code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/XavierCarrera/neural-network/blob/main/Neural_Network_Structure.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="JsSCGrCPZAAe"
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.datasets import load_iris
from sklearn.linear_model import Perceptron
# + id="vcaFCEdg5N8G" outputId="289bd812-881c-480e-b1df-8d8cb7062b4d" colab={"base_uri": "https://localhost:8080/", "height": 136}
iris = load_iris()
iris.target
# + id="vd_y1xsW5Uys" outputId="84f8c0b4-d480-47dd-b16f-5f3b07fdaaf9" colab={"base_uri": "https://localhost:8080/", "height": 102}
iris.data[:5, :]
# + id="HVyTkCPR6ujA" outputId="c4af2925-bd51-423c-ddd9-400d4e501767" colab={"base_uri": "https://localhost:8080/", "height": 388}
data = iris.data[:, (2, 3)]
labels = iris.target
plt.figure(figsize=(13,6))
plt.scatter(data[:, 0], data[:, 1], c=labels, cmap=plt.cm.Set1, edgecolor='face')
plt.xlabel('Petal Length (cm)')
plt.ylabel('Petal Width (cm)')
plt.show()
# + id="BOvElfZz7E4v" outputId="2e45dff8-68d9-4e63-bb35-a779e0d5c345" colab={"base_uri": "https://localhost:8080/", "height": 85}
X = iris.data[:, (2, 3)]
y = (iris.target == 2).astype(np.int)
test_perceptron = Perceptron()
test_perceptron.fit(X, y)
# + id="8YIq74LX7IOg" outputId="8bab46ad-46dd-4274-b6fc-e0e26830e010" colab={"base_uri": "https://localhost:8080/", "height": 34}
y1_pred = test_perceptron.predict([[5.1, 2]])
y2_pred = test_perceptron.predict([[1.4, 0.2]])
print('Prediction 1:', y1_pred, 'Prediction 2:', y2_pred)
| Neural_Network_Structure.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="e4cnBhHe-dlE"
# # Regression
#
# So far, in our exploration of machine learning, we have build sysems that predict discrete values: mountain bike or not mountain bike, democrat or republican. And not just binary choices, but, for example, deciding whether an image is of a particular digit:
#
# 
#
# or which of 1,000 categories does a picture represent.
#
#
# This lab looks at how we can build classifiers that predict **continuous** values and such classifiers are called regression classifiers.
#
# First, let's take a small detour into correlation.
#
# ## Correlation
# A correlation is the degree of association between two variables. One of my favorite books on this topic is:
#
# <img src="http://zacharski.org/files/courses/cs419/mg_statistics_big.png" width="250" />
#
#
# and they illustrate it by looking at
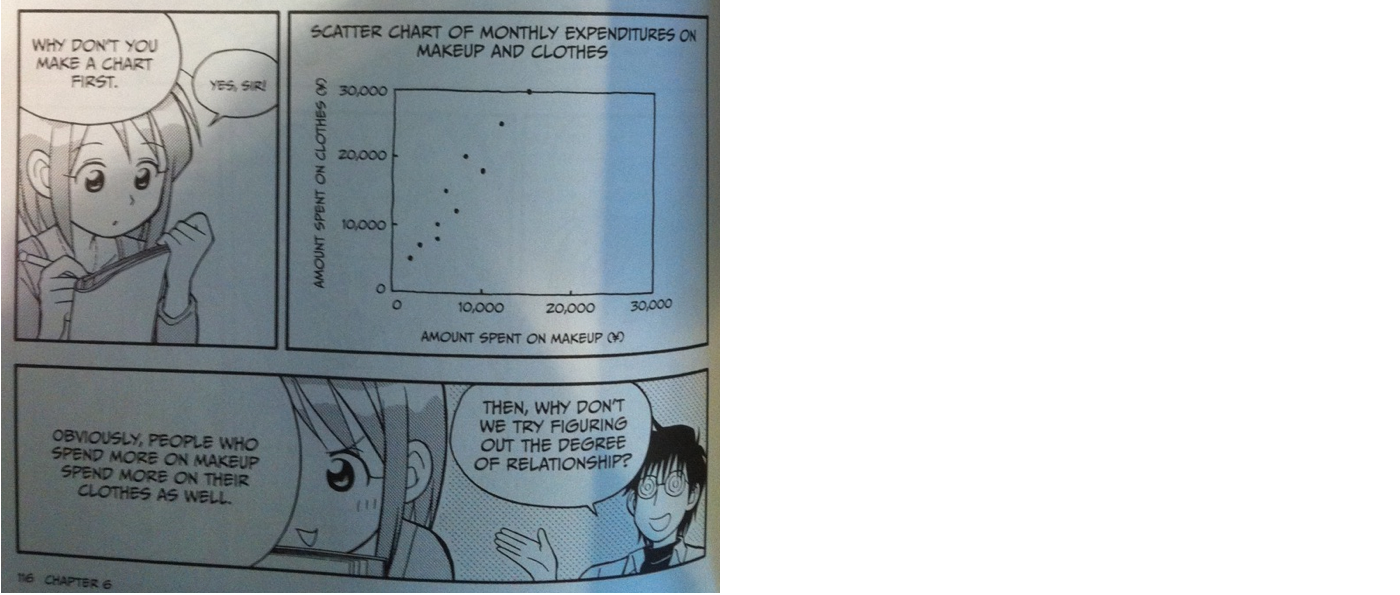
# ## Ladies expenditures on clothes and makeup
# 
# 
#
#
#
# So let's go ahead and create that data in Pandas and show the table:
# + colab={"base_uri": "https://localhost:8080/", "height": 357} colab_type="code" id="bqpDwvW3-dlF" outputId="971eb9f5-576f-41b4-c905-427b7bea3a0b"
import pandas as pd
from pandas import DataFrame
makeup = [3000, 5000, 12000, 2000, 7000, 15000, 5000, 6000, 8000, 10000]
clothes = [7000, 8000, 25000, 5000, 12000, 30000, 10000, 15000, 20000, 18000]
ladies = ['Ms A','Ms B','Ms C','Ms D','Ms E','Ms F','Ms G','Ms H','Ms I','Ms J',]
monthly = DataFrame({'makeup': makeup, 'clothes': clothes}, index= ladies)
monthly
# + [markdown] colab_type="text" id="_Yyce16P-dlJ"
# and let's show the scatterplot
# + colab={"base_uri": "https://localhost:8080/", "height": 617} colab_type="code" id="f5QIDuDg-dlJ" outputId="d300ceec-3855-4dc2-dfd2-8cb9e9c6210f"
from bokeh.plotting import figure, output_file, show
from bokeh.io import push_notebook, show, output_notebook
output_notebook()
x = figure(title="Montly Expenditures on Makeup and Clothes", x_axis_label="Money spent on makeup", y_axis_label="Money spent on clothes")
x.circle(monthly['makeup'], monthly['clothes'], size=10, color="navy", alpha=0.5)
output_file("stuff.html")
show(x)
# + [markdown] colab_type="text" id="qBH5fNxT-dlN"
# When the data points are close to a straight line going up, we say that there is a positive correlation between the two variables. So in the case of the plot above, it visually looks like a postive correlation. Let's look at a few more examples:
#
# ## Weight and calories consumed in 1-3 yr/old children
# This small but real dataset examines whether young children who weigh more, consume more calories
#
# + colab={"base_uri": "https://localhost:8080/", "height": 542} colab_type="code" id="bscoNATT-dlN" outputId="b6f143db-e830-41b4-98e0-78a785237335"
weight = [7.7, 7.8, 8.6, 8.5, 8.6, 9, 10.1, 11.5, 11, 10.2, 11.9, 10.4, 9.3, 9.1, 8.5, 11]
calories = [360, 400, 500, 370, 525, 800, 900, 1200, 1000, 1400, 1600, 850, 575, 425, 950, 800]
kids = DataFrame({'weight': weight, 'calories': calories})
kids
# + colab={"base_uri": "https://localhost:8080/", "height": 617} colab_type="code" id="Ngx0m8sN-dlT" outputId="d5a44bfc-0e0d-4bc2-ee79-02dc9b049cc6"
p = figure(title="Weight and calories in 1-3 yr.old children",
x_axis_label="weight (kg)", y_axis_label='weekly calories')
p.circle(kids['weight'], kids['calories'], size=10, color='navy', alpha=0.5)
show(p)
# + [markdown] colab_type="text" id="JfU5QT1G-dlW"
# And again, there appears to be a positive correlation.
#
# ## The stronger the correlation the closer to a straight line
# The closer the data points are to a straight line, the higher the correlation. A rising straight line (rising going left to right) would be perfect positive correlation. Here we are comparing the heights in inches of some NHL players with their heights in cm. Obviously, those are perfectly correlated.
# + colab={"base_uri": "https://localhost:8080/", "height": 234} colab_type="code" id="SJJ5k9Iu-dlX" outputId="06d2fea2-8efd-40e8-e313-f0ca88dc2877"
inches =[68, 73, 69,72,71,77]
cm = [173, 185, 175, 183, 180, 196]
nhlHeights = DataFrame({'heightInches': inches, 'heightCM': cm})
nhlHeights
# + colab={"base_uri": "https://localhost:8080/", "height": 617} colab_type="code" id="MWppQKt4-dla" outputId="8af36f32-ab60-447d-f1f1-bf3f1219a4d6"
p = figure(title="Comparison of Height in Inches and Height in CM",
x_axis_label="Height in Inches",
y_axis_label="Height in centimeters")
p.circle(nhlHeights['heightInches'], nhlHeights['heightCM'],
size=10, color='navy', alpha=0.5)
show(p)
# + [markdown] colab_type="text" id="h0vwF4yr-dld"
# ## No correlation = far from straight line
# On the opposite extreme, if the datapoints are scattered and no line is discernable, there is no correlation.
#
# Here we are comparing length of the player's hometown name to his height in inches. We are checking whether a player whose hometown name has more letters, tends to be taller. For example, maybe someone from Medicine Hat is taller than someone from Ledue. Obviously there should be no correlation.
#
#
# (Again, a small but real dataset)
# + colab={"base_uri": "https://localhost:8080/", "height": 820} colab_type="code" id="YYa58G0_-dle" outputId="5578cac4-92f3-41fa-a413-5b15a03196e1"
medicineHat = pd.read_csv('https://raw.githubusercontent.com/zacharski/machine-learning-notebooks/master/data/medicineHatTigers.csv')
medicineHat['hometownLength'] = medicineHat['Hometown'].str.len()
medicineHat
# + colab={"base_uri": "https://localhost:8080/", "height": 617} colab_type="code" id="duMhFyDu-dlg" outputId="83206323-deda-4c80-ebbe-15acdd75e498"
p = figure(title="Correlation of the number of Letters in the Hometown to Height",
x_axis_label="Player's Height", y_axis_label="Hometown Name Length")
p.circle(medicineHat['Height'], medicineHat['hometownLength'], size=10, color='navy', alpha=0.5)
show(p)
# + [markdown] colab_type="text" id="iMDBBcyM-dlj"
# And that does not look at all like a straight line.
#
# ## negative correlation has a line going downhill
# When the slope goes up, we say there is a positive correlation and when it goes down there is a negative correlation.
#
# #### the relationship of hair length to a person's height
# + colab={"base_uri": "https://localhost:8080/", "height": 617} colab_type="code" id="fF6MndKp-dlj" outputId="68446cbe-0e4e-47b0-8787-6caa51a8feff"
height =[62, 64, 65, 68, 69, 70, 67, 65, 72, 73, 74]
hairLength = [7, 10, 6, 4, 5, 4, 5, 8, 1, 1, 3]
cm = [173, 185, 175, 183, 180, 196]
people = DataFrame({'height': height, 'hairLength': hairLength})
p = figure(title="Correlation of hair length to a person's height",
x_axis_label="Person's Height", y_axis_label="Hair Length")
p.circle(people['height'], people['hairLength'], size=10, color='navy', alpha=0.5)
show(p)
# + [markdown] colab_type="text" id="VdRZxka9-dlm"
# There is a strong negative correlation between the length of someone's hair and how tall they are. That makes some sense. I am bald and 6'0" and my friend Sara is 5'8" and has long hair.
#
# # Numeric Represenstation of the Strength of the Correlation
#
# So far, we've seen a visual representation of the correlation, but we can also represent the degree of correlation numerically.
#
# ## Pearson Correlation Coefficient
#
# This ranges from -1 to 1.
# 1 is perfect positive correlation, -1 is perfect negative.
#
# $$r=\frac{\sum_{i=1}^n(x_i - \bar{x})(y_i-\bar{y})}{\sqrt{\sum_{i=1}^n(x_i - \bar{x})} \sqrt{\sum_{i=1}^n(y_i - \bar{y})}}$$
#
# In Pandas it is very easy to compute.
#
# ### Japanese ladies expensives on makeup and clothes
# Let's go back to our first example.
# First here is the data:
# + colab={"base_uri": "https://localhost:8080/", "height": 357} colab_type="code" id="sIowAZ8L-dln" outputId="391b27d1-6b60-4717-8169-e3d7613d8433"
monthly
# + colab={"base_uri": "https://localhost:8080/", "height": 617} colab_type="code" id="3t5jGS5f-dlp" outputId="b5567394-7ce8-49ba-8d3e-38ad1f01c574"
p = figure(title="Montly Expenditures on Makeup and Clothes",
x_axis_label="Money spent on makeup", y_axis_label="Money spent on clothes")
p.circle(monthly['makeup'], monthly['clothes'], size=10, color='navy', alpha=0.5)
show(p)
# + [markdown] colab_type="text" id="oxUIU2lE-dls"
# So that looks like a pretty strong positive correlation. To compute Pearson on this data we do:
# + colab={"base_uri": "https://localhost:8080/", "height": 110} colab_type="code" id="WvYUkvHD-dls" outputId="49d7af89-1bb2-4260-d847-f0d014545043"
monthly.corr()
# + [markdown] colab_type="text" id="3cdda0-9-dlv"
# There is no surprise that makeup is perfectly correlated with makeup and clothes with clothes (those are the 1.000 on the diagonal). The interesting bit is that the Pearson for makeup to clothes is 0.968. That is a pretty strong correlation.
#
# If you are interesting you can compute the Pearson values for the datasets above, but let's now move to ...
#
# #### Regression
#
# Let's say we know a young lady who spends about ¥22,500 per month on clothes (that's about $200/month). What do you think she spends on makeup, based on the chart below?
# + colab={"base_uri": "https://localhost:8080/", "height": 617} colab_type="code" id="tGwP03jC-dlw" outputId="98e981e4-4d9e-4fe0-c968-eef70d962354"
show(p)
# + [markdown] colab_type="text" id="l89HC2LH-dlz"
# I'm guessing you would predict she spends somewhere around ¥10,000 a month on makeup (almost $100/month). And how we do this is when looking at the graph we mentally draw an imaginary straight line through the datapoints and use that line for predictions. We are performing human linear regression. And as humans, we have the training set--the dots representing data points on the graph. and we **fit** our human classifier by mentally drawing that straight line. That straight line is our model. Once we have it, we can throw away the data points. When we want to make a prediction, we see where the money spent on clothes falls on that line.
#
# We just predicted a continuous value (money spent on makeup) from one factor (money spent on clothes).
#
# What happens when we want to predict a continuous value from 2 factors? Suppose we want to predict MPG based on the weight of a car and its horsepower.
#
# 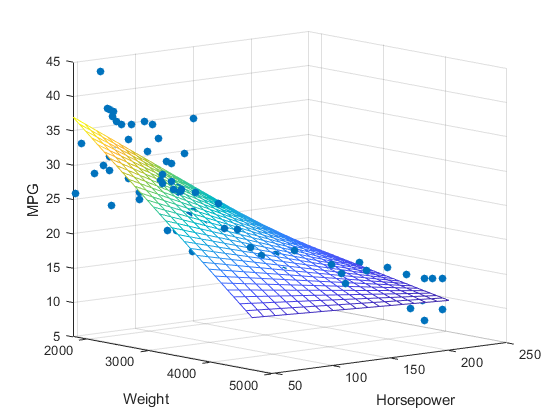
# from [Mathworks](https://www.mathworks.com/help/stats/regress.html)
#
# Now instead of a line representing the relationship we have a plane.
#
# Let's create a linear regression classifier and try this out!
#
# First, let's get the data.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 448} colab_type="code" id="nS-VZrlZ-dlz" outputId="2577e13e-f5c0-48c2-d164-860c8df5a8ab"
columnNames = ['mpg', 'cylinders', 'displacement', 'HP', 'weight', 'acceleration', 'year', 'origin', 'model']
cars = pd.read_csv('https://raw.githubusercontent.com/zacharski/ml-class/master/data/auto-mpg.csv',
na_values=['?'], names=columnNames)
cars = cars.set_index('model')
cars = cars.dropna()
cars
# + [markdown] colab_type="text" id="E0rh87Zf-dl2"
# Now divide the dataset into training and testing. And let's only use the horsepower and weight columns as features.
# + colab={"base_uri": "https://localhost:8080/", "height": 448} colab_type="code" id="R_JBLlrW-dl2" outputId="4355bca9-9a75-43b8-ae02-f552c2241b85"
from sklearn.model_selection import train_test_split
cars_train, cars_test = train_test_split(cars, test_size = 0.2)
cars_train_features = cars_train[['HP', 'weight']]
# cars_train_features['HP'] = cars_train_features.HP.astype(float)
cars_train_labels = cars_train['mpg']
cars_test_features = cars_test[['HP', 'weight']]
# cars_test_features['HP'] = cars_test_features.HP.astype(float)
cars_test_labels = cars_test['mpg']
cars_test_features
# + [markdown] colab_type="text" id="f62sESxd-dl6"
# ### SKLearn Linear Regression
#
# Now let's create a Linear Regression classifier and fit it.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="wJ5-BzSU-dl6" outputId="8adb5159-d67c-4956-90cf-ade0fbfeb089"
from sklearn.linear_model import LinearRegression
linclf = LinearRegression()
linclf.fit(cars_train_features, cars_train_labels)
# + [markdown] colab_type="text" id="qo_VNdhI-dl9"
# and finally use the trained classifier to make predictions on our test data
# + colab={} colab_type="code" id="gtNyLgia-dl9"
predictions = linclf.predict(cars_test_features)
# + [markdown] colab_type="text" id="jYsJvAiR-dl_"
# Let's take an informal look at how we did:
# + colab={"base_uri": "https://localhost:8080/", "height": 448} colab_type="code" id="9AnurGcX-dmC" outputId="30e7590c-dc59-48c6-f257-aace2dd5ae38"
results = cars_test_labels.to_frame()
results['Predicted']= predictions
results
# + [markdown] colab_type="text" id="rjOo1BLN-dmF"
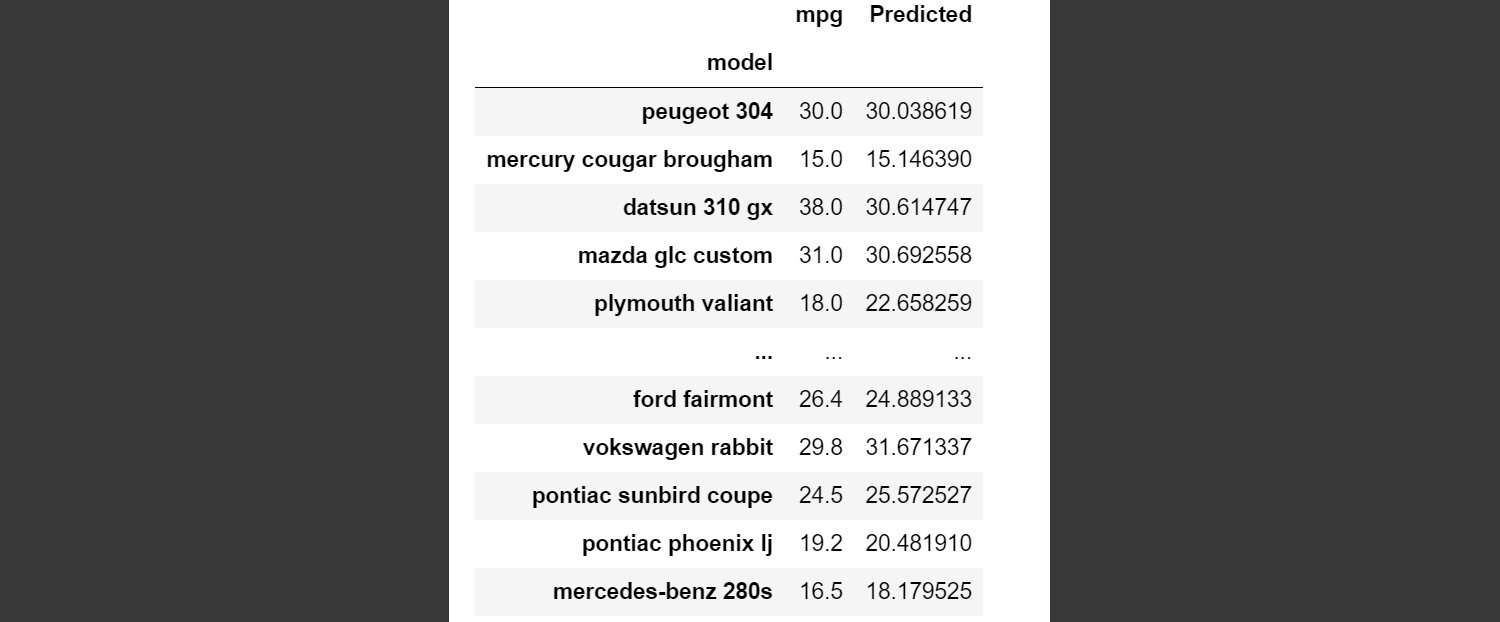
# Here is what my output looked like:
#
# 
#
# as you can see the first two predictions were pretty close as were a few others.
#
# ### Determining how well the classifier performed
#
# With categorical classifiers we used sklearn's accuracy_score:
#
# ```
# from sklearn.metrics import accuracy_score
# ```
# Consider a task of predicting whether an image is of a dog or a cat. We have 10 instances in our test set. After our classifier makes predictions, for each image we have the actual (true) value, and the value our classifier predicted:
#
# actual | predicted
# :-- | :---
# dog | dog
# **dog** | **cat**
# # # cat | cat
# dog | dog
# # # cat | cat
# **cat** | **dog**
# dog | dog
# # # cat | cat
# # # cat | cat
# dog | dog
#
# sklearn's accuracy score just counts how many predicted values matched the actual values and then divides by the total number of test instances. In this case the accuracy score would be .8000. The classifier was correct 80% of the time.
#
# We can't use this method with a regression classifier. In the image above, the actual MPG of the Peugeot 304 was 30 and our classifier predicted 30.038619. Does that count as a match or not? The actual mpg of a Pontiac Sunbird Coupe was 24.5 and we predicted 25.57. Does that count as a match? Instead of accuracy_score, there are different evaluation metrics we can use.
#
# #### Mean Squared Error and Root Mean Square Error
#
# A common metric is Mean Squared Error or MSE. MSE is a measure of the quality of a regression classifier. The closer MSE is to zero, the better the classifier. Let's look at some made up data to see how this works:
#
# vehicle | Actual MPG | Predicted MPG
# :---: | ---: | ---:
# Ram Promaster 3500 | 18.0 | 20.0
# Ford F150 | 20 | 19
# Fiat 128 | 33 | 33
#
# First we compute the error (the difference between the predicted and actual values)
#
# vehicle | Actual MPG | Predicted MPG | Error
# :---: | ---: | ---: | --:
# Ram Promaster 3500 | 18.0 | 20.0 | -2
# Ford F150 | 20 | 19 | 1
# Fiat 128 | 33 | 33 | 0
#
# Next we square the error and compute the average:
#
# vehicle | Actual MPG | Predicted MPG | Error | Error^2
# :---: | ---: | ---: | --: | ---:
# Ram Promaster 3500 | 18.0 | 20.0 | -2 | 4
# Ford F150 | 20 | 19 | 1 | 1
# Fiat 128 | 33 | 33 | 0 | 0
# MSE | - | - | - | 1.667
#
# **Root Mean Squared Error** is simply the square root of MSE. The advantage of RMSE is that it has the same units as what we are trying to predict. Let's take a look ...
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="z8DX6nRJ-dmF" outputId="829b2878-924d-4ad7-9642-4018bfba4234"
from sklearn.metrics import mean_squared_error
MSE = mean_squared_error(cars_test_labels, predictions)
RMSE = mean_squared_error(cars_test_labels, predictions, squared=False)
print("MSE: %5.3f. RMSE: %5.3f" %(MSE, RMSE))
# + [markdown] colab_type="text" id="W9ZgOu53-dmH"
# That RMSE tells us on average how many mpg we were off.
#
# ---
#
#
#
# ## So what kind of model does a linear regression classifier build?
#
# You probably know this if you reflect on grade school math classes you took.
#
# Let's go back and look at the young ladies expenditures on clothes and makeup.
# + colab={"base_uri": "https://localhost:8080/", "height": 617} colab_type="code" id="hpZ8csEm-dmI" outputId="ffa3271d-10a2-4e01-b7a0-2cf1ffa93ede"
p = figure(title="Montly Expenditures on Makeup and Clothes",
x_axis_label="Money spent on makeup", y_axis_label="Money spent on clothes")
p.circle(monthly['makeup'], monthly['clothes'], size=10, color='navy', alpha=0.5)
show(p)
# + [markdown] colab_type="text" id="pd0Kp1b0-dmL"
# When we talked about this example above, I mentioned that when we do this, we imagine a line. Let's see if we can use sklearns linear regression classifier to draw that line:
# + colab={"base_uri": "https://localhost:8080/", "height": 252} colab_type="code" id="iUPQmLI9-dmL" outputId="93f99ab9-eac7-4757-9d55-a707159639df"
regr = LinearRegression()
regr.fit(monthly[['clothes']], monthly['makeup'])
pred = regr.predict(monthly[['clothes']])
import matplotlib.pyplot as plt
# Plot outputs
plt.scatter(monthly['clothes'], monthly['makeup'], color='black')
plt.plot(monthly['clothes'], pred, color='blue', linewidth=3)
plt.xticks(())
plt.yticks(())
plt.show()
# + [markdown] colab_type="text" id="_lwOOz55-dmO"
# Hopefully that matches your imaginary line!
#
# The formula for the line is
#
# $$makeup=w_0clothes + y.intercept$$
#
# We can query our classifier for those values ($w_0$, and $i.intercept$):
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="_Qu6YBQa-dmO" outputId="5622fea3-934c-40e0-bf2d-867ec0ff3a9e"
print('w0 = %5.3f' % regr.coef_)
print('y intercept = %5.3f' % regr.intercept_)
# + [markdown] colab_type="text" id="4W7hfaH1-dmR"
# So the formula for this particular example is
#
# $$ makeup = 0.479 * clothes + 121.782$$
#
# So if a young lady spent ¥22,500 on clothes we would predict she spent the following on makeup:
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="o8H6PpL3-dmR" outputId="182a26ba-a23f-40fb-a608-f2538cf1768e"
makeup = regr.coef_[0] * 22500 + regr.intercept_
makeup
# + [markdown] colab_type="text" id="CWU_PMNW-dmT"
# The formula for regression in general is
#
# $$\hat{y}=\theta_0 + \theta_1x_1 + \theta_2x_2 + ... \theta_nx_n$$
#
# where $\theta_0$ is the y intercept. When you fit your classifier it is learning all those $\theta$'s. That is the model your classifier learns.
#
# It is important to understand this as it applies to other classifiers as well!
#
#
# ## Overfitting
# Consider two models for our makeup predictor. One is the straight line:
# + colab={"base_uri": "https://localhost:8080/", "height": 252} colab_type="code" id="QDX1lstU-dmU" outputId="b66454ce-2571-46af-b39f-2e48089e475c"
plt.scatter(monthly['clothes'], monthly['makeup'], color='black')
plt.plot(monthly['clothes'], pred, color='blue', linewidth=3)
plt.xticks(())
plt.yticks(())
plt.show()
# + [markdown] colab_type="text" id="K9pH023O-dmW"
# And the other looks like:
# + colab={"base_uri": "https://localhost:8080/", "height": 252} colab_type="code" id="G6yZNWN--dmX" outputId="af5531d3-cc89-4eed-8c77-48fb5b359d43"
monthly2 = monthly.sort_values(by='clothes')
plt.scatter(monthly2['clothes'], monthly2['makeup'], color='black')
plt.plot(monthly2['clothes'], monthly2['makeup'], color='blue', linewidth=3)
plt.xticks(())
plt.yticks(())
plt.show()
# + [markdown] colab_type="text" id="txLtJJZc-dma"
# The second model fits the training data perfectly. Is it better than the first? Here is what could happen.
#
# Let's say we have been tuning our model using our validation data set. Our error rates look like
#
# 
#
# As you can see our training error rate keeps going down, but at the very end our validation error increases. This is called **overfitting** the data. The model is highly tuned to the nuances of the training data. So much so, that it hurts the performance on new data--in this case, the validation data. This, obviously, is not a good thing.
#
# ---
# #### An aside
#
# Imagine preparing for a job interview for a position you really, really, want. Since we are working on machine learning, let's say it is a machine learning job. In their job ad they list a number of things they want the candidate to know:
#
# * Convolutional Neural Networks
# * Long Short Term Memory models
# * Recurrent Neural Networks
# * Generative Deep Learning
#
# And you spend all your waking hours laser focused on these topics. You barely get any sleep and you read articles on these topics while you eat. You know the tiniest intricacies of these topics. You are more than 100% ready.
#
# The day of the interview arrives. After of easy morning of chatting with various people, you are now in a conference room for the technical interview, standing at a whiteboard, ready to hopefully wow them with your wisdom. The first question they ask is for you to write the solution to the fizz buzz problem:
#
# > Write a program that prints the numbers from 1 to 100. But for multiples of three print “Fizz” instead of the number and for the multiples of five print “Buzz”. For numbers which are multiples of both three and five print “FizzBuzz”
#
# And you freeze. This is a simple request and a very common interview question. In fact, to prepare you for this job interview possibility, write it now:
#
#
#
#
# + colab={} colab_type="code" id="UMdU0j3ALXgU"
def fizzbuzz():
print("TO DO")
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="bDG7M1WGLgEj" outputId="1017545f-4b24-4bd6-f0b7-0b6b1011b7a4"
fizzbuzz()
# + [markdown] colab_type="text" id="k7Zt-eCZLgZe"
# Back to the job candidate freezing, this is an example of overfitting. You overfitting to the skills mentioned in the job posting.
#
# At dissertation defenses often faculty will ask the candidate questions outside of the candidate's dissertation. I heard of one case in a physics PhD defense where a faculty member asked "Why is the sky blue?" and the candidate couldn't answer.
#
# Anyway, back to machine learning.
#
# ---
#
#
# There are a number of ways to reduce the likelihood of overfitting including
#
# * We can reduce the complexity of the model. Instead of going with the model of the jagged line immediately above we can go with the simpler straight line model. We have seen this in decision trees where we limit the depth of the tree.
#
# * Another method is to increase the amount of training data.
#
# Let's examine the first. The process of reducing the complexity of a model is called regularization.
#
# The linear regression model we have just used tends to overfit the data and there are some variants that are better and these are called regularized linear models. These include
#
# * Ridge Regression
# * Lasso Regression
# * Elastic Net - a combination of Ridge and Lasso
#
# Let's explore Elastic Net. And let's use all the columns of the car mpg dataset:
# + colab={} colab_type="code" id="3nxItuCZ-dma"
newTrain_features = cars_train.drop('mpg', axis=1)
newTrain_labels = cars_train['mpg']
newTest_features = cars_test.drop('mpg', axis=1)
newTest_labels = cars_test['mpg']
# + [markdown] colab_type="text" id="t6GUbt83-dmc"
# First, let's try with our standard Linear Regression classifier:
# + colab={} colab_type="code" id="EwsP5LBo-dmd"
linclf = LinearRegression()
linclf.fit(newTrain_features, newTrain_labels)
predictions = linclf.predict(newTest_features)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="3iFVEwh_-dmf" outputId="61f6547e-0152-4d4f-bcb8-041593b01e4b"
MSE = mean_squared_error(newTest_labels, predictions)
RMSE = mean_squared_error(newTest_labels, predictions, squared=False)
print("MSE: %5.3f. RMSE: %5.3f" %(MSE, RMSE))
# + [markdown] colab_type="text" id="B7lJGcXQN1iI"
# Now let's try with [ElasticNet](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.ElasticNet.html)
# + colab={} colab_type="code" id="A0eTqpNS-dmk"
from sklearn.linear_model import ElasticNet
elastic_net = ElasticNet(alpha=0.1, l1_ratio=0.5)
elastic_net.fit(newTrain_features, newTrain_labels)
ePredictions = elastic_net.predict(newTest_features)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="PFjWu3Nx-dmm" outputId="ebb5f7d1-c936-469f-acd6-b298451bbe6c"
MSE = mean_squared_error(newTest_labels, ePredictions)
RMSE = mean_squared_error(newTest_labels, ePredictions, squared=False)
print("MSE: %5.3f. RMSE: %5.3f" %(MSE, RMSE))
# + [markdown] colab_type="text" id="oasWKk30OCui"
# I've run this a number of times. Sometimes linear regression is slightly better and sometimes ElasticNet is. Here are the results of one run:
#
# ##### RMSE
#
# Linear Regression | Elastic Net
# :---: | :---:
# 2.864 | 2.812
#
# So this is not the most convincing example.
#
# However, in general, it is always better to have some regularization, so (mostly) you should avoid the generic linear regression classifier.
#
# ## Happiness
# What better way to explore regression then to look at happiness.
#
# From a Zen perspective, happiness is being fully present in the current moment.
#
# But, ignoring that advice, let's see if we can predict happiness, or life satisfaction.
#
# 
#
# We are going to be investigating the [Better Life Index](https://stats.oecd.org/index.aspx?DataSetCode=BLI). You can download a csv file of that data from that site.
#
# 
#
# Now that you have the CSV data file on your laptop, you can upload it to Colab.
#
# In Colab, you will see a list of icons on the left.
#
# 
#
# Select the file folder icon.
#
# 
#
# Next, select the upload icon (the page with an arrow icon). And upload the file.
#
# Next, let's execute the Linux command `ls`:
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="HkxsnifT-dmp" outputId="2fa3cf4e-bbcd-4b90-ae87-aef1be18f243"
# !ls
# + [markdown] colab_type="text" id="nzw8N699SrPP"
# ### Load that file into a Pandas DataFrame
#
# We will load the file into Pandas Dataframe called `bli` for better life index:
# + colab={"base_uri": "https://localhost:8080/", "height": 900} colab_type="code" id="3tYTVthdMAPO" outputId="668b07db-8f9c-4e80-838c-556984a27f53"
# TO DO
# + [markdown] colab_type="text" id="R7T8rpt-YS3h"
# When examining the DataFrame we can see it has an interesting structure. So the first row we can parse:
#
# * The country is Australia
# * The feature is Labour market insecurity
# * The Inequality column tells us it is the **total** Labour market insecurity value.
# * The unit column tells the us the number is a percentage.
# * And the value is 5.40
#
# So, in English, the column is The total labor market insecurity for Australia is 5.40%.
#
# I am curious as to what values other than Total are in the Inequality column:
#
#
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="Sdjs0SLKlE5O" outputId="94caa3fe-3a00-413f-effc-4c1d62aff62b"
bli.Inequality.unique()
# + [markdown] colab_type="text" id="b7VnlbDZlJnm"
# Cool. So in addition to the total for each feature, we can get values for just men, just women, and the high and low.
#
# Let's get just the totals and then pivot the DataFrame so it is in a more usable format.
#
# In addition, there are a lot of NaN values in the data, let's replace them with the mean value of the column.
#
# We are just learning about regression and this is a very small dataset, so let's divide training and testing by hand ...
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="EQ2YbF3pMHhn" outputId="c3c5afbf-078c-48c5-a4fc-d4f5339e46f2"
bli = bli[bli["INEQUALITY"]=="TOT"]
bli = bli.pivot(index="Country", columns="Indicator", values="Value")
bli.fillna(bli.mean(), inplace=True)
bliTest = bli.loc['Greece':'Italy', :]
bliTrain = pd.concat([bli.loc[:'Germany' , :], bli.loc['Japan':, :]])
bliTrain
# + [markdown] colab_type="text" id="4uwI17DVr5SM"
# Now we need to divide both the training and test sets into features and labels.
# + colab={} colab_type="code" id="mqP1PbycaXTu"
# TO DO
# + [markdown] colab_type="text" id="GeUlBUDRsHN_"
# ### Create and Train an elastic net model
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="LXMzYK-9moqH" outputId="7ad04a73-02d2-41bd-f80d-166dd5041a99"
# TO DO
# + [markdown] colab_type="text" id="YGBEijsSsOgO"
# ### Use the trained model to make predictions on our tiny test set
# + colab={} colab_type="code" id="Uy79gKKGnhH5"
# TO DO
predictions = 'TODO'
# + [markdown] colab={} colab_type="code" id="Z55ROFZ1tGWf"
# Now let's visually compare the differences between the predictions and the actual values
# + colab={"base_uri": "https://localhost:8080/", "height": 264} colab_type="code" id="eAWEt8rNoj9A" outputId="7f9ee164-c2de-43f2-850b-4e2baf752144"
results = pd.DataFrame(bliTestLabels)
results['Predicted']= predictions
results
# + [markdown] colab_type="text" id="Bu5NtK7ltNrr"
# How did you do? For me Hungary was a lot less happy than what was predicted.
#
#
# # Prediction Housing Prices
#
# ## But first some wonkiness
# When doing one hot encoding, sometimes the original datafile has the same type of data in multiple columns. For example...
#
# Title | Genre 1 | Genre 2
# :--: | :---: | :---:
# Mission: Impossible - Fallout | Action | Drama
# Mama Mia: Here We Go Again | Comedy | Musical
# Ant-Man and The Wasp | Action | Comedy
# BlacKkKlansman | Drama | Comedy
#
#
# When we one-hot encode this we get something like
#
# Title | Genre1 Action | Genre1 Comedy | Genre1 Drama | Genre2 Drama | Genre2 Musical | Genre2 Comedy
# :--: | :--: | :--: | :--: | :--: | :--: | :--:
# Mission: Impossible - Fallout | 1 | 0 | 0 | 1 | 0 | 0
# Mama Mia: Here We Go Again | 0 | 1 | 0 | 0 | 1 | 0
# Ant-Man and The Wasp | 1 | 0 | 0 | 0 | 0 | 1
# BlacKkKlansman | 0 | 0 | 1 | 0 | 0 | 1
#
# But this isn't what we probably want. Instead this would be a better representation:
#
# Title | Action | Comedy | Drama | Musical
# :---: | :---: | :---: | :---: | :---: |
# Mission: Impossible - Fallout | 1 | 0 | 1 | 0
# Mama Mia: Here We Go Again | 0 | 1 | 0 | 1
# Ant-Man and The Wasp | 1 | 1 | 0 | 0
# BlacKkKlansman | 0 | 1 | 1 | 0
#
# Let's see how we might do this in code
#
# + colab={"base_uri": "https://localhost:8080/", "height": 171} colab_type="code" id="G_Y7mmEKo9Fg" outputId="78125e7b-324f-4b8c-f71c-0f85277220ba"
df = pd.DataFrame({'Title': ['Mission: Impossible - Fallout', 'Mama Mia: Here We Go Again',
'Ant-Man and The Wasp', 'BlacKkKlansman' ],
'Genre1': ['Action', 'Comedy', 'Action', 'Drama'],
'Genre2': ['Drama', 'Musical', 'Comedy', 'Comedy']})
df
# + colab={} colab_type="code" id="xDyEYXhHpEo7"
one_hot_1 = pd.get_dummies(df['Genre1'])
one_hot_2 = pd.get_dummies(df['Genre2'])
# + colab={"base_uri": "https://localhost:8080/", "height": 171} colab_type="code" id="aObvufy0vYwZ" outputId="d81366fd-a64c-4b7a-95b1-952571258687"
# now get the intersection of the column names
s1 = set(one_hot_1.columns.values)
s2 = set(one_hot_2.columns.values)
intersect = s1 & s2
only_s1 = s1 - intersect
only_s2 = s2 - intersect
# now logically or the intersect
logical_or = one_hot_1[list(intersect)] | one_hot_2[list(intersect)]
# then combine everything
combined = pd.concat([one_hot_1[list(only_s1)], logical_or, one_hot_2[list(only_s2)]], axis=1)
combined
### Now drop the two original columns and add the one hot encoded columns
df= df.drop('Genre1', axis=1)
df= df.drop('Genre2', axis=1)
df = df.join(combined)
df
# + [markdown] colab_type="text" id="sxM-Rw_ZwCMU"
# That looks more like it!!!
#
# ## The task: Predict Housing Prices
# Your task is to create a regession classifier that predicts house prices. The data and a description of
#
# * [The description of the data](https://raw.githubusercontent.com/zacharski/ml-class/master/data/housePrices/data_description.txt)
# * [The CSV file](https://raw.githubusercontent.com/zacharski/ml-class/master/data/housePrices/data.csv)
#
#
# Minimally, your classifier should be trained on the following columns:
# + colab={} colab_type="code" id="7RfTLx11vicC"
numericColumns = ['LotFrontage', 'LotArea', 'OverallQual', 'OverallCond', '1stFlrSF', '2ndFlrSF', 'GrLivArea',
'FullBath', 'HalfBath', 'Bedroom', 'Kitchen']
categoryColumns = ['MSZoning', 'Street', 'Alley', 'LotShape', 'LandContour',
'Utilities', 'LotConfig', 'LandSlope', 'Neighborhood', 'BldgType',
'HouseStyle', 'RoofStyle', 'RoofMatl', '' ]
# Using multicolumns is optional
multicolumns = [['Condition1', 'Condition2'], ['Exterior1st', 'Exterior2nd']]
# + [markdown] colab_type="text" id="PWcqhPixyfZI"
# You are free to use more columns than these. Also, you may need to process some of the columns.
# Here are the requirements:
#
# ### 1. Drop any data rows that contain Nan in a column.
# Once you do this you should have around 1200 rows.
# ### 2. Use the following train_test_split parameters
# ```
# train_test_split( originalData, test_size=0.20, random_state=42)
# ```
#
# ### 3. You are to compare Linear Regression and Elastic Net
# ### 4. You should use 10 fold cross validation (it is fine to use grid search)
# ### 5. When finished tuning your model, determine the accuracy on the test data using RMSE.
#
# # Performance Bonus
# You are free to adjust any hyperparameters but do so before you evaluate the test data. You may get up to 15xp bonus for improved accuracy.
#
# Good luck!
# + [markdown] colab_type="text" id="FBCp4tvCyCDr"
#
# + colab={} colab_type="code" id="MY9EdczzwAxT"
| labs/regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: twin-causal-model
# language: python
# name: twin-causal-model
# ---
# # Twin-Causal-Net Example
# ***
# The following notebook shows how to use the `twincausal` *Python* library.
# #### Import the libraries and generate synthetic uplift data
import twincausal.utils.data as twindata
from twincausal.model import twin_causal
from sklearn.model_selection import train_test_split
from twincausal.utils.performance import qini_curve, qini_barplot
X, T, Y = twindata.generator(scenario=3, random_state=123456) # Generate fake uplift data
X_train, X_test, T_train, T_test, Y_train, Y_test = train_test_split(X, T, Y, test_size=0.5, random_state=123)
# #### Initialize the model
input_size = X.shape[1] # Number of features, the model will handle automatically the treatment variable
# How to train a twin-neural model with 1-hidden layer
uplift = twin_causal(nb_features=input_size, # required parameter
# optional hyper-parameters for fine-tuning
nb_hlayers=1, nb_neurons=256, lrelu_slope=0.05, batch_size=256, shuffle=True,
max_iter=100, learningRate=0.009, reg_type=1, l_reg_constant=0.001,
prune=True, gpl_reg_constant=0.005, loss="uplift_loss",
# default parameters for reporting
learningCurves=True, save_model=True, verbose=False, logs=True,
random_state=1234)
uplift # Print model architecture
# #### Fitting the model
uplift.fit(X_train, T_train, Y_train, val_size=0.3) # You can control the proportion of obs to use for validation
# #### Predict and visualize
# +
# Uncomment the following if you want to load the "best" model based on the Qini coefficient obtained in the validation set
# Needs the arg save_model to be set to True
# import torch
# uplift.load_state_dict(torch.load("runs/Models/_twincausal/...")) # Change the path accordingly
# -
pred_our_loss = uplift.predict(X_test)
_, q = qini_curve(T_test, Y_test, pred_our_loss)
print('The Qini coefficient is:', q)
# #### Fit and compare models with no hidden layers and different losses
# +
# Change nb_hlayers to 0 and regularization to L2; keep default hyper-parameters
# -
twin = twin_causal(nb_features=input_size, nb_hlayers=0, reg_type=2)
twin.fit(X_train, T_train, Y_train)
pred_twin_loss = twin.predict(X_test)
_, q = qini_curve(T_test, Y_test, pred_twin_loss)
print('The Qini coefficient using the uplift loss is:', q)
# +
# Now change the loss type to use the usual binary cross entropy (or logistic_loss)
# -
logistic = twin_causal(nb_features=input_size, nb_hlayers=0, reg_type=2, loss="logistic_loss")
logistic.fit(X_train, T_train, Y_train)
pred_log_loss = logistic.predict(X_test)
_, q = qini_curve(T_test, Y_test, pred_log_loss)
print('The Qini coefficient using the logistic loss is:', q)
| examples/twin_causal_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Numpy <img align="right" src="../Supplementary_data/dea_logo.jpg">
#
# * [**Sign up to the DEA Sandbox**](https://docs.dea.ga.gov.au/setup/sandbox.html) to run this notebook interactively from a browser
# * **Compatibility**: Notebook currently compatible with both the `NCI` and `DEA Sandbox` environments
# * **Prerequisites**: Users of this notebook should have a basic understanding of:
# * How to run a [Jupyter notebook](01_jupyter_notebooks.ipynb)
# ## Background
# `Numpy` is a Python library which adds support for large, multi-dimension arrays and metrics, along with a large collection of high-level mathematical functions to operate on these arrays.
# More information about `numpy` arrays can be found [here](https://en.wikipedia.org/wiki/NumPy).
#
# ## Description
# This notebook is designed to introduce users to `numpy` arrays using Python code in Jupyter Notebooks via JupyterLab.
#
# Topics covered include:
#
# * How to use `numpy` functions in a Jupyter Notebook cell
# * Using indexing to explore multi-dimensional `numpy` array data
# * `Numpy` data types, broadcasting and booleans
# * Using `matplotlib` to plot `numpy` data
#
# ***
# ## Getting started
# To run this notebook, run all the cells in the notebook starting with the "Load packages" cell. For help with running notebook cells, refer back to the [Jupyter Notebooks notebook](01_Jupyter_notebooks.ipynb).
# ### Load packages
#
# In order to be able to use `numpy` we need to import the library using the special word `import`. Also, to avoid typing `numpy` every time we want to use one if its functions we can provide an alias using the special word `as`:
import numpy as np
# ### Introduction to Numpy
#
# Now, we have access to all the functions available in `numpy` by typing `np.name_of_function`. For example, the equivalent of `1 + 1` in Python can be done in `numpy`:
np.add(1, 1)
# Although this might not at first seem very useful, even simple operations like this one can be much quicker in `numpy` than in standard Python when using lots of numbers (large arrays).
#
# To access the documentation explaining how a function is used, its input parameters and output format we can press `Shift+Tab` after the function name. Try this in the cell below
np.add
# By default the result of a function or operation is shown underneath the cell containing the code. If we want to reuse this result for a later operation we can assign it to a variable:
a = np.add(2, 3)
# The contents of this variable can be displayed at any moment by typing the variable name in a new cell:
a
# ### Numpy arrays
# The core concept in `numpy` is the "array" which is equivalent to lists of numbers but can be multidimensional.
# To declare a `numpy` array we do:
np.array([1, 2, 3, 4, 5, 6, 7, 8, 9])
# Most of the functions and operations defined in `numpy` can be applied to arrays.
# For example, with the previous operation:
# +
arr1 = np.array([1, 2, 3, 4])
arr2 = np.array([3, 4, 5, 6])
np.add(arr1, arr2)
# -
# But a more simple and convenient notation can also be used:
arr1 + arr2
# #### Indexing
# Arrays can be sliced and diced. We can get subsets of the arrays using the indexing notation which is `[start:end:stride]`.
# Let's see what this means:
# +
arr = np.array([0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15])
print("6th element in the array:", arr[5])
print("6th element to the end of array", arr[5:])
print("start of array to the 5th element", arr[:5])
print("every second element", arr[::2])
# -
# Try experimenting with the indices to understand the meaning of `start`, `end` and `stride`.
# What happens if you don't specify a start?
# What value does `numpy` uses instead?
# Note that `numpy` indexes start on `0`, the same convention used in Python lists.
#
# Indexes can also be negative, meaning that you start counting from the end.
# For example, to select the last 2 elements in an array we can do:
arr[-2:]
# ### Multi-dimensional arrays
# `Numpy` arrays can have multiple dimensions. For example, we define a 2-dimensional `(1,9)` array using nested square bracket:
#
# <img src="../Supplementary_data/07_Intro_to_numpy/numpy_array_t.png" alt="drawing" width="600" align="left"/>
np.array([[1, 2, 3, 4, 5, 6, 7, 8, 9]])
# To visualise the shape or dimensions of a `numpy` array we can add the suffix `.shape`
print(np.array([1, 2, 3, 4, 5, 6, 7, 8, 9]).shape)
print(np.array([[1, 2, 3, 4, 5, 6, 7, 8, 9]]).shape)
print(np.array([[1], [2], [3], [4], [5], [6], [7], [8], [9]]).shape)
# Any array can be reshaped into different shapes using the function `reshape`:
np.array([1, 2, 3, 4, 5, 6, 7, 8]).reshape((2, 4))
# If you are concerned about having to type so many squared brackets, there are more simple and convenient ways of doing the same:
print(np.array([1, 2, 3, 4, 5, 6, 7, 8, 9]).reshape(1, 9).shape)
print(np.array([1, 2, 3, 4, 5, 6, 7, 8, 9]).reshape(9, 1).shape)
print(np.array([1, 2, 3, 4, 5, 6, 7, 8, 9]).reshape(3, 3).shape)
# Also there are shortcuts for declaring common arrays without having to type all their elements:
print(np.arange(9))
print(np.ones((3, 3)))
print(np.zeros((2, 2, 2)))
# ### Arithmetic operations
# `Numpy` has many useful arithmetic functions.
# Below we demonstrate a few of these, such as mean, standard deviation and sum of the elements of an array.
# These operation can be performed either across the entire array, or across a specified dimension.
arr = np.arange(9).reshape((3, 3))
print(arr)
print("Mean of all elements in the array:", np.mean(arr))
print("Std dev of all elements in the array:", np.std(arr))
print("Sum of all elements in the array:", np.sum(arr))
print("Mean of elements in array axis 0:", np.mean(arr, axis=0))
print("Mean of elements in array axis 1:", np.mean(arr, axis=1))
# ### Numpy data types
# `Numpy` arrays can contain numerical values of different types. These types can be divided in these groups:
#
# * Integers
# * Unsigned
# * 8 bits: `uint8`
# * 16 bits: `uint16`
# * 32 bits: `uint32`
# * 64 bits: `uint64`
# * Signed
# * 8 bits: `int8`
# * 16 bits: `int16`
# * 32 bits: `int32`
# * 64 bits: `int64`
#
# * Floats
# * 32 bits: `float32`
# * 64 bits: `float64`
#
# We can specify the type of an array when we declare it, or change the data type of an existing one with the following expressions:
# +
# Set datatype dwhen declaring array
arr = np.arange(5, dtype=np.uint8)
print("Integer datatype:", arr)
arr = arr.astype(np.float32)
print("Float datatype:", arr)
# -
# ### Broadcasting
#
# The term broadcasting describes how `numpy` treats arrays with different shapes during arithmetic operations.
# Subject to certain constraints, the smaller array is "broadcast" across the larger array so that they have compatible shapes.
# Broadcasting provides a means of vectorizing array operations so that looping occurs in C instead of Python.
# This can make operations very fast.
# +
a = np.zeros((3, 3))
print(a)
a = a + 1
print(a)
# +
a = np.arange(9).reshape((3, 3))
b = np.arange(3)
a + b
# -
# ### Booleans
# There is a binary type in `numpy` called boolean which encodes `True` and `False` values.
# For example:
# +
arr = arr > 0
print(arr)
arr.dtype
# -
# Boolean types are quite handy for indexing and selecting parts of images as we will see later.
# Many `numpy` functions also work with Boolean types.
# +
print("Number of 'Trues' in arr:", np.count_nonzero(arr))
# Create two boolean arrays
a = np.array([1, 1, 0, 0], dtype=bool)
b = np.array([1, 0, 0, 1], dtype=bool)
# Compare where they match
np.logical_and(a, b)
# -
# ### Introduction to Matplotlib
# This second part introduces `matplotlib`, a Python library for plotting `numpy` arrays as images.
# For the purposes of this tutorial we are going to use a part of `matplotlib` called `pyplot`.
# We import it by doing:
# +
# %matplotlib inline
import matplotlib.pyplot as plt
# -
# An image can be seen as a 2-dimensional array. To visualise the contents of a `numpy` array:
# +
arr = np.arange(100).reshape(10, 10)
print(arr)
plt.imshow(arr)
# -
# We can use the Pyplot library to load an image using the function `imread`:
im = np.copy(plt.imread("../Supplementary_data/07_Intro_to_numpy/africa.png"))
# #### Let's display this image using the `imshow` function.
plt.imshow(im)
# This is a [free stock photo](https://depositphotos.com/42725091/stock-photo-kilimanjaro.html) of Mount Kilimanjaro, Tanzania. A colour image is normally composed of three layers containing the values of the red, green and blue pixels. When we display an image we see all three colours combined.
# Let's use the indexing functionality of `numpy` to select a slice of this image. For example to select the top right corner:
plt.imshow(im[:100,-200:,:])
# We can also replace values in the 'red' layer with the value 255, making the image 'reddish'. Give it a try:
im[:, :, 0] = 255
plt.imshow(im)
# ## Recommended next steps
#
# For more advanced information about working with Jupyter Notebooks or JupyterLab, you can explore [JupyterLab documentation page](https://jupyterlab.readthedocs.io/en/stable/user/notebook.html).
#
# To continue working through the notebooks in this beginner's guide, the following notebooks are designed to be worked through in the following order:
#
# 1. [Jupyter Notebooks](01_Jupyter_notebooks.ipynb)
# 2. [Digital Earth Australia](02_DEA.ipynb)
# 3. [Products and Measurements](03_Products_and_measurements.ipynb)
# 4. [Loading data](04_Loading_data.ipynb)
# 5. [Plotting](05_Plotting.ipynb)
# 6. [Performing a basic analysis](06_Basic_analysis.ipynb)
# 7. **Introduction to Numpy (this notebook)**
# 8. [Introduction to Xarray](08_Intro_to_xarray.ipynb)
# 9. [Parallel processing with Dask](09_Parallel_processing_with_Dask.ipynb)
#
# Once you have you have completed the above nine tutorials, join advanced users in exploring:
#
# * The "DEA datasets" directory in the repository, where you can explore DEA products in depth.
# * The "Frequently used code" directory, which contains a recipe book of common techniques and methods for analysing DEA data.
# * The "Real-world examples" directory, which provides more complex workflows and analysis case studies.
# ***
# ## Additional information
#
# **License:** The code in this notebook is licensed under the [Apache License, Version 2.0](https://www.apache.org/licenses/LICENSE-2.0). Digital Earth Australia data is licensed under the Creative Commons by Attribution 4.0 license.
#
# **Contact:** If you need assistance, please post a question on the [Open Data Cube Slack channel](http://slack.opendatacube.org/) or on the [GIS Stack Exchange](https://gis.stackexchange.com/questions/ask?tags=open-data-cube) using the `open-data-cube` tag (you can view previously asked questions [here](https://gis.stackexchange.com/questions/tagged/open-data-cube)).
# If you would like to report an issue with this notebook, you can file one on [Github](https://github.com/GeoscienceAustralia/dea-notebooks).
#
# **Last modified:** September 2021
# ## Tags
# Browse all available tags on the DEA User Guide's [Tags Index]()
# + raw_mimetype="text/restructuredtext" active=""
# **Tags**: :index:`sandbox compatible`, :index:`NCI compatible`, :index:`numpy`, :index:`matplotlib`, :index:`plotting`, :index:`beginner`
| Beginners_guide/07_Intro_to_numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Libraries
# +
import pandas as pd
import numpy as np
import scipy.stats as stat
from math import sqrt
from mlgear.utils import show, display_columns
from surveyweights import normalize_weights
# -
# ## Load Processed Data
survey = pd.read_csv('responses_processed_national_weighted.csv').fillna('Not presented')
# ## Analysis
options = ['<NAME>, the Democrat', '<NAME>, the Republican']
survey_ = survey.loc[survey['vote_trump_biden'].isin(options)].copy()
options2 = ['<NAME>', '<NAME>']
survey_ = survey_.loc[survey_['vote2016'].isin(options2)].copy()
options3 = ['Can trust', 'Can\'t be too careful']
survey_ = survey_.loc[survey_['gss_trust'].isin(options3)].copy()
options4 = ['Disagree', 'Agree']
survey_ = survey_.loc[survey_['gss_spanking'].isin(options4)].copy()
survey_['lv_weight'] = normalize_weights(survey_['lv_weight'])
survey_['vote_trump_biden'].value_counts(normalize=True) * survey_.groupby('vote_trump_biden')['lv_weight'].mean() * 100
survey_['vote2016'].value_counts(normalize=True) * survey_.groupby('vote2016')['lv_weight'].mean() * 100
survey_['race'].value_counts(normalize=True) * survey_.groupby('race')['lv_weight'].mean() * 100
survey_['education'].value_counts(normalize=True) * survey_.groupby('education')['lv_weight'].mean() * 100
survey_['gss_trust'].value_counts(normalize=True) * survey_.groupby('gss_trust')['lv_weight'].mean() * 100
survey_['gss_spanking'].value_counts(normalize=True) * survey_.groupby('gss_spanking')['lv_weight'].mean() * 100
survey_['noncollege_white'].value_counts(normalize=True) * survey_.groupby('noncollege_white')['lv_weight'].mean() * 100
# +
print('## HIGH TRUST ##')
survey__ = survey_[survey_['gss_trust'] == 'Can trust']
survey__['lv_weight'] = normalize_weights(survey__['lv_weight'])
print(survey__['vote2016'].value_counts(normalize=True) * survey__.groupby('vote2016')['lv_weight'].mean() * 100)
print('-')
print(survey__['vote_trump_biden'].value_counts(normalize=True) * survey__.groupby('vote_trump_biden')['lv_weight'].mean() * 100)
print('-')
print('-')
print('## LOW TRUST ##')
survey__ = survey_[survey_['gss_trust'] == 'Can\'t be too careful']
survey__['lv_weight'] = normalize_weights(survey__['lv_weight'])
print(survey__['vote2016'].value_counts(normalize=True) * survey__.groupby('vote2016')['lv_weight'].mean() * 100)
print('-')
print(survey__['vote_trump_biden'].value_counts(normalize=True) * survey__.groupby('vote_trump_biden')['lv_weight'].mean() * 100)
# +
print('## NONCOLLEGE WHITE ##')
survey__ = survey_[survey_['noncollege_white']]
survey__['lv_weight'] = normalize_weights(survey__['lv_weight'])
print(survey__['vote2016'].value_counts(normalize=True) * survey__.groupby('vote2016')['lv_weight'].mean() * 100)
print('-')
print(survey__['vote_trump_biden'].value_counts(normalize=True) * survey__.groupby('vote_trump_biden')['lv_weight'].mean() * 100)
print('-')
print('-')
print('## NOT "NONCOLLEGE WHITE" ##')
survey__ = survey_[~survey_['noncollege_white']]
survey__['lv_weight'] = normalize_weights(survey__['lv_weight'])
print(survey__['vote2016'].value_counts(normalize=True) * survey__.groupby('vote2016')['lv_weight'].mean() * 100)
print('-')
print(survey__['vote_trump_biden'].value_counts(normalize=True) * survey__.groupby('vote_trump_biden')['lv_weight'].mean() * 100)
# +
print('## NONCOLLEGE WHITE, HIGH SOCIAL TRUST ##')
survey__ = survey_[survey_['noncollege_white'] & (survey_['gss_trust'] == 'Can trust')]
survey__['lv_weight'] = normalize_weights(survey__['lv_weight'])
print(survey__['vote2016'].value_counts(normalize=True) * survey__.groupby('vote2016')['lv_weight'].mean() * 100)
print('-')
print(survey__['vote_trump_biden'].value_counts(normalize=True) * survey__.groupby('vote_trump_biden')['lv_weight'].mean() * 100)
print('-')
print('-')
print('## NONCOLLEGE WHITE, LOW SOCIAL TRUST ##')
survey__ = survey_[survey_['noncollege_white'] & (survey_['gss_trust'] == 'Can\'t be too careful')]
survey__['lv_weight'] = normalize_weights(survey__['lv_weight'])
print(survey__['vote2016'].value_counts(normalize=True) * survey__.groupby('vote2016')['lv_weight'].mean() * 100)
print('-')
print(survey__['vote_trump_biden'].value_counts(normalize=True) * survey__.groupby('vote_trump_biden')['lv_weight'].mean() * 100)
print('-')
print('-')
print('## NOT "NONCOLLEGE WHITE", HIGH SOCIAL TRUST ##')
survey__ = survey_[~survey_['noncollege_white'] & (survey_['gss_trust'] == 'Can trust')]
survey__['lv_weight'] = normalize_weights(survey__['lv_weight'])
print(survey__['vote2016'].value_counts(normalize=True) * survey__.groupby('vote2016')['lv_weight'].mean() * 100)
print('-')
print(survey__['vote_trump_biden'].value_counts(normalize=True) * survey__.groupby('vote_trump_biden')['lv_weight'].mean() * 100)
print('-')
print('-')
print('## NOT "NONCOLLEGE WHITE", LOW SOCIAL TRUST ##')
survey__ = survey_[~survey_['noncollege_white'] & (survey_['gss_trust'] == 'Can\'t be too careful')]
survey__['lv_weight'] = normalize_weights(survey__['lv_weight'])
print(survey__['vote2016'].value_counts(normalize=True) * survey__.groupby('vote2016')['lv_weight'].mean() * 100)
print('-')
print(survey__['vote_trump_biden'].value_counts(normalize=True) * survey__.groupby('vote_trump_biden')['lv_weight'].mean() * 100)
| (K) Analysis - Noncollege White X Trust Shift.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.0 64-bit
# language: python
# name: python3
# ---
# # Objects
#
# In Python, objects are used to represent
# information. Every variable you use in a Python
# program is a reference to an object.
# The values you have been using so far –
# numbers, strings, dicts, lists, etc – are objects.
# They are among the built-in classes of Python,
# i.e., kinds of value that are already defined
# when you start the Python interpreter.
#
# You are not limited to those built-in classes.
# You can use them as a foundation to build your
# own.
#
# ## 1. Example: Points
#
# What if we wanted to define a new kind of value?
# For example, if we wanted to write a program
# to draw a graph, we might want to work with
# cartesian coordinates, representing each
# point as an (x,y) pair. We might represent the
# point as a tuple like `(5,7)`, or we could represent
# it as the list `[5, 7]`, or we could represent
# it as a dict `{"x": 5, "y": 7}`, and that
# might be satisfactory. If we wanted to represent moving a point (x,y) by some distance (dx, dy), we could define a a function like
#
# ```python
# def move(p, d):
# x,y = p
# dx, dy = d
# return (x+dx, y+dy)
# ```
#
# But if we are making a graphics program, we'll need to *move* functions for other graphical objects like rectangles and ovals,
# so instead of naming it `move` we'll need a more descriptive name
# like `move_point`. Also we should give the type contract for
# the function, which we can do with Python type hints. With these
# changes, we get something like this
# +
# Setup
from IPython.core.magic import register_cell_magic
from IPython.display import HTML, display
@register_cell_magic
def bgc(color, cell=None):
script = (
"var cell = this.closest('.jp-CodeCell');"
"var editor = cell.querySelector('.jp-Editor');"
"editor.style.background='{}';"
"this.parentNode.removeChild(this)"
).format(color)
display(HTML('<img src onerror="{}">'.format(script)))
# +
from typing import Tuple
from numbers import Number
def move_point(p: Tuple[Number, Number],
d: Tuple[Number, Number]) \
-> Tuple[Number, Number]:
x, y = p
dx, dy = d
return (x+dx, y+dy)
# -
# A simple test case increases our confidence that this works:
assert move_point((3,4),(5,6)) == (8,10)
#
# ### Can we do better?
#
# We aren't really satisfied with using tuples to
# represent points. What we'd really
# like is to express the concept of adding two points
# more concisely, as `(3,4) + (5,6)`. What would happen if we
# tried this?
(3,4) + (5,6)
# That's not what we wanted! Would it be better if we represented
# points as lists?
[3,4] + [5,6]
#
# No better. Maybe as dicts?
{"x": 3, "y": 4} + {"x": 5, "y": 6}
#
# That is not much of an improvement, although
# an error message is usually better than silently
# producing a bad result.
# What we really want is not to use one of the
# existing representations like lists or tuples or dicts,
# but to define a new representation for points.
#
# ## 2. A new representation
#
# Each data type in Python, including list, tuple,
# and dict, is defined as a *class* from which
# *objects* can be constructed. We can also define
# our own classes, to construct new kinds of objects.
# For example, we can make a new class `Point` to
# represent points.
#
# ```python
# class Point:
# """An (x,y) coordinate pair"""
# ```
#
# Inside the class we can define *methods*, which are like functions
# that are specialized for the new representation. The first
# method we should define is a *constructor* with the name `__init__`.
# The constructor describes how to create a new `Point` object:
#
# ```python
# class Point:
# """An (x,y) coordinate pair"""
# def __init__(self, x: Number, y: Number):
# self.x = x
# self.y = y
# ```
#
# ## 3. Instance variables
#
# Notice that the first argument to the constructor method is
# `self`, and within the method we refer to `self.x` and `self.y`.
# In a method that operates on some object *o*, the first argument
# to the method will always be `self`, which refers to the whole
# object *o*. Within the `self` object we can store *instance
# variables*, like `self.x` and `self.y`
# for the *x* and *y* coordinates of a point.
#
# ## 4. Methods
#
# What about defining an operation for moving a point? Instead of
# adding `_point` to the name of a `move` function, we can just
# put the function (now called a *method*) *inside* the `Point`
# class:
#
# ```python
# def move(self, d: Point) -> Point:
# """(x, y).move(dx, dy) = (x + dx, y + dy)"""
# x = self.x + d.x
# y = self.y + d.y
# return Point(x,y)
# ```
# ><span style='background :yellow' >**Type hints note:**</span> Because the Point class is a user-defined (not builtin) class, an extra type variable definition is required for the `Point` type hints to be recognized as valid type names:
from typing import TypeVar
Point = TypeVar('Point')
#
# Notice that the *instance variables*
# `self.x` and `self.y` we created in the constructor
# can be used in the `move` method. They are part of
# the object, and can be used by any method in the class.
# The instance variables of the other `Point` object `d`
# are also available
# in the `move` method. Let's look at how these objects
# are passed to the `move` method.
#
# Here is the complete `Point` code, including [type hints](https://towardsdatascience.com/python-type-hints-docstrings-7ec7f6d3416b), [docstrings](https://realpython.com/documenting-python-code/) and [doctests](https://www.digitalocean.com/community/tutorials/how-to-write-doctests-in-python).
# +
from typing import TypeVar
from numbers import Number
Point = TypeVar('Point')
class Point:
"""An (x,y) coordinate pair"""
def __init__(self, x: Number, y: Number):
"""Create a new Point at (x,y)
Args:
x: x coordinate
y: y coordinate
>>> p = Point(2, 3); (p.x, p.y)
(2, 3)
"""
self.x = x
self.y = y
def move(self, d: Point) -> Point:
"""(x, y).move(dx, dy) = (x + dx, y + dy)
Args:
d: a Point
Returns:
a new Point
>>> p = Point(2, 3).move(Point(5, 6)); (p.x, p.y)
(7, 9)
"""
x = self.x + d.x
y = self.y + d.y
return Point(x, y)
# -
# And a quick doctest check for the code so far:
import doctest
doctest.testmod(verbose=True)
#
# ## 5. Method calls
#
# Next we'll create two `Point` objects and call the `move` method
# to create a third `Point` object with the sums of their *x* and
# *y* coordinates:
# +
p = Point(3,4)
v = Point(5,6)
m = p.move(v)
assert m.x == 8 and m.y == 10
# -
#
# At first it may seem confusing that we defined the ``move`` method with two arguments, `self` and `d`, but
# it looks like we passed it only one argument, `v`. In fact
# we passed it both points: `p.move(v)` passes `p` as the `self` argument and `v` as the `d` argument. We use the variable
# before the `.`, like `p` in this case, in two different ways: To find the right method (function) to call, by looking inside the *class* to
# which `p` belongs, and to pass as the `self` argument to the method.
#
# The `move` method above returns a new `Point` object at the
# computed coordinates. A method can also change the values of
# instance variables. For example, suppose we add a `move_to`
# method to `Point`:
# +
# Remove the previous definition of Point from the global namespace:
# %reset -sf
from typing import TypeVar
from numbers import Number
Point = TypeVar('Point')
class Point:
"""An (x,y) coordinate pair"""
def __init__(self, x: Number, y: Number):
self.x = x
self.y = y
def move(self, d: Point) -> Point:
"""(x, y).move(dx, dy) = (x + dx, y + dy)"""
x = self.x + d.x
y = self.y + d.y
return Point(x,y)
def move_to(self, new_x, new_y):
"""Change the coordinates of this Point"""
self.x = new_x
self.y = new_y
# -
# The `move_to` method does not return a new point (it returns `None`), but
# it changes an existing point *object*.
# +
m = Point(8,10)
m.move_to(19,23)
assert m.x == 19 and m.y == 23
# -
#
# #### *Check your understanding*
#
# Consider class `Pet` and object `my_pet`.
# What are the *instance variables* of `my_pet`?
# What are the values of those instance variables
# after executing the code below?
#
# ```python
# class Pet:
# def __init__(self, kind: str, name: str):
# self.species = kind
# self.called = name
#
# def rename(self, new_name):
# self.called = new_name
#
# my_pet = Pet("canis familiaris", "fido")
# ```
#
# ## 6. A little magic
#
# We said above that what we really wanted was to express
# movement of points very compactly, as addition. We
# saw that addition of tuples or lists did not act as we
# wanted; instead of `(3,4) + (5,6)` giving us `(8,10)`, it
# gave us `(3,4,5,6)`. We can almost get what we want by describing
# how we want `+` to act on `Point` objects. We do this by
# defining a *special method* `__add__`:
#
# ```python
# def __add__(self, other: "Point"):
# """(x,y) + (dx, dy) = (x+dx, y+dy)"""
# return Point(self.x + other.x, self.y + other.y)
# ```
#
# Special methods are more commonly known as *magic methods*.
# They allow us to define how arithmetic operations like `+`
# and `-` work for each class of object, as well as
# comparisons like `<` and `==`, and some other operations.
# If `p` is a `Point` object, then `p + q` is interpreted as
# `p.__add__(q)`. So finally we get a very compact and
# readable notation:
#
# ```python
# p = Point(3,4)
# v = Point(5,6)
# m = p.move(v)
#
# assert m.x == 8 and m.y == 10
# ```
#
# More: [Appendix on magic methods](appendix_Special)
#
# ## Magic for printing
#
# Suppose we wanted to print a `Point` object. We
# could do it this way:
#
# ```python
# print(f"p is ({p.x}, {p.y})")
# ```
#
# That would give us a reasonable printed representation,
# like "p is (3, 4)", but it is tedious, verbose, and easy to
# get wrong. What if we just wrote
#
# ```python
# print(f"p is {p}")
# ```
#
# That would be simpler, but the result is not very
# useful:
#
# ```python
# p is <__main__.Point object at 0x10b4a22e0>
# ```
#
# ### `str()`d, not shaken
#
# If we want to print `Point` objects as simply
# as we print strings and numbers, but we want the
# printed representation to be readable, we will need
# to write additional methods to describe how a
# `Point` object should be converted to a string.
#
# In fact, in Python we normally write two magic
# methods for this: `__str__` describes how it
# should be represented by the `str()` function,
# which is the representation used in `print`
# or in an f-string like `f"it is {p}"`. We might
# decide that we want the object created by `Point(3,2)`
# to print as "(3, 2)". We would then write a
# `__str__` method in the `Point` class like this:
#
# ```python
# def __str__(self) -> str:
# return f"({self.x}, {self.y})"
# ```
#
# Now if we again execute
#
# ```python
# print(f"p is {p}")
# ```
#
# we get a more useful result:
#
# ```python
# p is (3, 4)
# ```
#
# ### A `repr()` for debugging
#
# Usually we will also want to provide a different
# string representation that is useful in debugging
# and at the Python command line interface. The
# string representation above may be fine for end users,
# but for the software developer it does not differentiate
# between a tuple `(3, 4)` and a `Point` object `(3, 4)`.
# We can define a `__repr__` method to give a string
# representation more useful in debugging. The function
# `repr(x)` is actually a call on the `__repr__` method
# of `x`, i.e., `x.__repr__()`.
#
# Although
# Python will permit us to write whatever `__repr__`
# method we choose, by accepted convention is to make
# it look like a call on the constructor, i.e., like
# Python code to create an identical object. Thus, for
# the `Point` class we might write:
#
# ```python
# def __repr__(self) -> str:
# return f"Point({self.x}, {self.y})"
# ```
#
# Now if we write
#
# ```python
# print(f"repr(p) is {repr(p)}")
# ```
#
# we will get
#
# ```python
# repr(p) is Point(3, 4)
# ```
#
# The `print` function automatically applies the `str` function to its
# arguments, so defining a good `__str__` method will ensure it
# is printed as you like in most cases. Oddly, though,
# the `__str__` method for `list` applies the `__repr__` method
# to each of its arguments, so if we write
#
# ```python
# print(p)
# print(v)
# print([p, v])
# ```
#
# we get
#
# ```
# (3, 4)
# (5, 6)
# [Point(3, 4), Point(5, 6)]
# ```
#
# #### *Check your understanding*
#
# Which of the following are legal, and what
# values do they return?
#
# * `str(5)`
# * `(5).str()`
# * `(5).__str__()`
# * `__str__(5)`
# * `repr([1, 2, 3])`
# * `[1, 2, 3].repr()`
# * `[1, 2, 3].__repr__()`
#
# What does the following little program print?
#
# ```python
# class Wrap:
# def __init__(self, val: str):
# self.value = val
#
# def __str__(self) -> str:
# return self.value
#
# def __repr__(self) -> str:
# return f"Wrap({self.value})"
#
# a = Wrap("alpha")
# b = Wrap("beta")
# print([a, b])
# ```
#
# ## Variables *refer* to objects
#
# Before reading on, try to predict what the following
# little program will print.
#
# ```python
# x = [1, 2, 3]
# y = x
# y.append(4)
# print(x)
# print(y)
# ```
#
# Now execute that program. Did you get the result you
# expected? If it surprised you, try visualizing it in
# PythonTutor (http://pythontutor.com/). You should get a
# diagram that looks like this:
#
# 
#
# `x` and `y` are distinct variables, but they are both references to the same list. When we change `y` by appending 4, we are changing the same object
# that `x` refers to. We say that `x` and `y` are *aliases*, two names for the same object.
#
# Note this is very different from the following:
#
# ```python
# x = [1, 2, 3]
# y = [1, 2, 3]
# y.append(4)
# print(x)
# print(y)
# ```
#
# Each time we create a list like `[1, 2, 3]`, we are creating a distinct
# list. In this seocond version of the program, `x` and `y` are not
# aliases.
#
# 
#
# It is essential to remember that variables hold *references* to objects, and
# there may be more than one reference to the same object. We can observe the
# same phenomenon with classes we add to Python. Consider this program:
#
# ```python
# class Point:
# """An (x,y) coordinate pair"""
# def __init__(self, x: int, y: int):
# self.x = x
# self.y = y
#
# def move(self, dx: int, dy: int):
# self.x += dx
# self.y += dy
#
# p1 = Point(3,5)
# p2 = p1
# p1.move(4,4)
# print(p2.x, p2.y)
# ```
#
# Once again we have created two variables that are *aliases*, i.e., they
# refer to the same object. PythonTutor illustrates:
#
# 
#
# Note that `Point` is a reference to the *class*, while `p1` and `p2` are references to the Point *object* we created from the Point class. When we call `p1.move`, the `move` method of class `Point` makes a change to
# the object that is referenced by both `p1` and `p2`. We often say that
# a method like `move` *mutates* an object.
#
# There is another way we could have written a method like `move`.
# Instead of *mutating* the object (changing the values of its *fields*
# `x` and `y`), we could have created a new Point object at the
# modified coordinates:
#
# ```python
# class Point:
# """An (x,y) coordinate pair"""
# def __init__(self, x: int, y: int):
# self.x = x
# self.y = y
#
# def moved(self, dx: int, dy: int) -> "Point":
# return Point(self.x + dx, self.y + dy)
#
# p1 = Point(3,5)
# p2 = p1
# p1 = p1.moved(4,4)
# print(p1.x, p1.y)
# print(p2.x, p2.y)
# ```
#
# Notice that method `moved`, unlike method `move` in the prior example,
# return a new Point object that is distinct from the Point object that was aliased. Initially `p1` and `p2` may be aliases, after `p2 = p1`:
#
# 
#
# But after `p1 = p1.moved(4,4)`, `p1` refers to a new, distinct object:
#
# 
#
# As we saw with the `list` example, aiasing applies to objects
# from the built-in Python classes as well as to objects
# from the classes that you will write. It just hasn't been
# apparent until now, because many of the built-in classes
# are *immutable*: They do not have any methods that
# change the values stored in an object. For example, when
# we write `3 + 5`, we are actually calling `(3).__add__(5)`;
# The `__add__` method does not change the value of `3` (that
# would be confusing!) but instead returns a new object `8`.
#
# We will write both *immutable* and *mutable* classes.
# Aliasing of mutable objects is often a mistake, but not always.
# Later we will intentionally created aliases to access
# mutable objects. The important thing is to be aware of it.
#
# ## Combining Objects: Composing
#
# The *instance variables* defined in a class and stored
# in the objects of that class can themselves be objects.
# We can make lists of objects, tuples of objects, etc.
#
# Often we will want to create a new class with instance
# variables that are objects created from classes that
# we have previously created. For example, if we create a
# new class `Rect` to represent rectangles, we might want
# to use `Point` objects to represent two corners of
# the rectangle:
#
# ```python
# class Rect:
# """A rectangle is represented by a pair of points
# (x_min, y_min), (x_max, y_max) at opposite corners.
# Whether (x_min, y_min) is lower left or upper left
# depends on the coordinate system.
# """
# def __init__(self, xy_min: Point, xy_max: Point):
# self.min_pt = xy_min
# self.max_pt = xy_max
#
# def area(self) -> Number:
# """Area is height * width"""
# height = self.max_pt.x - self.min_pt.x
# width = self.max_pt.y - self.min_pt.y
# return height * width
#
# def translate(self, delta: Point) -> "Rect":
# """New rectangle offset from this one by delta as movement vector"""
# return Rect(self.min_pt + delta, self.max_pt + delta)
#
# def __repr__(self) -> str:
# return f"Rect({repr(self.min_pt)}, {repr(self.max_pt)}"
#
# def __str__(self) -> str:
# return f"Rect({str(self.min_pt)}, {str(self.max_pt)})"
#
# p1 = Point(3,5)
# p2 = Point(8,7)
# r1 = Rect(p1, p2)
# mvmt = Point(4, 5)
# r2 = r1.translate(mvmt) # Treat Point(4,5) as (dx, dy)
# print(f"{r1} + {mvmt} => {r2}")
# print(f"Area of {r1} is {r1.area()}")
# ```
#
# Note that the `height` and `width` are local variables
# that exist only while method `area` is executing.
# `min_pt` and `max_pt`, on the other hand, are
# *instance variables* that are stored within the
# `Rect` object.
#
# #### *Check your understanding*
#
# Suppose we ran the above code in PythonTutor.
# (PythonTutor
# cannot import `Number`, but for the examples we could
# replace it with `int`.) What picture would it draw
# of `r1`? Would `height` and `width` in method
# `area` be included as instance variables? Why or
# why not?
#
# ## Wrapping and delegation
#
# Sometimes we want a class of objects that is almost
# like an existing class, but with a little extra
# information or a few new methods. One way to do this
# is to build a new class that _wraps_ an existing class,
# often a built-in class like `list` or `dict`. (In
# [the next chapter](02_1_Inheritance.md) we will
# see another approach.)
#
# Suppose we wanted objects that provide some of the same
# functionality as `list` objects, and also some new
# functionality or some restrictions. For example, we
# might want a method `area` that returns the sum of
# the areas of all the `Rect` objects in the `RectList`:
#
# ```python
# class RectList:
# """A collection of Rects."""
#
# def __init__(self):
# self.elements = [ ]
#
# def area(self) -> Number:
# total = 0
# for el in self.elements:
# total += el.area()
# return total
# ```
#
# That seems reasonable, but how do we add `Rect` objects to the
# `Rectlist`?
#
# We do *not* want to do it this way:
#
# ```python
# li = RectList()
# # DON'T DO THIS
# li.elements.append(Rect(Point(3,3), Point(5,7)))
# li.elements.append(Rect(Point(2,2), Point(3,3)))
# ```
#
# As a general rule, we should be cautious about accessing the instance
# variables of an object outside of methods of the object's class,
# and we should especially avoid modifying instance variables anywhere
# except in methods. Code that "breaks the abstraction", like the example
# above calling the `append` method of the `elements` instance variable, is
# difficult to read and maintain. So we want instead to give `RectList`
# it's own `append` method, so that we can write
#
# ```python
# li = RectList()
# li.append(Rect(Point(3,3), Point(5,7)))
# li.append(Rect(Point(2,2), Point(3,3)))
# print(f"Combined area is {li.area()}")
# ```
#
# The `append` method can be very simple!
#
# ```python
# def append(self, item: Rect):
# """Delegate to elements"""
# self.elements.append(item)
# ```
#
# We call this *delegation* because `append` method of `RectList` method
# just hands off the work to the `append` method of class `list`. When we
# write a *wrapper* class, we typically write several such
# trivial *delegation* methods.
#
# Wrapping and delegation work well when we want the wrapper class
# (like `RectList` in this example) to
# have a few of the same methods as the wrapped class (`list`). When
# we want the new collection class to have all or nearly all the methods
# of an existing collection, the *inheritance* approach introduced in the
# next chapter is more appropriate.
#
| notebooks/01_1_Objects.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import BOCD
import numpy as np
import matplotlib.pyplot as plt
from functools import partial
import pandas as pd
import warnings
import random
warnings.filterwarnings('ignore')
import matplotlib
import matplotlib.cm as cm
from Htests import online, Mood
import math
from itertools import cycle
from ruptures.utils import pairwise
import ruptures as rpt
# # Offline Ruptures Demo
# +
from math import log
from ruptures.base import BaseCost
from numpy.linalg import inv
from numpy.linalg import det
import ruptures as rpt
# Cost function 3 in Truong, Charles & Oudre, Laurent & <NAME>. (2018). A review of change point detection methods.
# Changes in mean and variance of gaussian data
class MyCost(BaseCost):
"""Custom cost"""
# The 2 following attributes must be specified for compatibility.
model = ""
min_size = 2
def fit(self, signal):
"""Set the internal parameter."""
self.signal = signal
return self
def error(self, start, end):
"""Return the approximation cost on the segment [start:end].
Args:
start (int): start of the segment
end (int): end of the segment
Returns:
float: segment cost
"""
sub = self.signal[start:end]
mean = sub.mean()
var = sub.var()
return len(sub)*log(var) + ((sub-mean)**2).sum()/var
# +
np.random.seed(1)
def generate_normal_time_series(num, minl=50, maxl=1000):
data = np.array([], dtype=np.float64)
partition = np.random.randint(minl, maxl, num)
for p in partition:
mean = np.random.normal()*1
scale = np.random.normal()*2
if scale < 0:
scale = scale * -1
tdata = np.random.normal(mean, scale, p)
data = np.concatenate((data, tdata))
return data, partition
data, partition = generate_normal_time_series(4, 100, 200)
actual_cps = partition.cumsum()[:-1]
# cost
algo = rpt.Pelt(custom_cost=MyCost(), jump = 1).fit(data)
cps = [cp - 1 for cp in algo.predict(pen=20)]
cps.pop(-1)
cps = np.insert(cps, 0, 0)
bkps = sorted(cps)
bkps = np.insert(bkps, len(bkps), len(data) - 1)
# plotting
color_cycle = cycle(["#4286f4", "#f44174"])
fig, (ax1) = plt.subplots(1, 1, figsize = (20, 5))
ax1.margins(x=0)
ax1.plot(data)
for (start, end), col in zip(pairwise(bkps), color_cycle):
ax1.axvspan(start, end, facecolor=col, alpha=0.2)
for y in actual_cps:
ax1.axvline(x = y, color = 'black')
plt.show()
# -
# # BOCD Demo
# +
R, cps, maxes_R, pred_mean, pred_var = BOCD.online_changepoint_detection(data, partial(BOCD.constant_hazard, 100), BOCD.GaussianUnknownMeanVar(1, 1, 1, 0))
bkps = sorted(cps) + [len(data)-1]
# plotting
color_cycle = cycle(["#4286f4", "#f44174"])
fig, (ax1, ax2) = plt.subplots(2, 1, figsize = (20, 10), sharex = True)
ax1.margins(x=0)
ax1.set_ylabel(r'$x_t$')
ax2.margins(x=0)
ax2.set_ylabel(r'$r_t$')
fig.subplots_adjust(hspace=0)
plt.setp([a.get_xticklabels() for a in fig.axes[:-1]], visible=False)
ax1.plot(data)
for (start, end), col in zip(pairwise(bkps), color_cycle):
ax1.axvspan(start, end, facecolor=col, alpha=0.2)
for y in actual_cps:
ax1.axvline(x = y, color = 'black')
sparsity = 1
ax2.pcolor(np.array(range(0, len(R[:,0]), sparsity)),
np.array(range(0, len(R[:,0]), sparsity)),
-np.log(R[0::sparsity, 0::sparsity]),
cmap=cm.gray, vmin=0, vmax=30)
ax2.plot(maxes_R, color = 'Red')
ax1.plot(pred_mean[:-1])
ax1.plot(pred_mean[:-1] + np.sqrt(pred_var[:-1]), linestyle = ':', color = 'black')
ax1.plot(pred_mean[:-1] - np.sqrt(pred_var[:-1]), linestyle = ':', color = 'black')
plt.show()
# -
# # HTest Demo
# Import thresholds
Mood_500 = np.genfromtxt('Mood500.txt')
# +
np.random.seed(2)
data = np.concatenate((np.random.normal(size = 200), np.random.normal(scale = 2, size = 200)))
cps, detected, change_type = online(data, Mood, Mood_500)
statistics = []
for i in range(19):
statistics.append(0)
for i in range(20, detected[1] + 2):
window = data[:i]
_, D = Mood(window)
statistics.append(D)
# Plotting
fig, (ax1, ax2) = plt.subplots(1, 2, figsize = (20 + 0.2, 5))
fig.subplots_adjust(wspace=0.2)
ax1.margins(x=0)
color_cycle = cycle(["#4286f4", "#f44174"])
bkps = sorted(cps) + [len(data) - 1]
ax1.plot(range(1, len(data) + 1), data)
for (start, end), col in zip(pairwise(bkps), color_cycle):
ax1.axvspan(start, end, facecolor=col, alpha=0.2)
ax1.set_ylabel(r'$x_t$', size = 14)
ax1.axvline(x = 200, color = 'Black')
bkps = sorted(cps) + [detected[1] + 2 - 1]
ax2.margins(x = 0)
ax2.plot(statistics)
ax2.plot(range(20, detected[1] + 2), Mood_500[19:detected[1] + 1], color = 'Black', linestyle = "--")
ax2.axvline(x = 200, color = 'Black')
ax2.set_ylabel(r'$M_{max, t}$', size = 14)
plt.show()
# -
| Change point detection methods demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Uber Data Analysis
# # Abstract
# Here we will be working with Uber datset.This dataset contains approximately 2 million trips. In first section I will describe the project and then mention the research wquestions and finally we will report our conclusions.
# ## Overview
#
# In this project we would like to understand customer trends in Uber data. Uber is one of the most popular transportantion app. In 2018 they released this dataset /////
# ## Research Questions
#
# In this project I will try to understand what is the most common travel day in uber data.
# In this project I have chosen my research dataset as uber dataset. I found this subject interesting it will show travel time of uber driver.How many he/ she has traveled during weekdays or weekends why there is such difference between that period how it is possible to distinguish their difference. Why it is important to know for simple users like us to know the travel time of the uber driver?It will be helpful for researchers to do make some sort of conclusions about this datset which we have.They can make conclusions based on that do they need services like "Uber" more or they dont need it.
# It is important what days are time pick period for people. Transport department could make conclusions for themselves like what transport are mostly in need for customers.If they analyze and conclude they could figure out for themselves uber is quite popular and it is important for everybody and it will imporve transport ways in different features for transportation.
# Also it will help to give genral overwiev about the destination of our passangers what locations require more stops and what trasportations are more needed on those locations.
# I got 
# this image which shows uber movement throught the city .
#
While I was doing this research analysis I have used histograms statistical element in order to view my results.
# ## Methodology & Analysis
import pandas as pd # here we import necessary libraries
uber_df = pd.read_csv("data/uber.csv", parse_dates=['date'], infer_datetime_format= True).sample(n = 10000)
uber_df.shape
uber_df.info()
# ## EDA (Exploratory Data Analysis)
uber_df.head()
# - check columns and what they stands for
import matplotlib.pyplot as plt
# +
plt.hist(uber_df.mean_travel_time)
plt.title("Distribution of Mean Travel Time")
plt.draw()
# -
# - It looks like most of the mean travel times are around 1000 seconds.
plt.boxplot(uber_df.mean_travel_time)
plt.draw()
# - Median travel time is around 1000 seconds. There are some outliers after 25000
uber_df.time_of_week.value_counts()
# In the first look it might seem surprising that the number of rides in weekends are not more than the weekdays but note that weekdays include M,T,W, Th, F.
# Goal: Notice that this data don't include the days of the trip. By using Python we can figure out the days by using date column in this dataset.
uber_df.date.dayofweek
import datetime
d = datetime.datetime.strptime('2016-03-03', '%Y-%m-%d')
uber_df.info()
uber_df['day'] = uber_df.date.map(lambda x: x.dayofweek)
uber_df
uber_df.sourceid.value_counts()
# Location 208 -- is the most common departure place.
uber_df.dstid.value_counts()
uber_df.dstid.plot(kind = 'hist')
# Location with id==170 looks like the most common terminal destination
uber_df.day.value_counts()
# Looks like the most popular trip day is a Friday with 1908 trips.
import matplotlib.pyplot as plt
plt.hist(uber_df.mean_travel_time)
uber_df.trip.value_counts()[:10]
uber_df.trip
select_trips =uber_df.index.tolist()
select_trips
select_trips = uber_df.index.tolist()
# # Data Analysis
# +
plt.hist(uber_df.mean_travel_time)
plt.title("Distribution of Mean Trip")
plt.draw()
# -
# Wow! The distribution of the mean_travel time is not normal and it is skewed. We can see the peak is around 1000 seconds.
| Data601_HW_1_Report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/PredictiveIntelligenceLab/JAX-BO/blob/master/jaxbo_colab.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="d1kVcs6oFjGV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="5112f8b9-9252-4091-ca3d-1c775f3c33f1"
import os
from getpass import getpass
# %cd '/content/'
# %rm -rf JAX-BO
user = getpass('GitHub user')
password = getpass('<PASSWORD>')
os.environ['GITHUB_AUTH'] = user + ':' + password
# !git clone https://$GITHUB_AUTH@github.com/PredictiveIntelligenceLab/JAX-BO.git
# %cd JAX-BO
# !pip uninstall --yes jaxbo
# !python setup.py install
# + id="k5jAEKxJIuVU" colab_type="code" colab={}
import numpy as onp
import jax.numpy as np
from jax import random, vmap
from jax.config import config
config.update("jax_enable_x64", True)
from scipy.optimize import minimize
from pyDOE import lhs
import matplotlib.pyplot as plt
from matplotlib import rc
from scipy.interpolate import griddata
from jaxbo.models import GP
from jaxbo.utils import normalize, compute_w_gmm
from jaxbo.test_functions import *
onp.random.seed(1234)
# + id="GUylj6N2J4bo" colab_type="code" colab={}
# Define test function
f, p_x, dim, lb, ub = oakley()
# Problem settings
N = 5
noise = 0.0
options = {'kernel': 'RBF',
'criterion': 'LW-LCB',
'input_prior': p_x,
'kappa': 2.0,
'nIter': 20}
gp_model = GP(options)
# Domain bounds
bounds = {'lb': lb, 'ub': ub}
# Initial training data
X = lb + (ub-lb)*lhs(dim, N)
y = vmap(f)(X)
y = y + noise*y.std(0)*onp.random.normal(y.shape)
# Test data
if dim == 1:
create_plots = True
nn = 1000
X_star = np.linspace(lb[0], ub[0], nn)[:,None]
y_star = vmap(f)(X_star)
elif dim == 2:
create_plots = True
nn = 80
xx = np.linspace(lb[0], ub[0], nn)
yy = np.linspace(lb[1], ub[1], nn)
XX, YY = np.meshgrid(xx, yy)
X_star = np.concatenate([XX.flatten()[:,None],
YY.flatten()[:,None]], axis = 1)
y_star = vmap(f)(X_star)
else:
create_plots = False
nn = 20000
X_star = lb + (ub-lb)*lhs(dim, nn)
y_star = vmap(f)(X_star)
# True location of global minimum
idx_true = np.argmin(y_star)
true_x = X_star[idx_true,:]
true_y = y_star.min()
dom_bounds = tuple(map(tuple, np.vstack((lb, ub)).T))
result = minimize(f, true_x, jac=None, method='L-BFGS-B', bounds = dom_bounds)
true_x, true_y = result.x, result.fun
# + id="EnVZ3wB8Wws5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="9a4c2b03-5369-4cf5-b6a1-f2c9d57ab131"
# Main Bayesian optimization loop
rng_key = random.PRNGKey(0)
for it in range(options['nIter']):
print('-------------------------------------------------------------------')
print('------------------------- Iteration %d/%d -------------------------' % (it+1, options['nIter']))
print('-------------------------------------------------------------------')
# Fetch normalized training data
norm_batch, norm_const = normalize(X, y, bounds)
# Train GP model
print('Train GP...')
rng_key = random.split(rng_key)[0]
opt_params = gp_model.train(norm_batch,
rng_key,
num_restarts = 10)
# Fit GMM
if options['criterion'] == 'LW-LCB' or options['criterion'] == 'LW-US':
print('Fit GMM...')
rng_key = random.split(rng_key)[0]
kwargs = {'params': opt_params,
'batch': norm_batch,
'norm_const': norm_const,
'bounds': bounds,
'rng_key': rng_key}
gmm_vars = gp_model.fit_gmm(**kwargs, N_samples = 10000)
else:
gmm_vars = None
# Compute next point via minimizing the acquisition function
print('Computing next acquisition point...')
kwargs = {'params': opt_params,
'batch': norm_batch,
'norm_const': norm_const,
'bounds': bounds,
'gmm_vars': gmm_vars}
new_X = gp_model.compute_next_point(num_restarts=10, **kwargs)
# Acquire data
new_y = vmap(f)(new_X)
new_y = new_y + noise*new_y.std(0)*onp.random.normal(new_y.shape)
# Augment training data
print('Updating data-set...')
X = np.concatenate([X, new_X], axis = 0)
y = np.concatenate([y, new_y], axis = 0)
# Print current best
idx_best = np.argmin(y)
best_x = X[idx_best,:]
best_y = y.min()
print('True location: ({}), True value: {}'.format(true_x, true_y))
print('Best location: ({}), Best value: {}'.format(best_x, best_y))
print('New location: ({}), New value: {}'.format(new_X, new_y))
# + id="x5pYL0MuQD44" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a0dd9e24-d8d4-4184-b09a-eae6fd7058dd"
# Test accuracy
mean, std = gp_model.predict(X_star, **kwargs)
lower = mean - 2.0*std
upper = mean + 2.0*std
# Check accuracy
error = np.linalg.norm(mean-y_star,2)/np.linalg.norm(y_star,2)
print("Relative L2 error u: %e" % (error))
# + id="ebJ1P5LzW1ka" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 638} outputId="62fba00a-5077-4d88-ad37-7ea9f5653fee"
if create_plots:
# Compute predictions
if options['criterion'] == 'LW-LCB' or options['criterion'] == 'LW-US':
w_pred = compute_w_gmm(X_star, **kwargs)
else:
w_pred = np.zeros(X_star.shape[0])
acq_fun = lambda x: gp_model.acquisition(x, **kwargs)
a_pred = vmap(acq_fun)(X_star)
x_new = gp_model.compute_next_point(num_restarts=10, **kwargs)
# Convert to NumPy
X_star = onp.array(X_star)
y_star = onp.array(y_star)
mean = onp.array(mean)
std = onp.array(std)
w_pred = onp.array(w_pred)
a_pred = onp.array(a_pred)
XX = onp.array(XX)
YY = onp.array(YY)
Y_star = griddata(X_star, y_star, (XX, YY), method='cubic')
Y_pred = griddata(X_star, mean, (XX, YY), method='cubic')
Y_std = griddata(X_star, std, (XX, YY), method='cubic')
W_star = griddata(X_star, w_pred, (XX, YY), method='cubic')
A_star = griddata(X_star, a_pred, (XX, YY), method='cubic')
# Plot
plt.rcParams.update({'font.size': 16})
plt.rcParams['axes.linewidth']=3
plt.figure(figsize = (16,8))
plt.subplot(1, 4, 1)
fig = plt.contourf(XX, YY, Y_star)
plt.plot(X[:,0], X[:,1], 'r.', ms = 6, alpha = 0.8)
# plt.plot(true_x[0], true_x[1], 'md', ms = 8, alpha = 1.0)
plt.xlabel(r'$x_1$')
plt.ylabel(r'$x_2$')
plt.title(r'Exact u(x)')
plt.axis('square')
plt.subplot(1, 4, 2)
fig = plt.contourf(XX, YY, Y_pred)
plt.xlabel(r'$x_1$')
plt.ylabel(r'$x_2$')
plt.title(r'Predicted mean')
plt.axis('square')
plt.subplot(1, 4, 3)
fig = plt.contourf(XX, YY, 2.0*Y_std)
plt.xlabel(r'$x_1$')
plt.ylabel(r'$x_2$')
plt.title(r'Two stds')
plt.axis('square')
plt.subplot(1, 4, 4)
fig = plt.contourf(XX, YY, np.abs(Y_star-Y_pred))
plt.xlabel(r'$x_1$')
plt.ylabel(r'$x_2$')
plt.title(r'Absolute error')
plt.axis('square')
plt.savefig('function_prediction.png', dpi = 300)
idx_max = np.argmin(a_pred)
plt.figure(figsize = (12,5))
plt.subplot(1, 2, 1)
fig = plt.contourf(XX, YY, W_star)
plt.xlabel(r'$x_1$')
plt.ylabel(r'$x_2$')
plt.title(r'$w_{GMM}(x)$')
plt.axis('square')
plt.subplot(1, 2, 2)
fig = plt.contourf(XX, YY, A_star)
plt.colorbar(fig)
# plt.plot(x0[:,0], x0[:,1], 'ms')
# plt.plot(X_star[idx_max,0], X_star[idx_max,1], 'md')
plt.plot(x_new[:,0], x_new[:,1], 'md', label = 'new X')
plt.legend(frameon = False)
plt.xlabel(r'$x_1$')
plt.ylabel(r'$x_2$')
plt.title(r'%s(x)' % (options['criterion']))
plt.axis('square')
plt.savefig('acquisition.png', dpi = 300)
| jaxbo_colab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
## Read polygons from file and apply as gee geometries
import fiona
from shapely.geometry import shape, MultiPolygon, MultiPoint, Polygon, Point
import glob
# current working directory
import os
cwd = os.getcwd()
# find files in shapefile folder
shape_file_path = '/media/chris/Data/GreenCityWatch/data/Ground_truth/*.shp'
shape_files = glob.glob(shape_file_path)
path = shape_files[1]
shape_files, cwd, path
# -
shp_AOIs = MultiPoint([shape(pol['geometry']) for pol in fiona.open(path)])
len(shp_AOIs)
from pprint import pprint
# +
diameters = []
for point in fiona.open(path):
diameter = point['properties']['diameter']
diameters.append(diameter)
# -
import numpy as np
diameters = np.array(diameters)
# +
import matplotlib.pyplot as plt
plt.hist(diameters[diameters < 94])
# -
plt.boxplot(diameters < 100)
plt.hist(diameters[diameters != 0])
| misc/Get_tree_inventory_info.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib
#matplotlib.use('Agg') # necessary for linux kernal
# %matplotlib inline
import matplotlib.pyplot as plt
import os
import numpy as np
import pandas as pd
from keras import models
from keras.models import load_model
from keras.layers import Input
from keras.models import Model
from mapgenlib.eval import readImg, rescaleImg, model_predict, save_prediction
def update_gan_generator_to_any_size(old_model):
# Remove the top layer and add input with no limit
old_model.layers.pop(0)
newInput = Input(shape=(None, None, 1)) # New image input
newOutputs = old_model(newInput)
newModel = Model(newInput, newOutputs)
return newModel
def predict_long_image(SavedModel, testPath, fn_input, out_path, scale, nr, overlap = 160):
image_arr = readImg(testPath + fn_input)
collect = np.zeros(image_arr.shape)
for i in range(int(image_arr.shape[1] / 2000)):
img_range = [0,4000,2000 * i - overlap,2000 * (i+1) + overlap]
img_insert = [0,4000,2000 * i,2000 * (i+1)]
if len(img_range) == 4: # If range was set
xmin, xmax, ymin, ymax = img_range
if ymin < 0:
ymin = 0
if ymax > image_arr.shape[1]:
ymax = image_arr.shape[1]
print(xmin, xmax, ymin, ymax)
image_sub = image_arr[xmin:xmax, ymin:ymax]
image_sub = rescaleImg(image_sub)
xmin_, xmax_, ymin_, ymax_ = img_insert
print(ymin_-ymin, ymax- ymax_)
pred_sub = model_predict(SavedModel, image_sub, num_runs = nr)
#if i%2==1:
# pred_sub = np.logical_not(pred_sub)
a,b = ymin_-ymin, ymin_-ymin + 2000 + ymax -ymax
collect[xmin_:xmax_, ymin_:ymax_] = pred_sub[:, a:b]
save_prediction(collect, out_path, fn_input[:-4], '_' + str(scale) + '_' + str(nr) + '_out.png')
# -
if 1: # Final Residual U-net with l2 loss and Adam 0.0004
#scale = 25
#modelPath = '../tmp_data/predictions/2019-02-28 11-05-31_25/'
#out_evaluation = r"../tmp_data/Evaluations/Runet_25k/"
#scale = 15
#modelPath = '../tmp_data/predictions/2019-02-26 12-55-00_15/'
#out_evaluation = r"../tmp_data/Evaluations/Runet_15k/"
scale = 10
modelPath = '../tmp_data/predictions/U128_2019-03-10 12-26-47_10/'
out_evaluation = r"../tmp_data/Evaluations/Runet_10k/"
modelname = "weights.h5"
saved_model = load_model(modelPath + modelname)
saved_model = update_gan_generator_to_any_size(saved_model)
tester_path = '../tmp_data/building/'
fn = r"big_hameln_label_50cm.tif"
predict_long_image(saved_model,tester_path, fn, out_evaluation, scale, nr = 1)
predict_long_image(saved_model,tester_path, fn, out_evaluation, scale, nr = 2)
| Evaluate_Hameln_20190610.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import glob
# +
# Read file into string
with open('data/1OLG.pdb', 'r') as f:
f_str = f.read()
# Let's look at the first 1000 characters
f_str[:1000]
# +
# Read contents of the file in as a list
with open('data/1OLG.pdb', 'r') as f:
f_list = f.readlines()
# Look at the list (first ten entries)
f_list[:10]
# -
# !head data/1OLG.pdb
#Print the first ten lines of the file
with open('data/1OLG.pdb', 'r') as f:
for i, line in enumerate(f):
print(line.rstrip())
if i >= 10:
break
# Print the first ten lines of the file
with open('data/1OLG.pdb', 'r') as f:
i = 0
while i < 10:
print(f.readline().rstrip())
i += 1
os.path.isfile('data/1OLG.pdb')
# +
if os.path.isfile('yogi.txt'):
raise RuntimeError('File yogi.txt already exists.')
with open('yogi.txt', 'w') as f:
f.write('When you come to a fork in the road, take it.')
f.write('You can observe a lot by just watching.')
f.write('I never said most of the things I said.')
# -
# !cat yogi.txt
with open('yogi.txt', 'w') as f:
f.write('When you come to a fork in the road, take it.\n')
f.write('You can observe a lot by just watching.\n')
f.write('I never said most of the things I said.\n')
# This will result in an exception
with open('gimme_phi.txt', 'w') as f:
f.write('The golden ratio is φ = ')
f.write(1.61803398875)
with open('gimme_phi.txt', 'w') as f:
f.write('The golden ratio is φ = ')
f.write('{phi:.8f}'.format(phi=1.61803398875))
# !cat gimme_phi.txt
with open('data/1OLG.pdb', 'r') as f, open('atoms_chain_A.txt', 'w') as f_out:
# Put the ATOM lines from chain A in new file
for line in f:
if len(line) > 21 and line[:4] == 'ATOM' and line[21] == 'A':
f_out.write(line)
# !head -10 atoms_chain_A.txt
# !tail -10 atoms_chain_A.txt
# +
file_list = glob.glob('data/*.pdb')
file_list
# +
# Dictionary to hold sequences
seqs = {}
# Loop through all matching files
for file_name in file_list:
# Extract PDB ID
pdb_id = file_name[file_name.find('/')+1:file_name.rfind('.')]
# Initialize sequence string, which we build as we go along
seq = ''
with open(file_name, 'r') as f:
for line in f:
if len(line) > 11 and line[:6] == 'SEQRES' and line[11] == 'A':
seq += line[19:].rstrip() + ' '
# Build sequence with dash-joined three letter codes
seq = '-'.join(seq.split())
# Store in the dictionary
seqs[pdb_id] = seq
# -
| lesson_11.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ejercicio Ridge, Lasso y ElasticNet
# Para este ejercicio vas a trabajar con datos de ventas de una empresa que fabrica muebles en España. Esta empresa es una Pyme que demanda conocer de antemano qué ingresos va a tener cada mes, ya que necesita pedir un anticipio a su entidad financiera de cara a poder abastecerse de materia prima al comenzar el mes. Como desconoce cuánto tiene que pedir, tira al alza y acaba pagando muchos intereses. El objetivo es reducir estos gastos.
#
# En las bases de datos de la empresa constan todos los gastos en publicidad y ventas, para cada uno de los meses desde su fundación (hace más de 15 años).
#
# Dado que los presupuestos de marketing se cierran al principio de cada mes, la empresa necesita un modelo predictivo que le anticipe las ventas que conseguirá a final de mes en función de los gastos que realizará en marketing.
#
# Para ello tendrás que utilizar tanto modelos de regresión normales, como regularizados.
#
# 1. Carga los datos y realiza un pequeño análisis exploratorio. Mira a ver cómo se relaciona las ventas con el resto de variables.
# 2. Crea varios modelos y modifica los hiperparámetros necesarios para mejorar el performance del modelo.
# ## Import libraries
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import warnings
warnings.filterwarnings("ignore")
# -
# ## Exploratory Data Analysis
# +
DATAPATH = 'data/Advertising.csv'
data = pd.read_csv(DATAPATH)
data.drop(columns=['Unnamed: 0'], inplace=True)
data.head()
# -
len(data)
data.columns
def scatter_plot(feature, target):
plt.figure(figsize=(16, 8))
plt.scatter(
data[feature],
data[target],
c='black'
)
plt.xlabel("Money spent on {} ads ($)".format(feature))
plt.ylabel("Sales")
plt.show()
scatter_plot('TV', 'sales')
scatter_plot('radio', 'sales')
scatter_plot('newspaper', 'sales')
data.corr()
import seaborn as sns
sns.distplot(data['sales']);
# ## Modelling
# +
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
X_train, X_test, y_train, y_test = train_test_split(data.drop(columns=['sales']),
data['sales'],
test_size = 0.20,
random_state=42)
# -
# ### Multiple linear regression
# +
from sklearn.linear_model import LinearRegression
lin_reg = LinearRegression()
lin_reg.fit(X_train, y_train)
print("MSE: ", mean_squared_error(y_test, lin_reg.predict(X_test)))
print("RMSE: ", np.sqrt(mean_squared_error(y_test, lin_reg.predict(X_test))))
# -
# ### Ridge regression
# +
from sklearn.linear_model import Ridge
ridge = Ridge(alpha=20)
ridge.fit(X_train, y_train)
print("MSE: ", mean_squared_error(y_test, ridge.predict(X_test)))
print("RMSE: ", np.sqrt(mean_squared_error(y_test, ridge.predict(X_test))))
# +
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import Ridge
alpha = [1e-15, 1e-10, 1e-8, 1e-4, 1e-3,1e-2, 1, 5, 10, 20]
ridge = Ridge()
parameters = {'alpha': [1e-15, 1e-10, 1e-8, 1e-4, 1e-3,1e-2, 1, 5, 10, 20]}
ridge_regressor = GridSearchCV(ridge, parameters,scoring='neg_mean_squared_error', cv=5)
ridge_regressor.fit(X_train, y_train)
# -
ridge_regressor.best_params_
-ridge_regressor.best_score_
# ### Lasso
# +
from sklearn.linear_model import Lasso
lasso = Lasso(alpha=1)
lasso.fit(X_train, y_train)
print("MSE: ", mean_squared_error(y_test, lasso.predict(X_test)))
print("RMSE: ", np.sqrt(mean_squared_error(y_test, lasso.predict(X_test))))
# +
from sklearn.linear_model import Lasso
lasso = Lasso()
parameters = {'alpha': [1e-15, 1e-10, 1e-8, 1e-4, 1e-3,1e-2, 1, 5, 10, 20]}
lasso_regressor = GridSearchCV(lasso, parameters, scoring='neg_mean_squared_error', cv = 5)
lasso_regressor.fit(X_train, y_train)
# -
lasso_regressor.best_params_
-lasso_regressor.best_score_
# ### ElasticNet
# +
from sklearn.linear_model import ElasticNet
elastic = ElasticNet(alpha=100, l1_ratio= 0.99)
elastic.fit(X_train, y_train)
print("MSE: ", mean_squared_error(y_test, elastic.predict(X_test)))
print("RMSE: ", np.sqrt(mean_squared_error(y_test, elastic.predict(X_test))))
# +
# %%time
from sklearn.linear_model import ElasticNet
elastic = ElasticNet()
parameters = {'alpha': [1e-5, 1e-4, 1e-3, 1e-2, 1e-1, 0.0, 1.0, 10.0, 100.0],
'l1_ratio': np.arange(0, 1, 0.01)}
elastic_regressor = GridSearchCV(elastic, parameters, scoring='neg_mean_squared_error', cv = 5)
elastic_regressor.fit(X_train, y_train)
# -
elastic_regressor.best_params_
-elastic_regressor.best_score_
# ### StandardScaler
# +
# %%time
from sklearn.linear_model import ElasticNet
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(X_train)
X_train_scaled = scaler.transform(X_train)
elastic = ElasticNet()
parameters = {'alpha': [1e-5, 1e-4, 1e-3, 1e-2, 1e-1, 0.0, 1.0, 10.0, 100.0],
'l1_ratio': np.arange(0, 1, 0.01)}
elastic_regressor = GridSearchCV(elastic, parameters, scoring='neg_mean_squared_error', cv = 5)
elastic_regressor.fit(X_train_scaled, y_train)
# -
-elastic_regressor.best_score_
elastic_regressor.coefs
| Bloque 3 - Machine Learning/01_Supervisado/3-Regularizacion/ejercicios/01_RESU_Advertising Ridge Lasso.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # MIDI to CSV Converter
import mido
import csv
import os
def cprint(s,b):
if b:
print(s)
def get_active_notes(mid):
active_notes = {}
tracks = mid.tracks
num_tracks = len(tracks)
all_notes = num_tracks * [None]
for i in range(0, num_tracks):
track = tracks[i]
time = 0
all_notes[i] = []
for msg in track:
msg_dict = msg.dict()
time += msg_dict['time']
if msg.type == 'note_on' or msg.type == 'note_off':
vel = msg_dict['velocity']
if vel > 0 and msg.type == 'note_on':
# Using a list for the active notes becuase note 71 in io.mid was definied twice at once
if active_notes.has_key(msg_dict['note']):
active_notes[msg_dict['note']].append({'time':time,'velocity':vel})
else:
active_notes[msg_dict['note']] = [{'time':time, 'velocity': vel}]
elif vel == 0 or msg.type == 'note_off':
note = msg_dict['note']
if len(active_notes[note])>0:
start_msg = active_notes[note].pop()
new_note = {'note': note, 'start': start_msg['time'],
'end': time, 'velocity': start_msg['velocity']}
all_notes[i].append(new_note)
return all_notes
def create_streams(all_notes):
streams = []
for notes in all_notes:
while notes != []:
stream = []
vel = 0
current_end = 0
for note in notes:
if note['start'] >= current_end:
if note['velocity'] != vel:
vel = note['velocity']
else:
del note['velocity']
stream.append(note)
current_end = note['end']
streams.append(stream)
for note in stream:
notes.remove(note)
return streams
midiNotes = ['C','C#','D','D#','E','F','F#','G','G#','A','A#','B']
def midi2str(midi):
return midiNotes[midi%12] + str(midi/12 -1)
def streams_to_cells(streams, speed, printing):
max_time = int(max([x['end'] for x in [item for sublist in streams for item in sublist]]))+1
start_cells = 'A2:A' + str(1+len(streams))
instructions = 'r m' + str(max_time-1)
turtles = [['!turtle(' + start_cells + ', ' + instructions + ', ' + str(speed) + ', 1)']]
for stream in streams:
cells = [""] * max_time
for note in stream:
start = int(note['start'])
cells[start] = midi2str(note['note'])
if note.has_key('velocity'):
cells[start] += (' ' + str(round(float(note['velocity'])/127,2)))
for rest_duration in range(1,int(note['length'])):
cells[start+rest_duration] = '-'
turtles.append(cells)
cprint(str(len(turtles)) + ' x ' + str(max([len(stream) for stream in turtles])), printing)
return turtles
def midi_to_excello(file_name, method=1, logging=False, printing=True):
# Fetch MIDI file
mid = mido.MidiFile(file_name)
tempo = [m.dict()['tempo'] for m in mid.tracks[0] if m.dict().has_key('tempo')][0]
ticks_per_beat = mid.ticks_per_beat
# Extract the notes from as onset, note, offset, volume from messages
all_notes = get_active_notes(mid)
# Split into the streams as played by individual turtles
streams = create_streams(all_notes)
all_notes = [item for sublist in streams for item in sublist]
cprint('Number of turtles: ' + str(len(streams)), printing)
# No Compression
if method == 0:
cprint("No Compression", printing)
difference_stat = 1
ratio_int = 1
for stream in streams:
for note in stream:
note['length'] = note['end'] - note['start']
#Compression
else:
differences = [(y['start']-x['start']) for x, y in zip(all_notes[:-1], all_notes[1:])]
lengths = [(x['end'] - x['start']) for x in [item for sublist in streams for item in sublist]]
# Mins
if method == 1:
cprint("Min Compression", printing)
difference_stat = min([x for x in differences if x > 1])
length_stat = min([x for x in lengths if x > 1])
# Modes
elif method == 2:
cprint("Mode Compression", printing)
difference_stat = max(set(differences), key=differences.count)
length_stat = max(set(lengths), key=lengths.count)
cprint('note difference stat: ' + str(difference_stat), printing)
cprint('note length stat: ' + str(length_stat), printing)
mode_ratio = (float(max(difference_stat, length_stat)) / min(difference_stat, length_stat))
cprint('mode ratio: ' + str(mode_ratio), printing)
ratio_int = int(mode_ratio)
cprint('integer ratio: ' + str(ratio_int), printing)
# ratio_correction = mode_ratio/ratio_int
# cprint('ratio correction: ' + str(ratio_correction), printing)
# Convert MIDI times to cell times
rounding_base = 0.1
for stream in streams:
for note in stream:
note['length'] = ((float(note['end']) - note['start'])/length_stat)
note['length'] = rounding_base * round(note['length']/rounding_base)
note['start'] = round(rounding_base * round((float(note['start'])/difference_stat*ratio_int)/rounding_base))
note['end'] = note['start'] + note['length']
speed = int(round((float(60*10**6)/tempo) * ticks_per_beat * (float(ratio_int)/difference_stat)))
cprint(speed, printing)
csv_name = file_name[::-1].replace('/','_',file_name.count('/')-2)[::-1]
csv_name = csv_name.replace('/midi','/csv/' + str(method)).replace('.mid','.csv')
with open(csv_name, "wb") as f:
writer = csv.writer(f)
writer.writerows(streams_to_cells(streams, speed, printing))
cprint("Written to " + csv_name, printing)
if logging:
cprint([csv_name, len(streams), int(max([x['end'] for x in [item for sublist in streams for item in sublist]]))], printing)
return [csv_name, len(streams), int(max([x['end'] for x in [item for sublist in streams for item in sublist]]))]
# # Converting
# 0: No Compression<br>
# 1: Compression using Minimum difference<br>
# 2: Compression using Modal difference
midi_to_excello('musescore/midi/Flute_Salad.mid', 1)
# # Corpus Conversion
datasets = ['piano-midi', 'bach', 'bach_chorales']
def convert_corpus(corpus, method):
midi_files = corpus + '/midi'
files = []
for r, _, f in os.walk(midi_files):
for file in f:
if '.mid' in file or '.MID' in file:
files.append(os.path.join(r, file))
if midi_files == 'bach/midi':
files.remove('bach/midi/suites/airgstr4.mid')
files = [ x for x in files if "wtcbki/" not in x ]
log = []
for f in files:
log.append(midi_to_excello(f, method, logging=True, printing=False)) # This also writes the file to disk.
log.sort(key=lambda x: x[2], reverse=False)
with open(midi_files.replace('/midi','/csv') + '/' + 'log' + str(method) + '.txt', mode="w") as outfile:
outfile.write('%s\n'% len(log))
for s in log:
outfile.write("%s\n" % s)
for corpus in datasets:
for method in [0,1,2]:
print(corpus, method)
convert_corpus(corpus, method)
# # MIDI note name conversion test
import audiolazy
for i in range(12,120):
print(audiolazy.midi2str(i) == midi2str(i))
| MIDI/MIDI_Conversion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/vasiliyeskin/bentrevett-pytorch-seq2seq_ru/blob/master/6%20-%20Attention%20is%20All%20You%20Need.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="sK-p0weReC_2"
# # 6 - Attention is All You Need
#
# В этом разделе мы будем реализовывать (слегка измененную версию) модели Transformer из статьи [Attention is All You Need](https://arxiv.org/abs/1706.03762). Все изображения в этой части взяты из статьи Transformer. Для получения дополнительной информации о Transformer обращайтесь [сюда](https://www.mihaileric.com/posts/transformers-attention-in-disguise/), [сюда](https://jalammar.github.io/illustrated-transformer/) [и сюда](http://nlp.seas.harvard.edu/2018/04/03/attention.html). На русском языке [здесь](https://habr.com/ru/post/486358/).
#
# 
#
# ## Введение
#
# Подобно свёрточной модели Sequence-to-Sequence, Transformer не использует никакой рекуррентности. Он также не использует свёрточные слои. Вместо этого модель полностью состоит из линейных слоев, механизмов внимания и нормализации.
#
# По состоянию на январь 2020 года трансформеры являются доминирующей архитектурой в NLP и используются для достижения передовых результатов во многих задач, и похоже, что они будут доминировать в ближайшем будущем в области обработки языков.
