code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.0 64-bit ('3.7')
# name: python3
# ---
# # Build a cuisine recommender
# ## Train classification model
# First, train a classification model using the cleaned cuisines dataset we used.
#
# - Start by importing useful libraries:
# +
# # !pip install skl2onnx
# -
# You need '*skl2onnx*' to help convert your Scikit-learn model to Onnx format.
import pandas as pd
# Then, work with your data by reading a CSV file using read_csv():
data = pd.read_csv('cleaned_cuisines.csv')
data.head()
# Remove the first two unnecessary columns and save the remaining data as 'X':
X = data.iloc[:,2:]
X.head()
# Save the labels as 'y':
y = data[['cuisine']]
y.head()
# ## Commence the training routine
# We will use the 'SVC' library which has good accuracy.
#
# - Import the appropriate libraries from Scikit-learn:
from sklearn.model_selection import train_test_split
from sklearn.svm import SVC
from sklearn.model_selection import cross_val_score
from sklearn.metrics import accuracy_score,precision_score,confusion_matrix,classification_report
# - Separate training and test sets:
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.3)
# - Build an SVC Classification model
model = SVC(kernel='linear', C=10, probability=True,random_state=0)
model.fit(X_train,y_train.values.ravel())
# - Now, test your model, calling predict():
y_pred = model.predict(X_test)
# - Print out a classification report to check the model's quality:
print(classification_report(y_test,y_pred))
# ## Convert your model to Onnx
# Make sure to do the conversion with the proper Tensor number. This dataset has 380 ingredients listed, so you need to notate that number in *FloatTensorType*:
#
# - Convert using a tensor number of 380.
# +
from skl2onnx import convert_sklearn
from skl2onnx.common.data_types import FloatTensorType
initial_type = [('float_input', FloatTensorType([None, 380]))]
options = {id(model): {'nocl': True, 'zipmap': False}}
onx = convert_sklearn(model, initial_types=initial_type, options=options)
with open("./model.onnx", "wb") as f:
f.write(onx.SerializeToString())
# -
# - Create the onx and store as a file model.onnx:
onx = convert_sklearn(model, initial_types=initial_type, options=options)
with open("./model.onnx", "wb") as f:
f.write(onx.SerializeToString())
# Now you are ready to use this neat model in a web app. Let's build an app that will come in handy when you look in your refrigerator and try to figure out which combination of your leftover ingredients you can use to cook a given cuisine, as determined by your model.
#
| notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# metadata:
# interpreter:
# hash: c51bd52eff1ef297cb7cf13ffa7cb0c7ccecc33c63cf5bbf48ca8c54127178f2
# name: python3
# ---
# # Convolutional Neural Networks with PyTorch
#
# "Deep Learning" is a general term that usually refers to the use of neural networks with multiple layers that synthesize the way the human brain learns and makes decisions. A convolutional neural network is a kind of neural network that extracts *features* from matrices of numeric values (often images) by convolving multiple filters over the matrix values to apply weights and identify patterns, such as edges, corners, and so on in an image. The numeric representations of these patterns are then passed to a fully-connected neural network layer to map the features to specific classes.
#
# There are several commonly used frameworks for creating CNNs. In this notebook, we'll build a simple example CNN using PyTorch.
#
# ## Import libraries
#
# First, let's install and import the PyTorch libraries we'll need.
# !pip install torch==1.7.1 torchvision==0.8.2 torchaudio==0.7.2 -f https://download.pytorch.org/whl/torch_stable.html
# + tags=[]
# Import PyTorch libraries
import torch
import torchvision
import torchvision.transforms as transforms
import torch.nn as nn
import torch.optim as optim
from torch.autograd import Variable
import torch.nn.functional as F
# Other libraries we'll use
import numpy as np
import os
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
# %matplotlib inline
print("Libraries imported - ready to use PyTorch", torch.__version__)
# -
# ## Explore the data
#
# In this exercise, you'll train a CNN-based classification model that can classify images of geometric shapes. Let's take a look at the classes of shape the model needs to identify.
# +
# The images are in the data/shapes folder
data_path = 'data/shapes/'
# Get the class names
classes = os.listdir(data_path)
classes.sort()
print(len(classes), 'classes:')
print(classes)
# Show the first image in each folder
fig = plt.figure(figsize=(8, 12))
i = 0
for sub_dir in os.listdir(data_path):
i+=1
img_file = os.listdir(os.path.join(data_path,sub_dir))[0]
img_path = os.path.join(data_path, sub_dir, img_file)
img = mpimg.imread(img_path)
a=fig.add_subplot(1, len(classes),i)
a.axis('off')
imgplot = plt.imshow(img)
a.set_title(img_file)
plt.show()
# -
# ## Load data
#
# PyTorch includes functions for loading and transforming data. We'll use these to create an iterative loader for training data, and a second iterative loader for test data (which we'll use to validate the trained model). The loaders will transform the image data into *tensors*, which are the core data structure used in PyTorch, and normalize them so that the pixel values are in a scale with a mean of 0.5 and a standard deviation of 0.5.
#
# Run the following cell to define the data loaders.
# + tags=[]
# Function to ingest data using training and test loaders
def load_dataset(data_path):
# Load all of the images
transformation = transforms.Compose([
# transform to tensors
transforms.ToTensor(),
# Normalize the pixel values (in R, G, and B channels)
transforms.Normalize(mean=[0.5, 0.5, 0.5], std=[0.5, 0.5, 0.5])
])
# Load all of the images, transforming them
full_dataset = torchvision.datasets.ImageFolder(
root=data_path,
transform=transformation
)
# Split into training (70% and testing (30%) datasets)
train_size = int(0.7 * len(full_dataset))
test_size = len(full_dataset) - train_size
train_dataset, test_dataset = torch.utils.data.random_split(full_dataset, [train_size, test_size])
# define a loader for the training data we can iterate through in 50-image batches
train_loader = torch.utils.data.DataLoader(
train_dataset,
batch_size=50,
num_workers=0,
shuffle=False
)
# define a loader for the testing data we can iterate through in 50-image batches
test_loader = torch.utils.data.DataLoader(
test_dataset,
batch_size=50,
num_workers=0,
shuffle=False
)
return train_loader, test_loader
# Get the iterative dataloaders for test and training data
train_loader, test_loader = load_dataset(data_path)
print('Data loaders ready')
# -
# ## Define the CNN
#
# In PyTorch, you define a neural network model as a class that is derived from the **nn.Module** base class. Your class must define the layers in your network, and provide a **forward** method that is used to process data through the layers of the network.
# + tags=[]
# Create a neural net class
class Net(nn.Module):
# Constructor
def __init__(self, num_classes=3):
super(Net, self).__init__()
# Our images are RGB, so input channels = 3. We'll apply 12 filters in the first convolutional layer
self.conv1 = nn.Conv2d(in_channels=3, out_channels=12, kernel_size=3, stride=1, padding=1)
# We'll apply max pooling with a kernel size of 2
self.pool = nn.MaxPool2d(kernel_size=2)
# A second convolutional layer takes 12 input channels, and generates 12 outputs
self.conv2 = nn.Conv2d(in_channels=12, out_channels=12, kernel_size=3, stride=1, padding=1)
# A third convolutional layer takes 12 inputs and generates 24 outputs
self.conv3 = nn.Conv2d(in_channels=12, out_channels=24, kernel_size=3, stride=1, padding=1)
# A drop layer deletes 20% of the features to help prevent overfitting
self.drop = nn.Dropout2d(p=0.2)
# Our 128x128 image tensors will be pooled twice with a kernel size of 2. 128/2/2 is 32.
# So our feature tensors are now 32 x 32, and we've generated 24 of them
# We need to flatten these and feed them to a fully-connected layer
# to map them to the probability for each class
self.fc = nn.Linear(in_features=32 * 32 * 24, out_features=num_classes)
def forward(self, x):
# Use a relu activation function after layer 1 (convolution 1 and pool)
x = F.relu(self.pool(self.conv1(x)))
# Use a relu activation function after layer 2 (convolution 2 and pool)
x = F.relu(self.pool(self.conv2(x)))
# Select some features to drop after the 3rd convolution to prevent overfitting
x = F.relu(self.drop(self.conv3(x)))
# Only drop the features if this is a training pass
x = F.dropout(x, training=self.training)
# Flatten
x = x.view(-1, 32 * 32 * 24)
# Feed to fully-connected layer to predict class
x = self.fc(x)
# Return class probabilities via a log_softmax function
return F.log_softmax(x, dim=1)
print("CNN model class defined!")
# -
# ## Train the model
#
# Now that we've defined a class for the network, we can train it using the image data.
#
# Training consists of an iterative series of forward passes in which the training data is processed in batches by the layers in the network, and the optimizer goes back and adjusts the weights. We'll also use a separate set of test images to test the model at the end of each iteration (or *epoch*) so we can track the performance improvement as the training process progresses.
#
# In the example below, we use 5 epochs to train the model using the batches of images loaded by the data loaders, holding back the data in the test data loader for validation. After each epoch, a loss function measures the error (*loss*) in the model and adjusts the weights (which were randomly generated for the first iteration) to try to improve accuracy.
#
# > **Note**: We're only using 5 epochs to minimze the training time for this simple example. A real-world CNN is usually trained over more epochs than this. CNN model training is processor-intensive, involving a lot of matrix and vector-based operations; so it's recommended to perform this on a system that can leverage GPUs, which are optimized for these kinds of calculation. This will take a while to complete on a CPU-based system - status will be displayed as the training progresses.
# + tags=[]
def train(model, device, train_loader, optimizer, epoch):
# Set the model to training mode
model.train()
train_loss = 0
print("Epoch:", epoch)
# Process the images in batches
for batch_idx, (data, target) in enumerate(train_loader):
# Use the CPU or GPU as appropriate
data, target = data.to(device), target.to(device)
# Reset the optimizer
optimizer.zero_grad()
# Push the data forward through the model layers
output = model(data)
# Get the loss
loss = loss_criteria(output, target)
# Keep a running total
train_loss += loss.item()
# Backpropagate
loss.backward()
optimizer.step()
# Print metrics for every 10 batches so we see some progress
if batch_idx % 10 == 0:
print('Training set [{}/{} ({:.0f}%)] Loss: {:.6f}'.format(
batch_idx * len(data), len(train_loader.dataset),
100. * batch_idx / len(train_loader), loss.item()))
# return average loss for the epoch
avg_loss = train_loss / (batch_idx+1)
print('Training set: Average loss: {:.6f}'.format(avg_loss))
return avg_loss
def test(model, device, test_loader):
# Switch the model to evaluation mode (so we don't backpropagate or drop)
model.eval()
test_loss = 0
correct = 0
with torch.no_grad():
batch_count = 0
for data, target in test_loader:
batch_count += 1
data, target = data.to(device), target.to(device)
# Get the predicted classes for this batch
output = model(data)
# Calculate the loss for this batch
test_loss += loss_criteria(output, target).item()
# Calculate the accuracy for this batch
_, predicted = torch.max(output.data, 1)
correct += torch.sum(target==predicted).item()
# Calculate the average loss and total accuracy for this epoch
avg_loss = test_loss/batch_count
print('Validation set: Average loss: {:.6f}, Accuracy: {}/{} ({:.0f}%)\n'.format(
avg_loss, correct, len(test_loader.dataset),
100. * correct / len(test_loader.dataset)))
# return average loss for the epoch
return avg_loss
# Now use the train and test functions to train and test the model
device = "cpu"
if (torch.cuda.is_available()):
# if GPU available, use cuda (on a cpu, training will take a considerable length of time!)
device = "cuda"
print('Training on', device)
# Create an instance of the model class and allocate it to the device
model = Net(num_classes=len(classes)).to(device)
# Use an "Adam" optimizer to adjust weights
# (see https://pytorch.org/docs/stable/optim.html#algorithms for details of supported algorithms)
optimizer = optim.Adam(model.parameters(), lr=0.001)
# Specify the loss criteria
loss_criteria = nn.CrossEntropyLoss()
# Track metrics in these arrays
epoch_nums = []
training_loss = []
validation_loss = []
# Train over 5 epochs (in a real scenario, you'd likely use many more)
epochs = 5
for epoch in range(1, epochs + 1):
train_loss = train(model, device, train_loader, optimizer, epoch)
test_loss = test(model, device, test_loader)
epoch_nums.append(epoch)
training_loss.append(train_loss)
validation_loss.append(test_loss)
# -
# ## View the loss history
#
# We tracked average training and validation loss for each epoch. We can plot these to verify that loss reduced as the model was trained, and to detect *over-fitting* (which is indicated by a continued drop in training loss after validation loss has levelled out or started to increase).
# +
# %matplotlib inline
from matplotlib import pyplot as plt
plt.plot(epoch_nums, training_loss)
plt.plot(epoch_nums, validation_loss)
plt.xlabel('epoch')
plt.ylabel('loss')
plt.legend(['training', 'validation'], loc='upper right')
plt.show()
# -
# ## Evaluate model performance
#
# You can see the final accuracy based on the test data, but typically you'll want to explore performance metrics in a little more depth. Let's plot a confusion matrix to see how well the model is predicting each class.
# + tags=[]
# Pytorch doesn't have a built-in confusion matrix metric, so we'll use SciKit-Learn
from sklearn.metrics import confusion_matrix
# Set the model to evaluate mode
model.eval()
# Get predictions for the test data and convert to numpy arrays for use with SciKit-Learn
print("Getting predictions from test set...")
truelabels = []
predictions = []
for data, target in test_loader:
for label in target.cpu().data.numpy():
truelabels.append(label)
for prediction in model.cpu()(data).data.numpy().argmax(1):
predictions.append(prediction)
# Plot the confusion matrix
cm = confusion_matrix(truelabels, predictions)
plt.imshow(cm, interpolation="nearest", cmap=plt.cm.Blues)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
plt.xlabel("Predicted Shape")
plt.ylabel("Actual Shape")
plt.show()
# -
# ## Save the Trained model
#
# Now that you've trained a working model, you can save it (including the trained weights) for use later.
# Save the model weights
model_file = 'models/shape_classifier.pt'
torch.save(model.state_dict(), model_file)
del model
print('model saved as', model_file)
# ## Use the trained model
#
# Now that we've trained and evaluated our model, we can use it to predict classes for new images.
# + tags=[]
import matplotlib.pyplot as plt
import os
from random import randint
# %matplotlib inline
# Function to predict the class of an image
def predict_image(classifier, image):
import numpy
# Set the classifer model to evaluation mode
classifier.eval()
# Apply the same transformations as we did for the training images
transformation = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(mean=[0.5, 0.5, 0.5], std=[0.5, 0.5, 0.5])
])
# Preprocess the image
image_tensor = transformation(image).float()
# Add an extra batch dimension since pytorch treats all inputs as batches
image_tensor = image_tensor.unsqueeze_(0)
# Turn the input into a Variable
input_features = Variable(image_tensor)
# Predict the class of the image
output = classifier(input_features)
index = output.data.numpy().argmax()
return index
# Function to create a random image (of a square, circle, or triangle)
def create_image (size, shape):
from random import randint
import numpy as np
from PIL import Image, ImageDraw
xy1 = randint(10,40)
xy2 = randint(60,100)
col = (randint(0,200), randint(0,200), randint(0,200))
img = Image.new("RGB", size, (255, 255, 255))
draw = ImageDraw.Draw(img)
if shape == 'circle':
draw.ellipse([(xy1,xy1), (xy2,xy2)], fill=col)
elif shape == 'triangle':
draw.polygon([(xy1,xy1), (xy2,xy2), (xy2,xy1)], fill=col)
else: # square
draw.rectangle([(xy1,xy1), (xy2,xy2)], fill=col)
del draw
return np.array(img)
# Create a random test image
classnames = os.listdir(os.path.join('data', 'shapes'))
classnames.sort()
shape = classnames[randint(0, len(classnames)-1)]
img = create_image ((128,128), shape)
# Display the image
plt.axis('off')
plt.imshow(img)
# Create a new model class and load the saved weights
model = Net()
model.load_state_dict(torch.load(model_file))
# Call the predction function
index = predict_image(model, img)
print(classes[index])
# -
# ## Further Reading
#
# To learn more about training convolutional neural networks with PyTorch, see the [PyTorch documentation](https://pytorch.org/).
#
# ## Challenge: Safari Image Classification
#
# Hopefully this notebook has shown you the main steps in training and evaluating a CNN. Why not put what you've learned into practice with our Safari image classification challenge in the [/challenges/05 - Safari CNN Challenge.ipynb](./challenges/05%20-%20Safari%20CNN%20Challenge.ipynb) notebook?
#
# > **Note**: The time to complete this optional challenge is not included in the estimated time for this exercise - you can spend as little or as much time on it as you like!
| 05b - Convolutional Neural Networks (PyTorch).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# +
import pandas as pd
import numpy as np
import lightgbm as lgb
import gc
from sklearn.model_selection import train_test_split,RandomizedSearchCV
from sklearn.metrics import confusion_matrix,classification_report,accuracy_score
from sklearn.preprocessing import StandardScaler
import joblib
import time
np.random.seed(2020)
def lgb_model_age(train_x, test_x,train_y, test_y):
model = lgb.LGBMClassifier (objective = 'multiclass',
num_class = 10,
n_estimators = 200
)
# model.fit(train_x,train_y,early_stopping_rounds=100,eval_set=eval_set=[(train_x,train_y),(test_x,test_y)],verbose = 10)
#设定搜索的xgboost参数搜索范围,值搜索XGBoost的主要6个参数
param_dist = {
'learning_rate':np.linspace(0.01,0.5,20),
'subsample':np.linspace(0.1,0.9,10),
'colsample_bytree':np.linspace(0.1,0.9,10),
'num_leaves' : range(32,128,6),
'reg_alpha' : np.linspace(0,0.1,10),
'reg_lambda' : np.linspace(0,0.1,10)
}
#RandomizedSearchCV参数说明,clf1设置训练的学习器
#param_dist字典类型,放入参数搜索范围
#scoring = 'neg_log_loss',精度评价方式设定为“neg_log_loss“
#n_iter=300,训练300次,数值越大,获得的参数精度越大,但是搜索时间越长
#n_jobs = -1,使用所有的CPU进行训练,默认为1,使用1个CPU
SearchCV = RandomizedSearchCV(model,param_dist,cv = 5,scoring = 'neg_log_loss',n_iter=10,n_jobs = -1,verbose = 10)
#在训练集上训练
SearchCV.fit(train_x,train_y)
# # 模型存储
joblib.dump(SearchCV, 'w2v_lgb_age_SearchCV.pkl')
# # 模型预测
y_t_pred = SearchCV.predict(test_x)
# print(model.get_score(importance_type='weight'))
cm = confusion_matrix(test_y, y_t_pred)
np.set_printoptions(precision=3) # 显示精度
cm_normalized = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis] # 将样本矩阵转化为比率
print('age************************************')
print('confusion_matrix is \n {:} \n '.format(cm_normalized))
print('test acc is \n {:} \n '.format(np.sum(test_y==y_t_pred)/len(test_y)))
print(classification_report(test_y,y_t_pred))
print('accuracy is %f , sen is %f,spe is %f ' % (accuracy_score(test_y, y_t_pred) * 100,
cm_normalized[0][0],cm_normalized[1][1] ))
return accuracy_score(test_y, y_t_pred)
def lgb_model_gender(train_x, test_x,train_y, test_y):
model = lgb.LGBMClassifier (objective = 'binary',
n_estimators = 100
)
# model.fit(train_x,train_y,early_stopping_rounds=100,eval_set=eval_set=[(train_x,train_y),(test_x,test_y)],verbose = 10)
#设定搜索的xgboost参数搜索范围,值搜索XGBoost的主要6个参数
param_dist = {
'learning_rate':np.linspace(0.01,0.5,20),
'subsample':np.linspace(0.1,0.9,10),
'colsample_bytree':np.linspace(0.1,0.9,10),
'num_leaves' : range(32,128,6),
'reg_alpha' : np.linspace(0,0.1,10),
'reg_lambda' : np.linspace(0,0.1,10)
}
#RandomizedSearchCV参数说明,clf1设置训练的学习器
#param_dist字典类型,放入参数搜索范围
#scoring = 'neg_log_loss',精度评价方式设定为“neg_log_loss“
#n_iter=300,训练300次,数值越大,获得的参数精度越大,但是搜索时间越长
#n_jobs = -1,使用所有的CPU进行训练,默认为1,使用1个CPU
SearchCV = RandomizedSearchCV(model,param_dist,cv = 5,scoring = 'neg_log_loss',n_iter=10,n_jobs = -1,verbose = 10)
#在训练集上训练
SearchCV.fit(train_x,train_y)
# # 模型存储
joblib.dump(SearchCV, 'w2v_lgb_gender_SearchCV.pkl')
# # 模型预测
y_t_pred = SearchCV.predict(test_x)
# print(model.get_score(importance_type='weight'))
cm = confusion_matrix(test_y, y_t_pred)
np.set_printoptions(precision=3) # 显示精度
cm_normalized = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis] # 将样本矩阵转化为比率
print('gender************************************')
print('confusion_matrix is \n {:} \n '.format(cm_normalized))
print('test acc is \n {:} \n '.format(np.sum(test_y==y_t_pred)/len(test_y)))
print(classification_report(test_y,y_t_pred))
print('accuracy is %f , sen is %f,spe is %f ' % (accuracy_score(test_y, y_t_pred) * 100,
cm_normalized[0][0],cm_normalized[1][1] ))
return accuracy_score(test_y, y_t_pred)
def load_data():
# user
data = pd.read_csv('w2v_feat_data/train_data.csv')
data = data.head(200000)
label = data[['age','gender']]
data = data.drop(['user_id','age','gender'],axis = 1)
return data,label
# -
data,label = load_data()
#划分age的训练和测试数据
train_x, test_x, train_y, test_y = train_test_split(data, label, test_size=0.8,random_state=2020)
# +
# star = time.time()
# acc_age = lgb_model_age(train_x, test_x,train_y['age'], test_y['age'])
# end = time.time()
# print('time spend ',end-star)
star = time.time()
acc_gender = lgb_model_gender(train_x, test_x,train_y['gender'], test_y['gender'])
end = time.time()
print('time spend ',end-star)
# print(acc_age+acc_gender)
# -
# 还是采用贝叶斯方法优化一下,大概取10,0000样本就可以
| lgb_w2v/w2v_lgb_randoncv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.0 64-bit (''Data-Analytics-and-Basics-of-Artificial-In-rUWBvdkx'':
# venv)'
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from IPython.display import display
df = pd.read_excel("data/tt.xlsx")
df.head()
# +
levels = {
1: "Peruskoulu",
2: "Toinen aste",
3: "Korkeakoulu",
4: "Ylempi korkeakoulu"
}
x = df["koulutus"].value_counts().sort_index().rename(index=levels).to_frame().rename(columns={"koulutus": "Lukumäärä"})
x["%"] = x["Lukumäärä"].apply(lambda n: n / len(df) * 100)
x
# -
x = df["koulutus"].value_counts().sort_index(ascending=False).rename(index=levels).to_frame().rename(columns={"koulutus": "Lukumäärä"})
x.plot(kind="barh", width=.5, legend=False)
| homework/w38/e01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.0 64-bit
# metadata:
# interpreter:
# hash: ac2eaa0ea0ebeafcc7822e65e46aa9d4f966f30b695406963e145ea4a91cd4fc
# name: python3
# ---
# The Happy Planet Index (HPI) is a measure of sustainable wellbeing. It compares how efficiently residents of different countries are using natural resources to achieve long, high wellbeing lives.
#
# Happy Planet Index = ( Life expectancy x Experienced wellbeing x Inequality of outcomes ) / Ecological Footprint
#
# [http://happyplanetindex.org/](http://happyplanetindex.org/)
import pandas as pd
import lux
df = pd.read_csv("https://raw.githubusercontent.com/lux-org/lux-datasets/master/data/hpi.csv")
df
# Now that we have seen some data with high correlation, let's rename and drop some similar columns
df = df.drop(columns=["AverageWellBeing", "AverageLifeExpectancy"])
df.rename(columns={"InequalityAdjustedWellbeing": "Average Wellbeing", "InequalityAdjustedLifeExpectancy": "Life Expectancy"})
df.intent = ["HappyLifeYears"]
df
| notebooks/exploratory_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + jupyter={"outputs_hidden": false}
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# -
from src.exploration.eda import *
# ## Loading Data
# mapping from id to emoji
mapping = pk.load(open(MAPPING_PATH, 'rb'))
# mapping from emoji to id
inv_map = {v: k for k, v in mapping.items()}
phraseVecModel = Phrase2Vec.from_word2vec_paths(300, W2V_PATH, E2V_PATH)
e2v = phraseVecModel.emojiVecModel
w2v = phraseVecModel.wordVecModel
# ## Emojis Descriptions Coherency
# <div class="alert alert-success">
# In order to assess whether the used features space is meaningful, we compute a dispersion metric across the multiple descriptions present for a single emoji.
# </div>
# +
desc_words_df = get_desc_emojis_df(phraseVecModel)
desc_words_df
# -
grouped_desc_df = gather_descs_vecs(desc_words_df,inv_map)
plot_num_desc_per_emoji(grouped_desc_df)
N = grouped_desc_df.shape[0]
N_1 = grouped_desc_df[grouped_desc_df.length ==1].shape[0]
print(f"Proportion of emojis with a single description: {N_1/N*100:.2f} %")
grouped_desc_df['dispersion'] = grouped_desc_df['vec'].apply(dispersion)
em_coherency_df = (grouped_desc_df[grouped_desc_df.dispersion > 0]
.sort_values('dispersion'))
# ### Most coherent emojis
best_emojis = em_coherency_df.head(5)
display_emoji_desc(best_emojis)
# ### Least Coherent Emojis
# TODO:
# * correlation matrixes between the subspaces of the vectors of the words in the description
# * try tfidf and stopwords removal
# * valuable information provided users
#
# First analyze the crowdsourcing output (dispersion for each emoji among humans)
# CORRELATION MATRIX ==> det == spread + ?random noise
worst_emojis = em_coherency_df.tail(5).iloc[::-1]
display_emoji_desc(worst_emojis)
# <div class="alert alert-success">
# The "worst emojis" rows seems to indicate that long descriptions don't always show an addition-friendly behavior: indeed, the semantic sense of each word added could land anywhere in the feature space. <br>
#
# Instead, using a method based on **crowdsourcing** coupled with a **tf-idf** selection could lead to better results.
# </div>
em_coherency_df.dispersion.hist(bins=40)
# <div class="alert alert-success">
# TODO : find the origin of this bimodal distribution
# </div>
# ## Similarity with words
# +
emoji_df = get_emoji_df(e2v,mapping)
face_emojis_df = emoji_df[emoji_df['em'].isin(emotions_faces)].copy()
face_emojis_df['top_sim10'] = face_emojis_df['em'].apply(lambda x:[i[0] for i in e2v.similar_by_word(x)])
face_emojis_df['top_sim5_faces'] = face_emojis_df['em'].apply(lambda em: get_10_faces(em,e2v))
face_emojis_df
# -
# ## T-SNE
# <div class="alert alert-success">
# T-sne results could be reproduced as in the emoji2vec paper.
# </div>
# <img src="../results/emojivec_eda/tsne/emojis_tsne.jpeg"></div>
# <div class="alert alert-success">
# Emotions Emojis were computed in order to check how the emotions behave in this feature space
# </div>
# <img src="../results/emojivec_eda/tsne/faces_tsne.jpeg"></div>
| notebooks/1_emojivec_EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# %reload_ext autoreload
# %autoreload 2
from synthetizer import anonymization_pipeline, RESOURCE_NAMES
from synthetizer.tools.paths import DATA_PATH
anonymization_pipeline(
resource_names = RESOURCE_NAMES,
id_suffix = "",
all_pages = True,
verbose = True,
output_dir = DATA_PATH,
)
| notebooks/demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from ipywidgets import interact
from matplotlib import pyplot as plt
import ffmpeg
import ipywidgets as widgets
import numpy as np
probe = ffmpeg.probe('in.mp4')
video_info = next(s for s in probe['streams'] if s['codec_type'] == 'video')
width = int(video_info['width'])
height = int(video_info['height'])
num_frames = int(video_info['nb_frames'])
# +
out, err = (
ffmpeg
.input('in.mp4')
.output('pipe:', format='rawvideo', pix_fmt='rgb24')
.run(capture_stdout=True)
)
video = (
np
.frombuffer(out, np.uint8)
.reshape([-1, height, width, 3])
)
@interact(frame=(0, num_frames))
def show_frame(frame=0):
plt.imshow(video[frame,:,:,:])
# +
from io import BytesIO
from PIL import Image
def extract_frame(stream, frame_num):
while isinstance(stream, ffmpeg.nodes.OutputStream):
stream = stream.node.incoming_edges[0].upstream_node.stream()
out, _ = (
stream
.filter_('select', 'gte(n,{})'.format(frame_num))
.output('pipe:', format='rawvideo', pix_fmt='rgb24', vframes=1)
.run(capture_stdout=True, capture_stderr=True)
)
return np.frombuffer(out, np.uint8).reshape([height, width, 3])
def png_to_np(png_bytes):
buffer = BytesIO(png_bytes)
pil_image = Image.open(buffer)
return np.array(pil_image)
def build_graph(
enable_overlay, flip_overlay, enable_box, box_x, box_y,
thickness, color):
stream = ffmpeg.input('in.mp4')
if enable_overlay:
overlay = ffmpeg.input('overlay.png')
if flip_overlay:
overlay = overlay.hflip()
stream = stream.overlay(overlay)
if enable_box:
stream = stream.drawbox(
box_x, box_y, 120, 120, color=color, t=thickness)
return stream.output('out.mp4')
def show_image(ax, stream, frame_num):
try:
image = extract_frame(stream, frame_num)
ax.imshow(image)
ax.axis('off')
except ffmpeg.Error as e:
print(e.stderr.decode())
def show_graph(ax, stream, detail):
data = ffmpeg.view(stream, detail=detail, pipe=True)
image = png_to_np(data)
ax.imshow(image, aspect='equal', interpolation='hanning')
ax.set_xlim(0, 1100)
ax.axis('off')
@interact(
frame_num=(0, num_frames),
box_x=(0, 200),
box_y=(0, 200),
thickness=(1, 40),
color=['red', 'green', 'magenta', 'blue'],
)
def f(
enable_overlay=True,
enable_box=True,
flip_overlay=True,
graph_detail=False,
frame_num=0,
box_x=50,
box_y=50,
thickness=5,
color='red'):
stream = build_graph(
enable_overlay,
flip_overlay,
enable_box,
box_x,
box_y,
thickness,
color
)
fig, (ax0, ax1) = plt.subplots(1, 2, figsize=(15,4))
plt.tight_layout()
show_image(ax0, stream, frame_num)
show_graph(ax1, stream, graph_detail)
# -
| examples/ffmpeg-numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Consider a measurable space $(\Omega, \mathcal{F})$. A map
# $Q\colon\Omega \times \mathcal{F} \to [0, +\infty)$ is a transition
# kernel if
#
# * $Q(\cdot, A)$ is measurable map for any $A \in \mathcal{F}$
#
# * $Q(\omega, \cdot)$ is a finite measure on $(\Omega, \mathcal{F})$
# for any $\omega \in \Omega$
#
# For any measurable map $f$ we can define a measurable map
#
# $$
# T f
# \colon x \mapsto \int f(\omega) \, Q(x, d\omega)
# \,. $$
#
#
# For a measure $\lambda$ on $(\Omega, \mathcal{F})$ we can similarly
# define a measure
#
# $$
# T^* \lambda
# \colon A \mapsto \int Q(\omega, A) \lambda(d\omega)
# \,. $$
#
# A dual pairing:
#
# $$
# \langle
# f, \lambda
# \rangle = \int f(\omega) \lambda(d\omega)
# \,. $$
#
# We can show that $
# \langle
# T f, \lambda
# \rangle = \langle
# f, T^* \lambda
# \rangle
# $ via Fubini (?) theorem:
#
# $$
# \int T f(x) \lambda(dx)
# = \int \int f(\omega) \, Q(x, d\omega) \lambda(dx)
# % = \iint f(\omega) \lambda(dx) Q(x, d\omega)
# = \int f(\omega) \int \lambda(dx) Q(x, d\omega)
# = \int f(\omega) (T^* \lambda)(d\omega)
# \,. $$
# +
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
# -
import torch
import torch.nn.functional as F
import gym
# A display device
# +
from IPython.display import clear_output
from time import sleep
def display(env, fps=15):
if fps > 0:
clear_output(wait=True)
env.render()
sleep(1. / fps)
# -
# Environment simulation loop:
# +
from tools.delayed import DelayedKeyboardInterrupt
def run(env, policy, *, fps=15):
state, terminate = env.reset(), False
display(env, fps)
with DelayedKeyboardInterrupt("ignore") as stop:
while not (terminate or stop):
# take an action and get a response form the environment
action = policy(env, state)
state_prime, reward, terminate, info = env.step(action)
# return the result
yield state, action, reward, state_prime, terminate
state = state_prime
# render the enviroment
display(env, fps)
env.close()
# -
# <br>
# +
from gym.envs.toy_text import FrozenLakeEnv, CliffWalkingEnv
env = FrozenLakeEnv(map_name="8x8", is_slippery=True)
# env = gym.make('CliffWalking-v0')
# -
# <br>
# ### Random policy
def random_policy(env):
return {
state: [(1. / env.nA, action) for action in kernel]
for state, kernel in env.P.items()
}
# +
def random(env, state=None):
return env.action_space.sample()
episode = [*run(env, random)]
# -
# <br>
# ## From Whiteson lecture on MLSS 2019
# MDP -- classical formal model of a sequential decision problem:
#
# * fully-observable, stationary, and possibly stochastic environment
#
# * discrete states $S$ and actions $A_s$ for each $s \in S$
#
# * transition kernel $s\to z \colon z\sim q(z\mid s, a)$ on $S$
#
# * reward distributuion $q(r\mid s, a, s')$ when transitioning $s \to s'$ under $a$
#
# * aplanning horizon or a discount factor $\gamma \in (0, 1)$
# Markov property
# $p(z_{t+1}, r_{t+1}\mid s_t, a_t) = p(z_{t+1}, r_{t+1}\mid s_t, a_t, s_{:t}, a_{:t})$
#
# * Reactive policies $a\sim \pi(a\mid s)$
# * deterministic policies
#
# **State-value** of a policy $
# v^\pi(s) = \mathbb{E}_\pi \bigl(
# \sum_{k\geq 1} \gamma r_{t+k+1}
# \big\vert s_t = s
# \bigr)
# $ and **action-value** $
# Q^\pi(s, a) = \mathbb{E}_\pi \bigl(
# \sum_{k\geq 1} \gamma r_{t+k+1}
# \big\vert s_t = s, a_t = a
# \bigr)
# $
# Bellman fixed-point equation for $v^\pi$:
# $$
# v^\pi(s)
# = \mathbb{E}_{a\sim \pi(s)} \mathbb{E}_{s' \sim q(s'\mid s, a)}
# r(s, a, s') + \gamma v^\pi(s')
# \,, $$
#
# and for $q^\pi$
#
# $$
# q^\pi(s, a)
# = \mathbb{E}_{s' \sim q(s'\mid s, a)}
# r(s, a, s') + \gamma \mathbb{E}_{a'\sim \pi(s')} q^\pi(s', a')
# = \mathbb{E}_{s' \sim q(s'\mid s, a)}
# r(s, a, s') + \gamma v^\pi(s')
# \,. $$
# Policies can be partially oredered by their value function. And
# all optimal policies share the same optimal state valeu function
# $v^*(\cdot) = \max_\pi v^\pi(\cdot)$.
# The Bellamn optimality conditions for $v^*$ and are
#
# $$
# v^*(s)
# = \max_{a\in A_s} \mathbb{E}_{s' \sim q(s'\mid s, a)}
# r(s, a, s') + \gamma v^*(s')
# \,, $$
#
# and
#
# $$
# q^*(s, a)
# = \mathbb{E}_{z \sim q(s'\mid s, a)}
# r(s, a, s') + \gamma \max_{a'\in A_{s'}} q^*(s', a')
# \,, $$
#
# respectively. The optimal policy is greedy with respect to $q$:
#
# $$
# \pi^*(s)
# = \delta_{a^*_s}
# \,, \text{ for }
# a^*_s = \arg\max_{a\in A_s} q^*(s, a)
# \,. $$
# <br>
# ## Policy Evaluation
# Bellman operator for a policy $\pi\colon \mathcal{S} \to \Delta_A$
# $$
# T_\pi(v)
# \colon s \mapsto \mathbb{E}_{a\sim \pi(a\mid s)}
# \mathbb{E}_{z\sim q(z\mid s, a)} r(s, a, z) + \gamma v(z)
# \,. $$
# +
def expected_state_reward(states, value, gamma=1.0):
# kernel -- list of next state-rewards with probabilities
return sum(
prob * (reward + gamma * value[state])
for prob, state, reward, term in states
)
def expected_action_reward(actions, kernel, value, gamma=1.0):
# policy -- list of actions with probabilities
return sum(
prob * expected_state_reward(kernel[action], value, gamma)
for prob, action in actions
)
# -
# The policy evaluation is performed via the fixed point iterations:
#
# * repeat $v_{t+1} \leftarrow T_\pi(v_t)$ until convergence in $\|\cdot\|_\infty$
#
def evaluate_policy(env, policy, gamma=1.0, atol=1e-8):
value, delta = {state: 0. for state in env.P}, float("+inf")
while delta > atol:
Tv = {
state: expected_action_reward(policy[state], kernel, value, gamma)
for state, kernel in env.P.items()
}
delta = max(abs(a - b) for a, b in zip(Tv.values(), value.values()))
value = Tv
return value
# The $q$-function of $v$ is
# $$
# q_\infty(s, a)
# = \mathbb{E}_{s'\sim q(s'\mid s, a)}
# r(s, a, s') + \gamma v_\infty(s')
# \,. $$
def q_fun(env, value, gamma=1.0):
return {
state: {
action: expected_state_reward(states, value, gamma)
for action, states in kernel.items()
}
for state, kernel in env.P.items()
}
# Let's evaluate a random exploration policy:
# +
gamma = 0.9
policy = random_policy(env)
value = evaluate_policy(env, policy, gamma=gamma)
# -
value
# The stationary $v$-function induces the following policy (consistent with it):
# $$
# \pi(s)
# = \delta_{a_s}
# \,, \text{ for } a_s = \arg \max_{a \in A_s} Q(s, a)
# \,. $$
def get_greedy_from_q(q):
def _policy(env, state):
actions, expected = zip(*q[state].items())
return actions[np.argmax(expected)]
return _policy
greedy = get_greedy_from_q(q_fun(env, value, gamma=gamma))
episode = [*run(env, greedy)]
# <br>
# ## Policy improvement
# The fixed point $v^\pi$ is the true value function of $\pi$. The associated $q$ function ca be used to reason about improvements in the policy $\pi$:
# if at some $s\in S$ we have $q^\pi(s, a_s) > v^\pi(s)$ for some $a_s \in A_s$ then the new policy $\hat{\pi}(\cdot) = \pi(\cdot)$ but $\hat{\pi}(s) = \delta_{a_s}$ is strictly better than $\pi$ (w.r.t $v$-function)
# Applying this to all states yields the **greedy** policy improvement:
#
# $$
# \pi_{t+1}(s) \in \arg\max_{a\in A_s} q^{\pi_t}(s, a)
# \,. $$
def better_policy(q):
policy = {}
for state, value in q.items():
# put equal mass on the actions with the maximal expected reward
v_max = max(value.values())
action = [a for a, v in value.items() if v >= v_max]
policy[state] = [(1. / len(action), a) for a in action]
return policy
# If $\pi_{t+1} = \pi_t$ then $v^{\pi_t} = v^{\pi_{t+1}} = v$, which
# satisfies the Bellamn Optimiality principle:
#
# * $T(v) = v$ for
# $$
# T(v)
# \colon S \to \mathbb{R}
# \colon s \mapsto \max_{a\in A_s}
# \mathbb{E}_{z\sim q(s'\mid s, a)} r(s, a, s') + \gamma v(s')
# \,. $$
def policy_iteration(env, gamma=1.0, atol=1e-8):
policy = random_policy(env)
value, delta = evaluate_policy(env, policy, gamma), float("+inf")
while delta > atol:
policy = better_policy(q_fun(env, value, gamma))
new = evaluate_policy(env, policy, gamma)
delta = max(abs(a - b) for a, b in zip(new.values(), value.values()))
value = new
return value, policy
value, policy = policy_iteration(env, gamma=gamma)
# Use a truly stochastic policy
def get_random_from_pi(policy):
def _policy(env, state):
probs, actions = zip(*policy[state])
return np.random.choice(actions, p=probs)
return _policy
episode = [*run(env, get_random_from_pi(policy))]
# Takes a stochastic policy and uses MAP action prediction
def get_greedy_from_pi(policy):
def _policy(env, state):
probs, actions = zip(*policy[state])
return actions[np.argmax(probs)]
return _policy
episode = [*run(env, get_greedy_from_pi(policy))]
# <br>
# ## Value iteration
# The operator in Bellaman's optimiality conditions acts on $v$ thus
#
# $$
# T(v)
# \colon S \to \mathbb{R}
# \colon s \mapsto \max_{a\in A_s} \mathbb{E}_{s'\sim q(s'\mid s, a)}
# r(s, a, s') + \gamma v(s')
# \,. $$
def optimal_action_reward(kernel, value, gamma=1.0):
return max(
expected_state_reward(states, value, gamma)
for action, states in kernel.items()
)
# It can be shown that $T$ is a contraction mapping w.r.t $\|\cdot\|_\infty$ for $\gamma \in (0, 1)$,
# and thus the fixed point iteration converges to a $v_*$:
# * repeat $v_{t+1} \leftarrow T(v_t)$ until convergence in $\|\cdot\|_\infty$
def value_iteration(env, gamma=1.0):
value, delta = {state: 0. for state in env.P}, float("+inf")
while delta > 1e-8:
# compute the operator
new = {state: optimal_action_reward(kernel, value, gamma)
for state, kernel in env.P.items()}
delta = max(abs(a - b) for a, b in zip(value.values(), new.values()))
value = new
return value
# Compute the $q$-function implied by the converged value function:
# $$
# Q(s, a)
# = \mathbb{E}_{z\sim q(z\mid s, a)} r(s, a, s') + \gamma v^*(z)
# \,. $$
# +
value = value_iteration(env, gamma=gamma)
q = q_fun(env, value, gamma=gamma)
# -
# The optimal value function implies an optimal (greedy) poilcy:
#
# $$
# \pi(s) \in \arg\max_{a\in A_s}
# \mathbb{E}_{z\sim q(z\mid s, a)} r(s, a, z) + \gamma v^*(z)
# \,. $$
episode = [*run(env, get_greedy_from_q(q))]
assert False
# ## Monte Carlo methods
# Provides amethod for finding an optimal policy without knowing the underlying MDP:
#
# * Learn $v^\pi$ without a model
#
# * unlike DP, MC uses the entire episode, and does not bootstrap (with $v^\pi(s')$
# * unlike DP, MC observes only one choice at each state $s\in S$
#
# Learn $q^\pi$ by averaging returns obtained when following $\pi$ after taking $a \in A_s$ at $s$:
# 1. (**Monte Carlo**) policy evaluation: use MC method to get $q^\pi$
# 2. (**greedy**) policy improvement: refine $\pi(s) \leftarrow g(s) = \arg \max_{a\in A_s} q^\pi(s, a)$
# * soften by mixing with a uniform policy:
# $
# \pi(a\vert s) = (1 - \varepsilon) \, \delta_{g(s)}(a) + \varepsilon \, \mathrm{U}_{A_s}(a)
# $
#
# PI theorem guarantees that and $\varepsilon$-greedy policy improves soft $\varepsilon$-greedy policy.
def soften(policy, epsilon=1e-1):
def _policy(env, state):
if np.random.uniform(1) <= epsilon:
return random(env, state)
return policy(env, state)
return _policy
# ### Off-policy MC control
#
# * **estimation policy** is evaluated on samples from
# **behaviour policy**, provided it is sufficiently exploratory
#
# * use importance sampling to re-weight returns:
# $
# \mathbb{E}_{z \sim P} h
# = \mathbb{E}_{z \sim Q} w \, h
# % \mathbb{E}_{z \sim P(z)} h(z)
# % = \mathbb{E}_{z \sim Q(z)} w_z h(z)
# $
# for $w = \tfrac{dP}{dQ}$ provided $P \ll Q$
# * $P$ comes from the estimation policy, $Q$ -- from the behavioural policy
# $$
# v^\pi
# \approx \frac{
# \hat{\mathbb{E}}_{i\sim D_s} \tfrac{p_i(s)}{p_i'(s)} R_i(s)
# }{
# \hat{\mathbb{E}}_{i\sim D_s} \tfrac{p_i(s)}{p_i'(s)}
# }
# \,, $$
# for
# $$
# \frac{p_i(s)}{p_i'(s)}
# = \frac{
# \prod_{k=t}^{T_i(s)-1} \pi(a_k \vert s_k) q(s_{k+1}\vert a_k, s_k)
# }{
# \prod_{k=t}^{T_i(s)-1} \pi'(a_k \vert s_k) q(s_{k+1}\vert a_k, s_k)
# }
# = \prod_{k=t}^{T_i(s)-1} \frac{\pi(a_k \vert s_k)}{\pi'(a_k \vert s_k)}
# \,, $$
# ### TD(0) algorithm
#
# Value function estimation from the experience (given by simulating from the behavioural policy)
#
# * $v(s_t) \leftarrow v(s_t) + \alpha (R_t + \gamma v(s_{t+1}) - v(s_t))$
# -- bootstrap from the existing estimate
#
# Samples and bootstraps
# +
from collections import defaultdict
def td_predict(env, policy, n_episodes=100, alpha=0.95, gamma=1.0, fps=15):
value = defaultdict(float)
# for each episode ...
with DelayedKeyboardInterrupt("ignore") as stop:
for n_episode in range(n_episodes):
# (lazily) play out the policy ...
episode = run(env, policy, fps=fps)
# run TD(0) updates ...
for state, action, reward, next_state, terminal in episode:
# ... for the value function of the policy
bootstrap = reward + gamma * value[next_state]
value[state] += alpha * (bootstrap - value[state])
if stop:
break
return value
# -
piter = get_random_from_pi(policy)
# policy = soften(get_greedy_from_q(q), epsilon=1e-1)
policy = soften(piter, epsilon=1e-1)
value = td_predict(env, policy, fps=0)
# ### TD: control SARSA
#
# Sarsa algorithm
def sarsa_evaluate(env, n_episodes=10000, alpha=0.01, gamma=1.0, fps=15):
q_fun = defaultdict(lambda: defaultdict(float))
def q_policy(env, state):
if q_fun[state]:
actions, expected = zip(*q_fun[state].items())
return actions[np.argmax(expected)]
return random(env, state)
# for each episode ...
for n_episode in range(n_episodes):
_feedback = soften(q_policy, epsilon=1e-2)
# play out the policy ...
episode = run(env, _feedback, fps=fps)
# run TD(0) updates ...
state, action, reward, next_state, terminal = next(episode)
while not terminal:
# ... for the q-function function of the policy
next_state, next_action, *rest = next(episode)
bootstrap = reward + gamma * q_fun[next_state][next_action]
q_fun[state][action] += alpha * (bootstrap - q_fun[state][action])
state, action = next_state, next_action
reward, next_state, terminal = rest
return q_fun
q_fun = sarsa_evaluate(env, fps=0)
q_fun
grp = {}
for (k, a), v in q_fun.items():
grp.setdefault(k , []).append((a, v))
q_fun = {k: v for k, v in zip(grp, map(dict, grp.values()))}
q_fun
assert False
# <br>
# ## Policy improvement and Policy Iteration
from matplotlib.collections import LineCollection
# +
s, *_ = zip(*episode)
segs = np.c_[np.unravel_index(s, (8, 8))]
# -
assert False
# <br>
# +
# # !pip install gym
# -
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
import torch
import torch.nn.functional as F
import gym
from gym.envs.classic_control import MountainCarEnv
# +
from gym import spaces
class ModifiedMountainCarEnv(MountainCarEnv):
def __init__(self, goal_velocity = 0):
self.min_position = -2.5
self.max_position = 2.5
self.max_speed = 0.07
self.goal_position = 2.0
self.goal_velocity = goal_velocity
self.force = 0.001
self.gravity = 0.0025
self.low = np.array([self.min_position, -self.max_speed])
self.high = np.array([self.max_position, self.max_speed])
self.viewer = None
self.action_space = spaces.Discrete(3)
self.observation_space = spaces.Box(self.low, self.high, dtype=np.float32)
self.seed()
# -
env = ModifiedMountainCarEnv()
def run(env, agent, fps=15):
state, terminated = env.reset(), False
history = []
while not terminated:
state, reward, terminated, info = env.step(agent(state))
history.append((state, reward))
return history
class BaseAgent(object):
def __init__(self, env):
self.env = env
def reset(self):
pass
def update(self, state, action, reward, next_state, terminated=False):
pass
def __call__(self, state=None):
return env.action_space.sample()
space = env.observation_space
from gym.spaces import Space
shape = (51, 71)
state = 0, 0
unit = (state - space.low) / (space.high - space.low)
class Discretizer(object):
def __init__(self, space, shape):
assert isinstance(space, Box) and len(space.shape) == 1
assert space.is_bounded
if not isinstance(n_states, (list, tuple)):
shape = space.shape[0] * [shape]
assert shape == space.shape[0]
self.space, self.shape = space, shape
def to_ix(self, state, flatten=False):
unit = (state - space.low) / (space.high - space.low)
ix = (unit * shape + 0.5).astype(int)
if flatten:
return np.unravel_index(ix, shape=self.shape)
return ix
def from_ix(self, *index):
return np.array(index) * (space.high - space.low) + space.low
# +
def rescale(obs, env):
space = env.observation_space
return (obs - space.low) / (space.high - space.low)
class TabularQLearner(BaseAgent):
def __init__(self, env, n_states=51):
super().__init__(env)
self.n_states, self.n_actions = n_states, env.action_space.n
self.reset()
def reset(self):
self.q_table = torch.zeros(self.n_states, self.n_states, n_actions)
def __call__(self, state=None):
return env.action_space.sample()
# -
| coding/notes_on_rl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:python3.5]
# language: python
# name: conda-env-python3.5-py
# ---
# Import packages
# +
import pandas as pd
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import scale, StandardScaler, MinMaxScaler
from sklearn.metrics import r2_score, mean_squared_error, mean_absolute_error
from math import sqrt
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import rcParams
from keras.layers import Input, Dense, Lambda, concatenate
from keras.models import Model, load_model, Sequential, model_from_json
from keras import backend as K
from keras import metrics, optimizers, regularizers
from keras.objectives import categorical_crossentropy, mean_squared_error
from keras.callbacks import ModelCheckpoint, TensorBoard, EarlyStopping
# fix random seed for reproducibility
tf.set_random_seed(123)
np.random.seed(123)
# -
# Load data
# load Concrete_Data.xls dataset, X: features, contains concrete mixes formula
df_formula = pd.read_excel('Concrete_Data.xls')
df_epd = pd.read_excel('environmental_impact.xlsx')
X = df_formula.values[:,:7]
# Samples separated into 6 groups <br>
# Group 0: < 3d <br>
# Group 1: 7d <br>
# Group 2: 14d <br>
# Group 3: 28d <br>
# Group 4: 56d <br>
# Group 5: > 90d
day_to_idx = {1:0,
3:0,
7:1,
14:2,
28:3,
56:4,
90:5,
91:5,
100:5,
120:5,
180:5,
270:5,
360:5,
365:5}
# Convert indices to one hot
# +
indices = []
day_raw = df_formula.values[:,7]
for i, day in enumerate(day_raw):
indices.append(day_to_idx[day])
indices = np.array(indices)
one_hot_vecs = np.zeros((day_raw.shape[0], 6))
one_hot_vecs[np.arange(day_raw.shape[0]), np.array(indices)] = 1
# -
# Get indices of samples from different groups, for later usage
group_0_train = np.where(indices == 0)[0]
group_1_train = np.where(indices == 1)[0]
group_2_train = np.where(indices == 2)[0]
group_3_train = np.where(indices == 3)[0]
group_4_train = np.where(indices == 4)[0]
group_5_train = np.where(indices == 5)[0]
# Stack up group info, strength, and environmental impact
Y = np.hstack((one_hot_vecs, df_formula.values[:,8].reshape(-1,1), df_epd.values[:,:12]))
# Setup scaler for X and Y
scaler_X = MinMaxScaler().fit(X)
scaler_Y = MinMaxScaler().fit(Y)
X = scaler_X.transform(X)
Y = scaler_Y.transform(Y)
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.2, random_state=42)
# Define input tensor
# +
n_x = X.shape[1]
n_y = Y.shape[1]
n_z = 2
# Q(z|X,y) -- encoder
formula = Input(shape=(n_x,))
cond = Input(shape=(n_y,))
# -
# Initialize encoder layers
inputs = concatenate([formula, cond])
enc_hidden_1 = Dense(30, activation='relu')(inputs)
enc_hidden_2 = Dense(25, activation='relu')(enc_hidden_1)
z_mean = Dense(n_z)(enc_hidden_2)
z_log_var = Dense(n_z)(enc_hidden_2)
# Define sampling function
def sample_z(args):
z_mean, z_log_var = args
epsilon = K.random_normal(shape=(K.shape(z_mean)[0], n_z), mean=0., stddev=1.0)
return z_mean + K.exp(z_log_var / 2) * epsilon
# Sample z ~ Q(z|X,y)
z = Lambda(sample_z)([z_mean, z_log_var])
z_cond = concatenate([z, cond])
# +
# P(X|z,y) -- decoder
dec_layer_1 = Dense(25, activation='relu')
dec_layer_2 = Dense(30, activation='relu')
dec_out = Dense(n_x, activation='sigmoid') #, activation='sigmoid'
dec_hidden_1 = dec_layer_1(z_cond)
dec_hidden_2 = dec_layer_2(dec_hidden_1)
reconstructed = dec_out(dec_hidden_2)
# +
# end-to-end autoencoder
cvae = Model([formula, cond], reconstructed)
# encoder, from inputs to latent space
encoder = Model([formula, cond], z)
# +
z_for_gen = Input(shape=(n_z,))
z_cond_for_gen = concatenate([z_for_gen, cond])
dec_hidden_1_for_gen = dec_layer_1(z_cond_for_gen)
dec_hidden_2_for_gen = dec_layer_2(dec_hidden_1_for_gen)
reconstructed_for_gen = dec_out(dec_hidden_2_for_gen)
# generator, from latent space to reconstructed inputs
generator = Model([z_for_gen, cond], reconstructed_for_gen)
# -
# Define loss
def cvae_loss(feature, reconstructed):
reconstruction_loss = mean_squared_error(feature, reconstructed)
kl_loss = - 0.5 * K.mean(1 + z_log_var - K.square(z_mean) - K.exp(z_log_var), axis=-1)
return reconstruction_loss + kl_loss
# Compile model
adam = optimizers.Adam(lr=0.001, beta_1=0.9, beta_2=0.999, epsilon=1e-08)
cvae.compile(optimizer=adam, loss=cvae_loss)
cvae.summary()
n_epoch = 100
n_batch = 10
# Train model
# +
checkpointer = ModelCheckpoint(filepath = "model_autoencoder.h5",
verbose = 0,
save_best_only = True)
tensorboard = TensorBoard(log_dir = './logs',
histogram_freq = 0,
write_graph = True,
write_images = True)
history = cvae.fit([X_train, Y_train], X_train,
epochs=n_epoch,
batch_size=n_batch,
shuffle=True,
callbacks = [EarlyStopping(patience = 5)],
validation_data=([X_test, Y_test], X_test))
# -
# Generate new samples
num_of_samples = 60000
day_vecs = np.zeros((num_of_samples, 6))
day_idx = np.random.choice(6, num_of_samples)
group_0 = np.where(day_idx == 0)[0]
group_1 = np.where(day_idx == 1)[0]
group_2 = np.where(day_idx == 2)[0]
group_3 = np.where(day_idx == 3)[0]
group_4 = np.where(day_idx == 4)[0]
group_5 = np.where(day_idx == 5)[0]
day_vecs = np.zeros((num_of_samples, 6))
day_vecs[np.arange(num_of_samples), day_idx] = 1
np.random.seed(1)
strength_and_environmental = np.random.uniform(0, 1, (num_of_samples, 13))
mean = [0,0]
cov = [[1, 0], [0, 1]]
np.random.seed(1)
Z_sampling = np.random.multivariate_normal(mean, cov, num_of_samples)
Y_sampling = np.hstack((day_vecs, strength_and_environmental))
samples_scaled = generator.predict([Z_sampling, Y_sampling])
generated_samples = scaler_X.inverse_transform(samples_scaled)
| Conditional_Variational_Autoencoder_for_Concrete Design.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf-tutorial
# language: python
# name: tf-tutorial
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %config InlineBackend.figure_format = 'svg'
# %matplotlib inline
pd.set_option('display.max_rows', None)
df_1 = pd.read_csv('clean_data_1.csv')
df_1.head()
df_2 = pd.read_csv('clean_data_2.csv')
df_2.head()
df = pd.concat([df_1, df_2], ignore_index=True)
df
# Remove total mass > 4000
df = df[df['total_mass'] < 4000]
df = df.sort_values(by = 'zT')
df
df.info()
# ## Preparing dataset
# +
# Train Test Split
from sklearn.model_selection import train_test_split
# X = df.iloc[:, 2]
# y = df.iloc[:, 3]
# X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0)
train, test = train_test_split(df, test_size=0.3, random_state=0)
# +
train = train.sort_values(by = 'zT')
X_train = train.iloc[:, 2]
y_train = train.iloc[:, 3]
X_test = test.iloc[:, 2]
y_test = test.iloc[:, 3]
# -
# Transform data
X_train = X_train.to_numpy().reshape(-1, 1)
X_test = X_test.to_numpy().reshape(-1, 1)
y_train = y_train.to_numpy().reshape(-1, 1)
y_test = y_test.to_numpy().reshape(-1, 1)
# ## 1. Linear Regression
from sklearn.linear_model import LinearRegression
# Creating and Training the Model
linear_regressor = LinearRegression()
linear_regressor.fit(X_train, y_train)
# +
# Plot points and fit line for training data
plt.scatter(X_train, y_train, color='teal', edgecolors='black', label='Train')
plt.plot(X_train, linear_regressor.predict(X_train), color='grey', label='Linear regressor')
plt.title('Linear Regression')
plt.xlabel('zT')
plt.ylabel('total_mass')
# plot scatter points for test data
plt.scatter(X_test, y_test, color='red', edgecolors='black', label='Test')
plt.legend()
plt.show()
# -
# ### Evaluation
predictions = linear_regressor.predict(X_test)
# +
from sklearn.metrics import mean_absolute_error
from math import sqrt
rmse_linear_regressor = sqrt(mean_absolute_error(y_test, predictions))
rmse_linear_regressor
# -
# ## 2. Polynomial Regression
from sklearn.preprocessing import PolynomialFeatures
# From here. we are simply generating the matrix for X^0, X^1 and X^2
poly_reg = PolynomialFeatures(degree = 2)
X_poly = poly_reg.fit_transform(X_train)
# polynomial regression model
poly_reg_model = LinearRegression()
poly_reg_model.fit(X_poly, y_train)
X_plot = poly_reg.fit_transform(X_train)
y_pred = poly_reg_model.predict(X_plot)
# +
plt.scatter(X_train, y_train, color='teal', edgecolors='black', label='Train')
plt.plot(X_train, y_pred, color='grey', label='Polynmial regressor')
plt.title('Polynmial Regression with degree = 2')
plt.xlabel('zT')
plt.ylabel('total_mass')
# plot scatter points for test data
plt.scatter(X_test, y_test, color='red', edgecolors='black', label='Test')
plt.legend()
plt.show()
# -
# ### Evaluation
X_test_poly = poly_reg.fit_transform(X_test)
poly_predictions = poly_reg_model.predict(X_test_poly)
rmse_poly = sqrt(mean_absolute_error(y_test, poly_predictions))
rmse_poly
# ## 3. Simple Vector Regression
from sklearn.svm import SVR
from sklearn.preprocessing import StandardScaler
# +
# Performing feature scaling
scaled_X = StandardScaler()
scaled_y = StandardScaler()
scaled_X_train = scaled_X.fit_transform(X_train)
scaled_y_train = scaled_y.fit_transform(y_train)
scaled_X_test = scaled_X.fit_transform(X_test)
scaled_y_test = scaled_y.fit_transform(y_test)
# -
svr_regressor = SVR(kernel='rbf', gamma='auto')
svr_regressor.fit(scaled_X_train, scaled_y_train.ravel())
# +
plt.scatter(scaled_X_train, scaled_y_train, color='teal', edgecolors='black', label='Train')
plt.plot(scaled_X_train, svr_regressor.predict(scaled_X_train), color='grey', label='SVR')
plt.title('Simple Vector Regression')
plt.xlabel('zT')
plt.ylabel('total_mass')
# plot scatter points for test data
plt.scatter(scaled_X_test, scaled_y_test, color='red', edgecolors='black', label='Test')
plt.legend()
plt.show()
# -
# ### Evaluation
# +
svr_predictions = svr_regressor.predict(scaled_X_test)
rmse_svr = sqrt(mean_absolute_error(scaled_y_test, svr_predictions))
rmse_svr
# -
# ## 4. Decision Tree Regression
from sklearn.tree import DecisionTreeRegressor
# No need to perform feature scaling. Since it will get taken care by the library itself.
tree_regressor = DecisionTreeRegressor(random_state = 0)
tree_regressor.fit(X_train, y_train)
# +
X_grid = np.arange(min(X_train), max(X_train), 0.01)
X_grid = X_grid.reshape(len(X_grid), 1)
plt.scatter(X_train, y_train, color='teal', edgecolors='black', label='Train')
plt.plot(X_grid, tree_regressor.predict(X_grid), color='grey', label='Tree regressor')
plt.title('Tree Regression')
plt.xlabel('zT')
plt.ylabel('total_mass')
# plot scatter points for test data
plt.scatter(X_test, y_test, color='red', edgecolors='black', label='Test')
plt.legend()
plt.show()
# -
# ### Evaluation
# +
tree_predictions = tree_regressor.predict(X_test)
tree_predictions
rmse_tree = sqrt(mean_absolute_error(y_test, tree_predictions))
rmse_tree
# -
# ## 5. Random Forest Regression
from sklearn.ensemble import RandomForestRegressor
forest_regressor = RandomForestRegressor(n_estimators = 300, random_state = 0)
forest_regressor.fit(X_train, y_train.ravel())
# +
X_grid = np.arange(min(X_train), max(X_train), 0.01)
X_grid = X_grid.reshape(len(X_grid), 1)
plt.scatter(X_train, y_train, color='teal', edgecolors='black', label='Train')
plt.plot(X_grid, forest_regressor.predict(X_grid), color='grey', label='Random Forest regressor')
plt.title('Random Forest Regression')
plt.xlabel('zT')
plt.ylabel('total_mass')
# plot scatter points for test data
plt.scatter(X_test, y_test, color='red', edgecolors='black', label='Test')
plt.legend()
plt.show()
# -
# ### Evaluation
# +
forest_predictions = forest_regressor.predict(X_test)
rmse_forest = sqrt(mean_absolute_error(y_test, forest_predictions))
rmse_forest
# -
# ## 6. Neural Network Regression
# ## 7. LASSO Regression
from sklearn.linear_model import LassoCV
lasso = LassoCV()
lasso.fit(X_train, y_train.ravel())
# +
plt.scatter(X_train, y_train, color='teal', edgecolors='black', label='Actual observation points')
plt.plot(X_train, lasso.predict(X_train), color='grey', label='LASSO regressor')
plt.title('LASSO Regression')
plt.xlabel('zT')
plt.ylabel('total_mass')
# plot scatter points for test data
plt.scatter(X_test, y_test, color='red', edgecolors='black', label='Test')
plt.legend()
plt.show()
# -
# ### Evalulation
# +
lasso_predictions = lasso.predict(X_test)
rmse_lasso = sqrt(mean_absolute_error(y_test, lasso_predictions))
rmse_lasso
# -
# ## 8. Ridge Regression
from sklearn.linear_model import RidgeCV
ridge = RidgeCV()
ridge.fit(X_train, y_train)
# +
plt.scatter(X_train, y_train, color='teal', edgecolors='black', label='Train')
plt.plot(X_train, ridge.predict(X_train), color='grey', label='Ridge regressor')
plt.title('Ridge Regression')
plt.xlabel('zT')
plt.ylabel('total_mass')
# plot scatter points for test data
plt.scatter(X_test, y_test, color='red', edgecolors='black', label='Test')
plt.legend()
plt.show()
# -
# ### Evaluation
# +
ridge_predictions = ridge.predict(X_test)
rmse_ridge = sqrt(mean_absolute_error(y_test, ridge_predictions))
rmse_ridge
# -
# ## 9. ElasticNet Regression
from sklearn.linear_model import ElasticNetCV
elasticNet = ElasticNetCV()
elasticNet.fit(X_train, y_train.ravel())
# +
plt.scatter(X_train, y_train, color='teal', edgecolors='black', label='Train')
plt.plot(X_train, elasticNet.predict(X_train), color='grey',label='ElasticNet regressor')
plt.title('ElasticNet Regression')
plt.xlabel('zT')
plt.ylabel('total_mass')
# plot scatter points for test data
plt.scatter(X_test, y_test, color='red', edgecolors='black', label='Test')
plt.legend()
plt.show()
# -
# ### Evaluation
# +
elasticNet_predictions = elasticNet.predict(X_test)
rmse_elasticNet = sqrt(mean_absolute_error(y_test, elasticNet_predictions))
rmse_elasticNet
# -
| Thermoelectric/regression analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
#export
"""
Bare example of how this module works::
import k1lib
class CbA(k1lib.Callback):
def __init__(self):
super().__init__()
self.initialState = 3
def startBatch(self):
print("startBatch - CbA")
def startPass(self):
print("startPass - CbA")
class CbB(k1lib.Callback):
def startBatch(self):
print("startBatch - CbB")
def endLoss(self):
print("endLoss - CbB")
# initialization
cbs = k1lib.Callbacks()
cbs.add(CbA()).add(CbB())
model = lambda xb: xb + 3
lossF = lambda y, yb: y - yb
# training loop
cbs("startBatch"); xb = 6; yb = 2
cbs("startPass"); y = model(xb); cbs("endPass")
cbs("startLoss"); loss = lossF(y, yb); cbs("endLoss")
cbs("endBatch")
print(cbs.CbA) # can reference the Callback object directly
So, point is, you can define lots of :class:`Callback` classes that
defines a number of checkpoint functions, like ``startBatch``. Then,
you can create a :class:`Callbacks` object that includes Callback
objects. When you do ``cbs("checkpoint")``, this will execute
``cb.checkpoint()`` of all the Callback objects.
Pretty much everything here is built upon this. The core training loop
has nothing to do with ML stuff. In fact, it's just a bunch of
``cbs("...")`` statements. Everything meaningful about the training
loop comes from different Callback classes. Advantage of this is that you
can tack on wildly different functions, have them play nicely with each
other, and remove entire complex functionalities by commenting out a
single line."""
import k1lib, time, os, logging, numpy as np, matplotlib.pyplot as plt
from typing import Set, List, Union, Callable, ContextManager, Iterator
from collections import OrderedDict
#export
__all__ = ["Callback", "Callbacks", "Cbs"]
#export
class Callback:
r"""Represents a callback. Define specific functions
inside to intercept certain parts of the training
loop. Can access :class:`k1lib.Learner` like this::
self.l.xb = self.l.xb[None]
This takes x batch of learner, unsqueeze it at
the 0 position, then sets the x batch again.
Normally, you will define a subclass of this and
define specific intercept functions, but if you
want to create a throwaway callback, then do this::
Callback().withCheckpoint("startRun", lambda: print("start running"))
You can use :attr:`~k1lib.callbacks.callbacks.Cbs` (automatically exposed) for
a list of default Callback classes, for any particular needs.
**order**
You can also use `.order` to set the order of execution of the callback.
The higher, the later it gets executed. Value suggestions:
- 7: pre-default runs, like LossLandscape
- 10: default runs, like DontTrainValid
- 13: custom mods, like ModifyBatch
- 15: pre-recording mod
- 17: recording mods, like Profiler.memory
- 20: default recordings, like Loss
- 23: post-default recordings, like ParamFinder
- 25: guards, like TimeLimit, CancelOnExplosion
Just leave as default (10) if you don't know what values to choose.
**dependsOn**
If you're going to extend this class, you can also specify dependencies
like this::
class CbC(k1lib.Callback):
def __init__(self):
super().__init__()
self.dependsOn = {"Loss", "Accuracy"}
This is so that if somewhere, ``Loss`` callback class is temporarily
suspended, then CbC will be suspended also, therefore avoiding errors.
**Suspension**
If your Callback is mainly dormant, then you can do something like this::
class CbD(k1lib.Callback):
def __init__(self):
super().__init__()
self.suspended = True
def startBatch(self):
# these types of methods will only execute
# if ``self.suspended = False``
pass
def analyze(self):
self.suspended = False
# do something that sometimes call ``startBatch``
self.suspended = True
cbs = k1lib.Callbacks().add(CbD())
# dormant phase:
cbs("startBatch") # does not execute CbD.startBatch()
# active phase
cbs.CbB.analyze() # does execute CbD.startBatch()
So yeah, you can easily make every checkpoint active/dormant by changing
a single variable, how convenient. See over :meth:`Callbacks.suspend`
for more."""
def __init__(self):
self.l = None; self.cbs = None; self.suspended = False
self.name = self.__class__.__name__; self.dependsOn:Set[str] = set()
self.order = 10 # can be modified by subclasses. A smaller order will be executed first
def suspend(self):
"""Checkpoint, called when the Callback is temporarily suspended. Overridable"""
pass
def restore(self):
"""Checkpoint, called when the Callback is back from suspension. Overridable"""
pass
def __getstate__(self):
state = dict(self.__dict__)
del state["l"]; del state["cbs"]; return state
def __setstate__(self, state): self.__dict__.update(state)
def __repr__(self): return f"{self._reprHead}, can...\n{self._reprCan}"
@property
def _reprHead(self): return f"Callback `{self.name}`"
@property
def _reprCan(self): return """- cb.something: to get specific attribute "something" from learner if not available
- cb.withCheckpoint(checkpoint, f): to quickly insert an event handler
- cb.detach(): to remove itself from its parent Callbacks"""
def withCheckpoint(self, checkpoint:str, f:Callable[["Callback"], None]):
"""Quickly set a checkpoint, for simple, inline-able functions
:param checkpoint: checkpoints like "startRun"
:param f: function that takes in the Callback itself"""
setattr(self, checkpoint, lambda: f(self)); return self
def __call__(self, checkpoint):
if not self.suspended and hasattr(self, checkpoint):
return getattr(self, checkpoint)() != None
def attached(self):
"""Called when this is added to a :class:`Callback`. Overrides this to
do custom stuff when this happens."""
pass
def detach(self):
"""Detaches from the parent :class:`Callbacks`"""
self.cbs.remove(self.name); return self
#export
Cbs = k1lib.Object()
Callback.lossCls = k1lib.Object()
#export
class Timings:
"""List of checkpoint timings. Not intended to be instantiated by the end user.
Used within :class:`~k1lib.callbacks.callbacks.Callbacks`, accessible via
:attr:`Callbacks.timings` to record time taken to execute a single
checkpoint. This is useful for profiling stuff."""
@property
def state(self):
answer = dict(self.__dict__); answer.pop("getdoc", None); return answer
@property
def checkpoints(self) -> List[str]:
"""List of all checkpoints encountered"""
return [cp for cp in self.state if k1lib.isNumeric(self[cp])]
def __getattr__(self, attr):
if attr.startswith("_"): raise AttributeError()
self.__dict__[attr] = 0; return 0
def __getitem__(self, idx): return getattr(self, idx)
def __setitem__(self, idx, value): setattr(self, idx, value)
def plot(self):
"""Plot all checkpoints' execution times"""
plt.figure(dpi=100); checkpoints = self.checkpoints
timings = np.array([self[cp] for cp in checkpoints])
maxTiming = timings.max()
if maxTiming >= 1:
plt.bar(checkpoints, timings); plt.ylabel("Time (s)")
elif maxTiming >= 1e-3 and maxTiming < 1:
plt.bar(checkpoints, timings*1e3); plt.ylabel("Time (ms)")
elif maxTiming >= 1e-6 and maxTiming < 1e-3:
plt.bar(checkpoints, timings*1e6); plt.ylabel("Time (us)")
plt.xticks(rotation="vertical"); plt.show()
def clear(self):
"""Clears all timing data"""
for cp in self.checkpoints: self[cp] = 0
def __repr__(self):
cps = '\n'.join([f'- {cp}: {self[cp]}' for cp in self.checkpoints])
return f"""Timings object. Checkpoints:\n{cps}\n
Can...
- t.startRun: to get specific checkpoint's execution time
- t.plot(): to plot all checkpoints"""
#export
_time = time.time
class Callbacks:
def __init__(self):
self._l: k1lib.Learner = None; self.cbsDict = {}; self._timings = Timings()
@property
def timings(self) -> Timings:
"""Returns :class:`~k1lib.callbacks.callbacks.Timings` object"""
return self._timings
@property
def l(self) -> "k1lib.Learner":
""":class:`k1lib.Learner` object. Will be set automatically when
you set :attr:`k1lib.Learner.cbs` to this :class:`Callbacks`"""
return self._l
@l.setter
def l(self, learner):
self._l = learner
for cb in self.cbs: cb.l = learner
@property
def cbs(self) -> List[Callback]:
"""List of :class:`Callback`"""
return [*self.cbsDict.values()] # convenience method for looping over stuff
def _sort(self) -> "Callbacks":
self.cbsDict = OrderedDict(sorted(self.cbsDict.items(), key=(lambda o: o[1].order))); return self
def add(self, cb:Callback, name:str=None):
"""Adds a callback to the collection."""
if cb in self.cbs: cb.l = self.l; cb.cbs = self; return self
cb.l = self.l; cb.cbs = self; name = name or cb.name
if name in self.cbsDict:
i = 0
while f"{name}{i}" in self.cbsDict: i += 1
name = f"{name}{i}"
cb.name = name; self.cbsDict[name] = cb; self._sort()
self._appendContext_append(cb); cb("attached"); return self
def __contains__(self, e:str) -> bool:
"""Whether a specific Callback name is in this :class:`Callback`."""
return e in self.cbsDict
def remove(self, *names:List[str]):
"""Removes a callback from the collection."""
for name in names:
if name not in self.cbsDict: return print(f"Callback `{name}` not found")
cb = self.cbsDict[name]; del self.cbsDict[name]; cb("detached")
self._sort(); return self
def removePrefix(self, prefix:str):
"""Removes any callback with the specified prefix"""
for cb in self.cbs:
if cb.name.startswith(prefix): self.remove(cb.name)
return self
def __call__(self, *checkpoints:List[str]) -> bool:
"""Calls a number of checkpoints one after another.
Returns True if any of the checkpoints return anything at all"""
self._checkpointGraph_call(checkpoints)
answer = False
for checkpoint in checkpoints:
beginTime = _time()
answer |= any([cb(checkpoint) for cb in self.cbs])
self._timings[checkpoint] += _time() - beginTime
return answer
def __getitem__(self, idx:Union[int, str]) -> Callback:
"""Get specific cbs.
:param idx: if :class:`str`, then get the Callback with this specific name,
if :class:`int`, then get the Callback in that index."""
return self.cbs[idx] if isinstance(idx, int) else self.cbsDict[idx]
def __iter__(self) -> Iterator[Callback]:
"""Iterates through all :class:`Callback`."""
for cb in self.cbsDict.values(): yield cb
def __len__(self):
"""How many :class:`Callback` are there in total?"""
return len(self.cbsDict)
def __getattr__(self, attr):
if attr == "cbsDict": raise AttributeError(attr)
if attr in self.cbsDict: return self.cbsDict[attr]
else: raise AttributeError(attr)
def __getstate__(self):
state = dict(self.__dict__); del state["_l"]; return state
def __setstate__(self, state):
self.__dict__.update(state)
for cb in self.cbs: cb.cbs = self
def __dir__(self):
answer = list(super().__dir__())
answer.extend(self.cbsDict.keys())
return answer
def __repr__(self):
return "Callbacks:\n" + '\n'.join([f"- {cbName}" for cbName in self.cbsDict if not cbName.startswith("_")]) + """\n
Use...
- cbs.add(cb[, name]): to add a callback with a name
- cbs("startRun"): to trigger a specific checkpoint, this case "startRun"
- cbs.Loss: to get a specific callback by name, this case "Loss"
- cbs[i]: to get specific callback by index
- cbs.timings: to get callback execution times
- cbs.checkpointGraph(): to graph checkpoint calling orders
- cbs.context(): context manager that will detach all Callbacks attached inside the context
- cbs.suspend("Loss", "Cuda"): context manager to temporarily prevent triggering checkpoints"""
def withBasics(self):
"""Adds a bunch of very basic Callbacks that's needed for everything. Also
includes Callbacks that are not necessary, but don't slow things down"""
self.add(Cbs.CoreNormal()).add(Cbs.Profiler()).add(Cbs.Recorder())
self.add(Cbs.ProgressBar()).add(Cbs.Loss()).add(Cbs.Accuracy()).add(Cbs.DontTrainValid())
return self.add(Cbs.CancelOnExplosion()).add(Cbs.ParamFinder())
def withQOL(self):
"""Adds quality of life Callbacks."""
return self
def withAdvanced(self):
"""Adds advanced Callbacks that do fancy stuff, but may slow things
down if not configured specifically."""
return self.add(Cbs.HookModule().withMeanRecorder().withStdRecorder()).add(Cbs.HookParam())
#export
@k1lib.patch(Callbacks)
def _resolveDependencies(self):
for cb in self.cbs:
cb._dependents:Set[Callback] = set()
cb.dependsOn = set(cb.dependsOn)
for cb in self.cbs:
for cb2 in self.cbs:
if cb2.__class__.__name__ in cb.dependsOn:
cb2._dependents.add(cb)
class SuspendContext:
def __init__(self, cbs:Callbacks, cbsNames:List[str], cbsClasses:List[str]):
self.cbs = cbs; self.cbsNames = cbsNames; self.cbsClasses = cbsClasses
self.cbs.suspendStack = getattr(self.cbs, "suspendStack", [])
def __enter__(self):
cbsClasses = set(self.cbsClasses); cbsNames = set(self.cbsNames)
self._resolveDependencies()
def explore(cb:Callback):
for dept in cb._dependents:
cbsClasses.add(dept.__class__.__name__); explore(dept)
[explore(cb) for cb in self.cbs if cb.__class__.__name__ in cbsClasses or cb.name in cbsNames]
stackFrame = {cb:cb.suspended for cb in self.cbs if cb.__class__.__name__ in cbsClasses or cb.name in cbsNames}
for cb in stackFrame: cb.suspend(); cb.suspended = True
self.suspendStack.append(stackFrame)
def __exit__(self, *ignored):
for cb, oldValue in self.suspendStack.pop().items():
cb.suspended = oldValue; cb.restore()
def __getattr__(self, attr): return getattr(self.cbs, attr)
@k1lib.patch(Callbacks)
def suspend(self, *cbNames:List[str]) -> ContextManager:
"""Creates suspension context for specified Callbacks. Matches callbacks with
their name. Works like this::
cbs = k1lib.Callbacks().add(CbA()).add(CbB()).add(CbC())
with cbs.suspend("CbA", "CbC"):
pass # inside here, only CbB will be active, and its checkpoints executed
# CbA, CbB and CbC are all active
.. seealso:: :meth:`suspendClasses`"""
return SuspendContext(self, cbNames, [])
@k1lib.patch(Callbacks)
def suspendClasses(self, *classNames:List[str]) -> ContextManager:
"""Like :meth:`suspend`, but matches callbacks' class names to the given list,
instead of matching names. Meaning::
cbs.k1lib.Callbacks().add(Cbs.Loss()).add(Cbs.Loss())
# cbs now has 2 callbacks "Loss" and "Loss0"
with cbs.suspendClasses("Loss"):
pass # now both of them are suspended"""
return SuspendContext(self, [], classNames)
@k1lib.patch(Callbacks)
def suspendEval(self, more:List[str]=[], less:List[str]=[]) -> ContextManager:
"""Same as :meth:`suspendClasses`, but suspend some default classes typical
used for evaluation callbacks. Just convenience method really. Currently includes:
- HookModule, HookParam, ProgressBar
- ParamScheduler, Loss, Accuracy, Autosave
- ConfusionMatrix
:param more: include more classes to be suspended
:param less: exclude classes supposed to be suspended by default"""
classes = {"HookModule", "HookParam", "ProgressBar", "ParamScheduler", "Loss", "Accuracy", "Autosave", "ConfusionMatrix"}
classes.update(more); classes -= set(less)
return self.suspendClasses(*classes)
#export
class AppendContext:
def __init__(self, cbs:Callbacks): self.cbs = cbs
def __enter__(self): self.cbs.contexts.append([])
def __exit__(self, *ignored):
[cb.detach() for cb in self.cbs.contexts.pop()]
@k1lib.patch(Callbacks)
def _appendContext_append(self, cb):
if "contexts" not in self.__dict__: self.contexts = [[]]
self.contexts[-1].append(cb)
@k1lib.patch(Callbacks)
def context(self) -> ContextManager:
"""Add context.
Works like this::
cbs = k1lib.Callbacks().add(CbA())
# CbA is available
with cbs.context():
cbs.add(CbB())
# CbA and CbB available
cbs.add(CbC())
# all 3 are available
# only CbA is available
"""
return AppendContext(self)
#export
@k1lib.patch(Callbacks)
def _checkpointGraph_call(self, checkpoints:List[str]):
if not hasattr(self, "_checkpointGraphDict"):
self._checkpointGraphDict = k1lib.Object().withAutoDeclare(lambda: k1lib.Object().withAutoDeclare(lambda: 0))
self._lastCheckpoint = "<root>"
for cp in checkpoints:
self._checkpointGraphDict[self._lastCheckpoint][cp] += 1
self._lastCheckpoint = cp
@k1lib.patch(Callbacks)
def checkpointGraph(self, highlightCb:Union[str, Callback]=None):
"""Graphs what checkpoints follows what checkpoints. Has to run at least once
first. Requires graphviz package though. Example::
cbs = Callbacks()
cbs("a", "b", "c", "d", "b")
cbs.checkpointGraph() # returns graph object. Will display image if using notebooks
.. image:: ../images/checkpointGraph.png
:param highlightCb: if available, will highlight the checkpoints the callback
uses. Can be name/class-name/class/self of callback."""
g = k1lib.digraph(); s = set()
for cp1, cp1o in self._checkpointGraphDict.state.items():
for cp2, v in cp1o.state.items():
g.edge(cp1, cp2, label=f" {v} "); s.add(cp2)
if highlightCb != None:
_cb = None
if isinstance(highlightCb, Callback): _cb = highlightCb
elif isinstance(highlightCb, type) and issubclass(highlightCb, Callback): # find cb that has the same class
for cbo in self.cbs:
if isinstance(cbo, highlightCb): _cb = cbo; break
if _cb is None: raise AttributeError(f"Can't find any Callback inside this Callbacks which is of type `{cb.__name__}`")
elif isinstance(highlightCb, str):
for cbName, cbo in self.cbsDict.items():
if cbName == highlightCb: _cb = cbo; break
if type(cbo).name == highlightCb: _cb = cbo; break
if _cb is None: raise AttributeError(f"Can't find any Callback inside this Callbacks with name or class `{cb}`")
else: raise AttributeError(f"Don't understand {cb}")
print(f"Highlighting callback `{_cb.name}`, of type `{type(_cb)}`")
for cp in s:
if hasattr(_cb, cp): g.node(cp, color="red")
return g
cbs = Callbacks()
cbs.add(Callback().withCheckpoint("a", lambda self: 0), "cb1")
cbs("a", "b", "c", "d", "b")
cbs.checkpointGraph("cb1")
# !../../export.py callbacks/callbacks
| k1lib/callbacks/callbacks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Data Collection
# Collecting data about the **`Copa America 2021's`** participants from **`FBref`**
# ## Import libraries
import pandas as pd
# ## Prepare data
countries = {
'Argentina': 'f9fddd6e',
'Bolivia': '1bd2760c',
'Brazil': '304635c3',
'Chile': '7fd9c2a2',
'Colombia': 'ab73cfe5',
'Ecuador': '123acaf8',
'Paraguay': 'd2043442',
'Peru': 'f711c854',
'Uruguay': '870e020f',
'Venezuela': 'df384984'
}
tables_order = {
0: 'standard',
3: 'advance_goalkeeping',
4: 'shooting',
5: 'passing',
6: 'pass_types',
7: 'goal_and_shot_creation',
8: 'defensive_actions',
9: 'possession',
11: 'misc'
}
# ## Read and process tables
def get_column_names(table):
new_cols = []
for c in table.columns.values:
if 'Unnamed' in c[0]:
new_cols.append(c[1].lower())
else:
c_new_name = c[0].replace(' ','').lower()
c_new_name = f"{c_new_name}_{c[1]}"
new_cols.append(c_new_name.lower())
return new_cols
for country, code in countries.items():
# read tables
print(f"Reading tables of {country}...")
tables = pd.read_html(f"https://fbref.com/en/squads/{code}/2021/{country.lower()}-Stats")
# process tables
print(f"Processing tables of {country}...")
for idx, table in enumerate(tables):
if idx in tables_order.keys():
new_cols = get_column_names(table)
table.columns = new_cols
print(f"Saving table {tables_order[idx]}...")
table.to_csv(f"data/{country.lower()}_{tables_order[idx]}.csv", index=False)
| data_collection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/agemagician/CodeTrans/blob/main/prediction/transfer%20learning%20fine-tuning/function%20documentation%20generation/ruby/small_model.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="c9eStCoLX0pZ"
# **<h3>Predict the documentation for ruby code using codeTrans transfer learning finetuning model</h3>**
# <h4>You can make free prediction online through this
# <a href="https://huggingface.co/SEBIS/code_trans_t5_small_code_documentation_generation_ruby_transfer_learning_finetune">Link</a></h4> (When using the prediction online, you need to parse and tokenize the code first.)
# + [markdown] id="6YPrvwDIHdBe"
# **1. Load necessry libraries including huggingface transformers**
# + colab={"base_uri": "https://localhost:8080/"} id="6FAVWAN1UOJ4" outputId="16e8533c-9b90-4670-a3e0-a3adb892df5b"
# !pip install -q transformers sentencepiece
# + id="53TAO7mmUOyI"
from transformers import AutoTokenizer, AutoModelWithLMHead, SummarizationPipeline
# + [markdown] id="xq9v-guFWXHy"
# **2. Load the token classification pipeline and load it into the GPU if avilabile**
# + colab={"base_uri": "https://localhost:8080/", "height": 316, "referenced_widgets": ["d5a66b0bffce496688cce3414a278c9b", "a46e494d44b34a759e8c28d637a0610c", "51d866add9ba4df19c8f48590675ec2a", "<KEY>", "008d6a82fa264c318c6d1c04e0fec6c2", "4cedc51c1ba74999a50cc476ae2783e8", "569c64d3e36043249a65f4297a2231cc", "b7039dc0e0d846b0a0f1743d8bf32ed6", "<KEY>", "fd40095843e548178e2fe3eed1c3f479", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "ca410b947d1d47f59b09c62d7a4229ba", "94ade8e6876c4addb6f4de750f225d64", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "899726e4a7fa47cc94148403bca1d5c5", "<KEY>", "0f87479fe5cf4829995fbc0e9f3a247e", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "0baa797833354b878ab3dfb5172e9f71", "1949658caf64492585aa0e41f8e0945a", "99165976006b45db9961364e5ab21942", "c5323ecdad494d1794ef6e3f812658e7", "7c82631295b840d29ed223f71649852c", "91ec07f673fd477c988478e8512da923", "ffe5db98602d45258088a511e97e1a3d", "b2971827ba26441894bc11026acde578"]} id="5ybX8hZ3UcK2" outputId="1491735d-be01-44e9-ea27-b78619c9b8fc"
pipeline = SummarizationPipeline(
model=AutoModelWithLMHead.from_pretrained("SEBIS/code_trans_t5_small_code_documentation_generation_ruby_transfer_learning_finetune"),
tokenizer=AutoTokenizer.from_pretrained("SEBIS/code_trans_t5_small_code_documentation_generation_ruby_transfer_learning_finetune", skip_special_tokens=True),
device=0
)
# + [markdown] id="hkynwKIcEvHh"
# **3 Give the code for summarization, parse and tokenize it**
# + id="nld-UUmII-2e"
code = "def add(severity, progname, &block)\n return true if io.nil? || severity < level\n message = format_message(severity, progname, yield)\n MUTEX.synchronize { io.write(message) }\n true\n end" #@param {type:"raw"}
# + id="cJLeTZ0JtsB5" colab={"base_uri": "https://localhost:8080/"} outputId="dd70d67f-b575-403c-a139-494252c3357f"
# !pip install tree_sitter
# !git clone https://github.com/tree-sitter/tree-sitter-ruby
# + id="hqACvTcjtwYK"
from tree_sitter import Language, Parser
Language.build_library(
'build/my-languages.so',
['tree-sitter-ruby']
)
RUBY_LANGUAGE = Language('build/my-languages.so', 'ruby')
parser = Parser()
parser.set_language(RUBY_LANGUAGE)
# + id="LLCv2Yb8t_PP"
def get_string_from_code(node, lines):
line_start = node.start_point[0]
line_end = node.end_point[0]
char_start = node.start_point[1]
char_end = node.end_point[1]
if line_start != line_end:
code_list.append(' '.join([lines[line_start][char_start:]] + lines[line_start+1:line_end] + [lines[line_end][:char_end]]))
else:
code_list.append(lines[line_start][char_start:char_end])
def my_traverse(node, code_list):
lines = code.split('\n')
if node.child_count == 0:
get_string_from_code(node, lines)
elif node.type == 'string':
get_string_from_code(node, lines)
else:
for n in node.children:
my_traverse(n, code_list)
return ' '.join(code_list)
# + id="BhF9MWu1uCIS" colab={"base_uri": "https://localhost:8080/"} outputId="b487bf5c-8223-41d2-fde6-6eba676282c7"
tree = parser.parse(bytes(code, "utf8"))
code_list=[]
tokenized_code = my_traverse(tree.root_node, code_list)
print("Output after tokenization: " + tokenized_code)
# + [markdown] id="sVBz9jHNW1PI"
# **4. Make Prediction**
# + colab={"base_uri": "https://localhost:8080/"} id="KAItQ9U9UwqW" outputId="d30bf764-4411-4d27-9398-df09b92bf2fb"
pipeline([tokenized_code])
| prediction/transfer learning fine-tuning/function documentation generation/ruby/small_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Differential correlation analysis
import scanpy as sc
import seaborn as sns
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import scipy.stats as stats
from pybedtools import BedTool
import pickle as pkl
# %matplotlib inline
import itertools
import sys
sys.path.append('/home/ssm-user/Github/scrna-parameter-estimation/dist/memento-0.0.6-py3.8.egg')
sys.path.append('/home/ssm-user/Github/misc-seq/miscseq')
import encode
import memento
pd.set_option('display.max_rows', None)
data_path = '/data_volume/memento/hbec/'
fig_path = '/data/home/Github/scrna-parameter-estimation/figures/fig6/'
import matplotlib
matplotlib.rcParams['pdf.fonttype'] = 42
matplotlib.rcParams['ps.fonttype'] = 42
import matplotlib.pylab as pylab
params = {'legend.fontsize': 'medium',
'axes.labelsize': 'medium',
'axes.titlesize':'medium',
'figure.titlesize':'medium',
'xtick.labelsize':'small',
'ytick.labelsize':'small'}
pylab.rcParams.update(params)
# ### Get canonical TFs
tf_df = pd.read_csv('../baseline/human_tf.txt', sep='\t')
tf_list = tf_df['Symbol'].tolist()
tf_list += ['CIITA', 'NLRC5']
# ### Read the processed RNA data
#
# Focus on the club and bc/club cells and type I interferons for now.
#
# Encode the timestamps to integers.
# +
# adata_processed = sc.read(data_path + 'HBEC_type_I_processed_deep.h5ad')
# -
adata = sc.read(data_path + 'HBEC_type_I_filtered_counts_deep.h5ad')
adata = adata[:, ~adata.var.index.str.startswith('MT-')].copy()
# adata.obs['cell_type'] = adata.obs['cell_type'].apply(lambda x: x if x != 'basal/club' else 'bc')
# adata.obs['cell_type'] = adata.obs['cell_type'].apply(lambda x: x if x != 'ionocyte/tuft' else 'ion-tuft')
# + active=""
# sc.pl.umap(adata_processed, color=['cell_type', 'time', 'stim'])
# -
converter = {'basal/club':'BC', 'basal':'B', 'ciliated':'C', 'goblet':'G', 'ionocyte/tuft':'IT', 'neuroendo':'N'}
adata.obs['ct'] = adata.obs['cell_type'].apply(lambda x: converter[x])
# ### Setup memento
def assign_q(batch):
if batch == 0:
return 0.387*0.25
elif batch == 1:
return 0.392*0.25
elif batch == 2:
return 0.436*0.25
else:
return 0.417*0.25
adata.obs['q'] = adata.obs['batch'].apply(assign_q)
memento.setup_memento(adata, q_column='q', trim_percent=0.1)
isg_classes = pd.read_csv('../isg_classes.csv')
t1_isg = isg_classes.query('overall_type =="type1"').gene.tolist()
t2_isg = isg_classes.query('overall_type =="type2"').gene.tolist()
shared_isg = isg_classes.query('overall_type == "shared"').gene.tolist()
all_isg = isg_classes[isg_classes.sum(axis=1) > 0].gene.tolist()
tf_isg = list(set(all_isg) & set(tf_list))
# ### Compare Type 1 vs 2 IFN's TFs
# +
ct = ['C']
adata_stim = adata.copy()[
adata.obs.ct.isin(ct) & \
adata.obs.stim.isin(['beta', 'gamma'])].copy()
adata_stim.obs['group'] = 1
# adata_stim.obs['time_step'] = adata_stim.obs['time']#.astype(int).apply(lambda x: time_converter[x])
# adata_stim.obs['time'] = 'tp_' + adata_stim.obs['time'].astype(str)#.apply(lambda x: time_converter[x])
memento.create_groups(adata_stim, label_columns=['donor', 'stim'])
memento.compute_1d_moments(adata_stim, min_perc_group=.9)
# moment_1d_df = memento.get_1d_moments(adata_stim, groupby='is_stim')
available_tfs = list(set(tf_isg) & set(adata_stim.var.index))
available_isgs= list(set(all_isg) & set(adata_stim.var.index))
memento.compute_2d_moments(adata_stim, list(itertools.product(available_tfs, available_isgs)))
corr_df = memento.get_2d_moments(adata_stim, groupby='stim')
m_df, v_df = memento.get_1d_moments(adata_stim, groupby='stim')
# memento.ht_2d_moments(
# adata_stim,
# formula_like='1 + donor',
# treatment_col='Inter',
# num_boot=10000,
# verbose=1,
# num_cpus=94,
# resampling='bootstrap',
# approx=False)
# moment_df = memento.get_2d_moments(adata_stim, groupby='group')
# rho = memento.get_2d_ht_result(adata_stim)
# -
m_df.query('gene in @tf_isg')
corr_df['diff'] = corr_df['stim_gamma'] - corr_df['stim_beta']
corr_df.groupby('gene_1')['diff'].mean().sort_values()
corr_df.sort_values('diff', ascending=False).head(30)
# ### Compare STAT1, STAT2, and STAT3
# +
# First, get correlations without adjustment
stim = 'gamma'
ct = ['C']
adata_stim = adata.copy()[
adata.obs.ct.isin(ct) & \
adata.obs.stim.isin([stim])].copy()
adata_stim.obs['group'] = 1
# adata_stim.obs['time_step'] = adata_stim.obs['time']#.astype(int).apply(lambda x: time_converter[x])
# adata_stim.obs['time'] = 'tp_' + adata_stim.obs['time'].astype(str)#.apply(lambda x: time_converter[x])
memento.create_groups(adata_stim, label_columns=['donor', 'group'])
memento.compute_1d_moments(adata_stim, min_perc_group=.9)
# moment_1d_df = memento.get_1d_moments(adata_stim, groupby='is_stim')
available_tfs = list(set(tf_list) & set(adata_stim.var.index))
available_isgs= list(set(all_isg) & set(adata_stim.var.index))
memento.compute_2d_moments(adata_stim, list(itertools.product(['STAT1', 'STAT2', 'IRF9', 'STAT3'], available_isgs)))
memento.ht_2d_moments(
adata_stim,
formula_like='1 + donor',
treatment_col='Inter',
num_boot=10000,
verbose=1,
num_cpus=94,
resampling='bootstrap',
approx=False)
moment_df = memento.get_2d_moments(adata_stim, groupby='group')
rho = memento.get_2d_ht_result(adata_stim)
# -
list(itertools.product(['STAT1', 'STAT2', 'IRF9', 'STAT3'], available_isgs))
# make data into wideform
wideform = pd.pivot(rho, index='gene_2', columns='gene_1', values=['corr_coef', 'corr_se'])
plt.scatter(wideform.loc[:, 'corr_coef'].loc[:,'STAT2'], wideform.loc[:, 'corr_coef'].loc[:,'STAT1'])
plt.plot([0,0.7], [0., 0.7])
wideform.head(50)
# +
# First, get correlations without adjustment
stim = 'gamma'
ct = ['C']
adata_stim = adata.copy()[
adata.obs.ct.isin(ct) & \
adata.obs.stim.isin([stim])].copy()
adata_stim.obs['group'] = 1
adata_stim.obs['med_expr'] = adata_stim[:, 'STAT1'].X.todense().A1
memento.create_groups(adata_stim, label_columns=['donor', 'group', 'med_expr'])
memento.compute_1d_moments(adata_stim, min_perc_group=.3)
# moment_1d_df = memento.get_1d_moments(adata_stim, groupby='is_stim')
available_genes = adata_stim.var.index.tolist()
candidates = candidates.query('gene_1 in @available_genes & gene_2 in @available_genes')
print(candidates.shape)
memento.compute_2d_moments(adata_stim, list(zip(candidates['gene_1'], candidates['gene_2'])))
memento.ht_2d_moments(
adata_stim,
formula_like='1 + donor + med_expr',
treatment_col='Inter',
num_boot=10000,
verbose=1,
num_cpus=94,
resampling='bootstrap',
approx=False)
rho_adj = memento.get_2d_ht_result(adata_stim)
# -
merged_rho = rho.merge(rho_adj, on=['gene_1', 'gene_2'], suffixes=('', '_adj'))
merged_rho['diff'] = merged_rho['corr_coef'] - merged_rho['corr_coef_adj']
merged_rho['diff_se'] = np.sqrt(merged_rho['corr_se']**2 + merged_rho['corr_se_adj']**2).values
merged_rho.query('gene_1 == "IRF1"').sort_values('diff', ascending=False)
merged_rho.head(2)
rho_adj
delta
for tf in (set(de_genes['gamma'])&set(tf_list)):
if tf in ['STAT1', 'ATF3'] or tf in adj_results: continue
delta, se = adjust(tf)
pv = stats.norm.sf(delta, loc=0, scale=se)
adj_results[tf] = (delta, se, pv)
def adjust(mediator):
adata_stim = adata.copy()[
adata.obs.cell_type.isin(ct) & \
adata.obs.stim.isin([stim])].copy()
adata_stim.obs['time_step'] = adata_stim.obs['time']#.astype(int).apply(lambda x: time_converter[x])
adata_stim.obs['time'] = 'tp_' + adata_stim.obs['time'].astype(str)#.apply(lambda x: time_converter[x])
adata_stim.obs['med_expr'] = adata_stim[:, mediator].X.todense().A1
memento.create_groups(adata_stim, label_columns=['donor', 'med_expr'])
memento.compute_1d_moments(adata_stim, min_perc_group=.5)
# moment_1d_df = memento.get_1d_moments(adata_stim, groupby='is_stim')
memento.compute_2d_moments(adata_stim, list(itertools.product(['STAT1'], c2_genes)))
# moment_df = memento.get_2d_moments(adata_stim, groupby='is_stim')
memento.ht_2d_moments(
adata_stim,
formula_like='1 + donor + med_expr',
treatment_col='Inter',
num_boot=10000,
verbose=0,
num_cpus=94,
resampling='permutation',
approx=False)
rho_adj = memento.get_2d_ht_result(adata_stim)
return (rho['corr_coef'] - rho_adj['corr_coef']).values, np.sqrt(rho['corr_se']**2 + rho_adj['corr_se']**2).values
delta, se = adjust('CIITA')
print(stats.norm.sf(delta, loc=0, scale=se))
for tf in (set(de_genes['gamma'])&set(tf_list)):
if tf in ['STAT1', 'ATF3'] or tf in adj_results: continue
delta, se = adjust(tf)
pv = stats.norm.sf(delta, loc=0, scale=se)
adj_results[tf] = (delta, se, pv)
result_df = []
for tf, val in adj_results.items():
a = pd.DataFrame()
a['gene'] = c2_genes
a['se'] = val[1]
a['delta_rho'] = val[0]
a['pv'] = val[2]
a['tf'] = tf
result_df.append(a)
result_df = pd.concat(result_df)
sns.barplot(y='tf', x='delta_rho', data=result_df.sort_values('delta_rho', ascending=False), errwidth=0)
sns.barplot(y='tf', x='delta_rho', data=result_df.groupby('tf')['delta_rho'].mean().sort_values(ascending=False).reset_index(name='delta_rho'))
plt.xlabel(r'$\Delta\rho_{\mathrm{ADJ}} - \Delta\rho$')
pv = result_df.groupby('tf')['pv'].apply(lambda x: stats.combine_pvalues(x)[1]).sort_values()
fdr = memento.util._fdrcorrect(pv)
fdr
delta, se = adjust('JUNB')
print(stats.norm.sf(delta, loc=0, scale=se))
delta, se = adjust('CIITA')
print(stats.norm.sf(delta, loc=0, scale=se))
# ### Run differential correlation between STAT1 and each stim's DEGs
ct = ['ciliated']
all_stims = ['gamma', 'lambda', 'alpha', 'beta']
# +
for stim in all_stims:
print('starting ', stim, '....')
adata_stim = adata.copy()[
adata.obs.cell_type.isin(ct) & \
adata.obs.stim.isin(['control', stim])].copy()
# time_converter={0:0, int('6'):1}
adata_stim.obs['time_step'] = adata_stim.obs['time']#.astype(int).apply(lambda x: time_converter[x])
adata_stim.obs['time'] = 'tp_' + adata_stim.obs['time'].astype(str)
memento.create_groups(adata_stim, label_columns=['time', 'donor'])
memento.compute_1d_moments(adata_stim, min_perc_group=.3)
moment_1d_df = memento.get_1d_moments(adata_stim, groupby='time')
# available_tfs = list(set(tf_list) & set(moment_1d_df[0].gene))
available_targets = list(set(de_genes[stim if stim not in ['alpha', 'beta'] else 'alpha/beta']) & set(moment_1d_df[0].gene))
memento.compute_2d_moments(adata_stim, list(itertools.product(['STAT1'], available_targets)))
# moment_df = memento.get_2d_moments(adata_stim, groupby='time')
# moment_df['max_corr'] = moment_df.iloc[:, 2:].max(axis=1).values
# candidates = moment_df.query('max_corr > 0.3 & gene_1 != gene_2')
# # top_tfs = candidates.groupby('gene_1').size().sort_values(ascending=False)
# # top_tfs = top_tfs[top_tfs > 15].index.tolist()
# # candidates = candidates.query('gene_1 in @top_tfs')
# candidates = list(zip(candidates['gene_1'], candidates['gene_2']))
# memento.compute_2d_moments(adata_stim, candidates)
memento.ht_2d_moments(
adata_stim,
formula_like='1 + time + donor',
treatment_col='time',
num_boot=10000,
verbose=1,
num_cpus=94,
resampling='permutation',
approx=False)
adata_stim.write(data_path + 'stat1_coex_test/{}.h5ad'.format(stim))
# +
for stim in all_stims:
print('starting ', stim, '....')
adata_stim = adata.copy()[
adata.obs.cell_type.isin(ct) & \
adata.obs.stim.isin(['control', stim])].copy()
time_converter={'0':0, '3':1, '6':2, '9':3, '24':4, '48':5}
adata_stim.obs['time_step'] = adata_stim.obs['time']#.astype(int).apply(lambda x: time_converter[x])
adata_stim.obs['time'] = 'tp_' + adata_stim.obs['time'].astype(str)#.apply(lambda x: time_converter[x])
adata_stim.obs['is_stim'] = (adata_stim.obs.stim==stim).astype(int)
memento.create_groups(adata_stim, label_columns=['is_stim','time', 'donor'])
memento.compute_1d_moments(adata_stim, min_perc_group=.3)
moment_1d_df = memento.get_1d_moments(adata_stim, groupby='time')
available_targets = list(set(de_genes[stim]) & set(moment_1d_df[0].gene))
memento.compute_2d_moments(adata_stim, list(itertools.product(['STAT1'], available_targets)))
moment_df = memento.get_2d_moments(adata_stim, groupby='time')
# moment_df['max_corr'] = moment_df.iloc[:, 2:].max(axis=1).values
# candidates = moment_df.query('max_corr > 0.3 & gene_1 != gene_2')
# candidates = list(zip(candidates['gene_1'], candidates['gene_2']))
# memento.compute_2d_moments(adata_stim, candidates)
memento.ht_2d_moments(
adata_stim,
formula_like='1 + is_stim + donor',
treatment_col='is_stim',
num_boot=10000,
verbose=1,
num_cpus=94,
resampling='permutation',
approx=False)
adata_stim.write(data_path + 'stat1_coex_test/{}_linear.h5ad'.format(stim))
# -
# ### Read DC results
def classify(x):
if x in shared_de_genes:
return 'shared'
elif x in shared_de_genes_lenient:
return 'partially_shared'
for stim in stims:
if x in uniq_de_genes[stim]:
return 'uniq_' + stim
return 'none-DE'
# +
deg_dc_results = {}
deg_2d_moments = {}
deg_m_moments = {}
deg_v_moments = {}
combined_results = {}
for stim in all_stims:
adata_stim = sc.read(data_path + 'stat1_coex_test/{}_linear.h5ad'.format(stim))
deg_dc_results[stim] = memento.get_2d_ht_result(adata_stim)
deg_2d_moments[stim] = memento.get_2d_moments(adata_stim, groupby='time')[['gene_1','gene_2'] + ['time_'+str(i) for i in [0, 1, 2, 3, 4, 5]]]
# deg_m_moments[stim],deg_v_moments[stim] = memento.get_1d_moments(adata_stim, groupby='time')
# deg_m_moments[stim] = deg_m_moments[stim][['gene'] + ['time_'+str(i) for i in [0, 3, 6, 9, 24, 48]]]
# deg_v_moments[stim] = deg_v_moments[stim][['gene'] + ['time_'+str(i) for i in [0, 3, 6, 9, 24, 48]]]
deg_dc_results[stim]['corr_fdr'] = memento.util._fdrcorrect(deg_dc_results[stim]['corr_pval'])
combined_results[stim] = deg_2d_moments[stim].merge(deg_dc_results[stim], on=['gene_1', 'gene_2'])
combined_results[stim]['deg_type'] = combined_results[stim]['gene_2'].apply(classify)
combined_results[stim]['sig'] = (combined_results[stim]['corr_fdr'] < 0.05) & (combined_results[stim]['corr_coef'] > 0.0)
# -
# ### Differential correlation and sharedness
a = deg_dc_results['lambda']
a['is_uniq'] = a['gene_2'].isin(uniq_de_genes['lambda'])
combined_results['lambda'].sort_values('corr_coef', ascending=False)
deg_dc_results['lambda']#.query('deg_type=="shared"')
for stim in all_stims:
contingency = pd.crosstab(
deg_dc_results[stim]['deg_type']=='shared',
deg_dc_results[stim]['sig'])
print(stim)
print(contingency)
print(stim, stats.fisher_exact(contingency))
ht_2d_df = memento.get_2d_ht_result(adata_stim)
ht_2d_df['corr_fdr'] = memento.util._fdrcorrect(ht_2d_df['corr_pval'])
sig_df = ht_2d_df.query('corr_fdr < 0.1')
tp_order = ['time_tp_' + str(i) for i in [0, 3, 6, 9, 24, 48]]
moment_1d_df[0].query('gene == "CIITA"')[tp_order]
dra=moment_df.merge(sig_df[['gene_1', 'gene_2']], on=['gene_1', 'gene_2'], how='inner')\
.query('gene_2.str.startswith("HLA-DRA")', engine='python')[['gene_1', 'gene_2'] + tp_order]
plt.plot(dra[tp_order].values.T, lw=1);
| analysis/ifn_hbec/version3/coexpression/pathway.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Machine Learning Engineer Nanodegree
# ## Model Evaluation & Validation
# ## Project: Predicting Boston Housing Prices
#
# Welcome to the first project of the Machine Learning Engineer Nanodegree! In this notebook, some template code has already been provided for you, and you will need to implement additional functionality to successfully complete this project. You will not need to modify the included code beyond what is requested. Sections that begin with **'Implementation'** in the header indicate that the following block of code will require additional functionality which you must provide. Instructions will be provided for each section and the specifics of the implementation are marked in the code block with a 'TODO' statement. Please be sure to read the instructions carefully!
#
# In addition to implementing code, there will be questions that you must answer which relate to the project and your implementation. Each section where you will answer a question is preceded by a **'Question X'** header. Carefully read each question and provide thorough answers in the following text boxes that begin with **'Answer:'**. Your project submission will be evaluated based on your answers to each of the questions and the implementation you provide.
#
# >**Note:** Code and Markdown cells can be executed using the **Shift + Enter** keyboard shortcut. In addition, Markdown cells can be edited by typically double-clicking the cell to enter edit mode.
