# Do this so that I can update the import file and then run this cell again so it updates here
import imp
imp.reload(tree)
import matplotlib.pyplot as plt
import time
import scipy.stats as st
# +
# Set the global variables, these will be explained
number_of_iterations = 500 # this number is related to the number of operations - if slow, then reduce this
z_range = 8
r = 0.5 # mean regression coefficient
r_s = 0.9 # standard deviation regression coefficient
mean_gen = 0
sd_gen = 1
k_val = -2
percent_step = 0.33
# plotting
fig_size = (12, 8)
# -
# Let's redefine the parent distribution with the normal number of iterations to make things more accurate
parent_distribution = tree.normal_distribution(number_of_iterations, z_range, mean_gen, sd_gen)
total_offspring_distribution1 = tree.final_superimposed_distribution_all_area_adj(parent_distribution, 1, r_s)
one_distribution = tree.one_offspring_distribution(parent_distribution, 250, r, r_s)
many_distribuitons = tree.offspring_distributions(parent_distribution, r, r_s)
super_distribution = tree.superimposed_offspring_distribution(many_distribuitons)
super_par_inc_distribution = tree.normalized_superimposed_distribution_to_parent_increment(super_distribution)
# +
print('PARENT')
print(parent_distribution[0], '\n')
# parent_dist_len = len(parent_distribution)
# print(parent_dist_len)
# parent_mid_index = (parent_dist_len - 1) // 2
# print(parent_distribution[parent_mid_index])
print('ONE OFFSPRING')
print(one_distribution[0])
print('(add parent mean to the end)\n')
print('MANY OFFSPRING')
print(many_distribuitons[0][0])
print('(add parent area to the end)\n')
print('SUPER DISTRIBUTION')
print(super_distribution[0])
print('(reorder things, add parent number and parent bound)\n')
print('SUPER PARENT INC DISTRIBUTION')
print(super_par_inc_distribution[0])
print('(do nothing)\n')
print('TOTAL')
print(total_offspring_distribution1[0])
print('(do nothing)\n')
# offspring_dist_len = len(total_offspring_distribution1)
# print(offspring_dist_len)
# offspring_mid_index = (offspring_dist_len - 1) // 2
# offspring_mean = total_offspring_distribution1[offspring_mid_index][0]
# print(offspring_mid_index)
# print(offspring_mean)
# -
total_offspring_distribution1
n_iterations_large = 500
parent_distribution_im = tree.normal_distribution(n_iterations_large, z_range, mean_gen, sd_gen)
# +
percent_step = 0.333
start = time.time()
step_percentile_five = tree.step_proportion_destined_percentile(parent_distribution_im, r, r_s, percent_step)
end = time.time()
print(end - start)
# +
start = time.time()
tree.step_proportion_attributable_percentile(parent_distribution_im, r, r_s, percent_step)
end = time.time()
print(end - start)
# +
start = time.time()
print(tree.proportion_destined_percentile(parent_distribution_im, r, r_s, 0, 0.2, 0.8, 1.0))
end = time.time()
print(end - start)
# +
start = time.time()
print(tree.proportion_attributable_percentile(parent_distribution_im, r, r_s, 0, 0.2, 0.8, 1.0))
end = time.time()
print(end - start)
# +
error_500 = 12.88387356225678 / 12.24162402732729
error_1000 = 25.83701167962977 / 24.869717301026885
error_2000 = 51.74334249467942 / 50.12570602236689
print(error_500)
print(error_1000)
print(error_2000)
# -
point_eight = st.norm.ppf(0.80)
one_point = 4
point_four = st.norm.ppf(0.4)
point_six = st.norm.ppf(0.60)
plt.figure(figsize=fig_size)
plt.xlim(-4.5, 4.5)
dis_dest = tree.offspring_distributions(parent_distribution, r, r_s, above_k_v_p=point_four, below_k_v_p=point_six)
tree.plot_distributions(dis_dest)
plt.figure(figsize=fig_size)
plt.xlim(-4.5, 4.5)
dis_att = tree.offspring_distributions(parent_distribution, r, r_s, above_k_v_o=point_eight, below_k_v_o=one_point)
tree.plot_distributions(dis_att)
len(dis_dest)
len(dis_att)
# +
plt.figure(figsize=fig_size)
plt.xlim(-4.5, 4.5)
tree.plot_distributions(tree.offspring_distributions(parent_distribution_im, r, r_s, above_k_v_p=-1, below_k_v_p=1))
tree.plot_distributions(tree.offspring_distributions(parent_distribution_im, r, r_s, above_k_v_p=-4, below_k_v_p=-1.5))
tree.plot_distributions(tree.offspring_distributions(parent_distribution_im, r, r_s, above_k_v_p=1.5, below_k_v_p=4))
# plt.savefig('cool_dist.png', dpi=300)
# +
n_parents_a = 1000
n_normal_a = 1000
n_att = n_parents_a * n_normal_a
n_att
# -
n_parents_d = 250
n_normal_d = 1000
n_des = n_parents_d * n_normal_d
n_des
n_att / n_des
# Add proportion attributable stepwise, find some good way of showing it. I'm thinking of five boxplots, or maybe a stack plot? **DONE**
#
# We need to make sure that it's also a good way of displaying the proportion destined stepwise. **DONE**
#
# We also need to talk about how the mean reg coeff is the actually a product of the inheritance coefficient and the parents correlation coefficient.
#
# Also, graph of r and r_s that maintain size of offspring
| archive/tree_project_work.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Before you turn this problem in, make sure everything runs as expected. First, **restart the kernel** (in the menubar, select Kernel$\rightarrow$Restart) and then **run all cells** (in the menubar, select Cell$\rightarrow$Run All).
#
# Make sure you fill in any place that says `YOUR CODE HERE` or "YOUR ANSWER HERE", as well as your name and collaborators below:
NAME = ""
COLLABORATORS = ""
# ---
# <!--NOTEBOOK_HEADER-->
# *This notebook contains material from [PyRosetta](https://RosettaCommons.github.io/PyRosetta.notebooks);
# content is available [on Github](https://github.com/RosettaCommons/PyRosetta.notebooks.git).*
# <!--NAVIGATION-->
# < [PyRosetta Google Drive Setup](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/01.01-PyRosetta-Google-Drive-Setup.ipynb) | [Contents](toc.ipynb) | [Index](index.ipynb) | [How to Get PyRosetta on Your Personal Computer](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/01.03-How-to-Get-Local-PyRosetta.ipynb) ><p><a href="https://colab.research.google.com/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/01.02-PyRosetta-Google-Drive-Usage-Example.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a>
# # PyRosetta Google Drive Usage Example
# After installing PyRosetta in Colab, here is an example for how you would initialize PyRosetta in Colab and use it.
# + colab={} colab_type="code" id="m1KtbxmLzV5p"
# Notebook setup
import sys
if 'google.colab' in sys.modules:
# !pip install pyrosettacolabsetup
import pyrosettacolabsetup
pyrosettacolabsetup.setup()
print ("Notebook is set for PyRosetta use in Colab. Have fun!")