#
# Самый популярный Transformer вариант это [BERT](https://arxiv.org/abs/1810.04805) (**B**idirectional **E**ncoder **R**epresentations from **T**ransformers) и предварительно обученные версии BERT обычно используются для замены слоёв эмбеддинга — если не больше - в NLP моделях.
#
# Распространенной библиотекой, используемой при работе с предварительно обученными трансформаторами, является библиотека [Transformers](https://huggingface.co/transformers/), смотрите [здесь](https://huggingface.co/transformers/pretrained_models.html) список всех доступных предварительно обученных моделей.
#
# Различия между реализацией в этой части и в статье:
# - мы используем обученную позиционную кодировку вместо статической
# - мы используем стандартный оптимизатор Adam со статической скоростью обучения вместо оптимизатора с динамически изменяющейся скоростью
# - мы не используем сглаживание меток
#
# Мы вносим все эти изменения, поскольку они близки к настройкам BERT и большинство вариантов Transformer используют аналогичную настройку.
# + [markdown] id="Igqzg51reC_3"
# ## Подготовка данных
#
# Как всегда, давайте импортируем все необходимые модули и зададим случайные начальные числа для воспроизводимости.
# + id="Ap3d_civeC_4"
import torch
import torch.nn as nn
import torch.optim as optim
import torchtext
from torchtext.legacy.datasets import Multi30k
from torchtext.legacy.data import Field, BucketIterator, TabularDataset
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import spacy
import numpy as np
import random
import math
import time
# + id="l9vyd9nIeC_5"
SEED = 1234
random.seed(SEED)
np.random.seed(SEED)
torch.manual_seed(SEED)
torch.cuda.manual_seed(SEED)
torch.backends.cudnn.deterministic = True
# + [markdown] id="Q0qCExV1eC_6"
# Для загрузки в Google Colab используем следующие команды (После загрузки обязательно перезапустите colab runtime! Наибыстрейший способ через короткую комаду: **Ctrl + M + .**):
# + pycharm={"name": "#%%\n"} id="dRk6kScVeC_6"
# !pip install -U spacy==3.0
# !python -m spacy download en_core_web_sm
# !python -m spacy download de_core_news_sm
# + [markdown] id="JpvSKHHteC_7"
# Затем мы создадим наши токенизаторы, как и раньше.
# + id="2cb9Q_txeC_8"
spacy_de = spacy.load('de_core_news_sm')
spacy_en = spacy.load('en_core_web_sm')
# + id="-dBOibhzeC_8"
def tokenize_de(text):
"""
Tokenizes German text from a string into a list of strings
"""
return [tok.text for tok in spacy_de.tokenizer(text)]
def tokenize_en(text):
"""
Tokenizes English text from a string into a list of strings
"""
return [tok.text for tok in spacy_en.tokenizer(text)]
def tokenize_text(text):
"""
Tokenizes English text from a string into a list of strings
"""
return [tok.text for tok in text.split(' ')]
# + [markdown] id="RyN4hrAGeC_9"
# Наши поля такие же, как и в предыдущих частях. Модель ожидает, что данные будут введены в первую очередь с размерностью батча, поэтому мы используем`batch_first = True`.
# + id="UKVJOamteC_-"
SRC = Field(tokenize = tokenize_de,
init_token = '<sos>',
eos_token = '<eos>',
lower = True,
batch_first = True)
TRG = Field(tokenize = tokenize_en,
init_token = '<sos>',
eos_token = '<eos>',
lower = True,
batch_first = True)
# + [markdown] id="ijABmpNPeC__"
# Затем мы загружаем набор данных Multi30k и создаем словарь.
# + id="H_hyhnU8eC__" outputId="a6f5039e-704d-4189-f896-f224d0b75173" colab={"base_uri": "https://localhost:8080/"}
train_data, valid_data, test_data = Multi30k.splits(exts = ('.de', '.en'),
fields = (SRC, TRG))
# + id="LXe-QUJBeDAA"
SRC.build_vocab(train_data, min_freq = 2)
TRG.build_vocab(train_data, min_freq = 2)
# + [markdown] id="URpN4-WIeDAA"
# Наконец, мы определяем устройство для обучения и итератор данных.
# + id="b3ujBl4beDAA"
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# + id="-MZp0w0heDAA"
BATCH_SIZE = 128
train_iterator, valid_iterator, test_iterator = BucketIterator.splits(
(train_data, valid_data, test_data),
batch_size = BATCH_SIZE,
device = device)
# + [markdown] id="sgInf_6_eDAB"
# ## Построение модели
#
# Далее мы построим модель. Как и в предыдущих частях, она состоит из *кодера* и *декодера*, с кодером *кодирующим* входное предложение (на немецком языке) в *вектор контекста* и декодера, который *декодирует* этот вектор контекста в выходное предложение (на английском языке).
#
# ### Кодировщик
#
# Подобно модели ConvSeq2Seq, кодировщик Transformer не пытается сжать всё исходное предложение $X = (x_1, ... ,x_n)$ в единый вектор контекста $z$. Вместо этого он создает последовательность векторов контекста $Z = (z_1, ... , z_n)$. Итак, если бы наша входная последовательность состояла из 5 токенов, у нас было бы $Z = (z_1, z_2, z_3, z_4, z_5)$. Почему мы называем этот тензор последовательностью контекстных векторов, а не последовательностью скрытых состояний? Скрытое состояние в момент времени $t$ в RNN видит только токены x_t и все токены, что были перед ним. Однако здесь каждый вектор контекста `видел` все токены во всех позициях входной последовательности.
#
# 
#
# Сначала токены проходят через стандартный слой эмбеддинга. Поскольку модель не является рекуррентной, она не имеет представления о порядке токенов в последовательности. Мы решаем эту проблему, используя второй слой эмбеддинга, называемый *позиционный слой эмбеддинга* (positional embedding layer). Это стандартный эмбеддинг, для которого входом является не сам токен, а позиция токена в последовательности, начиная с первого токена, токена `<sos>` (начало последовательности) в позиции 0. Позиционный эмбеддинг имеет размер словаря, равный 100, что означает, что наша модель может принимать предложения длиной до 100 токенов. Его можно увеличить, если мы хотим обрабатывать более длинные предложения.
#
# Оригинальная реализация Transformer в статье Attention is All You Need не обучала позиционный эмбеддинг. Вместо этого в ней использовался фиксированный статический эмбеддинг. Современные архитектуры Transformer, такие как BERT, вместо этого используют позиционные эмбеддинги, поэтому мы решили использовать их в этой реализации. Обратитесь [сюда](http://nlp.seas.harvard.edu/2018/04/03/attention.html#positional-encoding), чтобы узнать больше о позиционных эмбеддингах, используемых в исходной модели Transformer.
#
# Затем токен и результат прохождения позиционного эмбеддинга поэлементно суммируются для получения вектора, который содержит информацию о токене, а также его позицию в последовательности. Однако перед суммированием токенов эмбеддинга они умножаются на коэффициент масштабирования, равный $\sqrt{d_{model}}$, где $d_{model}$ размер скрытого измерения `hid_dim`. Это якобы уменьшает дисперсию эмбеддинга, и модель трудно обучить без этого коэффициента масштабирования. Затем применяется дропаут для комбинированного эмебеддинга.
#
# Комбинированный эмебеддинг затем пропускаются через $N$ *слоев кодировщика* для получения $Z$, для вывода и использования декодером.
#
# Исходная маска `src_mask` просто имеет ту же форму, что и исходное предложение, но имеет значение 1, когда токен в исходном предложении не является токеном `<pad>` и 0, когда это токен `<pad>`. Это используется в слоях кодировщика для маскировки механизмов многонаправленного внимания, которые используются для вычисления и применения внимания к исходному предложению, поэтому модель не обращает внимания на токены `<pad>`, которые не содержат полезной информации.
# + id="gPdb3zxNeDAC"
class Encoder(nn.Module):
def __init__(self,
input_dim,
hid_dim,
n_layers,
n_heads,
pf_dim,
dropout,
device,
max_length = 100):
super().__init__()
self.device = device
self.tok_embedding = nn.Embedding(input_dim, hid_dim)
self.pos_embedding = nn.Embedding(max_length, hid_dim)
self.layers = nn.ModuleList([EncoderLayer(hid_dim,
n_heads,
pf_dim,
dropout,
device)
for _ in range(n_layers)])
self.dropout = nn.Dropout(dropout)
self.scale = torch.sqrt(torch.FloatTensor([hid_dim])).to(device)
def forward(self, src, src_mask):
#src = [batch size, src len]
#src_mask = [batch size, 1, 1, src len]
batch_size = src.shape[0]
src_len = src.shape[1]
pos = torch.arange(0, src_len).unsqueeze(0).repeat(batch_size, 1).to(self.device)
#pos = [batch size, src len]
src = self.dropout((self.tok_embedding(src) * self.scale) + self.pos_embedding(pos))
#src = [batch size, src len, hid dim]
for layer in self.layers:
src = layer(src, src_mask)
#src = [batch size, src len, hid dim]
return src
# + [markdown] id="T09NNS4VeDAC"
# ### Слой кодировщика
#
# Слои кодировщика — это место, содержащее всю «соль» кодировщика. Сначала мы передаем исходное предложение и его маску в *слой многонаправленного внимания*, затем выполняем дропаут для его выхода, применяем остаточное соединение и передайте его через [слой нормализации](https://arxiv.org/abs/1607.06450). Затем результат мы пропускаем через слой *сети позиционно-зависимого прямого распространения* и снова применяем дропаут, остаточное соединение, слой нормализации, чтобы получить вывод этого слоя, который передается на следующий слой. Параметры не разделяются (не являются общими) между слоями.
#
# Слой многонаправленного внимания используется уровнем кодировщика для сосредоточения внимания на исходном предложению, то есть он вычисляет и применяет механизм внимание по отношению к себе, а не к другой последовательности, поэтому эту процедуру называются *внутренним вниманием*.
#
# [Эта](https://mlexplained.com/2018/01/13/weight-normalization-and-layer-normalization-explained-normalization-in-deep-learning-part-2/) статья подробно рассказывает о нормализации слоев. Суть в том, что в процедуре нормализации нормализуются значения признаков (то есть по скрытым измерениям), поэтому каждый признак имеет среднее значение 0 и стандартное отклонение 1. Это упрощает обучение нейронных сетей с большим количеством слоев, таких как Transformer.
# + id="35GS1I7ceDAD"
class EncoderLayer(nn.Module):
def __init__(self,
hid_dim,
n_heads,
pf_dim,
dropout,
device):
super().__init__()
self.self_attn_layer_norm = nn.LayerNorm(hid_dim)
self.ff_layer_norm = nn.LayerNorm(hid_dim)
self.self_attention = MultiHeadAttentionLayer(hid_dim, n_heads, dropout, device)
self.positionwise_feedforward = PositionwiseFeedforwardLayer(hid_dim,
pf_dim,
dropout)
self.dropout = nn.Dropout(dropout)
def forward(self, src, src_mask):
#src = [batch size, src len, hid dim]
#src_mask = [batch size, 1, 1, src len]
#self attention
_src, _ = self.self_attention(src, src, src, src_mask)
#dropout, residual connection and layer norm
src = self.self_attn_layer_norm(src + self.dropout(_src))
#src = [batch size, src len, hid dim]
#positionwise feedforward
_src = self.positionwise_feedforward(src)
#dropout, residual and layer norm
src = self.ff_layer_norm(src + self.dropout(_src))
#src = [batch size, src len, hid dim]
return src
# + [markdown] id="rUKKysfxeDAD"
# ### Слой многонаправленного внимания
#
# Одна из ключевых, новых концепций, представленных в статье о Transformer, - это *слой многонаправленного внимания*.
#
# 
#
# Внимание можно рассматривать как *запросы*, *ключи* и *значения* - где запрос используется с ключом для получения вектора внимания (обычно для вывода используется операция *softmax* и выходные величины имеют значения от 0 до 1, которые в сумме равны 1), используемый для получения взвешенной суммы значений.
#
# Трансформер использует *масштабированное скалярное произведение внимания*, для которого *запрос* и *ключ* объединяются путем взятия скалярного произведения между ними, затем применяя операцию softmax и масштабируя $d_k$ прежде чем, наконец, умножить на *значение*. $d_k$ это *размер направления*, `head_dim`, которое мы раскроем далее.
#
# $$ \text{Attention}(Q, K, V) = \text{Softmax} \big( \frac{QK^T}{\sqrt{d_k}} \big)V $$
#
# Это похоже на стандартное *скалярное произведение для внимания*, но масштабируется по $d_k$, которое, как говорится в статье, используется, чтобы остановить результаты скалярных произведений, становящиеся большими, что приводит к тому, что градиенты становятся слишком маленькими.
#
# Однако масштабированное скалярное произведение внимания применяется не просто к запросам, ключам и значениям. Вместо того, чтобы применять единственное внимание к запросам, ключам и значениям, их `hid_dim` разделить на $h$ *направлений* и масштабированное скалярное произведение внимания рассчитывается по всем направлениям параллельно. Это означает, что вместо того, чтобы уделять внимание одному понятию за одно применения внимания, мы обращаем внимание на $h$ понятий. Затем мы повторно объединяем направления в их `hid_dim` форму, таким образом, каждый `hid_dim` потенциально обращает внимание на $h$ разных понятий.
#
# $$ \text{MultiHead}(Q, K, V) = \text{Concat}(\text{head}_1,...,\text{head}_h)W^O $$
#
# $$\text{head}_i = \text{Attention}(QW_i^Q, KW_i^K, VW_i^V) $$
#
# $W^O$ это линейный слой, применяемый в конце слоя внимания с несколькими направлениями, `fc`. $W^Q, W^K, W^V$ линейные слои `fc_q`, `fc_k` и `fc_v`.
#
# Проходя по модулю, сначала вычисляем $QW^Q$, $KW^K$ и $VW^V$ с линейными слоями `fc_q`, `fc_k` и `fc_v`, дающие нам`Q`, `K` и `V`. Далее мы разбиваем `hid_dim` запроса, ключа и значения на `n_heads`, используя `.view` и правильно поменяв их порядок так, чтобы их можно было перемножить. Затем мы вычисляем `energy` (ненормализованное внимание) путем умножения `Q` и `K` вместе и масштабируя её на квадратный корень из `head_dim`, которое рассчитывается как `hid_dim // n_heads`. Затем мы маскируем энергию, чтобы не обращать внимания на элементы последовательности, на которые не следует сосредотачиваться, затем применяем softmax и дропаут. Затем мы применяем внимание к значениям направлений `V`, перед объединением `n_heads` вместе. Наконец, мы умножаем результат на $W^O$, представленное `fc_o`.
#
# Обратите внимание, что в нашей реализации длины ключей и значений всегда одинаковы, поэтому при матричном умножении выход softmax, `attention`, с `V` у нас всегда будут правильного размера для умножения матриц. Это умножение выполняется с использованием `torch.matmul` который, когда оба тензора > 2-мерны, выполняет пакетное матричное умножение по последним двум измерениям каждого тензора. Это будет **[query len, key len] x [value len, head dim]** умножение матрицы на размер пакета и каждое направление, которая обеспечивает результат вида **[batch size, n heads, query len, head dim]**.
#
# Поначалу кажется странным то, что дропаут применяется непосредственно к вниманию. Это означает, что наш вектор внимания, скорее всего, не будет суммироваться до 1, и мы можем уделять все внимание токену, но внимание к этому токену устанавливается на 0 в результате дропаута. Это никогда не объясняется и даже не упоминается в статье, однако используется в [официальное реализации](https://github.com/tensorflow/tensor2tensor/) и в каждой реализации Transformer, [включая BERT](https://github.com/google-research/bert/).
# + id="y-sQJFrXeDAE"
class MultiHeadAttentionLayer(nn.Module):
def __init__(self, hid_dim, n_heads, dropout, device):
super().__init__()
assert hid_dim % n_heads == 0
self.hid_dim = hid_dim
self.n_heads = n_heads
self.head_dim = hid_dim // n_heads
self.fc_q = nn.Linear(hid_dim, hid_dim)
self.fc_k = nn.Linear(hid_dim, hid_dim)
self.fc_v = nn.Linear(hid_dim, hid_dim)
self.fc_o = nn.Linear(hid_dim, hid_dim)
self.dropout = nn.Dropout(dropout)
self.scale = torch.sqrt(torch.FloatTensor([self.head_dim])).to(device)
def forward(self, query, key, value, mask = None):
batch_size = query.shape[0]
#query = [batch size, query len, hid dim]
#key = [batch size, key len, hid dim]
#value = [batch size, value len, hid dim]
Q = self.fc_q(query)
K = self.fc_k(key)
V = self.fc_v(value)
#Q = [batch size, query len, hid dim]
#K = [batch size, key len, hid dim]
#V = [batch size, value len, hid dim]
Q = Q.view(batch_size, -1, self.n_heads, self.head_dim).permute(0, 2, 1, 3)
K = K.view(batch_size, -1, self.n_heads, self.head_dim).permute(0, 2, 1, 3)
V = V.view(batch_size, -1, self.n_heads, self.head_dim).permute(0, 2, 1, 3)
#Q = [batch size, n heads, query len, head dim]
#K = [batch size, n heads, key len, head dim]
#V = [batch size, n heads, value len, head dim]
energy = torch.matmul(Q, K.permute(0, 1, 3, 2)) / self.scale
#energy = [batch size, n heads, query len, key len]
if mask is not None:
energy = energy.masked_fill(mask == 0, -1e10)
attention = torch.softmax(energy, dim = -1)
#attention = [batch size, n heads, query len, key len]
x = torch.matmul(self.dropout(attention), V)
#x = [batch size, n heads, query len, head dim]
x = x.permute(0, 2, 1, 3).contiguous()
#x = [batch size, query len, n heads, head dim]
x = x.view(batch_size, -1, self.hid_dim)
#x = [batch size, query len, hid dim]
x = self.fc_o(x)
#x = [batch size, query len, hid dim]
return x, attention
# + [markdown] id="nkiSMONWeDAE"
# ### Слой позиционно-зависимого прямого распространения
#
# Другой основной блок внутри уровня кодировщика — это *слой позиционно-зависимого прямого распространения*. Он устроен относительно просто по сравнению со слоем многонаправленного внимания. Вход преобразуется из `hid_dim` в `pf_dim`, где `pf_dim` обычно намного больше, чем `hid_dim`. Оригинальный Трансформер использовал `hid_dim` из 512 и `pf_dim` из 2048. Функция активации ReLU и дропаут применяются до того, как он снова преобразуется в представление `hid_dim`.
#
# Почему он используется? К сожалению, в статье это нигде не объясняется.
#
# BERT использует функцию активации [GELU](https://arxiv.org/abs/1606.08415), которую можно применить, просто переключив `torch.relu` на `F.gelu`. Почему они использовали GELU? Опять же, это никогда не объяснялось.
# + id="LxMkaf1PeDAE"
class PositionwiseFeedforwardLayer(nn.Module):
def __init__(self, hid_dim, pf_dim, dropout):
super().__init__()
self.fc_1 = nn.Linear(hid_dim, pf_dim)
self.fc_2 = nn.Linear(pf_dim, hid_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, x):
#x = [batch size, seq len, hid dim]
x = self.dropout(torch.relu(self.fc_1(x)))
#x = [batch size, seq len, pf dim]
x = self.fc_2(x)
#x = [batch size, seq len, hid dim]
return x
# + [markdown] id="Gd89Lm9-eDAF"
# ### Декодер
#
# Задача декодера — получить закодированное представление исходного предложения $Z$ и преобразовать его в предсказанные токены в целевом предложении $\hat{Y}$. Затем мы сравниваем $\hat{Y}$ с фактическими токенами в целевом предложении $Y$ для расчета потерь, которые будут использоваться для расчета градиентов параметров, а затем использованы в оптимизаторе для обновления весов с целью улучшить прогнозы.
#
# 
#
# Декодер похож на кодировщик, но в нём имеет два уровня внимания с несколькими направлениями. *Слой многонаправленного внимания с маскировкой* над целевой последовательностью и слоем многонаправленного внимания, который использует представление декодера в качестве запроса и представление кодера как ключ и значение.
#
# Декодер использует позиционный эмбеддинг и комбинирование - через поэлементную сумму - с масштабированными целевыми токенами, прошедшими эмбеддинг, после чего следует дропаут. Опять же, наши позиционные кодировки имеют «словарь» равный 100, что означает, что они могут принимать последовательности длиной до 100 токенов. При желании его можно увеличить.
#
# Комбинированные результаты эмбеддинга затем проходят через $N$ слоёв декодера, вместе с закодированным источником `enc_src`, а также исходной и целевой маской. Обратите внимание, что количество слоев в кодировщике не обязательно должно быть равно количеству слоев в декодере, даже если они оба обозначены как $N$.
#
# Представление декодера после $N$-го слоя затем пропускается через линейный слой `fc_out`. В PyTorch операция softmax содержится в нашей функции потерь, поэтому нам не нужно явно использовать здесь слой softmax.
#
# Помимо использования исходной маски, как мы это делали в кодировщике, чтобы наша модель не влияла на токен `<pad>`, мы также используем целевую маску. Это будет объяснено далее в `Seq2Seq` модели, которая инкапсулирует как кодер, так и декодер, но суть ее в том, что она выполняет ту же операцию, что и заполнение декодера в свёрточной модели sequence-to-sequence model. Поскольку мы обрабатываем все целевые токены одновременно и параллельно, нам нужен метод остановки декодера от «обмана», просто «глядя» на следующий токен в целевой последовательности и выводя его.
#
# Наш слой декодера также выводит нормализованные значения внимания, чтобы позже мы могли построить их график, чтобы увидеть, на что на самом деле обращает внимание наша модель.
# + id="XX2GuqlGeDAF"
class Decoder(nn.Module):
def __init__(self,
output_dim,
hid_dim,
n_layers,
n_heads,
pf_dim,
dropout,
device,
max_length = 100):
super().__init__()
self.device = device
self.tok_embedding = nn.Embedding(output_dim, hid_dim)
self.pos_embedding = nn.Embedding(max_length, hid_dim)
self.layers = nn.ModuleList([DecoderLayer(hid_dim,
n_heads,
pf_dim,
dropout,
device)
for _ in range(n_layers)])
self.fc_out = nn.Linear(hid_dim, output_dim)
self.dropout = nn.Dropout(dropout)
self.scale = torch.sqrt(torch.FloatTensor([hid_dim])).to(device)
def forward(self, trg, enc_src, trg_mask, src_mask):
#trg = [batch size, trg len]
#enc_src = [batch size, src len, hid dim]
#trg_mask = [batch size, 1, trg len, trg len]
#src_mask = [batch size, 1, 1, src len]
batch_size = trg.shape[0]
trg_len = trg.shape[1]
pos = torch.arange(0, trg_len).unsqueeze(0).repeat(batch_size, 1).to(self.device)
#pos = [batch size, trg len]
trg = self.dropout((self.tok_embedding(trg) * self.scale) + self.pos_embedding(pos))
#trg = [batch size, trg len, hid dim]
for layer in self.layers:
trg, attention = layer(trg, enc_src, trg_mask, src_mask)
#trg = [batch size, trg len, hid dim]
#attention = [batch size, n heads, trg len, src len]
output = self.fc_out(trg)
#output = [batch size, trg len, output dim]
return output, attention
# + [markdown] id="tHznByyGeDAG"
# ### Слой декодера
#
# Как упоминалось ранее, уровень декодера аналогичен уровню кодера, за исключением того, что теперь он имеет два уровня многонаправленного внимания `self_attention` и `encoder_attention`.
#
# Первый, как и в кодировщике, осуществляет внутреннее внимание, используя представление декодера, вплоть до запроса, ключа и значения. Затем следует дропаут, остаточное соединение и слой нормализации. Этот слой `self_attention` использует маску целевой последовательности `trg_mask`, чтобы предотвратить «обман» декодера, обращая внимание на токены, которые «опережают» тот, который он обрабатывает в настоящее время, поскольку он обрабатывает все токены в целевом предложении параллельно.
#
# Второй определяет как мы на самом деле подаём закодированное исходное предложение `enc_src` в наш декодер. В этом слое многонаправленного внимания запросы являются представлениями декодера, а ключи и значения — представлениями кодировщика. Здесь исходная маска `src_mask` используется для предотвращения того, чтобы слой многонаправленного внимания обращал внимание на токен `<pad>` в исходном предложении. Затем следуют уровни дропаута, остаточного соединения и уровень нормализации.
#
#
# Наконец, мы передаем результат через слой позиционно-зависимого прямого распространения и еще одна последовательность дропаута, остаточного соединения и уровень нормализации.
#
# Слой декодера не вводит никаких новых концепций, просто использует тот же набор слоев, что и кодировщик, но немного по-другому.
# + id="wjjqtbuEeDAG"
class DecoderLayer(nn.Module):
def __init__(self,
hid_dim,
n_heads,
pf_dim,
dropout,
device):
super().__init__()
self.self_attn_layer_norm = nn.LayerNorm(hid_dim)
self.enc_attn_layer_norm = nn.LayerNorm(hid_dim)
self.ff_layer_norm = nn.LayerNorm(hid_dim)
self.self_attention = MultiHeadAttentionLayer(hid_dim, n_heads, dropout, device)
self.encoder_attention = MultiHeadAttentionLayer(hid_dim, n_heads, dropout, device)
self.positionwise_feedforward = PositionwiseFeedforwardLayer(hid_dim,
pf_dim,
dropout)
self.dropout = nn.Dropout(dropout)
def forward(self, trg, enc_src, trg_mask, src_mask):
#trg = [batch size, trg len, hid dim]
#enc_src = [batch size, src len, hid dim]
#trg_mask = [batch size, 1, trg len, trg len]
#src_mask = [batch size, 1, 1, src len]
#self attention
_trg, _ = self.self_attention(trg, trg, trg, trg_mask)
#dropout, residual connection and layer norm
trg = self.self_attn_layer_norm(trg + self.dropout(_trg))
#trg = [batch size, trg len, hid dim]
#encoder attention
_trg, attention = self.encoder_attention(trg, enc_src, enc_src, src_mask)
#dropout, residual connection and layer norm
trg = self.enc_attn_layer_norm(trg + self.dropout(_trg))
#trg = [batch size, trg len, hid dim]
#positionwise feedforward
_trg = self.positionwise_feedforward(trg)
#dropout, residual and layer norm
trg = self.ff_layer_norm(trg + self.dropout(_trg))
#trg = [batch size, trg len, hid dim]
#attention = [batch size, n heads, trg len, src len]
return trg, attention
# + [markdown] id="d9SeKoymeDAG"
# ### Seq2Seq
#
# Наконец, у нас есть модуль `Seq2Seq`, который инкапсулирует кодировщик и декодер, а также управляет созданием масок.
#
# Исходная маска создается путем проверки того, что исходная последовательность не равна токену `<pad>`. Это 1, когда токен не является токеном `<pad>`, и 0, когда он является этим токеном. Затем он "разжимается", чтобы его можно было правильно транслировать при наложении маски на `energy`, которая имеет форму **_[batch size, n heads, seq len, seq len]_**.
#
# Целевая маска немного сложнее. Сначала мы создаем маску для токенов `<pad>`, как мы это делали для исходной маски. Далее, мы создаем «последующую» маску, `trg_sub_mask`, используя `torch.tril`. Таким образом создаётся диагональная матрица, в которой элементы над диагональю будут равны нулю, а элементы под диагональю будут установлены на любой входной тензор. В этом случае входным тензором будет тензор, заполненный единицами. Это означает, что наша `trg_sub_mask` будет выглядеть примерно так (для цели с 5 токенами):
#
# $$\begin{matrix}
# 1 & 0 & 0 & 0 & 0\\
# 1 & 1 & 0 & 0 & 0\\
# 1 & 1 & 1 & 0 & 0\\
# 1 & 1 & 1 & 1 & 0\\
# 1 & 1 & 1 & 1 & 1\\
# \end{matrix}$$
#
# Это показывает, на что может смотреть каждый целевой токен (строка) (столбец). Первый целевой токен имеет маску **_[1, 0, 0, 0, 0]_**, что означает, что он может смотреть только на первый целевой токен. Второй целевой токен имеет маску **_[1, 1, 0, 0, 0]_**, что означает, что он может просматривать как первый, так и второй целевые токены.
#
# Затем «последующая» маска логически дополняется маской заполнения, которая объединяет две маски, гарантируя, что ни последующие токены, ни маркеры заполнения не могут быть обработаны. Например, если бы последние два токена были токенами `<pad>`, маска выглядела бы так:
#
# $$\begin{matrix}
# 1 & 0 & 0 & 0 & 0\\
# 1 & 1 & 0 & 0 & 0\\
# 1 & 1 & 1 & 0 & 0\\
# 1 & 1 & 1 & 0 & 0\\
# 1 & 1 & 1 & 0 & 0\\
# \end{matrix}$$
#
# После того, как маски созданы, они используются с кодировщиком и декодером вместе с исходным и целевым предложениями, чтобы получить наше предсказанное целевое предложение, «output», вместе с вниманием декодера к исходной последовательности.
# + id="eDgCYHk6eDAH"
class Seq2Seq(nn.Module):
def __init__(self,
encoder,
decoder,
src_pad_idx,
trg_pad_idx,
device):
super().__init__()
self.encoder = encoder
self.decoder = decoder
self.src_pad_idx = src_pad_idx
self.trg_pad_idx = trg_pad_idx
self.device = device
def make_src_mask(self, src):
#src = [batch size, src len]
src_mask = (src != self.src_pad_idx).unsqueeze(1).unsqueeze(2)
#src_mask = [batch size, 1, 1, src len]
return src_mask
def make_trg_mask(self, trg):
#trg = [batch size, trg len]
trg_pad_mask = (trg != self.trg_pad_idx).unsqueeze(1).unsqueeze(2)
#trg_pad_mask = [batch size, 1, 1, trg len]
trg_len = trg.shape[1]
trg_sub_mask = torch.tril(torch.ones((trg_len, trg_len), device = self.device)).bool()
#trg_sub_mask = [trg len, trg len]
trg_mask = trg_pad_mask & trg_sub_mask
#trg_mask = [batch size, 1, trg len, trg len]
return trg_mask
def forward(self, src, trg):
#src = [batch size, src len]
#trg = [batch size, trg len]
src_mask = self.make_src_mask(src)
trg_mask = self.make_trg_mask(trg)
#src_mask = [batch size, 1, 1, src len]
#trg_mask = [batch size, 1, trg len, trg len]
enc_src = self.encoder(src, src_mask)
#enc_src = [batch size, src len, hid dim]
output, attention = self.decoder(trg, enc_src, trg_mask, src_mask)
#output = [batch size, trg len, output dim]
#attention = [batch size, n heads, trg len, src len]
return output, attention
# + [markdown] id="8cpVDTuOeDAH"
# ## Обучение модели Seq2Seq
#
# Теперь мы можем определить наш кодировщик и декодеры. Эта модель значительно меньше, чем Трансформеры, которые используются сегодня в исследованиях, но её можно быстро запустить на одном графическом процессоре.
# + id="BzAM-1EeeDAI"
INPUT_DIM = len(SRC.vocab)
OUTPUT_DIM = len(TRG.vocab)
HID_DIM = 256
ENC_LAYERS = 3
DEC_LAYERS = 3
ENC_HEADS = 8
DEC_HEADS = 8
ENC_PF_DIM = 512
DEC_PF_DIM = 512
ENC_DROPOUT = 0.1
DEC_DROPOUT = 0.1
enc = Encoder(INPUT_DIM,
HID_DIM,
ENC_LAYERS,
ENC_HEADS,
ENC_PF_DIM,
ENC_DROPOUT,
device)
dec = Decoder(OUTPUT_DIM,
HID_DIM,
DEC_LAYERS,
DEC_HEADS,
DEC_PF_DIM,
DEC_DROPOUT,
device)
# + [markdown] id="9-afo24OeDAI"
# Затем определяем и инкапсулирем модель sequence-to-sequence.
# + id="hC3RrufTeDAI"
SRC_PAD_IDX = SRC.vocab.stoi[SRC.pad_token]
TRG_PAD_IDX = TRG.vocab.stoi[TRG.pad_token]
model = Seq2Seq(enc, dec, SRC_PAD_IDX, TRG_PAD_IDX, device).to(device)
# + [markdown] id="ln8xZrNAeDAJ"
# Мы можем проверить количество параметров, заметив, что оно значительно меньше, чем 37M для модели свёрточной последовательности.
# + id="xP1Wa6w7eDAJ" colab={"base_uri": "https://localhost:8080/"} outputId="f7f84ed8-1637-4095-b9d4-4ef66acc2794"
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model):,} trainable parameters')
# + [markdown] id="3NSy_pbbeDAJ"
# В статье не упоминается, какая схема инициализации веса использовалась, однако форма Xavier, кажется, распространена среди моделей Transformer, поэтому мы используем ее здесь.
# + id="ioaWWCnveDAJ"
def initialize_weights(m):
if hasattr(m, 'weight') and m.weight.dim() > 1:
nn.init.xavier_uniform_(m.weight.data)
# + id="qQ2j_0oGeDAK"
model.apply(initialize_weights);
# + [markdown] id="TiStw6bneDAK"
# Оптимизатор, использованный в исходной статье Transformer, использует Adam со скоростью обучения, которая включает периоды «ускорения» и «торможения». BERT и другие модели Transformer используют Adam с фиксированной скоростью обучения, поэтому мы реализуем это. Проверьте [эту](http://nlp.seas.harvard.edu/2018/04/03/attention.html#optimizer) ссылку для получения более подробной информации о графике скорости обучения оригинального Transformer.
#
# Обратите внимание, что скорость обучения должна быть ниже, чем по умолчанию, используемой Адамом, иначе обучение будет нестабильным.
# + id="1UYMA88_eDAK"
LEARNING_RATE = 0.0005
optimizer = torch.optim.Adam(model.parameters(), lr = LEARNING_RATE)
# + [markdown] id="Fhl900ENeDAK"
# Next, we define our loss function, making sure to ignore losses calculated over `<pad>` tokens.
# + id="UICqTSymeDAL"
criterion = nn.CrossEntropyLoss(ignore_index = TRG_PAD_IDX)
# + [markdown] id="iGTfDmsceDAL"
# Затем мы определим наш цикл обучения. Это то же самое, что использовалось в предыдущей части.
#
# Поскольку мы хотим, чтобы наша модель предсказывала токен `<eos>`, но не выводила его в качестве выходных данных модели, мы просто отрезаем токен `<eos>` в конце последовательности. Таким образом:
#
# $$\begin{align*}
# \text{trg} &= [sos, x_1, x_2, x_3, eos]\\
# \text{trg[:-1]} &= [sos, x_1, x_2, x_3]
# \end{align*}$$
#
# $x_i$ обозначает фактический элемент целевой последовательности. Затем мы вводим это в модель, чтобы получить предсказанную последовательность, которая, мы надеемся, должна предсказывать токен `<eos>`:
#
# $$\begin{align*}
# \text{output} &= [y_1, y_2, y_3, eos]
# \end{align*}$$
#
# $y_i$ обозначает предсказанный элемент целевой последовательности. Затем мы вычисляем потери, используя исходный целевой тензор `trg` с токеном` <sos> `, отрезанным спереди, оставляя токен` <eos> `:
#
# $$\begin{align*}
# \text{output} &= [y_1, y_2, y_3, eos]\\
# \text{trg[1:]} &= [x_1, x_2, x_3, eos]
# \end{align*}$$
#
# Рассчитываем потери и обновляем параметры, как это обычно делается.
# + [markdown] id="ccRFgYeGNrlo"
# Для отображения прогресса в Google Colab устанавливаем tensorboard
# + pycharm={"name": "#%%\n"} id="LkzpXXiXNrlo"
# !pip install tensorboardX
# + pycharm={"name": "#%%\n"} id="DsZOmwRRNrlp"
from tensorboardX import SummaryWriter
import datetime
# Load the TensorBoard notebook extension
# %load_ext tensorboard
def train(model, iterator, optimizer, criterion, clip):
model.train()
epoch_loss = 0
for i, batch in enumerate(iterator):
src = batch.src
trg = batch.trg
optimizer.zero_grad()
output, _ = model(src, trg[:,:-1])
#output = [batch size, trg len - 1, output dim]
#trg = [batch size, trg len]
output_dim = output.shape[-1]
output = output.contiguous().view(-1, output_dim)
trg = trg[:,1:].contiguous().view(-1)
#output = [batch size * trg len - 1, output dim]
#trg = [batch size * trg len - 1]
loss = criterion(output, trg)
loss.backward()
torch.nn.utils.clip_grad_norm_(model.parameters(), clip)
optimizer.step()
epoch_loss += loss.item()
return epoch_loss / len(iterator)
# + [markdown] id="4cW7o40GeDAM"
# Цикл оценки такой же, как цикл обучения, только без вычислений градиента и обновления параметров.
# + id="0GqzApxLeDAM"
def evaluate(model, iterator, criterion):
model.eval()
epoch_loss = 0
with torch.no_grad():
for i, batch in enumerate(iterator):
src = batch.src
trg = batch.trg
output, _ = model(src, trg[:,:-1])
#output = [batch size, trg len - 1, output dim]
#trg = [batch size, trg len]
output_dim = output.shape[-1]
output = output.contiguous().view(-1, output_dim)
trg = trg[:,1:].contiguous().view(-1)
#output = [batch size * trg len - 1, output dim]
#trg = [batch size * trg len - 1]
loss = criterion(output, trg)
epoch_loss += loss.item()
return epoch_loss / len(iterator)
# + [markdown] id="Cscc-e_1eDAN"
# Затем мы определяем небольшую функцию, которую можем использовать, чтобы сообщить нам, сколько времени занимает эпоха.
# + id="DUyeAAqbeDAN"
def epoch_time(start_time, end_time):
elapsed_time = end_time - start_time
elapsed_mins = int(elapsed_time / 60)
elapsed_secs = int(elapsed_time - (elapsed_mins * 60))
return elapsed_mins, elapsed_secs
# + [markdown] id="gb5UqXIZNrlr"
# Запускаем tensorboard. Отображение данных начнётся после запуска обучения. Обновить данные можно кликнув по иконке справа вверху
# + pycharm={"name": "#%%\n"} id="BjpBlsspNrlr"
# Clear any logs from previous runs
# !rm -rf ./logs/
# %tensorboard --logdir runs
# + [markdown] id="Qbns68YAeDAN"
# Наконец, мы обучаем нашу фактическую модель. Эта модель почти в 3 раза быстрее, чем модель сверточной последовательности, а также обеспечивает меньшую сложность проверки!
# + id="_Bp64Iu_eDAN" colab={"base_uri": "https://localhost:8080/"} outputId="dc673303-e6a1-49a1-ac10-bb597e321003"
N_EPOCHS = 10
CLIP = 1
best_valid_loss = float('inf')
writer = SummaryWriter()
for epoch in range(N_EPOCHS):
start_time = time.time()
train_loss = train(model, train_iterator, optimizer, criterion, CLIP)
valid_loss = evaluate(model, valid_iterator, criterion)
end_time = time.time()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), 'tut6-model.pt')
print(f'Epoch: {epoch+1:02} | Time: {epoch_mins}m {epoch_secs}s')
print(f'\tTrain Loss: {train_loss:.3f} | Train PPL: {math.exp(train_loss):7.3f}')
print(f'\t Val. Loss: {valid_loss:.3f} | Val. PPL: {math.exp(valid_loss):7.3f}')
writer.add_scalar("Train Loss", train_loss, epoch+1)
writer.add_scalar("Train PPL", math.exp(train_loss), epoch+1)
writer.add_scalar("Val. Loss", valid_loss, epoch+1)
writer.add_scalar("Val. PPL", math.exp(valid_loss), epoch+1)
writer.close()
# + [markdown] id="FOkVvSSBeDAO"
# Мы загружаем наши «лучшие» параметры и добиваемся большей точности при тестировании, чем достигали все предыдущие модели.
# + id="_q4tv3UieDAO" colab={"base_uri": "https://localhost:8080/"} outputId="eb1ade78-0207-4f8d-d26c-b53ce9ef425d"
model.load_state_dict(torch.load('tut6-model.pt'))
test_loss = evaluate(model, test_iterator, criterion)
print(f'| Test Loss: {test_loss:.3f} | Test PPL: {math.exp(test_loss):7.3f} |')
# + [markdown] id="_sqZGLMheDAP"
# ## Вывод
#
# Теперь мы можем переводить с помощьюв нашей модели используя функцию `translate_sentence`.
#
# Были предприняты следующие шаги:
# - токенизируем исходное предложение, если оно не было токенизировано (является строкой)
# - добавляем токены `<sos>` и `<eos>`
# - оцифровываем исходное предложение
# - преобразовываем его в тензор и добавляем размер батча
# - создаём маску исходного предложения
# - загружаем исходное предложение и маску в кодировщик
# - создаём список для хранения выходного предложения, инициализированного токеном `<sos>`
# - пока мы не достигли максимальной длины
# - преобразовываем текущий прогноз выходного предложения в тензор с размерностью батча
# - создаём маску целевого предложения
# - поместим текущий выход, выход кодировщика и обе маски в декодер
# - получаем предсказание следующего выходного токена от декодера вместе с вниманием
# - добавляем предсказание к предсказанию текущего выходного предложения
# - прерываем, если предсказание было токеном `<eos>`
# - преобразовываем выходное предложение из индексов в токены
# - возвращаем выходное предложение (с удаленным токеном `<sos>`) и вниманием с последнего слоя
# + id="9vWuCZZmeDAP"
def translate_sentence(sentence, src_field, trg_field, model, device, max_len = 50):
model.eval()
if isinstance(sentence, str):
nlp = spacy.load('de_core_news_sm')
tokens = [token.text.lower() for token in nlp(sentence)]
else:
tokens = [token.lower() for token in sentence]
tokens = [src_field.init_token] + tokens + [src_field.eos_token]
src_indexes = [src_field.vocab.stoi[token] for token in tokens]
src_tensor = torch.LongTensor(src_indexes).unsqueeze(0).to(device)
src_mask = model.make_src_mask(src_tensor)
with torch.no_grad():
enc_src = model.encoder(src_tensor, src_mask)
trg_indexes = [trg_field.vocab.stoi[trg_field.init_token]]
for i in range(max_len):
trg_tensor = torch.LongTensor(trg_indexes).unsqueeze(0).to(device)
trg_mask = model.make_trg_mask(trg_tensor)
with torch.no_grad():
output, attention = model.decoder(trg_tensor, enc_src, trg_mask, src_mask)
pred_token = output.argmax(2)[:,-1].item()
trg_indexes.append(pred_token)
if pred_token == trg_field.vocab.stoi[trg_field.eos_token]:
break
trg_tokens = [trg_field.vocab.itos[i] for i in trg_indexes]
return trg_tokens[1:], attention
# + [markdown] id="W9rGNStpeDAP"
# Теперь мы определим функцию, которая отображает внимание к исходному предложению на каждом этапе декодирования. Поскольку у этой модели 8 направлений, мы можем наблюдать за вниманием для каждой из них.
# + id="bx-0QXdJeDAQ"
def display_attention(sentence, translation, attention, n_heads = 8, n_rows = 4, n_cols = 2):
assert n_rows * n_cols == n_heads
fig = plt.figure(figsize=(15,25))
for i in range(n_heads):
ax = fig.add_subplot(n_rows, n_cols, i+1)
_attention = attention.squeeze(0)[i].cpu().detach().numpy()
cax = ax.matshow(_attention, cmap='bone')
ax.tick_params(labelsize=12)
ax.set_xticklabels(['']+['<sos>']+[t.lower() for t in sentence]+['<eos>'],
rotation=45)
ax.set_yticklabels(['']+translation)
ax.xaxis.set_major_locator(ticker.MultipleLocator(1))
ax.yaxis.set_major_locator(ticker.MultipleLocator(1))
plt.show()
plt.close()
# + [markdown] id="dz6l-qBqeDAQ"
# Сначала возьмем пример из обучающей выборки.
# + id="ZROLKC-ceDAQ" colab={"base_uri": "https://localhost:8080/"} outputId="96444767-d07c-4e7e-b3a1-957d2ddc95b6"
example_idx = 8
src = vars(train_data.examples[example_idx])['src']
trg = vars(train_data.examples[example_idx])['trg']
print(f'src = {src}')
print(f'trg = {trg}')
# + [markdown] id="DJ6Wy6GheDAQ"
# Наш перевод выглядит неплохо, хотя наша модель меняет *is walking by* на *walks by*. Смысл все тот же.
# + id="vWcm12oMeDAR" colab={"base_uri": "https://localhost:8080/"} outputId="d37fcd5a-f42e-4d84-acea-215c222f664f"
translation, attention = translate_sentence(src, SRC, TRG, model, device)
print(f'predicted trg = {translation}')
# + [markdown] id="4R8PWHheeDAR"
# Мы можем видеть внимание каждого направления ниже. Конечно, все они разные, но трудно (возможно, невозможно) понять, на что отдельное направление научилось обращать внимание. Некоторые направления уделяют все внимание "eine" при переводе "a", некоторые совсем не сосредоточены, а другие слегка. Кажется, что все они следуют аналогичному шаблону «нисходящей лестницы», и внимание при выводе двух последних токенов равномерно распределяется между двумя последними токенами во входном предложении.
# + id="VG6hX0AYeDAR" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="e7ad7f88-779a-4152-d06b-56cff171d452"
display_attention(src, translation, attention)
# + [markdown] id="0jgegHsmeDAR"
# Затем давайте возьмем пример, на котором модель не обучалась, из набора для проверки.
# + id="L4E772bseDAR" colab={"base_uri": "https://localhost:8080/"} outputId="8f67ef8f-36e0-46ba-e685-bfef64623227"
example_idx = 6
src = vars(valid_data.examples[example_idx])['src']
trg = vars(valid_data.examples[example_idx])['trg']
print(f'src = {src}')
print(f'trg = {trg}')
# + [markdown] id="9GH716XmeDAR"
# Модель переводит это, переключая *is running* на просто *runs*, но это приемлемая замена.
# + id="5lveTOoJeDAS" colab={"base_uri": "https://localhost:8080/"} outputId="fd7e3096-591d-4190-d905-68a34e68f88d"
translation, attention = translate_sentence(src, SRC, TRG, model, device)
print(f'predicted trg = {translation}')
# + [markdown] id="VpjVZOX1eDAS"
# Опять же, некоторые направления внимания полностью обращают внимание на "ein", в то время как некоторые не обращают на него внимания. Опять же, при выводе точки и предложения <eos> в прогнозируемом целевом предложении кажется, что большинство направления внимания распределяют свое внимание как на точку, так и на токены `<eos>` в исходном предложении, хотя некоторые, похоже, обращают внимание на токены от начала предложения.
# + id="9p6MMbgweDAS" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="6f355355-ee8e-44cf-b234-ec19443d571d"
display_attention(src, translation, attention)
# + [markdown] id="8BKYqiBveDAT"
# Наконец, мы рассмотрим пример из тестовых данных.
# + id="IInGOZv3eDAT" colab={"base_uri": "https://localhost:8080/"} outputId="44c4ebc2-d96b-480e-d188-fe9608560afd"
example_idx = 10
src = vars(test_data.examples[example_idx])['src']
trg = vars(test_data.examples[example_idx])['trg']
print(f'src = {src}')
print(f'trg = {trg}')
# + [markdown] id="e-YO1E2peDAT"
# Прекрасный перевод!
# + id="XQeXDWPaeDAT" colab={"base_uri": "https://localhost:8080/"} outputId="5e08aeca-b32d-4617-8bf1-bc1f23e9441c"
translation, attention = translate_sentence(src, SRC, TRG, model, device)
print(f'predicted trg = {translation}')
# + id="zRM2zwMWeDAU" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="d48b04fe-6322-41f2-ddbc-b708daed586a"
display_attention(src, translation, attention)
# + [markdown] id="MK6Gy8w8eDAU"
# ## BLEU
#
# Наконец, мы рассчитываем оценку BLEU для трансформатора.
# + id="IqDQyipreDAU"
from torchtext.data.metrics import bleu_score
def calculate_bleu(data, src_field, trg_field, model, device, max_len = 50):
trgs = []
pred_trgs = []
for datum in data:
src = vars(datum)['src']
trg = vars(datum)['trg']
pred_trg, _ = translate_sentence(src, src_field, trg_field, model, device, max_len)
#cut off <eos> token
pred_trg = pred_trg[:-1]
pred_trgs.append(pred_trg)
trgs.append([trg])
return bleu_score(pred_trgs, trgs)
# + [markdown] id="mkGD72NUeDAU"
# Мы получили оценку BLEU 36,52, что превосходит ~ 34 для свёрточной модели sequence-to-sequence и ~ 28 для модели RNN, основанной на внимании. И все это с наименьшим количеством параметров и самым быстрым временем обучения!
# + id="tqtGecXVeDAV" colab={"base_uri": "https://localhost:8080/"} outputId="c91ef262-357f-4e9c-8787-41558dc25375"
bleu_score = calculate_bleu(test_data, SRC, TRG, model, device)
print(f'BLEU score = {bleu_score*100:.2f}')
# + [markdown] id="u5N0uto1eDAV"
# Поздравляем с окончанием этих уроков! Надеюсь, они были вам полезны.
#
# Если вы обнаружите какие-либо ошибки или захотите задать какие-либо вопросы по поводу какого-либо кода или используемых объяснений, не стесняйтесь отправлять проблему на GitHub, и я постараюсь исправить ее как можно скорее.
#
# ## Приложение
#
# Вышеупомянутая функция `calculate_bleu` не оптимизирована. Ниже представлена значительно более быстрая векторизованная версия, которую следует использовать при необходимости. Благодарим за реализацию [@azadyasar](https://github.com/azadyasar).
# + id="YS6PWyvJeDAV"
def translate_sentence_vectorized(src_tensor, src_field, trg_field, model, device, max_len=50):
assert isinstance(src_tensor, torch.Tensor)
model.eval()
src_mask = model.make_src_mask(src_tensor)
with torch.no_grad():
enc_src = model.encoder(src_tensor, src_mask)
# enc_src = [batch_sz, src_len, hid_dim]
trg_indexes = [[trg_field.vocab.stoi[trg_field.init_token]] for _ in range(len(src_tensor))]
# Even though some examples might have been completed by producing a <eos> token
# we still need to feed them through the model because other are not yet finished
# and all examples act as a batch. Once every single sentence prediction encounters
# <eos> token, then we can stop predicting.
translations_done = [0] * len(src_tensor)
for i in range(max_len):
trg_tensor = torch.LongTensor(trg_indexes).to(device)
trg_mask = model.make_trg_mask(trg_tensor)
with torch.no_grad():
output, attention = model.decoder(trg_tensor, enc_src, trg_mask, src_mask)
pred_tokens = output.argmax(2)[:,-1]
for i, pred_token_i in enumerate(pred_tokens):
trg_indexes[i].append(pred_token_i)
if pred_token_i == trg_field.vocab.stoi[trg_field.eos_token]:
translations_done[i] = 1
if all(translations_done):
break
# Iterate through each predicted example one by one;
# Cut-off the portion including the after the <eos> token
pred_sentences = []
for trg_sentence in trg_indexes:
pred_sentence = []
for i in range(1, len(trg_sentence)):
if trg_sentence[i] == trg_field.vocab.stoi[trg_field.eos_token]:
break
pred_sentence.append(trg_field.vocab.itos[trg_sentence[i]])
pred_sentences.append(pred_sentence)
return pred_sentences, attention
# + id="D1h_xidteDAW"
from torchtext.data.metrics import bleu_score
def calculate_bleu_alt(iterator, src_field, trg_field, model, device, max_len = 50):
trgs = []
pred_trgs = []
with torch.no_grad():
for batch in iterator:
src = batch.src
trg = batch.trg
_trgs = []
for sentence in trg:
tmp = []
# Start from the first token which skips the <start> token
for i in sentence[1:]:
# Targets are padded. So stop appending as soon as a padding or eos token is encountered
if i == trg_field.vocab.stoi[trg_field.eos_token] or i == trg_field.vocab.stoi[trg_field.pad_token]:
break
tmp.append(trg_field.vocab.itos[i])
_trgs.append([tmp])
trgs += _trgs
pred_trg, _ = translate_sentence_vectorized(src, src_field, trg_field, model, device)
pred_trgs += pred_trg
return pred_trgs, trgs, bleu_score(pred_trgs, trgs)
# + [markdown] id="h8jeoiKlNrl1"
# ## Обучение сети инвертированию предложения
#
# В конце приведу один из моих любимых тестов: тест на инверсию предложения. Очень простая для человека задача (ученики начальной школы обучаются за 10-15 примеров), но, порой, непреодолима для искусственных систем.
# + [markdown] id="crX4svL_QI5O"
# Для Google Colab скачаем обучающие последовательности
# + id="0XMaaybcRTFo"
# !wget https://raw.githubusercontent.com/vasiliyeskin/bentrevett-pytorch-seq2seq_ru/master/toy_revert/train.csv -P toy_revert
# !wget https://raw.githubusercontent.com/vasiliyeskin/bentrevett-pytorch-seq2seq_ru/master/toy_revert/val.csv -P toy_revert
# !wget https://raw.githubusercontent.com/vasiliyeskin/bentrevett-pytorch-seq2seq_ru/master/toy_revert/test.csv -P toy_revert
# + [markdown] id="edFokHttNrl1"
# В начале обучим сеть инверсии и посмотрим на результат.
#
# + colab={"base_uri": "https://localhost:8080/"} id="HKIUr8XaZcyi" outputId="1911b096-8343-4faa-8cfb-0125463b1909"
SRC = Field(tokenize="spacy",
init_token='<sos>',
eos_token='<eos>',
lower=True,
batch_first = True)
TRG = Field(tokenize="spacy",
init_token='<sos>',
eos_token='<eos>',
lower=True,
batch_first = True)
data_fields = [('src', SRC), ('trg', TRG)]
# load the dataset in csv format
train_data, valid_data, test_data = TabularDataset.splits(
path='toy_revert',
train='train.csv',
validation='val.csv',
test='test.csv',
format='csv',
fields=data_fields,
skip_header=True
)
SRC.build_vocab(train_data)
TRG.build_vocab(train_data)
# + pycharm={"name": "#%%\n"} id="5flcPxdENrl1" colab={"base_uri": "https://localhost:8080/"} outputId="7ed84c4c-09d8-4ea9-9d9d-aa493ae4ee01"
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
BATCH_SIZE = 32
train_iterator, valid_iterator, test_iterator = BucketIterator.splits(
(train_data, valid_data, test_data),
batch_size = BATCH_SIZE,
sort_key = lambda x: len(x.src),
sort_within_batch=True,
device = device)
################## create DNN Seq2Seq ###############################
INPUT_DIM = len(SRC.vocab)
OUTPUT_DIM = len(TRG.vocab)
HID_DIM = 64
ENC_LAYERS = 3
DEC_LAYERS = 3
ENC_HEADS = 8
DEC_HEADS = 8
ENC_PF_DIM = 512
DEC_PF_DIM = 512
ENC_DROPOUT = 0.1
DEC_DROPOUT = 0.1
enc = Encoder(INPUT_DIM,
HID_DIM,
ENC_LAYERS,
ENC_HEADS,
ENC_PF_DIM,
ENC_DROPOUT,
device)
dec = Decoder(OUTPUT_DIM,
HID_DIM,
DEC_LAYERS,
DEC_HEADS,
DEC_PF_DIM,
DEC_DROPOUT,
device)
SRC_PAD_IDX = SRC.vocab.stoi[SRC.pad_token]
TRG_PAD_IDX = TRG.vocab.stoi[TRG.pad_token]
model = Seq2Seq(enc, dec, SRC_PAD_IDX, TRG_PAD_IDX, device).to(device)
####################################################################
####### initial weights
model.apply(initialize_weights);
# print(model)
print(f'The model has {count_parameters(model):,} trainable parameters')
optimizer = optim.Adam(model.parameters(), lr = LEARNING_RATE)
criterion = nn.CrossEntropyLoss(ignore_index=TRG_PAD_IDX)
N_EPOCHS = 30
CLIP = 1
best_valid_loss = float('inf')
for epoch in range(N_EPOCHS):
start_time = time.time()
train_loss = train(model, train_iterator, optimizer, criterion, CLIP)
valid_loss = evaluate(model, valid_iterator, criterion)
end_time = time.time()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), 'tut3-model.pt')
print(f'Epoch: {epoch + 1:02} | Time: {epoch_mins}m {epoch_secs}s')
print(f'\tTrain Loss: {train_loss:.3f} | Train PPL: {math.exp(train_loss):7.3f}')
print(f'\t Val. Loss: {valid_loss:.3f} | Val. PPL: {math.exp(valid_loss):7.3f}')
# writer.add_scalar("Train_loss_average_per_epoch", train_loss, epoch)
# writer.add_scalar("Validate_loss_average_per_epoch", valid_loss, epoch)
model.load_state_dict(torch.load('tut3-model.pt'))
test_loss = evaluate(model, test_iterator, criterion)
print(f'| Test Loss: {test_loss:.3f} | Test PPL: {math.exp(test_loss):7.3f} |')
# + [markdown] id="r7JmvyFYNrl2"
# testing with the life example
# + id="kl2Hz5Sy9pUV"
def translate_sentence(sentence, src_field, trg_field, model, device, max_len = 20):
model.eval()
if isinstance(sentence, str):
nlp = spacy.load('de_core_news_sm')
tokens = [token.text.lower() for token in nlp(sentence)]
else:
tokens = [token.lower() for token in sentence]
tokens = [src_field.init_token] + tokens + [src_field.eos_token]
src_indexes = [src_field.vocab.stoi[token] for token in tokens]
src_tensor = torch.LongTensor(src_indexes).unsqueeze(0).to(device)
src_mask = model.make_src_mask(src_tensor)
with torch.no_grad():
enc_src = model.encoder(src_tensor, src_mask)
trg_indexes = [trg_field.vocab.stoi[trg_field.init_token]]
for i in range(max_len):
trg_tensor = torch.LongTensor(trg_indexes).unsqueeze(0).to(device)
trg_mask = model.make_trg_mask(trg_tensor)
with torch.no_grad():
output, attention = model.decoder(trg_tensor, enc_src, trg_mask, src_mask)
pred_token = output.argmax(2)[:,-1].item()
trg_indexes.append(pred_token)
if pred_token == trg_field.vocab.stoi[trg_field.eos_token]:
break
trg_tokens = [trg_field.vocab.itos[i] for i in trg_indexes]
return trg_tokens[1:], attention
# + id="V_Yp-8RyDCJt"
from torchtext.data.metrics import bleu_score
def calculate_bleu(data, src_field, trg_field, model, device, max_len = 20):
trgs = []
pred_trgs = []
for datum in data:
src = vars(datum)['src']
trg = vars(datum)['trg']
pred_trg, _ = translate_sentence(src, src_field, trg_field, model, device, max_len)
#cut off <eos> token
pred_trg = pred_trg[:-1]
pred_trgs.append(pred_trg)
trgs.append([trg])
return bleu_score(pred_trgs, trgs)
# + pycharm={"name": "#%%\n"} id="Y-0WOawtNrl2" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="64c21c14-ee56-46f5-b6f8-c79b868effd6"
example_idx = 10
src = vars(test_data.examples[example_idx])['src']
trg = vars(test_data.examples[example_idx])['trg']
print(f'src = {src}')
print(f'trg = {trg}')
print(f'source = {src}')
translation, attention = translate_sentence(src, SRC, TRG, model, device)
display_attention(src, translation, attention)
print(f'predicted trg = {translation}')
src = ['a', 'b', 'c', 'a', 'd']
print(f'source = {src}')
translation, attention = translate_sentence(src, SRC, TRG, model, device)
display_attention(src, translation, attention)
print(f'predicted trg = {translation}')
src = 'd b c d'.split(' ')
print(f'source = {src}')
translation, attention = translate_sentence(src, SRC, TRG, model, device)
display_attention(src, translation, attention)
print(f'predicted trg = {translation}')
src = ['a', 'a', 'a', 'a', 'd']
print(f'source = {src}')
translation, attention = translate_sentence(src, SRC, TRG, model, device)
print(f'predicted trg = {translation}')
src = ['d', 'b', 'c', 'a']
print(f'source = {src}')
translation, attention = translate_sentence(src, SRC, TRG, model, device)
print(f'predicted trg = {translation}')
src = ['d', 'd', 'd', 'd', 'd', 'd', 'd', 'd']
print(f'source = {src}')
translation, attention = translate_sentence(src, SRC, TRG, model, device)
print(f'predicted trg = {translation}')
bleu_score = calculate_bleu(test_data, SRC, TRG, model, device)
print(f'BLEU score = {bleu_score * 100:.2f}')
| 6 - Attention is All You Need.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ##### The following experiment is to convert bitcoin address, domains, nodes as an input vector and predict the unseen nodes, Here in this notebook Node2Vec method has used.
#
#
import pandas as pd
from tqdm.notebook import tqdm
import numpy as np
from tensorflow.keras import Model
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.layers import Dense, Conv1D, MaxPool1D, Dropout, Flatten
from tensorflow.keras.losses import binary_crossentropy
import tensorflow as tf
from sklearn import model_selection
from IPython.display import display, HTML
import stellargraph as sg
from stellargraph.data import EdgeSplitter
from stellargraph.mapper import PaddedGraphGenerator
from stellargraph.layer import DeepGraphCNN
from stellargraph import StellarGraph
from stellargraph import datasets
import networkx as nx
from stellargraph.data import BiasedRandomWalk
from sklearn.manifold import TSNE
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegressionCV
from sklearn.metrics import accuracy_score
import matplotlib.pyplot as plt
import seaborn as sns
import scikitplot as skplt
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.ensemble import AdaBoostClassifier
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.metrics import accuracy_score
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
from sklearn.metrics import roc_curve
from sklearn.metrics import auc
import os
import random
import stellargraph as sg
from stellargraph.data import UnsupervisedSampler
from stellargraph.mapper import Attri2VecLinkGenerator, Attri2VecNodeGenerator
from stellargraph.layer import Attri2Vec, link_classification
import warnings
warnings.filterwarnings("ignore")
from sklearn.pipeline import Pipeline
from sklearn.linear_model import LogisticRegressionCV
from sklearn.metrics import roc_auc_score
from sklearn.preprocessing import StandardScaler
LIMIT = 250
# +
bitcoin_address = pd.read_csv('bitcoin_address.csv',nrows=LIMIT)
bitcoin_address_link = pd.read_csv('bitcoin_address_link.csv',nrows=LIMIT)
page = pd.read_csv('page.csv',nrows=LIMIT)
page_link = pd.read_csv('page_link.csv',nrows=LIMIT)
domain = pd.read_csv('domain.csv',nrows=LIMIT)
# +
G1 = nx.Graph()
node_subjects = {}
#insert bitcoin address nodes
for index, row in tqdm(bitcoin_address.iterrows()):
G1.add_node(int(row.id))
node_subjects[row.id]=1
#insert pages
for index, row in tqdm(page.iterrows()):
G1.add_node(int(row.id))
node_subjects[row.id]=2
#insert page - bitcoin addres edges
for index, row in tqdm(bitcoin_address_link.iterrows()):
G1.add_edge(int(row.bitcoin_address), int(row.page))
#insert page - page edges
for index, row in tqdm(page_link.iterrows()):
G1.add_edge(int(row.link_from), int(row.link_to ))
#insert domain
for index, row in tqdm(domain.iterrows()):
G1.add_node(int(row.id))
node_subjects[row.id]=3
#insert page - domain edges
for index, row in tqdm(page.iterrows()):
G1.add_edge(int(row.id), int(row.domain))
# -
g1 = StellarGraph.from_networkx(G1)
print(g1.info())
# +
edge_splitter_test = EdgeSplitter(g1)
graph_test, examples_test, labels_test = edge_splitter_test.train_test_split(p=0.1, method="global")
print(graph_test.info())
# +
# Do the same process to compute a training subset from within the test graph
edge_splitter_train = EdgeSplitter(graph_test, g1)
graph_train, examples, labels = edge_splitter_train.train_test_split(p=0.1, method="global")
(examples_train,examples_model_selection,labels_train,labels_model_selection,) = train_test_split(examples, labels, train_size=0.75, test_size=0.25)
print(graph_train.info())
# -
pd.DataFrame({
'Split':['Training Set','Model Selection','Test set'],
'Number of Examples':[len(examples_train),len(examples_model_selection),len(examples_test)]
})
import multiprocessing
P = 1.0
Q = 1.0
DIMENSIONS = 128
NUMOFWALKS =10
WALKOFLENGTH = 80
WINDOWSIZE =10
NUMITER = 1
WORKERS = multiprocessing.cpu_count()
# +
from gensim.models import Word2Vec
def node2vec_embedding(graph, name):
rw = BiasedRandomWalk(graph)
walks = rw.run(graph.nodes(), n=NUMOFWALKS, length=WALKOFLENGTH, p=P, q=Q)
print(f"Number of random walks for '{name}': {len(walks)}")
model = Word2Vec(
walks,
size=DIMENSIONS,
window=WINDOWSIZE,
min_count=0,
sg=1,
workers=WORKERS,
iter=NUMITER,
)
get_embedding = lambda u: model.wv[u]
return get_embedding
# -
embedding_train = node2vec_embedding(graph_train, "Train Graph")
def linkExamplesToFeatures(linkExamples, transformNode, binaryOperator):
return [binaryOperator(transformNode(src), transformNode(dst)) for src, dst in linkExamples]
def trainLinkPredictionModel(linkExamples, linkLabels, getEmbedding, binaryOperator):
classif = linkPredictionClassifier()
linkFeatures = linkExamplesToFeatures(linkExamples, getEmbedding, binaryOperator)
classif.fit(linkFeatures, linkLabels)
return classif
def linkPredictionClassifier(max_iter=2000):
lr_clf = LogisticRegressionCV(Cs=10, cv=10, scoring="roc_auc", max_iter=max_iter)
return Pipeline(steps=[("sc", StandardScaler()), ("clf", lr_clf)])
def evaluateLinkPredictionModel(clf, link_examples_test, link_labels_test, get_embedding, binary_operator):
link_features_test = linkExamplesToFeatures(link_examples_test, get_embedding, binary_operator)
score = evaluate_ROC_AUC(clf, link_features_test, link_labels_test)
return score
def evaluate_ROC_AUC(clf, link_features, link_labels):
predicted = clf.predict_proba(link_features)
positive_column = list(clf.classes_).index(1)
return roc_auc_score(link_labels, predicted[:, positive_column])
def operator_hadamard(u, v):
return u * v
def operator_l1(u, v):
return np.abs(u - v)
def operator_l2(u, v):
return (u - v) ** 2
def operator_avg(u, v):
return (u + v) / 2.0
def run_link_prediction(binary_operator):
clf = trainLinkPredictionModel(examples_train, labels_train, embedding_train, binary_operator)
score = evaluateLinkPredictionModel(
clf,
examples_model_selection,
labels_model_selection,
embedding_train,
binary_operator,
)
return {
"classifier": clf,
"binary_operator": binary_operator,
"score": score,
}
binary_operators = [operator_hadamard, operator_l1, operator_l2, operator_avg]
# +
results = [run_link_prediction(op) for op in binary_operators]
best_result = max(results, key=lambda result: result["score"])
print(f"Best result from '{best_result['binary_operator'].__name__}'")
pd.DataFrame(
[(result["binary_operator"].__name__, result["score"]) for result in results],
columns=("name", "ROC AUC score"),
).set_index("name")
# -
embedding_test = node2vec_embedding(graph_test, "Test Graph")
test_score = evaluateLinkPredictionModel(best_result["classifier"],examples_test,labels_test,embedding_test,best_result["binary_operator"],)
print(f"ROC AUC score on test set using '{best_result['binary_operator'].__name__}': {test_score}")
# +
# print('LogisticRegressionCV',accuracy_score(y_test,y_pred),
# ' precision : ',precision_score(y_test,y_pred,average='micro'),
# ' recall : ',recall_score(y_test,y_pred,average='micro'),'\n')
# print(classification_report(y_test,y_pred))
# +
# plt.title("Confusion Matrix")
# sns.heatmap(confusion_matrix(y_test, y_pred),annot=True,cmap="Blues",fmt="g",cbar=False)
# plt.show()
| notebooks/node2vec-link-prediction [bitcoin address vs domains vs pages].ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="WPjSABTOpVUo" executionInfo={"status": "ok", "timestamp": 1638271520000, "user_tz": -60, "elapsed": 41568, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitatTDk380OCMvv2bDBT7Lsunn79WNpphzKI_dytU=s64", "userId": "08737126295476082217"}} outputId="e4fe8936-ea93-4472-9d97-3727e3137aed"
from IPython.display import clear_output
# https://www.dgl.ai/pages/start.html
# # !pip install dgl
# !pip install dgl-cu111 -f https://data.dgl.ai/wheels/repo.html # FOR CUDA VERSION
# !pip install dgllife
# !pip install rdkit-pypi
# !pip install --pre deepchem
# !pip install ipython-autotime
# !pip install gputil
# !pip install psutil
# !pip install humanize
# %load_ext autotime
clear = clear_output()
# + colab={"base_uri": "https://localhost:8080/"} id="AKaZdFr8pbzR" executionInfo={"status": "ok", "timestamp": 1638271528205, "user_tz": -60, "elapsed": 8215, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitatTDk380OCMvv2bDBT7Lsunn79WNpphzKI_dytU=s64", "userId": "08737126295476082217"}} outputId="730e6fdf-ee9f-40b6-ee92-b58466336b81"
import os
from os import path
import statistics
import warnings
import random
import time
import itertools
import psutil
import humanize
import GPUtil as GPU
import subprocess
from datetime import datetime, timedelta
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import tqdm
from tqdm import trange, tqdm_notebook, tnrange
import deepchem as dc
import rdkit
from rdkit import Chem
from rdkit.Chem.MolStandardize import rdMolStandardize
# embedding
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
from torch.optim.lr_scheduler import ReduceLROnPlateau
from torch.profiler import profile, record_function, ProfilerActivity
from torch.utils.tensorboard import SummaryWriter
import sklearn
from sklearn.metrics import (auc, roc_curve, roc_auc_score, average_precision_score,
accuracy_score, ConfusionMatrixDisplay, confusion_matrix, precision_recall_curve,
f1_score, PrecisionRecallDisplay)
from sklearn.ensemble import RandomForestClassifier
warnings.filterwarnings("ignore", message="DGLGraph.__len__")
DGLBACKEND = 'pytorch'
clear
def get_cmd_output(command):
return subprocess.check_output(command,
stderr=subprocess.STDOUT,
shell=True).decode('UTF-8')
# + colab={"base_uri": "https://localhost:8080/"} id="JCSXIJOJpgO0" executionInfo={"status": "ok", "timestamp": 1638271528206, "user_tz": -60, "elapsed": 24, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitatTDk380OCMvv2bDBT7Lsunn79WNpphzKI_dytU=s64", "userId": "08737126295476082217"}} outputId="9b671f35-b4f5-4631-db47-c478477ce372"
def load_dataset(dataset, bonds=False, feat='graph', create_new=False):
"""
dataset values: muv, tox21, dude-gpcr
feat values: graph, ecfp
"""
dataset_test_tasks = {
'tox21': ['SR-HSE', 'SR-MMP', 'SR-p53'],
'muv': ['MUV-832', 'MUV-846', 'MUV-852', 'MUV-858', 'MUV-859'],
'dude-gpcr': ['adrb2', 'cxcr4']
}
dataset_original = dataset
if bonds:
dataset = dataset + "_with_bonds"
if path.exists(f"{drive_path}/data/{dataset}_dgl.pkl") and not create_new:
# Load Dataset
print("Reading Pickle")
if feat == 'graph':
data = pd.read_pickle(f"{drive_path}/data/{dataset}_dgl.pkl")
else:
data = pd.read_pickle(f"{drive_path}/data/{dataset}_ecfp.pkl")
else:
# Create Dataset
df = pd.read_csv(f"{drive_path}/data/raw/{dataset_original}.csv")
if feat == 'graph':
data = create_dataset(df, f"{dataset}_dgl", bonds)
else:
data = create_ecfp_dataset(df, f"{dataset}_ecfp")
test_tasks = dataset_test_tasks.get(dataset_original)
drop_cols = test_tasks.copy()
drop_cols.extend(['mol_id', 'smiles', 'mol'])
train_tasks = [x for x in list(data.columns) if x not in drop_cols]
train_dfs = dict.fromkeys(train_tasks)
for task in train_tasks:
df = data[[task, 'mol']].dropna()
df.columns = ['y', 'mol']
# FOR BOND INFORMATION
if with_bonds:
for index, r in df.iterrows():
if r.mol.edata['edge_feats'].shape[-1] < 17:
df.drop(index, inplace=True)
train_dfs[task] = df
for key in train_dfs:
print(key, len(train_dfs[key]))
if feat == 'graph':
feat_length = data.iloc[0].mol.ndata['feats'].shape[-1]
print("Feature Length", feat_length)
if with_bonds:
feat_length = data.iloc[0].mol.edata['edge_feats'].shape[-1]
print("Feature Length", feat_length)
else:
print("Edge Features: ", with_bonds)
test_dfs = dict.fromkeys(test_tasks)
for task in test_tasks:
df = data[[task, 'mol']].dropna()
df.columns = ['y', 'mol']
# FOR BOND INFORMATION
if with_bonds:
for index, r in df.iterrows():
if r.mol.edata['edge_feats'].shape[-1] < 17:
df.drop(index, inplace=True)
test_dfs[task] = df
for key in test_dfs:
print(key, len(test_dfs[key]))
# return data, train_tasks, test_tasks
return train_dfs, test_dfs
# + [markdown] id="J6bY9GX7rR59"
# ## Initiate Training and Testing
# + colab={"base_uri": "https://localhost:8080/"} id="8MxSYcE4rRE5" executionInfo={"status": "ok", "timestamp": 1638271601512, "user_tz": -60, "elapsed": 73325, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitatTDk380OCMvv2bDBT7Lsunn79WNpphzKI_dytU=s64", "userId": "08737126295476082217"}} outputId="78cbee46-b58d-4a8a-8f4a-2a2b9a106900"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="g8AWDKNZpjAs" executionInfo={"status": "ok", "timestamp": 1638271601512, "user_tz": -60, "elapsed": 10, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitatTDk380OCMvv2bDBT7Lsunn79WNpphzKI_dytU=s64", "userId": "08737126295476082217"}} outputId="9f422790-53c0-46d6-952a-a6fbdb7abde6"
drive_path = "/content/drive/MyDrive/Colab Notebooks/MSC_21"
method_dir = "RandomForest"
log_path = f"{drive_path}/{method_dir}/logs/"
# PARAMETERS
# dude-gprc, tox21, muv
dataset = 'dude-gpcr'
with_bonds = False
rounds = 20
n_query = 64 # per class
episodes = 10000
lr = 0.001
balanced_queries = True
randomseed = 12
torch.manual_seed(randomseed)
np.random.seed(randomseed)
random.seed(randomseed)
torch.cuda.manual_seed(randomseed)
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
torch.backends.cudnn.is_available()
torch.backends.cudnn.benchmark = False # selects fastest conv algo
torch.backends.cudnn.deterministic = True
combinations = [
[10, 10],
[5, 10],
[1, 10],
[1, 5],
[1, 1]
]
cols = [
'DATE', 'CPU', 'CPU COUNT', 'GPU', 'GPU RAM', 'RAM', 'CUDA',
'REF', 'DATASET', 'ARCHITECTURE',
'SPLIT', 'TARGET', 'ACCURACY', 'ROC', 'PRC',
'TRAIN ROC', 'TRAIN PRC', 'EPISODES', 'TRAINING TIME', 'ROC_VALUES', 'PRC_VALUES'
]
# + colab={"base_uri": "https://localhost:8080/"} id="6Xdm-H_RrgQ1" executionInfo={"status": "ok", "timestamp": 1638271623776, "user_tz": -60, "elapsed": 6868, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitatTDk380OCMvv2bDBT7Lsunn79WNpphzKI_dytU=s64", "userId": "08737126295476082217"}} outputId="131a6af1-f76e-4848-89e7-0aa2ea997914"
train_dfs, test_dfs = load_dataset(dataset, bonds=with_bonds, feat='ecfp', create_new=False)
# + colab={"base_uri": "https://localhost:8080/"} id="c5C81CHZr_f6" executionInfo={"status": "ok", "timestamp": 1638271681317, "user_tz": -60, "elapsed": 46712, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitatTDk380OCMvv2bDBT7Lsunn79WNpphzKI_dytU=s64", "userId": "08737126295476082217"}} outputId="aac923c1-fff2-45fd-89ad-ab7613ef46d7"
dt_string = datetime.now().strftime("%d/%m/%Y %H:%M:%S")
cpu = get_cmd_output('cat /proc/cpuinfo | grep -E "model name"')
cpu = cpu.split('\n')[0].split('\t: ')[-1]
cpu_count = psutil.cpu_count()
cuda_version = get_cmd_output('nvcc --version | grep -E "Build"')
gpu = get_cmd_output("nvidia-smi -L")
general_ram_gb = humanize.naturalsize(psutil.virtual_memory().available)
gpu_ram_total_mb = GPU.getGPUs()[0].memoryTotal
for c in combinations:
n_pos = c[0]
n_neg = c[1]
results = pd.DataFrame(columns=cols)
for target in test_dfs.keys():
print(target)
running_roc = []
running_prc = []
for round in trange(rounds):
start_time = time.time()
df = test_dfs[target]
support_neg = df[df['y'] == 0].sample(n_neg)
support_pos = df[df['y'] == 1].sample(n_pos)
train_data = pd.concat([support_neg, support_pos])
test_data = df.drop(train_data.index)
train_data = train_data.sample(frac=1)
test_data = test_data.sample(frac=1)
train_X, train_y = list(train_data['mol'].to_numpy()), train_data['y'].to_numpy(dtype=np.int16)
test_X, test_y = list(test_data['mol'].to_numpy()), test_data['y'].to_numpy(dtype=np.int16)
model = RandomForestClassifier(n_estimators=100)
model.fit(train_X, train_y)
probs_y = model.predict_proba(test_X)
roc = roc_auc_score(test_y, probs_y[:, 1])
prc = average_precision_score(test_y, probs_y[:, 1])
running_roc.append(roc)
running_prc.append(prc)
end_time = time.time()
duration = str(timedelta(seconds=(end_time - start_time)))
rounds_roc = f"{statistics.mean(running_roc):.3f} \u00B1 {statistics.stdev(running_roc):.3f}"
rounds_prc = f"{statistics.mean(running_prc):.3f} \u00B1 {statistics.stdev(running_prc):.3f}"
rec = pd.DataFrame([[dt_string, cpu, cpu_count, gpu, gpu_ram_total_mb, general_ram_gb, cuda_version, "MSC",
dataset, method_dir, f"{n_pos}+/{n_neg}-", target, 0, rounds_roc, rounds_prc,
0, 0, 0, duration, running_roc, running_prc
]], columns=cols)
results = pd.concat([results, rec])
results.to_csv(f"{drive_path}/results/{dataset}_{method_dir}_pos{n_pos}_neg{n_neg}.csv", index=False)
# + colab={"base_uri": "https://localhost:8080/"} id="Lq-S6oaOs6kU" executionInfo={"status": "ok", "timestamp": 1637840745602, "user_tz": -60, "elapsed": 258, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitatTDk380OCMvv2bDBT7Lsunn79WNpphzKI_dytU=s64", "userId": "08737126295476082217"}} outputId="3cf05ea7-b99c-41c8-cc6d-909a9aef6c57"
# model.score(test_X, test_y)
# pred_y = model.predict(test_X)
# model.classes_
# + colab={"base_uri": "https://localhost:8080/"} id="U_1C5YfR1V-0" executionInfo={"status": "ok", "timestamp": 1638271682316, "user_tz": -60, "elapsed": 8, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitatTDk380OCMvv2bDBT7Lsunn79WNpphzKI_dytU=s64", "userId": "08737126295476082217"}} outputId="b59b5d2d-baad-424a-f06e-1d64bdea8d74"
from sklearn.metrics import confusion_matrix
preds = model.predict(test_X)
confusion_matrix(test_y, preds)
# + colab={"base_uri": "https://localhost:8080/", "height": 315} id="DCvJi8XW1r-1" executionInfo={"status": "ok", "timestamp": 1638271682782, "user_tz": -60, "elapsed": 471, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitatTDk380OCMvv2bDBT7Lsunn79WNpphzKI_dytU=s64", "userId": "08737126295476082217"}} outputId="1754319f-475a-4bda-efdc-58fef7a19c31"
ConfusionMatrixDisplay.from_predictions(test_y, preds)
| Random Forest Benchmark.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import plotly
plotly.offline.init_notebook_mode(connected=True)
# # Basic Sankey Chart
# +
data = dict(
type = 'sankey',
# 设置每个柱状条的颜色,宽度,边框等信息
node = dict(
thickness = 20,
line = dict(color = "black", width = 1),
label = ["A1", "A2", "B1", "B2", "C1", "C2"],
color = ["#e03636", "#edd08e", "#ff534d", "#23d58e", "#25c6fc", "#0000ff"]
),
# 设置能量流动条的宽度值
link = dict(
source = [0, 0, 1, 2, 3, 3],
target = [2, 3, 3, 4, 4, 5],
value = [8, 4, 2, 8, 4, 2]
)
)
layout = dict(
title = "基础桑吉图",
font = dict(size = 20)
)
fig = dict(data=[data], layout=layout)
plotly.offline.iplot(fig, validate=False)
# -
# # Advance Sankey Chart
# +
data_trace = dict(
type = 'sankey',
node = dict(
pad = 15,
thickness = 15,
line = dict(color = "black", width = 0.5),
label = ["专业A", "专业B", "专业C", "专业D", "专业E", "转到其他学院","留在本学院","国内升学","国外深造","直接就业","待就业"]
),
link = dict(
source = [0,0,1,1,2,2,3,3,4,4,6,6,6,6,5,5,5,5],
target = [5,6,5,6,5,6,5,6,5,6,7,8,9,10,7,8,9,10],
value = [5,25,6,25,4,40,7,42,6,40,62,37,70,3,5,15,7,1]
)
)
layout = dict(
title = "学院学生转专业统计 & 毕业去向统计",
font = dict(
size = 15,
color = 'black'
),
# 修改视图背景色
# plot_bgcolor = 'black',
# paper_bgcolor = 'black'
)
fig = dict(data=[data_trace], layout=layout)
plotly.offline.iplot(fig, validate=False)
| PythonPracticalCases/Plotly/sankey.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img style="float: right; margin: 0px 0px 15px 15px;"
# src="MontecarloImag.jpeg" width="300px" height="100px" />
# # Técnicas de reducción de varianza
#
# **Objetivos:**
# 1. Estudiar el método de integración montecarlos para aproximación de integrales.
# 2. Estudiar dos técnicas para reducción de varianza
# - Muestreo estratificado
# - Método de números aleatorios complementarios
#
# ## 1. Integración numérica Montecarlo
#
# En esta clase veremos una técnica alternativa de integración numérica (métodos numéricos): Integración Montecarlo
#
# ### Introducción
#
# En matempaticas financieras un problema frecuente es el valuar instrumentos financieros cuyos rendimientos son aleatorios. Por ejemplo los instrumentos de renta variable, las inversiones en la bolsa o los derivados, cuyos rendimientos dependen del comportamiento de una acción o de un bien como el oro o el petróleo. La valuación de estos instrumentos se reduce, al cálculo de una esperanza de una función continua de una variable aleatoria.
#
# Recordando algunos conceptos de probabilidad. Sea **$x$ una variable aleatoria continua**, si su función de densidad de probabilidad es $f(x)$, en un intervalo $[\alpha,\beta]$, entonces
#
# $$ P(X\leq c) = \int_\alpha^c f(x) dx$$
#
# $$ \textbf{Esperanza}\rightarrow E(g(x))=\int_\alpha^\beta g(x)f(x) dx $$
# $$ \textbf{Varianza}\rightarrow Var(g(x))=E(g(x)^2)-E(g(x))^2=\int_\alpha^\beta g(x)^2f(x) dx -E(x)^2$$
#
# Con frecuencia no es posible aplicar un método de integración para calcular en forma exacta la integral. En ese caso hay que aproximar la integral por medio de un método de integración numérico o por monte carlo.
# ## Método monte carlo
#
# Se sabe por la Ley de los Grandes Números que un buen estimador del valor esperado de una variable aleatoria continua $X$ con distribución $F$ es el valor promedio de una muestra finita de variables aleatorias, independientes con distribución $F$: Es decir
# $$ E(X)\approx \frac{1}{M}\sum_{i=1}^M X_i$$
#
# Como la esperanza de una variable aleatoria continua es una integral, la media muestral se puede usar para estimar el valor de una integral. Esta es la idea que está detrás del método de Monte-Carlo.
#
# Esta idea se puede generalizar para estimar el valor esperado de una función $G$ continua cuyo argumento es una variable aleatoria con distribución $F$: Si se tiene una muestra de variables aleatorias, independientes, idénticamente distribuidas con distribución $F$; entonces
#
# $$ E(G(X))\approx \frac{1}{M}\sum_{i=1}^M G(X_i)$$
# ### Aplicación al cálculo de integrales
#
# En el caso de interés, se desea estimar la integral de una función $G$ continua, esta integral puede verse como el cálculo del valor esperado de la función $G$ cuando se aplica a una variable aleatoria con distribución uniforme. Supongamos que el intervalo de integración es $[0, 1]$ y sea $x_1, x_2, \cdots, x_M$ una muestra de variables aleatorias, independientes
# con distribución uniforme en el intervalo $[0, 1]$ entonces:
#
# $$ \int_0^1G(x)dx = E(G(x)) \approx \frac{1}{M}\sum_{i=1}^M G(x_i)$$
#
# ** Todo el problema se reduce a generar la muestra**.
#
# Por otro lado observe que toda integral en el intervalo $[a,b]$ se puede transformar a una integral sobre el intervalo $[0,1]$, con el siguiente cambio de variable $u = \frac{x-a}{b-a} \rightarrow x = a+(b-a)u$ con $dx=(b-a)du$, entonces
#
# $\int_a^b G(x)dx= (b-a)\int\limits_0^1 G(a+(b-a)u)du\approx \frac{(b-a)}{M}\sum\limits_{i=1}^M G(\underbrace{a+(b-a)u_i}_{variable \ U\sim[a,b]})$
#
# con $u_i$ variables aleatorias uniformes en el intervalo $[0,1]$
# Escribamos una función que tenga como entradas:
# - la función a integrar $f$,
# - los límites de integración $a$ y $b$, y
# - los números aleatorios distribuidos uniformemente entre $[a,b]$,
#
# y que devuelva la aproximación montecarlo de la integral $\int_{a}^{b}f(x)\text{d}x$.
import numpy as np
from functools import reduce
import time
import matplotlib.pyplot as plt
import scipy.stats as st # Librería estadística
import pandas as pd
# Integración montecarlo
def int_montecarlo(f:'Función a integrar',
a:'Límite inferior de la integral',
b:'Límite superior de la integral',
U:'Muestra de números U~[a,b]'):
return (b-a)/len(U)*np.sum(f(U))
# ### Ejemplo
#
# Aproxime el valor de la siguiente integral usando el método monte carlo
#
# $$I=\int_{0}^{1}x^2\text{d}x=\left.\frac{x^3}{3}\right|_{x=0}^{x=1}=\frac{1}{3}\approx 0.33333$$
# +
I = 1/3
# intervalo de integración
a = 0; b = 1
# Cantidad de términos, en escala logarítmica
N = np.logspace(1,7,7,dtype=int)
# Definimos la tabla donde se mostrarán los resultados
df = pd.DataFrame(index=N,columns=['Valor_aproximacion', 'Error_relativo%'], dtype='float')
df.index.name = "Cantidad_terminos"
# Números aleatorios dependiente de la cantidad de términos N
ui = list(map(lambda N:np.random.uniform(a,b,N),N))
# Calculamos la aproximación por montecarlo dependiendo de la cantidad de
# términos que hayamos creado con ui
I_m = list(map(lambda Y:int_montecarlo(lambda x:x**2,a,b,Y),ui))
# Mostramos los resultados en la tabla previamente creada
df.loc[N,"Valor_aproximacion"] = I_m
df.loc[N,"Error_relativo%"] = np.abs(df.loc[N,"Valor_aproximacion"]-I)*100/I
df
# -
# ### Nota:
# Sean $I=E(g(x))$ y el estimado de $I$ es $\tilde I_M = \frac{1}{M}\sum\limits_{i=1}^M g(X_i)$. Tome $\sigma$ como la desviación estándar de $g(X)$ y $\tilde \sigma$ como la desviación estándar muestral. Por lo tanto, se tiene entonces que:
#
# $$ Var(\tilde I_M)=\frac{\sigma^2}{M} \longrightarrow std(\tilde I_M)=\frac{\sigma}{\sqrt{M}}\longrightarrow \text{Dispersión disminuye con la cantidad de datos}$$
#
# Típicamente no se tiene conocimiento de $\sigma$ por lo tanto se puede estimar por medio de la varianza muestral.
# $$\tilde \sigma^2=\frac{1}{M-1}\sum\limits_{i=1}^{M}(g(X_i)-\tilde I_M)^2$$
#
# ### Conclusión
#
# Si se desea disminuir el error de estimación de la integral (desviación estándar), hay dos caminos:
# 1. Aumentar la cantidad de muestras a evaluar en la función, **(Proceso lento)**, o
# 2. Utilizar técnicas para disminuir la varianza.
# # 2. Reducción de varianza
#
# ## a. Muestreo estratificado
#
# La idea radica en la observación de que la población puede ser **heterogénea** y consta de varios subgrupos homogéneos (como género, raza, estado económico social). Si deseamos aprender sobre toda la población (como por ejemplo, si a la gente en México le gustaría votar para las elecciones en 2018), podemos tomar una muestra aleatoria de toda la población para estimar esa cantidad. Por otro lado, sería más eficiente tomar muestras pequeñas de cada subgrupo y combinar las estimaciones en cada subgrupo según la fracción de la población representada por el subgrupo. Dado que podemos conocer la opinión de un subgrupo homogéneo con un tamaño de muestra relativamente pequeño, este procedimiento de muestreo estratificado sería más eficiente.
#
# Si nosotros deseamos estimar $E(X)$, donde $X$ depende de una variable aleatoria $S$ que toma uno de los valores en $\{1, ...,k\}$ con probabilidades conocidas, entonces la técnica de estratificación se ejecuta en k grupos, con el i-ésimo grupo que tiene $S = i$, tomando $\bar X_i$ es el valor promedio de $X$ cuando se estuvo en el grupo $S = i$, y luego se estima $E(x)$ por:
# $$E(x)=\sum\limits_{i=1}^{k}\underbrace{E(X|S=i)}_{\text{media muestral del i-ésimo grupo}}P(S=i)=\sum\limits_{i=1}^{k} \bar X_i P(S=i)$$
#
# Para ilustrar considere que queremos estimar $E(g(U))=\int_0^1g(x)dx$. Para esto vamos a considerar dos estimadores basados en una muestra de 2n corridas.
#
# 1.**Método estándar (visto anteriormente)**
# $$\textbf{media}\rightarrow \hat g=\frac{1}{2n}\sum\limits_{i=1}^{2n} g(U_i)$$
#
# $$\textbf{varianza}\rightarrow Var(\hat g)=\frac{1}{4n^2}\sum\limits_{i=1}^{2n} Var(g(U_i))=\frac{1}{2n}\bigg[\int_0^1g^2(x)dx-\bigg(\int_0^1g(x)dx\bigg)^2\bigg]$$
# Por otro lado nosotros podemos escribir
#
# $$E(g(U))=\int_0^{1/2}g(x)dx + \int_{1/2}^1g(x)dx$$
#
# - Seleccionamos $n$ U's de $[0,1/2]$
# - Seleccionamos $n$ U's de $[1/2,1]$
#
# 2.Con esta nueva selección construimos el **estimador estratificado**:
#
# $$\textbf{media}\rightarrow \hat g_s=\frac{1}{2n}\big[\sum\limits_{i=1}^{n} g(\underbrace{U_i/2}_{U\sim [0,1/2]}) + \sum\limits_{i=n+1}^{2n} g\big((\underbrace{U_i+1)/2}_{U\sim [1/2,1]}\big)\big]$$
#
# $$\textbf{varianza}\rightarrow Var(\hat g_s)=\frac{1}{4n^2}\big[\sum\limits_{i=1}^{n} Var(g(U_i/2)) + \sum\limits_{i=n+1}^{2n} Var(g\big((U_i+1)/2\big))\big]$$
#
# Realizando el cálculo directo tomando $U_i\sim U(0,1)$, se puede mostrar que:
#
# $$Var(g(U_i/2)) = 2 \int_0^{1/2} g^2(x)dx - 4m_1^2,$$
# $$Var(g((U_i+1)/2)) = 2 \int_{1/2}^1 g^2(x)dx - 4m_2^2,$$
#
# donde $m_1 = \int_0^{1/2}g(x)dx$ y $m_2 = \int_{1/2}^1g(x)dx$
#
# Realizando varias manipulaciones algebraicas se llega a que
#
# > $$Var(\hat g_s) = Var(\hat g)-\frac{1}{2n}(m_1-m_2)^2$$
#
# <font color ='red'> La varianza se disminuye con respecto a $Var(\hat g)$. Note también que en el ejemplo anterior se consideraron dos estratos, por eso únicamente aparecen $m_1$, $m_2$, pero si se hubieran tomado $n$ estratos, tendríamos que la reducción de varianza sería
# > $$Var(\hat g_s) = Var(\hat g)-\frac{1}{2n}(m_1-m_2-m_3-\cdots - m_n)^2$$
#
# > **Referencia**: Handbook in Monte Carlo simulation applications in financial engineering, risk management, and economics, pág. 97.
# ### Ejemplo
# Como ilustración de la estratificación, considere la simulación de números aleatorios normales $\mathcal{N}\sim (0,1)$, mediante el método estándar y el método de estratificación, respectivamente.
# +
np.random.seed(5555)
muestras = np.random.normal(size=10)
muestras2 = np.random.normal(np.random.rand(10))
muestras,muestras2
np.random.normal(.01),np.random.normal(1)
# +
np.random.seed(5555)
# Muestras
N = 10000
#### Comparar los resultados con cada una de las siguientes expresiones
muestras2 = np.random.normal(np.ones(N))
muestras = np.random.normal(np.random.rand(N))
# Cálculo de media
t1 = time.time()
g_hat = (sum(muestras))/len(muestras)
t2 = time.time()
print('Tiempo usando la fórmula teórica de la media=%2.6f, cuyo valor es=%2.5f' %(t2-t1,g_hat))
t1 = time.time()
g_hat2 = np.mean(muestras)
t2 = time.time()
print('Tiempo calculado usando la función de numpy=%2.6f, cuyo valor es=%2.5f' %(t2-t1,g_hat2))
# cálculo de la varianza
t1 = time.time()
# varg_hat = sum(list(map(lambda l:l**2,muestras-g_hat)))/len(muestras)
varg_hat = np.mean(list(map(lambda l:l**2,muestras-g_hat)))
t2 = time.time()
print('Tiempo calculado usando la varianza 1 (Teórica)=',t2-t1)
t1 = time.time()
var_numpy = np.var(muestras)
t2 = time.time()
print('Tiempo calculado usando la varianza 2 (numpy)=',t2-t1)
print('Var. con función creada = %2.6f y Var. con función de numpy %2.6f' %(varg_hat,var_numpy))
# Histogramas para las dos muestras
fig,(ax1,ax2) = plt.subplots(1,2)
fig.set_figwidth(15)
ax1.hist(muestras,100,density=True)
ax1.set_title('USando el comando random.normal(random.rand(N))')
ax2.hist(muestras2,100,density=True)
ax2.set_title('USando el comando random.normal(np.ones(N))')
plt.show()
# -
# # Esbozo del método de muestreo estratificado
# 
# +
N = 500
# (Método transformada inversa visto en clases pasadas)
# Método montecarlo estándar
montecarlo_standar = st.norm.ppf(np.random.rand(N))
plt.hist(montecarlo_standar,30,density=True)
plt.title('Montecarlo estándar')
plt.xlim((-3.5,3.5))
plt.show()
# Método muestreo estratificado (B estratos)
B = 300
U2 = np.random.rand(B)
i = np.arange(0,B)
v = (U2+i)/B
m_estratificado = st.norm.ppf(v)
plt.hist(m_estratificado,30,density=True)
plt.title('Muestreo estratificado')
plt.xlim((-3.5,3.5))
plt.show()
# -
# ### Ejemplo
# Estimar la media de la siguiente función de distribución de exponencial:
# $$\textbf{Densidad}\rightarrow f(x)=e^{-x}\rightarrow \text{media = 1}$$
# $$\textbf{Acumulada}\rightarrow F(x)=1-e^{-x}$$
#
# Usando el método de la transformada inversa que realizamos en clases pasadas, obtuvimos que para generar números aleatorios de esta distribución tenía que aplicarse $x_i=-ln(u_i)$ con $u_i \sim U[0,1]$.
#
# Suponga entonces que generamos 10 muestras y con eso intentamos estimar la media de esta función de distribución, como se muestra a continuación:
# +
np.random.seed(55555)
ui = np.random.rand(10)
i = np.arange(1,11)
# Definimos la tabla donde se mostrarán los resultados
df = pd.DataFrame(index= i,columns=['Numero_Aleatorio', 'Observacion_generada','Media_muestral'], dtype='float')
df.index.name = "Cantidad_terminos"
xi = -np.log(ui)
media_no_estrato = np.mean(xi)
# Mostramos los resultados en la tabla previamente creada
df.loc[len(i),"Media_muestral"] ='Media muestral = %f'%media_no_estrato
df.loc[i,"Numero_Aleatorio"] = ui
df.loc[i,"Observacion_generada"] = xi
# print('La media de la muestra es=',np.mean(xi)
df
# -
# ### Usando muestreo estratificado
#
# 
# 
# +
np.random.seed(5555)
r1 = np.random.uniform(0,0.64,4)
r2 = np.random.uniform(0.64,0.96,4)
r3 = np.random.uniform(0.96,1,2)
r = [r1,r2,r3]
m = range(len(r)) # Cantidad de estratos
w = [5/8,5/4,5]
xi = list(map(lambda r:-np.log(r),r))
muestras = list(map(lambda wi,xi:xi/wi,w,xi))
# muestras = np.concatenate([(xi[i]/w[i]).tolist() for i in m])
# Definimos la tabla donde se mostrarán los resultados
i = np.arange(1,11)
df = pd.DataFrame(index= i,columns=['Numero_Aleatorio_Estrato','Observacion_generada','xi/w'], dtype='float')
df.index.name = "términos"
# Mostramos los resultados en la tabla previamente creada
df.loc[i,"Numero_Aleatorio_Estrato"] = np.concatenate(r)
df.loc[i,"Observacion_generada"] = np.concatenate(xi)
df.loc[i,"xi/w"] = np.concatenate(muestras)
print('La media de la muestra es=',np.concatenate(muestras).mean())
df
# -
# np.concatenate(muestras).mean()
np.concatenate(muestras).mean()
muestras
media_estrato = np.concatenate(muestras).mean()
print('La media muestral estimada método estratificado es = ',media_estrato)
print('La media muestral estimada método no estratificado es = ',media_no_estrato)
print('Error relativo método estratificado = ',(1-media_estrato)*100)
print('Error relativo método no estratificado = ',(1-media_no_estrato)*100)
# # Tarea
#
# Aproxime el valor de la siguiente integral usando el método monte carlo crudo y método de reducción de varianza de muestreo estratíficado
#
# $$I=\int_{0}^{1}x^2\text{d}x=\left.\frac{x^3}{3}\right|_{x=0}^{x=1}=\frac{1}{3}\approx 0.33333$$
#
# Pasos
# 1. Cree una función que realice el método de muestreo estratíficado, recibiendo como único parámetro de entrada la cantidad de estratos y retornando las variables estratíficadas correspondientes.
# 2. Reporte los resultados de la aproximación de la integral usando montecarlo crudo y muestreo estratíficado, en un Dataframe con la información mostrada en la siguiente imagen:
# 
# ### Parámetros de entrega
# Se debe de entregar este ejercicio para el próximo martes 22 de octubre hasta las 11 pm, en un link que se habilitará en canvas.
# <script>
# $(document).ready(function(){
# $('div.prompt').hide();
# $('div.back-to-top').hide();
# $('nav#menubar').hide();
# $('.breadcrumb').hide();
# $('.hidden-print').hide();
# });
# </script>
#
# <footer id="attribution" style="float:right; color:#808080; background:#fff;">
# Created with Jupyter by <NAME>.
# </footer>
| TEMA-2/Clase13_MetodosDeReduccionDeVarianza.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import numpy as np
import matplotlib.pyplot as plt
img = cv2.imread('./samples/sample_receipt.png')
ret,thresh1 = cv2.threshold(img,210,255,cv2.THRESH_BINARY)
plt.imshow(thresh1, 'gray')
| .ipynb_checkpoints/Untitled-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Word2Seq Convolutional Neural Network
# Importing all the required libraries along with some instance variables initialization.
#
# It is important to note that **INPUT_SIZE** plays a big role in the model convergence.
# The value should be the average number of words per review.
# This shall yield the most optimum model state for faster convergence across the network.
#
# Tested values for *INPUT_SIZE* :
# * 700 (11 mins per epoch ) = 93.24%
#
# CNN able to handle the cushion values of input sequence not like RNN.
# RNN require more to specific number of INPUT_SIZE as compared to CNN.
# This shows the robustiness of CNN when dealing with variable length senteneces.
#
# **EPOCHS** is set to `10` only to select the best model with the best hyper-parameters configuration for the fastest network convergence.
# +
import pandas as pd
import numpy as np
import pickle
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
import tensorflow as tf
from keras.backend.tensorflow_backend import set_session
from tensorflow.python.keras.models import Sequential, load_model
from tensorflow.python.keras.layers import Dense, Embedding, Dropout, Flatten
from tensorflow.python.keras.layers import Conv1D, MaxPool1D
from tensorflow.python.keras import optimizers
from tensorflow.python.keras.preprocessing import text as keras_text, sequence as keras_seq
from tensorflow.python.keras.utils import to_categorical
from tensorflow.python.keras.callbacks import TensorBoard, ModelCheckpoint
from sklearn.metrics import confusion_matrix,accuracy_score,recall_score,precision_score,f1_score
from sklearn.utils import shuffle
from tensorflow import set_random_seed
import gc
import os
#myrand=np.random.randint(1, 99999 + 1)
myrand=58584
np.random.seed(myrand)
set_random_seed(myrand)
z=0
EMBEDDING_SIZE=32
WORDS_SIZE=8000
INPUT_SIZE=700
NUM_CLASSES=2
EPOCHS=10
# To allow dynamic GPU memory allowcation for model training
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
config.log_device_placement = True
sess = tf.Session(config=config)
set_session(sess)
# -
# Importing all the data from:
# * eRezeki
# * IMDB
# * Amazon
# * Yelp
#
# +
mydata = pd.read_csv('../../../../../Master (Sentiment Analysis)/Paper/Paper 3/Datasets/eRezeki/eRezeki_(text_class)_unclean.csv',header=0,encoding='utf-8')
mydata = mydata.loc[mydata['sentiment'] != "neutral"]
mydata['sentiment'] = mydata['sentiment'].map({'negative': 0, 'positive': 1})
mydata1 = pd.read_csv('../../../../../Master (Sentiment Analysis)/Paper/Paper 3/Datasets/IMDB/all_random.csv',header=0,encoding='utf-8')
mydata = mydata.append(mydata1)
mydata = shuffle(mydata)
mydata1 = pd.read_csv('../../../../../Master (Sentiment Analysis)/Paper/Paper 3/Datasets/Amazon(sports_outdoors)/Amazon_UCSD.csv',header=0,encoding='utf-8')
mydata1['feedback'] = mydata1['feedback'].astype(str)
mydata = mydata.append(mydata1)
mydata = shuffle(mydata)
mydata1 = pd.read_csv('../../../../../Master (Sentiment Analysis)/Paper/Paper 3/Datasets/Yelp(zhang_paper)/yelp_zhang.csv',header=0,encoding='utf-8')
mydata1['feedback'] = mydata1['feedback'].astype(str)
mydata = mydata.append(mydata1)
del(mydata1)
gc.collect()
mydata = shuffle(mydata)
mydata = shuffle(mydata)
mydata = shuffle(mydata)
# -
# Spllitting the data into training (70%) and testing (30%) sets.
x_train, x_test, y_train, y_test = train_test_split(mydata.iloc[:,0], mydata.iloc[:,1],
test_size=0.3,
random_state=myrand,
shuffle=True)
old_y_test = y_test
# Load the Word2Seq feature set as the tokneizer
tokenizer = keras_text.Tokenizer(char_level=False)
tokenizer.fit_on_texts(list(mydata['feedback']))
tokenizer.num_words=WORDS_SIZE
# Create sequence file from the tokkenizer for training and testing sets.
# +
## Tokkenizing train data and create matrix
list_tokenized_train = tokenizer.texts_to_sequences(x_train)
x_train = keras_seq.pad_sequences(list_tokenized_train,
maxlen=INPUT_SIZE,
padding='post')
x_train = x_train.astype(np.int64)
## Tokkenizing test data and create matrix
list_tokenized_test = tokenizer.texts_to_sequences(x_test)
x_test = keras_seq.pad_sequences(list_tokenized_test,
maxlen=INPUT_SIZE,
padding='post')
x_test = x_test.astype(np.int64)
# -
# Perform One Hot Encoding (OHE) to the labes of training and testing sets.
y_train = to_categorical(y_train, num_classes=NUM_CLASSES).astype(np.int64)
y_test = to_categorical(y_test, num_classes=NUM_CLASSES).astype(np.int64)
# Define and build the **Word2Seq CNN** model
# +
model = Sequential(name='Word2Seq CNN')
model.add(Embedding(input_dim =WORDS_SIZE,
output_dim=250,
input_length=INPUT_SIZE
))
model.add(Conv1D(filters=250, kernel_size=3, padding='same', activation='relu'))
model.add(MaxPool1D(pool_size=50))
model.add(Conv1D(filters=250, kernel_size=2, padding='same', activation='relu'))
model.add(MaxPool1D(pool_size=10))
model.add(Flatten())
model.add(Dense(250, activation='relu'))
model.add(Dense(2, activation='softmax'))
## Define multiple optional optimizers
sgd = optimizers.SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
adam = optimizers.Adam(lr=0.01, beta_1=0.9, beta_2=0.999, epsilon=1, decay=0.0, amsgrad=False)
## Compile model with metrics
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
print("Word2Seq CNN model built: ")
model.summary()
# -
# Create **TensorBoard callbacks** for:
# * Historical data
# * Training and evaluation gain/loss
#
# Also, create the **best-model callback** to save the **best** model every epoch.
# +
## Create TensorBoard callbacks
callbackdir= '/project/ten'
tbCallback = TensorBoard(log_dir=callbackdir,
histogram_freq=0,
batch_size=128,
write_graph=True,
write_grads=True,
write_images=True)
tbCallback.set_model(model)
mld = '/project/model/word2seq_cnn.hdf5_%s'%(z)
## Create best model callback
mcp = ModelCheckpoint(filepath=mld, monitor="val_acc",
save_best_only=True, mode='max', period=1, verbose=1)
# -
# **Train** the model
print('Training the Word2Seq CNN model')
history = model.fit(x = x_train,
y = y_train,
validation_data = (x_test, y_test),
epochs = EPOCHS,
batch_size = 128,
verbose =2,
callbacks=[mcp,tbCallback])
# **Predict** the testing set using the best model from the run.
print('\nPredicting the model')
model = load_model(mld)
results = model.evaluate(x_test, y_test, batch_size=128)
for num in range(0,2):
print(model.metrics_names[num]+': '+str(results[num]))
# **Evaluate** the performance of the model on the testing set using:
# * Confusion matrix
# * Accuracy
# * Precision
# * Recall
# * F-Measure
# +
print('\nConfusion Matrix')
predicted = model.predict_classes(x_test)
confusion = confusion_matrix(y_true=old_y_test, y_pred=predicted)
print(confusion)
## Performance measure
print('\nWeighted Accuracy: '+ str(accuracy_score(y_true=old_y_test, y_pred=predicted)))
print('Weighted precision: '+ str(precision_score(y_true=old_y_test, y_pred=predicted, average='weighted')))
print('Weighted recall: '+ str(recall_score(y_true=old_y_test, y_pred=predicted, average='weighted')))
print('Weighted f-measure: '+ str(f1_score(y_true=old_y_test, y_pred=predicted, average='weighted')))
# -
# **Plot** the model training history for training and evaluation sets.
# +
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs_range = range(len(acc))
plt.plot(epochs_range, acc, 'bo', label='Training acc')
plt.plot(epochs_range, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs_range, loss, 'bo', label='Training loss')
plt.plot(epochs_range, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
| Scripts/Word2Seq_CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # System and C/C++ Software Security
#
# ## Table of Contents
#
#
# ### Linux and Bash Scripting
# - **[Setup Hacking VM](./SetupVM.ipynb)**
# - **[Linux Command Fundamentals](https://github.com/rambasnet/Linux-Commands)**
# - **[Bash Script Fundamentals](https://github.com/rambasnet/Bash-Script-Fundamentals)**
#
# ### Python Programming
# - **[Python Basics](./PythonBasics.ipynb)**
# - **[Python In-Depth](https://github.com/rambasnet/Python-Fundamentals)**
# - **[Python Unittest](https://github.com/rambasnet/Kattis-Demos)**
#
# ### C++ Programming
# - **[CS1 Review](./CS1-Review.ipynb)**
# - **[Functions & Unit testing](./FunctionsAndUnittesting.ipynb)**
# - **[C Arrays](./C-Arrays.ipynb)**
# - **[Pointers & Applications](./Pointers.ipynb)**
# - **[C Strings - Buffer](./C-Strings.ipynb)**
# - **[Function Pointers](./Function-Pointers.ipynb)**
#
# #### C++ In-Depth
# - **[C++ Fundamentals Notebooks](https://github.com/rambasnet/CPP-Fundamentals)**
# - **[Data Structures with C++ Notebooks](https://github.com/rambasnet/Data-Structures)**
# - **[C++ STL Notebooks](https://github.com/rambasnet/CPP-STL)**
#
# ### Penetration Testing
# - Great resoures for hacking tips and resources: [https://www.hackers-arise.com/](https://www.hackers-arise.com/)
# - **[Hacking Tools](./HackingTools.ipynb)**
# - **[Pen Testing](./PenTesting.ipynb)**
#
# ### Under the Hood
# - **[C/C++ Program Memory Segments](./MemorySegments.ipynb)**
# - **[ELF Binary Format & Reverse Engineering](./ELF.ipynb)**
# - **[x86 Assembly and Stack](./x86-AssemblyAndStack.ipynb)**
# - **[GDB & PEDA](./GDB-Peda.ipynb)**
#
# ### Memory Corruption Vulnerabilities
# - **[Buffer Overflow Basics](./BufferOverflowBasics.ipynb)**
# - **[Detecting and Mitigating Memory Corruption Errors](./DetectingMemoryCorruptionErrors.ipynb)**
# - **[Buffer Overflow Protections](./BufferOverflowProtections.ipynb)**
#
# ### Stack Overflow Exploitations
# - **[Stack Overflow - Memory Corruption](./StackOverflow-MemoryCorruption.ipynb)**
# - **[Stack Overflow - Execution Flow Corruption](./StackOverflow-ExecutionFlow.ipynb)**
# - **[Stack Overflow - Remote Code Execution](./StackOverflow-RemoteCodeExecution.ipynb)**
# - **[Exploiting with Env Variable](./StackOverflow-EnvVariable.ipynb)**
#
# ### Exploit Code
# - **[Exploit code using Bash Script](./ExploitCode-UsingBashScript.ipynb)**
# - **[Exploit code using C++](./ExploitCode-UsingCPP.ipynb)**
# - **[Pwntools Tutorial](./PwntoolsTutorials.ipynb)**
# - **[Exploit code using Python Script](./ExploitCode-UsingPythonScript.ipynb)**
#
# ### Heap Overflow
# - **[Heap Overflow Exploitation](./HeapOverflowExploitation.ipynb)**
#
# ### Global Segment Overflow
# - **[Global Segment Overflow - Exploiting Function Pointers](./GlobalSegmentOverflow.ipynb)**
#
# ### Format String
# - **Will be covered after Network Exploitation**
# - **[Format String Vulnerability & Exploitations](./FormatStrings.ipynb)**
#
# ### SQL Database and Injection Attacks
# - **[SQLite3 with C++](./SQLite3-Database.ipynb)**
# - **[SQL Injection Attacks](./SQLInjection.ipynb)**
#
# ### Capture The Flag
# - **[Capture The Flag](./CTF.ipynb)**
# - **[Return Oriented Programming - ROP](./ROP.ipynb)**
| 00-TableOfContents.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import librosa as lb
import librosa.display
import scipy
import json
import numpy as np
import sklearn
from sklearn.metrics import classification_report
from sklearn.model_selection import train_test_split
import os
import keras
from keras.utils import np_utils
from keras import layers
from keras import models
from keras.models import Sequential
from keras.layers import Dense, Conv2D, MaxPool2D , Flatten, Dropout
from keras.preprocessing.image import ImageDataGenerator
from model_builder import build_example
from plotter import plot_history
import matplotlib.pyplot as plt
from pylab import plot, show, figure, imshow, xlim, ylim, title
# +
# CONSTANTS
DATA_DIR = "openmic-2018/"
CATEGORY_COUNT = 8
LEARNING_RATE = 0.00001
THRESHOLD = 0.5
# +
# LOAD DATA
OPENMIC = np.load(os.path.join(DATA_DIR, 'openmic-mel.npz'), allow_pickle=True)
print('OpenMIC keys: ' + str(list(OPENMIC.keys())))
X, Y_true, Y_mask, sample_key = OPENMIC['X'], OPENMIC['Y_true'], OPENMIC['Y_mask'], OPENMIC['sample_key']
print('X has shape: ' + str(X.shape))
print('Y_true has shape: ' + str(Y_true.shape))
print('Y_mask has shape: ' + str(Y_mask.shape))
print('sample_key has shape: ' + str(sample_key.shape))
# -
y, sr = lb.load(lb.util.example_audio_file(), duration = 10)
S = librosa.feature.melspectrogram(y=y, sr=sr)
plt.figure()
plt.figure(figsize=(5,5))
plt.subplot(1, 1, 1)
S_dB = librosa.power_to_db(S, ref=np.max)
librosa.display.specshow(S_dB, x_axis='time',
y_axis='mel', sr=sr,
fmax=8000)
plt.colorbar(format='%+2.0f dB')
# +
# LOAD LABELS
with open(os.path.join(DATA_DIR, 'class-map.json'), 'r') as f:
INSTRUMENTS = json.load(f)
print('OpenMIC instruments: ' + str(INSTRUMENTS))
# +
# SPLIT DATA (TRAIN - TEST - VAL)
# CHANGE X TO MEL
split_train, split_test, X_train, X_test, Y_true_train, Y_true_test, Y_mask_train, Y_mask_test = train_test_split(sample_key, X, Y_true, Y_mask)
split_val, split_test, X_val, X_test, Y_true_val, Y_true_test, Y_mask_val, Y_mask_test = train_test_split(split_test, X_test, Y_true_test, Y_mask_test, test_size=0.5)
train_set = np.asarray(set(split_train))
test_set = np.asarray(set(split_test))
print('# Train: {}, # Val: {}, # Test: {}'.format(len(split_train), len(split_test), len(split_val)))
# +
# DUPLICATE OF THE MODEL PREPROCESS
print(X_train.shape)
print(X_test.shape)
for instrument in INSTRUMENTS:
# Map the instrument name to its column number
inst_num = INSTRUMENTS[instrument]
print(instrument)
# TRAIN
train_inst = Y_mask_train[:, inst_num]
X_train_inst = X_train[train_inst]
X_train_inst = X_train_inst.astype('float16')
shape = X_train_inst.shape
X_train_inst = X_train_inst.reshape(shape[0],1, shape[1], shape[2])
Y_true_train_inst = Y_true_train[train_inst, inst_num] >= THRESHOLD
i = 0
for val in Y_true_train_inst:
i += val
print('TRAIN: ' + str(i) + ' true of ' + str(len(Y_true_train_inst)) + ' (' + str(round(i / len(Y_true_train_inst ) * 100,2)) + ' %)' )
# TEST
test_inst = Y_mask_test[:, inst_num]
X_test_inst = X_test[test_inst]
X_test_inst = X_test_inst.astype('float16')
shape = X_test_inst.shape
X_test_inst = X_test_inst.reshape(shape[0],1, shape[1], shape[2])
Y_true_test_inst = Y_true_test[test_inst, inst_num] >= THRESHOLD
i = 0
for val in Y_true_test_inst:
i += val
print('TEST: ' + str(i) + ' true of ' + str(len(Y_true_test_inst)) + ' (' + str(round(i / len(Y_true_test_inst ) * 100,2)) + ' %)' )
# VALIDATION
val_inst = Y_mask_val[:, inst_num]
X_val_inst = X_val[val_inst]
X_val_inst = X_val_inst.astype('float16')
shape = X_val_inst.shape
X_val_inst = X_val_inst.reshape(shape[0],1, shape[1], shape[2])
Y_true_val_inst = Y_true_val[val_inst, inst_num] >= THRESHOLD
i = 0
for val in Y_true_val_inst:
i += val
print('VALIDATION: ' + str(i) + ' true of ' + str(len(Y_true_val_inst)) + ' (' + str(round(i / len(Y_true_val_inst ) * 100,2)) + ' %)' )
# -
# <NAME>
len(Y_true_val_inst)
from keras.optimizers import SGD
# This dictionary will include the classifiers for each model
mymodels = dict()
# We'll iterate over all istrument classes, and fit a model for each one
# After training, we'll print a classification report for each instrument
for instrument in INSTRUMENTS:
# Map the instrument name to its column number
inst_num = INSTRUMENTS[instrument]
# Step 1: sub-sample the data
# First, we need to select down to the data for which we have annotations
# This is what the mask arrays are for
# Here, we're using the Y_mask_train array to slice out only the training examples
# for which we have annotations for the given class
# Again, we slice the labels to the annotated examples
# We thresold the label likelihoods at 0.5 to get binary labels
# TRAIN
train_inst = Y_mask_train[:, inst_num]
X_train_inst = X_train[train_inst]
X_train_inst = X_train_inst.astype('float16')
shape = X_train_inst.shape
X_train_inst = X_train_inst.reshape(shape[0],1, shape[1], shape[2])
Y_true_train_inst = Y_true_train[train_inst, inst_num] >= THRESHOLD
# TEST
test_inst = Y_mask_test[:, inst_num]
X_test_inst = X_test[test_inst]
X_test_inst = X_test_inst.astype('float16')
shape = X_test_inst.shape
X_test_inst = X_test_inst.reshape(shape[0],1, shape[1], shape[2])
Y_true_test_inst = Y_true_test[test_inst, inst_num] >= THRESHOLD
# VALIDATION
val_inst = Y_mask_val[:, inst_num]
X_val_inst = X_val[val_inst]
X_val_inst = X_val_inst.astype('float16')
shape = X_val_inst.shape
X_val_inst = X_val_inst.reshape(shape[0],1, shape[1], shape[2])
Y_true_val_inst = Y_true_val[val_inst, inst_num] >= THRESHOLD
# Step 3.
# Initialize a new classifier
model = models.Sequential()
model.add(Conv2D(input_shape=(1,128,430),data_format="channels_first",filters=32,kernel_size=(3,3),padding="same", activation="relu"))
model.add(Conv2D(filters=32,kernel_size=(3,3),padding="same", activation="relu"))
model.add(MaxPool2D(pool_size=(3,3),strides=(2,2)))
model.add(Dropout(0.25))
model.add(Conv2D(filters=128, kernel_size=(3,3), padding="same", activation="relu"))
model.add(Conv2D(filters=128, kernel_size=(3,3), padding="same", activation="relu"))
model.add(MaxPool2D(pool_size=(2,2),strides=(2,2)))
model.add(Dropout(0.25))
model.add(Conv2D(filters=256, kernel_size=(3,3), padding="same", activation="relu"))
model.add(MaxPool2D(pool_size=(2,2),strides=(2,2)))
model.add(layers.Flatten())
model.add(layers.Dense(units=512, activation='relu'))
model.add(layers.Dense(units=256, activation='relu'))
model.add(layers.Dense(units=1, activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer=SGD(lr=0.00001), metrics = ['accuracy'])
# model.summary()
# Step 4.
history = model.fit(X_train_inst,Y_true_train_inst , epochs=50, batch_size=32, validation_data=(X_val_inst,Y_true_val_inst))
plot_history()
loss, acc = model.evaluate(X_test_inst, Y_true_test_inst)
print('Test loss: {}'.format(loss))
print('Test accuracy: {:.2%}'.format(acc))
# Step 5.
# Finally, we'll evaluate the model on both train and test
Y_pred_train = model.predict(X_train_inst)
Y_pred_test = model.predict(X_test_inst)
Y_pred_train_bool = Y_pred_train > THRESHOLD #THRESHOLD (should be lower than 0.5)
Y_pred_test_bool = Y_pred_test > THRESHOLD #THRESHOLD (should be lower than 0.5)
print('-' * 52)
print(instrument)
print('\tTRAIN')
print(classification_report(Y_true_train_inst, Y_pred_train_bool))
print('\tTEST')
print(classification_report(Y_true_test_inst, Y_pred_test_bool))
# Store the classifier in our dictionary
mymodels[instrument] = model
# +
import matplotlib.pyplot as plt
from pylab import plot, show, figure, imshow, xlim, ylim, title
def plot_history():
plt.figure(figsize=(9,4))
plt.subplot(1,2,1)
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train accuracy', 'Validation accuracy'], loc='upper left')
plt.subplot(1,2,2)
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train loss', 'Validation loss'], loc='upper left')
plt.show()
# +
""""
# Step 3: simplify the data by averaging over time
# Instead of having time-varying features, we'll summarize each track by its mean feature vector over time
X_train_inst_sklearn = np.mean(X_train_inst, axis=1)
X_test_inst_sklearn = np.mean(X_test_inst, axis=1)
X_train_inst_sklearn = X_train_inst_sklearn.astype('float32')
X_train_inst_sklearn = lb.util.normalize(X_train_inst_sklearn)
"""
np.savez('models.npz',model=)
| sandbox.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div style="color:#777777;background-color:#ffffff;font-size:12px;text-align:right;">
# prepared by <NAME> (QuSoft@Riga) | November 07, 2018
# </div>
# <table><tr><td><i> I have some macros here. If there is a problem with displaying mathematical formulas, please run me to load these macros.</i></td></td></table>
# $ \newcommand{\bra}[1]{\langle #1|} $
# $ \newcommand{\ket}[1]{|#1\rangle} $
# $ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
# $ \newcommand{\inner}[2]{\langle #1,#2\rangle} $
# $ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
# $ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
# $ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
# $ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
# $ \newcommand{\mypar}[1]{\left( #1 \right)} $
# $ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
# $ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
# $ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
# $ \newcommand{\onehalf}{\frac{1}{2}} $
# $ \newcommand{\donehalf}{\dfrac{1}{2}} $
# $ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
# $ \newcommand{\vzero}{\myvector{1\\0}} $
# $ \newcommand{\vone}{\myvector{0\\1}} $
# $ \newcommand{\vhadamardzero}{\myvector{ \sqrttwo \\ \sqrttwo } } $
# $ \newcommand{\vhadamardone}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
# $ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
# $ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
# $ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
# $ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
# $ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
# $ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
# <h2> Coin Flip: A Probabilistic Bit </h2>
#
# <h3> A fair coin </h3>
#
# A coin has two sides: <i>Head</i> and <i>Tail</i>.
#
# After flipping a coin, we can get a Head or Tail. We can represent these two cases by a single bit:
# <ul>
# <li> 0 represents Head </li>
# <li> 1 represents Tail </li>
# </ul>
# <h3> Flipping a fair coin </h3>
#
# If our coin is fair, then the probability of obtaining a Head or Tail is equal:
#
# $ p= \dfrac{1}{2} = 0.5 $.
#
# Coin-flipping can be defined as an operator:
# <ul>
# <li> $ FairCoin(Head) = \frac{1}{2} Head + \frac{1}{2}Tail $ </li>
# <li> $ FairCoin(Tail) = \frac{1}{2} Head + \frac{1}{2}Tail $ </li>
# </ul>
# $
# FairCoin = \begin{array}{c|cc} & \mathbf{Head} & \mathbf{Tail} \\ \hline \mathbf{Head} & \dfrac{1}{2} & \dfrac{1}{2} \\ \mathbf{Tail} & \dfrac{1}{2} & \dfrac{1}{2} \end{array}
# $
#
# Or, by using 0 and 1:
#
# $
# FairCoin = \begin{array}{c|cc} & \mathbf{0} & \mathbf{1} \\ \hline \mathbf{0} & \dfrac{1}{2} & \dfrac{1}{2} \\ \mathbf{1} & \dfrac{1}{2} & \dfrac{1}{2} \end{array}
# $
# <h3> Task 1: Simulating FairCoin in Python</h3>
#
# Flip a fair coin 100 times. Calcuate the total number of heads and tails, and then compare them.
#
# Do the same experiment 1000 times.
#
# Do the same experiment 10,000 times.
#
# Do the same experiment 100,000 times.
#
# Do your results get close to the ideal case (the numbers of heads and tails are the same)?
# +
# first we import a procedure for picking a random number
from random import randrange
# randrange(m) returns a number randomly from the list {0,1,...,m-1}
# randrange(10) returns a number randomly from the list {0,1,...,9}
# here is an example
r=randrange(5)
print("I picked a random number between 0 and 4, which is ",r)
#
# your solution is here
#
# -
# <a href="..\bronze-solutions\B26_Coin_Flip_Solutions.ipynb#task1">click for our solution</a>
# <h3> Flipping a biased coin </h3>
#
# Our coin may have a bias.
#
# For example, the probability of getting head is greater than the probability of getting tail.
#
# Here is an example:
#
# $
# BiasedCoin = \begin{array}{c|cc} & \mathbf{Head} & \mathbf{Tail} \\ \hline \mathbf{Head} & 0.6 & 0.6 \\ \mathbf{Tail} & 0.4 & 0.4 \end{array}
# $
#
# Or, by using 0 and 1 as the states:
#
# $
# BiasedCoin = \begin{array}{c|cc} & \mathbf{0} & \mathbf{1} \\ \hline \mathbf{0} & 0.6 & 0.6\\ \mathbf{1} & 0.4 & 0.4 \end{array}
# $
# <h3> Task 2: Simulating BiasedCoin in Python</h3>
#
# Flip the following biased coin 100 times. Calcuate the total number of heads and tails, and then compare them.
#
# $
# BiasedCoin = \begin{array}{c|cc} & \mathbf{Head} & \mathbf{Tail} \\ \hline \mathbf{Head} & 0.6 & 0.6 \\ \mathbf{Tail} & 0.4 & 0.4 \end{array}
# $
#
#
# Do the same experiment 1000 times.
#
# Do the same experiment 10,000 times.
#
# Do the same experiment 100,000 times.
#
# Do your results get close to the ideal case $ \mypar{ \dfrac{ \mbox{# of heads} }{ \mbox{# of tails} } = \dfrac{0.6}{0.4} = 1.50000000 } $?
