code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pablocarreira-py39] *
# language: python
# name: conda-env-pablocarreira-py39-py
# ---
# <br>
#
# # Introdução
import os
import ast
import json
import folium
import requests
import numpy as np
import pandas as pd
import geopandas as gpd
# +
#import shutil
#import urllib.request
# +
# #!pip install traquitanas --upgrade
# -
import traquitanas.utils as tt
# <br>
#
# ## Paths
# +
# Inicialmente faz-se necessário criar uma pasta que receberão os dados do IBGE
data_path = os.path.join('..', 'data')
input_path = os.path.join(data_path, 'input')
output_path = os.path.join(data_path, 'output')
output_path_geo = os.path.join(output_path, 'geo')
output_path_tab = os.path.join(output_path, 'tab')
os.makedirs(data_path, exist_ok=True)
os.makedirs(input_path, exist_ok=True)
os.makedirs(output_path, exist_ok=True)
os.makedirs(output_path_geo, exist_ok=True)
os.makedirs(output_path_tab, exist_ok=True)
# -
# <br>
#
# # Dados Espaciais
#
# Com a estrutura de pastas criada, é possivel fazer o download dos arquivos disponiblizados pelo IBGE. Há uma infinidade de dados.
#
# - https://servicodados.ibge.gov.br/api/docs
# <br>
#
# ## <NAME>lo
# +
# Define qual o código IBGE do Estado
estado = 35
# Define URL
parameters = {
'formato': 'application/vnd.geo+json',
'resolucao': '5',
'qualidade': '4',
}
url = 'https://servicodados.ibge.gov.br/api/v2/malhas/{}'.format(estado)
r = requests.get(url, params=parameters)
print(r.url)
# Define o nome do arquivo que será salvo as informações do IBGE
geojson_file = os.path.join(input_path, 'sp_ibge.geojson')
# Save
with open(geojson_file, 'wb') as f:
f.write(r.content)
# -
file_encoding = tt.predict_encoding(geojson_file)
file_encoding
# +
gdf = gpd.read_file(
geojson_file
)
gdf.rename(
{'codarea': 'id_ibge'},
axis=1,
inplace=True
)
# Transforma Coordenadas
gdf = gdf.to_crs(epsg=4326)
# Salva Arquivo
gdf.to_file(
os.path.join(output_path_geo, 'sp_igbe.gpkg'),
layer='Limite Municipal',
driver='GPKG',
encoding='utf-8'
)
gdf
# +
# Map Object
m = folium.Map()
# Folium Object
folium.GeoJson(geojson_file).add_to(m)
# Fit and Plot map
m.fit_bounds(m.get_bounds())
m
# -
# <br>
#
# ## São Paulo (Memória)
# <br>
#
# Ou ainda, ao invés de baixar o arquivo, é possivel fazer com o que o mapa seja criado com a leitura dos dados diretamente do site do IBGE. Nessa função o *encoding* já foi definido, evitando o problema mencionado acima.
r = requests.get(url, params=parameters)
geojson = json.loads(r.text)
# +
# Map Object
m = folium.Map()
# Folium Object
folium.GeoJson(geojson).add_to(m)
# Fit and Plot map
m.fit_bounds(m.get_bounds())
m
# -
# <br>
#
# ## Estados
# +
# Define qual o código IBGE do Estado
estado = 'UF'
# Define URL
parameters = {
'formato': 'application/vnd.geo+json',
#'resolucao': '5',
#'qualidade': '4',
}
url = 'https://servicodados.ibge.gov.br/api/v3/malhas/paises/BR?intrarregiao={}'.format(estado)
r = requests.get(url, params=parameters)
print(r.url)
# Define o nome do arquivo que será salvo as informações do IBGE
geojson_file = os.path.join(input_path, 'br_estados_ibge.geojson')
# Save
with open(geojson_file, 'wb') as f:
f.write(r.content)
# +
gdf = gpd.read_file(
geojson_file
)
gdf.rename(
{'codarea': 'id_ibge'},
axis=1,
inplace=True
)
# Transforma Coordenadas
gdf = gdf.to_crs(epsg=4326)
# Salva Arquivo
gdf.to_file(
os.path.join(output_path_geo, 'br_igbe.gpkg'),
layer='Limite Municipal',
driver='GPKG',
encoding='utf-8'
)
gdf.head()
# +
# API
list_estados = []
for index, row in gdf.iterrows():
url = 'https://servicodados.ibge.gov.br/api/v1/localidades/estados/{UF}'.format(UF=row['id_ibge'])
r = requests.get(url)
dict_quotes = r.content.decode(r.encoding)
dict_quotes = ast.literal_eval(dict_quotes)
list_estados.append(dict_quotes)
# Transforma dictionary in Dataframe
df = pd.DataFrame(list_estados)
df.head()
# Adjust nested Dictionary
for i, row in df.iterrows():
df.loc[i, 'id_regiao'] = row['regiao']['id']
df.loc[i, 'sigla_regiao'] = row['regiao']['sigla']
df.loc[i, 'nome_regiao'] = row['regiao']['nome']
# Fim
df.drop(['regiao'], axis=1, inplace=True)
df['id_regiao'] = pd.to_numeric(df['id_regiao'], downcast='integer')
df.head()
# -
df.to_csv(
os.path.join(output_path_tab, 'tab_uf_ibge.csv'),
index=False
)
# +
# Map Object
m = folium.Map()
# Folium Object
folium.GeoJson(geojson_file).add_to(m)
# Fit and Plot map
m.fit_bounds(m.get_bounds())
m
# -
# <br>
#
# Uma vez com o mapa na mão, de qualquer que seja o meio que foi obtido, é possivel analisar a "tabela de atributos".
# Lá descobrimos que existe o par de coordenadas que define o centroide e, ainda, o 'codarea' que tem o código do IBGE do município.
# <br>
#
# # Dados Tabulares
# +
df_ufs = pd.read_csv(
os.path.join(output_path_tab, 'tab_ufs_ibge.csv'),
)
list_dfs = []
for i, row in df_ufs.iterrows():
estado = row['id']
url = 'http://servicodados.ibge.gov.br/api/v1/localidades/estados/{}/municipios'.format(estado)
df = pd.read_json(url)
list_dfs.append(df)
# Ajustes
df = pd.concat(list_dfs)
df = df[['id', 'nome']].copy()
# Salva
df.to_csv(
os.path.join(output_path_tab, 'tab_municipio_ibge.csv'),
index=False
)
df.head()
# +
# Defines IBGE Code State
estado = 35
# Defines URL
url = 'http://servicodados.ibge.gov.br/api/v1/localidades/estados/{}/municipios'.format(estado)
print('{}\n'.format(url))
# Create Dataframe
df = pd.read_json(url)
# Seleciona Colunas
df = df[['id', 'nome']]
# Renomeia Colunas
df = df.rename(columns=lambda x: x.replace('id', 'id_ibge'))
df
# -
# <br>
#
# # Mapa Categórico
# +
# Adjust dtypes
#df['id_ibge'] = df['id_ibge'].apply(lambda x: str(x))
# Add Random Colum
df['random'] = np.random.uniform(1, 6, df.shape[0])
# Results
print('{}\n'.format(df.dtypes))
display(df.head())
# +
# Geodataframe
gdf = gpd.read_file(
os.path.join(output_path_geo, 'sp_igbe.gpkg')
)
# Ajusta os Tipo
gdf['id_ibge'] = pd.to_numeric(gdf['id_ibge'], downcast='integer')
# Results
print('{}\n'.format(gdf.dtypes))
display(gdf.head())
# +
# Create Map
m = folium.Map()
highlight_function = lambda x: {
'fillColor': '#000000',
'color':'#000000',
'fillOpacity': 30,
'weight': 1
}
choropleth = folium.Choropleth(
geo_data=gdf,
name='choropleth',
data=df,
columns=['id_ibge', 'random', 'nome'],
key_on='feature.properties.id_ibge',
fill_color='YlGnBu',
fill_opacity=0.5,
line_opacity=0.1,
legend_name='Legenda',
smooth_factor=0,
show=False,
overlay=True,
highlight=True,
highlight_function=highlight_function,
nan_fill_color='White',
).add_to(m)
# Add Field in geodataframe
choropleth.geojson.add_child(
folium.features.GeoJsonTooltip(['id_ibge'], labels=False)
)
# Fit and Plot map
m.fit_bounds(m.get_bounds())
m
# -
| test/api.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Phase II Modeling
# +
import preprocessing
import wrangle
import model
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import confusion_matrix
from sklearn.metrics import recall_score
# ignore warnings
import warnings
warnings.simplefilter(action='ignore')
import os.path
from os import path
from sklearn.model_selection import cross_val_score, train_test_split, GridSearchCV
import re
from preprocessing_permits import permits_preprocessing_mother_function
import preprocessing_permits as pr
# -
# # Acquire
def rename_columns(df):
"""
Docstring
"""
# rename columns inplace
df.rename(
columns={
"Date": "survey_date",
"Code": "csa_code",
"Code.1": "cbsa_code",
"Unnamed: 3": "moncov",
"Name": "cbsa_name",
"Bldgs": "one_unit_bldgs_est",
"Units": "one_unit_units_est",
"Value": "one_unit_value_est",
"Bldgs.1": "two_units_bldgs_est",
"Units.1": "two_units_units_est",
"Value.1": "two_units_value_est",
"Bldgs.2": "three_to_four_units_bldgs_est",
"Units.2": "three_to_four_units_units_est",
"Value.2": "three_to_four_units_value_est",
"Bldgs.3": "five_or_more_units_bldgs_est",
"Units.3": "five_or_more_units_units_est",
"Value.3": "five_or_more_units_value_est",
"Bldgs.3": "five_or_more_units_bldgs_est",
"Units.3": "five_or_more_units_units_est",
"Value.3": "five_or_more_units_value_est",
"Bldgs.4": "one_unit_bldgs_rep",
"Units.4": "one_unit_units_rep",
"Value.4": "one_unit_value_rep",
"Bldgs.5": "two_units_bldgs_rep",
"Units.5": "two_units_units_rep",
"Value.5": "two_units_value_rep",
" Bldgs": "three_to_four_units_bldgs_rep",
"Units.6": "three_to_four_units_units_rep",
"Value.6": "three_to_four_units_value_rep",
"Bldgs.6": "five_or_more_units_bldgs_rep",
"Units.7": "five_or_more_units_units_rep",
"Value.7": "five_or_more_units_value_rep",
},
inplace=True,
)
return df
def acquire_building_permits():
"""
Docstring
"""
# conditional
if path.exists("building_permits.csv"):
# read csv
df = pd.read_csv("building_permits.csv", index_col=0)
else:
# create original df with 2019 data
df = pd.read_csv("https://www2.census.gov/econ/bps/Metro/ma2019a.txt", sep=",", header=1)
# rename columns
rename_columns(df)
for i in range(1980, 2019):
# read the txt file at url where i is the year in range
year_df = pd.read_csv(
f"https://www2.census.gov/econ/bps/Metro/ma{i}a.txt",
sep=",",
header=1,
names=df.columns.tolist(),
)
# append data to global df variable
df = df.append(year_df, ignore_index=True)
# make moncov into bool so that the null observations of this feature are not considered in the dropna below
df["moncov"] = np.where(df.moncov == "C", 1, 0)
# dropna inplace
df.dropna(inplace=True)
# chop off the succeding two digits after the year for survey_date
df["survey_date"] = df.survey_date.astype(str).apply(lambda x: re.sub(r"\d\d$", "", x))
# add a preceding "19" to any years where the length of the observation is 2 (e.i., "80"-"97")
df["survey_date"] = df.survey_date.apply(lambda x: "19" + x if len(x) == 2 else x)
# turn survey_date back into an int
df["survey_date"] = df.survey_date.astype(int)
# turn moncov back into a bool
df["moncov"] = df.moncov.astype(bool)
# sort values by survey_date
df.sort_values(by=["survey_date"])
# reset index inplace
df.reset_index(inplace=True)
# drop former index inplace
df.drop(columns=["index"], inplace=True)
# write df to disk as csv
df.to_csv("building_permits.csv")
return df
df_model = acquire_building_permits()
print(f"""Our DataFrame contains {df_model.shape[0]:,} observations and {df_model.shape[1]} features.""")
df_model
df_model.sort_values(by=["survey_date"], ascending=False)
df_model.survey_date.min(), df_model.survey_date.max()
df_model.survey_date.value_counts().sort_index(ascending=False)
df_model.groupby("cbsa_name").five_or_more_units_value_rep.mean()
df_model[(df_model.cbsa_name.str.contains("ABILENE"))]
df_model[df_model.survey_date > 1998].sort_values(by = "survey_date", ascending=True).cbsa_name.value_counts().tail(60)
# We need to find the cities that have the most number of continuous data.
(df_model.groupby("cbsa_name").survey_date.count().sort_values() == 17).sum()
df_model[df_model["five_or_more_units_value_est"] == 0.0].groupby("cbsa_name").three_to_four_units_value_rep.sum()
df_model.shape
# # Prepare
# Things that need to get done:
#
# * standarized the name of the cities so that they are all uniform. Need to make sure that when we do a groupby, cities aren't being overlook, or that there are duplicates.
# * Can we do a loop using `str.contains()` to add city names together in an effort to standarized the names. Use a double_split (" ") to create a split between the name and the state
df_model[df_model.cbsa_name.str.contains("Austin") == True]
df_model["city"] = df_model.cbsa_name.str.split(" ", 1, expand = True)[0]
df_model["state"] = df_model.cbsa_name.str.split(" ", 1, expand = True)[1]
df_model
df_model.state.str.split(" ", 1, expand= True)[1].value_counts()
df_model[df_model.state.str.contains(" AR") == True]
df_model[(df_model.city.str.contains("San Antonio") == True) & (df_model.survey_date > 2000)].sort_values(by="survey_date", ascending=True)
df_model["major_city"] = df_model.city.str.split("-", 1, expand=True)[0]
df_model["major_state"] = df_model.state.str.split("-", 1, expand=True)[0]
df_model["metropolitan_area"] = df_model.state.str.split("-", 1, expand=True)[1]
df_model["metropolitan_area"] = df_model.major_state.str.split(" ", 1, expand=True)[1]
df_model["major_state"] = df_model.major_state.str.split(" ", 1, expand=True)[0]
df = df_model.groupby(["major_city","major_state", "survey_date"]).sum().reset_index()
df[df.major_city == "Dallas"]
# +
def prep_building_permits(df_model):
df_model["city"] = df_model.cbsa_name.str.split(" ", 1, expand = True)[0]
df_model["state"] = df_model.cbsa_name.str.split(" ", 1, expand = True)[1]
df_model["major_city"] = df_model.city.str.split("-", 1, expand=True)[0]
df_model["major_state"] = df_model.state.str.split("-", 1, expand=True)[0]
df_model["metropolitan_area"] = df_model.state.str.split("-", 1, expand=True)[1]
df_model["metropolitan_area"] = df_model.major_state.str.split(" ", 1, expand=True)[1]
df_model["major_state"] = df_model.major_state.str.split(" ", 1, expand=True)[0]
df_model = df_model.groupby(["major_city","major_state", "survey_date"]).sum().reset_index()
return df_model
def filter_top_cities_building_permits(df):
df["city_state"] = df["major_city"] + "_" + df["major_state"]
city_mask = df.groupby("city_state").survey_date.count()
city_mask = city_mask[city_mask == 23]
# apply city mask to shrink the df
def in_city_mask(x):
return x in city_mask
df = df[df.city_state.apply(in_city_mask)]
df = df.sort_values(["major_city", "major_state", "survey_date"])
return df
# -
# # Explore
df_model = acquire_building_permits()
df_model = prep_building_permits(df_model)
df_model = filter_top_cities_building_permits(df_model)
df_model.major_state.value_counts().nlargest(10)
# +
# What is the state with the most number of new constructions?
df_model.groupby("major_state").five_or_more_units_units_est.sum().nlargest(10)
# +
# What is the state with the most money in new construction?
df_model.groupby("major_state").five_or_more_units_value_est.sum().nlargest(10)
# +
# Has the number of new buildings change over time?
df_model.groupby("survey_date").five_or_more_units_value_est.mean().plot.line(figsize=(15,5))
plt.title("Has the total value for multifamily housing change over time?")
plt.ylabel("Dolllars")
plt.xlabel("Year")
# +
# Has the value for buildings change over time?
df_model.groupby("survey_date").five_or_more_units_units_est.mean().plot.line(figsize=(15,5))
plt.title("Has the total value for multifamily housing change over time?")
plt.ylabel("Dolllars")
plt.xlabel("Year")
# -
# Are there any patterns in Houston 2009, Seattle 2010 and Dallas 2012?
houston = df_model[df_model.major_city == "Houston"]
plt.figure(figsize=(15,5))
sns.lineplot(data=houston, x="survey_date", y="five_or_more_units_value_est")
plt.title("Money spent in dense multifamily households for Houston")
plt.ylabel("Money Spent On Dense Multifamily households")
plt.xlabel("Year")
plt.axvline(2009, 0,.5, color="r")
plt.axvline(2010, 0,.5, color="r")
plt.axhline(houston.five_or_more_units_value_est.mean(), ls='--', color="Black")
plt.figure(figsize=(15,5))
sns.lineplot(data=houston, x="survey_date", y="five_or_more_units_units_est")
plt.title("Number of units in dense multifamily households for Houston")
plt.ylabel("Number of Dense Multifamily households built")
plt.xlabel("Year")
plt.axvline(2009, 0,.5, color="r")
plt.axvline(2010, 0,.5, color="r")
plt.axhline(houston.five_or_more_units_units_est.mean(), ls='--', color="Black")
# Are there any patterns in Houston 2009, Seattle 2010 and Dallas 2012?
seattle = df_model[df_model.major_city == "Seattle"]
plt.figure(figsize=(15,5))
sns.lineplot(data=seattle, x="survey_date", y="five_or_more_units_value_est")
plt.title("Money spent in dense multifamily households")
plt.ylabel("Money Spent On Dense Multifamily households")
plt.xlabel("Year")
plt.axvline(2010, 0,.5, color="r")
plt.axvline(2011, 0,.5, color="r")
plt.axhline(seattle.five_or_more_units_value_est.mean(), ls='--', color="Black")
plt.figure(figsize=(15,5))
sns.lineplot(data=seattle, x="survey_date", y="five_or_more_units_units_est")
plt.title("Number of units in dense multifamily households")
plt.ylabel("Number of Dense Multifamily households built")
plt.xlabel("Year")
plt.axvline(2010, 0,.5, color="r")
plt.axvline(2011, 0,.5, color="r")
plt.axhline(seattle.five_or_more_units_units_est.mean(), ls='--', color="Black")
# Are there any patterns in Houston 2009, Seattle 2010 and Dallas 2012?
dallas = df_model[df_model.major_city == "Dallas"]
plt.figure(figsize=(15,5))
sns.lineplot(data=dallas, x="survey_date", y="five_or_more_units_value_est")
plt.title("Money spent in dense multifamily households")
plt.ylabel("Money Spent On Dense Multifamily households")
plt.xlabel("Year")
plt.axvline(2011, 0,.5, color="r")
plt.axvline(2012, 0,.5, color="r")
plt.axhline(dallas.five_or_more_units_value_est.mean(), ls='--', color="Black")
plt.figure(figsize=(15,5))
sns.lineplot(data=dallas, x="survey_date", y="five_or_more_units_units_est")
plt.title("Number of units in dense multifamily households")
plt.ylabel("Number of Dense Multifamily households built")
plt.xlabel("Year")
plt.axvline(2011, 0,.5, color="r")
plt.axvline(2012, 0,.5, color="r")
plt.axhline(dallas.five_or_more_units_units_est.mean(), ls='--', color="Black")
def filter_top_cities_building_permits(df):
"""
This function masks df in two ways:
city_mask returns cities with only continuously reported data
threshold_mask returns cities where they had at least one "5 or more unit" building permit for every year
Returns 130 cities for modeling
"""
df["city_state"] = df["major_city"] + "_" + df["major_state"]
city_mask = df.groupby("city_state").survey_date.count()
city_mask = city_mask[city_mask == 23]
# apply city mask to shrink the df
def in_city_mask(x):
return x in city_mask
df = df[df.city_state.apply(in_city_mask)]
threshold_mask = df_model.groupby('city_state').five_or_more_units_bldgs_est.agg(lambda x: (x == 0).sum())
threshold_mask = threshold_mask[threshold_mask < 1].index.tolist()
# apply threshold mask to shrink the df
def in_threshold_mask(x):
return x in threshold_mask
df = df[df.city_state.apply(in_threshold_mask)]
df = df.sort_values(["major_city", "major_state", "survey_date"])
return df
df_model = filter_top_cities_building_permits(df_model)
df_model["avg_cost_per_building"] = df_model.five_or_more_units_value_est / df_model.five_or_more_units_bldgs_est
# Has the avg cost per building increased over time?
plt.figure(figsize=(15,5))
sns.lineplot(data=df_model, x="survey_date", y="avg_cost_per_building", ci=False)
plt.title("Avg cost per building over time")
df_model["avg_units"] = df_model.five_or_more_units_units_est / df_model.five_or_more_units_bldgs_est
# Has the avg number of units change over time?
plt.figure(figsize=(15,5))
sns.lineplot(data = df_model, x="survey_date", y="avg_units", ci=False)
plt.title("Avg number of units per building over time")
plt.figure(figsize=(15,5))
sns.lineplot(data = df_model, x="survey_date", y="five_or_more_units_units_est", ci=False)
plt.title("Number of units built over time")
# +
# What cities have the most dense buildings, on avg?
df_model.groupby("major_city").avg_units.mean().nlargest(10).plot.bar(figsize=(15,5))
plt.xticks(rotation=45)
# +
# What cities have the least dense buildings, on avg?
df_model.groupby("major_city").avg_units.mean().nsmallest(10).plot.bar(figsize=(15,5))
plt.xticks(rotation=45)
# -
# # Explore
#
# This exploration is for labeled data
df = permits_preprocessing_mother_function()
sns.barplot(data=df, x="should_enter", y="market_volume")
continuous_data = df.select_dtypes(include=["int", "float"])
continuous_data.info()
# +
f = plt.figure(figsize=(25,20))
continuous_data = df.select_dtypes(include=["int", "float"])
for count, element in enumerate(continuous_data):
f.add_subplot(4,5, count+1)
sns.barplot(data=df, x="should_enter", y=element, ci=False)
plt.tight_layout()
plt.show()
# -
df.info()
# # Model
features_for_modeling = ["city_state_high_density_value_delta_pct", "market_volume_delta_pct", "city_state_high_density_units_delta_pct", "ei"]
# features_for_modeling = ["ei", "city_state_high_density_bldgs_delta_pct", "avg_units_per_bldg", "value_per_bldg"]
label_feature = "should_enter"
train_scaled, validate_scaled, test_scaled, y_train, y_validate, y_test = permits_preprocessing_mother_function(modeling=True, features_for_modeling = features_for_modeling, label_feature= label_feature)
train_scaled.head()
# # Decision Tree
predictions = pd.DataFrame({"actual": y_train, "baseline": y_train.mode()[0]})
for i in range(1, 20):
clf, y_pred = model.run_clf(train_scaled, y_train, i)
score = clf.score(train_scaled, y_train)
validate_score = clf.score(validate_scaled, y_validate)
_, _, report = model.accuracy_report(clf, y_pred, y_train)
recall_score = report["True"].recall
print(f"Max_depth = {i}, accuracy_score = {score:.2f}. validate_score = {validate_score:.2f}, recall = {recall_score:.2f}")
clf, y_pred = model.run_clf(train_scaled, y_train, 4)
predictions["decision_tree"] = y_pred
accuracy_score, matrix, report = model.accuracy_report(clf, y_pred, y_train)
print(accuracy_score)
print(matrix)
report
coef = clf.feature_importances_
# We want to check that the coef array has the same number of items as there are features in our X_train dataframe.
assert(len(coef) == train_scaled.shape[1])
coef = clf.feature_importances_
columns = train_scaled.columns
df = pd.DataFrame({"feature": columns,
"feature_importance": coef,
})
df = df.sort_values(by="feature_importance", ascending=False)
sns.barplot(data=df, x="feature_importance", y="feature", palette="Blues_d")
plt.title("What are the most influencial features?")
# Interestingly, it seems that when it comes to decision tree, the `evolution_index` is actually the most indicative feature, along side the change in number of mortgage's approved. The total `quantity_of_mortgages_pop` doesn't seem to be as influencial in the predictions.
# # Random Forest
for i in range(1, 20):
rf, y_pred = model.run_rf(train_scaled, y_train, 1, i)
score = rf.score(train_scaled, y_train)
validate_score = rf.score(validate_scaled, y_validate)
_, _, report = model.accuracy_report(rf, y_pred, y_train)
recall_score = report["True"].recall
print(f"Max_depth = {i}, accuracy_score = {score:.2f}. validate_score = {validate_score:.2f}, recall = {recall_score:.2f}")
rf, y_pred = model.run_rf(train_scaled, y_train, 1, 3)
predictions["random_forest"] = y_pred
accuracy_score, matrix, report = model.accuracy_report(rf, y_pred, y_train)
print(accuracy_score)
print(matrix)
report
coef = rf.feature_importances_
columns = train_scaled.columns
df = pd.DataFrame({"feature": columns,
"feature_importance": coef,
})
df = df.sort_values(by="feature_importance", ascending=False)
sns.barplot(data=df, x="feature_importance", y="feature", palette="Blues_d")
plt.title("What are the most influencial features?")
# Interestingly, for the random_forest model, the delta of the number of loans approved by city where the most important or influencial indicator of whether a city would be `a hot martket` or not. The evolution index was the second most influencial feature. Again, the total `quantity_of_morgages_pop` was the least influencial feature.
# # KNN
for i in range(1, 20):
knn, y_pred = model.run_knn(train_scaled, y_train, i)
score = knn.score(train_scaled, y_train)
validate_score = knn.score(validate_scaled, y_validate)
_, _, report = model.accuracy_report(knn, y_pred, y_train)
recall_score = report["True"].recall
print(f"Max_depth = {i}, accuracy_score = {score:.2f}. validate_score = {validate_score:.2f}, recall = {recall_score:.2f}")
knn, y_pred = model.run_knn(train_scaled, y_train, 2)
predictions["knn"] = y_pred
accuracy_score, matrix, report = model.accuracy_report(knn, y_pred, y_train)
print(accuracy_score)
print(matrix)
report
# How do the different models compare on accuracy?
print("Accuracy Scores")
print("---------------")
for i in range(predictions.shape[1]):
report = model.create_report(predictions.actual, predictions.iloc[:,i])
print(f'{predictions.columns[i].title()} = {report.accuracy[0]:.2f}')
# How do the different models compare on recall?
print("Recall Scores")
print("---------------")
for i in range(predictions.shape[1]):
report = model.create_report(predictions.actual, predictions.iloc[:,i])
print(f'{predictions.columns[i].title()} = {report["True"].loc["recall"]:.2f}')
# How do the different models compare on recall?
print("Precision Scores")
print("---------------")
for i in range(predictions.shape[1]):
report = model.create_report(predictions.actual, predictions.iloc[:,i])
print(f'{predictions.columns[i].title()} = {report["True"].loc["precision"]:.2f}')
# ## Conclusion:
#
# Overall, we see that because we have optimized for *recall*, the accuracy scores are a bit lower than expected. However, our recall scores are really good. We will choose the KNN model as the most effective model, given that it consistently achieved the best scores (for accuracy, recall and precision).
#
# Just for reference:
#
# At 30% oversampling:
# * Best Model: KNN, K_n =1 (98% accuracy, 93% recall)
# * Accuracy diff between train and test = 12%
# * Recall diff between train and test = 0%
#
# At 50% oversampling:
# * Best Model: KNN, K_n = 1(100% accuracy, 100% recall)
# * Accuracy diff between train and test = 9%
# * Recall diff between train and test = 0%
# # Evaluate
knn, y_pred = model.run_knn(train_scaled, y_train, 1)
y_pred = knn.predict(test_scaled)
accuracy_score, matrix, report = model.accuracy_report(knn, y_pred, y_test)
print(accuracy_score)
print(matrix)
report
# ---
# +
# assert(1 == 2)
# -
# ----
#
# # Prediction
df = permits_preprocessing_mother_function()
df.head()
features_for_predicting = ["ei", "city_state_high_density_bldgs_delta_pct", "avg_units_per_bldg", "value_per_bldg"]
predictions = df[(df.year == 2020) | (df.year == 2019) | (df.year == 2018)].groupby("city_state")[features_for_predicting].mean()
predictions
# +
# Helper function used to updated the scaled arrays and transform them into usable dataframes
def return_values_prediction(scaler, df):
train_scaled = pd.DataFrame(scaler.transform(df), columns=df.columns.values).set_index([df.index.values])
return scaler, train_scaled
# Linear scaler
def min_max_scaler_prediction(df):
scaler = MinMaxScaler().fit(df)
scaler, df_scaled = return_values_prediction(scaler, df)
return scaler, df_scaled
# -
scaler, predictions_scaled = min_max_scaler_prediction(predictions)
predictions["label"] = knn.predict(predictions_scaled)
predictions
# +
city = predictions.reset_index().city_state.str.split("_", n=1, expand=True)[0]
state = predictions.reset_index().city_state.str.split("_", n=1, expand=True)[1]
# -
predictions = predictions.reset_index()
# +
predictions["city"] = city
predictions["state"] = state
# -
predictions
predictions[predictions.label == True].shape
predictions[predictions.label == True]
# +
# predictions.to_csv("predictions.csv")
# -
plt.figure(figsize=(15,5))
ax = sns.barplot(data=predictions, x="city", y="ei", hue="label")
plt.title("What markets will look like in 2021, based on evolution index")
plt.xticks(rotation=45, ha="right")
plt.xlabel("City")
plt.ylabel("Evolution Index (%)")
new_labels = ['Markets to not enter', 'Markets to enter']
h, l = ax.get_legend_handles_labels()
ax.legend(h, new_labels)
plt.show()
| additional_reference_notebooks/building_permits_modeling_notebook_daniel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Methods of Estimations
# In `Orbit`, we mainly support two methods to estimate model parameters (a.k.a posteriors in Bayesian).
#
# 1. Maximum a Posteriori (MAP)
# 2. Markov Chain Monte Carlo (MCMC)
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import orbit
from orbit.utils.dataset import load_iclaims
from orbit.models import ETS
from orbit.diagnostics.plot import plot_predicted_data
# -
print(orbit.__version__)
# load data
df = load_iclaims()
test_size = 52
train_df = df[:-test_size]
test_df = df[-test_size:]
response_col = 'claims'
date_col = 'week'
# ## Maximum a Posteriori (MAP)
#
# To use MAP method, one can simply specify `estimator='stan-map'` when instantiating a model. The advantage of MAP estimation is a faster computational speed. We also provide inference for MAP method, with the caveat that the uncertainty is mainly generated by bootstraping the noise process and as such we may not observe the uncertainty band from seasonality or other components.
# %%time
ets = ETS(
response_col=response_col,
date_col=date_col,
estimator='stan-map',
seasonality=52,
seed=8888,
)
ets.fit(df=train_df)
predicted_df = ets.predict(df=test_df)
_ = plot_predicted_data(train_df, predicted_df, date_col, response_col, title='Prediction with ETSMAP')
# To have the uncertainty from MAP, one can speicify `n_bootstrap_draws` (default to be -1).
# +
ets = ETS(
response_col=response_col,
date_col=date_col,
estimator='stan-map',
seasonality=52,
seed=8888,
n_bootstrap_draws=1e4
)
ets.fit(df=train_df)
predicted_df = ets.predict(df=test_df)
_ = plot_predicted_data(train_df, predicted_df, date_col, response_col, title='Prediction with ETSMAP')
# -
# ## MCMC
#
# To use MCMC method, one can specify `estimator='stan-mcmc'` (also the default) when instantiating a model. Compared to MAP, it usually takes longer time to fit a full Bayesian models where **No-U-Turn Sampler (NUTS)** [(<NAME> 2011)](https://arxiv.org/abs/1111.4246) is carried out under the hood. The advantage is that the inference and estimation are usually more robust.
# ### MCMC - Full
# %%time
ets = ETS(
response_col=response_col,
date_col=date_col,
estimator='stan-mcmc',
seasonality=52,
seed=8888,
num_warmup=400,
num_sample=400,
)
ets.fit(df=train_df)
predicted_df = ets.predict(df=test_df)
_ = plot_predicted_data(train_df, predicted_df, date_col, response_col, title='Prediction with ETSFull')
# You can also access the posterior samples by the attribute of `._posterior_samples` as a `dict`.
ets._posterior_samples.keys()
# ### MCMC - Point Estimation
# One can also have the point estimates via MCMC by specifying `point_method` as `mean` or `median` via `.fit`.
#
#
# Just like the full Bayesian method, it runs through the MCMC algorithm which is **NUTS** by default. The difference from a full model is that it aggregates the posterior samples first based on mean or median then does the prediction once using the aggreated posteriors.
# %%time
ets = ETS(
response_col=response_col,
date_col=date_col,
estimator='stan-mcmc',
seasonality=52,
seed=8888,
)
# specify point_method
ets.fit(df=train_df, point_method='mean')
predicted_df = ets.predict(df=test_df)
_ = plot_predicted_data(train_df, predicted_df, date_col, response_col,
title='Prediction with point method')
# Similarly, one can also specify `n_bootstrap_draws` to have the uncertainty for the point estimates.
# +
# %%time
ets = ETS(
response_col=response_col,
date_col=date_col,
estimator='stan-mcmc',
seasonality=52,
seed=8888,
n_bootstrap_draws=1e4,
)
# specify point_method
ets.fit(df=train_df, point_method='mean')
predicted_df = ets.predict(df=test_df)
_ = plot_predicted_data(train_df, predicted_df, date_col, response_col,
title='Prediction with point method')
| docs/tutorials/model_estimations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Summarising and Computing Descriptive Stats
# pandas objects are equipped with a set of common mathematical and statistical methods. Most of these fall into the category of reductions or summary statistics, methods that extract a single value from a Series or a Series of values from the rows and columns of a DataFrame
# - have built0in handling for missing data
import numpy as np
import pandas as pd
df = pd.DataFrame([[1.4, np.nan],[7.1, -4.5],
[np.nan, np.nan],[0.75,-1.3]],
index = ['a','b','c','d'],
columns=['one', 'two'])
df
df.sum()
df.sum(axis=1)
#NA values are excluded unless the entire slice is NA
#Skipping NA values can be disabled with the skipna option
df.mean(axis=1,skipna=False)
# #### The below method is extremely useful
df.describe()
# ## List of Descriptive and Summary Stat Methods
#
# - count: Number of non-NA values
# - describe: set of summary stats
# - min, max
# - argmin, argmax: compute index locations at which min and max value obtained, respectively
# - idxmin, idxmax: return index value where the max and min values are contained
# - quantile: compute sample quantile ranging from 0 to 1
# - sum
# - mean
# - median
# - mad: mean absolute devaition from the mean value
# - prod: product of all values
# - var: simple variance of all values
# - std
# - skew: sample skewness of values
# - cumsum: cumlative sum of values
# - diff: compute first arthmetic difference(useful for time series)
# - pct_change: Compute percent changes (valuable for time series analysis)
#
# ## Correlation & Covariance
# <b>Correlation:</b> a mutual relationship or connection between two or more things
#
# <b>Covariance:</b> In probability theory and statistics, covariance is a measure of the joint variability of two random variables. If the greater values of one variable mainly correspond with the greater values of the other variable, and the same holds for the lesser values, the covariance is positive
#
#
# Correlation and covaraince are computed for pairs of arguments. This example considers some DF’s of stock prices and volumes obtained from Yahoo! Finance using the add-on pandas-datareader package.
import pandas_datareader.data as web
# +
all_data = {ticker: web.get_data_yahoo(ticker)
for ticker in ['AAPL', 'IBM', 'MSFT', 'GOOG']}
price = pd.DataFrame({ticker: data['Adj Close']
for ticker, data in all_data.items()})
volume = pd.DataFrame({ticker: data['Volume']
for ticker, data in all_data.items()})
# -
#compute percent changes, a time series operation
returns = price.pct_change()
returns.tail()
# <b>.corr:</b> computes the correlation
#
# <b>.cov:</b> computes the covariance
returns['MSFT'].corr(returns['IBM'])
returns['MSFT'].cov(returns['IBM'])
#to return a full correaltion or covariance matrix
returns.corr()
returns.cov()
# <b>corrwith method</b>
#
# Using this, you can compute pairwise correlations. Can also pass axis=‘columns’ to get row-by-row results.
returns.corrwith(returns.IBM)
# ## Unique Values, Value Counts
#To return an array of the unique values in a Series, use .unique()
obj = pd.Series(['a','a','d','c','f','c','g','f'])
obj
uniques = obj.unique()
uniques
#To compute a Series containing value frequencies
obj.value_counts()
| Notes/Pandas/Getting Started with pandas/Summarising and Computing Descriptive Stats.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++17
// language: C++17
// name: xcpp17
// ---
// # Zeichen und Strings
// <div class="prereq">
// <h3>Das sollte man kennen ...</h3>
// <div>
// <a class="prereq" href="/user-redirect/algoviz/lessons/02_Grundlagen/01_Datentypen.ipynb">Datentypen</a>
// und <a class="prereq" href="/user-redirect/algoviz/lessons/02_Grundlagen/13_Zaehlschleifen.ipynb">Zählschleifen</a>.
// </div>
// </div>
// ## Zeichen
//
// In den bisherigen Lessons haben wir schon eine Reihe von **Strings** verwendet. Meistens in der Form von **konstanten Strings**, d.h. festen Texten die z.B. ausgegeben werden:
// +
#include <iostream>
using namespace::std; // Dadurch können uns das vorangestelte std:: sparen
cout << "Dies ist ein KONSTANTER STRING" << endl;
// -
// Ein String ist eine Sequenz von einzelnen **Zeichen**. Für diese steht der Datentyp `char` (Kurzform von character) zur Verfügung, den wir schon in der Lesson [Datentypen](/user-redirect/algoviz/lessons(02_Grundlagen/01_Datentypen.ipynb) kennengelernt haben. Im GEgensatz zu den Strings wird ein einzelnes Zeichen in **Einfache Anführungszeichen** gesetzt.
char zeichen = '*';
// Wie wir auch schon gesehen haben, entspricht jeder `char` einem 16-Bit ASCII Wert. Um zu sehen, welcher Wert das ist, können wir einfach den Typ **casten**, d.h. umwandeln. Dies geschieht auf die folgende Weise:
(int) zeichen
// Hierzu zwei Anmerkungen:
//
// 1) Dadurch das wir das Semikolon am Ende weglassen, wird in Jupyter Notebooks das Ergebnis direkt ausgegeben. Wir können uns also `cout << ...` sparen.
//
// 2) Das vorangestellte `(int)` besagt, dass der folgende Wert in einen `int`-Wert, also eine ganze Zahl umgewandelt werden soll. Daher ist das Ergebnis 42. Der Unicode des Zeichens `*`.
//
// Das mit dem Cast geht auch anders herum:
(char) 42
// ### Was man mit Zeichen machen kann und was nicht
//
// Man kann Werte vom Typ `char` miteinander vergleichen. `==` und `!=` prüfen auf Gleich- bzw. Ungleichheit. Die Relationen `<`, `>`, `<=` und `>=` vergleichen die ASCII-Werte der Zeichen miteinander.
'a' < 'b'
// Auch die Rechenoperation `+`, `-`, `*`, `/` und `%` verwenden einfach die ASCII-Werte. D.h. die Zeichen werden **implizit** in `int`-Werte umgewandelt. Daher sind die Ergebnisse auch vom Typ `int`.
'b' - 'a'
'a' + 'z'
// ## Strings
//
// Wie bereits gesagt, steht der Datentyp `std::string` (oder einfach `string`) für Sequenzen von vielen Zeichen. Er wird durch die Bibliothek `<string>` zur Verfügung gestellt. Häufig wird diese auch durch andere Bibliotheken (z.B. `<iostream>`) "miteingebunden", so dass man sich oft sparen kann es selbst zu tun.
// +
#include <string>
using namespace std;
// -
// Der Datentype `string` und alle damit verbundenen Operationen gehören zum namespace `std`, so dass man ihnen entweder `std::` voranstellen muss. Alternativ kann man sich das mit `using namespace std;` ersparen.
string text = "Dies ist eine Text";
// Konstante Strings werden in **doppelte Anführungszeichen** gesetzt. Verwendet man, wie bei `char` einfache, erhält man einen Fehler.
string text = 'Dies ist kein Text'; // Das wird etwas länger ...
// Umgekehrt übrigens auch:
char zeichen = "a"; // Die Fehlermeldung ist deutlich kürzer.
// Man kann Strings **konkatenieren**, d.h. aneinanderhängen:
string s = "Dies";
string t = "ist ein Text";
string r = s + " " + t;
// Man kann sie ausgeben:
cout << r << endl;
// Und man kann kann sie einlesen:
// +
string eingabe;
cout << "Geben Sie was ein : ";
cin >> eingabe;
cout << "Das haben Sie eingegeben : \"" << eingabe << "\"" << endl;
// -
// Der **Backslash** `\` vor den dopelten Anführungszeichen stellt sicher, das sie als Teil des Strings und nicht als Anfang oder Ende interpretiert werden. Man sagt, dass der Bakslash `\` das folgende Zeichen **schützt** es wird damit als Zeichen und nicht als syntaktisches Element interpretiert.
// Wenn nicht schon geschehen, probieren Sie mal in der Eingabe etwas mit Leerzeichen.
// Sie werden es nicht hinbekommen!
//
// Für solche Fälle wird der folgende Befehl benötigt:
// +
getline(cin,eingabe);
cout << "Das haben Sie eingegeben : \"" << eingabe << "\"" << endl;
// -
// ## Die Struktur von Strings
//
// Da ein String eine Sequenz von Zeichen ist, ist es nicht wirklich erstaunlich, dass man auf seine einzelnen Zeichen zugreifen kann. Sie sind, beginnend mit 0, durchnummeriert und können auch einzeln abgefragt werden. Dies geht entweder mit `at()` oder mit dem **Index-Operator** `[]`.
// +
string text = "Zeichenkette";
char zeichen = text.at(7); // Das Zeichen an Index 7 (also 'k')
cout << zeichen << endl;
zeichen = text[6]; // Das Zeichen in Index 6 (also 'n')
cout << zeichen << endl;
// -
// Verändern kann man die Zeichen ebenfalls einzeln:
// +
text[3] = 'X';
text.at(4) = 'Y';
cout << text << endl;
// -
// Die **Länge** eines Strings erhält man mittels `length()`.
cout << text.length() << endl;
// Damit können wir jetzt jeden String rückwärts ausgeben lassen.
//
// Dafür verwenden wir eine Zähl- oder For-Schleife und durchlaufen die indices von hinten nach vorne. Da die Nummerierung der Position bei 0 beginnt, hat das letzte Zeichen den Index `length()-1`. Also ist das die Intialisierung.
//
// Da wir rückwärts laufen, muss das Inkrement die Position immer um eins senken.
// +
string eingabe;
cout << "Geben Sie einen Text ein : ";
getline(cin,eingabe);
for ( int pos = eingabe.length()-1; pos > 0; pos = pos - 1 ) {
cout << eingabe[pos];
}
cout << endl;
// -
// <div class="task">
// <h3>Aufgabe</h3>
// <div>
// Das Programm enthält einen Fehler. Korrigieren Sie ihn!
// </div>
// </div>
// ### Was noch so geht
//
// Im Folgenden ein paar Beispiele, was man mit Strings so machen kann.
// #### Teilstrings finden
//
// Man kann Teilstrings finden.
// +
#include <iostream> // Falls Sie neustarten müssen
using namespace std;
string text = "Eine Nadel im Heuhaufen, und noch eine Nadel.";
int pos1 = text.find("Nadel");
int pos2 = text.find("Nadel", pos1+1);
int pos3 = text.find("Hammer");
cout << pos1 << endl << pos2 << endl << pos3 << endl;
// -
// Die Operation `int text.find(string teil, int start)` sucht das erste Vorkommen von `teil` in `text` und gibt die Anfangsposition zurück. Ist der zweite Parameter gegebn, beginnt die Suche bei diesem Index. Ist `teil` nicht enthalten, gibt `find` -1 zurück.
// #### Teilstrings kopieren
// +
#include <iostream> // Falls Sie neustarten müssen
using namespace std;
string text = "Eine Nadel im Heuhaufen.";
string teil = text.substr(5,5);
cout << teil << endl;
// -
// #### Teilstrings ersetzen
//
// Mit `text.replace` können Sie Teilstrings ersetzen.
// +
#include <iostream> // Falls Sie neustarten müssen
using namespace std;
string text = "Dies ist eine Mücke.";
cout << text << endl;
text.replace(12,8," Elefant"); // Der String wird geändert
cout << text << endl;
// -
// Die Argumente von `replace` sind der Index des ersten Zeichens das ersetzt werden soll, die Länge und der Ersatztext.
// <div class="prereq">
// <h3>Halt mal!</h3>
// <div>
// Aber das passt doch gar nicht! `"e Mücke"` ist nur sieben Zeichen lang!
// </div>
// </div>
// Und da sind wir wieder bei dem Problem mit den Umlauten. Schauen wir uns mal das `'ü'` genauer an und lassen es einfach mal so stehen:
'ü'
// Das `'ü'` ist zu groß für ein Zeichen. Es besteht in Wirklichkeit aus mehreren Zeichen.
string text = "ü";
text.length()
// Aber ...
text = "u";
text.length()
// Sehen wir uns die beiden Zeichen mal an:
string text="ü";
text[0]
text[1]
// Beide zusammen ergeben das `ü`. Für uns soll das hier genügen. Wir werden versuchen einfach keine Umlaute oder ähnliches zu nutzen.
//
// Wer mehr wissen will findet mehr Informationen untern dem Sticworten **Unicode** und **UTF-8** und in der Zusatz-Lesson [Unicode und andere Codierungen](/user-redirect/algoviz/lessons/02_Grundlagen/22_Unicode.ipynb) (sofern sie fertig ist).
// <div class="task">
// <h3>Aufgabe</h3>
// <div style="float:right"><a href="https://de.wikipedia.org/wiki/Terry_Pratchett"><img src="/user-redirect/algoviz/img/terrys-hat.png" width="40px"></a></div>
// <div>
// Machen Sie in dem folgenden Text aus allen Elefanten Mücken und aus jedem einzelnen Elefanten nur eine Mücke.
// <p>
// <b>Hinweise und Bemerkungen</b>
// <ul>
// <li>Die Backslashes schützen dei folgenden Newlines (unsichtbar). Damit gehören sie nicht zum String.</li>
// <li>Überlegen Sie sich gut in welcher Reihenfolge sie was ersetzen.</li>
// <li>Eine geeignete Schleife mit geeigneter Laufbedingung hilft weiter (eigentlich zwei).</li>
// </ul>
// </p>
// </div>
// </div>
// +
#include <iostream> // Falls Sie neustarten müssen
using namespace std;
string pratchett =
"Es heisst , die Welt sei flach und werde von vier Elefanten getragen, \
die auf dem Panzer einer riesigen Schildkroete stehen. Es heisst, \
die Elefanten haetten aufgrund ihrer Groesse Knochen aus Stein und \
Eisen, und Nerven aus Gold, weil diese ueber grosse Entfernungen \
hinweg besser leiten. Es heisst , dass der fuenfte Elefant vor langer \
Zeit heulend und trompetend durch die Luft der jungen Welt raste \
und hart genug landete, um Kontinente zu zerreissen und hohe \
Berge entstehen zu lassen. Niemand beobachtete die Landung, \
woraus sich eine interessante philosophische Frage ergibt: Wenn \
ein Millionen Tonnen schwerer zorniger Elefant vom Himmel faellt, \
ohne dass jemand da ist, der ihn hoert - verursacht er dann, \
philosophisch gesehen, irgendwelche Geraeusche? Und wenn ihn \
niemand sah - schlug er dann wirklich auf? Mit anderen Worten: \
War es nicht nur eine Geschichte fuer Kinder, um einige interessante \
natuerliche Ereignisse zu erklaeren? Was die Zwerge betrifft, von \
denen diese Legende stammt, und die tiefer graben als viele andere \
Leute: Sie meinen, die Geschichte enthalte ein Koernchen Wahrheit. \
\
An einem klaren Tag konnte man von einem geeigneten Ort in den \
Spitzhornbergen aus weit ueber die Ebene sehen. Im Hochsommer \
war es moeglich, die Staubwolken der Ochsengespanne zu zaehlen. \
Jedes Ochsenpaar bewegte sich mit einer Hoechstgeschwindigkeit \
von zwei Meilen in der Stunde und zog zwei Karren, jeweils mit vier \
Tonnen Fracht beladen. Die Fracht brauchte lange, um ihren \
Bestimmungsort zu erreichen, aber wenn sie dort ankam, gab es \
viel davon. Den Staedten am Runden Meer brachten die Karren \
Rohstoffe und manchmal auch Leute, die ihr Glueck versuchten und \
sich eine Hand voll Diamanten erhofften. Den Bergen brachten sie \
Fertigwaren, seltene Dinge von jenseits des Meeres und Leute, die \
Weisheit gefunden und ein paar Narben davongetragen hatten. Fuer \
gewoehnlich betrug der Abstand zwischen den Gespannen eine \
Tagesreise, wodurch sich die Landschaft in eine ausgebreitete \
Zeitmaschine verwandelte. An einem klaren Tag konnte man den \
letzten Dienstag sehen. Heliographen blitzten in der Ferne, als die \
Staubwolken Mitteilungen austauschten. Diese betrafen die \
Anwesenheit von Raeubern, Ladungen und Lokale, wo man doppeltes \
Spiegelei, eine dreifache Portion Bratkartoffeln und Steaks bekam, \
die auf allen Seiten ueber den Tellerrand ragten. Viele Leute waren mit \
den Karren unterwegs. Die Reise kostete nicht viel und war bequemer \
als ein Fussmarsch. Ausserdem erreichte man sein Ziel, frueher oder \
spaeter. Manche Leute reisten sogar gratis. Der Kutscher eines \
Karrens hatte Probleme mit seinen beiden Ochsen. Sie waren unruhig. \
In den Bergen haette er das erwartet, denn dort streiften Geschoepfe \
umher, die Ochsen fuer eine wandelnde Mahlzeit hielten. Aber hier \
gab es nichts Gefaehrlicheres als Kohl. Hinter ihm, in einer Luecke \
zwischen den Stapeln aus Bauholz schlief jemand. \
\
<NAME>, Der Fuenfte Elefant";
// +
int pos;
do {
pos = pratchett.find("Elefanten");
if ( pos >= 0 ) {
pratchett = pratchett.replace(pos,9,"Muecken");
}
} while (pos >= 0);
do {
pos = pratchett.find("Elefant");
if ( pos >= 0 ) {
pratchett = pratchett.replace(pos,7,"Muecke");
}
} while (pos >= 0);
cout << pratchett << endl;
// -
// Dann prüfen wir mal, ob Ihr Programm korrekt ist.
// +
int check = pratchett.find("Elefant");
if ( check == -1 ) {
cout << "Gut Gemacht! Ich habe keinen Elefanten gefunden!" << endl;
} else {
cout << "Da ist mindestens noch ein Elefant versteckt!" << endl;
}
// -
// Aber sind auch genügend Mücken drin?
// +
int zaehler = 0;
int muecke = -1;
do {
muecke = pratchett.find("Muecke",muecke+1);
if ( muecke >= 0 ) {
zaehler = zaehler+1;
}
} while ( muecke >= 0 );
if ( zaehler == 5 ) {
cout << "Genau richtig!" << endl;
} else if ( zaehler < 5 ) {
cout << "Zu wenig Mücken!" << endl;
} else if ( zaehler > 5 ) {
cout << "Eine Mückenplage! Zu viele!" << endl;
}
// -
// #### Einen Teilstring einfügen
//
// Man kann auch Teilstrings einfügen.
// +
#include <iostream> // Falls Sie neustarten müssen
using namespace std;
string text = "PenPineapplePen";
string complete = text.insert(12,"Apple");
cout << complete << endl;
// -
// <div style="float:right" title="Achten Sie darauf wohin Sie klicken!">✎<a href="https://www.youtube.com/watch?v=Ct6BUPvE2sM">🍍</a><a href="https://www.youtube.com/watch?v=m9DgT1lwulY">🍎</a>✎</div>
// ## Eine wichtige Beobachtung
//
// Ist Ihnen bei den diversen Befehlen etwas aufgefallen?
string s = "";
int pos = s.find("Nadel");
string t = s.substr(6,5);
string r = s.replace(13,4,"Elefant");
string u = s.insert(12,"Apple");
// In allen Fällen wir der String nicht als Parameter übergeben. Stattdessen werden die Operationen auf einer Variable vom Typ `string` aufgerufen. Die Variablke steht durch einen Punkt `.` getrennt davor. Das nennt man **Punktnotation**.
//
// Der Grund dafür ist, dass ein `string` ein [Objekt](/user-redirect/algoviz/lessons/02_Grundlagen/14_ErsterKontaktMitObjekten.ipynb) ist. Und damit funktionert er etwas anders als die sogenannten **elementaren** oder **atomaren** Typen `int`, `double`, `char` und `bool` (und ihre Varianten).
// <div class="followup">
// <h3>Weiter geht es mit ...</h3>
// <div>
// <a class="followup" href="/user-redirect/algoviz/lessons/02_Grundlagen/22_Caesar.ipynb">der Cäsar-Verschlüsselung</a> als Beispiel für einen Algorithmus auf Strings. Danach geht es zu den
// <a class="followup" href="/user-redirect/algoviz/lessons/03_Fortgeschritten/00_Arrays.ipynb">Arrays</a>.
// </div>
// </div>
| lessons/02_Grundlagen/16_Strings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 1 Polynomial curve fitting and regularization
# -------------------
# So far, linear model seems to be too simple to capture the variance in the data, which is actually generated from a flexible sinusoidal model as the latent function. Intuitively, you need more flexible model to capture data with large variantion. However, since you do not really know the latent model which generated the data, you do not want to make our model to be too flexible that leads to overfitting. Why? Because when the model is too flexible and tries to capture all variance in the data, the model would undesirably also capture the **variance in the noise**! It **overfits** the information in the data!
#
# In this section, you will see how model complexity influence model fitting. This will be illustrated through a **polynomial curve fitting**, where different orders of polynomial model will be trained to fit the above dummy dataset. You will notice how complex higher order polynomial model capture all the variance but overfit the data. To control the complexity of model and avoid overfitting, one option is to impose restrictions upon the model parameters **θ** while train the model to fit the data. A key idea in machine learning based regression, **regularization**, will thus be introduced. **Regularization** is an important technique for you to cope with model complexity. Various versions of **regularization**, such as the *lasso* and *ridge* regressions, are widely adopted.
# ### 1.0 Polynomial curve fitting
#
# Obviously, during training polynomial models to fit dummy dataset, the complexity of the model is determined by the orders of polynomial terms. For instance, polynomial model with order of 2 with polynomial terms (*x, x^2*) is less complex than that of order 3 with polynomial terms (*x, x^2, x^3*) and thus less flexible to be trained to capture the variance in the data.
#
# Explore the following code to see how model complexty affect the fitting results, and how complex model tries to fit everywhere in the dummy dataset.
# Recall how you generated the data.
# %matplotlib inline # In order to plot figures inline in Jupyter Notebook, we need to run this. But please ignore this in Colab.
# +
import numpy as np # You will be using Numpy module
import matplotlib.pyplot as plt # Module for visualization will also be used
# Define the true function for generating dummy dataset
def func(x):
# Generate sample dataset from a linearly transformed sinusoidal function: y=sin(x)+0.3*x+e (where 'e' is random noise)
return np.sin(x) + .3*x
# Define function for generating dummy dataset
def dummyData(func, sample_size, std):
# Artificially generate input x
x = np.random.rand(sample_size)[:,None]; x=x*15-5
# Call function to generate response y
y = func(x) + np.random.normal(scale=std, size=x.shape)
return x, y
# Generate dummy dataset
# 10 data points are created for model training, and another 5 are created to test the accuracy of the model
x_train, y_train = dummyData(func, 10, 0.25)
# Visualize the artificially generated dataset
plt.figure(figsize=(12,5))
plt.xlim([-6., 11])
plt.ylim([-4, 4])
plt.title('Data')
plt.plot(x_train,y_train,'kx',label='training data')
plt.legend()
# -
# Recall how you defined functions to train your model and make predictions.
# +
# Define a function to fit a linear model to the dummy data.
def fit(x, y):
# x : (N, D) np.ndarray
# y : (N,) np.ndarray
# Linear fit means a linear model in the form of M(x) = θ1 + θ2x is assumed
# Or, in the form of matrix multiplication M(X) = θ'X, where
# X is referred as designed vector/matrix in the form of [1, x]' and θ is [θ1, θ2]'
X_mat=np.hstack((np.ones(len(x)).reshape(-1,1), x)) # X_mat is a designed matrix
# Given the observed data y, a proper solution of θ to M(x) can be: θ* = argmin (||y - X @ θ||^2_2)
theta = np.linalg.inv(X_mat.T.dot(X_mat)).dot(X_mat.T).dot(y) # If you see an @ in the middle of a line, that's matrix multiplication.
var = np.mean(np.square(X_mat @ theta - y))
return theta, var
def predict(x, theta, var, return_std:bool=False):
X_mat=np.hstack((np.ones(len(x)).reshape(-1,1), x)) # X_mat is a designed matrix
y = X_mat @ theta
if return_std:
y_std = np.sqrt(var) + np.zeros_like(y)
return y, y_std
return y
# +
import functools
import itertools
# Define a fuction that project inputs into higher order polynomial inputs.
# For instance, when degree=3, x becomes to (x, x^2, x^3).
def transform(x, degree):
# Transforms input array with polynomial features
# x : (sample_size, n) ndarray
# Output : (sample_size, 1 + nC1 + ... + nCd) ndarray
if x.ndim == 1:
x = x[:, None]
x_t = x.transpose()
features = []
for degree in range(1, degree + 1):
for items in itertools.combinations_with_replacement(x_t, degree):
features.append(functools.reduce(lambda x, y: x * y, items))
return np.asarray(features).transpose()
x_all = np.linspace(-5, 10, 100).reshape(-1,1)
y_true = func(x_all)
for i, deg in enumerate([1, 2, 4, 8]):
plt.figure(figsize=(12,20))
plt.subplot(4, 1, i + 1)
x_poly = transform(x_train, deg) # Project original inputs into polynomial inputs
weights3, variance3 = fit(x_poly, y_train) # Fit data as a linear combination of the polynomial inputs
y_predict3 = predict(transform(x_all, deg), weights3, variance3) # Prediction
y_true = func(x_all) # True model values
plt.xlim([-6., 11])
plt.ylim([-4, 4])
plt.title('Data fitted: two models: '+'polynomial order = '+str(2**i))
plt.plot(x_train,y_train,'kx',label='training data') # Plot dummy dataset data points
plt.plot(x_all, y_true, linewidth=.5, label='true function curve') # True model values
plt.plot(x_all, y_predict3, '--m', linewidth=.5, label='predicted function curve') # Polynomial data fit
plt.legend()
# -
# ### 1.1 Control model complexity, regularization, and feature selection
#
# Given the order of the polynomial terms, the complexity of the model can be restricted or controlled by the parameters of the polynomial terms -- the **θ**. Indeed, there are multiple ways of **regularization** to control the parameters. You can either eliminate some *θ* of the higher order polynomial terms to retain only the lower order terms, or eliminate all **θ**.
#
# To achieve **regularization**, you need to define a new function that can control the impact of model weights **θ** to restrict the influence of terms in **X**. Technically, you can achieve this by adding a **regularization** term to our fitting function already defined in *section 0.1*.
#
# The **regularization term** restricts model weights **θ** within a certain range control the effects of regression terms, such as *(θ<sub>2</sub>, θ<sub>3</sub>,..., θ<sub>n+1</sub>)* of the polynomial terms *(x<sub>1</sub>, x<sub>2</sub>,..., x<sub>n</sub>)*. When the effects of these regression terms, especially the higher order ones, are restricted, the complexity of the model is controlled.
#
# You can of course design different forms of **regularization** to achieve different restriction effects. Popular **regularization** approaches have names such as **ridge regression** and **lasso regression**. The **regularization** term in **ridge regression** is introduced into the OLS loss function in the form of **||θ||^2_2**, thus the value of **θ** is also minimized while minimizing the loss function. It achieves an overall restriction for all **θ** equally, or for instance, for an 8 order polynomial model, *(θ<sub>2</sub>, θ<sub>3</sub>, θ<sub>4</sub>,..., θ<sub>9</sub>)* are all shrinking towards 0. **Lasso regression** imposes a **regularization** term in the form of **||θ||_1**, which eliminates **θ** in a way that some of them are eliminated more significant than the others. Ultimately, **lasso regression** reduces model complexity by deleting regressio terms, and acts as a **variable/feature selection** tool if you consider *(x<sub>1</sub>, x<sub>2</sub>, x<sub>3</sub>,..., x<sub>8</sub>)* as features in multivariable regression.
# +
# Here is how you train the linear model using the regularized Ordinary Least Squares (OLS)
# The regularization term restricts θ within ||θ||^2_2, which is known as the ridge regression.
def fitReg(x, y, alpha):
# x : (N, D) np.ndarray
# y : (N,) np.ndarray
# Similar to what has been defined in section 0.2, you need a designed matrix
X_mat=np.hstack((np.ones(len(x)).reshape(-1,1), x)) # X_mat is a designed matrix
# Now, a regularization term ||θ||^2_2 is introduced to control θ that restricts θ within certain range.
# The parameter alpha controls the extent of the restriction or regularization.
# Given the observed y, the solution of θ in section 0.2 becomes: θ* = argmin (||y - X @ θ||^2_2 + alpha * ||θ||^2_2)
eye = np.eye(np.size(X_mat, 1))
theta = np.linalg.solve(alpha * eye + X_mat.T @ X_mat, X_mat.T @ y)
#w = np.linalg.inv(X_mat.T.dot(X_mat)).dot(X_mat.T).dot(y) # If you see an @ in the middle of a line, that's matrix multiplication.
var = np.mean(np.square(X_mat @ theta - y))
return theta, var
deg = 8 # Polynomial model of order 8 already shown above
# Parameter that control the extent of restriction imposed upon model weights θ.
# Please try to experiment with this parameter to see how it affects the model fitting.
# As you can see when alpha is larger than 1e-1, the polynomial model with order 8 is already significantly restricted compared to that in the image above.
alpha = 1e-1
weights, variance = fitReg(transform(x_train, deg), y_train, alpha)
y_predict = predict(transform(x_all, deg), weights, variance)
plt.figure(figsize=(12,5))
plt.xlim([-6., 11])
plt.ylim([-4, 4])
plt.title('Data fitted')
plt.plot(x_train,y_train,'kx',label='training data')
plt.plot(x_all, y_true, linewidth=.5, label='true function curve')
plt.plot(x_all, y_predict, '--g', linewidth=.5, label='predicted function curve')
plt.legend()
# +
# By adding a regularization term to the loss function of OLS in the form of ||θ||_1, it becomes the lasso regression.
# And the loss function to solve θ in section 0.2 becomes: θ* = argmin (||y - X @ θ||^2_2 + alpha * ||θ||_1).
# In fact you don't need to write the function to implement every kind of regularized regression as for illustration shown above.
# There are already many great Python modules that implemented these widely used regression techniques.
# For instance, you can directly import the Lasso function provided by the "sklearn" module as shown below.
from sklearn.linear_model import Lasso
# By using Lasso function provided by "sklearn", you don't even have to generate the designed matrix from the input, the function takes care of the original input for you.
# But you still need to create the 8 order regression terms (x, x^2, x^3,..., x^8) by transforming the input to enforce a polynomial model fitting.
deg = 8 # Again, use the polynomial model of order 8 already shown above
# You can directly define your model by calling the Lasso function imported from "sklearn", and set the extent of regularization.
model_lasso = Lasso(alpha=1e-1) # Again, please try to experiment with this parameter to see how it affects the model fitting.
# Then, fit this model to the data.
model_lasso.fit(transform(x_train, deg), y_train)
# Make predictions every where to show the fitted curve.
y_predict = model_lasso.predict(transform(x_all, deg))
# Plot the data. Does the fitted curve look less complex than it was in section 1.1?
# Remember try to experiment with different alpha.
plt.figure(figsize=(12,5))
plt.xlim([-6., 11])
plt.ylim([-4, 4])
plt.title('Data fitted')
plt.plot(x_train,y_train,'kx',label='training data')
plt.plot(x_all, y_true, linewidth=.5, label='true function curve')
plt.plot(x_all, y_predict, '--g', linewidth=.5, label='predicted function curve')
plt.legend()
# -
| REG_1_polynomial_regularization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %config InlineBackend.figure_formats = ['svg']
# Helpers
from SimEx.Utilities.Units import meter, electronvolt, joule, radian
from SimEx.Analysis.DiffractionAnalysis import DiffractionAnalysis, plotResolutionRings, azimuthalIntegration
# Simple Beam Parameters
from SimEx.Parameters.PhotonBeamParameters import PhotonBeamParameters
# Diffraction
from SimEx.Parameters.DetectorGeometry import DetectorGeometry, DetectorPanel
# -
diffr_path = "../../../data/simulation/xstal/xstal_diffr200.h5"
analyzer = DiffractionAnalysis(input_path=diffr_path,
pattern_indices=1,
poissonize=True
)
print('npattern =',analyzer.npattern)
analyzer.plotPattern(logscale=True)
plotResolutionRings(analyzer.parameters)
# ls ../../../data/simulation/xstal/
| src/controller/crystFEL/plotDiffr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Library
# +
import numpy as np
import torch
import torch.nn as nn
from utils import *
from dataset import CollisionDataset
from torch.utils.data import DataLoader
# -
# # Model
class ResidualMLP(nn.Module):
def __init__(self, in_dim, out_dim):
super(ResidualMLP, self).__init__()
self.hidden_dim = 128
self.fc_1 = nn.Sequential(
nn.Linear(in_dim, self.hidden_dim),
nn.ReLU(inplace=True),
nn.Linear(self.hidden_dim, self.hidden_dim),
nn.ReLU(inplace=True),
)
self.fc_out = nn.Linear(self.hidden_dim, out_dim)
def forward(self, x, y_phy):
x = self.fc_1(x)
x = self.fc_out(x)
return x + y_phy
# +
def train_model(model, train_loader, test_loader, num_epochs, optimizer, scheduler, criterion):
model.train()
# Training the Model
min_test_dif = float('inf')
epoch_loss = []
for epoch in range(num_epochs):
batch_loss = []
for i, data in enumerate(train_loader):
# get the inputs
x = data['x']
y = data['y']
y_phy = data['y_phy']
x = x.cuda()
y = y.cuda()
y_phy = y_phy.cuda()
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
y_hat = net(x, y_phy)
loss = criterion(y_hat, y)
loss.backward()
optimizer.step()
batch_loss.append(loss.item())
# Results every epoch
cur_epoch_loss = sum(batch_loss) / len(batch_loss)
# Scheduler
scheduler.step(cur_epoch_loss)
# Test the network
train_dif = test_model(model, train_loader, criterion)
test_dif = test_model(model, test_loader, criterion)
# Print the result
print('Epoch: %d Train Loss: %f Train Dif: %f Test Dif: %f'
% (epoch, cur_epoch_loss, train_dif, test_dif))
epoch_loss.append(cur_epoch_loss)
if min_test_dif > test_dif:
min_test_dif = test_dif
print('Best')
return epoch_loss
def test_model(model, test_loader, criterion):
# Test the Model
model.eval()
batch_loss = []
for i, data in enumerate(test_loader):
# get the inputs
x = data['x']
y = data['y']
y_phy = data['y_phy']
x = x.cuda()
y = y.cuda()
y_phy = y_phy.cuda()
y_hat = net(x, y_phy)
loss = criterion(y_hat, y)
batch_loss.append(loss.item())
# Results every epoch
cur_epoch_loss = sum(batch_loss) / len(batch_loss)
model.train()
return cur_epoch_loss
# +
#################### Hyperparameters ####################
num_epochs = 50000
learning_rate = 0.001
weight_decay = 0
in_frames_num = 3
pre_frames_num = 15
factor = 0.95
patience = 40
batch_size = 16
#################### Hyperparameters ####################
net = ResidualMLP(in_dim=7, out_dim=2).cuda()
criterion = torch.nn.MSELoss()
train_set = CollisionDataset(
'./dataset/uIsPoint3/train',
sample_num=32
)
test_set = CollisionDataset(
'./dataset/uIsPoint3/test',
)
print(len(train_set), len(test_set))
train_loader = DataLoader(train_set, batch_size=batch_size, shuffle=True)
test_loader = DataLoader(test_set, batch_size=len(test_set), shuffle=False)
optimizer = torch.optim.Adam(net.parameters(), lr=learning_rate, weight_decay=weight_decay)
scheduler = torch.optim.lr_scheduler.ReduceLROnPlateau(
optimizer,
mode='min',
factor=factor,
patience=patience,
verbose=True,
threshold=1e-3
)
train_loss = train_model(
net,
train_loader,
test_loader,
num_epochs,
optimizer,
scheduler,
criterion
)
# -
| collision/Residual_physics_collision.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
# ***
#
# <br><h2>Session 10 | Combining PCA and Clustering</h2>
# <br><h3>THE ULTIMATE REVIEW FOR THE FINAL</h3>
# <h4>DAT-5303 | Machine Learning</h4>
# <NAME> - Faculty of Analytics<br>
# Hult International Business School<br><br><br>
#
# ***
# ***
# ***
# + deletable=false editable=false run_control={"frozen": true}
# <h3>Part I: Conceptual Review</h3><br>
# Let's start by reviewing our key unsupervised learning concepts.<br><br>
#
# <strong>Principal Component Analysis</strong><br>
# Focuses on the variance between explanatory variables (i.e. their covariance).<br><br>
#
# Three situations where PCA is useful:
# 1. Correlated Explanatory Variables (what's going on behind the scenes of the correlation)
# 2. Dimensionality Reduction (grouping large variable sets into a more manageable number of factors)
# 3. Latent Trait Exploration (measuring what cannot be measured directly)
#
#
# <br><br>
# <strong>Clustering</strong><br>
# Divides observations into groups (i.e. clusters). Observations can be grouped based on their similarities or their differences.
#
# <br>
# <h3><u>Don't forget!!!</u></h3>
#
# 1. Don't mix data concepts in the same algorithm (spending behavior, demographics, psychometrics, etc.).
# 2. Scale your data.
# 3. Interpretation is subjective, so spend ample time on this step.
#
# <br><br>
# <strong>Challenge 1</strong><br>
# Complete the code to import the necessary packages for this analysis.
# +
########################################
# importing packages
########################################
import pandas as pd # data science essentials
import matplotlib.pyplot as plt # fundamental data visualization
import seaborn as sns # enhanced visualizations
from sklearn.preprocessing import StandardScaler# standard scaler
from sklearn.decomposition import PCA # pca
from scipy.cluster.hierarchy import dendrogram, linkage # dendrograms
from sklearn.cluster import KMeans # k-means clustering
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
#
# <br>
# Run the following code to load the dataset and set print options.
# + deletable=false editable=false
########################################
# loading data and setting display options
########################################
# loading data
customers_df = pd.read_excel('top_customers_subset.xlsx')
# setting print options
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
pd.set_option('display.max_colwidth', 100)
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
#
# <br>
# <strong>User-Defined Functions</strong><br>
# Run the following code to load the user-defined functions used throughout this Notebook.
# + deletable=false editable=false
########################################
# inertia
########################################
def interia_plot(data, max_clust = 50):
"""
PARAMETERS
----------
data : DataFrame, data from which to build clusters. Dataset should be scaled
max_clust : int, maximum of range for how many clusters to check interia, default 50
"""
ks = range(1, max_clust)
inertias = []
for k in ks:
# INSTANTIATING a kmeans object
model = KMeans(n_clusters = k)
# FITTING to the data
model.fit(data)
# append each inertia to the list of inertias
inertias.append(model.inertia_)
# plotting ks vs inertias
fig, ax = plt.subplots(figsize = (12, 8))
plt.plot(ks, inertias, '-o')
# labeling and displaying the plot
plt.xlabel('number of clusters, k')
plt.ylabel('inertia')
plt.xticks(ks)
plt.show()
########################################
# scree_plot
########################################
def scree_plot(pca_object, export = False):
# building a scree plot
# setting plot size
fig, ax = plt.subplots(figsize=(10, 8))
features = range(pca_object.n_components_)
# developing a scree plot
plt.plot(features,
pca_object.explained_variance_ratio_,
linewidth = 2,
marker = 'o',
markersize = 10,
markeredgecolor = 'black',
markerfacecolor = 'grey')
# setting more plot options
plt.title('Scree Plot')
plt.xlabel('PCA feature')
plt.ylabel('Explained Variance')
plt.xticks(features)
if export == True:
# exporting the plot
plt.savefig('top_customers_correlation_scree_plot.png')
# displaying the plot
plt.show()
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
#
# <br>
# <strong>Challenge 2</strong><br>
# Drop demographic information and scale the data.
# +
# dropping demographic information
purchase_behavior = customers_df.drop(['Channel', 'Region'],
axis = 1)
# INSTANTIATING a StandardScaler() object
scaler = StandardScaler()
# FITTING the scaler with the data
scaler.fit(purchase_behavior)
# TRANSFORMING our data after fit
X_scaled = scaler.transform(purchase_behavior)
# converting scaled data into a DataFrame
purchases_scaled = pd.DataFrame(X_scaled)
# reattaching column names
purchases_scaled.columns = purchase_behavior.columns
# checking pre- and post-scaling variance
print(pd.np.var(purchase_behavior), '\n\n')
print(pd.np.var(purchases_scaled))
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
#
# <br>
#
# <h3>Part II: Principal Component Analysis</h3>
#
# Our process here is to:
# 1. Develop a PCA model with no limit to principal components
# 2. Analyze the <strong>explained_variance_ratio</strong> and the <strong>scree plot</strong>
# 3. Decide how many components to RETAIN
# 4. Build a new model with a limited number of principal components
# 5. Interpret your results (what does each PC represent)
#
# <br>
# Remember, there may be some niche opportunities in smaller principal components. Be sure to check this before moving on because this may lead to excellent market opportunities.
#
# <br><br>
# <strong>Challenge 3</strong><br>
# Develop a PCA object with no limit to principal components and analyze its scree plot.
# -
# INSTANTIATING a PCA object with no limit to principal components
pca = PCA(purchases_scaled,
random_state = 802)
help(PCA)
# +
# INSTANTIATING a PCA object with no limit to principal components
pca = PCA(n_components=None,
random_state = 802)
# FITTING and TRANSFORMING the scaled data
customer_pca = pca.fit_transform(purchases_scaled)
# calling the scree_plot function
scree_plot(pca_object = pca)
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
#
# <br>
# <strong>Challenge 4</strong><br>
# Reduce the number of principal components to a reasonable number based on the scree plot. Note that we do not need to rerun the scree plot. In this example, we will assume three PCs is a reasonable number based on the elbow in the scree plot. Also note that it would have been reasonable to retain enough PCs so that the cumulative explained variance ratio is greater than or equal to 0.80.
# +
# INSTANTIATING a new model using the first three principal components
pca_3 = PCA(n_components = 3,
random_state = 802)
# FITTING and TRANSFORMING the purchases_scaled
customer_pca_3 = pca_3.fit_transform(purchases_scaled)
# calling the scree_plot function
scree_plot(pca_object = pca_3)
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
#
# <br>
# <strong>OPTIONAL STEP</strong><br>Run the following code to compare the variance of the unlimited PCA model with the variance of the reduced PCA model. We are doing this in this script simply to show that the explain variance in each principal component does not change after dropping smaller PCs.
# + deletable=false editable=false
####################
### Max PC Model ###
####################
# transposing pca components (pc = MAX)
factor_loadings = pd.DataFrame(pd.np.transpose(pca.components_))
# naming rows as original features
factor_loadings = factor_loadings.set_index(purchases_scaled.columns)
##################
### 3 PC Model ###
##################
# transposing pca components (pc = 3)
factor_loadings_3 = pd.DataFrame(pd.np.transpose(pca_3.components_))
# naming rows as original features
factor_loadings_3 = factor_loadings_3.set_index(purchases_scaled.columns)
# checking the results
print(f"""
MAX Components Factor Loadings
------------------------------
{factor_loadings.round(2)}
3 Components Factor Loadings
------------------------------
{factor_loadings_3.round(2)}
""")
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
#
# <br>
# <strong>Challenge 5</strong><br>
# Name your principal components based on the latent traits they reflect.<br>
#
# In this step, make sure to develop a story behind what each PC name represents. This is an ideal method for bridging the gap between the technical and non-technical people you are working with. Remember, by doing a good job here you are putting analytics at the forefront of strategic decision making, which is a great way to boost your value within an organization.
# +
# naming each principal component
factor_loadings_3.columns = ['Herbivores',
'Fancy Diners',
'Winers']
# checking the result
factor_loadings_3
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
#
# <br>
# <strong>Challenge 6</strong><br>
# Analyze the factor loadings for each customer in the dataset. Do this by identifying groups of customers that have very high or very low factor loadings in any given principal component. A good heuristic is to look for factor loadings that are greater than one standard deviation from the mean in absolute value. Develop a strategy for key groups that you identify.<br><br>
#
# <strong>Don't forget</strong> to look at both the positive and negative loadings.<br>
# <strong>Don't forget</strong> to calculate the percentage of your audience effected by each loading when developing your targeting strategy/new ideas.<br>
# <strong>Don't forget</strong> to also consider the proportion of revenue generated by each group.
# +
# analyzing factor strengths per customer
X_pca_reduced = pca_3.transform(purchases_scaled)
# converting to a DataFrame
X_pca_df = pd.DataFrame(X_pca_reduced)
# checking the results
X_pca_df
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
#
# <br><h3>Part III: Clustering</h3><br>
# We are going to start by building an agglomerative clustering model. Remember, we are primarily interested in the <strong>dendrogram</strong> and the <strong>inertia plot</strong>. Our goal is to develop an idea as to how many clusters would be appropriate given our analysis of these tools, and then to apply this number of clusters to a k-Means model. Try to come away with 4-5 different numbers of clusters so that you have more options when applying k-Means. <strong>Before getting started, we need to rescale our data.</strong> The reason is that the variance amongst our features is no longer equal.
# + deletable=false editable=false
pd.np.var(X_pca_df)
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
#
# <br>
# <strong>Challenge 7</strong><br>
# Complete the code to prepare a scaled version of the factor loadings (i.e. principal components) dataset.
# +
# INSTANTIATING a StandardScaler() object
scaler = StandardScaler()
# FITTING the scaler with the data
scaler.fit(X_pca_df)
# TRANSFORMING our data after fit
X_scaled = scaler.transform(X_pca_df)
# converting scaled data into a DataFrame
pca_scaled = pd.DataFrame(X_scaled)
# reattaching column names
pca_scaled.columns = ['Herbivores',
'<NAME>',
'Winers']
# checking pre- and post-scaling variance
#print(pd.np.var(X_pca_df), '\n\n')
#print(pd.np.var(pca_scaled))
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
#
# <br>
# Run the following code to develop a dendrogram. Our goal here is to understand how many clusters to build using k-Means.
# + deletable=false editable=false
# grouping data based on Ward distance
standard_mergings_ward = linkage(y = pca_scaled,
method = 'ward')
# setting plot size
fig, ax = plt.subplots(figsize=(12, 12))
# developing a dendrogram
dendrogram(Z = standard_mergings_ward,
leaf_rotation = 90,
leaf_font_size = 6)
# saving and displaying the plot
plt.savefig('standard_hierarchical_clust_ward.png')
plt.show()
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
#
# <br>
# <strong>Challenge 8</strong><br>
# Develop a code to analyze the inertia plot. Our goal here is to develop more candidates for the number of clusters we might want to develop.
# -
# calling the inertia_plot() function
interia_plot(pca_scaled)
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
#
# <br>
# <strong>Challenge 9</strong><br>
# This is where we test our candidate number of clusters. When we find a clustering that we like, we move forward. For this example, let's assume we converged on a solution of three clusters.<br><br>
# <strong>Don't forget</strong> that the appropriate number of clusters does not have to be the same as the number of principal components that were retained.
# +
# INSTANTIATING a k-Means object with five clusters
customers_k_pca = KMeans(n_clusters = 3,
random_state = 802)
# fitting the object to the data
customers_k_pca.fit(pca_scaled)
# converting the clusters to a DataFrame
customers_kmeans_pca = pd.DataFrame({'Cluster': customers_k_pca.labels_})
# checking the results
print(customers_kmeans_pca.iloc[: , 0].value_counts())
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
#
# <br>
# <strong>Challenge 10</strong><br>
# Finish the code to display the centroids (mean values) for each cluster. Interpret their meaning. This is also a place where you may want to (optionally) name your clusters and develop back stories for ideal members of each group.
# +
# storing cluster centers
centroids_pca = customers_k_pca.cluster_centers_
# converting cluster centers into a DataFrame
centroids_pca_df = pd.DataFrame(centroids_pca)
# renaming principal components
centroids_pca_df.columns = ['Herbivores',
'Fancy Diners',
'Winers']
# checking results (clusters = rows, pc = columns)
centroids_pca_df.round(2)
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
#
# <br>
# <strong>Challenge 11</strong><br>
# Complete the code to concatenate channel, region, and PCA components into one DataFrame.
# +
# concatinating cluster memberships with principal components
clst_pca_df = pd.concat([customers_kmeans_pca,X_pca_df],
axis = 1)
# checking results
clst_pca_df
# concatenating demographic information with pca-clusters
final_pca_clust_df = pd.concat([customers_df.loc[ : , ['Channel', 'Region']],
clst_pca_df],
axis = 1)
# renaming columns
final_pca_clust_df.columns = ['Channel',
'Region',
'Cluster',
'Herbivores',
'Fancy Diners',
'Winers']
# checking the results
print(final_pca_clust_df.head(n = 5))
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
#
# <br>
# Run the following code to add labels to categorical variables. If you (optionally) named your clusters, make sure to label these as well.
# + deletable=false editable=false
# renaming channels
channel_names = {1 : 'Online',
2 : 'Mobile'}
final_pca_clust_df['Channel'].replace(channel_names, inplace = True)
# renaming regions
region_names = {1 : 'Alameda',
2 : 'San Francisco',
3 : 'Contra Costa'}
final_pca_clust_df['Region'].replace(region_names, inplace = True)
# renaming regions
cluster_names = {0 : 'Cluster 1',
1 : 'Cluster 2',
2 : 'Cluster 3'}
final_pca_clust_df['Cluster'].replace(cluster_names, inplace = True)
# adding a productivity step
data_df = final_pca_clust_df
# checking results
data_df
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
#
# <br>
# <h3>Part IV: Analyze with Demographics</h3><br>
# Now that we've completed all of our preparation through machine learning, we can analyze our results with demographics and other data.<br><br>
# <strong>Pause before this step</strong> so that you can consider all of the hypotheses and assumptions you have made up to this point. Also consider all of the assumptions your organization is making. For example, if the company is convinced of a particular trend, the following is a good opportunity to validate/negate that information.
# + deletable=false editable=false
########################
# Channel
########################
# Herbivores
fig, ax = plt.subplots(figsize = (12, 8))
sns.boxplot(x = 'Channel',
y = 'Herbivores',
hue = 'Cluster',
data = data_df)
plt.ylim(-10, 19)
plt.tight_layout()
plt.show()
# Fancy Diners
fig, ax = plt.subplots(figsize = (12, 8))
sns.boxplot(x = 'Channel',
y = 'Fancy Diners',
hue = 'Cluster',
data = data_df)
#plt.ylim(-3, 6)
plt.tight_layout()
plt.show()
# Winers
fig, ax = plt.subplots(figsize = (12, 8))
sns.boxplot(x = 'Channel',
y = 'Winers',
hue = 'Cluster',
data = data_df)
#plt.ylim(-5, 3)
plt.tight_layout()
plt.show()
########################
# Region
########################
# Herbivores
fig, ax = plt.subplots(figsize = (12, 8))
sns.boxplot(x = 'Region',
y = 'Herbivores',
hue = 'Cluster',
data = data_df)
#plt.ylim(-1, 8)
plt.tight_layout()
plt.show()
# Fancy Diners
fig, ax = plt.subplots(figsize = (12, 8))
sns.boxplot(x = 'Region',
y = 'Fancy Diners',
hue = 'Cluster',
data = data_df)
#plt.ylim(-3, 5)
plt.tight_layout()
plt.show()
# Winers
fig, ax = plt.subplots(figsize = (12, 8))
sns.boxplot(x = 'Region',
y = 'Winers',
hue = 'Cluster',
data = data_df)
#plt.ylim(-4, 3)
plt.tight_layout()
plt.show()
# + deletable=false editable=false run_control={"frozen": true}
# ***
# ***
#
# <br>
#
# ~~~
# __ __ _ _ _ _
# \ \ / / | | (_) | | |
# \ \_/ /__ _ _ _ __ ___ __ _ __| | ___ _| |_| |
# \ / _ \| | | | | '_ ` _ \ / _` |/ _` |/ _ \ | | __| |
# | | (_) | |_| | | | | | | | (_| | (_| | __/ | | |_|_|
# |_|\___/ \__,_| |_| |_| |_|\__,_|\__,_|\___| |_|\__(_)
#
#
# ~~~
# + deletable=false editable=false run_control={"frozen": true}
#
| S10 _ Combining PCA and Clustering - Guided.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import stanfordcorenlp
import spacy
nlp=spacy.load('en_core_web_sm')
s=u"Mr Robinson and Mr Lowth relied upon the views formed by Mr Bradley and Mr Ong about whether the Agreement applied to work carried out on the X50 work site."
d=nlp(s)
for i in d:
print i.text,i.tag_
from nltk.corpus import stopwords
st=stopwords("english")
print st,len(st)
nlt=stanfordcorenlp.StanfordCoreNLP('/home/judson/Documents/stanford-corenlp-full-2018-02-27/')
tr=nlt.dependency_parse(s)
(tr)
doc=nlp(s)
a=[]
for token in doc:
a.append([token.text,token.dep_, token.head.text, token.head.pos_,[child for child in token.children]])
#a={i[0]:i[1:] for i in a}
a
from nltk.parse.stanford import StanfordDependencyParser
dep_parser=StanfordDependencyParser(model_path="/home/judson/Documents/stanford-english-corenlp-2018-02-27-models/edu/stanford/nlp/models/lexparser/englishPCFG.ser.gz")
r=dep_parser.raw_parse(s)
from nltk.parse.corenlp import CoreNLPDependencyParser
d=CoreNLPDependencyParser()
# +
#for i in list(d.parse(s)):
# print i
# -
for i in doc:
if i.subtree:
print i,list(i.subtree),i.dep_,i.pos_,i.idx
else:
print i,i.dep_,i.pos_,i.idx
type(doc)
for token in doc:
print(token.text, token.dep_, token.head.text, token.head.pos_,[child for child in token.children])
d={tok.text:[tok.dep_,tok.head.text,tok.tag_,[child for child in tok.children]] for tok in doc}
print d
md= {i:j[3] for i,j in d.items()}
md
import spacy
nlp=spacy.load('en_core_web_sm')
def dict_conv(st):
doc=nlp(remove_stop_punct(st))
d={tok.text:[tok.dep_,tok.head.text,tok.tag_,[child for child in tok.children]] for tok in doc}
return d
data = {1 : [],2 : [],3 : [],4 : [3,2],5:[],6:[5],7:[],8:[4,6,9,1],9:[7]}
def dfs(data, path, paths = []):
datum = str(path[-1])
if datum in data:
if data[datum]==[]:
paths+=[path]
return paths
else:
for val in data[datum]:
new_path = path + [val]
paths = dfs(data, new_path, paths)
else:
paths += [path]
return paths
def parse_depend(dt):
md= {i:j[3] for i,j in dt.items()}
m=""
for i,j in dt.items():
if j[0]=="ROOT":
m=str(i)
ml=dfs(md,[m],[])
mpos=[[dt[str(j)][2] for j in i]for i in ml]
return mpos
print parse_depend(d)
def remove_stop_punct(s):
from nltk.corpus import stopwords
stopwords=stopwords.words('english')
doc=nlp(s)
#k=sorted([".","DT","TO","CC","IN"])
st=" ".join(t.text for t in doc if (t.pos_ not in ["PUNCT"] )and(t.text.lower() not in stopwords))
return st
#print remove_stop_punct(s)
print (dict_conv(s))
print parse_depend(dict_conv(s))
def deppars_matcher(s1,s2):
m1=parse_depend(dict_conv(unicode(s1, "utf-8")))
m2=parse_depend(dict_conv(unicode(s2, "utf-8")))
m3=[]
for i in m1:
if i in m2:
m3.append(i)
return m3
s2="I should interpolate to record that subsequent to reserving judgment on the question of leave to appeal, the Sharman applicants applied for a stay on Moore J's order requiring <NAME> to file that affidavit, and that I refused to grant that stay, but made orders requiring the affidavit to be left in a sealed envelope to be opened only upon further order of the Court."
s1="Those obligations were said to be no different in principle to any other interlocutory procedural order of the court, whether made pre-trial or during a trial, requiring parties to swear affidavits, to answer questions in cross-examination and to provide documentation."
deppars_matcher(s1,s2)
print parse_depend(dict_conv(unicode(s1, "utf-8")))
print parse_depend(dict_conv(unicode(s2, "utf-8")))
print dict_conv(unicode(s1,"utf-8"))
from nltk.corpus import stopwords
stopwords=stopwords.words('english')
sop=" ".join([i for i in s1.split() if i.lower() not in stopwords])
#print sop
dict_conv(unicode(sop,"utf-8"))
dd=nlp(u"this is a doctor,tent-tative from US in (1990).")
for df in dd:
print df.text,df.pos_
remove_stop_punct(u"this is a doctor,tent-tative from US in (1990).")
| Rough_implementation_ipynb/DepenPattern.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import pickle
import matplotlib.pyplot as plt
import numpy as np
# # Load Dataset
cifar_folder = os.path.join("/volumes/Data/cifar-10","cifar-10-batches-py")
allFiles = next(os.walk(cifar_folder))[2]
print("allFiles:", allFiles)
# ## load the pickle data
# +
# In[]
def unpickle(file):
with open(file, 'rb') as fo:
dict = pickle.load(fo, encoding='bytes')
return dict
label_data_path = os.path.join(cifar_folder, "batches.meta")
labels = unpickle(label_data_path)
print("labels:", labels)
train_dataset_1_path = os.path.join(cifar_folder, "data_batch_1")
train_dataset_1 = unpickle(train_dataset_1_path)
print("train dataset:", train_dataset_1.keys())
# -
# ## Show the data
# Notice the image stored in the pickle is the format `(R, R, R, ..., G, G, G, ..., B, B, B ...)` instead of `(R, G, B, R, G, B, ...)`.