# + colab={} colab_type="code" id="O4-UKoGw1CbX"
from pyrosetta import *
pyrosetta.init()
# + colab={} colab_type="code" id="GMsbR0atbGzM"
test = Pose() # to check that it works
# -
# <!--NAVIGATION-->
# < [PyRosetta Google Drive Setup](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/01.01-PyRosetta-Google-Drive-Setup.ipynb) | [Contents](toc.ipynb) | [Index](index.ipynb) | [How to Get PyRosetta on Your Personal Computer](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/01.03-How-to-Get-Local-PyRosetta.ipynb) ><p><a href="https://colab.research.google.com/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/01.02-PyRosetta-Google-Drive-Usage-Example.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a>
| student-notebooks/01.02-PyRosetta-Google-Drive-Usage-Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # LSTM IMPLEMENTATION
# %run Data_Preprocessing.ipynb
import torch
import torch.nn as nn
from torch.autograd import Variable
from sklearn.model_selection import train_test_split, StratifiedKFold
from sklearn.metrics import accuracy_score
from torch.utils.data import DataLoader, TensorDataset
from keras.layers import Dropout
from keras.models import Sequential
from keras.layers import Dense,LSTM
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
import pandas as pd
y_train.shape
# +
# Initialising the RNN
model = Sequential()
model.add(LSTM(units = 64, return_sequences = True, input_shape = (x_train.shape[1],7)))
model.add(Dropout(0.5))
# Adding a second LSTM layer and Dropout layer
model.add(LSTM(units = 64, return_sequences = True))
model.add(Dropout(0.5))
# Adding a third LSTM layer and Dropout layer
model.add(LSTM(units = 64, return_sequences = True))
model.add(Dropout(0.5))
# Adding a forth LSTM layer and Dropout layer
model.add(LSTM(units = 64, return_sequences = True))
model.add(Dropout(0.5))
# Adding a fifth LSTM layer and Dropout layer
model.add(LSTM(units = 64, return_sequences = True))
model.add(Dropout(0.5))
# Adding a sixth LSTM layer and Dropout layer
model.add(LSTM(units = 64, return_sequences = True))
model.add(Dropout(0.5))
# Adding a seventh LSTM layer and Dropout layer
model.add(LSTM(units = 64, return_sequences = True))
model.add(Dropout(0.2))
# Adding a eighth LSTM layer and Dropout layer
model.add(LSTM(units = 64, return_sequences = True))
model.add(Dropout(0.2))
# Adding a nineth LSTM layer and Dropout layer
model.add(LSTM(units = 64, return_sequences = True))
model.add(Dropout(0.2))
# Adding a tenth LSTM layer and and Dropout layer
model.add(LSTM(units = 64))
model.add(Dropout(0.2))
# Adding the output layer
model.add(Dense(units = 1))
# -
model.summary()
kf = StratifiedKFold(4,shuffle=False,random_state=42)
orignal_y = []
pedicted_y = []
fold = 0
# +
for train,test in kf(X,Y):
fold+=1
# print(f"Fold #{fold}")
x_trainv = X[train]
y_trainv = Y[train]
x_testv = X[test]
y_testv = Y[test]
model.compile(optimizer = 'adam', loss = 'mean_squared_error',metrics=['accuracy'])
history=model.fit(x_trainv, y_trainv, epochs = 30, batch_size = 50,validation_data=(x_testv,y_testv))
pred = model.predict(x_testv)
original_y.append(y_testv)
pred = np.argmax(pred,axis=1)
predicted_y.append(pred)
y_compare = np-argmax(y_testv,axis=1)
score = metrics.accuracy_score(y_compare,pred)
print(f"Fold score (accuracy): {score}")
original_y = np.concatenate(original_y)
predicted_y = np.concatenate(predicted_y)
original_y_compare = np.argmax(original_y,axis=1)
print(f"Final score (accuracy): {score}")
original_y = pd.DataFrame(original_y)
predicted_y = pd.DataFrame(predicted_y)
final_df = pd.concat([original_y,predicted_y],axis=1)
# -
#compile and fit the model on 30 epochs
model.compile(optimizer = 'adam', loss = 'mean_squared_error',metrics=['accuracy'])
history=model.fit(x_train, y_train, epochs = 30, batch_size = 50,validation_data=(x_test,y_test))
res = model.predict(x_test)
loss_l, accuracy_a = model.evaluate(x_test, y_test, verbose=1)
#loss_v, accuracy_v = model.evaluate(x_validate, y_validate, verbose=1)
#print("Validation: accuracy = %f ; loss_v = %f" % (accuracy_v, loss_v))
print("Test: accuracy = %f ; loss = %f" % (accuracy_a, loss_l))
print(accuracy_a, loss_l)
for i in res:
if i[0]>= 0.5:
i[0]=1
else:
i[0]=0
y_val = [0] * 31
for i in range(31):
y_val[i] = y_test[i][2]
plt.scatter(range(31),res,c="r")
plt.scatter(range(31),y_val,c="g")
plt.show()
plt.plot(history.history['loss'])
plt.show()
plt.plot(history.history['accuracy'])
plt.show()
plt.scatter(range(30),history.history['accuracy'],c="r")
plt.scatter(range(30),history.history['loss'],c="g")
plt.show()
t_loss = history.history['loss']
v_loss = history.history['val_loss']
epochs = range(0,30)
plt.plot(epochs, t_loss, 'g', label='Training loss')
plt.plot(epochs, v_loss, 'b', label='validation loss')
plt.title('Training and Validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
| LSTM_Implementation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook was prepared by [<NAME>](https://github.com/rishihot55). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# # Solution Notebook
# ## Problem: Find all valid combinations of n-pairs of parentheses.
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# ## Constraints
#
# * Is the input an integer representing the number of pairs?
# * Yes
# * Can we assume the inputs are valid?
# * No
# * Is the output a list of valid combinations?
# * Yes
# * Should the output have duplicates?
# * No
# * Can we assume this fits memory?
# * Yes
# ## Test Cases
#
# <pre>
# * None -> Exception
# * Negative -> Exception
# * 0 -> []
# * 1 -> ['()']
# * 2 -> ['(())', '()()']
# * 3 -> ['((()))', '(()())', '(())()', '()(())', '()()()']
# </pre>
# # Algorithm
#
# Let `l` and `r` denote the number of left and right parentheses remaining at any given point.
# The algorithm makes use of the following conditions applied recursively:
# * Left braces can be inserted any time, as long as we do not exhaust them i.e. `l > 0`.
# * Right braces can be inserted, as long as the number of right braces remaining is greater than the left braces remaining i.e. `r > l`. Violation of the aforementioned condition produces an unbalanced string of parentheses.
# * If both left and right braces have been exhausted i.e. `l = 0 and r = 0`, then the resultant string produced is balanced.
#
# The algorithm can be rephrased as:
# * Base case: `l = 0 and r = 0`
# - Add the string generated to the result set
# * Case 1: `l > 0`
# - Add a left parenthesis to the parentheses string.
# - Recurse (l - 1, r, new_string, result_set)
# * Case 2: `r > l`
# - Add a right parenthesis to the parentheses string.
# - Recurse (l, r - 1, new_string, result_set)
#
# Complexity:
# * Time: `O(4^n/n^(3/2))`, see [Catalan numbers](https://en.wikipedia.org/wiki/Catalan_number#Applications_in_combinatorics) - 1, 1, 2, 5, 14, 42, 132...