# <a href="..\bronze-solutions\B26_Coin_Flip_Solutions.ipynb#task2">click for our solution</a>
# first we import a procedure for picking a random number
from random import randrange
#
# your solution is here
#
| community/awards/teach_me_quantum_2018/bronze/bronze/.ipynb_checkpoints/B26_Coin_Flip-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ! pip install visualkeras numpy pandas seaborn tensorflow matplotlib
import os
import cv2
import visualkeras
import numpy as np
import pandas as pd
import seaborn as sns
import tensorflow as tf
import matplotlib.pyplot as plt
from sklearn.metrics import confusion_matrix
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPool2D
from tensorflow.keras.layers import BatchNormalization
from tensorflow.keras.optimizers import RMSprop,Adam,SGD,Adadelta
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.model_selection import train_test_split
from warnings import filterwarnings
filterwarnings('ignore')
# ## 1.Dataset checking
data_dir = (r'brain_tumor/Training/') #Path for our train directory
categories = ['glioma_tumor', 'meningioma_tumor', 'no_tumor', 'pituitary_tumor'] #Classes in our dataset
for i in categories:
path = os.path.join(data_dir, i)
for img in os.listdir(path):
img_array = cv2.imread(os.path.join(path,img))
# ## 2. Visualize samples from the dataset
print("Image dataset size :" ,img_array.shape)
new_size = 200
new_array = cv2.resize(img_array,(200,200))
plt.imshow(new_array,cmap = "gray")
# +
plt.figure(figsize=(20, 16))
fileNames = ['glioma_tumor/gg (10).jpg', 'meningioma_tumor/m (108).jpg', 'no_tumor/image (16).jpg', 'pituitary_tumor/p (12).jpg']
for i in range(4):
ax = plt.subplot(2, 2, i + 1)
img = cv2.imread(data_dir + fileNames[i])
img = cv2.resize(img, (new_size, new_size))
plt.imshow(img)
plt.title(categories[i])
# -
# ## 3.Data preparation and features visualization
# +
x_train=[]
y_train=[]
for i in categories:
train_path = os.path.join(data_dir,i)
for j in os.listdir(train_path):
img = cv2.imread(os.path.join(train_path,j),cv2.IMREAD_GRAYSCALE)
img = cv2.resize(img,(new_size,new_size))
x_train.append(img)
y_train.append(i)
# +
x_train=np.array(x_train)
x_train=x_train/255.0
x_train = x_train.reshape(-1,new_size,new_size,1)
y_train = np.array(y_train)
print('DataSet X shape (Data features):',x_train.shape)
print('DataSet Y shape (Gold label):',y_train.shape)
# -
#View the number of data samples in each categorie
sns.countplot(y_train)
#Splitting into training and validation sets at 20%
X_train,X_val,y_train,y_val = train_test_split(x_train,y_train,test_size=0.2,random_state=42)
# +
#Transform our multiclass label to one hot encode type : assign an integer for each category
y_train_new = []
for i in y_train:
y_train_new.append(categories.index(i))
y_train = y_train_new
y_train = tf.keras.utils.to_categorical(y_train)
y_val_new = []
for i in y_val:
y_val_new.append(categories.index(i))
y_val = y_val_new
y_val = tf.keras.utils.to_categorical(y_val)
# -
print('New shapes after spliting and encoding :')
print("TrainSet shape :",X_train.shape, y_train.shape)
print("ValidationSet shape :",X_val.shape, y_val.shape)
# ## 4.Data augmentation and preprocessing
#using the keras pre-build function to generate and process the data
datagen = ImageDataGenerator(
featurewise_center=False,
samplewise_center=False,
featurewise_std_normalization=False,
samplewise_std_normalization=False,
zca_whitening=False,
rotation_range=0,
zoom_range = 0,
width_shift_range=0,
height_shift_range=0,
horizontal_flip=True,
vertical_flip=False)
# ## 5.Model creation and compilation
# +
#Model creation (layers choice will be explained in the report)
model = Sequential()
model.add(Conv2D(filters=64, kernel_size=3, padding= 'Same', activation='relu', input_shape=(X_train.shape[1],X_train.shape[2],1)))
model.add(MaxPool2D(pool_size=(2,2)))
model.add(BatchNormalization())
model.add(Conv2D(filters=128, kernel_size=3, padding= 'Same', activation='relu'))
model.add(MaxPool2D(pool_size=(2,2)))
model.add(BatchNormalization())
model.add(Conv2D(filters=128, kernel_size=3, padding= 'Same', activation='relu'))
model.add(MaxPool2D(pool_size=(2,2)))
model.add(BatchNormalization())
model.add(Conv2D(filters=256, kernel_size=3, padding= 'Same', activation='relu'))
model.add(MaxPool2D(pool_size=(2,2)))
model.add(BatchNormalization())
model.add(Conv2D(filters=256, kernel_size=3, padding= 'Same', activation='relu'))
model.add(MaxPool2D(pool_size=(2,2)))
model.add(BatchNormalization())
model.add(Conv2D(filters=512, kernel_size=3, padding= 'Same', activation='relu'))
model.add(MaxPool2D(pool_size=(2,2)))
model.add(BatchNormalization())
model.add(Flatten())
model.add(Dense(1024, activation = "relu"))
model.add(Dropout(0.2))
model.add(Dense(512, activation = "relu"))
model.add(Dropout(0.2))
model.add(Dense(4, activation = "softmax"))
# -
#Model summary
model.summary()
#Model architecture
print('Model architecture :')
visualkeras.layered_view(model)
#Model compilation
optimizer = SGD(learning_rate=0.01)
model.compile(optimizer = optimizer, loss = "categorical_crossentropy", metrics=["accuracy"])
# ## 6.Training and Evaluation
#Setting the number of epochs
epochs = 50
batch_size = 64
history = model.fit_generator(datagen.flow(X_train,y_train, batch_size=batch_size),
epochs = epochs,
validation_data = (X_val,y_val))
# +
#accuracy and loss values during Training
plt.subplots( figsize=(20,17))
plt.subplot(1, 2, 1)
plt.plot(history.history["loss"],c = "purple")
plt.plot(history.history["val_loss"],c = "orange")
plt.legend(["train", "test"])
plt.title('Loss non-regularized model')
plt.subplot(1, 2, 2)
plt.plot(history.history["accuracy"],c = "purple")
plt.plot(history.history["val_accuracy"],c = "orange")
plt.legend(["train", "test"])
plt.title('Accuracy non-regularized model')
plt.show()
# -
#Model evaluation
y_pred = model.predict(X_val)
class_pred = [np.argmax(i) for i in y_pred]
val_labels =[np.argmax(i) for i in y_val]
#report = classification_report(val_labels, class_pred)
clf_report = classification_report(val_labels, class_pred,output_dict=True,labels=[0,1,2,3],target_names=categories)
sns.heatmap(pd.DataFrame(clf_report).iloc[:-1, :].T, annot=True)
plt.savefig('Model_CNN_Classification_report.png')
f,ax = plt.subplots(figsize=(10, 10))
confusion_mtx = confusion_matrix(val_labels, class_pred)
sns.set(font_scale=1.4)
sns.heatmap(confusion_mtx, annot=True, linewidths=0.01,cmap="Greens",linecolor="gray",ax=ax)
plt.xlabel("Predicted Label")
plt.ylabel("True Label")
plt.title("Confusion Matrix Validation set")
plt.show()
test_image = "tests/normal1.jpg"
#Read and prepare the image for the prediction (by resizing and applying the required parameters for testing)
img = cv2.imread(test_image,cv2.IMREAD_GRAYSCALE)
img = cv2.resize(img,(new_size,new_size))
img =np.array(img)
img =img/255.0
img_test =img.reshape(-1,new_size,new_size,1)
plt.imshow(img)
plt.show()
# +
#Using the input image predict the result/category
predicted_label = list(model.predict(img_test))
predicted_label = [np.argmax(i) for i in predicted_label]
print("Predicted image reesult:",categories[predicted_label[0]])
# -
#Save the model
model.save('Brain_cancer.h5')
| Deep_learning_part.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''dpln_env'': conda)'
# name: python3
# ---
# <br><center><span style='font-family: "DejaVu Sans"; font-size:48px; color:#FF7133'>Growing Pains Case</span></center>
# <center><span style='font-family: "DejaVu Sans"; font-size:28px'><i>Processing Solution Results</i></span></center>
# ---
# Standard libraries
from __future__ import print_function
import os
import pickle
from copy import deepcopy
from itertools import product, permutations, combinations
from collections import defaultdict
# 3rd party libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
from ipyleaflet import Map, Marker, MarkerCluster, AwesomeIcon, AntPath
# ---
# Import the file with outputs from the solver:
# +
soln_file = 'pickle/GPCaseResults_24Aug20_18-42.pickle'
data_file = 'pickle/gp_data.pickle'
with open(soln_file, 'rb') as f:
results = pickle.load(f)
with open(data_file, 'rb') as f:
gp_data = pickle.load(f)
# -
time_matrices = gp_data[0] # daily travel time matrices
demands = gp_data[1] # demand per location per day
travel_times = gp_data[2] # matrix of travel times between ALL locations (dataframe)
daily_orders = gp_data[3] # dataframe of daily orders for reference
daily_total_dels = gp_data[4] # summary of total orders by day
daily_total_vols = gp_data[5] # summary of total volume required by day
# <br> Import the store locations and process as was done before in the [EDA notebook](GrowingPains_EDA_DataPrep.ipynb).
# Store locations
store_locs = pd.read_csv('data/locations.csv', nrows=123) # last few rows in .csv are junk
# Change zipcodes to strings and pad with leading zeros
store_locs['ZIP'] = store_locs['ZIP'].apply(lambda x: str(int(x)).zfill(5))
store_locs.head(10)
# The statement below from the EDA and pre-processing notebook converts the zip ID's to coordinates that can be provided as input to an interactive map.
store_loc_coords = store_locs.loc[:, ['ZIPID', 'Y', 'X']].set_index('ZIPID').T.to_dict('list')
# store_loc_coords
# ---
# <br> <h3><u>Result Summaries</u></h3>
# The function below summarizes the overall totals making it easy to see at a glance the requirements for each day and compare different models.
# +
days = ['Mon', 'Tue', 'Wed', 'Thu', 'Fri']
totals = ['totalNumberOfVehicles',
'totalLoadAllVehicles',
'totalTimeAllVehicles',
'totalDistanceAllVehicles']
def result_summary(results=results, days=days, totals=totals):
results = deepcopy(results)
summary_dict = dict()
for day in days:
summary_dict[day] = {total: results.get(day).get(total) for total in totals}
return pd.DataFrame(summary_dict)
# -
result_summary()
# Loads are in cubic <i>ft<sup>3</sup></i> volume, total time is in <i>minutes</i>, and total distance is in <i>miles</i>.
# ---
# <br> <h3><u>Schedules</u></h3>
# The ultimate goal of this problem is to obtain schedules for each day. The class below processes the data from `results` and returns a summary for all the day's routes and a detailed schedule for every route.
# The first two functions are the functions from the [solution notebook](GrowingPains_ProblemSolution.ipynb) to convert the times from minutes to clock time:
# +
def clock_to_minutes(clock_time):
'''
Converts clock time to time of day in minutes
'''
t = clock_time.split(':')
return (int(t[0])*60 + int(t[1]))
def minutes_to_clock(time_in_minutes):
'''
Converts minutes of the day to clock time
'''
return ('{:02d}:{:02d}'.format(*divmod(time_in_minutes, 60)))
# -
# <br> This class returns the summaries for all routes in a day and a schedule for each route.
class route_scheduler(object):
'''
Returns route schedules from the solution results
'''
def __init__(self, day, results=results, demands=demands):
'''
day: day to schedule
results: The dict with solution results
demands: Dict with demands and orignal ZIPIDs
'''
self.day = day
# Important! Use deepcopy() else things can get messy
self.results = deepcopy(results.get(self.day))
self.demands = deepcopy(demands)
self.vehicles = [veh for veh in self.results.keys() \
if veh[-1].isnumeric()]
# map result ID's back to original ID's
zipIDs = list(self.demands.get(self.day).keys())[:]
# make sure depot (DC - ZIPID 20) is included and comes first
if 20 not in zipIDs:
zipIDs.insert(0, 20)
if zipIDs[0] != 20:
zipIDs.remove(20)
zipIDs.insert(0, 20)
self.stop_labels = dict(enumerate(zipIDs))
def get_vehicles(self):
'''Returns vehicle labels'''
return self.vehicles
def route_summaries(self):
'''
Get route summaries for given day
'''
summary = defaultdict(dict)
for vh in self.vehicles:
t = self.results.get(vh).get('totalTime')
d = self.results.get(vh).get('totalDistance')
L = self.results.get(vh).get('totalLoad')
summary[vh] = {'Total Duty Time': f'{t//60}hrs {t%60}min',
'Total Travel Distance': f'{d} miles',
'Total Driving Time': f'{d//40}hrs {int(((d%40)/40)*60)}min',
'Load Carried': f'{L} cuft.'}
return pd.DataFrame(summary)
def vehicle_schedule(self, vehicle):
'''Get complete schedule for a route'''
vh = self.results.get(vehicle)
stop_idx = vh.get('stops')
stops = []
for idx in stop_idx:
z = self.stop_labels.get(idx) # get ZIPID
cty = store_locs[store_locs.ZIPID == z].get('CITY').to_numpy()[0]
st = store_locs[store_locs.ZIPID == z].get('STATE').to_numpy()[0]
stop = "Distribution Center" if idx == 0 else " ".join([cty, st])
stops.append(stop)
# arrival times
t_arr = []
for i,t in enumerate(vh.get('arrive')):
if i == 0:
t_arr.append(None) # depot
else:
t_arr.append(minutes_to_clock(t))
# unloading time
unload = [(j-i) for i,j in\
zip(vh.get('arrive')[:-1], vh.get('depart'))]
unload.append(0) # add zero for end point (depot)
# deliveries
deliveries = vh.get('load')[:]
deliveries.append(0) # add zero for end point (depot)
# departures
t_dep = []
for t in vh.get('depart'):
t_dep.append(minutes_to_clock(t))
t_dep.append(None) # depot
# travel distance
d_tot = vh.get('distance')
dist = []
for i in range(len(d_tot)):
if i == 0: # route start
dist.append(d_tot[i])
else:
dist.append(d_tot[i] - d_tot[i-1])
schedule = pd.DataFrame({'Stop': stops,
'Arrive': t_arr,
'Unloading_time(min)': unload,
'Departure': t_dep,
'Delivery(cuft.)': deliveries,
'Travel_distance(mi)': dist})
return schedule
# <br> <b>Using the Route Scheduler</b>
# <br> An example showing route summaries and a few route schedules for <u>Thursday</u>:
# <b>Summaries:</b>
Thu_schedule = route_scheduler('Thu')
Thu_schedule.route_summaries()
# <b>Route schedules:</b>
# vehicle_2
Thu_schedule.vehicle_schedule('vehicle_2')
# vehicle_3
Thu_schedule.vehicle_schedule('vehicle_3')
# vehicle_4
Thu_schedule.vehicle_schedule('vehicle_4')
# ---
# <br> <h3><u>Visualizing Routes</u></h3>
# Visualizing the routes can be very helpful. This next class is used to display a map showing the locations visited by a given vehicle for a given day. The path on the map shows the connections and order of the stops.
if not os.path.exists('RouteMaps'):
os.makedirs('RouteMaps')
class route_map(object):
'''
Create a map object to show routes
'''
def __init__(self, day,
locations=store_loc_coords,
demands=demands,
results=results):
'''
day: Day to get routes
locations: dict with delivery location coordinates
results: dict object with solver results
'''
self.day = deepcopy(day)
self.locations = deepcopy(locations)
self.demands = deepcopy(demands)
self.results = deepcopy(results.get(self.day))
self.vehicles = [veh for veh in self.results.keys() \
if veh[-1].isnumeric()]
# map result ID's back to original ID's
zipIDs = list(self.demands.get(self.day).keys())[:]
# make sure depot (DC - ZIPID 20) is included and comes first
if 20 not in zipIDs:
zipIDs.insert(0, 20)
if zipIDs[0] != 20:
zipIDs.remove(20)
zipIDs.insert(0, 20)
self.stop_labels = dict(enumerate(zipIDs))
def __repr__(self):
return f'Show optimized routes for {day!r} on interactive map'
def getAllStopCoords(self):
'''
Maps node IDs for all stops of given day
to map coordinates of store
'''
self.stop_coords = {idx:tuple(self.locations.get(zipID)) \
for idx, zipID in self.stop_labels.items()}
def get_route_coords(self, vehicle):
'''Get the route XY coords'''
route = self.results.get(vehicle).get('stops')
return dict(enumerate([self.stop_coords.get(stop) for stop in route]))
def plot_route(self, vehicle, save_to_file=None):
'''
Plot route on an interactive map
---
save: path and filename to save html map
'''
route = self.get_route_coords(vehicle)
dc = route.get(0)
dc_icon = AwesomeIcon(name='home', marker_color='red')
# create map; centered roughly at mean of locations
_ = list(zip(*list(route.values())))
x, y = np.mean(_[0]), np.mean(_[1])
route_map = Map(center=(x, y), zoom=8)
# mark DC location
dc_marker = Marker(location=dc,
title='Depot-DC',
icon=dc_icon,
draggable=False)
route_map.add_layer(dc_marker)
# mark stops (1st and last are DC)
stops = list(route.values())[1:-1]
for i, loc in enumerate(stops):
marker = Marker(location=loc, title=str(i), draggable=False)
route_map.add_layer(marker)
# show route path
p = list(route.values())
p.append(dc)
path = AntPath(locations=p, weight=4)
route_map.add_layer(path)
if save_to_file is not None:
route_map.save(save_to_file, title=str(self.day) + '_' + str(vehicle))
return route_map
# <br> <b>Some route examples:</b>
Wed_routes = route_scheduler('Wed')
Wed_routes.route_summaries()
Wed_routes.vehicle_schedule('vehicle_1')
# +
Wed_map = route_map('Wed')
Wed_map.getAllStopCoords()
Wed_map.plot_route('vehicle_1', 'RouteMaps/Wed_veh1.html')
# -
| GrowingPains_ProcessingResults.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## ArcGIS Notebook to be demonstrate the usage of the built-in [Spark](https://spark.apache.org/) engine.
#
#
# This notebook will demonstrate the spatial binning of AIS data around the port of Miami using Apache Spark.
#
# The AIS broadcast data is in a FileGeodatabase that can be download from [here](https://marinecadastre.gov/ais). Note only data from 2010-2014 is in FileGeodatabase format.
# ### Import the required modules.
import os
import arcpy
from spark_esri import spark_start, spark_stop
# ### Start a Spark instance.
#
# Note the `config` argument to [configure the Spark instance](https://spark.apache.org/docs/latest/configuration.html).
config = {
"spark.driver.memory":"16G",
"spark.executor.memory":"16G"
}
spark = spark_start(config=config)
spark.version
# ### Read the selected Broadcast feature shapes in WebMercator SR.
#
# It is assumed that you added to the map the `Broadcast` point feature class from the download `miami.gdb`.
#
# Note that the `SearchCursor` is subject to the user selected features, and to an active query definition in the layer properties. For Example, set the query definition to `Stauts = 0: Under way using engine` to get the location of all moving ships, in such that we get a "heat map" of the port movement.
sp_ref = arcpy.SpatialReference(3857)
data = arcpy.da.SearchCursor("Broadcast", ["SHAPE@X","SHAPE@Y"], spatial_reference=sp_ref)
# ### Create a Spark data frame of the read data, and create a view named 'v0'.
spark\
.createDataFrame(data, "x double,y double")\
.createOrReplaceTempView("v0")
# ### Aggregate the data at 200x200 meters bins.
#
# The aggregation is performed by Spark as a SQL statement in a parallel share-nothing way and the resulting bins are collected back in the `rows` array variable.
#
# This is a nested SQL expression, where the inner expression is mapping the input `x` and `y` into `q` and `r` cell locations given a user defined bin size, and the outter expression is aggreating as a sum the `q` and `r` pairs. Finally, `q` and `r` are mapped back to `x` and `y` to enble the placement on a map.
# +
cell0 = 200.0 # meters
cell1 = cell0 * 0.5
rows = spark\
.sql(f"""
with t as (select cast(x/{cell0} as long) q,cast(y/{cell0} as long) r from v0)
select q*{cell0}+{cell1} x,r*{cell0}+{cell1} y,least(count(1),400) as pop
from t
group by q,r
""")\
.collect()
# -
# ### Report the number of bins
len(rows)
# ### Create an in-memory point feature class of the collected bins.
#
# The variable `rows` is an array of form `[[x0,y0,pop0],[x1,y1,pop1],...,[xN,yN,popN]]`.
# +
ws = "memory"
nm = "Bins"
fc = os.path.join(ws,nm)
arcpy.management.Delete(fc)
sp_ref = arcpy.SpatialReference(3857)
arcpy.management.CreateFeatureclass(ws,nm,"POINT",spatial_reference=sp_ref)
arcpy.management.AddField(fc, "POP", "LONG")
with arcpy.da.InsertCursor(fc, ["SHAPE@X","SHAPE@Y", "POP"]) as cursor:
for row in rows:
cursor.insertRow(row)
# -
# ### Apply a graduated colors symbology to highlight the bins.
# +
# _ = arcpy.ApplySymbologyFromLayer_management(fc, f"{nm}.lyrx")
# -
# ### Stop the spark instance.
spark_stop()
| notebooks/spark_esri.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hyperparameters and Model Validation
# 1. Choose a class of model
# 2. Choose model hyperparameters
# 3. Fit the model to the training data
# 4. Use the model to predict labels for new data
from sklearn.datasets import load_iris
iris = load_iris()
X = iris.data
y = iris.target
from sklearn.neighbors import KNeighborsClassifier
model = KNeighborsClassifier(n_neighbors=1)
model.fit(X, y)
y_model = model.predict(X)
from sklearn.metrics import accuracy_score
accuracy_score(y, y_model)
# ### Model validation:
# +
from sklearn.model_selection import train_test_split
# split the data with 50% in each set
X1, X2, y1, y2 = train_test_split(X, y, random_state=0,
train_size=0.5)
# fit the model on one set of data
model.fit(X1, y1)
# evaluate the model on the second set of data
y2_model = model.predict(X2)
accuracy_score(y2, y2_model)
# -
# ### Model validation via cross-validation
y2_model = model.fit(X1, y1).predict(X2)
y1_model = model.fit(X2, y2).predict(X1)
accuracy_score(y1, y1_model), accuracy_score(y2, y2_model)
from sklearn.model_selection import cross_val_score
cross_val_score(model, X, y, cv=5)
# # Grid search
from sklearn import svm, datasets
from sklearn.model_selection import GridSearchCV
help(svm.SVC)
iris = datasets.load_iris()
parameters = {'kernel':('linear', 'rbf'), 'C':[1, 10]}
svc = svm.SVC(gamma="scale")
clf = GridSearchCV(svc, parameters, cv=5)
clf.fit(iris.data, iris.target)
sorted(clf.cv_results_.keys())
clf.cv_results_['param_C'],clf.cv_results_['param_kernel']
clf.cv_results_['mean_test_score']
| SII/ML/3-Hyperparameters-and-Model-Validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Procesamiento Digital de Señales (DSP)
# Una de las ramas mas importantes de la electrónica y sistemas de control es el procesamiento Digital de señales. Son muchísimas las posibilidades de aplicacón al procesar señales de distintas fuentes: Sonido, transductores, señales eléctricas o electromagnéticas puras entre muchas otras.
#
# 
#
# Los manatíes no se pueden ver en ríos de aguas turbias, por lo que se identifican por sus sonidos. Al igual que las personas, los manatíes emiten sus sonidos con ligeras variaciones, aunque cada uno tiene una "voz" única. Por esto, contar cuántos manatíes hay en un río se puede realizar usando técnicas de clustering y DSP, procesando los sonidos que se captan, pensando en que cada cluster corresponde a un manatí en particular.
#
# En este caso se cuenta con una base de datos de muestras de sonidos de manatíes captados en un río.
#
# +
# Imports
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# %matplotlib inline
# -
# ## Datos a procesar
# Se lee el archivo csv con el dataset
manaties = pd.read_csv('../datasets/dataset_manaties.csv',engine='python')
manaties.info()
# Observar datos de forma rápida
manaties.head()
# ## Prepocesar los datos
# Ahora se tiene los datos en formato leide de csv, pero resulta mas facil trabajar con ellos ya normalizados, y en formato de dataset. Como se puede ver, se tienen 4 variables para cada muestra.
#
# Se obtienen tambien las metricas estadisticas para cada columna y las mas significativas.
manaties.describe()
# Normalizar los datos
manaties_normalizado = (manaties - manaties.min())/(manaties.max() - manaties.min())
manaties_normalizado.describe()
# Ahora que se tienen los datos normalizados entre 0 y 1, se pueden obtener sus metricas estadisticas en dos o tres variables PCA
# +
from sklearn.decomposition import PCA
pca = PCA(n_components=3) # tres componentes principales
pca_manaties = pca.fit_transform(manaties_normalizado)
DataFrame_manaties = pd.DataFrame(data= pca_manaties, columns=['Componente_1', 'Componente_2','Componente_3'])
# -
# ## Actividad propuesta:
# ### 1. Ahora que se tienen los datos procesados ordenados y normalizados, elija un método de agrupamiento y realice el análisis correspondiente, puede utilizar PCA (2 o 3 elementos) o usar los datos normalizados directamente.
# ### 2. Obtenga los diferentes clusters y cree un archivo csv con los resultados en la carpeta Results.
# ### 3. Realice gráficas del procedimiento y ajuste lo mejor posible el método. Justifique su elección.
# ### 4. Indique sus principales Conclusiones.
| IA_Clustering_Apps_Simulation/DSP estudio de manaties.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Time Series Analysis
# ---
# ## ARMA($p$,$q$) models
#
# A stochastic process process $\{x_t\}$ is an ARMA($p$,$q$) process (autoregressive, moving average process of order $p$ and $q$, respectively) if we have
#
# $$
# x_t = \mu + \phi_1 x_{t-1} + \phi_2 x_{t-2} + ... \phi_p x_{t-p} + \epsilon_{t} + \theta_1 \epsilon_{t-1} + \theta_2 \epsilon_{t-2} + ... + \theta_q \epsilon_{t-q}
# $$
#
# where $\epsilon_{t}$ is white noise, i.e. $E[\epsilon_{t}]=0$ and $Var[\epsilon_{t}]=\sigma^2$, and without loss of generality, we set $\mu$=0.
#
# If the coefficients $\phi_i \equiv 0$, then the ARMA($p$,$q$) process collapses to an MA($q$) process.
#
# Similarly, if $\theta_i\equiv 0$ then the ARMA($p$,$q$) process collapses to an AR($p$) process.
#
# A slightly more general formulation also used is given by the expression
#
# $$
# \sum_{i=0}^{p} \phi_i X_{t-i} = \sum_{i=0}^{q} \theta_i \varepsilon_{t-i}
# $$
#
# Note that this formulation introduces $\phi_0,\theta_0$ terms which were implicitly defined to be 1 above. Further, the sign of the AR coefficients is flipped from the first formulation. This is frequently how statistical computing packages treat ARMA($p$,$q$) processes.
#
#
# Let's simulate some ARMA($p$,$q$) processes. We will later examine both their autocorrelation functions (ACFs) and partial autocorrelation functions (PACFs).
# ### Data Simulation
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
def simulate_arma(ar=None, ma=None, nsample=100,
burnin=0, paths=1, seed=1):
"""Simulate ARMA data.
Assumption: White noise shocks are Gaussian with mean = 0, std = 1.
Args:
ar: list or None(Default) coefs for AR process, if None, then p=0.
ma: list or None(Default), coefs for MA process, if None, then q=0.
nsample: int, sample size, default=100.
burnin: int, number of sample for "warming start", default=1.
paths: int, number of time series, default=1.
seed: int, random seed for reproduction, default=1.
Returns:
y: np.array, size=[nsample, paths], generated ARMA process.
"""
if not ar:
ar = []
if not ma:
ma = []
# Seed the random number generator
np.random.seed(seed)
# numpy arrays are reversed for easier indexing:
ar, ma = np.array(ar[::-1]), np.array(ma[::-1])
# Orders (does not include zero lag)
p, q = len(ar), len(ma)
max_order = max(p, q)
# Total number of sample size
Nsim = nsample + burnin
# "Standard" Guassian shocks: Normal(0,1)
eps = np.random.randn(paths, Nsim)
# Initialize t < 0 with zeros
eps = np.concatenate((np.zeros((paths, max_order)), eps), axis=1)
y = np.zeros((paths, Nsim + max_order))
# Loop to construct the ARMA processes recursively.
for tt in range(max_order, Nsim + max_order):
y[:, tt] = np.sum(y[:, tt - p:tt] * ar, axis=1) \
+ np.sum(eps[:, tt - q:tt] * ma, axis=1) + eps[:, tt]
# Drop initial zeros and burnin and transpose for plotting.
y = y[:, max_order + burnin:].T
return y
# +
### Seed the random number generator
# (by default, the seed is set to 1 if no parameter is passed)
seed = np.random.seed(1)
### 1. White Noise
y_wn = simulate_arma(nsample=250,seed=seed)
### 2. AR(1) with persistence = 0.75
phi1_model1 = 0.75
y_ar1_model1 = simulate_arma(ar=[phi1_model1], nsample=250)
fig1, axes1 = plt.subplots(nrows=2, ncols=1, figsize=(12,10))
axes1[0].plot(y_wn)
axes1[0].set_xlabel('Time', fontsize=16)
axes1[0].set_title('White Noise', fontsize=18)
axes1[0].axhline(y=0, linewidth=0.4)
axes1[1].plot(y_ar1_model1)
axes1[1].set_xlabel('Time', fontsize=16)
axes1[1].set_title('AR($1$)' + ' with $\phi$ = ' + str(phi1_model1) , fontsize=18)
axes1[1].axhline(y=0, linewidth=0.4)
fig1.tight_layout()
# -
# `statsmodels` wrapper to use
# +
import statsmodels.api as sm
arparams = np.array([.75])
maparams = np.array([.65, .35])
ar = np.r_[arparams]
ma = np.r_[1]
y = sm.tsa.arma_generate_sample(ar, ma, 250)
# -
# ### ACFs/PACFs
#
# In this part, we focus on computing and understanding Autocorrelation Functions (ACFs), and Partial Autocorrelation Functions (PACFs).
#
# The ACFs can be defined as follows:
#
# \begin{align*}
# \text{Autocovariance function}: \gamma_j &\equiv \text{cov}(x_t,x_{t-j}) \\
# \text{Autocorrelation function}: \rho_j &\equiv \frac{\gamma_j}{\gamma_0}
# \end{align*}
#
# Both can be estimated using the Method of Moment, i.e. using their sample analogues:
#
# \begin{align*}
# \hat{\gamma_j} &\equiv \widehat{\text{cov}}(x_t,x_{t-j}) \\
# \hat{\rho_j} &\equiv \widehat{\text{corr}}(x_t,x_{t-j})
# \end{align*}
#
# This is what we will use today. However, the ACFs can also be obtained from a regression of $x_t$ on $x_{t-j}$ (we often add a constant to the regression):
#
# $$
# x_t = \alpha_j + \beta_j x_{t-j} + \epsilon_t
# \,\,\text{ where }\,\,
# \hat{\beta_j} = \frac{\text{cov}(x_t,x_{t-j})}{\text{var}(x_t)} = \hat{\rho_j}
# $$
#
# In a similar fashion, we can define the Partial Autocrrelation Functions (PACFs) as the correlation between $x_t$ and $x_{t-j}$ *conditional on controling for other lags* as follows (cf. Ruppert & Matteson 12.13 for details):
#
# $$
# x_t = \beta_{0,j} + \beta_{1,j} x_{t-1} + \ldots + \beta_{j,j} x_{t-j} + \epsilon_t
# \,\,\text{ where }\,\,
# \hat{\beta_{j,j}} = \widehat{PACF_j}
# $$
#
# Below we see how to implement the estimation of ACFs and PACFs via OLS, as well as the function in statsmodels.
# +
# 1. White Noise
y_wn_long = simulate_arma(nsample=10000).squeeze()
# 2. AR(1) with persistence = 0.75
phi1_model1 = 0.75
y_ar1_model1_long = simulate_arma(ar=[phi1_model1], nsample=10000).squeeze()
# +
# statsmodels function for ACF and PACF
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
fig, axes = plt.subplots(2, 2, figsize=(15, 10))
nlags = 15
plot_acf(y_wn_long, ax=axes[0,0], lags=nlags,
title="${\it statsmodels}$ ACF: White Noise")
plot_pacf(y_wn_long, ax=axes[1,0], lags=nlags,
title="${\it statsmodels}$ PACF: White Noise")
plot_acf(y_ar1_model1_long, ax=axes[0,1], lags=nlags,
title="${\it statsmodels}$ ACF: AR(1) with $\\phi$=" + str(phi1_model1))
plot_pacf(y_ar1_model1_long, ax=axes[1,1], lags=nlags,
title="${\it statsmodels}$ PACF: AR(1) with $\\phi$=" + str(phi1_model1))
fig.tight_layout()
# +
def calc_ols(y, x, addcon=True):
""" Calculate OLS coefficients from scratch. """
Nobs = y.shape[0]
if addcon:
X = np.c_[np.ones((Nobs,1)), x] # append the [Nobs x 1] columns of ones.
else:
X = x
XX = np.dot(X.T, X) # Construct sample average of E[X'X]
Xy = np.dot(X.T, y) # Construct sample average of E[X'Y]
beta_hat = np.linalg.solve(XX, Xy) # algebraic solution for OLS. beta_hat = (E[X'X]^-1)*(E[X'Y])
resids = y - np.dot(X, beta_hat) # residual eps_hat = y - beta_hat*X
return beta_hat, resids, X
def lag_mat(y, nlags, fill_vals=np.nan):
""" Create a matrix of lags of a given vector. """
y_lags = np.empty((y.shape[0], nlags + 1))
y_lags.fill(fill_vals)
# Include 0 lag
for lag in range(nlags + 1):
y_lags[lag:, lag] = np.roll(y, shift=lag)[lag:]
return y_lags
# ACF and PACF for a given lag
def calc_acf_lag_ols(y_lags, lag):
""" ACF for a given lag (OLS). """
if lag == 0:
return 1.
lhs = y_lags[lag:, 0]
rhs = y_lags[lag:, lag:lag + 1]
beta_hat, _, _ = calc_ols(y=lhs, x=rhs, addcon=True)
return beta_hat[-1]
def calc_pacf_lag_ols(y_lags, lag):
""" PACF for a given lag (OLS). """
if lag == 0:
return 1.
lhs = y_lags[lag:, 0]
# need y_lags[lag:, 1:lag+1] instead of y_lags[lag:,lag:lag+1] (unlike "calc_acf_lag_ols")
rhs = y_lags[lag:, 1:lag + 1]
beta_hat, _, _ = calc_ols(y=lhs, x=rhs, addcon=True)
return beta_hat[-1]
# ACF and PACF for all lags
def calc_acf_ols(y, nlags):
"""ACF for multiple lags."""
y_lags = lag_mat(y, nlags)
acf_list = [calc_acf_lag_ols(y_lags, lag) for lag in range(nlags + 1)]
return np.array(acf_list)
def calc_pacf_ols(y, nlags):
"""PACF for multiple lags."""
y_lags = lag_mat(y, nlags)
pacf_list = [calc_pacf_lag_ols(y_lags, lag) for lag in range(nlags + 1)]
return np.array(pacf_list)
# Plotting functions
def my_plot_acf(y, nlags=10, ax=None, title_string='',
title_fontsize=None, xlabel_string='Time'):
"""Plotting ACF with approx SEs."""
T = y.shape[0]
# approx SEs: scaling used in aysmptotic
# For j = 1: 1/sqrt(T) (i.e. H0: WN)
# For j > 1: sqrt((1 + 2*sum_{i=1}^{j-1} rho_i^2)/T)
# For details: cf. slide 23 of Lecture 1 and part on bartlett_confin on
# https://www.statsmodels.org/dev/generated/statsmodels.graphics.tsaplots.plot_acf.html
# Note: still slightly off compared to statsmodel, so to double-check
# se_approx = 1/np.sqrt(T)
se_approx = np.sqrt((np.insert(1 + 2 * np.cumsum(calc_acf_ols(y, nlags) ** 2), 0, 1, axis=0)[:-1]) / T)
# set up figure
if ax is None:
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(5, 5))
# ACF
ax.plot(calc_acf_ols(y, nlags), c='xkcd:true blue',
marker='o', markerfacecolor='xkcd:azure')
ax.fill_between(x=range(0, nlags + 1), y1=-1.96 * se_approx, y2=1.96 * se_approx, facecolor='blue', alpha=0.1)
ax.set_xlabel(xlabel_string)
if title_fontsize != None:
ax.set_title('${\it ACF}$: ' + title_string, fontsize=title_fontsize)
else:
ax.set_title('${\it ACF}$: ' + title_string)
def my_plot_pacf(y, nlags=10, ax=None, title_string='',
title_fontsize=None, xlabel_string='Time'):
"""Plotting PACF with approx SEs."""
T = y.shape[0]
# approx SEs: scaling used in aysmptotic
# To double-check
se_approx = 1 / np.sqrt(T)
# set up figure
if ax is None:
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(5, 5))
# PACF
ax.plot(calc_pacf_ols(y, nlags), c='xkcd:true blue',
marker='o', markerfacecolor='xkcd:azure')
ax.fill_between(x=range(0, nlags + 1), y1=-1.96 * se_approx, y2=1.96 * se_approx, facecolor='blue', alpha=0.1)
ax.set_xlabel(xlabel_string)
if title_fontsize != None:
ax.set_title('${\it PACF}$: ' + title_string, fontsize=title_fontsize)
else:
ax.set_title('${\it PACF}$: ' + title_string)
# +
nlags = 15
fig, axes = plt.subplots(nrows=2, ncols=2, figsize=(12,8))
my_plot_acf(y_wn_long, nlags , ax=axes[0,0], title_string="White Noise")
my_plot_pacf(y_wn_long, nlags , ax=axes[0,1], title_string="White Noise")
my_plot_acf(y_ar1_model1_long, nlags, ax=axes[1,0], title_string="AR(1) with $\\phi$=" + str(phi1_model1))
my_plot_pacf(y_ar1_model1_long, nlags , ax=axes[1,1], title_string="AR(1) with $\\phi$=" + str(phi1_model1))
fig.tight_layout()
# -
# #### Summary for ACF/PACF
#
# The Table below (Table 3.1 p. 101 in Shumway and Stoffer, 2016) summarizes what we can conclude from ACFs/PACFs about the order of an ARMA($p$,$q$) process.
#
# | Function | AR($p$) | MA($q$) | ARMA($p$,$q$) |
# |:--- | :---: |:---: |:---: |
# | ACF | Tails off | Cuts off for j > q | Tails off |
# | PACF | Cuts off for j > p | Tails off | Tails off |
#
# In practice, we have seen that it is sometimes tricky to distinguish close processes, especially in small samples. For those cases, using other and more formal criteria is then required. Some other like AIC/BIC, significance of lags, *etc.*.
# ### OLS Estimation, Diagnostics
#
# **<NAME>**
#
# Recall that the OLS estimate for AR(1) is biased downwards:
# $$
# \mathbb{E}_t[\hat{\phi}] \approx \phi - \frac{1+3\phi}{T}
# $$
#
# The bias increases with $\phi$ in absolute value (i.e. the estimate is biased downwards even more), and decreases with the sample size $T$. It is therefore a good idea to correct the estimate, especially when it is large, and the sample size is small.
#
# To do so, one often estimates the bias as $- \frac{1+3\hat{\phi}}{T}$ (i.e. by replacing the true value $\phi$ by its estimate $\hat{\phi}$, so that the bias-corrected estimate is:
# $$
# \hat{\phi} + \frac{1+3\hat{\phi}}{T}
# $$
# #### AIC, BIC for model selection
y_arma2_long = simulate_arma(ar=[0.3], nsample=1000).squeeze()
res = sm.tsa.arma_order_select_ic(y_arma2_long, ic=["aic", "bic"], trend="nc")
res
# #### Esimation by statsmodels
model = sm.tsa.arima.ARIMA(y_arma2_long, order=(1, 0, 0))
res = model.fit(cov_type='robust')
print(res.summary())
# #### Impulse response function
fig, ax = plt.subplots()
ax.plot(res.impulse_responses(steps=nlags))
ax.set_title('Impulse Response Functions')
# ### ADF test
# +
from statsmodels.tsa.adfvalues import mackinnonp, mackinnoncrit
def nth_moment(y, counts, center, n):
""" Calculates nth moment around 'center"""
return np.sum((y - center)**n) / np.sum(counts)
def calc_ols(y, x, addcon=True):
"""Calculate OLS coefficients, SEs, and other statistics from scratch"""
Nobs = y.shape[0]
if addcon:
X = np.c_[np.ones((Nobs, 1)), x] # append the [Nobs x 1] columns of ones.
else:
X = x
k = X.shape[1]
XX = np.dot(X.T, X) # Construct sample average of E[X'X]
Xy = np.dot(X.T, y) # Construct sample average of E[X'Y]
XX_inv = np.linalg.inv(XX)
### OLS estimator: algebraic solution for OLS. beta_hat = (E[X'X]^-1)*(E[X'Y])
beta_hat = np.linalg.solve(XX, Xy)
### R-squared (can use np.dot or @ for "dot product")
y_hat = np.dot(X, beta_hat)
resids = y - y_hat
sse = np.dot(resids.T, resids)
sst = (y - np.mean(y)).T@(y - np.mean(y))
r_squared = 1 - (sse/sst)
### Adjusted R-squared
r_squared_adj = r_squared - (1 - r_squared)*((k - 1)/(Nobs - k))
### Variance-Covariance matrix: non-robust homoskedastic SEs
sigma2_hat = sse / (Nobs - k)
sigma2_mle = sse / (Nobs - 1)
cov_matrix = np.dot(XX_inv, sigma2_hat)
se = cov_matrix.diagonal()**0.5
t_stat = beta_hat / se
### Variance-Covariance matrix: White (robust) heteroskedastic SEs:
cov_matrix_robust = np.dot(np.dot(np.dot(np.dot(XX_inv, X.T), np.diag(resids**2)),X),XX_inv)
cov_matrix_robust_unbiased = cov_matrix_robust
se_robust = cov_matrix_robust_unbiased.diagonal()**0.5
t_stat_robust = beta_hat / se_robust
### log-likelihood (assumption of normality:
loglikelihood = - (Nobs/2)*np.log(2*np.math.pi) - \
(Nobs/2)*np.log(sigma2_mle) - \
(1/(2*sigma2_mle))*sum(resids**2)
### Information criteria:
hannan_quinn = ((-2*loglikelihood)/Nobs) + ((2*(k - 1)*np.log(np.log(Nobs)))/Nobs)
aic = Nobs *(np.log(2*np.pi) + 1 + np.log((sum(resids**2)/(Nobs)))) + (((k - 1) + 1)*2)
bic = -2*loglikelihood + np.log(Nobs)*((k - 1) + 1)
### Higher-order moments of the residuals:
m1 = nth_moment(resids, resids.shape[0], center=0, n=1)
m2 = nth_moment(resids, resids.shape[0], center=m1, n=2)
m3 = nth_moment(resids, resids.shape[0], center=m1, n=3)
m4 = nth_moment(resids, resids.shape[0], center=m1, n=4)
skew = m3 / (m2**(3/2))
kurtosis = m4 / (m2**2)
ols_results = {'beta_hat': beta_hat,
'standard-errors': se,
't-stat': t_stat,
'standard-errors (robust)': se_robust,
't-stat (robust)': t_stat_robust}
stats = {"No. Observations": Nobs,
"Df Residuals": Nobs - k,
"Skewness of residuals:": skew,
"Kurtosis of residuals:": kurtosis,
"$R^2$": r_squared,
"adjusted-$R^2$": r_squared_adj,
"Log-likelihood": loglikelihood,
"Hannan-Quinn": hannan_quinn,
"AIC": aic,
"BIC": bic}
return ols_results, stats
def adf_test(y, nlags):
""" ADF test for time series stationarity.
Augmented Dickey-Fuller test with p-values obtained through
regression surface approximation from MacKinnon [1994], similar to
the statsmodels routine: statsmodels.tsa.stattools.adfuller
Args:
y: np.array, time series.
nlags: int, number of lags in ADF test.
"""
diff_z = np.diff(y, axis=0)
z = y[:-1]
ols_results, _ = calc_ols(diff_z, z)
df_stat = ols_results['t-stat'][1]
p_value = mackinnonp(df_stat, regression='c', N=1)
cr_val = mackinnoncrit(N=1, regression='c', nobs=len(y))
print('(A)DF critical values | 1%: {:.3f}, 5%:{:.3f}, 10%:{:.3f}'.format(cr_val[0], cr_val[1], cr_val[2]))
print('df test, (no lags) | df-stat: {:.3f}, p-value: {:.3f}'.format(df_stat, p_value))
for i in range(1, nlags + 1):
z = np.column_stack((z[1:], diff_z[:-i]))
ols_results, _ = calc_ols(diff_z[i:], z)
adf_stat = ols_results['t-stat'][1]
p_value = mackinnonp(adf_stat, regression='c', N=1)
print('ADF test, lag# = '+ str(i),' | ADF-stat: {:.3f}, p-value: {:.3f}'.format(adf_stat, p_value))
# -
# random walk
random_walk = simulate_arma(ar=[1], nsample=1000).squeeze()
adf_test(random_walk, nlags=1)
# The null hypothesis for ADF test is the process has a unit root. From the test result it can be seen that we cannot reject the null, and indeed the time series is nonstationary (random walk).
| notebooks/.ipynb_checkpoints/time_series_analysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="AF10Q1bvnh6_" outputId="7d7efa79-539f-42c6-be1b-c3584faf1021"
# !wget https://www.fs.usda.gov/rds/archive/products/RDS-2013-0009.4/RDS-2013-0009.4_GPKG.zip
# + colab={"base_uri": "https://localhost:8080/"} id="FmQPuqTfn8g5" outputId="77ba6220-cfeb-4a7e-8ab7-7736714c4486"
# !unzip RDS-2013-0009.4_GPKG.zip
# + id="36nydPcEkS1V"
# %%time
# Important library for many geopython libraries
# !apt install gdal-bin python-gdal python3-gdal
# Install rtree - Geopandas requirment
# !apt install python3-rtree
# Install Geopandas
# !pip install git+git://github.com/geopandas/geopandas.git
# Install descartes - Geopandas requirment
# !pip install descartes
# Install Folium for Geographic data visualization
# !pip install folium
# Install plotlyExpress
# !pip install plotly_express
# !pip3 install contextily
# + id="PgwX5_EZlqZa"
import pandas as pd
import geopandas
import matplotlib.pyplot as plt
import numpy as np
import os
# + id="ZbiOtDwPnOhv"
us_gdf = geopandas.read_file("/content/Data/FPA_FOD_20170508.gpkg", layer='Fires')
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="fnpekBHpoXug" outputId="596af85e-63a8-4da2-c04e-c92f2b0f303f"
us_gdf.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="4eXkbQkWu4Xy" outputId="f5ef53bb-e82c-4736-8afb-054d26f5b33d"
us_gdf.columns
# + colab={"base_uri": "https://localhost:8080/"} id="-6Q12tkWvIE1" outputId="6ad239b7-9f31-45cc-bd15-baae9b0af22b"
len(us_gdf)
# + colab={"base_uri": "https://localhost:8080/"} id="O4XPtG4-vOWY" outputId="7865e29c-90ce-4528-cc91-37426d10b9a1"
len(us_gdf.FOD_ID.unique())
# + id="PHKExnoWltLu"
us_gdf = us_gdf[['FOD_ID','FIRE_YEAR', 'DISCOVERY_DATE', 'DISCOVERY_DOY',
'STAT_CAUSE_DESCR', 'FIRE_SIZE', 'STATE', 'geometry']]
# + id="YPfIyCPRpL5m"
virginia=us_gdf[us_gdf['STATE']=='VA']
# + id="kYuBmKEX31qD"
del us_gdf
# + colab={"base_uri": "https://localhost:8080/", "height": 194} id="4cqk4K6Ep9m7" outputId="6fb3548b-ad05-474e-b9a4-a72116ec1222"
virginia.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 194} id="ZxEFXWJiqEEr" outputId="9cde1210-1eac-48d0-d732-b09b76768311"
virginia.drop('STATE', axis=1, inplace=True)
virginia.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 194} id="ywDh85SCqqjw" outputId="b0aefd3b-58a9-42a8-9e5d-e4cfe52a9a97"
virginia = virginia.dropna()
virginia.head()
# + id="m10M89gzw307"
virginia['lon'] = virginia.geometry.apply(lambda p: p.x)
virginia['lat'] = virginia.geometry.apply(lambda p: p.y)
# + colab={"base_uri": "https://localhost:8080/", "height": 214} id="HSeHBbzXzqII" outputId="bf29c882-199e-4b3a-bbb0-c993e8c2d9b8"
virginia.plot()
# + id="8AOxlLigyBxS"
virginia_nogeom=virginia.drop('geometry', axis=1)
# + id="sTZ7Q2tHq7yd"
virginia_nogeom.to_csv("virginia_cleaned.csv",index=False,mode='w')
# + colab={"base_uri": "https://localhost:8080/"} id="ul-lmW9lx9r5" outputId="1b386e53-06a1-4e69-9195-c52a9daf89fc"
len(virginia)
# + colab={"base_uri": "https://localhost:8080/", "height": 528} id="Mi8rLyDlyU8Y" outputId="a26aea46-9932-4884-bc14-e2ef78d7bdd7"
plt.figure(figsize=(8,8))
cause_df = virginia['STAT_CAUSE_DESCR'].value_counts()
plt.title('Distribution by cause', y=-0.15)
plt.pie(cause_df, labels=list(cause_df.index[:-2]) + ['', '']);
plt.axis('equal');
| Scripts/Process_FPA_FOD_Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mawhy/OpenCV/blob/master/Image_Registration.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="8tSrBKMUl2hQ" colab_type="text"
# # Image Processing CookBook
# ## Image Registration
# In Polish means "nakładanie obrazów"
# + id="H0kx8xEQ0dPn" colab_type="code" colab={}
# !git clone https://github.com/PacktPublishing/Python-Image-Processing-Cookbook.git
# %cp -av "/content/Python-Image-Processing-Cookbook/Chapter 05/images/" "/content/"
# %cp -av "/content/Python-Image-Processing-Cookbook/Chapter 05/models/" "/content/"
# %rm -rf "/content/Python-Image-Processing-Cookbook"
# + [markdown] id="Ygwzf0wnl2hR" colab_type="text"
# ### Medical Image Registration with SimpleITK
# + id="e_V5Qlmy0wkb" colab_type="code" colab={}
# !pip install SimpleITK
# + id="bs0SiA5XoBV2" colab_type="code" colab={}
# !pip install
# !pip install opencv-python==4.2.0.34
# !pip install opencv-contrib-python==4.2.0.34
# + id="aBgdSGPGl2hS" colab_type="code" colab={}
# https://stackoverflow.com/questions/41692063/what-is-the-difference-between-image-registration-and-image-alignment
# https://www.insight-journal.org/rire/download_training_data.php
# https://itk.org/Wiki/SimpleITK/Tutorials/MICCAI2015
# %matplotlib inline
import SimpleITK as sitk
import numpy as np
import matplotlib.pyplot as plt
fixed_image = sitk.ReadImage("images/ct_scan_11.jpg", sitk.sitkFloat32)
moving_image = sitk.ReadImage("images/mr_T1_06.jpg", sitk.sitkFloat32)
fixed_image_array = sitk.GetArrayFromImage(fixed_image)
moving_image_array = sitk.GetArrayFromImage(moving_image)
print(fixed_image_array.shape, moving_image_array.shape)
plt.figure(figsize=(20,10))
plt.gray()
plt.subplot(131), plt.imshow(fixed_image_array), plt.axis('off'), plt.title('CT-Scan Image', size=20)
plt.subplot(132), plt.imshow(moving_image_array), plt.axis('off'), plt.title('MRI-T1 Image', size=20)
plt.subplot(133), plt.imshow(0.6*fixed_image_array + 0.4*moving_image_array), plt.axis('off'), plt.title('Initial Alignment', size=20)
plt.show()
# + id="GpS4H-Wql2hb" colab_type="code" colab={}
np.random.seed(2)
registration_method = sitk.ImageRegistrationMethod()
initial_transform = sitk.CenteredTransformInitializer(fixed_image, moving_image, sitk.Similarity2DTransform())
# Similarity metric settings.
registration_method.SetMetricAsMattesMutualInformation(numberOfHistogramBins=50)
registration_method.SetMetricSamplingStrategy(registration_method.RANDOM)
registration_method.SetMetricSamplingPercentage(0.01)
registration_method.SetInterpolator(sitk.sitkLinear)
# Optimizer settings.
registration_method.SetOptimizerAsGradientDescent(learningRate=1.0, numberOfIterations=100, convergenceMinimumValue=1e-6, convergenceWindowSize=10)
registration_method.SetOptimizerScalesFromPhysicalShift()
# Setup for the multi-resolution framework.
registration_method.SetShrinkFactorsPerLevel(shrinkFactors = [4,2,1])
registration_method.SetSmoothingSigmasPerLevel(smoothingSigmas=[2,1,0])
registration_method.SmoothingSigmasAreSpecifiedInPhysicalUnitsOn()
# Don't optimize in-place, we would possibly like to run this cell multiple times.
registration_method.SetInitialTransform(initial_transform, inPlace=False)
final_transform = registration_method.Execute(sitk.Cast(fixed_image, sitk.sitkFloat32),
sitk.Cast(moving_image, sitk.sitkFloat32))
# + id="rC0i82K2l2hd" colab_type="code" colab={}
#print(final_transform)
resampler = sitk.ResampleImageFilter()
resampler.SetReferenceImage(fixed_image);
resampler.SetInterpolator(sitk.sitkLinear)
resampler.SetDefaultPixelValue(100)
resampler.SetTransform(final_transform)
out = resampler.Execute(moving_image)
simg1 = sitk.Cast(sitk.RescaleIntensity(fixed_image), sitk.sitkUInt8)
simg2 = sitk.Cast(sitk.RescaleIntensity(out), sitk.sitkUInt8)
cimg = sitk.Compose(simg1, simg2, simg1//2.+simg2//2.)
plt.figure(figsize=(20,10))
plt.gray()
plt.subplot(131), plt.imshow(fixed_image_array), plt.axis('off'), plt.title('CT-Scan Image', size=20)
plt.subplot(132), plt.imshow(sitk.GetArrayFromImage(out)), plt.axis('off'), plt.title('Transformed MRI-T1 Image', size=20)
plt.subplot(133), plt.imshow(sitk.GetArrayFromImage(cimg)), plt.axis('off'), plt.title('Final Alignment', size=20)
plt.show()
# + id="2PUVQ88Fl2hg" colab_type="code" colab={}
# https://www.insight-journal.org/rire/download_training_data.php
# https://itk.org/Wiki/SimpleITK/Tutorials/MICCAI2015
import SimpleITK as sitk
import matplotlib.pyplot as plt
fixed = sitk.ReadImage("images/mr_T1_01.jpg", sitk.sitkFloat32)
moving = sitk.ReadImage("images/mr_T1_01_trans.jpg", sitk.sitkFloat32)
R = sitk.ImageRegistrationMethod()
R.SetMetricAsMeanSquares()
R.SetOptimizerAsRegularStepGradientDescent(4.0, .01, 200 )
R.SetInterpolator(sitk.sitkLinear)
transfo = sitk.CenteredTransformInitializer(fixed, moving, sitk.Euler2DTransform())
R.SetInitialTransform(transfo)
outTx1 = R.Execute(fixed, moving)
print(outTx1)
print("Optimizer stop condition: {0}".format(R.GetOptimizerStopConditionDescription()))
print("Number of iterations: {0}".format(R.GetOptimizerIteration()))
R.SetMetricAsMattesMutualInformation(numberOfHistogramBins=50)
R.SetOptimizerAsRegularStepGradientDescent(4.0, .01, 200 )
R.SetInitialTransform(transfo)
outTx2 = R.Execute(fixed, moving)
print(outTx2)
print("Optimizer stop condition: {0}".format(R.GetOptimizerStopConditionDescription()))
print("Number of iterations: {0}".format(R.GetOptimizerIteration()))
#sitk.WriteTransform(outTx, 'transfo_final.tfm')
resampler = sitk.ResampleImageFilter()
resampler.SetReferenceImage(fixed)
resampler.SetInterpolator(sitk.sitkLinear)
resampler.SetDefaultPixelValue(100)
resampler.SetTransform(outTx1)
out1 = resampler.Execute(moving)
moving_image_array_trans1 = sitk.GetArrayFromImage(out1)
simg1 = sitk.Cast(sitk.RescaleIntensity(fixed), sitk.sitkUInt8)
simg2 = sitk.Cast(sitk.RescaleIntensity(out1), sitk.sitkUInt8)
cimg1_array = sitk.GetArrayFromImage(sitk.Compose(simg1, simg2, simg1//2.+simg2//2.))
resampler.SetTransform(outTx2)
out2 = resampler.Execute(moving)
moving_image_array_trans2 = sitk.GetArrayFromImage(out2)
simg1 = sitk.Cast(sitk.RescaleIntensity(fixed), sitk.sitkUInt8)
simg2 = sitk.Cast(sitk.RescaleIntensity(out2), sitk.sitkUInt8)
cimg2_array = sitk.GetArrayFromImage(sitk.Compose(simg1, simg2, simg1//2.+simg2//2.))
fixed_image_array = sitk.GetArrayFromImage(fixed)
moving_image_array = sitk.GetArrayFromImage(moving)
print(fixed_image_array.shape, moving_image_array.shape)
plt.figure(figsize=(20,30))
plt.gray()
plt.subplots_adjust(0,0,1,1,0.075,0.01)
plt.subplot(321), plt.imshow(fixed_image_array), plt.axis('off'), plt.title('MR-T1 Image', size=20)
plt.subplot(322), plt.imshow(moving_image_array), plt.axis('off'), plt.title('Shifted MR_T1 Image', size=20)
plt.subplot(323), plt.imshow(fixed_image_array - moving_image_array_trans1), plt.axis('off'), plt.title('Difference Images (MeanSquare)', size=20)
plt.subplot(324), plt.imshow(fixed_image_array - moving_image_array_trans2), plt.axis('off'), plt.title('Difference Images (MutualInformation)', size=20)
plt.subplot(325), plt.imshow(cimg1_array), plt.axis('off'), plt.title('Aligned Images (MeanSquare)', size=20)
plt.subplot(326), plt.imshow(cimg2_array), plt.axis('off'), plt.title('Aligned Images (MutualInformation)', size=20)
plt.show()
# + id="mAGFedoLl2hi" colab_type="code" colab={}
checkerboard = sitk.CheckerBoardImageFilter()
before_reg_image = checkerboard.Execute (fixed, moving)
after_reg_image = checkerboard.Execute (fixed, out2)
plt.figure(figsize=(20,10))
plt.gray()
plt.subplot(121), plt.imshow(sitk.GetArrayFromImage(before_reg_image)), plt.axis('off'), plt.title('Checkerboard before Registration Image', size=20)
plt.subplot(122), plt.imshow(sitk.GetArrayFromImage(after_reg_image)), plt.axis('off'), plt.title('Checkerboard After Registration Image', size=20)
plt.show()
# + [markdown] id="o64KqViKl2hl" colab_type="text"
# ### Image Alignment with ECC algorithm
# [Good articles](https://www.learnopencv.com/image-alignment-ecc-in-opencv-c-python/)
# + id="n4rRqlNMl2hm" colab_type="code" colab={}
import cv2
print(cv2.__version__)
# 4.2.0
import numpy as np
import matplotlib.pylab as plt
def compute_gradient(im) :
grad_x = cv2.Sobel(im,cv2.CV_32F,1,0,ksize=3)
grad_y = cv2.Sobel(im,cv2.CV_32F,0,1,ksize=3)
grad = cv2.addWeighted(np.absolute(grad_x), 0.5, np.absolute(grad_y), 0.5, 0)
return grad
im_unaligned = cv2.imread("images/me_unaligned.png")
height, width = im_unaligned.shape[:2]
print(height, width)
channels = ['B', 'G', 'R']
plt.figure(figsize=(30,12))
plt.gray()
plt.subplot(1,4,1), plt.imshow(cv2.cvtColor(im_unaligned, cv2.COLOR_BGR2RGB)), plt.axis('off'), plt.title('Unaligned Image', size=20)
for i in range(3):
plt.subplot(1,4,i+2), plt.imshow(im_unaligned[...,i]), plt.axis('off'), plt.title(channels[i], size=20)
plt.suptitle('Unaligned Image and Color Channels', size=30)
plt.show()
# Initialize the output image with a copy of the input image
im_aligned = im_unaligned.copy()
# Define motion model
warp_mode = cv2.MOTION_HOMOGRAPHY
# Set the warp matrix to identity.
warp_matrix = np.eye(3, 3, dtype=np.float32) if warp_mode == cv2.MOTION_HOMOGRAPHY else np.eye(2, 3, dtype=np.float32)
# Set the stopping criteria for the algorithm.
criteria = (cv2.TERM_CRITERIA_EPS | cv2.TERM_CRITERIA_COUNT, 500, 1e-6)
# The blue and green channels will be aligned to the red channel, so compute the gradient of the red channel first
im_grad2 = compute_gradient(im_unaligned[...,2])
# Warp the blue and green channels to the red channel
for i in range(2) :
print('Processing Channel {}...'.format(channels[i]))
(cc, warp_matrix) = cv2.findTransformECC (im_grad2, compute_gradient(im_unaligned[...,i]),warp_matrix, warp_mode, criteria, None, 5)
if warp_mode == cv2.MOTION_HOMOGRAPHY :
# Perspective warp - transformation is a Homography
im_aligned[...,i] = cv2.warpPerspective (im_unaligned[...,i], warp_matrix, (width,height), flags=cv2.INTER_LINEAR + cv2.WARP_INVERSE_MAP)
else :
# Affine warp - transformation is not a Homography
im_aligned[...,i] = cv2.warpAffine(im_unaligned[...,i], warp_matrix, (width, height), flags=cv2.INTER_LINEAR + cv2.WARP_INVERSE_MAP);
print (warp_matrix)
channels = ['B', 'G', 'R']
plt.figure(figsize=(30,12))
plt.subplot(1,4,1), plt.imshow(cv2.cvtColor(im_aligned, cv2.COLOR_BGR2RGB)), plt.axis('off'), plt.title('Aligned Image (ECC)', size=20)
for i in range(3):
plt.subplot(1,4,i+2), plt.imshow(im_aligned[...,i]), plt.axis('off'), plt.title(channels[i], size=20)
plt.suptitle('Aligned Image and Color Channels', size=30)
plt.show()
cv2.imwrite("images/me_aligned.png", im_aligned)
# + [markdown] id="Ez9mIo_vl2ho" colab_type="text"
# ### Face Alignment
# + id="9Egqyak6l2hp" colab_type="code" colab={}
from imutils.face_utils import FaceAligner
from imutils.face_utils import rect_to_bb
import imutils
import dlib
import cv2
import matplotlib.pylab as plt
# initialize dlib's face detector (HOG-based) and then create
# the facial landmark predictor and the face aligner
detector = dlib.get_frontal_face_detector()
predictor = dlib.shape_predictor('models/shape_predictor_68_face_landmarks.dat')
face_aligner = FaceAligner(predictor, desiredFaceWidth=256)
# load the input image, resize it, and convert it to grayscale
image = cv2.imread('images/scientists.png')
image = imutils.resize(image, width=800)
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
# show the original input image and detect faces in the grayscale image
plt.figure(figsize=(20,20))
plt.imshow(cv2.cvtColor(image, cv2.COLOR_BGR2RGB)), plt.axis('off')
plt.title('Original Image: Famous Indian Scientists', size=20)
plt.show()
rects = detector(gray, 2)
print('Number of faces detected:', len(rects))
i = 1
# loop over the face detections
plt.figure(figsize=(10,20))
plt.subplots_adjust(0,0,1,1,0.05,0.12)
for rect in rects:
# extract the ROI of the *original* face, then align the face
# using facial landmarks
(x, y, w, h) = rect_to_bb(rect)
face_original = imutils.resize(image[y:y + h, x:x + w], width=256)
face_aligned = face_aligner.align(image, gray, rect)
# display the output images
plt.subplot(9,4,i), plt.imshow(cv2.cvtColor(face_original, cv2.COLOR_BGR2RGB)), plt.title("Original", size=15), plt.axis('off')
plt.subplot(9,4,i+1), plt.imshow(cv2.cvtColor(face_aligned, cv2.COLOR_BGR2RGB)), plt.title("Aligned", size=15), plt.axis('off')
i += 2
plt.show()
# + id="AnyRjGe_l2hr" colab_type="code" colab={}
import dlib
import cv2
from imutils import face_utils
from skimage.transform import AffineTransform, warp
import numpy as np
import matplotlib.pylab as plt
# initialize dlib's face detector (HOG-based) and then create
# the facial landmark predictor and the face aligner
detector = dlib.get_frontal_face_detector()
predictor = dlib.shape_predictor('models/shape_predictor_68_face_landmarks.dat')
# load the input image, resize it, and convert it to grayscale
image = cv2.imread('images/monalisa.png')
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
rects = detector(gray, 2)
faces = []
face_landmarks = []
for (i, rect) in enumerate(rects):
shape = predictor(gray, rect)
shape = face_utils.shape_to_np(shape)
(left, top, w, h) = face_utils.rect_to_bb(rect)
faces.append(image[top:top+h, left:left+w])
landmark = []
for (x, y) in shape:
cv2.circle(image, (x, y), 1, (0, 255, 0), 2)
landmark.append([x-left,y-top])
face_landmarks.append(np.array(landmark))
plt.figure(figsize=(20,13))
plt.imshow(cv2.cvtColor(image, cv2.COLOR_BGR2RGB)), plt.axis('off'), plt.title('Original image with Facial landmarks', size=20)
plt.show()
plt.figure(figsize=(20,10))
plt.subplot(121), plt.imshow(cv2.cvtColor(faces[0], cv2.COLOR_BGR2RGB)), plt.axis('off'), plt.title('Right Face', size=20)
plt.subplot(122), plt.imshow(cv2.cvtColor(faces[1], cv2.COLOR_BGR2RGB)), plt.axis('off'), plt.title('Left Face', size=20)
plt.show()
transform = AffineTransform()
transform.estimate(face_landmarks[0], face_landmarks[1])
plt.figure(figsize=(10,10))
plt.gray()
plt.imshow(warp(cv2.cvtColor(faces[1], cv2.COLOR_BGR2RGB), transform, output_shape=faces[0].shape)), plt.axis('off'), plt.title('Warped right image on the left image', size=20)
plt.show()
# + [markdown] id="6zCBlX6gl2hu" colab_type="text"
# ### Face Morphing
# + id="1tn4v551l2hu" colab_type="code" colab={}
from scipy.spatial import Delaunay
from scipy import interpolate
from skimage.io import imread
import scipy.misc
import cv2
import dlib
import numpy as np
from matplotlib import pyplot as plt
# Find 68 face landmarks using dlib
def get_face_landmarks(image, predictor_path = 'models/shape_predictor_68_face_landmarks.dat'):
detector = dlib.get_frontal_face_detector()
predictor = dlib.shape_predictor(predictor_path)
try:
dets = detector(image, 1)
points = np.zeros((68, 2))
for k, d in enumerate(dets):
# get the landmarks for the face in box d.
shape = predictor(image, d)
for i in range(68):
points[i, 0] = shape.part(i).x
points[i, 1] = shape.part(i).y
except Exception as e:
print('Failed finding face points: ', e)
return []
points = points.astype(np.int32)
return points
def weighted_average_points(start_points, end_points, percent=0.5):
# Weighted average of two sets of supplied points
if percent <= 0: return end_points
elif percent >= 1: return start_points
else: return np.asarray(start_points*percent + end_points*(1-percent), np.int32)
def weighted_average(img1, img2, percent=0.5):
if percent <= 0: return img2
elif percent >= 1: return img1
else: return cv2.addWeighted(img1, percent, img2, 1-percent, 0)
# interpolates over every image channel
def bilinear_interpolate(img, coords):
int_coords = coords.astype(np.int32)
x0, y0 = int_coords
dx, dy = coords - int_coords
# 4 neighour pixels
q11, q21, q12, q22 = img[y0, x0], img[y0, x0+1], img[y0+1, x0], img[y0+1, x0+1]
btm = q21.T * dx + q11.T * (1 - dx)
top = q22.T * dx + q12.T * (1 - dx)
interpolated_pixels = top * dy + btm * (1 - dy)
return interpolated_pixels.T
# generate x,y grid coordinates within the ROI of supplied points
def get_grid_coordinates(points):
xmin, xmax = np.min(points[:, 0]), np.max(points[:, 0]) + 1
ymin, ymax = np.min(points[:, 1]), np.max(points[:, 1]) + 1
return np.asarray([(x, y) for y in range(ymin, ymax) for x in range(xmin, xmax)], np.uint32)
# warp each triangle from the src_image only within the ROI of the destination image (points in dst_points).
def process_warp(src_img, result_img, tri_affines, dst_points, delaunay):
roi_coords = get_grid_coordinates(dst_points)
# indices to vertices. -1 if pixel is not in any triangle
roi_tri_indices = delaunay.find_simplex(roi_coords)
for simplex_index in range(len(delaunay.simplices)):
coords = roi_coords[roi_tri_indices == simplex_index]
num_coords = len(coords)
out_coords = np.dot(tri_affines[simplex_index], np.vstack((coords.T, np.ones(num_coords))))
x, y = coords.T
result_img[y, x] = bilinear_interpolate(src_img, out_coords)
# calculate the affine transformation matrix for each triangle vertex (x,y) from dest_points to src_points
def gen_triangular_affine_matrices(vertices, src_points, dest_points):
ones = [1, 1, 1]
for tri_indices in vertices:
src_tri = np.vstack((src_points[tri_indices, :].T, ones))
dst_tri = np.vstack((dest_points[tri_indices, :].T, ones))
mat = np.dot(src_tri, np.linalg.inv(dst_tri))[:2, :]
yield mat
def warp_image(src_img, src_points, dest_points, dest_shape):
num_chans = 3
src_img = src_img[:, :, :3]
rows, cols = dest_shape[:2]
result_img = np.zeros((rows, cols, num_chans), np.uint8)
delaunay = Delaunay(dest_points)
tri_affines = np.asarray(list(gen_triangular_affine_matrices(delaunay.simplices, src_points, dest_points)))
process_warp(src_img, result_img, tri_affines, dest_points, delaunay)
return result_img, delaunay
def read_lion_landmarks():
with open("models/lion_face_landmark.txt") as key_file:
keypoints = [list(map(int, line.split())) for line in key_file]
return(np.array(keypoints))
# load images
src_path = 'images/me.png'
dst_path = 'images/lion.png'
src_img = imread(src_path)
dst_img = imread(dst_path)
size = dst_img.shape[:2]
src_img = cv2.resize(src_img[...,:3], size)
# define control points for warps
src_points = get_face_landmarks(src_img)
dst_points = read_lion_landmarks()
points = weighted_average_points(src_points, dst_points, percent=50)
src_face, src_delaunay = warp_image(src_img, src_points, points, size)
end_face, end_delaunay = warp_image(dst_img, dst_points, points, size)
print('here', len(src_points), len(dst_points))
fig = plt.figure(figsize=(20,10))
plt.subplot(121), plt.imshow(src_img)
for i in range(len(src_points)):
plt.plot(src_points[i,0], src_points[i,1], 'r.', markersize=20)
plt.title('Source image', size=20), plt.axis('off')
plt.subplot(122), plt.imshow(dst_img)
for i in range(len(dst_points)):
plt.plot(dst_points[i,0], dst_points[i,1], 'g.', markersize=20)
plt.title('Destination image', size=20), plt.axis('off')
plt.suptitle('Facial Landmarks computed for the images', size=30)
fig.subplots_adjust(wspace=0.01, left=0.1, right=0.9)
plt.show()
fig = plt.figure(figsize=(20,10))
plt.subplot(121), plt.imshow(src_img)
plt.triplot(src_points[:,0], src_points[:,1], src_delaunay.simplices.copy())
plt.plot(src_points[:,0], src_points[:,1], 'o', color='red'), plt.title('Source image', size=20), plt.axis('off')
plt.subplot(122), plt.imshow(dst_img)
plt.triplot(dst_points[:,0], dst_points[:,1], end_delaunay.simplices.copy())
plt.plot(dst_points[:,0], dst_points[:,1], 'o'), plt.title('Destination image', size=20), plt.axis('off')
plt.suptitle('Delaunay triangulation of the images', size=30)
fig.subplots_adjust(wspace=0.01, left=0.1, right=0.9)
plt.show()
fig = plt.figure(figsize=(18,20))
fig.subplots_adjust(top=0.925, bottom=0, left=0, right=1, wspace=0.01, hspace=0.08)
i = 1
for percent in np.linspace(1, 0, 16):
points = weighted_average_points(src_points, dst_points, percent)
src_face, src_delaunay = warp_image(src_img, src_points, points, size)
end_face, end_delaunay = warp_image(dst_img, dst_points, points, size)
average_face = weighted_average(src_face, end_face, percent)
plt.subplot(4,4,i), plt.imshow(average_face), plt.title('alpha=' + str(round(percent,4)), size=20), plt.axis('off')
i += 1
plt.suptitle('Face morphing', size=30)
plt.show()
# + [markdown] id="Qhyq4cRZl2hw" colab_type="text"
# ### Robust Matching with RANSAC Algorithm using Harris Corner Brief Descriptors
# [<NAME>](https://www.researchgate.net/publication/292995470_Image_Features_Detection_Description_and_Matching)
# + id="szF-Rg7Xl2hx" colab_type="code" colab={}
from skimage.feature import (corner_harris, corner_peaks, BRIEF, match_descriptors, plot_matches)
from skimage.transform import ProjectiveTransform, warp
from skimage.measure import ransac
from skimage.io import imread
from skimage.color import rgb2gray
import matplotlib.pylab as plt
np.random.seed(2)
img1 = rgb2gray(imread('images/victoria3.png'))
img2 = rgb2gray(imread('images/victoria4.png'))
keypoints1 = corner_peaks(corner_harris(img1), min_distance=1)
keypoints2 = corner_peaks(corner_harris(img2), min_distance=1)
extractor = BRIEF(patch_size=10)
extractor.extract(img1, keypoints1)
descriptors1 = extractor.descriptors
extractor.extract(img2, keypoints2)
descriptors2 = extractor.descriptors
matches = match_descriptors(descriptors1, descriptors2)
src_keypoints = keypoints1[matches[:,0]]
dst_keypoints = keypoints2[matches[:,1]]
homography = ProjectiveTransform()
homography.estimate(src_keypoints, dst_keypoints)
homography_robust, inliers = ransac((src_keypoints, dst_keypoints), ProjectiveTransform, min_samples=4,
residual_threshold=2, max_trials=500)
outliers = inliers == False
print(len(matches))
robust_matches = match_descriptors(descriptors1[matches[:,0]][inliers], descriptors2[matches[:,1]][inliers])
print(len(robust_matches))
fig, ax = plt.subplots(nrows=2, ncols=2, sharex=True, sharey=True, figsize=(20,15))
plt.gray()
plt.subplots_adjust(0,0,1,1,0.05,0.05)
plot_matches(ax[0,0], img1, img2, keypoints1, keypoints2, matches), ax[0,0].set_title('Matching without RANSAC', size=20)
ax[0,1].imshow(warp(img2, homography, output_shape=img2.shape)), ax[0,1].set_title('Homography without RANSAC', size=20)
plot_matches(ax[1,0], img1, img2, keypoints1, keypoints2, robust_matches), ax[1,0].set_title('Robust Matching with RANSAC', size=20)
ax[1,1].imshow(warp(img2, homography_robust, output_shape=img2.shape)), ax[1,1].set_title('Robust Homography with RANSAC', size=20)
for a in np.ravel(ax):
a.axis('off')
plt.show()
# + id="ehKTzvqZqmQ1" colab_type="code" colab={}
# !pip install opencv-python==3.4.2.16
# !pip install opencv-contrib-python==3.4.2.16
# + [markdown] id="Ty6LxJIFl2hz" colab_type="text"
# ### Image Mosaicing (Cylindrical Panorama)
# + id="B0AOwX9ql2h0" colab_type="code" colab={}
import cv2
# for this problem let's work with opencv 3.4.2.16
print(cv2.__version__)
# 3.4.2
# pip install opencv-contrib-python==3.4.2.16
# pip install opencv-python==3.4.2.16
import numpy as np
from matplotlib import pyplot as plt
import math
import glob
def compute_homography(image1, image2, bff_match=False):
sift = cv2.xfeatures2d.SIFT_create(edgeThreshold=10, sigma=1.5, contrastThreshold=0.08)
kp1, des1 = sift.detectAndCompute(image1, None)
kp2, des2 = sift.detectAndCompute(image2, None)
# Brute force matching
bf = cv2.BFMatcher()
matches = bf.knnMatch(des1, trainDescriptors=des2, k=2)
# Lowes Ratio
good_matches = []
for m, n in matches:
if m.distance < .75 * n.distance:
good_matches.append(m)
src_pts = np.float32([kp1[m.queryIdx].pt for m in good_matches])\
.reshape(-1, 1, 2)
dst_pts = np.float32([kp2[m.trainIdx].pt for m in good_matches])\
.reshape(-1, 1, 2)
if len(src_pts) > 4:
H, mask = cv2.findHomography(dst_pts, src_pts, cv2.RANSAC, 5)
else:
H = np.array([[0, 0, 0], [0, 0, 0], [0, 0, 0]])
return H
def warp_image(image, H):
image = cv2.cvtColor(image, cv2.COLOR_BGR2BGRA)
h, w, _ = image.shape
# Find min and max x, y of new image
p = np.array([[0, w, w, 0], [0, 0, h, h], [1, 1, 1, 1]])
p_prime = np.dot(H, p)
yrow = p_prime[1] / p_prime[2]
xrow = p_prime[0] / p_prime[2]
ymin = min(yrow)
xmin = min(xrow)
ymax = max(yrow)
xmax = max(xrow)
# Create a new matrix that removes offset and multiply by homography
new_mat = np.array([[1, 0, -1 * xmin], [0, 1, -1 * ymin], [0, 0, 1]])
H = np.dot(new_mat, H)
# height and width of new image frame
height = int(round(ymax - ymin))
width = int(round(xmax - xmin))
size = (width, height)
# Do the warp
warped = cv2.warpPerspective(src=image, M=H, dsize=size)
return warped, (int(xmin), int(ymin))
def cylindrical_warp_image(img, H):
h, w = img.shape[:2]
# pixel coordinates
y_i, x_i = np.indices((h, w))
X = np.stack([x_i,y_i,np.ones_like(x_i)],axis=-1).reshape(h*w, 3) # to homog
Hinv = np.linalg.inv(H)
X = Hinv.dot(X.T).T # normalized coords
# calculate cylindrical coords (sin\theta, h, cos\theta)
A = np.stack([np.sin(X[:,0]),X[:,1],np.cos(X[:,0])],axis=-1).reshape(w*h, 3)
B = H.dot(A.T).T # project back to image-pixels plane
# back from homog coords
B = B[:,:-1] / B[:,[-1]]
# make sure warp coords only within image bounds
B[(B[:,0] < 0) | (B[:,0] >= w) | (B[:,1] < 0) | (B[:,1] >= h)] = -1
B = B.reshape(h,w,-1)
img_rgba = cv2.cvtColor(img,cv2.COLOR_BGR2BGRA) # for transparent borders...
# warp the image according to cylindrical coords
return cv2.remap(img_rgba, B[:,:,0].astype(np.float32), B[:,:,1].astype(np.float32), cv2.INTER_AREA, borderMode=cv2.BORDER_TRANSPARENT)
def create_mosaic(images, origins):
# find central image
for i in range(0, len(origins)):
if origins[i] == (0, 0):
central_index = i
break
central_image = images[central_index]
central_origin = origins[central_index]
# zip origins and images together
zipped = list(zip(origins, images))
# sort by distance from origin (highest to lowest)
func = lambda x: math.sqrt(x[0][0] ** 2 + x[0][1] ** 2)
dist_sorted = sorted(zipped, key=func, reverse=True)
# sort by x value
x_sorted = sorted(zipped, key=lambda x: x[0][0])
# sort by y value
y_sorted = sorted(zipped, key=lambda x: x[0][1])
# determine the coordinates in the new frame of the central image
if x_sorted[0][0][0] > 0:
cent_x = 0 # leftmost image is central image
else:
cent_x = abs(x_sorted[0][0][0])
if y_sorted[0][0][1] > 0:
cent_y = 0 # topmost image is central image
else:
cent_y = abs(y_sorted[0][0][1])
# make a new list of the starting points in new frame of each image
spots = []
for origin in origins:
spots.append((origin[0]+cent_x, origin[1] + cent_y))
zipped = zip(spots, images)
# get height and width of new frame
total_height = 0
total_width = 0
for spot, image in zipped:
total_width = max(total_width, spot[0]+image.shape[1])
total_height = max(total_height, spot[1]+image.shape[0])
# print "height ", total_height
# print "width ", total_width
# new frame of panorama
stitch = np.zeros((total_height, total_width, 4), np.uint8)
# stitch images into frame by order of distance
for image in dist_sorted:
offset_y = image[0][1] + cent_y
offset_x = image[0][0] + cent_x
end_y = offset_y + image[1].shape[0]
end_x = offset_x + image[1].shape[1]
####
stitch_cur = stitch[offset_y:end_y, offset_x:end_x, :4]
stitch_cur[image[1]>0] = image[1][image[1]>0]
####
#stitch[offset_y:end_y, offset_x:end_x, :4] = image[1]
return stitch
def create_panorama(images, center):
h,w,_ = images[0].shape
f = 1000 # 800
H = np.array([[f, 0, w/2], [0, f, h/2], [0, 0, 1]])
for i in range(len(images)):
images[i] = cylindrical_warp_image(images[i], H)
panorama = None
for i in range(center):
print('Stitching images {}, {}'.format(i+1, i+2))
image_warped, image_origin = warp_image(images[i], compute_homography(images[i + 1], images[i]))
panorama = create_mosaic([image_warped, images[i+1]], [image_origin, (0,0)])
images[i + 1] = panorama
#print('Done left part')
for i in range(center, len(images)-1):
print('Stitching images {}, {}'.format(i+1, i+2))
image_warped, image_origin = warp_image(images[i+1], compute_homography(images[i], images[i + 1]))
panorama = create_mosaic([images[i], image_warped], [(0,0), image_origin])
images[i + 1] = panorama
#print('Done right part')
return panorama
images = [ cv2.cvtColor(cv2.imread(img), cv2.COLOR_RGB2RGBA) for img in glob.glob('images/victoria*.png')]
plt.figure(figsize=(20,4))
plt.subplots_adjust(top = 0.8, bottom = 0, right = 1, left = 0, hspace = 0, wspace = 0.05)
plt.margins(0,0)
for i in range(len(images)):
plt.subplot(1,len(images),i+1), plt.imshow(cv2.cvtColor(images[i], cv2.COLOR_BGR2RGB)), plt.axis('off'), plt.title('Image {}'.format(i+1), size=15)
plt.suptitle('Images to Stitch', size=20)
plt.show()
center = len(images) // 2
#print(len(images), center)
panorama = create_panorama(images, center)
plt.figure(figsize=(20,8))
plt.subplots_adjust(top = 0.9, bottom = 0, right = 1, left = 0, hspace = 0, wspace = 0)
plt.margins(0,0)
plt.imshow(cv2.cvtColor(panorama, cv2.COLOR_BGR2RGB)), plt.axis('off'), plt.title('Final Panorama Image', size=15)
plt.show()
# + [markdown] id="4IDidofTl2h2" colab_type="text"
# ### Panorama with opencv-python
# + id="SKaj8bcJl2h2" colab_type="code" colab={}
import numpy as np
import cv2
import glob
import matplotlib.pylab as plt
print(cv2.__version__)
# 3.4.2
# grab the paths to the input images and initialize our images list
print("Loading images...")
images = [ cv2.cvtColor(cv2.imread(img), cv2.COLOR_BGR2RGB) for img in glob.glob('images/victoria*.png')]
print('Number of images to stitch: {}'.format(len(images)))
fig = plt.figure(figsize=(20, 5))
for i in range(len(images)):
plt.subplot(1,len(images),i+1)
plt.imshow(images[i])
plt.axis('off')
fig.subplots_adjust(left=0, right=1, bottom=0, top=0.95, hspace=0.05, wspace=0.05)
plt.suptitle('Images to stich', size=25)
plt.show()
# initialize OpenCV's image sticher object and then perform the image
# stitching
print("Stitching images...")
stitcher = cv2.createStitcher()
(status, stitched) = stitcher.stitch(images)
print(status)
plt.figure(figsize=(20,10))
plt.imshow(stitched), plt.axis('off'), plt.title('Final Panorama Image', size=20)
plt.show()
# + [markdown] id="2wkwtpFEl2h4" colab_type="text"
# ### Finding similarity between an image and a set of images
# + id="8DzUZFIql2h5" colab_type="code" colab={}
import cv2
print(cv2.__version__)
# 3.4.2
import numpy as np
import glob
import matplotlib.pylab as plt
from collections import defaultdict
query = cv2.imread("images/query.png", cv2.CV_8U)
matched_images = defaultdict(list)
for image_file in glob.glob('images/search/*.png'):
search_image = cv2.imread(image_file, cv2.CV_8U)
sift = cv2.xfeatures2d.SIFT_create()
kp_1, desc_1 = sift.detectAndCompute(query, None)
kp_2, desc_2 = sift.detectAndCompute(search_image, None)
index_params = dict(algorithm=0, trees=5)
search_params = dict()
flann = cv2.FlannBasedMatcher(index_params, search_params)
matches = flann.knnMatch(desc_1, desc_2, k=2)
good_points = []
ratio = 0.6
for m, n in matches:
if m.distance < ratio*n.distance:
good_points.append(m)
num_good_points = len(good_points)
print('Image file = {}, Number of good matches = {}'.format(image_file, num_good_points))
if (num_good_points > 300) or (num_good_points < 10):
result = cv2.drawMatches(query, kp_1, search_image, kp_2, good_points, None)
plt.figure(figsize=(20,10))
plt.imshow(cv2.cvtColor(result, cv2.COLOR_BGR2RGB)), plt.axis('off')
plt.title(('Good match' if num_good_points > 300 else 'Poor match') + ' with {} matches'.format(num_good_points), size=20)
plt.show()
matched_images[len(good_points)].append(search_image)
plt.figure(figsize=(15,10))
plt.gray()
plt.imshow(query), plt.axis('off')
plt.title('Original (Query) Image', size=20)
plt.show()
i = 1
plt.figure(figsize=(20,35))
plt.subplots_adjust(left=0, right=1, bottom=0, top=0.925, wspace=0.02, hspace=0.1)
for num_matches in sorted(matched_images, reverse=True):
for image in matched_images[num_matches]:
plt.subplot(10, 4, i)
plt.imshow(image)
plt.axis('off')
plt.title('Image with {} good matches'.format(num_matches), size=15)
i += 1
plt.suptitle('Images matched with the Query Image ranked by the number of good matches', size=20)
plt.show()
# + id="exHTZwVyl2h7" colab_type="code" colab={}
query.shape
# + id="6-XlocOWl2iA" colab_type="code" colab={}
#https://www.kaggle.com/duttadebadri/image-classification/downloads/image-classification.zip/2
| Image_Registration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import pandas_profiling
import matplotlib.pyplot as plt
import scipy as stats
import matplotlib.ticker as ticker
import seaborn as sns
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_absolute_error,mean_squared_error,r2_score
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.datasets import load_boston
from sklearn import preprocessing
from sklearn.linear_model import LinearRegression, Ridge, Lasso
data=pd.read_csv('https://raw.githubusercontent.com/reddyprasade/Machine-Learning-Problems-DataSets/master/Regression/01%20Auto%20Insurance%20Total%20Claims%20Dataset/auto-insurance.csv',header=None)
data
data.columns=['number of claims','total payment for all the claims in thousands of Swedish Kronor']
data
data.head(2)
data.tail(2)
data.shape
data.info()
data.describe()
data.isna().sum()
sns.pairplot(data)
pandas_profiling.ProfileReport(data)
data.columns
x_data=data['number_of_claims']
x_data
y_data=data['total_payment_for_all_the_claims_in_thousands_of_Swedish_Kronor']
y_data
X_traine,X_test,Y_traine,Y_test,=train_test_split(x_data,y_data,test_size=0.20,random_state=5)
lr=LinearRegression()
lr.fit(X_traine.values.reshape(-1,1),Y_traine)
X_traine.shape
Y_traine.shape
X_test.shape
Y_test.shape
lr.coef_
lr.intercept_
train_score=lr.score(X_traine.values.reshape(-1,1),Y_traine)
train_score
test_score=lr.score(X_test.values.reshape(-1,1),Y_test)
test_score
rr=Ridge(alpha=0.01)
rr.fit(X_traine.values.reshape(-1,1),Y_traine)
rr.coef_
rr.intercept_
rr100=Ridge(alpha=100) #comparison with alpha value
rr100.fit(X_traine.values.reshape(-1,1),Y_traine)
rr100.coef_
rr100.intercept_
Ridge_train_score=rr.score(X_traine.values.reshape(-1,1),Y_traine)
Ridge_train_score
Ridge_test_score=rr.score(X_test.values.reshape(-1,1),Y_test)
Ridge_test_score
Ridge_train_score100=rr.score(X_traine.values.reshape(-1,1),Y_traine)
Ridge_train_score100
Ridge_test_score100=rr.score(X_test.values.reshape(-1,1),Y_test)
Ridge_test_score100
lo=Lasso(alpha=(0.01)**2)
lo.fit(X_traine.values.reshape(-1,1),Y_traine)
lo.coef_
lo.intercept_
lo100=Lasso(alpha=(100)**2)
lo100.fit(X_traine.values.reshape(-1,1),Y_traine)
lo100.coef_
lo100.intercept_
Lasso_train_score=lo.score(X_traine.values.reshape(-1,1),Y_traine)
Lasso_train_score
Lasso_test_score=lo.score(X_test.values.reshape(-1,1),Y_test)
Lasso_test_score
Lasso_train_score100=lo100.score(X_traine.values.reshape(-1,1),Y_traine)
Lasso_train_score100
Lasso_test_score100=lo100.score(X_test.values.reshape(-1,1),Y_test)
Lasso_test_score100
print("linear regression trine score",train_score)
print("linear regression test score",test_score)
print("linear regression trine score low_alpha",Ridge_train_score)
print("linear regression test score high_alpha",Ridge_test_score)
print("linear regression trine score low_alpha",Ridge_train_score100)
print("linear regression test score high_alpha",Ridge_test_score100)
print("linear regression trine score low_alpha",Lasso_train_score)
print("linear regression test score high_alpha",Lasso_test_score)
print("linear regression trine score low_alpha",Lasso_train_score100)
print("linear regression test score high_alpha",Lasso_test_score100)
# +
plt.figure(figsize=(16,9))
plt.plot(rr.coef_,alpha=0.7,linestyle='none',marker='*',markersize=15,color='red',
label=r'Ridge;$\alpha=0.01$',zorder=7)
#Zorder for ordering the markes
plt.plot(rr100.coef_,alpha=0.5,linestyle='none',marker='d',markersize=6,color='blue',
label=r'Ridge;$\alpha=100$',zorder=7)
plt.plot(lo.coef_,alpha=0.2,linestyle='none',marker='v',markersize=15,color='black',
label=r'Lasso;$\alpha=0.01$',zorder=7)
#Zorder for ordering the markes
plt.plot(lo100.coef_,alpha=0.9,linestyle='none',marker='8',markersize=6,color='yellow',
label=r'Lasso;$\alpha=100$',zorder=7)
#alpha here is for transparency
plt.plot(lr.coef_,alpha=0.4,marker='o',markersize=17,color='green',label='LinearRegression')
plt.xlabel('Coefficient Index',fontsize=16)
plt.ylabel('coefficient Magnitude',fontsize=16)
plt.legend(fontsize=13,loc=4)
plt.show()
# -
from sklearn.linear_model import LassoCV,RidgeCV,Lasso,Ridge,ElasticNet
Lasso_CV=LassoCV()
Ridge_CV=RidgeCV()
rr=Ridge()
Lo=Lasso()
ENET=ElasticNet()
Lasso_CV.fit(X_traine.values.reshape(-1,1),Y_traine)
Ridge_CV.fit(X_traine.values.reshape(-1,1),Y_traine)
rr.fit(X_traine.values.reshape(-1,1),Y_traine)
Lo.fit(X_traine.values.reshape(-1,1),Y_traine)
ENET.fit(X_traine.values.reshape(-1,1),Y_traine)
Predicate=pd.DataFrame({'Lasso_CV_pred':Lasso_CV.predict(X_test.values.reshape(-1,1)),
'Ridge_CV_pred':Ridge_CV.predict(X_test.values.reshape(-1,1)),
'rr':rr.predict(X_test.values.reshape(-1,1)),
'Lo':Lo.predict(X_test.values.reshape(-1,1)),'Actual_data':Y_test})
Predicate
#Recursive Feature Elimination:The Syntax
#import the class containing the feature selection method
from sklearn.feature_selection import RFE
#create an instance of the class
rfeMod=RFE(rr,n_features_to_select=7)
#fit the instance on the data and then predict the expected value
rfeMod=rfeMod.fit(X_traine,Y_traine)
Y_predict=rfeMod.predict(X_test)
rr
rr100
lo
lo100
| Optimization_techniques/Auto Insurance Total Claims Dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_excel("merge.xlsx")
df['일자'] = pd.to_datetime(df['일자'], format="%Y%m%d")
df = df.set_index('일자')
df.head()
# 60일 영업일 수익률
return_df = df.pct_change(60)
return_df.tail()
s = return_df.iloc[-1]
momentum_df = pd.DataFrame(s)
momentum_df.columns = ["모멘텀"]
momentum_df.head()
momentum_df["순위"] = momentum_df["모멘텀"].rank(ascending=False)
momentum_df.head(n=10)
momentum_df = momentum_df.sort_values(by='순위')
momentum_df.head()
momentum_df[:30].to_excel("momentum_list.xlsx")
| python-kiwoom-api/intermediate/section4/unit11/momentum.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Load and split the dataset
# - Load the train data and using all your knowledge of pandas try to explore the different statistical properties of the dataset.
# - Separate the features and target and then split the train data into train and validation set.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error, r2_score
import seaborn as sns
import warnings
warnings.filterwarnings("ignore")
# +
# Code starts here
train_data=pd.read_csv('train.csv')
test_data=pd.read_csv('test.csv')
test_data.head()
#np.mean(train['list_price'])
train=train_data[['ages','num_reviews','piece_count','play_star_rating','review_difficulty','star_rating','theme_name','val_star_rating','country','Id']]
validation= train_data['list_price']
train.head()
validation.head()
validation
# Code ends here.
# -
# ### Data Visualization
#
# - All the features including target variable are continuous.
# - Check out the best plots for plotting between continuous features and try making some inferences from these plots.
# +
# Code starts here
train.hist(column='star_rating',bins=10, figsize=(7,5), color='blue')
plt.show()
train.boxplot( 'num_reviews',showfliers=False, figsize=(10,5))
validation.hist('list_price',bins=10, figsize=(7,5), color='blue')
# Code ends here.
# -
# ### Feature Selection
# - Try selecting suitable threshold and accordingly drop the columns.
# +
# Code starts here
train.head()
test_data.head()
# Code ends here.
# -
# ### Model building
# +
# Code starts here
linearg = LinearRegression()
linearg.fit(train, validation)
pred= linearg.predict(test_data)
y_pred=pred.tolist()
y_pred
# Code ends here.
# -
# ### Residual check!
#
# - Check the distribution of the residual.
# +
# Code starts here
#rsqaured = r2_score(test_data, np.exp(pred))
# Code ends here.
# -
# ### Prediction on the test data and creating the sample submission file.
#
# - Load the test data and store the `Id` column in a separate variable.
# - Perform the same operations on the test data that you have performed on the train data.
# - Create the submission file as a `csv` file consisting of the `Id` column from the test data and your prediction as the second column.
# +
# Code starts here
id=test_data['Id']
id
my_dict={ 'Id': id, 'list_price': y_pred}
df=pd.DataFrame(my_dict)
df.set_index('Id')
df.to_csv('submission.csv')
df
# Code ends here.
| The-Lego-Collector's-Dilemma-Project/Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Customer Segmentation
# In this project, we will analyze a dataset containing data on various customers' annual spending
# amounts (reported in monetary units) of diverse product categories for internal structure. One goal of
# this project is to best describe the variation in the different types of customers that a wholesale
# distributor interacts with. Doing so would equip the distributor with insight into how to best structure
# their delivery service to meet the needs of each customer.
# ### Load in Data
#import libraries necessary for this project
import numpy as np
import pandas as pd
from IPython.display import display # Allows the use of display() for DataFrames
import seaborn
# Import supplementary visualizations code visuals.py
from vpython import *
# Pretty display for notebooks
# %matplotlib inline
import matplotlib.pyplot as plt
#Load the wholesale customers datase
data = pd.read_csv("Wholesale customers data.csv")
data.drop(['Region', 'Channel'], axis = 1, inplace = True)
data.head()
# ### Data Exploration
# Display a description of the dataset
display(data.describe())
# To get a better understanding of the customers and how their data will transform through the analysis,
# it would be best to select a few sample data points and explore them in more detail.
# Select three indices of your choice you wish to sample from the dataset
np.random.seed(40)
indices = np.random.randint(0,1,3).tolist()
# Create a DataFrame of the chosen samples
samples = pd.DataFrame(data.loc[indices], columns = data.keys()).reset_index(drop = True)
print("Chosen samples of wholesale customers dataset:")
display(samples)
data.apply(lambda col: col.drop_duplicates().reset_index(drop=True))
data = data.fillna(0)
#Find percentile of purchases for sample customers
from scipy.stats import percentileofscore
pct_samples = pd.DataFrame()
for item in samples:
pct_scores = []
for customer in samples.index:
pct_scores.append(round((percentileofscore(data[item], samples.loc[customer, item])),2))
pct_samples[item] = pct_scores
print("Percentile scores of purchases for sample customers: ")
display(pct_samples)
new_data = data.drop('Grocery', axis=1)
from sklearn.model_selection import train_test_split
# Split the data into training and testing sets using the given feature as the target
X_train, X_test, y_train, y_test = train_test_split(new_data,
data['Grocery'], test_size=0.25, random_state=42)
from sklearn.tree import DecisionTreeRegressor
# Create a decision tree regressor and fit it to the training set
regressor = DecisionTreeRegressor(random_state=42)
regressor.fit(X_train, y_train)
# Report the score of the prediction using the testing set
score = regressor.score(X_test, y_test)
print(score)
#Plot all categories vs Grocery purchases with trend line
fig = plt.figure(figsize=(10,10))
fig.subplots_adjust(hspace=0.5)
fig.subplots_adjust(wspace=0.5)
for i, item in enumerate(new_data):
z = np.polyfit(new_data[item], data['Grocery'], 1)
p = np.poly1d(z)
plt.subplot(3,2,i+1)
plt.scatter(x=new_data[item], y=data['Grocery'])
plt.plot(new_data[item], p(new_data[item]), "r-")
plt.xlabel(item + ' Units'), plt.ylabel('Grocery Units'),
plt.title(item + ' vs. Grocery Purchases')
# Find R^2 value (coefficient of determination) between other items and groceries
from scipy.stats import pearsonr
for category in new_data:
correlation_coef = pearsonr(new_data[category], data['Grocery'])[0]
print("Grocery and " + category + " R^2 value ={:.4f}".format(np.square(correlation_coef)))
#Produce a scatter matrix for each pair of features in the data
pd.plotting.scatter_matrix(data, alpha = 0.3, figsize = (16,12), diagonal = 'kde');
print('Grocery and Detergents_Paper R^2 score = {:.4f}'.format(np.square(pearsonr(data['Grocery'], data['Detergents_Paper'])[0])))
print('Grocery and Milk R^2 score = {:.4f}'.format(np.square(pearsonr(data['Grocery'], data['Milk'])[0])))
print( 'Detergents_Paper and Milk R^2 score = {:.4f}'.format(np.square(pearsonr(data['Detergents_Paper'], data['Milk'])[0])))
# Scale the data using the natural logarithm
log_data = np.log(data)
# Scale the sample data using the natural logarithm
log_samples = np.log(samples)
# Produce a scatter matrix for each pair of newly-transformed features
pd.plotting.scatter_matrix(log_data, alpha = 0.3, figsize = (16,12), diagonal ='kde');
#Display the log-transformed sample data
display(log_samples)
print('Grocery and Detergents_Paper R^2 score after transformation = {:.4f}'.format(np.square(pearsonr(log_data['Grocery'],
log_data['Detergents_Paper'])[0])))
print('Grocery and Milk R^2 score after transformation = {:.4f}'.format(np.
square(pearsonr(log_data['Grocery'], log_data['Milk'])[0])))
print( 'Detergents_Paper and Milk R^2 score after transformation = {:.4f}'.
format(np.square(pearsonr(log_data['Detergents_Paper'], log_data['Milk'])[0
])))
#For each feature find the data points with extreme high or low values
outlier_list = []
for feature in log_data.keys():
# Calculate Q1 (25th percentile of the data) for the given feature
Q1 = np.percentile(log_data[feature], 25)
# Calculate Q3 (75th percentile of the data) for the given feature
Q3 = np.percentile(log_data[feature], 75)
# Use the interquartile range to calculate an outlier step (1.5 times the interquartile range)
step = 1.5 * (Q3 - Q1)
# Display the outliers
print("Data points considered outliers for the feature '{}':".format(feature))
display(log_data[~((log_data[feature] >= Q1 - step) & (log_data[feature] <= Q3 + step))])
outlier_list.append(log_data[~((log_data[feature] >= Q1 - step) &
(log_data[feature] <= Q3 + step))].index.tolist())
# OPTIONAL: Select the indices for data points you wish to remove
outliers = []
# Remove the outliers, if any were specified
good_data = log_data.drop(log_data.index[outliers]).reset_index(drop = True)
#Flatten list of outliers and count occurences of each customer
outlier_list = [item for sublist in outlier_list for item in sublist]
# Add customer to outlier dictionary if they appear more than once
outlier_dict = {}
for i in outlier_list:
if outlier_list.count(i) > 1:
if i not in outlier_dict:
outlier_dict[i] = outlier_list.count(i)
# Find percentile scores of all outliers
outlier_pct = pd.DataFrame(columns=data.columns)
for customer_num in outlier_dict.keys():
pct_scores = []
for category in data:
pct_scores.append(round((percentileofscore(data[category], data.loc[
customer_num, category])),2))
outlier_pct.loc[customer_num] = pct_scores
print("Outlier number of appearances: ")
print(outlier_dict)
print("\nPercentile scores of outliers: ")
display(outlier_pct)
# OPTIONAL: Select the indices for data points you wish to remove
outliers = [154]
# Remove the outliers, if any were specified
good_data = log_data.drop(log_data.index[outliers]).reset_index(drop = True)
from sklearn.decomposition import PCA
# Apply PCA by fitting the good data with the same number of dimensions as features
pca = PCA(n_components=6).fit(good_data)
# TODO: Transform log_samples using the PCA fit above
pca_samples = pca.transform(log_samples)
# Generate PCA results plot
num_components = np.arange(1, 7)
cumulative_variance_explained = np.cumsum(pca.explained_variance_ratio_)
plt.plot(num_components, cumulative_variance_explained, '-o');
plt.xlabel('Number of PCs'); plt.ylabel('Cumulative Explained Variance');
plt.title('Cumulative Explained Variance vs. Number of PCs');
#Display sample log-data after having a PCA transformation applied
display(pd.DataFrame(np.round(pca_samples, 4), columns = pca_results.index.values))
print("Percentile scores of sample customers: ")
display(pct_samples)
#Apply PCA by fitting the good data with only two dimensions
pca = PCA(n_components=2).fit(good_data)
#Transform the good data using the PCA fit above
reduced_data = pca.transform(good_data)
# Transform log_samples using the PCA fit above
pca_samples = pca.transform(log_samples)
# Create a DataFrame for the reduced data
reduced_data = pd.DataFrame(reduced_data, columns = ['Dimension 1', 'Dimension 2'])
# Display sample log-data after applying PCA transformation in twodimensions
display(pd.DataFrame(np.round(pca_samples, 4), columns = ['Dimension 1', 'Dimension 2']))
# +
from mpl_toolkits.mplot3d import Axes3D
def discrete_cmap(N, base_cmap=None):
"""Create an N-bin discrete colormap from the specified input map
Source: https://gist.github.com/jakevdp/91077b0cae40f8f8244a"""
base = plt.cm.get_cmap(base_cmap)
color_list = base(np.linspace(0, 1, N))
cmap_name = base.name + str(N)
return plt.cm.get_cmap(base_cmap, N)
cmap = discrete_cmap(4, base_cmap = plt.cm.RdYlBu)
# +
# Plot each method
fig = plt.figure(figsize = (8, 8))
ax = fig.add_subplot(111, projection='3d')
p = ax.scatter(reduced_data['Dimension 1'], reduced_data['Dimension 1'],
cmap = cmap)
plt.title(f'{name.capitalize()}', size = 22)
fig.colorbar(p, aspect = 4, ticks = [1, 2, 3, 4])
# -
# ### Clusters (Gaussian Mixture)
from sklearn.mixture import GaussianMixture
from sklearn.metrics import silhouette_score
scores = []
for i in range(2, 11):
# TODO: Apply your clustering algorithm of choice to the reduced data
n_clusters = i
clusterer = GaussianMixture(n_components=n_clusters, random_state=42).fit(reduced_data)
# TODO: Predict the cluster for each data point
preds = clusterer.predict(reduced_data)
# TODO: Find the cluster centers
centers = clusterer.means_
# TODO: Predict the cluster for each transformed sample data point
sample_preds = clusterer.predict(pca_samples)
# TODO: Calculate the mean silhouette coefficient for the number of clusters chosen
score = silhouette_score(reduced_data, preds)
scores.append(score)
print('Clusters: {}; Silhouette Score = {:.4f}'.format(n_clusters, score))
plt.plot(np.arange(2,11), scores);
plt.xlabel('Num of Clusters'); plt.ylabel('Silhouette Score'); plt.title('Silhouette Score vs. Num of Clusters');
# Create clustering model with optimal number of components as measured bysilhouette score
clusterer = GaussianMixture(n_components=2, random_state=42)
# Fit to the processed data and make predictions
clusterer.fit(reduced_data)
preds = clusterer.predict(reduced_data)
reduced_data
sample_preds = clusterer.predict(pca_samples)
centers = clusterer.means_
# +
plt.scatter(reduced_data['Dimension 1'], reduced_data['Dimension 2'], c='red', s=50, cmap='viridis')
centers = clusterer.means_
plt.scatter(centers[:, 0], centers[:, 1], c='black', s=200, alpha=0.5);
# -
# Inverse transform the centers
log_centers = pca.inverse_transform(centers)
# Exponentiate the centers
true_centers = np.exp(log_centers)
# Display the true centers
segments = ['Segment {}'.format(i) for i in range(0,len(centers))]
true_centers = pd.DataFrame(np.round(true_centers), columns = data.keys())
true_centers.index = segments
print("Centers of clusters:")
display(true_centers)
centers_pct = pd.DataFrame(columns=data.columns)
for center in true_centers.index.tolist():
pct_scores = []
for category in data:
pct_scores.append(round((percentileofscore(data[category],true_centers.loc[center, category])),2))
centers_pct.loc[center] = pct_scores
print("Percentile scores of cluster centers: ")
display(centers_pct)
#Display the predictions
for i, pred in enumerate(sample_preds):
print("Sample point", i, "predicted to be in Cluster", pred)
| Customer Segmentation Part1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <h1>Theoretical Foundations, Continued</h1>
# <h1>Chapter 4: Classical Statistical Inference (4.1-4.4)</h1>
# <hr/>
#
# <h2>4.2 Maximum Likelihood Estimation (MLE)</h2>
#
# <h3>4.2.1-4.2.2 The Likelihood Function</h3>
#
# What's the idea? A set of data is a sample drawn from some distribution. As such, each datum has a probabilty; if we make the assumption that these probabilities are independent, we arrive fairly intuitively at the measure of likelihood for the sample:
#
# $$L \equiv p(\{x_i\}|M(\vec{\theta})) = {\displaystyle \prod_{i=1}^{n}} p(x_i|M(\vec{\theta})) \tag{4.1}$$
#
# where $M$ is the model (the distribution the data is drawn from), and $\vec{\theta}$ are the parameters the model takes.
#
# Note:
# <list>
# <li>$L$ is not a PDF as it is not normalized</li>
# <li>In fact, commonly $L\ll 1$, which leads to the use of Log-likelihood, $ln(L)$
# <li>$L$ can be considered both as a function of the model/distribution parameters, with fixed $\{x_i\}$, the case when trying to maximize it; or as a function of $x$, with fixed model parameters, when calculating the likelihood of some value.
# </list>
#
# "All" that needs now be done is take the derivative $L$ (or $ln(L)$), set to zero, and solve for the parameters giving the maximum. Once this is done, confidence estimates for the parameters can be determined, either analytically or (more likely) numerically. Finally, hypothesis tests/goodness of fit must be determined.
#
# <h3>4.2.4 Properties of Maximum Likelihood Estimators</h3>
#
# Assuming that the model $M$ truly is the correct class of distribution from which data ${x_i}$ are drawn, MLE's have several optimality properties.
# <list>
# <li>They are consistent, converging to the true value as data points increase</li>
# <li>They are asymptotically normal: the distibution of the parameter estimate approaches a normal distribution about the MLE as data points increase; the spread($\sigma$) of this distribution can be used as a confidence interval about the estimate.</li>
# <li>They achieve the minimum possible variance given the data at hand</li>
# </list>
#
# <h3>4.2.3 The Homoscedastic Gaussian Likelihood</h3>
#
# Given $N$ measurements, $\{x_i\}$, with a known, identical, normal error $\sigma$ the likelihood function becomes:
#
# $$L \equiv p(\{x_i\}|\mu,\sigma) = {\displaystyle \prod_{i=1}^{N}} \frac{1}{\sqrt{2\pi\sigma}} exp\left(\frac{-(x_i - \mu)^2}{2\sigma^2}\right) \tag{4.2}$$
#
# with only one parameter, $\vec{\theta}=(\mu)$, or simply $\theta=\mu$.
#
# Using the log-likelihood here is doubly useful; besides rendering tiny numbers more numerical/computationally managable, here, analytically, it turns multiplications into additions, and those additions are logs of exponentials, so that:
#
# $$ln(L)={\displaystyle \sum_{i=1}^{N}} ln\left(\frac{1}{\sqrt{2\pi\sigma}} exp\left(\frac{-(x_i - \mu)^2}{2\sigma^2}\right)\right) = {\displaystyle \sum_{i=1}^{N}}\left( ln\left(\frac{1}{\sqrt{2\pi\sigma}}\right) + \frac{-(x_i - \mu)^2}{2\sigma^2}\right) \tag{D1}$$
#
# or
#
# $$ln(L(\mu))=constant-{\displaystyle \sum_{i=1}^{N}} \frac{(x_i - \mu)^2}{2\sigma^2} \tag{4.4}$$
#
# Setting the derivative (by the only parameter, $\mu$) equal to zero gives:
#
# $$\frac{d~ln(L(\mu))}{d~\mu}=-{\displaystyle \sum_{i=1}^{N}} \frac{-2(x_i - \mu)}{2\sigma^2}=0 \implies {\displaystyle \sum_{i=1}^{N}} (x_i - \mu) = 0 \tag{D2}$$
#
# or
#
# $$\mu_{mle} = \frac{1}{N}{\displaystyle \sum_{i=1}^{N}}x_i \tag{4.5}$$
#
# As expected, as it should be.
#
# <h3>4.2.6 The Heteroscedastic Gaussian Likelihood</h3>
#
# Rather than the case of equation (4.4), we now have different errors per datum, $\sigma_i$:
#
# $$ln(L(\mu))=constant-{\displaystyle \sum_{i=1}^{N}} \frac{(x_i - \mu)^2}{2\sigma_i^2} \tag{4.8}$$
#
# from which, with $w_i = \sigma_i^{-2}$, and following (D2) above:
#
# $$\mu_{mle} = \frac{\displaystyle \sum_{i=1}^{N}w_i x_i}{\displaystyle \sum_{i=1}^{N}w_i} \tag{4.5}$$
#
# aka, simply the weighted mean.
#
# <h3>4.2.5 MLE Confidence Intervals</h3>
#
# Given a maximum likelihood estimate of e.g. $\mu_{mle}$ as above, what is its uncertainty?
#
# $$\sigma_{jk} = \left( - \frac{d^2 ~ln(L(\theta))}{d~\theta_j d\theta_k} \right)^{-1/2} \tag{4.6}$$
#
# for $\theta=\vec{\theta}_{mle}$. For why this is, the text refers the reader to the Wasserman textbooks (after a brief description.) Without the why: the diagnonal elements $\sigma_{ii}$ correspond to marginal errors for $\theta_i$, while the $\sigma_{ij}$ with $i \neq j$ indicate correlation of errors for $\theta_i$ and $\theta_j$.