# +
select_img_idx = 1
label = labels[b'label_names'][train_dataset_1[b'labels'][select_img_idx]]
print("Index: {}, Label: {}, Filename: {}".format(\
select_img_idx,
label, \
train_dataset_1[b'filenames'][select_img_idx]))
rawImgData = train_dataset_1[b'data'][select_img_idx]
rawImgData_r = rawImgData[0:1024].reshape(32,32,-1) / 255.0
rawImgData_g = rawImgData[1024:2048].reshape(32,32,-1) / 255.0
rawImgData_b = rawImgData[2048:].reshape(32,32,-1) / 255.0
imgData = np.dstack((rawImgData_r, rawImgData_g, rawImgData_b))
fig, ax = plt.subplots(1,1)
ax.set_title(label)
ax.imshow(imgData)
plt.show()
# -
| deep_learning/space_image/Cifar-10_Dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="7W_lSrg5H6dq" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 988} outputId="b0f6bda2-e2ab-4695-ea09-67706d50a28f" executionInfo={"status": "error", "timestamp": 1526405269832, "user_tz": -120, "elapsed": 8655, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "100979246453766324139"}}
# !pip install gputil
# !pip install psutil
# !pip install humanize
import psutil
import humanize
import os
import GPUtil as GPU
GPUs = GPU.getGPUs()
# XXX: only one GPU on Colab and isn’t guaranteed
gpu = GPUs[0]
def printm():
process = psutil.Process(os.getpid())
print("Gen RAM Free: " + humanize.naturalsize( psutil.virtual_memory().available ), " I Proc size: " + humanize.naturalsize( process.memory_info().rss))
print("GPU RAM Free: {0:.0f}MB | Used: {1:.0f}MB | Util {2:3.0f}% | Total {3:.0f}MB".format(gpu.memoryFree, gpu.memoryUsed, gpu.memoryUtil*100, gpu.memoryTotal))
printm()
# + id="rCj7B-azv_0D" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 499} outputId="bf93a2bf-3cca-4bd7-9b7d-150f7873ef2d" executionInfo={"status": "ok", "timestamp": 1526405341731, "user_tz": -120, "elapsed": 48548, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "100979246453766324139"}}
# !pip install imageio
# !pip install torch torchvision
# + id="jpbEz3JktpaP" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 35} outputId="5a01b5d7-5441-4f46-f772-649daffd8ece" executionInfo={"status": "ok", "timestamp": 1526405356802, "user_tz": -120, "elapsed": 15003, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "100979246453766324139"}}
import numpy as np
from sklearn.preprocessing import OneHotEncoder
from sklearn.utils import shuffle
from sklearn.model_selection import StratifiedShuffleSplit
import matplotlib.pyplot as plt
import pandas as pd
import urllib.request
import os, tarfile
import imageio
from scipy.io import loadmat
# from tensorflow.examples.tutorials.mnist import input_data
# %matplotlib inline
import tensorflow as tf
print(tf.test.gpu_device_name())
import torch
import torch.nn as nn
from torch.autograd import Variable
import torchvision.datasets as datasets
import torchvision.transforms as transforms
# + id="tEDytSfzts_2" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
SVHN_URL_TRAIN = 'https://www.dropbox.com/s/k02n8imqlqx3wk1/train_32x32.mat?dl=1'
SVHN_URL_TEST = 'https://www.dropbox.com/s/et2dulb99ld6fez/test_32x32.mat?dl=1'
# + [markdown] id="L0yQDzlctpcl" colab_type="text"
# # Utility functions
# + [markdown] id="8hVxGP31tpcr" colab_type="text"
# #### ToDos
# - Create a function to fetch data from a url.
# - Check if it is already downloaded.
# - Check if the file is csv or tar gz etc.
# - Add cross-validation code to be able to use sklearn cross_val_score function to quickly evaluate the performance.
# + id="9GsUFbnYLXXr" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
def fetch_data(URL, DOWNLOAD_FOLDER, DOWNLOAD_FILE):
if not os.path.isdir(DOWNLOAD_FOLDER):
os.makedirs(DOWNLOAD_FOLDER)
if not os.path.isfile(DOWNLOAD_FOLDER+DOWNLOAD_FILE):
print('Beginning file download...')
urllib.request.urlretrieve(URL, DOWNLOAD_FOLDER+DOWNLOAD_FILE)
print('Done.')
svhn_data = loadmat(DOWNLOAD_FOLDER+DOWNLOAD_FILE)
return svhn_data
# + id="LDR5Fhkttpcv" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
def split_train_test(XY, n_splits=1, test_size=0.2, random_state=42):
split = StratifiedShuffleSplit(n_splits=n_splits, test_size=test_size, random_state=random_state)
for train_index, test_index in split.split(XY[0], XY[1]):
X_train, Y_train = XY[0][train_index,:], XY[1][train_index]
X_test, Y_test = XY[0][test_index,:], XY[1][test_index]
return X_train, Y_train, X_test, Y_test
# + id="httI2oVntpdD" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
def get_svhn_data(url, download_folder, download_file, split_data=False):
svhn_dict = fetch_data(url, download_folder, download_file)
X = svhn_dict['X']
Y = svhn_dict['y']
Y_new = np.zeros(Y.shape, dtype=np.float32)
labels = np.arange(0,10,1)
for i in labels[1:]:
locs = np.where(Y==i)[0]
Y_new[locs,:] = Y[locs,:]
#locs = np.where(Y_train==10)[0]
#Y_new[locs,:] = Y[locs,:]
#X = (X - X.mean(axis=1, keepdims=True)) / X.std(axis=1, keepdims=True)
sz = X.shape
X_new = np.zeros((sz[3], sz[2], sz[0], sz[1]), dtype=np.float32)
for i in range(sz[3]):
for j in range(sz[2]):
X_new[i,j,:,:] = X[:,:,j,i] # <---- FOR PYTORCH (N x Channels x Width x Height)
if split_data:
return split_train_test((X_new, Y), n_splits=1, test_size=0.2, random_state=42)
return X_new, Y_new
# + id="kO2DdjKqfCar" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 35} outputId="d2f0f30b-d4d7-4af6-e5dc-346bf09e097a" executionInfo={"status": "ok", "timestamp": 1526405547522, "user_tz": -120, "elapsed": 923, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "100979246453766324139"}}
labels = np.arange(0,10,1)
print(labels[1:])
# + id="8VA8Ab7PtpdW" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
def one_hot_encoder(label):
encoder = OneHotEncoder(dtype=np.float32)
label_1hot = encoder.fit_transform(label.reshape(-1,1))
print('The labels are: {}'.format(np.unique(label)))
return label_1hot
# + [markdown] id="7r_M9DL7WzsY" colab_type="text"
# # Load data
# + id="RVTY55DhtpcV" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
root_folder = 'drive/app/svhn/'
# root_folder = 'D:/dev/data/'
# + id="07JKossstpdi" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 109} outputId="2f6b1789-ac4e-4abb-e226-85dff58ae86a" executionInfo={"status": "ok", "timestamp": 1526405571907, "user_tz": -120, "elapsed": 20194, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "100979246453766324139"}}
X_train, Y_train = get_svhn_data(SVHN_URL_TRAIN, root_folder, 'train_32x32.mat',
split_data=False)
X_test, Y_test = get_svhn_data(SVHN_URL_TEST, root_folder, 'test_32x32.mat',
split_data=False)
# X_train, Y_train = get_svhn_data(svhn_dict=svhn_data, split_data=True)
print("Train: [{}, {}], Test: [{}, {}]".format(X_train.shape, Y_train.shape, X_test.shape, Y_test.shape))
# print("Train: [{}, {}]".format(X_train.shape, Y_train.shape))
# + id="nCMfq3D_HtvV" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 72} outputId="ac844f6f-2611-4818-e9b2-7eb35fe78134" executionInfo={"status": "ok", "timestamp": 1526405573667, "user_tz": -120, "elapsed": 1690, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "100979246453766324139"}}
# !ls -l drive/app/svhn
# + id="tDmUQitEgKyn" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# #f, ax = plt.subplots(10,1,figsize=(150, 5))
# i=10
# #for i in range(10):
# idx = np.where(Y_train==i)[0][0]
# #print(idx)
# img = np.zeros([32,32,3])
# img[:,:,0] = X_train[idx,0,:,:]
# img[:,:,1] = X_train[idx,1,:,:]
# img[:,:,1] = X_train[idx,2,:,:]
# plt.imshow(img)
# plt.title(Y_train[idx,:])
# + id="0I4IA-eHtpd0" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 54} outputId="eb46f069-0318-4694-989d-d2b179186d76" executionInfo={"status": "ok", "timestamp": 1526405575359, "user_tz": -120, "elapsed": 669, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "100979246453766324139"}}
Y_train_1hot = one_hot_encoder(Y_train).toarray().view(np.float32)
Y_test_1hot = one_hot_encoder(Y_test).toarray().view(np.float32)
# print(Y_train_1hot[0:2])
# print(type(Y_train_1hot))
# + [markdown] id="KCFxdqUmK0Ix" colab_type="text"
# # IGNORE
# + id="2OlfKOzvtpd4" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
def init_weights(in_features, out_features):
W = np.random.randn(in_features, out_features) / sqrt(in_features)
b = np.zeros(out_features)
return W, b
# + id="yvWmqYMitpeA" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
class HiddenLinearLayer(object):
def __init__(self, in_features, out_features, activation_fn):
self.in_features = in_features
self.out_features = out_features
self.activation_fn = activation_fn
W, b = init_weights(in_features, out_features)
self.W = tf.Variable(W.astype(np.float32))
self.b = tf.Variable(b.astype(np.float32))
def forward(self, x):
return self.activation_fn(tf.matmul(x, self.W) + self.b)
# + id="oK8HO0XGtpeG" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
def weight_variable(shape):
initial = tf.truncated_normal(shape, stddev=0.1)
return tf.Variable(initial)
# + id="VD_ZIbQ_tpeL" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
def bias_variable(shape):
initial = tf.constant(0.1, shape=shape)
return tf.Variable(initial)
# + id="Y0pfrVIrtpeS" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
def conv2d(x, W):
return tf.nn.conv2d(x, W, strides=[1, 1, 1, 1], padding='SAME')
# + id="9n8vMJ_otpeY" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
def max_pool_2x2(x):
return tf.nn.max_pool(x, ksize=[1, 2, 2, 1],
strides=[1, 2, 2, 1], padding='SAME')
# + [markdown] id="flFb78F1LBa2" colab_type="text"
# # Model definition
# + id="9zj7hpiBtpeh" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
class CNN(nn.Module):
def __init__(self, width, height, n_channels):
super(CNN, self).__init__()
self.conv_1 = nn.Conv2d(in_channels=n_channels, out_channels=32, kernel_size=5, stride=1, padding=2)
self.conv_2 = nn.Conv2d(in_channels=32, out_channels=64, kernel_size=5, stride=1, padding=2)
self.conv_3 = nn.Conv2d(in_channels=64, out_channels=128, kernel_size=5, stride=1, padding=2)
self.conv_4 = nn.Conv2d(in_channels=128, out_channels=256, kernel_size=5, stride=1, padding=2)
self.fc5 = nn.Linear(in_features=256*8*8, out_features=1024)
self.fc6 = nn.Linear(in_features=1024, out_features=10)
self.activation = nn.ReLU()
self.maxpool = nn.MaxPool2d(kernel_size=2)
def forward(self, X):
out = self.conv_1(X)
out = self.activation(out)
out = self.conv_2(out)
out = self.activation(out)
#print(out.size())
out = self.maxpool(out)
#print(out.size())
out = self.conv_3(out)
out = self.activation(out)
out = self.conv_4(out)
out = self.activation(out)
#print(out.size())
out = self.maxpool(out)
#print(out.size())
out = out.view(out.size(0), -1)
out = self.fc5(out)
out = self.activation(out)
out = self.fc6(out)
return out
def fit(self, X, Y, criterion, optimizer, epochs, n_batches, batch_size, print_time):
X = torch.from_numpy(X).double()
Y = torch.from_numpy(Y).long()
train_data = torch.utils.data.TensorDataset(X, Y)
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size, shuffle=True)
iteration = 0
for epoch in range(epochs):
for i, (x, y) in enumerate(train_loader):
if torch.cuda.is_available():
x = Variable(x.cuda())
y = Variable(y.cuda())
else:
x = Variable(x)
y = Variable(y)
optimizer.zero_grad()
outputs = self.forward(x)
loss = criterion(outputs, y)
loss.backward()
optimizer.step()
iteration += 1
if iteration%print_time == 0:
print('Epoch: {}, Iteration: {}, Loss: {}'.format(epoch, iteration, loss))
#accuracy = predict(test_loader)
# for epoch in range(epoches):
# X_shuffled, Y_shuffled = shuffle(X, Y)
# for ibatch in range(n_batches):
# X_batch = torch.from_numpy(X_shuffled[ibatch*batch_size:(ibatch+1)*batch_size,:]).double()
# Y_batch = torch.from_numpy(Y_shuffled[ibatch*batch_size:(ibatch+1)*batch_size,:]).double()
# print(type(Y_batch))
# if torch.cuda.is_available():
# X_batch = Variable(X_batch.cuda()).float()
# Y_batch = Variable(Y_batch.cuda()).type(torch.cuda.LongTensor)
# else:
# X_batch = Variable(X_batch).float()
# Y_batch = Variable(Y_batch).type(torch.LongTensor)
# optimizer.zero_grad()
# outputs = self.forward(X_batch)
# loss = cost_fn(outputs, Y_batch)
# loss.backward()
# optimizer.step()
# if ibatch % print_time==0:
# print('Epoch\Batch: {}\{}, Train loss: {}'.format(epoch, ibatch, loss))
def predict(self, X, n_batches, batch_size):
correct = 0
test_cost = 0
total = 0
if ibatch%PRINT_TIME == 0:
for ibatch_test in range(n_batches):
X_batch = torch.from_numpy(X_test[ibatch_test*batch_size:(ibatch_test+1)*batch_size,:])
outputs = self.forward(X_batch)
if first == True:
predicted = torch.argmax(outputs, dim=1)
first = False
else:
predicted = torch.cat((predicted, torch.argmax(outputs, dim=1)))
return predicted
def score(self, Y, predicted):
#predicted = torch.argmax(predicted, axis=1)
accuracy = 100*np.mean(Y == predicted.data.numpy())
return accuracy
# + id="zS9Psa0Ctpes" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
WIDTH = 32
HEIGHT = 32
N_CHANNELS = 3
N_CLASSES = 10
BATCH_SIZE =32
MAX_ITER = 3
N_BATCHES = X_train.shape[0]//BATCH_SIZE
PRINT_TIME = N_BATCHES//N_BATCHES
TEST_N_BATCHES = X_test.shape[0]//BATCH_SIZE
# + [markdown] id="FxNsa9Xitpfa" colab_type="raw"
# W_conv1 = weight_variable([5, 5, 1, 32])
# b_conv1 = bias_variable([32])
# x_image = tf.reshape(x, [-1,28,28,1])
# h_conv1 = tf.nn.relu(conv2d(x_image, W_conv1) + b_conv1)
# h_pool1 = max_pool_2x2(h_conv1)
#
# W_conv2 = weight_variable([5, 5, 32, 64])
# b_conv2 = bias_variable([64])
#
# h_conv2 = tf.nn.relu(conv2d(h_pool1, W_conv2) + b_conv2)
# h_pool2 = max_pool_2x2(h_conv2)
#
# W_fc1 = weight_variable([7 * 7 * 64, 1024])
# b_fc1 = bias_variable([1024])
#
# h_pool2_flat = tf.reshape(h_pool2, [-1, 7*7*64])
# h_fc1 = tf.nn.relu(tf.matmul(h_pool2_flat, W_fc1) + b_fc1)
#
# keep_prob = tf.placeholder(tf.float32)
# h_fc1_drop = tf.nn.dropout(h_fc1, keep_prob)
#
# W_fc2 = weight_variable([1024, 10])
# b_fc2 = bias_variable([10])
#
# y_conv = tf.matmul(h_fc1_drop, W_fc2) + b_fc2
# + id="6dYP6pqOtpgN" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
ann = CNN(WIDTH, HEIGHT, N_CHANNELS)
ann = ann.double()
if torch.cuda.is_available():
ann.cuda()
cost_fn = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(ann.parameters(), lr=0.001)
# + id="xGgBO3sw2bCH" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 239} outputId="ce5d8ff3-968d-43ea-ecaf-c81f5cdb1748" executionInfo={"status": "ok", "timestamp": 1526393634619, "user_tz": -120, "elapsed": 491, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "100979246453766324139"}}
for parameter in list(ann.parameters()):
print(parameter.size())
# + id="HkZGTnFl2tnx" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 35} outputId="92d30a3d-653d-490b-bef6-96075ed4b5d3" executionInfo={"status": "ok", "timestamp": 1526405619177, "user_tz": -120, "elapsed": 1632, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "100979246453766324139"}}
_, _, X_Select, Y_Select = split_train_test((X_train, Y_train), n_splits=1, test_size=0.4, random_state=42)
print(X_Select.shape, Y_Select.shape)
# + id="1r23nqfhGg5x" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 562} outputId="32451069-98ee-46ae-995f-dbed570d907f" executionInfo={"status": "error", "timestamp": 1526405634142, "user_tz": -120, "elapsed": 2081, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "100979246453766324139"}}
ann.fit(X_train, np.squeeze(Y_train), cost_fn, optimizer, 10, N_BATCHES, BATCH_SIZE, PRINT_TIME)
# + id="9yGvGqCrp_Dq" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# !kill -9 -1
# + id="D8c1NOGH6mMn" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
| archive/cnn/pytorch/cnn_pytorch_svhn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Custom priors
#
# The prior probability is a critical element of Bayes theorem.
# However, to keep `uravu` straightforward to use, by default, a broad uniform prior probability is assigned to the `Relationship` object, or if bounds are present these are used as the limits.
#
# Of course this may be ignored and custom priors may be used (*and sometimes it may be necessary that this is done*).
# This tutorial will show **how** custom priors may be used with `uravu`.
#
# Let's start, as always, by producing some synthetic data
import numpy as np
import matplotlib.pyplot as plt
np.random.seed(2)
x = np.linspace(10, 50, 20)
y = .3 * x ** 2 - 1.4 * x + .2
y += y * np.random.randn(20) * 0.05
dy = 3 * x
plt.errorbar(x, y, dy, marker='o', ls='')
plt.show()
# The model for this data is a second order polynomial, below is a function that defines this.
# The `Relationship` object is also created.
def two_degree(x, a, b, c):
return c * x ** 2 + b * x + a
from uravu.relationship import Relationship
modeller = Relationship(two_degree, x, y, ordinate_error=dy)
modeller.max_likelihood('mini')
# The max likelihood (which makes no consideration of the prior) is found,
print(modeller.variable_modes)
# The default prior probabilities for these variables with `uravu` are uniform in the range $[x - 10, x + 10)$, where $x$ is the current value of the variable.
#
# However, if you wanted the prior probability to be a normal distribution, centred on the current value of the varible with a width of 1, it would be necessary to create a custom prior function.
# This function is shown below.
# +
from scipy.stats import norm
def custom_prior():
priors = []
for var in modeller.variable_medians:
priors.append(norm(loc=var, scale=1))
return priors
# -
# Note that the function returns a list of 'frozen' `scipy` RV objects that describe the shape of the priors.
#
# To make use of these priors, they must be passed to the `mcmc` or `nested_sampling` functions as the `prior_function` keyword argument.
modeller.mcmc(prior_function=custom_prior)
modeller.nested_sampling(prior_function=custom_prior)
print(modeller.ln_evidence)
# Any `scipy` [statistical function](https://docs.scipy.org/doc/scipy/reference/stats.html) that has a `logpdf` class method may be used in the definition of priors.
| docs/source/custom_priors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Materialien zu <i>zufall</i>
#
# Autor: <NAME> - <EMAIL>
#
# ## Aufgaben 15 - Simulation (Zahlenlotto)
# <br>
# <i>Die Aufgabe wurde entnommen aus<br>
# <br>
# LS Stochastik<br>
# Grundkurs<br>
# Klett Verlag 1986<br>
# S. 34 Aufgabe 11</i><br>
# <br>
# Mit welcher Wahrscheinlichkeit kommt beim Zahlenlotto "6 aus 49" mindestens ein<br>
# Paar aufeinanderfolgender Zahlen vor?
# <br>
# %run zufall/start
# Eine Ziehung wird dargestellt durch 6 Zufallszahlen zwischen 1 und 49, die ohne<br>
# Wiederholung ermittelt werden; zur einfacheren Behandlung werden die Zahlen sor-<br>
# tiert
ziehung = zuf_zahl((1, 49), 6, w=ohne, s=ja); ziehung
# Eine Funktion, die für eine Ziehung die geforderte Eigenschaft feststellt, ist die fol-<br>
# gende Prozedur, die $1$ (bei vorhandener Eigenschaft) bzw. $0$ (sonst) zurückgibt
# +
def f(x): # x - Liste
for i in range(5):
if x[i+1] == x[i]+1:
return 1
return 0
f(ziehung), f([2, 12, 13, 27, 34, 43]) # Test auf Funktionieren
# -
# Die <b>Simulation</b> wird mit $n$ Ziehungen durchgeführt
# +
n = 10000
sim = [ zuf_zahl((1, 49), 6, w=ohne, s=ja) for i in range(n) ] # n Ziehungen
sim1 = [ f(x) for x in sim ] # Anwenden der Funktion auf jede Ziehung
N(anzahl(1)(sim1) / n) # relative Häufigkeit
# -
# Der <b>theoretische Wert</b> ergibt sich über das Gegenereignis zu $\:
# 1 - \dfrac{\binom{44}{6}}{\binom{49}{6}} \approx 0.495$
| zufall/mat/aufgaben15.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.2
# language: julia
# name: julia-1.5
# ---
# # Terraform Registry to JSON
#
# ### 1. HTTP.get()
# ### 2. JSON.write()
using HTTP
using JSON
using Dates
function SaveJSONToFile(objectType::String, dictionaryOfAllRepos::Dict{String,Dict{String,Any}})
# Create a subfolder of this folder called "terraform-registry"
subfolderpath = joinpath(pwd(), "TfRegistry2JSON")
if !isdir(subfolderpath)
mkdir(subfolderpath)
end
# Name the file "(providers || modules)_(today's date).json"
filename =
objectType * "_" *
string(Dates.today()) *
".json"
filepath = joinpath(subfolderpath, filename)
# Write the JSON to the file
file = open(filepath, "w")
JSON.print(file, dictionaryOfAllRepos)
close(file)
end
function GetAllObjectsFromRegistry(objectType::String)
request = HTTP.get(
"https://registry.terraform.io/v2/" *
objectType *
"?page%5Bsize%5D=100")
dictionaryOfAllRegistryObjects = Dict{String, Dict{String,Any}}()
while true
# Parse HTTP payload into JSON
httpGetRegistryObjectsBodyString = String(request.body)
jsonRepos = JSON.parse(httpGetRegistryObjectsBodyString)
# Iterate through all JSON objects (max = 100)
for jsonRepo in jsonRepos["data"]
# Put the "id" attribute in our subobject
jsonRepo["attributes"]["id"] = jsonRepo["id"]
# Remove the prefix "https://github.com/" from the repoKey
usernameReponame = jsonRepo["attributes"]["source"][19:end]
# Put the user/repo key --> object
dictionaryOfAllRegistryObjects[usernameReponame] = jsonRepo["attributes"]
end
# Break the infinite loop!
if jsonRepos["links"]["next"] == nothing
break
end
# Or Get More!
sleep(1)
request = HTTP.get("https://registry.terraform.io/" * jsonRepos["links"]["next"])
end
# Save it all to file
SaveJSONToFile(objectType, dictionaryOfAllRegistryObjects)
end
@time GetAllObjectsFromRegistry("providers")
@time GetAllObjectsFromRegistry("modules")
| TfRegistry2JSON.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Text Analysis with NLTK
# + [markdown] slideshow={"slide_type": "notes"}
# Author: <NAME>
#
# Date: 2016/04/03
#
# Last review: 2017/04/21
# +
# # %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# import pylab
# Required imports
from wikitools import wiki
from wikitools import category
import nltk
from nltk.tokenize import word_tokenize
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer
from time import time
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from test_helper import Test
import gensim
# -
# ## 1. Corpus acquisition.
# In this notebook we will explore some tools for text analysis available from Python toolboxes.
#
# To do so, we will explore and analyze collections of Wikipedia articles from a given category, using `wikitools`, that makes the capture of content from wikimedia sites very easy.
#
# (*As a side note, there are many other available text collections to work with. In particular, the NLTK library has many examples, that you can explore using the `nltk.download()` tool*.
#
# import nltk
# nltk.download()
#
# *for instance, you can take the gutemberg dataset*
#
# Mycorpus = nltk.corpus.gutenberg
# text_name = Mycorpus.fileids()[0]
# raw = Mycorpus.raw(text_name)
# Words = Mycorpus.words(text_name)
#
# *Also, tools like Gensim or Sci-kit learn include text databases to work with*).
#
# In order to use Wikipedia data, we will select a single category of articles:
site = wiki.Wiki("https://en.wikipedia.org/w/api.php")
# Select a category with a reasonable number of articles (>100)
# cat = "Economics"
cat = "Pseudoscience"
print cat
# You can try with any other categories, but take into account that some categories may contain very few articles. Select a category with at least 100 articles. You can browse the wikipedia category tree here, https://en.wikipedia.org/wiki/Category:Contents, for instance, and select the appropriate one.
#
# We start downloading the text collection.
# +
# Loading category data. This may take a while
print "Loading category data. This may take a while..."
cat_data = category.Category(site, cat)
corpus_titles = []
corpus_text = []
for n, page in enumerate(cat_data.getAllMembersGen()):
print "\rLoading article {0}".format(n + 1),
corpus_titles.append(page.title)
corpus_text.append(page.getWikiText())
n_art = len(corpus_titles)
print "\nLoaded " + str(n_art) + " articles from category " + cat
# -
# Now, we have stored the whole text collection in two lists:
#
# * `corpus_titles`, which contains the titles of the selected articles
# * `corpus_text`, with the text content of the selected wikipedia articles
#
# You can browse the content of the wikipedia articles to get some intuition about the kind of documents that will be processed.
n = 5
print corpus_titles[n]
print corpus_text[n]
# ## 2. Corpus Processing
#
# Topic modelling algorithms process vectorized data. In order to apply them, we need to transform the raw text input data into a vector representation. To do so, we will remove irrelevant information from the text data and preserve as much relevant information as possible to capture the semantic content in the document collection.
#
# Thus, we will proceed with the following steps:
#
# 1. Tokenization
# 2. Homogeneization
# 3. Cleaning
# 4. Vectorization
# ### 2.1. Tokenization
#
# For the first steps, we will use some of the powerfull methods available from the [Natural Language Toolkit](http://www.nltk.org). In order to use the `word_tokenize` method from nltk, you might need to get the appropriate libraries using `nltk.download()`. You must select option "d) Download", and identifier "punkt"
# You can comment this if the package is already available.
# Select option "d) Download", and identifier "punkt"
nltk.download('punkt')
# **Task**: Insert the appropriate call to `word_tokenize` in the code below, in order to get the tokens list corresponding to each Wikipedia article:
# +
corpus_tokens = []
for n, art in enumerate(corpus_text):
print "\rTokenizing article {0} out of {1}".format(n + 1, n_art),
# This is to make sure that all characters have the appropriate encoding.
art = art.decode('utf-8')
# Tokenize each text entry.
# scode: tokens = <FILL IN>
tokens = word_tokenize(art)
# Add the new token list as a new element to corpus_tokens (that will be a list of lists)
# scode: <FILL IN>
corpus_tokens.append(tokens)
print "\n The corpus has been tokenized. Let's check some portion of the first article:"
print corpus_tokens[0][0:30]
# -
Test.assertEquals(len(corpus_tokens), n_art, "The number of articles has changed unexpectedly")
Test.assertTrue(len(corpus_tokens) >= 100,
"Your corpus_tokens has less than 100 articles. Consider using a larger dataset")
# ### 2.2. Homogeneization
#
# By looking at the tokenized corpus you may verify that there are many tokens that correspond to punktuation signs and other symbols that are not relevant to analyze the semantic content. They can be removed using the stemming tool from `nltk`.
#
# The homogeneization process will consist of:
#
# 1. Removing capitalization: capital alphabetic characters will be transformed to their corresponding lowercase characters.
# 2. Removing non alphanumeric tokens (e.g. punktuation signs)
# 3. Stemming/Lemmatization: removing word terminations to preserve the root of the words and ignore grammatical information.
# #### 2.2.1. Filtering
#
# Let us proceed with the filtering steps 1 and 2 (removing capitalization and non-alphanumeric tokens).
# **Task**: Convert all tokens in `corpus_tokens` to lowercase (using `.lower()` method) and remove non alphanumeric tokens (that you can detect with `.isalnum()` method). You can do it in a single line of code...
# +
corpus_filtered = []
for n, token_list in enumerate(corpus_tokens):
print "\rFiltering article {0} out of {1}".format(n + 1, n_art),
# Convert all tokens in token_list to lowercase, remove non alfanumeric tokens and stem.
# Store the result in a new token list, clean_tokens.
# scode: filtered_tokens = <FILL IN>
filtered_tokens = [token.lower() for token in token_list if token.isalnum()]
# Add art to corpus_filtered
# scode: <FILL IN>
corpus_filtered.append(filtered_tokens)
print "\nLet's check the first tokens from document 0 after filtering:"
print corpus_filtered[0][0:30]
# -
Test.assertTrue(all([c==c.lower() for c in corpus_filtered[23]]), 'Capital letters have not been removed')
Test.assertTrue(all([c.isalnum() for c in corpus_filtered[13]]), 'Non alphanumeric characters have not been removed')
# #### 2.2.2. Stemming vs Lemmatization
#
# At this point, we can choose between applying a simple stemming or ussing lemmatization. We will try both to test their differences.
# **Task**: Apply the `.stem()` method, from the stemmer object created in the first line, to `corpus_filtered`.
# +
# Select stemmer.
stemmer = nltk.stem.SnowballStemmer('english')
corpus_stemmed = []
for n, token_list in enumerate(corpus_filtered):
print "\rStemming article {0} out of {1}".format(n + 1, n_art),
# Convert all tokens in token_list to lowercase, remove non alfanumeric tokens and stem.
# Store the result in a new token list, clean_tokens.
# scode: stemmed_tokens = <FILL IN>
stemmed_tokens = [stemmer.stem(token) for token in token_list]
# Add art to the stemmed corpus
# scode: <FILL IN>
corpus_stemmed.append(stemmed_tokens)
print "\nLet's check the first tokens from document 0 after stemming:"
print corpus_stemmed[0][0:30]
# -
Test.assertTrue((len([c for c in corpus_stemmed[0] if c!=stemmer.stem(c)]) < 0.1*len(corpus_stemmed[0])),
'It seems that stemming has not been applied properly')
# Alternatively, we can apply lemmatization. For english texts, we can use the lemmatizer from NLTK, which is based on [WordNet](http://wordnet.princeton.edu). If you have not used wordnet before, you will likely need to download it from nltk
# You can comment this if the package is already available.
# Select option "d) Download", and identifier "wordnet"
nltk.download("wordnet")
# **Task**: Apply the `.lemmatize()` method, from the WordNetLemmatizer object created in the first line, to `corpus_filtered`.
# +
wnl = WordNetLemmatizer()
# Select stemmer.
corpus_lemmat = []
for n, token_list in enumerate(corpus_filtered):
print "\rLemmatizing article {0} out of {1}".format(n + 1, n_art),
# scode: lemmat_tokens = <FILL IN>
lemmat_tokens = [wnl.lemmatize(token) for token in token_list]
# Add art to the stemmed corpus
# scode: <FILL IN>
corpus_lemmat.append(lemmat_tokens)
print "\nLet's check the first tokens from document 0 after lemmatization:"
print corpus_lemmat[0][0:30]
# -
# One of the advantages of the lemmatizer method is that the result of lemmmatization is still a true word, which is more advisable for the presentation of text processing results and lemmatization.
#
# However, without using contextual information, lemmatize() does not remove grammatical differences. This is the reason why "is" or "are" are preserved and not replaced by infinitive "be".
#
# As an alternative, we can apply .lemmatize(word, pos), where 'pos' is a string code specifying the part-of-speech (pos), i.e. the grammatical role of the words in its sentence. For instance, you can check the difference between `wnl.lemmatize('is')` and `wnl.lemmatize('is, pos='v')`.
# ### 2.3. Cleaning
#
# The third step consists of removing those words that are very common in language and do not carry out usefull semantic content (articles, pronouns, etc).
#
# Once again, we might need to load the stopword files using the download tools from `nltk`
# You can comment this if the package is already available.
# Select option "d) Download", and identifier "stopwords"
nltk.download("stopwords")
# ** Task**: In the second line below we read a list of common english stopwords. Clean `corpus_stemmed` by removing all tokens in the stopword list.
# +
corpus_clean = []
stopwords_en = stopwords.words('english')
n = 0
for token_list in corpus_stemmed:
n += 1
print "\rRemoving stopwords from article {0} out of {1}".format(n, n_art),
# Remove all tokens in the stopwords list and append the result to corpus_clean
# scode: clean_tokens = <FILL IN>
clean_tokens = [token for token in token_list if token not in stopwords_en]
# scode: <FILL IN>
corpus_clean.append(clean_tokens)
print "\n Let's check tokens after cleaning:"
print corpus_clean[0][0:30]
# -
Test.assertTrue(len(corpus_clean) == n_art, 'List corpus_clean does not contain the expected number of articles')
Test.assertTrue(len([c for c in corpus_clean[0] if c in stopwords_en])==0, 'Stopwords have not been removed')
# ### 2.4. Vectorization
#
# Up to this point, we have transformed the raw text collection of articles in a list of articles, where each article is a collection of the word roots that are most relevant for semantic analysis. Now, we need to convert these data (a list of token lists) into a numerical representation (a list of vectors, or a matrix). To do so, we will start using the tools provided by the `gensim` library.
#
# As a first step, we create a dictionary containing all tokens in our text corpus, and assigning an integer identifier to each one of them.
# +
# Create dictionary of tokens
D = gensim.corpora.Dictionary(corpus_clean)
n_tokens = len(D)
print "The dictionary contains {0} tokens".format(n_tokens)
print "First tokens in the dictionary: "
for n in range(10):
print str(n) + ": " + D[n]
# -
# In the second step, let us create a numerical version of our corpus using the `doc2bow` method. In general, `D.doc2bow(token_list)` transform any list of tokens into a list of tuples `(token_id, n)`, one per each token in `token_list`, where `token_id` is the token identifier (according to dictionary `D`) and `n` is the number of occurrences of such token in `token_list`.
# ** Task**: Apply the `doc2bow` method from gensim dictionary `D`, to all tokens in every article in `corpus_clean`. The result must be a new list named `corpus_bow` where each element is a list of tuples `(token_id, number_of_occurrences)`.
# Transform token lists into sparse vectors on the D-space
# scode: corpus_bow = <FILL IN>
corpus_bow = [D.doc2bow(doc) for doc in corpus_clean]
Test.assertTrue(len(corpus_bow)==n_art, 'corpus_bow has not the appropriate size')
# At this point, it is good to make sure to understand what has happened. In `corpus_clean` we had a list of token lists. With it, we have constructed a Dictionary, `D`, which assign an integer identifier to each token in the corpus.
# After that, we have transformed each article (in `corpus_clean`) in a list tuples `(id, n)`.
print "Original article (after cleaning): "
print corpus_clean[0][0:30]
print "Sparse vector representation (first 30 components):"
print corpus_bow[0][0:30]
print "The first component, {0} from document 0, states that token 0 ({1}) appears {2} times".format(
corpus_bow[0][0], D[0], corpus_bow[0][0][1])
# Note that we can interpret each element of corpus_bow as a `sparse_vector`. For example, a list of tuples
#
# [(0, 1), (3, 3), (5,2)]
#
# for a dictionary of 10 elements can be represented as a vector, where any tuple `(id, n)` states that position `id` must take value `n`. The rest of positions must be zero.
#
# [1, 0, 0, 3, 0, 2, 0, 0, 0, 0]
#
# These sparse vectors will be the inputs to the topic modeling algorithms.
#
# Note that, at this point, we have built a Dictionary containing
print "{0} tokens".format(len(D))
# and a bow representation of a corpus with
print "{0} Wikipedia articles".format(len(corpus_bow))
# Before starting with the semantic analyisis, it is interesting to observe the token distribution for the given corpus.
# +
# SORTED TOKEN FREQUENCIES (I):
# Create a "flat" corpus with all tuples in a single list
corpus_bow_flat = [item for sublist in corpus_bow for item in sublist]
# Initialize a numpy array that we will use to count tokens.
# token_count[n] should store the number of ocurrences of the n-th token, D[n]
token_count = np.zeros(n_tokens)
# Count the number of occurrences of each token.
for x in corpus_bow_flat:
# Update the proper element in token_count
# scode: <FILL IN>
token_count[x[0]] += x[1]
# Sort by decreasing number of occurences
ids_sorted = np.argsort(- token_count)
tf_sorted = token_count[ids_sorted]
# -
# `ids_sorted` is a list of all token ids, sorted by decreasing number of occurrences in the whole corpus. For instance, the most frequent term is
print D[ids_sorted[0]]
# which appears
print "{0} times in the whole corpus".format(tf_sorted[0])
# In the following we plot the most frequent terms in the corpus.
# +
# SORTED TOKEN FREQUENCIES (II):
plt.rcdefaults()
# Example data
n_bins = 25
hot_tokens = [D[i] for i in ids_sorted[n_bins-1::-1]]
y_pos = np.arange(len(hot_tokens))
z = tf_sorted[n_bins-1::-1]/n_art
plt.barh(y_pos, z, align='center', alpha=0.4)
plt.yticks(y_pos, hot_tokens)
plt.xlabel('Average number of occurrences per article')
plt.title('Token distribution')
plt.show()
display()
# +
# SORTED TOKEN FREQUENCIES:
# Example data
plt.semilogy(tf_sorted)
plt.ylabel('Total number of occurrences')
plt.xlabel('Token rank')
plt.title('Token occurrences')
plt.show()
display()
# -
# ** Exercise**: There are usually many tokens that appear with very low frequency in the corpus. Count the number of tokens appearing only once, and what is the proportion of them in the token list.
# +
# scode: <WRITE YOUR CODE HERE>
# Example data
cold_tokens = [D[i] for i in ids_sorted if tf_sorted[i]==1]
print "There are {0} cold tokens, which represent {1}% of the total number of tokens in the dictionary".format(
len(cold_tokens), float(len(cold_tokens))/n_tokens*100)
# -
# ** Exercise**: Represent graphically those 20 tokens that appear in the highest number of articles. Note that you can use the code above (headed by `# SORTED TOKEN FREQUENCIES`) with a very minor modification.
# +
# scode: <WRITE YOUR CODE HERE>
# SORTED TOKEN FREQUENCIES (I):
# Count the number of occurrences of each token.
token_count2 = np.zeros(n_tokens)
for x in corpus_bow_flat:
token_count2[x[0]] += (x[1]>0)
# Sort by decreasing number of occurences
ids_sorted2 = np.argsort(- token_count2)
tf_sorted2 = token_count2[ids_sorted2]
# SORTED TOKEN FREQUENCIES (II):
# Example data
n_bins = 25
hot_tokens2 = [D[i] for i in ids_sorted2[n_bins-1::-1]]
y_pos2 = np.arange(len(hot_tokens2))
z2 = tf_sorted2[n_bins-1::-1]/n_art
plt.figure()
plt.barh(y_pos2, z2, align='center', alpha=0.4)
plt.yticks(y_pos2, hot_tokens2)
plt.xlabel('Number of articles')
plt.title('Token distribution')
plt.show()
display()
# -
# ** Exercise**: Count the number of tokens appearing only in a single article.
# +
# scode: <WRITE YOUR CODE HERE>
# -
# ** Exercise** (*All in one*): Note that, for pedagogical reasons, we have used a different `for` loop for each text processing step creating a new `corpus_xxx` variable after each step. For very large corpus, this could cause memory problems.
#
# As a summary exercise, repeat the whole text processing, starting from corpus_text up to computing the bow, with the following modifications:
#
# 1. Use a single `for` loop, avoiding the creation of any intermediate corpus variables.
# 2. Use lemmatization instead of stemming.
# 3. Remove all tokens appearing in only one document and less than 2 times.
# 4. Save the result in a new variable `corpus_bow1`.
# +
# scode: <WRITE YOUR CODE HERE>
# -
# ** Exercise** (*Visualizing categories*): Repeat the previous exercise with a second wikipedia category. For instance, you can take "communication".
#
# 1. Save the result in variable `corpus_bow2`.
# 2. Determine the most frequent terms in `corpus_bow1` (`term1`) and `corpus_bow2` (`term2`).
# 3. Transform each article in `corpus_bow1` and `corpus_bow2` into a 2 dimensional vector, where the first component is the frecuency of `term1` and the second component is the frequency of `term2`
# 4. Draw a dispersion plot of all 2 dimensional points, using a different marker for each corpus. Could you differentiate both corpora using the selected terms only? What if the 2nd most frequent term is used?
# +
# scode: <WRITE YOUR CODE HERE>
# -
# ** Exercise ** (bigrams): `nltk` provides an utility to compute n-grams from a list of tokens, in `nltk.util.ngrams`. Join all tokens in `corpus_clean` in a single list and compute the bigrams. Plot the 20 most frequent bigrams in the corpus.
# scode: <WRITE YOUR CODE HERE>
# Check the code below to see how ngrams works, and adapt it to solve the exercise.
# from nltk.util import ngrams
# sentence = 'this is a foo bar sentences and i want to ngramize it'
# sixgrams = ngrams(sentence.split(), 2)
# for grams in sixgrams:
# print grams
| TM1.IntrodNLP/NLP_py2_wikitools/databricks/TM1_NLP_db.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/hannsjaspherelalto/OOP-1-2/blob/main/OOP_Concepts_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="rniubDPje2QU" outputId="e6c51d0c-6ecf-4346-cba8-4f273dbf4c68"
class Birds:
def __init__(self,bird_name):
self.bird_name = bird_name
def flying_birds(self):
print(f"{self.bird_name} flies above the sky")
def non_flying_birds(self):
print(f"{self.bird_name} is the national bird of the Philippines")
vulture = Birds("Vulture")
crane = Birds("Crane")
emu = Birds("Emumu")
vulture.flying_birds()
crane.flying_birds()
emu.non_flying_birds()
# + colab={"base_uri": "https://localhost:8080/"} id="87yNH8gle5N6" outputId="0ce246e7-89ad-487e-f2d8-e8ab8001a08c"
class foo:
def __init__(self,a,b):
self.__a = a
self.__b = b
def add(self):
return self.__a +self.__b
number = foo(3,4)
number.add()
number.a = 7
number.add()
# + colab={"base_uri": "https://localhost:8080/"} id="pvN3bPqpe7hz" outputId="7c5b786e-c951-4720-d3b0-a7516b9d8727"
class Counter:
def __init__(self):
self.__current = 0
def increment(self):
self.__current +=1
def value(self):
return self.__current
def reset(self):
self.__current = 0
num = Counter()
num.increment()
num.increment()
num.increment()
num.counter = 1
num.value()
# + colab={"base_uri": "https://localhost:8080/"} id="8M3pcE41e9bS" outputId="7172404a-0f7f-4323-c403-b5f45d75e758"
class Person:
def __init__(self, firstname,surname):
self.firstname = firstname
self.surname = surname
def printname(self):
print(self.firstname,self.surname)
person = Person("Ana", "Santos")
person.printname()
class Teacher(Person):
pass
person2 = Teacher("Maria", "Sayo")
person2.printname()
class Student(Person):
pass
person3 = Student("Hanns", "Elalto")
person3.printname()
# + colab={"base_uri": "https://localhost:8080/"} id="Hb0xFFwDe_Cb" outputId="b7bd6726-70b5-4537-8d57-abf2b4e936fc"
class RegularPolygon:
def __init__(self,side):
self.side = side
class Square(RegularPolygon):
def area(self):
return self.side * self.side
class EquilateralTriangle(RegularPolygon):
def area(self):
return self.side * self.side * 0.433
object = Square(4)
print(object.area())
object2 = EquilateralTriangle(3)
print(object2.area())
# + colab={"base_uri": "https://localhost:8080/"} id="vZNiFc-YfZfN" outputId="bcce4609-8db5-4f73-c43e-7b65f2feb726"
class Person:
def __init__(self, name, pre, mid, fin):
self._name = name
self._pre = pre #<-encapsulation
self._mid = mid
self._fin = fin
class Student1(Person): #<-inheritance
def Grade(self): #<-polymorphism
ave = 0
ave = ((self._pre + self._mid + self._fin)/3)
print(("-"*20 + "\n") + "Preliminaries: " + str(self._pre))
print("Midterms : " + str(self._mid))
print("Finals : " + str(self._fin))
print("Average : " + str(round(ave,2)))
class Student2(Person): #<-inheritance
def Grade(self): #<-polymorphism
ave = 0
ave = ((self._pre*0.30) + (self._mid*0.30) + (self._fin*0.40))
print(("-"*20 + "\n") + "Preliminaries: " + str(self._pre))
print("Midterms : " + str(self._mid))
print("Finals : " + str(self._fin))
print(("-"*20 + "\n") + "Average : " + str(round(ave,2)))
class Student3(Person): #<-inheritance
def Grade(self): #<-polymorphism
ave = 0
ave = ((self._pre*0.25) + (self._mid*0.25) + (self._fin*0.50))
print(("-"*20 + "\n") + "Preliminaries: " + str(self._pre))
print("Midterms : " + str(self._mid))
print("Finals : " + str(self._fin))
print(("-"*20 + "\n") + "Average : " + str(round(ave,2)))
std1 = Student1("Hanns", 90, 87, 94)
std2 = Student2("Jaspher", 89, 86, 91)
std3 = Student3("Elalto", 100, 79, 73)
in_name = (input("Student Name: "))
if in_name == "Hanns":
std1.Grade() #student names are required
elif in_name == "Jaspher": #for the privacy of grades
std2.Grade()
elif in_name == "Elalto":
std3.Grade()
else:
print("The student's name is not on the database.")
| OOP_Concepts_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
import pandas as pd
import cPickle
import multiprocessing
import matplotlib.pyplot as plt
# %matplotlib inline
from IPython.display import clear_output
# ### Reading tweets
bank_train = pd.read_csv('../data/bank_train.csv', sep='\t', encoding='utf-8')
bank_train.head()
texts, labels = bank_train.text.values, bank_train.label.values
# ### Reading vocabulary and embeddings
word2id, embeddings = cPickle.load(open('../data/w2v/vectors_l.pkl', 'rb'))
# word2id, embeddings = cPickle.load(open('../data/w2v/parkin_vectors.pkl', 'rb'))
# + active=""
# # For a case when <EOS> symbol is not presented in downloaded embeddings
# word2id[u'</s>'] = embeddings.shape[0]
# embeddings = np.concatenate((embeddings, np.zeros((1, embeddings.shape[1]))))
# -
vocabulary = word2id.keys()
eos_id = word2id[u'</s>']
# ### Lemmatizing and replacing words with ids
# +
from nltk.tokenize import RegexpTokenizer
import pymorphy2
tokenizer = RegexpTokenizer(u'[а-яА-Я]+')
morph = pymorphy2.MorphAnalyzer()
def text2seq(text):
tokens_norm = [morph.parse(w)[0].normal_form for w in tokenizer.tokenize(text)]
return [word2id[w] for w in tokens_norm if w in vocabulary] + [eos_id]
sample = texts[49]
print sample
print u' '.join(tokenizer.tokenize(sample))
print u' '.join([morph.parse(w)[0].normal_form for w in tokenizer.tokenize(sample)])
print text2seq(sample)
# + active=""
# # TODO: adding new words to vocabulary
# from fuzzywuzzy import fuzz
# metric = fuzz.ratio
# if word not in vocabulary:
# max_score = 0
# closest_word = vocabulary[0]
# for w in vocabulary:
# score = metric(word, w)
# if score > max_score:
# max_score = score
# closest_word = w
# ...
# + active=""
# %%time
# pool = multiprocessing.Pool()
# X = pool.map(text2seq, texts)
# -
# Dumping
# + active=""
# cPickle.dump(X, open('../data/X_bank_train_full.pkl', 'wb'))
# -
# Loading
X = cPickle.load(open('../data/X_bank_train_full.pkl', 'rb'))
# Distribution of sequences' lengths
length_max = max(map(len, X))
plt.figure(figsize=(10,7))
plt.hist(map(len, X), bins=length_max)
plt.title(u"Банки", fontsize=30)
plt.xlabel(u"Кол-во слов в документе", fontsize=30)
plt.ylabel(u"Кол-во документов", fontsize=30)
plt.tick_params(labelsize=20)
plt.show()
sum(map(len, X)) / len(X)
# ### Zero padding
X = [x + [eos_id]*(length_max - len(x)) for x in X]
# ### Examples
for x in X[:4]:
print x
# ### One-hot encode labels
X = np.array(X)
y = bank_train.label.values
# +
def cls2probs(cls):
if cls == -1:
return [1., 0., 0.]
elif cls == 0:
return [0., 1., 0.]
else:
return [0., 0., 1.]
y = np.array([cls2probs(cls) for cls in y])
# -
# ### Split into train and validation sets
# +
from sklearn.model_selection import train_test_split
VAL_SIZE = 0.1
X_tr, X_val, y_tr, y_val = train_test_split(X, y, test_size=VAL_SIZE, random_state=40)
# + active=""
# # To use if y is one-dimensional array
# from collections import Counter
#
# print Counter(y_tr)
# print Counter(y_val)
# -
print "Train class frequencies:\t", [col.nonzero()[0].shape[0] for col in y_tr.transpose()]
print "Validation class frequencies:\t", [col.nonzero()[0].shape[0] for col in y_val.transpose()]
print "Constant classifier's validation accuracy:\t", [col.nonzero()[0].shape[0] for col in y_val.transpose()][1] * 1. / y_val.shape[0]
# ### Resampling
# + active=""
# # from imblearn.under_sampling import RandomUnderSampler
# # rus = RandomUnderSampler()
# # _y = np.argmax(y_tr, 1) - 1
# # X_resampled, y_resampled = rus.fit_sample(X_tr, _y)
# # y_resampled = np.array([cls2probs(cls) for cls in y_resampled])
#
# neutral_indices = np.random.choice(np.nonzero(y_tr[:,1])[0], np.nonzero(y_tr[:,0])[0].shape[0], replace=False)
# negative_indices = np.nonzero(y_tr[:,0])[0]
# positive_indices = np.nonzero(y_tr[:,2])[0]
#
# X_resampled = X_tr[np.concatenate([negative_indices, neutral_indices, positive_indices])]
# y_resampled = y_tr[np.concatenate([negative_indices, neutral_indices, positive_indices])]
# -
# ---
# # Network learning
# +
import tensorflow as tf
from tensorflow.python.ops.rnn_cell import LSTMCell, GRUCell
from tensorflow.python.ops.rnn import bidirectional_dynamic_rnn as bi_rnn
from tensorflow.contrib.layers import fully_connected
from utils import *
# -
y_pred_major = np.zeros(y_val.shape)
y_pred_major[:,0] = 1.
print "Constant classifier's F-score on validation set:", f_macro(y_val, y_pred_major)
# ### Bi-RNN
# +
EMBED_DIM = 300
HIDDEN_SIZE = 50
NUM_CLASSES = 3
tf.reset_default_graph()
batch_ph = tf.placeholder(tf.int32, [None, None])
target_ph = tf.placeholder(tf.float32, [None, NUM_CLASSES])
seq_len_ph = tf.placeholder(tf.int32, [None])
keep_prob_ph = tf.placeholder(tf.float32)
embeddings_ph = tf.placeholder(tf.float32, [len(vocabulary), EMBED_DIM])
embeddings_var = tf.Variable(tf.constant(0., shape=[len(vocabulary), EMBED_DIM]), trainable=False)
init_embeddings = embeddings_var.assign(embeddings_ph)
batch_embedded = tf.nn.embedding_lookup(embeddings_var, batch_ph)
# Bi-RNN layers
outputs,_ = bi_rnn(GRUCell(HIDDEN_SIZE), GRUCell(HIDDEN_SIZE),
inputs=batch_embedded,sequence_length=seq_len_ph, dtype=tf.float32, scope="bi_rnn1")
outputs = tf.concat(2, outputs)
# outputs2,_ = bi_rnn(GRUCell(HIDDEN_SIZE), GRUCell(HIDDEN_SIZE),
# inputs=outputs,sequence_length=seq_len_ph, dtype=tf.float32, scope="bi_rnn2")
# outputs2 = tf.concat(2, outputs2)
# Last output of Bi-RNN
output = outputs[:,0,:]
# Dropout
drop = tf.nn.dropout(output, keep_prob_ph)
# Fully connected layer
W = tf.Variable(tf.truncated_normal([HIDDEN_SIZE * 2, NUM_CLASSES], stddev=0.1), name="W")
b = tf.Variable(tf.constant(0., shape=[NUM_CLASSES]), name="b")
y_hat = tf.nn.xw_plus_b(drop, W, b, name="scores")
# +
# Adam parameters
LEARNING_RATE = 1e-4
EPSILON = 1e-5
BETA1 = 0.9
BETA2 = 0.9
# L2 regularization coefficient
BETA = 0
cross_entropy = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(y_hat, target_ph), name="cross_entropy")
l2_loss = tf.nn.l2_loss(W, name="l2_loss")
loss = cross_entropy + l2_loss * BETA
optimizer = tf.train.AdamOptimizer(learning_rate=LEARNING_RATE, beta1=BETA1, beta2=BETA2,
epsilon=EPSILON).minimize(loss)
# optimizer = tf.train.MomentumOptimizer(learning_rate=1e-1, momentum=0.1).minimize(loss)
accuracy = tf.reduce_mean(tf.cast(tf.equal(tf.argmax(target_ph, 1), tf.argmax(y_hat, 1)), tf.float32))
# +
DROPOUT = 0.6 # Probability of keeping a neuron
BATCH_SIZE = 256
EPOCHS = 100
results = []
train_batch_generator = batch_generator(X_tr, y_tr, BATCH_SIZE)
loss_tr_l = []
loss_val_l = []
ce_tr_l = [] # Cross-entropy
ce_val_l = []
acc_tr_l = [] # Accuracy
acc_val_l = []
f_macro_tr_l = []
f_macro_val_l = []
f_fair_tr_l = []
f_fair_val_l = []
min_ce = 0.32
with tf.Session() as sess:
sess.run(tf.initialize_all_variables())
sess.run(init_embeddings, feed_dict={embeddings_ph: embeddings})
print "Start learning..."
for epoch in range(EPOCHS):
for i in range(int(X_tr.shape[0] / BATCH_SIZE)):
x_batch, y_batch = train_batch_generator.next()
seq_len_tr = np.array([list(x).index(eos_id) + 1 for x in x_batch])
sess.run(optimizer, feed_dict={batch_ph: x_batch, target_ph: y_batch,
seq_len_ph: seq_len_tr, keep_prob_ph: DROPOUT})
y_pred_tr, ce_tr, loss_tr, acc_tr = sess.run([y_hat, cross_entropy, loss, accuracy],
feed_dict={batch_ph: x_batch, target_ph: y_batch,
seq_len_ph: seq_len_tr, keep_prob_ph: 1.0})
seq_len_val = np.array([list(x).index(eos_id) + 1 for x in X_val])
y_pred_val, ce_val, loss_val, acc_val = sess.run([y_hat, cross_entropy, loss, accuracy],
feed_dict={batch_ph: X_val, target_ph: y_val,
seq_len_ph: seq_len_val, keep_prob_ph: 1.0})
y_pred_tr = np.array([cls2probs(cls) for cls in np.argmax(y_pred_tr, 1) - 1])
y_pred_val = np.array([cls2probs(cls) for cls in np.argmax(y_pred_val, 1) - 1])
f_macro_tr, f_micro_tr = f_macro(y_batch, y_pred_tr), f_micro(y_batch, y_pred_tr)
f_macro_val, f_micro_val = f_macro(y_val, y_pred_val), f_micro(y_val, y_pred_val)
loss_tr_l.append(loss_tr)
loss_val_l.append(loss_val)
ce_tr_l.append(ce_tr)
ce_val_l.append(ce_val)
acc_tr_l.append(acc_tr)
acc_val_l.append(acc_val)
f_macro_tr_l.append(f_macro_tr)
f_macro_val_l.append(f_macro_val)
clear_output(wait=True)
print "epoch: {}".format(epoch)
print "\t Train loss: {:.3f}\t ce: {:.3f}\t acc: {:.3f}\t f_macro: {:.3f}".format(
loss_tr, ce_tr, acc_tr, f_macro_tr)
print "\t Valid loss: {:.3f}\t ce: {:.3f}\t acc: {:.3f}\t f_macro: {:.3f}".format(
loss_val, ce_val, acc_val, f_macro_val)
plt.figure(figsize=(15,10))
plt.plot(ce_tr_l, color='blue', label='ce_tr')
plt.plot(ce_val_l, color='red', label='ce_val')
plt.plot(f_macro_val_l, color='green', label='f_macro_val')
plt.xlim(0, EPOCHS - 1)
plt.ylim(0, 1)
plt.legend()
plt.show()
saver = tf.train.Saver()
saver.save(sess, 'model_birnn')
results.append([acc_val_l[-1], f_macro_val_l[-1]])
# -
# ### Cross-validation
# +
from sklearn.model_selection import StratifiedKFold, KFold
DROPOUT = 0.5 # Probability of keeping a neuron
BATCH_SIZE = 256
EPOCHS = 100
skf = KFold(10, shuffle=True, random_state=42)
results = []
for train_index, test_index in skf.split(X, y):
X_tr, X_val = X[train_index], X[test_index]
y_tr, y_val = y[train_index], y[test_index]
train_batch_generator = batch_generator(X_tr, y_tr, BATCH_SIZE)
loss_tr_l = []
loss_val_l = []
ce_tr_l = [] # Cross-entropy
ce_val_l = []
acc_tr_l = [] # Accuracy
acc_val_l = []
f_macro_tr_l = []
f_macro_val_l = []
f_fair_tr_l = []
f_fair_val_l = []
min_ce = 0.32
with tf.Session() as sess:
sess.run(tf.initialize_all_variables())
sess.run(init_embeddings, feed_dict={embeddings_ph: embeddings})
print "Start learning..."
for epoch in range(EPOCHS):
for i in range(int(X_tr.shape[0] / BATCH_SIZE)):
x_batch, y_batch = train_batch_generator.next()
seq_len_tr = np.array([list(x).index(eos_id) + 1 for x in x_batch])
sess.run(optimizer, feed_dict={batch_ph: x_batch, target_ph: y_batch,
seq_len_ph: seq_len_tr, keep_prob_ph: DROPOUT})
y_pred_tr, ce_tr, loss_tr, acc_tr = sess.run([y_hat, cross_entropy, loss, accuracy],
feed_dict={batch_ph: x_batch, target_ph: y_batch,
seq_len_ph: seq_len_tr, keep_prob_ph: 1.0})
seq_len_val = np.array([list(x).index(eos_id) + 1 for x in X_val])
y_pred_val, ce_val, loss_val, acc_val = sess.run([y_hat, cross_entropy, loss, accuracy],
feed_dict={batch_ph: X_val, target_ph: y_val,
seq_len_ph: seq_len_val, keep_prob_ph: 1.0})
y_pred_tr = np.array([cls2probs(cls) for cls in np.argmax(y_pred_tr, 1) - 1])
y_pred_val = np.array([cls2probs(cls) for cls in np.argmax(y_pred_val, 1) - 1])
f_macro_tr, f_micro_tr = f_macro(y_batch, y_pred_tr), f_micro(y_batch, y_pred_tr)
f_macro_val, f_micro_val = f_macro(y_val, y_pred_val), f_micro(y_val, y_pred_val)
loss_tr_l.append(loss_tr)
loss_val_l.append(loss_val)
ce_tr_l.append(ce_tr)
ce_val_l.append(ce_val)
acc_tr_l.append(acc_tr)
acc_val_l.append(acc_val)
f_macro_tr_l.append(f_macro_tr)
f_macro_val_l.append(f_macro_val)
clear_output(wait=True)
print "epoch: {}".format(epoch)
print "\t Train loss: {:.3f}\t ce: {:.3f}\t acc: {:.3f}\t f_macro: {:.3f}".format(
loss_tr, ce_tr, acc_tr, f_macro_tr)
print "\t Valid loss: {:.3f}\t ce: {:.3f}\t acc: {:.3f}\t f_macro: {:.3f}".format(
loss_val, ce_val, acc_val, f_macro_val)
plt.figure(figsize=(15,10))
plt.plot(ce_tr_l, color='blue', label='ce_tr')
plt.plot(ce_val_l, color='red', label='ce_val')
plt.plot(f_macro_val_l, color='green', label='f_macro_val')
plt.xlim(0, EPOCHS - 1)
plt.ylim(0, 1)
plt.legend()
plt.show()
results.append([acc_val_l[-1], f_macro_val_l[-1]])
# -
min(ce_val_l), max(acc_val_l), max(f_macro_val_l)
# ---
# ### Bi-RNN with Attention mechanism
# Attention mechanism formulas and notation
# $$
# \upsilon_{t}=\tanh{(W_{\omega}\left[\overrightarrow{h_{t}},\overleftarrow{h_{t}}\right]+b_{\omega})}\\
# \alpha_{t}=\frac{\exp{(\upsilon_{t}^{T}u_{\omega})}}{\sum_{j=1}^{T}\exp{(\upsilon_{j}^{T}u_{\omega})}}\\
# \upsilon=\sum_{t=1}^{T}\alpha_{t}h_{t}
# $$
# +
EMBED_DIM = 300
SEQ_LENGTH = length_max
HIDDEN_SIZE = 100
ATTENTION_SIZE = 100
NUM_CLASSES = 3
tf.reset_default_graph()
batch_ph = tf.placeholder(tf.int32, [None, SEQ_LENGTH])
target_ph = tf.placeholder(tf.float32, [None, NUM_CLASSES])
seq_len_ph = tf.placeholder(tf.int32, [None])
keep_prob_ph = tf.placeholder(tf.float32)
embeddings_ph = tf.placeholder(tf.float32, [len(vocabulary), EMBED_DIM])
embeddings_var = tf.Variable(tf.constant(0., shape=[len(vocabulary), EMBED_DIM]), trainable=False)
init_embeddings = embeddings_var.assign(embeddings_ph)
batch_embedded = tf.nn.embedding_lookup(embeddings_var, batch_ph)
# Bi-RNN layers
outputs,_ = bi_rnn(GRUCell(HIDDEN_SIZE), GRUCell(HIDDEN_SIZE),
inputs=batch_embedded,sequence_length=seq_len_ph, dtype=tf.float32, scope="bi_rnn1")
outputs = tf.concat(2, outputs)
# outputs2,_ = bi_rnn(GRUCell(HIDDEN_SIZE), GRUCell(HIDDEN_SIZE),
# inputs=outputs,sequence_length=seq_len_ph, dtype=tf.float32, scope="bi_rnn2")
# outputs2 = tf.concat(2, outputs2)
# Attention mechanism
W_omega = tf.Variable(tf.random_normal([2 * HIDDEN_SIZE, ATTENTION_SIZE], stddev=0.1))
b_omega = tf.Variable(tf.random_normal([ATTENTION_SIZE], stddev=0.1))
u_omega = tf.Variable(tf.random_normal([ATTENTION_SIZE], stddev=0.1))
v = tf.tanh(tf.matmul(tf.reshape(outputs, [-1, 2 * HIDDEN_SIZE]), W_omega) + tf.reshape(b_omega, [1, -1]))
vu = tf.matmul(v, tf.reshape(u_omega, [-1, 1]))
exps = tf.reshape(tf.exp(vu), [-1, SEQ_LENGTH])
alphas = exps / tf.reshape(tf.reduce_sum(exps, 1), [-1, 1])
# Output of Bi-RNN is reduced with attention vector
output = tf.reduce_sum(outputs * tf.reshape(alphas, [-1, SEQ_LENGTH, 1]), 1)
# Dropout
drop = tf.nn.dropout(output, keep_prob_ph)
# Fully connected layer
W = tf.Variable(tf.truncated_normal([HIDDEN_SIZE * 2, NUM_CLASSES], stddev=0.1), name="W")
b = tf.Variable(tf.constant(0., shape=[NUM_CLASSES]), name="b")
y_hat = tf.nn.xw_plus_b(drop, W, b, name="scores")
# +
# Adam parameters
LEARNING_RATE = 1e-4
EPSILON = 1e-5
BETA1 = 0.9
BETA2 = 0.9
# L2 regularization coefficient
BETA = 0
cross_entropy = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(y_hat, target_ph), name="cross_entropy")
l2_loss = tf.nn.l2_loss(W, name="l2_loss")
loss = cross_entropy + l2_loss * BETA
optimizer = tf.train.AdamOptimizer(learning_rate=LEARNING_RATE, beta1=BETA1, beta2=BETA2,
epsilon=EPSILON).minimize(loss)
# optimizer = tf.train.AdadeltaOptimizer(learning_rate=LEARNING_RATE, rho=0.7).minimize(loss)
# optimizer = tf.train.AdagradOptimizer(learning_rate=LEARNING_RATE, initial_accumulator_value=0.1).minimize(loss)
# optimizer = tf.train.MomentumOptimizer(learning_rate=1e-1, momentum=0.1).minimize(loss)
# optimizer = tf.train.RMSPropOptimizer(learning_rate=LEARNING_RATE, decay=0.9, momentum=0.1).minimize(loss)
accuracy = tf.reduce_mean(tf.cast(tf.equal(tf.argmax(target_ph, 1), tf.argmax(y_hat, 1)), tf.float32))
# +
from sklearn.model_selection import StratifiedKFold, KFold
DROPOUT = 0.5 # Probability of keeping a neuron
BATCH_SIZE = 256
EPOCHS = 100
results = []
results50 = []
skf = KFold(5, shuffle=True, random_state=42)
for train_index, test_index in skf.split(X, y):
X_tr, X_val = X_full[train_index], X_full[test_index]
y_tr, y_val = y_full[train_index], y_full[test_index]
train_batch_generator = batch_generator(X_tr, y_tr, BATCH_SIZE)
loss_tr_l = []
loss_val_l = []
ce_tr_l = [] # Cross-entropy
ce_val_l = []
acc_tr_l = [] # Accuracy
acc_val_l = []
f_macro_tr_l = []
f_macro_val_l = []
f_fair_tr_l = []
f_fair_val_l = []
min_ce = 0.3
with tf.Session() as sess:
sess.run(tf.initialize_all_variables())
sess.run(init_embeddings, feed_dict={embeddings_ph: embeddings})
print "Start learning..."
for epoch in range(EPOCHS):
for i in range(int(X_tr.shape[0] / BATCH_SIZE)):
x_batch, y_batch = train_batch_generator.next()
seq_len_tr = np.array([list(x).index(eos_id) + 1 for x in x_batch])
sess.run(optimizer, feed_dict={batch_ph: x_batch, target_ph: y_batch,
seq_len_ph: seq_len_tr, keep_prob_ph: DROPOUT})
y_pred_tr, ce_tr, loss_tr, acc_tr = sess.run([y_hat, cross_entropy, loss, accuracy],
feed_dict={batch_ph: x_batch, target_ph: y_batch,
seq_len_ph: seq_len_tr, keep_prob_ph: 1.0})
seq_len_val = np.array([list(x).index(eos_id) + 1 for x in X_val])
y_pred_val, ce_val, loss_val, acc_val = sess.run([y_hat, cross_entropy, loss, accuracy],
feed_dict={batch_ph: X_val, target_ph: y_val,
seq_len_ph: seq_len_val, keep_prob_ph: 1.0})
y_pred_tr = np.array([cls2probs(cls) for cls in np.argmax(y_pred_tr, 1) - 1])
y_pred_val = np.array([cls2probs(cls) for cls in np.argmax(y_pred_val, 1) - 1])
f_macro_tr, f_micro_tr = f_macro(y_batch, y_pred_tr), f_micro(y_batch, y_pred_tr)
f_macro_val, f_micro_val = f_macro(y_val, y_pred_val), f_micro(y_val, y_pred_val)
loss_tr_l.append(loss_tr)
loss_val_l.append(loss_val)
ce_tr_l.append(ce_tr)
ce_val_l.append(ce_val)
acc_tr_l.append(acc_tr)
acc_val_l.append(acc_val)
f_macro_tr_l.append(f_macro_tr)
f_macro_val_l.append(f_macro_val)
clear_output(wait=True)
print "epoch: {}".format(epoch)
print "\t Train loss: {:.3f}\t ce: {:.3f}\t acc: {:.3f}\t f_macro: {:.3f}".format(
loss_tr, ce_tr, acc_tr, f_macro_tr)
print "\t Valid loss: {:.3f}\t ce: {:.3f}\t acc: {:.3f}\t f_macro: {:.3f}".format(
loss_val, ce_val, acc_val, f_macro_val)
plt.figure(figsize=(15,10))
plt.plot(ce_tr_l, color='blue', label='ce_tr')
plt.plot(ce_val_l, color='red', label='ce_val')
plt.plot(f_macro_val_l, color='green', label='f_macro_val')
plt.xlim(0, EPOCHS - 1)
plt.ylim(0, 1)
plt.legend()
plt.show()
if epoch == 49:
results50.append([acc_val_l[-1], f_macro_val_l[-1]])
results.append([acc_val_l[-1], f_macro_val_l[-1]])
# if ce_val < min_ce:
# min_ce = ce_val
# saver = tf.train.Saver()
# saver.save(sess, 'model_attention_momentum')
# -
min(ce_val_l), max(acc_val_l), max(f_macro_val_l)
# ---
# ---
# ---
# # Testing
bank_test = pd.read_csv('../data/bank_test.csv', sep='\t', encoding='utf-8')
texts_test, labels_test = bank_test.text.values, bank_test.label.values
# +
from collections import Counter
print Counter(labels)
print Counter(labels_test)
print labels.shape[0]
print labels_test.shape[0]
# + active=""
# %%time
# pool = multiprocessing.Pool()
# X_test = pool.map(text2seq, texts_test)
# cPickle.dump(X_test, open('../data/X_bank_test.pkl', 'wb'))
# -
X_test = cPickle.load(open('../data/X_bank_test.pkl', 'rb'))
X_test = [x + [eos_id]*(length_max - len(x)) for x in X_test]
X_test = np.array(X_test)
y_test = np.array([cls2probs(cls) for cls in labels_test])
# + active=""
# # For experiments with mixing training set with test.
# X = np.vstack((X, X_test))
# y = np.vstack((y, y_test))
# -
with tf.Session() as sess:
new_saver = tf.train.import_meta_graph('model_birnn.meta')
new_saver.restore(sess, tf.train.latest_checkpoint('./'))
seq_len_test = np.array([list(x).index(eos_id) + 1 for x in X_test])
y_pred_test, ce_test, loss_test, acc_test = sess.run([y_hat, cross_entropy, loss, accuracy],
feed_dict={batch_ph: X_test, target_ph: y_test,
seq_len_ph: seq_len_test, keep_prob_ph: 1.0})
seq_len_val = np.array([list(x).index(eos_id) + 1 for x in X_val])
y_pred_val, ce_val, loss_val, acc_val = sess.run([y_hat, cross_entropy, loss, accuracy],
feed_dict={batch_ph: X_val, target_ph: y_val,
seq_len_ph: seq_len_val, keep_prob_ph: 1.0})
y_pred_val = np.array([cls2probs(cls) for cls in np.argmax(y_pred_val, 1) - 1])
f_macro_val, f_micro_val = f_macro(y_val, y_pred_val), f_micro(y_val, y_pred_val)
print f_macro_val, f_micro_val
y_pred_test = np.array([cls2probs(cls) for cls in np.argmax(y_pred_test, 1) - 1])
f_macro_test, f_micro_test = f_macro(y_test, y_pred_test), f_micro(y_test, y_pred_test)
print f_macro_test, f_micro_test
# Varying hidden size.
# bi-rnn, hid 20
print f_macro_val, f_micro_val, acc_val
print f_macro_test, f_micro_test, acc_test
# bi-rnn, hid 50
print f_macro_val, f_micro_val, acc_val
print f_macro_test, f_micro_test, acc_test
# bi-rnn, hid 100
print f_macro_val, f_micro_val, acc_val
print f_macro_test, f_micro_test, acc_test
# bi-rnn, hid 200
print f_macro_val, f_micro_val, acc_val
print f_macro_test, f_micro_test, acc_test
# bi-rnn, hid 300
print f_macro_val, f_micro_val, acc_val
print f_macro_test, f_micro_test, acc_test
# bi-rnn, hid 450
print f_macro_val, f_micro_val, acc_val
print f_macro_test, f_micro_test, acc_test
# Varying dropout parameter.
# bi-rnn, hid 50, keep_prob 0.1
print f_macro_val, f_micro_val, acc_val
print f_macro_test, f_micro_test, acc_test
# bi-rnn, hid 50, keep_prob 0.2
print f_macro_val, f_micro_val, acc_val
print f_macro_test, f_micro_test, acc_test
# bi-rnn, hid 50, keep_prob 0.3
print f_macro_val, f_micro_val, acc_val
print f_macro_test, f_micro_test, acc_test
# bi-rnn, hid 50, keep_prob 0.4
print f_macro_val, f_micro_val, acc_val
print f_macro_test, f_micro_test, acc_test
# bi-rnn, hid 50, keep_prob 0.6
print f_macro_val, f_micro_val, acc_val
print f_macro_test, f_micro_test, acc_test
| code/experiments_banks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# ### Import packages
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import pickle
from functools import reduce
from sklearn.dummy import DummyClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.cross_validation import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import accuracy_score
from sklearn.learning_curve import learning_curve
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.cross_validation import cross_val_score
from sklearn.metrics import precision_score
from sklearn.metrics import f1_score
from sklearn.metrics import recall_score
from sklearn.metrics import roc_auc_score, roc_curve
from sklearn.svm import LinearSVC
from sklearn.metrics import classification_report
from sklearn.model_selection import GridSearchCV
import imblearn.over_sampling
from sklearn.metrics import confusion_matrix
# -
df = pickle.load(open("clean_data.p", 'rb'))
# # Binary outcome - positive or negative, Dogs and Cats
# ### Create train and test data
# +
model_df = df.dropna()
X1 = model_df.drop(['happy_outcome', 'outcome_class'], axis=1)
y1 = model_df.happy_outcome
X_train, X_test, y_train, y_test = train_test_split(X1, y1, test_size=0.30)
# -
model_df.dog.value_counts()
y_train.value_counts()
# ### Basic logistic regression
# +
model_df = df.dropna()
X1 = model_df.drop(['happy_outcome', 'outcome_class'], axis=1)
y1 = model_df.happy_outcome
X_train, X_test, y_train, y_test = train_test_split(X1, y1, test_size=0.30)
lm_log_simple = LogisticRegression()
lm_log_simple.fit(X_train, y_train)
y_pred_log = lm_log_simple.predict(X_test)
print('\nLogistic Regression Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred_log))
print('Precision:', precision_score(y_test, y_pred_log))
print('F1:', f1_score(y_test, y_pred_log))
print('Recall:', recall_score(y_test, y_pred_log))
# +
log_confusion = confusion_matrix(y_test, lm_log_simple.predict(X_test))
plt.figure(dpi=150)
sns.heatmap(log_confusion, cmap=plt.cm.Blues, annot=True, square=True)
plt.xlabel('Predicted outcome')
plt.ylabel('Actual outcome')
plt.savefig('log_confusion_mat.png');
# -
param_grid = {'C': np.linspace(1e-7,1e2,11)}
grid = GridSearchCV(LogisticRegression(), param_grid=param_grid, cv=5, scoring='accuracy',n_jobs=-1)
grid.fit(X1,y1)
# +
import warnings
warnings.filterwarnings('ignore')
pd.DataFrame(grid.cv_results_)
# -
grid.best_estimator_
# +
model_df = df.dropna()
X1 = model_df.drop(['happy_outcome', 'outcome_class'], axis=1)
y1 = model_df.happy_outcome
X_train, X_test, y_train, y_test = train_test_split(X1, y1, test_size=0.30)
lm_log_simple = LogisticRegression(C=50.000000050000004, class_weight=None, dual=False,
fit_intercept=True, intercept_scaling=1, max_iter=100,
multi_class='ovr', n_jobs=1, penalty='l2', random_state=None,
solver='liblinear', tol=0.0001, verbose=0, warm_start=False)
lm_log_simple.fit(X_train, y_train)
y_pred_log = lm_log_simple.predict(X_test)
print('\nLogistic Regression Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred_log))
print('Precision:', precision_score(y_test, y_pred_log))
print('F1:', f1_score(y_test, y_pred_log))
print('Recall:', recall_score(y_test, y_pred_log))
# -
# ### Oversample from negative outcomes
ROS = imblearn.over_sampling.RandomOverSampler(ratio={0:16841*2,1:31480}, random_state=42)
X_train_rs, y_train_rs = ROS.fit_sample(X_train, y_train)
# #### Logistic regression on train data, score on test data: no resampling
# +
lm = LogisticRegression()
lm.fit(X_train, y_train)
y_pred = lm.predict(X_test)
print('\nLogistic Regression Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred))
print('Precision:', precision_score(y_test, y_pred))
print('F1:', f1_score(y_test, y_pred))
print('Recall:', recall_score(y_test, y_pred))
# -
# #### Logistic regression on train data, score on test data: oversampling negative outcome
# +
lm = LogisticRegression()
lm.fit(X_train_rs, y_train_rs)
y_pred = lm.predict(X_test)
print('\nLogistic Regression Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred))
print('Precision:', precision_score(y_test, y_pred))
print('F1:', f1_score(y_test, y_pred))
print('Recall:', recall_score(y_test, y_pred))
# +
lm_svc = LinearSVC()
lm_svc.fit(X_train, y_train)
y_pred_svc = lm_svc.predict(X_test)
print('\nLinear SVM Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred_svc))
print('Precision:', precision_score(y_test, y_pred_svc))
print('F1:', f1_score(y_test, y_pred_svc))
print('Recall:', recall_score(y_test, y_pred_svc))
# +
lm = LinearSVC()
lm.fit(X_train_rs, y_train_rs)
y_pred = lm.predict(X_test)
print('\nLinear SVM Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred))
print('Precision:', precision_score(y_test, y_pred))
print('F1:', f1_score(y_test, y_pred))
print('Recall:', recall_score(y_test, y_pred))
# -
# #### KNN
# +
lm_knn = KNeighborsClassifier(n_neighbors=2)
lm_knn.fit(X_train, y_train)
y_pred = lm.predict(X_test)
print('\nKNN Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred))
print('Precision:', precision_score(y_test, y_pred))
print('F1:', f1_score(y_test, y_pred))
print('Recall:', recall_score(y_test, y_pred))
fpr_knn, tpr_knn, thresholds = roc_curve(y_test, lm_knn.predict_proba(X_test)[:,1])
# -
# #### Random Forest
# +
lm_rf = RandomForestClassifier()
lm_rf.fit(X_train, y_train)
y_pred = lm.predict(X_test)
print('\nRandom Forest Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred))
print('Precision:', precision_score(y_test, y_pred))
print('F1:', f1_score(y_test, y_pred))
print('Recall:', recall_score(y_test, y_pred))
fpr_rf, tpr_rf, thresholds = roc_curve(y_test, lm_rf.predict_proba(X_test)[:,1])
# -
# #### Decision Tree
# +
lm_dt = DecisionTreeClassifier()
lm_dt.fit(X_train, y_train)
y_pred = lm.predict(X_test)
print('\nDecision Tree Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred))
print('Precision:', precision_score(y_test, y_pred))
print('F1:', f1_score(y_test, y_pred))
print('Recall:', recall_score(y_test, y_pred))
fpr_dt, tpr_dt, thresholds = roc_curve(y_test, lm_dt.predict_proba(X_test)[:,1])
# -
# #### Gaussian
# +
lm_g = GaussianNB()
lm_g.fit(X_train, y_train)
y_pred = lm.predict(X_test)
print('\nGaussian Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred))
print('Precision:', precision_score(y_test, y_pred))
print('F1:', f1_score(y_test, y_pred))
print('Recall:', recall_score(y_test, y_pred))
fpr_g, tpr_g, thresholds = roc_curve(y_test, lm_g.predict_proba(X_test)[:,1])
# -
# #### Dummy Classifier
# +
clf = DummyClassifier(strategy='most_frequent',random_state=4444) # what does random state do in DummyClassifier?
clf.fit(X_train, y_train)
y_pred_dummy = clf.predict(X_test)
print('\nDummy Classifier Test Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred_dummy))
print('Precision:', precision_score(y_test, y_pred_dummy))
print('F1:', f1_score(y_test, y_pred_dummy))
print('Recall:', recall_score(y_test, y_pred_dummy))
fpr_clf, tpr_clf, thresholds = roc_curve(y_test, clf.predict_proba(X_test)[:,1])
# +
fpr, tpr, thresholds = roc_curve(y_test, lm_log_simple.predict_proba(X_test)[:,1])
plt.rc('legend',**{'fontsize':16})
plt.figure(figsize=(10,5))
plt.plot(fpr, tpr,lw=2)
plt.plot(fpr_knn, tpr_knn,lw=2)
plt.plot(fpr_rf, tpr_rf,lw=2)
plt.plot(fpr_dt, tpr_dt,lw=2, c='black')
plt.plot(fpr_g, tpr_g,lw=2, c='purple')
plt.plot([0,1],[0,1],c='r',ls='--')
plt.xlim([-0.05,1.05])
plt.ylim([-0.05,1.05])
plt.xticks(size=16)
plt.yticks(size=16)
#plt.xlabel('False positive rate', size=16)
#plt.ylabel('True positive rate', size=16)
plt.legend(['Logistic Regression', 'KNN', 'Random Forest', 'Decision Tree', 'GaussionNB'])
plt.savefig('ROC_curve.png');
# -
# # Multiple Class Outcomes, Dogs and Cats
# ### Create train and test data
# +
model_df = df.dropna()
X1 = model_df.drop(['happy_outcome', 'outcome_class'], axis=1)
y1 = model_df.outcome_class
X_train, X_test, y_train, y_test = train_test_split(X1, y1, test_size=0.30)
# +
lm = LinearSVC()
lm.fit(X_train, y_train)
y_pred = lm.predict(X_test)
print('\nLinear SVM Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred))
print(classification_report(y_test, y_pred))
# +
lm = RandomForestClassifier()
lm.fit(X_train, y_train)
y_pred = lm.predict(X_test)
print('\nRandom Forest Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred))
print(classification_report(y_test, y_pred))
# +
clf = DummyClassifier(strategy='most_frequent',random_state=4444) # what does random state do in DummyClassifier?
clf.fit(X_train, y_train)
y_pred_dummy = clf.predict(X_test)
print('\nDummy Classifier Test Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred_dummy))
print(classification_report(y_test, y_pred))
# +
lm = LogisticRegression()
lm.fit(X_train, y_train)
y_pred = lm.predict(X_test)
print('\nLogistic Regression Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred))
print(classification_report(y_test, y_pred))
# -
# ## Dogs Only: binary outcome
# +
dog_mask = (df.dog == 1)
dog_df = df[dog_mask]
model_df = dog_df.dropna()
X1 = model_df.drop(['happy_outcome', 'outcome_class', 'dog'], axis=1)
y1 = model_df.happy_outcome
X_train, X_test, y_train, y_test = train_test_split(X1, y1, test_size=0.30)
# -
y1.value_counts()
# +
clf = DummyClassifier(strategy='most_frequent',random_state=4444) # what does random state do in DummyClassifier?
clf.fit(X_train, y_train)
y_pred_dummy = clf.predict(X_test)
print('\nDummy Classifier Test Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred_dummy))
print('Precision:', precision_score(y_test, y_pred_dummy))
print('F1:', f1_score(y_test, y_pred_dummy))
print('Recall:', recall_score(y_test, y_pred_dummy))
# +
lm = LogisticRegression()
lm.fit(X_train, y_train)
y_pred = lm.predict(X_test)
print('\nLogistic Regression Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred))
print('Precision:', precision_score(y_test, y_pred))
print('F1:', f1_score(y_test, y_pred))
print('Recall:', recall_score(y_test, y_pred))
# +
lm = RandomForestClassifier()
lm.fit(X_train, y_train)
y_pred = lm.predict(X_test)
print('\nRandom Forest Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred))
print('Precision:', precision_score(y_test, y_pred))
print('F1:', f1_score(y_test, y_pred))
print('Recall:', recall_score(y_test, y_pred))
# -
param_grid = {'C': np.linspace(1e-7,1e2,11)}
grid = GridSearchCV(LinearSVC(), param_grid=param_grid, cv=5, scoring='accuracy',n_jobs=-1)
grid.fit(X1,y1)
# +
import warnings
warnings.filterwarnings('ignore')
pd.DataFrame(grid.cv_results_)
# -
grid.best_estimator_
# +
lm = LinearSVC(C=9.9999999999999995e-08, class_weight=None, dual=True,
fit_intercept=True, intercept_scaling=1, loss='squared_hinge',
max_iter=1000, multi_class='ovr', penalty='l2', random_state=None,
tol=0.0001, verbose=0)
lm.fit(X_train, y_train)
y_pred = lm.predict(X_test)
print('\nLinear SVM Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred))
print('Precision:', precision_score(y_test, y_pred))
print('F1:', f1_score(y_test, y_pred))
print('Recall:', recall_score(y_test, y_pred))
# -
# ### Dogs only - multiple classifiers
# +
dog_mask = (df.dog == 1)
dog_df = df[dog_mask]
model_df = dog_df.dropna()
X1 = model_df.drop(['happy_outcome', 'outcome_class'], axis=1)
y1 = model_df.outcome_class
X_train, X_test, y_train, y_test = train_test_split(X1, y1, test_size=0.30)
# +
lm = LinearSVC()
lm.fit(X_train, y_train)
y_pred = lm.predict(X_test)
print('\nLinear SVM Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred))
print(classification_report(y_test, y_pred))
# +
lm = RandomForestClassifier()
lm.fit(X_train, y_train)
y_pred = lm.predict(X_test)
print('\nRandom Forest Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred))
print(classification_report(y_test, y_pred))
# +
clf = DummyClassifier(strategy='most_frequent',random_state=4444) # what does random state do in DummyClassifier?
clf.fit(X_train, y_train)
y_pred_dummy = clf.predict(X_test)
print('\nDummy Classifier Error Metrics\n')
print('Accuracy:', accuracy_score(y_test, y_pred))
print(classification_report(y_test, y_pred))
# -
| 02_fit_models_and_tune.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:light
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn import datasets
import statsmodels.api as sm
import seaborn as sns
from sklearn.cluster import KMeans
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score, roc_auc_score, roc_curve
from sklearn import tree
from sklearn.model_selection import train_test_split
from sklearn.decomposition import PCA
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import mean_squared_error
from sklearn.ensemble import BaggingClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.neighbors import KNeighborsClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn import svm
data1 = pd.read_csv('[Track1_데이터3] samp_cst_feat.csv',encoding = 'euc-kr')
data2 = pd.read_csv('[Track1_데이터2] samp_train.csv',encoding = 'euc-kr')
data1["MRC_ID_DI"] = data2["MRC_ID_DI"]
data1["MRC_ID_DI"] = data2["MRC_ID_DI"]
categories = ['VAR007','VAR015','VAR018','VAR026','VAR059',
'VAR066','VAR067','VAR070','VAR077','VAR078',
'VAR094','VAR096','VAR097','VAR098','VAR107',
'VAR111','VAR124','VAR127','VAR143','VAR144',
'VAR145','VAR148','VAR165','VAR177','VAR179',
'VAR199','VAR208',"MRC_ID_DI"]
data1[categories] = data1[categories].astype("int64")
data1.groupby(["MRC_ID_DI"]).size()
# #### 온라인 마켓 사용, 미사용으로 분류
data1["MRC_ID_DI"] = data1["MRC_ID_DI"].replace(range(1,11),1)
data1 = data1.drop(['cst_id_di'],axis = 1)
samsung = sm.add_constant(data1, has_constant = 'add')
samsung.head()
feature_columns = list(samsung.columns.difference(["MRC_ID_DI"]))
X = samsung[feature_columns]
y = samsung["MRC_ID_DI"]
print(y)
x_train, x_test, y_train, y_test = train_test_split(X, y,
train_size = 0.7, test_size = 0.3,
random_state = 100) #set_seed
print("x_train.shape = {}, x_test.shape = {}, y_train.shape = {}, y_test.shape = {}".format(x_train.shape, x_test.shape,
y_train.shape, y_test.shape))
model = sm.Logit(y_train, x_train)
results = model.fit(method = "newton")
results.summary()
results.params
np.exp(results.params)
results.aic
y_pred = results.predict(x_test)
y_pred
# +
def PRED(y, threshold):
Y = y.copy()
Y[Y > threshold] = 1
Y[Y <= threshold] = 0
return(Y.astype(int))
Y_pred = PRED(y_pred,0.5)
Y_pred
# -
# ### 오분류표
cfmat = confusion_matrix(y_test, Y_pred)
def acc(cfmat) :
acc = round((cfmat[0,0]+cfmat[1,1])/np.sum(cfmat),3)
return(acc)
acc(cfmat) # accuracy == 0.863
pca = PCA(n_components = 10)
pca.fit(X)
PCscore = pca.transform(X)
PCscore[:,0:5]
eigens_vector = pca.components_.transpose()
eigens_vector
# +
mX = np.matrix(X)
(mX * eigens_vector)[:, 0:5]
# -
print(PCscore)
plt.scatter(PCscore[:, 0], PCscore[:, 1], c = y)
print(PCscore[:,0])
plt.show()
# +
distortions = []
for i in range(1, 11) :
km = KMeans(n_clusters = i, random_state = 102)
km.fit(X)
distortions.append(km.inertia_)
plt.plot(range(1, 11), distortions, marker = 'o')
plt.xlabel("# of clusters")
plt.ylabel("Distortion")
plt.show()
# +
lr_clf = LogisticRegression(max_iter = 10000)
lr_clf.fit(x_train, y_train)
pred_lr = lr_clf.predict(x_test)
print(accuracy_score(y_test, pred_lr))
print(mean_squared_error(y_test, pred_lr))
# -
bag_clf = BaggingClassifier(base_estimator = lr_clf,
n_estimators = 5,
verbose = 1)
lr_clf_bag = bag_clf.fit(x_train, y_train)
pred_lr_bag = lr_clf_bag.predict(x_test)
pred_lr_bag
print(accuracy_score(y_test, pred_lr_bag))
print(mean_squared_error(y_test, pred_lr_bag))
# +
from sklearn.tree import DecisionTreeClassifier
dt_clf = DecisionTreeClassifier()
dt_clf.fit(x_train, y_train)
pred_dt = dt_clf.predict(x_test)
print(accuracy_score(y_test, pred_dt))
print(mean_squared_error(y_test, pred_dt))
# -
rf_clf = RandomForestClassifier(n_estimators = 5,
max_depth = 3,
random_state = 103,
verbose = 1)
rf_clf.fit(x_train, y_train)
pred = rf_clf.predict(x_test)
print(accuracy_score(y_test, pred))
rf_clf = RandomForestClassifier(n_estimators = 500,
max_depth = 3,
random_state = 103,
verbose = 1)
rf_clf.fit(x_train, y_train)
pred = rf_clf.predict(x_test)
print(accuracy_score(y_test, pred))
rf_clf = RandomForestClassifier(n_estimators = 500,
max_depth = 10,
random_state = 103,
verbose = 1)
rf_clf.fit(x_train, y_train)
pred = rf_clf.predict(x_test)
print(accuracy_score(y_test, pred))
rf_clf4 = RandomForestClassifier()
# +
params = { 'n_estimators' : [10, 100, 500, 1000],
'max_depth' : [3, 5, 10, 15]}
rf_clf4 = RandomForestClassifier(random_state = 103,
n_jobs = -1,
verbose = 1)
grid_cv = GridSearchCV(rf_clf4,
param_grid = params,
n_jobs = -1,
verbose = 1)
grid_cv.fit(x_train, y_train)
print('최적 하이퍼 파라미터: ', grid_cv.best_params_)
print('최고 예측 정확도: {:.4f}'.format(grid_cv.best_score_))
# +
test_acc = []
for n in range(1, 11):
clf = KNeighborsClassifier(n_neighbors = n)
clf.fit(x_train, y_train)
y_pred = clf.predict(x_test)
test_acc.append(accuracy_score(y_test, y_pred))
print("k : {}, 정확도 : {}".format(n, accuracy_score(y_test, y_pred)))
# -
test_acc
plt.figure()
plt.plot(range(1, 11), test_acc, label = 'test')
plt.xlabel("n_neighbors")
plt.ylabel("accuracy")
plt.xticks(np.arange(0, 11, step = 1))
plt.legend()
plt.show()
# +
clf_lin = svm.LinearSVC()
clf_lin.fit(x_train, y_train)
y_pred_lin = clf_lin.predict(x_test)
print(confusion_matrix(y_test, y_pred_lin))
print(accuracy_score(y_test, y_pred_lin))
# -
# #### 0(미사용), 1,6,8 Group shaping
group0, group1 = data1[data1["MRC_ID_DI"]==0], data1[data1["MRC_ID_DI"]==1]
group6, group8 = data1[data1["MRC_ID_DI"]==6], data1[data1["MRC_ID_DI"]==8]
print("group0.shape = {}, group1.shape = {}, group6.shape = {}, group8.shape = {}".format(group0.shape, group1.shape,
group6.shape, group8.shape))
group0, group1, group6, group8 = pd.get_dummies(group0), pd.get_dummies(group1), pd.get_dummies(group6), pd.get_dummies(group8)
# #### Dummy 변수 생성, group by shape
print("group0.shape = {}, group1.shape = {}, group6.shape = {}, group8.shape = {}".format(group0.shape, group1.shape,
group6.shape, group8.shape))
group0 = group0.T.drop(["MRC_ID_DI"]).T
group0.index = range(1,len(group0)+1)
group0
group0.corr(method = 'pearson')
for a in categories:
data1[a].value_counts().plot(kind= 'bar')
plt.title(a)
plt.show()
| Track 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (cv)
# language: python
# name: cv
# ---
# <i>Copyright (c) Microsoft Corporation. All rights reserved.</i>
#
# <i>Licensed under the MIT License.</i>
# # Fine-tuning an Action Recognition Model on HMDB51
#
# In this notebook, we demonstrate how to get the SOTA results for the [HMDB51](http://serre-lab.clps.brown.edu/resource/hmdb-a-large-human-motion-database/) human action dataset using the R(2+1)D model.
# ## Initialization
# Import all the functions we need
# +
import sys
sys.path.append("../../")
import time
import os
import numpy as np
from sklearn.metrics import accuracy_score
import torch
import torch.cuda as cuda
import torch.nn as nn
import torchvision
from utils_cv.action_recognition.model import VideoLearner
from utils_cv.action_recognition.dataset import VideoRecord, VideoDataset
from utils_cv.common.gpu import system_info
from utils_cv.common.data import data_path
system_info()
# -
# This shows your machine's GPUs (if it has any) and the computing device `torch/torchvision` is using.
# Ensure edits to libraries are loaded and plotting is shown in the notebook.
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
# Next, set some model runtime parameters.
# Number of consecutive frames used as input to the DNN. Recommended: 32 for high accuracy, 8 for inference speed.
MODEL_INPUT_SIZE = 8
# Batch size. Reduce if running out of memory.
BATCH_SIZE = 8
# Number of training epochs
EPOCHS = 1
# And specify where you downloaded HMDB51.
DATA_ROOT = os.path.join(str(data_path()), "misc/hmdb51")
VIDEO_DIR = os.path.join(DATA_ROOT, "videos")
# For this notebook, we'll try to reproduce the SOTA results on HMDB51 using a specific train/test split known simply as "split1". Download the split and specify where the download location.