# ## Getting Started
# In this project, you will evaluate the performance and predictive power of a model that has been trained and tested on data collected from homes in suburbs of Boston, Massachusetts. A model trained on this data that is seen as a *good fit* could then be used to make certain predictions about a home — in particular, its monetary value. This model would prove to be invaluable for someone like a real estate agent who could make use of such information on a daily basis.
#
# The dataset for this project originates from the [UCI Machine Learning Repository](https://archive.ics.uci.edu/ml/datasets/Housing). The Boston housing data was collected in 1978 and each of the 506 entries represent aggregated data about 14 features for homes from various suburbs in Boston, Massachusetts. For the purposes of this project, the following preprocessing steps have been made to the dataset:
# - 16 data points have an `'MEDV'` value of 50.0. These data points likely contain **missing or censored values** and have been removed.
# - 1 data point has an `'RM'` value of 8.78. This data point can be considered an **outlier** and has been removed.
# - The features `'RM'`, `'LSTAT'`, `'PTRATIO'`, and `'MEDV'` are essential. The remaining **non-relevant features** have been excluded.
# - The feature `'MEDV'` has been **multiplicatively scaled** to account for 35 years of market inflation.
#
# Run the code cell below to load the Boston housing dataset, along with a few of the necessary Python libraries required for this project. You will know the dataset loaded successfully if the size of the dataset is reported.
# +
# Import libraries necessary for this project
import numpy as np
import pandas as pd
from sklearn.cross_validation import ShuffleSplit
# Import supplementary visualizations code visuals.py
import visuals as vs
# Pretty display for notebooks
# %matplotlib inline
# Load the Boston housing dataset
data = pd.read_csv('housing.csv')
prices = data['MEDV']
features = data.drop('MEDV', axis = 1)
# Success
print "Boston housing dataset has {} data points with {} variables each.".format(*data.shape)
# -
# ## Data Exploration
# In this first section of this project, you will make a cursory investigation about the Boston housing data and provide your observations. Familiarizing yourself with the data through an explorative process is a fundamental practice to help you better understand and justify your results.
#
# Since the main goal of this project is to construct a working model which has the capability of predicting the value of houses, we will need to separate the dataset into **features** and the **target variable**. The **features**, `'RM'`, `'LSTAT'`, and `'PTRATIO'`, give us quantitative information about each data point. The **target variable**, `'MEDV'`, will be the variable we seek to predict. These are stored in `features` and `prices`, respectively.
# ### Implementation: Calculate Statistics
# For your very first coding implementation, you will calculate descriptive statistics about the Boston housing prices. Since `numpy` has already been imported for you, use this library to perform the necessary calculations. These statistics will be extremely important later on to analyze various prediction results from the constructed model.
#
# In the code cell below, you will need to implement the following:
# - Calculate the minimum, maximum, mean, median, and standard deviation of `'MEDV'`, which is stored in `prices`.
# - Store each calculation in their respective variable.
# +
# TODO: Minimum price of the data
minimum_price = np.min(prices)
# TODO: Maximum price of the data
maximum_price = np.max(prices)
# TODO: Mean price of the data
mean_price = np.mean(prices)
# TODO: Median price of the data
median_price = np.median(prices)
# TODO: Standard deviation of prices of the data
std_price = np.std(prices,ddof = 1)
# Mean of the three featurs, and standard deviations
avg_RM = np.mean(features['RM'])
std_RM = np.std(features['RM'])
avg_LSTAT = np.mean(features['LSTAT'])
std_LSTAT = np.std(features['LSTAT'])
avg_PTRATIO = np.mean(features['PTRATIO'])
std_PTRATIO = np.std(features['PTRATIO'])
# Show the calculated statistics
print "Statistics for Boston housing dataset:\n"
print "Minimum price: ${:,.2f}".format(minimum_price)
print "Maximum price: ${:,.2f}".format(maximum_price)
print "Mean price: ${:,.2f}".format(mean_price)
print "Median price ${:,.2f}".format(median_price)
print "Standard deviation of prices: ${:,.2f}".format(std_price)
print "Mean RM: {:,.2f}".format(avg_RM)
print "RM Standard deviation: {:,.2f}".format(std_RM)
print "Mean LSTAT: {:,.2f}".format(avg_LSTAT)
print "LSTAT Standard deviation: {:,.2f}".format(std_LSTAT)
print "Mean PTRATIO: {:,.2f}".format(avg_PTRATIO)
print "PTRATIO Standard Deviation: {:,.2f}".format(std_PTRATIO)
# -
# ### Question 1 - Feature Observation
# As a reminder, we are using three features from the Boston housing dataset: `'RM'`, `'LSTAT'`, and `'PTRATIO'`. For each data point (neighborhood):
# - `'RM'` is the average number of rooms among homes in the neighborhood.
# - `'LSTAT'` is the percentage of homeowners in the neighborhood considered "lower class" (working poor).
# - `'PTRATIO'` is the ratio of students to teachers in primary and secondary schools in the neighborhood.
#
# _Using your intuition, for each of the three features above, do you think that an increase in the value of that feature would lead to an **increase** in the value of `'MEDV'` or a **decrease** in the value of `'MEDV'`? Justify your answer for each._
# **Hint:** Would you expect a home that has an `'RM'` value of 6 be worth more or less than a home that has an `'RM'` value of 7?
# **Answer: ** I believe an increase in the RM feature would lead to an increase of the value of MEDV. This is based on the following assumptions:
#
# 1) As the number of rooms in a house increases, the size in square feet of the house is also increasing. In other words meaning that a larger number of rooms corressponds to a larger measured living area.
# 2) A larger house as measured in square feet is directly related to an increase in cost.
#
# I believe an increase in the LSTAT feature would lead to a decrease in the value of MEDV. This is based on the following assumptions:
#
# 1) Neighborhoods catering towards the working poor will price houses more affordably.
# 2) Homeowners in these neighborhoods are buying homes financially sustainable relative to their working wage. Where financially sustainable could mean the homeowners are able to pay off their home with a standard mortgage payments not taking out more than 30-40% of their income.
#
# I believe an increase in the PTRATIO feature would lead to a decrease in MEDV. This is based on the following assumptions:
#
# 1) A lower PTRATIO of schools is correlated with better student success measurements.
# 2) Homeowners think of better schools as an amenity, and this is corresspondingly taken into the value of a home by real estate agents.
#
# ----
#
# ## Developing a Model
# In this second section of the project, you will develop the tools and techniques necessary for a model to make a prediction. Being able to make accurate evaluations of each model's performance through the use of these tools and techniques helps to greatly reinforce the confidence in your predictions.
# ### Implementation: Define a Performance Metric
# It is difficult to measure the quality of a given model without quantifying its performance over training and testing. This is typically done using some type of performance metric, whether it is through calculating some type of error, the goodness of fit, or some other useful measurement. For this project, you will be calculating the [*coefficient of determination*](http://stattrek.com/statistics/dictionary.aspx?definition=coefficient_of_determination), R<sup>2</sup>, to quantify your model's performance. The coefficient of determination for a model is a useful statistic in regression analysis, as it often describes how "good" that model is at making predictions.
#
# The values for R<sup>2</sup> range from 0 to 1, which captures the percentage of squared correlation between the predicted and actual values of the **target variable**. A model with an R<sup>2</sup> of 0 is no better than a model that always predicts the *mean* of the target variable, whereas a model with an R<sup>2</sup> of 1 perfectly predicts the target variable. Any value between 0 and 1 indicates what percentage of the target variable, using this model, can be explained by the **features**. _A model can be given a negative R<sup>2</sup> as well, which indicates that the model is **arbitrarily worse** than one that always predicts the mean of the target variable._
#
# For the `performance_metric` function in the code cell below, you will need to implement the following:
# - Use `r2_score` from `sklearn.metrics` to perform a performance calculation between `y_true` and `y_predict`.
# - Assign the performance score to the `score` variable.
# TODO: Import 'r2_score'
from sklearn.metrics import r2_score
def performance_metric(y_true, y_predict):
""" Calculates and returns the performance score between
true and predicted values based on the metric chosen. """
# TODO: Calculate the performance score between 'y_true' and 'y_predict'
score = r2_score(y_true,y_predict)
# Return the score
return score
# ### Question 2 - Goodness of Fit
# Assume that a dataset contains five data points and a model made the following predictions for the target variable:
#
# | True Value | Prediction |
# | :-------------: | :--------: |
# | 3.0 | 2.5 |
# | -0.5 | 0.0 |
# | 2.0 | 2.1 |
# | 7.0 | 7.8 |
# | 4.2 | 5.3 |
# *Would you consider this model to have successfully captured the variation of the target variable? Why or why not?*
#
# Run the code cell below to use the `performance_metric` function and calculate this model's coefficient of determination.
# Calculate the performance of this model
score = performance_metric([3, -0.5, 2, 7, 4.2], [2.5, 0.0, 2.1, 7.8, 5.3])
print "Model has a coefficient of determination, R^2, of {:.3f}.".format(score)
# **Answer:** This model successfully captured the variation of the target variable. This is reflected in the high R^2 value showing that there is a 92.3% squared correlation between the actual and predicted values. This high percentage can be alternatively interpretd as the model being able to almost perfectly fit the data.
# ### Implementation: Shuffle and Split Data
# Your next implementation requires that you take the Boston housing dataset and split the data into training and testing subsets. Typically, the data is also shuffled into a random order when creating the training and testing subsets to remove any bias in the ordering of the dataset.
#
# For the code cell below, you will need to implement the following:
# - Use `train_test_split` from `sklearn.cross_validation` to shuffle and split the `features` and `prices` data into training and testing sets.
# - Split the data into 80% training and 20% testing.
# - Set the `random_state` for `train_test_split` to a value of your choice. This ensures results are consistent.
# - Assign the train and testing splits to `X_train`, `X_test`, `y_train`, and `y_test`.
# +
# TODO: Import 'train_test_split'
from sklearn.cross_validation import train_test_split
# TODO: Shuffle and split the data into training and testing subsets
X_train, X_test, y_train, y_test = train_test_split(features,prices, test_size = 0.2, random_state = 10)
# Success
print "Training and testing split was successful."
# -
# ### Question 3 - Training and Testing
# *What is the benefit to splitting a dataset into some ratio of training and testing subsets for a learning algorithm?*
# **Hint:** What could go wrong with not having a way to test your model?
# **Answer: **Splitting data into training and testing subsets allows one to test a learning algorithm on data that is different from data which it used to build a model. The performance of a learning algorithm on an independent tesing dataset can allow one to help determine if there is error due to a model underfitting/overfitting its predictions.
# ----
#
# ## Analyzing Model Performance
# In this third section of the project, you'll take a look at several models' learning and testing performances on various subsets of training data. Additionally, you'll investigate one particular algorithm with an increasing `'max_depth'` parameter on the full training set to observe how model complexity affects performance. Graphing your model's performance based on varying criteria can be beneficial in the analysis process, such as visualizing behavior that may not have been apparent from the results alone.
# ### Learning Curves
# The following code cell produces four graphs for a decision tree model with different maximum depths. Each graph visualizes the learning curves of the model for both training and testing as the size of the training set is increased. Note that the shaded region of a learning curve denotes the uncertainty of that curve (measured as the standard deviation). The model is scored on both the training and testing sets using R<sup>2</sup>, the coefficient of determination.
#
# Run the code cell below and use these graphs to answer the following question.
# Produce learning curves for varying training set sizes and maximum depths
vs.ModelLearning(features, prices)
# ### Question 4 - Learning the Data
# *Choose one of the graphs above and state the maximum depth for the model. What happens to the score of the training curve as more training points are added? What about the testing curve? Would having more training points benefit the model?*
# **Hint:** Are the learning curves converging to particular scores?
# **Answer: **I will discuss the graph labeled as max_depth = 6. As more data is added, the R^2 score declines, but begins to plateau with a score > 0.9. This decline can be due to the model being able to make predictions with little error when there are only a few points to use during the fitting of the model. The plateau at such a high R^2 shows that the model is good at describing the training data.
#
# The testing curve's score increases initially as the number of training points increase. However the score plateau's when the model is trained with 200-250 training points at a score higher than 0.7. The plateau of the testing curve shows that the models ability to generalize to test data will not get much better if further data is added.
# ### Complexity Curves
# The following code cell produces a graph for a decision tree model that has been trained and validated on the training data using different maximum depths. The graph produces two complexity curves — one for training and one for validation. Similar to the **learning curves**, the shaded regions of both the complexity curves denote the uncertainty in those curves, and the model is scored on both the training and validation sets using the `performance_metric` function.
#
# Run the code cell below and use this graph to answer the following two questions.
vs.ModelComplexity(X_train, y_train)
# ### Question 5 - Bias-Variance Tradeoff
# *When the model is trained with a maximum depth of 1, does the model suffer from high bias or from high variance? How about when the model is trained with a maximum depth of 10? What visual cues in the graph justify your conclusions?*
# **Hint:** How do you know when a model is suffering from high bias or high variance?
# **Answer: **The model suffers from high bias at a maximum depth of 1. The combination of a low R^2 score for both the training and testing curves at this depth, and the small difference in scores between these curves indicate that the model is not able to properly model trends in the data.
#
# The model suffers from high variance at a maximum depth of 10. The combination of a very high R^2 score for the training curve at this depth, and the much lower validation curve scoer indicates that the model may be overfitting to the training data.
# ### Question 6 - Best-Guess Optimal Model
# *Which maximum depth do you think results in a model that best generalizes to unseen data? What intuition lead you to this answer?*
# **Answer: ** I would think that a maximum depth of 3 would generate a model that best generalizes to unseen data. Based on the learning curves above it seems that with a maximum depth of 3, and about 300 data points we can get a score of about 0.8 which may be good enough depending on the specific application. This learning curve also shows a relatively small score difference between the training/testing curves at this area of plateau at around 300 Training Points. This can also be seen a bit more easily on the model complexity curve as the gap between training/testing curves increases above a maximum depth of 3. These two factors make it seem that a maximum depth of 3 has the best tradeoff between erros due to bias/variance.
# -----
#
# ## Evaluating Model Performance
# In this final section of the project, you will construct a model and make a prediction on the client's feature set using an optimized model from `fit_model`.
# ### Question 7 - Grid Search
# *What is the grid search technique and how it can be applied to optimize a learning algorithm?*
# **Answer: **The grid search technique is a method of trying an algorithm with multiple combinations of parameters to find which combination of parameters give the best performance. The best performance can be measured by cross-validating the algorithm with each parameter combination on training/testing data and using a metric depending on the type of model to 'score performance'. This metric could be something like R^2 above for a regression model.
# ### Question 8 - Cross-Validation
# *What is the k-fold cross-validation training technique? What benefit does this technique provide for grid search when optimizing a model?*
# **Hint:** Much like the reasoning behind having a testing set, what could go wrong with using grid search without a cross-validated set?
# **Answer: ** K-fold cross-validation is a technique that involves splitting your data into k bins. Where k-1 bins of data would be used for training, and the remaining bin for testing. This split is done multiple times with a different bin being the testing bin. An average performance metric is calculated from the test results of each individual run.
#
# Since this average performance metric was found by exposing the model to multiple test samples, this can be a better indicator of model's generalizability to unseen data. Furthermore, it limits the chance of the grid-search finding a parameter combination to be better than another combination just by chance due to being tested on more unseen data.
# ### Implementation: Fitting a Model
# Your final implementation requires that you bring everything together and train a model using the **decision tree algorithm**. To ensure that you are producing an optimized model, you will train the model using the grid search technique to optimize the `'max_depth'` parameter for the decision tree. The `'max_depth'` parameter can be thought of as how many questions the decision tree algorithm is allowed to ask about the data before making a prediction. Decision trees are part of a class of algorithms called *supervised learning algorithms*.
#
# In addition, you will find your implementation is using `ShuffleSplit()` for an alternative form of cross-validation (see the `'cv_sets'` variable). While it is not the K-Fold cross-validation technique you describe in **Question 8**, this type of cross-validation technique is just as useful!. The `ShuffleSplit()` implementation below will create 10 (`'n_splits'`) shuffled sets, and for each shuffle, 20% (`'test_size'`) of the data will be used as the *validation set*. While you're working on your implementation, think about the contrasts and similarities it has to the K-fold cross-validation technique.
#
# Please note that ShuffleSplit has different parameters in scikit-learn versions 0.17 and 0.18.
# For the `fit_model` function in the code cell below, you will need to implement the following:
# - Use [`DecisionTreeRegressor`](http://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeRegressor.html) from `sklearn.tree` to create a decision tree regressor object.
# - Assign this object to the `'regressor'` variable.
# - Create a dictionary for `'max_depth'` with the values from 1 to 10, and assign this to the `'params'` variable.
# - Use [`make_scorer`](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.make_scorer.html) from `sklearn.metrics` to create a scoring function object.
# - Pass the `performance_metric` function as a parameter to the object.
# - Assign this scoring function to the `'scoring_fnc'` variable.
# - Use [`GridSearchCV`](http://scikit-learn.org/0.17/modules/generated/sklearn.grid_search.GridSearchCV.html) from `sklearn.grid_search` to create a grid search object.
# - Pass the variables `'regressor'`, `'params'`, `'scoring_fnc'`, and `'cv_sets'` as parameters to the object.
# - Assign the `GridSearchCV` object to the `'grid'` variable.
# +
# TODO: Import 'make_scorer', 'DecisionTreeRegressor', and 'GridSearchCV'
from sklearn.tree import DecisionTreeRegressor
from sklearn.grid_search import GridSearchCV
#from sklearn.cross_validation import ShuffleSplit
def fit_model(X, y):
""" Performs grid search over the 'max_depth' parameter for a
decision tree regressor trained on the input data [X, y]. """
# Create cross-validation sets from the training data
cv_sets = ShuffleSplit(X.shape[0], test_size = 0.20, random_state = 0)
# TODO: Create a decision tree regressor object
regressor = DecisionTreeRegressor()
# TODO: Create a dictionary for the parameter 'max_depth' with a range from 1 to 10
params = {'max_depth': [1,2,3,4,5,6,7,8,9,10]}
# TODO: Transform 'performance_metric' into a scoring function using 'make_scorer'
scoring_fnc = make_scorer(performance_metric)
# TODO: Create the grid search object
grid = GridSearchCV(regressor,params,scoring_fnc,cv = cv_sets)
# Fit the grid search object to the data to compute the optimal model
grid = grid.fit(X, y)
# Return the optimal model after fitting the data
return grid.best_estimator_
# -
# ### Making Predictions
# Once a model has been trained on a given set of data, it can now be used to make predictions on new sets of input data. In the case of a *decision tree regressor*, the model has learned *what the best questions to ask about the input data are*, and can respond with a prediction for the **target variable**. You can use these predictions to gain information about data where the value of the target variable is unknown — such as data the model was not trained on.
# ### Question 9 - Optimal Model
# _What maximum depth does the optimal model have? How does this result compare to your guess in **Question 6**?_
#
# Run the code block below to fit the decision tree regressor to the training data and produce an optimal model.
# +
# Fit the training data to the model using grid search
reg = fit_model(X_train, y_train)
# Produce the value for 'max_depth'
print "Parameter 'max_depth' is {} for the optimal model.".format(reg.get_params()['max_depth'])
# -
# **Answer: ** The model has an optimal max_depth of 4. This was one max depth higher than my best guess. The difference in predictions can be attributed to GridSearch trying to optimize for the best R^2 score. While I had focused more on trying to find a tradeoff between bias/variance since the difference in testing scores was not very large between a max depth of 3 and 4.
# ### Question 10 - Predicting Selling Prices
# Imagine that you were a real estate agent in the Boston area looking to use this model to help price homes owned by your clients that they wish to sell. You have collected the following information from three of your clients:
#
# | Feature | Client 1 | Client 2 | Client 3 |
# | :---: | :---: | :---: | :---: |
# | Total number of rooms in home | 5 rooms | 4 rooms | 8 rooms |
# | Neighborhood poverty level (as %) | 17% | 32% | 3% |
# | Student-teacher ratio of nearby schools | 15-to-1 | 22-to-1 | 12-to-1 |
# *What price would you recommend each client sell his/her home at? Do these prices seem reasonable given the values for the respective features?*
# **Hint:** Use the statistics you calculated in the **Data Exploration** section to help justify your response.
#
# Run the code block below to have your optimized model make predictions for each client's home.
# +
# Produce a matrix for client data
client_data = [[5, 17, 15], # Client 1
[4, 32, 22], # Client 2
[8, 3, 12]] # Client 3
# Show predictions
for i, price in enumerate(reg.predict(client_data)):
print "Predicted selling price for Client {}'s home: ${:,.2f}".format(i+1, price)
# -
# **Answer: ** The above code recommended the following predictions for each client:
#
# Predicted selling price for Client 1's home: $406,933.33
# Predicted selling price for Client 2's home: $232,200.00
# Predicted selling price for Client 3's home: $938,053.85
#
# Below is the output of the original data exploration statistics with some additional feature means and standard deviations calculated (code for these calculations is in the same block):
#
# Minimum price: $105,000.00
# Maximum price: $1,024,800.00
# Mean price: $454,342.94
# Median price $438,900.00
# Standard deviation of prices: $165,340.28
# Mean RM: 6.24
# RM Standard deviation: 0.64
# Mean LSTAT: 12.94
# LSTAT Standard deviation: 7.07
# Mean PTRATIO: 18.52
# PTRATIO Standard Deviation: 2.11
#
#
# These prices seem reasonable in regards to the basic statistics of the dataset, and seem to be correct in terms of my intuition regarding the data's three features. The predictions are within the min and max prices of the dataset ensuring that we are not trying to extrapolate our model to unobserved feature/price trends.
#
# Looking at these statistics it seems that the values of Client 1's features most closely matches the mean value of each feature in the dataset. The predicted price of Client 1 is also within a standard deviation of the dataset's mean price.
#
# Based on my described intuition Client 2 should have a lower price due to a the client's house having a smaller RM, Higher LSTAT, and a higher PTRATIO. Looking at the feature statistics, each of Client 2's features differ by more than one standard deviation compared to the datasets means. The model agreed with these intuitions and observations by predicting a lower price compared to Client 1.
#
# Based on my described intuition Client 3 should have a higher price due to a the client's house having a larger RM, lower LSTAT, and a lower PTRATIO. Looking at the feature statistics, each of Client 3's features differ by more than one standard deviation compared to the datasets means. The model agreed with these intuitions and observations by predicting a higher price compared to Client 1.
#
#
#
#
#
#
#
# ### Sensitivity
# An optimal model is not necessarily a robust model. Sometimes, a model is either too complex or too simple to sufficiently generalize to new data. Sometimes, a model could use a learning algorithm that is not appropriate for the structure of the data given. Other times, the data itself could be too noisy or contain too few samples to allow a model to adequately capture the target variable — i.e., the model is underfitted. Run the code cell below to run the `fit_model` function ten times with different training and testing sets to see how the prediction for a specific client changes with the data it's trained on.
# +
vs.PredictTrials(features, prices, fit_model, client_data)
# -
# ### Question 11 - Applicability
# *In a few sentences, discuss whether the constructed model should or should not be used in a real-world setting.*
# **Hint:** Some questions to answering:
# - *How relevant today is data that was collected from 1978?*
# - *Are the features present in the data sufficient to describe a home?*
# - *Is the model robust enough to make consistent predictions?*
# - *Would data collected in an urban city like Boston be applicable in a rural city?*
# **Answer: **
#
# The model as it currently stands should not be used in a real world setting. Although the current dataset accounted for inflation, there are alot of other potential factors that may make this dataset inapplicable to the present housing market. The economic depression around 2008, policy changes since 1978, changes in infrastructure, and cultural preferences are just a few items that can question the validity of the dataset to the present. The high variability shown by the large range of $69,000 may indiciate that the model is also not robust enough to reliably make new predictions. Such a large range could result in clients losing out on a significant amount of profit when making a sale. This could potentially be due to not using the most important features for the model's foundation, or not enough features. Some examples of excluded features that could make an impact on housing prices and model robustness include the availability of amenities, and location. These questions require further research to better answer, and to know if the characteristics of the dataset would be similar enough to one collected in areas outside Boston (like a rural city) allowing for its more general use.
# > **Note**: Once you have completed all of the code implementations and successfully answered each question above, you may finalize your work by exporting the iPython Notebook as an HTML document. You can do this by using the menu above and navigating to
# **File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
| boston_housing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Statlog (Heart)
import pandas as pd
import ehrapy as ep
import numpy as np
import warnings
warnings.filterwarnings("ignore")
pd.set_option("display.max_columns", None)
# ## Getting the dataset
# !wget -nc http://archive.ics.uci.edu/ml/machine-learning-databases/statlog/heart/heart.dat -O statlog_heart.data
# !wget -nc http://archive.ics.uci.edu/ml/machine-learning-databases/statlog/heart/heart.doc
# # Data Wrangling
names = ['age', 'sex', 'chest pain type', 'resting blood pressure', 'serum cholestoral',
'fasting blood sugar', 'resting electrocardiographic results', 'maximum heart rate achieved', 'exercise induced angina', 'oldpeak',
'the slope of the peak exercise ST segment', 'number of major vessels', 'thal', 'class']
data = pd.read_csv("statlog_heart.data", sep=" ", names=names)
# Since the dataset does not have a patient ID we add one.
data["patient_id"] = range(1, len(data) + 1)
# Replace the original encoding (1, 2) to (0, 1) in class attribute
data["class"].replace((1, 2), (0, 1), inplace=True)
data.set_index("patient_id", inplace=True)
data.to_csv("statlog_heart_prepared.csv")
# ## ehrapy sanity check
adata = ep.io.read_csv("statlog_heart_prepared.csv",
index_column="patient_id")
adata.var_names
ep.ad.type_overview(adata)
ep.pp.knn_impute(adata)
ep.pp.norm_scale(adata)
ep.pp.pca(adata)
ep.pp.neighbors(adata)
ep.tl.umap(adata)
ep.tl.leiden(adata, resolution=0.5, key_added="leiden_0_5")
ep.pl.umap(adata, color=["leiden_0_5"], title="Leiden 0.5")
ep.pl.umap(adata, color=["class", "age", "sex"], ncols=1, wspace=0.75)
| statlog_heart/statlog_heart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Logging Concurrently
#
# Once we've got the hang of using `rubicon`, we can expand on our project from the *Iris Classifier* example.
# Let's see how a few other popular `scikit-learn` models perform with the Iris dataset. `rubicon` logging is totally
# thread-safe, so we can test a lot of model configurations at once.
# +
from rubicon import Rubicon
root_dir = "./rubicon-root"
rubicon = Rubicon(persistence="filesystem", root_dir=root_dir)
project = rubicon.create_project("Concurrent Experiments", description="Training multiple models in parallel")
print(project)
# -
# For a recap of the contents of the Iris dataset, check out `iris.DESCR` and `iris.data`. We'll put together
# a training dataset using a subset of the data.
# +
from datetime import datetime
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
random_state = int(datetime.utcnow().timestamp())
iris = load_iris()
iris_datasets = train_test_split(iris['data'], iris['target'], random_state=random_state)
# -
# We'll use `run_experiment` to log a new experiment to the provided `project` then train, run and log a model of type
# `classifier_cls` using the training and testing data in `iris_datasets`.
# +
import pandas as pd
from collections import namedtuple
SklearnTrainingMetadata = namedtuple("SklearnTrainingMetadata", "module_name method")
def run_experiment(project, classifier_cls, iris_datasets, **kwargs):
X_train, X_test, y_train, y_test = iris_datasets
experiment = project.log_experiment(
training_metadata=[SklearnTrainingMetadata("sklearn.datasets", "load_iris")],
model_name=classifier_cls.__name__,
tags=[classifier_cls.__name__],
)
for key, value in kwargs.items():
experiment.log_parameter(key, value)
n_features = len(iris.feature_names)
experiment.log_parameter("n_features", n_features)
for name in iris.feature_names:
experiment.log_feature(name)
classifier = classifier_cls(**kwargs)
classifier.fit(X_train, y_train)
classifier.predict(X_test)
accuracy = classifier.score(X_test, y_test)
experiment.log_metric("accuracy", accuracy)
if accuracy >= .95:
experiment.add_tags(["success"])
else:
experiment.add_tags(["failure"])
# -
# This time we'll take a look at two more classifiers, **decision tree** and **k-neighbors**, in addition to the **random forest** classifier we used in the last example. Each classifier will be ran across four sets of parameters (provided as `kwargs` to `run_experiment`), for a total of 12 experiments. Here, we'll build up a list of processes that will run each experiment in parallel.
# +
import multiprocessing
from sklearn.ensemble import RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
processes = []
for n_estimators in [10, 20, 30, 40]:
processes.append(multiprocessing.Process(
target=run_experiment, args=[project, RandomForestClassifier, iris_datasets],
kwargs={"n_estimators": n_estimators, "random_state": random_state},
))
for criterion in ["gini", "entropy"]:
for splitter in ["best", "random"]:
processes.append(multiprocessing.Process(
target=run_experiment, args=[project, DecisionTreeClassifier, iris_datasets],
kwargs={"criterion": criterion, "splitter": splitter, "random_state": random_state},
))
for n_neighbors in [5, 10, 15, 20]:
processes.append(multiprocessing.Process(
target=run_experiment, args=[project, KNeighborsClassifier, iris_datasets],
kwargs={"n_neighbors": n_neighbors},
))
# -
# Let's run all our experiments in parallel!
# +
for process in processes:
process.start()
for process in processes:
process.join()
# -
# Now we can validate that we successfully logged all 12 experiments to our project.
len(project.experiments())
# Let's see which experiments we tagged as successful and what type of model they used.
for e in project.experiments(tags=["success"]):
print(f"experiment {e.id} was successful using a {e.model_name}")
# We can also take a deeper look at any of our experiments.
# +
experiment = project.experiments()[0]
print(f"training_metadata: {SklearnTrainingMetadata(*experiment.training_metadata)}")
print(f"tags: {experiment.tags}")
print("parameters:")
for parameter in experiment.parameters():
print(f"\t{parameter.name}: {parameter.value}")
print("features:")
for feature in experiment.features():
print(f"\t{feature.name}")
print("metrics:")
for metric in experiment.metrics():
print(f"\t{metric.name}: {metric.value}")
# -
# Model developers can take advantage of `rubicon`'s thread-safety to test tons of models at once and collect results
# in a standardized format to analyze which ones performed the best. `rubicon` can even be run in more advanced
# distributed setups, like on a *Dask* cluster.
| notebooks/logging-concurrently.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 분류용 샘플 데이터
# ## 붓꽃 데이터
# - <NAME>의 붓꽃 분류 연구에 기반한 데이터. `load_iris()` 명령으로 로드하며 다음과 같이 구성되어 있다.
# - 타겟 데이터
# - setosa, versicolor, virginica의 세가지 붓꽃 종
# - 특징 자료
# - 꽃받침 길이 (Sepal Length)
# - 꽃받침 폭 (Sepal Width)
# - 꽃잎 길이 (Petal Length)
# - 꽃잎 폭 (Petal Width)
from sklearn.datasets import load_iris
iris = load_iris()
print(iris.DESCR)
df = pd.DataFrame(iris.data, columns = iris.feature_names)
sy = pd.Series(iris.target, dtype='category')
sy = sy.cat.rename_categories(iris.target_names)
df['species'] = sy
df.tail()
# %matplotlib inline
sns.pairplot(df, hue='species')
plt.show()
# ## 뉴스 그룹 텍스트 데이터
# - 20개의 뉴스 그룹 문서 데이터. `fetch_20newsgroups()` 명령으로 로드하며 다음과 같이 구성되어 있다.
# - 타겟 데이터
# - 문서가 속한 뉴스 그룹
# - 특징 데이터
# - 문서 텍스트
from sklearn.datasets import fetch_20newsgroups
newsgroups = fetch_20newsgroups(subset='all')
print(newsgroups.description)
print(newsgroups.keys())
from pprint import pprint
pprint(list(newsgroups.target_names))
print(newsgroups.data[1])
print('-' * 80)
print(newsgroups.target_names[newsgroups.target[1]])
# ## 올리베티 얼굴 사진 데이터
# - AT&T와 캠브리지 대학 전산 연구실에서 공동으로 제작한 얼굴 사진 데이터 베이스의 간략화된 버전. `fetch_olivetti_faces()` 명령으로 로드하며 다음과 같이 구성되어 있다.
# - 타겟 데이터
# - 40명의 개인을 나타내는 식별 번호
# - 특징 데이터
# - 각 개인의 얼굴을 찍은 64x64 해상도의 흑백 이미지
from sklearn.datasets import fetch_olivetti_faces
olivetti = fetch_olivetti_faces()
print(olivetti.DESCR)
print(olivetti.keys())
np.random.seed(0)
N=2; M=5;
fig = plt.figure(figsize=(8, 5))
plt.subplots_adjust(top=1, bottom=0, hspace=0, wspace =0.05)
klist = np.random.choice(range(len(olivetti.data)), N * M)
for i in range(N):
for j in range(M):
k = klist[i*M+j]
ax = fig.add_subplot(N, M, i*M+j+1)
ax.imshow(olivetti.images[k], cmap=plt.cm.bone);
ax.grid(False)
ax.xaxis.set_ticks([])
ax.yaxis.set_ticks([])
plt.title(olivetti.target[k])
plt.tight_layout()
plt.show()
| 2018_07_11_Classification_Sample_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 ('pro_env9')
# language: python
# name: python3
# ---
# <!--
# Copyright 2022 <NAME>
# -->
# # Finite-Size Scaling method by neural network
# The finite-size scaling (FSS) method is a powerful tool for getting universal information of critical phenomena. It estimates universal information from observables of critical phenomena at finite-size systems. In this document, we will introduce a FSS method based on a neural network (NN).
#
# - <NAME> and <NAME>: (preparation).
# ## Finite-size scaling law of a critical phase transition
# The finite-size scaling (FSS) law describes the observables of critical phenomena in a finite-size system. It is written as
# $$
# A(L, T) = L^{-c_2} f[(T-T_c)L^{c_1}],
# $$
# where T is a temperature, L is a system size, A is observable, and $f[]$ is a scaling function. $T_c$ is a critical temperature, and $c_1$ and $c_2$ are critical exponents. We want to find these parameters to describe observables. If we introduce new variables as
# \begin{align*}
# X &= (T-T_c)L^{c_1},\\
# Y &= A/L^{-c_2} = A L^{c_2},
# \end{align*}
# then the FSS law is
# $$
# Y = f[X].
# $$
# In usual, we write $c_1 = 1/\nu$. The $\nu$ is a critical exponent of a correlation length as
# $$
# \xi \propto (T-T_c)^{-\nu}.
# $$
# ## Modeling a scaling function by NN
# Although the scaling function is unclear in the FSS law, we model it in a neural network (NN) as $y=f(x)$. In the following, we use a feed-forward NN with two intermediate layers.
#
# We consider observables in the FSS law as stochastic variables as
# $$
# Y \approx f(X) \pm E
# \ \Rightarrow \ Y \approx N[ f(X), E^2]
# $$
# where $E$ is (an error of A) $\times L^{c_2}$. We optimize parameters $T_c, c_1, c_2$ and parameters of NN to maximize the likelihood.
# ## The Ising model on a square lattice
# For example, we will try to do a FSS analysis for the two-dimensional Ising model on a square lattice.
#
# The Hamiltonian $H$ of the Ising model is
# $$
# H=-\sum_{\langle ij \rangle} \sigma_i \sigma_j,
# $$
# where $\sigma$ is an Ising variable ($\pm 1$) and $\langle ij \rangle$ is a pair of nearest neighbor sites on a square lattice.
#
# The partition function $Z$ is
# $$
# Z = \sum_{\vec{\sigma}} \exp(-H/T),
# $$
# where $T$ is a temperature.
# The magnetization $M$ is
# $$
# M = \sum_i \sigma_i / L^2,
# $$
# where $L$ is a system size. This is an order parameter of the Ising phase transition.
# The susceptibility $\chi$ is
# $$
# \chi = \langle M^2 \rangle.
# $$
#
# The two-dimensional Ising model has a critical phase transition at a critical temperature.
# At the high temperatures, there is no order, and the averaged order parameter is zero. But, the order parameter continuously appears from a critical temperature. Below the critical temperature, the order parameter is finite. In particular, there is an exact solution of the square Ising model by Onsager (1944). Thus, the two-dimensional Ising model is the first non-trivial solvable model of a critical phase transition.
#
# In the case of the square Ising model, the critical temperature $T_c$ is $\frac{1}{2} \log[1+\sqrt{2}]$.
# The critical exponents $c_1$ and $c_2$ of the susceptibility $\chi$ are
# \begin{align*}
# &c_1 = 1/\nu = 1,\\
# &c_2 = -\frac{\gamma}{\nu} = -\frac{7}{4}.
# \end{align*}
#
import math
# Critical temperature of the square Ising model
tc_true = 0.5 * math.log(1+math.sqrt(2))
# For the susceptibility
c1_true, c2_true = (1.0, -1.75)
# ## Doing a FSS analysis by NN
# ### Preparation
# The following modules are necessary.
import torch
import torch.optim as optim
import matplotlib.pyplot as plt
# The following are special modules for doing the FSS method by the NN.
import sys, os
sys.path.append(os.path.abspath("../src"))
import fss_torch
#
# We first declare the dataset and transformer classes.
#
# ### Dataset class
# The dataset class stores the data of susceptibility. Each line of the data file consists of four values (system size, temperature, observable, statistical error). All data are automatically rescaled to do a finite-size scaling. In particular, the temperature is also transformed to a new scale. The two functions are prepared as "transform_t" and "inv_transform_t" in the dataset class.
dataset = fss_torch.fss.Dataset.fromFile(fname="./Data/Ising2D/ising-square-X.dat")
#
# ### Transformer class
# The transformer class defines the transformation of four variables to the new variables (X, Y, E). It needs the initial values of a critical temperature, $c_1$, and $c_2$. The $T_c$ should be inversely transformed to an original scale by "inv_transform_t" function in the dataset class.
rtc, rc1, rc2 = (0.98, 0.9, 0.9)
initial_values = [dataset.transform_t(tc_true * rtc), c1_true * rc1, c2_true * rc2]
transform = fss_torch.fss.Transform(initial_values)
# ### NN Model
# We define the NN model of a scaling function. Here, the number of neurons in intermediate layers is 50.
model = fss_torch.nsa_util.MLP(hidden_sizes=[50, 50])
# ### Optimizer
# We set the parameters of the NN model in an optimizer. The parameters of the transformer are also added in the optimizer.
optimizer = optim.Adam(model.parameters())
optimizer.add_param_group({"params": transform.parameters(), "lr": 0.01})
# ### Optimizing all paramters
# The optimization steps are
# 1. transform original values to (X, Y, E) in a scaling function.
# 2. calculate a loss function by the NN model.
# 3. update all parameters.
losses = []
results = []
loss_fn = torch.nn.GaussianNLLLoss()
for epoch in range(5000):
optimizer.zero_grad()
new_data = transform(dataset.data)
X = fss_torch.nsa_util.get_column(new_data, 0)
Y = fss_torch.nsa_util.get_column(new_data, 1)
E = fss_torch.nsa_util.get_column(new_data, 2)
loss = loss_fn(model(X), Y, E * E)
loss.backward()
optimizer.step()
losses.append(loss.item())
results.append([dataset.inv_transform_t(transform.tc), transform.c1, transform.c2])
# We check results of $T_c$, $c_1$ and $c_2$.
tc_results, c1_results, c2_results = (dataset.inv_transform_t(transform.tc), transform.c1, transform.c2)
print("Exact values : Tc = {}, c1 = {}, c2 = {}".format(tc_true, c1_true, c2_true))
print("Results of FSS : Tc = {}, c1 = {}, c2 = {}".format(tc_results, c1_results, c2_results))
# We plot a scaling function, loss and optimization process of paramters $T_c$ and $c_2$.
# +
fig, ax0 = plt.subplots(figsize=(5,5))
new_data = transform(dataset.data)
X, Y, E = new_data[:,0], new_data[:,1], new_data[:,2]
test_x = torch.linspace(-1, 1, 51).view(-1,1)
observed_pred = model(test_x)
ax0.set_title("Scaling function")
ax0.set_xlabel("X = (T-Tc)L^c1")
ax0.set_ylabel("Y = A L^c2")
ax0.plot(test_x, observed_pred.detach().numpy())
for L in [0.25, 0.5, 1.0]:
index_L = dataset.data[:,0]==L
ax0.scatter(X[index_L].detach().numpy(), Y[index_L].detach().numpy())
fig, ax1 = plt.subplots(figsize=(5,5))
ax1.set_title("Loss")
ax1.set_xlabel("epochs")
ax1.set_ylabel("loss")
ax1.set_xscale("log")
ax1.plot(losses)
fig, ax2 = plt.subplots(figsize=(5,5))
X = [r[0] for r in results[::100]]
Y = [r[2] for r in results[::100]]
ax2.set_title("Optimization of parameters")
ax2.set_xlabel("Tc")
ax2.set_ylabel("c2")
ax2.plot(X, Y, marker="o")
| examples/nsa.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Train Toxicity Model
# This notebook trains a model to detect toxicity in online comments. It uses a CNN architecture for text classification trained on the [Wikipedia Talk Labels: Toxicity dataset](https://figshare.com/articles/Wikipedia_Talk_Labels_Toxicity/4563973) and pre-trained GloVe embeddings which can be found at:
# http://nlp.stanford.edu/data/glove.6B.zip
# (source page: http://nlp.stanford.edu/projects/glove/).
#
# This model is a modification of [example code](https://github.com/fchollet/keras/blob/master/examples/pretrained_word_embeddings.py) found in the [Keras Github repository](https://github.com/fchollet/keras) and released under an [MIT license](https://github.com/fchollet/keras/blob/master/LICENSE). For further details of this license, find it [online](https://github.com/fchollet/keras/blob/master/LICENSE) or in this repository in the file KERAS_LICENSE.
# ## Usage Instructions
# (TODO: nthain) - Move to README
#
# Prior to running the notebook, you must:
#
# * Download the [Wikipedia Talk Labels: Toxicity dataset](https://figshare.com/articles/Wikipedia_Talk_Labels_Toxicity/4563973)
# * Download pre-trained [GloVe embeddings](http://nlp.stanford.edu/data/glove.6B.zip)
# * (optional) To skip the training step, you will need to download a model and tokenizer file. We are looking into the appropriate means for distributing these (sometimes large) files.
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import pandas as pd
from model_tool import ToxModel
# -
# ## Load Data
# +
SPLITS = ['train', 'dev', 'test']
wiki = {}
debias = {}
random = {}
for split in SPLITS:
wiki[split] = '../data/wiki_%s.csv' % split
debias[split] = '../data/wiki_debias_%s.csv' % split
random[split] = '../data/wiki_debias_random_%s.csv' % split
# -
# ## Train Models
hparams = {'epochs': 4}
# ### Random model
MODEL_NAME = 'cnn_debias_random_tox_v3'
debias_random_model = ToxModel(hparams=hparams)
debias_random_model.train(random['train'], random['dev'], text_column = 'comment', label_column = 'is_toxic', model_name = MODEL_NAME)
random_test = pd.read_csv(random['test'])
debias_random_model.score_auc(random_test['comment'], random_test['is_toxic'])
# ### Plain wikipedia model
MODEL_NAME = 'cnn_wiki_tox_v3'
wiki_model = ToxModel(hparams=hparams)
wiki_model.train(wiki['train'], wiki['dev'], text_column = 'comment', label_column = 'is_toxic', model_name = MODEL_NAME)
wiki_test = pd.read_csv(wiki['test'])
wiki_model.score_auc(wiki_test['comment'], wiki_test['is_toxic'])
# ### Debiased model
MODEL_NAME = 'cnn_debias_tox_v3'
debias_model = ToxModel(hparams=hparams)
debias_model.train(debias['train'], debias['dev'], text_column = 'comment', label_column = 'is_toxic', model_name = MODEL_NAME)
debias_test = pd.read_csv(debias['test'])
debias_model.score_auc(debias_test['comment'], debias_test['is_toxic'])
| unintended_ml_bias/Train_Toxicity_Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LETS GROW MORE
#
# ## LGM VIP - Data Science September-2021
#
# ### Name: <NAME>.
#
# #### Project Name : Image to Pencil Sketch with Python. ( BEGINNER LEVEL TASK: 4) .
#
#
#
# PROJECT DESCRIPTION : We need to read the image in RBG format and then convert it to a grayscale image. This will turn an image into a classic black and white photo. Then the next thing to do is invert the grayscale image also called negative image, this will be our inverted grayscale image. Inversion can be used to enhance details. Then we can finally create the pencil sketch by mixing the grayscale image with the inverted blurry image. This can be done by dividing the grayscale image by the inverted blurry image. Since images are just arrays, we can easily do this programmatically using the divide function from the cv2 library in Python.
# STEP 1) IMPORTING LIBRARIES.
# +
import cv2
cv2.__version__
import matplotlib.pyplot as plt
print("sucessfully imported the libraries")
# +
# Reading an image
from IPython import display
display.Image("C:/Users/LENOVO\Desktop/LETS GROW MORE TASKS/Shaunak task/BEGINNER TASKS/IMAGE TO SKETCH USING PYTHON/apj_abdul_kalam.jpg")
# +
img=cv2.imread('C:/Users/LENOVO\Desktop/LETS GROW MORE TASKS/Shaunak task/BEGINNER TASKS/IMAGE TO SKETCH USING PYTHON/apj_abdul_kalam.jpg')
# show image format (basically a 3-d array of pixel color info, in BGR format)
print(img)
# +
# converting the image to RGB color for matplotlib
img1=cv2.cvtColor(img,cv2.COLOR_BGR2RGB)
# showing the image with matplotlib
plt.imshow(img)
# -
plt.imshow(img1)
# +
# convert image to grayscale
gray_image=cv2.cvtColor(img,cv2.COLOR_RGB2GRAY)
# grayscale image represented as a 2-d array
print(gray_image)
print("\n")
print("--------------------------------------------------------------------")
# we have to change grayscale back to RGB for plt.imshow()
plt.imshow(gray_image)
# -
#invert the grayscale image
inverted_image =255-gray_image
plt.imshow(inverted_image)
#blur the inverted image
blurred_image =cv2.GaussianBlur(inverted_image,(21,21),0)
plt.imshow(blurred_image)
inverted_blurred=255-blurred_image
pencil_sketch=cv2.divide(gray_image,inverted_blurred,scale=255.0)
plt.imshow(pencil_sketch)
# * THE PENCIL SKETCH IS READY.
# * THE END..!!
| Image to Pencil Sketch with Python (BEGINNER LEVEL TASK : 4)/Image to Pencil Sketch with Python..ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sparse covariance estimation for Gaussian variables
#
# A derivative work by <NAME>, 5/22/2014.<br>
# Adapted (with significant improvements and fixes) from the CVX example of the same name, by <NAME>, 4/24/2008.
#
# Topic References:
#
# * Section 7.1.1, Boyd & Vandenberghe "Convex Optimization"
#
# ## Introduction
#
# Suppose $y \in \mathbf{\mbox{R}}^n$ is a Gaussian random variable with zero mean and
# covariance matrix $R = \mathbf{\mbox{E}}[yy^T]$, with sparse inverse $S = R^{-1}$
# ($S_{ij} = 0$ means that $y_i$ and $y_j$ are conditionally independent).
# We want to estimate the covariance matrix $R$ based on $N$ independent
# samples $y_1,\dots,y_N$ drawn from the distribution, and using prior knowledge
# that $S$ is sparse
#
# A good heuristic for estimating $R$ is to solve the problem
# $$\begin{array}{ll}
# \mbox{minimize} & \log \det(S) - \mbox{tr}(SY) \\
# \mbox{subject to} & \sum_{i=1}^n \sum_{j=1}^n |S_{ij}| \le \alpha \\
# & S \succeq 0,
# \end{array}$$
# where $Y$ is the sample covariance of $y_1,\dots,y_N$, and $\alpha$ is a sparsity
# parameter to be chosen or tuned.
#
# ## Generate problem data
# +
import cvxpy as cp
import numpy as np
import scipy as scipy
# Fix random number generator so we can repeat the experiment.
np.random.seed(0)
# Dimension of matrix.
n = 10
# Number of samples, y_i
N = 1000
# Create sparse, symmetric PSD matrix S
A = np.random.randn(n, n) # Unit normal gaussian distribution.
A[scipy.sparse.rand(n, n, 0.85).todense().nonzero()] = 0 # Sparsen the matrix.
Strue = A.dot(A.T) + 0.05 * np.eye(n) # Force strict pos. def.
# Create the covariance matrix associated with S.
R = np.linalg.inv(Strue)
# Create samples y_i from the distribution with covariance R.
y_sample = scipy.linalg.sqrtm(R).dot(np.random.randn(n, N))
# Calculate the sample covariance matrix.
Y = np.cov(y_sample)
# -
# ## Solve for several $\alpha$ values
# +
# The alpha values for each attempt at generating a sparse inverse cov. matrix.
alphas = [10, 2, 1]
# Empty list of result matrixes S
Ss = []
# Solve the optimization problem for each value of alpha.
for alpha in alphas:
# Create a variable that is constrained to the positive semidefinite cone.
S = cp.Variable(shape=(n,n), PSD=True)
# Form the logdet(S) - tr(SY) objective. Note the use of a set
# comprehension to form a set of the diagonal elements of S*Y, and the
# native sum function, which is compatible with cvxpy, to compute the trace.
# TODO: If a cvxpy trace operator becomes available, use it!
obj = cp.Maximize(cp.log_det(S) - sum([(S*Y)[i, i] for i in range(n)]))
# Set constraint.
constraints = [cp.sum(cp.abs(S)) <= alpha]
# Form and solve optimization problem
prob = cp.Problem(obj, constraints)
prob.solve(solver=cp.CVXOPT)
if prob.status != cp.OPTIMAL:
raise Exception('CVXPY Error')
# If the covariance matrix R is desired, here is how it to create it.
R_hat = np.linalg.inv(S.value)
# Threshold S element values to enforce exact zeros:
S = S.value
S[abs(S) <= 1e-4] = 0
# Store this S in the list of results for later plotting.
Ss += [S]
print('Completed optimization parameterized by alpha = {}, obj value = {}'.format(alpha, obj.value))
# -
# ## Result plots
# +
import matplotlib.pyplot as plt
# Show plot inline in ipython.
# %matplotlib inline
# Plot properties.
plt.rc('text', usetex=True)
plt.rc('font', family='serif')
# Create figure.
plt.figure()
plt.figure(figsize=(12, 12))
# Plot sparsity pattern for the true covariance matrix.
plt.subplot(2, 2, 1)
plt.spy(Strue)
plt.title('Inverse of true covariance matrix', fontsize=16)
# Plot sparsity pattern for each result, corresponding to a specific alpha.
for i in range(len(alphas)):
plt.subplot(2, 2, 2+i)
plt.spy(Ss[i])
plt.title('Estimated inv. cov matrix, $\\alpha$={}'.format(alphas[i]), fontsize=16)
| examples/notebooks/WWW/sparse_covariance_est.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
import sys
import seaborn as sns
from sklearn.metrics import silhouette_score, silhouette_samples
import warnings
warnings.simplefilter('ignore')
# Plotting settings
sns.set_style("white")
sns.set_style("ticks")
mpl.rcParams['font.sans-serif'] = 'DejaVu Sans'
mpl.rcParams['pdf.fonttype'] = 42
pkg_dir = '/home/mrossol/NaTGenPD'
#pkg_dir = '..'
sys.path.append(pkg_dir)
import NaTGenPD as npd
import NaTGenPD.cluster as cluster
data_dir = '/scratch/mrossol/CEMS'
#data_dir = '/Users/mrossol/Downloads/CEMS'
# + [markdown] heading_collapsed=true
# # Cluster dev
# + hidden=true
comb_file = os.path.join(data_dir, 'SMOKE_Clean_2016-2017.h5')
with npd.CEMS(comb_file, mode='r') as f:
#ng_ct = f['CT (NG)']
#boiler = f['Boiler (Coal)']
ng_cc = f['CC (NG)']
#oil_cc = f['CC (Oil)']
#oil_ct = f['CT (Oil)']
# + [markdown] hidden=true
# ### Oil CT
# + hidden=true
ct_df = oil_ct['1355_9']
#unit_df = ng_ct['120_CT5']
print(len(ct_df))
ct_df.plot.scatter(x='load', y='heat_rate',
xlim=(0, ct_df['load'].max() * 1.05),
ylim=(0, ct_df['heat_rate'].max() * 1.05))
plt.show()
c = cluster.SingleCluster(ct_df)
arr = c.get_data(['load', 'heat_rate'])
cluster_params = c._cluster(arr, min_samples=16, eps=.6)
print(cluster_params[1:])
labels = cluster_params[0]
c_df = ct_df[['load', 'heat_rate']].copy()
c_df['label'] = cluster_params[0]
sns.scatterplot(x='load', y='heat_rate', hue='label', data=c_df,
palette='Paired')
plt.show()
# + hidden=true
logger = npd.setup_logger('NaTGenPD.cluster', log_level='DEBUG')
ct_df = oil_ct['1355_9']
#unit_df = ng_ct['120_CT5']
print(len(ct_df))
ct_df.plot.scatter(x='load', y='heat_rate',
xlim=(0, ct_df['load'].max() * 1.05),
ylim=(0, ct_df['heat_rate'].max() * 1.05))
plt.show()
c = cluster.SingleCluster(ct_df)
arr = c.get_data(['load', 'heat_rate'])
cluster_params = c.optimize_clusters(min_samples=16)
print(cluster_params[1:])
labels = cluster_params[0]
c_df = ct_df[['load', 'heat_rate']].copy()
c_df['label'] = cluster_params[0]
sns.scatterplot(x='load', y='heat_rate', hue='label', data=c_df,
palette='Paired')
plt.show()
c_df.hist(column='load', bins=100)
plt.show()
# + [markdown] hidden=true
# ### Missing CCs in cc_map
# + hidden=true
path = os.path.join(data_dir, 'Mappings', 'CEMS_mergeguide.csv')
cc_map = pd.read_csv(path)
cc_map['cc_unit'] = cc_map['EIAPlant'].astype(str) + '_' + cc_map['EIAUnit'].astype(str)
ng_ccs = ng_cc._unit_dfs.size()
# + hidden=true
pos = ng_ccs.index.isin(cc_map['cc_unit'])
missing_cts = ng_ccs.loc[~pos]
missing_cts = pd.DataFrame({'CEMSUnit': missing_cts.index})
# + hidden=true
CEMS_ccs = pd.read_csv(path)
CEMS_ccs = pd.concat((CEMS_ccs, missing_cts), sort=True)
# + hidden=true
out_path = '/home/mrossol/CEMS_ccs.csv'
CEMS_ccs.to_csv(out_path, index=False)
# + [markdown] hidden=true
# ## Coal Boiler
# + hidden=true
boiler_df = boiler['1001_1']
print(len(boiler_df))
boiler_df.plot.scatter(x='load', y='heat_rate',
xlim=(0, boiler_df['load'].max() * 1.05),
ylim=(0, boiler_df['heat_rate'].max() * 1.05))
plt.show()
c = cluster.SingleCluster(boiler_df)
arr = c.get_data(['load', 'heat_rate'])
cluster_params = c.optimize_clusters(min_samples=13)
print(cluster_params[1:])
labels = cluster_params[0]
print('score = {:.4f}'.format(c.cluster_score(arr, labels)))
c_df = boiler_df[['load', 'heat_rate']].copy()
c_df['label'] = cluster_params[0]
sns.scatterplot(x='load', y='heat_rate', hue='label', data=c_df,
palette='Paired')
plt.show()
c_df.hist(column='load', bins=100)
plt.show()
# + hidden=true
boiler_df = boiler['991_5']
print(len(boiler_df))
boiler_df.plot.scatter(x='load', y='heat_rate',
xlim=(0, boiler_df['load'].max() * 1.05),
ylim=(0, boiler_df['heat_rate'].max() * 1.05))
plt.show()
c = cluster.SingleCluster(boiler_df)
arr = c.get_data(['load', 'heat_rate'])
cluster_params = c.optimize_clusters(min_samples=16)
print(cluster_params[1:])
labels = cluster_params[0]
print('score = {:.4f}'.format(c.cluster_score(arr, labels)))
c_df = boiler_df[['load', 'heat_rate']].copy()
c_df['label'] = cluster_params[0]
sns.scatterplot(x='load', y='heat_rate', hue='label', data=c_df,
palette='Paired')
plt.show()
c_df.hist(column='load', bins=100)
plt.show()
# + [markdown] hidden=true
# ## NG CT
# + hidden=true
ct_df = ng_ct['1239_12']
#unit_df = ng_ct['120_CT5']
print(len(ct_df))
ct_df.plot.scatter(x='load', y='heat_rate',
xlim=(0, ct_df['load'].max() * 1.05),
ylim=(0, ct_df['heat_rate'].max() * 1.05))
plt.show()
c = cluster.SingleCluster(ct_df)
arr = c.get_data(['load', 'heat_rate'])
cluster_params = c.optimize_clusters(min_samples=16)
print(cluster_params[1:])
labels = cluster_params[0]
c_df = ct_df[['load', 'heat_rate']].copy()
c_df['label'] = cluster_params[0]
sns.scatterplot(x='load', y='heat_rate', hue='label', data=c_df,
palette='Paired')
plt.show()
c_df.hist(column='load', bins=100)
plt.show()
# + hidden=true
ct_df = ng_ct['120_CT5']
print(len(ct_df))
ct_df.plot.scatter(x='load', y='heat_rate',
xlim=(0, ct_df['load'].max() * 1.05),
ylim=(0, ct_df['heat_rate'].max() * 1.05))
plt.show()
c = cluster.SingleCluster(ct_df)
arr = c.get_data(['load', 'heat_rate'])
cluster_params = c.optimize_clusters(min_samples=17)
labels = cluster_params[0]
c_df = ct_df[['load', 'heat_rate']].copy()
c_df['label'] = cluster_params[0]
sns.scatterplot(x='load', y='heat_rate', hue='label', data=c_df,
palette='Paired')
plt.show()
c_df.hist(column='load', bins=100)
plt.show()
# + [markdown] hidden=true
# ## NG CC
# + hidden=true
cc_df = ng_cc['55411_CC1']
print(len(cc_df))
cc_df.plot.scatter(x='load', y='heat_rate', c='cts',
colormap='rainbow',
xlim=(0, cc_df['load'].max() * 1.05),
ylim=(0, cc_df['heat_rate'].max() * 1.05))
plt.show()
c = cluster.ClusterCC(cc_df)
arr = c.unit_df[['load', 'heat_rate', 'cts']].values
labels = c.optimize_clusters(min_samples=17)
c_df = cc_df[['load', 'heat_rate']].copy()
c_df['label'] = labels
sns.scatterplot(x='load', y='heat_rate', hue='label', data=c_df,
palette='Paired')
plt.show()
# + hidden=true
cc_df = ng_cc['1007_CC1']
print(len(cc_df))
cc_df.plot.scatter(x='load', y='heat_rate', c='cts',
colormap='rainbow',
xlim=(0, cc_df['load'].max() * 1.05),
ylim=(0, cc_df['heat_rate'].max() * 1.05))
plt.show()
c = cluster.ClusterCC(cc_df)
arr = c.unit_df[['load', 'heat_rate', 'cts']].values
labels = c.optimize_clusters(min_samples=17)
c_df = cc_df[['load', 'heat_rate']].copy()
c_df['label'] = labels
sns.scatterplot(x='load', y='heat_rate', hue='label', data=c_df,
palette='Paired')
plt.show()
# + [markdown] heading_collapsed=true
# # Fit Filter
# + hidden=true
def round_to(data, val):
"""
round data to nearest val
Parameters
----------
data : 'ndarray', 'float'
Input data
perc : 'float'
Value to round to the nearest
Returns
-------
'ndarray', 'float
Rounded data, same type as data
"""
return data // val * val
# + hidden=true
fit_dir = os.path.join(data_dir, "CEMS_Fits")
hr_fits = npd.Fits(fit_dir)
# + hidden=true
for g_type in hr_fits.group_types:
group_df = hr_fits[g_type]
fit_units = np.sum(~group_df['a0'].isnull())
print('{}: {}'.format(g_type, fit_units))
# -
min_h
# +
group_fits = hr_fits['Boiler (Coal)']
stdev_multiplier=2
min_hr = group_fits.apply(npd.filter.FitFilter._get_hr_min, axis=1).dropna()
mean = min_hr.mean()
stdev = min_hr.std()
thresh = np.array([-stdev_multiplier, stdev_multiplier]) * stdev + mean
# + hidden=true
group_fits = hr_fits['Boiler (Coal)']
stdev_multiplier=2
min_hr = group_fits.apply(npd.filter.FitFilter._get_hr_min, axis=1).dropna()
mean = min_hr.mean()
stdev = min_hr.std()
thresh = np.array([-stdev_multiplier, stdev_multiplier]) * stdev + mean
print(thresh)
fig = plt.figure(figsize=(6, 4), dpi=100)
axis = fig.add_subplot(111)
axis.hist(min_hr, bins='auto')
axis.plot(thresh[[0, 0]], [0, 90], 'r--')
axis.plot(thresh[[1, 1]], [0, 90], 'r--')
for ax in ['top', 'bottom', 'left', 'right']:
axis.spines[ax].set_linewidth(1)
axis.tick_params(axis='both', labelsize=8, width=1,
length=4)
axis.set_xlabel('Min Heat Rate (mmBTU/MWh)', fontsize=10)
axis.set_ylabel('Counts', fontsize=10)
axis.set_xlim(0, 20)
fig.tight_layout()
plt.show()
plt.close()
# + hidden=true
group_fits = hr_fits['CT (NG)']
stdev_multiplier=2
min_hr = group_fits.apply(npd.filter.FitFilter._get_hr_min, axis=1).dropna()
mean = min_hr.mean()
stdev = min_hr.std()
thresh = np.array([-stdev_multiplier, stdev_multiplier]) * stdev + mean
print(thresh)
fig = plt.figure(figsize=(6, 4), dpi=100)
axis = fig.add_subplot(111)
axis.hist(min_hr, bins='auto')
axis.plot(thresh[[0, 0]], [0, 90], 'r--')
axis.plot(thresh[[1, 1]], [0, 90], 'r--')
for ax in ['top', 'bottom', 'left', 'right']:
axis.spines[ax].set_linewidth(1)
axis.tick_params(axis='both', labelsize=8, width=1,
length=4)
axis.set_xlabel('Min Heat Rate (mmBTU/MWh)', fontsize=10)
axis.set_ylabel('Counts', fontsize=10)
axis.set_xlim(0, 20)
fig.tight_layout()
plt.show()
plt.close()
# + hidden=true
cc_fits = hr_fits['CC (NG)']
cc_fits['cc_id'] = cc_fits['unit_id'].str.split('-').str[0]
cc_fits = cc_fits.set_index('cc_id')
cc_min_hr = cc_fits.apply(npd.filter.FitFilter._get_hr_min, axis=1).dropna().to_frame().reset_index()
min_hr = cc_min_hr.groupby('cc_id').min().values
fig = plt.figure(figsize=(6, 4), dpi=100)
axis = fig.add_subplot(111)
axis.hist(min_hr, bins='auto')
for ax in ['top', 'bottom', 'left', 'right']:
axis.spines[ax].set_linewidth(1)
axis.tick_params(axis='both', labelsize=12, width=1,
length=4)
axis.set_xlabel('Min Heat Rate (mmBTU/MWh)', fontsize=14)
axis.set_ylabel('Counts', fontsize=14)
axis.set_xlim(0, 15)
fig.tight_layout()
plt.show()
plt.close()
min_hr = min_hr[min_hr < 9.5]
mean = min_hr.mean()
stdev = min_hr.std()
thresh = np.array([-2, 2]) * stdev + mean
print(thresh)
fig = plt.figure(figsize=(6, 4), dpi=100)
axis = fig.add_subplot(111)
axis.hist(min_hr, bins='auto')
axis.plot(thresh[[0, 0]], [0, 90], 'r--')
axis.plot(thresh[[1, 1]], [0, 90], 'r--')
for ax in ['top', 'bottom', 'left', 'right']:
axis.spines[ax].set_linewidth(1)
axis.tick_params(axis='both', labelsize=12, width=1,
length=4)
axis.set_xlabel('Min Heat Rate (mmBTU/MWh)', fontsize=14)
axis.set_ylabel('Counts', fontsize=14)
axis.set_xlim(0, 10)
fig.tight_layout()
plt.show()
plt.close()
# + hidden=true
cc_df = hr_fits['CC (NG)']
cc_df['cc_id'] = cc_df['unit_id'].str.split('-').str[0]
cc_df = cc_df.set_index('cc_id')
cc_min_hr = cc_df.apply(npd.filter.FitFilter._get_hr_min, axis=1).dropna().to_frame().reset_index()
min_hr = cc_min_hr.groupby('cc_id').min()
# +
failed_units = npd.filter.FitFilter._min_hr_filter(min_hr, threshold=(None, 9))
filter_cols = [c for c in cc_df.columns
if c.startswith(('a', 'heat_rate', 'load'))
and c not in ['load_min', 'load_max']]
cc_df.loc[failed_units, filter_cols] = None
# -
cc_df
failed_units.shape
# + hidden=true
from scipy.optimize import curve_fit
def gauss(x, mu, sigma, A):
return A * np.exp(-(x - mu)**2 / (2 * sigma))
def bimodal(x, mu1, sigma1, A1, mu2, sigma2, A2):
return gauss(x, mu1, sigma1, A1) + gauss(x, mu2, sigma2, A2)
cc_fits = hr_fits['CC (NG)']
cc_fits['cc_id'] = cc_fits['unit_id'].str.split('-').str[0]
cc_fits = cc_fits.set_index('cc_id')
cc_min_hr = cc_fits.apply(npd.filter.FitFilter.get_hr_min, axis=1).dropna().to_frame().reset_index()
min_hr = cc_min_hr.groupby('cc_id').min().values
bad_min_hr = min_hr[min_hr < 10]
bin_s = 0.25
# Extract histogram values for bins of size bin_s
bins = np.arange(round_to(bad_min_hr.min(), bin_s) - bin_s/2,
round_to(bad_min_hr.max(), bin_s) + bin_s/2, bin_s)
y, _ = np.histogram(bad_min_hr, bins=bins)
x_g = (bins[1:] + bins[:-1])/2
expected = (np.mean(bad_min_hr), np.std(bad_min_hr), np.max(y))
g_params, _ = curve_fit(gauss, x_g, y, expected)
fit_g = gauss(x_g, *g_params)
print('Gaussian Fit')
print('\t {}'.format(g_params))
print('\t - Cut off = {:.2f}'.format(g_params[0] + 2 * g_params[1]))
# Extract histogram values for bins of size bin_s
bins = np.arange(round_to(min_hr.min(), bin_s) - bin_s/2,
round_to(min_hr.max(), bin_s) + bin_s/2, bin_s)
y, _ = np.histogram(min_hr, bins=bins)
x_b = (bins[1:] + bins[:-1])/2
expected = [np.mean(min_hr), np.std(min_hr), np.max(y)]
b_params, _ = curve_fit(bimodal, x_b, y, expected*2)
fit_b = bimodal(x_b, *b_params)
print('Bimodal Fit')
print('\t {}'.format(b_params))
print('\t - Cut off = {:.2f}'.format(b_params[0] + 2 * b_params[1]))
fig = plt.figure(figsize=(8, 6), dpi=100)
axis = fig.add_subplot(111)
mpl.rcParams['font.sans-serif'] = 'Arial'
mpl.rcParams['pdf.fonttype'] = 42
axis.hist(min_hr, bins=bins)
axis.plot(x_b, fit_b, '-r')
axis.plot(x_g, fit_g, '--g')
for ax in ['top', 'bottom', 'left', 'right']:
axis.spines[ax].set_linewidth(2)
axis.tick_params(axis='both', labelsize=16, width=2,
length=8)
axis.set_xlabel('Min Heat Rate (mmBTU/MWh)', fontsize=18)
axis.set_ylabel('Counts', fontsize=18)
axis.set_xlim(0, 20)
fig.tight_layout()
plt.show()
plt.close()
# + hidden=true
| notebooks/CEMS Heat Rate Dev.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # Probability
# :label:`sec_prob`
#
# In some form or another, machine learning is all about making predictions.
# We might want to predict the *probability* of a patient suffering a heart attack in the next year, given their clinical history. In anomaly detection, we might want to assess how *likely* a set of readings from an airplane's jet engine would be, were it operating normally. In reinforcement learning, we want an agent to act intelligently in an environment. This means we need to think about the probability of getting a high reward under each of the available actions. And when we build recommender systems we also need to think about probability. For example, say *hypothetically* that we worked for a large online bookseller. We might want to estimate the probability that a particular user would buy a particular book. For this we need to use the language of probability.
# Entire courses, majors, theses, careers, and even departments, are devoted to probability. So naturally, our goal in this section is not to teach the whole subject. Instead we hope to get you off the ground, to teach you just enough that you can start building your first deep learning models, and to give you enough of a flavor for the subject that you can begin to explore it on your own if you wish.
#
# We have already invoked probabilities in previous sections without articulating what precisely they are or giving a concrete example. Let us get more serious now by considering the first case: distinguishing cats and dogs based on photographs. This might sound simple but it is actually a formidable challenge. To start with, the difficulty of the problem may depend on the resolution of the image.
#
# 
# :width:`300px`
# :label:`fig_cat_dog`
#
# As shown in :numref:`fig_cat_dog`,
# while it is easy for humans to recognize cats and dogs at the resolution of $160 \times 160$ pixels,
# it becomes challenging at $40 \times 40$ pixels and next to impossible at $10 \times 10$ pixels. In
# other words, our ability to tell cats and dogs apart at a large distance (and thus low resolution) might approach uninformed guessing. Probability gives us a
# formal way of reasoning about our level of certainty.
# If we are completely sure
# that the image depicts a cat, we say that the *probability* that the corresponding label $y$ is "cat", denoted $P(y=$ "cat"$)$ equals $1$.
# If we had no evidence to suggest that $y =$ "cat" or that $y =$ "dog", then we might say that the two possibilities were equally
# *likely* expressing this as $P(y=$ "cat"$) = P(y=$ "dog"$) = 0.5$. If we were reasonably
# confident, but not sure that the image depicted a cat, we might assign a
# probability $0.5 < P(y=$ "cat"$) < 1$.
#
# Now consider the second case: given some weather monitoring data, we want to predict the probability that it will rain in Taipei tomorrow. If it is summertime, the rain might come with probability 0.5.
#
# In both cases, we have some value of interest. And in both cases we are uncertain about the outcome.
# But there is a key difference between the two cases. In this first case, the image is in fact either a dog or a cat, and we just do not know which. In the second case, the outcome may actually be a random event, if you believe in such things (and most physicists do). So probability is a flexible language for reasoning about our level of certainty, and it can be applied effectively in a broad set of contexts.
#
# ## Basic Probability Theory
#
# Say that we cast a die and want to know what the chance is of seeing a 1 rather than another digit. If the die is fair, all the six outcomes $\{1, \ldots, 6\}$ are equally likely to occur, and thus we would see a $1$ in one out of six cases. Formally we state that $1$ occurs with probability $\frac{1}{6}$.
#
# For a real die that we receive from a factory, we might not know those proportions and we would need to check whether it is tainted. The only way to investigate the die is by casting it many times and recording the outcomes. For each cast of the die, we will observe a value in $\{1, \ldots, 6\}$. Given these outcomes, we want to investigate the probability of observing each outcome.
#
# One natural approach for each value is to take the
# individual count for that value and to divide it by the total number of tosses.
# This gives us an *estimate* of the probability of a given *event*. The *law of
# large numbers* tell us that as the number of tosses grows this estimate will draw closer and closer to the true underlying probability. Before going into the details of what is going here, let us try it out.
#
# To start, let us import the necessary packages.
#
# + origin_pos=1 tab=["mxnet"]
# %matplotlib inline
import random
from mxnet import np, npx
from d2l import mxnet as d2l
npx.set_np()
# + [markdown] origin_pos=4
# Next, we will want to be able to cast the die. In statistics we call this process
# of drawing examples from probability distributions *sampling*.
# The distribution
# that assigns probabilities to a number of discrete choices is called the
# *multinomial distribution*. We will give a more formal definition of
# *distribution* later, but at a high level, think of it as just an assignment of
# probabilities to events.
#
# To draw a single sample, we simply pass in a vector of probabilities.
# The output is another vector of the same length:
# its value at index $i$ is the number of times the sampling outcome corresponds to $i$.
#
# + origin_pos=5 tab=["mxnet"]
fair_probs = [1.0 / 6] * 6
np.random.multinomial(1, fair_probs)
# + [markdown] origin_pos=8
# If you run the sampler a bunch of times, you will find that you get out random
# values each time. As with estimating the fairness of a die, we often want to
# generate many samples from the same distribution. It would be unbearably slow to
# do this with a Python `for` loop, so the function we are using supports drawing
# multiple samples at once, returning an array of independent samples in any shape
# we might desire.
#
# + origin_pos=9 tab=["mxnet"]
np.random.multinomial(10, fair_probs)
# + [markdown] origin_pos=12
# Now that we know how to sample rolls of a die, we can simulate 1000 rolls. We
# can then go through and count, after each of the 1000 rolls, how many times each
# number was rolled.
# Specifically, we calculate the relative frequency as the estimate of the true probability.
#
# + origin_pos=13 tab=["mxnet"]
counts = np.random.multinomial(1000, fair_probs).astype(np.float32)
counts / 1000
# + [markdown] origin_pos=16
# Because we generated the data from a fair die, we know that each outcome has true probability $\frac{1}{6}$, roughly $0.167$, so the above output estimates look good.
#
# We can also visualize how these probabilities converge over time towards the true probability.
# Let us conduct 500 groups of experiments where each group draws 10 samples.
#
# + origin_pos=17 tab=["mxnet"]
counts = np.random.multinomial(10, fair_probs, size=500)
cum_counts = counts.astype(np.float32).cumsum(axis=0)
estimates = cum_counts / cum_counts.sum(axis=1, keepdims=True)
d2l.set_figsize((6, 4.5))
for i in range(6):
d2l.plt.plot(estimates[:, i].asnumpy(),
label=("P(die=" + str(i + 1) + ")"))
d2l.plt.axhline(y=0.167, color='black', linestyle='dashed')
d2l.plt.gca().set_xlabel('Groups of experiments')
d2l.plt.gca().set_ylabel('Estimated probability')
d2l.plt.legend();
# + [markdown] origin_pos=20
# Each solid curve corresponds to one of the six values of the die and gives our estimated probability that the die turns up that value as assessed after each group of experiments.
# The dashed black line gives the true underlying probability.
# As we get more data by conducting more experiments,
# the $6$ solid curves converge towards the true probability.
#
# ### Axioms of Probability Theory
#
# When dealing with the rolls of a die,
# we call the set $\mathcal{S} = \{1, 2, 3, 4, 5, 6\}$ the *sample space* or *outcome space*, where each element is an *outcome*.
# An *event* is a set of outcomes from a given sample space.
# For instance, "seeing a $5$" ($\{5\}$) and "seeing an odd number" ($\{1, 3, 5\}$) are both valid events of rolling a die.
# Note that if the outcome of a random experiment is in event $\mathcal{A}$,
# then event $\mathcal{A}$ has occurred.
# That is to say, if $3$ dots faced up after rolling a die, since $3 \in \{1, 3, 5\}$,
# we can say that the event "seeing an odd number" has occurred.
#
# Formally, *probability* can be thought of as a function that maps a set to a real value.
# The probability of an event $\mathcal{A}$ in the given sample space $\mathcal{S}$,
# denoted as $P(\mathcal{A})$, satisfies the following properties:
#
# * For any event $\mathcal{A}$, its probability is never negative, i.e., $P(\mathcal{A}) \geq 0$;
# * Probability of the entire sample space is $1$, i.e., $P(\mathcal{S}) = 1$;
# * For any countable sequence of events $\mathcal{A}_1, \mathcal{A}_2, \ldots$ that are *mutually exclusive* ($\mathcal{A}_i \cap \mathcal{A}_j = \emptyset$ for all $i \neq j$), the probability that any happens is equal to the sum of their individual probabilities, i.e., $P(\bigcup_{i=1}^{\infty} \mathcal{A}_i) = \sum_{i=1}^{\infty} P(\mathcal{A}_i)$.
#
# These are also the axioms of probability theory, proposed by Kolmogorov in 1933.
# Thanks to this axiom system, we can avoid any philosophical dispute on randomness;
# instead, we can reason rigorously with a mathematical language.
# For instance, by letting event $\mathcal{A}_1$ be the entire sample space and $\mathcal{A}_i = \emptyset$ for all $i > 1$, we can prove that $P(\emptyset) = 0$, i.e., the probability of an impossible event is $0$.
#
#
# ### Random Variables
#
# In our random experiment of casting a die, we introduced the notion of a *random variable*. A random variable can be pretty much any quantity and is not deterministic. It could take one value among a set of possibilities in a random experiment.
# Consider a random variable $X$ whose value is in the sample space $\mathcal{S} = \{1, 2, 3, 4, 5, 6\}$ of rolling a die. We can denote the event "seeing a $5$" as $\{X = 5\}$ or $X = 5$, and its probability as $P(\{X = 5\})$ or $P(X = 5)$.
# By $P(X = a)$, we make a distinction between the random variable $X$ and the values (e.g., $a$) that $X$ can take.
# However, such pedantry results in a cumbersome notation.
# For a compact notation,
# on one hand, we can just denote $P(X)$ as the *distribution* over the random variable $X$:
# the distribution tells us the probability that $X$ takes any value.
# On the other hand,
# we can simply write $P(a)$ to denote the probability that a random variable takes the value $a$.
# Since an event in probability theory is a set of outcomes from the sample space,
# we can specify a range of values for a random variable to take.
# For example, $P(1 \leq X \leq 3)$ denotes the probability of the event $\{1 \leq X \leq 3\}$,
# which means $\{X = 1, 2, \text{or}, 3\}$. Equivalently, $P(1 \leq X \leq 3)$ represents the probability that the random variable $X$ can take a value from $\{1, 2, 3\}$.
#
# Note that there is a subtle difference between *discrete* random variables, like the sides of a die, and *continuous* ones, like the weight and the height of a person. There is little point in asking whether two people have exactly the same height. If we take precise enough measurements you will find that no two people on the planet have the exact same height. In fact, if we take a fine enough measurement, you will not have the same height when you wake up and when you go to sleep. So there is no purpose in asking about the probability
# that someone is 1.80139278291028719210196740527486202 meters tall. Given the world population of humans the probability is virtually 0. It makes more sense in this case to ask whether someone's height falls into a given interval, say between 1.79 and 1.81 meters. In these cases we quantify the likelihood that we see a value as a *density*. The height of exactly 1.80 meters has no probability, but nonzero density. In the interval between any two different heights we have nonzero probability.
# In the rest of this section, we consider probability in discrete space.
# For probability over continuous random variables, you may refer to :numref:`sec_random_variables`.
#
# ## Dealing with Multiple Random Variables
#
# Very often, we will want to consider more than one random variable at a time.
# For instance, we may want to model the relationship between diseases and symptoms. Given a disease and a symptom, say "flu" and "cough", either may or may not occur in a patient with some probability. While we hope that the probability of both would be close to zero, we may want to estimate these probabilities and their relationships to each other so that we may apply our inferences to effect better medical care.
#
# As a more complicated example, images contain millions of pixels, thus millions of random variables. And in many cases images will come with a
# label, identifying objects in the image. We can also think of the label as a
# random variable. We can even think of all the metadata as random variables
# such as location, time, aperture, focal length, ISO, focus distance, and camera type.
# All of these are random variables that occur jointly. When we deal with multiple random variables, there are several quantities of interest.
#
# ### Joint Probability
#
# The first is called the *joint probability* $P(A = a, B=b)$. Given any values $a$ and $b$, the joint probability lets us answer, what is the probability that $A=a$ and $B=b$ simultaneously?
# Note that for any values $a$ and $b$, $P(A=a, B=b) \leq P(A=a)$.
# This has to be the case, since for $A=a$ and $B=b$ to happen, $A=a$ has to happen *and* $B=b$ also has to happen (and vice versa). Thus, $A=a$ and $B=b$ cannot be more likely than $A=a$ or $B=b$ individually.
#
#
# ### Conditional Probability
#
# This brings us to an interesting ratio: $0 \leq \frac{P(A=a, B=b)}{P(A=a)} \leq 1$. We call this ratio a *conditional probability*
# and denote it by $P(B=b \mid A=a)$: it is the probability of $B=b$, provided that
# $A=a$ has occurred.
#
# ### Bayes' theorem
#
# Using the definition of conditional probabilities, we can derive one of the most useful and celebrated equations in statistics: *Bayes' theorem*.
# It goes as follows.
# By construction, we have the *multiplication rule* that $P(A, B) = P(B \mid A) P(A)$. By symmetry, this also holds for $P(A, B) = P(A \mid B) P(B)$. Assume that $P(B) > 0$. Solving for one of the conditional variables we get
#
# $$P(A \mid B) = \frac{P(B \mid A) P(A)}{P(B)}.$$
#
# Note that here we use the more compact notation where $P(A, B)$ is a *joint distribution* and $P(A \mid B)$ is a *conditional distribution*. Such distributions can be evaluated for particular values $A = a, B=b$.
#
# ### Marginalization
#
# Bayes' theorem is very useful if we want to infer one thing from the other, say cause and effect, but we only know the properties in the reverse direction, as we will see later in this section. One important operation that we need, to make this work, is *marginalization*.
# It is the operation of determining $P(B)$ from $P(A, B)$. We can see that the probability of $B$ amounts to accounting for all possible choices of $A$ and aggregating the joint probabilities over all of them:
#
# $$P(B) = \sum_{A} P(A, B),$$
#
# which is also known as the *sum rule*. The probability or distribution as a result of marginalization is called a *marginal probability* or a *marginal distribution*.
#
#
# ### Independence
#
# Another useful property to check for is *dependence* vs. *independence*.
# Two random variables $A$ and $B$ being independent
# means that the occurrence of one event of $A$
# does not reveal any information about the occurrence of an event of $B$.
# In this case $P(B \mid A) = P(B)$. Statisticians typically express this as $A \perp B$. From Bayes' theorem, it follows immediately that also $P(A \mid B) = P(A)$.
# In all the other cases we call $A$ and $B$ dependent. For instance, two successive rolls of a die are independent. In contrast, the position of a light switch and the brightness in the room are not (they are not perfectly deterministic, though, since we could always have a broken light bulb, power failure, or a broken switch).
#
# Since $P(A \mid B) = \frac{P(A, B)}{P(B)} = P(A)$ is equivalent to $P(A, B) = P(A)P(B)$, two random variables are independent if and only if their joint distribution is the product of their individual distributions.
# Likewise, two random variables $A$ and $B$ are *conditionally independent* given another random variable $C$
# if and only if $P(A, B \mid C) = P(A \mid C)P(B \mid C)$. This is expressed as $A \perp B \mid C$.
#
# ### Application
# :label:`subsec_probability_hiv_app`
#
# Let us put our skills to the test. Assume that a doctor administers an HIV test to a patient. This test is fairly accurate and it fails only with 1% probability if the patient is healthy but reporting him as diseased. Moreover,
# it never fails to detect HIV if the patient actually has it. We use $D_1$ to indicate the diagnosis ($1$ if positive and $0$ if negative) and $H$ to denote the HIV status ($1$ if positive and $0$ if negative).
# :numref:`conditional_prob_D1` lists such conditional probabilities.
#
# :Conditional probability of $P(D_1 \mid H)$.
#
# | Conditional probability | $H=1$ | $H=0$ |
# |---|---|---|
# |$P(D_1 = 1 \mid H)$| 1 | 0.01 |
# |$P(D_1 = 0 \mid H)$| 0 | 0.99 |
# :label:`conditional_prob_D1`
#
# Note that the column sums are all 1 (but the row sums are not), since the conditional probability needs to sum up to 1, just like the probability. Let us work out the probability of the patient having HIV if the test comes back positive, i.e., $P(H = 1 \mid D_1 = 1)$. Obviously this is going to depend on how common the disease is, since it affects the number of false alarms. Assume that the population is quite healthy, e.g., $P(H=1) = 0.0015$. To apply Bayes' theorem, we need to apply marginalization and the multiplication rule to determine
#
# $$\begin{aligned}
# &P(D_1 = 1) \\
# =& P(D_1=1, H=0) + P(D_1=1, H=1) \\
# =& P(D_1=1 \mid H=0) P(H=0) + P(D_1=1 \mid H=1) P(H=1) \\
# =& 0.011485.
# \end{aligned}
# $$
#
# Thus, we get
#
# $$\begin{aligned}
# &P(H = 1 \mid D_1 = 1)\\ =& \frac{P(D_1=1 \mid H=1) P(H=1)}{P(D_1=1)} \\ =& 0.1306 \end{aligned}.$$
#
# In other words, there is only a 13.06% chance that the patient
# actually has HIV, despite using a very accurate test.
# As we can see, probability can be counterintuitive.
#
# What should a patient do upon receiving such terrifying news? Likely, the patient
# would ask the physician to administer another test to get clarity. The second
# test has different characteristics and it is not as good as the first one, as shown in :numref:`conditional_prob_D2`.
#
#
# :Conditional probability of $P(D_2 \mid H)$.
#
# | Conditional probability | $H=1$ | $H=0$ |
# |---|---|---|
# |$P(D_2 = 1 \mid H)$| 0.98 | 0.03 |
# |$P(D_2 = 0 \mid H)$| 0.02 | 0.97 |
# :label:`conditional_prob_D2`
#
# Unfortunately, the second test comes back positive, too.
# Let us work out the requisite probabilities to invoke Bayes' theorem
# by assuming the conditional independence:
#
# $$\begin{aligned}
# &P(D_1 = 1, D_2 = 1 \mid H = 0) \\
# =& P(D_1 = 1 \mid H = 0) P(D_2 = 1 \mid H = 0) \\
# =& 0.0003,
# \end{aligned}
# $$
#
# $$\begin{aligned}
# &P(D_1 = 1, D_2 = 1 \mid H = 1) \\
# =& P(D_1 = 1 \mid H = 1) P(D_2 = 1 \mid H = 1) \\
# =& 0.98.
# \end{aligned}
# $$
#
# Now we can apply marginalization and the multiplication rule:
#
# $$\begin{aligned}
# &P(D_1 = 1, D_2 = 1) \\
# =& P(D_1 = 1, D_2 = 1, H = 0) + P(D_1 = 1, D_2 = 1, H = 1) \\
# =& P(D_1 = 1, D_2 = 1 \mid H = 0)P(H=0) + P(D_1 = 1, D_2 = 1 \mid H = 1)P(H=1)\\
# =& 0.00176955.
# \end{aligned}
# $$
#
# In the end, the probability of the patient having HIV given both positive tests is
#
# $$\begin{aligned}
# &P(H = 1 \mid D_1 = 1, D_2 = 1)\\
# =& \frac{P(D_1 = 1, D_2 = 1 \mid H=1) P(H=1)}{P(D_1 = 1, D_2 = 1)} \\
# =& 0.8307.
# \end{aligned}
# $$
#
# That is, the second test allowed us to gain much higher confidence that not all is well. Despite the second test being considerably less accurate than the first one, it still significantly improved our estimate.
#
#
#
# ## Expectation and Variance
#
# To summarize key characteristics of probability distributions,
# we need some measures.
# The *expectation* (or average) of the random variable $X$ is denoted as
#
# $$E[X] = \sum_{x} x P(X = x).$$
#
# When the input of a function $f(x)$ is a random variable drawn from the distribution $P$ with different values $x$,
# the expectation of $f(x)$ is computed as
#
# $$E_{x \sim P}[f(x)] = \sum_x f(x) P(x).$$
#
#
# In many cases we want to measure by how much the random variable $X$ deviates from its expectation. This can be quantified by the variance
#
# $$\mathrm{Var}[X] = E\left[(X - E[X])^2\right] =
# E[X^2] - E[X]^2.$$
#
# Its square root is called the *standard deviation*.
# The variance of a function of a random variable measures
# by how much the function deviates from the expectation of the function,
# as different values $x$ of the random variable are sampled from its distribution:
#
# $$\mathrm{Var}[f(x)] = E\left[\left(f(x) - E[f(x)]\right)^2\right].$$
#
#
# ## Summary
#
# * We can sample from probability distributions.
# * We can analyze multiple random variables using joint distribution, conditional distribution, Bayes' theorem, marginalization, and independence assumptions.
# * Expectation and variance offer useful measures to summarize key characteristics of probability distributions.
#
#
# ## Exercises
#
# 1. We conducted $m=500$ groups of experiments where each group draws $n=10$ samples. Vary $m$ and $n$. Observe and analyze the experimental results.
# 1. Given two events with probability $P(\mathcal{A})$ and $P(\mathcal{B})$, compute upper and lower bounds on $P(\mathcal{A} \cup \mathcal{B})$ and $P(\mathcal{A} \cap \mathcal{B})$. (Hint: display the situation using a [Venn Diagram](https://en.wikipedia.org/wiki/Venn_diagram).)
# 1. Assume that we have a sequence of random variables, say $A$, $B$, and $C$, where $B$ only depends on $A$, and $C$ only depends on $B$, can you simplify the joint probability $P(A, B, C)$? (Hint: this is a [Markov Chain](https://en.wikipedia.org/wiki/Markov_chain).)
# 1. In :numref:`subsec_probability_hiv_app`, the first test is more accurate. Why not run the first test twice rather than run both the first and second tests?
#
# + [markdown] origin_pos=21 tab=["mxnet"]
# [Discussions](https://discuss.d2l.ai/t/36)
#
| python/d2l-en/mxnet/chapter_preliminaries/probability.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from scipy import stats as sp
import sys, os
sys.path.append('../')
from lib import trace_analysis
from lib import plots_analysis
from lib import trace_statistics
from lib import trace_classification
import csv
from lib.functions import *
import warnings
warnings.filterwarnings('ignore')
# # Data Preparation
#
#
# Prepare the data to be used with kNN
# +
plots = set()
with open('traces/traces.csv') as csv_file:
csv_reader = csv.reader(csv_file, delimiter=',')
line_count = 0
for row in csv_reader:
if line_count == 0 or row[2].find('normal') >= 0:
line_count += 1
continue
else:
plots.add((row[1], row[2]))
plots = list(plots)
stats = None
win_25_stats = None
win_50_stats = None
win_100_stats = None
for row in plots:
experiment = row[1]
# Assign a label
if row[1].find('gh') >= 0:
label = 'GH'
else:
label = 'BH'
nodes, packets_node = trace_analysis.process_cooja2_traces(row[0], row[1])
# Update stats
if stats is None:
stats = trace_statistics.compute_labeled_statistics(nodes, packets_node, label, experiment)
win_25_stats = trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 25)
win_50_stats = trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 50)
win_100_stats = trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 100)
else:
stats = pd.concat([stats, trace_statistics.compute_labeled_statistics(nodes, packets_node, label, experiment)])
win_25_stats = pd.concat([win_25_stats, trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 25)])
win_50_stats = pd.concat([win_50_stats, trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 50)])
win_100_stats = pd.concat([win_100_stats, trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 100)])
stats = stats.reset_index(drop=True)
win_25_stats = win_25_stats.reset_index(drop=True)
win_50_stats = win_50_stats.reset_index(drop=True)
win_100_stats = win_100_stats.reset_index(drop=True)
# Create a dictionary containing all the statistics for each trace size
trace_stats = {200: stats, 25: win_25_stats, 50: win_50_stats, 100: win_100_stats}
stats.head(5)
# -
# Split topologies
# +
rnd1 = []
rnd2 = []
plot = []
for trace in plots:
if trace[0].find('rnd-1') >= 0:
rnd1.append(trace)
elif trace[0].find('rnd-2') >= 0:
rnd2.append(trace)
else:
plot.append(trace)
plot_stats = None
plot_win_25_stats = None
plot_win_50_stats = None
plot_win_100_stats = None
for row in plot:
experiment = row[1]
# Assign a label
if row[1].find('gh') >= 0:
label = 'GH'
else:
label = 'BH'
nodes, packets_node = trace_analysis.process_cooja2_traces(row[0], row[1])
# Update stats
if stats is None:
plot_stats = trace_statistics.compute_labeled_statistics(nodes, packets_node, label, experiment)
plot_win_25_stats = trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 25)
win_50_stats = trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 50)
plot_win_100_stats = trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 100)
else:
plot_stats = pd.concat([plot_stats, trace_statistics.compute_labeled_statistics(nodes, packets_node, label, experiment)])
plot_win_25_stats = pd.concat([plot_win_25_stats, trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 25)])
plot_win_50_stats = pd.concat([plot_win_50_stats, trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 50)])
plot_win_100_stats = pd.concat([plot_win_100_stats, trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 100)])
plot_stats = plot_stats.reset_index(drop=True)
plot_win_25_stats = plot_win_25_stats.reset_index(drop=True)
plot_win_50_stats = plot_win_50_stats.reset_index(drop=True)
plot_win_100_stats = plot_win_100_stats.reset_index(drop=True)
# Create a dictionary containing all the statistics for each trace size
plot_trace_stats = {200: plot_stats, 25: plot_win_25_stats, 50: plot_win_50_stats, 100: plot_win_100_stats}
plot_stats.head(5)
# +
rnd1_stats = None
rnd1_win_25_stats = None
rnd1_win_50_stats = None
rnd1_win_100_stats = None
for row in rnd1:
experiment = row[1]
# Assign a label
if row[1].find('gh') >= 0:
label = 'GH'
else:
label = 'BH'
nodes, packets_node = trace_analysis.process_cooja2_traces(row[0], row[1])
# Update stats
if stats is None:
rnd1_stats = trace_statistics.compute_labeled_statistics(nodes, packets_node, label, experiment)
rnd1_win_25_stats = trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 25)
rnd1_win_50_stats = trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 50)
rnd1_win_100_stats = trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 100)
else:
rnd1_stats = pd.concat([rnd1_stats, trace_statistics.compute_labeled_statistics(nodes, packets_node, label, experiment)])
rnd1_win_25_stats = pd.concat([rnd1_win_25_stats, trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 25)])
rnd1_win_50_stats = pd.concat([rnd1_win_50_stats, trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 50)])
rnd1_win_100_stats = pd.concat([rnd1_win_100_stats, trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 100)])
rnd1_stats = rnd1_stats.reset_index(drop=True)
rnd1_win_25_stats = rnd1_win_25_stats.reset_index(drop=True)
rnd1_win_50_stats = rnd1_win_50_stats.reset_index(drop=True)
rnd1_win_100_stats = rnd1_win_100_stats.reset_index(drop=True)
# Create a dictionary containing all the statistics for each trace size
rnd1_trace_stats = {200: rnd1_stats, 25: rnd1_win_25_stats, 50: rnd1_win_50_stats, 100: rnd1_win_100_stats}
rnd1_stats.head(5)
# +
rnd2_stats = None
rnd2_win_25_stats = None
rnd2_win_50_stats = None
rnd2_win_100_stats = None
for row in rnd2:
experiment = row[1]
# Assign a label
if row[1].find('gh') >= 0:
label = 'GH'
else:
label = 'BH'
nodes, packets_node = trace_analysis.process_cooja2_traces(row[0], row[1])
# Update stats
if stats is None:
rnd2_stats = trace_statistics.compute_labeled_statistics(nodes, packets_node, label, experiment)
rnd2_win_25_stats = trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 25)
rnd2_win_50_stats = trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 50)
rnd2_win_100_stats = trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 100)
else:
rnd2_stats = pd.concat([rnd2_stats, trace_statistics.compute_labeled_statistics(nodes, packets_node, label, experiment)])
rnd2_win_25_stats = pd.concat([rnd2_win_25_stats, trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 25)])
rnd2_win_50_stats = pd.concat([rnd2_win_50_stats, trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 50)])
rnd2_win_100_stats = pd.concat([rnd2_win_100_stats, trace_statistics.compute_window_labeled_statistics(nodes, packets_node, label, experiment, 100)])
rnd2_stats = rnd2_stats.reset_index(drop=True)
rnd2_win_25_stats = rnd2_win_25_stats.reset_index(drop=True)
rnd2_win_50_stats = rnd2_win_50_stats.reset_index(drop=True)
rnd2_win_100_stats = rnd2_win_100_stats.reset_index(drop=True)
# Create a dictionary containing all the statistics for each trace size
rnd2_trace_stats = {200: rnd2_stats, 25: rnd2_win_25_stats, 50: rnd2_win_50_stats, 100: rnd2_win_100_stats}
rnd2_stats.head(5)
# -
# Stats for single network
# +
net_stats = trace_statistics.compute_labeled_statistics_by_network(stats, 'loss', 9)
net_win_25_stats = trace_statistics.compute_window_labeled_statistics_by_network(win_25_stats, 'loss', 9, 25)
net_win_50_stats = trace_statistics.compute_window_labeled_statistics_by_network(win_50_stats, 'loss', 9, 50)
net_win_100_stats = trace_statistics.compute_window_labeled_statistics_by_network(win_100_stats, 'loss', 9, 100)
# Create a dictionary containing all the statistics for each trace size
network_stats = {200: net_stats, 25: net_win_25_stats, 50: net_win_50_stats, 100: net_win_100_stats}
net_stats.head(5)
# +
plot_net_stats = trace_statistics.compute_labeled_statistics_by_network(plot_stats, 'loss', 9)
plot_net_win_25_stats = trace_statistics.compute_window_labeled_statistics_by_network(plot_win_25_stats, 'loss', 9, 25)
plot_net_win_50_stats = trace_statistics.compute_window_labeled_statistics_by_network(plot_win_50_stats, 'loss', 9, 50)
plot_net_win_100_stats = trace_statistics.compute_window_labeled_statistics_by_network(plot_win_100_stats, 'loss', 9, 100)
# Create a dictionary containing all the statistics for each trace size
plot_network_stats = {200: plot_net_stats, 25: plot_net_win_25_stats, 50: plot_net_win_50_stats, 100: plot_net_win_100_stats}
plot_net_stats.head(5)
# +
rnd1_net_stats = trace_statistics.compute_labeled_statistics_by_network(rnd1_stats, 'loss', 9)
rnd1_net_win_25_stats = trace_statistics.compute_window_labeled_statistics_by_network(rnd1_win_25_stats, 'loss', 9, 25)
rnd1_net_win_50_stats = trace_statistics.compute_window_labeled_statistics_by_network(rnd1_win_50_stats, 'loss', 9, 50)
rnd1_net_win_100_stats = trace_statistics.compute_window_labeled_statistics_by_network(rnd1_win_100_stats, 'loss', 9, 100)
# Create a dictionary containing all the statistics for each trace size
rnd1_network_stats = {200: rnd1_net_stats, 25: rnd1_net_win_25_stats, 50: rnd1_net_win_50_stats, 100: rnd1_net_win_100_stats}
rnd1_net_stats.head(5)
# +
rnd2_net_stats = trace_statistics.compute_labeled_statistics_by_network(rnd2_stats, 'loss', 9)
rnd2_net_win_25_stats = trace_statistics.compute_window_labeled_statistics_by_network(rnd2_win_25_stats, 'loss', 9, 25)
rnd2_net_win_50_stats = trace_statistics.compute_window_labeled_statistics_by_network(rnd2_win_50_stats, 'loss', 9, 50)
rnd2_net_win_100_stats = trace_statistics.compute_window_labeled_statistics_by_network(rnd2_win_100_stats, 'loss', 9, 100)
# Create a dictionary containing all the statistics for each trace size
rnd2_network_stats = {200: rnd2_net_stats, 25: rnd2_net_win_25_stats, 50: rnd2_net_win_50_stats, 100: rnd2_net_win_100_stats}
rnd2_net_stats.head(5)
# -
# # Feature Selection
#
# Select the set of features and labels that we use to fit the algorithm
plots_analysis.random_forests_features_selection(trace_stats)
plots_analysis.random_forests_features_selection(plot_trace_stats)
plots_analysis.random_forests_features_selection(rnd1_trace_stats)
plots_analysis.random_forests_features_selection(rnd2_trace_stats)
# +
results = None # Results from each classification algorithm
cv_results = None # Cross validation results from each classification algorithm
net_results = None # Results from each classification algorithm
cv_net_results = None # Cross validation results from each classification algorithm
features_to_drop = ['node_id', 'experiment', 'label', 'loss', 'count', 'outliers']
net_features_to_drop = ['experiment', 'label']
# +
plot_results = None # Results from each classification algorithm
plot_cv_results = None # Cross validation results from each classification algorithm
plot_net_results = None # Results from each classification algorithm
plot_cv_net_results = None # Cross validation results from each classification algorithm
rnd1_results = None # Results from each classification algorithm
rnd1_cv_results = None # Cross validation results from each classification algorithm
rnd1_net_results = None # Results from each classification algorithm
rnd1_cv_net_results = None # Cross validation results from each classification algorithm
rnd2_results = None # Results from each classification algorithm
rnd2_cv_results = None # Cross validation results from each classification algorithm
rnd2_net_results = None # Results from each classification algorithm
rnd2_cv_net_results = None # Cross validation results from each classification algorithm