#
# For the Guassian cases,
#
# $$\sigma_{\mu} = \left(- \frac{d^2 ~ln(L(\mu))}{d~\mu^2} \right)^{-1/2} = \left({\displaystyle \sum_{i=1}^{N}\frac{1}{\sigma_i^2}}\right)^{-1/2} \tag{4.7/4.10}$$
#
# <h3>4.2.7 MLE in the Case of Truncated and Censored Data</h3>
#
# A variable to be measured, x, has some range of possible values; due to e.g. the measuring aparatus used, the range of values is not sampled uniformly. So: $S(x) \neq c$, where $S$ is the PDF for sampling x. In particular, for some values of $x$, $S(x)=0$. When this last is true for some $x<x_{min}$ and/or $x > x_{max}$, the data set is said to be truncated. (Censored data is where data has been removed for some reason.)
#
# If we take the Gaussian case, and the simple truncation $$S(x) = \left\{\begin{array}{ll}c & x_{min} \leq x \leq x_{max} \\ 0 & otherwise \\ \end{array} \right. $$ with $c$ a constant to make $S(x)$ sum to 1.
#
# The probability distribution for $x$ needs to be renormalized to account for this truncation: $p(x)$ needs to be scaled to become $C~p(x)$ such that $1 = {\displaystyle \int_{-\infty}^{\infty}} C~p(x)$ For this example case this is simple:
#
# $$C = C(\mu, \sigma, x_{min}, x_{max}) = \frac{1}{P(x_{max}|\mu, \sigma) - P(x_{min}|\mu, \sigma)} \tag{4.12}$$
#
# leading to a log-likelihood of:
#
# $$ln(L(\mu))=constant -{\displaystyle \sum_{i=1}^{N}} \frac{(x_i - \mu)^2}{2\sigma^2} + N~ln(C(\mu, \sigma, x_{min}, x_{max})) \tag{4.13}$$
#
# <h3>4.2.8 Beyond the Likelihood: Other Cost Functions and Robustness</h3>
#
# MLE represents the most common choice of cost functions. The expectation value of the cost function is called "risk." Minimizing risk is a way to obtain best-fit parameters.
#
# The mean integrated square error (MISE),
#
# $$MISE = \displaystyle \int_{-\infty}^{\infty} [f(x) - h(x)]^2 d~x \tag{4.14}$$
#
# is often used. MISE is based on Mean Squared Error (MSE), aka the $L_2$ norm. A cost function minimizing the absoluate deviation is called the $L_1$ norm. Many cost functions, with different properties; a particularly useful example of a property is robustness to outliers.
#
# In chapters 6-10 cost functions will be important for various methods; this is particularly true when formalizing the likelihood function is difficult, because an optimal solultion can still eb found by minimizing the risk.
#
# <h2>4.3 The Goodness of Fit and Model Selection</h2>
#
# MLE estimates the best-fit parameters and gives us their uncertainties, but does not tell us how good a fit the model/parameters are. What if a Gaussian model was choosen by e.g. the truth was Laplacian? And what if a polynomial is being fit: a higher order polynomial will always fit data better than a lower order polynomial, but is the higher order polynomial a better fit to the underlying process (e.g., are we just fitting noise or actually fitting additional complexity in the underlying distribution/process?)
#
# <h3>4.3.1 The Goodness of Fit for a Model</h3>
#
# In the Gaussian case, we have (4.4):
#
# $$ln(L(\mu))=constant-{\displaystyle \sum_{i=1}^{N}} \frac{(x_i - \mu)^2}{2\sigma^2} \tag{4.4}$$
#
# which may be re-written with $z_i=(x_i - \mu)/\sigma$ as
#
# $$ln(L(\mu))=constant-{\displaystyle \sum_{i=1}^{N}} z_i^2 = constant - \frac{1}{2}\chi^2 \tag{4.15}$$
#
# and hence the distibution of $ln(L)$ can be determined from the $\chi^2$ distribution with $N-k$ degrees of freedom, with $k$ model parameters. With an expectation value of $N-k$, for a "good fit" we should have $\chi_{dof}^2=\frac{\chi^2}{N-k} \approx 1$ (As in chapter 3, the warning here is that $\chi$ is very sensitive to outliers.)
#
# The probability that a certain ML value $L_{mle}$ arose by chance can only be evaluated by $\chi^2$ when the likelihood is Gaussian; otherwise $L_{mle}$ is still a measure of how well a model fits the data. Asssuming the same $k$, models can be ranked by their likelihood. But the $L_{mle}$ value(s) alone do not indicated in an <i>absolute</i> sense how well the model(s) fit the data; to know that requires knowing the distribution of $L_{mle}$, as given by $\chi^2$ for a Gaussian likelihood.
#
# <h3>4.3.2 Model Comparison</h3>
#
# The best way to compare models is cross-validation, but this topic is covered in detail in later chapters.
#
# The Aikake Information Criterion (AIC) is a simple method for comparing models that (attempts to) accounts for model complexity in addition to $L_{mle}$ when comparing models. AIC is defined as:
#
# $$AIC \equiv -2~ln(L_{mle}) + 2k + \frac{2k(k+1)}{N-k-1} \tag{4.17}$$
#
# or
#
# $$AIC = \frac{2~k~N}{N-(k+1)} - 2~ln(L_{mle})$$
#
# Out of multiple possible models, the one with the smallest AIC is the best.
# %pylab inline
def AIC(LNL, N):
return -2 * LNL + 2.0 * N / (N - 2.0)
def Gauss(X,M,S):
c = np.power((np.sqrt(2*np.pi)*S),-1)
e = -np.power(X-M,2)/(2*np.power(S,2))
return c*np.exp(e)
# +
#We will be needing the Gaussian error function
import scipy.special as spec
#Generating points from a gaussian distribution and truncating
D = np.random.normal(5,3,500)
D = D[D>=2]
D = D[D<=8]
#Creating triangle function likelihood terms
#Assuming we guessed X0 to be the value used in generation
t = np.zeros(len(D))
for i in range(len(D)):
if D[i] < 2 or D[i] > 8:
pass
elif D[i] < 5:
t[i] = np.log((D[i]-2)/9.0)
else:
t[i] = np.log((8-D[i])/9.0)
Taic = AIC(np.sum(t), len(D))
#Calculating (truncated) gaussian likelihood
#Assuming we guessed X0 to be the value used in generation
const = np.power(spec.erf(3.0/(np.sqrt(2)*3)),-1)
#(3.48) Simplified as abs(xmin-mu) = abs(xmax-mu)
Gaic = AIC(len(D) * np.log(1/(np.sqrt(2*np.pi)*3)) -
np.sum(np.power(D-5,2)/(2*np.power(3,2))) +
len(D) * np.log(const), len(D))
#Plotting data against probability densities
x1 = np.linspace(2,8,100)
y1 = const * Gauss(x1,5,3)
x2 = np.array([2,3,4,5,6,7,8])
y2 = np.array([0,1,2,3,2,1,0])/9.0
plt.figure(figsize = (8,6))
plt.hist(D, bins = 20, normed = True, histtype='stepfilled', alpha = 0.5)
plt.plot(x1,y1, linewidth = 2, label = "Gauss AIC = {:.2f}".format(Gaic))
plt.plot(x2,y2, linewidth = 2, label = "Tri AIC = {:.2f}".format(Taic))
plt.legend()
plt.show()
# -
# <h2>4.4 ML Applied to Gaussian Mixtures: The Expectation Maximization Algorithm</h2>
#
# A special case of a complex likelihood which can still be maximized simply (and treated analytically) is a mixture of Gaussians.
#
# <h3>4.4.1 Gaussian Mixture Model</h3>
#
# For a model made up of $M$ Gaussians the likelihood of a given datum $x_i$ is:
#
# $$p(x_i|\vec{\theta}) = {\displaystyle \sum_{i=1}^{M} \alpha_j ~ \mathcal{N}(\mu_j, \sigma_j)} \tag{4.18}$$
#
# where, because we require each point to be drawn from a true pdf, the normalization constants $\alpha_j$ must sum to 1. The log-likelihood is then:
#
# $$ln(L)={\displaystyle \sum_{i=1}^{N} ln \left( {\displaystyle \sum_{i=1}^{M} \alpha_j ~ \mathcal{N}(\mu_j, \sigma_j)} \right)} \tag{4.20}$$
#
# with $k=3M-1$ parameters.
#
# <h3>Class Labels and Hidden Variables</h3>
#
# A variety of more advanced methods are available for maximizing $ln(L)$, but a fast and and realatively easy method is "hidden variables". Each of the $M$ Guassians above are interpreted as a class such that any individual $x_i$ was generated by one and only one Gaussian. The hidden variable is $j$, identifying which class each $x_i$ belongs to. If each point's class is known, the problem resolves to $M$ separate MLE problems with Guassian models, as developed so far. The fraction of points in each class would be an estimator for the corresponding normalization factor, $\alpha_j$. When the class labels are known but the underlying distribution is not Gaussian, the "naive Bayesian classfier" ($\S$ 9.3.2) can be used.
#
# Continuing with the Gaussian case here, using Bayes' rule we find the probability of a given class for a given $x_i$:
#
# $$p(j|x_i)=\frac{\alpha_j ~ \mathcal{N}(\mu_j,\sigma_j)}{\displaystyle \sum_{j=1}^{M} \alpha_j ~ \mathcal{N}(\mu_j, \sigma_j)} \tag{4.21}$$
#
# or
#
# $$p(j|x_i) = \frac{\alpha_j ~ p(x_i|\mu_j,\sigma_j)}{\displaystyle \sum_{j=1}^{M} \alpha_j~p(x_i|\mu_j, \sigma_j)} = \frac{p(j) ~ p(x_i|\mu_j,\sigma_j)}{p(x_i)}
# \tag{D3}$$
#
# How to use (4.21) and (4.20) to come up with a way to handle this?
#
# <h3>4.4.3 The Basics of the Expectation Maximization Algorithm</h3>
#
# Replacing $\mathcal{N}(\mu_j, \sigma_j)$ with the general $p_j(x_i|\vec{\theta})$ in (4.20) and taking the partial derivative with respect to $\theta_j$, then rearranging gives:
#
# $$\frac{\partial~ln(L)}{\partial~\theta_j} = {\displaystyle \sum_{i=1}^{N} \left( \frac{\alpha_j~p_j(x_i|\vec{\theta})}{\displaystyle \sum_{j=1}^{M} \alpha_j~p(x_i|\vec{\theta})} \right)} \left( \frac{1}{p_j(x_i|\vec{\theta})} \frac{\partial~p_j(x_i|\vec{\theta})}{\partial~\theta_j} \right) \tag{4.24}$$
#
# where the first part is just (4.21/D3). For the EM algorithm we assume this is fixed during each iteration; this whole term is then replaced with $w_{ij}$. The second part is just a partial of the $ln(p_j(x_i|\vec{\theta}))$ and, when Gaussian as in our work so far, gives:
#
# $$\frac{\partial~ln(L)}{\partial~\theta_j} = -{\displaystyle \sum_{i=1}^{N}} w_{ij} \frac{\partial}{\partial~\theta_j} \left( ln(\sigma_j) + \frac{(x_i - \mu_j)^2}{2~\sigma_j^2} \right)$$
#
# and leads to the estimators for $\mu_j$, $\sigma_j$, and $\alpha_j$:
#
# $$\mu_j = \frac{\displaystyle \sum_{i=1}^{N} w_{ij} x_i}{\displaystyle \sum_{i=1}^{N} w_{ij}} \tag{4.26}$$
#
# $$\sigma_j^2 = \frac{\displaystyle \sum_{i=1}^{N} w_{ij}(x_i-\mu_j)^2}{\displaystyle \sum_{i=1}^{N} w_{ij}} \tag{4.27}$$
#
# and (somewhat circularly)
#
# $$\alpha_j = \frac{1}{N}{\displaystyle \sum_{i=1}^{N} w_{ij}} \tag{4.28}$$
#
# The EM algorithm starts with a guess for $w_{ij}$, then the maximization step (M-step) of evaluating (4.26-4.28), then a expectation step (E-step) of updating $w_{ij}$ based on the M-step outputs. The M-step and E-step are run iteratively until convergence, iteration limit, etc.
#
# Similar to overfitting with e.g. a high-degree polynomial, setting M too high will e.g. split data that should be classed together. Choosing the appropriate M is a case of model selection, and AIC (or BIC, later) should be applied.
#
# So far this has been for homoscedastic errors. Handling heteroscedastic errors can be done by replacing the $\sigma_j$ with $(sigma_j^2 + e_i^2)^(1/2)$ where $sigma_j$ is now the width of each class $j$ and $e_i$ the measurement error for each value. Now the M-step involves an explicit equation to update $\mu_j$ and an implicit equation for $\sigma_j$ that requires a numerical solution:
#
# $$\mu_j = \frac{\displaystyle \sum_{i=1}^{N} \frac{w_{ij}}{\sigma_j^2 + e_i^2} x_i}{\displaystyle \sum_{i=1}^{N} \frac{w_{ij}}{\sigma_j^2 + e_i^2}} \tag{4.30}$$
#
# and
#
# $${\displaystyle \sum_{i=1}^{N} \frac{w_{ij}}{\sigma_j^2 + e_i^2}} = {\displaystyle \sum_{i=1}^{N} \frac{w_{ij}}{(\sigma_j^2 + e_i^2)^2}(x_i - \mu_j)^2} \tag{4.31}$$
#
# Further discussion of this and similar problems (involving mixutre models) is dicussed in $\S$5.6.1 and in chapter 6.
# +
# Author: <NAME>
# License: BSD
# The figure produced by this code is published in the textbook
# "Statistics, Data Mining, and Machine Learning in Astronomy" (2013)
# For more information, see http://astroML.github.com
# To report a bug or issue, use the following forum:
# https://groups.google.com/forum/#!forum/astroml-general
from matplotlib import pyplot as plt
import numpy as np
from sklearn.mixture import GMM
#----------------------------------------------------------------------
# This function adjusts matplotlib settings for a uniform feel in the textbook.
# Note that with usetex=True, fonts are rendered with LaTeX. This may
# result in an error if LaTeX is not installed on your system. In that case,
# you can set usetex to False.
from astroML.plotting import setup_text_plots
setup_text_plots(fontsize=10, usetex=False)
#------------------------------------------------------------
# Set up the dataset.
# We'll use scikit-learn's Gaussian Mixture Model to sample
# data from a mixture of Gaussians. The usual way of using
# this involves fitting the mixture to data: we'll see that
# below. Here we'll set the internal means, covariances,
# and weights by-hand.
np.random.seed(1)
gmm = GMM(3, n_iter=1)
gmm.means_ = np.array([[-1], [0], [3]])
gmm.covars_ = np.array([[1.5], [1], [0.5]]) ** 2
gmm.weights_ = np.array([0.3, 0.5, 0.2])
X = gmm.sample(1000)
#------------------------------------------------------------
# Learn the best-fit GMM models
# Here we'll use GMM in the standard way: the fit() method
# uses an Expectation-Maximization approach to find the best
# mixture of Gaussians for the data
# fit models with 1-10 components
N = np.arange(1, 11)
models = [None for i in range(len(N))]
for i in range(len(N)):
models[i] = GMM(N[i]).fit(X)
# compute the AIC and the BIC
AIC = [m.aic(X) for m in models]
BIC = [m.bic(X) for m in models]
#------------------------------------------------------------
# Plot the results
# We'll use three panels:
# 1) data + best-fit mixture
# 2) AIC and BIC vs number of components
# 3) probability that a point came from each component
fig = plt.figure(figsize=(15, 5.1))
fig.subplots_adjust(left=0.12, right=0.97,
bottom=0.21, top=0.9, wspace=0.5)
# plot 1: data + best-fit mixture
ax = fig.add_subplot(131)
M_best = models[np.argmin(AIC)]
x = np.linspace(-6, 6, 1000)
logprob, responsibilities = M_best.score_samples(x) #GMM.eval is now GMM.score_samples
pdf = np.exp(logprob)
pdf_individual = responsibilities * pdf[:, np.newaxis]
ax.hist(X, 30, normed=True, histtype='stepfilled', alpha=0.4)
ax.plot(x, pdf, '-k')
ax.plot(x, pdf_individual, '--k')
ax.text(0.04, 0.96, "Best-fit Mixture",
ha='left', va='top', transform=ax.transAxes)
ax.set_xlabel('$x$')
ax.set_ylabel('$p(x)$')
# plot 2: AIC and BIC
ax = fig.add_subplot(132)
ax.plot(N, AIC, '-k', label='AIC')
ax.plot(N, BIC, '--k', label='BIC')
ax.set_xlabel('n. components')
ax.set_ylabel('information criterion')
ax.legend(loc=2)
# plot 3: posterior probabilities for each component
ax = fig.add_subplot(133)
p = M_best.predict_proba(x)
p = p[:, (1, 0, 2)] # rearrange order so the plot looks better
p = p.cumsum(1).T
ax.fill_between(x, 0, p[0], color='gray', alpha=0.3)
ax.fill_between(x, p[0], p[1], color='gray', alpha=0.5)
ax.fill_between(x, p[1], 1, color='gray', alpha=0.7)
ax.set_xlim(-6, 6)
ax.set_ylim(0, 1)
ax.set_xlabel('$x$')
ax.set_ylabel(r'$p({\rm class}|x)$')
ax.text(-5, 0.3, 'class 1', rotation='vertical')
ax.text(0, 0.5, 'class 2', rotation='vertical')
ax.text(3, 0.3, 'class 3', rotation='vertical')
plt.show()
# +
#Seting up random addition of 3 Gaussians
xgen = np.linspace(-10, 20, 30)
mgen = np.random.choice(np.arange(11), 3)
sgen = np.random.choice(np.linspace(0.5,3.5,50), 3)
agen = np.random.choice(np.linspace(5,25,50), 3)
ggen = np.zeros(len(xgen))
for i in range(3):
ggen = ggen + agen[i] * Gauss(xgen,mgen[i], sgen[i])
#Setting number of gaussians to fit
J = 3
#Setting starting guess values for mu, sigma, alpha
avg = np.sum(xgen*ggen) / np.sum(ggen)
var = np.sum(ggen*np.power(xgen-avg,2)) / (np.sum(ggen))
std = np.sqrt(var)
s = std * np.ones(J)
a = np.ones(J) / J
m = np.random.choice(xgen, J)
w = np.zeros((J,len(xgen)))
sig = np.zeros(J, dtype = bool)
mu = np.zeros(J, dtype = bool)
alph = np.zeros(J, dtype = bool)
#Start iterating over function
for k in range(50):
for j in range(J):
for i in range(len(xgen)):
#Creating omega matrix
w[j][i] = (a[j] * Gauss(xgen[i], m[j], s[j]) /
np.sum([a[l] * Gauss(xgen[i], m[l], s[l]) for l in range(J)]))
#Solving for new mu, sigma, alpha values & test against tolerance
m1 = np.sum(w[j]*xgen*ggen) / np.sum(w[j]*ggen)
if np.abs(m1 - m[j]) < 0.01:
mu[j] = True
m[j] = m1
s1 = np.sqrt(np.sum(w[j]*ggen*np.power(xgen-m[j],2)) /
np.sum(w[j]*ggen))
if np.abs(s1 - s[j]) < 0.01:
sig[j] = True
s[j] = s1
a1 = np.sum(w[j]*ggen) / np.sum(ggen)
if np.abs(a1 - a[j]) < 0.01:
alph[j] = True
a[j] = a1
if mu.all() and sig.all() and alph.all():
print('Convergence to tolerance after {} iterations:'.format(k))
break
#Plotting comparison to data
plt.figure(figsize = (14,8))
plt.plot(xgen,ggen, color = 'gray', linewidth = 6, label = 'data')
scale = np.trapz(ggen,xgen)
xout = np.linspace(-10,20,100)
g = np.sum([a[j] * Gauss(xout,m[j],s[j]) for j in range(J)], axis = 0)
plt.plot(xout,g * scale, color = 'black', linewidth = 4, label = 'EM Fit')
for i in range(J):
plt.plot(xout,a[i]*Gauss(xout,m[i],s[i])*scale, linewidth = 2, label = 'EM Gauss {}'.format(i+1))
plt.legend()
plt.show()
# +
#Using SDSS data via bossdata tool
import bossdata
#Getting flux data for object
finder = bossdata.path.Finder()
mirror = bossdata.remote.Manager()
Fiber = bossdata.spec.SpecFile(mirror.get(finder.get_spec_path(3953,55322,4)))
data = Fiber.get_valid_data()
wlen,flux,dflux = data['wavelength'][:],data['flux'][:],data['dflux'][:]
model = Fiber.hdulist[1]['model'][:]
offset = flux - model
#Choosing small subsample around multiple emission lines
peak = np.argmax(flux)
testy = np.copy(offset[peak-15:peak+20])
testx = np.copy(wlen[peak-15:peak+20])
#Setting number of gaussians to fit
J = 5
#Setting starting guess values for mu, sigma, alpha
avg = np.sum(testx*testy) / np.sum(testy)
var = np.sum(testy*np.power(testx-avg,2)) / (np.sum(testy)-1)
std = np.sqrt(var)
s = std * np.ones(J)
a = np.ones(J) / J
m = np.random.choice(testx, J)
w = np.zeros((J,len(testx)))
sig = np.zeros(J, dtype = bool)
mu = np.zeros(J, dtype = bool)
alph = np.zeros(J, dtype = bool)
#Start iterating over function
for k in range(50):
for j in range(J):
for i in range(len(testx)):
#Creating omega matrix
w[j][i] = (a[j] * Gauss(testx[i], m[j], s[j]) /
np.sum([a[l] * Gauss(testx[i], m[l], s[l]) for l in range(J)]))
#Solving for new mu, sigma, alpha values & test against tolerance
m1 = np.sum(w[j]*testx*testy) / np.sum(w[j]*testy)
if np.abs(m1 - m[j]) < 0.1:
mu[j] = True
else:
mu[j] = False
m[j] = m1
s1 = np.sqrt(np.sum(w[j]*testy*np.power(testx-m[j],2)) /
np.sum(w[j]*testy))
if np.abs(s1 - s[j]) < 0.05:
sig[j] = True
else:
sig[j] = False
s[j] = s1
a1 = np.sum(w[j]*testy) / np.sum(testy)
if np.abs(a1 - a[j]) < 0.1:
alph[j] = True
else:
alph[j] = False
a[j] = a1
if mu.all() and sig.all() and alph.all():
print('Convergence to tolerance after {} iterations:'.format(k))
break
#Ploting comparison to data
plt.figure(figsize = (14,8))
plt.plot(testx,testy, label = 'SDSS data', color = 'grey', linewidth = 5)
#plt.fill_between(testx,testy-dflux[peak-15:peak+20], testy + dflux[peak-15:peak+20])
g = np.sum([a[j]*Gauss(testx,m[j],s[j]) for j in range(J)], axis = 0)
plt.plot(testx,g * np.trapz(testy,testx), label = 'EM Fit', color = 'black', linewidth = 2)
plt.legend()
plt.show()
| Chapter4/Chapter 4 (4.1-4.4).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visual Analytics Coursework (Student Number - 2056224)
# +
#importing libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# -
data = pd.read_csv('Occupation by Sex.csv')
data
# # Data Pre-processing
# Following are the first steps towards data pre-processing:
#
# 1. The data is for year 2011, so there is no use of keeping the respective column with date. REMOVING COLUMN "date".
#
# 2. Check if columns "geography" and "geography code" are same. If yes, remove "geography code".
#
# 3. The words "measures: value" are same in every column. Removing just the respective names to make the column names look better and easily understandable.
#
# 4. The data has 'total' as Rural Urban, so there is no use of keeping the respective column. REMOVING COLUMN "Rural Urban".
data['date'].unique()
data['Rural Urban'].unique()
#Removing the date and rural urban column
data.drop('date', axis=1, inplace=True)
data.drop('Rural Urban', axis=1, inplace=True)
#Checing the columns geography and geography code are same or not?
if data['geography'].equals(data['geography code']) == True:
print("The columns have same values.")
else:
print("The columns does not have same values.")
#The values are same, so dropping the column geography code.
data.drop('geography code', axis=1, inplace=True)
# +
#Check if every column name ends with "measures: value".
name_lst = data.columns
for i in range(1, len(name_lst)):
if name_lst[i].endswith('measures: Value') == False:
print("{} does not ends with 'measures: value'. ".format(name_lst[i]))
# -
#Since the column names ends with the respective words, removing the respective from column names.
data.columns = data.columns.str.replace('; measures: Value', '')
data.columns
# The second step towards data pre-processing would be creating sub columns for a set of same category of column names.
def col_split(x : pd.Series):
y = x['variable'].split(';')
x['Sex'] = y[0]
x['Occupation'] = y[1]
return x
new_data = data.melt(id_vars=['geography']).apply(col_split, axis = 1)
#data.melt(id_vars=['geography']).apply(col_split, axis = 1)
#Dropping the column 'variable' as it is of no use now.
new_data.drop('variable', axis=1, inplace=True)
# +
#Removing the common name 'Sex: ' from 'Sex' column as it is of no use.
new_data['Sex'] = new_data['Sex'].str.replace('Sex: ', '')
#Removing the common name 'Occupation: ' from 'Occupation' column as it is of no use.
new_data['Occupation'] = new_data['Occupation'].str.replace('Occupation: ', '')
# -
#bringing the 'value' column at the end of dataframe
new_data['Counts'] = new_data['value']
new_data.drop('value', axis=1, inplace=True)
new_data['Sex'].unique()
new_data['Occupation'].unique()
new_data[['area code', 'area name']] = new_data['geography'].str.split('-', 1, expand=True)
new_data.head(10)
new_data.tail(10)
new_data.to_csv('Occupation by Sex clean.csv', index=False)
| Python Files/Occupation by Sex.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Podium Dataflow Dependency Tree
#
# ## Introduction
#
# This Jupyter Notebook demonstrates how it is possible to generate QDC dataflow / source / entity dependency tree from the QDC metadata database (in this case PostgreSQL) using Python.
#
# The starting point for the example output (below) is the name of a dataflow "prod_stg_compsych_member_t", this starting point should be the dataflow that you are intrested in determining it's predecessor dataflows.
#
# The process will look at the LOADERs in the starting point dataflow and look back up the dependency tree to determine the source of each of the LOADERs.
#
# These sources can be a QDC Source / Entity or the output from a prior dataflow.
#
# Where the source is the output from a prior dataflow, the process recurses on itself until:
#
# * There are only QDC Source / Entities remaining or
# * The source dataflow is in a stop list of dataflow names provided to the process.
#
# The result of the process is a [NetworkX](https://networkx.github.io) Graph where the nodes are dataflows, sources and entities, the edges reflect the relationships.
#
# The final Graph is converted to a [DOT](https://en.wikipedia.org/wiki/DOT_(graph_description_language)) file and rendered to the png shown here using the dot.exe command installed with [Graphviz](https://www.graphviz.org/).
#
# ```bash
# dot.exe -Tpng -o prod_stg_compsych_member_t.png prod_stg_compsych_member_t.dot
# ```
#
# ## Sample output
#
# 
# ## Python Notes
#
# Access to the PostgreSQL database is through Python SQLAlchemy, the coding style chosen here for SQLAlchemy was chosen for clarity and simplicity.
#
# I am sure that there are many improvements and optimizations that could be made.
#
# The Python version used here was 3.6.5 installed from the Anaconda distribution which means that all the packages imported were pre installed.
from sqlalchemy import create_engine
from sqlalchemy import MetaData , Table, Column, Integer, Numeric, String, DateTime, ForeignKey, Text
from sqlalchemy import select, desc
from sqlalchemy import and_, or_, not_
from sqlalchemy import text
from sqlalchemy.sql import func
import networkx as nx
import json
import datetime as dt
import os
import yaml
import sys
import pydot
from networkx.drawing.nx_pydot import write_dot
# ## QDC Database Connection and Styles
#
# The QDC repository database connection details are specified in a yaml format and parsed by the `get_podium_cfg()` function below.
#
# The input to the function is a stream, this can be a string or an open file handle.
#
# This version of the config file contains style attributes that are used in the resulting `dot.exe` generated output file.
#
# The expected yaml stream should be:
#
# ```yaml
# ---
# pd_connect:
# dr:
# user: 'user'
# pwd: '<PASSWORD>'
# db: 'podium_md'
# host: 'host.host.com'
# port: 5436
# dev:
# pd_connect:
# user: 'user'
# pwd: '<PASSWORD>'
# db: 'podium_md'
# host: 'host.host'
# port: 5436
#
# styles:
# meta:
# fontname: 'arial'
# fontsize: 10
# meta_edge:
# fontname: 'arial'
# fontsize: 10
# node:
# fontname: 'arial'
# fontsize: 12
# bgcolor: '"#ECF8B9"'
# source:
# shape: 'house'
# style: 'filled'
# fillcolor: '"#D4E4F7"'
# entity_source:
# shape: 'ellipse'
# style: 'filled'
# fillcolor: '"#3A84D9"'
# entity_target:
# shape: 'ellipse'
# style: 'filled'
# fillcolor: '"#DAC344"'
# bundle:
# shape: 'box'
# style: '"filled,rounded"'
# fillcolor: '"#E9446A"'
# edge:
# loader:
# color: '"#66B032"'
# store:
# color: '"#347B98"'
# source:
# color: '"#092834"'
#
# ```
# ## Helper Functions
#
# Most of these functions are helper functions in accessing the QDC metadata database, the function where the actual work is done is the `get_bundle()` function.
def get_podium_cfg(yaml_stream):
"""Loads the Podium connection parameters from the input stream"""
try:
pd_cfg = yaml.load(yaml_stream)
except:
raise "Unexpected error reading yaml stream"
return pd_cfg
def connect(user, password, db, host, port):
'''Returns a connection and a metadata object'''
url = f'postgresql+psycopg2://{user}:{password}@{host}:{port}/{db}'
# The return value of create_engine() is our connection object
con = create_engine(url, client_encoding='utf8')
# We then bind the connection to MetaData()
meta = MetaData(bind=con)
return con, meta
def get_source(con, meta, source_nid):
"""Retrieve Podium Source row by nid
Parameters
==========
con : SQLAlchemy connection
meta : SQL Alchemy Meta Object
source_nid : Integer source nid
Returns
=======
Uses ResultProxy first() to retrieve one row and close the result set.
"""
assert isinstance(source_nid, int)
pd_source = Table('podium_core.pd_source', meta)
s = pd_source.select()
s = s.where(pd_source.c.nid == source_nid)
r = con.execute(s)
assert r.rowcount == 1
return r.first()
def get_source_byname(con, meta, source_name, source_type = 'PODIUM_INTERNAL'):
"""Returns the Podium source row for a named Source"""
lc_source_name = source_name.lower()
pd_source = Table('podium_core.pd_source', meta)
s = select([pd_source.c.nid,
pd_source.c.sname])
s = s.where(and_(func.lower(pd_source.c.sname) == lc_source_name,
pd_source.c.source_type == source_type))
rp = con.execute(s)
return rp.first()
def get_entity(con, meta, entity_nid):
"""Fetches the entity record by the entity nid, returns 1 row at most"""
assert isinstance(entity_nid, int)
pd_entity = Table('podium_core.pd_entity', meta)
s = select([pd_entity.c.sname,
pd_entity.c.source_nid])
s = s.where(pd_entity.c.nid == entity_nid)
rp = con.execute(s)
return rp.first()
def get_entity_byname(con, meta, source_nid, entity_name):
"""Fetches the entity record by entity name, returns 1 row at most"""
assert isinstance(source_nid, int)
lc_entity_name = entity_name.lower()
pd_entity = Table('podium_core.pd_entity', meta)
s = select([pd_entity.c.nid,
pd_entity.c.sname])
s = s.where(and_(func.lower(pd_entity.c.sname) == lc_entity_name,
pd_entity.c.source_nid == source_nid))
rp = con.execute(s)
return rp.first()
def get_entity_store(con, meta, entity_id):
"""Fetches the dataflows (if any) that STORE the passed entity id"""
assert isinstance(entity_id, int)
pd_prep_package = Table('podium_core.pd_prep_package', meta)
pd_bundle = Table('podium_core.pd_bundle', meta)
s = select([pd_bundle.c.nid, pd_bundle.c.sname])
s = s.select_from(pd_prep_package.join(pd_bundle, pd_prep_package.c.bundle_nid == pd_bundle.c.nid))
s = s.where(and_(pd_prep_package.c.entity_id == entity_id,
pd_prep_package.c.package_type == 'STORE'))
rp = con.execute(s, eid=entity_id).fetchall()
return rp
def get_bundle_id(con, meta, sname):
"""Get the bundle id of the passed Prepare Workflow name.
This is a case insensitive match and can only return a single row
or None
"""
pd_bundle = Table('podium_core.pd_bundle', meta)
lc_sname = sname.lower()
s = pd_bundle.select()
s = s.where(func.lower(pd_bundle.c.sname) == lc_sname)
rp = con.execute(s)
r = rp.first()
return r
def get_bundle_gui_state(con, meta, nid):
"""Get the bundle gui state record datflow nid.
"""
pd_bundle_gui_state = Table('podium_core.pd_bundle_gui_state', meta)
gui_cols = [pd_bundle_gui_state.c.nid,
pd_bundle_gui_state.c.created_ttz,
pd_bundle_gui_state.c.modified_ttz,
pd_bundle_gui_state.c.version,
pd_bundle_gui_state.c.modifiedby,
pd_bundle_gui_state.c.createdby]
s = select(gui_cols)
s = s.where(pd_bundle_gui_state.c.nid == nid)
rp = con.execute(s)
return rp.first()
def get_bundle_last_execution(con, meta, bundle_nid, count=10):
"""Get the last count execution details of the specified bundle.
"""
pd_prepare_execution_workorder = Table('podium_core.pd_prepare_execution_workorder', meta)
wo_cols = [pd_prepare_execution_workorder.c.nid,
pd_prepare_execution_workorder.c.record_count,
pd_prepare_execution_workorder.c.start_time,
pd_prepare_execution_workorder.c.end_time]
e = select(wo_cols)
e = e.where(and_(pd_prepare_execution_workorder.c.bundle_nid == bundle_nid,
pd_prepare_execution_workorder.c.end_time.isnot(None),
pd_prepare_execution_workorder.c.workorder_status == "FINISHED"))
e = e.order_by(desc(pd_prepare_execution_workorder.c.end_time))
e = e.limit(count)
rp = con.execute(e)
r = rp.fetchall()
rp.close()
return r
def get_entity_last_load(con, meta, source_nid, entity_name, n=1):
"""Get the last execution details of the specified bundle.
"""
pd_source = Table('podium_core.pd_source', meta)
pd_entity = Table('podium_core.pd_entity', meta)
pd_workorder = Table('podium_core.pd_workorder', meta)
#print entity_name
parent_source = get_source(con, meta, source_nid)
src = pd_source.select()
src = src.where(pd_source.c.sname == parent_source.sname)
srp = con.execute(src)
orig_source_id = None
for r in srp:
print(f'Source: {r.sname}, Source Type: {r.source_type}, nid: {r.nid}')
if r.source_type != 'PODIUM_INTERNAL':
orig_source_id = r.nid
break
print(f'orig_source_id: {orig_source_id}')
if orig_source_id is None:
return None
ety = pd_entity.select()
ety = ety.where(and_(pd_entity.c.source_nid == orig_source_id,
pd_entity.c.sname == entity_name))
rp = con.execute(ety)
orig_entity = rp.first()
if orig_entity is not None:
orig_entity_nid = orig_entity.nid
wo = select([pd_workorder.c.nid,
pd_workorder.c.start_time,
pd_workorder.c.end_time,
pd_workorder.c.record_count,
pd_workorder.c.good_count,
pd_workorder.c.bad_count,
pd_workorder.c.ugly_count])
wo = wo.where(and_(pd_workorder.c.entity_nid == orig_entity_nid,
pd_workorder.c.workorder_status == 'FINISHED'))
wo = wo.order_by(desc(pd_workorder.c.end_time))
wo = wo.limit(n)
rp = con.execute(wo)
r = rp.first()
else:
r = None
return r
def get_package_nodes(con, meta, bundle_nid):
pd_prep_package = Table('podium_core.pd_prep_package', meta)
s = pd_prep_package.select()
s = s.where(pd_prep_package.c.bundle_nid == bundle_nid)
rp = con.execute(s)
r = rp.fetchall()
rp.close()
return r
# ## podium_core Tables Used
#
# 
prep_tables = ('pd_bundle',
'pd_bundle_gui_state',
'pd_prep_package',
'pd_entity',
'pd_source',
'pd_prepare_execution_workorder',
'pd_workorder'
)
# ## Establish connection to podium_core and fetch used tables metadata
#
# Enter the correct yaml file name (or stream) in the call to the `get_podium_cfg()` function.
# +
cfg = get_podium_cfg(open('pd_cfg.yaml', 'r'))
con_cfg = cfg['pd_connect']['dev']
# -
con, meta = connect(con_cfg['user'], con_cfg['pwd'], con_cfg['db'], con_cfg['host'], con_cfg['port'])
meta.reflect(bind=con, schema='podium_core', only=prep_tables)
# ## Main get_bundle() Function
#
# This function is called with a single dataflow name (sname) and for each LOADER in the dataflow recurses backwards looking for dataflows that STORE that entity.
#
# If a dataflow is found the function self recurses.
#
# To prevent being caught in circular references, as each dataflow is visited it's name is stored in the wf_list, this list is checked each time the function is entered.
#
# The stop list "STOPPERS[]" is a list of dataflow names that will also stop the recursion process.
#
# The Networkx DiGraph is built up throughout the process adding nodes for Sources, Entities and Dataflows as they are first encountered.
#
# The node ids are the nid's from the related `pd_bundle` (Dataflow), `pd_source` (Source) and `pd_entity` (Entity) tables prefixes with the characters `b_`, `s_` and `e_` respectively.
#
# Edges are created between nodes to show the node relationships.
def get_bundle(con, meta, sname: str, world_graph, wf_list, styles: dict, stop_wf = []):
"""Build bundle dependency digraph"""
# Check if dataflow is in stop list
if (sname.lower() in stop_wf):
print(f'Dataflow {sname} is in stop list\n')
return
source_styles = styles['source']
entity_styles = styles['entity_source']
target_styles = styles['entity_target']
bundle_styles = styles['bundle']
edge_styles = styles['edge']
print(bundle_styles)
bundle = get_bundle_id(con, meta, sname)
#import pdb; pdb.set_trace()
print(f'Current dataflow {bundle.sname} ({bundle.nid})')
if bundle:
bundle_nid = bundle.nid
bundle_description = bundle.description
bundle_sname = bundle.sname
bundle_gui_state = get_bundle_gui_state(con, meta, bundle.bundle_gui_state_nid)
if bundle_gui_state:
bundle_mod_dt = bundle_gui_state.modified_ttz
bundle_mod_by = bundle_gui_state.modifiedby
bundle_version = bundle_gui_state.version
else:
bundle_mod_dt = 'Unknown'
bundle_mod_by = 'Unknown'
bundle_version = 'Unknown'
# To-do - check if output file for version already exists
# if so then bypass
bundle_exec = get_bundle_last_execution(con, meta, bundle_nid)
if bundle_exec:
exec_stats = []
for i, r in enumerate(bundle_exec):
if i == 0:
last_record_count = r.record_count
last_start_time = r.start_time
last_end_time = r.end_time
exec_stats.append(({'start_time': str(r.start_time),
'end_time': str(r.end_time),
'records': r.record_count}))
else:
last_record_count = 0
last_start_time = ''
last_end_time = ''
print(f'\t{bundle_nid}, {bundle_description}, {bundle_sname} records {last_record_count}')
print(f'\tModified by: {bundle_mod_by}, Modified Date: {bundle_mod_dt}, Version: {bundle_version}')
print(f'\tLast Start: {last_start_time}, Last End: {last_end_time}')
else:
print(f'Package: {sname}, not found')
return None
# add bundle to "world" graph
bundle_node_key = f'b_{bundle_nid}'
W.add_node(bundle_node_key,
nid=bundle_nid,
# sname=bundle_sname,
label=bundle_sname,
n_type='bundle',
**bundle_styles)
# Add LOADER / STORE nodes
p = get_package_nodes(con, meta, bundle_nid)
# Add LOADER and STORE nodes to graph
for n in p:
id = n.nid
n_type = n.package_type
if n_type in ('LOADER','STORE'):
entity_id = n.entity_id
entity_node_key = f'e_{entity_id}'
if n_type == 'LOADER':
l = get_entity_store(con, meta, entity_id)
if len(l) == 0:
print(f'No STORE found for {entity_id}')
else:
for i, ldr in enumerate(l):
print(f'{entity_id} ({i}) STORE by {ldr.sname}')
if not (ldr.sname.lower() in wf_list):
wf_list.append(ldr.sname.lower())
get_bundle(con, meta, ldr.sname, world_graph, wf_list, styles, stop_wf)
if (not W.has_node(entity_node_key)):
entity = get_entity(con, meta, entity_id)
source_id = entity.source_nid
source = get_source(con, meta, source_id)
if n_type == 'LOADER':
W.add_node(entity_node_key,
n_type='entity',
nid=entity_id,
snid=source_id,
#sname=entity.sname,
label=entity.sname,
**entity_styles)
if n_type == 'STORE':
W.add_node(entity_node_key,
n_type='entity',
nid=entity_id,
snid=source_id,
#sname=entity.sname,
label=entity.sname,
**target_styles)
source_node_key = f's_{source_id}'
if (not W.has_node(source_node_key)):
W.add_node(source_node_key,
n_type='source',
nid=source_id,
#sname=source.sname,
label=source.sname,
**source_styles)
W.add_edge(source_node_key,
entity_node_key,
**edge_styles['source'])
else:
source_nid = W.node[entity_node_key]['snid']
source_node_key = f's_{source_nid}'
print(f"Graph already has entity {entity_id}, {W.node[source_node_key]['label']}.{W.node[entity_node_key]['label']}")
if (n_type == 'STORE'):
W.add_edge(bundle_node_key, entity_node_key, **edge_styles['store'])
elif (n_type == 'LOADER'):
W.add_edge(entity_node_key, bundle_node_key, **edge_styles['loader'])
else:
print(f'ERROR {bundle_node_key}, {source_node_key}, {entity_node_key}')
def subg(g, sg, n_type):
"""Create a record type subgraph of the passsed node type"""
label_list = [g.node[n]['label'] for n in g.nodes if g.node[n]['n_type'] == n_type]
label_list.sort(key=lambda x: x.lower())
# Start subgraph and header record of number of lines
print(f'subgraph {sg} {{')
print(f'r_{sg} [shape=record,label="{{')
print(f'{len(label_list)} {n_type}')
for i, label in enumerate(label_list):
print(f'| {label}')
# Close subgraph
print('}"];}')
def write_record_dot(g, output_file=None):
print("digraph g {")
subg(g, "s1", "bundle")
subg(g, "s2", "source")
subg(g, "s3", "entity")
print('}')
# ## Main
# +
# final dataflow name and "stop list"
SEED = "prod_stg_compsych_member_t" # "prod_stg_source_member_t" #
#STOPPERS = ('je70_chess_history_cdc_v',
# 'je70_chess_init_member_history_v',
# 'je70_chess_copy_member_history_v')
# The stop list should be zero or more dataflow names that are stoppers in the
# recursion. If a dataflow name in te STOPPER list is hit then the recursion will
# stop at that point.
STOPPERS = ()
# -
if __name__ == "__main__":
W = nx.DiGraph()
node_styles = cfg['styles']['node']
edge_styles = cfg['styles']['meta_edge']
meta_styles = cfg['styles']['meta']
now = dt.datetime.now()
meta_dict = {'shape': 'record', 'label': f'{{{now.strftime("%Y-%m-%d %H:%M")} | {{<f0> SEED | <f1> {SEED}}}}}', **meta_styles}
W.add_node('node', **node_styles)
W.add_node('meta', **meta_dict)
W.add_node('source', **cfg['styles']['source'])
W.add_node('entity', **cfg['styles']['entity_source'])
W.add_node('dataflow', **cfg['styles']['bundle'])
W.add_node('target', **cfg['styles']['entity_target'])
W.add_edges_from((('entity','source', {'label': 'belongs_to', **edge_styles}),
('entity','dataflow', {'label': 'LOADER', **edge_styles}),
('dataflow','target', {'label': 'STORE', **edge_styles})))
wf_list = []
wf_list.append(SEED.lower())
print(f'Processing starting point, dataflow {SEED}')
get_bundle(con, meta, SEED, W, wf_list, cfg['styles'], STOPPERS)
print(f'{len(W.nodes)} nodes added to DiGraph')
# Write output dot file
write_dot(W, f'{SEED}.dot')
# Write output GraphML file
with open(f"{SEED}.graphml", "wb") as ofile:
nx.write_graphml(W, ofile)
print("Finished")
| Prepare_Dependency_Gen.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="fu8NIB7u9TXq" outputId="d0880091-0d5e-4274-b7ed-8ae24f047257"
dia = 15
mes = 10
ano = 2015
print(dia, mes, ano, sep='/')
# + id="aNsdIWprflry"
# adivinhacao.py
import random
def jogar():
print("*********************************")
print("Bem vindo ao jogo de Adivinhação!")
print("*********************************")
numero_secreto = random.randrange(1, 100)
total_de_tentativas = 0
pontos = 1000
print("Qual o nível de dificuldade?")
print("(1) Fácil (2) Médio (3) Difícil")
nivel = int(input("Defina o nível: "))
if (nivel == 1):
total_de_tentativas = 20
elif (nivel == 2):
total_de_tentativas = 10
else:
total_de_tentativas = 5
for rodada in range(1, total_de_tentativas + 1):
print("Tentativa {} de {}".format(rodada, total_de_tentativas))
chute_str = input("Digite um número entre 1 e 100: ")
print("Você digitou: ", chute_str)
chute = int(chute_str)
if (chute < 1 or chute > 100):
print("Você deve digitar um número entre 1 e 100!")
continue
acertou = numero_secreto == chute
maior = chute > numero_secreto
menor = chute < numero_secreto
if (acertou):
print("Você acertou e fez {} pontos!".format(pontos))
break
else:
if (maior):
print("Você errou! O seu chute foi maior que o número secreto.")
elif (menor):
print("Você errou! O seu chute foi menor que o número secreto.")
pontos_perdidos = abs(numero_secreto - chute)
pontos = pontos - pontos_perdidos
print("Fim do jogo")
if (__name__ == "__main__"):
jogar()
## Se executar diretamente ele roda o programa com o nome, que no caso é o próprio arquivo
# + colab={"base_uri": "https://localhost:8080/"} id="--GM6AO-Qvlf" outputId="095dc59b-433a-4352-a72a-94c46149e945"
import random
sorteado = random.randrange(0,4)
print(sorteado)
if sorteado == 1:
print( "Paulo")
elif sorteado == 2:
print("Juliana")
else:
print("Tamires")
# + id="yHNtU7MZhrqD"
# forca.py
def jogar():
print("*********************************")
print("***Bem vindo ao jogo da Forca!***")
print("*********************************")
print("Fim do jogo")
if(__name__ == "__main__"):
jogar()
# + id="DKWedLQ_fo8R"
# jogos.py
import forca
import adivinhacao
def escolhe_jogo():
print("*********************************")
print("*******Escolha o seu jogo!*******")
print("*********************************")
print("(1) Forca (2) Adivinhação")
jogo = int(input("Qual jogo? "))
if (jogo == 1):
print("Jogando forca")
forca.jogar()
elif (jogo == 2):
print("Jogando adivinhação")
adivinhacao.jogar()
if (__name__ == "__main__"):
escolhe_jogo()
| Python_basics_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Interaction with other libraries
# ## Keras
#
# - It's a very romantic notion to think that we can come up with the best features
# to model our world. That notion has now been dispelled.
# - Most *object detection/labeling/segmentation/classification* tasks now have
# neural network equivalent algorithms that perform on-par with or better than
# hand-crafted methods.
# - One library that gives Python users particularly easy access to deep learning is Keras: https://github.com/fchollet/keras/tree/master/examples (it works with both Theano and TensorFlow).
# - **At SciPy2017:** "Fully Convolutional Networks for Image Segmentation", <NAME>, SciPy2017 (Friday 2:30pm)
# - Particularly interesting, because such networks can be applied to images of any size
# - ... and because Daniil is a scikit-image contributor ;)
# ### Configurations
#
# From http://www.asimovinstitute.org/neural-network-zoo/:
#
# <img src="neuralnetworks.png" style="width: 80%"/>
#
# E.g., see how to fine tune a model on top of InceptionV3:
#
# <img src="inception_v3_architecture.png"/>
#
# - https://keras.io/applications/#fine-tune-inceptionv3-on-a-new-set-of-classes
#
#
# - https://github.com/fchollet/keras/tree/master/examples
# - https://keras.io/scikit-learn-api/
#
#
# - In the Keras docs, you may read about `image_data_format`. By default, this is `channels-last`, which is
# compatible with scikit-image's storage of `(row, cols, ch)`.
# +
import numpy as np
from keras.models import Sequential
from keras.layers import Dense, Dropout
import matplotlib.pyplot as plt
# %matplotlib inline
## Generate dummy data
#X_train = np.random.random((1000, 2))
#y_train = np.random.randint(2, size=(1000, 1))
#X_test = np.random.random((100, 2))
#y_test = np.random.randint(2, size=(100, 1))
## Generate dummy data with some structure
from sklearn import datasets
from sklearn.model_selection import train_test_split
X, y = datasets.make_classification(n_features=2, n_samples=2000, n_redundant=0, n_informative=1,
n_clusters_per_class=1, random_state=42)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.1, random_state=42)
model = Sequential()
model.add(Dense(64, input_dim=2, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(64, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy',
optimizer='rmsprop',
metrics=['accuracy'])
model.fit(X_train, y_train,
epochs=20,
batch_size=128)
score = model.evaluate(X_test, y_test, batch_size=128)
print('\n\nAccuracy:', score[1]);
# -
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier()
rf.fit(X_train, y_train)
rf.score(X_test, y_test)
# +
f, (ax0, ax1, ax2) = plt.subplots(1, 3, figsize=(15, 5))
mask = (y_train == 0)
ax0.plot(X_train[mask, 0], X_train[mask, 1], 'b.')
ax0.plot(X_train[~mask, 0], X_train[~mask, 1], 'r.')
ax0.set_title('True Labels')
y_nn = model.predict_classes(X_test).flatten()
mask = (y_nn == 0)
ax1.plot(X_test[mask, 0], X_test[mask, 1], 'b.')
ax1.plot(X_test[~mask, 0], X_test[~mask, 1], 'r.')
ax1.set_title('Labels by neural net')
y_rf = rf.predict(X_test)
mask = (y_rf == 0)
ax2.plot(X_test[mask, 0], X_test[mask, 1], 'b.')
ax2.plot(X_test[~mask, 0], X_test[~mask, 1], 'r.');
ax2.set_title('Labels by random forest')
# -
from keras.applications.inception_v3 import InceptionV3, preprocess_input, decode_predictions
net = InceptionV3()
# +
from skimage import transform
def inception_predict(image):
# Rescale image to 299x299, as required by InceptionV3
image_prep = transform.resize(image, (299, 299, 3), mode='reflect')
# Scale image values to [-1, 1], as required by InceptionV3
image_prep = (img_as_float(image_prep) - 0.5) * 2
predictions = decode_predictions(
net.predict(image_prep[None, ...])
)
plt.imshow(image, cmap='gray')
for pred in predictions[0]:
(n, klass, prob) = pred
print(f'{klass:>15} ({prob:.3f})')
# -
from skimage import data, img_as_float
inception_predict(data.chelsea())
inception_predict(data.camera())
inception_predict(data.coffee())
# You can fine-tune Inception to classify your own classes, as described at
#
# https://keras.io/applications/#fine-tune-inceptionv3-on-a-new-set-of-classes
# ## SciPy: LowLevelCallable
#
# https://ilovesymposia.com/2017/03/12/scipys-new-lowlevelcallable-is-a-game-changer/
# +
import numpy as np
image = np.random.random((512, 512))
footprint = np.array([[0, 1, 0],
[1, 1, 1],
[0, 1, 0]], dtype=bool)
# -
from scipy import ndimage as ndi
# %timeit ndi.grey_erosion(image, footprint=footprint)
# %timeit ndi.generic_filter(image, np.min, footprint=footprint)
f'Slowdown is {825 / 2.85} times'
# %load_ext Cython
# + magic_args="--name=test9" language="cython"
#
# from libc.stdint cimport intptr_t
# from numpy.math cimport INFINITY
#
# cdef api int erosion_kernel(double* input_arr_1d, intptr_t filter_size,
# double* return_value, void* user_data):
#
# cdef:
# double[:] input_arr
# ssize_t i
#
# return_value[0] = INFINITY
#
# for i in range(filter_size):
# if input_arr_1d[i] < return_value[0]:
# return_value[0] = input_arr_1d[i]
#
# return 1
# +
from scipy import LowLevelCallable, ndimage
import sys
def erosion_fast(image, footprint):
out = ndimage.generic_filter(
image,
LowLevelCallable.from_cython(sys.modules['test9'], name='erosion_kernel'),
footprint=footprint
)
return out
# -
np.sum(
np.abs(
erosion_fast(image, footprint=footprint)
- ndi.generic_filter(image, np.min, footprint=footprint)
)
)
# %timeit erosion_fast(image, footprint=footprint)
# !pip install numba
# +
# Taken from <NAME>'s blog post:
# https://ilovesymposia.com/2017/03/12/scipys-new-lowlevelcallable-is-a-game-changer/
import numba
from numba import cfunc, carray
from numba.types import intc, CPointer, float64, intp, voidptr
from scipy import LowLevelCallable
def jit_filter_function(filter_function):
jitted_function = numba.jit(filter_function, nopython=True)
@cfunc(intc(CPointer(float64), intp, CPointer(float64), voidptr))
def wrapped(values_ptr, len_values, result, data):
values = carray(values_ptr, (len_values,), dtype=float64)
result[0] = jitted_function(values)
return 1
return LowLevelCallable(wrapped.ctypes)
# -
@jit_filter_function
def fmin(values):
result = np.inf
for v in values:
if v < result:
result = v
return result
# %timeit ndi.generic_filter(image, fmin, footprint=footprint)
# ## Parallel and batch processing
# [Joblib](https://pythonhosted.org/joblib/) (developed by scikit-learn) is used for:
#
#
# 1. transparent disk-caching of the output values and lazy re-evaluation (memoize pattern)
# 2. easy simple parallel computing
# 3. logging and tracing of the execution
# +
from sklearn.externals import joblib
from joblib import Memory
mem = Memory(cachedir='/tmp/joblib')
# +
from skimage import segmentation
@mem.cache
def cached_slic(image):
return segmentation.slic(image)
# -
from skimage import io
large_image = io.imread('../images/Bells-Beach.jpg')
# %time segmentation.slic(large_image)
# %time cached_slic(large_image)
# %time cached_slic(large_image)
# [Dask](https://dask.pydata.org) is a parallel computing library. It has two components:
#
# - Dynamic task scheduling optimized for computation. This is similar to Airflow, Luigi, Celery, or Make, but optimized for interactive computational workloads.
# - “Big Data” collections like parallel arrays, dataframes, and lists that extend common interfaces like NumPy, Pandas, or Python iterators to larger-than-memory or distributed environments. These parallel collections run on top of the dynamic task schedulers.
# - See <NAME>'s [blogpost](http://matthewrocklin.com/blog/work/2017/01/17/dask-images) for a more detailed example
| lectures/not_yet_booked/other_libraries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Segmenting and Clustering Neighborhoods in Toronto
#
# This is Peer-graded Assignment for Course Applied Data Science Capstone, Week 3
#
#
# It Contains three parts coresponding the three submit, just click the link below:
#
# - [Scrape neighborhoods in Toronto](#0)<br>
# - [Fetching Location data of each neighborhood](#2)<br>
# - [Neighbourhoods Clustering Analysis](#5)<br>
import pandas as pd
import numpy as np
import requests
# ## Scrape neighborhoods in Toronto <a id="0"></a>
# **Step(1)** We get all possible tables in the Wiki page via pandas function **read_html**:
url = 'https://en.wikipedia.org/wiki/List_of_postal_codes_of_Canada:_M'
res = requests.get(url)
dfs = pd.read_html(url)
# Let's have a general idea of what we got:
for idx, df in enumerate(dfs):
print('DataFrame[{}]:{}'.format(idx, df.shape))
# It's easy to guess that **ONLY** the first dataframe is what we need, which has 180 rows and 3 columns.
#
# Let's verify our thought by reviewing the first 5 rows:
dfs[0].head()
# Great!
#
# Let's rename the first column name to 'PostalCode' and save it into a new variable then we are done our first step.
nb_toronto = dfs[0].rename({'Postal code':'PostalCode'}, axis='columns')
nb_toronto.head()
# Now lets move to the data clearning procedures, as described below.
#
# **Step (2)** Ignore cells with a borough that is **Not assigned.**
nb_toronto.shape
nb_toronto[nb_toronto['Borough']=='Not assigned'].shape
nb_toronto = nb_toronto[nb_toronto['Borough']!='Not assigned']
nb_toronto.shape
nb_toronto.head()
# **Step (3)** Check duplications on postal code
nb_toronto['PostalCode'].unique().shape
# Since the unique number is same with the total row number, it's proved that there's no dupoication on column Postal Code.
#
# **Stpe (4)** replace '/' with ',' in neighbourhoods combination
#
nb_toronto.loc[:,'Neighborhood'] = nb_toronto.apply(lambda x: x['Neighborhood'].replace(' / ',','), axis=1)
nb_toronto.head()
# **Step (5)** copy borough to neighborhood if neighborhood is missing
#
nb_toronto[ nb_toronto['Neighborhood'] == 'Not assigned' ]
# Looks we do not have any rows with **Not assigned** neighbourhood.
#
# To be more safe, also check **None** value for the column:
nb_toronto[ nb_toronto['Neighborhood'] == None ]
nb_toronto.shape
# We are good for this part.
#
# **This is the end of the submition of part 1.**
#
# ---
# ## Fetching Location data of each neighborhood<a id="2"></a>
# Since the Geocoder package can be very unreliable, we use the provided csv file as our data source of the geographical coordinates of each postal code.
#
# **Step (1)** Read the data from the given URL.
url = 'http://cocl.us/Geospatial_data'
geo_df = pd.read_csv(url)
print(geo_df.shape)
geo_df.head()
geo_df.dtypes
# To keep consistance, we removed the space in column name 'Postal Code'.
geo_df.rename({'Postal Code':'PostalCode'}, axis=1, inplace=True)
geo_df.head()
# **Step (2)** Merge it into the neibourhood data frame **nb_toronto** which we already populated in previous part.
nb_toronto_geo = pd.merge(nb_toronto, geo_df, on = ['PostalCode'])
print(nb_toronto_geo.shape)
nb_toronto_geo.head()
#
# **This is the end of the submition of part 2.**
#
# ***
#
# ## Neighbourhoods Clustering Analysis<a id="5"></a>
#
# We performed the Clustering Analysis with following steps:
# - Step (1) General Idea for all Toronto neighbourhood
# - Step (2) Narrow down to Downtown Toronto for further analysis
# - Step (3) Define Foursqure Credentials and Version and explore the neighborhood in Downtown Toronto
# - Step (4) Prepare venue category for Clustering
# - Step (5) Clustering the neighborhood according venue categories count
# - Step (6) Visualize the resulting on map
#
# +
import matplotlib.cm as cm
import matplotlib.colors as colors
from sklearn.cluster import KMeans
import folium # map rendering library
# -
# **Step (1)** Let's first get a general idea about those neibourhoods in Toronto on map, just list all of them there.
# +
lat, lng = geo_df[['Latitude','Longitude']].max() + geo_df[['Latitude','Longitude']].min()
lat, lng = lat /2, lng /2
lat, lng
# +
# create map of New York using latitude and longitude values
map_toronto = folium.Map(location=[lat, lng], zoom_start=11)
# add markers to map
for idx, r in nb_toronto_geo.iterrows():
lat, lng, bor, nb = r['Latitude'], r['Longitude'],r['Borough'], r['Neighborhood']
label = '{}, {}'.format(nb, bor)
label = folium.Popup(label, parse_html=True)
folium.CircleMarker(
[lat, lng],
radius=5,
popup=label,
color='blue',
fill=True,
fill_color='#3186cc',
fill_opacity=0.7,
parse_html=False).add_to(map_toronto)
map_toronto
# -
# **Step (2)** Let's focus on the borough **Downtown Toronto** only to demostrate the analysis.
#
# First we slice the orinal data frame nb_toronto_geo to nb_york, we omit the geo from the name for all data frame will have geo information from now on.
#
dt_toronto = nb_toronto_geo[nb_toronto_geo['Borough']=='Downtown Toronto']
print(dt_toronto.shape)
dt_toronto.head()
# Now lets show it on map
# +
## first we find out the center of the map
lat, lng = dt_toronto[['Latitude','Longitude']].max() + dt_toronto[['Latitude','Longitude']].min()
lat, lng = lat /2, lng /2
lat, lng
# +
# create map of New York using latitude and longitude values
map_dt_toronto = folium.Map(location=[lat, lng], zoom_start=13)
# add markers to map
for idx, r in dt_toronto.iterrows():
lat, lng, bor, nb = r['Latitude'], r['Longitude'],r['Borough'], r['Neighborhood']
label = '{}, {}'.format(nb, bor)
label = folium.Popup(label, parse_html=True)
folium.CircleMarker(
[lat, lng],
radius=5,
popup=label,
color='blue',
fill=True,
fill_color='#3186cc',
fill_opacity=0.7,
parse_html=False).add_to(map_dt_toronto)
map_dt_toronto
# -
# **Step (3)** Define Foursqure Credentials and Version and explore the neighborhood in **Downtown Toronto**
#
CLIENT_ID = 'Y5FK5TTSXY24B0DDCUJBGCWCL<KEY>'
CLIENT_SECRET = '<KEY>'
VERSION = '20180605'
# We borrow the function **getNearbyVenues** from the course lab, but only keep the catogry for further analysis.
#
#
def getNearbyVenues(df, radius = 700, LIMIT = 70):
venues_list=[]
for idx, row in df.iterrows():
name, lat, lng = row['Neighborhood'], row['Latitude'], row['Longitude']
# create the API request URL
url = 'https://api.foursquare.com/v2/venues/explore?&client_id={}&client_secret={}&v={}&ll={},{}&radius={}&limit={}'.format(
CLIENT_ID,
CLIENT_SECRET,
VERSION,
lat,
lng,
radius,
LIMIT)
# make the GET request
results = requests.get(url).json()["response"]['groups'][0]['items']
# return only relevant information for each nearby venue
venues_list.append([(
name,
lat,
lng,
v['venue']['location']['lat'],
v['venue']['location']['lng'],
v['venue']['categories'][0]['name']) for v in results])
nearby_venues = pd.DataFrame([item for venue_list in venues_list for item in venue_list])
nearby_venues.columns = ['Neighborhood Name',
'Neighborhood Latitude',
'Neighborhood Longitude',
'Venue Latitude',
'Venue Longitude',
'Venue Category']
return(nearby_venues)
venues_dt_toronto = getNearbyVenues( dt_toronto )
print(venues_dt_toronto.shape)
# venues_dt_toronto.rename({'Neighborhood':'Neighborhood Name'}, axis = 1, inplace = True)
# venues_dt_toronto.columns
venues_dt_toronto.head()
# Lets have a whole picture by checking the numbers of venues for each neighborhood, and cateogry of venues.
venues_dt_toronto[['Neighborhood Name','Venue Category']].groupby('Neighborhood Name').count()
venues_dt_toronto[['Neighborhood Name','Venue Category']].groupby('Venue Category').count()\
.sort_values(by='Neighborhood Name', ascending = False).head(10)
# We can tell that most common venue Category is: **Coffee Shop, Dafe, Restaurant, Hotel**, and **Park.**
# **Step(4)** Prepare venue category for Clustering
# We need apply one-hot encoding to the category column first.
dt_toronto_onehot = pd.get_dummies(venues_dt_toronto[['Venue Category']], prefix = "", prefix_sep="")
print(dt_toronto_onehot.shape)
dt_toronto_clustering = dt_toronto_onehot.copy()
# Now we put **neighborhood name** back to one-hot data frame as first column and rename it as **Name** just for convenience.
dt_toronto_onehot = pd.concat( [venues_dt_toronto[['Neighborhood Name']], dt_toronto_onehot], axis= 1)
print(dt_toronto_onehot.shape)
dt_toronto_onehot.head()
# Since we are clustering the neiborhood, we can group the data by neighborhood and categories before we apply k-means clustering.
#
dt_toronto_groups = dt_toronto_onehot.groupby('Neighborhood Name').sum()
# **Step(5)** Clustering the neighborhood according venue categories count
#
# We will run the k-means to cluster the neighborhood into 5 clusters.
# +
kclusters = 5
kmeans = KMeans(n_clusters= kclusters, random_state = 0).fit(dt_toronto_groups)
kmeans.labels_
# -
# Now it's time to put the label and neiborhood name together.
#
dt_toronto_groups['Label'] = kmeans.labels_
tmp = dt_toronto_groups['Label'].reset_index().rename({'Neighborhood Name':'Neighborhood'}, axis = 1)
cluster_result = pd.merge(tmp, dt_toronto)
print(cluster_result.shape)
cluster_result
# **Step (6)** Visualize the resulting on map
#
# +
#find out the center point of the map
lat, lng = cluster_result[['Latitude','Longitude']].max() + cluster_result[['Latitude','Longitude']].min()
lat, lng = lat /2, lng /2
print(lat, lng)
# create map
map_clusters = folium.Map(location=[lat, lng], zoom_start=13)
# set color scheme for the clusters
x = np.arange(kclusters)
ys = [i + x + (i*x)**2 for i in range(kclusters)]
colors_array = cm.rainbow(np.linspace(0, 1, len(ys)))
rainbow = [colors.rgb2hex(i) for i in colors_array]
# add markers to the map
markers_colors = []
for lat, lon, poi, cluster in zip(cluster_result['Latitude'],
cluster_result['Longitude'],
cluster_result['Neighborhood'],
kmeans.labels_):
label = folium.Popup(str(poi) + ' Cluster ' + str(cluster), parse_html=True)
folium.CircleMarker(
[lat, lon],
radius=5,
popup=label,
color=rainbow[cluster-1],
fill=True,
fill_color=rainbow[cluster-1],
fill_opacity=0.7).add_to(map_clusters)
map_clusters
| IBM_DataScience/AppliedDataScienceCapstone/Assignments/DP0701EN-Week3-Assignment-Neighbourhoods-Toronto.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Эффективность работы кода на примере алгоритмов сортировки.
# ### Оценка сложности
# В программировании нотация большого «О» (О-нотация) используется в качестве меры измерения, помогающей программистам оценивать или предполагать эффективность написанного блока кода, скрипта или алгоритма. «Сколько времени потребуется на работу этого кода? Какова его сложность в привязке к тем данным, которые он обрабатывает?»
#
# Точное время работы скрипта или алгоритма определить сложно. Оно зависит от многих факторов, например, от скорости процессора и прочих характеристик компьютера, на котором запускается этот скрипт или алгоритм. Поэтому нотация большого «О» используется не для оценки конкретного времени работы кода. Ее используют для оценки того, как быстро возрастает время обработки данных алгоритмом в привязке к объему этих данных.
# ### Смешная история
# В 2009 году у одной компании в Южной Африке была проблема со скоростью интернета. У этой компании было два офиса в 50 милях друг от друга. Сотрудники решили провести занимательный эксперимент и посмотреть, не будет ли быстрее пересылать данные при помощи голубиной почты.
#
# Они поместили 4GB данных на флешку, прикрепили ее к голубю и выпустили его из офиса, чтобы он полетел в другой офис. И…
#
# Почтовый голубь оказался быстрее интернет-связи. Он победил с большим отрывом (иначе история не была бы такой забавной). Больше того, когда голубь долетел до второго офиса, а случилось это через два часа, через интернет загрузилось только 4% данных.
# ### Виды сложности
#
# * Временная сложность алгоритма определяет число шагов, которые должен предпринять алгоритм, в зависимости от объема входящих данных (n).
# * Пространственная сложность алгоритма определяет количество памяти, которое потребуется занять для работы алгоритма, в зависимости от объема входящих данных (n).
# ### Постоянное время: O(1)
#
# Обратите внимание, что в истории с голубем, рассказанной выше, голубь доставил бы и 5KB, и 10MB, и 2TB данных, хранящихся на флешке, за совершенно одинаковое количество времени. Время, необходимое голубю для переноса данных из одного офиса в другой, это просто время, необходимое, чтобы пролететь 50 миль.
#
# Если использовать О-нотацию, время, за которое голубь может доставить данные из офиса А в офис Б, называется постоянным временем и записывается как O(1). Оно не зависит от объема входящих данных.
# ### Пример алгоритма со сложностью О(1)
# +
lst1 = [1, 2, 3, 4, 5, 3, 4, 5, 6, 6]
lst2 = ['A', 'a', 'D', 'DDG', 'ddf', 'dfdf']
print(f"Length of lst1 is {len(lst1)}",
f"Length of lst2 is {len(lst2)}", sep='\n')
# -
# Почему так происходит? (структура данных "Список" такова, что длина любого массива может быть получена за О(1), как и любой элемент списка)
# ### Линейное время: O(n)
#
# В отличие от пересылки голубем, передача данных через интернет будет занимать все больше и больше времени по мере увеличения объема передаваемых данных.
#
# Если использовать О-нотацию, мы можем сказать, что время, нужное для передачи данных из офиса А в офис Б через интернет, возрастает линейно и прямо пропорционально количеству передаваемых данных. Это время записывается как O(n), где n — количество данных, которое нужно передать.
#
# Следует учитывать, что в программировании «О» большое описывает наихудший сценарий. Допустим, у нас есть массив чисел, где мы должны найти какое-то определенное число при помощи цикла for. Оно может быть найдено при любой итерации, и чем раньше, тем быстрее функция завершит работу. О-нотация всегда указывает на верхнюю границу, т. е., описывает случай, когда алгоритму придется осуществить максимальное количество итераций, чтобы найти искомое число. Как, например, в том случае, если это число окажется последним в перебираемом массиве.
# +
def find_element(x, lst):
'''
Функция ищет элемент x в списке lst
Если он есть, возвращает True, если нет - False
'''
# иди по списку
for i in lst:
# если текущий элемент нашелся, вернем True
if x == i:
return True
# если дошли до конца и элемент не нашелся, вернем False
return False
lst = [1, 2, 3, 4, 5, 87, 543, 34637, 547489]
find_element(543, lst)
# -
# Почему линейное? Представим, что искомый элемент - последний в массиве. Тогда в худшем случае нам придется пройти все элементы нашего массива.
# ### Экспоненциальное время: O(2^n)
#
# Если сложность алгоритма описывается формулой O(2^n), значит, время его работы удваивается с каждым дополнением к набору данных. Кривая роста функции O(2^n) экспоненциальная: сначала она очень пологая, а затем стремительно поднимается вверх. Примером алгоритма с экспоненциальной сложностью может послужить рекурсивный расчет чисел Фибоначчи:
# +
def fibonacci(n):
'''
Функция рекурсивно считает
n'ное число Фибоначчи
'''
# Первое и второе числа Фибоначчи равны 1
if n in (1, 2):
return 1
# Если число не 1 и не 2, считай число как сумму двух предыдущих
return fibonacci(n - 1) + fibonacci(n - 2)
fibonacci(10)
# -
# ### Логарифмическое время: O(log n)
#
# Логарифмическое время поначалу понять сложнее. Поэтому для объяснения я воспользуюсь распространенным примером: концепцией бинарного поиска.
#
# Бинарный поиск это алгоритм поиска в отсортированных массивах. Работает он следующим образом. В отсортированном наборе данных выбирается серединный элемент и сравнивается с искомым значением. Если значение совпадает, поиск завершен.
#
# Если искомое значение больше, чем значение серединного элемента, нижняя половина набора данных (все элементы с меньшими значениями, чем у нашего серединного элемента) отбрасывается и дальнейший поиск ведется тем же способом в верхней половине.
#
# Если искомое значение меньше, чем значение серединного элемента, дальнейший поиск ведется в нижней половине набора данных.
#
# Эти шаги повторяются, при каждой итерации отбрасывая половину элементов, пока не будет найдено искомое значение или пока оставшийся набор данных станет невозможно разделить напополам:
# +
from random import randint
# Сделали список
a = []
for i in range(15):
# Случайное целое число от 1 до 49 включительно
a.append(randint(1, 50))
# Отсортировали список
a.sort()
# Распечатали
print(a)
# Вводим с клавиатуры искомое число
value = int(input())
# Индекс середины списка
mid = len(a) // 2
# Индекс начала списка
low = 0
# Индекс конца списка
high = len(a) - 1
# Пока позиция "середины" не равна нашему значению
# и левый конце области, где мы ищем, меньше или равен правому концу:
while a[mid] != value and low <= high:
# Если наше значение больше значения в центре области поиска:
if value > a[mid]:
# Начинаем искать в области "справа от середины"
low = mid + 1
else:
# Иначе начинаем искать в области "слева от середины"
high = mid - 1
# Середина новой области поиска
mid = (low + high) // 2
if low > high:
print("No value")
else:
print("ID =", mid)
# -
# ### Пузырьковая сортировка (n^2)
#
# Этот простой алгоритм выполняет итерации по списку, сравнивая элементы попарно и меняя их местами, пока более крупные элементы не «всплывут» в начало списка, а более мелкие не останутся на «дне».
#
# **Алгоритм**
# Сначала сравниваются первые два элемента списка. Если первый элемент больше, они меняются местами. Если они уже в нужном порядке, оставляем их как есть. Затем переходим к следующей паре элементов, сравниваем их значения и меняем местами при необходимости. Этот процесс продолжается до последней пары элементов в списке.
#
# При достижении конца списка процесс повторяется заново для каждого элемента. Это крайне неэффективно, если в массиве нужно сделать, например, только один обмен. Алгоритм повторяется n² раз, даже если список уже отсортирован.
# Библиотека, чтоб время засекать
from datetime import datetime as dt
# +
def bubble_sort(nums):
# Устанавливаем swapped в True, чтобы цикл запустился хотя бы один раз
swapped = True
while swapped:
swapped = False
# Идем циклом по индексам наших элементов
for i in range(len(nums) - 1):
# Если текущий элемент слева больше своего элемента справа
if nums[i] > nums[i + 1]:
# Меняем элементы местами
nums[i], nums[i + 1] = nums[i + 1], nums[i]
# Устанавливаем swapped в True для следующей итерации
swapped = True
# По окончании первого прогона цикла for
# самый большой элемент "Всплывет" наверх
# Проверяем, что оно работает
random_list_of_nums = [9, 5, 2, 1, 8, 4, 3, 7, 6]
bubble_sort(random_list_of_nums)
print(random_list_of_nums)
# -
# ### Сортировка выбором
#
# Этот алгоритм сегментирует список на две части: отсортированную и неотсортированную. Наименьший элемент удаляется из второго списка и добавляется в первый.
#
# **Алгоритм**
# На практике не нужно создавать новый список для отсортированных элементов. В качестве него используется крайняя левая часть списка. Находится наименьший элемент и меняется с первым местами.
#
# Теперь, когда нам известно, что первый элемент списка отсортирован, находим наименьший элемент из оставшихся и меняем местами со вторым. Повторяем это до тех пор, пока не останется последний элемент в списке.
# +
def selection_sort(nums):
# Значение i соответствует кол-ву отсортированных значений
for i in range(len(nums)):
# Исходно считаем наименьшим первый элемент
lowest_value_index = i
# Этот цикл перебирает несортированные элементы
for j in range(i + 1, len(nums)):
if nums[j] < nums[lowest_value_index]:
lowest_value_index = j
# Самый маленький элемент меняем с первым в списке
nums[i], nums[lowest_value_index] = nums[lowest_value_index], nums[i]
# Проверяем, что оно работает
random_list_of_nums = [9, 5, 2, 1, 8, 4, 3, 7, 6]
selection_sort(random_list_of_nums)
print(random_list_of_nums)
# -
# #### Время сортировки
# Затраты времени на сортировку выбором в среднем составляют O(n²), где n — количество элементов списка.
# ## Эмпирическое сравнение скорости
#
# Давайте просто сгенерируем случайные данные и посмотрим, какой из алгоритмов будет быстрее работать
# +
import numpy as np
# Сюда будем писать время сортировки
lst_bubble = []
lst_selection = []
# На каждом шаге генерируем случайный список длины i
for i in range(10, 501):
l = list(np.random.rand(i))
l2 = l.copy()
# Засекаем время и сортируем его пузырьком
t0 = float(dt.utcnow().timestamp())
bubble_sort(l)
t1 = float(dt.utcnow().timestamp()) - t0
lst_bubble.append(t1)
# Засекаем время и сортируем его вставками
t0 = float(dt.utcnow().timestamp())
selection_sort(l2)
t1 = float(dt.utcnow().timestamp()) - t0
lst_selection.append(t1)
# +
# Библиотека для рисования
from matplotlib.pyplot import plot, legend
# plot(список значений по оси X, список значений по оси Y, label = название линии на графике)
plot(range(10, 501), lst_bubble, label='bubble')
plot(range(10, 501), lst_selection, label='selection')
legend()
# -
# ### Сортировка вставкой
#
# Этот алгоритм сегментирует список на две части: отсортированную и неотсортированную. Перебираются элементы в неотсортированной части массива. Каждый элемент вставляется в отсортированную часть массива на то место, где он должен находиться.
#
# **Алгоритм**
# Проходим по массиву слева направо и обрабатываем по очереди каждый элемент. Слева от очередного элемента наращиваем отсортированную часть массива, справа по мере процесса потихоньку испаряется неотсортированная. В отсортированной части массива ищется точка вставки для очередного элемента. Сам элемент отправляется в буфер, в результате чего в массиве появляется свободная ячейка — это позволяет сдвинуть элементы и освободить точку вставки.
def insertion(data):
# Идем по нашему списку от элемента номер 1 (по счету это второй) до конца
for i in range(1, len(data)):
# Индекс "предыдущего" для i элемента
j = i - 1
# Сохраним элемент номер i в "буфер"
key = data[i]
# Пока предыдущий для i-того элемент больше следующего и индекс "предыдущего" не дошел до 0
# сдвигаем j влево, пока не дойдем до места, на которое надо вставить элемент key
# вставляем его
while data[j] > key and j >= 0:
data[j + 1] = data[j]
j -= 1
data[j + 1] = key
return data
lst = [1,5,7,9,2]
insertion(lst)
# ### Сортировка слиянием
# Этот алгоритм относится к алгоритмам «разделяй и властвуй». Он разбивает список на две части, каждую из них он разбивает ещё на две и т. д. Список разбивается пополам, пока не останутся единичные элементы.
#
# Соседние элементы становятся отсортированными парами. Затем эти пары объединяются и сортируются с другими парами. Этот процесс продолжается до тех пор, пока не отсортируются все элементы.
#
# **Алгоритм**
# Список рекурсивно разделяется пополам, пока в итоге не получатся списки размером в один элемент. Массив из одного элемента считается упорядоченным. Соседние элементы сравниваются и соединяются вместе. Это происходит до тех пор, пока не получится полный отсортированный список.
#
# Сортировка осуществляется путём сравнения наименьших элементов каждого подмассива. Первые элементы каждого подмассива сравниваются первыми. Наименьший элемент перемещается в результирующий массив. Счётчики результирующего массива и подмассива, откуда был взят элемент, увеличиваются на 1.
# +
# Функция, чтобы объединить два отсортированных списка
def merge(left_list, right_list):
# Сюда будет записан результирующий отсортированный список
sorted_list = []
# поначалу оба индекса в нуле
left_list_index = right_list_index = 0
# Длина списков часто используется, поэтому создадим переменные для удобства
left_list_length, right_list_length = len(left_list), len(right_list)
# Проходим по всем элементам обоих списков. _ - просто обозначение неиспользуемой переменной
# Цикл нужен, чтобы пройти все left_list_length + right_list_length элементов
for _ in range(left_list_length + right_list_length):
if left_list_index < left_list_length and right_list_index < right_list_length:
# Сравниваем первые элементы в начале каждого списка
# Если первый элемент левого подсписка меньше, добавляем его
# в отсортированный массив
if left_list[left_list_index] <= right_list[right_list_index]:
sorted_list.append(left_list[left_list_index])
left_list_index += 1
# Если первый элемент правого подсписка меньше, добавляем его
# в отсортированный массив
else:
sorted_list.append(right_list[right_list_index])
right_list_index += 1
# Если достигнут конец левого списка, элементы правого списка
# добавляем в конец результирующего списка
elif left_list_index == left_list_length:
sorted_list.append(right_list[right_list_index])
right_list_index += 1
# Если достигнут конец правого списка, элементы левого списка
# добавляем в отсортированный массив
elif right_list_index == right_list_length:
sorted_list.append(left_list[left_list_index])
left_list_index += 1
return sorted_list
# Сортировка слиянием
def merge_sort(nums):
# Возвращаем список, если он состоит из одного элемента
if len(nums) <= 1:
return nums
# Для того чтобы найти середину списка, используем деление без остатка
# Индексы должны быть integer
mid = len(nums) // 2
# Сортируем и объединяем подсписки (до mid и после mid)
left_list = merge_sort(nums[:mid])
right_list = merge_sort(nums[mid:])
# Объединяем отсортированные списки в результирующий
return merge(left_list, right_list)
# Проверяем, что оно работает
random_list_of_nums = [120, 45, 68, 250, 176]
random_list_of_nums = merge_sort(random_list_of_nums)
print(random_list_of_nums)
# -
# ### Быстрая сортировка
# Этот алгоритм также относится к алгоритмам «разделяй и властвуй». При правильной конфигурации он чрезвычайно эффективен и не требует дополнительной памяти, в отличие от сортировки слиянием. Массив разделяется на две части по разные стороны от опорного элемента. В процессе сортировки элементы меньше опорного помещаются перед ним, а равные или большие —позади.
#
# #### Алгоритм
# Быстрая сортировка начинается с разбиения списка и выбора одного из элементов в качестве опорного. А всё остальное передвигается так, чтобы этот элемент встал на своё место. Все элементы меньше него перемещаются влево, а равные и большие элементы перемещаются вправо.
#
# #### Реализация
# Существует много вариаций данного метода. Способ разбиения массива, рассмотренный здесь, соответствует схеме Хоара (создателя данного алгоритма).
# Функция принимает на вход список nums
def quicksort(nums):
# Если его длина 0 и 1, возвращает его же - такой список всегда отсортирован :)
if len(nums) <= 1:
return nums
else:
# Если длина > 1 в качестве опорного выбирается случайный элемент из списка
q = random.choice(nums)
# Создается три списка:
# Сюда запишем элементы < q
s_nums = []
# Сюда запишем элементы > q
m_nums = []
# Сюда запишем элементы = q
e_nums = []
# Пишем
for n in nums:
if n < q:
s_nums.append(n)
elif n > q:
m_nums.append(n)
else:
e_nums.append(n)
# А теперь рекурсивно проделаем ту же процедуру с левым и правым списками: s_nums и m_nums
return quicksort(s_nums) + e_nums + quicksort(m_nums)
import random
quicksort([1, 2, 3, 4, 9, 6, 7, 5])
# Немного оптимизируем по памяти - мы много сохраняли лишней информации. За опорную точку выбирать будем начало списка.
# +
# Функция принимает на вход список и индексы его начала и конца
def partition(array, start, end):
# За опорный элемент выбрано начало (его индекс)
pivot = start
# Идем по списку от 1 элемента до конца
for i in range(start+1, end+1):
# Если элемент меньше или равен опорному
if array[i] <= array[start]:
#Идем по списку от 1 элемента до конца
pivot += 1
array[i], array[pivot] = array[pivot], array[i]
array[pivot], array[start] = array[start], array[pivot]
return pivot
def quick_sort(array, start=0, end=None):
if end is None:
end = len(array) - 1
def _quicksort(array, start, end):
if start >= end:
return
pivot = partition(array, start, end)
_quicksort(array, start, pivot-1)
_quicksort(array, pivot+1, end)
return _quicksort(array, start, end)
# -
array = [29, 19, 47, 11, 6, 19, 24, 12, 17,
23, 11, 71, 41, 36, 71, 13, 18, 32, 26]
quick_sort(array)
print(array)
| lect6/.ipynb_checkpoints/O_notation-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 1
#
# 1. Calculate the mean of the numbers 2, 3, and 10.
a=[2,3,10]
sum(a)/len(a)
# # Exercise 2
#
# 1 - Make a list with 5 things in it.
a=[i for i in range(0,5)]
len(a)
# 2 - Add two more things.
a.extend([0,0])
a
# # Exercise 2
#
# 1. Make a dictionary whose keys are the strings "zero" through "nine" and whose values are ints 0 through 9.
# +
digits = {}
# Your code goes here
digits["zero"]=0
digits["one"]=1
digits["two"]=2
digits["three"]=3
digits["four"]=4
digits["five"]=5
digits["six"]=6
digits["seven"]=7
digits["eight"]=8
digits["nine"]=9
digits
# -
# # Exercise 3
#
# 1. Make a dictionary and experiment using different types as keys. Can containers be keys?
d={}
d[(0,0)]=0
d["0"]=0
d[0]=0
#d[[0]]=0 error
#d[{"0":0}]=0 error
#some container can be keys (tuples), other cannot (arrays, dictionaries, sets)
d
# # Exercises 4
#
# 1. Write an if statement that prints whether x is even or odd
x = [-1,0,2,3,0.5]
for i in x:
if i%2==0:
print("even")
else:
print("odd")
# # Exercises 5
#
# 1. Using a loop, calculate the factorial of 42 (the product of all integers up to and including 42).
n=1
for i in range(2,43):
n*=i
n
# # Exercises 6
#
# 1. Create a function that returns the factorial of a given number.
given=42
n=1
for i in range(2,given+1):
n*=i
n
# # Exercise 7
#
# 7A. Read the file 'big_animals.txt' and print each line and the length of each line.
f=open('big_animals.txt',"r")
file=f.readlines()
for i in file:
print("length:",len(i),"line:",i.strip())
f.close()
# 7B. Read the file 'big_animals.txt' again and print each line where the number of animals sighted is greater than 10. We'll won't output the lines with fewer than 10 animals.
f=open('big_animals.txt',"r")
file=f.readlines()
for line in file:
line1=line.split()
if int(line1[-1])>10:
print(line.strip())
f.close()
# 7C. Read the file 'big_animals.txt' again and return a list of tuples.
f=open('big_animals.txt',"r")
file=f.readlines()
out=[]
for line in file:
line1=line.split()
out.append(tuple(line1))
f.close()
# 7D. Turn the code for #7C into a function and use it on the files 'merida_animals.txt' and 'fergus_animals.txt'.
# +
def FileToTuple(fileName):
f=open(fileName,"r")
file=f.readlines()
out=[]
for line in file:
line1=line.split()
out.append(tuple(line1))
f.close()
return out
print(FileToTuple('merida_animals.txt'))
print(FileToTuple('fergus_animals.txt'))
# -
# # Exercise 8
#
# 1. Check your answer to Exercise 5 (the value of 42!) using the **math.gamma()** function.
import math
n=1
for i in range(2,43):
n*=i
n/math.gamma(43)
| 04-containers-exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ai-utilities] *
# language: python
# name: ai-utilities
# ---
# Using the configuration library is simple. Import the configure_settings function and call it. The only requirement
# for successful collection is to have an existing project.yml file in the current directory of the notebooks.
#
# This example project has one, so we will call configure_settings from there.
#
# Further usage require us to load the settings. Instead of having the user have to import other libraries, we expose a
# second function from notebook_config called get_settings() which will return an instance of ProjectConfiguration.
#
# To complete this example, we will obtain an instance and print out settings values.
from azure_utils.configuration.notebook_config import get_or_configure_settings
# Now that the functions are imported, lets bring up the UI to configure the settings ONLY if the subscription_id
# setting has not been modified from it's original value of '<>'.
#
# 
#
settings_object = get_or_configure_settings()
# Finally, get an instance of the settings. You will do this in the main (configurable) notebook, and all follow on
# notebooks.
#
# From the default provided file we know the following settings are there.
#
# subscription_id, resource_group
# + pycharm={"name": "#%%\n"}
sub_id = settings_object.get_value('subscription_id')
rsrc_grp = settings_object.get_value('resource_group')
print(sub_id, rsrc_grp)
# -
# You have completed this sample notebook.
# You are now ready to move on to the [AutoML Local](01_DataPrep.ipynb) notebook.
| notebooks/exampleconfiguration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Navigation Project
# In this proejct, we will present an early approach to DQNs to play a simple game using a UnityEnvironment that simulates a world with yellow and blue bananas. Our agent’s task is to understand the dynamics of this world, by collecting yellow bananas that give +1 Reward as fast as possible, while avoiding blue bananas that give -1 Reward. Without a DQN, we can watch our agent using the equiprobable policy. During that phase we will see that the agent randomly selects actions, making it truly useless.
# In this notebook, I used the Unity ML-Agents environment for the first project of the [Deep Reinforcement Learning Nanodegree](https://www.udacity.com/course/deep-reinforcement-learning-nanodegree--nd893).
#
# ### 1. Start the Environment
#
# We begin by importing some necessary packages. If the code cell below returns an error, please revisit the project instructions to double-check that you have installed [Unity ML-Agents](https://github.com/Unity-Technologies/ml-agents/blob/master/docs/Installation.md) and [NumPy](http://www.numpy.org/).
# !pip -q install ./python
from unityagents import UnityEnvironment
import numpy as np
# Other imports
# +
import random
import time
from collections import namedtuple, deque
import matplotlib.pyplot as plt
# %matplotlib inline
import torch
import torch.nn.functional as F
import torch.optim as optim
# -
start = time.time()
from dqn_agent import Agent
# Next, we will start the environment! **_Before running the code cell below_**, change the `file_name` parameter to match the location of the Unity environment that you downloaded.
#
# - **Mac**: `"path/to/Banana.app"`
# - **Windows** (x86): `"path/to/Banana_Windows_x86/Banana.exe"`
# - **Windows** (x86_64): `"path/to/Banana_Windows_x86_64/Banana.exe"`
# - **Linux** (x86): `"path/to/Banana_Linux/Banana.x86"`
# - **Linux** (x86_64): `"path/to/Banana_Linux/Banana.x86_64"`
# - **Linux** (x86, headless): `"path/to/Banana_Linux_NoVis/Banana.x86"`
# - **Linux** (x86_64, headless): `"path/to/Banana_Linux_NoVis/Banana.x86_64"`
#
# For instance, if you are using a Mac, then you downloaded `Banana.app`. If this file is in the same folder as the notebook, then the line below should appear as follows:
# ```
# env = UnityEnvironment(file_name="Banana.app")
# ```
env = UnityEnvironment(file_name="/data/Banana_Linux_NoVis/Banana.x86_64")
# Now we have a link between our notebook and the actual Unity Environment. We can now start interacting with the environment and get feedback by our interactions.
# #### Brain
# Environments contain **_brains_** which are responsible for deciding the actions of their associated agents. Here we check for the first brain available, and set it as the default brain we will be controlling from Python.
# +
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
print(brain)
# -
# ### 2. Examine the State and Action Spaces
#
# The simulation contains a single agent that navigates a large environment. At each time step, it has four actions at its disposal:
# - `0` - walk forward
# - `1` - walk backward
# - `2` - turn left
# - `3` - turn right
#
# The state space has `37` dimensions and contains the agent's velocity, along with ray-based perception of objects around agent's forward direction. A reward of `+1` is provided for collecting a yellow banana, and a reward of `-1` is provided for collecting a blue banana.
#
# Run the code cell below to print some information about the environment.
# +
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents in the environment
print('Number of agents:', len(env_info.agents))
# number of actions
action_size = brain.vector_action_space_size
print('Number of actions:', action_size)
# examine the state space
state = env_info.vector_observations[0]
print('States look like:', state)
state_size = len(state)
print('States have length:', state_size)
# -
# ### 3. Take Random Actions in the Environment
#
# In the next code cell, we try to use the Python API to control the agent and receive feedback from the environment. Once this cell is executed, we can watch the agent's performance, if it selects an action (uniformly) at random with each time step. A window should pop up that allows you to observe the agent, as it moves through the environment.
#
# Of course, as part of the project, you'll have to change the code so that the agent is able to use its experience to gradually choose better actions when interacting with the environment!
def play_game(use_DQN = False):
env_info = env.reset(train_mode=False)[brain_name] # reset the environment
state = env_info.vector_observations[0] # get the current state
score = 0 # initialize the score
epsilon = 0
while True:
# if we play using DQN, the agent will act based on greedy-policy of the state
# otherwise we use the equiprobable policy to select randomly any of the available actions
action = np.int32(agent.act(state, epsilon)) if use_DQN else np.random.randint(action_size)
env_info = env.step(action)[brain_name] # send the action to the environment
next_state = env_info.vector_observations[0] # get the next state
reward = env_info.rewards[0] # get the reward
done = env_info.local_done[0] # see if episode has finished
score += reward # update the score
state = next_state # roll over the state to next time step
if done: # exit loop if episode finished
break
print("Score: {}".format(score))
play_game(use_DQN = False)
# ### 4. Interacting with the Environment to learn
#
# At this stage, we will put everything together so we can interact with the environment for several episodes, until we come up with a good policy estimation via our DQN network, and eventually solve the RL task. We will define a function DQN which will put everything together and drive the learning through interaction.
def dqn(n_episodes=2000, max_t=1000, eps_start=1.0, eps_end=0.01, eps_decay=0.995):
"""Deep Q-Learning.
Params
======
n_episodes (int): maximum number of training episodes
max_t (int): maximum number of timesteps per episode
eps_start (float): starting value of epsilon, for epsilon-greedy action selection
eps_end (float): minimum value of epsilon
eps_decay (float): multiplicative factor (per episode) for decreasing epsilon
"""
scores = [] # list containing scores from each episode
scores_window = deque(maxlen=100) # last 100 scores
moving_avgs = [] # list of moving averages
eps = eps_start # initialize epsilon
for i_episode in range(1, n_episodes+1):
env_info = env.reset(train_mode=True)[brain_name] # reset environment
state = env_info.vector_observations[0] # get current state
score = 0
for t in range(max_t):
action = agent.act(state, eps)
env_info = env.step(action)[brain_name] # send action to environment
next_state = env_info.vector_observations[0] # get next state
reward = env_info.rewards[0] # get reward
done = env_info.local_done[0] # see if episode has finished
agent.step(state, action, reward, next_state, done)
state = next_state
score += reward
if done:
break
scores.append(score) # save score for plotting
scores_window.append(score) # save score for moving average
moving_avg = np.mean(scores_window) # calculate moving average
moving_avgs.append(moving_avg) # save moving average
eps = max(eps_end, eps_decay*eps) # decrease epsilon
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, moving_avg), end="")
if i_episode % 100 == 0:
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, moving_avg))
if moving_avg >= 13.10:
print('\nEnvironment solved in {:d} episodes!\tAverage Score: {:.2f}'.format(i_episode-100, moving_avg))
torch.save(agent.qnetwork_local.state_dict(), 'checkpoint.pth')
break
return scores, moving_avgs
# Our DQN method will run for a maximum of 2000 episodes, after which if we have not reached our desired per 100 episodes score, the method will terminate.
# In a nutshell, at every episode we will
# * Reset the environment
# * For each state, loop by getting the current state
# * Follow the epsilon-greedy-policy so the agent acts on that state according policy and epsilon.
# * Get observation and reward
# * Use the agent's step function as described above to populate the experience buffer and possibly trigger the learn function.
#
# We keep doing this until our average score over 100 episodes is equal or higher than 13 (which relates to gathering 13 yellow bananas on time). After every 100 episodes we save a checkpoint if the average score has increased, but we also overwrite this checkpoint once we reach our goal. Now one might ask, what is the use to save intermediate scores. Well, sometimes you might interrupt this method earlier, and just want to check how your model performs with this "early" training (so it is there just for curiosity reasons).
# ### 5. Performance and Visualizing the scores per episode
start = time.time()
agent = Agent(state_size=state_size, action_size=action_size, seed=42)
scores, avgs = dqn(n_episodes=1000, eps_start=1.0, eps_end=0.02, eps_decay=0.95)
end = time.time()
elapsed = (end - start) / 60.0 # in minutes
print("Elapsed Time: {0:3.2f} mins.".format(elapsed))
# Given our existing approach, we reached the desired 13+ cumulative result (over 100 episodes) at episode 189 in 6:22 mins. And here is the visualization of the cumulative reward (over 100 episodes) all the way from episode one to 645.
# plot the scores
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(len(scores)), scores, label='DQN')
plt.plot(np.arange(len(scores)), avgs, c='r', label='average')
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.legend(loc='upper left');
plt.show()
end = time.time()
elapsed = (end - start) / 60.0 # in minutes
print("Elapsed Time: {0:3.2f} mins.".format(elapsed))
# ### 6. Testing the saved agent
# +
start = time.time()
# initialize the agent
agent = Agent(state_size=state_size, action_size=action_size, seed=42)
# load the weights from file
checkpoint = 'checkpoint.pth'
agent.qnetwork_local.load_state_dict(torch.load(checkpoint))
num_episodes = 10
scores = []
for i_episode in range(1,num_episodes+1):
env_info = env.reset(train_mode=False)[brain_name] # reset the environment
state = env_info.vector_observations[0] # get the current state
score = 0 # initialize the score
while True:
action = agent.act(state, eps=0) # select an action
env_info = env.step(action)[brain_name] # send the action to the env
next_state = env_info.vector_observations[0] # get the next state
reward = env_info.rewards[0] # get the reward
done = env_info.local_done[0] # is episode done?
score += reward # update the score
state = next_state # set state to next state
if done: # are we done yet?
scores.append(score)
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores)))
break
# plot the scores
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(len(scores)), scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
end = time.time()
elapsed = (end - start) / 60.0 # in minutes
print("\nElapsed Time: {0:3.2f} mins.".format(elapsed))
# -
# When finished, you can close the environment.
env.close()
# ### 7. Further Refinements
# There are several refinements to the existing ones that could add up to the performance of our smart DQN agent. Based on our existing setup, it is much likely that playing further with the hyperparameters we might came up with an agent that could learn faster.
# However, the agent could greatly benefit from additional RL-specific refinements which we could introduce to our existing setup.
# More specifically:
# * Prioritized learning could be used so rather than randomly sampling SARS tuples from our experience buffer, we could revisit more frequently the SARS tuples that better relate to rational actions over specific states.
# * A Double-DQN could be used to address the DQN problem of action value overestimation
# * The QNetwork pytorch model could be changed to adapt a dueling DQN architecture, which incorporates two streams. One for the state-values function and another for the advantage-values function.
| p1_navigation/Navigation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="ULdrhOaVbsdO"
# # Acme: Tutorial
#
# <a href="https://colab.research.google.com/github/deepmind/acme/blob/master/examples/tutorial.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
#
# This colab provides an overview of how Acme's modules can be stacked together to create reinforcement learning agents. It shows how to fit networks to environment specs, create actors, learners, replay buffers, datasets, adders, and full agents. It also highlights where you can swap out certain modules to create your own Acme based agents.
# + [markdown] colab_type="text" id="xaJxoatMhJ71"
# ## Installation
#
# In the first few cells we'll start by installing all of the necessary dependencies (and a few optional ones).
# + cellView="form" colab={} colab_type="code" id="KH3O0zcXUeun"
#@title Install necessary dependencies.
# # !sudo apt-get install -y xvfb ffmpeg
# # !pip install 'gym==0.10.11'
# # !pip install imageio
# # !pip install PILLOW
# # !pip install 'pyglet==1.3.2'
# # !pip install pyvirtualdisplay
# # !pip install dm-acme
# # !pip install dm-acme[reverb]
# # !pip install dm-acme[tf]
# # !pip install dm-acme[envs]
from IPython.display import clear_output
clear_output()
# + [markdown] colab_type="text" id="VEEj3Qw60y73"
# ### Install dm_control
#
# The next cell will install environments provided by `dm_control` _if_ you have an institutional MuJoCo license. This is not necessary, but without this you won't be able to use the `dm_cartpole` environment below and can instead follow this colab using `gym` environments. To do so simply expand the following cell, paste in your license file, and run the cell.
#
# Alternatively, Colab supports using a Jupyter kernel on your local machine which can be accomplished by following the guidelines here: https://research.google.com/colaboratory/local-runtimes.html. This will allow you to install `dm_control` by following instructions in https://github.com/deepmind/dm_control and using a personal MuJoCo license.
#
# + cellView="both" colab={} colab_type="code" id="IbZxYDxzoz5R"
# #@title Add your License
# #@test {"skip": true}
# mjkey = """
# """.strip()
# mujoco_dir = "$HOME/.mujoco"
# # Install OpenGL dependencies
# # !apt-get update && apt-get install -y --no-install-recommends \
# # libgl1-mesa-glx libosmesa6 libglew2.0
# # Get MuJoCo binaries
# # !wget -q https://www.roboti.us/download/mujoco200_linux.zip -O mujoco.zip
# # !unzip -o -q mujoco.zip -d "$mujoco_dir"
# # Copy over MuJoCo license
# # !echo "$mjkey" > "$mujoco_dir/mjkey.txt"
# # Install dm_control
# # !pip install dm_control
# Configure dm_control to use the OSMesa rendering backend
# %env MUJOCO_GL=osmesa
# Check that the installation succeeded
# try:
# from dm_control import suite
# env = suite.load('cartpole', 'swingup')
# pixels = env.physics.render()
# except Exception as e:
# raise e from RuntimeError(
# 'Something went wrong during installation. Check the shell output above '
# 'for more information.')
# else:
# from IPython.display import clear_output
# clear_output()
# del suite, env, pixels
# + [markdown] colab_type="text" id="c-H2d6UZi7Sf"
# ## Import Modules
#
# Now we can import all the relevant modules.
# + cellView="both" colab={} colab_type="code" id="HJ74Id-8MERq"
# %%capture
import copy
import pyvirtualdisplay
import imageio
import base64
import IPython
from acme import environment_loop
from acme.tf import networks
from acme.adders import reverb as adders
from acme.agents.tf import actors as actors
from acme.datasets import reverb as datasets
from acme.wrappers import gym_wrapper
from acme import specs
from acme import wrappers
from acme.agents.tf import d4pg
from acme.agents import agent
from acme.tf import utils as tf2_utils
from acme.utils import loggers
import gym
import dm_env
import matplotlib.pyplot as plt
import numpy as np
import reverb
import sonnet as snt
import tensorflow as tf
# Set up a virtual display for rendering OpenAI gym environments.
display = pyvirtualdisplay.Display(visible=0, size=(1400, 900)).start()
# -
import robosuite as suite
from robosuite.wrappers import GymWrapper
from robosuite.controllers import load_controller_config
from collections import OrderedDict
# + [markdown] colab_type="text" id="I6KuVGSk4uc9"
# ## Load an environment
#
# We can now load an environment. In what follows we'll create an environment in order to generate and visualize a single state from that environment. Just select the environment you want to use and run the cell.
# +
env_config = {
"control_freq": 20,
"env_name": "Lift",
"hard_reset": False,
"horizon": 500,
"ignore_done": False,
"reward_scale": 1.0,
"camera_names": "frontview",
"robots": [
"Panda"
]
}
controller_config = load_controller_config(default_controller="OSC_POSITION")
# -
env_suite = suite.make(**env_config,
has_renderer=False,
has_offscreen_renderer=False,
use_camera_obs=False,
reward_shaping=True,
controller_configs=controller_config,
)
keys = ["object-state"]
for idx in range(len(env_suite.robots)):
keys.append(f"robot{idx}_proprio-state")
# Wrap environment so it's compatible with Gym API
environment = GymWrapper(env_suite, keys=keys)
environment = gym_wrapper.GymWrapper(environment)
environment = wrappers.SinglePrecisionWrapper(environment)
def render(env):
return env.environment.render()
# +
# frame = render(environment)
# plt.imshow(frame)
# plt.axis('off')
# +
# observation_spec = OrderedDict()
# for key in keys:
# observation_spec[key] = specs.Array(environment.observation_spec()[key].shape, np.float32, key)
# reward_spec = specs.Array((), np.float32)
# discount_spec = specs.BoundedArray((), np.float32, 0.0, 1.0)
# +
# action_spec=specs.BoundedArray(
# shape=environment.action_spec[0].shape, dtype=np.float32, minimum=-1., maximum=1.)
# +
# observation_spec, action_spec, reward_spec, discount_spec
# + [markdown] colab_type="text" id="dQJprnmn41fP"
# ### Environment Spec
#
# We will later interact with the environment in a loop corresponding to the following diagram:
#
# <img src="https://github.com/deepmind/acme/raw/master/docs/diagrams/environment_loop.png" width="500" />
#
# But before we start building an agent to interact with this environment, let's first look at the types of objects the environment either returns (e.g. observations) or consumes (e.g. actions). The `environment_spec` will show you the form of the *observations*, *rewards* and *discounts* that the environment exposes and the form of the *actions* that can be taken.
#
# -
#environment_spec = specs.EnvironmentSpec(observations=observation_spec, actions=action_spec, rewards=reward_spec, discounts=discount_spec)
environment_spec = specs.make_environment_spec(environment)
# + colab={} colab_type="code" id="lph6pbmu7UfZ"
print('actions:\n', environment_spec.actions, '\n')
print('observations:\n', environment_spec.observations, '\n')
print('rewards:\n', environment_spec.rewards, '\n')
print('discounts:\n', environment_spec.discounts, '\n')
# + [markdown] colab_type="text" id="M0h2xojW9wT2"
# ## Build a policy network that maps observations to actions.
#
# The most important part of a reinforcement learning algorithm is potentially the policy that maps environment observations to actions. We can use a simple neural network to create a policy, in this case a simple feedforward MLP with layer norm. For our TensorFlow agents we make use of the `sonnet` library to specify networks or modules; all of the networks we will work with also have an initial batch dimension which allows for batched inference/learning.
#
# It is possible that the the observations returned by the environment are nested in some way: e.g. environments from the `dm_control` suite are frequently returned as dictionaries containing `position` and `velocity` entries. Our network is allowed to arbitarily map this dictionary to produce an action, but in this case we will simply concatenate these observations before feeding it through the MLP. We can do so using Acme's `batch_concat` utility to flatten the nested observation into a single dimension for each batch. If the observation is already flat this will be a no-op.
#
# Similarly, the output of the MLP is may have a different range of values than the action spec dictates. For this, we can use Acme's `TanhToSpec` module to rescale our actions to meet the spec.
# + colab={} colab_type="code" id="JZKWAFEQz5NP"
# Calculate how big the last layer should be based on total # of actions.
action_spec = environment_spec.actions
action_size = np.prod(action_spec.shape, dtype=int)
exploration_sigma = 0.3
# In order the following modules:
# 1. Flatten the observations to be [B, ...] where B is the batch dimension.
# 2. Define a simple MLP which is the guts of this policy.
# 3. Make sure the output action matches the spec of the actions.
policy_modules = [
tf2_utils.batch_concat,
networks.LayerNormMLP(layer_sizes=(300, 200, action_size)),
networks.TanhToSpec(spec=environment_spec.actions)]
policy_network = snt.Sequential(policy_modules)
# We will also create a version of this policy that uses exploratory noise.
behavior_network = snt.Sequential(
policy_modules + [networks.ClippedGaussian(exploration_sigma),
networks.ClipToSpec(action_spec)])
# + [markdown] colab_type="text" id="9FbmoOpZKwid"
# ## Create an actor
#
# An `Actor` is the part of our framework that directly interacts with an environment by generating actions. In more detail the earlier diagram can be expanded to show exactly how this interaction occurs:
#
# <img src="https://github.com/deepmind/acme/raw/master/docs/diagrams/actor_loop.png" width="500" />
#
# While you can always write your own actor, in Acme we also provide a number of useful premade versions. For the network we specified above we will make use of a `FeedForwardActor` that wraps a single feed forward network and knows how to do things like handle any batch dimensions or record observed transitions.
# + colab={} colab_type="code" id="rpTs49OWKvMV"
actor = actors.FeedForwardActor(policy_network)
# + [markdown] colab_type="text" id="oMxSAzDWYhC4"
# All actors have the following public methods and attributes:
# + colab={} colab_type="code" id="iKoARV3gYfcV"
[method_or_attr for method_or_attr in dir(actor) # pylint: disable=expression-not-assigned
if not method_or_attr.startswith('_')]
# + [markdown] colab_type="text" id="3qrb2ZGhAoR5"
# ## Evaluate the random actor's policy.
#
#
# Although we have instantiated an actor with a policy, the policy has not yet learned to achieve any task reward, and is essentially just acting randomly. However this is a perfect opportunity to see how the actor and environment interact. Below we define a simple helper function to display a video given frames from this interaction, and we show 500 steps of the actor taking actions in the world.
# + colab={} colab_type="code" id="OIJRbtAlxQVu"
def display_video(frames, filename='temp.mp4'):
"""Save and display video."""
# Write video
with imageio.get_writer(filename, fps=60) as video:
for frame in frames:
video.append_data(frame)
# Read video and display the video
video = open(filename, 'rb').read()
b64_video = base64.b64encode(video)
video_tag = ('<video width="320" height="240" controls alt="test" '
'src="data:video/mp4;base64,{0}">').format(b64_video.decode())
return IPython.display.HTML(video_tag)
# -
def render(env):
return env.render()
# + colab={} colab_type="code" id="wdCeuvHeUwwm"
# # Run the actor in the environment for desired number of steps.
# frames = []
# num_steps = 500
# timestep = environment.reset()
# for _ in range(num_steps):
# frames.append(render(environment))
# action = actor.select_action(timestep.observation)
# timestep = environment.step(action)
# # Save video of the behaviour.
# display_video(np.array(frames))
# + [markdown] colab_type="text" id="iz_OYw60MHmc"
# ## Storing actor experiences in a replay buffer
#
# Many RL agents utilize a data structure such as a replay buffer to store data from the environment (e.g. observations) along with actions taken by the actor. This data will later be fed into a learning process in order to update the policy. Again we can expand our earlier diagram to include this step:
#
# <img src="https://github.com/deepmind/acme/raw/master/docs/diagrams/batch_loop.png" width="500" />
#
# In order to make this possible, Acme leverages [Reverb](https://github.com/deepmind/reverb) which is an efficient and easy-to-use data storage and transport system designed for Machine Learning research. Below we will create the replay buffer before interacting with it.
# -
# + colab={} colab_type="code" id="uQ0NAMwGWHu6"
# Create a table with the following attributes:
# 1. when replay is full we remove the oldest entries first.
# 2. to sample from replay we will do so uniformly at random.
# 3. before allowing sampling to proceed we make sure there is at least
# one sample in the replay table.
# 4. we use a default table name so we don't have to repeat it many times below;
# if we left this off we'd need to feed it into adders/actors/etc. below.
replay_buffer = reverb.Table(
name=adders.DEFAULT_PRIORITY_TABLE,
max_size=1000000,
remover=reverb.selectors.Fifo(),
sampler=reverb.selectors.Uniform(),
signature=adders.NStepTransitionAdder.signature(
environment_spec),
rate_limiter=reverb.rate_limiters.MinSize(min_size_to_sample=1))
# Get the server and address so we can give it to the modules such as our actor
# that will interact with the replay buffer.
replay_server = reverb.Server([replay_buffer], port=None)
replay_server_address = 'localhost:%d' % replay_server.port
# + [markdown] colab_type="text" id="joJaFWKjNep-"
# We could interact directly with Reverb in order to add data to replay. However in Acme we have an additional layer on top of this data-storage that allows us to use the same interface no matter what kind of data we are inserting.
#
# This layer in Acme corresponds to an `Adder` which adds experience to a data table. We provide several adders that differ based on the type of information that is desired to be stored in the table, however in this case we will make use of an `NStepTransitionAdder` which stores simple transitions (if N=1) or accumulates N-steps to form an aggregated transition.
# + colab={} colab_type="code" id="9CozET1ENd_j"
# Create a 5-step transition adder where in between those steps a discount of
# 0.99 is used (which should be the same discount used for learning).
adder = adders.NStepTransitionAdder(
client=reverb.Client(replay_server_address),
n_step=10,
discount=0.999)
# + [markdown] colab_type="text" id="rMOgmNpsiyC5"
# We can either directly use the adder to add transitions to replay directly using the `add()` and `add_first()` methods as follows:
# -
from tqdm import tqdm
# + cellView="both" colab={} colab_type="code" id="ia16FN4dPWk4"
# # %%time
# num_episodes = 2 #@param
# for episode in tqdm(range(num_episodes)):
# timestep = environment.reset()
# adder.add_first(timestep)
# count = 0
# while not timestep.last():
# print(f"current step: {count}")
# count += 1
# if(count > 10):
# break
# action = actor.select_action(timestep.observation)
# timestep = environment.step(action)
# print(f"action: {action}, timestep: {timestep}")
# adder.add(action=action, next_timestep=timestep)
# + [markdown] colab_type="text" id="vxNHy-h2Wl9q"
# Since this is a common enough way to observe data, `Actor`s in Acme generally take an `Adder` instance that they use to define their observation methods. We saw earlier that the `FeedForwardActor` like all `Actor`s defines `observe` and `observe_first` methods. If we give the actor an `Adder` instance at init then it will use this adder to make observations.
# + colab={} colab_type="code" id="Xp-YYHaHWQg_"
actor = actors.FeedForwardActor(policy_network=behavior_network, adder=adder)
# + [markdown] colab_type="text" id="ertrRdjOZHZ3"
# Below we repeat the same process, but using `actor` and its `observe` methods. We note these subtle changes below.
# + cellView="both" colab={} colab_type="code" id="rJ0_UCLeZPOt"
# # %%time
# num_episodes = 2 #@param
# for episode in range(num_episodes):
# timestep = environment.reset()
# actor.observe_first(timestep) # Note: observe_first.
# while not timestep.last():
# action = actor.select_action(timestep.observation)
# timestep = environment.step(action)
# actor.observe(action=action, next_timestep=timestep) # Note: observe.
# + [markdown] colab_type="text" id="OVj1cBw8epeh"
# ## Learning from experiences in replay
# Acme provides multiple learning algorithms/agents. Here, we will use the Acme's D4PG learning algorithm to learn from the data collected by the actor. To do so, we first create a TensorFlow dataset from the Reverb table using the `make_dataset` function.
# + colab={} colab_type="code" id="IbDfFYTRqmoz"
# This connects to the created reverb server; also note that we use a transition
# adder above so we'll tell the dataset function that so that it knows the type
# of data that's coming out.
dataset = datasets.make_dataset(
server_address=replay_server_address,
batch_size=2048,
transition_adder=True)
dataset_iterator = iter(dataset)
# + [markdown] colab_type="text" id="G5VUXDBytDu9"
# In what follows we'll make use of D4PG, an actor-critic learning algorithm. D4PG is a somewhat complicated algorithm, so we'll leave a full explanation of this method to the accompanying paper (see the documentation).
#
# However, since D4PG is an actor-critic algorithm we will have to specify a critic for it (a value function). In this case D4PG uses a distributional critic as well. D4PG also makes use of online and target networks so we need to create copies of both the policy_network (from earlier) and the new critic network we are about to create.
#
# To build our critic networks, we use a ***multiplexer***, which is simply a neural network module that takes multiple inputs and processes them in different ways before combining them and processing further. In the case of Acme's `CriticMultiplexer`, the inputs are observations and actions, each with their own network torso. There is then a critic network module that processes the outputs of the observation network and the action network and outputs a tensor.
#
# Finally, in order to optimize these networks the learner must receive networks with the variables created. We have utilities in Acme to handle exactly this, and we do so in the final lines of the following code block.
# + colab={} colab_type="code" id="OgTk7IAKXKz0"
critic_network = snt.Sequential([
networks.CriticMultiplexer(
observation_network=tf2_utils.batch_concat,
action_network=tf.identity,
critic_network=networks.LayerNormMLP(
layer_sizes=(400, 300),
activate_final=True)),
# Value-head gives a 51-atomed delta distribution over state-action values.
networks.DiscreteValuedHead(vmin=-500., vmax=500., num_atoms=51)])
# Create the target networks
target_policy_network = copy.deepcopy(policy_network)
target_critic_network = copy.deepcopy(critic_network)
# We must create the variables in the networks before passing them to learner.
tf2_utils.create_variables(network=policy_network,
input_spec=[environment_spec.observations])
tf2_utils.create_variables(network=critic_network,
input_spec=[environment_spec.observations,
environment_spec.actions])
tf2_utils.create_variables(network=target_policy_network,
input_spec=[environment_spec.observations])
tf2_utils.create_variables(network=target_critic_network,
input_spec=[environment_spec.observations,
environment_spec.actions])
# + [markdown] colab_type="text" id="ENTZ_cj3GiLr"
# We can now create a learner that uses these networks. Note that here we're using the same discount factor as was used in the transition adder. The rest of the parameters are reasonable defaults.
#
# Note however that we will log output to the terminal at regular intervals. We have also turned off checkpointing of the network weights (i.e. saving them). This is usually used by default but can cause issues with interactive colab sessions.
# +
# d4pg.D4PGLearner?
# + colab={} colab_type="code" id="5DkJrgkSW94O"
learner = d4pg.D4PGLearner(policy_network=policy_network,
critic_network=critic_network,
target_policy_network=target_policy_network,
target_critic_network=target_critic_network,
dataset_iterator=dataset_iterator,
discount=0.999,
target_update_period=100,
policy_optimizer=snt.optimizers.Adam(1e-4),
critic_optimizer=snt.optimizers.Adam(1e-4),
# Log learner updates to console every 10 seconds.
logger=loggers.TerminalLogger(time_delta=10.),
checkpoint=False)
# + [markdown] colab_type="text" id="ANG_L3e1dGoT"
# Inspecting the learner's public methods we see that it primarily exists to expose its variables and update them. IE this looks remarkably similar to supervised learning.
# + colab={} colab_type="code" id="Tl3Hu9nHcyp5"
[method_or_attr for method_or_attr in dir(learner) # pylint: disable=expression-not-assigned
if not method_or_attr.startswith('_')]
# + [markdown] colab_type="text" id="oG9SglevFrmQ"
# The learner's `step()` method samples a batch of data from the replay dataset given to it, and performs optimization using the optimizer, logging loss metrics along the way. Note: in order to sample from the replay dataset, there must be at least 1000 elements in the replay buffer (which should already have from the actor's added experiences.)
# + colab={} colab_type="code" id="estCksXXFi2_"
# learner.step()
# + [markdown] colab_type="text" id="dL6U_Wi2HTA2"
# # Training loop
# Finally, we can put all of the pieces together and run some training steps in the environment, alternating the actor's experience gathering with learner's learning.
#
# This is a simple training loop that runs for `num_training_episodes` episodes where the actor and learner take turns generating and learning from experience respectively:
#
# - Actor acts in environment & adds experience to replay for `min_actor_steps_per_iteration` steps<br>
# - Learner samples from replay data and learns from it for `num_learner_steps_per_iteration` steps<br>
#
# Note: Since the learner and actor are sharing a policy network, any learning done on the learner, automatically is transferred to the actor's policy.
#
# + cellView="both" colab={} colab_type="code" id="zC0PgbeyHSSP"
# %%time
num_training_episodes = 100 # @param {type: "integer"}
min_actor_steps_before_learning = 3300 # @param {type: "integer"}
num_actor_steps_per_iteration = 2500 # @param {type: "integer"}
num_learner_steps_per_iteration = 1000 # @param {type: "integer"}
learner_steps_taken = 0
actor_steps_taken = 0
print_every = 1
for episode in range(num_training_episodes):
timestep = environment.reset()
actor.observe_first(timestep)
episode_return = 0
count = 0
while not timestep.last() and count < 500:
count += 1
# Get an action from the agent and step in the environment.
action = actor.select_action(timestep.observation)
next_timestep = environment.step(action)
# Record the transition.
actor.observe(action=action, next_timestep=next_timestep)
# Book-keeping.
episode_return += next_timestep.reward
actor_steps_taken += 1
timestep = next_timestep
# See if we have some learning to do.
if (actor_steps_taken >= min_actor_steps_before_learning and
actor_steps_taken % num_actor_steps_per_iteration == 0):
# Learn.
for learner_step in range(num_learner_steps_per_iteration):
learner.step()
learner_steps_taken += num_learner_steps_per_iteration
if((episode+1) % print_every) == 0:
# Log quantities.
print('Episode: %d | Return: %f | Learner steps: %d | Actor steps: %d'%(
episode, episode_return, learner_steps_taken, actor_steps_taken))
# + [markdown] colab_type="text" id="Z5w7y3iYYck7"
# ## Putting it all together: an Acme agent
#
# <img src="https://github.com/deepmind/acme/raw/master/docs/diagrams/agent_loop.png" width="500" />
#
# Now that we've used all of the pieces and seen how they can interact, there's one more way we can put it all together. In the Acme design scheme, an agent is an entity with both a learner and an actor component that will piece together their interactions internally. An agent handles the interchange between actor adding experiences to the replay buffer and learner sampling from it and learning and in turn, sharing its weights back with the actor.