TRAIN_SPLIT = os.path.join(DATA_ROOT, "hmdb51_vid_train_split_1.txt")
TEST_SPLIT = os.path.join(DATA_ROOT, "hmdb51_vid_val_split_1.txt")
# ---
#
# # Prepare HMDB51 using the Video Dataset
#
# We can load the downloaded HMBD51 dataset into our `VideoDataset` class by simply passing it the location of the folder. However, since we'll want to specifically test on "split1", we'll also want to pass it train and test reference files.
data = VideoDataset(
VIDEO_DIR,
train_split_file=TRAIN_SPLIT,
test_split_file=TEST_SPLIT,
batch_size=BATCH_SIZE,
video_ext="avi",
)
# Now that the data is loaded, we can take a quick look at some samples.
data.show_batch(rows=2)
# Lets also take a look at the (default) parameters that we're using:
print(f"""\
sample_length: {data.sample_length}
sample_step: {data.sample_step}
temporal_jitter: {data.temporal_jitter}
temporal_jitter_step: {data.temporal_jitter_step}
random_shift: {data.random_shift}
""")
# ## Setup the Video Learner
# Using the paramters below, one can reproduce published results on the HMDB-51 dataset. Video-clip accuracy of close to 80% can be obtained using as input to the DNN 32 consecutive frames (`MODEL_INPUT_SIZE`) and running 48 training epochs. Even using only 8 epochs the accuracy already reaches 75%. Inference and training speed can be reduced by a factor of 3-4 by setting `MODEL_INPUT_SIZE=8` and using higher batch size, however at a significant loss in accuracy.
learner = VideoLearner(data, num_classes=51)
learner.model
learner.dataset.train_ds.dataset.video_records[0].label
# Training configuration
train_cfgs = dict(
epochs=EPOCHS, # Number of training epochs
lr=0.0001, # Learning rate
lr_step_size=np.ceil(
2 / 3 * EPOCHS
), # Reduce learning rate by factor of 10 after this many epochs.
# No need to change these parameters
grad_steps=2, # Accumulate the gradients over multiple forward passes
mixed_prec=False, # Use mixed precision to speed up training
)
learner.fit(train_cfgs)
# ## Model Test
#
# Reported accuracy from "Closer look" paper: 74.5% (clip accuracy of 66.1% on split1 based on VMZ repo)
#
# 1. sample 10 clips uniformly sampled from each test video: [10 x 3 x (8 or 32) x 112 x 112]
# 2. calculate clip-level accuracy: Use 10 batch and infer
# 3. calculate video-level accuracy by averaging them
# 4. average over the clips
# +
# ### Load Saved Weights if needed
#learn.load(body_train_cfgs['model_name'] + "_032", body_train_cfgs['model_dir'])
# -
if cuda.is_available():
device = torch.device("cuda")
num_gpus = cuda.device_count()
# Look for the optimal set of algorithms to use in cudnn. Use this only with fixed-size inputs.
torch.backends.cudnn.benchmark = True
else:
device = torch.device("cpu")
num_gpus = 0
learner.dataset.test_ds.dataset.num_samples = 10
print(
f"{len(learner.dataset.test_ds)} samples of {learner.dataset.test_ds[0][0][0].shape}"
)
# +
# Initialize the model
model = learner.model
model.to(device)
if num_gpus > 1:
model = nn.DataParallel(model)
model.eval()
# Loop over all examples in the test set and compute accuracies
infer_times = []
video_preds = []
video_trues = []
clip_preds = []
clip_trues = []
report_every = 100
with torch.no_grad():
for i in range(
1, len(learner.dataset.test_ds)
): # [::10]: # Skip some examples to speed up accuracy computation
if i % report_every == 0:
print(
"Processsing {} of {} samples..".format(i, len(learner.dataset.test_ds))
)
# Get model inputs
inputs, label = learner.dataset.test_ds[i]
inputs = inputs.to(device, non_blocking=True)
# Run inference
start_time = time.time()
outputs = model(inputs)
outputs = outputs.cpu().numpy()
infer_time = time.time() - start_time
infer_times.append(infer_time)
# Store results
video_preds.append(outputs.sum(axis=0).argmax())
video_trues.append(label)
clip_preds.extend(outputs.argmax(axis=1))
clip_trues.extend([label] * learner.dataset.test_ds.dataset.num_segments)
print(
"Avg. inference time per video (10 clips) =",
np.array(infer_times).mean() * 1000,
"ms",
)
print("Video prediction accuracy =", accuracy_score(video_trues, video_preds))
print("Clip prediction accuracy =", accuracy_score(clip_trues, clip_preds))
| scenarios/action_recognition/02_training_hmdb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 304, "status": "ok", "timestamp": 1636037468369, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="jfX9Mo2IRKkS" outputId="72476e94-f6d5-4f3f-9289-95aba0fefb20"
# ! python -V
# -
# ! pip install transformers
# + executionInfo={"elapsed": 28083, "status": "ok", "timestamp": 1636037505252, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="JplCyr0VKrhP"
import random
import time
import copy
import numpy as np
import pandas as pd
from tqdm import tqdm
import torch
import torch.nn as nn
from torch.cuda.amp import autocast, GradScaler
from transformers import AutoTokenizer, AutoModel, AutoModelForSequenceClassification
from transformers import BertTokenizer, BertModel, AdamW
from transformers import RobertaTokenizer, RobertaModel
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder
from torch.utils.data import Dataset, DataLoader
# -
torch.cuda.is_available()
# + colab={"base_uri": "https://localhost:8080/", "height": 209, "referenced_widgets": ["adef346525b5428cb77ce85b67b7de02", "632a7169fbd74159b11d92ab6cbacfa7", "<KEY>", "<KEY>", "<KEY>", "a825221a71434b82be79352ad867c68a", "<KEY>", "<KEY>", "<KEY>", "769e20765ef64788b8de218a9853c99e", "<KEY>", "<KEY>", "d469fb245e2244848f21d12c9897e0e8", "4718ae66701049229c66c8506b3efe1b", "bddc64019a20420589041d59a8ac1e86", "158f6d85beec4b33bdc422ee9e115808", "d70b870ad108499e8651b57c9e9be34b", "<KEY>", "830958dc67714696a237fc7a051931bc", "2d185bee7972497296fc31ba5c0d54d6", "faaf710ce2cc471c9370811df2f54ba6", "2e6a2c5306764e31bea3a9e37121ff8c", "<KEY>", "d15f42cd6e9d427c99f12b45ba2b04e1", "8e60d43dcd7e4df9a554b7b8ed28c56d", "ebbe8ae171e94413be8fcc944c05b28b", "c280900ff8ba4cc8a2dbf30daad33a4b", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "4e0b4d1769974ba88e49d4fb02e2c4a2", "317c759beaf6494db054eae9162ccfb2", "<KEY>", "<KEY>", "6cd8338fff6a4f469d417911e8b8142b", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "e9d9038ad82440aebdb0fb714add647d", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "58564c36a43e4f3ab99a0e153cb5e480", "8d900d48dde840e0824e409582df0dd2", "5fa6528c3aa5487e87edb8797ebf465e", "d3ce46e504bd4daf9ac3763a7cdc222f", "487d23c4e2b64adda4a945df313d1ed1", "82b0759acdb845ea9859d2f96ce0e5d3", "e87825600783430391ed6af6be32c278", "<KEY>", "<KEY>", "972de0d98911484dad5f2c7b560d973c", "0344eb1638a44fc0bd8be3d2b5aafc99", "df27936694364ed583b70df9a0870a08", "7d83773323bd4e90bc86003c4220a662", "f543e115e78f40f7bc9813a76b5ba862", "<KEY>", "91647e75ea5e4c16a11046fda73ba026", "e2cbc04f6abc4b45b5374f424dd88efa"]} executionInfo={"elapsed": 4421, "status": "ok", "timestamp": 1636037509668, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="o24cy8ORGJjU" outputId="2c57e157-d12e-41c3-9631-cd59f18b134b"
#CHECKPOINT = "sberbank-ai/sbert_large_nlu_ru"
MAX_LEN = 256
TRAIN_BATCH_SIZE = 16
VALID_BATCH_SIZE = 16
EPOCHS = 5
LEARNING_RATE = 2e-05
#tokenizer = AutoTokenizer.from_pretrained(CHECKPOINT)
CHECKPOINT = "sberbank-ai/ruBert-base"
tokenizer = BertTokenizer.from_pretrained(CHECKPOINT)
def get_individual_labels(df):
labels = pd.get_dummies(df.label).rename({
"ДЖОУИ": "Joey", "МОНИКА": "Monica", "РЕЙЧЕЛ": "Rachel", "РОСС": "Ross",
"ФИБИ": "Phoebe", "ЧЕНДЛЕР": "Chandler"
}, axis=1)
return pd.concat([df, labels], axis=1)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 7, "status": "ok", "timestamp": 1636037511385, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="iuShp08vL62l" outputId="885c5ad6-61d9-40c9-ae33-5be1da37a048"
# %cd friends-classification/
# !mkdir models
# ! ls
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 459, "status": "ok", "timestamp": 1636037511841, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="acv7o-iyMo2n" outputId="e1180782-7199-4b5b-ef05-41f2f3d4d114"
df_train = pd.read_csv('train_data.csv').rename({'Category': 'label'}, axis=1)
df_train.other_speaker.fillna('', inplace=True)
df_val = pd.read_csv('val_data.csv')
df_val.other_speaker.fillna('', inplace=True)
df_test = pd.read_csv('test.csv')
df_test.other_speaker.fillna('', inplace=True)
df_train = get_individual_labels(df_train)
df_val = get_individual_labels(df_val)
# Encoding target variable
names_to_cats = LabelEncoder()
df_train['label_code'] = names_to_cats.fit_transform(df_train.label)
df_val['label_code'] = names_to_cats.transform(df_val.label)
df_fb_train = pd.read_csv('train_data_rus_fb_model.csv')
df_full = pd.concat([df_train, df_val])
print(df_train["label"].value_counts()/df_train.shape[0])
print()
print(df_val["label"].value_counts()/df_val.shape[0])
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 459, "status": "ok", "timestamp": 1636037511841, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="acv7o-iyMo2n" outputId="e1180782-7199-4b5b-ef05-41f2f3d4d114"
df_train = pd.read_csv('train_data_eng_fb_model.csv')
df_train.other_speaker.fillna('', inplace=True)
df_val = pd.read_csv('val_data_eng_fb_model.csv')
df_val.other_speaker.fillna('', inplace=True)
df_test = pd.read_csv('test_data_eng_fb_model.csv')
df_test.other_speaker.fillna('', inplace=True)
df_train2 = pd.read_csv('train_data_eng_helsinki_model.csv')
df_train2.other_speaker.fillna('', inplace=True)
df_val2 = pd.read_csv('val_data_eng_helsinki_model.csv')
df_val2.other_speaker.fillna('', inplace=True)
df_test2 = pd.read_csv('test_data_eng_helsinki_model.csv')
df_test2.other_speaker.fillna('', inplace=True)
df_train3 = pd.read_csv('train_data_eng_google_model.csv')
df_train3.other_speaker.fillna('', inplace=True)
df_val3 = pd.read_csv('val_data_eng_google_model.csv')
df_val3.other_speaker.fillna('', inplace=True)
df_test3 = pd.read_csv('test_data_eng_google_model.csv')
df_test3.other_speaker.fillna('', inplace=True)
# Encoding target variable
names_to_cats = LabelEncoder()
df_train['label_code'] = names_to_cats.fit_transform(df_train.label)
df_val['label_code'] = names_to_cats.transform(df_val.label)
df_fb_train = pd.read_csv('train_data_rus_fb_model.csv')
df_full = pd.concat([df_train, df_val])
print(df_train["label"].value_counts()/df_train.shape[0])
print()
print(df_val["label"].value_counts()/df_val.shape[0])
# -
df_train_2X = pd.concat([df_train, df_train2])
df_train_3X = pd.concat([df_train, df_train2, df_train3])
df_val_3X = pd.concat([df_val, df_val2, df_val3])
df_train_3X.drop_duplicates()
# + colab={"base_uri": "https://localhost:8080/", "height": 436} executionInfo={"elapsed": 9, "status": "ok", "timestamp": 1636037511842, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="q87GsjIbOl_e" outputId="382b8f47-59e7-47df-8e3c-c137d3c72bc1"
print(df_train.shape, df_val.shape, df_test.shape)
df_val
# + executionInfo={"elapsed": 264, "status": "ok", "timestamp": 1636041805080, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="6fwxjlLEGzIT"
class FriendsDataset(Dataset):
def __init__(self, dataframe, tokenizer, max_length=512, padding='max_length',
with_labels=True):
self.dataframe = dataframe # pandas dataframe
#Initialize the tokenizer
self.tokenizer = tokenizer
self.padding = padding
self.max_length = max_length
self.with_labels = with_labels
if 'label' not in self.dataframe.columns:
self.with_labels = False
def __len__(self):
return len(self.dataframe)
def __getitem__(self, index):
# Selecting sentence1 and sentence2 at the specified index in the data frame
sent1 = self.dataframe.other_speaker.iloc[index]
sent2 = self.dataframe.friend_response.iloc[index]
# Tokenize the pair of sentences to get token ids, attention masks and token type ids
encoded_pair = self.tokenizer(sent1, sent2,
padding=self.padding, # Pad to max_length
truncation=True, # Truncate to max_length
max_length=self.max_length,
return_tensors='pt') # Return torch.Tensor objects
token_ids = encoded_pair['input_ids'].squeeze(0) # tensor of token ids
attn_masks = encoded_pair['attention_mask'].squeeze(0) # binary tensor with "0" for padded values and "1" for the other values
token_type_ids = encoded_pair['token_type_ids'].squeeze(0) # binary tensor with "0" for the 1st sentence tokens & "1" for the 2nd sentence tokens
if self.with_labels: # True if the dataset has labels
label = self.dataframe.label_code.iloc[index]
label = self.dataframe.Phoebe.iloc[index]
return token_ids, attn_masks, token_type_ids, label
else:
return token_ids, attn_masks, token_type_ids
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 10, "status": "ok", "timestamp": 1636011617187, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="qJWu9ZEGYwui" outputId="b0920bbc-27cd-43ce-ebd6-15ab7ac953c5"
FriendsDataset(df_val, tokenizer)[2]
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 8, "status": "ok", "timestamp": 1636011617187, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="NCOVhXIwY8EQ" outputId="adfd41dd-9c00-431d-d1b1-afdd0d7428d9"
print(tokenizer.batch_encode_plus(df_val.other_speaker[:2].to_list(), padding=True))
# + colab={"base_uri": "https://localhost:8080/", "height": 89} executionInfo={"elapsed": 6, "status": "ok", "timestamp": 1636011621059, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="XeQGWQHShk6g" outputId="4f1b0520-0753-4b2b-c630-9f6d25aa5a68"
encoded_input = tokenizer(df_val.other_speaker[:2].to_list(), padding=True)
print(encoded_input)
tokenizer.decode(encoded_input["input_ids"][0])
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 273, "status": "ok", "timestamp": 1636011626459, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="A3MzcF4WhqPF" outputId="4510c98f-3b98-4733-ef1c-72fc36133c5a"
df_val.other_speaker[:2].to_list()
# + colab={"base_uri": "https://localhost:8080/", "height": 282} executionInfo={"elapsed": 544, "status": "ok", "timestamp": 1636011627375, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="ERDB6JyobclV" outputId="0fe39def-fc13-40f3-fb72-c8dcfd8ecafe"
seq_len = [len(i.split()) for i in df_train.other_speaker.fillna('')]
pd.Series(seq_len).hist(bins = 30)
# + colab={"base_uri": "https://localhost:8080/", "height": 469} executionInfo={"elapsed": 644, "status": "ok", "timestamp": 1636011628318, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="8vdS5P-2az0b" outputId="5e99479f-a945-443c-97a5-ec54b965acef"
dfs_all = pd.concat([df_train, df_val, df_test])
dfs_all['seq_len'] = dfs_all.apply(lambda row:
len(row['other_speaker'] + row['friend_response']), axis=1)
print(len(dfs_all), "number of all dialogs in train, validation and test")
[len(i.split()) for i in dfs_all.other_speaker]
dfs_all['seq_len'].hist(bins = 30)
dfs_all['seq_len'].value_counts().sort_index(ascending=True).cumsum().head(256)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 8, "status": "ok", "timestamp": 1635935619917, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="sbhksA3xcLna" outputId="107b86ad-8bb8-439f-b706-aa823fce7773"
df_train.other_speaker.describe()
# + id="7Mdv07RUakHR"
model = BertModel.from_pretrained(CHECKPOINT)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 17, "status": "ok", "timestamp": 1635926776189, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="iagLgdpUbxzU" outputId="8c46b13a-8ee1-4c58-8cd5-3f505169c983"
model.config
# + colab={"base_uri": "https://localhost:8080/", "height": 69} executionInfo={"elapsed": 15, "status": "ok", "timestamp": 1635926776190, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="eL0HT2PmSj3D" outputId="1f3b9b1d-90ad-40fd-ea98-e7c729eb4ebb"
df_val.other_speaker[2]
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1367, "status": "ok", "timestamp": 1635926777543, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="huooGHOoccF2" outputId="9dcaf530-386d-424e-cadd-ec33483c4ba1"
with torch.no_grad():
output = model(**tokenizer(df_val.other_speaker[2], return_tensors='pt'), )
print(output)
print(output.pooler_output.shape, output.last_hidden_state.shape)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 11, "status": "ok", "timestamp": 1635923888287, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="1BUrLzPQfm3e" outputId="4ffacb2f-5021-4cb1-e8bf-8b470ca8ed2e"
tokenizer(df_val.other_speaker[2], return_tensors='pt')
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 9, "status": "ok", "timestamp": 1635923888288, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="9osfOgxuaO43" outputId="51928964-2fc1-4125-ad46-a18bff032606"
tokenizer.special_tokens_map
# + executionInfo={"elapsed": 6, "status": "ok", "timestamp": 1636037511843, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="jLmUmq46a0D6"
class SentencePairClassifier(nn.Module):
def __init__(self, model=CHECKPOINT, freeze_model=True):
super(SentencePairClassifier, self).__init__()
# Instantiating BERT-based model object
# self.pretrained_layer = AutoModel.from_pretrained(CHECKPOINT)
self.pretrained_layer = BertModel.from_pretrained(CHECKPOINT)
#self.pretrained_layer = RobertaModel.from_pretrained(CHECKPOINT)
hidden_size = self.pretrained_layer.config.hidden_size
# Freeze model layers and only train the classification layer weights
if freeze_model:
for p in self.pretrained_layer.parameters():
p.requires_grad = False
print('All parameters frozen')
# Classification layer
self.cls_layer = nn.Linear(hidden_size, 2)
self.dropout = nn.Dropout(p=0.3)
@autocast() # run in mixed precision
def forward(self, input_ids, attn_masks, token_type_ids):
'''
Inputs:
-input_ids : Tensor containing token ids
-attn_masks : Tensor containing attention masks to be used to focus on non-padded values
-token_type_ids : Tensor containing token type ids to be used to identify sentence1 and sentence2
'''
# Feeding the inputs to the BERT-based model to obtain contextualized representations
output = self.pretrained_layer(input_ids, attn_masks, token_type_ids)
# Feeding to the classifier layer the last layer hidden-state of the [CLS] token further processed by a
# Linear Layer and a Tanh activation. The Linear layer weights were trained from the sentence order prediction (ALBERT) or next sentence prediction (BERT)
# objective during pre-training.
logits = self.cls_layer(self.dropout(output.pooler_output))
return logits
# + executionInfo={"elapsed": 6, "status": "ok", "timestamp": 1636037511843, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="JKiUIjwGiWEv"
def set_seed(seed):
""" Set all seeds to make results reproducible """
torch.manual_seed(seed)
torch.cuda.manual_seed_all(seed)
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
np.random.seed(seed)
random.seed(seed)
#os.environ['PYTHONHASHSEED'] = str(seed)
@autocast()
def evaluate_loss(net, device, criterion, dataloader):
net.eval()
n_correct = 0
mean_loss = 0
count = 0
with torch.no_grad():
for it, (seq, attn_masks, token_type_ids, labels) in enumerate(tqdm(dataloader)):
seq, attn_masks, token_type_ids, labels = \
seq.to(device), attn_masks.to(device), token_type_ids.to(device), labels.to(device)
logits = net(seq, attn_masks, token_type_ids)
mean_loss += criterion(logits.squeeze(-1), labels).item()
count += 1
max_logits, argmax_idx = torch.max(logits.data, dim=1)
n_correct += calcuate_accu(argmax_idx, labels)
del logits
return mean_loss / count, n_correct / len(dataloader.dataset)
# Function to calcuate the accuracy of the model
def calcuate_accu(big_idx, targets):
n_correct = (big_idx==targets).sum().item()
return n_correct
# + executionInfo={"elapsed": 271, "status": "ok", "timestamp": 1636040507768, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="NNBTL2IC8PMJ"
def train_bert(net, criterion, opti, lr, lr_scheduler, train_loader, val_loader, epochs, iters_to_accumulate):
best_loss = np.Inf
best_acc = 0
best_ep = 1
n_iterations = len(train_loader)
batch_size = train_loader.batch_size
print_every = 1000 // batch_size # print the training loss this many times per epoch
print_eval_iters = 10000 // batch_size
scaler = GradScaler()
for ep in range(epochs):
net.train()
curr_loss = 0.0
curr_n_correct = 0.
trailing_loss = 0.
trailing_n_correct = 0.
curr_n_tr_examples = 0
trainling_n_tr_examples = 0
for it, (seq, attn_masks, token_type_ids, labels) in enumerate(tqdm(train_loader)):
# Converting to cuda tensors
seq, attn_masks, token_type_ids, labels = \
seq.to(device), attn_masks.to(device), token_type_ids.to(device), labels.to(device)
# Enables autocasting for the forward pass (model + loss)
with autocast():
# Obtaining the logits from the model
pooled = net(seq, attn_masks, token_type_ids)
# Computing loss
loss = criterion(pooled.squeeze(-1), labels)
#print(loss, type(loss))
loss = loss / iters_to_accumulate # Normalize the loss because it is averaged
# Computing accuracy
#print(pooled.squeeze(-1), labels)
curr_loss += loss.item()
big_val, big_idx = torch.max(pooled.data, dim=1)
n_correct = calcuate_accu(big_idx, labels)
curr_n_correct += n_correct
trailing_loss += loss.item()
trailing_n_correct += n_correct
curr_n_tr_examples += labels.size(0)
trainling_n_tr_examples += labels.size(0)
# Backpropagating the gradients
# Scales loss. Calls backward() on scaled loss to create scaled gradients.
scaler.scale(loss).backward()
if (it + 1) % iters_to_accumulate == 0:
# Optimization step
# scaler.step() first unscales the gradients of the optimizer's assigned params.
# If these gradients do not contain infs or NaNs, opti.step() is then called,
# otherwise, opti.step() is skipped.
scaler.step(opti)
# Updates the scale for next iteration.
scaler.update()
# Adjust the learning rate based on the number of iterations.
lr_scheduler.step()
# Clear gradients
opti.zero_grad()
if (it + 1) % print_every == 0: # Print training loss information
print()
print("Batch {}/{} of epoch {} complete. Loss per last {} samples:: {} "
.format(it+1, n_iterations, ep+1, curr_n_tr_examples, curr_loss / print_every))
accu_step = (curr_n_correct*100) / curr_n_tr_examples
#print(f"Training Loss per 5000 steps: {loss_step}")
print(f"Training Accuracy per last {curr_n_tr_examples} samples: {accu_step}")
curr_loss = 0.0
curr_n_tr_examples = 0
curr_n_correct = 0
if (it + 1) % print_eval_iters == 0 or it == n_iterations - 1:
del pooled, loss
print("Epoch {}, batch {} complete! Training Loss : {}"
.format(ep+1, it+1, trailing_loss / (it+1)))
print("Epoch {}, batch {} complete! Training Accuracy : {}"
.format(ep+1, it+1, trailing_n_correct / trainling_n_tr_examples))
with autocast():
val_loss, val_accuracy = evaluate_loss(net, device, criterion, val_loader) # Compute validation loss
#print()
print("Epoch {}, batch {} complete! Validation Loss : {}".format(ep+1, it+1, val_loss))
print("Epoch {}, batch {} complete! Validation Accuracy : {}".format(ep+1, it+1,val_accuracy))
net.train()
#if val_loss < best_loss:
if val_accuracy > best_acc:
print("Validation loss changed from {} to {}".format(best_loss, val_loss))
print("Best validation accuracy improved from {} to {}".format(best_acc, val_accuracy))
print()
#net_copy = copy.deepcopy(net) # save a copy of the model
best_loss = val_loss
best_acc = val_accuracy
best_ep = ep + 1
# Saving the model
path_to_model='models/{}_lr_{}_val_acc_{}_ep_{}.pt'.format(time.ctime(), lr, round(best_acc, 4), best_ep)
torch.save(net.state_dict(), path_to_model)
print("The model has been saved in {}".format(path_to_model))
torch.cuda.empty_cache()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["3d8a373f8b1a4da3841d55ed5fd62935", "5435b27f9fab4fcd9f0cf0fe1f11db71", "b968066f8f924c309860b111ae0816a4", "98ff0c5dbe8049c588ef631dfbc08ef8", "121075183d2441ddb2e99f849d2e9686", "77b7765f0fe04c8a976230fb2eb01065", "88b743167cd44b318de6a3998f466556", "ef630b6fc27b4750839918ef7ac13c6c", "e55762599f664a0fa5288de998af11c1", "e3aeb80a52bf4ccdb69330b3ebce4355", "7f4f04df6382427288c4e9422e4fbf7d"]} executionInfo={"elapsed": 1839309, "status": "ok", "timestamp": 1636039351577, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="tV_3pDzJ81v9" outputId="5413e47a-47a4-4216-9f14-592c6d734425"
from transformers import get_linear_schedule_with_warmup
from transformers import get_constant_schedule
from sklearn.utils import compute_class_weight
# Set all seeds to make reproducible results
set_seed(1)
# Creating instances of training and validation set
print("Reading training data...")
train_set = FriendsDataset(dataframe=df_train, tokenizer=tokenizer, max_length=MAX_LEN)
#train_set = FriendsDataset(dataframe=df_full, tokenizer=tokenizer, max_length=MAX_LEN)
print("Reading validation data...")
val_set = FriendsDataset(dataframe=df_val, tokenizer=tokenizer, max_length=MAX_LEN)
# Creating instances of training and validation dataloaders
train_loader = DataLoader(train_set, batch_size=TRAIN_BATCH_SIZE, shuffle=True, num_workers=2)
val_loader = DataLoader(val_set, batch_size=VALID_BATCH_SIZE, shuffle=False, num_workers=2)
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
net = SentencePairClassifier(model=CHECKPOINT, freeze_model=False)
print(device)
if torch.cuda.device_count() > 1: # if multiple GPUs
print("Let's use", torch.cuda.device_count(), "GPUs!")
net = nn.DataParallel(net)
net.to(device)
class_weights = compute_class_weight(
'balanced',
classes=np.unique(df_train.Phoebe), y=df_train.Phoebe)
class_weights = torch.tensor(class_weights, dtype=torch.float)
class_weights = class_weights.to(device)
criterion = nn.CrossEntropyLoss(weight=class_weights)
opti = AdamW(net.parameters(), lr=LEARNING_RATE, weight_decay=1e-2)
num_warmup_steps = 0 # The number of steps for the warmup phase.
iters_to_accumulate = 2
num_training_steps = EPOCHS * len(train_loader) # The total number of training steps
t_total = (len(train_loader) // iters_to_accumulate) * EPOCHS # Necessary to take into account Gradient accumulation
#lr_scheduler = get_linear_schedule_with_warmup(optimizer=opti, num_warmup_steps=num_warmup_steps, num_training_steps=t_total)
lr_scheduler = get_constant_schedule(optimizer=opti)
# -
train_bert(net, criterion, opti, LEARNING_RATE, lr_scheduler, train_loader, val_loader, EPOCHS, iters_to_accumulate)
train_bert(net, criterion, opti, LEARNING_RATE, lr_scheduler, train_loader, val_loader, EPOCHS, iters_to_accumulate)
net.load_state_dict(torch.load('models/Mon Nov 15 03:12:52 2021_lr_2e-06_val_acc_0.3009_ep_4.pt'))
train_bert(net, criterion, opti, LEARNING_RATE, lr_scheduler, train_loader, val_loader, EPOCHS, iters_to_accumulate)
# +
LEARNING_RATE = 2e-6
opti = AdamW(net.parameters(), lr=LEARNING_RATE, weight_decay=1e-2)
num_warmup_steps = 0 # The number of steps for the warmup phase.
iters_to_accumulate = 2
num_training_steps = EPOCHS * len(train_loader) # The total number of training steps
t_total = (len(train_loader) // iters_to_accumulate) * EPOCHS # Necessary to take into account Gradient accumulation
#lr_scheduler = get_linear_schedule_with_warmup(optimizer=opti, num_warmup_steps=num_warmup_steps, num_training_steps=t_total)
lr_scheduler = get_constant_schedule(optimizer=opti)
train_bert(net, criterion, opti, LEARNING_RATE, lr_scheduler, train_loader, val_loader, EPOCHS, iters_to_accumulate)
# +
LEARNING_RATE = 2e-6
opti = AdamW(net.parameters(), lr=LEARNING_RATE, weight_decay=1e-2)
num_warmup_steps = 0 # The number of steps for the warmup phase.
iters_to_accumulate = 2
num_training_steps = EPOCHS * len(train_loader) # The total number of training steps
t_total = (len(train_loader) // iters_to_accumulate) * EPOCHS # Necessary to take into account Gradient accumulation
#lr_scheduler = get_linear_schedule_with_warmup(optimizer=opti, num_warmup_steps=num_warmup_steps, num_training_steps=t_total)
lr_scheduler = get_constant_schedule(optimizer=opti)
train_bert(net, criterion, opti, LEARNING_RATE, lr_scheduler, train_loader, val_loader, EPOCHS, iters_to_accumulate)
# -
tokenizer("Hello world")
torch.save(net.state_dict(), 'models/final_model.pt')
test_set = FriendsDataset(dataframe=df_test, tokenizer=tokenizer, max_length=MAX_LEN)
test_loader = DataLoader(test_set, batch_size=VALID_BATCH_SIZE, shuffle=False, num_workers=1)
def predict(net, device, dataloader):
net.eval()
predictions = []
with torch.no_grad():
for it, (seq, attn_masks, token_type_ids) in enumerate(tqdm(dataloader)):
seq, attn_masks, token_type_ids = \
seq.to(device), attn_masks.to(device), token_type_ids.to(device)
logits = net(seq, attn_masks, token_type_ids)
max_logits, argmax_idx = torch.max(logits.data, dim=1)
predictions.extend(argmax_idx.tolist())
del logits
return predictions
preds = predict(net, device, test_loader)
df_test
answers = pd.DataFrame(
names_to_cats.inverse_transform(preds),
index=df_test.Id, columns=["Category"])
answers.to_csv('submission1.csv')
answers
# + id="bxhl6QhJlbfE"
from transformers import get_constant_schedule
opti = AdamW(net.parameters(), lr=LEARNING_RATE, weight_decay=1e-2)
lr_scheduler = get_constant_schedule(optimizer=opti)
train_set = FriendsDataset(dataframe=df_train.iloc[5000:], tokenizer=tokenizer, max_length=MAX_LEN)
val_set = FriendsDataset(dataframe=df_val, tokenizer=tokenizer, max_length=MAX_LEN)
# Creating instances of training and validation dataloaders
train_loader = DataLoader(train_set, batch_size=TRAIN_BATCH_SIZE, shuffle=True, num_workers=2)
val_loader = DataLoader(val_set, batch_size=VALID_BATCH_SIZE, shuffle=False, num_workers=2)
train_bert(net, criterion, opti, LEARNING_RATE, lr_scheduler, train_loader, val_loader, EPOCHS, iters_to_accumulate)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 939, "status": "ok", "timestamp": 1636033344945, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="r6o9XkFVL6qD" outputId="2c4ae139-f975-4f8c-e7b0-88897023e72e"
val_loss, val_accuracy
# + executionInfo={"elapsed": 268, "status": "ok", "timestamp": 1636042247525, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="BnAB6Udp9Z-1"
import gc
gc.collect()
torch.cuda.empty_cache()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 448, "status": "ok", "timestamp": 1636024184045, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="VEInewq0pYTk" outputId="f8944075-b0f8-41a9-8378-f5fc6d1f90fb"
torch.cuda.get_device_properties(0).total_memory / 1e6
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 11448, "status": "ok", "timestamp": 1635932769348, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="zYNBGqTwpoHn" outputId="74b194fd-8d8b-4c9f-a35c-9d95859f2c6d"
# Check that we are using 100% of GPU memory footprint support libraries/code
# from https://github.com/patrickvonplaten/notebooks/blob/master/PyTorch_Reformer.ipynb
# !ln -sf /opt/bin/nvidia-smi /usr/bin/nvidia-smi
# !pip -q install gputil
# !pip -q install psutil
# !pip -q install humanize
import psutil
import humanize
import os
import GPUtil as GPU
GPUs = GPU.getGPUs()
# XXX: only one GPU on Colab and isn’t guaranteed
gpu = GPUs[0]
def printm():
process = psutil.Process(os.getpid())
print("Gen RAM Free: " + humanize.naturalsize( psutil.virtual_memory().available ), " | Proc size: " + humanize.naturalsize( process.memory_info().rss))
print("GPU RAM Free: {0:.0f}MB | Used: {1:.0f}MB | Util {2:3.0f}% | Total {3:.0f}MB".format(gpu.memoryFree, gpu.memoryUsed, gpu.memoryUtil*100, gpu.memoryTotal))
printm()
# #!kill -9 -1
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 452, "status": "ok", "timestamp": 1635932801898, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04422905123959661601"}, "user_tz": -420} id="e9gyehyY6n8C" outputId="0e7574a4-b8d7-44fa-8f06-717912d52d5a"
printm()
# + id="v1kSrbCvMNDf"
| Kaggle Sber's Friends Dialogues Classification Challenge/hypotheses/FriendsPredictNLP2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/blakelobato/DS-Unit-1-Sprint-1-Dealing-With-Data/blob/master/Blake_Lobato_DS_Unit_1_Sprint_Challenge_1_0919.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="NooAiTdnafkz" colab_type="text"
# # Data Science Unit 1 Sprint Challenge 1
#
# ## Loading, cleaning, visualizing, and analyzing data
#
# In this sprint challenge you will look at a dataset of the survival of patients who underwent surgery for breast cancer.
#
# http://archive.ics.uci.edu/ml/datasets/Haberman%27s+Survival
#
# Data Set Information:
# The dataset contains cases from a study that was conducted between 1958 and 1970 at the University of Chicago's Billings Hospital on the survival of patients who had undergone surgery for breast cancer.
#
# Attribute Information:
# 1. Age of patient at time of operation (numerical)
# 2. Patient's year of operation (year - 1900, numerical)
# 3. Number of positive axillary nodes detected (numerical)
# 4. Survival status (class attribute)
# -- 1 = the patient survived 5 years or longer
# -- 2 = the patient died within 5 year
#
# Sprint challenges are evaluated based on satisfactory completion of each part. It is suggested you work through it in order, getting each aspect reasonably working, before trying to deeply explore, iterate, or refine any given step. Once you get to the end, if you want to go back and improve things, go for it!
# + [markdown] id="DUjOBLFAr3A5" colab_type="text"
# ## Part 0 - Revert your version of Pandas right from the start
# I don't want any of you to get stuck because of Pandas bugs, so right from the get-go revert back to version `0.23.4`
# - Run the cell below
# - Then restart your runtime. Go to `Runtime` -> `Restart runtime...` in the top menu (or click the "RESTART RUNTIME" button that shows up in the output of the cell below).
#
# + id="PWq6GbkjsRYQ" colab_type="code" outputId="0bc2277e-0ef5-4f6d-b4fd-bcdaf1337c51" colab={"base_uri": "https://localhost:8080/", "height": 311}
# !pip install pandas==0.23.4
# + [markdown] id="5wch6ksCbJtZ" colab_type="text"
# ## Part 1 - Load and validate the data
#
# - Load the data as a `pandas` data frame.
# - Validate that it has the appropriate number of observations (you can check the raw file, and also read the dataset description from UCI).
# - Validate that you have no missing values.
# - Add informative names to the features.
# - The survival variable is encoded as 1 for surviving >5 years and 2 for not - change this to be 0 for not surviving and 1 for surviving >5 years (0/1 is a more traditional encoding of binary variables)
#
# At the end, print the first five rows of the dataset to demonstrate the above.
# + id="287TpoGKFRVK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 335} outputId="bb22ce16-05bf-48fe-aebf-9e1bf2ab0e18"
# TODO
import pandas as pd
import numpy as np
#df = pd.read_csv('http://archive.ics.uci.edu/ml/machine-learning-databases/haberman/haberman.data') #loads the data as a pandas data frame
col_names = ['Age', 'Year of Op', 'Num + Nodes Detected', 'Survival Class'] #creates column names form data description
df = pd.read_csv('http://archive.ics.uci.edu/ml/machine-learning-databases/haberman/haberman.data',names=col_names) #creates the dataframe with proper headers // informative names to the futures
df.head(10)
# + id="LESTFJkWnIVZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="246c96df-b28d-4a6a-b831-fb898621314e"
df.shape #checks with data description from UCI for total counts and columns
# + id="fohvv1HFnS6N" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 98} outputId="d9582def-392c-4a6d-98fe-9a3eeec7f5d1"
df.isnull().sum() #checks that there are no missing values
# + id="9gnSOjjFoyHH" colab_type="code" colab={}
def change_1_to_0(x): #define a function to change values from 1 to 0 and 2 to 1 in a traditional binary method
if x ==1:
return 0
else:
return 1
# + id="XP0WTFnypBe6" colab_type="code" colab={}
df['Survival Class'] = df['Survival Class'].apply(change_1_to_0) #change to traditional binary
# + id="leFx76aupMmp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 335} outputId="ba8193ac-4fed-4af2-ee91-8e231af9a419"
df.head(10)
# + [markdown] id="G7rLytbrO38L" colab_type="text"
# ## Part 2 - Examine the distribution and relationships of the features
#
# Explore the data - create at least *2* tables (can be summary statistics or crosstabulations) and *2* plots illustrating the nature of the data.
#
# This is open-ended, so to remind - first *complete* this task as a baseline, then go on to the remaining sections, and *then* as time allows revisit and explore further.
#
# Hint - you may need to bin some variables depending on your chosen tables/plots.
# + id="IAkllgCIFVj0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 277} outputId="5635e243-55ce-4c18-e4ff-7158c536e5c1"
import matplotlib.pyplot as plt
# TODO
df.describe() #table describing all the data
# + id="nr-0IcLhukFH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 115} outputId="cdb994ce-560f-438c-e79e-dcecf2731255"
df['Age'].value_counts(dropna=False).head() #shows the age counts in a table
# + id="wxmk1VBpqeMG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="f19f15af-8a58-4f84-9004-99b8307c22e8"
df['Age'].plot.density() #creates a density plot for age distribution
# + id="dtVhZ2S6qHSr" colab_type="code" colab={}
age_bin = pd.cut(df['Age'],5) #seperates the age into five evenly spaced bins
# + id="2FA5z2S9r_He" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 131} outputId="d3007072-4ac5-46b3-980c-23138a2e7d7e"
ct = pd.crosstab(age_bin, df['Survival Class']) #creates a cross tab table for the age groups bin and if they survived past 5 years or not
print(ct)
# + id="qLRMyqfnqSWZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 360} outputId="b6443202-eee8-46e7-c509-2778be05c973"
ct.plot(kind='bar') #creates bar graph on the cross tab data
# + id="vdZISyfdtlcr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 471} outputId="0ca78df7-0017-43ad-b59e-770e74c1d879"
node_bin = pd.cut(df['Num + Nodes Detected'],5) #one more graph and table to visualize the data
ct1 = pd.crosstab(node_bin,df['Survival Class'], normalize='columns') #creates a normalized crosstab
print(ct1)
ct2 = pd.crosstab(node_bin,df['Survival Class'])
ct2.plot(kind='bar')
# + [markdown] id="sDXMio-yiWPT" colab_type="text"
# ## Part 3 - DataFrame Filtering
#
# Use DataFrame filtering to subset the data into two smaller dataframes. You should make one dataframe for individuals who survived >5 years and a second dataframe for individuals who did not.
#
# Create a graph with each of the dataframes (can be the same graph type) to show the differences in Age and Number of Positive Axillary Nodes Detected between the two groups.
# + id="rLjGyJfGiedx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 191} outputId="98262f27-4b42-4b22-9aa1-4a47389686ce"
# TODO
df_1=df #creates another data set as a current copy of the original data frame
df_1.head(5)
# + id="ATwi91ZdvzVx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="0cf7fed5-1e5e-4a6e-84b0-84312c2e3199"
df_1.drop(index=1, columns='Survival Class') #takes all those who passed away in five years and removing their row contents from the new data frame
# + id="X9drHgG2winJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 191} outputId="3c040b78-b9fd-46be-a0fe-607066c151ea"
df_dropzero = df.drop(index=0, columns='Survival Class') #removes all those who lived more than five years in a new data frame called df_dropzero
df_dropzero.head()
# + id="HSBLNLoWw3Mg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 301} outputId="6dc19a47-d652-4d78-cad7-82f2c6a4e8a1"
plt.plot(df_1['Age'], df_1['Num + Nodes Detected'], 'co', df_dropzero['Age'], df_dropzero['Num + Nodes Detected'], 'b+',) #tries to plot both number of nodes detected versus age for the two data frames
# + id="X9DsTl0q-N0l" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 298} outputId="ee9793c8-05eb-4984-9d11-3a8293d4e785"
plt.plot(df_1['Age'], df_1['Num + Nodes Detected'], 'co')
plt.xlabel('Age')
plt.ylabel('+ Nodes Detected')
# + id="TkoHtQlx-awt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 298} outputId="98227a80-a3cb-40d7-9e77-7db383a3b518"
plt.plot(df_dropzero['Age'], df_dropzero['Num + Nodes Detected'], 'k+')
plt.xlabel('Age')
plt.ylabel('+ Nodes Detected')
# + id="TMFAYTuEyLsO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="b600026e-c381-4c4a-b7c4-82b4237249be"
df_1['Num + Nodes Detected'].plot.density() #creates a density plot of the dataframe with all the 1's taken out
# + id="tMgUMh7-yLwi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="caf599c7-44e1-4217-eb64-facea3aa81e7"
df_dropzero['Num + Nodes Detected'].plot.density() ##creates a density plot of the dataframe with all the 0's taken out
# + id="3qAmzN7jyY6I" colab_type="code" colab={}
#df.head(10)
#df_removedones = df.drop(index=1, columns='Survival Class')
#df_removedzeros = df.drop(index=0, columns='Survival Class')
# + id="ElF9rIiKyY9P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="3a2c4f70-2746-4fc9-c70a-ef04942aaa55"
df_removedones.plot.density() #creates a density plot of all headers for removedones DF
# + id="m2-yTY1jy0KY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="ec1df437-9e8b-4beb-c1bd-047a286e1043"
df_removedzeros.plot.density() #creates a density plot of all headers for removedzeros DF
# + id="tAqHbt_izBZB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 357} outputId="1f76627f-1f90-4247-a826-ea3540c3f636"
ct_surv = pd.crosstab(node_bin, df['Survival Class']) #creates cross tab of node bins versus survival class
ct_surv.plot(kind='bar') #Another analysis of the survival classes in a bar graph
# + id="Zqjuyofuz55t" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="b62d2fd9-cbdb-4891-ed51-1e93be6e2a8d"
ct_age = pd.crosstab(age_bin, df['Num + Nodes Detected'], normalize='columns') #creates cross tab for age bins and number of nodes detected then normalizes the data and makes a bar graph
print(ct_age)
plt.plot(df['Age'],df['Num + Nodes Detected'])
ct_age.plot(kind='bar')
# + [markdown] id="ZM8JckA2bgnp" colab_type="text"
# ## Part 4 - Analysis and Interpretation
#
# Now that you've looked at the data, answer the following questions:
#
# - What is at least one feature that looks to have a positive relationship with survival? (As that feature goes up in value rate of survival increases)
# - What is at least one feature that looks to have a negative relationship with survival? (As that feature goes down in value rate of survival increases)
# - How are those two features related with each other, and what might that mean?
#
# Answer with text, but feel free to intersperse example code/results or refer to it from earlier.
# + [markdown] id="6dKITTOVtHWo" colab_type="text"
# Your Text Answer Here
# + id="90OSBGhOjk1b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 530} outputId="85221cc2-0dd3-4430-9c9c-7ea5d25efbc8"
# Any Supplemental Code for Analysis Here
ct_op = pd.crosstab(df['Year of Op'], df['Survival Class'], normalize='columns') #cross tab evaluating survival with year of operation
print(ct_op)
ct_op.plot(kind='bar')
# + id="HssSLCcp2h5W" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 455} outputId="385bbc65-bd59-4db1-ed98-e302a7e79333"
ct_aux = pd.crosstab(node_bin, df['Survival Class'], normalize='columns') #node bin and survival class cross tab
ct_aux.plot(kind='bar')
print(ct_aux)
# + id="Yqa8Xdhl3W1Q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 360} outputId="505306a3-b948-4a86-b416-eebe5c3d6760"
ct_age = pd.crosstab(age_bin, df['Survival Class'], normalize='columns') #age bin and survival class cross tab
ct_age.plot(kind='bar')
# + id="VzZjQdok8yER" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 180} outputId="25155ba2-040c-47be-9c26-bae8a3c09963"
ctnew = pd.crosstab(age_bin, [df['Survival Class'], node_bin])
print(ctnew)
# + [markdown] id="uou5ZF9kznH9" colab_type="text"
#
# 4.1 What is at least one feature that looks to have a positive relationship with survival? (As that feature goes up in value rate of survival increases)
#
#
# The one feature that appears to have a positive relationship with survival rate is difficult to pinpoint using my tables and graphs. Even when I created two data frames that contained the values between those who had surivie over five years and those who didn't I did not see a large difference. I have tried to find an obvious relationship to no avail.
# It appears the younger patients have higher chances of surviving later. However, the year the operation occured does not show clear cut positive trend relating to the rate of survival chances.
# + [markdown] id="WawODJja0rWq" colab_type="text"
# 4.2 What is at least one feature that looks to have a negative relationship with survival? (As that feature goes down in value rate of survival increases)
#
# The one feature that appears to have a negative relationship with survival is the amount of positive auxillary nodes detected. As this value increases there appears to be more people who are unable to make it past five years.
#
# + [markdown] id="tm7knYrf5qs6" colab_type="text"
# 4. 3 How are those two features related with each other, and what might that mean?
#
# The two features are intermingled in several ways. First of all the younger you are the more the chance of survival. These features are related because as number of positve nodes increase the survival rate decreases. The trends in this data were very difficult to see with the naked eye so I highly recommend running regressions and statistics on the data to gain further insight and certainty.
| Blake_Lobato_DS_Unit_1_Sprint_Challenge_1_0919.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
print(sys.executable)
print(sys.version)
print(sys.version_info)
#
# # Sorting
# A frequent task in data science, engineering, etc. is the seemingly mundane task of
# sorting or ordering things. Here we explore a couple of simple sorting algorithms,
# just to show some of the thoughts that go into such a task, then will ultimately resort
# to the internal sorting routines built into Python.
#
# Sorting is frequently necessary when data are being handled; for example in integration
# and differentiation the data are usually presented to the various algorithms in ascending or
# descending order (at least on the x-axis).
# One may have tables of numbers, representing one or more explanatory variables, and one or
# more responses. At times we may need to arrange these tables in an order dictated by one
# or another of these various variables. Alternatively we may want to
# nd the median value
# or upper quartile of such a list { this task requires sorting.
# When sorting, one can also carry along operations to maintain correspondence with other
# lists (for lack of better name lets call this sort-and-carry).
# Tasks that fall under the broad category of sorting are:
# - Sort ; rearrange an array of numbers into numerical order (ascending or descending).
# - Sort and carry along ; rearrange an array of numbers into numerical order while per-
# forming the same rearrangement of one or more additional arrays so that the correspon-
# dence between elements in all arrays is maintained (the sets of arrays are essentially
# a relational database { so that each record (row) maintains the cross-record (
# elds;
# columns) relationship).
# - Index ; given an array, prepare an index table that is a table of pointers that indicates
# which number array element comes
# rst in numerical order, which is second, and so
# on.
# - Rank ; given an array, prepare a rank table that tells the numerical rank of an array
# element.
# The task of sorting `N` elements requires on the order of $K \cdot Nlog2N$ operations. The
# algorithm inventor tries to make $K$ as small as possible (understanding that $K = 0$ is
# practically impossible). Three useful sorting algorithms are:
#
# 1. Straight insertion sort;
# 2. Heapsort sort; and
# 3. Quicksort sort.
#
# The choice of method depends on the size of the list that needs to be sorted. If the list is
# short (perhaps $N < 50$ elements) then straight insertion is fast enough, concise, and simple
# to program. For a long list ($N > 1000$ elements) Quicksort is faster, but achieves the speed
# by use of extra memory. Heapsort is also good for large lists, and is an in-place routine.
# Python lists have a built-in `sort()` method that modifies the list in-place and a `sorted()`
# built-in function that builds a new sorted list from an iterable. So when sorting needs to
# be done, you should use the built-in tools. However, because it is a useful programming
# construct, the three sorting algorithms are presented as Python primitive codes.
# ### Bubble Sort
# The bubble sort is a place to start despite it's relative slowness. It is a pretty reviled
# algorithm (read the Wikipedia entry), but it is the algorithm that a naive programmer
# might cobble together in a hurry, and despite its shortcomings (it's really slow and
# inefficient), it is robust.
#
# Here is a description of the sorting task as described by <NAME> (2016)
# (pg. 65):
#
# Imagine you want to alphabetize your unsorted collection of books. A
# natural approach would be just to scan across the shelf looking for out-
# of-order pairs - Wallace followed by Pynchon, for instance - and
# flipping them around. Put Pynchon ahead of Wallace, then continue your scan,
# looping around to the beginning of the shelf each time you reach the end.
# When you make a complete pass without finding any more out-of-order
# pairs on the entire shelf, then you know the job is done.
# This process is a Bubble Sort, and it lands us in quadratic time. There
# are n books out of order, and each scan through the shelf can move each
# one at most one position. (We spot a tiny problem, make a tiny fix.) So
# in the worst case, where the shelf is perfectly backward, at least one book
# will need to be moved n positions. Thus a maximum of n passes through
# n books, which gives us O(n2) in the worst case. For instance, it
# means that sorting five shelves of books will take not five times as long as
# sorting a single shelf, but twenty-five times as long."
#
# Converting the word description into Python is fairly simple. We will have a vector of n
# numbers (we use a vector because its easy to step through the different positions),
# and we will scan through the vector once (and essentially find the smallest thing),
# and put it into the first position. Then we scan again from the second position and
# find the smallest thing remaining, and put it into the second position, and so on until
# the last scan which should have the remaining largest thing. If we desire a decreasing
# order, simply change the sense of the comparison.
#
# The algorithm defines an array and then sorts by repeated passes through the array.
# The program (outside of the sorting algorithm) is really quite simple.
# * Load contents into an array to be sorted.
# * Echo (print) the array (so we can verify the data are loaded as anticipated).
# * Loads the sorting function (the two loops)
# * Sort the array, put the results back into the array (an in-place sort).
# * Report the results.
#array = [7,11,5,8,9,13,66,99,223]
#array = [7,11,5]
array=[1003 ,3.2 ,55.5 , -0.0001 , -6 ,666.6 ,102]
howMany = len(array)
print("Item Count = : ",howMany)
print("Unsorted List : ", array)
# insertion sort
for irow in range(0, howMany-1) :
for jrow in range(0,(howMany-1-irow)) :
if array[jrow]> array[jrow+1] :
swap = array[jrow]
array[jrow]=array[jrow+1]
array[jrow+1]=swap
else:
continue
#results
print("Sorted List : ", array, end ="")
# In the script we see that the program (near the bottom of the file) assigns the values to the vector named array and
# the initial order of the array is ${1003, 3.2, 55.5,-0.0001,-6, 666.6, 102}$. The smallest
# value in the example is -6 and it appears in the 5-th position, not the 1-st as it
# should.
#
# The first pass through the array will move the largest value, 1003, in sequence to the
# right until it occupies the last position. Repeated passes through the array move the
# remaining largest values to the right until the array is ordered. One can consider the
# values of the array at each scan of the array as a series of transformations (irow-th
# scan) -- in practical cases we don't necessarily care about the intermediate values,
# but here because the size is manageable and we are trying to get our feet wet with
# algorithms, we can look at the values.
# The sequence of results (transformations) after each pass through the array is shown
# in the following list:
# 1. Initial value: [1003; 3,2; 55,5;-0,0001;-6; 666,6; 102].
# 2. First pass: [3,2; 55,5;-0,0001;-6; 666,6; 102; 1003].
# 3. Second pass: [3,2;-0,0001;-6; 55,5; 102; 666,6; 1003].
# 4. Third pass: [-0,0001;-6; 3,2; 55,5; 102; 666,6; 1003].
# 5. Fourth pass: [-6;-0,0001; 3,2; 55,5; 102; 666,6; 1003].
# 6. Fifth pass: [-6;-0,0001; 3,2; 55,5; 102; 666,6; 1003]. Sorted, fast scan.
# 7. Sixth pass: [-6;-0,0001; 3,2; 55,5; 102; 666,6; 1003]. Sorted, fast scan.
# We could probably add additional code to break from the scans when we have a single
# pass with no exchanges (like the last two scans) -- while meaningless in this example, for larger collections of
# things, being able to break out when the sorting is complete is a nice feature.
#
# ### Insertion Sort
# The next type of sorting would be to select one item and locate it either left or right
# of an adjacent item based on its size { like sorting a deck of cards, or perhaps a better
# description { again using the bookshelf analog from Christian and Griffths (2016)
# (pg. 65)
#
# You might take a different tack -- pulling all the books off the shelf
# and putting them back in place one by one. You'd put the ffrst book in
# the middle of the shelf, then take the second and compare it to the first,
# inserting it either to the right or to the left. Picking up the third book,
# you'd run through the books on the shelf from left to right until you found
# the right spot to tuck it in. Repeating this process, gradually all of the
# books would end up sorted on the shelf and you'd be done. Computer
# scientists call this, appropriately enough, Insertion Sort. The good news
# is that it's arguably even more intuitive than Bubble Sort and doesn't
# have quite the bad reputation. The bad news is that it's not actually that
# much faster. You still have to do one insertion for each book. And each
# insertion still involves moving past about half the books on the shelf, on
# average, to find the correct place.
# Although in practice Insertion Sort does run a bit faster than Bubble Sort,
# again we land squarely, if you will, in quadratic time. Sorting anything
# more than a single bookshelf is still an unwieldy prospect."
# Listing 8 is an R implementation of a straight insertion sort. The script is quite
# compact, and I used indentation and extra line spacing to keep track of the scoping
# delimiters. The sort works as follows, take the an element of the array (start with
# 2 and work to the right) and put it into a temporary location (called swap in my
# script). Then compare locations to the left of swap. If smaller, then break from the
# loop, exchange values, otherwise the values are currently ordered. Repeat (starting
# at the next element) , when all elements have been traversed the resulting vector is
# sorted. Here are the transformations for each pass through the outer loop:
#
# #### Straight Insertion
# The straight insertion sort is the algorithm a card player would use to sort cards. Pick out
# the second card and put it into order with respect to the first; then pick the third card
# and insert it into sequence with the first two; continue until the last card is picked out and
# inserted. Once the last card is sequenced, the result is a sorted deck (list).
# Python implementation of such an algorithm is:
#array = [7,11,5,8,9,13,66,99,223]
array = [7,11,5]
howMany = len(array)
print("Item Count = : ",howMany)
print("Unsorted List : ", array, end ="")
# insertion sort
for i in range(1, len(array)): # Traverse through 1 to len(arr)
key = array[i]
# Move elements of arr[0..i-1], that are
# greater than key, to one position ahead
# of their current position
j = i-1
while j >=0 and key < array[j] :
array[j+1] = array[j]
j -= 1
array[j+1] = key
#results
print("Sorted List : ", array, end ="")
# Probably useful to put into a functional structure:
# Function to do insertion sort
def insertionSort(array):
# Traverse through 1 to len(arr)
for i in range(1, len(array)):
key = array[i]
# Move elements of arr[0..i-1], that are
# greater than key, to one position ahead
# of their current position
j = i-1
while j >=0 and key < array[j] :
array[j+1] = array[j]
j -= 1
array[j+1] = key
return(array)
array = [7,11,5,8,9,13,66,99,223]
print("Unsorted List : ", array)
insertionSort(array)
print("Sorted List : ", array, end ="")
| 9-MyJupyterNotebooks/6-Sorting/6-Sorting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow_p36
# language: python
# name: conda_tensorflow_p36
# ---
import boto3
import pandas as pd
from sagemaker import get_execution_role
# +
role = get_execution_role()
bucket='ds6-yelp-reviews'
data_key = 'checkin.json'
data_location = 's3://{}/{}'.format(bucket, data_key)
df = pd.read_json(data_location, lines=True)
# -
df.head()
def visits(text):
return len(text.split(','))
df['no_visits'] = df['date'].apply(lambda x: visits(x))
import seaborn as sns
sns.distplot(df['no_visits'])
df['no_visits'].describe()
df.head()
df['business_id'].describe()
df[df['business_id'] == '70zgsd7ChWnaxAmdDWT0hQ'].values[0]
| exploratory-data-analysis-master/feature engineering/EDA-checkin-info.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sparkify Project - Feature Engineering and Selection
#
# This notebook is based on the work of Sparkify_data_analysis.ipynb. After the data is inspected and different behaviors between the user who churned and who did not are analyzed, further features are cr
# +
# import libraries
import datetime
import numpy as np
import time
import pandas as pd
import pyspark.sql.functions as F
from lightgbm import LGBMClassifier
from sklearn.feature_selection import SelectFromModel, RFE
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from pyspark.ml import Pipeline
from pyspark.ml.classification import RandomForestClassifier, LogisticRegression, GBTClassifier
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
from pyspark.ml.feature import StandardScaler, OneHotEncoderEstimator, StringIndexer, VectorAssembler # PCA, IDF,
from pyspark.ml.tuning import CrossValidator, ParamGridBuilder
from pyspark.sql.functions import col, udf
from pyspark.sql import SparkSession, Window
from pyspark.sql.types import IntegerType, StringType, TimestampType
# -
# create a Spark session
spark = SparkSession.builder \
.master("local") \
.appName("Sparkify") \
.getOrCreate()
# # Create Feature Dataframe
# Using the `mini_sparkify_event_data.json` to create the features relevant for churn prediction. Here we use the information gained during the data exploration and visualization.
events = spark.read.json("data/mini_sparkify_event_data.json")
# The userId contains values with an empty string. These entries need to be removed.
# events where a userId is an empty string are not valid, remove these
valid_events = events.where(col("userId") != "")
print("Number of total events: {}; number of valid events {}".format(events.count(), valid_events.count()))
print("Number of users: {}".format(valid_events.select("userId").distinct().count()))
# udf for transforming timestamp to datetime
get_date = udf(lambda x: datetime.datetime.fromtimestamp(x / 1000.0), TimestampType()) # udf to convert to timestamp/date
# create column with the date when the log entry was done
valid_events = valid_events.withColumn("log_date", get_date(col("ts")))
# udf that defines churn
find_churn = udf(lambda x: 1 if x == "Cancellation Confirmation" else 0, IntegerType())
# add a column "churn" to the dataframe indicating that a cancellation was confirmed
valid_events = valid_events.withColumn("churn", find_churn(col("page")))
# dataframe for the lengths users listen to music
user_length = valid_events.groupBy("userId").agg({"length": "mean"}) \
.withColumnRenamed("avg(length)", "avg_length")
# some values can be set into relation to a certain time period, e.g. the duration the user is active
# hence we need to create a df with all users and their active time period
# first find the first and last log entry for each user and how much log entries exist per user (all actions)
time_df = valid_events.groupBy(["userId"]) \
.agg(F.sum("churn").alias("churned"), F.min("log_date").alias("first_log"),
F.max("log_date").alias("last_log"), F.count("page").alias("log_counts"), F.max("ts").alias("last_ts"))
# +
def get_time_difference(date_1, date_2):
"""Caculates the time difference between to dates in days.
:param date_1: First date
:type: datetime instance
:param date_2: Second date
:type: datetime instance
:return: Difference between the two dates in days
"""
# difference between the dates
delta = date_2 - date_1
# minimum difference is one 1
if delta.days == 0:
return 1
else:
return delta.days
# register the function as a udf
get_time_difference_udf = udf(get_time_difference, IntegerType())
# add the duration as a time difference between first and last log date
time_df = time_df.withColumn("duration", get_time_difference_udf(col("first_log"), col("last_log"))) \
.drop("first_log", "last_log").withColumnRenamed("churned", "label")
# -
# create a dummy dataframe where each action (About, Thumbs Up, ...) from page is a new column with the number
# how often this action appeared in the data for each user
dummy_df = valid_events.select("userId", "page").groupBy("userId").pivot("page") \
.count().drop("Cancel", "Cancellation Confirmation")
# fill null values
dummy_df = dummy_df.na.fill(0)
# last valid user level
user_level = valid_events.orderBy("log_date", ascending=False).groupBy("userId").agg(F.first("level").alias('valid_level'))
# gender of the users
user_gender = valid_events.select(["userId", "gender"]).distinct()
# calculate the total amount of days the user listened to music
songs_per_date = valid_events.withColumn("date", F.to_date(col("log_date"))).where(col("page") == "NextSong") \
.groupBy(["userId", "date"]).agg(F.lit(1).alias("played_music"))
songs_per_day = songs_per_date.groupBy("userId").agg(F.sum("played_music").alias("music_days"))
# join all previous created dataframes
df = time_df.join(dummy_df, on="userId").join(user_level, on="userId") \
.join(user_gender, on="userId").join(user_length, on="userId").join(songs_per_day, on="userId")
# divide the actions by the amount of logs or the overall duration of their registration
def divide_columns_by(df, columns, value, appendix):
"""Generic function for dividing pyspark columns by a certain value.
:param df: Dataframe containing the columns to divide
:type: Pyspark Dataframe
:param columns: Name of the columns to divide
:type: list[str]
:param value: Value the columns are divided with
:type: int or float
:appendix: String that is added to the new columns
:type: str
:return: Pyspark dataframe with new columns containing the division
"""
for name in columns:
new_name = name+"_"+appendix
df = df.withColumn(new_name, col(name) / col(value))
return df
# variables which shall be divided by a certain value like the duration or log count
cols_to_divide = ['music_days', 'About', 'Add Friend', 'Add to Playlist', 'Downgrade', 'Error', 'Help', 'Home',
'Logout', 'NextSong', 'Roll Advert', 'Save Settings', 'Settings', 'Submit Downgrade',
'Submit Upgrade', 'Thumbs Down', 'Thumbs Up', 'Upgrade']
# dataframe with new columns for the values per duration
df_duration = divide_columns_by(df, cols_to_divide, "duration", "per_day")
# final feature dataframe also containing the values per log count
df_features = divide_columns_by(df_duration, cols_to_divide, "log_counts", "per_log")
# +
# calculate the number of sessions per user and the average number of sessions per day
user_sessions = valid_events.groupBy(["userId"]).agg(F.countDistinct("sessionId").alias("number_sessions"))
df_features = df_features.join(user_sessions, on="userId")
df_features = df_features.withColumn("sessions_per_day", col("number_sessions") / col("duration"))
# ratio of days the user actually listened to music
df_features = df_features.withColumn("ratio_music_days", col("music_days") / col("duration"))
# ratio of songs the user pressed the "Thumbs Up" button
df_features = df_features.withColumn("like_ratio", col("Thumbs Up") / col("NextSong"))
# -
# # Feature Selection
# Once the dataframe with all the features exist, the importance of the features is analyzed to find the ones that seem promising for the model. The feature importance is stored in a pandas dataframe and also saved in a csv-file. This csv-file is loaded before the modelling to decide which ones to keep and which ones to drop. For this 5 different features selection algorithms are implemented (see reference https://towardsdatascience.com/the-5-feature-selection-algorithms-every-data-scientist-need-to-know-3a6b566efd2).
#
# Selecting features by:
# * correlation
# * feature importance of random forest classifier (select from model)
# * recursive feature elimination
# * feature importance of logistic regression (select from model)
# * feature importance of light gbm classifier (select from model)
# convert the dataframe to pandas for feature analysis
pd_df = df_features.toPandas()
# maximum numbers of features to consider
num_features = 25
# split into numerical features and response variable
X = pd_df.drop(["label"], axis=1).select_dtypes(include=["int64", "int32", "float64", "datetime64[ns]"])
y = pd_df["label"]
def cor_selector(X, y,num_feats):
"""Selects features based on pearson correlation.
:param X: Matrix with the feature values
:type: Pandas Dataframe
:param y: Response vector
:type: Pandas Series
:num_featues: Number of features to keep
:type: Int
:return:
cor_support: Binary list for feature importance
cor_feature: List of feature names
"""
cor_list = []
feature_name = X.columns.tolist()
# calculate the correlation with y for each feature
for i in X.columns.tolist():
cor = np.corrcoef(X[i], y)[0, 1]
cor_list.append(cor)
# replace NaN with 0
cor_list = [0 if np.isnan(i) else i for i in cor_list]
# feature name
cor_feature = X.iloc[:,np.argsort(np.abs(cor_list))[-num_feats:]].columns.tolist()
# feature selection? 0 for not select, 1 for select
cor_support = [True if i in cor_feature else False for i in feature_name]
return cor_support, cor_feature
# select features on correlation
cor_support, cor_features = cor_selector(X, y, num_features)
def select_from_model(X, y, num_features):
"""Selects features based on random forest classifier.
:param X: Matrix with the feature values
:type: Pandas Dataframe
:param y: Response vector
:type: Pandas Series
:num_featues: Number of features to keep
:type: Int
:return:
embeded_rf_support: Binary list for feature importance
embeded_rf_feature: List of feature names
"""
embeded_rf_selector = SelectFromModel(RandomForestClassifier(n_estimators=100), max_features=num_features)
embeded_rf_selector.fit(X, y)
embeded_rf_support = embeded_rf_selector.get_support()
embeded_rf_feature = X.loc[:,embeded_rf_support].columns.tolist()
print(str(len(embeded_rf_feature)), 'selected features')
return embeded_rf_support, embeded_rf_feature
# select features from random forest model
embeded_rf_support, embeded_rf_feature = select_from_model(X, y, num_features)
def select_by_rfe(X, y, num_features):
"""Selects features based on recursive feature elimination.
:param X: Matrix with the feature values
:type: Pandas Dataframe
:param y: Response vector
:type: Pandas Series
:num_featues: Number of features to keep
:type: Int
:return:
rfe_support: Binary list for feature importance
rfe_feature: List of feature names
"""
X_norm=(X-X.mean())/X.std()
rfe_selector = RFE(estimator=LogisticRegression(), n_features_to_select=num_features, step=10, verbose=5)
rfe_selector.fit(X_norm, y)
rfe_support = rfe_selector.get_support()
rfe_feature = X.loc[:,rfe_support].columns.tolist()
print(str(len(rfe_feature)), 'selected features')
return rfe_support, rfe_feature
# select features from rfe
rfe_support, rfe_feature = select_by_rfe(X, y, num_features)
def select_from_model_lr(X, y, num_features):
"""Selects features based on logistic regression.
:param X: Matrix with the feature values
:type: Pandas Dataframe
:param y: Response vector
:type: Pandas Series
:num_featues: Number of features to keep
:type: Int
:return:
embeded_lr_support: Binary list for feature importance
embeded_lr_feature: List of feature names
"""
X_norm=(X-X.mean())/X.std()
embeded_lr_selector = SelectFromModel(LogisticRegression(penalty="l1"), max_features=num_features)
embeded_lr_selector.fit(X_norm, y)
embeded_lr_support = embeded_lr_selector.get_support()
embeded_lr_feature = X.loc[:,embeded_lr_support].columns.tolist()
print(str(len(embeded_lr_feature)), 'selected features')
return embeded_lr_support, embeded_lr_feature
# select features from logistic
embeded_lr_support, embeded_lr_feature = select_from_model_lr(X, y, num_features)
def select_by_lgbm(X, y, num_features):
"""Selects features based on LGBM Classifier.
:param X: Matrix with the feature values
:type: Pandas Dataframe
:param y: Response vector
:type: Pandas Series
:num_featues: Number of features to keep
:type: Int
:return:
embeded_lgb_support: Binary list for feature importance
embeded_lgb_feature: List of feature names
"""
lgbc=LGBMClassifier(n_estimators=500, learning_rate=0.05, num_leaves=32, colsample_bytree=0.2,
reg_alpha=3, reg_lambda=1, min_split_gain=0.01, min_child_weight=40)
embeded_lgb_selector = SelectFromModel(lgbc, max_features=num_features)
embeded_lgb_selector.fit(X, y)
embeded_lgb_support = embeded_lgb_selector.get_support()
embeded_lgb_feature = X.loc[:,embeded_lgb_support].columns.tolist()
print(str(len(embeded_lgb_feature)), 'selected features')
return embeded_lgb_support, embeded_lgb_feature
# select features from lgbm
embeded_lgb_support, embeded_lgb_feature = select_by_lgbm(X, y, num_features)
# prepare feature importance dataframe
feature_name = X.columns
feature_selection_df = pd.DataFrame({'Feature':feature_name, 'Pearson':cor_support, 'RFE':rfe_support, 'Logistics':embeded_lr_support,
'Random Forest':embeded_rf_support, 'LightGBM':embeded_lgb_support})
# count the selected times for each feature
feature_selection_df['Total'] = np.sum(feature_selection_df, axis=1)
# sort values according to importance
feature_selection_df = feature_selection_df.sort_values(['Total','Feature'] , ascending=False)
# display feature importance
feature_selection_df.reset_index().head(len(feature_name))
# store feature importance dataframe as csv-file
feature_selection_df.to_csv("data/feature_selection_df.csv")
| Sparkify_feature_importance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DS4000 Rigol Waveform Examples
#
# **<NAME>**
#
# **March 2021**
#
# This notebook illustrates shows how to extract signals from a `.wfm` file created by a the Rigol DS4000 scope. It also validates that the process works by comparing with `.csv` and screenshots.
#
# Two different `.wfm` files are examined one for the DS4022 scope and one for the DS4024 scope. The accompanying `.csv` files seem to have t=0 in the zero in the center of the waveform.
#
# *If RigolWFM is not installed, uncomment the following cell (i.e., delete the #) and run (shift-enter)*
# +
# #!pip install RigolWFM
# +
import numpy as np
import matplotlib.pyplot as plt
try:
import RigolWFM.wfm as rigol
except ModuleNotFoundError:
print('RigolWFM not installed. To install, uncomment and run the cell below.')
print('Once installation is successful, rerun this cell again.')
repo = "https://github.com/scottprahl/RigolWFM/raw/master/wfm/"
# -
# A list of Rigol scopes that should have the same file format is:
print(rigol.DS4000_scopes)
# ## DS4022 Waveform
# ### Look at a screen shot
#
# Start with a `.wfm` file from a Rigol DS4022 scope. It should looks something like this
#
# <img src="https://github.com/scottprahl/RigolWFM/raw/master/wfm/DS4022-A.png" width="70%">
# ### Import the `.wfm` data
# +
# raw=true is needed because this is a binary file
name = "DS4022-A.wfm"
wfm_filename = repo + name + "?raw=true"
w = rigol.Wfm.from_url(wfm_filename, '4000')
# -
# ### First a textual description.
description = w.describe()
print(description)
# Now for the actual signal
# +
toff=0.05
ch=w.channels[0]
plt.title("CH%d %.2fV/div %.2fVoff (%s %s)" % (1,ch.volt_per_division, ch.volt_offset, w.basename, w.firmware))
plt.plot(ch.times*1e3,ch.volts, color='blue', label='WFM')
plt.legend(loc='lower right')
plt.xlabel("Time (ms)")
plt.ylabel("Voltage (V)")
plt.show()
# -
# Note that CH3 and CH4 are both effectively zero, just displaced in the display graph above
w.plot()
plt.show()
# ## DS4024 Waveform
#
# ### Start with importing the `.csv` data
# +
filename = "DS4024-A.csv"
csv_filename = repo + filename
csv_data = np.genfromtxt(csv_filename, delimiter=',', skip_header=2).T
# need to do this separately because only the start and increment is given in the csv file
time = csv_data[0] * 2.000000e-06 - 1.400000e-03
# -
# ### Plot the `.csv` data
plt.plot(time*1000,csv_data[1], color='blue')
plt.plot(time*1000,csv_data[2], color='red')
plt.xlabel("time (ms)")
plt.ylabel("Volts (V)")
plt.title("DS4024-A")
plt.show()
# ### Import the `.wfm` data
# +
# raw=true is needed because this is a binary file
name = "DS4024-A.wfm"
wfm_filename = repo + name + "?raw=true"
w = rigol.Wfm.from_url(wfm_filename, '4')
# -
# ### Now describe the `.wfm` data
print(w.describe())
# ### Finally compare the `.wfm` data to the `.csv` data
# +
toff=0.05
ch=w.channels[0]
plt.title("CH%d %.2fV/div %.2fVoff (%s %s)" % (1,ch.volt_per_division, ch.volt_offset, w.basename, w.firmware))
plt.plot(ch.times*1e3,ch.volts, color='blue', label='WFM')
plt.plot(time*1e3+toff,csv_data[1], color='red', label='CSV')
plt.legend(loc='lower right')
plt.xlabel("Time (ms)")
plt.ylabel("Voltage (V)")
plt.xlim(-1.5,2)
plt.show()
# +
toff=0.0565
ch=w.channels[1]
plt.title("CH%d %.2fV/div %.2fVoff (%s %s)" % (2,ch.volt_per_division, ch.volt_offset, w.basename, w.firmware))
plt.plot(ch.times*1e3,ch.volts, color='blue', label='WFM')
plt.plot(time*1e3+toff,csv_data[2], color='red', label='CSV')
plt.legend(loc='lower right')
plt.xlabel("Time (ms)")
plt.ylabel("Voltage (V)")
plt.xlim(-0.5,1.0)
#plt.ylim(-0.02,0.02)
plt.show()
# -
| docs/1-DS4000-Waveforms.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Hyperparameter Tuning Your Own R Algorithm with Your Own Container in Amazon SageMaker
# _**Using Amazon SageMaker's Hyperparameter Tuning with a customer Docker container and R algorithm**_
#
# ---
#
# **Read before running the notebook:**
# - This notebook has been updated to SageMaker v2.0
# - Use Python3 kernel for this notebook.
# - Dockerfile has been updated to use [Amazon ECR Public Gallery](https://docs.aws.amazon.com/AmazonECR/latest/public/public-gallery.html)
#
#
# ## Contents
#
# 1. [Background](#Background)
# 1. [Setup](#Setup)
# 1. [Permissions](#Permissions)
# 1. [Code](#Code)
# 1. [Publish](#Publish)
# 1. [Data](#Data)
# 1. [Tune](#Tune)
# 1. [HPO Analysis](#HPO-Analysis)
# 1. [Host](#Host)
# 1. [Predict](#Predict)
# 1. [(Optional) Clean-up](#(Optional)-Clean-up)
# 1. [Wrap-up](#Wrap-up)
#
# ---
# ## Background
#
# R is a popular open source statistical programming language, with a lengthy history in Data Science and Machine Learning. The breadth of algorithms available as R packages is impressive and fuels a diverse community of users. In this example, we'll combine one of those algorithms ([Multivariate Adaptive Regression Splines](https://en.wikipedia.org/wiki/Multivariate_adaptive_regression_splines)) with SageMaker's hyperparameter tuning capabilities to build a simple model on the well-known [Iris dataset](https://en.wikipedia.org/wiki/Iris_flower_data_set).
#
#
# This notebook will focus mainly on the integration of hyperparameter tuning and a custom algorithm container, as well as hosting the tuned model and making inference using the endpoint. For more examples, please see this [notebook](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/advanced_functionality/r_bring_your_own/r_bring_your_own.ipynb).
#
# ---
# ## Setup
#
# _This notebook was created and tested on an ml.m4.xlarge notebook instance._
#
# Let's start by specifying:
#
# - The S3 bucket and prefix that you want to use for training and model data. This should be within the same region as the notebook instance, training, and hosting.
# - The IAM role arn used to give training and hosting access to your data. See the [documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/using-identity-based-policies.html) for more details on creating these. Note, if a role not associated with the current notebook instance, or more than one role is required for training and/or hosting, please replace `sagemaker.get_execution_role()` with a the appropriate full IAM role arn string(s).
# + isConfigCell=true
import sagemaker
bucket = sagemaker.Session().default_bucket()
prefix = "sagemaker/DEMO-hpo-r-byo"
role = sagemaker.get_execution_role()
# -
# Now we'll import the libraries we'll need for the remainder of the notebook.
import os
import boto3
import sagemaker
import pandas as pd
from sagemaker.tuner import (
IntegerParameter,
CategoricalParameter,
ContinuousParameter,
HyperparameterTuner,
)
# ### Permissions
#
# Running this notebook requires permissions in addition to the normal `SageMakerFullAccess` permissions. This is because we'll be creating a new repository in Amazon ECR. The easiest way to add these permissions is simply to add the managed policy `AmazonEC2ContainerRegistryFullAccess` to the role associated with your notebook instance. There's no need to restart your notebook instance when you do this, the new permissions will be available immediately.
#
# ---
# ## Code
#
# For this example, we'll need 3 supporting code files. We'll provide just a brief overview of what each one does. See the full R bring your own notebook for more details.
#
# - **Fit**: `mars.R` creates functions to train and serve our model.
# - **Serve**: `plumber.R` uses the [plumber](https://www.rplumber.io/) package to create a lightweight HTTP server for processing requests in hosting. Note the specific syntax, and see the plumber help docs for additional detail on more specialized use cases.
# - **Dockerfile**: This specifies the configuration for our docker container. Smaller containers are preferred for Amazon SageMaker as they lead to faster spin up times in training and endpoint creation, so this container is kept minimal. This docker file starts with base R image, installs `plumber` and `mda` libraries and their dependecies, then adds `mars.R` and `plumber.R`, and finally sets `mars.R` to run as the entrypoint when launched.
#
# - **Upodate:** The updated dockerfile leverages public R-Base image from [Amazon Public ECR Gallery](https://aws.amazon.com/about-aws/whats-new/2020/12/announcing-amazon-ecr-public-and-amazon-ecr-public-gallery/) which has been available since December 2020. Feel free to read more about this public gallery and browse for public images at https://gallery.ecr.aws/.
# ### Publish
# Now, to publish this container to ECR, we'll run the comands below.
#
# This command will take several minutes to run the first time.
algorithm_name = "rmars"
# + language="sh"
#
# # The name of our algorithm
# algorithm_name=rmars
#
# #set -e # stop if anything fails
# account=$(aws sts get-caller-identity --query Account --output text)
#
# # Get the region defined in the current configuration (default to us-west-2 if none defined)
# region=$(aws configure get region)
# region=${region:-us-east-1}
#
# fullname="${account}.dkr.ecr.${region}.amazonaws.com/${algorithm_name}:latest"
#
# # If the repository doesn't exist in ECR, create it.
# aws ecr describe-repositories --repository-names "${algorithm_name}" > /dev/null 2>&1
#
# if [ $? -ne 0 ]
# then
# aws ecr create-repository --repository-name "${algorithm_name}" > /dev/null
# fi
#
# # Get the login command from ECR and execute it directly
# $(aws ecr get-login-password --region ${region} --no-include-email)
#
# # Build the docker image locally with the image name and then push it to ECR
# # with the full name.
# docker build -t ${algorithm_name} .
# docker tag ${algorithm_name} ${fullname}
#
# docker push ${fullname}
# -
# ---
# ## Data
# For this illustrative example, we'll simply use `iris`. This a classic, but small, dataset used to test supervised learning algorithms. Typically the goal is to predict one of three flower species based on various measurements of the flowers' attributes. Further detail can be found [here](https://en.wikipedia.org/wiki/Iris_flower_data_set).
#
# Let's split the data to train and test datasets (70% / 30%) and then copy the data to S3 so that SageMaker training can access it.
data = pd.read_csv("iris.csv")
# Train/test split, 70%-30%
train_data = data.sample(frac=0.7, random_state=42)
test_data = data.drop(train_data.index)
test_data.head()
# Write to csv
train_data.to_csv("iris_train.csv", index=False)
test_data.to_csv("iris_test.csv", index=False)
# write to S3
train_file = "iris_train.csv"
boto3.Session().resource("s3").Bucket(bucket).Object(
os.path.join(prefix, "train", train_file)
).upload_file(train_file)
# _Note: Although we could do preliminary data transformations in the notebook, we'll avoid doing so, instead choosing to do those transformations inside the container. This is not typically the best practice for model efficiency, but provides some benefits in terms of flexibility._
# ---
# ## Tune
#
# Now, let's setup the information needed to train a Multivariate Adaptive Regression Splines model on `iris` data. In this case, we'll predict `Sepal.Length` rather than the more typical classification of `Species` in order to show how factors might be included in a model and to limit the use case to regression.
#
# First, we'll get our region and account information so that we can point to the ECR container we just created.
region = boto3.Session().region_name
account = boto3.client("sts").get_caller_identity().get("Account")
# Now we'll create an estimator using the [SageMaker Python SDK](https://github.com/aws/sagemaker-python-sdk). This allows us to specify:
# - The training container image in ECR
# - The IAM role that controls permissions for accessing the S3 data and executing SageMaker functions
# - Number and type of training instances
# - S3 path for model artifacts to be output to
# - Any hyperparameters that we want to have the same value across all training jobs during tuning
# +
estimator = sagemaker.estimator.Estimator(
image_uri="{}.dkr.ecr.{}.amazonaws.com/rmars:latest".format(account, region),
role=role,
instance_count=1,
instance_type="ml.m4.xlarge",
output_path="s3://{}/{}/output".format(bucket, prefix),
sagemaker_session=sagemaker.Session(),
hyperparameters={"degree": 2},
) # Setting constant hyperparameter
# target is by defauld "Sepal.Length". See mars.R where this is set.
# -
# Once we've defined our estimator we can specify the hyperparameters that we'd like to tune and their possible values. We have three different types of hyperparameters.
# - Categorical parameters need to take one value from a discrete set. We define this by passing the list of possible values to `CategoricalParameter(list)`
# - Continuous parameters can take any real number value between the minimum and maximum value, defined by `ContinuousParameter(min, max)`
# - Integer parameters can take any integer value between the minimum and maximum value, defined by `IntegerParameter(min, max)`
#
# *Note, if possible, it's almost always best to specify a value as the least restrictive type. For example, tuning `thresh` as a continuous value between 0.01 and 0.2 is likely to yield a better result than tuning as a categorical parameter with possible values of 0.01, 0.1, 0.15, or 0.2.*
# +
# to set the degree as a varying HP to tune, use: 'degree': IntegerParameter(1, 3) and remove it from the Estimator
hyperparameter_ranges = {
"thresh": ContinuousParameter(0.001, 0.01),
"prune": CategoricalParameter(["TRUE", "FALSE"]),
}
# -
# Next we'll specify the objective metric that we'd like to tune and its definition. This metric is output by a `print` statement in our `mars.R` file. Its critical that the format aligns with the regular expression (Regex) we then specify to extract that metric from the CloudWatch logs of our training job.
objective_metric_name = "mse"
metric_definitions = [{"Name": "mse", "Regex": "mse: ([0-9\\.]+)"}]
# Now, we'll create a `HyperparameterTuner` object, which we pass:
# - The MXNet estimator we created above
# - Our hyperparameter ranges
# - Objective metric name and definition
# - Whether we should maximize or minimize our objective metric (defaults to 'Maximize')
# - Number of training jobs to run in total and how many training jobs should be run simultaneously. More parallel jobs will finish tuning sooner, but may sacrifice accuracy. We recommend you set the parallel jobs value to less than 10% of the total number of training jobs (we'll set it higher just for this example to keep it short).
tuner = HyperparameterTuner(
estimator,
objective_metric_name,
hyperparameter_ranges,
metric_definitions,
objective_type="Minimize",
max_jobs=9,
max_parallel_jobs=3,
)
# And finally, we can start our hyperparameter tuning job by calling `.fit()` and passing in the S3 paths to our train and test datasets.
#
# *Note, typically for hyperparameter tuning, we'd want to specify both a training and validation (or test) dataset and optimize the objective metric from the validation dataset. However, because `iris` is a very small dataset we'll skip the step of splitting into training and validation. In practice, doing this could lead to a model that overfits to our training data and does not generalize well.*
tuner.fit({"train": "s3://{}/{}/train".format(bucket, prefix)}, wait=False)
# Let's just run a quick check of the hyperparameter tuning jobs status to make sure it started successfully and is `InProgress`.
# +
import time
status = boto3.client("sagemaker").describe_hyper_parameter_tuning_job(
HyperParameterTuningJobName=tuner.latest_tuning_job.job_name
)["HyperParameterTuningJobStatus"]
while status != "Completed":
status = boto3.client("sagemaker").describe_hyper_parameter_tuning_job(
HyperParameterTuningJobName=tuner.latest_tuning_job.job_name
)["HyperParameterTuningJobStatus"]
completed = boto3.client("sagemaker").describe_hyper_parameter_tuning_job(
HyperParameterTuningJobName=tuner.latest_tuning_job.job_name
)["TrainingJobStatusCounters"]["Completed"]
prog = boto3.client("sagemaker").describe_hyper_parameter_tuning_job(
HyperParameterTuningJobName=tuner.latest_tuning_job.job_name
)["TrainingJobStatusCounters"]["InProgress"]
print(f"{status}, Completed Jobs: {completed}, In Progress Jobs: {prog}")
time.sleep(30)
# -
# Wait until the HPO job is complete, and then run the following cell:
boto3.client("sagemaker").describe_hyper_parameter_tuning_job(
HyperParameterTuningJobName=tuner.latest_tuning_job.job_name
)["BestTrainingJob"]
# ---
#
# ## HPO Analysis
#
# Now that we've started our hyperparameter tuning job, it will run in the background and we can close this notebook. Once finished, we can use the [HPO Analysis notebook](https://github.com/awslabs/amazon-sagemaker-examples/tree/master/hyperparameter_tuning/analyze_results/HPO_Analyze_TuningJob_Results.ipynb) to determine which set of hyperparameters worked best.
#
# For more detail on Amazon SageMaker's Hyperparameter Tuning, please refer to the AWS documentation.
# ---
# ## Host
#
# Hosting the model we just tuned takes three steps in Amazon SageMaker. First, we define the model we want to host, pointing the service to the model artifact our training job just wrote to S3.
#
# We will use the results of the HPO for this purpose, but using `hyper_parameter_tuning_job` method.
best_training = boto3.client("sagemaker").describe_hyper_parameter_tuning_job(
HyperParameterTuningJobName=tuner.latest_tuning_job.job_name
)["BestTrainingJob"]
# Get the best trainig job and S3 location for the model file
best_model_s3 = boto3.client("sagemaker").describe_training_job(
TrainingJobName=best_training["TrainingJobName"]
)["ModelArtifacts"]["S3ModelArtifacts"]
best_model_s3
# +
import time
r_job = "DEMO-r-byo-" + time.strftime("%Y-%m-%d-%H-%M-%S", time.gmtime())
# +
r_hosting_container = {
"Image": "{}.dkr.ecr.{}.amazonaws.com/{}:latest".format(account, region, algorithm_name),
"ModelDataUrl": best_model_s3,
}
create_model_response = boto3.client("sagemaker").create_model(
ModelName=r_job, ExecutionRoleArn=role, PrimaryContainer=r_hosting_container
)
print(create_model_response["ModelArn"])
# -
# Next, let's create an endpoing configuration, passing in the model we just registered. In this case, we'll only use a few c4.xlarges.
# +
r_endpoint_config = "DEMO-r-byo-config-" + time.strftime("%Y-%m-%d-%H-%M-%S", time.gmtime())
print(r_endpoint_config)
create_endpoint_config_response = boto3.client("sagemaker").create_endpoint_config(
EndpointConfigName=r_endpoint_config,
ProductionVariants=[
{
"InstanceType": "ml.t2.medium",
"InitialInstanceCount": 1,
"ModelName": r_job,
"VariantName": "AllTraffic",
}
],
)
print("Endpoint Config Arn: " + create_endpoint_config_response["EndpointConfigArn"])
# -
# Finally, we'll create the endpoints using our endpoint configuration from the last step.
# +
# %%time
r_endpoint = "DEMO-r-endpoint-" + time.strftime("%Y%m%d%H%M", time.gmtime())
print(r_endpoint)
create_endpoint_response = boto3.client("sagemaker").create_endpoint(
EndpointName=r_endpoint, EndpointConfigName=r_endpoint_config
)
print(create_endpoint_response["EndpointArn"])
resp = boto3.client("sagemaker").describe_endpoint(EndpointName=r_endpoint)
status = resp["EndpointStatus"]
print("Status: " + status)
try:
boto3.client("sagemaker").get_waiter("endpoint_in_service").wait(EndpointName=r_endpoint)
finally:
resp = boto3.client("sagemaker").describe_endpoint(EndpointName=r_endpoint)
status = resp["EndpointStatus"]
print("Arn: " + resp["EndpointArn"])
print("Status: " + status)
if status != "InService":
raise Exception("Endpoint creation did not succeed")
# -
# ---
# ## Predict
# To confirm our endpoints are working properly, let's try to invoke the endpoint.
#
# _Note: The payload we're passing in the request is a CSV string with a header record, followed by multiple new lines. It also contains text columns, which the serving code converts to the set of indicator variables needed for our model predictions. Again, this is not a best practice for highly optimized code, however, it showcases the flexibility of bringing your own algorithm._
# +
import pandas as pd
import json
iris_test = pd.read_csv("iris_test.csv")
runtime = boto3.Session().client("runtime.sagemaker")
# +
# %%time
# there is a limit of max 500 samples at a time for invoking endpoints
payload = iris_test.drop(["Sepal.Length"], axis=1).to_csv(index=False)
response = runtime.invoke_endpoint(EndpointName=r_endpoint, ContentType="text/csv", Body=payload)
result = json.loads(response["Body"].read().decode())
display(result)
# -
# We can see the result is a CSV of predictions for our target variable. Let's compare them to the actuals to see how our model did.
# +
import matplotlib.pyplot as plt
import numpy as np
plt.scatter(iris_test["Sepal.Length"], np.fromstring(result[0], sep=","), alpha=0.4, s=50)
plt.xlabel("Sepal Length(Actual)")
plt.ylabel("Sepal Length(Prediction)")
x = np.linspace(*plt.xlim())
plt.plot(x, x, linestyle="--", color="g", linewidth=1)
plt.xlim(4, 8)
plt.ylim(4, 8)
plt.show()
# -
# ### (Optional) Clean-up
#
# If you're ready to be done with this notebook, please run the cell below. This will remove the hosted endpoint you created and avoid any charges from a stray instance being left on.
boto3.client("sagemaker").delete_endpoint(EndpointName=r_endpoint)
| r_examples/r_byo_r_algo_hpo/tune_r_bring_your_own.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Notebook com o trabalho final da disciplina de Métodos em Inferência Causal do Master em Jornalismo de Dados, Automação e Data Storytelling do Insper, ministrada por <NAME>.