# * Space complexity: `O(n)`, due to the implicit call stack storing a maximum of 2n function calls)
# ## Code
class Parentheses(object):
def find_pair(self, num_pairs):
if num_pairs is None:
raise TypeError('num_pairs cannot be None')
if num_pairs < 0:
raise ValueError('num_pairs cannot be < 0')
if not num_pairs:
return []
results = []
curr_results = []
self._find_pair(num_pairs, num_pairs, curr_results, results)
return results
def _find_pair(self, nleft, nright, curr_results, results):
if nleft == 0 and nright == 0:
results.append(''.join(curr_results))
else:
if nleft >= 0:
self._find_pair(nleft-1, nright, curr_results+['('], results)
if nright > nleft:
self._find_pair(nleft, nright-1, curr_results+[')'], results)
# ## Unit Test
# +
# %%writefile test_n_pairs_parentheses.py
from nose.tools import assert_equal, assert_raises
class TestPairParentheses(object):
def test_pair_parentheses(self):
parentheses = Parentheses()
assert_raises(TypeError, parentheses.find_pair, None)
assert_raises(ValueError, parentheses.find_pair, -1)
assert_equal(parentheses.find_pair(0), [])
assert_equal(parentheses.find_pair(1), ['()'])
assert_equal(parentheses.find_pair(2), ['(())',
'()()'])
assert_equal(parentheses.find_pair(3), ['((()))',
'(()())',
'(())()',
'()(())',
'()()()'])
print('Success: test_pair_parentheses')
def main():
test = TestPairParentheses()
test.test_pair_parentheses()
if __name__ == '__main__':
main()
# -
# %run -i test_n_pairs_parentheses.py
| recursion_dynamic/n_pairs_parentheses/n_pairs_parentheses_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Each major email provider has theit own SMTP(Simple mail Transfer protocol) server
#
# Provider | SMTP server domain name
# -- | --
# Gmail | smtp.gmail.com
# Yahoo mail | smtp.mail.yahoo.com
#
# For Gmail users, we will need to generate an app password instead of normal password.
# This let's gmail know that the python script attempting to access the account is authorized by you.
#
# ## Sending Emails
# Generate App password for your gmail
# https://support.google.com/accounts/answer/185833?hl=en
# +
## eysackwdogjanekv
# -
import smtplib
mail_server = smtplib.SMTP('smtp.gmail.com',587)
mail_server.ehlo()
mail_server.starttls()
password = input('Please provide your password:')
import getpass
email = getpass.getpass('Please provide the email:')
password = getpass.getpass('Please provide the password:')
mail_server.login(email,password)
from_address = email
to_address = '<EMAIL>'
subject = 'Hi-Python Script'
message = 'Test Script'
msg = 'Subject: '+subject+'\n'+message
mail_server.sendmail(from_address, to_address, msg)
mail_server.quit()
# ## Receiving mails
import imaplib
mail_server = imaplib.IMAP4_SSL('imap.gmail.com')
import getpass
email = getpass.getpass('Enter your email address:')
password = getpass.getpass('Enter your password:')
mail_server.login(email,password)
mail_server.list()
mail_server.select('INBOX')
typ, data = mail_server.search(None, 'SUBJECT Test-Python')
typ
data
email_id = data[0]
email_id
result, email_data = mail_server.fetch(email_id, '(RFC822)')
# +
## email_data
# -
result
raw_email = email_data[0][1]
raw_email_string = raw_email.decode('utf-8')
import email
email_message = email.message_from_string(raw_email_string)
email_message
for part in email_message.walk():
if part.get_content_type() == 'text/plain':
body = part.get_payload(decode = True)
print(body)
| docs/Notebooks/24.emails.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # First model with scikit-learn
#
# In this notebook, we present how to build predictive models on tabular
# datasets, with only numerical features.
#
# In particular we will highlight:
#
# * the scikit-learn API: `.fit(X, y)`/`.predict(X)`/`.score(X, y)`;
# * how to evaluate the statistical performance of a model with a train-test
# split.
#
# ## Loading the dataset with Pandas
#
# We will use the same dataset "adult_census" described in the previous
# notebook. For more details about the dataset see
# <http://www.openml.org/d/1590>.
#
# Numerical data is the most natural type of data used in machine learning and
# can (almost) directly be fed into predictive models. We will load a
# subset of the original data with only the numerical columns.
# +
import pandas as pd
adult_census = pd.read_csv("../datasets/adult-census-numeric.csv")
# -
# Let's have a look at the first records of this dataframe:
adult_census.head()
# We see that this CSV file contains all information: the target that we would
# like to predict (i.e. `"class"`) and the data that we want to use to train
# our predictive model (i.e. the remaining columns). The first step is to
# separate columns to get on one side the target and on the other side the
# data.
#
# ## Separate the data and the target
target_name = "class"
target = adult_census[target_name]
target
data = adult_census.drop(columns=[target_name, ])
data.head()
# We can now linger on the variables, also denominated features, that we will
# use to build our predictive model. In addition, we can also check how many
# samples are available in our dataset.
data.columns
print(f"The dataset contains {data.shape[0]} samples and "
f"{data.shape[1]} features")
# ## Fit a model and make predictions
#
# We will build a classification model using the "K-nearest neighbors"
# strategy. To predict the target of a new sample, a k-nearest neighbors takes
# into account its `k` closest samples in the training set and predicts the
# majority target of these samples.
#
# The `fit` method is called to train the model from the input
# (features) and target data.
#
# <div class="admonition caution alert alert-warning">
# <p class="first admonition-title" style="font-weight: bold;">Caution!</p>
# <p class="last">We use a K-nearest neighbors here. However, be aware that it is seldom useful
# in practice. We use it because it is an intuitive algorithm. In the next
# notebook, we will introduce better models.</p>
# </div>
# +
from sklearn.neighbors import KNeighborsClassifier
model = KNeighborsClassifier()
model.fit(data, target)
# -
# Learning can be represented as follows:
#
# 
#
# The method `fit` is composed of two elements: (i) a **learning algorithm**
# and (ii) some **model states**. The learning algorithm takes the training
# data and training target as input and set the model states. These model
# states will be used later to either predict (for classifier and regressor) or
# transform data (for transformers).
#
# Both the learning algorithm and the type of model states are specific to each
# type of models.
# <div class="admonition caution alert alert-warning">
# <p class="first admonition-title" style="font-weight: bold;">Caution!</p>
# <p class="last">Here and later, we use the name <tt class="docutils literal">data</tt> and <tt class="docutils literal">target</tt> to be explicit. In
# scikit-learn documentation, <tt class="docutils literal">data</tt> is commonly named <tt class="docutils literal">X</tt> and <tt class="docutils literal">target</tt> is
# commonly called <tt class="docutils literal">y</tt>.</p>
# </div>
# <div class="admonition tip alert alert-warning">
# <p class="first admonition-title" style="font-weight: bold;">Tip</p>
# <p>In the notebook, we will use the following terminology:</p>
# <ul class="last simple">
# <li>a predictor corresponds to a classifier or a regressor</li>
# <li>a predictive model (or model) corresponds to a succession of steps made of
# some preprocessing steps followed by a predictor. Sometimes, no
# preprocessing is required.</li>
# <li>an estimator corresponds to any scikit-learn object, transformer,
# classifier, or regressor.</li>
# </ul>
# </div>
# Let's use our model to make some predictions using the same dataset.
target_predicted = model.predict(data)