# -
# # Random Forests Classifier
#
# Let us experiment with random forests. First, let us select most relevant features.
results = pd.concat([results,
trace_classification.random_forest_classification(trace_stats, features_to_drop)
])
plot_results = pd.concat([plot_results,
trace_classification.random_forest_classification(plot_trace_stats, features_to_drop)
])
rnd1_results = pd.concat([rnd1_results,
trace_classification.random_forest_classification(rnd1_trace_stats, features_to_drop)
])
rnd2_results = pd.concat([rnd2_results,
trace_classification.random_forest_classification(rnd2_trace_stats, features_to_drop)
])
# We want to compute average accuracy for each trace size (i.e. for each window size from 200 - the entire window - to 100, 50 and 25)
cv_results = pd.concat([cv_results,
trace_classification.random_forest_cross_validation(trace_stats, features_to_drop)
])
# +
plot_cv_results = pd.concat([plot_cv_results,
trace_classification.random_forest_cross_validation(plot_trace_stats, features_to_drop)
])
rnd1_cv_results = pd.concat([rnd1_cv_results,
trace_classification.random_forest_cross_validation(rnd1_trace_stats, features_to_drop)
])
rnd2_cv_results = pd.concat([rnd2_cv_results,
trace_classification.random_forest_cross_validation(rnd2_trace_stats, features_to_drop)
])
# -
# Let's do the same for the whole network
net_results = pd.concat([net_results,
trace_classification.random_forest_classification(network_stats, net_features_to_drop)
])
# +
plot_net_results = pd.concat([plot_net_results,
trace_classification.random_forest_classification(plot_network_stats, net_features_to_drop)
])
rnd1_net_results = pd.concat([rnd1_net_results,
trace_classification.random_forest_classification(rnd1_network_stats, net_features_to_drop)
])
rnd2_net_results = pd.concat([rnd2_net_results,
trace_classification.random_forest_classification(rnd2_network_stats, net_features_to_drop)
])
# -
# Cross validation
cv_net_results = pd.concat([cv_net_results,
trace_classification.random_forest_cross_validation(network_stats, net_features_to_drop, cross_val=3)
])
# +
plot_cv_net_results = pd.concat([plot_cv_net_results,
trace_classification.random_forest_cross_validation(plot_network_stats, net_features_to_drop, cross_val=3)
])
rnd1_cv_net_results = pd.concat([rnd1_cv_net_results,
trace_classification.random_forest_cross_validation(rnd1_network_stats, net_features_to_drop, cross_val=3)
])
rnd2_cv_net_results = pd.concat([rnd2_cv_net_results,
trace_classification.random_forest_cross_validation(rnd2_network_stats, net_features_to_drop, cross_val=3)
])
# -
# # K-Nearest Neighbor (KNN) Classification
#
# Let us first observe the accuracies for different values of k
plots_analysis.knn_test_number_of_neighbors(trace_stats, 30)
plots_analysis.knn_test_number_of_neighbors(plot_trace_stats, 30)
plots_analysis.knn_test_number_of_neighbors(rnd1_trace_stats, 30)
plots_analysis.knn_test_number_of_neighbors(rnd2_trace_stats, 20)
# Let's build KNN classifier
results = pd.concat([results,
trace_classification.k_nearest_neighbor_classification(trace_stats, features_to_drop, n_neighbors=30)
])
# +
plot_results = pd.concat([plot_results,
trace_classification.k_nearest_neighbor_classification(plot_trace_stats, features_to_drop, n_neighbors=8)
])
rnd1_results = pd.concat([rnd1_results,
trace_classification.k_nearest_neighbor_classification(rnd1_trace_stats, features_to_drop, n_neighbors=7)
])
rnd2_results = pd.concat([rnd2_results,
trace_classification.k_nearest_neighbor_classification(rnd2_trace_stats, features_to_drop, n_neighbors=7)
])
# -
# We want to compute average accuracy for each trace size (i.e. for each window size from 200 - the entire window - to 100, 50 and 25)
cv_results = pd.concat([cv_results,
trace_classification.k_nearest_neighbor_cross_validation(trace_stats, features_to_drop, n_neighbors=30)
])
# +
plot_cv_results = pd.concat([plot_cv_results,
trace_classification.k_nearest_neighbor_cross_validation(plot_trace_stats, features_to_drop, n_neighbors=8)
])
rnd1_cv_results = pd.concat([rnd1_cv_results,
trace_classification.k_nearest_neighbor_cross_validation(rnd1_trace_stats, features_to_drop, n_neighbors=7)
])
rnd2_cv_results = pd.concat([rnd2_cv_results,
trace_classification.k_nearest_neighbor_cross_validation(rnd2_trace_stats, features_to_drop, n_neighbors=7)
])
# -
# Let's do the same for the whole network
net_results = pd.concat([net_results,
trace_classification.k_nearest_neighbor_classification(network_stats, net_features_to_drop)
])
# +
plot_net_results = pd.concat([plot_net_results,
trace_classification.k_nearest_neighbor_classification(plot_network_stats, net_features_to_drop)
])
rnd1_net_results = pd.concat([rnd1_net_results,
trace_classification.k_nearest_neighbor_classification(rnd1_network_stats, net_features_to_drop)
])
rnd2_net_results = pd.concat([rnd2_net_results,
trace_classification.k_nearest_neighbor_classification(rnd2_network_stats, net_features_to_drop, n_neighbors = 2)
])
# -
# Cross validation
cv_net_results = pd.concat([cv_net_results,
trace_classification.k_nearest_neighbor_cross_validation(network_stats, net_features_to_drop, cross_val=3)
])
# +
plot_cv_net_results = pd.concat([plot_cv_net_results,
trace_classification.k_nearest_neighbor_cross_validation(plot_network_stats, net_features_to_drop, cross_val=3)
])
rnd1_cv_net_results = pd.concat([rnd1_cv_net_results,
trace_classification.k_nearest_neighbor_cross_validation(rnd1_network_stats, net_features_to_drop, cross_val=3)
])
rnd2_cv_net_results = pd.concat([rnd2_cv_net_results,
trace_classification.k_nearest_neighbor_cross_validation(rnd2_network_stats, net_features_to_drop, cross_val=3, n_neighbors = 2)
])
# -
# # Support Vector Machines (SVM) Classification
#
# Let us experiment with another classifier
results = pd.concat([results,
trace_classification.support_vector_machines_classification(trace_stats, features_to_drop, kernel='rbf')
])
# +
plot_results = pd.concat([plot_results,
trace_classification.support_vector_machines_classification(plot_trace_stats, features_to_drop, kernel='rbf')
])
rnd1_results = pd.concat([rnd1_results,
trace_classification.support_vector_machines_classification(rnd1_trace_stats, features_to_drop, kernel='rbf')
])
rnd2_results = pd.concat([rnd2_results,
trace_classification.support_vector_machines_classification(rnd2_trace_stats, features_to_drop, kernel='rbf')
])
# -
# We want to compute average accuracy for each trace size (i.e. for each window size from 200 - the entire window - to 100, 50 and 25)
cv_results = pd.concat([cv_results,
trace_classification.support_vector_machines_cross_validation(trace_stats, features_to_drop, kernel='rbf')
])
# +
plot_cv_results = pd.concat([plot_cv_results,
trace_classification.support_vector_machines_cross_validation(plot_trace_stats, features_to_drop, kernel='rbf')
])
rnd1_cv_results = pd.concat([rnd1_cv_results,
trace_classification.support_vector_machines_cross_validation(rnd1_trace_stats, features_to_drop, kernel='rbf')
])
rnd2_cv_results = pd.concat([rnd2_cv_results,
trace_classification.support_vector_machines_cross_validation(rnd2_trace_stats, features_to_drop, kernel='rbf')
])
# -
# Let's do the same for the whole network
net_results = pd.concat([net_results,
trace_classification.support_vector_machines_classification(network_stats, net_features_to_drop, kernel='rbf')
])
# +
plot_net_results = pd.concat([plot_net_results,
trace_classification.support_vector_machines_classification(plot_network_stats, net_features_to_drop, kernel='rbf')
])
rnd1_net_results = pd.concat([rnd1_net_results,
trace_classification.support_vector_machines_classification(rnd1_network_stats, net_features_to_drop, kernel='rbf')
])
rnd2_net_results = pd.concat([rnd2_net_results,
trace_classification.support_vector_machines_classification(rnd2_network_stats, net_features_to_drop, kernel='rbf')
])
# -
# Cross validation
cv_net_results = pd.concat([cv_net_results,
trace_classification.support_vector_machines_cross_validation(network_stats, net_features_to_drop, cross_val=3)
])
# +
plot_cv_net_results = pd.concat([plot_cv_net_results,
trace_classification.support_vector_machines_cross_validation(plot_network_stats, net_features_to_drop, cross_val=3)
])
rnd1_cv_net_results = pd.concat([rnd1_cv_net_results,
trace_classification.support_vector_machines_cross_validation(rnd1_network_stats, net_features_to_drop, cross_val=3)
])
'''rnd2_cv_net_results = pd.concat([rnd2_cv_net_results,
trace_classification.support_vector_machines_cross_validation(rnd2_network_stats, net_features_to_drop, cross_val=3)
])'''
# -
# # One Vs The Rest Classifier
#
# SVM performs really well, but it is slow. Now we try to implement it in a most efficient way. The strategy consists in fitting one classifier per class. For each classifier, the class is fitted against all the other classes
results = pd.concat([results,
trace_classification.ensalble_svm_classification(trace_stats, features_to_drop, n_estimators=10)
])
# +
plot_results = pd.concat([plot_results,
trace_classification.ensalble_svm_classification(plot_trace_stats, features_to_drop, n_estimators=15)
])
rnd1_results = pd.concat([rnd1_results,
trace_classification.ensalble_svm_classification(rnd1_trace_stats, features_to_drop, n_estimators=15)
])
rnd2_results = pd.concat([rnd2_results,
trace_classification.ensalble_svm_classification(rnd2_trace_stats, features_to_drop, n_estimators=15)
])
# -
# Cross validate
cv_results = pd.concat([cv_results,
trace_classification.ensalble_svm_cross_validation(trace_stats, features_to_drop, n_estimators=15)
])
# +
plot_cv_results = pd.concat([plot_cv_results,
trace_classification.ensalble_svm_cross_validation(plot_trace_stats, features_to_drop, n_estimators=15)
])
rnd1_cv_results = pd.concat([rnd1_cv_results,
trace_classification.ensalble_svm_cross_validation(rnd1_trace_stats, features_to_drop, n_estimators=15)
])
rnd2_cv_results = pd.concat([rnd2_cv_results,
trace_classification.ensalble_svm_cross_validation(rnd2_trace_stats, features_to_drop, n_estimators=15)
])
# -
# Let's do the same for the whole network
net_results = pd.concat([net_results,
trace_classification.ensalble_svm_classification(network_stats, net_features_to_drop)
])
# +
plot_net_results = pd.concat([plot_net_results,
trace_classification.ensalble_svm_classification(plot_network_stats, net_features_to_drop)
])
'''rnd1_net_results = pd.concat([rnd1_net_results,
trace_classification.ensalble_svm_classification(rnd1_network_stats, net_features_to_drop)
])
rnd2_net_results = pd.concat([rnd2_net_results,
trace_classification.ensalble_svm_classification(rnd2_network_stats, net_features_to_drop)
])'''
# -
# Cross validation
cv_net_results = pd.concat([cv_net_results,
trace_classification.ensalble_svm_cross_validation(network_stats, net_features_to_drop, cross_val=3)
])
# +
plot_cv_net_results = pd.concat([plot_cv_net_results,
trace_classification.ensalble_svm_cross_validation(plot_network_stats, net_features_to_drop)
])
'''rnd1_cv_net_results = pd.concat([rnd1_cv_net_results,
trace_classification.ensalble_svm_cross_validation(rnd1_network_stats, net_features_to_drop)
])'''
'''rnd2_cv_net_results = pd.concat([rnd2_cv_net_results,
trace_classification.ensalble_svm_cross_validation(rnd2_network_stats, net_features_to_drop)
])'''
# -
# # Per Node Results
# Results from each model
results.reset_index(drop=True)
plot_results.reset_index(drop=True)
rnd1_results.reset_index(drop=True)
rnd2_results.reset_index(drop=True)
# Here we plot the average results for each model computed with cross validation
cv_results.reset_index(drop=True)
plot_cv_results.reset_index(drop=True)
rnd1_cv_results.reset_index(drop=True)
rnd2_cv_results.reset_index(drop=True)
# # Network Results
# Here we print the results from each model on the whole network
net_results.reset_index(drop=True)
plot_net_results.reset_index(drop=True)
rnd1_net_results.reset_index(drop=True)
rnd2_net_results.reset_index(drop=True)
# Here we plot the average results for each model computed with cross validation
cv_net_results.reset_index(drop=True)
plot_cv_net_results.reset_index(drop=True)
rnd1_cv_net_results.reset_index(drop=True)
rnd2_cv_net_results.reset_index(drop=True)
| module/data/cooja3-9nodes/BH-GH Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/African-Quant/WQU_MScFE_Capstone_Grp9/blob/master/Deployment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="OlsqE2t_PwOf" cellView="form" colab={"base_uri": "https://localhost:8080/"} outputId="eb751622-c0d1-4732-9dbb-b00c1075c6dc"
#@title Installation
# !pip install git+https://github.com/yhilpisch/tpqoa.git --upgrade --quiet
# ! pip install backtrader[plotting] --quiet
# + id="5xuLLhALeirJ" cellView="form"
#@title Imports { vertical-output: true }
import tpqoa
import numpy as np
import pandas as pd
# %matplotlib inline
from pylab import mpl, plt
plt.style.use('seaborn')
mpl.rcParams['savefig.dpi'] = 300
mpl.rcParams['font.family'] = 'serif'
from datetime import date, timedelta
# + id="4fa2AsNierg8" cellView="form"
#@title Oanda API
path = '/content/drive/MyDrive/Oanda_Algo/pyalgo.cfg'
api = tpqoa.tpqoa('/content/drive/MyDrive/Oanda_Algo/pyalgo.cfg')
# + id="qxFTnK32QZ94" cellView="form"
#@title Currency/Currency Pairs
inst = ['BCO_USD','EUR_USD', 'USD_JPY', 'GBP_USD', 'USD_CHF',
'AUD_USD', 'USD_CAD', 'NZD_USD', 'EUR_GBP', 'EUR_JPY',
'GBP_JPY', 'CHF_JPY', 'GBP_CHF', 'EUR_AUD', 'EUR_CAD',
'AUD_CAD', 'AUD_JPY', 'CAD_JPY', 'NZD_JPY', 'GBP_CAD',
'GBP_NZD', 'GBP_AUD', 'AUD_NZD', 'AUD_CHF', 'EUR_NZD',
'NZD_CHF', 'CAD_CHF', 'NZD_CAD', 'DE30_EUR', 'US30_USD',
'EUR_CHF', 'XAU_USD', 'XAG_USD']
currency = ['EUR', 'GBP', 'AUD', 'NZD', 'USD', 'CAD', 'CHF', 'JPY']
# + id="gIhN00HmQiXZ" cellView="form"
#@title get_data(instr, gran = 'D', td=1000)
def get_data(instr, gran = 'D', td=1000):
start = f"{date.today() - timedelta(td) }"
end = f"{date.today() - timedelta(1)}"
granularity = gran
price = 'M'
data = api.get_history(instr, start, end, granularity, price)
data.drop(['complete'], axis=1, inplace=True)
data.reset_index(inplace=True)
data.rename(columns = {'time':'Date','o':'Open','c': 'Close', 'h':'High', 'l': 'Low'}, inplace = True)
data.set_index('Date', inplace=True)
return data
# + colab={"base_uri": "https://localhost:8080/"} id="6nQnE_MhtFap" outputId="4b4334ea-c90a-4cf0-b231-cc427cacaceb"
d = {a:b for a, b in enumerate(inst)}
print(d)
# + colab={"base_uri": "https://localhost:8080/", "height": 235} id="u5-S7JJ7tb1N" outputId="f6a51e4e-d54e-4529-b613-2b0c75852bfe"
us30 = get_data(inst[29])
us30.tail()
# + [markdown] id="DKg1IH0BWDv7"
# Geting the indicator function
# + colab={"base_uri": "https://localhost:8080/"} id="KLbs5pu9-Tfz" outputId="618d40a2-b633-49d4-b57d-9eb8dd62abfa"
# %%writefile deploy.py
import os
import re
import tpqoa
import random
import numpy as np
from sklearn.metrics import accuracy_score, confusion_matrix, classification_report
from sklearn.model_selection import RandomizedSearchCV, GridSearchCV
import pandas as pd
from datetime import date, timedelta
import warnings
warnings.filterwarnings('ignore')
from sklearn.ensemble import RandomForestClassifier
from lightgbm import LGBMClassifier
from xgboost import XGBClassifier
pairs = ['EUR_USD', 'USD_JPY', 'GBP_USD', 'USD_CHF',
'AUD_USD', 'USD_CAD', 'NZD_USD', 'EUR_GBP', 'EUR_JPY',
'GBP_JPY', 'CHF_JPY', 'GBP_CHF', 'EUR_AUD', 'EUR_CAD',
'AUD_CAD', 'AUD_JPY', 'CAD_JPY', 'NZD_JPY', 'GBP_CAD',
'GBP_NZD', 'GBP_AUD', 'AUD_NZD', 'AUD_CHF', 'EUR_NZD',
'NZD_CHF', 'CAD_CHF', 'NZD_CAD', 'EUR_CHF']
def get_data(instr, gran = 'D', td=1000):
start = f"{date.today() - timedelta(td) }"
end = f"{date.today() - timedelta(1)}"
granularity = gran
price = 'M'
data = api.get_history(instr, start, end, granularity, price)
data.drop(['complete'], axis=1, inplace=True)
data.reset_index(inplace=True)
data.rename(columns = {'time':'Date','o':'Open','c': 'Close', 'h':'High', 'l': 'Low'}, inplace = True)
data.set_index('Date', inplace=True)
return data
# ATR
def eATR(df1,n=14):
"""This calculates the exponential Average True Range of of a dataframe of the open,
high, low, and close data of an instrument"""
df = df1[['Open', 'High', 'Low', 'Close']].copy()
# True Range
df['TR'] = 0
for i in range(len(df)):
try:
df.iloc[i, 4] = max(df.iat[i,1] - df.iat[i,2],
abs(df.iat[i,1] - df.iat[i-1,3]),
abs(df.iat[i,2] - df.iat[i-1,3]))
except ValueError:
pass
# eATR
df['eATR'] = df['TR'].ewm(span=n, adjust=False).mean()
return df['eATR']
def ssl(df1):
"""This function adds the ssl indicator as features to a dataframe
"""
df = df1.copy()
df['smaHigh'] = df['High'].rolling(window=10).mean()
df['smaLow'] = df['Low'].rolling(window=10).mean()
df['hlv'] = 0
df['hlv'] = np.where(df['Close'] > df['smaHigh'],1,np.where(df['Close'] < df['smaLow'],-1,df['hlv'].shift(1)))
df['sslDown'] = np.where(df['hlv'] < 0, df['smaHigh'], df['smaLow'])
df['sslUp'] = np.where(df['hlv'] < 0, df['smaLow'], df['smaHigh'])
df['sslPosition'] = np.where(df['Close'] > df['sslUp'], 1,
np.where(df['Close'] < df['sslDown'], -1, 0))
return df[['sslDown', 'sslUp', 'sslPosition']]
# <NAME>
def WAE(df1):
"""This function creates adds the indicator Waddah Attar features to a dataframe
"""
df = df1.copy()
# EMA
long_ema = df.loc[:,'Close'].ewm(span=40, adjust=False).mean()
short_ema = df.loc[:,'Close'].ewm(span=20, adjust=False).mean()
# MACD
MACD = short_ema - long_ema
# bBands
sma20 = df.loc[:,'Close'].rolling(window=20).mean() # 20 SMA
stddev = df.loc[:,'Close'].rolling(window=20).std() # 20 STDdev
lower_band = sma20 - (2 * stddev)
upper_band = sma20 + (2 * stddev)
#<NAME>
t1 = (MACD - MACD.shift(1))* 150
#t2 = MACD.shift(2) - MACD.shift(3)
df['e1'] = upper_band - lower_band
df['e2'] = -1 *df['e1']
#e2 = upper_band.shift(1) - lower_band.shift(1)
df['trendUp'] = np.where(t1 > 0, t1, 0)
df['trendDown'] = np.where(t1 < 0, t1, 0)
df['waePosition'] = np.where(df['trendUp'] > 0, 1,
np.where(df['trendDown'] < 0, -1, 0))
return df[['e1','e2','trendUp', 'trendDown', 'waePosition']]
def lag_feat(data1):
"""This function adds lag returns as features to a dataframe
"""
data = data1.copy()
lags = 8
cols = []
for lag in range(1, lags + 1):
col = f'lag_{lag}'
data[col] = data['ret'].shift(lag)
cols.append(col)
return data[cols]
def datepart_feat(df0, colname = 'Date'):
"""This function adds some common pandas date parts like 'year',
'month' etc as features to a dataframe
"""
df = df0.copy()
df.reset_index(inplace=True)
df1 = df.loc[:,colname]
nu_feats = ['Day', 'Dayofweek', 'Dayofyear']
targ_pre = re.sub('[Dd]ate$', '', colname)
for n in nu_feats:
df[targ_pre+n] = getattr(df1.dt,n.lower())
df[targ_pre+'week'] = df1.dt.isocalendar().week
df['week'] = np.int64(df['week'])
df[targ_pre+'Elapsed'] = df1.astype(np.int64) // 10**9
nu_feats.extend(['week', 'Elapsed'])
df.set_index(colname, inplace=True)
return df[nu_feats]
def gen_feat(pair):
df0 = get_data(pair)
df0['ret'] = df0['Close'].pct_change()
df0['dir'] = np.sign(df0['ret'])
eATR_ = eATR(df0).shift(1)
wae = WAE(df0).shift(1)
ssl1 = ssl(df0).shift(1)
datepart = datepart_feat(df0)
lags = lag_feat(df0)
return pd.concat([df0, eATR_, wae, ssl1, datepart, lags], axis=1).dropna()
# random forest
def rfc(xs, y, n_estimators=40, max_samples=100,
max_features=0.5, min_samples_leaf=5, **kwargs):
return RandomForestClassifier(n_jobs=-1, n_estimators=n_estimators,
max_samples=max_samples, max_features=max_features,
min_samples_leaf=min_samples_leaf, oob_score=True).fit(xs, y)
def rfc_deploy():
"""This function trains a Random Forest classifier and outputs the
out-of-sample performance from the validation and test sets
"""
df = pd.DataFrame()
for pair in pairs:
# retrieving the data and preparing the features
dataset = gen_feat(pair)
dataset.drop(['Open', 'High', 'Low', 'Close', 'volume'], axis=1, inplace=True)
# selecting the features to train on
cols = list(dataset.columns)
feats = cols[2:]
#splitting into training, validation and test sets
df_train = dataset.iloc[:-100,:]
train = df_train.copy()
df_test = dataset.iloc[-100:,:]
test = df_test.copy()
train_f = train.iloc[:-100,:]
valid = train.iloc[-100:,:]
#training the algorithm
m = rfc(train_f[feats], train_f['dir'])
# test sets
test_pred = m.predict(test[feats])
test_proba = m.predict_proba(test[feats])
df1 = pd.DataFrame(test_pred,columns=['prediction'], index=test.index)
proba_short = []
proba_long = []
for x in range(len(test_proba)):
proba_short.append(test_proba[x][0])
proba_long.append(test_proba[x][-1])
proba = {'proba_short': proba_short,
'proba_long': proba_long}
df2 = pd.DataFrame(proba, index=test.index)
df1['probability'] = np.where(df1['prediction'] == 1, df2['proba_long'],
np.where(df1['prediction'] == -1, df2['proba_short'], 0))
df1['signal'] = np.where((df1['probability'] >= .7) & (df1['prediction'] == 1), 'Go Long',
np.where((df1['probability'] >= 0.7) & (df1['prediction'] == -1), 'Go Short', 'Stand Aside'))
df1.reset_index(inplace=True)
df1['pair'] = pair
df1.set_index('pair', inplace=True)
entry_sig = df1[['probability', 'signal']].iloc[-1:]
# Merge
df = pd.concat([df, entry_sig], axis=0)
#output
return df
# Light GBM
def lgb(xs, y, learning_rate=0.15, boosting_type='gbdt',
objective='binary', n_estimators=50,
metric=['auc', 'binary_logloss'],
num_leaves=100, max_depth= 1,
**kwargs):
return LGBMClassifier().fit(xs, y)
def lgb_deploy():
"""This function trains a Light Gradient Boosting Method and outputs the
out-of-sample performance from the validation and test sets
"""
df = pd.DataFrame()
for pair in pairs:
# retrieving the data and preparing the features
dataset = gen_feat(pair)
dataset.drop(['Open', 'High', 'Low', 'Close', 'volume'], axis=1, inplace=True)
# selecting the features to train on
cols = list(dataset.columns)
feats = cols[2:]
#splitting into training, validation and test sets
df_train = dataset.iloc[:-100,:]
train = df_train.copy()
df_test = dataset.iloc[-100:,:]
test = df_test.copy()
train_f = train.iloc[:-100,:]
valid = train.iloc[-100:,:]
#training the algorithm
m = lgb(train_f[feats], train_f['dir']);
# test sets
test_pred = m.predict(test[feats])
test_proba = m.predict_proba(test[feats])
df1 = pd.DataFrame(test_pred,columns=['prediction'], index=test.index)
proba_short = []
proba_long = []
for x in range(len(test_proba)):
proba_short.append(test_proba[x][0])
proba_long.append(test_proba[x][-1])
proba = {'proba_short': proba_short,
'proba_long': proba_long}
df2 = pd.DataFrame(proba, index=test.index)
df1['probability'] = np.where(df1['prediction'] == 1, df2['proba_long'],
np.where(df1['prediction'] == -1, df2['proba_short'], 0))
df1['signal'] = np.where((df1['probability'] >= .7) & (df1['prediction'] == 1), 'Go Long',
np.where((df1['probability'] >= 0.7) & (df1['prediction'] == -1), 'Go Short', 'Stand Aside'))
df1.reset_index(inplace=True)
df1['pair'] = pair
df1.set_index('pair', inplace=True)
entry_sig = df1[['probability', 'signal']].iloc[-1:]
# Merge
df = pd.concat([df, entry_sig], axis=0)
#output
return df
# eXtreme Gradient Boosting
def xgb(xs, y):
return XGBClassifier().fit(xs, y)
def xgb_deploy():
"""This function trains a eXtreme Gradient Boosting Method and outputs the
out-of-sample performance from the validation and test sets
"""
df = pd.DataFrame()
for pair in pairs:
# retrieving the data and preparing the features
dataset = gen_feat(pair)
dataset.drop(['Open', 'High', 'Low', 'Close', 'volume'], axis=1, inplace=True)
# selecting the features to train on
cols = list(dataset.columns)
feats = cols[2:]
#splitting into training, validation and test sets
df_train = dataset.iloc[:-100,:]
train = df_train.copy()
df_test = dataset.iloc[-100:,:]
test = df_test.copy()
train_f = train.iloc[:-100,:]
valid = train.iloc[-100:,:]
#training the algorithm
m = xgb(train_f[feats], train_f['dir']);
# test sets
test_pred = m.predict(test[feats])
test_proba = m.predict_proba(test[feats])
df1 = pd.DataFrame(test_pred,columns=['prediction'], index=test.index)
proba_short = []
proba_long = []
for x in range(len(test_proba)):
proba_short.append(test_proba[x][0])
proba_long.append(test_proba[x][-1])
proba = {'proba_short': proba_short,
'proba_long': proba_long}
df2 = pd.DataFrame(proba, index=test.index)
df1['probability'] = np.where(df1['prediction'] == 1, df2['proba_long'],
np.where(df1['prediction'] == -1, df2['proba_short'], 0))
df1['signal'] = np.where((df1['probability'] >= .7) & (df1['prediction'] == 1), 'Go Long',
np.where((df1['probability'] >= 0.7) & (df1['prediction'] == -1), 'Go Short', 'Stand Aside'))
df1.reset_index(inplace=True)
df1['pair'] = pair
df1.set_index('pair', inplace=True)
entry_sig = df1[['probability', 'signal']].iloc[-1:]
# Merge
df = pd.concat([df, entry_sig], axis=0)
#output
return df
# + [markdown] id="Q9vcqlleo6zM"
# ### Deploying algorithm using Random Forest
# + colab={"base_uri": "https://localhost:8080/", "height": 906} id="QmumJpWbjLVb" outputId="b11e08ec-b3b2-4749-aa5a-7798bc90de2b"
rfc_deploy()
# + [markdown] id="2f0HRnZNpCyO"
# ### Deploying using Light Grandient Boosting
# + colab={"base_uri": "https://localhost:8080/", "height": 906} id="PyoYtWuDoszu" outputId="5c697c6a-214a-4632-d60c-82136cda5fbe"
lgb_deploy()
# + [markdown] id="7Tq_hBCwpMOl"
# ### Deploying using eXtreme Gradient Boosting method
# + colab={"base_uri": "https://localhost:8080/", "height": 906} id="BlH_y-QnovV9" outputId="2fb5788a-cbe1-49c9-b366-516cb851da0a"
xgb_deploy()
# + id="60KLu3-foyMX"
| Deployment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Cobwebbing
#
# <NAME>
#
# Created 4/20/21
#
# Based on a <a href="https://github.com/adam-rumpf/mathematica-class-demonstrations#cobwebbing" target="_blank">Mathematica class demonstration</a>.
#
# This is a standalone widget for playing around with cobweb diagrams for various dynamical systems. See the full notebook [here](./cobwebbing.ipynb).
#
# [Main Project Page](.././index.ipynb)
# + jupyter={"source_hidden": true} tags=[]
# %matplotlib widget
import ipywidgets as widgets
import matplotlib.pyplot as plt
import numpy as np
# Define parameters
COBWEB_MAX = 20 # number of cobweb iterations to generate
# Define functions
def lmap(x, lim=1.0, r=1.0, h=0.0):
"""Discrete logistic map with absolute harvesting.
Positional arguments:
x - input value
Keyword arguments:
lim (1.0) - population limit
r (1.0) - intrinsic growth rate
"""
return x + r*x*(1 - (x/lim)) - h
def tmap(x, lb=0.5, ub=1.0, r=1.0):
"""Discrete population growth with upper and lower stable population bounds.
Positional arguments:
x - input value
Keyword arguments:
lb (0.5) - lower population limit
ub (1.0) - upper population limit
r (1.0) - intrinsic growth rate
"""
return x + r*x*(x-lb)*(ub-x)
def mmap(x, r=0.0):
"""Discrete map meant to have an adjustable slope at one of its equlibria.
Positional arguments:
x - input value
Keyword arguments:
r (0.0) - slope at intermediate equilibrium point (<= 1.5)
"""
a = 4 - 4*r
return (a*x*x - 1.5*a*x + (1.0 + 0.5*a))*x
def cobweb_update(x0, cwx, cwy, lim=1.0, r=1.0, mode=0):
"""Updates the global cobweb lists. Lists are edited in-place.
Positional arguments:
x0 - initial population value
cwx - reference to a list of cobweb plot x-values
cwy - reference to a list of cobweb plot y-values
Keyword arguments:
lim (1.0) - population limit
r (1.0) - intrinsic growth rate
mode (0) - 0 for logistic with harvesting, 1 for bounded population, 2 for slope map
"""
# Generate cobweb coordinates
cwx[0] = x0
cwy[0] = 0.0
for i in range(0, 2*COBWEB_MAX, 2):
cwx[i+1] = cwx[i]
if mode == 0:
cwy[i+1] = max(lmap(cwx[i], r=r), 0.0)
elif mode == 1:
cwy[i+1] = max(tmap(cwx[i], r=r), 0.0)
elif mode == 2:
cwy[i+1] = max(mmap(cwx[i], r=r), 0.0)
cwx[i+2] = cwy[i+1]
cwy[i+2] = cwy[i+1]
cwx[-1] = cwx[-2]
if mode == 0:
cwy[-1] = max(lmap(cwx[-1], r=r), 0.0)
elif mode == 1:
cwy[-1] = max(tmap(cwx[-1], r=r), 0.0)
elif mode == 2:
cwy[-1] = max(mmap(cwx[-1], r=r), 0.0)
# Generate x- and n-values
x = np.linspace(0, 1.5, 101)
nval = [np.floor((n+1)/2) for n in range(2*COBWEB_MAX+2)]
# Set up side-by-side plots and initialize cobweb coordinate lists
figs1, ax1 = plt.subplots(1, 2, figsize=(10, 4))
cwx1 = np.zeros(2*COBWEB_MAX+2) # cobweb x-coordinates
cwy1 = np.zeros_like(cwx1) # cobweb y-coordinates
# Draw plot lines
@widgets.interact(step=(0, 2*COBWEB_MAX, 1), r=(0.5, 3.0, 0.01), x0=(0.0, 1.25, 0.01))
def update1(step=4, r=1.5, x0=0.25):
global ax1, cwx1, cwy1
# Cobweb plot
ax1[0].clear()
ax1[0].set_xlim([0, 1.5])
ax1[0].set_ylim([0, 1.5])
ax1[0].grid(False)
ax1[0].set_title("Cobweb Plot")
ax1[0].set_xlabel("$x_n$")
ax1[0].set_ylabel("$x_{n+1}$")
cobweb_update(x0, cwx1, cwy1, r=r, mode=0)
ax1[0].plot(x, lmap(x, r=r), color="C0")
ax1[0].plot(x, x, color="black")
ax1[0].plot(cwx1[:step+2], cwy1[:step+2], color="C1")
ax1[0].plot(cwx1[step:step+2], cwy1[step:step+2], color="red")
# Scatter plot
ax1[1].clear()
ax1[1].set_ylim([0, 1.5])
ax1[1].grid(False)
ax1[1].set_title("Time Series")
ax1[1].set_xlabel("$n$")
ax1[1].set_ylabel("$x_n$")
ax1[1].plot(np.append([0], nval[1:step+2:2]), np.append([x0], cwy1[1:step+2:2]), color="C0", marker=".", markersize=10)
ax1[1].plot(nval[step+1], cwy1[step+1], color="red", marker=".", markersize=10)
# + jupyter={"source_hidden": true} tags=[]
# Set up side-by-side plots and initialize cobweb coordinate lists
figs2, ax2 = plt.subplots(1, 2, figsize=(10, 4))
cwx2 = np.zeros(2*COBWEB_MAX+2) # cobweb x-coordinates
cwy2 = np.zeros_like(cwx2) # cobweb y-coordinates
# Draw plot lines
@widgets.interact(step=(0, 2*COBWEB_MAX, 1), r=(0.5, 4.0, 0.01), x0=(0.0, 1.25, 0.01))
def update2(step=4, r=1.5, x0=0.65):
global ax2, cwx2, cwy2
# Cobweb plot
ax2[0].clear()
ax2[0].set_xlim([0, 1.5])
ax2[0].set_ylim([0, 1.5])
ax2[0].grid(False)
ax2[0].set_title("Cobweb Plot")
ax2[0].set_xlabel("$x_n$")
ax2[0].set_ylabel("$x_{n+1}$")
cobweb_update(x0, cwx2, cwy2, r=r, mode=1)
ax2[0].plot(x, tmap(x, r=r), color="C0")
ax2[0].plot(x, x, color="black")
ax2[0].plot(cwx2[:step+2], cwy2[:step+2], color="C1")
ax2[0].plot(cwx2[step:step+2], cwy2[step:step+2], color="red")
# Scatter plot
ax2[1].clear()
ax2[1].set_ylim([0, 1.5])
ax2[1].grid(False)
ax2[1].set_title("Time Series")
ax2[1].set_xlabel("$n$")
ax2[1].set_ylabel("$x_n$")
ax2[1].plot(np.append([0], nval[1:step+2:2]), np.append([x0], cwy2[1:step+2:2]), color="C0", marker=".", markersize=10)
ax2[1].plot(nval[step+1], cwy2[step+1], color="red", marker=".", markersize=10)
# + tags=[]
| calc-diffeq-analysis/cobwebbing-standalone.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Epipolar Geometry
#
# _You can view [IPython Nootebook](README.ipynb) report._
#
# ----
#
# ## Contents
#
# - [GOAL](#GOAL)
# - [Basic Concepts](#Basic-Concepts)
# - [Code](#Code)
# - [Exercises](#Exercises)
#
# ## GOAL
#
# In this section:
#
# - We will learn about the basics of multiview geometry.
# - We will see what is epipole, epipolar lines, epipolar constraint etc.
#
# ## Basic Concepts
#
# When we take an image using pin-hole camera, we loose an important information, ie depth of the image. Or how far is each point in the image from the camera because it is a 3D-to-2D conversion. So it is an important question whether we can find the depth information using these cameras. And the answer is to use more than one camera. Our eyes works in similar way where we use two cameras (two eyes) which is called stereo vision. So let's see what OpenCV provides in this field.
#
# ### See also
#
# > [Learning OpenCV](https://www.amazon.com/Learning-OpenCV-Computer-Vision-Library/dp/0596516134) by **<NAME>** has a lot of information in this field.
#
# Before going to depth images, let's first understand some basic concepts in multiview geometry. In this section we will deal with epipolar geometry. See the image below which shows a basic setup with two cameras taking the image of same scene.
#
# 
#
# If we are using only the left camera, we can't find the 3D point corresponding to the point $ x $ in image because every point on the line $ OX $ projects to the same point on the image plane. But consider the right image also. Now different points on the line OX projects to different points ($ x′ $) in right plane. So with these two images, we can triangulate the correct 3D point. This is the whole idea.
#
# The projection of the different points on $ OX $ form a line on right plane (line $ l′ $). We call it **epiline** corresponding to the point $ x $. It means, to find the point $ x $ on the right image, search along this epiline. It should be somewhere on this line (Think of it this way, to find the matching point in other image, you need not search the whole image, just search along the epiline. So it provides better performance and accuracy). This is called **Epipolar Constraint**. Similarly all points will have its corresponding epilines in the other image. The plane $ XOO′ $ is called **Epipolar Plane**.
#
# $ O $ and $ O′ $ are the camera centers. From the setup given above, you can see that projection of right camera $ O′ $ is seen on the left image at the point, $ e $. It is called the **epipole**. Epipole is the point of intersection of line through camera centers and the image planes. Similarly $ e′ $ is the epipole of the left camera. In some cases, you won't be able to locate the epipole in the image, they may be outside the image (which means, one camera doesn't see the other).
#
# All the epilines pass through its epipole. So to find the location of epipole, we can find many epilines and find their intersection point.
#
# So in this session, we focus on finding epipolar lines and epipoles. But to find them, we need two more ingredients, **Fundamental Matrix (F)** and **Essential Matrix (E)**. Essential Matrix contains the information about translation and rotation, which describe the location of the second camera relative to the first in global coordinates. See the image below (Image courtesy: [Learning OpenCV](https://www.amazon.com/Learning-OpenCV-Computer-Vision-Library/dp/0596516134) by <NAME>):
#
# 
#
# But we prefer measurements to be done in pixel coordinates, right? Fundamental Matrix contains the same information as Essential Matrix in addition to the information about the intrinsics of both cameras so that we can relate the two cameras in pixel coordinates. (If we are using rectified images and normalize the point by dividing by the focal lengths, $ F=E $). In simple words, Fundamental Matrix F, maps a point in one image to a line (epiline) in the other image. This is calculated from matching points from both the images. A minimum of 8 such points are required to find the fundamental matrix (while using 8-point algorithm). More points are preferred and use RANSAC to get a more robust result.
#
# ## Code
#
# So first we need to find as many possible matches between two images to find the fundamental matrix. For this, we use SIFT descriptors with FLANN based matcher and ratio test.
#
# ```python
# import numpy as np
# import cv2 as cv
# from matplotlib import pyplot as plt
#
# img1 = cv.imread("../../data/left.jpg", 0) # Queryimage - left image
# img2 = cv.imread("../../data/right.jpg", 0) # Trainimage - right image
#
# # Initiate SIFT detector
# sift = cv.xfeatures2d.SIFT_create()
#
# # Find the keypoints and descriptors with SIFT
# kp1, des1 = sift.detectAndCompute(img1, None)
# kp2, des2 = sift.detectAndCompute(img2, None)
#
# # FLANN parameters
# FLANN_INDEX_KDTREE = 1
# index_params = dict(algorithm=FLANN_INDEX_KDTREE, trees=5)
# search_params = dict(checks=50)
# flann = cv.FlannBasedMatcher(index_params, search_params)
# matches = flann.knnMatch(des1, des2, k=2)
#
# # Ratio test as per Lowe's paper
# good = []
# pts1 = []
# pts2 = []
# for i, (m, n) in enumerate(matches):
# if m.distance < 0.8 * n.distance:
# good.append(m)
# pts2.append(kp2[m.trainIdx].pt)
# pts1.append(kp1[m.queryIdx].pt)
# ```
#
# Now we have the list of best matches from both the images. Let's find the Fundamental Matrix.
#
# ```python
# # We select only inlier points
# pts1 = np.int32(pts1)
# pts2 = np.int32(pts2)
# F, mask = cv.findFundamentalMat(pts1, pts2, cv.FM_LMEDS)
# pts1 = pts1[mask.ravel() == 1]
# pts2 = pts2[mask.ravel() == 1]
# ```
#
# Next we find the epilines. Epilines corresponding to the points in first image is drawn on second image. So mentioning of correct images are important here. We get an array of lines. So we define a new function to draw these lines on the images.
#
# ```python
# def drawlines(img1, img2, lines, pts1, pts2):
# """ img1 - image on which we draw the epilines for the points in img2
# lines - corresponding epilines """
# r, c = img1.shape
# img1 = cv.cvtColor(img1, cv.COLOR_GRAY2BGR)
# img2 = cv.cvtColor(img2, cv.COLOR_GRAY2BGR)
# for r, pt1, pt2 in zip(lines, pts1, pts2):
# color = tuple(np.random.randint(0, 255, 3).tolist())
# x0, y0 = map(int, [0, -r[2]/r[1]])
# x1, y1 = map(int, [c, -(r[2]+r[0]*c)/r[1]])
# img1 = cv.line(img1, (x0, y0), (x1, y1), color, 1)
# img1 = cv.circle(img1, tuple(pt1), 5, color, -1)
# img2 = cv.circle(img2, tuple(pt2), 5, color, -1)
# return img1, img2
# ```
#
# Now we find the epilines in both the images and draw them.
#
# ```python
# # Find epilines corresponding to points in left image (first image) and
# # drawing its lines on right image
# lines2 = cv.computeCorrespondEpilines(pts1.reshape(-1, 1, 2), 1, F)
# lines2 = lines2.reshape(-1, 3)
# img3, img4 = drawlines(img2, img1, lines2, pts2, pts1)
#
# # Find epilines corresponding to points in right image (second image) and
# # drawing its lines on left image
# lines1 = cv.computeCorrespondEpilines(pts2.reshape(-1, 1, 2), 2, F)
# lines1 = lines1.reshape(-1, 3)
# img5, img6 = drawlines(img1, img2, lines1, pts1, pts2)
#
# plt.subplot(121), plt.imshow(img5), plt.xticks([]), plt.yticks([])
# plt.subplot(122), plt.imshow(img3), plt.xticks([]), plt.yticks([])
# plt.subplots_adjust(left=0.01, right=0.99, wspace=0.02)
# plt.show()
# ```
#
# Below is the result we get:
#
# 
#
# You can see in the left image that all epilines are converging at a point outside the image at right side. That meeting point is the epipole.
#
# For better results, images with good resolution and many non-planar points should be used.
#
# ## Exercises
#
# 1. One important topic is the forward movement of camera. Then epipoles will be seen at the same locations in both with epilines emerging from a fixed point. [See this discussion](http://answers.opencv.org/question/17912/location-of-epipole/).
# 2. Fundamental Matrix estimation is sensitive to quality of matches, outliers etc. It becomes worse when all selected matches lie on the same plane. [Check this discussion](http://answers.opencv.org/question/18125/epilines-not-correct/).
#
| calibration-reconstruction/epipolar-geometry/README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="VvC5FVAg2-GM" colab_type="text"
# # Regression
#
# While classification is the problem of predicting the class of given data points, **regression** is the problem of learning the strength of association between features of a dataset and continuous outcomes. A **continuous** outcome is a real value such as an integer or floating point value often quantified as amounts and sizes. Simply, regression attempts to learn how strong the relationship is between features and outcomes. Formally, regression approximates a mapping function **f** from input variables **X** to a continuos variable **y**.
#
# In statistics, regression is typically defined as a measurement that attempts to determine the strength of the relationship between one dependent variable and a series of other changing variables known as independent variables. For those of you familiar with regression, you can think of independent variables as features and dependent variables as outcomes or outcome variables in the machine learning idiom.
#
# Since regression is used for predicting continous values, it can be referred to as **regression predictive modeling**. An algorithm capable of learning a regression predictive model is called a regression algorithm. Since regression predicts a quantity, *the performance must be measured as error* in those predictions.
#
# Performance of machine learning regression can be gauged in many ways. But, the most common are mean square error (MSE), mean absolute error (MAE), and root mean squared error (RMSE).
#
# **MSE** It is one of the most commonly used metrics, but least useful when a single bad prediction would ruin the entire model's predicting abilities. That is, when the dataset contains a lot of noise. It is most useful when the dataset contains outliers or unexpected values. Unexpected values are those that are too high or too low.
#
# **MAE** is not very sensitive to outliers in comparison to MSE since it doesn't punish huge errors. It is typically used when performance is measured on continuous variable data. It provides a linear value that averages the weighted individual differences equally.
#
# **RMSE** errors are squared before they are averaged. As such, RMSE assigns a higher weight to larger errors. So, RMSE is much more useful when large errors are present and they drastically affect the model's performance. A benefit of RMSE is that units of error score are the same as the predicted value.
# + [markdown] id="hK1Gi7H50Gox" colab_type="text"
# Enable the GPU (if not already enabled):
# 1. click **Runtime** in the top left menu
# 2. click **Change runtime type** from the drop-down menu
# 3. choose **GPU** from the *Hardware accelerator* drop-down menu
# 4. click **SAVE**
# + id="JW7eUCwg0hef" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7e26c04d-5a92-4907-f3aa-d62db7c64dc4"
import tensorflow as tf
# display tf version and test if GPU is active
tf.__version__, tf.test.gpu_device_name()
# + [markdown] id="-zhMawEn0TVZ" colab_type="text"
# Import the tensorflow library. If '/device:GPU:0' is displayed, the GPU is active. If '..' is displayed, the regular CPU is active.
# + [markdown] id="LedfTVyqJkJ1" colab_type="text"
# # Boston Housing Dataset
#
# The first regression dataset we explore is Boston Housing. **Boston Housing** is a dataset derived from information collected by the U.S. Census Service concerning housing in the area of Boston, Massachusetts. It was obtained from the StatLib archive (http://lib.stat.cmu.edu/datasets/boston), and has been used extensively throughout the machine learning literature to benchmark algorithms. The dataset is small in size with only 506 cases.
#
# The name for this dataset is simply **boston**. It contains 12 features and 1 outcome (or target). The features are as follows:
#
# 1. CRIM - per capita crime rate by town
# 2. ZN - proportion of residential land zoned for lots over 25,000 sq.ft.
# 3. INDUS - proportion of non-retail business acres per town.
# 4. CHAS - Charles River dummy variable (1 if tract bounds river; 0 otherwise)
# 5. NOX - nitric oxides concentration (parts per 10 million)
# 6. RM - average number of rooms per dwelling
# 7. AGE - proportion of owner-occupied units built prior to 1940
# 8. DIS - weighted distances to five Boston employment centres
# 9. RAD - index of accessibility to radial highways
# 10. TAX - full-value property-tax rate per $10,000
# 11. PTRATIO - pupil-teacher ratio by town
# 12. LSTAT - % lower status of the population
#
# The target is:
#
# * MEDV - median value of owner-occupied homes in $1000's
#
# Data was collected in the '70s, so don't be shocked by the low median value of homes.
# + [markdown] id="bDv_GGAXqARW" colab_type="text"
# ## Boston Data
#
# You can access any dataset for this book directly from GitHub with a few simple steps:
#
# 1. visit the book URL: https://github.com/paperd/tensorflow
# 2. locate the dataset and click on it
# 3. click the **Raw** button
# 4. copy the URL to Colab and assign it to a variable
# 5. read the dataset with Pandas **read_csv** method
#
# For convenience, we've already located the appropriate URL and assigned to a variable as so:
# + id="ktRCoDA2sAC6" colab_type="code" colab={}
url = 'https://raw.githubusercontent.com/paperd/tensorflow/\
master/chapter6/data/boston.csv'
# + [markdown] id="slf-ukRAzFvL" colab_type="text"
# Read the dataset into a Pandas DataFrame:
# + id="9UyY0SgjsRpE" colab_type="code" colab={}
import pandas as pd
data = pd.read_csv(url)
# + [markdown] id="E_ofnmbczlHL" colab_type="text"
# Verify that data was read properly:
# + id="Vqe-Yv8du5JX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 197} outputId="ad9d3915-a8fb-4860-b2a0-8414c7e446f2"
data.head()
# + [markdown] id="xXa89NmTw68J" colab_type="text"
# ## Explore the Dataset
# + [markdown] id="vrqd6hb9z-me" colab_type="text"
# Get datatypes:
# + id="eVcFWvh8wntF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="39ca63d0-e42a-45fa-de5a-13809b6f39a0"
data.dtypes
# + [markdown] id="WOQC0OnAw-C9" colab_type="text"
# Get general information:
# + id="n4ucq5OMw_n2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 372} outputId="32b912d3-aae8-469b-aece-94801582effe"
data.info()
# + [markdown] id="w5322Or9xCq-" colab_type="text"
# Create a DataFrame that holds basic statistics with the describe method and transpose it for easier viewing:
# + id="b0oPawBywvfh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 437} outputId="fb0f33b6-b431-47f7-945e-e5eee3969169"
data_t = data.describe()
desc = data_t.T
desc
# + [markdown] id="xuxYRUxrbdgx" colab_type="text"
# Target specific statistics from the transposed DataFrame:
# + id="jhUjNG-ubdpX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 437} outputId="bdbe74ac-5445-4a98-a54d-e38b21628178"
desc[['mean', 'std']]
# + [markdown] id="csqLKtgOxW95" colab_type="text"
# Describe a specific feature:
# + id="NFFLaTHbxGhu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 176} outputId="0b652e5e-455f-434f-9f46-16a3db848599"
data.describe().LSTAT
# + [markdown] id="f4S10R1VxsR9" colab_type="text"
# Or:
# + id="oSrZQR8CxoC9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 176} outputId="adbb48f8-87c6-4839-e511-54b845a09ee9"
data['LSTAT'].describe()
# + [markdown] id="9kX3uwtnvsyT" colab_type="text"
# Get columns:
# + id="Ma0GQx_9wc2N" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 247} outputId="e6c287dd-c252-4435-ea72-c1c81b3c1686"
cols = list(data)
cols
# + [markdown] id="dI7Nn6Nn2yFe" colab_type="text"
# ## Create Feature and Target Sets
#
# We need to create the feature set and target from the data, so let's do that now:
# + id="NEjZO7VD0W-T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 123} outputId="f83fcdbd-5e44-492d-a3dc-f15a1f8bea6e"
# create a copy of the DataFrame
df = data.copy()
# create the target
target = df.pop('MEDV')
print (target.head())
# + [markdown] id="eqPdXTW63W2R" colab_type="text"
# Verify that feature set is as expected:
# + id="Qa5MK7Hm3clR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 197} outputId="4b096133-8fa3-467a-f6d7-ad35da698d34"
df.head()
# + [markdown] id="b0qDofPL3lYJ" colab_type="text"
# ## Get Feature Names from the Features DataFrame
#
# It is easy to get the features since the target is no longer part of the DataFrame:
# + id="z2ekWP1K3qxL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 230} outputId="1f4f93d4-c0f7-4c71-8583-4f6d9861c65e"
feature_cols = list(df)
feature_cols
# + [markdown] id="C1W_h9RN4fCp" colab_type="text"
# Get number of features:
# + id="5P2L_2w_4gpH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="229f2e0a-185e-4c50-e643-ec9195841592"
len(feature_cols)
# + [markdown] id="N54kSxyv4oKo" colab_type="text"
# Or:
# + id="IXmW7klY4kgg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3c875abb-e474-479f-f8ba-9b8de6a85cc7"
len(df.columns)
# + [markdown] id="jEzC5rjb5HPM" colab_type="text"
# ## Convert Features and Labels to Numpy
#
# Convert Pandas DataFrame values to Numpy with the *values* method:
# + id="F2LPNUWl5Vs8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c4ab564a-43e5-4513-86af-67d4e81c3156"
features = df.values
labels = target.values
type(features), type(labels)
# + [markdown] id="N4TQdMlq5ZrT" colab_type="text"
# ## Split into train and test sets:
# + id="bMtTFidt5dfk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="6c98311a-3691-46ca-c127-ab30152ef530"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
features, labels, test_size=0.33, random_state=0)
br = '\n'
print ('X_train shape:', end=' ')
print (X_train.shape, br)
print ('X_test shape:', end=' ')
print (X_test.shape)
# + [markdown] id="49MtJKvH5yC3" colab_type="text"
# ## Scale Data and Create TensorFlow Tensors
#
# With image data, we scale by dividing each element by 255.0 to ensure that each input parameter (a pixel, in our case) has a similar data distribution. However, features represented by continous values are scaled differently. We rescale continuous data to have a mean (μ) of 0 and standard deviation (σ) of 1. A σ of 1 is called unit variance.
# + id="Wstb8WM756LP" colab_type="code" colab={}
# scale feature data and create TensorFlow tensors
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_train_std = scaler.fit_transform(X_train)
X_test_std = scaler.fit_transform(X_test)
train = tf.data.Dataset.from_tensor_slices(
(X_train_std, y_train))
test = tf.data.Dataset.from_tensor_slices(
(X_test_std, y_test))
# + [markdown] id="nSGJ74he77B0" colab_type="text"
# Let's view the first tensor:
# + id="b2S9LDDc79Kk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="30870082-4e2d-43aa-88e4-119741aaee6e"
def see_samples(data, num):
for feat, targ in data.take(num):
print ('Features: {}'.format(feat), br)
print ('Target: {}'.format(targ))
n = 1
see_samples(train, n)
# + [markdown] id="YIPqAUdr8xPO" colab_type="text"
# The first sample looks exactly as we expect.
# + [markdown] id="3hrGInpK5xNO" colab_type="text"
# ## Prepare Tensors for Training
# + id="yXJYOwMQ6gJB" colab_type="code" colab={}
BATCH_SIZE, SHUFFLE_BUFFER_SIZE = 16, 100
train_bs = train.shuffle(
SHUFFLE_BUFFER_SIZE).batch(BATCH_SIZE).prefetch(1)
test_bs = test.batch(BATCH_SIZE).prefetch(1)
# + [markdown] id="v1Asble50w05" colab_type="text"
# Inspect tensors:
# + id="JaPcW-OQ0y8X" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="80714859-c06e-4157-d943-90400c8d85e7"
train_bs, test_bs
# + [markdown] id="iNcHvDts7ksK" colab_type="text"
# ## Create a Model
#
# If we don't have a lot of training data, one technique to avoid overfitting is to create a small network with few hidden layers. We do just that!
#
# The 64 neuron input layer accommodates our 12 input features. We have one hidden layers with 64 neurons. The output layer has 1 neuron because we are using regression.
# + id="lIDe1Mxv7eSk" colab_type="code" colab={}
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Flatten
import numpy as np
# clear any previous model
tf.keras.backend.clear_session()
# generate a seed for replication purposes
np.random.seed(0)
tf.random.set_seed(0)
# notice input shape accommodates 12 features!
model = Sequential([
Dense(64, activation='relu', input_shape=[12,]),
Dense(64, activation='relu'),
Dense(1)
])
# + [markdown] id="UFyAY087wBl0" colab_type="text"
# ## Inspect the Model
# + id="mX7BmLnIwBvz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="05005227-d1a9-42fb-e2a0-6607c6808d63"
model.summary()
# + [markdown] id="ILxUyoN9bX0K" colab_type="text"
# ## Compile the Model
# + id="Qjpdb8QD-E5o" colab_type="code" colab={}
rmse = tf.keras.metrics.RootMeanSquaredError()
model.compile(loss='mse', optimizer='RMSProp',
metrics=[rmse, 'mae', 'mse'])
# + [markdown] id="X0kSUzSebW0T" colab_type="text"
# Mean Squared Error (MSE) is a common loss function used for regression problems. Mean Absolute Error (MAE) and RMSE are also common metrics. With some experimentation, we found that **RMSProp** performed pretty well with this dataset.
# + [markdown] id="VJpNRr7sbbwS" colab_type="text"
# ## Train the Model
# + id="vOHauLH1_264" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="c7845a27-6e8a-41b7-f184-cc60d3b394c1"
history = model.fit(train_bs, epochs=50,
validation_data=test_bs)
# + [markdown] id="VdbqyS21Z9py" colab_type="text"
# ## Visualize Training
#
# Let's try a different technique to visualize. Begin by creating variable **hist** that holds the model's history as a Pandas DataFrame. Create another variable **hist['epoch']** to hold epoch history. Display the last five rows to get an idea about performance.
# + id="6642hhv_bE6k" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 197} outputId="350cd6a3-6003-40a5-9906-13575b838cee"
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
hist.tail()
# + [markdown] id="PrjulWObh0MT" colab_type="text"
# Build the plots:
# + id="QVCSdFw7h2Ix" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 886} outputId="f387f995-84c5-4165-c949-4fae1eb306b3"
import matplotlib.pyplot as plt
def plot_history(history, limit1, limit2):
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
plt.figure()
plt.xlabel('epoch')
plt.ylabel('MAE [MPG]')
plt.plot(hist['epoch'], hist['mae'],
label='Train Error')
plt.plot(hist['epoch'], hist['val_mae'],
label = 'Val Error')
plt.ylim([0, limit1])
plt.legend()
plt.title('MAE by Epoch')
plt.show()
plt.clf()
plt.figure()
plt.xlabel('Epoch')
plt.ylabel('MSE [MPG]')
plt.plot(hist['epoch'], hist['mse'],
label='Train Error')
plt.plot(hist['epoch'], hist['val_mse'],
label = 'Val Error')
plt.ylim([0, limit2])
plt.legend()
plt.title('MSE by Epoch')
plt.show()
plt.clf()
plt.figure()
plt.xlabel('Epoch')
plt.ylabel('RMSE [MPG]')
plt.plot(hist['epoch'], hist['root_mean_squared_error'],
label='Train Error')
plt.plot(hist['epoch'], hist['val_root_mean_squared_error'],
label = 'Val Error')
plt.ylim([0, limit2])
plt.legend()
plt.title('RMSE by Epoch')
plt.show()
# set limits to make plot readable
mae_limit, mse_limit = 10, 100
plot_history(history, mae_limit, mse_limit)
# + [markdown] id="WXkMIR1NODys" colab_type="text"
# Since the validation error is a worse than the train error (for MAE and MSE at least), the model is overfitting. What can we do? The first step is to estimate when performance begins to degrade. From the visualizations, can you see when this happens?
# + [markdown] id="KhMb48ROjvRF" colab_type="text"
# ## Early Stopping
#
# With classification, our goal is to maximize accuracy. Of course, we also want to minimize loss. With regression, our goal is to minimize **MSE** or one of the other error metrics. From the visualizations, we see that our model is overfitting because validation error is higher than training error. We also see that once train and validation error cross, performance begins to degrade.
#
# There is one simple tuning experiment we can run to make this model more useful. We can stop the model when training and validation error are very close to each other. This technique is called early stopping. **Early stopping** is a widely used approach that stops training at the point when performance on a validation dataset starts to degrade.
#
# Let's modify our training experiment to automatically stop training when the validation score doesn't improve. We use an **EarlyStopping** callback that tests a training condition for every epoch. If a set amount of epochs elapse without showing improvement, training is automatically stopped.
#
# All we need to do is update our fit() method and rerun training:
# + id="N5KR6ta4mFCX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 230} outputId="8443c721-0048-44b8-cca1-551feecc95d8"
# clear the previous model
tf.keras.backend.clear_session()
# generate a seed for replication purposes
np.random.seed(0)
tf.random.set_seed(0)
# monitor 'val_loss' for early stopping
early_stop = tf.keras.callbacks.EarlyStopping(monitor='val_loss')
history = model.fit(train_bs, epochs=50,
validation_data=test_bs,
callbacks=[early_stop])
# + [markdown] id="XYjx3SYewatv" colab_type="text"
# Although we can get great results with early stopping, it is a good idea to add some control to ensure the best performance. For added control, we can add a parameter that forces the model to continue to a point that gives us the best performance. The **patience** parameter can be set to a given a number of epochs after which training will be stopped if there is no improvement. Let's try this and see what happens.
# + id="y9P-M8GiufXW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 407} outputId="f6a7da98-ebcc-4554-be95-1fcf267150a5"
# clear the previous model
tf.keras.backend.clear_session()
# generate a seed for replication purposes
np.random.seed(0)
tf.random.set_seed(0)
# set number of patience epochs
n = 4
early_stop = tf.keras.callbacks.EarlyStopping(
monitor='val_loss', patience=n)
history = model.fit(train_bs, epochs=50,
validation_data=test_bs,
callbacks=[early_stop])
# + [markdown] id="o_AMHiaEnN-N" colab_type="text"
# Now, we have a better model because it stops closer to the ideal performance.
#
# Let's plot:
# + id="gWQNPB0RnP5M" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 886} outputId="6e52d996-4e9c-48b9-b011-64808227bc59"
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
train_limit, test_limit = 10, 100
plot_history(history, train_limit, test_limit)
# + [markdown] id="bnGaZrlBnWcb" colab_type="text"
# ## Remove Bad Data
#
# Although early stopping worked as expected, we might be able to squeeze out a bit more performance with this dataset. The boston dataset has some bad data so we can modify it.
#
# What is wrong with the data? Prices of homes are capped at $50,000 because the Census Service censored the data. They decided to set the maximum value of the price variable to 50k USD, so no price can go beyond that value.
#
# What do we do? While maybe not ideal, we can remove data with prices at or above 50k USD. This is not ideal because we may be removing perfectly good data, but there is no way to know this. Another reason is because the dataset is so small to begin with. Neural nets are meant to perform at their best with larger datasets.
#
# Keep in mind that our book is an introduction, so we are not nearly as concerned with performance as we would be if working on real datasets. We are just trying to help you learn how to use TensorFlow 2.x. If you want to explore this topic further, we recommend this URL:
#
# https://towardsdatascience.com/things-you-didnt-know-about-the-boston-housing-dataset-2e87a6f960e8
# + [markdown] id="xVwxZFdHvAP6" colab_type="text"
# ## Get Data
#
# To make this happen, we have to return to the original data because we've processed the heck out of it to ready it for TensorFlow consumption. So let's get started:
# + id="OMWhR_VEsNFz" colab_type="code" colab={}
# get the raw data
url = 'https://raw.githubusercontent.com/paperd/tensorflow/\
master/chapter6/data/boston.csv'
boston = pd.read_csv(url)
# + [markdown] id="f1ZF45fpK54_" colab_type="text"
# Verify data:
# + id="mx9YONJKK6AL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 197} outputId="ca9ded36-1d7c-447e-cb99-567ad039a361"
boston.head()
# + [markdown] id="Dq-cW8LNxfc8" colab_type="text"
# ## Remove Noise
# + id="BqpBfLBiv3w0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="2b7b9f67-e143-4b0f-fa95-e20b61e089ca"
print ('data set before removing noise:', boston.shape)
# remove noise
noise = boston.loc[boston['MEDV'] >= 50]
data = boston.drop(noise.index)
print ('data set without noise:', data.shape)
# + [markdown] id="f9D-0FBBwGhb" colab_type="text"
# ## Create Feature and Target Data
#
# So, we now have a dataset without the possibly corrupted data. Split the data into feature and target sets:
# + id="xQixTaRjwXKE" colab_type="code" colab={}
# create a copy of the DataFrame
df = data.copy()
# create feature and target sets
target, features = df.pop('MEDV'), df.values
labels = target.values
# + [markdown] id="Ir-nAuxWwfO0" colab_type="text"
# ## Build the Input Pipeline
# + id="6J3pY8azwo4O" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(
features, labels, test_size=0.33, random_state=0)
# standardize feature data and create TensorFlow tensors
X_train_std = scaler.fit_transform(X_train)
X_test_std = scaler.fit_transform(X_test)
# slice data for TensorFlow consumption
train = tf.data.Dataset.from_tensor_slices(
(X_train_std, y_train))
test = tf.data.Dataset.from_tensor_slices(
(X_test_std, y_test))
# shuffle, batch, prefetch
BATCH_SIZE = 16
SHUFFLE_BUFFER_SIZE = 100
train_n = train.shuffle(
SHUFFLE_BUFFER_SIZE).batch(BATCH_SIZE).prefetch(1)
test_n = test.batch(BATCH_SIZE).prefetch(1)
# + [markdown] id="3LNo6kd61an0" colab_type="text"
# Inspect tensors:
# + id="aBRZcldO1b5O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="03661309-b8c1-4555-dd1f-58e8b02811bc"
train_n, test_n
# + [markdown] id="Nt5KAvKXeS3U" colab_type="text"
# ## Compile and Train
# + id="xzY0_Hd5eS-r" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 372} outputId="2ff17af4-369d-4f07-9c15-e6114352ae10"
rmse = tf.keras.metrics.RootMeanSquaredError()
model.compile(loss='mse', optimizer='RMSProp',
metrics=[rmse, 'mae', 'mse'])
tf.keras.backend.clear_session()
# generate a seed for replication purposes
np.random.seed(0)
tf.random.set_seed(0)
n = 4
early_stop = tf.keras.callbacks.EarlyStopping(
monitor='val_loss', patience=n)
history = model.fit(train_n, epochs=50,
validation_data=test_n,
callbacks=[early_stop])
# + [markdown] id="j9z83OyU0e7t" colab_type="text"
# ## Let's Visualize
# + id="yvR2d3XK0fF7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 886} outputId="d493e29d-fafc-44bc-91a3-e4bea172a87e"
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
train_limit, test_limit = 10, 100
plot_history(history, train_limit, test_limit)
# + [markdown] id="C_Jrdzdt0CMl" colab_type="text"
# Our model is not perfect, be we did improve performance.
# + [markdown] id="D4t7QFwN6LxV" colab_type="text"
# ## Generalize on Test Data
#
# Let's see how well the model generalizes with test data, which tells us how well we can expect the model to predict when we use it in the real world.
# + id="i5XL06mn6L7X" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="b9d21800-cb66-435f-f8f1-79ee761cc494"
loss, rmse, mae, mse = model.evaluate(test_n, verbose=2)
print ()
print('"Testing set Mean Abs Error: {:5.2f} thousand dollars'.
format(mae))
# + [markdown] id="aljDYGYD7kZd" colab_type="text"
# As we know, Mean Square Error (MSE) is a common loss function used for regression problems. Another common regression metric is Mean Absolute Error (MAE). MAE gives us a good idea how well our model performed in an easy to understand way. So, we can expect that our model predictions are off by the MAE value in thousands of dollars.
# + [markdown] id="FqS1sCkb5cFx" colab_type="text"
# ## Make Predictions
# + id="32UEqma95do9" colab_type="code" colab={}
predictions = model.predict(test_n)
# + [markdown] id="dV5Ruefl8QOC" colab_type="text"
# Display the first prediction:
# + id="3jE7Qw4p8QXY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="95850abd-3955-46c5-eb95-f05766b4a9ea"
# predicted housing price
first = predictions[0]
print ('predicted price:', first[0], 'thousand')
# actual housing price
print ('actual price:', y_test[0], 'thousand')
# + [markdown] id="iuW3tnCh9wQF" colab_type="text"
# Display the first five predictions:
# + id="QWDDb3vo8vUl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 123} outputId="6af69614-7cb5-4bb2-9050-eaef234204e1"
five = predictions[:5]
actuals = y_test[:5]
print ('pred', 'actual')
for i, p in enumerate(range(5)):
print (np.round(five[i][0],1), actuals[i])
# + [markdown] id="6GezQCoDAXzF" colab_type="text"
# ## Visualize Predictions
# + id="1WSnUbtBAYBD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 278} outputId="a6102605-2523-41bd-cbf4-03616dba0217"
fig, ax = plt.subplots()
ax.scatter(y_test, predictions)
ax.plot([y_test.min(), y_test.max()],\
[y_test.min(), y_test.max()], 'k--', lw=4)
ax.set_xlabel('Measured')
ax.set_ylabel('Predicted')
plt.show()
# + [markdown] id="L4bkl8gCC4ED" colab_type="text"
# The further a prediciton is away from the diagonal (true values), the more erroneous it is.
# + [markdown] id="PnYmzbJA0ynT" colab_type="text"
# # Load Boston Data from Scikit-Learn
#
# Since the Boston dataset is included in **sklearn.datasets**, let's load it from this environment.
# + id="3Q4SP-dJ1OEK" colab_type="code" colab={}
from sklearn import datasets
dataset = datasets.load_boston()
data, target = dataset.data, dataset.target
# + [markdown] id="Pyuhb3if1uB0" colab_type="text"
# It's easier to load because we don't have to access a CSV file. On the downside, the data is not loaded as a DataFrame. But, we can create a DataFrame with a few simple steps.
# + [markdown] id="Sd7XHVeXMRcA" colab_type="text"
# ## Build a DataFrame from a **sklearn** dataset
# + [markdown] id="O7GBBz1bKsNo" colab_type="text"
# First, access information about the dataset with the **keys()** function.
# + id="5mGWEh7yK9LY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d835995b-29ea-4636-bd83-1290ed739012"
dataset.keys()
# + [markdown] id="nS21SwD9LAXn" colab_type="text"
# Second, access the feature names.
# + id="mAO2vYiiK-8-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="534859d2-703a-498c-a5a2-76f5766d06cf"
feature_names = dataset.feature_names
feature_names
# + [markdown] id="sEEINWNHLiNQ" colab_type="text"
# Third, build a dataframe with feature and target data.
# + id="ZDAx1B0DKUlv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 197} outputId="38f947f3-4214-458e-bf4f-53d043febd07"
df_sklearn = pd.DataFrame(dataset.data, columns=feature_names)
df_sklearn['MEDV'] = dataset.target
df_sklearn.head()
# + [markdown] id="L5uk9b2Pnaxm" colab_type="text"
# Check information:
# + id="QacJ8roampKr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="b749931c-99c7-404d-9d63-80d025d6de1d"
df_sklearn.info()
# + [markdown] id="5su8IOmhKTAp" colab_type="text"
# ## Remove Noise
# + id="Q-YiPQnC1p4C" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="fb857cc1-7862-4ba9-8b1e-6cdd5e106262"
# remove noisy data
print ('data set before removing noise:', df_sklearn.shape)
noise = df_sklearn.loc[df_sklearn['MEDV'] >= 50]
df_clean = df_sklearn.drop(noise.index)
print ('data set without noise:', df_clean.shape)
# + [markdown] id="OOcbyYgt2mL1" colab_type="text"
# ## Build the Input Pipeline
# + id="tg4fWZWA2mYF" colab_type="code" colab={}
# create a copy of the DataFrame
df = df_clean.copy()
# create the target
target = df.pop('MEDV')
# convert features and target data
features = df.values
labels = target.values
# create train and test sets
X_train, X_test, y_train, y_test = train_test_split(
features, labels, test_size=0.33, random_state=0)
X_train_std = scaler.fit_transform(X_train)
X_test_std = scaler.fit_transform(X_test)
# slice data into a TensorFlow consumable form
train = tf.data.Dataset.from_tensor_slices(
(X_train_std, y_train))
test = tf.data.Dataset.from_tensor_slices(
(X_test_std, y_test))
# finalize the pipeline
BATCH_SIZE = 16
SHUFFLE_BUFFER_SIZE = 100
train_sk = train.shuffle(
SHUFFLE_BUFFER_SIZE).batch(BATCH_SIZE).prefetch(1)
test_sk = test.batch(BATCH_SIZE).prefetch(1)
# + [markdown] id="ZeKpqCRLW85R" colab_type="text"
# ## Create a New Model and Compile
#
# You may not have noticed, but the **sklearn** boston dataset has an extra feature column. So, we can either remove the feature or create a new model that accommodates it. Let's just create a new model and compile. Notice that we changed the input shape to accommodate the extra feature.
# + id="_vcoBFZyXYEh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 656} outputId="aebe07d9-50a8-49d4-91f0-a344274292c0"
# clear any previous model
tf.keras.backend.clear_session()
# generate a seed for replication purposes
np.random.seed(0)
tf.random.set_seed(0)
# new model with 13 input features
model = Sequential([
Dense(64, activation='relu', input_shape=[13,]),
Dense(64, activation='relu'),
Dense(1)
])
# compile the new model
rmse = tf.keras.metrics.RootMeanSquaredError()
model.compile(loss='mse', optimizer='RMSProp',
metrics=[rmse, 'mae', 'mse'])
# train
n = 4
early_stop = tf.keras.callbacks.EarlyStopping(monitor='val_loss',
patience=n)
history = model.fit(train_sk, epochs=50,
validation_data=test_sk,
callbacks=[early_stop])
# + [markdown] id="9cd64V-vEIVJ" colab_type="text"
# # Read from UCI Irvine Machine Learning Repository
#
# We can also read housing data directly from the UCI Irvine Machine Learning Repository.
# + id="9z-kYHTFEIrB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} outputId="37a5cd0d-f7f0-471a-8f2b-fda750536ab6"
dataset_path = tf.keras.utils.get_file('housing.data', 'https://archive.ics.uci.\
edu/ml/machine-learning-databases/housing/housing.data')
dataset_path
# + id="EkfTV4JeGCiZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 137} outputId="f6135546-f286-4b4a-92e2-e9154e0cf14e"
cols = ['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS',
'RAD', 'TAX', 'PTRATIO', 'B', 'LSTAT', 'MEDV']
uci_data = pd.read_csv(dataset_path, names=cols, sep=' ',
skipinitialspace=True)
uci_data.head(3)
# + id="4pL32EoarTxt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="3521925f-4690-4888-9e24-e2d0fa5a37f4"
uci_data.info()
# + [markdown] id="U966zKTUIlIp" colab_type="text"
# ## Create the Input Pipeline
# + id="qg9tPVjpIlS4" colab_type="code" colab={}
# create a copy of the DataFrame
df = uci_data.copy()
# create the target
target = df.pop('MEDV')
# prepare features and labels
features = df.values
labels = target.values
# create and train and test data
X_train, X_test, y_train, y_test = train_test_split(
features, labels, test_size=0.33, random_state=0)
# scale data
X_train_std = scaler.fit_transform(X_train)
X_test_std = scaler.fit_transform(X_test)
# prepare train and test data for TensorFlow consumption
train = tf.data.Dataset.from_tensor_slices((X_train_std, y_train))
test = tf.data.Dataset.from_tensor_slices((X_test_std, y_test))
# shuffle, batch, and prefetch
BATCH_SIZE = 16
SHUFFLE_BUFFER_SIZE = 100
train_ml = train.shuffle(SHUFFLE_BUFFER_SIZE).batch(BATCH_SIZE).prefetch(1)
test_ml = test.batch(BATCH_SIZE).prefetch(1)
# + [markdown] id="o4M9c2TaJyEr" colab_type="text"
# ## Compile and Model Data
# + id="5B4id2LlJkos" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="6ff92f2a-b007-48ef-827b-f35ab952af10"
# clear any previous model
tf.keras.backend.clear_session()
# plant a random seed for replication purposes
np.random.seed(0)
tf.random.set_seed(0)
# new model with 13 input features
model = Sequential([
Dense(64, activation='relu', input_shape=[13,]),
Dense(64, activation='relu'),
Dense(1)
])
# compile the new model
rmse = tf.keras.metrics.RootMeanSquaredError()
model.compile(loss='mse', optimizer='RMSProp',
metrics=[rmse, 'mae', 'mse'])
# train
n = 4
early_stop = tf.keras.callbacks.EarlyStopping(monitor='val_loss', patience=n)
history = model.fit(train_ml, epochs=50,
validation_data=test_ml,
callbacks=[early_stop])
# + [markdown] id="V3BgDTnS4Zv0" colab_type="text"
# # Model the Cars Dataset
#
# To get more practice, let's try another dataset.
# + [markdown] id="fpE90Z-lUXoF" colab_type="text"
# ## Get Cars Data from GitHub
#
# You can access any free standing dataset for this book directly from GitHub with a few simple steps:
#
# 1. visit the book URL: https://github.com/paperd/tensorflow
# 2. locate the dataset and click on it
# 3. click the Raw button
# 4. copy the URL to Colab and assign it to a variable
# 5. read the dataset with the Pandas read_csv method
# + [markdown] id="A5oSilX2Up0X" colab_type="text"
# We’ve already located the URL and assigned it to a variable:
# + id="x75xfFCeUsvv" colab_type="code" colab={}
cars_url = 'https://raw.githubusercontent.com/paperd/tensorflow/\
master/chapter6/data/cars.csv'
# + [markdown] id="7LqH7ZHXWwsH" colab_type="text"
# Read the dataset into a Pandas DataFrame:
# + id="1GhATmFIWw0N" colab_type="code" colab={}
cars = pd.read_csv(cars_url)
# + [markdown] id="LzsbvIfHW_eO" colab_type="text"
# Verify data:
# + id="xnfTcEviW_kn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 197} outputId="0671a217-b989-4a25-caa4-5fead115d20f"
cars.head()
# + [markdown] id="qE0z68OZZzy6" colab_type="text"
# Get information about dataset:
# + id="7gwYV3PDna2_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 289} outputId="0b0a86a2-9f31-4f49-d5ff-a866fc96a622"
cars.info()
# + [markdown] id="u9rK49_2oFMe" colab_type="text"
# ## Convert Categorical Column to Numeric
#
# Machine learning algorithms can only train numeric data. So, we must convert any non-numeric feature. The 'Origin' column is categorical, not numeric. To remedy, one solution is to encode the data as *one-hot*. **One hot encoding** is a process that converts categorical data into a numeric form for machine learning algorithm consumption.
#
# We start by slicing off the 'Origin' feature column from the original DataFrame into its own DataFrame. We then use this DataFrame as a template to build a new feature column in the original DataFrame for each category from the original 'Origin' feature.
# + id="-p9S_pG9qvYK" colab_type="code" colab={}
# create a copy of DataFrame
df = cars.copy()
origin = df.pop('Origin')
# + [markdown] id="bZWNEtdOrDSw" colab_type="text"
# Now, we define a feature column for 'US', 'Europe', and 'Japan' cars:
# + id="bbSkoUbDoFWM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 287} outputId="b22b8e1a-c3d8-416f-c796-ba30d5c03ea9"
df['US'] = (origin == 'US') * 1.0
df['Europe'] = (origin == 'Europe') * 1.0
df['Japan'] = (origin == 'Japan') * 1.0
df.tail(8)
# + [markdown] id="dq2cCot4rOnx" colab_type="text"
# So, for each entry in the 'US' feature column, we assign a '1.0' if the car is from the United States. Otherwise, we assign a '0.0'. We follow the same logic for the 'Europe' and 'Japan' feature columns based on the car's origin. Now, the algorithm can process the data properly.
# + [markdown] id="339fLZfpuFv7" colab_type="text"
# ## Slice Extraneous Data
#
# Since the name of each car has no impact on any predictions we might want to make, we can tuck it away into its own DataFrame in case we want to revisit it in the future.
# + id="evOZ7OnJvYGN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="29eeb98d-f7f8-46c1-bca3-a8d908588be1"
try:
name = df.pop('Car')
except:
print("An exception occurred")
# + [markdown] id="0c8psgMcvv-w" colab_type="text"
# If an exception occurrs, the *Car* column has already been removed.
# + [markdown] id="pVoMB5PlwL4z" colab_type="text"
# Verify:
# + id="-KwHEouxvaA9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 287} outputId="e064ee91-6554-4c70-89c1-d0e29224d45f"
df.tail(8)
# + [markdown] id="JGFD-H8ssvlz" colab_type="text"
# ## Create Features and Labels
#
# Our goal is to predict 'MPG' for cars in this dataset. So, the target is 'MPG' and features are the remaining feature columns.
# + [markdown] id="B3q_-5-UxrqV" colab_type="text"
# Create features and targets:
# + id="ZoFz6ESYxw5G" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="322aa075-865d-49ec-9b2a-08734910428c"
# create data sets
features = df.copy()
target = features.pop('MPG')
# get feature names
feature_cols = list(features)
print (feature_cols)
# get number of features
num_features = len(feature_cols)
print (num_features)
# convert feature and target data to float
features = features.values
labels = target.values
(type(features), type(labels))
# + [markdown] id="OHON9vQdsSg6" colab_type="text"
# ## Build the Input Pipeline
# + id="7TvDVxnus0-N" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="115d408a-dc08-40d9-c2ff-b2b783690110"
# split
X_train, X_test, y_train, y_test = train_test_split(
features, labels, test_size=0.33, random_state=0)
print ('X_train shape:', end=' ')
print (X_train.shape, br)
print ('X_test shape:', end=' ')
print (X_test.shape)
# scale
X_train_std = scaler.fit_transform(X_train)
X_test_std = scaler.fit_transform(X_test)
# slice
train = tf.data.Dataset.from_tensor_slices(
(X_train_std, y_train))
test = tf.data.Dataset.from_tensor_slices(
(X_test_std, y_test))
# shuffle, batch, prefetch
BATCH_SIZE = 16
SHUFFLE_BUFFER_SIZE = 100
train_cars = train.shuffle(
SHUFFLE_BUFFER_SIZE).batch(BATCH_SIZE).prefetch(1)
test_cars = test.batch(BATCH_SIZE).prefetch(1)
# + [markdown] id="k8oD8xtO2iHm" colab_type="text"
# Inspect tensors:
# + id="6Afhkdb12iNo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="692525a0-177e-4db7-d076-a1f325314cd6"
train_cars, test_cars
# + [markdown] id="FX5PjiJkvnDk" colab_type="text"
# ## Model Data
# + id="p5mbECFVvnPi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 941} outputId="86626154-d3d0-43f6-f8b2-1fca501c215c"
# clear any previous model
tf.keras.backend.clear_session()
# create the model
model = Sequential([
Dense(64, activation='relu', input_shape=[num_features]),
Dense(64, activation='relu'),
Dense(1)
])
# compile
rmse = tf.keras.metrics.RootMeanSquaredError()
optimizer = tf.keras.optimizers.RMSprop(0.001)
model.compile(loss='mse',
optimizer=optimizer,
metrics=[rmse, 'mae', 'mse'])
# train
tf.keras.backend.clear_session()
n = 4
early_stop = tf.keras.callbacks.EarlyStopping(
monitor='val_loss', patience=n)
car_history = model.fit(train_cars, epochs=100,
validation_data=test_cars,
callbacks=[early_stop])
# + [markdown] id="LQbREWjoyzc-" colab_type="text"
# ## Inspect the Model
# + id="XiFIJi0cyzll" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="07cf4bd1-27b6-4e9d-e83b-a81fa79559a9"
model.summary()
# + [markdown] id="joGMOHSXEmXh" colab_type="text"
# ## Visualize Training
# + id="C05oagoDEmga" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 886} outputId="31c332a8-d2dd-469d-cd31-bcdadf285136"
hist = pd.DataFrame(car_history.history)
hist['epoch'] = car_history.epoch
train_limit, test_limit = 10, 100
plot_history(history, train_limit, test_limit)
# + [markdown] id="4wpHwumxGrU8" colab_type="text"
# ## Generalize on Test Data
# + id="E11l_ablGsAk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="38f52d92-4838-4058-d695-3dc84c66dc3a"
loss, rmse, mae, mse = model.evaluate(test_cars, verbose=2)
print ()
print('"Testing set Mean Abs Error: {:5.2f} MPG'.format(mae))
# + [markdown] id="DhMF55-uG3nM" colab_type="text"
# ## Make Predictions
# + id="ip7M-YQ1G30K" colab_type="code" colab={}
predictions = model.predict(test_cars)
# + [markdown] id="Hc9brgz5HCjN" colab_type="text"
# ## Visualize Predictions
# + id="spb90N-1HCsF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 278} outputId="eb166990-ab4a-4591-d6be-82bfd922b393"
fig, ax = plt.subplots()
ax.scatter(y_test, predictions)
ax.plot([y_test.min(), y_test.max()],\
[y_test.min(), y_test.max()], 'k--', lw=4)
ax.set_xlabel('Measured')
ax.set_ylabel('Predicted')
plt.show()
# + [markdown] id="PSKIfgtPNZNZ" colab_type="text"
# # Read from UCI Irvine Machine Learning Repository
#
# We can also read car data directly from the UCI Irvine Machine Learning Repository.
# + id="yis3wFh5NZa_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} outputId="5c912952-f90d-438c-8510-b7f3f7daae7b"
dataset_path = tf.keras.utils.get_file("auto-mpg.data", "http://archive.ics.uci.\
edu/ml/machine-learning-databases/auto-mpg/auto-mpg.data")
dataset_path
# + [markdown] id="f1YifGCONsf3" colab_type="text"
# # Get Data into Pandas
# + id="N5w6YKACNsql" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 197} outputId="3c72e143-c848-4019-f76c-b2804427891c"
column_names = ['MPG','Cylinders','Displacement','Horsepower','Weight',
'Acceleration', 'Model Year', 'Origin']
uci_cars = pd.read_csv(dataset_path, names=column_names,
na_values = '?', comment='\t',
sep=' ', skipinitialspace=True)
uci_cars.head()
| class/lecture6/ch06.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.010474, "end_time": "2022-01-20T07:03:15.569876", "exception": false, "start_time": "2022-01-20T07:03:15.559402", "status": "completed"} tags=[]
# # Deep Neural Decision Forests
#
# Even though deep learning has attained trendendous success on data domains such as images, audio and texts.
# GDBT still rule the domain of tabular data.
#
# In this note we will discuss [Deep Neural Decision Forests](https://ieeexplore.ieee.org/document/7410529) for tabular deep learning
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 7.035528, "end_time": "2022-01-20T07:03:22.615390", "exception": false, "start_time": "2022-01-20T07:03:15.579862", "status": "completed"} tags=[]
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers as L
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler, OneHotEncoder
from sklearn.pipeline import Pipeline
from sklearn.model_selection import StratifiedKFold
import joblib
# + [markdown] papermill={"duration": 0.009059, "end_time": "2022-01-20T07:03:22.634086", "exception": false, "start_time": "2022-01-20T07:03:22.625027", "status": "completed"} tags=[]
# # Data
# + papermill={"duration": 0.36319, "end_time": "2022-01-20T07:03:23.006528", "exception": false, "start_time": "2022-01-20T07:03:22.643338", "status": "completed"} tags=[]
data = pd.read_csv('../input/song-popularity-prediction/train.csv')
print(data.shape)
data.head()
# + papermill={"duration": 0.07562, "end_time": "2022-01-20T07:03:23.092514", "exception": false, "start_time": "2022-01-20T07:03:23.016894", "status": "completed"} tags=[]
test = pd.read_csv('../input/song-popularity-prediction/test.csv')
X_test = test.drop(['id'], axis=1)
# + papermill={"duration": 0.021314, "end_time": "2022-01-20T07:03:23.124207", "exception": false, "start_time": "2022-01-20T07:03:23.102893", "status": "completed"} tags=[]
X = data.drop(['id', 'song_popularity'], axis=1)
y = data['song_popularity']
# + papermill={"duration": 0.017251, "end_time": "2022-01-20T07:03:23.151480", "exception": false, "start_time": "2022-01-20T07:03:23.134229", "status": "completed"} tags=[]
skf = StratifiedKFold(n_splits=5, shuffle=True, random_state=0)
# + [markdown] papermill={"duration": 0.009943, "end_time": "2022-01-20T07:03:23.171506", "exception": false, "start_time": "2022-01-20T07:03:23.161563", "status": "completed"} tags=[]
# # Model
# + papermill={"duration": 0.026675, "end_time": "2022-01-20T07:03:23.208380", "exception": false, "start_time": "2022-01-20T07:03:23.181705", "status": "completed"} tags=[]
class NeuralDecisionTree(keras.Model):
def __init__(self, depth, num_features, used_features_rate, num_classes):
super(NeuralDecisionTree, self).__init__()
self.depth = depth
self.num_leaves = 2 ** depth
self.num_classes = num_classes
num_used_features = int(num_features * used_features_rate)
one_hot = np.eye(num_features)
sampled_feature_indicies = np.random.choice(
np.arange(num_features), num_used_features, replace=False
)
self.used_features_mask = one_hot[sampled_feature_indicies]
self.pi = tf.Variable(
initial_value=tf.random_normal_initializer()(
shape=[self.num_leaves, self.num_classes]
),
dtype="float32",
trainable=True,
)
self.decision_fn = L.Dense(
units=self.num_leaves, activation="sigmoid", name="decision"
)
def call(self, features):
batch_size = tf.shape(features)[0]
features = tf.matmul(
features, self.used_features_mask, transpose_b=True
)
decisions = tf.expand_dims(
self.decision_fn(features), axis=2
)
decisions = L.concatenate(
[decisions, 1 - decisions], axis=2
)
mu = tf.ones([batch_size, 1, 1])
begin_idx = 1
end_idx = 2
for level in range(self.depth):
mu = tf.reshape(mu, [batch_size, -1, 1])
mu = tf.tile(mu, (1, 1, 2))
level_decisions = decisions[
:, begin_idx:end_idx, :
]
mu = mu * level_decisions
begin_idx = end_idx
end_idx = begin_idx + 2 ** (level + 1)
mu = tf.reshape(mu, [batch_size, self.num_leaves])
probabilities = keras.activations.softmax(self.pi)
outputs = tf.matmul(mu, probabilities)
return outputs
class NeuralDecisionForest(keras.Model):
def __init__(self, num_trees, depth, num_features, used_features_rate, num_classes):
super(NeuralDecisionForest, self).__init__()
self.ensemble = []
self.num_classes = num_classes
for _ in range(num_trees):
self.ensemble.append(
NeuralDecisionTree(depth, num_features, used_features_rate, num_classes)
)
def call(self, inputs):
batch_size = tf.shape(inputs)[0]
outputs = tf.zeros([batch_size, self.num_classes])
for tree in self.ensemble:
outputs += tree(inputs)
outputs /= len(self.ensemble)
return outputs
# + papermill={"duration": 0.016856, "end_time": "2022-01-20T07:03:23.235352", "exception": false, "start_time": "2022-01-20T07:03:23.218496", "status": "completed"} tags=[]
get_cat_pipeline = lambda: Pipeline([
('imputer', SimpleImputer(strategy='most_frequent')),
('encoder', OneHotEncoder(sparse=False))
])
get_num_pipeline = lambda: Pipeline([
('imputer', SimpleImputer(strategy='median')),
('scaler', StandardScaler())
])
# + papermill={"duration": 0.018558, "end_time": "2022-01-20T07:03:23.263992", "exception": false, "start_time": "2022-01-20T07:03:23.245434", "status": "completed"} tags=[]
class model_config:
NUMERIC_FEATURE_NAMES=[
'song_duration_ms', 'acousticness', 'danceability', 'energy', 'instrumentalness', 'liveness', 'loudness',
'speechiness', 'tempo', 'audio_valence'
]
CATEGORICAL_FEATURE_NAMES=[
'key','audio_mode','time_signature'
]
MAX_EPOCHS = 250
get_callbacks = lambda : [
keras.callbacks.EarlyStopping(min_delta=1e-4, patience=10, verbose=1, restore_best_weights=True),
keras.callbacks.ReduceLROnPlateau(patience=3, verbose=1)
]
# + [markdown] papermill={"duration": 0.009706, "end_time": "2022-01-20T07:03:23.283864", "exception": false, "start_time": "2022-01-20T07:03:23.274158", "status": "completed"} tags=[]
# # Training
# + _kg_hide-output=true papermill={"duration": 794.131461, "end_time": "2022-01-20T07:16:37.425268", "exception": false, "start_time": "2022-01-20T07:03:23.293807", "status": "completed"} tags=[]
preds_tree = []
preds_forest = []
for fold, (train_index, valid_index) in enumerate(skf.split(X, y)):
X_train, X_valid = X.iloc[train_index], X.iloc[valid_index]
y_train, y_valid = y.iloc[train_index], y.iloc[valid_index]
num_pipeline = get_num_pipeline().fit(X_train[model_config.NUMERIC_FEATURE_NAMES])
cat_pipeline = get_cat_pipeline().fit(X_train[model_config.CATEGORICAL_FEATURE_NAMES])
X_train = np.hstack((
num_pipeline.transform(X_train[model_config.NUMERIC_FEATURE_NAMES]),
cat_pipeline.transform(X_train[model_config.CATEGORICAL_FEATURE_NAMES])
))
X_valid = np.hstack((
num_pipeline.transform(X_valid[model_config.NUMERIC_FEATURE_NAMES]),
cat_pipeline.transform(X_valid[model_config.CATEGORICAL_FEATURE_NAMES])
))
X_test_ = np.hstack((
num_pipeline.transform(X_test[model_config.NUMERIC_FEATURE_NAMES]),
cat_pipeline.transform(X_test[model_config.CATEGORICAL_FEATURE_NAMES])
))
neural_decsion_tree = NeuralDecisionTree(
depth=5, num_features=X_train.shape[1], used_features_rate=1.0, num_classes=2
)
neural_decsion_tree.compile(
loss="sparse_categorical_crossentropy", optimizer="adam", metrics=['accuracy']
)
neural_decsion_tree.fit(
X_train, y_train, validation_data=(X_valid, y_valid), callbacks=get_callbacks(),
epochs=MAX_EPOCHS
)
preds_tree.append(neural_decsion_tree.predict(X_test_))
neural_decsion_forest = NeuralDecisionForest(
num_trees=10, depth=5, num_features=X_train.shape[1], used_features_rate=0.8, num_classes=2
)
neural_decsion_forest.compile(
loss="sparse_categorical_crossentropy", optimizer="adam", metrics=['accuracy']
)
neural_decsion_forest.fit(
X_train, y_train, validation_data=(X_valid, y_valid), callbacks=get_callbacks(),
epochs=MAX_EPOCHS
)
preds_forest.append(neural_decsion_forest.predict(X_test_))
# + [markdown] papermill={"duration": 2.936944, "end_time": "2022-01-20T07:16:43.245856", "exception": false, "start_time": "2022-01-20T07:16:40.308912", "status": "completed"} tags=[]
# # Submissions
# + papermill={"duration": 2.989026, "end_time": "2022-01-20T07:16:49.108373", "exception": false, "start_time": "2022-01-20T07:16:46.119347", "status": "completed"} tags=[]
submissions = pd.read_csv('../input/song-popularity-prediction/sample_submission.csv')
submissions['song_popularity'] = np.array([arr[:, 1] for arr in preds_tree]).mean(axis=0)
submissions.to_csv('preds_tree.csv', index=False)
submissions['song_popularity'] = np.array([arr[:, 1] for arr in preds_forest]).mean(axis=0)
submissions.to_csv('preds_forest.csv', index=False)
# + papermill={"duration": 2.874827, "end_time": "2022-01-20T07:16:54.944127", "exception": false, "start_time": "2022-01-20T07:16:52.069300", "status": "completed"} tags=[]
| Deep-Neural-Decision-Forests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
from IPython.display import HTML
HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide();
$('div.prompt').hide();
} else {
$('div.input').show();
$('div.prompt').show();
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
<form action="javascript:code_toggle()"><input type="submit" value="Code Toggle"></form>''')
# + [markdown] slideshow={"slide_type": "slide"}
# # Hackathon 3
#
# ** January 18-20, 2017 – Northwestern University **
#
# ## Install-A-Thon
#
# We’ll be having an Install-A-Thon (January 18), an eight-hour block in which a each person/group will go through the process of installing another group’s phase field software. If you successfully manage to install the software, you will be asked to use it to attempt to simulate a simple phase field problem. Note that the purpose of the exercise is just to see what is entailed in downloading, installing, and trying to use the code – whether you manage to finish the simulation or not is not so important. The next day, each group will report on their experiences, in terms of ease or difficulty, platform compatibility, available documentation, installation of library dependencies, learning curve to using the software, etc. Please add links to data, images and code in the Etherpad below.
#
# ## Benchmark Presentations
#
# We will have a section in which each group presents their results using the first set of benchmark problems (January 19) on spinodal decomposition and Ostwald ripening (see the website, https://pages.nist.gov/chimad-phase-field/, and the paper, http://dx.doi.org/10.1016/j.commatsci.2016.09.022, for more information). Please include as much metadata as possible, such as the computing platform, maximum memory usage, and runtime to whatever time you choose to halt the simulation (please include details regarding that criterion). Please generate microstructure images at the specified times and total free energy plots for most (hopefully all) of the computational domains and boundary conditions. We would like you to contribute your simulation data, input files, and results to ChiMaD website. We encourage you to brainstorm ways of using the benchmark problems that are unique to your own software and present that information if so. Please place data, images and code in a repository and add the link to the Etherpad below. Please add presentation slides to [Slideshare](http://www.slideshare.net) or [Speakerdeck](https://speakerdeck.com) and add the link in the Etherpad.
#
# ## Method of Manufactured Solutions Tutorial
#
# The workshop will end with a tutorial and a training section on using the Method of Manufactured Solutions (MMS) (January 20), where the attendees will have a chance to try their – or anybody else’s – software on an MMS problem.
#
# ## Etherpad
#
# Please add links to any repositories in the Etherpad below – https://etherpad.net/p/chimad-phase-field-hackathon-3
# +
from IPython.display import IFrame
IFrame("https://etherpad.net/p/chimad-phase-field-hackathon-3?showControls=true&showChat=true&showLineNumbers=true&useMonospaceFont=false",
width=700, height=800)
| hackathon3/index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.common.by import By
from selenium.webdriver.support.ui import Select
import time
import pandas as pd
from bs4 import BeautifulSoup
df = pd.read_csv('href_list.csv')
lst = df["0"]
#Wir starten den Browser auf
driver = webdriver.Firefox()
#Und nun sagen wir dem Browser, welche Seite er besuchen sollte.
count = 0
for url in lst:
driver.get(url)
time.sleep(3)
page = driver.page_source.encode('utf-8')
file = open("pages2/"+str(count)+".htm", "wb+")
file.write(page)
file.close()
count += 1
| 10 Selenium/03 Revisit new url .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # _Crash course_ de _conda_
# + [markdown] slideshow={"slide_type": "slide"}
# ## Objetivos
#
# - Aprender a utilizar o _conda_ para gerenciar a instalação de pacotes, dependências e ambientes;
# - Entender conceitos de distribuição, módulo e repositório;
# - Compreender comandos do _conda_ e praticá-los em terminal;
# + [markdown] slideshow={"slide_type": "slide"}
# ## Pré-requisitos
#
# - [Instalação](https://docs.anaconda.com/anaconda/install/) da distribuição Anaconda (recomenda-se Python 3.x);
# - Acesso simples a um terminal ou ao _Anaconda Prompt_;
# + [markdown] slideshow={"slide_type": "slide"}
# ## Introdução
#
# - _conda_ é um gerenciador de pacotes, dependências e ambientes para múltiplas linguagens;
#
# - Pacote de código aberto executável em Windows, macOS e Linux;
#
# > Objetivo: fornecer suporte à rápida instalação, desinstalação e atualização de pacotes
#
# - Hoje, ~ 7500 pacotes no [repo.anaconda.com](http://repo.anaconda.com)!
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Benefícios do _conda_ para cientistas de dados
#
# - Pacotes pré-construídos: evitam compiladores ou configurações específicas
#
# - Gestão: instalações mais difíceis com apenas um passo;
#
# - Reprodutibilidade: Permite que você forneça seu ambiente a outras pessoas em diferentes plataformas;
# + [markdown] slideshow={"slide_type": "slide"}
# ## Conceitos fundamentais
#
# - Anaconda: distribuição de código aberto, alto desempenho e otimizada para Python e R.
#
# - Anaconda Cloud: repositório de pacotes hospedado na web (nuvem)
#
# - Anaconda Navigator: interface gráfica incluída na distribuição para fácil gestão de pacotes, ambientes e canais.
# + [markdown] slideshow={"slide_type": "subslide"}
# - Canal: local dos repositórios onde o _conda_ procura por pacotes. Pode ser um repositório público, na web, ou privado, dentro da universidade, em uma empresa, na sua casa, etc.
# + [markdown] slideshow={"slide_type": "subslide"}
# - _conda_: gerenciador de pacotes e ambientes que vem incluído na distribuição.
# + [markdown] slideshow={"slide_type": "subslide"}
# - _conda environment_ (Ambiente): diretório que contém uma coleção específica de pacotes e dependências que pode ser administrado separadamente. Por exemplo, é possível manter um ambiente Python 2 e Python 3 totalmente isolados sem que um interfira em outro.
# + [markdown] slideshow={"slide_type": "subslide"}
# - _conda package_ (Pacote): arquivo comprimido que contém todos os elementos necessários para o funcionamento de um software: bibliotecas, módulos, executáveis e componentes.
# + [markdown] slideshow={"slide_type": "subslide"}
# - _conda repository_ (Repositório, ou _repo_): o repositório em nuvem mantido pela Anaconda.
# + [markdown] slideshow={"slide_type": "subslide"}
# - Miniconda: é uma versão menor da distribuição que inclui apenas pacotes essenciais, tais como `conda`, `pip`, `zlib` e outros considerados básicos. Pode ser expandido pela instalação de pacotes adicionais.
# + [markdown] slideshow={"slide_type": "subslide"}
# - _noarch package_ (Pacote independente de arquitetura): pacote que não contém nada específico à arquitetura de um sistema e que pode ser instalado em qualquer plataforma. `noarch` constitui um subdiretório em um canal.
# + [markdown] slideshow={"slide_type": "subslide"}
# - Repositório: qualquer local onde ativos de software são armazenados e podem ser baixados ou recuperados para instalação e uso em computadores.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Comandos fundamentais
#
# Ver [[Conda Cheat Sheet]](https://docs.conda.io/projects/conda/en/latest/_downloads/843d9e0198f2a193a3484886fa28163c/conda-cheatsheet.pdf)
#
# Aqui, dividiremos os comandos nos seguintes grupos:
#
# 1. informação e atualização
# 2. ambientes
# 3. pacotes e canais
# 4. adicionais
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Comandos para manutenção e atualização
#
# |comando|o que faz?|
# |---|---|
# |`conda --version` ou `conda -V`|verifica se conda está instalado |
# |`conda info`|verifica instalação e versão do conda|
# |`conda update -n base conda`|atualiza o gerenciador para a versão atual|
# |`conda update conda`|idem|
# |`conda update anaconda`|atualiza todos os pacotes da distribuição para versões estáveis|
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Comandos para trabalhar com ambientes
#
# |comando|o que faz?|
# |---|---|
# |`conda create --name AMB python=3.x "PKG1>v.s" PKG2`|cria novo ambiente com nome "AMB" para funcionar com a versão Python 3.x e instala neste ambiente os pacotes PKG1 e PKG2, sendo o primeiro na versão específica "v.s" e o outro a estável mais atual|
# + [markdown] slideshow={"slide_type": "subslide"}
# |comando|o que faz?|
# |---|---|
# |`conda activate AMB`|ativa o ambiente de nome AMB|
# |`conda activate /caminho/para/amb`|ativa um ambiente dado seu local|
# |`conda deactivate`|desativa o ambiente ativo|
# + [markdown] slideshow={"slide_type": "subslide"}
# |comando|o que faz?|
# |---|---|
# |`conda list`|lista todos os pacotes do ambiente ativo|
# |`conda list --name AMB`|lista todos os pacotes do ambiente AMB|
# + [markdown] slideshow={"slide_type": "subslide"}
# |comando|o que faz?|
# |---|---|
# |`conda remove --name AMB --all`|deleta todo o ambiente AMB|
# |`conda create --clone AMB --name NAMB`|faz um clone NAMB de AMB|
# + [markdown] slideshow={"slide_type": "subslide"}
# |comando|o que faz?|
# |---|---|
# |`conda env export --name AMB > amb.yml`|exporta configurações de AMB em um arquivo YAML|
# |`conda env create --file amb.yml`|cria AMB a partir de configurações contidas em um arquivo YAML|
# + [markdown] slideshow={"slide_type": "subslide"}
# > YAML (acrônimo para "YAML Ain't Markup Language") é uma linguagem de serialização de dados legível por humanos comumente usada para organizar arquivos de configuração. É utilizada em múltiplas linguagens. Veja [[YAML]](https://yaml.org).
# + [markdown] slideshow={"slide_type": "subslide"}
# >`conda activate` e `conda deactivate` somente funcionam a partir das versões 4.6 do `conda`. Para versões anteriores, no Windows usa-se `activate`/`deactivate` e no macOS, usa-se `source activate`/`source deactivate`.
#
# + [markdown] slideshow={"slide_type": "subslide"} tags=[]
# ### Comandos para trabalhar com pacotes e canais
#
# |comando|o que faz?|
# |---|---|
# |`conda search PCT=2.8 "PCT [version='>=2.8,<3.2']"`|procura pelo pacote PCT nos canais configurados cuja versão esteja no intervalo 2.8 <= v < 3.2|
# |`conda install PCT`|instala o pacote PCT, se disponível|
# |`conda install -c CH PCT`|instala o pacote AMB a partir do canal CH|
# |`conda install PCT==4.1.0`|instala o PCT com a versão especificada (4.1.0)|
# |`conda install "PCT[version='3.1.0\|3.1.1']"`|instala pacote com uma das versões especificadas (OU)|
# |`conda install "PCT>3.1,<3.5" `|instala uma das das versões do pacote especificadas (E)|
# + [markdown] slideshow={"slide_type": "subslide"} tags=[]
# ### Comandos adicionais
#
# |comando|o que faz?|
# |---|---|
# |`conda search AMB --info`|fornece informação detalhada sobre o pacote AMB|
# |`conda clean --all`|remove pacotes inutilizados|
# |`conda uninstall PCT --name AMB`|remove o pacote PCT do ambiente AMB|
# |`conda update --all --name AMB`|atualiza todos os pacotes do ambiente AMB|
# |`conda install --yes PCT1 PCT2`|instala pacotes sem exigir prompt do usuário|
# |`conda -h`|para obter ajuda sobre os comandos disponíveis do gerenciador|
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Exemplos
#
# - Criar ambiente chamado "dataScience" com versão Python 3.8 contendo os pacotes numpy, versão 1.19.1, e pandas, mais atual no repositório Anaconda.
#
# ```bash
# conda create --name dataScience python=3.8 numpy=1.19.1 pandas
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# - Alternar entre ambientes
#
# Abaixo, vamos reproduzir a mudança de um ambiente para outro em um Z Shell apontando para a pasta _ICD_ e mostrar que o pacote `scipy` está instalado em um ambiente, mas não em outro.
#
#
# ```bash
# # no ambiente 'base', procuramos pelo pacote 'scipy'
# (base) gustavo@GloryCrown ICD % conda list scipy
# ```
# ```
# # packages in environment at /Users/gustavo/opt/anaconda3:
# #
# # Name Version Build Channel
# scipy 1.6.2 py38hd5f7400_1
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# ```bash
# # ativamos um novo ambiente chamado 'lecture'
# (base) gustavo@GloryCrown ICD % conda activate lecture
# (lecture) gustavo@GloryCrown ICD %
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# ```bash
# # dentro do ambiente 'lecture', procuramos pelo pacote 'scipy'
# (lecture) gustavo@GloryCrown ICD % conda list scipy
# ```
#
# ```
# # packages in environment at /Users/gustavo/opt/anaconda3/envs/lecture:
# #
# # Name Version Build Channel
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# Nada é mostrado, significando que o pacote `scipy` está indisponível no ambiente `lecture`. Enfim, desativamos o ambiente ativo.
#
# ```bash
# # desativamos 'lecture' e voltamos para 'base'
# (lecture) gustavo@GloryCrown ICD % conda deactivate
# (base) gustavo@GloryCrown ICD %
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# - Criar arquivo YAML para construção de ambiente personalizado
#
# - Abra seu editor de texto preferido (sugestão: no Windows, `notepad++`; no Linux, `gedit`; no macOS, `TextEdit`);
# - Salve o arquivo como `icd.yml`;
# - Personalize o seu ambiente (use o modelo a seguir);
# - Use o comando `conda env create -f icd.yml` para criar o ambiente;
# - Verifique se o ambiente foi criado corretamente com `conda env list`. Você deve ver algo como:
# + [markdown] slideshow={"slide_type": "subslide"}
# ```
# (base) gustavo@GloryCrown ICD % conda env list
# # conda environments:
# #
# base * /Users/gustavo/opt/anaconda3
# icd /Users/gustavo/opt/anaconda3/envs/icd
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# ```yaml
# # Conteúdo do arquivo "icd.yaml"
# # para construir o ambiente 'icd'
# name: icd # nome do ambiente
# channels: # lista de canais a utilizar
# - defaults # canais padrão
# - conda-forge
# dependencies: # pacotes dependentes
# - numpy
# - scipy
# - sympy
# - matplotlib
# - pandas
# - seaborn
# ```
| _build/html/_sources/rise/02b-conda-rise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/OmdenaAI/RebootRx/blob/main/src/colab_example.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="hVLk2BI-ZkhP"
# # Using the local package in colab
# + id="kn_iB31bRCBX"
# get CUDA version (if using GPU)
# !nvidia-smi | grep -oP '(?<=CUDA Version: )(\d*\.\d*)'
# + id="NX4Ntq8WRIAQ"
from getpass import getpass
import os
os.environ['GIT_USER'] = input('Enter the user of your GitHub account: ')
os.environ['PASSWORD'] = getpass('Enter the password (or PAT if 2FA is enabled) of your GitHub account: ')
os.environ['GIT_AUTH'] = os.environ['GIT_USER'] + ':' + os.environ['PASSWORD']
print('Start installing git repo...')
# !pip install git+https://$GIT_AUTH@github.com/OmdenaAI/RebootRx.git@main > /dev/null
print('Package installed. Clear sensitive data...')
os.environ['PASSWORD'] = os.environ['GIT_AUTH'] = ""
# + id="Cz02JiZbGNJN" language="bash"
# pip install seqeval > /dev/null
# echo "Done!"
# + id="7cCEhOSD4UO3"
import os
import warnings
from datetime import datetime
from google.colab import drive
import matplotlib.pyplot as plt
import pandas as pd
from src.data_utils import ner
from src.model_utils import spacy_ner
from src import data_utils
from tqdm import tqdm
tqdm.pandas()
drive.mount('/content/drive')
ANNOTATIONS_PATH = "/content/drive/MyDrive/RebootRx/Official Folder of Reboot Rx Challenge/Task1 - Annotation/Final_datsets/RCT_Annotations_Final.csv"
MODEL_DIRPATH = "/content/drive/MyDrive/RebootRx/Official Folder of Reboot Rx Challenge/TASK3-MODELING/models/"
# + id="6lV2CKOAJ61_"
data = pd.read_csv(ANNOTATIONS_PATH)
data.info()
# + id="BrlUe-0mXkCM"
data = data_utils.labelbox(data)
data.head()
# + id="yW1vrA0OiUR4"
nlp = spacy_ner.create_blank_nlp(data["annotations"]) # specifying the tokenizer makes it much faster
# create a new column with zipped data and create TaggedCorpus object
data["tagged_corpus"] = pd.Series(zip(data["text"], data["annotations"]))
data["tagged_corpus"] = data["tagged_corpus"].progress_apply(
lambda x: ner.TaggedCorpus(text=x[0], annotations=x[1], tokenizer=nlp.tokenizer)
)
data.head()
# + [markdown] id="sLKzq_trYQXi"
# ## Modeling
#
# Use your model here!
# + id="aZaMSRTwjgT1"
_df = data.copy()
train = _df.sample(frac=0.9, random_state=42)
val = _df[~_df.index.isin(train.index)]
# + [markdown] id="Bxnre0xbQ6Ts"
# ## Evaluate
#
# Use [seqeval](https://github.com/chakki-works/seqeval) for evaluation
# + id="yz6nO7LbQ6Tt"
from seqeval.metrics import (
accuracy_score,
classification_report,
f1_score,
performance_measure,
precision_score,
recall_score,
)
# + id="0c_WIRZYQ6Tt"
# generate docs from validation set
docs_true = val.docs.to_list()
# generate iob list of tags from validation set
y_true = [ner.doc2ents(doc) for doc in docs_true]
# get the list of predictions from your model (`y_pred`) and run the evaluation below
# + id="uMeNB9PMQ6Tu"
print(classification_report(list(y_true), list(y_pred)))
# + [markdown] id="RNhYw6Paknvn"
# ### Log metrics, hyperparameters and models
# + id="Gc2Fv8FkAhBk"
from dagshub import dagshub_logger
metrics = {
"loss": training_loss[-1],
"loss_val": validation_loss[-1],
"accuracy": accuracy_score(y_true, y_pred),
"precision": precision_score(y_true, y_pred),
"recall": recall_score(y_true, y_pred),
"f1": f1_score(y_true, y_pred),
"class_report": classification_report(list(y_true), list(y_pred), output_dict=True),
**performance_measure(y_true, y_pred),
}
hp = {
"lib": "'spacy_ner'",
"n_epochs": n_epochs,
"dropout": dropout,
"n_train": len(train),
"n_test": len(val),
}
with dagshub_logger(
metrics_path=YOUR_METRICS_PATH,
hparams_path=YOUR_HYPERPARAMS_PATH,
) as logger:
# Metrics:
logger.log_metrics(metrics)
# Hyperparameters:
logger.log_hyperparams(hp)
# + id="_P2pTdRZQ6Tu"
if not os.path.exists(MODEL_PATH):
os.makedirs(MODEL_PATH)
model.save_to(MODEL_PATH) # this is model-variant
print("Saved model to", MODEL_PATH)
# # to load
# # model = spacy.load('model_name')
| src/colab_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
# This notebook demonstrates simble **Data Augumentation** combined with **ConvNet** and applied to [CIFAR-10](https://www.cs.toronto.edu/~kriz/cifar.html) dataset.