#
# Grupo: <NAME>, <NAME>, <NAME> e <NAME>
#
# Para a análise, selecionamos cidades de Minas Gerais que possuem barragem de rejeito e têm forte influência da mineração, lista das cidades e respectivos códigos do IBGE:
#
# Barão de Cocais: 3105400
# Brumadinho: 3109006
# Catas Altas: 3115359
# Itabira: 3131703
# Nova Lima: 3144805
# Mariana: 3140001
# Ouro Preto: 3146107
# São Gonçalo do Rio Abaixo: 3161908
# ## Preparando as bases
import pandas as pd
import numpy as np
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
rec12 = pd.read_csv('receitas_candidatos_2012_MG.txt', sep=';', encoding='latin_1', error_bad_lines=False)
rec16 = pd.read_csv('receitas_candidatos_2016_MG.txt', sep=';', encoding='latin_1', error_bad_lines=False)
rec20 = pd.read_csv('receitas_candidatos_2020_MG.csv', sep=';', encoding='latin_1', error_bad_lines=False)
rec12 = rec12.rename(columns={'Numero UE':'Sigla da UE','Municipio':'Nome da UE','CPF/CNPJ do doador':'CPFCNPJ do doador'})
rec16 = rec16.rename(columns={'Número partido doador':'Numero partido','CPF/CNPJ do doador':'CPFCNPJ do doador'})
rec20 = rec20.rename(columns={'ANO_ELEICAO':'Ano','SG_UF':'UF','SG_UE':'Sigla da UE',
'NM_UE':'Nome da UE','SQ_CANDIDATO':'Sequencial Candidato','NR_CANDIDATO':'Numero candidato',
'NM_CANDIDATO':'Nome candidato','DS_CARGO':'Cargo','NR_CPF_CANDIDATO':'CPF do candidato',
'SG_PARTIDO':'Sigla Partido','NR_PARTIDO':'Numero partido','NM_DOADOR':'Nome do doador',
'NM_DOADOR_RFB':'Nome do doador (Receita Federal)','NR_CPF_CNPJ_DOADOR':'CPFCNPJ do doador',
'DS_CNAE_DOADOR':'Setor econômico do doador','VR_RECEITA':'Valor receita','DS_ORIGEM_RECEITA':'Tipo receita',
'DS_RECEITA':'Descricao da receita','DS_FONTE_RECEITA':'Fonte recurso','DS_ESPECIE_RECEITA':'Especie recurso'})
rec12 = rec12.reindex(columns=['Ano','UF','Sigla da UE','Nome da UE','Sequencial Candidato',
'Numero candidato','Nome candidato', 'Cargo', 'CPF do candidato',
'Sigla Partido','Numero partido','Nome do doador', 'Nome do doador (Receita Federal)',
'CPFCNPJ do doador', 'Setor econômico do doador',
'Valor receita', 'Tipo receita', 'Descricao da receita', 'Fonte recurso', 'Especie recurso'])
rec16 = rec16.reindex(columns=['Ano','UF','Sigla da UE','Nome da UE','Sequencial Candidato',
'Numero candidato','Nome candidato', 'Cargo', 'CPF do candidato',
'Sigla Partido','Numero partido','Nome do doador', 'Nome do doador (Receita Federal)',
'CPFCNPJ do doador', 'Setor econômico do doador',
'Valor receita', 'Tipo receita', 'Descricao da receita', 'Fonte recurso', 'Especie recurso'])
rec20 = rec20.reindex(columns=['Ano','UF','Sigla da UE','Nome da UE','Sequencial Candidato',
'Numero candidato','Nome candidato', 'Cargo', 'CPF do candidato',
'Sigla Partido','Numero partido','Nome do doador', 'Nome do doador (Receita Federal)',
'CPFCNPJ do doador', 'Setor econômico do doador',
'Valor receita', 'Tipo receita', 'Descricao da receita', 'Fonte recurso', 'Especie recurso'])
rec16['Ano'] = '2016'
rec12['Ano'] = '2012'
rec12 = rec12.query('`Sigla da UE` == 52370 | `Sigla da UE` == 49212 | `Sigla da UE` == 47996 | `Sigla da UE` == 48950 | `Sigla da UE` == 46337 | `Sigla da UE` == 41840 | `Sigla da UE` == 41793 | `Sigla da UE` == 41076')
rec16 = rec16.query('`Sigla da UE` == 52370 | `Sigla da UE` == 49212 | `Sigla da UE` == 47996 | `Sigla da UE` == 48950 | `Sigla da UE` == 46337 | `Sigla da UE` == 41840 | `Sigla da UE` == 41793 | `Sigla da UE` == 41076')
rec20 = rec20.query('`Sigla da UE` == 52370 | `Sigla da UE` == 49212 | `Sigla da UE` == 47996 | `Sigla da UE` == 48950 | `Sigla da UE` == 46337 | `Sigla da UE` == 41840 | `Sigla da UE` == 41793 | `Sigla da UE` == 41076')
rec1 = rec12.append(rec16)
rec = rec1.append(rec20)
rec["Ano"] = rec["Ano"].astype("int64")
# ## Acrescentando dados
# +
#Inserindo dados para o campo 'População'
rec['População'] = '1000'
conditions = [
(rec['Nome da UE'] == 'NOVA LIMA'),
(rec['Nome da UE'] == 'ITABIRA'),
(rec['Nome da UE'] == 'OURO PRETO'),
(rec['Nome da UE'] == 'BRUMADINHO'),
(rec['Nome da UE'] == 'MARIANA'),
(rec['Nome da UE'] == 'BARÃO DE COCAIS'),
(rec['Nome da UE'] == 'SÃO GONÇALO DO RIO ABAIXO'),
(rec['Nome da UE'] == 'CATAS ALTAS'),
]
values = ['97378','120904','73994','39520','60142','32319','10818','5376']
rec['População'] = np.select(conditions, values)
rec['População'] = rec["População"].astype("float64")
# +
#Inserindo dados para o campo 'IDH'
rec['IDH'] = '0.1'
conditions = [
(rec['Nome da UE'] == 'NOVA LIMA'),
(rec['Nome da UE'] == 'ITABIRA'),
(rec['Nome da UE'] == 'OURO PRETO'),
(rec['Nome da UE'] == 'BRUMADINHO'),
(rec['Nome da UE'] == 'MARIANA'),
(rec['Nome da UE'] == 'BARÃO DE COCAIS'),
(rec['Nome da UE'] == 'SÃO GONÇALO DO RIO ABAIXO'),
(rec['Nome da UE'] == 'CATAS ALTAS'),
]
values = ['0.813','0.756','0.741','0.747','0.742','0.722','0.667','0.684']
rec['IDH'] = np.select(conditions, values)
rec['IDH'] = rec["IDH"].astype("float64")
# +
#Inserindo dados para o campo 'PIB per capita'
rec['PIB per capita'] = '0.1'
conditions = [
(rec['Nome da UE'] == 'NOVA LIMA'),
(rec['Nome da UE'] == 'ITABIRA'),
(rec['Nome da UE'] == 'OURO PRETO'),
(rec['Nome da UE'] == 'BRUMADINHO'),
(rec['Nome da UE'] == 'MARIANA'),
(rec['Nome da UE'] == 'BARÃO DE COCAIS'),
(rec['Nome da UE'] == 'SÃO GONÇALO DO RIO ABAIXO'),
(rec['Nome da UE'] == 'CATAS ALTAS'),
]
values = ['99000','56000','53000','40000','53000','20000','180000','48000']
rec['PIB per capita'] = np.select(conditions, values)
rec['PIB per capita'] = rec["PIB per capita"].astype("float64")
# +
#Transformando 'Valor receita' em float
rec["Valor receita"] = rec["Valor receita"].apply(lambda x: str(x).replace(",", "."))
rec["Valor receita"] = rec["Valor receita"].astype("float64")
#Renomeando partidos que mudaram de nome
rec["Sigla Partido"] = rec["Sigla Partido"].apply(lambda x: str(x).replace("PMDB", "MDB"))
rec["Sigla Partido"] = rec["Sigla Partido"].apply(lambda x: str(x).replace("PRB", "REPUBLICANOS"))
rec["Sigla Partido"] = rec["Sigla Partido"].apply(lambda x: str(x).replace("PPS", "CIDADANIA"))
rec["Sigla Partido"] = rec["Sigla Partido"].apply(lambda x: str(x).replace("PR", "PL"))
rec["Sigla Partido"] = rec["Sigla Partido"].apply(lambda x: str(x).replace("PT do B", "AVANTE"))
rec["Sigla Partido"] = rec["Sigla Partido"].apply(lambda x: str(x).replace("PTN", "PODE"))
rec["Sigla Partido"] = rec["Sigla Partido"].apply(lambda x: str(x).replace("PSDC", "DC"))
rec["Sigla Partido"] = rec["Sigla Partido"].apply(lambda x: str(x).replace("PEN", "PATRIOTA"))
rec["Sigla Partido"] = rec["Sigla Partido"].apply(lambda x: str(x).replace("SD", "SOLIDARIEDADE"))
rec["Sigla Partido"] = rec["Sigla Partido"].apply(lambda x: str(x).replace("PSOLIDARIEDADE", "PSD"))
rec["Sigla Partido"] = rec["Sigla Partido"].apply(lambda x: str(x).replace("PSOLIDARIEDADEB", "PSDB"))
rec["Sigla Partido"] = rec["Sigla Partido"].apply(lambda x: str(x).replace("PSOLIDARIEDADE", "PSD"))
# +
#Inserindo 'Numero Partido' nos campos NaN
conditions = [
(rec['Sigla Partido'] == 'PSDB'),
(rec['Sigla Partido'] == 'PODE'),
(rec['Sigla Partido'] == 'PPL'),
(rec['Sigla Partido'] == 'PC do B'),
(rec['Sigla Partido'] == 'CIDADANIA'),
(rec['Sigla Partido'] == 'PSC'),
(rec['Sigla Partido'] == 'PDT'),
(rec['Sigla Partido'] == 'PLOS'),
(rec['Sigla Partido'] == 'PT'),
(rec['Sigla Partido'] == 'REPUBLICANOS'),
(rec['Sigla Partido'] == 'AVANTE'),
(rec['Sigla Partido'] == 'SOLIDARIEDADE'),
(rec['Sigla Partido'] == 'MDB'),
(rec['Sigla Partido'] == 'PTC'),
(rec['Sigla Partido'] == 'PLTB'),
(rec['Sigla Partido'] == 'DC'),
(rec['Sigla Partido'] == 'PV'),
(rec['Sigla Partido'] == 'PL'),
(rec['Sigla Partido'] == 'PSD'),
(rec['Sigla Partido'] == 'REDE'),
(rec['Sigla Partido'] == 'PSB'),
(rec['Sigla Partido'] == 'PHS'),
(rec['Sigla Partido'] == 'PP'),
(rec['Sigla Partido'] == 'DEM'),
(rec['Sigla Partido'] == 'PTB'),
(rec['Sigla Partido'] == 'PMN'),
(rec['Sigla Partido'] == 'PATRIOTA'),
(rec['Sigla Partido'] == 'PSTU'),
(rec['Sigla Partido'] == 'PSL'),
(rec['Sigla Partido'] == 'PLP'),
(rec['Sigla Partido'] == 'PSOL'),
(rec['Sigla Partido'] == 'UP'),
(rec['Sigla Partido'] == 'PMB'),
]
values = ['45.', '19.', '54.', '65.', '23.', '20.', '12.', '90.', '13.', '10.', '70.', '77.',
'15.', '36.', '28.', '27.', '43.', '22.', '55.', '18.', '40.', '31.', '11.', '25.', '14.',
'33.', '51.', '16.', '17.', '44.', '50.', '80.', '35.']
rec['Numero partido'] = np.select(conditions, values)
# -
# # Conhecendo as cidades
#Quantos partidos disputando as eleições em cada ano?
#Índices gerais da cidade para dar contexto
#Cidades estão em ordem alfabética
rec.groupby(
['Nome da UE','População','IDH','PIB per capita','Ano']).agg(
{'Sigla Partido':['nunique']})
#Quantos candidatos a prefeito em cada eleição?
#Estão em ordem alfabética
rec.loc[rec['Cargo'] == 'Prefeito'].groupby(
['Nome da UE','Ano']).agg(
{'Nome candidato':['nunique']})
#Quantos candidatos a vereador em cada eleição?
rec.loc[rec['Cargo'] == 'Vereador'].groupby(
['Nome da UE','Ano']).agg(
{'Nome candidato':['nunique']}).sort_values(by=['Nome da UE','Ano'],ascending=True)
rec12['Tipo receita'].unique()
rec.to_csv("receitas.csv", sep=";")
# # Perguntas
#Qual eleição tem o maior somatório de doações de pessoas físicas ou jurídicas?
#Quantas foram naquele ano? Qual a média de doação no ano entre todas as cidades?
rec.loc[(rec['Tipo receita']=='Recursos de pessoas físicas') | (
rec['Tipo receita']=='Recursos de pessoas jurídicas')].groupby(
['Ano'])['Valor receita'].agg(['sum','count','mean']).sort_values(
by=['sum'],ascending=False)
#Qual cidade tem o maior somatório de doações nas três eleições?
#Qual a média de doação na cidade de pessoas físicas ou jurídicas?
rec.loc[(rec['Tipo receita']=='Recursos de pessoas físicas') | (
rec['Tipo receita']=='Recursos de pessoas jurídicas')].groupby(
['Nome da UE','PIB per capita', 'População', 'IDH'])['Valor receita'].agg(
['sum','mean']).sort_values(by=['sum'],ascending=False)
#Quais cidades tem as maiores médias de doações?
rec.loc[(rec['Tipo receita']=='Recursos de pessoas físicas') | (
rec['Tipo receita']=='Recursos de pessoas jurídicas')].groupby(
['Nome da UE','PIB per capita', 'População', 'IDH'])['Valor receita'].agg(
['mean']).sort_values(by=['mean'],ascending=False)
#Qual cidade tem o maior média de doações em cada ano?
rec.loc[(rec['Tipo receita']=='Recursos de pessoas físicas') | (
rec['Tipo receita']=='Recursos de pessoas jurídicas')].groupby(
['Ano','Nome da UE'])['Valor receita'].agg(
['mean']).sort_values(by=['Ano','mean'],ascending=False)
#Qual é a maior doação para cada cidade em cada eleição?
rec.loc[(rec['Tipo receita']=='Recursos de pessoas físicas') | (
rec['Tipo receita']=='Recursos de pessoas jurídicas')].groupby(
['Nome da UE','PIB per capita','Ano'])['Valor receita'].agg(
['max','mean']).sort_values(by=['Nome da UE'],ascending=False)
#Ranking das 10 maiores doações pessoas jurídicas (2012), em que cidade ocorreu e o nome do Doador
df = rec.loc[rec['Tipo receita']=='Recursos de pessoas jurídicas'].groupby(
['Ano','Nome da UE','PIB per capita','Nome do doador'])['Valor receita'].agg(
['max']).sort_values(by=['max'],ascending=False)
df.to_csv("maiores doações.csv", sep=";")
#Ranking das 10 maiores doações pessoas físicas (2016/20), em que cidade ocorreu e o nome do Doador
rec.loc[rec['Tipo receita']=='Recursos de pessoas físicas'].groupby(
['Ano','Nome da UE','PIB per capita','Nome do doador'])['Valor receita'].agg(
['max']).sort_values(by=['max'],ascending=False).head(10)
#Em que ano as cidades recolheram o maior total de doações?
rec.loc[(rec['Tipo receita']=='Recursos de pessoas físicas') | (
rec['Tipo receita']=='Recursos de pessoas jurídicas')].groupby(
['Nome da UE','Ano'])['Valor receita'].agg(
['count','sum','mean','median','min','max']).sort_values(
by=['Nome da UE','sum'],ascending=False)
#Quais partidos receberam mais doações e em que anos?
rec.loc[(rec['Tipo receita']=='Recursos de pessoas físicas') | (
rec['Tipo receita']=='Recursos de pessoas jurídicas')].groupby(
['Sigla Partido','Ano'])['Valor receita'].agg(
['count','sum','mean','median','min','max']).sort_values(
by=['sum'],ascending=False)
#Qual partido recebeu mais doações (sum) e em qual cidade? Ranking com os 20 primeiros.
rec.loc[(rec['Tipo receita']=='Recursos de pessoas físicas') | (
rec['Tipo receita']=='Recursos de pessoas jurídicas')].groupby(
['Sigla Partido','Nome da UE'])['Valor receita'].agg(
['count','sum','mean','median']).sort_values(
by=['sum'],ascending=False).head(20)
#Ranking com as 30 pessoas físicas que mais doaram em todas as cidades (sum) e em que ano foram as doações
rec.loc[rec['Tipo receita'] == 'Recursos de pessoas físicas'].groupby(
['Nome do doador','CPFCNPJ do doador','Nome da UE','Ano','PIB per capita'])['Valor receita'].agg(
['sum']).sort_values(
by=['sum'],ascending=False).head(30)
#Top 20 de candidatos com as doações unitárias mais altas (sort max), o cargo e em que ano foi a doação
rec.loc[(rec['Tipo receita']=='Recursos de pessoas físicas') | (
rec['Tipo receita']=='Recursos de pessoas jurídicas')].groupby(
['Nome candidato','Sigla Partido','Cargo','Nome da UE','Ano'])['Valor receita'].agg(
['max','count','sum']).sort_values(by=['max'],ascending=False).head(20)
#Top 10 de Candidatos que receberam mais dinheiro em doações (sort sum) e em que ano
rec.loc[(rec['Tipo receita']=='Recursos de pessoas físicas') | (
rec['Tipo receita']=='Recursos de pessoas jurídicas')].groupby(
["Nome candidato","Sigla Partido","Cargo",'Nome da UE','Ano'])['Valor receita'].agg(
['count','sum','mean','median','min','max']).sort_values(by=['sum'],ascending=False).head(10)
# +
#Agregando doadores
pessoasfisicasejuridicas = rec.loc[(rec['Tipo receita']=='Recursos de pessoas físicas') | (
rec['Tipo receita']=='Recursos de pessoas jurídicas')]
rec_count = pessoasfisicasejuridicas.groupby(['Nome do doador (Receita Federal)']).agg(
{'Valor receita':['sum']}).sort_values(('Valor receita', 'sum'), ascending=False)
#Filtrando quem fez um somatório de doações acima entre 50 mil reais
doadores = rec[rec['Nome do doador (Receita Federal)'].isin(
rec_count[rec_count['Valor receita', 'sum'] > 50000].index.tolist())]
#Reagrupando dataframe para exibição
doadores.groupby(['Nome do doador (Receita Federal)','CPFCNPJ do doador','Sigla Partido',
'Nome da UE','Ano']).agg({'Valor receita':['count','sum','mean']}).sort_values(
('Nome do doador (Receita Federal)'), ascending=True)
# -
doadores
| inferencia_causal/trabalho_final/notebook-todos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib as mlp
import matplotlib.pyplot as plt
from matplotlib import pyplot as plt
import seaborn as sns
import os
import requests
import shutil
import numpy as np
import statsmodels.api as sm
import math
from sklearn.model_selection import train_test_split
from sklearn import preprocessing
from sklearn.feature_selection import RFE
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error, r2_score, mean_absolute_error
from scipy import stats
from scipy.stats import kurtosis, skew
# %matplotlib inline
#Download the latest CSV file from the WHO website
url = 'https://covid19.who.int/WHO-COVID-19-global-data.csv'
r = requests.get(url, allow_redirects=True)
#Save the content with name.
open('WHO-COVID-19-global-data.csv', 'wb').write(r.content)
dataframe = pd.read_csv('WHO-COVID-19-global-data.csv')
dataframe.head
USA=dataframe[dataframe.Country == 'United States of America']
USA
plt.style.use('ggplot')
USA.plot(x='Date_reported', y=['New_cases','New_deaths'], title="USA New Cases and New Deaths from 1/3/2020 till 12/8/2020", subplots=True, legend=True, layout=(2,2), figsize=(15,15))
China=dataframe[dataframe.Country == 'China']
China
plt.style.use('ggplot')
China.plot(x='Date_reported', y=['New_cases','New_deaths'], title="China New Cases and New Deaths from 1/3/2020 till 12/8/2020", subplots=True, legend=True, layout=(2,2), figsize=(15,15))
#load the data
usa_data= pd.read_csv('USA1.csv')
#This will set my index as date column
usa_data.index= pd.to_datetime(usa_data['Date_reported'])
usa_data = usa_data.drop(['Date_reported'], axis=1)
usa_data.head()
#Check for missing data
usa_data.isna().any()
#define x & y data
x=usa_data['New_cases']
y=usa_data['Temperature']
plt.plot(x, y,'o', color='cadetblue', label = 'No of New Cases')
#make sure it is formatted
plt.title("USA")
plt.xlabel("New_cases")
plt.ylabel("Temperature")
plt.legend()
plt.show()
#measuring the coorelation
usa_data.corr()
#Check the data type of each column
usa_data.dtypes
#define x & y data
x=usa_data['New_cases']
y=usa_data['Cumulative_cases']
plt.plot(x, y,'o', color='cadetblue', label = 'Coorelation between No of new cases and current cases')
#make sure it is formatted
plt.title("Coorelation between No of new cases and current cases")
plt.xlabel("New_cases")
plt.ylabel("Cumulative_cases")
plt.legend()
plt.show()
usa_data.hist(grid= True, color= 'cadetblue')
plt.figure(figsize = (9,8))
USAcor=USA.corr()
sns.heatmap(USAcor, annot=True, cmap=plt.cm.RdYlBu)
plt.show()
#load the data
china_data= pd.read_csv('China1.csv')
#This will set my index as date column
china_data.index= pd.to_datetime(china_data['Date_reported'])
china_data = china_data.drop(['Date_reported'], axis=1)
china_data.head()
#define x & y data
x=china_data['New_cases']
y=china_data['Temperature']
plt.plot(x, y,'o', color='cadetblue', label = 'No of New Cases')
#make sure it is formatted
plt.title("China")
plt.xlabel("New_cases")
plt.ylabel("Temperature")
plt.legend()
plt.show()
#measuring the coorelation
china_data.corr()
#define x & y data
x=china_data['New_cases']
y=china_data['Cumulative_cases']
plt.plot(x, y,'o', color='cadetblue', label = 'Coorelation between No of new cases and current cases')
#make sure it is formatted
plt.title("Coorelation between No of new cases and current cases")
plt.xlabel("New_cases")
plt.ylabel("Cumulative_cases")
plt.legend()
plt.show()
#define x & y data
x=china_data['Cumulative_deaths']
y=china_data['Cumulative_cases']
plt.plot(x, y,'o', color='cadetblue', label = 'Coorelation between current cases and No of Death')
#make sure it is formatted
plt.title("Coorelation between current cases and total number of deaths")
plt.xlabel("Cumulative Deaths")
plt.ylabel("Cumulative_cases")
plt.legend()
plt.show()
plt.figure(figsize = (9,8))
Chinacor=China.corr()
sns.heatmap(Chinacor, annot=True, cmap=plt.cm.RdYlBu)
plt.show()
uk_df = dataframe[dataframe.Country == 'The United Kingdom']
uk_df.to_csv('uk.csv')
uk_df.head
#load the data
uk_data= pd.read_csv('uk.csv')
#This will set my index as date column
uk_data.index= pd.to_datetime(uk_data['Date_reported'])
uk_data = uk_data.drop(['Date_reported'], axis=1)
uk_data.head()
uk_data.corr()
#define x & y data
x=uk_data['New_cases']
y=uk_data['Cumulative_cases']
plt.plot(x, y,'o', color='cadetblue', label = 'Coorelation between No of new cases and current cases')
#make sure it is formatted
plt.title("Coorelation between No of new cases and current cases")
plt.xlabel("New_cases")
plt.ylabel("Cumulative_cases")
plt.legend()
plt.show()
plt.figure(figsize = (9,8))
UKcor=uk_df.corr()
sns.heatmap(UKcor, annot=True, cmap=plt.cm.RdYlBu)
plt.show()
| docs/report/fa20-523-342/project/code/Project5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (py38)
# language: python
# name: py38
# ---
# ### 6 m is mean nitricline depth and just below 10% light level
import matplotlib.pyplot as plt
import netCDF4 as nc
import numpy as np
import os
import glob
import datetime as dt
from salishsea_tools import viz_tools
from matplotlib.ticker import FormatStrFormatter
import cmocean
from salishsea_tools import viz_tools, evaltools as et
#import NorthNut as nn
import matplotlib.gridspec as gridspec
import pickle
import matplotlib as mpl
import matplotlib.patheffects as path_effects
mpl.rc('xtick', labelsize=8)
mpl.rc('ytick', labelsize=8)
mpl.rc('legend', fontsize=8)
mpl.rc('axes', titlesize=8)
mpl.rc('axes', labelsize=8)
mpl.rc('figure', titlesize=8)
mpl.rc('font', size=8)
mpl.rc('text', usetex=True)
mpl.rc('text.latex', preamble = r'''
\usepackage{txfonts}
\usepackage{lmodern}
''')
mpl.rc('font', family='sans-serif', weight='normal', style='normal')
from pandas.plotting import register_matplotlib_converters
register_matplotlib_converters()
# %matplotlib inline
with nc.Dataset('/ocean/eolson/MEOPAR/NEMO-forcing/grid/mesh_mask201702_noLPE.nc') as m:
tmask=np.copy(m.variables['tmask'][0,:,:,:])
fig,ax=plt.subplots(1,1,figsize=(3,5))
ax.contour(tmask[0,:,:],levels=[.5,])
ax.set_xlim(105,370)
ax.set_ylim(200,800)
ig0=0
ig1=398
jg0=0
jg1=800
tmask=tmask[:,jg0:jg1,ig0:ig1]
start=dt.datetime(2015,5,15) # originally 5/15-8/15, but changed to even number of fortnights (6, end is included)
end=dt.datetime(2015,8,20)
mod_basedir='/results/SalishSea/nowcast-green.201812/'
#/data/eolson/results/MEOPAR/SS36runs/linkHC201812/'
mod_nam_fmt='nowcast'
mod_flen=1
saveloc='/data/eolson/results/MEOPAR/SS36runs/calcFiles/NTransport/'
fver='HC201812Base'
tmask.shape
tmask2=np.expand_dims(tmask,0)
tmask2.shape
fliste3t=et.index_model_files(start,end,mod_basedir,mod_nam_fmt,mod_flen,'carp_T',1)
flistPP=et.index_model_files(start,end,mod_basedir,mod_nam_fmt,mod_flen,'ptrc_T',1)
t0=start
te=end
fformat0='%Y%m%d'
recalc=True
savepath=saveloc+'N6_LD_'+fver+'_'+t0.strftime(fformat0)+'-'+te.strftime(fformat0)+'.pkl'
times=[t0+dt.timedelta(hours=ii) for ii in range(0,int((te-t0).total_seconds()/3600)+24)]
if recalc==True:
## calculations
tf0=flistPP.loc[0,['t_0']].values[0]
tfe=flistPP.loc[len(flistPP)-1,['t_n']].values[0]-dt.timedelta(days=1)
IPPx=np.empty((int((tfe-tf0).days*24+24),jg1-jg0,ig1-ig0))
iii0=int((t0-tf0).total_seconds()/3600)
iiie=int((te-tf0).days*24+24)
ti=t0
for iif in range(0,len(flistPP)):
li0=iif*mod_flen*24
li1=(iif+1)*mod_flen*24
with nc.Dataset(flistPP.loc[iif,['paths']].values[0]) as fPP:
IPPx[li0:li1,...]=tmask2[:,6,:,:]*fPP.variables['nitrate'][:,6,jg0:jg1,ig0:ig1]
#constrain to correct times
IPP=IPPx[iii0:iiie,...]
data=dict()
data['NO3_6m']=IPP
pickle.dump(data,open(savepath,'wb'))
else:
data=pickle.load(open(savepath,'rb'))
IPP=data['NO3_6m']
fig,ax=plt.subplots(1,1,figsize=(3,4))
m=ax.pcolormesh(np.mean(IPP,0),vmin=0,vmax=.0005)
fig.colorbar(m)
| notebooks/revisions2/northDomainSummerN6-HC1812-LargeDomain.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="oQXxlpTD85az" colab_type="code" colab={}
# Deep Residual Learning for Image Recognition.
import torch
import torch.nn as nn
import torch.nn.functional as F
import torchvision
import torchvision.transforms as transforms
# + id="LEJCXxA-DEIQ" colab_type="code" colab={}
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True,
download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=4,
shuffle=True, num_workers=2)
testset = torchvision.datasets.CIFAR10(root='./data', train=False,
download=True, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=4,
shuffle=False, num_workers=2)
classes = ('plane', 'car', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
# + id="JVrq4mzNDRxe" colab_type="code" colab={}
# + id="daA7bADhpoCo" colab_type="code" colab={}
# viewing the image
import matplotlib.pyplot as plt
import numpy as np
# functions to show an image
def imshow(img):
img = img / 2 + 0.5 # unnormalize
npimg = img.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
# get some random training images
dataiter = iter(trainloader)
images, labels = dataiter.next()
# show images
imshow(torchvision.utils.make_grid(images))
# print labels
print(' '.join('%5s' % classes[labels[j]] for j in range(4)))
# + id="dGJuS6_R9I_1" colab_type="code" colab={}
class BasicBlock(nn.Module):
expansion = 1
def __init__(self, in_planes, planes, stride=1):
super(BasicBlock, self).__init__()
self.conv1 = nn.Conv2d(in_planes, planes, kernel_size=3, stride=stride, padding=1, bias=False)
self.bn1 = nn.BatchNorm2d(planes)
self.conv2 = nn.Conv2d(planes, planes, kernel_size=3, stride=1, padding=1, bias=False)
self.bn2 = nn.BatchNorm2d(planes)
self.shortcut = nn.Sequential()
if stride != 1 or in_planes != self.expansion*planes:
self.shortcut = nn.Sequential(
nn.Conv2d(in_planes, self.expansion*planes, kernel_size=1, stride=stride, bias=False),
nn.BatchNorm2d(self.expansion*planes)
)
def forward(self, x):
out = F.relu(self.bn1(self.conv1(x)))
out = self.bn2(self.conv2(out))
out += self.shortcut(x)
out = F.relu(out)
return out
# + id="iQ-vU98A9Olb" colab_type="code" colab={}
class Bottleneck(nn.Module):
expansion = 4
def __init__(self, in_planes, planes, stride=1):
super(Bottleneck, self).__init__()
self.conv1 = nn.Conv2d(in_planes, planes, kernel_size=1, bias=False)
self.bn1 = nn.BatchNorm2d(planes)
self.conv2 = nn.Conv2d(planes, planes, kernel_size=3, stride=stride, padding=1, bias=False)
self.bn2 = nn.BatchNorm2d(planes)
self.conv3 = nn.Conv2d(planes, self.expansion*planes, kernel_size=1, bias=False)
self.bn3 = nn.BatchNorm2d(self.expansion*planes)
self.shortcut = nn.Sequential()
if stride != 1 or in_planes != self.expansion*planes:
self.shortcut = nn.Sequential(
nn.Conv2d(in_planes, self.expansion*planes, kernel_size=1, stride=stride, bias=False),
nn.BatchNorm2d(self.expansion*planes)
)
def forward(self, x):
out = F.relu(self.bn1(self.conv1(x)))
out = F.relu(self.bn2(self.conv2(out)))
out = self.bn3(self.conv3(out))
out += self.shortcut(x)
out = F.relu(out)
return out
# + id="T_lbSdcZ9bpN" colab_type="code" colab={}
class ResNet(nn.Module):
def __init__(self, block, num_blocks, num_classes=10):
super(ResNet, self).__init__()
self.in_planes = 64
self.conv1 = nn.Conv2d(3, 64, kernel_size=3, stride=1, padding=1, bias=False)
self.bn1 = nn.BatchNorm2d(64)
self.layer1 = self._make_layer(block, 64, num_blocks[0], stride=1)
self.layer2 = self._make_layer(block, 128, num_blocks[1], stride=2)
self.layer3 = self._make_layer(block, 256, num_blocks[2], stride=2)
self.layer4 = self._make_layer(block, 512, num_blocks[3], stride=2)
self.linear = nn.Linear(512*block.expansion, num_classes)
def _make_layer(self, block, planes, num_blocks, stride):
strides = [stride] + [1]*(num_blocks-1)
layers = []
for stride in strides:
layers.append(block(self.in_planes, planes, stride))
self.in_planes = planes * block.expansion
return nn.Sequential(*layers)
def forward(self, x):
out = F.relu(self.bn1(self.conv1(x)))
out = self.layer1(out)
out = self.layer2(out)
out = self.layer3(out)
out = self.layer4(out)
out = F.avg_pool2d(out, 4)
out = out.view(out.size(0), -1)
out = self.linear(out)
return out
# + id="xQxOJRgl9kXr" colab_type="code" colab={}
def ResNet18():
return ResNet(BasicBlock, [2,2,2,2])
# + id="Fl9XjfU89lEt" colab_type="code" colab={}
def ResNet34():
return ResNet(BasicBlock, [3,4,6,3])
# + id="_6twa9EZ9oND" colab_type="code" colab={}
def ResNet50():
return ResNet(Bottleneck, [3,4,6,3])
# + id="qGg1aAaV9qKs" colab_type="code" colab={}
def ResNet101():
return ResNet(Bottleneck, [3,4,23,3])
# + id="QTpCxi-59sid" colab_type="code" colab={}
def ResNet152():
return ResNet(Bottleneck, [3,8,36,3])
# + id="MpsMGwmO9u6k" colab_type="code" colab={}
def test():
net = ResNet18()
y = net(torch.randn(1,3,32,32))
print(y.size())
# test()
# + id="M4xBbDROpoCv" colab_type="code" colab={}
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(net.parameters(), lr=0.001, momentum=0.9)
# + id="G2mDbcDw_mai" colab_type="code" colab={}
for epoch in range(2): # loop over the dataset multiple times
running_loss = 0.0
for i, data in enumerate(trainloader, 0):
# get the inputs
inputs, labels = data
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs = ResNet(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
# print statistics
running_loss += loss.item()
if i % 2000 == 1999: # print every 2000 mini-batches
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / 2000))
running_loss = 0.0
print('Finished Training')
| session_3/EVA4_Session8_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import torch
from torch import nn
from torch.nn import functional as F
from torchvision import transforms, datasets
# **NOTE**: it is recommended to watch [this link](https://drive.google.com/file/d/1jARX0gjNZwpkcMloOnE8HmngIYDQ6sIB/view?usp=sharing) about "Intoduction of how to code in Pytorch" instructed by <NAME> beforehand.
# ### What is Tensor?
# tensor is mostly same as numpy array (even its applications like broadcasting operation, indexing, slicing and etc), except for it brings us the opportunity to run operations on faster hardwares like GPU. let's see some tensor defintion
# +
arr = torch.zeros((256, 256), dtype=torch.int32)
# tensors are defined by default at CPU
print(arr.device)
# keep 'size', 'dtype' and 'device' same as arr, but fill with 1
arr2 = torch.ones_like(arr)
# keep 'dtype' and 'device' same as arr, but fill data arbitrarily
arr3 = arr.new_tensor([[1, 2], [3, 4]])
# -
# in order to feed tensors to deep-learning models, they should follow a customary shape form; `B C H W` for 4D tensors where `B` is batch size, `C` is channel dimension and `H W` are spatial dimensions.
# #### Device determination
# first we need to determine which device all torch tensors (including the input, learning weights and etc) are going to be allocated. basically, GPU is the first priority.
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# #### Pseudo random generation
# it is often recommended to generate **pseudo** random numbers as it provides fair comparison between different configs of deep learning model(s). torch provides this by `torch.manual_seed`.
# +
np.random.seed(12345)
# same seed on all devices; both CPU and CUDA
torch.manual_seed(12345)
# -
# ## Build a CNN model
# from now on, you will learn how to build and train a CNN model.
#
# pytorch models are defined as python classes inherited from `torch.nn.Module`. two functions are essential for model creation:
# 1. learning weights (parameters) and network layers are defined within `__init__()`.
# 2. forwarding procedure of the model is developed within `forward()`.
#
# so let's create a multi-classification CNN model (with ten ground-truth labels) containing the following layers: `Conv` -> `ReLU` -> `Batchnorm` -> `Conv` -> `ReLU` -> `Batchnorm` -> `Adaptive average pooling` -> `dropout` -> `fully connected`. suppose the input has only one channel and `forward()` will only return output of the model.
class Model(nn.Module):
def __init__(self):
super().__init__()
# your code here
def forward(self, x: torch.Tensor) -> torch.Tensor:
# your code here
return x
# #### set model device
# Previously, we have determined which device (GPU or CPU) is going to be used, although it has not been allocated yet to parameters of the model. Pytorch `.to(device)` Api provides this for us.
# +
model = Model()
model.to(device)
# -
# #### Model phases
# there are two phases for a Pytorch model: `.train()` and `.eval()`. models are by default at `.train()` phase, however the difference between these two is that in `eval()` phase, some layers change their behavior during inference; for instance dropout will be deactivated and batch normalization will not update estimated mean and variance and they will be used only for normalization, hence please note **`.eval()` will not block parameters to be updated**. therefore during evaluation, besides `model.eval()` we should assure that back propagation is temporarily deactivated and this is possible by `torch.no_grad()`. indeed disabling the gradient calculation enables us to use bigger batch sizes as it speeds up the computation and reduces memory usage.
# ## Data processing
# Before training, we need to prepare and process our dataset which is MNIST here.
# #### Data transformation
# PIL images should first be transformed to torch tensors. `torchvision.transforms.Compose` provides a pipeline of transforms. in the following 'converting to tensors' is only applied.
transform = transforms.Compose([
transforms.ToTensor()
])
# #### Download data
# as evaluation is not purpose of this notebook, you only need to load **train** set of MNIST dataset using `torchvision.datasets.MNIST`.
# your code here
train = None
# #### Data loader
# define train loader using `torch.utils.data.DataLoader`.
# +
batch_size = 32
# your code here
train_loader = None
# -
# ## Training
# here we are going to develop training process of MNIST classification.
# #### Optimizer
# define your optimizer, use `torch.optim`.
# your code here
optimizer = None
# #### Procedure
# implement the procedure of training in the following cell. please note **evaluation is not purpose of this notebook**, therefore only report the training loss changes which ought to be descending in general. consider cross entropy as loss function and compute it without using pre-defined APIs.
# the backpropagation consists of three sub-parts:
# 1. gradient computation
# 2. updating learning parameters
# 3. removing current computed gradients for next iteration
#
# fortunately we don't need to implement them from sctrach as pytorch provides APIs for them.
# +
num_epochs = 3
num_iters = len(train_loader)
train_losses = np.zeros((num_epochs, num_iters), dtype=np.float32)
for epoch in range(num_epochs):
for it, (X, y) in enumerate(train_loader):
## forward model
## compute loss
## backpropagation
| notebooks/basics/pytorch_intro/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def test_exercise_48_1(x) -> bool:
import sys
if "xlrd" in sys.modules:
return False
else:
return True
def test_exercise_48_2(x) -> bool:
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
df = pd.read_excel("../Sample - Superstore.xls")
df.head().equals(x)
def test_exercise_48_3(x) -> bool:
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
df = pd.read_excel("../Sample - Superstore.xls")
df.drop('Row ID',axis=1,inplace=True)
df.equals(x)
| Chapter04/unit_tests/.ipynb_checkpoints/Exercise 48-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''env2'': conda)'
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
from tralo.experiments import experiment
results = experiment('mnist_experiment.yaml')
results
# +
from tralo.visualize import plot_experiment
plot_experiment('mnist_experiment.yaml')
# -
# ## Individual Plots
df = results.dataframe()
# +
from matplotlib import pyplot as plt
plt.grid(True)
plt.scatter(df['mnist_loss'], df['mnist_accuracy'])
| doctest/experiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Membership Testing
# When it comes to container type objects, we are often interested in knowing whether the container contains a certain object or value.
# Membership operators are `in` and `not in`, and they work just as you might expect:
10 in [10, 20, 30]
100 in [10, 20, 30]
# But what happens when we are not dealing with the same object, but objects whose values compare equal (`is` vs `==`).
#
# Membership testing always uses `==` (which may fall back to using `is` in certain cases - but beyond the scope of this primer - for now just assume it will be using `==`).
# For example:
a = [100_000, 200_000]
x = 100_000
a[0] is x, a[0] == x
# So `a[0]` and `x` are not the same objects, but they do compare equal, and membership testing gives us this result:
x in a
# Tuples, sets, and even dictionaries support membership testing.
#
# Memrship testing in lists and tuples is very slow compared to membership testing in sets and dictionaries - so try to use those data types where possible if you are going to do a lot of membership tests. That's because memrship testing in a structure such as a list or tuple essentially has to iterate through the elements starting at index 0 until it finds (or does not find) what it is looking for. Sets and dictionaries on the other hand, are based on hash maps, which makes lookups extremely efficient in comparison (in fact that's one of the primary reasons for using dictionaries!)
# For dictionaries, membership testing actually just uses the keys, not the values:
d = {'a': 100, 'b': 200}
'a' in d
100 in d
| 06 - Membership.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import statsmodels.formula.api as smf
import statsmodels.api as sm
# %matplotlib inline
df = pd.read_csv('../Data_Files/CloakofInvisibility.dat', sep='\t')
df.head()
plt.figure(figsize=(10,6))
ax1 = plt.subplot(121)
_ = sns.boxplot(x='cloak', y='mischief1', data=df)
ax2 = plt.subplot(122, sharey=ax1)
_ = sns.boxplot(x='cloak', y='mischief2', data=df)
robust_anova= smf.ols('mischief2~mischief1+C(cloak)', data=df).fit()
table_robust = sm.stats.anova_lm(robust_anova, typ=2,robust='hc3')
table_robust
| Python/statistics_with_Python/11_GLM2_ANCOVA/Markdown_notebook/02_Robust_Ancova.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using critical sections
# A [critical section](https://en.wikipedia.org/wiki/Linearizability) is a region of code that should not run in parallel. For example, the increment of a variable is not considered an [atomic operation](https://en.wikipedia.org/wiki/Linearizability), so, it should be performed using [mutual exclusion](https://en.wikipedia.org/wiki/Mutual_exclusion).
# ## What happens when mutial exclusion is not used in critical sections?
#
# ### Using threads
# All Python’s built-in data structures (such as lists, dictionaries, etc.) are thread-safe. However, other user's data structures implemented by users, or simpler types like integers and floats, should not be accesed concurrently.
# +
# Two threads that have a critical section executed in parallel without mutual exclusion.
# This code does not work!
import threading
import time
counter = 10
def task_1():
global counter
for i in range(10**6):
counter += 1
def task_2():
global counter
for i in range(10**6+1):
counter -= 1
thread_1 = threading.Thread(target=task_1)
thread_2 = threading.Thread(target=task_2)
thread_1.start()
thread_2.start()
print("(Both threads started)")
thread_1.join()
thread_2.join()
print("\nBoth threads finished")
print('counter =', counter)
# -
# The same example, using mutual exclusion (using a [lock](https://docs.python.org/3/library/threading.html#lock-objects)):
# +
# Two threads that have a critical section executed sequentially.
import threading
import time
lock = threading.Lock()
counter = 10
def task_1():
global counter
for i in range(10**6):
with lock:
counter += 1
def task_2():
global counter
for i in range(10**6+1):
with lock:
counter -= 1
thread_1 = threading.Thread(target=task_1)
thread_2 = threading.Thread(target=task_2)
now = time.perf_counter() # Real time (not only user time)
thread_1.start()
thread_2.start()
print("Both threads started")
thread_1.join()
thread_2.join()
print("Both threads finished")
elapsed = time.perf_counter() - now
print(f"elapsed {elapsed:0.2f} seconds")
print('counter =', counter)
# -
# Notice that both tasks are CPU-bound. This means that using `threading` has not any wall time advantage compared to an iterative implementation of both taks.
# ### Using processes
# +
# Two processes that have a critical section executed sequentially
import multiprocessing
import time
import ctypes
def task_1(lock, counter):
for i in range(10000):
with lock:
counter.value += 1
def task_2(lock, counter):
for i in range(10001):
with lock:
counter.value -= 1
lock = multiprocessing.Lock()
manager = multiprocessing.Manager()
counter = manager.Value(ctypes.c_int, 10)
process_1 = multiprocessing.Process(target=task_1, args=(lock, counter))
process_2 = multiprocessing.Process(target=task_2, args=(lock, counter))
now = time.perf_counter()
process_1.start()
process_2.start()
print("Both tasks started")
process_1.join()
process_2.join()
print("Both tasks finished")
elapsed = time.perf_counter() - now
print(f"elapsed {elapsed:0.2f} seconds")
print('counter =', counter.value)
# -
# Unlike `threading`, `multiprocessing` is suitable for reducing the running times in the case of CPU-bound problems.
# ### Using coroutines
# Like threads, coroutines should be only used when the coroutines must wait (typically for an I/O transaction). Otherwhise, use `multiprocessing`.
# +
import asyncio
counter = 10
async def task_1():
global counter
for i in range(10):
print("o", end='', flush=True)
counter += 1
await task_2()
async def task_2():
global counter
print("O", end='', flush=True)
counter -= 1
await task_1()
print('\ncounter =', counter)
# +
import asyncio
import time
counter = 10
async def task_1():
global counter
for i in range(10**6):
counter += 1
await task_2()
async def task_2():
global counter
counter -= 1
now = time.perf_counter()
await task_1()
elapsed = time.perf_counter() - now
print(f"\nelapsed {elapsed:0.2f} seconds")
print('counter =', counter)
# -
# Coroutines are faster than threads, but not faster than the one-loop version of the task.
# +
import time
counter = 10
def task():
global counter
for i in range(10**6):
counter += 1
counter -= 1
now = time.perf_counter()
task()
elapsed = time.perf_counter() - now
print(f"\nelapsed {elapsed:0.2f} seconds")
print('counter =', counter)
| 04-concurrency/mutial_exclusion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="../images/qiskit_header.png" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" align="middle">
# # Qiskit Tutorials
#
# ***
#
#
# Welcome Qiskitters.
#
#
# These tutorials aim to explain how to use Qiskit. We assume you have installed Qiskit; if not, please look at [qiskit.org](http://www.qiskit.org) or the install [documentation](https://qiskit.org/documentation/install.html).
#
#
# We've collected a core reference set of notebooks in this section outlining the features of Qiskit. We will be keeping them up to date with the latest Qiskit version, currently 0.12. The focus of these notebooks is not on learning quantum computing. Instead we will be focused on how to use Qiskit, and will go into details only when needed. For those interested in learning about quantum computing we recommend the awesome [educational material](https://quantum-computing.ibm.com/support) we and the community have put together.
#
#
# Qiskit is made up of four elements: Terra, Aer, Ignis, and Aqua. Each element has its own goal, and together they make the full Qiskit framework.
#
# ## Getting started with Qiskit
#
# This section gives you the tools to make your first circuits, run them on real quantum systems and simulators, and view the data.
#
# 1. [Getting started with Qiskit](fundamentals/1_getting_started_with_qiskit.ipynb) - How to use Qiskit.
#
# 2. [Plotting data in Qiskit](fundamentals/2_plotting_data_in_qiskit.ipynb) - Illustrates the different ways of plotting data in Qiskit.
#
# 3. [The IBM Q Account](fundamentals/3_the_ibmq_account.ipynb) - Understanding the IBM Q account.
#
# 4. [Circuit Properties](fundamentals/4_quantum_circuit_properties.ipynb) - Important properties of quantum circuits.
#
# 5. [Using the Transpiler](fundamentals/5_using_the_transpiler.ipynb) - Mapping and optimizing circuits using the Qiskit transpiler.
#
# 6. [Jupyter Tools](fundamentals/6_qiskit_jupyter_tools.ipynb) - Qiskit functionality for Jupyter notebooks.
#
# 7. [Summary of quantum operations](fundamentals/7_summary_of_quantum_operations.ipynb) - List of quantum operations (gates, reset, measurements) in Qiskit Terra
#
#
# ## 2 Qiskit Terra
#
# Terra, the ‘earth’ element, is the foundation on which the rest of the software lies. Terra provides a bedrock for composing quantum programs at the level of circuits and pulses, to optimize them for the constraints of a particular device, and to manage the execution of batches of experiments on remote-access devices. Terra defines the interfaces for a desirable end-user experience, as well as the efficient handling of layers of optimization, pulse scheduling and backend communication.
#
# 1. [Advanced circuits](advanced/terra/1_advanced_circuits.ipynb) - Circuit building tools added including registerless declarations, composite gate updates and parameterized circuits.
# 2. [Operators overview](advanced/terra/2_operators_overview.ipynb) - Gives a summary of the features and uses of the Operator class.
# 3. [Advanced circuit visualization](advanced/terra/3_advanced_circuit_visualization.ipynb) - Details on drawing your quantum circuits.
# 4. [Transpiler passes and passmanager](advanced/terra/4_transpiler_passes_and_passmanager.ipynb) - How to use the transpiler passes, passmanger, and extend the transpiler with a new pass.
# 5. [Pulse schedules](advanced/terra/5_pulse_schedules.ipynb) - An introduction to working with pulse schedules.
# 6. [Creating a new provider](advanced/terra/6_creating_a_provider.ipynb) - A guide to integration of a new provider with Qiskit structures and interfaces
#
#
#
# ## 3 Qiskit Aer
#
# Aer, the ‘air’ element, permeates all Qiskit elements. To really speed up development of quantum computers, we need better simulators with the ability to model realistic noise processes that occur during computation on actual devices. Aer provides a high-performance simulator framework for studying quantum computing algorithms and applications in the noisy intermediate-scale quantum regime.
# 1. [Aer provider](advanced/aer/1_aer_provider.ipynb) - Gives a summary of the Qiskit Aer provider containing the Qasm, statevector, and unitary simulator
# 2. [Device noise simulation](advanced/aer/2_device_noise_simulation.ipynb) - Shows how to use the Qiskit Aer noise module to automatically generate a basic noise model for simulating hardware backends
# 3. [Building noise models](advanced/aer/3_building_noise_models.ipynb) - Shows how to use Qiskit Aer noise module to construct custom noise models for noisy simulations
# 4. [Custom gate noise](advanced/aer/4_custom_gate_noise.ipynb) - Shows to implement simulations using custom noisy gates.
# 5. [Noise transformations](advanced/aer/5_noise_transformation.ipynb) - Demonstrates the noise approximation utility functions to construct approximate Clifford noise models out of a general noise model
# 6. [Extended stabilizer tutorial](advanced/aer/6_extended_stabilizer_tutorial.ipynb) - Gives an overview of the *extended stabilizer* Qasm Simulator method
# 7. [Matrix Product State simulator](advanced/aer/7_matrix_product_state_method.ipynb) - Gives an overview of the *matrix product state* Simulator method
#
# ## 4 Qiskit Ignis
# Ignis, the ‘fire’ element, is dedicated to fighting noise and errors and to forging a new path. This includes better characterization of errors, improving gates, and computing in the presence of noise. Ignis is meant for those who want to design quantum error correction codes, or who wish to study ways to characterize errors through methods such as tomography and randomized benchmarking, or even to find a better way for using gates by exploring dynamical decoupling and optimal control. Ignis tutorials are found [here](advanced/ignis/) and include:
# 1. [Calibrating a qubit](advanced/ignis/1_calibrating_a_qubit.ipynb) - Using pulse to calibrate a "pi-pulse" gate by fitting a Rabi oscillation on a qubit. Using the "pi-pulse" measure the single-shot analog voltages that are returned by an experiment.
# 2. [Hamiltonian and gate characterizations](advanced/ignis/2_hamiltonian_and_gate_characterization.ipynb) - Sequences to measure ZZ rates between qubits and to measure rotation and angle errors in the gates.
# 3. [Relaxation and decoherence](advanced/ignis/3_relaxation_and_decoherence.ipynb) - How to measure coherence times on the real quantum hardware
# 4. [Measurement error mitigation](advanced/ignis/4_measurement_error_mitigation.ipynb) - How to peform calibration experiments for measurement errors and fed those calibrations into a "filter" that can be utilized to mitigate errors in subsequent experiments.
# 5. Randomized Benchmarking:
# * a. [Randomized benchmarking](advanced/ignis/5a_randomized_benchmarking.ipynb) - Randomized benchmarking (RB) is a technique used to measure the average gate error by measuring the outcomes of random Clifford circuits. This is used internally to report gate errors on our systems.
# * b. [Interleaved RB](advanced/ignis/5b_interleaved_rb.ipynb) - A variant of RB used to measure the error of a specific gate.
# * c. [Purity RB](advanced/ignis/5c_purity_rb.ipynb) - A variant of RB used to measure the *incoherent* error per gate.
# 6. Tomography:
# * a. [Quantum state tomography](advanced/ignis/6a_state_tomography.ipynb) - How to identify a quantum state using state tomography, in which the state is prepared repeatedly and measured in different bases.
# * b. [Quantum process tomography](advanced/ignis/6b_process_tomography.ipynb) - A method to reconstruct the quantum process matrix by preparing certain states, applying a gate, and then measuring the outcome in different bases.
# 7. [Quantum volume](advanced/ignis/7_quantum_volume.ipynb) - How to run quantum volume measurements on the quantum hardware.
# 8. [Repetition Code](advanced/ignis/8_repetition_code.ipynb) - How to run a simple error correction code, known as the repetition code. This can be used to characterize bit flip errors in the hardware.
# 9. [Logging](advanced/ignis/9_ignis_logging.ipynb) - An introduction to some of the logging features in Ignis, intended to be used to track characterization parameters.
#
#
# ## 5 Qiskit Aqua
# Aqua, the ‘water’ element, is the element of life. To make quantum computing live up to its expectations, we need to find real-world applications. Aqua is where algorithms for NISQ computers are built. These algorithms can be used to build applications for quantum computing.
# * [Amplitude Estimation](advanced/aqua/amplitude_estimation.ipynb) - Illustrates amplitude estimation, for a simple case, where the (assumed to be unknown) success probability *p* of a Bernoulli random variable is estimated
# * [HHL](advanced/aqua/linear_systems_of_equations.ipynb) - Solving linear systems of equations with the HHL algorithm
# * [Creating an Aqua algorithm](advanced/aqua/Aqua_how_to_build_a_pluggable_algorithm_components.ipynb) - Building an algorithm within the framework of Aqua
#
# Aqua is accessible to domain experts in *Artificial Intelligence*, *Chemistry*, *Finance* or *Optimization*, who want to explore the benefits of using quantum computers as accelerators for specific computational tasks, without needing to worry about how to translate the problem into the language of quantum machines:
#
# ### 5.1 Qiskit Artificial Intelligence
# [Qiskit AI](advanced/aqua/artificial_intelligence/index.ipynb) demonstates using quantum computers to tackle problems in the artificial intelliegence domain. These include using a quantum-enhanced support vector machine to experiment with classification problems on a quantum computer
#
# ### 5.2 Qiskit Chemistry
# [Qiskit Chemistry](advanced/aqua/chemistry/index.ipynb) - applications in the domain of quantum chemistry on quantum computers, including ground state energy, dipole moments and dissociation plots
#
# ### 5.3 Qiskit Finance
# [Qiskit Finance](advanced/aqua/finance/index.ipynb) - provides a collection of applications of quantum algorithms to use cases relevant in finance. This includes use cases like portfolio management, derivative pricing, or credit risk analysis.
#
# ### 5.4 Qiskit Optimization
# [Qiskit Optimization](advanced/aqua/optimization/index.ipynb) - using VQE (Variational Quantum Eigensolver) to experiment with optimization problems (max-cut and traveling salesman problem) on a quantum computer. Includes optimization problem modelling, using docplex, which can be automatically translated to input suitable for VQE.
#
import qiskit.tools.jupyter
# %qiskit_version_table
# %qiskit_copyright
| qiskit/1_start_here.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="mB5eSZXZIO9W"
# JAX-TensorFlow interoperation with JAX2TF
# ===========================================
#
# Link: go/jax2tf-colab
#
# The JAX2TF colab has been deprecated, and the example code has
# been moved to [jax2tf/examples](https://github.com/google/jax/tree/master/jax/experimental/jax2tf/examples).
#
| jax/experimental/jax2tf/JAX2TF_getting_started.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="BR3JXn5HU9rX"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# + id="hsg8dBEaU9rp"
df = pd.read_csv('train.csv')
# + colab={"base_uri": "https://localhost:8080/"} id="ybj7VbTfU9r5" outputId="9eead217-48ce-471c-9c0e-7937573c25b0"
df.shape
# + colab={"base_uri": "https://localhost:8080/"} id="bItpWysqU9sJ" outputId="5c353cff-e132-4ca4-b75c-d3649ecdd5e7"
df.columns[df.isnull().sum() > 0]
# + id="M5pnOVMsU9sW"
df.fillna(method='bfill',inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="EBbh8sjdU9sh" outputId="793d6f88-f687-49a1-f9d2-d4ea359611c9"
df.columns[df.isnull().sum() > 0]
# + id="ljoFuOYvU9st"
df.fillna(method='ffill',inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="vSdpRINJU9s7" outputId="fee759e3-b073-495e-9da2-96bcf7de115d"
df.columns[df.isnull().sum() > 0]
# + colab={"base_uri": "https://localhost:8080/"} id="2VeG9qEkU9tG" outputId="4b5b620b-ccb7-4e97-cbd7-f81fd0e50269"
Numeric_columns=df.select_dtypes(include=np.number).columns.tolist()
categorical_col=set(df.columns).difference(set(Numeric_columns))
print(categorical_col)
# + id="pOmPIsQfU9tV"
df['edjefe'] = df['edjefe'].replace({'no': 0, 'yes':1}).astype(float)
df['edjefa'] = df['edjefa'].replace({'no': 0, 'yes':1}).astype(float)
# + colab={"base_uri": "https://localhost:8080/"} id="3ioc78SCU9ta" outputId="a22cdc19-c7c2-48e6-96b7-a2525bbba952"
Numeric_columns=df.select_dtypes(include=np.number).columns.tolist()
categorical_col=set(df.columns).difference(set(Numeric_columns))
print(categorical_col)
# + id="SUeLDsazU9tz"
df['dependency'] = np.sqrt(df['SQBdependency'])
# + colab={"base_uri": "https://localhost:8080/"} id="ZGlLtv9gU9t-" outputId="3b16e8e0-53d2-4d3d-9f11-b35ede2fe604"
df.isna().sum().sum()
# + id="mvJDRbVBU9uI"
col_drops = ['Id','idhogar']
df.drop(col_drops,axis=1,inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="IcaKVSpbU9uT" outputId="90ad37d5-0ba8-47ad-c234-ad68a1364526"
# finding numeric and categorical columns
Numeric_columns=df.select_dtypes(include=np.number).columns.tolist()
categorical_col=set(df.columns).difference(set(Numeric_columns))
# numeric cols to numeric
print(categorical_col)
# + id="RSQ9MEt5U9ug"
train_set = df
# + id="OneyW2qyU9uu"
df['rent_per_adult'] = df['v2a1']/df['hogar_adul']
df['rent_per_person'] = df['v2a1']/df['hhsize']
df['overcrowding_room_and_bedroom'] = (df['hacdor'] + df['hacapo'])/2
df['no_appliances'] = df['refrig'] + df['computer'] + df['television']
# df1['home_condition_poor'] = (df1['epared1'] + df1['etecho1'] + df1['eviv1'])/3
# df1['home_condition_regular'] = (df1['epared2'] + df1['etecho2'] + df1['eviv2'])/3
# df1['home_condition_good'] = (df1['epared3'] + df1['etecho3'] + df1['eviv3'])/3
df['r4h1_percent_in_male'] = df['r4h1'] / df['r4h3']
df['r4m1_percent_in_female'] = df['r4m1'] / df['r4m3']
df['r4h1_percent_in_total'] = df['r4h1'] / df['hhsize']
df['r4m1_percent_in_total'] = df['r4m1'] / df['hhsize']
df['r4t1_percent_in_total'] = df['r4t1'] / df['hhsize']
df['adult'] = df['hogar_adul'] - df['hogar_mayor']
df['dependency_count'] = train_set['hogar_nin'] + df['hogar_mayor']
df['dependency'] = df['dependency_count'] / df['adult']
df['child_percent'] = df['hogar_nin']/df['hogar_total']
df['elder_percent'] = df['hogar_mayor']/df['hogar_total']
df['adult_percent'] = df['hogar_adul']/df['hogar_total']
df['rent_per_bedroom'] = df['v2a1']/df['bedrooms']
df['adults_per_bedroom'] = df['adult']/df['bedrooms']
df['child_per_bedroom'] = df['hogar_nin']/df['bedrooms']
df['male_per_bedroom'] = df['r4h3']/df['bedrooms']
df['female_per_bedroom'] = df['r4m3']/df['bedrooms']
df['bedrooms_per_person_household'] = df['hhsize']/df['bedrooms']
df['tablet_per_person_household'] = df['v18q1']/df['hhsize']
df['phone_per_person_household'] = df['qmobilephone']/df['hhsize']
df['age_12_19'] = df['hogar_nin'] - df['r4t1']
df['rent_per_room'] = df['v2a1']/df['rooms']
df['bedroom_per_room'] = df['bedrooms']/df['rooms']
df['elder_per_room'] = df['hogar_mayor']/df['rooms']
df['adults_per_room'] = df['adult']/df['rooms']
df['child_per_room'] = df['hogar_nin']/df['rooms']
df['male_per_room'] = df['r4h3']/df['rooms']
df['female_per_room'] = df['r4m3']/df['rooms']
df['room_per_person_household'] = df['hhsize']/df['rooms']
df['escolari_age'] = df['escolari']/df['age']
df['rez_esc_escolari'] = df['rez_esc']/df['escolari']
df['rez_esc_r4t1'] = df['rez_esc']/df['r4t1']
df['rez_esc_r4t2'] = df['rez_esc']/df['r4t2']
df['rez_esc_r4t3'] = df['rez_esc']/df['r4t3']
df['rez_esc_age'] = df['rez_esc']/df['age']
# + id="b0J0fSNyU9u-"
from lightgbm import LGBMClassifier
from sklearn.ensemble import RandomForestRegressor
from sklearn.naive_bayes import GaussianNB
# + id="RbJRp_xdU9ve"
from sklearn.model_selection import cross_val_score
from bayes_opt import BayesianOptimization
# + colab={"base_uri": "https://localhost:8080/"} id="bADfa1wTU9wC" outputId="bf557044-5ff8-4e2e-d1c1-f0ed39713f04"
pip install lightgbm
# + colab={"base_uri": "https://localhost:8080/"} id="nc6vcvs_U9wP" outputId="251c7a71-bde5-442f-850f-e4c431ae1557"
# !pip install bayesian-optimization
# + id="DycqVv83U9wo"
x_1 = df.drop('Target',axis=1)
y = df['Target']
# +
cols= [col for col in x_1.columns if x_1[col].isnull().any()]
reduced_x_1 = x_1.drop(cols, axis=1)
# reduced_X_valid = X_valid.drop(cols_with_missing, axis=1)
# + colab={"base_uri": "https://localhost:8080/"} id="UGyTJ72cU9wt" outputId="e5cb7fa8-5aec-42e7-bd1f-bdaa187ae79b"
df.shape
# + colab={"base_uri": "https://localhost:8080/"} id="9BbX-lrvU9w3" outputId="71140855-b723-4ee5-b9cc-ab5ce3cb1a53"
x_1.shape
# -
reduced_x_1.shape
# + id="CotxJMnNU9xN"
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import RepeatedKFold
from sklearn.model_selection import cross_validate,cross_val_score
# + id="GoaSvU_-nCQ_"
def optimize_lgb(data, targets):
def lgb_crossval(n_estimators, num_leaves, min_child_samples, subsample):
return lgb_cv(
n_estimators=int(n_estimators),
num_leaves = int(num_leaves),
min_child_samples=int(min_child_samples),
subsample=subsample,
data=data,
targets=targets,
)
optimizer = BayesianOptimization(
f=lgb_crossval,
pbounds={
"n_estimators": (100,500),
"num_leaves": (30,80),
"min_child_samples": (5,30),
"subsample": (0.6,1.0)
},
random_state=1234,
verbose=2
)
optimizer.maximize(n_iter=25, init_points=20)
print("Final result:", optimizer.max)
# + id="TicgIia4nCRA"
print("--- Optimizing Light GBM ---")
optimize_lgb(x_1, y)
# + id="Ry94-c0rnCRB" outputId="78ef3eda-0e0d-4c4f-fcb2-bfde4278977e"
Lgbm=LGBMClassifier(n_estimators=200,learning_rate=0.1,random_state=45,num_leaves=200)
model = GaussianNB()
# model = RandomForestRegressor(n_estimators=100, random_state=0)
# + id="qo-7u2A-nCRB"
from sklearn.model_selection import train_test_split
# + id="qbHtUdz3nCRC"
x_train,x_test,y_train,y_test = train_test_split(reduced_x_1,y,random_state=42)
#x_train, x_valid, y_train, y_valid = train_test_split(x_1, y, train_size=0.8, test_size=0.2,random_state=0)
# + id="y126p8eOnCRC" outputId="789ba820-8f54-4ed0-e5c1-240946b554fd"
#Lgbm.fit(x_train,y_train)
model.fit(x_train,y_train)
# model.fit(x_train,y_train)
# + id="q-XHijJ7nCRD" outputId="4eba132d-7fae-4b0f-f480-2a224343ad18"
Lgbm.score(x_test,y_test)
# + id="Vdty2GnQnCRD"
df1 = pd.read_csv('test.csv')
# + id="fPkLTy3onCRD"
df1.columns[df1.isnull().sum()> 0]
# + id="ckLilO-knCRE"
df1.fillna(method='bfill',inplace=True)
# + id="9yeE4yvAnCRE"
df1.columns[df1.isnull().sum()> 0]
# -
df1.fillna(method='ffill', inplace=True)
df1.columns[df1.isnull().sum()> 0]
# +
Numeric_columns=df1.select_dtypes(include=np.number).columns.tolist()
categorical_col=set(df1.columns).difference(set(Numeric_columns))
print(categorical_col)
# -
df1['edjefa'] = df1['edjefa'].replace({'no':0,'yes':1}).astype(float)
df1['edjefe'] = df1['edjefe'].replace({'no':0,'yes':1}).astype(float)
# +
Numeric_columns=df1.select_dtypes(include=np.number).columns.tolist()
categorical_col=set(df1.columns).difference(set(Numeric_columns))
print(categorical_col)
# -
df1['dependency'] = np.sqrt(df1['SQBdependency'])
# +
Numeric_columns=df1.select_dtypes(include=np.number).columns.tolist()
categorical_col=set(df1.columns).difference(set(Numeric_columns))
print(categorical_col)
# -
col_drops = ['Id','idhogar']
df1.drop(col_drops,inplace=True,axis=1)
# +
Numeric_columns=df1.select_dtypes(include=np.number).columns.tolist()
categorical_col=set(df1.columns).difference(set(Numeric_columns))
print(categorical_col)
# -
test_set = df1
# +
df1['rent_per_adult'] = df1['v2a1']/df1['hogar_adul']
df1['rent_per_person'] = df1['v2a1']/df1['hhsize']
df1['overcrowding_room_and_bedroom'] = (df1['hacdor'] + df1['hacapo'])/2
df1['no_appliances'] = df1['refrig'] + df1['computer'] + df1['television']
df1['r4h1_percent_in_male'] = df1['r4h1'] / df1['r4h3']
df1['r4m1_percent_in_female'] = df1['r4m1'] / df1['r4m3']
df1['r4h1_percent_in_total'] = df1['r4h1'] / df1['hhsize']
df1['r4m1_percent_in_total'] = df1['r4m1'] / df1['hhsize']
df1['r4t1_percent_in_total'] = df1['r4t1'] / df1['hhsize']
df1['adult'] = df1['hogar_adul'] - df1['hogar_mayor']
df1['dependency_count'] = df1['hogar_nin'] + df1['hogar_mayor']
df1['dependency'] = df1['dependency_count'] / df1['adult']
df1['child_percent'] = df1['hogar_nin']/df1['hogar_total']
df1['elder_percent'] = df1['hogar_mayor']/df1['hogar_total']
df1['adult_percent'] = df1['hogar_adul']/df1['hogar_total']
df1['rent_per_bedroom'] = df1['v2a1']/df1['bedrooms']
df1['adults_per_bedroom'] = df1['adult']/df1['bedrooms']
df1['child_per_bedroom'] = df1['hogar_nin']/df1['bedrooms']
df1['male_per_bedroom'] = df1['r4h3']/df1['bedrooms']
df1['female_per_bedroom'] = df1['r4m3']/df1['bedrooms']
df1['bedrooms_per_person_household'] = df1['hhsize']/df1['bedrooms']
df1['tablet_per_person_household'] = df1['v18q1']/df1['hhsize']
df1['phone_per_person_household'] = df1['qmobilephone']/df1['hhsize']
df1['age_12_19'] = df1['hogar_nin'] - df1['r4t1']
df1['rent_per_room'] = df1['v2a1']/df1['rooms']
df1['bedroom_per_room'] = df1['bedrooms']/df1['rooms']
df1['elder_per_room'] = df1['hogar_mayor']/df1['rooms']
df1['adults_per_room'] = df1['adult']/df1['rooms']
df1['child_per_room'] = df1['hogar_nin']/df1['rooms']
df1['male_per_room'] = df1['r4h3']/df1['rooms']
df1['female_per_room'] = df1['r4m3']/df1['rooms']
df1['room_per_person_household'] = df1['hhsize']/df1['rooms']
df1['escolari_age'] = df1['escolari']/df1['age']
df1['rez_esc_escolari'] = df1['rez_esc']/df1['escolari']
df1['rez_esc_r4t1'] = df1['rez_esc']/df1['r4t1']
df1['rez_esc_r4t2'] = df1['rez_esc']/df1['r4t2']
df1['rez_esc_r4t3'] = df1['rez_esc']/df1['r4t3']
df1['rez_esc_age'] = df1['rez_esc']/df1['age']
# -
x1 = test_set
x1.shape
y.shape
x1.shape
Lgbm1=LGBMClassifier(n_estimators=200,learning_rate=0.1,random_state=45,num_leaves=500)
x1_train,x1_test,y1_train,y1_test = train_test_split(x1[:9557],y,random_state=45)
Lgbm1.fit(x1_train,y1_train)
Lgbm1.score(x1_test,y1_test)
# + active=""
#
| Costa rican household poverty prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dual PET tracer de Pierro with motion
#
# Authors: <NAME>, <NAME>, <NAME>, <NAME>
# First version: 2nd of November 2019
#
# CCP PETMR Synergistic Image Reconstruction Framework (SIRF)
# Copyright 2019 University College London
# Copyright 2019 King's College London
#
# This is software developed for the Collaborative Computational
# Project in Positron Emission Tomography and Magnetic Resonance imaging
# (http://www.ccppetmr.ac.uk/).
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# # The challenge!
#
# This notebook is an open-ended look into using de Pierro MAPEM to reconstruct dual-PET acquisitions.
#
# - Imagine two different scans (FDG and amyloid) were performed in a short space of time on a single patient.
# - Your task is to implement an alternating reconstruction of the two scans using de Pierro's MAPEM algorithm!
#
# ## Suggested workflow - no motion
#
# - Inspire yourself from [de_Pierro_MAPEM.ipynb](de_Pierro_MAPEM.ipynb), in which Bowsher weights are calculated on some known side information.
# - Now, imagine that the side information is evolving with our image estimate
# - We'll probably want to perform an update on one of our images (image A)
# - Then recalculate the Bowsher weights of the second image (image B) with our newly-update image A
# - Then perform a normal de Pierro update on image B
# - Then recalculate the Bowsher weights of image A using our newly-updated image B
#
# ### But what about motion?
#
# It's possible that there's motion between the two images since they were acquired at different times. Once you've got everything working for dual PET reconstructions, it's time to add motion in just to complicate things! We've also given you a misaligned amyloid acquisition, `amyl_sino_noisy_misaligned.hs` and a corresponding $\mu$-map, `uMap_misaligned.hv`.
#
# - Imagine two different scans (FDG and amyloid) were performed in a short space of time on a single patient.
# - Your task is to implement an alternating reconstruction of the two scans using de Pierro's MAPEM algorithm!
# - Bear in mind that the two scans weren't performed at the same time, so the patient's head isn't necessarily in the same place...
#
# ## Suggested workflow - motion
#
# 1. Since we can't be sure of patient position, you should probably reconstruct each image individually
# 2. Then register them
# 3. Then modify your non-motion case, such that you resample each image into the others' space before calculating the Bowsher weights
#
# Hints:
# - For an implementation of de Pierro MAPEM, checkout the [de_Pierro_MAPEM.ipynb](de_Pierro_MAPEM.ipynb) notebook.
# - To go faster, rebin your sinograms (as per [de_Pierro_MAPEM.ipynb](de_Pierro_MAPEM.ipynb)!
# - For registration and resampling, check out the [../Reg/sirf_registration.ipynb](../Reg/sirf_registration.ipynb) notebook.
#
# ### One final word
#
# We've given you some pointers down below that you can fill bit by bit. The sections marked with astrisks won't be needed until you implement the motion case.
# # 0a. Some includes and imshow-esque functions
# +
# All the normal stuff you've already seen
# -
# # 0b. Input data
# +
# # copy files to working folder and change directory to where the output files are
brainweb_sim_data_path = exercises_data_path('working_folder', 'Synergistic', 'BrainWeb')
fname_FDG_sino = os.path.join(brainweb_sim_data_path, 'FDG_sino_noisy.hs')
fname_FDG_uMap = os.path.join(brainweb_sim_data_path, 'uMap_small.hv')
# No motion filenames
fname_amyl_sino = os.path.join(brainweb_sim_data_path, 'amyl_sino_noisy.hs')
fname_amyl_uMap = os.path.join(brainweb_sim_data_path, 'uMap_small.hv')
# Motion filenames
# fname_amyl_sino = 'amyl_sino_noisy_misaligned.hs'
# fname_amyl_uMap = 'uMap_misaligned.hv'
# -
# # 0c. Set up normal reconstruction stuff
# +
# Code to set up objective function and OSEM recontsructors
# -
# # 1. Two individual reconstructions *
# +
# Some code goes here
# -
# # 2. Register images *
# +
# Some more code goes here
# -
# # 3. A resample function? *
# +
# How about a bit of code here?
# -
# # 4. Maybe some de Pierro functions
# +
# A pinch more code here
# -
# # 5. Are we ready?
# +
# Final code!
| notebooks/Synergistic/Dual_PET.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## One Dimensional Motion
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib.ticker as tck
import matplotlib.patches as patches
# %matplotlib inline
# %config InlineBackend.figure_format = 'png2x'
# ## Velocity
#
# > The [velocity](https://en.wikipedia.org/wiki/Velocity) of an object is the rate of change of its position with respect to a frame of reference, and is a function of time. Velocity is equivalent to a specification of its speed and direction of motion (e.g. 60 km/h to the north). Velocity is an important concept in kinematics, the branch of classical mechanics that describes the motion of bodies.
#
# First, let's see how we can represent position in one dimension, starting with a single point `A` at the origin:
# +
def show_position(position, label, origin=(0,0)):
x, y = position, 0
ax = plt.axes()
plt.plot(x, y, 'o')
plt.annotate(label, xy=(x,y), xytext=(x + 0.1, -0.25))
plt.xlim(origin[0]-1.5, origin[0]+1.5)
plt.ylim(origin[1]-1.5, origin[1]+1.5)
plt.show()
show_position(0, '$A_0$')
# -
# Now, assume `A` moves relative to some reference frame and the position changes:
show_position(1, '$A_1$')
# The difference between the two positions is called displacement, defined as $ \Delta x = x_f - x_0 $, where
#
# - $ x_f $ is the final position
# - $ x_0 $ is the initial position
# - $ \Delta x $ is dispacement (also $ \vec{s} $)
# +
def show_displacement(x_0, x_f):
y = 0
ax = plt.axes()
plt.annotate('$x_0$', xy=(x_0,y), xytext=(x_0 + 0.1, -0.25))
plt.annotate('$x_f$', xy=(x_f,y), xytext=(x_f + 0.1, -0.25))
ax.add_patch(patches.FancyArrowPatch(
(x_0, y),
(x_f, y),
mutation_scale=20
))
plt.annotate('$\Delta x = x_f - x_0 = %.1f $' % (x_f - x_0), xy=((x_f-x_0)/2, y), xytext=((x_f-x_0)/2, 0.25))
plt.xlim(x_0-1.5, x_0+1.5)
plt.ylim(y-1.5, y+1.5)
plt.show()
show_displacement(0, 1)
# -
# Velocity is then defined as displacement over time:
#
# $$
# \vec{v} = \frac{\vec{s}}{t}
# $$
#
# One convenient way of representing change in position over time is using a Position vs. Time graph:
# +
def show_position_vs_time():
positions = [3, 7, 6, 0, -6, -7, -3]
times = range(0, len(positions))
ax = plt.axes()
ax.set_xlabel('$t$')
ax.set_ylabel('$x$')
plt.plot(times, positions, '-')
plt.show()
show_position_vs_time()
# -
# To find the instantaneous velocity at a given point in time, we only need to calculate the slope of our line at that time. Since the measurements in this case are discrete, all of the slopes are straight lines and can be calculated using two neighboring points.
#
# Consider if our graph was a continuous function like $ sin(x) $:
# +
def show_position_vs_time_sinx():
times = np.arange(0, 2 * np.pi, .01)
positions = np.sin(times)
ax = plt.axes()
ax.set_xlabel('$t$')
ax.set_ylabel('$x$')
ax.xaxis.set_major_formatter(tck.FormatStrFormatter('%g $\pi$'))
plt.plot(times / np.pi, positions, '-')
plt.show()
show_position_vs_time_sinx()
# -
# We can use differential calculus to find the instantaneous velocity at time $ t $:
#
# $$
# \vec{v}_t = \frac{d}{dx}\ sin(x_t)\ dx\ =\ cos(x_t)
# $$
#
# If our position vs. time graph is $ sin(x) $ this means our velocity vs. time graph is $ cos(x) $:
# +
def show_velocity_sinx():
times = np.arange(0, 2 * np.pi, .01)
positions = np.sin(times)
velocities = np.cos(times)
ax = plt.axes()
ax.set_xlabel('$t$')
ax.xaxis.set_major_formatter(tck.FormatStrFormatter('%g $\pi$'))
plt.plot(times / np.pi, positions, '-', label='$p$')
plt.plot(times / np.pi, velocities, '--', label='$\hat{v}$')
ax.legend()
plt.show()
show_velocity_sinx()
# -
# We could also take the second derivative to find the acceleration:
#
# $$
# \vec{a} = \frac{ \Delta{\vec{v}} }{ \Delta t }
# $$
#
# $$
# a_t = \frac{d}{dx}\ cos(x)\ =\ -sin(x)
# $$
# +
def show_accel_sinx():
times = np.arange(0, 2 * np.pi, .01)
positions = np.sin(times)
velocities = np.cos(times)
accelerations = -np.sin(times)
ax = plt.axes()
ax.set_xlabel('$t$')
ax.xaxis.set_major_formatter(tck.FormatStrFormatter('%g $\pi$'))
plt.plot(times / np.pi, positions, '-', label='$p$')
plt.plot(times / np.pi, velocities, '--', label='$\hat{v}$')
plt.plot(times / np.pi, accelerations, '--', label='$\hat{a}$')
ax.legend()
plt.show()
show_accel_sinx()
# -
| 01_mechanics/1.1-one-dimensional-motion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import xmitgcm as xm
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import plotly.graph_objects as go
import plotly.figure_factory as ff
import sys
import xarray as xr
# %matplotlib inline
# %config InlineBackend.figure_format='retina'
# -
# Note that 2-D and 3-D arrays should be index-ordered as $(y,x)$ and $(z, y, x)$ respectively, to be written in proper ordering for MITgcm.
# Writing utility
def writefield(fname, data, dtype=np.dtype("float32")):
dcopy = data.copy()
print("write to file: " + fname)
if dcopy.dtype != dtype:
dcopy = dcopy.astype(dtype)
print("changed dtype to " + str(dcopy.dtype))
if sys.byteorder == "little":
dcopy.byteswap(True)
fid = open(fname, "wb")
dcopy.tofile(fid)
fid.close()
def openfield(fname, shape, dtype=np.dtype("float32")):
data = np.fromfile(fname, dtype)
if sys.byteorder == "little":
data.byteswap(True)
return data.reshape(shape)
# # Set up grid
# +
Ho = -3982.2607 # ocean depth in meters
Lx = 1000
Ly = 2000
nx = 20 # number of gridpoints in x-direction - 1000km/5km
ny = 40 # number of gridpoints in y-direction - 2000km/5km
nz = 49 # number of vertical levels
dx = Lx / nx
dy = Ly / ny
X = np.linspace(0, Lx, nx, dtype="float32")
Y = np.linspace(0, Ly, ny, dtype="float32")
XX, YY = np.meshgrid(X, Y)
# -
# # Bathymetry
# Flat bottom at z=-Ho
h = -Ho * np.ones((ny, nx), dtype="float32")
h[(0, -1), :] = 0 # Walls at top and bottom of domain
# # Load pickup data
ds = xm.open_mdsdataset(data_dir="/scratch/daanr/ACC_run_coarse/Diags", prefix=["2D_diags", "GM_diags", "state"], calendar="360_days", read_grid=True,
geometry="cartesian")
float(ds.isel(time=69).time)/1e9 * 1000 / (360*24*60*60)
# # Export binaries for U, V, $\Theta$ & $\eta$
writefield("pickup_temp.bin", ds.isel(time=69).THETA.values)
writefield("pickup_uvel.bin", ds.isel(time=69).UVEL.values)
writefield("pickup_vvel.bin", ds.isel(time=69).VVEL.values)
writefield("pickup_etan.bin", ds.isel(time=69).ETAN.values)
# # Initial tracer (dummy salinity) setup
# We release the initial tracer profile at concentration 1 psu. Initial profile is padded (10 boxes on each side) with exponential decay to prevent spurious oscillations.
X_release = int(ds.XC[10].data)
Y_release = int(ds.YC[20].data)
print(X_release, Y_release)
XX, YY = np.meshgrid(ds.XC.data, ds.YC.data)
dist_to_release = np.sqrt(np.square(X_release - XX) + np.square(Y_release - YY))
# The initial tracer distribution is
# $$
# \begin{cases}
# \left(\frac{1}{(2 e)^{r/50,000}}-\frac{1}{2 e}\right)/\left(1-\frac{1}{2e}\right) &\qquad r<=50,000 \text{ meters}\\
# 0 &\qquad r>50,000 \text{ meters}
# \end{cases}
# $$
# where
# $$
# r = \sqrt{(X_r-X)^2 + (Y_r-Y)^2}
# $$
S_3D = np.zeros((nz, ny, nx), dtype='float32')
S_3D[24, :, :] = np.where(dist_to_release/50000<=1,(1/np.power(2*np.exp(1), dist_to_release/50000)- 1/(2*np.exp(1)))/(1-1/(2*np.exp(1))), 0)
plt.imshow(S_3D[24, :, :])
S_3D[24, :, :]
plt.plot(X, S_3D[24,20,:])
plt.xlabel('X [km]')
plt.ylabel(r'tracer concentration [$m^{-3}$]')
plt.show()
writefield("salinity_tracer_point.bin", S_3D)
| ACC_mitgcm_config/reentrant_coarse/input_ridge_5y_tracer_loca/pickup_field_creator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="J-SBMroaAsxu" colab_type="text"
# # Code to train LightGBM model
# This notebook was made on Google Colab ([link to notebook](https://colab.research.google.com/drive/1VtCe6znPTggCk5Bp_CJ1Hv9eRtgBdtT0?usp=sharing)). If you are running on your local machine remove the cells related to data retrieval and RDKit installation, and change the paths to the files as needed.
#
# + [markdown] id="AprnAEjfrD10" colab_type="text"
# #Software installation and data retrieval, etc.
# + id="iCRyXY2zqyrX" colab_type="code" colab={}
# %%capture
# Install RDKit
# takes 2-3 minutes
# !wget -c https://repo.continuum.io/miniconda/Miniconda3-latest-Linux-x86_64.sh
# !chmod +x Miniconda3-latest-Linux-x86_64.sh
# !time bash ./Miniconda3-latest-Linux-x86_64.sh -b -f -p /usr/local
# !time conda install -q -y -c conda-forge rdkit
# + id="xW0uiaZhBaQn" colab_type="code" colab={}
# %%capture
# ! rm -f lightgbm_density_train_data.csv lightgbm_barrier_train_data.csv
# ! wget https://sid.erda.dk/share_redirect/EwaEr2JMrb/all_230_billion/ml_data/lightgbm_density_train_data.csv
# ! wget https://sid.erda.dk/share_redirect/EwaEr2JMrb/all_230_billion/ml_data/lightgbm_barrier_train_data.csv
# + id="RPe4Es5arKvC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="e43a4456-4239-4731-b14d-bc38970ce211"
import sys
import re
from collections import OrderedDict
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# RDKit
sys.path.append('/usr/local/lib/python3.7/site-packages/')
from rdkit import Chem
from rdkit.Chem import AllChem
from rdkit.Chem import Descriptors
# ML
from sklearn.model_selection import train_test_split
import lightgbm as lgb
# + id="EzouqSGutgEz" colab_type="code" colab={}
plt.rcParams['xtick.labelsize'] = 16
plt.rcParams['ytick.labelsize'] = 16
plt.rcParams['axes.labelsize'] = 16
# + [markdown] id="ee9XLdj9rBbH" colab_type="text"
# # Helper Functions
# Functions to make RDKit object and the find number of substituents from gene names.
# + id="8IKN_h6ArASt" colab_type="code" colab={}
ligands = ['H', 'F', 'Cl', 'Br', 'C(F)(F)(F)', 'C#N',
'[N+](=O)([O-])', 'C(=O)[H]', 'C(=O)O',
'C(=O)C', 'C(=O)N', 'C#C', 'S(=O)(=O)(C)',
'C=N', 'O', 'OC', 'N', 'N(C)(C)', 'C',
'N(C(=O)(C))', 'SC', 'C3=CC=C(F)C=C3',
'C3=CC=C(Cl)C=C3', 'C3=CC=C(Br)C=C3',
'C3=CC=C(C(F)(F)(F))C=C3',
'C3=CC=C(C#N)C=C3', 'C3=CC=C([N+](=O)([O-]))C=C3',
'C3=CC=C(C(=O)[H])C=C3', 'C3=CC=C(C(=O)O)C=C3',
'C3=CC=C(C(=O)C)C=C3', 'C3=CC=C(C(=O)N)C=C3',
'C3=CC=C(C#C)C=C3', 'C3=CC=C(S(=O)(=O)(C))C=C3',
'C3=CC=C(C=N)C=C3', 'C3=CC=C(O)C=C3',
'C3=CC=C(OC)C=C3', 'C3=CC=C(N)C=C3',
'C3=CC=C(N(C)(C))C=C3', 'C3=CC=C(C)C=C3',
'C3=CC=C(N(C(=O)(C)))C=C3', 'C3=CC=C(SC)C=C3',
'C3=CC=CC=C3']
ligand_num2smiles = OrderedDict(zip(list(range(len(ligands))),ligands))
def get_mol(gene):
global ligands
gene = gene.split('-')
core = 'N#CC1(C#N)C(1)=C(2)C2=C(3)C(4)=C(5)C(6)=C(7)C21'
gene_smiles = core
for i, ligand_idx in enumerate(gene):
substr = ligands[int(ligand_idx)]
if substr == 'H':
continue
gene_smiles = gene_smiles.replace('('+str(i+1)+')', f'({substr})')
mol = Chem.MolFromSmiles(re.sub('\([0-9]\)', '', gene_smiles))
return mol
def num_subs(gene):
gene = [int(x) for x in gene.split('-')]
return 7 - gene.count(0)
def get_smiles(rdkit_mol):
return Chem.MolToSmiles(Chem.RemoveHs(rdkit_mol))
# + [markdown] id="i7cthSSErXH8" colab_type="text"
# # Prepare Data
# + id="I0fwjkZghKVt" colab_type="code" colab={}
df_density = pd.read_csv('lightgbm_density_train_data.csv', sep=',')
df_density['num_subs'] = df_density['gene'].apply(num_subs)
# + id="u8HHh3T0hmZi" colab_type="code" colab={}
df_barrier_scan = pd.read_csv('lightgbm_barrier_train_data.csv')
df_barrier_scan['num_subs'] = df_barrier_scan['gene'].apply(num_subs)
# + [markdown] id="lifT1S82r-P0" colab_type="text"
# # Train LightGBM Model
# + [markdown] id="gIOrWTjksscz" colab_type="text"
# ## Function for One-Hot Encoding
#
# + id="BW1qWOIbrpEV" colab_type="code" colab={}
def one_hot(gene):
"""Creates one-hot representation"""
one_hot = np.zeros((7,41), dtype=np.float32)
for i, base in enumerate([int(x) for x in gene.split('-')]):
if base == 0:
continue
one_hot[i,base - 1] = 1.
return one_hot
def training_data_one_hot(df, prop='density'):
""" Create Matrices and Vectors with representation and targets repectively """
Xs = np.empty((df.shape[0], 287))
ys = np.empty(df.shape[0])
for i, row in enumerate(df.itertuples()):
Xs[i] = one_hot(row.gene).flatten()
ys[i] = getattr(row, prop)
return Xs, ys
# + [markdown] id="2ZSG5Rmpv4vw" colab_type="text"
# ### Density Data
# + id="pGyxMp3EwgEZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3be1e8ac-bda5-44a2-892e-e70ed44df14e"
df_train_density, df_test_density = train_test_split(df_density, test_size=0.25, random_state=42)
df_test_density, df_val_density = train_test_split(df_test_density, test_size=0.50, random_state=42)
print(f"train size: {df_train_density.shape[0]}, validation size: {df_val_density.shape[0]}, test size: {df_test_density.shape[0]}")
# + id="2mHkomVUsS2D" colab_type="code" colab={}
train_data_density, train_label_density = training_data_one_hot(df_train_density, prop='xtb_density')
lgb_train_density = lgb.Dataset(train_data_density, label=train_label_density)
val_data_density, val_label_density = training_data_one_hot(df_val_density, prop='xtb_density')
lgb_val_density = lgb.Dataset(val_data_density, label=val_label_density)
test_data_density, test_label_density = training_data_one_hot(df_test_density, prop='xtb_density')
lgb_test_density = lgb.Dataset(test_data_density, label=test_label_density)
# + [markdown] id="5qkWAu_6v-hO" colab_type="text"
# ### Barrier Data
# + id="T26MqVcJwD1D" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2fa678ce-8d78-4fb6-9c22-5ae4b35a0220"
df_train_barrier, df_test_barrier = train_test_split(df_barrier_scan, test_size=0.17, random_state=42)
df_test_barrier, df_val_barrier = train_test_split(df_test_barrier, test_size=0.40, random_state=42)
print(f"train size: {df_train_barrier.shape[0]}, validation size: {df_val_barrier.shape[0]}, test size: {df_test_barrier.shape[0]}")
# + id="-mylUL-kzxW2" colab_type="code" colab={}
train_data_barrier, train_label_barrier = training_data_one_hot(df_train_barrier, prop='pm3_barrier_scan')
lgb_train_barrier = lgb.Dataset(train_data_barrier, label=train_label_barrier)
val_data_barrier, val_label_barrier = training_data_one_hot(df_val_barrier, prop='pm3_barrier_scan')
lgb_val_barrier = lgb.Dataset(val_data_barrier, label=val_label_barrier)
test_data_barrier, test_label_barrier = training_data_one_hot(df_test_barrier, prop='pm3_barrier_scan')
lgb_test_barrier = lgb.Dataset(test_data_barrier, label=test_label_barrier)
# + [markdown] id="NXhpwjOvwGrL" colab_type="text"
# ## Train Models
# + id="8aBq2dEQsTgF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 156} outputId="227a0754-7dbe-4822-d177-e5f5669a174c"
params = {
'boosting_type': 'gbdt',
'objective': 'regression',
'metric': ['l2', 'rmse'],
'num_leaves': 80,
'max_depth': 7,
'min_data_in_leaf': 50,
'max_bin': 100,
'learning_rate': 0.10,
'seed': 42}
num_round = 10000
model_lgb_density = lgb.train(params, lgb_train_density, num_round, valid_sets=[lgb_val_density],
early_stopping_rounds=200, verbose_eval=1000)
# + id="9UJMChBSwKBe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="b40ab081-f262-4443-a693-433d6525c6a8"
params = {
'boosting_type': 'gbdt',
'objective': 'regression',
'metric': ['l2', 'rmse'],
'num_leaves': 8,
'max_depth': 3,
'min_data_in_leaf': 80,
'max_bin': 100,
'learning_rate': .25,
'seed': 42}
max_num_trees = 10000
model_lgb_barrier = lgb.train(params, lgb_train_barrier, max_num_trees, valid_sets=[lgb_val_barrier],
early_stopping_rounds=100, verbose_eval=500)
# + [markdown] id="Cbluf-H3zGg9" colab_type="text"
# # Make Figure 5
# + id="0oVN6CbL076O" colab_type="code" colab={}
ypred_test_density = model_lgb_density.predict(test_data_density, num_iteration=model_lgb_density.best_iteration)
ypred_test_barrier = model_lgb_barrier.predict(test_data_barrier, num_iteration=model_lgb_barrier.best_iteration)
# + id="qa1-SF8TtIRf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 398} outputId="d0ac54b0-37f8-4480-b171-d08b2017f079"
fig, ax = plt.subplots(figsize=(13,6), ncols=2)
# Plot Density
sns.scatterplot(test_label_density, ypred_test_density, ax=ax[0])
ax[0].plot([0.0, 0.80], [0.0, 0.80], c='k', linestyle='--')
error_density = ypred_test_density - test_label_density
mae = abs(error_density).mean()
rms = (error_density).std()
ax[0].set(xlabel='xTB Storage Density [kJ/g]', ylabel='LightGBM Storage Density [kJ/g]')
s_density = f'RMS: {rms:.4f} \nMAE: {mae:.4f}'
ax[0].text(0.40,0.05, s_density, {'fontsize': 16})
# Plot Barrier
sns.scatterplot(test_label_barrier, ypred_test_barrier, ax=ax[1])
ax[1].plot([80, 350], [80, 350], c='k', linestyle='--')
error_barrier = ypred_test_barrier - test_label_barrier
mae = abs(error_barrier).mean()
rms = (error_barrier).std()
ax[1].set(xlabel='PM3 Scan Back Reaction [kJ/mol]', ylabel='LightGBM Scan Back Reaction [kJ/mol]')
s_barrier = f'RMS: {rms:.2f} \nMAE: {mae:.2f}'
ax[1].text(250,100, s_barrier, {'fontsize': 16});
| train_ml_models/train_lightgbm_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.012353, "end_time": "2021-03-02T06:04:23.932283", "exception": false, "start_time": "2021-03-02T06:04:23.919930", "status": "completed"} tags=[]
# # مروری کلی بر کرونا
# > مروری بر کل موارد کرونا ، مرگ و میر و موارد جدید براساس کشور.
#
# - comments: true
# - author: <NAME> <NAME>
# - categories: [مرور]
# - image: images/covid-overview.png
# - permalink: /covid-overview/
# - hide: false
# + papermill={"duration": 0.024498, "end_time": "2021-03-02T06:04:24.329343", "exception": false, "start_time": "2021-03-02T06:04:24.304845", "status": "completed"} tags=[]
#hide
print('''
Example of using jupyter notebook, pandas (data transformations), jinja2 (html, visual)
to create visual dashboards with fastpages
You see also the live version on https://gramener.com/enumter/covid19/
''')
# + papermill={"duration": 3.139528, "end_time": "2021-03-02T06:04:27.478425", "exception": false, "start_time": "2021-03-02T06:04:24.338897", "status": "completed"} tags=[]
#hide
# !pip install jdatetime
# !pip install persiantools
import numpy as np
import pandas as pd
from jinja2 import Template
import jdatetime as jd
import os
from persiantools import digits
from IPython.display import HTML
# + papermill={"duration": 0.133916, "end_time": "2021-03-02T06:04:27.621695", "exception": false, "start_time": "2021-03-02T06:04:27.487779", "status": "completed"} tags=[]
#hide
# FETCH
base_url = 'https://raw.githubusercontent.com/pratapvardhan/notebooks/master/covid19/'
dirpath = os.path.dirname(os.getcwd())
paths = {
'mapping': base_url + 'mapping_countries.csv',
'overview': dirpath + '/_templates/overview.tpl'
}
def get_mappings(url):
df = pd.read_csv(url)
return {
'df': df,
'replace.country': dict(df.dropna(subset=['Name']).set_index('Country')['Name']),
'map.continent': dict(df.set_index('Name')['Continent'])
}
mapping = get_mappings(paths['mapping'])
def get_template(path):
from urllib.parse import urlparse
if bool(urlparse(path).netloc):
from urllib.request import urlopen
return urlopen(path).read().decode('utf8')
return open(path).read()
def get_frame(name):
url = (
'https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/'
f'csse_covid_19_time_series/time_series_covid19_{name}_global.csv')
df = pd.read_csv(url)
# rename countries
df['Country/Region'] = df['Country/Region'].replace(mapping['replace.country'])
return df
def get_dates(df):
dt_cols = df.columns[~df.columns.isin(['Province/State', 'Country/Region', 'Lat', 'Long'])]
LAST_DATE_I = -1
# sometimes last column may be empty, then go backwards
for i in range(-1, -len(dt_cols), -1):
if not df[dt_cols[i]].fillna(0).eq(0).all():
LAST_DATE_I = i
break
return LAST_DATE_I, dt_cols
# + papermill={"duration": 0.367706, "end_time": "2021-03-02T06:04:27.999100", "exception": false, "start_time": "2021-03-02T06:04:27.631394", "status": "completed"} tags=[]
#hide
COL_REGION = 'Country/Region'
# Confirmed, Recovered, Deaths
df = get_frame('confirmed')
# dft_: timeseries, dfc_: today country agg
dft_cases = df
dft_deaths = get_frame('deaths')
dft_recovered = get_frame('recovered')
LAST_DATE_I, dt_cols = get_dates(df)
dt_today = dt_cols[LAST_DATE_I]
dt_5ago = dt_cols[LAST_DATE_I-5]
dfc_cases = dft_cases.groupby(COL_REGION)[dt_today].sum()
dfc_deaths = dft_deaths.groupby(COL_REGION)[dt_today].sum()
dfp_cases = dft_cases.groupby(COL_REGION)[dt_5ago].sum()
dfp_deaths = dft_deaths.groupby(COL_REGION)[dt_5ago].sum()
# + papermill={"duration": 0.057316, "end_time": "2021-03-02T06:04:28.066234", "exception": false, "start_time": "2021-03-02T06:04:28.008918", "status": "completed"} tags=[]
#hide
df_table = (pd.DataFrame(dict(Cases=dfc_cases, Deaths=dfc_deaths, PCases=dfp_cases, PDeaths=dfp_deaths))
.sort_values(by=['Cases', 'Deaths'], ascending=[False, False])
.reset_index())
df_table.rename(columns={'index': 'Country/Region'}, inplace=True)
for c in 'Cases, Deaths'.split(', '):
df_table[f'{c} (+)'] = (df_table[c] - df_table[f'P{c}']).clip(0) # DATA BUG
df_table['Fatality Rate'] = (100 * df_table['Deaths'] / df_table['Cases']).round(1)
df_table['Continent'] = df_table['Country/Region'].map(mapping['map.continent'])
df_table.head(15)
# + papermill={"duration": 0.01896, "end_time": "2021-03-02T06:04:28.095814", "exception": false, "start_time": "2021-03-02T06:04:28.076854", "status": "completed"} tags=[]
#hide
#delete problematic countries from table
df_table = df_table[~df_table['Country/Region'].isin(['Cape Verde', 'Cruise Ship', 'Kosovo'])]
# + papermill={"duration": 0.034128, "end_time": "2021-03-02T06:04:28.140415", "exception": false, "start_time": "2021-03-02T06:04:28.106287", "status": "completed"} tags=[]
#hide
# world, china, europe, us
metrics = ['Cases', 'Deaths', 'Cases (+)', 'Deaths (+)']
s_china = df_table[df_table['Country/Region'].eq('China')][metrics].sum().add_prefix('China ')
s_us = df_table[df_table['Country/Region'].eq('US')][metrics].sum().add_prefix('US ')
s_eu = df_table[df_table['Continent'].eq('Europe')][metrics].sum().add_prefix('EU ')
s_iran = df_table[df_table['Country/Region'].eq('Iran')][metrics].sum().add_prefix('Iran ')
summary = {'updated': pd.to_datetime(dt_today), 'since': pd.to_datetime(dt_5ago)}
summary = {**summary, **df_table[metrics].sum(), **s_china, **s_us, **s_eu, **s_iran}
summary
# + papermill={"duration": 0.041209, "end_time": "2021-03-02T06:04:28.192181", "exception": false, "start_time": "2021-03-02T06:04:28.150972", "status": "completed"} tags=[]
#hide
dft_ct_cases = dft_cases.groupby(COL_REGION)[dt_cols].sum()
dft_ct_new_cases = dft_ct_cases.diff(axis=1).fillna(0).astype(int)
dft_ct_new_cases.head()
# + papermill={"duration": 0.071639, "end_time": "2021-03-02T06:04:28.281343", "exception": false, "start_time": "2021-03-02T06:04:28.209704", "status": "completed"} tags=[]
#hide_input
template = Template(get_template(paths['overview']))
days = digits.en_to_fa(str((summary['updated'] - summary['since']).days))
g_date_updated = pd.to_datetime(summary['updated'])
jdate_updated = jd.date.fromgregorian(year=g_date_updated.year,month=g_date_updated.month,day=g_date_updated.day,locale='fa_IR').strftime("%d / %m / %Y")
jdate_updated = digits.en_to_fa(jdate_updated)
g_date_since = pd.to_datetime(summary['since'])
jdate_since = jd.date.fromgregorian(year=g_date_since.year,month=g_date_since.month,day=g_date_since.day,locale='fa_IR').strftime("%d / %m / %Y")
jdate_since = digits.en_to_fa(jdate_since)
html = template.render(
D=summary, table=df_table, # REMOVE .head(20) to see all values
newcases=dft_ct_new_cases.loc[:, dt_cols[LAST_DATE_I-50]:dt_cols[LAST_DATE_I]],
np=np, pd=pd, days=days, digits=digits,
jdate_updated=jdate_updated, jdate_since=jdate_since, enumerate=enumerate)
HTML(f'<div>{html}</div>')
# + [markdown] papermill={"duration": 0.012122, "end_time": "2021-03-02T06:04:28.305714", "exception": false, "start_time": "2021-03-02T06:04:28.293592", "status": "completed"} tags=[]
#
# تحلیل اصلی از [پراتاپ واردهان](https://twitter.com/PratapVardhan)[^۱]
#
# [^۱]: منبع: ["2019 Novel Coronavirus COVID-19 (2019-nCoV) Data Repository by Johns Hopkins CSSE"](https://systems.jhu.edu/research/public-health/ncov/) [GitHub repository](https://github.com/CSSEGISandData/COVID-19). لینک به صفحه [اصلی](https://github.com/pratapvardhan/notebooks/blob/master/covid19/covid19-compare-country-trajectories.ipynb).
| _notebooks/2020-03-21-covid19-overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn import preprocessing
from sklearn import metrics
import pandas as pd
from sklearn.model_selection import KFold
from IPython.display import display
from ipywidgets import IntProgress
import pandas as pd
import os
from sklearn.model_selection import RandomizedSearchCV
from time import time
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import make_scorer, roc_auc_score
from sklearn.neural_network import MLPRegressor
from sklearn.linear_model import Perceptron
if '__file__' in locals():
current_folder = os.path.dirname(os.path.abspath(__file__))
else:
current_folder = os.getcwd()
merge_features = '"{}"'.format(os.path.join(current_folder, '..', 'Features', 'Merge features.ipynb'))
calcular_auc = '"{}"'.format(os.path.join(current_folder, '..', 'Calcular AUC.ipynb'))
set_de_entrenamiento_testing_y_prediccion = '"{}"'.format(os.path.join(
current_folder,'..','Set de entrenamiento, testing y predicción.ipynb'))
predicciones_csv = os.path.join(current_folder, '..', 'predictions.csv')
pd.options.mode.chained_assignment = None
# %run $merge_features
assert(df_features.shape[0] == df['person'].unique().shape[0])
# +
# %run $set_de_entrenamiento_testing_y_prediccion
labels_with_features = labels.merge(df_features, how='inner', on='person')
data = labels_with_features.drop('label', axis=1)
target = labels_with_features['label']
# +
param_dist = {
'penalty' : [None,'l2','l1','elasticnet'],
'alpha' : np.arange(0.0001,0.005,0.001)
}
cv_splits = 100 # cantidad de splits en el cross validation
n_iter_search = 20 # cantidad de puntos, en total splits*n_iter_search
perceptron = Perceptron()
random_search = RandomizedSearchCV(perceptron, param_distributions=param_dist, iid=False, refit=True, verbose=10,
return_train_score=True, n_iter=n_iter_search, cv=cv_splits,
scoring='roc_auc', n_jobs=-1);
start = time()
random_search.fit(data, target)
print("RandomizedSearchCV took %.2f seconds for %d candidates"
" parameter settings." % ((time() - start), n_iter_search))
# -
random_search.cv_results_
random_search.best_params_
random_search.best_score_
| Algoritmos de ML/Perceptron_hiperparametros.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Apache Toree - Scala
// language: scala
// name: apache_toree_scala
// ---
// # 00: Spark Scala Tutorial
//
// 
//
// <NAME>, Ph.D.<br/>
// [Lightbend](http://lightbend.com)<br/>
// [<EMAIL>](mailto:<EMAIL>)<br/>
// [@deanwampler](https://twitter.com/deanwampler)
//
// This tutorial demonstrates how to write and run [Apache Spark](http://spark.apache.org) applications using Scala with some SQL. I also teach a little Scala as we go, but if you already know Spark and you are more interested in learning just enough Scala for Spark programming, see my other tutorial [Just Enough Scala for Spark](https://github.com/deanwampler/JustEnoughScalaForSpark).