# We can illustrate the prediction mechanism as follows:
#
# 
#
# To predict, a model uses a **prediction function** that will use the input
# data together with the model states. As for the learning algorithm and the
# model states, the prediction function is specific for each type of model.
# Let's now have a look at the computed predictions. For the sake of
# simplicity, we will look at the five first predicted targets.
target_predicted[:5]
# Indeed, we can compare these predictions to the actual data...
target[:5]
# ...and we could even check if the predictions agree with the real targets:
target[:5] == target_predicted[:5]
print(f"Number of correct prediction: "
f"{(target[:5] == target_predicted[:5]).sum()} / 5")
# Here, we see that our model makes a mistake when predicting for the first
# sample.
#
# To get a better assessment, we can compute the average success rate.
(target == target_predicted).mean()
# But, can this evaluation be trusted, or is it too good to be true?
#
# ## Train-test data split
#
# When building a machine learning model, it is important to evaluate the
# trained model on data that was not used to fit it, as generalization is
# more than memorization (meaning we want a rule that generalizes to new data,
# without comparing to data we memorized).
# It is harder to conclude on never-seen instances than on already seen ones.
#
# Correct evaluation is easily done by leaving out a subset of the data when
# training the model and using it afterwards for model evaluation.
# The data used to fit a model is called training data while the data used to
# assess a model is called testing data.
#
# We can load more data, which was actually left-out from the original data
# set.
adult_census_test = pd.read_csv('../datasets/adult-census-numeric-test.csv')
# From this new data, we separate out input features and the target to predict,
# as in the beginning of this notebook.
target_test = adult_census_test[target_name]
data_test = adult_census_test.drop(columns=[target_name, ])
# We can check the number of features and samples available in this new set.
print(f"The testing dataset contains {data_test.shape[0]} samples and "
f"{data_test.shape[1]} features")
# <div class="admonition note alert alert-info">
# <p class="first admonition-title" style="font-weight: bold;">Note</p>
# <p class="last">Scikit-learn provides a helper function <tt class="docutils literal">train_test_split</tt> which
# can be used to split the dataset into a training and a testing set. It will
# also ensure that the data are shuffled randomly before splitting the data.</p>
# </div>
#
# Instead of computing the prediction and manually computing the average
# success rate, we can use the method `score`. When dealing with classifiers
# this method returns their performance metric.
# +
accuracy = model.score(data_test, target_test)
model_name = model.__class__.__name__
print(f"The test accuracy using a {model_name} is "
f"{accuracy:.3f}")
# -
# Let's check the underlying mechanism when the `score` method is called:
#
# 
#
# To compute the score, the predictor first computes the predictions (using
# the `predict` method) and then uses a scoring function to compare the
# true target `y` and the predictions. Finally, the score is returned.
# If we compare with the accuracy obtained by wrongly evaluating the model
# on the training set, we find that this evaluation was indeed optimistic
# compared to the score obtained on an held-out test set.
#
# It shows the importance to always testing the statistical performance of
# predictive models on a different set than the one used to train these models.
# We will discuss later in more details how predictive models should be
# evaluated.
# <div class="admonition note alert alert-info">
# <p class="first admonition-title" style="font-weight: bold;">Note</p>
# <p class="last">In this MOOC, we will refer to <strong>statistical performance</strong> of a model when
# referring to the test score or test error obtained by comparing the
# prediction of a model and the true targets. Equivalent terms for
# <strong>statistical performance</strong> are predictive performance and generalization
# performance. We will refer to <strong>computational performance</strong> of a predictive
# model when accessing the computational costs of training a predictive model
# or using it to make predictions.</p>
# </div>
# In this notebook we:
#
# * fitted a **k-nearest neighbors** model on a training dataset;
# * evaluated its statistical performance on the testing data;
# * introduced the scikit-learn API `.fit(X, y)` (to train a model),
# `.predict(X)` (to make predictions) and `.score(X, y)`
# (to evaluate a model).
| notebooks/02_numerical_pipeline_introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
# +
# !mkdir tensorflow models
# !pip install -q numpy opencv-python matplotlib
# !wget --quiet https://www.dropbox.com/s/07p84k7q4kxwc02/tensorflow-1.13.2-cp37-cp37m-linux_x86_64.whl?dl=1 -O ./tensorflow/tensorflow-1.13.2-cp37-cp37m-linux_x86_64.whl
# !pip install -q ./tensorflow/tensorflow-1.13.2-cp37-cp37m-linux_x86_64.whl
# !curl --silent --header 'Host: raw.githubusercontent.com' --user-agent 'Mozilla/5.0 (X11; Ubuntu; Linux x86_64; rv:69.0) Gecko/20100101 Firefox/69.0' --header 'Accept: text/html,application/xhtml+xml,application/xml;q=0.9,*/*;q=0.8' --header 'Accept-Language: en-US,en;q=0.5' --referer 'https://github.com/google/mediapipe/blob/master/mediapipe/models/palm_detection.tflite' --header 'Upgrade-Insecure-Requests: 1' 'https://raw.githubusercontent.com/google/mediapipe/master/mediapipe/models/palm_detection.tflite' --output './models/palm_detection.tflite'
# !curl --silent --header 'Host: raw.githubusercontent.com' --user-agent 'Mozilla/5.0 (X11; Ubuntu; Linux x86_64; rv:69.0) Gecko/20100101 Firefox/69.0' --header 'Accept: text/html,application/xhtml+xml,application/xml;q=0.9,*/*;q=0.8' --header 'Accept-Language: en-US,en;q=0.5' --referer 'https://github.com/google/mediapipe/blob/master/mediapipe/models/hand_landmark.tflite' --header 'Upgrade-Insecure-Requests: 1' 'https://raw.githubusercontent.com/google/mediapipe/master/mediapipe/models/hand_landmark.tflite' --output './models/hand_landmark.tflite'
# !wget --quiet https://upload.wikimedia.org/wikipedia/commons/9/99/JPEG_20190317_010417.jpg -O ./data/test_img.jpg
# -
from hand_tracker import HandTracker
import cv2
import matplotlib.pyplot as plt
from matplotlib.patches import Polygon
# %matplotlib inline
palm_model_path = "./models/palm_detection.tflite"