# **Contents**
#
# * [CIFAR-10 Dataset](#CIFAR-10-Dataset) - load and preprocess dataset
# * [Data Aug. Setup](#Data-Aug.-Setup) - setup data augumentation
# * [Data Aug. Model](#Data-Aug.-Model) - make and train model
# * [Baseline Model](#Baseline-Model) - w/o data aug for comparison
# # Imports
import numpy as np
import matplotlib.pyplot as plt
# Limit TensorFlow GPU memory usage
#
# +
import tensorflow as tf
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
with tf.Session(config=config):
pass # init sessin with allow_growth
# -
# # CIFAR-10 Dataset
# Load dataset and show example images
(x_train_raw, y_train_raw), (x_test_raw, y_test_raw) = tf.keras.datasets.cifar10.load_data()
class2txt = ['airplane', 'automobile', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck']
# Show example images
fig, axes = plt.subplots(nrows=1, ncols=6, figsize=[16, 9])
for i in range(len(axes)):
axes[i].set_title(class2txt[y_train_raw[i, 0]])
axes[i].imshow(x_train_raw[i])
# Normalize features
x_train = (x_train_raw - x_train_raw.mean()) / x_train_raw.std()
x_test = (x_test_raw - x_train_raw.mean()) / x_train_raw.std()
print('x_train.shape', x_train.shape)
print('x_test.shape', x_test.shape)
# One-hot encode labels
y_train = tf.keras.utils.to_categorical(y_train_raw, num_classes=10)
y_test = tf.keras.utils.to_categorical(y_test_raw, num_classes=10)
print('y_train.shape', y_train.shape)
print(y_train[:3])
# # Data Aug. Setup
# Setup Keras image data augumentation
# +
from tensorflow.keras.preprocessing.image import ImageDataGenerator
img_data_gen = ImageDataGenerator(
rotation_range=10, # random rotation degrees
width_shift_range=0.1, # random shift 10%
height_shift_range=0.1,
horizontal_flip=True)
# -
# Show a horse
plt.imshow(x_train_raw[7]);
# Show more horses
fig, axes = plt.subplots(nrows=1, ncols=6, figsize=[16, 9])
for i, x_horse in enumerate(img_data_gen.flow(x_train_raw[7:8], batch_size=1)):
axes[i].imshow(x_horse.astype(int)[0])
if i >= len(axes)-1:
break
# **Note:** Keras ImageDataGenerator seems to run on *single CPU thread*, which makes it very slow. On my PC single epoch approx 20s
# +
import time
ts = time.time()
for i, (x_batch, y_batch) in enumerate(img_data_gen.flow(x_train, y_train, batch_size=250)):
if i >= 250:
break
print(time.time() - ts)
# -
# # Data Aug. Model
# Simple ConvNet, no changes here
# +
from tensorflow.keras.layers import InputLayer, Conv2D, MaxPooling2D, Activation, Flatten, Dense, Dropout, BatchNormalization
model = tf.keras.Sequential()
model.add(InputLayer(input_shape=[32, 32, 3]))
model.add(Conv2D(filters=16, kernel_size=3, padding='same', activation='elu'))
model.add(MaxPooling2D(pool_size=[2,2], strides=[2, 2], padding='same'))
model.add(Conv2D(filters=32, kernel_size=3, padding='same', activation='elu'))
model.add(MaxPooling2D(pool_size=[2,2], strides=[2, 2], padding='same'))
model.add(Conv2D(filters=64, kernel_size=3, padding='same', activation='elu'))
model.add(MaxPooling2D(pool_size=[2,2], strides=[2, 2], padding='same'))
model.add(Flatten())
model.add(Dropout(0.2))
model.add(Dense(512, activation='elu'))
model.add(Dropout(0.2))
model.add(Dense(10, activation='softmax'))
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
model.summary()
# -
# Train model with data augumentation (this will take a while)
hist = model.fit_generator(generator=img_data_gen.flow(x_train, y_train, batch_size=250),
steps_per_epoch=len(x_train)/250, epochs=10, verbose=2)
loss, acc = model.evaluate(x_train, y_train, batch_size=250, verbose=0)
print(f'Accuracy on train set: {acc:.3f}')
loss, acc = model.evaluate(x_test, y_test, batch_size=250, verbose=0)
print(f'Accuracy on test set: {acc:.3f}')
# Note only .014 difference between train/test accuracy
# # Baseline Model
# Simple ConvNet
# +
from tensorflow.keras.layers import InputLayer, Conv2D, MaxPooling2D, Activation, Flatten, Dense, Dropout, BatchNormalization
model = tf.keras.Sequential()
model.add(InputLayer(input_shape=[32, 32, 3]))
model.add(Conv2D(filters=16, kernel_size=3, padding='same', activation='elu'))
model.add(MaxPooling2D(pool_size=[2,2], strides=[2, 2], padding='same'))
model.add(Conv2D(filters=32, kernel_size=3, padding='same', activation='elu'))
model.add(MaxPooling2D(pool_size=[2,2], strides=[2, 2], padding='same'))
model.add(Conv2D(filters=64, kernel_size=3, padding='same', activation='elu'))
model.add(MaxPooling2D(pool_size=[2,2], strides=[2, 2], padding='same'))
model.add(Flatten())
model.add(Dropout(0.2))
model.add(Dense(512, activation='elu'))
model.add(Dropout(0.2))
model.add(Dense(10, activation='softmax'))
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
model.summary()
# -
# Train model
hist = model.fit(x=x_train, y=y_train, batch_size=250, epochs=10,
validation_data=(x_test, y_test), verbose=2)
# Final results
loss, acc = model.evaluate(x_train, y_train, batch_size=250, verbose=0)
print(f'Accuracy on train set: {acc:.3f}')
loss, acc = model.evaluate(x_test, y_test, batch_size=250, verbose=0)
print(f'Accuracy on test set: {acc:.3f}')
# Much higher difference between train/test accuracy. Data augumetion clearly helps as regularizer.
#
# Over model test accuracy is better here, but neither model converged.
| KerasNN/1250_Aug_CIFAR10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import logging
import os
import matplotlib
import matplotlib.pyplot as plt
from matplotlib import offsetbox
import numpy as np
import pandas as pd
import seaborn as sns
from scipy import stats
from statsmodels.stats.multicomp import MultiComparison
import divisivenormalization.analysis as analysis
import divisivenormalization.utils as helpers
from divisivenormalization.data import Dataset, MonkeySubDataset
helpers.config_ipython()
logging.basicConfig(level=logging.INFO)
sns.set()
sns.set_style("ticks")
# adjust sns paper context rc parameters
font_size = 8
rc_dict = {
"font.size": font_size,
"axes.titlesize": font_size,
"axes.labelsize": font_size,
"xtick.labelsize": font_size,
"ytick.labelsize": font_size,
"legend.fontsize": font_size,
"figure.figsize": (helpers.cm2inch(8), helpers.cm2inch(8)),
"figure.dpi": 300,
"pdf.fonttype": 42,
"savefig.transparent": True,
"savefig.bbox_inches": "tight",
}
sns.set_context("paper", rc=rc_dict)
class args:
num_best = 10
num_val = 10
fname_best_csv = "df_best.csv"
fname_val_csv = "df_val.csv"
weights_path = "weights"
train_logs_path = "train_logs"
orientation_binsize = np.deg2rad(10)
stim_full_size = 140 # full size of stimulus w/o subsampling and cropping
stim_subsample = 2
oriented_threshold = 0.125
# -
# ### Load data
# +
results_df = pd.read_csv("results.csv")
# Save a simplified version of the csv file, sorted by validation set performance
df_plain = helpers.simplify_df(results_df)
df_plain.to_csv("results_plain.csv")
data_dict = Dataset.get_clean_data()
data = MonkeySubDataset(data_dict, seed=1000, train_frac=0.8, subsample=2, crop=30)
# -
# ### Get and save FEV performance on test set
# Use the 10 best models for analysis.
# Split the csv files accordingly. Also, extract some weights to be used for later analysis and save
# them as pickle. As this operation requires model loading, we do it only if it was not done before.
# +
try:
df_best = pd.read_csv(args.fname_best_csv)
logging.info("loaded data from " + args.fname_best_csv)
except FileNotFoundError:
df_best = df_plain[0 : args.num_best].copy()
fev_lst = []
for i in range(args.num_best):
run_no = df_best.iloc[i]["run_no"]
logging.info("load run no " + str(run_no))
model = helpers.load_dn_nonspecific_model(run_no, results_df, data, args.train_logs_path)
fev = model.evaluate_fev_testset()
fev_lst.append(fev)
feve = model.evaluate_fev_testset_per_neuron()
helpers.pkl_dump(feve, run_no, "feve.pkl", args.weights_path)
# get weights and normalization input
(
features_chanfirst,
p,
pooled,
readout_feat,
u,
v,
dn_exponent,
) = helpers.get_weights(model)
norm_input = analysis.norm_input(pooled, p)
helpers.pkl_dump(features_chanfirst, run_no, "features_chanfirst.pkl", args.weights_path)
helpers.pkl_dump(p, run_no, "p.pkl", args.weights_path)
helpers.pkl_dump(pooled, run_no, "pooled.pkl", args.weights_path)
helpers.pkl_dump(norm_input, run_no, "norm_input.pkl", args.weights_path)
helpers.pkl_dump(readout_feat, run_no, "readout_feat_w.pkl", args.weights_path)
helpers.pkl_dump(u, run_no, "u.pkl", args.weights_path)
helpers.pkl_dump(v, run_no, "v.pkl", args.weights_path)
helpers.pkl_dump(dn_exponent, run_no, "dn_exponent.pkl", args.weights_path)
df_best["fev"] = fev_lst
df_best.to_csv(args.fname_best_csv)
# +
fev = df_best.fev.values * 100
print("Mean FEV", fev.mean())
print("SEM", stats.sem(fev, ddof=1))
print("max FEV", fev.max())
print("FEV of model with max correlation on validation set", fev[0])
# -
# ### Similarly oriented features contribute stronger
# +
sim_input_lst, dissim_input_lst = [], []
for i in range(args.num_best):
run_no = df_best.iloc[i].run_no
features = helpers.pkl_load(run_no, "features_chanfirst.pkl", args.weights_path)
norm_input = helpers.pkl_load(run_no, "norm_input.pkl", args.weights_path)
angles = analysis.angles_circ_var(features, args.oriented_threshold)
angles_diff = analysis.angle_diff(angles)
unor_mask, sim_mask, dissim_mask = analysis.orientation_masks(angles_diff)
sim_input = np.sum(norm_input[sim_mask])
dissim_input = np.sum(norm_input[dissim_mask])
sim_input_lst.append(sim_input)
dissim_input_lst.append(dissim_input)
fractions = [s / d for s, d in zip(sim_input_lst, dissim_input_lst)]
fraction_err = stats.sem(fractions, ddof=0)
mean = np.average(fractions)
conf_int = analysis.compute_confidence_interval(fractions)
print("Similar norm. input divided by dissimilar input", np.round(mean, 2))
print("Confidence interval", np.round(conf_int, 2))
print("Plus/minus", np.round(mean - conf_int[0], 2))
print(stats.wilcoxon(sim_input_lst, dissim_input_lst))
print("Cohen's d", np.round(analysis.cohens_d(sim_input_lst, dissim_input_lst), 1))
| nonspecific_divisive_net/analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **[Geospatial Analysis Home Page](https://www.kaggle.com/learn/geospatial-analysis)**
#
# ---
#
# # Introduction
#
# In this tutorial, you'll explore several techniques for **proximity analysis**. In particular, you'll learn how to do such things as:
# - measure the distance between points on a map, and
# - select all points within some radius of a feature.
#
# <div class="alert alert-block alert-info">
# To create your own copy of this notebook (where you can run the code yourself!), <b><a href="https://www.kaggle.com/kernels/fork/5832141">click here</a></b>.
# </div>
# + _kg_hide-input=true
import folium
from folium import Marker, GeoJson
from folium.plugins import HeatMap
import pandas as pd
import geopandas as gpd
# Function for displaying the map
def embed_map(m, file_name):
from IPython.display import IFrame
m.save(file_name)
return IFrame(file_name, width='100%', height='500px')
# -
# You'll work with a dataset from the US Environmental Protection Agency (EPA) that tracks releases of toxic chemicals in Philadelphia, Pennsylvania, USA.
releases = gpd.read_file("../input/geospatial-learn-course-data/toxic_release_pennsylvania/toxic_release_pennsylvania/toxic_release_pennsylvania.shp")
releases.head()
# You'll also work with a dataset that contains readings from air quality monitoring stations in the same city.
stations = gpd.read_file("../input/geospatial-learn-course-data/PhillyHealth_Air_Monitoring_Stations/PhillyHealth_Air_Monitoring_Stations/PhillyHealth_Air_Monitoring_Stations.shp")
stations.head()
# # Measuring distance
#
# To measure distances between points from two different GeoDataFrames, we first have to make sure that they use the same coordinate reference system (CRS). Thankfully, this is the case here, where both use EPSG 2272.
print(stations.crs)
print(releases.crs)
# We also check the CRS to see which units it uses (meters, feet, or something else). In this case, EPSG 2272 has units of feet. (_If you like, you can check this [here](https://epsg.io/2272)._)
#
# It's relatively straightforward to compute distances in GeoPandas. The code cell below calculates the distance (in feet) between a relatively recent release incident in `recent_release` and every station in the `stations` GeoDataFrame.
# +
# Select one release incident in particular
recent_release = releases.iloc[360]
# Measure distance from release to each station
distances = stations.geometry.distance(recent_release.geometry)
distances
# -
# Using the calculated distances, we can obtain statistics like the mean distance to each station.
print('Mean distance to monitoring stations: {} feet'.format(distances.mean()))
# Or, we can get the closest monitoring station.
print('Closest monitoring station ({} feet):'.format(distances.min()))
print(stations.iloc[distances.idxmin()][["ADDRESS", "LATITUDE", "LONGITUDE"]])
# # Creating a buffer
#
# If we want to understand all points on a map that are some radius away from a point, the simplest way is to create a buffer.
#
# The code cell below creates a GeoSeries `two_mile_buffer` containing 12 different Polygon objects. Each polygon is a buffer of 2 miles (or, 2\*5280 feet) around a different air monitoring station.
two_mile_buffer = stations.geometry.buffer(2*5280)
two_mile_buffer.head()
# We use `folium.GeoJson()` to plot each polygon on a map. Note that since folium requires coordinates in latitude and longitude, we have to convert the CRS to EPSG 4326 before plotting.
# +
# Create map with release incidents and monitoring stations
m = folium.Map(location=[39.9526,-75.1652], zoom_start=11)
HeatMap(data=releases[['LATITUDE', 'LONGITUDE']], radius=15).add_to(m)
for idx, row in stations.iterrows():
Marker([row['LATITUDE'], row['LONGITUDE']]).add_to(m)
# Plot each polygon on the map
GeoJson(two_mile_buffer.to_crs(epsg=4326)).add_to(m)
# Show the map
embed_map(m, 'm1.html')
# -
# Now, to test if a toxic release occurred within 2 miles of **any** monitoring station, we could run 12 different tests for each polygon (to check individually if it contains the point).
#
# But a more efficient way is to first collapse all of the polygons into a **MultiPolygon** object. We do this with the `unary_union` attribute.
# +
# Turn group of polygons into single multipolygon
my_union = two_mile_buffer.geometry.unary_union
print('Type:', type(my_union))
# Show the MultiPolygon object
my_union
# -
# We use the `contains()` method to check if the multipolygon contains a point. We'll use the release incident from earlier in the tutorial, which we know is roughly 3781 feet to the closest monitoring station.
# The closest station is less than two miles away
my_union.contains(releases.iloc[360].geometry)
# But not all releases occured within two miles of an air monitoring station!
# The closest station is more than two miles away
my_union.contains(releases.iloc[358].geometry)
# # Your turn
#
# In the **[final exercise](https://www.kaggle.com/kernels/fork/5832147)**, you'll investigate hospital coverage in New York City.
# ---
# **[Geospatial Analysis Home Page](https://www.kaggle.com/learn/geospatial-analysis)**
#
#
#
#
#
# *Have questions or comments? Visit the [Learn Discussion forum](https://www.kaggle.com/learn-forum) to chat with other Learners.*
| Geo_spatial/proximity-analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Project 'hard problems': traveling salesman problem
#
# *Selected Topics in Mathematical Optimization: 2017-2018*
#
# **<NAME>** ([email](<EMAIL>))
#
# 
# YOUR NAME(S) HERE
# +
# everything you need for the TSP!
from tsp_utils import *
# %matplotlib inline
# -
# ## The traveling salesman problem
#
# > **Traveling salesman problem (TSP)**: Given a list of cities and the distances between each pair of cities, find
# the *tour* with the lowest possible total cost that visits each city exactly once and returns to
# the origin city.
#
# - $n$ cities $x_1,\ldots,x_n$.
# - Cost matrix $C[x_i, x_j]$ (possibly symmetric and/or triangle inequality).
# - Search space is all permutations of cities: size of $(n-1)!$.
# - Objective function: sum of costs of the paths.
# For this problem, the 'cities' are represented as points on the 2D plane. The $x,y$-coordinates are stored in the Numpy array `coordinates` and the distances between two cities are found in the Numpy array `distances`.
coordinates[:10]
n = len(coordinates)
print('There are {} cities.'.format(n))
distances[:10,:10]
# Cities are referred by their respective index. A tour is implemented as a list of the permutation of the indices of $n$ cities, e.g. `[0, 1, 2, ..., n-1]`. Note that the cost is invariant w.r.t. cyclic permutations, i.e. the cost is the same independent from which city the tour starts.
# +
tour = list(range(n))
tour
# -
# Some simple function are provided to compute the length of a given tour and to plot the cities and a tour.
# +
cost = compute_tour_cost(tour, distances)
print('Cost of tour is {:.3f}'.format(cost))
fig, ax = plt.subplots(figsize=(8, 8))
plot_cities(ax, coordinates, color=blue) # plot cities as a scatter plot
plot_tour(ax, tour, coordinates, distances, color=red, title=True) # add the tour
# -
# A tour can be saved and loaded in a JSON file. You have to hand in your best tour!
# +
save_tour('Data/my_tour.json', tour)
loaded_tour = load_tour('Data/my_tour.json')
# -
# !rm Data/my_tour.json
# ## Assignments
#
# Implement *two* heuristic algorithms for finding a low-cost tour:
#
# - write a report in the notebook discussing your strategy and the final results;
# - embed the code with sufficient documentation;
# - plot your final best tour in your notebook with the total cost.
# ## Project
# DESCRIBE YOUR STRATEGY
# +
# CELLS FOR CODE!
# TINY EXAMPLE FROM MICHIEL
import itertools as it
def yield_some_permutations(tour, mtry):
"""
Yields mtry permutations of a tour.
"""
count = 0
for perm in it.permutations(tour):
yield perm
count += 1
if count > mtry:
break
def lazy_brute_force(distances, mtry=10000):
n, _ = distances.shape
return min(yield_some_permutations(list(range(n)), mtry),
key=lambda t : compute_tour_cost(t, distances))
# +
# %timeit
best_tour = lazy_brute_force(distances, mtry=10000)
# +
cost = compute_tour_cost(best_tour, distances)
print('Cost of tour is {:.3f}'.format(cost)) # improvement!
fig, ax = plt.subplots(figsize=(8, 8))
plot_cities(ax, coordinates, color=blue) # plot cities as a scatter plot
plot_tour(ax, best_tour, coordinates, distances, color=orange, title=True) # add the tour
# -
# DESCRIBE YOUR RESULTS
| Chapters/11.ProjectTSP/Project_TSP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: multiverse
# language: python
# name: multiverse
# ---
# + [markdown] id="GXZho2Mw5ako"
# This notebook can be run in two ways:
# - Run all cells from beginning to end. However, this is a time-consuming process that will take about 10hrs. Note that the maximal runtime of a colab notebook is 12hrs. If a section is time-consuming, then the estimated time will be reported at the beginning of the section.
#
# - Run the different sections independently. The necessary files are provided in the output folder and are saved as pickle at the end of each section.
#
# If not downloaded to the your machine all the new generated pickles will be lost once the colab session is terminated.
# If you want to download a file to your machine you can use:
# ```python
# from google.colab import files
# files.download(str(output_path / 'file_name.p'))
# ```
# + [markdown] id="lHW7cYvz0a74"
# # 1. Setting up the enviroment
# + id="BzOXabS-wOg3"
# Cloning the git repo with the data structure and complentary files
# Note: to run the code in this notebook, you will have to accept that the
# code was not developed by google.
# !git clone https://github.com/Mind-the-Pineapple/into-the-multiverse/
# + id="YtCBYS1Q07ZK"
# Install necessary python dependencies
# ! pip install -r into-the-multiverse/requirements.txt
# + [markdown] id="D0PRP8tFe_su"
# Note: Remember to restart the runtime by clicking on the button above to have the same version of matplotlib, mpl_toolkits, numpy as specified in the requirement.txt file.
# + [markdown] id="j1tU-46g1f72"
# ## Download the Data
# + [markdown] id="Nr9IDDki8O7P"
# All the data used for this project is [public availiable](https://figshare.com/articles/Data_for_Conservative_and_disruptive_modes_of_adolescent_change_in_human_brain_functional_connectivity_/11551602) and consists of 520 scans from 298 healthy [Váša et. al, 2020](https://www.pnas.org/content/117/6/3248) individuals (age 14-26, mean age = 19.24, see for
# details)
# + id="kiURCoijKCCM"
from pathlib import Path
PROJECT_ROOT = Path.cwd()
data_path = PROJECT_ROOT / 'into-the-multiverse' /'data' / 'age'
output_path = PROJECT_ROOT / 'into-the-multiverse' / 'output'/ 'age'
if not data_path.is_dir():
data_path.mkdir(parents=True)
# + id="WVOQ5bQxoL-e"
# !wget -O into-the-multiverse/data/age/nspn.fmri.main.RData https://ndownloader.figshare.com/files/20958708
# + id="16eWfSDM1uSU"
# !wget -O into-the-multiverse/data/age/nspn.fmri.gsr.RData https://ndownloader.figshare.com/files/20958699
# + id="YC9v-PY82lun"
# !wget -O into-the-multiverse/data/age/nspn.fmri.lowmot.RData https://ndownloader.figshare.com/files/20958702
# + id="RPaw6pyIhy4g"
# !wget -O into-the-multiverse/data/age/nspn.fmri.general.vars.RData https://ndownloader.figshare.com/files/20819796
# + [markdown] id="FBm5C8HB3Okj"
# ## Define key variables
# + [markdown] id="BAHxS6LfG5ez"
# As mentioned this notebook was written so that every section could be run separately if needed. But in order to make this work, this section ([Define key variables](https://colab.research.google.com/drive/1fdEMsbZtQiTAwioeSn-JMLsJqcHqDoxj?authuser=2#)) needs to run and the variables that are going to be required saved into memory.
# + id="65_d0CqVZowv"
# Add the into-the-multiverse folder to the Python path. This allows the helperfunction
# to be used
import sys
sys.path.insert(1, 'into-the-multiverse')
# + id="Ra4NkHs2gjlH"
import pickle
import random
import pyreadr
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.colorbar
import bct
from scipy import stats
from sklearn.metrics.pairwise import cosine_similarity
from tqdm import tqdm
from helperfunctions import gateway_coef_sign, analysis_space
import warnings
warnings.filterwarnings("ignore")
# + id="g6pjQl5L6Itv"
def get_variables_of_interest():
# Set the random seed
#np.random.seed(2)
rng = np.random.default_rng(2)
random.seed(2)
# Define paths - REMOTE
PROJECT_ROOT = Path.cwd()
data_path = PROJECT_ROOT / 'into-the-multiverse' /'data' / 'age'
output_path = PROJECT_ROOT / 'into-the-multiverse' / 'output'/ 'age'
# Load data
data1 = pyreadr.read_r(str(data_path / 'nspn.fmri.main.RData'))
data3 = pyreadr.read_r(str(data_path / 'nspn.fmri.lowmot.RData'))
genVar = pyreadr.read_r(str(data_path / 'nspn.fmri.general.vars.RData'))
data2 = pyreadr.read_r(str(data_path / 'nspn.fmri.gsr.RData'))
DataNames=['nspn.fmri.main.RData','nspn.fmri.gsr.RData','nspn.fmri.lowmot.RData']
#Dictionary of 16 graph theory measures taken from the Brain Connectivity Toolbox
BCT_models = {
'degree': bct.degrees_und,
'strength': bct.strengths_und,
'betweennness centrality': bct.betweenness_bin,
'clustering (bin.)': bct.clustering_coef_bu,
'clustering (wei.)': bct.clustering_coef_wu,
'eigenvector centrality': bct.eigenvector_centrality_und,
'sugraph centrality': bct.subgraph_centrality,
'local efficiency' : bct.efficiency_bin,
'modularity (louvain)': bct.modularity_louvain_und,
'modularity (probtune)': bct.modularity_probtune_und_sign,
'participation coefficient': bct.participation_coef,
'module degree z-score': bct.module_degree_zscore,
'pagerank centrality': bct.pagerank_centrality,
'diversity coefficient': bct.diversity_coef_sign,
'gateway degree': gateway_coef_sign,
'k-core centrality': bct.kcoreness_centrality_bu,
}
#Get info about brain regions and find Yeo network IDs; useful later on for
# graph metrics that need community labels.
KeptIDs = np.asarray(genVar['hcp.keep.id'])
YeoIDs = np.asarray(genVar['yeo.id.subc'])
KeptYeoIDs = YeoIDs[KeptIDs-1][:,0,0]
# Define some images properites
n_regions = 346
subject_array = 520
#Get motion regression functional connectivity data and reshape into
# region x region x subject array
FC = np.asarray(data1['fc.main'])
MainNoNan = np.nan_to_num(FC,copy=True,nan=1.0)
MainNoNanReshape = np.reshape(MainNoNan, [n_regions,n_regions,subject_array],
order='F')
#Get global signal regression functional connectivity data and reshape into
# region x region x subject array
FC=np.asarray(data2['fc.gsr'])
GSRNoNan = np.nan_to_num(FC,copy=True,nan=1.0)
GSRNoNanReshape = np.reshape(GSRNoNan, [n_regions,n_regions,subject_array],
order='F')
#Read in subject IDs and age
IDMain=np.asarray(data1['id.main'])
ages=np.asarray(data1['age.main'])
#Find unique subject IDs and index of first instance and find FC data
# corresponding to these indices
IDs,IDIndexUnique = np.unique(IDMain,return_index=True)
MainNoNanReshapeUnique = MainNoNanReshape[:,:,IDIndexUnique]
GSRNoNanReshapeUnique = GSRNoNanReshape[:,:,IDIndexUnique]
AgesUnique = ages[IDIndexUnique]
# Number of randomly selected subjects to be used to define the low-dimensional
# space then split FC data and age data into two: 50 for defining space and
#remaining 248 for subsequent prediction
SpaceDefineIdx = 50
LockBoxDataIdx = 100
RandomIndexes = rng.choice(IDs.shape[0], size=IDs.shape[0], replace=False)
MainNoNanModelSpace = MainNoNanReshapeUnique[:,:,RandomIndexes[0:SpaceDefineIdx]]
MainNoNanLockBoxData = MainNoNanReshapeUnique[:, :, RandomIndexes[SpaceDefineIdx:LockBoxDataIdx]]
MainNoNanPrediction = MainNoNanReshapeUnique[:,:,RandomIndexes[LockBoxDataIdx:]]
GSRNoNanModelSpace = GSRNoNanReshapeUnique[:,:,RandomIndexes[0:SpaceDefineIdx]]
GSRNoNanLockBoxData = GSRNoNanReshapeUnique[:,:,RandomIndexes[SpaceDefineIdx:LockBoxDataIdx]]
GSRNoNanPrediction = GSRNoNanReshapeUnique[:,:,RandomIndexes[LockBoxDataIdx:]]
AgesModelSpace = AgesUnique[RandomIndexes[0:SpaceDefineIdx]]
AgesLockBoxData = AgesUnique[RandomIndexes[SpaceDefineIdx:LockBoxDataIdx]]
AgesPrediction = AgesUnique[RandomIndexes[LockBoxDataIdx:]]
return output_path, BCT_models, KeptYeoIDs, \
AgesPrediction, MainNoNanPrediction, GSRNoNanPrediction, \
AgesModelSpace, MainNoNanModelSpace, GSRNoNanModelSpace, \
AgesLockBoxData, MainNoNanLockBoxData, GSRNoNanLockBoxData, \
n_regions, subject_array
# + id="s1Gv0TyC25o8"
output_path, BCT_models, KeptYeoIDs, \
AgesPrediction, MainNoNanPrediction, GSRNoNanPrediction, \
AgesModelSpace, MainNoNanModelSpace, GSRNoNanModelSpace, \
AgesLockBoxData, MainNoNanLockBoxData, GSRNoNanLockBoxData, \
n_regions, subject_array = get_variables_of_interest()
# + [markdown] id="UbdR2G1WgRSM"
# Note: Some times running the cell above throws the following error:
# ```
# UnicodeDecodeError: 'utf-8' codec can't decode byte 0x80 in position 0: invalid start byte
# ```
# If this error shows up, restart the kernel and re-run all cells on this section
# + [markdown] id="B5mCueTin-12"
# # 2. Run the different analysis to bild the space
# + [markdown] id="IzK2ld8I-RYj"
# This section will perform the exhaustive evaluation of all 544 (2 different analysis, 17 sparsity thresholds and 16 nodal graph theoretical
# metrics) analysis approaches.
# + id="puxCTglyiKVF"
BCT_Run = {}
Sparsities_Run= {}
Data_Run = {}
GroupSummary = {}
thresholds = [0.4,0.3,0.25,0.2,0.175,0.150,0.125,0.1,0.09,0.08,
0.07,0.06,0.05,0.04,0.03,0.02,0.01]
preprocessings = ['MRS', 'GRS']
n_thr = len(thresholds)
n_pre = len(preprocessings)
n_BCT = len(BCT_models.keys())
Results = np.zeros(((n_thr * n_pre * n_BCT), n_regions))
ResultsIndVar = np.zeros(((n_thr * n_pre * n_BCT), 1225))
count=0
with tqdm(range(n_thr * n_pre * n_BCT)) as pbar:
for pre_idx, DataPreproc in enumerate(preprocessings): # data preprocessing
if DataPreproc == 'MRS':
TempData = MainNoNanModelSpace
TotalSubjects = TempData.shape[2]
elif DataPreproc == 'GRS':
TempData = GSRNoNanModelSpace
TotalSubjects = TempData.shape[2]
for thr_idx, TempThreshold in enumerate(thresholds): # FC threshold level
for BCT_Num in BCT_models.keys(): # Graph theory measure
TempResults = np.zeros((TotalSubjects,n_regions))
for SubNum in range(TotalSubjects):
x = bct.threshold_proportional(TempData[:,:,SubNum],
TempThreshold, copy=True)
ss = analysis_space(BCT_Num, BCT_models, x, KeptYeoIDs)
#For each subject for each approach keep the 346 regional values.
TempResults[SubNum, :] = ss
BCT_Run[count] = BCT_Num;
Sparsities_Run[count] = TempThreshold
Data_Run[count] = DataPreproc
GroupSummary[count] ='Mean'
# Build an array of similarities between subjects for each
# analysis approach
cos_sim = cosine_similarity(TempResults, TempResults)
Results[count, :] = np.mean(TempResults, axis=0)
ResultsIndVar[count, :] = cos_sim[np.triu_indices(TotalSubjects, k=1)].T
count += 1
pbar.update(1)
ModelsResults={"Results": Results,
"ResultsIndVar": ResultsIndVar,
"BCT": BCT_Run,
"Sparsities": Sparsities_Run,
"Data": Data_Run,
"SummaryStat": GroupSummary}
pickle.dump( ModelsResults, open(str(output_path / "ModelsResults.p"), "wb" ) )
# + [markdown] id="BxVWimqHnyM0"
# # 3. Building and analysing the low-dimensional space
# + [markdown] id="HUYIJG675JWn"
# ## Different embeddings
# + [markdown] id="LkVxxGt8-Ci1"
# This section will use five different embedding algorithms to produce a low-dimension space that then be used for the active learning.
# + id="3ArZnzq35WAO"
from sklearn import manifold, datasets
from sklearn.preprocessing import StandardScaler
from collections import OrderedDict
from functools import partial
from time import time
import pickle
from mpl_toolkits.mplot3d import Axes3D
from matplotlib.ticker import NullFormatter
import matplotlib.patches as mpatches
import matplotlib.lines as mlines
import matplotlib.pyplot as plt
from umap.umap_ import UMAP
import phate
from sklearn.decomposition import PCA
# + id="SnseTiW87I53"
# Load the previous results
ModelResults = pickle.load(open(str(output_path / "ModelsResults.p"), "rb" ) )
Results = ModelResults['ResultsIndVar']
BCT_Run = ModelResults['BCT']
Sparsities_Run = ModelResults['Sparsities']
Data_Run = ModelResults['Data']
preprocessings = ['MRS', 'GRS']
# + id="M6sfdK1TiiVA"
#Scale the data prior to dimensionality reduction
scaler = StandardScaler()
X = scaler.fit_transform(Results.T)
X = X.T
n_neighbors = 20
n_components = 2 #number of components requested. In this case for a 2D space.
#Define different dimensionality reduction techniques
methods = OrderedDict()
LLE = partial(manifold.LocallyLinearEmbedding,
n_neighbors, n_components, eigen_solver='dense')
methods['LLE'] = LLE(method='standard', random_state=0)
methods['SE'] = manifold.SpectralEmbedding(n_components=n_components,
n_neighbors=n_neighbors, random_state=0)
methods['t-SNE'] = manifold.TSNE(n_components=n_components, init='pca',
random_state=0)
methods['UMAP'] = UMAP(random_state=40, n_components=2, n_neighbors=200,
min_dist=.8)
methods['PHATE'] = phate.PHATE()
methods['PCA'] = PCA(n_components=2)
# + id="dPvvEjch7Qju"
markers = ["x","s","o","*","D","1","v","p","H","+","|","_","3","^","4","<","X"]
colourmaps = {"MRS":"Oranges","GRS":"Purples"}
BCT = np.array(list(BCT_Run.items()))[:,1]
Sparsities = np.array(list(Sparsities_Run.items()))[:,1]
Data = np.array(list(Data_Run.items()))[:,1]
# Reduced dimensions
data_reduced = {}
gsDE, axs = plt.subplots(3,2, figsize=(16,16), constrained_layout=True)
axs = axs.ravel()
#Perform embedding and plot the results (including info about the approach in the color/intensity and shape).
for idx_method, (label, method) in enumerate(methods.items()):
Y = method.fit_transform(X)
# Save the results
data_reduced[label] = Y
Lines={}
for preprocessing in preprocessings:
BCTTemp=BCT[Data==preprocessing]
SparsitiesTemp=Sparsities[Data==preprocessing]
YTemp=Y[Data==preprocessing,:]
for idx_bct, bct_model in enumerate(BCT_models):
axs[idx_method].scatter(YTemp[:,0][BCTTemp==bct_model],
YTemp[:,1][BCTTemp==bct_model],
c=SparsitiesTemp[BCTTemp==bct_model],
marker=markers[idx_bct],
cmap=colourmaps[preprocessing], s=80)
Lines[idx_bct] = mlines.Line2D([], [], color='black', linestyle='None',
marker=markers[idx_bct], markersize=10,
label=bct_model)
# For visualisation purposes show the y and x labels only ons specific plots
if idx_method % 2 == 0:
axs[idx_method].set_ylabel('Dimension 1',fontsize=20)
if (idx_method == 4) or (idx_method == 5):
axs[idx_method].set_xlabel('Dimension 2',fontsize=20)
axs[idx_method].set_title("%s " % (label),fontsize=20, fontweight="bold")
axs[idx_method].axis('tight')
axs[idx_method].tick_params(labelsize=15)
OrangePatch = mpatches.Patch(color='orange', label='Motion Regression')
PurplePatch = mpatches.Patch(color='purple', label='Global Signal Regression')
OrangePatch = mpatches.Patch(color='orange', label='motion regression')
PurplePatch = mpatches.Patch(color=[85/255, 3/255, 152/255], label='global signal regression')
IntensityPatch1 = mpatches.Patch(color=[0.1, 0.1, 0.1], label='threshold: 0.4',
alpha=1)
IntensityPatch2 = mpatches.Patch(color=[0.1, 0.1, 0.1], label='threshold: 0.1',
alpha=0.4)
IntensityPatch3 = mpatches.Patch(color=[0.1, 0.1, 0.1], label='threshold: 0.01',
alpha=0.1)
BlankLine=mlines.Line2D([], [], linestyle='None')
gsDE.legend(handles=[OrangePatch, PurplePatch,BlankLine,IntensityPatch1,
IntensityPatch2, IntensityPatch3,BlankLine,
Lines[0],Lines[1],Lines[2],Lines[3],Lines[4],Lines[5],
Lines[6],Lines[7],Lines[8],Lines[9],Lines[10],Lines[11],
Lines[12],Lines[13],Lines[14],Lines[15]],fontsize=15,
frameon=False,bbox_to_anchor=(1.25, .7))
gsDE.savefig(str(output_path / 'DifferentEmbeddings.png'), dpi=300, bbox_inches='tight')
gsDE.savefig(str(output_path / 'DifferentEmbeddings.svg'), format="svg", bbox_inches='tight')
gsDE.show()
# + id="52KI031j495K"
methods['MDS'] = manifold.MDS(n_components, max_iter=100, n_init=10,
random_state=21, metric=True)
# + id="Ggt7wfAfkAiD"
#Do the same as above but for MDS
Y = methods['MDS'].fit_transform(X)
data_reduced['MDS'] = Y
figMDS = plt.figure(constrained_layout=False, figsize=(21,15))
gsMDS = figMDS.add_gridspec(nrows=15, ncols=20)
axs = figMDS.add_subplot(gsMDS[:,0:15])
idx_method = 0
for preprocessing in preprocessings:
BCTTemp=BCT[Data==preprocessing]
SparsitiesTemp=Sparsities[Data==preprocessing]
YTemp=Y[Data==preprocessing,:]
Lines={}
for idx_bct, bct_model in enumerate(BCT_models):
axs.scatter(YTemp[:,0][BCTTemp==bct_model],
YTemp[:,1][BCTTemp==bct_model],
c=SparsitiesTemp[BCTTemp==bct_model],
marker=markers[idx_bct],
norm=matplotlib.colors.Normalize(
vmin=np.min(SparsitiesTemp[BCTTemp==bct_model]),
vmax=np.max(SparsitiesTemp[BCTTemp==bct_model])),
cmap=colourmaps[preprocessing], s=120)
Lines[idx_bct] = mlines.Line2D([], [], color='black', linestyle='None',
marker=markers[idx_bct], markersize=10,
label=bct_model)
axs.spines['top'].set_linewidth(1.5)
axs.spines['right'].set_linewidth(1.5)
axs.spines['bottom'].set_linewidth(1.5)
axs.spines['left'].set_linewidth(1.5)
axs.set_xlabel('Dimension 2',fontsize=20,fontweight="bold")
axs.set_ylabel('Dimension 1',fontsize=20,fontweight="bold")
axs.tick_params(labelsize=15)
axs.set_title('Multi-dimensional Scaling', fontsize=25,fontweight="bold")
OrangePatch = mpatches.Patch(color='orange', label='motion regression')
PurplePatch = mpatches.Patch(color=[85/255, 3/255, 152/255], label='global signal regression')
IntensityPatch1 = mpatches.Patch(color=[0.1, 0.1, 0.1], label='threshold: 0.4',
alpha=1)
IntensityPatch2 = mpatches.Patch(color=[0.1, 0.1, 0.1], label='threshold: 0.1',
alpha=0.4)
IntensityPatch3 = mpatches.Patch(color=[0.1, 0.1, 0.1], label='threshold: 0.01',
alpha=0.1)
BlankLine=mlines.Line2D([], [], linestyle='None')
figMDS.legend(handles=[OrangePatch, PurplePatch,BlankLine,IntensityPatch1,
IntensityPatch2, IntensityPatch3,BlankLine,
Lines[0],Lines[1],Lines[2],Lines[3],Lines[4],Lines[5],
Lines[6],Lines[7],Lines[8],Lines[9],Lines[10],Lines[11],
Lines[12],Lines[13],Lines[14],Lines[15]],fontsize=15,
frameon=False,bbox_to_anchor=(1.4, 0.8),bbox_transform=axs.transAxes)
figMDS.savefig(str(output_path / 'MDSSpace.png'), dpi=300)
figMDS.savefig(str(output_path /'MDSSpace.svg'), format="svg")
# + id="5TV38MzhNhkn"
# Save results form the embedding to be used in the remaining analysis
pickle.dump(data_reduced, open(str(output_path / "embeddings.p"), "wb" ) )
# + [markdown] id="Y19DU5UH_EWC"
# ## Analyse the neighbours
# + id="2x25diSb_IFr"
from helperfunctions import (get_models_neighbours, get_dissimilarity_n_neighbours,
get_null_distribution)
# + id="VOAQ8O518_cP"
N = 544
n_neighbors_step = 10
neighbours_orig, adj_array = get_models_neighbours(N, n_neighbors_step, X)
# + id="XyfPmC_wNumB"
neighbours_tsne, _ = get_models_neighbours(N, n_neighbors_step,
data_reduced['t-SNE'])
diss_tsne = get_dissimilarity_n_neighbours(neighbours_orig, neighbours_tsne)
del neighbours_tsne
# + id="16BQTmevzoue"
neighbours_lle, _ = get_models_neighbours(N, n_neighbors_step,
data_reduced['LLE'])
diss_lle = get_dissimilarity_n_neighbours(neighbours_orig,neighbours_lle)
del neighbours_lle
# + id="phtJnBWCz1l6"
neighbours_se, _ = get_models_neighbours(N, n_neighbors_step,
data_reduced['SE'])
diss_se = get_dissimilarity_n_neighbours(neighbours_orig,neighbours_se)
del neighbours_se
# + id="r92E3fm3z6HC"
neighbours_mds, _ = get_models_neighbours(N, n_neighbors_step,
data_reduced['MDS'])
diss_mds = get_dissimilarity_n_neighbours(neighbours_orig,neighbours_mds)
del neighbours_mds
# + id="jpUpaN2YS5Nh"
neighbours_pca, _ = get_models_neighbours(N, n_neighbors_step,
data_reduced['PCA'])
diss_pca = get_dissimilarity_n_neighbours(neighbours_orig, neighbours_pca)
del neighbours_pca
# + id="gaxUNFasas6R"
null_distribution = get_null_distribution(N, n_neighbors_step)
# + id="L5XUNH1My_cW"
fig, ax = plt.subplots(figsize=(8, 6))
n_neighbours = range(2, N, n_neighbors_step)
ax.plot(n_neighbours, diss_tsne, label='t-SNE', color='#1DACE8')
ax.plot(n_neighbours, diss_lle, label='LLE', color='#E5C4A1')
ax.plot(n_neighbours, diss_se, label='SE', color='#F24D29')
ax.plot(n_neighbours, diss_mds, label='MDS', color='#1C366B')
ax.plot(n_neighbours, diss_pca, label='PCA', color='r')
plt.plot(n_neighbours, null_distribution, label='random', c='grey')
plt.ylim([0,1])
plt.xlim([0,N])
plt.legend(frameon=False)
plt.xlabel('$k$ Nearest Neighbors')
plt.ylabel('Dissimilarity $\epsilon_k$')
plt.savefig(str(output_path / 'dissimilarity_all.svg'))
plt.savefig(str(output_path / 'dissimilarity_all.png'), dpi=300)
plt.show()
# + id="M4sodQ_wC4pY"
# Download file to computer
from google.colab import files
files.download(str(output_path / 'dissimilarity_all.svg'))
# + [markdown] id="l8yM3HTB58LA"
# # 4. Exhaustive Search
#
# Exhaustive search for SVR prediction of age, so we know what "ground truth" is.
#
# Note: This step is time consuming and might take about 4hrs hrs to run.
# + id="2QNBlUjI6Nwt"
from bayes_opt import BayesianOptimization, UtilityFunction
from sklearn.gaussian_process.kernels import Matern, WhiteKernel
from sklearn.neighbors import NearestNeighbors
from sklearn.gaussian_process import GaussianProcessRegressor
from matplotlib import cm
from mpl_toolkits.mplot3d import Axes3D
from tqdm import tqdm
import warnings
warnings.filterwarnings("ignore")
from helperfunctions import objective_func_reg
# + id="eRa95C7d8lVu"
output_path, BCT_models, KeptYeoIDs, \
AgesPrediction, MainNoNanPrediction, GSRNoNanPrediction, \
AgesModelSpace, MainNoNanModelSpace, GSRNoNanModelSpace, \
AgesLockBoxData, MainNoNanLockBoxData, GSRNoNanLockBoxData, \
n_regions, subject_array = get_variables_of_interest()
# + id="VLz4M0z4Ljj9"
# Load embedding results. This cell is only necessary if you are running this
# part of the analysis separatly.
ModelEmbeddings = pickle.load(open(str(output_path / "embeddings.p"), "rb" ) )
ModelEmbedding = ModelEmbeddings['MDS']
# + id="Ib3ZKhDMmPcU"
PredictedAcc = np.zeros((len(Data_Run)))
for i in tqdm(range(len(Data_Run))):
tempPredAcc = objective_func_reg(i, AgesPrediction, Sparsities_Run, Data_Run,
BCT_models, BCT_Run, KeptYeoIDs, MainNoNanPrediction,
GSRNoNanPrediction)
PredictedAcc[i] = tempPredAcc
#Display how predicted accuracy is distributed across the low-dimensional space
plt.scatter(ModelEmbedding[0: PredictedAcc.shape[0], 0],
ModelEmbedding[0: PredictedAcc.shape[0], 1],
c=PredictedAcc, cmap='bwr')
plt.colorbar()
# + id="-aVnBYeLmQxa"
# Dump accuracies
pickle.dump(PredictedAcc, open(str(output_path / 'predictedAcc.pckl'), 'wb'))
# + id="LfOFZKFyCiJU"
# Download file to computer
from google.colab import files
files.download(str(output_path / 'predictedAcc.pckl'))
# + [markdown] id="v7UX0xYsKAdl"
# # 5. Active Learning
# + id="Q8aTNtvW3zp-"
from itertools import product
import pickle
from matplotlib import cm
import bct
from mpl_toolkits.mplot3d import Axes3D
from sklearn.svm import SVR
from sklearn.model_selection import permutation_test_score
from sklearn.gaussian_process import GaussianProcessRegressor
from scipy.stats import spearmanr
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
from helperfunctions import (initialize_bo, run_bo, posterior,
posteriorOnlyModels, display_gp_mean_uncertainty,
plot_bo_estimated_space, plot_bo_evolution,
analysis_space, plot_bo_repetions)
# + id="Ih9A4B4CIyYE"
# Load embedding results. This cell is only necessary if you are running this
# part of the analysis separatly.
ModelEmbeddings = pickle.load(open(str(output_path / "embeddings.p"), "rb" ))
ModelEmbedding = ModelEmbeddings['MDS']
PredictedAcc = pickle.load(open(str(output_path / "predictedAcc.pckl"), "rb"))
ModelResults = pickle.load(open(str(output_path / "ModelsResults.p"), "rb" ))
Results = ModelResults['ResultsIndVar']
BCT_Run = ModelResults['BCT']
Sparsities_Run = ModelResults['Sparsities']
Data_Run = ModelResults['Data']
preprocessings = ['MRS', 'GRS']
model_config = {}
model_config['Sparsities_Run'] = Sparsities_Run
model_config['Data_Run'] = Data_Run
model_config['BCT_models'] = BCT_models
model_config['BCT_Run'] = BCT_Run
model_config['CommunityIDs'] = KeptYeoIDs
model_config['MainNoNanPrediction'] = MainNoNanPrediction
model_config['GSRNoNanPrediction'] = GSRNoNanPrediction
model_config['MainNoNanLockBox'] = MainNoNanLockBoxData
model_config['GSRNoNanLockBox'] = GSRNoNanLockBoxData
ClassOrRegression = 'Regression'
# + [markdown] id="A3pdTLOC_WU4"
# ## Exploratory analysis
#
# Note: This step takes about 30min.
# + id="l81tOZ9Z33WD"
kappa = 10
# Define settings for the analysis
kernel, optimizer, utility, init_points, n_iter, pbounds, nbrs, RandomSeed = \
initialize_bo(ModelEmbedding, kappa)
# Perform optimization. Given that the space is continuous and the analysis
# approaches are not, we penalize suggestions that are far from any actual
# analysis approaches. For these suggestions the registered value is set to the
# lowest value from the burn in. These points (BadIters) are only used
# during search but exluded when recalculating the GP regression after search.
BadIter = run_bo(optimizer, utility, init_points,
n_iter, pbounds, nbrs, RandomSeed,
ModelEmbedding, model_config,
AgesPrediction,
ClassOrRegression,
MultivariateUnivariate=True, verbose=False)
# + id="qSUlVAr3LQa2"
x_exploratory, y_exploratory, z_exploratory, x, y, gp, vmax, vmin = \
plot_bo_estimated_space(kappa, BadIter,
optimizer, pbounds,
ModelEmbedding, PredictedAcc,
kernel, output_path, ClassOrRegression)
# + id="zQOakRcjLTCa"
# Display the results of the active search and the evolution of the search
# after 5, 10,20, 30 and 50 iterations.
corr = plot_bo_evolution(kappa, x_exploratory, y_exploratory, z_exploratory, x, y, gp,
vmax, vmin, ModelEmbedding, PredictedAcc, output_path, ClassOrRegression)
# + id="vLDCqGX2eVVA"
print(f'Spearman correlation {corr}')
# + [markdown] id="7gchNrIvexHA"
# ## Exploitatory analysis
# + id="nVoIxA3173sM"
kappa = .1
# Define settins for the analysis
kernel, optimizer, utility, init_points, n_iter, pbounds, nbrs, RandomSeed = \
initialize_bo(ModelEmbedding, kappa)
# Perform optimization. Given that the space is continuous and the analysis
# approaches are not, we penalize suggestions that are far from any actual
# analysis approaches. For these suggestions the registered value is set to the
# lowest value from the burn in. These points (BadIters) are only used
# during search but exluded when recalculating the GP regression after search.
BadIter = run_bo(optimizer, utility, init_points,
n_iter, pbounds, nbrs, RandomSeed,
ModelEmbedding, model_config,
AgesPrediction,
ClassOrRegression,
MultivariateUnivariate=True, verbose=False)
# + id="rijtkCYjfGa7"
x_exploratory, y_exploratory, z_exploratory, x, y, gp, vmax, vmin = \
plot_bo_estimated_space(kappa, BadIter,
optimizer, pbounds,
ModelEmbedding, PredictedAcc,
kernel, output_path, ClassOrRegression)
# + id="g7DbkSuNfJjh"
# Display the results of the active search and the evolution of the search
# after 5, 10,20, 30 and 50 iterations.
plot_bo_evolution(kappa, x_exploratory, y_exploratory, z_exploratory, x, y, gp,
vmax, vmin, ModelEmbedding, PredictedAcc, output_path, ClassOrRegression)
# + id="Zyc9XH6ufdzV"
# Download file to computer
from google.colab import files
files.download(str(output_path / 'BOptEvolutionK10.svg'))
files.download(str(output_path / 'BOptEvolutionK0.1.svg'))
files.download(str(output_path / 'BOptAndTrueK0.1.svg'))
files.download(str(output_path / 'BOptAndTrueK10.svg'))
# + [markdown] id="S8prbJVG3Z6M"
# ## Repetitions
# + [markdown] id="QsttJ2FeDsac"
# This is time consuming step and will take about 4 hrs to run.
# + id="yCGuS3mFDNmA"
from sklearn.metrics import mean_absolute_error
from sklearn.model_selection import PredefinedSplit
n_repetitions = 20
kappa = 10
TotalRegions = 346
n_permutations = 1000
BestModelGPSpace = np.zeros(n_repetitions)
BestModelGPSpaceModIndex = np.zeros(n_repetitions)
BestModelEmpirical = np.zeros(n_repetitions)
BestModelEmpiricalModIndex = np.zeros(n_repetitions)
ModelActualAccuracyCorrelation = np.zeros(n_repetitions)
CVPValBestModels = np.zeros(n_repetitions)
perm_scores = np.zeros((n_repetitions, n_permutations))
cv_mae = np.zeros(n_repetitions)
maes = np.zeros(n_repetitions)
#predictions = np.zeros((n_repetitions, len(AgesLockBoxData)))
for DiffInit in range(n_repetitions):
# Define settings for the analysis
kernel, optimizer, utility, init_points, n_iter, pbounds, nbrs, RandomSeed = \
initialize_bo(ModelEmbedding, kappa, repetitions=True,
DiffInit=DiffInit)
# Run BO on the Prediction again
FailedIters = run_bo(optimizer, utility, init_points,
n_iter, pbounds, nbrs, RandomSeed,
ModelEmbedding, model_config,
AgesPrediction,
ClassOrRegression,
MultivariateUnivariate=True, repetitions=True,
verbose=False)
gp = GaussianProcessRegressor(kernel=kernel, normalize_y=True,
n_restarts_optimizer=10)
x_temp = np.array([[res["params"]["b1"]] for res in optimizer.res])
y_temp = np.array([[res["params"]["b2"]] for res in optimizer.res])
z_temp = np.array([res["target"] for res in optimizer.res])
x_obs = x_temp[FailedIters==0]
y_obs = y_temp[FailedIters==0]
z_obs = z_temp[FailedIters==0]
muModEmb, sigmaModEmb, gpModEmb = posteriorOnlyModels(gp, x_obs, y_obs, z_obs,
ModelEmbedding)
BestModelGPSpace[DiffInit] = muModEmb.max()
BestModelGPSpaceModIndex[DiffInit] = muModEmb.argmax()
BestModelEmpirical[DiffInit] = z_obs.max()
Model_coord = np.array([[x_obs[z_obs.argmax()][-1], y_obs[z_obs.argmax()][-1]]])
BestModelEmpiricalModIndex[DiffInit] = nbrs.kneighbors(Model_coord)[1][0][0]
ModelActualAccuracyCorrelation[DiffInit] = spearmanr(muModEmb, PredictedAcc)[0]
TempModelNum = muModEmb.argmax()
TempThreshold = Sparsities_Run[TempModelNum]
BCT_Num = BCT_Run[TempModelNum]
# Load the Lockbox data
Y = AgesLockBoxData
CommunityIDs = KeptYeoIDs
if Data_Run[TempModelNum] == 'MRS':
TempDataLockBox = MainNoNanLockBoxData
TempDataPredictions = MainNoNanPrediction
elif Data_Run[TempModelNum] == 'GRS':
TempDataLockBox = GSRNoNanLockBoxData
TempDataPredictions = MainNoNanPrediction
TotalSubjectslock = TempDataLockBox.shape[2]
TotalSubjectsPredictions = TempDataPredictions.shape[2]
TempResultsLockData = np.zeros([TotalSubjectslock, n_regions])
for SubNum in range(0, TotalSubjectslock):
# Lock data
x = bct.threshold_proportional(TempDataLockBox[:, :, SubNum],
TempThreshold, copy=True)
ss = analysis_space(BCT_Num, BCT_models, x, KeptYeoIDs)
TempResultsLockData[SubNum, :] = ss
TempPredictionsData = np.zeros([TotalSubjectsPredictions, n_regions])
for SubNum in range(0, TotalSubjectsPredictions):
# Lock data
x = bct.threshold_proportional(TempDataPredictions[:, :, SubNum],
TempThreshold, copy=True)
ss = analysis_space(BCT_Num, BCT_models, x, KeptYeoIDs)
TempPredictionsData[SubNum, :] = ss
model = Pipeline([('scaler', StandardScaler()), ('svr', SVR())])
all_data = np.concatenate((TempPredictionsData, TempResultsLockData))
test_fold = np.concatenate((- np.ones(len(TempPredictionsData)),np.zeros(len(TempResultsLockData))))
all_ages = np.concatenate((AgesPrediction.ravel(), AgesLockBoxData.ravel()))
ps = PredefinedSplit(test_fold)
mae, perm_score, p_val = permutation_test_score(model, all_data, all_ages,
n_jobs=None, random_state=5, verbose=0,
groups=None, cv=ps, n_permutations=n_permutations,
scoring="neg_mean_absolute_error")
cv_mae[DiffInit] = mae
CVPValBestModels[DiffInit] = p_val
perm_scores[DiffInit, :] = perm_score
# + id="iMqeSsZlL28V"
plot_bo_repetions(ModelEmbedding, PredictedAcc, BestModelGPSpaceModIndex,
BestModelEmpiricalModIndex, BestModelEmpirical,
ModelActualAccuracyCorrelation, output_path, ClassOrRegression)
# + id="fU9lX346z3h1"
# Download image to computer
from google.colab import files
files.download(str(output_path / 'BOpt20Repeats.svg'))
# + id="LpuIeYBvKN0X"
import pandas as pd
# Obtain the list of 20 models that were defined as the best models
df = pd.DataFrame({'Data_Run': Data_Run,'sparsities': Sparsities_Run,
'bct': BCT_Run})
df_best = df.iloc[BestModelEmpiricalModIndex]
df_best['mae']= cv_mae
df_best['p-val'] = CVPValBestModels
df_best
# + id="8kXvhDOM--CW"
repetions_results = {
'dataframe': df_best,
'BestModelGPSpaceModIndex': BestModelGPSpaceModIndex,
'BestModelEmpiricalIndex': BestModelEmpiricalModIndex,
'BestModelEmpirical': BestModelEmpirical,
'ModelActualAccuracyCorrelation': ModelActualAccuracyCorrelation
}
pickle.dump( repetions_results, open(str(output_path / "repetitions_results.p"), "wb" ) )
# + id="jjXx0Fn2-3A-"
print(df_best.to_latex(index=False))
# + id="gZklq7xAOx2Q"
from google.colab import files
files.download(str(output_path / 'repetitions_results.p'))
# + id="lIqSVje1Nq68"
| notebooks/multiverse_analysis_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="Images/Splice_logo.jpeg" width="250" height="200" align="left" >
# # Using the Feature Store, and Database Deployment, for model deployment
# +
#Begin spark session
from pyspark.sql import SparkSession
spark = SparkSession.builder.getOrCreate()
#Create pysplice context. Allows you to create a Spark dataframe using our Native Spark DataSource
from splicemachine.spark import PySpliceContext
splice = PySpliceContext(spark)
#Initialize our Feature Store API
from splicemachine.features import FeatureStore
from splicemachine.features.constants import FeatureType
fs = FeatureStore(splice)
#Initialize MLFlow
from splicemachine.mlflow_support import *
mlflow.register_feature_store(fs)
mlflow.register_splice_context(splice)
# -
# # Deploy Machine Learning model as a table in the database
# <img src="Images/database_deployment.png" width="1000" align="left" >
# ## Benefits of Database Model Deployment
# - ### Fast
# - ### Easly to deploy and govern
# - ### Integreates with our Feature Store
from splicemachine.notebook import get_mlflow_ui
get_mlflow_ui()
# ## Create the deployment table
# Load in most relevant features generated in the previous notebook
# %store -r features_list
# %store -r features_str
# + language="sql"
# -- Create schema and drop table, if necessary
# CREATE SCHEMA IF NOT EXISTS deployed_models;
# DROP TABLE IF EXISTS deployed_models.twimlcon_regression;
#
# +
#Define the training data frame. Necessary so the model table knows what columns to make
training_df = fs.get_training_set_from_view('twimlcon_customer_lifetime_value').dropna()
#create the table itself
jobid = mlflow.deploy_db( db_schema_name='deployed_models',db_table_name='twimlcon_regression', run_id= '<replace with your run id>',
primary_key={'CUSTOMERID':'INTEGER','EVAL_TIME':'TIMESTAMP'},
df=training_df.select(features_list)
)
#watch the table creation logs
mlflow.watch_job(jobid)
# -
# ## Insert data into this empty table using the Feature Store
# <img src="Images/FS_tables.png" width="800" height="400" align="left" >
# ### Get most up to date Feature Values in milliseconds
# #### Return features as a Spark dataframe
feature_vector = fs.get_feature_vector(features=features_list, join_key_values={'customerid':'14235'})
feature_vector
# #### Return features using SQL
feature_vector_sql = fs.get_feature_vector(features=features_list, return_sql=True, join_key_values={'customerid':'14235'})
print(feature_vector_sql)
# %%time
# %%sql
{Insert SQL from previous cell here}
# ### Generate and retreive predictions using INSERT/SELECT sequence on a single row
# + language="sql"
# truncate table deployed_models.twimlcon_regression;
# -
# %%time
splice.execute(f"""
INSERT INTO deployed_models.twimlcon_regression ( CUSTOMERID, {features_str} )
SELECT fset2.CUSTOMERID, {features_str}
FROM twimlcon_fs.customer_lifetime fset2,
twimlcon_fs.customer_rfm_by_category fset1
WHERE fset2.CUSTOMERID = 15838 AND fset1.CUSTOMERID = 15838
union all
SELECT fset2.CUSTOMERID, {features_str}
FROM twimlcon_fs.customer_lifetime fset2,
twimlcon_fs.customer_rfm_by_category fset1
WHERE fset2.CUSTOMERID = 15839 AND fset1.CUSTOMERID = 15839""")
# + language="sql"
# SELECT * FROM deployed_models.twimlcon_regression;
# -
# ### Generate and retreive predictions using INSERT/SELECT sequence on a multiple rows
# #### This process will take about a minute given that this is a small cluster, it scales in performance with scale of the cluster.
# + language="sql"
# truncate table deployed_models.twimlcon_regression;
# -
# %%time
splice.execute(f"""
INSERT INTO deployed_models.twimlcon_regression ( EVAL_TIME, CUSTOMERID, {features_str} ) --splice-properties useSpark=False
SELECT fset2.ASOF_TS, fset2.CUSTOMERID, {features_str}
FROM twimlcon_fs.customer_lifetime_history fset2,
twimlcon_fs.customer_rfm_by_category_history fset1
WHERE fset2.CUSTOMERID = fset1.CUSTOMERID
AND fset2.ASOF_TS >=fset1.ASOF_TS AND fset2.ASOF_TS<fset1.UNTIL_TS
AND fset2.ASOF_TS BETWEEN '2020-10-01' and '2020-12-31'
""")
# + language="sql"
# SELECT * FROM deployed_models.twimlcon_regression ORDER BY EVAL_TIME {limit 10};
# -
spark.stop()
| twimlcon-workshop-materials/4 - Model Deployment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pandas data analysis - case air quality data
# + [markdown] slideshow={"slide_type": "slide"}
# Some imports:
# + slideshow={"slide_type": "-"}
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn
pd.options.display.max_rows = 8
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Some 'theory': the groupby operation (split-apply-combine)
#
# By "group by" we are referring to a process involving one or more of the following steps
#
# * **Splitting** the data into groups based on some criteria
# * **Applying** a function to each group independently
# * **Combining** the results into a data structure
#
# <img src="img/splitApplyCombine.png">
#
# Similar to SQL `GROUP BY`
# + [markdown] slideshow={"slide_type": "subslide"}
# The example of the image in pandas syntax:
# -
df = pd.DataFrame({'key':['A','B','C','A','B','C','A','B','C'],
'data': [0, 5, 10, 5, 10, 15, 10, 15, 20]})
df
# + slideshow={"slide_type": "subslide"}
df.groupby('key').aggregate('sum') # np.sum
# -
df.groupby('key').sum()
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Back to the air quality data
# -
# <div class="alert alert-success">
# <b>QUESTION</b>: how does the *typical monthly profile* look like for the different stations?
# </div>
# 1\. add a column to the dataframe that indicates the month (integer value of 1 to 12):
# + clear_cell=true
data['month'] = data.index.month
# + [markdown] slideshow={"slide_type": "subslide"}
# 2\. Now, we can calculate the mean of each month over the different years:
# + clear_cell=true
data.groupby('month').mean()
# -
# 3\. plot the typical monthly profile of the different stations:
# + clear_cell=true slideshow={"slide_type": "subslide"}
data.groupby('month').mean().plot()
# -
# <div class="alert alert-success">
# <b>QUESTION</b>: plot the weekly 95% percentiles of the concentration in 'BETR801' and 'BETN029' for 2011
# </div>
#
df2011 = data['2011']
df2011.groupby(df2011.index.week)[['BETN029', 'BETR801']].quantile(0.95).plot()
data = data.drop('month', axis=1)
# + [markdown] slideshow={"slide_type": "subslide"}
# <div class="alert alert-success">
# <b>QUESTION</b>: The typical diurnal profile for the different stations?
# </div>
# + slideshow={"slide_type": "fragment"}
data.groupby(data.index.hour).mean().plot()
# + [markdown] slideshow={"slide_type": "subslide"}
# <div class="alert alert-success">
# <b>QUESTION</b>: What is the difference in the typical diurnal profile between week and weekend days?
# </div>
# +
# data.index.weekday?
# -
data['weekday'] = data.index.weekday
# + [markdown] slideshow={"slide_type": "subslide"}
# Add a column indicating week/weekend
# -
data['weekend'] = data['weekday'].isin([5, 6])
data_weekend = data.groupby(['weekend', data.index.hour]).mean()
data_weekend.head()
# + slideshow={"slide_type": "subslide"}
data_weekend_FR04012 = data_weekend['FR04012'].unstack(level=0)
data_weekend_FR04012.head()
# + slideshow={"slide_type": "subslide"}
data_weekend_FR04012.plot()
# -
data = data.drop(['weekday', 'weekend'], axis=1)
# + [markdown] slideshow={"slide_type": "subslide"}
# <div class="alert alert-success">
# <b>QUESTION</b>: What are the number of exceedances of hourly values above the European limit 200 µg/m3 ?
# </div>
# -
exceedances = data > 200
# group by year and count exceedances (sum of boolean)
exceedances = exceedances.groupby(exceedances.index.year).sum()
exceedances
ax = exceedances.loc[2005:].plot(kind='bar')
ax.axhline(18, color='k', linestyle='--')
# <div class="alert alert-success">
# <b>QUESTION</b>: And are there exceedances of the yearly limit value of 40 µg/m3 since 200 ?
# </div>
yearly = data['2000':].resample('A')
(yearly > 40).sum()
yearly.plot()
plt.axhline(40, linestyle='--', color='k')
# + [markdown] slideshow={"slide_type": "subslide"}
# <div class="alert alert-success">
# <b>QUESTION</b>: Visualize the typical week profile for the different stations as boxplots.
# </div>
#
# Tip: the boxplot method of a DataFrame expects the data for the different boxes in different columns)
# + clear_cell=true slideshow={"slide_type": "fragment"}
# add a weekday and week column
data['weekday'] = data.index.weekday
data['week'] = data.index.week
data.head()
# + clear_cell=true slideshow={"slide_type": "subslide"}
# pivot table so that the weekdays are the different columns
data_pivoted = data['2012'].pivot_table(columns='weekday', index='week', values='FR04037')
data_pivoted.head()
# + clear_cell=true
box = data_pivoted.boxplot()
# + [markdown] slideshow={"slide_type": "subslide"}
# <div class="alert alert-success">
# <b>QUESTION</b>: Calculate the correlation between the different stations
# </div>
#
# + clear_cell=true
data[['BETR801', 'BETN029', 'FR04037', 'FR04012']].corr()
# + clear_cell=true
data[['BETR801', 'BETN029', 'FR04037', 'FR04012']].resample('D').corr()
# + [markdown] slideshow={"slide_type": "slide"}
# # Further reading
#
# - the documentation: http://pandas.pydata.org/pandas-docs/stable/
# - Wes McKinney's book "Python for Data Analysis"
# - lots of tutorials on the internet, eg http://github.com/jvns/pandas-cookbook
#
# -
# ## Acknowledgement
#
# > *© 2015, <NAME> and <NAME> (<mailto:<EMAIL>>, <mailto:<EMAIL>>). Licensed under [CC BY 4.0 Creative Commons](http://creativecommons.org/licenses/by/4.0/)*
| 04 - GroupBy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kmunve/APS/blob/master/Predict_aval_problem_combined.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="DQpuvUJRuqCJ" colab_type="text"
# # Predicting tomorrow's avalanche problem
#
# We use features from the last 3 days avalanche warnings to predict the main avalanche problem for the coming day.
# An avalanche problem describes why an avalanche danger exists and how severe it is.
#
# An avalanche problem contains a cause, a distribution, a potential avalanche size and a sensitivity of triggering. E.g.
#
#
#
# * Cause: Wind slabs
# * Distribution: Widespread
# * Size: Large
# * Sensitivity: Easy to trigger
#
# This is encoded as a 4-digit number where each digit encodes one of the four parameters, e.g. 5332.
#
#
# We use differnet decision tree approaches to predict these four elements.
# + [markdown] id="QAYi4teG5bwk" colab_type="text"
# ## Imports
# + id="f6kUT_Qqmxd5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="7f9789b3-efc3-4fff-b450-9a9cef840ff9"
import pandas as pd
import numpy as np
import json
import sklearn
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
import warnings
warnings.simplefilter('ignore')
print('Pandas:\t', pd.__version__)
print('Numpy:\t', np.__version__)
print('Scikit Learn:\t', sklearn.__version__)
print('Matplotlib:\t', matplotlib.__version__)
# + [markdown] id="NC1t4nM35hUm" colab_type="text"
# ## Data
# + id="BI_dP5x1zgbI" colab_type="code" outputId="a427404c-a18e-4a88-ecc6-86aad29c8255" colab={"base_uri": "https://localhost:8080/", "height": 84}
# define the decoder function for the 4-digit avalanche problem target
# !curl https://raw.githubusercontent.com/kmunve/APS/master/aps/config/snoskred_keys.json > snoskred_keys.json
def print_aval_problem_combined(aval_combined_int):
aval_combined_str = str(aval_combined_int)
with open('snoskred_keys.json') as jdata:
snoskred_keys = json.load(jdata)
type_ = snoskred_keys["Class_AvalancheProblemTypeName"][aval_combined_str[0]]
dist_ = snoskred_keys["Class_AvalDistributionName"][aval_combined_str[1]]
sens_ = snoskred_keys["Class_AvalSensitivityId"][aval_combined_str[2]]
size_ = snoskred_keys["DestructiveSizeId"][aval_combined_str[3]]
return f"{type_} : {dist_} : {sens_} : {size_}"
print(print_aval_problem_combined(6231))
# + id="KQ4LJCmdm5nr" colab_type="code" outputId="5db6cbec-77de-4e9b-9529-3eead4dabec5" colab={"base_uri": "https://localhost:8080/", "height": 386}
# get the data
### Dataset with previous forecasts and observations
v_df = pd.read_csv('https://raw.githubusercontent.com/hvtola/HTLA/master/varsom_ml_preproc_htla2.csv', index_col=0) # --- Added even more data from RegObs
# v_df = pd.read_csv('https://raw.githubusercontent.com/hvtola/HTLA/master/varsom_ml_preproc_htla.csv', index_col=0)
### Dataset with previous forecasts only
# v_df = pd.read_csv('https://raw.githubusercontent.com/kmunve/APS/master/aps/notebooks/ml_varsom/varsom_ml_preproc_3y.csv', index_col=0).drop_duplicates() # for some reason we got all rows twice in that file :-(
# v_df[['date', 'region_id', 'region_group_id', 'danger_level', 'avalanche_problem_1_cause_id']].head(791*4+10)
# v_df['region_id'].value_counts()
v_df['region_id'].value_counts()
# + id="2cw07yP5RiJS" colab_type="code" colab={}
# v_df['date'].value_counts()
# + id="OocLg0Ffm_0Q" colab_type="code" outputId="3381411b-07c1-4f42-87bb-a4218fae3799" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# keep only numeric columns
from pandas.api.types import is_numeric_dtype
num_cols = [var for var in v_df.columns.values if is_numeric_dtype(v_df[var])]
print(len(num_cols))
num_cols
# + id="hnyAeXahvGK0" colab_type="code" colab={}
# drop features that are related to the forecast we want to predict and features that should have no influence
drop_list = [
'danger_level',
'aval_problem_1_combined',
'avalanche_problem_1_cause_id',
'avalanche_problem_1_destructive_size_ext_id',
'avalanche_problem_1_distribution_id',
'avalanche_problem_1_exposed_height_1',
'avalanche_problem_1_exposed_height_2',
'avalanche_problem_1_ext_id',
'avalanche_problem_1_probability_id',
'avalanche_problem_1_problem_id',
'avalanche_problem_1_problem_type_id',
'avalanche_problem_1_trigger_simple_id',
'avalanche_problem_1_type_id',
'avalanche_problem_2_cause_id',
'avalanche_problem_2_destructive_size_ext_id',
'avalanche_problem_2_distribution_id',
'avalanche_problem_2_exposed_height_1',
'avalanche_problem_2_exposed_height_2',
'avalanche_problem_2_ext_id',
'avalanche_problem_2_probability_id',
'avalanche_problem_2_problem_id',
'avalanche_problem_2_problem_type_id',
'avalanche_problem_2_trigger_simple_id',
'avalanche_problem_2_type_id',
'avalanche_problem_3_cause_id',
'avalanche_problem_3_destructive_size_ext_id',
'avalanche_problem_3_distribution_id',
'avalanche_problem_3_exposed_height_1',
'avalanche_problem_3_exposed_height_2',
'avalanche_problem_3_ext_id',
'avalanche_problem_3_probability_id',
'avalanche_problem_3_problem_id',
'avalanche_problem_3_problem_type_id',
'avalanche_problem_3_trigger_simple_id',
'avalanche_problem_3_type_id',
'avalanche_problem_1_problem_type_id_class',
'avalanche_problem_1_sensitivity_id_class',
'avalanche_problem_1_trigger_simple_id_class',
'avalanche_problem_2_problem_type_id_class',
'avalanche_problem_2_sensitivity_id_class',
'avalanche_problem_2_trigger_simple_id_class',
'avalanche_problem_3_problem_type_id_class',
'avalanche_problem_3_sensitivity_id_class',
'avalanche_problem_3_trigger_simple_id_class',
'emergency_warning_Ikke gitt',
'emergency_warning_Naturlig utløste skred',
'author_Andreas@nve',
'author_Eldbjorg@MET',
'author_<NAME>',
'author_EspenN',
'author_Halvor@NVE',
'author_HåvardT@met',
'author_Ida@met',
'author_Ingrid@NVE',
'author_<NAME>',
'author_JonasD@ObsKorps',
'author_Julie@SVV',
'author_Jørgen@obskorps',
'author_Karsten@NVE',
'author_MSA@nortind',
'author_Matilda@MET',
'author_Odd-Arne@NVE',
'author_Ragnar@NVE',
'author_Ronny@NVE',
'author_Silje@svv',
'author_Tommy@NVE',
'author_ToreV@met',
'author_anitaaw@met',
'author_emma@nve',
'<EMAIL>',
'<EMAIL>',
'author_jan arild<EMAIL>',
'author_jegu@NVE',
'author_jostein<EMAIL>',
'author_kn<EMAIL>',
'author_magnush@met',
'author_martin@<EMAIL>',
'author_ragnhildn@met',
'author_rue@<EMAIL>',
'author_siri@<EMAIL>',
'author_solveig@NVE',
'author_to<EMAIL>',
'author_torolav@obskorps'
]
# + id="Suovm9CqVfZc" colab_type="code" colab={}
v_df.describe()
v_df = v_df.fillna(0) # be careful here !!!
# + id="yHIQCj19ndfP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="aff0cadb-c855-4c2e-9f5e-3198e5701265"
target_name = 'aval_problem_1_combined'
y_df = v_df[target_name]
y = y_df.values
X_df = v_df.filter(num_cols).drop(drop_list, axis='columns')
X = X_df.values
feature_names = X_df.columns.values
print(len(feature_names))
# + id="OpQ7kDImqAhn" colab_type="code" outputId="2eb200c6-8965-4ea0-a32d-86923432290d" colab={"base_uri": "https://localhost:8080/", "height": 34}
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=2442) # Fikk feilmelding med stratify=y
X_train.shape, y_train.shape, X_test.shape, y_test.shape
# + [markdown] id="wGSmuB-24Z39" colab_type="text"
# ## Decision tree
# + id="JHft6rvFqBh6" colab_type="code" outputId="aa133133-4a8d-4177-b9f5-1b4b5d0cba19" colab={"base_uri": "https://localhost:8080/", "height": 151}
from sklearn.tree import DecisionTreeClassifier
clf = DecisionTreeClassifier(max_depth=10)
# %time clf.fit(X_train, y_train)
# + id="c-kNA0jTq7ag" colab_type="code" outputId="317598c0-621c-4ee7-e29a-3584df40a376" colab={"base_uri": "https://localhost:8080/", "height": 34}
print('Decision tree with {0} leaves has a training score of {1} and a test score of {2}'.format(clf.tree_.max_depth, clf.score(X_train, y_train), clf.score(X_test, y_test)))
# + id="5wGNuTjir0-O" colab_type="code" outputId="ddbc4db9-170a-4ba1-eb60-ddd0ad07a0b3" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# just checking if the values make sense
k = 21 # error when using 1230
for i in range(len(feature_names)):
print(feature_names[i], ':\t', X_test[k, i])
prediction_ = clf.predict(X_test[k, :].reshape(1, -1))
print(target_name, ':\t', y_test[k], prediction_)
print(print_aval_problem_combined(prediction_[0])) # add information about dangerlevel
# + id="liG_Wv5HqOQP" colab_type="code" outputId="f9a0dd8c-b13f-4149-bbfc-e0a0919fa081" colab={"base_uri": "https://localhost:8080/", "height": 520}
# Finding the best parameters
s_test = []
s_train = []
ks = np.arange(1, 30, dtype=int)
for k in ks:
clf_ = DecisionTreeClassifier(max_depth = k)
clf_.fit(X_train, y_train)
s_train.append(clf_.score(X_train, y_train))
s_test.append(clf_.score(X_test, y_test))
#clf.score(X_train, y_train), clf.score(X_test, y_test)))
s_test = np.array(s_test)
print(s_test.max(), s_test.argmax())
plt.figure(figsize=(10, 8))
plt.plot(ks, s_test, color='red', label='test')
plt.plot(ks, s_train, color='blue', label='train')
plt.legend()
# + [markdown] id="Qz7COc684PRV" colab_type="text"
# ## Feature importance
# + id="n-ZhAx-7sstl" colab_type="code" outputId="862519ea-11d1-4730-bd5f-9e99e3126d5b" cellView="code" colab={"base_uri": "https://localhost:8080/", "height": 1000}
importance = clf.feature_importances_
feature_indexes_by_importance = importance.argsort()
for i in feature_indexes_by_importance:
print('{}-{:.2f}%'.format(feature_names[i], (importance[i] *100.0)))
# + id="ZlmMdc5XplCd" colab_type="code" outputId="25465538-fe67-45af-babb-75b6a1a29281" colab={"base_uri": "https://localhost:8080/", "height": 1000}
fig, ax = plt.subplots(figsize=(8,20))
y_pos = np.arange(len(feature_names))
ax.barh(y_pos, clf.feature_importances_*100, align='center')
ax.set_yticks(y_pos)
ax.set_yticklabels(feature_names)
ax.invert_yaxis() # labels read top-to-bottom
ax.set_xlabel('Feature importance')
ax.set_title('How much does each feature contribute?')
# + [markdown] id="TUflF7o2wX4f" colab_type="text"
# ## Using RandomForest
# + id="cm54kI64tVST" colab_type="code" outputId="d8c0272f-5c9f-45ac-bf94-7e67edf7f56b" colab={"base_uri": "https://localhost:8080/", "height": 168}
from sklearn.ensemble import RandomForestRegressor, RandomForestClassifier
rfc = RandomForestClassifier(n_estimators=30, min_samples_split=15)
rfc.fit(X_train, y_train)
predic_proba_rfc = rfc.predict_proba(X_test)
predictions_rfc = rfc.predict(X_test)
print('Random Forest Classifier with {0} leaves has a training score of {1} and a test score of {2}'.format(rf.max_depth, rf.score(X_train, y_train), rf.score(X_test, y_test)))
print(predictions_rfc)
print(predic_proba_rfc)
# + colab_type="code" outputId="4a2a81fb-c678-4c55-ea13-c5b6e3904fe9" id="qlwvzh7cHc6C" colab={"base_uri": "https://localhost:8080/", "height": 67}
rf = RandomForestRegressor(n_estimators=30, min_samples_split=15)
rf.fit(X_train, y_train)
predictions_rf = rf.predict(X_test)
print('Random Forest Regressor with {0} leaves has a training score of {1} and a test score of {2}'.format(rf.max_depth, rf.score(X_train, y_train), rf.score(X_test, y_test)))
print(predictions_rf)
# + id="uV07QOGiwk3x" colab_type="code" outputId="4b931ee0-a361-4c45-afe8-c755d66d3176" colab={"base_uri": "https://localhost:8080/", "height": 1000}
importance_rf = rf.feature_importances_
feature_indexes_by_importance_rf = importance_rf.argsort()
for i in feature_indexes_by_importance_rf:
print('{}-{:.2f}%'.format(feature_names[i], (importance_rf[i] *100.0)))
# + [markdown] id="q_RGZ7z-83vU" colab_type="text"
# ## Questions and further thoughts
# - why does the RandomForestRegressor perform better than the RandomForestClassifier?
# - how do I get the number of leaves of a RF?
# - are there other classic ML methods that would be suitable?
# - would the use of NN be beneficial? Can we use it to encode field observations?
# - sometimes we know a forecast (used for training) was right or wrong, often we don't. can we flag these cases and use it as a weighting?
# - use a similar feature set to get the nearest neighbor (date, region) from the forecasts in the database
# + id="jNdNgkLJwuPF" colab_type="code" colab={}
| Predict_aval_problem_combined.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/nmningmei/Deep_learning_fMRI_EEG/blob/master/10_1_searchlight_representational_similarity_analysis%2C_bonus_decoding.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Df-vPFlvQ8p3"
# # The script to illustrate a way to perform searchlight RSA.
# + [markdown] id="HXrHGp7VSWEv"
# # Get the extracted features and the mask files
# + id="xzt5iZvKPx0H" outputId="e68a1b94-d567-4fbe-db33-ec49800733b2" colab={"base_uri": "https://localhost:8080/"}
try:
# !git clone https://github.com/nmningmei/Extracted_features_of_Spanish_image_dataset.git
except:
# !ls Extracted_features_of_Spanish_image_dataset
# + [markdown] id="XaBuTFDoZQTv"
# # Get the fMRI data
# + id="uN2ilE3MYPWj"
# Import PyDrive and associated libraries.
# This only needs to be done once per notebook.
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
# Authenticate and create the PyDrive client.
# This only needs to be done once per notebook.
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
# Download a file based on its file ID.
#
# A file ID looks like: laggVyWshwcyP6kEI-y_W3P8D26sz
file_id = '1vLWSdXpOxqp3jOCypsWA27AEptsGdwav'
downloaded = drive.CreateFile({'id': file_id})
# + id="siP40EAcRdO_"
downloaded.GetContentFile('whole_bran.tar.gz')
# + id="9t4uX01wY6aU" outputId="e3b2ef0f-6f00-45bc-b124-bca00060f8ce" colab={"base_uri": "https://localhost:8080/"}
# !tar -xvf whole_bran.tar.gz
# + id="2olG3v16ZKVC" outputId="8e0bc03a-05d6-4db0-a72a-46f23ea0320d" colab={"base_uri": "https://localhost:8080/"}
# ls
# + [markdown] id="gAz9nOmO_-I5"
# # install and update some of the libraries if not
# + id="n9Rkj9z1ZOc0" outputId="7813d44d-ee0e-41b0-abf8-a5cc3379127a" colab={"base_uri": "https://localhost:8080/"}
try:
from nilearn.input_data import NiftiMasker
from nilearn.image import new_img_like
from brainiak.searchlight.searchlight import Searchlight
from brainiak.searchlight.searchlight import Ball
except:
# !pip install nilearn
# !python3 -m pip install -U brainiak
from nilearn.input_data import NiftiMasker
from nilearn.image import new_img_like
from brainiak.searchlight.searchlight import Searchlight
from brainiak.searchlight.searchlight import Ball
import os,gc
import pandas as pd
import numpy as np
from nibabel import load as load_fmri
from joblib import Parallel,delayed
from scipy.spatial import distance
from scipy.stats import spearmanr
# + id="WpT4JwEPlXV5"
def normalize(data,axis = 1):
return data - data.mean(axis).reshape(-1,1)
# Define voxel function
def sfn(l, msk, myrad, bcast_var):
"""
l: BOLD
msk: mask array
myrad: not use
bcast_var: label -- CNN features
"""
BOLD = l[0][msk,:].T.copy() # vectorize the voxel values in the sphere
model = bcast_var.copy() # vectorize the RDM
# pearson correlation
RDM_X = distance.pdist(normalize(BOLD),'correlation')
RDM_y = distance.pdist(normalize(model),'correlation')
D,p = spearmanr(RDM_X,RDM_y)
return D
def process_csv(file_name = 'whole_brain_conscious.csv'):
"""
to add some info to the event files to create better cross-validation folds
"""
df_data = pd.read_csv(file_name)
df_data['id'] = df_data['session'] * 1000 + df_data['run'] * 100 + df_data['trials']
df_data = df_data[df_data.columns[1:]]
return df_data
# + id="upBOZVVYnfDO"
radius = 3 # in mm, the data has voxel size of 2.4mm x 2.4mm x 2.4mm
feature_dir = 'Extracted_features_of_Spanish_image_dataset/computer_vision_features_no_background'
model_name = 'VGG19'
label_map = {'Nonliving_Things':[0,1],'Living_Things':[1,0]}
whole_brain_mask = 'Extracted_features_of_Spanish_image_dataset/combine_BOLD.nii.gz'
average = True
n_splits = 10 # recommend to perform the resampling for more than 500 times
n_jobs = -1
# + [markdown] id="CuOAEFxThxXc"
# # implementation of resampling
#
# 1. when the study is event-related but the events are not balanced, distribution is hard to measure, so resampling could avoid handling the assumption of normal distribution
# 2. by the law of large number, when the time of resampling is large enough (exhaust the permutation, i.e. 1-2-3, 2-3-1, 3-1-2, 1-3-2, 2-1-3, 3-2-1, or just a large number, like n = 1000), the average of the resampling estimate is an unbias estimate of the population measure, and the 95% credible interval (confidence interval for frequentist statitistics) contains the population with chance of 95%.
# 3. so, we gather one trial of each unique item, in total 96 trial, for both the BOLD signals (96, 88, 88, 66) and the CNN features (96, 300), and we compute the RDM of the BOLD signals in the searchlight sphere and the RDM of the CNN features regardless of the searchlight sphere. We then correlate the RDM of the BOLD in the searchlight sphere to the RDM of the CNN features.
# 4. repeat step 3. for different sampling
# + id="kScQ0txinypn" outputId="9a8c0ba2-8447-4af6-f984-0f59ad1c2d29" colab={"base_uri": "https://localhost:8080/"}
for conscious_state in ['unconscious','conscious']:
np.random.seed(12345)
df_data = process_csv(f'whole_brain_{conscious_state}.csv')
# load the data in the format of numpy but keep the 4D dimensions
BOLD_image = load_fmri(f'whole_brain_{conscious_state}.nii.gz')
print(f'{conscious_state}\tfMRI in {BOLD_image.shape} events in {df_data.shape}')
targets = np.array([label_map[item] for item in df_data['targets']])[:,-1]
# get the image names in the order of the experimental trials
images = df_data['paths'].apply(lambda x: x.split('.')[0] + '.npy').values
# get the CNN features (n_trial x 300)
CNN_feature = np.array([np.load(os.path.join(feature_dir,
model_name,
item)) for item in images])
groups = df_data['labels'].values
# define a function to create the folds first
def _proc(df_data):
"""
This is useful when the number of folds are thousands
"""
df_picked = df_data.groupby('labels').apply(lambda x: x.sample(n = 1).drop('labels',axis = 1)).reset_index()
df_picked = df_picked.sort_values(['targets','subcategory','labels'])
idx_test = df_picked['level_1'].values
return idx_test
print(f'partitioning data for {n_splits} folds')
idxs = Parallel(n_jobs = -1, verbose = 1)(delayed(_proc)(**{
'df_data':df_data,}) for _ in range(n_splits))
gc.collect() # free memory that is occupied by garbage
# define a function to run the RSA
def _searchligh_RSA(idx,
sl_rad = radius,
max_blk_edge = radius - 1,
shape = Ball,
min_active_voxels_proportion = 0,
):
# Brainiak function
sl = Searchlight(sl_rad = sl_rad,
max_blk_edge = max_blk_edge,
shape = shape,
min_active_voxels_proportion = min_active_voxels_proportion,
)
# distribute the data based on the sphere
## the first input is usually the BOLD signal, and it is in the form of
## lists not arrays, representing each subject
## the second input is usually the mask, and it is in the form of array
sl.distribute([np.asanyarray(BOLD_image.dataobj)[:,:,:,idx]],
np.asanyarray(load_fmri(whole_brain_mask).dataobj) == 1)
# broadcasted data is the data that remains the same during RSA
sl.broadcast(CNN_feature[idx])
# run searchlight algorithm
global_outputs = sl.run_searchlight(sfn,
pool_size = 1, # we run each RSA using a single CPU
)
return global_outputs
for _ in range(10):
gc.collect()
res = Parallel(n_jobs = -1,verbose = 1,)(delayed(_searchligh_RSA)(**{
'idx':idx}) for idx in idxs)
# save the data
results_to_save = np.zeros(np.concatenate([BOLD_image.shape[:3],[n_splits]]))
for ii,item in enumerate(res):
results_to_save[:,:,:,ii] = np.array(item, dtype=np.float)
results_to_save = new_img_like(BOLD_image,results_to_save,)
results_to_save.to_filename(f'RSA_{conscious_state}.nii.gz')
# + [markdown] id="mgSZDnr1anO9"
# # let's modify the code above to make it a searchlight decoding, without using nilearn
# + id="RGIC_Oc1b6F4"
# modify the voxel function for decoding
# like defining the decoder
# like putting the train-test pipeline in place
def sfn(l, msk, myrad, bcast_var):
"""
l: BOLD
msk: mask array
myrad: not use
bcast_var: label
"""
from sklearn.svm import LinearSVC
from sklearn.calibration import CalibratedClassifierCV
from sklearn.preprocessing import MinMaxScaler
from sklearn.pipeline import make_pipeline
from sklearn.metrics import roc_auc_score
BOLD = l[0][msk,:].T.copy() # vectorize the voxel values in the sphere
targets,idx_train,idx_test = bcast_var
# scaler the data to between 0 and 1, improve SVM decoding
scaler = MinMaxScaler()
# default with L2 regularization
svm = LinearSVC(class_weight = 'balanced',random_state = 12345)
# make the SVM to produce probabilistic predictions
svm = CalibratedClassifierCV(svm, cv = 5)
pipeline = make_pipeline(scaler,svm)
pipeline.fit(BOLD[idx_train],targets[idx_train])
y_pred = pipeline.predict_proba(BOLD[idx_test])[:,-1]
score = roc_auc_score(targets[idx_test],y_pred)
return score
# + id="j7CSwuBND7DC"
for conscious_state in ['unconscious','conscious']:
np.random.seed(12345)
df_data = process_csv(f'whole_brain_{conscious_state}.csv')
# load the data in the format of numpy but keep the 4D dimensions
BOLD_image = load_fmri(f'whole_brain_{conscious_state}.nii.gz')
print(f'{conscious_state}\tfMRI in {BOLD_image.shape} events in {df_data.shape}')
targets = np.array([label_map[item] for item in df_data['targets']])[:,-1]
# get the image names in the order of the experimental trials
images = df_data['paths'].apply(lambda x: x.split('.')[0] + '.npy').values
# get the CNN features (n_trial x 300)
CNN_feature = np.array([np.load(os.path.join(feature_dir,
model_name,
item)) for item in images])
groups = df_data['labels'].values
############################################################################
######## replace this part #################################################
# define a function to create the folds first
def _proc(df_data):
"""
This is useful when the number of folds are thousands
"""
df_picked = df_data.groupby('labels').apply(lambda x: x.sample(n = 1).drop('labels',axis = 1)).reset_index()
df_picked = df_picked.sort_values(['targets','subcategory','labels'])
idx_test = df_picked['level_1'].values
return idx_test
print(f'partitioning data for {n_splits} folds')
idxs = Parallel(n_jobs = -1, verbose = 1)(delayed(_proc)(**{
'df_data':df_data,}) for _ in range(n_splits))
gc.collect() # free memory that is occupied by garbage
############################################################################
######## with this #########################################################
from sklearn.model_selection import StratifiedShuffleSplit
cv = StraitifiedShuffleSplit(n_splits = 10, test_size = 0.2, random_state = 12345)
idxs_train,idxs_test = [],[]
for idx_train,idx_test in cv.split(df_data,targets):
idxs_train.append(idx_train)
idxs_test.append(idx_test)
############################################################################
##### end of modification No.1##############################################
############################################################################
# define a function to run the RSA
def _searchligh_RSA(idx,
sl_rad = radius,
max_blk_edge = radius - 1,
shape = Ball,
min_active_voxels_proportion = 0,
):
# get the train,test split of a given fold
idx_train,idx_test = idx
# Brainiak function
sl = Searchlight(sl_rad = sl_rad,
max_blk_edge = max_blk_edge,
shape = shape,
min_active_voxels_proportion = min_active_voxels_proportion,
)
# distribute the data based on the sphere
## the first input is usually the BOLD signal, and it is in the form of
## lists not arrays, representing each subject
## the second input is usually the mask, and it is in the form of array
sl.distribute([np.asanyarray(BOLD_image.dataobj)],
np.asanyarray(load_fmri(whole_brain_mask).dataobj) == 1)
########################################################################
##### second modification ##############################################
# broadcasted data is the data that remains the same during RSA
sl.broadcast([targets,idx_train,idx_test]) # <-- add the indices of training and testing
###### end of modification No.2 ########################################
# run searchlight algorithm
global_outputs = sl.run_searchlight(sfn,
pool_size = 1, # we run each RSA using a single CPU
)
return global_outputs
for _ in range(10):
gc.collect()
res = Parallel(n_jobs = -1,verbose = 1,)(delayed(_searchligh_RSA)(**{
'idx':idx}) for idx in idxs)
# save the data
results_to_save = np.zeros(np.concatenate([BOLD_image.shape[:3],[n_splits]]))
for ii,item in enumerate(res):
results_to_save[:,:,:,ii] = np.array(item, dtype=np.float)
results_to_save = new_img_like(BOLD_image,results_to_save,)
results_to_save.to_filename(f'searchlight_decoding_{conscious_state}.nii.gz')
| 10_1_searchlight_representational_similarity_analysis,_bonus_decoding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] cell_id="00000-65d397b6-62c5-4e7b-8f8e-a5676cddba9d" deepnote_cell_type="markdown" tags=[]
# # Homework 2
#
# List your name and the names of any collaborators at the top of this notebook.
#
# (Reminder: It's encouraged to work together; you can even submit the exact same homework as another student or two students, but you must list each other's names at the top.)
# + [markdown] cell_id="00023-25a4c333-9180-47fe-8bd9-11c1a27fbaad" deepnote_cell_type="markdown" tags=[]
# ## Exercise 1
#
# Using the Python function `getsizeof` (hint: you need to import a library to get access to it), find out how much space in memory the following take:
#
# 1. The integer 0.
# 1. An empty list.
# 1. A list containing the integers 1,2,3.
# 1. A list containing the strings 1,2,3.
# 1. A NumPy array containg the integers (or floats) 1,2,3. (We haven't talked about the data type of elements inside NumPy arrays, so don't worry about that.)
# 1. The `range` object `range(0,10**100,3)`.
#
# + [markdown] cell_id="00002-667b605d-0f9d-4644-81c9-954737848b60" deepnote_cell_type="markdown" tags=[]
# ## Exercise 2
#
# Write a function `make_arr` that takes as input a positive integer `n`, and as output returns a length `n` NumPy array containing random integers between 1 and 100 (inclusive). Use the `integers` method of an object produced by NumPy's `default_rng()`.
# + [markdown] cell_id="00001-bdd74195-31e4-40e3-b069-9bd18312834d" deepnote_cell_type="markdown" tags=[]
# ## Exercise 3
#
# Using your function `make_arr`, create a length one million array of random integers between 1 and 100 (inclusive). Save this array with the variable name `arr`.
# + [markdown] cell_id="00005-ef8450da-56d4-4e5a-954e-9f0265635395" deepnote_cell_type="markdown" tags=[]
# ## Exercise 4
#
# Compute the reciprocals of each element in `arr` by evaluating `1/arr`. How long does it take? (Use `%%timeit`.)
# + [markdown] cell_id="00007-96ed3942-96b4-45be-8b97-bebd8d08333c" deepnote_cell_type="markdown" tags=[]
# ## Exercise 5
#
# Convert `arr` into a list called `my_list`, and then use list comprehension to compute the reciprocals of each element in `my_list`. Time how long this takes using `%%timeit`. (Don't include the conversion to a list in the `%%timeit` cell; do the conversion before.)
# + [markdown] cell_id="00010-6fde3207-d343-4e61-9d9f-68584d034017" deepnote_cell_type="markdown" tags=[]
# ## Exercise 6
#
# In a markdown cell, indicate how the times compare for these two methods.
# + [markdown] cell_id="00005-b00eff0b-3c98-4d60-ba0f-36dc5d31a665" deepnote_cell_type="markdown" tags=[]
# ## Exercise 7
#
# What proportion of the elements in `arr` are equal to 100? Answer this question a few different ways; all these answers should be equal and should be very close to `0.01`).
#
# 1. Use `my_list` that you created above and the `count` method of a list to determine how often 100 occurs. Then divide by the total length. (To get the total length, use `len`, don't type out the length explicitly.)
#
# 1. Using list comprehension, make a list containing all the elements of `arr` which are equal to 100 (you don't need to use `my_list`; just pretend `arr` is a list and everything will work fine). Then compute the length of this new list divided by the length of `arr`. (This isn't a great strategy; it is mostly an excuse to practice with list comprehension.)
#
# 2. Make a Boolean array which is `True` whereever `arr` is 100 and which is `False` everywhere else. Then use the NumPy function `np.count_nonzero`, and then divide by the length of `arr`.
#
# 3. Convert the array into a pandas Series, and then apply the method `.value_counts()`, then compute `s[100]`, where `s` represents the output of `.value_counts()`, then divide by the length of `arr`.
# + [markdown] cell_id="00006-f1ec5c0a-8ead-4440-97d7-fd78402c9a67" deepnote_cell_type="markdown" tags=[]
# ## Exercise 8
#
# Repeat each of the previous computations, this time using `%%timeit` to see how long they take. In a markdown cell, report what answers you get. (For the pandas Series part, convert to the pandas Series outside of the `%%timeit` cell... It gives NumPy an unfair advantage if that conversion is included in the timing portion.)
# + [markdown] cell_id="00021-793b0d86-1388-4deb-bf3f-57a64fae514e" deepnote_cell_type="markdown" tags=[]
# ## Exercise 9
#
# In a markdown cell, answer the following question: Was one of the four methods significantly faster than the rest? Was one of the four methods significantly slower than the rest?
# + [markdown] cell_id="00021-766fa7be-de3a-4d02-878f-9a3828b72ed4" deepnote_cell_type="markdown" tags=[]
# ## Exercise 10
#
# Many of these exercises are about how to make various operations run faster by choosing appropriate data types. What do you think is one of the main reasons that this is relevant to data science?
# + [markdown] cell_id="00011-c71018e9-d115-4662-9314-fcdd124cf266" deepnote_cell_type="markdown" tags=[]
# ## Submission
#
# Download the .ipynb file for this notebook (click on the folder icon to the left, then the ... next to the file name) and upload the file on Canvas.
| _build/html/_sources/Week2/Homework2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] Collapsed="false"
# # Train a CNN
#
# In this notebook we will go through all the steps required to train a fully convolutional neural network. Because this takes a while and uses a lot of GPU RAM a separate command line script (`train_nn.py`) is also provided in the `src` directory.
# + Collapsed="false"
# %load_ext autoreload
# %autoreload 2
# + Collapsed="false"
# Depending on your combination of package versions, this can raise a lot of TF warnings...
import numpy as np
import matplotlib.pyplot as plt
import xarray as xr
import tensorflow as tf
import tensorflow.keras as keras
from tensorflow.keras.layers import *
import tensorflow.keras.backend as K
import seaborn as sns
import pickle
from src.score import *
from collections import OrderedDict
# -
tf.__version__
# + Collapsed="false"
def limit_mem():
"""By default TF uses all available GPU memory. This function prevents this."""
config = tf.compat.v1.ConfigProto()
config.gpu_options.allow_growth = True
tf.compat.v1.Session(config=config)
# + Collapsed="false"
limit_mem()
# + Collapsed="false"
sns.set_style('darkgrid')
sns.set_context('notebook')
# + Collapsed="false"
DATADIR = '/data/WeatherBench/5.625deg/'
PREDDIR = '/data/WeatherBench/predictions/'
# + [markdown] Collapsed="false"
# ## Create data generator
#
# First up, we want to write our own Keras data generator. The key advantage to just feeding in numpy arrays is that we don't have to load the data twice because our intputs and outputs are the same data just offset by the lead time. Since the dataset is quite large and we might run out of CPU RAM this is important.
# + Collapsed="false"
# Load the validation subset of the data: 2017 and 2018
z500_valid = load_test_data(f'{DATADIR}geopotential_500', 'z')
t850_valid = load_test_data(f'{DATADIR}temperature_850', 't')
valid = xr.merge([z500_valid, t850_valid])
# + Collapsed="false"
z = xr.open_mfdataset(f'{DATADIR}geopotential_500/*.nc', combine='by_coords')
t = xr.open_mfdataset(f'{DATADIR}temperature_850/*.nc', combine='by_coords').drop('level')
# + Collapsed="false"
# For the data generator all variables have to be merged into a single dataset.
datasets = [z, t]
ds = xr.merge(datasets)
# + Collapsed="false"
# In this notebook let's only load a subset of the training data
ds_train = ds.sel(time=slice('2015', '2016'))
ds_test = ds.sel(time=slice('2017', '2018'))
# + Collapsed="false"
class DataGenerator(keras.utils.Sequence):
def __init__(self, ds, var_dict, lead_time, batch_size=32, shuffle=True, load=True, mean=None, std=None):
"""
Data generator for WeatherBench data.
Template from https://stanford.edu/~shervine/blog/keras-how-to-generate-data-on-the-fly
Args:
ds: Dataset containing all variables
var_dict: Dictionary of the form {'var': level}. Use None for level if data is of single level
lead_time: Lead time in hours
batch_size: Batch size
shuffle: bool. If True, data is shuffled.
load: bool. If True, datadet is loaded into RAM.
mean: If None, compute mean from data.
std: If None, compute standard deviation from data.
"""
self.ds = ds
self.var_dict = var_dict
self.batch_size = batch_size
self.shuffle = shuffle
self.lead_time = lead_time
data = []
generic_level = xr.DataArray([1], coords={'level': [1]}, dims=['level'])
for var, levels in var_dict.items():
try:
data.append(ds[var].sel(level=levels))
except ValueError:
data.append(ds[var].expand_dims({'level': generic_level}, 1))
self.data = xr.concat(data, 'level').transpose('time', 'lat', 'lon', 'level')
self.mean = self.data.mean(('time', 'lat', 'lon')).compute() if mean is None else mean
self.std = self.data.std('time').mean(('lat', 'lon')).compute() if std is None else std
# Normalize
self.data = (self.data - self.mean) / self.std
self.n_samples = self.data.isel(time=slice(0, -lead_time)).shape[0]
self.init_time = self.data.isel(time=slice(None, -lead_time)).time
self.valid_time = self.data.isel(time=slice(lead_time, None)).time
self.on_epoch_end()
# For some weird reason calling .load() earlier messes up the mean and std computations
if load: print('Loading data into RAM'); self.data.load()
def __len__(self):
'Denotes the number of batches per epoch'
return int(np.ceil(self.n_samples / self.batch_size))
def __getitem__(self, i):
'Generate one batch of data'
idxs = self.idxs[i * self.batch_size:(i + 1) * self.batch_size]
X = self.data.isel(time=idxs).values
y = self.data.isel(time=idxs + self.lead_time).values
return X, y
def on_epoch_end(self):
'Updates indexes after each epoch'
self.idxs = np.arange(self.n_samples)
if self.shuffle == True:
np.random.shuffle(self.idxs)
# + Collapsed="false"
# then we need a dictionary for all the variables and levels we want to extract from the dataset
dic = OrderedDict({'z': None, 't': None})
# + Collapsed="false"
bs=32
lead_time=6
# + Collapsed="false"
# Create a training and validation data generator. Use the train mean and std for validation as well.
dg_train = DataGenerator(
ds_train.sel(time=slice('2015', '2015')), dic, lead_time, batch_size=bs, load=True)
dg_valid = DataGenerator(
ds_train.sel(time=slice('2016', '2016')), dic, lead_time, batch_size=bs, mean=dg_train.mean, std=dg_train.std, shuffle=False)
# + Collapsed="false"
dg_train.mean, dg_train.std
# + Collapsed="false"
# Now also a generator for testing. Impartant: Shuffle must be False!
dg_test = DataGenerator(ds_test, dic, lead_time, batch_size=bs, mean=dg_train.mean, std=dg_train.std, shuffle=False)
# + [markdown] Collapsed="false"
# ## Create and train model
#
# Next up, we need to create the model architecture. Here we will use a fully connected convolutional network. Because the Earth is periodic in longitude, we want to use a periodic convolution in the lon-direction. This is not implemented in Keras, so we have to do it manually.
# + Collapsed="false"
class PeriodicPadding2D(tf.keras.layers.Layer):
def __init__(self, pad_width, **kwargs):
super().__init__(**kwargs)
self.pad_width = pad_width
def call(self, inputs, **kwargs):
if self.pad_width == 0:
return inputs
inputs_padded = tf.concat(
[inputs[:, :, -self.pad_width:, :], inputs, inputs[:, :, :self.pad_width, :]], axis=2)
# Zero padding in the lat direction
inputs_padded = tf.pad(inputs_padded, [[0, 0], [self.pad_width, self.pad_width], [0, 0], [0, 0]])
return inputs_padded
def get_config(self):
config = super().get_config()
config.update({'pad_width': self.pad_width})
return config
class PeriodicConv2D(tf.keras.layers.Layer):
def __init__(self, filters,
kernel_size,
conv_kwargs={},
**kwargs, ):
super().__init__(**kwargs)
self.filters = filters
self.kernel_size = kernel_size
self.conv_kwargs = conv_kwargs
if type(kernel_size) is not int:
assert kernel_size[0] == kernel_size[1], 'PeriodicConv2D only works for square kernels'
kernel_size = kernel_size[0]
pad_width = (kernel_size - 1) // 2
self.padding = PeriodicPadding2D(pad_width)
self.conv = Conv2D(
filters, kernel_size, padding='valid', **conv_kwargs
)
def call(self, inputs):
return self.conv(self.padding(inputs))
def get_config(self):
config = super().get_config()
config.update({'filters': self.filters, 'kernel_size': self.kernel_size, 'conv_kwargs': self.conv_kwargs})
return config
# + Collapsed="false"
def build_cnn(filters, kernels, input_shape, dr=0):
"""Fully convolutional network"""
x = input = Input(shape=input_shape)
for f, k in zip(filters[:-1], kernels[:-1]):
x = PeriodicConv2D(f, k)(x)
x = LeakyReLU()(x)
if dr > 0: x = Dropout(dr)(x)
output = PeriodicConv2D(filters[-1], kernels[-1])(x)
return keras.models.Model(input, output)
# + Collapsed="false"
cnn = build_cnn([64, 64, 64, 64, 2], [5, 5, 5, 5, 5], (32, 64, 2))
# + Collapsed="false"
cnn.compile(keras.optimizers.Adam(1e-4), 'mse')
# + Collapsed="false"
cnn.summary()
# + Collapsed="false"
# Since we didn't load the full data this is only for demonstration.
cnn.fit(dg_train, epochs=100, validation_data=dg_valid,
callbacks=[tf.keras.callbacks.EarlyStopping(
monitor='val_loss',
min_delta=0,
patience=2,
verbose=1,
mode='auto'
)]
)
# + Collapsed="false"
# cnn.save_weights('/home/rasp/cube_home/tmp/test.h5')
# + Collapsed="false"
# Load weights from externally trained model
cnn.load_weights('/data/weather-benchmark/predictions/saved_models/fccnn_6h.h5')
# + [markdown] Collapsed="false"
# ## Create predictions
#
# Now that we have our model we need to create a prediction NetCDF file. This function does this.
#
# We can either directly predict the target lead time (e.g. 5 days) or create an iterative forecast by chaining together many e.g. 6h forecasts.
# + Collapsed="false"
def create_predictions(model, dg):
"""Create predictions for non-iterative model"""
preds = model.predict_generator(dg)
# Unnormalize
preds = preds * dg.std.values + dg.mean.values
fcs = []
lev_idx = 0
for var, levels in dg.var_dict.items():
if levels is None:
fcs.append(xr.DataArray(
preds[:, :, :, lev_idx],
dims=['time', 'lat', 'lon'],
coords={'time': dg.valid_time, 'lat': dg.ds.lat, 'lon': dg.ds.lon},
name=var
))
lev_idx += 1
else:
nlevs = len(levels)
fcs.append(xr.DataArray(
preds[:, :, :, lev_idx:lev_idx+nlevs],
dims=['time', 'lat', 'lon', 'level'],
coords={'time': dg.valid_time, 'lat': dg.ds.lat, 'lon': dg.ds.lon, 'level': levels},
name=var
))
lev_idx += nlevs
return xr.merge(fcs)
# + Collapsed="false"
fc = create_predictions(cnn, dg_test)
# + Collapsed="false"
compute_weighted_rmse(fc, valid).compute()
# + Collapsed="false"
def create_iterative_predictions(model, dg, max_lead_time=5*24):
state = dg.data[:dg.n_samples]
preds = []
for _ in range(max_lead_time // dg.lead_time):
state = model.predict(state)
p = state * dg.std.values + dg.mean.values
preds.append(p)
preds = np.array(preds)
lead_time = np.arange(dg.lead_time, max_lead_time + dg.lead_time, dg.lead_time)
das = []; lev_idx = 0
for var, levels in dg.var_dict.items():
if levels is None:
das.append(xr.DataArray(
preds[:, :, :, :, lev_idx],
dims=['lead_time', 'time', 'lat', 'lon'],
coords={'lead_time': lead_time, 'time': dg.init_time, 'lat': dg.ds.lat, 'lon': dg.ds.lon},
name=var
))
lev_idx += 1
else:
nlevs = len(levels)
das.append(xr.DataArray(
preds[:, :, :, :, lev_idx:lev_idx+nlevs],
dims=['lead_time', 'time', 'lat', 'lon', 'level'],
coords={'lead_time': lead_time, 'time': dg.init_time, 'lat': dg.ds.lat, 'lon': dg.ds.lon, 'level': levels},
name=var
))
lev_idx += nlevs
return xr.merge(das)
# + Collapsed="false"
fc_iter = create_iterative_predictions(cnn, dg_test)
# + Collapsed="false"
rmse = evaluate_iterative_forecast(fc_iter, valid)
# + Collapsed="false"
rmse.load()
# + Collapsed="false"
rmse.z_rmse.plot()
# + Collapsed="false"
rmse.t_rmse.plot()
# + [markdown] Collapsed="false"
# # The end
| notebooks/3-cnn-example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/AlyssonBatista/Codigos-python/blob/main/deep_learning_TensorFlow.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="i-tXJoljwHZT"
# # Curso de Deep Learning com TensorFlow
# + id="bZdAPCsjvzvC"
import tensorflow as tf
import numpy as np
from tensorflow import keras
# + id="qCa63mz3wbBZ"
# uma camada de entrada
model = keras.Sequential([keras.layers.Dense(units=1,input_shape=[1])]) #units é a quantida de neurônios e imput shape é o formato dos dados de entrada
model.compile(optimizer='sgd',loss='mean_squared_error')
xs = np.array([-1.0, 0.0, 1.0, 2.0, 3.0, 4.0], dtype=float)
ys = np.array([-3.0, -1.0, 1.0, 3.0, 5.0, 7.0], dtype=float)# y = (2 * x) - 1
model.fit(xs,ys,epochs=600)
print(model.predict([10.0]))
# + [markdown] id="5pQ-GHmqAAMI"
# ## Classificação de imagens
#
# + colab={"base_uri": "https://localhost:8080/", "height": 941} id="g9QV72fbAZfO" outputId="4fa8d983-c187-4541-9d00-3fd29273a126"
# TensorFlow e tf.keras
import tensorflow as tf
from tensorflow import keras
# Librariesauxiliares
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
'''
Label Classe
0 Camisetas/Top (T-shirt/top)
1 Calça (Trouser)
2 Suéter (Pullover)
3 Vestidos (Dress)
4 Casaco (Coat)
5 Sandálias (Sandal)
6 Camisas (Shirt)
7 Tênis (Sneaker)
8 Bolsa (Bag)
9 Botas (Ankle boot)
'''
def main():
fashion_mnist = keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
print(train_images.shape)
print(len(train_labels))
print(train_labels)
print(test_images.shape)
print(len(test_labels))
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
plt.figure()
plt.imshow(train_images[0])
plt.colorbar()
plt.grid(False)
plt.show()
train_images = train_images / 255.0
test_images = test_images / 255.0
plt.figure(figsize=(10, 10))
for i in range(25):
plt.subplot(5, 5, i + 1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
plt.xlabel(class_names[train_labels[i]])
plt.show()
if __name__ == "__main__":
main()
# + colab={"base_uri": "https://localhost:8080/"} id="lo9pdI7XAD2D" outputId="11c13e00-3dc3-4424-8f1a-b9a022e25b0b"
import keras
def main():
fashion_mnist = keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
print(train_images)
print(train_labels)
print(test_images)
print(test_labels)
if __name__ == "__main__":
main()
# + id="gHo-OmBbAlIQ"
| deep_learning_TensorFlow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Table of contents:
#
# * [The Paillier Cryptosystem](#paillier)
# * [Key Generation](#keygeneration)
# * [Random prime numbers](#twop)
# * [Calculate $l$, $g$ and $\mu$](#lgmu)
# * [Encryption function](#encryption)
# * [Decryption function](#decryption)
#
# Author: [<NAME>](https://github.com/sebastiaagramunt) for [OpenMined](https://www.openmined.org/) Privacy ML Series course.
#
#
#
# # The Paillier Cryptosystem <a class="anchor" id="paillier"></a>
#
# The [Paillier cryptosystem](https://en.wikipedia.org/wiki/Paillier_cryptosystem) was invented and named after Pascal Paillier in 1999. In this notebook we will implement the Paillier cryptosystem from scratch but in an forthcomming class we willl check Paillier's cryptosystem homomorphic properties.