//
// ## Introduction
//
// This top-level notebook will guide you through the tutorial. But first, let's discuss [Apache Spark](https://spark.apache.org), a distributed computing system written in Scala for distributed data programming. Besides Scala, you can program Spark using Java, Python, R, and SQL! This tutorial focuses on Scala and SQL.
//
// ### Spark Streaming and SQL
//
// Spark includes support for stream processing, using an older [DStream](https://spark.apache.org/docs/latest/streaming-programming-guide.html) or a newer [Structured Streaming](https://spark.apache.org/docs/latest/structured-streaming-programming-guide.html) backend, as well as more traditional batch-mode applications.
//
// > **Note:** The streaming examples in this tutorial use the older library. Newer examples are TODO.
//
// There is a [SQL](http://spark.apache.org/docs/latest/sql-programming-guide.html) module for working with data sets through SQL queries or a SQL-like API. It integrates the core Spark API with embedded SQL queries with defined schemas. It also offers [Hive](http://hive.apache.org) integration so you can query existing Hive tables, even create and delete them. Finally, it supports a variety of file formats, including CSV, JSON, Parquet, ORC, etc.
//
// There is also an interactive shell, which is an enhanced version of the Scala REPL (read, eval, print loop shell).
// ### Why Spark?
//
// By 2013, it became increasingly clear that a successor was needed for the venerable [Hadoop MapReduce](http://wiki.apache.org/hadoop/MapReduce) compute engine. MapReduce applications are difficult to write, but more importantly, MapReduce has significant performance limitations and it can't support event-streaming ("real-time") scenarios.
//
// Spark was seen as the best, general-purpose alternative, so all the major Hadoop vendors announced support for it in their distributions.
// ### Spark Clusters
//
// Let's briefly discuss the anatomy of a Spark cluster, adapting [this discussion (and diagram) from the Spark documentation](http://spark.apache.org/docs/latest/cluster-overview.html). Consider the following diagram:
//
// 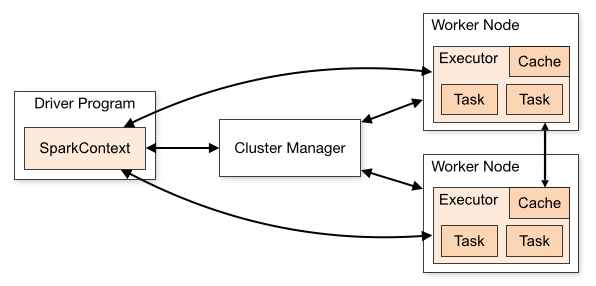
//
// Each program we'll write is a *Driver Program*. It uses a [SparkContext](http://spark.apache.org/docs/latest/api/scala/index.html#org.apache.spark.SparkContext) to communicate with the *Cluster Manager*, which is an abstraction over Hadoop YARN, Mesos, standalone (static cluster) mode, EC2, and local mode.
//
// The *Cluster Manager* allocates resources. An *Executor* JVM process is created on each worker node per client application. It manages local resources, such as the cache (see below) and it runs tasks, which are provided by your program in the form of Java jar files or Python scripts.
//
// Because each application has its own executor process per node, applications can't share data through the *Spark Context*. External storage has to be used (e.g., the file system, a database, a message queue, etc.)
// ### Resilient, Distributed Datasets
//
// 
//
// The data caching is one of the key reasons that Spark's performance is considerably better than the performance of MapReduce. Spark stores the data for the job in *Resilient, Distributed Datasets* (RDDs), where a logical data set is partitioned over the cluster.
//
// The user can specify that data in an RDD should be cached in memory for subsequent reuse. In contrast, MapReduce has no such mechanism, so a complex job requiring a sequence of MapReduce jobs will be penalized by a complete flush to disk of intermediate data, followed by a subsequent reloading into memory by the next job.
//
// RDDs support common data operations, such as *map*, *flatmap*, *filter*, *fold/reduce*, and *groupby*. RDDs are resilient in the sense that if a "partition" of data is lost on one node, it can be reconstructed from the original source without having to start the whole job over again.
//
// The architecture of RDDs is described in the research paper [Resilient Distributed Datasets: A Fault-Tolerant Abstraction for In-Memory Cluster Computing](https://www.usenix.org/system/files/conference/nsdi12/nsdi12-final138.pdf).
// ### SparkSQL
//
// [SparkSQL](http://spark.apache.org/docs/latest/sql-programming-guide.html) first introduced a new `DataFrame` type that wraps RDDs with schema information and the ability to run SQL queries on them. A successor called `Dataset` removes some of the type safety "holes" in the `DataFrame` API, although that API is still available.
//
// There is an integration with [Hive](http://hive.apache.org), the original SQL tool for Hadoop, which lets you not only query Hive tables, but run DDL statements too. There is convenient support for reading and writing various formats like [Parquet](http://parquet.io) and JSON.
// ### The Spark Version
//
// This tutorial uses Spark 2.2.0.
//
// The following documentation links provide more information about Spark:
//
// * [Documentation](http://spark.apache.org/docs/latest/).
// * [Scaladocs API](http://spark.apache.org/docs/latest/api/scala/index.html#org.apache.spark.package).
//
// The [Documentation](http://spark.apache.org/docs/latest/) includes a getting-started guide and overviews of the various major components. You'll find the [Scaladocs API](http://spark.apache.org/docs/latest/api/scala/index.html#org.apache.spark.package) useful for the tutorial.
// ## The Examples and Exercises
//
// Here is a list of the examples, some of which have exercises embedded as comments. Click each link to navigate to the corresopnding notebook. Note that each name ends with a number, indicating the order in which you should study them:
//
// | Example Notebook | Description |
// | :--------------- | :---------- |
// | <a href="01_Intro.ipynb" target="01_I">01_Intro</a> | The first example demonstrates several features of typical Spark jobs. The fact that it's easy to embed this code in a notebook demonstrates that it's easy to work with Spark interactively. |
// | <a href="02_WordCount.ipynb" target="02_WC">02_WordCount</a> | The *Word Count* algorithm: read a corpus of documents, tokenize it into words, and count the occurrences of all the words. A classic, simple algorithm used to learn many Big Data APIs. By default, it uses a file containing the King James Version (KJV) of the Bible. (The `data` directory has a [README](data/README.html) that discusses the sources of the data files.) |
// | <a href="03_WordCount.ipynb" target="03_WC">03_WordCount</a> | An alternative implementation of *Word Count* that uses a slightly different approach and also uses a library to handle input command-line arguments, demonstrating some idiomatic (but fairly advanced) Scala code. |
// | <a href="04_Matrix.ipynb" target="04_M">04_Matrix</a> | Demonstrates using explicit parallelism on a simplistic Matrix application. |
// | <a href="05a_Crawl.ipynb" target="05a_C">05a_Crawl</a> | Simulates a web crawler that builds an index of documents to words, the first step for computing the *inverse index* used by search engines. The documents "crawled" are sample emails from the Enron email dataset, each of which has been classified already as SPAM or HAM. |
// | <a href="05b_InvertedIndex.ipynb" target="05b_I">05b_InvertedIndex</a> | Using the crawl data, compute the index of words to documents (emails). |
// | <a href="06_NGrams.ipynb" target="06_N">06_NGrams</a> | Find all N-word ("NGram") occurrences matching a pattern. In this case, the default is the 4-word phrases in the King James Version of the Bible of the form `% love % %`, where the `%` are wild cards. In other words, all 4-grams are found with `love` as the second word. The `%` are conveniences; the NGram Phrase can also be a regular expression, e.g., `% hated? % %` finds all the phrases with `hate` and `hated`. |
// | <a href="07_Joins.ipynb" target="07_J">07_Joins</a> | Spark supports SQL-style joins as shown in this simple example. Note this RDD approach is obsolete; use the SparkSQL alternatives. |
// | <a href="08_SparkSQL.ipynb" target="08_S">08_SparkSQL</a> | Uses the SQL API to run basic queries over structured data in `DataFrames`, in this case, the same King James Version (KJV) of the Bible used in the previous tutorial. There is also a script version of this file. Using the _spark-shell_ to do SQL queries can be very convenient! |
// | <a href="09_SparkSQL-File-Formats.ipynb" target="09_S">09_SparkSQL-File-Formats</a> | Demonstrates writing and reading [Parquet](http://parquet.io)-formatted data, namely the data written in the previous example. |
// | <a href="10_SparkStreaming.ipynb" target="10_S">10_SparkStreaming</a> | The older _structured streaming_ (`DStream`) capability. Here it's used to construct a simplistic "echo" server. Running it is a little more involved, as discussed below. |
// ## Going Forward from Here
//
// To learn more, see the following resources:
//
// * [Lightbend's Fast Data Platform](http://lightbend.com/fast-data-platform) - a curated, fully-supported distribution of open-source streaming and microservice tools, like Spark, Kafka, HDFS, Akka Streams, etc.
// * The Apache Spark [website](http://spark.apache.org/).
// * [Talks from the Spark Summit conferences](http://spark-summit.org).
// * [Learning Spark](http://shop.oreilly.com/product/0636920028512.do), an excellent introduction from O'Reilly, if now a bit dated.
//
// ## Final Thoughts
//
// Thank you for working through this tutorial. Feedback and pull requests are welcome.
//
// [<NAME>](mailto:<EMAIL>)
//
| notebooks/00_Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
plt.rcParams['figure.figsize'] = (18, 10)
df = pd.read_csv('player-0.csv')
del df['Unnamed: 0']
from collections import Counter
letter_counts = Counter(df.y.values)
dfa = pd.DataFrame.from_dict(letter_counts, orient='index')
dfa.plot(title='Histogram of the diferent classes', kind='bar')
sns_plot = sns.pairplot(df.loc[:, df.columns!='segment'], hue='y')
sns_plot.savefig('pair_futbol_power.png')
# # Feature extraction
df.columns
# # MODEL
df.dropna(inplace=True)
X_train = df.iloc[:, :-2]
y_train = df['y'].values
# %%time
from imblearn.combine import SMOTETomek
smt = SMOTETomek()
X_train, y_train = smt.fit_resample(X_train, y_train)
# +
from xgboost import XGBClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import AdaBoostClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
model = KNeighborsClassifier(n_neighbors=30, n_jobs=-1)
model.fit(X_train, y_train)
# -
test = pd.read_csv('player-2.csv')
del test['Unnamed: 0']
test.dropna(inplace=True)
X_test = test.iloc[:, :-2]
y_test = test['y']
# +
from sklearn.metrics import classification_report
y_pred_class = model.predict(X_test.values)
print(classification_report(y_test, y_pred_class))
# +
from sklearn.metrics import accuracy_score
accuracy_score(y_test, y_pred_class)
# -
from joblib import dump, load
dump(model, 'ml_model.joblib')
| notebooks/ML.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Point data manipulation with PDAL
#
# http://pdal.io
#
# PDAL (the Point Data Abstraction Library) is an open source library for handling massive point data sets. It's name is derived from GDAL - since it aims to sit in the same space for point data.
#
# PDAL is actually a C library - if you're writing applications you can insert it into your code. It also has python bindings. Today we'll explore some of PDAL's capabilities using it's command line applications - which are mostly wrappers to PDAL's pipeline functions.
#
# We'll also us a sneaky bit of LibLAS: http://www.liblas.org
#
# ...but you'll hopefully see why we'd prefer PDAL in the end.
# ### Agenda for this session
#
# A lightning speed overview of point data handling and manipulation:
# 1. Getting information about a point dataset
# 2. Collecting a subset from a LiDAR survey
# 3. Requesting only a specific point class from a dataset
# 4. Classifying ground (in case you don't like the vendor's version of 'ground')
# 5. Requesting 'height above ground' instead of 'absolute height'
# 6. Generating a bare earth DEM and a DSM
#
# We will do all this on the command line, viewing results in CloudCompare or this notebook. These tasks are based on the PDAL workshop here: http://www.pdal.io/workshop/index.html, and are very much 'learn by doing'. PDAL is very well documented, please keep reading for more information.
#
# ...so feel free to zoom ahead, create and share!
# ### Set up
# ```
# module purge
# module load PDAL cloudcompare```
# ### Locate data
#
# We will use a LiDAR survey obtained over Merimbula in 2013. Here is it's catalogue entry:
#
# *THREDDS*
# http://dapds00.nci.org.au/thredds/catalog/rr1/Elevation/Merimbula0313/catalog.html
#
# The path to the data via the VDI is:
#
# ```
# /g/data1/Elevation/Merimbula0313/Tiles_2k_2k```
#
# ### 1. Basic information
# Try:
#
# ```
# pdal info /g/data1/Elevation/Merimbula0313/Tiles_2k_2k/Merimbula2013-C3-AHD_7605910_55_0002_0002.las ```
#
# ...and compare with:
#
# ```
# lasinfo /g/data1/Elevation/Merimbula0313/Tiles_2k_2k/Merimbula2013-C3-AHD_7605910_55_0002_0002.las ```
#
# <div class="panel panel-info">
# <div class="panel-heading">Lasinfo gives more compact results - but can only read LAS. PDAL's info function can tell you about dimensions in any dataset it has a schema for reading: http://www.pdal.io/stages/readers.html, which hints also that PDAL can <i>process</i> point data in a diverse range of data formats.</div></div>
# ### 2. Clipping point data with PDAL
#
# Straight into the fire! We're going straight to PDAL's **pipeline** architecture, which gives it an enormous amount of felxibility and power. A pipeline is a set of operations chained together and defined in a JSON file. You'll see it in action here!
#
# ### a. Why do we want to clip LAS data?
#
# LAS tiles are pretty hard to handle - you get a lot of extra data that you may not want, and they are pretty much always boringly square. If we only need a certain region, we can get just those points using PDAL.
#
# ### b. An example - selecting Merimbula town
#
# The image here shows that Merimbula town covers several LIDAR tiles. This is an extra challenge - it means some tiles have a **lot** of data we don't want.
#
# (QGIS screenshot pic)
#
# You can see a polygon around a region of interest - it's saved as geoJSON and looks like this:
#
# ```
# {
# "type": "FeatureCollection",
# "crs": { "type": "name", "properties": { "name": "urn:ogc:def:crs:EPSG::28355" } },
# "features": [
# { "type": "Feature", "properties": { "id": 0 }, "geometry": { "type": "Polygon", "coordinates": [ [ [ 759094.480855233967304, 5913008.271593709476292 ], [ 758464.699909413931891, 5912716.199270982295275 ], [ 757743.646362751838751, 5912898.744472593069077 ], [ 757716.26458250079304, 5913304.907546310685575 ], [ 757373.992329337401316, 5913418.998297326266766 ], [ 757018.029186049010605, 5913724.761510098353028 ], [ 757556.537531022448093, 5913784.088700683787465 ], [ 757828.153738587978296, 5913997.946465536952019 ], [ 757828.153738587396219, 5914326.52782854065299 ], [ 758357.534823469701223, 5914381.291389083489776 ], [ 758877.788648267393, 5914554.709330711513758 ], [ 758850.406868015765212, 5914810.272613044828176 ], [ 759042.079329782165587, 5914837.654393311589956 ], [ 759151.606450793216936, 5914673.363711818121374 ], [ 759370.660692813224159, 5914709.872752171941102 ], [ 759361.533432727912441, 5915102.34493575617671 ], [ 760593.713544093072414, 5915138.853976195678115 ], [ 761177.858189482591115, 5915047.581375411711633 ], [ 761123.094628979102708, 5914235.255227984860539 ], [ 761260.003530243760906, 5914007.07372591085732 ], [ 761570.33037310524378, 5913952.31016543880105 ], [ 761369.530651255394332, 5913559.837981833145022 ], [ 761141.349149147979915, 5913459.438120897859335 ], [ 760484.186423085397109, 5913377.292780089192092 ], [ 759817.896436938317493, 5913632.856062367558479 ], [ 759516.696854161447845, 5913550.710721591487527 ], [ 759416.29699323582463, 5913286.020179163664579 ], [ 759094.480855233967304, 5913008.271593709476292 ] ] ] } }
# ]
# }
#
# ```
#
# ...but PDAL needs WKT - using this website: http://rodic.fr/blog/online-conversion-between-geometric-formats/, we can get a WKT polygon:
#
# ```
# POLYGON((759094.480855234 5913008.2715937095,758464.6999094139 5912716.199270982,757743.6463627518 5912898.744472593,757716.2645825008 5913304.907546311,757373.9923293374 5913418.998297326,757018.029186049 5913724.761510098,757556.5375310224 5913784.088700684,757828.153738588 5913997.946465537,757828.1537385874 5914326.527828541,758357.5348234697 5914381.2913890835,758877.7886482674 5914554.7093307115,758850.4068680158 5914810.272613045,759042.0793297822 5914837.654393312,759151.6064507932 5914673.363711818,759370.6606928132 5914709.872752172,759361.5334327279 5915102.344935756,760593.7135440931 5915138.853976196,761177.8581894826 5915047.581375412,761123.0946289791 5914235.255227985,761260.0035302438 5914007.073725911,761570.3303731052 5913952.310165439,761369.5306512554 5913559.837981833,761141.349149148 5913459.438120898,760484.1864230854 5913377.292780089,759817.8964369383 5913632.856062368,759516.6968541614 5913550.7107215915,759416.2969932358 5913286.020179164,759094.480855234 5913008.2715937095))
# ```
#
# ### c. Making a list of LIDAR tiles.
#
# We need to know which tiles contain our data. The tile index shapefile ( ) will help us to figure out which tiles we need - here they are:
# ```
# Merimbula2013-C3-AHD_7565916_55_0002_0002.las
# Merimbula2013-C3-AHD_7565914_55_0002_0002.las
# Merimbula2013-C3-AHD_7565912_55_0002_0002.las
# Merimbula2013-C3-AHD_7585916_55_0002_0002.las
# Merimbula2013-C3-AHD_7585914_55_0002_0002.las
# Merimbula2013-C3-AHD_7585912_55_0002_0002.las
# Merimbula2013-C3-AHD_7605916_55_0002_0002.las
# Merimbula2013-C3-AHD_7605914_55_0002_0002.las
# Merimbula2013-C3-AHD_7605912_55_0002_0002.las
# ```
#
# ### d. Constructing a PDAL pipeline
#
# We create a JSON file which tells PDAL what to do:
#
# ```
# nano merimbula_pipeline.json
# ```
# ..and paste in the following:
#
# ```
# {
# "pipeline": [
# { "filename": "../merimbula2013/Tiles_2k_2k/Merimbula2013-C3-AHD_7585912_55_0002_0002.las",
# "tag": "A"
# },
# { "filename": "../merimbula2013/Tiles_2k_2k/Merimbula2013-C3-AHD_7585914_55_0002_0002.las",
# "tag": "B"
# },
# { "filename": "../merimbula2013/Tiles_2k_2k/Merimbula2013-C3-AHD_7605914_55_0002_0002.las",
# "tag": "C"
# },
# {
# "inputs": ["A", "B", "C"],
# "type": "filters.crop",
# "polygon": "POLYGON((759094.480855234 5913008.2715937095,758464.6999094139 5912716.199270982,757743.6463627518 5912898.744472593,757716.2645825008 5913304.907546311,757373.9923293374 5913418.998297326,757018.029186049 5913724.761510098,757556.5375310224 5913784.088700684,757828.153738588 5913997.946465537,757828.1537385874 5914326.527828541,758357.5348234697 5914381.2913890835,758877.7886482674 5914554.7093307115,758850.4068680158 5914810.272613045,759042.0793297822 5914837.654393312,759151.6064507932 5914673.363711818,759370.6606928132 5914709.872752172,759361.5334327279 5915102.344935756,760593.7135440931 5915138.853976196,761177.8581894826 5915047.581375412,761123.0946289791 5914235.255227985,761260.0035302438 5914007.073725911,761570.3303731052 5913952.310165439,761369.5306512554 5913559.837981833,761141.349149148 5913459.438120898,760484.1864230854 5913377.292780089,759817.8964369383 5913632.856062368,759516.6968541614 5913550.7107215915,759416.2969932358 5913286.020179164,759094.480855234 5913008.2715937095))",
# "outside": false
# },
# "./merimbulatown.las"
# ]
# }
#
# ```
#
# ### e. Apply our clipping operation
#
# Then we execute the task using:
#
# ```
# pdal pipeline merimbula_pipeline.json
# ```
#
# This will result in a set of points inside your polygon being written into a .LAS file at the location specified in the pipeline file. Now you have a template for doing this job with pretty much any LAS tiles!
#
# In a new terminal, type:
#
# ```
# cloudcompare &
# ```
#
# ...and use it's file/open menu to navigate to your newly made LAS file. Take a look at it there (hint - use the projections menu to convert the Z dimension to a scalar field to colour your points by height).
#
# <div class="panel panel-warning">
# <div class="panel-heading"><h3>Caution</h3></div>
# <div class="panel-body">If your polygon is quite large, or your points very dense, or both, you can still get a massive dataset! Use pdal info to get an estimate of how dense the data are, and figure out how much area you are clipping to estimate the final file size before going ahead.</div></div>
#
| .ipynb_checkpoints/PDAL on the VDI-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Time
# https://docs.python.org/3/library/time.html?highlight=time#module-time
#
# This is the lowest level library for dealing with times. It is not platform agnostic, so some functions are not available on all operating systems and even if they do, it might not give the same result.
import time
# The first function of note from the time library is the `time.time()` function. It will return the time in seconds since the epoch as a floating point number. The epoch generally referrs to January 1, 1970, 00:00:00 (UTC). You can check the epoch time on your system with `time.gmtime(0)`. The `time.gmtime()` function will turn the seconds-since-epoch time into something more human readable.
time.time()
time.gmtime(0)
time.gmtime(time.time())
# As you can see, I was working on this in June of 2019. These are relatively simple time functions.
#
# Another common time function that I want to point out is the `time.sleep()` function. It will suspend execution of the calling thread for the given number of seconds. There are a number of uses for it, and it's fairly standard in programming.
time.sleep(1)
# Aside from getting the current time and sleeping, you might also want to be able to create time objects from a string or turn time objects back into human readable strings. You can use the `time.strptime()` function to parse string dates and create time objects from them. Conversly, you can use `time.strftime()` to turn time objects into human readable strings. For more details on parsing them, visit the python docs at the link above.
#
# This is a more complicated way to format the epoch time into a human readable format. It does, at least, allow you to specifiy how you want it formatted.
time.strftime("%a, %d %b %Y %H:%M:%S +0000", time.gmtime(0))
# The easier way to print a time in an easily human-readable format. It is a default function and works like this:
time.asctime(time.gmtime(0))
# Now, let's take a look at some common date and time formats and how we can convert them to time objects.
time_string = '2009-06-15T13:45:30'
parse_format = '%Y-%m-%dT%H:%M:%S'
stripped = time.strptime(time_string, parse_format)
time.asctime(stripped)
time_string = '6/15/2009'
parse_format = '%m/%d/%Y'
stripped = time.strptime(time_string, parse_format)
time.asctime(stripped)
time_string = 'Monday, June 15, 2009'
parse_format = '%A, %B %d, %Y'
stripped = time.strptime(time_string, parse_format)
time.asctime(stripped)
time_string = 'Mon, Jun 15, 2009 1:45 PM'
parse_format = '%a, %b %d, %Y %I:%M %p'
stripped = time.strptime(time_string, parse_format)
time.asctime(stripped)
time_string = 'Monday, June 15, 09 13:45:30'
parse_format = '%A, %B %d, %y %H:%M:%S'
stripped = time.strptime(time_string, parse_format)
time.asctime(stripped)
time_string = '6/15/09 1:45:30 PM'
parse_format = '%m/%d/%y %I:%M:%S %p'
stripped = time.strptime(time_string, parse_format)
time.asctime(stripped)
time_string = '6/15/2009 13:45'
parse_format = '%m/%d/%Y %H:%M'
stripped = time.strptime(time_string, parse_format)
time.asctime(stripped)
# Manipulating times using the time functions is not easy, so it is better to use the datetime library.
#
# # Datetime
# https://docs.python.org/3/library/datetime.html?highlight=time#module-datetime
#
# From the docs: The datetime module supplies classes for manipulating dates and times in both simple and complex ways. While date and time arithmetic is supported, the focus of the implementation is on efficient attribute extraction for output formatting and manipulation.
import datetime
# The datetime package provides methods for dealing with the date and time separately, datetime (both together), and timedelta differences between datetimes. If you want the current time, you can use `datetime.datetime.now()` to retrieve it. Let's look at the current time, and then both the \_\_str\_\_ and \_\_repr\_\_ for it:
current_time = datetime.datetime.now()
print('str: ',current_time, '\trepr:', current_time.__repr__())
# Now you can work with dates, times, or datetimes, but I'm going to stick with the full datetime object for now, since that is what I most commonly use.
#
# Datetime objects can be timezone aware, as an example, you can get the current UTC time:
datetime.datetime.utcnow()
# You can also convert a `time` object into a datetime
datetime.datetime.fromtimestamp(time.time())
# The `strptime()` function is virtually the same as with `time`
datetime.datetime.strptime('2009-06-15T13:45:30', '%Y-%m-%dT%H:%M:%S')
# With more than one datetime object you can do comparisons, and create timedelta objects that represent distances between the times:
time1 = datetime.datetime.strptime('2009-06-15T13:45:30', '%Y-%m-%dT%H:%M:%S')
time2 = datetime.datetime.now()
# Also of note, you can extract the individual pieces of a datetime separately of one another:
time1.year, time1.month, time1.day, time1.hour, time1.tzinfo
# A few time difference and comparison operations:
my_timedelta = time2 - time1
my_timedelta
time2 + my_timedelta
time1 - my_timedelta
time1 < time2
# # Calendar
# https://docs.python.org/3/library/calendar.html#module-calendar
#
# From the docs: This module allows you to output calendars like the Unix cal program, and provides additional useful functions related to the calendar.
#
# I've honestly never used this once before. I've just not had a use case that called for it. This is really just me playing around with a few of the methods to see what it does!
import calendar
my_calendar = calendar.Calendar()
print(my_calendar)
for monthdate in my_calendar.itermonthdates(2019, 3):
print(monthdate.strftime("%a, %d %b %Y"))
# If you read the documents, you see that months start on a monday, hence why this returned values prior to the start of the specified month.
| Notebooks/Time Datetime and Calendar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:Anaconda3]
# language: python
# name: conda-env-Anaconda3-py
# ---
import pandas as pd
import numpy as np
def sessions(sessions):
p = sessions.groupby('user_id').size().reset_index()
p = p.rename(columns={0:'total_sessions'})
t = sessions.groupby('user_id').secs_elapsed.mean().reset_index()
t = t.rename(columns = {'secs_elapsed':'mean'})
p = pd.merge(p,t, on='user_id',how='left')
t = sessions.groupby('user_id').secs_elapsed.median().reset_index()
t = t.rename(columns = {'secs_elapsed':'median'})
p = pd.merge(p,t, on='user_id',how='left')
t = sessions.groupby('user_id').secs_elapsed.std().reset_index()
t = t.rename(columns = {'secs_elapsed':'std_dev'})
p = pd.merge(p,t, on='user_id',how='left')
t = sessions.groupby('user_id').secs_elapsed.min().reset_index()
t = t.rename(columns = {'secs_elapsed':'minimum'})
p = pd.merge(p,t, on='user_id',how='left')
t = sessions.groupby('user_id').secs_elapsed.max().reset_index()
t = t.rename(columns = {'secs_elapsed':'maximum'})
p = pd.merge(p,t, on='user_id',how='left')
for f in ['action','action_type','device_type','action_detail']:
#,'action_type','device_type','action_detail']:
t = sessions.groupby(['user_id',f]).size().reset_index()
t = t.rename(columns={0:'counts'})
print('hohohohohohohohohoho')
t = t.pivot(index='user_id',columns = f ,values='counts').reset_index()
p = pd.merge(p,t,on='user_id',how='left')
z = sessions
z['action_action_type_action_detail'] = z['action'] + z['action_type'] + z['action_detail']
t = z.groupby(['user_id','action_action_type_action_detail']).size().reset_index()
t = t.pivot(index='user_id',columns='action_action_type_action_detail',values=0).reset_index()
p=pd.merge(p,t,on='user_id',how='left')
return p
def f(data):
data = pd.get_dummies(data, columns=['gender','signup_method','signup_flow','language','affiliate_channel','affiliate_provider','first_affiliate_tracked', 'signup_app','first_device_type','first_browser'], drop_first=False)
dates = data.date_account_created.apply(lambda x: x.split('-'))
print(type(dates))
dates = dates.values
print(dates)
dates = np.vstack(dates)
print(dates)
year = dates[:,0].astype(int)
month = dates[:,1].astype(int)
day = dates[:,2].astype(int)
print(year)
print(month)
print(day)
print(len(year))
print(len(month))
print(len(day))
print(len(data))
data['day'] = day
data['month'] = month
data['year'] = year
print(data['year'])
tfa = np.vstack(data.timestamp_first_active.astype(str).apply(lambda x: list(map(int, [x[:4],x[4:6],x[6:8],x[8:10],x[10:12],x[12:14]]))).values)
data['tfa_year'] = tfa[:,0]
data['tfa_month'] = tfa[:,1]
data['tfa_day'] = tfa[:,2]
data = data.drop(['date_account_created'], axis=1)
data = data.drop(['timestamp_first_active'],axis=1)
data = data.drop(['date_first_booking'], axis=1)
av = data.age.values
data['age'] = np.where(np.logical_or(av<14, av>100), -1, av)
data.fillna(-1)
return data
import pandas as pd
sess = pd.read_csv('sessions.csv')
users = pd.read_csv('train_users_2.csv')
test = pd.read_csv('test_users.csv')
users['tt'] = 'train'
test['tt'] = 'test'
tt = pd.concat([users,test],ignore_index=True)
(len(users),len(test),len(tt)-len(users))
# +
#len(tt.iloc[213451:,:])
# -
test.shape
tt.shape
tt.head()
file = f(tt)
sess = sessions(sess)
data = pd.merge(file,sess,left_on='id',right_on='user_id',how='left')
data.shape
train1 = data.iloc[0:150000,:]
train2 = data.iloc[150000:213451,:]
train = pd.concat([train1,train2])
len(train)
train
test = data[data['tt'] == 'test']
test.shape
train.head()
test.head()
trf = train[train['year'] >= 2014]
import numpy as np
import pandas as pd
from sklearn.ensemble import RandomForestClassifier
from sklearn.preprocessing import LabelEncoder
'''lef = LabelEncoder()
lef.fit(trf.country_destination.values)
fretar = lef.transform(trf.country_destination.values)'''
leo = LabelEncoder()
leo.fit(train.country_destination.values)
fulltar=leo.transform(train.country_destination.values)
fretar = leo.transform(trf.country_destination.values)
len(fretar),len(fulltar)
test
(test.shape,trf.shape,train.shape)
test.head()
train.head()
test = test.drop('country_destination',axis=1)
train = train.drop('country_destination',axis=1)
trf = trf.drop('country_destination',axis=1)
(test.shape,train.shape,trf.shape)
train = train.drop('id',axis=1)
test = test.drop('id',axis=1)
trf = trf.drop('id',axis=1)
trf= trf.drop('tt',axis=1)
train = train.drop('tt',axis=1)
test = test.drop('tt',axis=1)
train =train.fillna(-1)
test=test.fillna(-1)
trf=trf.fillna(-1)
#trf= trf.drop('tt','axis'=1)
import csv
def tocsv(file,name):
return file.to_csv(name+'.csv', index=False)
tocsv(predictions,'pred1')
# +
#tocsv(trf,'ffresh')
# +
#tocsv(test,'ftest')
# +
#kk=pd.read_csv('ftest.csv')
# -
'''print(len(kk))
kk.tail()'''
# +
#mm = pd.read_csv('ffresh.csv')
# +
#len(mm)
# -
(train.shape,test.shape,trf.shape)
from sklearn.ensemble import RandomForestClassifier
from sklearn.preprocessing import LabelEncoder
from xgboost.sklearn import XGBClassifier
def perform_prediction(training, labels, testing, xgb_votes, rf_votes):
""" Perform prediction using a combination of XGB and RandomForests. """
predictions = np.zeros((len(testing), len(set(labels))))
# Predictions using xgboost.
for i in range(xgb_votes):
print('XGB vote %d' % i)
xgb = XGBClassifier(
max_depth=DEPTH_XGB, learning_rate=LEARNING_XGB,
n_estimators=ESTIMATORS_XGB, objective='multi:softprob',
subsample=SUBSAMPLE_XGB, colsample_bytree=COLSAMPLE_XGB,silent=False)
xgb.fit(training, labels)
predictions += xgb.predict_proba(testing)
# Predictions using RandomForestClassifier.
for i in range(rf_votes):
print('RandomForest vote %d' % i)
rand_forest = RandomForestClassifier(
n_estimators=ESTIMATORS_RF, criterion=CRITERION_RF, n_jobs=JOBS_RF,
max_depth=DEPTH_RF, min_samples_leaf=MIN_LEAF_RF, bootstrap=True)
rand_forest.fit(training, labels)
predictions += rand_forest.predict_proba(testing)
return predictions
predictions = np.zeros((len(test), len(leo.classes_)))
(leo.classes_)
predictions.shape
# +
DEPTH_XGB, ESTIMATORS_XGB, LEARNING_XGB, SUBSAMPLE_XGB, COLSAMPLE_XGB = (
7, 60, 0.2, 0.7, 0.6) # XGBoost parameters.
ESTIMATORS_RF, CRITERION_RF, DEPTH_RF, MIN_LEAF_RF, JOBS_RF = (
500, 'gini', 20, 8, 30) # RandomForestClassifier parameters.
XGB_ALL_VOTE, RF_ALL_VOTE, XGB_FRESH_VOTE, RF_FRESH_VOTE = (5, 2, 10, 4)
# +
#train = train.iloc[:,:].values
# +
#trf = trf.values
# -
test = test.drop('user_id',axis=1)
train = train.drop('user_id',axis=1)
trf = trf.drop('user_id',axis=1)
# +
#predictions = predictions + perform_prediction(train,fulltar,test,XGB_FRESH_VOTE, RF_FRESH_VOTE)
# -
(train.shape, test.shape, trf.shape)
# +
#train.user_id
# -
#leo,lef,train,trf, test,fretar,fulltar
predictions += perform_prediction(trf, fretar, test, XGB_FRESH_VOTE, RF_FRESH_VOTE)
predictions
# +
#predictions.values.to_csv('preds1.csv',index=False)
# -
predictions += perform_prediction(train, fulltar, test, XGB_ALL_VOTE, RF_ALL_VOTE)
predictions
predictions[0]
test
test_df=pd.read_csv('test_users.csv')
len(test_df)
# +
ids, countries = ([], [])
for i in range(len(test_df)):
idx = test_df.index[i]
ids += [idx] * 5
countries += leo.inverse_transform(
np.argsort(predictions[i])[::-1])[:5].tolist()
# Save prediction in CSV file.
sub = pd.DataFrame(np.column_stack((ids, countries)), columns=['id', 'country'])
sub.to_csv('1234.csv', index=False)
# -
a = pd.read_csv('SUBMISSION_CSV.csv')
test_df.set_index('id', inplace=True)
id_test = test_df.index.values
id_test
# +
######################################################################submission generation#####################################
#Taking the 5 classes with highest probabilities
ids = [] #list of ids
cts = [] #list of countries
for i in range(len(id_test)):
idx = id_test[i]
ids += [idx] * 5
cts += leo.inverse_transform(np.argsort(predictions[i])[::-1])[:5].tolist()
#Generate submission
print("Outputting final results...")
sub = pd.DataFrame(np.column_stack((ids, cts)), columns=['id', 'country'])
sub.to_csv('./submission1234121212.csv', index=False)
# -
ids
predictions[0]
predictions = np.vstack(predictions)
predictions
p = pd.DataFrame(predictions)
p.to_csv('predictions.csv')
# +
ids, countries = ([], [])
for i in range(len(test_df)):
idx = test_df.index[i]
ids += [idx] * 5
countries += leo.inverse_transform(
np.argsort(predictions[i])[::-1])[:5].tolist()
# Save prediction in CSV file.
sub = pd.DataFrame(np.column_stack((ids, countries)), columns=['id', 'country'])
sub.to_csv('SUBMISSION11111111111111111111_CSV.csv', index=False)
# -
len(countries)==len(ids)
| airbnb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Random Forest
# 앙상블ensemble은 여러 머신러닝 모델을 연결하여 더 강력한 모델을 만드는 기법이다.
# 앞서 확인한 것처럼 결정 트리의 주요 단점은 모형이 불안정하고 훈련 데이터에 과대적합되는 경향이 있다는 것이다. 랜덤 포레스트는 이 문제를 회피할 수 있는 방법이다. 랜덤 포레스트는 기본적으로 조금씩 다른 여러 결정 트리의 묶음이다.
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.datasets import load_breast_cancer
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# +
cancer = load_breast_cancer()
X_train, X_test, y_train, y_test = train_test_split(cancer.data, cancer.target, random_state=0)
# -
# ### Random Forest 모형의 생성
#
# - 랜덤 포레스트 모델을 만들려면 생성할 트리의 개수를 정해야 한다.(RandomForestRegressor나 RandomForestClassifier의 n_estimators 매개변수).
# - 트리를 만들기 위해 먼저 데이터의 부트스트랩 샘플bootstrap sample을 생성한다.
# - 그다음 이렇게 만든 데이터셋으로 결정 트리를 만든다.
# - 결정 트리 생성시 특성을 랜덤하게 선택한다. 몇 개의 특성을 고를지는 max_features 매개변수로 조정한다.
# - max_features를 n_features로 설정하면 트리의 각 분기에서 모든 특성을 고려하므로 특성 선택에 무작위성이 들어가지 않는다.
# - max_features=1로 설정하면 트리의 분기는 테스트할 특성을 고를 필요가 없게 되며 그냥 무작위로 선택한 특성의 임계값을 찾기만 하면 된다.
#
# **Parameter to Tune**
# - n_jobs - training에 사용할 core의 갯수 (n_jobs=-1, for all cores)
# - n_estimators - 생성할 tree의 갯수 (more is always better)
# - max_depth, 가지치기 수준
# - max_features, for randomization , 기본값
# - max_features = sqrt(n_features), for classification
# - max_features = log2(n_features), for regression
# +
forest = RandomForestClassifier(n_estimators=100, random_state=0)
forest.fit(X_train, y_train)
print('Accuracy on the training subset: {:.3f}'.format(forest.score(X_train, y_train)))
print('Accuracy on the test subset: {:.3f}'.format(forest.score(X_test, y_test)))
# -
n_features = cancer.data.shape[1]
plt.figure(figsize=(10,8))
plt.barh(range(n_features), forest.feature_importances_, align='center')
plt.yticks(np.arange(n_features), cancer.feature_names)
plt.xlabel('Feature Importance')
plt.ylabel('Feature')
plt.show()
| 03Supervised/04DecisionTree_RandomForest/03RandomForest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:snowmodelcal]
# language: python
# name: conda-env-snowmodelcal-py
# ---
import numpy as np
import geopandas as gpd
import codecs
import requests
import rasterio as rio
# +
#########################################################################
############################ USER INPUTS ################################
#########################################################################
# DOMAIN
# choose the modeling domain
domain = 'CO_S'
# PATHS
# path to domain data folder
dataPath = '/nfs/attic/dfh/Aragon2/CSOdmn/'+domain+'/'
#########################################################################
# -
def build_snowmodel_line(domain,dataPath):
# path to DEM ascii
DEMpath = dataPath+'DEM_'+domain+'.asc'
#path to SNOTEL gdf
gdfpath = dataPath+'CSO_SNOTEL_sites_'+domain+'.geojson'
#path to VEG .asc
VEGpath = dataPath+'NLCD2016_'+domain+'.asc'
#path to lat .asc
LATpath = dataPath+'grid_lat_'+domain+'.asc'
#path to lon .asc
LONpath = dataPath+'grid_lon_'+domain+'.asc'
# VEG outfile path
outVEGpath = dataPath+'NLCD2016_'+domain+'_line.asc'
# DEM outfile path
outDEMpath = dataPath+'DEM_'+domain+'_line.asc'
# Line outfile path
outFpath = dataPath+'snowmodel_line_pts.dat'
#lon outfile path
outLONpath = dataPath+'grid_lon_'+domain+'_line.asc'
#lat outfile path
outLATpath = dataPath+'grid_lat_'+domain+'_line.asc'
# station data
stn_gdf = gpd.read_file(gdfpath)
#path to CSO domain
domains_resp = requests.get("https://raw.githubusercontent.com/snowmodel-tools/preprocess_python/master/CSO_domains.json")
domains = domains_resp.json()
# CSO projection
mod_proj = domains[domain]['mod_proj']
#get metadata from .asc
def getmeta(file_name):
myvars = {}
lines = open(file_name, 'r').readlines()
for i in range(5):
line=lines[i]
name, var = line.partition("\t")[::2]
myvars[name.strip()] = int(var)
return myvars
myvars = getmeta(DEMpath)
#Build northing easting array
#Northing
# locate center of lower left cell
st = myvars['yllcorner']+(myvars['cellsize']/2)
#build northing array
north = np.arange(st,st+myvars['nrows']*myvars['cellsize'],myvars['cellsize'])
#Easting
# locate center of lower left cell
st = myvars['xllcorner']+(myvars['cellsize']/2)
#build easting array
east = np.arange(st,st+myvars['ncols']*myvars['cellsize'],myvars['cellsize'])
#fortran indexing starts at 1
#staion 1 to N of N stations
count = np.zeros(stn_gdf.shape[0])
#easting of pixel corresponding to station
stn_est = np.zeros(stn_gdf.shape[0])
#northing of pixel corresponding to station
stn_nor = np.zeros(stn_gdf.shape[0])
#index of pixel corresponding to station
est_idx = np.zeros(stn_gdf.shape[0])
#index of pixel corresponding to station
nor_idx = np.zeros(stn_gdf.shape[0])
for z in range(stn_gdf.shape[0]):
count[z] = z + 1
lons = abs(stn_gdf.easting[z]-east)
loIDX = [i for i, value in enumerate(lons) if value == np.min(abs(stn_gdf.easting[z]-east))]
stn_est[z] = east[loIDX[0]]
est_idx[z] = loIDX[0] + 1
lats = abs(stn_gdf.northing[z]-north)
laIDX = [i for i, value in enumerate(lats) if value == np.min(abs(stn_gdf.northing[z]-north))]
stn_nor[z] = north[laIDX[0]]
nor_idx[z] = laIDX[0] + 1
#Print out .dat file
f= open(outFpath,"w+")
for z in range(count.shape[0]):
f.write('{:08.0f}\t'.format(count[z])+'{:10.0f}\t'.format(est_idx[z])+'{:10.0f}\t'.format(nor_idx[z])+
'{:10.0f}\t'.format(stn_est[z])+'{:10.0f}\t\n'.format(stn_nor[z]))
f.close()
## Extract topo, veg, lat, and lon files for snowmodel_line
new = stn_gdf.to_crs(mod_proj)
with rio.open(DEMpath) as src:
rows, cols = rio.transform.rowcol(src.transform, new.geometry.centroid.x, new.geometry.centroid.y)
# function to extract .asc files at SNOTEL locations
def field2line(filepath,outpath):
with codecs.open(filepath, encoding='utf-8-sig') as f:
data = np.loadtxt(f,skiprows=6)
data_line=[]
for i in range(stn_gdf.shape[0]):
info = str(int(data[rows[i],cols[i]]))
data_line.append(info)
lines = open(filepath, 'r').readlines()
head = 'ncols\t1\nnrows\t'+str(stn_gdf.shape[0])+'\n'+lines[2]+lines[3]+lines[4]+lines[5]
data = ''
for s in data_line:
data += s +'\n'
f= open(outpath,"w+")
f.write(head+data)
f.close()
#extract DEM
field2line(DEMpath,outDEMpath)
#extract VEG
field2line(VEGpath,outVEGpath)
#extract LAT
field2line(LATpath,outLATpath)
#extract LON
field2line(LONpath,outLONpath)
build_snowmodel_line(domain,dataPath)
| 05_Build_SnowModel_line_file.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.1 32-bit
# language: python
# name: python38132bit01cfdaa7b6474f02875364eb1a0937bc
# ---
# # EDA with Python and applying Logistic Regression
# - For this lecture we will be working with the [Titanic dataset from Kaggle ](https://www.kaggle.com/c/titanic) . This is a very famous data set and very often is a student's first step in machine learning!
# - We'll be trying to predict a classification- survival or deceased.
# Let's begin our understanding of implementing Logistic Regression in Python for classification.
# - We'll use a \"semi-cleaned\" version of the titanic data set, if you use the data set hosted directly on Kaggle, you may need to do some additional cleaning not shown in this lecture notebook.
import numpy as np #importing the libraries
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
train=pd.read_csv("titanic_train.csv") #loading the dataset
train.head(7)
# ## Exploratory Data Analysis
#
# ### Step1: Missing data
train.isnull()
sns.heatmap(train.isnull(),yticklabels=False, cmap="afmhot") #create a heatmap using seaborn by loding the missing values dataset.
# Observation:
# - There are only two columns that contain the missing data ("Age" and "Cabin").We can see clearly that, "Cabin" has more missing data than "Age". With this, it's unable to do any basic level operation. We'll drop this later, or change it another feature like "Cabin known:1 or 0"
# - Let's visualize the data which is available.
sns.set_style("whitegrid") #Countplot is used to find range of count of values in a particular column or feature.
sns.countplot(x="Survived", data=train)
sns.set_style("whitegrid") #Countplot is used to find range of count of values in a particular column or feature.
sns.countplot(x="Survived", hue="Sex", data=train)
sns.set_style("whitegrid") #Countplot is used to find range of count of values in a particular column or feature.
sns.countplot(x="Survived", hue="Pclass", data=train)
sns.set_style("whitegrid") #Countplot is used to find range of count of values in a particular column or feature.
sns.countplot(x="Survived", hue="Embarked", data=train)
sns.set_style("whitegrid") #Distplot is used to see distribution of values in a particular column or feature.
sns.distplot(train["Age"].dropna(), kde=False, color="green")
sns.set_style("whitegrid")
sns.countplot(train["SibSp"]) #Countplot is used to find range of count of values in a particular column or feature.
sns.distplot(train["Fare"].dropna(),kde=False, color="green")#Distplot is used to see distribution of values in a particular column or feature.
# or train["Fare"].hist(color="green",bins=40) (this is another syntax for dsiplaying histogram for a feature or column)
# ## Data Cleaning
# We want to fill in the missing data of age column instead of dropping the entire age data rows.Oen way to do this filling the age by mean age of all passengers(imputation).Check the average age by passenger class.
plt.figure(figsize=(12,7)) #To find the average age of the people fall in types of Pclass
sns.boxplot(x="Pclass",y="Age", data=train, palette="winter")
# Obeservation:
# - we can see that old people tend to be in the class-1 and average age is around 37
# - we can also see that Classes (2 and 3) are having young people where class-2 averaging around 29 and class-3 averaging around 24.
# - So based on this, we have to fill the missing data with the average age values.
# +
def impute_age(cols): #Created a fucntion to fill the avergae age data in the missing values.
Age=cols[0]
Pclass=cols[1]
if pd.isnull(Age):
if (Pclass==1):
return 37
elif (Pclass==2):
return 37
else:
return 24
else:
return Age
# -
train['Age']=train[['Age','Pclass']].apply(impute_age,axis=1) #with the apply function we apply it to the columns
sns.heatmap(data=train.isnull(),yticklabels=False)
train.drop(columns='Cabin',inplace=True) #Droppping the cabin column because lot of feature eng' should be done.
train.head()
train.drop(columns="PassengerId",inplace=True) #dropping the unnecessary columns for the operations.
train.drop(columns='Ticket', inplace=True) #dropping the unnecessary columns for the operations.
# +
train.head()
# -
# #### NOTE:
# - We dropped the unnecessary columns, Now we have get dummies for the features(or columns) because it helps in getting the quick result after analyzing the data.
pclass=pd.get_dummies(train["Pclass"],drop_first=True) #Getting dummies for Pclass, Sex, Embarked
sex=pd.get_dummies(train["Sex"],drop_first=True)
embarked=pd.get_dummies(train["Embarked"],drop_first=True)
train.head()
train.drop(['Name','Sex','Embarked','Pclass'],axis=1, inplace=True)
train.head()
train=pd.concat([train,pclass,sex,embarked],axis=1)
train.head() # With this feature engineering for this dataset is done, we got the pefect and cleaned dataset to build a model.
# ## Building a Logistic Regression model
# - There are two types of datasets:
# - Train(Which is we used for feature engineering i.e cleaning, polishing the data)
# - Test (which is another file with which you can play or experiment for building the model and also for not ruining the train data)
# - we have to split our data into a training set and test set.
#
x_train=train.drop("Survived", axis=1)#Dropping the "Survived" column because it is a dependent feature.
y_train=train["Survived"]#Dependent feature.
from sklearn.model_selection import train_test_split #sklearn contains all the necessary libraries for testing and building the model.
X_train, X_test, Y_train, Y_test=train_test_split(x_train, y_train ,test_size=0.30,random_state=101) # inputting our test datasets
# ## Training and Predicting
from sklearn.linear_model import LogisticRegression
logmodel=LogisticRegression()
logmodel.fit(X_train, Y_train)
predictions=logmodel.predict(X_test)
from sklearn.metrics import confusion_matrix
cmatrix=confusion_matrix(Y_test,predictions)
cmatrix
from sklearn.metrics import accuracy_score
accuracy=accuracy_score(y_test,predictions)
accuracy
predictions
import matpllotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
| Exploratory Data Analysis (EDA) Practise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from funcionario import lagrange
# # Excesso de dados
#
# Já vimos que usar pontos demais para interpolar pode trazer problemas:
# ao usar um polinômio de grau muito grande, este pode oscilar demasiadamente;
# ao considerar muitos dados com um pouco de erro, este pode ser magnificado pela interpolação.
#
# Entretanto, possuir mais informação deveria nos dar **mais** entendimento sobre o nosso problema, não?
# Vamos, aqui, tentar aproveitar estes dados suplementares de outra forma.
# # Regressão linear
#
# O primeiro exemplo desta técnica é a regressão linear.
# Possuimos dados $(x_i,y_i)$ que estão "quase" alinhados.
# Se desejássemos passar um polinômio por todos eles, entretanto, seríamos forçados a ter algo extremamente oscilante.
# Mas podemos tentar achar uma reta que esteja "suficientemente perto" de todos (ou _a mais perto_ de todos).
np.random.seed(1)
a,b = np.random.rand(2)
xs = np.linspace(-1,1,30)
ys = a*xs + b + 0.1*np.random.rand(30)
a, b
# ## Exercício, em muitas partes:
#
# ### 1) Veja os dados
### Resposta aqui
plt.plot(xs,ys,'.')
plt.show()
# ### 2) Faça um gráfico do ajuste linear
# Estude a função polyfit
v = np.polyfit(xs,ys,deg=1)
### Resposta aqui
def poly(x,v):
poly = 0
vinv = v[::-1]
for n,coef in enumerate(vinv):
poly += coef*x**n
return poly
# +
# Agora, faça o gráfico do ajuste, e do erro
_, [ax1,ax2] = plt.subplots(ncols=2, figsize=(13,4))
### Resposta aqui
ax1.plot(xs,poly(xs,v))
ax1.plot(xs,ys,'.')
ax2.plot(xs,poly(xs,v)-ys,'.')
ax2.axhline(color='k', linewidth=1, linestyle='--')
ax2.set_title('Erro')
plt.show()
# -
# ### 3) Compare com o polinômio interpolador de grau 29
pol_int = lagrange(xs,ys)
# +
ts = np.linspace(-1,1,200)
_, [ax1, ax2] = plt.subplots(ncols=2, figsize=(13,4))
### Resposta aqui
ax1.plot(xs,ys,'.',label = 'Dados')
ax1.plot(ts,pol_int(ts),label = 'Polinomio grau 29')
ax1.plot(ts,poly(ts,v),label = 'Reta')
ax1.set_yscale('symlog')
ax1.legend()
ax2.plot(xs,ys,'.',label = 'Dados')
ax2.plot(ts,pol_int(ts),label = 'Polinomio grau 29')
ax2.plot(ts,poly(ts,v),label = 'Reta')
ax2.set_yscale('symlog')
ax2.set_ylim(-1/2,2)
ax2.legend()
plt.show()
# -
# # Regressão polinomial
# Podemos generalizar o procedimento que fizemos, e tentar ajustar um polinômio de grau mais alto.
# Neste caso, não será provavelmente muito útil, mas vejamos o que acontece!
# Estude a função polyfit e polyval
coefs = np.polyfit(xs,ys,deg=4)
my_poly = np.polyval(coefs, xs)
coefs
# ### 4) Faça novamente um gráfico comparativo!
# +
ts = np.linspace(-1,1,200)
_, [ax1, ax2, ax3] = plt.subplots(ncols=2, figsize=(13,4))
### Resposta aqui
ax1.plot(xs,ys,'.',label = 'Dados')
ax1.plot(ts,pol_int(ts),label = 'Polinomio grau 29')
ax1.plot(ts,poly(ts,v),label = 'Reta')
ax1.set_yscale('symlog')
ax1.legend()
ax2.plot(xs,ys,'.',label = 'Dados')
ax2.plot(ts,pol_int(ts),label = 'Polinomio grau 29')
ax2.plot(ts,poly(ts,v),label = 'Reta')
ax2.set_yscale('symlog')
ax2.set_ylim(-1/2,2)
ax2.legend()
plt.show()
# +
ts = np.linspace(-2,2,200)
_, [ax1, ax2] = plt.subplots(ncols=2, figsize=(13,4))
### Resposta aqui
ax1.plot(xs,ys,'.',label = 'Dados')
ax1.plot(ts,pol_int(ts),label = 'Polinomio grau 29')
ax1.plot(ts,poly(ts,v),label = 'Reta')
ax1.set_yscale('symlog')
ax1.legend()
ax2.plot(xs,ys,'.',label = 'Dados')
ax2.plot(ts,pol_int(ts),label = 'Polinomio grau 29')
ax2.plot(ts,poly(ts,v),label = 'Reta')
ax2.set_yscale('symlog')
ax2.set_ylim(-1/2,2)
ax2.legend()
plt.show()
# -
# # Modelos lineares gerais
#
# O que está por trás da função `polyfit` é que temos um modelo que é _linear_ nos parâmetros que desejamos encontrar.
# Um modelo pode ser pensado numa função $M_{par}(x)$, que dá uma aproximação para os pontos $(x_i, y_i)$.
# Em geral, emprega-se uma notação como $y \sim M_{par}(x)$.
#
# Por exemplo, se temos um modelo polinomial, os parâmetros são os coeficientes $p_i$ do polinômio,
# e portanto $M_{par}(x) = \sum_{i=0}^d p_i x^i$.
#
# Quando o modelo é linear nos parâmetros, ao escrevermos as 30 equações $M_{par}(x_k) = y_k$,
# obtemos um sistema linear $A \cdot par = y$.
# Se temos tantos parâmetros como equações (ou mais!), é razoável esperar que o sistema terá solução.
# Mas, nos casos acima, estamos na situação exatamente oposta:
# temos (muito!) menos variáveis do que equações,
# e o sistema será, muito provavelmente, impossível.
#
# Entretanto, podemos tentar encontrar coeficientes que sejam os "melhores possível"
# segundo algum critério.
# Um critério bastante comum e prático (ou seja, rápido para o computador executar)
# é o de mínimos quadrados:
#
# $$ \text{tomaremos $c$ tal que } \left\| Ac - y \right\| \text{ seja o menor possível.} $$
# Para minimizar $\left\| Ac - y \right\|$, vamos minimizar o quadrado, o que torna o problema mais fácil, já que:
#
# $$ \left\| Ac - y \right\|^2 = \langle Ac - y, Ac - y \rangle = c^T (A^T A) c - 2 c^T Ay + y^T y. $$
#
# Derivando em relação a $c$, obtemos as chamadas equações normais:
#
# $$ A^T A c = A^T y,$$
#
# que agora formam um sistema com tantas equações quanto a dimensão de $c$.
# ## Exemplo: um modelo misto
#
# Vamos criar "dados artificiais" para um modelo contendo tanto uma componente senoidal como uma exponencial e uma constante.
# Depois, veremos como uma abordagem de mínimos quadrados se comporta.
# Modelo artificial: $y \sim A \sin(2\pi t) + Be^{\lambda(t-1950)} + C$, com $\lambda = 0.03$.
# parâmetros
A = 20
B = 10
C = 150
l = 0.03
# dados artificiais
ts = np.arange(1950,2010,1/12)
ys = A*np.sin(2*np.pi*ts) + B*np.exp(l*(ts-1950)) + C
plt.figure(figsize=(13,4))
plt.plot(ts, ys, '.-')
plt.show()
# ### Exercício
#
# Faça uma função que monta a matriz correspondente ao modelo.
def model_M(ts):
# Retorna a matriz do sistema linear
### Resposta aqui
def f(x): return np.sin(2*np.pi*x)
def g(x): return np.exp(0.03*(x-1950))
def h(x): return 1
functions = [f,g,h]
dim = len(ts)
Matriz = []
for i in range(dim):
linha = [f(ts[i]) for f in functions]
Matriz.append(linha)
return np.array(Matriz)
M = model_M(ts)
M.shape
# E agora, encontre os parâmetros da série!
# +
def model_par(ts,ys):
# Calcula os parâmetros para os dados (t_i, y_i), resolvendo o sistema linear
M = model_M(ts)
#frst = np.dot(M.T,M)
#scnd = M.T*ys
#ret = np.linalg.solve(frst,scnd)
#ret = np.linalg.lstsq(M.T,ys)
#ys = np.array([ys])
#ys.shape()
frst = M.T @ M
#scnd = np.dot(M.T,ys)
ret = np.linalg.lstsq(M,ys)
return ret[0]
model_par(ts,ys)
# -
model_par(ts, ys)
# # Dados reais!
#
# Dados de emissão de $CO_2$, medidos em Mauna Loa.
dados = np.load('mounaloa.npz')
ts, ys = dados['ts'], dados['ys']
from matplotlib.mlab import movavg
ts_avg = movavg(ts,12)
ys_avg = movavg(ys,12)
plt.figure(figsize=(13,4))
plt.plot(ts, ys, '.-')
plt.plot(ts_avg, ys_avg)
plt.show()
plt.plot(np.diff(ys_avg))
plt.show()
# ### Um modelo intermediário
#
# Aqui, precisaríamos também estimar o $\lambda$.
# Isso é relativamente complicado, e vamos começar aproximando a exponencial por um polinômio de grau 1.
# Isso dá o seguinte modelo:
#
# $$y \sim B\sin(2\pi t) + A(t-1959) + C. $$
# +
def model_M(ts):
# Monte a matriz
### Resposta aqui
def model_par(ts,ys):
M = model_M(ts)
MMt = np.matmul(M,M.T)
return np.linalg.solve(MMt, np.dot(M,ys))
# -
def model_fun(par):
B,A,C = par
def mf(t):
# Função que avalia o modelo no tempo t
### Resposta aqui
return mf
par = model_par(ts, ys)
mf = model_fun(par)
print(par)
# +
plt.figure(figsize=(13,4))
plt.plot(ts, ys, 'x', label='data')
plt.plot(ts, mf(ts), label='fit')
plt.legend()
plt.show()
# +
plt.figure(figsize=(13,4))
plt.plot(ts, ys - mf(ts))
plt.title('Unexplained')
plt.show()
# -
# ### Um modelo com mais parâmetros:
#
# Vamos tentar ajustar melhor:
# - introduzindo um termo $\cos(2\pi t)$,
# - e aproximando a exponencial por um polinômio de grau 2.
#
# Fica melhor?
# +
def model_M(ts):
# Monte a matriz
### Resposta aqui
def model_par(ts,ys):
M = model_M(ts)
MMt = np.matmul(M,M.T)
return np.linalg.solve(MMt, np.dot(M,ys))
# -
def model_fun(par):
B,B_,A,A_,C = par
def mf(t):
# Função que avalia o modelo no tempo t
### Resposta aqui
return mf
par = model_par(ts, ys)
mf = model_fun(par)
print(par)
# +
plt.figure(figsize=(13,4))
plt.plot(ts, ys, 'x', label='data')
plt.plot(ts, mf(ts), label='fit')
plt.legend()
plt.show()
# +
plt.figure(figsize=(13,4))
plt.plot(ts, ys - mf(ts))
plt.title('Unexplained')
plt.show()
| comp-cientifica-I-2018-2/semana-10/Semana10-Parte4-Regressao-LstSq.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Elastix
#
# This notebooks show very basic image registration examples with on-the-fly generated binary images.
from itk import itkElastixRegistrationMethodPython
from itk import itkTransformixFilterPython
import itk
import numpy as np
import matplotlib.pyplot as plt
# ## Image generators
def image_generator(x1, x2, y1, y2, upsampled=False, bspline=False,
mask=False, artefact=False):
if upsampled:
image = np.zeros([1000, 1000], np.float32)
elif mask:
image = np.zeros([100, 100], np.uint8)
else:
image = np.zeros([100, 100], np.float32)
for x in range(x1, x2):
for y in range(y1, y2):
if bspline:
y += x
if x > 99 or y > 99:
pass
else:
image[x, y] = 1
else:
image[x, y] = 1
if artefact:
image[:, -10:] = 1
image = itk.image_view_from_array(image)
return image
# ## Size transformation test
# See example 12 for more explanation
# +
# Create small images for registration
fixed_image_small = image_generator(25,75,25,75)
fixed_image_small.SetSpacing([10,10])
moving_image_small = image_generator(0,50,10,60)
moving_image_small.SetSpacing([10,10])
# .. and a big moving image for transformation
moving_image_large = image_generator(0,500,100,600, upsampled=True)
# Import Default Parameter Map
parameter_object = itk.ParameterObject.New()
default_rigid_parameter_map = parameter_object.GetDefaultParameterMap('rigid',4)
parameter_object.AddParameterMap(default_rigid_parameter_map)
# Call elastix
result_image_small, result_transform_parameters = itk.elastix_registration_method(
fixed_image_small, moving_image_small,
parameter_object=parameter_object)
# Adjust parameter file with spacing and size of large image.
result_transform_parameters.SetParameter("Size", ['1000', '1000'])
result_transform_parameters.SetParameter("Spacing", ['1', '1'])
# Call transformix with procedural method
# Procedural method gives wrong result image
# result_image_large = itk.transformix_filter(
# moving_image=moving_image_large,
# transform_parameter_object=result_transform_parameters,
# log_to_console=True)
# Load Transformix Object
transformix_object = itk.TransformixFilter.New()
transformix_object.SetMovingImage(moving_image_large)
transformix_object.SetTransformParameterObject(result_transform_parameters)
# Update object (required)
transformix_object.UpdateLargestPossibleRegion()
# Results of Transformation
result_image_large = transformix_object.GetOutput()
# -
# ## Size transformation test visualization
#
# +
# %matplotlib inline
# Plot images
fig, axs = plt.subplots(1,3, sharey=True, figsize=[30,30])
plt.figsize=[100,100]
axs[0].imshow(result_image_small)
axs[0].set_title('Result Small', fontsize=30)
axs[1].imshow(fixed_image_small)
axs[1].set_title('Fixed Small', fontsize=30)
axs[2].imshow(moving_image_small)
axs[2].set_title('Moving Small', fontsize=30)
plt.show()
# -
fig, axs = plt.subplots(1,2, figsize=[30,30])
plt.figsize=[100,100]
axs[0].imshow(result_image_large)
axs[0].set_title('Result Large', fontsize=30)
axs[1].imshow(moving_image_large)
axs[1].set_title('Moving Large', fontsize=30)
plt.show()
| examples/ITK_UnitTestExample7_SizeTransformation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
#
# # Race classification
#
# <NAME> and <NAME> initially wrote this notebook. <NAME> reviwed the notebook, edited the markdown, and reproduced, commented on and made minor changes in the code.
#
# Racial demographic dialect predictions were made by the model developed by [<NAME>., <NAME>., & <NAME>. (2016)](https://arxiv.org/pdf/1608.08868.pdf). We modified their predict function in [the public Git repository](https://github.com/slanglab/twitteraae) to work in the notebook environment.