landmark_model_path = "./models/hand_landmark.tflite"
anchors_path = "./data/anchors.csv"
img = cv2.imread('./data/test_img.jpg')[:,:,::-1]
# box_shift determines
detector = HandTracker(palm_model_path, landmark_model_path, anchors_path,
box_shift=0.2, box_enlarge=1.3)
# +
kp, box = detector(img)
f,ax = plt.subplots(1,1, figsize=(10, 10))
ax.imshow(img)
ax.scatter(kp[:,0], kp[:,1])
ax.add_patch(Polygon(box, color="#00ff00", fill=False))
# -
| hand_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %pylab inline
from IPython.display import Audio
import librosa
import scipy as sp
from numba import jit
from scipy.spatial import distance
figsize(20,6)
prefix="baseline"
def filepre(nm):
return "tmp/"+prefix+"_"+nm
from IPython.display import clear_output
# -
import tensorflow as tf
tf.enable_eager_execution()
from tensorflow.python.client import device_lib
tfdevice='/device:GPU:0'
device_lib.list_local_devices()
# +
sr = 22050
def nrmse(output,target):
assert(output.shape[0] == target.shape[0])
combinedVar = 0.5 * (np.var(target, ddof=1) + np.var(output, ddof=1))
errorSignal = output - target
return np.sqrt(np.mean(errorSignal ** 2) / combinedVar)
def generateInternalWeights(nInternalUnits, connectivity):
success = False
internalWeights = 0
while success == False:
try:
internalWeights = np.random.randn(nInternalUnits,nInternalUnits) * (np.random.random((nInternalUnits,nInternalUnits)) < connectivity)
specRad = max(abs(np.linalg.eig(internalWeights)[0]))
if (specRad > 0):
internalWeights = internalWeights / specRad
success = True
except e:
print(e)
return internalWeights
pLoop = lambda n,p: p[n%p.size]
# -
from datetime import datetime
import os
resultsFolderName = "csynthResults/results" + str(datetime.now()) + "/"
os.mkdir(resultsFolderName)
relevant_path = "audios/ixi"
ixiFiles = [fn for fn in os.listdir(relevant_path)
if fn.endswith('wav')]
# ixiFiles = ['909a_22k.wav']
print(ixiFiles)
def ixistr(id):
return str(id) + "_" + ixiFiles[id] + "_"
def compareMFCCs(seq1, seq2):
fftSize=2048
hop=64
melspec = librosa.feature.melspectrogram(y=seq1, sr=sr, n_fft=fftSize,hop_length=hop)
mfccs = librosa.feature.mfcc(S=melspec,n_mfcc=20)[1:,:]
melspec2 = librosa.feature.melspectrogram(y=seq2, sr=sr, n_fft=fftSize,hop_length=hop)
mfccs2 = librosa.feature.mfcc(S=melspec2,n_mfcc=20)[1:,:]
return nrmse(mfccs.flatten(), mfccs2.flatten())
# +
def computeConceptor(p, net, i_pattern, alpha):
print('Computing conceptor, alpha: ', alpha)
# Cs = np.zeros((4, 1), dtype=np.object)
R = net['patternRs'][0,i_pattern]
[U,s,V] = svd(R)
# s = svd(R, compute_uv=False)
S = tf.diag(s)
# Snew = (S.dot(linalg.inv(S + pow(alpha, -2) * np.eye(p['N']))))
# C = U.dot(Snew).dot(U.T);
sinv = tf.matrix_inverse(tf.add(S, tf.multiply(double(pow(alpha, -2)), tf.eye(p['N'], dtype=float64))))
Snew = tf.matmul(S,sinv)
# Snew = tf.matmul(Snew, tf.eye(p['N'], dtype=float64))
# Snew = Snew.numpy()
# Snew = (S * linalg.inv(S + pow(alpha, -2) * np.eye(p['N'])))
tfU = tf.constant(U)
C = tf.matmul(tfU,Snew)
C = tf.matmul(C,tfU, adjoint_b=True)
return C
def testConceptor(p, C, net, recallTestLength, tfW, tfWbias):
with tf.device(tfdevice):
trials = 1
attens = np.zeros(trials)
LR = array(p['LR'])
LROneMinus = array(1.0 - p['LR'])
tfLR = tf.constant(LR)
tfLROneMinus = tf.constant(LROneMinus)
# tfWbias = tf.constant(net['Wbias'])
# tfW = tf.constant(net['W'])
# tfC = tf.constant(C)
for i_trial in range(trials):
x_CTestPL = np.zeros((p['N'], recallTestLength))
z_CTestPL = np.zeros((p['N'], recallTestLength))
# tfx_CTestPL = tf.TensorArray(tfW.dtype,p['N'])
# tfz_CTestPL = tf.TensorArray(tfW.dtype,p['N'])
x = tf.constant(0.5 * np.random.randn(p['N'],1))
for n in range(recallTestLength + p['washoutLength']):
xOld = tf.constant(x)
Wtarget = tf.matmul(tfW, x)
leakTerm = tf.multiply(LROneMinus,xOld)
newX =tf.tanh(tf.add(Wtarget, tfWbias))
newXLeaked = tf.multiply(LR,newX)
z = tf.add(leakTerm,newXLeaked)
x = tf.matmul(C,z)
if (n > p['washoutLength']):
# tfx_CTestPL.write(n-p['washoutLength'], tf.transpose(x))
# tfz_CTestPL.write(n-p['washoutLength'], tf.transpose(z))
x_CTestPL[:,n-p['washoutLength']] = tf.transpose(x).numpy()
z_CTestPL[:,n-p['washoutLength']] = tf.transpose(z).numpy()
# x_CTestPL = tfx_CTestPL.gather(tf.range(0,recallTestLength,1))
# z_CTestPL = tfz_CTestPL.gather(tf.range(0,recallTestLength,1))
attenuation = np.mean(pow(np.linalg.norm(z_CTestPL[:,:] - x_CTestPL[:,:], axis=1),2)) / np.mean(pow(np.linalg.norm(z_CTestPL[:,:], axis=1),2))
attens[i_trial] = attenuation
return np.mean(attens)
# -
def makeLoadedNetwork_v2(p):
Netconnectivity = 1
if p['N'] > 20:
Netconnectivity = 10.0/p['N'];
WstarRaw = generateInternalWeights(p['N'], Netconnectivity)
WinRaw = 2 * (np.random.rand(p['N'], 1) - 0.5)
WbiasRaw = 2 * (np.random.rand(p['N'], 1) - 0.5)
#Scale raw weights
Wstar = p['NetSR'] * WstarRaw;
Win = p['NetinpScaling'] * WinRaw;
Wbias = p['BiasScaling'] * WbiasRaw;
I = np.eye(p['N'])
x = np.zeros((p['N'],1))
allTrainxArgs = np.zeros((p['N'] + 1, 0));
allTrainOldxArgs = np.zeros((p['N'], 0));
allTrainWtargets = np.zeros((p['N'], 0));
allTrainOuts = np.zeros((1, 0));
patternRs = np.zeros((1, p['patts'].shape[0]), dtype=np.object)
print('Loading patterns: ', end='')
LR = array(p['LR'])
LROneMinus = array(1.0 - p['LR'])
for i_pattern in range(p['patts'].shape[0]):
print(i_pattern, " ", end='')
patt = p['patts'][i_pattern]
pattLearnLen = patt.size * p['learnLength']
xCollector = np.zeros((p['N'] + 1, pattLearnLen));
xOldCollector = np.zeros((p['N'], pattLearnLen));
WTargetCollector = np.zeros((p['N'], pattLearnLen));
pCollector = np.zeros((1, pattLearnLen));
x = np.zeros((p['N'], 1))
tfWstar = tf.constant(Wstar)
tfWin = tf.constant(Win)
with tf.device(tfdevice):
for n in range(p['washoutLength'] + pattLearnLen):
u = patt.take(n, mode='wrap')
xOld = x
Wtarget = (Wstar.dot(x)) + (Win.dot(u))
# wstarx=tf.matmul(tfWstar,x)
# winu = tf.multiply(tfWin,u)
# Wtarget = tf.add(wstarx, winu)
leakTerm = LROneMinus.dot(xOld)
newX =tanh(Wtarget + Wbias)
newXLeaked = LR.dot(newX)
x = leakTerm + newXLeaked
# xOldLR = tf.multiply(tf.constant(1.0-LR, dtype=float64), xOld)
# biasedTarget = tf.add(Wtarget, Wbias)
# biasedTarget = tf.tanh(biasedTarget)
# biasedTargetLR = tf.multiply(tf.constant(LR, dtype=float64), biasedTarget)
# x = tf.add(xOldLR, biasedTargetLR)
if n >= p['washoutLength']:
xCollector[:, n - p['washoutLength']] = np.concatenate((x[:,0], np.array([1])))
xOldCollector[:, n - p['washoutLength']] = xOld[:,0]