#
# Similar to how we used `num_actor_steps_per_iteration` and `num_learner_steps_per_iteration` parameters in our custom training loop above, the agent parameters `min_observations` and `observations_per_step` specify the structure of the agent's training loop.
# * `min_observations` specifies how many actor steps need to have happened to start learning.
# * `observations_per_step` specifies how many actor steps should occur in between each learner step.
# + colab={} colab_type="code" id="bqFpIHE-aRRg"
d4pg_agent = agent.Agent(actor=actor,
learner=learner,
min_observations=1000,
observations_per_step=8.)
# + [markdown] colab_type="text" id="o63pJeJl-Lv_"
# Of course we could have just used the `agents.D4PG` agent directly which sets
# all of this up for us. We'll stick with this agent we've just created, but most of the steps outlined in this tutorial can be skipped by just making use of a
# prebuilt agent and the environment loop.
# + [markdown] colab_type="text" id="F25PNc1GWV0R"
# ## Training the full agent
# + [markdown] colab_type="text" id="yRdwvHMQVoBB"
# To simplify collecting and storing experiences, you can also directly use Acme's `EnvironmentLoop` which runs the environment loop for a specified number of episodes. Each episode is itself a loop which interacts first with the environment to get an observation and then give that observation to the agent in order to retrieve an action. Upon termination of an episode a new episode will be started. If the number of episodes is not given then this will interact with the environment infinitely.
# + colab={} colab_type="code" id="evoLYDDagdv3"
# This may be necessary if any of the episodes were cancelled above.
adder.reset()
# We also want to make sure the logger doesn't write to disk because that can
# cause issues in colab on occasion.
logger = loggers.TerminalLogger(time_delta=10.)
# + colab={} colab_type="code" id="5Gu5D2obWTBc"
loop = environment_loop.EnvironmentLoop(environment, d4pg_agent, logger=logger)
loop.run(num_episodes=50)
# + [markdown] colab_type="text" id="EAn543WQfLNw"
# ## Evaluate the D4PG agent
#
# We can now evaluate the agent. Note that this will use the noisy behavior policy, and so won't quite be optimal. If we wanted to be absolutely precise we could easily replace this with the noise-free policy. Note that the optimal policy can get about 1000 reward in this environment. D4PG should generally get to that within 50-100 learner steps. We've cut it off at 50 and not dropped the behavior noise just to simplify this tutorial.
# -
agent.
# + colab={} colab_type="code" id="2__mFiraWND1"
# Run the actor in the environment for desired number of steps.
frames = []
num_steps = 1000
timestep = environment.reset()
for _ in range(num_steps):
frames.append(render(environment))
action = d4pg_agent.select_action(timestep.observation)
timestep = environment.step(action)
# Save video of the behaviour.
display_video(np.array(frames))
# -
| examples/panda_lift.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# https://www.fmf.uni-lj.si/~jazbinsek/Praktikum5/gammaenergies.pdf
#
# 
# Expected peaks (keV):
#
# U-238:
#
# 2204.2 (5) Bi-214
#
# 1764.5 (15.8) Bi-214
#
# 1238.1 (5.9) Bi-214
#
# 1120.3 (15.1) Bi-214
#
# 609.3*(46.3) Bi-214
#
# 352.0*(37.2) Pb-214
#
# 295.2 (19.2) Pb-214
#
# 92.6*(5.4) Th-234
#
# 63.3 (3.8) Th-234
#
# 46.5*(3.9) Pb-21
#
#
# U-235:
#
# 401.8 (6.5) Rn-219
#
# 271.2*(10.6) Rn-219
#
# 269.5*(13.6) Ra-223
#
# 236*(11.5) Th-227
#
# 210.6 (11.3) Th-227
#
# 185.7*(54) U-235
#
# Return table of channel number vs energy
# +
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import curve_fit
csv = np.genfromtxt('uranium_test_2019-02-19_D3S.csv', delimiter= ",").T
# -
summed = np.sum(csv, axis=1)
plt.plot(summed)
plt.yscale('log')
plt.show()
def fitFunc(x, a, m, s, c):
return a * np.exp(-(x - m)**2 / (2 * s**2)) + c
def linBgFitFunc(x, a, m, s, c, b):
return a * np.exp(-(x - m)**2 / (2 * s**2)) + c + b * x
def find(xSlice, xshift, trymax=20, trymu=200, trysig=100, trybg=5):
xmu = np.mean(xSlice)
xsig = np.std(xSlice)
xxdata = range(len(xSlice))
trydata = fitFunc(xSlice, np.max(xSlice), xmu, xsig, np.max(xSlice) + 50)
p0 = [trymax,trymu,trysig,trybg]
xpopt, xpcov = curve_fit(fitFunc, xxdata, xSlice, p0)
print(xpopt)
#plt.plot(xxdata, xSlice)
#plt.plot(xxdata, fitFunc(xxdata, *xpopt))
#plt.plot(int(xpopt[1]), fitFunc(xxdata, *xpopt)[int(xpopt[1])], 'ro')
xchannel = xshift + int(xpopt[1])
return xchannel
#plt.show()
def linBgFind(xSlice, xshift, trymax=20, trymu=200, trysig=100, trybg=5, trylin=-20):
xmu = np.mean(xSlice)
xsig = np.std(xSlice)
xxdata = range(len(xSlice))
#trydata = fitFunc(xSlice, np.max(xSlice), xmu, xsig, np.max(xSlice) + 50)
p0 = [trymax,trymu,trysig,trybg, trylin]
xpopt, xpcov = curve_fit(linBgFitFunc, xxdata, xSlice, p0)
print(xpopt)
#plt.plot(xxdata, xSlice)
#plt.plot(xxdata, fitFunc(xxdata, *xpopt))
#plt.plot(int(xpopt[1]), fitFunc(xxdata, *xpopt)[int(xpopt[1])], 'ro')
xchannel = xshift + int(xpopt[1])
return xchannel
def showFindFit(xSlice, xshift, trymax=20, trymu=200, trysig=100, trybg=5, lin=1):
xmu = np.mean(xSlice)
xsig = np.std(xSlice)
xxdata = range(len(xSlice))
#trydata = linBgFitFunc(1, np.max(xSlice), xmu, xsig, np.max(xSlice) + 50, lin)
p0 = [trymax,trymu,trysig,trybg, lin]
xpopt, xpcov = curve_fit(linBgFitFunc, xxdata, xSlice, p0)
print(xpopt)
#plt.plot(xxdata, xSlice)
#plt.plot(xxdata, fitFunc(xxdata, *xpopt))
#plt.plot(int(xpopt[1]), fitFunc(xxdata, *xpopt)[int(xpopt[1])], 'ro')
xchannel = xshift + int(xpopt[1])
return linBgFitFunc(xxdata, *xpopt)
#the plan is to just automate this block of code
Bi_shift = 2000
Bi_range = 400
Bi_slice = summed[Bi_shift:Bi_shift+Bi_range]
plt.plot(Bi_slice)
Bi_find = find(Bi_slice, Bi_shift)
print(Bi_find)
plt.plot(Bi_find- Bi_shift, Bi_slice[Bi_find- Bi_shift], 'ro')
plt.show()
#This block is redundant to the one above but we can see the fitting function here
Bi_mu = np.mean(Bi_slice)
Bi_sig = np.std(Bi_slice)
Bi_xdata = range(len(Bi_slice))
trydata = fitFunc(Bi_slice, np.max(Bi_slice), Bi_mu, Bi_sig, np.max(Bi_slice) + 50)
p0 = [Bi_mu,Bi_mu,100,5]
Bi_popt, Bi_pcov = curve_fit(fitFunc, Bi_xdata, Bi_slice, p0)
plt.plot(Bi_xdata, Bi_slice)
plt.plot(Bi_xdata, fitFunc(Bi_xdata, *Bi_popt))
plt.plot(int(Bi_popt[1]), fitFunc(Bi_xdata, *Bi_popt)[int(Bi_popt[1])], 'ro')
Bi_channel = Bi_shift + int(Bi_popt[1])
plt.show()
# +
Bi2_shift = 1600
Bi2_range = 300
Bi2_slice = summed[Bi2_shift:Bi2_shift+Bi2_range]
plt.plot(Bi2_slice)
Bi2_find = find(Bi2_slice, Bi2_shift)
print(Bi2_find)
plt.plot(Bi2_find-Bi2_shift, Bi2_slice[Bi2_find-Bi2_shift], 'ro')
plt.show()
# +
Bi3_shift = 1100
Bi3_range = 400
Bi3_slice = summed[Bi3_shift:Bi3_shift+Bi3_range]
plt.plot(Bi3_slice)
Bi3_find = find(Bi3_slice, Bi3_shift)
print(Bi3_find)
plt.plot(Bi3_find-Bi3_shift, Bi3_slice[Bi3_find-Bi3_shift], 'ro')
plt.show()
# +
Bi4_shift = 900
Bi4_range = 200
Bi4_slice = summed[Bi4_shift:Bi4_shift+Bi4_range]
plt.plot(Bi4_slice)
Bi4_find = find(Bi4_slice, Bi4_shift)
print(Bi4_find)
plt.plot(Bi4_find-Bi4_shift, Bi4_slice[Bi4_find-Bi4_shift], 'ro')
plt.show()
# +
Bi5_shift = 540
Bi5_range = 100
Bi5_slice = summed[Bi5_shift:Bi5_shift+Bi5_range]
plt.plot(Bi5_slice)
Bi5_find = find(Bi5_slice, Bi5_shift)
print(Bi5_find)
plt.plot(Bi5_find-Bi5_shift, Bi5_slice[Bi5_find-Bi5_shift], 'ro')
plt.show()
# +
Pb_shift = 250
Pb_range = 130
Pb_slice = summed[Pb_shift:Pb_shift+Pb_range]
plt.plot(Pb_slice)
Pb_find = linBgFind(Pb_slice, Pb_shift, 1200, 60, 80, 20)
#print(Pb_find)
plt.plot(Pb_find-Pb_shift, Pb_slice[Pb_find-Pb_shift], 'ro')
plt.plot(showFindFit(Pb_slice, Pb_shift, 1200, 60, 30, 500, -30))
plt.show()
# +
Th_shift = 60
Th_range = 150
Th_slice = summed[Th_shift:Th_shift+Th_range]
plt.plot(Th_slice)
Th_find = linBgFind(Th_slice, Th_shift, 1200, 60, 80, 20)
#print(Th_find)
plt.plot(Th_find-Th_shift, Th_slice[Th_find-Th_shift], 'ro')
#plt.plot(showFindFit(Th_slice, Th_shift, 1200, 60, 30, 500, -30))
plt.show()
# +
plt.plot(summed)
plt.plot(Bi_find, summed[Bi_find], 'ro') #2204.2 UNUSED
plt.plot(Bi2_find, summed[Bi2_find], 'bo') #1120.3
plt.plot(Bi3_find, summed[Bi3_find], 'r.') # UNUSED
plt.plot(Bi4_find, summed[Bi4_find], 'yo') #609.3
plt.plot(Bi5_find, summed[Bi5_find], 'mo') #352
plt.plot(Pb_find, summed[Pb_find], 'ko') #185.7
plt.plot(Th_find, summed[Th_find], 'co') #92.6
print(Bi2_find)
plt.yscale('log')
plt.show()
# +
channels = [Bi2_find, Bi4_find, Bi5_find, Pb_find, Th_find]
Th_channels = [390, 925, 3666]
#channels = channels + Th_channels
energies = [1120.3, 609.3, 352, 185.7, 92.6]
Th_energies = [238.6, 583.1, 2614.7]
plt.plot(channels, energies, 'ro')
plt.plot(Th_channels, Th_energies, 'bo')
plt.show()
print(channels)
# +
combChannels = channels + Th_channels
combEnergies = energies + Th_energies
plt.plot(combChannels, combEnergies, 'r.')
def linfit(x, m, b):
return m*x + b
def polyfit(x, m, b, r):
return r * x*x + m*x + b
p0 = [1, .6, 2]
xpopt, xpcov = curve_fit(polyfit, combChannels, combEnergies, p0)
print(xpopt)
plt.plot(polyfit(range(max(combChannels)), *xpopt))
plt.show()
# -
plt.plot([i*i*0.58 + 4.97*i + 3.5e-5 for i in range(len(summed))], [summed[i] for i in range(len(summed))])
plt.yscale('log')
plt.show()
i=len(summed)
i*i*0.58 + 4.97*i + 3.5e-5
| calibration/Uranium.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 付録2.2 Numpy入門
# ### NumPyの定義
# #### ライブラリインポート
# +
# ライブラリのインポート
import numpy as np
# numpyの浮動小数点の表示精度
np.set_printoptions(suppress=True, precision=4)
# -
# #### NumPyデータ定義
# +
# NumPyデータ定義
# 1次元配列変数の定義
a = np.array([1, 2, 3, 4, 5, 6, 7])
# 2次元配列変数の定義
b = np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9],[10,11,12]])
# -
# #### 内容表示
# +
# 内容表示
# 1次元配列変数の表示
print(a)
# 2次元配列変数の表示
print(b)
# -
# #### 要素数の確認
# +
# 要素数の確認
# 1次元配列変数の表示
print(a.shape)
# 2次元配列変数の表示
print(b.shape)
# len関数の利用
print(len(a))
print(len(b))
# -
# ### NumPyの操作
# #### 特定列の抽出
# +
# 特定列の抽出
# 0列
c = b[:,0]
print(c)
# [0,1]列
d = b[:,:2]
print(d)
# -
# #### reshape関数
# +
# reshape関数
l = np.array(range(12))
print(l)
# 3行4列に変換
m = l.reshape((3,4))
print(m)
# -
# #### 統計関数
# +
# 統計関数
print(f'元の変数: {a}')
a_sum = np.sum(a)
print(f'和: {a_sum}')
a_mean = np.mean(a)
print(f'平均: {a_mean}')
a_max = np.max(a)
print(f'最大値: {a_max}')
a_min = a.min()
print(f'最小値: {a_min}')
# -
# #### NumPy変数間の演算
# 二つのNumPy配列 ytとypの準備
yt = np.array([1, 1, 0, 1, 0, 1, 1, 0, 1, 1])
yp = np.array([1, 1, 0, 1, 0, 1, 1, 1, 1, 1])
print(yt)
print(yp)
# 配列の各要素を同時に比較する
w = (yt == yp)
print(w)
# 更にこの結果にsum関数をかける
print(w.sum())
# #### ブロードキャスト機能
# ブロードキャスト機能
print(a)
c = (a - np.min(a)) / (np.max(a) - np.min(a))
print(c)
# #### 数値配列の生成
# 数値配列の生成
x = np.linspace(-5, 5, 11)
print(x)
| notebooks/l2_01_numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="xBJWMeO0r-qy"
# # 이미지 준비
# + id="OzdgttRvp6F3" language="bash"
# apt install -y imagemagick
# [ ! -f flower_photos_300x200_small_train_test.zip ]&& wget https://raw.githubusercontent.com/Finfra/AI_CNN_RNN/main/data/flower_photos_300x200_small_train_test.zip
#
# rm -rf __MACOSX
# rm -rf flowers
# unzip -q flower_photos_300x200_small_train_test.zip
# mv flower_photos_300x200_small_train_test flowers
#
# cd flowers
# files=$(find |grep "\.jpg$\|\.png$")
# for i in $files; do
# # convert $i -quiet -resize 300x200^ -gravity center -extent 300x200 -colorspace Gray ${i%.*}.png
# convert $i -quiet -resize 300x200^ -gravity center -extent 300x200 -define png:color-type=2 ${i%.*}.png
#
# # identify ${i%.*}.png
# rm -f $i
# done
#
# find .|grep .DS_Store|xargs rm -f
# find .|head -n 10
#
# + [markdown] id="Dhow5w0Cyhu9"
# # CNN Example FromFile by Keras
#
# + id="bIcSLm6Cyhu-"
from __future__ import absolute_import, division, print_function, unicode_literals
import tensorflow as tf
from tensorflow.keras import datasets, layers, models
from tensorflow.keras.utils import to_categorical
import numpy as np
# + [markdown] id="sS-n7IiW7I0P"
# # 해당 Path의 이미지 Array로 리턴
# + id="z3_tr9TGyhvC"
from os import listdir
from os.path import isfile, join
from pylab import *
from numpy import *
def getFolder(thePath,isFile=True):
return [f for f in listdir(thePath) if isFile == isfile(join(thePath, f)) ]
def getImagesAndLabels(tPath,isGray=False):
labels=getFolder(tPath,False)
tImgDic={f:getFolder(join(tPath,f)) for f in labels}
tImages,tLabels=None,None
ks=sorted(list(tImgDic.keys()))
oh=np.identity(len(ks))
for label in tImgDic.keys():
for image in tImgDic[label]:
le=np.array([float(label)],ndmin=1)
img=imread(join(tPath,label,image))
if isGray:
img=img.reshape(img.shape+(1,))
img=img.reshape((1,) +img.shape)
if tImages is None:
tImages, tLabels =img, le
else:
tImages,tLabels = np.append(tImages,img,axis=0), np.append(tLabels,le ,axis=0)
return (tImages,to_categorical(tLabels) )
# + id="KERgFFAlQZOm"
w=300
h=200
color=3
# + id="W4s0Itg07vYk"
tPath='flowers/train'
train_images,train_labels=getImagesAndLabels(tPath)
tPath='flowers/test'
test_images,test_labels=getImagesAndLabels(tPath)
train_images, test_images = train_images / 255.0, test_images / 255.0
print(f"Shape of Train_images = {train_images.shape} , Shape of Train_labels = {train_labels.shape}")
# + id="Kk0CsU6LyhvI"
model = models.Sequential()
model.add(layers.Conv2D(96, (13, 13), activation='relu', padding='same', input_shape=(h,w, color)))
model.add(layers.Dropout(0.2))
model.add(layers.MaxPooling2D((3, 3)))
model.add(layers.Conv2D(64, (7, 7), activation='relu', padding='same'))
model.add(layers.Dropout(0.2))
model.add(layers.MaxPooling2D((3, 3)))
model.add(layers.Conv2D(64, (5, 5), activation='relu', padding='same'))
model.add(layers.Dropout(0.2))
model.add(layers.BatchNormalization())
model.add(layers.MaxPooling2D((3, 3)))
model.add(layers.Conv2D(64, (3, 3), activation='relu', padding='same'))
model.add(layers.MaxPooling2D((3, 3)))
model.add(layers.Flatten())
model.add(layers.Dense(200, activation='tanh',kernel_regularizer='l1'))
model.add(layers.Dense(2, activation='softmax'))
model.summary()
# + id="sjW_BUuXAI5G" language="bash"
# [ ! -d /content/ckpt/ ] && mkdir /content/ckpt/
#
#
# + id="2xmi4R6tBGnp"
epochs = 40
batch_size = 100
from tensorflow.keras.callbacks import ModelCheckpoint
filename = f'/contents/ckpt/checkpoint-epoch-{epochs}-batch-{batch_size}-trial-001.h5'
checkpoint = ModelCheckpoint(filename,
monitor='val_loss',
verbose=1,
save_best_only=True,
mode='auto'
)
# + id="Z8n7Q1jfyhvM"
model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
hist=model.fit(train_images,
train_labels,
batch_size=batch_size,
epochs=epochs,
validation_data=(test_images,test_labels),
callbacks=[checkpoint]
)
# + [markdown] id="wAEJQV6x5aLz"
# # Train history
# + id="nqLZ4c6F6zMA"
print('## training loss and acc ##')
fig, loss_ax = plt.subplots()
acc_ax = loss_ax.twinx()
loss_ax.plot(hist.history['loss'], 'y', label='train loss')
loss_ax.set_xlabel('epoch')
loss_ax.set_ylabel('loss')
loss_ax.legend(loc='upper left')
acc_ax.plot(hist.history['accuracy'], 'b', label='train acc')
acc_ax.set_ylabel('accuracy')
acc_ax.legend(loc='upper right')
plt.show()
# + [markdown] id="rMpYl9BByhvP"
#
# # Result
# + id="5k2PbbcG62d0"
score = model.evaluate(test_images, test_labels, verbose=0, batch_size=32)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
| 04.CNN/CNN_fromFile_flower_checkpoints.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import csv
# 1. you have to open a file in correct mode before you write or read to file
fileName='translator.txt'
accessMode='r'
fss=[]
# open myfile
with open(fileName,accessMode) as myfile:
allRowList=csv.reader(myfile)
for currentRow in allRowList:
fss.append(currentRow)
print(currentRow)
print(fss)
# -
fss
nss=[]
for fs in fss:
for f in fs:
ns=f.split()
nss.append(ns)
nss
mss=[['LOL','laughing out loud'],
['ROFL','Rolling on Floor Laughing']]
k=input("Enter the the sentence with short forms \n")
k="So funny LOL"
ls=k.split()
# ls
# +
# first looping through words of input sentence
for l in ls:
# print(l)
for ms in mss:
# print(ms[1])
for m in ms:
# print(m)
if l==m:
Acronymfound=True
print("\n Match Found WOW!")
print(l, "is a soruce match")
print(m, "is a replacement match")
print(ls.index(l))
print("\n In given sentence with acronym \" " + ls[ls.index(l)] + " \" at location " + str( ls.index(l)) )
print("\n The acronym \" " + ms[ms.index(m)] + " \" at location " + str( ms.index(m)) )
print(ms.index(m))
if (ms.index(m)==1):
print ("\n already transformed")
break
else:
print(ms[ms.index(m)+1])
print(type(ms[ms.index(m)+1]))
# ls[ls.index(l)]=ms[ms.index(m)+1]
(ls[ls.index(l)])=(ms[ms.index(m)+1])
print(l)
print(m)
else:
print(" \n\nmatch not found")
print( l + " is not equal to " + m)
print(ls)
outSentence=' '.join(ls)
outSentence
# -
# +
# input is list ls
# first looping through words of input sentence
for l in ls:
# print(l)
for ms in mss:
# print(ms[1])
for m in ms:
# print(m)
if l==m:
Acronymfound=True
print("\n Match Found WOW!")
print(l, "is a soruce match")
print(m, "is a replacement match")
print(ls.index(l))
print("\n In given sentence with acronym \" " + ls[ls.index(l)] + " \" at location " + str( ls.index(l)) )
print("\n The acronym \" " + ms[ms.index(m)] + " \" at location " + str( ms.index(m)) )
print(ms.index(m))
if (ms.index(m)==1):
print ("\n already transformed")
break
else:
print(ms[ms.index(m)+1])
print(type(ms[ms.index(m)+1]))
# ls[ls.index(l)]=ms[ms.index(m)+1]
(ls[ls.index(l)])=(ms[ms.index(m)+1])
print(l)
print(m)
else:
break
print(" \n\nmatch not found")
print( l + " is not equal to " + m)
#
print(ls)
outSentence=' '.join(ls)
print(outSentence)
# -
outSentence=' '.join(ls)
outSentence
outSentence=' '.join(ls)
outSentence
| challenge_15/text_converter_15-Copy2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="fluF3_oOgkWF"
# ##### Copyright 2020 The TensorFlow Authors.
# + cellView="form" id="AJs7HHFmg1M9" tags=[]
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="jYysdyb-CaWM"
# # Simple audio recognition: Recognizing keywords
# + [markdown] id="CNbqmZy0gbyE"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/audio/simple_audio">
# <img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
# View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/audio/simple_audio.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
# Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/audio/simple_audio.ipynb">
# <img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
# View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/audio/simple_audio.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="SPfDNFlb66XF"
# This tutorial demonstrates how to preprocess audio files in the WAV format and build and train a basic <a href="https://en.wikipedia.org/wiki/Speech_recognition" class="external">automatic speech recognition</a> (ASR) model for recognizing ten different words. You will use a portion of the [Speech Commands dataset](https://www.tensorflow.org/datasets/catalog/speech_commands) (<a href="https://arxiv.org/abs/1804.03209" class="external">Warden, 2018</a>), which contains short (one-second or less) audio clips of commands, such as "down", "go", "left", "no", "right", "stop", "up" and "yes".
#
# Real-world speech and audio recognition <a href="https://ai.googleblog.com/search/label/Speech%20Recognition" class="external">systems</a> are complex. But, like [image classification with the MNIST dataset](../quickstart/beginner.ipynb), this tutorial should give you a basic understanding of the techniques involved.
# + [markdown] id="Go9C3uLL8Izc"
# ## Setup
#
# Import necessary modules and dependencies. Note that you'll be using <a href="https://seaborn.pydata.org/" class="external">seaborn</a> for visualization in this tutorial.
# + id="dzLKpmZICaWN" tags=[]
import os
import pathlib
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
import tensorflow as tf
from tensorflow.keras import layers
from tensorflow.keras import models
from IPython import display
# Set the seed value for experiment reproducibility.
seed = 42
tf.random.set_seed(seed)
np.random.seed(seed)
# + [markdown] id="yR0EdgrLCaWR"
# ## Import the mini Speech Commands dataset
#
# To save time with data loading, you will be working with a smaller version of the Speech Commands dataset. The [original dataset](https://www.tensorflow.org/datasets/catalog/speech_commands) consists of over 105,000 audio files in the <a href="https://www.aelius.com/njh/wavemetatools/doc/riffmci.pdf" class="external">WAV (Waveform) audio file format</a> of people saying 35 different words. This data was collected by Google and released under a CC BY license.
#
# Download and extract the `mini_speech_commands.zip` file containing the smaller Speech Commands datasets with `tf.keras.utils.get_file`:
# + id="2-rayb7-3Y0I" tags=[]
DATASET_PATH = 'data/mini_speech_commands'
data_dir = pathlib.Path(DATASET_PATH)
if not data_dir.exists():
tf.keras.utils.get_file(
'mini_speech_commands.zip',
origin="http://storage.googleapis.com/download.tensorflow.org/data/mini_speech_commands.zip",
extract=True,
cache_dir='.', cache_subdir='data')
# + [markdown] id="BgvFq3uYiS5G"
# The dataset's audio clips are stored in eight folders corresponding to each speech command: `no`, `yes`, `down`, `go`, `left`, `up`, `right`, and `stop`:
# + id="70IBxSKxA1N9" tags=[]
commands = np.array(tf.io.gfile.listdir(str(data_dir)))
commands = commands[commands != 'README.md']
print('Commands:', commands)
# + [markdown] id="aMvdU9SY8WXN"
# Extract the audio clips into a list called `filenames`, and shuffle it:
# + id="hlX685l1wD9k" tags=[]
filenames = tf.io.gfile.glob(str(data_dir) + '/*/*')
filenames = tf.random.shuffle(filenames)
num_samples = len(filenames)
print('Number of total examples:', num_samples)
print('Number of examples per label:',
len(tf.io.gfile.listdir(str(data_dir/commands[0]))))
print('Example file tensor:', filenames[0])
# + [markdown] id="9vK3ymy23MCP"
# Split `filenames` into training, validation and test sets using a 80:10:10 ratio, respectively:
# + id="Cv_wts-l3KgD" tags=[]
train_files = filenames[:6400]
val_files = filenames[6400: 6400 + 800]
test_files = filenames[-800:]
print('Training set size', len(train_files))
print('Validation set size', len(val_files))
print('Test set size', len(test_files))
# + [markdown] id="g2Cj9FyvfweD"
# ## Read the audio files and their labels
# + [markdown] id="j1zjcWteOcBy"
# In this section you will preprocess the dataset, creating decoded tensors for the waveforms and the corresponding labels. Note that:
#
# - Each WAV file contains time-series data with a set number of samples per second.
# - Each sample represents the <a href="https://en.wikipedia.org/wiki/Amplitude" class="external">amplitude</a> of the audio signal at that specific time.
# - In a <a href="https://en.wikipedia.org/wiki/Audio_bit_depth" class="external">16-bit</a> system, like the WAV files in the mini Speech Commands dataset, the amplitude values range from -32,768 to 32,767.
# - The <a href="https://en.wikipedia.org/wiki/Sampling_(signal_processing)#Audio_sampling" class="external">sample rate</a> for this dataset is 16kHz.
#
# The shape of the tensor returned by `tf.audio.decode_wav` is `[samples, channels]`, where `channels` is `1` for mono or `2` for stereo. The mini Speech Commands dataset only contains mono recordings.
# + id="d16bb8416f90" tags=[]
test_file = tf.io.read_file(DATASET_PATH+'/down/0a9f9af7_nohash_0.wav')
test_audio, _ = tf.audio.decode_wav(contents=test_file)
test_audio.shape
# + [markdown] id="e6bb8defd2ef"
# Now, let's define a function that preprocesses the dataset's raw WAV audio files into audio tensors:
# + id="9PjJ2iXYwftD" tags=[]
def decode_audio(audio_binary):
# Decode WAV-encoded audio files to `float32` tensors, normalized
# to the [-1.0, 1.0] range. Return `float32` audio and a sample rate.
audio, _ = tf.audio.decode_wav(contents=audio_binary)
# Since all the data is single channel (mono), drop the `channels`
# axis from the array.
return tf.squeeze(audio, axis=-1)
# + [markdown] id="GPQseZElOjVN"
# Define a function that creates labels using the parent directories for each file:
#
# - Split the file paths into `tf.RaggedTensor`s (tensors with ragged dimensions—with slices that may have different lengths).
# + id="8VTtX1nr3YT-" tags=[]
def get_label(file_path):
parts = tf.strings.split(
input=file_path,
sep=os.path.sep)
# Note: You'll use indexing here instead of tuple unpacking to enable this
# to work in a TensorFlow graph.
return parts[-2]
# + [markdown] id="E8Y9w_5MOsr-"
# Define another helper function—`get_waveform_and_label`—that puts it all together:
#
# - The input is the WAV audio filename.
# - The output is a tuple containing the audio and label tensors ready for supervised learning.
# + id="WdgUD5T93NyT" tags=[]
def get_waveform_and_label(file_path):
label = get_label(file_path)
audio_binary = tf.io.read_file(file_path)
waveform = decode_audio(audio_binary)
return waveform, label
# + [markdown] id="nvN8W_dDjYjc"
# Build the training set to extract the audio-label pairs:
#
# - Create a `tf.data.Dataset` with `Dataset.from_tensor_slices` and `Dataset.map`, using `get_waveform_and_label` defined earlier.
#
# You'll build the validation and test sets using a similar procedure later on.
# -
type(files_ds)
# + id="0SQl8yXl3kNP" tags=[]
AUTOTUNE = tf.data.AUTOTUNE
files_ds = tf.data.Dataset.from_tensor_slices(train_files)
waveform_ds = files_ds.map(
map_func=get_waveform_and_label,
num_parallel_calls=AUTOTUNE)
# -
# + [markdown] id="voxGEwvuh2L7"
# Let's plot a few audio waveforms:
# + id="8yuX6Nqzf6wT" tags=[]
rows = 3
cols = 3
n = rows * cols
fig, axes = plt.subplots(rows, cols, figsize=(10, 12))
for i, (audio, label) in enumerate(waveform_ds.take(n)):
r = i // cols
c = i % cols
ax = axes[r][c]
ax.plot(audio.numpy())
ax.set_yticks(np.arange(-1.2, 1.2, 0.2))
label = label.numpy().decode('utf-8')
ax.set_title(label)
plt.show()
# + [markdown] id="EWXPphxm0B4m"
# ## Convert waveforms to spectrograms
#
# The waveforms in the dataset are represented in the time domain. Next, you'll transform the waveforms from the time-domain signals into the time-frequency-domain signals by computing the <a href="https://en.wikipedia.org/wiki/Short-time_Fourier_transform" class="external">short-time Fourier transform (STFT)</a> to convert the waveforms to as <a href="https://en.wikipedia.org/wiki/Spectrogram" clas="external">spectrograms</a>, which show frequency changes over time and can be represented as 2D images. You will feed the spectrogram images into your neural network to train the model.
#
# A Fourier transform (`tf.signal.fft`) converts a signal to its component frequencies, but loses all time information. In comparison, STFT (`tf.signal.stft`) splits the signal into windows of time and runs a Fourier transform on each window, preserving some time information, and returning a 2D tensor that you can run standard convolutions on.
#
# Create a utility function for converting waveforms to spectrograms:
#
# - The waveforms need to be of the same length, so that when you convert them to spectrograms, the results have similar dimensions. This can be done by simply zero-padding the audio clips that are shorter than one second (using `tf.zeros`).
# - When calling `tf.signal.stft`, choose the `frame_length` and `frame_step` parameters such that the generated spectrogram "image" is almost square. For more information on the STFT parameters choice, refer to <a href="https://www.coursera.org/lecture/audio-signal-processing/stft-2-tjEQe" class="external">this Coursera video</a> on audio signal processing and STFT.
# - The STFT produces an array of complex numbers representing magnitude and phase. However, in this tutorial you'll only use the magnitude, which you can derive by applying `tf.abs` on the output of `tf.signal.stft`.
# + id="_4CK75DHz_OR" tags=[]
def get_spectrogram(waveform):
# Zero-padding for an audio waveform with less than 16,000 samples.
input_len = 16000
waveform = waveform[:input_len]
zero_padding = tf.zeros(
[16000] - tf.shape(waveform),
dtype=tf.float32)
# Cast the waveform tensors' dtype to float32.
waveform = tf.cast(waveform, dtype=tf.float32)
# Concatenate the waveform with `zero_padding`, which ensures all audio
# clips are of the same length.
equal_length = tf.concat([waveform, zero_padding], 0)
# Convert the waveform to a spectrogram via a STFT.
spectrogram = tf.signal.stft(
equal_length, frame_length=255, frame_step=128)
# Obtain the magnitude of the STFT.
spectrogram = tf.abs(spectrogram)
# Add a `channels` dimension, so that the spectrogram can be used
# as image-like input data with convolution layers (which expect
# shape (`batch_size`, `height`, `width`, `channels`).
spectrogram = spectrogram[..., tf.newaxis]
return spectrogram
# + [markdown] id="5rdPiPYJphs2"
# Next, start exploring the data. Print the shapes of one example's tensorized waveform and the corresponding spectrogram, and play the original audio:
# + id="4Mu6Y7Yz3C-V" tags=[]
for waveform, label in waveform_ds.take(1):
label = label.numpy().decode('utf-8')
spectrogram = get_spectrogram(waveform)
print('Label:', label)
print('Waveform shape:', waveform.shape)
print('Spectrogram shape:', spectrogram.shape)
print('Audio playback')
display.display(display.Audio(waveform, rate=16000))
# + [markdown] id="xnSuqyxJ1isF"
# Now, define a function for displaying a spectrogram:
# + id="e62jzb36-Jog" tags=[]
def plot_spectrogram(spectrogram, ax):
if len(spectrogram.shape) > 2:
assert len(spectrogram.shape) == 3
spectrogram = np.squeeze(spectrogram, axis=-1)
# Convert the frequencies to log scale and transpose, so that the time is
# represented on the x-axis (columns).
# Add an epsilon to avoid taking a log of zero.
log_spec = np.log(spectrogram.T + np.finfo(float).eps)
height = log_spec.shape[0]
width = log_spec.shape[1]
X = np.linspace(0, np.size(spectrogram), num=width, dtype=int)
Y = range(height)
ax.pcolormesh(X, Y, log_spec)
# + [markdown] id="baa5c91e8603"
# Plot the example's waveform over time and the corresponding spectrogram (frequencies over time):
# + id="d2_CikgY1tjv" tags=[]
fig, axes = plt.subplots(2, figsize=(12, 8))
timescale = np.arange(waveform.shape[0])
axes[0].plot(timescale, waveform.numpy())
axes[0].set_title('Waveform')
axes[0].set_xlim([0, 16000])
plot_spectrogram(spectrogram.numpy(), axes[1])
axes[1].set_title('Spectrogram')
plt.show()
# + [markdown] id="GyYXjW07jCHA"
# Now, define a function that transforms the waveform dataset into spectrograms and their corresponding labels as integer IDs:
# + id="43IS2IouEV40" tags=[]
def get_spectrogram_and_label_id(audio, label):
spectrogram = get_spectrogram(audio)
label_id = tf.argmax(label == commands)
return spectrogram, label_id
# + [markdown] id="cf5d5b033a45"
# Map `get_spectrogram_and_label_id` across the dataset's elements with `Dataset.map`:
# + id="yEVb_oK0oBLQ" tags=[]
spectrogram_ds = waveform_ds.map(
map_func=get_spectrogram_and_label_id,
num_parallel_calls=AUTOTUNE)
# + [markdown] id="6gQpAAgMnyDi"
# Examine the spectrograms for different examples of the dataset:
# + id="QUbHfTuon4iF" tags=[]
rows = 3
cols = 3
n = rows*cols
fig, axes = plt.subplots(rows, cols, figsize=(10, 10))
for i, (spectrogram, label_id) in enumerate(spectrogram_ds.take(n)):
r = i // cols
c = i % cols
ax = axes[r][c]
plot_spectrogram(spectrogram.numpy(), ax)
ax.set_title(commands[label_id.numpy()])
ax.axis('off')
plt.show()
# + [markdown] id="z5KdY8IF8rkt"
# ## Build and train the model
#
# Repeat the training set preprocessing on the validation and test sets:
# + id="10UI32QH_45b" tags=[]
def preprocess_dataset(files):
files_ds = tf.data.Dataset.from_tensor_slices(files)
output_ds = files_ds.map(
map_func=get_waveform_and_label,
num_parallel_calls=AUTOTUNE)
output_ds = output_ds.map(
map_func=get_spectrogram_and_label_id,
num_parallel_calls=AUTOTUNE)
return output_ds
# + id="HNv4xwYkB2P6" tags=[]
train_ds = spectrogram_ds
val_ds = preprocess_dataset(val_files)
test_ds = preprocess_dataset(test_files)
# + [markdown] id="assnWo6SB3lR"
# Batch the training and validation sets for model training:
# + id="UgY9WYzn61EX" tags=[]
batch_size = 64
train_ds = train_ds.batch(batch_size)
val_ds = val_ds.batch(batch_size)
# -
for thing in train_ds.take(1):
print(thing[0].shape, thing[1].shape)
# + [markdown] id="GS1uIh6F_TN9"
# Add `Dataset.cache` and `Dataset.prefetch` operations to reduce read latency while training the model:
# + id="fdZ6M-F5_QzY" tags=[]
train_ds = train_ds.cache().prefetch(AUTOTUNE)
val_ds = val_ds.cache().prefetch(AUTOTUNE)
# + [markdown] id="rwHkKCQQb5oW"
# For the model, you'll use a simple convolutional neural network (CNN), since you have transformed the audio files into spectrogram images.
#
# Your `tf.keras.Sequential` model will use the following Keras preprocessing layers:
#
# - `tf.keras.layers.Resizing`: to downsample the input to enable the model to train faster.
# - `tf.keras.layers.Normalization`: to normalize each pixel in the image based on its mean and standard deviation.
#
# For the `Normalization` layer, its `adapt` method would first need to be called on the training data in order to compute aggregate statistics (that is, the mean and the standard deviation).
# + id="ALYz7PFCHblP" tags=[]
for spectrogram, _ in spectrogram_ds.take(1):
input_shape = spectrogram.shape
print('Input shape:', input_shape)
num_labels = len(commands)
# Instantiate the `tf.keras.layers.Normalization` layer.
norm_layer = layers.Normalization()
# Fit the state of the layer to the spectrograms
# with `Normalization.adapt`.
norm_layer.adapt(data=spectrogram_ds.map(map_func=lambda spec, label: spec))
model = models.Sequential([
layers.Input(shape=input_shape),
# Downsample the input.
layers.Resizing(32, 32),
# Normalize.
norm_layer,
layers.Conv2D(32, 3, activation='relu'),
layers.Conv2D(64, 3, activation='relu'),
layers.MaxPooling2D(),
layers.Dropout(0.25),
layers.Flatten(),
layers.Dense(128, activation='relu'),
layers.Dropout(0.5),
layers.Dense(num_labels),
])
model.summary()
# + [markdown] id="de52e5afa2f3"
# Configure the Keras model with the Adam optimizer and the cross-entropy loss:
# + id="wFjj7-EmsTD-" tags=[]
model.compile(
optimizer=tf.keras.optimizers.Adam(),
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'],
)
# + [markdown] id="f42b9e3a4705"
# Train the model over 10 epochs for demonstration purposes:
# + id="ttioPJVMcGtq" tags=[]
EPOCHS = 10
history = model.fit(
train_ds,
validation_data=val_ds,
epochs=EPOCHS,
callbacks=tf.keras.callbacks.EarlyStopping(verbose=1, patience=2),
)
# + [markdown] id="gjpCDeQ4mUfS"
# Let's plot the training and validation loss curves to check how your model has improved during training:
# + id="nzhipg3Gu2AY" tags=[]
metrics = history.history
plt.plot(history.epoch, metrics['loss'], metrics['val_loss'])
plt.legend(['loss', 'val_loss'])
plt.show()
# + [markdown] id="5ZTt3kO3mfm4"
# ## Evaluate the model performance
#
# Run the model on the test set and check the model's performance:
# + id="biU2MwzyAo8o" tags=[]
test_audio = []
test_labels = []
for audio, label in test_ds:
test_audio.append(audio.numpy())
test_labels.append(label.numpy())
test_audio = np.array(test_audio)
test_labels = np.array(test_labels)
# + id="ktUanr9mRZky" tags=[]
y_pred = np.argmax(model.predict(test_audio), axis=1)
y_true = test_labels
test_acc = sum(y_pred == y_true) / len(y_true)
print(f'Test set accuracy: {test_acc:.0%}')
# + [markdown] id="en9Znt1NOabH"
# ### Display a confusion matrix
#
# Use a <a href="https://developers.google.com/machine-learning/glossary#confusion-matrix" class="external">confusion matrix</a> to check how well the model did classifying each of the commands in the test set:
#
# + id="LvoSAOiXU3lL" tags=[]
confusion_mtx = tf.math.confusion_matrix(y_true, y_pred)
plt.figure(figsize=(10, 8))
sns.heatmap(confusion_mtx,
xticklabels=commands,
yticklabels=commands,
annot=True, fmt='g')
plt.xlabel('Prediction')
plt.ylabel('Label')
plt.show()
# + [markdown] id="mQGi_mzPcLvl"
# ## Run inference on an audio file
#
# Finally, verify the model's prediction output using an input audio file of someone saying "no". How well does your model perform?
# + id="zRxauKMdhofU" tags=[]
sample_file = data_dir/'no/01bb6a2a_nohash_0.wav'
sample_ds = preprocess_dataset([str(sample_file)])
for spectrogram, label in sample_ds.batch(1):
prediction = model(spectrogram)
plt.bar(commands, tf.nn.softmax(prediction[0]))
plt.title(f'Predictions for "{commands[label[0]]}"')
plt.show()
# + [markdown] id="VgWICqdqQNaQ"
# As the output suggests, your model should have recognized the audio command as "no".
# + [markdown] id="J3jF933m9z1J"
# ## Next steps
#
# This tutorial demonstrated how to carry out simple audio classification/automatic speech recognition using a convolutional neural network with TensorFlow and Python. To learn more, consider the following resources:
#
# - The [Sound classification with YAMNet](https://www.tensorflow.org/hub/tutorials/yamnet) tutorial shows how to use transfer learning for audio classification.
# - The notebooks from <a href="https://www.kaggle.com/c/tensorflow-speech-recognition-challenge/overview" class="external">Kaggle's TensorFlow speech recognition challenge</a>.
# - The
# <a href="https://codelabs.developers.google.com/codelabs/tensorflowjs-audio-codelab/index.html#0" class="external">TensorFlow.js - Audio recognition using transfer learning codelab</a> teaches how to build your own interactive web app for audio classification.
# - <a href="https://arxiv.org/abs/1709.04396" class="external">A tutorial on deep learning for music information retrieval</a> (Choi et al., 2017) on arXiv.
# - TensorFlow also has additional support for [audio data preparation and augmentation](https://www.tensorflow.org/io/tutorials/audio) to help with your own audio-based projects.
# - Consider using the <a href="https://librosa.org/" class="external">librosa</a> library—a Python package for music and audio analysis.
| capstone/notebooks/tutorial/simple_audio.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="unJwdVXqzSU7"
# # Counts, Frequencies, and Ngram Models
#
# Before you proceed, make sure to run the cell below.
# This will once again read in the cleaned up text files and store them as tokenized lists in the variables `hamlet`, `faustus`, and `mars`.
# If you get an error, make sure that you did the previous notebook and that this notebook is in a folder containing the files `hamlex_clean.txt`, `faustus_clean.txt`, and `mars_clean.txt` (which should be the case if you did the previous notebook).
# + colab={"base_uri": "https://localhost:8080/", "height": 199, "resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "headers": [["content-type", "application/javascript"]], "ok": true, "status": 200, "status_text": ""}}} id="1Iy5KNE6zSU7" outputId="0a48ebf9-d3a6-42fc-df7f-a8c505cfbf53"
from google.colab import files
#Import files
upload1 = files.upload()
upload2 = files.upload()
upload3 = files.upload()
hamlet_full = upload1['faustus_clean.txt'].decode('utf-8')
faustus_full = upload2['hamlet_clean.txt'].decode('utf-8')
mars_full = upload3['mars_clean.txt'].decode('utf-8')
# + id="BoKXUIAJ1awf"
import re
def tokenize(the_string):
"""Convert string to list of words"""
return re.findall(r"\w+", the_string)
# define a variable for each token list
hamlet = tokenize(hamlet_full)
faustus = tokenize(faustus_full)
mars = tokenize(mars_full)
# + [markdown] id="3Ha7kYZWzSU7"
# **Caution.**
# If you restart the kernel at any point, make sure to run all these previous cells again so that the variables `hamlet`, `faustus`, and `mars` are defined.
# + [markdown] id="pJtHy4zwzSU7"
# ## Counting words
#
# Python makes it very easy to count how often an element occurs in a list: the `collections` library provides a function `Counter` that does the counting for us.
# The `Counter` function takes as its only argument a list (like the ones produced by `re.findall` for tokenization).
# It then converts the list into a *Counter*.
# Here is what this looks like with a short example string.
# + colab={"base_uri": "https://localhost:8080/"} id="rFzd5xnLzSU7" outputId="3009b63d-c86f-4ee5-f434-02540175fb48"
import re
from collections import Counter # this allows us to use Counter instead of collections.Counter
test_string = "FTL is short for faster-than-light; we probably won't ever have space ships capable of FTL-travel."
# tokenize the string
tokens = re.findall(r"\w+", str.lower(test_string))
print("The list of tokens:", tokens)
# add an empty line
print()
# and now do the counting
counts = Counter(tokens)
print("Number of tokens for each word type:", counts)
# + [markdown] id="rcDv96NFzSU7"
# Let's take a quick peak at what the counts looks like for each text.
# We don't want to do this with something like `print(counts_hamlet)`, because the output would be so large that your browser might actually choke on it (it has happened to me sometimes).
# Instead, we will look at the 100 most common words.
# We can do this with the function `Counter.most_common`, which takes two arguments: a Counter, and a positive number.
# + colab={"base_uri": "https://localhost:8080/"} id="YhmGQPakzSU7" outputId="44cc3c82-456f-47ca-af17-8b08a29ae640"
from collections import Counter
# construct the counters
counts_hamlet = Counter(hamlet)
counts_faustus = Counter(faustus)
counts_mars = Counter(mars)
print("Most common Hamlet words:", Counter.most_common(counts_hamlet, 100))
print()
print("Most common Faustus words:", Counter.most_common(counts_faustus, 100))
print()
print("Most common John Carter words:", Counter.most_common(counts_mars, 100))
# + [markdown] id="GfRvTCwLzSU7"
# **Exercise.**
# The code below uses `import collections` instead of `from collections import Counter`.
# As you can test for yourself, the code now produces various errors.
# Fix the code so that the cell runs correctly.
# You must not change the `import` statement.
# + id="cvLRIWgazSU7"
import collections
# construct the counters
counts_hamlet = Counter(hamlet)
counts_faustus = Counter(faustus)
counts_mars = Counter(mars)
print("Most common Hamlet words:", Counter.most_common(counts_hamlet, 100))
print()
print("Most common Faustus words:", Counter.most_common(counts_faustus, 100))
print()
print("Most common John Carter words:", Counter.most_common(counts_mars, 100))
# + [markdown] id="xkBMKxhfzSU7"
# Python's output for `Counter.most_common` doesn't look too bad, but it is a bit convoluted.
# We can use the function `pprint` from the `pprint` library to have each word on its own line.
# The name *pprint* is short for *pretty-print*.
# + id="nsBd_2yxzSU7"
from pprint import pprint # we want to use pprint instead of pprint.pprint
from collections import Counter
# construct the counters
counts_hamlet = Counter(hamlet)
counts_faustus = Counter(faustus)
counts_mars = Counter(mars)
# we have to split lines now because pprint cannot take multiple arguments like print
print("Most common Hamlet words:")
pprint(Counter.most_common(counts_hamlet, 100))
print()
print("Most common Faustus words:")
pprint(Counter.most_common(counts_faustus, 100))
print()
print("Most common John Carter words:")
pprint(Counter.most_common(counts_mars, 100))
# + [markdown] id="E4UEP9w7zSU7"
# **Exercise.**
# What is the difference between the following two pieces of code?
# How do they differ in their output, and why?
# + colab={"base_uri": "https://localhost:8080/"} id="lUQc_Hj0zSU7" outputId="30cdc34b-284f-4917-dd25-f59b62cb3181"
from collections import Counter
counts = Counter(hamlet[:50])
print(counts)
# + colab={"base_uri": "https://localhost:8080/"} id="-6dGfBJezSU7" outputId="d89e3592-9bdf-4bb6-dacd-b28a1f0ab508"
from collections import Counter
count = Counter(hamlet)
print(Counter.most_common(count, 50))
# + [markdown] id="ejKIPswfzSU7"
# ## A problem
#
# If you look at the lists of 100 most common words for each text, you'll notice that they are fairly similar.
# For instance, all of them have *a*, *the*, and *to* among the most frequent ones.
# That's not a peculiarity of these few texts, it's a general property of English texts.
# This is because of **Zipf's law**: ranking words by their frequency, the n-th word will have a relative frequency of 1/n.
# So the most common word is twice as frequent as the second most common one, three times more frequent than the third most common one, and so on.
# As a result, a handful of words make up over 50% of all words in a text.
#
# Zipf's law means that word frequencies in a text give rise to a peculiar shape that we might call the Zipf dinosaur.
#
#
# A super-high neck, followed by a very long tail.
# For English texts, the distribution usually resembles the one below, and that's even though this graph only shows the most common words.
#
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 336} id="nhyUqgK97f3q" outputId="43d9337e-78db-4ee7-8a0d-2ecc5e7167ba"
from IPython.display import HTML
# Youtube
HTML('<iframe width="560" height="315" src="https://www.youtube.com/embed/fCn8zs912OE" frameborder="0" allowfullscreen></iframe>')
# + [markdown] id="5s7HUQsQ7o06"
# There is precious little variation between English texts with respect to which words are at the top.
# These common but uninformative words are called **stop words**.
# If we want to find any interesting differences between *Hamlet*, *Doctor Faustus*, and *Princess of Mars*, we have to filter out all these stop words.
# That's not something we can do by hand, but our existing box of tricks doesn't really seem to fit either.
# We could use a regular expression to delete all these words from the string before it even gets tokenized.
# But that's not the best solution:
#
# 1. A minor mistake in the regular expression might accidentally delete many things we want to keep.
# Odds are that this erroneous deletion would go unnoticed, possibly invalidating our stylistic analysis.
# 1. There's hundreds of stop words, so the regular expression would be very long.
# Ideally, our code should be compact and easy to read.
# A super-long regular expression is the opposite of that, and it's no fun to type either.
# And of course, the longer a regular expression, the higher the chance that you make a typo (which takes us back to point 1).
# 1. While regular expressions are fast, they are not as fast as most of the operations Python can perform on lists and counters.
# If there is an easy alternative to a regular expression, that alternative is worth exploring.
#
# Alright, so if regexes aren't the best solution, what's the alternative?
# Why, it's simple: 0.
# + [markdown] id="RWu5BuVkzSU7"
# ## Changing counts
#
# The values in a Python counter can be changed very easily.
# + colab={"base_uri": "https://localhost:8080/"} id="A2-NLvq6zSU7" outputId="10fef7d1-a53c-4d17-926c-8953b34652bc"
from collections import Counter
from pprint import pprint
# define a test counter and show its values
test = Counter(["John", "said", "that", "Mary", "said", "that", "Bill", "stinks"])
pprint(test)
# 'that' is a stop word; set its count to 0
test["that"] = 0
pprint(test)
# + [markdown] id="l8i-Tkv6zSU7"
# The code above uses the new notation `test['that']`.
#
# Counters are a subclass of dictionaries, so `test["that"]` points to the value for `"that"` in the counter `test`.
# We also say that `"that"` is a **key** that points to a specific **value**.
# The line
#
# ```python
# test["that"] = 0
# ```
#
# intstructs Python to set the value for the key `"that"` to `0`.
# + [markdown] id="Twy4-pFBzSU7"
# **Exercise.**
# Look at the code cell below.
# For each line, add a comment that briefly describes what it does (for instance, *set value of 'that' to 0*).
# If the line causes an error, fix the error and add two commments:
#
# 1. What caused the error?
# 1. What does the corrected line do?
#
# You might want to use `pprint` to look at how the counter changes after each line.
# + colab={"base_uri": "https://localhost:8080/", "height": 232} id="mmNFM7AQzSU7" outputId="71a31750-2d56-4251-8ef3-e90c318efa91"
from collections import Counter
# define a test counter and show its values
test = Counter(["John", "said", "that", "Mary", "said", "that", "Bill", "stinks"])
test["that"] = 0 # set value of 'that' to 0
test["Mary"] = test["that"]
test[John] = 10
test["said"] = test["John' - 'said"]
test["really"] = 0
# + [markdown] id="85JFA-LszSU7"
# Since we can change the values of keys in counters, stop words become very easy to deal with.
# Recall that the problem with stop words is not so much that they occur in the counter, but that they make up the large majority of high frequency words.
# Our intended fix was to delete them from the counter.
# But instead, we can just set the count of each stop word to 0.
# Then every stop word is still technically contained by the counter, but since its frequency is 0 it will no longer show up among the most common words, which is what we really care about.
#
# Alright, let's do that.
# + [markdown] id="D7didwYYzSU7"
# **Exercise.**
# Together with this notebook you found a figure which shows you the most common stop words of English (except for *whale*, you can ignore that one).
# Extend the code below so that the count for each one of the stop words listed in the figure is set to 0.
# Compare the output before and after stop word removal and ask yourself whether there has been significant progress.
# + id="lesEIVMjzSU7"
from collections import Counter
# construct the counters
counts_hamlet = Counter(hamlet)
# output with stop words
print("Most common Hamlet words before clean-up:\n", Counter.most_common(counts_hamlet, 25))
# set stop word counts to 0
# put your code here
# output without stop words
print("Most common Hamlet words after clean-up:\n", Counter.most_common(counts_hamlet, 25))
# + [markdown] id="W5bijSDYzSU7"
# Okay, this is an improvement, but it's really tedious.
# You have to write the same code over and over again, changing only the key.
# And you aren't even done yet, there's still many more stop words to be removed.
# But don't despair, you don't have to add another 100 lines of code.
# No, repetitive tasks like that are exactly why programming languages have **`for` loops**.
# + [markdown] id="8c5nXqM3zSU8"
# With a `for`-loop, setting the counts of stop words to 0 becomes a matter of just a few lines.
# + id="kl4B-VTQzSU8"
from collections import Counter
# construct the counters
counts_hamlet = Counter(hamlet)
counts_faustus = Counter(faustus)
counts_mars = Counter(mars)
stopwords = ["the", "of", "and", "a", "to", "in",
"that", "his", "it", "he", "but", "as",
"is", "with", "was", "for", "all", "this",
"at", "while", "by", "not", "from", "him",
"so", "be", "one", "you", "there", "now",
"had", "have", "or", "were", "they", "which",
"like"]
for word in stopwords:
counts_hamlet[word] = 0
counts_faustus[word] = 0
counts_mars[word] = 0
# + [markdown] id="y2B4nDw-zSU8"
# Okay, now we can finally compare the three texts based on their unigram counts.
# You can use the `Counter.most_common` function to see which words are most common in each text.
# We can also compare the overall frequency distribution.
# The code below will plot the counters, giving you a graphical representation of the frequency distribution, similar to the Zipf figures above.
#
# (Don't worry about what any of the code below does.
# Just run the cell and look at the pretty output.)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="Njq1zo6DzSU8" outputId="ddd33f87-ec7e-40e4-94e9-9311a3bf9ebf"
# %matplotlib inline
# import relevant matplotlib code
import matplotlib.pyplot as plt
# figsize(20, 10)
plt.figure(figsize=(20,10))
# the lines above are needed for Jupyter to display the plots in your browser
# do not remove them
# a little bit of preprocessing so that the data is ordered by frequency
def plot_preprocess(the_counter, n):
"""format data for plotting n most common items"""
sorted_list = sorted(the_counter.items(), key=lambda x: x[1], reverse=True)[:n]
words, counts = zip(*sorted_list)
return words, counts
for text in [counts_hamlet, counts_faustus, counts_mars]:
# you can change the max words value to look at more or fewer words in one plot
max_words = 10
words = plot_preprocess(text, max_words)[0]
counts = plot_preprocess(text, max_words)[1]
plt.bar(range(len(counts)), counts, align="center")
plt.xticks(range(len(words)), words)
plt.show()
# + [markdown] id="xYmdGy8tzSU8"
# So there you have it.
# Your first, fairly simple quantitative analysis of writing style.
# You can compare the three texts among several dimensions:
#
# 1. What are the most common words in each text?
# 1. Are the distributions very different?
# Perhaps one of them keeps repeating the same words over and over, whereas another author varies their vocabulary more and thus has a smoother curve that's not as much tilted towards the left?
#
# + id="sYaXPVIw-Hx4"
| 02_Ngrams/.ipynb_checkpoints/00_TextAnalysis_basic-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
from climpy.utils.refractive_index_utils import get_dust_ri
import climpy.utils.mie_utils as mie
from climpy.utils.aerosol_utils import get_Kok_dust_emitted_size_distribution
from climpy.utils.wrf_chem_made_utils import derive_m3s_from_mass_concentrations, get_wrf_sd_params
from climpy.utils.netcdf_utils import convert_time_data_impl
import climpy.utils.aeronet_utils as aeronet
import climpy.utils.file_path_utils as fpu
import numpy as np
import xarray as xr
import os
import netCDF4
import matplotlib.pyplot as plt
from climpy.utils.plotting_utils import save_figure, screen_width_inches, MY_DPI, save_figure_bundle
import scipy as sp
from scipy import special
# + [markdown] pycharm={"name": "#%% md\n"}
# # Verify that m3 and m0 sum up from individual components
# + pycharm={"name": "#%%\n"}
# Prepare smaller dataset for faster reading, for example, like this:
# cdo -P 2 seltimestep,24/24 -sellevidx,1/2 wrfout_d01_2017-06-15_00:00:00 debug/wrfout_d01_t24_l3
wrf_fp = '/work/mm0062/b302074/Data/AirQuality/AQABA/chem_100_v7/output/debug/wrfout_d01_t24_l3'
nc = netCDF4.Dataset(wrf_fp)
# nc = xr.open_dataset(wrf_fp)
# + pycharm={"name": "#%% derive m3 from individual aerosols concentrations\n"}
chem_opt = 100 # from namelist.input
m3_pp = derive_m3s_from_mass_concentrations(nc, chem_opt, wet=False)
m3_pp = m3_pp[np.newaxis, :] # add singleton time dimension to be consistent with m3
# + pycharm={"name": "#%% read m3 from output\n"}
sgs, dgs, m0s, m3s = get_wrf_sd_params(nc)
# individual aerosols masses should sum up to m3
# NOTE: m3i & m3j include water (h2oai, h2oaj)
m3 = np.stack(m3s).sum(axis=0) # d**3
# add inverse density for comparison
alt = nc.variables['ALT'][:]
#TODO: check
# m3 *= alt
# + pycharm={"name": "#%%\n"}
# Uncomment below, if you want to subtract the same aerosol type from both m3 and m3_pp
# In my case it is dust, which dominates
#print('{} will be subtracted from M3 and M3_pp'.format(aerosols_keys[37:41]))
#m3 -= aerosols_volumes_by_type[37:41]*6/np.pi
#m3_pp -= aerosols_volumes_by_type[37:41]*6/np.pi
# + pycharm={"name": "#%% Compute comparison diags\n"}
diff = m3 - m3_pp # this should be exactly zero
ratio = m3 / m3_pp
rel_diff = (m3 - m3_pp)/m3
#np.median(diff)
#np.median(ratio)
print('Median m3/m3_pp ratio is {}'.format(np.median(ratio)))
# + [markdown] pycharm={"name": "#%% md\n"}
# ### You may see differences in the regions with low m3
# ### This can be due to different np.pi (3.14) for example
# ### If you see considerable differences in the regions with high loading, then smth is wrong
# + pycharm={"name": "#%% plot diags\n"}
# dims: time, level, lat, lon
fig, axes = plt.subplots(constrained_layout=True, figsize=(9,12),
nrows=3, ncols=2)
plt.sca(axes[0,0])
plt.contourf(np.log10(m3[-1, 0]), levels=20)
plt.colorbar()
plt.title('log10(M3)')
plt.sca(axes[0,1])
plt.contourf(np.log10(m3_pp[-1, 0]), levels=20)
plt.colorbar()
plt.title('log10(M3_pp)')
plt.sca(axes[1,0])
plt.contourf(ratio[-1, 0], levels=20)
plt.colorbar()
plt.title('M3/M3_pp')
plt.sca(axes[1,1])
plt.contourf(diff[-1, 0], levels=20)
plt.colorbar()
plt.title('M3 - M3_pp')
plt.sca(axes[2,0])
plt.contourf(rel_diff[-1, 0], levels=20)
plt.colorbar()
plt.title('(M3-M3_pp)/M3')
plt.sca(axes[2,1])
plt.contourf(alt[-1, 0], levels=20)
plt.colorbar()
plt.title('ALT')
# + pycharm={"name": "#%% Scatter plot\n"}
plt.figure()
plt.scatter(m3[-1, 0].flatten(), m3_pp[-1, 0].flatten(), marker='.')
plt.xscale('log')
plt.yscale('log')
ax = plt.gca()
lims = [ np.min([ax.get_xlim(), ax.get_ylim()]), np.max([ax.get_xlim(), ax.get_ylim()])]
ax.plot(lims, lims, 'k-', alpha=0.75, zorder=0)
# plt.plot([0,1],[0,1], 'k', transform=ax.transAxes)
ax.set_aspect('equal')
# + pycharm={"name": "#%%\n"}
# yextra_add(1,6) = cblk(1,vnu3 )*(pirs/6.0)*1.0e9/blkdens(1) ! "n-m3-AP"/kg-air
# d**3 * pi/6 *10**9 * alt
# convfac2 = 1./alt(i,k,j)
# blkdens(blksize) = convfac2
| examples/wrf/wrf_made_aerosol_moment_diags.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# importing libraries
import pandas as pd
import numpy as np
import datetime
from collections import Iterable, defaultdict
# loading dataset
infect_afr = pd.read_excel("africa-covid19.xlsx", sheet_name = "Infected_per_day")
infect_afr.head(10)
# shape of dataframe
infect_afr.shape
# datatypes of columns
infect_afr.dtypes
# look for missing values
infect_afr.isna().any()
# replace missing values with 30222 in the entire dataframe
infect_afr.fillna(30222, inplace = True)
infect_afr.isna().any()
# create list of dataframe column names
legends = list(infect_afr.columns)
legends = legends[3:]
print(legends)
# convert datetime datatype to string datatype
legin = []
for lg in legends:
nouv = str(lg.strftime("%Y-%m-%d"))
legin.append(nouv)
print(legin)
# make country into a list
pays = list(infect_afr["COUNTRY_NAME"])
print(pays)
# identify length of lists pays and legin
print("The length of legin list: {}".format(str(len(legin))))
print("The length of pays list: {}".format(str(len(pays))))
# multiply values in legin list by 54
leginfivefour = []
n = 0
while n < 54:
for lg in legin:
leginfivefour.append(lg)
n += 1
print(len(leginfivefour))
# Function that multiplies values in list by 598
def multiplier598(listing):
column598 = []
for py in listing:
nchi = [py] * 598
for nc in nchi:
column598.append(nc)
return column598
# multiply values in pays by 598
pays598 = multiplier598(pays)
print(len(pays598))
# columns ISO and AFRICAN_REGION into list
ISO = list(infect_afr["ISO"])
AFR_REGION = list(infect_afr["AFRICAN_REGION"])
# multiply values in ISO and AFR_REGION list
ISO598 = multiplier598(ISO)
AFR_REGION598 = multiplier598(AFR_REGION)
print("Len ISO598: {}".format(str(len(ISO598))))
print("Len AFR_REGION598 {}".format(str(len(AFR_REGION598))))
# create a dictionary for the created lists and put them into a dataframe
hyb = {"ISO": ISO598,
"COUNTRY_NAME": pays598,
"AFRICAN_REGION": AFR_REGION598,
"DATE": leginfivefour}
hybrid = pd.DataFrame(hyb)
hybrid.head(10)
infect_afr.drop(['ISO', "COUNTRY_NAME", "AFRICAN_REGION"], axis = 1, inplace = True)
# values to be placed in nested list
values = infect_afr.values.tolist()
print(values)
# function to convert nested list to one-dimenisonal list
def flatten(lis):
for item in lis:
if isinstance (item, Iterable) and not isinstance (item, str):
for x in flatten(item):
yield x
else:
yield item
# apply function.
value598 = list(flatten(values))
print(len(value598))
# place value598 in hybrid dataframe
hybrid["infected"] = value598
hybrid.tail(10)
infect_afr.tail(10)
# RECOVERED_PER_DAY
# UPLOADING DATASET
recover_afr = pd.read_excel("africa-covid19.xlsx", sheet_name = "Recovered_per_day")
recover_afr.tail(10)
# shape of dataframe
recover_afr.shape
# number of countries and dates in dataset
nchi = list(recover_afr["COUNTRY_NAME"])
print("Number of countries are {}".format(str(len(nchi))))
reco = list(recover_afr.columns)
recover = reco[3:]
print("Number of days are {}".format(str(len(recover))))
# data pipeline
def transformer(dfking):
dfking.fillna(30222, inplace = True)
dfking.drop(["ISO", "COUNTRY_NAME", "AFRICAN_REGION"], axis = 1, inplace = True)
valley = dfking.values.tolist()
valley598 = list(flatten(valley))
return valley598
# recover_afr into pipeline
recover = transformer(recover_afr)
hybrid["recover"] = recover
hybrid.tail(10)
# DECEASED_PER_DAY
#
# uploading dataset
deceased_afr = pd.read_excel("africa-covid19.xlsx", sheet_name = "Deceased_per_day")
deceased_afr.tail(10)
# shape of dataframe
deceased_afr.shape
# data pipeline apply on deceased_afr
decease = transformer(deceased_afr)
hybrid["deceased"] = decease
hybrid.tail(10)
# CUMULATIVE_INFECTED
# uploading dataset
cum_infected = pd.read_excel("africa-covid19.xlsx", sheet_name = "Cumulative_infected")
cum_infected.tail(10)
# shape of dataset
cum_infected.shape
cumul_infected = transformer(cum_infected)
hybrid["cum_infected"] = cumul_infected
hybrid.tail(10)
# CUMULATIVE_RECOVERED
#
# UPLOADING DATASET
cum_recover = pd.read_excel("africa-covid19.xlsx", sheet_name = "Cumulative_recovered")
cum_recover.tail(10)
# shape of dataset
cum_recover.shape
# applying pipeline
cumul_recover = transformer(cum_recover)
hybrid["cum_recover"] = cumul_recover
hybrid.tail(10)
# CUMULATIVE_DECEASED
# UPLOADING DATASETS
cum_decease = pd.read_excel("africa-covid19.xlsx", sheet_name = "Cumulative_deceased")
cum_decease.tail(10)
# shape of dataframe
cum_decease.shape
# apply pipeline
cum_deceased = transformer(cum_decease)
hybrid["cum_deceased"] = cum_deceased
hybrid.tail(10)
# ## CLEANING, PROCESSING AND ENGINEERING
# shape of hybrid dataframe
hybrid.shape
# convert value 30222 into a missing value
hybrid.replace(30222, np.NaN, inplace = True)
# count the number of missing values in each column
hybrid.isna().sum()
# replace missing values with 0
hybrid.fillna(0, inplace = True)
# convert date datatype to datetime datatype
hybrid["DATE"] = pd.to_datetime(hybrid["DATE"])
# separate DATE column to day, month and year columns
hybrid["day"] = hybrid["DATE"].dt.day
hybrid["month"] = hybrid["DATE"].dt.month
hybrid["year"] = hybrid["DATE"].dt.year
# Delete DATE column
hybrid.drop(["DATE"], axis= 1, inplace= True)
hybrid.tail(10)
# Create month dictionary and list
month_dict = {
1: "Jan",
2: "Feb",
3: "Mar",
4: "Apr",
5: "May",
6: "Jun",
7: "Jul",
8: "Aug",
9: "Sep",
10: "Oct",
11: "Nov",
12: "Dec"
}
month_list = [x for x in range(1, 13)]
print(month_dict)
print(month_list)
# create new column with str month
hybrid["mois"] = "mapenzi"
# Replace int month with str month
for ml in month_list:
hybrid['mois'].loc[hybrid["month"] == ml] = month_dict[ml]
hybrid.tail(10)
population = {
"Nigeria": 206139589,
"Ethiopia": 114963588,
"Egypt": 102334404,
"Democratic Republic of the Congo": 89561403,
"South Africa": 59308690,
"Tanzania": 59734218,
"Kenya": 53771296,
"Uganda": 45741007,
"Algeria": 43851044,
"Sudan": 43849260,
"Morocco": 36910560,
"Angola": 32866272,
"Ghana": 31072940,
"Mozambique": 31255435,
"Madagascar": 27691018,
"Cameroon": 26545863,
"Cote d'Ivoire": 26378274,
"Niger": 24206644,
"Burkina Faso": 20903273,
"Mali": 20250833,
"Malawi": 19129952,
"Zambia": 18383955,
"Senegal": 16743927,
"Chad": 16425864,
"Somalia": 15893222,
"Zimbabwe": 14862924,
"Guinea": 13132795,
"Rwanda": 12952218,
"Benin": 12123200,
"Tunisia": 11818619,
"Burundi": 11890784,
"South Sudan": 11193725,
"Togo": 8278724,
"Sierra Leone": 7976983,
"Libya": 6871292,
"Congo": 5518087,
"Liberia": 5057681,
"Central African Republic": 4829767,
"Mauritania": 4649658,
"Eritrea": 3546421,
"Namibia": 2540905,
"Gambia": 2416668,
"Botswana": 2351627,
"Gabon": 2225734,
"Lesotho": 2142249,
"Guinea-Bissau": 1968001,
"Equatorial Guinea": 1402985,
"Mauritius": 1271768,
"Eswatini": 1160164,
"Djibouti": 988000,
"Comoros": 869601,
"Mayotte": 272815,
"Sao Tome and Principe": 219159,
"Seychelles": 98347
}
# list for keys
key_population = list(population.keys())
hybrid["population"] = 420
# new column population
for kp in key_population:
hybrid['population'].loc[hybrid["COUNTRY_NAME"] == kp] = population[kp]
hybrid.tail(10)
# SAVE TO CSV
hybrid.to_csv("hybrid.csv", index = False)
bygyttdtgyuhubgv
# MAKING OF CUMULATIVE.CSV
# upload the hybrid.csv
hype = pd.read_csv("hybrid.csv")
hype.tail(10)
# create dataframe containing rows with South Africa as its country
SA = pd.DataFrame(hype.loc[hype["COUNTRY_NAME"] == "South Africa"])
SA.head(10)
# drop unncessary columns
SA.drop(["ISO", "AFRICAN_REGION", "infected", "recover", "deceased"], axis = 1, inplace = True)
SA.tail(10)
# groupby ([month, year]) and place in dictionary month, year: cum_infected
infect_dict = SA.groupby(["month", "year"])["cum_infected"].apply(list).to_dict()
print(infect_dict)
# dictionary cum_recovered
recover_dict = SA.groupby(["month", "year"])["cum_recover"].apply(list).to_dict()
print(recover_dict)
# dictionary cumulative deceased
decease_dict = SA.groupby(["month", "year"])["cum_deceased"].apply(list).to_dict()
print(decease_dict)
# place keys of a dictionary into a list
key_in = list(infect_dict.keys())
print(key_in)
# function to replace values in dictionary
def racking(dct2, lst2):
for l in lst2:
derr = dct2[l]
last = derr[-1]
dct2[l] = last
return dct2
# implement function racking
infect_dict = racking(infect_dict, key_in)
recover_dict = racking(recover_dict, key_in)
decease_dict = racking(decease_dict, key_in)
print(decease_dict)
# function
def add_on(lst5, lst6, dct5):
val = []
for s in range(len(lst5)):
value = dct5[(lst5[s], lst6[s])]
val.append(value)
return val
# implement function
cum_infected = add_on(mois, annee, infect_dict)
cum_recovered = add_on(mois, annee, recover_dict)
cum_deceased = add_on(mois, annee, decease_dict)
# place into dataframe
cumulative = pd.DataFrame()
cumulative["months"] = mois
cumulative["year"] = annee
cumulative["infected"] = cum_infected
cumulative["recovered"] = cum_recovered
cumulative["deceased"] = cum_deceased
cumulative.tail(10)
# MAKE CUMULATIVE.CSV WITH ALL COUNTRIES
# place COUNTRY_NAME column into list
pays = list(hype["COUNTRY_NAME"])
print(len(pays))
# find unique values in list
res = defaultdict(list)
for ele in pays:
res[ele].append(ele)
uniquepays = list(res.keys())
print(uniquepays)
# multiply individual values by 2020
finalpays = []
for py in uniquepays:
nchi = [py] * 20
for nc in nchi:
finalpays.append(nc)
print(len(finalpays))
# place lists necessary
mois = ["Jan", "Feb", "Mar", "Apr", "May", "Jun", "Jul", "Aug", "Sep", "Oct", "Nov", "Dec",
"Jan", "Feb", "Mar", "Apr", "May", "Jun", "Jul", "Aug"]
annee = [2020, 2020, 2020, 2020, 2020, 2020, 2020, 2020, 2020, 2020, 2020, 2020,
2021, 2021, 2021, 2021, 2021, 2021, 2021, 2021]
# multiply lists mois and annee by 54 to fit length of uniquepays
def leginfivefour(lst8):
legin54 = []
n = 0
while n < 54:
for lg in lst8:
legin54.append(lg)
n += 1
return legin54
# implement function
mois54 = leginfivefour(mois)
annee54 = leginfivefour(annee)
print(len(mois54))
print(len(annee54))
# groupby ([COUNTRY_NAME, month, year]) [cum_infected] then place in dictionary
infect_dict = hype.groupby(["COUNTRY_NAME", "month", "year"])["cum_infected"].apply(list).to_dict()
print(infect_dict)
# groupby by cum_recovered
recover_dict = hype.groupby(["COUNTRY_NAME", "month", "year"])["cum_recover"].apply(list).to_dict()
print(recover_dict)
# groupby cum_deceased
decease_dict = hype.groupby(["COUNTRY_NAME", "month", "year"])["cum_deceased"].apply(list).to_dict()
print(decease_dict)
# keys to be put in a list
key_in = list(infect_dict.keys())
print(key_in)
# function that removes list value in dictionary and replaces with last value on list
def racking(dct2, lst2):
for l in lst2:
derr = dct2[l]
last = derr[-1]
dct2[l] = last
return dct2
# implement racking function
infect_dict = racking(infect_dict, key_in)
recover_dict = racking(recover_dict, key_in)
decease_dict = racking(decease_dict, key_in)
print(len(infect_dict))
# function that puts values of cum_infected, cum_recovered and cum_deceased in relation to COUNTRY_NAME, YEAR, MONTH
def add_on(lst5, lst6, lst7, dct5):
val = []
for s in range(1080):
value = dct5[(lst5[s], lst6[s], lst7[s])]
val.append(value)
return val
# implement function
cum_infected = add_on(finalpays, mois54, annee54, infect_dict)
cum_recovered = add_on(finalpays, mois54, annee54, recover_dict)
cum_deceased = add_on(finalpays, mois54, annee54, decease_dict)
print(cum_deceased)
# create dataframe
# place into dataframe
cumulative = pd.DataFrame()
cumulative["COUNTRY_NAME"] = finalpays
cumulative["months"] = mois54
cumulative["year"] = annee54
cumulative["infected"] = cum_infected
cumulative["recovered"] = cum_recovered
cumulative["deceased"] = cum_deceased
cumulative.tail(10)
cumulative["population"] = 420
# new column population
for kp in key_population:
cumulative['population'].loc[cumulative["COUNTRY_NAME"] == kp] = population[kp]
cumulative.tail(10)
#save to csv
cumulative.to_csv("cumulative.csv", index = False)
| PROJET TROIS AFRICA COVID-19.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
## Monday class: Markov chain
import random
random.choices(['hot','cold'],[0.1,0.9])
# -
random.choices(['hot','cold'],[0.1,0.9])
random.choices(['hot','cold'],[0.1,0.9])
# +
state = 'hot'
if state=='hot':
probs = [0.8,0.2]
else:
probs = [0.1,0.9]
state = random.choices(['hot','cold', probs])[0]
# -
state
| week08/week08 class exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# import the project1-prepareData notebook:
# !pip install ipynb
from ipynb.fs.full.project1_prepareData import *
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# +
#Hot encoding and Logistic regretion
from sklearn import linear_model
from sklearn.feature_extraction import DictVectorizer
def train(dataFrame, y):
# Hot Encoding
dicts = dataFrame.to_dict(orient="records")
dv = DictVectorizer(sparse=False)
X = dv.fit_transform(dicts)
# train
model = linear_model.LogisticRegression()
model.fit(X, y)
return dv, model
dv, model = train(df_train, y_train)
weights = model.coef_[0].round(3) # weights
w0 = model.intercept_ # bias, w0
weights_with_featureNames = dict(zip(dv.get_feature_names(), weights))
print("w0 =", w0)
# display(weights_with_featureNames)
display(pd.DataFrame([weights], index=["weight"], columns=dv.get_feature_names()))
# +
# Predict
def predict(dataFrame, dv, model):
dicts = dataFrame.to_dict(orient="records")
X = dv.transform(dicts)
y_pred = model.predict_proba(X)[:,1]
return y_pred
proba = predict(df_val, dv, model)
y_pred_val = proba #take column 1
low_salary_pred = (y_pred_val >= 0.5)
# customers_with_predicted_low_income = df_val[low_salary_pred]
# +
# Check accuracy
#check average accuracy on y_val
df_pred = pd.DataFrame()
df_pred["probability"] = y_pred_val
df_pred["prediction"] = low_salary_pred.astype(int)
df_pred["actual"] = y_val
df_pred["prediction_correct"] = df_pred.prediction == df_pred.actual
display(df_pred)
print("Accuracy % on y_val:",df_pred.prediction_correct.mean())
# +
# AUC and ROC curve
from sklearn import metrics
fpr, tpr, thresholds = metrics.roc_curve(y_val, y_pred_val)
auc = metrics.auc(fpr, tpr)
# print("thresholds:", thresholds)
print("AUC for LogisticRegresion:", auc)
import matplotlib.pyplot as plt
from sklearn import datasets, model_selection, svm
dicts = df_val.to_dict(orient="records")
X_val = dv.transform(dicts)
print("ROC curve:")
metrics.plot_roc_curve(model, X_val, y_val)
plt.show()
| project1/project1_logisticRegresion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Predicting Ki of Ligands to a Protein
# In this notebook, we analyze the BACE enyzme and build machine learning models for predicting the Ki of ligands to the protein. We will use the `deepchem` library to load this data into memory, split into train/test/validation folds, build and cross-validate models, and report statistics.
# %load_ext autoreload
# %autoreload 2
# %pdb off
# +
import os
import sys
import deepchem as dc
from deepchem.utils.save import load_from_disk
current_dir = os.path.dirname(os.path.realpath("__file__"))
dc.utils.download_url("https://s3-us-west-1.amazonaws.com/deepchem.io/datasets/desc_canvas_aug30.csv",
current_dir)
dataset_file = "desc_canvas_aug30.csv"
dataset = load_from_disk(dataset_file)
num_display=10
pretty_columns = (
"[" + ",".join(["'%s'" % column for column in dataset.columns.values[:num_display]])
+ ",...]")
dc.utils.download_url("https://s3-us-west-1.amazonaws.com/deepchem.io/datasets/crystal_desc_canvas_aug30.csv",
current_dir)
crystal_dataset_file = "crystal_desc_canvas_aug30.csv"
crystal_dataset = load_from_disk(crystal_dataset_file)
print("Columns of dataset: %s" % pretty_columns)
print("Number of examples in dataset: %s" % str(dataset.shape[0]))
print("Number of examples in crystal dataset: %s" % str(crystal_dataset.shape[0]))
# -
# To gain a visual understanding of compounds in our dataset, let's draw them using rdkit. We define a couple of helper functions to get started.
# +
import tempfile
from rdkit import Chem
from rdkit.Chem import Draw
from itertools import islice
from IPython.display import Image, display, HTML
def display_images(filenames):
"""Helper to pretty-print images."""
for filename in filenames:
display(Image(filename))
def mols_to_pngs(mols, basename="test"):
"""Helper to write RDKit mols to png files."""
filenames = []
for i, mol in enumerate(mols):
filename = "BACE_%s%d.png" % (basename, i)
Draw.MolToFile(mol, filename)
filenames.append(filename)
return filenames
# -
# Now, we display a compound from the dataset. Note the complex ring structures and polar structures.
num_to_display = 12
molecules = []
for _, data in islice(dataset.iterrows(), num_to_display):
molecules.append(Chem.MolFromSmiles(data["mol"]))
display_images(mols_to_pngs(molecules, basename="dataset"))
# Now let's picture the compounds in the crystal structure collection
num_to_display = 12
molecules = []
for _, data in islice(crystal_dataset.iterrows(), num_to_display):
molecules.append(Chem.MolFromSmiles(data["mol"]))
display_images(mols_to_pngs(molecules, basename="crystal_dataset"))
# Analyzing the distribution of pIC50 values in the dataset gives us a nice spread.
# +
# %matplotlib inline
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
pIC50s = np.array(dataset["pIC50"])
# Remove some dirty data from the dataset
pIC50s = [pIC50 for pIC50 in pIC50s if pIC50 != '']
n, bins, patches = plt.hist(pIC50s, 50, facecolor='green', alpha=0.75)
plt.xlabel('Measured pIC50')
plt.ylabel('Number of compounds')
plt.title(r'Histogram of pIC50 Values')
plt.grid(True)
plt.show()
# -
# We now featurize the data using the Canvas samples. To do so, we must specify the columns in the data input that correspond to the features. (Note that CanvasUID is excluded!)
user_specified_features = ['MW','AlogP','HBA','HBD','RB','HeavyAtomCount','ChiralCenterCount','ChiralCenterCountAllPossible','RingCount','PSA','Estate','MR','Polar','sLi_Key','ssBe_Key','ssssBem_Key','sBH2_Key','ssBH_Key','sssB_Key','ssssBm_Key','sCH3_Key','dCH2_Key','ssCH2_Key','tCH_Key','dsCH_Key','aaCH_Key','sssCH_Key','ddC_Key','tsC_Key','dssC_Key','aasC_Key','aaaC_Key','ssssC_Key','sNH3_Key','sNH2_Key','ssNH2_Key','dNH_Key','ssNH_Key','aaNH_Key','tN_Key','sssNH_Key','dsN_Key','aaN_Key','sssN_Key','ddsN_Key','aasN_Key','ssssN_Key','daaN_Key','sOH_Key','dO_Key','ssO_Key','aaO_Key','aOm_Key','sOm_Key','sF_Key','sSiH3_Key','ssSiH2_Key','sssSiH_Key','ssssSi_Key','sPH2_Key','ssPH_Key','sssP_Key','dsssP_Key','ddsP_Key','sssssP_Key','sSH_Key','dS_Key','ssS_Key','aaS_Key','dssS_Key','ddssS_Key','ssssssS_Key','Sm_Key','sCl_Key','sGeH3_Key','ssGeH2_Key','sssGeH_Key','ssssGe_Key','sAsH2_Key','ssAsH_Key','sssAs_Key','dsssAs_Key','ddsAs_Key','sssssAs_Key','sSeH_Key','dSe_Key','ssSe_Key','aaSe_Key','dssSe_Key','ssssssSe_Key','ddssSe_Key','sBr_Key','sSnH3_Key','ssSnH2_Key','sssSnH_Key','ssssSn_Key','sI_Key','sPbH3_Key','ssPbH2_Key','sssPbH_Key','ssssPb_Key','sLi_Cnt','ssBe_Cnt','ssssBem_Cnt','sBH2_Cnt','ssBH_Cnt','sssB_Cnt','ssssBm_Cnt','sCH3_Cnt','dCH2_Cnt','ssCH2_Cnt','tCH_Cnt','dsCH_Cnt','aaCH_Cnt','sssCH_Cnt','ddC_Cnt','tsC_Cnt','dssC_Cnt','aasC_Cnt','aaaC_Cnt','ssssC_Cnt','sNH3_Cnt','sNH2_Cnt','ssNH2_Cnt','dNH_Cnt','ssNH_Cnt','aaNH_Cnt','tN_Cnt','sssNH_Cnt','dsN_Cnt','aaN_Cnt','sssN_Cnt','ddsN_Cnt','aasN_Cnt','ssssN_Cnt','daaN_Cnt','sOH_Cnt','dO_Cnt','ssO_Cnt','aaO_Cnt','aOm_Cnt','sOm_Cnt','sF_Cnt','sSiH3_Cnt','ssSiH2_Cnt','sssSiH_Cnt','ssssSi_Cnt','sPH2_Cnt','ssPH_Cnt','sssP_Cnt','dsssP_Cnt','ddsP_Cnt','sssssP_Cnt','sSH_Cnt','dS_Cnt','ssS_Cnt','aaS_Cnt','dssS_Cnt','ddssS_Cnt','ssssssS_Cnt','Sm_Cnt','sCl_Cnt','sGeH3_Cnt','ssGeH2_Cnt','sssGeH_Cnt','ssssGe_Cnt','sAsH2_Cnt','ssAsH_Cnt','sssAs_Cnt','dsssAs_Cnt','ddsAs_Cnt','sssssAs_Cnt','sSeH_Cnt','dSe_Cnt','ssSe_Cnt','aaSe_Cnt','dssSe_Cnt','ssssssSe_Cnt','ddssSe_Cnt','sBr_Cnt','sSnH3_Cnt','ssSnH2_Cnt','sssSnH_Cnt','ssssSn_Cnt','sI_Cnt','sPbH3_Cnt','ssPbH2_Cnt','sssPbH_Cnt','ssssPb_Cnt','sLi_Sum','ssBe_Sum','ssssBem_Sum','sBH2_Sum','ssBH_Sum','sssB_Sum','ssssBm_Sum','sCH3_Sum','dCH2_Sum','ssCH2_Sum','tCH_Sum','dsCH_Sum','aaCH_Sum','sssCH_Sum','ddC_Sum','tsC_Sum','dssC_Sum','aasC_Sum','aaaC_Sum','ssssC_Sum','sNH3_Sum','sNH2_Sum','ssNH2_Sum','dNH_Sum','ssNH_Sum','aaNH_Sum','tN_Sum','sssNH_Sum','dsN_Sum','aaN_Sum','sssN_Sum','ddsN_Sum','aasN_Sum','ssssN_Sum','daaN_Sum','sOH_Sum','dO_Sum','ssO_Sum','aaO_Sum','aOm_Sum','sOm_Sum','sF_Sum','sSiH3_Sum','ssSiH2_Sum','sssSiH_Sum','ssssSi_Sum','sPH2_Sum','ssPH_Sum','sssP_Sum','dsssP_Sum','ddsP_Sum','sssssP_Sum','sSH_Sum','dS_Sum','ssS_Sum','aaS_Sum','dssS_Sum','ddssS_Sum','ssssssS_Sum','Sm_Sum','sCl_Sum','sGeH3_Sum','ssGeH2_Sum','sssGeH_Sum','ssssGe_Sum','sAsH2_Sum','ssAsH_Sum','sssAs_Sum','dsssAs_Sum','ddsAs_Sum','sssssAs_Sum','sSeH_Sum','dSe_Sum','ssSe_Sum','aaSe_Sum','dssSe_Sum','ssssssSe_Sum','ddssSe_Sum','sBr_Sum','sSnH3_Sum','ssSnH2_Sum','sssSnH_Sum','ssssSn_Sum','sI_Sum','sPbH3_Sum','ssPbH2_Sum','sssPbH_Sum','ssssPb_Sum','sLi_Avg','ssBe_Avg','ssssBem_Avg','sBH2_Avg','ssBH_Avg','sssB_Avg','ssssBm_Avg','sCH3_Avg','dCH2_Avg','ssCH2_Avg','tCH_Avg','dsCH_Avg','aaCH_Avg','sssCH_Avg','ddC_Avg','tsC_Avg','dssC_Avg','aasC_Avg','aaaC_Avg','ssssC_Avg','sNH3_Avg','sNH2_Avg','ssNH2_Avg','dNH_Avg','ssNH_Avg','aaNH_Avg','tN_Avg','sssNH_Avg','dsN_Avg','aaN_Avg','sssN_Avg','ddsN_Avg','aasN_Avg','ssssN_Avg','daaN_Avg','sOH_Avg','dO_Avg','ssO_Avg','aaO_Avg','aOm_Avg','sOm_Avg','sF_Avg','sSiH3_Avg','ssSiH2_Avg','sssSiH_Avg','ssssSi_Avg','sPH2_Avg','ssPH_Avg','sssP_Avg','dsssP_Avg','ddsP_Avg','sssssP_Avg','sSH_Avg','dS_Avg','ssS_Avg','aaS_Avg','dssS_Avg','ddssS_Avg','ssssssS_Avg','Sm_Avg','sCl_Avg','sGeH3_Avg','ssGeH2_Avg','sssGeH_Avg','ssssGe_Avg','sAsH2_Avg','ssAsH_Avg','sssAs_Avg','dsssAs_Avg','ddsAs_Avg','sssssAs_Avg','sSeH_Avg','dSe_Avg','ssSe_Avg','aaSe_Avg','dssSe_Avg','ssssssSe_Avg','ddssSe_Avg','sBr_Avg','sSnH3_Avg','ssSnH2_Avg','sssSnH_Avg','ssssSn_Avg','sI_Avg','sPbH3_Avg','ssPbH2_Avg','sssPbH_Avg','ssssPb_Avg','First Zagreb (ZM1)','First Zagreb index by valence vertex degrees (ZM1V)','Second Zagreb (ZM2)','Second Zagreb index by valence vertex degrees (ZM2V)','Polarity (Pol)','Narumi Simple Topological (NST)','Narumi Harmonic Topological (NHT)','Narumi Geometric Topological (NGT)','Total structure connectivity (TSC)','Wiener (W)','Mean Wiener (MW)','Xu (Xu)','Quadratic (QIndex)','Radial centric (RC)','Mean Square Distance Balaban (MSDB)','Superpendentic (SP)','Harary (Har)','Log of product of row sums (LPRS)','Pogliani (Pog)','Schultz Molecular Topological (SMT)','Schultz Molecular Topological by valence vertex degrees (SMTV)','Mean Distance Degree Deviation (MDDD)','Ramification (Ram)','Gutman Molecular Topological (GMT)','Gutman MTI by valence vertex degrees (GMTV)','Average vertex distance degree (AVDD)','Unipolarity (UP)','Centralization (CENT)','Variation (VAR)','Molecular electrotopological variation (MEV)','Maximal electrotopological positive variation (MEPV)','Maximal electrotopological negative variation (MENV)','Eccentric connectivity (ECCc)','Eccentricity (ECC)','Average eccentricity (AECC)','Eccentric (DECC)','Valence connectivity index chi-0 (vX0)','Valence connectivity index chi-1 (vX1)','Valence connectivity index chi-2 (vX2)','Valence connectivity index chi-3 (vX3)','Valence connectivity index chi-4 (vX4)','Valence connectivity index chi-5 (vX5)','Average valence connectivity index chi-0 (AvX0)','Average valence connectivity index chi-1 (AvX1)','Average valence connectivity index chi-2 (AvX2)','Average valence connectivity index chi-3 (AvX3)','Average valence connectivity index chi-4 (AvX4)','Average valence connectivity index chi-5 (AvX5)','Quasi Wiener (QW)','First Mohar (FM)','Second Mohar (SM)','Spanning tree number (STN)','Kier benzene-likeliness index (KBLI)','Topological charge index of order 1 (TCI1)','Topological charge index of order 2 (TCI2)','Topological charge index of order 3 (TCI3)','Topological charge index of order 4 (TCI4)','Topological charge index of order 5 (TCI5)','Topological charge index of order 6 (TCI6)','Topological charge index of order 7 (TCI7)','Topological charge index of order 8 (TCI8)','Topological charge index of order 9 (TCI9)','Topological charge index of order 10 (TCI10)','Mean topological charge index of order 1 (MTCI1)','Mean topological charge index of order 2 (MTCI2)','Mean topological charge index of order 3 (MTCI3)','Mean topological charge index of order 4 (MTCI4)','Mean topological charge index of order 5 (MTCI5)','Mean topological charge index of order 6 (MTCI6)','Mean topological charge index of order 7 (MTCI7)','Mean topological charge index of order 8 (MTCI8)','Mean topological charge index of order 9 (MTCI9)','Mean topological charge index of order 10 (MTCI10)','Global topological charge (GTC)','Hyper-distance-path index (HDPI)','Reciprocal hyper-distance-path index (RHDPI)','Square reciprocal distance sum (SRDS)','Modified Randic connectivity (MRC)','Balaban centric (BC)','Lopping centric (LC)','Kier Hall electronegativity (KHE)','Sum of topological distances between N..N (STD(N N))','Sum of topological distances between N..O (STD(N O))','Sum of topological distances between N..S (STD(N S))','Sum of topological distances between N..P (STD(N P))','Sum of topological distances between N..F (STD(N F))','Sum of topological distances between N..Cl (STD(N Cl))','Sum of topological distances between N..Br (STD(N Br))','Sum of topological distances between N..I (STD(N I))','Sum of topological distances between O..O (STD(O O))','Sum of topological distances between O..S (STD(O S))','Sum of topological distances between O..P (STD(O P))','Sum of topological distances between O..F (STD(O F))','Sum of topological distances between O..Cl (STD(O Cl))','Sum of topological distances between O..Br (STD(O Br))','Sum of topological distances between O..I (STD(O I))','Sum of topological distances between S..S (STD(S S))','Sum of topological distances between S..P (STD(S P))','Sum of topological distances between S..F (STD(S F))','Sum of topological distances between S..Cl (STD(S Cl))','Sum of topological distances between S..Br (STD(S Br))','Sum of topological distances between S..I (STD(S I))','Sum of topological distances between P..P (STD(P P))','Sum of topological distances between P..F (STD(P F))','Sum of topological distances between P..Cl (STD(P Cl))','Sum of topological distances between P..Br (STD(P Br))','Sum of topological distances between P..I (STD(P I))','Sum of topological distances between F..F (STD(F F))','Sum of topological distances between F..Cl (STD(F Cl))','Sum of topological distances between F..Br (STD(F Br))','Sum of topological distances between F..I (STD(F I))','Sum of topological distances between Cl..Cl (STD(Cl Cl))','Sum of topological distances between Cl..Br (STD(Cl Br))','Sum of topological distances between Cl..I (STD(Cl I))','Sum of topological distances between Br..Br (STD(Br Br))','Sum of topological distances between Br..I (STD(Br I))','Sum of topological distances between I..I (STD(I I))','Wiener-type index from Z weighted distance matrix - Barysz matrix (WhetZ)','Wiener-type index from electronegativity weighted distance matrix (Whete)','Wiener-type index from mass weighted distance matrix (Whetm)','Wiener-type index from van der waals weighted distance matrix (Whetv)','Wiener-type index from polarizability weighted distance matrix (Whetp)','Balaban-type index from Z weighted distance matrix - Barysz matrix (JhetZ)','Balaban-type index from electronegativity weighted distance matrix (Jhete)','Balaban-type index from mass weighted distance matrix (Jhetm)','Balaban-type index from van der waals weighted distance matrix (Jhetv)','Balaban-type index from polarizability weighted distance matrix (Jhetp)','Topological diameter (TD)','Topological radius (TR)','Petitjean 2D shape (PJ2DS)','Balaban distance connectivity index (J)','Solvation connectivity index chi-0 (SCIX0)','Solvation connectivity index chi-1 (SCIX1)','Solvation connectivity index chi-2 (SCIX2)','Solvation connectivity index chi-3 (SCIX3)','Solvation connectivity index chi-4 (SCIX4)','Solvation connectivity index chi-5 (SCIX5)','Connectivity index chi-0 (CIX0)','Connectivity chi-1 [Randic connectivity] (CIX1)','Connectivity index chi-2 (CIX2)','Connectivity index chi-3 (CIX3)','Connectivity index chi-4 (CIX4)','Connectivity index chi-5 (CIX5)','Average connectivity index chi-0 (ACIX0)','Average connectivity index chi-1 (ACIX1)','Average connectivity index chi-2 (ACIX2)','Average connectivity index chi-3 (ACIX3)','Average connectivity index chi-4 (ACIX4)','Average connectivity index chi-5 (ACIX5)','reciprocal distance Randic-type index (RDR)','reciprocal distance square Randic-type index (RDSR)','1-path Kier alpha-modified shape index (KAMS1)','2-path Kier alpha-modified shape index (KAMS2)','3-path Kier alpha-modified shape index (KAMS3)','Kier flexibility (KF)','path/walk 2 - Randic shape index (RSIpw2)','path/walk 3 - Randic shape index (RSIpw3)','path/walk 4 - Randic shape index (RSIpw4)','path/walk 5 - Randic shape index (RSIpw5)','E-state topological parameter (ETP)','Ring Count 3 (RNGCNT3)','Ring Count 4 (RNGCNT4)','Ring Count 5 (RNGCNT5)','Ring Count 6 (RNGCNT6)','Ring Count 7 (RNGCNT7)','Ring Count 8 (RNGCNT8)','Ring Count 9 (RNGCNT9)','Ring Count 10 (RNGCNT10)','Ring Count 11 (RNGCNT11)','Ring Count 12 (RNGCNT12)','Ring Count 13 (RNGCNT13)','Ring Count 14 (RNGCNT14)','Ring Count 15 (RNGCNT15)','Ring Count 16 (RNGCNT16)','Ring Count 17 (RNGCNT17)','Ring Count 18 (RNGCNT18)','Ring Count 19 (RNGCNT19)','Ring Count 20 (RNGCNT20)','Atom Count (ATMCNT)','Bond Count (BNDCNT)','Atoms in Ring System (ATMRNGCNT)','Bonds in Ring System (BNDRNGCNT)','Cyclomatic number (CYCLONUM)','Number of ring systems (NRS)','Normalized number of ring systems (NNRS)','Ring Fusion degree (RFD)','Ring perimeter (RNGPERM)','Ring bridge count (RNGBDGE)','Molecule cyclized degree (MCD)','Ring Fusion density (RFDELTA)','Ring complexity index (RCI)','Van der Waals surface area (VSA)','MR1 (MR1)','MR2 (MR2)','MR3 (MR3)','MR4 (MR4)','MR5 (MR5)','MR6 (MR6)','MR7 (MR7)','MR8 (MR8)','ALOGP1 (ALOGP1)','ALOGP2 (ALOGP2)','ALOGP3 (ALOGP3)','ALOGP4 (ALOGP4)','ALOGP5 (ALOGP5)','ALOGP6 (ALOGP6)','ALOGP7 (ALOGP7)','ALOGP8 (ALOGP8)','ALOGP9 (ALOGP9)','ALOGP10 (ALOGP10)','PEOE1 (PEOE1)','PEOE2 (PEOE2)','PEOE3 (PEOE3)','PEOE4 (PEOE4)','PEOE5 (PEOE5)','PEOE6 (PEOE6)','PEOE7 (PEOE7)','PEOE8 (PEOE8)','PEOE9 (PEOE9)','PEOE10 (PEOE10)','PEOE11 (PEOE11)','PEOE12 (PEOE12)','PEOE13 (PEOE13)','PEOE14 (PEOE14)']
# +
import deepchem as dc
import tempfile, shutil
featurizer = dc.feat.UserDefinedFeaturizer(user_specified_features)
loader = dc.data.UserCSVLoader(
tasks=["Class"], smiles_field="mol", id_field="mol",
featurizer=featurizer)
dataset = loader.featurize(dataset_file)
crystal_dataset = loader.featurize(crystal_dataset_file)
# -
# This data is already split into three subsets "Train" and "Test" with 20% and 80% respectively of the total data from the BACE enzyme. There is also a "Validation" set that contains data from a separate (but related assay). (Note that these names are really misnomers. The "Test" set would be called a validation set in standard machine-learning practice and the "Validation" set would typically be called an external test set.) Hence, we will rename the datasets after loading them.
splitter = dc.splits.SpecifiedSplitter(dataset_file, "Model")
train_dataset, valid_dataset, test_dataset = splitter.train_valid_test_split(
dataset)
#NOTE THE RENAMING:
valid_dataset, test_dataset = test_dataset, valid_dataset
# Let's quickly take a look at a compound in the validation set. (The compound displayed earlier was drawn from the train set).
print(valid_dataset.ids)
valid_mols = [Chem.MolFromSmiles(compound)
for compound in islice(valid_dataset.ids, num_to_display)]
display_images(mols_to_pngs(valid_mols, basename="valid_set"))
# Let's now write these datasets to disk
print("Number of compounds in train set")
print(len(train_dataset))
print("Number of compounds in validation set")
print(len(valid_dataset))
print("Number of compounds in test set")
print(len(test_dataset))
print("Number of compounds in crystal set")
print(len(crystal_dataset))
# The performance of common machine-learning algorithms can be very sensitive to preprocessing of the data. One common transformation applied to data is to normalize it to have zero-mean and unit-standard-deviation. We will apply this transformation to the pIC50 values (as seen above, the pIC50s range from 2 to 11).
# +
transformers = [
dc.trans.NormalizationTransformer(transform_X=True, dataset=train_dataset),
dc.trans.ClippingTransformer(transform_X=True, dataset=train_dataset)]
datasets = [train_dataset, valid_dataset, test_dataset, crystal_dataset]
for i, dataset in enumerate(datasets):
for transformer in transformers:
datasets[i] = transformer.transform(dataset)
train_dataset, valid_dataset, test_dataset, crystal_dataset = datasets
# -
# We now fit simple random forest models to our datasets.
# +
from sklearn.ensemble import RandomForestClassifier
def rf_model_builder(model_params, model_dir):
sklearn_model = RandomForestClassifier(**model_params)
return dc.models.SklearnModel(sklearn_model, model_dir)
params_dict = {
"n_estimators": [10, 100],
"max_features": ["auto", "sqrt", "log2", None],
}
metric = dc.metrics.Metric(dc.metrics.roc_auc_score)
optimizer = dc.hyper.HyperparamOpt(rf_model_builder)
best_rf, best_rf_hyperparams, all_rf_results = optimizer.hyperparam_search(
params_dict, train_dataset, valid_dataset, transformers,
metric=metric)
# +
import numpy.random
params_dict = {"learning_rate": np.power(10., np.random.uniform(-5, -3, size=1)),
"weight_decay_penalty": np.power(10, np.random.uniform(-6, -4, size=1)),
"nb_epoch": [40] }
n_features = train_dataset.get_data_shape()[0]
def model_builder(model_params, model_dir):
model = dc.models.MultiTaskClassifier(
1, n_features, layer_sizes=[1000], dropouts=.25,
batch_size=50, **model_params)
return model
optimizer = dc.hyper.HyperparamOpt(model_builder)
best_dnn, best_dnn_hyperparams, all_dnn_results = optimizer.hyperparam_search(
params_dict, train_dataset, valid_dataset, transformers,
metric=metric)
# -
# Now let's evaluate the best model on the validation and test sets and save the results to csv.
# +
from deepchem.utils.evaluate import Evaluator
rf_train_csv_out = "rf_train_regressor.csv"
rf_train_stats_out = "rf_train_stats_regressor.txt"
rf_train_evaluator = Evaluator(best_rf, train_dataset, transformers)
rf_train_score = rf_train_evaluator.compute_model_performance(
[metric], rf_train_csv_out, rf_train_stats_out)
print("RF Train set AUC %f" % (rf_train_score["roc_auc_score"]))
rf_valid_csv_out = "rf_valid_regressor.csv"
rf_valid_stats_out = "rf_valid_stats_regressor.txt"
rf_valid_evaluator = Evaluator(best_rf, valid_dataset, transformers)
rf_valid_score = rf_valid_evaluator.compute_model_performance(
[metric], rf_valid_csv_out, rf_valid_stats_out)
print("RF Valid set AUC %f" % (rf_valid_score["roc_auc_score"]))
rf_test_csv_out = "rf_test_regressor.csv"
rf_test_stats_out = "rf_test_stats_regressor.txt"
rf_test_evaluator = Evaluator(best_rf, test_dataset, transformers)
rf_test_score = rf_test_evaluator.compute_model_performance(
[metric], rf_test_csv_out, rf_test_stats_out)
print("RF Test set AUC %f" % (rf_test_score["roc_auc_score"]))
rf_crystal_csv_out = "rf_crystal_regressor.csv"
rf_crystal_stats_out = "rf_crystal_stats_regressor.txt"
rf_crystal_evaluator = Evaluator(best_rf, crystal_dataset, transformers)
rf_crystal_score = rf_crystal_evaluator.compute_model_performance(
[metric], rf_crystal_csv_out, rf_crystal_stats_out)
print("RF Crystal set R^2 %f" % (rf_crystal_score["roc_auc_score"]))
# +
dnn_train_csv_out = "dnn_train_classifier.csv"
dnn_train_stats_out = "dnn_train_classifier_stats.txt"
dnn_train_evaluator = Evaluator(best_dnn, train_dataset, transformers)
dnn_train_score = dnn_train_evaluator.compute_model_performance(
[metric], dnn_train_csv_out, dnn_train_stats_out)
print("DNN Train set AUC %f" % (dnn_train_score["roc_auc_score"]))
dnn_valid_csv_out = "dnn_valid_classifier.csv"
dnn_valid_stats_out = "dnn_valid_classifier_stats.txt"
dnn_valid_evaluator = Evaluator(best_dnn, valid_dataset, transformers)
dnn_valid_score = dnn_valid_evaluator.compute_model_performance(
[metric], dnn_valid_csv_out, dnn_valid_stats_out)
print("DNN Valid set AUC %f" % (dnn_valid_score["roc_auc_score"]))
dnn_test_csv_out = "dnn_test_classifier.csv"
dnn_test_stats_out = "dnn_test_classifier_stats.txt"
dnn_test_evaluator = Evaluator(best_dnn, test_dataset, transformers)
dnn_test_score = dnn_test_evaluator.compute_model_performance(
[metric], dnn_test_csv_out, dnn_test_stats_out)
print("DNN Test set AUC %f" % (dnn_test_score["roc_auc_score"]))
dnn_crystal_csv_out = "dnn_crystal_classifier.csv"
dnn_crystal_stats_out = "dnn_crystal_stats_classifier.txt"
dnn_crystal_evaluator = Evaluator(best_dnn, crystal_dataset, transformers)
dnn_crystal_score = dnn_crystal_evaluator.compute_model_performance(
[metric], dnn_crystal_csv_out, dnn_crystal_stats_out)
print("DNN Crystal set AUC %f" % (dnn_crystal_score["roc_auc_score"]))
# -
# Now, we construct regression models for the data.
#Make directories to store the raw and featurized datasets.
featurizer = dc.feat.UserDefinedFeaturizer(user_specified_features)
loader = dc.data.UserCSVLoader(
tasks=["pIC50"], smiles_field="mol", id_field="CID",
featurizer=featurizer)
dataset = loader.featurize(dataset_file)
crystal_dataset = loader.featurize(crystal_dataset_file)
splitter = dc.splits.SpecifiedSplitter(dataset_file, "Model")
train_dataset, valid_dataset, test_dataset = splitter.train_valid_test_split(
dataset)
#NOTE THE RENAMING:
valid_dataset, test_dataset = test_dataset, valid_dataset
print("Number of compounds in train set")
print(len(train_dataset))
print("Number of compounds in validation set")
print(len(valid_dataset))
print("Number of compounds in test set")
print(len(test_dataset))
print("Number of compounds in crystal set")
print(len(crystal_dataset))
# +
transformers = [
dc.trans.NormalizationTransformer(transform_X=True, dataset=train_dataset),
dc.trans.ClippingTransformer(transform_X=True, dataset=train_dataset)]
datasets = [train_dataset, valid_dataset, test_dataset, crystal_dataset]
for i, dataset in enumerate(datasets):
for transformer in transformers:
datasets[i] = transformer.transform(dataset)
train_dataset, valid_dataset, test_dataset, crystal_dataset = datasets
# +
from sklearn.ensemble import RandomForestRegressor
def rf_model_builder(model_params, model_dir):
sklearn_model = RandomForestRegressor(**model_params)
return dc.models.SklearnModel(sklearn_model, model_dir)
params_dict = {
"n_estimators": [10, 100],
"max_features": ["auto", "sqrt", "log2", None],
}
metric = dc.metrics.Metric(dc.metrics.r2_score)
optimizer = dc.hyper.HyperparamOpt(rf_model_builder)
best_rf, best_rf_hyperparams, all_rf_results = optimizer.hyperparam_search(
params_dict, train_dataset, valid_dataset, transformers,
metric=metric)
# +
import numpy.random
params_dict = {"learning_rate": np.power(10., np.random.uniform(-5, -3, size=2)),
"weight_decay_penalty": np.power(10, np.random.uniform(-6, -4, size=2)),
"nb_epoch": [20] }
n_features = train_dataset.get_data_shape()[0]
def model_builder(model_params, model_dir):
model = dc.models.MultiTaskRegressor(
1, n_features, layer_sizes=[1000], dropouts=[.25],
batch_size=50, **model_params)
return model
optimizer = dc.hyper.HyperparamOpt(model_builder)
best_dnn, best_dnn_hyperparams, all_dnn_results = optimizer.hyperparam_search(
params_dict, train_dataset, valid_dataset, transformers,
metric=metric)
# +
from deepchem.utils.evaluate import Evaluator
rf_train_csv_out = "rf_train_regressor.csv"
rf_train_stats_out = "rf_train_stats_regressor.txt"
rf_train_evaluator = Evaluator(best_rf, train_dataset, transformers)
rf_train_score = rf_train_evaluator.compute_model_performance(
[metric], rf_train_csv_out, rf_train_stats_out)
print("RF Train set R^2 %f" % (rf_train_score["r2_score"]))
rf_valid_csv_out = "rf_valid_regressor.csv"
rf_valid_stats_out = "rf_valid_stats_regressor.txt"
rf_valid_evaluator = Evaluator(best_rf, valid_dataset, transformers)
rf_valid_score = rf_valid_evaluator.compute_model_performance(
[metric], rf_valid_csv_out, rf_valid_stats_out)
print("RF Valid set R^2 %f" % (rf_valid_score["r2_score"]))
rf_test_csv_out = "rf_test_regressor.csv"
rf_test_stats_out = "rf_test_stats_regressor.txt"
rf_test_evaluator = Evaluator(best_rf, test_dataset, transformers)
rf_test_score = rf_test_evaluator.compute_model_performance(
[metric], rf_test_csv_out, rf_test_stats_out)
print("RF Test set R^2 %f" % (rf_test_score["r2_score"]))
rf_crystal_csv_out = "rf_crystal_regressor.csv"
rf_crystal_stats_out = "rf_crystal_stats_regressor.txt"
rf_crystal_evaluator = Evaluator(best_rf, crystal_dataset, transformers)
rf_crystal_score = rf_crystal_evaluator.compute_model_performance(
[metric], rf_crystal_csv_out, rf_crystal_stats_out)
print("RF Crystal set R^2 %f" % (rf_crystal_score["r2_score"]))
# +
dnn_train_csv_out = "dnn_train_regressor.csv"
dnn_train_stats_out = "dnn_train_regressor_stats.txt"
dnn_train_evaluator = Evaluator(best_dnn, train_dataset, transformers)
dnn_train_score = dnn_train_evaluator.compute_model_performance(
[metric], dnn_train_csv_out, dnn_train_stats_out)
print("DNN Train set R^2 %f" % (dnn_train_score["r2_score"]))
dnn_valid_csv_out = "dnn_valid_regressor.csv"
dnn_valid_stats_out = "dnn_valid_regressor_stats.txt"
dnn_valid_evaluator = Evaluator(best_dnn, valid_dataset, transformers)
dnn_valid_score = dnn_valid_evaluator.compute_model_performance(
[metric], dnn_valid_csv_out, dnn_valid_stats_out)
print("DNN Valid set R^2 %f" % (dnn_valid_score["r2_score"]))
dnn_test_csv_out = "dnn_test_regressor.csv"
dnn_test_stats_out = "dnn_test_regressor_stats.txt"
dnn_test_evaluator = Evaluator(best_dnn, test_dataset, transformers)
dnn_test_score = dnn_test_evaluator.compute_model_performance(
[metric], dnn_test_csv_out, dnn_test_stats_out)
print("DNN Test set R^2 %f" % (dnn_test_score["r2_score"]))
dnn_crystal_csv_out = "dnn_crystal_regressor.csv"
dnn_crystal_stats_out = "dnn_crystal_stats_regressor.txt"
dnn_crystal_evaluator = Evaluator(best_dnn, crystal_dataset, transformers)
dnn_crystal_score = dnn_crystal_evaluator.compute_model_performance(
[metric], dnn_crystal_csv_out, dnn_crystal_stats_out)
print("DNN Crystal set R^2 %f" % (dnn_crystal_score["r2_score"]))
# -
task = "pIC50"
rf_predicted_test = best_rf.predict(test_dataset)
rf_true_test = test_dataset.y
plt.scatter(rf_predicted_test, rf_true_test)
plt.xlabel('Predicted pIC50s')
plt.ylabel('Secondary Assay')
plt.title(r'RF predicted IC50 vs. Secondary Assay')
plt.xlim([2, 11])
plt.ylim([2, 11])
plt.plot([2, 11], [2, 11], color='k')
plt.show()
task = "pIC50"
dnn_predicted_test = best_dnn.predict(test_dataset, transformers)
dnn_true_test = test_dataset.y
plt.scatter(dnn_predicted_test, dnn_true_test)
plt.xlabel('Predicted pIC50s')
plt.ylabel('Secondary Assay')
plt.title(r'DNN predicted IC50 vs. Secondary Assay')
plt.xlim([2, 11])
plt.ylim([2, 11])
plt.plot([2, 11], [2, 11], color='k')
plt.show()
| examples/notebooks/BACE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="Auk-JYdPhiZa"
# # ***Block Deep Neural Network-Based Signal Detector for Generalized Spatial Modulation (Training)***
# + [markdown] id="MNZ76w3vetEE"
# This code is the training of the model studied in the paper "Block Deep Neural Network-Based Signal Detector for Generalized Spatial Modulation".