#
# Code has been changed to Python3 by <NAME>, <NAME> and <NAME> from intital Python 2 commitment.
# +
# Install uninstalled libs
import sys
# Import libraries
# #!pip3 install seaborn
import pandas as pd
import numpy as np
import re
import seaborn as sns
import matplotlib.pyplot as plt
## Language-demography model
import predict
## Custom functions
from clean_text import clean_tweet
# -
# ### Import tweets
# +
# Import file
tweets = pd.read_csv('./hatespeech_text_label_vote_RESTRICTED_100K.csv', sep='\t', header=None)
# Index variable
tweets.index.name = 'ID'
# Name columns
tweets.columns = ['Tweet', 'label', 'votes']
# First five rows
tweets.head()
# -
# ### Clean tweets
# +
# Clean text
tweets_clean = tweets.copy()
tweets_clean['Tweet'] = clean_tweet(tweets_clean['Tweet'])
# Convert string into unicode
tweets_clean['Tweet'] = tweets_clean['Tweet'] # Applied unicode for compatability with model
# First five rows
tweets_clean.head()
# -
# ### Apply predictions
# +
# Predict function from the language-demography model
predict.load_model()
def prediction(string):
return predict.predict(string.split())
# +
# Make predictions
predictions = tweets_clean['Tweet'].apply(prediction)
# +
# Save prediction results to a new column
tweets_clean['pred'] = predictions
# Fill tweets that have no predictions with NAs (NULL values)
tweets_clean = tweets_clean.fillna("NA")
# First five rows
tweets_clean.head()
# -
# - AAE: [African-American English](https://en.wikipedia.org/wiki/African-American_English)
# - WAE: White-aligned English
# +
def first_last(item):
if item is 'NA':
return 'NA'
return np.array([item[0], item[3]]) # item[0] = AAE, item[3] = WAE
tweets_clean['pred_aae_wae'] = tweets_clean['pred'].apply(first_last)
tweets_clean.head()
# +
# Predicting racial demographic information using a binary category (whites or blacks)
def detect_two(item):
if item is 'NA':
return None
if item[0] > item[1]:
return 0
else:
return 1
# Predicting racial demographic information using a multiclass category (whites, blacks, and others; Others indicate Asian Americans and Latinx)
def detect_all(item):
if item is "NA":
return None
if item[0] > item[1] and item[0] > item[2] and item[0] > item[3]:
return 0
elif item[3] > item[0] and item[3] > item[1] and item[3] > item[2]:
return 1
else:
return 2
# Same as Model2 except saving only AAE values as a constinuous variable
def detect_aae_cont(item):
if item is "NA":
return None
if item[0] > item[1] and item[0] > item[2] and item[0] > item[3]:
return item[0]
else:
return None
# Same as Model2 except saving only WAE values as a constinuous variable
def detect_wae_cont(item):
if item is "NA":
return None
if item[3] > item[0] and item[3] > item[1] and item[3] > item[2]:
return item[3]
else:
return None
# +
# Apply functions to the data
tweets_clean['race_bi'] = tweets_clean['pred_aae_wae'].apply(detect_two)
tweets_clean['race_all'] = tweets_clean['pred'].apply(detect_all)
tweets_clean['aae_cont'] = tweets_clean['pred'].apply(detect_aae_cont)
tweets_clean['wae_cont'] = tweets_clean['pred'].apply(detect_wae_cont)
# +
# Check
tweets_clean['aae_cont'].head()
# +
# Drop columns
final_tweets = tweets_clean.drop(columns=["pred", "pred_aae_wae"])
final_tweets['Tweet'] = tweets['Tweet']
final_tweets.head()
# -
# ### Export tweets to CSV
final_tweets.columns
final_tweets.to_csv('./race_predictions.csv', sep=',', encoding='utf-8',
header=["text", "label", "votes", "race_bi", "race_all", "aae_count", "wae_count"], index=True)
| code/race_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Kmb6Q6Z_Golu" colab_type="text"
# https://colab.research.google.com/drive/1ATDXbpOfeZik2bV4kBLz82aI1pKQlh0x
# + id="nLcfIezQELLU" colab_type="code" outputId="e95f9c93-c8d6-4958-83b9-7fb72f4dccb7" colab={"base_uri": "https://localhost:8080/", "height": 204}
# !wget https://d1p17r2m4rzlbo.cloudfront.net/wp-content/uploads/2017/04/a943287.csv
# + id="-ygQ0l5nENFQ" colab_type="code" colab={}
import pandas as pd, numpy as np
from skimage import io
# + id="_AT0as1FEONg" colab_type="code" outputId="394a7fc2-de4c-4e89-f7e9-bb381c92a0ad" colab={"base_uri": "https://localhost:8080/", "height": 224}
data = pd.read_csv('a943287.csv')
data.head()
# + id="8APQpw2fEPdp" colab_type="code" colab={}
data_male = data[data['please_select_the_gender_of_the_person_in_the_picture']=="male"].reset_index(drop='index')
data_female = data[data['please_select_the_gender_of_the_person_in_the_picture']=="female"].reset_index(drop='index')
# + id="Pafd6yBWEUCd" colab_type="code" outputId="5ec5eb21-c1d1-49c8-86d6-b85fc4a2c51e" colab={"base_uri": "https://localhost:8080/", "height": 34}
final_data = pd.concat([data_male[:1000],data_female[:1000]],axis=0).reset_index(drop='index')
final_data.shape
# + [markdown] id="Z83GjkeiUg-X" colab_type="text"
# # Scenario 1: Big images
# + id="escjW4KfEVmZ" colab_type="code" outputId="0f0a9d1a-afa2-4890-c9fc-d5aaf188d64b" colab={"base_uri": "https://localhost:8080/", "height": 357}
x = []
y = []
for i in range(final_data.shape[0]):
if(i%100==0):
print(i)
try:
image = io.imread(final_data.loc[i]['image_url'])
if(image.shape==(300,300,3)):
x.append(image)
y.append(final_data.loc[i]['please_select_the_gender_of_the_person_in_the_picture'])
except:
continue
# + id="24Ty9_ZtEXbA" colab_type="code" colab={}
x2 = []
y2 = []
for i in range(len(x)):
x2.append(x[i])
img_label = np.where(y[i]=="male",1,0)
y2.append(img_label)
# + id="6rsNV3gPHu_3" colab_type="code" outputId="50a7770e-e18c-4f6a-c9e6-3f62c21d3adf" colab={"base_uri": "https://localhost:8080/", "height": 34}
from keras.layers import Conv2D, MaxPooling2D, Flatten, Dense, Dropout
from keras.models import Sequential
# + id="XhPCiHJGH8OR" colab_type="code" colab={}
x2 = np.array(x2)
# + id="mCKvifQ9H9rj" colab_type="code" colab={}
x2 = x2.reshape(x2.shape[0],x2.shape[1],x2.shape[2],3)
# + id="FEGZeo1IH_Km" colab_type="code" outputId="e09b7958-b46d-43bd-e5fd-8d64ce935107" colab={"base_uri": "https://localhost:8080/", "height": 493}
model = Sequential()
model.add(Conv2D(64, kernel_size=(3, 3), activation='relu',input_shape=(300,300,3)))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(128, kernel_size=(3, 3), activation='relu',padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(256, kernel_size=(3, 3), activation='relu',padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(512, kernel_size=(3, 3), activation='relu',padding='same'))
model.add(Flatten())
model.add(Dense(100, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.summary()
# + id="tYCvMx70ICjO" colab_type="code" outputId="8d7cd474-4c3d-48a2-fb70-8541b6e7e72a" colab={"base_uri": "https://localhost:8080/", "height": 122}
model.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
# + id="X6SdyKEMIEDS" colab_type="code" colab={}
X = np.array(x2)/255
Y = np.array(y2)
# + id="1GG-cDsKIF5v" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,Y, test_size=0.1, random_state=42)
# + id="G59JT642IHbN" colab_type="code" outputId="8d944601-80ab-4f59-eb84-543e0db0f1e4" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(X_train.shape, X_test.shape, y_train.shape, y_test.shape)
# + id="LwtNucF5IxPC" colab_type="code" outputId="d4cde552-2f32-4ed4-8aa1-3f78b225bfbe" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.max(X_train)
# + id="qppUR5TeIIy-" colab_type="code" outputId="004a00a6-7026-4c34-f9a0-dc3f009e51f2" colab={"base_uri": "https://localhost:8080/", "height": 374}
history = model.fit(X_train, y_train, batch_size=32,epochs=10,verbose=1,validation_data = (X_test, y_test))
# + id="No1U3fbeILXK" colab_type="code" outputId="5d96b9f1-6c0b-4089-9b71-d7d103face3a" colab={"base_uri": "https://localhost:8080/", "height": 388}
import matplotlib.pyplot as plt
# %matplotlib inline
history_dict = history.history
loss_values = history_dict['loss']
val_loss_values = history_dict['val_loss']
acc_values = history_dict['acc']
val_acc_values = history_dict['val_acc']
epochs = range(1, len(val_loss_values) + 1)
plt.subplot(211)
plt.plot(epochs, history.history['loss'], 'r', label='Training loss')
plt.plot(epochs, val_loss_values, 'bo', label='Test loss')
plt.title('Training and test loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.grid('off')
plt.show()
plt.subplot(212)
plt.plot(epochs, history.history['acc'], 'r', label='Training accuracy')
plt.plot(epochs, val_acc_values, 'bo', label='Test accuracy')
plt.title('Training and test accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.gca().set_yticklabels(['{:.0f}%'.format(x*100) for x in plt.gca().get_yticks()])
plt.legend()
plt.grid('off')
plt.show()
# + id="evIdgTQ4KW8r" colab_type="code" colab={}
# + [markdown] id="FwZ8x6qEUkW4" colab_type="text"
# # Scenario 2: Small images
# + id="grv2y4C_JQ75" colab_type="code" colab={}
import cv2
x2 = []
y2 = []
for i in range(len(x)):
img = cv2.resize(x[i],(50,50))
x2.append(img)
img_label = np.where(y[i]=="male",1,0)
y2.append(img_label)
# + id="O6a8Wt3pJRAn" colab_type="code" outputId="f62e0812-87af-43f9-cd5e-a0cbe825099e" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.array(x2).shape
# + id="fMNBBG6WKW_w" colab_type="code" colab={}
x2 = np.array(x2)
x2 = x2.reshape(x2.shape[0],x2.shape[1],x2.shape[2],3)
# + id="xRmE_SmMJnl3" colab_type="code" outputId="4177d680-0cbf-4cf2-9aa8-3793a3de8e92" colab={"base_uri": "https://localhost:8080/", "height": 493}
model = Sequential()
model.add(Conv2D(64, kernel_size=(3, 3), activation='relu',input_shape=(50,50,3)))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(128, kernel_size=(3, 3), activation='relu',padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(256, kernel_size=(3, 3), activation='relu',padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(512, kernel_size=(3, 3), activation='relu',padding='same'))
model.add(Flatten())
model.add(Dense(100, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.summary()
# + id="CS3MU1nTJsVY" colab_type="code" outputId="273a081c-72b3-44d8-eb21-a280c9afa5e3" colab={"base_uri": "https://localhost:8080/", "height": 374}
model.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
X = np.array(x2)/255
Y = np.array(y2)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,Y, test_size=0.1, random_state=42)
history = model.fit(X_train, y_train, batch_size=32,epochs=10,verbose=1,validation_data = (X_test, y_test))
# + id="6rdC1vtTJnjG" colab_type="code" outputId="1b526c71-6e1f-4008-d9c3-c61b197c6a71" colab={"base_uri": "https://localhost:8080/", "height": 388}
import matplotlib.pyplot as plt
# %matplotlib inline
history_dict = history.history
loss_values = history_dict['loss']
val_loss_values = history_dict['val_loss']
acc_values = history_dict['acc']
val_acc_values = history_dict['val_acc']
epochs = range(1, len(val_loss_values) + 1)
plt.subplot(211)
plt.plot(epochs, history.history['loss'], 'r', label='Training loss')
plt.plot(epochs, val_loss_values, 'bo', label='Test loss')
plt.title('Training and test loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.grid('off')
plt.show()
plt.subplot(212)
plt.plot(epochs, history.history['acc'], 'r', label='Training accuracy')
plt.plot(epochs, val_acc_values, 'bo', label='Test accuracy')
plt.title('Training and test accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.gca().set_yticklabels(['{:.0f}%'.format(x*100) for x in plt.gca().get_yticks()])
plt.legend()
plt.grid('off')
plt.show()
# + id="yQiVImdfMH33" colab_type="code" colab={}
# + id="8b4sdn-VMH81" colab_type="code" colab={}
# + [markdown] id="zMSBY7V7UoZG" colab_type="text"
# # Aggressive pooling on big images
# + id="oRDod5ZlMICz" colab_type="code" colab={}
x2 = []
y2 = []
for i in range(len(x)):
x2.append(x[i])
img_label = np.where(y[i]=="male",1,0)
y2.append(img_label)
# + id="4cmrZx9UMIH9" colab_type="code" colab={}
x2 = np.array(x2)
x2 = x2.reshape(x2.shape[0],x2.shape[1],x2.shape[2],3)
# + id="5kIByaHYJngR" colab_type="code" outputId="f650c12a-86e4-4bbd-d8c0-a8dde0967108" colab={"base_uri": "https://localhost:8080/", "height": 493}
model = Sequential()
model.add(Conv2D(64, kernel_size=(3, 3), activation='relu',input_shape=(300,300,3)))
model.add(MaxPooling2D(pool_size=(3, 3)))
model.add(Conv2D(128, kernel_size=(3, 3), activation='relu',padding='same'))
model.add(MaxPooling2D(pool_size=(3, 3)))
model.add(Conv2D(256, kernel_size=(3, 3), activation='relu',padding='same'))
model.add(MaxPooling2D(pool_size=(3, 3)))
model.add(Conv2D(512, kernel_size=(3, 3), activation='relu',padding='same'))
model.add(Flatten())
model.add(Dense(100, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.summary()
# + id="2T80H4lnMSDI" colab_type="code" outputId="ceca256e-a25b-44a5-9308-30f22d1f7f0f" colab={"base_uri": "https://localhost:8080/", "height": 374}
model.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
X = np.array(x2)/255
Y = np.array(y2)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,Y, test_size=0.1, random_state=42)
history = model.fit(X_train, y_train, batch_size=32,epochs=10,verbose=1,validation_data = (X_test, y_test))
# + id="CLzT4ABwMSJe" colab_type="code" outputId="71ffbbde-e40c-4786-a7c1-4cf4fc28a572" colab={"base_uri": "https://localhost:8080/", "height": 388}
import matplotlib.pyplot as plt
# %matplotlib inline
history_dict = history.history
loss_values = history_dict['loss']
val_loss_values = history_dict['val_loss']
acc_values = history_dict['acc']
val_acc_values = history_dict['val_acc']
epochs = range(1, len(val_loss_values) + 1)
plt.subplot(211)
plt.plot(epochs, history.history['loss'], 'r', label='Training loss')
plt.plot(epochs, val_loss_values, 'bo', label='Test loss')
plt.title('Training and test loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.grid('off')
plt.show()
plt.subplot(212)
plt.plot(epochs, history.history['acc'], 'r', label='Training accuracy')
plt.plot(epochs, val_acc_values, 'bo', label='Test accuracy')
plt.title('Training and test accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.gca().set_yticklabels(['{:.0f}%'.format(x*100) for x in plt.gca().get_yticks()])
plt.legend()
plt.grid('off')
plt.show()
# + id="y3TRKpAUOWmo" colab_type="code" colab={}
# + id="SUKjtep7OWgz" colab_type="code" colab={}
# + [markdown] id="Q76v4wg2Uryy" colab_type="text"
# # VGG16 architecture
# + id="VUvQU6Y-KXCB" colab_type="code" colab={}
from keras.applications import vgg16
from keras.utils.vis_utils import plot_model
from keras.applications.vgg16 import preprocess_input
vgg16_model = vgg16.VGG16(include_top=False, weights='imagenet',input_shape=(300,300,3))
# + id="PGPW_PWYKXm2" colab_type="code" outputId="de90fd8e-8cb7-4c14-c819-3be58b9f31e3" colab={"base_uri": "https://localhost:8080/", "height": 799}
vgg16_model.summary()
# + id="RIM1RBZIKZ01" colab_type="code" outputId="3f458d00-ffb1-470c-c41d-d0581f3b2046" colab={"base_uri": "https://localhost:8080/", "height": 323}
import cv2
x2_vgg16 = []
for i in range(len(x)):
if(i%100 == 0):
print(i)
img = x[i]
img = preprocess_input(img.reshape(1,300,300,3))
img_new = vgg16_model.predict(img.reshape(1,300,300,3))
x2_vgg16.append(img_new)
# + id="o9MXRD6DK2fZ" colab_type="code" outputId="7aa24a00-2386-4fc9-bdc9-7fabd0c3a442" colab={"base_uri": "https://localhost:8080/", "height": 34}
x2_vgg16 = np.array(x2_vgg16)
x2_vgg16.shape
# + id="JCTbYIb_LmIu" colab_type="code" outputId="7d7eaf9a-2034-4329-8aca-a89125827dbd" colab={"base_uri": "https://localhost:8080/", "height": 34}
x2_vgg16= x2_vgg16.reshape(x2_vgg16.shape[0],x2_vgg16.shape[2],x2_vgg16.shape[3],x2_vgg16.shape[4])
x2_vgg16.shape
# + id="sVaG_te1LsOm" colab_type="code" colab={}
Y = np.array(y2)
# + id="u6qg85ScL52n" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(x2_vgg16,Y, test_size=0.1, random_state=42)
# + id="Si3WvsndL84I" colab_type="code" outputId="e1b79b2f-4c25-43c0-90fe-9cfbd830c89d" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(X_train.shape, X_test.shape, y_train.shape, y_test.shape)
# + id="c-zKSRUnNBPx" colab_type="code" outputId="a2ca8cae-3f9b-48ed-dc3d-8507d8dee1e9" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.max(X_train)
# + id="5uR4WnkBMDm7" colab_type="code" outputId="9101e788-719c-4eae-a510-c5ce0ff066e5" colab={"base_uri": "https://localhost:8080/", "height": 428}
model_vgg16 = Sequential()
model_vgg16.add(Conv2D(512, kernel_size=(3, 3), activation='relu',input_shape=(X_train.shape[1],X_train.shape[2],X_train.shape[3])))
model_vgg16.add(MaxPooling2D(pool_size=(2, 2)))
model_vgg16.add(Flatten())
model_vgg16.add(Dense(512, activation='relu'))
model_vgg16.add(Dropout(0.5))
model_vgg16.add(Dense(1, activation='sigmoid'))
model_vgg16.summary()
# + id="3yKSo3otMrEz" colab_type="code" colab={}
model_vgg16.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
# + id="TZ53V-bR2G1o" colab_type="code" outputId="944bf3e8-3d86-401b-bb85-071302935729" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.mean(y_train)
# + id="QKiM8uIpL-ft" colab_type="code" outputId="8fb9d896-8ab7-46b7-ad1d-dcfd59e86f57" colab={"base_uri": "https://localhost:8080/", "height": 374}
history_vgg16 = model_vgg16.fit(X_train/np.max(X_train), y_train, batch_size=16,epochs=10,verbose=1,validation_data = (X_test/np.max(X_train), y_test))
# + id="q-hPcBJfSPnj" colab_type="code" outputId="e420fc9b-dc60-4dab-a493-2df7ac5f0b00" colab={"base_uri": "https://localhost:8080/", "height": 388}
history_dict = history_vgg16.history
loss_values = history_dict['loss']
val_loss_values = history_dict['val_loss']
acc_values = history_dict['acc']
val_acc_values = history_dict['val_acc']
epochs = range(1, len(val_loss_values) + 1)
plt.subplot(211)
plt.plot(epochs, history_vgg16.history['loss'], 'ro', label='Training loss')
plt.plot(epochs, val_loss_values, 'b', label='Test loss')
plt.title('Training and test loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.grid('off')
plt.show()
plt.subplot(212)
plt.plot(epochs, history_vgg16.history['acc'], 'ro', label='Training accuracy')
plt.plot(epochs, val_acc_values, 'b', label='Test accuracy')
plt.title('Training and test accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.gca().set_yticklabels(['{:.0f}%'.format(x*100) for x in plt.gca().get_yticks()])
plt.legend()
plt.grid('off')
plt.show()
# + id="Jkgf7lMSQpMM" colab_type="code" outputId="777ab21c-c24d-4773-81bf-f3e4b2835f5e" colab={"base_uri": "https://localhost:8080/", "height": 34}
y_pred = model_vgg16.predict(X_test/np.max(X_train))
y_pred.shape
# + id="f2xjS0kTVvFF" colab_type="code" outputId="396b5d58-3445-44af-e948-5b477a9a5027" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.abs(y_pred.flatten()-y_test)[27]
# + id="Xd1TyyGXVhvn" colab_type="code" outputId="73bdfaef-2622-492e-bea2-5c23263d9c92" colab={"base_uri": "https://localhost:8080/", "height": 255}
np.argsort(np.abs(y_pred.flatten()-y_test))
# + id="OwCMpn5YMpJw" colab_type="code" colab={}
y_pred2 = np.where(y_pred>0.5,1,0)
# + id="b8Qz3iflRgOg" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
img_train, img_test, y_train, y_test = train_test_split(x,Y, test_size=0.1, random_state=42)
# + id="98EOCkk2WILN" colab_type="code" colab={}
y_test_char = np.where(y_test==0,'F','M')
# + id="m6hHqkMtWRzV" colab_type="code" colab={}
y_pred_char = np.where(y_pred>0.5,'M','F')
# + id="4U5nOEHSUJhF" colab_type="code" outputId="99a34e68-c9c1-4187-a7d1-fe27d01318e8" colab={"base_uri": "https://localhost:8080/", "height": 332}
import matplotlib.pyplot as plt
# %matplotlib inline
plt.subplot(221)
plt.imshow(img_test[85])
plt.title('Actual: '+str(y_test_char[85])+', '+'Predicted: '+str((y_pred_char[85][0])))
plt.grid('off')
plt.axis('off')
plt.subplot(222)
plt.title('Actual: '+str(y_test_char[32])+', '+'Predicted: '+str((y_pred_char[32][0])))
plt.imshow(img_test[32])
plt.grid('off')
plt.axis('off')
plt.subplot(223)
plt.title('Actual: '+str(y_test_char[83])+', '+'Predicted: '+str((y_pred_char[83][0])))
plt.imshow(img_test[83])
plt.grid('off')
plt.axis('off')
plt.subplot(224)
plt.title('Actual: '+str(y_test_char[46])+', '+'Predicted: '+str((y_pred_char[46][0])))
plt.imshow(img_test[46])
plt.grid('off')
plt.axis('off')
plt.show()
# + id="-3Hv1I0KSK30" colab_type="code" colab={}
# + id="JgkemAuGSK6o" colab_type="code" colab={}
# + id="wBWfhNCKSK9l" colab_type="code" colab={}
# + [markdown] id="_Kj1OpykUyLz" colab_type="text"
# # VGG19 architecture
# + id="iTDsK6pQREP_" colab_type="code" outputId="edc1f0f0-a8c2-482d-f3b7-a7912cf4714e" colab={"base_uri": "https://localhost:8080/", "height": 935}
from keras.applications import vgg19
from keras.utils.vis_utils import plot_model
from keras.applications.vgg19 import preprocess_input
vgg19_model = vgg19.VGG19(include_top=False, weights='imagenet',input_shape=(300,300,3))
vgg19_model.summary()
# + id="-1j9BedTSmg_" colab_type="code" outputId="f73f3ed0-9134-43da-de66-bc082ad43428" colab={"base_uri": "https://localhost:8080/", "height": 323}
import cv2
x2 = []
for i in range(len(x)):
if(i%100 == 0):
print(i)
img = x[i]
img = preprocess_input(img.reshape(1,300,300,3))
img_new = vgg19_model.predict(img.reshape(1,300,300,3))
x2.append(img_new)
# + id="1NCuhBFxXTIV" colab_type="code" outputId="1d71a512-47fa-4166-9926-21c8451a7f68" colab={"base_uri": "https://localhost:8080/", "height": 34}
x2 = np.array(x2)
x2.shape
# + id="yjgcDYmGXnpF" colab_type="code" outputId="46c756dd-9769-489b-8b6b-d893cf69b07e" colab={"base_uri": "https://localhost:8080/", "height": 34}
x2= x2.reshape(x2.shape[0],x2.shape[2],x2.shape[3],x2.shape[4])
x2.shape
# + id="ISGUnKFfXpYQ" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(x2,Y, test_size=0.1, random_state=42)
# + id="A_bK0OrMXqj8" colab_type="code" outputId="28fec3b0-5d07-4a1f-a3ec-eda7cb20f641" colab={"base_uri": "https://localhost:8080/", "height": 357}
model_vgg19 = Sequential()
model_vgg19.add(Conv2D(512, kernel_size=(3, 3), activation='relu',input_shape=(X_train.shape[1],X_train.shape[2],X_train.shape[3])))
model_vgg19.add(MaxPooling2D(pool_size=(2, 2)))
model_vgg19.add(Flatten())
model_vgg19.add(Dense(512, activation='relu'))
model_vgg19.add(Dropout(0.5))
model_vgg19.add(Dense(1, activation='sigmoid'))
model_vgg19.summary()
# + id="3Yff8PRgXr3I" colab_type="code" colab={}
model_vgg19.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
# + id="IScHi78sXteQ" colab_type="code" outputId="addf077a-aa2a-4f51-f965-bcac1704ccbc" colab={"base_uri": "https://localhost:8080/", "height": 374}
history_vgg19 = model_vgg19.fit(X_train/np.max(X_train), y_train, batch_size=16,epochs=10,verbose=1,validation_data = (X_test/np.max(X_train), y_test))
# + id="craIyFmbXvad" colab_type="code" outputId="366b4b87-0469-44d0-e519-b2c2e59d1463" colab={"base_uri": "https://localhost:8080/", "height": 388}
history_dict = history_vgg19.history
loss_values = history_dict['loss']
val_loss_values = history_dict['val_loss']
acc_values = history_dict['acc']
val_acc_values = history_dict['val_acc']
epochs = range(1, len(val_loss_values) + 1)
plt.subplot(211)
plt.plot(epochs, history_vgg19.history['loss'], 'ro', label='Training loss')
plt.plot(epochs, val_loss_values, 'b', label='Test loss')
plt.title('Training and test loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.grid('off')
plt.show()
plt.subplot(212)
plt.plot(epochs, history_vgg19.history['acc'], 'ro', label='Training accuracy')
plt.plot(epochs, val_acc_values, 'b', label='Test accuracy')
plt.title('Training and test accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.gca().set_yticklabels(['{:.0f}%'.format(x*100) for x in plt.gca().get_yticks()])
plt.legend()
plt.grid('off')
plt.show()
# + id="XZJ8aRwrX7X9" colab_type="code" colab={}
y_pred = model_vgg19.predict(X_test/np.max(X_train))
# + id="a6OKBGlQYCin" colab_type="code" outputId="a270ea4e-be5f-4cf8-d17f-03f9d8765350" colab={"base_uri": "https://localhost:8080/", "height": 255}
np.argsort(np.abs(y_pred.flatten() - y_test))
# + id="oS46ZA1c4Loe" colab_type="code" colab={}
y_pred_char = np.where(y_pred>0.5,'M','F')
y_test_char = np.where(y_test==0,'F','M')
# + id="CiBwEJG4YIKb" colab_type="code" outputId="3c4372ae-226d-4c33-a8cd-b99993e8c75a" colab={"base_uri": "https://localhost:8080/", "height": 332}
import matplotlib.pyplot as plt
# %matplotlib inline
plt.subplot(221)
plt.imshow(img_test[32])
plt.title('Actual: '+str(y_test_char[32])+', '+'Predicted: '+str((y_pred_char[32][0])))
plt.grid('off')
plt.axis('off')
plt.subplot(222)
plt.title('Actual: '+str(y_test_char[119])+', '+'Predicted: '+str((y_pred_char[119][0])))
plt.imshow(img_test[119])
plt.grid('off')
plt.axis('off')
plt.subplot(223)
plt.title('Actual: '+str(y_test_char[107])+', '+'Predicted: '+str((y_pred_char[107][0])))
plt.imshow(img_test[107])
plt.grid('off')
plt.axis('off')
plt.subplot(224)
plt.title('Actual: '+str(y_test_char[71])+', '+'Predicted: '+str((y_pred_char[71][0])))
plt.imshow(img_test[71])
plt.grid('off')
plt.axis('off')
plt.show()
# + id="TEjJgSMpYbpy" colab_type="code" colab={}
# + [markdown] id="8V_lVlP7U1TS" colab_type="text"
# # Inception v3 architecture
# + id="Do1Q4oYUYbtI" colab_type="code" outputId="f22a4a85-7221-4160-ba81-5f4e27584903" colab={"base_uri": "https://localhost:8080/", "height": 1000}
from keras.applications import inception_v3
from keras.applications.inception_v3 import preprocess_input
from keras.utils.vis_utils import plot_model
inception_model = inception_v3.InceptionV3(include_top=False, weights='imagenet',input_shape=(300,300,3))
inception_model.summary()
# + id="6-ZYZ1j9Y0XZ" colab_type="code" outputId="4cd9254d-0bbc-4d5e-a40a-ab1d91d54b6c" colab={"base_uri": "https://localhost:8080/", "height": 323}
import cv2
x2 = []
for i in range(len(x)):
if(i%100 == 0):
print(i)
img = x[i]
img = preprocess_input(img.reshape(1,300,300,3))
img_new = inception_model.predict(img.reshape(1,300,300,3))
x2.append(img_new)
x2 = np.array(x2)
x2.shape
x2= x2.reshape(x2.shape[0],x2.shape[2],x2.shape[3],x2.shape[4])
x2.shape
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(x2,Y, test_size=0.1, random_state=42)
# + id="4Vvy39CGYqoe" colab_type="code" outputId="7a232da6-55c8-48bb-83cc-ccc5d4a0be33" colab={"base_uri": "https://localhost:8080/", "height": 714}
model_inception_v3 = Sequential()
model_inception_v3.add(Conv2D(512, kernel_size=(3, 3), activation='relu',input_shape=(X_train.shape[1],X_train.shape[2],X_train.shape[3])))
model_inception_v3.add(MaxPooling2D(pool_size=(2, 2)))
model_inception_v3.add(Flatten())
model_inception_v3.add(Dense(512, activation='relu'))
model_inception_v3.add(Dropout(0.5))
model_inception_v3.add(Dense(1, activation='sigmoid'))
model_inception_v3.summary()
model_inception_v3.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
history_inception_v3 = model_inception_v3.fit(X_train/np.max(X_train), y_train, batch_size=16,epochs=10,verbose=1,validation_data = (X_test/np.max(X_train), y_test))
# + id="vnLVx_hBZRF1" colab_type="code" outputId="8c70d03d-f1fc-4fac-ed5a-4a3b4670395f" colab={"base_uri": "https://localhost:8080/", "height": 388}
history_dict = history_inception_v3.history
loss_values = history_dict['loss']
val_loss_values = history_dict['val_loss']
acc_values = history_dict['acc']
val_acc_values = history_dict['val_acc']
epochs = range(1, len(val_loss_values) + 1)
plt.subplot(211)
plt.plot(epochs, history_inception_v3.history['loss'], 'ro', label='Training loss')
plt.plot(epochs, val_loss_values, 'b', label='Test loss')
plt.title('Training and test loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.grid('off')
plt.show()
plt.subplot(212)
plt.plot(epochs, history_inception_v3.history['acc'], 'ro', label='Training accuracy')
plt.plot(epochs, val_acc_values, 'b', label='Test accuracy')
plt.title('Training and test accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.gca().set_yticklabels(['{:.0f}%'.format(x*100) for x in plt.gca().get_yticks()])
plt.legend()
plt.grid('off')
plt.show()
# + id="UP53f-S5ZmcM" colab_type="code" colab={}
# + id="Pg1KEWyFaKgX" colab_type="code" colab={}
# + [markdown] id="pkLzZS80U4W5" colab_type="text"
# # ResNet 50 architecture
# + id="d2JpkRRBaKkE" colab_type="code" outputId="7c3e5f4a-c113-4976-d596-5a77402ccb3a" colab={"base_uri": "https://localhost:8080/", "height": 1000}
from keras.applications import resnet50
from keras.applications.resnet50 import preprocess_input
resnet50_model = resnet50.ResNet50(include_top=False, weights='imagenet',input_shape=(300,300,3))
resnet50_model.summary()
# + id="v3pk68CdaXPD" colab_type="code" outputId="0cd76993-fb39-4a5b-c474-f54bc6ba0694" colab={"base_uri": "https://localhost:8080/", "height": 323}
import cv2
x2 = []
for i in range(len(x)):
if(i%100 == 0):
print(i)
img = x[i]
img = preprocess_input(img.reshape(1,300,300,3))
img_new = resnet50_model.predict(img.reshape(1,300,300,3))
x2.append(img_new)
# + id="BeksjCVgabnh" colab_type="code" outputId="7086bf5d-d7f4-4361-ec7d-28a8d784e35b" colab={"base_uri": "https://localhost:8080/", "height": 34}
x2 = np.array(x2)
x2.shape
# + id="jkdGm0DQahwx" colab_type="code" outputId="8a064e75-4243-4e24-f638-8426ebb9fab7" colab={"base_uri": "https://localhost:8080/", "height": 34}
x2= x2.reshape(x2.shape[0],x2.shape[2],x2.shape[3],x2.shape[4])
x2.shape
# + id="btaPYIlbt21r" colab_type="code" colab={}
Y = np.array(y2)
# + id="sTKP46Meaiu4" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(x2,Y, test_size=0.1, random_state=42)
# + id="APN7aQZgakDv" colab_type="code" outputId="80f4196f-1ada-4538-926e-803d4b50f8f7" colab={"base_uri": "https://localhost:8080/", "height": 425}
model_resnet50 = Sequential()
model_resnet50.add(Conv2D(512, kernel_size=(3, 3), activation='relu',input_shape=(X_train.shape[1],X_train.shape[2],X_train.shape[3])))
model_resnet50.add(MaxPooling2D(pool_size=(2, 2)))
model_resnet50.add(Conv2D(512, kernel_size=(3, 3), activation='relu'))
model_resnet50.add(MaxPooling2D(pool_size=(2, 2)))
model_resnet50.add(Flatten())
model_resnet50.add(Dense(512, activation='relu'))
model_resnet50.add(Dropout(0.5))
model_resnet50.add(Dense(1, activation='sigmoid'))
model_resnet50.summary()
# + id="7JhsBRlBaloW" colab_type="code" colab={}
model_resnet50.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
# + id="CoDqknOHmyaH" colab_type="code" outputId="df877050-6f59-485f-a1c2-742ebf229d82" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.max(X_train)
# + id="CxJsppZYaryk" colab_type="code" outputId="8b783e6c-2506-496d-ee81-bc1e1fa167b2" colab={"base_uri": "https://localhost:8080/", "height": 374}
history_resnet50 = model_resnet50.fit(X_train/np.max(X_train), y_train, batch_size=32,epochs=10,verbose=1,validation_data = (X_test/np.max(X_train), y_test))
# + id="Fo1A27MJmnzt" colab_type="code" outputId="f39b55fc-ff37-430f-ef2f-fa07c5e5e758" colab={"base_uri": "https://localhost:8080/", "height": 388}
import matplotlib.pyplot as plt
# %matplotlib inline
history_dict = history_resnet50.history
loss_values = history_dict['loss']
val_loss_values = history_dict['val_loss']
acc_values = history_dict['acc']
val_acc_values = history_dict['val_acc']
epochs = range(1, len(val_loss_values) + 1)
plt.subplot(211)
plt.plot(epochs, history_resnet50.history['loss'], 'ro', label='Training loss')
plt.plot(epochs, val_loss_values, 'b', label='Test loss')
plt.title('Training and test loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.grid('off')
plt.show()
plt.subplot(212)
plt.plot(epochs, history_resnet50.history['acc'], 'ro', label='Training accuracy')
plt.plot(epochs, val_acc_values, 'b', label='Test accuracy')
plt.title('Training and test accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.gca().set_yticklabels(['{:.0f}%'.format(x*100) for x in plt.gca().get_yticks()])
plt.legend()
plt.grid('off')
plt.show()
# + id="SSZ-a32puR0o" colab_type="code" colab={}
# + id="A6W6l9tF5yBD" colab_type="code" colab={}
# + [markdown] id="sVTtb-zUU99I" colab_type="text"
# # Visualizing the output intermediate layers
# + id="CTdEvrBp5yFK" colab_type="code" outputId="db6854e5-2f26-476e-f344-d3557c792981" colab={"base_uri": "https://localhost:8080/", "height": 320}
import matplotlib.pyplot as plt
# %matplotlib inline
plt.imshow(x[3])
plt.grid('off')
# + id="b17U1-6z6w6s" colab_type="code" outputId="630e30a5-7909-4dff-aecd-675499b74294" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.max(X_train)
# + id="DHc0iVGV6Pcv" colab_type="code" outputId="7dac3a50-9a7c-4232-cb74-139fc8fe2078" colab={"base_uri": "https://localhost:8080/", "height": 34}
from keras.applications.vgg16 import preprocess_input
model_vgg16.predict(vgg16_model.predict(preprocess_input(x[3].reshape(1,300,300,3)))/np.max(X_train))
# + id="zSbtvfWa7eQ5" colab_type="code" outputId="2c1f794f-dbda-4ae2-b953-345a6af66603" colab={"base_uri": "https://localhost:8080/", "height": 340}
for layer in vgg16_model.layers:
print(layer.name)
# + id="CLW5aGQB6Hyp" colab_type="code" colab={}
from keras import models
activation_model = models.Model(inputs=vgg16_model.input,outputs=vgg16_model.layers[1].output)
activations = activation_model.predict(preprocess_input(x[3].reshape(1,300,300,3)))
# + id="Swj3klEl7Kp8" colab_type="code" outputId="6845f7e3-ccdb-4bfa-af40-7431ff1dd89f" colab={"base_uri": "https://localhost:8080/", "height": 608}
fig, axs = plt.subplots(6, 6, figsize=(10, 10))
fig.subplots_adjust(hspace = .5, wspace=.5)
first_layer_activation = activations[0]
for i in range(6):
for j in range(6):
try:
axs[i,j].set_ylim((224, 0))
axs[i,j].contourf(first_layer_activation[:,:,((6*i)+j)],6,cmap='viridis')
axs[i,j].set_title('filter: '+str((6*i)+j))
axs[i,j].axis('off')
except:
continue
# + id="9AMdc26177Bo" colab_type="code" colab={}
# + id="VbYKby6k8tkO" colab_type="code" colab={}
activation_model = models.Model(inputs=vgg16_model.input,outputs=vgg16_model.layers[1].output)
activations = activation_model.predict(preprocess_input(np.array(x[:36]).reshape(36,300,300,3)))
# + id="1C79qYZ88wkU" colab_type="code" colab={}
fig, axs = plt.subplots(6, 6, figsize=(10, 10))
fig.subplots_adjust(hspace = .5, wspace=.5)
first_layer_activation = activations
for i in range(6):
for j in range(6):
try:
axs[i,j].set_ylim((224, 0))
axs[i,j].contourf(first_layer_activation[((6*i)+j),:,:,7],6,cmap='viridis')
axs[i,j].set_title('filter: '+str((6*i)+j))
axs[i,j].axis('off')
except:
continue
# + id="euxu7j3I9EJz" colab_type="code" colab={}
# + id="qVgibt9WWxP6" colab_type="code" colab={}
activation_model = models.Model(inputs=vgg16_model.input,outputs=vgg16_model.layers[-1].output)
activations = activation_model.predict(preprocess_input(x[3].reshape(1,300,300,3)))
# + id="wj4GdzaZamzU" colab_type="code" outputId="0653897e-85e0-417a-9d67-d67fa365d985" colab={"base_uri": "https://localhost:8080/", "height": 34}
last_layer_activation.shape
# + id="q7jr7Oh8W0so" colab_type="code" outputId="7b889fca-ed50-43a9-a0c2-a34ffc77448a" colab={"base_uri": "https://localhost:8080/", "height": 606}
fig, axs = plt.subplots(12, 11, figsize=(10, 10))
fig.subplots_adjust(hspace = .5, wspace=.5)
last_layer_activation = activations
count = 0
for i in range(12):
for j in range(11):
try:
count+=1
axs[i,j].set_ylim((6, 0))
axs[i,j].contourf(last_layer_activation[0,:,:,((12*i)+j)],11,cmap='viridis')
axs[i,j].set_title('filter: '+str(count))
axs[i,j].axis('off')
except:
continue
# + id="Fjh48LMcW2je" colab_type="code" colab={}
| Chapter05/Transfer_learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import csv
import mne
import pandas as pd
import matplotlib.pyplot as plt
# +
file_dir = "C:/Users/Bruger/Documents/Uni/6. Semester/BP data/data/v2.1.0/edf/01_tcp_ar/002/00000254/s007_2013_03_25/00000254_s007_t000.edf"
file = mne.io.read_raw_edf(file_dir, preload=True)
data = file.filter(0.1, 100)
data = data.notch_filter(60)
data
# +
file_dir = "C:/Users/Bruger/Documents/Uni/6. Semester/BP data/data/v2.1.0/edf/01_tcp_ar/002/00000254/s005_2010_11_15/00000254_s005_t000.edf"
file = mne.io.read_raw_edf(file_dir, preload=True)
data = file.filter(0.1, 100)
data = data.notch_filter(60)
data
# +
file_dir = "C:/Users/Bruger/Documents/Uni/6. Semester/BP data/data/v2.1.0/edf/03_tcp_ar_a/046/00004671/s017_2015_03_27/00004671_s017_t000.edf"
file = mne.io.read_raw_edf(file_dir, preload=True)
data = data.notch_filter(60)
data = file.filter(0.1, 100)
data
# +
#mne.export.export_raw("C:/Users/Bruger/Documents/Uni/6. Semester/BP/test2/data.edf", data, overwrite=True)
# -
df = data.to_data_frame()
df["EEG O1-REF"]
# %matplotlib
data.plot()
montage= {0: ["FP1-F7", "EEG FP1-REF", "EEG F7-REF"],
1: ["F7-T3", "EEG F7-REF", "EEG T3-REF"],
2: ["T3-T5", "EEG T3-REF", "EEG T5-REF"],
3: ["T5-O1", "EEG T5-REF", "EEG O1-REF"],
4: ["FP2-F8", "EEG FP2-REF", "EEG F8-REF"],
5: ["F8-T4", "EEG F8-REF", "EEG T4-REF"],
6: ["T4-T6", "EEG T4-REF", "EEG T6-REF"],
7: ["T6-O2", "EEG T6-REF", "EEG O2-REF"],
8: ["A1-T3", "EEG A1-REF", "EEG T3-REF"],
9: ["T3-C3", "EEG T3-REF", "EEG C3-REF"],
10: ["C3-CZ", "EEG C3-REF", "EEG CZ-REF"],
11: ["CZ-C4", "EEG CZ-REF", "EEG C4-REF"],
12: ["C4-T4", "EEG C4-REF", "EEG T4-REF"],
13: ["T4-A2", "EEG T4-REF", "EEG A2-REF"],
14: ["FP1-F3", "EEG FP1-REF", "EEG F3-REF"],
15: ["F3-C3", "EEG F3-REF", "EEG C3-REF"],
16: ["C3-P3", "EEG C3-REF", "EEG P3-REF"],
17: ["P3-O1", "EEG P3-REF", "EEG O1-REF"],
18: ["FP2-F4", "EEG FP2-REF", "EEG F4-REF"],
19: ["F4-C4", "EEG F4-REF", "EEG C4-REF"],
20: ["C4-P4", "EEG C4-REF", "EEG P4-REF"],
21: ["P4-O2", "EEG P4-REF", "EEG O2-REF"]}
montage
dir_rec = "C:/Users/Bruger/Documents/Uni/6. Semester/BP/data/v2.1.0/edf/01_tcp_ar/002/00000254/s005_2010_11_15/00000254_s005_t000.rec"
which_montages = set()
target = []
with open(dir_rec, "r") as file: # Læser csv-fil
ls = csv.reader(file)
for rows in ls:
target.append([int(rows[0]), float(rows[1]), float(rows[2]), int(rows[3])])
which_montages.add(int(rows[0]))
# Nu skal den nye data-frame laves:
first = True
for i in which_montages:
col_names = montage.get(i)
if first:
list1 = df[col_names[1]] # Get the first series
list2 = df[col_names[2]] # Get the second series
df_new = list1-list2
df_new = df_new.rename(col_names[0]) # Rename
first = False
else:
list1 = df[col_names[1]]
list2 = df[col_names[2]]
diff = list1-list2
diff = diff.rename(col_names[0]) # Rename
df_new = pd.concat([df_new, diff], axis=1, join='inner')
df_new
# Lets make the targets:
import torch
import math
import numpy as np
# +
tar = torch.zeros(df_new.shape[1], df_new.shape[0])
sorted_index = sorted(list(which_montages))
for i in target: # i = [montage_channel, start, end, type_artifact]
index = sorted_index.index(i[0]) # Find the correct index in the target
tar[index][250 * math.floor(i[1]): 250 * math.ceil(i[2])] = 1 # Make the artifacts = 1
# +
figure, axis = plt.subplots(2, 2)
# The artifacts on channel: FP1-F7
axis[0, 0].plot(df_new["FP1-F7"][(tar[0] == 0).numpy()], "b")
axis[0, 0].plot(df_new["FP1-F7"][(tar[0] == 1).numpy()], "ro", markersize=1)
axis[0, 0].set_title("FP1-F7")
axis[0, 0].axes.xaxis.set_visible(False) # remove digits on x-axis
# The artifacts on channel: F7-T3
axis[0, 1].plot(df_new["F7-T3"][(tar[1] == 0).numpy()], "b")
axis[0, 1].plot(df_new["F7-T3"][(tar[1] == 1).numpy()], "ro", markersize=1)
axis[0, 1].set_title("F7-T3")
axis[0, 1].axes.xaxis.set_visible(False) # remove digits on x-axis
# The artifacts on channel: FP2-F8
axis[1, 0].plot(df_new["FP2-F8"][(tar[2] == 0).numpy()], "b")
axis[1, 0].plot(df_new["FP2-F8"][(tar[2] == 1).numpy()], "ro", markersize=1)
axis[1, 0].set_title("FP2-F8")
axis[1, 0].axes.xaxis.set_visible(False) # remove digits on x-axis
# The artifacts on channel: F8-T4
axis[1, 1].plot(df_new["F8-T4"][(tar[3] == 0).numpy()], "b")
axis[1, 1].plot(df_new["F8-T4"][(tar[3] == 1).numpy()], "ro", markersize=1)
axis[1, 1].set_title("F8-T4")
axis[1, 1].axes.xaxis.set_visible(False) # remove digits on x-axis
# https://www.geeksforgeeks.org/plot-multiple-plots-in-matplotlib/
# -
plt.plot(df_new["FP1-F7"][(tar[0] == 1).numpy()], "ro", markersize=1)
# +
dir1_rec = "C:/Users/Bruger/Documents/Uni/6. Semester/BP/data/v2.1.0/lists/rec_01_tcp_ar.list"
dir2_rec = "C:/Users/Bruger/Documents/Uni/6. Semester/BP/data/v2.1.0/lists/rec_02_tcp_le.list"
dir3_rec = "C:/Users/Bruger/Documents/Uni/6. Semester/BP/data/v2.1.0/lists/rec_03_tcp_ar_a.list"
rec = []
with open(dir1_rec, "r") as file: # Læser csv-fil
ls = csv.reader(file)
for i in ls:
rec.append(i[0])
with open(dir2_rec, "r") as file: # Læser csv-fil
ls = csv.reader(file)
for i in ls:
rec.append(i[0])
with open(dir3_rec, "r") as file: # Læser csv-fil
ls = csv.reader(file)
for i in ls:
rec.append(i[0])
# +
artifacts = 0
for i in rec:
direct = "C:/Users/Bruger/Documents/Uni/6. Semester/BP/data/v2.1.0" + i[2:]
with open(direct, "r") as file: # Læser csv-fil
ls = csv.reader(file)
for rows in ls:
artifacts += 1
print(artifacts)
# -
def make_file_list(edf_dir: str, rec_dir: str, file_dir: list):
file1 = open(edf_dir)
file2 = open(rec_dir)
reader1 = csv.reader(file1)
reader2 = csv.reader(file2)
for i in zip(reader1, reader2):
first = "C:/Users/Bruger/Documents/Uni/6. Semester/BP/data/v2.1.0" + i[0][0][2:]
second = "C:/Users/Bruger/Documents/Uni/6. Semester/BP/data/v2.1.0" + i[1][0][2:]
file_dir.append([first, second])
file1.close()
file2.close()
return file_dir
# +
yeet = []
edf_dir = "C:/Users/Bruger/Documents/Uni/6. Semester/BP/data/v2.1.0/lists/edf_01_tcp_ar.list"
rec_dir = "C:/Users/Bruger/Documents/Uni/6. Semester/BP/data/v2.1.0/lists/rec_01_tcp_ar.list"
yeet = make_file_list(edf_dir, rec_dir, yeet)
len(yeet)
# +
file1 = open("C:/Users/Bruger/Documents/Uni/6. Semester/BP/data/v2.1.0/lists/rec_01_tcp_ar.list")
file2 = open("C:/Users/Bruger/Documents/Uni/6. Semester/BP/data/v2.1.0/lists/edf_01_tcp_ar.list")
reader1 = csv.reader(file1)
reader2 = csv.reader(file2)
for i in zip(reader1, reader2):
first = "C:/Users/Bruger/Documents/Uni/6. Semester/BP/data/v2.1.0" + i[0][0][2:]
second = "C:/Users/Bruger/Documents/Uni/6. Semester/BP/data/v2.1.0" + i[1][0][2:]
yeet.append([first, second])
file1.close()
file2.close()
# -
def increaser():
for i in range(10):
nr += 1
return nr
# +
nr = 0
increaser()
increaser()
increaser()
print(nr)
# -
| FourGraph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
x1 = 2
x2 = 5
print(x1, '+' ,x2 , "=" ,x1+x2 )
print(x1, '-' ,x2 , "=" ,x1-x2 )
print(x1, '*' ,x2 , "=" ,x1*x2 )
print(x1, '/' ,x2 , "=" ,x1/x2 )
print(x1, '//' ,x2 , "=" ,x1//x2)
print(x1, '%' ,x2 , "=" ,x1%x2 )
print(x1, '**' ,x2 , "=" ,x1**x2)
x1
x2
x1 = 50
| Week - 1/Calculator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="pnBBzbg0BezX"
# #Introducing optimization
# + colab={"base_uri": "https://localhost:8080/"} id="qei4RuK2BWOk" outputId="73c95bd3-edf8-4983-c2e3-d4e4014132c4"
# !pip install nnfs
# + id="O9-E0-EnCtR2"
import numpy as np
# + id="7aUt-uynCrEH"
# Common loss class
class Loss:
# Calculates the data and regularizations losses given model output and
# ground truth values
def calculate(self, output, y):
# Calculate sample losses
sample_losses = self.forward(output, y)
# Calculate mean loss
data_loss = np.mean(sample_losses)
# Return loss
return data_loss
# Cross-entropy loss
class Loss_CategoricalCrossentropy(Loss):
# Forward pass
def forward(self, y_pred, y_true):
# Number of samples in a batch
samples = len(y_pred)
# Clip data to prevent division by 0 when calculating log. We have to avoid
# calculating log(0). Then, we clip all the output values from softmax between
# 1e-7 and 1-1e-7.
y_pred_clipped = np.clip(y_pred, 1e-7, 1-1e-7)
# Probabilities for target values - only if categorical labels
# this is for sparse labels [0, 2, 5, 1], each value is a label
if len(y_true.shape) == 1:
correct_confidences = y_pred_clipped[range(samples), y_true]
# for one hot encoded labels, we multiply each softmax output
# by the label
elif len(y_true.shape) == 2:
correct_confidences = np.sum(y_pred_clipped * y_true, axis=1)
# Losses
negative_log_likelihoods = -np.log(correct_confidences)
return negative_log_likelihoods
# --------------------class layer dense----------------
class Layer_Dense:
def __init__(self, n_inputs, n_neurons):
"""
Initialise weights and biases
random.randn returns a matrix with n_inputs x n_neurons shape
it makes sense to create the weights matrix with inputs x neurons shape
NUMBER OF WEIGHTS VALUES = NUMBER OF INPUTS X NUMBER OF NEURONS
note that we define the shape as inputs X neurons and not neurons X inputs
to avoid transposing every time we do a forward pass
4 inputs, 3 neurons
wij -> weight, i -> input value, j -> neuron
[w11, w12, w13]
[w21, w22, w23]
[w31, w32, w33]
[w41, w42, w43]
each column is already a vector of weights of each neuron for the input
random.randn generates values from a Gaussian distribution with a mean of 0
and a variance of 1, which mean that it'll generate random numbers, positive and
negative [-1,1], centered at 0 and with the mean value close to 0.
we multiply here by 0.01 because we want to initialise weights with non-zero values
but these values have to be small because training updates will be smaller.
If weight values are very big, the training will last more time.
"""
self.weights = 0.01 * np.random.randn(n_inputs, n_neurons)
# ONE BIAS VALUE FOR EACH NEURON
self.biases = np.zeros((1, n_neurons))
# Forward pass
def forward(self, inputs):
# Calculate outputs values from inputs, weights and biases
self.output = np.dot(inputs, self.weights) + self.biases
# --------------------------------------------------------
# ReLU activation class
class Activation_ReLU:
# Forward pass
def forward(self, inputs):
# Calculate output values from input
self.output = np.maximum(0, inputs)
# -------------------------------------------------------
# Softmax activation class
class Activation_Softmax:
# Forward pass
def forward(self, inputs):
# Get unnormalized probabilities
exp_values = np.exp(inputs - np.max(inputs, axis=1, keepdims=True))
# Normalize them for each sample
probabilities = exp_values / np.sum(exp_values, axis=1, keepdims=True)
self.output = probabilities
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="ubU68qPECHUV" outputId="9c020e91-b9fb-49fe-e073-7c94a7375593"
import matplotlib.pyplot as plt
import nnfs
from nnfs.datasets import vertical_data, spiral_data
nnfs.init()
X, y = vertical_data(samples=100, classes=3)
plt.scatter(X[:, 0], X[:, 1], c=y, s=40, cmap='brg')
plt.show()
# + [markdown] id="IuBJXBwTDy4z"
# # Here we are testing if changing the weights randomly can help the neural network to achieve convergence
# + colab={"base_uri": "https://localhost:8080/"} id="OzYDDaNmC-sc" outputId="a644663c-ce7b-40b9-a09c-7011107ee9f1"
#create model
dense1 = Layer_Dense(2, 3)
activation1 = Activation_ReLU()
dense2 = Layer_Dense(3, 3)
activation2 = Activation_Softmax()
#create loss function
loss_function = Loss_CategoricalCrossentropy()
#create some variables to track the best loss and the associated weights and biases
lowest_loss = 9999999
best_dense1_weights = dense1.weights.copy()
best_dense1_biases = dense1.biases.copy()
best_dense2_weights = dense2.weights.copy()
best_dense2_biases = dense2.biases.copy()
# we initialize weights to random in each epoch, and later we update the weights
# if the loss decreases.
for iteration in range(10000):
#generate a new set of weights for iteration
dense1.weights = 0.05 * np.random.randn(2, 3)
dense1.biases = 0.05 * np.random.randn(1, 3)
dense2.weights = 0.05 * np.random.randn(3, 3)
dense2.biases = 0.05 * np.random.randn(1, 3)
# perform a forward pass of the training data through this layer
dense1.forward(X)
activation1.forward(dense1.output)
dense2.forward(activation1.output)
activation2.forward(dense2.output)
# calculating loss
loss = loss_function.calculate(activation2.output, y)
# calculate accuracy from output of activation2 and targets
predictions = np.argmax(activation2.output, axis=1)
accuracy = np.mean(predictions==y)
# if loss is smaller - print print and save weights and biases aside
if loss < lowest_loss:
print('New set of weights found, iteration:', iteration,
'loss:', loss, 'acc:', accuracy)
best_dense1_weights = dense1.weights.copy()
best_dense1_biases = dense1.biases.copy()
best_dense2_weights = dense2.weights.copy()
best_dense2_biases = dense2.biases.copy()
lowest_loss = loss
# + [markdown] id="rQEujPIBIg_k"
# # We can see that searching the weights randomly is not the proper way to do it.
# + [markdown] id="5G0Qr675I99U"
# Another option is to take a fraction of the weights and biases that produce the decrease in loss, instead of taking random values after decreasing the loss.
# + colab={"base_uri": "https://localhost:8080/"} id="pGMXJ8qhJIMK" outputId="8d36b007-22c1-475f-ec76-7694c6704756"
#create model
dense1 = Layer_Dense(2, 3)
activation1 = Activation_ReLU()
dense2 = Layer_Dense(3, 3)
activation2 = Activation_Softmax()
#create loss function
loss_function = Loss_CategoricalCrossentropy()
#create some variables to track the best loss and the associated weights and biases
lowest_loss = 9999999
best_dense1_weights = dense1.weights.copy()
best_dense1_biases = dense1.biases.copy()
best_dense2_weights = dense2.weights.copy()
best_dense2_biases = dense2.biases.copy()
# we initialize weights to random in each epoch, and later we update the weights
# if the loss decreases.
for iteration in range(10000):
#generate a new set of weights for iteration
# HERE WE TAKE A FRACTION OF THE WEIGHTS
dense1.weights += 0.05 * np.random.randn(2, 3)
dense1.biases += 0.05 * np.random.randn(1, 3)
dense2.weights += 0.05 * np.random.randn(3, 3)
dense2.biases += 0.05 * np.random.randn(1, 3)
# perform a forward pass of the training data through this layer
dense1.forward(X)
activation1.forward(dense1.output)
dense2.forward(activation1.output)
activation2.forward(dense2.output)
# calculating loss
loss = loss_function.calculate(activation2.output, y)
# calculate accuracy from output of activation2 and targets
predictions = np.argmax(activation2.output, axis=1)
accuracy = np.mean(predictions==y)
# if loss is smaller - print print and save weights and biases aside
if loss < lowest_loss:
print('New set of weights found, iteration:', iteration,
'loss:', loss, 'acc:', accuracy)
best_dense1_weights = dense1.weights.copy()
best_dense1_biases = dense1.biases.copy()
best_dense2_weights = dense2.weights.copy()
best_dense2_biases = dense2.biases.copy()
lowest_loss = loss
else:
dense1.weights = best_dense1_weights.copy()
dense1.biases = best_dense1_biases.copy()
dense2.weights = best_dense2_weights.copy()
dense2.biases = best_dense2_biases.copy()
# + [markdown] id="cpxtauqQLF3B"
# We see here that for this easy problem, this approach is just fine. Let's see it for the spiral dataset.
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="KL42i7vjLZVl" outputId="754c0c66-6ff2-4dfe-eac4-1f7b70b4f0b0"
X, y = spiral_data(samples=100, classes=3)
plt.scatter(X[:, 0], X[:, 1], c=y, s=40, cmap='brg')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="-CNFnmRQLPBb" outputId="ea28afd4-d8d0-4388-eeca-7a34441c4990"
#create model
dense1 = Layer_Dense(2, 3)
activation1 = Activation_ReLU()
dense2 = Layer_Dense(3, 3)
activation2 = Activation_Softmax()
#create loss function
loss_function = Loss_CategoricalCrossentropy()
#create some variables to track the best loss and the associated weights and biases
lowest_loss = 9999999
best_dense1_weights = dense1.weights.copy()
best_dense1_biases = dense1.biases.copy()
best_dense2_weights = dense2.weights.copy()
best_dense2_biases = dense2.biases.copy()
# we initialize weights to random in each epoch, and later we update the weights
# if the loss decreases.
for iteration in range(10000):
#generate a new set of weights for iteration
# HERE WE TAKE A FRACTION OF THE WEIGHTS
dense1.weights += 0.05 * np.random.randn(2, 3)
dense1.biases += 0.05 * np.random.randn(1, 3)
dense2.weights += 0.05 * np.random.randn(3, 3)
dense2.biases += 0.05 * np.random.randn(1, 3)
# perform a forward pass of the training data through this layer
dense1.forward(X)
activation1.forward(dense1.output)
dense2.forward(activation1.output)
activation2.forward(dense2.output)
# calculating loss
loss = loss_function.calculate(activation2.output, y)
# calculate accuracy from output of activation2 and targets
predictions = np.argmax(activation2.output, axis=1)
accuracy = np.mean(predictions==y)
# if loss is smaller - print print and save weights and biases aside
if loss < lowest_loss:
print('New set of weights found, iteration:', iteration,
'loss:', loss, 'acc:', accuracy)
best_dense1_weights = dense1.weights.copy()
best_dense1_biases = dense1.biases.copy()
best_dense2_weights = dense2.weights.copy()
best_dense2_biases = dense2.biases.copy()
lowest_loss = loss
else:
dense1.weights = best_dense1_weights.copy()
dense1.biases = best_dense1_biases.copy()
dense2.weights = best_dense2_weights.copy()
dense2.biases = best_dense2_biases.copy()
# + [markdown] id="KwrSGSN9MfE_"
# # We see here that this training is not succesful. Later, we'll learn that the most probable reason for this is a local minimum of loss.
| Chapters/Chapter6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Dependencies and Setup
# SQL Alchemy
from sqlalchemy import create_engine
# Pandas
import pandas as pd
# Matplotlib
import matplotlib.pyplot as plt
# NumPy
import numpy as np
# -
# 1.Import the SQL database into Pandas.
# Create Engine and Pass in Postgres Connection
# Setup to Connect to Database
engine = create_engine('postgresql://postgres:{password}:5432/Employees_DB')
conn = engine.connect()
from sqlalchemy import create_engine
#engine = create_engine('postgresql://localhost:5432/<your_db_name>')
#connection = engine.connect()
employees_data = pd.read_sql("SELECT * FROM employees", conn)
employees_data.head()
# 2.Create a histogram to visualize the most common salary ranges for employees.
salaries_data = pd.read_sql("SELECT salary FROM salaries",conn)
salaries_data.head()
print(salaries_data.min())
print(salaries_data.max())
salaries_data.hist(bins=3)
plt.ylabel("Frequency")
plt.xlabel("Employee Salary Range")
plt.savefig('SalaryRange_hist.png')
plt.show()
# +
#3. Create a bar chart of average salary by title.
# -
avg_salary_df = pd.read_sql("""
SELECT title, AVG(salary) as "AvgSalary"
FROM salaries AS s
INNER JOIN employees AS e ON s.emp_no = e.emp_no
INNER JOIN titles as t ON t.title_id = e.emp_title_id
GROUP by title
""", conn)
avg_salary_df.head()
# +
# Set x_axis, y_axis & Tick Locations
plt.figure(figsize=(10,5))
x_axis = avg_salary_df['title']
y_axis = avg_salary_df['AvgSalary']
ticks = np.arange(len(x_axis))
plt.bar(x_axis,y_axis,align='center',alpha=0.5, color=['gray','red','green','blue','cyan','purple','orange'])
# Create Ticks for Bar Chart's x_axis
plt.xticks(ticks, x_axis, rotation=45)
# Set Labels & Title
plt.ylabel("Average Salary")
plt.xlabel("Employee Title")
plt.title("Average Salary by Employee Title")
# Save Figure
plt.savefig("AvgSalaryByTitle_bar.png")
# Show plot
plt.show()
| EmployeeSQL/EmployeeSalary_bonus.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.insert(0,'C:\\Users\\<NAME>\\PycharmProjects\\NET_Solver\\')
import numpy as np
import torch
from geometry import *
from utils import Plot_Grid
from solver import *
from models import *
from mesh import *
from boundary import *
import matplotlib.pyplot as plt
# %matplotlib inline
from models.neural_net import _init_weights
nx, ny = 80, 80
xi, eta = np.linspace(0,1,nx), np.linspace(0,1,ny)
# xi, eta = np.meshgrid(xi_, eta_)
# anulus = Analytical_Annulus(1., 0.6, 0.5)
# grid = TFI(xi, eta, anulus)
# +
#Plot_Grid(grid.X()['x'],grid.Y()['y'],nx,ny)
# +
# Code the governing Equation
# give the value of tau, n and K
tau = 0.5
n = 0.5
K = 1.5
dpdz = 10
class Guv_Eq(PDE):
def PDELoss(self, inputs, outputs):
# Calculate the determinant
#jac_det = self.jacobian_det(computation)
# calculate alpha
#alpha = self.alpha(computation)
# calculate beta
#beta = self.beta(computation)
# calculate gamma
#gamma = self.gamma(computation)
# compute the gradient of U wrt inputs
gradient = self.compute_grad(outputs, inputs)
# seperate dudxi and dudeta
dudxi = gradient[:,0]
dudeta = gradient[:,1]
# calculate apparent viscosity
app_visc = self.HB_model(gradient, tau, n, K)
# code part(a) of the governing equation
left = (app_visc)*(dudxi)
# code part(b) of the governing equation
right = (app_visc)*(dudeta)
# differentiate part(a) wrt xi
d_left_dxi = self.compute_grad(left, inputs)[:,0]
# differentiate part(b) wrt eta
d_right_deta = self.compute_grad(right, inputs)[:,1]
# combine the left and right with dpdz for loss function
gov_eq = d_left_dxi + d_right_deta + dpdz
return {'PDE': gov_eq}
def shear_rate(self, gradient):
# calculate the determinant
#jac_det = self.jacobian_det(computation)
# seperate the input
#dxdxi, dxdeta, dydxi, dydeta = self.seperate_data(computation)
# seperate the gradients
dudxi = gradient[:,0]
dudeta = gradient[:,1]
# compute the shear rate
sr_ = (((dudxi)**2 + (dudeta)**2)**0.5)
#print(sr_)
sr = torch.where(sr_ >torch.from_numpy(np.array([1e-2])).float(),sr_ , torch.from_numpy(np.array([1e-2])).float())
return sr
def Modified_HB(self, inputs, outputs):
pass
def HB_model(self, gradients,tau, n, K):
# determine the shear rate
shear_rate = self.shear_rate(gradients)
# calculate the apparent viscosity
app_vis_ = torch.abs_((tau/shear_rate) + ((shear_rate)**(n-1))*K)
#app_vis = torch.where(app_vis_>torch.from_numpy(np.array([1e-2])).float(),app_vis_,torch.from_numpy(np.array([1e-2])).float())
return app_vis_
# initiate the PDE equation
pde = Guv_Eq(inputs=('xi', 'eta'), outputs='u')
# -
# define the meshing data
mesh = Mesh({'xi':xi, 'eta':eta})
pde.set_mesh(mesh)
# when u = 0 @ eta = 0,1
eta_ = np.array([0,1])
u_ = np.full(2*len(xi), 0)
pipe = Dirichlet({'xi':xi, 'eta':eta_}, {'u':u_}, name='Pipe')
pde.add_boundary(pipe)
# when dudxi = 0 @ xi = 0,1
xi_ = np.array([0,1])
class Neumann(Neumann):
def neumann_loss(self, inputs, outputs):
dudxi = self.compute_grad(outputs, inputs)[:,0]
return {'gradXi': dudxi}
outer = Neumann({'xi':xi_, 'eta':eta}, name='left_flank')
pde.add_boundary(outer)
# +
btach_size = 1024
lr = 0.0001
epochs = 1000
hid_dim = 256
layers = 21
in_dim = len(pde.input)
out_dim = len(pde.output)
#activation = Sine()
activation = torch.nn.Tanh()
net = MLP(in_dim, out_dim, layers, hid_dim, act=activation)
#net.apply(_init_weights)
optimizer = torch.optim.Adam(net.parameters())
scheduler = torch.optim.lr_scheduler.OneCycleLR(optimizer, max_lr = lr, pct_start=0.1, total_steps=epochs)
#scheduler = torch.optim.lr_scheduler.CyclicLR(optimizer, base_lr=0.01, max_lr=0.1,cycle_momentum=True)
pde.compile(net,optimizer, scheduler)
# -
# %time hist = pde.solve(epochs, btach_size)
plt.figure(figsize=(10,8))
for key, value in hist.items():
if key!= 'lr':
plt.plot(hist[key],label=key)
plt.legend()
else:
pass
plt.yscale('log')
# +
x = np.linspace(0,1,60)
y = np.linspace(0,1,60)
eval_mesh = Mesh({'x':x, 'y':y})
pred = pde.eval(eval_mesh).view(len(y),len(x)).numpy()
# -
xi, eta = np.meshgrid(x,y)
plt.figure(figsize=(10,8))
plt.contourf(xi, eta, pred, 100, cmap='jet')
plt.colorbar()
plt.axis('equal');
pred
| NET_Solver/annulus/debugging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# !pip install --upgrade covid19dh
import pandas as pd
import numpy as np
import datetime as dt
import dateutil.relativedelta as rd
# Display all the columns for the dataframes (not-truncated)
pd.set_option("display.max_columns", None)
# Format floats better
pd.options.display.float_format = '{:,.4f}'.format
from covid19dh import covid19
X, src = covid19(verbose = False)
X
X.to_csv("Resources/covid_data.csv")
| Data_Extraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Water Boiler Demo
import openmc
import water_boiler
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# +
sol_temp=303.
sol_conc=0.299;
U_enrch=0.1467;
cr_wd = 0.1; # cm, distance the control rod is withdrawn from the core
material, geometry, tallies, settings = water_boiler.generate_model(sol_temp,
sol_conc,
U_enrch,cr_wd);
# -
material.export_to_xml();
geometry.export_to_xml();
tallies.export_to_xml();
settings.export_to_xml();
openmc.run()
material, geometry, tallies, settings = water_boiler.generate_model(sol_temp,
sol_conc,U_enrch,
cr_wd=76.);
material.export_to_xml();
geometry.export_to_xml();
tallies.export_to_xml();
settings.export_to_xml();
openmc.run()
sp_filename = "statepoint." + str(settings.batches) + ".h5"
sp = openmc.StatePoint(sp_filename);
# +
flux = sp.get_tally(name='flux');
flux_df = flux.get_pandas_dataframe();
flux_vals = flux_df['mean'].to_numpy();
N = 1001;
energy_bins = np.logspace(-3,7,num=N);
energy_x = 0.5*(energy_bins[0:-1] + energy_bins[1:]);
plt.loglog(energy_x,flux_vals);
plt.grid();
plt.xlabel('Energy [eV]');
plt.ylabel('flux [n/cm**2-s]');
# +
OMC_DATA = "/home/sblair/OMC_DATA/endfb71_hdf5"
u238_path = OMC_DATA + "/U238.h5";
u238 = openmc.data.IncidentNeutron.from_hdf5(u238_path);
u238_capture = u238[102];
s32_path = OMC_DATA + "/S32.h5";
s32 = openmc.data.IncidentNeutron.from_hdf5(s32_path);
s32_capture = s32[102];
u234_path = OMC_DATA + "/U234.h5";
u234 = openmc.data.IncidentNeutron.from_hdf5(u234_path);
u234_capture = u234[102];
plt.rcParams['figure.figsize']=[12,8];
plt.loglog(energy_x,flux_vals,label='flux');
plt.loglog(s32_capture.xs['294K'].x,s32_capture.xs['294K'].y,label='S-32');
plt.loglog(u238_capture.xs['294K'].x,u238_capture.xs['294K'].y,label='U-238');
plt.loglog(u234_capture.xs['294K'].x,u234_capture.xs['294K'].y,label='U-234');
plt.grid();
plt.legend();
plt.xlabel('Energy [eV]');
plt.title('Capture Cross-Sections and Flux vs Energy');
# +
capture_by_nuclide = sp.get_tally(name='capture_by_nuclide');
capture_by_nuclide_df = capture_by_nuclide.get_pandas_dataframe();
capture_U234 = capture_by_nuclide_df[capture_by_nuclide_df['nuclide']=='U234']['mean'].to_numpy();
capture_U238 = capture_by_nuclide_df[capture_by_nuclide_df['nuclide']=='U238']['mean'].to_numpy();
capture_H1 = capture_by_nuclide_df[capture_by_nuclide_df['nuclide']=='H1']['mean'].to_numpy();
capture_S32 = capture_by_nuclide_df[capture_by_nuclide_df['nuclide']=='S32']['mean'].to_numpy();
capture_O16 = capture_by_nuclide_df[capture_by_nuclide_df['nuclide']=='O16']['mean'].to_numpy();
plt.rcParams['figure.figsize']=[12,8];
plt.loglog(energy_x,flux_vals,label='flux');
plt.loglog(energy_x,capture_U234,label='U-234');
plt.loglog(energy_x,capture_U238,label='U-238');
plt.loglog(energy_x,capture_H1,label='H-1');
plt.loglog(energy_x,capture_S32,label='S-32');
plt.loglog(energy_x,capture_O16,label='O-16');
plt.grid();
plt.legend();
plt.xlabel('Energy [eV]');
plt.title('Capture Reaction Rates and Flux vs. Energy');
# -
| examples/water_boiler/base_case/wb_demo/.ipynb_checkpoints/water_boiler_demo-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from matplotlib import style
style.use('fivethirtyeight')
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import datetime as dt
# # Reflect Tables into SQLAlchemy ORM
# Python SQL toolkit and Object Relational Mapper
import sqlalchemy
from sqlalchemy.ext.automap import automap_base
from sqlalchemy.orm import Session
from sqlalchemy import create_engine, func
engine = create_engine("sqlite:///Resources/hawaii.sqlite")
# reflect an existing database into a new model
Base = automap_base()
# reflect the tables
Base.prepare(engine, reflect=True)
# We can view all of the classes that automap found
Base.classes.keys()
# Save references to each table
Measurement = Base.classes.measurement
Station = Base.classes.station
# Create our session (link) from Python to the DB
session = Session(engine)
# # Exploratory Climate Analysis
# +
today_date = session.query(Measurement.date).order_by(Measurement.date.desc()).first()
print(today_date)
# +
today_date = list(np.ravel(today_date))[0]
today_date = dt.datetime.strptime(today_date, '%Y-%m-%d')
print(today_date)
# -
last_year = today_date - dt.timedelta(days=365)
print(last_year)
# +
shower = session.query(Measurement.date, Measurement.prcp).\
filter(Measurement.date > last_year).\
order_by(Measurement.date).all()
shower
# -
rain_df = pd.DataFrame(shower)
rain_df.set_index('date').head()
rain_df.plot('date', 'prcp')
plt.xlabel("Date")
plt.ylabel("Rain Per Inches")
plt.title("Pecipitation for Year")
plt.xticks(rotation=45)
plt.show()
# 
# +
# Use Pandas to calcualte the summary statistics for the precipitation data
rain_df.describe()
# -
# 
# Design a query to show how many stations are available in this dataset?
Stations = session.query(Measurement).group_by(Measurement.station).count()
print("There are {} stations.".format(Stations))
# +
Stations = session.query(Measurement).filter(Measurement.station)
print(Stations)
# +
MostActive_station = (session.query(Measurement.station, func.count(Measurement.tobs)).
group_by(Measurement.station).\
order_by(func.count(Measurement.tobs).desc()).all())
for station, count in MostActive_station:
print("Stations:",station," Weather Observations: ",count)
# -
# What are the most active stations? (i.e. what stations have the most rows)?
# List the stations and the counts in descending order.
# Using the station id from the previous query, calculate the lowest temperature recorded,
# highest temperature recorded, and average temperature most active station?
# +
# Calculate the highest temprature
stationID = MostActive_station[0][0]
highest_temp = (session.query(Measurement.tobs)
.filter(Measurement.station == stationID)
.order_by(Measurement.tobs.desc())
.first())[0]
print(f'Hightest Temperature: {highest_temp} ')
# +
# Calculate the lowest temprature
stationID = MostActive_station[0][0]
lowest_temp = (session.query(Measurement.tobs)
.filter(Measurement.station == stationID)
.order_by(Measurement.tobs.asc())
.first())[0]
print(f'lowest Temperature: {lowest_temp} ')
# +
# Calculate the Average temprature
avg_temp = (session.query(func.avg(Measurement.tobs))
.filter(Measurement.station == stationID))
print(f'Average Temperature:', avg_temp[0][0])
# -
# Choose the station with the highest number of temperature observations.
# Query the last 12 months of temperature observation data for this station and plot the results as a histogram
# +
# Choose the station with the highest number of temperature observations.
St_High_Temp = (session.query(Measurement.date, Measurement.tobs)
.filter(Measurement.date > last_year)
.filter(Measurement.station == stationID)
.order_by(Measurement.date)
.all())
# Query the last 12 months of temperature observation data for this station and plot the results as a histogram
df = pd.DataFrame(St_High_Temp)
temp = df.plot(kind='hist', bins=12, title=f'Temperature Frequency')
temp.set_xlabel('Temperature (F)')
temp = temp.get_figure()
plt.tight_layout()
# -
# 
# +
# This function called `calc_temps` will accept start date and end date in the format '%Y-%m-%d'
# and return the minimum, average, and maximum temperatures for that range of dates
def calc_temps(start_date, end_date):
"""TMIN, TAVG, and TMAX for a list of dates.
Args:
start_date (string): A date string in the format %Y-%m-%d
end_date (string): A date string in the format %Y-%m-%d
Returns:
TMIN, TAVE, and TMAX
"""
return session.query(func.min(Measurement.tobs), func.avg(Measurement.tobs), func.max(Measurement.tobs)).\
filter(Measurement.date >= start_date).filter(Measurement.date <= end_date).all()
# function usage example
print(calc_temps('2012-02-28', '2012-03-05'))
# +
# Use your previous function `calc_temps` to calculate the tmin, tavg, and tmax
# for your trip using the previous year's data for those same dates.
# +
# this is using the previous year data for those same dates:
date_arrived = dt.date(2011, 2, 28)
date_depart = dt.date(2011, 3, 5)
last_year = dt.timedelta(days=365)
temp_avg = (calc_temps((date_arrived-last_year), (date_depart-last_year)))
print(temp_avg)
# -
# Plot the results from your previous query as a bar chart.
# Use "Trip Avg Temp" as your Title
# Use the average temperature for the y value
# Use the peak-to-peak (tmax-tmin) value as the y error bar (yerr)
# +
# Plot the results from your previous query as a bar chart.
# Use "Trip Avg Temp" as your Title
# Use the average temperature for the y value
# Use the peak-to-peak (tmax-tmin) value as the y error bar (yerr)
y_error = temp_avg[0][2] - temp_avg[0][0]
y = [temp_avg[0][1]]
x = 0
fig = plt.figure(figsize=(20,10))
fig, ax = plt.subplots()
ax.set_ylabel("Temperature (F)", fontsize=20)
ax.set_title("Trip Avg Temp", fontsize=30)
ax.bar(x, y, width=1, color="orange", yerr=y_error)
ax.set_xlim(-1, 1)
ax.set_ylim(0, 80)
ax.set_xbound(lower= -1, upper= 1)
ax.tick_params(axis='x', which='both', bottom='off', top='off', labelbottom='off')
plt.show()
# -
# +
# Calculate the total amount of rainfall per weather station for your trip dates using the previous year's matching dates.
# Sort this in descending order by precipitation amount and list the station, name, latitude, longitude, and elevation
# -
# ## Optional Challenge Assignment
# +
# Create a query that will calculate the daily normals
# (i.e. the averages for tmin, tmax, and tavg for all historic data matching a specific month and day)
def daily_normals(date):
"""Daily Normals.
Args:
date (str): A date string in the format '%m-%d'
Returns:
A list of tuples containing the daily normals, tmin, tavg, and tmax
"""
sel = [func.min(Measurement.tobs), func.avg(Measurement.tobs), func.max(Measurement.tobs)]
return session.query(*sel).filter(func.strftime("%m-%d", Measurement.date) == date).all()
daily_normals("01-01")
# +
# calculate the daily normals for your trip
# push each tuple of calculations into a list called `normals`
# Set the start and end date of the trip
# Use the start and end date to create a range of dates
# Stip off the year and save a list of %m-%d strings
# Loop through the list of %m-%d strings and calculate the normals for each date
# -
# Load the previous query results into a Pandas DataFrame and add the `trip_dates` range as the `date` index
# Plot the daily normals as an area plot with `stacked=False`
| climate_starter - Copy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #Chopsticks!
#
# A few researchers set out to determine the optimal length of chopsticks for children and adults. They came up with a measure of how effective a pair of chopsticks performed, called the "Food Pinching Performance." The "Food Pinching Performance" was determined by counting the number of peanuts picked and placed in a cup (PPPC).
#
# ### An investigation for determining the optimum length of chopsticks.
# [Link to Abstract and Paper](http://www.ncbi.nlm.nih.gov/pubmed/15676839)
# *the abstract below was adapted from the link*
#
# Chopsticks are one of the most simple and popular hand tools ever invented by humans, but have not previously been investigated by [ergonomists](https://www.google.com/search?q=ergonomists). Two laboratory studies were conducted in this research, using a [randomised complete block design](http://dawg.utk.edu/glossary/whatis_rcbd.htm), to evaluate the effects of the length of the chopsticks on the food-serving performance of adults and children. Thirty-one male junior college students and 21 primary school pupils served as subjects for the experiment to test chopsticks lengths of 180, 210, 240, 270, 300, and 330 mm. The results showed that the food-pinching performance was significantly affected by the length of the chopsticks, and that chopsticks of about 240 and 180 mm long were optimal for adults and pupils, respectively. Based on these findings, the researchers suggested that families with children should provide both 240 and 180 mm long chopsticks. In addition, restaurants could provide 210 mm long chopsticks, considering the trade-offs between ergonomics and cost.
#
# ### For the rest of this project, answer all questions based only on the part of the experiment analyzing the thirty-one adult male college students.
# Download the [data set for the adults](https://www.udacity.com/api/nodes/4576183932/supplemental_media/chopstick-effectivenesscsv/download), then answer the following questions based on the abstract and the data set.
#
# **If you double click on this cell**, you will see the text change so that all of the formatting is removed. This allows you to edit this block of text. This block of text is written using [Markdown](http://daringfireball.net/projects/markdown/syntax), which is a way to format text using headers, links, italics, and many other options. You will learn more about Markdown later in the Nanodegree Program. Hit shift + enter or shift + return to show the formatted text.
# #### 1. What is the independent variable in the experiment?
# Chopstick length
# #### 2. What is the dependent variable in the experiment?
# The Food.Pinching.Efficiency (title in csv file) or PPPC (according to introduction) which is the measure of food-pinching performance.
#
# #### 3. How is the dependent variable operationally defined?
#
# The number of peanuts picked and placed in a cup. Presumably this is a rate per unit time since the values given in the csv file are not integers.
# #### 4. Based on the description of the experiment and the data set, list at least two variables that you know were controlled.
# Think about the participants who generated the data and what they have in common. You don't need to guess any variables or read the full paper to determine these variables. (For example, it seems plausible that the material of the chopsticks was held constant, but this is not stated in the abstract or data description.)
#
# Each group has the same gender and similar age (e.g. 31 male junior college students is one group). This means the age and gender were matched. Matching age could eliminate the effects of reduced flexibility, agility or mental focus that might be present in older subjects. Matching gender may be important because smaller hands (among women) may be better suited to smaller chopsticks.
# One great advantage of ipython notebooks is that you can document your data analysis using code, add comments to the code, or even add blocks of text using Markdown. These notebooks allow you to collaborate with others and share your work. For now, let's see some code for doing statistics.
# +
import pandas as pd
# pandas is a software library for data manipulation and analysis
# We commonly use shorter nicknames for certain packages. Pandas is often abbreviated to pd.
# hit shift + enter to run this cell or block of code
# +
path = r'/Users/pradau/Dropbox/temp/Downloads/chopstick-effectiveness.csv'
# Change the path to the location where the chopstick-effectiveness.csv file is located on your computer.
# If you get an error when running this block of code, be sure the chopstick-effectiveness.csv is located at the path on your computer.
dataFrame = pd.read_csv(path)
dataFrame
# -
# Let's do a basic statistical calculation on the data using code! Run the block of code below to calculate the average "Food Pinching Efficiency" for all 31 participants and all chopstick lengths.
dataFrame['Food.Pinching.Efficiency'].mean()
# This number is helpful, but the number doesn't let us know which of the chopstick lengths performed best for the thirty-one male junior college students. Let's break down the data by chopstick length. The next block of code will generate the average "Food Pinching Effeciency" for each chopstick length. Run the block of code below.
# +
meansByChopstickLength = dataFrame.groupby('Chopstick.Length')['Food.Pinching.Efficiency'].mean().reset_index()
meansByChopstickLength
# reset_index() changes Chopstick.Length from an index to column. Instead of the index being the length of the chopsticks, the index is the row numbers 0, 1, 2, 3, 4, 5.
# -
# #### 5. Which chopstick length performed the best for the group of thirty-one male junior college students?
# For the 31 male college students the best length was 240mm.
#
# +
# Causes plots to display within the notebook rather than in a new window
# %pylab inline
import matplotlib.pyplot as plt
plt.scatter(x=meansByChopstickLength['Chopstick.Length'], y=meansByChopstickLength['Food.Pinching.Efficiency'])
# title="")
plt.xlabel("Length in mm")
plt.ylabel("Efficiency in PPPC")
plt.title("Average Food Pinching Efficiency by Chopstick Length")
plt.show()
# -
# #### 6. Based on the scatterplot created from the code above, interpret the relationship you see. What do you notice?
#
# There appears to be a peak efficiency at 240mm and less efficiency for either smaller or longer lengths. It could be that the curve has a secondary peak at 300mm but it is difficult to determine without sampling at additional lengths.
# ### In the abstract the researchers stated that their results showed food-pinching performance was significantly affected by the length of the chopsticks, and that chopsticks of about 240 mm long were optimal for adults.
#
# #### 7a. Based on the data you have analyzed, do you agree with the claim?
# Yes.
#
# #### 7b. Why?
# First we must accept that the operational definition of food-pinching performance is reasonable, which it appears to be from this brief description. Then we can do a test to see if length shows any effect on efficiency.
# A paired t-test for the 240mm and 330mm lengths (2 tailed, alpha =0.01) is significant with p= 5.4e-6.
# Therefore the null hypothesis (mean efficiency in PPPC is equal for the 240mm and 330mm lengths) must be rejected, for this pair.
# We can conclude that chopstick does have a significant effect on efficiency in PPPC.
| Data_Analyst_ND_Project0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="kzAOca2rjxS3"
# # Clase 9. Plotly
# + colab={"base_uri": "https://localhost:8080/"} id="hG_TOzlZllcH" outputId="a222e6d6-a97c-4c11-b42a-88c299cee24a"
pip install wbgapi
# + colab={"base_uri": "https://localhost:8080/"} id="iGZt6olclxWx" outputId="85b6b61f-4fe2-408b-b7b8-2caa9953f318"
pip install wooldridge
# + id="kEMlRGGZloMz"
import wbgapi as wb
import wooldridge as wd
import pandas as pd
import numpy as np
import plotly.express as px
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="LYZ8kA8DluUC" outputId="60a9a39d-2687-4609-c281-df92e213d4f2"
wb.source.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 544} id="juh3Pl4al-Tk" outputId="26d1027e-8d33-4961-d5e1-68ffaac2c0d2"
wb.search('gini')
# + id="A_vnx_DxmeF3"
gini = wb.data.DataFrame('SI.POV.GINI', time=2018, labels=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="0z_0FxhLWNia" outputId="4dfdbf7f-6152-4f96-e83a-2eb9356b3780"
gini
# + id="6-B25NvIm0_H"
gini.dropna(inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="KtUkK3v8WYRX" outputId="14644e7d-3cd7-4e93-e928-4462e077cdbe"
gini
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="fly7sn8knIPl" outputId="4755a0ef-537e-4a85-c2be-77b370fafd6b"
wb.search('GDP')
# + id="zTHtLY9znZZl"
pib_per_capita = wb.data.DataFrame('NY.GDP.PCAP.KD', time=2018, labels=True)
# + colab={"base_uri": "https://localhost:8080/"} id="tb1M_U9PquPP" outputId="7df2afff-6494-47a3-a63d-e1c847ffc49b"
pib_per_capita['NY.GDP.PCAP.KD']
# + id="056_B0PCrE6d"
df = (pib_per_capita.
merge(gini, how='inner').
rename(columns={'NY.GDP.PCAP.KD':'PPC','SI.POV.GINI':'gini'}))
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="t8NGwZkSW6j2" outputId="10770604-d868-464e-965e-85484f17f983"
df.head()
# + id="OhpwLjTFU092"
wage = wd.data('wage1')
wage = (wage.
replace({'female':{1:'F',0:'M'}}).
rename(columns={'female':'sexo'}))
# + [markdown] id="P2LB3c7iU5GE"
# ## Scatter
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="G5RtpgD9sSGb" outputId="e58824a6-5af3-4aab-b433-b75f3b736c40"
fig = px.scatter(data_frame=df, x='PPC', y='gini')
fig.show()
# + id="rqnaBESYVN17"
fig = px.scatter(data_frame=wage, x='exper', y='lwage' )
fig.show()
# + id="srbymXXrVglM"
fig = px.scatter(data_frame=wage, x='exper', y='lwage', color='sexo', hover_data=['educ'])
fig.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="zHIY8RqdWsHI" outputId="d53a6b9e-1678-4eba-cadf-df715f0b45db"
fig = px.scatter(data_frame=df, x='PPC', y='gini')
fig.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="MintAzzCWr7a" outputId="254ba10c-8fa7-44ed-bf22-475e7981d811"
fig = px.scatter(data_frame=df, x='PPC', y='gini', text='Country')
fig.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="c71At2zBWznH" outputId="2d143e6e-0fc1-40f5-c3eb-9bce3618e784"
fig = px.scatter(df, x='PPC',y='gini', text= 'Country')
fig.update_traces(textposition='top center')
fig.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="bLp102KeVNmd" outputId="472b0bd0-b2aa-433c-e9da-408a6c0ed697"
fig = px.scatter(data_frame=df, x='PPC', y='gini', hover_data=['Country'])
fig.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 426} id="DrE9KYTFVNVl" outputId="0ea5c6f3-534e-4f6e-f199-8cc38ec88936"
wb.search('population')
# + id="X72FRPFhXuzv"
urb_pop = wb.data.DataFrame('SP.URB.TOTL', time=2018, labels=True)
urb_pop = (urb_pop.
rename(columns={'SP.URB.TOTL':'population'}))
df = df.merge(urb_pop, on='Country', how='inner')
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="XXDHTLfAYQUc" outputId="29988298-4dee-40d4-9407-5ff8a0edbb4b"
px.scatter(data_frame=df, x='PPC', y='gini', hover_data=['Country'],size='population')
# + id="VPiOgtmnae9k"
wb.search('expectancy')
# + id="KARbIvtdaesW"
life = wb.data.DataFrame('SP.DYN.LE00.IN', time=2018,labels=True)
life = (life.
rename(columns={'SP.DYN.LE00.IN':'life_expect'}).
dropna())
# + id="_nij164Ldjpm"
df = df.merge(life, how='inner', on='Country')
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="NfyCKiqUbULa" outputId="2e7d361a-1bfa-459e-93f8-ae5e8a038db7"
fig = px.scatter(data_frame=df,
x='PPC',
y='gini',
size='life_expect',
hover_data=['Country'])
fig.show()
# + id="TuR5T-bHewnx"
df['minmax_life'] = (df['life_expect'] - df['life_expect'].min()) / (df['life_expect'].max() - df['life_expect'].min())
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="rcU0b8C_e-Ze" outputId="8e961236-ef39-430e-8daa-7126ea2f38fb"
fig = px.scatter(data_frame=df,
x='PPC',
y='gini',
size='minmax_life',
hover_data=['Country', 'life_expect'])
fig.show()
# + [markdown] id="eWg11k43g-Bf"
# ## Histogramas
# + id="h2DE4LJHg9LY"
fig = px.histogram(data_frame = wage, x='lwage')
fig.show()
# + id="iO9G58bhg9BX"
fig = px.histogram(data_frame = wage, x='wage')
fig.show()
# + id="g-x0T01zg82s"
fig = px.histogram(data_frame = wage, x='wage', nbins=50)
fig.show()
# + id="cIh3zEYQg8q5"
fig = px.histogram(data_frame=wage,
x='lwage',
color='sexo'
)
fig.show()
# + id="JRw_SOiag8f5"
fig = px.histogram(data_frame=wage,
x='lwage',
color='sexo',
marginal= 'rug'
)
fig.show()
# + id="04lyF2sAg8V6"
fig = px.histogram(data_frame=wage,
x='lwage',
color='sexo',
marginal= 'box'
)
fig.show()
# + id="HT2Sjy_ug7_e"
fig = px.histogram(data_frame=wage,
x='lwage',
color='sexo',
marginal= 'violin'
)
fig.show()
# + [markdown] id="44yuFzxziY6Q"
# ## Line
# + id="KT_mIRCZ1hN0"
datos = px.data.gapminder()
datos.info()
# + id="WspmlfiHlxY0"
datos.head()
# + id="lOD6-xK2l0HP"
datos['continent'].unique()
# + id="qhFwYlicmAFD"
datos = (datos.query("continent=='Americas'").
query("country=='Colombia' or country=='Chile'"))
# + id="YyYk8F80nSVG"
datos
# + id="0l_ghiwtmi5k"
fig = px.line(datos, x='year',y='lifeExp',color='country')
fig.show()
# + [markdown] id="EevYF-7FoDqL"
# ## Box-plots
# + id="4qKWPp4snstg"
fig = px.box(wage, x='sexo', y='lwage')
fig.show()
# + id="CyvzLXrVkAix"
fig = px.box(wage, x='sexo', y='lwage', points='all')
fig.show()
| clase_9.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Function dftmatrix
#
# ## Synopse
#
# Kernel matrix for the 1-D Discrete Fourier Transform DFT.
#
# - **A = dftmatrix(N)**
#
# - **A**: Output image, square N x N, complex
#
#
# - **N**: Integer, number of points of the DFT
# + deletable=true editable=true
import numpy as np
def dftmatrix(N):
x = np.arange(N).reshape(N,1)
u = x
Wn = np.exp(-1j*2*np.pi/N)
A = (1./np.sqrt(N)) * (Wn ** u.dot(x.T))
return A
# + [markdown] deletable=true editable=true
# ## Examples
# + deletable=true editable=true
testing = (__name__ == "__main__")
if testing:
import numpy as np
import sys,os
ia898path = os.path.abspath('../../')
if ia898path not in sys.path:
sys.path.append(ia898path)
import ia898.src as ia
# + [markdown] deletable=true editable=true
# ### Example 1
# + deletable=true editable=true
if testing:
A = ia.dftmatrix(128)
ia.adshow(ia.normalize(A.real),'A.real')
ia.adshow(ia.normalize(A.imag),'A.imag')
# + [markdown] deletable=true editable=true
# Example 2
# ---------
# + deletable=true editable=true
if testing:
A = ia.dftmatrix(4)
print('A=\n', A.round(1))
print('A-A.T=\n', A - A.T)
print((np.abs(np.linalg.inv(A)-np.conjugate(A))).max() < 10E-15)
# + [markdown] deletable=true editable=true
# ### Example 3
#
# Showing the product $\mathbf{x}\mathbf{u}^T$:
# + deletable=true editable=true
if testing:
u = x = np.arange(10).reshape(10,1)
print('u xT=\n', u.dot(x.T))
# + [markdown] deletable=true editable=true
# ## Equation
#
#
# $$ \begin{matrix}
# W_N &=& \exp{\frac{-j2\pi}{N}} \\ A_N &=& \frac{1}{\sqrt{N}} (W_N)^{\mathbf{u} \mathbf{x}^T} \\ \mathbf{u} &=& \mathbf{x} = [0, 1, 2, \ldots, N-1]^T
# \end{matrix} $$
# + [markdown] deletable=true editable=true
# $$ \begin{matrix}
# A_N &=& A_N^T \ \mbox{symmetric} \\
# (A_N)^{-1} &=& (A_N)^*\ \mbox{column orthogonality, unitary matrix}
# \end{matrix} $$
# + [markdown] deletable=true editable=true
# ## See Also
#
# - `dft` - Discrete Fourier Transform
# - `dftmatrixexamples` - Visualization of the DFT matrix
#
# ## References
#
# - http://en.wikipedia.org/wiki/DFT_matrix
# + deletable=true editable=true
if testing:
print('testing dftmatrix')
print(repr(np.floor(0.5 + 10E4*ia.dftmatrix(4).real) / 10E4) == repr(np.array(
[[ 0.5, 0.5, 0.5, 0.5],
[ 0.5, 0. , -0.5, 0. ],
[ 0.5, -0.5, 0.5, -0.5],
[ 0.5, 0. , -0.5, 0. ]])))
print(repr(np.floor(0.5 + 10E4*ia.dftmatrix(4).imag) / 10E4) == repr(np.array(
[[ 0. , 0. , 0. , 0. ],
[ 0. , -0.5, 0. , 0.5],
[ 0. , 0. , 0. , 0. ],
[ 0. , 0.5, 0. , -0.5]])))
# + deletable=true editable=true
| src/dftmatrix.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## the code we wrote for creating the data.
# ## https://github.com/anryabrahamyan/Road_Traffic_Prediction/
# + colab={"base_uri": "https://localhost:8080/"} id="G00HVpGVjsHd" outputId="a78be7dc-2007-46fb-8e8c-7b3c8120fba2"
import pandas as pd
import matplotlib.pyplot as plt
from statsmodels.tsa.stattools import kpss,adfuller
from statsmodels.graphics.tsaplots import plot_acf,plot_pacf
from statsmodels.tsa.arima.model import ARIMA
from statsmodels.tsa.statespace.sarimax import SARIMAX
from statsmodels.tsa.holtwinters import ExponentialSmoothing
import statsmodels.api as sm
import pmdarima as pm
import numpy as np
from datetime import timedelta
# + colab={"base_uri": "https://localhost:8080/", "height": 418} id="gvtKyBDekUIK" outputId="423303b0-a641-4f80-cf3e-18b5e641a47c"
df = pd.read_csv("../dataset/data.csv")
df.datetime = pd.to_datetime(df.datetime,format = "%d/%m/%Y %H:%M:%S")
df.index = df.datetime
df.drop(columns = ['datetime'], inplace = True)
df.head()
# -
df = df.replace("None", np.nan)
df.iloc[72:144,45] = np.nan
df_cleaned =pd.DataFrame( index=pd.date_range(df.index.min(),df.index.max(),freq='min'))
df_cleaned.index = df_cleaned.index.round(freq='min')
df.index = df.index.round(freq='min')
df_without_na = df[~df['Vehicle_0.1'].isna()]
df_cleaned['Vehicle_0.1'] = np.nan
df_cleaned.loc[df_without_na.index,'Vehicle_0.1']=df_without_na['Vehicle_0.1']
df_cleaned.fillna(method = 'ffill',inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 345} id="HsPz8u64lQq0" outputId="119a6a6d-14d6-467d-f446-14195ebc4730"
plt.figure(figsize=(20,8))
plt.plot(df_cleaned["Vehicle_0.1"].astype(float))
plt.show()
# -
# From the plot we can see that there is no trend, however there is some seasonality because the frequency of the cars decreases after 12am.
# + colab={"base_uri": "https://localhost:8080/", "height": 414} id="hg5Ykt9ul_Rx" outputId="9bea1ec7-29cf-4571-b6f9-caf0237b0e5a"
X = df_cleaned["Vehicle_0.1"]
result = adfuller(X)
print('ADF Statistic: %f' % result[0])
print('p-value: %f' % result[1])
print('Critical Values:')
for key, value in result[4].items():
print('\t%s: %.3f' % (key, value))
# -
# We can see that p value of ADF statistics is almost 0 which is smaller than 0.05, thus we reject the null hypothesis and conclude that our series is stationary.
# + colab={"base_uri": "https://localhost:8080/", "height": 380} id="__d639REmfrH" outputId="020a5ce1-c67d-401e-ea2f-0ba82868597a"
result = kpss(X)
print('KPSS Statistic: %f' % result[0])
print('p-value: %f' % result[1])
print('number of lags: %f' % result[2])
print('Critical Values:')
for key, value in result[3].items():
print('\t%s: %.3f' % (key, value))
# -
# We have p-value equal to 0.1, which is larger than 0.05, thus we can say that we have stationary time series.
# + colab={"base_uri": "https://localhost:8080/", "height": 597} id="t1icigMkmrFf" outputId="3b0f938d-54ba-4660-950d-6920a56431f1"
plot_acf(X, lags=20)
plot_pacf(X, lags=20)
plt.show()
# -
# From the ACF plot we can see that all lags are significant up to lag 20 for the MA process but to reduce the complexity of the model, we will choose lag as 1 for the MA process. In PACF we have significant lags until lag 2 and then cuts off, so we will choose lag as 2.
# + id="jW3_4nM6muyY"
total_observations = len(X)
train = X[:int(0.8*total_observations)]
test = X[int(0.8*total_observations):]
# -
model = ARIMA(train.astype(float),order=(2, 0, 1)).fit()
print(model.summary())
predictions = model.forecast(len(test))
residuals = test.astype(float) - predictions
plt.figure(figsize=(10,4))
plt.plot(residuals)
plt.axhline(0, linestyle='--', color='red')
plt.title('Residuals from ARIMA Model', fontsize=22)
plt.ylabel('Error', fontsize=17)
Lj_Box = sm.stats.acorr_ljungbox(residuals, lags = 10,return_df = True)
plt.plot(Lj_Box.index, Lj_Box.lb_pvalue)
# We can see that the p-value for different lags is less than 0.05 and almost 0, so we can reject the null hyppthesis, meaning that the errors are not independent.