WTargetCollector[:, n - p['washoutLength']] = Wtarget[:,0]
pCollector[0, n - p['washoutLength']] = u
uOld = u
R = xCollector[0:-1].dot(xCollector[0:-1].T) / pattLearnLen
patternRs[0,i_pattern] = R
allTrainxArgs = np.concatenate((allTrainxArgs, xCollector), axis=1)
allTrainOldxArgs = np.concatenate((allTrainOldxArgs, xOldCollector), axis=1)
allTrainOuts = np.concatenate((allTrainOuts, pCollector), axis=1)
allTrainWtargets = np.concatenate((allTrainWtargets, WTargetCollector), axis=1)
Wout = (linalg.inv(allTrainxArgs.dot(allTrainxArgs.conj().T) +
(p['TychonovAlphaReadout'] * np.eye(p['N'] + 1))).dot(allTrainxArgs).dot(allTrainOuts.conj().T)).conj().T
outsRecovered = Wout.dot(allTrainxArgs);
NRMSE_readout = mean(nrmse(outsRecovered, allTrainOuts))
absSize_readout = mean(mean(abs(Wout), axis=0))
print("\nNRMSE readout: ", NRMSE_readout, " :: ", end='')
print("absSize readout: ", absSize_readout)
W = (linalg.inv(allTrainOldxArgs.dot(allTrainOldxArgs.conj().T) +
(p['TychonovAlpha'] * np.eye(p['N']))).dot(allTrainOldxArgs).dot(allTrainWtargets.conj().T)).conj().T
NRMSE_W = mean(nrmse(W.dot(allTrainOldxArgs), allTrainWtargets))
absSize_W = mean(mean(abs(W), axis=0))
print("NRMSE W: ", NRMSE_W, " :: ", end='')
print("absSize W: ", absSize_W)
data ={k: v for k, v in locals().items() if k in
('p','Win','Wstar', 'Wbias','NRMSE_W', 'absSize_W','patternRs','W',
'Wout','NRMSE_readout', 'absSize_readout')}
return data
# +
def render(p, patternCs, bestNet, lrMod=1.0, speed=1.0, xFade=0.05, srMod=1):
audio = np.zeros(0)
x = 0.5 * np.random.randn(p['N'],1)
C = patternCs[0]
LR = p['LR']
LR = array(LR * lrMod)
LROneMinus = array(1.0 - p['LR'])
Wmod = bestNet['W'] * srMod
#run to washout
for n in range(p['washoutLength']):
xOld = x
Wtarget = (Wmod.dot(x))
z = (LROneMinus.dot(xOld)) + (LR.dot(tanh(Wtarget + bestNet['Wbias'])))
x = C.dot(z)
for i_patt in range(p['patts'].shape[0]):
xFadeTime=int(p['patts'][i_patt].shape[0] * xFade)
for n in range(int(p['patts'][i_patt].shape[0] * speed)):
C = patternCs[i_patt]
v=int(p['patts'][i_patt].shape[0] * speed)
stepL = min(v - n - 1, xFadeTime)
stepU = min(n, xFadeTime)
m1 = 1.0
if(n > v-xFadeTime-1 and i_patt < p['patts'].shape[0]-1):
m1 = (stepL + stepU) / (2*xFadeTime)
nextC = patternCs[i_patt+1]
C = (m1 * C) + ((1.0-m1) * nextC)
else:
if (n < xFadeTime and i_patt > 0):
m1 = 0.5 - (n / (2*xFadeTime))
prevC = patternCs[i_patt-1]
C = (m1 * prevC) + ((1.0-m1) * C)
# else:
# C = cNet['Cs'][0,0]
xOld = x
Wtarget = (Wmod.dot(x))
z = (LROneMinus.dot(xOld)) + (LR.dot(tanh(Wtarget + bestNet['Wbias'])))
x = C.dot(z)
newSample = bestNet['Wout'].dot(np.concatenate((x[:,0], np.array([1]))))
audio = np.concatenate((audio, newSample))
return audio
# +
def evalModel(genome, patterns, patternLengths, orgAudio, N=900):
LR =genome[0]
modelParams = {'N':N, 'NetSR':1.5, 'NetinpScaling':1.2,'BiasScaling':0.3, 'TychonovAlpha':0.0001,
'washoutLength':50, 'learnLength':4, 'TychonovAlphaReadout':0.0001,
'LR': LR,
'patts':patterns
}
newNetwork = makeLoadedNetwork_v2(modelParams)
with tf.device(tfdevice):
tfWbias = tf.constant(newNetwork['Wbias'])
tfW = tf.constant(newNetwork['W'])
import scipy
def fitnessf(aperture, *args):
print('Pattern: ', args[0])
params = args[1]
net = args[2]
try:
C = computeConceptor(params, net, args[0], aperture)
except:
print("Exception when computing conceptor")
return 999
atten = testConceptor(params, C, net, params['patts'][args[0]].size * params['learnLength'], args[3], args[4])
return atten
apertures = [scipy.optimize.fminbound(fitnessf, 0,1000, disp=2, xtol=15, args = (x,modelParams, newNetwork, tfW, tfWbias))
for x in np.arange(modelParams['patts'].shape[0])]
#store conceptors with calculated apertures
patternCs = np.zeros(len(apertures), dtype=np.object)
for i_patt in range(patternCs.size):
patternCs[i_patt] = computeConceptor(modelParams, newNetwork, i_patt, apertures[i_patt]).numpy()
# figsize(20,3)
audio = render(modelParams, patternCs, newNetwork, 1.0,1.0, 0.05)
error = compareMFCCs(audio, orgAudio)
# plot(audio)
return {"error":error, "waveform":audio, 'apertures':apertures, 'net':newNetwork}
# -
def evalFitness(genome, data):
modelData = evalModel(genome, data['patterns'], data['patternLengths'], data['orgAudio'], data['N'])
if ("winner" in data):
if modelData['error'] < data['winner']['error']:
data['winner'] = modelData
else:
data['winner'] = modelData
return modelData['error']
# +
startTS = datetime.now()
testing = False
learningRates = linspace(0.05,0.95,10)
# for currentIxi in range(len(ixiFiles)):
for currentIxi in range(1) if testing else range(len(ixiFiles)):
def log(msg):
f = open(resultsFolderName + ixistr(currentIxi) + "searchLog.txt", "a")
f.write(str(datetime.now()) + ":")
f.write(msg)
f.write('\r\n')
f.close()
print(msg)
clear_output()
print("loading: ", ixiFiles[currentIxi])
y, sr = librosa.load("audios/" + ixiFiles[currentIxi], sr=22050)
y = y[:5000] / np.max(y) * 0.5
print(sr)
#divide out windows
patterns = []
minPatternSize = 9
lastCrossing=0
for i in range(y.shape[0]-1):
if (i-lastCrossing) > minPatternSize and y[i] >=0 and y[i+1] < 0:
print(i)
segment = y[lastCrossing:i]
patterns.append(segment)
lastCrossing = i
#convert to numpy
patterns = np.array(patterns, dtype=np.object)
maxPatterns = 150
patterns = patterns[:maxPatterns]
patternLengths = [x.shape[0] for x in patterns]
y = y[:sum(patternLengths)]
maxPatternLen = np.max(patternLengths)
minPatternLen = np.min(patternLengths)
print("max length: ", maxPatternLen)
print("min length: ", minPatternLen)
# for p in patterns:
# plot(p)
print(patterns.shape)
data = {'patterns':patterns, 'patternLengths':patternLengths, 'orgAudio':y}
def onEpochStart():
clear_output()
print('Runtime:', print(datetime.now() - startTS))
print(currentIxi, '/', len(ixiFiles), ' : ', ixiFiles[currentIxi])
log(str(scores))
# brute force search of learning rates
# do the search at low res with smaller N
data['N'] = 600
scores = zeros_like(learningRates)
for i,l in enumerate(learningRates):
trials = 5
trialScores = zeros(trials)
for trial in range(trials):
onEpochStart()
log("lo res trial" + str(trial))
log(str(scores))
log(str(trialScores))
trialScores[trial] = evalFitness(array([l]), data)
scores[i] = np.median(trialScores)
log(str(trialScores))
winningScore = np.min(scores)
log("Winning score: " + str(winningScore))
bestLR = learningRates[np.argmin(scores)]
# now generate the best of x at high res to find a good network
data['N'] = 900
trials = 10
scores = zeros(trials)
del data['winner']
for trial in range(trials):
onEpochStart()