# ## Key Generation <a class="anchor" id="keygeneration"></a>
#
# * Generate two random primes $p$ and $q$
# * Calculate $N$, the product of $p$ and $q$
# * if $N$ and $\phi(N)$ have common factors, go back to first step and generate new $p$ and $q$
# * if $N$ and $\phi(N)$ don't share common factors (i.e. gcd is 1) then:
# * calculate $l$, the least common multiple of $p-1$ and $q-1$
# * calculate $N^2$
# * draw a random number $g$ in between 1 and $N^2$
# * calculate $\mu$ as the inverse of $L(g^l \textit{mod }N^2, N)$ in modulo $N$ where $L(x, n)=(x-1)/n$
#
# The public key is ($N$, $g$) and the private key is ($N$, $l$, $\mu$)
# ### Drawing two prime numbers <a class="anchor" id="twop"></a>
# +
from crypto import RandomPrime
from crypto import xgcd
size_bits = 16
p = RandomPrime(size_bits, m=40)
q = RandomPrime(size_bits, m=40)
while p==q:
q = RandomPrime(size_bits, m=40)
N = p*q
gcd, _, _ = xgcd(N, (p-1)*(q-1))
print(f"p = {p}")
print(f"q = {q}")
print(f"gcd(N, (p-1)*(q-1))={gcd}")
# -
# ### Calculating $l$, $g$ and $\mu$ <a class="anchor" id="lgmu"></a>
# +
from random import randrange
from crypto import LCM, InverseMod
def _L(x, n):
return (x-1)//n
l = LCM(p-1, q-1)
nsq = N*N
g = randrange(1, nsq)
mu = InverseMod(_L(pow(g, l, nsq), N), N)
print(f"l = {l}")
print(f"N^2 = {nsq}")
print(f"mu = {mu}")
# +
PublicKey = (N, g)
PrivateKey = (N, l, mu)
print(f"PublicKey = (N, g) = ({N}, {g})")
print(f"PrivateKey = (N, l, mu) = ({N}, {l}, {mu})")
# -
# ## Encryption function
#
# Take the public key ($N$, $g$) and the message you want to send $m$. Find a random number $r<N$ such that it has no common factors with $N$. Then compute the ciphertext $c$ as:
#
# $$c = g^{m}*r^{N}(\text{mod }N^2)$$
# +
m = randrange(0, N)
N, g = PublicKey[0], PublicKey[1]
gcd = 2
while gcd!=1:
r = randrange(1, N)
gcd, _, _ = xgcd(r, N)
c = pow(g, m, N*N)*pow(r, N, N*N)%(N*N)
print(f"m: {m}")
print(f"c: {c}")
# -
# ## Decryption function
#
# Take the private key ($N$, $l$, $\mu$) and the ciphertext $c$ and compute:
#
# $$m = L(c^l(\text{mod }N^2), N)*\mu (\text{mod }N)$$
# +
N, l, mu = PrivateKey[0], PrivateKey[1], PrivateKey[2]
m2 = _L(pow(c, l, N*N), N)*mu%N
print(f"Recovered message: {m2}")
# +
from typing import Tuple
def PaillierKeyGenerator(size: int = 64):
'''
Implementation of Paillier Cryptosystem
This function generates p .
ublic and private keys
Input:
size: size in bits of the field
Output:
PublicKey: (n, g)
PrivateKey: (n, l, mu)
'''
gcd = 2
while gcd!=1:
p = RandomPrime(size, 40)
q = RandomPrime(size, 40)
N = p*q
gcd, _, _ = xgcd(N, (p-1)*(q-1))
if gcd==1:
l = LCM(p-1, q-1)
nsq = N*N
g = randrange(1, nsq)
mu = InverseMod(_L(pow(g, l, nsq), N), N)
return (N, g), (N, l, mu)
def PaillierEncrypt(m: int, PublicKey: Tuple[int, int]):
'''
Encrypts a message m using the Paillier public key
Input:
m: message (An integer message) (mod n)
PublicKey: A tuple (N, g)
Output:
c: Encrypted message
'''
N, g = PublicKey[0], PublicKey[1]
gcd = 2
while gcd!=1:
r = randrange(1, N)
gcd, _, _ = xgcd(r, N)
return pow(g, m, N*N)*pow(r, N, N*N)%(N*N)
def PaillierDecrypt(c: int, PrivateKey: Tuple[int, int, int]):
'''
Decrypts a ciphertext m using the Paillier private key
Input:
m: message (An integer message) (mod n)
PublicKey: A tuple (n, l, mu)
Output:
m: Decrypted message
'''
N, l, mu = PrivateKey[0], PrivateKey[1], PrivateKey[2]
return _L(pow(c, l, N*N), N)*mu%N
# +
PublicKey, PrivateKey = PaillierKeyGenerator(32)
print(f"PublicKey = {PublicKey}")
print(f"PrivateKey = {PrivateKey}")
# +
m = randrange(0, N)
c = PaillierEncrypt(m, PublicKey)
m2 = PaillierDecrypt(c, PrivateKey)
print(f"message: {m}")
print(f"ciphertext: {c}")
print(f"recovered_message: {m2}")
# -
| Foundations_of_Private_Computation/Asymmetric_Cryptographic_Systems/notebooks/Paillier_Crytposystem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <font color='blue'>Data Science Academy - Python Fundamentos - Capítulo 4</font>
#
# ## Download: http://github.com/dsacademybr
# Versão da Linguagem Python
from platform import python_version
print('Versão da Linguagem Python Usada Neste Jupyter Notebook:', python_version())
# ## Enumerate
# Criando uma lista
seq = ['a','b','c']
enumerate(seq)
# #### Enumera os elementos
list(enumerate(seq))
# Imprimindo os valores de uma lista com a função enumerate() e seus respectivos índices
for indice, valor in enumerate(seq):
print (indice, valor)
for indice, valor in enumerate(seq):
if indice >= 2:
break
else:
print (valor)
lista = ['Marketing', 'Tecnologia', 'Business']
for i, item in enumerate(lista):
print(i, item)
for i, item in enumerate('Isso é uma string'):
print(i, item)
for i, item in enumerate(range(10)):
print(i, item)
# # FIM
relatorio = [len(palavra.split()) for palavra in ["djksjhsjn jsksjk dddd jhjhdjj"]]
relatorio
# Interessado(a) em conhecer os cursos e formações da DSA? Confira aqui nosso catálogo de cursos:
#
# https://www.datascienceacademy.com.br/pages/todos-os-cursos-dsa
# ### Obrigado - Data Science Academy - <a href="http://facebook.com/dsacademybr">facebook.com/dsacademybr</a>
| Cap04/Notebooks/DSA-Python-Cap04-10-Enumerate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv
# language: python
# name: venv
# ---
# # Stock Price Prediction
#
# ### Task
# * 모델 및 하이퍼파라미터들을 바꿔가며 accuracy를 높혀 보자
# * 밑에 제시된 여러가지 시도를 해보자
# * The main flow of this code is referenced in this [blog](https://medium.com/@aniruddha.choudhury94/stock-market-prediction-by-recurrent-neural-network-on-lstm-model-56de700bff68)
# * LG전자의 주식데이터를 직접 다운 받아서 실제 예측을 해보자
# * Train data: 2017년 1월 1일 ~ 2018년 12월 31일 데이터
# * Test data: 2019년 1월 1일 ~ 2019년 1월 31일 데이터
# * Close price 예측 (baseline은 open price)
#
# ### Dataset
# * [Yahoo finance datasets](https://www.imdb.com/interfaces/)
# * 2-3년간 daily stock price 데이터를 이용하여 미래 한달의 주식가격을 예측
#
# ### Baseline code
# * Dataset: train, test로 split
# * Input data shape: (`batch_size`, `past_day`=60, 1)
# * Output data shape: (`batch_size`, 1)
# * Architecture:
# * `LSTM` - `Dense`
# * [`tf.keras.layers`](https://www.tensorflow.org/api_docs/python/tf/keras/layers) 사용
# * Training
# * `model.fit` 사용
# * Evaluation
# * `model.evaluate` 사용 for test dataset
#
# ### Try some techniques
# * Change model architectures (Custom model)
# * Use another cells (LSTM, GRU, etc.)
# * Use dropout layers
# * Change the `past_day`
# * Data augmentation (if possible)
# * Try Early stopping
# * Use various features (open, high, low, close prices and volume features)
# ## Import modules
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import os
import sys
import time
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
from IPython import display
import tensorflow as tf
from tensorflow.keras import layers
tf.enable_eager_execution()
tf.logging.set_verbosity(tf.logging.INFO)
os.environ["CUDA_VISIBLE_DEVICES"]="0"
# -
# ## Data Download
#
# * We colud download daliy stock price using `fix_yahoo_finance` library.
# * Some stock ticker symbols [NASDAQ] [link](http://eoddata.com/symbols.aspx)
# * `AAPL`: Apple Inc.
# * `AMZM`: Amazon.com Inc.
# * `GOOG`: Alphabet Class C (Google)
# * `MSFT`: Microsoft Corp.
# +
import fix_yahoo_finance as yf
dataset = yf.download(tickers='AAPL', start='2016-01-01', end='2018-01-01', auto_adjust=True)
# -
dataset.head()
# ### Data Preprocessing
#
# 1. Data discretization: Part of data reduction but with particular importance, especially for numerical data
# 2. Data transformation: Normalization.
# 3. Data cleaning: Fill in missing values.
# 4. Data integration: Integration of data files.
#
# After the dataset is transformed into a clean dataset, the dataset is divided into training and testing sets so as to evaluate. Creating a data structure with 60 timesteps and 1 output
#Data cleaning
dataset.isna().any()
dataset.info()
dataset['Open'].plot(figsize=(16, 6))
dataset.Close.plot(figsize=(16, 6))
plt.show()
# +
# convert column to float type when column type is an object
#dataset["Close"] = dataset["Close"].str.replace(',', '').astype(float)
# -
# 7 day rolling mean
dataset.rolling(7).mean().head(20)
dataset['Close: 30 Day Mean'] = dataset.Close.rolling(window=30).mean()
dataset[['Close', 'Close: 30 Day Mean']].plot(figsize=(16, 6))
plt.show()
# ### Make a training dataset
train_data = dataset['Open']
train_data = pd.DataFrame(train_data)
# Feature Scaling
from sklearn.preprocessing import MinMaxScaler
sc = MinMaxScaler(feature_range = (0, 1))
train_data_scaled = sc.fit_transform(train_data)
plt.figure(figsize=(16, 6))
plt.plot(train_data_scaled)
plt.show()
# +
# Creating a data structure with 60 timesteps and 1 output
past_days = 60
X_train = []
y_train = []
for i in range(past_days, len(train_data_scaled)):
X_train.append(train_data_scaled[i-past_days:i, 0])
y_train.append(train_data_scaled[i, 0])
X_train, y_train = np.array(X_train), np.array(y_train)
# Reshaping
X_train = np.reshape(X_train, (X_train.shape[0], X_train.shape[1], 1))
# Cast
X_train = X_train.astype(np.float32)
y_train = y_train.astype(np.float32)
# -
# We predict the price of next day given past 60 days prices
print(X_train.shape)
print(y_train.shape)
# ## Build a model
model = tf.keras.Sequential()
# +
# Adding the first LSTM layer
model.add(layers.LSTM(units=50, input_shape=(past_days, 1)))
# Adding the output layer
model.add(layers.Dense(units=1))
# -
model.summary()
# Check for model
model(X_train[0:2])[0]
# Compiling the RNN
model.compile(optimizer=tf.train.AdamOptimizer(0.001),
loss='mean_squared_error')
# Fitting the RNN to the Training set
model.fit(X_train, y_train, epochs=10, batch_size=32)
# ## Performance on Test-Set
#
# Now that the model has been trained we can calculate its mean squared error on the test-set.
# +
# Part 3 - Making the predictions and visualising the results
# Getting the real stock price of 2018
dataset_test = yf.download(tickers='AAPL', start='2018-01-01', end='2018-02-01', auto_adjust=True)
# -
dataset_test.head()
dataset_test.info()
test_data = dataset_test['Open']
test_data = pd.DataFrame(test_data)
test_data.info()
# Feature Scaling
from sklearn.preprocessing import MinMaxScaler
sc = MinMaxScaler(feature_range = (0, 1))
test_set_scaled = sc.fit_transform(test_data)
test_set_scaled = pd.DataFrame(test_set_scaled)
test_set_scaled.head()
# +
# Getting the predicted stock price of 2018
dataset_total = pd.concat((dataset['Open'], dataset_test['Open']), axis = 0)
inputs = dataset_total[len(dataset_total) - len(dataset_test) - past_days:].values
inputs = inputs.reshape(-1,1)
inputs = sc.transform(inputs)
X_test = []
y_test = []
prediction_days = dataset_test.shape[0]
for i in range(past_days, past_days + prediction_days):
X_test.append(inputs[i-past_days:i, 0])
y_test.append(inputs[i, 0])
X_test, y_test = np.array(X_test), np.array(y_test)
X_test = np.reshape(X_test, (X_test.shape[0], X_test.shape[1], 1))
# Cast
X_test = X_test.astype(np.float32)
y_test = y_test.astype(np.float32)
# -
predicted_stock_price = model.predict(X_test)
predicted_stock_price = sc.inverse_transform(predicted_stock_price)
# Visualising the results
real_stock_price = dataset_test.Open.values
plt.plot(real_stock_price, color='red', label='Real Stock Price')
plt.plot(predicted_stock_price, color='blue', label='Predicted Stock Price')
plt.title('Stock Price Prediction')
plt.xlabel('Time')
plt.ylabel('Stock Price')
plt.legend()
plt.show()
# ### Evalueate for test dataset
# %%time
result = model.evaluate(X_test, y_test)
print("Mean Squared Error: {0:.2%}".format(result))
| rnn/stock_price_prediction/stock_price_prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Power of IPython Notebook + Pandas + and Scikit-learn
# IPython Notebook, Numpy, Pandas, MongoDB, R — for the better part of a year now, I have been trying out these technologies as part of Udacity's [Data Analyst Nanodegree](https://www.udacity.com/course/data-analyst-nanodegree--nd002). My undergrad education barely touched on data visualization or more broadly data science, and so I figured being exposed to the aforementioned technologies would be fun. And fun it has been, with R's powerful IDE-powered data mundging and visualization techniques having been particularly revelatory. I learned enough of R to create [some complex visualizations](http://www.andreykurenkov.com/writing/fun-visualizations-of-stackoverflow/), and was impressed by how easy is to import data into its Dataframe representations and then transform and visualize that data. I also thought RStudio's paradigm of continuously intermixed code editing and execution was superior to my habitual workflow of just endlessly cycling between tweaking and executing of Python scripts.
#
# Still, R is a not-quite-general-purpose-language and I hit upon multiple instances in which simple things were hard to do. In such times, I could not help but miss the powers of Python, a language I have tons of experience with and which is about as general purpose as it gets. Luckily, the courses also covered the equivalent of an R implementation for Python: the Python Data Analysis Library, Pandas. This let me use the features of R I now liked — dataframes, powerful plotting methods, elegant methods for transforming data — with Python's lovely syntax and libraries I already knew and loved. And soon I got to do just that, using both Pandas and the supremely good Machine Learning package Scikit-learn for the final project of [Udacity's Intro to Machine Learning Course](https://www.udacity.com/course/intro-to-machine-learning--ud120). Not only that, but I also used IPython Notebook for RStudio-esque intermixed code editing and execution and nice PDF output.
#
# I had such a nice experience with this combination of tools that I decided to dedicate a post to it, and what follows is mostly a summation of that experience. Reading it should be sufficient to get a general idea for why these tools are useful, whereas a much more detailed introdution and tutorial for Pandas can be found elsewhere (for instance [here](http://nbviewer.jupyter.org/github/fonnesbeck/pytenn2014_tutorial/blob/master/Part%201.%20Data%20Wrangling%20with%20Pandas.ipynb)). Incidentally, this whole post was written in IPython Notebook and the source of that [can be found here](http://www.andreykurenkov.com/writing/files/2016-06-10-power-of-ipython-pandas-scikilearn/post.ipynb) with the produced HTML [here](http://www.andreykurenkov.com/writing/files/2016-06-10-power-of-ipython-pandas-scikilearn/post.html).
# ## Data Summarization
# First, a bit about the project. The task was to first explore and clean a given dataset, and then train classification models using it. The dataset contained dozens of features about roughly 150 important employees from the [notoriously corrupt](https://en.wikipedia.org/wiki/Enron_scandal) company Enron, witch were classified as either a "Person of Interest" or not based on the outcome of investigations into Enron's corruption. It's a tiny dataset and not what I would have chosen, but such were the instructions. The data was provided in a bunch of Python dictionaries, and at first I just used a Python script to change it into a CSV and started exploring it in RStudio. But, it soon dawned on me that I would be much better off just working entirely in Python, and the following code is taken verbatim from my final project submission.
#
# And so, the code. Following some imports and a '%matplotlib notebook' comment to allow plotting within IPython, I loaded the data using pickle and printed out some basic things about it (not yet using Pandas):
import matplotlib.pyplot as plt
import matplotlib
import pickle
import pandas as pd
import numpy as np
from IPython.display import display
# %matplotlib notebook
# +
enron_data = pickle.load(open("./ud120-projects/final_project/final_project_dataset.pkl", "rb"))
print("Number of people: %d"%len(enron_data.keys()))
print("Number of features per person: %d"%len(list(enron_data.values())[0]))
print("Number of POI: %d"%sum([1 if x['poi'] else 0 for x in enron_data.values()]))
# -
# But working with this set of dictionaries would not be nearly as fast or easy as a Pandas dataframe, so I soon converted it to that and went ahead and summarized all the features with a single method call:
# +
df = pd.DataFrame.from_dict(enron_data)
del df['TOTAL']
df = df.transpose()
numeric_df = df.apply(pd.to_numeric, errors='coerce')
del numeric_df['email_address']
numeric_df.describe()
# -
# Looking through these, I found one instance of a valid outlier - <NAME> (CEO of Enron), and removed him from the dataset.
#
# I should emphasize the benefits of doing all this in IPython Notebook. Being able to tweak parts of the code without reexecuting all of it and reloading all the data made iterating on ideas much faster, and iterating on ideas fast is essential for exploratory data analysis and development of machine learned models. It's no accident that the Matlab IDE and RStudio, both tools commonly used in the sciences for data processing and analysis, have essentially the same structure. I did not understand the benefits of IPython Notebook when I was first made to use it for class assignments in College, but now it has finally dawned on me that it fills the same role as those IDEs and became popular because it is similaly well suited for working with data.
# +
del numeric_df['loan_advances']
del numeric_df['restricted_stock_deferred']
del numeric_df['director_fees']
std = numeric_df.apply(lambda x: np.abs(x - x.mean()) / x.std())
std = std.fillna(std.mean())
std.describe()
# -
# This result suggested that most features have large outliers (larger than 3 standard deviations). In order to be careful not to remove any useful data, I manually inspected all rows with large outliers to see any values that seem appropriate for removal:
outliers = std.apply(lambda x: x > 5).any(axis=1)
outlier_df = pd.DataFrame(index=numeric_df[outliers].index)
for col in numeric_df.columns:
outlier_df[str((col,col+'_std'))] = list(zip(numeric_df[outliers][col],std[outliers][col]))
display(outlier_df)
numeric_df.drop('FREVERT MARK A',inplace=True)
df.drop('FREVERT MARK A',inplace=True)
# Looking through these, I found one instance of a valid outlier - <NAME> (CEO of Enron), and removed him from the dataset.
#
# I should emphasize the benefits of doing all this in IPython Notebook. Being able to tweak parts of the code without reexecuting all of it and reloading all the data made iterating on ideas much faster, and iterating on ideas fast is essential for exploratory data analysis and development of machine learned models. It's no accident that the Matlab IDE and RStudio, both tools commonly used in the sciences for data processing and analysis, have essentially the same structure. I did not understand the benefits of IPython Notebook when I was first made to use it for class assignments in College, but now it has finally dawned on me that it fills the same role as those IDEs and became popular because it is similaly well suited for working with data.
# ## Feature Visualization, Engineering and Selection
# The project also instructed me to choose a set of features, and to engineer some of my own. In order to get an initial idea of possible promising features and how I could use them to create new features, I computed the correlation of each feature to the Person of Interest classification:
corr = numeric_df.corr()
print('\nCorrelations between features to POI:\n ' +str(corr['poi']))
# The results indicated that 'exercised_stock_options', 'total_stock_value', and 'bonus' are the most promising features. Just for fun, I went ahead and plotted these features to see if I could visually verify their significance:
numeric_df.hist(column='exercised_stock_options',by='poi',bins=25,sharex=True,sharey=True)
plt.suptitle("exercised_stock_options by POI")
numeric_df.hist(column='total_stock_value',by='poi',bins=25,sharex=True,sharey=True)
plt.suptitle("total_stock_value by POI")
numeric_df.hist(column='bonus',by='poi',bins=25,sharex=True,sharey=True)
plt.suptitle("bonus by POI")
# As well as one that is not strongly correlated:
numeric_df.hist(column='to_messages',by='poi',bins=25,sharex=True,sharey=True)
plt.suptitle("to_messages by POI")
# The data and plots above indicated that the exercised_stock_options, total_stock_value, and restricted_stock, and to a lesser extent to payment related information (total_payments, salary, bonus, and expenses), are all correlated to Persons of Interest. Therefore, I created new features as sums and ratios of these ones. Working with Pandas made this incredibely easy due to vectorized operations, and though Numpy could similarly make this easy I think Pandas' Dataframe construct makes it especially easy.
#
# It was also easy to fix any problems with the data before starting to train machine learning models. In order to use the data for evaluation and training, I replaced null values with the mean of each feature so as to be able to use the dataset with Scikit-learn. I also scaled all features to a range of 1-0, to better work with Support Vector Machines:
# +
#Get rid of label
del numeric_df['poi']
poi = df['poi']
#Create new features
numeric_df['stock_sum'] = numeric_df['exercised_stock_options'] +\
numeric_df['total_stock_value'] +\
numeric_df['restricted_stock']
numeric_df['stock_ratio'] = numeric_df['exercised_stock_options']/numeric_df['total_stock_value']
numeric_df['money_total'] = numeric_df['salary'] +\
numeric_df['bonus'] -\
numeric_df['expenses']
numeric_df['money_ratio'] = numeric_df['bonus']/numeric_df['salary']
numeric_df['email_ratio'] = numeric_df['from_messages']/(numeric_df['to_messages']+numeric_df['from_messages'])
numeric_df['poi_email_ratio_from'] = numeric_df['from_poi_to_this_person']/numeric_df['to_messages']
numeric_df['poi_email_ratio_to'] = numeric_df['from_this_person_to_poi']/numeric_df['from_messages']
#Feel in NA values with 'marker' value outside range of real values
numeric_df = numeric_df.fillna(numeric_df.mean())
#Scale to 1-0
numeric_df = (numeric_df-numeric_df.min())/(numeric_df.max()-numeric_df.min())
# -
# Then, I scored features using Scikit-learn's SelectKBest to get an ordering of them to test with multiple algorithms afterward. Pandas Dataframes can be used directly with Scikit-learn, which is another great benefit of it:
from sklearn.feature_selection import SelectKBest
selector = SelectKBest()
selector.fit(numeric_df,poi.tolist())
scores = {numeric_df.columns[i]:selector.scores_[i] for i in range(len(numeric_df.columns))}
sorted_features = sorted(scores,key=scores.get, reverse=True)
for feature in sorted_features:
print('Feature %s has value %f'%(feature,scores[feature]))
# It appeared that several of my features are among the most useful, as 'poi_email_ratio_to', 'stock_sum', and 'money_total' are all ranked highly. But, since the data is so small I had no need to get rid of any of the features and went ahead with testing several classifiers with several sets of features.
# # Training and Evaluating Models
# Proceding with the project, I selected three algorithms to test and compare: Naive Bayes, Decision Trees, and Support Vector Machines. Naive Bayes is a good baseline for any ML task, and the other two fit well into the task of binary classification with many features and can both be automatically tuned using sklearn classes. A word on SkLearn: it is simply a very well designed Machine Learning toolkit, with great compatibility with Numpy (and therefore also Pandas) and an elegant and smart API structure that makes trying out different models and evaluating features and just about anything one might want short of Deep Learning easy.
#
# I think the code that follows will attest to that. I tested those three algorithms with a variable number of features, from one to all of them ordered by the SelectKBest scoring. Because the data is so small, I could afford an extensive validation scheme and did multiple random splits of the data into training and testing to get an average that best indicated the strength of each algorithm. I also went ahead and evaluated precision and recall besides accuracy, since those were to be the metric of performance. And all it took to do all that is maybe 50 lines of code:
# +
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
from sklearn.grid_search import RandomizedSearchCV, GridSearchCV
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import precision_score, recall_score, accuracy_score
from sklearn.cross_validation import StratifiedShuffleSplit
import scipy
import warnings
warnings.filterwarnings('ignore')
gnb_clf = GridSearchCV(GaussianNB(),{})
#No params to tune for for linear bayes, use for convenience
svc_clf = SVC()
svc_search_params = {'C': scipy.stats.expon(scale=1),
'gamma': scipy.stats.expon(scale=.1),
'kernel': ['linear','poly','rbf'],
'class_weight':['balanced',None]}
svc_search = RandomizedSearchCV(svc_clf,
param_distributions=svc_search_params,
n_iter=25)
tree_clf = DecisionTreeClassifier()
tree_search_params = {'criterion':['gini','entropy'],
'max_leaf_nodes':[None,25,50,100,1000],
'min_samples_split':[2,3,4],
'max_features':[0.25,0.5,0.75,1.0]}
tree_search = GridSearchCV(tree_clf,
tree_search_params,
scoring='recall')
search_methods = [gnb_clf,svc_search,tree_search]
average_accuracies = [[0],[0],[0]]
average_precision = [[0],[0],[0]]
average_recall = [[0],[0],[0]]
num_splits = 10
train_split = 0.9
indices = list(StratifiedShuffleSplit(poi.tolist(),
num_splits,
test_size=1-train_split,
random_state=0))
best_features = None
max_score = 0
best_classifier = None
num_features = 0
for num_features in range(1,len(sorted_features)+1):
features = sorted_features[:num_features]
feature_df = numeric_df[features]
for classifier_idx in range(3):
sum_values = [0,0,0]
#Only do parameter search once, too wasteful to do a ton
search_methods[classifier_idx].fit(feature_df.iloc[indices[0][0],:],
poi[indices[0][0]].tolist())
classifier = search_methods[classifier_idx].best_estimator_
for split_idx in range(num_splits):
train_indices, test_indices = indices[split_idx]
train_data = (feature_df.iloc[train_indices,:],poi[train_indices].tolist())
test_data = (feature_df.iloc[test_indices,:],poi[test_indices].tolist())
classifier.fit(train_data[0],train_data[1])
predicted = classifier.predict(test_data[0])
sum_values[0]+=accuracy_score(predicted,test_data[1])
sum_values[1]+=precision_score(predicted,test_data[1])
sum_values[2]+=recall_score(predicted,test_data[1])
avg_acc,avg_prs,avg_recall = [val/num_splits for val in sum_values]
average_accuracies[classifier_idx].append(avg_acc)
average_precision[classifier_idx].append(avg_prs)
average_recall[classifier_idx].append(avg_recall)
score = (avg_prs+avg_recall)/2
if score>max_score and avg_prs>0.3 and avg_recall>0.3:
max_score = score
best_features = features
best_classifier = search_methods[classifier_idx].best_estimator_
print('Best classifier found is %s \n\
with score (recall+precision)/2 of %f\n\
and feature set %s'%(str(best_classifier),max_score,best_features))
# -
# Then, I could go right back to Pandas to plot the results. Sure, I could do this with matplotlib just as well, but the flexibility and simplicity of the 'plot' function call on a DataFrame makes it much less annoying to use in my opinion.
results = pd.DataFrame.from_dict({'Naive Bayes': average_accuracies[0],
'SVC':average_accuracies[1],
'Decision Tree':average_accuracies[2]})
results.plot(xlim=(1,len(sorted_features)-1),ylim=(0,1))
plt.suptitle("Classifier accuracy by # of features")
results = pd.DataFrame.from_dict({'Naive Bayes': average_precision[0],
'SVC':average_precision[1],
'Decision Tree':average_precision[2]})
results.plot(xlim=(1,len(sorted_features)-1),ylim=(0,1))
plt.suptitle("Classifier precision by # of features")
results = pd.DataFrame.from_dict({'Naive Bayes': average_recall[0],
'SVC':average_recall[1],
'Decision Tree':average_recall[2]})
results.plot(xlim=(1,len(sorted_features)-1),ylim=(0,1))
plt.suptitle("Classifier recall by # of features")
# As output by my code, the best algorithm was consistently found to be Decision Trees and so I could finally finish up the project by submitting that as my model.
# ## Conclusion
# I did not much care for the project's dataset and overall structure, but I still greatly enjoyed completing it because of how fun it was to combine Pandas data processing with Scikit-learn model training in the process, with IPython Notebook making that process even more fluid. While not at all a well written introduction or tutorial for these packages, I do hope that this write up about a single project I finished using them might inspire some readers to try out doing that as well.
| writing/files/2016-06-10-power-of-ipython-pandas-scikilearn/post.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="hW1HXwdcEbyq"
# Linear regression : Linear regression is a basic and commonly used type of predictive analysis. The overall idea of
# regression is to examine two things: (1) does a set of predictor variables do a good job in predicting an outcome
# (dependent) variable? (2) Which variables in particular are significant predictors of the outcome variable, and in
# what way do they–indicated by the magnitude and sign of the beta estimates–impact the outcome variable? These regression estimates
# are used to explain the relationship between one dependent variable and one or more independent variables. The simplest form of the
# regression equation with one dependent and one independent variable is defined by the formula y = c + b*x, where y = estimated dependent
# variable score, c = constant, b = regression coefficient, and x = score on the independent variable.
# Linear regression in Python
# Step 1: Import packages and classes
# The first step is to import the package numpy and the class LinearRegression from sklearn.linear_model:
import numpy as np
from sklearn.linear_model import LinearRegression
# + id="iId_Tit9Emua"
# Step 2: Provide data
# The second step is defining data to work with. The inputs (regressors, 𝑥)
# and output (predictor, 𝑦) should be arrays (the instances of the class numpy.ndarray) or
# similar objects. This is the simplest way of providing data for regression:
x = np.array([5, 15, 25, 35, 45, 55]).reshape((-1, 1))
y = np.array([5, 20, 14, 32, 22, 38])
# Now, you have two arrays: the input x and output y. You should call .reshape() on x because this array
# is required to be two-dimensional, or to be more precise, to have one column and as many rows as necessary.
# + id="ZrHas74EHuib"
# Step 3: Create a model and fit it
# The next step is to create a linear regression model and fit it using the existing data.
# Let’s create an instance of the class LinearRegression, which will represent the regression model:
model = LinearRegression()
# This statement creates the variable model as the instance of LinearRegression. You can provide several optional parameters to LinearRegression:
# fit_intercept is a Boolean (True by default) that decides whether to calculate the intercept 𝑏₀ (True) or consider it equal to zero (False).
# normalize is a Boolean (False by default) that decides whether to normalize the input variables (True) or not (False).
# copy_X is a Boolean (True by default) that decides whether to copy (True) or overwrite the input variables (False).
# n_jobs is an integer or None (default) and represents the number of jobs used in parallel computation. None usually means one job and -1 to use all processors.
# + id="0CVyLferHzCX" outputId="78275138-307e-41b1-dbcd-7615fd305742" colab={"base_uri": "https://localhost:8080/"}
# Step 3: With .fit(), you calculate the optimal values of the weights 𝑏₀ and 𝑏₁, using the existing input
# and output (x and y) as the arguments. In other words, .fit() fits the model. It returns self, which is the variable model itself.
model.fit(x, y)
# + id="AuIDMb79H1-n" outputId="8ab06af1-3c1c-4d5b-e774-2994dfaa4d27" colab={"base_uri": "https://localhost:8080/"}
# Step 4: Get results
# Once you have your model fitted, you can get the results to check whether the model works satisfactorily and interpret it.
# You can obtain the coefficient of determination (𝑅²) with .score() called on model:
r_sq = model.score(x, y)
print('coefficient of determination:', r_sq)
# + id="d79dvs-uH7N5"
| Machine Learning/LinearRegression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="lwnGnFN2kGkQ" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1593517250607, "user_tz": -180, "elapsed": 652, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11873232984457614152"}}
import cv2
import matplotlib.pyplot as plt
import numpy as np
from os import walk
import xlwt
from xlwt import Workbook
import tensorflow as tf
from tensorflow import keras
from sklearn.cluster import KMeans
# + id="JUPR-vo6jS2J" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} executionInfo={"status": "ok", "timestamp": 1593517250878, "user_tz": -180, "elapsed": 909, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11873232984457614152"}} outputId="9df7dffc-2e15-4781-aaca-50e2c6013a17"
from keras.models import load_model
model = load_model("/content/drive/My Drive/project1/classifier2.h5")
# + id="SWi7BysMj8hk" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1593517251148, "user_tz": -180, "elapsed": 1169, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11873232984457614152"}}
model.compile(loss="categorical_crossentropy",optimizer=tf.keras.optimizers.Adam(learning_rate=0.0001),metrics=["accuracy"])
# + id="--Y2iT0CdLta" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1593517251148, "user_tz": -180, "elapsed": 1157, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11873232984457614152"}}
def detect_white_fg(image_part):
sayac = 0
for i in range(0,20,1):
for j in range(0,20,1):
point = np.array(image_part[i,j,:]).reshape((3,))
if ((point[0]<240 and point[0]>225) and (point[1]<240 and point[1]>225) and (point[2]<240 and point[2]>225)):
sayac += 1
if (sayac>110):
return True
else:
return False
# + id="G1WHnoOml-gL" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1593517251149, "user_tz": -180, "elapsed": 1147, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11873232984457614152"}}
def erase_bg(white_fg_c,centroids,hist):
n,m = centroids.shape
sayac = 0
sayac2 = 0
if (white_fg_c == True):
for i in range(0,n-1,1):
j = i+1
c1 = np.array(centroids[i-sayac2,:]).reshape((3,))
c2 = np.array(centroids[j-sayac2,:]).reshape((3,))
if (c1[0]-c2[0]>=0) and (c1[0]-c2[0]>=0) and (c1[0]-c2[0]>=0) and (c1.max()>=220 and c1.sum()>=680) and :
centroids = np.delete(centroids,j-sayac2,0)
hist = np.delete(hist,j-sayac2,0)
sayac2 += 1
else:
centroids = np.delete(centroids,i-sayac2,0)
hist = np.delete(hist,i-sayac2,0)
sayac2 += 1
else:
n,m = centroids.shape
for i in range(0,n,1):
k,l = centroids.shape
if (k==1):
break
c3 = np.array(centroids[i,:]).reshape((3,))
if (c3.sum()<=720 and c3.sum()>=680) and (c3.min()>=220):
centroids = np.delete(centroids,i-sayac,0)
hist = np.delete(hist,i-sayac,0)
sayac += 1
centroids = centroids.reshape(-1,3)
n,m = centroids.shape
if (n==3):
centroids = centroids[0:3,:]
hist = hist[0:3]
centroids = centroids.reshape(-1,3)
return (centroids,hist)
# + id="8B4bvs5HO2ku" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1593517251150, "user_tz": -180, "elapsed": 1130, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11873232984457614152"}}
def cluster_colors(image):
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
### VARIABLES
wn_clusters = 4
### WHITE FG DETECTION
image_part = image[60:80,50:70,:]
white_fg_c = detect_white_fg(image_part)
### KMEANS CLUSTERING
image = image.reshape((image.shape[0] * image.shape[1], 3))
clt = KMeans(n_clusters = wn_clusters)
clt.fit(image)
### HISTOGRAM
numLabels = np.arange(0, len(np.unique(clt.labels_)) + 1)
(hist, _) = np.histogram(clt.labels_, bins = numLabels)
hist = hist.astype("float")
### PLOTTING
centroids = clt.cluster_centers_
### BACKGROUND ELECTION
centroids,hist = erase_bg(white_fg_c,centroids,hist)
hist /= hist.sum()
bar = np.zeros((50, 300, 3), dtype = "uint8")
startX = 0
for (percent, color) in zip(hist, centroids):
# plot the relative percentage of each cluster
endX = startX + (percent * 300)
cv2.rectangle(bar, (int(startX), 0), (int(endX), 50),color.astype("uint8").tolist(), -1)
startX = endX
return (bar,centroids,hist)
# + id="OLxX4Oc4Rtos" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1593517251151, "user_tz": -180, "elapsed": 1122, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11873232984457614152"}}
def write_img(image,result,i):
image = cv2.cvtColor(image,cv2.COLOR_RGB2BGR)
if result == 0:
address = "/content/drive/My Drive/rtest3saves/gomlek/"+str(i)+".jpg"
elif result == 1:
address = "/content/drive/My Drive/rtest3saves/pantolon/"+str(i)+".jpg"
elif result == 2:
address = "/content/drive/My Drive/rtest3saves/t-shirt/"+str(i)+".jpg"
cv2.imwrite(address,image)
return address
# + id="OOWEHL30Cums" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1593517251152, "user_tz": -180, "elapsed": 1115, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11873232984457614152"}}
wb = Workbook()
sheet = wb.add_sheet("Rapor",cell_overwrite_ok=True)
def sheet_write(result,centroid,i,address,hist):
sheet.write(i+1,0,address)
if result == 0:
sheet.write(i+1,1, "gomlek")
elif result == 1:
sheet.write(i+1,1, "pantolon")
elif result == 2:
sheet.write(i+1,1, "t-shirt")
(n,m) = centroid.shape
sayac1=0
for countit in range(2,(3*2+2),3):
if (n==1 and sayac1==1):
sheet.write(i+1,countit,"0")
sheet.write(i+1,countit+1,"0")
sheet.write(i+1,countit+2,"0")
else:
sheet.write(i+1,countit,str(centroid[sayac1,0]))
sheet.write(i+1,countit+1,str(centroid[sayac1,1]))
sheet.write(i+1,countit+2,str(centroid[sayac1,2]))
sayac1 = sayac1 + 1
sayac2=0
for countit2 in range(3*2+2,7*2+2,4):
if (n==1 and sayac2==1):
sheet.write(i+1,countit2,"0")
sheet.write(i+1,countit2+1,"0")
sheet.write(i+1,countit2+2,"0")
sheet.write(i+1,countit2+3,"0")
else:
cmyk_codes = list(rgb_to_cmyk(centroid[sayac2,:]/255))
sheet.write(i+1,countit2,str(cmyk_codes[0]))
sheet.write(i+1,countit2+1,str(cmyk_codes[1]))
sheet.write(i+1,countit2+2,str(cmyk_codes[2]))
sheet.write(i+1,countit2+3,str(cmyk_codes[3]))
sayac2 = sayac2 + 1
if (n==1):
sheet.write(i+1,7*2+2,"1.0")
sheet.write(i+1,7*2+3,"0.0")
else:
sheet.write(i+1,7*2+2,str(hist[0]))
sheet.write(i+1,7*2+3,str(hist[1]))
sheet.write(i+1,7*2+4,str(n))
# + id="7weg1pogDW8j" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1593517251152, "user_tz": -180, "elapsed": 1105, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11873232984457614152"}}
def rgb_to_cmyk(centro):
k = 1 - centro.max()
c = (1 - centro[0] - k) / (1 - k)
m = (1 - centro[1] - k) / (1 - k)
y = (1 - centro[2] - k) / (1 - k)
return (c,m,y,k)
# + id="IYa6jWSkJPTN" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1593517285001, "user_tz": -180, "elapsed": 34945, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11873232984457614152"}}
path_list = []
for (dirpath, dirnames, filenames) in walk("/content/drive/My Drive/rtest3"):
path_list.extend(filenames)
break
results = []
for i in range(0,20,1):
path = dirpath + '/' + path_list[i]
image = cv2.imread(path)
bar,centroid,hist = cluster_colors(image)
image = cv2.cvtColor(image,cv2.COLOR_BGR2RGB)
image = cv2.resize(src=image,dsize=(125,100))
image = image.reshape((1,100,125,3))
result = model.predict_classes(image)
image = image.reshape((100,125,3))
address = write_img(image,result,i)
sheet_write(result,centroid,i,address,hist)
results.extend(result)
# + id="YTFUq_o3GXXo" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1593517285002, "user_tz": -180, "elapsed": 34935, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11873232984457614152"}}
wb.save("/content/drive/My Drive/project1/rapor44.xls")
| Project_MAIN/Old_Version/Project_V2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="LQNi8aIsijzk"
# # Identification of zero-inflated genes
# + [markdown] colab_type="text" id="W7cITGhKijzn"
# AutoZI is a deep generative model adapted from scVI allowing a gene-specific treatment of zero-inflation. For each gene $g$, AutoZI notably learns the distribution of a random variable $\delta_g$ which denotes the probability that gene $g$ is not zero-inflated. In this notebook, we present the use of the model on a PBMC dataset.
#
# More details about AutoZI can be found in : https://www.biorxiv.org/content/10.1101/794875v2
# + colab={"base_uri": "https://localhost:8080/", "height": 382} colab_type="code" id="kTvfZsr5jBT6" outputId="ec8f5e92-6210-4b12-8725-6f0d24fab440"
# !pip install --quiet scvi-colab
from scvi_colab import install
install()
# + [markdown] colab_type="text" id="mC-uVd3zijz7"
# ## Imports, data loading and preparation
# + colab={} colab_type="code" id="05Y2uUIxijz9"
import numpy as np
import pandas as pd
import anndata
import scanpy as sc
import scvi
# + colab={"base_uri": "https://localhost:8080/", "height": 330} colab_type="code" id="j27g4M4Uij0F" outputId="3af26218-d43a-437a-c5e4-a35ec5093177"
pbmc = scvi.data.pbmc_dataset()
pbmc.layers["counts"] = pbmc.X.copy()
sc.pp.normalize_total(pbmc, target_sum=10e4)
sc.pp.log1p(pbmc)
pbmc.raw = pbmc
scvi.data.poisson_gene_selection(
pbmc,
n_top_genes=1000,
batch_key="batch",
subset=True,
layer="counts",
)
scvi.model.AUTOZI.setup_anndata(
pbmc,
labels_key="str_labels",
batch_key="batch",
layer="counts",
)
# + [markdown] colab_type="text" id="cYPYNAwRij0M"
# ## Analyze gene-specific ZI
# + [markdown] colab_type="text" id="6ovhk47Oij0N"
# In AutoZI, all $\delta_g$'s follow a common $\text{Beta}(\alpha,\beta)$ prior distribution where $\alpha,\beta \in (0,1)$ and the zero-inflation probability in the ZINB component is bounded below by $\tau_{\text{dropout}} \in (0,1)$. AutoZI is encoded by the `AutoZIVAE` class whose inputs, besides the size of the dataset, are $\alpha$ (`alpha_prior`), $\beta$ (`beta_prior`), $\tau_{\text{dropout}}$ (`minimal_dropout`). By default, we set $\alpha = 0.5, \beta = 0.5, \tau_{\text{dropout}} = 0.01$.
#
# Note : we can learn $\alpha,\beta$ in an Empirical Bayes fashion, which is possible by setting `alpha_prior = None` and `beta_prior = None`
# + colab={} colab_type="code" id="khMZvSw5ij0O"
vae = scvi.model.AUTOZI(pbmc)
# + [markdown] colab_type="text" id="HXOSTMrLij0V"
# We fit, for each gene $g$, an approximate posterior distribution $q(\delta_g) = \text{Beta}(\alpha^g,\beta^g)$ (with $\alpha^g,\beta^g \in (0,1)$) on which we rely. We retrieve $\alpha^g,\beta^g$ for all genes $g$ (and $\alpha,\beta$, if learned) as numpy arrays using the method `get_alphas_betas` of `AutoZIVAE`.
# + colab={"base_uri": "https://localhost:8080/", "height": 173, "referenced_widgets": ["4d7ac4d4d06b46f68e47d51f45f3d93a", "086bfdcc78404f66be5ec5233b85067c"]} colab_type="code" id="__RPOTF2ij0W" outputId="3c17fcfb-f5fc-40e3-f13b-b88d953ec6b1"
vae.train(max_epochs=200, plan_kwargs = {'lr':1e-2})
# + colab={} colab_type="code" id="B-lEw2IWij0a"
outputs = vae.get_alphas_betas()
alpha_posterior = outputs['alpha_posterior']
beta_posterior = outputs['beta_posterior']
# + [markdown] colab_type="text" id="BXrym0Cgij0e"
# Now that we obtained fitted $\alpha^g,\beta^g$, different metrics are possible. Bayesian decision theory suggests us the posterior probability of the zero-inflation hypothesis $q(\delta_g < 0.5)$, but also other metrics such as the mean wrt $q$ of $\delta_g$ are possible. We focus on the former. We decide that gene $g$ is ZI if and only if $q(\delta_g < 0.5)$ is greater than a given threshold, say $0.5$. We may note that it is equivalent to $\alpha^g < \beta^g$. From this we can deduce the fraction of predicted ZI genes in the dataset.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="h2kXAcZLij0e" outputId="ea7f42b6-49f7-448b-cbaa-0b1b0bf5d9b7"
from scipy.stats import beta
# Threshold (or Kzinb/Knb+Kzinb in paper)
threshold = 0.5
# q(delta_g < 0.5) probabilities
zi_probs = beta.cdf(0.5, alpha_posterior, beta_posterior)
# ZI genes
is_zi_pred = (zi_probs > threshold)
print('Fraction of predicted ZI genes :', is_zi_pred.mean())
# + [markdown] colab_type="text" id="Ps5oykK0ij0k"
# We noted that predictions were less accurate for genes $g$ whose average expressions - or predicted NB means, equivalently - were low. Indeed, genes assumed not to be ZI were more often predicted as ZI for such low average expressions. A threshold of 1 proved reasonable to separate genes predicted with more or less accuracy. Hence we may want to focus on predictions for genes with average expression above 1.
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="6AFreZzjij0l" outputId="302b439d-028f-4264-e9c7-04ca6c194ba7"
mask_sufficient_expression = (np.array(pbmc.X.mean(axis=0)) > 1.).reshape(-1)
print('Fraction of genes with avg expression > 1 :', mask_sufficient_expression.mean())
print('Fraction of predicted ZI genes with avg expression > 1 :', is_zi_pred[mask_sufficient_expression].mean())
# + [markdown] colab_type="text" id="B1V3X_GSij0p"
# ## Analyze gene-cell-type-specific ZI
# + [markdown] colab_type="text" id="uUmJ6hStij0q"
# One may argue that zero-inflation should also be treated on the cell-type (or 'label') level, in addition to the gene level. AutoZI can be extended by assuming a random variable $\delta_{gc}$ for each gene $g$ and cell type $c$ which denotes the probability that gene $g$ is not zero-inflated in cell-type $c$. The analysis above can be extended to this new scale.
# + colab={"base_uri": "https://localhost:8080/", "height": 173, "referenced_widgets": ["a6621ea35c23422da082b161388e265c", "e76db661573a4d5cb20cc1e31c7717f5"]} colab_type="code" id="1pr5WP_Eij0q" outputId="5ca84813-93f1-4fd2-d1da-1b5d5e7402bb"
# Model definition
vae_genelabel = scvi.model.AUTOZI(
pbmc,
dispersion='gene-label',
zero_inflation='gene-label'
)
# Training
vae_genelabel.train(max_epochs=200, plan_kwargs = {'lr':1e-2})
# Retrieve posterior distribution parameters
outputs_genelabel = vae_genelabel.get_alphas_betas()
alpha_posterior_genelabel = outputs_genelabel['alpha_posterior']
beta_posterior_genelabel = outputs_genelabel['beta_posterior']
# + colab={"base_uri": "https://localhost:8080/", "height": 330} colab_type="code" id="1wu__h5lij0v" outputId="89ef3bd7-ddeb-4264-94f7-b1e41d92ceb6"
# q(delta_g < 0.5) probabilities
zi_probs_genelabel = beta.cdf(0.5,alpha_posterior_genelabel, beta_posterior_genelabel)
# ZI gene-cell-types
is_zi_pred_genelabel = (zi_probs_genelabel > threshold)
ct = pbmc.obs.str_labels.astype("category")
codes = np.unique(ct.cat.codes)
cats = ct.cat.categories
for ind_cell_type, cell_type in zip(codes, cats):
is_zi_pred_genelabel_here = is_zi_pred_genelabel[:,ind_cell_type]
print('Fraction of predicted ZI genes for cell type {} :'.format(cell_type),
is_zi_pred_genelabel_here.mean(),'\n')
# + colab={"base_uri": "https://localhost:8080/", "height": 486} colab_type="code" id="ItgDITuBij02" outputId="f99f45ce-a9fd-4839-c944-80d7c7152443"
# With avg expressions > 1
for ind_cell_type, cell_type in zip(codes, cats):
mask_sufficient_expression = (np.array(pbmc.X[pbmc.obs.str_labels.values.reshape(-1) == cell_type,:].mean(axis=0)) > 1.).reshape(-1)
print('Fraction of genes with avg expression > 1 for cell type {} :'.format(cell_type),
mask_sufficient_expression.mean())
is_zi_pred_genelabel_here = is_zi_pred_genelabel[mask_sufficient_expression,ind_cell_type]
print('Fraction of predicted ZI genes with avg expression > 1 for cell type {} :'.format(cell_type),
is_zi_pred_genelabel_here.mean(), '\n')
| AutoZI_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %load_ext nb_black
# -
import torch.nn.functional as F
import torch
torch.__version__
# ## Does nn.Conv2d init work well?
# [Jump_to lesson 9 video](https://course.fast.ai/videos/?lesson=9&t=21)
# +
#export
import sys
sys.path.append('../')
from exp.nb_02 import *
def get_data():
path = datasets.download_data(MNIST_URL, ext='.gz')
with gzip.open(path, 'rb') as f:
((x_train, y_train), (x_valid, y_valid),
_) = pickle.load(f, encoding='latin-1')
return map(tensor, (x_train, y_train, x_valid, y_valid))
def normalize(x, m, s):
return (x - m) / s
# +
# torch.nn.modules.conv._ConvNd.reset_parameters??
# +
# Signature: torch.nn.modules.conv._ConvNd.reset_parameters(self)
# Docstring: <no docstring>
# Source:
# def reset_parameters(self):
# init.kaiming_uniform_(self.weight, a=math.sqrt(5))
# if self.bias is not None:
# fan_in, _ = init._calculate_fan_in_and_fan_out(self.weight)
# bound = 1 / math.sqrt(fan_in)
# init.uniform_(self.bias, -bound, bound)
# File: ~/anaconda3/envs/dl-zoo/lib/python3.7/site-packages/torch/nn/modules/conv.py
# Type: function
# -
x_train, y_train, x_valid, y_valid = get_data()
train_mean, train_std = x_train.mean(), x_train.std()
x_train = normalize(x_train, train_mean, train_std)
x_valid = normalize(x_valid, train_mean, train_std)
# Reshape to C x h x w to use Conv layers
x_train = x_train.view(-1, 1, 28, 28)
x_valid = x_valid.view(-1, 1, 28, 28)
x_train.shape,x_valid.shape
n, *_ = x_train.shape
c = y_train.max() + 1
nh = 32
n, c
l1 = nn.Conv2d(1, nh, 5)
x = x_valid[:100]
x.shape
def stats(x):
return x.mean(), x.std()
# - weight shape in Conv2D is `out_channels x input_channels x filter_height x filter_width`.
l1.weight.shape
stats(l1.weight), stats(l1.bias)
t = l1(x)
stats(t)
# - Kaiming normal is generally used when the activation function is either **Relu** or **Leaky Reu**.
# - Since all pixels are nonnegative, so leaky relu is the same as **a = 1**.
init.kaiming_normal_(l1.weight, a=1.)
stats(l1(x))
# Looks like it is working.
def f1(x, a=0):
return F.leaky_relu(l1(x), a)
init.kaiming_normal_(l1.weight, a=0)
stats(f1(x))
# The mean is no longer zero because after normalizing the values and using Relu, we removed all values below zero and assign them a zero value. Therefore, the mean will no longer be zero.
l1 = nn.Conv2d(1, nh, 5)
stats(f1(x))
# The variance of default initialization is about **0.36** which is not good at all.
l1.weight.shape
# receptive field size (filter_height x filter_width)
rec_fs = l1.weight[0, 0].numel()
rec_fs
nf, ni= l1.weight.shape[:2]
nf,ni
fan_in = ni * rec_fs
fan_out = nf * rec_fs
fan_in, fan_out
# $$gain = \sqrt\frac{2}{1 + a ^2}$$
def gain(a):
return math.sqrt(2.0 / (1 + a ** 2))
# $a = 0$ is Relu.
gain(0), gain(0.01), gain(0.1), gain(1), gain(math.sqrt(5.))
# The first 3 gains are close to $\sqrt{2}$.
#
# PyTorch uses *kaiming_uniform* for initializing the weight tensors of Conv2D layer. The mean and std of a uniform distribution are $\frac{(b - a)}{2}$ and $\frac{(b - a)^2}{\sqrt{12}}$.
(1 - (-1)) / math.sqrt(12)
# The gain of $\sqrt{5}$ is the same as the std of uniform[-1, 1].
torch.zeros(10000).uniform_(-1, 1).std()
def kaiming2(x, a, use_fan_out=False):
nf, ni, *_ = x.shape
rec_fs = x[0, 0].shape.numel()
fan = nf * rec_fs if use_fan_out else ni * rec_fs
std = gain(a) / math.sqrt(fan)
bound = math.sqrt(3.) * std
x.data.uniform_(-bound, bound)
# since $std = \sqrt{\frac{(b - a)^2}{12}}$ and assume $a = -b$; therefore, $std = \sqrt\frac{4b^2}{12} = \frac{b}{\sqrt{3}}$
# $$\rightarrow bound = std * \sqrt{3}$$
kaiming2(l1.weight, a=0);
stats(f1(x))
kaiming2(l1.weight, a=math.sqrt(5.))
stats(f1(x))
class Flatten(nn.Module):
def forward(self, x):
return x.view(-1)
m = nn.Sequential(
nn.Conv2d(1, 8, 5, stride=2, padding=2), nn.ReLU(),
nn.Conv2d(8, 16, 3, stride=2, padding=1), nn.ReLU(),
nn.Conv2d(16, 32, 3, stride=2, padding=1), nn.ReLU(),
nn.Conv2d(32, 1, 3, stride=2, padding=1),
nn.AdaptiveAvgPool2d(1),
Flatten(),
)
y = y_valid[:100].float()
t = m(x)
stats(t)
# That is bad because the input had a std = 1, the first hidden layer had std = 0.4 and the 4th layer had a std ~ 0. That is not good at all because that means you can't learn anything with 4 layers and you can't create a model with more layers due to this issue.
l = mse(t,y)
l.backward()
stats(m[0].weight.grad)
# The gradients also have close to zero std.
# +
# init.kaiming_uniform_??
# -
for l in m:
if isinstance(l,nn.Conv2d):
init.kaiming_uniform_(l.weight)
l.bias.data.zero_()
t = m(x)
stats(t)
l = mse(t,y)
l.backward()
stats(m[0].weight.grad)
# The above gradients and weights std are much better in terms of std and are no longer close to zero.
# + [markdown] heading_collapsed=true
# ## Export
# + hidden=true
# !python ../src/notebook2script.py 02a_why_sqrt5.ipynb
| dl2/notebooks/02a_why_sqrt5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: elon_tweets
# language: python
# name: elon-tweets
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/jacobpad/Labs-Stuff/blob/master/Elon_Musk_twitter_followers_data.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={} colab_type="code" id="Klieh9osJbR1" jupyter={"outputs_hidden": true}
"""FOR USE IN COLAB"""
# # !pip install squarify
# # !python -m spacy download en_core_web_lg
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="WkdPc-hSJVEx" jupyter={"outputs_hidden": true} outputId="71d544c3-b344-4342-8401-25e14a533977"
# Base
from collections import Counter
import re
import pandas as pd
# Plotting
import squarify
import matplotlib.pyplot as plt
import seaborn as sns
# NLP Libraries
import spacy
from spacy.tokenizer import Tokenizer
from nltk.stem import PorterStemmer
# + colab={"base_uri": "https://localhost:8080/", "height": 419} colab_type="code" id="Zjk15SJe5liM" jupyter={"outputs_hidden": true} outputId="1ebf944f-be93-492f-a56a-342c9f3f6bc3"
import pandas as pd
# Establish the URL
url = 'https://raw.githubusercontent.com/jacobpad/Labs-Stuff/master/elonmusk_followers%20(1).json'
# Read in data
df = pd.read_json(url, orient='index')
df = df.reset_index()
df = df.rename(columns={'index':'date_time', 0:'tweet'})
# View
df
# + colab={} colab_type="code" id="xGhvvLNuCMIu" jupyter={"outputs_hidden": true}
# Apply a first round of text cleaning techniques
import re
import string
'''
# def clean_text_round1(text):
# """
# Make text lowercase, remove text in square brackets, remove punctuation
# and remove words containing numbers.
# """
# text = text.lower()
# text = re.sub('\[.*?\]', '', text)
# text = re.sub('[%s]' % re.escape(string.punctuation), '', text)
# text = re.sub('\w*\d\w*', '', text)
# text = re.sub('\n', '', text)
# return text
# round1 = lambda x: clean_text_round1(x)
# Let's take a look at the updated text
# df['data_clean'] = pd.DataFrame(df['tweet'].apply(round1))
# df
'''
# Tokenizer function
def tokenize(text):
"""
Parses a string into a list of semantic units (words)
Args:
text (str): The string that the function will tokenize.
Returns:
list: tokens parsed out by the mechanics of your choice
"""
# Removing url's
pattern = r"http\S+"
tokens = re.sub(pattern, "", text) # https://www.youtube.com/watch?v=O2onA4r5UaY
tokens = re.sub('[^a-zA-Z 0-9]', '', text)
tokens = tokens.lower().split()
return tokens
# + colab={"base_uri": "https://localhost:8080/", "height": 419} colab_type="code" id="RC1T2ONKJFFj" jupyter={"outputs_hidden": true} outputId="7df06938-aa3e-452c-d576-3ccd92a650e9"
# NLP Libraries
import spacy
from spacy.tokenizer import Tokenizer
from nltk.stem import PorterStemmer
nlp = spacy.load("en_core_web_lg")
# Tokenizer
tokenizer = Tokenizer(nlp.vocab)
# Apply tokenizer
df['tokens'] = df['tweet'].apply(tokenize)
# View
print(df)
# + colab={"base_uri": "https://localhost:8080/", "height": 187} colab_type="code" id="YvbhWeBvMjnQ" jupyter={"outputs_hidden": true} outputId="187da5cd-2bdc-40de-b09a-9c3215a2138d"
# The object `Counter` takes an iterable, but you can instaniate an empty one and update it.
word_counts = Counter()
# Update it based on a split of each of our documents
df['tokens_count'] = df['tokens'].apply(lambda x: word_counts.update(x))
# Print out the 10 most common words
word_counts.most_common(10)
# + colab={} colab_type="code" id="vrBE86cYEYF8" jupyter={"outputs_hidden": true}
# Count Function to count tokens
def count(docs):
word_counts = Counter()
appears_in = Counter()
total_docs = len(docs)
for doc in docs:
word_counts.update(doc)
appears_in.update(set(doc))
temp = zip(word_counts.keys(), word_counts.values())
wc = pd.DataFrame(temp, columns = ['word', 'count'])
wc['rank'] = wc['count'].rank(method='first', ascending=False)
total = wc['count'].sum()
wc['pct_total'] = wc['count'].apply(lambda x: x / total)
wc = wc.sort_values(by='rank')
wc['cul_pct_total'] = wc['pct_total'].cumsum()
t2 = zip(appears_in.keys(), appears_in.values())
ac = pd.DataFrame(t2, columns=['word', 'appears_in'])
wc = ac.merge(wc, on='word')
wc['appears_in_pct'] = wc['appears_in'].apply(lambda x: x / total_docs)
return wc.sort_values(by='rank')
# + colab={"base_uri": "https://localhost:8080/", "height": 419} colab_type="code" id="4jSfn_bmPR3c" jupyter={"outputs_hidden": true} outputId="7b1fbd9d-4aea-4db9-ba11-ec038a0598e7"
word_count_df = count(df['tokens'])
word_count_df
# + colab={"base_uri": "https://localhost:8080/", "height": 279} colab_type="code" id="HyZE-jT1PdzS" jupyter={"outputs_hidden": true} outputId="987a8fa2-7e26-45a7-f3b8-d4c4a3f40264"
# Cumulative Distribution Plot
sns.lineplot(x='rank', y='cul_pct_total', data=word_count_df);
# + colab={"base_uri": "https://localhost:8080/", "height": 248} colab_type="code" id="FpvH63rnPfYQ" jupyter={"outputs_hidden": true} outputId="09d10065-14a1-463f-9b56-264853097c28"
wc_top20 = word_count_df[word_count_df['rank'] <= 20]
squarify.plot(sizes=wc_top20['pct_total'], label=wc_top20['word'], alpha=.8)
plt.axis('off')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 419} colab_type="code" id="kFiovBSGgni6" jupyter={"outputs_hidden": true} outputId="4ed24df9-9c0c-4490-bf7b-eeb4dfd5f08c"
df
# + colab={} colab_type="code" id="y4thjul4gAB2" jupyter={"outputs_hidden": true}
# Customize stop words by adding to the default list
STOP_WORDS = nlp.Defaults.stop_words.union(['hi','\n','\n\n', '&', 'la',' ',
'que', 'de', 'o', 'y', 'en', 'para',
'.', 'el', 'un', 'los', 'le', 'por',
'un', 'el'])
# + colab={"base_uri": "https://localhost:8080/", "height": 669} colab_type="code" id="Mq25d-HpPlkD" jupyter={"outputs_hidden": true} outputId="88f05a69-60c0-4e31-bab7-14b0f0f50c45"
tokens = []
for doc in tokenizer.pipe(df['tweet'], batch_size=500):
doc_tokens = []
for token in doc:
if token.text.lower() not in STOP_WORDS:
doc_tokens.append(token.text.lower())
tokens.append(doc_tokens)
df['tokens'] = tokens
wc = count(df['tokens'])
wc.head(20)
# + colab={"base_uri": "https://localhost:8080/", "height": 248} colab_type="code" id="pjyvBlXTSIEZ" jupyter={"outputs_hidden": true} outputId="f9a8933a-5913-49cf-ea12-c9ce03bb2088"
wc_top20 = wc[wc['rank'] <= 20]
squarify.plot(sizes=wc_top20['pct_total'], label=wc_top20['word'], alpha=.8 )
plt.axis('off')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 771} colab_type="code" id="AztGJDBDpgcy" jupyter={"outputs_hidden": true} outputId="1d019480-9869-4682-8410-d040184e1a67"
wcdf = wc.head(20)
wcdf['top_words'] = wcdf['word'].to_list()
wcdf
# + colab={} colab_type="code" id="JvLGq6P_0blS" jupyter={"outputs_hidden": true}
# + colab={} colab_type="code" id="7xWjDC_i_Jlu" jupyter={"outputs_hidden": true}
# + colab={} colab_type="code" id="HOnQM_XF_Jhy" jupyter={"outputs_hidden": true}
# + colab={} colab_type="code" id="fHKjYL3O_JeV" jupyter={"outputs_hidden": true}
# + colab={} colab_type="code" id="nIXJwNA2_Jak" jupyter={"outputs_hidden": true}
# + colab={} colab_type="code" id="tYVG7Opb_JXT" jupyter={"outputs_hidden": true}
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="ikBVonLGBc0H" jupyter={"outputs_hidden": true} outputId="521ebe9e-2b3a-47e6-bd86-8e688f486642"
# Imports
import requests
import json
# Get the JSON
url = 'https://raw.githubusercontent.com/jacobpad/Labs-Stuff/master/elonmusk_followers_english_only.json'
r = requests.get(url)
df = r.json()
# Simple ormating to work with the JSON
df = pd.DataFrame(df.values())
col = ['original_tweet']
df = df.rename(columns={0:'original_tweet'})
# View it
df
# + colab={} colab_type="code" id="l5dNwq-5G11u" jupyter={"outputs_hidden": true}
# Make emoji free text
# Source: https://stackoverflow.com/questions/33404752/removing-emojis-from-a-string-in-python
import emoji
def give_emoji_free_text(text):
"""
Removes emoji's from tweets
Accepts:
Text (tweets)
Returns:
Text (emoji free tweets)
"""
emoji_list = [c for c in text if c in emoji.UNICODE_EMOJI]
clean_text = ' '.join([str for str in text.split() if not any(i in str for i in emoji_list)])
return clean_text
# Apply the function above and get tweets free of emoji's
call_emoji_free = lambda x: give_emoji_free_text(x)
# Apply `call_emoji_free` which calls the function to remove all emoji's
df['emoji_free_tweet'] = df['original_tweet'].apply(call_emoji_free)
df
# + jupyter={"outputs_hidden": true}
# Removing url's
def remove_url(text):
"""
Remove URL's
Accepts:
emoji_free_tweet
Returns:
emoji_free_tweet & url_free_tweet
Makes a new column
"""
# https://www.youtube.com/watch?v=O2onA4r5UaY
pattern = r"http\S+"
tokens = re.sub(pattern, "", text)
return tokens
# Make new url_free_tweet column by applying the function on emoji_free_tweet
df['url_free_tweet'] = df['emoji_free_tweet'].apply(remove_url)
# View
df
# + jupyter={"outputs_hidden": true}
# Customize stop words by adding to the default list
STOP_WORDS = nlp.Defaults.stop_words.union(['hi','\n','\n\n', '&', ' ', '.', '-', 'got', "it's", 'it’s', "i'm", 'i’m', 'im', 'want'])
tokens = []
for doc in tokenizer.pipe(df['url_free_tweet'], batch_size=500):
doc_tokens = []
for token in doc:
if token.text.lower() not in STOP_WORDS:
doc_tokens.append(token.text.lower())
tokens.append(doc_tokens)
# Makes tokens column
df['tokens'] = tokens
wc = count(df['tokens'])
wc.head(20)
tokens = []
for doc in tokenizer.pipe(df['url_free_tweet'], batch_size=500):
doc_tokens = []
for token in doc:
if token.text.lower() not in STOP_WORDS:
doc_tokens.append(token.text.lower())
tokens.append(doc_tokens)
df['tokens'] = tokens
wc = count(df['tokens'])
wc.head(20)
# + jupyter={"outputs_hidden": true}
df
# + jupyter={"outputs_hidden": true}
# The object `Counter` takes an iterable, but you can instaniate an empty one and update it.
word_counts = Counter()
# Update it based on a split of each of our documents
df['tokens_count'] = df['tokens'].apply(lambda x: word_counts.update(x))
# Print out the 10 most common words
word_counts.most_common(10)
# + jupyter={"outputs_hidden": true}
# Make the word counts a dataframe
wc = pd.DataFrame.from_dict(word_counts, orient='index')
wc = wc.reset_index()
wc = wc.rename(columns={'index':'word', 0:'count'})
wc[:100]
# + jupyter={"outputs_hidden": true}
# Make function to further edit words/tokens
def tokenize_further(text):
df['tokens2'] = df['tokens'].apply(lambda x: x.strip(','))
# + jupyter={"outputs_hidden": true}
wc_word_list = wc['word'].tolist()
wc_word_list
# + jupyter={"outputs_hidden": true}
# + jupyter={"outputs_hidden": true}
# + jupyter={"outputs_hidden": true}
# + jupyter={"outputs_hidden": true}
STOP, DONT RUN BELOW THIS POINT...
# + jupyter={"outputs_hidden": true}
--------
# Count Function to count tokens
def count(docs):
word_counts = Counter()
appears_in = Counter()
total_docs = len(docs)
for doc in docs:
word_counts.update(doc)
appears_in.update(set(doc))
temp = zip(word_counts.keys(), word_counts.values())
wc = pd.DataFrame(temp, columns = ['word', 'count'])
wc['rank'] = wc['count'].rank(method='first', ascending=False)
total = wc['count'].sum()
wc['pct_total'] = wc['count'].apply(lambda x: x / total)
wc = wc.sort_values(by='rank')
wc['cul_pct_total'] = wc['pct_total'].cumsum()
t2 = zip(appears_in.keys(), appears_in.values())
ac = pd.DataFrame(t2, columns=['word', 'appears_in'])
wc = ac.merge(wc, on='word')
wc['appears_in_pct'] = wc['appears_in'].apply(lambda x: x / total_docs)
return wc.sort_values(by='rank')
--------
word_count_df = count(df['tokens'])
word_count_df
--------
# Cumulative Distribution Plot
sns.lineplot(x='rank', y='cul_pct_total', data=word_count_df);
--------
wc_top20 = word_count_df[word_count_df['rank'] <= 20]
squarify.plot(sizes=wc_top20['pct_total'], label=wc_top20['word'], alpha=.8)
plt.axis('off')
plt.show()
--------
--------
--------
| Elon_twitter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 选择
# ## 布尔类型、数值和表达式
# 
# - 注意:比较运算符的相等是两个等到,一个等到代表赋值
# - 在Python中可以用整型0来代表False,其他数字来代表True
# - 后面还会讲到 is 在判断语句中的用发
0 == False
# ## 字符串的比较使用ASCII值
0< 10 <100
mun = eval(input("输入一个分数:"))
if mun > 90:
print("--A")
elif 80< mun <= 90 :
print("---B")
elif 60<=mun<=80:
print("---C")
elif mun< 60:
print("不合格")
# ## Markdown
# - https://github.com/younghz/Markdown
# ## EP:
# - <img src="../Photo/34.png"></img>
# - 输入一个数字,判断其实奇数还是偶数
# ## 产生随机数字
# - 函数random.randint(a,b) 可以用来产生一个a和b之间且包括a和b的随机整数
# +
#产生一个随机数,你去输入,如果你输入的树大于随机数,那么就告诉你太大了,反之,太小了。。。然后你一直输入,直到他满意为止。
import random
a = random.randint(3,10)
while True:
b = eval(input("输入一个数:"))
if a == b:
print("妈的,终于猜对了!")
break
elif a > b:
print("太小了")
elif a < b:
print("太大了")
# -
# ## 其他random方法
# - random.random 返回0.0到1.0之间前闭后开区间的随机浮点
# - random.randrange(a,b) 前闭后开
# ## EP:
# - 产生两个随机整数number1和number2,然后显示给用户,使用户输入数字的和,并判定其是否正确
# - 进阶:写一个随机序号点名程序
mun1,mun2 = (random.randint(1,2),random.randint(1,2))
sum_ = mun1 + mun2
while 1:
p = eval(input(">>"))
if p == sum_:
print("真棒!")
break
else:
print("加油!!!")
# ## if语句
# - 如果条件正确就执行一个单向if语句,亦即当条件为真的时候才执行if内部的语句
# - Python有很多选择语句:
# > - 单向if
# - 双向if-else
# - 嵌套if
# - 多向if-elif-else
#
# - 注意:当语句含有子语句的时候,那么一定至少要有一个缩进,也就是说如果有儿子存在,那么一定要缩进
# - 切记不可tab键和space混用,单用tab 或者 space
# - 当你输出的结果是无论if是否为真时都需要显示时,语句应该与if对齐
# ## EP:
# - 用户输入一个数字,判断其实奇数还是偶数
# - 进阶:可以查看下4.5实例研究猜生日
# ## 双向if-else 语句
# - 如果条件为真,那么走if内部语句,否则走else内部语句
age = eval(input("年龄:"))
if age <=30:
face = (input("长相:"))
if face == "好":
moey = (input("收入"))
if moey == "高":
work = (input("工作"))
if work == "公务员":
print("见面")
else:
print("不见")
else:
print("不高")
else:
print("太丑")
else:
print("too old")
# +
# -
# ## EP:
# - 产生两个随机整数number1和number2,然后显示给用户,使用户输入数字,并判定其是否正确,如果正确打印“you‘re correct”,否则打印正确错误
# ## 嵌套if 和多向if-elif-else
# 
# ## EP:
# - 提示用户输入一个年份,然后显示表示这一年的动物
# 
# - 计算身体质量指数的程序
# - BMI = 以千克为单位的体重除以以米为单位的身高
# 
# +
yera = eval(input("请输入年份:"))
if yera % 12 == 0:
print("猴年")
elif yera % 12 == 1:
print("鸡")
elif yera % 12 == 2:
print("狗")
elif yera % 12 == 3:
print("猪")
elif yera % 12 == 4:
print("鼠")
elif yera % 12 == 5:
print("牛")
elif yera % 12 == 6:
print("虎")
elif yera % 12 == 7:
print("兔")
elif yera % 12 == 8:
print("龙")
elif yera % 12 == 9:
print("蛇")
elif yera % 12 == 10:
print("马")
elif yera % 12 == 11:
print("羊")
else:
print("不知道")
# -
# ## 逻辑运算符
# 
a = [1,2,3]
1 in a #not False
not True
age = 20
(age < 30) #not False
# 
# 
# ## EP:
# - 判定闰年:一个年份如果能被4整除但不能被100整除,或者能被400整除,那么这个年份就是闰年
# - 提示用户输入一个年份,并返回是否是闰年
# - 提示用户输入一个数字,判断其是否为水仙花数
mun = eval(input("请输入年份:"))
if (mun%4 ==0 and mun%100 !=0) or (mun%400 ==0):
print("是闰月年")
else:
print("非闰月年")
mun = eval(input("请输入一个数字:"))
gewei =mun%10
shiwei=(mun//10)%10
baiwei=mun//100
sum1 = baiwei**3+shiwei**3+gewei**3
if sum1==mun:
print("是水仙花数!!")
else:
print("不是水仙花数!!")
# ## 实例研究:彩票
# 
import random
random_1 = random.randint(1,3)
random_2 = random.randint(0,3)
random_ =str(random_1)+str(random_2)
print(random_)
mun = (input("抽奖:"))
if mun ==random_:
print("恭喜您中一等奖")
elif random_[0]==mun[1] and random_[1]==mun[0]:
print("恭喜您中二等奖")
elif random_[0]==mun[0] or random_[0]==mun[1] or random_[1]==mun[0] or random_[1]==mun[1]:
print("恭喜您中三等奖")
else:
print("没有中奖")
# # Homework
# - 1
# 
import math
a,b,c = eval(input("Enter a,b,c:"))
m=math.pow(b,2)-4*a*c
if m>0:
r1 = (-1*b+math.sqrt(b**2-4*a*c))/2*a
r2 = (-1*b+math.sqrt(b**2-4*a*c))/2*a
print("The roots are "+r1,r2)
elif m==0:
r = (-1*b+math.sqrt(b**2-4*a*c))/2*a
print("The root is "+str(r))
else :
print("The equation has no real roots")
# - 2
# 
import random
a = random.randint(0,99)
b = random.randint(0,99)
c = a+b
print(a)
print(b)
print(c)
mun =eval(input("请输入你的答案"))
if mun==c:
print("恭喜您回答正确!")
else:
print("遗憾回答错误!")
# - 3
# 
d = eval(input("Enter today's day:"))
t = eval(input("Enter the munber of days elapsed since today:"))
m = d+t
if d==0:
a ="星期日"
elif d==1:
a="星期一"
elif d==2:
a=("星期二")
elif d==3:
a=("星期三")
elif d==4:
a=("星期四")
elif d==5:
a=("星期五")
elif d==6:
a=("星期六")
if m%7==0:
print( "Today is"+str(a)+"and the future "+str(t)+"is Sunday")
elif m%7==1:
print( "Today is"+str(a)+"and the future "+str(t)+"is Monday")
elif m%7==2:
print( "Today is"+str(a)+"and the future "+str(t)+"is Tuesday")
elif m%7==3:
print( "Today is"+str(a)+"and the future "+str(t)+"is Wednesday")
elif m%7==4:
print( "Today is"+str(a)+"and the future "+str(t)+"is Thursday")
elif m%7==5:
print( "Today is"+str(a)+"and the future "+str(t)+"is Firday")
elif m%7==6:
print( "Today is"+str(a)+"and the future "+str(t)+"is Saturday")
# - 4
# 
a,b,c = eval(input("请输入三个数:"))
d=[a,b,c]
d.sort()
print(d)
# - 5
# 
aw,ap = eval(input("Enter weight and price for package 1:"))
bw,bp = eval(input("Enter weight and price for package 2:"))
a = aw/ap
b = bw/bp
if a>b:
print("Package "+str(1)+" has the better price.")
else:
print("Package "+str(2)+" has the better price.")
# - 6
# 
# +
month = eval(input("请输入月份:"))
year = eval(input("请输入年份:"))
da = [1,3,5,7,10,12]
xiao = [4,6,8,9,11]
if (month in da):
print(year,"年",month,"月有31天")
elif (month in xiao):
print(year,"年",month,"月有30天")
else:
if (year%4==0 and year%100!=0) or (year%400==0):
print(year,"年为闰年",month,"月有29天")
else:
print(year,"年为平年",month,"月有28天")
# -
# - 7
# 
mun = eval(input("请输入一个猜测值(1代表正面,0代表反面):"))
a = random.randint(0,1)
if mun==a:
print("恭喜您猜对了!")
else:
print("遗憾您猜错了!")
"""dfsdfdf"""
# - 8
# 
mun = eval(input("scissor(0),rock(1),paper(2):"))
a = random.randint(0,2)
print(a)
if a==mun:
if a==mun==0:
print("he computer is scissor. You are scissor too.Tt is a draw")
elif a==mun==1:
print("he computer is rock. You are rock too.Tt is a draw")
else:
print("The computer is paper. You are paper too.Tt is a draw.")
elif a>mun:
if a==1 and mun==0:
print("The compuer is rock.You are scissor.You lose")
elif a==2 and mun==0:
print("The compuer is paper.You are scissor.You won")
else:
print("The compuer is paper.You are rock.You lose")
elif a<mun:
if a==0 and mun ==1:
print("The compuer is scissor.You are rock.You won")
elif a==0 and mun==2:
print("The compuer is scissor.You are paper.You lose")
elif a==1 and mun==2:
print("The compuer is rock.You are paper.You won")
# - 9
# 
# - 10
# 
# - 11
# 
mun = (input("请输入一个三位数:"))
if mun[0]==mun[2]:
print(str(mun)+"这个数是回文")
else:
print("这个数不是回文")
# - 12
# 
# +
a,b,c = eval(input("Enter three edges:"))
if (a+b>c)and(a+c>b)and(b+c>a):
l=a+b+c
print("它的周长是"+str(l))
else:
print("这个输入是非法的")
| 9.12.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sethtroisi/OEIS/blob/master/A037274/ProbPrimePrefix.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="_3fVAU_ZjHxK" colab_type="code" cellView="form" colab={}
#@title Look at probability of prime of form \<prefix>\<x digits>\[1379\]
# + id="U2WCLmkMcXmW" colab_type="code" colab={}
from IPython.display import clear_output
# !apt install libgmp-dev libmpfr-dev libmpc-dev
# !pip install gmpy2
clear_output()
# + id="DW0ZChHAa8WO" colab_type="code" cellView="form" colab={}
#@title Parameters { run: "auto" }
SIMULATIONS = 100000 #@param {type:"integer"}
num_digits = 80 #@param {type:"integer"}
# + id="1jtZCn_7anS3" colab_type="code" outputId="4838c3e9-edd6-4e0f-ba14-d216171f41b1" colab={"base_uri": "https://localhost:8080/", "height": 593}
import random
import gmpy2
import numpy as np
import math
import matplotlib.pyplot as plt
import seaborn as sns
import time
import tqdm
sns.set()
rand = random.Random(49)
base_rate = 1 / math.log(10 ** (num_digits + 0.5))
base_rate *= 2/1 * 5/4 # We filter these
prefixes = list(range(1, 99+1))
counts = [base_rate] + [0 for p in prefixes]
for simulation in range(SIMULATIONS+1):
middle = random.randint(10 ** (num_digits-4), 10 ** (num_digits-3) - 1)
for p in prefixes:
end = [1,3,7,9][simulation % 4]
test = str(p) + str(middle) + str(end)
assert len(test) == num_digits - 1 * (p <= 9)
if gmpy2.is_prime(int(test)):
counts[p] += 1
if simulation and simulation % 1000 == 0:
ratio = [counts[i] / simulation for i in range(len(counts))]
ratio[0] = base_rate
clear_output()
print ("After", simulation)
_ = plt.figure(figsize=(12, 8))
ax = sns.heatmap(np.reshape(ratio, (10, 10)), annot=True, fmt=".3f", cbar=False)
plt.show()
print ()
print ("Prime Number Theorum: {:.3}".format(ratio[0]))
print ("Avg: {:.3}".format(sum(ratio)/len(ratio)))
print ("Min: {:.3}".format(min(ratio)))
print ("Max: {:.3}".format(max(ratio)))
| A037274/ProbPrimePrefix.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="XJcXzC1IYiDX"
# # ECE 3 : Lab 4
# + [markdown] id="7EWvs0qtA9iq"
# ## Warm-Up
# + [markdown] id="6QRH_iyBBufA"
# #### (a) For matrix A use the np.sum() function to print:
#
# 1. The sum of all its elements
# 2. The sum of the elements for each one of its columns
# 3. The sum of the elements for each one of its rows
# + id="hoIlpx_pBqdy"
# Do not forget to import numpy
A = np.array([[1,2,3], [4, 2, 1], [6, 7, -5]])
# Insert your code below
# + [markdown] id="qRiihe-fCX7f"
# #### (b) Use the np.bincount() function to print the number of occurences of each value in vector x:
#
# + id="2ViwSFXHCfDk"
x = np.array([0, 1, 1, 1, 0, 1])
# Insert your code below
# + [markdown] id="jbyukgIcC4kG"
# #### (c) Use the np.argsort() function to print the indices that would sort vector x:
#
# 1. In ascending order
# 2. In descending order
# + id="KL-q_T3nDITU"
x = np.array([1, 6, 2, -1, 0])
# Insert your code below
# + [markdown] id="0ZhKyQlOX8sK"
# #### (d) Create a function `EvenList(n)` which generates a vector of even numbers from 2 to n (included). The function must return this vector.
#
# + id="JRrq3-zNU9H3" colab={"base_uri": "https://localhost:8080/", "height": 130} executionInfo={"status": "error", "timestamp": 1634671404412, "user_tz": 420, "elapsed": 14, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "00734435821983708962"}} outputId="1e883877-6b6e-4eef-ebf3-9ffed0dc4e57"
#Insert your code below
def EvenList(n):
return
# calling the function
print(EvenList(5)) #n=5
# + [markdown] id="iP1qpcO8ZMiw"
# #### (e) Write a function `BoundaryList(A)` which return the maximum and minimum value of the array `A`. The function should return a tuple `(min_val, max_val)`.
# + id="nd36IFkaZLdS"
# Insert your code below
def BoundaryList(A):
return
# calling the function
print(BoundaryList(np.array([-5,2,0,8,7])))
# + [markdown] id="SJrDeVZrbd90"
# #### (f) Generate the following pattern :
#
# \* * * * *
# \* * * *
# \* * *
# \* *
# \*
#
# *Hint : You will definitely need for loop(s).*
# + id="eJ0kU9Vuax_S"
# Insert your code below
# + [markdown] id="HxD67_WgcqoA"
# # ++ As an exercise, try coding a function which takes `n` as input and prints `n` rows of such a pattern. For example:
#
# GeneratePattern(3) will print:
#
# \* * *
# \* *
# \*
#
#
#
# GeneratePattern(4) will print:
#
# \* * * *
# \* * *
# \* *
# \*
#
# + id="Pl6LpceQcpUA"
# Insert your code below
def GeneratePattern(n):
return
# calling the function
GeneratePattern(3)
# + [markdown] id="a3w7PWGCYl6Y"
# ## Problem 1 - Classification using the k-NN algorithm (synthetic dataset)
# + [markdown] id="LYd766Ae7FrS"
# ### Some introductory ML terminology
#
# In engineering when we study a phenomenon we want to collect data about it and form a **dataset**. We do that by recording some measurable properties of the phenomenon, which we call **features**. The generated dataset consists of a number of **datapoints**, which are the measurements of said features. In some experiments we wish to categorize our data into classes, for that each datapoint is also accompanied by a **label** which indicates to which class it belongs.
# + [markdown] id="T_Q-GrM5Kr_T"
# ### Lemons vs limes
#
# Consider a farmers' co-op that receives mixed batches of lemons and limes from local producers and wants to separate them before distributing them to grocery stores. For that they install a camera system over their conveyor belt, which uses some advanced image processing algorithms and measures the volume and the hue of each fruit.
#
# They now need an algorithm that will classify the fruit as lemons or limes based on these two features. An engineer proposes that they use a machine learning solution. That means that instead of being explicitly programmed, the algorithm will "learn" a decision rule from a set of initial observations. For this, they measure the volume and the hue of 10 lemons and 10 limes using their camera system and assign the label "0" to lemons and the label "1" to limes.
#
# Because we will use this initial set of observations to train our algorithm, we call it the **training set**. The following block of code simulates the generation of the training set.
# + id="hrIMj1KaYlmW"
import numpy as np
# for now you don't need to understand how data are generated in this block
np.random.seed(0)
X_train = np.vstack((np.random.multivariate_normal([1, 5], [[1, 0], [0, 1]], size=10),
np.random.multivariate_normal([3, 3], [[1, 0], [0, 1]], size=10)))
y_train = np.hstack((np.zeros((10,), dtype=np.int8), np.ones((10,), dtype=np.int8))).T
# + [markdown] id="rcN3pll5tALd"
# Our training set contains 20 datapoints, each consisting of two feature measurements and a label. Let's take a look at a datapoint:
# + id="RZlyVzIeuT1r"
print("The training set containts {} datapoints. Each datapoint has {} features and belongs to one of {} classes.".format(X_train.shape[0], X_train.shape[1], len(np.unique(y_train))))
print("The first datapoint has hue {}, volume {} and belongs to class {}.".format(X_train[0, 0], X_train[0, 1], y_train[0]))
# + [markdown] id="lDA6zDdrygMp"
# Let's make a scatter plot of our training set:
# + id="n-5K8JSy2iFs"
from matplotlib import pyplot as plt
from matplotlib import colors
def scatter_plot(data, labels, class_names, class_colors, marker_type):
scatter = plt.scatter(data[:, 0], data[:, 1], c=labels, cmap=class_colors, marker=marker_type)
plt.legend(*scatter.legend_elements())
plt.legend(handles=scatter.legend_elements()[0], labels=class_names)
plt.xlabel('Hue')
plt.ylabel('Volume')
plt.grid()
names = ['Lemons', 'Limes']
marker_type = "o"
colormap = colors.ListedColormap(['gold', 'limegreen'])
scatter_plot(X_train, y_train, names, colormap, marker_type)
# + [markdown] id="U8rjZfHkD2Em"
# ### k Nearest Neighbors (k-NN) classification rule
#
# Now suppose that we have a set of 5 new datapoints that we don't know to which class they belong. We call that the test set:
# + id="YoUG9Xin5Md5"
X_test = np.array([[1.5, 3.0], [2.0, 6.0], [3.5, 4.5], [4.0, 3.0], [2.5, 4.0]])
y_test = np.array([2, 2, 2, 2, 2]).T # label 2 means uknown, it's not a new class
names = ['Lemons', 'Limes', 'Uknowns']
marker_type="o"
colormap = colors.ListedColormap(['gold', 'limegreen', 'red'])
scatter_plot(np.vstack((X_train, X_test)), np.hstack((y_train, y_test)),
names, colormap, marker_type)
# + [markdown] id="IGIOCswk2202"
# How do we decide if each of these new points is a lemon or a lime? How can we exploit the information our initial observations give us to make this decision?
#
# An answer to the above, is the k Nearest Neighbors algorithm. The k-NN classification rule assigns a datapoint to the class most common among its k nearest neighbors.
#
# * "Nearest" is defined with respect to some distance metric, usually that is the euclidean distance, however other distances are more appropriate for some applications
# * k is a parameter chosen by the engineer, it is taken to be odd to avoid a situation where we can't decide because of a tie
#
# In this exercise you will implement the k-NN rule, using the euclidean distance, to assign each datapoint from the test set to a class.
# + [markdown] id="YcI4hchM58W1"
# #### (a) Write a function that calculates the euclidean distance between two points:
# + id="NTofhpAW57Kr"
# Replace ... with your code
def eucl_dist(a, b):
return np.sum((...) ** ...) ** (...)
# + [markdown] id="EiAl8GZU_MkX"
# #### (b) Write a function that receives as arguments a point and the training set, and calculates the distance of that point from all the points of the training set. The function should return a vector of length equal to the number of points in the training set. You can opt to you use the previous function that you wrote or not.
#
#
#
# + id="SuSKmWn4-sRh"
# Replace ... with your code
def dist_from_training_set(a, X):
return np.sum((...) ** ..., axis=...) ** (...)
# + [markdown] id="NiOcye9WCNdD"
# #### (c) Write a function that receives as arguments a point, the training data, the training labels and an odd positive number k and implements that k-NN rule to assign the point into a class. The function should return the label assign to the point.
# + id="DOoIrSA7Buhb"
# Replace ... with your code
def kNN(a, X, y, k):
distances = dist_from_training_set(...)
nn_indices = distances.argsort()[:k]
nn_labels = y[...]
nn_occurences = np.bincount(...)
return np.argmax(...)
# + [markdown] id="T9uVr4fDB4Xq"
# #### (d) Using the 3-NN rule assign a label to each point of the test set. Store the results in a NumPy vector.
# + id="we0e3fETAaTT"
# Replace ... with your code
n_test = X_test.shape[0]
y_pred_3NN = np.empty(n_test,).T
for idx in range(n_test):
y_pred_3NN[idx] = kNN(...)
# + [markdown] id="Vq-uA3YcXisA"
# #### (e) Use sklearn's KNeighborsClassifier class to verify your result.
#
#
# + id="uMkFJYDjYk8j"
# Replace ... with your code
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors=...)
knn.fit(...)
y_pred = knn.predict(...)
print("User prediction : ", y_pred_3NN)
print("Package prediction : ", y_pred)
# + [markdown] id="FNgJ7QvnAaAM"
# #### (f) Repeat (d) but now use the 1-NN and the 5-NN rules. You may use yours or sklearn's implementation of the kNN algorithm. What do you observe?
# + id="KnGsLnACJOon"
# Replace ... with your code
y_pred_1NN, y_pred_5NN = np.empty(n_test,).T, np.empty(n_test,).T
for idx in range(...):
y_pred_1NN[...] = kNN(...)
y_pred_5NN[...] = kNN(...)
print(y_pred_1NN, y_pred_3NN, y_pred_5NN)
# + [markdown] id="7NwhlSWRM4TE"
# ### Plotting classification results
# + id="i4QXK78xM3JQ"
names_pred = ['Pred : 3NN']
colormap = colors.ListedColormap(['gold', 'limegreen'])
marker_test = "^"
scatter_plot(X_test, y_pred_3NN, None, colormap, marker_test)
names = ['Lemons', 'Limes']
colormap = colors.ListedColormap(['gold', 'limegreen'])
marker_train = "o"
scatter_plot(X_train, y_train, names, colormap, marker_train)
plt.grid()
# + [markdown] id="T7Y64HZZcRnC"
# ### Accuracy - A classificiation performance metric
#
# Now suppose that we know the ground truth labels for the examples in the test set. A metric for the performance of the classification algorithm is its accuracy which is defined as:
#
# $$accuracy = \frac{\text{number of correct predictions}}{\text{total number of predictions}}$$
# + [markdown] id="NQq-EKSicOW8"
# #### (g) Write a function that takes as arguments the ground truth labels and the predicted labels and outputs the classification accuracy. Use it to calculate the accuracy of the 3NN and 5NN rules on the test set.
# + id="mjep_bencN9n"
# Replace ... with your code
y_test = np.array([1, 0, 0, 1, 0]).T # this is the ground truth
def accuracy(y_true, y_pred):
n = y_true.shape[0]
return np.sum(...) / ...
acc_3NN = accuracy(...)
acc_5NN = accuracy(...)
print(acc_3NN, acc_5NN)
| lab-notebook/lab4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Result
#
# [](https://mybinder.org/v2/gh/danhtaihoang/hidden-variables/master?filepath=sphinx%2Fcodesource%2Fhidden.ipynb)
#
# We will demontrade in the following the performance of our method in analyzing binary data that is generated from the kinetic Ising model. Using configurations of a subset of variables, we will recover the interactions (including observed to observed, hidden to observed, observed to hidden, and hidden to hidden), the configurations of hidden variables, and the number of hidden variables.
#
# First of all, we import the necessary packages to the jupyter notebook:
# +
import numpy as np
import sys
import matplotlib.pyplot as plt
import simulate
import inference
# %matplotlib inline
np.random.seed(1)
# -
# We consider a system of `n0` variables interacting each others with interaction variability parameter `g`.
# +
# parameter setting:
n0 = 40 # number of variables
g = 4.0 # interaction variability parameter
w0 = np.random.normal(0.0,g/np.sqrt(n0),size=(n0,n0))
# -
# Using the function `simulate.generate_data`, we then generate a time series of variable states according to the kinetic Ising model with a data length `l`.
# +
l = int(2*(n0**2))
s0 = simulate.generate_data(w0,l)
# -
# A raster of variable configurations `s0` is plotted:
plt.figure(figsize=(8,6))
plt.title('raster of time series')
plt.imshow(s0.T[:,:400],cmap='gray',origin='lower')
plt.xlabel('time')
plt.ylabel('variable index')
plt.show()
# Using the configurations of entire system `s0`, we can infer sucessfully the interactions between variables:
w = inference.fem(s0)
# We plot the heat map of the actual interaction matrix `w0`, the inferred interaction matrix `w`, and the error inference, i.e., the discrepancy between `w0` and `w`.
# +
plt.figure(figsize=(14,3.2))
plt.subplot2grid((1,4),(0,0))
plt.title('actual interaction matrix')
plt.imshow(w0,cmap='rainbow',origin='lower')
plt.xlabel('j')
plt.ylabel('i')
plt.clim(-0.5,0.5)
plt.colorbar(fraction=0.045, pad=0.05,ticks=[-0.5,0,0.5])
plt.subplot2grid((1,4),(0,1))
plt.title('inferred interaction matrix')
plt.imshow(w,cmap='rainbow',origin='lower')
plt.xlabel('j')
plt.ylabel('i')
plt.clim(-0.5,0.5)
plt.colorbar(fraction=0.045, pad=0.05,ticks=[-0.5,0,0.5])
plt.subplot2grid((1,4),(0,2))
plt.title('error')
plt.imshow(w0-w,cmap='rainbow',origin='lower')
plt.xlabel('j')
plt.ylabel('i')
plt.clim(-0.5,0.5)
plt.colorbar(fraction=0.045, pad=0.05,ticks=[-0.5,0,0.5])
plt.subplot2grid((1,4),(0,3))
plt.title('inferred vs. actual')
plt.plot([-2.5,2.5],[-2.5,2.5],'r--')
plt.scatter(w0,w)
plt.xticks([-2,0,2])
plt.yticks([-2,0,2])
plt.xlabel('actual interactions')
plt.ylabel('inferred interactions')
plt.tight_layout(h_pad=1, w_pad=1.5)
plt.show()
# -
# However, in the real world data, it is likely that we do not observe the configurations of entire system, but only a subset of variables. For instance, there exists `nh0` hidden variables. So that, the number of observed variables is `n = n0 - nh0` and the observed configurations is `s = s0[:,:n]`.
nh0 = 10
n = n0 - nh0
s = s0[:,:n].copy()
# ## Ignoring hidden variables
# Let us see what happen if we ignore the existence of hidden variables and infer the interactions between the observations. Using the observed configurations `s`, we can infer the interactions between the observed variables:
w = inference.fem(s)
# The heat map of the actual interaction matrix `w0`, the inferred interaction matrix `w` (only from observed to observed), and the error between them are plotted.
# +
w_infer = np.zeros((n0,n0))
w_infer[:n,:n] = w
error = np.zeros((n0,n0))
error[:n,:n] = w0[:n,:n] - w
#-------------------------------------
plt.figure(figsize=(14,3.2))
plt.subplot2grid((1,4),(0,0))
plt.title('actual interaction matrix')
plt.imshow(w0,cmap='rainbow',origin='lower')
plt.xlabel('j')
plt.ylabel('i')
plt.clim(-0.5,0.5)
plt.colorbar(fraction=0.045, pad=0.05,ticks=[-0.5,0,0.5])
plt.subplot2grid((1,4),(0,1))
plt.title('inferred interaction matrix')
plt.imshow(w_infer,cmap='rainbow',origin='lower')
plt.xlabel('j')
plt.ylabel('i')
plt.clim(-0.5,0.5)
plt.colorbar(fraction=0.045, pad=0.05,ticks=[-0.5,0,0.5])
plt.subplot2grid((1,4),(0,2))
plt.title('error')
plt.imshow(error,cmap='rainbow',origin='lower')
plt.xlabel('j')
plt.ylabel('i')
plt.clim(-0.5,0.5)
plt.colorbar(fraction=0.045, pad=0.05,ticks=[-0.5,0,0.5])
plt.subplot2grid((1,4),(0,3))
plt.title('observed to observed')
plt.plot([-2.5,2.5],[-2.5,2.5],'r--')
plt.scatter(w0[:n,:n],w_infer[:n,:n])
plt.xticks([-2,0,2])
plt.yticks([-2,0,2])
plt.xlabel('actual interactions')
plt.ylabel('inferred interactions')
plt.tight_layout(h_pad=1, w_pad=1.5)
plt.show()
# -
# As expected, ignoring hidden variables leads to a poor inferred result.
#
# In the following, we will take into account the existance of hidden variables.
# ## Including hidden variables
# ### Inferring interactions
#
# We first assume that the number of hidden variables is known, `nh = nh0`. The interactions is then inferred by using the function `inference.infer_hidden`.
# +
nh = nh0
nrepeat = 100
cost_obs,w,sh = inference.infer_hidden(s,nh,nrepeat)
# -
# To compare the inferred interactions with the actual interactions, we use the function `inference.hidden_coordinate` to identify which inferred hidden variable corresponding to which actual hidden variable.
w,sh = inference.hidden_coordinate(w0,s0,w,sh)
# Now, we plot the the heat map of the actual interaction matrix `w0`, the inferred interaction matrix `w` (including observed to observed, hidden to observed, observed to hidden, and hidden to hidden), and the error between them.
# +
plt.figure(figsize=(11,3.2))
plt.subplot2grid((1,3),(0,0))
plt.title('actual interaction matrix')
plt.imshow(w0,cmap='rainbow',origin='lower')
plt.xlabel('j')
plt.ylabel('i')
plt.clim(-0.5,0.5)
plt.colorbar(fraction=0.045, pad=0.05,ticks=[-0.5,0,0.5])
plt.subplot2grid((1,3),(0,1))
plt.title('inferred interaction matrix')
plt.imshow(w,cmap='rainbow',origin='lower')
plt.xlabel('j')
plt.ylabel('i')
plt.clim(-0.5,0.5)
plt.colorbar(fraction=0.045, pad=0.05,ticks=[-0.5,0,0.5])
plt.subplot2grid((1,3),(0,2))
plt.title('error')
plt.imshow(w0-w,cmap='rainbow',origin='lower')
plt.xlabel('j')
plt.ylabel('i')
plt.clim(-0.5,0.5)
plt.colorbar(fraction=0.045, pad=0.05,ticks=[-0.5,0,0.5])
plt.tight_layout(h_pad=1, w_pad=1.5)
plt.show()
# +
plt.figure(figsize=(13.2,3.2))
plt.subplot2grid((1,4),(0,0))
plt.title('observed to observed')
plt.plot([-2.5,2.5],[-2.5,2.5],'r--')
plt.scatter(w0[:n,:n],w[:n,:n])
plt.xticks([-2,0,2])
plt.yticks([-2,0,2])
plt.xlabel('actual interactions')
plt.ylabel('inferred interactions')
plt.subplot2grid((1,4),(0,1))
plt.title('hidden to observed')
plt.plot([-2.5,2.5],[-2.5,2.5],'r--')
plt.scatter(w0[:n,n:],w[:n,n:])
plt.xticks([-2,0,2])
plt.yticks([-2,0,2])
plt.xlabel('actual interactions')
plt.ylabel('inferred interactions')
plt.subplot2grid((1,4),(0,2))
plt.title('observed to hidden')
plt.plot([-2.5,2.5],[-2.5,2.5],'r--')
plt.xticks([-2,0,2])
plt.yticks([-2,0,2])
plt.scatter(w0[n:,:n],w[n:,:n])
plt.xlabel('actual interactions')
plt.ylabel('inferred interactions')
plt.subplot2grid((1,4),(0,3))
plt.title('hidden to hidden')
plt.plot([-2.5,2.5],[-2.5,2.5],'r--')
plt.scatter(w0[n:,n:],w[n:,n:])
plt.xticks([-2,0,2])
plt.yticks([-2,0,2])
plt.xlabel('actual interactions')
plt.ylabel('inferred interactions')
plt.tight_layout(h_pad=1, w_pad=1.5)
plt.show()
# -
# So we can infer accurately the interactions, not only observed to observed, but also hidden to observed, observed to hidden, and hidden to hidden.
# ### Inferring configurations of hidden variables
# Our algorithm allows us to infer the configurations of hidden variables simultaneously with the interactions in the previous step. The acurracy of the inference is calculated as
accuracy = 1 - (np.abs(s0[:,n:]-sh)/2.).mean()
print('inference accuracy: %2.2f %%'%(accuracy*100))
# ### Inferring number of hidden variables
#
# In the above sections, we recovered the interactions and configurations of hidden variables by using the right number of hidden variables, `nh= nh0`. However, in the real world problems, the number of hidden variables may not be accessible. In the following, we apply our method for various number of hidden variables `nh` and compute the discrepancies of observed variables `cost_obs` and entire system `cost`.
# +
nh_list = [6,8,10,12,14,16]
nnh = len(nh_list)
def discrepancy(nh):
return inference.infer_hidden(s,nh,nrepeat)[0]
import multiprocessing
nPC = min(nnh,multiprocessing.cpu_count())
pool = multiprocessing.Pool(processes=nPC)
cost_obs_list = pool.map(discrepancy,nh_list)
pool.close()
cost_obs = np.array([cost_obs_list[i][-1] for i in range(nnh)])
cost = np.array([cost_obs[i]*(1+float(nh_list[i])/n) for i in range(nnh)])
# -
# We plot the discrepancies of observed variables `cost_obs` and entire system `cost` as function of hidden variables. The minimum of `cost` corresponds to the true value of number of hidden variables.
# +
plt.figure(figsize=(4,3.2))
plt.plot(nh_list,cost_obs,'ko-',linewidth=1,markersize=4,label='observed')
plt.plot(nh_list,cost,'r^-',linewidth=1,markersize=4,label='entire')
plt.xticks(nh_list)
plt.xlabel('number of hidden variables')
plt.ylabel('Discepancy')
plt.legend()
plt.show()
# -
| sphinx/ref_code/hidden_v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Implementing Back Propagation
#
# For this recipe, we will show how to do TWO separate examples, a regression example, and a classification example.
#
# To illustrate how to do back propagation with TensorFlow, we start by loading the necessary libraries and resetting the computational graph.
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
from tensorflow.python.framework import ops
ops.reset_default_graph()
# ### Create a Graph Session
sess = tf.Session()
# ## A Regression Example
#
# ------------------------------
#
# We create a regression example as follows. The input data will be 100 random samples from a normal (mean of 1.0, stdev of 0.1). The target will be 100 constant values of 10.0.
#
# We will fit the regression model: `x_data * A = target_values`
#
# Theoretically, we know that A should be equal to 10.0.
#
# We start by creating the data and targets with their respective placholders
x_vals = np.random.normal(1, 0.1, 100)
y_vals = np.repeat(10., 100)
x_data = tf.placeholder(shape=[1], dtype=tf.float32)
y_target = tf.placeholder(shape=[1], dtype=tf.float32)
# We now create the variable for our computational graph, `A`.
# Create variable (one model parameter = A)
A = tf.Variable(tf.random_normal(shape=[1]))
# We add the model operation to the graph. This is just multiplying the input data by A to get the output.
# Add operation to graph
my_output = tf.multiply(x_data, A)
# Next we have to specify the loss function. This will allow TensorFlow to know how to change the model variables. We will use the L2 loss function here. Note: to use the L1 loss function, change `tf.square()` to `tf.abs()`.
# Add L2 loss operation to graph
loss = tf.square(my_output - y_target)
# Now we initialize all our variables. For specificity here, this is initializing the variable `A` on our graph with a random standard normal number.
# Initialize variables
init = tf.global_variables_initializer()
sess.run(init)
# We need to create an optimizing operations. Here we use the standard `GradientDescentOptimizer()`, and tell TensorFlow to minimize the loss. Here we use a learning rate of `0.02`, but feel free to experiment around with this rate, and see the learning curve at the end. However, note that learning rates that are too large will result in the algorithm not converging.
# Create Optimizer
my_opt = tf.train.GradientDescentOptimizer(0.02)
train_step = my_opt.minimize(loss)
# ### Running the Regression Graph!
#
# Here we will run the regression computational graph for 100 iterations, printing out the A-value and loss every 25 iterations. We should see the value of A get closer and closer to the true value of 10, as the loss goes down.
# Run Loop
for i in range(100):
rand_index = np.random.choice(100)
rand_x = [x_vals[rand_index]]
rand_y = [y_vals[rand_index]]
sess.run(train_step, feed_dict={x_data: rand_x, y_target: rand_y})
if (i+1)%25==0:
print('Step #' + str(i+1) + ' A = ' + str(sess.run(A)))
print('Loss = ' + str(sess.run(loss, feed_dict={x_data: rand_x, y_target: rand_y})))
# ## Classification Example
#
# --------------------
#
# For the classification example, we will create an x-sample made of two different normal distribution inputs, `Normal(mean = -1, sd = 1)` and `Normal(mean = 3, sd = 1)`. For each of these the target will be the class `0` or `1` respectively.
#
# The model will fit the binary classification: If `sigmoid(x+A) < 0.5` then predict class `0`, else class `1`.
#
# Theoretically, we know that `A` should take on the value of the negative average of the two means: `-(mean1 + mean2)/2`.
#
# We start by resetting the computational graph:
ops.reset_default_graph()
# Start a graph session
# Create graph
sess = tf.Session()
# We generate the data that we will feed into the graph. Note that the `x_vals` are the combination of two separate normals, and the y_vals are the combination of two separate constants (two classes).
#
# We also create the relevant placeholders for the model.
# Create data
x_vals = np.concatenate((np.random.normal(-1, 1, 50), np.random.normal(3, 1, 50)))
y_vals = np.concatenate((np.repeat(0., 50), np.repeat(1., 50)))
x_data = tf.placeholder(shape=[1], dtype=tf.float32)
y_target = tf.placeholder(shape=[1], dtype=tf.float32)
# We now create the one model variable, used for classification. We also set the initialization function, a random normal, to have a mean far from the expected theoretical value.
#
# - Initialized to be around 10.0
# - Theoretically around -1.0
# Create variable (one model parameter = A)
A = tf.Variable(tf.random_normal(mean=10, shape=[1]))
# Now we add the model operation to the graph. This will be the adding of the variable `A` to the data. Note that the `sigmoid()` is left out of this operation, because we will use a loss function that has it built in.
#
# We also have to add the batch dimension to each of the target and input values to use the built in functions.
# +
# Add operation to graph
# Want to create the operstion sigmoid(x + A)
# Note, the sigmoid() part is in the loss function
my_output = tf.add(x_data, A)
# Now we have to add another dimension to each (batch size of 1)
my_output_expanded = tf.expand_dims(my_output, 0)
y_target_expanded = tf.expand_dims(y_target, 0)
# -
# Add classification loss (cross entropy)
xentropy = tf.nn.sigmoid_cross_entropy_with_logits(logits=my_output_expanded, labels=y_target_expanded)
# Now we declare the optimizer function. Here we will be using the standard gradient descent operator with a learning rate of `0.05`.
# Create Optimizer
my_opt = tf.train.GradientDescentOptimizer(0.05)
train_step = my_opt.minimize(xentropy)
# Next we create an operation to initialize the variables and then run that operation
# Initialize variables
init = tf.global_variables_initializer()
sess.run(init)
# ### Running the Classification Graph!
#
# Now we can loop through our classification graph and print the values of A and the loss values.
# Run loop
for i in range(1400):
rand_index = np.random.choice(100)
rand_x = [x_vals[rand_index]]
rand_y = [y_vals[rand_index]]
sess.run(train_step, feed_dict={x_data: rand_x, y_target: rand_y})
if (i+1)%200==0:
print('Step #' + str(i+1) + ' A = ' + str(sess.run(A)))
print('Loss = ' + str(sess.run(xentropy, feed_dict={x_data: rand_x, y_target: rand_y})))
# Now we can also see how well we did at predicting the data by creating an accuracy function and evaluating them on the known targets.