#WE do not consider the seasonal component because the order of dependence for 1 day is 1440 and it will take forever to fit that model. Further aggregations of the data will help with this problem.
model_auto=pm.auto_arima(
train,
test='adf',
max_p = 12,
max_d = 1,
max_q = 12,
trace=True,
error_action='ignore',
suppress_warnings=True,
stepwise=True,
n_fits=120,
maxiter=11,
n_jobs=-1)
print('MSE of the AUTOARIMA model is', np.mean((model_auto.predict(len(test))-test.astype("float"))**2))
print('MSE of the previously chosen model is', np.mean((model.forecast(len(test)) - test.astype("float")) **2))
# We can observe that the MSE of the previously chosen model is lower, meaning that it is better if we take into consideration MSEs.
model = ARIMA(X.astype(float),order=(5,0,1)).fit()
predictions = model.forecast(1440)#24 hours
fig = plt.figure(figsize = (10,10))
plt.plot(X,'blue')
plt.plot(predictions,c='red')
plt.show()
exp_smooth = ExponentialSmoothing(train.astype(float),trend=None, seasonal = 'add',initialization_method='estimated',seasonal_periods=120).fit()
# ## Car traffic is generally stationary in terms of trend, and the seasonality is additive because the variance is not increasing over time.
predictions = exp_smooth.forecast(len(test))
print('MSE of the exp_smoothing model is', np.mean((predictions.values-test.astype("float"))**2))
# ## The exponential smoothing model is worse than the Arima model from before.
exp_smooth = ExponentialSmoothing(df_cleaned.astype(float),trend=None, seasonal = 'add',initialization_method='estimated',seasonal_periods=120).fit()
ax = plt.subplot()
ax.plot(exp_smooth.forecast(120))
ax.plot(df_cleaned.astype(float))
ax.plot(model.forecast(120))
plt.show()
| predictor/HW4_project_part_without_aggregation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## load DII plain text of patents
import os,glob
def file_load_f(input_path):
'''input_path'''
file_path = []
for dir_name, dirs, files in os.walk(input_path):
if len(files) > 0:
for input_file in (glob.glob(dir_name + '\*.txt')):
file_path.append(input_file)
return file_path
# ## extract patent ab infor
def ab_collection_f(file):
ab_data =[]
ab_temp = []
for line in file.readlines():
if line[0:2] == 'AB':
is_in = 1
start_line = ([line[6:-1]])
ab_temp.append(start_line)
elif line[0:3] == ' ' and is_in == 1:
follow_line = line[3:-1]
ab_temp.append(start_line.append(follow_line))
else:
is_in = 0
for x in ab_temp:
if x != None :
ab_data.append(x)
return ab_data
# ## extract patent ti infor
def ti_collection_f(file_path):
file = open(file_path, encoding='utf-8')
ti_collection=[]
for line in file.readlines():
if line.startswith('TI '):
ti_collection.append(str(line[3:-1]))
return ti_collection
# ## divide the ab infor to "NOVELTY" "USE" "ADVANTAGE"
def ab_mess_process_f(ab_data_slice):
ad_extract=[]
i = 0
if len(ab_data_slice)<3:
mark = 3 - len(ab_data_slice)
for x in ab_data_slice:
if x.strip().startswith('(') and 'NOVELTY' in x.strip():
ad_extract.append(x.strip().split(')')[1].strip())
ad_extract.append('miss')
ad_extract.append('miss')
continue
if x.strip().startswith('NOVELTY'):
ad_extract.append(x)
elif x.strip().startswith('USE'):
ad_extract.append(x)
elif x.strip().startswith('ADVANTAGE'):
ad_extract.append(x)
else:
ad_extract.append('miss')
for i_add in range(mark):
ad_extract.append('miss')
else:
for x in ab_data_slice:
if x.strip().startswith('(') and 'NOVELTY' in x.strip():
ad_extract.append(x.strip().split(')')[1].strip())
ad_extract.append('miss')
ad_extract.append('miss')
continue
if x.strip().startswith('NOVELTY'):
ad_extract.append(x)
elif x.strip().startswith('USE'):
ad_extract.append(x)
elif x.strip().startswith('ADVANTAGE'):
ad_extract.append(x)
else:
ad_extract.append('miss')
i += 1
if i == 3:
break
return ad_extract
# ## massive ab infor process
def ab_process_f(ab_data):
ad_normmalize = []
for x in ab_data:
#print(len(x))
ad_normmalize.append(ab_mess_process_f(x))
return ad_normmalize
# ## create CSV file
def createListCSV_ab(fileName, header=None, data_List=[]):
with open(fileName, 'w', newline='') as csvFile:
import csv
csvWriter = csv.writer(csvFile)
if header:
csvWriter.writerow(header)
for data in data_List:
for y in data:
csvWriter.writerow([y[0].split('- ')[-1],y[1].split('- ')[-1],y[2].split('- ')[-1]])
csvFile.close
def createListCSV_ti(fileName, header=None, data_List=[]):
with open(fileName, 'w', newline='') as csvFile:
import csv
csvWriter = csv.writer(csvFile)
if header:
csvWriter.writerow(header)
for data in data_List:
if not isinstance(data,list):
csvWriter.writerow([data])
csvFile.close
# # application
input_path = r''#input_path--where your file is stored such as "C:\Users\bandk\Desktop\graphene_dii\2012 " a folder path
otput_path = r''#output_path--where your file will be output in CSV such as "C:\Users\bandk\Desktop\graphene_dii\2012_out_ab " a folder path
file_path =file_load_f(input_path)
ti_data =[]
for f in file_path:
file = open(f, 'r', encoding='UTF-8')
ab_data_temp = ab_process_f(ab_collection_f(file))
ab_data.append(ab_data_temp)
ti_data_temp = ti_collection_f(file)
ti_data = ti_data+ti_data_temp
header = ['NOVELTY','USE','ADVANTAGE']
createListCSV_ab(os.path.join(output_path,'ab_extract.csv'),header,ab_data)
header = ['TI']
createListCSV_ti(os.path.join(output_path,'ti_extract.csv'),header,ab_data)
| data_pre_process.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import time
import matplotlib.pyplot as plt
import numpy as np
import datetime
import pickle
import torch as torch
import torchvision
import mongoengine as mo
from torchvision import transforms
from torchvision import datasets
try:
from configcarrot import *
except ImportError:
login = '' # Please provide your login to MongoDB
password = '' # Please provide your password to MongoDB
host = '' # Please provide your host name to MongoDB
# +
transforms_image = transforms.Compose([transforms.Resize(32),
transforms.CenterCrop(32),
transforms.ToTensor()])
train_xray = torch.utils.data.DataLoader(datasets.ImageFolder('chest_xray/train',
transform=transforms_image),
batch_size=20, shuffle=True)
def imshow(img):
npimg = img.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# obtain one batch of training images
dataiter = iter(train_xray)
images, _ = dataiter.next() # _ for no labels
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(20, 4))
plot_size=20
for idx in np.arange(plot_size):
ax = fig.add_subplot(2, plot_size/2, idx+1, xticks=[], yticks=[])
imshow(images[idx])
# +
image_transforms = {'train': transforms.Compose([transforms.Resize(224),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225])]),
'valid': transforms.Compose([transforms.Resize(224),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225])]),
'test': transforms.Compose([transforms.Resize(224),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225])])
}
loaders = {'train': torch.utils.data.DataLoader(datasets.ImageFolder('chest_xray/train',
transform=image_transforms['train']),
batch_size=128, shuffle=True),
'valid': torch.utils.data.DataLoader(datasets.ImageFolder('chest_xray/val',
transform=image_transforms['valid']),
batch_size=128, shuffle=True),
'test': torch.utils.data.DataLoader(datasets.ImageFolder('chest_xray/test',
transform=image_transforms['test']),
batch_size=128, shuffle=True)}
# +
import torch.nn as nn
import torch.nn.functional as F
# define the CNN architecture
class Net(nn.Module):
### TODO: choose an architecture, and complete the class
def __init__(self):
super(Net, self).__init__()
## Define layers of a CNN
self.c1 = nn.Conv2d(in_channels=3, out_channels=32, kernel_size=3)
self.c2 = nn.Conv2d(in_channels=32, out_channels=64, kernel_size=3)
self.c3 = nn.Conv2d(in_channels=64, out_channels=128, kernel_size=3)
self.fc1 = nn.Linear(in_features=6272, out_features=512)
self.fc2 = nn.Linear(in_features=512, out_features=256)
self.fc3 = nn.Linear(in_features=256, out_features=412)
self.fc4 = nn.Linear(in_features=412, out_features=2)
def forward(self, x):
## Define forward behavior
x = F.relu(F.max_pool2d(self.c1(x), 3))
x = F.relu(F.max_pool2d(self.c2(x), 3))
x = F.relu(F.max_pool2d(self.c3(x), 3))
x = x.view(x.size(0), -1)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = F.relu(self.fc3(x))
x = self.fc4(x)
return x
# -
model = Net()
model.cuda()
# +
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr = 0.0005)
# -
mo.disconnect_all()
mo.connect(host='mongodb+srv://' + login + ':' + password + '@' + host,
authentication_source='admin')
class Carrot(mo.Document):
date_time = mo.DateTimeField(required=True)
epoch = mo.IntField()
train_loss = mo.FloatField()
test_loss = mo.FloatField()
train_accuracy = mo.FloatField()
test_accuracy = mo.FloatField()
parameters = mo.FileField()
gradients = mo.FileField()
def carrot(loader_train, loader_test, model=model, num_epoch=25, use_gpu = False):
# Checking GPU
if use_gpu:
model.cuda()
# Training
for epoch in range(1, num_epoch + 1):
train_loss = 0
test_loss = 0
train_accuracy = 0.0
test_accuracy = 0.0
parmeters = dict()
gradients = dict()
# Actual training
model.train()
for batch_idx, (data, target) in enumerate(loader_train):
if use_gpu:
data, target = data.cuda(), target.cuda()
output = model(data)
loss = criterion(output, target)
print('Loss: ', loss.item())
train_loss += loss.item()
optimizer.zero_grad()
loss.backward()
optimizer.step()
train_loss = train_loss / len(loader_train)
# Test accuracy calculation
model.eval()
print('Epoch: ', epoch)
total_correct = 0
total = 0
for batch_idx, (data, target) in enumerate(loader_train):
if use_gpu:
data, target = data.cuda(), target.cuda()
output = model(data)
loss = criterion(output, target)
print('Loss: ', loss.item())
max_arg_output = torch.argmax(output, dim=1)
total_correct += int(torch.sum(max_arg_output == target))
total += data.shape[0]
train_accuracy = total_correct/total
print('Training accuracy: {:.0%}'.format(train_accuracy))
total_correct = 0
total = 0
for batch_idx, (data, target) in enumerate(loader_test):
if use_gpu:
data, target = data.cuda(), target.cuda()
output = model(data)
loss = criterion(output, target)
test_loss += loss.item()
print('Loss: ', loss.item())
max_arg_output = torch.argmax(output, dim=1)
total_correct += int(torch.sum(max_arg_output == target))
total += data.shape[0]
test_loss = test_loss / len(loader_test)
test_accuracy = total_correct/total
print('Test accuracy: {:.0%}'.format(test_accuracy))
# Saving into MongoDB
td = Carrot(date_time=datetime.datetime.now())
td.epoch = epoch
td.train_loss = train_loss
td.test_loss = test_loss
td.train_accuracy = train_accuracy
td.test_accuracy = test_accuracy
parameters_dict = {k:v.cpu().numpy() for (k,v) in model.state_dict().items()}
td.parameters.put(pickle.dumps(parameters_dict), content_type = 'application/octet-stream')
keys = list(model.state_dict().keys())
gradients_dict = {keys[k]:v.grad.cpu().detach().numpy() for (k,v) in enumerate(model.parameters())}
td.gradients.put(pickle.dumps(gradients_dict), content_type = 'application/octet-stream')
td.save()
# You can save your model if you want to start training next time
# if total_correct/total > 0.8:
# torch.save(model.state_dict(), str(time.strftime("%Y%m%d_%H%M%S"))+'.pt')
carrot(model=model, loader_train=loaders['train'], loader_test=loaders['valid'], num_epoch=50, use_gpu = True)
# +
parameters_dict = 0
gradients_dict = 0
for batch_idx, (data, target) in enumerate(loaders['valid']):
if True:
data, target = data.cuda(), target.cuda()
output = model(data)
loss = criterion(output, target)
print('Loss: ', loss.item())
optimizer.zero_grad()
loss.backward()
optimizer.step()
parameters_dict = {k:v.cpu().numpy() for (k,v) in model.state_dict().items()}
keys = list(model.state_dict().keys())
gradients_dict = {keys[k]:v.grad.cpu().detach().numpy() for (k,v) in enumerate(model.parameters())}
# -
parameters_dict
gradients_dict
list(model.state_dict().keys())
| mongo_old/Carrot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cap_env
# language: python
# name: cap_env
# ---
# # This notebook contains a resumed table of the q-learners results. The results are the ones evaluated on the test set, with the learned actions (without learning on the test set)
# +
# Basic imports
import os
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import sys
from time import time
import pickle
# %matplotlib inline
# %pylab inline
pylab.rcParams['figure.figsize'] = (20.0, 10.0)
# %load_ext autoreload
# %autoreload 2
sys.path.append('../../')
# -
# # Predictor Results
pred_results_df = pd.DataFrame({
1.0: {'train_r2': 0.983486, 'train_mre': 0.008762, 'test_r2': 0.976241, 'test_mre': 0.013906},
7.0: {'train_r2': 0.906177, 'train_mre': 0.026232, 'test_r2': 0.874892, 'test_mre': 0.034764},
14.0: {'train_r2': 0.826779, 'train_mre': 0.037349, 'test_r2': 0.758697, 'test_mre': 0.051755},
28.0: {'train_r2': 0.696077, 'train_mre': 0.052396, 'test_r2': 0.515802, 'test_mre': 0.078545},
56.0: {'train_r2': 0.494079, 'train_mre': 0.073589, 'test_r2': 0.152134, 'test_mre': 0.108190},
}).T
pred_results_df = pred_results_df[['train_r2', 'test_r2', 'train_mre', 'test_mre']]
pred_results_df.index.name = 'ahead_days'
pred_results_df
# # Automatic Trader Results
features = [
'dyna',
'states',
'actions',
'training_days',
'epochs',
'predictor',
'random_decrease',
]
metrics =[
'sharpe_ratio',
'cumulative_return',
'epoch_time'
]
# +
names = [
'simple_q_learner',
'simple_q_learner_1000_states',
'simple_q_learner_1000_states_4_actions_full_training',
'simple_q_learner_1000_states_full_training',
'simple_q_learner_100_epochs',
'simple_q_learner_11_actions',
'simple_q_learner_fast_learner',
'simple_q_learner_fast_learner_1000_states',
'simple_q_learner_fast_learner_11_actions',
'simple_q_learner_fast_learner_3_actions',
'simple_q_learner_fast_learner_full_training',
'simple_q_learner_full_training',
'dyna_q_1000_states_full_training',
'dyna_q_learner',
'dyna_q_with_predictor',
'dyna_q_with_predictor_full_training',
'dyna_q_with_predictor_full_training_dyna1',
]
feat_data = np.array([
# (dyna, states, actions, training_days, epochs, predictor, random_decrease)
[0, 125, 2, 512, 15, False, 0.9999], # simple_q_learner
[0, 1000, 2, 512, 15, False, 0.9999], # simple_q_learner_1000_states
[0, 1000, 4, 5268, 7, False, 0.9999], # simple_q_learner_1000_states_4_actions_full_training
[0, 1000, 2, 5268, 7, False, 0.9999], # simple_q_learner_1000_states_full_training
[0, 125, 2, 512, 100, False, 0.9999], # simple_q_learner_100_epochs
[0, 125, 11, 512, 10, False, 0.9999], # simple_q_learner_11_actions
[0, 125, 2, 512, 4, False, 0.999], # simple_q_learner_fast_learner
[0, 1000, 2, 512, 4, False, 0.999], # simple_q_learner_fast_learner_1000_states
[0, 125, 11, 512, 4, False, 0.999], # simple_q_learner_fast_learner_11_actions
[0, 125, 3, 512, 4, False, 0.999], # simple_q_learner_fast_learner_3_actions
[0, 125, 2, 5268, 4, False, 0.999], # simple_q_learner_fast_learner_full_training
[0, 125, 2, 5268, 15, False, 0.9999], # simple_q_learner_full_training
[20, 1000, 2, 5268, 7, False, 0.9999], # dyna_q_1000_states_full_training
[20, 125, 2, 512, 4, False, 0.9999], # dyna_q_learner
[20, 125, 2, 512, 4, True, 0.9999], # dyna_q_with_predictor
[20, 125, 2, 5268, 4, True, 0.9999], # dyna_q_with_predictor_full_training
[1, 125, 2, 5268, 4, True, 0.9999], # dyna_q_with_predictor_full_training_dyna1
])
experiments_df = pd.DataFrame(feat_data, columns=features, index=names)
experiments_df.index.name = 'nb_name'
train_res_data = {
'simple_q_learner': {'sharpe': 1.9858481612185834, 'cum_ret': 0.38359700000000174, 'epoch_time': 18.330891609191895},
'simple_q_learner_1000_states': {'sharpe': 3.4470302925746776, 'cum_ret': 0.7292610000000004, 'epoch_time': 18.28188133239746},
'simple_q_learner_1000_states_4_actions_full_training': {'sharpe': 2.2430093688893264, 'cum_ret': 30.14936200000002, 'epoch_time': 157.69741320610046},
'simple_q_learner_1000_states_full_training': {'sharpe': 2.2911390004146073, 'cum_ret': 87.25733300000006, 'epoch_time': 125.35012602806091},
'simple_q_learner_100_epochs': {'sharpe': 4.093353629096188, 'cum_ret': 0.6627280000000009, 'epoch_time': 9.004882335662842},
'simple_q_learner_11_actions': {'sharpe': 1.5440407782808305, 'cum_ret': 0.2412700000000001, 'epoch_time': 11.08903431892395},
'simple_q_learner_fast_learner': {'sharpe': 2.8787265519379908, 'cum_ret':0.5468269999999986, 'epoch_time': 18.931288242340088},
'simple_q_learner_fast_learner_1000_states': {'sharpe': 2.031446601959524, 'cum_ret': 0.3971230000000021, 'epoch_time': 19.006957530975342},
'simple_q_learner_fast_learner_11_actions': {'sharpe': 3.241438316121647, 'cum_ret': 0.541966, 'epoch_time': 18.913504123687744},
'simple_q_learner_fast_learner_3_actions': {'sharpe': 2.9448069674427555, 'cum_ret': 0.4873689999999995, 'epoch_time': 18.46741485595703},
'simple_q_learner_fast_learner_full_training': {'sharpe': 1.0444534903132408, 'cum_ret': 0.7844770000000019, 'epoch_time': 143.5039553642273},
'simple_q_full_training': {'sharpe': 1.2592450659232495, 'cum_ret': 1.7391450000000006, 'epoch_time': 115.70198798179626},
'dyna_q_1000_states_full_training': {'sharpe': 2.2964510954840325, 'cum_ret': 94.75696199999993, 'epoch_time': 242.88240551948547},
'dyna_q_learner': {'sharpe': 3.706435588713091, 'cum_ret': 0.4938250000000006
, 'epoch_time': 18.87182092666626},
'dyna_q_with_predictor': {'sharpe': 3.2884867210125845, 'cum_ret': 0.5397989999999993, 'epoch_time': 458.8401937484741},
'dyna_q_with_predictor_full_training': {'sharpe': 1.0037137587999854, 'cum_ret': 2.565081999999997, 'epoch_time': 7850.391056537628},
'dyna_q_with_predictor_full_training_dyna1': {'sharpe': 0.48228187419119906, 'cum_ret': 0.1737430000000002, 'epoch_time': 730.5918335914612},
}
train_res_data_df = pd.DataFrame(train_res_data).T
test_res_data_no_learning = {
'simple_q_learner': {'sharpe': 0.3664203166030617, 'cum_ret': 0.06372499999999937, 'epoch_time': 17.75287628173828},
'simple_q_learner_1000_states': {'sharpe': -0.013747768227987086, 'cum_ret': -0.013047000000000142, 'epoch_time': 17.661759853363037},
'simple_q_learner_1000_states_4_actions_full_training': {'sharpe': 0.9400492987950515, 'cum_ret': 0.10791900000000054, 'epoch_time': 13.83948016166687},
'simple_q_learner_1000_states_full_training': {'sharpe': 1.3139195350252832, 'cum_ret': 0.2099470000000001, 'epoch_time': 8.874800443649292},
'simple_q_learner_100_epochs': {'sharpe': 0.6420028402682839, 'cum_ret': 0.10032399999999986, 'epoch_time': 9.116246461868286},
'simple_q_learner_11_actions': {'sharpe': 0.15616450321809833, 'cum_ret': 0.019991000000000758, 'epoch_time': 10.187344551086426},
'simple_q_learner_fast_learner': {'sharpe': 0.9643510680410812, 'cum_ret': 0.18794100000000125, 'epoch_time': 18.13912320137024},
'simple_q_learner_fast_learner_1000_states': {'sharpe': 0.8228017709095453, 'cum_ret': 0.16162700000000063, 'epoch_time': 19.452654361724854},
'simple_q_learner_fast_learner_11_actions': {'sharpe': 0.8238261816524384, 'cum_ret': 0.12766000000000033, 'epoch_time': 18.901001930236816},
'simple_q_learner_fast_learner_3_actions': {'sharpe': 0.6332862559879147, 'cum_ret': 0.08036399999999966, 'epoch_time': 19.221533060073853},
'simple_q_learner_fast_learner_full_training': {'sharpe': 1.2605807833904492, 'cum_ret': 0.056606000000000156, 'epoch_time': 11.412826538085938},
'simple_q_full_training': {'sharpe': -0.2562905901467118, 'cum_ret': -0.027945999999999693, 'epoch_time': 8.009900569915771},
'dyna_q_1000_states_full_training': {'sharpe': 0.4267994866360769, 'cum_ret': 0.0652820000000005, 'epoch_time': 14.224964618682861},
'dyna_q_learner': {'sharpe': 0.5191712068491942, 'cum_ret': 0.07307299999999883, 'epoch_time': 16.431984901428223},
'dyna_q_with_predictor': {'sharpe': 0.7435489843809434, 'cum_ret': 0.10403399999999974, 'epoch_time': 6.692898988723755},
'dyna_q_with_predictor_full_training': {'sharpe': -0.33503797163532956, 'cum_ret': -0.029740999999999795, 'epoch_time': 8.51533818244934},
'dyna_q_with_predictor_full_training_dyna1': {'sharpe': 0.20288841658633258, 'cum_ret': 0.008380000000000276, 'epoch_time': 10.236766338348389},
}
test_res_data_no_learning_df = pd.DataFrame(test_res_data_no_learning).T
test_res_data_learning = {
'simple_q_learner': {'sharpe': 0.9735950444291429, 'cum_ret': 0.1953619999999998, 'epoch_time': 18.097697019577026},
'simple_q_learner_1000_states': {'sharpe': -0.0867440896667206, 'cum_ret': -0.027372000000001173, 'epoch_time': 17.762672901153564},
'simple_q_learner_1000_states_4_actions_full_training': {'sharpe': 1.109613523501088, 'cum_ret': 0.12868000000000057, 'epoch_time': 9.899595499038696},
'simple_q_learner_1000_states_full_training': {'sharpe': 0.45880561807905557, 'cum_ret': 0.05954300000000079, 'epoch_time': 9.082789421081543},
'simple_q_learner_100_epochs': {'sharpe': 0.09274627213069256, 'cum_ret': 0.008058000000000565, 'epoch_time': 8.653764009475708},
'simple_q_learner_11_actions': {'sharpe': 0.4691456599751897, 'cum_ret': 0.07124699999999917, 'epoch_time': 10.827114582061768},
'simple_q_learner_fast_learner': {'sharpe': 0.6020182964860242, 'cum_ret': 0.09249299999999816, 'epoch_time': 17.882429122924805},
'simple_q_learner_fast_learner_1000_states': {'sharpe': 0.17618139275375405, 'cum_ret': 0.02545300000000017, 'epoch_time': 15.724592685699463},
'simple_q_learner_fast_learner_11_actions': {'sharpe': 0.9608337022400049, 'cum_ret': 0.1406880000000006, 'epoch_time': 17.67305564880371},
'simple_q_learner_fast_learner_3_actions': {'sharpe': 0.3254406127664859, 'cum_ret': 0.04086700000000043, 'epoch_time': 18.100637197494507},
'simple_q_learner_fast_learner_full_training': {'sharpe': 1.2605807833904492, 'cum_ret': 0.056606000000000156, 'epoch_time': 12.214732885360718},
'simple_q_full_training': {'sharpe': 0.3139835605580342, 'cum_ret': 0.02497299999999969, 'epoch_time': 7.958802700042725},
'dyna_q_1000_states_full_training': {'sharpe': 0.48863969848043476, 'cum_ret': 0.06846099999999988, 'epoch_time': 18.820592880249023},
'dyna_q_learner': {'sharpe': 0.0700928915599047, 'cum_ret': 0.004358999999999114, 'epoch_time': 18.085463523864746},
'dyna_q_with_predictor': {'sharpe': 0.6954014537549168, 'cum_ret': 0.09154599999999946, 'epoch_time': 338.36568880081177},
'dyna_q_with_predictor_full_training': {'sharpe': -0.8531759696425502, 'cum_ret': -0.07708900000000052, 'epoch_time': 375.830899477005},
'dyna_q_with_predictor_full_training_dyna1': {'sharpe': -0.15635735184097058, 'cum_ret': -0.006745999999999919, 'epoch_time': 38.24271035194397},
}
test_res_data_learning_df = pd.DataFrame(test_res_data_learning).T
train_benchmark_data = {
'simple_q_learner': {'sharpe_bench': 1.601691549431671, 'cum_ret_bench': 0.4244923418116293},
'simple_q_learner_1000_states': {'sharpe_bench': 1.601691549431671, 'cum_ret_bench': 0.4244923418116293},
'simple_q_learner_1000_states_4_actions_full_training': {'sharpe_bench': 0.4566770027925799, 'cum_ret_bench': 3.304502617801047},
'simple_q_learner_1000_states_full_training': {'sharpe_bench': 0.4566770027925799, 'cum_ret_bench': 3.304502617801047},
'simple_q_learner_100_epochs': {'sharpe_bench': 1.601691549431671, 'cum_ret_bench': 0.4244923418116293},
'simple_q_learner_11_actions': {'sharpe_bench': 1.601691549431671, 'cum_ret_bench': 0.4244923418116293},
'simple_q_learner_fast_learner': {'sharpe_bench': 1.601691549431671, 'cum_ret_bench': 0.4244923418116293},
'simple_q_learner_fast_learner_1000_states': {'sharpe_bench': 1.601691549431671, 'cum_ret_bench': 0.4244923418116293},
'simple_q_learner_fast_learner_11_actions': {'sharpe_bench': 1.601691549431671, 'cum_ret_bench': 0.4244923418116293},
'simple_q_learner_fast_learner_3_actions': {'sharpe_bench': 1.601691549431671, 'cum_ret_bench': 0.4244923418116293},
'simple_q_learner_fast_learner_full_training': {'sharpe_bench': 0.4566770027925799, 'cum_ret_bench': 3.304502617801047},
'simple_q_full_training': {'sharpe_bench': 0.4566770027925799, 'cum_ret_bench': 3.304502617801047},
'dyna_q_1000_states_full_training': {'sharpe_bench': 0.4566770027925799, 'cum_ret_bench': 3.304502617801047},
'dyna_q_learner': {'sharpe_bench': 1.601691549431671, 'cum_ret_bench': 0.4244923418116293},
'dyna_q_with_predictor': {'sharpe_bench': 1.601691549431671, 'cum_ret_bench': 0.4244923418116293},
'dyna_q_with_predictor_full_training': {'sharpe_bench': 0.4566770027925799, 'cum_ret_bench': 3.304502617801047},
'dyna_q_with_predictor_full_training_dyna1': {'sharpe_bench': 0.4566770027925799, 'cum_ret_bench': 3.304502617801047},
}
train_benchmark_data_df = pd.DataFrame(train_benchmark_data).T
test_benchmark_data = {
'simple_q_learner': {'sharpe_bench': 0.44271542660031676, 'cum_ret_bench': 0.1070225832012679},
'simple_q_learner_1000_states': {'sharpe_bench': 0.44271542660031676, 'cum_ret_bench': 0.1070225832012679},
'simple_q_learner_1000_states_4_actions_full_training': {'sharpe_bench': 0.44271542660031676, 'cum_ret_bench': 0.1070225832012679},
'simple_q_learner_1000_states_full_training': {'sharpe_bench': 0.44271542660031676, 'cum_ret_bench': 0.1070225832012679},
'simple_q_learner_100_epochs': {'sharpe_bench': 0.44271542660031676, 'cum_ret_bench': 0.1070225832012679},
'simple_q_learner_11_actions': {'sharpe_bench': 0.44271542660031676, 'cum_ret_bench': 0.1070225832012679},
'simple_q_learner_fast_learner': {'sharpe_bench': 0.44271542660031676, 'cum_ret_bench': 0.1070225832012679},
'simple_q_learner_fast_learner_1000_states': {'sharpe_bench': 0.44271542660031676, 'cum_ret_bench': 0.1070225832012679},
'simple_q_learner_fast_learner_11_actions': {'sharpe_bench': 0.44271542660031676, 'cum_ret_bench': 0.1070225832012679},
'simple_q_learner_fast_learner_3_actions': {'sharpe_bench': 0.44271542660031676, 'cum_ret_bench': 0.1070225832012679},
'simple_q_learner_fast_learner_full_training': {'sharpe_bench': 0.44271542660031676, 'cum_ret_bench': 0.1070225832012679},
'simple_q_full_training': {'sharpe_bench': 0.44271542660031676, 'cum_ret_bench': 0.1070225832012679},
'dyna_q_1000_states_full_training': {'sharpe_bench': 0.44271542660031676, 'cum_ret_bench': 0.1070225832012679},
'dyna_q_learner': {'sharpe_bench': 0.44271542660031676, 'cum_ret_bench': 0.1070225832012679},
'dyna_q_with_predictor': {'sharpe_bench': 0.2930367522823553, 'cum_ret_bench': 0.05002151977428149},
'dyna_q_with_predictor_full_training': {'sharpe_bench': 0.2930367522823553, 'cum_ret_bench': 0.05002151977428149},
'dyna_q_with_predictor_full_training_dyna1': {'sharpe_bench': 0.3772011734533203, 'cum_ret_bench': 0.07288030223327424},
}
test_benchmark_data_df = pd.DataFrame(test_benchmark_data).T
# -
# ## Features that were used in the experiments
print(experiments_df.shape)
experiments_df
experiments_df.to_csv('../../data/experiments_df.csv')
# ## Training Results
training_res_df = train_res_data_df.join(train_benchmark_data_df)
training_res_df.index.name = 'nb_name'
training_res_df['sharpe_increase'] = training_res_df['sharpe'] - training_res_df['sharpe_bench']
training_res_df['cum_ret_increase'] = training_res_df['cum_ret'] - training_res_df['cum_ret_bench']
print(training_res_df.shape)
training_res_df
training_res_df.to_csv('../../data/training_res_df.csv')
# ## Test Results without learning in the test set
test_no_learn_res_df = test_res_data_no_learning_df.join(test_benchmark_data_df)
test_no_learn_res_df.index.name = 'nb_name'
test_no_learn_res_df['sharpe_increase'] = test_no_learn_res_df['sharpe'] - test_no_learn_res_df['sharpe_bench']
test_no_learn_res_df['cum_ret_increase'] = test_no_learn_res_df['cum_ret'] - test_no_learn_res_df['cum_ret_bench']
print(test_no_learn_res_df.shape)
test_no_learn_res_df
test_no_learn_res_df.to_csv('../../data/test_no_learn_res_df.csv')
# ## Test Results with learning in the test set (always keeping causality)
test_learn_res_df = test_res_data_learning_df.join(test_benchmark_data_df)
test_learn_res_df.index.name = 'nb_name'
test_learn_res_df['sharpe_increase'] = test_learn_res_df['sharpe'] - test_learn_res_df['sharpe_bench']
test_learn_res_df['cum_ret_increase'] = test_learn_res_df['cum_ret'] - test_learn_res_df['cum_ret_bench']
print(test_learn_res_df.shape)
test_learn_res_df
test_learn_res_df.to_csv('../../data/test_learn_res_df.csv')
# ## Sharpe increases resumed
SHARPE_Q = 'sharpe_increase'
sharpe_q_df = pd.DataFrame(training_res_df[SHARPE_Q]).rename(columns={SHARPE_Q:'sharpe_i_train'})
sharpe_q_df = sharpe_q_df.join(test_no_learn_res_df[SHARPE_Q].rename('sharpe_i_test_no_learn'))
sharpe_q_df = sharpe_q_df.join(test_learn_res_df[SHARPE_Q].rename('sharpe_i_test_learn'))
print(sharpe_q_df.shape)
sharpe_q_df
# ## Best Agent
# ### The Agent with the best sharpe test increase (learning or not learning) was chosen as the "best".
best_agent_name = 'simple_q_learner_1000_states_full_training'
pd.DataFrame(experiments_df.loc[best_agent_name]).T
# +
indexes = ['training', 'test_no_learn', 'test_learn']
best_agent_df = pd.concat([
training_res_df.loc[best_agent_name],
test_no_learn_res_df.loc[best_agent_name],
test_learn_res_df.loc[best_agent_name],
], axis=1).T
best_agent_df.index = indexes
best_agent_df
# -
| notebooks/prod/n11_results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ieor4575
# language: python
# name: ieor4575
# ---
# +
import torch
from collections import deque
import numpy as np
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
import gym
# -
import wandb
wandb.login()
run=wandb.init(project="lab4", entity="ieor-4575", tags=["torch"])
# ## Policy Gradient
#
# In the previous lab, we talked about value based method for reinforcement learning. In this lab, we focus on policy based method.
#
# In policy based methods, intead of defining a value function $Q_\theta(s,a)$ and inducing a policy based on argmax, we parameterize a stochastic policy directly. The policy is parameterized as a categorical distribution over actions. Let it be $\pi_\phi(s)$ with parameter $\phi$, then the policy is defined by sampling actions $$a \sim \pi_\phi(s)$$
#
# The policy induces a probability $p(\tau)$ over trajectories $\tau = \{s_0,a_0,s_1,a_1,..\}$. The expected total discounted reward is
#
# $$\rho(\phi) = \mathbb{E}_{\tau \sim p(\tau)} \big[R(\tau)\big] = \mathbb{E}_{\pi_\phi} \big[\sum_{t=0}^\infty r_t \gamma^t \big]$$
#
# The aim is to find $\phi$ such that the expected reward induced by $\pi_\phi$ is maximized.
# ### Policy Gradient Computation
#
# We can derive policy gradient
#
# $$\nabla_\phi \rho(\phi) = \mathbb{E}_{\pi} \big[\sum_{t=0}^\infty Q^{\pi_\phi}(s_t,a_t) \nabla_\phi \log \pi_\phi(s_t, a_t) \big]$$
#
# To compute the gradient for update $\phi \leftarrow \phi + \alpha \nabla_\phi \rho(\phi)$, we need to estimate $Q^{\pi_\phi}(s,a)$. Since $Q^{\pi_\phi}(s,a)$ is usually not analytically accessible, it can be approximated by
# 1. Monte Carlo estimate
# 2. Train a value function $Q_\theta(s,a) \approx Q^{\pi_\phi}(s,a)$ and use it as a proxy
# 3. Mixture of both above
#
# Before estimating $Q^{\pi_\phi}(s,a)$, let us write a parameterized policy over actions. The policy $\pi_\phi(s)$ takes a state as input and outputs a categorical distribution over actions. For example, if we have two actions, the probability vector to output is of the form $\pi_\phi(s)=[0.6,0.4]$.
#
# **Loss function**
# Given samples of state action pairs $(s_i, a_i)$ and estimate $Q_i$ for $Q^{\pi_\phi}(s_i, a_i)$, for $i=1,\ldots, $ we set the loss function as
# $$-\frac{1}{N} \sum_{i=1}^N Q_i \log(\pi_\phi(s_i, a_i)) $$
# The loss function enables us to compute policy gradients in implementation. The negative gradient of the above loss function has the form
#
# $$\frac{1}{N} \sum_{i=1}^N Q_i \nabla_\phi \log \pi_\phi(s_i, a_i) $$
#
# where $Q_i$s are estimated and $\nabla_\phi \log\pi_\phi(s_i, a_i)$s are computed via backprop.
# +
# define neural net \pi_\phi(s) as a class
class Policy(object):
def __init__(self, obssize, actsize, lr):
"""
obssize: size of the states
actsize: size of the actions
"""
# TODO DEFINE THE MODEL
self.model = torch.nn.Sequential(
#input layer of input size obssize
torch.nn.Linear(obssize, 256),
torch.nn.ReLU(),
torch.nn.Linear(256, actsize)
)
# DEFINE THE OPTIMIZER
self.optimizer = torch.optim.Adam(self.model.parameters(), lr=lr)
# RECORD HYPER-PARAMS
self.obssize = obssize
self.actsize = actsize
# TEST
self.compute_prob(np.random.randn(obssize).reshape(1, -1))
def compute_prob(self, states):
"""
compute prob distribution over all actions given state: pi(s)
states: numpy array of size [numsamples, obssize]
return: numpy array of size [numsamples, actsize]
"""
states = torch.FloatTensor(states)
prob = torch.nn.functional.softmax(self.model(states), dim=-1)
return prob.cpu().data.numpy()
def _to_one_hot(self, y, num_classes):
"""
convert an integer vector y into one-hot representation
"""
scatter_dim = len(y.size())
y_tensor = y.view(*y.size(), -1)
zeros = torch.zeros(*y.size(), num_classes, dtype=y.dtype)
return zeros.scatter(scatter_dim, y_tensor, 1)
def train(self, states, actions, Qs):
"""
states: numpy array (states)
actions: numpy array (actions)
Qs: numpy array (Q values)
"""
states = torch.FloatTensor(states)
actions = torch.LongTensor(actions)
Qs = torch.FloatTensor(Qs)
# COMPUTE probability vector pi(s) for all s in states
logits = self.model(states)
prob = torch.nn.functional.softmax(logits, dim=-1)
# Compute probaility pi(s,a) for all s,a
action_onehot = self._to_one_hot(actions, actsize)
prob_selected = torch.sum(prob * action_onehot, axis=-1)
# FOR ROBUSTNESS
prob_selected += 1e-8
# TODO define loss function as described in the text above
# loss = ....
#loss_function = torch.nn.CrossEntropyLoss()
#loss = loss_function(prob, Qs.flatten())
loss = - torch.mean(Qs * torch.log(prob_selected))
# BACKWARD PASS
self.optimizer.zero_grad()
loss.backward()
# UPDATE
self.optimizer.step()
return loss.detach().cpu().data.numpy()
# -
# Try to rollout trajecories using the policy
# +
# Below is a set of template code for running a policy to interact with the environment
# It initializes a policy and runs it
# Note that you may not be able to run the code properly if there still some undefined components on the Policy class
env = gym.make("CartPole-v0")
obssize = env.observation_space.low.size
actsize = env.action_space.n
policyinit=Policy(obssize, actsize, 0.1)
obs = env.reset()
done = False
while not done:
prob = policyinit.compute_prob(np.expand_dims(obs,0))
prob /= np.sum(prob) #normalizing again to account for numerical errors
action = np.asscalar(np.random.choice(actsize, p=prob.flatten(), size=1)) #choose according distribution prob
obs, reward, done, info = env.step(action)
# -
# ### Estimate $Q^\pi(s,a)$
#
# To estimate $Q^\pi(s,a)$, we can rollout the policy until the episode ends and do monte carlo estimate. In particular, under policy $\pi$, we start from state action $(s_0,a_0)$ and rollout the policy to generate a trajectory $\{s_0,a_0,s_1,a_1...s_T,a_T\}$, with corresponding reward $r_0,r_1...r_T$. Monte carlo estimate is
#
# $$\hat{Q}_{MC}(s,a) = \sum_{t=0}^T r_t \gamma^t \approx Q^\pi(s,a)$$
#
# This estimate by itself is of high variance. Using pure monte carlo estimate may work but the gradient can have large variance and hence take the algorithm a long time to converge. We can reduce variance using baseline. Recall the derivation of PG
#
# $$\nabla_\phi \rho(\phi) = \mathbb{E}_{\pi_\phi} \big[\sum_{t=0}^\infty Q^{\pi_\phi}(s_t,a_t) \nabla_\phi \log \pi_\phi(s_t, a_t) \big] = \mathbb{E}_{\pi_\phi} \big[\sum_{t=0}^\infty ( Q^{\pi_\phi}(s_t,a_t) - b(s_t)) \nabla_\phi \log \pi_\phi(s_t, a_t) \big]$$
#
# where $b(s_t)$ can be any function of state $s_t$. $b(s_t)$ is called baseline. Optimal baseline function is hard to compute, but a good proxy is the value function $V^{\pi_\phi}(s_t)$. Hence the gradient has the form
# $$\nabla_\phi \rho(\pi_{\phi}) = \mathbb{E}_{\pi} \big[\sum_{t=0}^\infty A^{\pi_\phi}(s_t,a_t) \nabla_\phi \log \pi_\phi(s_t, a_t) \big]$$
#
# where $A^{\pi_\phi}(s,a)$ is the advantage. Hence we can train a value function $V^{\pi_\phi}(s)$ along side the policy and use it as baseline to reduce the variance of PG.
# Hence we also parameterize a value function $V_\theta(s) \approx V^{\pi_\phi}(s)$ with parameter $\theta$ to serve as baseline. The function takes as input the states $s$ and outputs a real value.
#
# Notice that unlike DQN, where $Q_\theta(s,a) \approx Q^\ast(s,a)$, now we have $V_\theta(s) \approx V^{\pi_\phi}(s)$. Therefore, we have a moving target to approximate, that changes with the current policy $\pi_\phi$. As $\phi$ is updated by PG, $\pi_\phi$ keeps changing, which changes $V^{\pi_\phi}(s)$ as well. We need to adapt $V_\theta(s)$ online to cater for the change in policy.
#
# Recall that to evaluate a policy $\pi$, we collect rollouts using $\pi$. If we start with state $s_0$, the reward following $\pi$ thereafter is $r_0...r_{T}$ then
#
# $$V^\pi(s_0) \approx \sum_{t=0}^{T} r_t \gamma^{t} = \hat{V}(s_0)$$
#
# In general, given a trajectory $(s_0, a_0, s_1, a_1, r_1, s_2, a_2, r_2, ..., s_{T+1})$
#
# $$\hat{V}(s_i) = \sum_{i=t}^{T} r_i \gamma^{i-t}$$
#
# And the objective to minimize over is
# $$\frac{1}{T+1} \sum_{i=0}^{T} (V_\theta(s_i) - \hat{V}(s_i))^2$$
#
# Since the policy keeps updating, we do not need to minimize the above objective to optimality. In practice, taking one gradient step w.r.t. above objective suffices.
#
# In the code cell below, define the neural network to learn value function estimate. The implementation is similar to Qfunction class in lab3, except that inputs are only states, and not actions.
# +
# TODO: define value function as a class. You need to define the model and set the loss.
class ValueFunction(object):
def __init__(self, obssize, lr):
"""
obssize: size of states
"""
# TODO DEFINE THE MODEL
self.model = torch.nn.Sequential(
#TODO
#input layer of input size obssize
#intermediate layers
#output layer of output size 1
torch.nn.Linear(obssize, 256),
torch.nn.ReLU(),
torch.nn.Linear(256, 1)
)
# DEFINE THE OPTIMIZER
self.optimizer = torch.optim.Adam(self.model.parameters(), lr=lr)
# RECORD HYPER-PARAMS
self.obssize = obssize
self.actsize = actsize
# TEST
self.compute_values(np.random.randn(obssize).reshape(1, -1))
def compute_values(self, states):
"""
compute value function for given states
states: numpy array of size [numsamples, obssize]
return: numpy array of size [numsamples]
"""
states = torch.FloatTensor(states)
return self.model(states).cpu().data.numpy()
def train(self, states, targets):
"""
states: numpy array
targets: numpy array
"""
states = torch.FloatTensor(states)
targets = torch.FloatTensor(targets)
# COMPUTE Value PREDICTIONS for states
v_preds = self.model(states)
#print(v_preds.shape)
# LOSS
# TODO: set LOSS as square error of predicted values compared to targets
#loss_function = torch.nn.MSELoss()
#loss = loss_function(v_preds, targets)
loss = torch.mean( (v_preds.flatten() - targets.flatten())**2 )
#print(diff.shape)
# BACKWARD PASS
self.optimizer.zero_grad()
loss.backward()
# UPDATE
self.optimizer.step()
return loss.detach().cpu().data.numpy()
# -
# ### Summary of pseudocode
#
# The critical components of the pseudocode are as follows.
#
# **Collect trajectories** Given current policy $\pi_\phi$, we can rollout using the policy by executing $a_t \sim \pi_\phi(s_t)$.
#
# **Update value function** Value function update is based on minimizing the L2 loss between predicted value function and estimated value functions. For each state $s_i, i=0,\ldots, T$ in a trajectory of length $T+1$, compute $\hat{V}(s_i)$ as dicounted reward over the rest of the path (as defined above).
# Then take one gradient step to update $\theta$ using the gradient of the following loss:
#
# $$\frac{1}{T+1} \sum_{i=0}^T(V_\theta(s_i) - \hat{V}(s_i))^2$$
#
# For your conveience, below we have provided a function discounted_rewards(r,gamma) that takes as inputs a list of $T$ rewards $r$ and computes all discounted rewards $\hat V(s_i), i=0, 1, 2, \ldots, T$.
#
# **Update policy using PG** To compute PG, we need to first monte carlo estimate action-value function $\hat{Q}(s_i,a_i)$. Given a trajectory with rewards $r=[r_0, r_1, r_2, \ldots, r_T]$, this can also be computed for all $s_i, a_i$ in this trajectory using the discounted_rewards(r, gamma) function below.
#
# Then use value function as a baseline to compute advantage
#
# $$\hat{A}(s_i,a_i) = \hat{Q}(s_i,a_i) - V_\theta(s_i)$$
#
# Then compute surrogate loss
#
# $$L = - \frac{1}{(T+1)}\sum_{i} \hat{A}(s_i,a_i) \log \pi(a_i|s_i) $$
#
# The policy is updated by $$\phi \leftarrow \phi - \alpha \nabla_\phi L \approx \phi + \alpha \nabla_\phi \rho(\pi_\phi)$$
def discounted_rewards(r, gamma):
""" take 1D float array of rewards and compute discounted reward """
discounted_r = np.zeros_like(r)
running_sum = 0
for i in reversed(range(0,len(r))):
discounted_r[i] = running_sum * gamma + r[i]
running_sum = discounted_r[i]
return list(discounted_r)
# ### Main implementation : Policy gradient algorithm
#
# Combine all the above steps and implement the policy gradient algorithm with value function baseline in the cell below. The use of baseline is optional.
# +
# %%wandb
#remove the above line if you do not want to see wandb plots in your notebook. You can always see them on the wandb website.
#You can change the code in this cell anyway you want
#However, just make sure per epsiode reward during the run of this algorithm is being recorded in list rrecord
#and logged on wandb as in the last few lines.
# parameter initializations (you can change any of these)
alpha = 3e-3 # learning rate for PG
beta = 3e-2 # learning rate for baseline
numtrajs = 6 # num of trajecories from the current policy to collect in each iteration
iterations = 600 # total num of iterations
envname = "CartPole-v0" # environment name
gamma = .99 # discount
# initialize environment
env = gym.make(envname)
obssize = env.observation_space.low.size
actsize = env.action_space.n
action_space = np.arange(env.action_space.n)
# initialize networks
actor = Policy(obssize, actsize, alpha) # policy initialization: IMPORTANT: this is the policy you will be scored on
baseline = ValueFunction(obssize, beta) # baseline initialization
#To record training reward for logging and plotting purposes
rrecord = []
steps = 200
# main iteration
for ite in range(iterations):
# To record traectories generated from current policy
OBS = [] # observations
ACTS = [] # actions
ADS = [] # advantages (to compute policy gradient)
VAL = [] # Monte carlo value predictions (to compute baseline, and policy gradient)
total_r = 0
for num in range(numtrajs):
# To keep a record of states actions and reward for each episode
obss = [] # states
acts = [] # actions
rews = [] # instant rewards
obs = env.reset()
done = False
# TODO: run one episode using the current policy "actor"
# TODO: record all observations (states, actions, rewards) from the epsiode in obss, acts, rews
for step in range(steps):
prob = actor.compute_prob(obs)
prob /= np.sum(prob)
action = np.random.choice(action_space, p=prob.flatten())
newobs, r, done, _ = env.step(action)
obss.append(list(obs))
acts.append(action)
rews.append(r)
total_r += r
obs = newobs
if done:
break
#Below is for logging training performance
rrecord.append(np.sum(rews))
# TODO: Use discounted_rewards function to compute \hat{V}s/\hat{Q}s from instant rewards in rews
V_hat = discounted_rewards(rews, gamma)
# TODO: record the computed \hat{V}s in VAL, states obss in OBS, and actions acts in ACTS, for batch update
OBS.extend(obss)
ACTS.extend(acts)
VAL.extend(list(V_hat))
# AFTER collecting numtrajs trajectories:
#1. TODO: train baseline
"""
Use the batch (OBS, VAL) of states and value predictions as targets to train baseline.
Use baseline.train : note that this takes as input numpy array, so you may have to convert
lists into numpy array using np.array()
"""
baseline.train(np.array(OBS), np.array(VAL))
# 2.TODO: Update the policy
"""
Compute baselines: use basline.compute_values for states in the batch OBS
Compute advantages ADS using VAL and computed baselines
Update policy using actor.train using OBS, ACTS and ADS
"""
v_theta = baseline.compute_values(np.array(OBS))
ADS = np.array(VAL) - np.concatenate(v_theta)
actor.train(np.array(OBS), np.array(ACTS), ADS)
if ite % 50 == 0:
print("iter:", ite, " avg reward: ", round(total_r/numtrajs, 2) )
#printing moving averages for smoothed visualization.
#Do not change below: this assume you recorded the sum of rewards in each episide in the list rrecord
fixedWindow=100
movingAverage=0
if len(rrecord) >= fixedWindow:
movingAverage=np.mean(rrecord[len(rrecord)-fixedWindow:len(rrecord)-1])
#wandb logging
wandb.log({ "training reward" : rrecord[-1], "training reward moving average" : movingAverage})
# -
# Finally, we evaluate the performance of the trained agent. We will evaluate the performance of the trained policy. The evaluation will be run for 100 epsiodes and print out the average performance across these episodes. Please **do not** change the code below.
# +
# DO NOT CHANGE CODE HERE
### DO NOT CHANGE
def evaluate(policy, env, episodes):
# main iteration
score = 0
for episode in range(episodes):
obs = env.reset()
done = False
rsum = 0
while not done:
p = policy.compute_prob(np.expand_dims(obs,0)).ravel()
p /= np.sum(p)
action = np.asscalar(np.random.choice(np.arange(2), size=1, p=p))
# env stepping forward
newobs, r, done, _ = env.step(action)
# update data
rsum += r
obs = newobs
wandb.log({"eval reward" : rsum})
score +=rsum
score = score/episodes
return score
# -
# DO NOT CHANGE CODE HERE
# after training, we will evaluate the performance of the learned policy "actor"
# on a target environment
env_test = gym.make(envname)
eval_episodes = 1000
score = evaluate(actor, env_test, eval_episodes)
wandb.run.summary["score"]=score
print("eval performance of the learned policy: {}".format(score))
run.finish()
| labs/lab4_policy_gradient.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Net Learning Master**
#
# This notebook runs
# * struc-learning-template.ipynb
# * param-learning-template.ipynb
#
# for different networks.
#
# See nbrun-example folder for an example of how to use nbrun (nbrun=notebook run)
# + [markdown] toc="true"
# # Table of Contents
# <p><div class="lev1 toc-item"><a href="#earthquake" data-toc-modified-id="earthquake-1"><span class="toc-item-num">1 </span>earthquake</a></div><div class="lev1 toc-item"><a href="#WetGrass" data-toc-modified-id="WetGrass-2"><span class="toc-item-num">2 </span>WetGrass</a></div><div class="lev1 toc-item"><a href="#asia_" data-toc-modified-id="asia_-3"><span class="toc-item-num">3 </span>asia_</a></div>
# -
# Change working directory to quantum-fog and add it to path environment variable.
import os
import sys
master_dir_path = os.getcwd()
print(master_dir_path)
os.chdir('../../')
qfog_path = os.getcwd()
print(qfog_path)
sys.path.insert(0,qfog_path)
# import nbrun.py
# +
# this doesn't work because of hyphen:
# from jupyter-notebooks.nbrun import run_notebook
import importlib
mod = importlib.import_module("jupyter-notebooks.nbrun")
# -
# import qfog modules
from learning.RandGen_NetParams import *
from graphs.BayesNet import *
# +
# if use_tempo_csv=True (False, resp.) will write training data
# into a tempo.csv (<bnet_name>.csv, resp.) file
def do_net(do_struc= True, do_param=True, use_tempo_csv=True):
in_bif = 'examples_cbnets/' + bnet_name + '.bif'
in_dot = 'examples_cbnets/' + bnet_name + '.dot'
in_csv = 'learning/training_data_c/'
if not generate_training_data:
in_csv += bnet_name + '.csv'
assert os.path.exists(in_csv), "no training data for this net in training_data_c folder"
else:
if use_tempo_csv:
in_csv += 'tempo.csv'
else:
in_csv += bnet_name + '.csv'
print('generating training data')
is_quantum = False
use_int_sts = True
bnet = BayesNet.read_bif(in_bif, is_quantum)
gen = RandGen_NetParams(is_quantum, bnet, num_samples, use_int_sts)
gen.write_csv(in_csv)
nb_kwargs = {'in_bif': in_bif, 'in_dot': in_dot, 'in_csv': in_csv, 'qfog_path': qfog_path}
# nbrun.py assumes cwd is the dir of the master notebook
os.chdir(master_dir_path)
if do_struc:
template_name = 'struc-learning-template'
mod.run_notebook(template_name, nb_suffix='-' + bnet_name, nb_kwargs=nb_kwargs)
if do_param:
template_name = 'param-learning-template'
mod.run_notebook(template_name, nb_suffix='-' + bnet_name, nb_kwargs=nb_kwargs)
# revert to preferred dir
os.chdir(qfog_path)
# -
# # earthquake
# +
bnet_name = 'earthquake'
generate_training_data = False # use False if training data already exists
num_samples = 5000
do_net()
# -
# # WetGrass
# +
bnet_name = 'WetGrass'
generate_training_data = False # use False if training data already exists
num_samples = 2000
do_net()
# -
# # asia_
# +
# asia_.bif is same as asia.bif but with hyphens replaced by underscores in state names
# because hyphens confuse current version of bnlearn.
bnet_name = 'asia_'
generate_training_data = True # use False if training data already exists
num_samples = 2000
do_net()
# + language="bash"
# rm -f learning/training_data_c/tempo.csv
| jupyter-notebooks/bnlearn-qfog-comparison/net-learning-master.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Least Squares Regression: Polynomial Curve Fitting
#
# This tutorial is meant to be a gentle introduction to machine learning. We present a simple polynomial fitting example using a least squares solution. We use this example to introduce important machine learning concepts.
#
# The goals of this tutorial are:
# - Explain how to develop an experiment. Split your data into development set and test set.
# - Introduce the concepts of *overfitting*, *underfitting*, and *model generalization*.
# - Introduce the *regularization* method for reducing *overfitting*.
#
#
# This tutorial is interactive and it corresponds to an adaptation of the example presented in chapter 1 of the book:
# **<NAME>. 2006. Pattern Recognition and Machine Learning (Information Science and Statistics). Springer-Verlag New York, Inc., Secaucus, NJ, USA.**
#
# ## Designing your experiment
#
# Machine learning consists of building models to explain your data. When designing your experiment, you need to split your data into a development set and a test set. The development set is split in 2 sets: a train set and validation set. The train set is used to learn the parameters of the different models you are training. The validation set is employed to select the best model among the different models you trained, therefore it has a bias and cannot be used as proof of generalization. The test set is used to see if the selected model generalizes well to unseen data.
#
# <img src="../Figures/train_val_test.png" alt="Drawing" style="width: 500px;"/>
#
# ## Generating synthetic data
# +
# Directive to make plots inline as opposed to having pop-up plots
# %matplotlib inline
import numpy as np # Import numpy with nickname np
import matplotlib.pylab as plt # plotting library
from ipywidgets import * # Interaction library
#Noise variance
var = 0.3
#Train set
train_size = 10
x_train = np.linspace(0,1,train_size)
#signal + noise
y_train = np.sin(2*np.pi*x_train) + np.random.normal(0,var,train_size)
#Val set
val_size = 100
x_val= np.linspace(0,1,val_size)
y = np.sin(2*np.pi*x_val)
y_val = y + np.random.normal(0,var,val_size) #signal + noise
plt.figure()
plt.plot(x_val,y,linewidth = 2.0,label = r'Model no noise: $sin(2 \pi x)$')
plt.scatter(x_train,y_train,color='red',label = "Model with noise")
plt.legend(loc = (0.02, 0.18))
plt.xlabel("x")
plt.ylabel("y")
plt.show()
# -
# ## Data
#
# Observations: $$\boldsymbol{X} =(x_1,x_2,...,x_N)^T$$
# Target: $$\boldsymbol{T} =(t_1,t_2,...,t_N)^T$$
#
#
# ## Model
#
# $$y(x,\boldsymbol{W})= w_0 + w_1x +w_2x^2+...+w_mx^m = \sum^M_{j=0}w_jx^j$$
#
# ## Cost Function
#
# Quadratic cost function: $$E(\boldsymbol{W})=\frac{1}{2}\sum_{n=1}^N\{y(x_n,\boldsymbol{W})-t_n\}^2$$
#
# Computing the derivative of the cost function and making it equal to zero, we can find the vector **W*** that minimizes the error:
# $$ \boldsymbol{W}^* = (\boldsymbol{A}^T\boldsymbol{A})^{-1}\boldsymbol{A} ^T\boldsymbol{T}$$
#
# Where **A** is defined by:
#
# $$\boldsymbol{A} = \begin{bmatrix}
# 1 & x_{1} & x_{1}^2 & \dots & x_{1}^M \\
# 1 & x_{2} & x_{2}^2 & \dots & x_{2}^M \\
# \vdots & \vdots & \vdots & \ddots & \vdots \\
# 1 & x_{N} & x_{N}^2 & \dots & x_{N}^M
# \end{bmatrix}$$
#Least squares polynomial fitting solution
# Implementation of the equation shown above in the explanation
def polynomial_fit(X,T,M):
A = np.power(X.reshape(-1,1),np.arange(0,M+1).reshape(1,-1))
T = T.reshape(-1,1)
W = np.dot(np.linalg.pinv(A),T)
return W.ravel()
# Plotting the least squares result varying the polynomial degree between 0 a 9. **Which model is a good model?**
# +
def plotmodel(M):
coefs = polynomial_fit(x_train, y_train, M)[::-1]
p = np.poly1d(coefs)
plt.figure()
plt.plot(x_val,y,linewidth = 2.0,label = 'Real')
plt.scatter(x_train,y_train,color='red',label= "Train")
plt.xlabel("x")
plt.ylabel(r'y')
y_fit = p(x_val)
plt.plot(x_val,y_fit,linewidth = 2.0,label ="Estimated")
plt.plot(x_val,y_val,'x',color='black',label = "Validation")
plt.legend(loc=(0.02,0.02))
plt.show()
interact(plotmodel,M=(0,9,1))
# -
# ## Root mean squared error
#
# $$E_{RMS}=\sqrt{2E(\boldsymbol{W^*})/N}$$
# +
# Computes RMS error
def rms_error(X,T,W):
p = np.poly1d(W)
T_fit = p(X)
E = np.sqrt(((T - T_fit)**2/T.size).sum())
return E
m = range(10)
train = []
val = []
for M in m:
W = polynomial_fit(x_train, y_train, M)[::-1]
error_train = rms_error(x_train,y_train,W)
error_val = rms_error(x_val,y_val,W)
train.append(error_train)
val.append(error_val)
plt.figure()
plt.plot(m,train,linewidth = 2.0,marker = 'o',markersize = 12,label = r'$E_{RMS}$ Train')
plt.plot(m,val,linewidth = 2.0,marker = 'x',markersize = 12,label = r'$E_{RMS}$ Validation')
plt.legend(loc = (0.02, 0.05))
plt.xlabel("Polynome degree")
plt.ylabel(r'$E_{RMS}$')
plt.show()
# -
# ## Cost function with regularization
#
# $$E(\boldsymbol{w})=\frac{1}{2}\sum_{n=1}^N\{y(x_n,\boldsymbol{w})-t_n\}^2 +\frac{\lambda}{2}||\boldsymbol{W}||^2$$
#
# The solution of the above equation is given by:
#
# $$\boldsymbol{W}^* = (\boldsymbol{A}^T\boldsymbol{A} + \lambda n\boldsymbol{I})^{-1}\boldsymbol{A} ^T\boldsymbol{T} $$
#Least square solution with regularization
def polynomial_fit_reg(X,T,M,lamb):
N = X.shape[0]
A = np.power(X.reshape(-1,1),np.arange(0,M+1).reshape(1,-1))
lambda_matrix = lamb*N*np.eye(M+1)
T = T.reshape(-1,1)
aux = np.dot(A.T,A) + lambda_matrix
aux = np.linalg.pinv(aux)
aux2 = np.dot(A.T,T)
W = np.dot(aux,aux2)
return W.ravel()
# In the demo below, we show the influence of $\lambda$ and $M$ in the polynomial fitting.
def plotmodel2(M,log_lamb):
lamb = np.exp(log_lamb)
coefs = polynomial_fit_reg(x_train, y_train, M,lamb)[::-1]
print "W:",coefs
print "Lambda", lamb
p = np.poly1d(coefs)
plt.figure()
plt.plot(x_val,y,linewidth = 2.0,label = 'Real')
plt.scatter(x_train,y_train,color='red',label= "Train")
plt.xlabel("x")
plt.ylabel(r'y')
y_fit = p(x_val)
plt.plot(x_val,y_fit,linewidth = 2.0,label ="Estimated")
plt.plot(x_val,y_val,'x',color='black',label = "Validation")
plt.legend(loc=(0.02,0.02))
plt.show()
interact(plotmodel2,M=(0,9,1),log_lamb = (-40,-9,.1))
# In the demo below, we show the influence of $\lambda$ in the fitting of a polynome of degree 9.
# +
log_lamb = range(-40,-9)
M = 9
train = []
val = []
for l in log_lamb:
lamb = np.exp(l)
coeffs = polynomial_fit_reg(x_train, y_train, M,lamb)[::-1]
error_train = rms_error(x_train,y_train,coeffs)
error_val = rms_error(x_val,y_val,coeffs)
train.append(error_train)
val.append(error_val)
plt.figure()
plt.plot(log_lamb,train,linewidth = 1.0,marker = 'o',markersize = 12,label = r'$E_{RMS}$ Train')
plt.plot(log_lamb,val,linewidth = 1.0,marker = 'x',markersize = 12,label = r'$E_{RMS}$ Validation')
plt.legend(loc = (0.02, 0.075))
plt.xlabel(r'$ln\lambda$')
plt.ylabel(r'$E_{RMS}$')
plt.title("Polynome degree 9")
plt.show()
# -
# At the beginning of this tutorial we said that when designing your experiment, you split your data into train, validation (train + validation = development) and test sets. We only illustrated the train and validation sets. The train data was used to find the parameters of the polynomial fitting. The validation set was used to select the model (i.e. the degree of the polynome). Now it is necessary to test the model on unseen data. See the suggested activities below.
#
# ## Activity suggestions
#
# - Create a test set with the same characteristics as the train and validation sets (signal shape and noise level) and test your selected model;
# - Change the input signal;
# - Change the noise intensity;
# - Use more data for training your model;
# - ...
| JNotebooks/overfitting_regularization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Preparing German Wikipedia to train a fast.ai (ULMFiT) model for German
# (should work with most other languages, too)
#
# *<NAME> <<EMAIL>>*
#
# The core idea of [Howard and Ruder's ULMFiT paper](https://arxiv.org/abs/1801.06146), see also https://nlp.fast.ai/, is to pretrain a language model on some corpus.
# Naturally, we also want such a thing for German. And happily I just launched [MathInf](https://mathinf.eu/), a great mathematical modelling, machine learning and actuarial consulting company, that allows me to do this type of research and make it public.
#
# [I have very raw info (and hope to add more description soon) on my blog](https://lernapparat.de/german-lm/). I'm making this available early at public request and hope it is useful to you to build great things, it is not as clean or well-commented I would love it to be, yet.
# I would love to hear from you if you make good use of it!
#
# So we take a wikipedia dump (`de_wikipedia_extracted dewiki-latest-pages-articles.xml.bz2` downloaded from [dumps.wikipedia.org](https://dumps.wikimedia.org/dewiki/latest/) and prepocessed by `wikiextractor/WikiExtractor.py -s --json -o de_wikipedia_extracted dewiki-latest-pages-articles.xml.bz2`) and make token files out of them.
#
# Note that the German Wikipedia contains more tokens (i.e. words) than recommended 100M to train the language model.
# I don't cut off much here, but only do this later when loading the tokens to start the training. That is a bit wasteful and follows a "keep as much data as long as you can" approach.
#
# Credit for all the good things in the Notebook likely belong to <NAME> ([see his notebook](https://github.com/sgugger/Deep-Learning/blob/master/Building%20a%20French%20LM.ipynb)) and <NAME> [see the original imdb notebook from his great course](https://github.com/fastai/fastai/blob/master/courses/dl2/imdb.ipynb), whose work I built on, all errors are my own.
#
#
# Enough talk, here is the data preparation.
# %matplotlib inline
# %reload_ext autoreload
# %autoreload 2
from fastai.text import *
import html
from matplotlib import pyplot
import numpy
import time
# +
BOS = 'xbos' # beginning-of-sentence tag
FLD = 'xfld' # data field tag
LANG='de'
datasetpath = Path('/home/datasets/nlp/wiki/')
# I ran this: wikiextractor/WikiExtractor.py -s --json -o de_wikipedia_extracted dewiki-latest-pages-articles.xml.bz2
work_path = Path('~/data/nlp/german_lm/data/de_wiki/tmp/').expanduser()
work_path.mkdir(exist_ok=True)
# -
# # Standarize format
#
# You can skip this entire section if you like the results. In this case continue at *Tokenize*.
LANG_FILENAMES = [str(f) for f in datasetpath.rglob("de_wikipedia_extracted/*/*")]
len(LANG_FILENAMES), LANG_FILENAMES[:5]
# +
LANG_TEXT = []
for fn in tqdm(LANG_FILENAMES):
for line in open(fn, encoding='utf8'):
LANG_TEXT.append(json.loads(line))
LANG_TEXT = pd.DataFrame(LANG_TEXT)
# -
LANG_TEXT.head()
# +
# Getting rid of the title name in the text field
def split_title_from_text(text):
words = text.split("\n\n", 1)
if len(words) == 2:
return words[1]
else:
return words[0]
LANG_TEXT['text'] = LANG_TEXT['text'].apply(lambda x: split_title_from_text(x))
# -
LANG_TEXT.head()
# Determine article lengths and only keep at most the largest million and only those with at least 2000 characters
LANG_TEXT['label'] = 0 # dummy
LANG_TEXT['length'] = LANG_TEXT['text'].str.len()
MAX_ARTICLES = 1_000_000
# keep at most 1 million articles and only those of more than 2000 characters
MIN_LENGTH_CHARS = max(2000, int(numpy.percentile(LANG_TEXT['length'], 100-min(100*MAX_ARTICLES/len(LANG_TEXT), 100))))
LANG_TEXT = LANG_TEXT[LANG_TEXT['length'] >= MIN_LENGTH_CHARS] # Chars not words...
LANG_TEXT.to_csv(datasetpath/'wiki_de.csv', header=True, index=False) # I must say, I think the header is good! If in doubt, you should listen to Jeremy though.
LANG_TEXT = pd.read_csv(datasetpath/'wiki_de.csv')
percentages = range(0,110,10)
print ('Article length percentiles' , ', '.join(['{}%: {}'.format(p, int(q)) for p,q in zip(percentages, numpy.percentile(LANG_TEXT['length'], percentages))]))
print ('Number of articles', len(LANG_TEXT))
#LANG_TEXT = LANG_TEXT.sort_values(by=['length'], ascending=False)
LANG_TEXT.head()
# Splitting 10% for validation.
df_trn,df_val = sklearn.model_selection.train_test_split(LANG_TEXT.pipe(lambda x: x[['label', 'text']]), test_size=0.1)
df_trn.to_csv(work_path/'train.csv', header=False, index=False)
df_val.to_csv(work_path/'valid.csv', header=False, index=False)
# I'm always trying to produce notebooks that you can run through in one go, so here is my attempt at getting rid of old stuff.
del LANG_TEXT
import gc
gc.collect()
# # Tokenize
#
# Note: be sure to care for your memory. I had all my memory allocated (for having several wikipedia copies in memory) and was swapping massively with the multiprocessing tokenization. My fix was to restart the notebook after after I had finished the above.
chunksize = 4000
N_CPUS = num_cpus() # I like to use all cores here, needs a patch to fast ai
# +
re1 = re.compile(r' +')
def fixup(x):
x = x.replace('#39;', "'").replace('amp;', '&').replace('#146;', "'").replace(
'nbsp;', ' ').replace('#36;', '$').replace('\\n', "\n").replace('quot;', "'").replace(
'<br />', "\n").replace('\\"', '"').replace('<unk>','u_n').replace(' @.@ ','.').replace(
' @-@ ','-').replace('\\', ' \\ ')
return re1.sub(' ', html.unescape(x))
# -
df_trn = pd.read_csv(work_path/'train.csv', header=None, chunksize=chunksize)
df_val = pd.read_csv(work_path/'valid.csv', header=None, chunksize=chunksize)
# +
def get_texts(df, n_lbls=1):
labels = df.iloc[:,range(n_lbls)].values.astype(np.int64)
texts = f'\n{BOS} {FLD} 1 ' + df[n_lbls].astype(str)
for i in range(n_lbls+1, len(df.columns)): texts += f' {FLD} {i-n_lbls} ' + df[i].astype(str)
texts = texts.apply(fixup).values.astype(str)
#tok = Tokenizer.proc_all(texts, lang=LANG) # use this if you have memory trouble
tok = Tokenizer.proc_all_mp(partition(texts, (len(texts)+N_CPUS-1)//N_CPUS), lang=LANG, ncpus=N_CPUS)
return tok, list(labels)
def get_all(df, name, n_lbls=1):
time_start = time.time()
for i, r in enumerate(df):
print("\r", i, end=" ")
if i > 0:
print ('time per chunk {}s'.format(int((time.time() - time_start) / i)), end="")
tok_, labels_ = get_texts(r, n_lbls)
#save the partial tokens instead of regrouping them in one big array.
np.save(work_path/f'{name}_tok{i}.npy', tok_)
# -
get_all(df_trn,'trn',1)
get_all(df_val,'val',1)
# # Numericalize
# Get the Counter object from all the splitted files.
def count_them_all(names):
cnt = Counter()
for name in names:
for file in work_path.glob(f'{name}_tok*'):
tok = np.load(file)
cnt_tok = Counter(word for sent in tok for word in sent)
cnt += cnt_tok
return cnt
cnt = count_them_all(['trn'])
cnt.most_common(25)
max_vocab = 60000
min_freq = 5
itos = [o for o,c in cnt.most_common(max_vocab) if c > min_freq]
itos.insert(0,'_pad_')
itos.insert(0,'_unk_')
len(itos)
pickle.dump(itos, open(work_path/'itos.pkl', 'wb'))
stoi = collections.defaultdict(int,{s:i for (i,s) in enumerate(itos)})
# Numericalize each partial file.
def numericalize(name):
results = []
for file in tqdm(work_path.glob(f'{name}_tok*')):
tok = np.load(file)
results.append(np.array([[stoi[word] for word in sent] for sent in tok]))
return np.concatenate(results)
trn_ids = numericalize('trn')
np.save(work_path/'trn_ids.npy', trn_ids)
val_ids = numericalize('val')
np.save(work_path/'val_ids.npy', val_ids)
# So now you have gread dumps to use with the [training program I published on my blog](https://lernapparat.de/german-lm/).
#
# As always, I would be honored by your feedback at <<EMAIL>>. I read and appreciate every mail.
#
# *Thomas*
| german_lm/German_LM_prepare.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LAS Demo
#
# This is a demonstration of how `run_exp.py` works and the result of example config file.
#
# ## Notes :
#
# 1. The goal is simply to reconstructed LAS and see if it works on TIMIT, all paremeters are NOT fine tuned.
#
# 2. I run the experiment once with unassigned random seed and get the result.
#
# 3. Run with smaller batch may help stablize testing performance. (but slowing down training process)
#
# 4. The model trained on example config can be find in `checkpoint/`
#
#
#
# ## Package & Config setting
# +
import yaml
from util.timit_dataset import load_dataset,create_dataloader
from util.functions import log_parser,batch_iterator, collapse_phn
from model.las_model import Listener,Speller
import numpy as np
from torch.autograd import Variable
import torch
import time
import matplotlib.pyplot as plt
% matplotlib inline
# Load example config file for experiment
config_path = 'config/las_example_config.yaml'
conf = yaml.load(open(config_path,'r'))
# Parameters loading
num_epochs = conf['training_parameter']['num_epochs']
training_msg = 'epoch_{:2d}_step_{:3d}_TrLoss_{:.4f}_TrWER_{:.2f}'
epoch_end_msg = 'epoch_{:2d}_TrLoss_{:.4f}_TrWER_{:.2f}_ValLoss_{:.4f}_ValWER_{:.2f}_time_{:.2f}'
listener_model_path = conf['meta_variable']['checkpoint_dir']+conf['meta_variable']['experiment_name']+'.listener'
speller_model_path = conf['meta_variable']['checkpoint_dir']+conf['meta_variable']['experiment_name']+'.speller'
verbose_step = conf['training_parameter']['verbose_step']
tf_rate_upperbound = conf['training_parameter']['tf_rate_upperbound']
tf_rate_lowerbound = conf['training_parameter']['tf_rate_lowerbound']
# -
# ## Prepare Dataset
# Load preprocessed TIMIT Dataset
# X : Padding to shape [num of sample, max_timestep, feature_dim]
# Y : Squeeze repeated label and apply one-hot encoding (preserve 0 for <sos> and 1 for <eos>)
X_train, y_train, X_val, y_val, X_test, y_test = load_dataset(**conf['meta_variable'])
train_set = create_dataloader(X_train, y_train, **conf['model_parameter'], **conf['training_parameter'], shuffle=True)
valid_set = create_dataloader(X_val, y_val, **conf['model_parameter'], **conf['training_parameter'], shuffle=False)
test_set = create_dataloader(X_test, y_test, **conf['model_parameter'], **conf['training_parameter'], shuffle=False)
# ## Construct Model
listener = Listener(**conf['model_parameter'])
speller = Speller(**conf['model_parameter'])
optimizer = torch.optim.Adam([{'params':listener.parameters()}, {'params':speller.parameters()}],
lr=conf['training_parameter']['learning_rate'])
# ## Training Progress
# +
best_ler = 1.0
traing_log = open(conf['meta_variable']['training_log_dir']+conf['meta_variable']['experiment_name']+'.log','w')
for epoch in range(num_epochs):
epoch_head = time.time()
tr_loss = 0.0
tr_ler = []
val_loss = 0.0
val_ler = []
# Teacher forcing rate linearly decay
tf_rate = tf_rate_upperbound - (tf_rate_upperbound-tf_rate_lowerbound)*(epoch/num_epochs)
# Training
for batch_index,(batch_data,batch_label) in enumerate(train_set):
batch_loss, batch_ler = batch_iterator(batch_data, batch_label, listener, speller, optimizer,
tf_rate, is_training=True, **conf['model_parameter'])
tr_loss += batch_loss
tr_ler.extend(batch_ler)
if (batch_index+1) % verbose_step == 0:
print(training_msg.format(epoch+1,batch_index+1,tr_loss[0]/(batch_index+1),sum(tr_ler)/len(tr_ler)),end='\r',flush=True)
training_time = float(time.time()-epoch_head)
# Validation
for _,(batch_data,batch_label) in enumerate(valid_set):
batch_loss, batch_ler = batch_iterator(batch_data, batch_label, listener, speller, optimizer,
tf_rate, is_training=False, **conf['model_parameter'])
val_loss += batch_loss
val_ler.extend(batch_ler)
# Logger
print(epoch_end_msg.format(epoch+1,tr_loss[0]/(batch_index+1),sum(tr_ler)/len(tr_ler),
val_loss[0]/len(valid_set),sum(val_ler)/len(val_ler),training_time),flush=True)
print(epoch_end_msg.format(epoch+1,tr_loss[0]/(batch_index+1),sum(tr_ler)/len(tr_ler),
val_loss[0]/len(valid_set),sum(val_ler)/len(val_ler),training_time),flush=True,file=traing_log)
# Checkpoint
if best_ler >= sum(val_ler)/len(val_ler):
best_ler = sum(val_ler)/len(val_ler)
torch.save(listener, listener_model_path)
torch.save(speller, speller_model_path)
# -
# # Result Visualization
#
# ## Learning Curve & Performance
#
# +
fig, axs = plt.subplots(1,2,figsize=(16,5))
tr_loss, tt_loss, tr_ler, tt_ler = log_parser('log/las_example.log')
axs[0].plot(tr_loss,label='Training');axs[0].plot(tt_loss,label='Validation');
axs[0].legend();axs[0].set_xlabel('training epochs');axs[0].set_ylabel('Loss');
axs[0].set_title('Learning Curve of LAS',size=15);axs[0].grid(True);
axs[1].plot(tr_ler,label='Training');axs[1].plot(tt_ler,label='Validation');axs[1].grid(True);
axs[1].legend();axs[1].set_xlabel('training epochs');axs[1].set_ylabel('Phoneme Error Rate');
axs[1].set_ylim(0,1)
axs[1].set_title('Performance of LAS',size=15)
plt.savefig('log/result.jpg')
# -
# ## Attention Visualization & Recognition Result
# +
target = 0
listener = torch.load(listener_model_path)
speller = torch.load(speller_model_path)
for batch_index,(batch_data,batch_label) in enumerate(test_set):
test_sample = batch_data[target:target+1]
test_label = batch_label[target]
break
feature = listener(Variable(test_sample.float()).cuda())
pred_seq, attention_score = speller(feature)
encoded_feature_length = int(sum(sum(list(np.sum(test_sample.numpy(),axis=-1)!=0)))/8)
pred = []
att_map = []
truth = []
for pred_phn,att_score,true_phn in zip(pred_seq, attention_score,test_label):
ground_truth = np.argmax(true_phn.numpy())
pred.append(np.argmax(pred_phn.cpu().data.numpy()))
att_map.append(att_score.cpu().data.numpy()[0,:encoded_feature_length])
truth.append(ground_truth)
if ground_truth == 1:
break
for idx,p in enumerate(pred):
if p == 1:
break
att_map = np.array(att_map)
pred = collapse_phn(pred[:idx],return_phn=True,drop_q=False)
pred.extend(['<eos>' for i in range(len(truth)-len(pred))])
truth = collapse_phn(truth[:-1],return_phn=True,drop_q=False)
truth.append('<eos>')
# +
fig,ax = plt.subplots(1,figsize=(15,13))
ax2 = ax.twinx()
plt.title('Attention Score During Decoding',size=30)
ax.set_xlabel('(Encoded) Input Sequence',size=20)
ax2.set_xlabel('(Encoded) Input Sequence',size=20)
ax.imshow(att_map, cmap='hot')
ax2.imshow(att_map, cmap='hot')
ax.set_yticks(range(len(truth)))
ax.set_yticklabels(pred,size=15)
ax.set_ylabel('Decoding Step',size=20)
ax2.set_yticks(range(len(truth)))
ax2.set_yticklabels(truth,size=15)
ax2.set_ylabel('Ground Truth',size=20)
plt.savefig('log/attention.jpg')
# -
# # Testing Performance
# +
from util.functions import LetterErrorRate
test_ler = []
for batch_index,(batch_data,batch_label) in enumerate(test_set):
feature = listener(Variable(batch_data.float()).cuda())
pred_seq, _ = speller(feature)
batch_ler = LetterErrorRate(np.argmax(np.array([p.data.cpu().numpy() for p in pred_seq]).transpose(1,0,2),axis=-1),
np.argmax(batch_label.numpy(),axis=-1))
test_ler.extend(batch_ler)
print('Testing Phoneme Error Rate',sum(test_ler)/len(test_ler))
# -
| las_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
with open('C:\\Users\\void\\conda projects\\diary_parser\\diaries\\all_links') as f:
data = f.readlines()
data = [x[:-1] for x in data]
def extract_username(link):
return link.replace('https://www.diary.ru/' ,'').split('/')[0][1:]
import os
links_by_user = {}
for link in data:
user = extract_username(link)
if not user in links_by_user:
links_by_user[user] = []
links_by_user[user].append(link)
for user in links_by_user:
path = 'C:\\Users\\void\\conda projects\\diary_parser\\diaries\\' + user
os.makedirs(path)
with open(path + '\\links', 'w') as f:
for link in links_by_user[user]:
f.write(link + '\n')
| Group links by user.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Scikit-learn DBSCAN OD Clustering
#
# <img align="right" src="https://anitagraser.github.io/movingpandas/assets/img/movingpandas.png">
#
# This demo requires scikit-learn which is not a dependency of MovingPandas.
# %matplotlib inline
# +
import urllib
import os
import numpy as np
import pandas as pd
from geopandas import GeoDataFrame, read_file
from shapely.geometry import Point, LineString, Polygon, MultiPoint
from datetime import datetime, timedelta
import matplotlib.pyplot as plt
from sklearn.cluster import DBSCAN
from geopy.distance import great_circle
import sys
sys.path.append("..")
import movingpandas as mpd
import warnings
warnings.simplefilter("ignore")
# -
# ## Ship movements (AIS data)
df = read_file('../data/ais.gpkg')
df['t'] = pd.to_datetime(df['Timestamp'], format='%d/%m/%Y %H:%M:%S')
df = df.set_index('t')
df = df[df.SOG>0]
MIN_LENGTH = 100 # meters
TRIP_ID = 'MMSI'
traj_collection = mpd.TrajectoryCollection(df, TRIP_ID, min_length=MIN_LENGTH)
print("Finished creating {} trajectories".format(len(traj_collection)))
trips = mpd.ObservationGapSplitter(traj_collection).split(gap=timedelta(minutes=5))
print("Extracted {} individual trips from {} continuous vessel tracks".format(len(trips), len(traj_collection)))
KMS_PER_RADIAN = 6371.0088
EPSILON = 0.1 / KMS_PER_RADIAN
trips.get_start_locations()
# +
def make_od_line(row, od_clusters):
return LineString([od_clusters.loc[row['od'][0]].geometry, od_clusters.loc[row['od'][-1]].geometry])
def get_centermost_point(cluster):
centroid = (MultiPoint(cluster).centroid.x, MultiPoint(cluster).centroid.y)
centermost_point = min(cluster, key=lambda point: great_circle(point, centroid).m)
return Point(tuple(centermost_point)[1], tuple(centermost_point)[0])
def extract_od_gdf(trips):
origins = trips.get_start_locations()
origins['type'] = '0'
origins['traj_id'] = [trip.id for trip in trips]
destinations = trips.get_end_locations()
destinations['type'] = '1'
destinations['traj_id'] = [trip.id for trip in trips]
od = origins.append(destinations)
od['lat'] = od.geometry.y
od['lon'] = od.geometry.x
return od
def dbscan_cluster_ods(od_gdf, eps):
matrix = od_gdf[['lat', 'lon']].to_numpy()
db = DBSCAN(eps=eps, min_samples=1, algorithm='ball_tree', metric='haversine').fit(np.radians(matrix))
cluster_labels = db.labels_
num_clusters = len(set(cluster_labels))
clusters = pd.Series([matrix[cluster_labels == n] for n in range(num_clusters)])
return cluster_labels, clusters
def extract_od_clusters(od_gdf, eps):
cluster_labels, clusters = dbscan_cluster_ods(od_gdf, eps)
od_gdf['cluster'] = cluster_labels
od_by_cluster = pd.DataFrame(od_gdf).groupby(['cluster'])
clustered = od_by_cluster['ShipType'].unique().to_frame(name='types')
clustered['n'] = od_by_cluster.size()
clustered['symbol_size'] = clustered['n']*10 # for visualization purposes
clustered['sog'] = od_by_cluster['SOG'].mean()
clustered['geometry'] = clusters.map(get_centermost_point)
clustered = clustered[clustered['n']>0].sort_values(by='n', ascending=False)
return clustered
def extract_od_matrix(trips, eps, directed=True):
od_gdf = extract_od_gdf(trips)
matrix_nodes = extract_od_clusters(od_gdf, eps)
od_by_traj_id = pd.DataFrame(od_gdf).sort_values(['type']).groupby(['traj_id']) # Groupby preserves the order of rows within each group.
od_by_traj_id = od_by_traj_id['cluster'].unique().to_frame(name='clusters') # unique() preserves input order according to https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.unique.html
if directed:
od_matrix = od_by_traj_id.groupby(od_by_traj_id['clusters'].apply(tuple)).count().rename({'clusters':'n'}, axis=1)
else:
od_matrix = od_by_traj_id.groupby(od_by_traj_id['clusters'].apply(sorted).apply(tuple)).count().rename({'clusters':'n'}, axis=1)
od_matrix['od'] = od_matrix.index
od_matrix['geometry'] = od_matrix.apply(lambda x: make_od_line(row=x, od_clusters=matrix_nodes), axis=1 )
return od_matrix, matrix_nodes
od_matrix, matrix_nodes = extract_od_matrix(trips, EPSILON*2, directed=True)
# -
np.max(od_matrix.n)
# +
from holoviews import dim
( GeoDataFrame(od_matrix).hvplot(title='OD flows', geo=True, tiles='OSM', line_width=dim('n'), alpha=0.5, frame_height=600, frame_width=600) *
GeoDataFrame(matrix_nodes).hvplot(c='sog', size='symbol_size', hover_cols=['cluster', 'n'], geo=True, cmap='RdYlGn')
)
# -
# ## Bird migration data
df = read_file('../data/gulls.gpkg')
df['t'] = pd.to_datetime(df['timestamp'])
df = df.set_index('t')
traj_collection = mpd.TrajectoryCollection(df, 'individual-local-identifier', min_length=MIN_LENGTH)
print("Finished creating {} trajectories".format(len(traj_collection)))
trips = mpd.TemporalSplitter(traj_collection).split(mode='month')
print("Extracted {} individual trips from {} continuous tracks".format(len(trips), len(traj_collection)))
# +
EPSILON = 100 / KMS_PER_RADIAN
def extract_od_gdf(trips):
origins = trips.get_start_locations()
origins['type'] = '0'
origins['traj_id'] = [trip.id for trip in trips]
destinations = trips.get_end_locations()
destinations['type'] = '1'
destinations['traj_id'] = [trip.id for trip in trips]
od = origins.append(destinations)
od['lat'] = od.geometry.y
od['lon'] = od.geometry.x
return od
def extract_od_clusters(od_gdf, eps):
cluster_labels, clusters = dbscan_cluster_ods(od_gdf, eps)
od_gdf['cluster'] = cluster_labels
od_by_cluster = pd.DataFrame(od_gdf).groupby(['cluster'])
clustered = od_by_cluster.size().to_frame(name='n')
clustered['geometry'] = clusters.map(get_centermost_point)
clustered = clustered[clustered['n']>0].sort_values(by='n', ascending=False)
return clustered
od_matrix, matrix_nodes = extract_od_matrix(trips, EPSILON, directed=False)
# -
( GeoDataFrame(od_matrix).hvplot(title='OD flows', geo=True, tiles='OSM', hover_cols=['n'], line_width=dim('n')*0.05, alpha=0.5, frame_height=600, frame_width=600) *
GeoDataFrame(matrix_nodes).hvplot(c='n', size=dim('n')*0.1, hover_cols=['cluster', 'n'], geo=True, cmap='RdYlGn')
)
# ### Comparing OD flows and TrajectoryCollectionAggregator
# +
aggregator = mpd.TrajectoryCollectionAggregator(trips, max_distance=1000000, min_distance=100000, min_stop_duration=timedelta(minutes=5))
flows = aggregator.get_flows_gdf()
clusters = aggregator.get_clusters_gdf()
# -
( flows.hvplot(title='Generalized aggregated trajectories', geo=True, hover_cols=['weight'], line_width='weight', alpha=0.5, color='#1f77b3', tiles='OSM', frame_height=600, frame_width=400) *
clusters.hvplot(geo=True, color='red', size='n')
+
GeoDataFrame(od_matrix).hvplot(title='OD flows', geo=True, tiles='OSM', hover_cols=['n'], line_width=dim('n')*0.05, alpha=0.5, frame_height=600, frame_width=400) *
GeoDataFrame(matrix_nodes).hvplot(c='n', size=dim('n')*0.1, hover_cols=['cluster', 'n'], geo=True, cmap='RdYlGn')
)
| 3-tech-demos/scikit-learn-od-clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Mixed Effects Models
# +
# %matplotlib inline
import numpy as np
import statsmodels.api as sm
import statsmodels.formula.api as smf
# -
# %load_ext rpy2.ipython
# %R library(lme4)
# Comparing R lmer to Statsmodels MixedLM
# =======================================
#
# The Statsmodels imputation of linear mixed models (MixedLM) closely follows the approach outlined in Lindstrom and Bates (JASA 1988). This is also the approach followed in the R package LME4. Other packages such as Stata, SAS, etc. should also be consistent with this approach, as the basic techniques in this area are mostly mature.
#
# Here we show how linear mixed models can be fit using the MixedLM procedure in Statsmodels. Results from R (LME4) are included for comparison.
#
# Here are our import statements:
# ## Growth curves of pigs
#
# These are longitudinal data from a factorial experiment. The outcome variable is the weight of each pig, and the only predictor variable we will use here is "time". First we fit a model that expresses the mean weight as a linear function of time, with a random intercept for each pig. The model is specified using formulas. Since the random effects structure is not specified, the default random effects structure (a random intercept for each group) is automatically used.
data = sm.datasets.get_rdataset('dietox', 'geepack').data
md = smf.mixedlm("Weight ~ Time", data, groups=data["Pig"])
mdf = md.fit()
print(mdf.summary())
# Here is the same model fit in R using LMER:
# + language="R"
# data(dietox, package='geepack')
# -
# %R print(summary(lmer('Weight ~ Time + (1|Pig)', data=dietox)))
# Note that in the Statsmodels summary of results, the fixed effects and random effects parameter estimates are shown in a single table. The random effect for animal is labeled "Intercept RE" in the Statmodels output above. In the LME4 output, this effect is the pig intercept under the random effects section.
#
# There has been a lot of debate about whether the standard errors for random effect variance and covariance parameters are useful. In LME4, these standard errors are not displayed, because the authors of the package believe they are not very informative. While there is good reason to question their utility, we elected to include the standard errors in the summary table, but do not show the corresponding Wald confidence intervals.
#
# Next we fit a model with two random effects for each animal: a random intercept, and a random slope (with respect to time). This means that each pig may have a different baseline weight, as well as growing at a different rate. The formula specifies that "Time" is a covariate with a random coefficient. By default, formulas always include an intercept (which could be suppressed here using "0 + Time" as the formula).
md = smf.mixedlm("Weight ~ Time", data, groups=data["Pig"], re_formula="~Time")
mdf = md.fit()
print(mdf.summary())
# Here is the same model fit using LMER in R:
# %R print(summary(lmer("Weight ~ Time + (1 + Time | Pig)", data=dietox)))
# The random intercept and random slope are only weakly correlated $(0.294 / \sqrt{19.493 * 0.416} \approx 0.1)$. So next we fit a model in which the two random effects are constrained to be uncorrelated:
.294 / (19.493 * .416)**.5
# +
md = smf.mixedlm("Weight ~ Time", data, groups=data["Pig"],
re_formula="~Time")
free = sm.regression.mixed_linear_model.MixedLMParams.from_components(np.ones(2),
np.eye(2))
mdf = md.fit(free=free)
print(mdf.summary())
# -
# The likelihood drops by 0.3 when we fix the correlation parameter to 0. Comparing 2 x 0.3 = 0.6 to the chi^2 1 df reference distribution suggests that the data are very consistent with a model in which this parameter is equal to 0.
#
# Here is the same model fit using LMER in R (note that here R is reporting the REML criterion instead of the likelihood, where the REML criterion is twice the log likeihood):
# %R print(summary(lmer("Weight ~ Time + (1 | Pig) + (0 + Time | Pig)", data=dietox)))
# ## Sitka growth data
#
# This is one of the example data sets provided in the LMER R library. The outcome variable is the size of the tree, and the covariate used here is a time value. The data are grouped by tree.
data = sm.datasets.get_rdataset("Sitka", "MASS").data
endog = data["size"]
data["Intercept"] = 1
exog = data[["Intercept", "Time"]]
# Here is the statsmodels LME fit for a basic model with a random intercept. We are passing the endog and exog data directly to the LME init function as arrays. Also note that endog_re is specified explicitly in argument 4 as a random intercept (although this would also be the default if it were not specified).
md = sm.MixedLM(endog, exog, groups=data["tree"], exog_re=exog["Intercept"])
mdf = md.fit()
print(mdf.summary())
# Here is the same model fit in R using LMER:
# + language="R"
# data(Sitka, package="MASS")
# print(summary(lmer("size ~ Time + (1 | tree)", data=Sitka)))
# -
# We can now try to add a random slope. We start with R this time. From the code and output below we see that the REML estimate of the variance of the random slope is nearly zero.
# %R print(summary(lmer("size ~ Time + (1 + Time | tree)", data=Sitka)))
# If we run this in statsmodels LME with defaults, we see that the variance estimate is indeed very small, which leads to a warning about the solution being on the boundary of the parameter space. The regression slopes agree very well with R, but the likelihood value is much higher than that returned by R.
exog_re = exog.copy()
md = sm.MixedLM(endog, exog, data["tree"], exog_re)
mdf = md.fit()
print(mdf.summary())
# We can further explore the random effects struture by constructing plots of the profile likelihoods. We start with the random intercept, generating a plot of the profile likelihood from 0.1 units below to 0.1 units above the MLE. Since each optimization inside the profile likelihood generates a warning (due to the random slope variance being close to zero), we turn off the warnings here.
# +
import warnings
with warnings.catch_warnings():
warnings.filterwarnings("ignore")
likev = mdf.profile_re(0, 're', dist_low=0.1, dist_high=0.1)
# -
# Here is a plot of the profile likelihood function. We multiply the log-likelihood difference by 2 to obtain the usual $\chi^2$ reference distribution with 1 degree of freedom.
import matplotlib.pyplot as plt
plt.figure(figsize=(10,8))
plt.plot(likev[:,0], 2*likev[:,1])
plt.xlabel("Variance of random slope", size=17)
plt.ylabel("-2 times profile log likelihood", size=17)
# Here is a plot of the profile likelihood function. The profile likelihood plot shows that the MLE of the random slope variance parameter is a very small positive number, and that there is low uncertainty in this estimate.
# +
re = mdf.cov_re.iloc[1, 1]
likev = mdf.profile_re(1, 're', dist_low=.5*re, dist_high=0.8*re)
plt.figure(figsize=(10, 8))
plt.plot(likev[:,0], 2*likev[:,1])
plt.xlabel("Variance of random slope", size=17)
plt.ylabel("-2 times profile log likelihood", size=17)
# -
| examples/notebooks/mixed_lm_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from numpy import linalg as LA
# -
def square_matrix_ridge_regression(X,Y,Astar,lmda,beeta):
t=1+X.shape[1]
D=lmda*np.array([[1,0,0,0],[0,1,0,0],[0,0,1,0],[0,0,0,1]])
C=lmda*Astar
for s in range(1,t):
xs=X[:,s-1]
ys=Y[:,s-1]
C=C+(1/beeta)*np.outer(ys,xs)
D=D+(1/beeta)*np.outer(xs,xs)
return C.dot(np.linalg.inv(D))
A=np.random.rand(2,4)
Astar=A+np.random.normal(0,np.sqrt(0.09),A.shape)
X=np.zeros((4,1000000))
Y=np.zeros((2,1000000))
for t in range(1000000):
X[:,t]=np.random.rand(4)
Y[:,t]=A.dot(X[:,t])+np.random.normal(0,np.sqrt(0.0001),2)
A_regression=square_matrix_ridge_regression(X,Y,Astar,100000,100000)
LA.norm((A-A_regression),'fro')
print(A)
print(A_regression)
print(A-A_regression)
print(Astar)
| Ordinary LQR/Matrix Regression Checking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="lbH1bRBGWLmC"
# # ALPR using YOLO-Net as LP detector and Modified YOLO as OCR
# + [markdown] colab_type="text" id="3koVM2Y-WN1f"
# ## Introduction
# + [markdown] colab_type="text" id="su7SkjYwe_3i"
# In this notebook, we will implement Automatic License Plate Recognition (ALPR) system composed by
# 1. Vehicle Detection using **YOLOv2** network trained on PASCAL-VOC dataset
# 1. License Plate (LP) Detection using **YOLO-Net** trained on [OpenALPR benchmark](https://platerecognizer.com/number-plate-datasets/) dataset
# 1. Optical Character Recognition (OCR) using **Modified YOLO** network, with the same architecture presented in **Real-Time Brazilian License Plate Detection and Recognition Using Deep Convolutional Neural Networks** by <NAME> and <NAME> [[paper](http://sibgrapi.sid.inpe.br/archive.cgi/sid.inpe.br/sibgrapi/2017/08.18.12.21)][[webpage](http://www.inf.ufrgs.br/~smsilva/real-time-brazilian-alpr/)]
#
# The ALPR implementation process involve,
# 1. Vehicle Detection
# ```
# 1.1 Download weights and config file of YOLOv2 network trained on PASCAL-VOC dataset
# 1.2 Utility functions
# 1.3 Detect vehicles
# ```
# 2. License Plate Detection
# ```
# 2.1 Download weights and config file of pretrained YOLO-Net
# 2.2 Utility functions
# 2.3 Detect license plates
# ```
# 3. Optical Character Recognition
# ```
# 3.1 Download weights and config file of pretrained Modified YOLO
# 3.2 Utility functions
# 3.3 Recognize characters
# ```
# 4. Inference
# ```
# 4.1 Download the test image
# 4.2 Utility functions
# 4.3 Infer on the test image
# 4.4 Display inference
# 4.5 Observations
# ```
# + [markdown] colab_type="text" id="2wZwOBS68q-t"
# ## 1. Vehicle Detection
# + [markdown] colab_type="text" id="OOO4RrWMbvHB"
# Vehicle detection is the first step in ALPR system. Vehicles are one of the underlying objects present in many classical detection and recognition datasets, such as PASCAL-VOC, ImageNet, and COCO.
#
# We will use pretrained YOLOv2 network trained on PASCAL-VOC dataset to perform vehicle detection.
# We will use weights and config file of YOLOv2 from [here](http://www.inf.ufrgs.br/~smsilva/alpr-unconstrained/data/vehicle-detector/), which is same as used by <NAME> an author of **License Plate Detection and Recognition in Unconstrained Scenarios**. The model was trained for 20 different object classes. The full list of class names can be found [here](http://www.inf.ufrgs.br/~smsilva/alpr-unconstrained/data/vehicle-detector/voc.names).
#
# We will not perform any change or refinement to YOLOv2, just we will use the network as a black box, merging the outputs related to vehicles (i.e. cars and buses), and ignoring the other classes.
#
#
#
#
# + [markdown] colab_type="text" id="NMIgaDlYLYoC"
# ### 1.1 Download weights and config file of YOLOv2 network trained on PASCAL-VOC dataset
# + colab={} colab_type="code" id="MO0hXprLKQ45"
# !wget -c -N www.inf.ufrgs.br/~smsilva/alpr-unconstrained/data/vehicle-detector/yolo-voc.cfg -P vehicle-detector/
# !wget -c -N www.inf.ufrgs.br/~smsilva/alpr-unconstrained/data/vehicle-detector/voc.data -P vehicle-detector/
# !wget -c -N www.inf.ufrgs.br/~smsilva/alpr-unconstrained/data/vehicle-detector/yolo-voc.weights -P vehicle-detector/
# !wget -c -N www.inf.ufrgs.br/~smsilva/alpr-unconstrained/data/vehicle-detector/voc.names -P vehicle-detector/
# + [markdown] colab_type="text" id="WgMdsA19G6Y4"
# ### 1.2 Utility functions
# + [markdown] colab_type="text" id="br274ZMdqgVu"
# Here, we define few utility functions like,
# - `getOutputsNames`: Get the names of the output layers for given input neural network.
# - `postprocess`: to get rid of detected bounding box with low confidence
# - `drawPred:` to draw the predicted bounding box
# - `crop_region:` to crop out specified region from given input image
#
# We also define `Label` a bounding box class. All detected bounding boxs are stored as an object of this class.
# + cellView="form" colab={} colab_type="code" id="LDamrM56L7Of"
#@title
# Get the names of the output layers
def getOutputsNames(net):
""" Get the names of the output layers.
Generally in a sequential CNN network there will be
only one output layer at the end. In the YOLOv3
architecture, there are multiple output layers giving
out predictions. This function gives the names of the
output layers. An output layer is not connected to
any next layer.
Args
net : neural network
"""
# Get the names of all the layers in the network
layersNames = net.getLayerNames()
# Get the names of the output layers, i.e. the layers with unconnected outputs
return [layersNames[i[0] - 1] for i in net.getUnconnectedOutLayers()]
import cv2 as cv
# Remove the bounding boxes with low confidence using non-maxima suppression
def postprocess(frame, outs, confThreshold, nmsThreshold=0.4):
frameHeight = frame.shape[0]
frameWidth = frame.shape[1]
classIds = []
confidences = []
boxes = []
# Scan through all the bounding boxes output from the network and keep only the
# ones with high confidence scores. Assign the box's class label as the class with the highest score.
classIds = []
confidences = []
boxes = []
predictions = []
for out in outs:
for detection in out:
scores = detection[5:]
classId = np.argmax(scores)
confidence = scores[classId]
if confidence > confThreshold:
center_x = int(detection[0] * frameWidth)
center_y = int(detection[1] * frameHeight)
width = int(detection[2] * frameWidth)
height = int(detection[3] * frameHeight)
left = int(center_x - width / 2)
top = int(center_y - height / 2)
classIds.append(classId)
confidences.append(float(confidence))
boxes.append([left, top, width, height])
# Perform non maximum suppression to eliminate redundant overlapping boxes with
# lower confidences.
if nmsThreshold:
indices = cv.dnn.NMSBoxes(boxes, confidences, confThreshold, nmsThreshold)
else:
indices = [[x] for x in range(len(boxes))]
for i in indices:
i = i[0]
box = boxes[i]
left = box[0]
top = box[1]
width = box[2]
height = box[3]
predictions.append([classIds[i], confidences[i], [left, top, left + width, top + height]])
return predictions
import cv2 as cv
# Draw the predicted bounding box
def drawPred(frame, pred):
classId = pred[0]
conf = pred[1]
box = pred[2]
left, top, right, bottom = box[0], box[1], box[2], box[3]
# draw bounding box
cv.rectangle(frame, (left, top), (right, bottom), (0, 255, 0), 3)
import numpy as np
class Label:
def __init__(self,cl=-1,tl=np.array([0.,0.]),br=np.array([0.,0.]),prob=None):
self.__tl = tl
self.__br = br
self.__cl = cl
self.__prob = prob
def __str__(self):
return 'Class: %d, top_left(x:%f,y:%f), bottom_right(x:%f,y:%f)' % (self.__cl, self.__tl[0], self.__tl[1], self.__br[0], self.__br[1])
def copy(self):
return Label(self.__cl,self.__tl,self.__br)
def wh(self): return self.__br-self.__tl
def cc(self): return self.__tl + self.wh()/2
def tl(self): return self.__tl
def br(self): return self.__br
def tr(self): return np.array([self.__br[0],self.__tl[1]])
def bl(self): return np.array([self.__tl[0],self.__br[1]])
def cl(self): return self.__cl
def area(self): return np.prod(self.wh())
def prob(self): return self.__prob
def set_class(self,cl):
self.__cl = cl
def set_tl(self,tl):
self.__tl = tl
def set_br(self,br):
self.__br = br
def set_wh(self,wh):
cc = self.cc()
self.__tl = cc - .5*wh
self.__br = cc + .5*wh
def set_prob(self,prob):
self.__prob = prob
def crop_region(I,label,bg=0.5):
wh = np.array(I.shape[1::-1])
ch = I.shape[2] if len(I.shape) == 3 else 1
tl = np.floor(label.tl()*wh).astype(int)
br = np.ceil (label.br()*wh).astype(int)
outwh = br-tl
if np.prod(outwh) == 0.:
return None
outsize = (outwh[1],outwh[0],ch) if ch > 1 else (outwh[1],outwh[0])
if (np.array(outsize) < 0).any():
pause()
Iout = np.zeros(outsize,dtype=I.dtype) + bg
offset = np.minimum(tl,0)*(-1)
tl = np.maximum(tl,0)
br = np.minimum(br,wh)
wh = br - tl
Iout[offset[1]:(offset[1] + wh[1]),offset[0]:(offset[0] + wh[0])] = I[tl[1]:br[1],tl[0]:br[0]]
return Iout
# + [markdown] colab_type="text" id="BBgTXhEfG894"
# ### 1.3 Detect vehicles
# + [markdown] colab_type="text" id="zgYtMlKPgov4"
# Let's define the `vehicle_detection` function which takes an image as input and return `Icars` list of cropped images of vehicles as well as `Lcars` list of bouding box around vehicles.
#
# We use `postprocess` utility function to get rid of detected bounding box with low confidence. The `postprocess` utility function internally uses `cv.dnn.NMSBoxes` which perform non maximum suppression to eliminate redundant overlapping boxes with lower confidences. We keep only those bounding boxs whose corresponding `classId` is either `car` (class number 6) or `bus` (class number 7), since this two `classId` are related to vehicles.