log("hi res trial " + str(trial))
scores[trial] = evalFitness(array([bestLR]), data)
log(str(scores))
plt.close()
figsize(20,4)
plt.xlabel("Time (samples)", fontsize=20)
plt.ylabel("Amplitude", fontsize=20)
plt.xticks(fontsize=18, rotation=0)
plt.yticks(fontsize=18, rotation=0)
plot(y, label='Original', alpha=0.6)
plot(data['winner']['waveform'], alpha=1.0, label='Reconstruction')
plt.legend(fontsize=20)
plt.savefig(resultsFolderName + ixistr(currentIxi) + "compare.pdf", bbox_inches='tight')
librosa.output.write_wav(resultsFolderName + ixistr(currentIxi) + "org.wav",y, sr)
librosa.output.write_wav(resultsFolderName + ixistr(currentIxi) + "recon.wav", data['winner']['waveform'], sr)
import dill as pickle
with open(resultsFolderName + ixistr(currentIxi) + r"model.dill.pickled", "wb") as output_file:
pickle.dump({'winner':data['winner'], 'original':y, 'patterns':patterns}, output_file, protocol=0)
# -
| conceptorSynthesisExpt_GPU.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Benchmarking your code
def fun():
max(range(1000))
# Using magic functions of Jupyter and `timeit`
#
# * https://docs.python.org/3.5/library/timeit.html
# * https://ipython.org/ipython-doc/3/interactive/magics.html#magic-time
# %%timeit
fun()
# %%time
fun()
# ## Exercises
# 1. What is the fastest way to download 100 pages from index.hu?
# 2. How to calculate the factors of 1000 random integers effectively using `factorize_naive` function below?
import requests
def get_page(url):
response = requests.request(url=url, method="GET")
return response
get_page("http://index.hu")
def factorize_naive(n):
""" A naive factorization method. Take integer 'n', return list of
factors.
"""
if n < 2:
return []
factors = []
p = 2
while True:
if n == 1:
return factors
r = n % p
if r == 0:
factors.append(p)
n = n // p
elif p * p >= n:
factors.append(n)
return factors
elif p > 2:
# Advance in steps of 2 over odd numbers
p += 2
else:
# If p == 2, get to 3
p += 1
assert False, "unreachable"
| AdvancedPython/StandardLibrary/Concurrency exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.cluster import KMeans
from sklearn.svm import SVC
from sklearn.decomposition import PCA
from sklearn import metrics
# from mlxtend.plotting import plot_decision_regions
from sklearn import preprocessing
from sklearn.linear_model import LogisticRegression
from ast import literal_eval
import warnings
import numpy as np
from collections import OrderedDict
from lob_data_utils import lob, db_result, model, roc_results
from lob_data_utils.svm_calculation import lob_svm
import os
sns.set_style('whitegrid')
warnings.filterwarnings('ignore')
# -
data_length = 10000
rs_params = [(0.1, 1.0)]
stocks = list(roc_results.result_cv_10000.keys())
data_dir = 'res_pca_gdf_que3'
# +
def get_mean_scores(scores: dict) -> dict:
mean_scores = {}
for k, v in scores.items():
mean_scores[k] = np.mean(v)
return mean_scores
def get_score_for_clf(clf, df_test):
x_test = df_test[['queue_imbalance']]
y_test = df_test['mid_price_indicator'].values
return model.test_model(clf, x_test, y_test)
def get_logistic_regression(stock, data_length):
df, df_test = lob.load_prepared_data(
stock, data_dir='../gaussian_filter/data', cv=False, length=data_length)
clf = LogisticRegression()
train_x = df[['queue_imbalance']]
scores = model.validate_model(clf, train_x, df['mid_price_indicator'])
res = {
**get_mean_scores(scores),
'stock': stock,
'kernel': 'logistic',
}
test_scores = get_score_for_clf(clf, df_test)
return {**res, **test_scores}
# -
df_res = pd.DataFrame()
for stock in stocks:
for r, s in rs_params:
#pd.read_csv('svm_features_{}_len{}_r{}_s{}.csv'.format(stock, data_length, r, s))
filename = data_dir + '/svm_pca_gdf_{}_len{}_r{}_s{}_K20-30.csv'.format(stock, data_length, r, s)
if os.path.exists(filename):
df_temp = pd.read_csv(filename)
df_temp['r'] = [r] * len(df_temp)
df_temp['s'] = [s] * len(df_temp)
df_res = df_res.append(df_temp)
#df_res.drop(columns=['Unnamed: 0'], inplace=True)
columns = ['C', 'f1', 'features', 'gamma', 'kappa',
'matthews', 'roc_auc', 'stock',
'test_f1', 'test_kappa', 'test_matthews', 'test_roc_auc', 'r', 's']
df_res[columns].sort_values(by='matthews', ascending=False).groupby('stock').head(1)
log_res = []
for stock in stocks:
log_res.append(get_logistic_regression(stock, data_length))
df_log_res = pd.DataFrame(log_res)
df_log_res['stock'] = df_log_res['stock'].values.astype(np.int)
df_log_res.index = df_log_res['stock'].values.astype(np.int)
df_gdf_best = df_res[columns].sort_values(by='matthews', ascending=False).groupby('stock').head(1)
df_gdf_best['stock'] = df_gdf_best['stock'].values.astype(np.int)
df_gdf_best.index = df_gdf_best['stock'].values.astype(np.int)
df_all = pd.merge(df_gdf_best, df_log_res, on='stock', suffixes=['_svm', '_log'])
all_columns = ['matthews_svm', 'matthews_log', 'test_matthews_svm', 'test_matthews_log',
'roc_auc_svm', 'roc_auc_log', 'test_roc_auc_svm', 'test_roc_auc_log', 'stock' ]
df_all[all_columns]
df_all['matthews_diff'] = df_all['matthews_svm'] - df_all['matthews_log']
df_all['matthews_test_diff'] = df_all['test_matthews_svm'] - df_all['test_matthews_log']
sns.distplot(df_all['matthews_diff'], label='training')
sns.distplot(df_all['matthews_test_diff'], label='testing')
plt.legend()
sns.distplot(df_all['matthews_svm'], label='svm')
sns.distplot(df_all['matthews_log'], label='log')
plt.legend()
df_all['matthews_diff'].sum(), df_all['matthews_test_diff'].sum()
len(df_all[df_all['matthews_svm'] > df_all['matthews_log']][all_columns]), len(df_all[df_all['test_matthews_svm'] > df_all['test_matthews_log']]), len(df_all)
df_all[df_all['test_matthews_svm'] < df_all['test_matthews_log']][all_columns]
len(df_all[df_all['roc_auc_svm'] > df_all['roc_auc_log']][all_columns]), len(df_all[df_all['test_roc_auc_svm'] > df_all['test_roc_auc_log']][all_columns]), len(df_all)
df_all[df_all['test_matthews_svm'] < df_all['test_matthews_log']][all_columns]
df_all[df_all['test_roc_auc_svm'] < df_all['test_roc_auc_log']][all_columns]
| gdf_pca/junk/results-pca_gdf_20-30_que3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kathleenmei/CPEN-21A-ECE-2-1/blob/main/Lab1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="sq5k8ORl2LCD"
# #Laboratory 1
# + colab={"base_uri": "https://localhost:8080/"} id="33CwuNgp2QR-" outputId="a0466914-48f3-4281-a486-ebf2d21af298"
a = "Welcome to Python Programming"
x = "Name:"
y = "Address:"
z = "Age:"
print(a)
print(x +" " "<NAME>")
print(y +" " "#37 Bliss, Biga II, Silang, Cavite")
print(z +" " "19 years old")