# +
# Evaluate Predictions
predictions = []
for i in range(len(x_vals)):
x_val = [x_vals[i]]
prediction = sess.run(tf.round(tf.sigmoid(my_output)), feed_dict={x_data: x_val})
predictions.append(prediction[0])
accuracy = sum(x==y for x,y in zip(predictions, y_vals))/100.
print('Ending Accuracy = ' + str(np.round(accuracy, 2)))
# -
| 02_TensorFlow_Way/05_Implementing_Back_Propagation/05_back_propagation.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python3
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from funcionario import euler_npts
# # Uma EDO "instável"
#
# Considere a EDO $y' = -10y$.
#
# ## Qual é a solução desta EDO?
#
# ## Observando a solução numérica
#
# Vamos ver como o método de Euler se comporta para esta equação:
# +
def G(t,y):
return -10*y
def ansG(t,t0,y0): return np.exp(-10*(t-t0))*y0
# +
_, [ax1, ax2, ax3] = plt.subplots(ncols=3, figsize=(16,4))
ns = np.logspace(5,10,num=6,base=2,dtype=int)
for n in ns:
ts, ys = euler_npts(G, [0,5], y0=1.2, npts=n, retpts=True)
ax1.plot(ts, ys, label=str(n))
ax2.semilogy(ts, np.abs(ys - ansG(ts,0,1)), label=str(n))
ax3.semilogy(ts, np.abs(1 - ys/ansG(ts,0,1)), label=str(n))
for a in [ax1, ax2, ax3]:
a.set_xlabel('t')
a.legend(title='nstep')
ax1.set_title('Euler explícito: solução')
ax2.set_title('Euler explícito: erro absoluto')
ax3.set_title('Euler explícito: erro relativo')
plt.show()
# -
# ## Observando mais de perto
#
# Vamos dar um "zoom" na "transição de fase" para ver o que aconteceu entre 32 e 64 pontos:
# +
_, [ax1, ax2, ax3] = plt.subplots(ncols=3, figsize=(16,4))
# Copiar, mudar ns
### Resposta aqui
for n in ns:
ts, ys = euler_npts(G, [0,5], y0=1.2, npts=n, retpts=True)
ax1.plot(ts, ys, label=str(n))
ax2.semilogy(ts, np.abs(ys - ansG(ts,0,1)), label=str(n))
ax3.semilogy(ts, np.abs(1 - ys/ansG(ts,0,1)), label=str(n))
for a in [ax1, ax2, ax3]:
a.set_xlabel('t')
a.legend(title='nstep')
ax1.set_title('Euler explícito: solução')
ax2.set_title('Euler explícito: erro absoluto')
ax3.set_title('Euler explícito: erro relativo')
plt.show()
# -
# # Analisando o que aconteceu
#
# Temos $y_{next} = y_{curr} - 10y_{curr} h = (1 - 10h)y_{curr}$, e se $10h > 1$ teremos uma solução oscilatória.
# Que é qualitativamente diferente da solução real!
| comp-cientifica-I-2018-2/semana-4/raw_files/Euler instavel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Employee Churn Analysis
# # What is Employee Churn ?
# When an employee leaves the organization is known as churn.
# Employee churn will be affected by age, tenure, pay, job satisfaction, salary, working conditions, growth potential
# and employee’s perceptions of fairness.
# Some other variables such as age, gender, ethnicity, education, and marital status, were essential factors
# in the prediction of employee churn.
# Predicting the employee churn will help some proactive measures to be taken for employee recruitment and prevention.
# For the given HR dataset, Do the Employee Churn Analysis
#
# # About the attributes of the dataset - HR.csv
# 1. satisfaction_level: It is employee satisfaction point, which ranges from 0-1.
# 2. last_evaluation: It is evaluated performance by the employer, which also ranges from 0-1.
# 3. number_projects: How many numbers of projects assigned to an employee?
# 4. average_monthly_hours: How many average numbers of hours worked by an employee in a month?
# 5. time_spent_company: time_spent_company means employee experience. The number of years spent by an employee in the company.
# 6. work_accident: Whether an employee has had a work accident or not.
# 7. promotion_last_5years: Whether an employee has had a promotion in the last 5 years or not.
# 8. Departments: Employee's working department/division.
# # Stmt: Find out the Pattern of Employee churn and Predict the future churn.
# Hints:
# 1. Filter out the employees who has left
# 2. Find out the minimum 3 group of employees based on satisfaction level and the evaluation of the employee
# 3. Group 1 : high satisfaction and high evalation
# Group 2: Low satisfaction and high Evaluation
# Group 3: moderate satisfaction and moderate evaluation
# 4. Find out how many in each category fits on to 3 groups.
# 5. Filter out the employees who has not left
# 6. Predict how many employees in the organization may leave according to the pattern modelled earlier with 3 groups
import numpy as np
import pandas as pd
df=pd.read_csv('C:/Users/mozhi/Desktop/DA 2019/Assessment/ISA2/HR.csv')
df.head()
#import module
from sklearn.cluster import KMeans
# Filter data
left_emp = df[['satisfaction_level', 'last_evaluation']][df.left == 1]
# Create groups using K-means clustering.
kmeans = KMeans(n_clusters = 3, random_state = 0).fit(left_emp)
# Add new column "label" annd assign cluster labels.
import matplotlib.pyplot as plt
left_emp['label'] = kmeans.labels_
# Draw scatter plot
plt.scatter(left_emp['satisfaction_level'], left_emp['last_evaluation'], c=left_emp['label'])
plt.xlabel('Satisfaction Level')
plt.ylabel('Last Evaluation')
plt.title('3 Clusters of employees who left')
plt.show()
left_emp['label'].value_counts()
about_to_leave = df[['satisfaction_level', 'last_evaluation']][df.left == 0]
clus=kmeans.predict(about_to_leave)
clus
about_to_leave['labels']=clus
about_to_leave.head()
about_to_leave['labels'].value_counts()
# Draw scatter plot
plt.scatter(about_to_leave['satisfaction_level'], about_to_leave['last_evaluation'], c=about_to_leave['labels'])
plt.xlabel('Satisfaction Level')
plt.ylabel('Last Evaluation')
plt.title('3 Clusters of employees who is about_to_leave')
plt.show()
# +
#Conclusion: 32 % may leave the organization
| Sem4/DataAnalytics/DA Lab - students copy/Clustering/.ipynb_checkpoints/ISA2-Lab solution-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# I am creating all inputs for scrapy and crawl
# mkdir code
# cd code
import scrapy
# This below creates requirements for scrapy and crawl
# %%cmd
scrapy startproject tutorial
# cd tutorial
# %%cmd
scrapy genspider nba nba.com
# cd tutorial
# +
# # %load items.py
# Define here the models for your scraped items
#
# See documentation in:
# https://doc.scrapy.org/en/latest/topics/items.html
import scrapy
class TutorialItem(scrapy.Item):
# define the fields for your item here like:
# name = scrapy.Field()
pass
# -
# cd C:\Users\Sandeep\big-data-python-class\Homeworks\Homework8\code\tutorial\tutorial\spiders
# Here I take the initially corresponding url's of www.nba.com and will start crawling web page urls into different categories for future usage of numpys.
# +
# # %load nba.py
import scrapy
class nbaItem(scrapy.Item):
title = scrapy.Field()
link = scrapy.Field()
resp = scrapy.Field()
class NbaSpider(scrapy.Spider):
name = 'ba'
allowed_domains = ['amazon.com']
start_urls = ['http://www.nba.com/scores#/','https://stats.nba.com/schedule/','http://www.nba.com/news','https://stats.nba.com/']
def parse(self, response):
res = scrapy.Selector(response)
titles = res.xpath('//ul/li')
items = []
for title in titles:
item = nflItem()
item["title"] = title.xpath("a/text()").extract()
item["link"] = title.xpath("a/@href").extract()
item["resp"] = response
if item["title"] != []:
items.append(item)
return items
# -
# This will crawl all the webpages recursively going into deeper url's
# %%cmd
scrapy crawl nba
# Moving all crawl pages to csv
# %%cmd
scrapy crawl nba -o results.csv -t csv
import pandas as pd
temp =[]
data=pd.read_csv("results.csv")
data['link']='http://www.nba.com'+data['link']
x=data['resp'][0]
print x
temp.append((x.split()[-1]).split('>')[-2])
print temp
for i in range(len(data)):
if data['resp'][i] == x:
temp.append(data['link'][i])
else:
x=data['resp'][i]
nba_items=list(pd.DataFrame(temp)[0].unique())
links = []
length = len(temp)
for i, val in enumerate(temp):
if i < length-1:
links.append((temp[i], temp[i+1]))
# Stochastic Matrix
import numpy as np
n = pd.DataFrame(index=nba_items, columns=nba_items)
m = n.replace(np.NaN, 0)
for i in links:
m.loc[i] = 1.0
arr = np.array(m)
v = arr.sum(axis=1)
res = arr/v[:, np.newaxis]
sophist_matrix = np.nan_to_num(res)
sophist_matrix
# # PageRank Algorithm
# +
import numpy as np
from scipy.sparse import csc_matrix
def pageRank(G, s = .85, maxerr = .001):
"""
Computes the pagerank for each of the n states.
Used in webpage ranking and text summarization using unweighted
or weighted transitions respectively.
Args
----------
G: Stochastic Matrix
Kwargs
----------
s(theta): probability of following a transition. 1-s probability of teleporting
to another state. Defaults to 0.85
maxerr: if the sum of pageranks between iterations is bellow this we will
have converged. Defaults to 0.001
"""
n = G.shape[0]
# transform G into markov matrix M
M = csc_matrix(G,dtype=np.float)
rsums = np.array(M.sum(1))[:,0]
ri, ci = M.nonzero()
M.data /= rsums[ri]
# bool array of sink states
sink = rsums==0
# Compute pagerank r until we converge
ro, r = np.zeros(n), np.ones(n)
while np.sum(np.abs(r-ro)) > maxerr:
ro = r.copy()
# calculate each pagerank at a time
for i in xrange(0,n):
# inlinks of state i
Ii = np.array(M[:,i].todense())[:,0]
# account for sink states
Si = sink / float(n)
# account for teleportation to state i
Ti = np.ones(n) / float(n)
r[i] = ro.dot( Ii*s + Si*s + Ti*(1-s) )
# return normalized pagerank
return r/sum(r)
#print pageRank(pd.DataFrame(pagerank(sophist_matrix)),s=.86)
# -
Rank_val=pd.DataFrame(pageRank(sophist_matrix,s=.86))
#Rank_val = pd.DataFrame(pagerank(sophist_matrix))
Rank_sort = (Rank_val.sort_values(0)).head(10)
Rank_lt = list(Rank_sort.index)
print Rank_lt
url = pd.DataFrame(nba_items)
print "Top 10 page URLs:"
for i in Rank_lt:
print url[0][i]
# # Hits Algorithm
def hits(A):
n= len(A)
Au= dot(transpose(A),A)
Hu = dot(A,transpose(A))
a = ones(n);
h = ones(n)
#print a,h
for j in range(5):
a = dot(a,Au)
a= a/sum(a)
h = dot(h,Hu)
h = h/ sum(h)
return h
hit_val = pd.DataFrame(hits(sophist_matrix))
hit_sort = (hit_val.sort_values(0, ascending=False)).head(10)
hit_lt = list(hit_sort.index)
url = pd.DataFrame(nba_items)
print "Top 10 page URLs:"
for i in hit_lt:
print url[0][i]
| Homeworks/Rongali_linkanalysis_2018/Rongali_linkanalysis_2018.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] id="q388X88DDV9W" colab_type="text"
#
# + id="i0y4JGUuDYSu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 241} outputId="4e455cca-143f-40ca-e176-6fe83d59174f"
REPO_NAME = 'Random-Forest-forecasting' #@param {type:"string"}
from google.colab import drive
drive.mount('/content/gdrive')
# %run /content/gdrive/My\ Drive/lib/InitLib.ipynb
# + id="gM6JtpqJIwBj" colab_type="code" colab={}
# + id="nm_Qx4xfDR7a" colab_type="code" colab={}
import pandas as pd
import numpy as np
import matplotlib.pylab as plt
from statsmodels.tsa.seasonal import seasonal_decompose
from statsmodels.tsa.ar_model import AR
from sklearn.metrics import r2_score
# %matplotlib inline
plt.rcParams['figure.figsize']=(20,10)
plt.style.use('ggplot')
# + id="aznVZmm8DR8W" colab_type="code" colab={}
sales_data = pd.read_csv(REPO_DIR + '/Autoregression/retail_sales.csv')
sales_data['date']=pd.to_datetime(sales_data['date'])
sales_data.set_index('date', inplace=True)
# + id="p1P5ybTHDR83" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="c4699916-f993-4fbb-b49c-b0617936496e"
sales_data.head()
# + id="gghhnyPUDR9m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 649} outputId="42590904-3100-41a9-8961-64b330bca788"
sales_data.plot()
# + id="srz_Px-jDR-E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 749} outputId="ec3aab96-c2a9-4d52-911c-306a8bdabdad"
decomposed = seasonal_decompose(sales_data['sales'], model='additive')
x =decomposed.plot() #See note below about this
# + id="ycAqHuKIDR-i" colab_type="code" colab={}
sales_data['stationary']=sales_data['sales'].diff()
# + id="ETkcZAVuDR-5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="7b518762-9a91-4321-d791-8d6ef9ad9c41"
sales_data.head()
# + id="R5vVecyPDR_U" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 649} outputId="17c80d01-28af-468e-8aa2-8b7865a78660"
sales_data['stationary'].plot()
# + id="iVyBLBNADR_w" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 749} outputId="eafb52d9-2544-484a-c622-de80a32f97aa"
decomposed = seasonal_decompose(sales_data['stationary'].dropna(), model='additive')
x =decomposed.plot() #See note below about this
# + id="8IAqp3MIDSAO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 649} outputId="32f2021e-d52a-4b18-fae8-6e3bed8205f3"
pd.plotting.lag_plot(sales_data['sales'])
# + id="tFeHShmpDSAq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 650} outputId="1c6f3252-b2c6-4ec8-a498-a490de482159"
pd.plotting.autocorrelation_plot(sales_data['sales'])
# + id="H5WNMK5mDSBF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8f24368c-fc13-453b-ba5c-d2b97dceb452"
sales_data['sales'].corr(sales_data['sales'].shift(12))
# + id="5E8R3t89DSBk" colab_type="code" colab={}
#create train/test datasets
X = sales_data['stationary'].dropna()
train_data = X[1:len(X)-12]
test_data = X[X[len(X)-12:]]
# + id="Y-CCR9kRDSB5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="c5876f86-aa70-4493-d007-c68ee8954975"
#train the autoregression model
model = AR(train_data)
model_fitted = model.fit()
# + id="5oWKtFEYDSDE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="834d6e21-413a-4389-d6c4-93f90c6e8a6c"
print('The lag value chose is: %s' % model_fitted.k_ar)
# + id="2WcLIv5SDSDj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="fc9acc9c-c19e-4845-d6d5-25ca3410fa1e"
print('The coefficients of the model are:\n %s' % model_fitted.params)
# + id="bzsxKJLMDSEG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} outputId="e7363a57-179f-411e-c3dc-9744b13280cd"
# make predictions
predictions = model_fitted.predict(
start=len(train_data),
end=len(train_data) + len(test_data)-1,
dynamic=False)
# create a comparison dataframe
compare_df = pd.concat(
[sales_data['stationary'].tail(12),
predictions], axis=1).rename(
columns={'stationary': 'actual', 0:'predicted'})
# + id="n8SAExZqDSEi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 421} outputId="2e107f11-c077-4d99-ec10-643ae7fd2a4d"
compare_df
# + id="26Ls5QanDSFK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 644} outputId="762d5096-f7db-44d9-c0a6-3d557c74ee00"
compare_df.plot()
# + id="aoe-b2DMDSFn" colab_type="code" colab={}
r2 = r2_score(sales_data['stationary'].tail(12), predictions)
# + id="kg8v3LmDDSGH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a624aa4f-e8a6-4dfb-cd46-5682ad29cf68"
r2
# + id="waECLj8oDSGk" colab_type="code" colab={}
| Autoregression/Autoregression_retail_sales.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Map of sub-domain of ANHA12
# +
import matplotlib.pyplot as plt
import netCDF4 as nc
import numpy as np
import cmocean
import pickle
# %matplotlib inline
# -
# #### Parameters:
# +
# domain dimensions:
imin, imax = 1479, 2179
jmin, jmax = 159, 799
# Resolution of grid cells to show in figure:
grid_reso = 10
# -
# #### Load files:
# ANHA12 grid
mesh = nc.Dataset('/ocean/brogalla/GEOTRACES/data/ANHA12/ANHA12_mesh1.nc')
mesh_lon = np.array(mesh.variables['nav_lon'])
mesh_lat = np.array(mesh.variables['nav_lat'])
mesh_bathy = np.array(mesh.variables['hdept'][0])
# #### Figure:
# +
fig, ax1, proj1, ax2, proj2 = pickle.load(open('/ocean/brogalla/GEOTRACES/pickles/surface-land-map-globe.pickle','rb'))
# Globe: --------------------------------------------------------------------------------------
x, y = proj2(mesh_lon, mesh_lat)
bath = proj2.contourf(x, y, mesh_bathy, 20, cmap=cmocean.cm.tempo, vmin=0, vmax=6000, zorder=1)
# Sub-domain
proj2.plot(x[imin:imax,jmin], y[imin:imax,jmin], 'w-', lw=0.9, zorder=3)
proj2.plot(x[imin:imax,jmax], y[imin:imax,jmax], 'w-', lw=0.9, zorder=3)
proj2.plot(x[imin,jmin:jmax], y[imin,jmin:jmax], 'w-', lw=0.9, zorder=3)
proj2.plot(x[imax,jmin:jmax], y[imax,jmin:jmax], 'w-', lw=0.9, zorder=3)
# full domain
proj2.plot(x[460:-1:grid_reso,0] , y[460:-1:grid_reso,0] , 'w--', lw=0.8, zorder=3, dashes=(2, 2))
proj2.plot(x[0,370:-1:grid_reso] , y[0,370:-1:grid_reso] , 'w--', lw=0.8, zorder=3, dashes=(2, 2))
proj2.plot(x[100:-1:grid_reso,-1] , y[100:-1:grid_reso,-1] , 'w--', lw=0.8, zorder=3, dashes=(2, 2))
proj2.plot(x[-1,0:-1:grid_reso] , y[-1,0:-1:grid_reso] , 'w--', lw=0.8, zorder=3, dashes=(2, 2))
xline_add = [x[500,0], x[700,70], x[1100,0], x[0,370]]
yline_add = [y[240,-1], y[85,-80], y[80,0], y[0,370]]
proj2.plot(xline_add, yline_add, 'w--', lw=0.8, zorder=3, dashes=(2, 2))
# Sub-domain map: ---------------------------------------------------------------------------
x_sub, y_sub = proj1(mesh_lon, mesh_lat)
proj1.plot(x_sub[imin:imax,jmax] , y_sub[imin:imax,jmax] , 'w-', lw=1.2, zorder=5)
proj1.plot(x_sub[imin:imax,jmin] , y_sub[imin:imax,jmin] , 'w-', lw=1.2, zorder=5)
proj1.plot(x_sub[imin,jmin:jmax] , y_sub[imin,jmin:jmax] , 'w-', lw=1.2, zorder=5)
proj1.plot(x_sub[imax,jmin:jmax] , y_sub[imax,jmin:jmax] , 'w-', lw=1.2, zorder=5)
proj1.plot(x_sub[imin:imax+grid_reso:grid_reso,jmin:jmax+grid_reso:grid_reso], \
y_sub[imin:imax+grid_reso:grid_reso,jmin:jmax+grid_reso:grid_reso], 'w-', lw=0.4, zorder=3);
proj1.plot(x_sub[imin:imax+grid_reso:grid_reso,jmin:jmax+grid_reso:grid_reso].T, \
y_sub[imin:imax+grid_reso:grid_reso,jmin:jmax+grid_reso:grid_reso].T, 'w-', lw=0.4, zorder=3);
# Color bar: --------------------------------------------------------------------------------
cbaxes = fig.add_axes([0.92, 0.18, 0.02, 0.35]);
CBar = plt.colorbar(bath, ax=ax1, cax=cbaxes);
CBar.set_label('Ocean depth [m]', fontsize=8)
CBar.ax.tick_params(axis='y', length=0, labelsize=8)
fig.savefig('/ocean/brogalla/GEOTRACES/figures/paper1-202110/M1-sub-domain.png', bbox_inches='tight', dpi=300)
fig.savefig('/ocean/brogalla/GEOTRACES/figures/paper1-202110/M1-sub-domain.svg', bbox_inches='tight', format='svg', dpi=300)
# -
| paper-materials/M1-sub-domain-map.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # このノートブックについて
#
# [ノートブックへ](https://www.dropbox.com/s/vsfglb1bkth10i3/20200622-kw-request-handler.ipynb?dl=0)
#
# Bokeh のウィジェットを利用した場合、そのコールバックは原則的に JavaScript で記述する。使い慣れた Python でコッルバックを書きたい、あるいは Python 側にあるデータをアクセスしたい、サーバ側のデータベースにアクセスしたいなどの理由から Python でコールバックを書きたいことも多い。
#
# このノートブックは、JavaScript で書かれた小さなコールバックから、Python の関数を起動し、**あたかも** その関数がウィジェット用のコールバックのように振る舞う仕掛けを作っている。
#
# Python のコールバック関数はウィジェットを直接的に監視することはできない。そこで、この関数が監視できるデータソースを用意し、それを JavaScript のコールバックに受け渡し、コールバックのなかでその内容を更新する。Bokeh は、この JavaScript 側での更新を Python 側の対応するデータソースへの更新として反映する。この Python 側での更新により、Python 側のデータソースを監視している Python で書かれたコールバック関数が駆動される。
import numpy as np
from bokeh.plotting import figure, output_notebook, show
from bokeh.models import HoverTool, ColumnDataSource, CustomJS
# 以下の JavaScript 用のコッルバックでは、Hover された点の ID を JavaScript のデータ構造に保存された選択されているデータ群 (`cb_data.index.indices`) をPython のデータソースに移植しているらしい。
# - (`console.log` で確認したところ)`js_callback` は、Hover とは無関係にマウスの移動のごとに呼ばれるらしい。
# - `if` 文はなにを確認しているのか?
# - `data_source` が確かに Python のデータソースに該当することを確認すること。
#
JS_CODE = '''
if (cb_data.renderer.id==glyph.id) {
selection.selected.indices = cb_data.index.indices;
}'''
# - `selection` は空のデータソース。ここに Hover ツールで選択されたデータの情報が書き込まれる。
def ask_js_callback_to_call(hover_tool, glyph, py_callback):
selection = ColumnDataSource()
selection.selected.on_change('indices', py_callback)
# 選択された領域が更新されたときに JS コールバックを読む仕組み
js_callback = CustomJS(args={'selection': selection, 'glyph': glyph}, code=JS_CODE)
print(hover_tool.callback)
#hover_tool.callback = js_callback if hover_tool.callback is None else hover_tool.callback + [js_callback]
hover_tool.callback = js_callback
# ## Bokeh ドキュメントの構成
#
# - $[0,1]^2$の領域に無作為に青い点を$B$個、赤い点を$R$個ずつ配置する。
# - 一見すると赤い点がホバーされたときのコールバック(`py_callback`)が Hover ツールから呼ばれるように見える。しかし、実際に設定されているのは JavaScript のコールバック (`js_callback`) である。`js_callback` が裏に隠れたデータソースの更新 (`indices`イベント) を介して、間接的に `py_callback` が呼ばれるように仕向ける仕掛けになっている。
def Document(doc):
(B, R) = 5, 5
fig = figure()
blue_points = fig.circle(x='x', y='y', fill_color='blue', line_color=None,
source=ColumnDataSource(dict(x=np.random.rand(B), y=np.random.rand(B))))
red_points = fig.circle(x='x', y='y', fill_color='red', line_color=None,
source=ColumnDataSource(dict(x=np.random.rand(R), y=np.random.rand(R))))
def py_callback(action, _, i):
if len(i) > 0:
print(f'{action}({i[0]}) -> x={red_points.data_source.data["x"][i][0]:.3f}')
hover_tool = HoverTool()
ask_js_callback_to_call(hover_tool, red_points, py_callback)
fig.add_tools(hover_tool)
doc.add_root(fig)
output_notebook()
show(Document)
| bokeh/interaction/20200622-kw-request-handler.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # DLA Abund Cookbook (v1.0)
# +
# imports
# suppress warnings for these examples
import warnings
warnings.filterwarnings('ignore')
import json, io
import numpy as np
import astropy.units as u
from linetools.isgm.abscomponent import AbsComponent
from linetools.lists.linelist import LineList
from linetools.spectra.io import readspec
import linetools
from linetools.abund.relabund import RelAbund
from pyigm.abssys.dla import DLASystem
# -
# ## 0. Analyze NHI, Continuum fit the spectrum (or spectra)
#
# These steps are not described here.
#
# For continuum fitting, we recommend lt_continuum from linetools
#
# lt_continuum spec.fits
# ## 1. Generate a simple DLA system
#
# Requires $z$, $N_{\rm HI}$ and the coordinates
#
# $v_{\rm lim}$ defaults to 500 km/s if input as None
# PH957
vlim = None
dla = DLASystem(('01:03:11.38','+13:16:16.7'), 2.309, vlim, 21.37, sig_NHI=[0.08,0.08])
dla
# Write
dla.write_json()
# ## 2. AbsKin GUI for Line IDs
#
# * Requires xastropy for now
# * Best run off the JSON file
# * And from the command line
#
# lt_xabssys PH957_f.fits J010311.38+131616.7_z2.309.json -outfile J010311.38+131616.7_z2.309_HIRES.json
#
#
#
# * User modifies the velocity limits
# * Identifies blends
# * Sets upper/lower limits as desired
# * Rejects lines that are not worth analysis
# Load
json_fil = '../../pyigm/abssys/tests/files/J010311.38+131616.7_z2.309_ESI.json'
dla = DLASystem.from_json(json_fil)
# ## 3. Measure EWs and AODM columns
#
# Load spectrum from linetools
spec_fil = linetools.__path__[0]+'/spectra/tests/files/PH957_f.fits'
spec = readspec(spec_fil)
# EW
dla.measure_restew(spec=spec)
# AODM
dla.measure_aodm(spec=spec)
# View
dla.fill_trans()
dla._trans
# ## 4. Update Components
#
# The fill_ionN method synthesizes multiple components of the same ion (and with the
# same $E_j$). These are then pushed into the _ionN table.
dla.update_component_colm()
dla.fill_ionN()
dla._ionN
# ## 5. Relative Abundances (relative to Solar)
#
# The following method takes all column densities measured
# from low-ion transitions from the ground state. It assumes
# no ionization corrections.
dla.XY = RelAbund.from_ionclm_table((1,dla.NHI, dla.sig_NHI[0]), dla._ionN)
dla.XY.elements['H'].name
dla.XY.table()
dla.XY.table('Fe')
| docs/examples/DLA_abund_cookbook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Import libraries
# +
from matplotlib import pyplot as plt
import numpy as np
# %matplotlib inline
# -
# ## Define function
# The goal is to create a plot of the following function
# \begin{equation}
# f(x)=0.2+0.4x^2+0.3x\cdot\sin(15x)+0.05\cos(50x)
# \end{equation}
x = np.linspace(0, 1, 100)
y = 0.2+0.4*x**2+0.3*x*np.sin(15*x)+0.05*np.cos(50*x)
# ## Produce figure
plt.figure(figsize=(6, 6))
plt.plot(x, y)
plt.show()
| function-plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="awABtmzB5KNF" colab_type="text"
# ##############################################################################
#
# This notebook uses the twarc tool, to hydrate tweet IDs
#
# These tweet IDs are collected via the Covid19 Streaming endpoint.
#
# ##############################################################################
# + id="LyyQIn8ue420" colab_type="code" colab={}
from google.colab import drive
drive.mount('/content/drive')
# + id="m14P-EDSfHyw" colab_type="code" colab={}
# cd 'drive/My Drive/coronawhy'
# + id="XaLfwlG4fY41" colab_type="code" colab={}
# !pip install twarc
# + id="3dJvpnwDfatf" colab_type="code" colab={}
# !pip install tqdm
# + id="KcRZZC5-fch0" colab_type="code" colab={}
# !twarc configure
# + id="613oxyTpfgfw" colab_type="code" colab={}
# !python hydra.py
# + id="P2boEnoLgB9_" colab_type="code" colab={}
| Hydrating_notebook_from_CLI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/BButa/dw_matrix_car/blob/master/day4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="fKh-JntxDs5w" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 493} outputId="11919269-4829-458c-bc1f-625843005540"
# !pip install --upgrade tables
# !pip install eli5
# !pip install xgboost
# + id="xANGjHdjD5dD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 168} outputId="300374e0-b324-490b-d7b3-abfd36af1302"
import pandas as pd
import numpy as np
from sklearn.dummy import DummyRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
import xgboost as xgb
from sklearn.metrics import mean_absolute_error as mae
from sklearn.model_selection import cross_val_score, KFold
import eli5
from eli5.sklearn import PermutationImportance
# + id="EqLUym3sERyQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7c203930-e363-4d39-ddb9-ffe54bcaeaf7"
# cd "/content/drive/My Drive/Colab Notebooks/dw_matrix/matrix_two/dw_matrix_car"
# + id="3u4196tIEVdR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3f76a258-5f02-480f-e7c8-5b103acb8345"
df = pd.read_hdf('data/car.h5')
df.shape
# + id="seuTrRVDEc3A" colab_type="code" colab={}
# + [markdown] id="c37eWkNvEfm3" colab_type="text"
# Feature Engineering
# + id="jXXFY4-hEkI5" colab_type="code" colab={}
SUPFIX_CAT = '__cat'
for feat in df.columns:
if isinstance(df[feat][0], list): continue
factorized_values=df[feat].factorize()[0]
if SUPFIX_CAT in feat:
df[feat] = factorized_values
else:
df[feat + SUPFIX_CAT] =factorized_values
# + id="Ib6gD9ZPEm_a" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2169000d-717b-4560-8021-7fa354d0d74c"
cat_feats = [ x for x in df.columns if SUPFIX_CAT in x ]
cat_feats = [x for x in cat_feats if 'price' not in x]
len(cat_feats)
# + id="JCSAFuVpEqhJ" colab_type="code" colab={}
def run_model(model, feats):
X = df[feats].values
y= df['price_value'].values
scores = cross_val_score(model, X, y, cv=3, scoring='neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
# + [markdown] id="wwb2aHrrFjBt" colab_type="text"
# Decision Tree
# + id="S7nltuLSEuQh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="726e675e-4b6a-4150-ea48-5e5dd6aa4291"
run_model( DecisionTreeRegressor(max_depth=5), cat_feats)
# + [markdown] id="Tfo_L2CeFq3W" colab_type="text"
# ## Random Forest
#
# + id="gw_iIWDzFdzS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="69b52ae4-4436-45b2-88f5-1ce5cc3d784e"
model = RandomForestRegressor(max_depth=5,n_estimators=50, random_state=0)
run_model(model, cat_feats)
# + id="WXm5VPARF8e8" colab_type="code" colab={}
# + [markdown] id="jkdByIkxIVi1" colab_type="text"
# ## XGBoost
# + id="532qLcedIX5y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="487564ca-e0bd-4a15-98d4-17df9ff8bcaf"
xgb_params={
'max_deapth': 5,
'n_estimators': 50,
'learning_rate': 0.5,
'seed': 0
}
run_model( xgb.XGBRegressor(**xgb_params), cat_feats)
# + id="t7BEaD1dPxq_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 408} outputId="909cfd90-3bdc-45be-94af-89a45fe00f42"
m = xgb.XGBRegressor(max_depth=5, n_estimators=50, learning_rate=0.1, seed=0)
m.fit(X,y)
imp = PermutationImportance(m, random_state=0).fit(X,y)
eli5.show_weights(imp,feature_names =cat_feats)
# + id="yLXRkRzSUZ6X" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="830fcd09-69a1-436c-b8e7-014965028924"
len(cat_feats)
# + id="qwICm-5qQmkP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="09dd9046-2dc7-47c5-a528-173d1f847718"
feats = [
'param_napęd__cat',
'param_rok-produkcji__cat',
'param_stan__cat',
'param_skrzynia-biegów__cat',
'param_faktura-vat__cat',
'param_moc__cat',
'param_marka-pojazdu__cat',
'feature_kamera-cofania__cat',
'param_typ__cat',
'param_pojemność-skokowa__cat',
'seller_name__cat',
'feature_wspomaganie-kierownicy__cat',
'param_model-pojazdu__cat',
'param_wersja__cat',
'param_kod-silnika__cat',
'feature_system-start-stop__cat',
'feature_asystent-pasa-ruchu__cat',
'feature_czujniki-parkowania-przednie__cat',
'feature_łopatki-zmiany-biegów__cat',
'feature_regulowane-zawieszenie__cat',]
run_model( xgb.XGBRegressor(**xgb_params), feats)
# + id="J0hcDrHlUfgV" colab_type="code" colab={}
df['param_rok-produkcji'] =df['param_rok-produkcji'].map(lambda x: -1 if str(x)== 'None' else int(x) )
# + id="wWmLtbxrU5iG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="a715fa96-5319-49a6-f51b-135337c1ef29"
feats = [
'param_napęd__cat',
'param_rok-produkcji',
'param_stan__cat',
'param_skrzynia-biegów__cat',
'param_faktura-vat__cat',
'param_moc__cat',
'param_marka-pojazdu__cat',
'feature_kamera-cofania__cat',
'param_typ__cat',
'param_pojemność-skokowa__cat',
'seller_name__cat',
'feature_wspomaganie-kierownicy__cat',
'param_model-pojazdu__cat',
'param_wersja__cat',
'param_kod-silnika__cat',
'feature_system-start-stop__cat',
'feature_asystent-pasa-ruchu__cat',
'feature_czujniki-parkowania-przednie__cat',
'feature_łopatki-zmiany-biegów__cat',
'feature_regulowane-zawieszenie__cat',]
run_model( xgb.XGBRegressor(**xgb_params), feats)
# + id="Xi_m5bV6VcWG" colab_type="code" colab={}
df['param_moc'] = df['param_moc'].map(lambda x: -1 if str(x)== 'None' else int(str(x).split(' ')[0]))
# + id="yQXFsiAoWKZf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="547b0750-e796-42f6-86e2-1baef1d529c8"
feats = [
'param_napęd__cat',
'param_rok-produkcji',
'param_stan__cat',
'param_skrzynia-biegów__cat',
'param_faktura-vat__cat',
'param_moc',
'param_marka-pojazdu__cat',
'feature_kamera-cofania__cat',
'param_typ__cat',
'param_pojemność-skokowa__cat',
'seller_name__cat',
'feature_wspomaganie-kierownicy__cat',
'param_model-pojazdu__cat',
'param_wersja__cat',
'param_kod-silnika__cat',
'feature_system-start-stop__cat',
'feature_asystent-pasa-ruchu__cat',
'feature_czujniki-parkowania-przednie__cat',
'feature_łopatki-zmiany-biegów__cat',
'feature_regulowane-zawieszenie__cat',]
run_model( xgb.XGBRegressor(**xgb_params), feats)
# + id="Npmnb1uqWTg1" colab_type="code" colab={}
df['param_pojemność-skokowa'] = df['param_pojemność-skokowa'].map(lambda x: -1 if str(x)== 'None' else int(str(x).split('cm')[0].replace(' ','')))
# + id="8z3q_132dOXq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="0d56a4e1-c0e5-4208-a1c5-8950d57f0bd2"
feats = [
'param_napęd__cat',
'param_rok-produkcji',
'param_stan__cat',
'param_skrzynia-biegów__cat',
'param_faktura-vat__cat',
'param_moc',
'param_marka-pojazdu__cat',
'feature_kamera-cofania__cat',
'param_typ__cat',
'param_pojemność-skokowa',
'seller_name__cat',
'feature_wspomaganie-kierownicy__cat',
'param_model-pojazdu__cat',
'param_wersja__cat',
'param_kod-silnika__cat',
'feature_system-start-stop__cat',
'feature_asystent-pasa-ruchu__cat',
'feature_czujniki-parkowania-przednie__cat',
'feature_łopatki-zmiany-biegów__cat',
'feature_regulowane-zawieszenie__cat',]
run_model( xgb.XGBRegressor(**xgb_params), feats)
# + id="Nw0DI6WfeCEY" colab_type="code" colab={}
| day4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Announcements
# - Please familiarize yourself with the term projects, and sign up for your (preliminary) choice using [this form](https://forms.gle/ByLLpsthrpjCcxG89). _You may revise your choice, but I'd recommend settling on a choice well before Thanksgiving._
# - Recommended reading on ODEs: [Lecture notes by Prof. Hjorth-Jensen (University of Oslo)](https://www.asc.ohio-state.edu/physics/ntg/6810/readings/hjorth-jensen_notes2013_08.pdf)
# - Problem Set 4 posted on D2L, due Oct 5.
# - __Outlook__: completing ODE examples this week, then Partial Differential Equations (PDEs)
# - Conference for Undergraduate Women in Physics: online event in 2021, [applications accepted until 10/25](https://www.aps.org/programs/women/cuwip/)
# # Special Considerations: Symplectic Integrators
# Some systems of ODEs have special properties, for example constants of motion, that should be
# conserved by the system. Simple examples in mechanics are the energy, linear momentum, and angular
# momentum. Sometimes it is more important to build in to the integration scheme these conservation
# laws than to achieve a high accuracy per step size. Normally, the error in the solution grows as the
# solution evolves, bounded by the number of steps times the error per step. For certain schemes,
# however, while they still make errors, the errors will be of such a form that the conserved quantities
# will be better preserved.
#
# In Lecture 15 we derived the __velocity form__ of the Verlet algorithm
# \begin{align}
# x_{n+1} &= x_n + v_n\delta t + {1\over 2}a_n(\delta t)^2 + \mathcal{O}(\delta t^3) \tag{1a}\\
# a_{n+1} &= a(x_{n+1}) \label{eq:Vv}\tag{1b}\\
# v_{n+1} &= v_n + {1\over 2}\left(a_n + a_{n+1}\right)\delta t + \mathcal{O}(\delta t^2).\tag{1c}
# \end{align}
#
# While the velocity computed in this way is only
# first-order accurate, if the acceleration depends only upon position, then the
# velocity appears only where multiplied by $\delta t$ in determining the positions and so
# second-order accuracy is preserved. Of course, if determining the velocity to
# high accuracy is required, then one might turn to a different scheme. On the
# other hand, if the acceleration depends upon the velocity (for example, a drag
# term), there are fewer constants of the motion to be conserved anyway, and one
# of the higher-order schemes developed previously would serve.
# <span style="color:red"> Note that Eq.(\ref{eq:Vv}) evaluates $a_{n+1}$ before $v_{n+1}$ can be computed, so the acceleration should only be a function of position, not velocity. _This is by design, as the derivation of the Verlet algorithm was based on conservation of energy._ </span>
#
# Here's one implementation of the velocity Verlet algorithm. Note that we compute the acceleration $a_n$ and $a_{n+1}$ in each call; this is not optimal computationally, but easier to read and use with our existing `odeSolve` code.
import numpy as np
def vVerlet_step(g, t, x,v, h, *P):
"""Implements a full step h of the velocity Verlet algorithm
Parameters:
g: function. RHS of ODE, with signature g(t,y[0],*P).
x: Current position x.
v: Current velocity v.
h: float. Step length.
*P: tuple, additional parameters for g function
Returns:
x,v: numpy arrays: next position, next velocity
"""
a_current = g(x,*P)
#advance the position: x_n -> x_n+1
x += v*h+a_current*h**2
#compute a_n+1
a_next = g(x,*P)
#advance the velocity v_n -> v_n+1
v +=(a_current+a_next)*h**2
return x,v
# ## Example: Planetary Motion
# Today we will study the orbits of two masses interacting through gravity.
#
# In general, the gravitational force on object $j$ with mass $m_j$ and position $\vec{r}_j$ through its gravitational interaction with objects $i$ with mass $m_i$ and position $\vec{x}_i$ is given by
# $$
# \vec F_j = -\sum_{i\ne j}\frac{G m_j m_i}{|\vec x_i - \vec x_j|^2} \times \frac{\left(\vec x_i - \vec x_j\right)}{|\vec x_i - \vec x_j|}\,
# $$
# with the unit vector $\hat{\vec{x}} = \frac{\vec{x}}{|x|}$. _Note the direction of this force, gravity should pull two particles closer to each other! If that's not the case in your implementation, this is a good point to start..._
#
# In the __special case of a two-particle system__, we can rewrite this in terms of the separation vector $\vec r = \vec x_1 - \vec x_2$ and total mass $M= m_1+m_2$ such that
# the gravitational acceleration $\vec{a}_i=\vec{F}_i/m_i$ is given by
# \begin{align}
# \vec{a}(\vec r) &= -\frac{G M}{|r|^2} \hat{\vec{r}}\,.\label{eq:aG}\tag{2}
# \end{align}
# In this case, in the center of mass reference frame, the kinetic energy is given by
# $$T = \frac{1}{2}M\dot{\vec r}^2$$
# and the potential energy is given by
# $$U(\vec r) =-\frac{G m_1 m_2 }{|r|} $$
# _Compared to previous lectures, we will now work with vectorized positions and velocities (rather than writing a separate ODE for each coordinate component). This will later require a small adaption to our function `odeSolve`:_
#
# <span style="color:blue">
# Now use `vVerlet_step` to __integrate simplified planetary orbits__:
# Set $GM = 1$ and try initial conditions
# $$
# x(0) = 0.5, \quad y(0)=0, \quad v_x(0)=0, \quad v_y(0)=1.63\,.
# $$
# You will need to define a function `a_gravity` to evaluate the gravitational force, and later on you will need a function to calculate the potential energy `U_gravity` as well.
#
# ## Velocity Verlet vs RK4: Energy conservation
# Assess the stability of `RK4` and `Velocity Verlet` by checking energy conservation over longer simulation times.
#
# <span style="color:blue"> In order to use `R4K` for integrating this problem, turn Equation(\ref{eq:aG}) into a system of ODEs for the vectorized position and velocity, so that you can reuse the code below. </span>
# +
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
def a_gravity(r, G=1, M=1):
rr = np.sum(r*r)
rhat = r/np.sqrt(rr)
return - G*M/rr * rhat
def U_gravity(r, m=1, G=1, M=1):
#this assumes the r is a 1D vector, not a 2D array
return -G*m*M/np.sqrt(np.sum(r*r))
def analyze_Energy(r,v, m=1, G=1, M=1):
if (len(r.shape) == 1): # r is simple vector
return 0.5*np.sum(v*v)*M+U_gravity(r,m=m,G=G,M=M)
else:
n_steps, ndim = r.shape
E = np.zeros(n_steps)
for i in range(n_steps):
E[i] = 0.5*np.sum(v[i,:]*v[i,:])*M+U_gravity(r[i,:],m=m,G=G,M=M)
def simplified_verlet_wrapper(tmax =2000, h = 1):
dim = 3
r0 = np.array([0.5, 0,0])
v0 = np.array([0, 1.63,0])
mass = 1
nsteps = int(t_max/h)
r = np.zeros((nsteps, dim))
v = np.zeros_like(r)
r[0, :] = r0
v[0, :] = v0
for i in range(nsteps-1):
#write your Verlet integration here, without worrying about consistency with odeSolve conventiosn
return r,v
# -
r,v = simplified_verlet_wrapper()
rx, ry = r[:,0],r[:,1]
ax = plt.subplot(1,1,1)
ax.set_aspect(1)
ax.plot(rx, ry)
# <span style="color:blue"> Write down the system of ODEs that you want to solve with RK4, and define the corresponding `a_gravity_RK4` function.
# </span>
# code copied from Lecture 12/13
# you don't need to change anything in this cell
def RK4_step(t, y, h, g, *P):
"""
Implements a single step of a fourth-order, explicit Runge-Kutta scheme
"""
thalf = t + 0.5*h
k1 = h * g(t, y, *P)
k2 = h * g(thalf, y + 0.5*k1, *P)
k3 = h * g(thalf, y + 0.5*k2, *P)
k4 = h * g(t + h, y + k3, *P)
return y + (k1 + 2*k2 + 2*k3 + k4)/6
# <span style="color:blue"> Next, we will consider the conservation of energy of the velocity Verlet algorithm and RK4.
# Plot the total energy for both solvers as a function of time, for varying step sizes.</span>
| Lectures/Lecture 16/Lecture16_ExampleVerletIntegration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# "Geo Data Science with Python"
# ### Notebook Exercise 6e
#
# ---
#
# # Downloading Science Data with pydap
#
# If you work in teams, please indicate your colaborators below!
NAME = ""
COLLABORATORS = ""
# ---
# # Task description for working with pydap
#
# (20 points)
#
# - On the OPeNDAP test server: http://test.opendap.org/dap/data/nc, find the file: 20070917-MODIS_A-JPL-L2P-A2007260000000.L2_LAC_GHRSST-v01.nc
#
# - Generate a download URL, only for the lat, lon, time and sea_surface_temperature variables
# - Use the pydap package to connect to and inspect the dataset
# - Using pydap, write code to answer the questions:
# - What is the shape of the sea_surface_temperature variable?
# - What is the code to retrieve the fill value of the dataset?
# - What is the shape of the lat/lon variables?
# - What is the code for downloading only the first 500x500 datapoints from all variables?
# - Would you need to create a meshgrid to plot the variable?
#
# - Write a **decent** report below your code, answering all the qustions (up to you if you want to use only markup cells or a combination of code and markup cells.
# - Make a simple plot of the 500x500 subset of the sst variable with plt.scatter()
#
| Exercise06e_pydap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# __Word Alignment Assignment__
#
# Your task is to learn word alignments for the data provided with this Python Notebook.
#
# Start by running the 'train' function below and implementing the assertions which will fail. Then consider the following improvements to the baseline model:
# * Is the TranslationModel parameterized efficiently?
# * What form of PriorModel would help here? (Currently the PriorModel is uniform.)
# * How could you use a Hidden Markov Model to model word alignment indices? (There's an implementation of simple HMM below to help you start.)
# * How could you initialize more complex models from simpler ones?
# * How could you model words that are not aligned to anything?
#
# Grades will be assigned as follows*:
#
# AER below on blinds | Points
# ----------|-------------
# 0.40 - 0.50 | 3
# 0.35 - 0.40 | 6
# 0.30 - 0.35 | 9
# 0.25 - 0.30 | 12
# 0.00 - 0.25 | 15
#
# You should save the notebook with the final scores for 'dev' and 'test' test sets.
#
#
# +
# This cell contains the generative models that you may want to use for word alignment.
# Currently only the TranslationModel is at all functional.
import numpy as np
from collections import defaultdict
class TranslationModel:
"Models conditional distribution over trg words given a src word."
def __init__(self, src_corpus, trg_corpus):
self._trg_given_src_probs = defaultdict(lambda : defaultdict(lambda : 0.0001))
self._src_trg_counts = defaultdict(lambda : defaultdict(lambda : 0.0))
def get_params(self):
return self._trg_given_src_probs
def get_conditional_prob(self, src_token, trg_token):
"Return the conditional probability of trg_token given src_token."
return self._trg_given_src_probs[src_token][trg_token]
def get_parameters_for_sentence_pair(self, src_tokens, trg_tokens):
"Returns matrix with t[i][j] = p(f_j|e_i)."
return np.array([[self._trg_given_src_probs[src_token][trg_token]
for trg_token in trg_tokens] for src_token in src_tokens])
def collect_statistics(self, src_tokens, trg_tokens, posterior_matrix):
"Accumulate counts of translations from: posterior_matrix[i][j] = p(a_j=i|e, f)"
assert posterior_matrix.shape == (len(src_tokens), len(trg_tokens))
assert False, "Implement collection of statistics here."
def recompute_parameters(self):
"Reestimate parameters and reset counters."
self._trg_given_src_probs = defaultdict(lambda : defaultdict(lambda : 0.0))
assert False, "Reestimation parameters from counters here."
class PriorModel:
"Models the prior probability of an alignment given only the sentence lengths and token indices."
def __init__(self, src_corpus, trg_corpus):
"Add counters and parameters here for more sophisticated models."
self._distance_counts = {}
self._distance_probs = {}
def get_parameters_for_sentence_pair(self, src_length, trg_length):
return np.ones((src_length, trg_length)) * 1.0 / src_length
def get_prior_prob(self, src_index, trg_index, src_length, trg_length):
"Returns a uniform prior probability."
return 1.0 / src_length
def collect_statistics(self, src_length, trg_length, posterior_matrix):
"Extract the necessary statistics from this matrix if needed."
pass
def recompute_parameters(self):
"Reestimate the parameters and reset counters."
pass
class TransitionModel:
"Models the prior probability of an alignment conditioned on previous alignment."
def __init__(self, src_corpus, trg_corpus):
"Add counters and parameters here for more sophisticated models."
pass
def get_parameters_for_sentence_pair(self, src_length):
"Retrieve the parameters for this sentence pair: A[k, i] = p(a_{j} = i|a_{j-1} = k)"
pass
def collect_statistics(self, src_length, bigram_posteriors):
"Extract statistics from the bigram_posterior[i][j]: p(a_{t-1} = i, a_{t} = j| e, f)"
pass
def recompute_parameters(self):
"Recompute the transition matrix"
pass
# +
# This cell contains the framework for training and evaluating a model using EM.
from utils import read_parallel_corpus, extract_test_alignments, score_alignments, write_aligned_corpus
def infer_posteriors(src_tokens, trg_tokens, prior_model, translation_model):
"Compute the posterior probability p(a_j=i | f, e) for each target token f_j given e and f."
# P[i][j] = P(a_j=i| I, J): prior alignment prob given only sentence lengths.
P = prior_model.get_parameters_for_sentence_pair(len(src_tokens), len(trg_tokens))
# Emissions prob of seeing f_j given e_i: T[i][j] = P(f_j|e_i)
T = translation_model.get_parameters_for_sentence_pair(src_tokens, trg_tokens)
# posterior[i][j] = P(a_j = i | e, f)
assert False, "Compute the posterior distribution over src indices for each trg word."
return posteriors, log_likelihood
def collect_expected_statistics(src_corpus, trg_corpus, prior_model, translation_model):
"E-step: infer posterior distribution over each sentence pair and collect statistics."
corpus_log_likelihood = 0.0
for src_tokens, trg_tokens in zip(src_corpus, trg_corpus):
# Infer posterior
posteriors, log_likelihood = infer_posteriors(src_tokens, trg_tokens, prior_model, translation_model)
# Collect statistics in each model.
prior_model.collect_statistics(len(src_tokens), len(trg_tokens), posteriors)
translation_model.collect_statistics(src_tokens, trg_tokens, posteriors)
# Update log prob
corpus_log_likelihood += log_likelihood
return corpus_log_likelihood
def estimate_models(src_corpus, trg_corpus, prior_model, translation_model, num_iterations):
"Estimate models iteratively using EM."
for iteration in range(num_iterations):
# E-step
corpus_log_likelihood = collect_expected_statistics(src_corpus, trg_corpus, prior_model, translation_model)
# M-step
prior_model.recompute_parameters()
translation_model.recompute_parameters()
print("corpus log likelihood: %1.3f" % corpus_log_likelihood)
return prior_model, translation_model
def get_alignments_from_posterior(posteriors):
"Returns the MAP alignment for each target word given the posteriors."
alignments = {}
for src_index, trg_index in enumerate(np.argmax(posteriors, 0)):
if src_index not in alignments:
alignments[src_index] = {}
alignments[src_index][trg_index] = '*'
return alignments
def align_corpus(src_corpus, trg_corpus, prior_model, translation_model):
"Align each sentence pair in the corpus in turn."
aligned_corpus = []
for src_tokens, trg_tokens in zip(src_corpus, trg_corpus):
posteriors, _ = infer_posteriors(src_tokens, trg_tokens, prior_model, translation_model)
alignments = get_alignments_from_posterior(posteriors)
aligned_corpus.append((src_tokens, trg_tokens, alignments))
return aligned_corpus
def initialize_models(src_corpus, trg_corpus):
prior_model = PriorModel(src_corpus, trg_corpus)
translation_model = TranslationModel(src_corpus, trg_corpus)
return prior_model, translation_model
def normalize(src_corpus, trg_corpus):
# Consider applying some normalization here to reduce the numbers of parameters.
return src_corpus, trg_corpus
def train(num_iterations):
# Load data
src_corpus, trg_corpus, _ = read_parallel_corpus('en-cs.all')
# Normalize it.
src_corpus, trg_corpus = normalize(src_corpus, trg_corpus)
# Initialize models
prior_model, translation_model = initialize_models(src_corpus, trg_corpus)
# Train them with EM
prior_model, translation_model = estimate_models(src_corpus, trg_corpus, prior_model, translation_model, num_iterations)
# Align the corpus
aligned_corpus = align_corpus(src_corpus, trg_corpus, prior_model, translation_model)
# Return alignments on test sets.
return extract_test_alignments(aligned_corpus)
def evaluate(test_set_alignments):
src_dev, trg_dev, wa_dev = read_parallel_corpus('en-cs-wa.dev', has_alignments=True)
src_test, trg_test, wa_test = read_parallel_corpus('en-cs-wa.test', has_alignments=True)
print('recall %1.3f; precision %1.3f; aer %1.3f' % score_alignments(wa_dev, test_set_alignments['dev']))
print('recall %1.3f; precision %1.3f; aer %1.3f' % score_alignments(wa_test, test_set_alignments['test']))
# -
# Start by running the following cell and fixing some failures:
test_set_alignments = train(10)
evaluate(test_set_alignments)
| resources/_under_construction/19/week06_mt/wa/word_alignment_homework.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import os, shutil, numpy as np, glob, time, pickle as pkl, imars3d
from imars3d.jnbui import ct_wizard, imageslider
from imars3d.ImageFile import ImageFile
# Be patient, this may take a little while too
# # %matplotlib notebook
# %matplotlib inline
from matplotlib import pyplot as plt
# -
# # Input Settings
data_folder = '/Volumes/G-DRIVE/IPTS/IPTS-25519-iMars3D-command-line/raw/ct_scans/2021_07_21_1in/'
ob_folder = '/Volumes/G-DRIVE/IPTS/IPTS-25519-iMars3D-command-line/raw/ob/2021_07_21_1in/'
df_folder = '/Volumes/G-DRIVE/IPTS/IPTS-25519-iMars3D-command-line/raw/df/2021_07_19/'
ct_sig = "treated_1inch"
ct_scan_root = "/Volumes/G-DRIVE/IPTS/IPTS-25519-iMars3D-command-line/raw/ct_scans/"
ct_dir = "/Volumes/G-DRIVE/IPTS/IPTS-25519-iMars3D-command-line/raw/ct_scans/2021_07_21_1in/"
iptsdir = "/Volumes/G-DRIVE/IPTS/IPTS-25519-iMars3D-command-line/"
outdir = "/Volumes/G-DRIVE/IPTS/IPTS-25519-iMars3D-command-line/shared/processed_data/2021_07_21_1in"
instrument = "CG1D"
ob_dir = "/Volumes/G-DRIVE/IPTS/IPTS-25519-iMars3D-command-line/raw/ob/"
data_dir = "/Volumes/G-DRIVE/IPTS/IPTS-25519-iMars3D-command-line/raw/"
workdir = "/Volumes/G-DRIVE/IPTS/sandbox/work.imars3d/2021_07_21_1in"
df_files = glob.glob(os.path.join(df_folder, "*.tiff"))
scan = "2021_07_21_1in"
ipts = 25519
ob_files = glob.glob(os.path.join(ob_folder, "*.tiff"))
ct_subdir = "2021_07_21_1in"
df_dir = "/Volumes/G-DRIVE/IPTS/IPTS-25519-iMars3D-command-line/raw/df/"
facility = "HFIR"
ct_scans_subdir = glob.glob(os.path.dirname(data_folder) + "/*")
# # Create CT data object
from imars3d.CT import CT
ct = CT(data_dir,
CT_subdir=ct_dir,
CT_identifier=ct_sig,
workdir=workdir,
outdir=outdir,
ob_files=ob_files,
df_files=df_files)
# # preprocess
# %%time
ppd = ct.preprocess()
xmin = 500
ymin = 0
xmax = 1600
ymax = 2047
# %%time
ct.recon(crop_window=(xmin, ymin, xmax, ymax))
| notebooks/CT_reconstruction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pwd
# %cd ..
# +
import matplotlib.pyplot as plt
import pickle
import numpy as np
import tools
from pylab import *
import matplotlib.animation as animation
import matplotlib as mpl
import numpy as np
import os
import glob
import standard.analysis as sa
import tools
import matplotlib.pyplot as plt
import task
import tensorflow as tf
from model import FullModel
from dict_methods import *
# %matplotlib inline
# -
mpl.rcParams['font.size'] = 15
mpl.rcParams['pdf.fonttype'] = 42
mpl.rcParams['ps.fonttype'] = 42
mpl.rcParams['font.family'] = 'arial'
mpl.rcParams['image.cmap']='jet'
d = os.path.join(os.getcwd(), 'files')
files = glob.glob(os.path.join(d,'cluster_simple', 'cluster_simple50'))
print(len(files))
res = defaultdict(list)
for f in files:
temp = tools.load_all_results(f, argLast = False)
chain_defaultdicts(res, temp)
plt.figure(figsize=(8,8))
plt.plot(res['lin_bins'][0,:500],res['lin_hist'][:,-1,:500].T)
plt.ylim([0, 10000])
peak_inds = np.zeros_like(res['kc_prune_threshold']).astype(np.bool)
for i, thres in enumerate(res['kc_prune_threshold']):
x = np.where(res['lin_bins'][i,:-1] > res['kc_prune_threshold'][i])[0][0]
peak = np.argmax(res['lin_hist'][i, -1, x-1:])
if peak > 10:
peak_inds[i] = True
else:
peak_inds[i] = False
peak_inds
res['lr'][np.invert(peak_inds)]
plt.figure(figsize=(8,8))
plt.plot(res['lin_bins'][0,:500],res['lin_hist'][peak_inds,-1,:500].T)
plt.ylim([0, 2000])
ind = np.where(np.invert(peak_inds))[0]
ind
plt.figure(figsize=(8,8))
plt.plot(res['lin_bins'][0,:500],res['lin_hist'][ind,-1,:500].T)
plt.ylim([0, 10000])
acc_ind = res['train_acc'][:,-1] > .6
res['lr'][np.invert(acc_ind)]
badkc_ind = res['bad_KC'][:,-1] < .2
res['separate_lr'][np.invert(badkc_ind)]
ind = np.logical_and(badkc_ind, acc_ind, peak_inds)
res['separate_lr'][ind]
#filter
for k, v in res.items():
res[k] = v[peak_inds]
def _get_K(res):
n_model, n_epoch = res['sparsity'].shape[:2]
Ks = np.zeros((n_model, n_epoch))
bad_KC = np.zeros((n_model, n_epoch))
for i in range(n_model):
for j in range(n_epoch):
sparsity = res['sparsity'][i, j]
Ks[i, j] = sparsity[sparsity>0].mean()
bad_KC[i,j] = np.sum(sparsity==0)/sparsity.size
res['K'] = Ks
res['bad_KC'] = bad_KC
_get_K(res)
def simple_plot(xkey, ykey, filter_dict = None):
if filter_dict is not None:
temp = filter(res, filter_dict=filter_dict)
x = temp[xkey]
y = temp[ykey][:,-1]
plt.figure()
plt.plot(np.log(x), y, '*')
plt.xticks(np.log(x),x)
plt.xlabel(xkey)
plt.ylabel(ykey)
#if filter_dict is not None:
# plt.legend('{} = {}'.format(filter_dict.key[0],filter_dict.value[0]))
def marginal_plot(xkey, ykey, varykey, marginals=None):
plt.figure()
for i in np.unique(res[varykey]):
temp = filter(res, {varykey:i})
if marginals:
temp = filter(temp, marginals)
x = temp[xkey]
y = temp[ykey][:,-1]
plt.plot(np.log(x), y, '*')
x = np.unique(res[xkey])
plt.xticks(np.log(x),x)
plt.xlabel(xkey)
plt.ylabel(ykey)
plt.legend(np.unique(res[varykey]))
plt.title(marginals)
#marginal_plot('lr', 'coding_level', 'kc_prune_threshold')
marginal_plot('lr', 'K', 'kc_prune_threshold', {'N_KC':2500})
#marginal_plot('lr', 'train_acc', 'kc_prune_threshold')
#marginal_plot('lr', 'K', 'N_KC', {'kc_prune_threshold':0.08})
marginal_plot('lr', 'coding_level', 'separate_lr', {'N_KC':2500})
marginal_plot('lr', 'K', 'separate_lr', {'N_KC':2500})
marginal_plot('lr', 'train_acc', 'separate_lr', {'N_KC':2500})
marginal_plot('lr', 'coding_level', 'N_KC', {'kc_prune_threshold':0.08})
marginal_plot('lr', 'K', 'N_KC', {'kc_prune_threshold':0.08})
marginal_plot('lr', 'train_acc', 'N_KC', {'kc_prune_threshold':0.08})
# +
x = filter(res, {'kc_prune_threshold':0.08, 'N_KC':2500})
plt.plot(x['K'].T)
plt.legend(np.unique(x['N_KC']))
plt.ylim([0, 11])
plt.figure(figsize=(8,8))
plt.plot(x['lin_bins'][0,:-1],x['lin_hist'][:,-1].T)
plt.ylim([0, 1000])
plt.legend(x['lr'])
# +
x = filter(x, {'N_KC':2500, 'kc_prune_threshold':0.08})
plt.plot(x['log_val_loss'].T)
plt.legend(np.unique(x['separate_lr']))
plt.legend(x['lr'])
plt.figure(figsize=(8,8))
plt.plot(x['lin_bins'][0,:500],x['lin_hist'][:,-1,:500].T)
plt.ylim([0, 2000])
plt.legend(x['lr'])
# -
| notebooks/analyze_K_big.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import os
from argparse import ArgumentParser
from sklearn.externals import joblib
from tictacs import from_recipe
from pan import ProfilingDataset
import dill
import cPickle as pickle
# from sklearn.neighbors import KNeighborsClassifier
# from sklearn.metrics import accuracy_score, confusion_matrix
infolder = "../DATA/pan16-author-profiling-training-dataset-2016-04-25/pan16-author-profiling-training-dataset-english-2016-02-29/"
outfolder = "models/"
print('Loading dataset->Grouping User texts.\n')
dataset = ProfilingDataset(infolder)
print('Loaded {} users...\n'.format(len(dataset.entries)))
# get config
config = dataset.config
tasks = config.tasks
print('\n--------------- Thy time of Running ---------------')
for task in tasks:
print('Learning to judge %s..' % task)
# load data
X, y = dataset.get_data(task)
# -
Instance
# +
from pan import ProfilingDataset, createDocProfiles, create_target_prof_trainset
from pan import preprocess
task = 'gender'
docs = createDocProfiles(dataset)
X, y = create_target_prof_trainset(docs, task)
print len(X)
#X = preprocess.preprocess(X)
# -
Profile
#reload(preprocess)
#reload(features)
from pan import features
from pan import preprocess
X, y = dataset.get_data('age')
#X, y = dataset.get_data('gender')
print len(X)
#print X[0]
#X = preprocess.preprocess(X)
#print "$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$$"
#print X[0]
from nltk.internals import find_binary
_megam_bin = None
def config_megam(bin=None):
"""
Configure NLTK's interface to the ``megam`` maxent optimization
package.
:param bin: The full path to the ``megam`` binary. If not specified,
then nltk will search the system for a ``megam`` binary; and if
one is not found, it will raise a ``LookupError`` exception.
:type bin: str
"""
global _megam_bin
_megam_bin = find_binary(
'megam', bin,
env_vars=['MEGAM'],
binary_names=['megam.opt', 'megam', 'megam_686', 'megam_i686.opt'],
url='http://www.umiacs.umd.edu/~hal/megam/index.html')
from nltk import MaxentClassifier
from sklearn.pipeline import Pipeline, FeatureUnion
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.svm import SVC
from pan.features import SOA_Model2
soac = features.SOAC_Model2(max_df=1.0, min_df=1, tokenizer_var='sklearn', max_features=None)
svm = SVC(kernel='rbf', C=0.1, gamma=1, class_weight='balanced', probability=False)
#combined = FeatureUnion([('count_tokens', countTokens), ('count_hash', countHash),
# ('count_urls', countUrls), ('count_replies', countReplies),
# ('soa', soa), ('soac', soac)])+
#combined = FeatureUnion([('count_tokens', countTokens), ('count_hash', countHash),
# ('count_urls', countUrls), ('count_replies', countReplies)])
pipe1 = Pipeline([('soac',soac), ('svm', svm)])
pipe1.steps
from sklearn.cross_validation import train_test_split
from sklearn.metrics import accuracy_score, confusion_matrix,precision_recall_fscore_support, classification_report
from sklearn.cross_validation import train_test_split
from sklearn.grid_search import GridSearchCV
num_folds = 4
split = 0.25
X_train, X_cv, y_train, y_cv = train_test_split(X, y, test_size=split, stratify=y)
for i, x in enumerate(X_train):
if len(x)==0:
X_train.remove(x)
y_train.remove(y_train[i])
for i, x in enumerate(X_cv):
if len(x)==0:
X_cv.remove(x)
y_cv.remove(y_cv[i])
print len(X_train), len(X_cv), len(X_cv) + len(X_train), len(X)
X_meta, X_cv, y_meta, y_cv = train_test_split(X_cv, y_cv, test_size=0.5, stratify=y_cv)
print len(X_train), len(X_cv), len(X_meta), len(X_cv) + len(X_train) + len(X_meta), len(X)
params = {'svm__C': [0.001, 0.01, 0.1, 1, 10, 100]}
params = {}
grid_cv = GridSearchCV(pipe1, param_grid=params, verbose=1, n_jobs=-1, cv=num_folds, refit=True)
grid_cv.fit(X_train, y_train)
#grid_cv.fit(X_train, y_train)
print(grid_cv.best_score_)
print(grid_cv.best_params_)
pipe1 = grid_cv.best_estimator_
# +
from nltk.corpus import names
from nltk import MaxentClassifier
from nltk import classify
import random
names = ([(name, 'male') for name in names.words('male.txt')] + [(name, 'female') for name in names.words('female.txt')])
random.shuffle(names)
def gender_features3(name):
features = {}
features["fl"] = name[0].lower()
features["ll"] = name[-1].lower()
features["fw"] = name[:2].lower()
features["lw"] = name[-2:].lower()
return features
featuresets = [(gender_features3(n), g) for (n, g) in names]
train_set, test_set = featuresets[500:], featuresets[:500]
me3_megam_classifier = MaxentClassifier.train(train_set, "megam")
classify.accuracy(me3_megam_classifier, test_set)
# +
import regex as re
import nltk
import numpy
from textblob.tokenizers import WordTokenizer
from sklearn.base import BaseEstimator, TransformerMixin
from pan.misc import _twokenize
def tokenization2(text):
import re
emoticons_str = r"""
(?:
[:=;] # Eyes
[oO\-]? # Nose (optional)
[D\)\]\(\]/\\OpP] # Mouth
)"""
regex_str = [
emoticons_str,
#r'<[^>]+>', # HTML tags
#r'(?:@[\w_]+)', # @-mentions
#r"(?:\#+[\w_]+[\w\'_\-]*[\w_]+)", # hash-tags
#r'http[s]?://(?:[a-z]|[0-9]|[$-_@.&+]|[!*\(\),]|(?:%[0-9a-f][0-9a-f]))+', # URLs
#r'(?:(?:\d+,?)+(?:\.?\d+)?)', # numbers
#r"(?:[a-z][a-z'\-_]+[a-z])", # words with - and '
#r'(?:[\w_]+)', # other words
#r'(?:\S)' # anything else
]
tokens_re = re.compile(r'('+'|'.join(regex_str)+')', re.VERBOSE | re.IGNORECASE)
emoticon_re = re.compile(r'^'+emoticons_str+'$', re.VERBOSE | re.IGNORECASE)
return [token if emoticon_re.search(token) else token.lower() for token in tokens_re.findall(text)]
class CountHash(BaseEstimator, TransformerMixin):
""" Model that extracts a counter of twitter-style hashes. """
pat = re.compile(r'(?<=\s+|^)#\w+', re.UNICODE)
def fit(self, X, y=None):
return self
def transform(self, texts):
""" transform data
:texts: The texts to count hashes in
:returns: list of counts for each text
"""
return [[len(CountHash.pat.findall(text)) / float(len(text))] for text in texts]
class CountReplies(BaseEstimator, TransformerMixin):
""" Model that extracts a counter of twitter-style @replies. """
pat = re.compile(r'(?<=\s+|^)@\w+', re.UNICODE)
def fit(self, X, y=None):
return self
def transform(self, texts):
""" transform data
:texts: The texts to count replies in
:returns: list of counts for each text
"""
return [[len(CountReplies.pat.findall(text)) / float(len(text))] for text in texts]
class CountURLs(BaseEstimator, TransformerMixin):
""" Model that extracts a counter of URL links from text. """
pat = re.compile(r'((https?|ftp)://[^\s/$.?#].[^\s]*)')
def fit(self, X, y=None):
return self
def transform(self, texts):
""" transform data
:texts: The texts to count URLs in
:returns: list of counts for each text
"""
return [[len(CountURLs.pat.findall(text)) / float(len(text))] for text in texts]
class CountCaps(BaseEstimator, TransformerMixin):
""" Model that extracts a counter of capital letters from text. """
def fit(self, X, y=None):
return self
def transform(self, texts):
""" transform data
:texts: The texts to count capital letters in
:returns: list of counts for each text
"""
return [[sum(c.isupper() for c in text)] for text in texts]
class CountWordCaps(BaseEstimator, TransformerMixin):
""" Model that extracts a counter of capital words from text. """
def fit(self, X, y=None):
return self
def transform(self, texts):
""" transform data
:texts: The texts to count capital words in
:returns: list of counts for each text
"""
return [[sum(w.isupper() for w in nltk.word_tokenize(text))]
for text in texts]
class CountWordLength(BaseEstimator, TransformerMixin):
""" Model that extracts a counter of word length from text. """
def __init__(self, span):
""" Initialize this feature extractor
:span: tuple - range of lengths to count
"""
self.span = span
def fit(self, X, y=None):
return self
def transform(self, texts):
""" transform data
:texts: The texts to count word lengths in
:returns: list of counts for each text
"""
mini = self.span[0]
maxi = self.span[1]
num_counts = maxi - mini
# wt = WordTokenizer()
tokens = [tokenization(text) for text in texts]
text_len_dist = []
for line_tokens in tokens:
counter = [0] * num_counts
for word in line_tokens:
word_len = len(word)
if mini <= word_len <= maxi:
counter[word_len - 1] += 1
text_len_dist.append([each for each in counter])
return text_len_dist
class CountTokens(BaseEstimator, TransformerMixin):
""" Model that extracts a counter of capital words from text. """
def __init__(self):
self.l = ['a', 'b', 'c', 'd', 'e', 'f', 'g', 'h', 'i', 'j',
'k', 'l', 'm', 'n', 'o', 'p', 'q', 'r', 's', 't',
'u', 'v', 'w', 'x', 'y', 'z', '0', '1', '2', '3',
'4', '5', '6', '7', '8', '9', '!', '.', ':', '?',#]
';', ',', ')', '(', '-', '%', '$', '#', '@', '^',
'&', '*', '=', '+', '/', '"', "'", '<', '>', '|',
'~', '`']
def fit(self, X, y=None):
return self
def transform(self, texts):
""" transform data
:texts: The texts to count capital words in
:returns: list of counts for each text
"""
l = self.l
return [[text.lower().count(token)/float(len(text)) for token in l]
for text in texts]
# +
import re
POSITIVE = ["*O", "*-*", "*O*", "*o*", "* *",
":P", ":D", ":d", ":p",
";P", ";D", ";d", ";p",
":-)", ";-)", ":=)", ";=)",
":<)", ":>)", ";>)", ";=)",
"=}", ":)", "(:;)",
"(;", ":}", "{:", ";}",
"{;:]",
"[;", ":')", ";')", ":-3",
"{;", ":]",
";-3", ":-x", ";-x", ":-X",
";-X", ":-}", ";-=}", ":-]",
";-]", ":-.)",
"^_^", "^-^"]
NEGATIVE = [":(", ";(", ":'(",
"=(", "={", "):", ");",
")':", ")';", ")=", "}=",
";-{{", ";-{", ":-{{", ":-{",
":-(", ";-(",
":,)", ":'{",
"[:", ";]"
]
class EmoticonsCounter(object):
""" Model that extracts a counter emoticons """
def __init__(self, POSITIVE, NEGATIVE):
self.l = POSITIVE + NEGATIVE
def fit(self, X, y=None):
return self
def transform(self, texts):
""" transform data
:texts: The texts to count capital words in
:returns: list of counts for each text
"""
l = self.l
#print texts
return [[text.count(emo) for emo in l]
for text in texts]
# -
pt = ParseTweet(X[3])
pt.getAttributeEmoticon(X[3])
# +
from sklearn.feature_extraction.text import TfidfVectorizer
tf = TfidfVectorizer(min_df=5, max_df=0.9, max_features=100, tokenizer=_twokenize.tokenizeRawTweetText)
# -
a = CountTokens()
a = EmoticonsCounter(POSITIVE, NEGATIVE)
dd = numpy.array(a.transform(X))
dd.shape
import pandas as pd
data = pd.DataFrame(dd, columns=a.l)
data["class"] = y
print(data.describe())
# +
grouped = data.groupby('class')
means = grouped.mean().T
import matplotlib.pyplot as plt
### BAR PLOTS OF MEAN VALUE OF FEATURES FOR EACH CLASS ######
grouped = data.groupby('class')
plt.figure()
grouped.mean().T.plot(kind='barh', figsize=(60,10))
plt.savefig('test1.png')
plt.show()
# -
def feat_to_fi(feat_vector, names):
features = {}
#print feat_vector
#print len(feat_vector)
for i in xrange(len(feat_vector)):
features[names[i]] = feat_vector[i]
return features
#trained_soac = pipe1.steps[0][1]
#Transf = trained_soac.transform(X_train + X_cv)
Transf = a.transform(X_train+X_meta +X_cv)
#Transf = tf.fit_transform(X_train + X_cv).todense()
#from sklearn.preprocessing import normalize
#Transf = normalize(Transf, axis=1, norm='l1')
Transf = numpy.array(Transf)
print Transf[0,:]
y_all = y_train + y_meta + y_cv
featuresets = ([(feat_to_fi(Transf[i,:], a.l), y_all[i]) for i in xrange(Transf.shape[0]) ])
train_set, test_set = featuresets[:len(X_train)], featuresets[len(X_train):]
meta_set, test_set = test_set[:len(X_meta)], test_set[len(X_meta):]
#mec = MaxentClassifier.train(train_set, "megam")
#mec = nltk.classify.MaxentClassifier.train(train_set, 'GIS', trace=0, max_iter=1000)
tf.get_feature_names()
scores = []
gauss = []
for i in xrange(1000):
gauss.append(random.random()*random.randint(0,10))
mec = MaxentClassifier.train(train_set, "megam", gaussian_prior_sigma=gauss[-1])
scores.append(classify.accuracy(mec, meta_set))
print "Best prior: %0.3f with score: %0.2f " % (gauss[scores.index(max(scores))],max(scores))
#mec = nltk.classify.MaxentClassifier.train(train_set, 'GIS', trace=0, max_iter=1000)
#classify.accuracy(mec, test_set)
mec = MaxentClassifier.train(train_set, "megam", gaussian_prior_sigma=gauss[scores.index(max(scores))])
mec.show_most_informative_features(n=100)
mec.explain(train_set[0][0])
mec.classify_many([test_sample[0] for test_sample in test_set])
len(y_cv), len(meta_set)
from sklearn.metrics import classification_report, accuracy_score, confusion_matrix
#predict = model.predict(X_cv)
predict = mec.classify_many([test_sample[0] for test_sample in test_set])
acc = accuracy_score(y_cv, predict)
conf = confusion_matrix(y_cv, predict, labels=sorted(list(set(y))))
rep = classification_report(y_cv, predict, target_names=sorted(list(set(y))))
print('Accuracy : {}'.format(acc))
print('Confusion matrix :\n {}'.format(conf))
print('Classification report :\n {}'.format(rep))
| Entropy_Features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using Jupyter notebook for interactive development
#
# url: http://jupyter.org/
#
# 
# # How to run this notebook?
#
# Click **Cell** and then **Run All**
#
# # How to run it online?
# [mybinder.org/repo/hainm/notebook-pytraj](http://mybinder.org/repo/hainm/notebook-pytraj)
#
# # See also
# [protein viewer example](./3pqr.ipynb)
# ## Install
#
# ```bash
# conda install parmed -c ambermd # Python package for topology editing and force field development
# conda install pytraj-dev -c ambermd # Python interface for cpptraj (MD trajectory data analysis)
# conda install pysander -c ambermd # Python interface for SANDER
# # all above will be also available in AMBER16 release (next few months)
#
# conda install nglview -c ambermd # Protein/DNA/RAN viewer in notebook
#
# # notebook
# conda install jupyter notebook
#
# ```
# ## ParmEd: Cross-program parameter and topology file editor and molecular mechanical simulator engine.
#
# url: https://github.com/ParmEd/ParmEd (AMBER16)
# +
import warnings
warnings.filterwarnings("ignore", category=DeprecationWarning)
warnings.filterwarnings("ignore", category=UserWarning)
import parmed as pmd
x = pmd.load_file('tz2.pdb')
[res.name for res in x.residues]
# -
[atom.name for atom in x.residues[0]]
# ## PYTRAJ: Interactive data analysis for molecular dynamics simulations
#
# url: https://github.com/Amber-MD/pytraj (AMBER 16)
# ### Compute distances and plot
# +
import pytraj as pt
traj = pt.load('tz2.nc', 'tz2.parm7')
distances = pt.distances(traj, ':1 :12', dtype='dataframe')
distances.head()
# +
# %matplotlib inline
distances.hist()
# -
# ### Compute multiple dihedrals
dihedrals = pt.multidihedral(traj, resrange='1-3', dtype='dataframe')
dihedrals.head(3) # show only first 3 snapshots
# +
# %matplotlib inline
from matplotlib import pyplot as plt
plt.plot(dihedrals['phi_2'], dihedrals['psi_2'], '-bo', linewidth=0)
plt.xlim([-180, 180])
plt.ylim([-180, 180])
# -
# ### get help?
help(pt.multidihedral)
# ## Protein/DNA/RNA viewer in notebook
#
# - Written in Python/Javascript
# - super light (~3 MB)
# - super easy to install (pip install nglview)
# +
import warnings
warnings.filterwarnings('ignore')
import nglview as nv
view = nv.show_pytraj(traj)
view
# -
view.representations = []
view.add_representation('cartoon', color='residueindex')
view.add_representation('licorice')
t0 = pt.fetch_pdb('3pqr')
view0 = pt.view.to_nglview(t0)
view0
view0.representations = []
view0.add_representation('cartoon', selection='protein', color='residueindex')
view0.add_representation('surface', selection='protein', opacity='0.2')
# ### Not yet ported to Python
#
# 
# ## Phenix wishlist (for developers)
#
# - phenix.conda install jupyter notebook
# - phenix.ipython
| phenix2016.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.5 64-bit (''.env'': virtualenv)'
# name: python37564bitenvvirtualenv0dc96e775ff048369257ba6f47949af5
# ---
# + id="qENuCNYVEveh" colab_type="code" colab={}
from google.colab import drive
drive.mount('/content/drive')
# + id="anO2d1D1oF1C" colab_type="code" colab={}
# !pip install rasterio
# !pip install nvector
# + id="OXPNii7Xno1e" colab_type="code" colab={}
# See https://stackoverflow.com/questions/55821982/is-there-a-way-to-import-future-in-colaboratory
# from __future__ import annotations
import numpy as np
import rasterio
import nvector as nv
import matplotlib.pyplot as plt
import json
from typing import Tuple, List, Dict
from tqdm import tqdm
from numba import jit, cuda
# %matplotlib inline
# + [markdown] id="3jhEpVC2no1i" colab_type="text"
# # Calculate tops of the world in Google Colab
# + id="3PP0S_I3urzm" colab_type="code" colab={}
class LatLongElev:
def __init__(self, latitude: float, longitude: float, elevation: float=0.0):
self.latitude = latitude
self.longitude = longitude
self.elevation = elevation
class XYZ:
def __init__(self, x: float, y: float, z: float):
self.x = x
self.y = y
self.z = z
def __eq__(self, xyz) -> bool:
return self.x == xyz.x and self.y == xyz.y and self.z == xyz.z
def __str__(self) -> str:
return "x={}, y={}, z={}".format(self.x, self.y, self.z)
def __hash__(self) -> int:
return hash((self.x, self.y, self.z))
def project_onto_line(self, _a, _b):
# See https://gamedev.stackexchange.com/questions/72528/how-can-i-project-a-3d-point-onto-a-3d-line
p = self.to_np_array()
a = _a.to_np_array()
b = _b.to_np_array()
ap = p-a
ab = b-a
result = a + np.dot(ap, ab)/np.dot(ab, ab) * ab
return XYZ(result[0], result[1], result[2])
def get_chunk_index(self, chunk_size: float):
x = self.x - self.x % chunk_size
y = self.y - self.y % chunk_size
z = self.z - self.z % chunk_size
return XYZ(x, y, z)
def to_np_array(self) -> np.array:
return np.array([self.x, self.y, self.z])
# + id="7P3a1vk0no1j" colab_type="code" colab={}
def plotGlobe(xyz_list: List[XYZ]):
x = [xyz.x for xyz in xyz_list]
y = [xyz.y for xyz in xyz_list]
z = [xyz.z for xyz in xyz_list]
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.scatter(x, y, z)
plt.show()
DEEPEST_DEPRESSION_ON_EARTH = -418.0
def get_latlongelev_list_from_tif_image(image_file: str, step: int=1) -> List[LatLongElev]:
latlongelev_list = []
with rasterio.open(image_file) as image:
elevation_values = image.read(1)
for y in range(0, image.height, step):
for x in range(0, image.width, step):
longitude, latitude = image.xy(y, x)
elevation = elevation_values[y, x]
# Clean under the sea elevations
elevation = elevation if elevation > DEEPEST_DEPRESSION_ON_EARTH else DEEPEST_DEPRESSION_ON_EARTH
latlongelev_list.append(LatLongElev(latitude, longitude, elevation))
return latlongelev_list
@jit(forceobj=True)
def latlongelev_list_to_xyz_list(latlongelev_list: List[LatLongElev]) -> List[XYZ]:
# See https://github.com/pbrod/nvector#example-4-geodetic-latitude-to-ecef-vector
wgs84 = nv.FrameE(name='WGS84')
xyz_list = []
for latlongelev in latlongelev_list:
depth = latlongelev.elevation * (-1)
pointB = wgs84.GeoPoint(latitude=latlongelev.latitude, longitude=latlongelev.longitude, z=depth, degrees=True)
p_EB_E = pointB.to_ecef_vector()
x, y, z = p_EB_E.pvector.ravel()[0], p_EB_E.pvector.ravel()[1], p_EB_E.pvector.ravel()[2]
xyz_list.append(XYZ(x, y, z))
return xyz_list
@jit(forceobj=True)
def filter_only_tops(xyz_list: List[XYZ]) -> List[XYZ]:
progress_bar = tqdm(total=len(xyz_list))
top_list = []
for xyz in xyz_list:
top = get_top_for_direction(xyz, xyz_list)
# Store only unique tops
if top not in top_list:
top_list.append(top)
progress_bar.update(1)
progress_bar.close()
return top_list
CENTER_OF_THE_EARTH = XYZ(0, 0, 0)
@jit(forceobj=True)
def get_top_for_direction(direction: XYZ, xyz_list) -> XYZ:
xyz_projection_list = []
for xyz in xyz_list:
xyz_projection = xyz.project_onto_line(CENTER_OF_THE_EARTH, direction)
if are_on_the_same_side_relative_to_center(xyz_projection, direction):
xyz_projection_list.append((xyz, xyz_projection))
# Lambda cannot be passed as an argument while using JIT
# See https://github.com/numba/numba/issues/4481#issuecomment-524914268
top = max(xyz_projection_list, key=_distance_wrapper)[0]
return top
def are_on_the_same_side_relative_to_center(a: XYZ, b: XYZ) -> bool:
signs = np.sign([a.x, b.x, a.y, b.y, a.z, b.z])
for axis in range(0, 6, 2):
if signs[0+axis] != signs[1+axis]:
return False
return True
def _distance_wrapper(xyz_tuple: Tuple[XYZ, XYZ]) -> float:
return _distance(xyz_tuple[1])
def _distance(xyz: XYZ) -> float:
# The square root function is monotonic, so it can be discarded
return xyz.x**2 + xyz.y**2 + xyz.z**2
CHUNK_SIZE = 200000
def save_results_to_local_file(tops: List[XYZ], output_file: str):
tops_dto = convert_tops_to_dto(tops, CHUNK_SIZE)
with open(output_file, 'w') as file:
json.dump(tops_dto, file)
def convert_tops_to_dto(tops: List[XYZ], chunk_size: int) -> List[Dict]:
chunks = {}
for top in tops:
index = top.get_chunk_index(chunk_size)
if chunks.get(index):
chunks[index]["tops"].append(top)
else:
chunks[index] = {"index": index, "tops": [top]}
dto = [{
"index": vars(chunk["index"]),
"tops": [vars(top) for top in chunk["tops"]]
} for _, chunk in chunks.items()]
return dto
# + id="Svz50yHLpdrt" colab_type="code" colab={}
latlongelev_list = get_latlongelev_list_from_tif_image(
'/content/drive/My Drive/Colab Notebooks/calculate_tops/srtm_40_02.tif',
step=200)
xyz_list = latlongelev_list_to_xyz_list(latlongelev_list)
# + id="HYuaIXrMqkhl" colab_type="code" outputId="34e482e2-48ed-4c82-e0a7-c448ec22c2b0" colab={"base_uri": "https://localhost:8080/", "height": 248}
plotGlobe(xyz_list)
# + id="e4M5Ibcvqw86" colab_type="code" outputId="b6cdba94-a836-4940-9c51-ca295b63e971" colab={"base_uri": "https://localhost:8080/", "height": 34}
tops = filter_only_tops(xyz_list)
# + id="g9K1ZfYdq7Ll" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 248} outputId="6d0b52e7-fff9-49f2-84ed-920a97678938"
plotGlobe(tops)
# + id="HOd2oUcIo0Tm" colab_type="code" colab={}
save_results_to_local_file(tops, '/content/drive/My Drive/Colab Notebooks/calculate_tops/result.json')
# + id="kojvvP1Hve4K" colab_type="code" colab={}
| data_mining/calculate_tops_in_colab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generalized Poisson Distribution
# The **Poisson** distribution models count data where the mean is equal to the variance.
#
# The **Negative Binomial** distribution allows us to model overdispersed count data. 2 parameters:
# - $\mu > 0$ mean
# - $\alpha > 0$ overdispersion parameter: as $\alpha \rightarrow \infty$, Neg Bin converges to Poisson
#
# We want our model to be able to fit underdispersed count data so that we can get tighter interval estimates. The **Generalized Poisson** distribution is flexible enough to handle both overdispersion and underdispersion. It has the following PMF:
#
# $$f(y | \theta, \lambda) = \frac{\theta (\theta + \lambda y)^{y-1} e^{-\theta - \lambda y}}{y!}, y = 0,1,2,...$$
# where $\theta > 0$ and $\max(-1, -\frac{\theta}{4}) \leq \lambda \leq 1$
#
# - When $\lambda = 0$, the Gen Poisson reduces to the standard Poisson with $\mu = \theta$
# - When $\lambda < 0$, the model has underdispersion
# - When $\lambda > 0$, the model has overdispersion
#
# This notebook gives a sketch of the code to implement this distribution in PyMC3. The model is the same as what I show in the AR model notebook, but with the Generalized Poisson in place of the Poisson.
# ## References
# General info on the generalized poisson:
# - https://www.tandfonline.com/doi/pdf/10.1080/03610929208830766
# - https://journals.sagepub.com/doi/pdf/10.1177/1536867X1201200412
# - https://towardsdatascience.com/generalized-poisson-regression-for-real-world-datasets-d1ff32607d79
#
# Random sampling algorithms:
# - https://www.tandfonline.com/doi/abs/10.1080/01966324.1997.10737439?journalCode=umms20
#
# Creating a custom distribution in PyMC3:
# - https://discourse.pymc.io/t/examples-of-random-in-custom-distribution/1263
# - https://docs.pymc.io/notebooks/getting_started.html#Arbitrary-deterministics
# %matplotlib inline
import pymc3 as pm
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import theano.tensor as tt
from pymc3.distributions.dist_math import bound, logpow, factln
from pymc3.distributions.distribution import draw_values, generate_samples
# ## Log Likelihood Function
#
# $$\log f(y | \theta, \lambda) = \log\theta + \log\left((\theta + \lambda y)^{y-1}\right) - (\theta + \lambda y) - \log(y!),\,y = 0,1,2,...$$
# where $\theta > 0$ and $\max(-1, -\frac{\theta}{4}) \leq \lambda \leq 1$
def genpoisson_logp(theta, lam, value):
log_prob = bound(np.log(theta) + logpow(theta + lam * value, value - 1)
- (theta + lam * value) - factln(value),
theta >= 0,
-1 <= lam, -theta/4 <= lam, lam <= 1,
value >= 0)
# Return zero when value > m, where m is the largest pos int for which theta + m * lam > 0 (when lam < 0)
return tt.switch(theta + value * lam <= 0,
0, log_prob)
# ## Generator Function
# The function defined below is meant to be analogous to `scipy.stats.<DIST_NAME>.rvs` which generates random samples from the distribution with the given parameters.
#
# ### Inversion Algorithm
# (presented in <NAME> (1997) Generalized Poisson Random Variate Generation)
#
# Initialize $\omega \leftarrow e^{-\lambda}$
# 1. $X \leftarrow 0$
# 2. $S \leftarrow e^{-\theta}$ and $P \leftarrow S$
# 3. Generate $U$ from uniform distribution on $(0,1)$.
# 4. While $U > S$, do
# 1. $X \leftarrow X + 1$
# 2. $C \leftarrow \theta - \lambda + \lambda X$
# 3. $P \leftarrow \omega \cdot C (1 + \frac{\lambda}{C})^{X-1} P X^{-1}$
# 4. $S \leftarrow S + P$
# 5. Deliver $X$
def genpoisson_rvs(theta, lam, size=None):
if size is not None:
assert size == theta.shape
else:
size = theta.shape
lam = lam[0]
omega = np.exp(-lam)
X = np.full(size, 0)
S = np.exp(-theta)
P = np.copy(S)
for i in range(size[0]):
U = np.random.uniform()
while U > S[i]:
X[i] += 1
C = theta[i] - lam + lam * X[i]
P[i] = omega * C * (1 + lam/C)**(X[i]-1) * P[i] / X[i]
S[i] += P[i]
return X
# ## Generalized Poisson Distribution class definition
class GenPoisson(pm.Discrete):
def __init__(self, theta, lam, *args, **kwargs):
super(GenPoisson, self).__init__(*args, **kwargs)
self.theta = theta
self.lam = lam
def logp(self, value):
theta = self.theta
lam = self.lam
return genpoisson_logp(theta, lam, value)
def random(self, point=None, size=None):
theta, lam = draw_values([self.theta, self.lam], point=point, size=size)
return generate_samples(genpoisson_rvs,
theta=theta, lam=lam,
size=size)
# ## Using the Generalized Poisson likelihood in our GAR model
#
# Suppose that $y$ is GenPoisson distributed over the exponential of a latent time series modeled by an autoregressive process with $W$ lags, i.e.,
#
# $$y_t \sim \text{GenPoisson}( \theta = \exp(f_t), \lambda )$$
#
# where for each $t$, $f_t$ is a linear combination of the past $W$ timesteps, i.e.,
#
# $$f_t \sim N(\beta_0 + \beta_1 * f_{t-1} + ... + \beta_W * f_{t-W}, \tau^2)$$
#
# ### Priors
# Bias weight $$\beta_0 \sim N(0,0.1)$$
# Weight on most recent timestep $$\beta_1 \sim N(1,0.1)$$
# Weights on all other previous timesteps $$\beta_2, ..., \beta_W \sim N(0,0.1)$$
# Standard deviation $$\tau \sim \text{HalfNormal}(0.1)$$
# Dispersion parameter $$\lambda \sim \text{TruncatedNormal}(0, 0.1, \text{lower}=-1, \text{upper}=1)$$
df = pd.read_csv('../mass_dot_gov_datasets/boston_medical_center_2020-04-29_to_2020-07-06.csv')
y = df['hospitalized_total_covid_patients_suspected_and_confirmed_including_icu'].astype(float)
T = len(y)
W = 2 # window size
F = 7 # num days of forecasts
with pm.Model() as model:
bias = pm.Normal('beta[0]', mu=0, sigma=0.1)
beta_recent = pm.Normal('beta[1]', mu=1, sigma=0.1)
rho = [bias, beta_recent]
for i in range(2, W+1):
beta = pm.Normal(f'beta[{i}]', mu=0, sigma=0.1)
rho.append(beta)
tau = pm.HalfNormal('tau', sigma=0.1)
f = pm.AR('f', rho, sigma=tau, constant=True, shape=T+F)
lam = pm.TruncatedNormal('lam', mu=0, sigma=0.1, lower=-1, upper=1)
y_past = GenPoisson('y_past', theta=tt.exp(f[:T]), lam=lam, observed=y)
y_logp = pm.Deterministic('y_logp', y_past.logpt)
with model:
trace = pm.sample(5000, tune=1000, target_accept=0.99, max_treedepth=15,
chains=2, cores=1, init='adapt_diag', random_seed=42)
pm.traceplot(trace);
summary = pm.summary(trace)['mean'].to_dict()
for i in range(W+1):
print(f'beta[{i}]', summary[f'beta[{i}]'])
print('lambda', summary['lam'])
print('\nTraining score:')
print('Chain 1:', np.log(np.mean(np.exp(trace.get_values('y_logp', chains=0)))) / T)
print('Chain 2:', np.log(np.mean(np.exp(trace.get_values('y_logp', chains=1)))) / T)
with model:
y_future = GenPoisson('y_future', theta=tt.exp(f[-F:]), lam=lam, shape=F, testval=1)
forecasts = pm.sample_posterior_predictive(trace, vars=[y_future], random_seed=42)
samples = forecasts['y_future']
# +
low = np.zeros(F)
high = np.zeros(F)
mean = np.zeros(F)
median = np.zeros(F)
for i in range(F):
low[i] = np.percentile(samples[:,i], 2.5)
high[i] = np.percentile(samples[:,i], 97.5)
median[i] = np.percentile(samples[:,i], 50)
mean[i] = np.mean(samples[:,i])
plt.figure(figsize=(8,6))
x_future = np.arange(1,F+1)
plt.errorbar(x_future, median,
yerr=[median-low, high-median],
capsize=2, fmt='.', linewidth=1,
label='2.5, 50, 97.5 percentiles');
plt.plot(x_future, mean, 'x', label='mean');
x_past = np.arange(-4,1)
plt.plot(x_past, y[-5:], 's', label='observed')
plt.legend();
plt.title('Forecasts');
plt.xlabel('Days ahead');
# -
| notebooks/generalized_poisson.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <font size="+5">#01. Data Tables, Plots & Basic Concepts of Programming</font>
# <ul>
# <li>Doubts? → Ask me in <img src="https://discord.com/assets/f9bb9c4af2b9c32a2c5ee0014661546d.png" style="height: 1em; vertical-align: middle;"> <a href="https://discord.gg/cmB3KGsqMy">Discord</a></li>
# <li>Tutorials → <img src="https://openmoji.org/php/download_asset.php?type=emoji&emoji_hexcode=E044&emoji_variant=color" style="height: 1em; vertical-align: middle;"> <a href="https://www.youtube.com/channel/UCovCte2I3loteQE_kRsfQcw">YouTube</a></li>
# <li>Book Private Lessons → <span style="color: orange">@</span> <a href="https://sotastica.com/reservar">sotastica</a></li>
# </ul>
# # Define a varaible
# > Asign an `object` (numbers, text) to a `variable`.
pepa = 652417896
pepa
pepa/39
pepa+3
nombre = alfonso
# si no esta definido? en algun lugar deberia estarlo?
#
# donde?
#
# - el registro (environment)
pepa
hola
adios
34
3.34
# # The Registry (_aka The Environment_)
# > Place where Python goes to **recognise what we type**.
hola
"hola"
nombre = "alfonso"
alfonso
nombre
type(34)
type(34.3)
type('alfonso')
type([])
type({})
type
# # Use of Functions
# ## Predefined Functions in Python (_Built-in_ Functions)
# > https://docs.python.org/3/library/functions.html
len('ajwoifej wjf oie')
len('hola')
# ## Discipline to Search Solutions in Google
# > Apply the following steps when **looking for solutions in Google**:
# >
# > 1. **Necesity**: How to load an Excel in Python?
# > 2. **Search in Google**: by keywords
# > - `load excel python`
# > - ~~how to load excel in python~~
# > 3. **Solution**: What's the `function()` that loads an Excel in Python?
# > - A Function to Programming is what the Atom to Phisics.
# > - Every time you want to do something in programming
# > - **You will need a `function()`** to make it
# > - Theferore, you must **detect parenthesis `()`**
# > - Out of all the words that you see in a website
# > - Because they indicate the presence of a `function()`.
# ## External Functions
# > Download [this Excel](https://github.com/sotastica/data/raw/main/internet_usage_spain.xlsx).
# > Apply the above discipline and make it happen 🚀
# > I want to see the table, c'mon 👇
read_excel()
import pandas
read_excel()
pandas.factorize
pandas.fly()
pandas.read_excel
pandas.read_excel()
pandas.read_excel(io=internet_usage_spain.xlsx)
archivo = 'internet_usage_spain.xlsx'
archivo
pandas.read_excel(io=archivo)
pandas.read_excel(io='internet_usage_spain.xlsx')
import pandas
internet_usage_spain = 'internet_usage_spain'
pandas.read_excel(io=internet_usage_spain, sheet_name=1)
import pandas
pandas.read_excel(io='internet_usage_spain.xlsx', sheet_name=1)
# ## Discipline to Search Solutions in Google
# > Apply the following steps when **looking for solutions in Google**:
# >
# > 1. **Necesity**: How to load an Excel in Python?
# > 2. **Search in Google**: by keywords
# > - `load excel python`
# > - ~~how to load excel in python~~
# > 3. **Solution**: What's the `function()` that loads an Excel in Python?
# > - A Function to Programming is what the Atom to Phisics.
# > - Every time you want to do something in programming
# > - **You will need a `function()`** to make it
# > - Theferore, you must **detect parenthesis `()`**
# > - Out of all the words that you see in a website
# > - Because they indicate the presence of a `function()`.
# ## External Functions
# > Download [this Excel](https://github.com/sotastica/data/raw/main/internet_usage_spain.xlsx).
# > Apply the above discipline and make it happen 🚀
# > I want to see the table, c'mon 👇
read_excel()
read_excel = 'asdf'
read_excel()
import panda
import pandas
read_excel()
pandas.api()
pandas.api.extensions.register_extension_dtype
pandas.read_excel()
pandas.read_excel(io=internet_usage_spain.xlsx)
# `internet_usage_spain = ?`
89
34.3
'wjfioew'
weroij
pandas.read_excel(io='internet_usage_spain.xlsx')
pandas.read_excel(io='internet_usage_spain.xlsx',sheet_name=1)
# ## The Elements of Programming
# > - `Library`: where the code of functions are stored.
# > - `Function`: execute several lines of code with one `word()`.
# > - `Parameter`: to **configure** the function's behaviour.
# > - `Object`: **data structure** to store information.
# ## Code Syntax
# **What happens inside the computer when we run the code?**
#
# > In which order Python reads the line of code?
# > - From left to right.
# > - From up to down.
# > Which elements are being used in the previous line of code?
pandas.factorize
pandas.fly()
# 1. `libreria`
# 2. `.` **DOT NOTATION** accedes dentro de la libreria
# 3. `funcion()`
# 4. pasamos `objetos` a los `argumentos`
# 5. ejecutamos
#
# 6. la funcion nos devuelve un objeto
pandas.read_excel(io='internet_usage_spain.xlsx',sheet_name=1)
pandas.read_excel(io=89,sheet_name=1)
pandas.read_excel(io='89',sheet_name=1)
pandas.read_excel(io='internet_usage_spain.xlsx',sheet_name=1)
pandas.read_excel(io='internet_usage_spain.xlsx',sheet_name=1)
import os
os.getcwd()
os.listdir()
os.listdir('datos')
pandas.read_excel(io='datos/internet_usage_spain.xlsx',sheet_name=1)
# ## Functions inside Objects
# > - The `dog` makes `guau()`: `dog.guau()`
# > - The `cat` makes `miau()`: `cat.miau()`
# > - What could a `DataFrame` make? `object.` + `[tab key]`
var = 89
pepa=pandas.read_excel(io='datos/internet_usage_spain.xlsx',sheet_name=1)
89
pepa
pepa
pepa.mean()
pepa.mean(numeric_only=True)
pepa.sum()
pepa.age.sum()
pepa.age.mean()
# # Accessing `Objects`
# > Objects are **data structures** that store information.
# > Which **syntax** do we use to access the information?
# + [markdown] tags=[]
# ## Dot Notation `.`
# -
pepa.age
# ## Square Brackets `[]`
pepa['age']
pepa['age'].mean()
pepa[age]
# # `DataFrame` Manipulation
# > Could we solve the following questions with a `DataFrame` **object**?
# ## How many people are in each study category?
pepa.education.value_counts()
pepa.education.value_counts()
# ## Age average by internet usage
# ## Age average by internet usage and gender
# # Data Visualization with Python
# ## Load Another Dataset
# > We load this [csv file](https://raw.githubusercontent.com/mwaskom/seaborn-data/master/tips.csv).
# > ¿What does it mean `csv`?
# > ¿Why don't we use an `Excel` to better see the data?
# ## Scatterplot
# ### With `Matplotlib` library
# > Scatterplot
# ### With `Seaborn` library
# > Scatterplot
# ### With `Plotly` library
# > Scatterplot
# ## Other Data Visualization Figures
# ### Histogram
# ### Boxplot
# ### Bar Chart
# ### Pie Plot
# ### Maps
| #01. Data Tables & Basic Concepts of Programming/01session.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Output feedback control for the harmonic oscillator
# The harmonic oscillator is a useful model for systems that have a dominating resonance frequency with no, or very little damping. An example of such systems is the sideway movement of a container hanging from a gantry crane moving containers on and off ships.
# 
# <font size="2">by Tosaka, from Wikimedia.org</font>
# + [markdown] slideshow={"slide_type": "slide"}
# Consider a container of mass $m=10^4$ kg, hanging from a wire of length $l=40$ m. We can control the system by applying an acceleration $u$ of the cart on top of the crane. The ODE describing the dynamics of the container is obtained by considering a reference frame fixed in the cart
# $$ ml^2 \ddot{\theta} = -lmg\sin\theta + lm\cos\theta u + l\cos\theta w,$$
# where $\theta$ is the angle of the wires to the vertical, and $w$ is a disturbance force from wind blowing on the container. The small-angle approximation $\sin\theta \approx \theta$ works well in this case, giving the model
# $$\ddot{\theta} = -\frac{g}{l}\theta + \frac{1}{l}u + \frac{1}{lm}w.$$
# Write $y=\theta$ and the model in the s-domain becomes
# $$ Y(s) = \frac{1}{s^2 + \omega^2}\Big(bU(s) + kW(s)\big),$$
# where $\omega^2 = \frac{g}{l}= \frac{9.8}{40} \approx 0.25$ and $b=1/l= 0.025$ and $k=1/(lm) = 2.5\times 10^{-5}$
# + [markdown] slideshow={"slide_type": "slide"}
# The system can be written on state-space form as
# \begin{align}
# \dot{x} &= \underbrace{\begin{bmatrix} 0 & -\omega^2\\1 & 0 \end{bmatrix}}_{A}x + \underbrace{\begin{bmatrix}1\\0\end{bmatrix}}_{B}bu + \begin{bmatrix}1\\0\end{bmatrix}kw\\
# y &= \underbrace{\begin{bmatrix} 0 & 1 \end{bmatrix}}_{C}x
# \end{align}
# + [markdown] slideshow={"slide_type": "slide"}
# ## Discrete-time state-space model
# The discrete-time state-space model using a sampling period $h$ is
# \begin{align}
# x(k+1) &= \Phi(h)x(k) + \Gamma(h)u + \Gamma(h)v\\
# y(k) &= Cx(k)
# \end{align}
# where
# $$ \Phi(h) = \mathrm{e}^{Ah} = \begin{bmatrix} \cos(h\omega) & -\omega\sin(h\omega)\\\frac{1}{\omega}\sin(h\omega) & \cos(h\omega) \end{bmatrix}$$
# and
# $$ \Gamma(h) = \int_0^h \mathrm{e}^{As}B ds = \begin{bmatrix} \frac{1}{\omega}\sin(h\omega)\\\frac{1}{\omega^2} \big(1-\cos(h\omega)\big) \end{bmatrix}.$$
# ### Verification by symbolic computation
# + slideshow={"slide_type": "subslide"}
import numpy as np
import sympy as sy
sy.init_printing(use_latex='mathjax', order='lex')
h,omega = sy.symbols('h,omega', real=True, positive=True)
A = sy.Matrix([[0,-omega**2], [1,0]])
B = sy.Matrix([[1],[0]])
Phi = sy.simplify(sy.exp(A*h).rewrite(sy.sin))
Phi
# + slideshow={"slide_type": "subslide"}
s = sy.symbols('s',real=True, positive=True)
Gamma = sy.simplify(sy.integrate(sy.exp(A*s)*B, (s, 0, h)).rewrite(sy.cos))
Gamma
# + [markdown] slideshow={"slide_type": "slide"}
# ### Choosing the sampling ratio $h$
# We may use the rule-of-thumb $\omega h \approx 0.2\, \text{to} \, 0.6$ for choosing the sampling period. For our specific case we also have $\omega^2 = 0.25$. Let's choose $\omega h = \pi/6 \approx 0.53$, so that $\cos(h\omega) = \frac{\sqrt{3}}{2} \approx 0.866$ and $\sin(h\omega) = 0.5.$ This gives the discrete-time system (ignoring the disturbance for now)
# \begin{align}
# x(k+1) &= \begin{bmatrix} \frac{\sqrt{3}}{2} & -0.25 \\ 1 & \frac{\sqrt{3}}{2} \end{bmatrix} + \begin{bmatrix} 1\\4-2\sqrt{3}\end{bmatrix}0.025u(k)\\
# y(k) &= \begin{bmatrix} 0 & 1\end{bmatrix} x
# \end{align}
# + slideshow={"slide_type": "subslide"}
omegaval = 0.5
hval = np.pi/6/omegaval
Phi_np = np.array(Phi.subs({h:hval, omega:omegaval})).astype(np.float64)
Phi_np
# + slideshow={"slide_type": "subslide"}
Gamma_np = np.array(Gamma.subs({h:hval, omega:omegaval})).astype(np.float64)
Gamma_np
# + slideshow={"slide_type": "skip"}
4-2*np.sqrt(3)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Reachability
# The controllability matrix for this second order system becomes
# $$ W_c = \begin{bmatrix} \Gamma & \Phi\Gamma \end{bmatrix} = \begin{bmatrix} 1 & 0.732\\ 0.536 & 1.464 \end{bmatrix}, $$
# with determinant
# $$\det W_c = 1.072 \neq 0.$$
# + slideshow={"slide_type": "subslide"}
Wc_n = np.hstack((Gamma_np, np.dot(Phi_np,Gamma_np)))
Wc_n
# + slideshow={"slide_type": "subslide"}
np.linalg.det(Wc_n)
# + [markdown] slideshow={"slide_type": "slide"}
# ## State feedback
# Introducing the state-feedback control law
# $$ u = -l_1x_1 - l_2 x_2 + l_0y_{ref} = -Lx + l_0y_{ref}$$
# gives the closed-loop state-space system
# \begin{align}
# x(k+1) &= \Phi x(k) +\Gamma\big(-Lx(k) + l_0y_{ref}(k)\big) + \Gamma v(k) = \left( \Phi - \Gamma L \right) x(k) + l_0\Gamma y_{ref}(k) + \Gamma v(k)\\
# y(k) &= C x(k),
# \end{align}
# where
# $$ \Gamma L = \begin{bmatrix} 1\\0.536\end{bmatrix}\begin{bmatrix}l_1 & l_2\end{bmatrix} = \begin{bmatrix} l_1 & l_2\\0.536 l_1 & 0.536 l_2 \end{bmatrix} $$
# and
# $$ \Phi - \Gamma L = \begin{bmatrix} 0.866 & -0.25 \\ 1 & 0.866\end{bmatrix} - \begin{bmatrix} l_1 & l_2\\0.536 l_1 & 0.536 l_2 \end{bmatrix} = \begin{bmatrix} 0.866-l_1 & -0.25-l_2\\ 1 - 0.536l_1 & 0.866 - 0.536l_2\end{bmatrix}$$
# + slideshow={"slide_type": "skip"}
l1,l2 = sy.symbols('l1,l2')
L = sy.Matrix([[l1, l2]])
Phi_c=Phi.subs({h:hval, omega:omegaval}) - Gamma.subs({h:hval, omega:omegaval})*L
# + [markdown] slideshow={"slide_type": "subslide"}
# with characteristic polynomial given by
# \begin{align}
# \det \left( zI - (\Phi-\Gamma L) \right) &= \det \left( \begin{bmatrix} z & 0\\0 & z \end{bmatrix} - \begin{bmatrix} 1 & h\\0 & 1 \end{bmatrix} + \begin{bmatrix} l_1\frac{h^2}{2} & l_2\frac{h^2}{2}\\ l_1h & l_2h \end{bmatrix} \right)\\
# &= \det \begin{bmatrix} z-1+l_1\frac{h^2}{2} & -h+l_2\frac{h^2}{2}\\l_1h & z-1+l_2h
# \end{bmatrix}\\
# &= (z-1+l_1\frac{h^2}{2})(z-1+l_2h) - l_1h(-h + l_2\frac{h^2}{2})\\
# &= z^2 + (-1+l_2h-1+l_1\frac{h^2}{2}) z + (1-l_2h - l_1\frac{h^2}{2} + l_1l_2\frac{h^3}{2} +l_1h^2 -l_1l_2\frac{h^3}{2})\\
# &= z^2 + (l_1\frac{h^2}{2}+l_2h-2) z + (1 +l_1\frac{h^2}{2} -l_2h)
# \end{align}
# ### Verification by symbolic computation
# -
l1, l2 = sy.symbols('l1, l2', real=True)
z = sy.symbols('z')
L = sy.Matrix([[l1, l2]])
ch_poly = sy.Poly((z*sy.eye(2) - (Phi - Gamma*L)).det(), z)
ch_poly.as_expr()
# ### Desired closed-loop characteristic polynomial
# Here we are interested in designing a deadbeat controller, so the desired closed-loop poles are
# $$ p_1 = 0, \qquad p_2=0,$$
# and the desired characteristic polynomial is
# $$ A_c(z) = (z-p_1)(z-p_2) = z^2. $$
# In the same spirit as when designing an RST controller using the polynomial approach, we set the calculated characteristic polynomial - obtained when introducing the linear state feedback- equal to the desired characteristic polynomial.