#
# We will use `vehicle_detection` function as first step in our ALPR system implementation.
# + cellView="both" colab={} colab_type="code" id="IQLn5oe7bT3a"
# Import necessary modules
import cv2 as cv
import numpy as np
# Initialize the parameters
vehicle_threshold = .5
vehicle_weights = 'vehicle-detector/yolo-voc.weights'
vehicle_netcfg = 'vehicle-detector/yolo-voc.cfg'
# Load the model
vehicle_net = cv.dnn.readNetFromDarknet(vehicle_netcfg, vehicle_weights)
vehicle_net.setPreferableBackend(cv.dnn.DNN_BACKEND_OPENCV)
vehicle_net.setPreferableTarget(cv.dnn.DNN_TARGET_CPU)
def vehicle_detection(frame):
# Create a 4D blob from a frame.
blob = cv.dnn.blobFromImage(frame, 1/255, (416, 416), [0,0,0], 1, crop=False)
# Sets the input to the network
vehicle_net.setInput(blob)
# Runs the forward pass to get output of the output layers
outs = vehicle_net.forward(getOutputsNames(vehicle_net))
# Remove the bounding boxes with low confidence
R = postprocess(frame, outs, vehicle_threshold)
Icars = []
Lcars = []
if len(R):
WH = np.array(frame.shape[1::-1], dtype=float)
for i, r in enumerate(R):
# if classId in ['car', 'bus'] and confidence > vehicle_threshold
if r[0] in [6, 7] and r[1] > vehicle_threshold:
box = r[2]
x1,y1,x2,y2 = (np.array(r[2])/np.concatenate((WH,WH))).tolist()
tl = np.array([x1, y1])
br = np.array([x2, y2])
label = Label(0,tl,br)
Lcars.append(label)
Icar = crop_region(frame,label)
Icars.append(Icar.astype(np.uint8))
return Icars, Lcars
# + [markdown] colab_type="text" id="xKDLlZcdGr0n"
# ## 2. License Plate Detection
# + [markdown] colab_type="text" id="Ok7qp6HysjAb"
# License Plate Detection is the second step in ALPR system. For each detected vehicle in the first step, we apply this second step to detect region of license plate in detected vehicle region.
#
# We will use YOLO-Net - a YOLOv3 Network trained in previous notebook, for LP detection.
# + [markdown] colab_type="text" id="_Pxrp8X2_jBC"
# ### 2.1 Download weights and config file of pretrained YOLO-Net
# + colab={} colab_type="code" id="qeWSyPjSX-gF"
# !wget -c -N https://www.dropbox.com/s/vw9omi6tjntp6vr/yolov3-LP-train_best.weights -P lp-detector/
# !wget -c -N https://www.dropbox.com/sh/5y72h8ul8654y9i/AAAFwOwOl7bsQ4BmuxraKBRta?dl=0 -O lp-detector/yolov3_LP.zip
# %cd lp-detector
# !unzip -q yolov3_LP.zip
# %cd ..
# + [markdown] colab_type="text" id="V4ul6eGnmWYt"
# ### 2.2 Detect license plates
# + [markdown] colab_type="text" id="sVtXkqI_t8AQ"
# Let's define a `lp_detection` function which takes vehicle image as input and return `Llps` list of bounding boxs around detected license plates and `Ilps` list of croppsed images of detected license plates.
#
# Again, we use `postprocess` utility function to remove bouding box with low threshold.
# + colab={} colab_type="code" id="Y0Taw1zSmY4l"
# Import necessary modules
import cv2 as cv
import numpy as np
import time
# Initialize the parameters
lp_threshold = .6
yolo_lp_confi_path = 'lp-detector/yolov3-LP-test.cfg'
yolo_lp_weights_path = 'lp-detector/yolov3-LP-train_best.weights'
# load the network
lp_net = cv.dnn.readNetFromDarknet(yolo_lp_confi_path, yolo_lp_weights_path)
lp_net.setPreferableBackend(cv.dnn.DNN_BACKEND_OPENCV)
lp_net.setPreferableTarget(cv.dnn.DNN_TARGET_CPU)
def lp_detection(vehicle_img):
# Create a 4D blob from a frame.
blob = cv.dnn.blobFromImage(vehicle_img, 1/255, (416, 416), [0,0,0], 1, crop=False)
# Sets the input to the network
lp_net.setInput(blob)
# Runs the forward pass to get output of the output layers
start = time.time()
outs = lp_net.forward(getOutputsNames(lp_net))
elapsed = time.time() - start
# Remove the bounding boxes with low confidence
R = postprocess(vehicle_img, outs, lp_threshold)
Llps = []
Ilps = []
if len(R):
WH = np.array(vehicle_img.shape[1::-1], dtype=float)
for i, r in enumerate(R):
if r[1] > lp_threshold:
box = r[2]
x1,y1,x2,y2 = (np.array(r[2])/np.concatenate((WH,WH))).tolist()
tl = np.array([x1, y1])
br = np.array([x2, y2])
label = Label(0,tl,br)
Llps.append(label)
Ilp = crop_region(vehicle_img,label)
Ilps.append(Ilp.astype(np.uint8))
return Llps, Ilps, elapsed
# + [markdown] colab_type="text" id="sANQtYzpHjtO"
# ## 3. Optical Character Recognition
# + [markdown] colab_type="text" id="Y-SEN7Upu52b"
# OCR is third and last step in ALPR system. In this step, for each detected license plate we apply OCR for a) character segmentation and b) character recognition.
#
# We will perform character segmentation and recognition over the detected LP using a Modified YOLO network, with the same architecture presented in **Real-Time Brazilian License Plate Detection and Recognition Using Deep Convolutional Neural Networks** by <NAME> and <NAME> [[paper](http://sibgrapi.sid.inpe.br/archive.cgi/sid.inpe.br/sibgrapi/2017/08.18.12.21)][[webpage](http://www.inf.ufrgs.br/~smsilva/real-time-brazilian-alpr/)].
#
# The network structure of Modified YOLO
#
# 
#
# We will use weights and config file of Modified YOLO from [here](http://www.inf.ufrgs.br/~smsilva/alpr-unconstrained/data/ocr/), which is same as used by <NAME> an author of **License Plate Detection and Recognition in Unconstrained Scenarios**. The training dataset for Modified YOLO was considerably enlarged by using synthetic and augmented data to cope with LP characteristics of different regions around the world (Europe, United States and Brazil).
#
# The artificially created data consist of pasting a string of seven characters onto a textured background and then performing random transformations, such as rotation, translation, noise, and blur.
#
# Some generated samples and a short overview of the pipeline for synthetic data generation are shown in figure below.
#
# .
#
#
#
#
# + [markdown] colab_type="text" id="J1ws6m5GJRiE"
# ### 3.1 Download weights and config file of pretrained Modified YOLO
# + colab={} colab_type="code" id="Zo5TVA684aL2"
# !wget -c -N www.inf.ufrgs.br/~smsilva/alpr-unconstrained/data/ocr/ocr-net.cfg -P ocr/
# !wget -c -N www.inf.ufrgs.br/~smsilva/alpr-unconstrained/data/ocr/ocr-net.names -P ocr/
# !wget -c -N www.inf.ufrgs.br/~smsilva/alpr-unconstrained/data/ocr/ocr-net.weights -P ocr/
# !wget -c -N www.inf.ufrgs.br/~smsilva/alpr-unconstrained/data/ocr/ocr-net.data -P ocr/
# + [markdown] colab_type="text" id="Ex-RJOxq2gD_"
# ### 3.2 Utility functions
# + [markdown] colab_type="text" id="P1Uf8yGNemAE"
# Let's define a utility function `dknet_label_conversion` to convert detected character bounding box into an object of `Label` class.
# + cellView="both" colab={} colab_type="code" id="6PTwqd2y0aqh"
#@title
# Import necessary modules
import numpy as np
def dknet_label_conversion(R,img_width,img_height, classes):
WH = np.array([img_width,img_height],dtype=float)
L = []
for r in R:
center = np.array(r[2][:2])/WH
wh2 = (np.array(r[2][2:])/WH)*.5
L.append(Label(ord(classes[r[0]]),tl=center-wh2,br=center+wh2,prob=r[1]))
return L
# + [markdown] colab_type="text" id="iZXmukRCH5sA"
# ### 3.3 Recognize characters
# + [markdown] colab_type="text" id="VhsKusP7v7_s"
# Let's define a `lp_ocr` function which takes image of license plate as input, apply Modified YOLO OCR, converts license plate text into string and return that string as output.
#
# We also use `postprocess` utility function to remove character detections with low confidence.
# + colab={} colab_type="code" id="oZ64epbmFpVp"
# Import necessary modules
import cv2 as cv
# Initialize the parameters
ocr_threshold = .4
ocr_weights_path = 'ocr/ocr-net.weights'
ocr_confi_path = 'ocr/ocr-net.cfg'
ocr_classes_path = 'ocr/ocr-net.names'
ocr_classes = None
with open(ocr_classes_path, 'rt') as f:
ocr_classes = f.read().rstrip('\n').split('\n')
# load the network
ocr_net = cv.dnn.readNetFromDarknet(ocr_confi_path, ocr_weights_path)
ocr_net.setPreferableBackend(cv.dnn.DNN_BACKEND_OPENCV)
ocr_net.setPreferableTarget(cv.dnn.DNN_TARGET_CPU)
def lp_ocr(lp_img):
h,w,_ = lp_img.shape
# Create a 4D blob from a frame
blob = cv.dnn.blobFromImage(lp_img, 1/255, (240, 80), [0,0,0], 1, crop=False)
# Sets the input to the network
ocr_net.setInput(blob)
# Runs the forward pass to get output of the output layers
outs = ocr_net.forward(getOutputsNames(ocr_net))
# Remove the bounding boxes with low confidence
R = postprocess(lp_img, outs, ocr_threshold, None)
lp_str = ''
if len(R):
L = dknet_label_conversion(R, w, h, ocr_classes)
L.sort(key=lambda x: x.tl()[0])
lp_str = ''.join([chr(l.cl()) for l in L])
return lp_str
# + [markdown] colab_type="text" id="02Hl-s5DF6Mu"
# ## 4. Inference
#
#
# + [markdown] colab_type="text" id="vztcY-VOxASz"
# We have already defined 1) `vehicle_detection` 2) `lp_detection` and 3) `lp_ocr` to perform vehicle detection, license plate detection and OCR respectively.
#
# Let's implement our ALPR systen which is compised all these three function in a sequential pipeline.
# + [markdown] colab_type="text" id="v2q6BMfiKdkD"
# ### 4.1 Download the test image
# + colab={} colab_type="code" id="hvUSHzxQK0gY"
# !wget "https://raw.githubusercontent.com/sergiomsilva/alpr-unconstrained/master/samples/test/03066.jpg" -O test_img.jpg
# + [markdown] colab_type="text" id="p2IG4NPfVEJX"
# ### 4.2 Utility functions
# + [markdown] colab_type="text" id="3V3_sb7m0aKc"
# Here, we define few utility functions like,
# - `draw_label`: to draw bounding box using object of `Label` class as input
# - `draw_losangle`: to draw bouding box using a set of four corner points.
# - `write2img`: to write text on a input image around given bounding box.
# + cellView="form" colab={} colab_type="code" id="dB5N7ujBPPRR"
#@title
import numpy as np
import cv2 as cv
def draw_label(I,l,color=(255,0,0),thickness=1):
wh = np.array(I.shape[1::-1]).astype(float)
tl = tuple((l.tl()*wh).astype(int).tolist())
br = tuple((l.br()*wh).astype(int).tolist())
cv.rectangle(I,tl,br,color,thickness=thickness)
def draw_losangle(I,pts,color=(1.,1.,1.),thickness=1):
assert(pts.shape[0] == 2 and pts.shape[1] == 4)
for i in range(4):
pt1 = tuple(pts[:,i].astype(int).tolist())
pt2 = tuple(pts[:,(i+1)%4].astype(int).tolist())
cv.line(I,pt1,pt2,color,thickness)
def write2img(Img,label,strg,txt_color=(0,0,0),bg_color=(255,255,255),font_size=1):
wh_img = np.array(Img.shape[1::-1])
font = cv.FONT_HERSHEY_SIMPLEX
wh_text,v = cv.getTextSize(strg, font, font_size, 3)
bl_corner = label.tl()*wh_img
tl_corner = np.array([bl_corner[0],bl_corner[1]-wh_text[1]])/wh_img
br_corner = np.array([bl_corner[0]+wh_text[0],bl_corner[1]])/wh_img
bl_corner /= wh_img
if (tl_corner < 0.).any():
delta = 0. - np.minimum(tl_corner,0.)
elif (br_corner > 1.).any():
delta = 1. - np.maximum(br_corner,1.)
else:
delta = 0.
tl_corner += delta
br_corner += delta
bl_corner += delta
tpl = lambda x: tuple((x*wh_img).astype(int).tolist())
cv.rectangle(Img, tpl(tl_corner), tpl(br_corner), bg_color, -1)
cv.putText(Img,strg,tpl(bl_corner),font,font_size,txt_color,3)
# + [markdown] colab_type="text" id="Guk8LR6IRkr_"
# ### 4.3 Infer on the test image
#
# + [markdown] colab_type="text" id="h4qM8nfGx6O4"
# To infer on the test image we apply
# - first, `vehicle_detection` to detect all vehicles in an input test image. Output of this step is `Icars` a list of cropped vehicles regions as well as `Lcars` a list of bounding boxs around detected vehicles.
#
# - second, `lp_detection` on each cropped vehicle regions in `Icars` to detect license plates. Output of this step is `Llps` a list of bouding boxs around detected license plate as well as `Ilps` cropped images of license plate in given vehicle image.
#
# - third, `lo_ocr` on each cropped license plate region in `Ilps` to convert licese plat text in it to a string.
#
# - finally, `write2img` to write recognized license plat charactes on the input test image.
#
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="O9febVzWb3q9" outputId="e4357826-bd50-41ce-93da-a316751b31a1"
# Import necessary modules
import numpy as np
import cv2 as cv
# read test image
test_img = cv.imread('test_img.jpg')
# detect cars
Icars, Lcars = vehicle_detection(test_img)
print('# vehicle detected: {}'.format(len(Icars)))
# for each detected car in test image
for Icar, Lcar in zip(Icars, Lcars):
# draw car bounding box on test image
draw_label(test_img,Lcar,color=(0,255,255),thickness=3)
# detect LP in detected car
Llps, Ilps, elapsed = lp_detection(Icar)
# for each detected LP in the detected car image
for Llp, Ilp in zip(Llps, Ilps):
# draw LP bounding box on test image
tl = Llp.tl() * Lcar.wh() + Lcar.tl()
br = Llp.br() * Lcar.wh() + Lcar.tl()
draw_label(test_img,Label(0,tl,br),color=(255,0,0),thickness=3)
# Recognize characters
lp_str = lp_ocr(Ilp)
# write text on test image
write2img(test_img,Label(0,tl,br),lp_str)
# + [markdown] colab_type="text" id="AimbDxuwRkse"
# ### 4.4 Display inference
# + colab={"base_uri": "https://localhost:8080/", "height": 470} colab_type="code" id="YezwhstGRksj" outputId="122ce949-1f56-47b8-b336-0827db7d2f64"
# Import necessary modules
import matplotlib
import matplotlib.pyplot as plt
# Display inference
fig=plt.figure(figsize=(10, 10))
plt.imshow(test_img[:,:,::-1])
plt.show()
# + [markdown] colab_type="text" id="Hi4yj9G7o5EO"
# ### 4.5 Observations
#
# + [markdown] colab_type="text" id="t1jYYfEkLfR2"
# From the above result we can make following observations,
#
# - There is improvement in performance of text recognition for Modified YOLO as compared to tesseract
#
# - Even now, if the images are given in unconstrained scenarios like more rotated and different view of the car's license plate, our YOLO-Net fails to give accurate bounding box around license plate which may affect the performace of OCR.
#
# In next notebook, we will use WPOD-Net to get more tight bounding box around license plate and see how it improve overall ALPR performance.
| ALPR/4_ALPR_using_YOLO_Net_LP_Modified_YOLO_OCR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # RNA-Seq Analysis Training Demo
# ## Overview
# This short tutorial demonstrates how to run an RNA-Seq workflow using a prokaryotic data set. Steps in the workflow include read trimming, read QC, read mapping, and counting mapped reads per gene to quantitate gene expression.
#
# 
# ### STEP 1: Install Mambaforge and then install snakemake using bioconda.
# First install Mambaforge.
#
# !curl -L -O https://github.com/conda-forge/miniforge/releases/latest/download/Mambaforge-$(uname)-$(uname -m).sh
# !bash Mambaforge-$(uname)-$(uname -m).sh -b -u -p $HOME/mambaforge
# !export PATH="$HOME/mambaforge/bin:$PATH"
# Next, install Trimmomatic using bioconda.
#
# !$HOME/mambaforge/bin/mamba install -y -c conda-forge -c bioconda trimmomatic
# !$HOME/mambaforge/bin/mamba install -y -c conda-forge -c bioconda fastqc
# !$HOME/mambaforge/bin/mamba install -y -c conda-forge -c bioconda multiqc
# !$HOME/mambaforge/bin/mamba install -y -c conda-forge -c bioconda salmon
# ### STEP 2: Setup Environment
# Create a set of directories to store the reads, reference sequence files, and output files.
#
# !cd $HOMEDIR
# !echo $PWD
# !mkdir -p data
# !mkdir -p data/raw_fastq
# !mkdir -p data/trimmed
# !mkdir -p data/fastqc
# !mkdir -p data/aligned
# !mkdir -p data/reference
# ### STEP 2: Copy FASTQ Files
# In order for this tutorial to run quickly, we will only analyze 50,000 reads from a sample from both sample groupsinstead of analyzing all the reads from all six samples. These files have been posted on a Google Storage Bucket that we made publicly accessible.
#
# !curl https://storage.googleapis.com/me-inbre-rnaseq-pipelinev2/data/raw_fastqSub/SRR13349122_1.fastq --output data/raw_fastq/SRR13349122_1.fastq
# !curl https://storage.googleapis.com/me-inbre-rnaseq-pipelinev2/data/raw_fastqSub/SRR13349122_2.fastq --output data/raw_fastq/SRR13349122_2.fastq
# !curl https://storage.googleapis.com/me-inbre-rnaseq-pipelinev2/data/raw_fastqSub/SRR13349128_1.fastq --output data/raw_fastq/SRR13349128_1.fastq
# !curl https://storage.googleapis.com/me-inbre-rnaseq-pipelinev2/data/raw_fastqSub/SRR13349128_2.fastq --output data/raw_fastq/SRR13349128_2.fastq
# ### STEP 3: Copy reference transcriptome files that will be used by Salmon
# Salmon is a tool that aligns RNA-Seq reads to a set of transcripts rather than the entire genome.
# !curl https://storage.googleapis.com/me-inbre-rnaseq-pipelinev2/data/reference/M_chelonae_transcripts.fasta --output data/reference/M_chelonae_transcripts.fasta
# !curl https://storage.googleapis.com/me-inbre-rnaseq-pipelinev2/data/reference/decoys.txt --output data/reference/decoys.txt
# ### STEP 4: Copy data file for Trimmomatic
# !curl https://storage.googleapis.com/me-inbre-rnaseq-pipelinev2/config/TruSeq3-PE.fa --output TruSeq3-PE.fa
# ### STEP 5: Run Trimmomatic
# Trimmomatic will trim off any adapter sequences or low quality sequence it detects in the FASTQ files.
# !trimmomatic PE -threads 2 data/raw_fastq/SRR13349122_1.fastq data/raw_fastq/SRR13349122_2.fastq data/trimmed/SRR13349122_1_trimmed.fastq data/trimmed/SRR13349122_2_trimmed.fastq data/trimmed/SRR13349122_1_trimmed_unpaired.fastq data/trimmed/SRR13349122_2_trimmed_unpaired.fastq ILLUMINACLIP:TruSeq3-PE.fa:2:30:10:2:keepBothReads LEADING:3 TRAILING:3 MINLEN:36
# !trimmomatic PE -threads 2 data/raw_fastq/SRR13349128_1.fastq data/raw_fastq/SRR13349128_2.fastq data/trimmed/SRR13349128_1_trimmed.fastq data/trimmed/SRR13349128_2_trimmed.fastq data/trimmed/SRR13349128_1_trimmed_unpaired.fastq data/trimmed/SRR13349128_2_trimmed_unpaired.fastq ILLUMINACLIP:TruSeq3-PE.fa:2:30:10:2:keepBothReads LEADING:3 TRAILING:3 MINLEN:36
# ### STEP 6: Run FastQC
# FastQC is an invaluable tool that allows you to evaluate whether there are problems with a set of reads. For example, it will provide a report of whether there is any bias in the sequence composition of the reads.
# !fastqc -o data/fastqc data/trimmed/SRR13349122_1_trimmed.fastq
# !fastqc -o data/fastqc data/trimmed/SRR13349128_1_trimmed.fastq
# ### STEP 7: Run MultiQC
# MultiQC reads in the FastQQ reports and generate a compiled report for all the analyzed FASTQ files.
# !multiqc -f data/fastqc
# ### STEP 8: Index the Transcriptome so that Trimmed Reads Can Be Mapped Using Salmon
# !salmon index -t data/reference/M_chelonae_transcripts.fasta -p 8 -i data/reference/transcriptome_index --decoys data/reference/decoys.txt -k 31 --keepDuplicates
# ### STEP 9: Run Salmon to Map Reads to Transcripts and Quantify Expression Levels
# Salmon aligns the trimmed reads to the reference transcriptome and generates the read counts per transcript. In this analysis, each gene has a single transcript.
# !salmon quant -i data/reference/transcriptome_index -l SR -r data/trimmed/SRR13349122_1_trimmed.fastq -p 8 --validateMappings -o data/quants/SRR13349122_quant
# !salmon quant -i data/reference/transcriptome_index -l SR -r data/trimmed/SRR13349128_1_trimmed.fastq -p 8 --validateMappings -o data/quants/SRR13349128_quant
# ### STEP 10: Report the top 10 most highly expressed genes in the samples
# Top 10 most highly expressed genes in the wild-type sample.
#
# !sort -nrk 4,4 data/quants/SRR13349122_quant/quant.sf | head -10
# Top 10 most highly expressed genes in the double lysogen sample.
#
# !sort -nrk 4,4 data/quants/SRR13349128_quant/quant.sf | head -10
# ### STEP 11: Report the expression of a putative acyl-ACP desaturase (BB28_RS16545) that was downregulated in the double lysogen relative to wild-type
# A acyl-transferase was reported to be downregulated in the double lysogen as shown in the table of the top 20 upregulated and downregulated genes from the paper describing the study.
# 
# Use `grep` to report the expression in the wild-type sample. The fields in the Salmon `quant.sf` file are as follows. The level of expression is reported in the Transcripts Per Million (`TPM`) and number of reads (`NumReads`) fields:
# `Name Length EffectiveLength TPM NumReads`
# !grep 'BB28_RS16545' data/quants/SRR13349122_quant/quant.sf
# Use `grep` to report the expression in the double lysogen sample. The fields in the Salmon `quant.sf` file are as follows. The level of expression is reported in the Transcripts Per Million (`TPM`) and number of reads (`NumReads`) fields:
# `Name Length EffectiveLength TPM NumReads`
# !grep 'BB28_RS16545' data/quants/SRR13349128_quant/quant.sf
# ### The next tutorial will focus on the analysis of the gene counts
# Now that you have read counts per gene, the next tutorial will show you the following workflow that is used to generate the list of differentially expressed genes.
#
# 
| notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# -
data = pd.read_csv("raw_data1_with_labels.csv", index_col = 0)
data.head()
# +
# Getting the labels and the title column
y = data['y']
text = data['title']
# +
# It'll be create functions to process the data like the data processing notebook
def getting_numeric_features(data):
df = data.copy()
features = pd.DataFrame(index = df.index)
features['date'] = pd.to_datetime(df['upload_date'])
features['views'] = df['view_count']
features['time_since_pub'] = (pd.to_datetime("2020-12-12") - features['date']) / np.timedelta64(1, 'D')
features['views_per_day'] = features['views'] / features['time_since_pub']
features.drop(columns = ["time_since_pub", "date"], inplace = True)
return features
# -
features = getting_numeric_features(data)
features.head()
# +
# splitting the data into train and validation
mask1 = data['upload_date'] <= "2020-08-31" # 63% - be use to train the model
mask2 = data['upload_date'] > "2020-08-31" # 27% - be use to test the model
X_train, X_val = features[mask1], features[mask2]
y_train, y_val = y[mask1], y[mask2]
X_train.shape, y_train.shape, X_val.shape, y_val.shape
# +
from sklearn.feature_extraction.text import TfidfVectorizer
from scipy.sparse import hstack
def getting_text_features(text, mask1, mask2, min_df = 2, ngram_range = (1, 3)):
X_train_text = np.squeeze(text[mask1])
X_val_text = np.squeeze(text[mask2])
title_vec = TfidfVectorizer(min_df=min_df, ngram_range=ngram_range)
X_train_bow = title_vec.fit_transform(X_train_text)
X_val_bow = title_vec.transform(X_val_text)
return X_train_bow, X_val_bow, title_vec
def final_data(x_train, x_val, x_train_bow, x_val_bow):
return hstack([x_train, x_train_bow]), hstack([x_val, x_val_bow])
# +
x_train_bow, x_val_bow, title_vec_rf = getting_text_features(text, mask1, mask2)
x_train_bow.shape, x_val_bow.shape
# -
X_train_with_title, X_val_with_title = final_data(X_train, X_val, x_train_bow, x_val_bow)
X_train_with_title.shape, X_val_with_title.shape
# *After getting the data prepared to feed into the model, we are building some models to compare their metrics.*
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
# ### Random Forest Classifier
mdl1 = RandomForestClassifier(n_estimators = 1000, random_state = 0, n_jobs = -1, class_weight = "balanced", min_samples_leaf= 1)
mdl1.fit(X_train_with_title, y_train)
# +
from sklearn.metrics import roc_auc_score, average_precision_score
p_rf = mdl1.predict_proba(X_val_with_title)[:, 1]
average_precision_score(y_val, p_rf), roc_auc_score(y_val, p_rf)
# -
#
# (0.4398304045590999, 0.6974981833281428) - n_estimators = 1000 - min_samples_leaf= 1 - n_gram (1,2)
# (0.4281493555354776, 0.6715716806809924) - n_estimators = 1000 - min_samples_leaf= 2 - n_gram (1,2)
# (0.41291598480012126, 0.6572978303747534) - n_estimators = 100 - min_samples_leaf= 2 - n_gram (1,2)
# (0.4390506690818257, 0.6829648084708814) - n_estimators = 1000 - min_samples_leaf= 1 - n_gram (1,2)
# (0.47479006692854603, 0.7139520398629711) - n_estimators = 1000 - min_samples_leaf= 1 - n_gram (1,3) ~ best
# ### LGBM Classifier
# +
from lightgbm import LGBMClassifier
mdl2 = LGBMClassifier(random_state = 0, class_weight="balanced", n_jobs = -1)
mdl2.fit(X_train, y_train)
# -
p = mdl2.predict_proba(X_val)[:, 1]
average_precision_score(y_val, p), roc_auc_score(y_val, p)
# +
# Now we will use the scikit-optimize library (skopt) to tune the lgbm classifier
from skopt import forest_minimize
# -
title_train = np.squeeze(text[mask1])
title_val = np.squeeze(text[mask2])
def tune_lgbm(params):
lr = params[0]
max_depth = params[1]
min_child_samples = params[2]
subsample = params[3]
colsample_bytree = params[4]
n_estimators = params[5]
min_df = params[6]
ngram_range = (1, params[7])
title_vec = TfidfVectorizer(min_df = min_df, ngram_range = ngram_range)
title_bow_train = title_vec.fit_transform(title_train)
title_bow_val = title_vec.transform(title_val)
X_train_with_title = hstack([X_train, title_bow_train])
X_val_with_title = hstack([X_val, title_bow_val])
mdl = LGBMClassifier(learning_rate = lr, max_depth=max_depth,
min_child_samples=min_child_samples, subsample=subsample,
colsample_bytree=colsample_bytree, bagging_freq = 1, n_estimators= n_estimators,
random_state =0, class_weight = "balanced", n_jobs=-1)
mdl.fit(X_train_with_title, y_train)
p = mdl.predict_proba(X_val_with_title)[:, 1]
print(roc_auc_score(y_val, p))
return -average_precision_score(y_val, p)
# +
space = [(1e-3, 1e-1, 'log-uniform'), #lr
(1, 20), # max_depth
(1, 20), #min_child_samples
(0.05, 1.), #subsample
(0.05, 1.), #colsample_bytree
(100, 1000), # n_estimators
(1, 5), # min_df
(1, 5)] #n_gram range
res = forest_minimize(tune_lgbm, space, random_state = 160475, n_random_starts = 20, n_calls = 50, verbose = 1)
# -
print(f'Best parameters: {res.x}')
# ### Best Model
x_train_bow, x_val_bow, title_vec_lgbm = getting_text_features(text, mask1, mask2, min_df=4, ngram_range=(1, 1))
X_train_with_title, X_val_with_title = final_data(X_train, X_val, x_train_bow, x_val_bow)
X_train_with_title.shape,
mdl2 = LGBMClassifier(random_state = 0, class_weight="balanced", n_jobs = -1, learning_rate=0.015286972843636785, max_depth = 14,
min_child_samples = 1, subsample = 0.5605787546434184, colsample_bytree=0.9289814346488457,
n_estimators = 539)
mdl2.fit(X_train_with_title, y_train)
p_lgbm = mdl2.predict_proba(X_val_with_title)[:, 1]
average_precision_score(y_val, p_lgbm), roc_auc_score(y_val, p_lgbm)
# ## Ensemble
# LGBM Classifier - (0.469143426580733, 0.715275615073186)
#
#
# Random Forest - (0.47479006692854603, 0.7139520398629711)
pd.DataFrame({"LR" : p_rf, "LGBM": p_lgbm}).corr()
# *the low correlation value indicates that getting this two models together will improve our model*
p = 0.5*p_rf + 0.5*p_lgbm
average_precision_score(y_val, p), roc_auc_score(y_val, p)
# ## Save our models
# +
import joblib as jb
jb.dump(mdl2, "mdl_lgbm.pkl.z")
jb.dump(mdl1, "mdl_random_forest.pkl.z")
# -
jb.dump(title_vec_rf, "title_vectorizer_rf.pkl.z")
jb.dump(title_vec_lgbm, "title_vectorizer_lgbm.pkl.z")
| Data Modeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kwbt-kzk/github-slideshow/blob/main/excercise_3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="kwVUgf-_xiYz"
## Repltの黒い部分にが相変わらず入力できませんでしたので、先生にご提案していただいたこのファイルで提出いたします。
## お手数をおかけいたします
## GitHubのIDは kwbt-kzk です
## 学籍番号は 15719024 です
## 氏名は 川端一輝 です
# + id="xOQKmQac-cz_"
##Excercise 3.1
##Part 1
basename="Station"
filenames=[]
##Part 2
for value in range(21):
filenames.append(basename+''+str(value)+'.txt')
print(filenames)
# + id="5v1zAdBVQKMJ"
##Excercise 3.2
temperatures = [
-5.4, 1.0, -1.3, -4.8, 3.9, 0.1, -4.4, 4.0, -2.2, -3.9, 4.4, -2.5, -4.6,
5.1, 2.1, -2.4, 1.9, -3.3, -4.8, 1.0, -0.8, -2.8, -0.1, -4.7, -5.6, 2.6,
-2.7, -4.6, 3.4, -0.4, -0.9, 3.1, 2.4, 1.6, 4.2, 3.5, 2.6, 3.1, 2.2, 1.8,
3.3, 1.6, 1.5, 4.7, 4.0, 3.6, 4.9, 4.8, 5.3, 5.6, 4.1, 3.7, 7.6, 6.9, 5.1,
6.4, 3.8, 4.0, 8.6, 4.1, 1.4, 8.9, 3.0, 1.6, 8.5, 4.7, 6.6, 8.1, 4.5, 4.8,
11.3, 4.7, 5.2, 11.5, 6.2, 2.9, 4.3, 2.8, 2.8, 6.3, 2.6, -0.0, 7.3, 3.4,
4.7, 9.3, 6.4, 5.4, 7.6, 5.2
]
##Part 1
cold=[]
slippery=[]
comfortable=[]
warm=[]
##Part 2
for temperaure in temperatures:
if temperature < -2:
cold.append(temperature);
elif -2 <= temperature < 2:
Slippery.append(temperature)
elif 2 <= temperature < 15:
Comfortable.append(temperature)
elif 15 < temperature:
Warm.append(temperature)
##Part 3
slippery_times='XXX'
Slippery_times=len(slippery)
print("In April 2013 it was slippery", slippery_times, "times.")
warm_times='XXX'
cold_times='XXX'
cold_times=len(cold)
print("In April 2013 it was cold", cold_times, "times.")
# + colab={"base_uri": "https://localhost:8080/"} id="APb8D2C6YsTv" outputId="a7f49196-8c4f-4f2c-c302-e872a3c6eb8f"
##Excercise 3.3
stations = ['Hanko Russarö', 'Heinola Asemantaus', 'Helsinki Kaisaniemi',
'Helsinki Malmi airfield', 'Hyvinkää Hyvinkäänkylä', 'Joutsa Savenaho',
'Juuka Niemelä', 'Jyväskylä airport', 'Kaarina Yltöinen', 'Kauhava airfield',
'Kemi Kemi-Tornio airport', 'Kotka Rankki', 'Kouvola Anjala',
'Kouvola Utti airport', 'Kuopio Maaninka', 'Kuusamo airport',
'Lieksa Lampela', 'Mustasaari Valassaaret', 'Parainen Utö', 'Pori airport',
'Rovaniemi Apukka', 'Salo Kärkkä', 'Savonlinna Punkaharju Laukansaari',
'Seinäjoki Pelmaa', 'Siikajoki Ruukki', 'Siilinjärvi Kuopio airport',
'Tohmajärvi Kemie', 'Utsjoki Nuorgam', 'Vaala Pelso', 'Vaasa airport',
'Vesanto Sonkari', 'Vieremä Kaarakkala', 'Vihti Maasoja', 'Ylitornio Meltosjärvi']
lats = [59.77, 61.2, 60.18, 60.25, 60.6, 61.88, 63.23, 62.4,
60.39, 63.12, 65.78, 60.38, 60.7, 60.9, 63.14, 65.99,
63.32, 63.44, 59.78, 61.47, 66.58, 60.37, 61.8, 62.94,
64.68, 63.01, 62.24, 70.08, 64.501, 63.06, 62.92, 63.84,
60.42, 66.53]
lons = [22.95, 26.05, 24.94, 25.05, 24.8, 26.09, 29.23, 25.67,
22.55, 23.04, 24.58, 26.96, 26.81, 26.95, 27.31, 29.23,
30.05, 21.07, 21.37, 21.79, 26.01, 23.11, 29.32, 22.49,
25.09, 27.8, 30.35, 27.9, 26.42, 21.75, 26.42, 27.22,
24.4, 24.65]
north_south_cutoff = 64.5
east_west_cutoff = 26.3
##Part 1
north_west=[]
north_east=[]
south_west=[]
south_east=[]
print(north_west, south_west)
##Part 2
n=len(stations)
print("In the data, there are", n, "stations.")
##Part 3
##飛ばします
##Part 4
for number in range(n):
if lats[number] > north_south_cutoff:
if lons[number] > east_west_cutoff:
north_east.append(stations[number])
else:
north_west.append(stations[number])
else:
if lons[number] > east_west_cutoff:
south_east.append(stations[number])
else:
south_west.append(stations[number])
# This test print should print out station names in North West
# Hint: there should be 4 stations in this class
print("The names of the Northwest stations are:\n", north_west)
# This test print should print out station names in North Eest
# Hint: there should be 3 stations in this class
print("The names of the Northeast stations are:\n", north_east)
# This test print should print out station names in South West
# Hint: there should be 16 stations in this class
print("The names of the Southwest stations are:\n", south_west)
# This test print should print out station names in South East
# Hint: there should be 11 stations in this class
print("The names of the Southeast stations are:\n", south_east)
##Part 5
## 1.Almost no problem.
## 2.I did.
##Part 6
print("Northwest contains{share: .0f} % of all stations.".format(share=north_west_share))
print("Northeast contains{share: .0f} % of all stations.".format(share=north_east_share))
print("Southwest contains{share: .0f} % of all stations.".format(share=south_west_share))
print("Southeast contains{share: .0f} % of all stations.".format(share=south_east_share))
| excercise_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
import numpy as np
# +
# Get the numbers of columns in the csv:
csv_in = open("ex.csv", "r") # open the csv
ncol = len(csv_in.readline().split(",")) # read the first line and count the # of columns
csv_in.close() # close the csv
print("Number of columns in the csv: " + str(ncol)) # print the # of columns
# +
# WORKING VERSION
# Create random column order list (dataset) + iterator
col_list = tf.data.Dataset.range(ncol).shuffle(buffer_size=ncol)
col_next = col_list.make_one_shot_iterator().get_next()
#def scale_zscore(vector):
# mean, var = tf.nn.moments(vector, axes=[0])
# normalized_col = tf.map_fn(lambda x: (x - mean)/tf.sqrt(var), vector)
# return normalized_col
# Launch of graph
with tf.Session() as sess:
while True: # Loop on 'col_next', the queue of column iterator
try:
index = sess.run(col_next)
dataset = tf.contrib.data.CsvDataset( # Creates a dataset of the current csv column
"ex.csv",
[tf.float32],
select_cols=[index] # Only parse last three columns
)
next_element = dataset.make_one_shot_iterator().get_next() # Creates an iterator
print('Current column to be full pass: ' + str(index))
current_col = []
while True:
try:
current_col.append(sess.run(next_element)[0]) # Full pass
except tf.errors.OutOfRangeError: # End of full pass
print(current_col)
current_col = tf.convert_to_tensor([current_col])
mean, var = tf.nn.moments(current_col, axes=[0])
normalized_col = tf.map_fn(lambda x: (x - mean)/tf.sqrt(var), current_col)
print(normalized_col)
print('\n')
break
except tf.errors.OutOfRangeError:
break
# -
| notebooks/dev.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/porcelainruler/Kaggle/blob/main/Titanic.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="TG7Xa1XsmI-y" outputId="2db808a0-68d2-4d7e-fcf7-bf730f7b6c1d"
# # ! pip install -q tensorflow
import tensorflow as tf
print(tf.__version__)
import pandas as pd
import numpy as np
import seaborn as sbs
import re
# To plot within notebook
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
# Setting figure size
from matplotlib.pylab import rcParams
rcParams['figure.figsize'] = 20,10
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, LSTM
from tensorflow.keras import regularizers
# For normalizing data
import sklearn
from sklearn.tree import DecisionTreeClassifier
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
scaler = MinMaxScaler(feature_range=(0, 1))
import warnings
warnings.filterwarnings('ignore')
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 128} id="3d0Ri5hwmcjK" outputId="547f1d6e-068f-47b0-e9aa-daa8a1819ffe"
from google.colab import files
uploaded1 = files.upload()
uploaded2 = files.upload()
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="X2hEzbwonun-" outputId="9d0f3e29-b5e2-434d-c52c-91fae3cf15dd"
import io
train = pd.read_csv(io.BytesIO(uploaded1['train.csv']))
test = pd.read_csv(io.BytesIO(uploaded2['test.csv']))
all = [train, test]
train.head()
test.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 212} id="QLGDcQAarVKz" outputId="6b272b61-d9b8-4891-d783-98d07abe854b"
'''
df = df.fillna(value=0)
# df = df.drop(columns=['Name'])
# df = df.drop(columns=['Ticket'])
print(df.iloc[0, 3])
for idx in range(len(df)):
if df.iloc[idx, 3] == 'male':
df.iloc[idx, 3] = 0
else:
df.iloc[idx, 3] = 1
if str(df.iloc[idx, 8])[0] == 'C':
df.iloc[idx, 8] = str(df.iloc[idx, 8])[1:]
df.head()
'''
# + colab={"base_uri": "https://localhost:8080/", "height": 246} id="CZkMjaQ3_z1r" outputId="dab8acdf-c985-4117-f0c7-1b72d9d864bd"
# Creating FamilySize Feature
for data in all:
data['family_size'] = data['SibSp'] + data['Parch'] + 1
# Creating isAlone Feature
for data in all:
data['is_alone'] = 0
data.loc[data['family_size'] == 1, 'is_alone'] = 1
# NaN Value in Embarked Column
for data in all:
data['Embarked'] = data['Embarked'].fillna('S')
# NaN Value in Fare Column
for data in all:
data['Fare'] = data['Fare'].fillna(data['Fare'].median())
train['category_fare'] = pd.qcut(train['Fare'], 4)
# Missing Age Value Handling
for data in all:
age_avg = data['Age'].mean()
age_std = data['Age'].std()
age_null = data['Age'].isnull().sum()
random_list = np.random.randint(age_avg - age_std, age_avg + age_std , size = age_null)
data['Age'][np.isnan(data['Age'])] = random_list
data['Age'] = data['Age'].astype(int)
train['category_age'] = pd.cut(train['Age'], 5)
#Feature 7: Name
def get_title(name):
title_search = re.search(' ([A-Za-z]+)\. ', name)
if title_search:
return title_search.group(1)
return ""
for data in all:
data['title'] = data['Name'].apply(get_title)
for data in all:
data['title'] = data['title'].replace(['Lady', 'Countess','Capt', 'Col','Don', 'Dr', 'Major', 'Rev', 'Sir', 'Jonkheer', 'Dona'],'Rare')
data['title'] = data['title'].replace('Mlle','Miss')
data['title'] = data['title'].replace('Ms','Miss')
data['title'] = data['title'].replace('Mme','Mrs')
train.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="4ABugcX5EnDN" outputId="3c423cd1-095d-4c93-a7f3-d2578cfca12b"
#Map Data
for data in all:
#Mapping Sex
sex_map = { 'female':0 , 'male':1 }
data['Sex'] = data['Sex'].map(sex_map).astype(int)
#Mapping Title
title_map = {'Mr':1, 'Miss':2, 'Mrs':3, 'Master':4, 'Rare':5}
data['title'] = data['title'].map(title_map)
data['title'] = data['title'].fillna(0)
#Mapping Embarked
embark_map = {'S':0, 'C':1, 'Q':2}
data['Embarked'] = data['Embarked'].map(embark_map).astype(int)
#Mapping Fare
data.loc[ data['Fare'] <= 7.91, 'Fare'] = 0
data.loc[(data['Fare'] > 7.91) & (data['Fare'] <= 14.454), 'Fare'] = 1
data.loc[(data['Fare'] > 14.454) & (data['Fare'] <= 31), 'Fare'] = 2
data.loc[ data['Fare'] > 31, 'Fare'] = 3
data['Fare'] = data['Fare'].astype(int)
#Mapping Age
data.loc[ data['Age'] <= 16, 'Age'] = 0
data.loc[(data['Age'] > 16) & (data['Age'] <= 32), 'Age'] = 1
data.loc[(data['Age'] > 32) & (data['Age'] <= 48), 'Age'] = 2
data.loc[(data['Age'] > 48) & (data['Age'] <= 64), 'Age'] = 3
data.loc[ data['Age'] > 64, 'Age'] = 4
#Mapping Age*Title
data['Age*Title'] = data['Age'] * data['title']
#Feature Selection
#Create list of columns to drop
drop_elements = ["Name", "Ticket", "Cabin", "SibSp", "Parch", "family_size"]
#Drop columns from both data sets
train = train.drop(drop_elements, axis = 1)
train = train.drop(['PassengerId','category_fare', 'category_age'], axis = 1)
test = test.drop(drop_elements, axis = 1)
train.head()
# + id="0PWiKl3yt9Ve"
X_train = train.drop(columns=['Survived'], axis=1)
Y_train = train['Survived']
X_test = test.drop("PassengerId", axis=1).copy()
# X.head()
# Y.head()
# X_norm = scaler.fit_transform(X)
# X_norm[1][2]
X_train, X_val_and_test, Y_train, Y_val_and_test = train_test_split(X_train, Y_train, test_size=0.25)
x_val, x_test, y_val, y_test = train_test_split(X_val_and_test, Y_val_and_test, test_size=0.5)
# + colab={"base_uri": "https://localhost:8080/", "height": 402} id="mcRoIu7iGp2f" outputId="dd859398-d402-4a91-e00a-d830c503a18c"
X_train
# X_test
# + id="MVzUKabLmczZ"
# Data Plot
# df = sbs.load_dataset('data')
# print(sbs.pairplot(data=data))
sbs.pairplot(data=data)
plt.show()
# Data Correlation
# -- 1st Tech
plt.matshow(data.corr())
plt.show()
# -- 2nd Tech
'''
f = plt.figure(figsize=(7, 7))
plt.matshow(data.corr(), fignum=f.number)
plt.xticks(range(data.shape[1]), data.columns, fontsize=14, rotation=45)
plt.yticks(range(data.shape[1]), data.columns, fontsize=14)
cb = plt.colorbar()
cb.ax.tick_params(labelsize=14)
plt.title('Correlation Matrix', fontsize=16);
plt.show()
'''
# -- 3rd Tech
'''
corr = data.corr()
f = plt.figure(figsize=(8,8))
corr.style.background_gradient(cmap='coolwarm')
plt.matshow(corr, fignum=f.number)
plt.show(corr)
'''
# + colab={"base_uri": "https://localhost:8080/"} id="AxG6NpBqTaxw" outputId="117cce8d-85b4-4ade-936c-44680744f10d"
# Decision Tree Model
decision_tree = DecisionTreeClassifier()
decision_tree.fit(X_train, Y_train)
Y_pred = decision_tree.predict(X_test)
accuracy = round(decision_tree.score(X_train, Y_train) * 100, 2)
print("Model Accuracy: ",accuracy)
# + colab={"base_uri": "https://localhost:8080/"} id="4DmeVQd_uucs" outputId="13c04a4c-b8af-4ef5-b103-1e6fa386ee61"
model = Sequential()
model.add(Dense(48, activation='selu', kernel_initializer='lecun_normal', kernel_regularizer=regularizers.l2(0.01), input_shape=(8,)))
model.add(Dropout(0.25))
model.add(Dense(32, activation='selu', kernel_initializer='lecun_normal', kernel_regularizer=regularizers.l2(0.01)))
model.add(Dropout(0.25))
model.add(Dense(1, activation='sigmoid', kernel_regularizer=regularizers.l2(0.01)))
model.compile(optimizer='adam', loss='binary_crossentropy', metrics=['accuracy'])
hist = model.fit(X_train, Y_train, batch_size=32, epochs=100, validation_data=(x_val, y_val))
# print(model.summary())
# + colab={"base_uri": "https://localhost:8080/"} id="qMU0HVnIztcu" outputId="5030be12-a861-4027-d926-85dfe7aba770"
model.evaluate(x_test, y_test)[1]
# + colab={"base_uri": "https://localhost:8080/", "height": 621} id="i2JDzwuaz8M8" outputId="a2df4811-424b-43b1-9506-16ecf461f0db"
plt.plot(hist.history['loss'])
plt.plot(hist.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Val'], loc='upper right')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="JF7iLqM_eHGQ" outputId="f0102be3-bd1c-4d10-f554-b1a76a87e008"
# Create and Fit the LSTM network - Not Working
x_train, y_train = np.array(X_train), np.array(Y_train)
x_train = np.reshape(x_train, (x_train.shape[0],x_train.shape[1],1))
model = Sequential()
model.add(LSTM(units=50, return_sequences=True, input_shape=(x_train.shape[1],1)))
model.add(LSTM(units=50))
model.add(Dense(1))
model.compile(optimizer='adam', loss='binary_crossentropy', metrics=['accuracy'])
model.fit(x_train, y_train, epochs=100, batch_size=16, verbose=2)
# + colab={"base_uri": "https://localhost:8080/"} id="m8toeKKGQM8m" outputId="0f945049-95e1-4adf-cdb3-ac897fdb5aba"
Y = model.predict(X_test, batch_size=32)
Y_dt = Y_pred
Y[:20]
#Y_dt[:20]
# + colab={"base_uri": "https://localhost:8080/", "height": 975} id="ffbTAA_b1-c8" outputId="eb765498-e4e0-4e91-8e9b-5b5edddcf730"
Y = model.predict(X_test, batch_size=32)
Y_dt = Y_pred
out = pd.DataFrame(index=range(len(X)), columns=['PassengerID', 'Survived'])
for idx in range(len(X)):
out.iloc[idx, 0] = X.iloc[idx, 0]
if idx < 20:
print(out.iloc[idx, 0], Y[idx])
if Y[idx] > 0.5:
out.iloc[idx, 1] = 1
else:
out.iloc[idx, 1] = 0
out.head(20)
# + colab={"base_uri": "https://localhost:8080/"} id="a0yVdZKd1oMM" outputId="ff1bcefb-86af-482f-9851-04450393e24a"
Y[:20]
# + colab={"base_uri": "https://localhost:8080/"} id="0iRtfbrB53HD" outputId="26c0fd6c-258b-44a0-ebf3-b64423b11eff"
from google.colab import drive
drive.mount('drive')
# + id="wT0UOJxq6Ghf"
out.to_csv('titanic_NN_3D_Selu_L2-1-Reg_result.csv', index=False)
# !cp titanic_NN_3D_Selu_L2-1-Reg_result.csv "drive/My Drive/"
# + colab={"base_uri": "https://localhost:8080/"} id="FdawVwuB7ckf" outputId="8067193c-9e87-46dd-ca91-4d7698d89b6b"
print(len(out))
| Titanic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from selenium import webdriver
from bs4 import BeautifulSoup as bs
from time import sleep
# ## practice 1
# +
ch_driver = webdriver.Chrome('chromedriver_path')
url = 'https://www.tripadvisor.com/Restaurants-g191-United_States.html'
ch_driver.get(url)
ch_driver.find_element_by_class_name('continue').click()
cities = ch_driver.find_elements_by_class_name('geo_name')
# for check
len(cities)
# -
cities[0].find_element_by_tag_name('a').click()
sleep(3)
soup = bs(ch_driver.page_source, "html.parser")
results = soup.find('div', attrs={'id':'EATERY_SEARCH_RESULTS'})
restaurant = results.find('div', attrs={'data-index': '1'})
# get restaurant name
title_div = restaurant.find('div', attrs={'class': 'title'})
title = title_div.find('a').text.strip('\n')
title
# get rating
rating = restaurant.find('span', attrs={'class' : 'ui_bubble_rating bubble_45'}).attrs['alt']
rating
# get review count
count = restaurant.find('span', attrs={'class' : 'reviewCount'}).find('a').text.strip('\n')
count
# get price
try:
price = restaurant.find('span', attrs={'class' : 'item price'}).text
except:
price = "N/A"
price
# get cuisine
try:
cuisine_div = restaurant.find('div', attrs={'class' : 'cuisines'}).find_all('a')
cuisine = []
for div in cuisine_div:
cuisine.append(div.text)
except:
cuisine = []
cuisine
# +
review_ul = restaurant.find('ul', attrs={'class' : 'review_stubs review_snippets rebrand'})
review_a = review_ul.find_all('a')
review = []
for tag in review_a:
review.append(tag.text)
review
# -
# ## practice 2
ch_driver.back()
cities_div = ch_driver.find_elements_by_class_name('geo_name')
city_list = []
for item in cities_div:
name = item.find_element_by_tag_name('a').text
city_list.append(name)
city_list
#city name slicing
for city in city_list:
print(city[:-12])
cities_div[0].find_element_by_tag_name('a').click()
soup = bs(ch_driver.page_source, "html.parser")
results = soup.find('div', attrs={'id':'EATERY_SEARCH_RESULTS'})
rank = 1
restaurant = results.find('div', attrs={'data-index': '{rank}'.format(rank = rank)})
# +
cuisine_div = restaurant.find('div', attrs={'class' : 'cuisines'}).find_all('a')
cuisine_list = []
for div in cuisine_div:
cuisine_list.append(div.text)
cuisine = ", ".join(cuisine_list)
cuisine
# +
review_ul = restaurant.find('ul', attrs={'class' : 'review_stubs review_snippets rebrand'})
review_a = review_ul.find_all('a')
review_list = []
for tag in review_a:
review_list.append(tag.text)
review = ", ".join(review_list)
review
# -
def scraper():
result_list = []
for i in range(1, 30+1):
soup = bs(ch_driver.page_source, "html.parser")
results = soup.find('div', attrs={'id':'EATERY_SEARCH_RESULTS'})
rank = str(i)
restaurant = results.find('div', attrs={'data-index': '{rank}'.format(rank = rank)})
title_div = restaurant.find('div', attrs={'class' : 'title'})
title = title_div.find('a').text.strip('\n')
rating_div = restaurant.find('div', attrs={'class' : 'rating rebrand'})
rating = rating_div.find('span').attrs['alt']
count = restaurant.find('span', attrs={'class' : 'reviewCount'}).find('a').text.strip('\n')
try:
price = restaurant.find('span', attrs={'class' : 'item price'}).text
cuisine_div = restaurant.find('div', attrs={'class' : 'cuisines'}).find_all('a')
cuisine_list = []
for div in cuisine_div:
cuisine_list.append(div.text)
cuisine = ", ".join(cuisine_list)
except:
price = "N/A"
cuisine = "N/A"
review_ul = restaurant.find('ul', attrs={'class' : 'review_stubs review_snippets rebrand'})
review_a = review_ul.find_all('a')
review_list = []
for tag in review_a:
review_list.append(tag.text)
review = ", ".join(review_list)
result_list.append((title, rating, count, price, cuisine, review))
return result_list
scraper()[0]
| TripAdvisor_WebScraping_Practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="T_Qy0mr5TQBN"
# Montando dataframe
# + id="JuYddin9S4q-"
import pandas as pd
# + id="Q_Najr02THuJ"
base = pd.read_csv('house_prices.csv')
# + id="_aUNAdbOTTrk"
x = base.iloc[:, 3:19].values
y = base.iloc[:, 2].values
# + [markdown] id="Z2u-zV5KU5H2"
# Divisão de dataframe em teste e treinamento
# + id="vpjWQfqyTkvh"
from sklearn.model_selection import train_test_split
x_trein, x_teste, y_trein, y_teste = train_test_split(x, y, test_size=0.3, random_state=0)
# + [markdown] id="F3_Cgi70U9h7"
# Regressão Random Forest
# + id="3jereyo9Uwih" colab={"base_uri": "https://localhost:8080/"} outputId="811d1edd-a7ce-4592-b6e2-746d72b666a6"
from sklearn.tree import DecisionTreeRegressor
regressor = DecisionTreeRegressor()
regressor.fit(x_trein, y_trein)
# + colab={"base_uri": "https://localhost:8080/"} id="l9e8fZKgV3I7" outputId="75000aa7-cabe-4024-b0c0-0da780ecf994"
regressor.score(x_trein, y_trein)
# + id="42T2MqbKWMiR"
previsoes = regressor.predict(x_teste)
# + colab={"base_uri": "https://localhost:8080/"} id="kw7_5_ZAYy8E" outputId="92f11c9e-2a55-42aa-db1c-45da08786000"
from sklearn.metrics import mean_absolute_error
mae = mean_absolute_error(y_teste, previsoes)
mae
# + id="I9iXMUFHZFnK" colab={"base_uri": "https://localhost:8080/"} outputId="cd228c3c-7b3b-43f3-cf2a-ec24d3e2f9ce"
regressor.score(x_teste, y_teste)
| regressao_arvore_casa.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + id="k92C-qiliYLw"
#Import modules
# %matplotlib inline
import pandas as pd
import numpy as np
from scipy import stats
import matplotlib.pyplot as plt
import seaborn as sns
import re
# + colab={"base_uri": "https://localhost:8080/", "height": 81} id="cDBtXtXxiYL0" outputId="0d34fd9d-615e-4ece-83e6-bf5df1ad24b6"
url = "https://raw.githubusercontent.com/buzzCraft/Dave3625-21-Lab/main/Lab2/data/Titanic.csv"
df = pd.read_csv(url, sep=',')
df.head(1)
# + colab={"base_uri": "https://localhost:8080/"} id="B1c_WEgCiYL2" outputId="b2ba0893-8a85-4ef4-af68-d604c149e0ff"
df.isna().sum()
# + id="dYQR7VV5iYL2"
#Fill missing values - for this lab, lets just use teh median() function
df["Age"] = df["Age"].fillna(df["Age"].median())
df["Fare"] = df["Fare"].fillna(df["Fare"].median())
# + id="tTciJJO7iYL3" outputId="83f87cfd-5ded-40e2-e807-32da56b7678b"
#Add a new column for HasCabin
# ~ invert the bool value, so that no cabin = False
df['HasCabin'] = ~df.Cabin.isnull()
df.head()
# + id="nawyIXNziYL3"
#Lets add a new column called Title
df['Title'] = df.Name.apply(lambda x: re.search(' ([A-Z][a-z]+)\.', x).group(1))
# + colab={"base_uri": "https://localhost:8080/", "height": 309} id="WZrdC9BmiYL4" outputId="707bee1b-fdc8-4ab7-eaa8-3b273ae37b2b"
sns.countplot(x='Title', data=df);
plt.xticks(rotation=45);
# + colab={"base_uri": "https://localhost:8080/"} id="PXB_M1wETY1w" outputId="8c02c17c-52d0-4f60-8ef4-308e7c742a53"
df["Title"].value_counts()
# + id="szrjQtWLiYL5" outputId="da5c1788-6958-44f6-e288-30be22e50dd4"
# Grouping titles to
df['Title'] = df['Title'].replace({'Mlle':'Miss', 'Mme':'Mrs', 'Ms':'Miss'})
df['Title'] = df['Title'].replace(['Don', 'Dona', 'Rev', 'Dr',
'Major', 'Lady', 'Sir', 'Col', 'Capt', 'Countess', 'Jonkheer'],'Unique')
sns.countplot(x='Title', data=df);
plt.xticks(rotation=45);
# -
# Prepearing for binary data conversion
# + id="xRKJxVwziYL6" outputId="c1df5ee6-eaa6-4b9a-80c8-4b4044e1755c"
# Creating numerical columns for age and fare
df['CatAge'] = pd.qcut(df.Age, q=4, labels=False )
df['CatFare']= pd.qcut(df.Fare, q=4, labels=False)
df.head()
# + id="7vIS45p8iYL6" outputId="30c1f757-563a-42e2-8585-eba11e2abf12"
#Dropping columns that wont make sence in binary
df = df.drop(["Age", "Fare", "Cabin", "Name", "Ticket"], axis=1)
df.head()
# + id="P8SswsgtiYL8" outputId="3ce467eb-863c-4e00-8bfc-df6811df59b5"
# Converting to binary values
df_dum = pd.get_dummies(df, drop_first=True)
df_dum.head()
# + id="wUJYoWbdiYL8"
# + id="cUU2MyLeiYL9"
| Lab2/solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pyvizenv] *
# language: python
# name: conda-env-pyvizenv-py
# ---
# # Ensemble Learning
#
# ## Initial Imports
''' in this study we are analysing loan data to determine who is in risk of default their loan,
after cleaning the data set and balancig the scaled fetures, we introduce our train features to train and we take the diferent metrics
to get our results using the random forest classifier and the easy ensable classifier'''
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import pandas as pd
from pathlib import Path
from collections import Counter
from sklearn.metrics import balanced_accuracy_score
from sklearn.metrics import confusion_matrix, accuracy_score
from imblearn.metrics import classification_report_imbalanced
from sklearn.model_selection import train_test_split, cross_validate
from sklearn.preprocessing import StandardScaler
from sklearn.ensemble import RandomForestClassifier
from imblearn.ensemble import EasyEnsembleClassifier
from sklearn import preprocessing
# ## Read the CSV and Perform Basic Data Cleaning
# +
# Load the data
file_path = Path('Resources/LoanStats_2019Q1.csv')
df = pd.read_csv(file_path)
# Preview the data
df.dtypes
# -
# ## Split the Data into Training and Testing
# +
# Create our features
x_cols = [i for i in df.columns if i not in ('hardship_flag', 'debt_settlement_flag','home_ownership', 'verification_status', 'issue_d', 'pymnt_plan', 'loan_status')]
X = df[x_cols].dropna()
# Create our target
y = df['loan_status']
# -
# Check the balance of our target values
y.value_counts()
X.dtypes
# +
# Create X_train, X_test, y_train, y_test
le = preprocessing.LabelEncoder()
X = X.apply(le.fit_transform)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=1, stratify=y)
#check the number of records
print(f"the number of records in the training dataset is:", X_train.shape[0])
print(f"the number of records in the test dataset is", X_test.shape[0])
print(f"the number of records in the y_train dataset is:", y_train.shape[0])
# -
# ## Data Pre-Processing
#
# Scale the training and testing data using the `StandardScaler` from `sklearn`. Remember that when scaling the data, you only scale the features data (`X_train` and `X_testing`).
# +
# Create the StandardScaler instance
X_scaler = StandardScaler()
# +
# Fit the Standard Scaler with the training data
# When fitting scaling functions, only train on the training dataset
X_scaler.fit(X_train)
# -
# Scale the training and testing data
StandardScaler(copy=True, with_mean=True, with_std=True)
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
# ## Ensemble Learners
#
# In this section, you will compare two ensemble algorithms to determine which algorithm results in the best performance. You will train a Balanced Random Forest Classifier and an Easy Ensemble classifier . For each algorithm, be sure to complete the folliowing steps:
#
# 1. Train the model using the training data.
# 2. Calculate the balanced accuracy score from sklearn.metrics.
# 3. Display the confusion matrix from sklearn.metrics.
# 4. Generate a classication report using the `imbalanced_classification_report` from imbalanced-learn.
# 5. For the Balanced Random Forest Classifier only, print the feature importance sorted in descending order (most important feature to least important) along with the feature score
#
# Note: Use a random state of 1 for each algorithm to ensure consistency between tests
# ### Balanced Random Forest Classifier
# +
# Resample the training data with the BalancedRandomForestClassifier
rf_model = RandomForestClassifier(n_estimators=500, random_state=1)
rf_model = rf_model.fit(X_train_scaled, y_train)
#check the model performance
y_pred = rf_model.predict(X_test_scaled)
# +
# Calculated the balanced accuracy score
bac = balanced_accuracy_score(y_test, y_pred)
print(f"balance accuracy :", bac)
# +
# Display the confusion matrix
tn, fp, fn, tp = confusion_matrix(y_test, y_pred).ravel()
cm = confusion_matrix(y_test, y_pred)
cm_df = pd.DataFrame(
cm, index=["Actual 0", "Actual 1"], columns=["Predicted 0", "Predicted 1"]
)
recall = tp /(tp+fn)
# -
print("confusion matrix")
display(cm_df)
print(F"the recall| of this model is:", recall)
# Print the imbalanced classification report
print("Classification Report")
print(classification_report_imbalanced(y_test, y_pred))
# +
# Calculating the accuracy score
acc_score = accuracy_score(y_test, y_pred)
print(f"Accuracy Score : {acc_score}")
# -
# List the features sorted in descending order by feature importance
y_pred = pd.DataFrame(y_pred)
y_pred.sort_values(by=, axis=0, ascending=True)
y_pred.head()
# ### Easy Ensemble Classifier
# +
# Train the Classifier
eec = EasyEnsembleClassifier(random_state=1)
eec_model = eec.fit(X_train, y_train)
y_pred= eec_model.predict(X_test)
# +
# Calculated the balanced accuracy score
balanced_accuracy_score(y_test, y_pred)
# +
# Display the confusion matrix
confusion_matrix(y_test, y_pred)
print(F"the recall of this model is:", recall)
# +
# Print the imbalanced classification report
print(classification_report_imbalanced(y_test, y_pred))
# + active=""
# ### Final Questions
#
# 1. Which model had the best balanced accuracy score?
#
# the same recall score
#
# 2. Which model had the best recall score?
#
# the same recall score
#
# 3. Which model had the best geometric mean score?
#
# Easy Ensemble Classifier
#
# 4. What are the top three features?
#
# YOUR ANSWER HERE.
# -
| Starter_Code/credit_risk_ensemble.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: minst-ensemble
# language: python
# name: minst-ensemble
# ---
# +
import os
import sys
import importlib
import itertools
import numpy as np
import sklearn.decomposition as decomp
import seaborn as sb
# add project modules to the path
path_to_module = os.path.abspath(os.path.join(os.getcwd(), "..", "src/"))
sys.path.append(path_to_module)
import src.models.train_model as train_model
# +
path_to_data = os.path.abspath(os.path.join(os.getcwd(),
"..",
"data/processed/"
))
scaled_data = train_model.load_processed_data(file_path=path_to_data,
augmented_suffix="",
blurred_suffix=""
)
augmented_scaled_data = train_model.load_processed_data(file_path=path_to_data,
blurred_suffix=""
)
blurred_scaled_data = train_model.load_processed_data(file_path=path_to_data,
augmented_suffix=""
)
aug_blurred_scaled_data =train_model.load_processed_data(file_path=path_to_data)
# +
X_scaled = scaled_data["X_train"]
y_scaled = scaled_data["y_train"]
X_aug_scaled = augmented_scaled_data["X_train"]
y_aug_scaled = augmented_scaled_data["y_train"]
X_blur_scaled = blurred_scaled_data["X_train"]
y_blur_scaled = blurred_scaled_data["y_train"]
X_aug_blur_scaled = aug_blurred_scaled_data["X_train"]
y_aug_blur_scaled = aug_blurred_scaled_data["y_train"]
# -
pca = decomp.PCA(2)
scaled_2d = pca.fit_transform(X_scaled)
sb.scatterplot(x=scaled_2d[:,0],
y=scaled_2d[:,1],
hue=y_scaled,
palette="Set1",
size=0.1,
marker="o",
alpha=0.2,
legend="full",
edgecolor="none"
)
# there's some seperation here. especially 1s appear very bunched.
augmented_scaled_2d = pca.fit_transform(X_aug_scaled)
sb.scatterplot(x=augmented_scaled_2d[:,0],
y=augmented_scaled_2d[:,1],
hue=y_aug_scaled,
palette="Set1",
size=0.1,
marker="o",
alpha=0.2,
legend="full",
edgecolor="none"
)
# suspect the blob in the middle represents the non augmented data, and the other parts of the
blurred_scaled_2d = pca.fit_transform(X_blur_scaled)
sb.scatterplot(x=blurred_scaled_2d[:,0],
y=blurred_scaled_2d[:,1],
hue=y_blur_scaled,
palette="Set1",
size=0.1,
marker="o",
alpha=0.2,
legend="full",
edgecolor="none"
)
aug_blurred_scaled_2d = pca.fit_transform(X_aug_blur_scaled)
sb.scatterplot(x=aug_blurred_scaled_2d[:,0],
y=aug_blurred_scaled_2d[:,1],
hue=y_aug_blur_scaled,
palette="Set1",
size=0.1,
marker="o",
alpha=0.2,
legend="full",
edgecolor="none"
)
| notebooks/01-er-exploration-pca_visualisation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Convert a pandas dataframe to geojson for web-mapping
#
# Author: <NAME>
#
# Original: [pandas-to-geojson](https://github.com/gboeing/urban-data-science/blob/dc86c9c89b73f87f97301883d7456f1f814589f5/17-Leaflet-Web-Mapping/pandas-to-geojson.ipynb)
import pandas as pd, requests, json
# First download data from the city of Berkeley's API. You can use Socrata's $limit parameter to specify how many rows to grab (otherwise the default is 1,000 rows of data): https://dev.socrata.com/docs/paging.html
#
# Example request: https://data.cityofberkeley.info/resource/k489-uv4i.json?$limit=5
# API endpoint for city of Berkeley's 311 calls
endpoint_url = 'https://data.cityofberkeley.info/resource/bscu-qpbu.json?$limit=20&$where=latitude%20%3C%3E%20%22%22'
# fetch the URL and load the data
response = requests.get(endpoint_url)
data = response.json()
# Next, turn the json data into a dataframe and clean it up a bit: drop unnecessary columns and any rows that lack lat-long data. We want to make our json file as small as possible (prefer under 5 mb) so that it can be loaded over the Internet to anyone viewing your map, without taking forever to download a huge file.
# +
# turn the json data into a dataframe and see how many rows and what columns we have
df = pd.DataFrame(data)
print('We have {} rows'.format(len(df)))
str(df.columns.tolist())
# -
# convert lat-long to floats and change address from ALL CAPS to regular capitalization
df['latitude'] = df['latitude'].astype(float)
df['longitude'] = df['longitude'].astype(float)
df['street_address'] = df['street_address'].str.title()
# we don't need all those columns - only keep useful ones
cols = ['request_detail', 'request_subcategory', 'latitude', 'longitude', 'street_address', 'case_status']
df_subset = df[cols]
# +
# drop any rows that lack lat/long data
df_geo = df_subset.dropna(subset=['latitude', 'longitude'], axis=0, inplace=False)
print('We have {} geotagged rows'.format(len(df_geo)))
df_geo.tail()
# -
# what is the distribution of issue types?
df_geo['request_subcategory'].value_counts()
# Finally, convert each row in the dataframe to a geojson-formatted feature and save the result as a file. The format is pretty simple and you can see it here: http://geojson.org/
def df_to_geojson(df, properties, lat='latitude', lon='longitude'):
# create a new python dict to contain our geojson data, using geojson format
geojson = {'type':'FeatureCollection', 'features':[]}
# loop through each row in the dataframe and convert each row to geojson format
for _, row in df.iterrows():
# create a feature template to fill in
feature = {'type':'Feature',
'properties':{},
'geometry':{'type':'Point',
'coordinates':[]}}
# fill in the coordinates
feature['geometry']['coordinates'] = [row[lon],row[lat]]
# for each column, get the value and add it as a new feature property
for prop in properties:
feature['properties'][prop] = row[prop]
# add this feature (aka, converted dataframe row) to the list of features inside our dict
geojson['features'].append(feature)
return geojson
cols = [
'street_address',
'request_detail',
'request_subcategory',
'case_status'
]
geojson = df_to_geojson(df_geo, cols)
# In [nteract](https://github.com/nteract/nteract), we can display geojson directly with the built-in leaflet renderer.
import IPython
IPython.display.display({'application/geo+json': geojson}, raw=True)
# _Known temporary [issue](https://github.com/nteract/nteract/issues/2034): leaflet fails to render_
| python/pandas-to-geojson.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Matplotlib: treemap
# ======================================================================
#
# Treemaps are a nice way of showing tree information not based on a
# connected node approach.
#
# See <http://www.cs.umd.edu/hcil/treemap/>
#
# 
# +
"""
Treemap builder using pylab.
Uses algorithm straight from http://hcil.cs.umd.edu/trs/91-03/91-03.html
<NAME> 29/7/2006
"""
import pylab
from matplotlib.patches import Rectangle
class Treemap:
def __init__(self, tree, iter_method, size_method, color_method):
"""create a tree map from tree, using itermethod(node) to walk tree,
size_method(node) to get object size and color_method(node) to get its
color"""
self.ax = pylab.subplot(111,aspect='equal')
pylab.subplots_adjust(left=0, right=1, top=1, bottom=0)
self.ax.set_xticks([])
self.ax.set_yticks([])
self.size_method = size_method
self.iter_method = iter_method
self.color_method = color_method
self.addnode(tree)
def addnode(self, node, lower=[0,0], upper=[1,1], axis=0):
axis = axis % 2
self.draw_rectangle(lower, upper, node)
width = upper[axis] - lower[axis]
try:
for child in self.iter_method(node):
upper[axis] = lower[axis] + (width * float(self.size_method(child))) / self.size_method(node)
self.addnode(child, list(lower), list(upper), axis + 1)
lower[axis] = upper[axis]
except TypeError:
pass
def draw_rectangle(self, lower, upper, node):
print lower, upper
r = Rectangle( lower, upper[0]-lower[0], upper[1] - lower[1],
edgecolor='k',
facecolor= self.color_method(node))
self.ax.add_patch(r)
if __name__ == '__main__':
# example using nested lists, iter to walk and random colors
size_cache = {}
def size(thing):
if isinstance(thing, int):
return thing
if thing in size_cache:
return size_cache[thing]
else:
size_cache[thing] = reduce(int.__add__, [size(x) for x in thing])
return size_cache[thing]
import random
def random_color(thing):
return (random.random(),random.random(),random.random())
tree= ((5,(3,5)), 4, (5,2,(2,3,(3,2,2)),(3,3)), (3,2) )
Treemap(tree, iter, size, random_color)
pylab.show()
| ipython/Matplotlib_TreeMap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Tools for NOAA TMAX Record Frequency Analysis
# +
# <help>
# +
# <api>
from IPython.display import display, Image
from IPython.html.widgets import interact_manual
import matplotlib.pyplot as plt
import os
import struct
import glob
import seaborn as sns
import pandas as pd
import datetime as dt
## Use this global variable to specify the path for station summary files.
NOAA_STATION_SUMMARY_PATH = "/resources/noaa-hdta/data/derived/15mar2015/summaries/"
## Build Station List
station_detail_colnames = ['StationID','State','Name',
'Latitude','Longitude','QueryTag']
station_detail_rec_template = {'StationID': "",
'State': "",
'Name': "",
'Latitude': "",
'Longitude': "",
'QueryTag': ""
}
STATION_DETAIL = '/resources/ghcnd-stations.txt'
def get_filename(pathname):
'''Fetch filename portion of pathname.'''
plist = pathname.split('/')
fname, fext = os.path.splitext(plist[len(plist)-1])
return fname
def fetch_station_list():
station_list = []
raw_files = os.path.join(NOAA_STATION_SUMMARY_PATH,'','*_sum.csv')
for index, fname in enumerate(glob.glob(raw_files)):
f = get_filename(fname).split('_')[0]
station_list.append(str(f))
return station_list
def process_station_detail(fname,stations):
'''Return dataframe of station detail.'''
station_list = []
with open(fname,'r') as f:
lines = f.readlines()
f.close()
for line in lines:
r = noaa_gather_station_detail(line,stations)
station_list += r
return pd.DataFrame(station_list,columns=station_detail_colnames)
def noaa_gather_station_detail(line,slist):
'''Build a list of stattion tuples.'''
station_tuple_list = []
station_id_key = line[0:3]
if station_id_key == 'USC' or station_id_key == 'USW':
fields = struct.unpack('12s9s10s7s2s30s', line[0:70])
if fields[0].strip() in slist:
station_tuple = dict(station_detail_rec_template)
station_tuple['StationID'] = fields[0].strip()
station_tuple['State'] = fields[4].strip()
station_tuple['Name'] = fields[5].strip()
station_tuple['Latitude'] = fields[1].strip()
station_tuple['Longitude'] = fields[2].strip()
qt = "{0} at {1} in {2}".format(fields[0].strip(),fields[5].strip(),fields[4].strip())
station_tuple['QueryTag'] = qt
station_tuple_list.append(station_tuple)
return station_tuple_list
# Exploration Widget
month_abbrev = { 1: 'Jan', 2: 'Feb', 3: 'Mar', 4: 'Apr',
5: 'May', 6: 'Jun', 7: 'Jul', 8: 'Aug',
9: 'Sep', 10: 'Oct', 11: 'Nov', 12: 'Dec'
}
def compute_years_of_station_data(df):
yrs = dt.date.today().year-min(df['FirstYearOfRecord'])
print("This weather station has been in service and collecting data for {0} years.").format(yrs)
return
def compute_tmax_record_quantity(df,freq):
threshold = int(freq)
df_result = df.query('(TMaxRecordCount > @threshold)')
return df_result
def fetch_station_data(stationid):
fname = os.path.join(NOAA_STATION_SUMMARY_PATH,'',stationid+'_sum.csv')
return pd.DataFrame.from_csv(fname)
def create_day_identifier(month,day):
return str(day)+'-'+month_abbrev[int(month)]
def create_date_list(mlist,dlist):
mv = mlist.values()
dv = dlist.values()
new_list = []
for index, value in enumerate(mv):
new_list.append(create_day_identifier(value,dv[index]))
return new_list
def create_record_date_list(mlist,dlist,ylist):
mv = mlist.values()
dv = dlist.values()
yv = ylist.values()
new_list = []
for index, value in enumerate(mv):
new_list.append(dt.date(yv[index],value,dv[index]))
return new_list
def compute_max_record_durations(df):
dates = create_date_list(df['Month'].to_dict(),df['Day'].to_dict())
s_dates = pd.Series(dates)
s_values = pd.Series(df['MaxDurTMaxRecord'].to_dict().values())
df_new = pd.concat([s_dates, s_values], axis=1)
df_new.columns = {"Duration","Date"}
return df_new
def plot_tmax_record_results(df):
dates = create_record_date_list(df['Month'].to_dict(),
df['Day'].to_dict(),
df['TMaxRecordYear'].to_dict()
)
s_dates = pd.Series(dates)
s_tempvalues = pd.Series(df['TMax'].to_dict().values())
df_new = pd.concat([s_dates,s_tempvalues], axis=1)
df_new.columns = {"RecordDate","RecordHighTemp"}
plt.figure(figsize = (9,9), dpi = 72)
plt.xticks(rotation=90)
sns.pointplot(df_new["RecordDate"],df_new["RecordHighTemp"])
return df_new
def plot_duration_results(df):
plt.figure(figsize = (9,9), dpi = 72)
plt.xlabel('Day')
plt.ylabel('Record Duration in Years')
plt.title('Maximum Duration for TMax Records')
ax = plt.gca()
colors= ['r', 'b']
df.plot(kind='bar',color=colors, alpha=0.75, ax=ax)
ax.xaxis.set_ticklabels( ['%s' % i for i in df.Date.values] )
plt.grid(b=True, which='major', linewidth=1.0)
plt.grid(b=True, which='minor')
return
def explore_tmaxfreq(station,hirec):
df_station_detail = fetch_station_data(station)
df_station_address_detail = process_station_detail(STATION_DETAIL,fetch_station_list())
df_station_name = df_station_address_detail.query("(StationID == @station)")
qt = df_station_name.iloc[0]["QueryTag"]
print("Historical high temperature record analysis for weather station {0}.").format(qt)
display(df_station_name)
print("Station detail, quick glimpse.")
display(df_station_detail.head())
compute_years_of_station_data(df_station_detail)
df_record_days = compute_tmax_record_quantity(df_station_detail,hirec)
if not df_record_days.empty:
print("This station has experienced {0} days of new record highs where a new record has been set more than {1} times throughout the operation of the station.").format(len(df_record_days),hirec)
display(df_record_days.head(10))
print("Displayed above are the details for up to the first 10 new record high events. All records are ploted below.")
df_rec_results = plot_tmax_record_results(df_record_days)
df_durations = compute_max_record_durations(df_record_days)
plot_duration_results(df_durations)
else:
print("This weather station has not experienced any days with greater than {0} new record highs.").format(hirec)
return
def noaaquery(renderer=lambda station,hirec : explore_tmaxfreq(station,hirec)):
'''
Creates an interactive query widget with an optional custom renderer.
station: Weather Station ID
hirec: Query indicator for the maximum number of TMAX records for a given day.
'''
df_station_detail = process_station_detail(STATION_DETAIL,fetch_station_list())
station_vals = tuple(df_station_detail.StationID)
hirec_vals = tuple(map(str, range(1,51)))
@interact_manual(station=station_vals, hirec=hirec_vals)
def noaaquery(station, hirec):
'''Inner function that gets called when the user interacts with the widgets.'''
try:
station_id = station.strip()
high_rec_freq = hirec
except Exception as e:
print("Widget Error: {0}").format(e.message)
renderer(station_id, high_rec_freq)
| noaa/tmaxfreq/noaaquery_tmaxfreq_tools.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## part 1 ##
import numpy as np
testlines = '''7,4,9,5,11,17,23,2,0,14,21,24,10,16,13,6,15,25,12,22,18,20,8,19,3,26,1
22 13 17 11 0
8 2 23 4 24
21 9 14 16 7
6 10 3 18 5
1 12 20 15 19
3 15 0 2 22
9 18 13 17 5
19 8 7 25 23
20 11 10 24 4
14 21 16 12 6
14 21 17 24 4
10 16 15 9 19
18 8 23 26 20
22 11 13 6 5
2 0 12 3 7'''.split('\n')
testlines[-1]
with open('day4.txt') as fp:
puzzlelines = fp.read().split('\n')[:-1]
puzzlelines[-1]
def parse_lines(lines):
draws = [int(n) for n in lines[0].split(',')]
size = len(lines[2].split())
num_cards = (len(lines) - 1) // (size + 1)
cards = []
for n in range(num_cards):
card = np.zeros((size, size), int)
first_line = n*(size + 1) + 2
for i, line in enumerate(lines[first_line:first_line+size]):
card[i, :] = [int(s) for s in line.split()]
cards.append(card)
return draws, size, cards
test_draws, test_size, test_cards = parse_lines(testlines)
puzzle_draws, puzzle_size, puzzle_cards = parse_lines(puzzlelines)
def check_card(card, drawn):
size = card[0,:].size
for row in range(size):
if all(i in drawn for i in card[row,:]):
return True
for col in range(size):
if all(j in drawn for j in card[:, col]):
return True
return False
def score(card, drawn):
return sum(i for i in np.nditer(card) if i not in drawn)*drawn[-1]
def process(draws, cards):
curr = []
for num in draws:
curr.append(num)
for card in cards:
if check_card(card, curr):
return score(card, curr)
return None
process(test_draws, test_cards)
process(puzzle_draws, puzzle_cards)
# ## part 2 ##
def process2(draws, cards):
curr = []
winning_cards = []
for num in draws:
curr.append(num)
for i, card in enumerate(cards):
if i in winning_cards:
continue
if check_card(card, curr):
winning_cards.append(i)
if len(winning_cards) == len(cards):
return score(card, curr)
return None
process2(test_draws, test_cards)
process2(puzzle_draws, puzzle_cards)
| day4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Geocoding and Network Analysis Demo
# ## What is the Network Analysis?
#
# The ArcGIS Network Analyst allows you to solve common `network` problems
#
#
#
# ### Examples:
#
# - finding the best route across a city
# - finding the closest emergency vehicle or facility,
# - identifying a service area around a location
# - servicing a set of orders with a fleet of vehicles
# - choosing the best facilities to open or close.
# # Constructing drive time based service areas
# This sample shows how the `network` module of the ArcGIS API for Python can be used to construct service areas. In this sample, we generate service areas for two of the fire stations in central Tokyo, Japan. We later observe how the service area varies by time of day for a fire station in the city of Los Angeles.
# + [markdown] heading_collapsed=true
# ##### Connect to the GIS
# + hidden=true
from datetime import datetime
from IPython.display import HTML
import pandas as pd
from arcgis.gis import GIS
my_gis = GIS(profile='agol_profile')
# -
# ### Create a Network Layer
from arcgis.network import ServiceAreaLayer
service_area_url = my_gis.properties.helperServices.serviceArea.url
service_area_url
sa_layer = ServiceAreaLayer(service_area_url, gis=my_gis)
# ## Calculate service area from the Resort
from arcgis.geocoding import geocode
ps_geocode = geocode('Hilton, Berlin, DE', as_featureset=True)
sdf = ps_geocode.sdf
sdf.head()
# # Display the Hilton Hotels in Berlin
map1 = my_gis.map()
map1.zoomLevel = 10
map1.extent = sdf.spatial.full_extent
map1
ps_geocode.sdf.spatial.plot(map_widget=map1)
# ## Compute the service area
#
# +
travel_modes = sa_layer.retrieve_travel_modes()
for t in travel_modes['supportedTravelModes']:
print(t['name'])
# -
truck_mode = [t for t in travel_modes['supportedTravelModes'] if t['name'] == 'Driving Time'][0]
sdf = ps_geocode.sdf
s = sdf.spatial
s.to_feature_collection()
from arcgis.features import analysis
drive_time_job = analysis.create_drive_time_areas(s.to_feature_collection(),
break_values=[5, 10, 15],
travel_mode=truck_mode,
future=True)
drive_time_job
# ## Checking Job Status
drive_time_job.status
# ## Get the Job Result
fc = drive_time_job.result()
fc
# ### Visualize the service area on the map
#
# Visualize the results on the map widget.
#
map1.add_layer(fc)
# Click the drive time areas to explore their attributes. Because the content of the pop-ups may include HTML source code, it is also possible to have the pop-up windows include other resources such as tables and images.
# ### Driving directions
from arcgis.network import RouteLayer
route_service_url = my_gis.properties.helperServices.route.url
route_service_url
route_service = RouteLayer(route_service_url, gis=my_gis)
route_service
stops = [ 'Zufahrt zum Flughafen Tegel, 13405 Berlin, Germany', 'Mohrenstraße 30, 10117 Berlin, Germany']
from arcgis.geocoding import geocode, batch_geocode
stops_geocoded = batch_geocode(stops)
# #### data wrangling
stops_geocoded = [item['location'] for item in stops_geocoded]
stops_geocoded
stops_geocoded2 = '{},{};{},{}'.format(stops_geocoded[0]['x'],stops_geocoded[0]['y'],
stops_geocoded[1]['x'],stops_geocoded[1]['y'])
stops_geocoded2
# ### Driving directions
result = route_service.solve(stops_geocoded2,
return_routes=True,
return_stops=True,
return_directions=True,
return_barriers=False,
return_polygon_barriers=False,
return_polyline_barriers=False)
ext = {'spatialReference': {'latestWkid': 4326},
'xmin': 13.309110351000072,
'ymin': 52.55763923600006,
'xmax': 13.392801147000057,
'ymax': 52.51226653600003}
# +
map2 = my_gis.map('Berlin, Germany', zoomlevel=12)
map2
# -
from arcgis.features import Feature, FeatureSet
features = result['routes']['features']
routes = FeatureSet(features)
stop_features = result['stops']['features']
stop_fset = FeatureSet(stop_features)
len(result['stops'])
stop_features = result['stops']['features']
stop_fset = FeatureSet(stop_features)
ln_symbol = { "type": "simple-line", "width": 2, "color": [255, 0, 0, 1] }
symbol = {
"type" : "simple-marker",
"style" : "square",
"outline" : {
"style": "dash-dot"
},
"color": [255, 211, 127, 1]
}
map2.draw(routes, symbol=ln_symbol)
map2.draw(stop_fset, symbol=symbol)
# # Visualizing the Instructions
import pandas as pd
text = [att['attributes'] for att in result['directions'][0]['features']]
df = pd.DataFrame(text)[['text', 'length','time']]
df
| 04. Performing Analysis with the Python API/06 - network/07 - networks-service-areas-driving-directions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/vigneshwaran-dev/CV-research-timeline/blob/main/GoogLeNet/notebook.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="shQIYIt6JbcA"
import tensorflow as tf
from tensorflow.keras import Model
from tensorflow.keras.layers import Dense, Activation, Dropout, Flatten, Conv2D, MaxPooling2D, AveragePooling2D, Lambda, Input, GlobalAveragePooling2D
from tensorflow.keras.losses import categorical_crossentropy
from tensorflow.nn import local_response_normalization
from tensorflow.keras.optimizers import SGD
from tensorflow.keras.callbacks import LearningRateScheduler
# + [markdown] id="nMVPxhNGl-MI"
# Defining the Model as per the Original Paper
# + id="wNHRgxhVKRJ_"
def inception_block(model, filter_map):
branch_0 = Conv2D(filters=filter_map[0], kernel_size=(1, 1), padding='same')(model)
branch_0 = Activation('relu')(branch_0)
branch_1 = Conv2D(filters=filter_map[1][0], kernel_size=(1, 1), padding='same')(model)
branch_1 = Activation('relu')(branch_1)
branch_1 = Conv2D(filters=filter_map[1][1], kernel_size=(1, 1), padding='same')(branch_1)
branch_1 = Activation('relu')(branch_1)
branch_2 = Conv2D(filters=filter_map[2][0], kernel_size=(1, 1), padding='same')(model)
branch_2 = Activation('relu')(branch_2)
branch_2 = Conv2D(filters=filter_map[2][1], kernel_size=(1, 1), padding='same')(branch_2)
branch_2 = Activation('relu')(branch_2)
branch_3 = MaxPooling2D(pool_size=(3, 3), strides=(1, 1), padding='same')(model)
branch_3 = Conv2D(filters=filter_map[3], kernel_size=(1, 1), padding='same')(branch_3)
branch_3 = Activation('relu')(branch_3)
return tf.concat([branch_0, branch_1, branch_2, branch_3], axis=3)
# Initial Convolutional Block
_input = Input(shape=(224, 224, 3))
model = Conv2D(filters=64, kernel_size=(7, 7), strides=(2, 2), padding='same')(_input)
model = Activation('relu')(model)
model = MaxPooling2D(pool_size=(3, 3), strides=(2, 2))(model)
model = Lambda(local_response_normalization)(model)
model = Conv2D(filters=64, kernel_size=(1, 1), strides=(1, 1), padding='same')(model)
model = Activation('relu')(model)
model = Conv2D(filters=192, kernel_size=(3, 3), strides=(1, 1), padding='same')(model)
model = Activation('relu')(model)
model = Lambda(local_response_normalization)(model)
model = MaxPooling2D(pool_size=(3, 3), strides=(2, 2))(model)
# 1st Sequence of Inceptions
model = inception_block(model, [64, [96, 128], [16, 32], 32])
model = inception_block(model, [128, [128, 192], [32, 96], 64])
model = MaxPooling2D(pool_size=(3, 3), strides=(2, 2))(model)
model = inception_block(model, [192, [96, 208], [16, 48], 64])
# 1st Output Branch
output_1 = AveragePooling2D(pool_size=(5, 5), strides=(3, 3))(model)
output_1 = Conv2D(filters=128, kernel_size=(1, 1), strides=(1, 1), padding='same')(output_1)
output_1 = Activation('relu')(output_1)
output_1 = Flatten()(output_1)
output_1 = Dense(1024)(output_1)
output_1 = Activation('relu')(output_1)
output_1 = Dropout(0.7)(output_1)
output_1 = Dense(1000)(output_1)
output_1 = Activation('softmax')(output_1)
# 2nd Sequence of Inceptions
model = inception_block(model, [160, [112, 224], [24, 64], 64])
model = inception_block(model, [128, [128, 256], [24, 64], 64])
model = inception_block(model, [112, [144, 288], [32, 64], 64])
# 2nd Output Branch
output_2 = AveragePooling2D(pool_size=(5, 5), strides=(3, 3))(model)
output_2 = Conv2D(filters=128, kernel_size=(1, 1), strides=(1, 1), padding='same')(output_2)
output_2 = Activation('relu')(output_2)
output_2 = Flatten()(output_2)
output_2 = Dense(1024)(output_2)
output_2 = Activation('relu')(output_2)
output_2 = Dropout(0.7)(output_2)
output_2 = Dense(1000)(output_2)
output_2 = Activation('softmax')(output_2)
# 3rd Sequence of Inceptions
model = inception_block(model, [256, [160, 320], [32, 128], 128])
model = MaxPooling2D(pool_size=(3, 3), strides=(2, 2))(model)
model = inception_block(model, [256, [160, 320], [32, 128], 128])
model = inception_block(model, [384, [192, 384], [48, 128], 128])
# Final Output
model = GlobalAveragePooling2D()(model)
model = Dropout(0.4)(model)
model = Dense(1000)(model)
model = Activation('softmax')(model)
model = Model(inputs=_input, outputs=[model, output_1, output_2])
# + colab={"base_uri": "https://localhost:8080/"} id="dgmKxaerN-OP" outputId="7f9e5acf-9f56-470b-ecfc-16cb8c935a01"
model.summary()
# + id="mPJ6al6pLn5k"
model.compile(loss=[categorical_crossentropy, categorical_crossentropy, categorical_crossentropy],
optimizer=SGD(learning_rate=0.01, momentum=0.9),
metrics=['accuracy'])
# + [markdown] id="etzsyGSKmGn7"
# Considering the data to be present in TRAIN_DATA_LOCATION and VALIDATION_DATA_LOCATION directories and running them through data generators to perform live data augumentation during the training process
# + id="BSvuN4XSdzNI"
from tensorflow.keras.preprocessing.image import ImageDataGenerator
train_dir = 'TRAIN_DATA_LOCATION'
valid_dir = 'VALIDATION_DATA_LOCATION'
BATCH_SIZE = 32
train_datagen = ImageDataGenerator(rescale=1./255,
rotation_range=10,
width_shift_range=0.1,
height_shift_range=0.1,
shear_range=0.1,
zoom_range=0.1)
train_generator = train_datagen.flow_from_directory(train_dir,
target_size=(224, 224),
color_mode='rgb',
batch_size=BATCH_SIZE,
seed=1,
shuffle=True,
class_mode='categorical')
valid_datagen = ImageDataGenerator(rescale=1.0/255.0)
valid_generator = valid_datagen.flow_from_directory(valid_dir,
target_size=(224, 224),
color_mode='rgb',
batch_size=BATCH_SIZE,
seed=7,
shuffle=True,
class_mode='categorical')
train_num = train_generator.samples
# + [markdown] id="vCkprKt6mbm3"
# Training the Model
# + id="QlUCsMp-kzXJ"
import datetime
log_dir = 'logs/fit/' + datetime.datetime.now().strftime('%Y%m%d-%H%M%S')
tensorboard_callback = tf.keras.callbacks.TensorBoard(log_dir=log_dir)
callback_list = [tensorboard_callback]
def reduce_lr(epoch):
lr = 0.01
reduction = epoch % 8
for _ in range(1, reduction):
lr -= (lr / 100) * 80
return lr
callback_list.append(LearningRateScheduler(reduce_lr))
model.fit(train_generator,
epochs=1,
steps_per_epoch=train_num // BATCH_SIZE,
validation_data=valid_generator,
validation_steps=valid_num // BATCH_SIZE,
callbacks=callback_list,
verbose=1)
model.save('googlenet.h5')
# + [markdown] id="5Bj7xKU4md-y"
# Visualizing the performance using Tensorboard
# + id="W6pvAAeylQiC"
# %load_ext tensorboard
# %tensorboard --logdir logs/fit
# + [markdown] id="QFxjcbSdmjtK"
# Prediction
# + id="MEhIFtiEmjMl"
x_valid, label_batch = next(iter(valid_generator))
prediction_values = model.predict_classes(x_valid)
print(prediction_values)
| GoogLeNet/notebook.ipynb |