| Lab1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.2
# language: julia
# name: julia-1.4
# ---
# # State-of-the-art model composition in MLJ (Machine Learning in Julia)
# In this script we use [model
# stacking](https://alan-turing-institute.github.io/DataScienceTutorials.jl/getting-started/stacking/)
# to demonstrate the ease with which machine learning models can be
# combined in sophisticated ways using MLJ. In the future MLJ will
# have a canned version of stacking. For now we show how to stack
# using MLJ's generic model composition syntax, which is an extension
# of the normal fit/predict syntax.
DIR = @__DIR__
include(joinpath(DIR, "setup.jl"))
# ## Stacking is hard
# [Model
# stacking](https://alan-turing-institute.github.io/DataScienceTutorials.jl/getting-started/stacking/),
# popular in Kaggle data science competitions, is a sophisticated way
# to blend the predictions of multiple models.
# With the python toolbox
# [scikit-learn](https://scikit-learn.org/stable/) (or its [julia
# wrap](https://github.com/cstjean/ScikitLearn.jl)) you can use
# pipelines to combine composite models in simple ways but (automated)
# stacking is beyond its capabilities.
# One python alternative is to use
# [vecstack](https://github.com/vecxoz/vecstack). The [core
# algorithm](https://github.com/vecxoz/vecstack/blob/master/vecstack/core.py)
# is about eight pages (without the scikit-learn interface):
# .
# ## Stacking is easy (in MLJ)
# Using MLJ's [generic model composition
# API](https://alan-turing-institute.github.io/MLJ.jl/dev/composing_models/)
# you can build a stack in about a page.
# Here's the complete code needed to define a new model type that
# stacks two base regressors and one adjudicator in MLJ. Here we use
# three folds to create the base-learner [out-of-sample
# predictions](https://alan-turing-institute.github.io/DataScienceTutorials.jl/getting-started/stacking/)
# to make it easier to read. You can make this generic with little fuss.
# +
using MLJ
folds(data, nfolds) =
partition(1:nrows(data), (1/nfolds for i in 1:(nfolds-1))...);
model1 = @load LinearRegressor pkg=MLJLinearModels
model2 = @load LinearRegressor pkg=MLJLinearModels
judge = @load LinearRegressor pkg=MLJLinearModels
X = source()
y = source()
folds(X::AbstractNode, nfolds) = node(XX->folds(XX, nfolds), X)
MLJ.restrict(X::AbstractNode, f::AbstractNode, i) =
node((XX, ff) -> restrict(XX, ff, i), X, f);
MLJ.corestrict(X::AbstractNode, f::AbstractNode, i) =
node((XX, ff) -> corestrict(XX, ff, i), X, f);
f = folds(X, 3)
m11 = machine(model1, corestrict(X, f, 1), corestrict(y, f, 1))
m12 = machine(model1, corestrict(X, f, 2), corestrict(y, f, 2))
m13 = machine(model1, corestrict(X, f, 3), corestrict(y, f, 3))
y11 = predict(m11, restrict(X, f, 1));
y12 = predict(m12, restrict(X, f, 2));
y13 = predict(m13, restrict(X, f, 3));
m21 = machine(model2, corestrict(X, f, 1), corestrict(y, f, 1))
m22 = machine(model2, corestrict(X, f, 2), corestrict(y, f, 2))
m23 = machine(model2, corestrict(X, f, 3), corestrict(y, f, 3))
y21 = predict(m21, restrict(X, f, 1));
y22 = predict(m22, restrict(X, f, 2));
y23 = predict(m23, restrict(X, f, 3));
y1_oos = vcat(y11, y12, y13);
y2_oos = vcat(y21, y22, y23);
X_oos = MLJ.table(hcat(y1_oos, y2_oos))
m_judge = machine(judge, X_oos, y)
m1 = machine(model1, X, y)
m2 = machine(model2, X, y)
y1 = predict(m1, X);
y2 = predict(m2, X);
X_judge = MLJ.table(hcat(y1, y2))
yhat = predict(m_judge, X_judge)
@from_network machine(Deterministic(), X, y; predict=yhat) begin
mutable struct MyStack
regressor1=model1
regressor2=model2
judge=judge
end
end
my_stack = MyStack()
# -
# For the curious: Only the last block defines the new model type. The
# rest defines a *[learning network]()* - a kind of working prototype
# or blueprint for the type. If the source nodes `X` and `y` wrap some
# data (instead of nothing) then the network can be trained and tested
# as you build it.
# ## Composition plays well with other work-flows
# We did not include standardization of inputs and target (with
# post-prediction inversion) in our stack. However, we can add these
# now, using MLJ's canned pipeline composition:
pipe = @pipeline Standardizer my_stack target=Standardizer
# Want to change a base learner and adjudicator?
pipe.my_stack.regressor2 = @load DecisionTreeRegressor pkg=DecisionTree
pipe.my_stack.judge = @load KNNRegressor
# Want a CV estimate of performance of the complete model on some data?
X, y = @load_boston;
mach = machine(pipe, X, y)
evaluate!(mach, resampling=CV(), measure=[mae, rms])
# Want to inspect the learned parameters of the adjudicator?
fp = fitted_params(mach);
fp.my_stack.judge
# What about the first base-learner of the stack? There are four sets
# of learned parameters! One for each fold to make an out-of-sample
# prediction, and one trained on all the data:
fp.my_stack.regressor1
fp.my_stack.regressor1[1].coefs
# Want to tune multiple (nested) hyperparameters in the stack? Tuning is a
# model wrapper (for better composition!):
# +
r1 = range(pipe, :(my_stack.regressor2.max_depth), lower = 1, upper = 25)
r2 = range(pipe, :(my_stack.judge.K), lower=1, origin=10, unit=10)
import Distributions.Poisson
tuned_pipe = TunedModel(model=pipe,
ranges=[r1, (r2, Poisson)],
tuning=RandomSearch(),
resampling=CV(),
measure=rms,
n=100)
mach = machine(tuned_pipe, X, y) |> fit!
best_model = fitted_params(mach).best_model
K = fitted_params(mach).best_model.my_stack.judge.K;
max_depth = fitted_params(mach).best_model.my_stack.regressor2.max_depth
@show K max_depth;
# -
# Visualize tuning results:
using Plots
pyplot()
plot(mach)
# ---
#
# *This notebook was generated using [Literate.jl](https://github.com/fredrikekre/Literate.jl).*
| wow.ipynb |