# \begin{align}
# z^1: \qquad l_1\frac{h^2}{2} + l_2h -2 &= 0\\
# z^0: \qquad l_1\frac{h^2}{2} - l_2h+1 &= 0
# \end{align}
# which can be written as the system of equations
# $$ \underbrace{\begin{bmatrix} \frac{h^2}{2} & h\\\frac{h^2}{2} & -h \end{bmatrix}}_{M} \underbrace{\begin{bmatrix} l_1\\l_2\end{bmatrix}}_{L^T} = \underbrace{\begin{bmatrix}2\\-1\end{bmatrix}}_{b} $$
# with solution given by
#
# $$L^T = M^{-1}b = \frac{1}{-h^3} \begin{bmatrix} -h & -h\\-\frac{h^2}{2} & \frac{h^2}{2} \end{bmatrix} \begin{bmatrix} 2\\-1 \end{bmatrix}$$
# $$ = -\frac{1}{h^3} \begin{bmatrix} -2h+h\\-h^2-\frac{h^2}{2}\end{bmatrix} = \begin{bmatrix} \frac{1}{h^2}\\\frac{3}{2h} \end{bmatrix} $$
# ### Verification by symbolic calculation
des_ch_poly = sy.Poly(z*z, z)
dioph_eqn = ch_poly - des_ch_poly
sol = sy.solve(dioph_eqn.coeffs(), (l1,l2))
sol
# In the system of equations $ML^T=b$ above, note that the matrix $M$ can be written
# $$ M = \begin{bmatrix} \frac{h^2}{2} & h\\\frac{h^2}{2} & -h \end{bmatrix} = \begin{bmatrix}1 & 0\\-2 & 1\end{bmatrix}\underbrace{\begin{bmatrix} \frac{h^2}{2} & h \\ \frac{3h^2}{2} & h\end{bmatrix}}_{W_c^T}, $$
# so $M$ will be invertible if and only if $\det W_c^T = \det W_c \neq 0$.
# ## The resulting closed-loop system
# So, we have found the control law
# $$ u(k) = -Lx(k) + l_0y_{ref}(k) = -\begin{bmatrix} \frac{1}{h^2} & \frac{3}{2h} \end{bmatrix}x(k) + l_0 y_{ref}(k)$$
# which gives a closed-loop system with poles in the origin, i.e. deadbeat control. The closed-loop system becomes
# \begin{align*}
# x(k+1) &= \big( \Phi - \Gamma L \big) x(k) + \Gamma l_0 y_{ref}(k) + \Gamma v(k)\\
# &= \left( \begin{bmatrix} 1 & h\\0 & 1\end{bmatrix} - \begin{bmatrix} \frac{h^2}{2}\\h\end{bmatrix}\begin{bmatrix} \frac{1}{h^2} & \frac{3}{2h} \end{bmatrix} \right) x(k) + \Gamma l_0 y_{ref}(k) + \Gamma v(k)\\
# &= \left( \begin{bmatrix} 1 & h\\0 & 1\end{bmatrix} - \begin{bmatrix} \frac{1}{2} & \frac{3h}{4}\\ \frac{1}{h} & \frac{3}{2}\end{bmatrix}\right) x(k) + \Gamma l_0 y_{ref}(k) + \Gamma v(k)\\
# &= \underbrace{\begin{bmatrix} \frac{1}{2} & \frac{h}{4} \\-\frac{1}{h} & -\frac{1}{2}\end{bmatrix}}_{\Phi_c}x(k) + \begin{bmatrix}\frac{h^2}{2}\\h\end{bmatrix} l_0 y_{ref}(k) + \begin{bmatrix}\frac{h^2}{2}\\h\end{bmatrix} v(k)\\
# y(k) &= \begin{bmatrix} 1 & 0 \end{bmatrix} x(k)
# \end{align*}
# ### Verification using symbolic computations
L = sy.Matrix([[sol[l1], sol[l2]]])
Phic = Phi - Gamma*L
Phic
# ## Determining the reference signal gain $l_0$
# Consider the steady-state solution for a unit step in the reference signal. We set $y_{ref}=1$ and $v = 0$. This gives
# $$ x(k+1) = \Phi_c x(k) + \Gamma l_0. $$
# In steady-state there is no change in the state, so $x(k+1)=x(k)=x_{ss}$, which leads to
# $$ x_{ss} = \Phi_c x_{ss} + \Gamma l_0$$
# $$ (I - \Phi_c)x_{ss} = \Gamma l_0$$
# \begin{align}
# x_{ss} &= (I - \Phi_c)^{-1}\Gamma l_0\\
# &= \begin{bmatrix} \frac{1}{2} &-\frac{h}{4}\\ \frac{1}{h} & \frac{3}{2} \end{bmatrix}^{-1} \begin{bmatrix} \frac{h^2}{2}\\h \end{bmatrix} l_0\\
# &= \begin{bmatrix}\frac{3}{2} & \frac{h}{4}\\-\frac{1}{h} & \frac{1}{2} \end{bmatrix} \begin{bmatrix} \frac{h^2}{2}\\h\end{bmatrix} l_0\\
# &= \begin{bmatrix}\frac{3h^2}{4} + \frac{h^2}{4}\\-\frac{h}{2} + \frac{h}{2} \end{bmatrix}l_0= \begin{bmatrix}h^2\\ 0 \end{bmatrix}l_0\\
# \end{align}
# which means that the steady-state velocity $\dot{z}(\infty) = x_2(\infty) = 0$. This makes sense.
#
# We can now determine $l_0$. Since $y(k)=x_1(k)$ then $y_{ss} = h^2 l_0$ for a unit step in the reference signal. We would like the steady-state value $y_{ss}$ to be the same as the reference signal (which is equal to one, of course) so this gives
# $$ h^2l_0 = 1 \quad \Rightarrow \quad l_0 = \frac{1}{h^2}. $$
# ## Simulate step responses (symbolically)
# ### Step response from the reference
l0 = 1/(h*h)
C = sy.Matrix([[1,0]])
x = sy.Matrix([[0],[0]]) # Initial state
yref = sy.Matrix([[1]])
xs = [x] # List to hold state trajectory
us = [[0]] # and control signal
ys = [[0]] # and system output
for k in range(6): # No need to simulate too long. It is deadbeat control after all
us.append(-L*x + l0*yref)
x = Phic*x + Gamma*l0*yref
xs.append(x)
ys.append(C*x)
xs
us
# ### Step response from the disturbance
x = sy.Matrix([[0],[0]]) # Initial state
yref = sy.Matrix([[0]])
v = sy.Matrix([[1]])
xs = [x] # List to hold state trajectory
us = [[0]] # and control signal
ys = [[0]] # and system output
for k in range(6): # No need to simulate too long. It is deadbeat control after all
us.append(-L*x + l0*yref)
x = Phic*x + Gamma*l0*yref + Gamma*v
xs.append(x)
ys.append(C*x)
xs
# ## Simulate step-responses (numerically)
import control as ctrl
import matplotlib.pyplot as plt
# Convert to from sympy matrices to numpy
hval = .1
Phi_np = np.array(Phi.subs({h:hval})).astype(np.float64)
Gamma_np = np.array(Gamma.subs({h:hval})).astype(np.float64)
L_np = np.array(L.subs({h:hval})).astype(np.float64)
l0_np = np.array(l0.subs({h:hval})).astype(np.float64)
Phic_np = Phi_np - Gamma_np*L_np
C_np = np.array(C).astype(np.float64)
D_np = np.array([[0]])
sys_c = ctrl.ss(Phic_np, Gamma_np*l0_np, C_np, D_np, hval) # From ref signal
sys_cv = ctrl.ss(Phic_np, Gamma_np, C_np, D_np, hval) # From disturbance signal
tvec = np.asarray(np.arange(8))*hval
T, yout = ctrl.step_response(sys_c, tvec)
T, yout_v = ctrl.step_response(sys_cv, tvec)
plt.figure(figsize=(14,3))
plt.step(tvec, yout.flatten())
plt.figure(figsize=(14,3))
plt.step(tvec, yout_v.flatten())
# # Exercises
# ## Design a less agressive controller
# Consider to let the closed-loop poles be less fast. Choose something reasonable, for instance a double pole in $z=0.5$, or a pair of complex-conjugated poles in $z=0.6 \pm i0.3$. Redo the design, following the example above. Find the state feedback and simulate step-responses.
# ## Design a deadbeat controller for the DC-motor
# From the textbook (Åström & Wittenmark) Appendix:
# 
# 1. Use symbolic calculations to find the discrete-time state-space model for arbitrary sampling period $h$.
# 2. Design a deadbeat controller for arbitrary sampling period.
# 3. Assume a disturbance is acting on the input to the system, as an unknown torque on the motor shaft. This means that the disturbance enters into the system in the same way as the disturbance on the mass on frictionless surface analyzed above. Simulate step-responses for the closed-loop system.
| state-space/.ipynb_checkpoints/Under construction - Output feedback controller for the harmonic oscillator-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:southern_ocean] *
# language: python
# name: conda-env-southern_ocean-py
# ---
# ## <u>Notebook Synopsis</u>:<br>
#
# Here I develop a set of models similar to that of Step-1-, here substituting the single trend component for a piecewise trend sub-model. Specifically I:
#
# * Load the training data generated and saved in previous NB.
# * Develop and combine piecewise trend, seasonal, and residual noise submodels similar to previous NB.
# * Compare models using WAIC or PSIS-LOOCV.
# * Retain and save models predicted to perform better.
# +
import pickle
import pathlib
from platform import python_version as pyver
import pandas as pd
import numpy as np
import pymc3 as pm
import theano.tensor as tt
from sklearn.preprocessing import MinMaxScaler
import arviz as ar
import matplotlib.pyplot as pl
import matplotlib.dates as mdates
from matplotlib import rcParams
# -
def print_ver(pkg, name=None):
try:
print(f'{pkg.__name__}: {pkg.__version__}')
except AttributeError:
print(f'{name}: {pkg}')
print_ver(pyver(), 'python')
for pi in [np, pd, pm, ar]:
print_ver(pi)
# +
# %matplotlib inline
years = mdates.YearLocator(day=1)
months = mdates.MonthLocator(bymonthday=1)
rcParams['xtick.major.size'] = 8
rcParams['xtick.minor.size'] = 4
rcParams['xtick.minor.visible'] = True
rcParams['xtick.labelsize'] = 16
rcParams['ytick.labelsize'] = 16
rcParams['axes.labelsize'] = 16
rcParams['axes.titlesize'] = 18
rcParams['axes.formatter.limits'] = (-3, 2)
# -
df = pd.read_csv('../../dataJar/manning.csv')
df.head()
df['ds'] = pd.to_datetime(df['ds'])
df['y_scaled'] = df.y / df.y.max()
df['t'] = (df.ds - df.ds.min()) / (df.ds.max() - df.ds.min())
df.plot(x='ds', y='y', figsize=(16, 6),)
# ### <u>Modeling a Piecewise Trend</u>:
#
# Within the context of Generalized Additive Models(GAMs), which arise from the simple additive combination of submodels, I develop here a set of models following $$y(t) = g(t) + s(t) + ar1(t)$$
# where \\(y(t)\\) is the modeled signal (chlorophyll in the AOSTZ sector), \\(g(t) \\) is the trend (i.e *rate of change*) sub-model, \\(s(t)\\) is the seasonal sub-model, \\(ar1(t)\\) is the AR1 residual.
#
# The piecewise model is implemented by inserting a fixed number of changepoints such that $$g(t) = (k + a(t)^T\delta)t + (m + a(t)^T\gamma)$$
# where \\(k\\) is the base trend, modified by preset changpoints stored in a vector \\(s\\). At each unique changepoint \\(s_j\\) the trend is adjusted by \\(\delta_j\\), stored in a vector \\(\delta\\), everytime \\(t\\) surpasses a changepoint \\(s_j\\). Used for this purpose, \\(a(t)\\) is basically a vectorized switchboard that turns on for a given switchpoint such that
# \begin{equation}
# a(t) =
# \begin{cases} 1 , & \text{if $t \geq s_j$} \\
# 0 , & \text{otherwise}
# \end{cases}
# \end{equation}
# The second part, \\( m + a(t)^T\gamma\\) ensures the segments defined by the switchpoints are connected. Here, \\(m\\) is an offset parameter, and \\(\gamma_j\\) is set to \\(-s_j\delta_j\\).
#
# The issue though is to find the right number of preset changepoint that will capture actual changepoints while not bogging down the inference. Moreover, for the sake of practicality, these will need to be regularly spaced. Here I try several setups including, one change point at the beginning of the year, and one for every season (4pts/year), one every two months (6pts/year), and one for every month (as many changepoints as data points). The idea is then to put a rather restrictive Laplace prior on \\(\delta\\) to rule out unlikely changepoints, effectively setting the corresponding \\(\delta_j\\) to 0.
#
# First is to define some [helper functions as in the previous notebook](./Step-1-Modeling_AOSTZ_with_pymc3-simpletrend_fourier_seasonality_ar1_residual.ipynb#helpers):
# +
def fourier_series(t, p=12, n=1):
"""
input:
------
t [numpy array]: vector of time index
p [int]: period
n [int]: number of fourier components
output:
-------
sinusoids [numpy array]: 2D array of cosines and sines
"""
p = p / t.size
wls = 2 * π * np.arange(1, n+1) / p
x_ = wls * t[:, None]
sinusoids = np.concatenate((np.cos(x_), np.sin(x_)), axis=1)
return sinusoids
def seasonality(mdl, n_fourier, t):
"""
m [pymc3 Model class]: model object
n_fourier [int]: number of fourier components
t [numpy array]: vector of time index
"""
with mdl:
σ = pm.Exponential('σ', 1)
f_coefs = pm.Normal('fourier_coefs', 0, sd=1, shape=(n_fourier*2))
season = tt.dot(fourier_series(t, n=n_fourier), f_coefs)
return season
def piecewise_trend(mdl, s, t, a_t, obs,
k_prior_scale=5, δ_prior_scale=0.05, m_prior_scale=5):
"""
input:
------
mdl [pymc3 Model class]: model object
s [numpy array]: changepoint vector
t [numpy array]: time vector
obs [numpy array]: vector of observations
a_t [numpy int array]: 2D (t*s) adjustment indicator array
k_prior_scale [float]: base trend normal prior scale parameter (default=5)
δ_prior_scale [float]: trend adjustment laplace prior scale param. (default=0.05)
m_prior_scale [float]: base offset normal prior scale param. (default=5)
"""
with mdl:
# Priors:
k = pm.Normal('k', 0, k_prior_scale) # base trend prior
if δ_prior_scale is None:
δ_prior_scale = pm.Exponential('τ', 1.5)
δ = pm.Laplace('δ', 0, δ_prior_scale, shape=s.size) # rate of change prior
m = pm.Normal('m', 0, m_prior_scale) # offset prior
γ = -s * δ
trend = (k + tt.dot(a_t, δ)) * t + (m + tt.dot(a_t, γ))
return trend
def ar1_residual(mdl, n_obs):
with mdl:
k_ = pm.Uniform('k', -1.1, 1.1)
tau_ = pm.Gamma('tau', 10, 3)
ar1 = pm.AR1('ar1', k=k_, tau_e=tau_, shape=n_obs)
return ar1
def changepoint_setup(t, n_changepoints, s_start=None, s=None, changepoint_range=1):
"""
input:
------
t [numpy array]: time vector
n_changepoints [int]: number of changepoints to consider
s [numpy array]: user-specified changepoint vector (default=None)
s_start [int]: changepoint start index (default=0)
changepoint_range[int]: adjustable time proportion (default=1)
output:
-------
s [numpy array]: changepoint vector
a_t [numpy int array]: 2D (t*s) adjustment indicator array
"""
if s is None:
if s_start is None:
s = np.linspace(start=0, stop=changepoint_range*t.max(),
num=n_changepoints+1)[1:]
else:
s = np.linspace(start=s_start, stop=changepoint_range*t.max(),
num=n_changepoints)
a_t = (t[:,None] > s) * 1
return a_t, s
def model_runner(t_, obs_s, add_trend=False, add_season=False, add_AR1=False,
**payload):
mdl = pm.Model()
a_t, s = None, None
with mdl:
y_ = 0
σ = pm.HalfCauchy('σ', 2.5)
if add_trend:
n_switches = payload.pop('n_switches', t_.size)
s_start = payload.pop('s_start', None)
s = payload.pop('s', None)
chg_pt_rng = payload.pop('changepoint_range', 1)
k_prior_scale = payload.pop('k_prior_scale', 5)
δ_prior_scale = payload.pop('δ_prior_scale', 0.05)
m_prior_scale = payload.pop('m_prior_scale', 5)
a_t, s = changepoint_setup(t_, n_switches, s_start=s_start, s=s,
changepoint_range=chg_pt_rng)
trend_ = piecewise_trend(mdl, s, t_, a_t, obs_s,
k_prior_scale, δ_prior_scale, m_prior_scale)
y_ += trend_
if add_season:
n_fourier = payload.pop('n_fourier', 4)
season = seasonality(mdl, n_fourier=n_fourier, t=t_)
y_ += season
if add_AR1:
ar1 = ar1_residual(mdl, obs_s.size)
y_ += ar1
pm.Normal('obs', mu=y_, sd=σ, observed=obs_s)
return mdl, a_t, s
def sanity_check(m, df):
"""
:param m: (pm.Model)
:param df: (pd.DataFrame)
"""
# Sample from the prior and check of the model is well defined.
y = pm.sample_prior_predictive(model=m, vars=['obs'])['obs']
pl.figure(figsize=(16, 6))
pl.plot(y.mean(0), label='mean prior')
pl.fill_between(np.arange(y.shape[1]), -y.std(0), y.std(0), alpha=0.25, label='standard deviation')
pl.plot(df['y_scaled'], label='true value')
pl.legend()
# -
mdl_trend_only, A, s_pts = model_runner(df.t, df.y_scaled, add_trend=True,
n_switches=25, changepoint_range=0.8)
# And run the sanity check
sanity_check(mdl_trend_only, df)
with mdl_trend_only:
aprox = pm.find_MAP()
# +
def det_trend(k, m, delta, t, s, A):
return (k + np.dot(A, delta)) * t + (m + np.dot(A, (-s * delta)))
g = det_trend(aprox['k'], aprox['m'], aprox['δ'],
df['t'], s_pts, A) * df['y'].max()
pl.figure(figsize=(16, 6))
pl.title('$g(t)$')
pl.plot(g)
pl.scatter(np.arange(df.shape[0]), df.y, s=0.5, color='black')
# -
mdl_trend_season, A, s_pts = model_runner(df.t, df.y_scaled, add_trend=True,
n_switches=25, changepoint_range=0.8,
δ_prior_scale=None, n_fourier=4)
sanity_check(mdl_trend_season, df)
| Part-2-PyMC3-modeling/ipynb/GAM/Step-2-Modeling_AOSTZ_with_pymc3-Piecewise_trend_model-testing_w_manning_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
adult.dtypes
adult.income.values
#Only numeric data for regression
X=adult[["age ","fnlwgt","education-num","capital-gain","capital-loss","hours-per-week"]].values
X
from sklearn import datasets ## imports datasets from scikit-learn
data = datasets.load_boston() ## loads Boston dataset from datasets library
print (data.DESCR)
import numpy as np
import pandas as pd
# +
# define the data/predictors as the pre-set feature names
df = pd.DataFrame(data.data, columns=data.feature_names)
# Put the target (housing value -- MEDV) in another DataFrame
target = pd.DataFrame(data.target, columns=["MEDV"])
# +
## Without a constant
import statsmodels.api as sm
X = df["RM"]
y = target["MEDV"]
# -
# Note the difference in argument order
model = sm.OLS(y, X).fit()
predictions = model.predict(X) # make the predictions by the model
# Print out the statistics
model.summary()
mtcars=pd.read_csv("https://raw.githubusercontent.com/vincentarelbundock/Rdatasets/master/csv/datasets/mtcars.csv")
mtcars
# +
X = mtcars[["disp","wt","qsec"]]
y = mtcars["mpg"]
# -
# Note the difference in argument order
model = sm.OLS(y, X).fit()
predictions = model.predict(X) # make the predictions by the model
# Print out the statistics
model.summary()
X
y
#http://scikit-learn.org/stable/auto_examples/linear_model/plot_ols.html
import matplotlib.pyplot as plt
import numpy as np
from sklearn import datasets, linear_model
from sklearn.metrics import mean_squared_error, r2_score
# Split the data into training/testing sets
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
#data_X_train = data_X[:-20]
#data_X_test = data_X[-20:]
X_train
X_test
# +
# Create linear regression object
regr = linear_model.LinearRegression()
# Train the model using the training sets
regr.fit(X_train, y_train)
# +
# Make predictions using the testing set
y_pred = regr.predict(X_test)
# The coefficients
print('Coefficients: \n', regr.coef_)
# The mean squared error
print("Mean squared error: %.2f"
% mean_squared_error(y_test, y_pred))
# Explained variance score: 1 is perfect prediction
print('Variance score: %.2f' % r2_score(y_test, y_pred))
# -
| linear regression using statsmodel and scikit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Strategy Risk
#
# Most of the functions below can be found under:
#
# * Backtest/stats_measure
# * Sample_data/make_data
#
# > Strategy risk is different from portfolio risk.
# >
# > Advance in Financial Machine Learning [2018], <NAME>.
#
# Strategy risk is inherent within investment management style, method and technique, reflected as a mismatch between expectation and eventual outcome.
#
# While portfolio risk is directly derived from underlying assets itself.
#
# If you are keen on generating synthetic data for your research, copy the code snippets [Generate synthetic raw data](https://gist.github.com/boyboi86/5e00faf48f60abfdbe838fbdee269471) in my gist.
#
# Contact: <EMAIL>
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import research as rs
from scipy.stats import norm, kurtosis
from scipy.stats import skew as Skew
# %matplotlib inline
# +
def _target_sr(p: float = 0.5, freq: int = 52, ptsl: list = [0.02,-0.02], seed: int = None):
if seed is not None:
np.random.seed(seed)
rnd=np.random.binomial(n=1,p=p, size = freq)
x = [ptsl[0] if i == 1 else ptsl[1] for i in rnd]
mean = np.mean(x)
std = np.std(x)
return (mean, std, mean/std)
def target_sr(p: float = 0.5, freq: int = 52, ptsl: list = [0.02,-0.02], n_run: int = 1000000, seed: int = None):
mean, std, sr = 0, 0, []
for n in np.arange(n_run):
_mean, _std, _sr = _target_sr(p = p, freq = freq, ptsl = ptsl, seed = seed)
mean += _mean
std += _std # std is only 0.1, because variance root
if _sr <= .2: sr.append(_sr)
mean = mean/n_run # var = 0.01 but std = 0.1
std = std/n_run
im_sr = mean/std
print("Mean: {0:.6f}\nStd: {1:.6f}\nSharpe Ratio: {2:.6f}".format(mean, std, im_sr))
if len(sr) >0:
p =len(sr)/n_run
print("Probability of getting SR < 2.: {0}%".format(100 * p))
return (mean, std, im_sr, sr)
else:
print("All SR >= 2")
return (mean, std, im_sr)
def im_p(freq: int = 52, trgt_sr: int = 2., ptsl: list = [0.02, -0.02]):
pt, sl = ptsl[0], ptsl[1]
a = (freq + trgt_sr ** 2) * (pt - sl) ** 2
b = (2 * freq * sl - trgt_sr ** 2 * (pt-sl)) * (pt-sl)
c = freq * sl ** 2
p = (-b+(b ** 2 - 4*a*c) ** .5)/ (2*a)
print("\nImplied Precision Rate Required: {0:.6f}".format(p))
return p
def im_freq(p: float = 0.6, trgt_sr: int = 2., ptsl: list = [0.02, -0.02]):
pt, sl = ptsl[0], ptsl[1]
freq = (trgt_sr * (pt - sl)) ** 2*p*(1-p)/((pt-sl)*p+sl)**2
print("\nImplied Frequency Required: {0:.6f}".format(freq))
return int(freq)
def im_pt(freq: int, trgt_sr: float, p: float, sl: float):
pt = (sl * freq**(1/2)) / (trgt_sr * (p*(1-p))**(1/2) - p * freq**(1/2)) + sl
print("\nImplied Profit-taking level: {0:.6f}".format(pt))
return pt
def im_sl(freq: int, p: float, pt: float, trgt_sr: float):
sl = (pt * (trgt_sr * (p*(1-p))**(1/2) - p * freq**(1/2))) / \
(trgt_sr * (p*(1-p))**(1/2) - p * freq**(1/2) + freq**(1/2))
print("\nImplied Stop-loss limit: {0:.6f}".format(sl))
return sl
# -
# **Note**
#
# According to the book, it was using random without a seed.
#
# Hence Monte-Carlos seems appropriate, but if you notice without a seed, it's impossible to get a consistant result.
#
# In order to see if the outcome is reliable, I included a probability measure to see if the return sharpe ratio is reliable.
#
# At least one of the criteria has to be fulfilled:
#
# * As long as Probability of getting SR below 2. is less than 50%.
# * All SR >= 2.
#
# The idea is simple, if we manage to reduce the probability of not attaining SR > 2, it would reflect in our probability.
#
# Likewise, if we were to improve SR, the probability of not getting SR < 2 will decrease (Since we manage to pull SR from origin).
#
# Hence, we can safely assume the SR is somewhat reliable (At least 2 >=).
#
# **Note**
#
# Before you intend to "Annualize" the return.
#
# Kindly refer to this [Stackoverflow](https://quant.stackexchange.com/questions/2260/how-to-annualize-sharpe-ratio)
# +
trgt_sr = 2.
n_run = 100000
p = 0.6
freq = 52
ptsl = [.02,-.02]
trgt = target_sr(p = p,
freq = freq,
ptsl = ptsl,
n_run = n_run,
seed = None)
# +
_p = im_p(freq = freq, trgt_sr = trgt_sr, ptsl = ptsl)
p_trgt = target_sr(p = _p, #use implied precision
freq = freq,
ptsl = ptsl,
n_run = n_run,
seed = None)
chg_p = (_p - 0.6)/ 0.6
# +
_freq = im_freq(p = p, trgt_sr = trgt_sr, ptsl = ptsl)
f_trgt = target_sr(p = p,
freq = _freq, #use implied freq
ptsl = ptsl,
n_run = n_run,
seed = None)
chg_f = (_freq - 52)/52
# +
_pt = im_pt(freq = freq, trgt_sr = trgt_sr, p = p, sl = ptsl[1])
pt_trgt = target_sr(p = p,
freq = freq,
ptsl = [_pt, ptsl[1]], #use implied profit taking
n_run = n_run,
seed = None)
chg_pt = (_pt - 0.02)/0.02
# +
_sl = im_sl(freq = freq, p = p, pt = ptsl[0], trgt_sr = trgt_sr)
sl_trgt = target_sr(p = p,
freq = freq,
ptsl = [ptsl[0], _sl], #use implied stop loss
n_run = n_run,
seed = None)
chg_sl = (_sl + 0.02)/ 0.02
# -
# **Note**
#
# SR required is 0.2 (Equal to SR 2 in the book)
#
# It is not possible to achieve a SR 2, after running Monte-Carlos 100,000 times for theoriginal input.
#
# The Average Sharpe Ratio was around 2.
#
# On top of that it has a high risk of not achieve the intended amount with a low precision rate 0.6.
#
# We can assume that Sharpe Ratio of 2 not possible, probably below 2 with the given parameter. (Not viable Strategy)
#
# **Note**
#
# Minimal precision required: 0.6336306209562121
#
# Otherwise, our we will have high probability SR < 2.
#
# **Note**
#
# If the strategy was run on a daily basis. Trading days = 252.
#
# Min Bets Frequency Required: 96.0
#
# **Note**
#
# Profit-taking level should be around 2.4% at least.
#
# Optimal Profit-taking level: 0.023092
#
# **Note**
#
# Alternate Stop loss level should be around 1.8% at least to attain sharpe ratio 2.
#
# optimal stop loss limit: -0.017322
# +
p_sr, f_sr = (p_trgt[2] * 10 - trgt_sr), (f_trgt[2] * 10 - trgt_sr)
pt_sr, sl_sr = (pt_trgt[2] * 10 - trgt_sr), (sl_trgt[2] * 10 - trgt_sr)
print("\n1 % change in precision will result in {0:.3f}% change in SR\n".format(p_sr/ chg_p))
print("1 % change in frequency will result in {0:.3f}% change in SR\n".format(f_sr/ chg_f))
print("1 % change in profit-taking will result in {0:.3f}% change in SR\n".format(pt_sr/ chg_pt))
print("1 % change in stop-loss will result in {0:.3f}% change in SR\n".format(sl_sr/ chg_sl))
# -
# ### Conclusion
#
# Precision rate will result in the most change for SR in all 4 factors.
#
# Lowest hang fruit is practically the precision rate.
#
# Precision Rate> Stop Loss> Profit Taking > Frequency
#
# Precision and Frequency will definitely affect the profit-taking and stop-loss.
#
# Ultimately based on the algorithm:
#
# * Low precision rate with high frequency = frequent stop-loss
# * High Precision rate with high frequency = frquenct profit-taking
# * Low precision with low frequency = less stop-loss triggered
# * High precision with low frequency = less profit-taking triggered
#
# With SR in mind, precision rate along with frequency play a major role, profit-taking and stop-loss limit is more "reactive" to the former 2.
#
# If you have low precision rate, having higher stop-loss and lower profit-taking limits might be a good idea.
#
# However, there is some trade-off between frequency and precision rate. But the changes might not be significant unless frequency is really large.
#
# **Note**
#
# If you are still not convinced, kindly refer to the mathematical formula above.
# +
def mix_gauss(mu1: float, mu2: float, sig1: float, sig2: float, p: float, n_obs: int):
rtn1 = np.random.normal(mu1, sig1, size=int(n_obs * p))
rtn2 = np.random.normal(mu2,sig2, size=int(n_obs) - rtn1.shape[0])
rtn = np.append(rtn1,rtn2, axis = 0)
np.random.shuffle(rtn)
return rtn
def prob_failure(rtn: float, freq: int, trgt_sr: float):
pos_rtn, neg_rtn = rtn[rtn>0].mean(), rtn[rtn<=0].mean()
p = rtn[rtn>0].shape[0]/ float(rtn.shape[0])
ptsl = [pos_rtn, neg_rtn]
threshold = im_p(freq = freq, trgt_sr = trgt_sr, ptsl = ptsl)
risk = norm.cdf(threshold, p, p * (1 - p))
print("Predicted Precision Pate: {0:.6f}\n".format(p))
return risk
def strategy_failure(mu1: float, mu2: float, sig1: float, sig2: float, p: float, n_obs: int, freq: int, trgt_sr: float):
rtn = mix_gauss(mu1 = mu1,
mu2 = mu2,
sig1 = sig1,
sig2 = sig2,
p = p,
n_obs = n_obs)
_proba_failure = prob_failure(rtn = rtn,
freq = freq,
trgt_sr = trgt_sr)
print("Strategy Failure Probability: {0:.5f}".format(_proba_failure))
if _proba_failure> 0.05:
print("Discard Strategy; High risk indicated")
else:
print("Accept Strategy; Moderate risk indicated")
# -
# **Note**
#
# The first 4 moments can be calculated:
#
# 1. Mean
# 2. Variance
# 3. Skewness
# 4. Kurtosis
#
# **Note**
#
# Please learn to differentiate between raw moments and central moments.
# +
mu1, mu2, sig1, sig2, p, n_obs = -.1, 0.06, 0.12, .03, .15, 12 * 2
rtn = mix_gauss(mu1 = mu1,
mu2 = mu2,
sig1 = sig1,
sig2 = sig2,
p = p,
n_obs = n_obs)
mean, std, skew, kurt = np.mean(rtn), np.std(rtn), Skew(rtn), kurtosis(rtn)
print("\n1st mts: {0}\n2nd mts: {1}\n3rd mts: {2}\n4th mts: {3}\n".format(mean, std, skew, kurt))
sr1 = rs.sharpe_ratio(rtn = rtn, rf_param = .0, t_days = 12 * 2)
benchmark = mean/ (std * (12 * 2) ** 0.5)
print("Annualized Sharpe Ratio: {0:.6f}\nBenchmark: {1:.6f}\n".format(sr1, benchmark))
proba = rs.proba_sr(obs_sr = sr1,
benchmark_sr = benchmark,
num_returns = 12 * 2,
skew_returns = skew,
kurt_returns = kurt)
if proba > 0.95:
print("At 5% significance level, Sharpe Ratio: {0:.6f}".format(sr1))
print("Accept Strategy, as indicated by PSR")
trgt_sr = sr1
else:
print("At 5% significance level, Sharpe Ratio: {0:.6f}".format(sr1))
print("Discard Strategy, as indicated by PSR")
trgt_sr = 2.
# -
strategy_failure(mu1 = mu1,
mu2 = mu2,
sig1 = sig1,
sig2 = sig2,
p = p,
n_obs = n_obs,
freq = n_obs,
trgt_sr= trgt_sr) # depends if they pass first test
# **Note**
#
# Because the data is randomly generated, hence the conclusion may not always be in line with learning outcome.
#
# But the "correct" answer seems to be accept strategy for PSR and discard strategy for Strat-Risk Metric.
# ### Probabilistic SR vs Strategy Risk Metrics
#
# Which method is better? It depends.
#
# As demostrated in [AFML 14.1](https://github.com/boyboi86/AFML/blob/master/AFML%2014.1.ipynb)
#
# PSR tends to favor "normal" distributed returns.
#
# As a result, it will reject "Too good or too bad to be true returns". Hence it follows a gaussian shape as a measure based on benchmark provided.
#
# Strategy risk metric focus on key factor which determined the risk/ return that intended strategy might faced.
#
# As demostrated earlier in this exercise, key factor is precision rate which will impact around 14% SR per 1% PR change.
#
# Return distribution would most likely display asymmetrical returns and tends to be bias. (SR-maximization/ Return-driven)
#
# **Consider the below graph**
#
# The two graphs are generated with different properties:
#
# * Asymmetrical return with higher bias (Higher frequency/ lower variance)
# * Symmetrical return with higher variance (Lower Frequency/ lower bias)
#
# Notice there is an overlapping area between these 2 distributions, therefore there is a mutually inclusive area which both can agreed upon.
#
# As long as the back-test return distribution can fall within the overlapped region (The sweet spot between SR-maximization and realistic outcome).
#
# It will be favored and endorsed by both metrics.
#
# As such, these 2 methods can and are considered complementary to each other.
psr_favor = np.random.normal(0, 2, 10000)
strat_risk_favor = np.random.normal(2,1.5,10000)
df = pd.DataFrame({'PSR Favor': psr_favor, 'SRM Favor': strat_risk_favor})
df.plot(kind='kde', figsize=(12,8), grid=True)
plt.axvline(x=0,ls='--',c='r')
plt.show()
| AFML 15.1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Part 1 - Introduction to Grid
#
# ##### Grid is a platform to **train**, **share** and **manage** models and datasets in a **distributed**, **collaborative** and **secure way**.
#
# Grid platform aims to be a secure peer to peer platform. It was created to use pysyft's features to perform federated learning processes without the need to manage distributed workers directly. Nowadays, to perform machine learning process with PySyft library, the user needs to manage directly all the workers' stuff (start nodes, manage node connections, turn off nodes, etc). Grid platform solves this in a transparent way. The user won't need to know about how the nodes are connected or where is some specific dataset.
#
# Authors:
# - <NAME> - Github: [IonesioJunior](https://github.com/IonesioJunior)
#
# ## Why should we use grid?
# As mentioned before, the grid is basically a platform that uses PySyft library to manage distributed workers providing some special features.
#
# **We should use grid to:**
# - Train models using datasets that we've never seen (without getting access to its real values).
# - Train a model with encrypted datasets.
# - Provide Secure MLaaS running encrypted model inferences across grid network.
# - We can serve an encrypted model without giving its real weights to anyone.
# - We can run encrypted inferences without sending our private data to anyone.
# - Mitigate risks and impacts using Federated Learning's **"privacy by design"** property.
# - Manage the privacy level of datasets stored at grid network allowing/disallowing access to them.
#
# ## How it works?
# We have two concepts of grid: **Private Grid Platform** and **Public Grid Platform**
# ### Private Grid
# ###### Private Grid is used to build private's grid platform.
# It will empower you with the control to manage the entire platform, you'll be able to create, remove and manage all nodes connected on your grid network. However, with power and control, you'll need to take care of the grid platform by yourself.
#
# - To build it, you'll need to know previously where is each grid node that you want to use in your infrastructure.
# - You will need to configure scale up/scale down routines (nº of nodes) by yourself.
# - You can add pr remove nodes.
# - You will be connected directly with these nodes.
#
# <p align="center">
# <img height="600px" width="600px" src="https://github.com/OpenMined/rfcs/blob/master/20190821-grid-platform/DHT-grid.png?raw=true">
# </p>
#
# +
import syft as sy
import torch as th
from syft.grid.clients.data_centric_fl_client import DataCentricFLClient
hook = sy.TorchHook(th)
# +
# How to build / use a private grid network
# 1 - Start the grid nodes.
# 2 - Connect to them directly
# 3 - Create Private Grid using their instances.
# We need to know the address of every node.
node1 = DataCentricFLClient(hook, "ws://localhost:3000")
node2 = DataCentricFLClient(hook, "ws://localhost:3001")
node3 = DataCentricFLClient(hook, "ws://localhost:3002")
node4 = DataCentricFLClient(hook, "ws://localhost:3003")
my_grid = sy.PrivateGridNetwork(node1,node2,node3,node4)
# -
# ### Public Grid
# ###### Public Grid offers the oportunity to work as a real collaborative platform.
# Unlike the private grid, anyone has the power to control all nodes connected to the public grid, the platform will be managed by grid gateway. This component will update the network automatically and perform queries through the nodes. It's important to note that the grid gateway can **only perform non-privileged commands** on grid nodes, it will avoid some vulnerabilities.
#
# Therefore, anyone can register a new node, upload new datasets using their nodes to share it with everyone in a secure way.
#
#
# Public Grid should work as a **Secure Data Science platform** (such as Kaggle, but using Privacy-Preserving concepts):
# - We send pointers to datasets instead of real datasets.
# - We can share our models across the network in an encrypted way.
# - We can run inferences using our sensitive datasets without send the real value of it to anyone.
# <p align="center">
# <img height="600px" width="600px" src="https://github.com/OpenMined/rfcs/blob/master/20190821-grid-platform/partially_grid.png?raw=true">
# </p>
# +
# How to build/use a public grid network
# 1 - Start the grid nodes
# 2 - Register them at grid gateway component
# 3 - Use grid gateway to perform queries.
# You just need to know the address of grid gateway.
my_grid = sy.PublicGridNetwork(hook, "http://localhost:5000")
# -
# # Congratulations!!! - Time to Join the Community!
#
# Congratulations on completing this notebook tutorial! If you enjoyed this and would like to join the movement toward privacy preserving, decentralized ownership of AI and the AI supply chain (data), you can do so in the following ways!
#
# ### Star PySyft on GitHub
#
# The easiest way to help our community is just by starring the GitHub repos! This helps raise awareness of the cool tools we're building.
#
# - [Star PySyft](https://github.com/OpenMined/PySyft)
#
# ### Join our Slack!
#
# The best way to keep up to date on the latest advancements is to join our community! You can do so by filling out the form at [http://slack.openmined.org](http://slack.openmined.org)
#
# ### Join a Code Project!
#
# The best way to contribute to our community is to become a code contributor! At any time you can go to PySyft GitHub Issues page and filter for "Projects". This will show you all the top level Tickets giving an overview of what projects you can join! If you don't want to join a project, but you would like to do a bit of coding, you can also look for more "one off" mini-projects by searching for GitHub issues marked "good first issue".
#
# - [PySyft Projects](https://github.com/OpenMined/PySyft/issues?q=is%3Aopen+is%3Aissue+label%3AProject)
# - [Good First Issue Tickets](https://github.com/OpenMined/PySyft/issues?q=is%3Aopen+is%3Aissue+label%3A%22good+first+issue%22)
#
# ### Donate
#
# If you don't have time to contribute to our codebase, but would still like to lend support, you can also become a Backer on our Open Collective. All donations go toward our web hosting and other community expenses such as hackathons and meetups!
#
# [OpenMined's Open Collective Page](https://opencollective.com/openmined)
| examples/tutorials/grid/Part 01 - Intro to Grid Platform.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img align="left" src="https://ithaka-labs.s3.amazonaws.com/static-files/images/tdm/tdmdocs/CC_BY.png"><br />
#
# Created by [<NAME>](http://nkelber.com) and Ted Lawless for [JSTOR Labs](https://labs.jstor.org/) under [Creative Commons CC BY License](https://creativecommons.org/licenses/by/4.0/)<br />
# For questions/comments/improvements, email <EMAIL>.<br />
# ___
# # Python Basics 2
#
# **Description:** This lesson describes the basics of [flow control statements](https://docs.constellate.org/key-terms/#flow-control-statement) including:
# * [Boolean values](https://docs.constellate.org/key-terms/#boolean-value)
# * [Boolean operators](https://docs.constellate.org/key-terms/#boolean-operator)
# * [Comparison operators](https://docs.constellate.org/key-terms/#comparison-operator)
# * `if` statements
# * `else` statements
# * `elif` statements
# * `while` and `for` loop statements
# * Handling errors with `try` and `except`
#
# and the basics of writing functions:
#
# * `def` statements
# * [Local scope](https://docs.constellate.org/key-terms/#local-scope)
# * [Global scope](https://docs.constellate.org/key-terms/#global-scope)
#
# This is part 2 of 3 in the series *Python Basics* that will prepare you to do text analysis using the [Python](https://docs.constellate.org/key-terms/#python) programming language.
#
# **Use Case:** For Learners (Detailed explanation, not ideal for researchers)
#
# **Difficulty:** Beginner
#
# **Completion Time:** 90 minutes
#
# **Knowledge Required:**
# * [Getting Started with Jupyter Notebooks](./getting-started-with-jupyter.ipynb)
# * [Python Basics 1](./python-basics-1.ipynb)
#
# **Knowledge Recommended:** None
#
# **Data Format:** None
#
# **Libraries Used:** `random` to generate random numbers
#
# **Research Pipeline:** None
# ___
# [](https://www.youtube.com/watch?v=yzApQqZkJ2c)
# ## Flow Control Statements
#
# In *Python Basics 1*, you learned about [expressions](https://docs.constellate.org/key-terms/#expression), [operators](https://docs.constellate.org/key-terms/#operator), [variables](https://docs.constellate.org/key-terms/#variable), and a few native [Python](https://docs.constellate.org/key-terms/#python) [functions](https://docs.constellate.org/key-terms/#). We wrote programs that executed line-by-line, starting at the top and running to the bottom. This approach works great for simple programs that may execute a few tasks, but as you begin writing programs that can do multiple tasks you'll need a way for your programs to decide which action comes next. We can control when (or if) code gets executed with [flow control statements](https://docs.constellate.org/key-terms/#flow-control-statement). If a program is a set of steps for accomplishing a task, then [flow control statements](https://docs.constellate.org/key-terms/#flow-control-statement) help the program decide the next action.
#
# [Flow control statements](https://docs.constellate.org/key-terms/#flow-control-statement) work like a flowchart. For example, let's say your goal is to hang out and relax with friends. There are a number of steps you might take, depending on whether your friends are available or you feel like making some new friends.
#
# 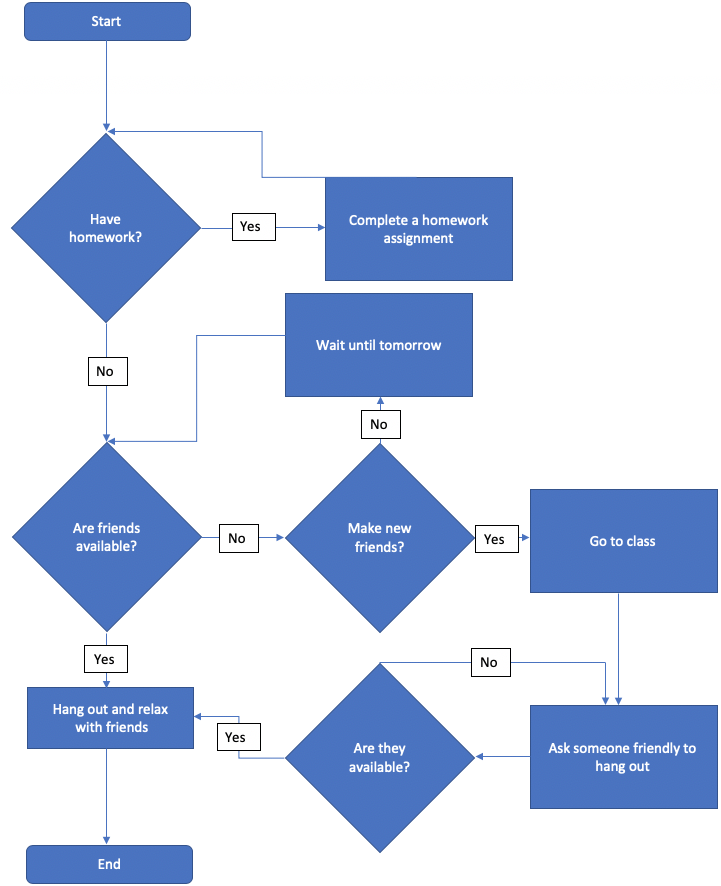
#
# Each diamond in our flowchart represents a decision that has to be made about the best step to take next. This is the essence of [flow control statements](https://docs.constellate.org/key-terms/#flow-control-statement). They help a program decide what the next step should be given the current circumstances.
# ### Boolean Values
#
# One way we to create [flow control statements](https://docs.constellate.org/key-terms/#flow-control-statement) is with [boolean values](https://docs.constellate.org/key-terms/#boolean-value) that have two possible values: **True** or **False**. In our example above, we could consider a "Yes" to be "True" and a "No" to be "False." When we have the data we need to answer each question, we could store that answer in a variable, like:
#
# * ```are_friends_available = False```
# * ```make_new_friends = True```
# * ```new_friend_available = True```
#
# This would allow us to determine which action to take next. When we assign [boolean values](https://docs.constellate.org/key-terms/#boolean-value) to a [variable](https://docs.constellate.org/key-terms/#variable), the first letter must be capitalized:
# Note, the first letter of a boolean value must always be capitalized in Python
are_friends_available = false
print(are_friends_available)
# The boolean values **True** and **False** cannot be used for variable names.
# Treating the boolean value True as a variable will create an error
True = 7
# +
# But we can store Boolean values in a variable
pizza_is_life = False
tacos_are_life = True
# -
# ### Comparison Operators
# Now that we have a way to store [integers](https://docs.constellate.org/key-terms/#integer), [floats](https://docs.constellate.org/key-terms/#float), [strings](https://docs.constellate.org/key-terms/#string), and [boolean values](https://docs.constellate.org/key-terms/#boolean-value) in [variables](https://docs.constellate.org/key-terms/#variable), we can use a [comparison operator](https://docs.constellate.org/key-terms/#comparison-operator) to help make decisions based on those values. We used the [comparison operator](https://docs.constellate.org/key-terms/#comparison-operator) `==` in *Python Basics 1*. This operator asks whether two [expressions](https://docs.constellate.org/key-terms/#expression) are equal to each other.
# Comparing two values with the comparison operator ==
67 == 67
# Note, a comparison operator uses ==
# Do not confuse with variable assignment statement which uses a single =
67 = 67
# There are additional [comparison operators](https://docs.constellate.org/key-terms/#comparison-operator) that can help us with [flow control statements](https://docs.constellate.org/key-terms/#flow-control-statement).
#
# |Operator|Meaning|
# |---|---|
# |==|Equal to|
# |!=|Not equal to|
# |<|Less than|
# |>|Greater than|
# |<=|Less than or equal to|
# |>=|Greater than or equal to|
# Use the "Not equal to" operator
# Use the "equal to" operator with a string
# A string cannot be equal to a float or an integer.
# Try using the "equal to" operator to compare a string with an integer
# But an integer can be equal to a float.
# Try using the "equal to" operator to compare an integer with a float
# We can use the comparison operator with variables.
# +
# Using a comparison operator on a variable
# Create a variable number_of_dogs and assign the value of zero
# Check whether number_of_dogs is greater than or equal to 1
# -
# ### Boolean Operators (and/or/not)
# We can also use [Boolean operators](https://docs.constellate.org/key-terms/#boolean-operator) (**and**/**or**/**not**) to create [expressions](https://docs.constellate.org/key-terms/#expression) that evaluate to a single [Boolean value](https://docs.constellate.org/key-terms/#boolean-value) (**True**/**False**).
#
# #### Using the Boolean Operator `and`
# The `and` operator determines whether *both* conditions are **True**.
# If condition one is True AND condition two is True
# What will the evaluation be?
True and True
# If condition one is True AND condition two is False
# What will the evaluation be?
True and False
# In order for an ```and``` [expression](https://docs.constellate.org/key-terms/#expression) to evaluate to **True**, every condition must be **True**. Here is the "Truth Table" for every pair:
#
# |Expression|Evaluation|
# |---|---|
# |True and True|True|
# |True and False|False|
# |False and True|False|
# |False and False|False|
#
# Since `and` [expressions](https://docs.constellate.org/key-terms/#expression) require all conditions to be **True**, they can easily result in **False** evaluations.
#
# #### Using the Boolean Operator ```or```
# The ```or``` operator determines whether *any* condition is **True**.
# Is expression one True OR is expression two True?
True or False
# Is condition one True OR is condition two True?
False or False
# An ```or``` [expression](https://docs.constellate.org/key-terms/#expression) evaluates to **True** if *any* condition is **True**. Here is the "Truth Table" for every pair:
#
# |Expression|Evaluation|
# |---|---|
# |True or True|True|
# |True or False|True|
# |False or True|True|
# |False or False|False|
#
# Since ```or``` [expressions](https://docs.constellate.org/key-terms/#expression) only require a single condition to be **True**, they can easily result in **True** evaluations.
#
# #### Using the Boolean Operator ```not```
# The```not``` operator only operates on a single expression, essentially flipping **True** to **False** or **False** to **True**.
# The not operator flips a True to False
not False
# #### Combining Boolean and Comparison Operators
#
# We can combine [Boolean operators](https://docs.constellate.org/key-terms/#boolean-operator) and [comparison operators](https://docs.constellate.org/key-terms/#comparison-operator) to create even more nuanced **Truth** tests.
# Evaluating two conditions with integers at once
(3 < 22) and (60 == 34) # What does each condition evaluate to?
# Evaluating two conditions with integers at once
(3 == 45) or (3 != 7) # What does each condition evaluate to?
# So far, we have evaluated one or two conditions at once, but we could compare even more at once. (In practice, this is rare since it creates code that can be difficult to read.) [Boolean operators](https://docs.constellate.org/key-terms/#boolean-operator) also have an order of operations like mathematical [operators](https://docs.constellate.org/key-terms/#operator). They resolve in the order of `not`, `and`, then `or`.
# ## Writing a Flow Control Statement
#
# The general form of a [flow control statement](https://docs.constellate.org/key-terms/#flow-control-statement) in [Python](https://docs.constellate.org/key-terms/#python) is a condition followed by an action clause:
#
# `In this condition:`<br />
# `perform this action`
#
# Let's return to part of our flowchart for hanging out with friends.
#
# 
#
# We can imagine a [flow control statement](https://docs.constellate.org/key-terms/#flow-control-statement) that would look something like:
#
# `if have_homework == True:`<br />
# `complete assignment`
#
# The condition is given followed by a colon (:). The action clause then follows on the next line, indented into a [code block](https://docs.constellate.org/key-terms/#code-block).
#
# * If the condition is fulfilled (evaluates to **True**), the action clause in the block of code is executed.
# * If the condition is not fulfilled (evaluates to **False**), the action clause in the block of code is skipped over.
# ### Code Blocks
# A [code block](https://docs.constellate.org/key-terms/#code-block) is a snippet of code that begins with an indentation. A [code block](https://docs.constellate.org/key-terms/#code-block) can be a single line or many lines long. Blocks can contain other blocks forming a hierarchal structure. In such a case, the second block is indented an additional degree. Any given block ends when the number of indentations in the current line is less than the number that started the block.
#
# 
#
# Since the level of indentation describes which code block will be executed, improper indentations will make your code crash. When using indentations to create code blocks, look carefully to make sure you are working in the code block you intend. Each indentation for a code block is created by pressing the tab key.
# ## Types of Flow Control Statements
#
# The code example above uses an `if` statement, but there are other kinds of [flow control statements](https://docs.constellate.org/key-terms/#flow-control-statement) available in [Python](https://docs.constellate.org/key-terms/#python).
#
# |Statement|Means|Condition for execution|
# |---|---|---|
# |`if`|if|if the condition is fulfilled|
# |`elif`|else if|if no previous conditions were met *and* this condition is met|
# |`else`|else|if no condition is met (no condition is supplied for an `else` statement)|
# |`while`|while|while condition is true|
# |`for`|for|execute in a loop for this many times|
# |`try`|try|try this and run the `except` code if an error occurs|
#
# Let's take a look at each of these [flow control statement](https://docs.constellate.org/key-terms/#flow-control-statement) types.
# ### `if` Statements
#
# An `if` statement begins with an [expression](https://docs.constellate.org/key-terms/#expression) that evaluates to **True** or **False**.
#
# * if **True**, then perform this action
# * if **False**, skip over this action
#
# In practice, the form looks like this:
#
# `if this is True:` <br />
# `perform this action`
#
# Let's put an `if` statement into practice with a very simple program that asks the user how their day is going and then responds. We can visualize the flow of the program in a flowchart.
#
# 
#
# Our program will use a single `if` statement. If the user types "Yes" or "yes", then our program will send a response.
# +
# A program that responds to a user having a good day
having_good_day = input('Are you having a good day? (Yes or No) ') # Define a variable having_good_day to hold the user's input in a string
if having_good_day == 'Yes' or having_good_day == 'yes': # If the user has input the string 'Yes' or 'yes'
print('Glad to hear your day is going well!') # Print: Glad to hear your day is going well!
# -
# Our program works fairly well so long as the user inputs 'Yes' or 'yes'. If they type 'no' or something else, it simply ends. If we want to have our program still respond, we can use an `else` statement.
# ### `else` Statements
#
# An `else` statement *does not require a condition* to evaluate to **True** or **False**. It simply executes when none of the previous conditions are met. The form looks like this:
#
# `else:` <br />
# `perform this action`
#
# Our updated flowchart now contains a second branch for our program.
#
# 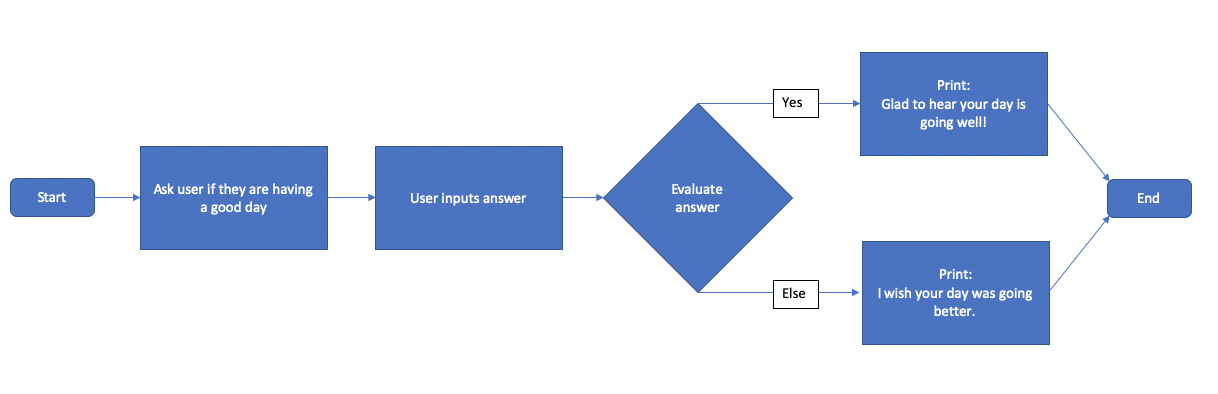
#
#
# +
# A program that responds to whether the user is having a good or bad day
having_good_day = input('Are you having a good day? (Yes or No) ') # Define a variable having_good_day to hold the user's input
if having_good_day == 'Yes' or having_good_day == 'yes': # If the user has input the string 'Yes' or 'yes'
print('Glad to hear your day is going well!') # Print: Glad to hear your day is going well!
# Write an else statement here
# -
# Our new program is more robust. The new `else` statement still gives the user a response if they do not respond "Yes" or "yes". But what if we wanted to add an option for when a user says "No"? Or when a user inputs something besides "Yes" or "No"? We could use a series of `elif` statements.
# ### `elif` Statements
#
# An `elif` statement, short for "else if," allows us to create a list of possible conditions where one (and only one) action will be executed. `elif` statements come after an initial `if` statement and before an `else` statement:
#
# `if condition A is True:` <br />
# `perform action A` <br />
# `elif condition B is True:` <br />
# `perform action B` <br />
# `elif condition C is True:` <br />
# `perform action C` <br />
# `elif condition D is True:` <br />
# `perform action D` <br />
# `else:` <br />
# `perform action E`
#
# For example, we could add an `elif` statement to our program so it responds to both "Yes" and "No" with unique answers. We could then add an `else` statement that responds to any user input that is not "Yes" or "No".
#
# 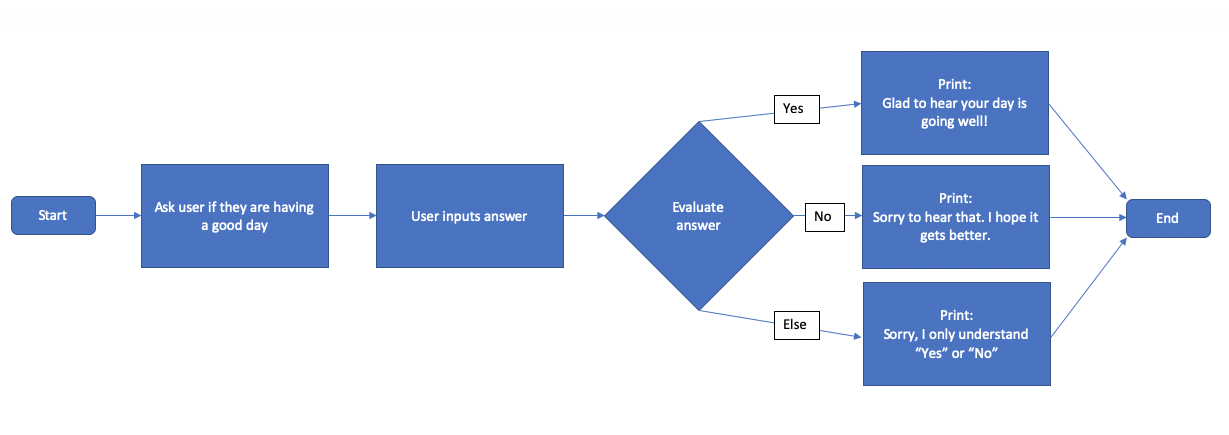
# +
# A program that responds to whether the user is having a good or bad day
having_good_day = input('Are you having a good day? (Yes or No) ') # Define a variable having_good_day to hold the user's input
if having_good_day == 'Yes' or having_good_day == 'yes': # If the user has input the string 'Yes' or 'yes'
print('Glad to hear your day is going well!') # Print: Glad to hear your day is going well!
# Write and elif statement for having_good_day == 'No'
# An else statement that catches if the answer is not 'yes' or 'no'
else: # Execute this if none of the other branches executes
print('Sorry, I only understand "Yes" or "No"') # Note that we can use double quotations in our string because it begins and ends with single quotes
# -
# #### The difference between`elif` and `if`?
#
# When an `elif` condition is met, all other `elif` statements are skipped over. This means that one (and only one) [flow control statement](https://docs.constellate.org/key-terms/#flow-control-statement) is executed when using `elif` statements. The fact that only one `elif` statement is executed is important because it may be possible for multiple [flow control statements](https://docs.constellate.org/key-terms/#flow-control-statement) to evaluate to **True**. A series of `elif` statements evaluates from top-to-bottom, only executing the first `elif` statement whose condition evaluates to **True**. The rest of the `elif` statements are skipped over (whether they are **True** or **False**).
#
# In practice, a set of mutually exclusive `if` statements will result in the same actions as an `if` statement followed by `elif` statements. There are a few good reasons, however, to use `elif` statements:
#
# 1. A series of `elif` statements helps someone reading your code understand that a single flow control choice is being made.
# 2. Using `elif` statements will make your program run faster since other conditional statements are skipped after the first evaluates to **True**. Otherwise, every `if` statement has to be evaluated before the program moves to the next step.
# 3. Writing a mutually exclusive set of `if` statements can be very complex.
#
# Expanding on the concept of our "How is your day going?" program, let's take a look at an example that asks the user "How is your week going?" It will take two inputs: the day of the week (`day_of_week`) and how the user feels the week is going (`having_good_week`).
#
# 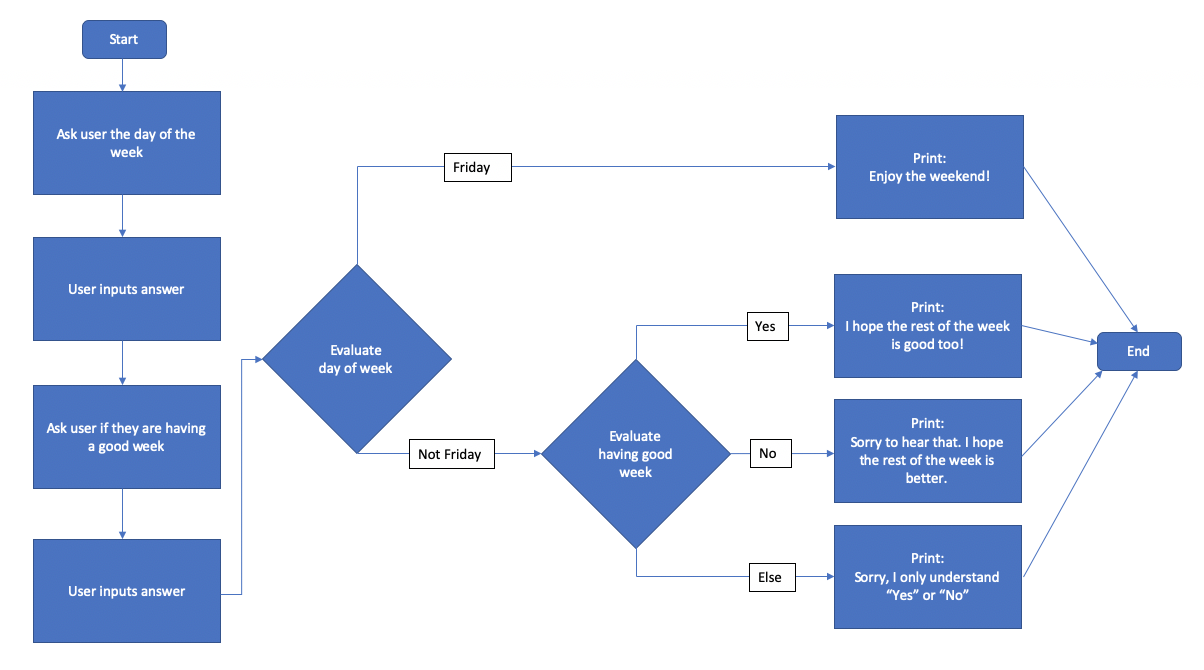
# +
# A program that responds to the user's input for the day of the week and how their week is going.
day_of_week = input('What day of the week is it? ')
having_good_week = input('Are you having a good week? ')
if day_of_week == 'Friday' or day_of_week == 'friday':
print('Enjoy the weekend!')
elif having_good_week == 'Yes' or having_good_week == 'yes':
print('I hope the rest of the week is good too!')
elif having_good_week == 'No' or having_good_week == 'no':
print('Sorry to hear that. I hope the rest of the week is better.')
else:
print('Sorry, I only understand "Yes" or "No"')
# -
# In the program above, try changing the `elif` statements to `if` statements. What happens if the user inputs 'Friday' and 'Yes'?
# ### `while` Loop Statements
#
# So far, we have used [flow control statements](https://docs.constellate.org/key-terms/#flow-control-statement) like decision-making branches to decide what action should be taken next. Sometimes, however, we want a particular action to loop (or repeat) until some condition is met. We can accomplish this with a `while` loop statement that takes the form:
#
# `while condition is True:` <br />
# `take this action`
#
# After the [code block](https://docs.constellate.org/key-terms/#code-block) is executed, the program loops back to check and see if the `while` loop condition has changed from **True** to **False**. The code block stops looping when the condition becomes **False**.
#
# In the following program, the user will guess a number until they get it correct.
#
# 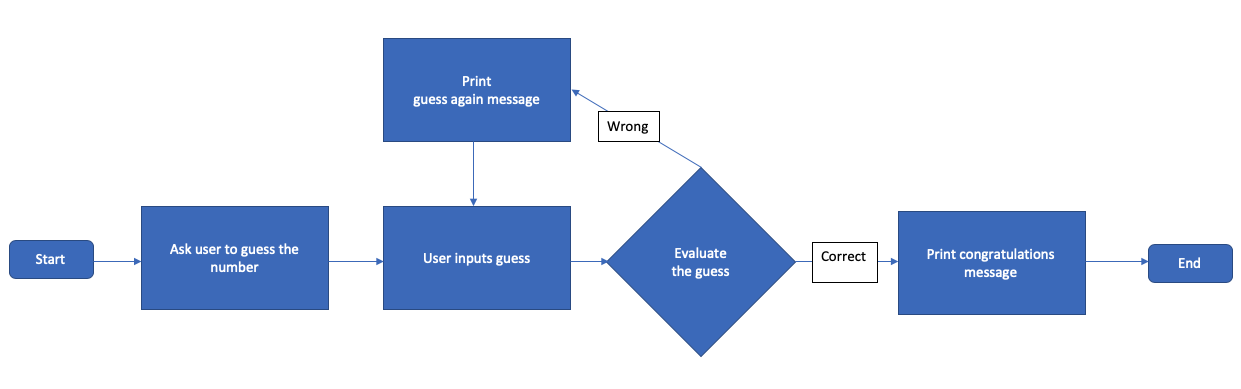
# +
# A program that asks the user to guess a number
# The secret number is set here by the programmer.
secret_number = str(4) # We convert the integer to a string to compare easily with a user input string
# Ask the user to make a guess and take input in a string
guess = input('I am thinking of a number between 1-10. Can you guess it? ') # Take the user's first guess
# Check to see if the user guess matches our secret number
while guess != secret_number: # While the users guess does not equal secret_number
guess = input('Nope. Guess again! ') # Allow the user to change the value of guess
print('You guessed the secret number, ' + secret_number) # Print a congratulations message with the secret number
# -
# #### Stopping Accidental Infinite Loops
# When using a `while` loop, it is possible to accidentally create an infinite loop that never ends. This happens because the `while` condition *never* becomes **False**.
#
# If you accidentally write code that infinitely repeats, you can stop the execution by selecting **Interrupt** from the **Kernel** menu. (Alternatively, you can press the letter **i** twice on your keyboard.) You may also want to remove the output of the rogue cell. You can do this from the **Cell** menu.
# * Clearing output from a single cell: **Cell** ▶ **Current Outputs** ▶ **Clear**
# * Clearing output from all cells: **Cell** ▶ **All Output** ▶ **Clear**
#
# 
# Run this infinite loop then interrupt the kernel
# Can you clear the cell output?
while True:
print('Oh noes!')
# #### A Repeating `while` Loop
#
# In the program above, the `while` loop checked to see if the user guessed a particular number. We could also use a `while` loop to repeat a [code block](https://docs.constellate.org/key-terms/#code-block) a particular number of times.
# A simple program that prints out 1, 2, 3
number = 0
while number < 3:
number = number + 1 # We can also write an equivalent shortcut: number += 1
print(number)
# ### `for` Loop Statements with a `range()` Function
#
# An abbreviated way to write a `while` loop that repeats a specified number of times is using a `for` loop with a `range()` function. This loop takes the form:
#
# `for i in range(j):` <br />
# `take this action`
#
# where `i` is a generic [variable](https://docs.constellate.org/key-terms/#variable) for counting the number of iterations and `j` is the number of times you want the [code block](https://docs.constellate.org/key-terms/#code-block) to repeat.
#
# The starting value of `i` is 0. After each loop, `i` increases by one until it reaches `j`. The loop then stops. The [variable](https://docs.constellate.org/key-terms/#variable) names `i` and `j` are merely conventions. Using a different name may make the purpose of your code clearer to readers.
# +
# A `for` loop that repeats 'What?' three times.
# How could you make this repeat five times?
# What happens if you change the name of variable `i`?
for i in range(3):
print('What?')
# -
# A `for` loop that prints the value of the current iteration, here called `i`.
for i in range(5):
print(i)
# In the examples above, the `range()` [function](https://docs.constellate.org/key-terms/#function) takes a single argument that specifies the number of loops. The assumption is our `range()` [function](https://docs.constellate.org/key-terms/#function) will start at 0, but we can specify any starting [integer](https://docs.constellate.org/key-terms/#integer) by adding an additional [argument](https://docs.constellate.org/key-terms/#argument).
# A `for` loop that starts looping at 27 and stops looping at 32
for i in range(27, 32):
print(i)
# A `for` loop that starts looping at -3 and stops looping at 7
for i in range(-3, 7):
print(i)
# We can also specify the size of each step for the `range()` [function](https://docs.constellate.org/key-terms/#function). In the above examples, the [function](https://docs.constellate.org/key-terms/#function) adds one for each increment step, but we can add larger numbers or even specify negative numbers to have the `range()` [function](https://docs.constellate.org/key-terms/#function) count down. The general form is:
#
# `for i in range(start, stop, increment):` <br />
# `take this action`
# A `for` loop that counts down from ten to one, followed by printing `Go!`
for i in range(10, 0, -1):
print(i)
print('Go!')
# ### `Continue` and `Break` Statements
# `while` loops and `for` loops can also use `continue` and `break` statements to affect flow control.
# * A `continue` statement immediately restarts the loop.
# * A `break` statement immediately exits the loop.
#
# Let's return to our secret number guessing program. We will write the same program workflow using `continue` and `break` statements.
#
# 
# +
# A program that asks the user to guess a number
# Initialize the variables `guess` and `secret_number`
guess = 0
secret_number = str(4) # The secret number is set here by the programmer. Notice it is turned into a string so it can be easily compared with user inputs.
# Ask the user to make a guess
print('I am thinking of a number between 1-10.')
# Test whether the user guess matches `secret_number`
while True:
guess = input('What is your guess? ')
if guess == secret_number:
break
else:
continue
# After loop ends, print a congratulations message with the secret number
print('You guessed the secret number, ' + secret_number)
# -
# ### Exception Handling with `try` and `except`
#
# When running code that may create an error, we can use `try` and `except` statements to stop a program from crashing.
# +
# Try running the first code block, if there's an error then run the `except` code
# Divide 100 by a number the user chooses
user_number = input('100 divided by ... ')
try:
print(100 / int(user_number))
except:
print("You can't divide by zero.")
# -
# ___
# <h2 style="color:red; display:inline">Coding Challenge! < / > </h2>
#
# **Using your knowledge of flow control statements, can you write a program that asks a user for their name, then prints out a response depending on whether they are old enough to drive? Check the end of this notebook for example solutions.**
# ___
# +
# Level 1 Challenge
# Ask the user for their name and store it in a variable `user_name`
# Ask the user for their age and store it in a variable `user_age`
# Write an if, elif, else statement that checks to see if the user is driving age.
# If the user is driving age, print out "user_name is old enough to drive."
# If the user is not driving age, print out "user_name is not old enough to drive."
# +
# Level 2 Challenge
# Improve your flow control to consider whether the user has input a realistic age
# If the user inputs an age over 120, print "Is `user_name` a human or a turtle?"
# If the user inputs an age less than 5, print "`user_name` is not even ready for a bicycle!"
# +
# Level 3 Challenge
# A program that checks to see if a user is old enough to drive.
# Verifies user has input a number and it is realistic.
# Find a solution to address when a user enters text that is not a number.
# -
# ## Functions
#
# We have used several [Python](https://docs.constellate.org/key-terms/#python) [functions](https://docs.constellate.org/key-terms/#function) already, including `print()`, `input()`, and `range()`. You can identify a function by the fact that it ends with a set of parentheses `()` where [arguments](https://docs.constellate.org/key-terms/#argument) can be **passed** into the function. Depending on the [function](https://docs.constellate.org/key-terms/#function) (and your goals for using it), a [function](https://docs.constellate.org/key-terms/#function) may accept no [arguments](https://docs.constellate.org/key-terms/#argument), a single [argument](https://docs.constellate.org/key-terms/#argument), or many [arguments](https://docs.constellate.org/key-terms/#argument). For example, when we use the `print()` [function](https://docs.constellate.org/key-terms/#function), a [string](https://docs.constellate.org/key-terms/#string) (or a variable containing a [string](https://docs.constellate.org/key-terms/#string)) is passed as an [argument](https://docs.constellate.org/key-terms/#argument).
#
# [Functions](https://docs.constellate.org/key-terms/#function) are a convenient shorthand, like a mini-program, that makes our code more **modular**. We don't need to know all the details of how the `print()` [function](https://docs.constellate.org/key-terms/#function) works in order to use it. [Functions](https://docs.constellate.org/key-terms/#function) are sometimes called "black boxes", in that we can put an [argument](https://docs.constellate.org/key-terms/#argument) into the box and a **return value** comes out. We don't need to know the inner details of the "black box" to use it. (Of course, as you advance your programming skills, you may become curious about how certain [functions](https://docs.constellate.org/key-terms/#function) work. And if you work with sensitive data, you may *need* to peer in the black box to ensure the security and accuracy of the output.)
#
# ### Libraries and Modules
#
# While [Python](https://docs.constellate.org/key-terms/#python) comes with many [functions](https://docs.constellate.org/key-terms/#function), there are thousands more that others have written. Adding them all to [Python](https://docs.constellate.org/key-terms/#python) would create mass confusion, since many people could use the same name for [functions](https://docs.constellate.org/key-terms/#function) that do different things. The solution then is that [functions](https://docs.constellate.org/key-terms/#function) are stored in [modules](https://docs.constellate.org/key-terms/#module) that can be **imported** for use. A [module](https://docs.constellate.org/key-terms/#module) is a [Python](https://docs.constellate.org/key-terms/#python) file (extension ".py") that contains the definitions for the [functions](https://docs.constellate.org/key-terms/#function) written in [Python](https://docs.constellate.org/key-terms/#python). These [modules](https://docs.constellate.org/key-terms/#module) (individual [Python](https://docs.constellate.org/key-terms/#python) files) can then be collected into even larger groups called [packages](https://docs.constellate.org/key-terms/#package) and [libraries](https://docs.constellate.org/key-terms/#library). Depending on how many [functions](https://docs.constellate.org/key-terms/#function) you need for the program you are writing, you may import a single [module](https://docs.constellate.org/key-terms/#module), a [package](https://docs.constellate.org/key-terms/#package) of [modules](https://docs.constellate.org/key-terms/#module), or a whole [library](https://docs.constellate.org/key-terms/#library).
#
# The general form of importing a [module](https://docs.constellate.org/key-terms/#module) is:
# `import module_name`
#
# You may recall from the "Getting Started with Jupyter Notebooks" lesson, we imported the `time` [module](https://docs.constellate.org/key-terms/#module) and used the `sleep()` [function](https://docs.constellate.org/key-terms/#function) to wait 5 seconds.
# A program that waits five seconds then prints "Done"
print('Waiting 5 seconds...')
import time # We import the `time` module
time.sleep(5) # We run the sleep() function from the time module using `time.sleep()`
print('Done')
# We can also just import the `sleep()` [function](https://docs.constellate.org/key-terms/#function) without importing the whole `time` [module](https://docs.constellate.org/key-terms/#module).
# A program that waits five seconds then prints "Done"
print('Waiting 5 seconds...')
from time import sleep # We import just the sleep() function from the time module
sleep(5) # Notice that we just call the sleep() function, not time.sleep
print('Done')
# ## Writing a Function
#
# In the above examples, we **called** a [function](https://docs.constellate.org/key-terms/#function) that was already written. To call our own [functions](https://docs.constellate.org/key-terms/#function), we need to define our [function](https://docs.constellate.org/key-terms/#function) first with a **function definition statement** followed by a [code block](https://docs.constellate.org/key-terms/#code-block):
#
# `def my_function():` <br />
# `"""Description what the functions does"""`<br />
# `do this task`
#
#
# After the [function](https://docs.constellate.org/key-terms/#function) is defined, we can **call** on it to do us a favor whenever we need by simply executing the [function](https://docs.constellate.org/key-terms/#function) like so:
#
# `my_function()`
#
# After the function is defined, we can call it as many times as we want without having to rewrite its code. In the example below, we call `my_function` twice.
# +
# Creating a simple function to double a number
def my_function():
"""Doubles any number the user enters"""
outputnumber = int(inputnumber) * 2
print(outputnumber)
inputnumber = input('I will double any number. Give me a number. ')
my_function()
inputnumber = input('Give me a new number. ')
my_function()
inputnumber = input('Give me one last number. ')
my_function()
# -
# Using [functions](https://docs.constellate.org/key-terms/#function) also makes it easier for us to update our code. Let's say we wanted to change our program to square our `inputnumber` instead of doubling it. We can simply change the [function](https://docs.constellate.org/key-terms/#function) definition one time to make the change everywhere. See if you can make the change. (Remember to also change your program description in the first line!)
# +
# Creating a simple function to raise a number to the second power.
def my_function():
"""Squares any number the user enters"""
outputnumber = int(inputnumber) ** 2
print(outputnumber)
inputnumber = input('I will raise any number to the second power. Give me a number. ')
my_function()
inputnumber = input('Give me a new number. ')
my_function()
inputnumber = input('Give me one last number. ')
my_function()
# -
# By changing our [function](https://docs.constellate.org/key-terms/#function) one time, we were able to make our program behave differently in three different places. Generally, it is good practice to avoid duplicating program code to avoid having to change it in multiple places. When programmers edit their code, they may spend time **deduplicating** to make the code easier to read and maintain.
# ### Parameters vs. Arguments
#
# When we write a [function](https://docs.constellate.org/key-terms/#function) definition, we can define a [parameter](https://docs.constellate.org/key-terms/#parameter) to work with the [function](https://docs.constellate.org/key-terms/#function). We use the word [parameter](https://docs.constellate.org/key-terms/#parameter) to describe the [variable](https://docs.constellate.org/key-terms/#variable) in parentheses within a [function](https://docs.constellate.org/key-terms/#function) definition:
#
# `def my_function(input_variable):` <br />
# `do this task`
#
# In the pseudo-code above, `input_variable` is a [parameter](https://docs.constellate.org/key-terms/#parameter) because it is being used within the context of a [function](https://docs.constellate.org/key-terms/#function) *definition*. When we actually call and run our [function](https://docs.constellate.org/key-terms/#function), the actual [variable](https://docs.constellate.org/key-terms/#variable) or value we pass to the [function](https://docs.constellate.org/key-terms/#function) is called an [argument](https://docs.constellate.org/key-terms/#argument).
# +
# A program to greet the user by name
def greeting_function(user_name): #`user_name` here is a parameter since it is in the definition of the `greeting_function`
print('Hello ' + user_name)
greeting_function('Sam') # 'Sam' is an argument that is being passed into the `greeting_function`
# -
# In the above example, we passed a [string](https://docs.constellate.org/key-terms/#string) into our [function](https://docs.constellate.org/key-terms/#function), but we could also pass a [variable](https://docs.constellate.org/key-terms/#variable).
# +
# A program to greet the user by name
def greeting_function(user_name): #`user_name` here is a parameter since it is in the definition of the `greeting_function`
print('Hello ' + user_name)
answer = input('What is your name? ')
greeting_function(answer) # `answer` is an argument that is being passed into the `greeting_function`
# -
# ### Function Return Values
#
# Whether or not a [function](https://docs.constellate.org/key-terms/#function) takes an [argument](https://docs.constellate.org/key-terms/#argument), it will return a value. If we do not specify that return value in our [function](https://docs.constellate.org/key-terms/#function) definition, it is automatically set to `None`, a special value like the Boolean `True` and `False` that simply means null or nothing. (`None` is not the same thing as, say, the integer `0`.) We can also specify return values for our [function](https://docs.constellate.org/key-terms/#function) using a [flow control statement](https://docs.constellate.org/key-terms/#flow-control-statement) followed by `return` in the [code block](https://docs.constellate.org/key-terms/#code-block).
#
# Let's write a function for telling fortunes. We can call it `fortune_picker` and it will accept a number (1-6) then return a string for the fortune.
# +
# A fortune-teller program that contains a function `fortune_picker`
# `fortune_picker` accepts an integer (1-6) and returns a fortune string
def fortune_picker(fortune_number): # A function definition statement that has a parameter `fortune_number`
if fortune_number == 1:
return 'You will have six children.'
elif fortune_number == 2:
return 'You will become very wise.'
elif fortune_number == 3:
return 'A new friend will help you find yourself.'
elif fortune_number == 4:
return 'Do not eat the sushi.'
elif fortune_number == 5:
return 'That promising venture... it is a trap.'
elif fortune_number == 6:
return 'Sort yourself out then find love.'
print(fortune_picker(5))
# -
# In our example, we passed the [argument](https://docs.constellate.org/key-terms/#argument) `3` that returned the [string](https://docs.constellate.org/key-terms/#string) `'A new friend will help you find yourself'`. To change the fortune, we would have to pass a different [integer](https://docs.constellate.org/key-terms/#integer) into the [function](https://docs.constellate.org/key-terms/#function). To make our fortune-teller random, we could import the [function](https://docs.constellate.org/key-terms/#function) `randint()` that chooses a random number between two [integers](https://docs.constellate.org/key-terms/#integer). We pass the two [integers](https://docs.constellate.org/key-terms/#integer) as [arguments](https://docs.constellate.org/key-terms/#argument) separated by a comma.
# +
# A fortune-teller program that uses a random integer
from random import randint # import the randint() function from the random module
def fortune_picker(fortune_number): # A function definition statement that has a parameter `fortune_number`
if fortune_number == 1:
return 'You will have six children.'
elif fortune_number == 2:
return 'You will become very wise.'
elif fortune_number == 3:
return 'A new friend will help you find yourself.'
elif fortune_number == 4:
return 'Do not eat the sushi.'
elif fortune_number == 5:
return 'That promising venture... it is a trap.'
elif fortune_number == 6:
return 'Sort yourself out then find love.'
random_number = randint(1, 6) # Choose a random number between 1 and 6 and assign it to a new variable `random_number`
print(fortune_picker(random_number))
# -
# ### Local and Global Scope
#
# We have seen that [functions](https://docs.constellate.org/key-terms/#function) make maintaining code easier by avoiding duplication. One of the most dangerous areas for duplication is [variable](https://docs.constellate.org/key-terms/#variable) names. As programming projects become larger, the possibility that a [variable](https://docs.constellate.org/key-terms/#variable) will be re-used goes up. This can cause weird errors in our programs that are hard to track down. We can alleviate the problem of duplicate [variable](https://docs.constellate.org/key-terms/#variable) names through the concepts of [local scope](https://docs.constellate.org/key-terms/#local-scope) and [global scope](https://docs.constellate.org/key-terms/#global-scope).
#
# We use the phrase [local scope](https://docs.constellate.org/key-terms/#local-scope) to describe what happens within a [function](https://docs.constellate.org/key-terms/#function). The [local scope](https://docs.constellate.org/key-terms/#local-scope) of a [function](https://docs.constellate.org/key-terms/#function) may contain [local variable](https://docs.constellate.org/key-terms/#local-variable), but once that [function](https://docs.constellate.org/key-terms/#function) has completed the [local variable](https://docs.constellate.org/key-terms/#local-variable) and their contents are erased.
#
# On the other hand, we can also create [global variables](https://docs.constellate.org/key-terms/#global-variable) that persist at the top-level of the program *and* within the [local scope](https://docs.constellate.org/key-terms/#local-scope) of a [function](https://docs.constellate.org/key-terms/#function).
#
# * In the [global scope](https://docs.constellate.org/key-terms/#global-scope), [Python](https://docs.constellate.org/key-terms/#python) does not recognize any [local variable](https://docs.constellate.org/key-terms/#local-variable) from within [functions](https://docs.constellate.org/key-terms/#function)
# * In the [local scope](https://docs.constellate.org/key-terms/#local-scope) of a [function](https://docs.constellate.org/key-terms/#function), [Python](https://docs.constellate.org/key-terms/#python) can recognize and modify any [global variables](https://docs.constellate.org/key-terms/#global-variable)
# * It is possible for there to be a [global variable](https://docs.constellate.org/key-terms/#global-variable) and a [local variable](https://docs.constellate.org/key-terms/#local-variable) with the same name
#
# Ideally, [Python](https://docs.constellate.org/key-terms/#python) programs should limit the number of [global variables](https://docs.constellate.org/key-terms/#global-variable) and create most [variables](https://docs.constellate.org/key-terms/#variable) in a [local scope](https://docs.constellate.org/key-terms/#local-scope).
# +
# Demonstration of global variable being use in a local scope
# The program crashes when a local variable is used in a global scope
global_secret_number = 7
def share_number():
local_secret_number = 13
print(f'The global secret number is {global_secret_number}')
print(f'The local secret number is {local_secret_number} ')
share_number()
print(f'The global secret number is {global_secret_number}')
print(f'The local secret number is {local_secret_number}')
# -
# The code above defines a [global variable](https://docs.constellate.org/key-terms/#global-variable) `global_secret_number` with the value of 7. A [function](https://docs.constellate.org/key-terms/#function), called `share_number`, then defines a [local variable](https://docs.constellate.org/key-terms/#local-variable) `local_secret_number` with a value of 13. When we call the `share_number` [function](https://docs.constellate.org/key-terms/#function), it prints the [local variable](https://docs.constellate.org/key-terms/#local-variable) and the [global variable](https://docs.constellate.org/key-terms/#global-variable). After the `share_number()` [function](https://docs.constellate.org/key-terms/#function) completes we try to print both variables in a [global scope](https://docs.constellate.org/key-terms/#global-scope). The program prints `global_secret_number` but crashes when trying to print `local_secret_number` in a [global scope](https://docs.constellate.org/key-terms/#global-scope).
#
# It's a good practice not to name a [local variable](https://docs.constellate.org/key-terms/#local-variable) the same thing as a [global variable](https://docs.constellate.org/key-terms/#global-variable). If we define a [variable](https://docs.constellate.org/key-terms/#variable) with the same name in a [local scope](https://docs.constellate.org/key-terms/#local-scope), it becomes a [local variable](https://docs.constellate.org/key-terms/#local-variable) within that scope. Once the [function](https://docs.constellate.org/key-terms/#function) is closed, the [global variable](https://docs.constellate.org/key-terms/#global-variable) retains its original value.
# +
# A demonstration of global and local scope using the same variable name
secret_number = 7
def share_number():
secret_number = 10
print(secret_number)
share_number()
print(secret_number)
# -
# ### The Global Statement
#
# A [global statement](https://docs.constellate.org/key-terms/#global-statement) allows us to modify a [global variable](https://docs.constellate.org/key-terms/#global-variable) in a [local scope](https://docs.constellate.org/key-terms/#local-scope).
# +
# Using a global statement in a local scope to change a global variable locally
secret_number = 7
def share_number():
global secret_number # The global statement indicates this the global variable, not a local variable
secret_number = 10
print(secret_number)
share_number()
print(secret_number)
# -
# ___
# ## Lesson Complete
# Congratulations! You have completed *Python Basics 2*. There is only one more lesson in *Python Basics*:
#
# * *Python Basics 3*
#
# ### Start Next Lesson: [Python Basics 3](./python-basics-3.ipynb)
#
# ### Coding Challenge! Solutions
#
# There are often many ways to solve programming problems. Here are a few possible ways to solve the challenges, but there are certainly more!
# +
# Level 1 Solution
# A program that checks to see if a user is old enough to drive.
# Ask the user for their name and store it in a variable `user_name`
# Ask the user for their age and store it in a variable `user_age`
# Write an if, elif, else statement that checks to see if the user is driving age.
# If the user is driving age, print out "user_name is old enough to drive."
# If the user is not driving age, print out "user_name is not old enough to drive."
user_name = input('What is your name? ')
user_age = int(input('And what is your age? '))
if user_age >= 16:
print(f'{user_name} is old enough to drive.')
elif user_age < 16:
print(f'{user_name} is not old enough to drive.')
# +
# Level 2 Solution
# A program that checks to see if a user is old enough to drive.
# Now with improved ability to check if the age number is realistic.
# Improve your flow control to consider whether the user has input a realistic age
# If the user inputs an age over 120, print "Is `user_name` a human or a turtle?"
# If the user inputs an age less than 5, print "`user_name` is not even ready for a bicycle!"
user_name = input('What is your name? ')
user_age = int(input('And what is your age? '))
if user_age > 120:
print(f'Is {user_name} a human or a turtle?')
elif user_age < 5:
print(f'{user_name} is not even ready for a bicycle!')
elif user_age >= 16:
print(f'{user_name} is old enough to drive.')
elif user_age < 16:
print(f'{user_name} is not old enough to drive.')
# +
# Level 3 Solution
# A program that checks to see if a user is old enough to drive.
# Verifies user has input a number and it is realistic.
# Find a solution to address when a user enters text that is not a number.
user_name = input('What is your name? ')
user_age = None
while user_age == None:
try:
user_age = int(input('And what is your age? '))
except:
print('Please input a natural number for your age.')
if user_age > 120:
print(f'Is {user_name} a human or a turtle?')
elif user_age < 5:
print(f'{user_name} is not even ready for a bicycle!')
elif user_age >= 16:
print(f'{user_name} is old enough to drive.')
elif user_age < 16:
print(f'{user_name} is not old enough to drive.')
# +
# Level 3 Solution (using continue/break)
# A program that checks to see if a user is old enough to drive.
# Verifies user has input a number and it is realistic.
# Find a solution to address when a user enters text that is not a number.
user_name = input('What is your name? ')
while True:
user_age = input('What is your age? ')
try:
user_age = int(user_age)
break
except:
print('Please input a number for your age.')
continue
if user_age > 120:
print(f'Is {user_name} a human or a turtle?')
elif user_age < 5:
print(f'{user_name} is not even ready for a bicycle!')
elif user_age >= 16:
print(f'{user_name} is old enough to drive.')
elif user_age < 16:
print(f'{user_name} is not old enough to drive.')
# -
| python-basics-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding: utf-8 -*-
### EXEMPLO DA AULA
### Data: 21-07-2020
### By: <NAME>
# +
import pandas as pd
dataset = pd.read_csv('../00_datasets/credit_data.csv')
# -
media = dataset['age'][dataset['age'] > 0].mean()
dataset.loc[ dataset.age < 0, 'age' ] = media
previsores = dataset.iloc[ :, 1:4 ].values
classe = dataset.iloc[ :, 4 ].values
# +
import numpy as np
from sklearn.impute import SimpleImputer
imputer = SimpleImputer( missing_values=np.nan, strategy='mean' )
imputer = imputer.fit( previsores[:, 0:3] )
previsores[:, 0:3] = imputer.transform( previsores[:, 0:3] )
# +
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
previsores = scaler.fit_transform(previsores)
# +
from sklearn.model_selection import train_test_split
previsores_treinamento, previsores_teste, classe_treinamento, classe_teste = train_test_split( previsores, classe, test_size=0.25,random_state=0 )
# +
from sklearn.neighbors import KNeighborsClassifier
classificador = KNeighborsClassifier( n_neighbors=5, metric='minkowski', p=2 )
classificador.fit(previsores_treinamento, classe_treinamento)
previsoes = classificador.predict( previsores_teste )
# +
from sklearn.metrics import accuracy_score
precisao = accuracy_score( classe_teste, previsoes )
precisao
# -
| 04_aprendizagem-baseada-em-instancias/KNN_credit_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Writing Functions
#
# ---
# ## Questions:
# - How can I create my own functions?
#
# ## Learning Objectives:
# - Explain and identify the difference between function definition and function call.
# - Write a function that takes a small, fixed number of arguments and produces a single result.
#
# ---
# ## Break programs down into functions to make them easier to understand.
#
# * Human beings can only keep a few items in working memory at a time.
# * Understand larger/more complicated ideas by understanding and combining pieces.
# * Components in a machine.
# * Lemmas when proving theorems.
# * Functions serve the same purpose in programs.
# * *Encapsulate* complexity so that we can treat it as a single "thing".
# * Also enables *re-use*.
# * Write one time, use many times.
# ## Define a function using `def` with a name, parameters, and a block of code.
#
# * Begin the definition of a new function with `def`.
# * Followed by the name of the function.
# * Must obey the same rules as variable names.
# * Then *parameters* in parentheses.
# * Empty parentheses if the function doesn't take any inputs.
# * We will discuss this in detail in a moment.
# * Then a colon.
# * Then an indented block of code.
#
# ~~~python
# def print_greeting():
# print('Hello!')
# ~~~
# ## Defining a function does not run it.
#
# * Defining a function does not run it.
# * Like assigning a value to a variable.
# * Must call the function to execute the code it contains.
#
# ~~~python
# print_greeting()
# ~~~
# ## Arguments in call are matched to parameters in definition.
#
# * Functions are most useful when they can operate on different data.
# * Specify *parameters* when defining a function.
# * These become variables when the function is executed.
# * Are assigned the arguments in the call (i.e., the values passed to the function).
# * If you don't name the arguments when using them in the call, the arguments will be matched to parameters in the order the parameters are defined in the function.
#
# ~~~python
# def print_date(year, month, day):
# joined = str(year) + '/' + str(month) + '/' + str(day)
# print(joined)
#
# print_date(1871, 3, 19)
# ~~~
# Or, we can name the arguments when we call the function, which allows us to
# specify them in any order:
# ~~~python
# print_date(month=3, day=19, year=1871)
# ~~~
# * Via [Twitter](https://twitter.com/minisciencegirl/status/693486088963272705):
# `()` contains the ingredients for the function while the body contains the recipe.
#
# ## Functions may return a result to their caller using `return`.
#
# * Use `return ...` to give a value back to the caller.
# * May occur anywhere in the function.
# * But functions are easier to understand if `return` occurs:
# * At the start to handle special cases.
# * At the very end, with a final result.
#
# ~~~python
# def average(values):
# if len(values) == 0:
# return None
# return sum(values) / len(values)
# ~~~
# ~~~python
# a = average([1, 3, 4])
# print('average of actual values:', a)
# ~~~
# ~~~python
# print('average of empty list:', average([]))
# ~~~
# * Remember: [every function returns something]({{ page.root }}/04-built-in/).
# * A function that doesn't explicitly `return` a value automatically returns `None`.
#
# ~~~python
# result = print_date(1871, 3, 19)
# print('result of call is:', result)
# ~~~
#
# ## Identifying Syntax Errors
#
# 1. Read the code below and try to identify what the errors are
# *without* running it.
# 2. Run the code and read the error message.
# Is it a `SyntaxError` or an `IndentationError`?
# 3. Fix the error.
# 4. Repeat steps 2 and 3 until you have fixed all the errors.
#
# ~~~python
# def another_function
# print("Syntax errors are annoying.")
# print("But at least python tells us about them!")
# print("So they are usually not too hard to fix.")
# ~~~
# +
### BEGIN SOLUTION
def another_function():
print("Syntax errors are annoying.")
print("But at least Python tells us about them!")
print("So they are usually not too hard to fix.")
### END SOLUTION
# -
# ## Definition and Use
#
# What does the following program print?
#
# ~~~python
# def report(pressure):
# print('pressure is', pressure)
#
# print('calling', report, 22.5)
# ~~~
# A function call always needs parenthesis, otherwise you get memory address of the function object. So, if we wanted to call the function named report, and give it the value 22.5 to report on, we could have our function call as follows
# ~~~python
# print("calling")
# report(22.5)
# ~~~
# ## Order of Operations
#
# 1. What's wrong in this example?
#
# ~~~python
# result = print_date(1871,3,19)
#
# def print_date(year, month, day):
# joined = str(year) + '/' + str(month) + '/' + str(day)
# print(joined)
# ~~~
# 2. After fixing the problem above, explain why running this example code:
#
# ~~~python
# result = print_date(1871, 3, 19)
# print('result of call is:', result)
# ~~~
#
# Gives this output:
#
# ~~~python
# 1871/3/19
# result of call is: None
# ~~~
#
# Why is the result of the call `None`?
# ## Solution
#
# 1. The problem with the example is that the function `print_date()` is defined *after* the call to the function is made. Python
# doesn't know how to resolve the name `print_date` since it hasn't been defined yet and will raise a `NameError` e.g.,
# `NameError: name 'print_date' is not defined`
#
# 2. The first line of output (`1871/3/19`) is from the print function inside `print_date()`, while the second line
# is from the print function below the function call. All of the code inside `print_date()` is executed first, and
# the program then "leaves" the function and executes the rest of the code.
#
# 3. `print_date()` doesn't explicitly `return` a value, so it automatically returns `None`.
#
# ## Encapsulation
#
# Fill in the blanks to create a function that takes a single filename as an argument,
# loads the data in the file named by the argument,
# and returns the minimum value in that data.
#
# ~~~python
# import pandas as pd
#
# def min_in_data(____):
# data = ____
# return ____
# ~~~
# +
### BEGIN SOLUTION
import pandas as pd
def min_in_data(filename):
data = pd.read_csv(filename)
return data.min()
### END SOLUTION
# -
# ## Find the First
#
# Fill in the blanks to create a function that takes a list of numbers as an argument
# and returns the first negative value in the list.
# What does your function do if the list is empty?
#
# ~~~python
# def first_negative(values):
# for v in ____:
# if ____:
# return ____
# ~~~
# +
### BEGIN SOLUTION
def first_negative(values):
for v in values:
if v<0:
return v
### END SOLUTION
# -
# If an empty list is passed to this function, it returns `None`:
# ~~~python
# my_list = []
# print(first_negative(my_list))
# ~~~
# ## Calling by Name
#
# Earlier we saw this function:
#
# ~~~python
# def print_date(year, month, day):
# joined = str(year) + '/' + str(month) + '/' + str(day)
# print(joined)
# ~~~
#
# We saw that we can call the function using *named arguments*, like this:
#
# ~~~python
# print_date(day=1, month=2, year=2003)
# ~~~
#
# 1. What does `print_date(day=1, month=2, year=2003)` print?
# 2. When have you seen a function call like this before?
# 3. When and why is it useful to call functions this way?
# +
### BEGIN SOLUTION
# 1. `2003/2/1`
# 2. We saw examples of using *named arguments* when working with the pandas library. For example, when reading in a dataset
# using `data = pd.read_csv('data/gapminder_gdp_europe.csv', index_col='country')`, the last argument `index_col` is a
# named argument.
# 3. Using named arguments can make code more readable since one can see from the function call what name the different arguments have inside the function. It can also reduce the chances of passing arguments in the wrong order, since by using named arguments the order doesn't matter.
### END SOLUTION
# -
# ## Encapsulation of an If/Print Block
#
# The code below will run on a label-printer for chicken eggs. A digital scale will report a chicken egg mass (in grams)
# to the computer and then the computer will print a label.
#
# Please re-write the code so that the if-block is folded into a function.
#
# ~~~python
# import random
#
# for i in range(10):
# # simulating the mass of a chicken egg
# # the (random) mass will be 70 +/- 20 grams
# mass = 70 + 20.0 * (2.0 * random.random() - 1.0)
#
# print(mass)
#
# # egg sizing machinery prints a label
# if mass >= 85:
# print("jumbo")
# elif mass >= 70:
# print("large")
# elif mass < 70 and mass >= 55:
# print("medium")
# else:
# print("small")
# ~~~
# The simplified program follows. What function definition will make it functional?
#
# ~~~python
# # revised version
# import random
# for i in range(10):
# # simulating the mass of a chicken egg
# # the (random) mass will be 70 +/- 20 grams
# mass = 70 + 20.0 * (2.0 * random.random() - 1.0)
#
# print(mass, print_egg_label(mass))
#
# ~~~
# 1. Create a function definition for `print_egg_label()` that will work with the revised program above. Note, the function's return value will be significant. Sample output might be `71.23 large`.
# 2. A dirty egg might have a mass of more than 90 grams, and a spoiled or broken egg will probably have a mass that's less than 50 grams. Modify your `print_egg_label()` function to account for these error conditions. Sample output could be `25 too light, probably spoiled`.
# +
### BEGIN SOLUTION
def print_egg_label(mass):
#egg sizing machinery prints a label
if mass >= 90:
return "warning: egg might be dirty"
elif mass >= 85:
return "jumbo"
elif mass >= 70:
return "large"
elif mass < 70 and mass >= 55:
return "medium"
elif mass < 50:
return "too light, probably spoiled"
else:
return "small"
### END SOLUTION
# -
# ## Encapsulating Data Analysis
#
# Assume that the following code has been executed:
#
# ~~~python
# import pandas as pd
#
# df = pd.read_csv('data/gapminder_gdp_asia.csv', index_col=0)
# japan = df.loc['Japan']
# ~~~
#
# 1. Complete the statements below to obtain the average GDP for Japan
# across the years reported for the 1980s.
#
# ~~~python
# year = 1983
# gdp_decade = 'gdpPercap_' + str(year // ____)
# avg = (japan.loc[gdp_decade + ___] + japan.loc[gdp_decade + ___]) / 2
# ~~~
### BEGIN SOLUTION
year = 1983
gdp_decade = 'gdpPercap_' + str(year // 10)
avg = (japan.loc[gdp_decade + '2'] + japan.loc[gdp_decade + '7']) / 2
### END SOLUTION
# 2. Abstract the code above into a single function.
#
# ~~~python
# def avg_gdp_in_decade(country, continent, year):
# df = pd.read_csv('data/gapminder_gdp_'+___+'.csv',delimiter=',',index_col=0)
# ____
# ____
# ____
# return avg
# ~~~
# +
### BEGIN SOLUTION
def avg_gdp_in_decade(country, continent, year):
df = pd.read_csv('data/gapminder_gdp_' + continent + '.csv', index_col=0)
c = df.loc[country]
gdp_decade = 'gdpPercap_' + str(year // 10)
avg = (c.loc[gdp_decade + '2'] + c.loc[gdp_decade + '7'])/2
return avg
### END SOLUTION
# -
# 3. How would you generalize this function
# if you did not know beforehand which specific years occurred as columns in the data?
# For instance, what if we also had data from years ending in 1 and 9 for each decade?
# (Hint: use the columns to filter out the ones that correspond to the decade,
# instead of enumerating them in the code.)
# +
### BEGIN SOLUTION
def avg_gdp_in_decade(country, continent, year):
df = pd.read_csv('data/gapminder_gdp_' + continent + '.csv', index_col=0)
c = df.loc[country]
gdp_decade = 'gdpPercap_' + str(year // 10)
total = 0.0
num_years = 0
for yr_header in c.index: # c's index contains reported years
if yr_header.startswith(gdp_decade):
total = total + c.loc[yr_header]
num_years = num_years + 1
return total/num_years
### END SOLUTION
# -
# The function can now be called by:
#
# ~~~python
# avg_gdp_in_decade('Japan','asia',1983)
# ~~~
# ## Simulating a dynamical system
#
# In mathematics, a [dynamical system](https://en.wikipedia.org/wiki/Dynamical_system) is a system
# in which a function describes the time dependence of a point in a geometrical space. A canonical
# example of a dynamical system is the [logistic map](https://en.wikipedia.org/wiki/Logistic_map),
# a growth model that computes a new population density (between 0 and 1) based on the current
# density. In the model, time takes discrete values 0, 1, 2, ...
#
# 1. Define a function called `logistic_map` that takes two inputs: `x`, representing the current
# population (at time _t_), and a parameter `r=1`. This function should return a value
# representing the state of the system (population) at time _t_+1, using the mapping function: $x(t+1) = r * x(t) * [1 - x(t)]$
# +
### BEGIN SOLUTION
def logistic_map(x, r):
return r * x * (1 - x)
### END SOLUTION
# -
# 2. Using a `for` or `while` loop, iterate the `logistic_map` function defined in part 1, starting
# from an initial population of 0.5, for a period of time `t_final=10`. Store the intermediate
# results in a list so that after the loop terminates you have accumulated a sequence of values
# representing the state of the logistic map at times _t_=0,1,...,_t_final_. Print this list to
# see the evolution of the population.
# +
### BEGIN SOLUTION
initial_population = 0.5
t_final = 10
r = 1.0
population = [initial_population]
for t in range(1, t_final):
population.append( logistic_map(population[t-1], r) )
### END SOLUTION
# -
# 3. Encapsulate the logic of your loop into a function called `iterate` that takes the initial
# population as its first input, the parameter `t_final` as its second input and the parameter
# `r` as its third input. The function should return the list of values representing the state of
# the logistic map at times _t_=0,1,...,_t_final_. Run this function for periods `t_final=100`
# and `1000` and print some of the values. Is the population trending toward a steady state?
# +
### BEGIN SOLUTION
def iterate(initial_population, t_final, r):
population = [initial_population]
for t in range(1, t_final):
population.append( logistic_map(population[t-1], r) )
return population
for period in (10, 100, 1000):
population = iterate(0.5, period, 1)
print(population[-1])
### END SOLUTION
# + [markdown] jupyter={"outputs_hidden": false}
# ---
# ## Summary of Key Points:
# - Break programs down into functions to make them easier to understand.
# - Define a function using `def` with a name, parameters, and a block of code.
# - Defining a function does not run it.
# - Arguments in call are matched to parameters in definition.
# - Functions may return a result to their caller using `return`.
# -
# ---
# This lesson is adapted from the [Software Carpentry](https://software-carpentry.org/lessons/) [Plotting and Programming in Python](http://swcarpentry.github.io/python-novice-gapminder/) workshop.