# + [markdown] id="b5ASyoSgiFsm"
# ***Libraries***
# + id="5Z5vzTOHhzDD"
from tensorflow import keras
from tensorflow.keras.layers import Dense, BatchNormalization
from tensorflow.keras.optimizers import SGD
from tensorflow.keras.metrics import Accuracy
import numpy as np
import WirelessCommLib as wcl
import time
# + [markdown] id="pcYHQ-bBiMG8"
# ***Functions***
# + id="AYl9InPCiP85"
# =====================================================================================================
# 1. Deep Learning Model
#
# ARGUMENTS
# 1-) n_x: Number of input nodes (Data Type: int)
# 2-) n_y: Number of output nodes (Data Type: int)
# 3-) n_h_list: List of the number of nodes of each hidden layer (Data Type: numpy.ndarray or list |
# Shape: (3,))
# 4-) Np: Number of active transmit antennas (Data Type: int)
#
# OUTPUT
# - model: Deep learning model (Data Type: tensorflow.python.keras.engine.functional.Functional)
# =====================================================================================================
def Model(n_x, n_y, n_h_list, Np):
num_hidden_layers = len(n_h_list)
input_layer = keras.Input(shape=(n_x,), name="Input Layer")
model_outputs = []
model_losses = []
model_metrics = []
for active_antenna_index in range(Np):
n_h1 = n_h_list[0]
dense_layer_name = "Dense_AAI" + str(active_antenna_index + 1) + "_HLI" + str(1)
hidden_layer = Dense(n_h1, kernel_regularizer=keras.regularizers.l2(l=0.001), activation="relu", name=dense_layer_name)(input_layer)
bn_layer_name = "BatchNormalization_AAI" + str(active_antenna_index + 1) + "_HLI" + str(1)
hidden_layer = BatchNormalization(name=bn_layer_name)(hidden_layer)
for hidden_layer_index in range(1, num_hidden_layers):
n_h = n_h_list[hidden_layer_index]
dense_layer_name = "Dense_AAI" + str(active_antenna_index + 1) + "_HLI" + str(hidden_layer_index + 1)
hidden_layer = Dense(n_h, kernel_regularizer=keras.regularizers.l2(l=0.001), activation="relu", name=dense_layer_name)(hidden_layer)
bn_layer_name = "BatchNormalization_AAI" + str(active_antenna_index + 1) + "_HLI" + str(hidden_layer_index + 1)
hidden_layer = BatchNormalization(name=bn_layer_name)(hidden_layer)
output_layer_name = "Output_AAI" + str(active_antenna_index + 1)
output_layer = Dense(n_y, kernel_regularizer=keras.regularizers.l2(l=0.001), activation="softmax", name=output_layer_name)(hidden_layer)
model_outputs.append(output_layer)
model_losses.append("categorical_crossentropy")
metric_name = "Accuracy" + str(active_antenna_index + 1)
model_metrics.append(Accuracy(name=metric_name))
model = keras.Model(inputs=input_layer, outputs=model_outputs, name="B_DNN_Model")
SGD_optimizer = SGD(lr=0.005, nesterov=True)
model.compile(optimizer=SGD_optimizer, loss=model_losses, metrics=model_metrics)
return model
# =====================================================================================================
# =====================================================================================================
# 2. Number Of Nodes In Each Hidden Layer
#
# ARGUMENT
# - M: Constellation size (Data Type: int | Condition: Power of 2)
#
# OUTPUT
# n_h_list: List of the number of nodes of each hidden layer (Data Type: numpy.ndarray or list | Shape:
# (3,))
# =====================================================================================================
def HiddenLayerNodes(M):
if M == 2:
n_h_list = [128, 64, 32]
elif M == 4:
n_h_list = [256, 128, 64]
elif M == 16:
n_h_list = [512, 256, 128]
return n_h_list
# =====================================================================================================
# + [markdown] id="BnXSzRsFkNAB"
# ***GSM Parameters***
# + id="T4o0XUNpkU3x"
Ns = 15000000 # Number of training time slots
Nt = 2 # Number of transmit antennas
fig_name = "3a"
if fig_name == "3a":
Np = 2 # Number of active transmit antennas
Nr = 2 # Number of receive antennas
M = 4 # Constellation size
mod_type = "PSK" # Modulation type
FVG_type = "SFVG"
elif fig_name == "3b":
Np = 2
Nr = 4
M = 4
mod_type = "PSK"
FVG_type = "SFVG"
elif fig_name == "3c":
Np = 2
Nr = 64
M = 4
mod_type = "PSK"
FVG_type = "SFVG"
elif fig_name == "4a-S":
Np = 2
Nr = 2
M = 2
mod_type = "PSK"
FVG_type = "SFVG"
elif fig_name == "4a-J":
Np = 2
Nr = 2
M = 2
mod_type = "PSK"
FVG_type = "JFVG"
elif fig_name == "4a-C":
Np = 2
Nr = 2
M = 2
mod_type = "PSK"
FVG_type = "CFVG"
elif fig_name == "4b-2":
Np = 2
Nr = 2
M = 2
mod_type = "PSK"
FVG_type = "SFVG"
elif fig_name == "4b-4":
Np = 2
Nr = 2
M = 4
mod_type = "PSK"
FVG_type = "SFVG"
elif fig_name == "4b-16":
Np = 2
Nr = 2
M = 16
mod_type = "QAM"
FVG_type = "SFVG"
else: # Enter your own parameters
Np = 2
Nr = 2
M = 2
mod_type = "PSK"
FVG_type = "SFVG"
N_tot = wcl.Combination(Nt, Np) # Number of total transmit antenna combinations (TACs)
ns = int(np.floor(np.log2(N_tot))) # Number of spatial bits transmitted during a time-slot
m = int(np.log2(M)) # Number of information bits transmitted from a single active antenna during a time-slot
ni = Np * m # Number of total information bits transmitted during a time-slot
n_tot = ns + ni # Number of total bits transmitted during a time-slot
N = 2 ** ns # Number of illegitimate TACs
is_normalized = True
ss = wcl.Constellation(M, mod_type, is_normalized) # Signal set
TAC_set = wcl.OptimalTAC_Set(Nt, Np, N) # Optimal TAC set
# + [markdown] id="5PrCJvxxkdRv"
# ***Data Preprocessing***
# + [markdown] id="gVvsN_VMkidT"
# Input Data
# + id="aDi6O1NYkjv9"
bit_matrix = np.random.randint(2, size=(n_tot, Ns))
if FVG_type == "SFVG":
train_input_data = np.zeros((Ns, 2 * Nr + 2 * Nr * Np))
elif FVG_type == "JFVG":
train_input_data = np.zeros((Ns, 1 + Np * Np))
elif FVG_type == "CFVG":
train_input_data = np.zeros((Ns, Nr + Nr * Np))
for j in range(Ns):
bit_array = bit_matrix[:, j]
x = wcl.EncodeBits(bit_array, ss, TAC_set, ns, m, Nt, Np) # Transmitted vector
H = wcl.Channel([Nr, Nt]) # Rayleigh fading channel
y = np.matmul(H, x) # Received signal vector
train_input_data[j, :] = np.concatenate((wcl.FVG(y, FVG_type), wcl.FVG(H, FVG_type)))[:, 0]
# + [markdown] id="hNNf-iQLkpWo"
# Output Data
# + id="TpB_njTakovK"
train_output_data = []
for active_antenna_index in range(Np):
current_active_antenna_labels = np.zeros((Ns, M))
for j in range(Ns):
start_bit_index = ns + active_antenna_index * m
stop_bit_index = ns + (active_antenna_index + 1) * m
current_time_slot_bits = bit_matrix[start_bit_index : stop_bit_index, j]
current_active_antenna_labels[j, wcl.Bin2Dec(current_time_slot_bits)] = 1
train_output_data.append(current_active_antenna_labels)
# + [markdown] id="E5shoez_ksg2"
# ***Deep Learning Model***
# + id="b9H9Ai__kurc"
if FVG_type == "SFVG":
n_x = 2 * Nr + 2 * Nr * Np
elif FVG_type == "JFVG":
n_x = 1 + Np * Np
elif FVG_type == "CFVG":
n_x = Nr + Nr * Np
n_y = M
n_h_list = HiddenLayerNodes(M)
B_DNN_model = Model(n_x, n_y, n_h_list, Np)
B_DNN_model.summary()
# + [markdown] id="JAqCWzKAkxCE"
# ***Training The Model***
# + id="jFRcrll2k1UW"
start_time = time.time()
B_DNN_model.fit(train_input_data, train_output_data, validation_split=0.25, batch_size=512, epochs=50, shuffle=True)
finish_time = time.time()
training_duration = finish_time - start_time
print("Training Time: ", training_duration, " seconds")
# Enter the path of the "Block-DNN" folder in order to save the model
B_DNN_folder_path = ""
model_path = B_DNN_folder_path + "/Trained Models/"
model_name = "B_DNN_model_Np" + str(Np) + "_Nr" + str(Nr) + "_M" + str(M) + mod_type + "_" + FVG_type + ".h5"
B_DNN_model.save(model_path + model_name)
| B-DNN_Training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.7 64-bit
# language: python
# name: python37764bit9b831f803f76425e8df3192c920ce51f
# ---
# # How to solve optimization problem in Pyleecan
#
# This tutorial explains how to use Pyleecan to solve **constrained global optimization** problem.
#
# The notebook related to this tutorial is available on [GitHub](https://github.com/Eomys/pyleecan/tree/master/Tutorials/tuto_Optimization.ipynb).
#
# The tutorial introduces the different objects to define that enable to parametrize each aspect of the optimization. To do so, we will present an example to maximize the average torque and minimize the first torque harmonic by varying the stator slot opening and the rotor external radius and by adding a constraint on torque ripple.
#
# ## Problem definition
#
# The object [**OptiProblem**](https://www.pyleecan.org/pyleecan.Classes.OptiObjFunc.html) contains all the optimization problem characteristics:
#
# - the output that contains the simulation default parameters
# - the design variable to vary some parameters of the simulation (e.g. input current, topology of the machine)
# - the objective functions to minimize for the simulation
# - some constraints (optional)
#
# ### Default Output definition
#
# To define the problem, we first define an output which contains the default simulation and its results. The optimization process will be based on this output, each evaluation will copy the simulation, set the value of the design variables and run the new simulation.
#
# For this example, we use the simulation defined in the tutorial [How to define a simulation to call FEMM](https://www.pyleecan.org/tuto_Simulation_FEMM.html), but we decrease the precision of the FEMM mesh to speed up the calculations.
# +
# Change of directory to have pyleecan in the path
from os import chdir
chdir('..')
from numpy import ones, pi, array, linspace
from pyleecan.Classes.Simu1 import Simu1
from pyleecan.Classes.Output import Output
from pyleecan.Classes.InputCurrent import InputCurrent
from pyleecan.Classes.MagFEMM import MagFEMM
from pyleecan.Functions.load import load
# Import the machine from a script
IPMSM_A = load('pyleecan/Data/Machine/IPMSM_A.json')
rotor_speed = 2000 # [rpm]
# Create the Simulation
mySimu = Simu1(name="EM_SIPMSM_AL_001", machine=IPMSM_A)
# Defining Simulation Input
mySimu.input = InputCurrent()
# time discretization [s]
mySimu.input.time.value= linspace(start=0, stop=60/rotor_speed, num=16, endpoint=False)# 16 timesteps
# Angular discretization along the airgap circonference for flux density calculation
mySimu.input.angle.value = linspace(start = 0, stop = 2*pi, num=2048, endpoint=False) # 2048 steps
# Rotor speed as a function of time [rpm]
mySimu.input.Nr.value = ones(16) * rotor_speed
# Stator currents as a function of time, each column correspond to one phase [A]
mySimu.input.Is.value = array(
[
[ 1.77000000e+02, -8.85000000e+01, -8.85000000e+01],
[ 5.01400192e-14, -1.53286496e+02, 1.53286496e+02],
[-1.77000000e+02, 8.85000000e+01, 8.85000000e+01],
[-3.25143725e-14, 1.53286496e+02, -1.53286496e+02],
[ 1.77000000e+02, -8.85000000e+01, -8.85000000e+01],
[ 2.11398201e-13, -1.53286496e+02, 1.53286496e+02],
[-1.77000000e+02, 8.85000000e+01, 8.85000000e+01],
[-3.90282030e-13, 1.53286496e+02, -1.53286496e+02],
[ 1.77000000e+02, -8.85000000e+01, -8.85000000e+01],
[ 9.75431176e-14, -1.53286496e+02, 1.53286496e+02],
[-1.77000000e+02, 8.85000000e+01, 8.85000000e+01],
[-4.33634526e-13, 1.53286496e+02, -1.53286496e+02],
[ 1.77000000e+02, -8.85000000e+01, -8.85000000e+01],
[ 4.55310775e-13, -1.53286496e+02, 1.53286496e+02],
[-1.77000000e+02, 8.85000000e+01, 8.85000000e+01],
[-4.76987023e-13, 1.53286496e+02, -1.53286496e+02]
]
)
# Definition of the magnetic simulation (is_mmfr=False => no flux from the magnets)
mySimu.mag = MagFEMM(
type_BH_stator=0, # 0 to use the B(H) curve,
# 1 to use linear B(H) curve according to mur_lin,
# 2 to enforce infinite permeability (mur_lin =100000)
type_BH_rotor=0, # 0 to use the B(H) curve,
# 1 to use linear B(H) curve according to mur_lin,
# 2 to enforce infinite permeability (mur_lin =100000)
angle_stator=0, # Angular position shift of the stator
file_name = "", # Name of the file to save the FEMM model
is_symmetry_a=True, # 0 Compute on the complete machine, 1 compute according to sym_a and is_antiper_a
sym_a = 4, # Number of symmetry for the angle vector
is_antiper_a=True, # To add an antiperiodicity to the angle vector
Kmesh_fineness = 0.2, # Decrease mesh precision
Kgeo_fineness = 0.2, # Decrease mesh precision
)
# We only use the magnetic part
mySimu.force = None
mySimu.struct = None
# Set the default output for the optimization
defaultOutput = Output(simu=mySimu)
# -
# ### Minimization problem definition
#
# To setup the optimization problem, we define some objective functions using the [**OptiObjFunc**](https://www.pyleecan.org/pyleecan.Classes.OptiObjFunc.html) object.
#
# Each objective function takes an output object in argument and returns a float to **minimize**.
# We gather the objective functions into a dictionnary.
# +
from pyleecan.Classes.OptiObjFunc import OptiObjFunc
import numpy as np
# Objective functions
def tem_av(output):
"""Return the average torque opposite (opposite to be maximized)"""
return -abs(output.mag.Tem_av)
def harm1(output):
"""Return the first torque harmonic """
N = output.mag.time.size
# Compute the real fft of the torque
sp = 2 / N * np.abs(np.fft.rfft(output.mag.Tem))
# Return the first torque harmonic
return sp[1]
my_obj = {
"Opposite average torque (Nm)": OptiObjFunc(
description="Maximization of the average torque", func=tem_av
),
"First torque harmonic (Nm)": OptiObjFunc(
description="Minimization of the first torque harmonic", func=harm1
),
}
# -
# ### Design variables
# We use the object [**OptiDesignVar**](https://www.pyleecan.org/pyleecan.Classes.OptiDesignVar.html) to define the design variables.
#
#
# To define a design variable, we have to specify different attributes:
#
# - *name* to access to the variable in the output object. This attribute **must begin by "output"**.
# - *type_var* to specify the variable "type":
# - *interval* for continuous variables
# - *set* for discrete variables
# - *space* to set the variable bound
# - *function* to define the variable for the first generation, the function takes the space in argument and returns the variable value
#
# We store the design variables in a dictionnary that will be in argument of the problem. For this example, we define two design variables:
#
# 1. Stator slot opening: can be any value between 0 and the slot width.
# 2. Rotor external radius: can be one of the four value specified \[99.8%, 99.9%, 100%, 100.1%\] of the default rotor external radius
# +
from pyleecan.Classes.OptiDesignVar import OptiDesignVar
import random
# Design variables
my_design_var = {
"Stator slot opening": OptiDesignVar(
name="output.simu.machine.stator.slot.W0",
type_var="interval",
space=[
0 * defaultOutput.simu.machine.stator.slot.W2,
defaultOutput.simu.machine.stator.slot.W2,
],
function=lambda space: random.uniform(*space),
),
"Rotor ext radius": OptiDesignVar(
name="output.simu.machine.rotor.Rext",
type_var="set",
space=[
0.998 * defaultOutput.simu.machine.rotor.Rext,
0.999 * defaultOutput.simu.machine.rotor.Rext,
defaultOutput.simu.machine.rotor.Rext,
1.001 * defaultOutput.simu.machine.rotor.Rext,
],
function=lambda space: random.choice(space),
),
}
# -
# ### Constraints
#
# The class [**OptiConstraint**](https://www.pyleecan.org/pyleecan.Classes.OptiConstraint.html) enables to define some constraint. For each constraint, we have to define the following attributes:
#
# - name
# - type_const: type of constraint
# - "=="
# - "<="
# - "<"
# - ">="
# - ">"
# - value: value to compare
# - get_variable: function which takes output in argument and returns the constraint value
#
# We also store the constraints into a dict.
from pyleecan.Classes.OptiConstraint import OptiConstraint
my_constraint = dict(
const1 = OptiConstraint(
name = "const1",
type_const = "<=",
value = 2.17,
get_variable = lambda output: abs(output.mag.Tem_rip),
)
)
# ### Evaluation function
#
#
# We can create our own evaluation function if needed by defining a function which only take an output in argument.
#
# For this example we keep the default one which calls the `Output.simu.run` method.
# +
from pyleecan.Classes.OptiProblem import OptiProblem
# Problem creation
my_prob = OptiProblem(
output=defaultOutput,
design_var=my_design_var,
obj_func=my_obj,
constraint = my_constraint,
eval_func = None # To keep the default evaluation function
)
# -
# ## Solver
#
# The class [**OptiGenAlgNsga2**](https://www.pyleecan.org/pyleecan.Classes.OptiGenAlgNsga2Deap.html) enables to solve our problem using [NSGA-II](https://www.iitk.ac.in/kangal/Deb_NSGA-II.pdf) genetical algorithm. The algorithm takes several parameters:
#
# |Parameter|Description|Type|Default Value|
# | :-: | :- | :-: | :-: |
# |*problem*|Problem to solve|**OptiProblem**|mandatory|
# |*size\_pop*| Population size per generation|**int**|40|
# |*nb\_gen*|Generation number|**int**|100|
# |*p\_cross*|Crossover probability|**float**|0.9|
# |*p\_mutate*|Mutation probability|**float**|0.1|
#
#
# The `solve` method performs the optimization and returns an [**OutputMultiOpti**](https://www.pyleecan.org/pyleecan.Classes.OutputMultiOpti.html) object which contains the results.
# +
from pyleecan.Classes.OptiGenAlgNsga2Deap import OptiGenAlgNsga2Deap
# Solve problem with NSGA-II
solver = OptiGenAlgNsga2Deap(problem=my_prob, size_pop=16, nb_gen=8, p_mutate=0.5)
res = solver.solve()
# -
# During the algorithm the object displays some data containing:
#
# - number of errors: failure during the objective function execution
# - number of infeasible: number of individual with constraints violations
# ## Plot results
#
# **OutputMultiOpti** has several methods to display some results:
#
# - `plot_generation`: to plot individuals for two objective functions
# - `plot_pareto_front`: to plot the pareto front for two objective functions
# - `plot_pareto_front_design_space`: to plot the pareto front in the design space according to two design variables
# - `plot_generation_design_space`: to plot individuals in the design space according to two design variables
#
# %matplotlib notebook
res.plot_generation(
obj1 = "Opposite average torque (Nm)", #label of the first objective function
obj2 = "First torque harmonic (Nm)", #label of the second objective function
)
# %matplotlib notebook
res.plot_generation_design_space(
dvar1 = "Rotor ext radius",
dvar2 = "Stator slot opening"
)
# %matplotlib notebook
res.plot_pareto(
obj1 = "Opposite average torque (Nm)", #label of the first objective function
obj2 = "First torque harmonic (Nm)", #label of the second objective function
)
res.plot_pareto_design_space(
dvar1 = "Rotor ext radius",
dvar2 = "Stator slot opening"
)
| Tutorials/tuto_Optimization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.DataFrame({'col1': [0, pd.np.nan, pd.np.nan, 3, 4],
'col2': [pd.np.nan, 1, 2, pd.np.nan, pd.np.nan],
'col3': [4, pd.np.nan, pd.np.nan, 7, 10]})
print(df)
print(df.interpolate())
print(df.interpolate(axis=1))
print(df.interpolate(limit=1))
print(df.interpolate(limit=1, limit_direction='forward'))
print(df.interpolate(limit=1, limit_direction='backward'))
print(df.interpolate(limit=1, limit_direction='both'))
print(df.interpolate(limit_direction='both'))
print(df.interpolate(limit_area='inside'))
print(df.interpolate(limit_area='outside'))
print(df.interpolate(limit_area='outside', limit_direction='both'))
df_copy = df.copy()
df_copy.interpolate(inplace=True)
print(df_copy)
s = pd.Series([0, pd.np.nan, pd.np.nan, pd.np.nan, 4, pd.np.nan, pd.np.nan],
index=[0, 2, 5, 6, 8, 10, 14])
print(s)
print(s.interpolate())
print(s.interpolate('index'))
print(s.interpolate('values'))
s.index = list('abcdefg')
print(s)
print(s.interpolate())
# +
# print(s.interpolate('values'))
# TypeError: Cannot cast array data from dtype('O') to dtype('float64') according to the rule 'safe'
# -
s = pd.Series([0, 10, pd.np.nan, pd.np.nan, 4, pd.np.nan, pd.np.nan],
index=[0, 2, 5, 6, 8, 10, 14])
print(s.interpolate('spline', order=2))
s.index = range(7)
print(s.interpolate('spline', order=2))
s.index = list('abcdefg')
# +
# print(s.interpolate('spline', order=2))
# TypeError: unsupported operand type(s) for -: 'str' and 'str'
| notebook/pandas_interpolate.ipynb |
# +
def check_goldbach_for_num(n,primes_set) :
'''gets an even integer- n, and a set of primes- primes_set. Returns whether there're two primes which their sum is n'''
relevant_primes_set={p for p in primes_set if p<n}
for prime in primes_set :
if (n-prime) in relevant_primes_set:
return True
return False
# -
from sympy.ntheory.generate import primerange
primes = set(primerange(50,1000))
check_goldbach_for_num(116, primes)
| demo/goldbach/submissions/2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding: utf-8 -*-
"""
Spyder Editor
This is a temporary script file.
"""
import numpy as np
import pandas as pd
from sklearn.metrics import mean_squared_error
file = pd.read_csv('BlackFriday.csv')
# +
#printing dataset
#print(file)
file.info()
# -
file.describe()
# +
file.isnull().any().sum()
file.shape
total_miss = file.isnull().sum()
perc_miss = total_miss/file.isnull().count()*100
missing_data = pd.DataFrame({'Total missing':total_miss,
'% missing':perc_miss})
missing_data.sort_values(by='Total missing',
ascending=False).head(3)
# -
# distribution of Amount
import seaborn as sns
amount = [file['Purchase'].values]
sns.distplot(amount)
# +
import matplotlib.pyplot as plt
import seaborn as sns
spent_byage = file.groupby(by='Age').sum()['Purchase']
plt.figure(figsize=(12,6))
sns.barplot(x=spent_byage.index,y=spent_byage.values, palette="Blues_d")
plt.title('Mean Purchases per Age Group')
plt.show()
# -
for column in file.columns:
print(column)
print(len(file[column][file[column].isna()]))
data = file.fillna(0)
for column in data.columns:
print(column)
print(len(data[column][data[column].isna()]))
Y = data['Purchase']
# +
X = data.drop(columns=['Purchase','User_ID', 'Product_ID'])#,'Age', 'User_ID', 'Product_ID','Gender','City_Category', 'Occupation', 'Product_Category_1','Product_Category_2','Product_Category_3'])
X_original = X
# -
len(pd.unique(data['User_ID'])), len(pd.unique(data['Product_ID']))
# +
def run_model(model, X_train, Y_train, X_test = None, Y_test = None):
print("Model: ", model.__class__.__name__)
model.fit(X_train, Y_train)
#model.coef_
#a= model.predict(X_test)
print("Training accuracy", model.score(X_train, Y_train))
#print(a)
if X_test is not None and Y_test is not None:
print("Testing accuracy", model.score(X_test, Y_test))
# -
coef5=pd.Series(model.feature_importances_, X_original.columns).sort_values(ascending=False)
coef5.plot(kind='bar',title='Feature Importances')
params = {'n_estimators': 500, 'max_depth': 4, 'min_samples_split': 2,
'learning_rate': 0.01, 'loss': 'ls'}
def preprocess(X,Y,split=True):
if 'Age' in X.columns:
X['Min_Age'], X['Max_Age'] = X['Age'].str.split('-', 1).str
X['Max_Age'].fillna('100', inplace=True)
X['Min_Age'] = X['Min_Age'].str.replace('+', '')
X['Min_Age'] = pd.to_numeric(X['Min_Age'])
X['Max_Age'] = pd.to_numeric(X['Max_Age'])
X = X.drop(columns=['Age'])
if 'Stay_In_Current_City_Years' in X.columns:
X['Stay_In_Current_City_Years'] = X['Stay_In_Current_City_Years'].str.replace('+', '')
X['Stay_In_Current_City_Years'] = pd.to_numeric(X['Stay_In_Current_City_Years'])
for column in ['Occupation', 'Marital_Status', 'Product_Category_1', 'Product_Category_2', 'Product_Category_3']:
if column in X.columns:
X[column] = pd.to_numeric(X[column])
X.index.name = 'key'
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
le = LabelEncoder()
if 'Gender' in X.columns:
X.Gender = le.fit_transform(X.Gender)
if 'City_Category' in X.columns:
X.City_Category = le.fit_transform(X.City_Category)
for column in ['Gender', 'City_Category', 'Occupation', 'Product_Category_1', 'Product_Category_2', 'Product_Category_3']:
if column in X.columns:
# Get one hot encoding of columns B
#print(column)
one_hot = pd.get_dummies(X[column])
one_hot.index.name='key'#, inplace=True)
# Drop column B as it is now encoded
X = X.drop(column, axis = 1)
# Join the encoded df
X = X.join(one_hot, on='key', how='left', lsuffix='_left', rsuffix='_right')
#print("done")
from sklearn.model_selection import train_test_split
if split:
return train_test_split(X, Y, test_size=.20, random_state=42)
else:
return X,Y
# +
for column in X_original.columns:
print("Trying with column", column)
singleX = X_original[column]
X = pd.DataFrame(singleX)
X_train, X_test, y_train, y_test = preprocess(X,Y)
from sklearn.metrics import mean_squared_error
from sklearn.linear_model import LinearRegression
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.linear_model import Ridge
params = {'n_estimators': 500, 'max_depth': 4, 'min_samples_split': 2,
'learning_rate': 0.01, 'loss': 'ls'}
for model in [LinearRegression(normalize=True), DecisionTreeRegressor(), GradientBoostingRegressor(**params), Ridge(alpha=0.05,normalize=True)]:
run_model(model, X_train, y_train, X_test, y_test)
mse = mean_squared_error(y_test, model.predict(X_test))
rmse = np.sqrt(mse)
predicted_label = model.predict(X_test)
print("predicted labels =",predicted_label)
print("RootMeanSquareError =",rmse)
# +
for column in X_original.columns:
print("Trying without column", column)
X = X_original.drop(columns=[column])
from sklearn.linear_model import LinearRegression
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import GradientBoostingRegressor
for model in [LinearRegression(normalize=True), DecisionTreeRegressor(), GradientBoostingRegressor(**params),Ridge(alpha=0.05,normalize=True)]:
run_model(model, X_train, y_train, X_test, y_test)
mse = mean_squared_error(y_test, model.predict(X_test))
rmse = np.sqrt(mse)
predicted_label = model.predict(X_test)
print("predicted labels =",predicted_label)
print("RootMeanSquareError =",rmse)
print()
# -
for column in X_original.columns:
print("Trying with column", column)
singleX = X_original[column]
X = pd.DataFrame(singleX)
X_train, X_test, y_train, y_test =preprocess(X,Y)
from sklearn.linear_model import LinearRegression
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import GradientBoostingRegressor
for model in [ GradientBoostingRegressor()]:
run_model(model, X_train, y_train, X_test, y_test)
print()
# +
X,y = preprocess(X_original,Y,False)
from sklearn.linear_model import Ridge
alg2=Ridge(alpha=0.05,normalize=True)
from sklearn.metrics import mean_squared_error
from math import sqrt
from sklearn.model_selection import train_test_split
from sklearn.model_selection import train_test_split
from sklearn.cross_validation import cross_val_score
from sklearn.model_selection import KFold
s_avg = 0
k_fold = KFold(n_splits=10, shuffle=False, random_state=43)
for train_index, test_index in k_fold.split(X,y):
print(len(X_train), len(X_test))
X_train, X_test = X.iloc[train_index], X.iloc[test_index]
y_train, y_test = y.iloc[train_index], y.iloc[test_index]
alg2.fit(X_train,y_train)
original_values=y_test
predicted=alg2.predict(X_test)
s = alg2.score(X_test, y_test)
print(s)
s_avg = s_avg + s
s_avg = s_avg/10
print(s_avg)
# -
X
# +
model_DTR = DecisionTreeRegressor()
X_train, X_test, y_train, y_test = preprocess(X_original, Y)
model_DTR.fit(X_train, y_train)
mse = mean_squared_error(y_test, model_DTR.predict(X_test))
rmse = np.sqrt(mse)
predicted_label = model_DTR.predict(X_test)
print("predicted labels =",predicted_label)
print("RootMeanSquareError =",rmse)
model_DTR.score(X_train, y_train), model_DTR.score(X_test, y_test)
# +
model_GBR = GradientBoostingRegressor()
X_train, X_test, y_train, y_test= preprocess(X_original, Y)
model_GBR.fit(X_train, y_train)
mse = mean_squared_error(y_test, model_GBR.predict(X_test))
rmse = np.sqrt(mse)
predicted_label = model_GBR.predict(X_test)
print("predicted labels =",predicted_label)
print("RootMeanSquareError =",rmse)
model_GBR.score(X_train, y_train), model_GBR.score(X_test, y_test)
model_GBR.score(X_train, y_train), model_GBR.score(X_test, y_test)
# -
| ML_Project4/Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 0.09204, "end_time": "2021-03-22T14:46:30.902459", "exception": false, "start_time": "2021-03-22T14:46:30.810419", "status": "completed"} tags=[]
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# -
pip install tensorflow
# + papermill={"duration": 5.953084, "end_time": "2021-03-22T14:46:36.901990", "exception": false, "start_time": "2021-03-22T14:46:30.948906", "status": "completed"} tags=[]
import tensorflow as tf
print("Num GPUs Available: ", len(tf.config.list_physical_devices('GPU')))
# + papermill={"duration": 19.48794, "end_time": "2021-03-22T14:46:56.488893", "exception": false, "start_time": "2021-03-22T14:46:37.000953", "status": "completed"} tags=[]
# !pip install bs4
# !pip install contractions
# + papermill={"duration": 0.56967, "end_time": "2021-03-22T14:46:57.115166", "exception": false, "start_time": "2021-03-22T14:46:56.545496", "status": "completed"} tags=[]
import seaborn as sns
import matplotlib.pyplot as plt
# + papermill={"duration": 7.128261, "end_time": "2021-03-22T14:47:04.298072", "exception": false, "start_time": "2021-03-22T14:46:57.169811", "status": "completed"} tags=[]
reviews = pd.read_csv('../dataset/Reviews.csv')
# + papermill={"duration": 0.060369, "end_time": "2021-03-22T14:47:04.412849", "exception": false, "start_time": "2021-03-22T14:47:04.352480", "status": "completed"} tags=[]
# reviews = reviews.iloc[:20000]
# + papermill={"duration": 0.082449, "end_time": "2021-03-22T14:47:04.548876", "exception": false, "start_time": "2021-03-22T14:47:04.466427", "status": "completed"} tags=[]
reviews.head()
# + papermill={"duration": 0.064836, "end_time": "2021-03-22T14:47:04.667865", "exception": false, "start_time": "2021-03-22T14:47:04.603029", "status": "completed"} tags=[]
reviews.iloc[10].Text
# + papermill={"duration": 0.065477, "end_time": "2021-03-22T14:47:04.787856", "exception": false, "start_time": "2021-03-22T14:47:04.722379", "status": "completed"} tags=[]
reviews.iloc[10].Summary
# + papermill={"duration": 0.356246, "end_time": "2021-03-22T14:47:05.199041", "exception": false, "start_time": "2021-03-22T14:47:04.842795", "status": "completed"} tags=[]
reviews.info()
# + [markdown] papermill={"duration": 0.056258, "end_time": "2021-03-22T14:47:05.312243", "exception": false, "start_time": "2021-03-22T14:47:05.255985", "status": "completed"} tags=[]
# ## Get Total unique Products
# + papermill={"duration": 1.156218, "end_time": "2021-03-22T14:47:06.524458", "exception": false, "start_time": "2021-03-22T14:47:05.368240", "status": "completed"} tags=[]
reviews.nunique()
# + papermill={"duration": 0.360055, "end_time": "2021-03-22T14:47:06.940487", "exception": false, "start_time": "2021-03-22T14:47:06.580432", "status": "completed"} tags=[]
reviews.isnull().sum()
# ProfileName doesnt matter since UserId and ProfileName is same. So If UserId is present then we should not care about ProfileName
# + papermill={"duration": 0.120211, "end_time": "2021-03-22T14:47:07.118268", "exception": false, "start_time": "2021-03-22T14:47:06.998057", "status": "completed"} tags=[]
# users who didnt provide Summary
reviews[reviews.Summary.isnull()].UserId.value_counts()
# + papermill={"duration": 0.071409, "end_time": "2021-03-22T14:47:07.246885", "exception": false, "start_time": "2021-03-22T14:47:07.175476", "status": "completed"} tags=[]
# How many users provided, respective ratings
reviews.Score.value_counts()
# + papermill={"duration": 0.223106, "end_time": "2021-03-22T14:47:07.527749", "exception": false, "start_time": "2021-03-22T14:47:07.304643", "status": "completed"} tags=[]
reviews.Score.value_counts().plot.bar()
# + papermill={"duration": 0.066539, "end_time": "2021-03-22T14:47:07.652747", "exception": false, "start_time": "2021-03-22T14:47:07.586208", "status": "completed"} tags=[]
# Most of the reviews are positive, So ppl do like food , Average of the Score is 4.1
reviews.Score.mean()
# + papermill={"duration": 0.845977, "end_time": "2021-03-22T14:47:08.559131", "exception": false, "start_time": "2021-03-22T14:47:07.713154", "status": "completed"} tags=[]
import datetime
lol = lambda x: datetime.datetime.fromtimestamp(x)
reviews['Date'] = reviews.Time.apply(lol)
# + papermill={"duration": 0.352345, "end_time": "2021-03-22T14:47:08.986703", "exception": false, "start_time": "2021-03-22T14:47:08.634358", "status": "completed"} tags=[]
reviews['weekDay_Time'] = reviews.Date.dt.weekday
reviews['month'] = reviews.Date.dt.month
reviews['quarter'] = reviews.Date.dt.quarter
# + papermill={"duration": 0.143688, "end_time": "2021-03-22T14:47:09.244275", "exception": false, "start_time": "2021-03-22T14:47:09.100587", "status": "completed"} tags=[]
reviews.head()
# + papermill={"duration": 0.602436, "end_time": "2021-03-22T14:47:09.976042", "exception": false, "start_time": "2021-03-22T14:47:09.373606", "status": "completed"} tags=[]
sns.countplot(data=reviews, x='weekDay_Time')
# + papermill={"duration": 0.260265, "end_time": "2021-03-22T14:47:10.321182", "exception": false, "start_time": "2021-03-22T14:47:10.060917", "status": "completed"} tags=[]
sns.countplot(data=reviews, x='month')
# + papermill={"duration": 0.209935, "end_time": "2021-03-22T14:47:10.592183", "exception": false, "start_time": "2021-03-22T14:47:10.382248", "status": "completed"} tags=[]
sns.countplot(data=reviews, x='quarter')
# + papermill={"duration": 0.289147, "end_time": "2021-03-22T14:47:10.943426", "exception": false, "start_time": "2021-03-22T14:47:10.654279", "status": "completed"} tags=[]
reviews = reviews[~reviews.Summary.isnull()]
reviews=reviews.reset_index(drop=True)
# + [markdown] papermill={"duration": 0.06246, "end_time": "2021-03-22T14:47:11.068729", "exception": false, "start_time": "2021-03-22T14:47:11.006269", "status": "completed"} tags=[]
# ## Lets do Some EDA on the Text Data
# + [markdown] papermill={"duration": 0.062193, "end_time": "2021-03-22T14:47:11.193975", "exception": false, "start_time": "2021-03-22T14:47:11.131782", "status": "completed"} tags=[]
# ### Lets preprocess convert everything to Lower Case
# + papermill={"duration": 1.067023, "end_time": "2021-03-22T14:47:12.323362", "exception": false, "start_time": "2021-03-22T14:47:11.256339", "status": "completed"} tags=[]
reviews.Text = reviews.Text.str.lower()
reviews.Summary = reviews.Summary.str.lower()
# + papermill={"duration": 0.081201, "end_time": "2021-03-22T14:47:12.467671", "exception": false, "start_time": "2021-03-22T14:47:12.386470", "status": "completed"} tags=[]
reviews.head()
# + papermill={"duration": 0.069855, "end_time": "2021-03-22T14:47:12.600302", "exception": false, "start_time": "2021-03-22T14:47:12.530447", "status": "completed"} tags=[]
Text_data = reviews.Text.values
Summary = reviews.Summary.values
# + [markdown] papermill={"duration": 0.063068, "end_time": "2021-03-22T14:47:12.726546", "exception": false, "start_time": "2021-03-22T14:47:12.663478", "status": "completed"} tags=[]
# ## do we have similar Summariaes?
# + papermill={"duration": 0.348606, "end_time": "2021-03-22T14:47:13.139690", "exception": false, "start_time": "2021-03-22T14:47:12.791084", "status": "completed"} tags=[]
reviews[reviews.Text.duplicated()]
# + papermill={"duration": 0.173626, "end_time": "2021-03-22T14:47:13.378363", "exception": false, "start_time": "2021-03-22T14:47:13.204737", "status": "completed"} tags=[]
Total_len = len(Summary)
Total_distinct_Summary = len(set(Summary))
print(Total_len)
print(Total_distinct_Summary)
# + [markdown] papermill={"duration": 0.064303, "end_time": "2021-03-22T14:47:13.509232", "exception": false, "start_time": "2021-03-22T14:47:13.444929", "status": "completed"} tags=[]
# ## do we have similar Text Data?
# + papermill={"duration": 0.198671, "end_time": "2021-03-22T14:47:13.771883", "exception": false, "start_time": "2021-03-22T14:47:13.573212", "status": "completed"} tags=[]
Total_len = len(Text_data)
Total_distinct_Text_data = len(set(Text_data))
print(Total_len)
print(Total_distinct_Text_data)
# + [markdown] papermill={"duration": 0.064786, "end_time": "2021-03-22T14:47:13.903803", "exception": false, "start_time": "2021-03-22T14:47:13.839017", "status": "completed"} tags=[]
# ## ok lets drop the similar Text Data
# + papermill={"duration": 0.757604, "end_time": "2021-03-22T14:47:14.726438", "exception": false, "start_time": "2021-03-22T14:47:13.968834", "status": "completed"} tags=[]
reviews = reviews.drop_duplicates('Text')
reviews = reviews.reset_index(drop=True)
# + papermill={"duration": 0.074586, "end_time": "2021-03-22T14:47:14.868766", "exception": false, "start_time": "2021-03-22T14:47:14.794180", "status": "completed"} tags=[]
Text_data = reviews.Text.values
Summary = reviews.Summary.values
# -
pip install wordcloud
# + papermill={"duration": 7.511038, "end_time": "2021-03-22T14:47:22.444296", "exception": false, "start_time": "2021-03-22T14:47:14.933258", "status": "completed"} tags=[]
from wordcloud import WordCloud, STOPWORDS
summaryWordCloud = ' '.join(Summary).lower()
wordcloud2 = WordCloud().generate(summaryWordCloud)
plt.imshow(wordcloud2)
# + papermill={"duration": 0.074169, "end_time": "2021-03-22T14:47:22.586770", "exception": false, "start_time": "2021-03-22T14:47:22.512601", "status": "completed"} tags=[]
# TextWordCloud = ' '.join(Text_data).lower()
# wordcloud2 = WordCloud().generate(TextWordCloud)
# plt.imshow(wordcloud2)
# + [markdown] papermill={"duration": 0.06727, "end_time": "2021-03-22T14:47:22.721756", "exception": false, "start_time": "2021-03-22T14:47:22.654486", "status": "completed"} tags=[]
# ## Word Cloud for our Data. Looks COOL
# + [markdown] papermill={"duration": 0.067936, "end_time": "2021-03-22T14:47:22.857350", "exception": false, "start_time": "2021-03-22T14:47:22.789414", "status": "completed"} tags=[]
# ## Remove HTML Tags
# + papermill={"duration": 88.166366, "end_time": "2021-03-22T14:48:51.091608", "exception": false, "start_time": "2021-03-22T14:47:22.925242", "status": "completed"} tags=[]
from bs4 import BeautifulSoup
soup = lambda text:BeautifulSoup(text)
passText = lambda text:soup(text).get_text()
reviews.Text = reviews.Text.apply(passText)
# + [markdown] papermill={"duration": 0.117915, "end_time": "2021-03-22T14:48:51.327728", "exception": false, "start_time": "2021-03-22T14:48:51.209813", "status": "completed"} tags=[]
# ## Histogram for the length of reviews and summary
# + papermill={"duration": 0.562706, "end_time": "2021-03-22T14:48:52.005945", "exception": false, "start_time": "2021-03-22T14:48:51.443239", "status": "completed"} tags=[]
reviews.Summary.apply(lambda x:len(x.split(' '))).plot(kind='hist')
# + papermill={"duration": 2.047915, "end_time": "2021-03-22T14:48:54.128460", "exception": false, "start_time": "2021-03-22T14:48:52.080545", "status": "completed"} tags=[]
reviews.Text.apply(lambda x:len(x.split(' '))).plot(kind='hist')
# + papermill={"duration": 1.879591, "end_time": "2021-03-22T14:48:56.082854", "exception": false, "start_time": "2021-03-22T14:48:54.203263", "status": "completed"} tags=[]
reviews.Text.apply(lambda x:len(x.split(' '))).quantile(0.95)
# + papermill={"duration": 0.079417, "end_time": "2021-03-22T14:48:56.235356", "exception": false, "start_time": "2021-03-22T14:48:56.155939", "status": "completed"} tags=[]
input_characters = set()
target_characters = set()
# + papermill={"duration": 0.077767, "end_time": "2021-03-22T14:48:56.389808", "exception": false, "start_time": "2021-03-22T14:48:56.312041", "status": "completed"} tags=[]
import re
# + papermill={"duration": 0.745965, "end_time": "2021-03-22T14:48:57.205689", "exception": false, "start_time": "2021-03-22T14:48:56.459724", "status": "completed"} tags=[]
from nltk.tokenize import TweetTokenizer
tweet = TweetTokenizer()
# -
pip install contractions
# + papermill={"duration": 0.084076, "end_time": "2021-03-22T14:48:57.360855", "exception": false, "start_time": "2021-03-22T14:48:57.276779", "status": "completed"} tags=[]
import contradiction
# + papermill={"duration": 10.663715, "end_time": "2021-03-22T14:49:08.095128", "exception": false, "start_time": "2021-03-22T14:48:57.431413", "status": "completed"} tags=[]
sampleReview = reviews.iloc[:20000].copy()
#Fix Contradiction
sampleReview.Text = sampleReview.Text.apply(lambda x: contractions.fix(x))
sampleReview.Summary = sampleReview.Summary.apply(lambda x: contractions.fix(x))
sampleReview.Text = sampleReview.Text.apply(lambda x: tweet.tokenize(x))
sampleReview.Summary = sampleReview.Summary.apply(lambda x: tweet.tokenize(x))
# + papermill={"duration": 0.094154, "end_time": "2021-03-22T14:49:08.261323", "exception": false, "start_time": "2021-03-22T14:49:08.167169", "status": "completed"} tags=[]
sampleReview.head()
# + papermill={"duration": 0.519446, "end_time": "2021-03-22T14:49:08.852227", "exception": false, "start_time": "2021-03-22T14:49:08.332781", "status": "completed"} tags=[]
sampleReview.Text = sampleReview.Text.apply(lambda tokens: [word for word in tokens if word.isalpha()])
sampleReview.Summary = sampleReview.Summary.apply(lambda tokens: [word for word in tokens if word.isalpha()])
# -
pip install nltk
pip install stopwords
# + papermill={"duration": 0.100687, "end_time": "2021-03-22T14:49:09.024918", "exception": false, "start_time": "2021-03-22T14:49:08.924231", "status": "completed"} tags=[]
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
stop_words = stopwords.words()
stop_words = set(stop_words)
# + papermill={"duration": 0.07936, "end_time": "2021-03-22T14:49:09.175874", "exception": false, "start_time": "2021-03-22T14:49:09.096514", "status": "completed"} tags=[]
len(sampleReview.Text)
# + papermill={"duration": 0.304213, "end_time": "2021-03-22T14:49:09.551408", "exception": false, "start_time": "2021-03-22T14:49:09.247195", "status": "completed"} tags=[]
# %%time
sampleReview.Text = sampleReview.Text.apply(lambda tokens: [word for word in tokens if word not in stop_words][:100])
# + papermill={"duration": 0.120262, "end_time": "2021-03-22T14:49:09.744516", "exception": false, "start_time": "2021-03-22T14:49:09.624254", "status": "completed"} tags=[]
sampleReview.head()
# + papermill={"duration": 0.126352, "end_time": "2021-03-22T14:49:09.997920", "exception": false, "start_time": "2021-03-22T14:49:09.871568", "status": "completed"} tags=[]
# from nltk.corpus import wordnet
# sampleReview.Text = sampleReview.Text.apply(lambda tokens: [word for word in tokens if wordnet.synsets(word)])
# sampleReview.Summary = sampleReview.Summary.apply(lambda tokens: [word for word in tokens if wordnet.synsets(word)])
# + papermill={"duration": 0.161517, "end_time": "2021-03-22T14:49:10.282806", "exception": false, "start_time": "2021-03-22T14:49:10.121289", "status": "completed"} tags=[]
sampleReview.Summary = sampleReview.Summary.apply(lambda x:['<BOS>']+x+['<EOS>'])
# + papermill={"duration": 0.151327, "end_time": "2021-03-22T14:49:10.523162", "exception": false, "start_time": "2021-03-22T14:49:10.371835", "status": "completed"} tags=[]
sampleReview.Text = sampleReview.Text.apply(lambda x:' '.join(x))
sampleReview.Summary = sampleReview.Summary.apply(lambda x:' '.join(x))
# + papermill={"duration": 0.083229, "end_time": "2021-03-22T14:49:10.680444", "exception": false, "start_time": "2021-03-22T14:49:10.597215", "status": "completed"} tags=[]
sampleReview.Summary
# + papermill={"duration": 0.082649, "end_time": "2021-03-22T14:49:10.837805", "exception": false, "start_time": "2021-03-22T14:49:10.755156", "status": "completed"} tags=[]
input_texts = sampleReview.Text.values
target_texts = list(sampleReview.Summary.values)
# + papermill={"duration": 0.596274, "end_time": "2021-03-22T14:49:11.510213", "exception": false, "start_time": "2021-03-22T14:49:10.913939", "status": "completed"} tags=[]
sampleReview.to_csv('filtered_data.csv', index=False)
# -
pip install keras
# + papermill={"duration": 1.794354, "end_time": "2021-03-22T14:49:13.379591", "exception": false, "start_time": "2021-03-22T14:49:11.585237", "status": "completed"} tags=[]
from keras.preprocessing.text import Tokenizer
VOCAB_SIZE = 50000
tokenizerText = Tokenizer(num_words=VOCAB_SIZE, oov_token='<OOV>')
tokenizerText.fit_on_texts(input_texts)
tokenizerSummary = Tokenizer(num_words=VOCAB_SIZE, oov_token='<OOV>')
tokenizerSummary.fit_on_texts(target_texts)
def text2seq(encoder_text, decoder_text, VOCAB_SIZE):
encoder_sequences = tokenizerText.texts_to_sequences(encoder_text)
decoder_sequences = tokenizerSummary.texts_to_sequences(decoder_text)
return encoder_sequences, decoder_sequences
encoder_sequences, decoder_sequences = text2seq(input_texts, target_texts, VOCAB_SIZE)
# + papermill={"duration": 0.085631, "end_time": "2021-03-22T14:49:13.541054", "exception": false, "start_time": "2021-03-22T14:49:13.455423", "status": "completed"} tags=[]
textVocabSize = len(tokenizerText.word_index)
summaryVocabSize = len(tokenizerSummary.word_index)
textVocabSize, summaryVocabSize
# + papermill={"duration": 0.102491, "end_time": "2021-03-22T14:49:13.718134", "exception": false, "start_time": "2021-03-22T14:49:13.615643", "status": "completed"} tags=[]
def vocab_creater(text_lists, VOCAB_SIZE, tokenizer):
dictionary = tokenizer.word_index
word2idx = {}
idx2word = {}
for k, v in dictionary.items():
if v < VOCAB_SIZE:
word2idx[k] = v
idx2word[v] = k
if v >= VOCAB_SIZE-1:
continue
return word2idx, idx2word
word2idxText, idx2wordText = vocab_creater(input_texts, textVocabSize, tokenizerText)
word2idxSummary, idx2wordSummary = vocab_creater(target_texts, summaryVocabSize, tokenizerSummary)
# + papermill={"duration": 0.081808, "end_time": "2021-03-22T14:49:13.875281", "exception": false, "start_time": "2021-03-22T14:49:13.793473", "status": "completed"} tags=[]
EMBEDDING_DIM=100
maxLenText=100
maxLenSummary=40
# + papermill={"duration": 0.422169, "end_time": "2021-03-22T14:49:14.373259", "exception": false, "start_time": "2021-03-22T14:49:13.951090", "status": "completed"} tags=[]
from keras.preprocessing.sequence import pad_sequences
def padding(encoder_sequences, decoder_sequences, maxLenText, maxLenSummary):
encoder_input_data = pad_sequences(encoder_sequences, maxlen=maxLenText, dtype='int32', padding='post', truncating='post')
decoder_input_data = pad_sequences(decoder_sequences, maxlen=maxLenSummary, dtype='int32', padding='post', truncating='post')
return encoder_input_data, decoder_input_data
encoder_input_data, decoder_input_data = padding(encoder_sequences, decoder_sequences,maxLenText, maxLenSummary)
# + papermill={"duration": 0.083092, "end_time": "2021-03-22T14:49:14.532028", "exception": false, "start_time": "2021-03-22T14:49:14.448936", "status": "completed"} tags=[]
encoder_input_data.shape
# + papermill={"duration": 0.08319, "end_time": "2021-03-22T14:49:14.691177", "exception": false, "start_time": "2021-03-22T14:49:14.607987", "status": "completed"} tags=[]
def glove_100d_dictionary():
embeddings_index = {}
f = open('../input/glove-global-vectors-for-word-representation/glove.6B.100d.txt')
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
f.close()
return embeddings_index
# + papermill={"duration": 16.814012, "end_time": "2021-03-22T14:49:31.581280", "exception": false, "start_time": "2021-03-22T14:49:14.767268", "status": "completed"} tags=[]
embeddings_index = glove_100d_dictionary()
# + papermill={"duration": 0.129413, "end_time": "2021-03-22T14:49:31.786633", "exception": false, "start_time": "2021-03-22T14:49:31.657220", "status": "completed"} tags=[]
def embedding_matrix_creater(embedding_dimention, tokenizer):
embedding_matrix = np.zeros((len(tokenizer.word_index) + 1, embedding_dimention))
for word, i in tokenizer.word_index.items():
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
# words not found in embedding index will be all-zeros.
embedding_matrix[i] = embedding_vector
return embedding_matrix
# + papermill={"duration": 0.230107, "end_time": "2021-03-22T14:49:32.164039", "exception": false, "start_time": "2021-03-22T14:49:31.933932", "status": "completed"} tags=[]
embeddingMatrixText = embedding_matrix_creater(100, tokenizerText)
embeddingMatrixSummary = embedding_matrix_creater(100, tokenizerSummary)
# + papermill={"duration": 0.084714, "end_time": "2021-03-22T14:49:32.372170", "exception": false, "start_time": "2021-03-22T14:49:32.287456", "status": "completed"} tags=[]
embeddingMatrixText.shape
# + papermill={"duration": 0.097338, "end_time": "2021-03-22T14:49:32.544895", "exception": false, "start_time": "2021-03-22T14:49:32.447557", "status": "completed"} tags=[]
from keras.layers import Embedding
encoder_embedding_layer = Embedding(input_dim = textVocabSize+1,
output_dim = 100,
input_length = maxLenText,
mask_zero=True,
weights = [embeddingMatrixText],
trainable = False)
decoder_embedding_layer = Embedding(input_dim = summaryVocabSize+1,
output_dim = 100,
input_length = maxLenSummary,
mask_zero=True,
weights = [embeddingMatrixSummary],
trainable = False)
# + papermill={"duration": 0.082622, "end_time": "2021-03-22T14:49:32.702802", "exception": false, "start_time": "2021-03-22T14:49:32.620180", "status": "completed"} tags=[]
# sampleReview.Text = sampleReview.Text.apply(lambda tokens: [contractions[word] for word in tokens if word.isalpha()])
# sampleReview.Summary = sampleReview.Summary.apply(lambda tokens: [contractions[word] for word in tokens if word.isalpha()])
# -
pip install pydot
# + papermill={"duration": 0.091559, "end_time": "2021-03-22T14:49:32.870113", "exception": false, "start_time": "2021-03-22T14:49:32.778554", "status": "completed"} tags=[]
from numpy.random import seed
seed(1)
from sklearn.model_selection import train_test_split
import logging
import matplotlib.pyplot as plt
import pandas as pd
import pydot
import keras
from keras import backend as k
k.set_learning_phase(1)
from keras.preprocessing.text import Tokenizer
from keras import initializers
from keras.optimizers import RMSprop,Adam
from keras.models import Sequential,Model
from keras.layers import Dense,LSTM,Dropout,Input,Activation,Add,concatenate, Embedding, RepeatVector
from keras.layers.advanced_activations import LeakyReLU,PReLU
from keras.callbacks import ModelCheckpoint
from keras.models import load_model
from keras.optimizers import Adam
from keras.layers import TimeDistributed
# + papermill={"duration": 0.084592, "end_time": "2021-03-22T14:49:33.032826", "exception": false, "start_time": "2021-03-22T14:49:32.948234", "status": "completed"} tags=[]
MAX_LEN = 100
EMBEDDING_DIM = 100
HIDDEN_UNITS = 300
textVocabSize = textVocabSize+1
summaryVocabSize = summaryVocabSize+1
LEARNING_RATE = 0.002
BATCH_SIZE = 8
EPOCHS = 5
# + papermill={"duration": 0.099054, "end_time": "2021-03-22T14:49:33.208900", "exception": false, "start_time": "2021-03-22T14:49:33.109846", "status": "completed"} tags=[]
# input_characters = sorted(list(input_characters))+[' ']
# target_characters = sorted(list(target_characters))+[' ']
# num_encoder_tokens = len(encoder_input_data)
# num_decoder_tokens = len(decoder_input_data)
num_encoder_tokens = textVocabSize
num_decoder_tokens = summaryVocabSize
max_encoder_seq_length = max([len(txt) for txt in encoder_input_data])
max_decoder_seq_length = max([len(txt) for txt in decoder_input_data])
print("Number of samples:", len(input_texts))
print("Number of unique input tokens:", num_encoder_tokens)
print("Number of unique output tokens:", num_decoder_tokens)
print("Max sequence length for inputs:", max_encoder_seq_length)
print("Max sequence length for outputs:", max_decoder_seq_length)
# + papermill={"duration": 0.084686, "end_time": "2021-03-22T14:49:33.370833", "exception": false, "start_time": "2021-03-22T14:49:33.286147", "status": "completed"} tags=[]
# encoder_input_data = np.zeros(
# (len(input_texts), max_encoder_seq_length, num_encoder_tokens), dtype="float32"
# )
# decoder_input_data = np.zeros(
# (len(input_texts), max_decoder_seq_length, num_decoder_tokens), dtype="float32"
# )
# decoder_target_data = np.zeros(
# (len(input_texts), max_decoder_seq_length, num_decoder_tokens), dtype="float32"
# )
# + papermill={"duration": 0.08335, "end_time": "2021-03-22T14:49:33.533206", "exception": false, "start_time": "2021-03-22T14:49:33.449856", "status": "completed"} tags=[]
# for i, seqs in enumerate(encoder_input_data):
# for j, seq in enumerate(seqs):
# decoder_target_data[i, j, seq] = 1.0
# + papermill={"duration": 0.124181, "end_time": "2021-03-22T14:49:33.734368", "exception": false, "start_time": "2021-03-22T14:49:33.610187", "status": "completed"} tags=[]
from tensorflow.keras.layers import Input, LSTM, Embedding, Dense, Concatenate, TimeDistributed, Bidirectional
# + papermill={"duration": 4.872511, "end_time": "2021-03-22T14:49:38.684338", "exception": false, "start_time": "2021-03-22T14:49:33.811827", "status": "completed"} tags=[]
"""
Chatbot Inspired Encoder-Decoder-seq2seq
"""
encoder_inputs = Input(shape=(maxLenText, ), dtype='int32',)
encoder_embedding = encoder_embedding_layer(encoder_inputs)
encoder_LSTM = LSTM(HIDDEN_UNITS, return_state=True,return_sequences=True)
encoder_outputs1, state_h, state_c = encoder_LSTM(encoder_embedding)
encoder_lstm2 = LSTM(HIDDEN_UNITS,return_sequences=True,return_state=True)
encoder_output2, state_h2, state_c2 = encoder_lstm2(encoder_outputs1)
encoder_lstm3= LSTM(HIDDEN_UNITS, return_state=True)
encoder_outputs, state_h, state_c= encoder_lstm3(encoder_output2)
decoder_inputs = Input(shape=(maxLenSummary, ), dtype='int32',)
decoder_embedding = decoder_embedding_layer(decoder_inputs)
decoder_LSTM = LSTM(HIDDEN_UNITS, return_state=True, return_sequences=True)
decoder_outputs, _, _ = decoder_LSTM(decoder_embedding, initial_state=[state_h, state_c])
# attn_out, attn_states = tf.keras.layers.Attention()([encoder_outputs, decoder_outputs])
# decoder_concat_input = Concatenate(axis=-1, name='concat_layer')([decoder_outputs, attn_out])
# dense_layer = Dense(VOCAB_SIZE, activation='softmax')
decoder_time = TimeDistributed(Dense(summaryVocabSize, activation='softmax'))
outputs = decoder_time(decoder_outputs)
model = Model([encoder_inputs, decoder_inputs], outputs)
# + papermill={"duration": 0.095617, "end_time": "2021-03-22T14:49:38.858033", "exception": false, "start_time": "2021-03-22T14:49:38.762416", "status": "completed"} tags=[]
rmsprop = RMSprop(lr=0.01, clipnorm=1.)
model.compile(loss='categorical_crossentropy', optimizer=rmsprop, metrics=["accuracy"])
# + papermill={"duration": 0.093811, "end_time": "2021-03-22T14:49:39.029984", "exception": false, "start_time": "2021-03-22T14:49:38.936173", "status": "completed"} tags=[]
model.summary()
# + papermill={"duration": 0.085337, "end_time": "2021-03-22T14:49:39.194182", "exception": false, "start_time": "2021-03-22T14:49:39.108845", "status": "completed"} tags=[]
import numpy as np1
num_samples = len(decoder_sequences)
decoder_output_data = np.zeros((num_samples, maxLenSummary, summaryVocabSize), dtype="int32")
# + papermill={"duration": 0.085843, "end_time": "2021-03-22T14:49:39.358547", "exception": false, "start_time": "2021-03-22T14:49:39.272704", "status": "completed"} tags=[]
num_samples
# + papermill={"duration": 0.410247, "end_time": "2021-03-22T14:49:39.848409", "exception": false, "start_time": "2021-03-22T14:49:39.438162", "status": "completed"} tags=[]
for i, seqs in enumerate(decoder_sequences):
for j, seq in enumerate(seqs):
if j > 0:
decoder_output_data[i][j-1][seq] = 1
# + papermill={"duration": 0.094456, "end_time": "2021-03-22T14:49:40.022998", "exception": false, "start_time": "2021-03-22T14:49:39.928542", "status": "completed"} tags=[]
art_train, art_test, sum_train, sum_test = train_test_split(encoder_input_data, decoder_input_data, test_size=0.2)
train_num = art_train.shape[0]
target_train = decoder_output_data[:train_num]
target_test = decoder_output_data[train_num:]
# + papermill={"duration": 0.0862, "end_time": "2021-03-22T14:49:40.188921", "exception": false, "start_time": "2021-03-22T14:49:40.102721", "status": "completed"} tags=[]
import tensorflow as tf
# + papermill={"duration": 0.088337, "end_time": "2021-03-22T14:49:40.355750", "exception": false, "start_time": "2021-03-22T14:49:40.267413", "status": "completed"} tags=[]
class My_Custom_Generator(keras.utils.Sequence) :
def __init__(self, art_train, sum_train, decoder_output, batch_size) :
self.art = art_train
self.sum = sum_train
self.decoder = decoder_output
self.batch_size = batch_size
def __len__(self) :
return (np.ceil(len(self.art) / float(self.batch_size))).astype(np.int)
def __getitem__(self, idx) :
batch_x1 = self.art[idx * self.batch_size : (idx+1) * self.batch_size]
batch_x2 = self.sum[idx * self.batch_size : (idx+1) * self.batch_size]
batch_y = self.decoder[idx * self.batch_size : (idx+1) * self.batch_size]
return [np.array(batch_x1),np.array(batch_x2)],np.array(batch_y)
# + papermill={"duration": 0.085056, "end_time": "2021-03-22T14:49:40.519366", "exception": false, "start_time": "2021-03-22T14:49:40.434310", "status": "completed"} tags=[]
batch_size = 64
my_training_batch_generator = My_Custom_Generator(art_train, sum_train, target_train, batch_size)
my_validation_batch_generator = My_Custom_Generator(art_test, sum_test, target_test,batch_size)
# + papermill={"duration": 1291.726571, "end_time": "2021-03-22T15:11:12.324735", "exception": false, "start_time": "2021-03-22T14:49:40.598164", "status": "completed"} tags=[]
callback = tf.keras.callbacks.EarlyStopping(monitor='accuracy', patience=4)
model.fit_generator(generator=my_training_batch_generator,
steps_per_epoch = int(16000 // batch_size),
epochs = 100,
verbose = 1,callbacks=[callback])
# + papermill={"duration": 2.773596, "end_time": "2021-03-22T15:11:18.234445", "exception": false, "start_time": "2021-03-22T15:11:15.460849", "status": "completed"} tags=[]
model.save_weights('nmt_weights_100epochs.h5')
# + papermill={"duration": 2.866686, "end_time": "2021-03-22T15:11:23.920505", "exception": false, "start_time": "2021-03-22T15:11:21.053819", "status": "completed"} tags=[]
model.load_weights('nmt_weights_100epochs.h5')
# + papermill={"duration": 3.35559, "end_time": "2021-03-22T15:11:29.941953", "exception": false, "start_time": "2021-03-22T15:11:26.586363", "status": "completed"} tags=[]
encoder_states = [state_h, state_c]
encoder_model = Model(encoder_inputs, encoder_states)
thought_input = [Input(shape=(HIDDEN_UNITS, )), Input(shape=(HIDDEN_UNITS, ))]
decoder_embedding = decoder_embedding_layer(decoder_inputs)
decoder_outputss, state_h, state_c = decoder_LSTM(decoder_embedding, initial_state=thought_input)
decoder_states = [state_h, state_c]
decoder_outputs = decoder_time(decoder_outputss)
decoder_model = Model(inputs=[decoder_inputs]+thought_input, outputs=[decoder_outputs]+decoder_states)
# + papermill={"duration": 2.73243, "end_time": "2021-03-22T15:11:35.614858", "exception": false, "start_time": "2021-03-22T15:11:32.882428", "status": "completed"} tags=[]
# encoder_states = [encoder_outputs, state_h, state_c]
# encoder_model = Model(encoder_inputs, encoder_states)
# decoder_embedding = decoder_embedding_layer(decoder_inputs)
# decoder_outputss, state_h, state_c = decoder_LSTM(decoder_embedding, initial_state=thought_input)
# decoder_states = [state_h, state_c]
# decoder_outputs = decoder_time(decoder_outputss)
# decoder_model = Model(inputs=[decoder_inputs]+thought_input, outputs=[decoder_outputs]+decoder_states)
# + papermill={"duration": 2.728603, "end_time": "2021-03-22T15:11:41.035415", "exception": false, "start_time": "2021-03-22T15:11:38.306812", "status": "completed"} tags=[]
decoder_model.summary()
# + papermill={"duration": 3.154252, "end_time": "2021-03-22T15:11:47.148840", "exception": false, "start_time": "2021-03-22T15:11:43.994588", "status": "completed"} tags=[]
def decode_sequence(input_seq):
states_value = encoder_model.predict(input_seq)
target_seq = np.zeros((100,1))
target_seq[0, 0] = word2idxSummary['bos']
stop_condition = False
decoded_sentence = ''
while not stop_condition:
output_tokens, h, c = decoder_model.predict([target_seq] + states_value)
sampled_token_index = np.argmax(output_tokens[0, -1, :])
sampled_word =idx2wordSummary[sampled_token_index]
decoded_sentence += ' '+ sampled_word
if (sampled_word == 'eos' or
len(decoded_sentence) > 40):
stop_condition = True
target_seq = np.zeros((100,1))
target_seq[0, 0] = sampled_token_index
states_value = [h, c]
return decoded_sentence
# + papermill={"duration": 6.739858, "end_time": "2021-03-22T15:11:56.632350", "exception": false, "start_time": "2021-03-22T15:11:49.892492", "status": "completed"} tags=[]
print("Text->",tokenizerText.sequences_to_texts([art_train[10]]))
print("\n\n\n")
print("Summary->",tokenizerSummary.sequences_to_texts([art_test[10]]))
print("\n\n\n")
print("using model->",decode_sequence([art_test[10]]))
# -
import pickle
filename = 'model.pkl'
pickle.dump(decode_sequence, open(filename, 'wb'))
model_columns = list(X.columns)
with open('columns.pkl','wb') as file:
pickle.dump(model_columns, file)
| modelNotebook/nlp-summarization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div style="text-align: center;">
# <h2>INFSCI 2595 Machine Learning - Spring 2019 </h2>
# <h1 style="font-size: 250%;">Assignment #1</h1>
# <h3>Due: Feb 17, 2019</h3>
# <h3>Total points: 100 </h3>
# </div>
# Type in your information in the double quotes
firstName = ""
lastName = ""
pittID = ""
#Libraries
# %matplotlib inline
import numpy as np
import pandas as pd
import seaborn as sns
import statsmodels.api as sm
import matplotlib.pyplot as plt
from IPython.display import Image
import statsmodels.formula.api as smf
from sklearn.datasets import load_boston
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn import datasets
# <h3> Problem #1. K-nearest neighbors [13 points] </h3>
#
# In this problem, do not use build-in functions for classification (i.e. do not use sklearn.neighbors). You will be required to do the calculations and make the predictions based on your understanding of how the technique works. You can use pen and paper to do the calculations (and upload image of your solutions) or write the code of detailed calculations and prediction.
#
# The table below provides a training data set containing six observations, three predictors, and one qualitative response variable.
# Suppose we wish to use this data set to make a prediction for Y when X1 = -1, X2 = 1 X3 = 2 using K-nearest neighbors.
# |Obs.|X1|X2|X3|Y
# |--|-------------------------------|
# |1 |0|3|0|Green|
# |2 |2|0|0|Green|
# |3|0|1|3|Green|
# |4|0|1|2|Red|
# |5|-1|0|1|Red|
# |6|1|1|1|Green|
# <b>Problem #1-1.</b> Compute the Euclidean distance between each observation and the test point. Your output should be a vector, where each number in the vector represents the distance between the observations and the test point [5 pts].
#
# Answer code
# here is all data
all_points = np.array([[0,3,0],[2,0,0],[0,1,3],[0,1,2],[-1,0,1],[1,1,1]])
test_point = np.array([[-1,1,2]])
data = np.concatenate((all_points, test_point), axis=0)
# here colomn names for data frame
names = np.array(['Obs.1','Obs.2','Obs.3','Obs.4','Obs.5','Obs.6','T_Point'])
# creating empty 2d array
array = [[0 for x in range(len(data))] for y in range(len(data))]
#### Calculating Euclidean distance
i = 0
while i < len(data)-1:
j = 1
while j < len(data):
dist = (((data[i][0]-data[j][0])**2)+ ((data[i][1]-data[j][1])**2)+((data[i][2]-data[j][2])**2))**(.5)
array[i][j] = dist
j= j+1
i = i+1
### Creating symemtical matrix
for i in range(len(data)):
for j in range(i, len(data)):
array[j][i] = array[i][j]
# printing
print(" Euclidean distance table:")
print (pd.DataFrame(array,columns=names, index=names) )
# <b>Problem #1-2 [4 points].</b> What is our prediction with K = 1?
#Answer code
from sklearn.neighbors import KNeighborsClassifier
Y = np.array([0,0,0,1,1,0]) # Where 0 represents Green color and 1 represents Red color.
all_points = np.array([[0,3,0],[2,0,0],[0,1,3],[0,1,2],[-1,0,1],[1,1,1]])
test_point = np.array([[-1,1,2]])
def printcolor(x,k):
print ("When k is equal to %d, prediction for Y when X1 = X2 = X3 = 0 is:" %k )
if x == [0]:
print ("Green")
elif x == [1]:
print ("Red")
else:
print ("Something went wrong sorry.....")
k = 1
knn=KNeighborsClassifier(n_neighbors= k)
knn.fit(all_points,Y)
predict = knn.predict(test_point)
print ("Using KNeighborsClassifier")
printcolor(predict,k)
print ("==========================================================================")
print ("Using Euclidean distance table")
df = pd.DataFrame(np.vstack((np.delete(pd.DataFrame(array,columns=names, index=names)["T_Point"].values, 6, 0),Y)).T,columns= ["Distance","Y" ] )
#predict = df["Distance"].min()
Small = df.nsmallest(1, 'Distance')
Y = Small['Y'].values
printcolor(Y,k)
# <b>Problem #1-3. [4 points]</b> What is our prediction with K = 3? K = 5?
# +
# Answer code
from sklearn.neighbors import KNeighborsClassifier
k = 3
Y = np.array([0,0,0,1,1,0])
knn=KNeighborsClassifier(n_neighbors= k)
knn.fit(all_points,Y)
predict = knn.predict(test_point)
print ("Using KNeighborsClassifier")
printcolor(predict,k)
print ("==========================================================================")
print ("Using Euclidean distance table")
from scipy import stats
Y= np.array(stats.mode(df.nsmallest(k, 'Distance')['Y'].values))[0]
printcolor(Y,k)
# +
from sklearn.neighbors import KNeighborsClassifier
k = 5
Y = np.array([0,0,0,1,1,0])
knn=KNeighborsClassifier(n_neighbors= k)
knn.fit(all_points,Y)
predict = knn.predict(test_point)
print ("Using KNeighborsClassifier")
printcolor(predict,k)
print ("==========================================================================")
print ("Using Euclidean distance table")
from scipy import stats
Y= np.array(stats.mode(df.nsmallest(k, 'Distance')['Y'].values))[0]
printcolor(Y,k)
# -
# <b>Problem #1-4 [4 points].</b> For KNN classification, discuss the impact of choosing the parameter K on the model performance. Highlight the different trade-offs.
# <h3>Answer: </h3> <br>
# **In the KNN model: **
# The value of K has a significant impact on prediction. Changing K may lead to different predictions. We try to avoid very low values of K as they may overfit the model, and very high values of K would underfit the model.
#
# <h3> Problem #2. Linear regression calculations of coefficients and $R^2$ [17 points] </h3>
# - You should not use built-in functions for linear regression in this question's parts (2-1) and (2-2). Do the calculations manually or explicitly by code. <br>
# - Feel free to select any type of submission you are comfortable with (Since it may require some mathematical formula and symbols; MS Word, or scanned version of your writing will be fine)
# <b>Problem #2-1.</b> Find the least squared fit of a linear regression model using the following traning data. Coefficients are obtained using the formulas [7pts]:
# \begin{array} \\
# \hat{\beta}_0 = \bar{y} - \hat{\beta}_1\bar{x}, \\ \\
# \hat{\beta}_1 = \frac{\sum_{i=1}^n y_i x_i - \bar{y}\sum_{i=1}^n x_i}{\sum_{i=1}^n x^2 - \bar{x}\sum_{i=1}^n x_i} \\ \\
# \text{where }\bar{x} = \frac{\sum_{i=1}^y x_i}{n} \text{ and } \bar{y} = \frac{\sum_{i=1}^y y_i}{n} \\
# \end{array}
# |Smoke index(x)|Lung cancer mortality index(y)|
# |:--:|:-------------------------------:|
# |127|115|
# |121|128|
# |94|128|
# |126|156|
# |102|101|
# |111|128|
# |95|115|
# |89|105|
# |101|87|
# +
Image(filename='image1.png')
# -
# <b>Problem #2-2.</b> Given the test data below, compute the TSS, RSS and the R-squared metric of the fitted model [6pts]
# |Xi|Yi|
# |:--:|:-------------------------------:|
# |90|103|
# |106|131|
# |105|85|
# |115|99|
# |113|144|
# +
Image(filename='image3.png')
# -
# <b>Problem #2-3.</b> Check your calculations by finding the coefficients and the $R^2$ score using scikit-learn library (from sklearn.linear_model import LinearRegression) [4pts]
# +
X_train=np.array([127,121,94,126,102,111,96,89,101])
Y_train=np.array([115,128,128,156,101,128,115,105,87])
#from sklearn.linear_model import LinearRegression
X_train=X_train.reshape(-1,1); X_train=X_train.reshape(-1,1)
linreg= LinearRegression().fit(X_train, Y_train)
print("The intercept using sklearn: ", linreg.intercept_)
print("The coefficient using sklearn:",linreg.coef_)
Image(filename='image1.png')
X_test=np.array([90,106,105,115,113])
Y_test=np.array([103,131,85,99,144])
X_test=X_test.reshape(-1,1); X_test=X_test.reshape(-1,1)
scoreR=linreg.score(X_test, Y_test)
print("The R sqaured score is using sklearn,", scoreR)
print("Answers are almost the same as the calculated one, minor differences due to numbers approaximations in coefficient values.")
# -
# <h3> Problem #3. Linear Regression Optimization with Gradient Descent [20 points] </h3>
#
# In this question, you will create a synthetic dataset, then use gradient descent to find the coefficients' estimates. We will see how the RSS changes with iterations of the gradient descent.
#
#
# <b>Problem #3-1.</b> Create a label $y$, which is related to a feature $X$, such that $y=10 + 5 X + \epsilon$. $\epsilon$ is a Gaussian noise (normal distribution) with mean 0 and unit variance. X is generated from a uniform distribution as given below. The number of observations to be generated is 100 [4pts].
# +
NumberObservations=100
minVal=1
maxVal=20
X = np.random.uniform(minVal,maxVal,(NumberObservations,1))
print(X.shape)
#Add you code below to define error and Y based on the information above
# +
beta0=10
beta1=5
stdDeviation=1
ErrMean=0
error = np.random.normal(loc=ErrMean, scale=stdDeviation,size=(NumberObservations,1))
y=beta0+beta1*X +error
# -
# <b>Problem #3-2 [12 Points].</b> Now assume you do not know the actual relationship between X and y, use the generated noisy data and find the coefficients of a linear regression model obtained by gradient descent. Do not use any built-in functions for gradient descent. Write the code based on your understanding of the algorithm. The cost function is the RSS, set the number of iterations to 10000 and the learning rate to 0.00001.
#
# - Print the estimated coefficients of the model obtained using gradient descent (7 pts)
# - Record the RSS in each iteration of the gradient descent and plot the last 6000 values of the RSS. Comment on figure. (3 pts)
# - Comment on how close the obtained coefficients are from the actual ones (3pts)
# +
# write your code here.
#Make sure that x has two indices
n_iter=10000
eta=0.00001 #learning rate =10^-4 worked well
RSS=[]
w=np.random.normal(loc=0, scale=stdDeviation,size=(1 + X.shape[1],1))
# random initialization, with size equals to size of x + 1 (since we also have bias)
print(w.shape)
for i in range(n_iter):
output = np.dot(X, w[1:]) + w[0]
output=output.reshape(-1,1)
errors = (output-y)
w[1:] =w[1:]- eta * 2*np.dot(X.T,errors)
w[0] = w[0] - eta * 2*errors.sum()
RSS1 = (errors**2).sum()
RSS.append(RSS1)
print(w)
plt.plot(RSS[len(RSS)-6000:])
# -
# <b>Problem #3-1 [5 Points].</b> Repeat (3-1) and (3-2) but increase the standard deviation of the error to 100. Comment on output coefficient estimates. (3pts)
# +
beta0=10
beta1=5
stdDeviation=100
ErrMean=0
error = np.random.normal(loc=ErrMean, scale=stdDeviation,size=(NumberObservations,1))
y=beta0+beta1*X +error
n_iter=10000
eta=0.00001 #learning rate =10^-4 worked well
RSS=[]
w=np.random.normal(loc=0, scale=stdDeviation,size=(1 + X.shape[1],1))
# random initialization, with size equals to size of x + 1 (since we also have bias)
print(w.shape)
for i in range(n_iter):
output = np.dot(X, w[1:]) + w[0]
output=output.reshape(-1,1)
errors = (output-y)
w[1:] =w[1:]- eta * 2*np.dot(X.T,errors)
w[0] = w[0] - eta * 2*errors.sum()
RSS1 = (errors**2).sum()
RSS.append(RSS1)
print(w)
plt.plot(RSS[len(RSS)-6000:])
# -
# Due to the increased variance of the error the coefficent estimate becomes far from the actual ones.
# ======================================================================================================================= <br>=======================================================================================================================
# <h3> Problem #4. This question involves the use of multiple linear regression on the Boston dataset [30 points]</h3> <br>
# In this part, you should download and analyze **"Boston House Prices"** dataset. <br>
# - Whole dataset contains 14 attributes, (13 numeric/categorical predictive and 1 target value) what is a abbreviation of the target value?
# - The target value is MEDV => Median value of owner-occupied homes in $1000's
#
# Use a code below to download the dataset:
dataset = load_boston()
print(dataset.keys())
# Print and read the description of the dataset then answer the following questions.
#
#
#Code
print(dataset.DESCR)
# =======================================================================================================================
# <b>Problem #4-1 </b> Generate descriptive statistics using DataFrame. (hint: use "DataFrame . describe" method)<br>
#
# Follow two steps to answer questions [4pts].
# - Create a DataFrame usnig "data" from the dataset with columns using "feature_names".
# - Generate descriptive statistics
#
# <b> Answer the following questions:</b>
#
# - Which feature has the lowest range (minimum and maximum value)?
#
# - Which feature has the higest mean?
#
# - Which feature has a highest variance?
#Code
DataFrame = pd.DataFrame(dataset.data, columns= dataset.feature_names)
DataFrame.describe()
list_range = []
for i in range(len(DataFrame.columns)):
list_range.append(max(DataFrame.iloc[:,i])-min(DataFrame.iloc[:,i]))
indexes = [k for k,x in enumerate(list_range) if x == min(list_range)]
indexes
list_var = []
for i in range(len(DataFrame.columns)):
list_var.append(np.var(DataFrame.iloc[:,i]))
indexes = [k for k,x in enumerate(list_var) if x == max(list_var)]
indexes
# <h3>Answer: </h3> <br>
# <b>
# Which feature has the lowest range (minimum and maximum value)?
#
# - Feature *NOX* has the lowest range
#
# Which feature has the higest mean?
#
# - Feature *TAX* has the higest mean
#
# Which feature has a lowest standard deviation?
#
# - Feature *TAX* has a highest variance
#
# </b>
# =======================================================================================================================
# <b>Problem #4-2</b> Feature Scaling
#
# <b> Answer the following questions [4pts]:</b>
#
# - From the information above, Do you recommend **feature scaling** ? Explain.
#
# - What is a difference between MinMaxScaler and StandardScaler?
# <h3>Answer: </h3> <br>
# <b>
#
# - It is recommended to try feature scaling, especially if we will try non-parametric models, since the different features have different scale. This is also essential if we use gradient descent. For OLS, scaling features is not expected to matter since coefficents will take care of this issue.
#
# - MinMaxScaler scales a feature to the range of 0 to 1.
#
# - StandardScaler standardizes the features with zero mean and unit variance.
# </b>
# =======================================================================================================================
# <b>Problem #4-3</b> Calculate and report **correlations** between variables
#
# Follow the following steps to answer questions.
#
# - Add target to the dataFrame.
#
# - Find the correlation matrix that shows the correlation between each pair of variables in the dataframe. <br>
#
# - Plot a correlation matrix (heatmap) You can check: https://seaborn.pydata.org/generated/seaborn.heatmap.html
#
#
#
#
# <b> Answer the following questions [4pts]:</b>
#
# - What is a difference between positive and negative numbers on the correlation table? Explain.
# - What is the feature that is most correlated with the target? Do you think it is the most or the least helpful features in predicting the target class? Explain.
#
# - What is the correlation between the feature RM and the LSTAT?
# - What does this value of correlation indicate?
df = (pd.DataFrame(dataset.data, columns= dataset.feature_names))
df['target'] = dataset.target
corr = df.corr()
print (corr)
# <h3>Answer: </h3> <br>
# <b>
# - A negative number means that features are negatively correlated; if one variable increases, the other variable decreases. A positive number means positive correlation; If one variable increases second variable increase as well.
#
# - The feature that is most correlated with the target is LSTAT. They are negatively correlated.
# Features that have strong correlation with the target (positive or negative) are expected to be helpful in prediction. We need check the how strong the association is by obtaining the p-values.
#
# - The correlation between the feature RM and the LSTAT is -0.613808
#
# - This value indicates a high negative correlation between the feature RM and the LSTAT.
# </b>
# =======================================================================================================================
# for generating heatmap, you can use the code below or write your own
# %matplotlib notebook
dataset = load_boston()
sns.set(style="white")
# Get correlation matrix
df = (pd.DataFrame(dataset.data, columns= dataset.feature_names))
df['target'] = dataset.target
corr = df.corr()
# Generate a mask for the upper triangle
mask = np.zeros_like(corr, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(11, 9))
# Generate a custom diverging colormap
cmap = sns.diverging_palette(220, 10, as_cmap=True)
# Draw the heatmap with the mask and correct aspect ratio
sns.heatmap(corr, mask=mask, cmap=cmap, vmax=1, center=0, square=True, linewidths=.9, cbar_kws={"shrink": .7})
# =======================================================================================================================
#
#
# <b>Problem #4-4 </b> Scatter plot [4 points]. <br>
#
# - Plot RM versus target variable
# - Plot TAX versus target variable
# - Plot LSTAT versus target variable <br>
# - Comment on each of the above plots. Is the relationship linear or non linear?
#
# **Plot RM versus target variable**
# +
Y = dataset.target
X = dataset.data[:,[5]]
plt.scatter(X,Y,marker='o')
plt.xlabel('Feature value (X, RM)'); plt.ylabel('Target value (Y)')
plt.show()
# -
# **Plot TAX versus target variable**
# +
Y = dataset.target
X = dataset.data[:,[9]]
plt.scatter (X,Y,marker='o')
plt.xlabel('Feature value (X, TAX)'); plt.ylabel('Target value (Y)')
plt.show()
# -
# **Plot LSTAT versus target variable **
# +
Y = dataset.target
X = dataset.data[:,[12]]
plt.scatter(X,Y,marker='o')
plt.xlabel('Feature value (X, LSTAT)'); plt.ylabel('Target value (Y)')
plt.show()
# -
# <h3>Answer: </h3> <br>
# <b>
# - Plot RM versus target variable (approximately linear)
# - Plot TAX versus target variable (nonlinear)
# - Plot LSTAT versus target variable (approximately quadratic)
#
# =======================================================================================================================
# <b>Problem #4-5.</b>
#
# - Feel free to use scikit-learn (sklearn library) or write your own code to answer the following questions..
# - Use train_test_split() with "random_state=0
#
# Answer the following questions [10 pts]:
#
# 1. Fit a linear regression model with RM and LSTAT features only. Find the R-squared metric on train and test sets.
# 2. Fit a linear regression model using RM, LSTAT and include the interaction term (RM * LSTAT). How R-squared metric differs from the previous model without interaction term?
# 3. Fit a linear regression model using LSTAT and include the polynomial term ( $LSTAT^2$). Find the R-squared metric.
# 4. Fit linear regression model using LSTAT and include the polynomial term ( $LSTAT^2$ and $LSTAT^4$ ). Find the R-squared metric.
# - How does R-squared metric differ in the previous models ? <br> Comment your observation.
# **1. Fit a linear regression model with RM and LSTAT features only**
# +
# Data preprocessing
X_train,X_test,Y_train,Y_test=train_test_split(dataset.data,dataset.target,random_state=0)
pd_feature = pd.DataFrame(X_train, columns=dataset.feature_names)
pd_target = pd.DataFrame(Y_train, columns=['target'])
# Concatenate target and feature dataset
frames = [pd_feature, pd_target]
pd_dataset = pd.concat(frames, axis=1)
# using Statsmodels
# modelsmf = smf.ols('target ~ RM+LSTAT', pd_dataset)
# #modelsmf.fit().summary() #optional if you wanna see whole summary
# print ("Statsmodels model result of R-squared metric with RM and LSTAT features only:\n",modelsmf.fit().rsquared)
#=========================================================================================================================
# using Sklearn model
df = (pd.DataFrame(dataset.data, columns= dataset.feature_names))
df['target'] = dataset.target
Y = df['target']
X = df[["RM", "LSTAT"]]
X_train,X_test,Y_train,Y_test=train_test_split(X,Y,random_state=0)
# Here we are creting dataset using only train data.
frames = [X_train, Y_train]
pd_dataset = pd.concat(frames, axis=1)
modelLR = LinearRegression().fit(X_train,Y_train)
print("Sklearn model result of train R-squared metric with RM and LSTAT features only:\n", modelLR.score(X_train, Y_train))
print ("Use Test: Sklearn model result of R-squared metric with RM and LSTAT features only:\n", modelLR.score(X_test, Y_test))
# -
# **2 Fit a linear regression model using RM, LSTAT and include the interaction term (RM * LSTAT)**
# +
# using Statsmodels
# modelsmf = smf.ols('target ~ RM+LSTAT+LSTAT*RM', pd_dataset)
# # modelsmf.fit().summary() #optional if you wanna see whole summary
# print ("Statsmodels model result of R-squared metric with RM LSTAT and term (RM * LSTAT):\n",modelsmf.fit().rsquared)
#=========================================================================================================================
# using Sklearn model
# Data preprocessing
df = (pd.DataFrame(dataset.data, columns= dataset.feature_names))
df['target'] = dataset.target
Y = df['target']
df['RMLSTAT'] = (df['RM']*df['LSTAT'])
X = df[['RM', 'LSTAT','RMLSTAT']]
X_train,X_test,Y_train,Y_test=train_test_split(X,Y,random_state=0)
modelLR2 = LinearRegression().fit(X_train,Y_train)
print("Sklearn model result of train R-squared metric with RM and LSTAT term (RM * LSTAT):\n", modelLR2.score(X_train, Y_train))
print("Use Test: Sklearn model result of R-squared metric with RM and LSTAT features only:\n", modelLR2.score(X_test, Y_test))
# -
# **3 Fit a linear regression model using LSTAT and include the polynomial term ( LSTAT^2 )**
# using Statsmodels
# modelsmf = smf.ols('target ~ LSTAT+I(LSTAT**2)', pd_dataset)
# # modelsmf.fit().summary() #optional if you wanna see whole summary
# print ("Statsmodels model result of R-squared metric with LSTAT and polynomial term ( LSTAT^2):\n",modelsmf.fit().rsquared)
#=========================================================================================================================
# using Sklearn model
# Data preprocessing
df = (pd.DataFrame(dataset.data, columns= dataset.feature_names))
df['target'] = dataset.target
Y = df['target']
df['LSTATSQ']=(df['LSTAT']*df['LSTAT'])
X = df[['LSTAT','LSTATSQ']]
X_train,X_test,Y_train,Y_test=train_test_split(X,Y,random_state=0)
modelLRSQ = LinearRegression().fit(X_train,Y_train)
print("Sklearn model result of R-squared metric with LSTAT and polynomial term ( LSTAT^2):\n", modelLRSQ.score(X_train, Y_train))
print("Use Test: Sklearn model result of R-squared metric with RM and LSTAT features only:\n", modelLRSQ.score(X_test, Y_test))
# **4. Fit linear regression model using LSTAT and include the polynomial term ( LSTAT^2 and LSTAT^4)**
# using Statsmodels
# modelsmf = smf.ols('target ~ LSTAT+I(LSTAT**2)+I(LSTAT**4)', pd_dataset)
# # modelsmf.fit().summary() #optional if you wanna see whole summary
# print ("Statsmodels model result of R-squared metric with LSTAT and polynomial term (LSTAT^2 and LSTAT^4):\n",modelsmf.fit().rsquared)
#=========================================================================================================================
# using Sklearn model
# Data preprocessing
df = (pd.DataFrame(dataset.data, columns= dataset.feature_names))
df['target'] = dataset.target
Y = df['target']
df['LSTATSQ']=(df['LSTAT']*df['LSTAT'])
df['LSTAT4']=(df['LSTAT']*df['LSTAT']*df['LSTAT']*df['LSTAT'])
X = df[['LSTAT','LSTATSQ','LSTAT4']]
X_train,X_test,Y_train,Y_test=train_test_split(X,Y,random_state=0)
modelLRSQ = LinearRegression().fit(X_train,Y_train)
print("Sklearn model result of R-squared metric with LSTAT and polynomial term (LSTAT^2 and LSTAT^4):\n", modelLRSQ.score(X_train, Y_train))
print("Use Test: Sklearn model result of R-squared metric with RM and LSTAT features only:\n", modelLRSQ.score(X_test, Y_test))
# =========================================================================================================================
# <b>Problem #4-6 .</b> Fit all features (13 features) in the dataset to a multiple linear regression model, and report p-values of all feature.
#
# <b> Answer the following questions [4 pts]:</b>
#
# - What does p-value represent?
# - What are the features that have strong association with the target? what are the ones that seem to have week association? <br>
#
# Data preprocessing
pd_feature = pd.DataFrame(dataset.data, columns=dataset.feature_names)
pd_target = pd.DataFrame(dataset.target, columns=['target'])
# Concatenate target and feature dataset
frames = [pd_feature, pd_target]
pd_dataset = pd.concat(frames, axis=1)
full_model = smf.ols('target ~ CRIM+ZN+INDUS+CHAS+NOX+RM+AGE+DIS+RAD+TAX+PTRATIO+B+LSTAT', pd_dataset)
full_model_result = full_model.fit()
print(full_model_result.summary())
# <h3>Answer: </h3> <br>
# <b>
# - A small p-value (typically ≤ 0.05) indicates strong evidence against the null hypothesis, hence strong association with the target.
# - Most of the features look important (with low p-values). There are two insignificant features: INDUS and AGE.
# <h3> Problem #5. Regularization [20 points] </h3>
# - Will be covered in lecture on Feb 11.
#
#
# - In this problem, we will use the same dataset as in previous question -- the Boston data set.
# - Scale the features with StandardScaler
# <b>Problem #5-1 [5 points].</b> Use Ridge regression model with tuning parameter set to 1 (alpha =1). Find the test $R^2$ score and number of non zero coefficients.
#
# <b>Problem #5-2 [5 points].</b> Use Lasso regression instead of Ridge regression, also set the tuning parameter to 1. Find the test $R^2$ score and number of non zero coefficients.
#
# <b>Problem #5-3 [5 points].</b> Change the tuning parameter of the Lasso model to a very low value (alpha =0.001). What is the $R^2$ score.
#
# <b>Problem #4-4 [5 points].</b> Comment on your result.
#
# +
#try with scaling
from sklearn.datasets import load_boston
from sklearn.linear_model import Ridge
from sklearn.linear_model import Lasso
from sklearn import preprocessing
dataset = load_boston()
X=dataset.data
Y=dataset.target
X_train, X_test, Y_train, Y_test= train_test_split(X, Y, random_state= 0)
scaler=preprocessing.StandardScaler().fit(X_train)
X_train_transformed=scaler.transform(X_train)
X_test_transformed=scaler.transform(X_test)
#A) Ridge regression, using tuning parameter of 100
RidgeModel100=Ridge(alpha=1).fit(X_train_transformed, Y_train)
#find the R2 metric with the .score
print("Score of Ridge Regression with tuning parameter =1 is: ", RidgeModel100.score(X_test_transformed,Y_test))
print("number of coef. that are not equal to zero with Ridge regression", np.sum(RidgeModel100.coef_!=0))
#B) Lasso regression, using tuning parameter of 100
LassoModel100=Lasso(alpha=1).fit(X_train_transformed, Y_train)
print("Score of Lasso Regression with tuning parameter =1 is: ", LassoModel100.score(X_test_transformed,Y_test))
print("number of coef. that are not equal to zero with Lasso regression when alpha =100 is: ", np.sum(LassoModel100.coef_!=0))
print(LassoModel100.coef_)
#C) Lasso regression, using very small tuning parameter
LassoModel001=Lasso(alpha=0.001).fit(X_train_transformed, Y_train)
print("Score of Lasso Regression with tuning parameter =0.001 is: ", LassoModel001.score(X_test_transformed,Y_test))
print("number of coef. that are not equal to zero with Lasso regression when alpha =0.001 is: ", np.sum(LassoModel001.coef_!=0))
print(LassoModel001.coef_)
# -
# ##### Comment
#
# - It is clear from the results above that with Ridge regression, non of the coefficients is zero. Using Lasso regression with the same value of the tuning paramter, 8 coefficients are equal to zero.
#
# - With a low value for the tuning paramter, non of the coefficients are equal to zero with Lasso. It is expected that this result will be simiar to OLS with no regularization (you can check that in a straightforward manner).
#
# - Results implies that most features are important in predicting the response.
#
# ==========================================================================+============================================== <br> =========================================================================================================================
# ### Submission
# Once you completed the assignment, <br>
# Name your file in the format of <b style='color:red'>LASTNAME-PITTID-Assignment1.ipynb</b>, and submit it on the courseweb
| Assignment1Spring_2019-Soln.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from pandas.plotting import scatter_matrix
def missing_values_table(df):
mis_val = df.isnull().sum()
mis_val_percent = 100 * df.isnull().sum()/len(df)
mis_val_table = pd.concat([mis_val, mis_val_percent], axis=1)
mis_val_table_ren_columns = mis_val_table.rename(
columns = {0 : 'Valores ausentes', 1 : '% do valor total'})
return mis_val_table_ren_columns.loc[(mis_val_table_ren_columns!=0).any(axis=1)]
#load data
df = pd.read_csv("data/train.csv")
# ## Scatter plot geral - validar se os dados se apresentam de forma esparsa
df
# ## Visualização
from mpl_toolkits.mplot3d import Axes3D
from sklearn import decomposition
import seaborn as sn
df2 = df[df.columns.difference(['ID_code','target'])].copy()
def pca_view(df2, n_components=4):
pca = decomposition.PCA(n_components=n_components)
pca.fit(df2)
X = pca.transform(df2)
print("Soma dos 3 primeiros componentes:",np.sum(pca.explained_variance_ratio_[0:3]))
sn.barplot(list(range(1,len(pca.components_)+1)), 1*pca.explained_variance_ratio_)
plt.plot(np.cumsum(pca.explained_variance_ratio_), )
plt.show()
fig = plt.figure(1, figsize=(10, 10))
plt.clf()
ax = Axes3D(fig, rect=[0, 0, .95, 1], elev=18, azim=134)
plt.cla()
ax.scatter(X[:, 0], X[:, 1], X[:, 2], edgecolor='k')
ax.w_xaxis.set_ticklabels([])
ax.w_yaxis.set_ticklabels([])
ax.w_zaxis.set_ticklabels([])
fig = plt.figure(figsize = (8,8))
ax = fig.add_subplot(1,1,1)
ax.set_xlabel('Componente Principal 1', fontsize = 15)
ax.set_ylabel('Componente Principal 2', fontsize = 15)
ax.scatter(X[:, 0], X[:, 1], s = 50)
plt.show()
return X
df2.boxplot(figsize=(20, 20), rot=90)
# #### Em alguns atributos, é possível dectectar a presença de outiliers, nesse caso é necessário um pouco de atenção e verificar a necessidade de normalizar os dados.
from sklearn.preprocessing import StandardScaler, RobustScaler, Normalizer, QuantileTransformer, MinMaxScaler
x_st = Normalizer().fit_transform(df2)
print("Normalizados:")
pca_view(x_st, 10) # normalizado
#print("Dados originais:")
#pca_view(df2, 10) # dados reais
classes = df[['ID_code','target']].groupby(['target']).count()
classes
classes.plot.pie(y='ID_code', autopct='%1.0f%%')
from imblearn.over_sampling import SMOTE, ADASYN
sm = SMOTE(random_state=42)
ada = ADASYN(random_state=42)
# ### Aparentemente a disposição dos dados está mais para não linear do que uma apresentação linear, conforme visto anteriormente na técnica PCA, vou confirmar isso também utilizando um SVM sem kernel.
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
from sklearn.svm import LinearSVC, SVC
from sklearn.neural_network import MLPClassifier
from sklearn.model_selection import train_test_split, cross_val_score, cross_validate
import sklearn
import xgboost
from sklearn.model_selection import GridSearchCV
df3 = df.copy()
df3 = df3.sample(frac=1, replace=True, random_state=1)
X = df3[df3.columns.difference(['ID_code','target'])].copy()
y = df3['target'].values
X_res, y_res = sm.fit_resample(X, y)
X_train, X_test, y_train, y_test = train_test_split(X_res, y_res, train_size=0.75, test_size=0.25)
# +
#X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=0.75, test_size=0.25)
# +
# svm = SVC()
# svm.fit(X_train, y_train)
# print(svm_lin.score(X_test, y_test))
#print("R2",cross_val_score(svm_lin, X_test, y_test, cv=5, scoring='r2'))
# +
#print(sklearn.metrics.classification_report(y_test, svm.predict(X_test)))
# -
# ## RandomForest
rf = RandomForestClassifier()
rf.fit(X_train, y_train)
print(rf.score(X_test, y_test))
#print("R2",cross_validate(rf, X, y, cv=5))
print(sklearn.metrics.classification_report(y_test, rf.predict(X_test)))
# ## GradientBoosting
gb = GradientBoostingClassifier()
gb.fit(X_train, y_train)
print(gb.score(X_test, y_test))
#print("R2",cross_validate(gb, X, y, cv=5, scoring=('r2')))
# ## XGBoost
xgb = xgboost.XGBClassifier()
xgb.fit(X_train, y_train)
print(xgb.score(X_test, y_test))
#print("R2",cross_validate(xgb, X, y, cv=5, scoring=('r2')))
print(xgb.score(X_test, y_test))
print(sklearn.metrics.classification_report(y_test, xgb.predict(X_test)))
from imblearn.ensemble import RUSBoostClassifier, BalancedRandomForestClassifier
clf = BalancedRandomForestClassifier(random_state=0)
clf.fit(X_train, y_train)
print(sklearn.metrics.classification_report(y_test, clf.predict(X_test)))
# ## MLP
mlp = MLPClassifier(hidden_layer_sizes=(100,), alpha=0.1)
mlp.fit(X_train, y_train)
print(mlp.score(X_test, y_test))
print(sklearn.metrics.classification_report(y_test, mlp.predict(X_test)))
parameters = {'solver': ['lbfgs'], 'max_iter': [1000,1100,1200,1300,1400,1500,1600,1700,1800,1900,2000 ], 'alpha': 10.0 ** -np.arange(1, 10), 'hidden_layer_sizes':np.arange(10, 15), 'random_state':[0,1,2,3,4,5,6,7,8,9]}
clf = GridSearchCV(MLPClassifier(), parameters, n_jobs=-1, verbose=10)
clf.fit(X_train, y_train)
print(clf.score(X_test, y_test))
print(clf.best_params_)
print(sklearn.metrics.classification_report(y_test, clf.predict(X_test)))
mlp = MLPClassifier(hidden_layer_sizes=12, alpha=0.1, max_iter=1000, random_state=4, solver='lbfgs')
mlp.fit(X_train, y_train)
print(mlp.score(X_test, y_test))
print(sklearn.metrics.classification_report(y_test, mlp.predict(X_test)))
print(sklearn.metrics.classification_report(y_test, mlp.predict(X_test)))
from ultimate.mlp import MLP
param = {
'loss_type': 'mse',
'layer_size': [X_train.shape[1],16,16,16,1],
'activation': 'relu',
'output_range': [0, 1],
'output_shrink': 0.001,
'importance_mul': 0.0001,
'importance_out': True,
'rate_init': 0.02,
'rate_decay': 0.9,
'epoch_train': 50,
'epoch_decay': 1,
'verbose': 1,
}
param = (layer_size=[X_train.shape[1], 8, 8, 8, 1], rate_init=0.02, loss_type="mse", epoch_train=100, epoch_decay=10, verbose=1)
mlp2 = MLP(param).fit(X_train, y_train)
# #### Até agora, as técnicas XGBoost, GB, RF e SVM linear estão apresentando uma melhor performance
# A próxima abordagem é ajustar os hiper parâmetros, geralmente o comum é utilizar um técnica de GridSearch.
from sklearn.model_selection import RandomizedSearchCV, GridSearchCV
params = {
'min_child_weight': [1, 5, 10, 20, 30],
'gamma': [0.5, 1, 1.5, 2, 5, 10, 20, 30, 40, 50, 90, 100, 150],
'subsample': [0.6, 0.8, 1.0],
'colsample_bytree': [0.6, 0.8, 1.0],
'max_depth': [3, 4, 5, 10, 15, 50]
}
param_comb = 5
params = {
'min_child_weight': [1, 5, 10],
'gamma': [0.5, 1, 1.5, 2, 5],
'subsample': [0.6, 0.8, 1.0],
'colsample_bytree': [0.6, 0.8, 1.0],
'max_depth': [3, 4, 5]
}
random_search = RandomizedSearchCV(xgb, param_distributions=params, n_iter=param_comb, scoring='r2', n_jobs=4, cv=5, random_state=1001 )
random_search.fit(X_train, y_train)
print('\n All results:')
print(random_search.cv_results_)
print('\n Best estimator:')
print(random_search.best_estimator_)
print('\n Best normalized gini score for %d-fold search with %d parameter combinations:' % (5, param_comb))
print(random_search.best_score_ * 2 - 1)
print('\n Best hyperparameters:')
print(random_search.best_params_)
results_hp = cross_validate(random_search.best_estimator_, X, y, cv=5, scoring=('r2'))
results_nm = cross_validate(xgb, X, y, cv=5, scoring=('r2'))
print("XGB BH:", np.mean(results_hp['test_score']))
print("XGB NM:", np.mean(results_nm['test_score']))
fig, ax = plt.subplots(figsize=(20, 20))
xgboost.plot_importance(xgb, ax=ax)
fig, ax = plt.subplots(figsize=(20, 20))
xgboost.plot_importance(xgb, ax=ax)
test_df = pd.read_csv('data/test.csv')
test_df = pre_process(test_df)
test_df2 = test_df[test_df.columns.difference(['ID_code','target'])].copy()
pca_view(test_df2)
y_test = mlp.predict(test_df2)
results_df = pd.DataFrame(data={'ID_code':test_df['ID_code'], 'target':y_test})
results_df.to_csv('submission-mlp-05.csv', index=False)
| kaggle_santander/DataAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python3
# ---
# +
from devito import *
from examples.seismic.source import WaveletSource, TimeAxis
from examples.seismic import plot_image
import numpy as np
from sympy import init_printing, latex
init_printing(use_latex=True)
# -
# Initial grid: 1km x 1km, with spacing 100m
extent = (2000., 2000.)
shape = (81, 81)
x = SpaceDimension(name='x', spacing=Constant(name='h_x', value=extent[0]/(shape[0]-1)))
z = SpaceDimension(name='z', spacing=Constant(name='h_z', value=extent[1]/(shape[1]-1)))
grid = Grid(extent=extent, shape=shape, dimensions=(x, z))
# +
class DGaussSource(WaveletSource):
def wavelet(self, f0, t):
a = 0.004
return -2.*a*(t - 1./f0) * np.exp(-a * (t - 1./f0)**2)
# Timestep size from Eq. 7 with V_p=6000. and dx=100
t0, tn = 0., 200.
dt = 1e2*(1. / np.sqrt(2.)) / 60.
time_range = TimeAxis(start=t0, stop=tn, step=dt)
src = DGaussSource(name='src', grid=grid, f0=0.01, time_range=time_range)
src.coordinates.data[:] = [1000., 1000.]
# +
#NBVAL_SKIP
src.show()
# -
# Now we create the velocity and pressure fields
p = TimeFunction(name='p', grid=grid, staggered=NODE, space_order=2, time_order=1)
vx= TimeFunction(name='vx', grid=grid, staggered=x, space_order=2, time_order=1)
vz = TimeFunction(name='vz', grid=grid, staggered=z, space_order=2, time_order=1)
# +
t = grid.stepping_dim
time = grid.time_dim
# We need some initial conditions
V_p = 4.0
#V_s = 1.0
density = 3.
dx = 100.
rox = 1/density * dt
roz = 1/density * dt
l2m = V_p*V_p*density * dt
c1 = 9.0/8.0;
c2 = -1.0/24.0;
# The source injection term
src_p = src.inject(field=p.forward, expr=src)
# 2nd order acoustic according to fdelmoc
u_vx_2 = Eq(vx.forward, vx + rox * p.dx)
u_vz_2 = Eq(vz.forward, vz + roz * p.dz)
u_p_2 = Eq(p.forward, p + l2m * (vx.forward.dx + vz.forward.dz))
op_2 = Operator([u_vx_2, u_vz_2, u_p_2] + src_p)
# 4th order acoustic according to fdelmoc
# Now we create the velocity and pressure fields
p4 = TimeFunction(name='p', grid=grid, staggered=NODE,space_order=4, time_order=1)
vx4= TimeFunction(name='vx', grid=grid, staggered=x, space_order=4, time_order=1)
vz4 = TimeFunction(name='vz', grid=grid, staggered=z, space_order=4, time_order=1)
u_vx_2 = Eq(vx4.forward, vx4 + rox * p4.dx)
u_vz_2 = Eq(vz4.forward, vz4 + roz * p4.dz)
u_p_2 = Eq(p4.forward, p4 + l2m * (vx4.forward.dx + vz4.forward.dz))
op_4 = Operator([u_vx_2, u_vz_2, u_p_2] + src_p)
# +
### 2nd order acoustic
# Reset the fields
vx.data[:] = 0.
vz.data[:] = 0.
p.data[:] = 0.
# +
#NBVAL_IGNORE_OUTPUT
# Propagate the source
op_2(time=src.time_range.num-1)
# +
#NBVAL_SKIP
# Let's see what we got....
plot_image(vx.data[0])
plot_image(vz.data[0])
plot_image(p.data[0])
# +
#NBVAL_IGNORE_OUTPUT
### 4th order acoustic
# Propagate the source
op_4(time=src.time_range.num-1)
# +
#NBVAL_SKIP
# Let's see what we got....
plot_image(vx4.data[0])
plot_image(vz4.data[0])
plot_image(p4.data[0])
| examples/seismic/tutorials/05_staggered_acoustic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=false editable=false
# Initialize OK
from client.api.notebook import Notebook
ok = Notebook('lab09.ok')
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_context("talk")
# %matplotlib inline
import plotly.offline as py
py.init_notebook_mode(connected=True)
import plotly.graph_objs as go
import plotly.figure_factory as ff
import cufflinks as cf
cf.set_config_file(offline=False, world_readable=True, theme='ggplot')
# -
# # Logistic Regression
#
# In this lab we will be covering a very popular classification technique known as **logistic regression**.
#
# # Real Data
#
# For this lecture we will use the Wisconsin Breast Cancer Dataset which we can obtain from [scikit learn](http://scikit-learn.org/stable/datasets/index.html#breast-cancer-wisconsin-diagnostic-database).
import sklearn.datasets
data_dict = sklearn.datasets.load_breast_cancer()
data = pd.DataFrame(data_dict['data'], columns=data_dict['feature_names'])
# Target data_dict['target'] = 0 is malignant 1 is benign
data['malignant'] = (data_dict['target'] == 0)
data.columns
points = go.Scatter(x=data['mean radius'], y = 1.*data['malignant'], mode="markers")
layout = dict(xaxis=dict(title="Mean Radius"),yaxis=dict(title="Malignant"))
py.iplot(go.Figure(data=[points], layout=layout))
# This is a clear example of over-plotting. We can improve the above plot by jittering the data:
jitter_y = data['malignant'] + 0.1 * np.random.rand(data['malignant'].size) -0.05
points = go.Scatter(x=data['mean radius'], y = jitter_y,
mode="markers",
marker=dict(opacity=0.5))
py.iplot(go.Figure(data=[points], layout=layout))
# Perhaps a better way to visualize the data is using stacked histograms.
py.iplot(ff.create_distplot(
[data.loc[~data['malignant'], 'mean radius'],
data.loc[data['malignant'], 'mean radius']],
group_labels=["Benign","Malignant"],
bin_size=0.5))
# + [markdown] deletable=false editable=false
# ### Prediction rule
# Looking at the above histograms could you describe a rule to predict whether or a cell is malignant?
#
# <!--
# BEGIN QUESTION
# name: q1a
# manual: true
# -->
# <!-- EXPORT TO PDF -->
# -
# *Write your answer here, replacing this text.*
# # Least Squares Regression
#
# **Goal:** We would like to predict whether the tumor is malignant from the size of the tumor. We will be using least square regression to build a classifier that can achieve the objective.
# + [markdown] deletable=false editable=false
#
#
# ## Part 1a- Preparing the data Train-Test Split
# Always split your data into training and test groups. The model learns from the training examples and then we test our model on the test set. In this example we will first split the data using the train_test_split from sklearn. Keep 75% of the data for training and the remaining 25% for testing.
# <!--
# BEGIN QUESTION
# name: q1b
# -->
# -
from sklearn.model_selection import train_test_split
data_tr, data_te =...
...
print("Training Data Size: ", len(data_tr))
print("Test Data Size: ", len(data_te))
# + deletable=false editable=false
ok.grade("q1b");
# -
# ## Question 1b- Setting labels and Values
# Now let us visualize the data.
# We will define $X$ and $Y$ as variables containing the training features and labels.
# +
print(data_tr.head(5))
# + [markdown] deletable=false editable=false
# The row mean radius gives us the radius of each tumor. You will now be selecting the values from the mean radius and storing it in the data variable X. Similary "malignant" column tells us whether the tumor is malignant or not. In order to prepare the training labels you will store these values in the float format where 0 stands for false and 1 stands for true. This will be stored in variable Y.
#
# <!--
# BEGIN QUESTION
# name: q1b
# -->
# -
X=...
Y=...
...
# + deletable=false editable=false
ok.grade("q1b");
# + [markdown] deletable=false editable=false
# ## Part 2a- Fitting a least square regression module
# Once we are done with the basics of data modelling you can fit a least square regression module on the data. Follow the given instructions:
# 1. Use the `LinearRegression()` function to create a model for least square linear regression
# 2. Use the `fit()` function to fit the data $(X,Y)$
# <!--
# BEGIN QUESTION
# name: q2a
# -->
# +
import sklearn.linear_model as linear_model
# Call the linear regression model
least_squares_model = ...
# Now use the fit function
...
# + deletable=false editable=false
ok.grade("q2a");
# -
# # How is our fit?
jitter_y = Y + 0.1*np.random.rand(len(Y)) - 0.05
points = go.Scatter(name="Jittered Data",
x=np.squeeze(X), y = jitter_y,
mode="markers", marker=dict(opacity=0.5))
X_plt = np.linspace(np.min(X), np.max(X), 10)
model_line = go.Scatter(name="Least Squares",
x=X_plt, y=least_squares_model.predict(X_plt[:,np.newaxis]),
mode="lines", line=dict(color="orange"))
py.iplot([points, model_line])
# + [markdown] deletable=false editable=false
# ## Questions:
# 1. Are we happy with the fit?
# 2. What is the meaning of predictions that are neither 0 or 1?
# 3. Could we use this to make a decision?
#
# <!--
# BEGIN QUESTION
# name: q2a
# manual: true
# -->
# <!-- EXPORT TO PDF -->
# -
# *Write your answer here, replacing this text.*
# + [markdown] deletable=false editable=false
# ## Part 2b- What is the Root Mean Squared Error?
# Calcualte the mean squared error by using the mse module and predict function.
# <!--
# BEGIN QUESTION
# name: q2b
# -->
#
# -
from sklearn.metrics import mean_squared_error as mse
rmse=...
...
print("Training RMSE:",rmse )
# + deletable=false editable=false
ok.grade("q2b");
# -
#
#
# # Part 3-Classification Error
#
# This is a classification problem, so we probably want to measure how often we predict the correct value. This is sometimes called the zero-one loss (or error):
#
# $$ \large
# \textbf{ZeroOneLoss} = \frac{1}{n} \sum_{i=1}^n \textbf{I}\left[ y_i \neq f_\theta(x) \right]
# $$
#
# However, to use the classification error we need to define a decision rule that maps $f_\theta(x)$ to the $\{0,1\}$ classification values.
# + [markdown] deletable=false editable=false
# ---
#
#
# # Question 3a Simple Decision Rule
#
# Therefore, in order to solve the issue, we instituted the following simple decision rule:
#
# $$\Large
# \text{If } f_\theta(x) > 0.5 \text{ predict 1 (malignant) else predict 0 (benign).}
# $$
#
# This simple **decision rule** is deciding that a tumor is malignant if our model predicts a value above 0.5 (closer to 1 than zero).
#
# We will now be developing a classifier based on this simple rule. The output results are stored as boolean outcomes and are set to True for all values that are greater than 0.5 and False for all values that are less than 0.5.
#
# <!--
# BEGIN QUESTION
# name: q3a
# -->
# -
ind_mal=...
...
# + deletable=false editable=false
ok.grade("q3a");
# +
jitter_y = Y + 0.1*np.random.rand(len(Y)) - 0.05
ind_mal = least_squares_model.predict(X) > 0.5
mal_points = go.Scatter(name="Classified as Malignant",
x=np.squeeze(X[ind_mal]), y = jitter_y[ind_mal],
mode="markers", marker=dict(opacity=0.5, color="red"))
ben_points = go.Scatter(name="Classified as Benign",
x=np.squeeze(X[~ind_mal]), y = jitter_y[~ind_mal],
mode="markers", marker=dict(opacity=0.5, color="blue"))
dec_boundary = (0.5 - least_squares_model.intercept_)/least_squares_model.coef_[0]
dec_line = go.Scatter(name="Least Squares Decision Boundary",
x = [dec_boundary,dec_boundary], y=[-0.5,1.5], mode="lines",
line=dict(color="black", dash="dot"))
py.iplot([mal_points, ben_points, model_line,dec_line])
# + [markdown] deletable=false editable=false
# # Compute `ZeroOneLoss`
# You will now be computing tht zero one loss and predicting the fraction of the data that is incorrect.
# <!--
# BEGIN QUESTION
# name: q3b
# -->
# -
from sklearn.metrics import zero_one_loss
zerooneloss=...
...
print("Training Fraction incorrect:", zerooneloss)
# + deletable=false editable=false
ok.grade("q3b");
# + [markdown] deletable=false editable=false
# **Questions**
#
# 1. Are we happy with this error level?
# 1. What error would we get if we just guessed the label?
#
#
# <!--
# BEGIN QUESTION
# name: q3a
# manual: true
# -->
# <!-- EXPORT TO PDF -->
# -
# *Write your answer here, replacing this text.*
# # Guessing the Majority Class
#
# This is the simplest baseline we could imagine and one you should always compare against. Let's start by asking what is the majority class
print("Fraction of Malignant Samples:", np.mean(Y))
# If we guess the majority class **benign**, what accuracy would we get?
# You can figure this out from the above number
print("Guess Majority:", zero_one_loss(Y, np.zeros(len(Y))))
# This is standard example of a common problem in classification (and perhaps modern society): **class imbalance**.
#
#
#
#
#
#
#
#
#
# + [markdown] deletable=false editable=false
#
# # Part 4 Cross Validation of Zero-One Error
#
# You will now be performing one of the most popular techniques for evaluating a classification model. The techniques is known as cross-validation. Cross-validation refers to breaking the entire data-set into $n$ parts where the $n-1$ parts are used for training and one of the parts is used for validation. The cycle is repeated for each part. Finally, the overall error is calculated for each part.
#
# You will be performing a 3-fold cross validation in this section. Do the following
# 1. Call the linear regression model and fit it on `tr_ind` for X and Y
# 2. Predict the outcome of the model using `model.predict` for `te_ind` and store it in outcome
# 3. Calculate the zero one loss for the predicted values
# <!--
# BEGIN QUESTION
# name: q4a
# -->
# +
from sklearn.model_selection import KFold
kfold = KFold(3,shuffle=True, random_state=42)
linreg_errors = []
models = []
for tr_ind, te_ind in kfold.split(X):
# Create a linear regression model and fit it with the training data and indices
model=...
...
models.append(model)
# Predict the outcome on the test data
outcome = ...
# Calculate the zero one loss for the predicted solution
zerooneloss = ...
# Append the zerooneloss to linreg_errors variable
...
print("Min Validation Error: ", np.min(linreg_errors))
print("Median Validation Error:", np.median(linreg_errors))
print("Max Validation Error: ", np.max(linreg_errors))
print(models)
# + deletable=false editable=false
ok.grade("q4a");
# -
# We can visualize all the models and their decisions
# +
dec_lines = [
go.Scatter(name="Decision Boundary",
x = [(0.5 - m.intercept_)/m.coef_[0]]*2,
y=[-0.5,1.5], mode="lines",
line=dict(dash="dot"))
for m in models]
X_plt = np.linspace(np.min(X), np.max(X), 10)
model_lines = [
go.Scatter(name="Least Squares " + str(zero_one_loss(Y, m.predict(X) > 0.5)),
x=X_plt, y=m.predict(np.array([X_plt]).T),
mode="lines")
for m in models]
py.iplot([points] + model_lines + dec_lines)
# -
#
#
# # Can we think of the line as a _"probability"_?
#
#
# Not really. Probabilities are constrained between 0 and 1. How could we learn a model that captures this probabilistic interpretation?
#
#
# # Could we just truncate the line?
#
# Maybe.
#
# We can define the probability as:
#
# $$ \large
# p_i = \min\left(\max \left( x^T \theta , 0 \right), 1\right)
# $$
#
# which would look like:
def bound01(z):
u = np.where(z > 1, 1, z)
return np.where(u < 0, 0, u)
X_plt = np.linspace(np.min(X), np.max(X), 100)
p_line = go.Scatter(name="Truncated Least Squares",
x=X_plt, y=bound01(least_squares_model.predict(np.array([X_plt]).T)),
mode="lines", line=dict(color="green", width=8))
py.iplot([mal_points, ben_points, model_line, p_line, dec_line], filename="lr-06")
# So far, least squares regression seems pretty reasonable and we can "force" the predicted values to be bounded between 0 and 1.
#
#
# **Can we interpret the truncated values as probabilities?**
#
# Perhaps, but it would depend on how the model is estimated (more on this soon).
#
#
#
# # An Issue with Extreme Points
#
# It seems like large tumor sizes are indicative of malignant tumors. Suppose we observed a very large malignant tumor that is 100mm in mean radius. What would this do to our model?
#
#
# Let's add an extra data point and see what happens:
X_ex = np.vstack([X, [100]])
Y_ex = np.hstack([Y, 1.])
least_squares_model_ex = linear_model.LinearRegression()
least_squares_model_ex.fit(X_ex, Y_ex)
# +
X_plt = np.linspace(np.min(X)-5, np.max(X)+5, 100)
extreme_point = go.Scatter(
name="Extreme Point", x=[100], y=[1], mode="markers",
marker=dict(color="green", size=10))
model_line.line.color = "gray"
model_line_ex = go.Scatter(name="New Least Squares",
x=X_plt, y=least_squares_model_ex.predict(np.array([X_plt]).T),
mode="lines", line=dict(color="orange"))
dec_line.line.color = "gray"
dec_boundary_ex = (0.5 - least_squares_model_ex.intercept_)/least_squares_model_ex.coef_[0]
dec_line_ex = go.Scatter(
name="Decision Boundary",
x = [dec_boundary_ex, dec_boundary_ex], y=[-0.5,1.5], mode="lines",
line=dict(color="black", dash="dash"))
py.iplot([mal_points, ben_points,model_line, model_line_ex, dec_line, dec_line_ex, extreme_point])
# -
# ## Observing the resulting RMSE
print("Before:",
zero_one_loss(Y_ex, least_squares_model.predict(X_ex) > 0.5))
print("After:",
zero_one_loss(Y_ex, least_squares_model_ex.predict(X_ex) > 0.5))
# + [markdown] deletable=false editable=false
# Looking at the above results, explain what you observed.
#
# <!--
# BEGIN QUESTION
# name: q4b
# manual: true
# -->
# <!-- EXPORT TO PDF -->
# -
# *Write your answer here, replacing this text.*
# + [markdown] deletable=false editable=false
# # Submit
# Make sure you have run all cells in your notebook in order before running the cell below, so that all images/graphs appear in the output.
# **Please save before submitting!**
#
# <!-- EXPECT 4 EXPORTED QUESTIONS -->
# + deletable=false editable=false
# Save your notebook first, then run this cell to submit.
import jassign.to_pdf
jassign.to_pdf.generate_pdf('lab09.ipynb', 'lab09.pdf')
ok.submit()
| lab/lab09/lab09.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
score = np.random.randint(40,100,(10,5))
test_score = score[:5,:]
# # ndarry运算
test_score[test_score > 60] = 1
test_score
np.any(test_score > 90)
np.all(test_score > 90)
temp = score[:4 ,:4]
temp
np.where(temp>60 , 1, 0)
temp1 = score[:4,:4]
temp1
np.max(temp,axis=1)
np.argmax(temp, axis=1)
# # 数组间的运算
arr1 = np.array([[1,2,3,4],[5,6,7,8]])
arr1 + 1
lista = [1 ,2, 3, 4]
lista + 1
arr1 / 2
lista / 2
arr1 = np.array([[1,2,3,4],[5,6,7,8]])
arr2 = np.array([[1,2,3,4],[5,6,7,8]])
arr1 + arr2
| numpytest/jupyter/test7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Searching optimal parallelism configurations
#
# The objective of this notebook is to find the best possible configuration of the downloader. In general we will be interested in optimizing two KPIs:
# - the overall download time of a given set of byte ranges
# - the time to get the first chunks so that we can get started with processing
#
# We have observed in the `cold_latencies` notebook that download times between consecutive invocations of the same container do not change significantly, so we ignore that dimension in this analysis.
# +
import os
region_name="us-east-2"
binary_name="lambda"
aws_profile=os.environ["AWS_PROFILE"] # Specify the profile you want to use from your .aws/credentials file with the AWS_PROFILE env variable
MEGA = 1024*1024
# -
# #### Get the name of the lambda function deployed with the `infra` notebook
# lambda_name = !docker run \
# --rm \
# -v $HOME/.aws/credentials:/creds:ro \
# -v cloud-reader-tf:/mnt/state_vol \
# cloudfuse/cloud-reader-terraform output lambda_arn
lambda_name = lambda_name[0][1:len(lambda_name[0])-1]
print('lambda_name:', lambda_name)
# #### Define the function invokation routines and plotting
#
# We launch multiple invokations in parallel (~100) to get good averages statistics in different locations of the datacenters.
# +
from joblib import Parallel, delayed
import boto3
import json
import base64
import pandas as pd
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings('once')
def q90(x):
return x.quantile(0.9)
def q99(x):
return x.quantile(0.99)
# call the AWS Lambda functions with the reader binary
def invoke_function(initial_permits, release_rate, max_parallel, ranges, show_logs = False):
session = boto3.Session(profile_name=aws_profile)
client = session.client('lambda', region_name = region_name)
inputParams = {
"region": region_name,
"bucket": "cloudfuse-taxi-data",
"key": "synthetic/pattern-3gb",
"size": 1024*1024*16*200,
"ranges": ranges,
"max_parallel": max_parallel,
"initial_permits": initial_permits,
"release_rate": release_rate,
}
response = client.invoke(
FunctionName = lambda_name,
InvocationType = 'RequestResponse',
Payload = json.dumps(inputParams),
LogType='Tail' if show_logs else 'None'
)
if show_logs:
print(base64.b64decode(response['LogResult']).decode("utf-8") )
return json.load(response['Payload'])
# run `nb_execution` instances in parallel, each trying out all download parallelisms between 1 and `max_max_parallel`
def run_different_parallelism(chunk_megabytes, nb_execution, max_max_parallel):
def invoke_batch():
results = []
for max_parallel in range(1, max_max_parallel+1):
results.append({
"response": invoke_function(max_parallel, 1, max_parallel, [{"start": i*chunk_megabytes*MEGA, "length": chunk_megabytes*MEGA} for i in range(0,20)]),
"max_parallel": max_parallel,
"initial_permits": max_parallel,
"release_rate": 1,
})
return results
return [item for sublist in Parallel(n_jobs=nb_execution)(delayed(invoke_batch)() for i in range(nb_execution)) for item in sublist]
# run `nb_execution` instances in parallel, with a configuration that increases the number of parallel downloads from 1 to 16 gradually
def run_progressive_release(chunk_megabytes, nb_execution):
def invoke_one():
return [{
"response": invoke_function(1, 2, 16, [{"start": i*chunk_megabytes*MEGA, "length": chunk_megabytes*MEGA} for i in range(0,20)]),
"max_parallel": 16,
"initial_permits": 1,
"release_rate": 2,
}]
return [item for sublist in Parallel(n_jobs=nb_execution)(delayed(invoke_one)() for i in range(nb_execution)) for item in sublist]
# flatten the results to a pandas and plot them
def plot_bench(results):
downloads = []
for res in results:
for dl in res['response']['cache_stats']:
downloads.append({
"dl_duration": dl['dl_duration'],
"first_read": res['response']['range_durations'][0],
"last_read": res['response']['range_durations'][-1],
"initial_permits": res['initial_permits'],
"release_rate": res['release_rate'],
"max_parallel": res['max_parallel'],
})
df = pd.DataFrame(downloads)
df_grouped = df.groupby(['initial_permits', 'release_rate']).agg({
'dl_duration': 'mean',
'first_read': ['mean', q90, q99],
'last_read': ['mean', q90, q99]
}).reset_index(level='release_rate')
plt.figure();
different_parallelism_df = df_grouped[df_grouped['release_rate']==1].drop(columns=['release_rate'])
ax = different_parallelism_df.plot(figsize=[20,10], color=['blue', 'lightgreen', 'green', 'darkgreen', 'pink', 'red', 'brown'], grid=True)
progressive_release_df = df_grouped[df_grouped['release_rate']==2].drop(columns=['release_rate'])
progressive_release_df.plot(figsize=[20,10], color=['blue', 'lightgreen', 'green', 'darkgreen', 'pink', 'red', 'brown'], grid=True, ax=ax, marker='o', legend=False, use_index=False)
# -
# ### 10MB chunks
#
# **Note:** We ran the same experience at different points in time to check if the results where consistent.
# !date
res = run_different_parallelism(chunk_megabytes = 10, nb_execution = 100, max_max_parallel = 16)
res.extend(run_progressive_release(chunk_megabytes = 10, nb_execution = 100))
plot_bench(res)
# !date
res = run_different_parallelism(chunk_megabytes = 10, nb_execution = 100, max_max_parallel = 16)
res.extend(run_progressive_release(chunk_megabytes = 10, nb_execution = 100))
plot_bench(res)
# ### 100MB chunks
#
# We also check that the results are similar with larger chunks
# !date
res = run_different_parallelism(chunk_megabytes = 100, nb_execution = 50, max_max_parallel = 16)
res.extend(run_progressive_release(chunk_megabytes = 100, nb_execution = 50))
plot_bench(res)
# ### Conclusions:
#
# In earlier tests that can be found in the literature such as https://arxiv.org/abs/1911.11727 or the performance guildelines https://docs.aws.amazon.com/AmazonS3/latest/userguide/optimizing-performance-guidelines.html, each individual download request to S3 had a limited bandwidth. This made it very beneficial to have multiple downloads in parallel to increase the total throughput.
#
# These new results tell us a different story. A single connection to S3 mostly fills the maximum available bandwidth between the Lambda and S3 at ~100MB/s. We can see that whatever parallelism we choose, the time to download the whole file (indicated by the `last_read` KPI) remains more or less the same. Having some parallelism slightly helps to fully utilize the bandwidth, but it seems that a parallelism of **2** is enough.
#
# In this setup, the only KPI that remains to be optimized is the time to first read, to allow us to start the processing of the data as quickly as possible. Having a single download at the beginning then scaling up parallelism progressively helps maintaining a very high bandwidth for the first downloads to get the first chunks quickly, while benefiting from the optimal bandwidth utilization of parallel downloads.
#
# Interestingly, this hybrid approach also provides better total download performances (time to last read), especially with smaller chunks (10MB).
| notebooks/optimal_parallelism.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: loco
# language: python
# name: loco
# ---
# # Converts RGB, segmentation maps into COCO
segm_dir = '/home/locobotm/AVD/habitat_data_with_seg/seg_p' # folder with *.npy segm maps to be converted to COCO
img_dir = "/home/locobotm/AVD/habitat_data_with_seg/rgb"
habitat_semantic_json = "/home/locobotm/AVD/habitat_data_with_seg/info_semantic.json"
coco_file_name = "coco_habitat_apart0.json" # Dumps annotations to this file
# +
import numpy as np
import sys
if '/opt/ros/kinetic/lib/python2.7/dist-packages' in sys.path:
sys.path.remove('/opt/ros/kinetic/lib/python2.7/dist-packages')
import cv2
import random
import os
import numpy as np
import json
from matplotlib import pyplot as plt
from PIL import Image
from pycococreatortools import pycococreatortools
from IPython import embed
from tqdm import tqdm
from IPython.core.display import display, HTML
# -
fs = [x.split('.')[0] +'.jpg' for x in os.listdir(segm_dir)]
print("Creating COCO annotations for {} images".format(len(fs)))
# ### Create coco json for `fs`
# +
with open(habitat_semantic_json, "r") as f:
habitat_semantic_data = json.load(f)
INFO = {}
LICENSES = [{}]
# create categories out of it
CATEGORIES = []
for obj_cls in habitat_semantic_data["classes"]:
CATEGORIES.append({"id": obj_cls["id"], "name": obj_cls["name"], "supercategory": "shape"})
if obj_cls['name'] in ('floor', 'wall', 'ceiling', 'wall-plug'):
print(obj_cls['id'], obj_cls['name'])
coco_output = {
"info": INFO,
"licenses": LICENSES,
"categories": CATEGORIES,
"images": [],
"annotations": [],
}
# +
count = 0
for x in tqdm(fs):
image_id = int(x.split('.')[0])
# load the annotation file
try:
prop_path = os.path.join(segm_dir, "{:05d}.npy".format(image_id))
annot = np.load(prop_path).astype(np.uint8)
except Exception as e:
print(e)
continue
img_filename = "{:05d}.jpg".format(image_id)
img = Image.open(os.path.join(img_dir, img_filename))
image_info = pycococreatortools.create_image_info(
image_id, os.path.basename(img_filename), img.size
)
coco_output["images"].append(image_info)
# for each annotation add to coco format
for i in np.sort(np.unique(annot.reshape(-1), axis=0)):
try:
category_info = {"id": habitat_semantic_data["id_to_label"][i], "is_crowd": False}
if category_info["id"] < 1 or category_info["id"] in (31, 40, 93, 95):
# Exclude wall, ceiling, floor, wall-plug
continue
except:
print("label value doesnt exist for", i)
continue
binary_mask = (annot == i).astype(np.uint8)
annotation_info = pycococreatortools.create_annotation_info(
count, image_id, category_info, binary_mask, img.size, tolerance=2
)
if annotation_info is not None:
coco_output["annotations"].append(annotation_info)
count += 1
with open(coco_file_name, "w") as output_json:
json.dump(coco_output, output_json)
# -
# # Visualize using detectron2
# +
from detectron2.data import DatasetCatalog, MetadataCatalog
from detectron2.data.datasets import register_coco_instances
from detectron2.utils.visualizer import Visualizer, ColorMode
DatasetCatalog.clear()
MetadataCatalog.clear()
register_coco_instances('foobar', {}, coco_file_name, img_dir)
MetadataCatalog.get('foobar')
dataset_dicts = DatasetCatalog.get('foobar')
plt.figure(figsize=(12 , 8))
for d in random.sample(dataset_dicts, 5):
img = cv2.imread(d["file_name"])
visualizer = Visualizer(img[:, :, ::-1], metadata=MetadataCatalog.get('foobar'), scale=0.5)
vis = visualizer.draw_dataset_dict(d)
img = vis.get_image()
plt.imshow(img)
plt.show()
# -
| examples_and_tutorials/notebooks/active_vision/coco_creator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# !pip install scipy
# !pip install gtts
import pyttsx3
import pyaudio
import speech_recognition as sr
import webbrowser
import datetime
import pywhatkit
import os
import wave
import ipywidgets as widgets
from gtts import gTTS
# +
#Listen to our microphone and return the audio as text using google
q = ""
def transform():
r = sr.Recognizer()
with sr.Microphone() as source:
print("Mendengarkan")
audio = r.listen(source)
q = ""
try:
q = r.recognize_google(audio, language = 'id', show_all = False)
print("Saya Mendengarkan: " + q)
except sr.UnknownValueError:
print("Maaf saya tidak mengerti")
return "Saya Menunggu"
except sr.RequestError as e:
print("Permintaan Gagal; {0}".format(e))
return q
# -
transform ()
# +
transform()
# q = "tolong"
if q == "jangan mendekat" or "menjauh" or "lepaskan" or "lepasin" or "berhenti" or "tolong":
import sounddevice
from scipy.io.wavfile import write
frekuensi_sample = 44100
# second = int(input("Enter time duration in seconds: "))
# second = 80000 # sama dengan 2 hari
waktu_rekam = 10
print("Recording.....\n")
record_voice = sounddevice.rec( int ( waktu_rekam * frekuensi_sample ) , samplerate = frekuensi_sample , channels = 2 )
sounddevice.wait()
write("out2.wav",frekuensi_sample,record_voice)
print("Finished.....\nPlease check your output file")
# -
| I-Safe.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# +
import json
import matplotlib.pylab as plt
import numpy as np
import os
import lib.helper as hl
import lib.cell_plot as cplt
% matplotlib inline
# +
# plotting current at the synapse(s) of a cell from cell_file
region_type = 'hippocamp'
neuron_type = 'exc'
current_dir = os.getcwd()
dir_name = os.path.join(current_dir, 'data', 'results')
file_name = '2018_12_5_14_1_all'
cell_file = 'c10861.CNG1.npz'
save_cell_figs_dir = os.path.join(dir_name,file_name,'cell/figs')
hl.create_folder(save_cell_figs_dir)
ext = '.png'
# +
params_cell_file = os.path.join(dir_name, file_name) + '.json'
with file(params_cell_file, 'r') as fid:
params_cell = json.load(fid)
results_cell = os.path.join(dir_name, file_name, 'cell', 'results', cell_file)
results = np.load(results_cell)
# access current at the synapses
synapse = results['ppt_vecs']
v_soma = results['v_soma']
dt = params_cell['timestep']
timeline = np.linspace(0, (len(v_soma))*dt, len(v_soma))
ax = plt.subplot(111)
plt.plot(timeline,synapse.T[:]*(-1),lw=2)
plt.ylabel('current at the synapse (nA)')
plt.xlabel('time')
plt.xlim([timeline[0], timeline[-1]])
plt.ylim([(np.max(np.abs(synapse))+np.max(np.abs(synapse))*0.3)*(-1), (np.max(np.abs(synapse))+np.max(np.abs(synapse))*0.3)])
plt.savefig(save_cell_figs_dir+'/synapses'+cell_file[:-4]+ext)
cplt.clean_plot(ax)
print 'saved fig: ' +save_cell_figs_dir+ '/synapses'+cell_file+ext
# -
| plot_cell_results.py.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Plotting alignment data
# +
# %matplotlib inline
import pandas as pd
import utils.db_utils as db
import utils.plot_utils as plot
from config import getConfig
############################################
# get configuration
cfg = getConfig() # configure values in config.js
############################################
targetBibleType = cfg['targetBibleType']
origLangPathGreek = cfg['origLangPathGreek']
origLangPathHebrew = cfg['origLangPathHebrew']
targetLanguagePath = cfg['targetLanguagePath']
dbPath = cfg['dbPath']
testamentStr = cfg['testamentStr']
baseDataPath = cfg['baseDataPath']
targetLang = 'en'
connections = db.initAlignmentDB(dbPath)
connection = db.getConnectionForTable(connections, 'default')
connection_owi = db.getConnectionForTable(connections, db.original_words_index_table)
# + pycharm={"name": "#%%\n"}
# find all alignments for this lemma
def findAlignmentsForWord(connection_owi, word, minAlignments, searchLemma=False, maxRows=None):
alignmentsByWord = db.findAlignmentsFromIndexDbForOrigWord(connection_owi, word, searchLemma, maxRows)
alignmentsList, rejectedAlignmentsList = db.filterAlignments(alignmentsByWord, minAlignments)
return pd.DataFrame(alignmentsList)
word = 'θεός' # Found 1354
# word = 'καί' # Found 8995
# word = 'ὁ' # Found 20377
# word = 'αὐτός' # Found 5573
minAlignments = 0
lemmaAlignments = findAlignmentsForWord(connection_owi, word, minAlignments, searchLemma=True)
print(f"Found {len(lemmaAlignments)} alignments")
# + pycharm={"name": "#%%\n"}
# find all alignments for this original word
# word = 'Θεός' # found 69
# word = 'Θεὸς' # found 239
word = 'Θεοῦ' # found 712
# word = 'καί' # Found 32
# word = 'καὶ' #Found 8961
# word = 'τὸ' # Found 1715
# word = 'αὐτοῦ' # Found 1415
origAlignments = findAlignmentsForWord(connection_owi, word, minAlignments, searchLemma = False)
print(f"Found {len(origAlignments)} alignments")
# + pycharm={"name": "#%%\n"}
db.describeAlignments(origAlignments)
# -
# ### Analysis of alignments for Θεοῦ in the en_ult:
# #### Frequency of alignments:
# + pycharm={"name": "#%%\n"}
frequency = origAlignments['alignmentText'].value_counts()
print(frequency)
# -
# ##### Notes:
# - the left column is the specific alignment, and the right column is the number of times that specific alignment has been made so far in the NT.
# - alignments that contain more words are more suspect.
# - in future will combine "God s" to "God's" before doing analysis
# <p></p>
# + pycharm={"name": "#%%\n"}
plot.plotFieldFrequency(frequency, "", 'alignment', title="Frequency of Alignments", xNumbers=False, xShowTicks=False)
# -
# ### Analysis:
# #### Analysis of numerical metrics:
# + pycharm={"name": "#%%\n"}
descr = origAlignments.describe()
print(f"Alignments description:\n{descr}")
# -
# #### Analysis of original language word count:
# + pycharm={"name": "#%%\n"}
field = 'origWordsCount'
field_frequency = origAlignments[field].value_counts().sort_index()
print(f"\nFrequency of {field}:\n{field_frequency}")
# -
# ##### Notes:
# - this field analysis suggests for θεός nearly all the original language word counts are tight. The word counts of 3 may need review. So we could probaby use that as a threshold for to flag for review.
# <p></p>
# + [markdown] pycharm={"name": "#%% md\n"}
# #### Analysis of target language word count:
# + pycharm={"name": "#%%\n"}
field = 'targetWordsCount'
field_frequency = origAlignments[field].value_counts().sort_index()
print(f"\nFrequency of {field}:\n{field_frequency}")
# -
# ##### Notes:
# - this field analysis suggests that for θεός likely all the target language word counts are tight. The word count of 3 probably good for English (`of a god`). But still we could probaby use that as a threshold for to flag for review.
# <p></p>
# #### Analysis of count of extra unaligned words between aligned original language words:
# + pycharm={"name": "#%%\n"}
field = 'origWordsBetween'
field_frequency = origAlignments[field].value_counts().sort_index()
print(f"\nFrequency of {field}:\n{field_frequency}")
plot.plotFieldFrequency(field_frequency, field, f"Words Between", max=10)
# -
# ##### Notes:
# - this field analysis suggests that most original language alignments probably good. Probably the cases of a word between (count > 0) aligned words should be reviewed.
# <p></p>
# #### Analysis of count of extra unaligned words between aligned target language words:
# + pycharm={"name": "#%%\n"}
field = 'targetWordsBetween'
field_frequency = origAlignments[field].value_counts().sort_index()
print(f"\nFrequency of {field}:\n{field_frequency}")
plot.plotFieldFrequency(field_frequency, field, f"Words Between", max=10)
# + [markdown] pycharm={"name": "#%% md\n"}
# ##### Notes:
# - this field analysis suggests that most target language alignments probably good. Large gaps between aligned words are likely due to wordmap suggesting wrong occurence of a word and the user selecting. Probably the cases of a word between (count > 0) aligned words should be reviewed.
# <p></p>
# + pycharm={"name": "#%%\n"}
# + pycharm={"name": "#%%\n"}
| plot_original_word_alignments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Functions to export
def column_distplot(dataframe, column, xlabel=None):
'''Plot the histogram of a column using Seaborn'''
# Set Figure
sns.set(rc={'figure.figsize':(10,5)},style="white", context="talk")
# Plot
column = column
data = dataframe[column][~dataframe[column].isna()]
ax = sns.distplot(data);
# Title and Axis
ax.set_title("Histogram of the fighters' {}".format(column));
if type(xlabel)==str:
ax.set_xlabel(xlabel)
else:
ax.set_xlabel(str(column).capitalize())
sns.despine()
col_cap = str(column).capitalize()
print("{} Skewness: {}".format(col_cap, round(dataframe[column].skew(), 2)))
print("{} Kurtosis: {}".format(col_cap, round(dataframe[column].kurt(), 2)))
print('{} Mean: {}'.format(col_cap, round(data.mean(), 2)))
print('{} Median: {}'.format(col_cap, data.median()))
def parse_height(height):
'''Splits the height value into feet and inches'''
ht_ = height.split(" ")
ft_ = float(ht_[0])
in_ = float(ht_[1])
return (12*ft_) + in_
# ## Cleaning fighter_home_country data
def generalize_columns(df):
'''Make the column headings lowercase and generalized (remove the B_ and R_)'''
# Make lowercase
df.columns = df.columns.str.lower()
# Split on the _
df_columns = df.columns.str.split("_")
# Take the last word in the inner list
df_columns = [el[-1] for el in df_columns]
# Change the names of columns
df.columns = df_columns
return df
def clean_hometown(data_both_fighters):
'''Cleans up the hometown column and returns the home_country alongside the name'''
# Split the hometown column into
data_both_fighters['hometown'] = data_both_fighters['hometown'].str.split(" ")
data_both_fighters.dropna(subset=['hometown'], inplace=True)
for row in data_both_fighters['hometown']:
# If the list ends in a space, pop it off
if len(row[-1]) == 0:
row.pop()
# Replace [United, States] with [USA]
if row[-1] == 'States':
row[-1] = 'USA'
# Change 'Michigan' to 'USA'
if row[-1] == 'Michigan':
row[-1] = 'USA'
# Strip out the row to only the last element
data_both_fighters['home_country']=list(map(lambda x: x[-1], data_both_fighters['hometown']))
# Drop the 'hometown column'
data_both_fighters.drop(columns='hometown', axis=1, inplace=True)
# Reset the index
data_both_fighters.reset_index(drop=True, inplace=True)
return data_both_fighters
def clean_fighter_data(data):
'''Cleans up the dataframe and returns the df with unique fighters names, home_country'''
# Split dataframe into dataframe for the Blue fighter and another dataframe for the Red fighter
data_b_fighter = data[['B_Name', 'B_HomeTown']]
data_r_fighter = data[['R_Name', 'R_HomeTown']]
# Change the column headings to lowercase and remove the B_ and R_
[generalize_columns(df) for df in [data_b_fighter, data_r_fighter]]
# Concatenate the Blue fighters data to the Red fighters data
data_both_fighters = pd.concat([data_b_fighter, data_r_fighter], ignore_index=True)
# Filter the data to only unique names
data_both_fighters.drop_duplicates(subset='name', keep='first', inplace=True)
clean_hometown(data_both_fighters)
return data_both_fighters
def show_na_cols(dataframe):
if dataframe.isna().sum().any():
print('The columns with na values in the dataframe:')
for column in dataframe.columns:
cols_na = dataframe[column].isna().sum()
if cols_na:
print('\t{}:\t'.format(column), cols_na)
else:
print('There are no columns with na values in the dataframe')
def calc_age_at_fight(data, new_col_name, fighter_dob, date='date',):
'''Calculate the age of a fighter at the time of the fight, based on their date of birth'''
# Calculate the time difference between 2 dates (in days)
data[new_col_name] = data['date'] - data[fighter_dob]
# Remove the 'days' value in column
data[new_col_name] /= np.timedelta64(1, 'D')
# Convert form days to years
data[new_col_name] //= 365
# Doesnt work
def drop_bouts_with_no_fighter_stats(data, fighters_data):
'''Take out the bouts where we do not have data on both fighters'''
original_bout_length = len(data)
# Create a list of all the fighters' names
name_list = fighters_data.index.tolist()
# Create booleans for whether the name in fighter1/figther2 column appears in our list of fighter names
f1_in_name_list = data['fighter1'].isin(name_list)
f2_in_name_list = data['fighter2'].isin(name_list)
# Drop out the rows where at least 1 of names does not appear in our fighter names list
data = data[(f1_in_name_list) & (f2_in_name_list)]
# Print the action we have carried out
new_bout_length = len(data)
print('We have reduced the number of bouts from {} to {}, as we do not have statistics for {} fighters'
.format(original_bout_length, new_bout_length, original_bout_length-new_bout_length))
return data
# # EDA & Data Cleaning - [Your Project Name Here]
# ## Local Code Imports - Do not delete
# + tags=["hidecode"]
# DO NOT REMOVE THESE
# %load_ext autoreload
# %autoreload 2
# -
# DO NOT REMOVE This
# %reload_ext autoreload
# +
## DO NOT REMOVE
## import local src module -
## src in this project will contain all your local code
## clean_data.py, model.py, visualize.py, custom.py
from src import make_data as mk
from src import visualize as viz
from src import model as mdl
from src import pandas_operators as po
from src import custom as cm
def test_src():
mk.test_make_data()
viz.test_viz()
mdl.test_model()
po.test_pandas()
return 1
# -
test_src()
# ## Code Imports
# +
# Dataframes
import numpy as np
import pandas as pd
# Visualization
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
# Calculating age of fighters
import datetime
# -
# # Project Overview
#
# ## Background
# UFC BACKGROUND
#
# PROJECT OBJECTIVE
# # Data Understanding
# + [markdown] heading_collapsed=true
# ## Bouts Data
# + [markdown] hidden=true
# ### Bouts Data - Description
# + [markdown] hidden=true
# RESERVED FOR BOUTS DATA DESCRIPTION
# + [markdown] hidden=true
# #### Bouts Data - Data Dictionary
# + [markdown] hidden=true
# RESERVED SPACE FOR BOUTS DATA DICTIONARY
# + [markdown] hidden=true
# ### Bouts Data - Import and Summary
# + hidden=true
data_bouts = pd.read_csv('../data/raw/ufc_bouts.csv', parse_dates=['date'])
data_bouts.head(3)
# + hidden=true
print('DataFrame Shape is {}'.format(data_bouts.shape))
print('This represents {} fights'.format(data_bouts.shape[0]))
# + hidden=true
data_bouts.info()
# + [markdown] hidden=true
# ### Bouts Data - Exploration & Cleaning
# + hidden=true
if data_bouts.shape[0] == data_bouts['bout_id'].nunique():
print('Each sample (row) in our bouts data is unique')
# + [markdown] hidden=true
# #### Results column - drop nc and draw rows
# + hidden=true
data_bouts.result.value_counts()
# + [markdown] hidden=true
# - The no contests are either due to illegal moves or failed drugs tests.
# - Example below between <NAME> and <NAME> was a no contest due to <NAME> failing a drugs test
# - https://www.mmamania.com/2015/2/6/7992549/no-contest-anderson-silvas-victory-over-nick-diaz-at-ufc-183
# + hidden=true
# Example of no contest between <NAME> and <NAME>
bout_nc = data_bouts['result']=='nc'
bout_anderson_silva = data_bouts['fighter1']=='<NAME>'
data_bouts[(bout_nc) & (bout_anderson_silva)]
# + hidden=true
print('Result Column Value Counts:\n', data_bouts.result.value_counts(), '\n')
# Drop NC rows as they dont help us predict the outcome
data_bouts = data_bouts[data_bouts['result']!='nc']
# drop the draw rows for now, so that we have a binary outcome of win or lose
data_bouts = data_bouts[data_bouts['result']!='draw']
print('Result Column Value Counts After Dropping nc AND draw:\n', data_bouts.result.value_counts())
# + [markdown] hidden=true
# #### Location column - shorten
# + hidden=true
# Strip the location column down to only the country
data_bouts['location'] = data_bouts['location'].str.split(" ").str.get(-1)
# Rename the location column to bout_location
data_bouts.rename({'location': 'bout_location'}, inplace=True)
data_bouts.head(2)
# + [markdown] hidden=true
# #### Method column - Reduce number of categories & Dummy code
# + hidden=true
print('- There were originally {} categories of win method\n'.format(data_bouts.method.value_counts().count()))
# Reduce the categories down
data_bouts['method'] = data_bouts['method'].str.split("-").str.get(0)
print('- Now there are only {} categories of win method: \n{}'.format(data_bouts.method.value_counts().count(), data_bouts.method.value_counts()))
# + hidden=true
bout_methods = pd.get_dummies(data_bouts['method'], prefix='method', prefix_sep='_')
data_bouts = pd.concat([data_bouts, bout_methods], axis=1)
data_bouts.drop(columns='method', inplace=True)
data_bouts.head(2)
# + [markdown] hidden=true
# #### Shuffle the fighter1 and fighter2
# + hidden=true
fighter1_is_winner = data_bouts['fighter1'].equals(data_bouts['winner'])
if fighter1_is_winner:
print('The wiiner is always fighter1. We need to randomize the winner between fighter1 and fighter2')
# + hidden=true
# Randomly choose half of the observations to swap its fighter1 value with its fighter2 value
bout_rows_to_shuffle = np.random.choice(len(data_bouts), size=len(data_bouts) // 2, replace=False)
# Column location of fighter1 and fighter2
f1_loc = data_bouts.columns.get_loc('fighter1')
f2_loc = data_bouts.columns.get_loc('fighter2')
# Swap the values
data_bouts.iloc[bout_rows_to_shuffle, [f1_loc, f2_loc]] = data_bouts.iloc[bout_rows_to_shuffle, [f2_loc, f1_loc]].values
# + [markdown] hidden=true
# #### title_fight column to binary
# + hidden=true
data_bouts['title_fight'] = data_bouts['title_fight'] == 't'
data_bouts['title_fight'] = data_bouts['title_fight'].astype(int)
# + [markdown] hidden=true
# #### winner column to binary
# + [markdown] hidden=true
# - To avoid the confusion of a value of 0 meaning fighter1 wins, and value of 1 meaning fighter2 wins, we need to change the winner column name
# + hidden=true
# Change winner column name
data_bouts.rename(index=str, columns={'winner': 'winner_is_fighter1'}, inplace=True)
# Change to binary
data_bouts.winner_is_fighter1 = data_bouts.winner_is_fighter1==data_bouts.fighter1
data_bouts.winner_is_fighter1 = data_bouts.winner_is_fighter1*1
# + [markdown] hidden=true
# #### Keep only useful columns
# + hidden=true
cols_to_keep = ['date', 'location', 'fighter1', 'fighter2', 'winner_is_fighter1', 'title_fight', 'method_DEC', 'method_DQ', 'method_KO/TKO', 'method_SUB']
data_bouts = data_bouts[cols_to_keep]
data_bouts.head(3)
# + [markdown] hidden=true
# #### Reset index
# + hidden=true
data_bouts.reset_index(inplace=True, drop=True)
# + [markdown] hidden=true
# ### Save the cleaned bouts df to processed data file
# + hidden=true
data_bouts.to_csv('../data/processed/bouts_cleaned', index=False)
# + [markdown] hidden=true
# - With so many columns, the .info method doesn't give us much insight. We need to look at different sections of the dataframe individually.
# -
# ## Fighters Data
# ### Fighters Data - Description
# RESERVED FOR FIGHTERS DATA DESCRIPTION
# #### Fighters Data - Data Dictionary
# - SLpM - Significant Strikes Landed per Minute
# - Str_Acc - Significant Striking Accuracy
# - SApM - Significant Strikes Absorbed per Minute
# - Str_Def - Significant Strike Defence (the % of opponents strikes that did not land)
# - TD_Avg - Average Takedowns Landed per 15 minutes
# - TD_Acc - Takedown Accuracy
# - TD_Def - Takedown Defense (the % of opponents TD attempts that did not land)
# - Sub_Avg - Average Submissions Attempted per 15 minutes
# ### Fighters Data - Import and Summary
data_fighters.shape
data_fighters = pd.read_csv('../data/raw/ufc_fighters.csv', parse_dates=['dob'])
data_fighters.head(3)
print('DataFrame Shape is {}'.format(data_fighters.shape))
print('This represents {} fighters'.format(data_fighters.shape[0]))
data_fighters.info()
# - We can see that there are few features which should be in a number format, but are shown as objects (in most cases they are strings).
# - These features are height, weight, Str_Acc, Str_Def, TD_Acc and TD_Def
# - We will need to clean these features before we can explore them further
# ### Fighters Data - Exploration & Cleaning
# #### Drop % signs
pct_cols_to_change = ['Str_Acc', 'Str_Def', 'TD_Acc', 'TD_Def']
for col in pct_cols_to_change:
data_fighters[col] = data_fighters[col].str.replace('%','')
data_fighters.head(2)
# #### NaN values
# Most of the computational tools that we will be using cannot handle missing values, or at the very least produce unpredictable results. We must therefore address these missing values in our dataset. We have several options for this
# - eliminate missing values: This is a very simple method however we may end up removing too many samples, which would make it impossible for our model to distinguish between classes.
# - impute missing values: This involves guessing what the values could be using values such as mean, median or even mode.
data_fighters.isna().sum().sort_values(ascending=False)[:6]
# - We will deal with these NaN values further down
# #### Reach column
# +
# Strip the " sign and convert to type int
data_fighters['reach'] = data_fighters['reach'].str.replace('"','')
# Convert from type string, to type float
data_fighters['reach'] = data_fighters['reach'].astype(float)
# -
column_distplot(data_fighters, 'reach')
# - As this reach distribution is approximately normally distributed with no extreme outliers, we will fill the na values with the mean value of reach
data_fighters.reach = data_fighters.reach.fillna(round(data_fighters.reach.mean(), 0))
# #### Stance column
cm.column_countplot(data_fighters, 'stance', show_count=True)
# - The vast majority of fighters fight in the orthodox stance, so it is easiest to fill the na values with this modal average
stance_mode_avg = data_fighters.stance.mode()[0]
data_fighters.stance.fillna(stance_mode_avg, inplace=True)
# #### Dummy code Stance column
data_fighters = pd.get_dummies(data=data_fighters, columns=['stance'])
# rename so the column heading is 1 word
data_fighters.rename(columns = {'stance_Open Stance': 'stance_Open_Stance'}, inplace = True)
# #### Calc average age to deal with NaN in dob column
# need to keep one of these tables to easily add it in at the end of cleaning
# How many dobs are missing?
missing_dobs = data_fighters['dob'].isna().sum() / data_fighters.shape[1]
print('{}% of fighter data of births are missing'.format(round(missing_dobs, 2)))
bouts_dobs = cm.calc_average_age(data_fighters, data_bouts, drop_na=True)
bouts_dobs.head(3)
# Make 2 lists of ages into 1 list
fighter_ages = list(bouts_dobs['fighter1_age']) + list(bouts_dobs['fighter2_age'])
average_age = cm.plot_sns_displot_ages(fighter_ages,
'Age of Fighters',
plot_median=True)
# Fill NaN age values with median age
bouts_dobs = cm.calc_average_age(data_fighters, data_bouts, drop_na=False)
median = average_age['median']
bouts_dobs.fillna(value={'fighter1_age': median, 'fighter2_age': median}, inplace=True)
# #### dob column
# +
# # Create the year of birth column (dob_year)
# data_fighters['dob_year'] = 0
# for index, value in enumerate(data_fighters['dob']):
# data_fighters['dob_year'][index] = data_fighters['dob'][index].year
# today = datetime.date.today()
# # Fighter's age_today = difference in today's year and their year of birth
# data_fighters['age_today'] = today.year - data_fighters.dob_year
# +
# column_distplot(data_fighters, 'age_today', xlabel='Age of Fighters Today')
# -
# - The distribution of fighter ages is fairly normal, but due to the skewness and kurtosis we will choose to use the median age to fill in the NaN values.
# +
# # Calculate the median date of birth
# dates = list(data_fighters.sort_values('dob')['dob'])
# median_dob = dates[len(dates)//2]
# print('The median date of birth is {}-{}-{}'.format(median_dob.year, median_dob.month, median_dob.day))
# # Fill na values with this median date of birth
# data_fighters.dob.fillna(median_dob, inplace=True)
# -
# #### height column
# Drop ' and " signs
data_fighters['height'] = data_fighters['height'].str.replace("'","")
data_fighters['height'] = data_fighters['height'].str.replace('"',"")
# Calculate height in inches based off the feet and inches in height column
# Note that we can only perform the operation on columns that are not NaN
height_not_na = ~data_fighters['height'].isna()
data_fighters['height_inches'] = data_fighters['height'][height_not_na].apply(lambda x: parse_height(x))
column_distplot(data_fighters, 'height_inches')
# - We will use the mean height to fill in NaN values for height
# +
height_inches_not_na = data_fighters['height_inches'][~data_fighters['height_inches'].isna()]
height_mean = height_inches_not_na.mean()
print('The mean height of a fighter is {} inches'.format(round(height_mean, 1)))
# Fill na values with this mean height
data_fighters.height_inches.fillna(height_mean, inplace=True)
# + [markdown] heading_collapsed=true
# #### Remove fighters with limited statistics
# + hidden=true
# Original number of fighters
total_fighters = len(data_fighters)
print('We have {} fighters in our original data.'.format(total_fighters))
# Fighters with no recorded stats
fight_statistics = ['SLpM', 'Str_Acc', 'SApM']
no_stat_fighters = len(data_fighters[(data_fighters[fight_statistics]==0).any(axis=1)])
print('There are {} fighters that have no statistics for the columns: {}.'.format(no_stat_fighters, fight_statistics))
# Reduced dataset size
data_fighters = data_fighters[~(data_fighters[fight_statistics]==0).any(axis=1)]
stat_fighters = len(data_fighters)
print('We therefore reduce the number of fighters in our data to {}.'.format(stat_fighters))
# Reset the index
data_fighters.reset_index(drop=True, inplace=True)
# + [markdown] heading_collapsed=true
# #### Check for Duplicate Names
# + hidden=true
if len(data_fighters) == data_fighters.fighter_id.nunique():
print('The fighter_id column has only unique values')
# + hidden=true
# calculate how many duplicated fighter names there are
non_unique_names = data_fighters.shape[0] - data_fighters.name.nunique()
print('The {} name(s) in the dataframe that is/are not unique are:'.format(non_unique_names))
# print out duplicated figher names
duplicate_names = list(data_fighters['name'][data_fighters.duplicated(subset="name")])
for i, name in zip(range(1,len(duplicate_names)+1), duplicate_names):
print('\t{})'.format(i), name)
# + hidden=true
data_fighters[data_fighters.duplicated(subset="name", keep=False)]
# + [markdown] hidden=true
# - It appears that each fighter is unique in the table above, as statistics such as date of birth (dob) and weight differ.
# - Upon further investigation, these are indeed two different fighters, with the 1st fighter (born 1988) changing his ring name to <NAME> Ma to avoid the confusion with <NAME> (born 1981)
# - We shall change the name of the 1st fighter <NAME> (born 1988) to <NAME>
# + hidden=true
# Check if there are any Dong Hyun Ma in the dataset
if len(data_fighters[data_fighters.name=='<NAME>']):
print('Dong Hyun Ma already exists')
else:
# Find the right <NAME> Kim, who is now <NAME> Ma
DongHyunMa = data_fighters['name']=='<NAME>'
DongHyunMa_dob = data_fighters['dob']=='1988-09-09'
# Change name to <NAME>
data_fighters['name'][DongHyunMa & (DongHyunMa_dob)] = '<NAME>'
print('<NAME> 1988 has been renamed as <NAME>')
# + hidden=true
# -
# #### Drop Unecessary Columns
# We now need to drop variables that have no likely impact on the outcome of a fight.
# - Weight: Although in reality a fighter's weight is extremely important, within a single fight, both fighter's weights will be within a narrow window and very similar to each other, i.e. any difference will be negligible
# +
# Columns to drop
cols_to_drop = ['fighter_id', 'nc', 'height', 'weight', 'last_updated']
# Drop the columns
data_fighters.drop(cols_to_drop, axis=1, inplace=True)
# View the narrower fighters dataframe
data_fighters.head(3)
# -
# #### Add total_bouts column
# don't include nc values, as they do not useful in predicting a winner
# Also exclude draws as we want a binary outcome of win or lose
data_fighters['total_bouts'] = data_fighters['win'] + data_fighters['lose']
# #### Add win rate column
data_fighters['win_rate'] = round(data_fighters['win'] / data_fighters['total_bouts'], 2)
# #### Rearrange columns and group similar columns together
fighter_columns = data_fighters.columns
fighter_columns
data_fighters = data_fighters[['name', 'dob',
'win', 'lose', 'draw',
'total_bouts', 'win_rate',
'height_inches', 'reach',
'stance_Open_Stance', 'stance_Orthodox',
'stance_Sideways', 'stance_Southpaw',
'stance_Switch',
'SLpM', 'Str_Acc', 'SApM', 'Str_Def',
'TD_Avg', 'TD_Acc', 'TD_Def', 'Sub_Avg']]
# #### Change all column headings to lower
data_fighters.columns = data_fighters.columns.str.lower()
# ### Fighters Data - Further Exploration
np.round(data_fighters.describe(), 2)
data_fighters[data_fighters.lose>30]
# #### set name column into index
data_fighters.set_index('name', inplace=True)
# #### Save the cleaned fighters df to processed data file
data_fighters.to_csv('../data/processed/fighters_cleaned')
# ## Home Country Data
fighter_stats_each_bout = pd.read_csv('../data/raw/fighter_stats_each_bout.csv.zip')
fighter_stats_each_bout.head()
fighter_home_country = clean_fighter_data(fighter_stats_each_bout)
fighter_home_country.to_csv('../data/processed/fighter_home_country', index=False)
# ## Combine Dataframes
# - Here we combine the bouts data with the fighters data, into 1 dataframe that will be used for the machine learning process
# - We can only look at bouts where we have information on both fighters
bouts_cleaned = pd.read_csv('../data/processed/bouts_cleaned')
fighters_cleaned = pd.read_csv('../data/processed/fighters_cleaned',
index_col=0)
bouts_cleaned.location.value_counts()
bouts_cleaned.head(3)
# ### Drop bouts where we have no data on fighters
# +
original_bout_length = len(bouts_cleaned)
# Create a list of all the fighters' names
name_list = fighters_cleaned.index.tolist()
# booleans if name in fighter1/figther2 column is in list of fighter names
f1_in_name_list = bouts_cleaned['fighter1'].isin(name_list)
f2_in_name_list = bouts_cleaned['fighter2'].isin(name_list)
# Drop rows where at least 1 of names does not appear in fighter names list
bouts_cleaned = bouts_cleaned[(f1_in_name_list) & (f2_in_name_list)]
# Print the action we have carried out
new_bout_length = len(bouts_cleaned)
print('We have reduced the number of bouts from {} to {}, as we do not have statistics for {} fighters'
.format(original_bout_length, new_bout_length, original_bout_length-new_bout_length))
# -
# ### Create 2 new dataframes for fighter1 data and fighter2 data
# +
# Pull in the statistics for each fighter1
data_fighter1 = fighters_cleaned.loc[bouts_cleaned["fighter1"]]
# Pull in the statistics for each fighter2
data_fighter2 = fighters_cleaned.loc[bouts_cleaned["fighter2"]]
# Add suffix to each new dataframe's columns so we can merge them
data_fighter1 = data_fighter1.add_prefix('fighter1_')
data_fighter2 = data_fighter2.add_prefix('fighter2_')
# Reset the indices so we can merge them
bouts_cleaned.reset_index(inplace=True, drop=True)
data_fighter1.reset_index(inplace=True)
data_fighter2.reset_index(inplace=True)
# -
# ### Add Home Country to fighter1 and fighter2
data_fighter1 = data_fighter1.merge(fighter_home_country, how='left')
data_fighter2 = data_fighter2.merge(fighter_home_country, how='left')
# +
# Rename columns so we can keep track of whos home country it is
data_fighter1.rename(index=str, columns={'home_country': 'fighter1_home_country'}, inplace=True)
data_fighter2.rename(index=str, columns={'home_country': 'fighter2_home_country'}, inplace=True)
data_fighter1.index = data_fighter1.index.astype('int64')
data_fighter2.index = data_fighter2.index.astype('int64')
# -
# ### Create a combined dataframe of bouts with details of fighter1 and fighter2
# Create a merged dataframe
data_combined = pd.concat([bouts_cleaned, data_fighter1, data_fighter2], axis=1)
data_combined.tail(3)
# Drop the name columns (2 of them)
data_combined.drop(columns='name', inplace=True)
# ### Add fighter age at time of fight
data_combined = data_combined.join(bouts_dobs[['fighter1_age', 'fighter2_age']])
# +
# # Convert appropriate columns to datetime
# date_cols = ['date', 'fighter1_dob', 'fighter2_dob']
# for col in date_cols:
# data_combined[col] = pd.to_datetime(data_combined[col], format='%Y-%m-%d', errors='coerce')
# +
# # Calculate age of fighters at time of fight
# new_cols = ['fighter1_age_at_fight', 'fighter2_age_at_fight']
# dobs = ['fighter1_dob', 'fighter2_dob']
# for i in list(range(len(new_cols))):
# calc_age_at_fight(data_combined, new_cols[i], dobs[i])
# -
# ### Add home advantage column
# +
# 1 if the fighter is fighting at home, 0 if they do not
data_combined['fighter1_fight_at_home'] = (data_combined['fighter1_home_country'] == data_combined['location']) * 1
data_combined['fighter2_fight_at_home'] = (data_combined['fighter2_home_country'] == data_combined['location']) * 1
# 1 if fighter1 has a home advantage, 0.5 if its even, 0 if fighter2 has a home advantage
home_difference = data_combined['fighter1_fight_at_home'] - data_combined['fighter2_fight_at_home']
data_combined['fighter1_home_advantage'] = ((home_difference) + 1) / 2
# +
# Bouts with information of home for fighter1 or fighter2
f1_home_not_na = ~data_combined['fighter1_home_country'].isna()
f2_home_not_na = ~data_combined['fighter2_home_country'].isna()
# Bouts with information on both fighters' home country
bouts_with_both_fighters_homes = len(data_combined[f1_home_not_na][f2_home_not_na])
total_bouts = len(data_combined)
# Percentage of bouts with information on both fighters' home country
pct_rows_with_homes = bouts_with_both_fighters_homes / total_bouts * 100
print('{}% of the bouts have information for home of both fighters'.format(round( pct_rows_with_homes, 0)))
# -
# - As the data is so limited for where a fighter is from, we cannot use this data for now
# ### Remove columns that are not needed for modelling
cols_to_drop = ['location',
'fighter1_draw', 'fighter1_dob',
'fighter1_home_country', 'fighter1_fight_at_home',
'fighter2_draw', 'fighter2_dob',
'fighter2_home_country', 'fighter2_fight_at_home',
'fighter1_home_advantage',
'method_DEC', 'method_DQ', 'method_KO/TKO', 'method_SUB']
data_combined.drop(labels=cols_to_drop, axis=1, inplace=True)
show_na_cols(data_combined)
# ### Save combined dataframe to csv
data_combined.to_csv('../data/processed/combined', index=False)
| notebooks/cleaning_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import warnings
warnings.filterwarnings("ignore")
from IPython.core.display import HTML as Center
Center(""" <style>.output_png {display: table-cell; text-align: center; vertical-align: middle;}</style> """)
import cv2
import numpy as np
import skimage.io
import os
import sys
import matplotlib.pyplot as plt
import math
from scipy import ndimage
import pandas as pd
import seaborn as sns
from random import randint, uniform
from tqdm import tqdm
from collections import Counter
from itertools import permutations, product
from imblearn.under_sampling import RandomUnderSampler
import pickle
from sklearn.metrics import precision_score, recall_score, accuracy_score, confusion_matrix, f1_score
from sklearn.model_selection import train_test_split
import xgboost as xgb
from tensorflow.keras.datasets import mnist
sys.path.append('../src/inference')
from utils import *
test_data = '../data/test/img'
model_dir = '../model'
# -
# # MNIST dataset
(X_train_mnist, y_train_mnist), (X_test_mnist, y_test_mnist) = mnist.load_data()
# +
data_mnist = np.concatenate((X_train_mnist, X_test_mnist), axis=0)
data_mnist = data_mnist.reshape(data_mnist.shape[0],-1)
target_mnist = np.concatenate((y_train_mnist, y_test_mnist), axis=0)
col = np.unique(target_mnist)
digits_no = len(col)
counts_mnist = Counter(target_mnist)
plt.figure(figsize=(10,6))
plt.bar(counts_mnist.keys(), counts_mnist.values(), width = 0.8)
plt.tick_params(labelsize = 14)
plt.xticks(list(counts_mnist.keys()))
plt.xlabel("Digits",fontsize=16)
plt.ylabel("Frequency",fontsize=16)
plt.title('%s datapoints in MNIST dataset'%(target_mnist.shape[0]))
plt.show()
# -
# # Resampling of MNIST dataset to train/val/test format
X_train, X, y_train, y = train_test_split(data_mnist, target_mnist, test_size = 0.2, stratify=target_mnist)
X_val, X_test, y_val, y_test = train_test_split(X, y, test_size = 0.5, stratify = y)
# +
plt.figure(figsize=(30,10))
for idx in range(digits_no):
index = np.where(target_mnist==idx)[0][0]
img = data_mnist[index]
plt.subplot(2,5,idx+1)
plt.title('Labelled as %s'%(idx), fontsize=40)
plt.imshow(img.reshape(28,28))
plt.axis('off')
plt.tight_layout()
# -
# # Digit classification using xgboost
# +
# Hyperparameters of xgb classifier
params_xgb = {'num_class': digits_no,
'learning_rate': 0.05,
'objective': 'multi:softprob',
'n_estimators': 1000}
# XGBoost setting and training
cls = xgb.XGBClassifier(**params_xgb).fit(X_train,
y_train,
early_stopping_rounds = 5,
eval_metric = ['merror','mlogloss'],
eval_set = [(X_train, y_train), (X_val, y_val)],
verbose = False)
pickle.dump(cls, open(os.path.join(model_dir,'xgb_mnist.pkl'), "wb"))
# -
# # Training metrics
# +
# Training metrics
results = cls.evals_result()
epochs = len(results['validation_0']['mlogloss'])
x_axis = range(0, epochs)
# Plot log loss
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(20,5))
ax1.plot(x_axis, results['validation_0']['mlogloss'], label='Train')
ax1.plot(x_axis, results['validation_1']['mlogloss'], label='Val')
ax1.legend(fontsize=15)
ax1.set_xlabel('epoch', fontsize=15)
ax1.set_ylabel('Log Loss', fontsize=15)
# Plot classification error
ax2.plot(x_axis, [1-x for x in results['validation_0']['merror']], label='Train')
ax2.plot(x_axis, [1-x for x in results['validation_1']['merror']], label='Val')
ax2.legend(fontsize=15)
ax2.set_xlabel('epoch', fontsize=15)
ax2.set_ylabel('Accuracy', fontsize=15)
plt.tight_layout()
# -
# # Evaluation
# +
preds = cls.predict(X_test)
print("Precision = {:.4f}".format(precision_score(np.squeeze(y_test), preds, average='macro')))
print("Recall = {:.4f}".format(recall_score(np.squeeze(y_test), preds, average='macro')))
print("F1-score = {:.4f}".format(f1_score(np.squeeze(y_test), preds, average='macro')))
print("Accuracy = {:.4f}".format(accuracy_score(np.squeeze(y_test), preds)))
# +
res = confusion_matrix(y_test, preds)
confusion = pd.DataFrame(res, columns=col, index = col)
confusion.index.name, confusion.columns.name = 'Actual', 'Predicted'
plt.figure(figsize = (10,8))
sns.heatmap(confusion, annot=True, cmap='Blues')
plt.title('Test accuracy: %.4f' % (accuracy_score(np.squeeze(y_test), preds)))
plt.show()
# -
# # Test over sudoku tile data
# +
img_sudoku = cv2.imread(os.path.join(test_data,'clear.jpg'))[:, :, [2, 1, 0]]
gray_sudoku = cv2.cvtColor(img_sudoku, cv2.COLOR_RGB2GRAY)
centroid, (grid, vis), bbox = get_res(img_sudoku)
fig = plt.figure(figsize=(20,10))
plt.subplot(1,3,1)
plt.imshow(img_sudoku)
plt.title('Input image')
plt.axis('off')
plt.subplot(1,3,2)
plt.imshow(grid)
plt.title('Grid retrieval')
plt.axis('off')
plt.subplot(1,3,3)
plt.imshow(vis)
plt.title('Centroid retrieval')
plt.axis('off')
plt.tight_layout()
plt.show()
# +
plt.figure(figsize=(20,20))
for idx, bb in enumerate(bbox):
x0, x1, y0, y1 = bb
inv = np.uint8(np.invert(gray_sudoku[y0:y1,x0:x1]))
img_resize = cv2.resize(inv, (28,28), interpolation=cv2.INTER_LINEAR).reshape(1,-1)
l = None if np.max(inv) < 150 else cls.predict(img_resize)[0]
plt.subplot(9,9,idx+1)
plt.imshow(inv)
plt.axis('off')
plt.title('Pred = %s'%(l),fontsize=20)
plt.tight_layout()
# -
# It appears that training on MNIST dataset does not generalize well over computer created digits and performs poorly over sudoku data.
# ## Data creation
def text_on_img(text, size=3, center=False):
font = ['DejaVu Sans','Arial','sans-serif', 'fantasy', 'monospace', 'sans', 'sans serif', 'serif']
fig, ax = plt.subplots()
ax.text(0+uniform(0.0, 0.6 if not center else 0.25),
0+uniform(0.0, 0.6 if not center else 0.25),
'%s'%(text),
fontsize=250+randint(0,50),
fontweight=100+randint(-50,50),
fontname=font[randint(0,len(font)-1)])
plt.axis('off')
fig.canvas.draw()
data = np.frombuffer(fig.canvas.tostring_rgb(), dtype=np.uint8)
w, h = fig.canvas.get_width_height()
plt.close(fig)
vis = np.invert(data.reshape((int(h), int(w), -1))[:, :, [2, 1, 0]])
vis = cv2.cvtColor(vis,cv2.COLOR_RGB2GRAY)
vis = cv2.blur(vis,(9,9))
vis = cv2.resize(vis,(28,28))
return vis.reshape(-1,784)
# +
s = 6000 # Number of datapoint to be created per digit
balance = [(idx,s) for idx in range(digits_no)] # Digits-volume dictionnary
data_editor = np.zeros((digits_no*s,784)) # Flatten image 28x28 pixels
target_editor = np.zeros((digits_no*s,1)) # Label data
k=0
for (key, val) in balance:
print('Creating training data for digit %s'%(key))
for j in tqdm(range(val), position = 0):
data_editor[k,:] = text_on_img(str(key))
target_editor[k,:] = np.array(key)
k+=1
target_editor = np.squeeze(target_editor)
# -
# # Data preparation & visualization
# As creating twice the amount of data for being able to recognize both handritten and editor created digits, I decided to reduce the number of data taken from MNIST as high accuracy can be expected with fewer number of datapoints.
#
# In order to make my dataset totally balanced, I downsampled each class of MNIST dataset to 6.000 datapoints with their corresponding image and combine it with artificailly created images.
# +
undersample = RandomUnderSampler({i:s for i in list(range(digits_no))}, random_state=0)
data_mnist, target_mnist = undersample.fit_resample(data_mnist, target_mnist)
data = np.concatenate((data_editor, data_mnist), axis=0)
target = np.concatenate((target_editor, target_mnist + 10), axis=0) # +10 → Stratify mnist/editor digits categories
X_train, X, y_train, y = train_test_split(data, target, test_size=0.2, random_state=0, stratify=target)
X_val, X_test, y_val, y_test = train_test_split(X, y, test_size=0.5, random_state=0, stratify=y)
# Stratification from both handwritten and editor based should be fixed so training data contains
# equal number of handwritten data and text-editor generated data
y_train[np.where(y_train > 9)] -= 10
y_val[np.where(y_val > 9)] -= 10
y_test[np.where(y_test > 9)] -= 10
# +
editor = list(Counter(target_editor).values())
mnist = list(Counter(target_mnist).values())
plt.figure(figsize=(10,6))
mnist_plot = plt.bar(np.arange(digits_no), mnist, 0.35)
editor_plot = plt.bar(np.arange(digits_no), editor, 0.35, bottom=mnist)
plt.tick_params(labelsize = 14)
plt.xticks(list(range(digits_no)))
plt.xlabel("Digits", fontsize=16)
plt.ylabel('Occurence', fontsize=16)
plt.legend((mnist_plot[0], editor_plot[0]), ('MNIST', 'Editor'))
plt.title('%s datapoints'%(target.shape[0]))
plt.show()
# -
# # Train
# XGBoost setting and training
cls_all = xgb.XGBClassifier(**params_xgb).fit(X_train,
y_train,
early_stopping_rounds = 5,
eval_metric = ['merror','mlogloss'],
eval_set = [(X_train, y_train), (X_val, y_val)],
verbose = False)
pickle.dump(cls_all, open(os.path.join(model_dir,'xgb_mnist_and_editor.pkl'), "wb"))
# # Training metrics
# +
# Training metrics
results = cls_all.evals_result()
epochs = len(results['validation_0']['mlogloss'])
x_axis = range(0, epochs)
# Plot log loss
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(20,5))
ax1.plot(x_axis, results['validation_0']['mlogloss'], label='Train')
ax1.plot(x_axis, results['validation_1']['mlogloss'], label='Val')
ax1.legend(fontsize=15)
ax1.set_xlabel('epoch', fontsize=15)
ax1.set_ylabel('Log Loss', fontsize=15)
# Plot classification error
ax2.plot(x_axis, [1-x for x in results['validation_0']['merror']], label='Train')
ax2.plot(x_axis, [1-x for x in results['validation_1']['merror']], label='Val')
ax2.legend(fontsize=15)
ax2.set_xlabel('epoch', fontsize=15)
ax2.set_ylabel('Accuracy', fontsize=15)
plt.tight_layout()
# -
# ## Evaluation
# +
preds = cls_all.predict(X_test)
print("Precision = {:.4f}".format(precision_score(np.squeeze(y_test), preds, average='macro')))
print("Recall = {:.4f}".format(recall_score(np.squeeze(y_test), preds, average='macro')))
print("F1-score = {:.4f}".format(f1_score(np.squeeze(y_test), preds, average='macro')))
print("Accuracy = {:.4f}".format(accuracy_score(np.squeeze(y_test), preds)))
# +
res = confusion_matrix(y_test, preds)
confusion = pd.DataFrame(res, columns=col, index = col)
confusion.index.name, confusion.columns.name = 'Actual', 'Predicted'
plt.figure(figsize = (10,8))
sns.heatmap(confusion, annot=True, cmap='Blues')
plt.title('Test accuracy: %.4f' % (accuracy_score(np.squeeze(y_test), preds)))
# -
# # Test batch data
# +
display_col = 5
plt.figure(figsize=(20,30))
for digit, idx in product(range(digits_no), range(display_col)):
img = X_test[np.where(y_test==digit)[0][idx]]
pred = int(cls_all.predict(img.reshape(1,-1))[0])
plt.subplot(digits_no + 1, display_col, digit*display_col + idx + 1)
plt.title('pred = %s'%(pred), fontsize=30)
plt.imshow(img.reshape(28,28))
plt.axis('off')
plt.tight_layout()
# -
# # Visualization over sudoku data
# +
plt.figure(figsize=(20,20))
for idx, bb in enumerate(bbox):
x0, x1, y0, y1 = bb
inv = np.uint8(np.invert(gray_sudoku[y0:y1,x0:x1]))
img_resize = cv2.resize(inv, (28,28), interpolation=cv2.INTER_LINEAR)
l = None if np.max(inv) < 150 else int(cls_all.predict(img_resize.reshape(1,-1))[0])
plt.subplot(9,9,idx+1)
plt.imshow(img_resize)
plt.axis('off')
plt.title('Pred = %s'%(l),fontsize=20)
plt.tight_layout()
# -
| notebook/digits_recognition_ML.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Semantic file formats
# ## The dream of a semantic web
# So how can we fulfill the dream of a file-format which is **self-documenting**:
# universally unambiguous and interpretable?
# (Of course, it might not be true, but we don't have capacity to discuss how to model reliability
# and contested testimony.)
# By using URIs to define a controlled vocabulary, we can be unambiguous.
#
# But the number of different concepts to be labelled is huge: so we need a **distributed** solution:
# a global structure of people defining ontologies, (with methods for resolving duplications and inconsistencies.)
# Humanity has a technology that can do this: the world wide web. We've seen how many different
# actors are defining ontologies.
# We also need a shared semantic structure for our file formats. XML allows everyone to define their
# own schema. Our universal file format requires a restriction to a basic language, which allows us
# to say the things we need:
# ## The Triple
# We can then use these defined terms to specify facts, using a URI for the subject, verb, and object of our sentence.
# +
# %%writefile reaction.ttl
<http://dbpedia.org/ontology/water>
<http://purl.obolibrary.org/obo/PATO_0001681>
"18.01528"^^<http://purl.obolibrary.org/obo/UO_0000088>
.
# -
# * [Water](http://dbpedia.org/ontology/water)
# * [Molar mass](http://purl.obolibrary.org/obo/PATO_0001681)
# * [Grams per mole](http://purl.obolibrary.org/obo/UO_0000088)
# This is an unambiguous statement, consisting of a subject, a verb, and an object, each of which is either a URI or a literal value. Here, the object is a *literal* with a type.
# ## RDF file formats
# We have used the RDF **semantic** format, in its "Turtle" syntactic form:
#
# ```
# subject verb object .
# subject2 verb2 object2 .
# ```
# We can parse it:
# +
from rdflib import Graph
graph = Graph()
graph.parse("reaction.ttl", format="ttl")
len(graph) # prints 2
for statement in graph:
print(statement)
# -
# The equivalent in **RDF-XML** is:
print(graph.serialize(format='xml').decode())
# We can also use namespace prefixes in Turtle:
print(graph.serialize(format='ttl').decode())
# ## Normal forms and Triples
# How do we encode the sentence "water has two hydrogen atoms" in RDF?
# See [Defining N-ary Relations on the Semantic Web](https://www.w3.org/TR/swbp-n-aryRelations/) for the definitive story.
# I'm not going to search carefully here for existing ontologies for the relationships we need:
# later we will understand how to define these as being the same as or subclasses of concepts
# in other ontologies. That's part of the value of a distributed approach: we can define
# what we need, and because the Semantic Web tools make rigorous the concepts of `rdfs:sameAs` and `subclassOf`
# `rdfs:subclassOf` this will be OK.
#
# However, there's a problem. We can do:
# +
# %%writefile reaction.ttl
@prefix disr: <http://www.hep.ucl.ac.uk/cdt-dis/ontologies/reactions/> .
@prefix dbo: <http://dbpedia.org/ontology/> .
@prefix obo: <http://purl.obolibrary.org/obo/> .
dbo:water obo:PATO_0001681 "18.01528"^^obo:UO_0000088 ;
disr:containsElement obo:CHEBI_33260 .
# -
# * [ElementalHydrogen](http://www.ebi.ac.uk/chebi/searchId.do?chebiId=CHEBI:33260)
# We've introduced the semicolon in Turtle to say two statements about the same entity. The equivalent RDF-XML is:
graph = Graph()
graph.parse("reaction.ttl", format="ttl")
print(graph.serialize(format='xml').decode())
# However, we can't express "hasTwo" in this way without making an infinite number of properties!
#
# RDF doesn't have a concept of adverbs. Why not?
# It turns out there's a fundamental relationship between the RDF triple and a RELATION in
# the relational database model.
# * The **subject** corresponds to the relational primary key.
# * The **verb** (RDF "property") corresponds to the relational column name.
# * The **object** corresponds to the value in the corresponding column.
# We already found out that to model the relationship of atoms to molecules we needed a join table, and the
# number of atoms was metadata on the join.
# So, we need an entity type (RDF **class**) which describes an ElementInMolecule.
# Fortunately, we don't have to create a universal URI for every single relatioship, thanks to
# RDF's concept of an anonymous entity. (Uniquely defined only by its relationships.)
# Imagine if we had to make a URN for oxygen-in-water, hydrogen-in-water etc!
# +
# %%writefile reaction.ttl
@prefix disr: <http://www.hep.ucl.ac.uk/cdt-dis/ontologies/reactions/> .
@prefix dbo: <http://dbpedia.org/ontology/> .
@prefix obo: <http://purl.obolibrary.org/obo/> .
@prefix xs: <http://www.w3.org/2001/XMLSchema> .
dbo:water obo:PATO_0001681 "18.01528"^^obo:UO_0000088 ;
disr:containsElement obo:CHEBI_33260 ;
disr:hasElementQuantity [
disr:countedElement obo:CHEBI_33260 ;
disr:countOfElement "2"^^xs:integer ] .
# -
# Here we have used [ ] to indicate an anonymous entity, with no subject. We then define
# two predicates on that subject, using properties corresponding to our column names in the join table.
# Another turtle syntax for an anonymous "blank node" is this:
# +
# %%writefile reaction.ttl
@prefix disr: <http://www.hep.ucl.ac.uk/cdt-dis/ontologies/reactions/> .
@prefix dbo: <http://dbpedia.org/ontology/> .
@prefix obo: <http://purl.obolibrary.org/obo/> .
@prefix xs: <http://www.w3.org/2001/XMLSchema> .
dbo:water obo:PATO_0001681 "18.01528"^^obo:UO_0000088 ;
disr:containsElement obo:CHEBI_33260 ;
disr:hasElementQuantity _:a .
_:a disr:countedElement obo:CHEBI_33260 ;
disr:countOfElement "2"^^xs:integer .
# -
# ## Serialising to RDF
# Here's code to write our model to Turtle:
# +
# %%writefile chemistry_turtle_template.mko
@prefix disr: <http://www.hep.ucl.ac.uk/cdt-dis/ontologies/reactions/> .
@prefix obo: <http://purl.obolibrary.org/obo/> .
@prefix xs: <http://www.w3.org/2001/XMLSchema> .
[
# %for reaction in reactions:
disr:hasReaction [
# %for molecule in reaction.reactants.molecules:
disr:hasReactant [
% for element in molecule.elements:
disr:hasElementQuantity [
disr:countedElement [
a obo:CHEBI_33259;
disr:symbol "${element.symbol}"^^xs:string
] ;
disr:countOfElement "${molecule.elements[element]}"^^xs:integer
];
% endfor
a obo:CHEBI_23367
] ;
# %endfor
# %for molecule in reaction.products.molecules:
disr:hasProduct [
% for element in molecule.elements:
disr:hasElementQuantity [
disr:countedElement [
a obo:CHEBI_33259;
disr:symbol "${element.symbol}"^^xs:string
] ;
disr:countOfElement "${molecule.elements[element]}"^^xs:integer
] ;
% endfor
a obo:CHEBI_23367
] ;
# %endfor
a disr:reaction
] ;
# %endfor
a disr:system
].
# -
# "a" in Turtle is an always available abbreviation for http://www.w3.org/1999/02/22-rdf-syntax-ns#type
#
# We've also used:
#
# * [Molecular entity](http://www.ebi.ac.uk/chebi/searchId.do?chebiId=CHEBI%3A23367)
# * [Elemental molecular entity](http://www.ebi.ac.uk/chebi/searchId.do?chebiId=CHEBI%3A33259)
# I've skipped serialising the stoichiometries : to do that correctly I also need to create a
# relationship class for molecule-in-reaction.
# And we've not attempted to relate our elements to their formal definitions, since our model
# isn't recording this at the moment. We could add this statement later.
# +
import mako
from parsereactions import parser
from IPython.display import display, Math
system=parser.parse(open('system.tex').read())
display(Math(str(system)))
# +
from mako.template import Template
mytemplate = Template(filename='chemistry_turtle_template.mko')
with open('system.ttl','w') as ttlfile:
ttlfile.write((mytemplate.render( **vars(system))))
# -
# !cat system.ttl
graph = Graph()
graph.parse("system.ttl", format="ttl")
# We can see why the group of triples is called a *graph*: each node is an entity and each arc a property relating entities.
# Note that this format is very very verbose. It is **not** designed to be a nice human-readable format.
#
# Instead, the purpose is to maximise the capability of machines to reason with found data.
# ## Formalising our ontology: RDFS
# Our http://www.hep.ucl.ac.uk/cdt-dis/ontologies/reactions/ namespace now contains the following properties:
# * disr:hasReaction
# * disr:hasReactant
# * disr:hasProduct
# * disr:containsElement
# * disr:countedElement
# * disr:hasElementQuantity
# * disr:countOfElement
# * disr:symbol
# And two classes:
# * disr:system
# * disr:reaction
# We would now like to find a way to formally specify some of the relationships between these.
#
# The **type** (`http://www.w3.org/1999/02/22-rdf-syntax-ns#type` or `a`) of the subject of hasReaction
# must be `disr:system`.
#
#
# [RDFS](https://www.w3.org/TR/rdf-schema/) will allow us to specify which URNs define classes and which properties,
# and the domain and range (valid subjects and objects) of our properties.
# For example:
# +
# %%writefile cdt_dis_ontology.ttl
@prefix disr: <http://www.hep.ucl.ac.uk/cdt-dis/ontologies/reactions/> .
@prefix obo: <http://purl.obolibrary.org/obo/> .
@prefix xs: <http://www.w3.org/2001/XMLSchema> .
@prefix rdfs: <http://www.w3.org/2000/01/rdf-schema#> .
@prefix rdf: <http://www.w3.org/1999/02/22-rdf-syntax-ns#> .
disr:system a rdfs:Class .
disr:reaction a rdfs:Class .
disr:hasReaction a rdf:Property .
disr:hasReaction rdfs:domain disr:system .
disr:hasReaction rdfs:range disr:reaction .
# -
# This will allow us to make our file format briefer: given this schema, if
#
# `_:a hasReaction _:b`
#
# then we can **infer** that
#
# `_:a a disr:system .
# _:b a disr:reaction .`
#
# without explicitly stating it.
#
# Obviously there's a lot more to do to define our other classes, including defining a class for our anonymous element-in-molecule nodes.
# This can get very interesting:
# +
# %%writefile cdt_dis_ontology.ttl
@prefix disr: <http://www.hep.ucl.ac.uk/cdt-dis/ontologies/reactions/> .
@prefix obo: <http://purl.obolibrary.org/obo/> .
@prefix xs: <http://www.w3.org/2001/XMLSchema> .
@prefix rdfs: <http://www.w3.org/2000/01/rdf-schema#> .
@prefix rdf: <http://www.w3.org/1999/02/22-rdf-syntax-ns#> .
disr:system a rdfs:Class .
disr:reaction a rdfs:Class .
disr:hasReaction a rdf:Property .
disr:hasReaction rdfs:domain disr:system .
disr:hasReaction rdfs:range disr:reaction .
disr:hasParticipant a rdf:Property .
disr:hasReactant rdfs:subPropertyOf disr:hasParticipant .
disr:hasProduct rdfs:subPropertyOf disr:hasParticipant
# -
# [OWL](https://www.w3.org/TR/owl-ref/) extends RDFS even further.
# Inferring additional rules from existing rules and schema is very powerful: an interesting branch of AI. (Unfortunately the [python tool](https://github.com/RDFLib/OWL-RL) for doing this automatically is currently not updated to python 3 so I'm not going to demo it. Instead, we'll see in a moment how to apply inferences to our graph to introduce new properties.)
# ## SPARQL
# So, once I've got a bunch of triples, how do I learn anything at all from them? The language
# is so verbose it seems useless!
# SPARQL is a very powerful language for asking questions of knowledge bases defined in RDF triples:
# +
results=graph.query(
"""SELECT DISTINCT ?asymbol ?bsymbol
WHERE {
?molecule disr:hasElementQuantity ?a .
?a disr:countedElement ?elementa .
?elementa disr:symbol ?asymbol .
?molecule disr:hasElementQuantity ?b .
?b disr:countedElement ?elementb .
?elementb disr:symbol ?bsymbol
}""")
for row in results:
print("Elements %s and %s are found in the same molecule" % row)
# -
# We can see how this works: you make a number of statements in triple-form, but with some
# quantities as dummy-variables. SPARQL finds all possible subgraphs of the triple graph which
# are compatible with the statements in your query.
#
#
# We can also use SPARQL to specify **inference rules**:
graph.update(
"""INSERT { ?elementa disr:inMoleculeWith ?elementb }
WHERE {
?molecule disr:hasElementQuantity ?a .
?a disr:countedElement ?elementa .
?elementa disr:symbol ?asymbol .
?molecule disr:hasElementQuantity ?b .
?b disr:countedElement ?elementb .
?elementb disr:symbol ?bsymbol
}"""
)
# +
graph.query("""
SELECT DISTINCT ?asymbol ?bsymbol
WHERE {
?moleculea disr:inMoleculeWith ?moleculeb .
?elementa disr:symbol ?asymbol .
?elementb disr:symbol ?bsymbol
}""")
for row in results:
print("Elements %s and %s are found in the same molecule" % row)
# -
# Exercise for reader: express "If x is the subject of a hasReaction relationship, then x must be a system"
# in SPARQL.
# Exercise for reader: search for a SPARQL endpoint knowledge base in your domain.
#
# Connect to it using [Python RDFLib's SPARQL endpoint wrapper](https://github.com/RDFLib/sparqlwrapper) and ask it a question.
| ch09fileformats/12SemanticModels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Enums and Dataclasses
#
# Often we would like to be able to refer to a fixed set of values as constants.
#
# Consider, for example, a program that works with the points of the compass. We could refer to them as string values, but this would leave us exposed to spelling errors, differences of convention (should `North West` be hyphenated? Could it be referred to as `NW`?).
#
# Instead let us define the points of the compass and see how we might use them when writing some classes.
# +
from enum import Enum
from dataclasses import dataclass, field
from typing import List
class Direction(Enum):
NORTH = 1
NORTH_WEST = 2
NORTH_EAST = 3
SOUTH = 4
SOUTH_WEST = 5
SOUTH_EAST = 6
EAST = 7
WEST = 8
@dataclass
class NavigationInstruction:
direction: Direction
distance: int
def print_instruction(self):
print(f'Move {self.distance}m {self.direction.name}')
@dataclass
class Path:
instructions: List[NavigationInstruction] = field(default_factory=list)
def print_all(self):
for instruction in self.instructions:
instruction.print_instruction()
# -
path = Path(instructions=[NavigationInstruction(Direction.NORTH, 50),
NavigationInstruction(Direction.EAST, 100),
NavigationInstruction(Direction.NORTH_WEST, 75)])
path.print_all()
# ## Some notes
#
# Python will still allow us to pass any values in when constructing `NavigationInstruction`. This is because Python doesn't do any type checking at run time. We can use `MyPy` to run static analysis of our code and ensure that it we aren't passing invalid values in (this is definitely a good thing to do) but the greater value comes from communicating our intent clearly to our future selves and collaborators.
#
# When using a `list` or a `dict` in a dataclass we need to provide a value to the `default_factory`
# ## Further Reading
#
# [Enums](https://docs.python.org/3.7/library/enum.html)
#
# [Dataclasses](https://docs.python.org/3.7/library/dataclasses.html)
| notebooks/05_Enums_and_Dataclasses.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
from bs4 import BeautifulSoup
import re
import json
import pickle
import datetime
def name_formatter(name):
name = re.sub("_", " ", name).strip().title()
return(name)
# need to do pip3 install lxml and then reload the jupyter file
def ar_lookup(name):
"""
Convert names from English to Arabic using European Media Monitors and
code modified from <NAME>.
"""
name = name_formatter(name)
base_url = "http://emm.newsexplorer.eu/NewsExplorer/search/en/entities?query="
url = base_url + name
try:
page = requests.get(url)
soup = BeautifulSoup(page.content, "lxml")
name_url = soup.find("p", {"class" : "center_headline"}).find("a")['href']
except Exception as e:
#print("Couldn't get page of results back: ", e)
return []
try:
base = "http://emm.newsexplorer.eu/NewsExplorer/search/en/"
name_url = base + name_url
name_page = requests.get(name_url)
soup = BeautifulSoup(name_page.content, "lxml")
# check to make sure in list???? Take in alt names???/petrarch2/petrarch2/data/dictionaries
names = soup.find("td", {"colspan" : "1"}).find_all("p")
names = [i.text for i in names][1:]
names_en = [i for i in names if re.search("\(.*?Eu|\(.*?en", i)]
names_en = [re.sub("\s+?\(.+?\)", "", name) for name in names_en]
#print("Found match. Matched English name: ", names_en[0])
names_ar = [i for i in names if re.search("\(.*?ar", i)]
names_ar = [re.sub("\s+?\(.+?\)", "", name) for name in names_ar]
return names_ar
except Exception:
traceback.print_exc()
return []
ar_lookup("ANGELA_MERKEL")
# +
def clean_line(line):
# Take out extra space, underscores, comments, etc.
cleaned = re.sub("_* .+", "", line).strip()
cleaned = re.sub("_$", "", cleaned, flags=re.MULTILINE)
return cleaned
def ingest_dictionary(dict_path):
"""
Read in the country (or other) actor dictionaries.
"""
with open(dict_path) as f:
country_file = f.read()
split_file = country_file.split("\n")
dict_dict = []
key_name = ""
alt_names = []
roles = []
for line in split_file:
if not line:
pass
elif line[0] == "#":
pass
elif re.match("[A-Z]", line[0]):
# handle the previous
entry = {"actor_en" : key_name,
"alt_names_en" : alt_names,
"roles" : roles}
dict_dict.append(entry)
# zero everything out
alt_names = []
roles = []
# make new key name
key_name = clean_line(line)
# check to see if the role is built in
if bool(re.search("\[[A-Z]{3}\]", line)):
roles = re.findall("\[(.+?)\]", line)
elif line[0] == "+":
cleaned = clean_line(line[1:])
alt_names.append(cleaned)
elif re.match("\s", line):
roles.append(line.strip())
return dict_dict
dp = "./Phoenix.Countries.actors.txt"
dict_dict = ingest_dictionary(dp)
# -
len(dict_dict)
dict_dict[1123]
ar_lookup("Mohammad_Najibullah")
#dict_dict here is just one object, not a list of object
def eng_to_ar(dict_dict):
"""
Update an English language dictionary entry with Arabic names
"""
ar_names = ar_lookup(dict_dict["actor_en"])
if not ar_names:
#print("No ar name match found for "+ str(dict_dict["actor_en"]))
raise Exception(dict_dict["actor_en"])
#return dict_dict
dict_dict['actor_ar'] = ar_names[0]
if len(ar_names) > 1:
dict_dict['alt_names_ar'] = ar_names[1:]
return dict_dict
def grabAllTHeEnglishNamesThatNoArName(dict_dict):
noFindList=[]
for item in dict_dict:
try:
eng_to_ar(item)
except Exception as e:
noFindList.append(e)
#print(e)
# %%time
notfind=grabAllTHeEnglishNamesThatNoArName(dict_dict)
#then dump the data to pickle
try:
with open("noFindWord", 'wb') as f:
pickle.dump(nofind, f, pickle.HIGHEST_PROTOCOL)
except:
print("failed to save the result to disk")
pass
def hack_wiki(eng_name):
base_url="https://en.wikipedia.org/wiki/"+eng_name
try:
page=requests.get(base_url)
soup=BeautifulSoup(page.content,"lxml")
name=soup.find(id="firstHeading").contents
ar_url=soup.find("li",{"class":"interwiki-ar"}).find("a")['href']
#print(ar_url)
ar_page=requests.get(ar_url)
ar_soup=BeautifulSoup(ar_page.content,"lxml")
ar_name=ar_soup.find(id="firstHeading").contents
print(ar_name)
#print("name "+name+" url: "+str(ar_url))
except Exception as e:
print(e)
def eng_to_ar(dict_dict):
"""
Update an English language dictionary entry with Arabic names
"""
ar_names = ar_lookup(dict_dict["actor_en"])
if not ar_names:
print("No ar name match found.")
return dict_dict
dict_dict['actor_ar'] = ar_names[0]
if len(ar_names) > 1:
dict_dict['alt_names_ar'] = ar_names[1:]
return dict_dict
test=eng_to_ar(dict_dict[7777])
dic=pickle.load(open('countrycode.pkl', 'rb'))
countrycode=[key for key,value in dic.items()]
# #Format stored in our db. { "_id" : ObjectId("5772026ca78a30ce5acbec43"), "sentenceId" : "5771f20f0dfcd69f645102fa", "word" : "President Obama", "countryCode" : "USA", "firstRoleCode" : "GOV", "secondRoleCode" : "LEG", "dateStart" : "Sun Jun 01 2008 00:00:00 GMT-0500 (CDT)", "dateEnd" : "Wed Jun 01 2016 00:00:00 GMT-0500 (CDT)", "confidenceFlag" : true, "userId" : "577201cea78a30ce5acbec41", "userName" : "guest", "taggingTime" : ISODate("2016-06-28T04:51:56.092Z"), "__v" : 0 }
test
haha='[IRQELI 620101-030901]'
hahaha=haha.split(" ")
hahaha[1][0]
from pymongo import MongoClient
client=MongoClient()
client=MongoClient('mongodb://portland.cs.ou.edu:23755/')
db=client['lexisnexis']
db.authenticate('boomer', 'burritos_for_breakfast')
secondroles=db.secondroles
firstroles=db.agents
secondrolelist=[]
for item in secondroles.find():
secondrolelist.append(item['id'])
firstrolelist=[]
for item in firstroles.find():
firstrolelist.append(item['id'])
def transferStringToDateTime(inputtime):
if(str(inputtime)!=''):
dd=datetime.datetime.strptime(inputtime,'%y%m%d').date()
#this is for 62 return 1962 not 2062:
if dd.year>2017:
new=dd.replace(year=dd.year-100)
return new
else:
return dd
else:
return ''
transferStringToDateTime("620101")
#if it is first role it will return 1, if it is in secondRole list it will reture 2, and it it does not exist in both it return 3
def checkBelongToWhichRole(code):
if code in firstrolelist:
return 1;
elif code in secondrolelist:
return 2;
else:
return 3;
def checkIfCountrycodeExist(code):
if code in countrycode:
return 1
else:
return 0
# [AFGGOVMIL >050101] this format of stuff needs to be handled
strtest="IRQELI 620101-030901"
strtest.split(" ")
len("yan")
strtest[1:4]
test
# +
#extract the roles and parse the string of each role to get the role format we like for our fajita website
def extract_actor_roles(dict):
roles=[]
#it could be ['actor_ar'] is not written since we did not find the translation
if 'actor_ar' in dict:
wordlist=[]
wordlist.append(dict['actor_ar'])
if 'alt_names_ar' in dict:
for name in dict['alt_names_ar']:
wordlist.append(name)
for word in wordlist:
for item in dict['roles']:
temp=[]
splitresults=item.split(" ")
#if it has both firstRole and secondRole "[" will be include in teh str
len1=len(splitresults[0])
country=''
if(len1==4):
country=splitresults[0][1:4]
role1=''
role2=''
elif(len1==7):
country=splitresults[0][1:4]
role1=splitresults[0][4:7]
role2=''
elif(len1==10):
country=splitresults[0][1:4]
role1=splitresults[0][4:7]
role2=splitresults[0][7:10]
temp.append(country)
temp.append(checkIfCountrycodeExist(country))
temp.append(role1)
temp.append(checkBelongToWhichRole(role1))
temp.append(role2)
temp.append(checkBelongToWhichRole(role2))
if(len(splitresults)>1):
timerange=splitresults[1]
if(timerange[0]!=">" and timerange[0]!="<"):
temp.append(transferStringToDateTime(timerange[0:6]))
temp.append("startdate")
temp.append(transferStringToDateTime(timerange[7:13]))
temp.append("enddate")
elif(timerange[0]==">"):
temp.append(transferStringToDateTime(timerange[1:7]))
temp.append("startdate")
temp.append("")
temp.append("enddate")
elif(timerange[0]==">"):
temp.append("")
temp.append("startdate")
temp.append(transferStringToDateTime(timerange[1:7]))
temp.append("enddate")
else:
temp.append("")
temp.append("startdate")
temp.append("")
temp.append("enddate")
temp.append(word)
roles.append(temp)
return roles
#ok, the format will be
#["contry",'1',"role1","1","role2","2","starttime","something","endtime","something","word"]
# -
midresult=extract_actor_roles(test)
#generate a list of json object for each element in the list
def generateFajitaObjectsList(midresult):
finalResult=[]
for item in midresult:
tempjson={}
#insert in all the deault stuff first
#should be false, the flag is false means we are confident about the word being tagged in fajita.
tempjson["confidenceFlag"]=False
tempjson["sentenceId"]="10000"
tempjson["userName"]="existEnglishDictionary"
tempjson["userId"]="existEnglishDictionary"
tempjson["countryCode"]=item[0]
tempjson["firstRoleCode"]=""
tempjson["secondRoleCode"]=""
tempjson["taggingTime"]= datetime.datetime.now().strftime('%Y-%m-%d')
tempjson["dateStart"]=str(item[6])
tempjson["dateEnd"]=str(item[7])
tempjson["word"]=item[10]
if(item[3]==1):
tempjson["firstRoleCode"]=item[2]
elif(item[3]==2):
tempjson["secondRoleCode"]=item[2]
else:
pass
if(item[5]==1):
tempjson["firstRoleCode"]=item[4]
elif(item[5]==2):
tempjson["secondRoleCode"]=item[4]
else:
pass
finalResult.append(tempjson)
return finalResult
final=generateFajitaObjectsList(midresult)
final
# #now need to insert the value back to the database
for item in final:
db.sourcedictionaries.insert_one(
{'confidenceFlag': True,
'countryCode': 'FG',
'dateEnd': '',
'dateStart': '',
'firstRoleCode': '',
'secondRoleCode': '',
'sentenceId': '10000',
'taggingTime': '2017-06-24',
'userId': 'existEnglishDictionary',
'userName': 'existEnglishDictionary',
'word': 'حامد كرزاي'}
)
# +
#filter out the items that does not have roles defined in there:
def iterativeInsertAll(dict_dict):
insertRecordCount=0
insertDistinctPersonCount=0
for item in dict_dict:
try:
#print(insertRecordCount)
temptest=eng_to_ar(item)
tempmidresult=extract_actor_roles(temptest)
finalresult=generateFajitaObjectsList(tempmidresult)
for insertrocord in finalresult:
try:
db.sourcedictionaries.insert_one(insertrocord)
insertRecordCount=insertRecordCount+1
except:
pass
insertDistinctPersonCount=insertDistinctPersonCount+1
except Exception as e:
print(e)
pass
return {"recordinsert":insertRecordCount,"distinctPersonCount":insertDistinctPersonCount}
# -
# [AFGGOVMIL >050101]
# fromat like this need to handled,
# maybe have two actors and have the time format with > or < there.
# %%time
totalinsert=iterativeInsertAll(dict_dict)
totalinsert["recordinsert"]
totalinsert["distinctPersonCount"]
| otherHelperCode/english_to_arabic_dictionary/.ipynb_checkpoints/Actors_to_Arabic _extension-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--BOOK_INFORMATION-->
# <img align="left" style="padding-right:10px;" src="figures/PDSH-cover-small.png">
# *This notebook contains an excerpt from the [Python Data Science Handbook](http://shop.oreilly.com/product/0636920034919.do) by <NAME>; the content is available [on GitHub](https://github.com/jakevdp/PythonDataScienceHandbook).*
#
# *The text is released under the [CC-BY-NC-ND license](https://creativecommons.org/licenses/by-nc-nd/3.0/us/legalcode), and code is released under the [MIT license](https://opensource.org/licenses/MIT). If you find this content useful, please consider supporting the work by [buying the book](http://shop.oreilly.com/product/0636920034919.do)!*
# <!--NAVIGATION-->
# < [In Depth: k-Means Clustering](05.11-K-Means.ipynb) | [Contents](Index.ipynb) | [In-Depth: Kernel Density Estimation](05.13-Kernel-Density-Estimation.ipynb) >
# # In Depth: Gaussian Mixture Models
# The *k*-means clustering model explored in the previous section is simple and relatively easy to understand, but its simplicity leads to practical challenges in its application.
# In particular, the non-probabilistic nature of *k*-means and its use of simple distance-from-cluster-center to assign cluster membership leads to poor performance for many real-world situations.
# In this section we will take a look at Gaussian mixture models (GMMs), which can be viewed as an extension of the ideas behind *k*-means, but can also be a powerful tool for estimation beyond simple clustering.
#
# We begin with the standard imports:
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns; sns.set()
import numpy as np
# ## Motivating GMM: Weaknesses of k-Means
#
# Let's take a look at some of the weaknesses of *k*-means and think about how we might improve the cluster model.
# As we saw in the previous section, given simple, well-separated data, *k*-means finds suitable clustering results.
#
# For example, if we have simple blobs of data, the *k*-means algorithm can quickly label those clusters in a way that closely matches what we might do by eye:
# Generate some data
from sklearn.datasets.samples_generator import make_blobs
X, y_true = make_blobs(n_samples=400, centers=4,
cluster_std=0.60, random_state=0)
X = X[:, ::-1] # flip axes for better plotting
# Plot the data with K Means Labels
from sklearn.cluster import KMeans
kmeans = KMeans(4, random_state=0)
labels = kmeans.fit(X).predict(X)
plt.scatter(X[:, 0], X[:, 1], c=labels, s=40, cmap='viridis');
# From an intuitive standpoint, we might expect that the clustering assignment for some points is more certain than others: for example, there appears to be a very slight overlap between the two middle clusters, such that we might not have complete confidence in the cluster assigment of points between them.
# Unfortunately, the *k*-means model has no intrinsic measure of probability or uncertainty of cluster assignments (although it may be possible to use a bootstrap approach to estimate this uncertainty).
# For this, we must think about generalizing the model.
#
# One way to think about the *k*-means model is that it places a circle (or, in higher dimensions, a hyper-sphere) at the center of each cluster, with a radius defined by the most distant point in the cluster.
# This radius acts as a hard cutoff for cluster assignment within the training set: any point outside this circle is not considered a member of the cluster.
# We can visualize this cluster model with the following function:
# +
from sklearn.cluster import KMeans
from scipy.spatial.distance import cdist
def plot_kmeans(kmeans, X, n_clusters=4, rseed=0, ax=None):
labels = kmeans.fit_predict(X)
# plot the input data
ax = ax or plt.gca()
ax.axis('equal')
ax.scatter(X[:, 0], X[:, 1], c=labels, s=40, cmap='viridis', zorder=2)
# plot the representation of the KMeans model
centers = kmeans.cluster_centers_
radii = [cdist(X[labels == i], [center]).max()
for i, center in enumerate(centers)]
for c, r in zip(centers, radii):
ax.add_patch(plt.Circle(c, r, fc='#CCCCCC', lw=3, alpha=0.5, zorder=1))
# -
kmeans = KMeans(n_clusters=4, random_state=0)
plot_kmeans(kmeans, X)
# An important observation for *k*-means is that these cluster models *must be circular*: *k*-means has no built-in way of accounting for oblong or elliptical clusters.
# So, for example, if we take the same data and transform it, the cluster assignments end up becoming muddled:
# +
rng = np.random.RandomState(13)
X_stretched = np.dot(X, rng.randn(2, 2))
kmeans = KMeans(n_clusters=4, random_state=0)
plot_kmeans(kmeans, X_stretched)
# -
# By eye, we recognize that these transformed clusters are non-circular, and thus circular clusters would be a poor fit.
# Nevertheless, *k*-means is not flexible enough to account for this, and tries to force-fit the data into four circular clusters.
# This results in a mixing of cluster assignments where the resulting circles overlap: see especially the bottom-right of this plot.
# One might imagine addressing this particular situation by preprocessing the data with PCA (see [In Depth: Principal Component Analysis](05.09-Principal-Component-Analysis.ipynb)), but in practice there is no guarantee that such a global operation will circularize the individual data.
#
# These two disadvantages of *k*-means—its lack of flexibility in cluster shape and lack of probabilistic cluster assignment—mean that for many datasets (especially low-dimensional datasets) it may not perform as well as you might hope.
#
# You might imagine addressing these weaknesses by generalizing the *k*-means model: for example, you could measure uncertainty in cluster assignment by comparing the distances of each point to *all* cluster centers, rather than focusing on just the closest.
# You might also imagine allowing the cluster boundaries to be ellipses rather than circles, so as to account for non-circular clusters.
# It turns out these are two essential components of a different type of clustering model, Gaussian mixture models.
# ## Generalizing E–M: Gaussian Mixture Models
#
# A Gaussian mixture model (GMM) attempts to find a mixture of multi-dimensional Gaussian probability distributions that best model any input dataset.
# In the simplest case, GMMs can be used for finding clusters in the same manner as *k*-means:
from sklearn.mixture import GMM
gmm = GMM(n_components=4).fit(X)
labels = gmm.predict(X)
plt.scatter(X[:, 0], X[:, 1], c=labels, s=40, cmap='viridis');
# But because GMM contains a probabilistic model under the hood, it is also possible to find probabilistic cluster assignments—in Scikit-Learn this is done using the ``predict_proba`` method.
# This returns a matrix of size ``[n_samples, n_clusters]`` which measures the probability that any point belongs to the given cluster:
probs = gmm.predict_proba(X)
print(probs[:5].round(3))
# We can visualize this uncertainty by, for example, making the size of each point proportional to the certainty of its prediction; looking at the following figure, we can see that it is precisely the points at the boundaries between clusters that reflect this uncertainty of cluster assignment:
size = 50 * probs.max(1) ** 2 # square emphasizes differences
plt.scatter(X[:, 0], X[:, 1], c=labels, cmap='viridis', s=size);
# Under the hood, a Gaussian mixture model is very similar to *k*-means: it uses an expectation–maximization approach which qualitatively does the following:
#
# 1. Choose starting guesses for the location and shape
#
# 2. Repeat until converged:
#
# 1. *E-step*: for each point, find weights encoding the probability of membership in each cluster
# 2. *M-step*: for each cluster, update its location, normalization, and shape based on *all* data points, making use of the weights
#
# The result of this is that each cluster is associated not with a hard-edged sphere, but with a smooth Gaussian model.
# Just as in the *k*-means expectation–maximization approach, this algorithm can sometimes miss the globally optimal solution, and thus in practice multiple random initializations are used.
#
# Let's create a function that will help us visualize the locations and shapes of the GMM clusters by drawing ellipses based on the GMM output:
# +
from matplotlib.patches import Ellipse
def draw_ellipse(position, covariance, ax=None, **kwargs):
"""Draw an ellipse with a given position and covariance"""
ax = ax or plt.gca()
# Convert covariance to principal axes
if covariance.shape == (2, 2):
U, s, Vt = np.linalg.svd(covariance)
angle = np.degrees(np.arctan2(U[1, 0], U[0, 0]))
width, height = 2 * np.sqrt(s)
else:
angle = 0
width, height = 2 * np.sqrt(covariance)
# Draw the Ellipse
for nsig in range(1, 4):
ax.add_patch(Ellipse(position, nsig * width, nsig * height,
angle, **kwargs))
def plot_gmm(gmm, X, label=True, ax=None):
ax = ax or plt.gca()
labels = gmm.fit(X).predict(X)
if label:
ax.scatter(X[:, 0], X[:, 1], c=labels, s=40, cmap='viridis', zorder=2)
else:
ax.scatter(X[:, 0], X[:, 1], s=40, zorder=2)
ax.axis('equal')
w_factor = 0.2 / gmm.weights_.max()
for pos, covar, w in zip(gmm.means_, gmm.covars_, gmm.weights_):
draw_ellipse(pos, covar, alpha=w * w_factor)
# -
# With this in place, we can take a look at what the four-component GMM gives us for our initial data:
gmm = GMM(n_components=4, random_state=42)
plot_gmm(gmm, X)
# Similarly, we can use the GMM approach to fit our stretched dataset; allowing for a full covariance the model will fit even very oblong, stretched-out clusters:
gmm = GMM(n_components=4, covariance_type='full', random_state=42)
plot_gmm(gmm, X_stretched)
# This makes clear that GMM addresses the two main practical issues with *k*-means encountered before.
# ### Choosing the covariance type
#
# If you look at the details of the preceding fits, you will see that the ``covariance_type`` option was set differently within each.
# This hyperparameter controls the degrees of freedom in the shape of each cluster; it is essential to set this carefully for any given problem.
# The default is ``covariance_type="diag"``, which means that the size of the cluster along each dimension can be set independently, with the resulting ellipse constrained to align with the axes.
# A slightly simpler and faster model is ``covariance_type="spherical"``, which constrains the shape of the cluster such that all dimensions are equal. The resulting clustering will have similar characteristics to that of *k*-means, though it is not entirely equivalent.
# A more complicated and computationally expensive model (especially as the number of dimensions grows) is to use ``covariance_type="full"``, which allows each cluster to be modeled as an ellipse with arbitrary orientation.
#
# We can see a visual representation of these three choices for a single cluster within the following figure:
# 
# [figure source in Appendix](06.00-Figure-Code.ipynb#Covariance-Type)
# ## GMM as *Density Estimation*
#
# Though GMM is often categorized as a clustering algorithm, fundamentally it is an algorithm for *density estimation*.
# That is to say, the result of a GMM fit to some data is technically not a clustering model, but a generative probabilistic model describing the distribution of the data.
#
# As an example, consider some data generated from Scikit-Learn's ``make_moons`` function, which we saw in [In Depth: K-Means Clustering](05.11-K-Means.ipynb):
from sklearn.datasets import make_moons
Xmoon, ymoon = make_moons(200, noise=.05, random_state=0)
plt.scatter(Xmoon[:, 0], Xmoon[:, 1]);
# If we try to fit this with a two-component GMM viewed as a clustering model, the results are not particularly useful:
gmm2 = GMM(n_components=2, covariance_type='full', random_state=0)
plot_gmm(gmm2, Xmoon)
# But if we instead use many more components and ignore the cluster labels, we find a fit that is much closer to the input data:
gmm16 = GMM(n_components=16, covariance_type='full', random_state=0)
plot_gmm(gmm16, Xmoon, label=False)
# Here the mixture of 16 Gaussians serves not to find separated clusters of data, but rather to model the overall *distribution* of the input data.
# This is a generative model of the distribution, meaning that the GMM gives us the recipe to generate new random data distributed similarly to our input.
# For example, here are 400 new points drawn from this 16-component GMM fit to our original data:
Xnew = gmm16.sample(400, random_state=42)
plt.scatter(Xnew[:, 0], Xnew[:, 1]);
# GMM is convenient as a flexible means of modeling an arbitrary multi-dimensional distribution of data.
# ### How many components?
#
# The fact that GMM is a generative model gives us a natural means of determining the optimal number of components for a given dataset.
# A generative model is inherently a probability distribution for the dataset, and so we can simply evaluate the *likelihood* of the data under the model, using cross-validation to avoid over-fitting.
# Another means of correcting for over-fitting is to adjust the model likelihoods using some analytic criterion such as the [Akaike information criterion (AIC)](https://en.wikipedia.org/wiki/Akaike_information_criterion) or the [Bayesian information criterion (BIC)](https://en.wikipedia.org/wiki/Bayesian_information_criterion).
# Scikit-Learn's ``GMM`` estimator actually includes built-in methods that compute both of these, and so it is very easy to operate on this approach.
#
# Let's look at the AIC and BIC as a function as the number of GMM components for our moon dataset:
# +
n_components = np.arange(1, 21)
models = [GMM(n, covariance_type='full', random_state=0).fit(Xmoon)
for n in n_components]
plt.plot(n_components, [m.bic(Xmoon) for m in models], label='BIC')
plt.plot(n_components, [m.aic(Xmoon) for m in models], label='AIC')
plt.legend(loc='best')
plt.xlabel('n_components');
# -
# The optimal number of clusters is the value that minimizes the AIC or BIC, depending on which approximation we wish to use. The AIC tells us that our choice of 16 components above was probably too many: around 8-12 components would have been a better choice.
# As is typical with this sort of problem, the BIC recommends a simpler model.
#
# Notice the important point: this choice of number of components measures how well GMM works *as a density estimator*, not how well it works *as a clustering algorithm*.
# I'd encourage you to think of GMM primarily as a density estimator, and use it for clustering only when warranted within simple datasets.
# ## Example: GMM for Generating New Data
#
# We just saw a simple example of using GMM as a generative model of data in order to create new samples from the distribution defined by the input data.
# Here we will run with this idea and generate *new handwritten digits* from the standard digits corpus that we have used before.
#
# To start with, let's load the digits data using Scikit-Learn's data tools:
from sklearn.datasets import load_digits
digits = load_digits()
digits.data.shape
# Next let's plot the first 100 of these to recall exactly what we're looking at:
def plot_digits(data):
fig, ax = plt.subplots(10, 10, figsize=(8, 8),
subplot_kw=dict(xticks=[], yticks=[]))
fig.subplots_adjust(hspace=0.05, wspace=0.05)
for i, axi in enumerate(ax.flat):
im = axi.imshow(data[i].reshape(8, 8), cmap='binary')
im.set_clim(0, 16)
plot_digits(digits.data)
# We have nearly 1,800 digits in 64 dimensions, and we can build a GMM on top of these to generate more.
# GMMs can have difficulty converging in such a high dimensional space, so we will start with an invertible dimensionality reduction algorithm on the data.
# Here we will use a straightforward PCA, asking it to preserve 99% of the variance in the projected data:
from sklearn.decomposition import PCA
pca = PCA(0.99, whiten=True)
data = pca.fit_transform(digits.data)
data.shape
# The result is 41 dimensions, a reduction of nearly 1/3 with almost no information loss.
# Given this projected data, let's use the AIC to get a gauge for the number of GMM components we should use:
n_components = np.arange(50, 210, 10)
models = [GMM(n, covariance_type='full', random_state=0)
for n in n_components]
aics = [model.fit(data).aic(data) for model in models]
plt.plot(n_components, aics);
# It appears that around 110 components minimizes the AIC; we will use this model.
# Let's quickly fit this to the data and confirm that it has converged:
gmm = GMM(110, covariance_type='full', random_state=0)
gmm.fit(data)
print(gmm.converged_)
# Now we can draw samples of 100 new points within this 41-dimensional projected space, using the GMM as a generative model:
data_new = gmm.sample(100, random_state=0)
data_new.shape
# Finally, we can use the inverse transform of the PCA object to construct the new digits:
digits_new = pca.inverse_transform(data_new)
plot_digits(digits_new)
# The results for the most part look like plausible digits from the dataset!
#
# Consider what we've done here: given a sampling of handwritten digits, we have modeled the distribution of that data in such a way that we can generate brand new samples of digits from the data: these are "handwritten digits" which do not individually appear in the original dataset, but rather capture the general features of the input data as modeled by the mixture model.
# Such a generative model of digits can prove very useful as a component of a Bayesian generative classifier, as we shall see in the next section.
# <!--NAVIGATION-->
# < [In Depth: k-Means Clustering](05.11-K-Means.ipynb) | [Contents](Index.ipynb) | [In-Depth: Kernel Density Estimation](05.13-Kernel-Density-Estimation.ipynb) >
testing complete; Gopal
| tests/ml-books/05.12-Gaussian-Mixtures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# +
from collections import defaultdict
import warnings
import logging
import gffutils
import pybedtools
import pandas as pd
import copy
import re
from gffutils.pybedtools_integration import tsses
logging.basicConfig(level=logging.INFO)
# -
gtf = '/home/cmb-panasas2/skchoudh/genomes/sacCerR64/annotation/Saccharomyces_cerevisiae.R64-1-1.91.gtf'
gtf_db = '/home/cmb-panasas2/skchoudh/genomes/sacCerR64/annotation/Saccharomyces_cerevisiae.R64-1-1.91.gtf.db'
prefix = '/home/cmb-panasas2/skchoudh/genomes/sacCerR64/annotation/Saccharomyces_cerevisiae.R64-1-1.91.gffutils'
chrsizes = '/home/cmb-panasas2/skchoudh/genomes/sacCerR64/fasta/Saccharomyces_cerevisiae.R64-1-1.dna.toplevel.sizes'
db = gffutils.create_db(gtf, dbfn=gtf_db, merge_strategy='merge',
disable_infer_transcripts=True,
disable_infer_genes = True,
force=True)
# +
def create_gene_dict(db):
'''
Store each feature line db.all_features() as a dict of dicts
'''
gene_dict = defaultdict(lambda: defaultdict(lambda: defaultdict(list)))
for line_no, feature in enumerate(db.all_features()):
gene_ids = feature.attributes['gene_id']
feature_type = feature.featuretype
if feature_type == 'gene':
if len(gene_ids)!=1:
logging.warning('Found multiple gene_ids on line {} in gtf'.format(line_no))
break
else:
gene_id = gene_ids[0]
gene_dict[gene_id]['gene'] = feature
else:
transcript_ids = feature.attributes['transcript_id']
for gene_id in gene_ids:
for transcript_id in transcript_ids:
gene_dict[gene_id][transcript_id][feature_type].append(feature)
return gene_dict
for x in db.featuretypes():
print x
# -
gene_dict = create_gene_dict(db)
# +
def get_gene_list(gene_dict):
return list(set(gene_dict.keys()))
def get_UTR_regions(gene_dict, gene_id, transcript, cds):
if len(cds)==0:
return [], []
utr5_regions = []
utr3_regions = []
utrs = gene_dict[gene_id][transcript]['UTR']
first_cds = cds[0]
last_cds = cds[-1]
for utr in utrs:
## Push all cds at once
## Sort later to remove duplicates
strand = utr.strand
if strand == '+':
if utr.stop < first_cds.start:
utr.feature_type = 'five_prime_UTR'
utr5_regions.append(utr)
elif utr.start > last_cds.stop:
utr.feature_type = 'three_prime_UTR'
utr3_regions.append(utr)
else:
raise RuntimeError('Error with cds')
elif strand == '-':
if utr.stop < first_cds.start:
utr.feature_type = 'three_prime_UTR'
utr3_regions.append(utr)
elif utr.start > last_cds.stop:
utr.feature_type = 'five_prime_UTR'
utr5_regions.append(utr)
else:
raise RuntimeError('Error with cds')
return utr5_regions, utr3_regions
def create_bed(regions, bedtype='0'):
'''Create bed from list of regions
bedtype: 0 or 1
0-Based or 1-based coordinate of the BED
'''
bedstr = ''
for region in regions:
assert len(region.attributes['gene_id']) == 1
## GTF start is 1-based, so shift by one while writing
## to 0-based BED format
if bedtype == '0':
start = region.start - 1
else:
start = region.start
bedstr += '{}\t{}\t{}\t{}\t{}\t{}\n'.format(region.chrom,
start,
region.stop,
re.sub('\.\d+', '', region.attributes['gene_id'][0]),
'.',
region.strand)
return bedstr
def rename_regions(regions, gene_id):
regions = list(regions)
if len(regions) == 0:
return []
for region in regions:
region.attributes['gene_id'] = gene_id
return regions
def merge_regions(db, regions):
if len(regions) == 0:
return []
merged = db.merge(sorted(list(regions), key=lambda x: x.start))
return merged
def merge_regions_nostrand(db, regions):
if len(regions) == 0:
return []
merged = db.merge(sorted(list(regions), key=lambda x: x.start), ignore_strand=True)
return merged
# +
gene_bed = ''
exon_bed = ''
intron_bed = ''
start_codon_bed = ''
stop_codon_bed = ''
cds_bed = ''
gene_list = []
for gene_id in get_gene_list(gene_dict):
gene_list.append(gene_dict[gene_id]['gene'])
exon_regions, intron_regions = [], []
star_codon_regions, stop_codon_regions = [], []
cds_regions = []
for feature in gene_dict[gene_id].keys():
if feature == 'gene':
continue
cds = list(gene_dict[gene_id][feature]['CDS'])
exons = list(gene_dict[gene_id][feature]['exon'])
merged_exons = merge_regions(db, exons)
introns = db.interfeatures(merged_exons)
exon_regions += exons
intron_regions += introns
cds_regions += cds
merged_exons = merge_regions(db, exon_regions)
renamed_exons = rename_regions(merged_exons, gene_id)
merged_introns = merge_regions(db, intron_regions)
renamed_introns = rename_regions(merged_introns, gene_id)
merged_cds = merge_regions(db, cds_regions)
renamed_cds = rename_regions(merged_cds, gene_id)
exon_bed += create_bed(renamed_exons)
intron_bed += create_bed(renamed_introns)
cds_bed += create_bed(renamed_cds)
gene_bed = create_bed(gene_list)
gene_bedtool = pybedtools.BedTool(gene_bed, from_string=True)
exon_bedtool = pybedtools.BedTool(exon_bed, from_string=True)
intron_bedtool = pybedtools.BedTool(intron_bed, from_string=True)
cds_bedtool = pybedtools.BedTool(cds_bed, from_string=True)
gene_bedtool.remove_invalid().sort().saveas('{}.genes.bed'.format(prefix))
exon_bedtool.remove_invalid().sort().saveas('{}.exon.bed'.format(prefix))
intron_bedtool.remove_invalid().sort().saveas('{}.intron.bed'.format(prefix))
cds_bedtool.remove_invalid().sort().saveas('{}.cds.bed'.format(prefix))
# -
| notebooks/archive/Gffutils sacCerR64.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# - week 5
#
# - remove storage corssborad
#
# - firm physical interruptible definition
#
# - how to change kw/h to m3/h
#
#
# # about dataset
# ## entsog series:
# * entsog_2019_dataset : selbst extract dataset from entsog capacity talbe 2019
# * lisa's table: same source like talbe above
# * api table: firm capacity firm+interruptable capacity
#
# ## IGG dataset:
# * IGG+norway dataset+ lisa's convert equation
# +
import pandas as pd
import re
from tqdm import tqdm
import time
from multiprocessing import Pool
import multiprocessing
original_hour = 'https://transparency.entsog.eu/api/v1/operationaldatas.csv?pointDirection={}&from=2019-10-01&to=2020-04-01&indicator=Physical%20Flow&periodType=hour&timezone=CET&limit=-1&dataset=1'
firm_interrupt_hour='https://transparency.entsog.eu/api/v1/operationalData.csv?pointDirection={}&from=2020-01-01&to=2020-10-13&indicator=Firm%20Technical,Interruptible%20Total&periodType=hour&timezone=CET&limit=-1&dataset=1'
interrupt_hour='https://transparency.entsog.eu/api/v1/operationalData.csv?pointDirection={}&from=2020-01-01&to=2020-10-13&indicator=Interruptible%20Total&periodType=hour&timezone=CET&limit=-1&dataset=1'
firm_hour='https://transparency.entsog.eu/api/v1/operationalData.csv?pointDirection={}&from=2020-01-01&to=2020-10-13&indicator=Firm%20Technical&periodType=hour&timezone=CET&limit=-1&dataset=1'
#kWh/h
# or kWh/d
#need divide 10e6
# +
# compare data from lisa
#-----------------------
compare_data = pd.read_excel('Capacities_ENTSOG_2020.xlsx')
compare_data.set_index('Unnamed: 0', inplace=True)
m=compare_data.stack().reset_index()
m.columns=['From','To','V']
# GWh/d
# +
#load data from api
#------------------
operationaldata=pd.read_csv('https://transparency.entsog.eu/api/v1/Interconnections.csv?limit=-1')
fullpointdata=pd.read_csv('https://transparency.entsog.eu/api/v1/connectionpoints.csv?limit=-1',index_col=False)
direction=pd.read_csv('https://transparency.entsog.eu/api/v1/operatorpointdirections.csv?limit=-1')
# only EU
simple_direction_dataset = direction[['id', 'pointKey',
'pointLabel', 'directionKey',
'relatedPoints', 'tsoItemIdentifier',
'tpTsoCountry', 'adjacentCountry',
'tpEURelationship', 'tpCrossBorderPointType',
'tpTsoGCVMax','virtualReverseFlow']].copy()
# clean id ,remove some strange things in string
simple_direction_dataset.id = simple_direction_dataset.id.str[1:].str.lower()
# what id looks like
# de-tso-0009itp-00047exit
# solve id not so clean problem
simple_direction_dataset.id = simple_direction_dataset.id.str.replace(
'exit', 'split')
simple_direction_dataset.id = simple_direction_dataset.id.str.replace(
'entry', 'split')
simple_direction_dataset.id = simple_direction_dataset.id.str.split(
'split').str[0]
simple_direction_dataset.id = simple_direction_dataset.id + \
simple_direction_dataset.directionKey
# limit to crossborder
p = simple_direction_dataset[simple_direction_dataset.adjacentCountry !=
simple_direction_dataset.tpTsoCountry]
#only keep physical direction flow
#virtualReverseFlow dont have nan
p=p[p.virtualReverseFlow=='No']
len(p)
# -
direction[direction.tsoItemIdentifier=='21Z0000000004839']
# +
#list of transmission point
#--------------------------
transmission_point_list=list(operationaldata[operationaldata.fromInfrastructureTypeLabel=='Transmission'].pointKey.unique())
# -
# # Further filtering about point
# ## problem only fullpointdata have type of point
# * but direction stilll have some point that not show up
# ['UGS-00273', 'ITP-00111', 'UGS-00050', 'ITP-00494', 'DIS-00003',
# 'LNG-00003', 'VTP-00030', 'ITP-00062', 'ITP-00162', 'ITP-00255',
# 'ITP-00037', 'ITP-00114', 'ITP-00147', 'ITP-00519', 'ITP-00131',
# 'ITP-00125', 'ITP-00026', 'ITP-00034', 'ITP-00075', 'ITP-00105',
# 'UGS-00292', 'UGS-00293', 'UGS-00296', 'ITP-00056', 'ITP-00073',
# 'ITP-00083', 'ITP-00081', 'ITP-00086', 'ITP-00109', 'ITP-00188',
# 'ITP-00491', 'ITP-00057', 'ITP-00068', 'ITP-00006', 'ITP-00031',
# 'ITP-00047', 'ITP-00060', 'ITP-00066', 'ITP-00069', 'ITP-00080',
# 'ITP-00108', 'ITP-00126', 'UGS-00359', 'UGS-00365', 'ITP-00211',
# 'ITP-00250', 'ITP-00251', 'ITP-00452', 'ITP-00247', 'ITP-00297',
# 'ITP-00451', 'ITP-00454', 'ITP-00053', 'ITP-00100', 'LNG-00025',
# 'LNG-00020', 'ITP-00495', 'ITP-00160', 'ITP-00161', 'ITP-00301',
# 'ITP-00302', 'UGS-00253', 'ITP-00431', 'ITP-00432', 'ITP-00433',
# 'ITP-00434', 'UGS-00182', 'LNG-00053', 'LNG-00054', 'LNG-00055',
# 'UGS-00280', 'UGS-00281', 'UGS-00282', 'UGS-00405', 'UGS-00406',
# 'UGS-00407', 'UGS-00408', 'UGS-00409', 'ITP-00496']
# not in fullpointdata list
#
# * according to statistic data ITP is mostly transmission and UGS is mostly storage
# ## solving
# * method to filter:
# - choose all points in direction, which are transmission type in fullpointdata
#
# - points that not show up in fullpointdata, according to first three code of pointkey to filter
# * how : keep ITP, LNG
# relationship between point types and first three point key
fullpointdata.reset_index(inplace=True)
fullpointdata['kurz']=fullpointdata['pointKey'].str[:3]
fullpointdata['number']=1
fullpointdata[['kurz','infrastructureLabel','number']].groupby(['infrastructureLabel','kurz']).sum()
# +
# point should in transmission_point_list or start with ITP or LNG
ITP_LNG_in_p=list(p[p.pointKey.str[:3].isin(['ITP','LNG'])].pointKey.unique())
select_list=list(set(transmission_point_list+ITP_LNG_in_p))
p_after_selected=p[p.pointKey.isin(select_list)]
print('before selected: ', len(p),'\n after selected:',len(p_after_selected))
#create geodata
p_with_geodata=p_after_selected.merge(fullpointdata[['pointKey','tpMapX','tpMapY']].drop_duplicates(),left_on='pointKey',right_on='pointKey',how='left')
#there is duplicate, remove it
p_with_geodata.drop_duplicates(inplace=True)
print('after drop duplicate:',len(p_with_geodata))
# -
#id also unique
len(p_with_geodata.id.unique())
# +
#load data and clean data
#------------------------
entsog_2019_dataset=pd.read_excel('Capacities for Transmission Capacity Map RTS008_NS - DWH_final.xlsx',sheet_name='Capacity Map',skiprows=range(11))
#clean dataset drop nan
entsog_2019_dataset.dropna(axis=1,how='all',inplace=True)
entsog_2019_dataset.dropna(axis=0,how='all',inplace=True)
#-------choose only cross border capacity
entsog_2019_dataset.reset_index(drop=True,inplace=True)
#entsog_2019_dataset[entsog_2019_dataset.Number.notna()&entsog_2019_dataset.iloc[:,1:].isna().T.all()]
#1-172 crossborder capacity with eu export to on eu
#193-238 corssborder with no eu export to eu
#[193:239]
entsog_2019_dataset=entsog_2019_dataset.iloc[list(range(1,173))+list(range(193,239))]
#only keep rows with capacity
entsog_2019_dataset=entsog_2019_dataset[entsog_2019_dataset['Technical physical capacity (GWh/d)'].notna()&entsog_2019_dataset['Technical physical capacity (GWh/d)']>0]
#fill point name
entsog_2019_dataset.Point.fillna(method='ffill',inplace=True)
#remove viural pipe
entsog_2019_dataset=entsog_2019_dataset[entsog_2019_dataset['Unnamed: 3'].isna()]
entsog_2019_dataset.dropna(axis=1,how='all',inplace=True)
#choose useful data
entsog_2019_dataset=entsog_2019_dataset[['Point','Technical physical capacity (GWh/d)','From Identifier','To Identifier','From CC','To CC']]
#drop that a country that not in eu and also some capacity to storage
entsog_2019_dataset=entsog_2019_dataset[entsog_2019_dataset['From CC'].notna()&entsog_2019_dataset['To CC'].notna()]
#capacity must not be nan
entsog_2019_dataset=entsog_2019_dataset[entsog_2019_dataset['Technical physical capacity (GWh/d)'].notna()]
#clean point name
entsog_2019_dataset.Point=entsog_2019_dataset.Point.str.split('\\n').str[0]
# .str means not string but means dealing with single cell not as series
# +
#add point key information
#-------------------------
entsog_2019_dataset=entsog_2019_dataset.merge(operationaldata[['pointKey','pointLabel','pointTpMapX','pointTpMapY']].drop_duplicates(),left_on='Point',right_on='pointLabel',how='left')
entsog_2019_dataset.fillna('NULL',inplace=True)
#replace nan with 'NULL'
#nan is hard to deal with for single cell value
#deal with 38ZEEG
def deal_with_38ZEEG(df):
if df['From Identifier']=='38ZEEG-0007107-I' or df['To Identifier']=='38ZEEG-0007107-I':
df['pointKey']=fullpointdata.set_index('pointEicCode').loc['38ZEEG-0007107-I']['pointKey']
df['Point']=fullpointdata.set_index('pointEicCode').loc['38ZEEG-0007107-I']['pointKey']
df['pointLabel']=fullpointdata.set_index('pointEicCode').loc['38ZEEG-0007107-I']['pointLabel']
df['pointTpMapX']=fullpointdata.set_index('pointEicCode').loc['38ZEEG-0007107-I']['tpMapX']
df['pointTpMapY']=fullpointdata.set_index('pointEicCode').loc['38ZEEG-0007107-I']['tpMapY']
return df
entsog_2019_dataset=entsog_2019_dataset.apply(deal_with_38ZEEG,axis=1)
try:
entsog_2019_dataset.drop('pointLabel',axis=1,inplace=True)
except KeyError:
pass
entsog_2019_dataset.columns=['Point','Capacity(GWh/d)','From_ID','To_ID','From','To','PointKey','Longitude','Latitude']
#show the rows not match
entsog_2019_dataset[entsog_2019_dataset.PointKey=='NULL']
# +
#statistic
#---------
#remove inside country point
entsog_2019_dataset=entsog_2019_dataset[entsog_2019_dataset.From!=entsog_2019_dataset.To]
#remove not transmission type points, store in another dataframe
only_transmission_entsog_2019=entsog_2019_dataset[entsog_2019_dataset.PointKey.isin(transmission_point_list)]
self_extract_data=only_transmission_entsog_2019.groupby(['From','To']).sum()['Capacity(GWh/d)']
self_extract_data=self_extract_data.reset_index()
self_extract_data
# +
#compare table from same source
compare_table=m.merge(self_extract_data,left_on=['From','To'],right_on=['From','To'],how='left')
compare_table['diff_percent']=(compare_table.V-compare_table['Capacity(GWh/d)'])/compare_table.V
compare_table.diff_percent=compare_table.diff_percent
compare_table[(compare_table.diff_percent.abs()>0.1)|compare_table.diff_percent.isna()]
#table from lisa have more information
#selbst extract data from same source 62.5% are almost the same (different<10%)
#selbst extract data are often have larger capacity then lisa's table, when difference are greater than 10%
# -
len(entsog_2019_dataset[['PointKey','From','To']].drop_duplicates())-len(entsog_2019_dataset)
#this three parameters value combination is unique key
# ## Physical hour data
# +
'''
Multiprocessor version
'''
def get_data_hour(inputs):
df=inputs[0]
api_url=inputs[1]
process_number=inputs[2]
answer=[]
notsuccessful=0
for i in df.id:
#print('-----')
trytime=0
while trytime<2:
try:
url=api_url.format(i)
#get data
temp_table=pd.read_csv(url)
#united id
temp_table.id=i
#get max Physically flow
temp_table=temp_table[['id','value']].groupby('id').max()
#set index
temp_table.reset_index(inplace=True)
answer.append(temp_table)
break
except:
trytime+=1
if trytime==2:
#record how many rows fail to get data from server
notsuccessful+=1
print('process{} is finish with '.format(process_number),notsuccessful,'not successful')
#return a dataset
return pd.concat(answer)
def split_df(df,Processors=2):
df = df.sample(frac=1).reset_index(drop=True)
stride=(len(df)//Processors)+1
return [df.iloc[x*stride:x*stride+stride,:].copy() for x in range(Processors)]
def fix_direction(dataframe):
if dataframe['directionKey']=='entry':
temp_value=dataframe['tpTsoCountry']
dataframe['tpTsoCountry']=dataframe['adjacentCountry']
dataframe['adjacentCountry']=temp_value
return dataframe
#direction always is from tp to adj
#which means if is exit then dont need to change, entry need to be change
#enrty means gas enter tpTso
#split dataset which can be use three times
#p_split=split_df(p,multiprocessing.cpu_count())
# -
# # only firm capacity daily
# +
#load from internet
#------------------
processors=multiprocessing.cpu_count()
p_split=split_df(p_with_geodata,processors)
#get data
args=[[i,original_hour,x] for x,i in enumerate(p_split)]
with Pool(processes=processors) as pool:
physical_capacity=pool.map(get_data_hour,args)
physical_capacity=pd.concat(physical_capacity)
#physical_capacity=get_data_hour(p_with_geodata,firm_hour)
physical_capacity.to_csv('physical_capacity',index=False)
#load from local
#-----------------
#physical_capacity=pd.read_csv('physical_capacity')
#physical_capacity=physical_capacity[physical_capacity.id.isin(real_flow_list)]
#physical_capacity.drop('Unnamed: 0',axis=1,inplace=True)
physical_capacity=p_with_geodata.merge(physical_capacity,left_on='id',right_on='id')
physical_capacity=physical_capacity.apply(fix_direction,axis=1)
physical_capacity.drop('directionKey',axis=1,inplace=True)
physical_capacity_country_level=physical_capacity[['tpTsoCountry','adjacentCountry','value']].groupby(by=['tpTsoCountry','adjacentCountry']).sum().reset_index()
#from kWh/h to GWh/d
physical_capacity_country_level.value=physical_capacity_country_level.value/10e6
physical_capacity_country_level.columns=['tpTsoCountry','adjacentCountry','firm_value']
physical_capacity_country_level.head()
# -
# # something datat physical data (because of pipe in pipe)
# # firm data some times have missing data
# # physical data will be processed by entsog, one point same direction(from country a to country b) all operators willl have almost the same physical flow
# # so physical data need to be process twice
#
#
# # firm capacity
#
# # some times interruptable data have infintie
#
# # final data will be groupby pointkey+fromcountry+tocountry
# +
#some times firm capacity have no data,which means 0, so it is better to use firm with physical capacity
# +
#internet
#--------------
processors=multiprocessing.cpu_count()
p_split=split_df(p_with_geodata,processors)
#get data
args=[[i,firm_hour,x] for x,i in enumerate(p_split)]
with Pool(processes=processors) as pool:
firm_capacity=pool.map(get_data_hour,args)
firm_capacity=pd.concat(firm_capacity)
#firm_capacity=get_data_hour(p_with_geodata,firm_hour)
firm_capacity.to_csv('firm_capacity',index=False)
#local
#---------------
#firm_capacity=pd.read_csv('firm_capacity')
#firm_capacity=firm_capacity[firm_capacity.id.isin(real_flow_list)]
#firm_capacity.drop('Unnamed: 0',axis=1,inplace=True)
firm_capacity=p_with_geodata.merge(firm_capacity,left_on='id',right_on='id')
firm_capacity=firm_capacity.apply(fix_direction,axis=1)
firm_capacity.drop('directionKey',axis=1,inplace=True)
firm_capacity_country_level=firm_capacity[['tpTsoCountry','adjacentCountry','value']].groupby(by=['tpTsoCountry','adjacentCountry']).sum().reset_index()
#from kWh/h to GWh/d
firm_capacity_country_level.value=firm_capacity_country_level.value/10e6
firm_capacity_country_level.columns=['tpTsoCountry','adjacentCountry','firm_value']
firm_capacity_country_level.head()
# -
# interruptable capacity
# ------------
#
# they are different reccords with different labels, just easy to sum up
# +
processors=multiprocessing.cpu_count()
p_split=split_df(p_with_geodata,processors)
#get data
args=[[i,interrupt_hour,x] for x,i in enumerate(p_split)]
with Pool(processes=processors) as pool:
inter_capacity=pool.map(get_data_hour,args)
inter_capacity=pd.concat(inter_capacity)
inter_capacity.to_csv('inter_capacity',index=False)
#inter_capacity=pd.read_csv('inter_capacity')
#inter_capacity=inter_capacity[inter_capacity.id.isin(real_flow_list)]
#inter_capacity.drop('Unnamed: 0',axis=1,inplace=True)
inter_capacity=p_with_geodata.merge(inter_capacity,left_on='id',right_on='id')
inter_capacity=inter_capacity.apply(fix_direction,axis=1)
inter_capacity.drop('directionKey',axis=1,inplace=True)
inter_capacity_country_level=inter_capacity[['tpTsoCountry','adjacentCountry','value']].groupby(by=['tpTsoCountry','adjacentCountry']).sum().reset_index()
#from kWh/h to GWh/d
inter_capacity_country_level.value=inter_capacity_country_level.value/10e6
inter_capacity_country_level.value=inter_capacity_country_level.value.round()
inter_capacity_country_level.columns=['tpTsoCountry','adjacentCountry','inter_value']
inter_capacity_country_level.head()
# +
#there is problem, same different direction id means the same flow and the same pipes, need to drop duplicate
#solve remove id and drop duplicates
def drop_id_drop_duplicate(df):
df.drop('id',axis=1,inplace=True)
len_before=len(df)
df.drop_duplicates(inplace=True)
len_after=len(df)
print('before: ',len_before,' after: ',len_after)
return df
physical_capacity=drop_id_drop_duplicate(physical_capacity)
firm_capacity=drop_id_drop_duplicate(firm_capacity)
inter_capacity=drop_id_drop_duplicate(inter_capacity)
# -
x=physical_capacity[['pointKey','tpTsoCountry','adjacentCountry']].copy()
x['m']=1
x=x.groupby(['pointKey','tpTsoCountry','adjacentCountry']).count().reset_index()
x.sort_values('m')
final_capacity=inter_capacity.merge(firm_capacity,how='outer',left_on=['tpTsoCountry','adjacentCountry'],right_on=['tpTsoCountry','adjacentCountry'])
#final_capacity=final_capacity.merge(physical_capacity,how='outer',left_on=['tpTsoCountry','adjacentCountry'],right_on=['tpTsoCountry','adjacentCountry'])
final_capacity['day']=final_capacity.firm_value*24
def add_compare(dataframe):
return compare_data[dataframe['adjacentCountry']][dataframe['tpTsoCountry']]
final_capacity['compare_value']=final_capacity.apply(add_compare,axis=1)
ak=final_capacity.merge(self_extract_data,left_on=[ 'tpTsoCountry','adjacentCountry'],right_on=['From','To'],how='outer')
#GWh/d
ak[ak.tpTsoCountry.notna()]
# +
entsog family
igg
change the unit
pdf about diff dataset
capacity per hour + start and end highest resolution (city longtitude and lagtitude)
try with the different pressure samll area ;country; all
check
segement error
# +
#physical
#arg=[original_hour for i in p_split]
#with Pool(multiprocessing.cpu_count()) as pool:
# physical_capacity=pool.starmap(get_data_hour,zip(p_split,arg))
#-----------------------
##get data from internet
#physical_capacity=get_data_hour(p,original_hour)
#physical_capacity.to_csv('physical_capacity')
#-----------------------
#load from local
physical_capacity=pd.read_csv('physical_capacity')
physical_capacity=physical_capacity[physical_capacity.id.isin(real_flow_list)]
physical_capacity.drop('Unnamed: 0',axis=1,inplace=True)
physical_capacity.drop_duplicates(inplace=True)
physical_capacity_with_geodata=p_with_geodata.drop_duplicates().merge(physical_capacity,left_on='id',right_on='id')
physical_capacity_with_geodata=physical_capacity_with_geodata.apply(fix_direction,axis=1)
physical_capacity=p.merge(physical_capacity,left_on='id',right_on='id')[['directionKey','tpTsoCountry','adjacentCountry','value']]
physical_capacity=physical_capacity.apply(fix_direction,axis=1)
physical_capacity.drop('directionKey',axis=1,inplace=True)
physical_capacity=physical_capacity.groupby(by=['tpTsoCountry','adjacentCountry']).sum().reset_index()
#from kWh/h to GWh/d
physical_capacity.value=physical_capacity.value/10e6
physical_capacity.columns=['tpTsoCountry','adjacentCountry','physical_value']
physical_capacity.head()
| gas_network_exploring/notebook/get_Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
##IMPORTS
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler
from sklearn.cluster import KMeans
from sklearn.preprocessing import MinMaxScaler, RobustScaler
from sklearn.model_selection import train_test_split
import warnings
warnings.filterwarnings('ignore')
#Modeling Tools
from sklearn.model_selection import train_test_split
from sklearn.impute import SimpleImputer
import statsmodels.api as sm
from statsmodels.formula.api import ols
from datetime import date
from scipy import stats
#Custom functions
from env import host, user, password #Database credentials
import wrangle4
import env
import eval_model
import explore2
import model
## Evaluation tools
from sklearn.metrics import mean_squared_error
from sklearn.metrics import r2_score
from math import sqrt
# -
train, X_train, y_train, X_validate, y_validate, X_test, y_test=wrangle4.wrangle()
df=train
import lux
df
df.intent =['logerror']
df
from lux.vis.Vis import Vis
Vis(["Region=New England","MedianEarnings"],df)
| lux.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Daten umwandeln - Strings und Zahlen
# Daten umzuwandeln ist wahrscheinlich, das was du am meisten machen wirst. Deshalb ist es praktisch es schon jetzt kennen zu lernen.