#
# Licensed under [CC-BY 4.0](https://creativecommons.org/licenses/by/4.0/) 2021 by [SURGE](https://github.com/surge-dalhousie)
| session_next/writing-functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# import necessary packages
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
from mpl_finance import candlestick_ohlc
# for offline plotly
#from plotly import tools
#import plotly.offline as py
#py.init_notebook_mode(connected=True)
#import plotly.graph_objs as go
# for online plotly
import plotly
plotly.tools.set_credentials_file(username='luceven', api_key='xos4ZlMxDvmYt7zNlouS')
import plotly.plotly as py
import plotly.graph_objs as go
from plotly import tools
import seaborn as sns
# default figure size
#sns.set()
#sns.set(rc={"figure.figsize": (12, 6)})
# default style
sns.set_style('whitegrid')
import datetime as dt
# %matplotlib inline
# -
# read in raw data
raw_data = pd.read_csv("./data/crypto-markets.csv")
raw_data.head(20)
# +
# since slug, symbol, and name represent the same thing, we drop slug and symbol
mydata = raw_data.drop(['slug', 'symbol'], axis = 1)
# change date from string to datetime
mydata['date'] = pd.to_datetime(mydata['date'], format='%Y-%m-%d')
# -
mydata.head(20)
# take out the first three ranked data
btc_data = mydata[mydata['ranknow'] == 1]
other_t10_sum = mydata[(mydata['ranknow'] >= 2) & (mydata['ranknow'] <= 10)].groupby('date', as_index=False).sum()
# +
# plot ohlc of Bitcoin VS sum of all other top 10 cryptocurrency
# ref: https://plot.ly/python/ohlc-charts/
def plot_ohlc(data1, data2):
trace0 = go.Ohlc(x = data1.date,
open = data1.open,
high = data1.high,
low = data1.low,
close = data1.close,
name = 'Bitcoin')
trace1 = go.Ohlc(x = data2.date,
open = data2.open,
high = data2.high,
low = data2.low,
close = data2.close,
name = 'Other Top 10',
increasing=dict(line=dict(color= '#17BECF')),
decreasing=dict(line=dict(color= '#7F7F7F')))
data = [trace0, trace1]
py.iplot(data, filename = 'OHLC for Bitcoin VS ALL Other')
plot_ohlc(btc_data, other_t10_sum)
# +
# we can tell that nearly all the dramatic changes start at 2017.
recent_data = mydata[mydata['date'] >= dt.date(2017, 1, 1)]
# take average of high low and close
recent_data['hlc_average'] = (recent_data['high'] + recent_data['low'] + recent_data['close']) / 3
# +
# Bitcoin VS others
btc_mean = recent_data[recent_data['ranknow'] == 1]
other_t10_mean = recent_data[(recent_data['ranknow'] >= 2) & (recent_data['ranknow'] <= 10)].groupby('date', as_index=False).mean()
minor_mean = recent_data[recent_data['ranknow'] > 10].groupby('date', as_index=False).mean()
# plot price and volume
def plot_price_vol_compare(data1, data2, data3):
fig = tools.make_subplots(rows=1, cols=2, subplot_titles=(
'Crypto Currency Price', 'Transaction Volume'
))
trace0 = go.Scatter(x=data1['date'], y=data1['hlc_average'], name='Bitcoin')
fig.append_trace(trace0, 1, 1)
trace1 = go.Scatter(x=data1['date'], y=data1['volume'], name='Bitcoin')
fig.append_trace(trace1, 1, 2)
trace2 = go.Scatter(x=data2['date'], y=data2['hlc_average'], name='Others')
fig.append_trace(trace2, 1, 1)
trace3 = go.Scatter(x=data2['date'], y=data2['volume'], name='Others')
fig.append_trace(trace3, 1, 2)
trace4 = go.Scatter(x=data3['date'], y=data3['hlc_average'], name='Minor ones')
fig.append_trace(trace4, 1, 1)
trace5 = go.Scatter(x=data3['date'], y=data3['volume'], name='Minor ones')
fig.append_trace(trace5, 1, 2)
fig['layout'].update(title='BitCoin vs others')
fig['layout'].update(showlegend=False)
fig['layout']['yaxis1'].update(title='USD')
fig['layout']['yaxis2'].update(title='Transactions')
fig['layout']['xaxis1'].update(nticks=6)
fig['layout']['xaxis2'].update(nticks=6)
py.iplot(fig, filename='bitcoin-vs-others')
plot_price_vol_compare(btc_mean, other_t10_mean, minor_mean)
# +
# Look at Bitcoin specificly
currency = recent_data[recent_data['name'] == 'Bitcoin'].copy()
def plot_candlestick(data):
increasing_color = '#17BECF'
decreasing_color = '#7F7F7F'
plot_data = []
layout = {
'xaxis': {
'rangeselector': {
'visible': True
}
},
# Adding a volume bar chart for candlesticks is a good practice usually
'yaxis': {
'domain': [0, 0.2],
'showticklabels': False
},
'yaxis2': {
'domain': [0.2, 0.8]
},
'legend': {
'orientation': 'h',
'y': 0.9,
'yanchor': 'bottom'
},
'margin': {
't': 40,
'b': 40,
'r': 40,
'l': 40
}
}
# Defining main chart
trace0 = go.Candlestick(
x=data['date'], open=data['open'], high=data['high'],
low=data['low'], close=data['close'],
yaxis='y2', name='Bitcoin',
increasing=dict(line=dict(color=increasing_color)),
decreasing=dict(line=dict(color=decreasing_color)),
)
plot_data.append(trace0)
# Adding some range buttons to interact
rangeselector = {
'visible': True,
'x': 0,
'y': 0.8,
'buttons': [
{'count': 1, 'label': 'reset', 'step': 'all'},
{'count': 6, 'label': '6 mo', 'step': 'month', 'stepmode': 'backward'},
{'count': 3, 'label': '3 mo', 'step': 'month', 'stepmode': 'backward'},
{'count': 1, 'label': '1 mo', 'step': 'month', 'stepmode': 'backward'},
]
}
layout['xaxis'].update(rangeselector=rangeselector)
# Setting volume bar chart colors
colors = []
for i, _ in enumerate(data['date']):
if i != 0:
if data['close'].iloc[i] > data['close'].iloc[i-1]:
colors.append(increasing_color)
else:
colors.append(decreasing_color)
else:
colors.append(decreasing_color)
trace1 = go.Bar(
x=data['date'], y=data['volume'],
marker=dict(color=colors),
yaxis='y', name='Volume'
)
plot_data.append(trace1)
# Adding Moving Average
def moving_average(interval, window_size=10):
window = np.ones(int(window_size)) / float(window_size)
return np.convolve(interval, window, 'same')
trace2 = go.Scatter(
x=data['date'][5:-5], y=moving_average(data['close'])[5:-5],
yaxis='y2', name='Moving Average',
line=dict(width=1)
)
plot_data.append(trace2)
# Adding boilinger bands
def bollinger_bands(price, window_size=10, num_of_std=5):
rolling_mean = price.rolling(10).mean()
rolling_std = price.rolling(10).std()
upper_band = rolling_mean + (rolling_std * 5)
lower_band = rolling_mean - (rolling_std * 5)
return upper_band, lower_band
bb_upper, bb_lower = bollinger_bands(data['close'])
trace3 = go.Scatter(
x=data['date'], y=bb_upper,
yaxis='y2', line=dict(width=1),
marker=dict(color='#ccc'), hoverinfo='none',
name='Bollinger Bands',
legendgroup='Bollinger Bands'
)
plot_data.append(trace3)
trace4 = go.Scatter(
x=data['date'], y=bb_lower,
yaxis='y2', line=dict(width=1),
marker=dict(color='#ccc'), hoverinfo='none',
name='Bollinger Bands', showlegend=False,
legendgroup='Bollinger Bands'
)
plot_data.append(trace4)
fig = go.Figure(data=plot_data, layout=layout)
py.iplot(fig, filename='Bitcoin-candlestick')
plot_candlestick(currency)
# -
# some exploration by matplotlib and seaborn
market_data = mydata
# create two column that has unit in billions
market_data['market_billion'] = market_data['market'] / 1000000000
market_data['volume_million'] = market_data['volume'] / 1000000000
# +
# pairplot of Bitcoin
def plot_pair(df):
dfBTC=df[df['name']== 'Bitcoin']
df2=dfBTC.drop(['ranknow','high','low','close_ratio','spread'],axis=1)
df2['date'] = pd.to_datetime(df2.date)
df2.set_index('date', inplace=True)
sns.pairplot(df2[['close','volume','market']])
plt.savefig("./plots/pairplot_btc.jpg")
plot_pair(market_data)
# -
# Let's prepare one dataframe where we will observe closing prices for each currency
wide_format = market_data.groupby(['date', 'name'])['close'].last().unstack()
# +
# plot market cap for top 10 cryptocurrencies
def plot_market_cap(data):
plt.figure(figsize = (10, 8))
ax = data.groupby(['name'])['market_billion'].last().sort_values(ascending=False).head(10).sort_values().plot(kind='barh')
ax.set_xlabel("Market cap (in Billion USD)")
plt.title("Top 10 Currencies by Market Cap")
plt.savefig("./plots/market_cap.jpg")
plot_market_cap(market_data)
# +
# plot the transaction volumn
def plot_trans_vol(data):
plt.figure(figsize = (10, 8))
ax = data.groupby(['name'])['volume_million'].last().sort_values(ascending=False).head(10).sort_values().plot(kind='barh')
ax.set_xlabel("Transaction Volume (in Millions)")
plt.title("Top 10 Currencies by Transaction Volume")
plt.savefig("./plots/transaction_volumn.jpg")
plot_trans_vol(market_data)
# +
# Look at top 10 currency
def top_10_trend(data):
top_10_currency_names = data.groupby(['name'])['market'].last().sort_values(ascending=False).head(10).index
data_top_10_currencies = data[data['name'].isin(top_10_currency_names)]
plt.rcParams['figure.figsize']=(20, 12)
ax = data_top_10_currencies.groupby(['date', 'name'])['close'].mean().unstack().plot()
ax.set_ylabel("Price per unit (in USD)")
ax.set_xlabel("Year")
plt.title("Price per unit of currency")
plt.savefig("./plots/top_10_price_unit.jpg")
top_10_trend(market_data)
# +
# From obove result we can see compare to the top3, the other cryptocurrencies has not that much impact.
# We will only look at top 3 from now on
def top_3_market_cap(data):
top_3_currency_names = data.groupby(['name'])['market'].last().sort_values(ascending=False).head(3).index
data_top_3_currencies = data[data['name'].isin(top_3_currency_names)]
plt.rcParams['figure.figsize']=(20, 12)
ax = data_top_3_currencies.groupby(['date', 'name'])['market_billion'].mean().unstack().plot()
ax.set_xlabel("Year")
ax.set_ylabel("Market Cap (in billion USD)")
plt.title("Market cap per Currency")
plt.savefig("./plots/top_3_market_cap.jpg")
top_3_market_cap(market_data)
# +
# Like the previous observation, we are interested in the trends after 2017
def plot_trend_recent(data):
top_3_currency_names = data.groupby(['name'])['market'].last().sort_values(ascending=False).head(3).index
data_top_3_currencies = data[data['name'].isin(top_3_currency_names)]
plt.rcParams['figure.figsize']=(12, 6)
ax = data_top_3_currencies[data_top_3_currencies.date.dt.year >= 2017].groupby(['date', 'name'])['close'].mean().unstack().plot()
ax.set_xlabel("Month")
ax.set_ylabel("Price per unit (in USD)")
plt.title("Price per unit of currency (from 2017)")
plt.savefig("./plots/top_3_trend_recent.jpg")
plot_trend_recent(market_data)
# +
def plot_market_cap_recent(data):
top_3_currency_names = data.groupby(['name'])['market'].last().sort_values(ascending=False).head(3).index
data_top_3_currencies = data[data['name'].isin(top_3_currency_names)]
plt.rcParams['figure.figsize']=(12, 6)
ax = data_top_3_currencies[data_top_3_currencies.date.dt.year >= 2017].groupby(['date', 'name'])['volume'].mean().unstack().plot()
ax.set_xlabel("Month")
ax.set_ylabel("Market Cap (in billion USD)")
plt.title("Market cap per Currency (from 2017)")
plt.savefig("./plots/market_cap_recent.jpg")
plot_market_cap_recent(market_data)
# +
# find the correlation between cryptocurrencies
def plot_corr_top10(data):
top_10_currency_names = data.groupby(['name'])['market'].last().sort_values(ascending=False).head(10).index
data_top_10_currencies = data[data['name'].isin(top_10_currency_names)]
plt.figure(figsize=(12, 12))
sns.heatmap(wide_format[top_10_currency_names].corr(),vmin=0, vmax=1, cmap='coolwarm', annot=True)
plt.title("Correlation between Top 10 Cryptocurrencies")
plt.savefig("./plots/corr_top_10.jpg")
plot_corr_top10(market_data)
# -
# find market data for each year
market_2015 = market_data[(market_data['date'] >= dt.date(2015, 1, 1)) & (market_data['date'] <= dt.date(2015, 12, 31))]
market_2016 = market_data[(market_data['date'] >= dt.date(2016, 1, 1)) & (market_data['date'] <= dt.date(2016, 12, 31))]
market_2017 = market_data[(market_data['date'] >= dt.date(2017, 1, 1)) & (market_data['date'] <= dt.date(2017, 12, 31))]
market_2018 = market_data[(market_data['date'] >= dt.date(2018, 1, 1)) & (market_data['date'] <= dt.date(2018, 12, 31))]
# plot pie chart of market share
def plot_pie_chart(data):
market_val = pd.DataFrame({'market_share' : data.groupby(["name"])['market_billion'].mean().sort_values(ascending=False)}).reset_index()
market_sum = market_val['market_share'].sum()
plot_data = market_val.head(10)
other_share = market_sum - plot_data['market_share'].sum()
plot_data.loc[10] = ['All Other', other_share]
#labels = plot_data.sort_values(ascending=False).index
#values = plot_data.sort_values(ascending=False)
labels = plot_data['name']
values = plot_data['market_share']
trace = go.Pie(labels=labels, values=values)
return labels, values, trace
#py.iplot([trace], filename='basic_pie_chart')
def subplot_pie_chart(data1, data2, data3, data4):
label1, value1, trace1 = plot_pie_chart(data1)
label2, value2, trace2 = plot_pie_chart(data2)
label3, value3, trace3 = plot_pie_chart(data3)
label4, value4, trace4 = plot_pie_chart(data4)
fig = {
'data': [
{
'labels': label1,
'values': value1,
'type': 'pie',
'name': 'Market Share 2015',
'domain': {'x': [0, .48],
'y': [.51, 1]},
'hoverinfo':'label+percent+name',
},
{
'labels': label2,
'values': value2,
'type': 'pie',
'name': 'Market Share 2016',
'domain': {'x': [.52, 1],
'y': [.51, 1]},
'hoverinfo':'label+percent+name',
},
{
'labels': label3,
'values': value3,
'type': 'pie',
'name': 'Market Share 2017',
'domain': {'x': [0, .48],
'y': [0, .49]},
'hoverinfo':'label+percent+name',
},
{
'labels': label4,
'values': value4,
'type': 'pie',
'name':'Market Share 2018',
'domain': {'x': [.52, 1],
'y': [0, .49]},
'hoverinfo':'label+percent+name',
}
],
}
py.iplot(fig, filename='pie_chart_subplots')
subplot_pie_chart(market_2015, market_2016, market_2017, market_2018)
# +
#from datetime import datetime, timedelta
# ref: https://www.kaggle.com/taniaj/cryptocurrency-market-analysis
# Bitcoin price change recent 10 month
def plot_btc_price_change(df):
# Candlestick chart for Rank 1 currency (limited to 10 Months)
rank = 1
months = 10
name = df[df.ranknow == rank].iloc[-1]['name']
filtered_df = df[(df['ranknow'] == rank) & (df['date'] > (max(df['date']) - dt.timedelta(days=30*months)))]
OHLCfiltered_df = filtered_df[['date','open','high','low','close']]
OHLCfiltered_df['date'] = mdates.date2num(OHLCfiltered_df['date'].dt.date)
f,ax=plt.subplots(figsize=(18,11))
ax.xaxis_date()
candlestick_ohlc(ax, OHLCfiltered_df.values, width=0.5, colorup='g', colordown='r',alpha=0.75)
plt.xlabel("Date")
ax.xaxis.set_major_formatter(mdates.DateFormatter('%Y-%m-%d'))
plt.gcf().autofmt_xdate()
plt.title(name + " price")
plt.ylabel("Price")
plt.savefig("./plots/candlestick_btc.jpg")
plt.show()
plot_btc_price_change(market_data)
# +
# find out moving averages of Bitcoin
def plot_moving_avg(df):
# Moving average chart for Rank 1 currency (10 months)
rank = 1
months = 10
name = df[df.ranknow == rank].iloc[-1]['name']
filtered_df = df[(df['ranknow'] == rank) & (df['date'] > (max(df['date']) - dt.timedelta(days=30*months)))]
filtered_df.set_index('date', inplace=True)
f, ax = plt.subplots(figsize=(18,11))
filtered_df.close.plot(label='Raw', ax=ax)
filtered_df.close.rolling(20).mean().plot(label='20D MA', ax=ax)
filtered_df.close.ewm(alpha=0.03).mean().plot(label='EWMA($\\alpha=.03$)', ax=ax)
plt.title(name + " price with Moving Averages")
plt.legend()
plt.xlabel("Month")
plt.gcf().autofmt_xdate()
plt.ylabel("Close ($)")
plt.savefig("./plots/moving_avg_btc.jpg")
plt.show()
plot_moving_avg(market_data)
# -
# plot pie chart for market share of 2017
plot_pie_chart(market_2015)
# plot pie chart for market share of 2016
plot_pie_chart(market_2016)
# plot pie chart for market share of 2017
plot_pie_chart(market_2017)
# plot pie chart for market share of 2018
plot_pie_chart(market_2018)
| HW8/.ipynb_checkpoints/crypto_market-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # fGSEA benchmark
#
# This jupyter notebook calculates the speed of fGSEA as well as the p-value consistency. We install the newest version of fGSEA from github.
library(devtools)
install_github("ctlab/fgsea")
# +
library(fgsea)
library(data.table)
library(ggplot2)
library(BiocParallel)
register(SerialParam())
# -
sig = read.table("test/aging_muscle_gtex.tsv", sep=",", header=T)
signature = sig[,2]
names(signature) = sig[,1]
# ## Initializing gene set library
#
# The gene set library can be downloaded from Enrichr.
ll = readLines("test/GO_Biological_Process_2021.txt")
genes = list()
gmt = list()
for(l in ll){
sp = unlist(strsplit(l, "\t"))
pname = sp[1]
gmt[[pname]] = sp[3:length(sp)]
}
# ## Run fGSEA
# +
st = Sys.time()
fgseaRes <- fgsea(pathways = gmt,
stats = signature,
eps = 0.0,
minSize = 5,
maxSize = 4000)
print(Sys.time() - st)
# -
# # Consistency
#
# Compare consistency of log p-values.
ll = readLines("test/KEGG_2019_Mouse.txt")
genes = list()
gmt = list()
for(l in ll){
sp = unlist(strsplit(l, "\t"))
pname = sp[1]
gmt[[pname]] = sp[3:length(sp)]
}
res = list()
for(i in 1:10){
fgseaRes <- fgsea(pathways = gmt,
stats = signature,
minSize = 5,
maxSize = 4000
)
res[[length(res)+1]] = log(fgseaRes[,"pval"])
}
# ## Calulate error
#
# The calculated p-value error will be used in the p-value consistency benchmark notebook to compare GSEApy, blitzGSEA, and fGSEA
pp=do.call(cbind, res)
print(dim(pp))
ee=(1-cor(pp))
diag(ee) = NaN
paste(unlist(rowMeans(ee, na.rm=T)), collapse=",")
| testing/fgsea_benchmark.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from src.import_file import *
from src import data_read
from src import high_dim_feature_with_labels
from src import PCA_self
from src import LDA_self
'''Check results with PCA inbuilt'''
img_list,img_targets=data_read.get_data_()
N=len(img_targets)
X,Y=high_dim_feature_with_labels.norm_highx_y_(img_list,img_targets)
'K is the lower dimension'
for K in range(2,20):
X_low=PCA_self.pca_scratch_high_dim(X,K,N)
LDA_self.FiscLDA(X_low,Y)
# +
def pca_scratch_high_dim(X,K,N):
'''
Here I apply High_Dimensional_PCA for dimensionality
reduction.
Function inputs N images each flattened to the dimension high_dim=10201
which gets reduced to dimension low_dim=K.
X - N x high_dim
K - low_dim
N - no.of.samples
function returns array of dim N*low_dim
'''
'''sample_dim_matrix is of dimensions N*N
Its Eigenvectors will be of dimension N*1
Preserving top K principal components
==> Keeping K eigenvectors corresponding to the top K eigenvalues
'''
mat_N=np.dot(X,X.T)/N
eig_val,eig_vec_N=LA.eig(mat_N)
increase_idx=np.argsort(eig_val)
decrease_idx=np.flip(increase_idx)
eig_vec_D_ls=[( 1/(N*eig_val[i])**0.5 )*np.dot(X.T,eig_vec_N[:,i]) for i in range(N)]
eig_vec_D=np.stack(eig_vec_D_ls)
K_feature_ls=[np.squeeze(np.dot(eig_vec_D[0:K],X[i].reshape(-1,1))) for i in range(N)]
K_feature_arr=np.stack(K_feature_ls)
return eig_vec_D
# -
pda=PCA_scratch(1)
pda.fit(X)
# +
import numpy as np
from numpy import linalg as LA
class PCA_scratch:
def __init__(self,n_components):
self.K=n_components
def fit(self,X):
N=X.shape[0]
self.N=N
if(self.K>N):
raise ValueError('no.of.components should be less than no. of samples')
mat_N=np.dot(X,X.T)/N
eig_val,eig_vec_N=LA.eig(mat_N)
increase_idx=np.argsort(eig_val)
decrease_idx=np.flip(increase_idx)
eig_vec_D_ls=[( 1/(N*eig_val[i])**0.5 )*np.dot(X.T,eig_vec_N[:,i]) for i in range(self.N)]
eig_vec_D=np.stack(eig_vec_D_ls)
self.eig_vec_D=eig_vec_D
def fit_predict(self,X):
self.fit(X)
K_feature_ls=[np.squeeze(np.dot(self.eig_vec_D[0:self.K],X[i].reshape(-1,1))) for i in range(self.N)]
K_feature_arr=np.stack(K_feature_ls)
return K_feature_arr
def predict(self,Xtest):
K_feature_ls=[np.squeeze(np.dot(self.eig_vec_D[0:self.K],Xtest[i].reshape(-1,1))) for i in range(self.N)]
K_feature_arr=np.stack(K_feature_ls)
return K_feature_arr
# -
X=np.ones((20,10201))
X.shape
# +
pca_sl=PCA_scratch(15)
pca_sl.fit(X)
pca_sl.predict(X)[0]
# +
pca_sl=PCA_scratch(15)
# pca_sl.fit(X)
# pc
# pca_sl.eig_vec_D
pca_sl.fit_predict(X)[0]
# -
pca_sl.N
A=pca_scratch_high_dim(X,12,X.shape[0])
A[0:12].shape
X[0].shape
np.squeeze(np.dot(A[0:12],X[0].reshape(-1,1))).shape
| Combined.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python rcnn_ns
# language: python
# name: rcnn_ns
# ---
# +
import gym
from gym import wrappers
import qlearning
import numpy
import matplotlib.pyplot as plt
NUM_EPISODES = 2000
N_BINS = [8, 8, 8, 8]
MAX_STEPS = 200
FAIL_PENALTY = -100
EPSILON = 0.5
EPSILON_DECAY = 0.99
LEARNING_RATE = 0.05
DISCOUNT_FACTOR = 0.9
RECORD = False
MIN_VALUES = [-0.5, -2.0, -0.5, -3.0]
MAX_VALUES = [0.5, 2.0, 0.5, 3.0]
BINS = [numpy.linspace(MIN_VALUES[i], MAX_VALUES[i], N_BINS[i])
for i in range(4)]
# +
def discretize(obs):
return tuple([int(numpy.digitize(obs[i], BINS[i])) for i in range(4)])
def train(agent, env, history, num_episodes=NUM_EPISODES):
for i in range(NUM_EPISODES):
if i % 100:
print ("Episode {}".format(i + 1))
obs = env.reset()
cur_state = discretize(obs)
for t in range(MAX_STEPS):
action = agent.get_action(cur_state)
observation, reward, done, info = env.step(action)
next_state = discretize(observation)
if done:
reward = FAIL_PENALTY
agent.learn(cur_state, action, next_state, reward, done)
print("Episode finished after {} timesteps".format(t + 1))
history.append(t + 1)
break
agent.learn(cur_state, action, next_state, reward, done)
cur_state = next_state
if t == MAX_STEPS - 1:
history.append(t + 1)
print("Episode finished after {} timesteps".format(t + 1))
return agent, history
# +
env = gym.make('CartPole-v0')
if RECORD:
env = wrappers.Monitor(env, '/tmp/cartpole-experiment-1', force=True)
def get_actions(state):
return [0, 1]
agent = qlearning.QLearningAgent(get_actions,
epsilon=EPSILON,
alpha=LEARNING_RATE,
gamma=DISCOUNT_FACTOR,
epsilon_decay=EPSILON_DECAY)
history = []
agent, history = train(agent, env, history)
if RECORD:
env.monitor.close()
avg_reward = [numpy.mean(history[i*100:(i+1)*100]) for i in range(int(len(history)/100))]
f_reward = plt.figure(1)
plt.plot(numpy.linspace(0, len(history), len(avg_reward)), avg_reward)
plt.ylabel('Rewards')
f_reward.show()
print ('press enter to continue')
input()
plt.close()
# Display:
print ('press ctrl-c to stop')
while True:
obs = env.reset()
cur_state = discretize(obs)
done = False
t = 0
while not done:
env.render()
t = t+1
action = agent.get_action(cur_state)
observation, reward, done, info = env.step(action)
next_state = discretize(observation)
if done:
reward = FAIL_PENALTY
agent.learn(cur_state, action, next_state, reward, done)
print("Episode finished after {} timesteps".format(t+1))
history.append(t+1)
break
agent.learn(cur_state, action, next_state, reward, done)
cur_state = next_state
# -
| run_qlearning_legal.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Импорт необходимых модулей
import sys
sys.path.append('..')
from source.code.utils import preprocessing
from source.code.evaluate import my_cross_validation
from source.code.visualize import plot_results
from source.code.visualize import plot_mean_results
import pandas as pd
import numpy as np
from surprise import SVD
from surprise import NMF
from surprise import SVDpp
from surprise.prediction_algorithms.knns import KNNBasic
from surprise.model_selection import GridSearchCV
from surprise.model_selection import cross_validate
from surprise import Dataset
from surprise import Reader
# # Загрузка данных
ratings = pd.read_csv('data/BX-Book-Ratings.csv', sep=';', header=0, error_bad_lines=False, encoding='Windows-1251', low_memory=False)
books = pd.read_csv('data/BX-Books.csv', sep=';', header=0, error_bad_lines=False, encoding='Windows-1251', low_memory=False)
users = pd.read_csv('data/BX-Users.csv', sep=';', header=0, error_bad_lines=False, encoding='Windows-1251', low_memory=False)
data_dict = {}
data_dict['books'] = books
data_dict['users'] = users
data_dict['ratings'] = ratings
# # Предварительная обработка
preprocessed_data_dict = preprocessing(data_dict=data_dict, is_explicit=True, book_ratings_count_threshold=2, user_ratings_count_threshold=2)
preprocessed_data_dict['ratings'] = preprocessed_data_dict['ratings'].rename({'User-ID': 'userID', 'ISBN': 'itemID', 'Book-Rating': 'rating'}, axis='columns')
reader = Reader(rating_scale=(1, 10))
data = Dataset.load_from_df(preprocessed_data_dict['ratings'][['userID', 'itemID', 'rating']], reader)
# # Тестирование различных алгоритмов на кросс-валидации
# ## Метрики - RMSE, MAE
# ### SVD
algo_svd = SVD()
svd_cv = cross_validate(algo_svd, data, measures=['RMSE', 'MAE'], cv=5, verbose=True)
# ### NMF
algo_nmf = NMF()
nmf_cv = cross_validate(algo_nmf, data, measures=['RMSE', 'MAE'], cv=5, verbose=True)
# ### KNN (item-based)
sim_options = {
'name': 'cosine',
'user_based': False
}
algo_knn = KNNBasic(k=5, sim_options=sim_options)
knn_item_based_cv = cross_validate(algo_knn, data, measures=['RMSE', 'MAE'], cv=5, verbose=True, n_jobs=1)
# ### KNN (user-based)
sim_options = {
'name': 'cosine',
'user_based': True
}
algo_knn = KNNBasic(k=5, sim_options=sim_options)
knn_user_based_cv = cross_validate(algo_knn, data, measures=['RMSE', 'MAE'], cv=5, verbose=True, n_jobs=1)
# ## Результаты тестирования
plot_results(
[svd_cv, nmf_cv, knn_item_based_cv, knn_user_based_cv],
['SVD', 'NMF', 'Item-based', 'User-based'],
['Fold 1', 'Fold 2', 'Fold 3', 'Fold 4', 'Fold 5']
)
plot_mean_results(
[svd_cv, nmf_cv, knn_item_based_cv, knn_user_based_cv],
['SVD', 'NMF', 'Item-based', 'User-based']
)
# ## Метрики - MAP@k, MAR@k
# ### SVD
algo_svd = SVD()
svd_cv = my_cross_validation(algo_svd, data, k=5, threshold=7, n_splits=5, verbose=True)
# ### NMF
algo_nmf = NMF()
nmf_cv = my_cross_validation(algo_nmf, data, k=5, threshold=7, n_splits=5, verbose=True)
# ### KNN (item-based)
sim_options = {
'name': 'cosine',
'user_based': False
}
algo_knn = KNNBasic(k=5, sim_options=sim_options)
knn_item_based_cv = my_cross_validation(algo_knn, data, k=5, threshold=7, n_splits=5, verbose=True)
# ### KNN (user-based)
sim_options = {
'name': 'cosine',
'user_based': True
}
algo_knn = KNNBasic(k=5, sim_options=sim_options)
knn_user_based_cv = my_cross_validation(algo_knn, data, k=5, threshold=7, n_splits=5, verbose=True)
# ## Результаты тестирования
plot_results(
[svd_cv, nmf_cv, knn_item_based_cv, knn_user_based_cv],
['SVD', 'NMF', 'Item-based', 'User-based'],
['Fold 1', 'Fold 2', 'Fold 3', 'Fold 4', 'Fold 5']
)
plot_mean_results(
[svd_cv, nmf_cv, knn_item_based_cv, knn_user_based_cv],
['SVD', 'NMF', 'Item-based', 'User-based']
)
# # Тестирование SVD-разложения с учетом implicit-feedback
algo = SVDpp()
reader = Reader(rating_scale=(1, 10))
preprocessed_data_dict = preprocessing(data_dict, is_explicit=False, book_ratings_count_threshold=2, user_ratings_count_threshold=2)
preprocessed_data_dict['ratings'] = preprocessed_data_dict['ratings'].rename(
{
'User-ID': 'userID',
'ISBN': 'itemID',
'Book-Rating': 'rating'
},
axis='columns'
)
data = Dataset.load_from_df(preprocessed_data_dict['ratings'][['userID', 'itemID', 'rating']], reader)
svd_cv = my_cross_validation(algo, data, k=5, threshold=7, n_splits=5, verbose=True)
# # Использование Grid search для тюнинга гиперпараметров
# ## Метрики - RMSE, MAE
# ### Загрузка данных
preprocessed_data_dict = preprocessing(data_dict=data_dict, is_explicit=True, book_ratings_count_threshold=2, user_ratings_count_threshold=2)
preprocessed_data_dict['ratings'] = preprocessed_data_dict['ratings'].rename({'User-ID': 'userID', 'ISBN': 'itemID', 'Book-Rating': 'rating'}, axis='columns')
reader = Reader(rating_scale=(1, 10))
data = Dataset.load_from_df(preprocessed_data_dict['ratings'][['userID', 'itemID', 'rating']], reader)
# ### SVD
param_grid = {
'n_epochs': [5, 10],
'lr_all': [0.002, 0.005],
'reg_all': [0.4, 0.6]
}
gs = GridSearchCV(SVD, param_grid, measures=['rmse', 'mae'], cv=5)
gs.fit(data)
print(gs.best_score['rmse'])
print(gs.best_params['rmse'])
# ### NMF
param_grid = {
'n_epochs': [5, 10],
'lr_bu': [0.002, 0.005],
'lr_bi': [0.4, 0.6]
}
gs = GridSearchCV(NMF, param_grid, measures=['rmse', 'mae'], cv=5)
gs.fit(data)
print(gs.best_score['rmse'])
print(gs.best_params['rmse'])
# ### KNN (item-based)
param_grid = {
'bsl_options': {
'method': ['als', 'sgd'],
'reg': [1, 2]
},
'k': [2, 3],
'sim_options': {
'name': ['msd', 'cosine'],
'min_support': [1, 5],
'user_based': [False]
}
}
gs = GridSearchCV(KNNBasic, param_grid, measures=['rmse', 'mae'], cv=5, n_jobs=1)
gs.fit(data)
print(gs.best_score['rmse'])
print(gs.best_params['rmse'])
# ### KNN (user-based)
param_grid = {
'bsl_options': {
'method': ['als', 'sgd'],
'reg': [1, 2]
},
'k': [2, 3],
'sim_options': {
'name': ['msd', 'cosine'],
'min_support': [1, 5],
'user_based': [True]
}
}
gs = GridSearchCV(KNNBasic, param_grid, measures=['rmse', 'mae'], cv=5, n_jobs=1)
gs.fit(data)
print(gs.best_score['rmse'])
print(gs.best_params['rmse'])
| notebooks/EXPLICIT_TASK_SOLUTION.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ```py
# # -*- conding:utf-8 -*-
#
# import logging
# import logging.config
# import os
#
# __all__ = ['getLogger']
#
# DIR_DEBUG = './logs/debug'
# DIR_INFO = './logs/info'
# DIR_ERROR = './logs/error'
#
# os.makedirs(DIR_DEBUG, exist_ok=True)
# os.makedirs(DIR_INFO, exist_ok=True)
# os.makedirs(DIR_ERROR, exist_ok=True)
#
# IS_INIT = False
#
# LOGGING = {
# "version": 1,
# "disable_existing_loggers": False,
# # 日志输出格式设置
# "formatters": {
# "simple": {
# 'format': '%(asctime)s [%(name)s:%(lineno)d] [%(levelname)s]- %(message)s'
# },
# 'standard': {
# 'format': '%(asctime)s [%(threadName)s:%(thread)d] [%(name)s:%(lineno)d] [%(levelname)s]- %(message)s'
# },
# },
#
# "handlers": {
# # 控制台处理器,控制台输出
# "console": {
# "class": "logging.StreamHandler",
# "level": "DEBUG",
# "formatter": "simple",
# "stream": "ext://sys.stdout"
# },
# # 默认处理器, 文件大小滚动
# "debug": {
# "class": "logging.handlers.RotatingFileHandler",
# "level": "DEBUG",
# "formatter": "simple",
# 'filename': os.path.join(DIR_DEBUG, 'debug.log'),
# "maxBytes": 1024 * 1024 * 5, # 5 MB
# "backupCount": 50,
# "encoding": "utf8"
# },
# # 错误日志, 文件大小滚动
# 'error': {
# 'level': 'ERROR',
# 'class': 'logging.handlers.RotatingFileHandler',
# 'filename': os.path.join(DIR_ERROR, 'error.log'),
# 'maxBytes': 1024 * 1024 * 2,
# 'backupCount': 10,
# 'formatter': 'standard',
# 'encoding': 'utf-8',
# },
# # info 及以上级别, 日期滚动
# 'info': {
# 'class': 'logging.handlers.TimedRotatingFileHandler',
# 'level': 'INFO',
# "formatter": "simple",
# 'filename': os.path.join(DIR_INFO, 'info.log'),
# 'when': 'D', # 单位: 天
# 'interval': 1, # 滚动周期
# 'backupCount': 30, # 备份30天
# "encoding": "utf8"
# },
# },
# 'loggers': {
# 'websockets': {
# 'handlers': ['debug', 'console'],
# 'level': "ERROR",
# 'propagate': False
# },
# },
# "root": {
# 'handlers': ['debug', 'error', 'info', 'console'],
# 'level': "DEBUG",
# 'propagate': False
# }
# }
#
#
# def initLogConf():
# """
# 配置日志
# """
# global IS_INIT
# if IS_INIT:
# return
# logging.config.dictConfig(LOGGING)
# logging.debug("Init logging")
# IS_INIT = True
#
#
# def getLogger(name):
# return logging.getLogger(name)
#
#
# initLogConf()
#
# if __name__ == '__main__':
# logger = getLogger('aaa')
# logger.debug('debug')
# logger.info('info')
# logger.error('error')
# ```
| 开发常用/日志-logging-常用配置文件.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="cyrwiLlGODRv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="9cf4e427-f3c3-465e-cd09-f040524743df"
# !git clone https://github.com/tensorflow/models.git
# + id="ZyOXCEzZAQuG" colab_type="code" colab={}
# + id="TSGF2LnbO10P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="32b2fdac-70c0-4ed2-f35f-d9665310cb17"
# %%shell
# cd ..
# ls
# + id="kax2NQWmO_OT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 150} outputId="a6f1317b-f6bd-4ae6-af28-93ba5ce62a07"
# %cd ..
# %ls
# + id="JVsOA6h7kkHo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 873} outputId="a54b7fb0-13f6-44de-c490-3f4149da17be"
# !pip install tensorflow==1.15.0
# + id="7_7bsxygO_Qi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="d1329185-0f45-462a-a21d-144714a77919"
# %cd ..
# !apt-get install protobuf-compiler python-pil python-lxml python-tk
# !pip install Cython
# !pip install jupyter
# !pip install matplotlib
# %cd /content/models/research
# !protoc object_detection/protos/*.proto --python_out=.
# %set_env PYTHONPATH=/content/models/research:/content/models/research/slim
# !python object_detection/builders/model_builder_test.py
# + id="0S_blA8yHXGO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 154} outputId="d093b6ad-db1f-499b-bbfc-69d5fab8d77b"
# !pip install tf_slim
# + id="GRW4MllQO_S0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="2b18e99a-2400-4c79-8207-6106ccab0eb1"
# !python object_detection/legacy/train.py \
# --train_dir=/content/models/research/object_detection/main_files/CP \
# --pipeline_config_path=/content/models/research/object_detection/main_files/pipeline.config
# + id="B3mpLX60e-or" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="819cc4d5-5354-4e98-b129-9119b4c2a1da"
# !python object_detection/export_inference_graph.py \
# --input_type=image_tensor \
# --pipeline_config_path=/content/models/research/object_detection/main_files/pipeline.config \
# --trained_checkpoint_prefix=/content/models/research/object_detection/main_files/CP/model.ckpt-2000 \
# --output_directory=/content/models/research/object_detection/main_files/IG
# + id="PUXrdUHzO_VI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 117} outputId="1ef34205-3e3f-4767-8ed6-cc8ee85abd3f"
# FIND OUT INPUT AND OUTPUT TENSOR NAMES
import tensorflow as tf
gf = tf.GraphDef()
m_file = open('/content/models/research/object_detection/main_files/IG/frozen_inference_graph.pb','rb')
gf.ParseFromString(m_file.read())
with open('somefile.txt', 'a') as the_file:
for n in gf.node:
the_file.write(n.name+'\n')
file = open('somefile.txt','r')
data = file.readlines()
print ("output name = ")
print (data[len(data)-1])
print ("Input name = ")
file.seek ( 0 )
print (file.readline())
# + id="dPb-spmEaLOo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="0c603974-850e-418c-851a-16e7adea5aed"
# %%shell
tflite_convert \
--graph_def_file=/content/models/research/object_detection/main_files/IG/frozen_inference_graph.pb \
--output_file=/content/models/research/object_detection/main_files/IG/ssd_v1.tflite \
--output_format=TFLITE \
--input_shapes=1,300,300,3 \
--input_arrays=normalized_input_image_tensor \
--output_arrays='TFLite_Detection_PostProcess','TFLite_Detection_PostProcess:1','TFLite_Detection_PostProcess:2','TFLite_Detection_PostProcess:3' \
--inference_type=QUANTIZED_UINT8 \
--mean_values=128 \
--std_dev_values=127 \
--change_concat_input_ranges=false \
--allow_custom_ops
# + id="B-ryhOeMcFD9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="d3582365-851e-4bbb-c020-88a1807e95ca"
# !tflite_convert --graph_def_file=/content/models/research/object_detection/main_files/IG/frozen_inference_graph.pb --output_file=/content/models/research/object_detection/main_files/IG/ssd_v1.tflite --output_format=TFLITE --input_shapes=1,300,300,3 --input_arrays=image_tensor --output_arrays=raw_detection_scores --inference_type=QUANTIZED_UINT8 --mean_values=128 --std_dev_values=127 --change_concat_input_ranges=false --allow_custom_ops
# + id="88rmYBKAeak2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="d0298bc2-1d4b-44d1-9ef2-02e83a27e14d"
# !toco --graph_def_file=/content/models/research/object_detection/main_files/IG/frozen_inference_graph.pb --output_file=./ssd_v1.tflite --input_format=TENSORFLOW_GRAPHDEF --output_format=TFLITE –-input_shape=1,299,299,3 --input_array=image_tensor --output_array=raw_detection_scores --inference_type=FLOAT --input_type=FLOAT
| Mahdi_Paper_detection_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: default:Python
# language: python
# name: conda-env-default-py
# ---
# !wget -O 'download-img-reddit.py' https://raw.githubusercontent.com/nephre/reddit-media-downloader/master/reddit.py
# +
# # !python 'download-img-reddit.py' -r yerbamate -t 4 --images-only
# +
# # !cd downloads/yerbamate/ && ls -1 | wc -l
| 0-create-dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
# Code to calculate absorption coefficient and reflection values for polymer films as measured on bare mirror
#Import necessary packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sb
# Define custom functions to calculate c and alpha via T = c + exp(-alpha*t) where T is transmission, c is reflection,
# alpha is extinciton coefficient, and t is thickness
def getC(transmission, alpha, thickness):
return(transmission-np.exp(-alpha*thickness))
def getAlpha(transmission, c, thickness):
return(-np.log(transmission-c)/thickness)
path = 'C:\\Users\\acarr.BNL\\Documents\\Data\\Optical Transmission\\'
cEVAtrichloro=[]; alphaEVAtrichloro=[]; transmissionEVAtrichloro=[];
cEVAtoluene=[]; alphaEVAtoluene=[]; transmissionEVAtoluene=[];
cEVAacetone=[]; alphaEVAacetone=[]; transmissionEVAacetone=[];
# load transmission data into lists, convert to arrays
for n in range(1,7):
samp='p_EVA_trichlorobenzene_'+str(n)+'_fixed.csv';
A=pd.read_csv(path+'Polymer_Backgrounds\\'+samp);
transmissionEVAtrichloro.append(A.mean().mean())
if n==6:
transmissionEVAtrichloro=np.asarray(transmissionEVAtrichloro)
for n in range(1,7):
samp='p_EVA_toluene_'+str(n)+'_fixed.csv';
A=pd.read_csv(path+'Polymer_Backgrounds\\'+samp);
transmissionEVAtoluene.append(A.mean().mean())
if n==6:
transmissionEVAtoluene=np.asarray(transmissionEVAtoluene)
for n in range(1,7):
samp='p_EVA_acetone_'+str(n)+'_fixed.csv';
A=pd.read_csv(path+'Polymer_Backgrounds\\'+samp);
transmissionEVAacetone.append(A.mean().mean())
if n==6:
transmissionEVAacetone=np.asarray(transmissionEVAacetone)
thicknessEVAtrichloro=np.asarray([0.172,0.090,0.097,0.090,0.102,0.126])/10;
thicknessEVAtoluene=np.asarray([0.165,0.207,0.195,0.189,0.160,0.208])/10;
thicknessEVAacetone=np.asarray([0.184,0.219,0.250,0.181,0.215,0.245])/10;
# Calculate constant c using lit alpha of ~ 90. Then calculate actual alpha using average c
for n in range(0,6):
cEVAtrichloro.append(getC(transmissionEVAtrichloro[n], 90, thicknessEVAtrichloro[n]))
cEVAtoluene.append(getC(transmissionEVAtoluene[n], 90, thicknessEVAtoluene[n]))
cEVAacetone.append(getC(transmissionEVAacetone[n], 90, thicknessEVAacetone[n]))
if n==5:
cEVAtrichloro=np.asarray(cEVAtrichloro);cEVAtoluene=np.asarray(cEVAtoluene);cEVAacetone=np.asarray(cEVAacetone);
for n in range(0,6):
alphaEVAtrichloro.append(getAlpha(transmissionEVAtrichloro[n], cEVAtrichloro.mean(), thicknessEVAtrichloro[n]))
alphaEVAtoluene.append(getAlpha(transmissionEVAtoluene[n], cEVAtoluene.mean(), thicknessEVAtoluene[n]))
alphaEVAacetone.append(getAlpha(transmissionEVAacetone[n], cEVAacetone.mean(), thicknessEVAacetone[n]))
if n==5:
alphaEVAtrichloro=np.asarray(alphaEVAtrichloro);
alphaEVAtoluene=np.asarray(alphaEVAtoluene);
alphaEVAacetone=np.asarray(alphaEVAacetone);
# Print all results for averages and std
print('EVA/acetone avg transmission = '+str(transmissionEVAacetone[~np.isnan(transmissionEVAacetone)].mean())...
+' +/- '+str(transmissionEVAacetone[~np.isnan(transmissionEVAacetone)].std()))
print('EVA/acetone avg alpha = '+str(alphaEVAacetone[~np.isnan(alphaEVAacetone)].mean())+...
' +/- '+str(alphaEVAacetone[~np.isnan(alphaEVAacetone)].std()))
print('EVA/acetone avg c = '+str(cEVAacetone[~np.isnan(cEVAacetone)].mean())+...
' +/- '+str(cEVAacetone[~np.isnan(cEVAacetone)].std())+'\n')
print('EVA/toluene avg transmission = '+str(transmissionEVAtoluene[~np.isnan(transmissionEVAtoluene)].mean())+...
' +/- '+str(transmissionEVAtoluene[~np.isnan(transmissionEVAtoluene)].std()))
print('EVA/toluene avg alpha = '+str(alphaEVAtoluene[~np.isnan(alphaEVAtoluene)].mean())+...
' +/- '+str(alphaEVAtoluene[~np.isnan(alphaEVAtoluene)].std()))
print('EVA/toluene avg c = '+str(cEVAtoluene[~np.isnan(cEVAtoluene)].mean())+...
' +/- '+str(cEVAtoluene[~np.isnan(cEVAtoluene)].std())+'\n')
print('EVA/trichloro avg transmission = '+str(transmissionEVAtrichloro[~np.isnan(transmissionEVAtrichloro)].mean())+...
' +/- '+str(transmissionEVAtrichloro[~np.isnan(transmissionEVAtrichloro)].std()))
print('EVA/trichloro avg alpha = '+str(alphaEVAtrichloro[~np.isnan(alphaEVAtrichloro)].mean())+...
' +/- '+str(alphaEVAtrichloro[~np.isnan(alphaEVAtrichloro)].std()))
print('EVA/trichloro avg c = '+str(cEVAtrichloro[~np.isnan(cEVAtrichloro)].mean())+...
' +/- '+str(cEVAtrichloro[~np.isnan(cEVAtrichloro)].std())+'\n')
# +
# %matplotlib inline
# Code to plot heatmaps and kernel density estimation curves for polymer films as measured on the bare mirror and in
# in between 2 quartz slides
#Import necessary packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
plt.rcParams.update({'font.size': 18})
import seaborn as sb
path ='C:\\Users\\acarr.BNL\\Documents\\Data\\Optical Transmission\\';
# Load data
samp='p_EVA_acetone_1_fixed.csv'; polymer1=pd.read_csv(path+'Polymer_Backgrounds\\'+samp);
samp='p_EVA_acetone_2_fixed.csv'; polymer2=pd.read_csv(path+'Polymer_Backgrounds\\'+samp);
samp='p_EVA_acetone_3_fixed.csv'; polymer3=pd.read_csv(path+'Polymer_Backgrounds\\'+samp);
samp='p_EVA_acetone_1_2slides_fixed.csv'; polymer1slides=pd.read_csv(path+samp);
samp='p_EVA_acetone_2_2slides_fixed.csv'; polymer2slides=pd.read_csv(path+samp);
samp='p_EVA_acetone_3_2slides_fixed.csv'; polymer3slides=pd.read_csv(path+samp);
# Make figure with correct subplot axes
fig = plt.figure(figsize = (15,15))
ax1 = plt.subplot2grid((3,3), (0,0)); ax2 = plt.subplot2grid((3,3), (0,1)); ax3 = plt.subplot2grid((3,3), (0,2));
ax4 = plt.subplot2grid((3,3), (1,0)); ax5 = plt.subplot2grid((3,3), (1,1)); ax6 = plt.subplot2grid((3,3), (1,2));
ax7 = plt.subplot2grid((3,3), (2,0)); ax8 = plt.subplot2grid((3,3), (2,1)); ax9 = plt.subplot2grid((3,3), (2,2));
# Show heatmaps and kernel density estimations on axes
sb.heatmap(polymer1, ax=ax1); sb.heatmap(polymer1slides, ax=ax2);
sb.kdeplot(polymer1.values.flatten(), label='EVA', ax=ax3);
sb.kdeplot(polymer1slides.values.flatten(), label='EVA + slides', ax=ax3);
sb.heatmap(polymer2, ax=ax4); sb.heatmap(polymer2slides, ax=ax5);
sb.kdeplot(polymer2.values.flatten(),label='EVA',ax=ax6);
sb.kdeplot(polymer2slides.values.flatten(),label='EVA + slides',ax=ax6);
sb.heatmap(polymer3, ax=ax7); sb.heatmap(polymer3slides, ax=ax8);
sb.kdeplot(polymer3.values.flatten(),label='EVA',ax=ax9);
sb.kdeplot(polymer3slides.values.flatten(),label='EVA + slides',ax=ax9);
# Generate axes labels
ax=[ax3,ax6,ax9];
for n in ax:
n.set_ylabel('Probability Density'); n.set_xlabel('Det/Inc');
plt.tight_layout(); plt.show()
# +
# %matplotlib inline
# Code to calculate absorption coefficient and reflection values for polymer films as measured on bare mirror and in
# between two quartz slides
#Import necessary packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sb
# Define custom functions to calculate c and alpha via T = c + exp(-alpha*t) where T is transmission, c is reflection,
# alpha is extinciton coefficient, and t is thickness
def getC(transmission, alpha, thickness):
return(transmission-np.exp(-alpha*thickness))
def getAlpha(transmission, c, thickness):
return(-np.log(transmission-c)/thickness)
path = 'C:\\Users\\acarr.BNL\\Documents\\Data\\Optical Transmission\\'
cEVAnoslides=[]; alphaEVAnoslides=[]; transmissionEVAnoslides=[];
cEVAslides=[]; alphaEVAslides=[]; transmissionEVAslides=[];
# load transmission data into lists, convert to arrays
for n in range(1,4):
samp='p_EVA_acetone_'+str(n)+'_fixed.csv';
A=pd.read_csv(path+'Polymer_Backgrounds\\'+samp);
transmissionEVAnoslides.append(A.mean().mean())
if n==3:
transmissionEVAnoslides=np.asarray(transmissionEVAnoslides)
for n in range(1,4):
samp='p_EVA_acetone_'+str(n)+'_2slides_fixed.csv';
A=pd.read_csv(path+samp);
transmissionEVAslides.append(A.mean().mean())
if n==3:
transmissionEVAslides=np.asarray(transmissionEVAslides)
thicknessEVA=np.asarray([0.184,0.219,0.250])/10;
# Calculate constant c using lit alpha of ~ 90. Then calculate actual alpha using average c
for n in range(0,3):
cEVAnoslides.append(getC(transmissionEVAnoslides[n], 90, thicknessEVA[n]))
cEVAslides.append(getC(transmissionEVAslides[n], 90, thicknessEVA[n]))
if n==2:
cEVAnoslides=np.asarray(cEVAnoslides);cEVAslides=np.asarray(cEVAslides)
for n in range(0,3):
alphaEVAnoslides.append(getAlpha(transmissionEVAnoslides[n], cEVAnoslides.mean(), thicknessEVA[n]))
alphaEVAslides.append(getAlpha(transmissionEVAslides[n], cEVAslides.mean(), thicknessEVA[n]))
if n==2:
alphaEVAnoslides=np.asarray(alphaEVAnoslides); alphaEVAslides=np.asarray(alphaEVAslides);
# Print results
print('EVA/noslides avg transmission = '+str(transmissionEVAnoslides[~np.isnan(transmissionEVAnoslides)].mean())+' +/- '+str(transmissionEVAnoslides[~np.isnan(transmissionEVAnoslides)].std()))
print('EVA/noslides avg alpha = '+str(alphaEVAnoslides[~np.isnan(alphaEVAnoslides)].mean())+' +/- '+str(alphaEVAnoslides[~np.isnan(alphaEVAnoslides)].std()))
print('EVA/noslides avg c = '+str(cEVAnoslides[~np.isnan(cEVAnoslides)].mean())+' +/- '+str(cEVAnoslides[~np.isnan(cEVAnoslides)].std())+'\n')
print('EVA/slides avg transmission = '+str(transmissionEVAslides[~np.isnan(transmissionEVAslides)].mean())+' +/- '+str(transmissionEVAslides[~np.isnan(transmissionEVAslides)].std()))
print('EVA/slides avg alpha = '+str(alphaEVAslides[~np.isnan(alphaEVAslides)].mean())+' +/- '+str(alphaEVAslides[~np.isnan(alphaEVAslides)].std()))
print('EVA/slides avg c = '+str(cEVAslides[~np.isnan(cEVAslides)].mean())+' +/- '+str(cEVAslides[~np.isnan(cEVAslides)].std())+'\n')
# +
# %matplotlib inline
# Code to calculate absorption coefficient and reflection values for polymer films with and without graphene as measured
# between two quartz slides
#Import necessary packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sb
# Define custom functions to calculate c and alpha via T = c + exp(-alpha*t) where T is transmission, c is reflection,
# alpha is extinciton coefficient, and t is thickness
def getC(transmission, alpha, thickness):
return(transmission-np.exp(-alpha*thickness))
def getAlpha(transmission, c, thickness):
return(-np.log(transmission-c)/thickness)
path = 'C:\\Users\\acarr.BNL\\Documents\\Data\\Optical Transmission\\'
cEVAGrnoslides=[]; alphaEVAGrnoslides=[]; transmissionEVAGrnoslides=[];
cEVAGrslides=[]; alphaEVAGrslides=[]; transmissionEVAGrslides=[];
thicknessEVA=np.asarray([0.265,0.178,0.234])/10;
# load transmission data into lists, convert to arrays
letters = ['CK','CJ','CL']
for n, lett in enumerate(letters):
samp='Gr_p_EVA_acetone_'+lett+'_fixed.csv';
A=pd.read_csv(path+'2019_03_13\\'+samp); transmissionEVAGrnoslides.append(A.mean().mean())
samp='Gr_p_EVA_acetone_'+lett+'_2slides_fixed.csv';
A=pd.read_csv(path+'2019_04_26\\'+samp); transmissionEVAGrslides.append(A.mean().mean())
if n==2:
transmissionEVAGrnoslides=np.asarray(transmissionEVAGrnoslides);
transmissionEVAGrslides=np.asarray(transmissionEVAGrslides)
# Calculate constant c using lit alpha of ~ 90. Then calculate actual alpha using average c
for n in range(0,3):
cEVAGrnoslides.append(getC(transmissionEVAGrnoslides[n], 90, thicknessEVA[n]))
cEVAGrslides.append(getC(transmissionEVAGrslides[n], 90, thicknessEVA[n]))
if n==2:
cEVAGrnoslides=np.asarray(cEVAGrnoslides);cEVAGrslides=np.asarray(cEVAGrslides)
for n in range(0,3):
alphaEVAGrnoslides.append(getAlpha(transmissionEVAGrnoslides[n], cEVAGrnoslides.mean(), thicknessEVA[n]))
alphaEVAGrslides.append(getAlpha(transmissionEVAGrslides[n], cEVAGrslides.mean(), thicknessEVA[n]))
if n==2:
alphaEVAGrnoslides=np.asarray(alphaEVAGrnoslides); alphaEVAGrslides=np.asarray(alphaEVAGrslides);
# Print results
print('Gr/EVA/noslides avg transmission = '+str(transmissionEVAGrnoslides[~np.isnan(transmissionEVAGrnoslides)].mean())+' +/- '+str(transmissionEVAGrnoslides[~np.isnan(transmissionEVAGrnoslides)].std()))
print('Gr/EVA/noslides avg alpha = '+str(alphaEVAGrnoslides[~np.isnan(alphaEVAGrnoslides)].mean())+' +/- '+str(alphaEVAGrnoslides[~np.isnan(alphaEVAGrnoslides)].std()))
print('Gr/EVA/noslides avg c = '+str(cEVAGrnoslides[~np.isnan(cEVAGrnoslides)].mean())+' +/- '+str(cEVAGrnoslides[~np.isnan(cEVAGrnoslides)].std())+'\n')
print('Gr/EVA/slides avg transmission = '+str(transmissionEVAGrslides[~np.isnan(transmissionEVAGrslides)].mean())+' +/- '+str(transmissionEVAGrslides[~np.isnan(transmissionEVAGrslides)].std()))
print('Gr/EVA/slides avg alpha = '+str(alphaEVAGrslides[~np.isnan(alphaEVAGrslides)].mean())+' +/- '+str(alphaEVAGrslides[~np.isnan(alphaEVAGrslides)].std()))
print('Gr/EVA/slides avg c = '+str(cEVAGrslides[~np.isnan(cEVAGrslides)].mean())+' +/- '+str(cEVAGrslides[~np.isnan(cEVAGrslides)].std())+'\n')
# +
# %matplotlib inline
# Code to plot heatmaps and kernel density estimation curves for polymer films with and without graphene as measured
# in between 2 quartz slides
#Import necessary packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
plt.rcParams.update({'font.size': 18})
import seaborn as sb
# Load heatmap data
path ='C:\\Users\\acarr.BNL\\Documents\\Data\\Optical Transmission\\';
samp='Gr_p_EVA_acetone_CK_fixed.csv'; Grpolymer1=pd.read_csv(path+'2019_03_13\\'+samp);
samp='Gr_p_EVA_acetone_CJ_fixed.csv'; Grpolymer2=pd.read_csv(path+'2019_03_13\\'+samp);
samp='Gr_p_EVA_acetone_CL_fixed.csv'; Grpolymer3=pd.read_csv(path+'2019_03_13\\'+samp);
samp='Gr_p_EVA_acetone_CK_2slides_fixed.csv'; Grpolymer1slides=pd.read_csv(path+'2019_04_26\\'+samp);
samp='Gr_p_EVA_acetone_CJ_2slides_fixed.csv'; Grpolymer2slides=pd.read_csv(path+'2019_04_26\\'+samp);
samp='Gr_p_EVA_acetone_CL_2slides_fixed.csv'; Grpolymer3slides=pd.read_csv(path+'2019_04_26\\'+samp);
# Generate figure with correct subplot axes
fig = plt.figure(figsize = (15,15))
ax1 = plt.subplot2grid((3,3), (0,0)); ax2 = plt.subplot2grid((3,3), (0,1)); ax3 = plt.subplot2grid((3,3), (0,2));
ax4 = plt.subplot2grid((3,3), (1,0)); ax5 = plt.subplot2grid((3,3), (1,1)); ax6 = plt.subplot2grid((3,3), (1,2));
ax7 = plt.subplot2grid((3,3), (2,0)); ax8 = plt.subplot2grid((3,3), (2,1)); ax9 = plt.subplot2grid((3,3), (2,2));
# Plot heatmaps and kde on correct axes
sb.heatmap(Grpolymer1, ax=ax1); sb.heatmap(Grpolymer1slides, ax=ax2);
sb.kdeplot(Grpolymer1.values.flatten(), label='Gr/EVA', ax=ax3);
sb.kdeplot(Grpolymer1slides.values.flatten(), label='Gr/EVA + slides', ax=ax3);
sb.heatmap(Grpolymer2, ax=ax4); sb.heatmap(Grpolymer2slides, ax=ax5);
sb.kdeplot(Grpolymer2.values.flatten(),label='Gr/EVA',ax=ax6);
sb.kdeplot(Grpolymer2slides.values.flatten(),label='Gr/EVA + slides',ax=ax6);
sb.heatmap(Grpolymer3, ax=ax7); sb.heatmap(Grpolymer3slides, ax=ax8);
sb.kdeplot(Grpolymer3.values.flatten(),label='Gr/EVA',ax=ax9);
sb.kdeplot(Grpolymer3slides.values.flatten(),label='Gr/EVA + slides',ax=ax9);
# Add axes labels
ax=[ax3,ax6,ax9];
for n in ax:
n.set_ylabel('Probability Density'); n.set_xlabel('Det/Inc'); n.legend(loc=8)
plt.tight_layout(); plt.show()
# +
# %matplotlib inline
# Code to plot heatmaps and kernel density estimation curves for PMMA samples measured on different/same days
#Import necessary packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
plt.rcParams.update({'font.size': 18})
import seaborn as sb
# Load heatmap data
path ='C:\\Users\\acarr.BNL\\Documents\\Data\\Optical Transmission\\2019_05_13\\';
samp='PMMA_180_4';
Grpolymer1=pd.read_csv(path+samp+'_fixed.csv'); Grpolymer2=pd.read_csv(path+samp+'_check_fixed.csv');
# Generate figure with correct subplot axes
fig = plt.figure(figsize = (15,15))
ax1 = plt.subplot2grid((2,4), (0,0), colspan = 2); ax2 = plt.subplot2grid((2,4), (0,2), colspan = 2);
ax3 = plt.subplot2grid((2,4), (1,1), colspan = 2);
# Plot heatmaps and kde on correct axes
sb.heatmap(Grpolymer1, ax=ax1);
sb.distplot(Grpolymer1.values.flatten(), label='Scan 1', ax=ax3);
sb.heatmap(Grpolymer2, ax=ax2);
sb.distplot(Grpolymer2.values.flatten(),label='Scan 2',ax=ax3);
# Add axes labels
ax3.set_ylabel('Probability Density'); ax3.set_xlabel('Det/Inc'); #n.legend(loc=8)
ax3.legend(); plt.tight_layout(); plt.show()
# +
# %matplotlib inline
# %run get_heatmap.ipynb
# Code to calculate number of layers of graphene transferred as measured between two quartz slides
#Import necessary packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sb
def trans_curve(x, alpha, c):
# where x is film thickness [cm], y is optical transmission, aplha is absorption coefficient [1/cm], and c is both
# back and front reflection plus other constants
return(c + np.exp(-alpha*x))
def getC(transmission, alpha, thickness):
return(transmission-np.exp(-alpha*thickness))
def getAlpha(transmission, c, thickness):
return(-np.log(transmission-c)/thickness)
# Load heatmap data
path ='C:\\Users\\acarr.BNL\\Documents\\Data\\Optical Transmission\\';
samp='Gr_p_EVA_acetone_CK_2slides_fixed.csv'; stackPower1=pd.read_csv(path+'2019_04_26\\'+samp);
samp='Gr_p_EVA_acetone_CJ_2slides_fixed.csv'; stackPower2=pd.read_csv(path+'2019_04_26\\'+samp);
samp='Gr_p_EVA_acetone_CL_2slides_fixed.csv'; stackPower3=pd.read_csv(path+'2019_04_26\\'+samp);
samp='Gr_p_EVA_acetone_BH_2slides_fixed.csv'; stackPower4=pd.read_csv(path+'2019_04_30\\'+samp);
samp='Gr_p_EVA_acetone_BG_2slides_fixed.csv'; stackPower5=pd.read_csv(path+'2019_04_30\\'+samp);
samp='Gr_p_EVA_acetone_BF_2slides_fixed.csv'; stackPower6=pd.read_csv(path+'2019_04_30\\'+samp);
c=-0.006135; alpha=56.94;
bkg1 = trans_curve(0.0265, alpha, c); bkg2 = trans_curve(0.0178, alpha, c); bkg3 = trans_curve(0.0234, alpha, c);
bkg4 = trans_curve(0.0132, alpha, c); bkg5 = trans_curve(0.0126, alpha, c); bkg6 = trans_curve(0.0171, alpha, c);
# Calculate number of layers of graphene assuming each layer of graphene absorbs 4.6% of HeNe laser roundtrip, i.e.
# light passes through each layer twice, for the Gr/polymer stack
stackLayers1=(bkg1 - stackPower1)/0.046;
stackLayers1=stackLayers1[stackLayers1>0]; stackLayers1=stackLayers1[stackLayers1<2]
stackLayers2=(bkg2 - stackPower2)/0.046
stackLayers2=stackLayers2[stackLayers2>0]; stackLayers2=stackLayers2[stackLayers2<2]
stackLayers3=(bkg3 - stackPower3)/0.046
stackLayers3=stackLayers3[stackLayers3>0]; stackLayers3=stackLayers3[stackLayers3<2]
stackLayers4=(bkg4 - stackPower4)/0.046;
stackLayers4=stackLayers4[stackLayers4>0]; stackLayers4=stackLayers4[stackLayers4<2]
stackLayers5=(bkg5 - stackPower5)/0.046
stackLayers5=stackLayers5[stackLayers5>0]; stackLayers5=stackLayers2[stackLayers2<2]
stackLayers6=(bkg6 - stackPower6)/0.046
stackLayers6=stackLayers6[stackLayers6>0]; stackLayers6=stackLayers3[stackLayers3<2]
idx1 = 0; idx2 = 25; idx3 = 0; idx4 = 25;
#idx1 = 2; idx2 = 10; idx3 = 2; idx4 = 12;
plt.title('Det/Inc w background 1')
plt.show(sb.heatmap(stackPower1, annot = False, cmap = 'coolwarm'))
print('avg Det/Inc = ' + str(stackPower1.mean().mean()))
print('min Det/Inc = ' +str(stackPower1.min().min())); print('max Det/Inc = ' +str(stackPower1.max().max()))
plt.title('Det/Inc w background 2')
plt.show(sb.heatmap(stackPower2, annot = False, cmap = 'coolwarm'))
print('avg Det/Inc = ' + str(stackPower2.mean().mean()))
print('min Det/Inc = ' +str(stackPower2.min().min())); print('max Det/Inc = ' +str(stackPower2.max().max()))
plt.title('Det/Inc w background 3')
plt.show(sb.heatmap(stackPower3, annot = False, cmap = 'coolwarm'))
print('avg Det/Inc = ' + str(stackPower3.mean().mean()))
print('min Det/Inc = ' +str(stackPower3.min().min())); print('max Det/Inc = ' +str(stackPower3.max().max()))
plt.title('Det/Inc w background 4')
plt.show(sb.heatmap(stackPower4, annot = False, cmap = 'coolwarm'))
print('avg Det/Inc = ' + str(stackPower4.mean().mean()))
print('min Det/Inc = ' +str(stackPower4.min().min())); print('max Det/Inc = ' +str(stackPower4.max().max()))
plt.title('Det/Inc w background 5')
plt.show(sb.heatmap(stackPower5, annot = False, cmap = 'coolwarm'))
print('avg Det/Inc = ' + str(stackPower5.mean().mean()))
print('min Det/Inc = ' +str(stackPower5.min().min())); print('max Det/Inc = ' +str(stackPower5.max().max()))
plt.title('Det/Inc w background 6')
plt.show(sb.heatmap(stackPower6, annot = False, cmap = 'coolwarm'))
print('avg Det/Inc = ' + str(stackPower6.mean().mean()))
print('min Det/Inc = ' +str(stackPower6.min().min())); print('max Det/Inc = ' +str(stackPower6.max().max()))
background = np.zeros_like(stackPower1) + bkg1; stackPower1_nobkg = background - stackPower1
background = np.zeros_like(stackPower2) + bkg2; stackPower2_nobkg = background - stackPower2
background = np.zeros_like(stackPower3) + bkg3; stackPower3_nobkg = background - stackPower3
background = np.zeros_like(stackPower4) + bkg4; stackPower4_nobkg = background - stackPower4
background = np.zeros_like(stackPower5) + bkg5; stackPower5_nobkg = background - stackPower5
background = np.zeros_like(stackPower6) + bkg6; stackPower6_nobkg = background - stackPower6
plt.title('Det/Inc wo background 1')
plt.show(sb.heatmap(stackPower1_nobkg, annot = False, cmap = 'coolwarm'))
print('background = ' + str(bkg1))
plt.title('Det/Inc wo background 2')
plt.show(sb.heatmap(stackPower2_nobkg, annot = False, cmap = 'coolwarm'))
print('background = ' + str(bkg2))
plt.title('Det/Inc wo background 3')
plt.show(sb.heatmap(stackPower3_nobkg, annot = False, cmap = 'coolwarm'))
print('background = ' + str(bkg3))
plt.title('Det/Inc wo background 4')
plt.show(sb.heatmap(stackPower4_nobkg, annot = False, cmap = 'coolwarm'))
print('background = ' + str(bkg4))
plt.title('Det/Inc wo background 5')
plt.show(sb.heatmap(stackPower5_nobkg, annot = False, cmap = 'coolwarm'))
print('background = ' + str(bkg5))
plt.title('Det/Inc wo background 6')
plt.show(sb.heatmap(stackPower6_nobkg, annot = False, cmap = 'coolwarm'))
print('background = ' + str(bkg6))
plt.title('Layers 1')
plt.show(sb.heatmap(stackLayers1, annot = False, cmap='coolwarm'))
print('Avg layers = ' + str(stackLayers1.mean().mean()))
plt.title('Layers 2')
plt.show(sb.heatmap(stackLayers2, annot = False, cmap='coolwarm'))
print('Avg layers = ' + str(stackLayers2.mean().mean()))
plt.title('Layers 3')
plt.show(sb.heatmap(stackLayers3, annot = False, cmap='coolwarm'))
print('Avg layers = ' + str(stackLayers3.mean().mean()))
plt.title('Layers 4')
#plt.show(sb.heatmap(stackLayers4, annot = False, cmap='coolwarm'))
print('Avg layers = ' + str(stackLayers4.mean().mean()))
plt.title('Layers 5')
plt.show(sb.heatmap(stackLayers5, annot = False, cmap='coolwarm'))
print('Avg layers = ' + str(stackLayers5.mean().mean()))
plt.title('Layers 6')
plt.show(sb.heatmap(stackLayers6, annot = False, cmap='coolwarm'))
print('Avg layers = ' + str(stackLayers6.mean().mean()))
# +
# %matplotlib inline
# %run get_heatmap.ipynb
# Code to calculate number of layers of graphene transferred as measured between two quartz slides as a funciton of alpha
# and c
#Import necessary packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sb
def trans_curve(x, alpha, c):
# where x is film thickness [cm], y is optical transmission, aplha is absorption coefficient [1/cm], and c is both
# back and front reflection plus other constants
return(c + np.exp(-alpha*x))
def getC(transmission, alpha, thickness):
return(transmission-np.exp(-alpha*thickness))
def getAlpha(transmission, c, thickness):
return(-np.log(transmission-c)/thickness)
# Load heatmap data
path ='C:\\Users\\acarr.BNL\\Documents\\Data\\Optical Transmission\\';
samp='Gr_p_PE_hexane_O_2slides_fixed.csv'; stackPower1slides=pd.read_csv(path+'2019_05_08\\'+samp);
samp='Gr_p_PE_hexane_M_2slides_fixed.csv'; stackPower2slides=pd.read_csv(path+'2019_05_08\\'+samp);
samp='Gr_p_PE_hexane_N_2slides_fixed.csv'; stackPower3slides=pd.read_csv(path+'2019_05_08\\'+samp);
samp='Gr_p_PE_hexane_CP_2slides_fixed.csv'; stackPower4slides=pd.read_csv(path+'2019_05_08\\'+samp);
samp='Gr_p_PE_hexane_CQ_2slides_fixed.csv'; stackPower5slides=pd.read_csv(path+'2019_05_08\\'+samp);
samp='Gr_p_PE_hexane_CR_2slides_fixed.csv'; stackPower6slides=pd.read_csv(path+'2019_05_08\\'+samp);
samp='Gr_p_PE_hexane_O_fixed.csv'; stackPower1noslides=pd.read_csv(path+'2019_03_11\\'+samp);
samp='Gr_p_PE_hexane_M_fixed.csv'; stackPower2noslides=pd.read_csv(path+'2019_03_11\\'+samp);
samp='Gr_p_PE_hexane_N_fixed.csv'; stackPower3noslides=pd.read_csv(path+'2019_03_11\\'+samp);
samp='Gr_p_PE_hexane_CP_fixed.csv'; stackPower4noslides=pd.read_csv(path+'2019_03_18\\'+samp);
samp='Gr_p_PE_hexane_CQ_fixed.csv'; stackPower5noslides=pd.read_csv(path+'2019_03_18\\'+samp);
samp='Gr_p_PE_hexane_CR_fixed.csv'; stackPower6noslides=pd.read_csv(path+'2019_03_18\\'+samp);
percentSlidesC1=[];percentSlidesC2=[];percentSlidesC3=[];percentSlidesC4=[];percentSlidesC5=[];percentSlidesC6=[];
percentSlidesAlpha1=[];percentSlidesAlpha2=[];percentSlidesAlpha3=[];
percentSlidesAlpha4=[];percentSlidesAlpha5=[];percentSlidesAlpha6=[];
percentNoSlidesC1=[];percentNoSlidesC2=[];percentNoSlidesC3=[];
percentNoSlidesC4=[];percentNoSlidesC5=[];percentNoSlidesC6=[];
percentNoSlidesAlpha1=[];percentNoSlidesAlpha2=[];percentNoSlidesAlpha3=[];
percentNoSlidesAlpha4=[];percentNoSlidesAlpha5=[];percentNoSlidesAlpha6=[];
avgNoSlidesAlpha=[];avgNoSlidesC=[];avgSlidesAlpha=[];avgSlidesC=[];
c=0.1; alpha=50;
for n in range(0,100,1):
A=pd.DataFrame();B=A;C=A;D=A;E=A;F=A;
bkg1=trans_curve(0.0163, n, c);stackLayers1slides=(bkg1-stackPower1slides)/0.046;
bkg2=trans_curve(0.0132, n, c);stackLayers2slides=(bkg2-stackPower2slides)/0.046;
bkg3=trans_curve(0.0201, n, c);stackLayers3slides=(bkg3-stackPower3slides)/0.046;
bkg4=trans_curve(0.0189, n, c);stackLayers4slides=(bkg4-stackPower4slides)/0.046;
bkg5=trans_curve(0.0151, n, c);stackLayers5slides=(bkg5-stackPower5slides)/0.046;
bkg6=trans_curve(0.0180, n, c);stackLayers6slides=(bkg6-stackPower6slides)/0.046;
A=stackLayers1slides[stackLayers1slides>0]; A=A[A<2]; A=A.values.flatten();
percentSlidesAlpha1.append((len(A[~np.isnan(A)])/len(A)*100));
B=stackLayers2slides[stackLayers2slides>0]; B=B[B<2]; B=B.values.flatten();
percentSlidesAlpha2.append((len(B[~np.isnan(B)])/len(B)*100));
C=stackLayers3slides[stackLayers3slides>0]; C=C[C<2]; C=C.values.flatten();
percentSlidesAlpha3.append((len(C[~np.isnan(C)])/len(C)*100))
D=stackLayers4slides[stackLayers4slides>0]; D=D[D<2]; D=D.values.flatten();
percentSlidesAlpha4.append((len(D[~np.isnan(D)])/len(D)*100));
E=stackLayers5slides[stackLayers5slides>0]; E=E[E<2]; E=E.values.flatten();
percentSlidesAlpha5.append((len(E[~np.isnan(E)])/len(E)*100));
F=stackLayers6slides[stackLayers6slides>0]; F=F[F<2]; F=F.values.flatten();
percentSlidesAlpha6.append((len(F[~np.isnan(F)])/len(F)*100))
A=pd.DataFrame();B=A;C=A;D=A;E=A;F=A;
stackLayers1noslides=(bkg1-stackPower1noslides)/0.046;
stackLayers2noslides=(bkg2-stackPower2noslides)/0.046;
stackLayers3noslides=(bkg3-stackPower3noslides)/0.046;
stackLayers4noslides=(bkg4-stackPower4noslides)/0.046;
stackLayers5noslides=(bkg5-stackPower5noslides)/0.046;
stackLayers6noslides=(bkg6-stackPower6noslides)/0.046;
A=stackLayers1noslides[stackLayers1noslides>0]; A=A[A<2]; A=A.values.flatten();
percentNoSlidesAlpha1.append((len(A[~np.isnan(A)])/len(A)*100));
B=stackLayers2noslides[stackLayers2noslides>0]; B=B[B<2]; B=B.values.flatten();
percentNoSlidesAlpha2.append((len(B[~np.isnan(B)])/len(B)*100));
C=stackLayers3noslides[stackLayers3noslides>0]; C=C[C<2]; C=C.values.flatten();
percentNoSlidesAlpha3.append((len(C[~np.isnan(C)])/len(C)*100))
D=stackLayers4noslides[stackLayers4noslides>0]; D=D[D<2]; D=D.values.flatten();
percentNoSlidesAlpha4.append((len(D[~np.isnan(D)])/len(D)*100));
E=stackLayers5noslides[stackLayers5noslides>0]; E=E[E<2]; E=E.values.flatten();
percentNoSlidesAlpha5.append((len(E[~np.isnan(E)])/len(E)*100));
F=stackLayers6noslides[stackLayers6noslides>0]; F=F[F<2]; F=F.values.flatten();
percentNoSlidesAlpha6.append((len(F[~np.isnan(F)])/len(F)*100))
for n in np.linspace(0,1,100):
A=pd.DataFrame();B=A;C=A;D=A;E=A;F=A;
bkg1=trans_curve(0.0163, alpha, n);stackLayers1slides=(bkg1-stackPower1slides)/0.046;
bkg2=trans_curve(0.0132, alpha, n);stackLayers2slides=(bkg2-stackPower2slides)/0.046;
bkg3=trans_curve(0.0201, alpha, n);stackLayers3slides=(bkg3-stackPower3slides)/0.046;
bkg4=trans_curve(0.0189, alpha, n);stackLayers4slides=(bkg4-stackPower4slides)/0.046;
bkg5=trans_curve(0.0151, alpha, n);stackLayers5slides=(bkg5-stackPower5slides)/0.046;
bkg6=trans_curve(0.0180, alpha, n);stackLayers6slides=(bkg6-stackPower6slides)/0.046;
A=stackLayers1slides[stackLayers1slides>0]; A=A[A<2]; A=A.values.flatten();
percentSlidesC1.append((len(A[~np.isnan(A)])/len(A)*100));
B=stackLayers2slides[stackLayers2slides>0]; B=B[B<2]; B=B.values.flatten();
percentSlidesC2.append((len(B[~np.isnan(B)])/len(B)*100));
C=stackLayers3slides[stackLayers3slides>0]; C=C[C<2]; C=C.values.flatten();
percentSlidesC3.append((len(C[~np.isnan(C)])/len(C)*100))
D=stackLayers4slides[stackLayers4slides>0]; D=D[D<2]; D=D.values.flatten();
percentSlidesC4.append((len(D[~np.isnan(D)])/len(D)*100));
E=stackLayers5slides[stackLayers5slides>0]; E=E[E<2]; E=E.values.flatten();
percentSlidesC5.append((len(E[~np.isnan(E)])/len(E)*100));
F=stackLayers6slides[stackLayers6slides>0]; F=F[F<2]; F=F.values.flatten();
percentSlidesC6.append((len(F[~np.isnan(F)])/len(F)*100))
A=pd.DataFrame();B=A;C=A;D=A;E=A;F=A;
stackLayers1noslides=(bkg1-stackPower1noslides)/0.046;
stackLayers2noslides=(bkg2-stackPower2noslides)/0.046;
stackLayers3noslides=(bkg3-stackPower3noslides)/0.046;
stackLayers4noslides=(bkg4-stackPower4noslides)/0.046;
stackLayers5noslides=(bkg5-stackPower5noslides)/0.046;
stackLayers6noslides=(bkg6-stackPower6noslides)/0.046;
A=stackLayers1noslides[stackLayers1noslides>0]; A=A[A<2]; A=A.values.flatten();
percentNoSlidesC1.append((len(A[~np.isnan(A)])/len(A)*100));
B=stackLayers2noslides[stackLayers2noslides>0]; B=B[B<2]; B=B.values.flatten();
percentNoSlidesC2.append((len(B[~np.isnan(B)])/len(B)*100));
C=stackLayers3noslides[stackLayers3noslides>0]; C=C[C<2]; C=C.values.flatten();
percentNoSlidesC3.append((len(C[~np.isnan(C)])/len(C)*100))
D=stackLayers4noslides[stackLayers4noslides>0]; D=D[D<2]; D=D.values.flatten();
percentNoSlidesC4.append((len(D[~np.isnan(D)])/len(D)*100));
E=stackLayers5noslides[stackLayers5noslides>0]; E=E[E<2]; E=E.values.flatten();
percentNoSlidesC5.append((len(E[~np.isnan(E)])/len(E)*100));
F=stackLayers6noslides[stackLayers6noslides>0]; F=F[F<2]; F=F.values.flatten();
percentNoSlidesC6.append((len(F[~np.isnan(F)])/len(F)*100))
percentNoSlidesAlpha1=np.asarray(percentNoSlidesAlpha1); percentNoSlidesC1=np.asarray(percentNoSlidesC1);
percentNoSlidesAlpha2=np.asarray(percentNoSlidesAlpha2); percentNoSlidesC2=np.asarray(percentNoSlidesC2);
percentNoSlidesAlpha3=np.asarray(percentNoSlidesAlpha3); percentNoSlidesC3=np.asarray(percentNoSlidesC3);
percentNoSlidesAlpha4=np.asarray(percentNoSlidesAlpha4); percentNoSlidesC4=np.asarray(percentNoSlidesC4);
percentNoSlidesAlpha5=np.asarray(percentNoSlidesAlpha5); percentNoSlidesC5=np.asarray(percentNoSlidesC5);
percentNoSlidesAlpha6=np.asarray(percentNoSlidesAlpha6); percentNoSlidesC6=np.asarray(percentNoSlidesC6);
percentSlidesAlpha1=np.asarray(percentSlidesAlpha1); percentSlidesC1=np.asarray(percentSlidesC1);
percentSlidesAlpha2=np.asarray(percentSlidesAlpha2); percentSlidesC2=np.asarray(percentSlidesC2);
percentSlidesAlpha3=np.asarray(percentSlidesAlpha3); percentSlidesC3=np.asarray(percentSlidesC3);
percentSlidesAlpha4=np.asarray(percentSlidesAlpha4); percentSlidesC4=np.asarray(percentSlidesC4);
percentSlidesAlpha5=np.asarray(percentSlidesAlpha5); percentSlidesC5=np.asarray(percentSlidesC5);
percentSlidesAlpha6=np.asarray(percentSlidesAlpha6); percentSlidesC6=np.asarray(percentSlidesC6);
fig, ax = plt.subplots(2,2, figsize=(15,15));
ax[0,0].set_xlabel('alpha');ax[1,0].set_xlabel('alpha');ax[0,1].set_xlabel('c');ax[1,1].set_xlabel('c');
ax[0,0].set_ylabel('0 < % layers transferred < 2');ax[0,1].set_ylabel('0 < % layers transferred < 2');
ax[1,0].set_ylabel('0 < % layers transferred < 2');ax[1,1].set_ylabel('0 < % layers transferred < 2');
cX = np.linspace(0,1,100); alphaX = np.arange(0,100,1);
ax[0,0].scatter(alphaX,percentNoSlidesAlpha1,color='C0'); ax[0,1].scatter(cX, percentNoSlidesC1,color='C0');
ax[0,0].scatter(alphaX,percentNoSlidesAlpha2,color='C1'); ax[0,1].scatter(cX, percentNoSlidesC2,color='C1');
ax[0,0].scatter(alphaX,percentNoSlidesAlpha3,color='C2'); ax[0,1].scatter(cX, percentNoSlidesC3,color='C2');
ax[0,0].scatter(alphaX,percentNoSlidesAlpha4,color='C3'); ax[0,1].scatter(cX, percentNoSlidesC4,color='C3');
ax[0,0].scatter(alphaX,percentNoSlidesAlpha5,color='C4'); ax[0,1].scatter(cX, percentNoSlidesC5,color='C4');
ax[0,0].scatter(alphaX,percentNoSlidesAlpha6,color='C5'); ax[0,1].scatter(cX, percentNoSlidesC6,color='C5');
ax[1,0].scatter(alphaX,percentSlidesAlpha1,color='C0'); ax[1,1].scatter(cX, percentSlidesC1,color='C0');
ax[1,0].scatter(alphaX,percentSlidesAlpha2,color='C1'); ax[1,1].scatter(cX, percentSlidesC2,color='C1');
ax[1,0].scatter(alphaX,percentSlidesAlpha3,color='C2'); ax[1,1].scatter(cX, percentSlidesC3,color='C2');
ax[1,0].scatter(alphaX,percentSlidesAlpha4,color='C3'); ax[1,1].scatter(cX, percentSlidesC4,color='C3');
ax[1,0].scatter(alphaX,percentSlidesAlpha5,color='C4'); ax[1,1].scatter(cX, percentSlidesC5,color='C4');
ax[1,0].scatter(alphaX,percentSlidesAlpha6,color='C5'); ax[1,1].scatter(cX, percentSlidesC6,color='C5');
ax[0,0].legend(('1','2','3','4','5','6'))
avgSlidesAlpha.append(alphaX[[i for i, j in enumerate(percentSlidesAlpha1) if j == percentSlidesAlpha1.max()]]);
avgSlidesAlpha.append(alphaX[[i for i, j in enumerate(percentSlidesAlpha2) if j == percentSlidesAlpha2.max()]]);
avgSlidesAlpha.append(alphaX[[i for i, j in enumerate(percentSlidesAlpha3) if j == percentSlidesAlpha3.max()]]);
avgSlidesAlpha.append(alphaX[[i for i, j in enumerate(percentSlidesAlpha4) if j == percentSlidesAlpha4.max()]]);
avgSlidesAlpha.append(alphaX[[i for i, j in enumerate(percentSlidesAlpha5) if j == percentSlidesAlpha5.max()]]);
avgSlidesAlpha.append(alphaX[[i for i, j in enumerate(percentSlidesAlpha6) if j == percentSlidesAlpha6.max()]]);
avgNoSlidesAlpha.append(alphaX[[i for i, j in enumerate(percentNoSlidesAlpha1) if j == percentNoSlidesAlpha1.max()]]);
avgNoSlidesAlpha.append(alphaX[[i for i, j in enumerate(percentNoSlidesAlpha2) if j == percentNoSlidesAlpha2.max()]]);
avgNoSlidesAlpha.append(alphaX[[i for i, j in enumerate(percentNoSlidesAlpha3) if j == percentNoSlidesAlpha3.max()]]);
avgNoSlidesAlpha.append(alphaX[[i for i, j in enumerate(percentNoSlidesAlpha4) if j == percentNoSlidesAlpha4.max()]]);
avgNoSlidesAlpha.append(alphaX[[i for i, j in enumerate(percentNoSlidesAlpha5) if j == percentNoSlidesAlpha5.max()]]);
avgNoSlidesAlpha.append(alphaX[[i for i, j in enumerate(percentNoSlidesAlpha6) if j == percentNoSlidesAlpha6.max()]]);
avgSlidesC.append(cX[[i for i, j in enumerate(percentSlidesC1) if j == percentSlidesC1.max()]]);
avgSlidesC.append(cX[[i for i, j in enumerate(percentSlidesC2) if j == percentSlidesC2.max()]]);
avgSlidesC.append(cX[[i for i, j in enumerate(percentSlidesC3) if j == percentSlidesC3.max()]]);
avgSlidesC.append(cX[[i for i, j in enumerate(percentSlidesC4) if j == percentSlidesC4.max()]]);
avgSlidesC.append(cX[[i for i, j in enumerate(percentSlidesC5) if j == percentSlidesC5.max()]]);
avgSlidesC.append(cX[[i for i, j in enumerate(percentSlidesC6) if j == percentSlidesC6.max()]]);
avgNoSlidesC.append(cX[[i for i, j in enumerate(percentNoSlidesC1) if j == percentNoSlidesC1.max()]]);
avgNoSlidesC.append(cX[[i for i, j in enumerate(percentNoSlidesC2) if j == percentNoSlidesC2.max()]]);
avgNoSlidesC.append(cX[[i for i, j in enumerate(percentNoSlidesC3) if j == percentNoSlidesC3.max()]]);
avgNoSlidesC.append(cX[[i for i, j in enumerate(percentNoSlidesC4) if j == percentNoSlidesC4.max()]]);
avgNoSlidesC.append(cX[[i for i, j in enumerate(percentNoSlidesC5) if j == percentNoSlidesC5.max()]]);
avgNoSlidesC.append(cX[[i for i, j in enumerate(percentNoSlidesC6) if j == percentNoSlidesC6.max()]]);
avgSlidesC = np.asarray([y for x in avgSlidesC for y in x]);
avgSlidesAlpha = np.asarray([y for x in avgSlidesAlpha for y in x]);
avgNoSlidesC = np.asarray([y for x in avgNoSlidesC for y in x]);
avgNoSlidesAlpha = np.asarray([y for x in avgNoSlidesAlpha for y in x]);
plt.show();
print('avg alpha no slides = '+str(avgNoSlidesAlpha.mean()) +' +/- '+str(avgNoSlidesAlpha.std()));
print('avg alpha slides = '+str(avgSlidesAlpha.mean()) +' +/- '+str(avgSlidesAlpha.std()));
print('avg c no slides = '+str(avgNoSlidesC.mean()) +' +/- '+str(avgNoSlidesC.std()));
print('avg c slides = '+str(avgSlidesC.mean()) +' +/- '+str(avgSlidesC.std()))
# -
| Optical transmission calculate transmission through background.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/BaishaliChetia/CapsNet-Keras/blob/master/c_mnistTf2Normalized.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="jeoBe9bpMlPR"
# Original implementation at:
#
# https://github.com/ageron/handson-ml/blob/master/extra_capsnets.ipynb
#
# Geron's model doesn't use the keras functional API. In the keras functional API, you don't need to give the batchsize.
#
# When you print the model, you get this:
#
# ```
# Layer (type) Output Shape Param #
# _________________________________________________________________
# input (InputLayer) [(None, 28, 28, 1)] 0
# _________________________________________________________________
# conv_layer_1 (Conv2D) (None, 20, 20, 256) 20992
# _________________________________________________________________
# conv_layer_2 (Conv2D) (None, 6, 6, 256) 5308672
# _________________________________________________________________
# reshape_layer_1 (Reshape) (None, 1, 1152, 8) 0
# _________________________________________________________________
# caps1_output_layer (SquashLa (None, 1, 1152, 8) 0
# _________________________________________________________________
# Total params: 5,329,664
# Trainable params: 5,329,664
# Non-trainable params: 0
# ```
#
# Notice that the Input-layer has shape (None, 28, 28, 1), but we only specified (28, 28, 1). It added None implicitly and that takes care of the batch.
#
# So for anywhere Geron uses the batch size explicitly, you don't need to do anything and tensorflow will take care of.
#
# Also note that tensorflow 1 APIs are still provided with the compat layer. I used the reduce_sum from TF1 in the squash layer, that allowed me to use Geron's code.
#
# Documentation on how to migrate from TF1 to TF2 can be found here:
#
# https://www.tensorflow.org/guide/migrate
#
# + colab={"base_uri": "https://localhost:8080/"} id="sY0OS1x1qLp2" outputId="479f7f3d-a536-437a-a69d-52ef31ff7e64"
from google.colab import drive
drive.mount('/content/drive')
# + id="KCCmDd7lMlPU"
import numpy as np
import tensorflow as tf
import pandas as pd
import tensorflow.keras as K
# + id="-UydL5gJMlPV"
caps1_n_maps = 32
caps1_n_caps = caps1_n_maps * 6 * 6 # 1152 primary capsules
caps1_n_dims = 8
caps2_n_caps = 10
caps2_n_dims = 16
tf.random.set_seed(500000)
# + id="sukwGEY4MlPV"
class SquashLayer(K.layers.Layer):
def __init__(self, axis=-1, **kwargs):
super(SquashLayer, self).__init__(**kwargs)
self.axis = axis
def build(self, input_shapes):
pass
def call(self, inputs):
EPSILON = 1.0e-9
squared_norm = tf.compat.v1.reduce_sum(tf.square(inputs),\
axis=self.axis,\
keepdims=True)
safe_norm = tf.sqrt(squared_norm + EPSILON)
squash_factor = squared_norm / (1. + squared_norm)
unit_vector = inputs / safe_norm
return squash_factor * unit_vector
def get_config(self):
config = super(SquashLayer, self).get_config()
config.update({"axis": self.axis})
return config
# + id="T_qIVI_u2i-s"
class SafeNorm(K.layers.Layer):
def __init__(self, axis=-1, keep_dims = False, **kwargs):
super(SafeNorm, self).__init__(**kwargs)
self.axis = axis
self.keep_dims = keep_dims
def build(self, input_shapes):
pass
def call(self, input):
EPSILON = 1.0e-9
squared_norm = tf.compat.v1.reduce_sum(tf.square(inputs),\
axis=self.axis,\
keepdims= self.keep_dims)
safe_norm = tf.sqrt(squared_norm + EPSILON)
return safe_norm
def get_config(self):
config = super(SafeNorm, self).get_config()
config.update({"axis": self.axis, "keep_dims": self.keep_dims})
return config
# + id="qECOObckMlPW"
# This should be the part where the digit layer, and where we tile things
# This is incomplete, and work in progress
# TODO: Complete this
class MyDigitCapsLayer(K.layers.Layer):
def __init__(self, **kwargs):
super(MyDigitCapsLayer, self).__init__(**kwargs)
def build(self, input_shapes):
init_sigma = 0.1 # TODO: use
self.kernel = self.add_weight(\
"kernel",\
(caps1_n_caps, caps2_n_caps, caps2_n_dims, caps1_n_dims),\
initializer="random_normal",\
dtype=tf.float32)
# To debug this function, I used prints to print the shape
# expand_dims just adds an exis, so if you say expand_dims(inshape=(5, 3), -1),
# you get the output shape (5, 3, 1), it just adds an axis at the end
# Then tile just multiplies one of the dimensions (that is it stacks along that direction N times)
# so tile(inshape=(5, 3, 1), [1, 1, 1000]) will yield a shape (5, 3, 1000)
#
# Notice I didn't tile in build, but in call, Most probaly this is the right thing to do
# but we'll only figure out when we actually train
def call(self, inputs):
# Add a dimension at the end
exp1 = tf.expand_dims(inputs, -1, name="caps1_output_expanded")
# add a dimension along 3rd axis
exp1 = tf.expand_dims(exp1, 2, name="caps2_output_espanced")
# tile along 3rd axis
tile = tf.tile(exp1, [1, 1, caps2_n_caps, 1, 1], name="caps1_output_tiled")
caps2_predicted = tf.matmul(self.kernel, tile, name="caps2_predicted")
return caps2_predicted
def get_config(self):
return super(MyDigitCapsLayer, self).get_config()
# + id="Pg6qxAU3h0hv"
# https://www.tensorflow.org/api_docs/python/tf/keras/losses/Loss
class MarginLoss(K.losses.Loss):
def __init__(self, **kwargs):
super(MarginLoss, self).__init__(**kwargs)
def get_config(self):
config = super(MarginLoss, self).get_config()
return config
def safe_norm(self, input, axis=-2, epsilon=1e-7, keep_dims=False, name=None):
squared_norm = tf.reduce_sum(tf.square(input), axis=axis,
keepdims=keep_dims)
return tf.sqrt(squared_norm + epsilon)
def call(self,y_true, input):
# print(f"y_true.shape = {y_true.shape}, y_pred.shape = {y_pred.shape}")
# return K.losses.MeanSquaredError()(y_true, y_pred)
#y_true = K.Input(shape=[], dtype=tf.int64, name="y")
m_plus = 0.9
m_minus = 0.1
lambda_ = 0.5
#y_true one hot encode y_train
T = tf.one_hot(y_true, depth=caps2_n_caps, name="T")
caps2_output_norm = self.safe_norm(input, keep_dims = True)
present_error_raw = tf.square(\
tf.maximum(0., m_plus - caps2_output_norm),
name="present_error_raw")
present_error = tf.reshape(\
present_error_raw, shape=(-1, 10),
name="present_error")
absent_error_raw = tf.square(\
tf.maximum(0., caps2_output_norm - m_minus),
name="absent_error_raw")
absent_error = tf.reshape(\
absent_error_raw, shape=(-1, 10),
name="absent_error")
L = tf.add(\
T * present_error,\
lambda_ * (1.0 - T) * absent_error,
name="L")
marginLoss = tf.reduce_mean(\
tf.reduce_sum(L, axis=1),\
name="margin_loss")
return marginLoss
# + id="kpXMBYOeWlDd"
class RoutingByAgreement(K.layers.Layer):
def __init__(self, round_number=-1, **kwargs):
super(RoutingByAgreement, self).__init__(**kwargs)
self.round_number = round_number
def get_config(self):
config = super(RoutingByAgreement, self).get_config()
config.update({"round_number": self.round_number})
return config
def build(self, input_shapes):
self.raw_weights_1 = self.add_weight("raw_weights", \
(caps1_n_caps, caps2_n_caps, 1, 1), \
initializer = "zeros", \
dtype=tf.float32,)
#print("Routing layer: self.raw_weights = ", self.raw_weights.shape, "input_shape = ", input_shapes)
@staticmethod
def squash(inputs):
EPSILON = 1.0e-7
squared_norm = tf.compat.v1.reduce_sum(tf.square(inputs),\
keepdims=True)
safe_norm = tf.sqrt(squared_norm + EPSILON)
squash_factor = squared_norm / (1. + squared_norm)
unit_vector = inputs / safe_norm
return squash_factor * unit_vector
def single_round_routing(self, inputs, raw_weights, agreement):
raw_weights = tf.add(raw_weights, agreement)
routing_wt = tf.nn.softmax(raw_weights, axis=2)
wt_predictions = tf.multiply(routing_wt, inputs)
wt_sum = tf.reduce_sum(wt_predictions, axis=1, keepdims=True)
return wt_sum
def call(self, inputs):
agreement = tf.zeros(shape=self.raw_weights_1.shape)
sqsh_wt_sum = None
x = inputs
for i in range(2):
wt_sum = self.single_round_routing(inputs, self.raw_weights_1, agreement)
sqsh_wt_sum = RoutingByAgreement.squash(wt_sum)
sqsh_wt_sum_tiled = tf.tile(\
sqsh_wt_sum ,\
[1, caps1_n_caps, 1, 1, 1],\
name="caps2_output_round_1_tiled")
agreement = tf.matmul(\
x, \
sqsh_wt_sum_tiled,\
transpose_a=True,\
name="agreement")
return sqsh_wt_sum
# + id="aSEe-231jn49" colab={"base_uri": "https://localhost:8080/"} outputId="219103f6-b3e2-49e0-fe5c-b1342f057708"
(x_train, y_train,), (x_test, y_test) = K.datasets.mnist.load_data()
print(x_train.shape, x_test.shape)
# + id="fcHaaMo8db4K"
x_train = x_train/255.0
x_test = x_test/255.0
#print(x_train[500])
# + colab={"base_uri": "https://localhost:8080/"} id="UnmSudqTMlPX" outputId="9edc7ce7-b73f-492f-d134-0278ec19ff80"
class Model:
@staticmethod
def build(inshape=(28, 28, 1)):
inp = K.Input(shape=inshape, dtype=tf.float32, name='input')
# Primary capsules
# For each digit in the batch
# 32 maps, each 6x6 grid of 8 dimensional vectors
# First Conv layer
conv1_params = \
{
"filters": 256,
"kernel_size": 9,
"strides": 1,
"padding": "valid",
"activation": tf.nn.relu,
}
x = K.layers.Conv2D(**conv1_params, name="conv_layer_1")(inp)
# Second conv layer
conv2_params = \
{
"filters": caps1_n_maps * caps1_n_dims, # 256 convolutional filters
"kernel_size": 9,
"strides": 2,
"padding": "valid",
"activation": tf.nn.relu
}
x = K.layers.Conv2D(**conv2_params, name="conv_layer_2")(x)
# Reshape
x = K.layers.Reshape(\
(caps1_n_caps, caps1_n_dims),\
name="reshape_layer_1")(x)
x = SquashLayer(name="caps1_output_layer")(x)
x = MyDigitCapsLayer(name = "caps2_predicted")(x)
caps2_predicted = x # Save this value for later
#routing by agreement (2 rounds)
x = RoutingByAgreement(name="routing1", round_number=2)(x)
return K.Model(inputs=inp, outputs=x, name='my')
m = Model.build()
print(m.summary())
# + id="lw3j_zWQMlPZ" colab={"base_uri": "https://localhost:8080/"} outputId="967dfad9-cd0e-4951-d59a-d22267c2461d"
y_train_train = tf.one_hot(y_train, depth=caps2_n_caps, name="T")
print(y_train_train.shape)
#print(y_train)
# + id="6zxRXCYIz_HQ"
class MyAccuracy(K.metrics.Metric):
def __init__(self, **kwargs):
super(MyAccuracy, self).__init__(**kwargs)
self.acc_obj = None
self.state = 0
def get_config(self):
config = super(MyAccuracy, self).get_config()
config.update({"acc_obj": None, "state": self.state})
return config
def safe_norm(self, input, axis=-2, epsilon=1e-7, keep_dims=True, name=None):
squared_norm = tf.reduce_sum(tf.square(input), axis=axis,
keepdims=keep_dims)
return tf.sqrt(squared_norm + epsilon)
def update_state(self, y_true, input, sample_weight=None):
if self.acc_obj is None:
self.acc_obj = K.metrics.Accuracy()
y_proba = self.safe_norm(input, axis=-2)
y_proba_argmax = tf.argmax(y_proba, axis=2)
y_pred = tf.squeeze(y_proba_argmax, axis=[1,2], name="y_pred")
y_true = tf.reshape(y_true, (y_true.shape[0], ))
y_true = tf.cast(y_true, dtype=tf.int64)
self.acc_obj.update_state(y_true, y_pred, sample_weight)
def reset_state(self):
self.acc_obj.reset_state()
def result(self):
return self.acc_obj.result()
# + id="ohNmlixkrAek"
from keras.callbacks import ModelCheckpoint, CSVLogger
comparison_metric = MyAccuracy()
#checkpoint_filepath = "/content/drive/MyDrive/Weights/weights-improvement-{epoch:02d}-{val_my_accuracy:.2f}.hdf5"
model_checkpoint_callback = tf.keras.callbacks.ModelCheckpoint(
filepath = "/content/drive/MyDrive/WeightsMnist/11_best_weights1.hdf5",
save_weights_only=True,
monitor=f"val_{comparison_metric.name}",
mode='max',
save_best_only=True)
model_checkpoint_callback2 = tf.keras.callbacks.ModelCheckpoint(
filepath = "/content/drive/MyDrive/WeightsMnist/11_latest_weights1.hdf5",
save_weights_only=True,
monitor=f"val_{comparison_metric.name}",
mode='max',
save_best_only=False)
log_csv = CSVLogger("/content/drive/MyDrive/WeightsMnist/11_mylogs1.csv", separator = ",", append = False)
callback_list = [model_checkpoint_callback, model_checkpoint_callback2, log_csv]
# + id="cahMgtOqMlPa"
m.compile(optimizer='adam', loss=MarginLoss(), metrics=[MyAccuracy()])
history = m.fit(x_train, y_train, batch_size=32, epochs=2, verbose= 1, validation_split=0.2, callbacks = callback_list)
# + id="tyUFx-dcMlPa" colab={"base_uri": "https://localhost:8080/", "height": 605} outputId="b0fffd32-2c91-49eb-e7f6-7d383d673cab"
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (30, 10)
plt.rcParams["font.size"] = 20
fig, ax = plt.subplots(1, 2)
ax[0].plot(history.history['my_accuracy_1'])
ax[0].plot(history.history['val_my_accuracy_1'])
ax[0].set_title('Model Accuracy')
ax[0].set_ylabel('Accuracy')
ax[0].set_xlabel('Epoch')
ax[0].legend(['Training Accuracy', 'Validation Accuracy'], loc='best')
ax[1].plot(history.history['loss'])
ax[1].plot(history.history['val_loss'])
ax[1].set_title('Model Loss')
ax[1].set_ylabel('Loss')
ax[1].set_xlabel('Epoch')
ax[1].legend(['Training Loss', 'Validation Loss'], loc='best')
plt.show()
# + id="8bnBQabNNKLH"
print(f'Best Validation Accuracy = {np.max(history.history["val_my_accuracy_1"])}')
print(f'Best Training Accuracy = {np.max(history.history["my_accuracy_1"])}')
# + id="9pOcuSJGSvLY"
m.save("/content/drive/MyDrive/WeightsMnist/save.tf", save_format='tf')
# + id="WLeqoVJkjPKB"
#Extra layer for evaluate
class DimensionCorrection(K.layers.Layer):
def __init__(self, **kwargs):
super(DimensionCorrection, self).__init__(**kwargs)
def safe_norm(self, input, axis=-2, epsilon=1e-7, keep_dims=False, name=None):
squared_norm = tf.reduce_sum(tf.square(input), axis=axis,
keepdims=keep_dims)
return tf.sqrt(squared_norm + epsilon)
def call(self,y_pred):
y_proba = self.safe_norm(y_pred, axis=-2)
y_proba_argmax = tf.argmax(y_proba, axis=2)
y_pred = tf.squeeze(y_proba_argmax, axis=[1,2], name="y_pred")
return y_pred
# + colab={"base_uri": "https://localhost:8080/"} id="R3s5KL2po5FN" outputId="b596dfb9-e99a-4612-d596-90ac73cb004b"
y_test = tf.cast(y_test, dtype= tf.int64)
print(y_test.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 816} id="8cJcH3YlsAVt" outputId="0b4f9bca-a978-4fc8-8fda-d13e76db066c"
m = Model.build()
m.load_weights('/content/drive/MyDrive/WeightsMnist/latest_weights1.hdf5')
m.compile(optimizer='Adam', loss=MarginLoss)
newmodel = K.models.Sequential(\
[\
m,\
DimensionCorrection(),\
]\
)
newmodel.summary()
m.trainable = False
newmodel.compile(optimizer='adam')
y_pred = newmodel.predict(x_test)
import sklearn
from sklearn.metrics import confusion_matrix, accuracy_score, roc_auc_score
print(confusion_matrix(y_test, y_pred))
print(f"accuracy = {accuracy_score(y_test, y_pred)}")
# + id="_lbZNCJZkZ46"
y_pred_eval = DimensionCorrection()
m2 = Model.build()
m2.load_weights('/content/drive/MyDrive/WeightsMnist/latest_weights1.hdf5')
# + id="N-vE6yGAoBPI"
m3 = K.models.Sequential()
m3.add(m2)
m3.add(y_pred_eval)
m3.build()
m3.compile(optimizer='adam', loss=MarginLoss(), metrics=[MyAccuracy()])
m3.evaluate(x_test, y_test, batch_size= 32, verbose= 1)
#m3.evaluate(x_test, y_test, batch_size= 32, verbose= 1)
# + id="h9vf-xeFUcgi"
#converter = tf.lite.TFLiteConverter.from_keras_model(m)
# + id="VuULPnldermd"
# + id="wdrYq9SwUr-N"
# converter.optimizations = [tf.lite.Optimize.DEFAULT]
# converter.target_spec.supported_types = [tf.float16]
# quantize_model = converter.convert()
# + colab={"base_uri": "https://localhost:8080/"} id="d5Vm_QpbUry8" outputId="0ed2fc82-702e-415e-a578-21da15a79b43"
mm = K.models.load_model('/content/drive/MyDrive/WeightsMnist/save.tf',\
custom_objects=\
{\
"SquashLayer": SquashLayer,\
"SafeNorm": SafeNorm,\
"MyDigitCapsLayer": MyDigitCapsLayer,\
"RoutingByAgreement": RoutingByAgreement,\
"MyAccuracy": MyAccuracy,\
"MarginLoss": MarginLoss,\
})
print(type(mm), mm.summary())
print(mm.weights[0][0][0][0][0:5])
e = mm.load_weights('/content/drive/MyDrive/WeightsMnist/latest_weights1.hdf5')
print(mm.weights[0][0][0][0][0:5])
# Create the .tflite file
tflite_model_file = "/content/drive/MyDrive/WeightsMnist/compressed.tflite"
converter = tf.lite.TFLiteConverter.from_keras_model(mm)
converter.optimizations = [tf.lite.Optimize.DEFAULT]
converter.target_spec.supported_ops = [
tf.lite.OpsSet.TFLITE_BUILTINS, # enable TensorFlow Lite ops.
tf.lite.OpsSet.SELECT_TF_OPS # enable TensorFlow ops.
]
tflite_model = converter.convert()
with open(tflite_model_file, "wb") as f:
f.write(tflite_model)
# + [markdown] id="F1c026FiF3Hb"
#
# + colab={"base_uri": "https://localhost:8080/"} id="eVrB1GkTFItQ" outputId="03fe35c9-ab59-4f96-848c-7fdc97f6f0d6"
# !du -sh /content/drive/MyDrive/WeightsMnist/*
# + [markdown] id="cTzAAQHpL51b"
# # Use this tutorial for pruning
# quantization has already been done earlier
#
# https://www.tensorflow.org/model_optimization/guide/pruning/pruning_with_keras
#
# + id="aJLicHf3J1G-"
| c_mnistTf2Normalized.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Imports
from splinter import Browser
from bs4 import BeautifulSoup
import pandas as pd
import time
# !which chromedriver
executable_path = {'executable_path': '/usr/local/bin/chromedriver'}
browser = Browser('chrome', **executable_path, headless=True)
# +
# url = "https://mars.nasa.gov/news/?page=0&per_page=40&order=publish_date+desc%2Ccreated_at+desc&search=&category=19%2C165%2C184%2C204&blank_scope=Latest"
# browser.visit(url)
# html = browser.html
# soup = BeautifulSoup(html, "html.parser")
# # print(soup.prettify())
# # results = soup.find_all('li', class_='slide')
# for result in results:
# try:
# title = result.find(class_='content_title').text
# news_p= result.find(class_='article_teaser_body').text
# if (title and news_p):
# print('-------------------------')
# print(title)
# print(news_p)
# except AttributeError as e:
# print(e)
# -
def init_browser():
executable_path = {'executable_path': '/usr/local/bin/chromedriver'}
return Browser('chrome', **executable_path, headless=True)
# +
def nasa():
browser = init_browser()
url = "https://mars.nasa.gov/news/?page=0&per_page=40&order=publish_date+desc%2Ccreated_at+desc&search=&category=19%2C165%2C184%2C204&blank_scope=Latest"
browser.visit(url)
html = browser.html
soup = BeautifulSoup(html, "html.parser")
results = soup.find_all('li', class_='slide')
news_title=[]
news_p=[]
for result in results:
news_title.append(result.find(class_='content_title').get_text())
news_p.append(result.find(class_='article_teaser_body').get_text())
return news_title, news_p
# -
nasa()
def jpl():
browser = init_browser()
url = 'https://www.jpl.nasa.gov/spaceimages/?search=&category=Mars'
browser.visit(url)
html = browser.html
soup = BeautifulSoup(html, "html.parser")
results = soup.find_all('a', class_='fancybox')
featured_image_url=[]
for result in results:
featured_image_url.append('https://www.jpl.nasa.gov'+result['data-fancybox-href'])
return featured_image_url
jpl()
# +
# url ='https://twitter.com/marswxreport?lang=en'
# browser.visit(url)
# time.sleep(5)
# html = browser.html
# soup = BeautifulSoup(html, "html.parser")
# # print(soup.prettify())
# results3 = soup.find_all('article', attrs ={'role':'article'})
# weather_text=results3[0].find_all('span', attrs = {'class':'css-901oao css-16my406 r-1qd0xha r-ad9z0x r-bcqeeo r-qvutc0'})
# weather_text_cleaned=weather_text[4].text.replace('\n', ' ')
# -
def twit():
url ='https://twitter.com/marswxreport?lang=en'
browser.visit(url)
time.sleep(5)
html = browser.html
soup = BeautifulSoup(html, "html.parser")
results3 = soup.find_all('article', attrs ={'role':'article'})
weather_text=results3[0].find_all('span', attrs = {'class':'css-901oao css-16my406 r-1qd0xha r-ad9z0x r-bcqeeo r-qvutc0'})
weather_text_cleaned=weather_text[4].text.replace('\n', ' ')
return weather_text_cleaned
twit()
url = 'https://space-facts.com/mars/'
tables = pd.read_html(url)
facts=tables[0]
facts
# +
#Removing the colon
facts[0]=facts[0].str.strip(":")
#Adding column labels
facts.columns=['Facts', 'Values']
#Add Index
facts_df=facts.set_index('Facts')
facts_df
# -
html_table=facts_df.to_html()
html_table
facts_df.to_html('mars_fact_table.html')
# +
# def facts():
# url = 'https://space-facts.com/mars/'
# tables = pd.read_html(url)
# facts=tables[0]
# #Remobing the colon
# facts[0]=facts[0].str.strip(":")
# #Adding column labels
# facts.columns=['Facts', 'Values']
# #Add Index
# facts_df=facts.set_index('Facts')
# facts_df
# return facts_df
# +
#Opening Broswer
base_url='https://astrogeology.usgs.gov'
url ='https://astrogeology.usgs.gov/search/results?q=hemisphere+enhanced&k1=target&v1=Mars'
browser.visit(url)
time.sleep(5)
html = browser.html
soup = BeautifulSoup(html, "html.parser")
results4 = soup.find_all('div', class_='description')
title=[]
links=[]
#Saving title/initial address for image
for result in results4:
title.append(result.find('h3').text)
links.append(base_url+result.find('a')['href'])
img_url=[]
#Saving full image link
for link in links:
url = link
browser.visit(url)
html = browser.html
soup = BeautifulSoup(html, "html.parser")
img_url.append(base_url + soup.find('img', class_="wide-image")['src'])
#Creating a dictionary
hemisphere_image_urls ={'title':title,
'img_url':img_url}
# -
| Missions_to_Mars/mission_to_mars.ipynb |