# ### Einen String in eine Zahl umwandeln
# Dazu wenden wir den **int()**-Befehl auf einen String (direkt oder in einer Variable gespeichert) an: **int(string)**
#
# int steht für Integer, den englischen Begriff für Ganzzahl.
# +
a = "5"
b = "6"
print(int(a) + int(b))
# -
# ### Einen String in eine Kommazahl umwandeln
# Dazu wenden wir den **float()**-Befehl auf einen String (direkt oder in einer Variable gespeichert) an: **float(string)**
#
#
# Der Name float kommt daher, dass man Kommazahlen auch Fließkommazahlen nennt.
# +
a = "5.5"
b = "6.6"
print(float(a) + float(b))
# -
# ### Eine Zahl in einen String umwandeln
# Dazu wenden wir den **str()**-Befehl auf eine Ganzzahl oder Kommazahl an (direkt oder in einer Variable gespeichert): **str(zahl)**
age = 21
print("Ich bin " + str(age) + " Jahre alt")
# ## Übung
#
# - Speichere die Zahl PI mit 4 Nachkommastellen in der Variable a.
# - Multipliziere sie mit 2.
# - Wandle die Zahl in einen String um und gebe sie aus als Satz: "Das doppelter der Zahl Pi ist: a"
#
#
# # Daten umwandeln - Listen und Strings
# Wir können mit Python Elemente zu einer Liste zusammenfügen oder eine Liste in einzelne Elemente zerlegen.
# ### Strings aus einer Liste zu einem String zusammenfügen
#
# Mit dem **join()**-Befehl, der auf einen String angewendet wird, verbinden wir die Strings aus einer Liste zu einem neuen String: **string.join(liste)**
#
# Der String, auf den join() angewendet wird, bildet dabei die Nahtstelle: Dieser String wird als Verbindung zwischen den einzelnen Listenelementen im neuen String gesetzt.
students = ["Max", "Monika", "Erik", "Franziska"]
print(", ".join(students))
students_as_string = ", ".join(students)
print("An unserer Uni studieren: " + students_as_string)
students = ["Max", "Monika", "Erik", "Franziska"]
print(" - ".join(students))
# ### Einen String in eine Liste aufspalten
#
# Mit dem **split()**-Befehl, der auf einen String angewendet wird, wird dieser String an seinen Leerzeichen aufgespalten und die daraus resultierenden Einzelstrings in einer Liste gespeichert: **string.split()**
i = "Max, Monika, Erik, Franziska"
print(i.split())
# Wir können sogar noch genauer festlegen, an welchen Stellen der String von split() aufgespaltet werden soll.
print(i.split(", "))
print(i.split("a"))
# Insbesondere können wir auch mehrere der Befehle, die wir schon kennen gelernt haben, miteinander kombinieren:
# +
# Hier zählen wir die Anzahl der Wörter des Satzes s
s = "Ich bin ein Satz mit vielen Wörtern"
print(len(s.split()))
# -
# ## Übung 1
# - Nimm einen Artikel aus 20 minuten.
# - Kopiere ihn in einen String.
# - Zerlege den Artikel in Wörter.
# - Zähle die Wörter.
# - Gib aus " Der Artikel xyz enthält x wörter"
# ## Übung 2
#
# - Lass dir vom Nachbarn seine Email geben.
# - Wenn die Mailadresse _<EMAIL>_ lautet, sollst du _Max-Mustermann_ ausgeben; wenn die Mailadresse _<EMAIL>_ heisst, sollst du _KlaraKlarnamen_ ausgeben.
# ## Übung 3
#
# Aktuell legen alle Kunden (`mail1`, `mail2`, `mail3`) als separate Variable vor. Wir möchten daraus jetzt eine Liste bauen, sodass wir die Möglichkeit hätten, später noch weitere Kunden in diese Liste hinzuzufügen.
#
# Überführe deswegen die Kunden `mail1`, `mail2` und `mail3` in die Liste `clients` und lasse dir anschließend die Anzahl der Elemente der Liste `clients` mit Hilfe von Python ausgeben.
# +
mail1 = "<EMAIL>"
mail2 = "<EMAIL>"
mail3 = "<EMAIL>"
clients = []
# Füge hier mail1, mail2, mail3 zur clients - Liste hinzu
# Zähle wieviele elemente die liste hat.
print(clients)
| 02 Python Teil 1/05 Daten umwandeln.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="N0ruWqaR37Oc" colab_type="text"
# # Hur påverkades Bokskogen av torkan 2018?
# I denna uppgift studerar vi [Bokskogen](https://sv.wikipedia.org/wiki/Torups_rekreationsomr%C3%A5de) i centrala Skåne och dess närliggande områdes växtlighet med hjälp av satellitbilder. Bilderna är alla tagna under första halvan av augusti, men från tre olika år. Kan vi se någon skillnad efter en sommar med torka? Vi kommer använda teori om NDVI (Normalized Difference Vegetation Index) som är ett mått på hur mycket levande växtlighet det finns. Vi börjar med ett kort teoriavsnitt för att få mer känsla för vad NDVI är.
#
# + [markdown] id="V1UxA4EIMK2u" colab_type="text"
# ## Normalized Difference Vegetation Index (NDVI)
# [NDVI](https://sv.wikipedia.org/wiki/NDVI) är ett värde som kan beräknas från hur mycket ljus som reflekteras i en bilds våglängdsband i nära-infrarött och rött ljus. Indexet används oftast för bilder från satelliter rymden, och indikerar mängden levande vegetation. En mer detaljerad förklaring över hur NDVI används för detta syfte finns [här](https://earthobservatory.nasa.gov/features/MeasuringVegetation/measuring_vegetation_2.php).
#
#
# Frisk vegetation ([klorofyll](https://sv.wikipedia.org/wiki/Klorofyll)) reflekterar mer nära-infrarött (NIR) och grönt ljus i jämförelse med andra våglängder, men absorberar mer rött och blått ljus. NDVI beräknas genom att ta foto på NIR-våglängder och röda våglängder och sedan för varje pixel beräkna NDVI med formeln nedan.
#
# <br>
#
# $$NDVI = \frac{(NIR-Red)}{(NIR+Red)}.$$
#
# <br>
#
# Ovanstående formel genererar resultat mellan -1 och 1. Låg reflektion (eller låga värden) i det röda bandet och hög reflektion i det nära-infraröda bandet producerar ett högt NDVI värde. Och vice versa. I allmänhet, indikerar höga NDVI-värden på frisk vegetation medan låga NDVI-värden pekar på mindre eller ingen vegetation.
#
#
# NDVI kan även användas som ett verktyg för att identifiera torka genom att kolla på skillnader från år till år. Man kan analysera detta numeriskt men i denna uppgiften ska vi endast se visuellt vad som händer genom att plotta NDVI för tre år. Vill man se mer sofistikerade metoder kan man kolla in fortsättningsuppgiften till denna: Torkan 2.
#
# + [markdown] id="3BDFrVp9SwpU" colab_type="text"
# ## 1. Växtligheten 2015
#
# Vi börjar med att kolla på sensommaren 2015. I mappen Bokskogen ligger det tre filer som alla heter `data_1X.npz`. Detta är numpy-arrays som innehåller ett heltal för varje pixel, som tillsammans kan skapa en bild. Kör följande kod för att ladda in alla nödvändiga paket, npz-filer och en färgbild över området vi kommer kolla på:
# + id="2PxkNQM43OoS" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
from IPython.display import Image
#Download the data used in the exercise.
# !wget https://github.com/lunduniversity/schoolprog-satellite-data/raw/master/drought/bokskogen/data_15.npz --no-verbose
# !wget https://github.com/lunduniversity/schoolprog-satellite-data/raw/master/drought/bokskogen/data_17.npz --no-verbose
# !wget https://github.com/lunduniversity/schoolprog-satellite-data/raw/master/drought/bokskogen/data_18.npz --no-verbose
# !wget https://github.com/lunduniversity/schoolprog-satellite/raw/master/exercises/drought/tci_hd.png --no-verbose
# + [markdown] id="9rB3pKplypF9" colab_type="text"
# Nu ska vi visa bilden vi laddade ned, samt läsa in filen som innehåller bilder från 2015.
# + id="z65aKXqwy3kV" colab_type="code" colab={}
#Visa bilden
display(Image(filename="tci_hd.png", width = 700, height = 700))
#Ladda in filen från 2015
bands15 = np.load('data_15.npz')
# + [markdown] id="GBwHyYRaXGAz" colab_type="text"
# Vår `bands15`-variabel fungerar nu ungefär som en `dict` och innehåller data för både det röda och det nära infraröda bandet. För att plocka ut något från `bands15` måste vi använda nycklar som vi gjort i tidigare uppgifter med `dict`.
#
# **Uppdrag:** Kan du själv ta reda på vad nycklarna till `bands15` är?
# + id="-aodDVpO65sJ" colab_type="code" colab={}
# + [markdown] id="ZKKOYgfp66OT" colab_type="text"
#
# <details>
# <summary markdown="span">
# Tips
# </summary>
# <p>Du kan använda funktionerna <code>print(list())</code> tillsammans med metoden <code>keys()</code>
# </p>
# </details>
#
# <details>
# <summary markdown="span">
# Svar
# </summary>
# <p><code>print(list(bands15.keys()))</code> ger nycklarna <code>"red"</code> och <code>"nir"</code> (nir = near-infrared)</p>
# </details>
# + [markdown] id="AZXd4i0eXQtl" colab_type="text"
#
# För att lättare kunna använda datan vill vi nu skapa var sin variabel för de båda banden.
#
# **Uppdrag:** Skapa variablerna `red15` och `nir15` med hjälp av `bands15`.
# + id="XcR6GCX7W3hG" colab_type="code" colab={}
red15 =
nir15 =
# + [markdown] id="lFhKDiQ8YHJv" colab_type="text"
# <details>
# <summary markdown="span">
# Svar
# </summary>
# <p><pre>red15 = bands15['red']
# nir15 = bands15['nir']</pre>
# </p>
# </details>
#
# + [markdown] id="uSRhKkWjYQhu" colab_type="text"
# Vi har nu delat upp datan för de båda banden var för sig. För att få en känsla av vad det faktiskt är för data vi har att göra med skulle vi kunna testa plotta dem.
#
# **Uppdrag:** Plotta en eller båda variablerna vi precis skapade. Vad visar plotten?
#
#
# + id="78f5GZI_XmWD" colab_type="code" colab={}
# + [markdown] id="IXMw3PHIYlT-" colab_type="text"
# <details>
# <summary markdown="span">
# Tips
# </summary>
# <p><pre>plt.figure(figsize=(10,10))
# plt.imshow(red15)
# plt.savefig('red15.png')</pre>
# </p>
# </details>
#
# <details>
# <summary markdown="span">
# Svar
# </summary>
# <p>Plotten visar en bild över Bokskogen med närliggande åkrar och sjöar. Bilderna består endast av ljus från röda respektive nära infraröda bandet, vilket gör att det blir svårt att se vad de föreställer.
# </p>
# </details>
# + [markdown] id="wNlN4AjFZIS0" colab_type="text"
# För att uträkningar ska bli mer exakta vill vi egentligen ha våra värden i form av flyttal. Gör om `red15` och `nir15` till flyttal med hjälp av följande kod:
# + id="JIbwD3ijYlsH" colab_type="code" colab={}
red15 = red15.astype(float)
nir15 = nir15.astype(float)
# + [markdown] id="hCsBLZD0ZTcU" colab_type="text"
# För att lättare kunna tolka bilden vill vi räkna ut NDVI. Detta kommer göra att partier med mycket växtligher skiljer sig i färg gentemot partier utan växtlighet när vi plottar bilden.
#
# **Uppdrag:** Skapa en variabel `ndvi15` och räkna ut den med hjälp av `red15` och `nir15`.
# + id="RPn3OShFZLq0" colab_type="code" colab={}
ndvi15 =
# + [markdown] id="BZTUW76DZZPh" colab_type="text"
# <details>
# <summary markdown="span">
# Tips
# </summary>
# <p>
# NDVI kan bestämmas genom ekvationen <code>ndvi = (nir-red)/(nir+red)</code>
# </p>
# </details>
#
# <details>
# <summary markdown="span">
# Svar
# </summary>
# <p>
# <code>ndvi15 = (nir15-red15)/(nir15+red15)</code>
# </p>
# </details>
# + [markdown] id="Cbtx6_VDZimP" colab_type="text"
# Vi skulle nu kunna plotta bilden som innan, men för att göra det tydligare använder vi några speciella inställningar. Kör koden nedan:
# + id="9wN6znisZZv6" colab_type="code" colab={}
plt.figure(figsize=(10,10))
plt.pcolormesh(ndvi15, cmap='PiYG')
plt.ylim(ndvi15.shape[0], 0)
plt.clim(-1.0, 1.0)
plt.colorbar(label='NDVI')
plt.savefig('colormap.png')
# + [markdown] id="SiCafXTdZpJw" colab_type="text"
# **Uppdrag:** Förstår du ungefär vad de olika metoderna gör? Testa ta bort någon del och se vad som händer!
#
# <details>
# <summary markdown="span">
# Svar
# </summary>
# <p><ul>
# <li><code>plt.figure(figsize=(10,10))</code> skapar en ny figur som har storleken 10*10 inches (tum).</li>
# <li><code>plt.pcolormesh(ndvi15, cmap='PiYG')</code> skapar en färgplot av arrayen som matas in, i detta fallet <code>ndvi15</code>. En färgplot innebär att programmet tar arrayen och läser in den som en bild, och varje värde anger färgen på en pixel. Hur färgerna fördelas kan man ange med <code>cmap</code>-parametern.
# </p></li>
# <li><code>plt.ylim(ndvi.shape[0], 0)</code> sätter gränserna för y-axeln. Om vi inte skulle använt detta skulle bilden hamnat upp och ner (testa att ta bort denna och se vad som händer). Detta är för att x-axeln och y-axeln för <code>pcolormesh()</code> börjar nere i vänstra hörnet och ökar till höger respektivt uppåt. Men arrayer börjar uppe i vänstra hörnet och ökar till höger respektive nedåt. <code>ndvi.shape[0]</code> ger oss antalet rader i <code>ndvi</code> så vad vi egentligen gör med <code>plt.ylim(ndvi.shape[0], 0)</code> är att säga till programmet att börja plotta rad <code>ndvi.shape[0]</code> till <code>0</code>, vilket gör bilden rättvänd.</li>
# <li><code>plt.clim(-1.0, 1.0)</code> sätter gränserna för färgskalan. Anledningen till att vi sätter -1 till 1 är att NDVI-värden alltid är mellan -1 och 1. Skulle vi inte gjort detta är det de största repektive minsta värdena i <code>ndvi15</code> som sätter gränserna för färgskalan. Detta ger en omotiverad kontrast. </li>
# <li><code>plt.colorbar(label='NDVI')</code> gör att vi får ett fält på sidan grafen som illustrerar färgskalan. Vi anger också att det ska stå NDVI bredvid fältet.</li>
# <li>Slutligen använder vi <code>plt.savefig('colormap.png')</code> som helt enkelt sparar grafen till filen <code>colormap.png</code>.</li>
# </ul>
#
# </details>
# + [markdown] id="7q7TExmjZyPH" colab_type="text"
# **Uppdrag:** Vad visar bilden? Kan du se vilka bönder som inte har skördat sina åkrar än?
#
# <details>
# <summary markdown="span">
# Svar
# </summary>
# <p>
# En grön åker tyder på att det finns växtlighet där, medan en vitare åker tyder på att det finns låg växtlighet och att åkern troligtvis är skördad.
# </p>
# </details>
# + [markdown] id="tPTngrXohG15" colab_type="text"
# ## 2. Jämför växtlighet
# För att kunna jämföra åren måste vi nu egentligen göra samma sak för 2017 och 2018. Vi skulle kunna skriva ungefär samma kod som vi precis skrivit ytterligare två gånger, men detta är inte särskilt effektivt eftersom vi i princip skriver samma sak tre gånger med ett fåtal utbytta ord. För ändamål som dessa är därför funktioner väldigt användbara.
# + [markdown] id="SQV_bLN1hIPe" colab_type="text"
# **Uppdrag:** Skriv en funktion `plot_ndvi(file_name, save_name)` som tar in en sträng `file_name`, som är namnet på `.npz`-filen du vill läsa in datan från, och en sträng `save_name`, som blir namnet på den bild som plottas. Bilden som plottas ska vara baserad på NDVI med liknande inställningar som innan.
# + id="A9dCU8EsZs62" colab_type="code" colab={}
def plot_ndvi(file_name, save_name):
#Skriv din kod här..
# + [markdown] id="Pe0ifcF4hfxb" colab_type="text"
# <details>
# <summary markdown="span">
# Tips
# </summary>
# <p>
# Du kan återanvända väldigt stor del av den kod du redan skrivit.
# </p>
# </details>
#
# <details><summary markdown="span">Lösning</summary>
# <p>
# <pre><code>def plot_ndvi(file_name, save_name):
# bands = np.load(file_name)
# red = bands['red'].astype(float)
# nir = bands['nir'].astype(float)
# ndvi = (nir-red) / (nir+red)
# plt.figure(figsize=(10,10))
# plt.pcolormesh(ndvi, cmap='PiYG')
# plt.ylim(ndvi.shape[0], 0)
# plt.clim(-1.0, 1.0)
# plt.colorbar(label='NDVI')
# plt.savefig(save_name)</code></pre>
# </details>
#
# + [markdown] id="x5XtdmS_hoO7" colab_type="text"
# Nu när du har en smidig funktion för att plotta NDVI kan du enkelt göra detta för alla tre år.
#
# **Uppdrag:** Plotta NDVI för 2015, 2017 och 2018. Jämför bilderna. Kan du se någon skillnad? Om du ser skillnad, vilken och varför?
#
# + id="3dB2dfXp8vv0" colab_type="code" colab={}
# + [markdown] id="KK-UgRaA8wjd" colab_type="text"
# <details><summary markdown="span">Lösning</summary>
# <p>
# <pre><code>plot_ndvi("data_15.npz", "bok15.png")
# plot_ndvi("data_17.npz", "bok17.png")
# plot_ndvi("data_18.npz", "bok18.png")
# </code></pre>
# </details>
#
# <details><summary markdown="span">Svar</summary>
# <p>
# Torkan 2018 hade sin påverkan på växtligheten. Om du kollar på åkrarna ser du att växtligheten inte är i närheten av de tidigare åren. Det ser ut som att många bönder blev tvugna att skörda de grödor som klarade sig mycket tidigare på grund av torkan. Man kan även se på själva skogen att växtligheten har sjunkit då den inte framstår som lika grön.
# </details>
# + [markdown] id="9r1x_ynT9OC-" colab_type="text"
# # Fortsättningsuppgifter
# - Modifiera funktionen `plot_ndvi()` så att du kan ange en titel för figuren och plotta figurerna igen fast med beskrivande titlar.
| exercises/drought/Bokskogen.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit ('3.8')
# name: python3
# ---
from tqdm.notebook import tqdm
import json
import collections
import pandas as pd
import numpy as np
import glob
# ## Load JSON files
sorted(glob.glob('./../data/*/*.json'))
with open('./../data/projects/project_ids.json', 'rb') as f:
project_ids = json.load(f)
len(set(project_ids))
with open('./../data/activity_facts/2021-06-07 13:00:40.json', 'rb') as f:
activity_facts_dict = json.load(f)
activities = pd.DataFrame(activity_facts_dict).drop_duplicates()
with open('./../data/analyses/2021-06-07 13:01:54.json', 'r') as f:
analyses_dict = json.load(f)
analyses = pd.DataFrame(analyses_dict).drop_duplicates()
with open('./../data/html_pages/2021-06-07 13:13:26.json', 'r') as f:
html_pages_dict = json.load(f)
html_pages = pd.DataFrame(html_pages_dict).drop_duplicates().set_index('project_id')
# # Fix data types
activities.month = pd.to_datetime(activities.month).dt.tz_localize(None)
for column in set(activities.columns) - {'month', 'project_id'}:
activities[column] = activities[column].astype(int)
activities.dtypes
# +
analyses.min_month = pd.to_datetime(analyses.min_month).dt.tz_localize(None)
analyses.max_month = pd.to_datetime(analyses.max_month).dt.tz_localize(None)
analyses.updated_at = pd.to_datetime(analyses.updated_at).dt.tz_localize(None)
analyses.oldest_code_set_time = pd.to_datetime(analyses.oldest_code_set_time).dt.tz_localize(None)
projects = analyses.groupby('project_id')[['updated_at', 'min_month', 'max_month']].max()
projects.dtypes
# -
projects.shape[0]
# # Trim & filter
trimmed_activities = activities[(activities.month.dt.year>=1991) & (activities.month.dt.year<=2020)]
duplicate_projects = html_pages[html_pages.original_project_name.notnull()]
trimmed_activities_no_duplicates = trimmed_activities[~trimmed_activities.project_id.isin(set(duplicate_projects.index))]
valid_project_ids = set(trimmed_activities_no_duplicates.project_id)
valid_projects = projects[projects.index.isin(valid_project_ids)]
valid_projects.to_hdf('../data/openhub.h5', key='valid_projects')
# # Fill all activities
def fill(project_id, df, last_seen):
df = df.set_index('month')
if last_seen:
ts = min(last_seen.to_datetime64(), np.datetime64('2020-12-01T00:00:00.000000000'))
try:
df = df.append(new_entry, verify_integrity=True, sort=False).sort_index()
except: # entry (month) does already exist
pass
df = df.asfreq('MS', fill_value=0)
df['project_id'] = project_id
return df
last_seen_dict = dict((valid_projects.updated_at.astype('datetime64[M]')) - pd.DateOffset(months=1))
groups = list(trimmed_activities_no_duplicates.groupby('project_id'))
filled_activities = [fill(*group, last_seen_dict.get(group[0])) for group in tqdm(groups)]
final = pd.concat(filled_activities).reset_index().rename(columns={'index': 'month'})
final.set_index(['month', 'project_id']).sort_index().to_hdf('../data/openhub.h5', key='filled_activities')
# # Overview
# ## Projects
projects.index.nunique()
activities.project_id.nunique()
trimmed_activities.project_id.nunique()
trimmed_activities_no_duplicates.project_id.nunique()
# ## Duplicates
trimmed_activities.project_id.nunique() - trimmed_activities_no_duplicates.project_id.nunique()
# Linux kernel is `'e206a54e97690cce50cc872dd70ee896'`
html_pages[html_pages.original_project_name=='e206a54e97690cce50cc872dd70ee896'].original_project_name.count()
valid_projects.shape
| notebooks/02 Preprocessing.ipynb |
# -*- coding: utf-8 -*-
# # Automatic generation of Notebook using PyCropML
# This notebook implements a crop model.
# ### Model UpdateCalendar
# +
# coding: utf8
from pycropml.units import u
from copy import copy
from array import array
import numpy
from math import *
from datetime import datetime
def model_UpdateCalendar(cumulTT = 741.510096671757,
calendarMoments = ["Sowing"],
calendarDates = [datetime(2007, 3, 21)],
calendarCumuls = [0.0],
currentdate = datetime(2007, 3, 27),
phase = 1.0):
"""
- Name: UpdateCalendar -Version: 1.0, -Time step: 1
- Description:
* Title: Calendar Model
* Author: <NAME>
* Reference: Modeling development phase in the
Wheat Simulation Model SiriusQuality.
See documentation at http://www1.clermont.inra.fr/siriusquality/?page_id=427
* Institution: INRA Montpellier
* Abstract: Lists containing for each stage the date it occurs as well as a copy of all types of cumulated thermal times
- inputs:
* name: cumulTT
** description : cumul thermal times at current date
** variablecategory : auxiliary
** datatype : DOUBLE
** min : -200
** max : 10000
** default : 741.510096671757
** unit : °C d
** inputtype : variable
* name: calendarMoments
** description : List containing apparition of each stage
** variablecategory : state
** datatype : STRINGLIST
** default : ['Sowing']
** unit :
** inputtype : variable
* name: calendarDates
** description : List containing the dates of the wheat developmental phases
** variablecategory : state
** datatype : DATELIST
** default : ['2007/3/21']
** unit :
** inputtype : variable
* name: calendarCumuls
** description : list containing for each stage occured its cumulated thermal times
** variablecategory : state
** datatype : DOUBLELIST
** default : [0.0]
** unit : °C d
** inputtype : variable
* name: currentdate
** description : current date
** variablecategory : auxiliary
** datatype : DATE
** default : 2007/3/27
** unit :
** inputtype : variable
* name: phase
** description : the name of the phase
** variablecategory : state
** datatype : DOUBLE
** min : 0
** max : 7
** default : 1
** unit :
** inputtype : variable
- outputs:
* name: calendarMoments
** description : List containing apparition of each stage
** variablecategory : state
** datatype : STRINGLIST
** unit :
* name: calendarDates
** description : List containing the dates of the wheat developmental phases
** variablecategory : state
** datatype : DATELIST
** unit :
* name: calendarCumuls
** description : list containing for each stage occured its cumulated thermal times
** variablecategory : state
** datatype : DOUBLELIST
** unit : °C d
"""
if phase >= 1.0 and phase < 2.0 and "Emergence" not in calendarMoments:
calendarMoments.append("Emergence")
calendarCumuls.append(cumulTT)
calendarDates.append(currentdate)
elif phase >= 2.0 and phase < 3.0 and "FloralInitiation" not in calendarMoments:
calendarMoments.append("FloralInitiation")
calendarCumuls.append(cumulTT)
calendarDates.append(currentdate)
elif phase >= 3.0 and phase < 4.0 and "Heading" not in calendarMoments:
calendarMoments.append("Heading")
calendarCumuls.append(cumulTT)
calendarDates.append(currentdate)
elif phase == 4.0 and "Anthesis" not in calendarMoments:
calendarMoments.append("Anthesis")
calendarCumuls.append(cumulTT)
calendarDates.append(currentdate)
elif phase == 4.5 and "EndCellDivision" not in calendarMoments:
calendarMoments.append("EndCellDivision")
calendarCumuls.append(cumulTT)
calendarDates.append(currentdate)
elif phase >= 5.0 and phase < 6.0 and "EndGrainFilling" not in calendarMoments:
calendarMoments.append("EndGrainFilling")
calendarCumuls.append(cumulTT)
calendarDates.append(currentdate)
elif phase >= 6.0 and phase < 7.0 and "Maturity" not in calendarMoments:
calendarMoments.append("Maturity")
calendarCumuls.append(cumulTT)
calendarDates.append(currentdate)
return (calendarMoments, calendarDates, calendarCumuls)
# -
from pycropml.units import u
params= model_updatecalendar(
cumulTT = 112.330110409888*u.°C d,
calendarMoments = ["Sowing"]*u.,
calendarDates = [datetime(2007, 3, 21) ,]*u.,
calendarCumuls = [0.0]*u.°C d,
phase = 1*u.,
)
calendarMoments_estimated = params[0]
calendarMoments_computed = ["Sowing", "Emergence"]*u.
assert calendarMoments_computed == calendarMoments_estimated
calendarDates_estimated = params[1]
calendarDates_computed = ["2007/3/21", "2007/3/27"]*u.
assert calendarDates_computed == calendarDates_estimated
calendarCumuls_estimated =params[2].round(2)*params[2].units
calendarCumuls_computed = [ 0.0 ,112.33]*u.°C d
assert numpy.all(calendarCumuls_estimated == calendarCumuls_computed)
| src/pycropml/transpiler/antlr_py/csharp/examples/pheno_pkg/test/py/UpdateCalendar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
s = pd.Series(np.random.randn(4), name='daily returns')
s
s.describe()
s.index
s.index = ['AMZN', 'AAPL', 'MSFT', 'GOOG']
s
import urllib.request
source = urllib.request.urlopen('http://research.stlouisfed.org/fred2/series/UNRATE/downloaddata/UNRATE.csv')
data = pd.read_csv(source, index_col=0, parse_dates=True, header=None)
type(data)
data.head()
data.describe()
from pandas_datareader import data,wb
import datetime as dt
start, end = dt.datetime(2006, 1, 1), dt.datetime(2016, 12, 31)
data2 = data.DataReader('UNRATE', 'fred', start, end)
data2.plot()
plt.show()
# ##### Exercise 1
# +
ticker_list = {'INTC': 'Intel',
'MSFT': 'Microsoft',
'IBM': 'IBM',
'BHP': 'BHP',
'TM': 'Toyota',
'AAPL': 'Apple',
'AMZN': 'Amazon',
'BA': 'Boeing',
'QCOM': 'Qualcomm',
'KO': 'Coca-Cola',
'GOOG': 'Google',
'SNE': 'Sony',
'PTR': 'PetroChina'}
start = dt.datetime(2013, 1, 1)
end = dt.datetime.today()
price_change = {}
for ticker in ticker_list:
prices = data.DataReader(ticker, 'yahoo', start, end)
closing_prices = prices['Close']
change = 100 * (closing_prices[-1] - closing_prices[0]) / closing_prices[0]
name = ticker_list[ticker]
price_change[name] = change
pc = pd.Series(price_change)
pc.sort_values(inplace=True)
fig, ax = plt.subplots(figsize=(10,8))
pc.plot(kind='bar', ax=ax)
plt.show()
# -
| Part1-Python/Pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="kA4Pu2sPr_eG" colab_type="code" outputId="cf48f5a3-15a9-48d9-e6cf-c0a0cf639043" executionInfo={"status": "ok", "timestamp": 1590678647212, "user_tz": 180, "elapsed": 4883, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjWI4QgYDUyC8ZcW5KHpuBwcY-MZWVqbPWcZSzxIg=s64", "userId": "06256612867315483887"}} colab={"base_uri": "https://localhost:8080/", "height": 318}
# !nvidia-smi
# + id="rOXNne0OoLTd" colab_type="code" outputId="7779eb3a-5d5d-47ba-be0d-4f290e7c3e74" executionInfo={"status": "ok", "timestamp": 1590678661772, "user_tz": 180, "elapsed": 12120, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjWI4QgYDUyC8ZcW5KHpuBwcY-MZWVqbPWcZSzxIg=s64", "userId": "06256612867315483887"}} colab={"base_uri": "https://localhost:8080/", "height": 105}
# !pip --quiet install transformers
# !pip --quiet install tokenizers
# + id="htKChfMHoYRb" colab_type="code" outputId="1e486edf-ff8e-40a0-c6ee-6bfe75e9b7c0" executionInfo={"status": "ok", "timestamp": 1590678696281, "user_tz": 180, "elapsed": 46617, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjWI4QgYDUyC8ZcW5KHpuBwcY-MZWVqbPWcZSzxIg=s64", "userId": "06256612867315483887"}} colab={"base_uri": "https://localhost:8080/", "height": 125}
from google.colab import drive
drive.mount('/content/drive')
# + id="xtDZkoyzod_S" colab_type="code" colab={}
# !cp -r '/content/drive/My Drive/Colab Notebooks/Tweet Sentiment Extraction/Scripts/.' .
# + id="38KKgQn7owCr" colab_type="code" colab={}
COLAB_BASE_PATH = '/content/drive/My Drive/Colab Notebooks/Tweet Sentiment Extraction/'
MODEL_BASE_PATH = COLAB_BASE_PATH + 'Models/Files/205-roBERTa_base/'
import os
os.mkdir(MODEL_BASE_PATH)
# + [markdown] id="MOE8CNSAnvq-" colab_type="text"
# ## Dependencies
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _kg_hide-input=true _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" id="x48gxo3vnvrA" colab_type="code" outputId="c8637f4a-4034-4fbd-8c5e-e3cc8ad48472" executionInfo={"status": "ok", "timestamp": 1590678708202, "user_tz": 180, "elapsed": 58519, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjWI4QgYDUyC8ZcW5KHpuBwcY-MZWVqbPWcZSzxIg=s64", "userId": "06256612867315483887"}} colab={"base_uri": "https://localhost:8080/", "height": 72}
import json, warnings, shutil
from scripts_step_lr_schedulers import *
from tweet_utility_scripts import *
from tweet_utility_preprocess_roberta_scripts_aux import *
from transformers import TFRobertaModel, RobertaConfig
from tokenizers import ByteLevelBPETokenizer
from tensorflow.keras.models import Model
from tensorflow.keras import optimizers, metrics, losses, layers
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint, LearningRateScheduler
SEED = 0
seed_everything(SEED)
warnings.filterwarnings("ignore")
pd.set_option('max_colwidth', 120)
# + [markdown] id="6HW0GQzPnvrF" colab_type="text"
# # Load data
# + id="jS858zlQNNCs" colab_type="code" colab={}
# Unzip files
# !tar -xf '/content/drive/My Drive/Colab Notebooks/Tweet Sentiment Extraction/Data/complete_64_clean/fold_1.tar.gz'
# !tar -xf '/content/drive/My Drive/Colab Notebooks/Tweet Sentiment Extraction/Data/complete_64_clean/fold_2.tar.gz'
# !tar -xf '/content/drive/My Drive/Colab Notebooks/Tweet Sentiment Extraction/Data/complete_64_clean/fold_3.tar.gz'
# !tar -xf '/content/drive/My Drive/Colab Notebooks/Tweet Sentiment Extraction/Data/complete_64_clean/fold_4.tar.gz'
# !tar -xf '/content/drive/My Drive/Colab Notebooks/Tweet Sentiment Extraction/Data/complete_64_clean/fold_5.tar.gz'
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _kg_hide-input=true _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" id="hVhA98wenvrG" colab_type="code" outputId="4ecf2a02-cbb7-499a-8e5a-87a05c15f970" executionInfo={"status": "ok", "timestamp": 1590678723830, "user_tz": 180, "elapsed": 74137, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjWI4QgYDUyC8ZcW5KHpuBwcY-MZWVqbPWcZSzxIg=s64", "userId": "06256612867315483887"}} colab={"base_uri": "https://localhost:8080/", "height": 568}
database_base_path = COLAB_BASE_PATH + 'Data/complete_64_clean/'
k_fold = pd.read_csv(database_base_path + '5-fold.csv')
print(f'Training samples: {len(k_fold)}')
display(k_fold.head())
# + [markdown] id="b0bUTMPynvrM" colab_type="text"
# # Model parameters
# + id="Paexn4ywnvrM" colab_type="code" colab={}
vocab_path = COLAB_BASE_PATH + 'qa-transformers/roberta/roberta-base-vocab.json'
merges_path = COLAB_BASE_PATH + 'qa-transformers/roberta/roberta-base-merges.txt'
base_path = COLAB_BASE_PATH + 'qa-transformers/roberta/'
config = {
"MAX_LEN": 64,
"BATCH_SIZE": 32,
"EPOCHS": 5,
"LEARNING_RATE": 3e-5,
"ES_PATIENCE": 1,
"N_FOLDS": 5,
"question_size": 4,
"base_model_path": base_path + 'roberta-base-tf_model.h5',
"config_path": base_path + 'roberta-base-config.json'
}
with open(MODEL_BASE_PATH + 'config.json', 'w') as json_file:
json.dump(json.loads(json.dumps(config)), json_file)
# + [markdown] id="D_8F5nxDnvrW" colab_type="text"
# # Tokenizer
# + _kg_hide-output=true id="6qeQlxrDnvrX" colab_type="code" colab={}
tokenizer = ByteLevelBPETokenizer(vocab_file=vocab_path, merges_file=merges_path,
lowercase=True, add_prefix_space=True)
# + [markdown] id="S3FN1pb1nvrb" colab_type="text"
# ## Learning rate schedule
# + _kg_hide-input=true id="XUYfnwPRnvrc" colab_type="code" outputId="620d0bb5-3238-4d2e-94d8-5ddef543a3c4" executionInfo={"status": "ok", "timestamp": 1590678734112, "user_tz": 180, "elapsed": 84401, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjWI4QgYDUyC8ZcW5KHpuBwcY-MZWVqbPWcZSzxIg=s64", "userId": "06256612867315483887"}} colab={"base_uri": "https://localhost:8080/", "height": 440}
LR_MIN = 1e-6
LR_MAX = config['LEARNING_RATE']
LR_EXP_DECAY = .5
@tf.function
def lrfn(epoch):
lr = LR_MAX * LR_EXP_DECAY**epoch
if lr < LR_MIN:
lr = LR_MIN
return lr
rng = [i for i in range(config['EPOCHS'])]
y = [lrfn(x) for x in rng]
fig, ax = plt.subplots(figsize=(20, 6))
plt.plot(rng, y)
print("Learning rate schedule: {:.3g} to {:.3g} to {:.3g}".format(y[0], max(y), y[-1]))
# + [markdown] id="3aHwZDF9nvrR" colab_type="text"
# # Model
# + id="7gIO5wkQnvrS" colab_type="code" colab={}
module_config = RobertaConfig.from_pretrained(config['config_path'], output_hidden_states=False)
def model_fn(MAX_LEN):
input_ids = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='input_ids')
attention_mask = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='attention_mask')
base_model = TFRobertaModel.from_pretrained(config['base_model_path'], config=module_config, name="base_model")
sequence_output = base_model({'input_ids': input_ids, 'attention_mask': attention_mask})
last_state = sequence_output[0]
x_start = layers.Dropout(.1)(last_state)
x_start = layers.Dense(1)(x_start)
x_start = layers.Flatten()(x_start)
y_start = layers.Activation('softmax', name='y_start')(x_start)
x_end = layers.Dropout(.1)(last_state)
x_end = layers.Dense(1)(x_end)
x_end = layers.Flatten()(x_end)
y_end = layers.Activation('softmax', name='y_end')(x_end)
model = Model(inputs=[input_ids, attention_mask], outputs=[y_start, y_end])
return model
# + [markdown] id="jy24FlCgnvrj" colab_type="text"
# # Train
# + _kg_hide-input=true _kg_hide-output=true id="RU2sa648nvrj" colab_type="code" outputId="fbfd63a0-6b95-41ba-dc6d-05586e9db169" executionInfo={"status": "ok", "timestamp": 1590682960578, "user_tz": 180, "elapsed": 4310845, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjWI4QgYDUyC8ZcW5KHpuBwcY-MZWVqbPWcZSzxIg=s64", "userId": "06256612867315483887"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
AUTO = tf.data.experimental.AUTOTUNE
history_list = []
for n_fold in range(config['N_FOLDS']):
n_fold +=1
print('\nFOLD: %d' % (n_fold))
# Load data
base_data_path = 'fold_%d/' % (n_fold)
x_train = np.load(base_data_path + 'x_train.npy')
y_train = np.load(base_data_path + 'y_train.npy')
x_valid = np.load(base_data_path + 'x_valid.npy')
y_valid = np.load(base_data_path + 'y_valid.npy')
step_size = x_train.shape[1] // config['BATCH_SIZE']
valid_step_size = x_valid.shape[1] // config['BATCH_SIZE']
# Train model
model_path = 'model_fold_%d.h5' % (n_fold)
model = model_fn(config['MAX_LEN'])
optimizer = optimizers.Adam(lr=config['LEARNING_RATE'])
model.compile(optimizer, loss={'y_start': losses.CategoricalCrossentropy(label_smoothing=0.2),
'y_end': losses.CategoricalCrossentropy(label_smoothing=0.2)})
es = EarlyStopping(monitor='val_loss', mode='min', patience=config['ES_PATIENCE'],
restore_best_weights=True, verbose=1)
checkpoint = ModelCheckpoint((MODEL_BASE_PATH + model_path), monitor='val_loss', mode='min',
save_best_only=True, save_weights_only=True)
lr_schedule = LearningRateScheduler(lrfn)
history = model.fit(list(x_train), list(y_train),
validation_data=(list(x_valid), list(y_valid)),
batch_size=config['BATCH_SIZE'],
callbacks=[checkpoint, es, lr_schedule],
epochs=config['EPOCHS'],
verbose=2).history
history_list.append(history)
# Make predictions
# model.load_weights(MODEL_BASE_PATH + model_path)
predict_eval_df(k_fold, model, x_train, x_valid, get_test_dataset, decode, n_fold, tokenizer, config, config['question_size'])
# + [markdown] id="_59x5WnGnvro" colab_type="text"
# # Model loss graph
# + _kg_hide-input=true id="Tlq60Zvenvro" colab_type="code" cellView="form" outputId="b15dad8a-84a6-4604-a9b0-de39bbe10ff3" executionInfo={"status": "ok", "timestamp": 1590682963986, "user_tz": 180, "elapsed": 4314188, "user": {"displayName": "<NAME>", "photoUrl": "https://<KEY>", "userId": "06256612867315483887"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
#@title
for n_fold in range(config['N_FOLDS']):
print('Fold: %d' % (n_fold+1))
plot_metrics(history_list[n_fold])
# + [markdown] id="xxbg-_R_p2dE" colab_type="text"
# # Model evaluation
# + id="KgsapEFup3zY" colab_type="code" cellView="form" outputId="8eff0ff1-7130-45ff-ad7a-06526a2be492" executionInfo={"status": "ok", "timestamp": 1590682967566, "user_tz": 180, "elapsed": 4317762, "user": {"displayName": "<NAME>", "photoUrl": "https://<KEY>", "userId": "06256612867315483887"}} colab={"base_uri": "https://localhost:8080/", "height": 302}
#@title
display(evaluate_model_kfold(k_fold, config['N_FOLDS']).style.applymap(color_map))
# + [markdown] id="yP3xG-Cenvry" colab_type="text"
# # Visualize predictions
# + _kg_hide-input=true id="cGxcMVRMnvrz" colab_type="code" cellView="form" outputId="a83b6711-073b-4056-cc70-ef5a2ccd6e65" executionInfo={"status": "ok", "timestamp": 1590682967567, "user_tz": 180, "elapsed": 4317755, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjWI4QgYDUyC8ZcW5KHpuBwcY-MZWVqbPWcZSzxIg=s64", "userId": "06256612867315483887"}} colab={"base_uri": "https://localhost:8080/", "height": 581}
#@title
k_fold['jaccard_mean'] = (k_fold['jaccard_fold_1'] + k_fold['jaccard_fold_2'] +
k_fold['jaccard_fold_3'] + k_fold['jaccard_fold_4'] +
k_fold['jaccard_fold_4']) / 5
display(k_fold[['text', 'selected_text', 'sentiment', 'text_tokenCnt',
'selected_text_tokenCnt', 'jaccard', 'jaccard_mean']].head(15))
| Model backlog/Train/205-Tweet-Train-5Fold-roBERTa 91.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,jl:hydrogen
# text_representation:
# extension: .jl
# format_name: hydrogen
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.8.0-DEV
# language: julia
# name: julia-1.8
# ---
# %% [markdown]
# Since there is no minimal working example (MWE), I could not figure out what you really wanted to do.
# (Please read https://discourse.julialang.org/t/please-read-make-it-easier-to-help-you/14757)
#
# However, I thought that possibly you would like to do something like this. Please take a look at the MWE below.
# %%
module SCI
export Problem, Algorithm_A, Algorithm_B, solve
abstract type AbstractProblem end
struct Problem{T} <: AbstractProblem x::T end
abstract type AbstractAlgorithm end
Base.@kwdef struct Algorithm_A{T} <: AbstractAlgorithm a::T = 2.0 end
Base.@kwdef struct Algorithm_B{T} <: AbstractAlgorithm a::T = 3.0 end
default_algorithm(prob::Problem) = Algorithm_A()
struct Solution{R, P<:AbstractProblem, A<:AbstractAlgorithm} result::R; prob::P; alg::A end
solve(prob::AbstractProblem) = solve(prob, default_algorithm(prob))
solve(prob::AbstractProblem, alg::AbstractAlgorithm) = Solution(alg.a * prob.x, prob, alg)
"""
Here, `Base` module plays the role of `COM` package.
SCI module can define the method of
`Base.show` = `COM.show` function for `SCI.Solution` type.
"""
function Base.show(io::IO, sol::Solution)
result = """
Problem: $(sol.prob)
Algorithm: $(sol.alg)
Result: $(sol.result)
"""
print(io, result)
end
end
using .SCI
@show prob = Problem(1.5)
println()
println(solve(prob))
println(solve(prob, Algorithm_B()))
# %%
module SCI2 # extension of SCI module
export Problem2, Algorithm_C
using ..SCI: SCI, AbstractProblem, AbstractAlgorithm, Solution
struct Problem2{T} <: AbstractProblem x::T; y::T end
Base.@kwdef struct Algorithm_C{T} <: AbstractAlgorithm a::T = 2.0; b = 3.0 end
SCI.default_algorithm(prob::Problem2) = Algorithm_C()
SCI.solve(prob::Problem2, alg::Algorithm_C) = Solution(alg.a * prob.x + alg.b * prob.y, prob, alg)
end
using .SCI2
@show prob2 = Problem2(10.0, 1.0)
println()
println(solve(prob2))
# %% [markdown]
# In the above example I don't define `AbstractSolution` type, nor did I make `Solution` a subtype of it. But you can define it, and then define another Solution type as a subtype of it in `SCI2` module, and also define how to display objects of another Solution type. Many other extensions are also possible.
#
# See also https://discourse.julialang.org/t/function-depending-on-the-global-variable-inside-module/64322/10
# %%
| 0011/Handling of package inter-dependency.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# # Question 5d
# +
# Create x, y points
num_points = 5000
X, Y = np.random.uniform(low=-1.0, high=1.0, size=(num_points)), np.random.uniform(low=-1.0, high=1.0, size=(num_points))
# Create figure
fig, ax = plt.subplots(1)
ax.scatter(X, Y, marker='.', s=1)
ax.add_patch(plt.Circle((0, 0), 1, color='b', fill=False))
ax.axis('equal')
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_title('Sampling (X, Y) Points in Circle')
plt.savefig('Sampling (X, Y) Points in Circle.pdf')
# -
# # Question 5e
# +
# Create x, y points
num_points = 5000
R, T = np.random.uniform(low=0, high=1.0, size=(num_points)), np.random.uniform(low=0, high=2*np.pi, size=(num_points))
X, Y = R * np.cos(T), R * np.sin(T)
# Create figure
fig, ax = plt.subplots(1)
ax.scatter(X, Y, marker='.', s=1)
ax.add_patch(plt.Circle((0, 0), 1, color='b', fill=False))
ax.axis('equal')
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_title('Sampling (R, T) Points in Circle')
plt.savefig('Sampling (R, T) Points in Circle.pdf')
# -
# # Question 6a
#
# The running average appears to converge to 0 as n grows.
# Create X points and calculate running average
num_points = 10000
mean = 0
std_dev = 1
X = np.random.normal(loc=mean, scale=std_dev, size=(num_points))
X_running_average = np.zeros_like(X)
for i in range(len(X_running_average)):
X_running_average[i] = np.sum(X[:(i+1)]) / len(X[:(i+1)])
# Create figure
fig, ax = plt.subplots(1)
ax.plot(np.arange(len(X_running_average)), X_running_average)
ax.set_xlabel('n')
ax.set_ylabel('Normal Running Average')
ax.set_title('Normal Running Average vs n')
plt.savefig('Normal Running Average vs n.pdf')
# # Question 6b
#
# The running average does not appear to converge to 0, and does not seem to converge at all. The Cauchy distribution occassionally has enormous numbers that occur frequently enough to dominate the expectation.
# Create X points and calculate running average
from tqdm import tqdm
num_points = 10000
Y = np.random.uniform(low=-np.pi/2, high=np.pi/2, size=(num_points))
X = np.tan(Y)
X_running_average = np.zeros_like(X)
for i in tqdm(range(len(X_running_average))):
X_running_average[i] = np.sum(X[:(i+1)]) / len(X[:(i+1)])
# Create figure
fig, ax = plt.subplots(1)
ax.plot(np.arange(len(X_running_average)), X_running_average)
ax.set_xlabel('n')
ax.set_ylabel('Cauchy Running Average')
ax.set_title('Cauchy Running Average vs n')
plt.savefig('Cauchy Running Average vs n.pdf')
| Assignment 4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.9 64-bit (''learn-env'': conda)'
# language: python
# name: python36964bitlearnenvcondae7e6328cec2744cc9785efcdf88db667
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sam-thurman/asl_alphabet_image_classification/blob/master/notebooks/model_create/edge_model_create.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="L_Me7KhpRsz7" colab_type="code" outputId="b1d4a362-d871-4cb8-9323-c1953aa07de5" colab={"base_uri": "https://localhost:8080/", "height": 122}
from google.colab import drive
drive.mount('/content/drive')
# + id="n4fX4yD_Ru_f" colab_type="code" outputId="5d0bb3a0-cf02-4481-cb61-125a98b17c98" colab={"base_uri": "https://localhost:8080/", "height": 34}
# %cd drive/My\ Drive/
# + id="pEjiAn1IReDV" colab_type="code" colab={}
import os
import pathlib
import tarfile
import boto3
from botocore.handlers import disable_signing
from scipy.io import loadmat
from imageio import imread
import numpy as np
from skimage.transform import resize
def list_files(base_path, validExts=(".jpg", ".jpeg", ".png", ".bmp"), contains=None):
# loop over the directory structure
for (rootDir, dirNames, filenames) in os.walk(base_path):
# loop over the filenames in the current directory
for filename in filenames:
# if the contains string is not none and the filename does not contain
# the supplied string, then ignore the file
if contains is not None and filename.find(contains) == -1:
continue
# determine the file extension of the current file
ext = filename[filename.rfind("."):].lower()
# check to see if the file is an image and should be processed
if ext.endswith(validExts):
# construct the path to the image and yield it
imagePath = os.path.join(rootDir, filename).replace(" ", "\\ ")
yield imagePath
class BSDS500(object):
BUCKET = 'i008data'
FN = 'BSR_bsds500.tgz'
STORE_FN = os.path.join(os.environ['HOME'], 'BSR.tgz')
HOME = os.environ['HOME']
def __init__(self, path_to_bsds=None, images_to_gray=False, target_size=None, masks_to_binary=True):
if not path_to_bsds:
self.BSDS_BASE = self.get_bsds()
else:
self.BSDS_BASE = path_to_bsds
print(self.BSDS_BASE)
self.images_to_gray = images_to_gray
self.target_size = target_size
self.masks_to_binary = masks_to_binary
self.TRAIN_PATH = os.path.join(self.BSDS_BASE, 'BSDS500/data/images/train/')
self.TEST_PATH = os.path.join(self.BSDS_BASE, 'BSDS500/data/images/test/')
self.VALID_PATH = os.path.join(self.BSDS_BASE, 'BSDS500/data/images/val/')
self.GROUND_TRUTH_TRAIN = os.path.join(self.BSDS_BASE, 'BSDS500/data/groundTruth/train/')
self.GROUND_TRUTH_TEST = os.path.join(self.BSDS_BASE, 'BSDS500/data/groundTruth/test/')
self.GROUND_TRUTH_VALID = os.path.join(self.BSDS_BASE, 'BSDS500/data/groundTruth/val/')
def get_bsds(self):
if not pathlib.Path(self.STORE_FN).exists():
print("DOWNLOADING BSDS500 DATA BE PATIENT")
s3_resource = boto3.resource('s3')
s3_resource.meta.client.meta.events.register('choose-signer.s3.*', disable_signing)
bucket = s3_resource.Bucket(self.BUCKET)
bucket.download_file(self.FN, self.STORE_FN)
ds_dir = self.STORE_FN.split('.')[0]
if not pathlib.Path(ds_dir).is_dir():
tar = tarfile.open(self.STORE_FN)
tar.extractall(self.HOME)
# dir_path = os.path.dirname(os.path.realpath(__file__))
dir_path = 'bsds500/data'
return os.path.join(dir_path, self.STORE_FN.split('.')[0])
def load_ground_truth(self, gt_path):
ground_truth_paths = sorted(list(list_files(gt_path, validExts=('.mat'))))
file_id = []
cnts = []
sgmnts = []
for gt_path in ground_truth_paths:
file_name = os.path.basename(gt_path).split('.')[0]
gt = loadmat(gt_path)
gt = gt['groundTruth'][0]
for annotator in gt:
contours = annotator[0][0][1] # 1-> contours
segments = annotator[0][0][0] # 0 -> segments
if self.target_size:
contours = resize(contours.astype(float), output_shape=self.target_size)
segments = resize(segments, output_shape=self.target_size)
if self.masks_to_binary:
contours[contours > 0] = 1
file_id.append(file_name)
cnts.append(contours)
sgmnts.append(segments)
cnts = np.concatenate([np.expand_dims(a, 0) for a in cnts])
sgmnts = np.concatenate([np.expand_dims(a, 0) for a in sgmnts])
cnts = cnts[..., np.newaxis]
sgmnts = sgmnts[..., np.newaxis]
return file_id, cnts, sgmnts
def load_images(self, list_of_files):
processed_images = []
for i, f in enumerate(list_of_files):
if self.images_to_gray:
im = imread(f, mode='L')
else:
im = imread(f)
if self.target_size:
im = resize(im, output_shape=self.target_size)
processed_images.append(np.expand_dims(im, 0))
processed_images = np.concatenate(processed_images)
if self.images_to_gray:
processed_images = processed_images[..., np.newaxis]
return processed_images
def get_train(self):
file_ids, cnts, sgmnts = self.load_ground_truth(self.GROUND_TRUTH_TRAIN)
image_paths = [self.TRAIN_PATH + f_id + '.jpg' for f_id in file_ids]
images = self.load_images(image_paths)
return file_ids, cnts, sgmnts, images
def get_test(self):
file_ids, cnts, sgmnts = self.load_ground_truth(self.GROUND_TRUTH_TEST)
image_paths = [self.TEST_PATH + f_id + '.jpg' for f_id in file_ids]
images = self.load_images(image_paths)
return file_ids, cnts, sgmnts, images
def get_val(self):
file_ids, cnts, sgmnts = self.load_ground_truth(self.GROUND_TRUTH_VALID)
image_paths = [self.VALID_PATH + f_id + '.jpg' for f_id in file_ids]
images = self.load_images(image_paths)
return file_ids, cnts, sgmnts, images
# + id="6gM_eUUrRFT5" colab_type="code" colab={}
import pandas as pd
import numpy as np
import scipy as math
import matplotlib.pyplot as plt
import seaborn as sns
import pickle
# from PIL import Image
from sklearn.model_selection import train_test_split
import keras
from keras.preprocessing.image import ImageDataGenerator, array_to_img, img_to_array, load_img
from keras.models import Model, Sequential
from keras import layers
from keras.layers import Dense, Input, Dropout, GlobalAveragePooling2D, Flatten, Conv2D, BatchNormalization, Activation, MaxPooling2D, concatenate, UpSampling2D
from keras import optimizers
from keras.optimizers import Adam
import sys
import os
# from bsds500 import BSDS500
from keras.callbacks import CSVLogger
# + id="K7l9f1vzRFT-" colab_type="code" colab={}
# + id="_YZqwQr_RFUB" colab_type="code" colab={}
# + id="3K9zWHluRFUD" colab_type="code" outputId="a37f1794-b313-4b0f-cd53-626b9a0a7bb2" colab={"base_uri": "https://localhost:8080/", "height": 34}
TARGET_SHAPE = (192, 192)
bsds = BSDS500(target_size=TARGET_SHAPE)
ids, contours_train, sgmnts, train_images = bsds.get_train()
ids, contours_test, sgmnts, test_images = bsds.get_train()
C = np.concatenate([contours_test, contours_train])
I = np.concatenate([test_images, train_images])
# + id="x6LCCHyVRFUH" colab_type="code" colab={}
def get_unet(img_dim, channels):
inputs = Input((img_dim, img_dim, channels))
conv1 = Conv2D(32, (3, 3), activation='relu', padding='same')(inputs)
conv1 = Conv2D(32, (3, 3), activation='relu', padding='same')(conv1)
pool1 = MaxPooling2D(pool_size=(2, 2))(conv1)
conv2 = Conv2D(64, (3, 3), activation='relu', padding='same')(pool1)
conv2 = Conv2D(64, (3, 3), activation='relu', padding='same')(conv2)
pool2 = MaxPooling2D(pool_size=(2, 2))(conv2)
conv3 = Conv2D(128, (3, 3), activation='relu', padding='same')(pool2)
conv3 = Conv2D(128, (3, 3), activation='relu', padding='same')(conv3)
pool3 = MaxPooling2D(pool_size=(2, 2))(conv3)
conv4 = Conv2D(256, (3, 3), activation='relu', padding='same')(pool3)
conv4 = Conv2D(256, (3, 3), activation='relu', padding='same')(conv4)
pool4 = MaxPooling2D(pool_size=(2, 2))(conv4)
conv5 = Conv2D(512, (3, 3), activation='relu', padding='same')(pool4)
conv5 = Conv2D(512, (3, 3), activation='relu', padding='same')(conv5)
up6 = UpSampling2D(size=(2, 2))(conv5)
conv6 = Conv2D(256, (3, 3), activation='relu', padding='same')(up6)
conv6 = Conv2D(256, (3, 3), activation='relu', padding='same')(conv6)
up7 = UpSampling2D(size=(2, 2))(conv6)
conv7 = Conv2D(128, (3, 3), activation='relu', padding='same')(up7)
conv7 = Conv2D(128, (3, 3), activation='relu', padding='same')(conv7)
up8 = UpSampling2D(size=(2, 2))(conv7)
conv8 = Conv2D(64, (3, 3), activation='relu', padding='same')(up8)
conv8 = Conv2D(64, (3, 3), activation='relu', padding='same')(conv8)
up9 = UpSampling2D(size=(2, 2))(conv8)
conv9 = Conv2D(32, (3, 3), activation='relu', padding='same')(up9)
conv9 = Conv2D(32, (3, 3), activation='relu', padding='same')(conv9)
conv10 = Conv2D(1, (1, 1), activation='sigmoid')(conv9)
model = Model(inputs=[inputs], outputs=[conv10])
model.compile(optimizer=Adam(lr=1e-4), loss='binary_crossentropy')
return model
# + id="sRnWFeB2RFUJ" colab_type="code" outputId="29c738f4-6e60-4c74-b719-23270b7bb17d" colab={"base_uri": "https://localhost:8080/", "height": 34}
I[0].shape, C[0].shape
# + id="ulhkGHZ8RFUM" colab_type="code" outputId="07325d4f-da7d-4bcc-9995-7623f208cc7d" colab={"base_uri": "https://localhost:8080/", "height": 1000}
csv_callback = CSVLogger('history.log', append=True)
unet = get_unet(192, 3)
history = unet.fit(I, C, verbose=2, epochs=200, validation_split=0.1, callbacks=[csv_callback])
# + id="QPsaRLZ7VkiC" colab_type="code" colab={}
unet.save('unet.h5')
# + id="t7vxg06EiVD8" colab_type="code" outputId="fc6fea57-0592-4478-de4d-68f4a2b0baa7" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# %ls
# + id="XjCDvjXMRFUP" colab_type="code" colab={}
del unet
# + id="4hN9lbSTVxrr" colab_type="code" colab={}
from keras.models import load_model
unet = load_model('unet.h5')
# + id="BSQ7jPvYiYUG" colab_type="code" colab={}
del unet
# + id="PGlcu8BbihDZ" colab_type="code" outputId="d9d48496-ea42-4b55-cf5c-c2f21b0de595" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# %ls
# + id="yoP9RpFHi6xY" colab_type="code" colab={}
| notebooks/model_create/edge_model_create.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Classification and Segmentation
#
# ## CS194-26 Project 4, Spring 2020
#
# ### by <NAME>
#
# #### <EMAIL> -- cs194-ahb
# +
from IPython.core.display import HTML
HTML("""
<style>
div.cell { /* Tunes the space between cells */
margin-top:1em;
margin-bottom:1em;
}
div.text_cell_render h1 { /* Main titles bigger, centered */
font-size: 2.2em;
line-height:0.9em;
}
div.text_cell_render h2 { /* Parts names nearer from text */
margin-bottom: -0.4em;
}
div.text_cell_render { /* Customize text cells */
font-family: 'Georgia';
font-size:1.2em;
line-height:1.4em;
padding-left:3em;
padding-right:3em;
}
.output_png {
display: table-cell;
text-align: center;
vertical-align: middle;
}
</style>
<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide();
} else {
$('div.input').show();
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
""")
# -
# ## Part 1 – Image Classification
# The first part of this project is an image classifier for the Fashion-MNIST dataset, built in PyTorch. The network I created for classification is a 5 layer network, with 2 Convolution layers and 3 Fully Connected layers. The architecture resembles the following:
#
# $$\big(\text{Conv} \longrightarrow \text{ReLU} \longrightarrow \text{MaxPool}\big) \times 2 \longrightarrow \big(\text{FC} \longrightarrow \text{ReLU}\big) \times 2 \longrightarrow \text{FC}$$
#
# The parameters for the Conv and FC layers can be seen below.
# +
import torch
import torchvision
import torchvision.transforms as transforms
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import numpy as np
import matplotlib.pyplot as plt
# +
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
# 1 input image channel, 6 output channels, 3x3 square convolution
# kernel
self.conv1 = nn.Conv2d(1, 32, 3)
self.conv2 = nn.Conv2d(32, 32, 3)
# an affine operation: y = Wx + b
self.fc1 = nn.Linear(32 * 5 * 5, 120) # 6*6 from image dimension
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
def forward(self, x):
# Max pooling over a (2, 2) window
x = F.max_pool2d(F.relu(self.conv1(x)), 2)
# If the size is a square you can only specify a single number
x = F.max_pool2d(F.relu(self.conv2(x)), 2)
x = x.view(-1, np.prod(x.size()[1:]))
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
net = Net()
print(net)
# +
trainset = torchvision.datasets.FashionMNIST(root='./data', train=True,
download=True, transform=transforms.ToTensor())
train_size = int(0.8*len(trainset))
train_set, val_set = torch.utils.data.random_split(trainset, [train_size, len(trainset)-train_size])
trainloader = torch.utils.data.DataLoader(train_set, batch_size=50,
shuffle=True, num_workers=2)
valloader = torch.utils.data.DataLoader(val_set, batch_size=50,
shuffle=True, num_workers=2)
testset = torchvision.datasets.FashionMNIST(root='./data', train=False,
download=True, transform=transforms.ToTensor())
testloader = torch.utils.data.DataLoader(testset, batch_size=50,
shuffle=False, num_workers=2)
classes = {v: k for k, v in trainset.class_to_idx.items()}
# -
# A quick visualization of our dataset, with the class labels:
dataiter = iter(trainloader)
images, labels = dataiter.next()
plt.figure(figsize=(15,10))
plt.imshow(np.transpose(torchvision.utils.make_grid(images[:4,...]).numpy(), (1, 2, 0)))
plt.xlabel(' | '.join('%5s' % classes[labels[j].item()] for j in range(4)), size=12)
plt.xticks([30*i for i in range(5)])
plt.yticks([])
plt.show()
# +
# Run this code to train. To load, run the loading code below.
num_epochs = 10
optimizer = optim.Adam(net.parameters(), lr=0.001)
criterion = nn.CrossEntropyLoss()
train_acc = np.zeros(num_epochs)
val_acc = np.zeros(num_epochs)
for epoch in range(num_epochs):
running_loss = 0
for i, (images, labels) in enumerate(trainloader):
optimizer.zero_grad() # zero the gradient buffers
output = net(images)
loss = criterion(output, labels)
loss.backward()
optimizer.step() # Does the update
running_loss += loss.item()
if i % 100 == 0:
print ('Iteration %05d Train loss %f' % (i, running_loss/100), '\r', end='')
running_loss=0
with torch.no_grad():
total = 0
correct = 0
for i, (images, labels) in enumerate(trainloader):
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
train_acc[epoch] = correct/total*100
total = 0
correct = 0
for i, (images, labels) in enumerate(valloader):
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
val_acc[epoch] = correct/total*100
# +
# Saving data
# torch.save(net.state_dict(), './mnist_net.pth') # Saving state dict
# np.save("train_acc", train_acc) # Saving train acc
# np.save("val_acc", val_acc) # Saving train acc
# -
# Loading data
net = Net()
net.load_state_dict(torch.load('./mnist_net.pth'))
train_acc = np.load("train_acc.npy")
val_acc = np.load("val_acc.npy")
# A plot of the training and validation accuracy during training (calculated very epoch). As we can see in the plot below, we overfit a bit to the training data, but the network still performs well on the validation set.
plt.figure(figsize=(15,10))
plt.plot(range(1,11),train_acc, label="train acc", marker="o")
plt.plot(range(1,11), val_acc, label="val acc", marker="o")
plt.xlabel("Number of Epochs")
plt.ylabel("Accuracy (%)")
plt.title("Accuracy vs. Epochs")
plt.legend()
plt.show()
correct = 0
total = 0
with torch.no_grad():
for (images, labels) in testloader:
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
# Accuracy of the network on the test set:
print("Accuracy of the network on the {0} test images: {1:.2f} %".format(total,100 * correct / total))
# +
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
with torch.no_grad():
for data in testloader:
images, labels = data
outputs = net(images)
_, predicted = torch.max(outputs, 1)
c = (predicted == labels).squeeze()
for i in range(50):
label = labels[i]
class_correct[label] += c[i].item()
class_total[label] += 1
# -
print("Per Class Accuracy")
print("-"*25)
for i in range(10):
print("{0:^11s} : {1:.2f} %".format(classes[i], 100 * class_correct[i] / class_total[i]))
# As we can see in the Per Class Accuracy table above, the network performs extremely well for the Bag and Ankle Boot classes, but not as well for the Shirt and Coat classes. This is probably due to the high variance in the input dataset for the poorly performing classes.
# +
correct_ex = np.zeros((10, 2, 1, 28, 28))
correct_ex_tally = np.zeros(10, dtype=int)
incorrect_ex = np.zeros((10, 2, 1, 28, 28))
incorrect_ex_tally = np.zeros(10, dtype=int)
with torch.no_grad():
for data in testloader:
if all([tally==2 for tally in correct_ex_tally]) and all([tally==2 for tally in incorrect_ex_tally]):
break
images, labels = data
outputs = net(images)
_, predicted = torch.max(outputs, 1)
c = (predicted == labels).squeeze()
for i in range(50):
label = labels[i]
classified_correct = c[i].item()
if classified_correct and correct_ex_tally[label] != 2:
correct_ex[label, correct_ex_tally[label]] = images[i]
correct_ex_tally[label] += 1
elif not classified_correct and incorrect_ex_tally[label] != 2:
incorrect_ex[label, incorrect_ex_tally[label]] = images[i]
incorrect_ex_tally[label] += 1
if all([tally==2 for tally in correct_ex_tally]) and all([tally==2 for tally in incorrect_ex_tally]):
break
# -
# Below is a visualization of data points (images) that the network classifies correctly and incorrectly.
fig = plt.figure(figsize=(10,20))
fig.suptitle("Correctly and Incorrectly Classified Examples", fontsize=15, ha="center", va="center", y=0.93)
plt.figtext(0.32,0.9,"Correct Classification", va="center", ha="center", size=15)
plt.figtext(0.72,0.9,"Incorrect Classification", va="center", ha="center", size=15)
for class_no in range(10):
for i in range(2):
plt.subplot(10, 4, 4*class_no+i+1)
plt.imshow(correct_ex[class_no, i, 0], cmap="gray")
# plt.title("Correct Ex {}".format(i+1))
plt.xticks([])
plt.yticks([])
if i == 0:
plt.ylabel(classes[class_no])
for i in range(2):
plt.subplot(10, 4, 4*class_no+3+i)
plt.imshow(incorrect_ex[class_no, i, 0], cmap="gray")
# plt.title("Incorrect Ex {}".format(i+1))
plt.axis("off")
# Below is a visualization of the filters learned in the first Convolution layer of the network.
filters = np.transpose(net.conv1.weight.data, (0, 2, 3, 1))[...,0]
fig = plt.figure(figsize=(15,10))
fig.suptitle("Learned Filters for conv1", fontsize=16, y=0.93)
for i, filt in enumerate(filters, 1):
plt.subplot(4, 8, i)
plt.imshow(filt)
plt.xticks([])
plt.yticks([])
# plt.tight_layout()
# ## Part 2 – Image Segmentation
# Semantic Segmentation refers to labeling each pixel in the image to its correct object class. For this part, I used a Mini Facade dataset, which consists of images of different cities around the world and diverse architectural styles and their semantic segmentation labels in 5 different classes: balcony, window, pillar, facade and others. Using this dataset, I trained a deep neural network of my design to perform semantic segmentation.
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.insert(0, './part2')
import part2.dataset
import part2.train
# +
net = part2.train.Net()
net.load_state_dict(torch.load('./part2/models/model_starter_net.pth',map_location=torch.device('cpu')))
ap_data = part2.dataset.FacadeDataset(dataDir='part2/starter_set',flag='test_dev', data_range=(0,114), onehot=True)
ap_loader = torch.utils.data.DataLoader(ap_data, batch_size=1)
train_loss = np.load("part2/train_loss.npy")
val_loss = np.load("part2/val_loss.npy")
# -
# My network architecture is shown below. The general idea was to increase channel depth as I decrease the spatial dimension, and then to decrease channel depth as I increase the spatial dimensions.
#
# I used Cross Entropy Loss as my loss function, and the ADAM optimizer with a learning rate of $10^{-3}$ and a weight decay of $10^{-5}$ as my solver. I did a 90-10 split for my training-validation datasets. I trained my network for $15$ epochs and tuned my networks parameters based on the results on the validation set.
print(net)
# A plot of the training and validation loss during training (calculated very epoch) is shown below.
plt.figure(figsize=(15,10))
plt.plot(range(1,16),train_loss, label="train loss")
plt.plot(range(1,16), val_loss, label="val loss")
plt.xlabel("Number of Epochs")
plt.ylabel("Loss")
plt.title("Loss vs. Epochs")
plt.legend()
plt.show()
# The Average Precision, per class and the average across classes is below.
aps = part2.train.cal_AP(ap_loader, net, nn.CrossEntropyLoss(), "cpu")
print("Average AP across classes: {}".format(np.mean(aps)))
im_no = 65
folder = "part2/output_test/"
image_types = ["x", "gt", "y"]
images = [plt.imread(folder+im_t+str(im_no)+".png") for im_t in image_types]
# A sample data point (image) from the dataset, its ground truth, and the network output is shown below. As a reference, here is a table of the 5 classes and their corresponding colors: 
#
#
# As we can see, the network performs decently well for such a challenging task. It more or less classifies the broad regions correctly, but it is clear that the classification is very coarse.
#
# Additionally, it performs quite well for the facade and windows, but quite poorly for the pillars.
titles = ["Original Image", "Ground Truth Segmentation", "Network Output"]
plt.figure(figsize=(20,20))
for i, image in enumerate(images):
plt.subplot(1, 3, i+1)
plt.imshow(image)
plt.axis("off")
plt.title(titles[i])
| resources/cs194-projs/proj4_website/proj4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import print_function, division
import pandas as pd
import tabula
import os
from tabula import wrapper
# -
# ## View the raw data
# ! cd ../data/raw && ls -h
# ## Add the titles to a list fr later use
# +
directory_in_str = '../data/raw/'
directory = os.fsencode(directory_in_str)
pdfiles = []
for file in os.listdir(directory):
filename = os.fsdecode(file)
if filename.endswith(".pdf"):
pdfiles.append(filename)
else:
continue
# -
print(pdfiles)
# ## For Each File Convert it to CSV and save it to /data/interim
# print p just to see how far we are
for p in pdfiles:
print(p)
# df = wrapper.read_pdf('../data/raw/' + str(p), skip = 0, removeempty= 'FALSE', pages='all')
bp = wrapper.convert_into('../data/raw/' + str(p), '../data/interim/' + str(p) + ".csv", output_format="csv", removeempty= 'FALSE', pages="all")
# ## We now have CSVs of the files without the constituency information
#
# Instructions to complete dataset
# Extract the constituency name from each original pdf then for each empty line insert it
# OR add a new colomn named constituency and between each empty row insert the constituency name.
#
# This might get you started
#
# # !pdftotext 'NATIONAL ASSEMBLY MIDLANDS PROVINCE.pdf' - | grep 'Constituency' | sed -n 's/In respect of //p' | sed -n 's/the results are as follows://p' > const.txt
| notebooks/1-tjm-quickandclean-pdf2csv-moreworktobedone.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="../Pierian-Data-Logo.PNG">
# <br>
# <strong><center>Copyright 2019. Created by <NAME>.</center></strong>
# # Full Artificial Neural Network Code Along
# In the last section we took in four continuous variables (lengths) to perform a classification. In this section we'll combine continuous and categorical data to perform a regression. The goal is to estimate the cost of a New York City cab ride from several inputs. The inspiration behind this code along is a recent <a href='https://www.kaggle.com/c/new-york-city-taxi-fare-prediction'>Kaggle competition</a>.
#
# <div class="alert alert-success"><strong>NOTE:</strong> In this notebook we'll perform a regression with one output value. In the next one we'll perform a binary classification with two output values.</div>
#
# ## Working with tabular data
# Deep learning with neural networks is often associated with sophisticated image recognition, and in upcoming sections we'll train models based on properties like pixels patterns and colors.
#
# Here we're working with tabular data (spreadsheets, SQL tables, etc.) with columns of values that may or may not be relevant. As it happens, neural networks can learn to make connections we probably wouldn't have developed on our own. However, to do this we have to handle categorical values separately from continuous ones. Make sure to watch the theory lectures! You'll want to be comfortable with:
# * continuous vs. categorical values
# * embeddings
# * batch normalization
# * dropout layers
# ## Perform standard imports
# +
import torch
import torch.nn as nn
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ## Load the NYC Taxi Fares dataset
# The <a href='https://www.kaggle.com/c/new-york-city-taxi-fare-prediction'>Kaggle competition</a> provides a dataset with about 55 million records. The data contains only the pickup date & time, the latitude & longitude (GPS coordinates) of the pickup and dropoff locations, and the number of passengers. It is up to the contest participant to extract any further information. For instance, does the time of day matter? The day of the week? How do we determine the distance traveled from pairs of GPS coordinates?
#
# For this exercise we've whittled the dataset down to just 120,000 records from April 11 to April 24, 2010. The records are randomly sorted. We'll show how to calculate distance from GPS coordinates, and how to create a pandas datatime object from a text column. This will let us quickly get information like day of the week, am vs. pm, etc.
#
# Let's get started!
df = pd.read_csv('../Data/NYCTaxiFares.csv')
df.head()
df['fare_amount'].describe()
# From this we see that fares range from \\$2.50 to \\$49.90, with a mean of \\$10.04 and a median of \\$7.70
# ## Calculate the distance traveled
# The <a href='https://en.wikipedia.org/wiki/Haversine_formula'>haversine formula</a> calculates the distance on a sphere between two sets of GPS coordinates.<br>
# Here we assign latitude values with $\varphi$ (phi) and longitude with $\lambda$ (lambda).
#
# The distance formula works out to
#
# ${\displaystyle d=2r\arcsin \left({\sqrt {\sin ^{2}\left({\frac {\varphi _{2}-\varphi _{1}}{2}}\right)+\cos(\varphi _{1})\:\cos(\varphi _{2})\:\sin ^{2}\left({\frac {\lambda _{2}-\lambda _{1}}{2}}\right)}}\right)}$
#
# where
#
# $\begin{split} r&: \textrm {radius of the sphere (Earth's radius averages 6371 km)}\\
# \varphi_1, \varphi_2&: \textrm {latitudes of point 1 and point 2}\\
# \lambda_1, \lambda_2&: \textrm {longitudes of point 1 and point 2}\end{split}$
def haversine_distance(df, lat1, long1, lat2, long2):
"""
Calculates the haversine distance between 2 sets of GPS coordinates in df
"""
r = 6371 # average radius of Earth in kilometers
phi1 = np.radians(df[lat1])
phi2 = np.radians(df[lat2])
delta_phi = np.radians(df[lat2]-df[lat1])
delta_lambda = np.radians(df[long2]-df[long1])
a = np.sin(delta_phi/2)**2 + np.cos(phi1) * np.cos(phi2) * np.sin(delta_lambda/2)**2
c = 2 * np.arctan2(np.sqrt(a), np.sqrt(1-a))
d = (r * c) # in kilometers
return d
df['dist_km'] = haversine_distance(df,'pickup_latitude', 'pickup_longitude', 'dropoff_latitude', 'dropoff_longitude')
df.head()
# ## Add a datetime column and derive useful statistics
# By creating a datetime object, we can extract information like "day of the week", "am vs. pm" etc.
# Note that the data was saved in UTC time. Our data falls in April of 2010 which occurred during Daylight Savings Time in New York. For that reason, we'll make an adjustment to EDT using UTC-4 (subtracting four hours).
df['EDTdate'] = pd.to_datetime(df['pickup_datetime'].str[:19]) - pd.Timedelta(hours=4)
df['Hour'] = df['EDTdate'].dt.hour
df['AMorPM'] = np.where(df['Hour']<12,'am','pm')
df['Weekday'] = df['EDTdate'].dt.strftime("%a")
df.head()
df['EDTdate'].min()
df['EDTdate'].max()
# ## Separate categorical from continuous columns
df.columns
cat_cols = ['Hour', 'AMorPM', 'Weekday']
cont_cols = ['pickup_latitude', 'pickup_longitude', 'dropoff_latitude', 'dropoff_longitude', 'passenger_count', 'dist_km']
y_col = ['fare_amount'] # this column contains the labels
# <div class="alert alert-info"><strong>NOTE:</strong> If you plan to use all of the columns in the data table, there's a shortcut to grab the remaining continuous columns:<br>
# <pre style='background-color:rgb(217,237,247)'>cont_cols = [col for col in df.columns if col not in cat_cols + y_col]</pre>
#
# Here we entered the continuous columns explicitly because there are columns we're not running through the model (pickup_datetime and EDTdate)</div>
#
# ## Categorify
# Pandas offers a <a href='https://pandas.pydata.org/pandas-docs/stable/user_guide/categorical.html'><strong>category dtype</strong></a> for converting categorical values to numerical codes. A dataset containing months of the year will be assigned 12 codes, one for each month. These will usually be the integers 0 to 11. Pandas replaces the column values with codes, and retains an index list of category values. In the steps ahead we'll call the categorical values "names" and the encodings "codes".
# Convert our three categorical columns to category dtypes.
for cat in cat_cols:
df[cat] = df[cat].astype('category')
df.dtypes
# We can see that <tt>df['Hour']</tt> is a categorical feature by displaying some of the rows:
df['Hour'].head()
# Here our categorical names are the integers 0 through 23, for a total of 24 unique categories. These values <em>also</em> correspond to the codes assigned to each name.
#
# We can access the category names with <tt>Series.cat.categories</tt> or just the codes with <tt>Series.cat.codes</tt>. This will make more sense if we look at <tt>df['AMorPM']</tt>:
df['AMorPM'].head()
df['AMorPM'].cat.categories
df['AMorPM'].head().cat.codes
df['Weekday'].cat.categories
df['Weekday'].head().cat.codes
# <div class="alert alert-info"><strong>NOTE: </strong>NaN values in categorical data are assigned a code of -1. We don't have any in this particular dataset.</div>
# Now we want to combine the three categorical columns into one input array using <a href='https://docs.scipy.org/doc/numpy/reference/generated/numpy.stack.html'><tt>numpy.stack</tt></a> We don't want the Series index, just the values.
# +
hr = df['Hour'].cat.codes.values
ampm = df['AMorPM'].cat.codes.values
wkdy = df['Weekday'].cat.codes.values
cats = np.stack([hr, ampm, wkdy], 1)
cats[:5]
# -
# <div class="alert alert-info"><strong>NOTE:</strong> This can be done in one line of code using a list comprehension:
# <pre style='background-color:rgb(217,237,247)'>cats = np.stack([df[col].cat.codes.values for col in cat_cols], 1)</pre>
#
# Don't worry about the dtype for now, we can make it int64 when we convert it to a tensor.</div>
#
# ## Convert numpy arrays to tensors
# +
# Convert categorical variables to a tensor
cats = torch.tensor(cats, dtype=torch.int64)
# this syntax is ok, since the source data is an array, not an existing tensor
cats[:5]
# -
# We can feed all of our continuous variables into the model as a tensor. Note that we're not normalizing the values here; we'll let the model perform this step.
# <div class="alert alert-info"><strong>NOTE:</strong> We have to store <tt>conts</tt> and <tt>y</tt> as Float (float32) tensors, not Double (float64) in order for batch normalization to work properly.</div>
# Convert continuous variables to a tensor
conts = np.stack([df[col].values for col in cont_cols], 1)
conts = torch.tensor(conts, dtype=torch.float)
conts[:5]
conts.type()
# +
# Convert labels to a tensor
y = torch.tensor(df[y_col].values, dtype=torch.float).reshape(-1,1)
y[:5]
# -
cats.shape
conts.shape
y.shape
# ## Set an embedding size
# The rule of thumb for determining the embedding size is to divide the number of unique entries in each column by 2, but not to exceed 50.
# This will set embedding sizes for Hours, AMvsPM and Weekdays
cat_szs = [len(df[col].cat.categories) for col in cat_cols]
emb_szs = [(size, min(50, (size+1)//2)) for size in cat_szs]
emb_szs
# ## Define a TabularModel
# This somewhat follows the <a href='https://docs.fast.ai/tabular.models.html'>fast.ai library</a> The goal is to define a model based on the number of continuous columns (given by <tt>conts.shape[1]</tt>) plus the number of categorical columns and their embeddings (given by <tt>len(emb_szs)</tt> and <tt>emb_szs</tt> respectively). The output would either be a regression (a single float value), or a classification (a group of bins and their softmax values). For this exercise our output will be a single regression value. Note that we'll assume our data contains both categorical and continuous data. You can add boolean parameters to your own model class to handle a variety of datasets.
# <div class="alert alert-info"><strong>Let's walk through the steps we're about to take. See below for more detailed illustrations of the steps.</strong><br>
#
# 1. Extend the base Module class, set up the following parameters:
# * <tt>emb_szs: </tt>list of tuples: each categorical variable size is paired with an embedding size
# * <tt>n_cont: </tt>int: number of continuous variables
# * <tt>out_sz: </tt>int: output size
# * <tt>layers: </tt>list of ints: layer sizes
# * <tt>p: </tt>float: dropout probability for each layer (for simplicity we'll use the same value throughout)
#
# <tt><font color=black>class TabularModel(nn.Module):<br>
# def \_\_init\_\_(self, emb_szs, n_cont, out_sz, layers, p=0.5):<br>
# super().\_\_init\_\_()</font></tt><br>
#
# 2. Set up the embedded layers with <a href='https://pytorch.org/docs/stable/nn.html#modulelist'><tt><strong>torch.nn.ModuleList()</strong></tt></a> and <a href='https://pytorch.org/docs/stable/nn.html#embedding'><tt><strong>torch.nn.Embedding()</strong></tt></a><br>Categorical data will be filtered through these Embeddings in the forward section.<br>
# <tt><font color=black> self.embeds = nn.ModuleList([nn.Embedding(ni, nf) for ni,nf in emb_szs])</font></tt><br><br>
# 3. Set up a dropout function for the embeddings with <a href='https://pytorch.org/docs/stable/nn.html#dropout'><tt><strong>torch.nn.Dropout()</strong></tt></a> The default p-value=0.5<br>
# <tt><font color=black> self.emb_drop = nn.Dropout(emb_drop)</font></tt><br><br>
# 4. Set up a normalization function for the continuous variables with <a href='https://pytorch.org/docs/stable/nn.html#batchnorm1d'><tt><strong>torch.nn.BatchNorm1d()</strong></tt></a><br>
# <tt><font color=black> self.bn_cont = nn.BatchNorm1d(n_cont)</font></tt><br><br>
# 5. Set up a sequence of neural network layers where each level includes a Linear function, an activation function (we'll use <a href='https://pytorch.org/docs/stable/nn.html#relu'><strong>ReLU</strong></a>), a normalization step, and a dropout layer. We'll combine the list of layers with <a href='https://pytorch.org/docs/stable/nn.html#sequential'><tt><strong>torch.nn.Sequential()</strong></tt></a><br>
# <tt><font color=black> self.bn_cont = nn.BatchNorm1d(n_cont)<br>
# layerlist = []<br>
# n_emb = sum((nf for ni,nf in emb_szs))<br>
# n_in = n_emb + n_cont<br>
# <br>
# for i in layers:<br>
# layerlist.append(nn.Linear(n_in,i)) <br>
# layerlist.append(nn.ReLU(inplace=True))<br>
# layerlist.append(nn.BatchNorm1d(i))<br>
# layerlist.append(nn.Dropout(p))<br>
# n_in = i<br>
# layerlist.append(nn.Linear(layers[-1],out_sz))<br>
# <br>
# self.layers = nn.Sequential(*layerlist)</font></tt><br><br>
# 6. Define the forward method. Preprocess the embeddings and normalize the continuous variables before passing them through the layers.<br>Use <a href='https://pytorch.org/docs/stable/torch.html#torch.cat'><tt><strong>torch.cat()</strong></tt></a> to combine multiple tensors into one.<br>
# <tt><font color=black> def forward(self, x_cat, x_cont):<br>
# embeddings = []<br>
# for i,e in enumerate(self.embeds):<br>
# embeddings.append(e(x_cat[:,i]))<br>
# x = torch.cat(embeddings, 1)<br>
# x = self.emb_drop(x)<br>
# <br>
# x_cont = self.bn_cont(x_cont)<br>
# x = torch.cat([x, x_cont], 1)<br>
# x = self.layers(x)<br>
# return x</font></tt>
# </div>
# <div class="alert alert-danger"><strong>Breaking down the embeddings steps</strong> (this code is for illustration purposes only.)</div>
# This is our source data
catz = cats[:4]
catz
# This is passed in when the model is instantiated
emb_szs
# This is assigned inside the __init__() method
selfembeds = nn.ModuleList([nn.Embedding(ni, nf) for ni,nf in emb_szs])
selfembeds
list(enumerate(selfembeds))
# This happens inside the forward() method
embeddingz = []
for i,e in enumerate(selfembeds):
embeddingz.append(e(catz[:,i]))
embeddingz
# We concatenate the embedding sections (12,1,4) into one (17)
z = torch.cat(embeddingz, 1)
z
# This was assigned under the __init__() method
selfembdrop = nn.Dropout(.4)
z = selfembdrop(z)
z
# <div class="alert alert-danger"><strong>This is how the categorical embeddings are passed into the layers.</strong></div>
class TabularModel(nn.Module):
def __init__(self, emb_szs, n_cont, out_sz, layers, p=0.5):
super().__init__()
self.embeds = nn.ModuleList([nn.Embedding(ni, nf) for ni,nf in emb_szs])
self.emb_drop = nn.Dropout(p)
self.bn_cont = nn.BatchNorm1d(n_cont)
layerlist = []
n_emb = sum((nf for ni,nf in emb_szs))
n_in = n_emb + n_cont
for i in layers:
layerlist.append(nn.Linear(n_in,i))
layerlist.append(nn.ReLU(inplace=True))
layerlist.append(nn.BatchNorm1d(i))
layerlist.append(nn.Dropout(p))
n_in = i
layerlist.append(nn.Linear(layers[-1],out_sz))
self.layers = nn.Sequential(*layerlist)
def forward(self, x_cat, x_cont):
embeddings = []
for i,e in enumerate(self.embeds):
embeddings.append(e(x_cat[:,i]))
x = torch.cat(embeddings, 1)
x = self.emb_drop(x)
x_cont = self.bn_cont(x_cont)
x = torch.cat([x, x_cont], 1)
x = self.layers(x)
return x
torch.manual_seed(33)
model = TabularModel(emb_szs, conts.shape[1], 1, [200,100], p=0.4)
model
# ## Define loss function & optimizer
# PyTorch does not offer a built-in <a href='https://en.wikipedia.org/wiki/Root-mean-square_deviation'>RMSE Loss</a> function, and it would be nice to see this in place of MSE.<br>
# For this reason, we'll simply apply the <tt>torch.sqrt()</tt> function to the output of MSELoss during training.
criterion = nn.MSELoss() # we'll convert this to RMSE later
optimizer = torch.optim.Adam(model.parameters(), lr=0.001)
# ## Perform train/test splits
# At this point our batch size is the entire dataset of 120,000 records. This will take a long time to train, so you might consider reducing this. We'll use 60,000. Recall that our tensors are already randomly shuffled.
# +
batch_size = 60000
test_size = int(batch_size * .2)
cat_train = cats[:batch_size-test_size]
cat_test = cats[batch_size-test_size:batch_size]
con_train = conts[:batch_size-test_size]
con_test = conts[batch_size-test_size:batch_size]
y_train = y[:batch_size-test_size]
y_test = y[batch_size-test_size:batch_size]
# -
len(cat_train)
len(cat_test)
# ## Train the model
# Expect this to take 30 minutes or more! We've added code to tell us the duration at the end.
# +
import time
start_time = time.time()
epochs = 300
losses = []
for i in range(epochs):
i+=1
y_pred = model(cat_train, con_train)
loss = torch.sqrt(criterion(y_pred, y_train)) # RMSE
losses.append(loss)
# a neat trick to save screen space:
if i%25 == 1:
print(f'epoch: {i:3} loss: {loss.item():10.8f}')
optimizer.zero_grad()
loss.backward()
optimizer.step()
print(f'epoch: {i:3} loss: {loss.item():10.8f}') # print the last line
print(f'\nDuration: {time.time() - start_time:.0f} seconds') # print the time elapsed
# -
# ## Plot the loss function
plt.plot(range(epochs), losses)
plt.ylabel('RMSE Loss')
plt.xlabel('epoch');
# ## Validate the model
# Here we want to run the entire test set through the model, and compare it to the known labels.<br>
# For this step we don't want to update weights and biases, so we set <tt>torch.no_grad()</tt>
# TO EVALUATE THE ENTIRE TEST SET
with torch.no_grad():
y_val = model(cat_test, con_test)
loss = torch.sqrt(criterion(y_val, y_test))
print(f'RMSE: {loss:.8f}')
# This means that on average, predicted values are within ±$3.31 of the actual value.
#
# Now let's look at the first 50 predicted values:
print(f'{"PREDICTED":>12} {"ACTUAL":>8} {"DIFF":>8}')
for i in range(50):
diff = np.abs(y_val[i].item()-y_test[i].item())
print(f'{i+1:2}. {y_val[i].item():8.4f} {y_test[i].item():8.4f} {diff:8.4f}')
# So while many predictions were off by a few cents, some were off by \\$19.00. Feel free to change the batch size, test size, and number of epochs to obtain a better model.
# ## Save the model
# We can save a trained model to a file in case we want to come back later and feed new data through it. The best practice is to save the state of the model (weights & biases) and not the full definition. Also, we want to ensure that only a trained model is saved, to prevent overwriting a previously saved model with an untrained one.<br>For more information visit <a href='https://pytorch.org/tutorials/beginner/saving_loading_models.html'>https://pytorch.org/tutorials/beginner/saving_loading_models.html</a>
# Make sure to save the model only after the training has happened!
if len(losses) == epochs:
torch.save(model.state_dict(), 'TaxiFareRegrModel.pt')
else:
print('Model has not been trained. Consider loading a trained model instead.')
# ## Loading a saved model (starting from scratch)
# We can load the trained weights and biases from a saved model. If we've just opened the notebook, we'll have to run standard imports and function definitions. To demonstrate, restart the kernel before proceeding.
# +
import torch
import torch.nn as nn
import numpy as np
import pandas as pd
def haversine_distance(df, lat1, long1, lat2, long2):
r = 6371
phi1 = np.radians(df[lat1])
phi2 = np.radians(df[lat2])
delta_phi = np.radians(df[lat2]-df[lat1])
delta_lambda = np.radians(df[long2]-df[long1])
a = np.sin(delta_phi/2)**2 + np.cos(phi1) * np.cos(phi2) * np.sin(delta_lambda/2)**2
c = 2 * np.arctan2(np.sqrt(a), np.sqrt(1-a))
return r * c
class TabularModel(nn.Module):
def __init__(self, emb_szs, n_cont, out_sz, layers, p=0.5):
super().__init__()
self.embeds = nn.ModuleList([nn.Embedding(ni, nf) for ni,nf in emb_szs])
self.emb_drop = nn.Dropout(p)
self.bn_cont = nn.BatchNorm1d(n_cont)
layerlist = []
n_emb = sum((nf for ni,nf in emb_szs))
n_in = n_emb + n_cont
for i in layers:
layerlist.append(nn.Linear(n_in,i))
layerlist.append(nn.ReLU(inplace=True))
layerlist.append(nn.BatchNorm1d(i))
layerlist.append(nn.Dropout(p))
n_in = i
layerlist.append(nn.Linear(layers[-1],out_sz))
self.layers = nn.Sequential(*layerlist)
def forward(self, x_cat, x_cont):
embeddings = []
for i,e in enumerate(self.embeds):
embeddings.append(e(x_cat[:,i]))
x = torch.cat(embeddings, 1)
x = self.emb_drop(x)
x_cont = self.bn_cont(x_cont)
x = torch.cat([x, x_cont], 1)
return self.layers(x)
# -
# Now define the model. Before we can load the saved settings, we need to instantiate our TabularModel with the parameters we used before (embedding sizes, number of continuous columns, output size, layer sizes, and dropout layer p-value).
emb_szs = [(24, 12), (2, 1), (7, 4)]
model2 = TabularModel(emb_szs, 6, 1, [200,100], p=0.4)
# Once the model is set up, loading the saved settings is a snap.
model2.load_state_dict(torch.load('TaxiFareRegrModel.pt'));
model2.eval() # be sure to run this step!
# Next we'll define a function that takes in new parameters from the user, performs all of the preprocessing steps above, and passes the new data through our trained model.
def test_data(mdl): # pass in the name of the new model
# INPUT NEW DATA
plat = float(input('What is the pickup latitude? '))
plong = float(input('What is the pickup longitude? '))
dlat = float(input('What is the dropoff latitude? '))
dlong = float(input('What is the dropoff longitude? '))
psngr = int(input('How many passengers? '))
dt = input('What is the pickup date and time?\nFormat as YYYY-MM-DD HH:MM:SS ')
# PREPROCESS THE DATA
dfx_dict = {'pickup_latitude':plat,'pickup_longitude':plong,'dropoff_latitude':dlat,
'dropoff_longitude':dlong,'passenger_count':psngr,'EDTdate':dt}
dfx = pd.DataFrame(dfx_dict, index=[0])
dfx['dist_km'] = haversine_distance(dfx,'pickup_latitude', 'pickup_longitude',
'dropoff_latitude', 'dropoff_longitude')
dfx['EDTdate'] = pd.to_datetime(dfx['EDTdate'])
# We can skip the .astype(category) step since our fields are small,
# and encode them right away
dfx['Hour'] = dfx['EDTdate'].dt.hour
dfx['AMorPM'] = np.where(dfx['Hour']<12,0,1)
dfx['Weekday'] = dfx['EDTdate'].dt.strftime("%a")
dfx['Weekday'] = dfx['Weekday'].replace(['Fri','Mon','Sat','Sun','Thu','Tue','Wed'],
[0,1,2,3,4,5,6]).astype('int64')
# CREATE CAT AND CONT TENSORS
cat_cols = ['Hour', 'AMorPM', 'Weekday']
cont_cols = ['pickup_latitude', 'pickup_longitude', 'dropoff_latitude',
'dropoff_longitude', 'passenger_count', 'dist_km']
xcats = np.stack([dfx[col].values for col in cat_cols], 1)
xcats = torch.tensor(xcats, dtype=torch.int64)
xconts = np.stack([dfx[col].values for col in cont_cols], 1)
xconts = torch.tensor(xconts, dtype=torch.float)
# PASS NEW DATA THROUGH THE MODEL WITHOUT PERFORMING A BACKPROP
with torch.no_grad():
z = mdl(xcats, xconts)
print(f'\nThe predicted fare amount is ${z.item():.2f}')
# ## Feed new data through the trained model
# For convenience, here are the max and min values for each of the variables:
# <table style="display: inline-block">
# <tr><th>Column</th><th>Minimum</th><th>Maximum</th></tr>
# <tr><td>pickup_latitude</td><td>40</td><td>41</td></tr>
# <tr><td>pickup_longitude</td><td>-74.5</td><td>-73.3</td></tr>
# <tr><td>dropoff_latitude</td><td>40</td><td>41</td></tr>
# <tr><td>dropoff_longitude</td><td>-74.5</td><td>-73.3</td></tr>
# <tr><td>passenger_count</td><td>1</td><td>5</td></tr>
# <tr><td>EDTdate</td><td>2010-04-11 00:00:00</td><td>2010-04-24 23:59:42</td></tr>
# <strong>Use caution!</strong> The distance between 1 degree of latitude (from 40 to 41) is 111km (69mi) and between 1 degree of longitude (from -73 to -74) is 85km (53mi). The longest cab ride in the dataset spanned a difference of only 0.243 degrees latitude and 0.284 degrees longitude. The mean difference for both latitude and longitude was about 0.02. To get a fair prediction, use values that fall close to one another.
z = test_data(model2)
# ## Great job!
| 04a-Full-ANN-Code-Along-Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#hide
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# +
# default_exp perturb
# -
# # Perturb
#
# > Functions for perturbing a graph
#export
from nbdev.showdoc import *
import networkx as nx
import numpy as np
import pandas as pd
from grapht.graphtools import non_pendant_edges, has_isolated_nodes
from grapht.sampling import khop_subgraph, sample_edges, sample_nodes
# ## Edge additions and deletions
#export
def randomly_perturb(G, add=0, remove=0):
"""Randomly add and remove edges."""
Gp = G.copy()
edges_to_remove = sample_edges(G, remove)
edges_to_add = []
while len(edges_to_add) < add:
edge = sample_nodes(G, 2)
if edge not in G.edges:
edges_to_add.append(edge)
Gp.remove_edges_from(edges_to_remove)
Gp.add_edges_from(edges_to_add)
return Gp
# ## Edge deletions
#export
def khop_remove(G, k, r, max_iter=np.Inf, enforce_connected=False, enforce_no_isolates=True):
"""Removes r edges which are in a k-hop neighbourhood of a random node.
Args:
G: A nx.Graph to remove edges from.
k: If None then remove edges uniformly, else remove in a k-hop neighbourhood.
r: The number of edges to remove.
max_iter: The maximum number of attempts to find a valid perturbation.
enforce_connected: If True the perturbed graph will be connected.
enforce_no_isolates: If True the perturbed graph will not contain isolated nodes.
Returns:
solution: a perturbed graph.
edges: a list of edges which were removed.
node: the node which the k-hop neighbourhood was taken around.
"""
solution = None
attempts = 0
while solution is None:
# generate subgraph
if k is not None:
subgraph, node = khop_subgraph(G, k)
else:
subgraph, node = G, None
# check subgraph can yield a solution
if not enforce_no_isolates and len(subgraph.edges()) < r:
continue
if enforce_no_isolates and len(non_pendant_edges(subgraph)) < r:
continue
# perturb graph
edges = sample_edges(subgraph, r, non_pendant=enforce_no_isolates)
Gp = G.copy()
Gp.remove_edges_from(edges)
# check its valid
if enforce_connected:
if nx.is_connected(Gp):
solution = Gp
else:
if enforce_no_isolates:
if not has_isolated_nodes(Gp):
solution = Gp
else:
solution = Gp
# timeout counter
attempts += 1
if attempts >= max_iter:
break
# return solution if found
if solution is None:
return None
else:
edge_info = pd.DataFrame(edges, columns=['u', 'v'])
edge_info['type'] = 'remove'
return solution, edge_info, node
# ## Rewiring
# +
#export
def khop_rewire(G, k, r, max_iter=100):
"""Rewire the graph in place where edges which are rewired are in a k-hop neighbourhood.
A random k-hop neighbourhood is selected in G and r edges are rewired.
If the graph contains an isolated node this procedure is repeated.
If `max_iter` attempted do not give a graph without isolated nodes `None` is returned.
Returns:
solution (nx.Graph): the rewired graph
rewire_info (pd.DataFrame): a dataframe describing which edges were added or removed
node: The node from which the k-hop neighbourhood was taken around
"""
solution = None
for _ in range(max_iter):
if k is not None:
subgraph, node = khop_subgraph(G, k)
else:
subgraph, node = G, None
if len(subgraph.edges()) < r:
continue
edges = sample_edges(subgraph, r, non_pendant=False)
Gp = G.copy()
rewire_info = rewire(Gp, edges)
if not has_isolated_nodes(Gp):
solution = Gp
if solution is None:
return None
else:
return solution, rewire_info, node
def rewire(G, edges):
"""Rewires `edges` in `G` inplace and returns a dataframe with the edges which were added or removed.
All edges are broken into stubs and then stubs are randomly joined together.
Self loops are removed after the rewiring step.
A dataframe is returned where each row is (u, v, 'add') or (u, v, 'remove').
The dataframe will include entries (u, u, 'add') if self loops were added but these won't appear in the graph.
"""
edges = np.array(edges)
new_edges = np.reshape(np.random.permutation(edges.flatten()), (-1, 2))
G.remove_edges_from(edges.tolist())
G.add_edges_from(new_edges.tolist())
G.remove_edges_from(nx.selfloop_edges(G))
df_remove = pd.DataFrame(edges, columns = ['u', 'v'])
df_remove['type'] = 'remove'
df_add = pd.DataFrame(new_edges, columns = ['u', 'v'])
df_add['type'] = 'add'
return pd.concat([df_remove, df_add], ignore_index=True)
# -
#hide
from nbdev.export import notebook2script
notebook2script()
| nbs/03_perturb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import random
import datetime
df = pd.read_csv("../datasets/transactions_raw.csv", parse_dates=['created_at'])
clients = pd.read_csv("../datasets/clients.csv", parse_dates=['created_at'])
items = pd.read_csv("../datasets/items.csv")
df.head()
clients.head()
items.head()
sold = random.sample(items.item_id.tolist(), int(.05*items.shape[0]))*15 + random.sample(items.item_id.tolist(), int(.8*items.shape[0]))*10
df['item_id'] = [random.choice(sold) for x in range(df.shape[0])]
df
fclients = clients.sample(int(.42*clients.shape[0]))
# +
# df['client_id'] = [random.choice(fclients) for x in range(df.shape[0])]
# -
df
def fill_client_id(created_at):
try:
filtered_clients = clients[clients.created_at<created_at]
potential_buyers = filtered_clients.client_id.tolist()
if (random.randint(0, 2)==0):
return potential_buyers[-1]
return random.choice(potential_buyers)
except:
return 0
df['client_id'] = df.created_at.apply(fill_client_id)
df
df[df.client_id!=0].to_csv('../datasets/transactions.csv', index=False)
| notebooks/4_transactions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # pandas.DataFrame.select_dtypes¶
#
# DataFrame.select_dtypes(include=None, exclude=None)
import pandas as pd
import numpy as np
# Create a data Frame
df = pd.DataFrame({ 'a': [1, 2, 3] * 2,
'b': [True, False] *3 ,
'c': [1.0, 2.0, 4.0] * 2,
'd': ['A','X','Z','B','Z','A']})
df.dtypes
df.select_dtypes(exclude=['int64'])
df.select_dtypes(exclude=['object'])
print("All column Names",df.columns)
print("Numeric Columns",df._get_numeric_data().columns)
print("Object Type Columns ",df.select_dtypes(include=['object']).columns)
print("NOT Object Type Columns ",df.select_dtypes(exclude=['object']).columns)
numerics = df.select_dtypes(exclude=['object']).columns
print("Numerics Columns ",numerics)
non_numeric = df.select_dtypes(include=['object']).columns
print("Non Numeric Columns ",non_numeric)
| content/Pandas Operations/Select Columns of specified Data Type.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:calphad-dev-2]
# language: python
# name: conda-env-calphad-dev-2-py
# ---
# # Exploring `calculate` and `equilibrium` xarray Datasets
#
# ## xarray Datasets
#
# Results returned from calling `calculate` or `equilibrium` in pycalphad are [xarray](http://xarray.pydata.org/en/stable/) Datasets. An xarray Dataset is a data structure that represents N-dimensional tabular data. It is an N-dimensional analog to the Pandas DataFrame.
#
# This notebook will walk through the structure of xarray Datasets in pycalphad and some basics of using them. For more in-depth tutorials and documentation on using xarray Datasets and DataArray's fully, see the [xarray documentation](http://xarray.pydata.org/en/stable/index.html).
#
# ## Dataset structure
#
# Each Dataset stores the conditions that properties are calculated at and the values of the properties as a function of the different conditions. There are three key terms:
#
# * `Dimensions`: these are the conditions that are calculated over, e.g. pressure (P) and temperature (T). They are essentially labels.
# * `Coordinates`: these are the actual *values* that are taken on by the dimensions.
# * `Data variables`: these are the properties calculated by pycalphad, such as the Gibbs energy, mixing energy, composition, etc.
#
# ## `calculate()` results
#
# Calculate is used to sample properties of a single phase. There are five dimensions/coordinates:
#
# * `P`: pressures (in Pa).
# * `T`: temperatures (in K).
# * `component`: the string names of the components in the system
# * `internal_dof`: The internal_dof (internal degrees of freedom) is the index of the site in any phase's site fraction array. Below the FCC_A1 phase has the sublattice model (AL, ZN) and thus the internal_dof are integers 0 and 1 referring to the AL site (index 0) and the ZN site (index 1).
# * `points`: By default, the calculate function samples points over all of the internal degrees of freedom. Each coordinate point simply represents the index is a list of all configurations of the internal_dof sampled. There is no underlying physical meaning or order.
#
# There are also at least four Data variables:
#
# * `Phase`: The string name of the phase. For `calculate`, this will always be the phase name passed.
# * `X`: The composition of each component in mole fraction as a function of the temperature, pressure, and the index of the points (there is one composition for each point).
# * `Y`: The site fraction of each index in the internal_dof array for the given temperature, pressure and point.
# * `output`: "output" is always whatever property is calculated by the output keyword passed to `calculate`. The default is the molar Gibbs energy, GM.
# +
# %matplotlib inline
from pycalphad import Database, calculate, equilibrium, variables as v
dbf = Database('alzn_mey.tdb')
comps = ['AL', 'ZN', 'VA']
calc_result = calculate(dbf, comps, 'FCC_A1', P=101325, T=[500, 1000])
print(calc_result)
# -
# We can manipulate this by selecting data by value (of a coordinate) using `sel` or index (of a coordinate) using `isel` similar to a Pandas array. Below we get the site fraction of ZN (internal_dof index of 1 selected by index) at 1000K (selected by value) for the 50th point (selected by index).
#
# The results of selecting over Data variables gives an xarray DataArray which is useful for plotting or performing computations on (see [DataArrays vs Datasets](http://xarray.pydata.org/en/stable/data-structures.html)).
print(calc_result.Y.isel(internal_dof=1, points=49).sel(T=1000))
# accessing the `values` attribute on any on any DataArray returns the multidimensional NumPy array
print(calc_result.X.values)
# ## `equilibrium()` results
#
# The Datasets returned by equilibrium are very similar to calculate, however there are several key differences worth discussing. In equilibrium Datasets, there are six dimensions/coordinates:
#
# * `P`: pressures (in Pa).
# * `T`: temperatures (in K).
# * `component`: (Same as calculate) The string names of the components in the system.
# * `internal_dof`: (Same as calculate, except it will be the longest possible internal_dof for all phases) The internal_dof (internal degrees of freedom) is the index of the site in any phase's site fraction array. Below the FCC_A1 phase has the sublattice model (AL, ZN) and thus the internal_dof are integers 0 and 1 referring to the AL site (index 0) and the ZN site (index 1).
# * `X_ZN`: This is the composition of the species that was passed into the conditions array. Since we passed `v.X('ZN')` to the conditions dictionary, this is `X_ZN`.
# * `vertex`: The vertex is the index of the phase in equilibrium. The vertex has no inherent physical meaning. There will automatically be enough to describe the number of phases present in any equilibria calculated, implying that vertex can never be large enough to invalidate Gibbs phase rule.
#
# There are also at least six Data variables:
#
# * `Phase`: The string name of the phase in equilibrium at the conditions. There are as many as `len(vertex)` phases. Any time there are fewer phases in equilibrium than the indices described by `vertex`, the values of phase are paded by `''`, e.g. for a single phase region for FCC_A1, the values of Phase will be `['FCC_A1', '']`. When more than one phase is present, it is important to note that they are not necessarily sorted.
# * `NP`: Phase fraction of each phase in equilibrium. When there is no other equilibrium phase (e.g. single phase `['FCC_A1', '']`) then the value of `NP` will be `nan` for the absence of a phase, rather than 0.
# * `MU`: The chemical potentials of each component for the conditions calculated.
# * `X`: The equilibrium composition of each element in each phase for the calculated conditions.
# * `Y`: The equilibrium site fraction of each site in each phase for the calculated conditions.
# * `GM`: Same as `output` for `calculate`. It is always reported no matter the value of `output`.
# * `output`: (optional) "output" is always whatever equilibrium property is calculated by the output keyword passed to `equilibrium`. Unlike `calculate`, this will be in addition to the `GM` because `GM` is always reported.
phases = ['LIQUID', 'FCC_A1', 'HCP_A3']
eq_result = equilibrium(dbf, comps , phases, {v.X('ZN'):(0,1,0.05), v.T: (500, 1000, 100), v.P:101325}, output='HM')
print(eq_result)
# A common operation might be to find the phase fractions of the HCP_A3 phase as a function of composition for T=800.
#
# However, the only way we can access the values of the phase fraction is by either the indices or values of the coordinates, we would have to know which index the HCP_A3 phase is in before hand to use the `sel` or `isel` commands.
#
# Since we do not know this, we can do what is called [masking](http://xarray.pydata.org/en/stable/indexing.html#masking-with-where) to find the data values that match a condition (the Phase is FCC_A1):
print(eq_result.NP.where(eq_result.Phase=='FCC_A1').sel(P=101325, T=800))
| examples/UsingCalculationResults.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Ridge Regularization on Housing Dataset
import numpy as np
import pandas as pd
from numpy.linalg import inv
def get_data(column_names):
train_df = pd.read_csv('./data/housing_train.txt', delim_whitespace=True, header = None)
test_df = pd.read_csv('./data/housing_test.txt', delim_whitespace=True, header = None)
test_df.columns = column_names
train_df.columns = column_names
return train_df, test_df
column_names = ['CRIM','ZN','INDUS','CHAS','NOX','RM','AGE','DIS','RAD','TAX','PTRATIO','B','LSTAT','MEDV']
train_data, test_data = get_data(column_names)
train_data.describe()
test_data.describe()
def normalize(dataset):
means = dataset.mean(axis = 0)
dataset = dataset - means
return dataset, means
def get_regularized_weights(train_data, reg_strength):
x = train_data.drop(['MEDV'], axis = 1).values
y = train_data['MEDV'].values
# center the data
z, means = normalize(x)
z = np.append(np.ones([len(z),1]),z,1)
I = np.eye(len(z[0]))
I[0,0] = y.mean()
inverse = inv(np.dot(z.T,z) + reg_strength * I)
w = np.dot(np.dot(inverse, z.T), y)
return w, means
weights, means = get_regularized_weights(train_data, 0)
def predict(test_data, weights, means):
test_data = test_data.drop(['MEDV'], axis = 1).values
test_data = test_data - means
test_data = np.append(np.ones([len(test_data),1]),test_data,1)
preds = {}
for i in range(len(test_data)):
preds[i] = np.dot(weights, test_data[i])
return preds
def get_mse(test_data, preds):
test_labels = test_data['MEDV'].values
errors = []
for i, label in enumerate(test_labels):
errors.append(np.square(label - preds[i]))
mse = pd.Series(errors).mean()
return mse
preds = predict(test_data, weights, means)
print('MSE test for Housing Dataset with Ridge Regularization: {}'.format(get_mse(test_data, preds)))
preds_train = predict(train_data, weights, means)
print('MSE train for Housing Dataset with Ridge Regularization: {}'.format(get_mse(train_data, preds_train)))
| Regularization/Housing_Ridge_Regularization.ipynb |